code
stringlengths 38
801k
| repo_path
stringlengths 6
263
|
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# +
import tensorflow as tf
import keras
import keras.backend as K
from scipy.signal import resample
from sklearn.utils import shuffle
from sklearn.metrics import classification_report, confusion_matrix, accuracy_score, f1_score
from collections import Counter
from keras import regularizers
from keras.models import Sequential, Model, load_model, model_from_json
from keras.utils import to_categorical
from keras.layers import Input, Dense, Flatten, Reshape, Concatenate, Dropout
from keras.layers import Conv2D, MaxPooling2D, UpSampling2D, Conv2DTranspose
from keras.layers.normalization import BatchNormalization
from keras.callbacks import ModelCheckpoint
from keras.utils import np_utils
from keras.layers.advanced_activations import LeakyReLU
def get_class_weights(y):
counter = Counter(y)
majority = max(counter.values())
return {cls: float(majority/count) for cls, count in counter.items()}
class Estimator:
l2p = 0.001
@staticmethod
def early_layers(inp, fm = (1,3), hid_act_func="relu"):
# Start
x = Conv2D(32, fm, padding="same", kernel_regularizer=regularizers.l2(Estimator.l2p), activation=hid_act_func)(inp)
x = BatchNormalization()(x)
x = MaxPooling2D(pool_size=(1, 2))(x)
x = Dropout(0.25)(x)
# 1
x = Conv2D(32, fm, padding="same", kernel_regularizer=regularizers.l2(Estimator.l2p), activation=hid_act_func)(x)
x = BatchNormalization()(x)
x = MaxPooling2D(pool_size=(1, 2))(x)
x = Dropout(0.25)(x)
return x
@staticmethod
def late_layers(inp, num_classes, fm = (1,3), act_func="softmax", hid_act_func="relu", b_name="Identifier"):
# 2
x = Conv2D(32, fm, padding="same", kernel_regularizer=regularizers.l2(Estimator.l2p), activation=hid_act_func)(inp)
x = BatchNormalization()(x)
x = MaxPooling2D(pool_size=(1, 2))(x)
x = Dropout(0.25)(x)
# End
x = Flatten()(x)
x = Dense(128, kernel_regularizer=regularizers.l2(Estimator.l2p), activation=hid_act_func)(x)
x = BatchNormalization()(x)
x = Dropout(0.5)(x)
x = Dense(32, kernel_regularizer=regularizers.l2(Estimator.l2p), activation=hid_act_func)(x)
x = BatchNormalization()(x)
x = Dropout(0.5)(x)
x = Dense(num_classes, activation=act_func, name = b_name)(x)
return x
@staticmethod
def build(height, width, num_classes, name, fm = (1,3), act_func="softmax",hid_act_func="relu"):
inp = Input(shape=(height, width, 1))
early = Estimator.early_layers(inp, fm, hid_act_func=hid_act_func)
late = Estimator.late_layers(early, num_classes, fm, act_func=act_func, hid_act_func=hid_act_func)
model = Model(inputs=inp, outputs=late ,name=name)
return model
# +
import numpy as np
import pandas as pd
from pandas.plotting import autocorrelation_plot
import matplotlib.pyplot as plt
def get_ds_infos():
"""
Read the file includes data subject information.
Data Columns:
0: code [1-24]
1: weight [kg]
2: height [cm]
3: age [years]
4: gender [0:Female, 1:Male]
Returns:
A pandas DataFrame that contains inforamtion about data subjects' attributes
"""
dss = pd.read_csv("data_subjects_info.csv")
print("[INFO] -- Data subjects' information is imported.")
return dss
def set_data_types(data_types=["userAcceleration"]):
"""
Select the sensors and the mode to shape the final dataset.
Args:
data_types: A list of sensor data type from this list: [attitude, gravity, rotationRate, userAcceleration]
Returns:
It returns a list of columns to use for creating time-series from files.
"""
dt_list = []
for t in data_types:
if t != "attitude":
dt_list.append([t+".x",t+".y",t+".z"])
else:
dt_list.append([t+".roll", t+".pitch", t+".yaw"])
return dt_list
def creat_time_series(dt_list, act_labels, trial_codes, mode="mag", labeled=True, combine_grav_acc=False):
"""
Args:
dt_list: A list of columns that shows the type of data we want.
act_labels: list of activites
trial_codes: list of trials
mode: It can be "raw" which means you want raw data
for every dimention of each data type,
[attitude(roll, pitch, yaw); gravity(x, y, z); rotationRate(x, y, z); userAcceleration(x,y,z)].
or it can be "mag" which means you only want the magnitude for each data type: (x^2+y^2+z^2)^(1/2)
labeled: True, if we want a labeld dataset. False, if we only want sensor values.
combine_grav_acc: True, means adding each axis of gravity to corresponding axis of userAcceleration.
Returns:
It returns a time-series of sensor data.
"""
num_data_cols = len(dt_list) if mode == "mag" else len(dt_list*3)
if labeled:
dataset = np.zeros((0,num_data_cols+7)) # "7" --> [act, code, weight, height, age, gender, trial]
else:
dataset = np.zeros((0,num_data_cols))
ds_list = get_ds_infos()
print("[INFO] -- Creating Time-Series")
for sub_id in ds_list["code"]:
for act_id, act in enumerate(act_labels):
for trial in trial_codes[act_id]:
fname = 'A_DeviceMotion_data/'+act+'_'+str(trial)+'/sub_'+str(int(sub_id))+'.csv'
raw_data = pd.read_csv(fname)
raw_data = raw_data.drop(['Unnamed: 0'], axis=1)
vals = np.zeros((len(raw_data), num_data_cols))
if combine_grav_acc:
raw_data["userAcceleration.x"] = raw_data["userAcceleration.x"].add(raw_data["gravity.x"])
raw_data["userAcceleration.y"] = raw_data["userAcceleration.y"].add(raw_data["gravity.y"])
raw_data["userAcceleration.z"] = raw_data["userAcceleration.z"].add(raw_data["gravity.z"])
for x_id, axes in enumerate(dt_list):
if mode == "mag":
vals[:,x_id] = (raw_data[axes]**2).sum(axis=1)**0.5
else:
vals[:,x_id*3:(x_id+1)*3] = raw_data[axes].values
vals = vals[:,:num_data_cols]
if labeled:
lbls = np.array([[act_id,
sub_id-1,
ds_list["weight"][sub_id-1],
ds_list["height"][sub_id-1],
ds_list["age"][sub_id-1],
ds_list["gender"][sub_id-1],
trial
]]*len(raw_data))
vals = np.concatenate((vals, lbls), axis=1)
dataset = np.append(dataset,vals, axis=0)
cols = []
for axes in dt_list:
if mode == "raw":
cols += axes
else:
cols += [str(axes[0][:-2])]
if labeled:
cols += ["act", "id", "weight", "height", "age", "gender", "trial"]
dataset = pd.DataFrame(data=dataset, columns=cols)
return dataset
#________________________________
#________________________________
def ts_to_secs(dataset, w, s, standardize = False, **options):
data = dataset[dataset.columns[:-7]].values
act_labels = dataset["act"].values
id_labels = dataset["id"].values
trial_labels = dataset["trial"].values
mean = 0
std = 1
if standardize:
## Standardize each sensor’s data to have a zero mean and unity standard deviation.
## As usual, we normalize test dataset by training dataset's parameters
if options:
mean = options.get("mean")
std = options.get("std")
print("[INFO] -- Test Data has been standardized")
else:
mean = data.mean(axis=0)
std = data.std(axis=0)
print("[INFO] -- Training Data has been standardized: the mean is = "+str(mean)+" ; and the std is = "+str(std))
data -= mean
data /= std
else:
print("[INFO] -- Without Standardization.....")
## We want the Rows of matrices show each Feature and the Columns show time points.
data = data.T
m = data.shape[0] # Data Dimension
ttp = data.shape[1] # Total Time Points
number_of_secs = int(round(((ttp - w)/s)))
## Create a 3D matrix for Storing Sections
secs_data = np.zeros((number_of_secs , m , w ))
act_secs_labels = np.zeros(number_of_secs)
id_secs_labels = np.zeros(number_of_secs)
k=0
for i in range(0 , ttp-w, s):
j = i // s
if j >= number_of_secs:
break
if id_labels[i] != id_labels[i+w-1]:
continue
if act_labels[i] != act_labels[i+w-1]:
continue
if trial_labels[i] != trial_labels[i+w-1]:
continue
secs_data[k] = data[:, i:i+w]
act_secs_labels[k] = act_labels[i].astype(int)
id_secs_labels[k] = id_labels[i].astype(int)
k = k+1
secs_data = secs_data[0:k]
act_secs_labels = act_secs_labels[0:k]
id_secs_labels = id_secs_labels[0:k]
return secs_data, act_secs_labels, id_secs_labels, mean, std
##________________________________________________________________
ACT_LABELS = ["dws","ups", "wlk", "jog", "std", "sit"]
TRIAL_CODES = {
ACT_LABELS[0]:[1,2,11],
ACT_LABELS[1]:[3,4,12],
ACT_LABELS[2]:[7,8,15],
ACT_LABELS[3]:[9,16],
ACT_LABELS[4]:[6,14],
ACT_LABELS[5]:[5,13],
}
# -
class SSA(object):
__supported_types = (pd.Series, np.ndarray, list)
def __init__(self, tseries, L, save_mem=True):
"""
Decomposes the given time series with a singular-spectrum analysis. Assumes the values of the time series are
recorded at equal intervals.
Parameters
----------
tseries : The original time series, in the form of a Pandas Series, NumPy array or list.
L : The window length. Must be an integer 2 <= L <= N/2, where N is the length of the time series.
save_mem : Conserve memory by not retaining the elementary matrices. Recommended for long time series with
thousands of values. Defaults to True.
Note: Even if an NumPy array or list is used for the initial time series, all time series returned will be
in the form of a Pandas Series or DataFrame object.
"""
# Tedious type-checking for the initial time series
if not isinstance(tseries, self.__supported_types):
raise TypeError("Unsupported time series object. Try Pandas Series, NumPy array or list.")
# Checks to save us from ourselves
self.N = len(tseries)
if not 2 <= L <= self.N/2:
raise ValueError("The window length must be in the interval [2, N/2].")
self.L = L
self.orig_TS = pd.Series(tseries)
self.K = self.N - self.L + 1
# Embed the time series in a trajectory matrix
self.X = np.array([self.orig_TS.values[i:L+i] for i in range(0, self.K)]).T
# Decompose the trajectory matrix
self.U, self.Sigma, VT = np.linalg.svd(self.X)
self.d = np.linalg.matrix_rank(self.X)
self.TS_comps = np.zeros((self.N, self.d))
if not save_mem:
# Construct and save all the elementary matrices
self.X_elem = np.array([ self.Sigma[i]*np.outer(self.U[:,i], VT[i,:]) for i in range(self.d) ])
# Diagonally average the elementary matrices, store them as columns in array.
for i in range(self.d):
X_rev = self.X_elem[i, ::-1]
self.TS_comps[:,i] = [X_rev.diagonal(j).mean() for j in range(-X_rev.shape[0]+1, X_rev.shape[1])]
self.V = VT.T
else:
# Reconstruct the elementary matrices without storing them
for i in range(self.d):
X_elem = self.Sigma[i]*np.outer(self.U[:,i], VT[i,:])
X_rev = X_elem[::-1]
self.TS_comps[:,i] = [X_rev.diagonal(j).mean() for j in range(-X_rev.shape[0]+1, X_rev.shape[1])]
self.X_elem = "Re-run with save_mem=False to retain the elementary matrices."
# The V array may also be very large under these circumstances, so we won't keep it.
self.V = "Re-run with save_mem=False to retain the V matrix."
# Calculate the w-correlation matrix.
self.calc_wcorr()
def components_to_df(self, n=0):
"""
Returns all the time series components in a single Pandas DataFrame object.
"""
if n > 0:
n = min(n, self.d)
else:
n = self.d
# Create list of columns - call them F0, F1, F2, ...
cols = ["F{}".format(i) for i in range(n)]
return pd.DataFrame(self.TS_comps[:, :n], columns=cols, index=self.orig_TS.index)
def reconstruct(self, indices):
"""
Reconstructs the time series from its elementary components, using the given indices. Returns a Pandas Series
object with the reconstructed time series.
Parameters
----------
indices: An integer, list of integers or slice(n,m) object, representing the elementary components to sum.
"""
if isinstance(indices, int): indices = [indices]
ts_vals = self.TS_comps[:,indices].sum(axis=1)
return pd.Series(ts_vals, index=self.orig_TS.index)
def calc_wcorr(self):
"""
Calculates the w-correlation matrix for the time series.
"""
# Calculate the weights
w = np.array(list(np.arange(self.L)+1) + [self.L]*(self.K-self.L-1) + list(np.arange(self.L)+1)[::-1])
def w_inner(F_i, F_j):
return w.dot(F_i*F_j)
# Calculated weighted norms, ||F_i||_w, then invert.
F_wnorms = np.array([w_inner(self.TS_comps[:,i], self.TS_comps[:,i]) for i in range(self.d)])
F_wnorms = F_wnorms**-0.5
# Calculate Wcorr.
self.Wcorr = np.identity(self.d)
for i in range(self.d):
for j in range(i+1,self.d):
self.Wcorr[i,j] = abs(w_inner(self.TS_comps[:,i], self.TS_comps[:,j]) * F_wnorms[i] * F_wnorms[j])
self.Wcorr[j,i] = self.Wcorr[i,j]
def plot_wcorr(self, min=None, max=None):
"""
Plots the w-correlation matrix for the decomposed time series.
"""
if min is None:
min = 0
if max is None:
max = self.d
if self.Wcorr is None:
self.calc_wcorr()
ax = plt.imshow(self.Wcorr,interpolation = 'none')
plt.xlabel(r"$\tilde{F}_i$")
plt.ylabel(r"$\tilde{F}_j$")
plt.colorbar(ax.colorbar, fraction=0.045)
ax.colorbar.set_label("$W_{i,j}$")
plt.clim(0,1)
# For plotting purposes:
if max == self.d:
max_rnge = self.d-1
else:
max_rnge = max
plt.xlim(min-0.5, max_rnge+0.5)
plt.ylim(max_rnge+0.5, min-0.5)
#https://stackoverflow.com/a/45305384/5210098
def f1_metric(y_true, y_pred):
def recall(y_true, y_pred):
"""Recall metric.
Only computes a batch-wise average of recall.
Computes the recall, a metric for multi-label classification of
how many relevant items are selected.
"""
true_positives = K.sum(K.round(K.clip(y_true * y_pred, 0, 1)))
possible_positives = K.sum(K.round(K.clip(y_true, 0, 1)))
recall = true_positives / (possible_positives + K.epsilon())
return recall
def precision(y_true, y_pred):
"""Precision metric.
Only computes a batch-wise average of precision.
Computes the precision, a metric for multi-label classification of
how many selected items are relevant.
"""
true_positives = K.sum(K.round(K.clip(y_true * y_pred, 0, 1)))
predicted_positives = K.sum(K.round(K.clip(y_pred, 0, 1)))
precision = true_positives / (predicted_positives + K.epsilon())
return precision
precision = precision(y_true, y_pred)
recall = recall(y_true, y_pred)
return 2*((precision*recall)/(precision+recall+K.epsilon()))
# +
## Here we set parameter to build labeld time-series from dataset of "(A)DeviceMotion_data"
## attitude(roll, pitch, yaw); gravity(x, y, z); rotationRate(x, y, z); userAcceleration(x,y,z)
results ={}
sdt = ["rotationRate","userAcceleration"]
mode = "mag"
cga = True # Add gravity to acceleration or not
print("[INFO] -- Selected sensor data types: "+str(sdt)+" -- Mode: "+str(mode)+" -- Grav+Acc: "+str(cga))
act_labels = ACT_LABELS [0:4]
print("[INFO] -- Selected activites: "+str(act_labels))
trial_codes = [TRIAL_CODES[act] for act in act_labels]
dt_list = set_data_types(sdt)
dataset = creat_time_series(dt_list, act_labels, trial_codes, mode=mode, labeled=True, combine_grav_acc = cga)
print("[INFO] -- Shape of time-Series dataset:"+str(dataset.shape))
#*****************
TRAIN_TEST_TYPE = "subject" # "subject" or "trial"
#*****************
if TRAIN_TEST_TYPE == "subject":
test_ids = [4,9,11,21]
print("[INFO] -- Test IDs: "+str(test_ids))
test_ts = dataset.loc[(dataset['id'].isin(test_ids))]
train_ts = dataset.loc[~(dataset['id'].isin(test_ids))]
else:
test_trail = [11,12,13,14,15,16]
print("[INFO] -- Test Trials: "+str(test_trail))
test_ts = dataset.loc[(dataset['trial'].isin(test_trail))]
train_ts = dataset.loc[~(dataset['trial'].isin(test_trail))]
print("[INFO] -- Shape of Train Time-Series :"+str(train_ts.shape))
print("[INFO] -- Shape of Test Time-Series :"+str(test_ts.shape))
print("___________Train_VAL____________")
val_trail = [11,12,13,14,15,16]
val_ts = train_ts.loc[(train_ts['trial'].isin(val_trail))]
train_ts = train_ts.loc[~(train_ts['trial'].isin(val_trail))]
print("[INFO] -- Training Time-Series :"+str(train_ts.shape))
print("[INFO] -- Validation Time-Series :"+str(val_ts.shape))
print("___________________________________________________")
print(train_ts.head())
## This Variable Defines the Size of Sliding Window
## ( e.g. 100 means in each snapshot we just consider 100 consecutive observations of each sensor)
w = 128 # 50 Equals to 1 second for MotionSense Dataset (it is on 50Hz samplig rate)
## Here We Choose Step Size for Building Diffrent Snapshots from Time-Series Data
## ( smaller step size will increase the amount of the instances and higher computational cost may be incurred )
s = 10
train_data, act_train, id_train, train_mean, train_std = ts_to_secs(train_ts.copy(),
w,
s,
standardize = True)
s = 10
val_data, act_val, id_val, val_mean, val_std = ts_to_secs(val_ts.copy(),
w,
s,
standardize = True,
mean = train_mean,
std = train_std)
s = 10
test_data, act_test, id_test, test_mean, test_std = ts_to_secs(test_ts.copy(),
w,
s,
standardize = True,
mean = train_mean,
std = train_std)
print("[INFO] -- Training Sections: "+str(train_data.shape))
print("[INFO] -- Validation Sections: "+str(val_data.shape))
print("[INFO] -- Test Sections: "+str(test_data.shape))
id_train_labels = to_categorical(id_train)
id_val_labels = to_categorical(id_val)
id_test_labels = to_categorical(id_test)
act_train_labels = to_categorical(act_train)
act_val_labels = to_categorical(act_val)
act_test_labels = to_categorical(act_test)
# -
## Here we add an extra dimension to the datasets just to be ready for using with Convolution2D
train_data = np.expand_dims(train_data,axis=3)
print("[INFO] -- Shape of Training Sections:", train_data.shape)
val_data = np.expand_dims(val_data,axis=3)
print("[INFO] -- Validation Sections:"+str(val_data.shape))
test_data = np.expand_dims(test_data,axis=3)
print("[INFO] -- Shape of Training Sections:", test_data.shape)
# +
import sys
window = 10 # SSA window == number of components
ssa_train_data = train_data.copy()
ssa_val_data = val_data.copy()
ssa_test_data = test_data.copy()
ssa_train_0 = []
ssa_train_1 = []
ssa_val_0 = []
ssa_val_1 = []
ssa_test_0 = []
ssa_test_1 = []
print("\n Train \n")
for i in range(len(ssa_train_data)):
ssa_train_0.append(SSA(ssa_train_data[i,0,:,0], window))
ssa_train_1.append(SSA(ssa_train_data[i,1,:,0], window))
if(i%100==1):
sys.stdout.write("\rNow: "+str(np.round(i*100/len(ssa_train_data), 2))+"%")
sys.stdout.flush()
print("\n Val \n")
for i in range(len(ssa_val_data)):
ssa_val_0.append(SSA(ssa_val_data[i,0,:,0], window))
ssa_val_1.append(SSA(ssa_val_data[i,1,:,0], window))
if(i%100==1):
sys.stdout.write("\rNow: "+str(np.round(i*100/len(ssa_val_data), 2))+"%")
sys.stdout.flush()
print("\n Test \n")
for i in range(len(ssa_test_data)):
ssa_test_0.append(SSA(ssa_test_data[i,0,:,0], window))
ssa_test_1.append(SSA(ssa_test_data[i,1,:,0], window))
if(i%100==1):
sys.stdout.write("\rNow: "+str(np.round(i*100/len(ssa_test_data), 2))+"%")
sys.stdout.flush()
# -
act_history = {}
ep = 32
for num_comps in range(1,11):
ssa_train_data = train_data.copy()
ssa_val_data = val_data.copy()
ssa_test_data = test_data.copy()
print("With "+str(num_comps)+" components:")
for i in range(len(ssa_train_data)):
ssa_train_data[i,0,:,0] = ssa_train_0[i].reconstruct(list(range(0,num_comps)))
ssa_train_data[i,1,:,0] = ssa_train_1[i].reconstruct(list(range(0,num_comps)))
for i in range(len(ssa_val_data)):
ssa_val_data[i,0,:,0] = ssa_val_0[i].reconstruct(list(range(0,num_comps)))
ssa_val_data[i,1,:,0] = ssa_val_1[i].reconstruct(list(range(0,num_comps)))
for i in range(len(ssa_test_data)):
ssa_test_data[i,0,:,0] = ssa_test_0[i].reconstruct(list(range(0,num_comps)))
ssa_test_data[i,1,:,0] = ssa_test_1[i].reconstruct(list(range(0,num_comps)))
height = train_data.shape[1]
width = train_data.shape[2]
id_class_numbers = 24
act_class_numbers = 4
fm = (2,5)
print("___________________________________________________")
## Callbacks
eval_metric= "val_f1_metric"
early_stop = keras.callbacks.EarlyStopping(monitor=eval_metric, mode='max', patience = 7)
filepath="XXACT.best.hdf5"
checkpoint = ModelCheckpoint(filepath, monitor=eval_metric, verbose=1, save_best_only=True, mode='max')
callbacks_list = [checkpoint, early_stop]
## Callbacks
eval_act = Estimator.build(height, width, act_class_numbers, name ="EVAL_ACT", fm=fm, act_func="softmax",hid_act_func="relu")
eval_act.compile( loss="categorical_crossentropy", optimizer='adam', metrics=['acc',f1_metric])
print("Model Size = "+str(eval_act.count_params()))
eval_act.fit(ssa_train_data, act_train_labels,
validation_data = (ssa_val_data, act_val_labels),
epochs = ep,
batch_size = 128,
verbose = 0,
class_weight = get_class_weights(np.argmax(act_train_labels,axis=1)),
callbacks = callbacks_list
)
eval_act.load_weights("XXACT.best.hdf5")
eval_act.compile( loss="categorical_crossentropy", optimizer='adam', metrics=['acc',f1_metric])
result1 = eval_act.evaluate(ssa_test_data, act_test_labels, verbose = 2)
act_acc = result1[1].round(4)*100
print("***[RESULT]*** ACT Accuracy: "+str(act_acc))
preds = eval_act.predict(ssa_test_data)
preds = np.argmax(preds, axis=1)
conf_mat = confusion_matrix(np.argmax(act_test_labels, axis=1), preds)
conf_mat = conf_mat.astype('float') / conf_mat.sum(axis=1)[:, np.newaxis]
print("***[RESULT]*** ACT Confusion Matrix")
print(np.array(conf_mat).round(3)*100)
f1act = f1_score(np.argmax(act_test_labels, axis=1), preds, average=None).mean()
print("***[RESULT]*** ACT Averaged F-1 Score : "+str(f1act))
act_history[num_comps] = f1act
act_history
|
msda/msda_res_fig_15_bottom_act.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Nucleus-segmentation
# language: python
# name: other-env
# ---
# + deletable=true editable=true
import numpy as np
import keras
from keras.models import Model, Sequential
from keras.layers import Input, add, BatchNormalization, Activation, Lambda
from keras.layers.core import Dense, Dropout, Activation, Flatten, Layer
from keras.layers.convolutional import Conv2D, MaxPooling2D, UpSampling2D
from keras.layers.core import Dropout, Lambda
from keras.utils import np_utils, generic_utils
from keras.optimizers import Adam, SGD
from keras import backend as K
from keras.metrics import categorical_accuracy
import keras.backend as K
import keras.layers as KL
# from sklearn.metrics import f1_score
# from .base_network import BaseNetwork
# from .custom_layers.SpatialTransformLayer import SpatialTransformLayer
# from skimage.transform import resize
# from skimage.io import imread
# import matplotlib.pyplot as plt
# class MicroblinkBaseNet(BaseNetwork):
# def __init__(self, output_directory, checkpoint_directory, config_dict, preprocessor, name = "MicroblinkBaseNet", train = True):
# BaseNetwork.__init__(self, output_directory, checkpoint_directory, config_dict, preprocessor, name=name, train = train)
# y, x = config_dict['STNOutputSize'].split(',')
# x = int(x)
# y = int(y)
# self.STN_output_size = (y, x)
# self.train_localization = bool(config_dict['TrainLocalization'])
# # def get_localization_network(self, inputs):
# # # initial weights for localization network for identity transform
# # b = np.zeros((2, 3), dtype='float32')
# # b[0, 0] = 1
# # b[1, 1] = 1
# # W = np.zeros((50, 6), dtype='float32')
# # weights = [W, b.flatten()]
# # # initial weights for localization network zoom on top of image
# # # b = np.zeros((2, 3), dtype='float32')
# # # b[0, 0] = 1
# # # b[1, 1] = 0.3
# # # b[1, 2] = -0.7
# # # W = np.zeros((50, 6), dtype='float32')
# # # weights = [W, b.flatten()]
# # # s = Lambda(lambda x: x / 255.) (inputs)
# # locnet = Convolution2D(16, (11, 11), activation='relu', name = 'localization_conv_1')(inputs)
# # locnet = Convolution2D(16, (1, 11), activation='relu', name = 'localization_conv_2')(inputs)
# # locnet = MaxPooling2D(pool_size=(2,2), name = 'localization_maxpool_1')(locnet)
# # locnet = Convolution2D(32, (7, 7), activation='relu', name = 'localization_conv_3')(locnet)
# # locnet = Convolution2D(32, (7, 7), activation='relu', name = 'localization_conv_4')(locnet)
# # locnet = MaxPooling2D(pool_size=(2,2), name = 'localization_maxpool_2')(locnet)
# # locnet = Convolution2D(64, (5, 5), activation='relu', name = 'localization__conv_5')(locnet)
# # locnet = Convolution2D(64, (5, 5), activation='relu', name = 'localization_conv_6')(locnet)
# # locnet = MaxPooling2D(pool_size=(2,2), name = 'localization_maxpool_3')(locnet)
# # locnet = Convolution2D(32, (3, 3), activation='relu', name = 'localization_conv_7')(locnet)
# # locnet = Convolution2D(32, (3, 3), activation='relu', name = 'localization_conv_8')(locnet)
# # locnet = MaxPooling2D(pool_size=(2,2), name = 'localization_maxpool_4')(locnet)
# # locnet = Convolution2D(16, (3, 3), activation='relu', name = 'localization_conv_9')(locnet)
# # locnet = Convolution2D(16, (3, 3), activation='relu', name = 'localization_conv_10')(locnet)
# # locnet = MaxPooling2D(pool_size=(2,2), name = 'localization_maxpool_5')(locnet)
# # locnet = Flatten()(locnet)
# # locnet = Dense(50, activation = 'relu', name = 'localization_dense_1')(locnet)
# # locnet = Dense(6, weights=weights, name = 'localization_dense_affine')(locnet)
# # locnet = Model(inputs = [inputs], outputs = [locnet])
# # return locnet
# def get_network(self):
# inputs = Input(self.preprocessor.get_shape())
# # self.locnet = self.get_localization_network(inputs)
# # for layer in self.locnet.layers:
# # layer.trainable = self.train_localization
# # outputs = SpatialTransformLayer(localization_net=self.locnet,
# # output_size=self.STN_output_size, name='spatial_layer_1')(inputs)
# # s = Lambda(lambda x: x / 255.) (outputs)
# outputs = Convolution2D(32, (3, 3), padding='same', activation = 'relu', name = 'classification_conv_1')(outputs)
# outputs = MaxPooling2D(pool_size=(2,2), name = 'classification_maxpool_1')(outputs)
# outputs = Convolution2D(32, (3, 3), padding='same', activation = 'relu', name = 'classification_conv_2')(outputs)
# outputs = MaxPooling2D(pool_size=(2,2), name = 'classification_maxpool_2')(outputs)
# outputs = Convolution2D(32, (3, 3), padding='same', activation = 'relu', name = 'classification_conv_3')(outputs)
# outputs = MaxPooling2D(pool_size=(2,2), name = 'classification_maxpool_3')(outputs)
# outputs = Convolution2D(32, (3, 3), padding='same', activation = 'relu', name = 'classification_conv_4')(outputs)
# outputs = MaxPooling2D(pool_size=(2,2), name = 'classification_maxpool_4')(outputs)
# outputs = Convolution2D(32, (3, 3), padding='same', activation = 'relu', name = 'classification_conv_5')(outputs)
# outputs = MaxPooling2D(pool_size=(2,2), name = 'classification_maxpool_5')(outputs)
# outputs = Convolution2D(32, (3, 3), padding='same', activation = 'relu', name = 'classification_conv_6')(outputs)
# outputs = MaxPooling2D(pool_size=(2,2), name = 'classification_maxpool_6')(outputs)
# outputs = Flatten()(outputs)
# outputs = Dense(256, activation = 'relu', name = 'classification_dense_1')(outputs)
# outputs = Dense(self.number_of_classes, activation='softmax', name = 'classification_dense_probs')(outputs)
# model = Model(inputs=[inputs], outputs=[outputs])
# model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=[categorical_accuracy])
# model.summary()
# X_in = model.input
# X_transformed = model.layers[1].output
# print(model.layers[1].name)
# self.transformation_operation = K.function([X_in], [X_transformed])
# return model
# def get_additional_callbacks(self):
# return []#[OutputsCallback(self.transformation_operation, self.preprocessor.X_train, self.preprocessor.IMG_HEIGHT, self.preprocessor.IMG_WIDTH, self.preprocessor.IMG_CHANNELS)] #return array of new callbacks [EarlyStopping(..), ..]
# +
def conv_block(feat_maps_out, prev):
prev = BatchNormalization()(prev) # Specifying the axis and mode allows for later merging
prev = Activation('relu')(prev)
prev = Conv2D(feat_maps_out, (3, 3), padding='same')(prev)
prev = BatchNormalization()(prev) # Specifying the axis and mode allows for later merging
prev = Activation('relu')(prev)
prev = Conv2D(feat_maps_out, (3, 3), padding='same')(prev)
return prev
def skip_block(feat_maps_in, feat_maps_out, prev):
if feat_maps_in != feat_maps_out:
# This adds in a 1x1 convolution on shortcuts that map between an uneven amount of channels
prev = Conv2D(feat_maps_out, (1, 1), padding='same')(prev)
return prev
def Residual(feat_maps_in, feat_maps_out, prev_layer):
'''
A customizable residual unit with convolutional and shortcut blocks
Args:
feat_maps_in: number of channels/filters coming in, from input or previous layer
feat_maps_out: how many output channels/filters this block will produce
prev_layer: the previous layer
'''
skip = skip_block(feat_maps_in, feat_maps_out, prev_layer)
conv = conv_block(feat_maps_out, prev_layer)
return add([skip, conv]) # the residual connection
def ResidualAttention(inputs, p = 1, t = 2, r = 1):
channel_axis = -1
num_channels = inputs._keras_shape[channel_axis]
first_residuals = inputs
for i in range(p):
first_residuals = Residual(num_channels, num_channels, first_residuals)
output_trunk = first_residuals
for i in range(t):
output_trunk = Residual(num_channels, num_channels, output_trunk)
output_soft_mask = MaxPooling2D(pool_size=(2,2))(first_residuals)
for i in range(r):
output_soft_mask = Residual(num_channels, num_channels, output_soft_mask)
#skip connection
output_skip_connection = Residual(num_channels, num_channels, output_soft_mask)
#2r residual blocks and first upsampling
output_soft_mask = MaxPooling2D(pool_size=(2,2))(output_soft_mask)
for i in range(2*r):
output_soft_mask = Residual(num_channels, num_channels, output_soft_mask)
output_soft_mask = UpSampling2D([2, 2])(output_soft_mask)
#addition of the skip connection
output_soft_mask = add([output_soft_mask, output_skip_connection])
#last r blocks of residuals and upsampling
for i in range(r):
output_soft_mask = Residual(num_channels, num_channels, output_soft_mask)
output_soft_mask = UpSampling2D([2, 2])(output_soft_mask)
#final attention output
output_soft_mask = Conv2D(num_channels, (1,1), activation='relu')(output_soft_mask)
#final attention output
output_soft_mask = Conv2D(num_channels, (1,1), activation='sigmoid')(output_soft_mask)
output = Lambda(lambda x:(1 + x[0]) * x[1])([output_soft_mask,output_trunk])
for i in range(p):
output = Residual(num_channels, num_channels, output)
return output
img_rows = 224
img_cols = 224
inp = Input((img_rows, img_cols, 1))
cnv1 = Conv2D(64, (7, 7), strides=[2,2], activation='relu', padding='same')(inp)
r1 = Residual(64, 128, cnv1)
# An example residual unit coming after a convolutional layer. NOTE: the above residual takes the 64 output channels
# from the Convolutional2D layer as the first argument to the Residual function
r2 = Residual(128, 128, r1)
r3 = Residual(128, 256, r2)
r3 = ResidualAttention(r3)
out = Conv2D(1, (1, 1), padding='same', activation='sigmoid')(r3)
print(out)
model = Model(inputs = [inp], outputs = [out])
model.compile(optimizer='adam', loss='mean_squared_error')
model.summary()
# -
|
scripts/Residual.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: nlp
# language: python
# name: nlp
# ---
# +
# default_exp seg_unet_training
# -
#export
from fastai import *
from fastai.vision import *
from fastai.callbacks.hooks import *
from ocr.seg_dataset_isri_unlv import isri_unlv_config
from pathlib import PosixPath
from ocr.core import save_inference, load_inference
#export
codes = list(isri_unlv_config.cat2id.keys())
print(codes)
path_lbl = isri_unlv_config.SEG_DIR
path_img = isri_unlv_config.IMG_DIR
a = os.listdir(path_lbl)
b = os.listdir(path_img)
for x in b:
if x not in a:
print(x)
fnames = get_image_files(path_img)
fnames[:3]
lbl_names = get_image_files(path_lbl)
lbl_names[:3]
img_f = fnames[0]
img = open_image(img_f)
img.show(figsize=(5,5))
get_y_fn = lambda x: path_lbl/f'{x.stem}.png'
mask = open_mask(get_y_fn(img_f))
mask.show(figsize=(5,5), alpha=1)
src_size = np.array(mask.shape[1:])
src_size,mask.data.unique()
# w = 360
w = 480
size = np.array([int(w*1.3), w])
bs = 4
src = (SegmentationItemList.from_folder(path_img)
.split_by_rand_pct(0.1)
.label_from_func(get_y_fn, classes=codes))
# +
#export
train_transforms = [
# crop_pad(),
rotate(degrees=(-10, 10), p=0.9),
symmetric_warp(magnitude=(-0.1, 0.1), p=0.9),
# dihedral_affine(p=1), # (flips image), will cause problems, because top left corner will be for example bottom right
# rand_zoom(scale=(.5,1.), p=0.9),
brightness(change=(0.4, 0.6), p=0.8),
contrast(scale=(0.8,1.2), p=0.8),
]
valid_transforms = [
rotate(degrees=(-1, 1), p=0.2)
]
transforms = (train_transforms, valid_transforms)
# transforms = get_transforms()
# -
data = (src.transform(transforms, size=size, tfm_y=True)
.databunch(bs=bs)
.normalize(imagenet_stats))
data
data.show_batch(2, figsize=(10,7))
# +
#export
name2id = {v:k for k,v in enumerate(codes)}
void_code = name2id['Background']
def acc_camvid(input, target):
target = target.squeeze(1)
mask = target != void_code
return (input.argmax(dim=1)[mask]==target[mask]).float().mean()
# -
metrics=acc_camvid
# metrics=accuracy
wd=1e-2
learn = unet_learner(data, models.resnet34, metrics=metrics, wd=wd)
learn.path = PosixPath('.')
learn.callbacks.append(ShowGraph(learn))
lr_find(learn)
learn.recorder.plot()
lr=1e-4
learn.fit_one_cycle(5, slice(lr))
save_inference(learn, 'unet_layout')
learn.show_results(rows=3, figsize=(20,20))
learn = load_inference('unet_layout')
learn.data = data
learn.unfreeze()
learn.fit_one_cycle(5, slice(lr/50, lr/10))
save_inference(learn, 'unet_layout')
learn.show_results(rows=3, figsize=(20,20))
learn.fit_one_cycle(30, 1e-5)
learn.fit_one_cycle(1, 1e-6)
save_inference(learn, 'unet_layout')
learn.show_results(rows=3, figsize=(20,20))
|
nbs/11_seg_unet_training.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import torch
import os
from datetime import datetime
import time
import random
import cv2
import pandas as pd
import numpy as np
import albumentations as A
import matplotlib.pyplot as plt
from albumentations.pytorch.transforms import ToTensorV2
from sklearn.model_selection import StratifiedKFold
from torch.utils.data import Dataset,DataLoader
from torch.utils.data.sampler import SequentialSampler, RandomSampler
from glob import glob
from tqdm import tqdm
SEED = 42
def seed_everything(seed):
random.seed(seed)
os.environ['PYTHONHASHSEED'] = str(seed)
np.random.seed(seed)
torch.manual_seed(seed)
torch.cuda.manual_seed(seed)
torch.backends.cudnn.deterministic = True
torch.backends.cudnn.benchmark = False
seed_everything(SEED)
# -
# # EDA number of labels
video_labels = pd.read_csv('train_labels.csv').fillna(0)
video_labels.head()
# +
video_labels_with_impact = video_labels[video_labels['impact'] > 0]
videos = video_labels.video.unique()
num = []
for video in videos[1:]:
v = video_labels[(video_labels["impact"]==1) & (video_labels["video"]==video)]
print("number of labels: {} in video {}".format(len(v), video))
num.append(len(v))
print("mean:{} max:{} min:{}".format(np.mean(num), np.max(num), np.min(num)))
# -
# # Make labels with all labels
video_labels = pd.read_csv('train_labels.csv').fillna(0)
# filter low-confident
video_labels_with_impact = video_labels[(video_labels['impact'] > 0)*(video_labels["confidence"]>1)*(video_labels["visibility"]>0)]
# +
video_labels['image_name'] = video_labels['video'].str.replace('.mp4', '') + '_' + video_labels['frame'].astype(str).str.zfill(3) + '.png'
video_labels['impact'] = video_labels['impact'].astype(int)+1
video_labels['x'] = video_labels['left']
video_labels['y'] = video_labels['top']
video_labels['w'] = video_labels['width']
video_labels['h'] = video_labels['height']
video_labels.to_csv("train_labels_all.csv")
video_labels.head()
# -
# # Generate impact only labels
video_labels = pd.read_csv('train_labels.csv').fillna(0)
video_labels_with_impact = video_labels[(video_labels['impact'] > 0)*(video_labels["confidence"]>1)*(video_labels["visibility"]>0)]
for row in tqdm(video_labels_with_impact[['video','frame','label']].values):
frames = np.array([-4,-3,-2,-1,1,2,3,4])+row[1]
video_labels.loc[(video_labels['video'] == row[0])
& (video_labels['frame'].isin(frames))
& (video_labels['label'] == row[2]), 'impact'] = 1
video_labels['image_name'] = video_labels['video'].str.replace('.mp4', '') + '_' + video_labels['frame'].astype(str).str.zfill(3) + '.png'
video_labels = video_labels[video_labels.groupby('image_name')['impact'].transform("sum") > 0].reset_index(drop=True)
video_labels['impact'] = video_labels['impact'].astype(int)+1
video_labels['x'] = video_labels['left']
video_labels['y'] = video_labels['top']
video_labels['w'] = video_labels['width']
video_labels['h'] = video_labels['height']
video_labels.head()
video_labels.to_csv("train_labels_4.csv")
video_labels = pd.read_csv('train_labels.csv').fillna(0)
video_labels_with_impact = video_labels[(video_labels['impact'] > 0)*(video_labels["confidence"]>1)*(video_labels["visibility"]>0)]
for row in tqdm(video_labels_with_impact[['video','frame','label']].values):
frames = np.array([-3,-2,-1,1,2,3])+row[1]
video_labels.loc[(video_labels['video'] == row[0])
& (video_labels['frame'].isin(frames))
& (video_labels['label'] == row[2]), 'impact'] = 1
video_labels['image_name'] = video_labels['video'].str.replace('.mp4', '') + '_' + video_labels['frame'].astype(str).str.zfill(3) + '.png'
video_labels = video_labels[video_labels.groupby('image_name')['impact'].transform("sum") > 0].reset_index(drop=True)
video_labels['impact'] = video_labels['impact'].astype(int)+1
video_labels['x'] = video_labels['left']
video_labels['y'] = video_labels['top']
video_labels['w'] = video_labels['width']
video_labels['h'] = video_labels['height']
video_labels.head()
video_labels.to_csv("train_labels_3.csv")
video_labels = pd.read_csv('train_labels.csv').fillna(0)
video_labels_with_impact = video_labels[(video_labels['impact'] > 0)*(video_labels["confidence"]>1)*(video_labels["visibility"]>0)]
for row in tqdm(video_labels_with_impact[['video','frame','label']].values):
frames = np.array([-2,-1,1,2,])+row[1]
video_labels.loc[(video_labels['video'] == row[0])
& (video_labels['frame'].isin(frames))
& (video_labels['label'] == row[2]), 'impact'] = 1
video_labels['image_name'] = video_labels['video'].str.replace('.mp4', '') + '_' + video_labels['frame'].astype(str).str.zfill(3) + '.png'
video_labels = video_labels[video_labels.groupby('image_name')['impact'].transform("sum") > 0].reset_index(drop=True)
video_labels['impact'] = video_labels['impact'].astype(int)+1
video_labels['x'] = video_labels['left']
video_labels['y'] = video_labels['top']
video_labels['w'] = video_labels['width']
video_labels['h'] = video_labels['height']
video_labels.head()
video_labels.to_csv("train_labels_2.csv")
# # Generate images from video
def mk_images(video_name, video_labels, video_dir, out_dir, only_with_impact=True):
video_path=f"{video_dir}/{video_name}"
video_name = os.path.basename(video_path)
vidcap = cv2.VideoCapture(video_path)
if only_with_impact:
boxes_all = video_labels.query("video == @video_name")
print(video_path, boxes_all[boxes_all.impact > 1.0].shape[0])
else:
print(video_path)
frame = 0
while True:
it_worked, img = vidcap.read()
if not it_worked:
break
frame += 1
if only_with_impact:
boxes = video_labels.query("video == @video_name and frame == @frame")
boxes_with_impact = boxes[boxes.impact > 1.0]
if boxes_with_impact.shape[0] == 0:
continue
img_name = f"{video_name}_frame{frame}"
image_path = f'{out_dir}/{video_name}'.replace('.mp4',f'_{str(frame).zfill(3)}.png')
_ = cv2.imwrite(image_path, img)
uniq_video = video_labels.video.unique()
video_dir = 'train'
out_dir = 'train_images'
# !mkdir -p $out_dir
for video_name in uniq_video:
mk_images(video_name, video_labels, video_dir, out_dir)
uniq_video = video_labels.video.unique()
video_dir = 'train'
out_dir = 'train_images'
# !mkdir -p $out_dir
for video_name in uniq_video:
mk_images(video_name, video_labels, video_dir, out_dir, False)
|
train-prepare-labels.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from PIL import Image
import numpy as np
from matplotlib.pyplot import imshow
import imageio
import cv2
import os
from os import path
path="./garfield_3paneldata/"
path_to_save="./garfield_data_test/"
if not os.path.exists(path_to_save):
os.mkdir(path_to_save)
for i in range(1991,2020,1):
if not os.path.exists(path_to_save+str(i)):
os.mkdir(path_to_save+str(i))
p1=path+str(i)+"/"
p2=path_to_save+str(i)+'/'
for j in range(367):
if os.path.exists(p1+str(j)+".gif"):
gif = imageio.mimread(p1+str(j)+".gif")
img = cv2.cvtColor(gif[0], cv2.COLOR_RGB2BGR)
cv2.imwrite(p2+str(j)+".jpeg",img)
p="./garfield_data_single/"
if not os.path.exists(p):
os.mkdir(p)
for i in range(1991,2020,1):
if not os.path.exists(p+str(i)):
os.mkdir(p+str(i))
p1=path_to_save+str(i)+"/"
p2=p+str(i)+'/'
dim = (256,256)
for j in range(367):
if os.path.exists(p1+str(j)+".jpeg"):
img = cv2.imread(p1+str(j)+".jpeg")
x=img.shape[1]
if img.shape[0]<500:
crop_img1 = cv2.resize(img[:, :int(x/3)], dim)
crop_img2 = cv2.resize(img[:, int(x/3):int(2*x/3)], dim)
crop_img3 = cv2.resize(img[:, int(2*x/3):], dim)
cv2.imwrite(p2+str(j)+"_1.jpeg",crop_img1)
cv2.imwrite(p2+str(j)+"_2.jpeg",crop_img2)
cv2.imwrite(p2+str(j)+"_3.jpeg",crop_img3)
# cv2.waitKey(0)
p="./garfield_data_2_1/"
if not os.path.exists(p):
os.mkdir(p)
for i in range(1991,2020,1):
if not os.path.exists(p+str(i)):
os.mkdir(p+str(i))
p1=path_to_save+str(i)+"/"
p2=p+str(i)+'/'
dim = (512,256)
for j in range(367):
if os.path.exists(p1+str(j)+".jpeg"):
img = cv2.imread(p1+str(j)+".jpeg")
x=img.shape[1]
if img.shape[0]<500:
# crop_img1 = cv2.resize(img[:, :int(x/3)], dim)
crop_img2 = cv2.resize(img[:, :int(2*x/3)], dim)
crop_img3 = cv2.resize(img[:, int(2*x/3):], dim)
# cv2.imwrite(p2+str(j)+"_1.jpeg",crop_img1)
cv2.imwrite(p2+str(j)+"_1.jpeg",crop_img2)
cv2.imwrite(p2+str(j)+"_2.jpeg",crop_img3)
|
Data_cleaner.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 选择
# ## 布尔类型、数值和表达式
# 
# - 注意:比较运算符的相等是两个等到,一个等到代表赋值
# - 在Python中可以用整型0来代表False,其他数字来代表True
# - 后面还会讲到 is 在判断语句中的用发
# ## 字符串的比较使用ASCII值
# ## Markdown
# - https://github.com/younghz/Markdown
# ## EP:
# - <img src="../Photo/34.png"></img>
# - 输入一个数字,判断其实奇数还是偶数
# ## 产生随机数字
# - 函数random.randint(a,b) 可以用来产生一个a和b之间且包括a和b的随机整数
import math,random
num = random.randint(0,10)
while True :
num_ = eval(input('>>'))
if num_ > num :
print('大了')
elif num < num_:
print('小了')
else:
break
print('等于')
# ## 其他random方法
# - random.random 返回0.0到1.0之间前闭后开区间的随机浮点
# - random.randrange(a,b) 前闭后开
# ## EP:
# - 产生两个随机整数number1和number2,然后显示给用户,使用户输入数字的和,并判定其是否正确
# - 进阶:写一个随机序号点名程序
# ## if语句
# - 如果条件正确就执行一个单向if语句,亦即当条件为真的时候才执行if内部的语句
# - Python有很多选择语句:
# > - 单向if
# - 双向if-else
# - 嵌套if
# - 多向if-elif-else
#
# - 注意:当语句含有子语句的时候,那么一定至少要有一个缩进,也就是说如果有儿子存在,那么一定要缩进
# - 切记不可tab键和space混用,单用tab 或者 space
# - 当你输出的结果是无论if是否为真时都需要显示时,语句应该与if对齐
# ## EP:
# - 用户输入一个数字,判断其实奇数还是偶数
# - 进阶:可以查看下4.5实例研究猜生日
# ## 双向if-else 语句
# - 如果条件为真,那么走if内部语句,否则走else内部语句
# ## EP:
# - 产生两个随机整数number1和number2,然后显示给用户,使用户输入数字,并判定其是否正确,如果正确打印“you‘re correct”,否则打印正确错误
# ## 嵌套if 和多向if-elif-else
# 
# ## EP:
# - 提示用户输入一个年份,然后显示表示这一年的动物
# 
# - 计算身体质量指数的程序
# - BMI = 以千克为单位的体重除以以米为单位的身高
# 
# +
hight = eval(input('身高'))
weight = eval(input('体重'))
BMI = weight/hight
if BMI < 18.5:
print('超轻')
elif BMI >= 30.0:
print('痴肥')
elif 25.0<= BMI < 30.0:
print('超重')
elif 18.5<= BMI < 25.0:
print('标准')
else:
print('标准')
# -
# ## 逻辑运算符
# 
# 
# 
# ## EP:
# - 判定闰年:一个年份如果能被4整除但不能被100整除,或者能被400整除,那么这个年份就是闰年
# - 提示用户输入一个年份,并返回是否是闰年
# - 提示用户输入一个数字,判断其是否为水仙花数
year = eval(input('输入年份'))
if (year%4 == 0 and year % 100 !=0) or year % 400 == 0 :
print('是闰年')
else:
print('不是闰年')
# +
num = eval(input('输入一个3位数'))
a = num // 100
b = (num // 10) % 10
c = num % 10
d = a*a*a+b*b*b+c*c*c
if d == num :
print('是水仙花数')
else:
print('不是水仙花数')
# -
# ## 实例研究:彩票
# 
# # Homework
# - 1
# 
import math
a,b,c = eval(input('输入a,b,c值'))
judge = b**2-4*a*c
if judge > 0:
r1 =(-b+math.sqrt(judge))/2*a
r2 =(-b-math.sqrt(judge))/2*a
print(r1,r2)
elif judge == 0:
r3 =(-b+math.sqrt(judge))/2*a
print(r1)
else:
print("The equation has no real roots")
# - 2
# 
import math,random
num_1 = random.randint(0,99)
num_2 = random.randint(0,99)
num_3 =num_1+num_2
print(num_1,num_2)
sum = eval(input('输入的和'))
if sum == num_3:
print('Ture')
else:
print('False')
# - 3
# 
# - 4
# 
n1,n2,n3=eval(input('输入三个整数,以逗号隔开'))
if n1 < n2 :
n4 = n1
n1 = n2
n2 = n4
elif n1 < n3 :
n4 = n1
n1 = n3
n3 = n4
elif n2 < n3:
n4 = n2
n2 = n3
n3 = n4
print(n3,n2,n1)
# - 5
# 
weight_1,price_1 = eval(input('重量1,价钱1:'))
weight_2,price_2 = eval(input('重量2,价钱2:'))
aver_1 = price_1/weight_1
aver_2 = price_2/weight_2
if aver_1 > aver_2:
print('Package 2 has the better price')
elif aver_1 < aver_2:
print('Package 1 has the better price')
else:
print('1 = 2')
# - 6
# 
# - 7
# 
import random
x = random.randint(0,1)
x0 =["正","反"]
x_ =input('猜测是哪面:')
if x_ == x0[x] :
print('Ture')
else:
print('False')
# - 8
# 
import random
x = random.randint(0,2)
x_ =eval(input('输入0、1、2:'))
if x == x_ :
print('平')
elif x == 0 and x_ == 1:
print('你赢了')
elif x == 1 and x_ == 2:
print('你赢了')
elif x == 0 and x_ == 2:
print('你输了')
elif x == 1 and x_ == 0:
print('你输了')
elif x == 2 and x_ == 1:
print('你输了')
elif x == 2 and x_ == 0:
print('你赢了')
# - 9
# 
# - 10
# 
import random
x = random.randint(0,13)
y = random.randint(0,3)
x_ = ["Ace","2","3","4","5","6","7","8","9","10","jack","Queen","King"]
y_ = ["梅花","红桃","方块","黑桃"]
print("你抽中的牌是:"+str(y_[y])+str(x_[x]))
# - 11
# 
num = input("输入一个三位数:")
if num[0] == num[2]:
print(num+"是一个回文数")
else:
print(num+"不是一个回文数")
# - 12
# 
a,b,c = eval(input("请输入三角形的三条边长:"))
if (a+b>c and a-b<c):
s = a+b+c
print("周长:"+str(s))
else:
print("输入的3条边长不能构成三角形")
|
rxl 9.12.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# # Sentiment Analysis Web App
#
# _Deep Learning Nanodegree Program | Deployment_
#
# ---
#
# In this notebook we will use Amazon's SageMaker service to construct a random tree model to predict the sentiment of a movie review. In addition, we will deploy this model to an endpoint and construct a very simple web app which will interact with our model's deployed endpoint.
#
# ## General Outline
#
# Typically, when using a notebook instance with SageMaker, you will proceed through the following steps. Of course, not every step will need to be done with each project. Also, there is quite a lot of room for variation in many of the steps, as you will see throughout these lessons.
#
# 1. Download or otherwise retrieve the data.
# 2. Process / Prepare the data.
# 3. Upload the processed data to S3.
# 4. Train a chosen model.
# 5. Test the trained model (typically using a batch transform job).
# 6. Deploy the trained model.
# 7. Use the deployed model.
#
# In this notebook we will progress through each of the steps above. We will also see that the final step, using the deployed model, can be quite challenging.
# ## Step 1: Downloading the data
#
# The dataset we are going to use is very popular among researchers in Natural Language Processing, usually referred to as the [IMDb dataset](http://ai.stanford.edu/~amaas/data/sentiment/). It consists of movie reviews from the website [imdb.com](http://www.imdb.com/), each labeled as either '**pos**itive', if the reviewer enjoyed the film, or '**neg**ative' otherwise.
#
# > <NAME>., et al. [Learning Word Vectors for Sentiment Analysis](http://ai.stanford.edu/~amaas/data/sentiment/). In _Proceedings of the 49th Annual Meeting of the Association for Computational Linguistics: Human Language Technologies_. Association for Computational Linguistics, 2011.
#
# We begin by using some Jupyter Notebook magic to download and extract the dataset.
# %mkdir ../data
# !wget -O ../data/aclImdb_v1.tar.gz http://ai.stanford.edu/~amaas/data/sentiment/aclImdb_v1.tar.gz
# !tar -zxf ../data/aclImdb_v1.tar.gz -C ../data
# ## Step 2: Preparing and Processing the data
#
# The data we have downloaded is split into various files, each of which contains a single review. It will be much easier going forward if we combine these individual files into two large files, one for training and one for testing.
# +
import os
import glob
def read_imdb_data(data_dir='../data/aclImdb'):
data = {}
labels = {}
for data_type in ['train', 'test']:
data[data_type] = {}
labels[data_type] = {}
for sentiment in ['pos', 'neg']:
data[data_type][sentiment] = []
labels[data_type][sentiment] = []
path = os.path.join(data_dir, data_type, sentiment, '*.txt')
files = glob.glob(path)
for f in files:
with open(f) as review:
data[data_type][sentiment].append(review.read())
# Here we represent a positive review by '1' and a negative review by '0'
labels[data_type][sentiment].append(1 if sentiment == 'pos' else 0)
assert len(data[data_type][sentiment]) == len(labels[data_type][sentiment]), \
"{}/{} data size does not match labels size".format(data_type, sentiment)
return data, labels
# -
data, labels = read_imdb_data()
print("IMDB reviews: train = {} pos / {} neg, test = {} pos / {} neg".format(
len(data['train']['pos']), len(data['train']['neg']),
len(data['test']['pos']), len(data['test']['neg'])))
# +
from sklearn.utils import shuffle
def prepare_imdb_data(data, labels):
"""Prepare training and test sets from IMDb movie reviews."""
#Combine positive and negative reviews and labels
data_train = data['train']['pos'] + data['train']['neg']
data_test = data['test']['pos'] + data['test']['neg']
labels_train = labels['train']['pos'] + labels['train']['neg']
labels_test = labels['test']['pos'] + labels['test']['neg']
#Shuffle reviews and corresponding labels within training and test sets
data_train, labels_train = shuffle(data_train, labels_train)
data_test, labels_test = shuffle(data_test, labels_test)
# Return a unified training data, test data, training labels, test labets
return data_train, data_test, labels_train, labels_test
# -
train_X, test_X, train_y, test_y = prepare_imdb_data(data, labels)
print("IMDb reviews (combined): train = {}, test = {}".format(len(train_X), len(test_X)))
train_X[100]
# ## Processing the data
#
# Now that we have our training and testing datasets merged and ready to use, we need to start processing the raw data into something that will be useable by our machine learning algorithm. To begin with, we remove any html formatting and any non-alpha numeric characters that may appear in the reviews. We will do this in a very simplistic way using Python's regular expression module. We will discuss the reason for this rather simplistic pre-processing later on.
# +
import re
REPLACE_NO_SPACE = re.compile("(\.)|(\;)|(\:)|(\!)|(\')|(\?)|(\,)|(\")|(\()|(\))|(\[)|(\])")
REPLACE_WITH_SPACE = re.compile("(<br\s*/><br\s*/>)|(\-)|(\/)")
def review_to_words(review):
words = REPLACE_NO_SPACE.sub("", review.lower())
words = REPLACE_WITH_SPACE.sub(" ", words)
return words
# -
review_to_words(train_X[100])
# +
import pickle
cache_dir = os.path.join("../cache", "sentiment_web_app") # where to store cache files
os.makedirs(cache_dir, exist_ok=True) # ensure cache directory exists
def preprocess_data(data_train, data_test, labels_train, labels_test,
cache_dir=cache_dir, cache_file="preprocessed_data.pkl"):
"""Convert each review to words; read from cache if available."""
# If cache_file is not None, try to read from it first
cache_data = None
if cache_file is not None:
try:
with open(os.path.join(cache_dir, cache_file), "rb") as f:
cache_data = pickle.load(f)
print("Read preprocessed data from cache file:", cache_file)
except:
pass # unable to read from cache, but that's okay
# If cache is missing, then do the heavy lifting
if cache_data is None:
# Preprocess training and test data to obtain words for each review
#words_train = list(map(review_to_words, data_train))
#words_test = list(map(review_to_words, data_test))
words_train = [review_to_words(review) for review in data_train]
words_test = [review_to_words(review) for review in data_test]
# Write to cache file for future runs
if cache_file is not None:
cache_data = dict(words_train=words_train, words_test=words_test,
labels_train=labels_train, labels_test=labels_test)
with open(os.path.join(cache_dir, cache_file), "wb") as f:
pickle.dump(cache_data, f)
print("Wrote preprocessed data to cache file:", cache_file)
else:
# Unpack data loaded from cache file
words_train, words_test, labels_train, labels_test = (cache_data['words_train'],
cache_data['words_test'], cache_data['labels_train'], cache_data['labels_test'])
return words_train, words_test, labels_train, labels_test
# -
# Preprocess data
train_X, test_X, train_y, test_y = preprocess_data(train_X, test_X, train_y, test_y)
# ### Extract Bag-of-Words features
#
# For the model we will be implementing, rather than using the reviews directly, we are going to transform each review into a Bag-of-Words feature representation. Keep in mind that 'in the wild' we will only have access to the training set so our transformer can only use the training set to construct a representation.
# +
import numpy as np
from sklearn.feature_extraction.text import CountVectorizer
# from sklearn.externals import joblib
# joblib is an enhanced version of pickle that is more efficient for storing NumPy arrays
import joblib
def extract_BoW_features(words_train, words_test, vocabulary_size=5000,
cache_dir=cache_dir, cache_file="bow_features.pkl"):
"""Extract Bag-of-Words for a given set of documents, already preprocessed into words."""
# If cache_file is not None, try to read from it first
cache_data = None
if cache_file is not None:
try:
with open(os.path.join(cache_dir, cache_file), "rb") as f:
cache_data = joblib.load(f)
print("Read features from cache file:", cache_file)
except:
pass # unable to read from cache, but that's okay
# If cache is missing, then do the heavy lifting
if cache_data is None:
# Fit a vectorizer to training documents and use it to transform them
# NOTE: Training documents have already been preprocessed and tokenized into words;
# pass in dummy functions to skip those steps, e.g. preprocessor=lambda x: x
vectorizer = CountVectorizer(max_features=vocabulary_size)
features_train = vectorizer.fit_transform(words_train).toarray()
# Apply the same vectorizer to transform the test documents (ignore unknown words)
features_test = vectorizer.transform(words_test).toarray()
# NOTE: Remember to convert the features using .toarray() for a compact representation
# Write to cache file for future runs (store vocabulary as well)
if cache_file is not None:
vocabulary = vectorizer.vocabulary_
cache_data = dict(features_train=features_train, features_test=features_test,
vocabulary=vocabulary)
with open(os.path.join(cache_dir, cache_file), "wb") as f:
joblib.dump(cache_data, f)
print("Wrote features to cache file:", cache_file)
else:
# Unpack data loaded from cache file
features_train, features_test, vocabulary = (cache_data['features_train'],
cache_data['features_test'], cache_data['vocabulary'])
# Return both the extracted features as well as the vocabulary
return features_train, features_test, vocabulary
# -
# Extract Bag of Words features for both training and test datasets
train_X, test_X, vocabulary = extract_BoW_features(train_X, test_X)
len(train_X[100])
# ## Step 3: Upload data to S3
#
# Now that we have created the feature representation of our training (and testing) data, it is time to start setting up and using the XGBoost classifier provided by SageMaker.
#
# ### Writing the datasets
#
# The XGBoost classifier that we will be using requires the dataset to be written to a file and stored using Amazon S3. To do this, we will start by splitting the training dataset into two parts, the data we will train the model with and a validation set. Then, we will write those datasets to a file locally and then upload the files to S3. In addition, we will write the test set to a file and upload that file to S3. This is so that we can use SageMakers Batch Transform functionality to test our model once we've fit it.
# +
import pandas as pd
# Earlier we shuffled the training dataset so to make things simple we can just assign
# the first 10 000 reviews to the validation set and use the remaining reviews for training.
val_X = pd.DataFrame(train_X[:10000])
train_X = pd.DataFrame(train_X[10000:])
val_y = pd.DataFrame(train_y[:10000])
train_y = pd.DataFrame(train_y[10000:])
# -
# The documentation for the XGBoost algorithm in SageMaker requires that the training and validation datasets should contain no headers or index and that the label should occur first for each sample.
#
# For more information about this and other algorithms, the SageMaker developer documentation can be found on __[Amazon's website.](https://docs.aws.amazon.com/sagemaker/latest/dg/)__
# First we make sure that the local directory in which we'd like to store the training and validation csv files exists.
data_dir = '../data/sentiment_web_app'
if not os.path.exists(data_dir):
os.makedirs(data_dir)
# +
pd.DataFrame(test_X).to_csv(os.path.join(data_dir, 'test.csv'), header=False, index=False)
pd.concat([val_y, val_X], axis=1).to_csv(os.path.join(data_dir, 'validation.csv'), header=False, index=False)
pd.concat([train_y, train_X], axis=1).to_csv(os.path.join(data_dir, 'train.csv'), header=False, index=False)
# +
# To save a bit of memory we can set text_X, train_X, val_X, train_y and val_y to None.
test_X = train_X = val_X = train_y = val_y = None
# -
# ### Uploading Training / Validation files to S3
#
# Amazon's S3 service allows us to store files that can be accessed by both the built-in training models such as the XGBoost model we will be using as well as custom models such as the one we will see a little later.
#
# For this and most other tasks we will be doing using SageMaker, there are two methods we could use. The first is to use the low level functionality of SageMaker which requires knowing each of the objects involved in the SageMaker environment. The second is to use the high level functionality in which certain choices have been made on the user's behalf. The low level approach benefits from allowing the user a great deal of flexibility while the high level approach makes development much quicker. For our purposes we will opt to use the high level approach although using the low-level approach is certainly an option.
#
# Recall the method `upload_data()` which is a member of the object representing our current SageMaker session. What this method does is upload the data to the default bucket (which is created if it does not exist) into the path described by the key_prefix variable. To see this for yourself, once you have uploaded the data files, go to the S3 console and look to see where the files have been uploaded.
#
# For additional resources, see the __[SageMaker API documentation](http://sagemaker.readthedocs.io/en/latest/)__ and in addition the __[SageMaker Developer Guide.](https://docs.aws.amazon.com/sagemaker/latest/dg/)__
# +
import sagemaker
session = sagemaker.Session() # Store the current SageMaker session
# S3 prefix (which folder will we use)
prefix = 'sentiment-web-app'
test_location = session.upload_data(os.path.join(data_dir, 'test.csv'), key_prefix=prefix)
val_location = session.upload_data(os.path.join(data_dir, 'validation.csv'), key_prefix=prefix)
train_location = session.upload_data(os.path.join(data_dir, 'train.csv'), key_prefix=prefix)
# -
# ## Step 4: Creating the XGBoost model
#
# Now that the data has been uploaded it is time to create the XGBoost model. To begin with, we need to do some setup. At this point it is worth discussing what a model is in SageMaker. It is easiest to think of a model of comprising three different objects in the SageMaker ecosystem, which interact with one another.
#
# - Model Artifacts
# - Training Code (Container)
# - Inference Code (Container)
#
# The Model Artifacts are what you might think of as the actual model itself. For example, if you were building a neural network, the model artifacts would be the weights of the various layers. In our case, for an XGBoost model, the artifacts are the actual trees that are created during training.
#
# The other two objects, the training code and the inference code are then used to manipulate the training artifacts. More precisely, the training code uses the training data that is provided and creates the model artifacts, while the inference code uses the model artifacts to make predictions on new data.
#
# The way that SageMaker runs the training and inference code is by making use of Docker containers. For now, think of a container as being a way of packaging code up so that dependencies aren't an issue.
# +
from sagemaker import get_execution_role
# Our current execution role is required when creating the model as the training
# and inference code will need to access the model artifacts.
role = get_execution_role()
# -
role
# +
# We need to retrieve the location of the container which is provided by Amazon for using XGBoost.
# As a matter of convenience, the training and inference code both use the same container.
from sagemaker.amazon.amazon_estimator import get_image_uri
container = get_image_uri(session.boto_region_name, 'xgboost')
# +
# First we create a SageMaker estimator object for our model.
xgb = sagemaker.estimator.Estimator(container, # The location of the container we wish to use
role, # What is our current IAM Role
train_instance_count=1, # How many compute instances
train_instance_type='ml.m4.xlarge', # What kind of compute instances
output_path='s3://{}/{}/output'.format(session.default_bucket(), prefix),
sagemaker_session=session)
# And then set the algorithm specific parameters.
xgb.set_hyperparameters(max_depth=5,
eta=0.2,
gamma=4,
min_child_weight=6,
subsample=0.8,
silent=0,
objective='binary:logistic',
early_stopping_rounds=10,
num_round=500)
# -
# ### Fit the XGBoost model
#
# Now that our model has been set up we simply need to attach the training and validation datasets and then ask SageMaker to set up the computation.
s3_input_train = sagemaker.s3_input(s3_data=train_location, content_type='csv')
s3_input_validation = sagemaker.s3_input(s3_data=val_location, content_type='csv')
xgb.fit({'train': s3_input_train, 'validation': s3_input_validation})
# ## Step 5: Testing the model
#
# Now that we've fit our XGBoost model, it's time to see how well it performs. To do this we will use SageMakers Batch Transform functionality. Batch Transform is a convenient way to perform inference on a large dataset in a way that is not realtime. That is, we don't necessarily need to use our model's results immediately and instead we can perform inference on a large number of samples. An example of this in industry might be performing an end of month report. This method of inference can also be useful to us as it means that we can perform inference on our entire test set.
#
# To perform a Batch Transformation we need to first create a transformer objects from our trained estimator object.
xgb_transformer = xgb.transformer(instance_count = 1, instance_type = 'ml.m4.xlarge')
# Next we actually perform the transform job. When doing so we need to make sure to specify the type of data we are sending so that it is serialized correctly in the background. In our case we are providing our model with csv data so we specify `text/csv`. Also, if the test data that we have provided is too large to process all at once then we need to specify how the data file should be split up. Since each line is a single entry in our data set we tell SageMaker that it can split the input on each line.
xgb_transformer.transform(test_location, content_type='text/csv', split_type='Line')
# Currently the transform job is running but it is doing so in the background. Since we wish to wait until the transform job is done and we would like a bit of feedback we can run the `wait()` method.
xgb_transformer.wait()
# Now the transform job has executed and the result, the estimated sentiment of each review, has been saved on S3. Since we would rather work on this file locally we can perform a bit of notebook magic to copy the file to the `data_dir`.
# !aws s3 cp --recursive $xgb_transformer.output_path $data_dir
# The last step is now to read in the output from our model, convert the output to something a little more usable, in this case we want the sentiment to be either `1` (positive) or `0` (negative), and then compare to the ground truth labels.
predictions = pd.read_csv(os.path.join(data_dir, 'test.csv.out'), header=None)
predictions = [round(num) for num in predictions.squeeze().values]
from sklearn.metrics import accuracy_score
accuracy_score(test_y, predictions)
# ## Step 6: Deploying the model
#
# Once we construct and fit our model, SageMaker stores the resulting model artifacts and we can use those to deploy an endpoint (inference code). To see this, look in the SageMaker console and you should see that a model has been created along with a link to the S3 location where the model artifacts have been stored.
#
# Deploying an endpoint is a lot like training the model with a few important differences. The first is that a deployed model doesn't change the model artifacts, so as you send it various testing instances the model won't change. Another difference is that since we aren't performing a fixed computation, as we were in the training step or while performing a batch transform, the compute instance that gets started stays running until we tell it to stop. This is important to note as if we forget and leave it running we will be charged the entire time.
#
# In other words **If you are no longer using a deployed endpoint, shut it down!**
xgb_predictor = xgb.deploy(initial_instance_count = 1, instance_type = 'ml.m4.xlarge')
# ### Testing the model (again)
#
# Now that we have deployed our endpoint, we can send the testing data to it and get back the inference results. We already did this earlier using the batch transform functionality of SageMaker, however, we will test our model again using the newly deployed endpoint so that we can make sure that it works properly and to get a bit of a feel for how the endpoint works.
#
# When using the created endpoint it is important to know that we are limited in the amount of information we can send in each call so we need to break the testing data up into chunks and then send each chunk. Also, we need to serialize our data before we send it to the endpoint to ensure that our data is transmitted properly. Fortunately, SageMaker can do the serialization part for us provided we tell it the format of our data.
# +
from sagemaker.predictor import csv_serializer
# We need to tell the endpoint what format the data we are sending is in so that SageMaker can perform the serialization.
xgb_predictor.content_type = 'text/csv'
xgb_predictor.serializer = csv_serializer
# +
# We split the data into chunks and send each chunk seperately, accumulating the results.
def predict(data, rows=512):
split_array = np.array_split(data, int(data.shape[0] / float(rows) + 1))
predictions = ''
for array in split_array:
predictions = ','.join([predictions, xgb_predictor.predict(array).decode('utf-8')])
return np.fromstring(predictions[1:], sep=',')
# +
test_X = pd.read_csv(os.path.join(data_dir, 'test.csv'), header=None).values
predictions = predict(test_X)
predictions = [round(num) for num in predictions]
# -
# Lastly, we check to see what the accuracy of our model is.
from sklearn.metrics import accuracy_score
accuracy_score(test_y, predictions)
# And the results here should agree with the model testing that we did earlier using the batch transform job.
#
# ### Cleaning up
#
# Now that we've determined that deploying our model works as expected, we are going to shut it down. Remember that the longer the endpoint is left running, the greater the cost and since we have a bit more work to do before we are able to use our endpoint with our simple web app, we should shut everything down.
xgb_predictor.delete_endpoint()
# ## Step 7: Putting our model to work
#
# As we've mentioned a few times now, our goal is to have our model deployed and then access it using a very simple web app. The intent is for this web app to take some user submitted data (a review), send it off to our endpoint (the model) and then display the result.
#
# However, there is a small catch. Currently the only way we can access the endpoint to send it data is using the SageMaker API. We can, if we wish, expose the actual URL that our model's endpoint is receiving data from, however, if we just send it data ourselves we will not get anything in return. This is because the endpoint created by SageMaker requires the entity accessing it have the correct permissions. So, we would need to somehow authenticate our web app with AWS.
#
# Having a website that authenticates to AWS seems a bit beyond the scope of this lesson so we will opt for an alternative approach. Namely, we will create a new endpoint which does not require authentication and which acts as a proxy for the SageMaker endpoint.
#
# As an additional constraint, we will try to avoid doing any data processing in the web app itself. Remember that when we constructed and tested our model we started with a movie review, then we simplified it by removing any html formatting and punctuation, then we constructed a bag of words embedding and the resulting vector is what we sent to our model. All of this needs to be done to our user input as well.
#
# Fortunately we can do all of this data processing in the backend, using Amazon's Lambda service.
#
# <img src="Web App Diagram.svg">
#
# The diagram above gives an overview of how the various services will work together. On the far right is the model which we trained above and which will be deployed using SageMaker. On the far left is our web app that collects a user's movie review, sends it off and expects a positive or negative sentiment in return.
#
# In the middle is where some of the magic happens. We will construct a Lambda function, which you can think of as a straightforward Python function that can be executed whenever a specified event occurs. This Python function will do the data processing we need to perform on a user submitted review. In addition, we will give this function permission to send and recieve data from a SageMaker endpoint.
#
# Lastly, the method we will use to execute the Lambda function is a new endpoint that we will create using API Gateway. This endpoint will be a url that listens for data to be sent to it. Once it gets some data it will pass that data on to the Lambda function and then return whatever the Lambda function returns. Essentially it will act as an interface that lets our web app communicate with the Lambda function.
#
# ### Processing a single review
#
# For now, suppose we are given a movie review by our user in the form of a string, like so:
test_review = "Nothing but a disgusting materialistic pageant of glistening abed remote control greed zombies, totally devoid of any heart or heat. A romantic comedy that has zero romantic chemestry and zero laughs!"
# How do we go from this string to the bag of words feature vector that is expected by our model?
#
# If we recall at the beginning of this notebook, the first step is to remove any unnecessary characters using the `review_to_words` method. Remember that we intentionally did this in a very simplistic way. This is because we are going to have to copy this method to our (eventual) Lambda function (we will go into more detail later) and this means it needs to be rather simplistic.
test_words = review_to_words(test_review)
print(test_words)
# Next, we need to construct a bag of words embedding of the `test_words` string. To do this, remember that a bag of words embedding uses a `vocabulary` consisting of the most frequently appearing words in a set of documents. Then, for each word in the vocabulary we record the number of times that word appears in `test_words`. We constructed the `vocabulary` earlier using the training set for our problem so encoding `test_words` is relatively straightforward.
def bow_encoding(words, vocabulary):
bow = [0] * len(vocabulary) # Start by setting the count for each word in the vocabulary to zero.
for word in words.split(): # For each word in the string
if word in vocabulary: # If the word is one that occurs in the vocabulary, increase its count.
bow[vocabulary[word]] += 1
return bow
test_bow = bow_encoding(test_words, vocabulary)
print(test_bow)
len(test_bow)
# So now we know how to construct a bag of words encoding of a user provided review, how to we send it to our endpoint? First, we need to start the endpoint back up.
xgb_predictor = xgb.deploy(initial_instance_count = 1, instance_type = 'ml.m4.xlarge')
# At this point we could just do the same thing that we did earlier when we tested our deployed model and send `test_bow` to our endpoint using the `xgb_predictor` object. However, when we eventually construct our Lambda function we won't have access to this object, so how do we call a SageMaker endpoint?
#
# It turns out that Python functions that are used in Lambda have access to another Amazon library called `boto3`. This library provides an API for working with Amazon services, including SageMaker. To start with, we need to get a handle to the SageMaker runtime.
# +
import boto3
runtime = boto3.Session().client('sagemaker-runtime')
# -
# And now that we have access to the SageMaker runtime, we can ask it to make use of (invoke) an endpoint that has already been created. However, we need to provide SageMaker with the name of the deployed endpoint. To find this out we can print it out using the `xgb_predictor` object.
xgb_predictor.endpoint
# Using the SageMaker runtime and the name of our endpoint, we can invoke the endpoint and send it the `test_bow` data.
response = runtime.invoke_endpoint(EndpointName = xgb_predictor.endpoint, # The name of the endpoint we created
ContentType = 'text/csv', # The data format that is expected
Body = test_bow)
# So why did we get an error?
#
# Because we tried to send the endpoint a list of integers but it expected us to send data of type `text/csv`. So, we need to convert it.
response = runtime.invoke_endpoint(EndpointName = xgb_predictor.endpoint, # The name of the endpoint we created
ContentType = 'text/csv', # The data format that is expected
Body = ','.join([str(val) for val in test_bow]).encode('utf-8'))
print(response)
# As we can see, the response from our model is a somewhat complicated looking dict that contains a bunch of information. The bit that we are most interested in is `'Body'` object which is a streaming object that we need to `read` in order to make use of.
response = response['Body'].read().decode('utf-8')
print(response)
# Now that we know how to process the incoming user data we can start setting up the infrastructure to make our simple web app work. To do this we will make use of two different services. Amazon's Lambda and API Gateway services.
#
# Lambda is a service which allows someone to write some relatively simple code and have it executed whenever a chosen trigger occurs. For example, you may want to update a database whenever new data is uploaded to a folder stored on S3.
#
# API Gateway is a service that allows you to create HTTP endpoints (url addresses) which are connected to other AWS services. One of the benefits to this is that you get to decide what credentials, if any, are required to access these endpoints.
#
# In our case we are going to set up an HTTP endpoint through API Gateway which is open to the public. Then, whenever anyone sends data to our public endpoint we will trigger a Lambda function which will send the input (in our case a review) to our model's endpoint and then return the result.
# ### Setting up a Lambda function
#
# The first thing we are going to do is set up a Lambda function. This Lambda function will be executed whenever our public API has data sent to it. When it is executed it will receive the data, perform any sort of processing that is required, send the data (the review) to the SageMaker endpoint we've created and then return the result.
#
# #### Part A: Create an IAM Role for the Lambda function
#
# Since we want the Lambda function to call a SageMaker endpoint, we need to make sure that it has permission to do so. To do this, we will construct a role that we can later give the Lambda function.
#
# Using the AWS Console, navigate to the **IAM** page and click on **Roles**. Then, click on **Create role**. Make sure that the **AWS service** is the type of trusted entity selected and choose **Lambda** as the service that will use this role, then click **Next: Permissions**.
#
# In the search box type `sagemaker` and select the check box next to the **AmazonSageMakerFullAccess** policy. Then, click on **Next: Review**.
#
# Lastly, give this role a name. Make sure you use a name that you will remember later on, for example `LambdaSageMakerRole`. Then, click on **Create role**.
# #### Part B: Create a Lambda function
#
# Now it is time to actually create the Lambda function. Remember from earlier that in order to process the user provided input and send it to our endpoint we need to gather two pieces of information:
#
# - The name of the endpoint, and
# - the vocabulary object.
#
# We will copy these pieces of information to our Lambda function after we create it.
#
# To start, using the AWS Console, navigate to the AWS Lambda page and click on **Create a function**. When you get to the next page, make sure that **Author from scratch** is selected. Now, name your Lambda function, using a name that you will remember later on, for example `sentiment_analysis_xgboost_func`. Make sure that the **Python 3.6** runtime is selected and then choose the role that you created in the previous part. Then, click on **Create Function**.
#
# On the next page you will see some information about the Lambda function you've just created. If you scroll down you should see an editor in which you can write the code that will be executed when your Lambda function is triggered. Collecting the code we wrote above to process a single review and adding it to the provided example `lambda_handler` we arrive at the following.
#
# ```python
# # We need to use the low-level library to interact with SageMaker since the SageMaker API
# # is not available natively through Lambda.
# import boto3
#
# # And we need the regular expression library to do some of the data processing
# import re
#
# REPLACE_NO_SPACE = re.compile("(\.)|(\;)|(\:)|(\!)|(\')|(\?)|(\,)|(\")|(\()|(\))|(\[)|(\])")
# REPLACE_WITH_SPACE = re.compile("(<br\s*/><br\s*/>)|(\-)|(\/)")
#
# def review_to_words(review):
# words = REPLACE_NO_SPACE.sub("", review.lower())
# words = REPLACE_WITH_SPACE.sub(" ", words)
# return words
#
# def bow_encoding(words, vocabulary):
# bow = [0] * len(vocabulary) # Start by setting the count for each word in the vocabulary to zero.
# for word in words.split(): # For each word in the string
# if word in vocabulary: # If the word is one that occurs in the vocabulary, increase its count.
# bow[vocabulary[word]] += 1
# return bow
#
#
# def lambda_handler(event, context):
#
# vocab = "*** ACTUAL VOCABULARY GOES HERE ***"
#
# words = review_to_words(event['body'])
# bow = bow_encoding(words, vocab)
#
# # The SageMaker runtime is what allows us to invoke the endpoint that we've created.
# runtime = boto3.Session().client('sagemaker-runtime')
#
# # Now we use the SageMaker runtime to invoke our endpoint, sending the review we were given
# response = runtime.invoke_endpoint(EndpointName = '***ENDPOINT NAME HERE***',# The name of the endpoint we created
# ContentType = 'text/csv', # The data format that is expected
# Body = ','.join([str(val) for val in bow]).encode('utf-8')) # The actual review
#
# # The response is an HTTP response whose body contains the result of our inference
# result = response['Body'].read().decode('utf-8')
#
# # Round the result so that our web app only gets '1' or '0' as a response.
# result = round(float(result))
#
# return {
# 'statusCode' : 200,
# 'headers' : { 'Content-Type' : 'text/plain', 'Access-Control-Allow-Origin' : '*' },
# 'body' : str(result)
# }
# ```
# Once you have copy and pasted the code above into the Lambda code editor, replace the `**ENDPOINT NAME HERE**` portion with the name of the endpoint that we deployed earlier. You can determine the name of the endpoint using the code cell below.
xgb_predictor.endpoint
# In addition, you will need to copy the vocabulary dict to the appropriate place in the code at the beginning of the `lambda_handler` method. The cell below prints out the vocabulary dict in a way that is easy to copy and paste.
print(str(vocabulary))
# Once you have added the endpoint name to the Lambda function, click on **Save**. Your Lambda function is now up and running. Next we need to create a way for our web app to execute the Lambda function.
# ### Setting up API Gateway
#
# Now that our Lambda function is set up, it is time to create a new API using API Gateway that will trigger the Lambda function we have just created.
#
# Using AWS Console, navigate to **Amazon API Gateway** and then click on **Get started**.
#
# On the next page, make sure that **New API** is selected and give the new api a name, for example, `sentiment_analysis_web_app`. Then, click on **Create API**.
#
# Now we have created an API, however it doesn't currently do anything. What we want it to do is to trigger the Lambda function that we created earlier.
#
# Select the **Actions** dropdown menu and click **Create Method**. A new blank method will be created, select its dropdown menu and select **POST**, then click on the check mark beside it.
#
# For the integration point, make sure that **Lambda Function** is selected and click on the **Use Lambda Proxy integration**. This option makes sure that the data that is sent to the API is then sent directly to the Lambda function with no processing. It also means that the return value must be a proper response object as it will also not be processed by API Gateway.
#
# Type the name of the Lambda function you created earlier into the **Lambda Function** text entry box and then click on **Save**. Click on **OK** in the pop-up box that then appears, giving permission to API Gateway to invoke the Lambda function you created.
#
# The last step in creating the API Gateway is to select the **Actions** dropdown and click on **Deploy API**. You will need to create a new Deployment stage and name it anything you like, for example `prod`.
#
# You have now successfully set up a public API to access your SageMaker model. Make sure to copy or write down the URL provided to invoke your newly created public API as this will be needed in the next step. This URL can be found at the top of the page, highlighted in blue next to the text **Invoke URL**.
# ## Step 7: Deploying our web app
#
# Now that we have a publicly available API, we can start using it in a web app. For our purposes, we have provided a simple static html file which can make use of the public api you created earlier.
#
# In the `website` folder there should be a file called `index.html`. Download the file to your computer and open that file up in a text editor of your choice. There should be a line which contains **\*\*REPLACE WITH PUBLIC API URL\*\***. Replace this string with the url that you wrote down in the last step and then save the file.
#
# Now, if you open `index.html` on your local computer, your browser will behave as a local web server and you can use the provided site to interact with your SageMaker model.
#
# If you'd like to go further, you can host this html file anywhere you'd like, for example using github or hosting a static site on Amazon's S3. Once you have done this you can share the link with anyone you'd like and have them play with it too!
#
# > **Important Note** In order for the web app to communicate with the SageMaker endpoint, the endpoint has to actually be deployed and running. This means that you are paying for it. Make sure that the endpoint is running when you want to use the web app but that you shut it down when you don't need it, otherwise you will end up with a surprisingly large AWS bill.
# ### Delete the endpoint
#
# Remember to always shut down your endpoint if you are no longer using it. You are charged for the length of time that the endpoint is running so if you forget and leave it on you could end up with an unexpectedly large bill.
xgb_predictor.delete_endpoint()
# ## Optional: Clean up
#
# The default notebook instance on SageMaker doesn't have a lot of excess disk space available. As you continue to complete and execute notebooks you will eventually fill up this disk space, leading to errors which can be difficult to diagnose. Once you are completely finished using a notebook it is a good idea to remove the files that you created along the way. Of course, you can do this from the terminal or from the notebook hub if you would like. The cell below contains some commands to clean up the created files from within the notebook.
# +
# First we will remove all of the files contained in the data_dir directory
# !rm $data_dir/*
# And then we delete the directory itself
# !rmdir $data_dir
# Similarly we remove the files in the cache_dir directory and the directory itself
# !rm $cache_dir/*
# !rmdir $cache_dir
# -
|
Tutorials/IMDB Sentiment Analysis - XGBoost - Web App.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# # Réseau de neurones pour la régression
# Nous allons ici appliquer un réseau de neurones à un problème de régression. Nous allons essayer de faire un modèle pour estimer les prix des appartements en Suisse. Pour cela, nous utilisons un jeu de données construit à partir d'annonces d'appartements pour la location. Ce jeu de données contient notamment le nombre de pièces, la surface habitable en mètres carrés, la latitude, longitude et évidemment le loyer par mois en francs.
#
# Pour comparaison, nous allons également faire un modèle de régression linéaire.
# ## Lecture des données
d = read.table(file="data/appartements.tsv", header=TRUE, sep="\t")
head(d)
summary(d)
nrow(d)
# Nous avons les données pour presque 15'000 appartements.
plot(d$loyer_mois, d$surface_habitable)
# ## Préparation du jeu de données
# Il est important de normaliser les données avant d'entraîner un réseau de neurones. Si les données ne sont pas normalisées, le réseau a plus de difficultés de converger vers un résultat acceptable.
#
# Il y a beaucoup de façons de normaliser les données dans R. Le but est de faire en sorte que pour chaque variable, les valeurs se situent entre 0 et 1 (ou parfois entre -1 et 1).
#
# Dans un premier temps, nous calculons les valeurs minimum et maximum pour chaque variable dans notre jeu de données (à part la première colonne qui est l'identifiant unique de l'appartement):
maxs = apply(d[,2:6], 2, max)
mins = apply(d[,2:6], 2, min)
maxs
mins
# Nous pouvons maintenant normaliser nos données. En gros, pour une valeur `x` donnée, nous faisons le calcul `(x - min) / (max - min)`, et ceci pour chaque valeur et pour toutes les variables (les valeurs minimales et maximales sont par variable évidemment).
#
# La fonction `scale` permet de faire cette opération pour toutes les variables en même temps:
dnorm = as.data.frame(scale(d[,2:6], center=mins, scale=maxs-mins))
head(dnorm)
# ## Régression linéaire
# Essayons de faire une simple régression multiple pour estimer le loyer des appartements. En principe, nous pouvons utiliser les données sans normalisation pour une régression linéaire. Cepedant, puisque nous allons comparer le résultat avec celui d'un réseau de neurones, nous utilisons dores et déjà les valeurs normalisées.
regmul = lm(loyer_mois ~ surface_habitable + nombre_pieces, data=dnorm)
summary(regmul)
# Les résultat est une régression avec un $R^2$ de 0.56. Nous voyons par ailleurs que le facteur principal pour le loyer d'un appartement et la surface habitable. Le nombre de pièces a un petit effect négatif (un appartement avec plus de pièces étant un peu moins cher qu'un appartement de même taille mais moins de pièces).
# Nous pouvons encore essayer d'inclure l'effet de la localisation. En effet, nous pouvons penser qu'en fonction de la localisation de l'appartement, le loyer pourrait varier. Nous pouvons faire ceci en simplement incluant les coordonnées géographiques:
regmulloc = lm(loyer_mois ~ surface_habitable + nombre_pieces + lat + lng, data=dnorm)
summary(regmulloc)
# On arrive effectivement à expliquer un peu plus de variance, avec un $R^2$ de 0.57. Cependant, la différence est quand-même minime. Nous voyons aussi que la longitude n'a quasiment pas d'effet, et la latitude seulement un effet minimal. Plus on va au Sud, plus les appartements sont un poil plus cher selon ce modèle.
# ## Réseau de neurones
# Avec un réseau de neurones on doit séparer notre jeu de données en jeu d'entrainement et jeu de test, comme on l'avait déjà fait pour la régression logistique. Vu que nous avons un nombre plus important de données, nous pouvons réserver 25% des données pour le test, et utiliser seulement 75% pour l'entrainement:
idx = sample(nrow(dnorm), size=0.75*nrow(dnorm))
dtrain = dnorm[idx,]
dtest = dnorm[-idx,]
c(nrow(dtrain), nrow(dtest))
# Nous avons donc environ 11'000 appartements pour l'entrainement et 3600 pour tester les modèle.
#
# Nous allons utiliser le package `neuralnet` pour construire notre réseau de neurones. Nous allons utiliser un autre package, `caret` pour calculer le $R^2$ à travers la fonction `postResample`.
library(neuralnet)
library(caret)
# Nous construisons un réseau avec deux couches, la première avec 8 neurones et la deuxième avec 4 neurones. L'entrainement peut prendre un petit moment...
nn = neuralnet(
loyer_mois ~ surface_habitable + nombre_pieces,
data=dtrain,
hidden=c(30,10),
linear.output=TRUE
)
# Nous calculons maintenant les prédictions (donc l'estimation du loyer) pour le jeu de donneés test.
pred_nn = predict(nn, dtest)
# Nous allons maintenant calculer le $R^2$ pour pouvoir comparer avec la régression multiple. Pour cela, nous allons calculer le loyer par mois non normalisé, à la fois les valeurs estimées par le modèle et les valeurs réelles, pour le jeu de données test:
yhat = pred_nn * (max(d$loyer_mois) - min(d$loyer_mois)) + min(d$loyer_mois)
y = dtest[,5] * (max(d$loyer_mois) - min(d$loyer_mois)) + min(d$loyer_mois)
# Nous pouvons maintenant estimer le $R^2$ avec la fonction `postResample`:
postResample(yhat, y)
# Nous obtenons une valeur d'environ 0.58, donc un peu meilleur mais loin d'être bon.
# À partir de là, vous pouvez essayer de changer un peu l'architecture du réseau, ou encore d'enlever des variables, afin d'optimiser le modèle.
|
60-ann/ann.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Atul's super simple neural net
#
# _October 8, 2017_
#
# This workbook contains an implementation of a ridiculously simple neural net with one hidden layer that has two units, and one unit in the output layer. The sigmoid activation function is used in both layers.
import numpy as np
from numpy.testing import assert_almost_equal
# First, let's define the sigmoid function and its gradient...
# +
def sigmoid(x):
return 1 / (1 + np.power(np.e, -x))
# Sanity check.
assert sigmoid(0) == 0.5
assert_almost_equal(sigmoid(-100), 0)
assert_almost_equal(sigmoid(100), 1)
def sigmoid_gradient(x):
return sigmoid(x) * (1 - sigmoid(x))
# Sanity check.
assert_almost_equal(sigmoid_gradient(-100), 0)
assert_almost_equal(sigmoid_gradient(100), 0)
# -
# Now let's define a function that runs forward propagation on a neural net, given the weights and biases for both layers.
#
# In implementing this, I found Coursera's deeplearning.ai course on [Neural Networks and Deep Learning](https://www.coursera.org/learn/neural-networks-deep-learning) useful; all notation is generally taken from that class.
# +
def forward_prop_nn(w1, b1, w2, b2, x):
z1 = np.dot(w1, x) + b1
a1 = sigmoid(z1)
z2 = np.dot(w2, a1) + b2
a2 = sigmoid(z2)
return (z1, a1, z2, a2)
def predict_nn(w1, b1, w2, b2, x):
return forward_prop_nn(w1, b1, w2, b2, x)[-1]
# -
# Here we will manually create a neural net to run the XNOR boolean operation. XNOR is just the negation of XOR, i.e. it will be true if _both_ its inputs are true or false, and false otherwise.
xnor_w_1 = np.array([
[-20, -20], # Weights for "(NOT x[0]) AND (NOT x[1])"
[ 20, 20], # Weights for "x[0] AND x[1]"
]) * 10
xnor_b_1 = np.array([
[ 10], # Bias for "(NOT x[0]) AND (NOT x[1])"
[-30], # Bias for "x[0] AND x[1]"
]) * 10
xnor_w_2 = np.array([
[ 20, 20], # Weights for "x[0] OR x[1]"
]) * 10
xnor_b_2 = np.array([
[-10], # Bias for "x[0] OR x[1]"
]) * 10
# Now let's make sure our manually-constructed NN matches our intuitive expectations of XNOR.
# +
# Define a "truth table" for our XNOR function. We'll use this to make sure our NN
# works, and we'll also use it later as training data.
boolean_xnor_truth_table = [
# x[0] x[1] y
[(True , True ), True ],
[(False, False), True ],
[(False, True ), False],
[(True , False), False]
]
# This is a numpy-friendly version of our truth table, where each item is
# a tuple consisting of a 2x1 array representing the input (x) and a 1x1
# array representing the output (y).
xnor_truth_table = [
(np.array(x, dtype=float).reshape(2, 1),
np.array([[y]], dtype=float))
for (x, y) in boolean_xnor_truth_table
]
# Test our NN to make sure everything works.
for x, y in xnor_truth_table:
assert_almost_equal(predict_nn(xnor_w_1, xnor_b_1, xnor_w_2, xnor_b_2, x), y)
# -
# Now let's define a function to train a neural net!
#
# This is intentionally un-vectorized because I wanted to make sure I understood the algorithm before dealing with vectorization. Thus the `examples` parameter is just a Python list of tuples containing a 2x1 numpy array and an expected 1x1 output.
#
# The Coursera class' [Backpropagation intuition](https://www.coursera.org/learn/neural-networks-deep-learning/lecture/6dDj7/backpropagation-intuition-optional) lecture was particularly helpful in understanding the math behind this. I supplemented my understanding with [Khan Academy's AP Calculus AB](https://www.khanacademy.org/math/ap-calculus-ab) when needed because I am very rusty at Calculus.
# +
def cost_func(predicted_y, y):
return -y * np.log(predicted_y) - (1 - y) * np.log(1 - predicted_y)
def train_nn(examples, iterations, learning_rate, check_gradient=None, print_cost=True):
m = len(examples)
np.random.seed(1)
# Initialize our weights and biases. Note that the weights need to
# be randomly initialized so we can break symmetry.
w1 = np.random.rand(2, 2)
b1 = np.zeros([2, 1])
w2 = np.random.rand(1, 2)
b2 = np.zeros([1, 1])
for i in range(iterations):
dw1 = np.zeros([2, 2])
db1 = np.zeros([2, 1])
dw2 = np.zeros([1, 2])
db2 = np.zeros([1, 1])
cost = np.zeros([1, 1])
for x, y in examples:
# Forward propagation.
z1, a1, z2, a2 = forward_prop_nn(w1, b1, w2, b2, x)
# Calculate the cost of our output by comparing it to the
# expected output.
cost += cost_func(a2, y)
# Back propagation.
dz2 = a2 - y
dw2 += np.dot(dz2, a1.T)
db2 += dz2
dz1 = np.dot(w2.T, dz2) * sigmoid_gradient(z1)
dw1 += np.dot(dz1, x.T)
db1 += dz1
dw1 /= m
db1 /= m
dw2 /= m
db2 /= m
cost /= m
if check_gradient is not None:
check_gradient(w1, b1, w2, b2, examples, dw1, db1, dw2, db2)
w1 -= learning_rate * dw1
b1 -= learning_rate * db1
w2 -= learning_rate * dw2
b2 -= learning_rate * db2
if i % 100 == 0 and print_cost:
print(f"cost at iteration {i}: {cost[0][0]}")
return (w1, b1, w2, b2)
# -
# Now let's train a neural net to learn the XNOR operation.
#
# This is obviously a stupid use of a neural net, but I wanted a trivial use case to make sure I understood how things work.
# +
print("Training neural net...\n")
# Reuse our truth table as our training data.
w1, b1, w2, b2 = train_nn(xnor_truth_table, 5000, 1)
# Test our NN to make sure it produces the same responses as our truth table.
# Note that normally a NN classifier would use some sort of thresholding
# to determine whether its outputs are true or false, but here we'll just
# directly compare its output to the expected truth table value to two
# decimal places, because our NN happens to be that awesome.
print(f"\nTraining complete. Verifying predictions...\n")
for x, y in xnor_truth_table:
print(f"{x} should be approximately {float(y)}...")
y_hat = predict_nn(w1, b1, w2, b2, x)
assert_almost_equal(y_hat, y, decimal=2)
print(f" ✓ Prediction is {float(y_hat)}, hooray!\n")
# -
# Hooray! But how do we know our gradient descent math is correct?
#
# We can figure this out with gradient checking!
# +
def check_gradient(w1, b1, w2, b2, examples, check_dw1, check_db1, check_dw2, check_db2):
epsilon = 0.0001
m = len(examples)
theta = np.concatenate((
w1.reshape(-1, 1),
b1.reshape(-1, 1),
w2.reshape(-1, 1),
b2.reshape(-1, 1),
))
costs_left = np.zeros([len(theta), 1])
costs_right = np.copy(costs_left)
def unrolled_predict_nn(theta, x):
w1 = theta[0:4].reshape(2, 2)
b2 = theta[4:6].reshape(2, 1)
w2 = theta[6:8].reshape(1, 2)
b2 = theta[8:9].reshape(1, 1)
return predict_nn(w1, b1, w2, b2, x)
for x, y in examples:
for i in range(len(theta)):
theta_left = np.copy(theta)
theta_left[i] -= epsilon
theta_right = np.copy(theta)
theta_right[i] += epsilon
costs_left[i] += cost_func(unrolled_predict_nn(theta_left, x), y)[0]
costs_right[i] += cost_func(unrolled_predict_nn(theta_right, x), y)[0]
costs_left /= m
costs_right /= m
theta_prime = (costs_right - costs_left) / (2 * epsilon)
dw1 = theta_prime[0:4].reshape(2, 2)
db1 = theta_prime[4:6].reshape(2, 1)
dw2 = theta_prime[6:8].reshape(1, 2)
db2 = theta_prime[8:9].reshape(1, 1)
assert_almost_equal(dw1, check_dw1)
assert_almost_equal(db1, check_db1, decimal=2) # TODO: Why the disparity here?
assert_almost_equal(dw2, check_dw2)
assert_almost_equal(db2, check_db2)
train_nn(xnor_truth_table, 10, 1, check_gradient=check_gradient, print_cost=False)
print("Gradients check out OK!")
# -
|
super-simple-nn.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="SB93Ge748VQs" colab_type="text"
# ##### Copyright 2019 The TensorFlow Authors.
# + id="0sK8X2O9bTlz" colab_type="code" cellView="both" colab={}
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="HEYuO5NFwDK9" colab_type="text"
# # Examining the TensorFlow Graph
#
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/tensorboard/graphs"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/tensorboard/blob/master/docs/graphs.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/tensorboard/blob/master/docs/graphs.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# </table>
# + [markdown] id="56V5oun18ZdZ" colab_type="text"
# ## Overview
#
# TensorBoard’s **Graphs dashboard** is a powerful tool for examining your TensorFlow model. You can quickly view a conceptual graph of your model’s structure and ensure it matches your intended design. You can also view a op-level graph to understand how TensorFlow understands your program. Examining the op-level graph can give you insight as to how to change your model. For example, you can redesign your model if training is progressing slower than expected.
# + [markdown] id="TOSJ-4nteBYG" colab_type="text"
# This tutorial presents a quick overview of how to generate graph diagnostic data and visualize it in TensorBoard’s Graphs dashboard. You’ll define and train a simple Keras Sequential model for the Fashion-MNIST dataset and learn how to log and examine your model graphs. You will also use a tracing API to generate graph data for functions created using the new `tf.function` annotation.
# + [markdown] id="zNI1-dflrAo0" colab_type="text"
# ## Setup
# + id="6B95Hb6YVgPZ" colab_type="code" colab={}
try:
# # %tensorflow_version only exists in Colab.
# %tensorflow_version 2.x
except Exception:
pass
# Load the TensorBoard notebook extension.
# %load_ext tensorboard
# + id="_wqSAZExy6xV" colab_type="code" outputId="4558e63e-513d-4f64-dc1c-92937710e68c" colab={"base_uri": "https://localhost:8080/", "height": 34}
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
from datetime import datetime
from packaging import version
import tensorflow as tf
from tensorflow import keras
print("TensorFlow version: ", tf.__version__)
assert version.parse(tf.__version__).release[0] >= 2, \
"This notebook requires TensorFlow 2.0 or above."
# + id="Ao7fJW1Pyiza" colab_type="code" colab={}
# Clear any logs from previous runs
# !rm -rf ./logs/
# + [markdown] id="e25E37vd1xEW" colab_type="text"
# ## Define a Keras model
#
# In this example, the classifier is a simple four-layer Sequential model.
# + id="skqORzvE3Egy" colab_type="code" colab={}
# Define the model.
model = keras.models.Sequential([
keras.layers.Flatten(input_shape=(28, 28)),
keras.layers.Dense(32, activation='relu'),
keras.layers.Dropout(0.2),
keras.layers.Dense(10, activation='softmax')
])
model.compile(
optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
# + [markdown] id="qbjuoz9E3VC_" colab_type="text"
# Download and prepare the training data.
# + id="6TDmc41z3g38" colab_type="code" colab={}
(train_images, train_labels), _ = keras.datasets.fashion_mnist.load_data()
train_images = train_images / 255.0
# + [markdown] id="8DV0xibO3bRC" colab_type="text"
# ## Train the model and log data
#
# Before training, define the [Keras TensorBoard callback](https://www.tensorflow.org/api_docs/python/tf/keras/callbacks/TensorBoard), specifying the log directory. By passing this callback to Model.fit(), you ensure that graph data is logged for visualization in TensorBoard.
# + id="TU_L_u9SqQdH" colab_type="code" outputId="210b42ed-25ea-4c78-d4c1-2ddab616b175" colab={"base_uri": "https://localhost:8080/", "height": 204}
# Define the Keras TensorBoard callback.
logdir="logs/fit/" + datetime.now().strftime("%Y%m%d-%H%M%S")
tensorboard_callback = keras.callbacks.TensorBoard(log_dir=logdir)
# Train the model.
model.fit(
train_images,
train_labels,
batch_size=64,
epochs=5,
callbacks=[tensorboard_callback])
# + [markdown] id="IRX5OIsi4TTV" colab_type="text"
# ## Op-level graph
#
# Start TensorBoard and wait a few seconds for the UI to load. Select the Graphs dashboard by tapping “Graphs” at the top.
# + id="PFgFjlPEqXb9" colab_type="code" colab={}
# %tensorboard --logdir logs
# + [markdown] id="EGlOqRp54ufD" colab_type="text"
# By default, TensorBoard displays the **op-level graph**. (On the left, you can see the “Default” tag selected.) Note that the graph is inverted; data flows from bottom to top, so it’s upside down compared to the code. However, you can see that the graph closely matches the Keras model definition, with extra edges to other computation nodes.
#
# Graphs are often very large, so you can manipulate the graph visualization:
#
# * Scroll to **zoom** in and out
# * Drag to **pan**
# * Double clicking toggles **node expansion** (a node can be a container for other nodes)
#
# You can also see metadata by clicking on a node. This allows you to see inputs, outputs, shapes and other details.
#
# + [markdown] id="F-2yw5qd7OpK" colab_type="text"
# <img class="tfo-display-only-on-site" src="https://github.com/tensorflow/tensorboard/blob/master/docs/images/graphs_computation.png?raw=1"/>
# + [markdown] id="jDRynpVw53SJ" colab_type="text"
# <img class="tfo-display-only-on-site" src="https://github.com/tensorflow/tensorboard/blob/master/docs/images/graphs_computation_detail.png?raw=1"/>
# + [markdown] id="Oj9FSPdz6SO2" colab_type="text"
# ## Conceptual graph
#
# In addition to the execution graph, TensorBoard also displays a **conceptual graph**. This is a view of just the Keras model. This may be useful if you’re reusing a saved model and you want to examine or validate its structure.
#
# To see the conceptual graph, select the “keras” tag. For this example, you’ll see a collapsed **Sequential** node. Double-click the node to see the model’s structure:
# + [markdown] id="Qw9rbEcE6eZB" colab_type="text"
# <img class="tfo-display-only-on-site" src="https://github.com/tensorflow/tensorboard/blob/master/docs/images/graphs_tag_selection.png?raw=1"/> <br/>
# <img class="tfo-display-only-on-site" src="https://github.com/tensorflow/tensorboard/blob/master/docs/images/graphs_conceptual.png?raw=1"/>
# + [markdown] id="zVuaKBifu-qF" colab_type="text"
# ## Graphs of tf.functions
#
# The examples so far have described graphs of Keras models, where the graphs have been created by defining Keras layers and calling Model.fit().
#
# You may encounter a situation where you need to use the `tf.function` annotation to ["autograph"](https://www.tensorflow.org/guide/function), i.e., transform, a Python computation function into a high-performance TensorFlow graph. For these situations, you use **TensorFlow Summary Trace API** to log autographed functions for visualization in TensorBoard.
# + [markdown] id="JIuhJnQ8w-dT" colab_type="text"
# To use the Summary Trace API:
#
# * Define and annotate a function with `tf.function`
# * Use `tf.summary.trace_on()` immediately before your function call site.
# * Add profile information (memory, CPU time) to graph by passing `profiler=True`
# * With a Summary file writer, call `tf.summary.trace_export()` to save the log data
#
# You can then use TensorBoard to see how your function behaves.
#
# + id="woI67Stgv_uY" colab_type="code" colab={}
# The function to be traced.
@tf.function
def my_func(x, y):
# A simple hand-rolled layer.
return tf.nn.relu(tf.matmul(x, y))
# Set up logging.
stamp = datetime.now().strftime("%Y%m%d-%H%M%S")
logdir = 'logs/func/%s' % stamp
writer = tf.summary.create_file_writer(logdir)
# Sample data for your function.
x = tf.random.uniform((3, 3))
y = tf.random.uniform((3, 3))
# Bracket the function call with
# tf.summary.trace_on() and tf.summary.trace_export().
tf.summary.trace_on(graph=True, profiler=True)
# Call only one tf.function when tracing.
z = my_func(x, y)
with writer.as_default():
tf.summary.trace_export(
name="my_func_trace",
step=0,
profiler_outdir=logdir)
# + id="zCArnWzP0VuZ" colab_type="code" colab={}
# %tensorboard --logdir logs/func
# + [markdown] id="WDl1PBFQ64xi" colab_type="text"
# <img class="tfo-display-only-on-site" src="https://github.com/tensorflow/tensorboard/blob/master/docs/images/graphs_autograph.png?raw=1"/>
# + [markdown] id="1pLRaf3q6Nku" colab_type="text"
# You can now see the structure of your function as understood by TensorBoard. Click on the "Profile" radiobutton to see CPU and memory statistics.
|
docs/graphs.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
print('Hello world')
S0=100.0 # spot stock price
K=100.0 # strike
T=1.0 # maturity
r=0.1 # risk free rate
sig=0.2 # diffusion coefficient or volatility
# -
#
# # Hello world
# Are
# $$
# a, b \in \mathbb{R}, then applies (a+b)^{2} = a^{2} + ab + b^{2}
# $$
#
# +
rint('hello world21')
|
tests/b.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import sys
sys.path.append(os.path.join('..', 'src'))
import pickle
import time
import numpy as np
import matplotlib.pyplot as plt
plt.rcParams['figure.figsize'] = (7, 5)
import seaborn as sns
sns.set()
# -
DATA_DIR = os.path.join('..', 'data', 'processed', 'filtering')
# +
with open(os.path.join(DATA_DIR, 'user_to_items.pickle'), 'rb') as file:
user_to_items = pickle.load(file)
with open(os.path.join(DATA_DIR, 'item_to_users.pickle'), 'rb') as file:
item_to_users = pickle.load(file)
with open(os.path.join(DATA_DIR, 'train_ratings.pickle'), 'rb') as file:
train_ratings = pickle.load(file)
with open(os.path.join(DATA_DIR, 'test_ratings.pickle'), 'rb') as file:
test_ratings = pickle.load(file)
# -
ratings = np.fromiter(train_ratings.values(), dtype=np.float32)
sns.countplot(ratings, color='#a2c5e3')
plt.xlabel("Rating value")
plt.ylabel("Number of ratings")
plt.title("Distribution of ratings")
plt.savefig("rate-dist.png", pad_inches=0.1)
user_ratings_number = np.array([len(items) for items in user_to_items.values()])
sns.distplot(user_ratings_number)
plt.xlabel("Number of ratings")
plt.ylabel("Fraction of users")
plt.title("Distribution of number of ratings per user")
plt.savefig("user_dist.png")
item_ratings_number = np.array([len(users) for users in item_to_users.values()])
sns.distplot(item_ratings_number, kde=True)
plt.xlabel("Number of ratings")
plt.ylabel("Fraction of items")
plt.title("Distribution of number of ratings per item")
plt.savefig("item_dist.png")
item_ratings_average = np.array([
np.mean([
train_ratings[(user, item)] for user in users
])
for item, users in item_to_users.items()
])
sns.distplot(item_ratings_average, kde=False)
plt.xlabel("Mean rating")
plt.ylabel("Number of items")
plt.title("Distribution of average ratings per item")
user_ratings_average = np.array([
np.mean([
train_ratings[(user, item)] for item in items
])
for user, items in user_to_items.items()
])
sns.distplot(user_ratings_average, kde=False)
plt.xlabel("Mean rating")
plt.ylabel("Number of users")
plt.title("Distribution of average ratings per user")
|
notebook/DataExploration.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # SVM - Evaluation On Datasets
# This notebook analyzes on multiple datasets. This notebook
# considers whether classifiers in ensembles were trained on
# features with values compatible with the tests vector.
# **TO DO**
# 1. Check the data to see if trained on Normoxia.
# 1. Also, was training done on individual replicas? Are the data all features or just regulators?
# 1. Bar plot presentation of evaluations
# 1. Filter ensembles if test_vector is not contained in the training data.
# # Preliminaries
# +
import init
from common import constants as cn
from common.trinary_data import TrinaryData
from common.data_provider import DataProvider
from common_python.plots import util_plots
from common_python.classifier import classifier_ensemble
from common_python.classifier import classifier_collection
from common import transform_data
import collections
import os
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import sklearn
from sklearn.decomposition import PCA
from sklearn import svm
from sklearn.model_selection import cross_val_score
# %matplotlib inline
# -
# # Data
# Data used in the analysis.
DATA = TrinaryData(is_regulator=False, is_averaged=True, is_dropT1=True)
DATA.df_X.head()
df_sampleAM = transform_data.trinaryReadsDF(
csv_file="AM_MDM_Mtb_transcripts_DEseq.csv", is_time_columns=False, is_display_errors=False)
df_sampleAW = transform_data.trinaryReadsDF(csv_file="AW_plus_v_AW_neg_Mtb_transcripts_DEseq.csv",
is_time_columns=False, is_display_errors=False)
df_sampleAM = df_sampleAM.T
df_sampleAW.columns
df_sampleAW = df_sampleAW.T
df_sampleAW.head()
# # Classification Validations of Controlled Samples
# Classify T2-T25 and see if result is same as original class. Use 5-fold cross validation, where there is a holdout for each class and the selection is random.
#
accuracy_dct = {}
for rank in [1, 2, 4, 8, 16, 32]:
accuracy_dct[rank] = classifier_ensemble.ClassifierEnsemble.crossValidate(
DATA, num_iter=10, num_holdout=1, filter_high_rank=rank)
plt.plot(list(accuracy_dct.keys()), list(accuracy_dct.values()))
plt.ylim([0, 1.1])
plt.xlabel("No classifiers in ensemble")
plt.ylabel("accuracy")
# # Classification of Lab Samples
svm_ensemble = classifier_ensemble.ClassifierEnsemble(
classifier_ensemble.ClassifierDescriptorSVM(), filter_high_rank=15, size=30)
df_X = DATA.df_X.copy()
df_X.columns = DATA.features
svm_ensemble.fit(df_X, DATA.ser_y)
svm_ensemble.predict(df_sampleAM)
svm_ensemble.predict(df_sampleAW)
# # Support for Prediction
|
notebooks/SVM - Evaluations on datasets.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3-azureml
# kernelspec:
# display_name: Python 3.7.9 64-bit
# metadata:
# interpreter:
# hash: 04635d289a519a1410467dd0afb0db42f9184808881ca68b2eb5a687a20a5a94
# name: Python 3.7.9 64-bit
# ---
# + [markdown] nteract={"transient": {"deleting": false}}
# Copyright (c) Microsoft Corporation. All rights reserved. Licensed under the MIT License. # Use Azure Percept MM Python SDK in Jupyter Notebook
#
# Before Azure Percept MM Python SDK is formally published, you need to install it from a local wheel file, which can be found under the same folder where this workbook resides.
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}} gather={"logged": 1599976565124}
import sys
# !$sys.executable -m pip uninstall -y sczpy
# !$sys.executable -m pip install sczpy-0.0.7-py3-none-any.whl
# + [markdown] nteract={"transient": {"deleting": false}}
# Then, you need to declare a few environment variables that defines the Azure Service Principal to be used for authentication. If you've used the PowerShell script to provision your Azure Percept MM server, you should use the same service principal provisioned by the script.
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
# %env AZURE_CLIENT_ID=
# %env AZURE_CLIENT_SECRET=
# %env AZURE_TENANT_ID=
# + [markdown] nteract={"transient": {"deleting": false}}
# Next, create a new Azure Percept MM client using your Azure Percept MM server URL
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}} gather={"logged": 1599976920088}
import sczpy
server_url=
client = sczpy.SCZClient(server_url)
# + [markdown] nteract={"transient": {"deleting": false}}
# An AI model version needs to be registered with the Azure Percept MM server:
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}} gather={"logged": 1599977851607}
model_name = "my-model"
model_version = "v1"
client.register_model(model_name, model_version)
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}} gather={"logged": 1599977523456}
client.encrypt(model_name, model_version, "model.txt", "model.txt.enc")
client.decrypt(model_name, model_version, "model.txt.enc", "model.decrypted.txt")
client.upload_model(model_name, model_version, "model.txt.enc")
client.download_model(model_name, model_version, "downloaded.txt.enc")
client.decrypt(model_name, model_version, "downloaded.txt.enc", "downloaded.decrypted.txt")
|
Sample-Scripts-and-Notebooks/Official/Secured Locker/jupyter-basics/sczpy-basics.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="9zF7SQGjbxbJ" colab_type="code" outputId="8b5f9039-7870-44b4-8715-284cab2dd933" executionInfo={"status": "ok", "timestamp": 1581719190505, "user_tz": -60, "elapsed": 9209, "user": {"displayName": "<NAME>", "photoUrl": "https://lh6.googleusercontent.com/-Mk6sDG3z8U8/AAAAAAAAAAI/AAAAAAAATPI/zF3jRAXRQtg/s64/photo.jpg", "userId": "13278186009456793751"}} colab={"base_uri": "https://localhost:8080/", "height": 269}
# !pip install eli5
# + id="4lw-o6fYbzm_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 166} outputId="fb8294ea-8d23-4950-b101-bd73f7013e3e" executionInfo={"status": "ok", "timestamp": 1581719196001, "user_tz": -60, "elapsed": 4108, "user": {"displayName": "<NAME>", "photoUrl": "https://lh6.googleusercontent.com/-Mk6sDG3z8U8/AAAAAAAAAAI/AAAAAAAATPI/zF3jRAXRQtg/s64/photo.jpg", "userId": "13278186009456793751"}}
import pandas as pd
import numpy as np
from sklearn.tree import DecisionTreeRegressor
from sklearn.ensemble import RandomForestRegressor
from sklearn.metrics import mean_absolute_error
from sklearn.model_selection import cross_val_score
import eli5
from eli5.sklearn import PermutationImportance
from ast import literal_eval
from tqdm import tqdm_notebook
# + id="LW0OX8swcvpM" colab_type="code" outputId="34436c21-b3c0-467b-eaa0-e8c4bc84a769" executionInfo={"status": "ok", "timestamp": 1581719202880, "user_tz": -60, "elapsed": 1607, "user": {"displayName": "<NAME>", "photoUrl": "https://lh6.googleusercontent.com/-Mk6sDG3z8U8/AAAAAAAAAAI/AAAAAAAATPI/zF3jRAXRQtg/s64/photo.jpg", "userId": "13278186009456793751"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# cd "/content/drive/My Drive/Colab Notebooks/dw_matrix"
# + id="1xQtGKfQc34N" colab_type="code" colab={}
df = pd.read_csv('data/men_shoes.csv', low_memory=False)
# + id="AnBAAIn4c456" colab_type="code" colab={}
def run_model(feats, model = DecisionTreeRegressor(max_depth=5)):
X = df[feats].values
y = df['prices_amountmin'].values
model = DecisionTreeRegressor(max_depth=5)
scores = cross_val_score(model, X, y, scoring='neg_mean_absolute_error')
return np.mean(scores), np.std(scores)
# + id="p49gxxZQ9fqj" colab_type="code" outputId="e0fb1689-a806-4e3b-d36c-86e60132323d" executionInfo={"status": "ok", "timestamp": 1581719217348, "user_tz": -60, "elapsed": 1075, "user": {"displayName": "<NAME>", "photoUrl": "https://lh6.googleusercontent.com/-Mk6sDG3z8U8/AAAAAAAAAAI/AAAAAAAATPI/zF3jRAXRQtg/s64/photo.jpg", "userId": "13278186009456793751"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# df['brand_cat'] = df['brand'].factorize()[0]
# run_model(['brand_cat'])
df['brand_cat'] = df['brand'].map(lambda x: str(x).lower()).factorize()[0]
run_model(['brand_cat'])
# + id="Vunqyc4v9zuv" colab_type="code" outputId="35ee7278-e602-47d3-97d3-6d3959845911" executionInfo={"status": "ok", "timestamp": 1581719220503, "user_tz": -60, "elapsed": 1052, "user": {"displayName": "<NAME>", "photoUrl": "https://lh6.googleusercontent.com/-Mk6sDG3z8U8/AAAAAAAAAAI/AAAAAAAATPI/zF3jRAXRQtg/s64/photo.jpg", "userId": "13278186009456793751"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
model = RandomForestRegressor(max_depth=5, n_estimators=100, random_state=0)
run_model(['brand_cat'], model)
# + id="IuVLIq5b-PQv" colab_type="code" outputId="19cd4999-f0d9-4401-af94-2e45934967d8" executionInfo={"status": "ok", "timestamp": 1581719223504, "user_tz": -60, "elapsed": 1485, "user": {"displayName": "<NAME>", "photoUrl": "https://lh6.googleusercontent.com/-Mk6sDG3z8U8/AAAAAAAAAAI/AAAAAAAATPI/zF3jRAXRQtg/s64/photo.jpg", "userId": "13278186009456793751"}} colab={"base_uri": "https://localhost:8080/", "height": 138}
df.features.head().values
# + id="iEC1OUIE-hmf" colab_type="code" outputId="9f22549c-ea96-4132-a1bc-9569e6b9afee" executionInfo={"status": "ok", "timestamp": 1581719226176, "user_tz": -60, "elapsed": 1508, "user": {"displayName": "<NAME>", "photoUrl": "https://lh6.googleusercontent.com/-Mk6sDG3z8U8/AAAAAAAAAAI/AAAAAAAATPI/zF3jRAXRQtg/s64/photo.jpg", "userId": "13278186009456793751"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
test = {'key': 'value'}
test['key']
str(test)
# + id="WvhBZBm9_10h" colab_type="code" outputId="e2bc83a4-20ad-4e00-e7ce-2d93decbc75c" executionInfo={"status": "ok", "timestamp": 1581719228226, "user_tz": -60, "elapsed": 979, "user": {"displayName": "<NAME>", "photoUrl": "https://lh6.googleusercontent.com/-Mk6sDG3z8U8/AAAAAAAAAAI/AAAAAAAATPI/zF3jRAXRQtg/s64/photo.jpg", "userId": "13278186009456793751"}} colab={"base_uri": "https://localhost:8080/", "height": 118}
str_dict = '[{"key":"Gender","value":["Men"]},{"key":"Shoe Size","value":["M"]},{"key":"Shoe Category","value":["Men\'s Shoes"]},{"key":"Color","value":["Multicolor"]},{"key":"Manufacturer Part Number","value":["8190-W-NAVY-7.5"]},{"key":"Brand","value":["Josmo"]}]'
literal_eval(str_dict)
# + id="nUypGu4NAQKx" colab_type="code" outputId="6fe1706c-e325-4900-8522-ac0f735f60be" executionInfo={"status": "ok", "timestamp": 1581719230842, "user_tz": -60, "elapsed": 981, "user": {"displayName": "<NAME>", "photoUrl": "https://lh6.googleusercontent.com/-Mk6sDG3z8U8/AAAAAAAAAAI/AAAAAAAATPI/zF3jRAXRQtg/s64/photo.jpg", "userId": "13278186009456793751"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
literal_eval(str_dict)[0]['key']
# + id="DU-HDRS1Ahpw" colab_type="code" outputId="a8af9598-08ab-4e08-cf7b-71eaa432f7cc" executionInfo={"status": "ok", "timestamp": 1581719233383, "user_tz": -60, "elapsed": 1067, "user": {"displayName": "<NAME>", "photoUrl": "https://lh6.googleusercontent.com/-Mk6sDG3z8U8/AAAAAAAAAAI/AAAAAAAATPI/zF3jRAXRQtg/s64/photo.jpg", "userId": "13278186009456793751"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
literal_eval(str_dict)[0]['value'][0]
# + id="8Lb8SOFqAmoU" colab_type="code" outputId="4a90b293-b96d-4a43-d248-70d107c58cbb" executionInfo={"status": "ok", "timestamp": 1581719236704, "user_tz": -60, "elapsed": 2693, "user": {"displayName": "<NAME>", "photoUrl": "https://lh6.googleusercontent.com/-Mk6sDG3z8U8/AAAAAAAAAAI/AAAAAAAATPI/zF3jRAXRQtg/s64/photo.jpg", "userId": "13278186009456793751"}} colab={"base_uri": "https://localhost:8080/", "height": 138}
def parse_features(x):
if str(x) == 'nan': return[]
return literal_eval(x.replace('\\"', '"'))
df['features_parsed'] = df['features'].map(parse_features)
df['features_parsed'].head().values
# + id="jwKmhCnYBjTn" colab_type="code" colab={}
def parse_features(x):
output_dict = {}
if str(x) == 'nan': return output_dict
features = literal_eval(x.replace('\\"', '"'))
for item in features:
# {'key': 'Gender', 'value': ['Men']}
key = item['key'].lower().strip()
value = item['value'][0].lower().strip()
output_dict[key] = value
return output_dict
df['features_parsed'] = df['features'].map(parse_features)
# + id="CrmmAh15dKRc" colab_type="code" outputId="bacfd615-7ebf-4880-ddfc-490cfc8f8e03" executionInfo={"status": "ok", "timestamp": 1581719245943, "user_tz": -60, "elapsed": 1426, "user": {"displayName": "<NAME>", "photoUrl": "https://lh6.googleusercontent.com/-Mk6sDG3z8U8/AAAAAAAAAAI/AAAAAAAATPI/zF3jRAXRQtg/s64/photo.jpg", "userId": "13278186009456793751"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
keys = set()
df['features_parsed'].map(lambda x: keys.update(x.keys()))
len(keys)
# + id="kCyj3NPVft_F" colab_type="code" outputId="154996a0-cfe9-454b-c121-8381eb2ffdff" executionInfo={"status": "ok", "timestamp": 1581719248869, "user_tz": -60, "elapsed": 1459, "user": {"displayName": "<NAME>", "photoUrl": "https://lh6.googleusercontent.com/-Mk6sDG3z8U8/AAAAAAAAAAI/AAAAAAAATPI/zF3jRAXRQtg/s64/photo.jpg", "userId": "13278186009456793751"}} colab={"base_uri": "https://localhost:8080/", "height": 118}
df.features_parsed.head().values
# + id="yanDejpjdpR7" colab_type="code" outputId="f3b18e69-2e29-4f3b-b7e1-9b93e88a68cf" executionInfo={"status": "ok", "timestamp": 1581719254743, "user_tz": -60, "elapsed": 3962, "user": {"displayName": "<NAME>", "photoUrl": "https://lh6.googleusercontent.com/-Mk6sDG3z8U8/AAAAAAAAAAI/AAAAAAAATPI/zF3jRAXRQtg/s64/photo.jpg", "userId": "13278186009456793751"}} colab={"base_uri": "https://localhost:8080/", "height": 66, "referenced_widgets": ["58758183e871429e9216197db506e293", "0b1583a9978d463b9efe752ed4edfa91", "ad1a4ca2e4804d699d5f5102975f5e54", "85e7d9e7346548ea8978e5c93d286bb3", "635e9ad7ca874c5a870fc1d537471d8f", "70fae058f69148b79de5b82656e991ba", "8ebc1703737a4478a69608f1a181323e", "915cd2ba8c3942bd9621ff734cfa0a43"]}
def get_name_feat(key):
return 'feat_' + key
for key in tqdm_notebook(keys):
df[get_name_feat(key)] = df.features_parsed.map(lambda feats: feats[key] if key in feats else np.nan)
# + id="7lRXURk0g-sv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 134} outputId="e289d00c-031b-4d46-b0a6-1a48f4ba7775" executionInfo={"status": "ok", "timestamp": 1581719271718, "user_tz": -60, "elapsed": 917, "user": {"displayName": "<NAME>", "photoUrl": "https://lh6.googleusercontent.com/-Mk6sDG3z8U8/AAAAAAAAAAI/AAAAAAAATPI/zF3jRAXRQtg/s64/photo.jpg", "userId": "13278186009456793751"}}
df.columns
# + id="yZxtF_l60LCe" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="42ae7e78-f91a-4c11-cdfa-ee10f7e5cc4a" executionInfo={"status": "ok", "timestamp": 1581720003883, "user_tz": -60, "elapsed": 1497, "user": {"displayName": "<NAME>", "photoUrl": "https://lh6.googleusercontent.com/-Mk6sDG3z8U8/AAAAAAAAAAI/AAAAAAAATPI/zF3jRAXRQtg/s64/photo.jpg", "userId": "13278186009456793751"}}
df[df['feat_athlete'].isnull()].shape
# + id="v-z76q0z0Xvb" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="8ef45c87-ffcc-48fe-e769-d2e2190e71ca" executionInfo={"status": "ok", "timestamp": 1581719372285, "user_tz": -60, "elapsed": 918, "user": {"displayName": "<NAME>", "photoUrl": "https://lh6.googleusercontent.com/-Mk6sDG3z8U8/AAAAAAAAAAI/AAAAAAAATPI/zF3jRAXRQtg/s64/photo.jpg", "userId": "13278186009456793751"}}
df.shape[0]
# + id="m69b-4E60fl1" colab_type="code" colab={}
keys_stat = {}
for key in keys:
keys_stat[key] = df[False == df[get_name_feat(key)].isnull()].shape[0] / df.shape[0] * 100
# + id="CSjeNPt50urG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 101} outputId="31ed9203-54fd-436a-cfa2-96a5d97c324e" executionInfo={"status": "ok", "timestamp": 1581720230146, "user_tz": -60, "elapsed": 1427, "user": {"displayName": "<NAME>", "photoUrl": "https://lh6.googleusercontent.com/-Mk6sDG3z8U8/AAAAAAAAAAI/AAAAAAAATPI/zF3jRAXRQtg/s64/photo.jpg", "userId": "13278186009456793751"}}
{k:v for k,v in keys_stat.items() if v>30}
# + id="KquC1r8r3Wp9" colab_type="code" colab={}
df['feat_brand_cat'] = df['feat_brand'].factorize()[0]
df['feat_color_cat'] = df['feat_color'].factorize()[0]
df['feat_gender_cat'] = df['feat_gender'].factorize()[0]
df['feat_manufacturer_cat'] = df['feat_manufacturer'].factorize()[0]
df['feat_material_cat'] = df['feat_material'].factorize()[0]
df['feat_sport_cat'] = df['feat_sport'].factorize()[0]
df['feat_style_cat'] = df['feat_style'].factorize()[0]
for key in keys:
df[ get_name_feat(key) + '_cat'] = df[get_name_feat(key)].factorize()[0]
# + id="7yfPDCGd4Y8C" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="c64167f3-b995-4a64-ee52-c0414c7553f9" executionInfo={"status": "ok", "timestamp": 1581721641095, "user_tz": -60, "elapsed": 1528, "user": {"displayName": "<NAME>", "photoUrl": "https://lh6.googleusercontent.com/-Mk6sDG3z8U8/AAAAAAAAAAI/AAAAAAAATPI/zF3jRAXRQtg/s64/photo.jpg", "userId": "13278186009456793751"}}
df['brand'] = df['brand'].map(lambda x: str(x).lower() )
df[df.brand==df.feat_brand].shape
# + id="amgyy47g4f83" colab_type="code" colab={}
feats = ['']
# + id="JqQO1JAB5lSo" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="e76f7110-0dba-4288-bffd-79d784a74fbc" executionInfo={"status": "ok", "timestamp": 1581720735440, "user_tz": -60, "elapsed": 674, "user": {"displayName": "<NAME>", "photoUrl": "https://lh6.googleusercontent.com/-Mk6sDG3z8U8/AAAAAAAAAAI/AAAAAAAATPI/zF3jRAXRQtg/s64/photo.jpg", "userId": "13278186009456793751"}}
model = RandomForestRegressor(max_depth=5, n_estimators=100)
run_model(['brand_cat'],model)
# + id="8GooxbSz9Q2O" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="43f05fe2-9d53-4de0-a3fe-c8bb478c1392" executionInfo={"status": "ok", "timestamp": 1581721695222, "user_tz": -60, "elapsed": 1268, "user": {"displayName": "<NAME>", "photoUrl": "https://lh6.googleusercontent.com/-Mk6sDG3z8U8/AAAAAAAAAAI/AAAAAAAATPI/zF3jRAXRQtg/s64/photo.jpg", "userId": "13278186009456793751"}}
feats_cat = [x for x in df.columns if 'cat' in x]
feats_cat
# + id="O3g8ZPrH5wk1" colab_type="code" colab={}
feats = ['brand_cat', 'feat_metal type_cat', 'feat_shape_cat', 'feat_gender_cat', 'feat_material_cat', 'feat_brand_cat', 'feat_style_cat', 'feat_sport_cat']
# feats += feats_cat
# feats = list(set(feats))
model = RandomForestRegressor(max_depth=5, n_estimators=100)
result = run_model(feats,model)
# + id="2ILpM5_t6HDv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 185} outputId="8e3a5015-14d2-45b8-858a-c182646fa568" executionInfo={"status": "ok", "timestamp": 1581722238869, "user_tz": -60, "elapsed": 5340, "user": {"displayName": "<NAME>", "photoUrl": "https://lh6.googleusercontent.com/-Mk6sDG3z8U8/AAAAAAAAAAI/AAAAAAAATPI/zF3jRAXRQtg/s64/photo.jpg", "userId": "13278186009456793751"}}
X = df[feats].values
y = df['prices_amountmin'].values
m = RandomForestRegressor(max_depth=5, n_estimators=100, random_state=0)
m.fit(X, y)
print(result)
perm = PermutationImportance(m, random_state=1).fit(X, y);
eli5.show_weights(perm, feature_names=feats)
# + id="0-zKkv896x-g" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 218} outputId="529758d4-0e70-425f-9630-cddc0e99b365" executionInfo={"status": "ok", "timestamp": 1581721149750, "user_tz": -60, "elapsed": 1486, "user": {"displayName": "<NAME>", "photoUrl": "https://lh6.googleusercontent.com/-Mk6sDG3z8U8/AAAAAAAAAAI/AAAAAAAATPI/zF3jRAXRQtg/s64/photo.jpg", "userId": "13278186009456793751"}}
df['brand'].value_counts(normalize=True)
# + id="JhRGtBSn7Q5k" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 121} outputId="8bfc23a0-3072-4ce4-cfb3-5942a8c58a3c" executionInfo={"status": "ok", "timestamp": 1581721424657, "user_tz": -60, "elapsed": 1506, "user": {"displayName": "<NAME>", "photoUrl": "https://lh6.googleusercontent.com/-Mk6sDG3z8U8/AAAAAAAAAAI/AAAAAAAATPI/zF3jRAXRQtg/s64/photo.jpg", "userId": "13278186009456793751"}}
df[df['brand']=='nike'].features_parsed.sample(5).values
# + id="5X9mDJnc7b05" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 50} outputId="18d4acbd-edde-44e7-80d2-4b728c2122e0" executionInfo={"status": "ok", "timestamp": 1581722427504, "user_tz": -60, "elapsed": 4302, "user": {"displayName": "<NAME>", "photoUrl": "https://lh6.googleusercontent.com/-Mk6sDG3z8U8/AAAAAAAAAAI/AAAAAAAATPI/zF3jRAXRQtg/s64/photo.jpg", "userId": "13278186009456793751"}}
# ls
# + id="QSpTxMKxAM8x" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="cb977d1d-21f3-4f62-ac0c-3fca6d7e4453" executionInfo={"status": "ok", "timestamp": 1581722451246, "user_tz": -60, "elapsed": 4052, "user": {"displayName": "<NAME>", "photoUrl": "https://lh6.googleusercontent.com/-Mk6sDG3z8U8/AAAAAAAAAAI/AAAAAAAATPI/zF3jRAXRQtg/s64/photo.jpg", "userId": "13278186009456793751"}}
# ls 'matrix_one (2)'
# + id="PtgFzCmHAS0D" colab_type="code" colab={}
# !git add
|
matrix_one (2)/day5.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:filter_pipeline]
# language: python
# name: conda-env-filter_pipeline-py
# ---
# This notebook creates the input for the mutation rate models of each observable signature 5 contribution per patient of each subset (private primary, private relapse and shared) of clonal SNVs.
#
# With this data the phylogenetic trees are also drawn which corresponds to Figure 3b of the paper
#
# This piece of code relies on a workspace directory structure such as:
# ```
# cohort/
# patientID/
# DxTumorID_vs_normalID/
# ReTumorID_vs_normalID/ (sometimes)
#
# ```
# patientID, DxTumorID etc can be found in ../ext_files/all_cohort_clinical_groups.tsv
#
# Be aware that the filtered mafs with clonal classification and joined mutations after running the scripts in ```filter/``` have the following file name: ```TumorID_vs_normalID + _strelka_uniq_all_anno_vep92_categories_filt_snps_cluster.maf```
# .This file name is used in the following code.
# +
import sys, os
os.environ["PATH"] = os.path.dirname(sys.executable) + os.pathsep + os.environ["PATH"]
import matplotlib
matplotlib.use('Agg')
import pandas as pd
import os
import matplotlib.pyplot as plt
import matplotlib.gridspec as gridspec
from matplotlib import collections as mc
from aux_functions import stage_mapping, get_context_rev,add_pyrimidine_type, df_to_dict,count_variant_type, get_muts_x_signature
from aux_data_in_pyvar import PATS_DIRS, config_rcparams
pd.set_option('display.max_columns', None)
pd.set_option('display.max_rows', None)
pd.set_option('display.max_colwidth', -1)
# %load_ext autoreload
# %autoreload 2
# -
config_rcparams()
plt.rcParams['font.size'] = 14
def figure_phylo_tree(df,sig, out_dir, clonality):
# PLOT
fig = plt.figure(figsize=(20, 80))
outer = gridspec.GridSpec(len(df['PATIENT'].unique()), 1, wspace=0, hspace=0)
j = 0
num_pat = len(df['PATIENT'].unique())
for i, rw in df.iterrows():
# CREATE SEGMENTS
# lines as proportion in percentatge
if round(df.loc[i, 'PRIVATE_PRY_'+sig]) > round(
df.loc[i, 'PRIVATE_REL_'+sig]):
total_scale = df.loc[i, 'TRUNK_'+sig] + df.loc[i, 'PRIVATE_PRY_'+sig]
else:
total_scale = df.loc[i, 'TRUNK_'+sig] + df.loc[i, 'PRIVATE_REL_'+sig]
line_trunk = (df.loc[i, 'TRUNK_'+sig] / total_scale) * 100
line_pry = (df.loc[i, 'PRIVATE_PRY_'+sig] / total_scale) * 100
line_rel = (df.loc[i, 'PRIVATE_REL_'+sig] / total_scale) * 100
lines = [[(0, 1), (line_trunk, 1)], [(line_trunk, 1), (line_trunk + line_rel, 2.4)],
[(line_trunk, 1), (line_trunk + line_pry, 0.2)]]
c = ['#006837', '#fd8d3c','#2c7fb8']
# PLOT
line_segments = mc.LineCollection(lines, linewidths=20,
colors=c, linestyle='solid')
ax = plt.subplot(outer[j, 0])
ax.add_collection(line_segments)
ax.scatter(line_trunk, 1, s=1000, color='r', zorder=3)
ax.set_ylabel('{} \n {} years,\n{}, {}'.format(df.loc[i, 'PATIENT'],
int(df.loc[i, 'DIAGNOSIS_AGE_YEARS']),
df.loc[i, 'SEX'],
df.loc[i, 'IMMUNO_CLASS']), fontsize=60, labelpad=50)
ax.set_ylim([-3, 5])
ax.set_xlim([0, 100])
ax.spines["right"].set_visible(False)
ax.annotate('{}'.format(int(round(df.loc[i, 'TRUNK_'+sig]))),
(int(round(line_trunk/3)), -1), textcoords='data',
size=40, horizontalalignment='center', rotation=90)
ax.annotate('{}'.format(int(round(df.loc[i, 'PRIVATE_PRY_'+sig]))),
(int(round((line_trunk + line_pry) - (line_pry / 2))), -1.2), textcoords='data',
size=40, verticalalignment='center', rotation=90, ha='right')
ax.annotate('{}'.format(int(round(df.loc[i, 'PRIVATE_REL_'+sig]))),
(int(round((line_trunk + line_pry) - (line_pry / 2))), 3), textcoords='data', size=40,
verticalalignment='center', rotation=90, ha='left')
ax.annotate('{} days'.format(int(df.loc[i, 'PRIMARY_TO_RELAPSE_AGE_DAYS'])),
(100, 1), textcoords='data', size=40, verticalalignment='center', rotation=90)
if j < (num_pat-1):
fig.add_subplot(ax)
plt.yticks([], [])
plt.xticks([], [])
else:
fig.add_subplot(ax)
plt.tick_params(axis='x', which='major', labelsize=50)
plt.yticks([], [])
j = j + 1
plt.tight_layout()
plt.xlabel("Molecular time (%)", rotation=180, fontsize=80)
plt.xticks(rotation=90)
fig.savefig(os.path.join(out_dir,
"phylo_tree_muts_"+clonality+"_"+sig+".svg"),
dpi=500, bbox_inches='tight',
orientation='portrait')
plt.show()
plt.close()
clinical = pd.read_csv("", sep='\t') # Additional file 1 Table S1
clinical = clinical[~clinical['Patient_id'].isin(['PAT3', 'PAT4'])]
clinical.head()
# ### Signatures contribution to each subset
# Path to the results from fitting of signatures of deconstructSigs run. Here we expect the results from
# all samples (primary and relapse) together. If you check ../ext_runs/run_deconstructSig/make_inputs_fitting_adults.ipynb
# it should correspond to the results file signatures_weight.csv of a run with folder named run_subsets_together/
deconstruct_run = ""
clonal = True
# +
signature_subsets = pd.DataFrame()
for pat in clinical['Patient_id'].unique():
# GET INFO
pat_clinical = clinical[clinical['Patient_id'] == pat].reset_index()
com_pry = pat_clinical.loc[0, 'Primary_seq_id']+'_vs_'+pat_clinical.loc[0, 'Remission_seq_id']
com_rel = pat_clinical.loc[0, 'Relapse_seq_id']+'_vs_'+pat_clinical.loc[0, 'Remission_seq_id']
print(pat)
# READ MUTATION DATASET
df_pry = pd.read_table(os.path.join(PATS_DIRS[pat], pat, com_pry,
com_pry+'_strelka_uniq_all_anno_vep92_categories_filt_snps_cluster.maf'),
sep='\t',low_memory=False)
df_rel = pd.read_table(os.path.join(PATS_DIRS[pat], pat, com_rel,
com_rel+'_strelka_uniq_all_anno_vep92_categories_filt_snps_cluster.maf'),
sep='\t',low_memory=False)
# GET CLONALS and SNVS
df_pry = df_pry[df_pry['mut_type'] == 'snv']
df_rel = df_rel[df_rel['mut_type'] == 'snv']
print(len(df_pry))
print(len(df_rel))
# SEPARATE CLONALS SUBCLONALS
if clonal == True:
df_pry = df_pry[df_pry['clonal_classification'] == 'clonal']
df_rel = df_rel[df_rel['clonal_classification'] == 'clonal']
# GET SUBSETS
all_pry_variants = set(df_pry['Variant'].unique())
all_rel_variants = set(df_rel['Variant'].unique())
print(len(all_pry_variants))
print(len(all_rel_variants))
shared_variants = all_pry_variants.intersection(all_rel_variants)
private_pry_variants = all_pry_variants.difference(shared_variants)
private_rel_variants = all_rel_variants.difference(shared_variants)
df_shared = df_pry[df_pry['Variant'].isin(shared_variants)]
df_private_pry = df_pry[df_pry['Variant'].isin(private_pry_variants)]
df_private_rel = df_rel[df_rel['Variant'].isin(private_rel_variants)]
for signature in ['SBS1', 'SBS5', 'SBS18']:
count_pp, count_pr, count_sh = get_muts_x_signature(sh=df_shared, pp=df_private_pry,
pr=df_private_rel, pat=pat, sig=signature,
prob_file_path=deconstruct_run)
signature_subsets.set_value(index=pat, col='PRIVATE_PRY_'+signature, value=count_pp)
signature_subsets.set_value(index=pat, col='PRIVATE_REL_'+signature, value=count_pr)
signature_subsets.set_value(index=pat, col='TRUNK_'+signature, value=count_sh)
# ADD info for phylotree
signature_subsets.set_value(index=pat, col='DIAGNOSIS_AGE_YEARS', value=pat_clinical.loc[0, 'Primary_diagnosis_age'])
signature_subsets.set_value(index=pat, col='PRIMARY_TO_RELAPSE_AGE_DAYS', value=pat_clinical.loc[0, 'days_between_pry_rel'])
signature_subsets.set_value(index=pat, col='IMMUNO_CLASS', value=pat_clinical.loc[0,'Primary_immunoclassification'])
signature_subsets.set_value(index=pat, col='SEX', value=pat_clinical.loc[0,'Sex'])
# -
signature_subsets.reset_index(inplace=True)
signature_subsets.rename(columns={'index':'PATIENT'}, inplace=True)
signature_subsets.sort_values('DIAGNOSIS_AGE_YEARS', ascending=True, inplace=True) # sort by age
signature_subsets.to_csv("../intermediate_files/signature_counts.tsv", sep='\t', index=False)
# ### phylotree plot
signature = 'SBS5'
output_path = "" # output path for the figure
figure_phylo_tree(signature_subsets, signature, output_path, 'clonal')
len(signature_subsets)
signature_subsets
|
notebook_figures/counts_with_probability_exposure_assignments_phylotree.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Brewing Logistic Regression then Going Deeper
#
# While Caffe is made for deep networks it can likewise represent "shallow" models like logistic regression for classification. We'll do simple logistic regression on synthetic data that we'll generate and save to HDF5 to feed vectors to Caffe. Once that model is done, we'll add layers to improve accuracy. That's what Caffe is about: define a model, experiment, and then deploy.
# +
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
import os
os.chdir('..')
import sys
sys.path.insert(0, './python')
import caffe
import os
import h5py
import shutil
import tempfile
import sklearn
import sklearn.datasets
import sklearn.linear_model
import pandas as pd
# -
# Synthesize a dataset of 10,000 4-vectors for binary classification with 2 informative features and 2 noise features.
# +
X, y = sklearn.datasets.make_classification(
n_samples=10000, n_features=4, n_redundant=0, n_informative=2,
n_clusters_per_class=2, hypercube=False, random_state=0
)
# Split into train and test
X, Xt, y, yt = sklearn.cross_validation.train_test_split(X, y)
# Visualize sample of the data
ind = np.random.permutation(X.shape[0])[:1000]
df = pd.DataFrame(X[ind])
_ = pd.scatter_matrix(df, figsize=(9, 9), diagonal='kde', marker='o', s=40, alpha=.4, c=y[ind])
# -
# Learn and evaluate scikit-learn's logistic regression with stochastic gradient descent (SGD) training. Time and check the classifier's accuracy.
# +
# %%timeit
# Train and test the scikit-learn SGD logistic regression.
clf = sklearn.linear_model.SGDClassifier(
loss='log', n_iter=1000, penalty='l2', alpha=5e-4, class_weight='auto')
clf.fit(X, y)
yt_pred = clf.predict(Xt)
print('Accuracy: {:.3f}'.format(sklearn.metrics.accuracy_score(yt, yt_pred)))
# -
# Save the dataset to HDF5 for loading in Caffe.
# +
# Write out the data to HDF5 files in a temp directory.
# This file is assumed to be caffe_root/examples/hdf5_classification.ipynb
dirname = os.path.abspath('./examples/hdf5_classification/data')
if not os.path.exists(dirname):
os.makedirs(dirname)
train_filename = os.path.join(dirname, 'train.h5')
test_filename = os.path.join(dirname, 'test.h5')
# HDF5DataLayer source should be a file containing a list of HDF5 filenames.
# To show this off, we'll list the same data file twice.
with h5py.File(train_filename, 'w') as f:
f['data'] = X
f['label'] = y.astype(np.float32)
with open(os.path.join(dirname, 'train.txt'), 'w') as f:
f.write(train_filename + '\n')
f.write(train_filename + '\n')
# HDF5 is pretty efficient, but can be further compressed.
comp_kwargs = {'compression': 'gzip', 'compression_opts': 1}
with h5py.File(test_filename, 'w') as f:
f.create_dataset('data', data=Xt, **comp_kwargs)
f.create_dataset('label', data=yt.astype(np.float32), **comp_kwargs)
with open(os.path.join(dirname, 'test.txt'), 'w') as f:
f.write(test_filename + '\n')
# -
# Let's define logistic regression in Caffe through Python net specification. This is a quick and natural way to define nets that sidesteps manually editing the protobuf model.
# +
from caffe import layers as L
from caffe import params as P
def logreg(hdf5, batch_size):
# logistic regression: data, matrix multiplication, and 2-class softmax loss
n = caffe.NetSpec()
n.data, n.label = L.HDF5Data(batch_size=batch_size, source=hdf5, ntop=2)
n.ip1 = L.InnerProduct(n.data, num_output=2, weight_filler=dict(type='xavier'))
n.accuracy = L.Accuracy(n.ip1, n.label)
n.loss = L.SoftmaxWithLoss(n.ip1, n.label)
return n.to_proto()
train_net_path = 'examples/hdf5_classification/logreg_auto_train.prototxt'
with open(train_net_path, 'w') as f:
f.write(str(logreg('examples/hdf5_classification/data/train.txt', 10)))
test_net_path = 'examples/hdf5_classification/logreg_auto_test.prototxt'
with open(test_net_path, 'w') as f:
f.write(str(logreg('examples/hdf5_classification/data/test.txt', 10)))
# -
# Now, we'll define our "solver" which trains the network by specifying the locations of the train and test nets we defined above, as well as setting values for various parameters used for learning, display, and "snapshotting".
# +
from caffe.proto import caffe_pb2
def solver(train_net_path, test_net_path):
s = caffe_pb2.SolverParameter()
# Specify locations of the train and test networks.
s.train_net = train_net_path
s.test_net.append(test_net_path)
s.test_interval = 1000 # Test after every 1000 training iterations.
s.test_iter.append(250) # Test 250 "batches" each time we test.
s.max_iter = 10000 # # of times to update the net (training iterations)
# Set the initial learning rate for stochastic gradient descent (SGD).
s.base_lr = 0.01
# Set `lr_policy` to define how the learning rate changes during training.
# Here, we 'step' the learning rate by multiplying it by a factor `gamma`
# every `stepsize` iterations.
s.lr_policy = 'step'
s.gamma = 0.1
s.stepsize = 5000
# Set other optimization parameters. Setting a non-zero `momentum` takes a
# weighted average of the current gradient and previous gradients to make
# learning more stable. L2 weight decay regularizes learning, to help prevent
# the model from overfitting.
s.momentum = 0.9
s.weight_decay = 5e-4
# Display the current training loss and accuracy every 1000 iterations.
s.display = 1000
# Snapshots are files used to store networks we've trained. Here, we'll
# snapshot every 10K iterations -- just once at the end of training.
# For larger networks that take longer to train, you may want to set
# snapshot < max_iter to save the network and training state to disk during
# optimization, preventing disaster in case of machine crashes, etc.
s.snapshot = 10000
s.snapshot_prefix = 'examples/hdf5_classification/data/train'
# We'll train on the CPU for fair benchmarking against scikit-learn.
# Changing to GPU should result in much faster training!
s.solver_mode = caffe_pb2.SolverParameter.CPU
return s
solver_path = 'examples/hdf5_classification/logreg_solver.prototxt'
with open(solver_path, 'w') as f:
f.write(str(solver(train_net_path, test_net_path)))
# -
# Time to learn and evaluate our Caffeinated logistic regression in Python.
# +
# %%timeit
caffe.set_mode_cpu()
solver = caffe.get_solver(solver_path)
solver.solve()
accuracy = 0
batch_size = solver.test_nets[0].blobs['data'].num
test_iters = int(len(Xt) / batch_size)
for i in range(test_iters):
solver.test_nets[0].forward()
accuracy += solver.test_nets[0].blobs['accuracy'].data
accuracy /= test_iters
print("Accuracy: {:.3f}".format(accuracy))
# -
# Do the same through the command line interface for detailed output on the model and solving.
# !./build/tools/caffe train -solver examples/hdf5_classification/logreg_solver.prototxt
# If you look at output or the `logreg_auto_train.prototxt`, you'll see that the model is simple logistic regression.
# We can make it a little more advanced by introducing a non-linearity between weights that take the input and weights that give the output -- now we have a two-layer network.
# That network is given in `nonlinear_auto_train.prototxt`, and that's the only change made in `nonlinear_logreg_solver.prototxt` which we will now use.
#
# The final accuracy of the new network should be higher than logistic regression!
# +
from caffe import layers as L
from caffe import params as P
def nonlinear_net(hdf5, batch_size):
# one small nonlinearity, one leap for model kind
n = caffe.NetSpec()
n.data, n.label = L.HDF5Data(batch_size=batch_size, source=hdf5, ntop=2)
# define a hidden layer of dimension 40
n.ip1 = L.InnerProduct(n.data, num_output=40, weight_filler=dict(type='xavier'))
# transform the output through the ReLU (rectified linear) non-linearity
n.relu1 = L.ReLU(n.ip1, in_place=True)
# score the (now non-linear) features
n.ip2 = L.InnerProduct(n.ip1, num_output=2, weight_filler=dict(type='xavier'))
# same accuracy and loss as before
n.accuracy = L.Accuracy(n.ip2, n.label)
n.loss = L.SoftmaxWithLoss(n.ip2, n.label)
return n.to_proto()
train_net_path = 'examples/hdf5_classification/nonlinear_auto_train.prototxt'
with open(train_net_path, 'w') as f:
f.write(str(nonlinear_net('examples/hdf5_classification/data/train.txt', 10)))
test_net_path = 'examples/hdf5_classification/nonlinear_auto_test.prototxt'
with open(test_net_path, 'w') as f:
f.write(str(nonlinear_net('examples/hdf5_classification/data/test.txt', 10)))
solver_path = 'examples/hdf5_classification/nonlinear_logreg_solver.prototxt'
with open(solver_path, 'w') as f:
f.write(str(solver(train_net_path, test_net_path)))
# +
# %%timeit
caffe.set_mode_cpu()
solver = caffe.get_solver(solver_path)
solver.solve()
accuracy = 0
batch_size = solver.test_nets[0].blobs['data'].num
test_iters = int(len(Xt) / batch_size)
for i in range(test_iters):
solver.test_nets[0].forward()
accuracy += solver.test_nets[0].blobs['accuracy'].data
accuracy /= test_iters
print("Accuracy: {:.3f}".format(accuracy))
# -
# Do the same through the command line interface for detailed output on the model and solving.
# !./build/tools/caffe train -solver examples/hdf5_classification/nonlinear_logreg_solver.prototxt
# Clean up (comment this out if you want to examine the hdf5_classification/data directory).
shutil.rmtree(dirname)
|
examples/brewing-logreg.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
#
# # Importing data from EEG devices
#
#
# MNE includes various functions and utilities for reading EEG
# data and electrode locations.
# :depth: 2
#
#
#
# BrainVision (.vhdr, .vmrk, .eeg)
# ================================
#
# The BrainVision file format consists of three separate files:
#
# 1. A text header file (``.vhdr``) containing meta data
# 2. A text marker file (``.vmrk``) containing information about events in the
# data
# 3. A binary data file (``.eeg``) containing the voltage values of the EEG
#
# Both text files are based on the
# `Microsoft Windows INI format <https://en.wikipedia.org/wiki/INI_file>`_
# consisting of:
#
# * sections marked as ``[square brackets]``
# * comments marked as ``; comment``
# * key-value pairs marked as ``key=value``
#
# A documentation for core BrainVision file format is provided by Brain Products.
# You can view the specification
# `here <https://docs.google.com/viewer?url=https://raw.githubusercontent.com/sappelhoff/brainvision-validator/master/doc/BrainVisionCoreFileFormat.pdf>`_
#
# BrainVision EEG files can be read in using :func:`mne.io.read_raw_brainvision`
# with the .vhdr header file as an input.
#
# <div class="alert alert-danger"><h4>Warning</h4><p>Renaming BrainVision files can be problematic due to their
# multifile structure. See this
# `example <https://mne-tools.github.io/mne-bids/auto_examples/rename_brainvision_files>`_
# for an instruction.</p></div>
#
#
#
# European data format (.edf)
# ===========================
#
# EDF and EDF+ files can be read using :func:`mne.io.read_raw_edf`.
#
# `EDF (European Data Format) <http://www.edfplus.info/specs/edf.html>`_ and
# `EDF+ <http://www.edfplus.info/specs/edfplus.html>`_ are 16-bit formats.
#
# The EDF+ files may contain an annotation channel which can be used to store
# trigger information. These annotations are available in ``raw.annotations``.
#
# Saving EDF files is not supported natively yet. This
# `gist <https://gist.github.com/skjerns/bc660ef59dca0dbd53f00ed38c42f6be>`__
# can be used to save any mne.io.Raw into EDF/EDF+/BDF/BDF+.
#
#
#
# BioSemi data format (.bdf)
# ==========================
#
# The `BDF format <http://www.biosemi.com/faq/file_format.htm>`_ is a 24-bit
# variant of the EDF format used by EEG systems manufactured by BioSemi. It can
# be imported with :func:`mne.io.read_raw_bdf`.
#
# BioSemi amplifiers do not perform "common mode noise rejection" automatically.
# The signals in the EEG file are the voltages between each electrode and CMS
# active electrode, which still contain some CM noise (50 Hz, ADC reference
# noise, etc., see `the BioSemi FAQ <https://www.biosemi.com/faq/cms&drl.htm>`__
# for further detail).
# Thus, it is advisable to choose a reference (e.g., a single channel like Cz,
# average of linked mastoids, average of all electrodes, etc.) on import of
# BioSemi data to avoid losing signal information. The data can be re-referenced
# later after cleaning if desired.
#
# <div class="alert alert-danger"><h4>Warning</h4><p>The data samples in a BDF file are represented in a 3-byte
# (24-bit) format. Since 3-byte raw data buffers are not presently
# supported in the fif format these data will be changed to 4-byte
# integers in the conversion.</p></div>
#
#
#
# General data format (.gdf)
# ==========================
#
# GDF files can be read in using :func:`mne.io.read_raw_gdf`.
#
# `GDF (General Data Format) <https://arxiv.org/abs/cs/0608052>`_ is a flexible
# format for biomedical signals that overcomes some of the limitations of the
# EDF format. The original specification (GDF v1) includes a binary header
# and uses an event table. An updated specification (GDF v2) was released in
# 2011 and adds fields for additional subject-specific information (gender,
# age, etc.) and allows storing several physical units and other properties.
# Both specifications are supported in MNE.
#
#
#
# Neuroscan CNT data format (.cnt)
# ================================
#
# CNT files can be read in using :func:`mne.io.read_raw_cnt`.
# The channel locations can be read from a montage or the file header. If read
# from the header, the data channels (channels that are not assigned to EOG, ECG,
# EMG or misc) are fit to a sphere and assigned a z-value accordingly. If a
# non-data channel does not fit to the sphere, it is assigned a z-value of 0.
#
# <div class="alert alert-danger"><h4>Warning</h4><p>Reading channel locations from the file header may be dangerous, as the
# x_coord and y_coord in ELECTLOC section of the header do not necessarily
# translate to absolute locations. Furthermore, EEG-electrode locations that
# do not fit to a sphere will distort the layout when computing the z-values.
# If you are not sure about the channel locations in the header, use of a
# montage is encouraged.</p></div>
#
#
#
# EGI simple binary (.egi)
# ========================
#
# EGI simple binary files can be read in using :func:`mne.io.read_raw_egi`.
# The EGI raw files are simple binary files with a header and can be exported
# from using the EGI Netstation acquisition software.
#
#
#
# EGI MFF (.mff)
# ==============
# These files can also be read with :func:`mne.io.read_raw_egi`.
#
#
#
# EEGLAB set files (.set)
# =======================
#
# EEGLAB .set files can be read in using :func:`mne.io.read_raw_eeglab`
# and :func:`mne.read_epochs_eeglab`.
#
#
#
# Nicolet (.data)
# ===============
# These files can be read with :func:`mne.io.read_raw_nicolet`.
#
#
#
# eXimia EEG data (.nxe)
# ======================
#
# EEG data from the Nexstim eXimia system can be read in using the
# :func:`mne.io.read_raw_eximia` function.
#
# Setting EEG references
# ======================
#
# The preferred method for applying an EEG reference in MNE is
# :func:`mne.set_eeg_reference`, or equivalent instance methods like
# :meth:`raw.set_eeg_reference() <mne.io.Raw.set_eeg_reference>`. By default,
# an average reference is used. See `tut-set-eeg-ref` for more information.
#
# Reading electrode locations and head shapes for EEG recordings
# ==============================================================
#
# Some EEG formats (EGI, EDF/EDF+, BDF) neither contain electrode location
# information nor head shape digitization information. Therefore, this
# information has to be provided separately. For that purpose all raw instances
# have a :meth:`mne.io.Raw.set_montage` method to set electrode locations.
#
# When using the locations of the fiducial points the digitization data
# are converted to the MEG head coordinate system employed in the
# MNE software, see `coordinate_systems`.
#
|
dev/_downloads/8de2adebc31e5c03b3dea5dae8841c18/plot_20_reading_eeg_data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Plot fig. 7.7
#
# Temperature response to forcing 1750-2019
#
# Theme Song: Arguing With Thermometers<br>
# Artist: <NAME><br>
# Album: A Flash flood of Colour<br>
# Released: 2012
# +
import numpy as np
import scipy.stats as st
import pandas as pd
import matplotlib.pyplot as pl
import os
from matplotlib import gridspec, rc
from matplotlib.lines import Line2D
import matplotlib.patches as mp
from netCDF4 import Dataset
import warnings
from ar6.utils.h5 import *
# +
# TODO - sort out figure sizing
pl.rcParams['figure.figsize'] = (18/2.54, 11/2.54)
pl.rcParams['font.size'] = 11
pl.rcParams['font.family'] = 'Arial'
pl.rcParams['xtick.direction'] = 'out'
pl.rcParams['xtick.minor.visible'] = True
pl.rcParams['ytick.major.left'] = True
pl.rcParams['ytick.major.size'] = 0
pl.rcParams['xtick.top'] = True
# -
results = load_dict_from_hdf5('../data_output_large/twolayer_AR6-historical.h5')
results.keys()
results['AR6-historical']['surface_temperature'].shape
results['AR6-historical']['surface_temperature'][0].mean()
forcings = ['co2', 'ch4', 'n2o', 'other_wmghg', 'o3', 'h2o_stratospheric',
'contrails', 'aerosol-radiation_interactions', 'aerosol-cloud_interactions',
'bc_on_snow', 'land_use', 'volcanic', 'solar', 'wmghgs', 'aerosol', 'albedo', 'anthro', 'natural']
for forcing in forcings:
print(forcing, results['remove_%s' % forcing]['surface_temperature'][0].mean())
# +
AR6_forc = {}
AR6_ecsforc = {}
forcings = ['co2', 'ch4', 'n2o', 'other_wmghg', 'o3', 'h2o_stratospheric',
'contrails', 'aerosol-radiation_interactions', 'aerosol-cloud_interactions',
'bc_on_snow', 'land_use', 'volcanic', 'solar', 'wmghgs', 'aerosol', 'albedo', 'anthro', 'natural']
for forcing in forcings:
AR6_forc[forcing] = np.zeros(5)
AR6_forc[forcing] = np.percentile(
(results['AR6-historical']['surface_temperature'][-1] - results['AR6-historical']['surface_temperature'][0])-
(results['remove_%s' % forcing]['surface_temperature'][-1] - results['remove_%s' % forcing]['surface_temperature'][0]), (5,16,50,84,95)
)
AR6_ecsforc[forcing] = np.zeros(5)
AR6_ecsforc[forcing] = np.percentile(
(results['AR6-historical_climuncert']['surface_temperature'][-1] - results['AR6-historical_climuncert']['surface_temperature'][0])-
(results['remove_%s_climuncert' % forcing]['surface_temperature'][-1] - results['remove_%s_climuncert' % forcing]['surface_temperature'][0]), (5,16,50,84,95)
)
# create some new aggregated categories
non_co2_ghgs = (
(
(results['AR6-historical']['surface_temperature'][-1] - results['AR6-historical']['surface_temperature'][0]) -
(results['remove_ch4']['surface_temperature'][-1] - results['remove_ch4']['surface_temperature'][0])
) +
(
(results['AR6-historical']['surface_temperature'][-1] - results['AR6-historical']['surface_temperature'][0]) -
(results['remove_n2o']['surface_temperature'][-1] - results['remove_n2o']['surface_temperature'][0])
) +
(
(results['AR6-historical']['surface_temperature'][-1] - results['AR6-historical']['surface_temperature'][0]) -
(results['remove_other_wmghg']['surface_temperature'][-1] - results['remove_other_wmghg']['surface_temperature'][0])
)
)
AR6_forc['ch4+n2o+cfc'] = np.percentile(non_co2_ghgs, (5,16,50,84,95))
non_co2_ghgs = (
(
(results['AR6-historical_climuncert']['surface_temperature'][-1] - results['AR6-historical_climuncert']['surface_temperature'][0]) -
(results['remove_ch4_climuncert']['surface_temperature'][-1] - results['remove_ch4_climuncert']['surface_temperature'][0])
) +
(
(results['AR6-historical_climuncert']['surface_temperature'][-1] - results['AR6-historical_climuncert']['surface_temperature'][0]) -
(results['remove_n2o_climuncert']['surface_temperature'][-1] - results['remove_n2o_climuncert']['surface_temperature'][0])
) +
(
(results['AR6-historical_climuncert']['surface_temperature'][-1] - results['AR6-historical_climuncert']['surface_temperature'][0]) -
(results['remove_other_wmghg_climuncert']['surface_temperature'][-1] - results['remove_other_wmghg_climuncert']['surface_temperature'][0])
)
)
AR6_ecsforc['ch4+n2o+cfc'] = np.percentile(non_co2_ghgs, (5,16,50,84,95))
AR6_forc['total'] = np.percentile(results['AR6-historical']['surface_temperature'][-1] - results['AR6-historical']['surface_temperature'][0], (5,16,50,84,95))
AR6_ecsforc['total'] = np.percentile(results['AR6-historical_climuncert']['surface_temperature'][-1] - results['AR6-historical_climuncert']['surface_temperature'][0], (5,16,50,84,95))
# -
print(AR6_ecsforc['natural'])
print(AR6_ecsforc['anthro'])
print(AR6_ecsforc['total'])
# +
fig, ax = pl.subplots()
gs = gridspec.GridSpec(1,2, width_ratios=[5,1])
ax = pl.subplot(gs[0,0])
# errorbar for owmghg is going to be a mess so pre-calculate it
#hot = ['#f03b20','#feb24c','#feeda0']
#hot = ['#de2d26','#fc9274','#fee0d2']
#cool=['#3182bd','#9ecae1']
hot = ['#cc404a','#ed8037','#ecd151']
cool=['#304fbf','#369ce8']
ax.barh(0, AR6_ecsforc['co2'][2], color=hot[0], lw=1, edgecolor='k')
eb = ax.errorbar(AR6_forc['co2'][2], 0.18, xerr=np.array([AR6_forc['co2'][2]-AR6_forc['co2'][0],AR6_forc['co2'][4]-AR6_forc['co2'][2]]).reshape((2,1)), capsize=3, color='k')
eb[-1][0].set_linestyle(':')
eb = ax.errorbar(AR6_ecsforc['co2'][2], -0.18, xerr=np.array([AR6_ecsforc['co2'][2]-AR6_ecsforc['co2'][0],AR6_ecsforc['co2'][4]-AR6_ecsforc['co2'][2]]).reshape((2,1)), capsize=3, color='k')
ax.barh(-1, AR6_ecsforc['ch4'][2], color=hot[0], lw=1, edgecolor='k')
ax.barh(-1, AR6_ecsforc['n2o'][2], color=hot[1], left=AR6_ecsforc['ch4'][2], lw=1, edgecolor='k')
ax.barh(-1, AR6_ecsforc['other_wmghg'][2], color=hot[2], left=AR6_ecsforc['ch4'][2]+AR6_ecsforc['n2o'][2], lw=1, edgecolor='k')
eb = ax.errorbar(AR6_forc['ch4+n2o+cfc'][2], -0.82, xerr=np.array([AR6_forc['ch4+n2o+cfc'][2]-AR6_forc['ch4+n2o+cfc'][0],AR6_forc['ch4+n2o+cfc'][4]-AR6_forc['ch4+n2o+cfc'][2]]).reshape((2,1)), capsize=3, color='k')
eb[-1][0].set_linestyle(':')
eb = ax.errorbar(AR6_ecsforc['ch4+n2o+cfc'][2], -1.18, xerr=np.array([AR6_ecsforc['ch4+n2o+cfc'][2]-AR6_ecsforc['ch4+n2o+cfc'][0],AR6_ecsforc['ch4+n2o+cfc'][4]-AR6_ecsforc['ch4+n2o+cfc'][2]]).reshape((2,1)), capsize=3, color='k')
ax.barh(-2, AR6_ecsforc['o3'][2], color=hot[0], lw=1, edgecolor='k')
eb = ax.errorbar(AR6_forc['o3'][2], -1.82, xerr=np.array([AR6_forc['o3'][2]-AR6_forc['o3'][0],AR6_forc['o3'][4]-AR6_forc['o3'][2]]).reshape((2,1)), capsize=3, color='k')
eb[-1][0].set_linestyle(':')
eb = ax.errorbar(AR6_ecsforc['o3'][2], -2.18, xerr=np.array([AR6_ecsforc['o3'][2]-AR6_ecsforc['o3'][0],AR6_ecsforc['o3'][4]-AR6_ecsforc['o3'][2]]).reshape((2,1)), capsize=3, color='k')
ax.barh(-3, AR6_ecsforc['h2o_stratospheric'][2], color=hot[0], lw=1, edgecolor='k')
eb = ax.errorbar(AR6_forc['h2o_stratospheric'][2], -2.82, xerr=np.array([AR6_forc['h2o_stratospheric'][2]-AR6_forc['h2o_stratospheric'][0],AR6_forc['h2o_stratospheric'][4]-AR6_forc['h2o_stratospheric'][2]]).reshape((2,1)), capsize=3, color='k')
eb[-1][0].set_linestyle(':')
eb = ax.errorbar(AR6_ecsforc['h2o_stratospheric'][2], -3.18, xerr=np.array([AR6_ecsforc['h2o_stratospheric'][2]-AR6_ecsforc['h2o_stratospheric'][0],AR6_ecsforc['h2o_stratospheric'][4]-AR6_ecsforc['h2o_stratospheric'][2]]).reshape((2,1)), capsize=3, color='k')
ax.barh(-4, AR6_ecsforc['land_use'][2], color=cool[0], lw=1, edgecolor='k')
eb = ax.errorbar(AR6_forc['land_use'][2], -3.82, xerr=np.array([AR6_forc['land_use'][2]-AR6_forc['land_use'][0],AR6_forc['land_use'][4]-AR6_forc['land_use'][2]]).reshape((2,1)), capsize=3, color='k')
eb[-1][0].set_linestyle(':')
eb = ax.errorbar(AR6_ecsforc['land_use'][2], -4.18, xerr=np.array([AR6_ecsforc['land_use'][2]-AR6_ecsforc['land_use'][0],AR6_ecsforc['land_use'][4]-AR6_ecsforc['land_use'][2]]).reshape((2,1)), capsize=3, color='k')
ax.barh(-4, AR6_ecsforc['bc_on_snow'][2], color=hot[0], lw=1, edgecolor='k')
eb = ax.errorbar(AR6_forc['bc_on_snow'][2], -3.82, xerr=np.array([AR6_forc['bc_on_snow'][2]-AR6_forc['bc_on_snow'][0],AR6_forc['bc_on_snow'][4]-AR6_forc['bc_on_snow'][2]]).reshape((2,1)), capsize=3, color='k')
eb[-1][0].set_linestyle(':')
eb = ax.errorbar(AR6_ecsforc['bc_on_snow'][2], -4.18, xerr=np.array([AR6_ecsforc['bc_on_snow'][2]-AR6_ecsforc['bc_on_snow'][0],AR6_ecsforc['bc_on_snow'][4]-AR6_ecsforc['bc_on_snow'][2]]).reshape((2,1)), capsize=3, color='k')
ax.barh(-5, AR6_ecsforc['contrails'][2], color=hot[0], lw=1, edgecolor='k')
eb = ax.errorbar(AR6_forc['contrails'][2], -4.82, xerr=np.array([AR6_forc['contrails'][2]-AR6_forc['contrails'][0],AR6_forc['contrails'][4]-AR6_forc['contrails'][2]]).reshape((2,1)), capsize=3, color='k')
eb[-1][0].set_linestyle(':')
eb = ax.errorbar(AR6_ecsforc['contrails'][2], -5.18, xerr=np.array([AR6_ecsforc['contrails'][2]-AR6_ecsforc['contrails'][0],AR6_ecsforc['contrails'][4]-AR6_ecsforc['contrails'][2]]).reshape((2,1)), capsize=3, color='k')
ax.barh(-6, AR6_ecsforc['aerosol-radiation_interactions'][2], color=cool[0], lw=1, edgecolor='k')
ax.barh(-6, AR6_ecsforc['aerosol-cloud_interactions'][2], color=cool[1], left=AR6_ecsforc['aerosol-radiation_interactions'][2], lw=1, edgecolor='k')
eb = ax.errorbar(AR6_forc['aerosol'][2], -5.82, xerr=np.array([AR6_forc['aerosol'][2]-AR6_forc['aerosol'][0],AR6_forc['aerosol'][4]-AR6_forc['aerosol'][2]]).reshape((2,1)), capsize=3, color='k')
eb[-1][0].set_linestyle(':')
eb = ax.errorbar(AR6_ecsforc['aerosol'][2], -6.18, xerr=np.array([AR6_ecsforc['aerosol'][2]-AR6_ecsforc['aerosol'][0],AR6_ecsforc['aerosol'][4]-AR6_ecsforc['aerosol'][2]]).reshape((2,1)), capsize=3, color='k')
ax.barh(-7, AR6_ecsforc['solar'][2], color=cool[0], lw=1, edgecolor='k')
eb = ax.errorbar(AR6_forc['solar'][2], -6.82, xerr=np.array([AR6_forc['solar'][2]-AR6_forc['solar'][0],AR6_forc['solar'][4]-AR6_forc['solar'][2]]).reshape((2,1)), capsize=3, color='k')
eb[-1][0].set_linestyle(':')
eb = ax.errorbar(AR6_ecsforc['solar'][2], -7.18, xerr=np.array([AR6_ecsforc['solar'][2]-AR6_ecsforc['solar'][0],AR6_ecsforc['solar'][4]-AR6_ecsforc['solar'][2]]).reshape((2,1)), capsize=3, color='k')
ax.barh(-8, AR6_ecsforc['volcanic'][2], color=cool[0], lw=1, edgecolor='k')
eb = ax.errorbar(AR6_forc['volcanic'][2], -7.82, xerr=np.array([AR6_forc['volcanic'][2]-AR6_forc['volcanic'][0],AR6_forc['volcanic'][4]-AR6_forc['volcanic'][2]]).reshape((2,1)), capsize=3, color='k')
eb[-1][0].set_linestyle(':')
eb = ax.errorbar(AR6_ecsforc['volcanic'][2], -8.18, xerr=np.array([AR6_ecsforc['volcanic'][2]-AR6_ecsforc['volcanic'][0],AR6_ecsforc['volcanic'][4]-AR6_ecsforc['volcanic'][2]]).reshape((2,1)), capsize=3, color='k')
#ax.barh(-9, AR6_ecsforc['anthro'][2], color=hot[0], lw=1, edgecolor='k')
#ax.barh(-9, AR6_ecsforc['natural'][2], color=cool[0], lw=1, edgecolor='k')
ax.barh(-9, AR6_ecsforc['total'][2], color=hot[0], lw=1, edgecolor='k')
eb = ax.errorbar(AR6_forc['total'][2], -8.82, xerr=np.array([AR6_forc['total'][2]-AR6_forc['total'][0],AR6_forc['total'][4]-AR6_forc['total'][2]]).reshape((2,1)), capsize=3, color='k')
eb[-1][0].set_linestyle(':')
eb = ax.errorbar(AR6_ecsforc['total'][2], -9.18, xerr=np.array([AR6_ecsforc['total'][2]-AR6_ecsforc['total'][0],AR6_ecsforc['total'][4]-AR6_ecsforc['total'][2]]).reshape((2,1)), capsize=3, color='k')
# labels and prettiness
ax.text(1.0, -1, 'CH$_4$',ha='center',va='center', color=hot[0], size=9)
ax.text(1.28, -1,'N$_2$O',ha='center',va='center', color=hot[1], size=9)
ax.text(1.45, -1, 'Halogens', va='center', color=hot[2], size=9)
#ax.text(1.1, -2, 'Stratospheric', va='center', ha='left', color=hot[1])
#ax.text(0.5, -2, 'Tropospheric', va='center', ha='left', color=hot[0])
ax.text(-0.23,-4, 'Land use', va='center', ha='right', color=cool[0], size=9)
ax.text(0.16, -4, 'Light absorbing particles\non snow and ice', va='center', ha='left', color=hot[0], size=9)
ax.text(0.05, -6, 'Aerosol-cloud', va='center', ha='left', color=cool[1], size=9)
ax.text(0.7, -6, 'Aerosol-radiation', va='center', ha='left', color=cool[0], size=9)
#ax.text(-0.75, -9, 'Anthropogenic', va='center', ha='center', color=hot[0], size=9)
#ax.text(-0.2, -9, 'Natural', va='center', ha='center', color=hot[1], size=9)
ax.text(1.91, -3, 'Anthropogenic', va='center', ha='center', rotation=270)
ax.text(1.91, -7.5, 'Natural', va='center', ha='center', rotation=270)
ax.set_ylim(-9.5,0.5)
ax.set_xlim(-1.2,2.0)
ax.axvline(0, color='k', lw=0.6)
ax.axhline(-6.5, color='k', lw=0.6)
ax.axhline(-8.5, color='k', lw=0.6)
ax.set_yticks(np.arange(-9,1));
ax.set_xlabel('$^{\circ}$C')
ax.set_title('')
ax.set_yticklabels(['Carbon dioxide','Other well-mixed\ngreenhouse gases','Ozone','Stratospheric\nwater vapour', 'Albedo', 'Contrails & aviation-\ninduced cirrus', 'Aerosol', 'Solar', 'Volcanic', 'Total'][::-1]);
ax.set_title('Simulated temperature contributions in 2019 relative to 1750', size=11)
# legend
eb = ax.errorbar(0.5, -7.2, xerr=0.1, color='k', capsize=2, lw=0.8)
eb[-1][0].set_linestyle(':')
eb = ax.errorbar(0.5, -7.75, xerr=0.1, color='k', capsize=2, lw=0.8)
ax.text(0.68, -7.185, 'Forcing uncertainty', va='center', fontsize=8);
ax.text(0.68, -7.735, 'Forcing + climate \nsensitivity uncertainty', va='center', fontsize=8);
rect = mp.Rectangle((0.35, -8.17), 1.3, 1.28, facecolor='None', edgecolor='k', lw=0.5)
ax.add_patch(rect)
# values
ax_values = pl.subplot(gs[0,1])
ax_values.text(0.5,0,'{:.2f} [{:.2f} to {:.2f}]'.format(AR6_ecsforc['co2'][2], AR6_ecsforc['co2'][0], AR6_ecsforc['co2'][4]), ha='center', va='center', size=9)
ax_values.text(0.5,-0.6,'{:.2f} [{:.2f} to {:.2f}]'.format(AR6_ecsforc['ch4'][2], AR6_ecsforc['ch4'][0], AR6_ecsforc['ch4'][4]), ha='center', va='center', size=9, color=hot[0])
ax_values.text(0.5,-1.0,'{:.2f} [{:.2f} to {:.2f}]'.format(AR6_ecsforc['n2o'][2], AR6_ecsforc['n2o'][0], AR6_ecsforc['n2o'][4]), ha='center', va='center', size=9, color=hot[1])
ax_values.text(0.5,-1.4,'{:.2f} [{:.2f} to {:.2f}]'.format(AR6_ecsforc['other_wmghg'][2], AR6_ecsforc['other_wmghg'][0], AR6_ecsforc['other_wmghg'][4]), ha='center', va='center', size=9, color=hot[2])
ax_values.text(0.5,-2,'{:.2f} [{:.2f} to {:.2f}]'.format(AR6_ecsforc['o3'][2], AR6_ecsforc['o3'][0], AR6_ecsforc['o3'][4]), ha='center', va='center', size=9)
ax_values.text(
0.5,-3,'{:.2f} [{:.2f} to {:.2f}]'.format(
AR6_ecsforc['h2o_stratospheric'][2],
AR6_ecsforc['h2o_stratospheric'][0] if abs(AR6_ecsforc['h2o_stratospheric'][0]) >= 0.005 else abs(AR6_ecsforc['h2o_stratospheric'][0]),
AR6_ecsforc['h2o_stratospheric'][4]), ha='center', va='center', size=9
)
ax_values.text(0.5,-3.8,'{:.2f} [{:.2f} to {:.2f}]'.format(AR6_ecsforc['land_use'][2], AR6_ecsforc['land_use'][0], AR6_ecsforc['land_use'][4]), ha='center', va='center', size=9, color=cool[0])
ax_values.text(
0.5,-4.2,'{:.2f} [{:.2f} to {:.2f}]'.format(
AR6_ecsforc['bc_on_snow'][2],
AR6_ecsforc['bc_on_snow'][0] if abs(AR6_ecsforc['bc_on_snow'][0]) >= 0.005 else abs(AR6_ecsforc['bc_on_snow'][0]),
AR6_ecsforc['bc_on_snow'][4]), ha='center', va='center', size=9, color=hot[0]
)
ax_values.text(0.5,-5,'{:.2f} [{:.2f} to {:.2f}]'.format(AR6_ecsforc['contrails'][2], AR6_ecsforc['contrails'][0], AR6_ecsforc['contrails'][4]), ha='center', va='center', size=9)
ax_values.text(0.5,-5.8,'{:.2f} [{:.2f} to {:.2f}]'.format(AR6_ecsforc['aerosol-radiation_interactions'][2], AR6_ecsforc['aerosol-radiation_interactions'][0], AR6_ecsforc['aerosol-radiation_interactions'][4]), ha='center', va='center', size=9, color=cool[0])
ax_values.text(0.5,-6.2,'{:.2f} [{:.2f} to {:.2f}]'.format(AR6_ecsforc['aerosol-cloud_interactions'][2], AR6_ecsforc['aerosol-cloud_interactions'][0], AR6_ecsforc['aerosol-cloud_interactions'][4]), ha='center', va='center', size=9, color=cool[1])
ax_values.text(
0.5,-7,'{:.2f} [{:.2f} to {:.2f}]'.format(
AR6_ecsforc['solar'][2] if abs(AR6_ecsforc['solar'][2]) >= 0.005 else abs(AR6_ecsforc['solar'][2]),
AR6_ecsforc['solar'][0] if abs(AR6_ecsforc['solar'][0]) >= 0.005 else abs(AR6_ecsforc['solar'][0]),
AR6_ecsforc['solar'][4] if abs(AR6_ecsforc['solar'][4]) >= 0.005 else abs(AR6_ecsforc['solar'][4]),
), ha='center', va='center', size=9)
ax_values.text(0.5,-8,'{:.2f} [{:.2f} to {:.2f}]'.format(AR6_ecsforc['volcanic'][2], AR6_ecsforc['volcanic'][0], AR6_ecsforc['volcanic'][4]), ha='center', va='center', size=9)
ax_values.text(0.5,-9,'{:.2f} [{:.2f} to {:.2f}]'.format(AR6_ecsforc['total'][2], AR6_ecsforc['total'][0], AR6_ecsforc['total'][4]), ha='center', va='center', size=9)
ax_values.set_title('$^{\circ}$C', size=9)
ax_values.set_xlim(0,1)
ax_values.set_ylim(-9.5, 0.5)
ax_values.spines['left'].set_visible(False)
ax_values.spines['right'].set_visible(False)
ax_values.spines['bottom'].set_visible(False)
ax_values.spines['top'].set_visible(False)
ax_values.set_yticklabels([])
ax_values.set_xticks([])
# Create the figure
fig.subplots_adjust(left=0.2)
pl.tight_layout()
pl.savefig('../figures/fig7.7.png', dpi=300)
pl.savefig('../figures/fig7.7.pdf')
# +
rebased_climuncert = results['AR6-historical_climuncert']['surface_temperature'] - np.mean(results['AR6-historical_climuncert']['surface_temperature'][100:151,:], axis=0)
rebased = results['AR6-historical']['surface_temperature'] - np.mean(results['AR6-historical']['surface_temperature'][100:151,:], axis=0)
pl.fill_between(np.arange(1750,2020),
np.percentile(rebased_climuncert, 5, axis=1),
np.percentile(rebased_climuncert, 95, axis=1),
alpha=0.3)
pl.plot(np.arange(1750,2020),np.median(rebased_climuncert, axis=1), label='emulated - climuncert');
pl.fill_between(np.arange(1750,2020),
np.percentile(rebased, 5, axis=1),
np.percentile(rebased, 95, axis=1),
alpha=0.3)
pl.plot(np.arange(1750,2020),np.median(rebased, axis=1), label='emulated - forcing only');
xl = pd.read_excel('../data_input/observations/AR6 FGD assessment time series - GMST and GSAT.xlsx', skiprows=1, skipfooter=28)
temp_gmst=xl['4-set mean'].values
temp_year=xl['Unnamed: 0'].values
pl.plot(temp_year, temp_gmst, label='chapter 2 GMST')
pl.legend()
temp_gmst[-1]
# +
rebased_climuncert = results['AR6-historical_climuncert']['surface_temperature'] - results['remove_volcanic_climuncert']['surface_temperature']
rebased = results['AR6-historical']['surface_temperature'] - results['remove_volcanic']['surface_temperature']
pl.fill_between(np.arange(1750,2020),
np.percentile(rebased_climuncert-rebased_climuncert[0,:], 5, axis=1),
np.percentile(rebased_climuncert-rebased_climuncert[0,:], 95, axis=1),
alpha=0.3)
pl.plot(np.arange(1750,2020),np.median(rebased_climuncert-rebased_climuncert[0,:], axis=1), label='emulated - climuncert');
pl.fill_between(np.arange(1750,2020),
np.percentile(rebased-rebased[0,:], 5, axis=1),
np.percentile(rebased-rebased[0,:], 95, axis=1),
alpha=0.3)
pl.plot(np.arange(1750,2020),np.median(rebased-rebased[0,:], axis=1), label='emulated - climuncert');
np.median(rebased_climuncert[-1,:]-rebased_climuncert[0,:])
pl.axhline(0, ls=':', color='k')
# -
AR6_forc['total'], AR6_ecsforc['total']
# ## numbers for ES
AR6_ecsforc['anthro'][[0,2,4]]
AR6_ecsforc['total'][[0,2,4]]
AR6_ecsforc['wmghgs'][[0,2,4]]
AR6_ecsforc['o3'][[0,2,4]]
AR6_ecsforc['aerosol'][[0,2,4]]
AR6_ecsforc['albedo'][[0,2,4]]
AR6_ecsforc['natural'][[0,2,4]]
# ## numbers for chapter 1
np.percentile(
(results['AR6-historical_climuncert']['surface_temperature'][100:151].mean(axis=0) - results['AR6-historical_climuncert']['surface_temperature'][0])-
(results['remove_anthro_climuncert']['surface_temperature'][100:151].mean(axis=0) - results['remove_anthro_climuncert']['surface_temperature'][0]), (5,16,50,84,95)
)
|
notebooks/220_chapter7_fig7.7.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exceptions
# Exceptions which are events that can modify the *flow* of control through a program.
#
# In Python, exceptions are triggered automatically on errors, and they can be triggered and intercepted by your code.
#
# They are processed by **four** statements we’ll study in this notebook, the first of which has two variations (listed separately here) and the last of which was an optional extension until Python 2.6 and 3.0:
#
# * `try/except`:
# * Catch and recover from exceptions raised by Python, or by you
#
# * `try/finally`:
# * Perform cleanup actions, whether exceptions occur or not.
#
# * `raise`:
# * Trigger an exception manually in your code.
#
# * `assert`:
# * Conditionally trigger an exception in your code.
#
# * `with/as`:
# * Implement context managers in Python 2.6, 3.0, and later (optional in 2.5).
# # `try/except` Statement
# ```
# try:
# statements # Run this main action first
# except name1:
# # Run if name1 is raised during try block
# statements
# except (name2, name3):
# # Run if any of these exceptions occur
# statements
# except name4 as var:
# # Run if name4 is raised, assign instance raised to var
# statements
# except: # Run for all other exceptions raised
# statements
# else:
# statements # Run if no exception was raised during try block
# ```
list_of_numbers = [number for number in range(1, 100)]
print(list_of_numbers)
# +
dictionary_of_numbers = {}
for number in list_of_numbers:
dictionary_of_numbers[number**2] = number
try:
index = list_of_numbers.index(2)
value = dictionary_of_numbers[index]
except (ValueError, KeyError):
print('Error Raised, but Controlled! ')
else:
# This executes ONLY if no exception is raised
print('Getting number at position %d : %d' % (index, value))
finally:
# Do cleanup operations
print('Cleaning UP')
# -
# # `try/finally` Statement
# The other flavor of the try statement is a specialization that has to do with finalization (a.k.a. termination) actions. If a finally clause is included in a try, Python will always run its block of statements “on the way out” of the try statement, whether an exception occurred while the try block was running or not.
#
# In it's general form, it is:
#
# ```
# try:
# statements # Run this action first
# finally:
# statements # Always run this code on the way out
# ```
# <a name="ctx"></a>
# # `with/as` Context Managers
# Python 2.6 and 3.0 introduced a new exception-related statement—the with, and its optional as clause. This statement is designed to work with context manager objects, which support a new method-based protocol, similar in spirit to the way that iteration tools work with methods of the iteration protocol.
# ## Context Manager Intro
# ### Basic Usage:
#
# ```
# with expression [as variable]:
# with-block
# ```
# ### Classical Usage
#
# ```python
#
# with open(r'C:\misc\data') as myfile:
# for line in myfile:
# print(line)
# # ...more code here...
# ```
#
# ... even using multiple context managers:
#
# ```python
# with open('script1.py') as f1, open('script2.py') as f2:
# for (linenum, (line1, line2)) in enumerate(zip(f1, f2)):
# if line1 != line2:
# print('%s\n%r\n%r' % (linenum, line1, line2))
# ```
# ### How it works
#
# 1. The expression is evaluated,resulting in an object known as a **context manager** that must have `__enter__` and `__exit__` methods
#
# 2. The context manager’s `__enter__` method is called. The value it returns is assigned to the variable in the as clause if present, or simply discarded otherwise
#
# 3. The code in the nested with block is executed.
#
# 4. If the with block raises an exception, the `__exit__(type,value,traceback)` method is called with the exception details. These are the same three values returned by `sys.exc_info` (Python function). If this method returns a `false` value, the exception is **re-raised**; otherwise, the exception is terminated. The exception should normally be reraised so that it is propagated outside the with statement.
#
# 5. If the with block does not raise an exception, the `__exit__` method is still called, but its type, value, and traceback arguments are all passed in as `None`.
# ## Usage with Exceptions
class TraceBlock:
def message(self, arg):
print('running ' + arg)
def __enter__(self):
print('starting with block')
return self
def __exit__(self, exc_type, exc_value, exc_tb):
if exc_type is None:
print('exited normally\n')
else:
print('raise an exception! ' + str(exc_type))
return False # Propagate
with TraceBlock() as action:
action.message('test 1')
print('reached')
with TraceBlock() as action:
action.message('test 2')
raise TypeError()
print('not reached')
# ## User Defined Exceptions
# +
class AlreadyGotOne(Exception):
pass
def gail():
raise AlreadyGotOne()
# -
try:
gail()
except AlreadyGotOne:
print('got exception')
# +
class Career(Exception):
def __init__(self, job, *args, **kwargs):
super(Career, self).__init__(*args, **kwargs)
self._job = job
def __str__(self):
return 'So I became a waiter of {}'.format(self._job)
raise Career('Engineer')
|
L0/08 Exceptions.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/akashkulkarni1192/AI_N-SlidingPuzzle-Solver/blob/master/Attendee_Intro_to_Visualizations_with_Python.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="cpwsxUdOthZh" colab_type="text"
# Import the California Housing dataset.
#
# + id="fuynGZ8jDsca" colab_type="code" colab={}
DATASETS_URL = "https://github.com/ageron/handson-ml/raw/master/datasets"
import os
import tarfile
from six.moves import urllib
HOUSING_PATH = "datasets/housing"
HOUSING_URL = DATASETS_URL + "/housing/housing.tgz"
def fetch_housing_data(housing_url=HOUSING_URL, housing_path=HOUSING_PATH):
if not os.path.exists(housing_path):
os.makedirs(housing_path)
tgz_path = os.path.join(housing_path, "housing.tgz")
urllib.request.urlretrieve(housing_url, tgz_path)
housing_tgz = tarfile.open(tgz_path)
housing_tgz.extractall(path=housing_path)
housing_tgz.close()
fetch_housing_data()
import pandas as pd
def load_housing_data(housing_path=HOUSING_PATH):
csv_path = os.path.join(housing_path, "housing.csv")
return pd.read_csv(csv_path)
housing = load_housing_data()
# + id="QSgmIYFNcMGc" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 297} outputId="f0a0a63c-d4b1-4f51-9ec2-ba8d073b858f"
housing.describe()
# + id="u1MDtO8qdcGG" colab_type="code" colab={}
import seaborn as sns
import matplotlib.pyplot as plt
# + id="42Nbd22VdlQ-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 446} outputId="2db7b09f-5a52-41e4-c778-330ff61a8a8c"
plt.xlabel('freq')
plt.ylabel('median house val')
plt.hist(housing['median_house_value'])
# + id="kmIWc65Reszt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 347} outputId="36fa00fc-3cd1-492d-ec08-a8785636b9ca"
housing['ocean_proximity'].value_counts().plot(kind='pie')
# + id="9zXyPegde26P" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 420} outputId="958bd795-9401-417b-e120-7a850e9bd990"
housing['ocean_proximity'].value_counts().plot(kind='bar')
# + id="WosdC2Qq1t2e" colab_type="code" outputId="af80b7d0-6c88-4b82-9907-169b14bf0fc0" colab={"base_uri": "https://localhost:8080/", "height": 102}
h_ocean = 9136/20640
inland = 6551/20640
n_ocean = 2658/20640
n_bay = 2290/20640
island = 5/20640
print(h_ocean)
print(inland)
print(n_ocean)
print(n_bay)
print(island)
# + id="hNfIJI1Y1TRY" colab_type="code" outputId="c1b52853-f7d3-4549-a48e-708934d7a297" colab={"base_uri": "https://localhost:8080/", "height": 403}
# Pie chart
labels = ['<1H OCEAN', 'INLAND', 'NEAR OCEAN', 'NEAR BAY', 'ISLAND']
sizes = [44, 32, 13, 11, 0.2]
#colors
colors = ['#66b3ff','#A5D6A7', '#D7CCC8', '#FDD835', '#263238']
fig1, ax1 = plt.subplots()
patches, texts, autotexts = ax1.pie(sizes, colors = colors, labels=labels, autopct='%1.1f%%', startangle=90)
for text in texts:
text.set_color('black')
for autotext in autotexts:
autotext.set_color('black')
# Equal aspect ratio ensures that pie is drawn as a circle
ax1.axis('equal')
plt.tight_layout()
plt.legend(loc = 'upper left')
plt.show()
# + id="_aArOFeQg3VM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 364} outputId="01c2f1a5-b51b-4d34-fec0-ad0027c9ee93"
plt.scatter(housing['median_income'], housing['median_house_value'])
# + id="enEwG1IthP5A" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 328} outputId="1f6e7f53-0884-4a0f-f56c-0a2f07257ddd"
housing.corr()
# + id="XZ8QikTEhm7R" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 187} outputId="a405f36c-c3dd-49ca-c316-8eab741fb977"
housing.corr()['median_house_value'].sort_values(ascending=False)
# + [markdown] id="P1i1ZlpF8w_G" colab_type="text"
# Let's move on to another type of plot that is useful for showing correlations. We would expect to see a correlation between the income and the house value, as more income would most likely mean a nicer home.
# + id="TQ3EUT1q1s0B" colab_type="code" outputId="ff85fdb6-9259-414e-b591-8885f623e533" colab={"base_uri": "https://localhost:8080/", "height": 701}
# Set up the matplotlib figure
f, ax = plt.subplots(figsize=(12, 10))
plt.title('Pearson Correlation of Housing Features')
# Draw the heatmap using seaborn
sns.heatmap(housing.corr(),linewidths=0.25, square=True, cmap = "BrBG", linecolor='black', annot=True)
# + id="yWTP3oDqiP1I" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 435} outputId="aa3e9e75-c463-4341-d1fb-12d03cdfc70d"
housing.plot(kind='scatter', x = 'longitude', y='latitude')
# + id="QuzLZqTVi0rN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 415} outputId="deb8c1fc-5d45-4880-9ac3-487e0e424ea8"
housing.plot(kind='scatter', x = 'longitude', y='latitude', alpha=0.4)
# + id="y9xPkQ_0i8h3" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 432} outputId="468a39b6-00ce-4f03-8d4b-736c2e267c1f"
housing.plot(kind='scatter', x = 'longitude', y='latitude', alpha=0.4, s= housing['population']/100, label='population', figsize=(10,7), c='median_house_value', cmap = plt.get_cmap('jet'), colorbar=True)
plt.legend()
# shows higher house value around SanFrancisco and Los Angeles areas
# + id="W9W89mGCjayk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 725} outputId="66a6682d-eaf7-4f8d-ab11-8e3119be50da"
ocean_proimixty_graph = sns.lmplot(data=housing, x='longitude', y='latitude', fit_reg=False, hue='ocean_proximity', size=10, aspect=1.5, scatter_kws={'s':200})
# plotting ocean proximity on top of the scatter plot. This plot looks okay. If INLAND(red color) was seen around ocean areas - that would mean some of the data points were mislabelled and we need to clean it
# + id="mZ4mrzMlkO1W" colab_type="code" colab={}
|
Attendee_Intro_to_Visualizations_with_Python.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Python Fundamentals
#
# - Variables, Expressions and Statements
#
# Reference: https://www.py4e.com/lessons
# 2.3 Write a program to prompt the user for hours and rate per hour using input to compute gross pay. Use 35 hours and a rate of 2.75 per hour to test the program (the pay should be 96.25). You should use input to read a string and float() to convert the string to a number. Do not worry about error checking or bad user data.
# +
hrs = input("Enter Hours:")
type(hrs)
rph = input("Enter rate per hour:")
type(rph)
# convert rate per hour to float
rph= float(rph)
pay = hrs* rph
print("Your pay is: ", pay)
# -
# See this [SO post](https://stackoverflow.com/questions/485789/why-do-i-get-typeerror-cant-multiply-sequence-by-non-int-of-type-float/27488582)
hrs = input("Enter Hours:")
print ("Data type is ",type(hrs))
rph = input("Enter rate per hour:")
print ("Data type is ",type(rph))
# +
hrs = float(input("Enter Hours:"))
rph = float(input("Enter rate per hour:"))
# convert rate per hour to float
pay = hrs* rph
print("Your pay is: ", pay)
# -
# ### Conditional Execution
#
# IF Statement
# 3.1 Write a program to prompt the user for hours and rate per hour using input to compute gross pay. Pay the hourly rate for the hours up to 40 and 1.5 times the hourly rate for all hours worked above 40 hours. Use 45 hours and a rate of 10.50 per hour to test the program (the pay should be 498.75). You should use input to read a string and float() to convert the string to a number. Do not worry about error checking the user input - assume the user types numbers properly.
# +
hrs = float(input("Enter Hours:"))
rph = float(input("Enter rate per hour:"))
# convert rate per hour to float
pay = hrs* rph
if (hrs<=40):
pay = hrs*rph
if (hrs>40):
#rph = 1.5*rph
pay = hrs*1.5*rph
print("Your pay is: ", pay)
# -
# 3.3 Write a program to prompt for a score between `0.0` and `1.0`. If the score is out of range, print an error. If the score is between `0.0` and `1.0`, print a grade using the following table:
#
# Score Grade
#
# `>= 0.9` A
#
# `>= 0.8` B
#
# `>= 0.7` C
#
# `>= 0.6` D
#
# `< 0.6` F
#
# If the user enters a value out of range, print a suitable error message and exit. For the test, enter a score of 0.85.
# +
score = float(input("Enter Score: "))
type(score)
if (score>1.0):
print ("Sorry! Out of range value")
if (score<= 1.0 and score >=0.0):
if(score >= 0.9):
print ("A")
elif(score >= 0.8):
print ("B")
elif(score >= 0.7):
print ("C")
elif(score >= 0.6):
print ("D")
elif(score < 0.6):
print ("F")
else:
print ("Logic is wrong")
# -
|
python/fundamentals/python fundamentals.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Backtest a Single Model
#
# The way to gauge the performance of a time-series model is through re-training models with different historic periods and check their forecast within certain steps. This is similar to a time-based style cross-validation. More often, we called it `backtest` in time-series modeling.
#
# The purpose of this notebook is to illustrate how to do 'backtest' on a single model using `BackTester`
#
# `BackTester` will compose a `TimeSeriesSplitter` within it, but `TimeSeriesSplitter` is useful as a standalone, in case there are other tasks to perform that requires splitting but not backtesting. You can also retrieve the composed `TimeSeriesSplitter` object from `BackTester` to utilize the additional methods in `TimeSeriesSplitter`
#
# Currently, there are two schemes supported for the back-testing engine: expanding window and rolling window.
#
# * expanding window: for each back-testing model training, the train start date is fixed, while the train end date is extended forward.
# * rolling window: for each back-testing model training, the training window length is fixed but the window is moving forward.
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
from orbit.models import DLT
from orbit.diagnostics.backtest import BackTester, TimeSeriesSplitter
from orbit.diagnostics.plot import plot_bt_predictions
from orbit.diagnostics.metrics import smape, wmape
from orbit.utils.dataset import load_iclaims
from orbit.utils.plot import get_orbit_style
# -
# %load_ext autoreload
# %autoreload 2
# ## Load data
# +
raw_data = load_iclaims()
data = raw_data.copy()
print(data.shape)
data.head(5)
# -
# ## Create a BackTester
# +
# instantiate a model
dlt = DLT(date_col='week',
response_col='claims',
regressor_col=['trend.unemploy', 'trend.filling', 'trend.job'],
seasonality=52,
estimator='stan-map')
# -
bt = BackTester(model=dlt,
df=data,
min_train_len=100,
incremental_len=100,
forecast_len=20)
# ## Backtest Fit and Predict
#
# The most expensive portion of backtesting is fitting the model iteratively. Thus, we separate the API calls for `fit_predict` and `score` to avoid redundant computation for multiple metrics or scoring methods.
# + tags=[]
bt.fit_predict();
# -
# Once `fit_predict()` is called, the fitted models and predictions can be easily retrieved from `BackTester`. Here the data is grouped by the date, split_key, and whether or not that observation is part of the training or test data.
# + jupyter={"outputs_hidden": true} tags=[]
predicted_df = bt.get_predicted_df()
predicted_df.head()
# -
# We also provide a plotting utility to visualize the predictions against the actuals for each split.
plot_bt_predictions(predicted_df, metrics=smape, ncol=2, include_vline=True);
# Users might find this useful for any custom computations that may need to be performed on the set of predicted data. Note that the columns are renamed to generic and consistent names.
# Sometimes, it might be useful to match the data back to the original dataset for ad-hoc diagnostics. This can easily be done by merging back to the orignal dataset
predicted_df.merge(data, left_on='date', right_on='week')
# ## Backtest Scoring
#
# The main purpose of `BackTester` are the evaluation metrics. Some of the most widely used metrics are implemented and built into the `BackTester` API.
#
# The default metric list is **smape, wmape, mape, mse, mae, rmsse**.
bt.score()
# It is possible to filter for only specific metrics of interest, or even implement your own callable and pass into the `score()` method. For example, see this function that uses last observed value as a predictor and computes the `mse`. Or `naive_error` which computes the error as the delta between predicted values and the training period mean.
#
# Note these are not really useful error metrics, just showing some examples of callables you can use ;)
# + code_folding=[]
def mse_naive(test_actual):
actual = test_actual[1:]
predicted = test_actual[:-1]
return np.mean(np.square(actual - predicted))
def naive_error(train_actual, test_predicted):
train_mean = np.mean(train_actual)
return np.mean(np.abs(test_predicted - train_mean))
# -
bt.score(metrics=[mse_naive, naive_error])
# It doesn't take additional time to refit and predict the model, since the results are stored when `fit_predict()` is called. Check docstrings for function criteria that is required for it to be supported with this API.
# In some cases, we may want to evaluate our metrics on both train and test data. To do this you can call score again with the following indicator
bt.score(include_training_metrics=True)
# ## Backtest Get Models
#
# In cases where `BackTester` doesn't cut it or for more custom use-cases, there's an interface to export the `TimeSeriesSplitter` and predicted data, as shown earlier. It's also possible to get each of the fitted models for deeper diving.
fitted_models = bt.get_fitted_models()
# +
model_1 = fitted_models[0]
model_1.get_regression_coefs()
# -
# ### Get TimeSeriesSplitter
#
# BackTester composes a TimeSeriesSplitter within it, but TimeSeriesSplitter can also be created on its own as a standalone object. See section below on TimeSeriesSplitter for more details on how to use the splitter.
#
# All of the additional TimeSeriesSplitter args can also be passed into BackTester on instantiation
ts_splitter = bt.get_splitter()
ts_splitter.plot()
plt.grid();
# ## Appendix
# ### Create a TimeSeriesSplitter
# #### Expanding window
min_train_len = 380
forecast_len = 20
incremental_len = 20
ex_splitter = TimeSeriesSplitter(df=data,
min_train_len=min_train_len,
incremental_len=incremental_len,
forecast_len=forecast_len,
window_type='expanding',
date_col='week')
print(ex_splitter)
# +
ex_splitter.plot()
plt.grid();
# -
# #### Rolling window
roll_splitter = TimeSeriesSplitter(df=data,
min_train_len=min_train_len,
incremental_len=incremental_len,
forecast_len=forecast_len,
window_type='rolling',
date_col='week')
roll_splitter.plot()
plt.grid();
# #### Specifying number of splits
# User can also define number of splits using `n_splits` instead of specifying minimum training length. That way, minimum training length will be automatically calculated.
ex_splitter2 = TimeSeriesSplitter(df=data,
min_train_len=min_train_len,
incremental_len=incremental_len,
forecast_len=forecast_len,
n_splits=5,
window_type='expanding',
date_col='week')
# +
ex_splitter2.plot()
plt.grid();
|
examples/backtest.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Importing Neccessary Libraries
# + id="ptx3OFDpsTsz"
import tensorflow as tf
from tensorflow.keras import Sequential
from tensorflow.keras.layers import Conv2D,MaxPool2D,Dense,Flatten,BatchNormalization,Dropout
from tensorflow.keras.optimizers import Adam
from tensorflow.keras.preprocessing import image
# + id="dw_QhGxftEN0"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# + id="dYKU9llftPit"
from sklearn.model_selection import train_test_split
from tqdm import tqdm
# -
# # Reading the Dataset
# + id="ZQGbkFZvuwDK"
df = pd.read_csv('input/train.csv')
# -
# Taking 30 percentage of the dataset for this project as using whole dataset crashes the memory threshold of kaggle kernels
df = df.head(2300)
# + executionInfo={"elapsed": 83822, "status": "ok", "timestamp": 1579525193086, "user": {"displayName": "<NAME>", "photoUrl": "https://lh6.googleusercontent.com/-t7IqSC7tKDY/AAAAAAAAAAI/AAAAAAAAhPo/INNzCzvCptg/s64/photo.jpg", "userId": "04862212736858688671"}, "user_tz": -330} id="Vnd8IQNHu6Pt" outputId="70cef3fa-a3c1-42b2-9e08-b0f0f20662ae"
df.head()
# -
# # Converting the images into Numpy array to train the CNN
# + executionInfo={"elapsed": 180313, "status": "ok", "timestamp": 1579525289590, "user": {"displayName": "<NAME>", "photoUrl": "https://lh6.googleusercontent.com/-t7IqSC7tKDY/AAAAAAAAAAI/AAAAAAAAhPo/INNzCzvCptg/s64/photo.jpg", "userId": "04862212736858688671"}, "user_tz": -330} id="WzqMmm4Gu7Gj" outputId="837cff8c-cd8a-47af-9d24-e3b3e9b3e9a6"
width = 350
height = 350
X = []
for i in tqdm(range(df.shape[0])):
path = 'input/Images/'+df['Id'][i]+'.jpg'
img = image.load_img(path,target_size=(width,height,3))
img = image.img_to_array(img)
img = img/255.0
X.append(img)
X = np.array(X)
# -
X.shape
# + executionInfo={"elapsed": 180292, "status": "ok", "timestamp": 1579525289594, "user": {"displayName": "<NAME>", "photoUrl": "https://lh6.googleusercontent.com/-t7IqSC7tKDY/AAAAAAAAAAI/AAAAAAAAhPo/INNzCzvCptg/s64/photo.jpg", "userId": "04862212736858688671"}, "user_tz": -330} id="9F1Iu__BxxLi" outputId="177d1261-60c0-456e-9754-a7143f0ec6d1"
y = df.drop(['Id','Genre'],axis=1)
y = y.to_numpy()
y.shape
# + id="vcEBnVf5x_St"
X_train,X_test,y_train,y_test=train_test_split(X,y,test_size=0.1)
# -
# # Preparing the model
# + id="almDiwamzS_T"
model = Sequential()
model.add(Conv2D(16,kernel_size=(3,3),activation='relu',input_shape=X_train[0].shape))
model.add(BatchNormalization())
model.add(MaxPool2D(2,2))
model.add(Dropout(0.3))
model.add(Conv2D(32,kernel_size=(3,3),activation='relu'))
model.add(BatchNormalization())
model.add(MaxPool2D(2,2))
model.add(Dropout(0.3))
model.add(Conv2D(64,kernel_size=(3,3),activation='relu'))
model.add(BatchNormalization())
model.add(MaxPool2D(2,2))
model.add(Dropout(0.4))
model.add(Flatten())
model.add(Dense(128,activation='relu'))
model.add(BatchNormalization())
model.add(Dropout(0.5))
model.add(Dense(25,activation='sigmoid'))
# + executionInfo={"elapsed": 203386, "status": "ok", "timestamp": 1579525312719, "user": {"displayName": "<NAME>", "photoUrl": "https://lh6.googleusercontent.com/-t7IqSC7tKDY/AAAAAAAAAAI/AAAAAAAAhPo/INNzCzvCptg/s64/photo.jpg", "userId": "04862212736858688671"}, "user_tz": -330} id="T0ziT8VFzXic" outputId="1fd74132-6a84-4c61-cc93-5b8bc010f6a1"
model.summary()
# + id="KfRxMPmU0lW1"
model.compile(optimizer='adam',loss='binary_crossentropy',metrics=['accuracy'])
# + executionInfo={"elapsed": 285550, "status": "ok", "timestamp": 1579525394894, "user": {"displayName": "<NAME>", "photoUrl": "https://lh6.googleusercontent.com/-t7IqSC7tKDY/AAAAAAAAAAI/AAAAAAAAhPo/INNzCzvCptg/s64/photo.jpg", "userId": "04862212736858688671"}, "user_tz": -330} id="KcD4VgAe0xgB" outputId="13a3c68a-d657-4ac3-8942-a377840d4cd2"
history = model.fit(X_train,y_train,epochs=5,validation_data=(X_test,y_test))
# + id="Uki_4QlT09Hx"
def plotLearningCurve(history,epochs):
epochRange = range(1,epochs+1)
plt.plot(epochRange,history.history['accuracy'])
plt.plot(epochRange,history.history['val_accuracy'])
plt.title('Model Accuracy')
plt.xlabel('Epoch')
plt.ylabel('Accuracy')
plt.legend(['Train','Validation'],loc='best')
plt.show()
plt.plot(epochRange,history.history['loss'])
plt.plot(epochRange,history.history['val_loss'])
plt.title('Model Loss')
plt.xlabel('Epoch')
plt.ylabel('Loss')
plt.legend(['Train','Validation'],loc='best')
plt.show()
# + executionInfo={"elapsed": 285525, "status": "ok", "timestamp": 1579525394896, "user": {"displayName": "<NAME>", "photoUrl": "https://lh6.googleusercontent.com/-t7IqSC7tKDY/AAAAAAAAAAI/AAAAAAAAhPo/INNzCzvCptg/s64/photo.jpg", "userId": "04862212736858688671"}, "user_tz": -330} id="rw2kGq-z34kS" outputId="2a3867d9-7167-4c4d-f375-2688e900459d"
plotLearningCurve(history,5)
# -
# # Applying the trained model to Predict the Genre of the input image
# + executionInfo={"elapsed": 1680, "status": "ok", "timestamp": 1579525492044, "user": {"displayName": "<NAME>", "photoUrl": "https://lh6.googleusercontent.com/-t7IqSC7tKDY/AAAAAAAAAAI/AAAAAAAAhPo/INNzCzvCptg/s64/photo.jpg", "userId": "04862212736858688671"}, "user_tz": -330} id="ngsqISoW37h4" outputId="efe04dc8-645c-43fe-d466-3d1d838867e3"
img = image.load_img('input/Images/tt0088247.jpg',target_size=(width,height,3))
plt.imshow(img)
img = image.img_to_array(img)
img = img/255.0
img = img.reshape(1,width,height,3)
classes = df.columns[2:]
y_pred = model.predict(img)
top3=np.argsort(y_pred[0])[:-4:-1]
for i in range(3):
print(classes[top3[i]])
# -
# # As you can see from the above output Model predicted the "Drama", "Thriller" and "Action" genre of the film from the poster of the image.
|
Genre Prediction from the Movie Poster/.ipynb_checkpoints/Genre_prediction_poster-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/jacobpad/DS-Unit-2-Linear-Models/blob/master/module2-regression-2/LS_DS12L_212.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="XmOQQWETlITN"
# Lambda School Data Science
#
# *Unit 2, Sprint 1, Module 2*
#
# ---
# + [markdown] colab_type="text" id="zmFCg6UArzn6"
# # Regression 2
# - Do train/test split
# - Use scikit-learn to fit a multiple regression
# - Understand how ordinary least squares regression minimizes the sum of squared errors
# - Define overfitting/underfitting and the bias/variance tradeoff
# + [markdown] colab_type="text" id="TLcAHk5arzn8"
# ### Setup
#
# Run the code cell below. You can work locally (follow the [local setup instructions](https://lambdaschool.github.io/ds/unit2/local/)) or on Colab.
#
# Libraries:
# - matplotlib
# - numpy
# - pandas
# - plotly
# - scikit-learn
# + colab_type="code" id="gtJg1IJZrzn9" colab={}
import sys
# If you're on Colab:
if 'google.colab' in sys.modules:
DATA_PATH = 'https://raw.githubusercontent.com/LambdaSchool/DS-Unit-2-Applied-Modeling/master/data/'
# If you're working locally:
else:
DATA_PATH = '../data/'
# Ignore this Numpy warning when using Plotly Express:
# FutureWarning: Method .ptp is deprecated and will be removed in a future version. Use numpy.ptp instead.
import warnings
warnings.filterwarnings(action='ignore', category=FutureWarning, module='numpy')
# + [markdown] colab_type="text" id="0LxiLwQC5age"
# # Do train/test split
# + [markdown] colab_type="text" id="ZLte4DsmmZgC"
# ## Overview
# + [markdown] colab_type="text" id="V2ui5w4C5agf"
# ### Predict Elections! 🇺🇸🗳️
# + [markdown] colab_type="text" id="7bf0OR0J5agf"
# How could we try to predict the 2020 US Presidential election?
#
# According to Douglas Hibbs, a political science and economics professor, you can [explain elections with just two features, "Bread and Peace":](https://douglas-hibbs.com/background-information-on-bread-and-peace-voting-in-us-presidential-elections/)
#
# > Aggregate two-party vote shares going to candidates of the party holding the presidency during the postwar era are well explained by just two fundamental determinants:
# >
# > (1) Positively by weighted-average growth of per capita real disposable personal income over the term.
# > (2) Negatively by cumulative US military fatalities (scaled to population) owing to unprovoked, hostile deployments of American armed forces in foreign wars.
# + [markdown] colab_type="text" id="xNDqNff_0N6m"
# Let's look at the data that Hibbs collected and analyzed:
# + colab_type="code" id="VRh-FSMTrzoG" outputId="d5dd14d5-e7fa-4590-cad5-c7d40efb3c5e" colab={"base_uri": "https://localhost:8080/", "height": 587}
import pandas as pd
df = pd.read_csv(DATA_PATH+'elections/bread_peace_voting.csv')
df
# + [markdown] colab_type="text" id="PggiQ7pm5agj"
# Data Sources & Definitions
#
# - 1952-2012: Douglas Hibbs, [2014 lecture at Deakin University Melbourne](http://www.douglas-hibbs.com/HibbsArticles/HIBBS-PRESVOTE-SLIDES-MELBOURNE-Part1-2014-02-26.pdf), Slide 40
# - 2016, Vote Share: [The American Presidency Project](https://www.presidency.ucsb.edu/statistics/elections)
# - 2016, Recent Growth in Personal Incomes: [The 2016 election economy: the "Bread and Peace" model final forecast](https://angrybearblog.com/2016/11/the-2016-election-economy-the-bread-and-peace-model-final-forecast.html)
# - 2016, US Military Fatalities: Assumption that Afghanistan War fatalities in 2012-16 occured at the same rate as 2008-12
#
# > Fatalities denotes the cumulative number of American military fatalities per millions of US population the in Korea, Vietnam, Iraq and Afghanistan wars during the presidential terms preceding the 1952, 1964, 1968, 1976 and 2004, 2008 and 2012 elections. —[Hibbs](http://www.douglas-hibbs.com/HibbsArticles/HIBBS-PRESVOTE-SLIDES-MELBOURNE-Part1-2014-02-26.pdf), Slide 33
# + [markdown] colab_type="text" id="ULwtMf5s5agj"
# Here we have data from the 1952-2016 elections. We could make a model to predict 1952-2016 election outcomes — but do we really care about that?
#
# No, not really. We already know what happened, we don't need to predict it.
# + [markdown] colab_type="text" id="Q1ADKjNU5agk"
# This is explained in [_An Introduction to Statistical Learning_](http://faculty.marshall.usc.edu/gareth-james/ISL/), Chapter 2.2, Assessing Model Accuracy:
#
# > In general, we do not really care how well the method works training on the training data. Rather, _we are interested in the accuracy of the predictions that we obtain when we apply our method to previously unseen test data._ Why is this what we care about?
# >
# > Suppose that we are interested in developing an algorithm to predict a stock’s price based on previous stock returns. We can train the method using stock returns from the past 6 months. But we don’t really care how well our method predicts last week’s stock price. We instead care about how well it will predict tomorrow’s price or next month’s price.
# >
# > On a similar note, suppose that we have clinical measurements (e.g. weight, blood pressure, height, age, family history of disease) for a number of patients, as well as information about whether each patient has diabetes. We can use these patients to train a statistical learning method to predict risk of diabetes based on clinical measurements. In practice, we want this method to accurately predict diabetes risk for _future patients_ based on their clinical measurements. We are not very interested in whether or not the method accurately predicts diabetes risk for patients used to train the model, since we already know which of those patients have diabetes.
# + [markdown] colab_type="text" id="9203LqC05agk"
# So, we're really interested in the 2020 election — but we probably don't want to wait until then to evaluate our model.
#
# There is a way we can estimate now how well our model will generalize in the future. We can't fast-forward time, but we can rewind it...
#
# We can split our data in **two sets.** For example:
# 1. **Train** a model on elections before 2008.
# 2. **Test** the model on 2008, 2012, 2016.
#
# This "backtesting" helps us estimate how well the model will predict the next elections going forward, starting in 2020.
# + [markdown] colab_type="text" id="4v28ozcC5agl"
# This is explained in [_Forecasting,_ Chapter 3.4,](https://otexts.com/fpp2/accuracy.html) Evaluating forecast accuracy:
#
# > The accuracy of forecasts can only be determined by considering how well a model performs on new data that were not used when fitting the model.
# >
# >When choosing models, it is common practice to separate the available data into two portions, training and test data, where the training data is used to estimate any parameters of a forecasting method and the test data is used to evaluate its accuracy. Because the test data is not used in determining the forecasts, it should provide a reliable indication of how well the model is likely to forecast on new data.
# >
# >
# >
# >The size of the test set is typically about 20% of the total sample, although this value depends on how long the sample is and how far ahead you want to forecast. The following points should be noted.
# >
# >- A model which fits the training data well will not necessarily forecast well.
# >- A perfect fit can always be obtained by using a model with enough parameters.
# >- Over-fitting a model to data is just as bad as failing to identify a systematic pattern in the data.
# >
# >Some references describe the test set as the “hold-out set” because these data are “held out” of the data used for fitting. Other references call the training set the “in-sample data” and the test set the “out-of-sample data”. We prefer to use “training data” and “test data” in this book.
# + [markdown] id="suRHvjK8MYRE" colab_type="text"
# **How should we split: Randomly? Before/after a given date?**
#
# I recommend you all read a great blog post, [How (and why) to create a good validation set](https://www.fast.ai/2017/11/13/validation-sets/), by fast.ai cofounder <NAME>.
#
# She gives great examples to answer the question “When is a random subset not good enough?” I’m not as opposed to random splits as R<NAME> seems to be. But it’s worth thinking about the trade-offs!
#
# Time-based and random splits can both be useful, and you’ll get repeated hands-on practice with both during this unit! (She also talks about the distinction between validation & test sets, which we’ll introduce in the last lesson of this Sprint.)
# + [markdown] colab_type="text" id="N01ZcGp75agl"
# ## Follow Along
#
# Split the data in two sets:
# 1. Train on elections before 2008.
# 2. Test on 2008 and after.
# + colab_type="code" id="3ymL6tTE5agm" colab={}
train = df[df['Year'] < 2008]
# + id="mYnW52LkReKh" colab_type="code" colab={}
test = df[df['Year'] >= 2008]
# + [markdown] colab_type="text" id="AYzA8Snk5ago"
# How many observations (rows) are in the train set? In the test set?
# + colab_type="code" id="IwsFNRRl5ago" outputId="9d96ae26-8505-40ab-915a-5821414d5801" colab={"base_uri": "https://localhost:8080/", "height": 35}
train.shape, test.shape
# + [markdown] colab_type="text" id="WgBB3mUS5agq"
# Note that this volume of data is at least two orders of magnitude smaller than we usually want to work with for predictive modeling.
#
# There are other validation techniques that could be used here, such as [time series cross-validation](https://scikit-learn.org/stable/modules/cross_validation.html#time-series-split), or [leave-one-out cross validation](https://scikit-learn.org/stable/modules/cross_validation.html#leave-one-out-loo) for small datasets. However, for this module, let's start simpler, with train/test split.
#
# Using a tiny dataset is intentional here. It's good for learning because we can see all the data at once.
# + [markdown] colab_type="text" id="FX3C2Fef5agr"
# ## Challenge
#
# In your assignment, you will do train/test split, based on date.
# + [markdown] colab_type="text" id="0jqNcyDg5agr"
# # Use scikit-learn to fit a multiple regression
# + [markdown] colab_type="text" id="BzEcjXpp5ags"
# ## Overview
#
# We've done train/test split, and we're ready to fit a model.
#
# We'll proceed in 3 steps. The first 2 are review from the previous module. The 3rd is new.
#
# - Begin with baselines (0 features)
# - Simple regression (1 feature)
# - Multiple regression (2 features)
# + [markdown] colab_type="text" id="9ChZZMtu5qMU"
# ## Follow Along
# + [markdown] colab_type="text" id="6mYp_0Aq5ags"
# ### Begin with baselines (0 features)
# + [markdown] colab_type="text" id="d_0nPUcQ0RpB"
# What was the average Incumbent Party Vote Share, in the 1952-2004 elections?
# + colab_type="code" id="mjm2CBV1ty35" outputId="71f49e93-1b68-4f55-c000-26b162046d69" colab={"base_uri": "https://localhost:8080/", "height": 35}
train['Incumbent Party Vote Share'].mean()
# + [markdown] colab_type="text" id="2aYkCqM_5agv"
# What if we guessed this number for every election? How far off would this be on average?
# + id="lCwUsXJnMYRU" colab_type="code" colab={}
# Arrange y target vectors
target = 'Incumbent Party Vote Share'
y_train = train[target]
y_test = test[target]
# + id="qUQYL2jQMYRW" colab_type="code" outputId="4a4ee8c8-2d33-4090-aa44-45b8d0674587" colab={"base_uri": "https://localhost:8080/", "height": 52}
# Get mean baseline
print('Mean Baseline (using 0 features)')
guess = y_train.mean()
print(guess)
# + id="JaijmBAXMYRY" colab_type="code" outputId="9dc8cf56-fa6b-4106-98af-18578d7126a5" colab={"base_uri": "https://localhost:8080/", "height": 35}
# Train Error
from sklearn.metrics import mean_absolute_error
y_pred = [guess] * len(y_train)
mae = mean_absolute_error(y_train, y_pred)
print(f'Train Error (1952-2004 elections): {mae:.2f} percentage points')
# + colab_type="code" id="3ma2uzUD5agv" outputId="206b0bfa-4c92-4d16-a530-5a53f0bdd1b1" colab={"base_uri": "https://localhost:8080/", "height": 35}
# Test Error
y_pred = [guess] * len(y_test)
mae = mean_absolute_error(y_test, y_pred)
print(f'Test Error (2008-16 elections): {mae:.2f} percentage points')
# + [markdown] colab_type="text" id="QlAAlCA_5agx"
# ### Simple regression (1 feature)
# + [markdown] colab_type="text" id="RaQj9J7P5agy"
# Make a scatterplot of the relationship between 1 feature and the target.
#
# We'll use an economic feature: Average Recent Growth in Personal Incomes. ("Bread")
# + colab_type="code" id="88JNPm46rzoD" outputId="f981115e-c9d9-4c40-fd88-32d83d9d5ec2" colab={"base_uri": "https://localhost:8080/", "height": 542}
import pandas as pd
import plotly.express as px
px.scatter(
train,
x='Average Recent Growth in Personal Incomes',
y='Incumbent Party Vote Share',
text='Year',
title='US Presidential Elections, 1952-2004',
trendline='ols', # Ordinary Least Squares
)
# + [markdown] colab_type="text" id="XJbyyFBj5ag1"
# 1952 & 1968 are outliers: The incumbent party got fewer votes than predicted by the regression. What do you think could explain those years? We'll come back to this soon, but first...
# + [markdown] colab_type="text" id="JTaTwOCN5ag2"
# Use scikit-learn to fit the simple regression with one feature.
#
# Follow the [5 step process](https://jakevdp.github.io/PythonDataScienceHandbook/05.02-introducing-scikit-learn.html#Basics-of-the-API), and refer to [Scikit-Learn LinearRegression documentation](https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.LinearRegression.html).
# + id="urIHJoCTMYRi" colab_type="code" colab={}
# 1. Import the appropriate estimator class from Scikit-Learn
from sklearn.linear_model import LinearRegression
# + id="OOI24cZKMYRl" colab_type="code" colab={}
# 2. Instantiate this class
model = LinearRegression()
# + id="DoAPSq_3MYRo" colab_type="code" outputId="b2393844-4bf1-412d-804b-71a468bc5562" colab={"base_uri": "https://localhost:8080/", "height": 35}
# 3. Arrange X features matrices (already did y target vectors)
features = ['Average Recent Growth in Personal Incomes']
X_train = train[features]
X_test = test[features]
print(f'Linear Regression, dependent on: {features}')
# + id="np_C9tGWMYRq" colab_type="code" outputId="c28c1df7-2caf-4479-f5ee-7a1a601b5434" colab={"base_uri": "https://localhost:8080/", "height": 35}
# 4. Fit the model
model.fit(X_train, y_train)
y_pred = model.predict(X_train)
mae = mean_absolute_error(y_train, y_pred)
print(f'Train Error: {mae:.2f} percentage points')
# + colab_type="code" id="1AD_jJHm5ag2" outputId="44dd7f07-34e6-47fa-9640-e6b98dfb3ab3" colab={"base_uri": "https://localhost:8080/", "height": 35}
# 5. Apply the model to new data
y_pred = model.predict(X_test)
mae = mean_absolute_error(y_test, y_pred)
print(f'Test Error: {mae:.2f} percentage points')
# + [markdown] colab_type="text" id="pMW4wSmC5ag4"
# How does the error compare to the baseline?
# + [markdown] colab_type="text" id="8PBgsYaa5ag4"
# ### Multiple regression (2 features)
# + [markdown] colab_type="text" id="H9fEbVTE0ZMe"
# Make a scatterplot of the relationship between 2 features and the target.
#
# We'll add another feature: US Military Fatalities per Million. ("Peace" or the lack thereof.)
#
# Rotate the scatterplot to explore the data. What's different about 1952 & 1968?
# + colab_type="code" id="XWNNK5r-t_pF" outputId="7ab00772-30ef-4e22-a1f3-581329d75d7f" colab={"base_uri": "https://localhost:8080/", "height": 542}
px.scatter_3d(
train,
x='Average Recent Growth in Personal Incomes',
y='US Military Fatalities per Million',
z='Incumbent Party Vote Share',
text='Year',
title='US Presidential Elections, 1952-2004'
)
# + [markdown] colab_type="text" id="MNoTuf9v7l6A"
# Use scikit-learn to fit a multiple regression with two features.
# + id="VvbvcvzVMYR0" colab_type="code" outputId="1f9d1384-20d6-4c07-cb7d-f095f6ba017c" colab={"base_uri": "https://localhost:8080/", "height": 35}
# TODO: Complete this cell
# Re-arrange X features matrices
features = ['Average Recent Growth in Personal Incomes',
'US Military Fatalities per Million']
print(f'Linear Regression, dependent on: {features}')
X_train = train[features]
X_test = test[features]
# + id="5CcrvO7tMYR2" colab_type="code" outputId="8c1e4f2e-a630-4a3e-bc87-de40c7913cb9" colab={"base_uri": "https://localhost:8080/", "height": 35}
# TODO: Fit the model
model.fit(X_train, y_train)
# + id="NoRjz3oBZwx-" colab_type="code" colab={}
y_pred = model.predict(X_train)
# + id="5uqVYVrEZ-pS" colab_type="code" outputId="19dcdcee-5251-47a4-abe9-ab758474150e" colab={"base_uri": "https://localhost:8080/", "height": 35}
# Calculating Training Mean Absolute Error manually,
# just for demonstration purposes
(y_pred - y_train).abs().mean()
# + id="wfE1dP4IaafW" colab_type="code" outputId="a64fec29-4e38-4c90-a233-9d20a16a2619" colab={"base_uri": "https://localhost:8080/", "height": 35}
mae = mean_absolute_error(y_train, y_pred)
print(f'Train Error: {mae:.2f} percentage points')
# + colab_type="code" id="_Mijdiua5ag8" outputId="6f9961c6-5672-<PASSWORD>" colab={"base_uri": "https://localhost:8080/", "height": 35}
# TODO: Apply the model to new data
y_pred = model.predict(X_test)
mae = mean_absolute_error(y_test, y_pred)
print(f'Test Error: {mae:.2f} percentage points')
# + [markdown] colab_type="text" id="9zKDY_CJ5ag-"
# How does the error compare to the prior model?
# + [markdown] colab_type="text" id="qZq7sRk45ag_"
# ### Plot the plane of best fit
# + [markdown] colab_type="text" id="t27N85U85ahA"
# For a regression with 1 feature, we plotted the line of best fit in 2D.
#
# (There are many ways to do this. Plotly Express's `scatter` function makes it convenient with its `trendline='ols'` parameter.)
#
# For a regression with 2 features, we can plot the plane of best fit in 3D!
#
# (Plotly Express has a `scatter_3d` function but it won't plot the plane of best fit for us. But, we can write our own function, with the same "function signature" as the Plotly Express API.)
# + colab_type="code" id="MQ73YaIF0gJC" colab={}
import itertools
import numpy as np
import plotly.express as px
import plotly.graph_objs as go
from sklearn.linear_model import LinearRegression
def regression_3d(df, x, y, z, num=100, **kwargs):
"""
Visualize linear regression in 3D: 2 features + 1 target
df : Pandas DataFrame
x : string, feature 1 column in df
y : string, feature 2 column in df
z : string, target column in df
num : integer, number of quantiles for each feature
"""
# Plot data
fig = px.scatter_3d(df, x, y, z, **kwargs)
# Fit Linear Regression
features = [x, y]
target = z
model = LinearRegression()
model.fit(df[features], df[target])
# Define grid of coordinates in the feature space
xmin, xmax = df[x].min(), df[x].max()
ymin, ymax = df[y].min(), df[y].max()
xcoords = np.linspace(xmin, xmax, num)
ycoords = np.linspace(ymin, ymax, num)
coords = list(itertools.product(xcoords, ycoords))
# Make predictions for the grid
predictions = model.predict(coords)
Z = predictions.reshape(num, num).T
# Plot predictions as a 3D surface (plane)
fig.add_trace(go.Surface(x=xcoords, y=ycoords, z=Z))
return fig
# + colab_type="code" id="f2TMPMM4u_5I" outputId="270b0072-58a0-4de6-a6ca-a95e463acc2a" colab={"base_uri": "https://localhost:8080/", "height": 542}
regression_3d(
train,
x='Average Recent Growth in Personal Incomes',
y='US Military Fatalities per Million',
z='Incumbent Party Vote Share',
text='Year',
title='US Presidential Elections, 1952-2004'
)
# + [markdown] colab_type="text" id="GXOl-Aut5ahE"
# Where are 1952 & 1968 in relation to the plane? Which elections are the biggest outliers now?
# + [markdown] colab_type="text" id="vHAHyrNC5ahE"
# Roll over points on the plane to see predicted incumbent party vote share (z axis), dependent on personal income growth (x axis) and military fatatlies per capita (y axis).
# + [markdown] colab_type="text" id="SJ490-qEvfSr"
# ### Get and interpret coefficients
# + [markdown] colab_type="text" id="8YR1p7eexIf6"
# During the previous module, we got the simple regression's coefficient and intercept. We plugged these numbers into an equation for the line of best fit, in slope-intercept form: $y = mx + b$
#
# Let's review this objective, but now for multiple regression.
#
# What's the equation for the plane of best fit?
#
# $y = \beta_0 + \beta_1x_1 + \beta_2x_2$
#
# Can you relate the intercept and coefficients to what you see in the plot above?
# + colab_type="code" id="lFhJqqSS-i9d" outputId="6988b49b-83a2-4087-a13a-53c2ce00a4e5" colab={"base_uri": "https://localhost:8080/", "height": 35}
model.intercept_, model.coef_
# + colab_type="code" id="dxXoa0FW5ahH" outputId="465f9cac-ebe9-4d49-c00a-044a0b79e010" colab={"base_uri": "https://localhost:8080/", "height": 35}
beta0 = model.intercept_
beta1, beta2 = model.coef_
print(f'y = {beta0} + {beta1}x1 + {beta2}x2')
# + colab_type="code" id="_DivQmKEDcit" outputId="dea7923b-2399-4880-b107-b0823f9329b2" colab={"base_uri": "https://localhost:8080/", "height": 69}
# This is easier to read
print('Intercept', model.intercept_)
coefficients = pd.Series(model.coef_, features)
print(coefficients.to_string())
# + [markdown] colab_type="text" id="Vf8s9srD5ahM"
# One of the coefficients is positive, and the other is negative. What does this mean?
# + [markdown] id="67YlhFOJMYSL" colab_type="text"
# Let's look at some scenarios. We'll see that one unit's change in an independent variable results in a coefficient worth of change in the dependent variable.
# + [markdown] colab_type="text" id="TAQjaiHoxR-7"
# What does the model predict if income growth=0%, fatalities=0
# + colab_type="code" id="Rup9q-6gAIZD" outputId="9d3c319c-9d4d-4dd3-d81d-8c0a5195549b" colab={"base_uri": "https://localhost:8080/", "height": 35}
model.predict([[0, 0]])
# + [markdown] colab_type="text" id="Itt56qYNxYIb"
# Income growth = 1% (fatalities = 0)
# + colab_type="code" id="1onFls9AAh5Z" outputId="c460e9dc-7aee-4534-975f-6807f09af910" colab={"base_uri": "https://localhost:8080/", "height": 35}
model.predict([[1, 0]])
# + [markdown] colab_type="text" id="V5ZMUDjGxdhe"
# The difference between these predictions = ?
# + colab_type="code" id="el1gAUyGApnV" outputId="3855a808-377d-44a9-dcba-964431f6e689" colab={"base_uri": "https://localhost:8080/", "height": 35}
model.predict([[1, 0]]) - model.predict([[0, 0]])
# + [markdown] colab_type="text" id="1U1o9L65xn6_"
# What if... income growth = 2% (fatalities = 0)
# + colab_type="code" id="stfnvUc_A3pM" outputId="1f2ba70c-6289-4eac-9cd4-df6ea7a8b482" colab={"base_uri": "https://localhost:8080/", "height": 35}
model.predict([[2, 0]])
# + [markdown] colab_type="text" id="bjp2kDm5xq79"
# The difference between these predictions = ?
# + colab_type="code" id="Zywdu1SJA_hP" outputId="e5e41673-5d81-4210-93d6-61fd2f67a144" colab={"base_uri": "https://localhost:8080/", "height": 35}
model.predict([[2, 0]]) - model.predict([[1, 0]])
# + [markdown] colab_type="text" id="klON2yUxxu1_"
# What if... (income growth=2%) fatalities = 100
# + colab_type="code" id="YTFqg0ixBcIH" outputId="2eb8d180-3315-4287-b4f4-1810a8234214" colab={"base_uri": "https://localhost:8080/", "height": 35}
model.predict([[2, 100]])
# + [markdown] colab_type="text" id="AfqzTR6dxyQ-"
# The difference between these predictions = ?
# + colab_type="code" id="VHtDzQT-Bxel" outputId="9aa6c71b-e984-44c4-e50d-82c952dda590" colab={"base_uri": "https://localhost:8080/", "height": 35}
model.predict([[2, 100]]) - model.predict([[2, 0]])
# + [markdown] colab_type="text" id="6-3Whu9gx2ac"
# What if income growth = 3% (fatalities = 100)
# + colab_type="code" id="2ge-czWaCAWQ" outputId="7fcc3ec5-ad33-447a-8408-dfdfd8d917e4" colab={"base_uri": "https://localhost:8080/", "height": 35}
model.predict([[3, 100]])
# + [markdown] colab_type="text" id="x7wDd61ax6Kk"
# The difference between these predictions = ?
# + colab_type="code" id="ZDWbZ1duCJLS" outputId="18be46c3-b28a-486d-b5d9-068a79272509" colab={"base_uri": "https://localhost:8080/", "height": 35}
model.predict([[3, 100]]) - model.predict([[2, 100]])
# + [markdown] colab_type="text" id="WXaeJe9px9d1"
# What if (income growth = 3%) fatalities = 200
# + colab_type="code" id="MP6UnCT1CXoD" outputId="1c7fdea2-c76c-46cd-9545-dd0b998b7011" colab={"base_uri": "https://localhost:8080/", "height": 35}
model.predict([[3, 200]])
# + [markdown] colab_type="text" id="MuioZsRUyAud"
# The difference between these predictions = ?
# + colab_type="code" id="OpiDm-QjCZqi" outputId="f508e5aa-aa50-4915-861a-7805947f623a" colab={"base_uri": "https://localhost:8080/", "height": 35}
model.predict([[3, 200]]) - model.predict([[3, 100]])
# + [markdown] colab_type="text" id="-h8Tl7HOwBM7"
# ## Challenge
#
# In your assignment, you'll fit a Linear Regression with at least 2 features.
# + [markdown] colab_type="text" id="o1_b4j6ZtZgb"
# # Understand how ordinary least squares regression minimizes the sum of squared errors
# + [markdown] colab_type="text" id="D8moJTC6uDuh"
# ## Overview
#
# So far, we've evaluated our models by their absolute error. It's an intuitive metric for regression problems.
#
# However, ordinary least squares doesn't directly minimize absolute error. Instead, it minimizes squared error.
#
#
#
# + [markdown] colab_type="text" id="ARY8zz7J6EtO"
# In this section, we'll introduce two new regression metrics:
#
# - Squared error
# - $R^2$
#
# + [markdown] colab_type="text" id="cwqJlR1M6Crw"
# We'll demostrate two possible methods to minimize squared error:
#
# - Guess & check
# - Linear Algebra
# + [markdown] colab_type="text" id="zspnHmo7wbKF"
# ## Follow Along
# + [markdown] colab_type="text" id="ebHzgmJfG80i"
# ### Guess & Check
#
# This function visualizes squared errors. We'll go back to simple regression with 1 feature, because it's much easier to visualize.
#
# Use the function's m & b parameters to "fit the model" manually. Guess & check what values of m & b minimize squared error.
# + colab_type="code" id="BmdIG_W_8tTE" colab={}
from matplotlib.patches import Rectangle
import matplotlib.pyplot as plt
import numpy as np
from sklearn.metrics import mean_absolute_error, mean_squared_error, r2_score
def squared_errors(df, feature, target, m, b):
"""
Visualize linear regression, with squared errors,
in 2D: 1 feature + 1 target.
Use the m & b parameters to "fit the model" manually.
df : Pandas DataFrame
feature : string, feature column in df
target : string, target column in df
m : numeric, slope for linear equation
b : numeric, intercept for linear requation
"""
# Plot data
fig = plt.figure(figsize=(7,7))
ax = plt.axes()
df.plot.scatter(feature, target, ax=ax)
# Make predictions
x = df[feature]
y = df[target]
y_pred = m*x + b
# Plot predictions
ax.plot(x, y_pred)
# Plot squared errors
xmin, xmax = ax.get_xlim()
ymin, ymax = ax.get_ylim()
scale = (xmax-xmin)/(ymax-ymin)
for x, y1, y2 in zip(x, y, y_pred):
bottom_left = (x, min(y1, y2))
height = abs(y1 - y2)
width = height * scale
ax.add_patch(Rectangle(xy=bottom_left, width=width, height=height, alpha=0.1))
# Print regression metrics
mse = mean_squared_error(y, y_pred)
rmse = np.sqrt(mse)
mae = mean_absolute_error(y, y_pred)
r2 = r2_score(y, y_pred)
print('Mean Squared Error:', mse)
print('Root Mean Squared Error:', rmse)
print('Mean Absolute Error:', mae)
print('R^2:', r2)
# + [markdown] colab_type="text" id="x0313AWF5ahp"
# Here's what the mean baseline looks like:
# + colab_type="code" id="8bsq0zoE5ahq" outputId="7f5f8ec9-299c-4bdb-ba35-adcdb3ff4771" colab={"base_uri": "https://localhost:8080/", "height": 513}
feature = 'Average Recent Growth in Personal Incomes'
squared_errors(train, feature, target, m=0, b=y_train.mean())
# + [markdown] colab_type="text" id="5RVz4qYR5ahr"
# Notice that $R^2$ is exactly zero.
#
# [$R^2$ represents the proportion of the variance for a dependent variable that is explained by the independent variable(s).](https://en.wikipedia.org/wiki/Coefficient_of_determination)
#
# The mean baseline uses zero independent variables and explains none of the variance in the dependent variable, so its $R^2$ score is zero.
#
# The highest possible $R^2$ score is 1. The lowest possible *Train* $R^2$ score with ordinary least squares regression is 0.
#
# In this demo, it's possible to get a negative Train $R^2$, if you manually set values of m & b that are worse than the mean baseline. But that wouldn't happen in the real world.
#
# However, in the real world, it _is_ possible to get a negative *Test/Validation* $R^2$. It means that your *Test/Validation* predictions are worse than if you'd constantly predicted the mean of the *Test/Validation* set.
# + [markdown] colab_type="text" id="KaJZqg1r5ahs"
# ---
#
# Now that we've visualized the squared errors for the mean baseline, let's guess & check some better values for the m & b parameters:
# + colab_type="code" id="TkyDSs8f6stD" outputId="f168660e-3d79-4bc6-a643-3814b2180526" colab={"base_uri": "https://localhost:8080/", "height": 513}
squared_errors(train, feature, target, m=3, b=46)
# + [markdown] colab_type="text" id="oxp-rvWC5aht"
# You can run the function repeatedly, with different values for m & b.
#
# How do you interpret each metric you see?
#
# - Mean Squared Error
# - Root Mean Squared Error
# - Mean Absolute Error
# - $R^2$
#
# Does guess & check really get used in machine learning? Sometimes! Some complex functions are hard to minimize, so we use a sophisticated form of guess & check called "gradient descent", which you'll learn about in Unit 4.
#
# (Also, GridSearchCV and RandomizedSearchCV for hyperparameter optimization are a form of guess & check, which you'll learn about later this unit.)
#
# Fortunately, we don't need to use guess & check for ordinary least squares regression. We have a solution, using linear algebra!
#
# + [markdown] colab_type="text" id="u6biDIhlrzoI"
# ### Linear Algebra
#
# The same result that is found by minimizing the sum of the squared errors can be also found through a linear algebra process known as the "Least Squares Solution:"
#
# \begin{align}
# \hat{\beta} = (X^{T}X)^{-1}X^{T}y
# \end{align}
#
# Before we can work with this equation in its linear algebra form we have to understand how to set up the matrices that are involved in this equation.
#
# #### The $\beta$ vector
#
# The $\beta$ vector represents all the parameters that we are trying to estimate, our $y$ vector and $X$ matrix values are full of data from our dataset. The $\beta$ vector holds the variables that we are solving for: $\beta_0$ and $\beta_1$
#
# Now that we have all of the necessary parts we can set them up in the following equation:
#
# \begin{align}
# y = X \beta + \epsilon
# \end{align}
#
# Since our $\epsilon$ value represents **random** error we can assume that it will equal zero on average.
#
# \begin{align}
# y = X \beta
# \end{align}
#
# The objective now is to isolate the $\beta$ matrix. We can do this by pre-multiplying both sides by "X transpose" $X^{T}$.
#
# \begin{align}
# X^{T}y = X^{T}X \beta
# \end{align}
#
# Since anything times its transpose will result in a square matrix, if that matrix is then an invertible matrix, then we should be able to multiply both sides by its inverse to remove it from the right hand side. (We'll talk tomorrow about situations that could lead to $X^{T}X$ not being invertible.)
#
# \begin{align}
# (X^{T}X)^{-1}X^{T}y = (X^{T}X)^{-1}X^{T}X \beta
# \end{align}
#
# Since any matrix multiplied by its inverse results in the identity matrix, and anything multiplied by the identity matrix is itself, we are left with only $\beta$ on the right hand side:
#
# \begin{align}
# (X^{T}X)^{-1}X^{T}y = \hat{\beta}
# \end{align}
#
# We will now call it "beta hat" $\hat{\beta}$ because it now represents our estimated values for $\beta_0$ and $\beta_1$
#
# #### Lets calculate our $\beta$ parameters with numpy!
# + colab_type="code" id="ouidNhDprzoI" outputId="4596ee10-2c18-4f94-ad57-01aeba10d8ab" colab={"base_uri": "https://localhost:8080/", "height": 607}
# This is NOT something you'll be tested on. It's just a demo.
# X is a matrix. Add column of constants for fitting the intercept.
def add_constant(X):
constant = np.ones(shape=(len(X),1))
return np.hstack((constant, X))
X = add_constant(train[features].values)
print('X')
print(X)
# y is a column vector
y = train[target].values[:, np.newaxis]
print('y')
print(y)
# Least squares solution in code
X_transpose = X.T
X_transpose_X = X_transpose @ X
X_transpose_X_inverse = np.linalg.inv(X_transpose_X)
X_transpose_y = X_transpose @ y
beta_hat = X_transpose_X_inverse @ X_transpose_y
print('Beta Hat')
print(beta_hat)
# + colab_type="code" id="jDTgv1lKrzoK" outputId="89578285-c6b4-4236-e0a7-4f03646f7957" colab={"base_uri": "https://localhost:8080/", "height": 35}
# Scikit-learn gave the exact same results!
model.intercept_, model.coef_
# + [markdown] colab_type="text" id="RaLUPbonrzoM"
# # Define overfitting/underfitting and the bias/variance tradeoff
# + [markdown] colab_type="text" id="W-40FQscwfD5"
# ## Overview
# + [markdown] colab_type="text" id="W9Aboy6MrzoR"
# Read [_Python Data Science Handbook,_ Chapter 5.3](https://jakevdp.github.io/PythonDataScienceHandbook/05.03-hyperparameters-and-model-validation.html#The-Bias-variance-trade-off). <NAME> explains overfitting & underfitting:
#
# > Fundamentally, the question of "the best model" is about finding a sweet spot in the tradeoff between bias and variance. Consider the following figure, which presents two regression fits to the same dataset:
# >
# >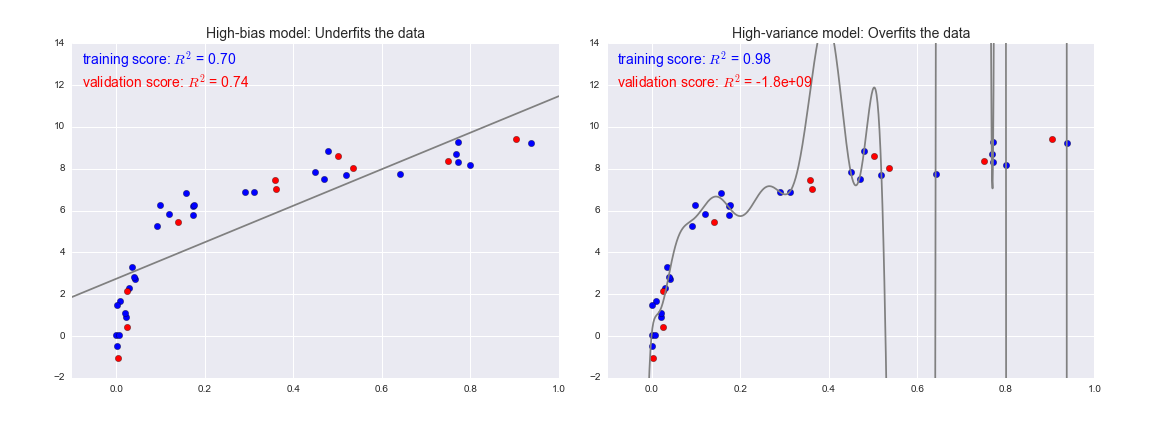
# >
# > The model on the left attempts to find a straight-line fit through the data. Because the data are intrinsically more complicated than a straight line, the straight-line model will never be able to describe this dataset well. Such a model is said to _underfit_ the data: that is, it does not have enough model flexibility to suitably account for all the features in the data; another way of saying this is that the model has high _bias_.
# >
# > The model on the right attempts to fit a high-order polynomial through the data. Here the model fit has enough flexibility to nearly perfectly account for the fine features in the data, but even though it very accurately describes the training data, its precise form seems to be more reflective of the particular noise properties of the data rather than the intrinsic properties of whatever process generated that data. Such a model is said to _overfit_ the data: that is, it has so much model flexibility that the model ends up accounting for random errors as well as the underlying data distribution; another way of saying this is that the model has high _variance_.
# + [markdown] colab_type="text" id="Q6smEDli6REF"
# VanderPlas goes on to connect these concepts to the "bias/variance tradeoff":
#
# > From the scores associated with these two models, we can make an observation that holds more generally:
# >
# >- For high-bias models, the performance of the model on the validation set is similar to the performance on the training set.
# >
# >- For high-variance models, the performance of the model on the validation set is far worse than the performance on the training set.
# >
# > If we imagine that we have some ability to tune the model complexity, we would expect the training score and validation score to behave as illustrated in the following figure:
# >
# >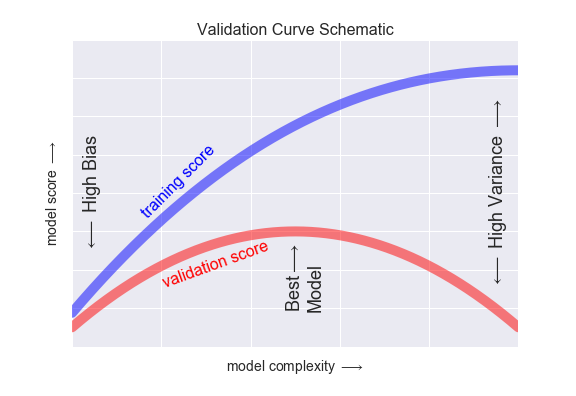
# >
# > The diagram shown here is often called a validation curve, and we see the following essential features:
# >
# >- The training score is everywhere higher than the validation score. This is generally the case: the model will be a better fit to data it has seen than to data it has not seen.
# >- For very low model complexity (a high-bias model), the training data is under-fit, which means that the model is a poor predictor both for the training data and for any previously unseen data.
# >- For very high model complexity (a high-variance model), the training data is over-fit, which means that the model predicts the training data very well, but fails for any previously unseen data.
# >- For some intermediate value, the validation curve has a maximum. This level of complexity indicates a suitable trade-off between bias and variance.
# >
# >The means of tuning the model complexity varies from model to model.
# + [markdown] colab_type="text" id="TWRkoX-75ahz"
# So far, our only "means of tuning the model complexity" has been selecting one feature or two features for our linear regression models. But we'll quickly start to select more features, and more complex models, with more "hyperparameters."
#
# This is just a first introduction to underfitting & overfitting. We'll continue to learn about this topic all throughout this unit.
# + [markdown] colab_type="text" id="bQ8rflrBwgYj"
# ## Follow Along
# + [markdown] colab_type="text" id="eeqQm4JM5ah0"
# Let's make our own Validation Curve, by tuning a new type of model complexity: polynomial degrees in a linear regression.
# + [markdown] colab_type="text" id="8ajsC5CS9nYI"
# Go back to the the NYC Tribeca condo sales data
# + colab_type="code" id="DzCSC1zsrzoN" colab={}
# Read NYC Tribeca condo sales data, from first 4 months of 2019.
# Dataset has 90 rows, 9 columns.
df = pd.read_csv(DATA_PATH+'condos/tribeca.csv')
assert df.shape == (90, 9)
# Arrange X features matrix & y target vector
features = ['GROSS_SQUARE_FEET']
target = 'SALE_PRICE'
X = df[features]
y = df[target]
# + [markdown] colab_type="text" id="VccY7fno9sA8"
# Do random [train/test split](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.train_test_split.html)
# + colab_type="code" id="ki0qmlk69uWr" colab={}
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=11)
# + [markdown] colab_type="text" id="HpQnoWJV9zkr"
# Repeatedly fit increasingly complex models, and keep track of the scores
# + colab_type="code" id="YShWjZrIrzoP" outputId="e8e43b54-72ae-40ba-a65a-90a6463394a9" colab={"base_uri": "https://localhost:8080/", "height": 1000}
from IPython.display import display, HTML
from sklearn.linear_model import LinearRegression
from sklearn.metrics import r2_score
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import PolynomialFeatures
# Credit for PolynomialRegression: <NAME>, Python Data Science Handbook, Chapter 5.3
# https://jakevdp.github.io/PythonDataScienceHandbook/05.03-hyperparameters-and-model-validation.html#Validation-curves-in-Scikit-Learn
def PolynomialRegression(degree=2, **kwargs):
return make_pipeline(PolynomialFeatures(degree),
LinearRegression(**kwargs))
polynomial_degrees = range(1, 10, 2)
train_r2s = []
test_r2s = []
for degree in polynomial_degrees:
model = PolynomialRegression(degree)
display(HTML(f'Polynomial degree={degree}'))
model.fit(X_train, y_train)
train_r2 = model.score(X_train, y_train)
test_r2 = model.score(X_test, y_test)
display(HTML(f'<b style="color: blue">Train R2 {train_r2:.2f}</b>'))
display(HTML(f'<b style="color: red">Test R2 {test_r2:.2f}</b>'))
plt.scatter(X_train, y_train, color='blue', alpha=0.5)
plt.scatter(X_test, y_test, color='red', alpha=0.5)
plt.xlabel(features)
plt.ylabel(target)
x_domain = np.linspace(X.min(), X.max())
curve = model.predict(x_domain)
plt.plot(x_domain, curve, color='blue')
plt.show()
display(HTML('<hr/>'))
train_r2s.append(train_r2)
test_r2s.append(test_r2)
display(HTML('Validation Curve'))
plt.plot(polynomial_degrees, train_r2s, color='blue', label='Train')
plt.plot(polynomial_degrees, test_r2s, color='red', label='Test')
plt.xlabel('Model Complexity (Polynomial Degree)')
plt.ylabel('R^2 Score')
plt.legend()
plt.show()
# + [markdown] colab_type="text" id="VxzxiDxz5ah4"
# As model complexity increases, what happens to Train $R^2$ and Test $R^2$?
# + [markdown] colab_type="text" id="bO298NwR5ah5"
# # Review
#
# In your assignment, you'll continue to **predict how much it costs to rent an apartment in NYC,** using the dataset from renthop.com.
#
#
# - Do train/test split. Use data from April & May 2016 to train. Use data from June 2016 to test.
# - Engineer at least two new features.
# - Fit a linear regression model with at least two features.
# - Get the model's coefficients and intercept.
# - Get regression metrics RMSE, MAE, and $R^2$, for both the train and test sets.
#
# You've been provided with a separate notebook for your assignment, which has all the instructions and stretch goals. What's the best test MAE you can get? Share your score and features used with your cohort on Slack!
# + [markdown] colab_type="text" id="4WcLE44XwpOL"
# # Sources
#
# #### Train/Test Split
# - <NAME>, Hastie, Tibshirani, [_An Introduction to Statistical Learning_](http://faculty.marshall.usc.edu/gareth-james/ISL/), Chapter 2.2, Assessing Model Accuracy
# - Hyndman, Athanasopoulos, [_Forecasting,_ Chapter 3.4,](https://otexts.com/fpp2/accuracy.html) Evaluating forecast accuracy
# - <NAME>, [How (and why) to create a good validation set](https://www.fast.ai/2017/11/13/validation-sets/)
#
# #### Bias-Variance Tradeoff
# - <NAME>, [_Python Data Science Handbook,_ Chapter 5.3](https://jakevdp.github.io/PythonDataScienceHandbook/05.03-hyperparameters-and-model-validation.html#The-Bias-variance-trade-off), Hyperparameters and Model Validation
# - StatQuest, [Machine Learning Fundamentals: Bias and Variance](https://youtu.be/EuBBz3bI-aA) (6.5 minutes)
#
# #### "Bread and Peace" Background
# - <NAME>, [Background Information on the ‘Bread and Peace’ Model of Voting in Postwar US Presidential Elections](https://douglas-hibbs.com/background-information-on-bread-and-peace-voting-in-us-presidential-elections/)
# - <NAME>, [What Do Economic Models Really Tell Us About Elections?](https://fivethirtyeight.com/features/what-do-economic-models-really-tell-us-about-elections/)
#
#
# #### "Bread and Peace" Data Sources & Definitions
# - 1952-2012: <NAME>, [2014 lecture at Deakin University Melbourne](http://www.douglas-hibbs.com/HibbsArticles/HIBBS-PRESVOTE-SLIDES-MELBOURNE-Part1-2014-02-26.pdf), Slide 40
# - 2016, Vote Share: [The American Presidency Project](https://www.presidency.ucsb.edu/statistics/elections)
# - 2016, Recent Growth in Personal Incomes: [The 2016 election economy: the "Bread and Peace" model final forecast](https://angrybearblog.com/2016/11/the-2016-election-economy-the-bread-and-peace-model-final-forecast.html)
# - 2016, US Military Fatalities: Assumption that Afghanistan War fatalities in 2012-16 occured at the same rate as 2008-12
#
# > Fatalities denotes the cumulative number of American military fatalities per millions of US population the in Korea, Vietnam, Iraq and Afghanistan wars during the presidential terms preceding the 1952, 1964, 1968, 1976 and 2004, 2008 and 2012 elections. —[Hibbs](http://www.douglas-hibbs.com/HibbsArticles/HIBBS-PRESVOTE-SLIDES-MELBOURNE-Part1-2014-02-26.pdf), Slide 33
|
module2-regression-2/LS_DS12L_212.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # APIs
#
# Instead of downloading World Bank data via a csv file, you're going to download the data using the World Bank APIs. The purpose of this exercise is to gain experience with another way of extracting data.
#
# API is an acronym that stands for application programming interface. API’s provide a standardized way for two applications to talk to each other. For this project, the applications communicating with each other are the server application where World Bank stores data and your Jupyter notebook.
#
# If you wanted to pull data directly from the World Bank’s server, you’d have to know what database system the World Bank was using. You’d also need permission to log in directly to the server, which would be a security risk for the World Bank. And if the World Bank ever migrated its data to a new system, you would have to rewrite all of your code again. The API allows you to execute code on the World Bank server without getting direct access.
#
# # Before there were APIs
#
# Before there were APIs, there was web scraping. People would download html directly from a website and then parse the results programatically. This practice is in a legal grey area. One reason that APIs became popular was so that companies could provide data to users and discourage web scraping.
#
# Here are a few articles about the legality of web scraping.
#
# * [QVC Can't Stop Web Scraping](https://www.forbes.com/sites/ericgoldman/2015/03/24/qvc-cant-stop-web-scraping/#120db59b3ca3)
# * [Quora - Legality of Web Scraping](https://www.quora.com/What-is-the-legality-of-web-scraping)
#
# All sorts of companies have public facing APIs including Facebook, Twitter, Google and Pinterest. You can pull data from these companies to create your own applications.
#
# In this notebook, you’ll get practice using Python to pull data from the World Bank indicators API.
#
# Here are links to information about the World Bank indicators and projects APIs if you want to learn more:
# * [World Bank Indicators API](world bank projects api)
# * [World Bank Projects API](http://search.worldbank.org/api/v2/projects)
# # Using APIs
#
# In general, you access APIs via the web using a web address. Within the web address, you specify the data that you want. To know how to format the web address, you need to read the documentation. Some APIs also require that you send login credentials as part of your request. The World Bank APIs are public and do not require login credentials.
#
# The Python requests library makes working with APIs relatively simple.
# # Example Indicators API
#
# Run the code example below to request data from the World Bank Indicators API. According to the documntation, you format your request url like so:
#
# `http://api.worldbank.org/v2/countries/` + list of country abbreviations separated by ; + `/indicators/` + indicator name + `?` + options
#
# where options can include
# * per_page - number of records to return per page
# * page - which page to return - eg if there are 5000 records and 100 records per page
# * date - filter by dates
# * format - json or xml
#
# and a few other options that you can read about [here](https://datahelpdesk.worldbank.org/knowledgebase/articles/898581-api-basic-call-structure).
# +
import requests
import pandas as pd
url = 'http://api.worldbank.org/v2/countries/br;cn;us;de/indicators/SP.POP.TOTL/?format=json&per_page=1000'
r = requests.get(url)
r.json()
# -
# This json data isn't quite ready for a pandas data frame. Notice that the json response is a list with two entries. The first entry is
# ```
# {'lastupdated': '2018-06-28',
# 'page': 1,
# 'pages': 1,
# 'per_page': 1000,
# 'total': 232}
# ```
#
# That first entry is meta data about the results. For example, it says that there is one page returned with 232 results.
#
# The second entry is another list containing the data. This data would need some cleaning to be used in a pandas data frame. That would happen later in the transformation step of an ETL pipeline. Run the cell below to read the results into a dataframe and see what happens.
# +
###
# Run this cell that converts the json into a dataframe
# Note that you do not need the pd.read_json() method because this is not a file or a string containing json
##
pd.DataFrame(r.json()[1])
# -
# There are some issues with this dataframe. The country and indicator variables don't look particularly useful in their current form. Again, dealing with those issues would come in the transformation phase of a pipeline, which comes later in the lesson.
# # Exercise Indicators API
#
# Use the Indicators API to request rural population data for Switzerland in the years 1995 through 2001. Here are a few helpful resources:
# * [documentation included how to filter by year](https://datahelpdesk.worldbank.org/knowledgebase/articles/898581-api-basic-call-structure)
# * [2-character iso country codes](https://www.nationsonline.org/oneworld/country_code_list.htm)
# * [search box for World Bank indicators](https://data.worldbank.org)
#
# To find the indicator code, first search for the indicator here: https://data.worldbank.org
# Click on the indicator name. The indicator code is in the url. For example, the indicator code for total population is SP.POP.TOTL, which you can see in the link [https://data.worldbank.org/indicator/SP.RUR.TOTL](https://data.worldbank.org/indicator/SP.RUR.TOTL).
# +
# TODO: get the url ready
url = None
# TODO: send the request
r = None
# TODO: output the json using the json method like in the previous example
|
Pipelines/ETLPipelines/4_api_exercise/.ipynb_checkpoints/4_api_exercise-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="b89ce15b"
import sys
# Find jVMC package
sys.path.append(sys.path[0] + "/..")
import jax
from jax.config import config
config.update("jax_enable_x64", True)
import jax.random as random
import jax.numpy as jnp
import flax.linen as nn
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# + colab={"base_uri": "https://localhost:8080/"} id="d22bbd8b" outputId="42386334-83bf-4486-c3e1-a71ab9800123"
# install the pip package and import jVMC
# !pip install jVMC
import jVMC
# + id="058f0001"
L = 10
g = -0.7
GPU_avail = True
# Initialize net
if GPU_avail:
# reproduces results in Fig. 3 of the paper
# estimated run_time in colab (GPU enabled): ~26 minutes
net = jVMC.nets.CNN(F=(L,), channels=(16,), strides=(1,), periodicBoundary=True)
n_steps = 1000
n_Samples = 40000
else:
# may be used to obtain results on Laptop CPUs
# estimated run_time: ~100 seconds
net = jVMC.nets.CpxRBM(numHidden=8, bias=False)
n_steps = 300
n_Samples = 5000
psi = jVMC.vqs.NQS(net, seed=1234) # Variational wave function
# + colab={"base_uri": "https://localhost:8080/"} id="7f4e192b" outputId="7709c307-8753-4e85-cee1-1d16b1a3f46c"
def energy_single_p_mode(h_t, P):
return np.sqrt(1 + h_t**2 - 2 * h_t * np.cos(P))
def ground_state_energy_per_site(h_t, N):
Ps = 0.5 * np.arange(- (N - 1), N - 1 + 2, 2)
Ps = Ps * 2 * np.pi / N
energies_p_modes = np.array([energy_single_p_mode(h_t, P) for P in Ps])
return - 1 / N * np.sum(energies_p_modes)
exact_energy = ground_state_energy_per_site(g, L)
print(exact_energy)
# + id="f6989de3"
# Set up hamiltonian
hamiltonian = jVMC.operator.BranchFreeOperator()
for l in range(L):
hamiltonian.add(jVMC.operator.scal_opstr(-1., (jVMC.operator.Sz(l), jVMC.operator.Sz((l + 1) % L))))
hamiltonian.add(jVMC.operator.scal_opstr(g, (jVMC.operator.Sx(l), )))
# + id="bfd19ac7"
# Set up sampler
sampler = jVMC.sampler.MCSampler(psi, (L,), random.PRNGKey(4321), updateProposer=jVMC.sampler.propose_spin_flip_Z2,
numChains=100, sweepSteps=L,
numSamples=n_Samples, thermalizationSweeps=25)
# Set up TDVP
tdvpEquation = jVMC.util.tdvp.TDVP(sampler, rhsPrefactor=1.,
svdTol=1e-8, diagonalShift=10, makeReal='real')
stepper = jVMC.util.stepper.Euler(timeStep=1e-2) # ODE integrator
# + colab={"base_uri": "https://localhost:8080/"} id="564ab5ad" outputId="2fd1a78c-efbc-4df0-c67b-644133fb2333"
res = []
for n in range(n_steps):
dp, _ = stepper.step(0, tdvpEquation, psi.get_parameters(), hamiltonian=hamiltonian, psi=psi, numSamples=None)
psi.set_parameters(dp)
print(n, jax.numpy.real(tdvpEquation.ElocMean0) / L, tdvpEquation.ElocVar0 / L)
res.append([n, jax.numpy.real(tdvpEquation.ElocMean0) / L, tdvpEquation.ElocVar0 / L])
# + colab={"base_uri": "https://localhost:8080/", "height": 355} id="b9d0a52b" outputId="446b1d5a-e9b6-431f-fc1b-ebb8975715aa"
res = np.array(res)
fig, ax = plt.subplots(2,1, sharex=True, figsize=[4.8,4.8])
ax[0].semilogy(res[:, 0], res[:, 1] - exact_energy, '-', label=r"$L="+str(L)+"$")
ax[0].set_ylabel(r'$(E-E_0)/L$')
ax[1].semilogy(res[:, 0], res[:, 2], '-')
ax[1].set_ylabel(r'Var$(E)/L$')
ax[0].legend()
plt.xlabel('iteration')
plt.tight_layout()
plt.savefig('gs_search.pdf')
|
examples/ex0_ground_state_search.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/phamxuansang241/ML-from-scratch/blob/Data-Preparation/Algorithm_Evaluation_Methods.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="crPqtR-InMfc"
from random import seed
from random import randrange
# + [markdown] id="jnjvHF_En3On"
# ### Train and Test Split
# + id="_tf5Za3koE-g"
# Split a dataset into a train and test set
def train_test_split(dataset, split = 0.60):
train = list()
train_size = split * len(dataset)
dataset_copy = list(dataset)
while len(train) < train_size:
index = randrange(len(dataset_copy))
train.append(dataset_copy.pop(index))
return train, dataset_copy
# + [markdown] id="sBPhy-Y9pDa1"
# ### k-fold Cross-Validation Split
# + id="KMNMnCqjpBxM"
# Split a dataset into $k$ folds
def cross_validation_split(dataset, folds = 3):
dataset_split = list()
dataset_copy = list(dataset)
fold_size = int(len(dataset_copy) / folds)
for i in range(folds):
fold = list()
while len(fold) < fold_size:
index = randrange(len(dataset_copy))
fold.append(dataset_copy.pop(index))
dataset_split.append(fold)
return dataset_split
# + [markdown] id="TlD6CaNbqrpc"
# ### Main
# + colab={"base_uri": "https://localhost:8080/"} id="KgcTraP3qpdg" outputId="15b7b2b1-d269-49b8-b52a-02bbd20c7fbb"
# Test train and test split
seed(1)
dataset = [[1], [2], [3], [4], [5], [6], [7], [8], [9], [10]]
train, test = train_test_split(dataset, split = 0.7)
print("Train set: ", train)
print("Test set: ", test)
# + colab={"base_uri": "https://localhost:8080/"} id="Nbo9_UJxrNhe" outputId="b501f4b6-1339-4009-92b3-b7fc93f61646"
# Test cross-validation split
seed(1)
dataset = [[1], [2], [3], [4], [5], [6], [7], [8], [9], [10]]
folds = cross_validation_split(dataset, folds = 3)
len_folds = len(folds)
for i in range(len_folds):
print(f"Fold {i}: {folds[i]}")
# + id="TtL4GC47rhWT"
|
Algorithm_Evaluation_Methods.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import string
df = pd.read_csv('C:\\Users\\<NAME>\\OneDrive\\Desktop\\python_basic\\Ml\\00 ml_course\\DATA\\tips.csv')
# # Series Basics
ser = pd.Series(data = np.random.randint(1,200,26),index=[x for x in string.ascii_lowercase])
df.head()
ser.head()
s = pd.Series(data = {'Sammy':5,'Frank':10,'Spike':7}) # automatically assign keys as index
s
s['Sammy'] # call my key
s[0]# call by index
q1 = {'Japan': 80, 'China': 450, 'India': 200, 'USA': 250}
q2 = {'Brazil': 100, 'India': 210,'USA': 260,'China': 500}
data1 = pd.Series(data=q1)
data2 = pd.Series(data =q2)
data1
data2
data1 + data2 # automatically finds missing and puts null in uncomman nan + data = nan
data1.add(data2,fill_value = 0) #automatically finds missing and puts 0 in uncomman 0+data = data
data1.sub(data2,fill_value = 0)
data1*2 # broadcasting
data1/2
data1.add_prefix('hello')
data1.aggregate(func=np.mean)
# # DataFrames
df.head()
df.describe() # all continous columns
df.info()
len(df)
df1 = pd.DataFrame(data=np.random.randint(2,200,(5,5)),index=['a','b','c','d','e'],columns=['a','b','c','d','e'])
df.index
df.columns
# # Selection and Indexing
# * 1 Column
# * 2 rows
df['tip'].head(3)
df.loc[2] # all values of second row 'use if named index'
df.iloc[2] # use if numeric index
df = df.set_index('Payment ID')
df.head(2)
df.loc['Sun2959']
df.iloc[0]
df = df.reset_index()
df[['tip','CC Number']].head()
df.iloc[[0,1,2]][['tip','CC Number']]
df.drop([0,1],axis=0).head()
df.drop('Payment ID', axis=1).head(3)
# # conditional Filtering
# 1 single condition
# 2 multiple condition
df[df['tip']>7]
# +
# and &
# or |
# not ~
# -
df[(df['tip']>5) & (df['sex'] == 'Female') & (df['smoker']=='Yes')]
options = ['Yes','Male','Sun',5]
df[df['sex'].isin(options) & df['smoker'].isin(options)].head()
# # Apply
def tips(x):
return np.round(x)
df['tip'].apply(tips)
df['tip'].apply(lambda x : np.round(x))
def take(x,y):
return x+y
df[['tip','total_bill']].apply(lambda df : take(df['tip'],df['total_bill']),axis=1)
df['total']= np.vectorize(take)(df['tip'],df['total_bill'])
df.sort_values('tip',ascending=False).head(2)
df.value_counts('time') # used for cateogires
df.corr()
df.max()
df.min()
df['total'].idxmax()
df.iloc[170]
df['sex'].replace(['Female','Male'],['F','M'])
d = {'Female':'F' , 'Male' : 'M'}
df['sex'].map(d)
df['time'].unique()
df['time'].nunique()
df.nlargest(5,columns=['tip','price_per_person'],keep = 'all')
df['tip'].between(2,3,inclusive = True)
simple_df = pd.DataFrame([1,2,2],['a','b','c'])
simple_df
simple_df.duplicated()
simple_df.drop_duplicates()
# # Missing Values
# * Keep null
# * drop null
# * replace null
df = pd.read_csv('C:\\Users\\<NAME>\\OneDrive\\Desktop\\python_basic\\Ml\\00 ml_course\\DATA\\movie_scores.csv')
df.isnull() # Similar to df.isna()
df.notna() # similar to df.notnull()
df[df['first_name'].notna()]
df.dropna() # drop all null
df.dropna(how = 'all')
df.dropna(how='any')
df
df.fillna(value = 'x',axis = 1)
df['first_name'].fillna('name',inplace=True)
df
df['pre_movie_score'].interpolate()
#
|
02 Pandas/Revision_01.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/gptix/DS-Unit-2-Linear-Models/blob/master/module4-logistic-regression/Jud_Taylor_assignment_regression_classification_4.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="n_nyfetxaMnT" colab_type="text"
# Lambda School Data Science
#
# *Unit 2, Sprint 1, Module 4*
#
# ---
# + [markdown] colab_type="text" id="7IXUfiQ2UKj6"
# # Logistic Regression
#
#
# # Assignment 🌯
#
# You'll use a [**dataset of 400+ burrito reviews**](https://srcole.github.io/100burritos/). How accurately can you predict whether a burrito is rated 'Great'?
#
#
# + [markdown] id="tVLrLQa68WP1" colab_type="text"
# ## Prepare import path based on environment.
# + colab_type="code" id="o9eSnDYhUGD7" colab={}
# %%capture
import sys
# If you're on Colab:
if 'google.colab' in sys.modules:
DATA_PATH = 'https://raw.githubusercontent.com/LambdaSchool/DS-Unit-2-Linear-Models/master/data/'
# !pip install category_encoders==2.*
# If you're working locally:
else:
DATA_PATH = '../data/'
# + [markdown] id="UWrJYDwt8hLg" colab_type="text"
# ## Import useful libraries.
# + id="duBJV7cGbDBE" colab_type="code" colab={}
# Import useful libraries.
import pandas as pd
import numpy as np
import category_encoders as ce
from sklearn.linear_model import LogisticRegression
from sklearn.impute import SimpleImputer
from sklearn.linear_model import LogisticRegressionCV
from sklearn.preprocessing import StandardScaler
# + [markdown] id="NGnJjh9P8m_4" colab_type="text"
# ## Load raw data.
# + id="_mSCJytXaMne" colab_type="code" colab={}
# Load data downloaded from https://srcole.github.io/100burritos/
raw_df = pd.read_csv(DATA_PATH+'burritos/burritos.csv')
# + [markdown] id="-rjOIRZw8yNI" colab_type="text"
# ## Make a working copy of raw data.
# + id="ePKdX-iAa2Ea" colab_type="code" colab={}
# Make a working copy of the raw df.
df = raw_df
# + [markdown] id="1eUV31qt8sFb" colab_type="text"
# ## Review dataframe.
# + id="iGa-jcGjbNt9" colab_type="code" outputId="494bbbbc-c7d2-4ea0-db3e-1d4873854658" colab={"base_uri": "https://localhost:8080/", "height": 34}
# Review dataframe.
# df
# df.describe()
# df.describe().T
# df.head()
# df.tail()
# df.columns
# df.dtypes
df.shape
# df.columns.isna()
# df.isna()
# + [markdown] id="VQudMNuJao-n" colab_type="text"
# > We have developed a 10-dimensional system for rating the burritos in San Diego. ... Generate models for what makes a burrito great and investigate correlations in its dimensions.
#
#
# + [markdown] id="UbjlwGvC9Nwd" colab_type="text"
# ## Engineer data.
# + id="JajdYoWonunT" colab_type="code" colab={}
# Replace spaces in column names to underbars
df.columns = [col.replace(' ', '_') for col in df]
# df.columns
# df.head()
# + id="vc3iHtkSt9ry" colab_type="code" colab={}
# Convert ingredients columns to one-hot encodings
ingredients = ['Beef','Pico','Guac','Cheese','Fries','Sour_cream','Pork','Chicken',
'Shrimp','Fish','Rice','Beans','Lettuce','Tomato','Bell_peper',
'Carrots','Cabbage','Sauce','Salsa.1','Cilantro','Onion',
'Taquito','Pineapple','Ham','Chile_relleno','Nopales','Lobster',
'Queso','Egg','Mushroom','Bacon','Sushi','Avocado','Corn',
'Zucchini']
# quick utility
def check_uniques():
'''List unique values in columns.'''
for i in ingredients:
print(i + str(df[i].unique()))
# + id="4TvCLULI6Dkm" colab_type="code" colab={}
# check_uniques()
# + id="uyzsyLw94RHu" colab_type="code" colab={}
# Replace non-zeroes in columns with ones
for i in ingredients:
df[i] = df[i].map({'x': 1, 'X': 1})
# check_uniques()
# + id="nGdLzRm63hT0" colab_type="code" colab={}
# Replace NaN's with 0
for i in ingredients:
df[i] = df[i].fillna(0)
# check_uniques()
# + id="ARuob7ySaMnj" colab_type="code" outputId="0fabfdee-8ec9-4406-9f76-4c54b14e9716" colab={"base_uri": "https://localhost:8080/", "height": 139}
# We will use an 'overall' value as 'Great', which will be encoded as True/1
# Drop unrated burritos.
df = df.dropna(subset=['overall'])
# Add a feature to hold TF for 'Great'
df['Great'] = df['overall'] >= 4
# + id="W3yS-IVT-YKS" colab_type="code" colab={}
# df.Great
# + id="Htobs9Y8aMno" colab_type="code" outputId="412a90dd-dda8-4ee5-f501-73bfa43875a8" colab={"base_uri": "https://localhost:8080/", "height": 241}
# Clean/combine the Burrito categories
# make all text in 'Burrito' column lowercase, to simplify manipulaton.
df['Burrito'] = df['Burrito'].str.lower()
# define predicates (Y/N tests)
california = df['Burrito'].str.contains('california')
asada = df['Burrito'].str.contains('asada')
surf = df['Burrito'].str.contains('surf')
carnitas = df['Burrito'].str.contains('carnitas')
# Apply predicates, and apply catchall for remainder
df.loc[california, 'Burrito'] = 'California'
df.loc[asada, 'Burrito'] = 'Asada'
df.loc[surf, 'Burrito'] = 'Surf & Turf'
df.loc[carnitas, 'Burrito'] = 'Carnitas'
df.loc[~california & ~asada & ~surf & ~carnitas, 'Burrito'] = 'Other'
# + id="xtmR7PAs_Tya" colab_type="code" colab={}
# df.Burrito
# + id="IO_DhxjcaMnt" colab_type="code" colab={}
# Drop some high cardinality categoricals
df = df.drop(columns=['Notes', 'Location', 'Reviewer', 'Address', 'URL', 'Neighborhood'])
# + id="ih9XVbZNaMny" colab_type="code" colab={}
# Drop some columns to prevent "leakage"
df = df.drop(columns=['Rec', 'overall'])
# + [markdown] id="qFdLT-8i9AjT" colab_type="text"
# ## Split into Train, Validate, Test sets
# - [ ] Do train/validate/test split. Train on reviews from 2016 & earlier. Validate on 2017. Test on 2018 & later.
# + id="NZ0MMp4UcYC_" colab_type="code" outputId="60226854-9a72-45b1-e98b-25b613896300" colab={"base_uri": "https://localhost:8080/", "height": 221}
# Create a field to hold year.
df['Year'] = pd.DatetimeIndex( pd.to_datetime(df['Date'], format='%m/%d/%Y') ).year
df['Year']
# + id="RJBfUqAwg0qe" colab_type="code" outputId="8705a2ab-f70f-422a-b628-6936254d696e" colab={"base_uri": "https://localhost:8080/", "height": 1000}
train = df[df['Year'] <= 2016]
val = df[df['Year'] == 2017]
test = df[df['Year'] >= 2018]
shapes = train.shape, val.shape, test.shape
df.shape
# shapes
df.isna().sum()
# + id="s0yWOlPN6BTu" colab_type="code" outputId="942f9a3c-3da8-4f66-a929-051cf25464b4" colab={"base_uri": "https://localhost:8080/", "height": 34}
# Drop columns with large numbers of NaN's
# drop_because_too_many_NaNs = ['Yelp', 'Google', 'Chips', 'Mass_(g)',
# 'Density_(g/mL)', 'Length', 'Circum', 'Volume',
# 'Unreliable', 'NonSD']
# df = df.drop(columns=drop_because_too_many_NaNs)
df.shape
# + id="IexCqDUp8CpB" colab_type="code" outputId="5b0a14a9-3d58-41f3-a0d5-fbf973d0fe1e" colab={"base_uri": "https://localhost:8080/", "height": 1000}
df.isna().sum()
# df['Temp'].isna().count()
df.dtypes
# + id="4JKrQo7_9WeI" colab_type="code" outputId="9723c6e3-02d0-4aa3-9a18-840f34b32c3f" colab={"base_uri": "https://localhost:8080/", "height": 34}
# Drop rows with NaN
df = df.dropna()
df.shape
# + [markdown] id="kRYE-kfUAeOa" colab_type="text"
# ## Baseline classification
# - [ ] Begin with baselines for classification.
# + id="4ySTvhpEBJel" colab_type="code" outputId="991af44a-be42-4e02-c7e0-b0182b5c26af" colab={"base_uri": "https://localhost:8080/", "height": 68}
target = 'Great'
y_train = train[target]
print(y_train.shape)
split_train = y_train.value_counts(normalize=True)
great_percentage_train = split_train[1]*100
baseline = y_train.mode().values[0]
print("Baseline: is it great? - " + str(baseline))
print("Percentage of great burritos: " + str(round(great_percentage_train, 2)))
# + id="8vEOY2weDAcx" colab_type="code" outputId="cce20ba1-ac69-4fa8-ecee-2e24666e8ca9" colab={"base_uri": "https://localhost:8080/", "height": 51}
# Test baseline against val set
y_val = val[target]
print(y_val.shape)
split_val = y_val.value_counts(normalize=True)
# y_val.value_counts(normalize=True)
great_percentage_val = split_val[1]*100
print("Percentage of great burritos: " + str(round(great_percentage_val, 2)))
# print("Difference between ")
# + [markdown] id="RArybLnnEboH" colab_type="text"
# ## Logistic Regression
# - [ ] Use scikit-learn for logistic regression.
# + id="MCZCCcxHEWvW" colab_type="code" outputId="f51848a4-321f-4262-ea44-c57ec44c09ce" colab={"base_uri": "https://localhost:8080/", "height": 224}
features = ['Burrito', 'Cost', 'Hunger', 'Tortilla', 'Temp', 'Meat', 'Fillings',
'Uniformity', 'Salsa', 'Synergy', 'Wrap', 'Beef', 'Pork',
'Chicken', 'Shrimp', 'Fish']
target = 'Great'
X_train = train[features]
y_train = train[target]
X_val = val[features]
y_val = val[target]
log_reg = LogisticRegression(solver='lbfgs')
X_train.shape, y_train.shape, X_val.shape, y_val.shape
X_train.head()
# + id="bCiF6fGKMMvz" colab_type="code" colab={}
# log_reg.fit(X_train, y_train)
# print('Validation Accuracy', log_reg.score(X_val_imputed, y_val))
# + id="uCUYsxehO_49" colab_type="code" colab={}
encoder = ce.OneHotEncoder(use_cat_names=True)
X_train_encoded = encoder.fit_transform(X_train)
X_val_encoded = encoder.transform(X_val)
X_train_encoded.head()
# + id="9VUd3T-pQxUe" colab_type="code" colab={}
imputer = SimpleImputer()
X_train_imputed = imputer.fit_transform(X_train_encoded)
X_val_imputed = imputer.transform(X_val_encoded)
X_train_imputed[:5]
# + id="6jF_o4tYQ4i4" colab_type="code" colab={}
scaler = StandardScaler()
X_train_scaled = scaler.fit_transform(X_train_imputed)
X_val_scaled = scaler.transform(X_val_imputed)
# + id="zZJJFlc5SwI7" colab_type="code" colab={}
X_train_scaled[:5]
# + id="jgPTwoTyUB0I" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 102} outputId="34462b8f-5c0c-477d-b199-575a91776a72"
model= LogisticRegressionCV(cv=5)
model.fit(X_train_scaled, y_train)
# + [markdown] id="ME_sZ2nbQX4P" colab_type="text"
# - [ ] Get your model's validation accuracy. (Multiple times if you try multiple iterations.)
#
# + id="6yfKD2FpU0Nw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="72d41d96-e07c-42c6-e233-67ab69d022f6"
print('Validtion accuracy', model.score(X_val_scaled, y_val))
# + [markdown] id="pOwV8Rf3lWjs" colab_type="text"
# - [ ] Get your model's test accuracy. (One time, at the end.)
# + id="P80De2pJW-7U" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="de5bd0d5-ae7e-40b2-9c14-bf62e493d36e"
X_test = test[features]
y_test = test[target]
X_test_encoded = encoder.fit_transform(X_test)
X_test_imputed = imputer.fit_transform(X_test_encoded)
X_test_scaled = scaler.fit_transform(X_test_imputed)
print('Test accuracy', model.score(X_test_scaled, y_test))
# + id="cpmUmkUrVQIv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 282} outputId="cf638df8-6532-4f39-8454-075a5ae507c0"
pd.Series(model.coef_[0], X_train_encoded.columns).sort_values().plot.barh()
# + [markdown] id="hEsd4KYbVHYw" colab_type="text"
# - [ ] Commit your notebook to your fork of the GitHub repo.
# **Done**
# + [markdown] id="w1bwk5tKVJ1R" colab_type="text"
# ## Stretch Goals
#
# - [ ] Add your own stretch goal(s) **Done (trnsformed fillings to numeric by hnd)**
# - [ ] Make exploratory visualizations. **Done**
# - [ ] Do one-hot encoding. **Done**
# - [ ] Do [feature scaling](https://scikit-learn.org/stable/modules/preprocessing.html). **Done**
# - [ ] Get and plot your coefficients. **Done**
# - [ ] Try [scikit-learn pipelines](https://scikit-learn.org/stable/modules/compose.html).
# + id="IRrFNL7LaMn2" colab_type="code" colab={}
|
module4-logistic-regression/Jud_Taylor_assignment_regression_classification_4.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # cRNN Lorenz-96 Demo
# - In this notebook, we train a cRNN model on data simulated from a Lorenz-96 system.
import numpy as np
import torch
from models.crnn import cRNN, train_model_gista
from synthetic import simulate_lorenz_96
import matplotlib.pyplot as plt
# For GPU acceleration
device = torch.device('cuda')
# Simulate data
p = 10
X_np, GC = simulate_lorenz_96(p, T=1000)
X = torch.tensor(X_np[np.newaxis], dtype=torch.float32, device=device)
# Plot data
fig, axarr = plt.subplots(1, 2, figsize=(16, 5))
axarr[0].plot(X_np)
axarr[1].plot(X_np[:100])
plt.show()
# Set up model
crnn = cRNN(p, hidden=10).cuda(device=device)
# Train with GISTA
check_every = 100
train_loss_list, train_mse_list = train_model_gista(
crnn, X, lam=6.3, lam_ridge=1e-4, lr=0.005, max_iter=20000, check_every=check_every, truncation=5)
# +
# Loss function plot
fig, axarr = plt.subplots(1, 2, figsize=(16, 5))
axarr[0].plot(check_every * np.arange(len(train_loss_list)), train_loss_list)
axarr[0].set_title('Train loss')
axarr[1].plot(check_every * np.arange(len(train_mse_list)), train_mse_list)
axarr[1].set_title('Train MSE')
plt.show()
# +
# Verify learned Granger causality
GC_est = crnn.GC().cpu().data.numpy()
print('True variable usage = %.2f%%' % (100 * np.mean(GC)))
print('Estimated variable usage = %.2f%%' % (100 * np.mean(GC_est)))
print('Accuracy = %.2f%%' % (100 * np.mean(GC == GC_est)))
# Make figures
fig, axarr = plt.subplots(1, 2, figsize=(10, 5))
axarr[0].imshow(GC, cmap='Blues')
axarr[0].set_title('GC actual')
axarr[0].set_ylabel('Affected series')
axarr[0].set_xlabel('Causal series')
axarr[0].set_xticks([])
axarr[0].set_yticks([])
axarr[1].imshow(GC_est, cmap='Blues', vmin=0, vmax=1, extent=(0, p, p, 0))
axarr[1].set_ylabel('Affected series')
axarr[1].set_xlabel('Causal series')
axarr[1].set_xticks([])
axarr[1].set_yticks([])
# Mark disagreements
for i in range(p):
for j in range(p):
if GC[i, j] != GC_est[i, j]:
rect = plt.Rectangle((j, i-0.05), 1, 1, facecolor='none', edgecolor='red', linewidth=1)
axarr[1].add_patch(rect)
plt.show()
# -
|
Neural-GC/crnn_lorenz_demo.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
a = 3
type(a)
a = 3.2
type(a)
a = "a"
type(a)
id(a)
a = True
type(a)
a = input("enter number")
type(a)
b = int(a)
type(b)
c = str(b)
type(c)
a =float(3)
a
b =bool(0)
type(b)
b
b = bool(1)
type(b)
b
b = bool(-1)
type(b)
b
b = bool(3)
type(b)
b
a = [1,"sameer",1.3,True]
a
a[1]
a[1][0]
a.append(35)
a
a.insert(2,"qasim")
a
a[1]="zohaib"
a
b = a[0:3]
b
b = a[1:]
b
b = a[:3]
b
a
del a[1]
a
a
a.remove(35)
a
a =[1,2,3,4,5,6,7,8.9]
a
b = a.pop()
b
a = 3
3 == 2
2 == 2
4<=3
4>=3
3>=3
3 != 2
False or False
True and True
False and False
False and True
a = [1,2,3,4, 5 ,6]
a
a is 3
2 is 2
for b in a:
if b is 7:
print("true")
elif b is 4:
print("4 is in list")
break
else:
print("false")
a
for b in a:
print(b)
c = input("ENter Your age")
c = int(c)
d = input("Enter Your Gender")
# +
if c<=25:
if d=="m":
print("congratulation you selected")
# -
for a in range(1,10):
print(a)
a = [1,2,3,4,5,6,7]
for b in a:
print(b)
i = 0
while i<=10:
print(i)
i+=1
for i in range(0,10):
print(i)
if i != 11:
continue
|
Q1/batch1/Notebooks files/PRACTICE SAMEER.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Machine Learning Engineer Nanodegree
# ## Supervised Learning
# ## Project: Finding Donors for *CharityML*
# Welcome to the second project of the Machine Learning Engineer Nanodegree! In this notebook, some template code has already been provided for you, and it will be your job to implement the additional functionality necessary to successfully complete this project. Sections that begin with **'Implementation'** in the header indicate that the following block of code will require additional functionality which you must provide. Instructions will be provided for each section and the specifics of the implementation are marked in the code block with a `'TODO'` statement. Please be sure to read the instructions carefully!
#
# In addition to implementing code, there will be questions that you must answer which relate to the project and your implementation. Each section where you will answer a question is preceded by a **'Question X'** header. Carefully read each question and provide thorough answers in the following text boxes that begin with **'Answer:'**. Your project submission will be evaluated based on your answers to each of the questions and the implementation you provide.
#
# >**Note:** Please specify WHICH VERSION OF PYTHON you are using when submitting this notebook. Code and Markdown cells can be executed using the **Shift + Enter** keyboard shortcut. In addition, Markdown cells can be edited by typically double-clicking the cell to enter edit mode.
# ## Getting Started
#
# In this project, you will employ several supervised algorithms of your choice to accurately model individuals' income using data collected from the 1994 U.S. Census. You will then choose the best candidate algorithm from preliminary results and further optimize this algorithm to best model the data. Your goal with this implementation is to construct a model that accurately predicts whether an individual makes more than $50,000. This sort of task can arise in a non-profit setting, where organizations survive on donations. Understanding an individual's income can help a non-profit better understand how large of a donation to request, or whether or not they should reach out to begin with. While it can be difficult to determine an individual's general income bracket directly from public sources, we can (as we will see) infer this value from other publically available features.
#
# The dataset for this project originates from the [UCI Machine Learning Repository](https://archive.ics.uci.edu/ml/datasets/Census+Income). The datset was donated by <NAME> and <NAME>, after being published in the article _"Scaling Up the Accuracy of Naive-Bayes Classifiers: A Decision-Tree Hybrid"_. You can find the article by <NAME> [online](https://www.aaai.org/Papers/KDD/1996/KDD96-033.pdf). The data we investigate here consists of small changes to the original dataset, such as removing the `'fnlwgt'` feature and records with missing or ill-formatted entries.
# ----
# ## Exploring the Data
# Run the code cell below to load necessary Python libraries and load the census data. Note that the last column from this dataset, `'income'`, will be our target label (whether an individual makes more than, or at most, $50,000 annually). All other columns are features about each individual in the census database.
# +
# !python3 --version
# Import libraries necessary for this project
import numpy as np
import pandas as pd
from time import time
from IPython.display import display # Allows the use of display() for DataFrames
# Import supplementary visualization code visuals.py
import visuals as vs
# Pretty display for notebooks
# %matplotlib inline
# Load the Census dataset
data = pd.read_csv("census.csv")
# Success - Display the first record
display(data.head(n=1))
# -
# ### Implementation: Data Exploration
# A cursory investigation of the dataset will determine how many individuals fit into either group, and will tell us about the percentage of these individuals making more than \$50,000. In the code cell below, you will need to compute the following:
# - The total number of records, `'n_records'`
# - The number of individuals making more than \$50,000 annually, `'n_greater_50k'`.
# - The number of individuals making at most \$50,000 annually, `'n_at_most_50k'`.
# - The percentage of individuals making more than \$50,000 annually, `'greater_percent'`.
#
# ** HINT: ** You may need to look at the table above to understand how the `'income'` entries are formatted.
# +
# TODO: Total number of records
n_records = data.shape[0]
# TODO: Number of records where individual's income is more than $50,000
n_greater_50k = data[data['income'] == '>50K'].shape[0]
# TODO: Number of records where individual's income is at most $50,000
n_at_most_50k = data[data['income'] == '<=50K'].shape[0]
# TODO: Percentage of individuals whose income is more than $50,000
greater_percent = (n_greater_50k*100)/n_records
# Print the results
print("Total number of records: {}".format(n_records))
print("Individuals making more than $50,000: {}".format(n_greater_50k))
print("Individuals making at most $50,000: {}".format(n_at_most_50k))
print("Percentage of individuals making more than $50,000: {}%".format(greater_percent))
# -
# ** Featureset Exploration **
#
# * **age**: continuous.
# * **workclass**: Private, Self-emp-not-inc, Self-emp-inc, Federal-gov, Local-gov, State-gov, Without-pay, Never-worked.
# * **education**: Bachelors, Some-college, 11th, HS-grad, Prof-school, Assoc-acdm, Assoc-voc, 9th, 7th-8th, 12th, Masters, 1st-4th, 10th, Doctorate, 5th-6th, Preschool.
# * **education-num**: continuous.
# * **marital-status**: Married-civ-spouse, Divorced, Never-married, Separated, Widowed, Married-spouse-absent, Married-AF-spouse.
# * **occupation**: Tech-support, Craft-repair, Other-service, Sales, Exec-managerial, Prof-specialty, Handlers-cleaners, Machine-op-inspct, Adm-clerical, Farming-fishing, Transport-moving, Priv-house-serv, Protective-serv, Armed-Forces.
# * **relationship**: Wife, Own-child, Husband, Not-in-family, Other-relative, Unmarried.
# * **race**: Black, White, Asian-Pac-Islander, Amer-Indian-Eskimo, Other.
# * **sex**: Female, Male.
# * **capital-gain**: continuous.
# * **capital-loss**: continuous.
# * **hours-per-week**: continuous.
# * **native-country**: United-States, Cambodia, England, Puerto-Rico, Canada, Germany, Outlying-US(Guam-USVI-etc), India, Japan, Greece, South, China, Cuba, Iran, Honduras, Philippines, Italy, Poland, Jamaica, Vietnam, Mexico, Portugal, Ireland, France, Dominican-Republic, Laos, Ecuador, Taiwan, Haiti, Columbia, Hungary, Guatemala, Nicaragua, Scotland, Thailand, Yugoslavia, El-Salvador, Trinadad&Tobago, Peru, Hong, Holand-Netherlands.
# ----
# ## Preparing the Data
# Before data can be used as input for machine learning algorithms, it often must be cleaned, formatted, and restructured — this is typically known as **preprocessing**. Fortunately, for this dataset, there are no invalid or missing entries we must deal with, however, there are some qualities about certain features that must be adjusted. This preprocessing can help tremendously with the outcome and predictive power of nearly all learning algorithms.
# ### Transforming Skewed Continuous Features
# A dataset may sometimes contain at least one feature whose values tend to lie near a single number, but will also have a non-trivial number of vastly larger or smaller values than that single number. Algorithms can be sensitive to such distributions of values and can underperform if the range is not properly normalized. With the census dataset two features fit this description: '`capital-gain'` and `'capital-loss'`.
#
# Run the code cell below to plot a histogram of these two features. Note the range of the values present and how they are distributed.
# +
# Split the data into features and target label
income_raw = data['income']
features_raw = data.drop('income', axis = 1)
# Visualize skewed continuous features of original data
vs.distribution(data)
# -
# For highly-skewed feature distributions such as `'capital-gain'` and `'capital-loss'`, it is common practice to apply a <a href="https://en.wikipedia.org/wiki/Data_transformation_(statistics)">logarithmic transformation</a> on the data so that the very large and very small values do not negatively affect the performance of a learning algorithm. Using a logarithmic transformation significantly reduces the range of values caused by outliers. Care must be taken when applying this transformation however: The logarithm of `0` is undefined, so we must translate the values by a small amount above `0` to apply the the logarithm successfully.
#
# Run the code cell below to perform a transformation on the data and visualize the results. Again, note the range of values and how they are distributed.
# +
# Log-transform the skewed features
skewed = ['capital-gain', 'capital-loss']
features_log_transformed = pd.DataFrame(data = features_raw)
features_log_transformed[skewed] = features_raw[skewed].apply(lambda x: np.log(x + 1))
# Visualize the new log distributions
vs.distribution(features_log_transformed, transformed = True)
# -
# ### Normalizing Numerical Features
# In addition to performing transformations on features that are highly skewed, it is often good practice to perform some type of scaling on numerical features. Applying a scaling to the data does not change the shape of each feature's distribution (such as `'capital-gain'` or `'capital-loss'` above); however, normalization ensures that each feature is treated equally when applying supervised learners. Note that once scaling is applied, observing the data in its raw form will no longer have the same original meaning, as exampled below.
#
# Run the code cell below to normalize each numerical feature. We will use [`sklearn.preprocessing.MinMaxScaler`](http://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.MinMaxScaler.html) for this.
# +
# Import sklearn.preprocessing.StandardScaler
from sklearn.preprocessing import MinMaxScaler
# Initialize a scaler, then apply it to the features
scaler = MinMaxScaler() # default=(0, 1)
numerical = ['age', 'education-num', 'capital-gain', 'capital-loss', 'hours-per-week']
features_log_minmax_transform = pd.DataFrame(data = features_log_transformed)
features_log_minmax_transform[numerical] = scaler.fit_transform(features_log_transformed[numerical])
# Show an example of a record with scaling applied
display(features_log_minmax_transform.head(n = 5))
# -
# ### Implementation: Data Preprocessing
#
# From the table in **Exploring the Data** above, we can see there are several features for each record that are non-numeric. Typically, learning algorithms expect input to be numeric, which requires that non-numeric features (called *categorical variables*) be converted. One popular way to convert categorical variables is by using the **one-hot encoding** scheme. One-hot encoding creates a _"dummy"_ variable for each possible category of each non-numeric feature. For example, assume `someFeature` has three possible entries: `A`, `B`, or `C`. We then encode this feature into `someFeature_A`, `someFeature_B` and `someFeature_C`.
#
# | | someFeature | | someFeature_A | someFeature_B | someFeature_C |
# | :-: | :-: | | :-: | :-: | :-: |
# | 0 | B | | 0 | 1 | 0 |
# | 1 | C | ----> one-hot encode ----> | 0 | 0 | 1 |
# | 2 | A | | 1 | 0 | 0 |
#
# Additionally, as with the non-numeric features, we need to convert the non-numeric target label, `'income'` to numerical values for the learning algorithm to work. Since there are only two possible categories for this label ("<=50K" and ">50K"), we can avoid using one-hot encoding and simply encode these two categories as `0` and `1`, respectively. In code cell below, you will need to implement the following:
# - Use [`pandas.get_dummies()`](http://pandas.pydata.org/pandas-docs/stable/generated/pandas.get_dummies.html?highlight=get_dummies#pandas.get_dummies) to perform one-hot encoding on the `'features_log_minmax_transform'` data.
# - Convert the target label `'income_raw'` to numerical entries.
# - Set records with "<=50K" to `0` and records with ">50K" to `1`.
# +
# TODO: One-hot encode the 'features_log_minmax_transform' data using pandas.get_dummies()
features_final = pd.get_dummies(features_log_minmax_transform)
# TODO: Encode the 'income_raw' data to numerical values
income = income_raw.apply(lambda x:1 if x=='>50K' else 0)
# # Print the number of features after one-hot encoding
encoded = list(features_final.columns)
print("{} total features after one-hot encoding.".format(len(encoded)))
# Uncomment the following line to see the encoded feature names
print(encoded)
# -
# ### Shuffle and Split Data
# Now all _categorical variables_ have been converted into numerical features, and all numerical features have been normalized. As always, we will now split the data (both features and their labels) into training and test sets. 80% of the data will be used for training and 20% for testing.
#
# Run the code cell below to perform this split.
# +
# Import train_test_split
from sklearn.model_selection import train_test_split
# Split the 'features' and 'income' data into training and testing sets
X_train, X_test, y_train, y_test = train_test_split(features_final,
income,
test_size = 0.2,
random_state = 0)
# Show the results of the split
print("Training set has {} samples.".format(X_train.shape[0]))
print("Testing set has {} samples.".format(X_test.shape[0]))
# -
# ----
# ## Evaluating Model Performance
# In this section, we will investigate four different algorithms, and determine which is best at modeling the data. Three of these algorithms will be supervised learners of your choice, and the fourth algorithm is known as a *naive predictor*.
# ### Metrics and the Naive Predictor
# *CharityML*, equipped with their research, knows individuals that make more than \$50,000 are most likely to donate to their charity. Because of this, *CharityML* is particularly interested in predicting who makes more than \$50,000 accurately. It would seem that using **accuracy** as a metric for evaluating a particular model's performace would be appropriate. Additionally, identifying someone that *does not* make more than \$50,000 as someone who does would be detrimental to *CharityML*, since they are looking to find individuals willing to donate. Therefore, a model's ability to precisely predict those that make more than \$50,000 is *more important* than the model's ability to **recall** those individuals. We can use **F-beta score** as a metric that considers both precision and recall:
#
# $$ F_{\beta} = (1 + \beta^2) \cdot \frac{precision \cdot recall}{\left( \beta^2 \cdot precision \right) + recall} $$
#
# In particular, when $\beta = 0.5$, more emphasis is placed on precision. This is called the **F$_{0.5}$ score** (or F-score for simplicity).
#
# Looking at the distribution of classes (those who make at most \$50,000, and those who make more), it's clear most individuals do not make more than \$50,000. This can greatly affect **accuracy**, since we could simply say *"this person does not make more than \$50,000"* and generally be right, without ever looking at the data! Making such a statement would be called **naive**, since we have not considered any information to substantiate the claim. It is always important to consider the *naive prediction* for your data, to help establish a benchmark for whether a model is performing well. That been said, using that prediction would be pointless: If we predicted all people made less than \$50,000, *CharityML* would identify no one as donors.
#
#
# #### Note: Recap of accuracy, precision, recall
#
# ** Accuracy ** measures how often the classifier makes the correct prediction. It’s the ratio of the number of correct predictions to the total number of predictions (the number of test data points).
#
# ** Precision ** tells us what proportion of messages we classified as spam, actually were spam.
# It is a ratio of true positives(words classified as spam, and which are actually spam) to all positives(all words classified as spam, irrespective of whether that was the correct classificatio), in other words it is the ratio of
#
# `[True Positives/(True Positives + False Positives)]`
#
# ** Recall(sensitivity)** tells us what proportion of messages that actually were spam were classified by us as spam.
# It is a ratio of true positives(words classified as spam, and which are actually spam) to all the words that were actually spam, in other words it is the ratio of
#
# `[True Positives/(True Positives + False Negatives)]`
#
# For classification problems that are skewed in their classification distributions like in our case, for example if we had a 100 text messages and only 2 were spam and the rest 98 weren't, accuracy by itself is not a very good metric. We could classify 90 messages as not spam(including the 2 that were spam but we classify them as not spam, hence they would be false negatives) and 10 as spam(all 10 false positives) and still get a reasonably good accuracy score. For such cases, precision and recall come in very handy. These two metrics can be combined to get the F1 score, which is weighted average(harmonic mean) of the precision and recall scores. This score can range from 0 to 1, with 1 being the best possible F1 score(we take the harmonic mean as we are dealing with ratios).
# ### Question 1 - Naive Predictor Performace
# * If we chose a model that always predicted an individual made more than $50,000, what would that model's accuracy and F-score be on this dataset? You must use the code cell below and assign your results to `'accuracy'` and `'fscore'` to be used later.
#
# ** Please note ** that the the purpose of generating a naive predictor is simply to show what a base model without any intelligence would look like. In the real world, ideally your base model would be either the results of a previous model or could be based on a research paper upon which you are looking to improve. When there is no benchmark model set, getting a result better than random choice is a place you could start from.
#
# ** HINT: **
#
# * When we have a model that always predicts '1' (i.e. the individual makes more than 50k) then our model will have no True Negatives(TN) or False Negatives(FN) as we are not making any negative('0' value) predictions. Therefore our Accuracy in this case becomes the same as our Precision(True Positives/(True Positives + False Positives)) as every prediction that we have made with value '1' that should have '0' becomes a False Positive; therefore our denominator in this case is the total number of records we have in total.
# * Our Recall score(True Positives/(True Positives + False Negatives)) in this setting becomes 1 as we have no False Negatives.
# +
TP = np.sum(income) # Counting the ones as this is the naive case. Note that 'income' is the 'income_raw' data
# encoded to numerical values done in the data preprocessing step.
FP = income.count() - TP # Specific to the naive case
TN = 0 # No predicted negatives in the naive case
FN = 0 # No predicted negatives in the naive case
# TODO: Calculate accuracy, precision and recall
accuracy = TP/(TP+FP)
recall = TP/(TP+FN)
precision = TP/(TP+FP)
# TODO: Calculate F-score using the formula above for beta = 0.5 and correct values for precision and recall.
beta = 0.5
fscore = (1 + beta * beta) * ((precision * recall)/((beta * beta * precision) + recall))
# Print the results
print("Naive Predictor: [Accuracy score: {:.4f}, F-score: {:.4f}]".format(accuracy, fscore))
# -
# ### Supervised Learning Models
# **The following are some of the supervised learning models that are currently available in** [`scikit-learn`](http://scikit-learn.org/stable/supervised_learning.html) **that you may choose from:**
# - Gaussian Naive Bayes (GaussianNB)
# - Decision Trees
# - Ensemble Methods (Bagging, AdaBoost, Random Forest, Gradient Boosting)
# - K-Nearest Neighbors (KNeighbors)
# - Stochastic Gradient Descent Classifier (SGDC)
# - Support Vector Machines (SVM)
# - Logistic Regression
# ### Question 2 - Model Application
# List three of the supervised learning models above that are appropriate for this problem that you will test on the census data. For each model chosen
#
# - Describe one real-world application in industry where the model can be applied.
# - What are the strengths of the model; when does it perform well?
# - What are the weaknesses of the model; when does it perform poorly?
# - What makes this model a good candidate for the problem, given what you know about the data?
#
# ** HINT: **
#
# Structure your answer in the same format as above^, with 4 parts for each of the three models you pick. Please include references with your answer.
# **Answer: **
#
# AdaBoost:
# - Head gesture recognition for hands‐free control of an intelligent wheelchair, detection of traffic signs, predicting subscriber dissatisfaction and improving retention in the wireless telecommunications industry. [3][4][5]
# - Can achieve similar classification results that more complex models with less tweaking of parameters os settings and is very simple. Can be used to feature selection on very large sets of features. [6]
# - Can be sensitive to noisy data and outliers. [6]
# - This classificator is often referred to as the best out-of-the-box classifier and as described in the strenghts, it can perform well in complex models with a good performance. [7]
#
# Random Forest:
# - It can be used for quality assessment of Wikipedia articles and Machine Fault Diagnosis. [9][10]
# - Combine predictions from many indiviual trees and can lean non-linear relationships. It is robust to outliers. [8]
# - Unconstrained, individual trees are prone to overfiting because they can keep branching until they memorize the training data. When used for regression they cannot predict beyond the range in the training data. [8]
# - It can deal with non-linear relationships on this data set, making it a good canditate to this scenario.
#
# GaussianNB:
# - It can be used in text categorization (i.e. spam detection) and automatic medical diagnosis. [1]
# - Easy to implement and eficient when there is a large number of features. [2]
# - When features are related or the order counts a lot the NB does a poor job due to this naive characteristic. Easily beaten by more complex tuned models. [2]
# - There is a total of 103 features on this data set, a naive aproach can perform well in this scenario and will be really fast to test the results.
#
# [1] https://en.wikipedia.org/wiki/Naive_Bayes_classifier
#
# [2] https://www.youtube.com/watch?v=nfbKTrufPOs - Udacity video
#
# [3] https://doi.org/10.1108/01439910710718469 - <NAME>, <NAME>, <NAME>, <NAME>, (2007) "Head gesture recognition for hands‐free control of an intelligent wheelchair", Industrial Robot: An International Journal, Vol. 34 Issue: 1, pp.60-68
#
# [4] http://ieeexplore.ieee.org/stamp/stamp.jsp?tp=&arnumber=6706807&isnumber=6706705 - <NAME>, <NAME>, <NAME>, <NAME> and <NAME>, "Detection of traffic signs in real-world images: The German traffic sign detection benchmark," The 2013 International Joint Conference on Neural Networks (IJCNN), Dallas, TX, 2013, pp. 1-8.
#
# [5] http://ieeexplore.ieee.org/stamp/stamp.jsp?tp=&arnumber=846740&isnumber=18371 - <NAME>, <NAME>, <NAME>, <NAME> and <NAME>, "Predicting subscriber dissatisfaction and improving retention in the wireless telecommunications industry," in IEEE Transactions on Neural Networks, vol. 11, no. 3, pp. 690-696, May 2000.
#
# [6] http://www.nickgillian.com/wiki/pmwiki.php/GRT/AdaBoost
#
# [7] https://codesachin.wordpress.com/tag/adaboost/
#
# [8] https://elitedatascience.com/machine-learning-algorithms
#
# [9] https://link.springer.com/chapter/10.1007/978-3-319-26762-3_27 - <NAME>., <NAME>. (2015) Modelling the Quality of Attributes in Wikipedia Infoboxes. In: Abr<NAME>. (eds) Business Information Systems Workshops. BIS 2015. Lecture Notes in Business Information Processing, vol 228. Springer, Cham
#
# [10] https://link.springer.com/chapter/10.1007/978-1-84628-814-2_82 - <NAME>., <NAME>., <NAME>. (2006) Application of Random Forest Algorithm in Machine Fault Diagnosis. In: <NAME>., <NAME>., <NAME>., <NAME>., <NAME>. (eds) Engineering Asset Management. Springer, London
# ### Implementation - Creating a Training and Predicting Pipeline
# To properly evaluate the performance of each model you've chosen, it's important that you create a training and predicting pipeline that allows you to quickly and effectively train models using various sizes of training data and perform predictions on the testing data. Your implementation here will be used in the following section.
# In the code block below, you will need to implement the following:
# - Import `fbeta_score` and `accuracy_score` from [`sklearn.metrics`](http://scikit-learn.org/stable/modules/classes.html#sklearn-metrics-metrics).
# - Fit the learner to the sampled training data and record the training time.
# - Perform predictions on the test data `X_test`, and also on the first 300 training points `X_train[:300]`.
# - Record the total prediction time.
# - Calculate the accuracy score for both the training subset and testing set.
# - Calculate the F-score for both the training subset and testing set.
# - Make sure that you set the `beta` parameter!
# TODO: Import two metrics from sklearn - fbeta_score and accuracy_score
from sklearn.metrics import accuracy_score
from sklearn.metrics import fbeta_score
def train_predict(learner, sample_size, X_train, y_train, X_test, y_test):
'''
inputs:
- learner: the learning algorithm to be trained and predicted on
- sample_size: the size of samples (number) to be drawn from training set
- X_train: features training set
- y_train: income training set
- X_test: features testing set
- y_test: income testing set
'''
results = {}
# TODO: Fit the learner to the training data using slicing with 'sample_size' using .fit(training_features[:], training_labels[:])
start = time() # Get start time
learner = learner.fit(X_train[:sample_size], y_train[:sample_size])
end = time() # Get end time
# TODO: Calculate the training time
results['train_time'] = end-start
# TODO: Get the predictions on the test set(X_test),
# then get predictions on the first 300 training samples(X_train) using .predict()
start = time() # Get start time
predictions_test = learner.predict(X_test)
predictions_train = learner.predict(X_train[:300])
end = time() # Get end time
# TODO: Calculate the total prediction time
results['pred_time'] = end-start
# TODO: Compute accuracy on the first 300 training samples which is y_train[:300]
results['acc_train'] = accuracy_score(y_train[:300], predictions_train)
# TODO: Compute accuracy on test set using accuracy_score()
results['acc_test'] = accuracy_score(y_test, predictions_test)
# TODO: Compute F-score on the the first 300 tra}ining samples using fbeta_score()
results['f_train'] = fbeta_score(y_train[:300], predictions_train, beta=0.5)
# TODO: Compute F-score on the test set which is y_test
results['f_test'] = fbeta_score(y_test, predictions_test, beta=0.5)
# Success
print("{} trained on {} samples.".format(learner.__class__.__name__, sample_size))
# Return the results
return results
# ### Implementation: Initial Model Evaluation
# In the code cell, you will need to implement the following:
# - Import the three supervised learning models you've discussed in the previous section.
# - Initialize the three models and store them in `'clf_A'`, `'clf_B'`, and `'clf_C'`.
# - Use a `'random_state'` for each model you use, if provided.
# - **Note:** Use the default settings for each model — you will tune one specific model in a later section.
# - Calculate the number of records equal to 1%, 10%, and 100% of the training data.
# - Store those values in `'samples_1'`, `'samples_10'`, and `'samples_100'` respectively.
#
# **Note:** Depending on which algorithms you chose, the following implementation may take some time to run!
# +
# TODO: Import the three supervised learning models from sklearn
from sklearn.ensemble import AdaBoostClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.naive_bayes import GaussianNB
# TODO: Initialize the three models
clf_A = GaussianNB()
clf_B = AdaBoostClassifier(random_state=13)
clf_C = RandomForestClassifier(random_state=13)
# TODO: Calculate the number of samples for 1%, 10%, and 100% of the training data
# HINT: samples_100 is the entire training set i.e. len(y_train)
# HINT: samples_10 is 10% of samples_100 (ensure to set the count of the values to be `int` and not `float`)
# HINT: samples_1 is 1% of samples_100 (ensure to set the count of the values to be `int` and not `float`)
samples_100 = len(y_train)
samples_10 = int(samples_100/10)
samples_1 = int(samples_10/10)
# Collect results on the learners
results = {}
for clf in [clf_A, clf_B, clf_C]:
clf_name = clf.__class__.__name__
results[clf_name] = {}
for i, samples in enumerate([samples_1, samples_10, samples_100]):
results[clf_name][i] = \
train_predict(clf, samples, X_train, y_train, X_test, y_test)
# Run metrics visualization for the three supervised learning models chosen
vs.evaluate(results, accuracy, fscore)
# -
# ----
# ## Improving Results
# In this final section, you will choose from the three supervised learning models the *best* model to use on the student data. You will then perform a grid search optimization for the model over the entire training set (`X_train` and `y_train`) by tuning at least one parameter to improve upon the untuned model's F-score.
# ### Question 3 - Choosing the Best Model
#
# * Based on the evaluation you performed earlier, in one to two paragraphs, explain to *CharityML* which of the three models you believe to be most appropriate for the task of identifying individuals that make more than \$50,000.
#
# ** HINT: **
# Look at the graph at the bottom left from the cell above(the visualization created by `vs.evaluate(results, accuracy, fscore)`) and check the F score for the testing set when 100% of the training set is used. Which model has the highest score? Your answer should include discussion of the:
# * metrics - F score on the testing when 100% of the training data is used,
# * prediction/training time
# * the algorithm's suitability for the data.
# **Answer: **
#
# The most appropriate model to identify individuals that make more than $50,000 is AdaBoost due the F-score of the model. The training and prediction times are higher than other models, but still a short time and it fits well on the dataset.
# ### Question 4 - Describing the Model in Layman's Terms
#
# * In one to two paragraphs, explain to *CharityML*, in layman's terms, how the final model chosen is supposed to work. Be sure that you are describing the major qualities of the model, such as how the model is trained and how the model makes a prediction. Avoid using advanced mathematical jargon, such as describing equations.
#
# ** HINT: **
#
# When explaining your model, if using external resources please include all citations.
# **Answer: **
#
# Adaboost use simpler learners (weak learners) emsembled in a more robust learner (strong learner). The _adaptative_ in the Adaboost name is in the sense that subsequent weak learners are tweaked in favor o f those instances misclassified by previous classifiers. In other words, we ensemble weak learners that attempts to correct the errors from the previous classifiers.
#
# The predictions are made by calculating the weighted average of the weak classifiers. For a new input instance, each weak learner calculates a predicted values as either +1.0 or -1.0, then the prediction is taken as a the sum of the wieghted predictions.
# ### Implementation: Model Tuning
# Fine tune the chosen model. Use grid search (`GridSearchCV`) with at least one important parameter tuned with at least 3 different values. You will need to use the entire training set for this. In the code cell below, you will need to implement the following:
# - Import [`sklearn.grid_search.GridSearchCV`](http://scikit-learn.org/0.17/modules/generated/sklearn.grid_search.GridSearchCV.html) and [`sklearn.metrics.make_scorer`](http://scikit-learn.org/stable/modules/generated/sklearn.metrics.make_scorer.html).
# - Initialize the classifier you've chosen and store it in `clf`.
# - Set a `random_state` if one is available to the same state you set before.
# - Create a dictionary of parameters you wish to tune for the chosen model.
# - Example: `parameters = {'parameter' : [list of values]}`.
# - **Note:** Avoid tuning the `max_features` parameter of your learner if that parameter is available!
# - Use `make_scorer` to create an `fbeta_score` scoring object (with $\beta = 0.5$).
# - Perform grid search on the classifier `clf` using the `'scorer'`, and store it in `grid_obj`.
# - Fit the grid search object to the training data (`X_train`, `y_train`), and store it in `grid_fit`.
#
# **Note:** Depending on the algorithm chosen and the parameter list, the following implementation may take some time to run!
# +
# TODO: Import 'GridSearchCV', 'make_scorer', and any other necessary libraries
from sklearn.tree import DecisionTreeClassifier
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import make_scorer
# TODO: Initialize the classifier
clf = AdaBoostClassifier(base_estimator=DecisionTreeClassifier(), random_state=13)
# TODO: Create the parameters list you wish to tune, using a dictionary if needed.
# HINT: parameters = {'parameter_1': [value1, value2], 'parameter_2': [value1, value2]}
parameters = {
'learning_rate': [0.1, 0.25, 0.5],
'n_estimators': np.arange(30, 60, 10),
'base_estimator__min_samples_split': np.arange(2, 8, 2),
'base_estimator__max_depth': np.arange(1, 5, 1)
}
# TODO: Make an fbeta_score scoring object using make_scorer()
scorer = make_scorer(fbeta_score, beta=0.5)
# TODO: Perform grid search on the classifier using 'scorer' as the scoring method using GridSearchCV()
grid_obj = GridSearchCV(clf, parameters, scoring=scorer)
# TODO: Fit the grid search object to the training data and find the optimal parameters using fit()
grid_fit = grid_obj.fit(X_train, y_train)
# Get the estimator
best_clf = grid_fit.best_estimator_
# Make predictions using the unoptimized and model
predictions = (clf.fit(X_train, y_train)).predict(X_test)
best_predictions = best_clf.predict(X_test)
# Report the before-and-afterscores
print("Unoptimized model\n------")
print("Accuracy score on testing data: {:.4f}".format(accuracy_score(y_test, predictions)))
print("F-score on testing data: {:.4f}".format(fbeta_score(y_test, predictions, beta = 0.5)))
print("\nOptimized Model\n------")
print("Final accuracy score on the testing data: {:.4f}".format(accuracy_score(y_test, best_predictions)))
print("Final F-score on the testing data: {:.4f}".format(fbeta_score(y_test, best_predictions, beta = 0.5)))
# -
# ### Question 5 - Final Model Evaluation
#
# * What is your optimized model's accuracy and F-score on the testing data?
# * Are these scores better or worse than the unoptimized model?
# * How do the results from your optimized model compare to the naive predictor benchmarks you found earlier in **Question 1**?_
#
# **Note:** Fill in the table below with your results, and then provide discussion in the **Answer** box.
# #### Results:
#
# | Metric | Unoptimized Model | Optimized Model | Naive Predictor |
# | :------------: | :---------------: | :-------------: | :-------------: |
# | Accuracy Score | 0.8367 | 0.8615 | 0.2478 |
# | F-score | 0.6679 | 0.7416 | 0.2917 |
# **Answer: **
#
# - The optimized model accuracy and F-score on the testing data are 0.8615 and 0.7416.
# - The scores are **2.964% and 11.0345% better** that the unoptimized model.
# - The scores are **247.6594% and 154.2338% better** that the naive predictor.
# ----
# ## Feature Importance
#
# An important task when performing supervised learning on a dataset like the census data we study here is determining which features provide the most predictive power. By focusing on the relationship between only a few crucial features and the target label we simplify our understanding of the phenomenon, which is most always a useful thing to do. In the case of this project, that means we wish to identify a small number of features that most strongly predict whether an individual makes at most or more than \$50,000.
#
# Choose a scikit-learn classifier (e.g., adaboost, random forests) that has a `feature_importance_` attribute, which is a function that ranks the importance of features according to the chosen classifier. In the next python cell fit this classifier to training set and use this attribute to determine the top 5 most important features for the census dataset.
# ### Question 6 - Feature Relevance Observation
# When **Exploring the Data**, it was shown there are thirteen available features for each individual on record in the census data. Of these thirteen records, which five features do you believe to be most important for prediction, and in what order would you rank them and why?
# **Answer:**
#
# 1. **age**: An older person is more likely to have a high income and savings and a younger person didn't have enough time to build a carreer, have a PhD. So I think thats the most important feature because people tend to get richer as the life goes by.
# 2. **hours-per-week**: Usually people who work more hours per week receive more money. Half-time jobs are unlikely to pay well.
# 3. **capital-loss**: How much a person lost in the given year will influence on the decision of make a donation, the more the person lost, more likely to not become a donator.
# 4. **education-num**: This feature follow the Age logic. People who are high educated tend to earn more money, but its not as good as the age feature to measure this.
# 5. **occupation**: The occupation can tell a lot about the people earnings and his behavior. Some occupations tend to have a more "human" side and this can influence on the model's decision too.
# ### Implementation - Extracting Feature Importance
# Choose a `scikit-learn` supervised learning algorithm that has a `feature_importance_` attribute availble for it. This attribute is a function that ranks the importance of each feature when making predictions based on the chosen algorithm.
#
# In the code cell below, you will need to implement the following:
# - Import a supervised learning model from sklearn if it is different from the three used earlier.
# - Train the supervised model on the entire training set.
# - Extract the feature importances using `'.feature_importances_'`.
# +
# TODO: Import a supervised learning model that has 'feature_importances_'
# from sklearn.ensemble import AdaBoostClassifier # already imported
# TODO: Train the supervised model on the training set using .fit(X_train, y_train)
model = AdaBoostClassifier().fit(X_train, y_train)
# TODO: Extract the feature importances using .feature_importances_
importances = model.feature_importances_
# Plot
vs.feature_plot(importances, X_train, y_train)
# -
# ### Question 7 - Extracting Feature Importance
#
# Observe the visualization created above which displays the five most relevant features for predicting if an individual makes at most or above \$50,000.
# * How do these five features compare to the five features you discussed in **Question 6**?
# * If you were close to the same answer, how does this visualization confirm your thoughts?
# * If you were not close, why do you think these features are more relevant?
# **Answer:**
#
# It was very close from my guess. The only one that is different is capital-gain, in mine I put occupation. The visualization confirm most of my thoughts that what would be the most important features. The order is different from what I thought, but makes sense that capital loss is more important even than age, as people with a big capital-loss most likely are not making great money and following this path is natural that capital-gain is such a important feature too.
# ### Feature Selection
# How does a model perform if we only use a subset of all the available features in the data? With less features required to train, the expectation is that training and prediction time is much lower — at the cost of performance metrics. From the visualization above, we see that the top five most important features contribute more than half of the importance of **all** features present in the data. This hints that we can attempt to *reduce the feature space* and simplify the information required for the model to learn. The code cell below will use the same optimized model you found earlier, and train it on the same training set *with only the top five important features*.
# +
# Import functionality for cloning a model
from sklearn.base import clone
# Reduce the feature space
X_train_reduced = X_train[X_train.columns.values[(np.argsort(importances)[::-1])[:5]]]
X_test_reduced = X_test[X_test.columns.values[(np.argsort(importances)[::-1])[:5]]]
# Train on the "best" model found from grid search earlier
clf = (clone(best_clf)).fit(X_train_reduced, y_train)
# Make new predictions
reduced_predictions = clf.predict(X_test_reduced)
# Report scores from the final model using both versions of data
print("Final Model trained on full data\n------")
print("Accuracy on testing data: {:.4f}".format(accuracy_score(y_test, best_predictions)))
print("F-score on testing data: {:.4f}".format(fbeta_score(y_test, best_predictions, beta = 0.5)))
print("\nFinal Model trained on reduced data\n------")
print("Accuracy on testing data: {:.4f}".format(accuracy_score(y_test, reduced_predictions)))
print("F-score on testing data: {:.4f}".format(fbeta_score(y_test, reduced_predictions, beta = 0.5)))
# -
# ### Question 8 - Effects of Feature Selection
#
# * How does the final model's F-score and accuracy score on the reduced data using only five features compare to those same scores when all features are used?
# * If training time was a factor, would you consider using the reduced data as your training set?
# **Answer:**
#
# | Metric | Optimized Model | Reduced data |
# | :------------: | :-------------: | :-------------: |
# | Accuracy Score | 0.8615 | 0.8399 |
# | F-score | 0.7416 | 0.6969 |
#
# The Accuracy and F-score of the model on the reduced data was **2.5072% and 6.0275% worse** than the final model trained with all features. If training time was a factor the model using only five features is a way to go, because the lost in the accuracy and f-score was low and the training time was a lot faster.
# > **Note**: Once you have completed all of the code implementations and successfully answered each question above, you may finalize your work by exporting the iPython Notebook as an HTML document. You can do this by using the menu above and navigating to
# **File -> Download as -> HTML (.html)**. Include the finished document along with this notebook as your submission.
|
finding_donors/finding_donors.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.4.2
# language: julia
# name: julia-1.4
# ---
# # Loops
# Topics:
# 1. `while` loops
# 2. `for` loops
# ## while loops
#
# The syntax for a `while` is
# ```julia
# while *condition*
# *loop body*
# end
# ```
# For example, we could use `while` to count or to iterate over an array.
n = 0
while n < 10
global n += 1
println(n)
end
n
println(n)
myfriends = ["Ted", "Robyn", "Barney", "Lily", "marshall"]
i = 1
while i <= length(myfriends)
friend = myfriends[i]
println("Hi $friend, it's great to see you!")
global i += 1
end
# ## for loops
#
# The syntax for a `for` loop is
#
# ```julia
# for *var* in *loop iterable*
# *loop body*
# end
# ```
# We cound use a for loop to generate the same results as either of the examples above:
for n in 1:10
println(n)
end
myfriends = ["Ted", "Robyn", "Barney", "Lily", "Marshall"]
for friend in myfriends
println("Hi $friend, it's great to see you!")
end
# Now let's use `for` loops to create some addition tables, where the value of every entry is the sum of its orw and column indices. <br>
#
# Note that we iterate over this array via column-major loops in order to get the best performance. More information about fast indexing of multidimensional arrays inside nested loops can be found at https://docs.julialang.org/en/v1/manual/performance-tips/#Access-arrays-in-memory-order,-along-columns-1
# First, we initialize an array with zeros.
# +
m, n = 5, 5
A = fill(0, (m, n))
for j in 1:n
for i in 1:m
A[i, j] = i+j
end
end
println(A)
# -
# Here's some syntactic sugar for the same nested `for` loop
# +
B = fill(0, (m, n))
for j in 1:n, i in 1:m
B[i, j] = i + j
end
println(B)
# -
# The more "Julia" way to create this addition table would have been with an *array comprehension*.
C = [i + j for i in 1:m, j in 1:n]
# ### Exercises
# #### 4.1
# Loop over integers between 1 and 100 and print their squares.
# #### 4.2
# Add to the code above a bit to create a dictionary, `squres` that holds integers and their squares as key, value pairs such that
# ```julia
# squares[10] == 100
# ```
# #### 4.3
# Use an array comprehension to create an array `squares_arr` that stores the squares for all integers between 1 and 100.
# ### Solutions
# #### 4.1
for i in 1:100
println(i * i)
end
# #### 4.2
squares = Dict{Integer, Integer}()
for i in 1:100
squares[i] = i * i
end
@assert squares[10] == 100
@assert squares[11] == 121
# #### 4.3
squares_arr = [i * i for i in 1:100]
@assert length(squares_arr) == 100
@assert sum(squares_arr) == 338350
|
Loops.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Final grade
# ___
#
# Using exercises spreadsheets from Designing and Analyzing Language Tests by Oxford. The purpose of this notebook is to create a gradebook - that is, a record of every student's grade n every quiz, test, or assignment in the course, and their end-of-course grade.
#
# <br>
#
# #### General Setup
# ___
# import libraries
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# styling for plots
plt.style.use('seaborn-whitegrid')
plt.rcParams['figure.figsize'] = (7,5)
# <br>
#
# #### Load the data
# ___
# load the data set and check the first 5 rows
test_res = pd.read_excel('Data/overall_results.xlsx')
test_res.head()
test_res.tail()
# data set info
test_res.info()
# The dataset shows test results for 32 students. There are 8 quizzes, 1 midterm, 1 project and 1 final test. The dataset also has some missing values. We will ignore it and leave them blank since changing them to 0.0 may reduce the overall grade.
# check stats
test_res.describe()
# <br>
#
# #### EDA
# ___
#
# The course grades are based on the average quiz grade (30%), the course project (30%), the midterm exam (20%), and the final exam (20%).
# calculate the mean for quizes for each student
test_res['Quiz Avg'] = np.round(test_res.drop(['Name', 'Midterm', 'Project', 'Final'], axis=1).mean(axis=1), 2)
test_res.head()
# calculate weighted average
test_res['Wtd. Avg'] = np.round(test_res['Quiz Avg'] * 0.3 + test_res['Midterm'] * 0.2 + test_res['Project'] * 0.3 + test_res['Final'] * 0.2, 2)
test_res.head()
# calculate mean for each column
averages = pd.DataFrame({'Average': np.round(test_res.loc[:, test_res.columns != 'Name'].mean(), 2)}).T
averages
# We can now see that the class average for the aveall course (the average of the weighted averages) is around 78.0.
# We can curve the grades - in that if the class average is below 85%, points will be added to every the student's final score in order to raise the average to this target.
# calculate the curve
curve = 85 - averages.values[:,-1][0]
np.around(curve)
# The result is the number of points added to each student's course grade.
# calculate the curved avg. only apply if curve is above 0
if curve > 0:
test_res['Curved Avg'] = test_res['Wtd. Avg'] + curve
test_res.head(6)
else:
test_res['Curved Avg'] = test_res['Wtd. Avg']
# calculate the rounded avg
test_res['Rounded Avg'] = np.around(test_res['Curved Avg']).astype(int)
test_res.head()
# <br>
#
# #### Grade lookup
# ___
# load grade dataset
grades = pd.read_excel('Data/grades.xlsx')
grades
# create a list of ranges from the min and max columns
val = grades.loc[:,'Min':'Max'].apply(tuple, 1).tolist()
val
# turn into an interval
indx = pd.IntervalIndex.from_tuples(val, closed='both')
indx
# add a column to the test res data set by looking up the grade
test_res['Grade'] = grades.loc[indx.get_indexer(test_res['Rounded Avg']), 'Grade'].values
test_res.head()
# <br>
#
# #### Organising dataframes and saving them as Excel
# ___
# calculate mean for each column
averages = pd.DataFrame({'Average': np.round(test_res.drop(['Name', 'Grade'], axis=1).mean(), 2)}).T
averages
# calculate frequency of each grade
freq = pd.DataFrame(test_res.Grade.value_counts())
freq['Frequency'] = freq.Grade
freq.drop('Grade', axis=1, inplace=True)
freq.index.name = 'Grade'
freq
# +
# frequency
fig, ax = plt.subplots()
fig.suptitle('Frequency of Grades', y=1.02, weight='bold', fontsize=13)
ax = sns.barplot(x=freq.index, y='Frequency', data=freq, palette="Blues_d")
# save the plot
plt.savefig('Data/frequency_of_grades.png', bbox_inches='tight');
# +
# histograms and frequency polygon
fig, [ax0, ax1] = plt.subplots(1,2, figsize=(14,5))
fig.suptitle('Distribution of Weighted Averages', y=1.02, weight='bold', fontsize=13)
# weighted scores hist
ax0.hist(test_res['Wtd. Avg'], bins=40)
ax0.set(title='Histogram of Weighted Averages',
xlabel='Weighted Average',
ylabel='Frequency')
# weighted scores freq polygon
ax1.plot(test_res['Wtd. Avg'],marker='.', linestyle='solid', markersize=20, markerfacecolor='lightyellow')
ax1.set(title='Frequency Polygon for Weighted Averages',
xlabel='Weighted Average',
ylabel='Frequency')
# save the plot
plt.savefig('Data/distribution_of_weighted_averages.png', bbox_inches='tight');
# -
# <br>
#
# #### Saving results to Excel
# ___
# sort the table by weighted avg in descending order
sorted_scores = test_res.sort_values('Wtd. Avg', ascending=False, kind='stable')
sorted_scores.head()
# +
# write and save 3 dataframes to the excel file
writer = pd.ExcelWriter('Data/final_exam_analysis.xlsx', engine='xlsxwriter')
sorted_scores.to_excel(writer, index = False)
averages.to_excel(writer, startrow=len(test_res)+1, index=True, header=False)
freq.to_excel(writer, startcol=len(test_res.columns)+1, index=True)
# insert the image into the worksheet
workbook = writer.book
worksheet = writer.sheets['Sheet1']
worksheet.insert_image('A38', 'Data/distribution_of_weighted_averages.png')
worksheet.insert_image('V2', 'Data/frequency_of_grades.png')
# styling
column_settings = [{'header': column} for column in sorted_scores.columns]
(max_row, max_col) = sorted_scores.shape
worksheet.add_table(0, 0, max_row, max_col - 1, {'columns': column_settings})
writer.save()
# -
# <br>
#
# ___
# #### End
|
Data analytics/Language tests/Final grade.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Setup
# #### Imports
from PW_explorer.load_worlds import load_worlds
from PW_explorer.run_clingo import run_clingo
from PW_explorer.visualize import PWEVisualization
from PW_explorer.helper import pw_slicer, rel_slicer, rel_name_remapper
from PW_explorer.export import PWEExport
import pandas as pd
import os
from misc import *
import networkx as nx
from nxpd import draw
import nxpd
from nxpd import nxpdParams
nxpdParams['show'] = 'ipynb'
# %load_ext PWE_NB_Extension
# #### Helper Functions
def save_query_figs(query_analysis, folder='3hop_all_patterns_new'):
fnames = {}
for pw_id in range(1, len(query_analysis['pw_objs'])+1):
pw_rel_dfs, pw_objs = pw_slicer(query_analysis['pw_rel_dfs'], query_analysis['pw_objs'], [pw_id])
pw_obj = pw_objs[0]
node_fig_g = get_pattern_graph3(pw_rel_dfs, pw_obj, silent=True,
q_head='binary-node-highlight',
chain_eq_node_labels=False)
q_str = print_rewritten_query_string(pw_rel_dfs, html=False, display_q=False, include_ineqls=False)
q_str = q_str.replace('thop', 'q')
node_fig_g.graph['label'] = q_str
node_fig_g.graph['labelloc'] = 't' # options 't', 'b'
fname = '{}.{}'.format('{}/pw_{}'.format(folder, pw_id), 'png')
draw(node_fig_g, filename=fname, show=False)
fnames[pw_id] = fname
return fnames
# #### Supplementary ASP Encodings
# +
# %%clingo --donot-display_input --donot-run -lci query_analysis_suppl
% schema ruleOccTrue(ATOM, OCC)
ruleOccTrue(R, OC) :- ruleOcc(R, OC).
% schema ruleHTrue(HEAD)
ruleHTrue(H) :- ruleH(H).
% schema newVar(VAR, DOMAIN)
newVar(VAR, TYPE) :- var(VAR, TYPE), not eqOrd(_,VAR,TYPE).
% schema eqOrdMinimal(VAR1, VAR2, DOMAIN)
eqOrdMinimal(A,B,TYPE) :- eqOrd(A,B,TYPE), newVar(A,TYPE).
% schema neqOrd(VAR1, VAR2, DOMAIN)
neqOrd(A,B,TYPE) :- newVar(A,TYPE), newVar(B,TYPE), A<B.
% New inferred edges
% schema e(NODE1, NODE2)
e(V1,V2) :- newArc(V1, 1, "e", OCC), newArc(V2, 2, "e", OCC).
% schema e(OCC, NODE1, NODE2)
e(OCC,V1,V2) :- newArc(V1, 1, "e", OCC), newArc(V2, 2, "e", OCC).
# %graphviz graph graph_type=directed rankdir=LR
# %graphviz edge newArc(HEAD, _, TAIL) label=$2
# %graphviz node ruleOccTrue(N) color=green
#show ruleOccTrue/2.
#show ruleHTrue/1.
#show var/2.
#show newVar/2.
#show ruleOcc/2.
#show ruleH/1.
#show eqOrd/3.
#show neqOrd/3.
#show eqOrdMinimal/3.
#show arc/4.
#show newArc/4.
#show hArc/3.
#show newHArc/3.
#show e/2.
#show e/3.
# -
# ### Query Analysis
# Say we want to statically analyze the well-studied 3hop/2 query:
#
# __thop(X,Y) :- e(X,Z1), e(Z1,Z2), e(Z2,Y).__
#
# To do this, we first encode the query as a incidence graph as shown below:
# +
# %%clingo --donot-display_input --donot-run -lci thop_query_desc
% Q: thop(X,Y) :- e(X,Z1), e(Z1,Z2), e(Z2,Y).
% Declare the variables appearing in the query:
% schema var(VAR, DOMAIN).
var("X", node).
var("Y", node).
var("Z1", node).
var("Z2", node).
% Declare the logical atoms in the body of the query:
% schema ruleOcc(ATOM, OCC).
ruleOcc("e", 1..3).
% Declare the logical atom in the head of the query:
% schema ruleH(HEAD).
ruleH("thop").
% Declare the edges of the incidence graph:
% schema arc(VAR, POS, ATOM, OCC).
arc("X", 1, "e", 1). % This means X is in the 1st position in the 1st "e" rule-atom
arc("Z1", 2, "e", 1). % ...
arc("Z1", 1, "e", 2). % ...
arc("Z2", 2, "e", 2). % This means Z2 is in the 2nd position in the 2nd "e" rule-atom
arc("Z2", 1, "e", 3). % and so on....
arc("Y", 2, "e", 3).
% Declare the edges from the "query-head" node
% Use the rules below to change the rule head.
% schema hArc(VAR, POS, HEAD).
hArc("X", 1, "thop") :- ruleH("thop").
hArc("Y", 2, "thop") :- ruleH("thop").
# -
# Next we encode the query analysis code that uses a query description such as the one above.
#
# Within this analysis, we create Bell-Number(n) (i.e. B<sub>n</sub>) many partitions where n is the number of distinct variable in the query.
#
# Then we recreate a new incidence graph where the we substitute each disjoint group in the partition with a representative variable (for simplicity, we use the lexicographically smallest variable).
# +
# %%clingo --donot-display_input -lci query_analysis_enc --donot-run
% Create variable partitions
% schema eq(VAR1, VAR2, DOMAIN)
eq(A,B,TYPE) ; not eq(A,B,TYPE) :- var(A, TYPE), var(B, TYPE), A!=B.
eq(A,B,TYPE) :- eq(B,A,TYPE).
eq(A,B,TYPE) :- eq(A,C,TYPE), eq(C,B,TYPE), A!=B, B!=C, A!=C.
% Order the individual partitions
% schema eqOrd(VAR1, VAR2, DOMAIN)
eqOrd(A,B,TYPE) :- eq(A,B,TYPE), A<B.
% Find the new arcs, getting rid of the redundant variables as per the orderings
% schema newArc(VAR, POS, ATOM, OCC)
newArc(VAR, POS, ATOM, OCC) :- arc(VAR_ORIG, POS, ATOM, OCC), eqOrd(VAR,VAR_ORIG,TYPE), not eqOrd(_,VAR,TYPE).
newArc(VAR, POS, ATOM, OCC) :- arc(VAR, POS, ATOM, OCC), not eqOrd(_,VAR,_).
% schema newHArc(VAR, POS, HEAD)
newHArc(VAR, POS, H) :- hArc(VAR_ORIG,POS,H), eqOrd(VAR,VAR_ORIG,TYPE), not eqOrd(_,VAR,TYPE).
newHArc(VAR, POS, H) :- hArc(VAR, POS, H), not eqOrd(_,VAR,_).
# -
# With these two encodings, we can create the specialized queries that make up the original query.
#
# For clarity, let's project out the resultant e/2 and thop/2 relation that make up the query:
# +
# %%clingo --donot-display_input -l query_analysis_enc thop_query_desc
e(V1,V2) :- newArc(V1, 1, "e", OCC), newArc(V2, 2, "e", OCC).
thop(V1,V2) :- newHArc(V1,1,"thop"), newHArc(V2,2,"thop").
#show e/2.
#show thop/2.
# -
# As we can see, we get B<sub>4</sub> (= 15) PWs. The outputs above showcase a canonical database that would satisfy the specialized query that these PWs represent.
#
# First we load these PWs in using the PWE Tool.
# %clingo --donot-display_input --donot-display_output -l query_analysis_enc thop_query_desc query_analysis_suppl -exp query_analysis
query_analysis['pw_rel_dfs'], query_analysis['rel_schema'], query_analysis['pw_objs'] =load_worlds(query_analysis['asp_soln'],
query_analysis['meta_data'])
# With some simple python and ASP magic, we can create these specialized queries, their canonical databases and visualize the graph patterns these canonical databases represent.
for i in range(1, len(query_analysis['pw_objs'])+1):
print('-'*60)
print("\nPW:", i)
pw_rel_dfs, pw_objs = pw_slicer(query_analysis['pw_rel_dfs'], query_analysis['pw_objs'], [i])
pw_obj = pw_objs[0]
print("\nPartitioning Description:")
print_fancy_rewrite(pw_rel_dfs)
print("Specialized Query:")
print_explicit_rewritten_query_string(pw_rel_dfs, chain_eq=False)
print("Canonical DB:")
pw_edge_facts = get_edge_facts(pw_obj)
pw_head_facts = get_query_head_facts(pw_obj)
print(" ".join(pw_edge_facts+pw_head_facts))
g = get_pattern_graph3(pw_rel_dfs, pw_obj, silent=True, q_head='hyperedge')
# draw(g, filename='{}.{}'.format('3hop_all_patterns_new/pw_{}'.format(i), 'png'), show=False)
display(draw(g))
print('-'*60)
# Above, we can see for each PW, the variable partitions (in blue), followed by the specialized query the PW represents, followed by a textual and visual representation of the canonical database of the specialized query.
#
# These visual representations of the canonical databases are what we call ___graph-patterns___.
#
# One outstanding issue is that these specialized queries/patterns aren't disjoint, _yet_, i.e. some of them are equivalent. To find the set of disjoint set of these specialized queries/patterns, we check for isomorphisms between these canonical databases. If there exists an isomorphism between two canonical databases (and hence the graph patterns), the two specialized queries (and the graph patterns) are equivalent, and can be grouped together.
#
# Below is the encoding that checks if there exists such an isomorphism for two such canonical databases:
# +
# %%clingo --donot-display_input -lci qep_eq_check --donot-run
% e(X,Y) :- e(Y,X). --> only if undirected
gnode1(X) :- e1(X,_).
gnode1(X) :- e1(_,X).
gnode2(X) :- e2(X,_).
gnode2(X) :- e2(_,X).
vmap(X,Y) ; not vmap(X,Y) :- gnode1(X), gnode2(Y).
:- vmap(X1,Y1), vmap(X2,Y2), e1(X1,X2), not e2(Y1,Y2).
:- vmap(X1,Y1), vmap(X2,Y2), not e1(X1,X2), e2(Y1,Y2).
% 3hop Query
:- vmap(X1,Y1), vmap(X2,Y2), thop1(X1,X2), not thop2(Y1,Y2).
:- vmap(X1,Y1), vmap(X2,Y2), not thop1(X1,X2), thop2(Y1,Y2).
:- gnode1(X), #count {Y: vmap(X,Y)} != 1.
:- gnode2(Y), #count {X: vmap(X,Y)} != 1.
#show.
# -
# We can now use PWE to compute the disjoint groups efficiently using the above encoding:
sets = get_equivalent_sets(query_analysis['pw_objs'], lambda pw1, pw2: are_equivalent_patterns(pw1,pw2,qep_eq_check))
for s in sets:
print({pw.pw_id for pw in s})
len(sets)
# As we can see, there are 14 disjoint specialized queries (as opposed to 15), since specialized queries 13 and 14 are equivalent. Let's take a look at their graph patterns again to see why that's the case:
for i in [13,14]:
print('-'*60)
print("\nPW:", i)
pw_rel_dfs, pw_objs = pw_slicer(query_analysis['pw_rel_dfs'], query_analysis['pw_objs'], [i])
pw_obj = pw_objs[0]
print("Canonical DB:")
pw_edge_facts = get_edge_facts(pw_obj)
pw_head_facts = get_query_head_facts(pw_obj)
print(" ".join(pw_edge_facts+pw_head_facts))
g = get_pattern_graph3(pw_rel_dfs, pw_obj, silent=True, q_head='binary-edge')
display(draw(g))
print('-'*60)
# As we can verify, the two are indeed equivalent. We can swap "Z1" with "Z2" in the first one or vice-versa in the second one, to get the other canonical database, meaning there exists an isomorphism between the two.
# ### Building a hierarchy of _semi_-specialized queries
#
# In the specialized queries above, we explicitly encode the equalities and in/dis-equalities between the variables in the query. Say we only encode the equalities but not the in/dis-equalities. This gives us a set of _semi_-specialized queries which can be arranged in a hierarchy based on query containment relations. We build these queries and their hierarchy below.
# Below is an ASP encoding to check for query containment between two queries Q1 and Q2 (is Q2 contained in Q1?) based on their canonical databases:
# +
# %%clingo --donot-display_input -lci qep_containment_check --donot-run
% Project out the nodes of the two queries
gnode1(X) :- e1(X,_).
gnode1(X) :- e1(_,X).
gnode2(X) :- e2(X,_).
gnode2(X) :- e2(_,X).
% Generate a mapping from variables of Q1 to variables of Q2:
vmap(X,Y) ; not vmap(X,Y) :- gnode1(X), gnode2(Y).
% Test that for every edge e1(X1,Y1) and thop(X1,Y1) in Q1 there is an edge e2(Y1,Y2) and thop(Y1,Y2) in Q2 s.t.
% X1 maps to Y1 and X2 maps to Y2
:- vmap(X1,Y1), vmap(X2,Y2), e1(X1,X2), not e2(Y1,Y2).
:- vmap(X1,Y1), vmap(X2,Y2), thop1(X1,X2), not thop2(Y1,Y2).
% Ensure that it is a many-to-one/one-to-one mapping
:- gnode1(X), #count {Y: vmap(X,Y)} != 1.
#show.
# -
# Function to test whether query Q1 (represented by pw1) contains query Q2 (represented by pw2)
# using the above encoding
def q1_contains_q2(pw1, pw2):
return are_equivalent_patterns(pw1, pw2, qep_containment_check)
# Save figures for each PW (to build the hierarchy)
query_fig_fnames = save_query_figs(query_analysis, folder='3hop_all_patterns_new')
# Next, we create the hierarchy of the queries:
g = nx.DiGraph()
for pw_id in range(1, len(query_analysis['pw_objs'])+1):
fname = query_fig_fnames[pw_id]
g.add_node(pw_id, image=os.path.abspath(fname),
shape='box', fixedsize='true', imagescale='true', size=2, label=" ")
# Let's add edges based on query containment between all the queries:
# +
for pw_id1 in range(1, len(query_analysis['pw_objs'])+1):
for pw_id2 in range(1, len(query_analysis['pw_objs'])+1):
if pw_id1 == pw_id2:
continue
if q1_contains_q2(query_analysis['pw_objs'][pw_id1-1], query_analysis['pw_objs'][pw_id2-1]):
g.add_edge(pw_id1, pw_id2, dir='back')
draw(g)
# -
# As we can see above, PW-13, PW-14 and PW-15 have bi-directional containment, meaning they are equivalent. Recall that we already knew this from the isomorphism test earlier.
#
# We can hence contract PW-13, PW-14 and PW-15 into a single node:
# +
keep_running = True
while keep_running:
keep_running=False
for eu, ev in list(g.edges):
if (ev,eu) in list(g.edges):
print(f"Merging {eu} and {ev}")
g = nx.contracted_nodes(g, min(eu,ev), max(eu,ev), self_loops=False)
keep_running = True
break
draw(g)
# -
# To make the hierarchy easier to analyze, we remove the redundant transitive edges using a transitive reduction.
# +
g_ = nx.algorithms.dag.transitive_reduction(g)
# Copy over the style properties (the nx.algorithms.dag.transitive_reduction loses these)
for n in g.nodes:
for k,v in g.nodes[n].items():
g_.nodes[n][k] = v
for e in g_.edges:
g_.edges[e]['dir'] = 'back'
draw(g_)
# -
# The arrows can be iterpreted as ___'isa'___ relations.
# +
# draw(g_, filename='3hop_query_hierarchy.gv', show=False)
# draw(g_, filename='3hop_query_hierarchy.pdf', show=False)
# -
|
Query Analysis -- 3hop-Minimal.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# #### Detecting missing values with np.nan
import pandas as pd
import numpy as np
df=pd.DataFrame({'col':[1,np.nan]})
df
df==np.nan
# This is because comparing missing value to anything results in a False - instead of this you should
# use
df=pd.DataFrame({'col':[1,np.nan]})
df.isnull()
# #### TIP :
#
# ##### Pandas don't support missing in attributes of type integer. For example if you have missings in the grade column:
#
# +
#df= pd.read_csv("data.csv", dtype={'grade': int})
# error: Integer column has NA values
# -
# In this case you just should use float instead of integers or set the object dtype.
# ### Automatic Data Alignment (index-awared behaviour)
#
# #### If you want to append a series of values [1,2] to the column of dataframe df, you will get NaNs:
# +
import pandas as pd
series=pd.Series([1,2])
df=pd.DataFrame(index=[3,4])
df['col']=series
df
# -
# because setting a new column automatically aligns the data by the indexe, and your values 1 and
# 2 would get the indexes 0 and 1, and not 3 and 4 as in your data frame:
df=pd.DataFrame(index=[1,2])
df['col']=series
df
# #### If you want to ignore index, you should set the .values at the end:
df['col']=series.values
df
|
13.NaN values.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# [Return to Home Page](START_HERE.ipynb)
# # References
#
# - https://www.kaggle.com/
# - https://rapids.ai/about.html
# - https://distributed.dask.org/en/latest/worker.html#
# - https://elitedatascience.com/imbalanced-classes
# - https://machinelearningmastery.com/train-test-split-for-evaluating-machine-learning-algorithms/
# - https://github.com/zronaghi/nasa-ml-workshop
# - https://github.com/rapidsai/notebooks
#
# # Other Bootcamps
# The contents of this Bootcamp originates from [OpenACC GPU Bootcamp Github](https://github.com/gpuhackathons-org/gpubootcamp). Here are some additional Bootcamp which might be of interest:
#
# - [Intelligent Video Analytic using DeepStream](https://github.com/gpuhackathons-org/gpubootcamp/tree/master/ai/DeepStream)
#
# [Return to Home Page](START_HERE.ipynb)
|
ai/RAPIDS/English/Python/jupyter_notebook/References.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + deletable=true editable=true
"""Convolutional Neural Network Estimator for MNIST, built with tf.layers."""
# FROM : https://www.tensorflow.org/tutorials/layers#building_the_cnn_mnist_classifier
# CODE : https://www.tensorflow.org/code/tensorflow/examples/tutorials/layers/cnn_mnist.py
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
import pickle
import tensorflow as tf
from tensorflow.contrib import learn
from tensorflow.contrib.learn.python.learn.estimators import model_fn as model_fn_lib
tf.logging.set_verbosity(tf.logging.INFO) # Quite a lot...
#tf.logging.set_verbosity(tf.logging.WARN) # This prevents Logging ...
do_training = False
# -
import sys
print(sys.version)
print('Tensorflow:',tf.__version__)
# Expecting:
# ```
# Tensorflow: 1.0.0
# 3.5.2 (default, Sep 14 2016, 11:28:32)
# [GCC 6.2.1 20160901 (Red Hat 6.2.1-1)]
# ```
# + deletable=true editable=true
def cnn_model_fn(features, integer_labels, mode):
"""Model function for CNN."""
#print("Run cnn_model_fn, mode=%s" % (mode,))
if type(features) is dict:
#print("New-style feature input")
features_images=features['images']
else:
print("OLD-style feature input (DEPRECATED)")
features_images=features
# Input Layer
# Reshape X to 4-D tensor: [batch_size, width, height, channels]
# MNIST images are 28x28 pixels, and have one color channel
input_layer = tf.reshape(features_images, [-1, 28, 28, 1], name='input_layer')
# Convolutional Layer #1
# Computes 32 features using a 5x5 filter with ReLU activation.
# Padding is added to preserve width and height.
# Input Tensor Shape: [batch_size, 28, 28, 1]
# Output Tensor Shape: [batch_size, 28, 28, 32]
conv1 = tf.layers.conv2d(
inputs=input_layer,
filters=32,
kernel_size=[5, 5],
padding="same",
activation=tf.nn.relu)
# Pooling Layer #1
# First max pooling layer with a 2x2 filter and stride of 2
# Input Tensor Shape: [batch_size, 28, 28, 32]
# Output Tensor Shape: [batch_size, 14, 14, 32]
pool1 = tf.layers.max_pooling2d(inputs=conv1, pool_size=[2, 2], strides=2)
# Convolutional Layer #2
# Computes 64 features using a 5x5 filter.
# Padding is added to preserve width and height.
# Input Tensor Shape: [batch_size, 14, 14, 32]
# Output Tensor Shape: [batch_size, 14, 14, 64]
conv2 = tf.layers.conv2d(
inputs=pool1,
filters=64,
kernel_size=[5, 5],
padding="same",
activation=tf.nn.relu)
# Pooling Layer #2
# Second max pooling layer with a 2x2 filter and stride of 2
# Input Tensor Shape: [batch_size, 14, 14, 64]
# Output Tensor Shape: [batch_size, 7, 7, 64]
pool2 = tf.layers.max_pooling2d(inputs=conv2, pool_size=[2, 2], strides=2)
# Flatten tensor into a batch of vectors
# Input Tensor Shape: [batch_size, 7, 7, 64]
# Output Tensor Shape: [batch_size, 7 * 7 * 64]
pool2_flat = tf.reshape(pool2, [-1, 7 * 7 * 64])
# Dense Layer
# Densely connected layer with 1024 neurons
# Input Tensor Shape: [batch_size, 7 * 7 * 64]
# Output Tensor Shape: [batch_size, 1024]
dense = tf.layers.dense(inputs=pool2_flat, units=1024, activation=tf.nn.relu)
# Add dropout operation; 0.6 probability that element will be kept
dropout = tf.layers.dropout(
inputs=dense, rate=0.4, training= (mode == learn.ModeKeys.TRAIN) )
# Logits layer
# Input Tensor Shape: [batch_size, 1024]
# Output Tensor Shape: [batch_size, 10]
logits = tf.layers.dense(inputs=dropout, units=10)
#logits = tf.Print(logits, [input_layer.get_shape(), integer_labels.get_shape()], "Debug size information : ", first_n=1)
#logits = tf.layers.dense(inputs=dense, units=10)
loss = None
train_op = None
# Calculate Loss (for both TRAIN and EVAL modes)
if mode != learn.ModeKeys.INFER:
onehot_labels = tf.one_hot(indices=tf.cast(integer_labels, tf.int32), depth=10)
loss = tf.losses.softmax_cross_entropy(logits=logits, onehot_labels=onehot_labels)
#loss = tf.nn.sparse_softmax_cross_entropy_with_logits(logits=logits, labels=[ cls_targets[0] ])
# Configure the Training Op (for TRAIN mode)
if mode == learn.ModeKeys.TRAIN:
train_op = tf.contrib.layers.optimize_loss(
loss=loss,
global_step=tf.contrib.framework.get_global_step(),
learning_rate=0.001,
#optimizer="SGD")
optimizer="Adam")
# Generate Predictions
predictions = {
"classes": tf.argmax(input=logits, axis=1),
"probabilities": tf.nn.softmax(logits, name="softmax_tensor"),
"logits": logits,
#"before_and_after":( input_layer, logits ),
#"before_and_after":dict(input_layer=input_layer, logits=logits),
}
# For OLD-STYLE inputs (needs wierd 'evaluate' metric)
if mode == model_fn_lib.ModeKeys.EVAL:
predictions['input_grad'] = tf.gradients(loss, [input_layer])[0]
# For NEW-STYLE inputs (can smuggle in extra parameters)
if type(features) is dict and 'fake_targets' in features:
loss_vs_target = tf.nn.sparse_softmax_cross_entropy_with_logits(
logits=logits,
labels=features['fake_targets']
)
predictions['image_gradient_vs_fake_target'] = tf.gradients(loss_vs_target, [input_layer])[0]
# Return a ModelFnOps object
return model_fn_lib.ModelFnOps(
mode=mode, predictions=predictions, loss=loss, train_op=train_op)
# + deletable=true editable=true
# Create the Estimator : https://www.tensorflow.org/extend/estimators
mnist_classifier = learn.Estimator(
model_fn=cnn_model_fn, model_dir="mnist_model/cnn") # This is relative to the ipynb
# Check : the checkpoints file in 'mnist_model/cnn' has filenames that are in same directory
# -
if False:
print( mnist_classifier.get_variable_names() )
#mnist_classifier.get_variable_value('conv2d/bias')
#mnist_classifier.save()
#tf.get_variable('input_layer')
print( tf.global_variables() )
print( tf.get_collection(tf.GraphKeys.GLOBAL_VARIABLES) )
print( [n.name for n in tf.get_default_graph().as_graph_def().node] )
# + deletable=true editable=true
# Load training and eval data
mnist = learn.datasets.load_dataset("mnist")
train_data = mnist.train.images # Returns np.array
train_labels = np.asarray(mnist.train.labels, dtype=np.int32)
eval_data = mnist.test.images # Returns np.array
eval_labels = np.asarray(mnist.test.labels, dtype=np.int32)
#print(eval_labels[7])
print("Data Loaded")
#https://www.tensorflow.org/get_started/input_fn#passing_input_fn_data_to_your_model
def mnist_batch_input_fn(dataset, batch_size=100, seed=None, num_epochs=1):
# If seed is defined, this will shuffle data into batches
if False: # This is the idea (but numpy, rather than Tensors)
feature_dict = dict( images = dataset.images )
labels = np.asarray( dataset.labels, dtype=np.int32)
return feature_dict, labels # but batch_size==EVERYTHING_AT_ONCE, unless we batch it up...
np_labels = np.asarray( dataset.labels, dtype=np.int32)
# Instead, build a Tensor dict
all_images = tf.constant( dataset.images, shape=dataset.images.shape, verify_shape=True )
all_labels = tf.constant( np_labels, shape=np_labels.shape, verify_shape=True )
print("mnist_batch_input_fn sizing : ",
dataset.images.shape,
np.asarray( dataset.labels, dtype=np.int32).shape,
np.asarray( [dataset.labels], dtype=np.int32).T.shape,
)
# And create a 'feeder' to batch up the data appropriately...
image, label = tf.train.slice_input_producer( [all_images, all_labels],
num_epochs=num_epochs,
shuffle=(seed is not None), seed=seed,
)
dataset_dict = dict( images=image, labels=label ) # This becomes pluralized into batches by .batch()
batch_dict = tf.train.batch( dataset_dict, batch_size,
num_threads=1, capacity=batch_size*2,
enqueue_many=False, shapes=None, dynamic_pad=False,
allow_smaller_final_batch=False,
shared_name=None, name=None)
batch_labels = batch_dict.pop('labels')
# Return :
# 1) a mapping of feature columns to Tensors with the corresponding feature data, and
# 2) a Tensor containing labels
return batch_dict, batch_labels
batch_size=100
# + deletable=true editable=true
if do_training:
# Set up logging for predictions
# Log the values in the "Softmax" tensor with label "probabilities"
tensors_to_log = {"probabilities": "softmax_tensor"}
logging_hook = tf.train.LoggingTensorHook( tensors=tensors_to_log, every_n_secs=20 ) #every_n_iter=1000 )
# Train the model
epochs=5
if False:
mnist_classifier.fit(
x=train_data,
y=train_labels,
batch_size=batch_size,
steps=train_labels.shape[0]/batch_size * epochs,
monitors=[logging_hook]
)
mnist_classifier.fit(
input_fn=lambda: mnist_batch_input_fn(mnist.train, batch_size=batch_size, seed=42, num_epochs=epochs),
#steps=train_labels.shape[0] / batch_size * epochs,
#monitors=[logging_hook],
)
# + deletable=true editable=true
if False: # This should log 'hi[1]' to the console (not to the Jupyter window...)
# http://stackoverflow.com/questions/37898478
# /is-there-a-way-to-get-tensorflow-tf-print-output-to-appear-in-jupyter-notebook-o
a = tf.constant(1.0)
a = tf.Print(a, [a], 'hi')
sess = tf.Session()
a.eval(session=sess)
# + deletable=true editable=true
# Configure the accuracy metric for evaluation
cnn_metrics = {
"accuracy":
learn.MetricSpec(
metric_fn=tf.metrics.accuracy, prediction_key="classes"),
}
# Evaluate the model and print results
#cnn_eval_results = mnist_classifier.evaluate( x=eval_data, y=eval_labels, metrics=cnn_metrics)
cnn_eval_results = mnist_classifier.evaluate(
input_fn=lambda: mnist_batch_input_fn(mnist.test, batch_size=batch_size),
metrics=cnn_metrics,
#steps=eval_labels.shape[0]/batch_size,
)
print(cnn_eval_results)
# -
# Ok, so the built Estimator gets ~99% accuracy on the test set in <20 secs on CPU.
# ### Adversarial Images
#
# Let's create some adversarial digits for MNIST that fool the original Estimator
# +
train_offset = 17
image_orig = train_data[train_offset] # This is a flat numpy array with an image in it
label_orig = train_labels[train_offset] # This the digit label for that image
#label_target = (label_orig+1) % 10
label_target = 3
label_orig, label_target
# +
if False: # Works, but 'old-style'
#class_predictions = mnist_classifier.predict( x=np.array([image_orig]), batch_size=1, as_iterable=False)
class_predictions = mnist_classifier.predict( x=image_orig, as_iterable=False)
class_predictions['probabilities'][0]
#class_predictions = mnist_classifier.predict( x=image_orig, outputs=['probabilities'], as_iterable=False)
#class_predictions
def mnist_direct_data_input_fn(features_np_dict, targets_np):
features_dict = { k:tf.constant(v) for k,v in features_np_dict.items()}
targets = None if targets_np is None else tf.constant(targets_np)
return features_dict, targets
class_predictions_generator = mnist_classifier.predict(
input_fn=lambda: mnist_direct_data_input_fn(dict(images=np.array([image_orig])), None),
outputs=['probabilities'],
)
for class_predictions in class_predictions_generator:
break # Get the first one...
class_predictions['probabilities']
# -
# ### Intuition behind 'gradient' for explicit inception version ...
# +
## Set the graph for the Inception model as the default graph,
## so that all changes inside this with-block are done to that graph.
#with model.graph.as_default():
# # Add a placeholder variable for the target class-number.
# # This will be set to e.g. 300 for the 'bookcase' class.
# pl_cls_target = tf.placeholder(dtype=tf.int32)
#
# # Add a new loss-function. This is the cross-entropy.
# # See Tutorial #01 for an explanation of cross-entropy.
# loss = tf.nn.sparse_softmax_cross_entropy_with_logits(logits=y_logits, labels=[pl_cls_target])
#
# # Get the gradient for the loss-function with regard to
# # the resized input image.
# gradient = tf.gradients(loss, resized_image)
# -
# This is the way to do it 'OLD style', where we smuggle out the information during an EVALUATE() call
if False:
# FIGURING-IT-OUT STEP : WORKS
def metric_accuracy(cls_targets, predictions):
return tf.metrics.accuracy(cls_targets, predictions)
# FIGURING-IT-OUT STEP : WORKS
def metric_accuracy_here(labels, predictions, weights=None, metrics_collections=None, updates_collections=None, name=None):
if labels.dtype != predictions.dtype:
predictions = tf.cast(predictions, labels.dtype)
is_correct = tf.to_float(tf.equal(predictions, labels))
return tf.metrics.mean(is_correct, weights, metrics_collections, updates_collections, name or 'accuracy')
# FIGURING-IT-OUT STEP : WORKS
def metric_mean_here(labels, predictions, weights=None, metrics_collections=None, updates_collections=None, name=None):
return tf.metrics.mean(labels, weights, metrics_collections, updates_collections, name or 'gradient_mean')
# FINALLY! :: WORKS
def metric_concat_here(labels, predictions, weights=None, metrics_collections=None, updates_collections=None, name=None):
return tf.contrib.metrics.streaming_concat(labels, axis=0, max_size=None,
metrics_collections=metrics_collections,
updates_collections=updates_collections,
name = name or 'gradient_concat')
model_gradient = {
# "accuracy": learn.MetricSpec(metric_fn=tf.metrics.accuracy, prediction_key="classes"), # WORKS
# "accuracy": learn.MetricSpec(metric_fn=metric_accuracy, prediction_key="classes"), # WORKS
# "accuracy": learn.MetricSpec(metric_fn=metric_accuracy_here, prediction_key="classes"), # WORKS
# "accuracy": learn.MetricSpec(metric_fn=metric_mean_here, prediction_key="classes"), # WORKS
"gradient": learn.MetricSpec(metric_fn=metric_concat_here, prediction_key="input_grad"), # WORKS!
}
# Evaluate the model and print results OLD-STYLE
cnn_gradient = mnist_classifier.evaluate(
x=np.array([ image_orig ], dtype='float32'), y=np.array([ label_target ], dtype='int32'),
batch_size=1,
#input_fn = (lambda: (np.array([ image_orig ], dtype='float32'), np.array([7], dtype='int32'))),
metrics=model_gradient)
#cnn_gradient = mnist_classifier.evaluate( x=image_orig, y=np.int32(7), metrics=model_gradient)
cnn_gradient['gradient'].shape
# +
# NEW-STYLE : We can get the data from a .PREDICT() directly (outputs=[xyz] is passed through)
def mnist_direct_data_input_fn(features_np_dict, targets_np):
features_dict = { k:tf.constant(v) for k,v in features_np_dict.items()}
targets = None if targets_np is None else tf.constant(targets_np)
return features_dict, targets
tensor_prediction_generator = mnist_classifier.predict(
input_fn=lambda: mnist_direct_data_input_fn(
dict(
images=np.array([ image_orig ]),
fake_targets=np.array([ label_target ], dtype=np.int),
), None),
outputs=['image_gradient_vs_fake_target'],
)
for tensor_predictions in tensor_prediction_generator:
break # Get the first one...
grads = tensor_predictions['image_gradient_vs_fake_target']
grads.shape,grads.min(),grads.max()
# -
# Plot the gradients
plt.figure(figsize=(12,3))
for i in range(1):
plt.subplot(1, 10, i+1)
plt.imshow(((grads+8.)/11.).reshape((28, 28)), cmap='gray', interpolation='nearest')
plt.axis('off')
def find_adversarial_noise(image_np, cls_target, model,
pixel_max=255, noise_limit=None,
required_score=0.99, max_iterations=50):
"""
Find the noise that must be added to the given image so
that it is classified as the target-class by the given model.
image_np: numpy image in correct 'picture-like' format
cls_target: Target class-number (integer between 0-n_classes).
noise_limit: Limit for pixel-values in the noise (scaled for 0...255 image)
required_score: Stop when target-class 'probabilty' reaches this.
max_iterations: Max number of optimization iterations to perform.
"""
# Initialize the noise to zero.
noise = np.zeros_like( image_np )
# Perform a number of optimization iterations to find
# the noise that causes mis-classification of the input image.
for i in range(max_iterations):
print("Iteration:", i)
# The noisy image is just the sum of the input image and noise.
noisy_image = image_np + noise
# Ensure the pixel-values of the noisy image are between
# 0 and pixel_max like a real image. If we allowed pixel-values
# outside this range then maybe the mis-classification would
# be due to this 'illegal' input breaking the Inception model.
noisy_image = np.clip(a=noisy_image, a_min=0.0, a_max=float(pixel_max))
# Calculate the predicted class-scores as well as the gradient.
#pred, grad = session.run([y_pred, gradient], feed_dict=feed_dict)
tensor_prediction_generator = model.predict(
input_fn=lambda: mnist_direct_data_input_fn(
dict(
images=np.array([ noisy_image ]),
fake_targets=np.array([ cls_target ], dtype=np.int),
), None),
outputs=['probabilities','logits','image_gradient_vs_fake_target'],
)
for tensor_predictions in tensor_prediction_generator:
break # Get the first one...
#tensor_predictions['image_gradient_vs_fake_target'].shape
pred = tensor_predictions['probabilities']
logits = tensor_predictions['logits']
grad = tensor_predictions['image_gradient_vs_fake_target']
print( ','.join([ ("%.4f" % p) for p in pred ]))
#print(pred.shape, grad.shape)
# The scores (probabilities) for the source and target classes.
# score_source = pred[cls_source]
score_target = pred[cls_target]
# The gradient now tells us how much we need to change the
# noisy input image in order to move the predicted class
# closer to the desired target-class.
# Calculate the max of the absolute gradient values.
# This is used to calculate the step-size.
grad_absmax = np.abs(grad).max()
# If the gradient is very small then use a lower limit,
# because we will use it as a divisor.
if grad_absmax < 1e-10:
grad_absmax = 1e-10
# Calculate the step-size for updating the image-noise.
# This ensures that at least one pixel colour is changed by 7 out of 255
# Recall that pixel colours can have 255 different values.
# This step-size was found to give fast convergence.
step_size = 7/255.0*pixel_max / grad_absmax
# Print the score etc. for the source-class.
#msg = "Source score: {0:>7.2%}, class-number: {1:>4}, class-name: {2}"
#print(msg.format(score_source, cls_source, name_source))
# Print the score etc. for the target-class.
print("Target class (%d) score: %7.4f" % (cls_target, score_target, ))
# Print statistics for the gradient.
msg = "Gradient min: {0:>9.6f}, max: {1:>9.6f}, stepsize: {2:>9.6f}"
print(msg.format(grad.min(), grad.max(), step_size))
# Newline.
print()
# If the score for the target-class is not high enough.
if score_target < required_score:
# Update the image-noise by subtracting the gradient
# scaled by the step-size.
noise -= step_size * grad
# Ensure the noise is within the desired range.
# This avoids distorting the image too much.
if noise_limit is not None:
noise = np.clip(a = noise,
a_min = -noise_limit/255.0*pixel_max,
a_max = noise_limit/255.0*pixel_max)
else:
# Abort the optimization because the score is high enough.
break
return (
noisy_image, noise, score_target, logits
#name_source, name_target, \
#score_source, score_source_org, score_target
)
np.min(image_orig), np.max(image_orig)
# +
print(label_orig, label_target)
image_orig_sq = np.reshape(image_orig, (28,28,1))
res = find_adversarial_noise(image_orig_sq, label_target, mnist_classifier,
pixel_max=1.0, # for 0.0 ... 1.0 images (MNIST)
#pixel_max=255.0, # for 0..255 images (ImageNet)
#noise_limit=7.0,
required_score=0.99, max_iterations=50)
adversarial_image, adversarial_noise, adversarial_score, adversarial_logits = res
# Plot the image, alterted image and noise
plt.figure(figsize=(12,3))
for i,im in enumerate( [image_orig, adversarial_image, adversarial_noise] ):
plt.subplot(1, 10, 1+i)
plt.imshow(im.reshape((28, 28)), cmap='gray', interpolation='nearest')
plt.axis('off')
# +
# tf.getDefaultGraph().finalize()
# + [markdown] deletable=true editable=true
# ### Next Steps
#
# Let's :
#
# * go through the training set and store the logits for [the valid?] training examples;
#
# * build an AutoEncoder on the logits, which minimises reconstruction error;
#
# * histogram the reconstruction error to find a bound above which we can reject an input image;
#
# * attempt to create adversarial examples on an updated network that includes the autoencoder bound as a gating function on the rest of the outputs;
#
# * create an infoGAN network for MNIST that allows us to create digits that are 'between' two classes;
#
# * score the reconstruction error of the between images to look at the rejection regions (which hopefully isolate the islands of acceptance from one another)
# -
# #### Get logit representation for all training examples
# +
# Evaluate the model and gather the results. NB: no seed, since we want to preserve the ordering
# Predictions take ~ 60secs
predictions = mnist_classifier.predict(
input_fn=lambda: mnist_batch_input_fn(mnist.train, batch_size=batch_size),
outputs=['logits'],
as_iterable=True)
train_data_logits = np.array([ p['logits'] for p in predictions ])
predictions = mnist_classifier.predict(
input_fn=lambda: mnist_batch_input_fn(mnist.test, batch_size=batch_size),
outputs=['logits'],
as_iterable=True)
eval_data_logits = np.array([ p['logits'] for p in predictions ])
train_data_logits.shape, eval_data_logits.shape
# +
# Optionally save the logits for quicker iteration...
logits_filename = './mnist_model/logits.pkl'
if not tf.gfile.Exists(logits_filename):
logits_saver = ( train_data_logits, train_labels, eval_data_logits, eval_labels )
pickle.dump(logits_saver, open(logits_filename,'wb'), protocol=pickle.HIGHEST_PROTOCOL)
# -
# #### Explore the logit representations
# Load the logits
if True:
res = pickle.load( open(logits_filename, 'rb'), encoding='iso-8859-1')
train_data_logits, train_labels, eval_data_logits, eval_labels = res
# Show an example #s, target_classes, and logits
print(" %s" % ( ', '.join(["%7s" % l for l in range(10)]),) )
for train_data_example in [99, 98, 84]: # all have a true label of '6'
print("#%4d : '%d' [ %s ]" % (
train_data_example,
train_labels[train_data_example],
', '.join(["%+7.3f" % l for l in train_data_logits[train_data_example,:]]),
))
# +
# Ok, so how about the reconstruction error for the training logits that it gets wrong?
# Create an indicator function that is 1 iff the label doesn't match the best logit answer
train_labels_predicted = np.argmax( train_data_logits, axis=1 )
print("train_labels_predicted.shape :", train_labels_predicted.shape)
print( 'predicted : ',train_labels_predicted[80:100], '\nactual : ', train_labels[80:100] )
#train_error_indices = np.where( train_labels_predicted == train_labels, 0, 1)
train_error_indices = train_labels_predicted != train_labels
print( "Total # of bad training examples : ", np.sum( train_error_indices ) ) # [80:90]
# Gather the 'badly trained logits'
train_error_logits = train_data_logits[train_error_indices]
print("train_error_logits.shape :", train_error_logits.shape)
train_valid_indices = train_labels_predicted == train_labels
train_valid_logits = train_data_logits[train_valid_indices]
# +
# Histogram various pre-processings of the input logits
#def n(x): return x
#def n(x): return ( (x - x.mean(axis=1, keepdims=True))/x.std(axis=1, keepdims=True) )
#def n(x): return ((x - x.min(axis=1, keepdims=True))/(x.max(axis=1, keepdims=True) - x.min(axis=1, keepdims=True) + 0.0001))
#def n(x): return np.fabs(x)
def n(x):
len_em = len_except_max = (x.shape[1]-1)
x_max = x.max(axis=1, keepdims=True)
x_argmax = x.argmax(axis=1)
mean_em = (x.sum(axis=1, keepdims=True) - x_max) / len_em
sumsq_em = np.sum(np.square(x - mean_em), axis=1, keepdims=True) - np.square(x_max - mean_em)
std_em = np.sqrt( sumsq_em / len_em )
y = (x - mean_em) / std_em
y = np.clip(y, -4.0, +4.0)
y[np.arange(x.shape[0]), x_argmax]=5.0
return y
count, bins, patches = plt.hist(n(train_valid_logits).flatten(), 50, normed=1, facecolor='green', alpha=1.0)
count, bins, patches = plt.hist(n(train_error_logits).flatten(), 50, normed=1, facecolor='blue', alpha=0.5)
plt.xlabel('logit')
plt.ylabel('density')
#plt.title(r'$\mathrm{Histogram\ of\ IQ:}\ \mu=100,\ \sigma=15$')
#plt.axis([-4, 6, 0, 0.8])
plt.grid(True)
plt.show()
# -
# ### Build an autoencoder for the preprocessed logits
# Let's build an autoencoder 'regression' model with a hidden layer 'fewer' units
def autoencoder_model_fn(features, unused_labels, mode):
logits_dim = 10
#hidden_dim = logits_dim
hidden_dim = int(logits_dim*.75)
input_layer = features['logits']
# One-hot on the input logit that's > 4.5
one_hot = tf.div( tf.add( tf.sign( tf.subtract(input_layer, 4.5) ), 1.0), 2.0)
one_hot = tf.Print(one_hot, [one_hot], message="one_hot: ", first_n=1, summarize=30 )
# This summary is the inputs with the 'top-1' set to zero
input_remainder = tf.subtract( input_layer, tf.multiply(one_hot, 5.0) )
input_summary = tf.layers.dense(inputs=input_layer, units=int(logits_dim*.5), activation=tf.nn.relu)
combined = tf.concat( [input_summary, one_hot], 1)
# Encoder Dense Layer
#dense1 = tf.layers.dense(inputs=input_layer, units=hidden_dim, activation=tf.nn.relu)
#dense1 = tf.layers.dense(inputs=input_layer, units=logits_dim, activation=tf.nn.relu)
#dense = tf.layers.dense(inputs=input_layer, units=hidden_dim, activation=tf.nn.elu) # ELU!
#dense1 = tf.layers.dense(inputs=input_layer, units=hidden_dim, activation=tf.nn.tanh)
#dense1 = tf.layers.dense(inputs=input_layer, units=logits_dim, activation=tf.nn.tanh)
#dense1 = tf.layers.dense(inputs=combined, units=logits_dim, activation=tf.nn.tanh)
#dense2 = tf.layers.dense(inputs=dense1, units=hidden_dim, activation=tf.nn.tanh)
#dense2 = tf.layers.dense(inputs=dense1, units=logits_dim*2, activation=tf.nn.tanh)
#dense2 = tf.layers.dense(inputs=dense1, units=logits_dim, activation=tf.nn.tanh)
#dense2 = dense1
dense2 = combined
# Add dropout operation; 0.6 probability that element will be kept
#dropout = tf.layers.dropout(
# inputs=dense2, rate=0.9, training=mode == learn.ModeKeys.TRAIN)
# Decoder Dense Layer
#output_layer = tf.layers.dense(inputs=dropout, units=logits_dim)
output_layer = tf.layers.dense(inputs=dense2, units=logits_dim) # Linear activation
loss = None
train_op = None
## Calculate Loss (for both TRAIN and EVAL modes)
#if mode != learn.ModeKeys.INFER:
# loss = tf.losses.mean_squared_error( input_layer, output_layer )
if False:
loss = tf.losses.mean_squared_error( input_layer, output_layer )
if True:
weighted_diff = tf.multiply( tf.subtract(1.0, one_hot), tf.subtract(input_layer, output_layer) )
#weighted_diff = tf.multiply( 1.0, tf.subtract(input_layer, output_layer) )
loss = tf.reduce_mean( tf.multiply (weighted_diff, weighted_diff) )
# Configure the Training Op (for TRAIN mode)
if mode == learn.ModeKeys.TRAIN:
train_op = tf.contrib.layers.optimize_loss(
loss=loss,
global_step=tf.contrib.framework.get_global_step(),
learning_rate=0.001,
optimizer="Adam")
# Generate Predictions
predictions = {
"mse": loss,
"regenerated":output_layer,
"gradient": tf.gradients(loss, input_layer),
}
# Return a ModelFnOps object
return model_fn_lib.ModelFnOps(
mode=mode, predictions=predictions, loss=loss, train_op=train_op)
mnist_autoencoder = learn.Estimator(
model_fn=autoencoder_model_fn, model_dir="mnist_model/autoencoder")
# +
def mnist_logit_batch_input_fn(logits, batch_size=100, seed=None, num_epochs=1):
# If seed is defined, this will shuffle data into batches
all_logits = tf.constant( logits, shape=logits.shape, verify_shape=True )
fake_labels = tf.constant( np.zeros((logits.shape[0],)) )
print("mnist_logit_batch_input_fn sizing : ", all_logits.shape, )
# And create a 'feeder' to batch up the data appropriately...
logit, label = tf.train.slice_input_producer( [ all_logits, fake_labels ],
num_epochs=num_epochs,
shuffle=(seed is not None), seed=seed,
)
dataset_dict = dict( logits=logit, labels=label ) # This becomes pluralized into batches by .batch()
batch_dict = tf.train.batch( dataset_dict, batch_size,
num_threads=1, capacity=batch_size*2,
enqueue_many=False, shapes=None, dynamic_pad=False,
allow_smaller_final_batch=False,
shared_name=None, name=None)
batch_labels = batch_dict.pop('labels')
#batch_labels = batch_dict.pop('logits')
# Return :
# 1) a mapping of feature columns to Tensors with the corresponding feature data, and
# 2) fake_labels (all 0)
return batch_dict, batch_labels
autoenc_batch_size, autoenc_epochs = 100, 20
# +
# Fit the autoencoder to the logits
mnist_autoencoder.fit(
input_fn=lambda: mnist_logit_batch_input_fn( n(train_valid_logits), #train_data_logits,
batch_size=autoenc_batch_size,
seed=42,
num_epochs=autoenc_epochs),
)
# -
# * n/2 hidden INFO:tensorflow:Saving checkpoints for 25000 into mnist_model/autoencoder/model.ckpt.
# * n/2 hidden INFO:tensorflow:Loss for final step: 1.2686.
#
# * 2xReLU INFO:tensorflow:Saving checkpoints for 25000 into mnist_model/autoencoder/model.ckpt.
# * 2xReLU INFO:tensorflow:Loss for final step: 1.47784e-05.
#
# * ELU+ReLU INFO:tensorflow:Saving checkpoints for 5000 into mnist_model/autoencoder/model.ckpt.
# * ELU+ReLU INFO:tensorflow:Loss for final step: 0.00331942.
# +
# Configure the accuracy metric for evaluation
def metric_mean_here(labels, predictions, weights=None, metrics_collections=None, updates_collections=None, name=None):
return tf.metrics.mean(labels, weights, metrics_collections, updates_collections, name or 'gradient_mean')
autoenc_metrics = {
"loss":learn.MetricSpec(metric_fn=metric_mean_here, prediction_key="mse"),
}
# Evaluate the model and print results
#autoencoder_eval_results = mnist_autoencoder.evaluate( x=eval_data_logits, y=eval_data_logits, metrics=auto_metrics)
autoencoder_train_results = mnist_autoencoder.evaluate(
input_fn=lambda: mnist_logit_batch_input_fn(n(train_valid_logits), # train_data_logits,
batch_size=train_valid_logits.shape[0],
),
metrics=autoenc_metrics)
print(autoencoder_train_results)
autoencoder_eval_results = mnist_autoencoder.evaluate(
input_fn=lambda: mnist_logit_batch_input_fn(n(eval_data_logits),
batch_size=eval_data_logits.shape[0],
),
metrics=autoenc_metrics)
print(autoencoder_eval_results)
# -
# * {'loss': 1.1115935e-06, 'global_step': 18250} => This autoencoder thing works
if False: # Double up train_error_logits to check whether mean() is working
train_error_logits = np.vstack( [train_error_logits,train_error_logits] )
train_error_logits.shape
# +
# What is the mean reconstruction error for the incorrectly trained digits?
autoencoder_error_results = mnist_autoencoder.evaluate(
input_fn=lambda: mnist_logit_batch_input_fn(n(train_error_logits),
batch_size=train_error_logits.shape[0],
),
metrics=autoenc_metrics)
print(autoencoder_error_results)
# -
adversarial_logits
# +
autoencoder_adversarial_results = mnist_autoencoder.evaluate(
input_fn=lambda: mnist_logit_batch_input_fn(n(np.array([
#train_data_logits[84],
adversarial_logits,
])),
batch_size=1,
),
metrics=autoenc_metrics)
print(autoencoder_adversarial_results)
# -
# ?tf.reduce_sum()
|
notebooks/work-in-progress/adversarial/MNIST-CNN_learn-layers.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import geopandas
import matplotlib.pyplot as plt
import cartopy.crs as ccrs
from requests.utils import requote_uri
import urllib.parse
covid_sc_url = 'https://scdhec.gov/sites/default/files/afs-covid/county-dashboard/Suppressed County Data by Week.csv'
covid_sc_url = requote_uri(covid_sc_url)
covid_sc = pd.read_csv(covid_sc_url)
covid_sc
covid_sc['Day Week Ends On'] = pd.to_datetime(covid_sc['Day Week Ends On'])
most_recent = covid_sc['Day Week Ends On'].max()
most_recent_data = covid_sc.loc[covid_sc['Day Week Ends On'] == most_recent]
most_recent_data
usa_counties = geopandas.read_file('cb_2017_us_county_20m/cb_2017_us_county_20m.shp')
sc_counties = usa_counties.loc[usa_counties['STATEFP'] == '45']
sc_counties.head()
merge = pd.merge(left=sc_counties[['geometry', 'NAME']],
right=most_recent_data,
left_on='NAME',
right_on='County',).drop(columns=['NAME'])
merge.head()
# Convert confirmed case columns integers
merge['Count Confirmed Cases Suppressed if 1-4'] = merge['Count Confirmed Cases Suppressed if 1-4'].astype(int)
# +
# Make map
fig = plt.figure(figsize=(1280/72, 720/72))
ax = fig.add_axes([0, 0, 1, 1], projection=ccrs.PlateCarree(),
frameon=False)
ax.patch.set_visible(False)
ax.set_title('South Carolina COVID Cases', fontweight='bold', fontsize='36')
merge.plot(column='Count Confirmed Cases Suppressed if 1-4',
cmap='OrRd', edgecolor='black',
scheme='quantiles', legend=True,
figsize=(1280/72, 720/72),
ax=ax)
fig.savefig('sc_covid.png', bbox_inches='tight')
# -
|
python/covid.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import os
import torch
import torch.nn as nn
import torch.nn.functional as F
from torch import optim
import time
from torch.utils.data import Dataset, DataLoader
import torchaudio
import torchvision.transforms as transforms
import matplotlib.pyplot as plt
from sklearn.preprocessing import label_binarize
from sklearn.utils.multiclass import unique_labels
from sklearn.metrics import confusion_matrix
from sklearn.metrics import roc_curve, auc
from itertools import cycle
# -
device = torch.device('cuda')
class CNN_1D(nn.Module):
def __init__(self):
super(CNN_1D, self).__init__()
self.conv1 = nn.Conv1d(1, 64, 50, stride=5, padding=25)
self.bn1 = nn.BatchNorm1d(64)
self.activation1 = nn.ReLU()
self.maxpool1 = nn.MaxPool1d(kernel_size=10, stride=5)
##########################################################################
self.conv2a_1 = nn.Conv1d(64, 256, 1, stride=1, padding=0)
self.bn2a_1 = nn.BatchNorm1d(256)
self.conv2a_2a = nn.Conv1d(64, 64, 1, stride=1, padding=0)
self.bn2a_2a = nn.BatchNorm1d(64)
self.activation2a_2a = nn.ReLU()
self.conv2a_2b = nn.Conv1d(64, 64, 10, stride=1, padding=5)
self.bn2a_2b = nn.BatchNorm1d(64)
self.activation2a_2b = nn.ReLU()
self.conv2a_2c = nn.Conv1d(64, 256, 1, stride=1, padding=0)
self.bn2a_2c = nn.BatchNorm1d(256)
self.activation2a = nn.ReLU()
##########################################################################
self.conv2b_2a = nn.Conv1d(256, 64, 1, stride=1, padding=0)
self.bn2b_2a = nn.BatchNorm1d(64)
self.activation2b_2a = nn.ReLU()
self.conv2b_2b = nn.Conv1d(64, 64, 10, stride=1, padding=5)
self.bn2b_2b = nn.BatchNorm1d(64)
self.activation2b_2b = nn.ReLU()
self.conv2b_2c = nn.Conv1d(64, 256, 1, stride=1, padding=0)
self.bn2b_2c = nn.BatchNorm1d(256)
self.activation2b = nn.ReLU()
##########################################################################
self.conv2c_2a = nn.Conv1d(256, 64, 1, stride=1, padding=0)
self.bn2c_2a = nn.BatchNorm1d(64)
self.activation2c_2a = nn.ReLU()
self.conv2c_2b = nn.Conv1d(64, 64, 10, stride=1, padding=5)
self.bn2c_2b = nn.BatchNorm1d(64)
self.activation2c_2b = nn.ReLU()
self.conv2c_2c = nn.Conv1d(64, 256, 1, stride=1, padding=0)
self.bn2c_2c = nn.BatchNorm1d(256)
self.activation2c = nn.ReLU()
##########################################################################
self.conv3a_1 = nn.Conv1d(256, 512, 1, stride=5, padding=0)
self.bn3a_1 = nn.BatchNorm1d(512)
self.conv3a_2a = nn.Conv1d(256, 128, 1, stride=5, padding=0)
self.bn3a_2a = nn.BatchNorm1d(128)
self.activation3a_2a = nn.ReLU()
self.conv3a_2b = nn.Conv1d(128, 128, 10, stride=1, padding=5)
self.bn3a_2b = nn.BatchNorm1d(128)
self.activation3a_2b = nn.ReLU()
self.conv3a_2c = nn.Conv1d(128, 512, 1, stride=1, padding=0)
self.bn3a_2c = nn.BatchNorm1d(512)
self.activation3a = nn.ReLU()
##########################################################################
self.conv3b_2a = nn.Conv1d(512, 128, 1, stride=1, padding=0)
self.bn3b_2a = nn.BatchNorm1d(128)
self.activation3b_2a = nn.ReLU()
self.conv3b_2b = nn.Conv1d(128, 128, 10, stride=1, padding=5)
self.bn3b_2b = nn.BatchNorm1d(128)
self.activation3b_2b = nn.ReLU()
self.conv3b_2c = nn.Conv1d(128, 512, 1, stride=1, padding=0)
self.bn3b_2c = nn.BatchNorm1d(512)
self.activation3b = nn.ReLU()
##########################################################################
self.conv3c_2a = nn.Conv1d(512, 128, 1, stride=1, padding=0)
self.bn3c_2a = nn.BatchNorm1d(128)
self.activation3c_2a = nn.ReLU()
self.conv3c_2b = nn.Conv1d(128, 128, 10, stride=1, padding=5)
self.bn3c_2b = nn.BatchNorm1d(128)
self.activation3c_2b = nn.ReLU()
self.conv3c_2c = nn.Conv1d(128, 512, 1, stride=1, padding=0)
self.bn3c_2c = nn.BatchNorm1d(512)
self.activation3c = nn.ReLU()
##########################################################################
self.conv3d_2a = nn.Conv1d(512, 128, 1, stride=1, padding=0)
self.bn3d_2a = nn.BatchNorm1d(128)
self.activation3d_2a = nn.ReLU()
self.conv3d_2b = nn.Conv1d(128, 128, 10, stride=1, padding=5)
self.bn3d_2b = nn.BatchNorm1d(128)
self.activation3d_2b = nn.ReLU()
self.conv3d_2c = nn.Conv1d(128, 512, 1, stride=1, padding=0)
self.bn3d_2c = nn.BatchNorm1d(512)
self.activation3d = nn.ReLU()
##########################################################################
self.conv4a_1 = nn.Conv1d(512, 1024, 1, stride=5, padding=0)
self.bn4a_1 = nn.BatchNorm1d(1024)
self.conv4a_2a = nn.Conv1d(512, 256, 1, stride=5, padding=0)
self.bn4a_2a = nn.BatchNorm1d(256)
self.activation4a_2a = nn.ReLU()
self.conv4a_2b = nn.Conv1d(256, 256, 10, stride=1, padding=5)
self.bn4a_2b = nn.BatchNorm1d(256)
self.activation4a_2b = nn.ReLU()
self.conv4a_2c = nn.Conv1d(256, 1024, 1, stride=1, padding=0)
self.bn4a_2c = nn.BatchNorm1d(1024)
self.activation4a = nn.ReLU()
##########################################################################
self.conv4b_2a = nn.Conv1d(1024, 256, 1, stride=1, padding=0)
self.bn4b_2a = nn.BatchNorm1d(256)
self.activation4b_2a = nn.ReLU()
self.conv4b_2b = nn.Conv1d(256, 256, 10, stride=1, padding=5)
self.bn4b_2b = nn.BatchNorm1d(256)
self.activation4b_2b = nn.ReLU()
self.conv4b_2c = nn.Conv1d(256, 1024, 1, stride=1, padding=0)
self.bn4b_2c = nn.BatchNorm1d(1024)
self.activation4b = nn.ReLU()
##########################################################################
self.conv4c_2a = nn.Conv1d(1024, 256, 1, stride=1, padding=0)
self.bn4c_2a = nn.BatchNorm1d(256)
self.activation4c_2a = nn.ReLU()
self.conv4c_2b = nn.Conv1d(256, 256, 10, stride=1, padding=5)
self.bn4c_2b = nn.BatchNorm1d(256)
self.activation4c_2b = nn.ReLU()
self.conv4c_2c = nn.Conv1d(256, 1024, 1, stride=1, padding=0)
self.bn4c_2c = nn.BatchNorm1d(1024)
self.activation4c = nn.ReLU()
##########################################################################
self.conv4d_2a = nn.Conv1d(1024, 256, 1, stride=1, padding=0)
self.bn4d_2a = nn.BatchNorm1d(256)
self.activation4d_2a = nn.ReLU()
self.conv4d_2b = nn.Conv1d(256, 256, 10, stride=1, padding=5)
self.bn4d_2b = nn.BatchNorm1d(256)
self.activation4d_2b = nn.ReLU()
self.conv4d_2c = nn.Conv1d(256, 1024, 1, stride=1, padding=0)
self.bn4d_2c = nn.BatchNorm1d(1024)
self.activation4d = nn.ReLU()
##########################################################################
self.conv4e_2a = nn.Conv1d(1024, 256, 1, stride=1, padding=0)
self.bn4e_2a = nn.BatchNorm1d(256)
self.activation4e_2a = nn.ReLU()
self.conv4e_2b = nn.Conv1d(256, 256, 10, stride=1, padding=5)
self.bn4e_2b = nn.BatchNorm1d(256)
self.activation4e_2b = nn.ReLU()
self.conv4e_2c = nn.Conv1d(256, 1024, 1, stride=1, padding=0)
self.bn4e_2c = nn.BatchNorm1d(1024)
self.activation4e = nn.ReLU()
##########################################################################
self.conv4f_2a = nn.Conv1d(1024, 256, 1, stride=1, padding=0)
self.bn4f_2a = nn.BatchNorm1d(256)
self.activation4f_2a = nn.ReLU()
self.conv4f_2b = nn.Conv1d(256, 256, 10, stride=1, padding=5)
self.bn4f_2b = nn.BatchNorm1d(256)
self.activation4f_2b = nn.ReLU()
self.conv4f_2c = nn.Conv1d(256, 1024, 1, stride=1, padding=0)
self.bn4f_2c = nn.BatchNorm1d(1024)
self.activation4f = nn.ReLU()
##########################################################################
self.conv5a_1 = nn.Conv1d(1024, 2048, 1, stride=5, padding=0)
self.bn5a_1 = nn.BatchNorm1d(2048)
self.conv5a_2a = nn.Conv1d(1024, 512, 1, stride=5, padding=0)
self.bn5a_2a = nn.BatchNorm1d(512)
self.activation5a_2a = nn.ReLU()
self.conv5a_2b = nn.Conv1d(512, 512, 10, stride=1, padding=5)
self.bn5a_2b = nn.BatchNorm1d(512)
self.activation5a_2b = nn.ReLU()
self.conv5a_2c = nn.Conv1d(512, 2048, 1, stride=1, padding=0)
self.bn5a_2c = nn.BatchNorm1d(2048)
self.activation5a = nn.ReLU()
##########################################################################
self.conv5b_2a = nn.Conv1d(2048, 512, 1, stride=1, padding=0)
self.bn5b_2a = nn.BatchNorm1d(512)
self.activation5b_2a = nn.ReLU()
self.conv5b_2b = nn.Conv1d(512, 512, 10, stride=1, padding=5)
self.bn5b_2b = nn.BatchNorm1d(512)
self.activation5b_2b = nn.ReLU()
self.conv5b_2c = nn.Conv1d(512, 2048, 1, stride=1, padding=0)
self.bn5b_2c = nn.BatchNorm1d(2048)
self.activation5b = nn.ReLU()
##########################################################################
self.conv5c_2a = nn.Conv1d(2048, 512, 1, stride=1, padding=0)
self.bn5c_2a = nn.BatchNorm1d(512)
self.activation5c_2a = nn.ReLU()
self.conv5c_2b = nn.Conv1d(512, 512, 10, stride=1, padding=5)
self.bn5c_2b = nn.BatchNorm1d(512)
self.activation5c_2b = nn.ReLU()
self.conv5c_2c = nn.Conv1d(512, 2048, 1, stride=1, padding=0)
self.bn5c_2c = nn.BatchNorm1d(2048)
self.activation5c = nn.ReLU()
##########################################################################
self.AvgPool = nn.AvgPool1d(21)
def forward(self, x):
x = self.conv1(x)
x = self.bn1(x)
x = self.activation1(x)
x = self.maxpool1(x)
res = x
res = self.conv2a_1(res)
res = self.bn2a_1(res)
x = self.conv2a_2a(x)
x = self.bn2a_2a(x)
x = self.activation2a_2a(x)
x = self.conv2a_2b(x)
x = self.bn2a_2b(x)
x = self.activation2a_2b(x)
x = self.conv2a_2c(x)
x = self.bn2a_2c(x)
x = x[:,:,:-1] + res
x = self.activation2a(x)
res = x
x = self.conv2b_2a(x)
x = self.bn2b_2a(x)
x = self.activation2b_2a(x)
x = self.conv2b_2b(x)
x = self.bn2b_2b(x)
x = self.activation2b_2b(x)
x = self.conv2b_2c(x)
x = self.bn2b_2c(x)
x = x[:,:,:-1] + res
x = self.activation2b(x)
res = x
x = self.conv2c_2a(x)
x = self.bn2c_2a(x)
x = self.activation2c_2a(x)
x = self.conv2c_2b(x)
x = self.bn2c_2b(x)
x = self.activation2c_2b(x)
x = self.conv2c_2c(x)
x = self.bn2c_2c(x)
x = x[:,:,:-1] + res
x = self.activation2c(x)
res = x
res = self.conv3a_1(res)
res = self.bn3a_1(res)
x = self.conv3a_2a(x)
x = self.bn3a_2a(x)
x = self.activation3a_2a(x)
x = self.conv3a_2b(x)
x = self.bn3a_2b(x)
x = self.activation3a_2b(x)
x = self.conv3a_2c(x)
x = self.bn3a_2c(x)
x = x[:,:,:-1] + res
x = self.activation3a(x)
res = x
x = self.conv3b_2a(x)
x = self.bn3b_2a(x)
x = self.activation3b_2a(x)
x = self.conv3b_2b(x)
x = self.bn3b_2b(x)
x = self.activation3b_2b(x)
x = self.conv3b_2c(x)
x = self.bn3b_2c(x)
x = x[:,:,:-1] + res
x = self.activation3b(x)
res = x
x = self.conv3c_2a(x)
x = self.bn3c_2a(x)
x = self.activation3c_2a(x)
x = self.conv3c_2b(x)
x = self.bn3c_2b(x)
x = self.activation3c_2b(x)
x = self.conv3c_2c(x)
x = self.bn3c_2c(x)
x = x[:,:,:-1] + res
x = self.activation3c(x)
res = x
x = self.conv3d_2a(x)
x = self.bn3d_2a(x)
x = self.activation3d_2a(x)
x = self.conv3d_2b(x)
x = self.bn3d_2b(x)
x = self.activation3d_2b(x)
x = self.conv3d_2c(x)
x = self.bn3d_2c(x)
x = x[:,:,:-1] + res
x = self.activation3d(x)
res = x
res = self.conv4a_1(res)
res = self.bn4a_1(res)
x = self.conv4a_2a(x)
x = self.bn4a_2a(x)
x = self.activation4a_2a(x)
x = self.conv4a_2b(x)
x = self.bn4a_2b(x)
x = self.activation4a_2b(x)
x = self.conv4a_2c(x)
x = self.bn4a_2c(x)
x = x[:,:,:-1] + res
x = self.activation4a(x)
res = x
x = self.conv4b_2a(x)
x = self.bn4b_2a(x)
x = self.activation4b_2a(x)
x = self.conv4b_2b(x)
x = self.bn4b_2b(x)
x = self.activation4b_2b(x)
x = self.conv4b_2c(x)
x = self.bn4b_2c(x)
x = x[:,:,:-1] + res
x = self.activation4b(x)
res = x
x = self.conv4c_2a(x)
x = self.bn4c_2a(x)
x = self.activation4c_2a(x)
x = self.conv4c_2b(x)
x = self.bn4c_2b(x)
x = self.activation4c_2b(x)
x = self.conv4c_2c(x)
x = self.bn4c_2c(x)
x = x[:,:,:-1] + res
x = self.activation4c(x)
res = x
x = self.conv4d_2a(x)
x = self.bn4d_2a(x)
x = self.activation4d_2a(x)
x = self.conv4d_2b(x)
x = self.bn4d_2b(x)
x = self.activation4d_2b(x)
x = self.conv4d_2c(x)
x = self.bn4d_2c(x)
x = x[:,:,:-1] + res
x = self.activation4d(x)
res = x
x = self.conv4e_2a(x)
x = self.bn4e_2a(x)
x = self.activation4e_2a(x)
x = self.conv4e_2b(x)
x = self.bn4e_2b(x)
x = self.activation4e_2b(x)
x = self.conv4e_2c(x)
x = self.bn4e_2c(x)
x = x[:,:,:-1] + res
x = self.activation4e(x)
res = x
x = self.conv4f_2a(x)
x = self.bn4f_2a(x)
x = self.activation4f_2a(x)
x = self.conv4f_2b(x)
x = self.bn4f_2b(x)
x = self.activation4f_2b(x)
x = self.conv4f_2c(x)
x = self.bn4f_2c(x)
x = x[:,:,:-1] + res
x = self.activation4f(x)
res = x
res = self.conv5a_1(res)
res = self.bn5a_1(res)
x = self.conv5a_2a(x)
x = self.bn5a_2a(x)
x = self.activation5a_2a(x)
x = self.conv5a_2b(x)
x = self.bn5a_2b(x)
x = self.activation5a_2b(x)
x = self.conv5a_2c(x)
x = self.bn5a_2c(x)
x = x[:,:,:-1] + res
x = self.activation5a(x)
res = x
x = self.conv5b_2a(x)
x = self.bn5b_2a(x)
x = self.activation5b_2a(x)
x = self.conv5b_2b(x)
x = self.bn5b_2b(x)
x = self.activation5b_2b(x)
x = self.conv5b_2c(x)
x = self.bn5b_2c(x)
x = x[:,:,:-1] + res
x = self.activation5b(x)
res = x
x = self.conv5c_2a(x)
x = self.bn5c_2a(x)
x = self.activation5c_2a(x)
x = self.conv5c_2b(x)
x = self.bn5c_2b(x)
x = self.activation5c_2b(x)
x = self.conv5c_2c(x)
x = self.bn5c_2c(x)
x = x[:,:,:-1] + res
x = self.activation5c(x)
x = self.AvgPool(x)
x = x.view(x.size(0), -1)
return x
model_CNN = CNN_1D()
class MyModel(nn.Module):
def __init__(self):
super(MyModel, self).__init__()
self.cnn = model_CNN
self.fc1 = nn.Linear(6144,4096)
self.activation_fc1 = nn.ReLU()
self.dropout1 = nn.Dropout(p=0.5)
self.fc2 = nn.Linear(4096,2048)
self.activation_fc2 = nn.ReLU()
self.dropout2 = nn.Dropout(p=0.5)
self.fc3 = nn.Linear(2048,1024)
self.activation_fc3 = nn.ReLU()
self.fc4 = nn.Linear(1024,2)
def forward(self, x1, x2):
x1 = self.cnn(x1)
x2 = self.cnn(x2)
x_add = x1+x2
x_minus = x1-x2
x_multiply = x1*x2
x = torch.cat((x_add, x_minus, x_multiply),-1)
x = self.fc1(x)
x = self.activation_fc1(x)
x = self.dropout1(x)
x = self.fc2(x)
x = self.activation_fc2(x)
x = self.dropout2(x)
x = self.fc3(x)
x = self.activation_fc3(x)
x = self.fc4(x)
return x
model = MyModel()
sample1 = torch.rand([1,1,256*256])
sample2 = torch.rand([1,1,256*256])
model(sample1,sample2).size()
sample1.size()
def random_sampling_and_normalization(sample,sampling_length=256*256,padding=10):
length = sample.size(1)
if length<sampling_length:
pad = int((sampling_length-length)/2)
sample = torch.cat((torch.zeros((1,pad)),sample,torch.zeros((1,pad))),-1)
sample = torch.cat((torch.zeros((1,padding)),sample,torch.zeros((1,padding))),-1)
length = sample.size(1)
random_num = np.random.randint(low=0, high=(length-sampling_length-1))
sample = sample[:,random_num:random_num+sampling_length]
#normalization
#channel=(channel-mean)/std
mean = torch.mean(sample)
std = torch.std(sample)
sample = (sample-mean)/std
return sample
def center_sampling_and_normalization(sample,sampling_length=256*256):
length = sample.size(1)
if length<sampling_length:
pad = int(sampling_length-length)
sample = torch.cat((sample,torch.zeros((1,pad))),-1)
sample = sample[:,:sampling_length]
#normalization
#channel=(channel-mean)/std
mean = torch.mean(sample)
std = torch.std(sample)
sample = (sample-mean)/std
return sample
class my_dataset(Dataset):
def __init__(self, df_path, train = False):
self.df = pd.read_csv(df_path)
self.train = train
def __len__(self):
return len(self.df)
def __getitem__(self,idx):
sample_1_name = self.df.iloc[idx]['sample 1']
sample_1_path = '/scratch/cz2064/myjupyter/Time_Series/Data/data_VoxCeleb/wav/'+sample_1_name
sample_1,_ = torchaudio.load(sample_1_path)
sample_2_name = self.df.iloc[idx]['sample 2']
sample_2_path = '/scratch/cz2064/myjupyter/Time_Series/Data/data_VoxCeleb/wav/'+sample_2_name
sample_2,_ = torchaudio.load(sample_2_path)
if self.train:
sample_1_tensor = random_sampling_and_normalization(sample_1)
sample_2_tensor = random_sampling_and_normalization(sample_2)
else:
sample_1_tensor = center_sampling_and_normalization(sample_1)
sample_2_tensor = center_sampling_and_normalization(sample_2)
label = self.df.loc[idx,'True or False']
label = torch.tensor(label, dtype=torch.long)
sample = {'x1': sample_1_tensor, 'x2': sample_2_tensor, 'y': label}
return sample
train_df_path = '/scratch/cz2064/myjupyter/Time_Series/notebook/train.csv'
val_df_path = '/scratch/cz2064/myjupyter/Time_Series/notebook/val.csv'
test_df_path = '/scratch/cz2064/myjupyter/Time_Series/notebook/test.csv'
BATCH_SIZE = 128
train_sampler = torch.utils.data.sampler.RandomSampler(my_dataset(train_df_path,train = True)\
,num_samples=50000,replacement=True)
train_loader = DataLoader(my_dataset(train_df_path,train = True), batch_size=BATCH_SIZE, \
sampler = train_sampler,num_workers=16)
val_loader = DataLoader(my_dataset(val_df_path), batch_size=BATCH_SIZE, shuffle=True,num_workers=16)
test_loader = DataLoader(my_dataset(test_df_path), batch_size=BATCH_SIZE, shuffle=True)
sample = next(iter(train_loader))
sample['y'].type()
# ## Train
# +
def train(model, train_loader=train_loader, val_loader=val_loader, learning_rate=5e-5, num_epoch=1000):
start_time = time.time()
loss_fn = nn.CrossEntropyLoss()
optimizer = optim.Adam(model.parameters(),lr=learning_rate)
train_loss_return = []
train_acc_return = []
val_loss_return = []
val_acc_return = []
best_acc = 0
for epoch in range(num_epoch):
# Training steps
correct = 0
total = 0
predictions = []
truths = []
model.train()
train_loss_list = []
for i, (sample) in enumerate(train_loader):
sample_1 = sample['x1'].to(device)
sample_2 = sample['x2'].to(device)
labels = sample['y'].to(device)
outputs = model(sample_1,sample_2)
pred = outputs.data.max(-1)[1]
predictions += list(pred.cpu().numpy())
truths += list(labels.cpu().numpy())
total += labels.size(0)
correct += (pred == labels).sum()
model.zero_grad()
loss = loss_fn(outputs, labels)
print(loss)
train_loss_list.append(loss.item())
loss.backward()
optimizer.step()
# report performance
acc = (100 * correct / total)
train_acc_return.append(acc)
train_loss_every_epoch = np.average(train_loss_list)
train_loss_return.append(train_loss_every_epoch)
print('----------Epoch{:2d}/{:2d}----------'.format(epoch+1,num_epoch))
print('Train set | Loss: {:6.4f} | Accuracy: {:4.2f}% '.format(train_loss_every_epoch, acc))
# Evaluate after every epochh
correct = 0
total = 0
model.eval()
predictions = []
truths = []
val_loss_list = []
with torch.no_grad():
for i, (sample) in enumerate(val_loader):
sample_1 = sample['x1'].to(device)
sample_2 = sample['x2'].to(device)
labels = sample['y'].to(device)
outputs = model(sample_1,sample_2)
loss = loss_fn(outputs, labels)
val_loss_list.append(loss.item())
pred = outputs.data.max(-1)[1]
predictions += list(pred.cpu().numpy())
truths += list(labels.cpu().numpy())
total += labels.size(0)
correct += (pred == labels).sum()
# report performance
acc = (100 * correct / total)
val_acc_return.append(acc)
val_loss_every_epoch = np.average(val_loss_list)
val_loss_return.append(val_loss_every_epoch)
if acc > best_acc:
best_acc = acc
best_model_wts = model.state_dict()
save_model(model,train_loss_return,train_acc_return,val_loss_return,val_acc_return,best_model_wts)
elapse = time.strftime('%H:%M:%S', time.gmtime(int((time.time() - start_time))))
print('Test set | Loss: {:6.4f} | Accuracy: {:4.2f}% | time elapse: {:>9}'\
.format(val_loss_every_epoch, acc,elapse))
return model,train_loss_return,train_acc_return,val_loss_return,val_acc_return,best_model_wts
def save_model(model,train_loss_return,train_acc_return,val_loss_return,val_acc_return,best_model_wts):
state = {'best_model_wts':best_model_wts, 'model':model, \
'train_loss':train_loss_return, 'train_acc':train_acc_return,\
'val_loss':val_loss_return, 'val_acc':val_acc_return}
torch.save(state, 'checkpoint_CNN.pt')
return None
# -
model = MyModel().to(device)
train(model)
# ## Evaluate
model = MyModel().to(device)
model.load_state_dict(torch.load('/scratch/cz2064/myjupyter/Time_Series/notebook/python_files/\
Model_CNN_50/Second Train/checkpoint_CNN.pt',map_location=torch.device(device))['best_model_wts'])
test_df_path = '/scratch/cz2064/myjupyter/Time_Series/notebook/test.csv'
test_loader = DataLoader(my_dataset(test_df_path), batch_size=1, shuffle=False)
def evaluate_model(model, dataloader):
model.eval()
y_true = []
y_score = []
y_pre = []
model.eval()
for i, (sample) in enumerate(dataloader):
sample_1 = sample['x1'].to(device)
sample_2 = sample['x2'].to(device)
label = sample['y'].to('cpu',dtype=torch.long)
y_true += label.tolist()
output = model(sample_1,sample_2)
output = F.softmax(output,dim=1)
output = output.to('cpu')
if y_score == []:
y_score = np.array(output.detach().numpy())
else:
y_score = np.concatenate((y_score,output.detach().numpy()),axis = 0)
for i in y_score:
y_pre.append(list(i).index(max(i)))
return y_true,y_pre,y_score
y_true,y_pre,y_score = evaluate_model(model, test_loader)
test_df = pd.read_csv(test_df_path)
test_df['pre'] = y_pre
test_df['score'] = np.expand_dims(np.array(y_score),2)[:,1]
test_df.head()
test_df.to_csv('Pre_ResNet_50.csv', index = False)
def ROC_curve(y_test,y_score):
fpr = dict()
tpr = dict()
roc_auc = dict()
n_classes = y_test.shape[1]
for i in range(n_classes):
fpr[i], tpr[i], _ = roc_curve(y_test[:, i], y_score[:, i])
roc_auc[i] = auc(fpr[i], tpr[i])
colors = cycle(['aqua', 'darkorange', 'cornflowerblue'])
lw = 2
for i, color in zip(range(n_classes), colors):
plt.plot(fpr[i], tpr[i], color=color, lw=lw,
label='ROC curve of class {0} (area = {1:0.2f})'
''.format(i, roc_auc[i]))
plt.plot([0, 1], [0, 1], 'k--', lw=lw)
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('Some extension of Receiver operating characteristic to multi-class')
plt.legend(loc="lower right")
plt.show()
return None
ROC_curve(np.expand_dims(np.array(y_true),1),np.expand_dims(np.array(y_score),2)[:,1])
|
3_CNN_50.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: genet
# language: python
# name: genet
# ---
# # Modifying the `Network` object
# `GeNet` supports some simple modifications like adding, reindexing and removing nodes and links and some involved modifications like changing the data stored under nodes or links. All of these changes get recorded in `n.change_log`.
# +
from genet import read_matsim
import os
path_to_matsim_network = '../example_data/pt2matsim_network'
network = os.path.join(path_to_matsim_network, 'network.xml')
schedule = os.path.join(path_to_matsim_network, 'schedule.xml')
vehicles = os.path.join(path_to_matsim_network, 'vehicles.xml')
n = read_matsim(
path_to_network=network,
epsg='epsg:27700',
path_to_schedule=schedule,
path_to_vehicles=vehicles
)
# you don't need to read the vehicles file, but doing so ensures all vehicles
# in the schedule are of the expected type and the definition of the vehicle
# is preserved
n.print()
# -
# ## Adding nodes/links
n.add_link(link_id='proposed_index', u='4356572310', v='5811263955')
n.add_node(node='proposed_index', attribs={'data':'some_data'})
# The index passed is only a proposition. If a node or link under this link exists, a new, unique index will be generated.
actual_link_id_added = n.add_link(link_id='proposed_index', u='4356572310', v='5811263955')
# ## Reindexing
#
# To reindex a node or link:
n.reindex_node('proposed_index', 'another_index')
n.reindex_link('proposed_index', 'another_index')
# ## Removing nodes/links
#
# To remove a link or node:
n.remove_links(['another_index', actual_link_id_added])
n.remove_node('another_index')
# ## Modifying data stored on nodes or edges:
#
# Let's say you have extracted `genet.Network` link ids of interest (See Section on Using Network - Accessing Data) and now you want to make changes to the network. Let's make changes to the nested OSM data stored on the links. We will replace the highway tags from `'primary'` to `'SOMETHING'`.
# +
from genet import graph_operations
links = n.extract_links_on_edge_attributes(
conditions= {'attributes': {'osm:way:highway': {'text': 'primary'}}},
)
links[:5]
# -
n.link(links[0])
# We create a dictionary which maps same changes to all links in the list using:
#
# ```python
# {link_id: {'attributes': {'osm:way:highway': {'text': 'SOMETHING'}}} for link_id in links}
# ```
n.apply_attributes_to_links({link_id: {'attributes': {'osm:way:highway': {'text': 'SOMETHING'}}} for link_id in links})
n.link(links[0])
n.change_log.head()
n.change_log.loc[618, :]['old_attributes']
n.change_log.loc[618, :]['new_attributes']
n.change_log.loc[618, :]['diff']
# Another useful method is the `apply_function_to_links`/`nodes`. This function takes a user-defined function with the variable that is the data dictionary stored on the links or nodes respectively. This function does not support `conditions`, but this is something that can be encoded in you function, for example:
# +
def divide_capacity(link_attributes):
if 'car' in link_attributes['modes']:
return link_attributes['capacity']/link_attributes['permlanes']
n.apply_function_to_links(divide_capacity, 'base_capacity')
# -
# This method will apply results of this function to links for which a value can be computed. For use here, that means that for any link which has a `'car'` value in `'modes'` a value of capacity divided by the number of lanes will be computed. We specify that the results will be stored under `base_capacity` in the links attribute dictionary.
n.link(links[0])
# # Modifying the `Schedule` object
#
# GeNet has a number of methods to change the Schedule objects. Make sure you validate the end result.
from genet import Stop, Route, Service
# ## Adding Routes, Services
#
# You can add `Route`s and `Service`s. To add a `Route`, you need to identify which existing `Service` it should come under.
# +
route = Route(
route_short_name='N55',
mode='bus',
trips={'trip_id': ['fun_trip_1', 'fun_trip_2'],
'trip_departure_time': ['03:53:00', '16:23:00'],
'vehicle_id': ['fun_bus_1', 'fun_bus_2']},
arrival_offsets=['00:00:00', '00:02:00', '00:04:00', '00:06:00'],
departure_offsets=['00:00:00', '00:02:00', '00:04:00', '00:06:00'],
id='new_route',
# route= ['834', '1573', '3139', '3141', '574', '3154', '979', '980', '981'],
await_departure= [True, True, True, True],
stops=[n.schedule.stop('490000235X.link:834'),
Stop(id='new_stop', x=529500, y=181300,
name='New Stop', epsg='epsg:27700'),
Stop(id='other_new_stop', x=529502, y=181302,
name='Other New Stop', epsg='epsg:27700'),
n.schedule.stop('490010689KB.link:981')]
)
n.schedule.add_route('20274', route)
# -
# Alternatively, you can create and add a new `Service`.
n.schedule.add_service(Service(id='new_service', routes=[route]))
# You can also refer to existing stops in the `Schedule` when creating a`Route` to be added. You can either just pass Stop IDs as strings or use a stop method on the schedule to take and use that stop object. Note that in the case of the former (passing ID strings), the route will not have the spatial information for those stops until it is added to the Schedule.
r = Route(
route_short_name='N55',
mode='bus',
trips={'trip_id': ['some_trip_1'],
'trip_departure_time': ['16:23:00'],
'vehicle_id': ['some_bus_2']},
arrival_offsets=['00:00:00', '00:06:00'],
departure_offsets=['00:00:00', '00:06:00'],
id='another_new_route',
# route= ['834', '1573', '3139', '3141', '574', '3154', '979', '980', '981'],
await_departure= [True, True],
stops=['490000235X.link:834',
'490010689KB.link:981']
)
r.graph().nodes['490000235X.link:834']
n.schedule.add_route('20274', r)
r.graph().nodes['490000235X.link:834']
r = Route(
route_short_name='N55',
mode='bus',
trips={'trip_id': ['some_trip_1'],
'trip_departure_time': ['16:23:00'],
'vehicle_id': ['some_bus_2']},
arrival_offsets=['00:00:00', '00:06:00'],
departure_offsets=['00:00:00', '00:06:00'],
id='another_new_route_2',
# route= ['834', '1573', '3139', '3141', '574', '3154', '979', '980', '981'],
await_departure= [True, True],
stops=[n.schedule.stop('490000235X.link:834'),
n.schedule.stop('490010689KB.link:981')]
)
n.schedule.add_route('20274', r)
# Note that for a Schedule to be a valid MATSim network, each stop referred to by a route needs a `linkRefId` attribute which links the stop to the `Network`.
#
# Trying to add Stops with IDs already in the Schedule will result in an error, unless the `force=True` is set. The added route/service will inherit the data stored under those stops. The idea is that you can either specify the Stops in the route/service to be added correctly, or if they are to be changed, you use a dedicated method - check section 'Modifying data' below.
# +
route = Route(
route_short_name='N55',
mode='bus',
trips={'trip_id': ['fun_trip_1', 'fun_trip_2'],
'trip_departure_time': ['03:53:00', '16:23:00'],
'vehicle_id': ['fun_bus_1', 'fun_bus_2']},
arrival_offsets=['00:00:00', '00:02:00', '00:04:00', '00:06:00'],
departure_offsets=['00:00:00', '00:02:00', '00:04:00', '00:06:00'],
id='another_new_route_3',
# route= ['834', '1573', '3139', '3141', '574', '3154', '979', '980', '981'],
await_departure= [True, True, True, True],
stops=[
Stop(id='490000235X.link:834', x=529981, y=181412, epsg='epsg:27700'),
Stop(id='new_stop', x=529500, y=181300, epsg='epsg:27700', name='New Stop'),
Stop(id='other_new_stop', x=529502, y=181302, epsg='epsg:27700', name='Other New Stop'),
Stop(id='490010689KB.link:981', x=529166, y=181256, epsg='epsg:27700')
]
)
n.schedule.add_route('20274', route, force=True)
# -
# Note the message above: `The following stops will inherit the data currently stored under those Stop IDs in the Schedule: ['490000235X.link:834', '490010689KB.link:981'].`
#
# NOTE: adding routes and services results in new vehicles (unless you reuse the ones already in the Schedule---beware that the same vehicle cannot service multiple trips at the same time, genet does not currently have checks for this, the user needs to be mindful of the physics of shared vehicles). New vehicles need definitions, you can add them yourself to `schedule.vehicles['vehicle_id'] = {'type': 'bus'}` ensuring this vehicle type is defined in `schedule.vehicle_types['bus']`, or you can use a genet method to generate those vehicles, the type will be derived from the mode of the route. Then you can check if all of the types that vehicles are referring to have definitions.
len(n.schedule.vehicles)
n.schedule.generate_vehicles()
n.schedule.validate_vehicle_definitions()
len(n.schedule.vehicles)
n.schedule.change_log().head()
# There are no methods to add `Stop` objects on their own. They are added to the Schedule with `Route` and `Service` objects.
# ## Reindexing
n.schedule['new_service'].reindex(new_id='more_appropriate_id')
n.schedule.route('new_route').reindex(new_id='more_appropriate_route_id')
n.schedule.change_log().head()
# ## Removing Stops, Routes, Services
n.schedule.remove_service('more_appropriate_id')
n.schedule.remove_route('more_appropriate_route_id')
for route_id in {'another_new_route', 'another_new_route_2', 'another_new_route_3'}:
n.schedule.remove_route(route_id)
n.schedule.change_log().tail()
# You can also remove `Stop`s. This will disconnect `Route`s and `Service`s using that `Stop`s and likely render them invalid. The method will warn you which `Route`s and `Servce`s are affected.
n.schedule.remove_stop('new_stop')
# Likewise, removing `Route`s and `Service`s is likely to result in a number of unused stops. There is a method to remove such stops.
n.schedule.remove_unsused_stops()
# ## Modifying data stored for Stops, Routes, Services
#
# ### Applying known or pre-computed changes
# Applying changes or new attributes to Services, Routes and Stops can be done via Schedule level methods. They all work with a dictionary where the keys are the object IDs and the values are dictionaries holding attribute names and values. The method to extract a DataFrame on attributes comes in handy here. E.g.
df = n.schedule.service_attribute_data(keys='name')
df.head()
# DataFrames are easy to work with. Youcould for exmaple manipulate the names or use other data to change these. For demonstration here, let's just set the names to something easy.
df['name'] = df['name'].apply(lambda x: f'Service_{x}')
df.head()
# You can then convert this to a dictionary and pass it to the `apply_attributes_to_services` method.
n.schedule.apply_attributes_to_services(df.T.to_dict())
n.schedule.change_log().tail()
# You can do the same for `Routes` and `Stops`. Your dictionaries cannot however hold changes to indices. You will encounter an error and should use `reindex` methods for such operations.
n.schedule.apply_attributes_to_routes(
{'VJ375a660d47a2aa570aa20a8568012da8497ffecf': {
'name': 'my_favourite_route',
'mode': 'piggyback'
}}
)
n.schedule.apply_attributes_to_stops(
{'490000235YB.link:574': {'new_attribute': 'hello!'}}
)
n.schedule.change_log().tail()
# You can use `route_trips_to_dataframe` to extract all of the trips, their departures and vehicle IDs associated with the trips in the schedule. Trip ids need not be unique, route IDs provide a secondary index. Associated service IDs are also given for convenience.
trips = n.schedule.route_trips_to_dataframe(gtfs_day='20210101')
trips.head()
# Let's change all of the trip ids to something shorter
trips['trip_id'] = 'trip_' + trips.index.to_series().astype(str)
trips.head()
# You can `set_route_trips_dataframe` which takes this dataframe and applies changes to all route trips based on the data in the dataframe. This means you can generate this DataFrame as shown below, manipulate trips (delete them, add new ones), change their departure times or change their vehicle ids to be shared for differnt trips, perhaps on some temporal logic and as long as the dataframe has the same schema, you can use it to set new trips in the schedule. This will appear in the changelog as a route level modify event.
#
# Nb removing all trips of the same route from the dataframe will have no effect when being applied. If there is data in the dataframe for a route, all of its trips will be replaced by the data in the dataframe, and if there is no data for a route in the frame, no changes will be applied to that route (i.e. the trips attribute for routes missing from the dataframe will not be set as empty).
n.schedule.set_route_trips_dataframe(trips)
n.schedule.route_attribute_data(keys=[{'trips': 'trip_id'}])
# ### Applying changes using functions or mappings
#
# If you have some logic that can be written into a function of object's attributes, you can pass this to `apply_function_to_x` methods. You need to select `location`, which refers to the name of the attribute the result should be stored under. It can already exist and be overwritten. The function passed, is not expected to work with all objects. It will fail silently, only evaluating and generating outputs where possible.
# +
from shapely.geometry import Point
def add_shapely_geometry_points(stop_attribs):
return Point(stop_attribs['x'], stop_attribs['y'])
n.schedule.apply_function_to_stops(add_shapely_geometry_points, location='geometry')
# -
n.schedule.change_log().tail(2)
n.schedule.stop_attribute_data(keys=['name', 'x', 'y', 'geometry']).head()
from geopandas import GeoDataFrame
GeoDataFrame(n.schedule.stop_attribute_data(keys='geometry')).plot()
n.schedule.stop('490000235YB.link:574').__dict__
# For this let's say we want to reduce the number of trips. For simplicity of demonstration we don't have about which trips we delete, but logic around timings of trips can be added in this function, as the trips are saved as one of routes attributes (check out the summary methods)
# before
len(n.schedule.route_trips_to_dataframe())
# +
def reduce_trips(attribs):
# just delete any other trip
attribs['trips']['trip_id'] = attribs['trips']['trip_id'][::2]
attribs['trips']['trip_departure_time'] = attribs['trips']['trip_departure_time'][::2]
attribs['trips']['vehicle_id'] = attribs['trips']['vehicle_id'][::2]
return attribs['trips']
n.schedule.apply_function_to_routes(reduce_trips, 'trips')
# -
# after
len(n.schedule.route_trips_to_dataframe())
# Note, this could also be done using the `route_trips_to_dataframe` and `set_route_trips_dataframe` mentioned above.
#
# Let's give an example of using a mapping. We can re-use the service name DataFrame we generated above.
df['new_name'] = 'Brand_new_name' + df['name']
df.head()
name_map = dict(zip(df['name'], df['new_name']))
name_map
# In this case, `location` refers to the attribute to be mapped.
n.schedule.apply_function_to_services(name_map, location='name')
n.schedule.change_log().tail()
|
notebooks/5.1. Modifying Network - Basics.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ## Test a class for the easy retrieval of postgres data via Jupyter widgets
#
# the following two lines indicate that external functions are auto-reloaded as soon as they change.
# %load_ext autoreload
# %autoreload 2
# Print statements
from __future__ import print_function # Python 2.x
# +
# General stuff:
import sys
import argparse
import os
import json
import numpy as np
import math
import psycopg2
import cPickle
import numpy as np
import pandas as pd
from datetime import date
from tqdm import tqdm_notebook
# Plotting:
from matplotlib import pyplot as plt
import seaborn as sns
import matplotlib.cm as cm
import matplotlib as mpl
# %matplotlib inline
# External functions from subfolder /database_helpers.
# as soon as you change something in there and press save, it will auto reload on next execution.
from database_helpers.psql_start import *
from database_helpers.create_tables import *
from database_helpers.write2tables import *
from postgres_analysis_helpers.general import *
from postgres_analysis_helpers.get_data import *
# register pickle type to retrieve binary data from database
psycopg2.extensions.register_type(psycopg2.extensions.new_type(psycopg2.BINARY.values, 'BINARY-PICKLE', cast_pickle))
# -
db_status = test_connect()
if db_status == False:
print('Grrr... no database connection could be established.')
else:
print('Yippiyeah! Database connection is established!')
# ### Make an empty dataframe and call the retrieval function
base_dataframe = pd.DataFrame() # create empty dataframe or feed in a base dataframe from before!
psql = psql_neuroballs(base_dataframe)
psql.retrieve_data()
base_dataframe = psql.data()
base_dataframe.columns
base_dataframe = base_dataframe[(base_dataframe.mvl > 0.35) & (base_dataframe.gridscore > 0.35)]
base_dataframe.sort_values(by='gridscore', ascending=False,inplace=True)
len(base_dataframe)
# ### get individual cells ...
dict1 = {}
dict1['animal_id'] = '70113'
dict1['n_drive_user'] = 'davidcr'
dict1['session_name'] = '02022017s1'
dict1['tetrode_no'] = 6
dict1['cluster_no'] = 36
params = config()
dictio = dict1
#sql_cmd = "SELECT * FROM BNT_tb_screen WHERE animal_id = '{}' AND n_drive_user = '{}' AND session_name = '{}' and tetrode_no = {} and cluster_no = {};".format(dictio['animal_id'],dictio['n_drive_user'],
sql_cmd = "SELECT * FROM BNT_tb_screen WHERE animal_id = '{}' AND n_drive_user = '{}';".format(dictio['animal_id'],dictio['n_drive_user'])
sql_db_pd = pd.read_sql_query(sql_cmd, psycopg2.connect(**params), index_col=None,parse_dates=['session_ts','analysis_ts'])
sql_db_pd = sql_db_pd[sql_db_pd.session_name=='01022017s1']
draw_ratemaps(base_dataframe,'masked_ratemap',19,50)
draw_path_spike_plots(base_dataframe,base_dataframe,19,50)
draw_hd_tuning_curves(base_dataframe,19,50)
figure= plt.figure(figsize=(5,5))
sns.set_style('white')
plt.imshow(base_dataframe.masked_ratemap[0],interpolation='nearest',cmap='jet')
base_dataframe.columns
inhib_p_thresh = 0.001
base_dataframe = base_dataframe.dropna(subset=['tetrode_no','cluster_no'])
# sort dataframe!
base_dataframe.sort_values('inhib_lowest_p',ascending=True, inplace=True)
# drop duplicates (maintaining the most significant / highest inhibited session)
base_dataframe.drop_duplicates(subset=['session_ts','animal_id','tetrode_no','cluster_no'],inplace=True)
# reset indices
base_dataframe.reset_index(drop=True,inplace=True)
# refresh "inhibited" label column based on inhib_p_thresh (see above)
base_dataframe['inhibited'] = (base_dataframe.inhib_lowest_p < inhib_p_thresh) & (base_dataframe.inhibited == True)
len(base_dataframe)
psql = psql_neuroballs(base_dataframe)
psql.retrieve_data()
base_dataframe = psql.data()
len(base_dataframe)
# ### Now you have a filtered base dataset with one session per cluster
# Use this to retrieve the base session data
# drop session_name column
base_dataframe = base_dataframe.loc[:,~base_dataframe.columns.duplicated()]
base_dataframe.drop('session_name', 1,inplace=True)
len(base_dataframe)
# +
#create_spike_plots_stimulus(base_dataframe,25,[5,5])
# -
basesessions_ = psql_neuroballs(base_dataframe)
basesessions_.retrieve_data()
# get the new dataframe:
basesessions_df = basesessions_.data()
len(basesessions_df)
draw_ratemaps(basesessions_df,'masked_ratemap',25,0)
draw_hd_tuning_curves(basesessions_df,25,0)
# ### Save dataset for later retrieval
export_path_pickle = r"C:\work\python\klusta_analysis\postgres_notebooks\export_dataframes"
export_path_pickle = "/".join(export_path_pickle.split("\\"))
#basesessions_df.to_pickle(export_path_pickle + "/dave_arch.pkl")
# #### and retrieve
basesessions_df = pd.read_pickle(export_path_pickle + "/dave_arch.pkl")
len(basesessions_df)
basesessions_ = psql_neuroballs(basesessions_df)
basesessions_.retrieve_data()
basesessions_df = basesessions_.data()
basesessions_df.to_pickle(export_path_pickle + "/dave_arch.pkl")
|
Notebook archive/psql_neuroballs_tests.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# This IPython Notebook introduces the use of the `openmc.mgxs` module to calculate multi-group cross sections for an infinite homogeneous medium. In particular, this Notebook introduces the the following features:
#
# * **General equations** for scalar-flux averaged multi-group cross sections
# * Creation of multi-group cross sections for an **infinite homogeneous medium**
# * Use of **tally arithmetic** to manipulate multi-group cross sections
# ## Introduction to Multi-Group Cross Sections (MGXS)
# Many Monte Carlo particle transport codes, including OpenMC, use continuous-energy nuclear cross section data. However, most deterministic neutron transport codes use *multi-group cross sections* defined over discretized energy bins or *energy groups*. An example of U-235's continuous-energy fission cross section along with a 16-group cross section computed for a light water reactor spectrum is displayed below.
from IPython.display import Image
Image(filename='images/mgxs.png', width=350)
# A variety of tools employing different methodologies have been developed over the years to compute multi-group cross sections for certain applications, including NJOY (LANL), MC$^2$-3 (ANL), and Serpent (VTT). The `openmc.mgxs` Python module is designed to leverage OpenMC's tally system to calculate multi-group cross sections with arbitrary energy discretizations for fine-mesh heterogeneous deterministic neutron transport applications.
#
# Before proceeding to illustrate how one may use the `openmc.mgxs` module, it is worthwhile to define the general equations used to calculate multi-group cross sections. This is only intended as a brief overview of the methodology used by `openmc.mgxs` - we refer the interested reader to the large body of literature on the subject for a more comprehensive understanding of this complex topic.
# ### Introductory Notation
# The continuous real-valued microscopic cross section may be denoted $\sigma_{n,x}(\mathbf{r}, E)$ for position vector $\mathbf{r}$, energy $E$, nuclide $n$ and interaction type $x$. Similarly, the scalar neutron flux may be denoted by $\Phi(\mathbf{r},E)$ for position $\mathbf{r}$ and energy $E$. **Note**: Although nuclear cross sections are dependent on the temperature $T$ of the interacting medium, the temperature variable is neglected here for brevity.
# ### Spatial and Energy Discretization
# The energy domain for critical systems such as thermal reactors spans more than 10 orders of magnitude of neutron energies from 10$^{-5}$ - 10$^7$ eV. The multi-group approximation discretization divides this energy range into one or more energy groups. In particular, for $G$ total groups, we denote an energy group index $g$ such that $g \in \{1, 2, ..., G\}$. The energy group indices are defined such that the smaller group the higher the energy, and vice versa. The integration over neutron energies across a discrete energy group is commonly referred to as **energy condensation**.
#
# Multi-group cross sections are computed for discretized spatial zones in the geometry of interest. The spatial zones may be defined on a structured and regular fuel assembly or pin cell mesh, an arbitrary unstructured mesh or the constructive solid geometry used by OpenMC. For a geometry with $K$ distinct spatial zones, we designate each spatial zone an index $k$ such that $k \in \{1, 2, ..., K\}$. The volume of each spatial zone is denoted by $V_{k}$. The integration over discrete spatial zones is commonly referred to as **spatial homogenization**.
# ### General Scalar-Flux Weighted MGXS
# The multi-group cross sections computed by `openmc.mgxs` are defined as a *scalar flux-weighted average* of the microscopic cross sections across each discrete energy group. This formulation is employed in order to preserve the reaction rates within each energy group and spatial zone. In particular, spatial homogenization and energy condensation are used to compute the general multi-group cross section $\sigma_{n,x,k,g}$ as follows:
#
# $$\sigma_{n,x,k,g} = \frac{\int_{E_{g}}^{E_{g-1}}\mathrm{d}E'\int_{\mathbf{r} \in V_{k}}\mathrm{d}\mathbf{r}\sigma_{n,x}(\mathbf{r},E')\Phi(\mathbf{r},E')}{\int_{E_{g}}^{E_{g-1}}\mathrm{d}E'\int_{\mathbf{r} \in V_{k}}\mathrm{d}\mathbf{r}\Phi(\mathbf{r},E')}$$
#
# This scalar flux-weighted average microscopic cross section is computed by `openmc.mgxs` for most multi-group cross sections, including total, absorption, and fission reaction types. These double integrals are stochastically computed with OpenMC's tally system - in particular, [filters](https://mit-crpg.github.io/openmc/pythonapi/filter.html) on the energy range and spatial zone (material, cell or universe) define the bounds of integration for both numerator and denominator.
# ### Multi-Group Scattering Matrices
# The general multi-group cross section $\sigma_{n,x,k,g}$ is a vector of $G$ values for each energy group $g$. The equation presented above only discretizes the energy of the incoming neutron and neglects the outgoing energy of the neutron (if any). Hence, this formulation must be extended to account for the outgoing energy of neutrons in the discretized scattering matrix cross section used by deterministic neutron transport codes.
#
# We denote the incoming and outgoing neutron energy groups as $g$ and $g'$ for the microscopic scattering matrix cross section $\sigma_{n,s}(\mathbf{r},E)$. As before, spatial homogenization and energy condensation are used to find the multi-group scattering matrix cross section $\sigma_{n,s,k,g \to g'}$ as follows:
#
# $$\sigma_{n,s,k,g\rightarrow g'} = \frac{\int_{E_{g'}}^{E_{g'-1}}\mathrm{d}E''\int_{E_{g}}^{E_{g-1}}\mathrm{d}E'\int_{\mathbf{r} \in V_{k}}\mathrm{d}\mathbf{r}\sigma_{n,s}(\mathbf{r},E'\rightarrow E'')\Phi(\mathbf{r},E')}{\int_{E_{g}}^{E_{g-1}}\mathrm{d}E'\int_{\mathbf{r} \in V_{k}}\mathrm{d}\mathbf{r}\Phi(\mathbf{r},E')}$$
#
# This scalar flux-weighted multi-group microscopic scattering matrix is computed using OpenMC tallies with both energy in and energy out filters.
# ### Multi-Group Fission Spectrum
# The energy spectrum of neutrons emitted from fission is denoted by $\chi_{n}(\mathbf{r},E' \rightarrow E'')$ for incoming and outgoing energies $E'$ and $E''$, respectively. Unlike the multi-group cross sections $\sigma_{n,x,k,g}$ considered up to this point, the fission spectrum is a probability distribution and must sum to unity. The outgoing energy is typically much less dependent on the incoming energy for fission than for scattering interactions. As a result, it is common practice to integrate over the incoming neutron energy when computing the multi-group fission spectrum. The fission spectrum may be simplified as $\chi_{n}(\mathbf{r},E)$ with outgoing energy $E$.
#
# Unlike the multi-group cross sections defined up to this point, the multi-group fission spectrum is weighted by the fission production rate rather than the scalar flux. This formulation is intended to preserve the total fission production rate in the multi-group deterministic calculation. In order to mathematically define the multi-group fission spectrum, we denote the microscopic fission cross section as $\sigma_{n,f}(\mathbf{r},E)$ and the average number of neutrons emitted from fission interactions with nuclide $n$ as $\nu_{n}(\mathbf{r},E)$. The multi-group fission spectrum $\chi_{n,k,g}$ is then the probability of fission neutrons emitted into energy group $g$.
#
# Similar to before, spatial homogenization and energy condensation are used to find the multi-group fission spectrum $\chi_{n,k,g}$ as follows:
#
# $$\chi_{n,k,g'} = \frac{\int_{E_{g'}}^{E_{g'-1}}\mathrm{d}E''\int_{0}^{\infty}\mathrm{d}E'\int_{\mathbf{r} \in V_{k}}\mathrm{d}\mathbf{r}\chi_{n}(\mathbf{r},E'\rightarrow E'')\nu_{n}(\mathbf{r},E')\sigma_{n,f}(\mathbf{r},E')\Phi(\mathbf{r},E')}{\int_{0}^{\infty}\mathrm{d}E'\int_{\mathbf{r} \in V_{k}}\mathrm{d}\mathbf{r}\nu_{n}(\mathbf{r},E')\sigma_{n,f}(\mathbf{r},E')\Phi(\mathbf{r},E')}$$
#
# The fission production-weighted multi-group fission spectrum is computed using OpenMC tallies with both energy in and energy out filters.
#
# This concludes our brief overview on the methodology to compute multi-group cross sections. The following sections detail more concretely how users may employ the `openmc.mgxs` module to power simulation workflows requiring multi-group cross sections for downstream deterministic calculations.
# ## Generate Input Files
# +
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import openmc
import openmc.mgxs as mgxs
# -
# First we need to define materials that will be used in the problem. Before defining a material, we must create nuclides that are used in the material.
# Instantiate some Nuclides
h1 = openmc.Nuclide('H1')
o16 = openmc.Nuclide('O16')
u235 = openmc.Nuclide('U235')
u238 = openmc.Nuclide('U238')
zr90 = openmc.Nuclide('Zr90')
# With the nuclides we defined, we will now create a material for the homogeneous medium.
# Instantiate a Material and register the Nuclides
inf_medium = openmc.Material(name='moderator')
inf_medium.set_density('g/cc', 5.)
inf_medium.add_nuclide(h1, 0.028999667)
inf_medium.add_nuclide(o16, 0.01450188)
inf_medium.add_nuclide(u235, 0.000114142)
inf_medium.add_nuclide(u238, 0.006886019)
inf_medium.add_nuclide(zr90, 0.002116053)
# With our material, we can now create a `Materials` object that can be exported to an actual XML file.
# Instantiate a Materials collection and export to XML
materials_file = openmc.Materials([inf_medium])
materials_file.export_to_xml()
# Now let's move on to the geometry. This problem will be a simple square cell with reflective boundary conditions to simulate an infinite homogeneous medium. The first step is to create the outer bounding surfaces of the problem.
# Instantiate boundary Planes
min_x = openmc.XPlane(boundary_type='reflective', x0=-0.63)
max_x = openmc.XPlane(boundary_type='reflective', x0=0.63)
min_y = openmc.YPlane(boundary_type='reflective', y0=-0.63)
max_y = openmc.YPlane(boundary_type='reflective', y0=0.63)
# With the surfaces defined, we can now create a cell that is defined by intersections of half-spaces created by the surfaces.
# +
# Instantiate a Cell
cell = openmc.Cell(cell_id=1, name='cell')
# Register bounding Surfaces with the Cell
cell.region = +min_x & -max_x & +min_y & -max_y
# Fill the Cell with the Material
cell.fill = inf_medium
# -
# OpenMC requires that there is a "root" universe. Let us create a root universe and add our square cell to it.
# Instantiate Universe
root_universe = openmc.Universe(universe_id=0, name='root universe')
root_universe.add_cell(cell)
# We now must create a geometry that is assigned a root universe and export it to XML.
# +
# Create Geometry and set root Universe
openmc_geometry = openmc.Geometry()
openmc_geometry.root_universe = root_universe
# Export to "geometry.xml"
openmc_geometry.export_to_xml()
# -
# Next, we must define simulation parameters. In this case, we will use 10 inactive batches and 40 active batches each with 2500 particles.
# +
# OpenMC simulation parameters
batches = 50
inactive = 10
particles = 2500
# Instantiate a Settings object
settings_file = openmc.Settings()
settings_file.batches = batches
settings_file.inactive = inactive
settings_file.particles = particles
settings_file.output = {'tallies': True}
# Create an initial uniform spatial source distribution over fissionable zones
bounds = [-0.63, -0.63, -0.63, 0.63, 0.63, 0.63]
uniform_dist = openmc.stats.Box(bounds[:3], bounds[3:], only_fissionable=True)
settings_file.source = openmc.source.Source(space=uniform_dist)
# Export to "settings.xml"
settings_file.export_to_xml()
# -
# Now we are ready to generate multi-group cross sections! First, let's define a 2-group structure using the built-in `EnergyGroups` class.
# Instantiate a 2-group EnergyGroups object
groups = mgxs.EnergyGroups()
groups.group_edges = np.array([0., 0.625, 20.0e6])
# We can now use the `EnergyGroups` object, along with our previously created materials and geometry, to instantiate some `MGXS` objects from the `openmc.mgxs` module. In particular, the following are subclasses of the generic and abstract `MGXS` class:
#
# * `TotalXS`
# * `TransportXS`
# * `AbsorptionXS`
# * `CaptureXS`
# * `FissionXS`
# * `KappaFissionXS`
# * `ScatterXS`
# * `ScatterMatrixXS`
# * `Chi`
# * `ChiPrompt`
# * `InverseVelocity`
# * `PromptNuFissionXS`
#
# Of course, we are aware that the fission cross section (`FissionXS`) can sometimes be paired with the fission neutron multiplication to become $\nu\sigma_f$. This can be accomodated in to the `FissionXS` class by setting the `nu` parameter to `True` as shown below.
#
# Additionally, scattering reactions (like (n,2n)) can also be defined to take in to account the neutron multiplication to become $\nu\sigma_s$. This can be accomodated in the the transport (`TransportXS`), scattering (`ScatterXS`), and scattering-matrix (`ScatterMatrixXS`) cross sections types by setting the `nu` parameter to `True` as shown below.
#
# These classes provide us with an interface to generate the tally inputs as well as perform post-processing of OpenMC's tally data to compute the respective multi-group cross sections. In this case, let's create the multi-group total, absorption and scattering cross sections with our 2-group structure.
# +
# Instantiate a few different sections
total = mgxs.TotalXS(domain=cell, groups=groups)
absorption = mgxs.AbsorptionXS(domain=cell, groups=groups)
scattering = mgxs.ScatterXS(domain=cell, groups=groups)
# Note that if we wanted to incorporate neutron multiplication in the
# scattering cross section we would write the previous line as:
# scattering = mgxs.ScatterXS(domain=cell, groups=groups, nu=True)
# -
# Each multi-group cross section object stores its tallies in a Python dictionary called `tallies`. We can inspect the tallies in the dictionary for our `Absorption` object as follows.
absorption.tallies
# The `Absorption` object includes tracklength tallies for the 'absorption' and 'flux' scores in the 2-group structure in cell 1. Now that each `MGXS` object contains the tallies that it needs, we must add these tallies to a `Tallies` object to generate the "tallies.xml" input file for OpenMC.
# +
# Instantiate an empty Tallies object
tallies_file = openmc.Tallies()
# Add total tallies to the tallies file
tallies_file += total.tallies.values()
# Add absorption tallies to the tallies file
tallies_file += absorption.tallies.values()
# Add scattering tallies to the tallies file
tallies_file += scattering.tallies.values()
# Export to "tallies.xml"
tallies_file.export_to_xml()
# -
# Now we a have a complete set of inputs, so we can go ahead and run our simulation.
# Run OpenMC
openmc.run()
# ## Tally Data Processing
# Our simulation ran successfully and created statepoint and summary output files. We begin our analysis by instantiating a `StatePoint` object.
# Load the last statepoint file
sp = openmc.StatePoint('statepoint.50.h5')
# In addition to the statepoint file, our simulation also created a summary file which encapsulates information about the materials and geometry. By default, a `Summary` object is automatically linked when a `StatePoint` is loaded. This is necessary for the `openmc.mgxs` module to properly process the tally data.
# The statepoint is now ready to be analyzed by our multi-group cross sections. We simply have to load the tallies from the `StatePoint` into each object as follows and our `MGXS` objects will compute the cross sections for us under-the-hood.
# Load the tallies from the statepoint into each MGXS object
total.load_from_statepoint(sp)
absorption.load_from_statepoint(sp)
scattering.load_from_statepoint(sp)
# Voila! Our multi-group cross sections are now ready to rock 'n roll!
# ## Extracting and Storing MGXS Data
# Let's first inspect our total cross section by printing it to the screen.
total.print_xs()
# Since the `openmc.mgxs` module uses [tally arithmetic](https://mit-crpg.github.io/openmc/pythonapi/examples/tally-arithmetic.html) under-the-hood, the cross section is stored as a "derived" `Tally` object. This means that it can be queried and manipulated using all of the same methods supported for the `Tally` class in the OpenMC Python API. For example, we can construct a [Pandas](http://pandas.pydata.org/) `DataFrame` of the multi-group cross section data.
df = scattering.get_pandas_dataframe()
df.head(10)
# Each multi-group cross section object can be easily exported to a variety of file formats, including CSV, Excel, and LaTeX for storage or data processing.
absorption.export_xs_data(filename='absorption-xs', format='excel')
# The following code snippet shows how to export all three `MGXS` to the same HDF5 binary data store.
total.build_hdf5_store(filename='mgxs', append=True)
absorption.build_hdf5_store(filename='mgxs', append=True)
scattering.build_hdf5_store(filename='mgxs', append=True)
# ## Comparing MGXS with Tally Arithmetic
# Finally, we illustrate how one can leverage OpenMC's [tally arithmetic](https://mit-crpg.github.io/openmc/pythonapi/examples/tally-arithmetic.html) data processing feature with `MGXS` objects. The `openmc.mgxs` module uses tally arithmetic to compute multi-group cross sections with automated uncertainty propagation. Each `MGXS` object includes an `xs_tally` attribute which is a "derived" `Tally` based on the tallies needed to compute the cross section type of interest. These derived tallies can be used in subsequent tally arithmetic operations. For example, we can use tally artithmetic to confirm that the `TotalXS` is equal to the sum of the `AbsorptionXS` and `ScatterXS` objects.
# +
# Use tally arithmetic to compute the difference between the total, absorption and scattering
difference = total.xs_tally - absorption.xs_tally - scattering.xs_tally
# The difference is a derived tally which can generate Pandas DataFrames for inspection
difference.get_pandas_dataframe()
# -
# Similarly, we can use tally arithmetic to compute the ratio of `AbsorptionXS` and `ScatterXS` to the `TotalXS`.
# +
# Use tally arithmetic to compute the absorption-to-total MGXS ratio
absorption_to_total = absorption.xs_tally / total.xs_tally
# The absorption-to-total ratio is a derived tally which can generate Pandas DataFrames for inspection
absorption_to_total.get_pandas_dataframe()
# +
# Use tally arithmetic to compute the scattering-to-total MGXS ratio
scattering_to_total = scattering.xs_tally / total.xs_tally
# The scattering-to-total ratio is a derived tally which can generate Pandas DataFrames for inspection
scattering_to_total.get_pandas_dataframe()
# -
# Lastly, we sum the derived scatter-to-total and absorption-to-total ratios to confirm that they sum to unity.
# +
# Use tally arithmetic to ensure that the absorption- and scattering-to-total MGXS ratios sum to unity
sum_ratio = absorption_to_total + scattering_to_total
# The scattering-to-total ratio is a derived tally which can generate Pandas DataFrames for inspection
sum_ratio.get_pandas_dataframe()
|
examples/jupyter/mgxs-part-i.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import pdb
import os
import json
import numpy as np
import matplotlib.pyplot as plt
colors = ['#F0080F','#0A00D7','#00F0F0','#700FD4','#F000F0','#00050C', '#00E404','#F00F00','#006347','#00C00B', \
'#F00500','#008000','#0000FF','#F0F0F0','#7C0C00','#E0F00F','#00B000','#000FF0','#ADD806', '#320032', \
'#48000C', '#C00085','#F5000A', '#00E4B5','#0000E6','#0070D6','#D00003','#DD0000','#FF0000','#2E0057',]
IOU_THRESHOLD=0.5
path_dt = 'saved_model/cross_validation/'
path_gt = 'saved_model/val_gt.json'
plt.rcParams['figure.dpi'] = 300
plt.figure(1)
plt.title('Miss Rate')
plt.xlabel('FPPI')
plt.ylabel('miss rate')
list_epoch=list()
list_mr=list()
total_fppi_list=list()
total_miss_list=list()
total_p_list=list()
total_r_list=list()
ap=list()
res_dt=sorted(os.listdir(path_dt))
for dt_name in res_dt:
with open(path_gt, 'r') as g:
gt_all=json.load(g)
with open(path_dt + dt_name, 'r') as d:
dt=json.load(d)
total_fg = 0
gt_dict={}
for i in range(len(gt_all['annotations'])):
if gt_all['annotations'][i]['vis_ratio']>0.65 and gt_all['annotations'][i]['ignore']==0 and gt_all['annotations'][i]['height']>50:
total_fg+=1
if int(gt_all['annotations'][i]['image_id']) in gt_dict:
gt_dict[int(gt_all['annotations'][i]['image_id'])].append(gt_all['annotations'][i])
else:
gt_dict[int(gt_all['annotations'][i]['image_id'])]=[]
gt_dict[int(gt_all['annotations'][i]['image_id'])].append(gt_all['annotations'][i])
dt_sorted = sorted(dt, key=lambda k: k['score'], reverse=True)
false_num=0
miss_num=total_fg
true_num=0
total_fppi=[]
total_miss=[]
p=[]
r=[]
def IOU(dt):
if int(dt['image_id']) not in gt_dict:
return 'fp'
gt = gt_dict[dt['image_id']]
iou=list()
for i in range(len(gt)):
inter_xmin = max(dt['bbox'][0], gt[i]['bbox'][0])
inter_ymin = max(dt['bbox'][1], gt[i]['bbox'][1])
inter_xmax = min(dt['bbox'][0]+dt['bbox'][2], gt[i]['bbox'][0]+gt[i]['bbox'][2])
inter_ymax = min(dt['bbox'][1]+dt['bbox'][3], gt[i]['bbox'][1]+gt[i]['bbox'][3])
Iw = max(inter_xmax - inter_xmin, 0)
Ih = max(inter_ymax - inter_ymin, 0)
I = Iw * Ih
U = dt['bbox'][2]*dt['bbox'][3] + gt[i]['bbox'][2]*gt[i]['bbox'][3] - I
iou.append(float(I)/float(U))
return iou
def match(dt_sorted):
global gt_dict
iou = IOU(dt_sorted)
if iou == 'fp':
return 'fp'
iou_index = np.argsort(-np.array(iou))
iou_sorted = sorted(iou, reverse=True)
iou_fg=list()
for i in range(len(iou_sorted)):
if iou_sorted[i]>IOU_THRESHOLD:
iou_fg.append(iou_sorted[i])
else:
break
if len(iou_fg)<1:
return 'fp'
for i in range(len(iou_fg)):
if gt_dict[dt_sorted['image_id']][iou_index[i]]['ignore']==0 and gt_dict[dt_sorted['image_id']][iou_index[i]]['height']>50 and gt_dict[dt_sorted['image_id']][iou_index[i]]['vis_ratio']>0.65:
gt_dict[dt_sorted['image_id']][iou_index[i]]['ignore']=2
return 'tp'
for i in range(len(iou_fg)):
if gt_dict[dt_sorted['image_id']][iou_index[i]]['ignore']==0:
gt_dict[dt_sorted['image_id']][iou_index[i]]['ignore']=2
return 'ignore'
return 'fp'
for i in range(len(dt_sorted)):
matched = match(dt_sorted[i])
if matched == 'tp':
miss_num-=1
true_num+=1
p.append(true_num/(true_num+false_num))
r.append(true_num/total_fg)
elif matched == 'fp':
miss_rate=float(miss_num)/total_fg
fppi=false_num/500.0
total_fppi.append(fppi)
total_miss.append(miss_rate)
false_num+=1
elif matched == 'ignore':
pass
total_mr=0.0
total_num=0
for i in range(len(total_fppi)):
if total_fppi[i]<1 and total_fppi[i]>0.01:
total_mr+=total_miss[i]
total_num+=1
list_mr.append(total_mr/total_num)
total_fppi_list.append(total_fppi)
total_miss_list.append(total_miss)
total_p_list.append(p)
total_r_list.append(r)
ap.append(sum(p)/(len(r)))
for i in range(len(total_fppi_list)):
plt.loglog(total_fppi_list[i], total_miss_list[i], colors[i], label=test_dir[i][:-4]+': ' + str(round(list_mr[i], 3)))
plt.legend(fontsize=5)
plt.figure(2)
plt.title('mAP')
plt.xlabel('recall')
plt.ylabel('precision')
for i in range(len(total_p_list)):
plt.plot(total_r_list[i], total_p_list[i], colors[i], label=test_dir[i][:-4]+': ' + str(round(ap[i], 3)))
plt.legend(fontsize=5)
# -
|
evaluation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Optimizing the number of layers in an LSTM Autoencoder
#
# This is a continuation of the regular autoencoder for rare event classification presented in
# https://towardsdatascience.com/lstm-autoencoder-for-extreme-rare-event-classification-in-keras-ce209a224cfb
# and code present in
# https://github.com/cran2367/lstm_autoencoder_classifier
#
# The dataset used here is taken from here,
#
# **Dataset: Rare Event Classification in Multivariate Time Series** https://arxiv.org/abs/1809.10717 (please cite this article, if using the dataset).
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import seaborn as sns
import pandas as pd
import numpy as np
from pylab import rcParams
import tensorflow as tf
from keras import optimizers, Sequential
from keras.models import Model
from keras.utils import plot_model
from keras.layers import Dense, LSTM, RepeatVector, TimeDistributed, Dropout
from keras.callbacks import ModelCheckpoint, TensorBoard
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import train_test_split
from sklearn.metrics import confusion_matrix, precision_recall_curve
from sklearn.metrics import recall_score, classification_report, auc, roc_curve
from sklearn.metrics import precision_recall_fscore_support, f1_score
from numpy.random import seed
seed(7)
from tensorflow import set_random_seed
set_random_seed(11)
from sklearn.model_selection import train_test_split
SEED = 123 #used to help randomly select the data points
DATA_SPLIT_PCT = 0.2
rcParams['figure.figsize'] = 8, 6
LABELS = ["Normal","Break"]
# -
# ## Reading and preparing data
# The data is taken from https://arxiv.org/abs/1809.10717. Please use this source for any citation.
'''
Download data here:
https://docs.google.com/forms/d/e/1FAIpQLSdyUk3lfDl7I5KYK_pw285LCApc-_RcoC0Tf9cnDnZ_TWzPAw/viewform
'''
df = pd.read_csv("data/processminer-rare-event-mts - data.csv")
df.head(n=5) # visualize the data.
# ### Shift the data
#
# This is a timeseries data in which we have to predict the event (y = 1) ahead in time. In this data, consecutive rows are 2 minutes apart. We will shift the labels in column `y` by 2 rows to do a 4 minute ahead prediction.
# +
sign = lambda x: (1, -1)[x < 0]
def curve_shift(df, shift_by):
'''
This function will shift the binary labels in a dataframe.
The curve shift will be with respect to the 1s.
For example, if shift is -2, the following process
will happen: if row n is labeled as 1, then
- Make row (n+shift_by):(n+shift_by-1) = 1.
- Remove row n.
i.e. the labels will be shifted up to 2 rows up.
Inputs:
df A pandas dataframe with a binary labeled column.
This labeled column should be named as 'y'.
shift_by An integer denoting the number of rows to shift.
Output
df A dataframe with the binary labels shifted by shift.
'''
vector = df['y'].copy()
for s in range(abs(shift_by)):
tmp = vector.shift(sign(shift_by))
tmp = tmp.fillna(0)
vector += tmp
labelcol = 'y'
# Add vector to the df
df.insert(loc=0, column=labelcol+'tmp', value=vector)
# Remove the rows with labelcol == 1.
df = df.drop(df[df[labelcol] == 1].index)
# Drop labelcol and rename the tmp col as labelcol
df = df.drop(labelcol, axis=1)
df = df.rename(columns={labelcol+'tmp': labelcol})
# Make the labelcol binary
df.loc[df[labelcol] > 0, labelcol] = 1
return df
# +
'''
Shift the data by 2 units, equal to 4 minutes.
Test: Testing whether the shift happened correctly.
'''
print('Before shifting') # Positive labeled rows before shifting.
one_indexes = df.index[df['y'] == 1]
display(df.iloc[(one_indexes[0]-3):(one_indexes[0]+2), 0:5].head(n=5))
# Shift the response column y by 2 rows to do a 4-min ahead prediction.
df = curve_shift(df, shift_by = -2)
print('After shifting') # Validating if the shift happened correctly.
display(df.iloc[(one_indexes[0]-4):(one_indexes[0]+1), 0:5].head(n=5))
# -
# If we note here, we moved the positive label at 5/1/99 8:38 to t-1 and t-2 timestamps, and dropped row t. There is a time difference of more than 2 minutes between a break row and the next row because in the data consecutive break rows are deleted. This was done to prevent a classification model learn predicting a break after the break has happened. Refer https://arxiv.org/abs/1809.10717 for details.
# Remove time column, and the categorical columns
df = df.drop(['time', 'x28', 'x61'], axis=1)
# # Prepare data for LSTM models
# LSTM is a bit more demanding than other models. Significant amount of time and attention goes in preparing the data that fits an LSTM.
#
# First, we will create the 3-dimensional arrays of shape: (samples x timesteps x features). Samples mean the number of data points. Timesteps is the number of time steps we look back at any time t to make a prediction. This is also referred to as lookback period. The features is the number of features the data has, in other words, the number of predictors in a multivariate data.
# +
input_X = df.loc[:, df.columns != 'y'].values # converts the df to a numpy array
input_y = df['y'].values
n_features = input_X.shape[1] # number of features
# -
def temporalize(X, y, lookback):
output_X = []
output_y = []
for i in range(len(X)-lookback-1):
t = []
for j in range(1,lookback+1):
# Gather past records upto the lookback period
t.append(X[[(i+j+1)], :])
output_X.append(t)
output_y.append(y[i+lookback+1])
return output_X, output_y
# In LSTM, to make prediction at any time t, we will look at data from (t-lookback):t. In the following, we have an example to show how the input data are transformed with the `temporalize` function with `lookback=5`. For the modeling, we may use a longer lookback.
# +
'''
Test: The 3D tensors (arrays) for LSTM are forming correctly.
'''
print('First instance of y = 1 in the original data')
display(df.iloc[(np.where(np.array(input_y) == 1)[0][0]-5):(np.where(np.array(input_y) == 1)[0][0]+1), ])
lookback = 5 # Equivalent to 10 min of past data.
# Temporalize the data
X, y = temporalize(X = input_X, y = input_y, lookback = lookback)
print('For the same instance of y = 1, we are keeping past 5 samples in the 3D predictor array, X.')
display(pd.DataFrame(np.concatenate(X[np.where(np.array(y) == 1)[0][0]], axis=0 )))
# -
# The two tables are the same. This testifies that we are correctly taking 5 samples (= lookback), X(t):X(t-5) to predict y(t).
# ### Divide the data into train, valid, and test
X_train, X_test, y_train, y_test = train_test_split(np.array(X), np.array(y), test_size=DATA_SPLIT_PCT, random_state=SEED)
X_train, X_valid, y_train, y_valid = train_test_split(X_train, y_train, test_size=DATA_SPLIT_PCT, random_state=SEED)
X_train.shape
# +
X_train_y0 = X_train[y_train==0]
X_train_y1 = X_train[y_train==1]
X_valid_y0 = X_valid[y_valid==0]
X_valid_y1 = X_valid[y_valid==1]
# -
X_train_y0.shape
# #### Reshaping the data
# The tensors we have here are 4-dimensional. We will reshape them into the desired 3-dimensions corresponding to sample x lookback x features.
# +
X_train = X_train.reshape(X_train.shape[0], lookback, n_features)
X_train_y0 = X_train_y0.reshape(X_train_y0.shape[0], lookback, n_features)
X_train_y1 = X_train_y1.reshape(X_train_y1.shape[0], lookback, n_features)
X_test = X_test.reshape(X_test.shape[0], lookback, n_features)
X_valid = X_valid.reshape(X_valid.shape[0], lookback, n_features)
X_valid_y0 = X_valid_y0.reshape(X_valid_y0.shape[0], lookback, n_features)
X_valid_y1 = X_valid_y1.reshape(X_valid_y1.shape[0], lookback, n_features)
# -
# ### Standardize the data
# It is usually better to use a standardized data (transformed to Gaussian, mean 0 and sd 1) for autoencoders.
#
# One common mistake is: we normalize the entire data and then split into train-test. This is not correct. Test data should be completely unseen to anything during the modeling. We should normalize the test data using the feature summary statistics computed from the training data. For normalization, these statistics are the mean and variance for each feature.
#
# The same logic should be used for the validation set. This makes the model more stable for a test data.
#
# To do this, we will require two UDFs.
#
# - `flatten`: This function will re-create the original 2D array from which the 3D arrays were created. This function is the inverse of `temporalize`, meaning `X = flatten(temporalize(X))`.
# - `scale`: This function will scale a 3D array that we created as inputs to the LSTM.
# +
def flatten(X):
'''
Flatten a 3D array.
Input
X A 3D array for lstm, where the array is sample x timesteps x features.
Output
flattened_X A 2D array, sample x features.
'''
flattened_X = np.empty((X.shape[0], X.shape[2])) # sample x features array.
for i in range(X.shape[0]):
flattened_X[i] = X[i, (X.shape[1]-1), :]
return(flattened_X)
def scale(X, scaler):
'''
Scale 3D array.
Inputs
X A 3D array for lstm, where the array is sample x timesteps x features.
scaler A scaler object, e.g., sklearn.preprocessing.StandardScaler, sklearn.preprocessing.normalize
Output
X Scaled 3D array.
'''
for i in range(X.shape[0]):
X[i, :, :] = scaler.transform(X[i, :, :])
return X
# -
# Initialize a scaler using the training data.
scaler = StandardScaler().fit(flatten(X_train_y0))
X_train_y0_scaled = scale(X_train_y0, scaler)
X_train_y1_scaled = scale(X_train_y1, scaler)
X_train_scaled = scale(X_train, scaler)
'''
Test: Check if the scaling is correct.
The test succeeds if all the column means
and variances are 0 and 1, respectively, after
flattening.
'''
a = flatten(X_train_y0_scaled)
print('colwise mean', np.mean(a, axis=0).round(6))
print('colwise variance', np.var(a, axis=0))
# The test succeeded. Now we will _scale_ the validation and test sets.
# +
X_valid_scaled = scale(X_valid, scaler)
X_valid_y0_scaled = scale(X_valid_y0, scaler)
X_test_scaled = scale(X_test, scaler)
# -
# ## LSTM Autoencoder training
# First we will initialize the Autoencoder architecture. We are building a simple autoencoder. More complex architectures and other configurations should be explored.
# +
timesteps = X_train_y0_scaled.shape[1] # equal to the lookback
n_features = X_train_y0_scaled.shape[2] # 59
epochs = 100
batch_size = 64
lr = 0.0001
# +
lstm_autoencoder = Sequential()
# Encoder
lstm_autoencoder.add(LSTM(32, activation='linear', input_shape=(timesteps, n_features), return_sequences=True))
lstm_autoencoder.add(LSTM(4, activation='linear', return_sequences=False, dropout = 0.5))
lstm_autoencoder.add(RepeatVector(timesteps))
# Decoder
lstm_autoencoder.add(LSTM(4, activation='linear', return_sequences=True))
lstm_autoencoder.add(LSTM(32, activation='linear', return_sequences=True))
lstm_autoencoder.add(TimeDistributed(Dense(n_features)))
lstm_autoencoder.summary()
# -
# As a rule-of-thumb, look at the number of parameters. If not using any regularization, keep this less than the number of samples. If using regularization, depending on the degree of regularization you can let more parameters in the model that is greater than the sample size. For example, if using dropout with 0.5, you can have up to double the sample size (loosely speaking).
# +
adam = optimizers.Adam(lr)
lstm_autoencoder.compile(loss='mse', optimizer=adam)
cp = ModelCheckpoint(filepath="lstm_autoencoder_classifier.h5",
save_best_only=True,
verbose=0)
tb = TensorBoard(log_dir='./logs',
histogram_freq=0,
write_graph=True,
write_images=True)
lstm_autoencoder_history = lstm_autoencoder.fit(X_train_y0_scaled, X_train_y0_scaled,
epochs=epochs,
batch_size=batch_size,
validation_data=(X_valid_y0_scaled, X_valid_y0_scaled),
verbose=2).history
# -
plt.plot(lstm_autoencoder_history['loss'], linewidth=2, label='Train')
plt.plot(lstm_autoencoder_history['val_loss'], linewidth=2, label='Valid')
plt.legend(loc='upper right')
plt.title('Model loss')
plt.ylabel('Loss')
plt.xlabel('Epoch')
plt.show()
# ### Sanity check
# Doing a sanity check by validating the reconstruction error
# on the train data. Here we will reconstruct the entire train
# data with both 0 and 1 labels.
#
# **Expectation**: the reconstruction error of 0 labeled data should
# be smaller than 1.
#
# **Caution**: do not use this result for model evaluation. It may
# result into overfitting issues.
# +
train_x_predictions = lstm_autoencoder.predict(X_train_scaled)
mse = np.mean(np.power(flatten(X_train_scaled) - flatten(train_x_predictions), 2), axis=1)
error_df = pd.DataFrame({'Reconstruction_error': mse,
'True_class': y_train.tolist()})
groups = error_df.groupby('True_class')
fig, ax = plt.subplots()
for name, group in groups:
ax.plot(group.index, group.Reconstruction_error, marker='o', ms=3.5, linestyle='',
label= "Break" if name == 1 else "Normal")
ax.legend()
plt.title("Reconstruction error for different classes")
plt.ylabel("Reconstruction error")
plt.xlabel("Data point index")
plt.show();
# -
# ## Predictions using the Autoencoder
# +
valid_x_predictions = lstm_autoencoder.predict(X_valid_scaled)
mse = np.mean(np.power(flatten(X_valid_scaled) - flatten(valid_x_predictions), 2), axis=1)
error_df = pd.DataFrame({'Reconstruction_error': mse,
'True_class': y_valid.tolist()})
precision_rt, recall_rt, threshold_rt = precision_recall_curve(error_df.True_class, error_df.Reconstruction_error)
plt.plot(threshold_rt, precision_rt[1:], label="Precision",linewidth=5)
plt.plot(threshold_rt, recall_rt[1:], label="Recall",linewidth=5)
plt.title('Precision and recall for different threshold values')
plt.xlabel('Threshold')
plt.ylabel('Precision/Recall')
plt.legend()
plt.show()
# +
test_x_predictions = lstm_autoencoder.predict(X_test_scaled)
mse = np.mean(np.power(flatten(X_test_scaled) - flatten(test_x_predictions), 2), axis=1)
error_df = pd.DataFrame({'Reconstruction_error': mse,
'True_class': y_test.tolist()})
threshold_fixed = 0.55
groups = error_df.groupby('True_class')
fig, ax = plt.subplots()
for name, group in groups:
ax.plot(group.index, group.Reconstruction_error, marker='o', ms=3.5, linestyle='',
label= "Break" if name == 1 else "Normal")
ax.hlines(threshold_fixed, ax.get_xlim()[0], ax.get_xlim()[1], colors="r", zorder=100, label='Threshold')
ax.legend()
plt.title("Reconstruction error for different classes")
plt.ylabel("Reconstruction error")
plt.xlabel("Data point index")
plt.show();
# -
pred_y = [1 if e > threshold_fixed else 0 for e in error_df.Reconstruction_error.values]
# +
conf_matrix = confusion_matrix(error_df.True_class, pred_y)
plt.figure(figsize=(6, 6))
sns.heatmap(conf_matrix, xticklabels=LABELS, yticklabels=LABELS, annot=True, fmt="d");
plt.title("Confusion matrix")
plt.ylabel('True class')
plt.xlabel('Predicted class')
plt.show()
# +
false_pos_rate, true_pos_rate, thresholds = roc_curve(error_df.True_class, error_df.Reconstruction_error)
roc_auc = auc(false_pos_rate, true_pos_rate,)
plt.plot(false_pos_rate, true_pos_rate, linewidth=5, label='AUC = %0.3f'% roc_auc)
plt.plot([0,1],[0,1], linewidth=5)
plt.xlim([-0.01, 1])
plt.ylim([0, 1.01])
plt.legend(loc='lower right')
plt.title('Receiver operating characteristic curve (ROC)')
plt.ylabel('True Positive Rate')
plt.xlabel('False Positive Rate')
plt.show()
# -
# ### Obtaining the intermediate layer
intermediate_layer = Model(inputs=lstm_autoencoder.inputs, outputs=lstm_autoencoder.layers[1].output)
# time_dist_layer = Model(inputs=encoder_decoder.inputs, outputs=encoder_decoder.layers[5].output)
intermediate_output = intermediate_layer.predict(X_train_y0_scaled)
intermediate_output.shape
intermediate_output
aa = np.array(intermediate_output)
cc = np.cov(aa.T)
np.linalg.eig(cc)[0]
plt.plot(np.linalg.eig(cc)[0])
# * Run PCA on original data.
# * See how the eigenvalues are distributed.
# * Run PCA on the scores. Expectation: their eig is all equal.
# *
|
lstm-autoencoder-pca.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R [conda env:scanpy_r]
# language: R
# name: conda-env-scanpy_r-r
# ---
library(readr)
library(tximport)
samples <- read.table('../cross-species-data/SRP007412_sample_info.tsv', header = TRUE)
unique(samples$species)
samples
unique(samples$species)
samples <- samples[samples$species=="gallus_gallus",]
species_tx2gene_map = list("bos_taurus"= "/home//cmb-panasas2/skchoudh/genomes/bos_taurus/annotation/Bos_taurus.UMD3.1.94.tx2gene.tsv",
"rattus_norvegicus"="/home/cmb-panasas2/skchoudh/genomes/rattus_norvegicus/annotation/Rattus_norvegicus.Rnor_6.0.94.tx2gene.tsv",
"pongo_abelii"="/home/cmb-panasas2/skchoudh/genomes/pongo_abelii/annotation/Pongo_abelii.PPYG2.94.tx2gene.tsv",
"monodelphis_domestica"="/home/cmb-panasas2/skchoudh/genomes/monodelphis_domestica/annotation/Monodelphis_domestica.monDom5.94.tx2gene.tsv",
"macaca_mulatta"="/home/cmb-panasas2/skchoudh/genomes/macaca_mulatta/annotation/Macaca_mulatta.Mmul_8.0.1.94.tx2gene.tsv",
"pan_troglodytes" ="/home/cmb-panasas2/skchoudh/genomes/pan_troglodytes/annotation/Pan_troglodytes.Pan_tro_3.0.94.tx2gene.tsv",
"mus_musculus"="/home/cmb-panasas2/skchoudh/genomes/mus_musculus/annotation/Mus_musculus.GRCm38.94.tx2gene.tsv",
"homo_sapiens" = "/home/cmb-panasas2/skchoudh/genomes/homo_sapiens/annotation/Homo_sapiens.GRCh38.94.tx2gene.tsv",
"gallus_gallus" = "/home/cmb-panasas2/skchoudh/genomes/gallus_gallus//annotation//Gallus_gallus.Gallus_gallus-5.0.94.tx2gene.tsv",
"ornithorhynchus_anatinus" = "/home/cmb-panasas2/skchoudh/genomes/ornithorhynchus_anatinus/annotation/Ornithorhynchus_anatinus.OANA5.94.tx2gene.tsv",
"gorilla_gorilla" = "/home/cmb-panasas2/skchoudh/genomes/gorilla_gorilla/annotation/Gorilla_gorilla.gorGor4.94.tx2gene.tsv",
"pan_paniscus" = "/home/cmb-panasas2/skchoudh/genomes/pan_paniscus/annotation//Pan_paniscus.panpan1.1.94.tx2gene.tsv"
)
names(species_tx2gene_map)
collapse_to_gene <- function (species, srp, sampleInfo){
files <- file.path('../cross-species-data/kallisto-tables', species, srp, '/counts/', sampleInfo$sample, "abundance.tsv")
names(files) <- sampleInfo$tissue
print(species_tx2gene_map[[species]])
tx2gene <- read.table(species_tx2gene_map[[species]], header=FALSE)
txi.kallisto <- tximport(files, type = "kallisto", ignoreTxVersion = TRUE, tx2gene = tx2gene)
df <- as.data.frame(txi.kallisto$abundance)
#df$genes <- rownames(df)
write.table(df, file=file.path('../cross-species-data/tpm_tables/', paste0(srp, '-', species, '_kallisto_gene_tables.tsv')), quote=F, sep='\t', row.names=T, col.names=T)
}
samples
collapse_to_gene('gallus_gallus', 'SRP016501', samples)
# +
samples <- read.table('../cross-species-data/SRP016501_sample_info.tsv', header = TRUE)
for (species in unique(samples$species)){
collapse_to_gene(species, 'SRP016501', samples[samples$species==species,])
}
# +
samples <- read.table('../cross-species-data/SRP007412_sample_info.tsv', header = TRUE)
for (species in unique(samples$species)){
collapse_to_gene(species, 'SRP007412', samples[samples$species==species,])
}
# +
samples <- read.table('../cross-species-data/SRP136499_sample_info.tsv', header = TRUE, sep='\t')
for (species in unique(samples$species)){
collapse_to_gene(species, 'SRP136499', samples[samples$species==species,])
}
# -
species <- 'pan_troglodytes'
srp <- 'SRP136499'
sampleInfo<- samples[samples$species==species,]
files <- file.path('/staging/as/skchoudh/rna-seq-output/', species, srp, '/counts/', sampleInfo$sample, "abundance.tsv")
names(files) <- sampleInfo$tissue
files
|
notebooks/18.tximport-export-tpm.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <b>Calule a derivada da função dada. Simplifique as respostas.</b>
# <b> 7. $y = \sqrt{2x}$<b>
# <b>Aplicando a regra do radical: $ \sqrt{a} = a^{\frac{1}{2}}$<br><br>
# $ \sqrt{2x} \, =\, ( 2x )^{\frac{1}{2}}$<br><br>
# <b>Aplique a regra da potência: </b> $\frac{d}{dx}(x^a) = a\cdot x^{a-1}$<br><br>
# $\frac{d}{dx}(\sqrt{2x}) = \frac{1}{2} \cdot (2x)^{\frac{1}{2}-1}$<br>
# $\frac{d}{dx}(\sqrt{2x}) = \frac{1}{2} \cdot (2x)^{-\frac{1}{2}}$<br><br>
#
# <b>Aplicando a regra do expoente: $ a^{-b} = \frac{1}{a^b}$<br><br>
#
# $\frac{d}{dx}(\sqrt{2x}) = \frac{1}{2} \cdot \frac{1}{(2x)^{\frac{1}{2}}}$<br><br>
#
# <b>Aplicando a regra do radical: $ \sqrt{a} = a^{\frac{1}{2}}$<br><br>
#
# $\frac{d}{dx}(\sqrt{2x}) = \frac{1}{2} \cdot \frac{1}{\sqrt{2x}}$<br><br>
# $\frac{d}{dx}(\sqrt{2x}) = \frac{1}{2\sqrt{2x}}$<br>
#
|
Problemas 2.2/07.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:py37]
# language: python
# name: conda-env-py37-py
# ---
# ### Ingest and Translate MTRduino Field Data
#
# python: >= 3.7
# author: bell
# Current setup has 5 samples for each 10min measurement.
# Calibration constants are determined prior to (and occasionally post) deployment.
# +
import pandas as pd
import numpy as np
# %matplotlib inline
# -
infile = '/Users/bell/Downloads/newmtrfrom17bs4/5009 MTR.TXT'
# +
#Calibration Coefficients
unit_id_str = '5009'
cal_date_str = '2017-July-13'
AA = 1.19337E-03
BB = 5.29331E-04
CC = 1.30828E-06
# -
df = pd.read_csv(infile,delimiter=',',names=['date_time','sample_1','sample_2','sample_3','sample_4','sample_5','ref_thermistor','unused'],
parse_dates=['date_time'],index_col='date_time')
def steinhardt_hart(sample,AA,BB,CC):
return (1 / (AA + BB * np.log10(sample) + CC * (np.log10(sample))**3) - 273.15)
#apply steinhardt_hart equation to each sample
df['T1'] = df.apply(lambda row: steinhardt_hart(row['sample_1'],AA,BB,CC), axis=1)
df['T2'] = df.apply(lambda row: steinhardt_hart(row['sample_2'],AA,BB,CC), axis=1)
df['T3'] = df.apply(lambda row: steinhardt_hart(row['sample_3'],AA,BB,CC), axis=1)
df['T4'] = df.apply(lambda row: steinhardt_hart(row['sample_4'],AA,BB,CC), axis=1)
df['T5'] = df.apply(lambda row: steinhardt_hart(row['sample_5'],AA,BB,CC), axis=1)
#quicklook of first couple of samples
df.head()
#quicklook of the first of 5 samples, final data will do statistical averageing
df.T1.plot(figsize=(11,4.25))
# calculate mean and std of 5 sample suite
df['ave'] = df[['T1', 'T2', 'T3', 'T4', 'T5']].mean(axis=1)
df['median'] = df[['T1', 'T2', 'T3', 'T4', 'T5']].median(axis=1)
df['std'] = df[['T1', 'T2', 'T3', 'T4', 'T5']].std(axis=1)
df.to_csv(unit_id_str + '.csv', columns=['ave','median','std'])
## resample if necessary, to 10min intervals, interpolating linearly forward but only one step
# doing it on the averaged data is ok especially if the std is low or 0
df['median'].resample('10T').mean().interpolate(limit=1)
|
MTRduino/MTRduino_raw2engr.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Face and Facial Keypoint detection
#
# After you've trained a neural network to detect facial keypoints, you can then apply this network to *any* image that includes faces. The neural network expects a Tensor of a certain size as input and, so, to detect any face, you'll first have to do some pre-processing.
#
# 1. Detect all the faces in an image using a face detector (we'll be using a Haar Cascade detector in this notebook).
# 2. Pre-process those face images so that they are grayscale, and transformed to a Tensor of the input size that your net expects. This step will be similar to the `data_transform` you created and applied in Notebook 2, whose job was tp rescale, normalize, and turn any iimage into a Tensor to be accepted as input to your CNN.
# 3. Use your trained model to detect facial keypoints on the image.
#
# ---
# In the next python cell we load in required libraries for this section of the project.
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
# %matplotlib inline
# #### Select an image
#
# Select an image to perform facial keypoint detection on; you can select any image of faces in the `images/` directory.
# +
import cv2
# load in color image for face detection
image = cv2.imread('images/obamas.jpg')
# switch red and blue color channels
# --> by default OpenCV assumes BLUE comes first, not RED as in many images
image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
# plot the image
fig = plt.figure(figsize=(9,9))
plt.imshow(image)
# -
# ## Detect all faces in an image
#
# Next, you'll use one of OpenCV's pre-trained Haar Cascade classifiers, all of which can be found in the `detector_architectures/` directory, to find any faces in your selected image.
#
# In the code below, we loop over each face in the original image and draw a red square on each face (in a copy of the original image, so as not to modify the original). You can even [add eye detections](https://docs.opencv.org/3.4.1/d7/d8b/tutorial_py_face_detection.html) as an *optional* exercise in using Haar detectors.
#
# An example of face detection on a variety of images is shown below.
#
# <img src='images/haar_cascade_ex.png' width=80% height=80%/>
#
# +
# load in a haar cascade classifier for detecting frontal faces
face_cascade = cv2.CascadeClassifier('detector_architectures/haarcascade_frontalface_default.xml')
# run the detector
# the output here is an array of detections; the corners of each detection box
# if necessary, modify these parameters until you successfully identify every face in a given image
faces = face_cascade.detectMultiScale(image, 1.2, 2)
# make a copy of the original image to plot detections on
image_with_detections = image.copy()
# loop over the detected faces, mark the image where each face is found
for (x,y,w,h) in faces:
# draw a rectangle around each detected face
# you may also need to change the width of the rectangle drawn depending on image resolution
cv2.rectangle(image_with_detections,(x,y),(x+w,y+h),(255,0,0),3)
fig = plt.figure(figsize=(9,9))
plt.imshow(image_with_detections)
# -
# ## Loading in a trained model
#
# Once you have an image to work with (and, again, you can select any image of faces in the `images/` directory), the next step is to pre-process that image and feed it into your CNN facial keypoint detector.
#
# First, load your best model by its filename.
# +
import torch
from models import Net
net = Net()
## TODO: load the best saved model parameters (by your path name)
## You'll need to un-comment the line below and add the correct name for *your* saved model
# net.load_state_dict(torch.load('saved_models/keypoints_model_1.pt'))
net.load_state_dict(torch.load('saved_models/supreeth_model_1.pt'))
## print out your net and prepare it for testing (uncomment the line below)
net.eval()
# -
# ## Keypoint detection
#
# Now, we'll loop over each detected face in an image (again!) only this time, you'll transform those faces in Tensors that your CNN can accept as input images.
#
# ### TODO: Transform each detected face into an input Tensor
#
# You'll need to perform the following steps for each detected face:
# 1. Convert the face from RGB to grayscale
# 2. Normalize the grayscale image so that its color range falls in [0,1] instead of [0,255]
# 3. Rescale the detected face to be the expected square size for your CNN (224x224, suggested)
# 4. Reshape the numpy image into a torch image.
#
# **Hint**: The sizes of faces detected by a Haar detector and the faces your network has been trained on are of different sizes. If you find that your model is generating keypoints that are too small for a given face, try adding some padding to the detected `roi` before giving it as input to your model.
#
# You may find it useful to consult to transformation code in `data_load.py` to help you perform these processing steps.
#
#
# ### TODO: Detect and display the predicted keypoints
#
# After each face has been appropriately converted into an input Tensor for your network to see as input, you can apply your `net` to each face. The ouput should be the predicted the facial keypoints. These keypoints will need to be "un-normalized" for display, and you may find it helpful to write a helper function like `show_keypoints`. You should end up with an image like the following with facial keypoints that closely match the facial features on each individual face:
#
# <img src='images/michelle_detected.png' width=30% height=30%/>
#
#
#
# +
def showpoints(image,keypoints):
plt.figure()
keypoints = keypoints.data.numpy()
keypoints = keypoints * 60.0 + 68
keypoints = np.reshape(keypoints, (68, -1))
plt.imshow(image, cmap='gray')
plt.scatter(keypoints[:, 0], keypoints[:, 1], s=50, marker='.', c='r')
from torch.autograd import Variable
image_copy = np.copy(image)
# loop over the detected faces from your haar cascade
for (x,y,w,h) in faces:
# Select the region of interest that is the face in the image
roi = image_copy[y:y+h,x:x+w]
## TODO: Convert the face region from RGB to grayscale
roi = cv2.cvtColor(roi, cv2.COLOR_RGB2GRAY)
image = roi
## TODO: Normalize the grayscale image so that its color range falls in [0,1] instead of [0,255]
roi = roi/255.0
## TODO: Rescale the detected face to be the expected square size for your CNN (224x224, suggested)
roi = cv2.resize(roi, (224,224))
## TODO: Reshape the numpy image shape (H x W x C) into a torch image shape (C x H x W)
roi = np.expand_dims(roi, 0)
roi = np.expand_dims(roi, 0)
## TODO: Make facial keypoint predictions using your loaded, trained network
roi_torch = Variable(torch.from_numpy(roi))
roi_torch = roi_torch.type(torch.FloatTensor)
keypoints = net(roi_torch)
## TODO: Display each detected face and the corresponding keypoints
showpoints(image,keypoints)
# -
|
3. Facial Keypoint Detection, Complete Pipeline.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Notebook version 1.0, 31 Aug 2021. Written by <NAME> / CSC - IT Center for Science Ltd. <EMAIL>
#
# Licensed under the MIT license: http://www.opensource.org/licenses/mit-license.php
#
# Tested on Kvasi, running QLM version 1.2.1: https://research.csc.fi/-/kvasi
# ***
# # Applying Noise to QAOA
# This notebook explains how to apply noise to QAOA for the MaxCut algorithm. It shows how to extract the circuit from the MaxCut problem class and how to measure the expectation value of an observable on a noisy QPU. We are going to use the Clifford gate set $(H, S, RX, RY, RZ, CNOT)$ with the addition of the $PH$ gate so that no circuit conversion is needed. The phase gate (PH) can be implemented in practice using the other Clifford $+ T$ gates but that is omitted here.
## defining the networkx Graph
import networkx as nx
nodes = 6
graph = nx.generators.random_graphs.erdos_renyi_graph(n=nodes, p=0.5)
nx.draw(graph,with_labels=True) # vizualization of the randomly generated graph
# +
## defining the Max-Cut problem
from qat.vsolve.qaoa import MaxCut
problem = MaxCut(graph) # Instatiate MaxCut problem, with the above graph as an input
H = problem.get_observable() # Get the problem Hamiltonian
p = 2 # Depth of QAOA
ansatz = problem.qaoa_ansatz(depth = p) # creates ansatz with one pair of 'QAOA blocks' applied
# extract a corresponding circuit object from the ansatz
circuit = ansatz.circuit
# %qatdisplay circuit
## In case the circuit is too big to draw, we can use qat.core.util.statistics to
# see how the circuit is built.
from qat.core.util import statistics
print(statistics(circuit))
# -
# ### Noisy Simulation of the Algorithm
# Now that we know what gates our QAOA circuit uses, we can define noise models for this circuit the same way as with any other circuit. We need to define noisy Hadamards (H), phase gates (PH), X-rotations (RX), and CNOT gates. In addition, we define amplitude damping and pure dephasing noise to model environmental noise. This noise affects qubits during the whole duration of the algorithm. Learn more here: https://arxiv.org/abs/1904.06560
# +
## Specifications for gate fidelities and gate times. Change these
# values to observe how the they affect the results of the algorithm
# X rotation gate
Xrot_fidelity = 0.995 # probability of success
Xrot_time = 30 # gate duration in nanoseconds
# H gate
H_fidelity = 0.99
H_time = 30
# CNOT gate
CNOT_fidelity = 0.98
CNOT_time = 280
# PH gate
PH_fidelity = 0.99
PH_time = 30
## Environmetal noise parameters
T1 = 120000 # qubit's energy relaxation time in nanoseconds
T2 = 190000 # qubit's dephasing time in nanoseconds
# -
# Here we define quantum channels for the gates we use in the circuit. In order to do this, one needs to know how to represent gates in their matrix form. These can be found under "The AQASM format" section in the QLM documentation, for example.
# +
from qat.quops import QuantumChannelKraus ## Kraus representation of quantum channels
import numpy as np # numpy is used to generate matrices
# Noisy X-rotation
px = Xrot_fidelity # probability that the RX(theta) rotation succeeds
noisy_RX = lambda theta : QuantumChannelKraus([np.sqrt(px)*np.array([[np.cos(theta/2), -np.sin(theta/2)*1j],
[-np.sin(theta/2)*1j, np.cos(theta/2)]]), # the RX(pi/2) gate -> gate succeeds
np.sqrt(1-px)*np.identity(2)], # the identity -> nothing happens
name="noisy RX(pi/2)") # name of the quantum operation/channel
# Noisy Hadamard
ph = H_fidelity
noisy_H = QuantumChannelKraus([np.sqrt(ph)*np.array([[1, 1],
[1, -1]]/np.sqrt(2)),
np.sqrt(1-ph)*np.identity(2)],
name="noisy H")
# Noisy CNOT
pcnot = CNOT_fidelity # probability that the CNOT rotation succeeds
noisy_CNOT = QuantumChannelKraus([np.sqrt(pcnot)*np.array([[1, 0, 0, 0],
[0, 1, 0, 0],
[0, 0, 0, 1],
[0, 0, 1, 0]]), # the CNOT gate -> gate succeeds
np.sqrt(1-pcnot)*np.identity(4)],# the identity -> nothing happens
name="noisy CNOT")
# Noisy PH
pph = PH_fidelity
noisy_PH = lambda theta : QuantumChannelKraus([np.sqrt(pph)*np.array([[1,0],
[0, np.exp(theta*1j)]]),
np.sqrt(1-pph)*np.identity(2)], name="noisy PH")
# +
from qat.quops import ParametricPureDephasing, ParametricAmplitudeDamping
## Amplitude Damping characterized by T_1
amp_damp = ParametricAmplitudeDamping(T_1 = T1)
## Pure Dephasing characterized by T_phi. The contribution of amplitude damping (T1) is removed
## from transverse relaxation (T2) to give pure dephasing only (T_phi)
pure_deph = ParametricPureDephasing(T_phi = 1/(1/T2 - 1/(2*T1)))
from itertools import product
## Amplitude damping channel for two qubits. We build a new 4x4 quantun channel whose Kraus operators
## are pairwise Kronecker products of the Kraus operators of single qubit amplitude damping channels.
## The gate duration of CNOT is given as parameter 'tau' to amp_damp.
two_qbit_amp_damp = QuantumChannelKraus([np.kron(k1, k2)
for k1, k2 in product(amp_damp(tau=CNOT_time).kraus_operators,
amp_damp(tau=CNOT_time).kraus_operators)])
## Pure dephasing channel for two qubits. The logic is exactly same as above.
two_qbit_pure_deph = QuantumChannelKraus([np.kron(k1, k2)
for k1, k2 in product(pure_deph(tau=CNOT_time).kraus_operators,
pure_deph(tau=CNOT_time).kraus_operators)])
# +
## Let's define environmental noise during gates.
## The series concatenation of quantum channels is implemented in pyAQASM by the '*' operator.
## We set amplitude damping and pure dephasing channels to follow all gate operations
## (this is approximating environmental noise during these gates).
gates_noise = {"RX" : lambda _: amp_damp(tau=Xrot_time)*pure_deph(tau=Xrot_time),
"H" : lambda : amp_damp(tau=H_time)*pure_deph(tau=H_time),
"CNOT" : lambda : two_qbit_amp_damp*two_qbit_pure_deph,
"PH" : lambda _: amp_damp(tau=PH_time)*pure_deph(tau=PH_time)}
# This dictionary connects the noisy quantum channels to gate names
quantum_channels = {"RX": noisy_RX,"H": noisy_H, "CNOT": noisy_CNOT, 'PH': noisy_PH}
## Dictionary connecting the gate durations to the gate's name
gate_times = {"H": H_time,"RX": Xrot_time, "CNOT": CNOT_time, 'PH': PH_time}
from qat.hardware import GatesSpecification
## instanciate a GatesSpecification with gate times and the corresponding quantum channels
gates_spec = GatesSpecification(gate_times, quantum_channels)
from qat.hardware.default import HardwareModel
hw_model = HardwareModel(gates_spec, gate_noise=gates_noise, idle_noise=[amp_damp, pure_deph])
## Display the circuit as a timeline of quantum operations where colored boxes represent noisy quantum gates
## and white boxes noisy idling periods.
# %qatdisplay circuit --hardware hw_model
# -
# ### Optimizing Circuit parameters
# +
from qat.plugins import ScipyMinimizePlugin # import the plugin for optimization
## Instantiate the classical optimizer using Constrained Optimization BY Linear Approximation (COBYLA) method
optimizer = ScipyMinimizePlugin(method="COBYLA", # the optimization method
tol=1e-2, # tolerance is a threshold which, if crossed,
# stops the iterations of an optimizer
options={"maxiter":150}) # maximum optimization iterations.
# Needs to be large enough for result to converge
# (see below)
# -
# Noisy simulations require another plugin called ObservableSplitter to be added to the stack before the optimization step. This is because calculating the expectation value of a given observable is not implemented for NoisyQProc which we use for noisy simulations.
# +
from qat.plugins import ObservableSplitter
from qat.qpus import NoisyQProc
## Now we can stack the classical optimizer, ObservableSplitter and NoisyQProc
stack = optimizer | ObservableSplitter() | NoisyQProc(hw_model)
## now the job is set to calculate the expectation value for H.
job = circuit.to_job('OBS', observable=H, nbshots=0)
noisy_result = stack.submit(job)
print('Optimized noisy value: ', noisy_result.value)
# -
# Let's perform the same measurement on an ideal circuit to confirm that our noise definitions had an effect on the energy measurement of the observable. The energy reading given by the ideal circuit should be lower than that of the noisy circuit.
# +
from qat.qpus import LinAlg
qpu = LinAlg()
ideal_stack = optimizer | qpu ## NOTE: ObservableSplitter is not needed here.
job = circuit.to_job('OBS', observable=H, nbshots=0)
result = ideal_stack.submit(job)
print('Optimized ideal value', result.value)
# -
# ## Retrieving the maximum cut
# The goal in MaxCut is to partition the vertices (modelled as qubits) to two subgraphs such that the disagreement between neighbors is maximized which is equivalent to maximizing the number of cut edges. By measuring $|\gamma_{optimal},\beta_{optimal}>$ in the computational basis, we can divide the qubits into the two different subgraphs. This is because qubits belonging to different subgraphs will have different measurement results.
# +
## Fetch the parameters
params = eval(noisy_result.meta_data['parameters']) # the optimal parameters are in the metadata of the result
gammas = params[0:p] # gammas are the first p values of the parameter array, where p is the depth of the ansatz
betas = params[p:] # betas are the last p values of the parameter array, where p is the depth of the ansatz
# Bind the parameters into the QAOA circuit
circuit_optimal = circuit.bind_variables({key : var for key, var in zip(circuit.get_variables(),params)})
# %qatdisplay circuit_optimal
job2 = circuit_optimal.to_job(nbshots=0)
result2 = NoisyQProc(hw_model).submit(job2)
print("Samples from measurements of the state with optimal parameters:")
for sample in result2:
if sample.probability > 0.01: # only consider samples with reasonable probability
print(sample.state.value[0],", probability:", sample.probability)
## To plot the optimal graph partition, let's find the optimal solution that occurs with highest probability
solutions = {} # initialize a Python dictionary
## loop over samples to collect the measurement outcomes and their probabilities to the dictionary
for sample in result2:
solutions.update({sample.state.value[0] : sample.probability})
most_probable_solution = max(solutions, key=solutions.get) # find the key i.e. state that has maximum probability
print("-------------------------------")
print("The most probable solutions is")
print(most_probable_solution)
# -
# ### All states and their probabilities
# +
import matplotlib.pyplot as plt
## collect result states and their probabilities in two lists
states = [str(sample.state.value[0]) for sample in result2 if sample.probability >= 0.000]
probabilities = [sample.probability for sample in result2 if sample.probability > 0.000]
plt.figure(figsize=(12, 6))
plt.rcParams['font.size'] = 12
plt.bar(states, probabilities)
plt.xlabel('state')
plt.ylabel('probability')
plt.xticks(rotation=90)
plt.show()
# -
# ### Ideal Reference:
# +
## Compare noisy results to the ideal results
ideal_result = qpu.submit(job2)
states = [str(sample.state.value[0]) for sample in ideal_result if sample.probability >= 0.000]
probabilities = [sample.probability for sample in ideal_result if sample.probability > 0.000]
plt.figure(figsize=(12, 6))
plt.rcParams['font.size'] = 12
plt.bar(states, probabilities)
plt.xlabel('state')
plt.ylabel('probability')
plt.xticks(rotation=90)
plt.show()
# -
# ### Visualize the graph partition
## We need to extract the indices of the 0 and 1 partitions
print("Solution configuration: \n" + str(most_probable_solution) + "\n")
indices_0 = [i for i, value in enumerate(most_probable_solution) if value == '0'] # build array holding the indices of 0's in the solution confiquration
print("The nodes in the first subgraph:\n" + str(indices_0) + "\n")
indices_1 = [i for i, value in enumerate(most_probable_solution) if value == '1'] # build array holding the indices of 1's in the solution confiquration
print("The nodes in the second subgraph:\n" + str(indices_1) )
# +
## Here we visualize the solution of the MaxCut problem.
import matplotlib.pyplot as plt
nodes_positions = nx.spring_layout(graph)
plt.figure(figsize=(12,8))
node_size = 440
font_size = 14
nx.draw_networkx(graph, # the nodes of '0'-partition will be colored blue
pos=nodes_positions,
nodelist=indices_0,
node_color='steelblue',
node_size=node_size,
font_size=font_size)
nx.draw_networkx(graph, # the nodes of '1'-partition will be colored red
pos=nodes_positions,
nodelist=indices_1,
node_color='red',
node_size=node_size,
font_size=font_size)
plt.show()
# -
|
Noise-modelling/Tutorial - Noise simulation for QAOA.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7
# language: python
# name: python3
# ---
# +
# The code was removed by Watson Studio for sharing.
# -
# 
#
# ## Setup your notebook file stystem on Watson Studio
#
# All of the notebooks in these courses are written to run locally on your computer running a Jupyter notebook server. If you wish to run the notebooks in Watson Studio in the IBM Cloud, you will need to add some modifications to each notebook.
#
# Why? Because once you import a course notebook and the data files for that notebook into a Watson Studio project, the data files are no longer available to the notebook!
# This is simply due to the fact that the imported data files are stored in an IBM Cloud Object Storage (COS) bucket. The notebook does not have access to those objects in the COS bucket. Thus, if you import a notebook and its data files into a Studio project then try to run it, the notebook will return "File not found" errors.
# In order to make the data files available to your notebook, you will need to run some code in your notebook to:
#
# 1 - Access the correct COS bucket <br>
# 2 - Read your data file from the bucket into a byte stream object <br>
# 3 - Write that byte stream object to the virtual disk of the container running the notebook.
# +
# cos2file - takes an object from Cloud Object Storage and writes it to file on container file system.
# Uses the IBM project_lib library.
# See https://dataplatform.cloud.ibm.com/docs/content/wsj/analyze-data/project-lib-python.html
# Arguments:
# p: project object defined in project token
# data_path: the directory to write the file
# filename: name of the file in COS
import os
def cos2file(p,data_path,filename):
data_dir = p.project_context.home + data_path
if not os.path.exists(data_dir):
os.makedirs(data_dir)
open( data_dir + '/' + filename, 'wb').write(p.get_file(filename).read())
# -
# Calling cos2file allows to make the data files available to the notebook
cos2file(project, '/data', 'aavail-target.csv')
# 
#
# # CASE STUDY - Unsupervised Learning
#
# %%capture
# ! pip install -U scikit-learn
# ! pip install -U imblearn
# +
import os
import time
import numpy as np
import pandas as pd
import scipy.stats as stats
import matplotlib.pyplot as plt
from sklearn.utils import shuffle
from sklearn.model_selection import train_test_split, GridSearchCV
from sklearn.compose import ColumnTransformer
from sklearn.base import BaseEstimator, TransformerMixin
from sklearn.impute import SimpleImputer
from sklearn.cluster import KMeans, SpectralClustering
from sklearn.preprocessing import StandardScaler, OneHotEncoder, OrdinalEncoder
from sklearn.pipeline import Pipeline
from sklearn.metrics import classification_report, f1_score
from sklearn.metrics import silhouette_score
from sklearn.ensemble import RandomForestClassifier
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.mixture import BayesianGaussianMixture
from sklearn.svm import SVC
import imblearn.pipeline as pl
from imblearn.pipeline import make_pipeline
from imblearn.over_sampling import RandomOverSampler
from imblearn.over_sampling import SMOTE, SVMSMOTE
plt.style.use('seaborn')
# %matplotlib inline
DATA_DIR = os.path.join("..","data")
# -
# ## Synopsis
#
# > We are now going to predict customer retention. There are many models and many transforms to consider. Use your
# knowledge of pipelines and functions to ensure that your code makes it easy to compare and iterate over.
#
# > Marketing has asked you to make a report on customer retention. They would like you to come up with information that can be used to improve current marketing strategy efforts. The current plan is for marketing at AAVAiL to
# collect more features on subscribers the and they would like to use your report as a proof-of-concept in order to get buyin for this effort.
#
# ## Outline
#
# 1. Create a churn prediction baseline model
# 2. Use clustering as part of your prediction pipeline
# 3. Run and experiment to see if re-sampling techniques improve your model
#
# ## Data
#
# Here we load the data as we have already done.
#
# `aavail-target.csv`
# +
df = pd.read_csv(os.path.join(DATA_DIR, r"aavail-target.csv"))
## pull out the target and remove uneeded columns
_y = df.pop('is_subscriber')
y = np.zeros(_y.size)
y[_y==0] = 1
df.drop(columns=['customer_id', 'customer_name'], inplace=True)
df.head()
# -
# ### QUESTION 1
#
# Using the train_test_split() function, create a stratified train test split of the data
## YOUR CODE HERE
X_train,X_test,y_train,y_test = train_test_split(df,y,test_size = 0.25,stratify = y,random_state = 42)
# ### QUESTION 2
#
# Create a baseline model. We are going to test whether clustering followed by a model improves the results. Then, we will test whether re-sampling techniques provide improvements. Use a pipeline or another method, but create a baseline model given the data. Here is the ColumnTransformer we have used before:
# +
## preprocessing pipeline
numeric_features = ['age', 'num_streams']
numeric_transformer = Pipeline(steps=[
('imputer', SimpleImputer(strategy='mean')),
('scaler', StandardScaler())])
categorical_features = ['country', 'subscriber_type']
categorical_transformer = Pipeline(steps=[
('imputer', SimpleImputer(strategy='constant', fill_value='missing')),
('encod', OrdinalEncoder())])
preprocessor = ColumnTransformer(
transformers=[
('num', numeric_transformer, numeric_features),
('cat', categorical_transformer, categorical_features)])
# +
# YOUR CODE HERE (Replace the #<> symbols with your code)
# Create an instance of a binary classifier.
clf = RandomForestClassifier()
# Create a pipeline that binds the preprocessing transformer and the classifier estimator.
pipe = Pipeline(steps=[('pre', preprocessor),
('rf', clf)])
# Here we apply a grid search to optimize the hyperparamters of the classifier.
param_grid = {
'rf__n_estimators': [20, 50, 100, 150],
'rf__max_depth': [4, 5, 6, 7, 8],
'rf__criterion': ['gini', 'entropy']
}
grid = GridSearchCV(pipe, param_grid=param_grid, cv=3, n_jobs=-1, scoring='f1')
# Fit the pipeline to the training data.
grid.fit(X_train, y_train)
best_params = grid.best_params_
# Predict the dependent variable of the test set.
y_pred = grid.predict(X_test)
# Print the f1_score of the prediction.
print("f1_score", round(f1_score(y_test, y_pred, average='binary'), 3))
# -
# ### QUESTION 3
#
# The next part is to create a version of the classifier that uses identified clusters. Here is a class to get you started. It is a transformer like those that we have been working with. There is an example of how to use it just below. In this example 4 clusters were specified and their one-hot encoded versions were appended to the feature matrix. Now using pipelines and/or functions compare the performance using cluster profiling as part of your matrix to the baseline. You may compare multiple models and multiple clustering algorithms here.
# +
class KmeansTransformer(BaseEstimator, TransformerMixin):
def __init__(self, n_clusters=4):
self.n_clusters = n_clusters
self.km = KMeans(n_clusters=self.n_clusters, n_init=20)
def transform(self, X, *_):
labels = self.km.predict(X)
return np.hstack((X, labels.reshape(-1, 1)))
def fit(self, X, y=None, *_):
self.km.fit(X)
labels = self.km.predict(X)
self.silhouette_score = round(silhouette_score(X, labels, metric='mahalanobis'), 3)
return self
class GmmTransformer(BaseEstimator, TransformerMixin):
def __init__(self, n_clusters=4):
self.n_clusters = n_clusters
self.gmm = BayesianGaussianMixture(n_components=self.n_clusters, covariance_type='full',
max_iter=500, n_init=10, warm_start=True)
def transform(self, X,*_):
probs = self.gmm.predict_proba(X) + np.finfo(float).eps
return np.hstack((X, probs))
def fit(self, X, y=None, *_):
self.gmm.fit(X)
labels = self.gmm.predict(X)
self.silhouette_score = round(silhouette_score(X, labels, metric='mahalanobis'), 3)
return self
## example for kmeans
preprocessor.fit(X_train)
X_train_pre = preprocessor.transform(X_train)
kt = KmeansTransformer(4)
kt.fit(X_train_pre)
X_train_kmeans = kt.transform(X_train_pre)
print(X_train_pre.shape)
print(X_train_kmeans.shape)
## example for GMM
preprocessor.fit(X_train)
X_train_pre = preprocessor.transform(X_train)
gt = GmmTransformer(4)
gt.fit(X_train_pre)
X_train_gmm = gt.transform(X_train_pre)
print(X_train_pre.shape)
print(X_train_gmm.shape)
# +
## YOUR CODE HERE (Replace the #<> symbols by your code)
def run_clustering_pipeline(umodel):
"""
This function evaluates different Pipelines comprised of the preprocessing transfomer,
a clustering transformer and a classifier estimator.
INPUT : The name of the clustering transformer : 'gmm' or 'kmeans'
OUTPUT : The list of f1_scores of the pipeline on the test set for the different number of clusters
"""
fscores= [] # this list will store the f1_score of the different models that we will train
for n_clusters in np.arange(3, 8):
# Create an instance of a binary classifier (The same as the one you trained in the previous question)
estimator = RandomForestClassifier()
param_grid = {
'n_estimators': [20, 50, 100, 150],
'max_depth': [4, 5, 6, 7, 8],
'criterion': ['gini', 'entropy']
}
clf = GridSearchCV(estimator, param_grid=param_grid, cv=3, n_jobs=-1, scoring='f1')
if umodel == 'gmm':
# Create an instance of the Gmm transformer with n_clusters clusters
cluster = GmmTransformer(n_clusters)
elif umodel == 'kmeans':
# Create an instance of the Kmean transformer with n_clusters clusters
cluster = KmeansTransformer(n_clusters)
else:
raise Exception("invalid unsupervised learning model")
# Create a Pipeline that binds the preprocessing transformer, the clustering transformer and the classifier estimator
pipe = Pipeline(steps=[('pre', preprocessor),
('clustering', cluster),
('classifier', clf)])
# Fit the pipeline on training set
pipe.fit(X_train, y_train)
# Predict the test set
y_pred = pipe.predict(X_test)
# Compute the f1 score and add this score to the fscores list.
score = round(f1_score(y_test, y_pred, average='binary'), 3)
fscores.append(score)
return fscores
## run the different iteration of the model
cp_results = {}
cp_results['kmeans'] = run_clustering_pipeline('kmeans')
cp_results['gmm'] = run_clustering_pipeline('gmm')
## display table of results
df_cp = pd.DataFrame(cp_results)
df_cp["n_clusters"] = [str(i) for i in np.arange(3,8)]
df_cp.set_index("n_clusters", inplace=True)
df_cp.head(n=10)
# -
# ## QUESTION 4
#
# Run an experiment to see if you can you improve on your workflow with the addition of re-sampling techniques? For instance, you can copy the structure of the function created in the previous question and add a re-sampling transformer to the pipeline.
# +
## YOUR CODE HERE
# This cell might take several minutes to run
def run_clustering_pipeline(umodel):
"""
This function evaluates different Pipelines constituated of the preprocessing transfomer,
a clustering transformer, a re-sampling transformer and a classifier estimator.
INPUT : The name of the clustering transformer : 'gmm' or 'kmeans'
OUTPUT : The list of f1_scores of the pipeline on the test set for the different number of clusters.
"""
fscores = [] # this list will store the f1_score of the different models that we will train
for n_clusters in np.arange(3,8):
# Create an instance of a binary classifier (The same as the one you trained in the previous question)
estimator = RandomForestClassifier()
param_grid = {
'n_estimators': [20, 50, 100, 150],
'max_depth': [4, 5, 6, 7, 8],
'criterion': ['gini', 'entropy']
}
clf = GridSearchCV(estimator, param_grid=param_grid, cv=3, n_jobs=-1, scoring='f1')
if umodel == 'gmm':
# Create an instance of the Gmm transformer with n_clusters clusters
cluster = GmmTransformer(n_clusters)
elif umodel == 'kmeans':
# Create an instance of the Kmean transformer with n_clusters clusters
cluster = KmeansTransformer(n_clusters)
else:
raise Exception("invalid unsupervised learning model")
# Create a Pipeline that binds the preprocessing transformer, the clustering transformer,
# the re-sampling transformer and the classifier
pipe = pl.Pipeline(steps=[('pre', preprocessor),
('clustering', cluster),
('smote', SMOTE(random_state=42)),
('classifier', clf)])
# Fit the pipeline on training set
pipe.fit(X_train,y_train)
# Predict the test set
y_pred = pipe.predict(X_test)
# Compute the f1 score and add this score to the fscores list.
score = round(f1_score(y_test, y_pred,average='binary'),3)
fscores.append(score)
return(fscores)
## Run the different iteration of the model
cp_results = {}
cp_results['kmeans'] = run_clustering_pipeline('kmeans')
cp_results['gmm'] = run_clustering_pipeline('gmm')
## Display table of results
df_cp = pd.DataFrame(cp_results)
df_cp["n_clusters"] = [str(i) for i in np.arange(3,8)]
df_cp.set_index("n_clusters",inplace=True)
df_cp.head(n=10)
# -
# Usando Smote y atributos generados mediante Clustering, se logró mejorar los resultados.
|
Aprendizaje No Supervisado/Clustering_2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# [](https://colab.research.google.com/github/comp-hci-lab/BDSI_2021_ML/blob/master/A1%20-%20Decision%20Boundary.ipynb)
# ## BDSI ML 2021
# # A1 - Neural Nets: Decision Boundary (demo)
# - Two synthetic datasets in 2D
# - Visualize the decision boundary and output contour of single-layer or multi-layer neural nets
#
# Similar visualizations can be found at: https://playground.tensorflow.org/
# + colab={} colab_type="code" id="2M0PqNXXZW2v"
# GPU support
import torch
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
print('using device:', device)
# -
import numpy as np
from matplotlib import pyplot as plt
from tqdm import tqdm
import torch
import torch.nn as nn
plt.rcParams['figure.figsize'] = (8.0, 8.0)
plt.rcParams['lines.markersize'] = 6.0
# +
#@title Helper functions for plotting. { display-mode: "form" }
from matplotlib import cm, ticker
MARKERS = ['o', 'x', 'v']
COLORS = ['red', 'green', 'blue']
def count_parameters(model):
return sum(p.numel() for p in model.parameters() if p.requires_grad)
def plot_points(X, y):
for i, label in enumerate(set(y)):
points = X[y == label]
marker = MARKERS[i % len(MARKERS)]
color = COLORS[i % len(COLORS)]
plt.scatter(points[:,0], points[:,1], marker=marker, color=color)
def predict(model, x):
with torch.no_grad():
o = model(torch.from_numpy(x).float().to(device))
return o.cpu().numpy()
def plot_boundary(X, pred):
try:
x_min, x_max = plt.gca().get_xlim()
y_min, y_max = plt.gca().get_ylim()
except:
x_min, x_max = X[:,0].min() - .1, X[:,0].max() + .1
y_min, y_max = X[:,1].min() - .1, X[:,1].max() + .1
xs, ys = np.meshgrid(
np.linspace(x_min, x_max, 200),
np.linspace(y_min, y_max, 200)
)
xys = np.column_stack([xs.ravel(), ys.ravel()])
zs = pred(xys).reshape(xs.shape)
plt.contour(xs, ys, (zs >= 0.5).astype(int), cmap='Greys')
plt.imshow(zs, cmap="PiYG", vmin=-.2, vmax=1.2, alpha=0.4, origin='lower', extent=[x_min, x_max, y_min, y_max])
# +
#@title Two synthetic datasets in 2D. { display-mode: "form" }
from torch.utils.data import Dataset, DataLoader
class D1(Dataset):
def __init__(self, N=50):
super().__init__()
np.random.seed(0)
self.X = np.r_[2.0 * np.random.randn(N//2, 2) - [2, 2], 2.0 * np.random.randn(N//2, 2) + [2, 2]]
self.y = np.array([0] * (N//2) + [1] * (N//2))
def __getitem__(self, idx):
return torch.from_numpy(self.X[idx]).float(), torch.FloatTensor([self.y[idx]])
def __len__(self):
return len(self.X)
class D2(Dataset):
def __init__(self, N=50):
super().__init__()
np.random.seed(0)
X = 2.0 * np.random.randn(N, 2)
y = np.array([0] * N)
y[np.diag(X@X.T)>6] = 1
self.X = X
self.y = y
def __getitem__(self, idx):
return torch.from_numpy(self.X[idx]).float(), torch.FloatTensor([self.y[idx]])
def __len__(self):
return len(self.X)
# -
# ## Load data
# Try `d = D1(100)` or `d = D2(100)`
d = D1(100)
plot_points(d.X, d.y)
# ## Define the neural network:
torch.random.manual_seed(2)
net = nn.Sequential(
nn.Linear(2,1),
nn.Sigmoid(),
)
net.to(device)
criterion = nn.BCELoss()
optimizer = torch.optim.SGD(net.parameters(), lr=0.01)
dataloader = DataLoader(d, batch_size=4)
print('Number of float-valued parameters:', count_parameters(net))
# ## Before training...
plot_points(d.X, d.y)
plot_boundary(d.X, lambda x: predict(net, x))
# ## After training...
# Trainer loop
for epoch in tqdm(range(1000)):
for X, y in dataloader:
X, y = X.to(device), y.to(device)
optimizer.zero_grad()
output = net(X)
loss = criterion(output, y)
loss.backward()
optimizer.step()
plot_points(d.X, d.y)
plot_boundary(d.X, lambda x: predict(net, x))
|
A1 - Decision Boundary.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# 
# # Running LISFLOOD model using eWaterCycle package (on SURF Research Cloud)
#
# This notebooks shows how to run [LISFLOOD](https://ec-jrc.github.io/lisflood-model/) model. Please note that the [lisflood-grpc4bmi docker image](https://hub.docker.com/r/ewatercycle/lisflood-grpc4bmi) in eWaterCycle is compatible only with forcing data and parameter set on [eWaterCycle infrastructure](https://github.com/eWaterCycle/infra) like a server on the SURF Research Cloud. More information about data, configuration and installation instructions can be found in the [System setup](https://ewatercycle.readthedocs.io/en/latest/system_setup.html) in the eWaterCycle documentation.
# +
import logging
import warnings
logger = logging.getLogger("grpc4bmi")
logger.setLevel(logging.WARNING)
warnings.filterwarnings("ignore", category=UserWarning)
# +
import pandas as pd
import ewatercycle.forcing
import ewatercycle.models
import ewatercycle.parameter_sets
# -
# ## Load forcing data
#
# For this example notebook, `lisflood_ERA-Interim_*_1990_1990.nc` data are copied
# from `/projects/0/wtrcycle/comparison/forcing/lisflood` to `/scratch/shared/ewatercycle/lisflood_example/lisflood_forcing_data`.
# Also the lisvap output files 'e0', 'es0' and 'et0' are generated and stored in the same directory.
# These data are made by running ESMValTool recipe and lisvap. We can now use those files to run the Lisflood model.
forcing = ewatercycle.forcing.load_foreign(
target_model="lisflood",
directory="/mnt/data/forcing/lisflood_ERA5_1990_global-masked",
start_time="1990-01-01T00:00:00Z",
end_time="1990-12-31T00:00:00Z",
forcing_info={
"PrefixPrecipitation": "lisflood_ERA5_pr_1990_1990.nc",
"PrefixTavg": "lisflood_ERA5_tas_1990_1990.nc",
"PrefixE0": "lisflood_ERA5_e0_1990_1990.nc",
"PrefixES0": "lisflood_ERA5_es0_1990_1990.nc",
"PrefixET0": "lisflood_ERA5_et0_1990_1990.nc",
},
)
print(forcing)
# + [markdown] tags=[]
# ## Load parameter set
# + [markdown] tags=[]
# This example uses parameter set on Cartesius machine of SURFsara.
# + tags=[]
parameterset = ewatercycle.parameter_sets.ParameterSet(
name="Lisflood01degree_masked",
directory="/mnt/data/parameter-sets/lisflood_global-masked_01degree",
config="/mnt/data/parameter-sets/lisflood_global-masked_01degree/settings_lisflood_ERA5.xml",
target_model="lisflood",
)
print(parameterset)
# + [markdown] tags=[]
# ## Set up the model
# + [markdown] tags=[]
# To create the model object, we need to select a version.
# -
ewatercycle.models.Lisflood.available_versions
model = ewatercycle.models.Lisflood(
version="20.10", parameter_set=parameterset, forcing=forcing
)
print(model)
model.parameters
# Setup model with model_mask, IrrigationEfficiency of 0.8 instead of 0.75 and an earlier end time, making total model time just 1 month.
# +
model_mask = "/mnt/data/climate-data/aux/LISFLOOD/model_mask.nc"
config_file, config_dir = model.setup(
IrrigationEfficiency="0.8", end_time="1990-1-31T00:00:00Z", MaskMap=model_mask
)
print(config_file)
print(config_dir)
# -
model.parameters
# Initialize the model with the config file:
model.initialize(config_file)
# Get model variable names
model.output_var_names
# ## Run the model
# Store simulated values at one target location until model end time. In this example, we use the coordinates of Merrimack observation station as the target coordinates.
# +
target_longitude = [-71.35]
target_latitude = [42.64]
target_discharge = []
time_range = []
end_time = model.end_time
while model.time < end_time:
model.update()
target_discharge.append(
model.get_value_at_coords(
"Discharge", lon=target_longitude, lat=target_latitude
)[0]
)
time_range.append(model.time_as_datetime.date())
print(model.time_as_isostr)
# -
# Store simulated values for all locations of the model grid at end time.
discharge = model.get_value_as_xarray("Discharge")
model.finalize()
# ## Inspect the results
#
# The discharge time series at Merrimack observation station:
simulated_target_discharge = pd.DataFrame(
{"simulation": target_discharge}, index=pd.to_datetime(time_range)
)
simulated_target_discharge.plot(figsize=(12, 8))
# The lisflood output has a global extent. In this example, we plot the discharge values in Merrimack catchment and at the last time step.
lc = discharge.coords["longitude"]
la = discharge.coords["latitude"]
discharge_map = discharge.loc[
dict(longitude=lc[(lc > -73) & (lc < -70)], latitude=la[(la > 42) & (la < 45)])
].plot(robust=True, cmap="GnBu", figsize=(12, 8))
discharge_map.axes.scatter(
target_longitude, target_latitude, s=250, c="r", marker="x", lw=2
)
|
docs/examples/lisflood.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import os
raw_data_path = os.path.join(os.path.pardir,'data','raw')
train_file_path = os.path.join(raw_data_path,'train.csv')
test_file_path = os.path.join(raw_data_path,'test.csv')
print(train_file_path)
train_df = pd.read_csv(train_file_path,index_col="PassengerId")
test_df = pd.read_csv(test_file_path,index_col="PassengerId")
type(train_df)
# # Basic structure
train_df.info()
test_df.info()
test_df["Survived"] = -888
df = pd.concat((train_df,test_df),axis=0)
df.info()
df.head()
df.head(10)
df.tail()
df.tail(10)
df.Name
df['Name']
df[['Name','Age']]
df.loc[5:10]
df.loc[5:10,"Age":"Pclass"]
df.loc[5:10, ["Age","Fare","Embarked"]]
df.iloc[5:10, 3:8]
male_passengers = df.loc[df["Sex"] == 'male',:]
print("'Number of Male passenger {0}".format(len(male_passengers)))
male_passengers_first_class = df.loc[((df["Sex"] == 'male') & (df["Pclass"] == 1)),:]
male_passengers_first_class = df.loc[((df.Sex == 'male') & (df.Pclass == 1)),:]
print("Male Passengers in first class {0}".format(len(male_passengers_first_class)))
# # Summary Statistics
df.describe()
print("Minimum fare {0}".format(df.Fare.min()))
print("max fare {0}".format(df.Fare.max()))
print("median fare {0}".format(df.Fare.median()))
print("median fare {0}".format(df.Fare.median()))
print("25 percentile {0}".format(df.Fare.quantile(.25)))
print("50 percentile {0}".format(df.Fare.quantile(.5)))
print("75 percentile {0}".format(df.Fare.quantile(.75)))
print("Variance {0}".format(df.Fare.var()))
print("Std deviation fare {0}".format(df.Fare.std()))
df.Fare.plot(kind='box',title='Whisker plot')
# %matplotlib inline
# ## Categorical Statistics
df.describe(include='all')
df.Sex.value_counts()
df.Sex.value_counts(normalize=True)
df[df.Survived != -888].Survived.value_counts()
df[df.Survived != -888].Survived.value_counts(normalize=True)
df.Pclass.value_counts()
df.Pclass.value_counts().plot(kind='bar')
df.Pclass.value_counts().plot(kind='bar',rot=0,title='Class wise passenger count');
# ## Distribution
#
df.Age.plot(kind='hist',title='Histogram for Age')
df.Age.plot(kind='hist',title='Histogram for Age',bins=20);
df.Age.plot(kind='kde',title='Density plot for age')
df.Fare.plot(kind='hist',title='Fare distribution')
print('skewness for age :{0:2f}'.format(df.Age.skew()))
print('skewness for age :{0:2f}'.format(df.Fare.skew()))
# # Scatter plot
df.plot.scatter(x="Age",y="Fare",title="Age vs Fare")
df.plot.scatter(x="Age",y="Fare",alpha=0.1)
df.plot.scatter(x="Pclass",y="Fare",title="Class vs Fare",alpha=0.15)
# # Grouping data
df.groupby("Sex").Age.median()
df.groupby("Sex").Age.mean()
df.groupby("Pclass").Age.median()
df.groupby("Pclass")["Fare","Age"].median()
df.groupby("Pclass")["Age","Fare"].median()
df.groupby("Pclass").agg({'Fare':'mean','Age':'median'})
aggregations = {
'Fare':{
'mean_fare':'mean',
'median_fare':'median',
'max_fare':'max',
'min_fare':np.min,
},
'Age':{
'mean_age':'mean',
'median_age':'median',
'max_age':'max',
'min_age':'min',
'range_age':lambda x: max(x) - min(x)
}
}
df.groupby('Pclass').agg(aggregations)
df.groupby(['Pclass','Embarked']).Fare.median()
# # Crosstabs
pd.crosstab(df.Pclass,df.Sex)
pd.crosstab(df.Sex,df.Pclass)
pd.crosstab(df.Sex,df.Pclass).plot(kind='bar')
pd.crosstab(df.Sex,df.Pclass).plot(kind='hist')
# # Pivot Table
df.pivot_table(index="Sex",columns="Pclass",values="Age",aggfunc="mean")
df.groupby(['Sex','Pclass']).Age.mean()
df.groupby(['Sex','Pclass']).Age.mean().unstack()
df.groupby(['Sex','Pclass']).Age.mean().plot(kind='box')
# # Data munging
#
# ### missing values treatment
# Data issues
# - Missing values - one more attributes corresponding to observ
# - Extreme values (outlier)
# - Erroneous values
#
# +
### Missing values - Values not known for features or observations
# +
## Solutions Deletion, Imputation
# +
## Mean imputation - fill missing values from mean value (not reliable should avoid)
# -
## Median imputation - fill missing values from median value
# +
## Mode imputation - categoric feature ( replace the highest value)
# +
##Technqiues - Forward/Backward Fill, previous or next values will be replaces
# +
# Predictive model can be used
# -
df.info()
# ## Feature : Embarked
df[df.Embarked.isnull()]
df.Embarked.value_counts()
pd.crosstab(df[df.Survived != -888].Survived,df[df.Survived != -888].Embarked)
df.groupby(['Pclass','Embarked']).Fare.median()
df.Embarked.fillna('C',inplace=True)
df[df.Embarked.isnull()]
df.info()
df[df.Fare.isnull()]
median_fare = df.loc[(df.Embarked == 'S') & (df.Pclass==3),'Fare'].median()
print(median_fare)
df.Fare.fillna(median_fare,inplace=True)
df[df.Fare.isnull()]
df.info()
df[df.Age.isnull()]
pd.options.display.max_rows = 15
df[df.Age.isnull()]
df.Age.plot(kind='hist',bins=20,title='Age distribution')
df.Age.mean()
df.Age.median()
df.groupby('Sex').Age.median()
df[df.Age.notnull()].boxplot('Age','Sex')
# +
# Replace values
# sex_age_median = df.groupby('Sex').Age.transform('median')
# df.Age.fillna(sex_age_median,inplace=True)
# -
df[df.Age.notnull()].boxplot("Age","Pclass")
# +
# we could replace by grouping pclass
# pclass_median_age = df.groupby("PClass").Age.transform('median)
# df.Age.fillna(pclass_median_age,inplace=True)
# -
df.Name
def GetTitle(name):
first_name_with_title = name.split(',')[1]
title = first_name_with_title.split('.')[0]
title = title.strip().lower()
return title
df.Name.map(lambda x: GetTitle(x))
df.Name.map(lambda x: GetTitle(x)).unique()
def GetTitle(name):
title_groups = {
"mr":"Mr",
"mrs":"Mrs",
"miss":"Miss",
"master":"Master",
"don":"Sir",
"rev":"Sir",
"dr":"Officer",
"mme":"Mrs",
"ms":"Mrs",
"major":"Officer",
"lady":"Lady",
"sir":"Sir",
"mlle":"Miss",
"col":"Officer",
"capt":"Officer",
"the countess":"Lady",
"jonkheer":"Sir",
"dona":"Lady"
}
first_name_with_title = name.split(',')[1]
title = first_name_with_title.split('.')[0]
title = title.strip().lower()
return title_groups[title]
df.Name.map(lambda x: GetTitle(x)).unique()
df['Title'] = df.Name.map(lambda x: GetTitle(x))
df.head()
df[df.Age.notnull()].boxplot("Age","Title")
title_median_age = df.groupby('Title').Age.transform('median')
df.Age.fillna(title_median_age,inplace=True)
df.head()
df.info()
# # Working with outliers
#
# ### Different from normal
#
# #### Multiple source
#
# - Data entry
# - Data processing
# - Natural
#
# #### Issues
#
# - Biased analysis
# - Biased models
#
#
#
# ### Outlier Detection
#
# - Using Histgram
# - Boxplot
# - Scatter plot
#
# ##### Outlier Treatments
#
# - Removal -
# - Transformation - Log and sqrt
# - Binning - seperate bin
# - Imputation - replace with reasonable values
#
df.Age.plot(kind='hist',bins=20)
df.loc[df.Age > 70]
# ## Fare
#
df.Fare.plot(kind='hist',bins=20)
#Transformation using log function
df.Fare.plot(kind='box')
# Look into the outliers
df.loc[df.Fare == df.Fare.max()]
np.log(2)
np.exp(0.6931471805599453)
LogFare = np.log(df.Fare + 1.0)
## Transformation for extreme values
LogFare.plot(kind='hist',bins=20)
# +
#Binnig
# -
pd.qcut(df.Fare,4)
pd.qcut(df.Fare,4,labels=["verylow","low","high","veryhigh"])
df["Fare_Bin"] = pd.qcut(df.Fare,4,labels=["verylow","low","high","veryhigh"])
df.Fare_Bin.value_counts().plot(kind='bar')
# # Feature Engineering
#
# process of transforming raw data to better representative features in order to create better predictive models
#
# - Transformation
# - Creation (using domain expertise -)
# - Selection
#
# Domain knowlege and technical expertise
# +
# Feature creation using exisitng data
# -
df['AgeState'] = np.where(df.Age >= 18,'Adult','Child')
df.AgeState.value_counts()
pd.crosstab(df.loc[(df.Survived != -888)].Survived,df.loc[(df.Survived != -888)].AgeState)
# ### Feature: FamilySize
df['FamilySize'] = df.Parch + df.SibSp
df['FamilySize'].plot(kind='hist')
pd.crosstab(df.loc[df.Survived != -888].Survived,df.loc[df.Survived != -888].FamilySize)
df.loc[df.FamilySize == df.FamilySize.max(),['Name','Survived','FamilySize','Ticket']]
# ## Feature: IsMother
df['IsMother'] = np.where((df.Sex == 'female' ) & (df.Parch > 0) & (df.Age > 18) & (df.Title != 'Miss'),1,0)
df.IsMother.value_counts()
pd.crosstab(df.loc[df.Survived != -888].Survived,df.loc[df.Survived != -888].IsMother)
# ## Feature: Deck
#
# Kevin attribute
df.Cabin
df.Cabin.unique()
df.loc[df.Cabin=='T']
df.loc[df.Cabin == 'T','Cabin'] = np.NaN
df.Cabin.unique()
def get_deck(cabin):
return np.where(pd.notnull(cabin),str(cabin)[0],'Z')
df['Deck'] = df['Cabin'].map(lambda x : get_deck(x))
df.Deck.value_counts()
pd.crosstab(df[df.Survived != -888].Survived,df[df.Survived != -888].Deck)
df.info()
# ### Categorical feature
#
# converting categorical feature to numerical feature
#
# - Binary encoding ( change to 1 or 0 when 2 categories possible)
# - Label encoding (when more than 2 you can change encode value to 1,2,3..) when is ordered value (low,high,very high)
# - One-Hot Encoding - each category become a new feature which holds 1 or 0 if you have more categories more features to be genereated - this is popular way of encoding in ML
#
#
#
# +
## Categorical Feature encoding
# -
df['IsMale'] = np.where(df.Sex == 'Female',0,1)
df = pd.get_dummies(df,columns=['Deck','Title','Fare_Bin','Embarked','AgeState'])
print(df.info())
# Drop columns
df.drop(labels=['Cabin','Name','Parch','Sex','Ticket','SibSp'],axis=1,inplace=True)
coloums = [column for column in df.columns if column != "Survived" ]
coloums = ['Survived'] + coloums
df = df[coloums]
df.info()
# ## Save Processed dataset
processed_data_path = os.path.join(os.pardir,'data','processed')
processed_train_path = os.path.join(processed_data_path,'train.csv')
processed_test_path = os.path.join(processed_data_path,'test.csv')
#train data
df.loc[df.Survived != -888].to_csv(processed_train_path)
#test data
columns = [column for column in df.columns if column != "Survived"]
df.loc[df.Survived == -888,coloums].to_csv(processed_test_path)
# ## Building the data processing script
get_processed_data_script_file = os.path.join(os.pardir,'src','data','get_processed_data.py' )
# +
# %%writefile $get_processed_data_script_file
import numpy as np
import pandas as pd
import os
def read_data():
raw_data_path = os.path.join(os.pardir,'data','raw')
train_file_path = os.path.join(raw_data_path,'train.csv')
test_file_path = os.path.join(raw_data_path,'test.csv')
# read the data with all default parameters
train_df = pd.read_csv(train_file_path,index_col="PassengerId")
test_df = pd.read_csv(test_file_path,index_col="PassengerId")
test_df['Survived'] = -888
df = pd.concat((train_df,test_df),axis=0)
return df
def process_data(df):
return(df
## create title attribute
.assign(Title = lambda x: x.Name.map(get_title))
#working with missing values
.pipe(fill_missing_values)
## create fare bin attribute
.assign(Fare_Bin = lambda x:
pd.qcut(x.Fare,4,labels=['very_low','low','high','very_high']))
##create age state
.assign(AgeState = lambda x: np.where(x.Age >18,'Adult','Child'))
## Assign family size
.assign(FamilySize = lambda x:x.Parch + x.SibSp +1)
.assign(IsMother= lambda x: np.where(((x.Sex == 'female')
& (x.Age > 18) &
(x.Parch > 0) &
(x.Title != 'Miss')),1,0))
.assign(Cabin = lambda x:np.where(x.Cabin =='T',np.NaN,x.Cabin))
.assign(Deck= lambda x: x.Cabin.map(get_deck))
.assign(IsMale = lambda x: np.where(x.Sex == 'Male',1,0))
.pipe(pd.get_dummies,
columns=['Deck','Pclass','Title','Fare_Bin','Embarked','AgeState'])
.drop(['Cabin','Name','Ticket','Parch','SibSp','Sex'],axis=1)
.pipe(reorder_columns)
)
def reorder_columns(df):
coloums = [column for column in df.columns if column != "Survived" ]
coloums = ['Survived'] + coloums
return df[coloums]
def fill_missing_values(df):
#Embarked
df.Embarked.fillna('C',inplace=True)
#Fare
median_fare = df[(df.Pclass==3) & (df.Embarked=='S')]['Fare'].median()
df.Fare.fillna(median_fare,inplace=True)
#age
title_age_median = df.groupby('Title').Age.transform('median')
df.Age.fillna(title_age_median,inplace=True)
return df
def get_deck(cabin):
return np.where(pd.notnull(cabin),str(cabin)[0],'Z')
def write_data(df):
processed_data_path = os.path.join(os.pardir,'data','processed')
write_train_path = os.path.join(processed_data_path,'train.csv')
write_test_path = os.path.join(processed_data_path,'test.csv')
#train data
df[df.Survived != -888].to_csv(write_train_path)
#test data
columns = [column for column in df.columns if column != "Survived"]
df[df.Survived == -888][columns].to_csv(write_test_path)
def get_title(name):
title_groups = {
"mr":"Mr",
"mrs":"Mrs",
"miss":"Miss",
"master":"Master",
"don":"Sir",
"rev":"Sir",
"dr":"Officer",
"mme":"Mrs",
"ms":"Mrs",
"major":"Officer",
"lady":"Lady",
"sir":"Sir",
"mlle":"Miss",
"col":"Officer",
"capt":"Officer",
"the countess":"Lady",
"jonkheer":"Sir",
"dona":"Lady"
}
first_name_with_title = name.split(',')[1]
title = first_name_with_title.split('.')[0]
title = title.strip().lower()
return title_groups[title]
if __name__ == '__main__':
df = read_data()
df = process_data(df)
write_data(df)
# -
# !python $get_processed_data_script_file
# # Advanced visualisation using matplotlib
import matplotlib.pyplot as plt
# %matplotlib inline
plt.hist(df.Age)
plt.hist(df.Age,bins=20)
plt.hist(df.Age,bins=20)
plt.show()
plt.hist(df.Age,bins=20)
plt.title(str('Hist: Age'))
plt.xlabel('Age')
plt.ylabel('Counts')
plt.show()
f, ax = plt.subplots()
ax.hist(df.Age)
ax.set_title("Hist: Age")
ax.set_xlabel("Age")
ax.set_ylabel("Counts")
plt.show()
f, ax = plt.subplots()
ax.boxplot(df.Age)
ax.set_title("Hist: Age")
ax.set_xlabel("Age")
ax.set_ylabel("Counts")
plt.show()
# +
f,ax = plt.subplots(2,2)
ax[0,0].hist(df.Age)
ax[0,1].boxplot(df.Age)
ax[1,0].hist(df.Fare)
ax[1,1].axis('off')
plt.tight_layout()
# -
# # Building and Evaluvating predictive models #1
# ## Data science project cycle
#
# - Machine learning concepts
# ****Learn from data or examples
# - supervised -input and output features (train,test)
# - unsupervised - Clustering - customer segmentation
#
# Classification - Discrete values - (Binary classication,Multiclass classification)
# Regresssion - Continuous values
# **** Classifier - more than 3 features (seperating hyperplane)
# Logistic Regression
# SVM
# NN
# RF
# Performace metrics
# - Accuarcy = correct_count/total_count
# - Precision = confusion_matrix TP/(TP+FP)
# What fraction of positive predictions are correct?
# - Recall = TP/(TP + FN)
# What fraction of positive cases you predicted correctly?
# - Classifier evaluation - Train and Test Split
#
# - Basline model
# - Output majory class
# - Predictive model should have better performance than baseline
#
# - Predictive model
# - Fine Tune
# - Model persistence
#
# ==> analyse + Model ==> Presentation
#
#
|
notebooks/Exploring and Processing data - part 1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### PYT-DS SAISOFT
#
# * [Overview 1](https://github.com/4dsolutions/Python5/blob/master/OverviewNotes_PYTDS.ipynb)
# * [Overview 2](https://github.com/4dsolutions/Python5/blob/master/OverviewNotes_PYTDS_2.ipynb)
#
# <a data-flickr-embed="true" href="https://www.flickr.com/photos/kirbyurner/27963484878/in/album-72157693427665102/" title="Barry at Large"><img src="https://farm1.staticflickr.com/969/27963484878_b38f0db42a_m.jpg" width="240" height="180" alt="Barry at Large"></a><script async src="//embedr.flickr.com/assets/client-code.js" charset="utf-8"></script>
#
# # DATA SCIENCE WITH PYTHON
#
#
# [Click here](http://nbviewer.jupyter.org/github/4dsolutions/Python5/blob/master/OverviewNotes_PYTDS_3.ipynb) to view this JN on nbviewer.
#
# In the concluding sessions of this course, I have shifted from talking about the data pipeline, to the functions at the end of the tunnel, our Machine Learning algorithms, which I've also likened to a stable of horses, in terms of how we "race" them to find the best. Choosing the best horse for your application takes experience. Don't expect to become a data scientist overnight.
#
# In our sequence below, I start with a famous, oft used dataset, made of 28 by 28 numpy arrays, representing grayscale images of the numerals 0 through 9, quite a few specimens of each. They're labeled rows. We know the digits. Lets take a look.
import numpy as np
from sklearn.datasets import load_digits
digits = load_digits()
print(digits.data.shape)
print(digits.DESCR)
import matplotlib.pyplot as plt
% matplotlib inline
plt.gray() # gray reversed shown below
_ = plt.matshow(digits.images[0])
# [Another way](http://scikit-learn.org/stable/auto_examples/datasets/plot_digits_last_image.html).
plt.figure(1, figsize=(3, 3))
plt.imshow(digits.images[0], cmap=plt.cm.gray_r, interpolation='nearest')
plt.show()
_ = plt.matshow(digits.images[108])
digits.data[108]
# Remember how we think in machine learning. We have a multifaceted (multi-featured) set of samples, rows with many columns, and then a single column of correct results, an "answer key" if you will.
#
# We often call this answer key column the "target" and then measure "error" as divergence between guesses and target.
#
# Decreasing divergence bespeaks of a learning rate as the model trains on, or fits the training data. Whether we control this learning rate as a hyperparameter, or leave it to the algorithm to work at some built-in speed, depends on which machine learner type we've selected. Below we're looking at [KNN](http://scikit-learn.org/stable/modules/generated/sklearn.neighbors.KNeighborsClassifier.html) and then a neural net.
digits.target[108]
# That's a very poor rendering of the numeral 7 and we're immediately forgiving if our Machine Learning algorithm gets some wrong, with training data of such abysmal quality. As seen from ```digits.data```, the 64 bits used to represent a digit are hardly enough. Other datasets come with at least 28 x 28 bits for each numeral. We're truly at the low end with this skimpy number of bits per digit.
#
# Neverthesless, we press on... I'm making only minor changes to [this open source script on Github](https://gist.github.com/fabiosato/2a2014cd386d475fd037), by [Fabiosato](https://gist.github.com/fabiosato).
#
# Remember how KNN works:
from IPython.display import YouTubeVideo
YouTubeVideo("MDniRwXizWo")
# Remember to distinguish KNN from K-Means. You might use the latter to create the clusters whereby you could then fit the former. Here's [a paper on LinkedIn](https://www.linkedin.com/pulse/classification-clustering-knn-vs-k-means-raymond-rashid/) suggesting doing that. Once you have the clusters (voters), a new data point is "claimed" by one or more clusters.
#
# [Hierarchical clustering algorithms](http:github/4dsolutions/Python5/blob/master/OverviewNotes_PYTDS_3.ipynbPYT_DS_Clustering.ipynb) compete with K-Means. The latter does better for spherical or globular clusters.
from IPython.display import YouTubeVideo
YouTubeVideo("3vHqmPF4VBA")
# +
from sklearn.metrics import confusion_matrix, accuracy_score
from sklearn.model_selection import train_test_split
from sklearn import neighbors # http://scikit-learn.org/stable/modules/classes.html#module-sklearn.neighbors
# prepare datasets from training and for validation
X_train, X_test, y_train, y_test = train_test_split(digits.data, digits.target,
test_size=0.4, random_state=0)
# runs the kNN classifier for even number of neighbors from 1 to 10
for n in range(1, 10, 2):
clf = neighbors.KNeighborsClassifier(n)
# instance based learning
clf.fit(X_train, y_train)
# our 'ground truth'
y_true = y_test
# predict
y_pred = clf.predict(X_test)
# learning metrics
cm = confusion_matrix(y_true, y_pred)
acc = accuracy_score(y_true, y_pred)
print ("Neighbors: %d" % n)
print ("Confusion Matrix")
print (cm)
print ("Accuracy score: %f" % accuracy_score(y_true, y_pred))
print ()
# -
# Discerning digits within a blizzard of data points streaming in, or other patterns, may be described as a process of identifying clusters or [neighborhoods](http://scikit-learn.org/stable/modules/neighbors.html). Even before we name the clusters we claim to find, we need to find them, and this is where dimensionality reduction comes in handy, as if we can get the dimensions down to three, we have some axes we might use.
#
# "Dimensionality reduction" involves finding eigenvectors, the most efficient at singling out cells in not containing redundant info, forming a basis. An idea of ranking eigenvectors, in the sense of "most significant digits", allows us to cluster data by just the first few eigenvector coordinates.
#
# One might usefully compare this process to discovering the desmomap, or binary tree resulting from bottom-up progressive agglomeration into larger groups. One may then place a threshold cut through the data to vary the number of clusters one wishes to regard as separate. There's a sense of binning and/or pigeon-holing, where the hyperparameter is the degree of subdivisioning.
#
# Does a [neural network](http://scikit-learn.org/stable/modules/generated/sklearn.neural_network.MLPClassifier.html) fare better? Let's admit, the KNN machine learner did a great job. Fast horse!
# +
from sklearn.neural_network import MLPClassifier
# runs the MLP classifier, all with same hyperparameters
for n in range(1, 10, 2):
clf = MLPClassifier()
# instance based learning
clf.fit(X_train, y_train)
# our 'ground truth'
y_true = y_test
# predict
y_pred = clf.predict(X_test)
# learning metrics
cm = confusion_matrix(y_true, y_pred)
acc = accuracy_score(y_true, y_pred)
# print ("Neighbors: %d" % n)
print ("Confusion Matrix")
print (cm)
print ("Accuracy score: %f" % accuracy_score(y_true, y_pred))
print ()
# -
# I'd say these two are competitive, but award KNN first prize in this case. On the other hand, I did not try varying the hyperparameters available to me with [the MLP classifier](http://scikit-learn.org/stable/modules/neural_networks_supervised.html). Lets say the results so far are inconclusive. More research needed.
|
OverviewNotes_PYTDS_3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_mxnet_p36
# language: python
# name: conda_mxnet_p36
# ---
# # SageMaker built-in BlazingText(Supervised) example
# 1. [Introduction](#Introduction)
# 2. [Development Environment and Data Preparation](#Development-Environment-and-Data-Preparation)
# 1. [Setup](#Setup)
# 2. [Data Preparation](#Data-Preparation)
# 3. [Preprocessing](#Preprocessing)
# 3. [Starting Sagemaker Training Job](#Starting-Sagemaker-Training-Job)
# 1. [Create BlazingText Container](#Create-BlazingText-Container)
# 2. [Creating an Estimator and start a training job](#Creating-an-Estimator-and-start-a-training-job)
# 3. [Training](#Training)
# 4. [Inference](#Inference)
# 1. [(Optional) Batch Inference](#(Optional)-Batch-Inference)
# 1. [Hosting / Inference](#Hosting-/-Inference)
# 5. [Stop / Close the Endpoint](#Stop-/-Close-the-Endpoint)
# ## Introduction
#
# ビルトインアルゴリズムの一つであるBlazingTextでは、教師なしアルゴリズムであるWord2Vecと教師ありの分類アルゴリズムを提供しています。
#
# ビルトインアルゴリズムを使用する場合、学習とデプロイに関連するコードのほとんどを開発者が意識する必要がなくなる点も利点となります。
#
# このノートブックでは、Amazon の商品レビューに対する感情分析、つまり、そのレビューが Positive (Rating が 5 or 4) か、Negative (Rating が 1 or 2)なのかを判定します。これは、文書を Positive か Negative に分類する2クラスの分類問題なので、**BlazingText**による教師あり学習を適用することができます。
# ## Data Preparation
#
# Amazon の商品レビューデータセットは[Registry of Open Data on AWS](https://registry.opendata.aws/)で公開されており、 以下からダウンロード可能です。
# このノートブックでは、日本語のデータセットをダウンロードします。
#
# - データセットの概要
# https://registry.opendata.aws/amazon-reviews/
#
# - 日本語のデータセット(readme.htmlからたどることができます)
# https://s3.amazonaws.com/amazon-reviews-pds/tsv/amazon_reviews_multilingual_JP_v1_00.tsv.gz
#
# 以下では、データをダウンロードして解凍 (unzip) します。
# +
import urllib.request
import os
import gzip
import shutil
download_url = "https://s3.amazonaws.com/amazon-reviews-pds/tsv/amazon_reviews_multilingual_JP_v1_00.tsv.gz"
dir_name = "data"
file_name = "amazon_review.tsv.gz"
tsv_file_name = "amazon_review.tsv"
file_path = os.path.join(dir_name,file_name)
tsv_file_path = os.path.join(dir_name,tsv_file_name)
os.makedirs(dir_name, exist_ok=True)
if os.path.exists(file_path):
print("File {} already exists. Skipped download.".format(file_name))
else:
urllib.request.urlretrieve(download_url, file_path)
print("File downloaded: {}".format(file_path))
if os.path.exists(tsv_file_path):
print("File {} already exists. Skipped unzip.".format(tsv_file_name))
else:
with gzip.open(file_path, mode='rb') as fin:
with open(tsv_file_path, 'wb') as fout:
shutil.copyfileobj(fin, fout)
print("File uznipped: {}".format(tsv_file_path))
# -
# ## Preprocessing
#
# ダウンロードしたデータには学習に不要なデータや直接利用できないデータもあります。以下の前処理で利用できるようにします。
#
# 1. ダウンロードしたデータには不要なデータも含まれているので削除します。
# 2. 2クラス分類 (positive が 1, negative が 0)となるように評価データを加工し、レビューデータをMeCabを使ってスペース区切りのデータにします。
# 3. 学習データ、バリデーションデータ、テストデータに分けて、学習用にS3にデータをアップロードします。
#
# ### データの確認
#
# タブ区切りの tsv ファイルを読んで1行目を表示してみます。
# +
import pandas as pd
df = pd.read_csv(tsv_file_path, sep ='\t')
df.head()
# -
# ### 不要なデータの削除
#
# 今回利用しないデータは以下の2つです。必要なデータだけ選んで保存します。
#
# - 評価データ `star_rating` と レビューのテキストデータ `review_body` 以外のデータ
# - 評価が 3 のデータ (positive でも negative でもないデータ)
df_pos_neg = df.loc[:, ["star_rating", "review_body"]]
df_pos_neg = df_pos_neg[df_pos_neg.star_rating != 3]
df_pos_neg.head()
# ### MeCab のインストール
#
# BlazingText は、文章をそのまま学習・推論に利用することはできず、語ごとにスペースで区切って利用する必要があります。これはスペースで区切られている英語などでは問題ありませんが、スペースで区切られていない日本語では追加の処理が必要になります。
#
# ここでは、形態素とよばれる語の単位に分解(分かち書き)する形態素解析ツール MeCab を利用します。MeCab は pip でインストールして利用することができます。冒頭に`!`を入れることで、シェルコマンドを実行できます。`import MeCab` としても問題ないか確認しましょう。
# +
import sys
# !{sys.executable} -m pip install --upgrade pip
# !{sys.executable} -m pip install mecab-python3
# !{sys.executable} -m pip install unidic-lite
import MeCab
# -
# ### データの加工
#
# BlazingText では以下のようなデータが必要です。
#
# ```
# __label__1 私 は これ が 好き です 。
# __label__0 私 は これ が きらい です 。
# ```
#
# `__label__数字` は文書のラベルを表します。negative `__label__0`、positive なら `__label__1` とします。ラベル以降は、文書をスペース区切りにしたものですので、各文に対して MeCab による形態素解析を実行します。全文の処理に2, 3分必要になる場合があります。
# +
mecab = MeCab.Tagger("-Owakati")
def func_to_row(x):
if x["star_rating"] < 3:
label = '0'
else:
label = '1'
x["star_rating"] = "__label__" + label
x["review_body"] = mecab.parse(x["review_body"].replace('<br />', '')).replace('\n', '')
return x
labeled_df = df_pos_neg.apply(lambda x: func_to_row(x), axis =1)
# -
# ### データの分割
#
# すべてのデータを学習データとすると、データを使って作成したモデルが良いのか悪いのか評価するデータが別途必要になります。
# そこで、データを学習データ、バリデーションデータ、テストデータに分割して利用します。学習データはモデルの学習に利用し、バリデーションデータは学習時のモデルの評価に利用します。最終的に作成されたモデルに対してテストデータによる評価を行います。
#
# `train_ratio` で設定した割合のデータを学習データとし、残ったデータをバリデーションとデータテストデータに分割して利用します。学習に利用する学習データとバリデーションデータは、後にSageMakerで利用するために、`savetxt` を利用してスペース区切りの csv に保存します。
# +
import numpy as np
data_size = len(labeled_df.index)
train_ratio = 0.8
train_index = np.random.choice(data_size, int(data_size*train_ratio), replace=False)
other_index = np.setdiff1d(np.arange(data_size), train_index)
valid_index = np.random.choice(other_index, int(len(other_index)/2), replace=False)
test_index = np.setdiff1d(np.arange(data_size), np.concatenate([train_index, valid_index]))
np.savetxt('train.csv',labeled_df.iloc[train_index].values, fmt="%s %s", delimiter=' ')
np.savetxt('validation.csv',labeled_df.iloc[valid_index].values, fmt="%s %s", delimiter=' ')
print("Data is splitted into:")
print("Training data: {} records.".format(len(train_index)))
print("Validation data: {} records.".format(len(valid_index)))
print("Test data: {} records.".format(len(test_index)))
# -
# ### Uploading data to `sagemaker_session_bucket`
#
# SageMaker での学習に利用するために、学習データとバリデーションデータを S3 にアップロードします。SageMaker Python SDK の upload_data を利用すると、S3 にファイルをアップロードできます。アップロード先のバケットは `sagemaker-{リージョン名}-{アカウントID}`で、バケットがない場合は自動作成されます。もし存在するバケットにアップロードする場合は、このバケット名を引数で指定できます。
#
# アップロードが終われば、TrainingInput を利用して、アップロードしたファイルの content_type などを指定します。
# +
import sagemaker
from sagemaker.inputs import TrainingInput
sess = sagemaker.Session()
s3_train_data = sess.upload_data(path='train.csv', key_prefix='amazon-review-data')
s3_validation_data = sess.upload_data(path='validation.csv', key_prefix='amazon-review-data')
print("Training data is uploaded to {}".format(s3_train_data))
print("Validation data is uploaded to {}".format(s3_validation_data))
train_data = TrainingInput(s3_train_data, distribution='FullyReplicated', content_type='text/plain', s3_data_type='S3Prefix')
validation_data = TrainingInput(s3_validation_data, distribution='FullyReplicated', content_type='text/plain', s3_data_type='S3Prefix')
data_channels = {'train': train_data, 'validation': validation_data}
# -
# ## Starting Sagemaker Training Job
#
# BlazingText はビルトインアルゴリズムなので、アルゴリズムの実装は不要です。BlazingTextのコンテナイメージを呼び出して実行します。`get_image_uri` を利用すればコンテナイメージの URI を取得することができます。 取得した URI とそれを実行するインスタンスなどを指定して、Estimator を呼び出すことで学習の設定を行うことができます。
#
# ビルトインアルゴリズムでは、実行内容を設定するいくつかのハイパーパラメータを設定する必要があります。BlazingText では `mode` のハイパーパラメータが必須です。テキスト分類を行う場合は `mode="supervised"` の指定が必要です。
#
# BlazingTextは、FastTextテキスト分類器を拡張し、カスタムCUDAカーネルを使用してGPUアクセラレーションを活用しています。このモデルは、マルチコアCPUやGPUを使って数分で10億語以上の単語を学習することができ、最先端の深層学習テキスト分類アルゴリズムと同等のパフォーマンスを実現しています。詳細については、[algorithm documentation](https://docs.aws.amazon.com/sagemaker/latest/dg/blazingtext.html)または[the text classification notebook](https://github.com/awslabs/amazon-sagemaker-examples/blob/master/introduction_to_amazon_algorithms/blazingtext_text_classification_dbpedia/blazingtext_text_classification_dbpedia.ipynb)をご参照ください。
#
# 要約すると、BlazingTextでは以下のモードが、異なるタイプのインスタンスでサポートされています。
#
#
# | Modes | cbow (supports subwords training) | skipgram (supports subwords training) | batch_skipgram | supervised |
# |:----------------------: |:----: |:--------: |:--------------: | :--------------: |
# | Single CPU instance | ✔ | ✔ | ✔ | ✔ |
# | Single GPU instance | ✔ | ✔ | - | ✔ (Instance with 1 GPU only) |
# | Multiple CPU instances | - | - | ✔ | - | |
#
# 最後に S3 のデータを指定して fit を呼べば学習を始めることができます。
# ## Create BlazingText Container
# +
import boto3
region_name = boto3.Session().region_name
container = sagemaker.image_uris.retrieve("blazingtext", region_name)
print('Using SageMaker BlazingText container: {} ({})'.format(container, region_name))
# -
# ## Creating an Estimator and start a training job
# +
bt_model = sagemaker.estimator.Estimator(
container,
role=sagemaker.get_execution_role(),
instance_count=1,
instance_type='ml.m5.xlarge',
input_mode= 'File',
sagemaker_session=sess
)
bt_model.set_hyperparameters(
mode="supervised",
epochs=10,
vector_dim=10,
early_stopping=True,
patience=4,
min_epochs=5
)
# -
# ## Training
bt_model.fit(inputs=data_channels, logs=True)
# ## Inference
# ## (Optional) Batch Inference
# バッチ変換処理を使用してファイルに対して一括で推論を実行します。
# ここではすでにトークナイズ、単語の頻度表現へ変換済みの学習データを使用しますが、新規のデータへ適用する場合は別途実行する必要があります。
'''
# ビルトインのBatch推論は1ファイル6MBまでの制限があるため、データを小さくします(実運用で大きなデータを扱う際はデータを分割する必要があります)
test_data = labeled_df.iloc[test_index][:1000]
print(test_data.shape)
test_data.head()
'''
# データのフォーマットは学習時とは異なり、json形式 or jsonline形式となります
#
# `content-type: application/json`
# ```json
# {
# "instances": ["the movie was excellent", "i did not like the plot ."]
# }
# ```
# top-kを予測したい場合は以下のように変更します。
# ```json
# {
# "instances": ["the movie was excellent", "i did not like the plot ."],
# "configuration": {"k": 2}
# }
# ```
#
# `content-type: application/jsonlines`
# ```jsonline
# {"source": "source_0"}
# {"source": "source_1"}
# ```
# top-kを予測したい場合は以下のように変更します。
# ```jsonline
# {"source": "source_0", "k": 2}
# {"source": "source_1", "k": 3}
# ```
'''
# json形式
import json
d = list(test_data.review_body)
with open('test.json', 'w') as f:
for line in d:
json.dump({"source": line}, f, ensure_ascii=False)
'''
'''
# jsonline形式
import json
d = {"instances": list(test_data.review_body)}
with open('test.json', 'w') as f:
json.dump(d, f, ensure_ascii=False)
'''
'''
s3_test_data = sess.upload_data(path='test.json', key_prefix='amazon-review-data')
print("Test data is uploaded to {}".format(s3_test_data))
'''
'''
bucket = sess.default_bucket()
output_path = f's3://{bucket}/amazon-review-data/output/blazingtext_batch_transform'
'''
'''
transformer = bt_model.transformer(
instance_count = 1,
instance_type = 'ml.m5.xlarge',
output_path = output_path,
strategy = "MultiRecord",
)
transformer.transform(
data = s3_test_data,
data_type = "S3Prefix",
content_type = "application/json",
split_type = "Line",
)
'''
# ### Download-predict-file-from-s3
# バッチ推論されたアウトプットファイルをS3からダウンロードして、精度を確認します。
'''
from sagemaker.s3 import S3Downloader, s3_path_join
# creating s3 uri for result file -> input file + .out
output_file = "test.json.out"
output_path = s3_path_join(output_path, output_file)
# download file
S3Downloader.download(output_path, '.')
'''
'''
with open(output_file) as f:
output = json.load(f)
print(output)
'''
# ## Hosting / Inference
#
# 学習が終わると、作成されたモデルをデプロイして、推論を実行することができます。デプロイは deploy を呼び出すだけでできます。`---`といった出力があるときはデプロイ中で、`!`が出力されるとデプロイが完了です。
#
# エンドポイントは json 形式でリクエストを受け付けますので、serializer の content_type に `application/json` を指定します。
text_classifier = bt_model.deploy(initial_instance_count = 1,instance_type = 'ml.m5.xlarge')
text_classifier.serializer = sagemaker.serializers.IdentitySerializer(content_type = 'application/json')
# デプロイが終わったら推論を実行してみましょう。ここでは negative なレビューを 5件、 positive なレビューを 5件ランダムに選択して推論を実行します。
# +
import json
num_test = 5
test_data = labeled_df.iloc[test_index]
neg_test_data = test_data[test_data.star_rating == '__label__0']
pos_test_data = test_data[test_data.star_rating == '__label__1']
neg_index = np.random.choice(neg_test_data.index, num_test)
pos_index = np.random.choice(pos_test_data.index, num_test)
neg_test_sentences = [text for text in neg_test_data.loc[neg_index]["review_body"].values]
payload = {"instances" : neg_test_sentences}
response = text_classifier.predict(json.dumps(payload))
predictions = json.loads(response)
for i, pred in enumerate(predictions):
print("Ground Truth: {}, Prediction: {} (probability: {})"
.format(0, pred["label"][0][-1], pred["prob"]))
print(neg_test_sentences[i].replace(' ', ''))
print()
pos_test_sentences = [text for text in pos_test_data.loc[pos_index]["review_body"].values]
payload = {"instances" : pos_test_sentences}
response = text_classifier.predict(json.dumps(payload))
predictions = json.loads(response)
for i, pred in enumerate(predictions):
print("Ground Truth: {}, Prediction: {} (probability: {})"
.format(1, pred["label"][0][-1], pred["prob"]))
print(pos_test_sentences[i].replace(' ', ''))
print()
# -
# ## Stop / Close the Endpoint
text_classifier.delete_endpoint()
|
BlazingText/Supervised/AmazonSageMaker_BlazingText(Supervised).ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: mypydev
# language: python
# name: python3
# ---
# # WeatherPy
# ----
#
# #### Note
# * Instructions have been included for each segment. You do not have to follow them exactly, but they are included to help you think through the steps.
# +
# Dependencies and Setup
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import requests
import time
from scipy.stats import linregress
import json
import scipy.stats as st
import os
# Import API key
from api_keys import weather_api_key
# Incorporated citipy to determine city based on latitude and longitude
from citipy import citipy
# Output File (CSV)
output_data_file = os.path.join("output_data", "cities.csv")
# Range of latitudes and longitudes
lat_range = (-90, 90)
lng_range = (-180, 180)
# -
# ## Generate Cities List
# +
# List for holding lat_lngs and cities
lat_lngs = []
cities = []
# Create a set of random lat and lng combinations
lats = np.random.uniform(lat_range[0], lat_range[1], size=1500)
lngs = np.random.uniform(lng_range[0], lng_range[1], size=1500)
lat_lngs = zip(lats, lngs)
# Identify nearest city for each lat, lng combination
for lat_lng in lat_lngs:
city = citipy.nearest_city(lat_lng[0], lat_lng[1]).city_name
# If the city is unique, then add it to a our cities list
if city not in cities:
cities.append(city)
# Print the city count to confirm sufficient count
len(cities)
# -
# ### Perform API Calls
# * Perform a weather check on each city using a series of successive API calls.
# * Include a print log of each city as it'sbeing processed (with the city number and city name).
#
# +
city_name_list = []
cloudiness_list = []
country_list = []
date_list = []
humidity_list = []
lat_list = []
lng_list = []
max_temp_list = []
wind_speed_list = []
index_counter = 0
set_counter = 1
print("Beginning Data Retrieval ")
print("-----------------------------")
base_url = "http://api.openweathermap.org/data/2.5/weather?"
units = "imperial"
query_url = f"{base_url}appid={weather_api_key}&units={units}&q="
# For each city name in cities list, do below things...
for index, city in enumerate(cities, start = 1):
try:
response = requests.get(query_url + city).json()
city_name_list.append(response["name"])
cloudiness_list.append(response["clouds"]["all"])
country_list.append(response["sys"]["country"])
date_list.append(response["dt"])
humidity_list.append(response["main"]["humidity"])
lat_list.append(response["coord"]["lat"])
lng_list.append(response["coord"]["lon"])
max_temp_list.append(response['main']['temp_max'])
wind_speed_list.append(response["wind"]["speed"])
if index_counter > 49:
index_counter = 0
set_counter = set_counter + 1
else:
index_counter = index_counter + 1
print(f"Processing Record {index_counter} of Set {set_counter} : {city}")
except(KeyError, IndexError):
print("City not found. Skipping...")
print("-----------------------------")
print("Data Retrieval Complete")
print("-----------------------------")
# -
# ### Convert Raw Data to DataFrame
# * Export the city data into a .csv.
# * Display the DataFrame
# +
# Create a panda data frame using data retrieved
weather_df = pd.DataFrame({
"City" : city_name_list,
"Cloudiness" : cloudiness_list,
"Country" : country_list,
"Date" : date_list,
"Humidity" : humidity_list,
"Lat" : lat_list,
"Lng" : lng_list,
"Max Temp" : max_temp_list,
"Wind Speed" : wind_speed_list
})
# Save city data into a csv file
path = os.path.join("output_data", "cities.csv")
weather_df.to_csv(path, index = False)
weather_df.head()
# -
weather_df.describe()
# ## Inspect the data and remove the cities where the humidity > 100%.
# ----
# Skip this step if there are no cities that have humidity > 100%.
# +
# Based on the table above, the maximum humidity is 100 so we have nothing to do here
# -
# ## Plotting the Data
# * Use proper labeling of the plots using plot titles (including date of analysis) and axes labels.
# * Save the plotted figures as .pngs.
# ## Latitude vs. Temperature Plot
# +
# Create Latitude vs. Temperature Plot scatter plot
plt.scatter(weather_df["Lat"], weather_df["Max Temp"], facecolor = "steelblue", edgecolor = "black")
# Set title
plt.title("Latitude vs. Temperature")
# Set x axis label
plt.xlabel("Laitude")
# Set y axis label
plt.ylabel("Max Temperature (F)")
# Set grid line
plt.grid(linestyle='-', linewidth=1, alpha = 0.5)
# Save the plotted figure as .pngs
path = os.path.join("Images", "Latitude vs. Temperature Plot.png")
plt.savefig(path)
print("We are analyzing latitude vs temperature and we can realize that once the latitude passes 20, the temperatures starts dropping.")
# -
# ## Latitude vs. Humidity Plot
# +
# Create Latitude vs. Humidity Plot scatter plot
plt.scatter(weather_df["Lat"], weather_df["Humidity"], facecolor = "steelblue", edgecolor = "black")
# Set title
plt.title("Latitude vs. Humidity")
# Set x axis label
plt.xlabel("Laitude")
# Set y axis label
plt.ylabel("Humidity (%)")
# Set grid line
plt.grid(linestyle='-', linewidth=1, alpha = 0.5)
# Save the plotted figure as .pngs
path = os.path.join("Images", "Latitude vs. Humidity Plot.png")
plt.savefig(path)
print("We are analyzing latitude vs humidity and we can notice that humidity is low when the latitude is around zero")
# -
# ## Latitude vs. Cloudiness Plot
# +
# Create Latitude vs. Cloudiness Plot scatter plot
plt.scatter(weather_df["Lat"], weather_df["Cloudiness"], facecolor = "steelblue", edgecolor = "black")
# Set title
plt.title("Latitude vs. Cloudiness")
# Set x axis label
plt.xlabel("Laitude")
# Set y axis label
plt.ylabel("Cloudiness (%)")
# Set y axis limit
plt.ylim(-5,105)
# Set grid line
plt.grid(linestyle='-', linewidth=1, alpha = 0.5)
# Save the plotted figure as .pngs
path = os.path.join("Images", "Latitude vs. Cloudiness Plot.png")
plt.savefig(path)
print("We are analyzing latitude vs cloudiness and we realize that cloudiness is clustered either at zero or at a hundred")
# -
# ## Latitude vs. Wind Speed Plot
# +
# Create Latitude vs. Wind Speed scatter plot
plt.scatter(weather_df["Lat"], weather_df["Wind Speed"], facecolor = "steelblue", edgecolor = "black")
plt.title("Latitude vs. Wind Speed")
# Set x axis label
plt.xlabel("Laitude")
# Set y axis label
plt.ylabel("Wind Speed (%)")
# Set y axis limit
plt.ylim(-2,50)
# Set grid line
plt.grid(linestyle='-', linewidth=1, alpha = 0.5)
# Save the plotted figure as .pngs
path = os.path.join("Images", "Latitude vs. Wind Speed Plot.png")
plt.savefig(path)
print("We are analyzing latitude vs wind speed and we can realize that wind speed is relatively the same across different latitudes")
# -
# ## Linear Regression
# +
# Create Northern and Southern Hemisphere DataFrames
northern_hemisphere = weather_df.loc[weather_df["Lat"] >= 0]
southern_hemisphere = weather_df.loc[weather_df["Lat"] < 0]
# Define function for creating linear agression and scatter plot
def linear_agression(x,y):
print(f"The r-squared is : {round(st.pearsonr(x, y)[0],2)}")
(slope, intercept, rvalue, pvalue, stderr) = linregress(x, y)
regress_values = x * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
plt.scatter(x, y)
plt.plot(x,regress_values,"r-")
return line_eq
# Define a fuction for annotating
def annotate(line_eq, a, b):
plt.annotate(line_eq,(a,b),fontsize=15,color="red")
# -
# #### Northern Hemisphere - Max Temp vs. Latitude Linear Regression
# +
equation = linear_agression(northern_hemisphere["Lat"], northern_hemisphere["Max Temp"])
annotate(equation, 0, 0)
# Set a title
plt.title("Northern Hemisphere - Max Temp vs. Latitude Linear Regression")
# Set xlabel
plt.xlabel("Latitude")
# Set ylabel
plt.ylabel("Max Temp (F)")
# Save the figure
path = os.path.join("Images", "Northern Hemisphere - Max Temp vs. Latitude Linear Regression Plot.png")
plt.savefig(path)
# -
# #### Southern Hemisphere - Max Temp vs. Latitude Linear Regression
# +
equation = linear_agression(southern_hemisphere["Lat"],southern_hemisphere["Max Temp"])
annotate(equation, -30, 50)
# Set a title
plt.title("Southern Hemisphere - Max Temp vs. Latitude Linear Regression")
# Set xlabel
plt.xlabel("Latitude")
# Set ylabel
plt.ylabel("Max Temp (F)")
# Save the figure
path = os.path.join("Images", "Southern Hemisphere - Max Temp vs. Latitude Linear Regression Plot.png")
plt.savefig(path)
print("After looking at both graphs we can see that they are going in oposite directions. As the latitude increases above zero, the temperature drops ")
print("and as the latitude decreases below zero, the temperature increases.")
# -
# #### Northern Hemisphere - Humidity (%) vs. Latitude Linear Regression
# +
equation = linear_agression(northern_hemisphere["Lat"], northern_hemisphere["Humidity"])
annotate(equation, 40, 15)
# Set a title
plt.title("Northern Hemisphere - Humidity (%) vs. Latitude Linear Regression")
# Set xlabel
plt.xlabel("Latitude")
# Set ylabel
plt.ylabel("Humidity (%)")
# Save the figure
path = os.path.join("Images", "Northern Hemisphere - Humidity (%) vs. Latitude Linear Regression Plot.png")
plt.savefig(path)
# -
# #### Southern Hemisphere - Humidity (%) vs. Latitude Linear Regression
# +
equation = linear_agression(southern_hemisphere["Lat"], southern_hemisphere["Humidity"])
annotate(equation, -40, 50)
# Set title
plt.title("Southern Hemisphere - Humidity (%) vs. Latitude Linear Regression")
# Set xlabel
plt.xlabel("Latitude")
# Set ylabel
plt.ylabel("Humidity (%)")
# Save the figure
path = os.path.join("Images", "Southern Hemisphere - Humidity (%) vs. Latitude Linear Regression Plot.png")
plt.savefig(path)
print("After looking at both graphs, we notice that humidity changes in a similar way in both the southern and northern hemispheres")
# -
# #### Northern Hemisphere - Cloudiness (%) vs. Latitude Linear Regression
# +
equation = linear_agression(northern_hemisphere["Lat"], northern_hemisphere["Cloudiness"])
annotate(equation, 30, 40)
# Set a title
plt.title("Northern Hemisphere - Cloudiness (%) vs. Latitude Linear Regression")
# Set xlabel
plt.xlabel("Latitude")
# Set ylabel
plt.ylabel("Cloudiness (%)")
# Save the figure
path = os.path.join("Images", "Northern Hemisphere - Cloudiness (%) vs. Latitude Linear Regression Plot.png")
plt.savefig(path)
# -
# #### Southern Hemisphere - Cloudiness (%) vs. Latitude Linear Regression
# +
equation = linear_agression(southern_hemisphere["Lat"], southern_hemisphere["Cloudiness"])
annotate(equation, -30, 40)
# Set a title
plt.title("Southern Hemisphere - Cloudiness (%) vs. Latitude Linear Regression")
# Set xlabel
plt.xlabel("Latitude")
# Set ylabel
plt.ylabel("Cloudiness (%)")
# Save the figure
path = os.path.join("Images", "Southern Hemisphere - Cloudiness (%) vs. Latitude Linear Regression Plot.png")
plt.savefig(path)
print("After looking at both graphs, we notice that cloudiness changes in a similar way in both the southern and northern hemispheres ")
print("however, the points in the southern hemisphere are slightly shifted to the right")
# -
# #### Northern Hemisphere - Wind Speed (mph) vs. Latitude Linear Regression
# +
equation = linear_agression(northern_hemisphere["Lat"], northern_hemisphere["Wind Speed"])
annotate(equation, 40, 20)
# Set title
plt.title("Northern Hemisphere - Wind Speed (mph) vs. Latitude Linear Regression")
# Set xlabel
plt.xlabel("Latitude")
# Set ylabel
plt.ylabel("Wind Speed (mph)")
# Save the figure
path = os.path.join("Images", "Northern Hemisphere - Wind Speed vs. Latitude Linear Regression Plot.png")
plt.savefig(path)
# -
# #### Southern Hemisphere - Wind Speed (mph) vs. Latitude Linear Regression
# +
equation = linear_agression(southern_hemisphere["Lat"], southern_hemisphere["Wind Speed"])
annotate(equation, -30, 15)
# Set title
plt.title("Southern Hemisphere - Wind Speed (mph) vs. Latitude Linear Regression")
# Set xlabel
plt.xlabel("Latitude")
# Set ylabel
plt.ylabel("Wind Speed (mph)")
# Save the figure
path = os.path.join("Images", "Southern Hemisphere - Wind Speed vs. Latitude Linear Regression Plot.png")
plt.savefig(path)
print("After looking at both graphs we can see that they are going in oposite directions. As the latitude increases above zero, the wind speed increases ")
print("and as the latitude decreases below zero, the wind speed decreases.")
# -
|
WeatherPy/WeatherPy.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <div class="alert alert-block alert-info">
# __Name__: heidelburg_train_classifiers<br/>
# __Description__: Assess and visualize FOX amr prediction performance in S. Heidelburg <br/>
# __Author__: <NAME> matthew dot whiteside at canada dot ca<br/>
# __Date__: Nov 6, 2017<br/>
# __TODO__:<br/>
# </div>
# %load_ext autoreload
# %autoreload 2
import numpy as np
import pandas as pd
from sklearn.externals import joblib
from sklearn.metrics import f1_score, classification_report
import xgboost as xgb
import os
os.chdir('../pangenome')
import utils
import classify
import config
amr = joblib.load(config.SH['amr'])
amr_list = joblib.load(config.SH['amr_list'])
sample_index = joblib.load(config.SH['sample_index'])
pg = joblib.load(config.SH['pg'])
locus_list = joblib.load(config.SH['locus_list'])
test_train_index = joblib.load(config.SH['test_train_index'])
rfc = joblib.load(config.SH['fox_rfc'])
gbc = joblib.load(config.SH['fox_gbc'])
xbc = joblib.load(config.SH['fox_xbc'])
X_train = pg[test_train_index == 'Training',:].toarray()
X_test = pg[test_train_index == 'Validation',:].toarray()
y_train = amr[test_train_index == 'Training', amr_list == 'FOX']
y_test = amr[test_train_index == 'Validation', amr_list == 'FOX']
predictions = xbc.predict(X_test)
print(classification_report(predictions, y_test))
import matplotlib as plt
xgb.plot_importance(xbc)
plt.pyplot.show()
# +
xbc._Booster.dump_model('trees.txt', with_stats=True)
# -
locus_list[np.array(xbc._Booster.feature_names) == 'f5144']
xbc
locus_list[4653]
locus_list[4652]
|
jupyter/heidelburg_fox_performance.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Programming Exercise 2: Logistic Regression
# +
# imports here
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
# -
# ## Logistic Regression
# In this part of the exercise, you will build a logistic regression model to predict whether a student gets admitted into a university.
#
# Suppose that you are the administrator of a university department and you want to determine each applicant’s chance of admission based on their results on two exams. You have historical data from previous applicants that you can use as a training set for logistic regression. For each training example, you have the applicant’s scores on two exams and the admissions decision.
#
# Your task is to build a classification model that estimates an applicant’s probability of admission based the scores from those two exams.
# ### Visualizing the data
# Before starting to implement any learning algorithm, it is always good to visualize the data if possible.
# +
def plot_data():
from numpy import loadtxt, where
from pylab import scatter, show, legend, xlabel, ylabel
#load the dataset
data = loadtxt('ex2data1.txt', delimiter=',')
X = data[:, 0:2]
y = data[:, 2]
pos = where(y == 1)
neg = where(y == 0)
scatter(X[pos, 0], X[pos, 1], marker='o', c='y')
scatter(X[neg, 0], X[neg, 1], marker='x', c='b')
xlabel('Exam 1 score')
ylabel('Exam 2 score')
legend(['Admitted', 'Not Admitted'])
plt.title('Scatter plot of training data')
show()
plot_data()
# -
data = pd.read_csv("ex2data1.txt", sep=",", header=None, names=['X1', 'X2', 'y'])
data.head()
X = data[['X1', 'X2']]
y = data['y']
# +
plt.scatter(data['X1'],data['X2'],c=y)
plt.xlabel('Exam 1 score')
plt.ylabel('Exam 2 score')
plt.title('Scatter plot of training data')
plt.show()
# -
# ## Implementation
# ### Sigmoid function
# Before you start with the actual cost function, recall that the logistic regression hypothesis is defined as: $h_{\theta}(x) = g(\theta^{T}x)$, where function g is the sigmoid function. The sigmoid function is defined as: $g(z) = \frac{1}{1+e^{-z}}$ .
# +
def sigmoid(z):
return 1 / (1 + np.exp(-z))
sigmoid(0)
# -
# ### Cost function and gradient
# Now you will implement the cost function and gradient for logistic regression. Recall that the cost function in logistic regression is $J(\theta) = \frac{1}{m}\sum_{i=1}^{m}[-y^{(i)}log(h_{\theta}(x^{(i)})) - (1-y^{(i)})log(1-h_{\theta}(x^{(i)}))]$ and the gradient of the cost is a vector of the same length as $\theta$ where the j th element (for j = 0, 1, . . . , n) is defined as follows: $\frac{\partial J(\theta )}{\partial \theta _{j}} = \frac{1}{m}\sum_{i=1}^{m}(h_{\theta }(x^{(i)})-y^{(i)})x_{j}^{(i)}$ . Note that while this gradient looks identical to the linear regression gradient, the formula is actually different because linear and logistic regression have different definitions of $h_{\theta }(x)$.
def costFunction(theta, X, y):
m = y.shape[0]
h_theta = sigmoid(np.matmul(X, theta))
error = np.matmul(y.T, np.log(h_theta)) + np.matmul((1-y).T, np.log(1 - h_theta))
J = -error / m
grad = np.matmul(X.T, (h_theta-y)) / m
return (J, grad)
m, n = X.shape # Number of features
ones = pd.DataFrame(np.ones((m, 1)), columns=['ones'])
X = pd.concat([ones, X], axis=1) # Add a column of ones to X
X.head()
# +
initial_theta = np.zeros((n + 1, 1))
cost, grad = costFunction(initial_theta, X, y.values.reshape((m, 1)))
print(cost)
print(grad)
# -
# ### Learning parameters
# +
import scipy.optimize as opt
result = opt.fmin_tnc(func=costFunction, x0=initial_theta, args=(X, y))
# -
result
# ### Evaluating logistic regression
# For a student with an Exam 1 score of 45 and an Exam 2 score of 85, you should expect to see an admission probability of 0.776.
theta = result[0]
probability = sigmoid(np.matmul([1, 45, 85], theta.T))
probability
# +
def classifierPredict(theta, X):
"""
take in numpy array of theta and X and predict the class
"""
predictions = np.matmul(X, theta)
return predictions > 0
p = classifierPredict(theta, X)
print("Train Accuracy: {} %".format(sum(p==y)))
# -
# ### Plotting the decision boundary
# +
# Only need 2 points to define a line, so choose two endpoints
def plotDecisionBoundary(theta, X, y):
plt.scatter(X.X1, X.X2, c=y)
plt.xlabel('Exam 1 score')
plt.ylabel('Exam 2 score')
plt.title('Scatter plot of training data')
plot_X1 = np.array([min(X.X1), max(X.X1)])
plot_X2 = -(theta[0] + theta[1] * plot_X1)/theta[2]
plt.plot(plot_X1, plot_X2)
plt.show()
plotDecisionBoundary(theta, X, y)
# +
def plotDecisionBoundary(theta, X, y):
from numpy import loadtxt, where
from pylab import scatter, show, legend, xlabel, ylabel
#load the dataset
data = loadtxt('ex2data1.txt', delimiter=',')
X = data[:, 0:2]
y = data[:, 2]
pos = where(y == 1)
neg = where(y == 0)
scatter(X[pos, 0], X[pos, 1], marker='o', c='y')
scatter(X[neg, 0], X[neg, 1], marker='x', c='b')
xlabel('Exam 1 score')
ylabel('Exam 2 score')
plt.title('Scatter plot of training data')
plot_X1 = np.array([min(X[:, 0]), max(X[:, 0])])
plot_X2 = -(theta[0] + theta[1] * plot_X1)/theta[2]
plt.plot(plot_X1, plot_X2)
legend(['Decision Boundary', 'Admitted', 'Not Admitted'])
show()
plotDecisionBoundary(theta, X, y)
# -
|
Week 3/Notebook-ex2/Programming Exercise 2- Logistic Regression.ipynb
|
(* --- *)
(* jupyter: *)
(* jupytext: *)
(* text_representation: *)
(* extension: .ml *)
(* format_name: light *)
(* format_version: '1.5' *)
(* jupytext_version: 1.14.4 *)
(* kernelspec: *)
(* display_name: OCaml cs3110-finalproj *)
(* language: OCaml *)
(* name: ocaml-jupyter *)
(* --- *)
#require "jupyter-archimedes"
#use "k_means.ml"
(* +
let rec gen_clusters acc n k r =
if n = 0 then acc
else
let rec helper acc c k =
if k = 0 then acc
else
let theta = Random.float Float.pi in
let r' = Random.float r in
let x = r' *. Float.cos theta in
let y = r' *. Float.sin theta in
helper (((fst c) +. x, (snd c) +. y) :: acc) c (k - 1) in
gen_clusters (acc @ (helper [] (Random.float 60.0, Random.float 60.0) k)) (n - 1) k r
let pp = gen_clusters [] 3 40 12.0
let pp_train = construct pp
let res = classify pp_train 3 100
let centroids = fst res |> to_list
let pp_labeled = snd res
(* -
let vp = A. init ~w:600. ~h:600. ["jupyter"] in
A.Axes.box vp ;
for i = 0 to pred (Array.length pp_labeled) do
if pp_labeled.(i).tag = 0 then
A.set_color vp A.Color.red ;
if pp_labeled.(i).tag = 1 then
A.set_color vp A.Color.blue ;
if pp_labeled.(i).tag = 2 then
A.set_color vp A.Color.green ;
A.List.xy_pairs vp [pp_labeled.(i).pos];
done;
A.set_color vp A.Color.black ;
A.List.xy_pairs vp centroids ;
A.close vp
|
k_means.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: fenotebook
# language: python
# name: fenotebook
# ---
#
# ## Feature Creation: Combine with reference feature
#
# The CombineWithReferenceFeature() applies combines a group of variables with a group of reference variables utilising mathematical operations ['sub', 'div','add','mul'], returning one or more additional features as a result.
#
# For this demonstration, we use the UCI Wine Quality Dataset.
#
# The data is publicly available on [UCI repository](https://archive.ics.uci.edu/ml/datasets/Wine+Quality)
#
# <NAME>, <NAME>, <NAME>, <NAME> and <NAME>. Modeling wine preferences by data mining from physicochemical properties. In Decision Support Systems, Elsevier, 47(4):547-553, 2009.
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import (
accuracy_score,
roc_curve,
roc_auc_score,
classification_report,
confusion_matrix,
)
from sklearn.pipeline import Pipeline as pipe
from sklearn.preprocessing import StandardScaler
from feature_engine.creation import CombineWithReferenceFeature
from feature_engine.imputation import MeanMedianImputer
pd.set_option('display.max_columns', None)
# +
# Read data
data = pd.read_csv('winequality-red.csv', sep=';')
data.head()
# -
# **This Data contains 11 features, all numerical, with no missing values.**
# +
# Let's transform the Target, i.e Wine Quality into a binary classification problem:
bins = [0,5,10]
labels = [0, 1] # 'low'=0, 'high'=1
data['quality_range']= pd.cut(x=data['quality'], bins=bins, labels=labels)
data[['quality_range','quality']].head(5)
# +
# drop original target
data.drop('quality', axis=1, inplace = True)
# -
# ### Sub and Div Combinators:
#
# Let's create two new variables:
#
# - non_free_sulfur_dioxide = total sulfur dioxide - free sulfur dioxide
# - percentage_free_sulfur = free sulfur dioxide / total sulfur dioxide
# +
# Create the Combinators
# this transformer substracts free sulfur from total sulfur
sub_with_reference_feature = CombineWithReferenceFeature(
variables_to_combine=['total sulfur dioxide'],
reference_variables=['free sulfur dioxide'],
operations=['sub'],
new_variables_names=['non_free_sulfur_dioxide']
)
# this transformer divides free sulfur by total sulfur
div_with_reference_feature = CombineWithReferenceFeature(
variables_to_combine=['free sulfur dioxide'],
reference_variables=['total sulfur dioxide'],
operations=['div'],
new_variables_names=['percentage_free_sulfur']
)
# Fit the Sub Combinator on training data
sub_with_reference_feature.fit(data)
# perform the substraction
data_t = sub_with_reference_feature.transform(data)
# perform division
# We can combine both steps in a single call with ".fit_transform()" method
data_t = div_with_reference_feature.fit_transform(data_t)
# +
# Note the additional variables at the end of the dataframe
data_t.head()
# -
# #### Combine with more than 1 operation
#
# We can also combine the variables with more than 1 mathematical operation. And the transformer has the option to create variable names automatically.
#
# Here we will create the following variables:
#
# - ratio_fixed_to_volatile_acidity = fixed acidity / volatile acidity
# - total_acidity = fixed acidity + volatile acidity
# +
# Create the Combinator
multiple_combinator = CombineWithReferenceFeature(
variables_to_combine=['fixed acidity'],
reference_variables=['volatile acidity'],
operations=['div', 'add'],
new_variables_names=['ratio_fixed_to_volatile', 'total_acidity']
)
# +
# Fit the Combinator to the training data
multiple_combinator.fit(data_t)
# +
# Transform the data
data_t = multiple_combinator.transform(data_t)
# +
# Note the additional variables at the end of the dataframe
data_t.head()
# -
# ### Pipeline Example
#
# We can put all these transformations into single pipeline:
#
# Create new variables scale features and train a Logistic Regression model to predict the wine quality range.
#
# See more on how to use Feature-engine within Scikit-learn Pipelines in these [examples](https://github.com/solegalli/feature_engine/tree/master/examples/Pipelines)
# +
X = data.drop(['quality_range'], axis=1)
y = data.quality_range
X_train, X_test, y_train, y_test = train_test_split(X,
y,
test_size=0.1,
random_state=0,
shuffle=True,
stratify=y
)
X_train.shape, X_test.shape
# -
value_pipe = pipe([
# Create new features
('subtraction', CombineWithReferenceFeature(
variables_to_combine=['total sulfur dioxide'],
reference_variables=['free sulfur dioxide'],
operations=['sub'],
new_variables_names=['non_free_sulfur_dioxide']
)
),
('ratio', CombineWithReferenceFeature(
variables_to_combine=['free sulfur dioxide'],
reference_variables=['total sulfur dioxide'],
operations=['div'],
new_variables_names=['percentage_free_sulfur']
)
),
('acidity', CombineWithReferenceFeature(
variables_to_combine=['fixed acidity'],
reference_variables=['volatile acidity'],
operations=['div', 'add'],
new_variables_names=['ratio_fixed_to_volatile', 'total_acidity']
)
),
# scale features
('scaler', StandardScaler()),
# Logistic Regression
('LogisticRegression', LogisticRegression())
])
value_pipe.fit(X_train, y_train)
pred_train = value_pipe.predict(X_train)
pred_test = value_pipe.predict(X_test)
# +
print('Logistic Regression Model train accuracy score: {}'.format(
accuracy_score(y_train, pred_train)))
print()
print('Logistic Regression Model test accuracy score: {}'.format(
accuracy_score(y_test, pred_test)))
# -
print('Logistic Regression Model test classification report: \n\n {}'.format(
classification_report(y_test, pred_test)))
# +
score = round(accuracy_score(y_test, pred_test), 3)
cm = confusion_matrix(y_test, pred_test)
sns.heatmap(cm, annot=True, fmt=".0f")
plt.xlabel('Predicted Values')
plt.ylabel('Actual Values')
plt.title('Accuracy Score: {0}'.format(score), size=15)
plt.show()
# +
# Predict probabilities for the test data
probs = value_pipe.predict_proba(X_test)[:, 1]
# Get the ROC Curve
fpr, tpr, thresholds = roc_curve(y_test, probs)
# Plot ROC curve
plt.figure(figsize=(8, 5))
plt.plot([0, 1], [0, 1], 'k--')
plt.plot(fpr, tpr)
plt.xlabel('False Positive Rate = 1 - Specificity Score')
plt.ylabel('True Positive Rate = Recall Score')
plt.title('ROC Curve')
plt.show()
# -
|
examples/creation/CombineWithReferenceFeature.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <script async src="https://www.googletagmanager.com/gtag/js?id=UA-59152712-8"></script>
# <script>
# window.dataLayer = window.dataLayer || [];
# function gtag(){dataLayer.push(arguments);}
# gtag('js', new Date());
#
# gtag('config', 'UA-59152712-8');
# </script>
#
# # `NRPyPlusTOVID`: An Einstein Toolkit Thorn for Piecewise-Polytrope TOV neutron star initial data
#
# ## Author: <NAME> and <NAME>
#
# [comment]: <> (Abstract: TODO)
#
# **Notebook Status:** <font color='orange'><b> Partially Validated </b></font>
#
# **Validation Notes:** NRPy+ TOV initial data generation module validated against [J<NAME>'s TOV initial data solver](https://ccrg.rit.edu/~jfaber/BNSID/TOV/), as described in the [NRPy+ implementation notes of the TOV solution for piecewise-polytrope neutron stars](Tutorial-TOV-Piecewise_Polytrope_EOSs.ipynb).
#
# ### NRPy+ Source Code for this module: [TOV/TOV_Solver.py](../edit/TOV/TOV_Solver.py) [\[tutorial\]](Tutorial-Tutorial-ADM_Initial_Data-TOV.ipynb) Constructs numerical solution to TOV equations for neutron stars with piecewise polytrope equations of state
#
# ## Introduction:
# In this part of the tutorial, we will construct an Einstein Toolkit (ETK) thorn (module) that will set up [TOV initial data](https://en.wikipedia.org/wiki/Tolman–Oppenheimer–Volkoff_equation) for an equilibrium neutron star. As documented in the [Piecewise Polytrope NRPy+ tutorial](Tutorial-TOV-Piecewise_Polytrope_EOSs.ipynb), piecewise-polytrope equations of state are supported, which closely approximate realistic nuclear equations of state appropriate for neutron star matter. In the [Tutorial-Tutorial-ADM_Initial_Data-TOV](Tutorial-Tutorial-ADM_Initial_Data-TOV.ipynb) tutorial notebook, we used NRPy+ to construct the SymPy expressions for these initial data.
#
# We will construct this thorn in two steps.
#
# 1. Call on NRPy+ to convert the SymPy expressions for the initial data into one C-code kernel.
# 1. Write the C code and linkages to the Einstein Toolkit infrastructure (i.e., the .ccl files) to complete this Einstein Toolkit module.
# <a id='toc'></a>
#
# # Table of Contents
# $$\label{toc}$$
#
# This notebook is organized as follows
#
# 1. [Step 1](#initializenrpy): **Call on NRPy+ to generate the TOV solution given a piecewise-polytrope equation of state; output the data to a text file**
# 1. [Step 2](#initial_data): **Converting TOV initial data so that it can be used by the Einstein Toolkit**
# 1. [Step 2.a](#initial_data__interpolation): Interpolate the TOV data file as needed
# 1. [Step 2.b](#initial_data__tov_to_adm_sph): Converting the TOV variables to ADM variables in Spherical coordinates
# 1. [Step 2.c](#initial_data__convert_adm_sph_to_admbase): Convert Spherical ADM quantities to `ADMBase` (Cartesian) variables $\left\{\alpha,\beta^i,\gamma_{ij},K_{ij}\right\}$
# 1. [Step 2.d](#initial_data__convert_to_hydrobase): Convert TOV solution quantities to `HydroBase` variables $\left\{P,\rho_{\rm baryonic},\epsilon,v_{\rm (n)}^i\right\}$
# 1. [Step 3](#einstein): **Interfacing with the Einstein Toolkit**
# 1. [Step 3.a](#einstein_c): Constructing the Einstein Toolkit C-code calling functions that include the C code kernels
# 1. [Step 3.b](#einstein_ccl): CCL files - Define how this module interacts and interfaces with the larger Einstein Toolkit infrastructure
# 1. [Step 3.c](#einstein_list): Add the C code to the Einstein Toolkit compilation list
# 1. [Step 4](#latex_pdf_output): **Output this notebook to $\LaTeX$-formatted PDF**
# <a id='initializenrpy'></a>
#
# # Step 1: Call on NRPy+ to generate the TOV solution given a piecewise-polytrope equation of state; output the data to a text file \[Back to [top](#toc)\]
# $$\label{initializenrpy}$$
#
#
# +
# Step 1: Import needed core NRPy+ modules
from outputC import lhrh # NRPy+: Core C code output module
import sympy as sp # SymPy: The Python computer algebra package upon which NRPy+ depends
import finite_difference as fin # NRPy+: Finite difference C code generation module
import NRPy_param_funcs as par # NRPy+: Parameter interface
import grid as gri # NRPy+: Functions having to do with numerical grids
import loop as lp # NRPy+: Generate C code loops
import indexedexp as ixp # NRPy+: Symbolic indexed expression (e.g., tensors, vectors, etc.) support
import reference_metric as rfm # NRPy+: Reference metric support
import cmdline_helper as cmd # NRPy+: Multi-platform Python command-line interface
import shutil, os, sys # Standard Python modules for multiplatform OS-level functions
# Create directory for NRPyPlusTOVID thorn & subdirectories in case they don't exist.
outrootdir = "NRPyPlusTOVID/"
cmd.mkdir(os.path.join(outrootdir))
outdir = os.path.join(outrootdir,"src") # Main C code output directory
cmd.mkdir(outdir)
# Step 1.a: This is an Einstein Toolkit (ETK) thorn. Here we
# tell NRPy+ that gridfunction memory access will
# therefore be in the "ETK" style.
par.set_parval_from_str("grid::GridFuncMemAccess","ETK")
par.set_parval_from_str("grid::DIM", 3)
DIM = par.parval_from_str("grid::DIM")
# Step 1.b: NRPyPlusTOVID uses Cartesian coordinates, so
# we specify the reference metric to be Cartesian here:
par.set_parval_from_str("reference_metric::CoordSystem","Cartesian")
rfm.reference_metric()
# ADJUST THIS PARAMETER IF DESIRED.
# "Single" = Single Polytrope
# "APR4" = APR4 Piecewise Polytrope
# "SLY" = SLy Piecewise Polytrope
# .---------------------------------------------.
# | For all available names please look in the |
# | TOV/Piecewise_Polytrope__dict.py NRPy+ file |
# .---------------------------------------------.
# vvvvvvvvvvvvvvvv
EOSname = "Single"
# EOSname = "SLy"
# EOSname = "APR4"
# ^^^^^^^^^^^^^^^^
# Import our TOV solver, which supports both single
# and piecewise polytropic EOSs
import TOV.TOV_Solver as TOV
import TOV.Polytropic_EOSs as poly
if EOSname=="Single":
# Set neos = 1 (single polytrope)
neos = 1
# Set rho_poly_tab (not needed for a single polytrope)
rho_poly_tab = []
# Set Gamma_poly_tab
Gamma_poly_tab = [2.0]
# Set K_poly_tab0
K_poly_tab0 = 1. # ZACH NOTES: CHANGED FROM 100.
rhob_central = 0.129285309 # M/R_Schw = 1.468770268913230e-01
# Set the eos quantities
eos = poly.set_up_EOS_parameters__complete_set_of_input_variables(neos,rho_poly_tab,Gamma_poly_tab,K_poly_tab0)
import time
start = time.time()
TOV.TOV_Solver(eos,
outfile="outputTOVpolytrope.txt",
rho_baryon_central=rhob_central,
verbose = True)
print("Single Polytrope TOV solution generated in: "+str(time.time()-start)+" s")
print("Initial data file: outputTOVpolytrope.txt")
else:
# Set up the EOS parameters
eos = poly.set_up_EOS_parameters__Read_et_al_input_variables(EOSname)
# Set up the initial condition for the pressure by
# selecting a central baryon density
# rhob_central = 2.0 # M/R_Schw = 3.303692404611947e-01
# rhob_central = 1.0 # M/R_Schw = 2.051637558540178e-01
rhob_central = 0.8 # M/R_Schw = 1.470662481999595e-01
# Solve the TOV equations given our EOS and central density
import time
start = time.time()
outfilename = "outputTOVpolytrope-"+EOSname+".txt"
TOV.TOV_Solver(eos,outfile=outfilename,rho_baryon_central=rhob_central,verbose=True)
print("PPEOS "+EOSname+" TOV solution generated in: "+str(time.time()-start)+" s")
print("Initial data file: "+outfilename)
# -
# <a id='initial_data'></a>
#
# # Step 2: Converting TOV initial data so that it can be used by the Einstein Toolkit \[Back to [top](#toc)\]
# $$\label{initial_data}$$
#
# Main driver function:
#
# * Looping over all gridpoints:
# * Read in `const CCTK_REAL rr = r[CCTK_GFINDEX3D(cctkGH,i0,i1,i2)];`
# * **Given this radius call interpolation driver to get all the base TOV quantities**
# * **Convert TOV spacetime quantities to ADM quantities in *spherical* basis**
# * Call the Cartesian ADMBase converter
# * Call the HydroBase converter
#
# <a id='initial_data__interpolation'></a>
#
# ## Step 2.a: Interpolate the TOV data file as needed \[Back to [top](#toc)\]
# $$\label{initial_data__interpolation}$$
#
# We start by interpolating the TOV data file to the gridpoints used by ETK, using the [tov_interp.h](../edit/TOV/tov_interp.h) file, which using Lagrange polynomial interpolation (for more details on the usage of this interpolation file, please look at the [start-to-finish TOV initial data tutorial notebook](Tutorial-Start_to_Finish-BSSNCurvilinear-Setting_up_TOV_initial_data.ipynb)).
#
# Keep in mind that the TOV data file just written stored $\left(r,\rho(r),\rho_{\text{baryonic}}(r),P(r),M(r),e^{\nu(r)}\right)$, where $\rho(r)$ is the total mass-energy density (cf. $\rho_{\text{baryonic}}$).
# +
shutil.copy(os.path.join("TOV","tov_interp.h"),outdir)
with open(os.path.join(outdir,"interpolate_TOV_solution_to_point.h"), "w") as file:
file.write("""
/* Load the TOV_interpolate_1D() function */
#include "tov_interp.h"
/* This function returns the TOV quantities at point rr
* by interpolating the data in the TOV initial data file.
*/
void interpolate_TOV_solution_to_point(const CCTK_REAL rr, ID_inputs other_inputs,
CCTK_REAL *exp_4phi, CCTK_REAL *expnu,
CCTK_REAL *Pressure, CCTK_REAL *rho_baryon, CCTK_REAL *rho__total_energy_density) {
/* The mass valus is not used, but we have to
* store it in this dummy variable because the
* initial data file contains it.
*/
CCTK_REAL M;
/* Perform the interpolation, returning:
* - rho__total_energy_density
* - rho_baryon
* - Pressure
* - Mass (dummy variable, unused)
* - exp(nu)
* - exp(4phi)
*/
TOV_interpolate_1D(rr,other_inputs.Rbar,other_inputs.Rbar_idx,other_inputs.interp_stencil_size,
other_inputs.numlines_in_file,
other_inputs.r_Schw_arr,other_inputs.rho_arr,other_inputs.rho_baryon_arr,other_inputs.P_arr,other_inputs.M_arr,
other_inputs.expnu_arr,other_inputs.exp4phi_arr,other_inputs.rbar_arr,
rho__total_energy_density,rho_baryon,Pressure,&M,expnu,exp_4phi);
}\n""")
# -
# <a id='initial_data__tov_to_adm_sph'></a>
#
# ## Step 2.b: Converting the TOV variables to ADM variables in Spherical coordinates \[Back to [top](#toc)\]
# $$\label{initial_data__tov_to_adm_sph}$$
#
# Now we perform the interpolation of the TOV quantities to ADM quantities in spherical coordinates, using (see [the TOV initial data tutorial notebook](Tutorial-ADM_Initial_Data-TOV.ipynb) for more details):
#
#
# \begin{equation}
# \boxed{
# \begin{aligned}
# \alpha &= e^{\nu(\bar{r})/2} \\
# \beta^k &= 0 \\
# \gamma_{\bar{r}\bar{r}} &= e^{4\phi}\\
# \gamma_{\theta\theta} &= e^{4\phi} \bar{r}^2 \\
# \gamma_{\phi\phi} &= e^{4\phi} \bar{r}^2 \sin^2 \theta \\
# \end{aligned}
# }
# \end{equation}
with open(os.path.join(outdir,"convert_TOV_spacetime_vars_to_ADM_vars.h"), "w") as file:
file.write("""
/* This function converts TOV quantities into
* ADM quantities in Spherical coordinates.
*/
void convert_TOV_spacetime_vars_to_ADM_vars( const CCTK_REAL rr, const CCTK_REAL th,
const CCTK_REAL IDexp_4phi, const CCTK_REAL IDexpnu,
CCTK_REAL *IDalpha,
CCTK_REAL *IDgammaDD00, CCTK_REAL *IDgammaDD01, CCTK_REAL *IDgammaDD02,
CCTK_REAL *IDgammaDD11, CCTK_REAL *IDgammaDD12, CCTK_REAL *IDgammaDD22) {
/***************************************************************
* Convert TOV quantities to ADM quantities in Spherical basis *
***************************************************************
*
* First we convert the lapse function:
* .------------------.
* | alpha = e^(nu/2) |
* .------------------.
*/
*IDalpha = sqrt(IDexpnu);
/* Next we convert the metric function:
* .----------------------------------------.
* | gamma_{00} = e^{4phi} |
* .----------------------------------------.
* | gamma_{11} = e^{4phi} r^2 |
* .----------------------------------------.
* | gamma_{22} = e^{4phi} r^2 sin^2(theta) |
* .----------------------------------------.
* | All other components are zero. |
* .----------------------------------------.
*/
*IDgammaDD00 = IDexp_4phi;
*IDgammaDD11 = IDexp_4phi * rr * rr;
*IDgammaDD22 = IDexp_4phi * rr * rr * sin(th) * sin(th);
*IDgammaDD01 = 0.0;
*IDgammaDD02 = 0.0;
*IDgammaDD12 = 0.0;
}\n""")
# <a id='initial_data__convert_adm_sph_to_admbase'></a>
#
# ## Step 2.c: Convert Spherical ADM quantities to `ADMBase` (Cartesian) variables $\left\{\alpha,\beta^i,\gamma_{ij},K_{ij}\right\}$ \[Back to [top](#toc)\]
# $$\label{initial_data__convert_adm_sph_to_admbase}$$
#
# The [TOV line element](https://en.wikipedia.org/wiki/Tolman%E2%80%93Oppenheimer%E2%80%93Volkoff_equation) in *Schwarzschild coordinates* is written (in the $-+++$ form):
# $$
# ds^2 = - c^2 e^\nu dt^2 + \left(1 - \frac{2GM}{rc^2}\right)^{-1} dr^2 + r^2 d\Omega^2.
# $$
#
# In *isotropic coordinates* with $G=c=1$ (i.e., the initial coordinate slicing and units we prefer to use), the ($-+++$ form) line element is written:
# $$
# ds^2 = - e^{\nu} dt^2 + e^{4\phi} \left(d\bar{r}^2 + \bar{r}^2 d\Omega^2\right),
# $$
# where $\phi$ here is the *conformal factor*.
#
# The ADM 3+1 line element for this diagonal metric in isotropic spherical coordinates is given by:
# $$
# ds^2 = (-\alpha^2 + \beta_k \beta^k) dt^2 + \gamma_{\bar{r}\bar{r}} d\bar{r}^2 + \gamma_{\theta\theta} d\theta^2+ \gamma_{\phi\phi} d\phi^2,
# $$
#
# from which we can immediately read off the ADM quantities:
# \begin{align}
# \alpha &= e^{\nu(\bar{r})/2} \\
# \beta^k &= 0 \\
# \gamma_{\bar{r}\bar{r}} &= e^{4\phi}\\
# \gamma_{\theta\theta} &= e^{4\phi} \bar{r}^2 \\
# \gamma_{\phi\phi} &= e^{4\phi} \bar{r}^2 \sin^2 \theta \\
# \end{align}
# +
thismodule = __name__
IDalpha = par.Cparameters("REAL", thismodule, "IDalpha", 1e300) # IDalpha must be set in C
IDbetaU = ixp.zerorank1() # beta^i is zero
IDgammaDD = ixp.zerorank2()
for i in range(3):
for j in range(i,3):
IDgammaDD[i][j] = par.Cparameters("REAL", thismodule, "IDgammaDD"+str(i)+str(j), 1e300) # IDgammaDD must be set in C
IDgammaDD[j][i] = IDgammaDD[i][j]
IDKDD = ixp.zerorank2() # K_{ij} is zero
# -
# As this ETK module expects Cartesian coordinates, and the TOV solution above is in the spherical basis, we next perform the Jacobian transformations necessary to convert into the Cartesian basis:
#
# All ADM tensors and vectors are in the Spherical coordinate basis $x^i_{\rm Sph} = (r,\theta,\phi)$, but we need them in the Cartesian coordinate basis $x^i_{\rm Cart}=$`(xx0,xx1,xx2)` set by the `"reference_metric::CoordSystem"` variable. Empirically speaking, it is far easier to write `(x(xx0,xx1,xx2),y(xx0,xx1, xx2),z(xx0,xx1,xx2))` than the inverse, so we will compute the Jacobian matrix
#
# $$
# {\rm Jac\_dUSph\_dDrfmUD[i][j]} = \frac{\partial x^i_{\rm Sph}}{\partial x^j_{\rm Cart}},
# $$
#
# via exact differentiation (courtesy SymPy), and the inverse Jacobian
# $$
# {\rm Jac\_dUrfm\_dDSphUD[i][j]} = \frac{\partial x^i_{\rm Cart}}{\partial x^j_{\rm Sph}},
# $$
#
# using NRPy+'s `generic_matrix_inverter3x3()` function.
#
# In terms of these, the transformation of ADM tensors from Spherical to `"reference_metric::CoordSystem==Cartesian"` coordinates may be written:
#
# \begin{align}
# \gamma^{\rm Cart}_{ij} &=
# \frac{\partial x^\ell_{\rm Cart}}{\partial x^i_{\rm Sph}}
# \frac{\partial x^m_{\rm Cart}}{\partial x^j_{\rm Sph}} \gamma^{\rm Sph}_{\ell m}
# \end{align}
#
# Since $\beta^i=K_{ij}=0$ in this case, and $\alpha$ is not a tensor, only the above Jacobian transformation need be performed:
# +
# Transform initial data to our coordinate system:
# First compute Jacobian and its inverse
drrefmetric__dx_0UDmatrix = sp.Matrix([[sp.diff(rfm.xxSph[0],rfm.xx[0]), sp.diff(rfm.xxSph[0],rfm.xx[1]), sp.diff(rfm.xxSph[0],rfm.xx[2])],
[sp.diff(rfm.xxSph[1],rfm.xx[0]), sp.diff(rfm.xxSph[1],rfm.xx[1]), sp.diff(rfm.xxSph[1],rfm.xx[2])],
[sp.diff(rfm.xxSph[2],rfm.xx[0]), sp.diff(rfm.xxSph[2],rfm.xx[1]), sp.diff(rfm.xxSph[2],rfm.xx[2])]])
dx__drrefmetric_0UDmatrix = drrefmetric__dx_0UDmatrix.inv()
# Declare as gridfunctions the final quantities we will output for the initial data
alpha = gri.register_gridfunctions("EVOL","alpha")
betaU = ixp.register_gridfunctions_for_single_rank1("EVOL","betaU")
gammaDD = ixp.register_gridfunctions_for_single_rank2("EVOL","gammaDD","sym01")
KDD = ixp.register_gridfunctions_for_single_rank2("EVOL","KDD","sym01")
alpha = IDalpha # No Jacobian necessary!
betaU = IDbetaU # Because beta^i = 0
KDD = IDKDD # Because K_{ij} = 0
for i in range(3):
for j in range(3):
# Matrices are stored in row, column format, so (i,j) <-> (row,column)
gammaDD[i][j] = 0
for k in range(3):
for l in range(3):
gammaDD[i][j] += drrefmetric__dx_0UDmatrix[(k,i)]*drrefmetric__dx_0UDmatrix[(l,j)]*IDgammaDD[k][l]
# -={ Spacetime quantities: Generate C code from expressions and output to file }=-
ADMQuantities_to_print = [\
lhrh(lhs=gri.gfaccess("out_gfs","alpha"),rhs=alpha),\
lhrh(lhs=gri.gfaccess("out_gfs","betaU0"),rhs=betaU[0]),\
lhrh(lhs=gri.gfaccess("out_gfs","betaU1"),rhs=betaU[1]),\
lhrh(lhs=gri.gfaccess("out_gfs","betaU2"),rhs=betaU[2]),\
lhrh(lhs=gri.gfaccess("out_gfs","gammaDD00"),rhs=gammaDD[0][0]),\
lhrh(lhs=gri.gfaccess("out_gfs","gammaDD01"),rhs=gammaDD[0][1]),\
lhrh(lhs=gri.gfaccess("out_gfs","gammaDD02"),rhs=gammaDD[0][2]),\
lhrh(lhs=gri.gfaccess("out_gfs","gammaDD11"),rhs=gammaDD[1][1]),\
lhrh(lhs=gri.gfaccess("out_gfs","gammaDD12"),rhs=gammaDD[1][2]),\
lhrh(lhs=gri.gfaccess("out_gfs","gammaDD22"),rhs=gammaDD[2][2]),\
lhrh(lhs=gri.gfaccess("out_gfs","KDD00"),rhs=KDD[0][0]),\
lhrh(lhs=gri.gfaccess("out_gfs","KDD01"),rhs=KDD[0][1]),\
lhrh(lhs=gri.gfaccess("out_gfs","KDD02"),rhs=KDD[0][2]),\
lhrh(lhs=gri.gfaccess("out_gfs","KDD11"),rhs=KDD[1][1]),\
lhrh(lhs=gri.gfaccess("out_gfs","KDD12"),rhs=KDD[1][2]),\
lhrh(lhs=gri.gfaccess("out_gfs","KDD22"),rhs=KDD[2][2])
]
with open(os.path.join(outdir,"ADMQuantities.h"),"w") as file:
ADMQuantities_CcodeKernel = fin.FD_outputC("returnstring",ADMQuantities_to_print,
params="outCverbose=False,includebraces=False,preindent=1")
file.write("""
static inline
void ADMQuantities(const cGH* restrict const cctkGH, const CCTK_INT i0,const CCTK_INT i1,const CCTK_INT i2,
const CCTK_REAL *restrict xx0GF,const CCTK_REAL *restrict xx1GF,const CCTK_REAL *restrict xx2GF,
const CCTK_REAL IDalpha,
const CCTK_REAL IDgammaDD00,const CCTK_REAL IDgammaDD01, const CCTK_REAL IDgammaDD02,
const CCTK_REAL IDgammaDD11,const CCTK_REAL IDgammaDD12, const CCTK_REAL IDgammaDD22,
CCTK_REAL *alphaGF,CCTK_REAL *betaU0GF,CCTK_REAL *betaU1GF,CCTK_REAL *betaU2GF,
CCTK_REAL *gammaDD00GF, CCTK_REAL *gammaDD01GF, CCTK_REAL *gammaDD02GF,
CCTK_REAL *gammaDD11GF, CCTK_REAL *gammaDD12GF, CCTK_REAL *gammaDD22GF,
CCTK_REAL *KDD00GF, CCTK_REAL *KDD01GF, CCTK_REAL *KDD02GF,
CCTK_REAL *KDD11GF, CCTK_REAL *KDD12GF, CCTK_REAL *KDD22GF) {
const CCTK_REAL xx0 = xx0GF[CCTK_GFINDEX3D(cctkGH, i0, i1, i2)];
const CCTK_REAL xx1 = xx1GF[CCTK_GFINDEX3D(cctkGH, i0, i1, i2)];
const CCTK_REAL xx2 = xx2GF[CCTK_GFINDEX3D(cctkGH, i0, i1, i2)];
"""+ADMQuantities_CcodeKernel+"""
}
""")
# -
# <a id='initial_data__convert_to_hydrobase'></a>
#
# ## Step 2.d: Convert TOV solution quantities to `HydroBase` variables $\left\{P,\rho_{\rm baryonic},\epsilon,v_{\rm (n)}^i\right\}$ \[Back to [top](#toc)\]
# $$\label{initial_data__convert_to_hydrobase}$$
#
#
# The TOV solver outputs pressure $P$, the *total* energy density $\rho$, and the baryonic density $\rho_{\rm baryonic}$ as a function of the stellar radius (in isotropic coordinates by default).
#
# Then, the `HydroBase` quantities $\rho^{\rm HB}_{\rm baryonic}$, internal energy $\epsilon^{\rm HB}$, and pressure $P^{\rm HB}$ are given in terms of these variables via
#
# \begin{align}
# P^{\rm HB} &= P; \\
# \rho^{\rm HB}_{\rm baryonic} &= \rho_{\rm baryonic}, \\
# \rho &= \rho_{\rm baryonic} \left(1 + \epsilon_{\rm cold}\right) \\
# \implies \epsilon_{\rm cold} &= \frac{\rho}{\rho_{\rm baryonic}} - 1\\
# \epsilon^{\rm HB} &= \epsilon_{\rm cold}, \\
# \end{align}
# [the NRPy+ piecewise polytrope tutorial notebook](Tutorial-TOV-Piecewise_Polytrope_EOSs.ipynb#rhob_from_pcold). Note that $\rho_{\rm baryonic}$ will be floored to a nonzero atmosphere value, so that computing $\epsilon$ will never involve a division by zero.
#
# The TOV star is motionless, with all spatial components of the 4-velocity $u^i=0$ and (as seen above) zero shift $\beta^i$. Thus the Valencia 3-velocity (i.e., the 3-velocity normal to the spatial slice) $v_{\rm (n)}^i$ is given by
#
# $$
# v_{\rm (n)}^{i,{\rm HB}} = 0
# $$
# +
IDValencia3velocityU = ixp.zerorank1() # Valencia 3-velocity is zero
IDPressure = par.Cparameters("REAL", thismodule, "IDPressure", 1e300) # IDPressure must be set in C
IDrho_baryonic = par.Cparameters("REAL", thismodule, "IDrho_baryonic", 1e300) # IDrho_baryonic must be set in C
IDrho__total_energy_density = par.Cparameters("REAL", thismodule, "IDrho__total_energy_density", 1e300) # IDrho__total_energy_density must be set in C
# Declare as gridfunctions the final quantities we will output for the initial data
Valencia3velocityU = ixp.register_gridfunctions_for_single_rank1("EVOL","Valencia3velocityU")
Pressure, rho_baryonic, epsilon = gri.register_gridfunctions("EVOL",["Pressure", "rho_baryonic", "epsilon"])
Valencia3velocityU = IDValencia3velocityU # Because all components of Valencia3velocityU are *zero*
Pressure = IDPressure
rho_baryonic = IDrho_baryonic
epsilon = IDrho__total_energy_density / IDrho_baryonic - sp.sympify(1)
# -={ Spacetime quantities: Generate C code from expressions and output to file }=-
HydroQuantities_to_print = [\
lhrh(lhs=gri.gfaccess("out_gfs","Pressure"),rhs=Pressure),\
lhrh(lhs=gri.gfaccess("out_gfs","rho_baryonic"),rhs=rho_baryonic),\
lhrh(lhs=gri.gfaccess("out_gfs","epsilon"),rhs=epsilon),\
lhrh(lhs=gri.gfaccess("out_gfs","Valencia3velocityU0"),rhs=Valencia3velocityU[0]),\
lhrh(lhs=gri.gfaccess("out_gfs","Valencia3velocityU1"),rhs=Valencia3velocityU[1]),\
lhrh(lhs=gri.gfaccess("out_gfs","Valencia3velocityU2"),rhs=Valencia3velocityU[2])
]
with open(os.path.join(outdir,"HydroQuantities.h"),"w") as file:
HydroQuantities_CcodeKernel = fin.FD_outputC("returnstring",HydroQuantities_to_print,
params="outCverbose=False,includebraces=False,preindent=2")
file.write("""
static inline
void HydroQuantities(const cGH* restrict const cctkGH, const CCTK_INT i0,const CCTK_INT i1,const CCTK_INT i2,
const CCTK_REAL IDPressure, const CCTK_REAL IDrho_baryonic,
const CCTK_REAL IDrho__total_energy_density,
CCTK_REAL *PressureGF,CCTK_REAL *rho_baryonicGF,
CCTK_REAL *epsilonGF,
CCTK_REAL *Valencia3velocityU0GF,
CCTK_REAL *Valencia3velocityU1GF,
CCTK_REAL *Valencia3velocityU2GF) {
DECLARE_CCTK_PARAMETERS;
if(IDrho__total_energy_density <= 0 || IDrho_baryonic <= 0 || IDPressure <= 0) {
rho_baryonicGF[CCTK_GFINDEX3D(cctkGH, i0, i1, i2)] = rho_atmosphere;
PressureGF[CCTK_GFINDEX3D(cctkGH, i0, i1, i2)] = K_atmosphere*pow(rho_atmosphere,Gamma_atmosphere);
epsilonGF[CCTK_GFINDEX3D(cctkGH, i0, i1, i2)] = 0;
Valencia3velocityU0GF[CCTK_GFINDEX3D(cctkGH, i0, i1, i2)] = 0;
Valencia3velocityU1GF[CCTK_GFINDEX3D(cctkGH, i0, i1, i2)] = 0;
Valencia3velocityU2GF[CCTK_GFINDEX3D(cctkGH, i0, i1, i2)] = 0;
} else {
"""+HydroQuantities_CcodeKernel+"""
// Apply pressure depletion.
PressureGF[CCTK_GFINDEX3D(cctkGH, i0, i1, i2)] *= (1.0 - Pressure_depletion_factor);
}
}
""")
# -
# <a id='einstein'></a>
#
# # Step 3: Interfacing with the Einstein Toolkit \[Back to [top](#toc)\]
# $$\label{einstein}$$
#
# <a id='einstein_c'></a>
#
# ## Step 3.a: Constructing the Einstein Toolkit C-code calling functions that include the C code kernels \[Back to [top](#toc)\]
# $$\label{einstein_c}$$
#
# We will write another C file with the functions we need here.
with open(os.path.join(outdir,"InitialData.c"), "w") as file:
file.write("""
#include <math.h>
#include <stdio.h>
#include <stdbool.h>
#include "cctk.h"
#include "cctk_Parameters.h"
#include "cctk_Arguments.h"
// Declare initial data input struct:
// stores data from initial data solver,
// so they can be put on the numerical grid.
typedef struct __ID_inputs {
CCTK_REAL Rbar;
int Rbar_idx;
int interp_stencil_size;
int numlines_in_file;
CCTK_REAL *r_Schw_arr,*rho_arr,*rho_baryon_arr,*P_arr,*M_arr,*expnu_arr,*exp4phi_arr,*rbar_arr;
} ID_inputs;
#include "ADMQuantities.h"
#include "HydroQuantities.h"
#include "interpolate_TOV_solution_to_point.h"
#include "convert_TOV_spacetime_vars_to_ADM_vars.h"
// Alias for "vel" vector gridfunction:
#define velx (&vel[0*cctk_lsh[0]*cctk_lsh[1]*cctk_lsh[2]])
#define vely (&vel[1*cctk_lsh[0]*cctk_lsh[1]*cctk_lsh[2]])
#define velz (&vel[2*cctk_lsh[0]*cctk_lsh[1]*cctk_lsh[2]])
void read_TOV_input_data_from_file(ID_inputs *TOV_in) {
DECLARE_CCTK_PARAMETERS;
// Step 1: Set up TOV initial data
// Step 1.a: Read TOV initial data from data file
// Open the data file:
char filename[100];
sprintf(filename,"%s",TOV_filename); // TOV_filename is a CCTK_PARAMETER
FILE *in1Dpolytrope = fopen(filename, "r");
if (in1Dpolytrope == NULL) {
fprintf(stderr,"ERROR: could not open file %s\\n",filename);
exit(1);
}
// Count the number of lines in the data file:
int numlines_in_file = count_num_lines_in_file(in1Dpolytrope);
// Allocate space for all data arrays:
CCTK_REAL *r_Schw_arr = (CCTK_REAL *)malloc(sizeof(CCTK_REAL)*numlines_in_file);
CCTK_REAL *rho_arr = (CCTK_REAL *)malloc(sizeof(CCTK_REAL)*numlines_in_file);
CCTK_REAL *rho_baryon_arr = (CCTK_REAL *)malloc(sizeof(CCTK_REAL)*numlines_in_file);
CCTK_REAL *P_arr = (CCTK_REAL *)malloc(sizeof(CCTK_REAL)*numlines_in_file);
CCTK_REAL *M_arr = (CCTK_REAL *)malloc(sizeof(CCTK_REAL)*numlines_in_file);
CCTK_REAL *expnu_arr = (CCTK_REAL *)malloc(sizeof(CCTK_REAL)*numlines_in_file);
CCTK_REAL *exp4phi_arr = (CCTK_REAL *)malloc(sizeof(CCTK_REAL)*numlines_in_file);
CCTK_REAL *rbar_arr = (CCTK_REAL *)malloc(sizeof(CCTK_REAL)*numlines_in_file);
// Read from the data file, filling in arrays.
// read_datafile__set_arrays() may be found in TOV/tov_interp.h
if(read_datafile__set_arrays(in1Dpolytrope, r_Schw_arr,rho_arr,rho_baryon_arr,P_arr,M_arr,expnu_arr,exp4phi_arr,rbar_arr) == 1) {
fprintf(stderr,"ERROR WHEN READING FILE %s!\\n",filename);
exit(1);
}
fclose(in1Dpolytrope);
REAL Rbar = -100;
int Rbar_idx = -100;
for(int i=1;i<numlines_in_file;i++) {
if(rho_arr[i-1]>0 && rho_arr[i]==0) { Rbar = rbar_arr[i-1]; Rbar_idx = i-1; }
}
if(Rbar<0) {
fprintf(stderr,"Error: could not find rbar=Rbar from data file.\\n");
exit(1);
}
TOV_in->Rbar = Rbar;
TOV_in->Rbar_idx = Rbar_idx;
const int interp_stencil_size = 12;
TOV_in->interp_stencil_size = interp_stencil_size;
TOV_in->numlines_in_file = numlines_in_file;
TOV_in->r_Schw_arr = r_Schw_arr;
TOV_in->rho_arr = rho_arr;
TOV_in->rho_baryon_arr = rho_baryon_arr;
TOV_in->P_arr = P_arr;
TOV_in->M_arr = M_arr;
TOV_in->expnu_arr = expnu_arr;
TOV_in->exp4phi_arr = exp4phi_arr;
TOV_in->rbar_arr = rbar_arr;
/* END TOV INPUT ROUTINE */
}
void NRPyPlusTOVID_ET_InitialData(CCTK_ARGUMENTS) {
DECLARE_CCTK_ARGUMENTS;
DECLARE_CCTK_PARAMETERS;
ID_inputs TOV_in;
read_TOV_input_data_from_file(&TOV_in);
#pragma omp parallel for
for(CCTK_INT i2=0;i2<cctk_lsh[2];i2++) for(CCTK_INT i1=0;i1<cctk_lsh[1];i1++) for(CCTK_INT i0=0;i0<cctk_lsh[0];i0++) {
CCTK_INT idx = CCTK_GFINDEX3D(cctkGH,i0,i1,i2);
CCTK_REAL rr = r[idx];
CCTK_REAL th = acos(z[idx]/rr);
CCTK_REAL IDexp_4phi,IDnu,IDPressure,IDrho_baryonic,IDrho__total_energy_density;
interpolate_TOV_solution_to_point(rr, TOV_in, &IDexp_4phi,&IDnu,
&IDPressure,&IDrho_baryonic,&IDrho__total_energy_density);
CCTK_REAL IDalpha,IDgammaDD00,IDgammaDD01,IDgammaDD02,IDgammaDD11,IDgammaDD12,IDgammaDD22;
convert_TOV_spacetime_vars_to_ADM_vars(rr, th, IDexp_4phi,IDnu,
&IDalpha,&IDgammaDD00,&IDgammaDD01,&IDgammaDD02,&IDgammaDD11,&IDgammaDD12,&IDgammaDD22);
HydroQuantities(cctkGH, i0,i1,i2,
IDPressure,IDrho_baryonic,IDrho__total_energy_density,
press,rho,eps,velx,vely,velz);
ADMQuantities(cctkGH,i0,i1,i2,
x,y,z,
IDalpha,IDgammaDD00,IDgammaDD01,IDgammaDD02,IDgammaDD11,IDgammaDD12,IDgammaDD22,
alp,betax,betay,betaz,
gxx,gxy,gxz,gyy,gyz,gzz,
kxx,kxy,kxz,kyy,kyz,kzz);
}
free(TOV_in.r_Schw_arr);
free(TOV_in.rho_arr);
free(TOV_in.rho_baryon_arr);
free(TOV_in.P_arr);
free(TOV_in.M_arr);
free(TOV_in.expnu_arr);
free(TOV_in.exp4phi_arr);
free(TOV_in.rbar_arr);
}
""")
# <a id='einstein_ccl'></a>
#
# ## Step 3.b: CCL files - Define how this module interacts and interfaces with the larger Einstein Toolkit infrastructure \[Back to [top](#toc)\]
# $$\label{einstein_ccl}$$
#
# Writing a module ("thorn") within the Einstein Toolkit requires that three "ccl" files be constructed, all in the root directory of the thorn:
#
# 1. `interface.ccl}`: defines the gridfunction groups needed, and provides keywords denoting what this thorn provides and what it should inherit from other thorns. Specifically, this file governs the interaction between this thorn and others; more information can be found in the [official Einstein Toolkit documentation](http://einsteintoolkit.org/usersguide/UsersGuidech12.html#x17-178000D2.2).
# With "implements", we give our thorn its unique name. By "inheriting" other thorns, we tell the Toolkit that we will rely on variables that exist and are declared "public" within those functions.
# %%writefile $outrootdir/interface.ccl
implements: NRPyPlusTOVID
inherits: admbase grid hydrobase
# 2. `param.ccl`: specifies free parameters within the thorn, enabling them to be set at runtime. It is required to provide allowed ranges and default values for each parameter. More information on this file's syntax can be found in the [official Einstein Toolkit documentation](http://einsteintoolkit.org/usersguide/UsersGuidech12.html#x17-183000D2.3).
# +
# %%writefile $outrootdir/param.ccl
shares: grid
shares: ADMBase
USES CCTK_INT lapse_timelevels
USES CCTK_INT shift_timelevels
USES CCTK_INT metric_timelevels
USES KEYWORD metric_type
EXTENDS KEYWORD initial_data
{
"NRPyPlusTOVID" :: "Initial data from NRPyPlusTOVID solution"
}
EXTENDS KEYWORD initial_lapse
{
"NRPyPlusTOVID" :: "Initial lapse from NRPyPlusTOVID solution"
}
EXTENDS KEYWORD initial_shift
{
"NRPyPlusTOVID" :: "Initial shift from NRPyPlusTOVID solution"
}
EXTENDS KEYWORD initial_dtlapse
{
"NRPyPlusTOVID" :: "Initial dtlapse from NRPyPlusTOVID solution"
}
EXTENDS KEYWORD initial_dtshift
{
"NRPyPlusTOVID" :: "Initial dtshift from NRPyPlusTOVID solution"
}
shares: HydroBase
EXTENDS KEYWORD initial_hydro
{
"NRPyPlusTOVID" :: "Initial GRHD data from NRPyPlusTOVID solution"
}
#["r_in","r_at_max_density","a","M"] A_b, kappa, gamma
restricted:
CCTK_STRING TOV_filename "Which interpolator should I use"
{
".+" :: "Any nonempty string"
} "outputTOVpolytrope.txt"
restricted:
CCTK_REAL rho_atmosphere "Atmosphere baryonic density"
{
0:* :: "Physical values"
-1 :: "forbidden value to make sure it is explicitly set in the parfile"
} -1
restricted:
CCTK_REAL K_atmosphere "Polytropic K to be used with the EOS corresponding to rho_atmosphere"
{
0:* :: "Physical values"
-1 :: "forbidden value to make sure it is explicitly set in the parfile"
} -1
restricted:
CCTK_REAL Gamma_atmosphere "Polytropic Gamma to be used with the EOS corresponding to rho_atmosphere"
{
0:* :: "Physical values"
-1 :: "forbidden value to make sure it is explicitly set in the parfile"
} -1
restricted:
CCTK_REAL Pressure_depletion_factor "Pressure depletion factor = Pdf: P => (1-Pdf)*P"
{
0:* :: "Greater than or equal to zero, where zero is no depletion and default."
} 0.0
# -
# 3. `schedule.ccl`: allocates storage for gridfunctions, defines how the thorn's functions should be scheduled in a broader simulation, and specifies the regions of memory written to or read from gridfunctions. $\text{schedule.ccl}$'s official documentation may be found [here](http://einsteintoolkit.org/usersguide/UsersGuidech12.html#x17-186000D2.4).
#
# We specify here the standardized ETK "scheduling bins" in which we want each of our thorn's functions to run.
# +
# %%writefile $outrootdir/schedule.ccl
STORAGE: ADMBase::metric[metric_timelevels], ADMBase::curv[metric_timelevels], ADMBase::lapse[lapse_timelevels], ADMBase::shift[shift_timelevels]
schedule NRPyPlusTOVID_ET_InitialData IN HydroBase_Initial
{
LANG: C
READS: grid::x(Everywhere)
READS: grid::y(Everywhere)
READS: grid::y(Everywhere)
WRITES: admbase::alp(Everywhere)
WRITES: admbase::betax(Everywhere)
WRITES: admbase::betay(Everywhere)
WRITES: admbase::betaz(Everywhere)
WRITES: admbase::kxx(Everywhere)
WRITES: admbase::kxy(Everywhere)
WRITES: admbase::kxz(Everywhere)
WRITES: admbase::kyy(Everywhere)
WRITES: admbase::kyz(Everywhere)
WRITES: admbase::kzz(Everywhere)
WRITES: admbase::gxx(Everywhere)
WRITES: admbase::gxy(Everywhere)
WRITES: admbase::gxz(Everywhere)
WRITES: admbase::gyy(Everywhere)
WRITES: admbase::gyz(Everywhere)
WRITES: admbase::gzz(Everywhere)
WRITES: hydrobase::vel[0](Everywhere)
WRITES: hydrobase::vel[1](Everywhere)
WRITES: hydrobase::vel[2](Everywhere)
WRITES: hydrobase::rho(Everywhere)
WRITES: hydrobase::eps(Everywhere)
WRITES: hydrobase::press(Everywhere)
} "Set up general relativistic hydrodynamic (GRHD) fields for TOV initial data"
# -
# <a id='einstein_list'></a>
#
# ## Step 3.c: Add the C code to the Einstein Toolkit compilation list \[Back to [top](#toc)\]
# $$\label{einstein_list}$$
#
# We will also need `make.code.defn`, which indicates the list of files that need to be compiled. This thorn only has the one C file to compile.
# %%writefile $outdir/make.code.defn
SRCS = InitialData.c
# <a id='latex_pdf_output'></a>
#
# # Step 4: Output this notebook to $\LaTeX$-formatted PDF file \[Back to [top](#toc)\]
# $$\label{latex_pdf_output}$$
#
# The following code cell converts this Jupyter notebook into a proper, clickable $\LaTeX$-formatted PDF file. After the cell is successfully run, the generated PDF may be found in the root NRPy+ tutorial directory, with filename
# [Tutorial-ETK_thorn-NRPyPlusTOVID.pdf](Tutorial-ETK_thorn-NRPyPlusTOVID.pdf) (Note that clicking on this link may not work; you may need to open the PDF file through another means.)
import cmdline_helper as cmd # NRPy+: Multi-platform Python command-line interface
cmd.output_Jupyter_notebook_to_LaTeXed_PDF("Tutorial-ETK_thorn-NRPyPlusTOVID")
|
Tutorial-ETK_thorn-NRPyPlusTOVID.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Compute the derivative for f(x)=x^2
# +
def get_derivative(func, x):
"""Compute the derivative of `func` at the location `x`."""
h = 0.0001 # step size
return (func(x+h) - func(x)) / h # rise-over-run
def f(x): return x**2 # some test function f(x)=x^2
x = 3 # the location of interest
computed = get_derivative(f, x)
actual = 2*x
computed, actual # = 6.0001, 6 # pretty close if you ask me...
# -
|
Calculus/Derivative.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: SageMath 8.1
# language: ''
# name: sagemath
# ---
# + deletable=true editable=true language="latex"
#
# Recall that the h-polynomial of a polytope P is $h(t) = f_0+f_1(t−1) +· · ·+f_d(t−1)^d$
# where $f_i=$ number of i-dimensional faces of P.
# Compare the coefficients of these polynomials with
# the values $μ(\hat{0},[n]^j)$ of the Mobius functions of $\Pi^w_{\Gamma_n}$
#
# + deletable=true editable=true
# N=5
# P= graphs.PathGraph(n)
# adj= P.adjacency_matrix()
# sampmat=adj
# subsets=Subsets(list(range(1,N+1))).list()
# print(subsets)
# subsets.remove(subsets[0])
# subsetn=subsets[:]
# marked=[]
# for i,sub in enumerate(subsetn):
# testblock=list(sub)
# testblock[:]=[x-1 for x in testblock]
# testadj=sampmat[testblock,testblock]
# if disconnected(testadj,len(testblock)):
# marked.append(sub)
# else:
# continue
# for i in marked:
# subsetn.remove(i)
# subsetn
# graphs.StarGraph(4).adjacency_matrix()
subsets=Subsets(list(range(1,4))).list()
list(range(5))
# + deletable=true editable=true
def path(N):
return graphs.PathGraph(N).adjacency_matrix()
def star(N):
if N<4:
return graphs.PathGraph(N).adjacency_matrix()
else:
return graphs.StarGraph(N-1).adjacency_matrix()
def disconnected(AdjMat,b): #b is the size of of the testblock
ans= sum(sum(AdjMat))/2
return ans!=b-1
def tubes(n,AdjMat):
subsets=Subsets(list(range(n))).list()
subsets.remove(subsets[0])
subsetn=subsets[:]
marked=[]
for i,sub in enumerate(subsetn):
testblock=list(sub)
testadj=AdjMat[testblock,testblock]
if disconnected(testadj,len(testblock)):
marked.append(sub)
else:
continue
for i in marked:
subsetn.remove(i)
return subsetn[:-1]
def IDG(arg,Adjmat): #short for Induces Disconnected Graph
testblock=list(arg)
testblock[:]=[x-1 for x in testblock]
testadj=Adjmat[testblock,testblock]
if disconnected(testadj,len(testblock)):
return True
else:
return False
def containment(a,b):
if len(a) != len(b)-1:
return False
if not (a).issubset(b):
return False
return True
# P= graphs.PathGraph(N)
# S=graphs.StarGraph(N-1) #the Sage's Star Graph actually graphs N+1 vertices instead of N so I put N-1 here to account for that
# pathadj= P.adjacency_matrix() #default ordering of rows and columns is from 0 to N
# staradj= S.adjacency_matrix()
# n (tubes+1) Path graph data
# 3 5
# 4 9
# 5 14
# 6 20
# 7 27
# 8 35
# 9 44
#possible OEIS sequences
## Sum of the integers from 1 to n, excluding the perfect sixth powers.
## a(n) = n*(n+3)/2.
# n (tubes+1) star graph data
# 3 5
# 4 10
# 5 14
# 6 20
# 7 27
# 8 35
# 9 44
##########################
##this influences the REST of the program so choose wisely
N=5
##########################
P= graphs.PathGraph(N)
S=graphs.StarGraph(N-1) #the Sage's Star Graph actually graphs N+1 vertices instead of N so I put N-1 here to account for that
pathadj= P.adjacency_matrix() #default ordering of rows and columns is from 0 to N
staradj= S.adjacency_matrix()
if N<4:
staradj=pathadj
# print(len(tubes(N,staradj)))
# print(len(tubes(N,pathadj)))
# + deletable=true editable=true
# %%time
pathtubesdata=[]
startubesdata=[]
for i in range(3,13):
pathtubesdata.append(len(tubes(i,path(i))))
startubesdata.append(len(tubes(i,star(i))))
print("n=3 to 12",pathtubesdata) ##a(n) = n-1*(n+2)/2 by oeis
print("n=3 to 12",startubesdata) ##a(n) = 2^(n-1) + n - 2. (row sums of triangles)?
print(tubes(6,path(6)))
# ('n=3 to 12', [5, 9, 14, 20, 27, 35, 44, 54, 65, 77])
# ('n=3 to 12', [5, 10, 19, 36, 69, 134, 263, 520, 1033, 2058])
# CPU times: user 15.1 s, sys: 1.05 s, total: 16.1 s
# Wall time: 19.9 s
# + deletable=true editable=true
#reminding myself of the set operations in python
# print {1,2,3}|({2,3,4}) #union
# print{1,2,3}&({2,3,4}) #intersection
# print{2,3,4}<=({2,3,4}) #subset
# print{2,3,4}>={2,3,4}
# print {2,3,4}<{2,3,4} #proper subset
#set operations in sage are a little different
# print Set([1,2,3])|Set([2,3,4])
## this only returns the tubings of length 2...
## recursion for pairwise comparison as the tubings get larger?
def tubings(n,Adjmat):
tubings=[]
marked=[]
T = tubes(n,Adjmat)
print("|T|= %d" %len(T))
set_of_pairs = [(T[p1], T[p2]) for p1 in range(len(T)) for p2 in range(p1+1,len(T))]
print("number of pairs of T = %d" %len(set_of_pairs)) #equal to len(T) choose 2 as you would expect
for pair in set_of_pairs:
if len(pair[0])==n or len(pair[1])==n:
next
# print pair,(pair[0]).intersection(pair[1]).is_empty(),len((pair[0])&(pair[1]))==len(pair[0]),(pair[1])&(pair[0]) #best way to debug at the moment is to print pair,condition
elif ((pair[0]).intersection(pair[1]).is_empty()==False) and (len(pair[0]&pair[1])==len(pair[0]) or len(pair[1]&pair[0])==len(pair[1])):
tubings.append(list(pair))
elif (pair[0]).intersection(pair[1]).is_empty() and IDG(pair[0]|pair[1],Adjmat)==True: ##IDG== True means its not a tube
tubings.append(list(pair))
else:
continue
#insert recursion here?
return (tubings)
adj1=pathadj
adj2=staradj
tubs=tubings(N,adj1)
tubs2=tubings(N,adj2)
def intersect(lst1, lst2):
# Use of hybrid method
lst3 = [value for value in lst1 if value in lst2]
# for elm in lst1:
# if elm&
return lst3
#print('experiment',intersect([{1},{3,4}],[{1},{1,2,3,4}]))
#do not count maximum tubes
# + deletable=true editable=true
def tube(n,AdjMat):
subsets=Subsets(list(range(n))).list()
subsets.remove(subsets[0])
subsetn=subsets[:]
marked=[]
for i,sub in enumerate(subsetn):
testblock=list(sub)
testadj=AdjMat[testblock,testblock]
if disconnected(testadj,len(testblock)):
marked.append(sub)
else:
continue
for i in marked:
subsetn.remove(i)
return subsetn[:-1]
len(tubes(5,path(5)))
# + deletable=true editable=true
# %%time
def C(n,r):
if r>n:
return 0
else:
return (factorial(n))/(factorial(r)*factorial(n-r))
def h(n):
from math import factorial
hpoly=[0]*(n+1)
for i in range(1,n+1):
hpoly[i]=C(n,i)*C(n,i-1)/(n)
#try // instead?
return hpoly[1:]
h(4)
def ht(n):
from math import factorial
hpoly=[0]*(n)
for i in range(n):
hpoly[i]=C(n,i+1)*C(n,i)/(n)
#try // instead?
return hpoly
print(ht(4),h(4))
# + deletable=true editable=true
##example of how to write the cover relation for a poset
elms = SetPartitions(4)
def fcn(A, B):
if len(A) != len(B)+1:
return False
for a in A:
if not any(set(a).issubset(b) for b in B):
return False
return True
P=Poset((elms, fcn))#, cover_relations=True)
index1=P[0]
index2=P[len(P)-1]
mobfunc=P.moebius_function(index1, index2)
print("mobius func on {} , {} ={}".format(index1,index2,mobfunc))
P.plot()
#(polytopes.simplex().plot())
# + deletable=true editable=true
##TODO
##Fix tubings fnc to include more than just 2 dimensions in the tubes
##Organize into poset
## plot polytope?
tubes(N,pathadj)
tubings(N,pathadj)
|
H_Polynomials-Copy1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.mlab as mlab
from scipy.stats import norm
plt.style.use('fivethirtyeight')
traffic = pd.read_csv('train_revised_11.09.2018/train_revised.csv')
traffic.columns
t = traffic[['ride_id', 'travel_date', 'travel_time','travel_from', 'car_type', 'max_capacity']]
t.info()
t.dtypes
t.describe(include='all')
t.head()
# Here I wanted to have a look at the fields that had 2 unique values.
print(t['car_type'].value_counts())
print(t['max_capacity'].value_counts())
# So if you look at the describe cell above you'll find something curious with the travel times. Despite the fact that there over 6249 trips, and there's only 78 unquie times, graphed below. Looking at the values, majority of the first 10 are in the morning at 7, tells me that I would make sense to simplify this field to an hour field instead.
t['travel_time'].value_counts().plot.bar()
t['travel_time'].value_counts()
# Creating the new variable. I'm just going to call it hour_booked
t['hour_booked'] = pd.to_numeric(t['travel_time'].str.extract(r'(^\d*)').loc[:,0])
t.head()
t['hour_booked'].value_counts().sort_index().plot.bar()
((t['hour_booked'].value_counts()/t['hour_booked'].value_counts().sum())*100).sort_index()
# Wow that's most people booking tickets at 7 am. Over 55%. I'm thinking that I might change this into a categorical variable when I do my models.
#
# Next I think we can look at dates. We have 149 days, let's make that field more configurable
t.head()
t['travel_date'] = pd.to_datetime(t['travel_date'])
# So first I looked at the year assuming they came from same year but that would have been a mistakes at the data is from over 2017, 2018. Also the busiest months are the end of the year and the beginning. Added variable for month/year for further exploration.
t['m_y'] = t['travel_date'].dt.strftime('%Y-%m')
t['day'] = t['travel_date'].dt.strftime('%w')
t['m_y'].value_counts().sort_index().plot.bar()
# So it seems that for a lot of months in 2017, bookings were made on single days. That might be a problem. Given that the test data for this is for 2018. We'll have to keep an eye on whether removing those months makes sense. I'd say some of this is an articially made dataset.
for x in t['m_y'].value_counts().sort_index().index:
print('For month: {}'.format(x))
print(t[t['m_y']==x]['day'].value_counts().shape)
t.head()
# Let's seperate the dataset based on the car type
b = t[t['car_type']=='Bus']
s = t[t['car_type']=='shuttle']
# So let's see how often buses travel without filling.
(b['ride_id'].value_counts().value_counts().sort_index(ascending=False)/b['ride_id'].value_counts().value_counts().sort_index(ascending=False).sum()).plot.bar()
(s['ride_id'].value_counts().value_counts().sort_index(ascending=False)/s['ride_id'].value_counts().value_counts().sort_index(ascending=False).sum()).plot.bar()
# Looks like a lot of the buses are leaving with only a few passengers. This tells me that they're not only using this platform to book people. Either that or this dataset is not complete or artificially made. Anyway lemme continue exploring the hypothesis I was looking at before this happened. What times is a driver likely to leave without filling his ride.
b.head()
bpf = (b['ride_id'].value_counts()/49).round(2)
spf = (s['ride_id'].value_counts()/11).round(2)
for x in bpf.index:
b.loc[b['ride_id'].isin([x]), 'p_filled'] = bpf[x]
b.head()
for x in spf.index:
s.loc[s['ride_id'].isin([x]), 'p_filled'] = spf[x]
s.head()
s['hour_booked'].value_counts().sort_index().index
s[s['hour_booked']==5]['p_filled'].value_counts().sort_index().plot.bar()
s[s['hour_booked']==6]['p_filled'].value_counts().sort_index().plot.bar()
s[s['hour_booked']==7]['p_filled'].value_counts().sort_index().plot.bar()
s[s['hour_booked']==8]['p_filled'].value_counts().sort_index().plot.bar()
s[s['hour_booked']==9]['p_filled'].value_counts().sort_index().plot.bar()
s[s['hour_booked']==10]['p_filled'].value_counts().sort_index().plot.bar()
s[s['hour_booked']==11]['p_filled'].value_counts().sort_index().plot.bar()
(s['p_filled'].value_counts().sort_index()/s['p_filled'].value_counts().sum()).plot.bar()
# So from these plots you can see that shuttle drivers are more likely to leave the closer their rides get to filled(exponentially), this could be used to come up with a real time prediction system one day. Let's look at the larger buses
n = s[['hour_booked','p_filled']]
n
for x in s['hour_booked'].value_counts().sort_index().index:
print(s[s['hour_booked']==x]['p_filled'].mean())
b['hour_booked'].value_counts().sort_index().index
b[b['hour_booked']==5]['p_filled'].value_counts().sort_index().plot.bar()
b[b['hour_booked']==6]['p_filled'].value_counts().sort_index().plot.bar()
b[b['hour_booked']==7]['p_filled'].value_counts().sort_index().plot.bar()
b[b['hour_booked']==8]['p_filled'].value_counts().sort_index().plot.bar()
(b[b['hour_booked']==19]['p_filled'].value_counts().sort_index()/b[b['hour_booked']==19]['p_filled'].value_counts().sum()).plot.bar()
b[b['hour_booked']==23]['p_filled'].value_counts().sort_index().plot.bar()
# It seems that bus drivers are less concern about filling their rides before they leave
# The first time I went through this data set I spent so much time looking at the seats only to realise that the test set did not have any data on that. Also it doesn't have payment method either. Sigh, that's a whole morning gone. Always remember to look at the client brief people.
|
EDA.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.6 64-bit (''azureml-customerchurn'': conda)'
# name: python37664bitazuremlcustomerchurnconda333f9d2e7dd246df8e2a65ddfa476e59
# ---
import pandas as pd
from pandas_profiling import ProfileReport
df_customerchurn = pd.read_csv('../data/raw/customer-churn.csv', na_values=[' '])
df_customerchurn.info()
ProfileReport(df_customerchurn).to_file('../reports/customer-churn.html')
|
notebooks/explore-data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python37
# language: python
# name: python37
# ---
# +
from sentence_transformers import SentenceTransformer
from datetime import datetime
import logging
import re
import numpy as np
import pandas as pd
import hdbscan
import umap
import matplotlib.pyplot as plt
# -
import warnings
warnings.filterwarnings(action='ignore')
model_path = "kpfsbert-base" # sbert
cluster_mode = 'title'
# Data Loading to clustering
DATA_PATH = 'data/newstrust/newstrust_20210601_samlple.json'
df = pd.read_json(DATA_PATH, encoding='utf-8')
len(df)
#kpfSBERT 모델 로딩
model = SentenceTransformer(model_path)
# UMAP 차원축소 실행
def umap_process(corpus_embeddings, n_components=5):
umap_embeddings = umap.UMAP(n_neighbors=15,
n_components=n_components,
metric='cosine').fit_transform(corpus_embeddings)
return umap_embeddings
# HDBSCAN 실행
def hdbscan_process(corpus, corpus_embeddings, min_cluster_size=15, min_samples=10, umap=True, n_components=5, method='eom'):
if umap:
umap_embeddings = umap_process(corpus_embeddings, n_components)
else:
umap_embeddings = corpus_embeddings
cluster = hdbscan.HDBSCAN(min_cluster_size=min_cluster_size,
min_samples=10,
allow_single_cluster=True,
metric='euclidean',
core_dist_n_jobs=1,# knn_data = Parallel(n_jobs=self.n_jobs, max_nbytes=None) in joblib
cluster_selection_method=method).fit(umap_embeddings) #eom leaf
docs_df = pd.DataFrame(corpus, columns=["Doc"])
docs_df['Topic'] = cluster.labels_
docs_df['Doc_ID'] = range(len(docs_df))
docs_per_topic = docs_df.groupby(['Topic'], as_index = False).agg({'Doc': ' '.join})
return docs_df, docs_per_topic
# +
# 카테고리별 클러스터링
start = datetime.now()
print('작업 시작시간 : ', start)
previous = start
bt_prev = start
tot_df = pd.DataFrame()
print(' processing start... with cluster_mode :', cluster_mode)
category = df.category.unique()
df_category = []
for categ in category:
df_category.append(df[df.category==categ])
cnt = 0
rslt = []
topics = []
#순환하며 데이터 만들어 df에 고쳐보자
for idx, dt in enumerate(df_category):
corpus = dt[cluster_mode].values.tolist()
# '[보통공통된꼭지제목]' 형태를 제거해서 클러스터링시 품질을 높인다.
for i, cp in enumerate(corpus):
corpus[i] = re.sub(r'\[(.*?)\]', '', cp)
# print(corpus[:10])
corpus_embeddings = model.encode(corpus, show_progress_bar=True)
docs_df, docs_per_topic = hdbscan_process(corpus, corpus_embeddings,
umap=False, n_components=15, #연산량 줄이기 위해 umap 사용시 True
method='leaf',
min_cluster_size=5,
min_samples=30,
)
cnt += len(docs_df)
rslt.append(docs_df)
topics.append(docs_per_topic)
dt['cluster'] = docs_df['Topic'].values.tolist()
tot_df = pd.concat([tot_df,dt])
bt = datetime.now()
print(len(docs_df), 'docs,', len(docs_per_topic)-1 ,'clusters in', category[idx], ', 소요시간 :', bt - bt_prev)
bt_prev = bt
now = datetime.now()
print(' Total docs :', cnt,'in', len(rslt), 'Categories', ', 소요시간 :', now - previous)
previous = now
#cluster update
df['cluster'] = tot_df['cluster'].astype(str)
end = datetime.now()
print('작업 종료시간 : ', end, ', 총 소요시간 :', end - start)
# -
categ = '사회'
condition = (df.category == categ) & (df.cluster == '2') # 조건식 작성
test = df[condition]
print(len(test))
test.title.values.tolist()
# +
# 클러스터별 주제어 추출확인
from sklearn.feature_extraction.text import CountVectorizer
def c_tf_idf(documents, m, ngram_range=(1, 1)):
count = CountVectorizer(ngram_range=ngram_range, stop_words="english").fit(documents)
t = count.transform(documents).toarray()
w = t.sum(axis=1)
tf = np.divide(t.T, w)
sum_t = t.sum(axis=0)
idf = np.log(np.divide(m, sum_t)).reshape(-1, 1)
tf_idf = np.multiply(tf, idf)
return tf_idf, count
def extract_top_n_words_per_topic(tf_idf, count, docs_per_topic, n=20):
words = count.get_feature_names_out()
labels = list(docs_per_topic.Topic)
tf_idf_transposed = tf_idf.T
indices = tf_idf_transposed.argsort()[:, -n:]
top_n_words = {label: [(words[j], tf_idf_transposed[i][j]) for j in indices[i]][::-1] for i, label in enumerate(labels)}
return top_n_words
def extract_topic_sizes(df):
topic_sizes = (df.groupby(['Topic'])
.Doc
.count()
.reset_index()
.rename({"Topic": "Topic", "Doc": "Size"}, axis='columns')
.sort_values("Size", ascending=False))
return topic_sizes
# -
# 카테고리별 클러스터별 주제어 추출확인
category_id = 3
# +
tf_idf, count = c_tf_idf(topics[category_id].Doc.values, m=len(corpus))
top_n_words = extract_top_n_words_per_topic(tf_idf, count, topics[category_id], n=20)
topic_sizes = extract_topic_sizes(rslt[category_id]); topic_sizes.head(10)
# -
len(topic_sizes)
top_n_words[2][:10]
|
kpfSBERT_clustering.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Box Chart
import matplotlib.pyplot as plt
import numpy
data = numpy.random.randint(0, 1000, 6)
# +
plt.title("Box Chart")
plt.ylabel("Random Values")
plt.boxplot(data)
|
data-visualization/charts/box chart.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Import required libraries
import pandas as pd
from sklearn.datasets import load_iris
from factor_analyzer import FactorAnalyzer
import matplotlib.pyplot as plt
import numpy as np
import seaborn as sns
from sklearn.cluster import SpectralClustering
from sklearn.cluster import KMeans
# set the address of the file on your disk
# this table has factors (output from the factor analysis code)
pathname = (r"C:\MyFiles\Dropbox\Teaching\Advanced_GIS_for_Environmental_Planning\Data\NYC\CensusTracts\Census_Tracts_with_Data_two_factor_brooklyn3.csv")
df = pd.read_csv(pathname)
df.sample(2)
# lets' get the factors
X=df[['factor_1','factor_2','factor 3','factor 4']]
# some scatter plots
sns.displot(data=df,x ='factor_1',y ='factor 3',alpha=0.9)
# creating the cluster object
k_means = KMeans(n_clusters=3)
k_means.fit(X)
# getting the cluster lables and put them in one column
df['cluster'] = k_means.labels_
len(k_means.labels_)
df
# plotting the factors with clusters
sns.displot(data=df,x ='factor_1',y ='factor_2',hue='cluster',alpha=0.9)
# more plots to show the clusters
sns.jointplot(data=df,x ='factor_1',y ='factor 4',hue='cluster',alpha=0.9,palette=sns.color_palette("hls", 3))
# more clustering from SpectralClustering algorithm
model = SpectralClustering(n_clusters=3, affinity='nearest_neighbors',
assign_labels='kmeans')
labels = model.fit_predict(X)
# create a column to hod the labels
df['cluster_new']=labels
sns.jointplot(data=df,x ='factor_1',y ='factor_2',hue='cluster_new',alpha=0.9,palette=sns.color_palette("hls", 3))
# saving the clusters in a table
df.to_csv(r"C:\MyFiles\Dropbox\Teaching\Advanced_GIS_for_Environmental_Planning\Data\NYC\CensusTracts\Census_Tracts_with_Data_two_factor_brooklyn3_withClusters3.csv")
|
K-means Clustering.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Nikhila-padmanabhan/Machine-Learning-Concepts/blob/master/Digitdataset.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="XwNcYBERVT0i" colab_type="code" colab={}
from sklearn.datasets import load_digits
# + id="9cw0Ql9yVW6l" colab_type="code" colab={}
Data = load_digits()
# + id="8gMeqacXVc_l" colab_type="code" colab={}
Data.data
# + id="9eo-0lavVg1B" colab_type="code" colab={}
Data.images
# + id="mVOY4S2nVrdy" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="8cc93d02-1bb4-4ac0-938d-82dd475e70b4"
Data.target
# + id="-opf6eQ-VwcE" colab_type="code" colab={}
print(Data.DESCR)
# + id="LAspZ6oZV1tl" colab_type="code" colab={}
# + id="PCz55pxzVyLU" colab_type="code" colab={}
|
Digitdataset.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import linearsolve as ls
import matplotlib.pyplot as plt
plt.style.use('classic')
# %matplotlib inline
# # Discussion: Week 7
#
#
# ## Exercise: The Baseline RBC Model without Labor
#
# The equilibrium conditions for the RBC model without labor are:
#
# \begin{align}
# \frac{1}{C_t} & = \beta E_t \left[\frac{\alpha A_{t+1}K_{t+1}^{\alpha-1} +1-\delta }{C_{t+1}}\right]\\
# K_{t+1} & = I_t + (1-\delta) K_t\\
# Y_t & = A_t K_t^{\alpha}\\
# Y_t & = C_t + I_t\\
# \log A_{t+1} & = \rho \log A_t + \epsilon_{t+1}
# \end{align}
#
# where $\epsilon_{t+1} \sim \mathcal{N}(0,\sigma^2)$.
#
# The objective is use `linearsolve` to create a stochastic simulation of the RBC model. USe the following parameter values for the simulation:
#
# | $$\rho$$ | $$\sigma$$ | $$\beta$$ | $$\alpha$$ | $$\delta $$ |
# |----------|------------|-----------|------------|-------------|
# | 0.75 | 0.006 | 0.99 | 0.35 | 0.025 |
#
#
# ### Model Preparation
#
# Before proceding, let's recast the model in the form required for `linearsolve`. Write the model with all variables moved to the lefthand side of the equations and dropping the expecations operator $E_t$ and the exogenous shock $\epsilon_{t+1}$:
#
# \begin{align}
# 0 & = \beta\left[\frac{\alpha A_{t+1}K_{t+1}^{\alpha-1} +1-\delta }{C_{t+1}}\right] - \frac{1}{C_t}\\
# 0 & = A_t K_t^{\alpha} - Y_t\\
# 0 & = I_t + (1-\delta) K_t - K_{t+1}\\
# 0 & = C_t + I_t - Y_t\\
# 0 & = \rho \log A_t - \log A_{t+1}
# \end{align}
#
# Remember, capital and TFP are called *state variables* because they're $t+1$ values are predetermined. Output, consumption, and investment are called a *costate* or *control* variables. Note that the model as 5 equations in 5 endogenous variables.
#
#
# ### Initialization, Approximation, and Solution
#
# The next several cells initialize the model in `linearsolve` and then approximate and solve it.
# +
# Create a variable called 'parameters' that stores the model parameter values in a Pandas Series. CELL PROVIDED
parameters = pd.Series(dtype=float)
parameters['rho'] = .75
parameters['sigma_squared'] = 0.006**2
parameters['beta'] = 0.99
parameters['alpha'] = 0.35
parameters['delta'] = 0.025
# Create variable called 'var_names' that stores the variable names in a list with state variables ordered first
var_names = ['a','k','y','c','i']
# Create variable called 'shock_names' that stores an exogenous shock name for each state variable.
shock_names = ['e_a','e_k']
# Define a function that evaluates the equilibrium conditions of the model solved for zero.
def equilibrium_equations(variables_forward,variables_current,parameters):
# Parameters. PROVIDED
p = parameters
# Current variables. PROVIDED
cur = variables_current
# Forward variables. PROVIDED
fwd = variables_forward
# Euler equation
euler_equation = p.beta*(p.alpha*fwd.a*fwd.k**(p.alpha-1)+1-p.delta)/fwd.c - 1/cur.c
# Production function
production_function = cur.a*cur.k**p.alpha - cur.y
# Capital evolution
capital_evolution = cur.i + (1 - p.delta)*cur.k - fwd.k
# Market clearing
market_clearing = cur.c+cur.i - cur.y
# Exogenous tfp
tfp_process = p.rho*np.log(cur.a) - np.log(fwd.a)
# Stack equilibrium conditions into a numpy array
return np.array([
euler_equation,
production_function,
capital_evolution,
market_clearing,
tfp_process
])
# Initialize the model into a variable named 'rbc_model'
rbc_model = ls.model(equations = equilibrium_equations,
n_states=2,
var_names=var_names,
shock_names=shock_names,
parameters=parameters)
# Compute the steady state numerically using .compute_ss() method of rbc_model
guess = [1,4,1,1,1]
rbc_model.compute_ss(guess)
# Find the log-linear approximation around the non-stochastic steady state and solve using .approximate_and_solve() method of rbc_model
rbc_model.approximate_and_solve()
# -
# ### Stochastic Simulation
#
# Compute a 201 period stochastic simulation of the model. Set the seed for the simulation to 126.
#
# Recall that `linearsolve` handles the model *as if* there is an exogenous shock for every state variable. In this case, that means two exogenous shocks $\epsilon_t^a$ and $\epsilon^k_t$. For the stochastic simulation, we need to specify the *covariance matrix* for the two shocks. The covariance matrix has the variance of each shock in the diagonal elements and covariances on the off-diagonal elements. Since there is no shock to capital in our model,
#
# \begin{align}
# \text{Covariance matrix} & = \left[\begin{array}{cc}\sigma^2 & 0\\ 0 & 0\end{array}\right]
# \end{align}
# Compute the stochastic simulation using the .stoch_sim() method of rbc_model
rbc_model.stoch_sim(T=201,seed=126,cov_mat = [[parameters['sigma_squared'],0],[0,0]])
# Print the first 5 rows of rbc_model.simulated
print(rbc_model.simulated.head())
# Print the first 5 rows of simulated output
print(rbc_model.simulated['y'].head())
# ## Exercise: Analyze Simulation Results
#
# On a single axis, plot the simulated values for output, consumption and investment.
# Plot simulated results
fig = plt.figure(figsize=(12,4))
ax = fig.add_subplot(1,1,1)
rbc_model.simulated[['y','c','i']].plot(ax=ax,lw=3,grid=True,alpha=0.75)
ax.legend(loc='center left', bbox_to_anchor=(1, 0.5))
# **Questions**
#
# 1. What is the exogenous *cause* of the fluctuations in output, consumption and investment in your simulation?
# 2. Looking at the plot of your simulated data, which quantity — output, consumption or investment — has the smallest fluctuations from the steady state? What feature of the model leads to this?
# 3. Which quantity — output, consumption or investment — has the largest fluctuations from the steady state? What feature of the model leads to this?
# **Answers**
#
# 1. Exogenous shocks to TFP. That is, exogenous shocks to the economy's production function.<!-- answer -->
# 2. Consumption. The household's Euler equation implies that the household wants to *smooth* consumption over time.<!-- answer -->
# 3. Investment. Again, the Euler equation. The household uses investment fluctuations to smooth consumption. When output is high, the household raises investment in order to accumulate capital and save some of the high production for later. Similarly, when output is low, the household reduces investment in order to deplete capital and therefore borrow against future output.<!-- answer -->
|
Discussion Notebooks/Econ126_Discussion_Week_07.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Peakfit class example
# Fitting is based on the LMfit library: https://lmfit.github.io/lmfit-py/index.html
# +
from nasagamma import spectrum as sp
import numpy as np
import pandas as pd
from nasagamma import peaksearch as ps
from nasagamma import peakfit as pf
# dataset 1
file = "data/SSR-mcnp.hdf"
df = pd.read_hdf(file, key='data')
df = df.iloc[1:,:]
cts_np = df.cts.to_numpy() * 1e8
erg = np.array(df.index)
# Required input parameters (in channels)
fwhm_at_0 = 1.0
ref_fwhm = 31
ref_x = 1220
min_snr = 1
# -
# instantiate a Spectrum and a peaearch object
spect = sp.Spectrum(counts=cts_np, energies=erg)
search = ps.PeakSearch(spect, ref_x, ref_fwhm, fwhm_at_0, min_snr=min_snr)
# We can now define a background type: linear, quadratic, exponential, or any other n-degree polynomial.
# We also define a range of x-values to perform the fit
# peakfit class
bkg0 = 'poly2'
xrange = [2.6, 4]
fit = pf.PeakFit(search, xrange, bkg=bkg0)
fit.plot(plot_type="full", legend='on')
# One can try changing the x-range and background type and see how things change.
|
examples/3.peakfitting_example.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
os.chdir('../..')
# +
import pandas as pd
import numpy as np
from datetime import datetime, date
import matplotlib.pyplot as plt
import json
data = 'data/'
# -
atms = pd.read_csv(data + 'atms.small.csv')
clients = pd.read_csv(data + 'clients.small.csv')
companies = pd.read_csv(data + 'companies.small.csv')
transactions = pd.read_csv(data + 'transactions.small.csv')
months = {}
days = {}
counts = 0
dates = []
for i in transactions.index:
date = transactions.loc[i]['date']
month = int(date.split('/')[1])
day = int(date.split('/')[0])
if month not in months.keys():
months[month] = 0
months[month] += 1
if day not in days.keys():
days[day] = 0
days[day] += 1
if day > 12 and month > 12:
counts += 1
counts/len(transactions)
months
days
# ## Extract time and flow patterns
threshold_minutes = 15
threshold_amount = 0.1
# +
groups = {'source': {'time': [], 'flow': []},
'target': {'time': [], 'flow': []}}
ids = {'source': transactions['source'].unique(), 'target': transactions['target'].unique()}
for key in ids.keys():
print('Check for {}'.format(key))
for id_ in ids[key]:
df = transactions[transactions[key] == id_]
df.index = range(len(df))
times = np.zeros((len(df), len(df)), dtype=np.bool)
amounts = np.zeros((len(df), len(df)), dtype=np.bool)
for j in range(len(df)):
for k in range(j, len(df)):
# Time diff
tj = datetime.strptime(df.loc[j]['time'], '%H:%M:%S').time()
tk = datetime.strptime(df.loc[k]['time'], '%H:%M:%S').time()
diff = np.abs(((datetime.combine(date.today(), tj) - datetime.combine(date.today(), tk)).total_seconds()))
if diff < threshold_minutes*60:
times[j][k] = True
times[k][j] = True
else:
times[j][k] = False
times[k][j] = False
# Amount diff
amj = df.loc[j]['amount']
amk = df.loc[k]['amount']
diff = np.abs(amj-amk)
avg = np.mean([amj, amk])
if diff < threshold_amount*avg:
amounts[j][k] = True
amounts[k][j] = True
else:
amounts[j][k] = False
amounts[k][j] = False
mat = amounts*times
j=0
while j <= len(df)-1:
arr = [i for i, x in enumerate(mat[j, :]) if x]
j = arr[-1]+1
if len(arr) > 2:
linked = [df.loc[k]['id'] for k in arr]
if len(df[df['id'].isin(linked)]['date'].unique()) == 1:
groups[key]['flow'].append(linked)
else:
groups[key]['time'].append(linked)
# -
for key in groups.keys():
print('{}:'.format(key))
for type_ in groups[key].keys():
print(' {}: {}'.format(type_, len(groups[key][type_])))
# # Check for similar transactions
for key in groups.keys():
for type_ in groups[key].keys():
ids = []
dup = []
for i, g in enumerate(groups[key][type_]):
df = transactions[transactions['id'].isin(g)]
id_ = df[key].iloc[0]
if id_ in ids:
dup.append([ids.index(id_), i])
ids.append(id_)
merge = []
for pair in dup:
amount1 = np.mean(transactions[transactions['id'].isin(groups[key][type_][pair[0]])]['amount'])
amount2 = np.mean(transactions[transactions['id'].isin(groups[key][type_][pair[1]])]['amount'])
delta = np.abs(amount1-amount2)
if delta < threshold_amount*np.mean([amount1, amount2]):
merge.append(pair)
to_del = []
for pair in merge:
groups[key][type_][pair[0]] = list(np.append(groups[key][type_][pair[0]], groups[key][type_][pair[1]]))
to_del.append(pair[1])
for index in sorted(to_del, reverse=True):
del groups[key][type_][index]
for key in groups.keys():
print('{}:'.format(key))
for type_ in groups[key].keys():
print(' {}: {}'.format(type_, len(groups[key][type_])))
with open(data + 'groups.json', 'w') as outfile:
json.dump(groups, outfile)
# # Extract JSON for d3js
# +
with open(data + 'groups.json', 'r') as outfile:
groups = json.load(outfile)
source_ids = []
for g in groups['source']['time']:
susp_node = transactions[transactions['id'].isin(g)]['source'].iloc[0]
source_ids.append(susp_node)
target_ids = []
for g in groups['target']['time']:
susp_node = transactions[transactions['id'].isin(g)]['source'].iloc[0]
target_ids.append(susp_node)
# -
transactions[transactions['id'].isin(groups['source']['flow'][0])]
transactions[transactions['id'].isin(groups['target']['flow'][0])]
# +
# group 0 = suspect
# group 1 = accomplice
# group 2 = accomplice edge
# group 3 = normal
# group 4 = normal edge
# -
for id_ in range(3):
data = {'nodes': [], 'links': [], 'type': None}
susp_node = transactions[transactions['id'].isin(groups['source']['time'][id_])]['source'].iloc[0]
data['nodes'].append({'id': susp_node, 'tag': 'suspect', 'type': 'suspect'})
susp_index = list(transactions[transactions['id'].isin(groups['source']['time'][id_])].index)
# get the out nodes
out = transactions[transactions['source'] == susp_node]
out_nodes = []
for i in out.index:
row = out.loc[i]
acc = True if i in susp_index else False
if acc:
tag = 'accomplice'
else:
tag = 'usual'
if row['target'] not in out_nodes:
data['nodes'].append({'id': row['target'], 'tag': tag, 'type': 'target'})
out_nodes.append(row['target'])
data['links'].append({'source': susp_node, 'target': row['target'], 'tag': tag, 'date': row['date'], 'time': row['time'], 'amount': row['amount'], 'currency': row['currency']})
in_ = transactions[transactions['target'] == susp_node]
in_nodes = []
if susp_node in target_ids:
susp_in = True
data['type'] = 'Time Pattern Outflow + Inflow'
idx = target_ids.index(susp_node)
susp_index_in = list(transactions[transactions['id'].isin(groups['target']['time'][idx])].index)
else:
data['type'] = 'Time Pattern Outflow'
susp_in = False
for i in in_.index:
row = in_.loc[i]
if susp_in:
acc = True if i in susp_index_in else False
if acc:
tag = 'accomplice'
else:
tag = 'usual'
else:
tag = 'usual'
if row['source'] not in in_nodes:
data['nodes'].append({'id': row['source'], 'tag': tag, 'type': 'source'})
in_nodes.append(row['source'])
data['links'].append({'source': row['source'], 'target': susp_node, 'tag': tag, 'date': row['date'], 'time': row['time'], 'amount': row['amount'], 'currency': row['currency']})
# Save the file
fn = str(susp_node) + '.json'
with open('json/'+fn, 'w') as outfile:
json.dump(data, outfile)
# +
list_ = []
for id_ in range(3):
data = {'nodes': [], 'links': [], 'type': None}
susp_node = transactions[transactions['id'].isin(groups['source']['time'][id_])]['source'].iloc[0]
score = np.random.random()
list_.append({'id': susp_node, 'score': score})
list_ = sorted(list_, key=lambda k: k['score'], reverse=True)
with open('json/ranked_list.json', 'w') as outfile:
json.dump(list_, outfile)
# -
list_
|
code/gael/data_analysis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + jupyter={"outputs_hidden": false}
# %pylab inline
# -
# Работа с распределением Вейбулла <http://numpy.readthedocs.io/en/latest/reference/generated/numpy.random.RandomState.weibull.html>
# + jupyter={"outputs_hidden": false}
# Draw samples from the distribution:
a = 5. # shape
s = np.random.weibull(a, 1000)
# Display the histogram of the samples, along with
# the probability density function:
#import matplotlib.pyplot as plt
x = np.arange(1,100.)/50.
def weib(x,n,a):
return (a / n) * (x / n)**(a - 1) * np.exp(-(x / n)**a)
count, bins, ignored = plt.hist(np.random.weibull(5.,1000))
x = np.arange(1,100.)/50.
scale = count.max()/weib(x, 1., 5.).max()
# plt.plot(x, weib(x, 1., 5.)*scale)
plot(x, weib(x, 1., 5.)*scale)
# plt.show()
# + jupyter={"outputs_hidden": false}
# Draw samples from the distribution:
a = 1. # задает само распределение
s = np.random.weibull(a, 1000)
#import matplotlib.pyplot as plt
x = np.arange(1,100.)/50.
def weib(x,n,a):
return (a / n) * (x / n)**(a - 1) * np.exp(-(x / n)**a)
count, bins, ignored = plt.hist(np.random.weibull(5.,1000))
x = np.arange(1,100.)/50.
scale = count.max()/weib(x, 1., 5.).max()
# plt.plot(x, weib(x, 1., 5.)*scale)
plot(x, weib(x, 1., 5.)*scale)
# plt.show()
# + jupyter={"outputs_hidden": false}
plot(s)
# + jupyter={"outputs_hidden": false}
sum = 0
for i in s:
sum+=i
print(sum/len(s))
# + jupyter={"outputs_hidden": false}
a = 1. # задает само распределение
s = np.random.weibull(a, 1000)
s
plot(s)
# -
|
random/.ipynb_checkpoints/02_base_weibull-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from DatasetHandler.BiwiBrowser import *
biwi = readBIWIDataset(subjectList = [s for s in range(1, 2)])#
frames, labelsList = [], []
for inputMatrix, labels in biwi:
frames.append(inputMatrix)
labelsList.append(labels)
inputMatrix.shape
inputMatrix, labels = frames[0], labelsList[0]
from keras.preprocessing.image import load_img
from keras.preprocessing.image import img_to_array
from keras.applications.vgg16 import preprocess_input
from keras.applications.vgg16 import decode_predictions
from keras.applications.vgg16 import VGG16
import keras
# Inspect the model
vgg16.summary()
# Verify things look as expected
full_model.summary()
patch_size = BIWI_Frame_Shape #(48, 64, 3)#
from keras.layers import *
from keras import Model
keras.backend.clear_session()
def getFinalModel():
dense_layer_1 = 1#int((patch_size[0] * patch_size[1]) / 1)0010#00000
dense_layer_2 = 8
num_classes = 6
vgg16 = VGG16(weights='imagenet', include_top=False, input_shape= patch_size)
block5_pool = vgg16.get_layer('block5_pool').output
gate = (int(d) for d in block5_pool.shape[1:])
#vgg16.summary()
# This shape has to match the last layer in VGG16 (without top)
dense_input = Input(shape=gate) #patch_size) #(15, 20, 512)
dense_output = Flatten(name='flatten')(dense_input)
dense_output = Dense(dense_layer_1, activation='relu', name='fc1')(dense_output)
dense_output = Dense(dense_layer_2, activation='relu', name='fc2')(dense_output)
dense_output = Dense(num_classes, activation='softmax', name='predictions')(dense_output)
top_model = Model(inputs=dense_input, outputs=dense_output, name='top_model')
# Now combine the two models
full_output = top_model(block5_pool)
full_model = Model(inputs=vgg16.input, outputs=full_output)
for layer in full_model.layers[:15]:
layer.trainable = False
full_model.compile(optimizer='adam', loss='mean_squared_error', metrics=['accuracy'])
return full_model
full_model = getFinalModel()
full_model.fit(inputMatrix, labels, batch_size=10, epochs=1, verbose=2)
|
DeepRL_For_HPE/Older_VGG16Runner_Notebooks/.ipynb_checkpoints/VGG16RunnerOnMSI-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# coding: utf-8
import numpy as np
def smooth_curve(x):
"""損失関数のグラフを滑らかにするために用いる
参考:http://glowingpython.blogspot.jp/2012/02/convolution-with-numpy.html
"""
window_len = 11
s = np.r_[x[window_len-1:0:-1], x, x[-1:-window_len:-1]]
w = np.kaiser(window_len, 2)
y = np.convolve(w/w.sum(), s, mode='valid')
return y[5:len(y)-5]
def shuffle_dataset(x, t):
"""データセットのシャッフルを行う
Parameters
----------
x : 訓練データ
t : 教師データ
Returns
-------
x, t : シャッフルを行った訓練データと教師データ
"""
permutation = np.random.permutation(x.shape[0])
x = x[permutation,:] if x.ndim == 2 else x[permutation,:,:,:]
t = t[permutation]
return x, t
def conv_output_size(input_size, filter_size, stride=1, pad=0):
return (input_size + 2*pad - filter_size) / stride + 1
def im2col(input_data, filter_h, filter_w, stride=1, pad=0):
"""
Parameters
----------
input_data : (データ数, チャンネル, 高さ, 幅)の4次元配列からなる入力データ
filter_h : フィルターの高さ
filter_w : フィルターの幅
stride : ストライド
pad : パディング
Returns
-------
col : 2次元配列
"""
N, C, H, W = input_data.shape
out_h = (H + 2*pad - filter_h)//stride + 1
out_w = (W + 2*pad - filter_w)//stride + 1
img = np.pad(input_data, [(0,0), (0,0), (pad, pad), (pad, pad)], 'constant')
col = np.zeros((N, C, filter_h, filter_w, out_h, out_w))
for y in range(filter_h):
y_max = y + stride*out_h
for x in range(filter_w):
x_max = x + stride*out_w
col[:, :, y, x, :, :] = img[:, :, y:y_max:stride, x:x_max:stride]
col = col.transpose(0, 4, 5, 1, 2, 3).reshape(N*out_h*out_w, -1)
return col
def col2im(col, input_shape, filter_h, filter_w, stride=1, pad=0):
"""
Parameters
----------
col :
input_shape : 入力データの形状(例:(10, 1, 28, 28))
filter_h :
filter_w
stride
pad
Returns
-------
"""
N, C, H, W = input_shape
out_h = (H + 2*pad - filter_h)//stride + 1
out_w = (W + 2*pad - filter_w)//stride + 1
col = col.reshape(N, out_h, out_w, C, filter_h, filter_w).transpose(0, 3, 4, 5, 1, 2)
img = np.zeros((N, C, H + 2*pad + stride - 1, W + 2*pad + stride - 1))
for y in range(filter_h):
y_max = y + stride*out_h
for x in range(filter_w):
x_max = x + stride*out_w
img[:, :, y:y_max:stride, x:x_max:stride] += col[:, :, y, x, :, :]
return img[:, :, pad:H + pad, pad:W + pad]
|
common/util.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.5.6 64-bit (''dcscn'': conda)'
# language: python
# name: python3
# ---
# !python3 train.py --dataset=0.25@0.25km_train --test_dataset=0.25@0.25km_valid --max_value=1366 --scale=4 --layers=7 --filters=32 --min_filters=8 --filters_decay_gamma=1.2 --nin_filters=24 --nin_filters2=8 --reconstruct_layers=0 --self_ensemble=1 --training_images=228688 --batch_image_size=32 --build_batch=True --do_benchmark=True --pixel_shuffler_filters=1
# !python3 evaluate.py --scale=4 --layers=7 --filters=32 --min_filters=8 --filters_decay_gamma=1.2 --nin_filters=24 --nin_filters2=8 --reconstruct_layers=0 --self_ensemble=1 --batch_image_size=32 --pixel_shuffler_filters=1 --test_dataset=0.25@0.25km_valid
# !python3 train.py --dataset=0.25@0.25km_train --test_dataset=0.25@0.25km_valid --max_value=1366 --scale=16 --layers=7 --filters=32 --min_filters=8 --filters_decay_gamma=1.2 --nin_filters=24 --nin_filters2=8 --reconstruct_layers=0 --self_ensemble=1 --training_images=228688 --batch_image_size=45 --build_batch=True --pixel_shuffler_filters=1
|
test.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="eIrvnAbGZ1wP"
# ##### Copyright 2019 The TensorFlow Authors.
# + cellView="form" colab_type="code" id="_A4IPZ-WZ9H7" colab={}
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="zfiHTzhkmNwd"
# # Keras RNN API
#
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/guide/keras/rnn">
# <img src="https://www.tensorflow.org/images/tf_logo_32px.png" />
# View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/guide/keras/rnn.ipynb">
# <img src="https://www.tensorflow.org/images/colab_logo_32px.png" />
# Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/guide/keras/rnn.ipynb">
# <img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />
# View source on GitHub</a>
# </td>
# <td>
# <a target="_blank" href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/guide/keras/rnn.ipynb">
# <img src="https://www.tensorflow.org/images/download_logo_32px.png" />
# Download notebook</a>
# </td>
# </table>
#
# + [markdown] colab_type="text" id="jfOdaQLhXLDR"
# Recurrent neural networks (RNN) are a class of neural networks that is powerful for modeling sequence data such as time series or natural language.
#
# Schematically, a RNN layer uses a `for` loop to iterate over the timesteps of a sequence, while maintaining an internal state that encodes information about the timesteps it has seen so far.
#
# The Keras RNN API is designed with a focus on:
#
# - **Ease of use**: the built-in `tf.keras.layers.RNN`, `tf.keras.layers.LSTM`, `tf.keras.layers.GRU` layers enable you to quickly build recurrent models without having to make difficult configuration choices.
#
# - **Ease of customization**: You can also define your own RNN cell layer (the inner part of the `for` loop) with custom behavior, and use it with the generic `tf.keras.layers.RNN` layer (the `for` loop itself). This allows you to quickly prototype different research ideas in a flexible way with minimal code.
#
# + [markdown] colab_type="text" id="QGJH5EKYoSHZ"
# ## Setup
# + colab_type="code" id="wJEBe8hTlB6W" colab={}
from __future__ import absolute_import, division, print_function, unicode_literals
import collections
import matplotlib.pyplot as plt
import numpy as np
try:
# # %tensorflow_version only exists in Colab.
# %tensorflow_version 2.x
except Exception:
pass
import tensorflow as tf
from tensorflow.keras import layers
# + [markdown] colab_type="text" id="DznzjxWCilt4"
# ## Build a simple model
#
# + [markdown] colab_type="text" id="H5tPG7KJirBj"
# There are three built-in RNN layers in Keras:
#
# 1. `tf.keras.layers.SimpleRNN`, a fully-connected RNN where the output from previous timestep is to be fed to next timestep.
#
# 2. `tf.keras.layers.GRU`, first proposed in [Learning Phrase Representations using RNN Encoder-Decoder for Statistical Machine Translation](https://arxiv.org/abs/1406.1078).
#
# 3. `tf.keras.layers.LSTM`, first proposed in [Long Short-Term Memory](https://www.bioinf.jku.at/publications/older/2604.pdf).
#
# In early 2015, Keras had the first reusable open-source Python implementations of LSTM and GRU.
#
# Here is a simple example of a `Sequential` model that processes sequences of integers, embeds each integer into a 64-dimensional vector, then processes the sequence of vectors using a `LSTM` layer.
# + colab_type="code" id="QHdAFEATnFpn" colab={}
model = tf.keras.Sequential()
# Add an Embedding layer expecting input vocab of size 1000, and
# output embedding dimension of size 64.
model.add(layers.Embedding(input_dim=1000, output_dim=64))
# Add a LSTM layer with 128 internal units.
model.add(layers.LSTM(128))
# Add a Dense layer with 10 units and softmax activation.
model.add(layers.Dense(10, activation='softmax'))
model.summary()
# + [markdown] colab_type="text" id="sVT4R7O3qDXM"
# ## Outputs and states
# + [markdown] colab_type="text" id="IOQnPR9eqLwk"
# By default, the output of a RNN layer contain a single vector per sample. This vector is the RNN cell output corresponding to the last timestep, containing information about the entire input sequence. The shape of this output is `(batch_size, units)` where `units` corresponds to the `units` argument passed to the layer's constructor.
#
# A RNN layer can also return the entire sequence of outputs for each sample (one vector per timestep per sample), if you set `return_sequences=True`. The shape of this output is `(batch_size, timesteps, units)`.
# + colab_type="code" id="wNlkR8oXpNEx" colab={}
model = tf.keras.Sequential()
model.add(layers.Embedding(input_dim=1000, output_dim=64))
# The output of GRU will be a 3D tensor of shape (batch_size, timesteps, 256)
model.add(layers.GRU(256, return_sequences=True))
# The output of SimpleRNN will be a 2D tensor of shape (batch_size, 128)
model.add(layers.SimpleRNN(128))
model.add(layers.Dense(10, activation='softmax'))
model.summary()
# + [markdown] colab_type="text" id="1HagyjYos5rD"
# In addition, a RNN layer can return its final internal state(s). The returned states can be used to resume the RNN execution later, or [to initialize another RNN](https://arxiv.org/abs/1409.3215). This setting is commonly used in the encoder-decoder sequence-to-sequence model, where the encoder final state is used as the initial state of the decoder.
#
# To configure a RNN layer to return its internal state, set the `return_state` parameter to `True` when creating the layer. Note that `LSTM` has 2 state tensors, but `GRU` only has one.
#
# To configure the initial state of the layer, just call the layer with additional keyword argument `initial_state`.
# Note that the shape of the state needs to match the unit size of the layer, like in the example below.
# + colab_type="code" id="2_HsGBrDvaea" colab={}
encoder_vocab = 1000
decoder_vocab = 2000
encoder_input = layers.Input(shape=(None, ))
encoder_embedded = layers.Embedding(input_dim=encoder_vocab, output_dim=64)(encoder_input)
# Return states in addition to output
output, state_h, state_c = layers.LSTM(
64, return_state=True, name='encoder')(encoder_embedded)
encoder_state = [state_h, state_c]
decoder_input = layers.Input(shape=(None, ))
decoder_embedded = layers.Embedding(input_dim=decoder_vocab, output_dim=64)(decoder_input)
# Pass the 2 states to a new LSTM layer, as initial state
decoder_output = layers.LSTM(
64, name='decoder')(decoder_embedded, initial_state=encoder_state)
output = layers.Dense(10, activation='softmax')(decoder_output)
model = tf.keras.Model([encoder_input, decoder_input], output)
model.summary()
# + [markdown] colab_type="text" id="kJDJSXjZ2VaY"
# ## RNN layers and RNN cells
# + [markdown] colab_type="text" id="hQRxLRSS2gDf"
# In addition to the built-in RNN layers, the RNN API also provides cell-level APIs. Unlike RNN layers, which processes whole batches of input sequences, the RNN cell only processes a single timestep.
#
# The cell is the inside of the `for` loop of a RNN layer. Wrapping a cell inside a `tf.keras.layers.RNN` layer gives you a layer capable of processing batches of sequences, e.g. `RNN(LSTMCell(10))`.
#
# Mathemetically, `RNN(LSTMCell(10))` produces the same result as `LSTM(10)`. In fact, the implementation of this layer in TF v1.x was just creating the corresponding RNN cell and wrapping it in a RNN layer. However using the built-in `GRU` and `LSTM` layers enables the use of CuDNN and you may see better performance.
#
# There are three built-in RNN cells, each of them corresponding to the matching RNN layer.
#
# - `tf.keras.layers.SimpleRNNCell` corresponds to the `SimpleRNN` layer.
#
# - `tf.keras.layers.GRUCell` corresponds to the `GRU` layer.
#
# - `tf.keras.layers.LSTMCell` corresponds to the `LSTM` layer.
#
# The cell abstraction, together with the generic `tf.keras.layers.RNN` class, make it very easy to implement custom RNN architectures for your research.
#
# + [markdown] id="veiCKSUU-ina" colab_type="text"
# ## Cross-batch statefulness
# + [markdown] colab_type="text" id="EvAaiMJbWR2A"
# When processing very long sequences (possibly infinite), you may want to use the pattern of **cross-batch statefulness**.
#
# Normally, the internal state of a RNN layer is reset every time it sees a new batch (i.e. every sample seen by the layer is assume to be independent from the past). The layer will only maintain a state while processing a given sample.
#
# If you have very long sequences though, it is useful to break them into shorter sequences, and to feed these shorter sequences sequentially into a RNN layer without resetting the layer's state. That way, the layer can retain information about the entirety of the sequence, even though it's only seeing one sub-sequence at a time.
#
# You can do this by setting `stateful=True` in the constructor.
#
# If you have a sequence `s = [t0, t1, ... t1546, t1547]`, you woud split it into e.g.
#
# ```
# s1 = [t0, t1, ... t100]
# s2 = [t101, ... t201]
# ...
# s16 = [t1501, ... t1547]
# ```
#
# Then you would process it via:
#
# ```python
# lstm_layer = layers.LSTM(64, stateful=True)
# for s in sub_sequences:
# output = lstm_layer(s)
# ```
#
# When you want to clear the state, you can use `layer.reset_states()`.
#
#
# > Note: In this setup, sample `i` in a given batch is assumed to be the continuation of sample `i` in the previous batch. This means that all batches should contain the same number of samples (batch size). E.g. if a batch contains `[sequence_A_from_t0_to_t100, sequence_B_from_t0_to_t100]`, the next batch should contain `[sequence_A_from_t101_to_t200, sequence_B_from_t101_to_t200]`.
#
#
#
#
# Here is a complete example:
#
#
# + colab_type="code" id="E6TsLXJ0X3Xd" colab={}
paragraph1 = np.random.random((20, 10, 50)).astype(np.float32)
paragraph2 = np.random.random((20, 10, 50)).astype(np.float32)
paragraph3 = np.random.random((20, 10, 50)).astype(np.float32)
lstm_layer = layers.LSTM(64, stateful=True)
output = lstm_layer(paragraph1)
output = lstm_layer(paragraph2)
output = lstm_layer(paragraph3)
# reset_states() will reset the cached state to the original initial_state.
# If no initial_state was provided, zero-states will be used by default.
lstm_layer.reset_states()
# + [markdown] colab_type="text" id="7AtPur5BDzb4"
# ##Bidirectional RNNs
# + [markdown] colab_type="text" id="OsdEIXXREL_N"
# For sequences other than time series (e.g. text), it is often the case that a RNN model can perform better if it not only processes sequence from start to end, but also backwards. For example, to predict the next word in a sentence, it is often useful to have the context around the word, not only just the words that come before it.
#
# Keras provides an easy API for you to build such bidirectional RNNs: the `tf.keras.layers.Bidirectional` wrapper.
# + colab_type="code" id="MNhYIAXqYl3B" colab={}
model = tf.keras.Sequential()
model.add(layers.Bidirectional(layers.LSTM(64, return_sequences=True),
input_shape=(5, 10)))
model.add(layers.Bidirectional(layers.LSTM(32)))
model.add(layers.Dense(10, activation='softmax'))
model.summary()
# + [markdown] colab_type="text" id="ThwlodTjZCU0"
# Under the hood, `Bidirectional` will copy the RNN layer passed in, and flip the `go_backwards` field of the newly copied layer, so that it will process the inputs in reverse order.
#
# The output of the `Bidirectional` RNN will be, by default, the sum of the forward layer output and the backward layer output. If you need a different merging behavior, e.g. concatenation, change the `merge_mode` parameter in the `Bidirectional` wrapper constructor. For more details about `Bidirectional`, please check [the API docs](https://www.tensorflow.org/versions/r2.0/api_docs/python/tf/keras/layers/Bidirectional).
# + [markdown] colab_type="text" id="ANGN956w6FRs"
# ## Performance optimization and CuDNN kernels in TensorFlow 2.0
# + [markdown] colab_type="text" id="76xAs7epaX21"
# In Tensorflow 2.0, the built-in LSTM and GRU layers have been updated to leverage CuDNN kernels by default when a GPU is available. With this change, the prior `keras.layers.CuDNNLSTM/CuDNNGRU` layers have been deprecated, and you can build your model without worrying about the hardware it will run on.
#
# Since the CuDNN kernel is built with certain assumptions, this means the layer **will not be able to use the CuDNN kernel if you change the defaults of the built-in LSTM or GRU layers**. E.g.:
#
# - Changing the `activation` function from `tanh` to something else.
# - Changing the `recurrent_activation` function from `sigmoid` to something else.
# - Using `recurrent_dropout` > 0.
# - Setting `unroll` to True, which forces LSTM/GRU to decompose the inner `tf.while_loop` into an unrolled `for` loop.
# - Setting `use_bias` to False.
# - Using masking when the input data is not strictly right padded (if the mask corresponds to strictly right padded data, CuDNN can still be used. This is the most common case).
#
# For the detailed list of contraints, please see the documentation for the [LSTM](https://www.tensorflow.org/versions/r2.0/api_docs/python/tf/keras/layers/LSTM) and [GRU](https://www.tensorflow.org/versions/r2.0/api_docs/python/tf/keras/layers/GRU) layers.
# + [markdown] colab_type="text" id="ybd73JmvqLp4"
# ### Using CuDNN kernels when available
#
# Let's build a simple LSTM model to demonstrate the performance difference.
#
# We'll use as input sequences the sequence of rows of MNIST digits (treating each row of pixels as a timestep), and we'll predict the digit's label.
#
# + colab_type="code" id="m9kM9hwRsxMx" colab={}
batch_size = 64
# Each MNIST image batch is a tensor of shape (batch_size, 28, 28).
# Each input sequence will be of size (28, 28) (height is treated like time).
input_dim = 28
units = 64
output_size = 10 # labels are from 0 to 9
# Build the RNN model
def build_model(allow_cudnn_kernel=True):
# CuDNN is only available at the layer level, and not at the cell level.
# This means `LSTM(units)` will use the CuDNN kernel,
# while RNN(LSTMCell(units)) will run on non-CuDNN kernel.
if allow_cudnn_kernel:
# The LSTM layer with default options uses CuDNN.
lstm_layer = tf.keras.layers.LSTM(units, input_shape=(None, input_dim))
else:
# Wrapping a LSTMCell in a RNN layer will not use CuDNN.
lstm_layer = tf.keras.layers.RNN(
tf.keras.layers.LSTMCell(units),
input_shape=(None, input_dim))
model = tf.keras.models.Sequential([
lstm_layer,
tf.keras.layers.BatchNormalization(),
tf.keras.layers.Dense(output_size, activation='softmax')]
)
return model
# + [markdown] colab_type="text" id="uuztNezFh0BL"
# ### Load MNIST dataset
# + colab_type="code" id="m_kZTLDobchi" colab={}
mnist = tf.keras.datasets.mnist
(x_train, y_train), (x_test, y_test) = mnist.load_data()
x_train, x_test = x_train / 255.0, x_test / 255.0
sample, sample_label = x_train[0], y_train[0]
# + [markdown] colab_type="text" id="UXF8elCuib8k"
# ### Create a model instance and compile it
# We choose `sparse_categorical_crossentropy` as the loss function for the model. The output of the model has shape of `[batch_size, 10]`. The target for the model is a integer vector, each of the integer is in the range of 0 to 9.
# + colab_type="code" id="klgv6dfK0KNb" colab={}
model = build_model(allow_cudnn_kernel=True)
model.compile(loss='sparse_categorical_crossentropy',
optimizer='sgd',
metrics=['accuracy'])
# + colab_type="code" id="qzeeo65r25CU" colab={}
model.fit(x_train, y_train,
validation_data=(x_test, y_test),
batch_size=batch_size,
epochs=5)
# + [markdown] colab_type="text" id="kvCWAssZjsdW"
# ### Build a new model without CuDNN kernel
# + colab_type="code" id="H2JfHDOhOFtx" colab={}
slow_model = build_model(allow_cudnn_kernel=False)
slow_model.set_weights(model.get_weights())
slow_model.compile(loss='sparse_categorical_crossentropy',
optimizer='sgd',
metrics=['accuracy'])
slow_model.fit(x_train, y_train,
validation_data=(x_test, y_test),
batch_size=batch_size,
epochs=1) # We only train for one epoch because it's slower.
# + [markdown] colab_type="text" id="Zx8QLf81dTVr"
# As you can see, the model built with CuDNN is much faster to train compared to the model that use the regular TensorFlow kernel.
#
# The same CuDNN-enabled model can also be use to run inference in a CPU-only environment. The `tf.device` annotation below is just forcing the device placement. The model will run on CPU by default if no GPU is available.
#
# You simply don't have to worry about the hardware you're running on anymore. Isn't that pretty cool?
# + colab_type="code" id="z_z1eRh1fMBL" colab={}
with tf.device('CPU:0'):
cpu_model = build_model(allow_cudnn_kernel=True)
cpu_model.set_weights(model.get_weights())
result = tf.argmax(cpu_model.predict_on_batch(tf.expand_dims(sample, 0)), axis=1)
print('Predicted result is: %s, target result is: %s' % (result.numpy(), sample_label))
plt.imshow(sample, cmap=plt.get_cmap('gray'))
# + [markdown] colab_type="text" id="2mCetBoTiqcB"
# ## RNNs with list/dict inputs, or nested inputs
#
# Nested structures allow implementers to include more information within a single timestep. For example, a video frame could have audio and video input at the same time. The data shape in this case could be:
#
# `[batch, timestep, {"video": [height, width, channel], "audio": [frequency]}]`
#
# In another example, handwriting data could have both coordinates x and y for the current position of the pen, as well as pressure information. So the data representation could be:
#
# `[batch, timestep, {"location": [x, y], "pressure": [force]}]`
#
# The following code provides an example of how to build a custom RNN cell that accepts such structured inputs.
#
# + [markdown] colab_type="text" id="A1IkIxWykSZQ"
# ### Define a custom cell that support nested input/output
# + colab_type="code" id="6yOT8nSqzp4A" colab={}
NestedInput = collections.namedtuple('NestedInput', ['feature1', 'feature2'])
NestedState = collections.namedtuple('NestedState', ['state1', 'state2'])
class NestedCell(tf.keras.layers.Layer):
def __init__(self, unit_1, unit_2, unit_3, **kwargs):
self.unit_1 = unit_1
self.unit_2 = unit_2
self.unit_3 = unit_3
self.state_size = NestedState(state1=unit_1,
state2=tf.TensorShape([unit_2, unit_3]))
self.output_size = (unit_1, tf.TensorShape([unit_2, unit_3]))
super(NestedCell, self).__init__(**kwargs)
def build(self, input_shapes):
# expect input_shape to contain 2 items, [(batch, i1), (batch, i2, i3)]
input_1 = input_shapes.feature1[1]
input_2, input_3 = input_shapes.feature2[1:]
self.kernel_1 = self.add_weight(
shape=(input_1, self.unit_1), initializer='uniform', name='kernel_1')
self.kernel_2_3 = self.add_weight(
shape=(input_2, input_3, self.unit_2, self.unit_3),
initializer='uniform',
name='kernel_2_3')
def call(self, inputs, states):
# inputs should be in [(batch, input_1), (batch, input_2, input_3)]
# state should be in shape [(batch, unit_1), (batch, unit_2, unit_3)]
input_1, input_2 = tf.nest.flatten(inputs)
s1, s2 = states
output_1 = tf.matmul(input_1, self.kernel_1)
output_2_3 = tf.einsum('bij,ijkl->bkl', input_2, self.kernel_2_3)
state_1 = s1 + output_1
state_2_3 = s2 + output_2_3
output = [output_1, output_2_3]
new_states = NestedState(state1=state_1, state2=state_2_3)
return output, new_states
# + [markdown] colab_type="text" id="BJHOrrybk6Zy"
# ### Build a RNN model with nested input/output
#
# Let's build a Keras model that uses a `tf.keras.layers.RNN` layer and the custom cell we just defined.
# + colab_type="code" id="itrDe0Y2qPjP" colab={}
unit_1 = 10
unit_2 = 20
unit_3 = 30
input_1 = 32
input_2 = 64
input_3 = 32
batch_size = 64
num_batch = 100
timestep = 50
cell = NestedCell(unit_1, unit_2, unit_3)
rnn = tf.keras.layers.RNN(cell)
inp_1 = tf.keras.Input((None, input_1))
inp_2 = tf.keras.Input((None, input_2, input_3))
outputs = rnn(NestedInput(feature1=inp_1, feature2=inp_2))
model = tf.keras.models.Model([inp_1, inp_2], outputs)
model.compile(optimizer='adam', loss='mse', metrics=['accuracy'])
# + [markdown] colab_type="text" id="2MaihTM2mDcp"
# ### Train the model with randomly generated data
#
# Since there isn't a good candidate dataset for this model, we use random Numpy data for demonstration.
# + colab_type="code" id="lN-imRqElz2S" colab={}
input_1_data = np.random.random((batch_size * num_batch, timestep, input_1))
input_2_data = np.random.random((batch_size * num_batch, timestep, input_2, input_3))
target_1_data = np.random.random((batch_size * num_batch, unit_1))
target_2_data = np.random.random((batch_size * num_batch, unit_2, unit_3))
input_data = [input_1_data, input_2_data]
target_data = [target_1_data, target_2_data]
model.fit(input_data, target_data, batch_size=batch_size)
# + [markdown] colab_type="text" id="oDdrwgBWnjYp"
# With the Keras `tf.keras.layers.RNN` layer, You are only expected to define the math logic for individual step within the sequence, and the `tf.keras.layers.RNN` layer will handle the sequence iteration for you. It's an incredibly powerful way to quickly prototype new kinds of RNNs (e.g. a LSTM variant).
#
# For more details, please visit the [API docs](https://www.tensorflow.org/versions/r2.0/api_docs/python/tf/keras/layers/RNN).
|
site/en/guide/keras/rnn.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Dependencies
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from scipy import stats
# +
# Read in data
# +
# Run the t-test
# +
# Report the data
# +
# Plot sample means with error bars
# +
# Plot mean height of players
# -
|
05-Matplotlib/3/Activities/08-Stu_Students_t_test/Unsolved/ttest.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
videos = """
https://www.youtube.com/watch?v=cyU3Qgox3K4&t=131s&ab_channel=TanahMelayu
https://www.youtube.com/watch?v=68bH2c04v7o&ab_channel=TanahMelayu
https://www.youtube.com/watch?v=9ITPO6ooNSk&ab_channel=TanahMelayu
https://www.youtube.com/watch?v=sw5h9hlityE&t=1284s&ab_channel=TanahMelayu
https://www.youtube.com/watch?v=rwhXalBKmbE&ab_channel=TanahMelayu
https://www.youtube.com/watch?v=O4mJ_Fufp0o&ab_channel=TanahMelayu
https://www.youtube.com/watch?v=_9ZBUQjJfMI&ab_channel=TanahMelayu
https://www.youtube.com/watch?v=NKSyshvDHWY&ab_channel=TanahMelayu
https://www.youtube.com/watch?v=hIjJGdzOhk8&ab_channel=TanahMelayu
https://www.youtube.com/watch?v=trY5KYqaiXw&ab_channel=TanahMelayu
https://www.youtube.com/watch?v=6f-8js73O1k
https://www.youtube.com/watch?v=NSRWj5Zz7Qo
https://www.youtube.com/watch?v=glR3xP2HTS0
https://www.youtube.com/watch?v=ydoo5KXzJbg
https://www.youtube.com/watch?v=ph_fuYS7ucg&ab_channel=AbuKhadijah
https://www.youtube.com/watch?v=RnNs7-49jmE&ab_channel=AbuKhadijah
https://www.youtube.com/watch?v=PhYkAGmhgTQ&ab_channel=AbuKhadijah
https://www.youtube.com/watch?v=hCWxKltGsWc&ab_channel=AbuKhadijah
https://www.youtube.com/watch?v=W6FXt5p9V3U&ab_channel=AbuKhadijah
https://www.youtube.com/watch?v=VFC4UDw8ju0&ab_channel=AbuKhadijah
https://www.youtube.com/watch?v=R4DHxsDHzF8&ab_channel=AbuKhadijah
https://www.youtube.com/watch?v=rQ_P9xo6pfo&ab_channel=AbuKhadijah
https://www.youtube.com/watch?v=QW3GndtSb1k
https://www.youtube.com/watch?v=arZATVyf8j8
https://www.youtube.com/watch?v=KkjtaMemWIc
https://www.youtube.com/watch?v=qDNwTIZlJRo
https://www.youtube.com/watch?v=rARJlDAJEgE
https://www.youtube.com/watch?v=Z9v2NDMciIk
https://www.youtube.com/watch?v=N3tDIGVt9d8
https://www.youtube.com/watch?v=PXuIhQevtqM
https://www.youtube.com/watch?v=038_W7kvhI0
https://www.youtube.com/watch?v=P4TN4HDiEPk
https://www.youtube.com/watch?v=CR-JLqCElrY
https://www.youtube.com/watch?v=DXDML7x2FQs
https://www.youtube.com/watch?v=NB4UnmMFsUE
https://www.youtube.com/watch?v=IFhlmN0bDyI
https://www.youtube.com/watch?v=0ca50QUW5OI
https://www.youtube.com/watch?v=TwKDWH9hSfQ
https://www.youtube.com/watch?v=Fv8g4WBYL5k
https://www.youtube.com/watch?v=RHkqyW16nAM
https://www.youtube.com/watch?v=TSW_LB7p_rk
https://www.youtube.com/watch?v=bDHdtkmA-Q0
https://www.youtube.com/watch?v=dF7HRsk95aY
https://www.youtube.com/watch?v=HYf4V_0T1rk
https://www.youtube.com/watch?v=QSWRFVfc55k
https://www.youtube.com/watch?v=ip2EWKhmOYY
https://www.youtube.com/watch?v=s4hfxsYVnpE
https://www.youtube.com/watch?v=xw2IYlEVSn8
https://www.youtube.com/watch?v=ds3-U1BeznE
https://www.youtube.com/watch?v=-t8sUAMQ94M
https://www.youtube.com/watch?v=uZ4KTG1gQoI
https://www.youtube.com/watch?v=Cno40z2_-oU
https://www.youtube.com/watch?v=INq9ouq6EJc
https://www.youtube.com/watch?v=ZFw2QoiRi68
https://www.youtube.com/watch?v=gynmylUhxLg
https://www.youtube.com/watch?v=pu1B9RTGRfk
https://www.youtube.com/watch?v=rBB0rlZTL0o
https://www.youtube.com/watch?v=Wm-8ZeLyux0
https://www.youtube.com/watch?v=EP2qcEFmFD8
https://www.youtube.com/watch?v=o1Gxi86NM3M
https://www.youtube.com/watch?v=32GoCGuk4R4
https://www.youtube.com/watch?v=zlpmlFBj83M
https://www.youtube.com/watch?v=NPwVuX5T-fU
https://www.youtube.com/watch?v=mQuUzU8xQ44
https://www.youtube.com/watch?v=aHYimafirpg
https://www.youtube.com/watch?v=DfflcWqHY9E
https://www.youtube.com/watch?v=5Yu8ZklAV1Q
https://www.youtube.com/watch?v=jW6GYlyus4A
https://www.youtube.com/watch?v=crqask4aeJ0
https://www.youtube.com/watch?v=QkJQ4LeZwcI
https://www.youtube.com/watch?v=dpfE6ek6sEg
https://www.youtube.com/watch?v=wGY7jUCuU3Y
https://www.youtube.com/watch?v=GKQqdCXb8l8
https://www.youtube.com/watch?v=e5cIN57hJ_0
https://www.youtube.com/watch?v=Xwhd8aOFafs&ab_channel=BuletinTV3
https://www.youtube.com/watch?v=VdBxCz-Fw_0&ab_channel=BuletinTV3
https://www.youtube.com/watch?v=AVt7WL-ZR4Y
https://www.youtube.com/watch?v=bRUjmsnE5ZU&t=155s&ab_channel=PurePixels
https://www.youtube.com/watch?v=nPsVmw-GF4A&t=600s
https://www.youtube.com/watch?v=BWU8IJBEjTk&t=948s
https://www.youtube.com/watch?v=Am7A-vpCER0&ab_channel=PurePixels
https://www.youtube.com/watch?v=gsRtHPYjQCw
https://www.youtube.com/watch?v=gUOp5gBhz2I
https://www.youtube.com/watch?v=i7liW-7TTOI&ab_channel=PurePixels
https://www.youtube.com/watch?v=yRJWsihQ60s
https://www.youtube.com/watch?v=QoA1y20tyOw&ab_channel=Ajar
https://www.youtube.com/watch?v=_ZnpSEffVus&ab_channel=LearningWithShirley
https://www.youtube.com/watch?v=6XVf5bmw_PU&ab_channel=Bing-BahasaMelayu
https://www.youtube.com/watch?v=AyoBUd469wA&ab_channel=Bing-BahasaMelayu
https://www.youtube.com/watch?v=UtzK54Pb7Xc&ab_channel=Bing-BahasaMelayu
https://www.youtube.com/watch?v=EXgPLTHDM-c&ab_channel=Bing-BahasaMelayu
https://www.youtube.com/watch?v=qNqK6_fjBGE
https://www.youtube.com/watch?v=dn0mDC9XaoE
https://www.youtube.com/watch?v=gYpA-GwuMco
https://www.youtube.com/watch?v=FLRVppiXPu8&ab_channel=Ajar
https://www.youtube.com/watch?v=PKJshdf_FnM&ab_channel=Ajar
https://www.youtube.com/watch?v=DnOIVS6ylqs
https://www.youtube.com/watch?v=nWCATzTi0HU&ab_channel=Ajar
https://www.youtube.com/watch?v=5VP6w6ilB04&ab_channel=Ajar
https://www.youtube.com/watch?v=VNYyHVdO8_U
https://www.youtube.com/watch?v=Z2KnOKHBWFY
https://www.youtube.com/watch?v=mXFQb1ghFco
https://www.youtube.com/watch?v=xbYDNw0ZV78
https://www.youtube.com/watch?v=yyRdpqr45eo
https://www.youtube.com/watch?v=XRREYTMwLL0
https://www.youtube.com/watch?v=jCoNNZA8s8c
https://www.youtube.com/watch?v=C_PJ2FHFjbo
https://www.youtube.com/watch?v=lTQJminlKn0
https://www.youtube.com/watch?v=F7Zp1yMq2UM
https://www.youtube.com/watch?v=ULwl3_VSMS8
https://www.youtube.com/watch?v=uLcd4-qE-FQ
https://www.youtube.com/watch?v=Cu-UEBq8QhU
https://www.youtube.com/watch?v=Ktf542-Jeac
https://www.youtube.com/watch?v=FhLi9mWRBRQ
https://www.youtube.com/watch?v=0aiJynNT2Fk
https://www.youtube.com/watch?v=Gw59dGbVzBA
https://www.youtube.com/watch?v=u481vfr8q3c
https://www.youtube.com/watch?v=CPzDF8FUdcU
https://www.youtube.com/watch?v=sf2wUigdiuU
https://www.youtube.com/watch?v=n-0KdpNJzHQ
https://www.youtube.com/watch?v=bjXTuQrpkC8
https://www.youtube.com/watch?v=CubnJGvZFpk
https://www.youtube.com/watch?v=b59TtMqYusw
https://www.youtube.com/watch?v=gPappt9MHPc
https://www.youtube.com/watch?v=e6FH8wtaltw
https://www.youtube.com/watch?v=YKe-mzGU0JU
https://www.youtube.com/watch?v=1v-n23HAi7M
https://www.youtube.com/watch?v=WC0_yaR0jY4
https://www.youtube.com/watch?v=QO-YWGQvb1Q
https://www.youtube.com/watch?v=o7MmHE7Hmg4
https://www.youtube.com/watch?v=HGFCASE37OQ
https://www.youtube.com/watch?v=2naYfcudqss
https://www.youtube.com/watch?v=TwOgCpknOUU
https://www.youtube.com/watch?v=iqxyI-eYK3M
https://www.youtube.com/watch?v=4sYNZ3vkIbg
https://www.youtube.com/watch?v=opAw8b75NQM
https://www.youtube.com/watch?v=SVViu1hC4UI
https://www.youtube.com/watch?v=At-WKdApIhs&t=1227s&ab_channel=SEISMIKTV
https://www.youtube.com/watch?v=bXVvXLHpmS8&t=1027s&ab_channel=naddyepul
https://www.youtube.com/watch?v=_bfH3pxuOoM&t=1235s&ab_channel=khairuljamain
https://www.youtube.com/watch?v=e-eMNNxW3vo&t=1507s
https://www.youtube.com/watch?v=uCg5WeQ34rM&t=765s&ab_channel=AmirArsad
https://www.youtube.com/watch?v=VlmIyZllgz0
https://www.youtube.com/watch?v=4ZiC_Tg6hmU&t=774s&ab_channel=ONZZ
https://www.youtube.com/watch?v=0UHRMfS3Vqo&ab_channel=khairuljamain
https://www.youtube.com/watch?v=XJTfe7fVaGc&t=366s
https://www.youtube.com/watch?v=nzxL4rgpUpY&t=427s
https://www.youtube.com/watch?v=hOpZQQG2qg4
https://www.youtube.com/watch?v=sSWZVKSsIlU
https://www.youtube.com/watch?v=BoX_hMvaXWM
https://www.youtube.com/watch?v=riEwi1I6KKg
https://www.youtube.com/watch?v=mNZRQE-QDiM
https://www.youtube.com/watch?v=_Zsx149RtiU&t=632s
https://www.youtube.com/watch?v=WKi6VUFzjuo
https://www.youtube.com/watch?v=4jlZgyvBq7c
https://www.youtube.com/watch?v=uPruQ1sr3o8
https://www.youtube.com/watch?v=LtARertQkUo
https://www.youtube.com/watch?v=e7xE7V11Vwc
https://www.youtube.com/watch?v=eJE2FaX8CPE
https://www.youtube.com/watch?v=JXcolry-4Pk
https://www.youtube.com/watch?v=6VAfydhtJGg
https://www.youtube.com/watch?v=uq8eKjAy0us
https://www.youtube.com/watch?v=hXDFROt0dFI
https://www.youtube.com/watch?v=aZcwqUZI2rA&t=994s
https://www.youtube.com/watch?v=a34vZ2fwa4o
https://www.youtube.com/watch?v=Oeof60EnP0s
https://www.youtube.com/watch?v=WbP1SCK0nQQ
https://www.youtube.com/watch?v=MXScwqRvvxc&ab_channel=WanAnimation
https://www.youtube.com/watch?v=qkDvoyZDEh8&ab_channel=WanAnimation
https://www.youtube.com/watch?v=r86fA5i3PaE
https://www.youtube.com/watch?v=rmiKORIv0uw
https://www.youtube.com/watch?v=iW946gD8BcI
https://www.youtube.com/watch?v=dNgrEYnaa0U
https://www.youtube.com/watch?v=uLQDcX9i2XM
https://www.youtube.com/watch?v=enDHYM-0INM
https://www.youtube.com/watch?v=-q36CzKlwbQ
https://www.youtube.com/watch?v=0ZKxMaNusTA
https://www.youtube.com/watch?v=4ZiC_Tg6hmU&t=780s&ab_channel=ONZZ
https://www.youtube.com/watch?v=4Ga8I27AFZg
https://www.youtube.com/watch?v=S8zARg4V6VM
https://www.youtube.com/watch?v=DCNRWKiahbE
https://www.youtube.com/watch?v=LWDEe2fHJpk
https://www.youtube.com/watch?v=rpbjezVZ2GI
https://www.youtube.com/watch?v=fW1sUT-3VMM
https://www.youtube.com/watch?v=I9eEz2OvI0E&t=52s
https://www.youtube.com/watch?v=FQDcHVgZCTE
https://www.youtube.com/watch?v=4DCkzN-pdyw&t=77s
https://www.youtube.com/watch?v=Sau4UqPjBBk
https://www.youtube.com/watch?v=_gQ-uHjCdBs
https://www.youtube.com/watch?v=Q5kDJPXJPbc
https://www.youtube.com/watch?v=JIOEzlTO6Kg&t=232s
https://www.youtube.com/watch?v=JKnaHxIYHm8&ab_channel=SlowTalkPodcast
https://www.youtube.com/watch?v=l-NmUn-FLLI
https://www.youtube.com/watch?v=-Lc21hhbAfE
https://www.youtube.com/watch?v=fQyh4O6Hp_E
https://www.youtube.com/watch?v=qbFORSE48cQ&ab_channel=EduwebTV
https://www.youtube.com/watch?v=t__QLuABpvw&ab_channel=EduwebTV
https://www.youtube.com/watch?v=_Bw0Ah2-rMM&ab_channel=EduwebTV
https://www.youtube.com/watch?v=UPp7Rx6D4tg
https://www.youtube.com/watch?v=S_A4zDZD2qc
https://www.youtube.com/watch?v=sCtvZPgGZCg&ab_channel=EduwebTV
https://www.youtube.com/watch?v=IjrjbjyIHF8&ab_channel=EduwebTV
https://www.youtube.com/watch?v=Te-funnFw3s&ab_channel=EduwebTV
https://www.youtube.com/watch?v=D7VNROr3GQs&ab_channel=EduwebTV
https://www.youtube.com/watch?v=aBO1inBeQRI&ab_channel=EduwebTV
https://www.youtube.com/watch?v=gqo3pB3FVmg
https://www.youtube.com/watch?v=NeiArFP2Itc
https://www.youtube.com/watch?v=-Sd927cAvnQ&ab_channel=EduwebTV
https://www.youtube.com/watch?v=QOCbT-XH36I
https://www.youtube.com/watch?v=YNuLH_RP_v0
https://www.youtube.com/watch?v=ARNSWWs1x6g&ab_channel=EduwebTV
https://www.youtube.com/watch?v=L9urGAc2084&ab_channel=EduwebTV
https://www.youtube.com/watch?v=8ajsI0iCBMo&ab_channel=EduwebTV
https://www.youtube.com/watch?v=6G0N1OrIR4k&ab_channel=EduwebTV
https://www.youtube.com/watch?v=oJ0xApNXRBo
https://www.youtube.com/watch?v=_Y0zX_Uh2eI&ab_channel=EduwebTV
https://www.youtube.com/watch?v=VCysrXUWc_w
https://www.youtube.com/watch?v=YNuLH_RP_v0
https://www.youtube.com/watch?v=4UCEW_MX2p8
https://www.youtube.com/watch?v=VCysrXUWc_w
https://www.youtube.com/watch?v=un7dcUafjSc&t=1828s
https://www.youtube.com/watch?v=f2xNnNcwmK8
https://www.youtube.com/watch?v=00spujx3b70
https://www.youtube.com/watch?v=MZCJCoBwuho
https://www.youtube.com/watch?v=LcSKIOdHbbc
https://www.youtube.com/watch?v=9aoNnGW8hto
https://www.youtube.com/watch?v=36_nkekh-gE
https://www.youtube.com/watch?v=6GwVOXHYZlQ
https://www.youtube.com/watch?v=Q0ICpOpSZKU&ab_channel=EduwebTV
https://www.youtube.com/watch?v=At-WKdApIhs&t=654s&ab_channel=SEISMIKTV
https://www.youtube.com/watch?v=I9eEz2OvI0E&t=3s&ab_channel=SlowTalkPodcast
https://www.youtube.com/watch?v=fQyh4O6Hp_E
https://www.youtube.com/watch?v=rpbjezVZ2GI
https://www.youtube.com/watch?v=fW1sUT-3VMM
https://www.youtube.com/watch?v=neCmuMPg_IE
https://www.youtube.com/channel/UCmBSFLogevWdrid3MM5xReg/videos
https://www.youtube.com/watch?v=Sau4UqPjBBk
https://www.youtube.com/watch?v=4DCkzN-pdyw
https://www.youtube.com/watch?v=_gQ-uHjCdBs
https://www.youtube.com/watch?v=Q5kDJPXJPbc
https://www.youtube.com/watch?v=JIOEzlTO6Kg
https://www.youtube.com/watch?v=kTrI0LaiY8U
https://www.youtube.com/watch?v=JKnaHxIYHm8
https://www.youtube.com/watch?v=RdqVSxXwQIg
https://www.youtube.com/watch?v=l-NmUn-FLLI
https://www.youtube.com/watch?v=-Lc21hhbAfE
https://www.youtube.com/watch?v=I3hpV0ZnaNE&ab_channel=IdealystTV
https://www.youtube.com/watch?v=OqNWrMU6mwQ&t=5s&ab_channel=AisKacang
https://www.youtube.com/watch?v=OqNWrMU6mwQ&t=5s&ab_channel=AisKacang
https://www.youtube.com/watch?v=QDL74a4Uibk&ab_channel=AisKacang
https://www.youtube.com/watch?v=t_6UO2Vi5nk&ab_channel=AisKacang
https://www.youtube.com/watch?v=f6H3ZsFy9dY&ab_channel=AisKacang
https://www.youtube.com/watch?v=vjuFRuWXabc&ab_channel=AisKacang
https://www.youtube.com/watch?v=SP77L5Z9JNw&ab_channel=AisKacang
https://www.youtube.com/watch?v=GpGFouytwFA&ab_channel=AisKacang
https://www.youtube.com/watch?v=2to1DeBZyiE&ab_channel=AisKacang
https://www.youtube.com/watch?v=-GZZGYGPJGY&t=192s&ab_channel=AisKacang
https://www.youtube.com/watch?v=VlmIyZllgz0
https://www.youtube.com/watch?v=bRUjmsnE5ZU&ab_channel=PurePixels
https://www.youtube.com/watch?v=BWU8IJBEjTk&ab_channel=PurePixels
https://www.youtube.com/watch?v=nPsVmw-GF4A&ab_channel=PurePixels
https://www.youtube.com/watch?v=bXVvXLHpmS8&t=87s&ab_channel=naddyepul
https://www.youtube.com/watch?v=cOuIF6LuVmw&ab_channel=THELAKI
https://www.youtube.com/watch?v=lcMZYdMnq-E&ab_channel=AmirArsad
https://www.youtube.com/watch?v=uCg5WeQ34rM&ab_channel=AmirArsad
https://www.youtube.com/watch?v=g95Rcydh0fs&t=5705s&ab_channel=MasBulan
https://www.youtube.com/watch?v=cBcTg1xyQgM&ab_channel=SUSUTVC
https://www.youtube.com/watch?v=aZcwqUZI2rA&t=989s&ab_channel=khairuljamain
https://www.youtube.com/watch?v=_Zsx149RtiU&ab_channel=khairuljamain
https://www.youtube.com/watch?v=tcqzySvke84&ab_channel=SkolarMalaysia
https://www.youtube.com/watch?v=wQXgHxeeZ2c&ab_channel=SkolarMalaysia
https://www.youtube.com/watch?v=lFr8JRu4Vj0
https://www.youtube.com/watch?v=bUXOSgBduvo
https://www.youtube.com/watch?v=GurzFbHVCI4
https://www.youtube.com/watch?v=YutaPWokeiE
https://www.youtube.com/watch?v=x0HhvgjUAjY
https://www.youtube.com/watch?v=pwFJaiaHBmA
https://www.youtube.com/watch?v=lyr89SJagaM
https://www.youtube.com/watch?v=9Rm1omhPz7c
https://www.youtube.com/watch?v=Vz3rCaQVarM
https://www.youtube.com/watch?v=TGSfdP5JiTs&ab_channel=khairuljamain
https://www.youtube.com/watch?v=Z1FpCckeSsk&ab_channel=khairuljamain
https://www.youtube.com/watch?v=cEIf11YM6iU
https://www.youtube.com/watch?v=riEwi1I6KKg
https://www.youtube.com/watch?v=BoX_hMvaXWM
https://www.youtube.com/watch?v=sSWZVKSsIlU
https://www.youtube.com/watch?v=7aIM0DhAQMc
https://www.youtube.com/watch?v=XJTfe7fVaGc&ab_channel=khairuljamain
https://www.youtube.com/watch?v=ysC2rYKKt20&ab_channel=SUSUTVC
https://www.youtube.com/watch?v=U7oaQ_tR8sQ
https://www.youtube.com/watch?v=pB-1y9Zsrj4
https://www.youtube.com/watch?v=eK3YtkMiqbM
https://www.youtube.com/watch?v=HGdtUpZ8qpQ
https://www.youtube.com/watch?v=F69FiR60K78
https://www.youtube.com/watch?v=sw5h9hlityE&t=876s&ab_channel=TanahMelayu
https://www.youtube.com/watch?v=4ZiC_Tg6hmU&t=768s&ab_channel=ONZZ
https://www.youtube.com/watch?v=C0CVcdzfkuQ&t=1281s&ab_channel=ONZZ
https://www.youtube.com/watch?v=C0CVcdzfkuQ&t=1281s&ab_channel=ONZZ
https://www.youtube.com/watch?v=WSHL2nyN-wA
https://www.youtube.com/watch?v=GSd5Tbo5fnE
https://www.youtube.com/watch?v=wEZaPeMCLL4
https://www.youtube.com/watch?v=2dlGt0FgmMo
https://www.youtube.com/watch?v=WoM2j5bwxVk
https://www.youtube.com/watch?v=KRDKcKiJbMc
https://www.youtube.com/watch?v=bGmzcVR8ms0
https://www.youtube.com/watch?v=OMk_Bb67lsk&t=860s
https://www.youtube.com/watch?v=u9kyC-ObWWM
https://www.youtube.com/watch?v=2EBDHR-7Rmw&t=1808s
https://www.youtube.com/watch?v=UT9VX6udGf8&t=951s&ab_channel=SterkProduction
https://www.youtube.com/watch?v=bMX6Nq8hveM&t=921s&ab_channel=SterkProduction
https://www.youtube.com/watch?v=7uHpElfSpes&t=843s&ab_channel=SterkProduction
https://www.youtube.com/watch?v=s0QwIMD123Y&t=2072s&ab_channel=SterkProduction
https://www.youtube.com/watch?v=_QsLUdHUytY&t=1056s&ab_channel=SterkProduction
https://www.youtube.com/watch?v=O_Mr_TG9aQs&t=546s&ab_channel=SterkProduction
https://www.youtube.com/watch?v=zbWJgXwcRes&ab_channel=SterkProduction
https://www.youtube.com/watch?v=hRUVMlq6PYk&ab_channel=SterkProduction
https://www.youtube.com/watch?v=-PfFTGk6-WM&ab_channel=THELAKI
https://www.youtube.com/watch?v=F1d0Huv54Ro&ab_channel=THELAKI
https://www.youtube.com/watch?v=Q1oB0OtOGkE&ab_channel=THELAKI
https://www.youtube.com/watch?v=2DqS7RigSLU
https://www.youtube.com/watch?v=V91NKEeCurI&t=1064s
https://www.youtube.com/watch?v=fcTQVSxcOf0&ab_channel=ERA
"""
videos = list(set(filter(None, videos.split('\n'))))
len(videos)
import youtube_dl
# +
import mp
from tqdm import tqdm
def loop(urls):
urls = urls[0]
ydl_opts = {
'format': 'bestaudio/best',
'postprocessors': [{
'key': 'FFmpegExtractAudio',
'preferredcodec': 'mp3',
'preferredquality': '192',
}],
'no-check-certificate': True
}
for i in tqdm(range(len(urls))):
try:
with youtube_dl.YoutubeDL(ydl_opts) as ydl:
ydl.download([urls[i]])
except:
pass
# -
import mp
mp.multiprocessing(videos, loop, cores = 12, returned = False)
# !mkdir malay-video
# !mv *.mp3 malay-video
|
data/semisupervised-malay/download-videos.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# http://ataspinar.com/2017/05/26/classification-with-scikit-learn/
# +
#This is a notebook containing the code of blogpost http://ataspinar.com/2017/05/26/classification-with-scikit-learn/
#Although I'll also give a short description in this notebook, for a full explanation you should read the blog.
# Lets import some modules for basic computation
import time
import pandas as pd
import numpy as np
import pickle
# Some modules for plotting and visualizing
import seaborn as sns
import matplotlib.pyplot as plt
from IPython.display import display
# And some Machine Learning modules from scikit-learn
from sklearn.decomposition import PCA
from sklearn.preprocessing import StandardScaler, LabelEncoder
from sklearn.linear_model import LogisticRegression
from sklearn.svm import SVC
from sklearn.neighbors import KNeighborsClassifier
from sklearn import tree
from sklearn.neural_network import MLPClassifier
from sklearn.neighbors import KNeighborsClassifier
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.gaussian_process.kernels import RBF
from sklearn.ensemble import RandomForestClassifier
from sklearn.naive_bayes import GaussianNB
#These Classifiers have been commented out because they take too long and do not give more accuracy as the other ones.
#from sklearn.ensemble import AdaBoostClassifier
#from sklearn.discriminant_analysis import QuadraticDiscriminantAnalysis
#from sklearn.gaussian_process import GaussianProcessClassifier
# +
dict_classifiers = {
"Logistic Regression": LogisticRegression(),
"Nearest Neighbors": KNeighborsClassifier(),
"Linear SVM": SVC(),
"Gradient Boosting Classifier": GradientBoostingClassifier(n_estimators=1000),
"Decision Tree": tree.DecisionTreeClassifier(),
"Random Forest": RandomForestClassifier(n_estimators=1000),
"Neural Net": MLPClassifier(alpha = 1),
"Naive Bayes": GaussianNB(),
#"AdaBoost": AdaBoostClassifier(),
#"QDA": QuadraticDiscriminantAnalysis(),
#"Gaussian Process": GaussianProcessClassifier()
}
def batch_classify(X_train, Y_train, X_test, Y_test, no_classifiers = 5, verbose = True):
"""
This method, takes as input the X, Y matrices of the Train and Test set.
And fits them on all of the Classifiers specified in the dict_classifier.
The trained models, and accuracies are saved in a dictionary. The reason to use a dictionary
is because it is very easy to save the whole dictionary with the pickle module.
Usually, the SVM, Random Forest and Gradient Boosting Classifier take quiet some time to train.
So it is best to train them on a smaller dataset first and
decide whether you want to comment them out or not based on the test accuracy score.
"""
dict_models = {}
for classifier_name, classifier in list(dict_classifiers.items())[:no_classifiers]:
t_start = time.clock()
classifier.fit(X_train, Y_train)
t_end = time.clock()
t_diff = t_end - t_start
train_score = classifier.score(X_train, Y_train)
test_score = classifier.score(X_test, Y_test)
dict_models[classifier_name] = {'model': classifier, 'train_score': train_score, 'test_score': test_score, 'train_time': t_diff}
if verbose:
print("trained {c} in {f:.2f} s".format(c=classifier_name, f=t_diff))
return dict_models
def label_encode(df, list_columns):
"""
This method one-hot encodes all column, specified in list_columns
"""
for col in list_columns:
le = LabelEncoder()
col_values_unique = list(df[col].unique())
le_fitted = le.fit(col_values_unique)
col_values = list(df[col].values)
le.classes_
col_values_transformed = le.transform(col_values)
df[col] = col_values_transformed
def expand_columns(df, list_columns):
for col in list_columns:
colvalues = df[col].unique()
for colvalue in colvalues:
newcol_name = "{}_is_{}".format(col, colvalue)
df.loc[df[col] == colvalue, newcol_name] = 1
df.loc[df[col] != colvalue, newcol_name] = 0
df.drop(list_columns, inplace=True, axis=1)
def get_train_test(df, y_col, x_cols, ratio):
"""
This method transforms a dataframe into a train and test set, for this you need to specify:
1. the ratio train : test (usually 0.7)
2. the column with the Y_values
"""
mask = np.random.rand(len(df)) < ratio
df_train = df[mask]
df_test = df[~mask]
Y_train = df_train[y_col].values
Y_test = df_test[y_col].values
X_train = df_train[x_cols].values
X_test = df_test[x_cols].values
return df_train, df_test, X_train, Y_train, X_test, Y_test
def display_dict_models(dict_models, sort_by='test_score'):
cls = [key for key in dict_models.keys()]
test_s = [dict_models[key]['test_score'] for key in cls]
training_s = [dict_models[key]['train_score'] for key in cls]
training_t = [dict_models[key]['train_time'] for key in cls]
df_ = pd.DataFrame(data=np.zeros(shape=(len(cls),4)), columns = ['classifier', 'train_score', 'test_score', 'train_time'])
for ii in range(0,len(cls)):
df_.loc[ii, 'classifier'] = cls[ii]
df_.loc[ii, 'train_score'] = training_s[ii]
df_.loc[ii, 'test_score'] = test_s[ii]
df_.loc[ii, 'train_time'] = training_t[ii]
display(df_.sort_values(by=sort_by, ascending=False))
def display_corr_with_col(df, col):
correlation_matrix = df.corr()
correlation_type = correlation_matrix[col].copy()
abs_correlation_type = correlation_type.apply(lambda x: abs(x))
desc_corr_values = abs_correlation_type.sort_values(ascending=False)
y_values = list(desc_corr_values.values)[1:]
x_values = range(0,len(y_values))
xlabels = list(desc_corr_values.keys())[1:]
fig, ax = plt.subplots(figsize=(8,8))
ax.bar(x_values, y_values)
ax.set_title('The correlation of all features with {}'.format(col), fontsize=20)
ax.set_ylabel('Pearson correlatie coefficient [abs waarde]', fontsize=16)
plt.xticks(x_values, xlabels, rotation='vertical')
plt.show()
# -
# # 1. The glass - dataset
filename_glass = '../datasets/glass.csv'
# +
df_glass = pd.read_csv(filename_glass)
print("This dataset has nrows, ncols: {}".format(df_glass.shape))
display(df_glass.head())
display(df_glass.describe())
# -
# ## 1.3 Classification
# +
y_col_glass = 'Type'
x_cols_glass = list(df_glass.columns.values)
x_cols_glass.remove(y_col_glass)
train_test_ratio = 0.7
df_train, df_test, X_train, Y_train, X_test, Y_test = get_train_test(df_glass, y_col_glass, x_cols_glass, train_test_ratio)
dict_models = batch_classify(X_train, Y_train, X_test, Y_test, no_classifiers = 8)
display_dict_models(dict_models)
# -
# # 2. Mushroom dataset (containing categorical data)
filename_mushrooms = '../datasets/mushrooms.csv'
df_mushrooms = pd.read_csv(filename_mushrooms)
display(df_mushrooms.head())
# ## 2.1 Preprocessing the Dataset
for col in df_mushrooms.columns.values:
print(col, df_mushrooms[col].unique())
# ### 2.1.1 Remove columns with only 1 value
for col in df_mushrooms.columns.values:
if len(df_mushrooms[col].unique()) <= 1:
print("Removing column {}, which only contains the value: {}".format(col, df_mushrooms[col].unique()[0]))
# ### 2.1.2 Handling columns with missing or incorrect values
# Some datasets contain missing values like NaN, null, NULL, '?', '??' etc
#
# It could be that all missing values are of type NaN, or that some columns contain NaN and other columns contain missing data in the form of '??'.
#
# It is up to your best judgement to decide what to do with these missing values.
# What is most effective, really depends on the type of data, the type of missing data and the ratio between missing data and non-missing data.
#
# - If the number of rows containing missing data is only a few percent of the total dataset, the best option could be to drop those rows.
#
# - If there is a column which contains almost all missing data, it will not have much added value and it might be best to drop that column.
#
# - It could be that a value not being filled in also is information which helps with the classification and it is best to leave it like it is.
#
# - etc
# ### 2.1.2.1 Drop rows with missing values
print("Number of rows in total: {}".format(df_mushrooms.shape[0]))
print("Number of rows with missing values in column 'stalk-root': {}".format(df_mushrooms[df_mushrooms['stalk-root'] == '?'].shape[0]))
df_mushrooms_dropped_rows = df_mushrooms[df_mushrooms['stalk-root'] != '?']
# ### 2.1.2.2 Drop column with more than X percent missing values
# +
drop_percentage = 0.8
df_mushrooms_dropped_cols = df_mushrooms.copy(deep=True)
df_mushrooms_dropped_cols.loc[df_mushrooms_dropped_cols['stalk-root'] == '?', 'stalk-root'] = np.nan
for col in df_mushrooms_dropped_cols.columns.values:
no_rows = df_mushrooms_dropped_cols[col].isnull().sum()
percentage = no_rows / df_mushrooms_dropped_cols.shape[0]
if percentage > drop_percentage:
del df_mushrooms_dropped_cols[col]
print("Column {} contains {} missing values. This is {} percent. Dropping this column.".format(col, no_rows, percentage))
# -
# ### 2.1.2.3 Fill missing values with zero / -1
df_mushrooms_zerofill = df_mushrooms.copy(deep = True)
df_mushrooms_zerofill.loc[df_mushrooms_zerofill['stalk-root'] == '?', 'stalk-root'] = np.nan
df_mushrooms_zerofill.fillna(0, inplace=True)
# ### 2.1.2.4 Fill missing values with backward fill
df_mushrooms_bfill = df_mushrooms.copy(deep = True)
df_mushrooms_bfill.loc[df_mushrooms_bfill['stalk-root'] == '?', 'stalk-root'] = np.nan
df_mushrooms_bfill.fillna(method='bfill', inplace=True)
# ### 2.1.2.5 Fill missing values with forward fill
df_mushrooms_ffill = df_mushrooms.copy(deep = True)
df_mushrooms_ffill.loc[df_mushrooms_ffill['stalk-root'] == '?', 'stalk-root'] = np.nan
df_mushrooms_ffill.fillna(method='ffill', inplace=True)
# ## 2.2 Handling columns with categorical data
# When it comes to columns with categorical data, you can do two things.
#
# - 1) One-hot encode the columns such that they are converted to numerical values.
# - 2) Expand the column into N different columns containing binary values.
#
#
#
# ** Example: **
#
# Let assume that we have a column called 'FRUIT' with unique values ['ORANGE', 'APPLE', PEAR'].
# - In the first case it would be converted to the unique values [0, 1, 2]
# - In the second case it would be converted into three different columns called ['FRUIT_IS_ORANGE', 'FRUIT_IS_APPLE', 'FRUIT_IS_PEAR'] and after this the original column 'FRUIT' would be deleted. The three new columns contain the values 1 or 0 depending on the value of the original column.
#
#
# When using the first method, you should pay attention to the fact that some classifiers will try to make sense of the numerical value of the one-hot encoded column. For example the Nearest neighbour algorithm assumes that the value 1 is closer to 0 than the value 2. But the numerical values have no meaning in the case of one-hot encoded columns (an APPLE is not closer to an ORANGE than a PEAR is.). And the results therefore can be misleading.
#
#
# ### 2.2.1 One-Hot encoding the columns with categorical data
df_mushrooms_ohe = df_mushrooms.copy(deep=True)
to_be_encoded_cols = df_mushrooms_ohe.columns.values
label_encode(df_mushrooms_ohe, to_be_encoded_cols)
display(df_mushrooms_ohe.head())
# +
## Now lets do the same thing for the other dataframes
df_mushrooms_dropped_rows_ohe = df_mushrooms_dropped_rows.copy(deep = True)
df_mushrooms_zerofill_ohe = df_mushrooms_zerofill.copy(deep = True)
df_mushrooms_bfill_ohe = df_mushrooms_bfill.copy(deep = True)
df_mushrooms_ffill_ohe = df_mushrooms_ffill.copy(deep = True)
label_encode(df_mushrooms_dropped_rows_ohe, to_be_encoded_cols)
label_encode(df_mushrooms_zerofill_ohe, to_be_encoded_cols)
label_encode(df_mushrooms_bfill_ohe, to_be_encoded_cols)
label_encode(df_mushrooms_ffill_ohe, to_be_encoded_cols)
# -
# ### 2.2.2 Expanding the columns with categorical data
# +
y_col = 'class'
to_be_expanded_cols = list(df_mushrooms.columns.values)
to_be_expanded_cols.remove(y_col)
df_mushrooms_expanded = df_mushrooms.copy(deep=True)
label_encode(df_mushrooms_expanded, [y_col])
expand_columns(df_mushrooms_expanded, to_be_expanded_cols)
display(df_mushrooms_expanded.head())
# +
## Now lets do the same thing for all other dataframes
df_mushrooms_dropped_rows_expanded = df_mushrooms_dropped_rows.copy(deep = True)
df_mushrooms_zerofill_expanded = df_mushrooms_zerofill.copy(deep = True)
df_mushrooms_bfill_expanded = df_mushrooms_bfill.copy(deep = True)
df_mushrooms_ffill_expanded = df_mushrooms_ffill.copy(deep = True)
label_encode(df_mushrooms_dropped_rows_expanded, [y_col])
label_encode(df_mushrooms_zerofill_expanded, [y_col])
label_encode(df_mushrooms_bfill_expanded, [y_col])
label_encode(df_mushrooms_ffill_expanded, [y_col])
expand_columns(df_mushrooms_dropped_rows_expanded, to_be_expanded_cols)
expand_columns(df_mushrooms_zerofill_expanded, to_be_expanded_cols)
expand_columns(df_mushrooms_bfill_expanded, to_be_expanded_cols)
expand_columns(df_mushrooms_ffill_expanded, to_be_expanded_cols)
# -
# ## 2.4 Classifying the dataset
# We have seen that there are two different ways to handle columns with categorical data, and many different ways to handle missing values.
#
# Since computation power is cheap, it is easy to try out all of these ways on all of the classifiers present in the scikit-learn.
#
# After we have seen which method and which classifier has the highest accuracy initially we can continue in that direction.
dict_dataframes = {
"df_mushrooms_ohe": df_mushrooms_ohe,
"df_mushrooms_dropped_rows_ohe": df_mushrooms_dropped_rows_ohe,
"df_mushrooms_zerofill_ohe": df_mushrooms_zerofill_ohe,
"df_mushrooms_bfill_ohe": df_mushrooms_bfill_ohe,
"df_mushrooms_ffill_ohe": df_mushrooms_ffill_ohe,
"df_mushrooms_expanded": df_mushrooms_expanded,
"df_mushrooms_dropped_rows_expanded": df_mushrooms_dropped_rows_expanded,
"df_mushrooms_zerofill_expanded": df_mushrooms_zerofill_expanded,
"df_mushrooms_bfill_expanded": df_mushrooms_bfill_expanded,
"df_mushrooms_ffill_expanded": df_mushrooms_ffill_expanded
}
# +
y_col = 'class'
train_test_ratio = 0.7
for df_key, df in dict_dataframes.items():
x_cols = list(df.columns.values)
x_cols.remove(y_col)
df_train, df_test, X_train, Y_train, X_test, Y_test = get_train_test(df, y_col, x_cols, train_test_ratio)
dict_models = batch_classify(X_train, Y_train, X_test, Y_test, no_classifiers = 8, verbose=False)
print()
print(df_key)
display_dict_models(dict_models)
print("-------------------------------------------------------")
# -
# ## 2.5 Improving upon the Classifier: hyperparameter optimization
# After you have determined with a quick and dirty method type of filling missing values and which classifier performs best for your dataset, you can improve upon the Classifier by optimizing its hyperparameters.
#
# Since the mushroom dataset already has a high accuracy on the test set, there is not much to improve upon. So demonstrate hyperparameter optimization we'll use the glass dataset again.
# +
GDB_params = {
'n_estimators': [100, 500, 1000],
'learning_rate': [0.5, 0.1, 0.01, 0.001],
'criterion': ['friedman_mse', 'mse', 'mae']
}
df_train, df_test, X_train, Y_train, X_test, Y_test = get_train_test(df_glass, y_col_glass, x_cols_glass, 0.6)
for n_est in GDB_params['n_estimators']:
for lr in GDB_params['learning_rate']:
for crit in GDB_params['criterion']:
clf = GradientBoostingClassifier(n_estimators=n_est,
learning_rate = lr,
criterion = crit)
clf.fit(X_train, Y_train)
train_score = clf.score(X_train, Y_train)
test_score = clf.score(X_test, Y_test)
print("For ({}, {}, {}) - train, test score: \t {:.5f} \t-\t {:.5f}".format(n_est, lr, crit[:4], train_score, test_score))
# -
# # 3. Understanding complex datasets
# ## 3.1 Correlation Matrix
# Some datasets contain a lot of features / columns, and it is not immediatly clear which of these features are helping with the Classification / Regression, and which of these features are only adding more noise.
#
# To have a better understanding of this, you could make a correlation matrix of the data, and plot all features by descending order of correlation value.
correlation_matrix = df_glass.corr()
plt.figure(figsize=(10,8))
ax = sns.heatmap(correlation_matrix, vmax=1, square=True, annot=True,fmt='.2f', cmap ='GnBu', cbar_kws={"shrink": .5}, robust=True)
plt.title('Correlation matrix between the features', fontsize=20)
plt.show()
display_corr_with_col(df_glass, 'Type')
# ## 3.2 Cumulative Explained Variance
#
# The Cumulative explained variance shows how much of the variance is captures by the first x features.
#
# Below we can see that the first 4 features (i.e. the four features with the largest correlation) already capture 90% of the variance.
#
# If you have low accuracy values for your Regression / Classification model, you could decide to stepwise remove the features with the lowest correlation, (or stepwise add features with the highest correlation).
# +
X = df_glass[x_cols_glass].values
X_std = StandardScaler().fit_transform(X)
pca = PCA().fit(X_std)
var_ratio = pca.explained_variance_ratio_
components = pca.components_
#print(pca.explained_variance_)
plt.plot(np.cumsum(var_ratio))
plt.xlim(0,9,1)
plt.xlabel('Number of Features', fontsize=16)
plt.ylabel('Cumulative explained variance', fontsize=16)
plt.show()
# -
# ## 3.3 Pairwise relationships between the features
#
# In addition to the correlation matrix, you can plot the pairwise relationships between the features, to see **how** these features are correlated.
ax = sns.pairplot(df_glass, hue='Type')
plt.title('Pairwise relationships between the features')
plt.show()
|
notebooks/scikit_classification.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import tensorflow as tf
from tensorflow.keras.layers import Dense, Flatten
from tensorflow.keras import Model
import matplotlib.pyplot as plt
import numpy as np
# # Download a dataset
# (x_train, y_train), (x_test, y_test) = tf.keras.datasets.mnist.load_data()
# # Batch and shuffle the data
# train_ds = tf.data.Dataset.from_tensor_slices(
# (x_train.astype('float32') / 255, y_train)).shuffle(1024).batch(32)
# test_ds = tf.data.Dataset.from_tensor_slices(
# (x_test.astype('float32') / 255, y_test)).batch(32)
# +
from nus_wide_data_util import get_labeled_data, get_top_k_labels
class_num = 5
top_k = get_top_k_labels('', top_k=class_num)
print(top_k)
data_X_image, data_X_text, data_Y = get_labeled_data('', top_k, 60000)
print(type(data_X_image), type(data_X_text), type(data_Y))
train_num = 50000
x_train, x_test, y_train, y_test = (np.array(data_X_image[:train_num]).astype('float32'), np.array(data_X_text[:train_num]).astype('float32')), \
(np.array(data_X_image[train_num:]).astype('float32'), np.array(data_X_text[train_num:]).astype('float32')), \
np.array(data_Y[:train_num]).astype('float32'), np.array(data_Y[train_num:]).astype('float32')
# Batch and shuffle the data
train_ds = tf.data.Dataset.from_tensor_slices(
(x_train, y_train)).shuffle(1024).batch(32)
test_ds = tf.data.Dataset.from_tensor_slices(
(x_test, y_test)).batch(32)
# -
np.sum(y_test, axis=0)
class VFLPassiveModel(Model):
def __init__(self):
super(VFLPassiveModel, self).__init__()
self.flatten = Flatten()
self.d1 = Dense(class_num, name="dense1")
def call(self, x):
x = self.flatten(x)
return self.d1(x)
# +
import numpy as np
class VFLActiveModel(Model):
def __init__(self):
super(VFLActiveModel, self).__init__()
self.added = tf.keras.layers.Add()
def call(self, x):
x = self.added(x)
return tf.keras.layers.Softmax()(x)
# +
# normal training
passive_model_image = VFLPassiveModel()
passive_model_text = VFLPassiveModel()
active_model = VFLActiveModel()
loss_object = tf.keras.losses.CategoricalCrossentropy()
optimizer = tf.keras.optimizers.Adam()
train_loss = tf.keras.metrics.Mean(name='train_loss')
train_accuracy = tf.keras.metrics.CategoricalAccuracy(name='train_accuracy')
test_loss = tf.keras.metrics.Mean(name='test_loss')
test_accuracy = tf.keras.metrics.CategoricalAccuracy(name='test_accuracy')
EPOCHS = 5
for epoch in range(EPOCHS):
# For each batch of images and labels
for (images, texts), labels in train_ds:
with tf.GradientTape() as passive_tape:
# passive_model sends passive_output to active_model
passive_image_output = passive_model_image(images)
passive_text_output = passive_model_text(texts)
with tf.GradientTape() as active_tape:
active_tape.watch(passive_image_output)
active_tape.watch(passive_text_output)
active_output = active_model([passive_image_output, passive_text_output])
loss = loss_object(labels, active_output)
# active_model sends passive_output_gradients back to passive_model
[active_image_gradients, active_text_gradients] = active_tape.gradient(loss, [passive_image_output, passive_text_output])
passive_image_loss = tf.multiply(passive_image_output, active_image_gradients.numpy())
passive_text_loss = tf.multiply(passive_text_output, active_text_gradients.numpy())
[passive_image_gradients, passive_text_gradients] = \
passive_tape.gradient([passive_image_loss, passive_text_loss], \
[passive_model_image.trainable_variables, passive_model_text.trainable_variables])
optimizer.apply_gradients(zip(passive_image_gradients, passive_model_image.trainable_variables))
optimizer.apply_gradients(zip(passive_text_gradients, passive_model_text.trainable_variables))
train_loss(loss)
train_accuracy(labels, active_output)
for (test_images, test_texts), test_labels in test_ds:
passive_output = [passive_model_image(test_images), passive_model_text(test_texts)]
active_output = active_model(passive_output)
t_loss = loss_object(test_labels, active_output)
test_loss(t_loss)
test_accuracy(test_labels, active_output)
template = 'Epoch {}, Loss: {}, Accuracy: {}, Test Loss: {}, Test Accuracy: {}'
print(template.format(epoch+1,
train_loss.result(),
train_accuracy.result()*100,
test_loss.result(),
test_accuracy.result()*100))
# Reset the metrics for the next epoch
train_loss.reset_states()
train_accuracy.reset_states()
test_loss.reset_states()
test_accuracy.reset_states()
# +
import numpy as np
def get_possioned_gradients(passive_output_gradients, passive_output, N, class1, class2, alpha = 1.0):
#passive_output_gradients -= passive_output
attack_mat = np.eye(N, dtype='float32')
attack_mat[:, class2] += attack_mat[:, class1]*alpha
attack_mat[:, class1] -= attack_mat[:, class1]*alpha
passive_output_gradients = tf.matmul(passive_output_gradients, attack_mat)
#passive_output_gradients += passive_output
return passive_output_gradients
# +
# backdoor training
passive_model_image = VFLPassiveModel()
passive_model_text = VFLPassiveModel()
active_model = VFLActiveModel()
loss_object = tf.keras.losses.CategoricalCrossentropy()
optimizer = tf.keras.optimizers.Adam()
optimizer_attack = tf.keras.optimizers.Adam(lr=0.1)
train_loss = tf.keras.metrics.Mean(name='train_loss')
train_accuracy = tf.keras.metrics.CategoricalAccuracy(name='train_accuracy')
test_loss = tf.keras.metrics.Mean(name='test_loss')
test_accuracy = tf.keras.metrics.CategoricalAccuracy(name='test_accuracy')
EPOCHS = 5
for epoch in range(EPOCHS):
# For each batch of images and labels
for (images, texts), labels in train_ds:
with tf.GradientTape() as passive_tape:
# passive_model sends passive_output to active_model
passive_image_output = passive_model_image(images)
passive_text_output = passive_model_text(texts)
with tf.GradientTape() as active_tape:
active_tape.watch(passive_image_output)
active_tape.watch(passive_text_output)
active_output = active_model([passive_image_output, passive_text_output])
loss = loss_object(labels, active_output)
# active_model sends passive_output_gradients back to passive_model
[active_image_gradients, active_text_gradients] = active_tape.gradient(loss, [passive_image_output, passive_text_output])
active_image_gradients = get_possioned_gradients(active_image_gradients, passive_image_output, 5, 3, 4)
#active_text_gradients = get_possioned_gradients(active_text_gradients, passive_text_output, 5, 3, 4)
passive_image_loss = tf.multiply(passive_image_output, active_image_gradients.numpy())
passive_text_loss = tf.multiply(passive_text_output, active_text_gradients.numpy())
[passive_image_gradients, passive_text_gradients] = \
passive_tape.gradient([passive_image_loss, passive_text_loss], \
[passive_model_image.trainable_variables, passive_model_text.trainable_variables])
#optimizer.apply_gradients(zip(passive_image_gradients, passive_model_image.trainable_variables))
optimizer_attack.apply_gradients(zip(passive_image_gradients, passive_model_image.trainable_variables))
optimizer.apply_gradients(zip(passive_text_gradients, passive_model_text.trainable_variables))
#optimizer_attack.apply_gradients(zip(passive_text_gradients, passive_model_text.trainable_variables))
train_loss(loss)
train_accuracy(labels, active_output)
for (test_images, test_texts), test_labels in test_ds:
passive_output = [passive_model_image(test_images), passive_model_text(test_texts)]
active_output = active_model(passive_output)
t_loss = loss_object(test_labels, active_output)
test_loss(t_loss)
test_accuracy(test_labels, active_output)
template = 'Epoch {}, Loss: {}, Accuracy: {}, Test Loss: {}, Test Accuracy: {}'
print(template.format(epoch+1,
train_loss.result(),
train_accuracy.result()*100,
test_loss.result(),
test_accuracy.result()*100))
# Reset the metrics for the next epoch
train_loss.reset_states()
train_accuracy.reset_states()
test_loss.reset_states()
test_accuracy.reset_states()
# +
import matplotlib.pyplot as plt
import numpy as np
label_test = np.argmax(y_test, axis=1)
(image_test, text_test) = x_test
print(image_test.shape, text_test.shape)
image_val = image_test[label_test==1]
text_val = text_test[label_test==1]
print(image_val.shape, text_val.shape)
y_val = y_test[label_test==1]
passive_output = [passive_model_image(image_val), passive_model_text(text_val)]
active_output = active_model(passive_output)
output_distribution = np.sum(active_output, axis=0)
print(output_distribution)
n = 5
X = np.arange(n)
plt.bar(X, output_distribution)
plt.show()
# +
class MyLinearModel(Model):
def __init__(self):
super(MyLinearModel, self).__init__()
self.flatten = Flatten()
self.d1 = Dense(class_num, activation='softmax', name="dense1")
def call(self, x):
x = self.flatten(x)
return self.d1(x)
model = MyLinearModel()
# +
loss_object = tf.keras.losses.SparseCategoricalCrossentropy()
optimizer = tf.keras.optimizers.Adam()
train_loss = tf.keras.metrics.Mean(name='train_loss')
train_accuracy = tf.keras.metrics.SparseCategoricalAccuracy(name='train_accuracy')
test_loss = tf.keras.metrics.Mean(name='test_loss')
test_accuracy = tf.keras.metrics.SparseCategoricalAccuracy(name='test_accuracy')
EPOCHS = 5
for epoch in range(EPOCHS):
# For each batch of images and labels
for images, labels in train_ds:
with tf.GradientTape() as tape:
predictions = model(images)
loss = loss_object(labels, predictions)
gradients = tape.gradient(loss, model.trainable_variables)
print(gradients[0].shape, gradients[1].shape)
optimizer.apply_gradients(zip(gradients, model.trainable_variables))
train_loss(loss)
train_accuracy(labels, predictions)
for test_images, test_labels in test_ds:
predictions = model(test_images)
t_loss = loss_object(test_labels, predictions)
test_loss(t_loss)
test_accuracy(test_labels, predictions)
template = 'Epoch {}, Loss: {}, Accuracy: {}, Test Loss: {}, Test Accuracy: {}'
print(template.format(epoch+1,
train_loss.result(),
train_accuracy.result()*100,
test_loss.result(),
test_accuracy.result()*100))
# Reset the metrics for the next epoch
train_loss.reset_states()
train_accuracy.reset_states()
test_loss.reset_states()
test_accuracy.reset_states()
# -
with tf.GradientTape() as tape:
a = tf.constant(2.)
b = tf.constant(1.)
tape.watch(a)
tape.watch(b)
c = tf.multiply(a, b)
g = tape.gradient(c, [a, b])
print(g)
|
vertical_backdoor_attack.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Implement a Singly Linked List
#
# For this interview problem, create a node class and show how it can be used to create a Singly Linked List
class LinkedListNode(object):
def __init__(self, value):
self.value = value
self.nextnode = None
# +
# Create a Singly Linked List here
# -
# # Test Your Solution
#
# Note that there is no test for this solution (because it would give away the answer structure).
#
# Check out the Implement a Linked List Solution Notebook for the answer to this interview problem, as well as the answer for the implementation of a doubly linked list.
a = LinkedListNode(1)
b = LinkedListNode(2)
c = LinkedListNode(3)
a.nextnode = b
b.nextnode = c
a
a.value
a.nextnode.value
|
Linked List Interview Problems /Implement a Singly Linked List.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/johnanisere/30-seconds-of-css/blob/master/gaussian_code_exercise.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="8k3p5S4AJ1Rl" colab_type="text"
# # Gaussian Code Exercise
#
#
# The 'numbers.txt' file is read in by the read_data_file() method.
# + id="a4xzb3d2J1Rn" colab_type="code" colab={}
import math
import matplotlib.pyplot as plt
class Gaussian():
""" Gaussian distribution class for calculating and
visualizing a Gaussian distribution.
Attributes:
mean (float) representing the mean value of the distribution
stdev (float) representing the standard deviation of the distribution
data_list (list of floats) a list of floats extracted from the data file
"""
def __init__(self, mu = 0, sigma = 1):
self.mean = mu
self.stdev = sigma
self.data = []
def calculate_mean(self):
"""Method to calculate the mean of the data set.
Args:
None
Returns:
float: mean of the data set
"""
#TODO: Calculate the mean of the data set. Remember that the data set is stored in self.data
# Change the value of the mean attribute to be the mean of the data set
# Return the mean of the data set
def calculate_stdev(self, sample=True):
"""Method to calculate the standard deviation of the data set.
Args:
sample (bool): whether the data represents a sample or population
Returns:
float: standard deviation of the data set
"""
# TODO:
# Calculate the standard deviation of the data set
#
# The sample variable determines if the data set contains a sample or a population
# If sample = True, this means the data is a sample.
# Keep the value of sample in mind for calculating the standard deviation
#
# Make sure to update self.stdev and return the standard deviation as well
def read_data_file(self, file_name, sample=True):
"""Method to read in data from a txt file. The txt file should have
one number (float) per line. The numbers are stored in the data attribute.
After reading in the file, the mean and standard deviation are calculated
Args:
file_name (string): name of a file to read from
Returns:
None
"""
# This code opens a data file and appends the data to a list called data_list
with open(file_name) as file:
data_list = []
line = file.readline()
while line:
data_list.append(int(line))
line = file.readline()
file.close()
# TODO:
# Update the self.data attribute with the data_list
# Update self.mean with the mean of the data_list.
# You can use the calculate_mean() method with self.calculate_mean()
# Update self.stdev with the standard deviation of the data_list. Use the
# calcaulte_stdev() method.
def plot_histogram(self):
"""Method to output a histogram of the instance variable data using
matplotlib pyplot library.
Args:
None
Returns:
None
"""
# TODO: Plot a histogram of the data_list using the matplotlib package.
# Be sure to label the x and y axes and also give the chart a title
def pdf(self, x):
"""Probability density function calculator for the gaussian distribution.
Args:
x (float): point for calculating the probability density function
Returns:
float: probability density function output
"""
# TODO: Calculate the probability density function of the Gaussian distribution
# at the value x. You'll need to use self.stdev and self.mean to do the calculation
def plot_histogram_pdf(self, n_spaces = 50):
"""Method to plot the normalized histogram of the data and a plot of the
probability density function along the same range
Args:
n_spaces (int): number of data points
Returns:
list: x values for the pdf plot
list: y values for the pdf plot
"""
#TODO: Nothing to do for this method. Try it out and see how it works.
# + id="2iIyIpPzJ1Rq" colab_type="code" colab={}
# Unit tests to check your solution
import unittest
class TestGaussianClass(unittest.TestCase):
def setUp(self):
self.gaussian = Gaussian(25, 2)
def test_initialization(self):
self.assertEqual(self.gaussian.mean, 25, 'incorrect mean')
self.assertEqual(self.gaussian.stdev, 2, 'incorrect standard deviation')
def test_pdf(self):
self.assertEqual(round(self.gaussian.pdf(25), 5), 0.19947,\
'pdf function does not give expected result')
def test_meancalculation(self):
self.gaussian.read_data_file('numbers.txt', True)
self.assertEqual(self.gaussian.calculate_mean(),\
sum(self.gaussian.data) / float(len(self.gaussian.data)), 'calculated mean not as expected')
def test_stdevcalculation(self):
self.gaussian.read_data_file('numbers.txt', True)
self.assertEqual(round(self.gaussian.stdev, 2), 92.87, 'sample standard deviation incorrect')
self.gaussian.read_data_file('numbers.txt', False)
self.assertEqual(round(self.gaussian.stdev, 2), 88.55, 'population standard deviation incorrect')
tests = TestGaussianClass()
tests_loaded = unittest.TestLoader().loadTestsFromModule(tests)
unittest.TextTestRunner().run(tests_loaded)
# + id="qzyYpxggJ1Ru" colab_type="code" colab={}
|
gaussian_code_exercise.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python36
# ---
# # Semantic Segmentation with Keras
#
# In this exercise, you'll use the U-Net network to perform binary classification and segmentation for images of planes.
#
# > **Important**: Using the U-Net model is resource-intensive. before running the code in this notebook, shut down all other notebooks in this library (In each open notebook other than this one, on the **File** menu, click **Close and Halt**). If you experience and Out-of-Memory (OOM) error when running code in this notebook, shut down this entire library, and then reopen it and open only this notebook.
# ## Install Keras
#
# To begin with, we'll install the latest version of Keras.
# !pip install --upgrade keras
# ## Explore the Training Data
#
# The training data for a U-Net model consists of two kinds of input:
#
# - **Image files**: The images that represent the *features* on which we want to train the model.
# - **Mask files**: Images of the object masks that the network will be trained to predict - these are the *labels*.
#
# In this example, we're going to use U-Net for binary classification of airplanes images, so there's only one class of object - and therefore one class of mask. We've deliberately made this example as simple as possible, partly to make it easier to understand what's going on, and partly to ensure it can be run in a resource-constrained environment.
#
# Let's take a look at the training images and masks:
# +
import os
from matplotlib import pyplot as plt
import skimage.io as io
import numpy as np
# %matplotlib inline
fig = plt.figure(figsize=(12, 60))
train_dir = '../../data/segmentation/train'
image_dir = os.path.join(train_dir,"image/plane")
mask_dir = os.path.join(train_dir,"mask/plane")
files = os.listdir(image_dir)
rows = len(files)
cell = 0
for file in files:
cell += 1
# Open the image and mask files
img_path = os.path.join(image_dir, file)
img = io.imread(img_path, as_gray = True)
mask_path = os.path.join(mask_dir, file)
mask = io.imread(mask_path, as_gray = True)
# plot the image
a=fig.add_subplot(rows,3,cell)
imgplot=plt.imshow(img, "gray")
cell += 1
# plot the mask
a=fig.add_subplot(rows,3,cell)
imgplot=plt.imshow(mask, "gray")
cell += 1
# Plot them overlaid
a=fig.add_subplot(rows,3,cell)
imgplot=plt.imshow(img, "gray")
imgplot=plt.imshow(mask, "gray", alpha=0.4)
plt.show()
# -
# ## Import the U-Net Code
#
# The code to implement U-Net is provided in two python files:
#
# - **model.py**: This file contains the code that implements the U-Net model
# - **data.py**: This file contains functions to help load and prepare training data.
#
# > **Tip**: You should explore the code in these files to get a better understanding of the way the model works.
#
from unet_keras.data import *
from unet_keras.model import *
# The ouput from the code above shows the training images with their corresponding mask labels, and finally the mask overlaid on the image so you can clearly see that the masks represent the pixels that belong to the plane objects in the images.
#
# > **Note**: We deliberately chose images in which the plane objects are clearly contrasted with the background to make it easier to train with a very small number of training images and a very small amount of training!
# ## Load the Training Data
# We have a very small number of training images, so we'll apply some data augmentation to randomly flip, zoom, shear, and otherwise transform the images for each batch.
data_gen_args = dict(rotation_range=0.2,
width_shift_range=0.05,
height_shift_range=0.05,
shear_range=0.05,
zoom_range=0.05,
horizontal_flip=True,
fill_mode='nearest')
train_images = trainGenerator(2,train_dir,'image','mask',data_gen_args,save_to_dir = None)
# ## Download the Model Weights
# The model has already been partially trained, so we'll download the trained weights as a starting point.
# !wget "https://aka.ms/unet.h5" -O ~/unet.h5
# ## Train the Model
#
# Now we're ready to train the U-Net model. We'll train it from the training generator we created, and save the model weights after each epoch if the loss has improved. In this example, to reduce the required compute resources we'll train it for just one epoch with minimal batches. In reality, you'd need to train the model over several epochs on a GPU-based computer.
#
# > _**Note**: This will take a while on a non-GPU machine - go get some coffee!_
model = unet()
home = os.path.expanduser("~")
weights_file = os.path.join(home, "unet.h5")
model.load_weights(weights_file)
model_checkpoint = ModelCheckpoint(weights_file, monitor='loss',verbose=1, save_best_only=True)
model.fit_generator(train_images,steps_per_epoch=1,epochs=1,callbacks=[model_checkpoint])
# ## Test the Trained Model
#
# OK, let's see how well our trained model does with some images of airplanes it hasn't seen.
# +
import os
from matplotlib import pyplot as plt
import skimage.io as io
import numpy as np
# %matplotlib inline
model = unet()
model.load_weights(weights_file)
fig = plt.figure(figsize=(12, 60))
test_dir = '../../data/segmentation/test'
files = os.listdir(test_dir)
rows = len(files)
cell = 0
for file in files:
cell += 1
# Open the file
img_path = os.path.join(test_dir, file)
img = io.imread(img_path, as_gray = True)
src_img = img
a=fig.add_subplot(rows,3,cell)
imgplot=plt.imshow(src_img, "gray")
cell += 1
img = np.reshape(img,img.shape+(1,))
mask_predictions = model.predict([[img]])
mask = mask_predictions[0]
img_mask = mask.reshape(mask.shape[0], mask.shape[1])
a=fig.add_subplot(rows,3,cell)
imgplot=plt.imshow(img_mask, "gray")
cell += 1
a=fig.add_subplot(rows,3,cell)
imgplot=plt.imshow(src_img, "gray")
imgplot=plt.imshow(img_mask, "binary", alpha=0.6)
plt.show()
# -
# It's not fantastic, largely because we used such a small amount of data; but hopefully it serves to demonstrate the principles of semantic segmentation with U-Net.
#
# ## Acknowledgements and Citations
#
# The U-Net architecture is documented by its inventors (<NAME>, <NAME>, and <NAME>), at https://arxiv.org/abs/1505.04597.
#
# The Keras implementation of U-Net used in this exercise is based on zhixuhao's work at https://github.com/zhixuhao/unet, with some simplifications.
#
# The data used in this exercise includes images adapted from the PASCAL Visual Object Classes Challenge (VOC2007) dataset at http://host.robots.ox.ac.uk/pascal/VOC/voc2007/.
#
#
# @misc{pascal-voc-2007,
# author = "<NAME> <NAME> <NAME>.",
# title = "The {PASCAL} {V}isual {O}bject {C}lasses {C}hallenge 2007 {(VOC2007)} {R}esults",
# howpublished = "http://www.pascal-network.org/challenges/VOC/voc2007/workshop/index.html"}
#
#
|
Mod04/02-Unet/U-Net (Keras).ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: doggo
# language: python
# name: doggo
# ---
import os
print(os.getcwd())
def update_working_directory():
from pathlib import Path
p = Path(os.getcwd()).parents[0]
os.chdir(p)
print(p)
update_working_directory()
# # Importing the required libraries
import numpy as np
import pandas as pd
import gym
import time
import math
import statistics
from tqdm import tqdm
import random
from collections import defaultdict
import matplotlib.pyplot as plt
import dill
# # Building the environment
# ## Parameters specific to the environment
decimals_state = 2
gamma = 0.95 # discount for future rewards (also called decay factor)
# ## Observation & Action spaces
# +
n_states = (10**decimals_state+1)**3
action_dict = {
0: 'NO ACTION',
1: 'WALKING',
2: 'EATING',
3: 'PLAYING'
}
n_actions= len(action_dict)
print(n_states, n_actions)
# -
# ## Reset
def get_state_id(dog_state):
return '{:01.4f}_{:01.4f}_{:01.4f}_{}'.format(
dog_state['food'], dog_state['fat'], dog_state['affection'], dog_state['can_action_be_taken'])
def env_reset():
dog_state = {
'food': 0.5,
'fat': 0,
'affection': 0.5,
'last_action_taken': 0,
'minutes_since_last_action': 0,
'can_action_be_taken': True
}
dog_state['state_id'] = get_state_id(dog_state)
return dog_state
env_reset()
# ## Next state
# +
WALKING_TIME = 15
EATING_TIME = 1
PLAYING_TIME = 4
food_consumption_rate = 1.0 / (30 * 3600)
affection_consumption_rate = 1.0 / (50 * 3600)
walking_fat_converge_rate = 0.2
walking_affection_converge_rate = 0.4
playing_fat_converge_rate = 0.1
playing_affection_converge_rate = 0.20
eating_food_increase = 0.6
eating_fat_increase = 0.25
# -
def round_up(n, decimals=0):
multiplier = 10 ** decimals
return math.ceil(n * multiplier) / multiplier
def round_down(n, decimals=0):
multiplier = 10 ** decimals
return math.floor(n * multiplier) / multiplier
# +
def apply_decreasing_rate(value: float, rate: float) -> float:
"""
Apply a decreasing rate to a value
:param value: current value
:param rate: per second
:return: updated value
"""
return value - (60 * rate)
def converge(value: float, target: float, ratio: float) -> float:
diff: float = (target - value) * ratio
return value + diff
def update_food(dog_state):
update_food = apply_decreasing_rate(dog_state['food'], food_consumption_rate)
return round_down(max(0.0, update_food), decimals=decimals_state)
def update_fat(dog_state):
update_fat = dog_state['fat']
return update_fat
def update_affection(dog_state):
update_affection = apply_decreasing_rate(dog_state['affection'], affection_consumption_rate)
return round_down(max(0.0, update_affection), decimals=decimals_state)
def update_if_walking(dog_state):
update_fat = round_down(converge(dog_state['fat'], 0.0, walking_fat_converge_rate), decimals=decimals_state)
update_affection = round_up(converge(dog_state['affection'], 1.0, walking_affection_converge_rate), decimals=decimals_state)
return (update_fat, update_affection)
def update_if_feeding(dog_state):
update_food = round_up(min(dog_state['food'] + eating_food_increase, 1.0), decimals=decimals_state)
update_fat = round_up(min(dog_state['fat'] + eating_fat_increase, 1.0), decimals=decimals_state)
return (update_food, update_fat)
def update_if_playing(dog_state):
update_fat = round_down(converge(dog_state['fat'], 0.0, playing_fat_converge_rate), decimals=decimals_state)
update_affection = round_up(converge(dog_state['affection'], 1.0, playing_affection_converge_rate), decimals=decimals_state)
return (update_fat, update_affection)
def get_happiness(dog_state):
happiness = min(dog_state['food'], 1.0 - dog_state['fat'], dog_state['affection'])
return happiness
def update_done(dog_state):
happiness = get_happiness(dog_state)
return happiness <= 0.0
# -
# state2, reward1, done, info = env.step(action1)
def env_step(state1, action):
state2 = state1.copy()
reward_penalty = 0
# Affect of time
state2['food'] = update_food(state2)
state2['fat'] = update_fat(state2)
state2['affection'] = update_affection(state2)
state2['minutes_since_last_action'] += 1
# Applying action
if action != 0:
if state2['can_action_be_taken']:
reward_penalty += 0.1
state2['can_action_be_taken'] = False
state2['minutes_since_last_action'] = 0
state2['last_action_taken'] = action
else:
reward_penalty += 0.5
# Affect of actions
if (state2['last_action_taken'] == 1) & (state2['minutes_since_last_action'] == WALKING_TIME):
state2['fat'], state2['affection'] = update_if_walking(state2)
state2['can_action_be_taken'] = True
if (state2['last_action_taken'] == 2) & (state2['minutes_since_last_action'] == EATING_TIME):
state2['food'], state2['fat'] = update_if_feeding(state2)
state2['can_action_be_taken'] = True
if (state2['last_action_taken'] == 3) & (state2['minutes_since_last_action'] == PLAYING_TIME):
state2['fat'], state2['affection'] = update_if_playing(state2)
state2['can_action_be_taken'] = True
done = update_done(state2)
if done:
reward = -10
else:
reward = min(state2['food'], 1.0 - state2['fat'], state2['affection']) - reward_penalty
info = None
state2['state_id'] = get_state_id(state2)
return (state2, reward, done, info)
# ## Render
def env_render(dog_state, action, Q):
print(dog_state)
print(action)
print(Q[dog_state['state_id']])
# # Defining utility functions to be used in the learning process
# ## Initialising Q
def init_Q(n_actions, init_Q_type="ones"):
"""
@param n_actions the number of actions
@param type random, ones or zeros for the initialization
"""
if init_Q_type == "ones":
default_Q_values = np.ones(n_actions)
elif init_Q_type == "random":
default_Q_values = np.random.random(n_actions)
elif init_Q_type == "zeros":
default_Q_values = np.zeros(n_actions)
def get_default_Q_values():
return default_Q_values
return defaultdict(get_default_Q_values)
# ## Initialising N
def init_N(n_actions):
"""
@param n_actions the number of actions
"""
default_N_values = np.zeros(n_actions)
def get_default_N_values():
return default_N_values
return defaultdict(get_default_N_values)
# ## Choose an action
# Numpy generator
rng = np.random.default_rng() # Create a default Generator.
def select_best_action(Q_state):
winner = np.argwhere(Q_state == np.amax(Q_state))
winner_list = winner.flatten().tolist()
action = random.choice(winner_list)
return action
# ### $\epsilon$-Greedy
def epsilon_greedy(Q, state_id, n_actions, epsilon):
"""
@param Q Q values {state, action} -> value
@param epsilon for exploration
@param n_actions number of actions
@param state state at time t
"""
if rng.uniform(0, 1) < epsilon:
action = np.random.randint(0, n_actions)
else:
action = select_best_action(Q[state_id])
return action
# ## Discounted reward
# +
def get_discounted_reward(t, l_rewards_episode, gamma):
l_discounted_reward_episode = [t_prime_reward*(gamma**t_prime) for (t_prime, t_prime_reward) in enumerate(l_rewards_episode[t:])]
G_k_t = sum(l_discounted_reward_episode)
return G_k_t
def add_discounted_reward(steps_episode, gamma):
l_rewards_episode = [step_episode['reward'] for step_episode in steps_episode]
for (t, step_episode) in enumerate(steps_episode):
step_episode['discounted_reward'] = get_discounted_reward(t, l_rewards_episode, gamma)
# -
# ## Update N-matrice
def update_N_MC(N, step_episode, method_monte_carlo, states_already_visited=[]):
state_id = step_episode['state']['state_id']
action = step_episode['action']
previous_N_value_state = N[state_id].copy()
if method_monte_carlo == 'first_visit':
if not state_id in states_already_visited:
new_N_value = N[state_id][action] + 1
previous_N_value_state[action] = new_N_value
if method_monte_carlo == 'every_visit':
new_N_value = N[state_id][action] + 1
previous_N_value_state[action] = new_N_value
N[state_id] = previous_N_value_state
# ## Update Q-matrice (state-action value function)
# ### Monte-Carlo
def update_Q_MC(Q, N, step_episode, method_monte_carlo, states_already_visited=[]):
state_id = step_episode['state']['state_id']
action = step_episode['action']
G_k_t = step_episode['discounted_reward']
previous_Q_value_state = Q[state_id].copy()
if method_monte_carlo == 'first_visit':
if not state_id in states_already_visited:
new_Q_value = Q[state_id][action] + (G_k_t - Q[state_id][action]) / N[state_id][action]
previous_Q_value_state[action] = new_Q_value
if method_monte_carlo == 'every_visit':
new_Q_value = Q[state_id][action] + (G_k_t - Q[state_id][action]) / N[state_id][action]
previous_Q_value_state[action] = new_Q_value
Q[state_id] = previous_Q_value_state
# ### SARSA
# Function to learn the Q-value - Is it temporal-difference?
def update_Q_SARSA(Q, state1_id, action1, reward1, state2_id, action2, expected=False):
previous_Q_value_state1 = Q[state1_id].copy()
predict = Q[state1_id][action1]
target = reward1 + gamma * Q[state2_id][action2]
if expected:
expected_value = np.mean(Q[state2_id])
target = reward1 + gamma * expected_value
new_Q_value = Q[state1_id][action1] + alpha * (target - predict)
previous_Q_value_state1[action1] = new_Q_value
Q[state1_id] = previous_Q_value_state1
return Q
# ### Q-learning
# Function to learn the Q-value
def update_Q_Qlearning(Q, state1_id, action1, reward1, state2_id, action2, expected=False):
previous_Q_value_state1 = Q[state1_id].copy()
predict = Q[state1_id][action1]
target = reward1 + gamma * Q[state2_id][action2]
new_Q_value = Q[state1_id][action1] + alpha * (target - predict)
previous_Q_value_state1[action1] = new_Q_value
Q[state1_id] = previous_Q_value_state1
return Q
# + [markdown] heading_collapsed=true
# ## Updating parameters
# + [markdown] hidden=true
# ### Epsilon $\epsilon$ - Exploration rate
# + hidden=true
def get_epsilon(episode, init_epsilon):
n_epsilon = init_epsilon/(episode+1)
return n_epsilon
# + [markdown] hidden=true
# ### Alpha $\alpha$ - Learning rate
# + hidden=true
def get_alpha(episode, init_alpha):
n_alpha = init_alpha/(episode+1)
return n_alpha
# -
# ## Plots Reward / Steps / Happiness
# +
import matplotlib.colors as colors
def get_list_colors():
colors_list = ['r','g','b','k','darkorange','y','lime','c','m'] + list(colors._colors_full_map.values())
return colors_list
colors = get_list_colors()
# -
import numpy as np
def running_mean(x, N):
cumsum = np.cumsum(np.insert(x, 0, 0))
return (cumsum[N:] - cumsum[:-N]) / float(N)
# ### Reward
# +
import numpy as np
import matplotlib.pyplot as plt
def plot_evolution_reward(evolution_reward, method_name):
n_moving_points = int(np.ceil(len(evolution_reward)/100))
y = running_mean(evolution_reward,n_moving_points)
x = range(len(y))
info_parameters = '{} method - {} steps \n {} init_epsilon - {} init_alpha - {} gamma - {} nmax_steps'.format(
method_name, len(evolution_reward), init_epsilon, init_alpha, gamma, nmax_steps)
plt.plot(x, y)
plt.title('Evolution of Avg Reward per step per episode over time \n (smoothed over window size {})'.format(n_moving_points))
plt.xlabel('Episode \n '+ info_parameters)
plt.ylabel('Avg Reward per step per episode (Smoothed)')
plt.grid(True)
plt.tight_layout()
plt.savefig('data/figures/{}__reward.png'.format(method_name), format='png', dpi=500)
plt.show()
# -
def plot_comparison_evolution_reward(evo_training__evo_avg_reward_per_step):
n_episodes = len(list(evo_training__evo_avg_reward_per_step.values())[0])
info_parameters = 'All methods - {} episodes \n {} init_epsilon - {} init_alpha - {} gamma - {} nmax_steps'.format(
n_episodes, init_epsilon, init_alpha, gamma, nmax_steps)
fig = plt.figure()
cnt=0
for method in list(evo_training__evo_avg_reward_per_step.keys()):
n_moving_points = int(np.ceil(len(evo_training__evo_avg_reward_per_step[method])/100))
y = running_mean(evo_training__evo_avg_reward_per_step[method], n_moving_points)
x = range(len(y))
plt.plot(
x
, y
, label=method
, marker='', color=colors[cnt], linewidth=1, alpha=0.75
)
cnt += 1
plt.title('Evolution of Avg Reward per step per episode over time \n (smoothed over window size {})'.format(n_moving_points))
plt.xlabel('Episode \n '+ info_parameters)
plt.ylabel('Avg Reward per step \n per episode (Smoothed)')
plt.legend(bbox_to_anchor=(0.5,-0.10), loc="lower center",
bbox_transform=fig.transFigure, ncol=4, fancybox=True, shadow=True, borderpad=1)
plt.grid(True)
plt.tight_layout()
plt.savefig('data/figures/Comparison__reward.png', format='png', dpi=500)
plt.show()
# ### Number of steps
# +
import numpy as np
import matplotlib.pyplot as plt
def plot_evolution_steps(evolution_steps, method_name):
n_moving_points = int(np.ceil(len(evolution_steps)/100))
y = running_mean(evolution_steps,n_moving_points)
x = range(len(y))
info_parameters = '{} method - {} steps \n {} init_epsilon - {} init_alpha - {} gamma - {} nmax_steps'.format(
method_name, len(evolution_steps), init_epsilon, init_alpha, gamma, nmax_steps)
plt.plot(x, y)
plt.title('Episode Length over time \n (smoothed over window size {})'.format(n_moving_points))
plt.axhline(nmax_steps, color = 'r')
plt.axhline(0, color = 'b')
plt.ylim([-10, nmax_steps*1.05])
plt.xlabel('Episode \n '+ info_parameters)
plt.ylabel('Episode Length (Smoothed)')
plt.grid(True)
plt.tight_layout()
plt.savefig('data/figures/{}__steps.png'.format(method_name), format='png', dpi=500)
plt.show()
# -
def plot_comparison_evolution_steps(evo_training__evo_n_steps):
n_episodes = len(list(evo_training__evo_n_steps.values())[0])
info_parameters = 'All methods - {} episodes \n {} init_epsilon - {} init_alpha - {} gamma - {} nmax_steps'.format(
n_episodes, init_epsilon, init_alpha, gamma, nmax_steps)
fig = plt.figure()
cnt=0
for method in list(evo_training__evo_n_steps.keys()):
n_moving_points = int(np.ceil(len(evo_training__evo_n_steps[method])/100))
y = running_mean(evo_training__evo_n_steps[method], n_moving_points)
x = range(len(y))
plt.plot(
x, y, label=method
, marker='', color=colors[cnt], linewidth=1, alpha=0.75
)
cnt += 1
plt.title('Episode Length over time \n (smoothed over window size {})'.format(n_moving_points))
plt.axhline(nmax_steps, color = 'r')
plt.axhline(0, color = 'b')
plt.ylim([-10, nmax_steps*1.05])
plt.xlabel('Episode \n '+ info_parameters)
plt.ylabel('Episode Length (Smoothed)')
plt.legend(bbox_to_anchor=(0.5,-0.10), loc="lower center",
bbox_transform=fig.transFigure, ncol=4, fancybox=True, shadow=True, borderpad=1)
plt.grid(True)
plt.tight_layout()
plt.savefig('data/figures/Comparison__steps.png', format='png', dpi=500)
plt.show()
# ### Happiness
# +
import numpy as np
import matplotlib.pyplot as plt
def plot_evolution_happiness(evolution_happiness_all, method_name):
n_moving_points = int(np.ceil(len(evolution_happiness_all)/100))
y = running_mean(evolution_happiness_all,n_moving_points)
x = range(len(y))
info_parameters = '{} method - {} steps \n {} init_epsilon - {} init_alpha - {} gamma - {} nmax_steps'.format(
method_name, len(evolution_happiness_all), init_epsilon, init_alpha, gamma, nmax_steps)
plt.plot(x, y)
plt.title('Happiness over time \n (smoothed over window size {})'.format(n_moving_points))
plt.ylim([-0.05, 1.05])
plt.xlabel('Episode \n '+ info_parameters)
plt.ylabel('Happiness (Smoothed)')
plt.grid(True)
plt.tight_layout()
plt.savefig('data/figures/{}__happiness.png'.format(method_name), format='png', dpi=500)
plt.show()
# -
def plot_comparison_evolution_happiness(evo_training__evo_avg_happiness):
n_episodes = len(list(evo_training__evo_avg_happiness.values())[0])
info_parameters = 'All methods - {} episodes \n {} init_epsilon - {} init_alpha - {} gamma - {} nmax_steps'.format(
n_episodes, init_epsilon, init_alpha, gamma, nmax_steps)
fig = plt.figure()
cnt=0
for method in list(evo_training__evo_avg_happiness.keys()):
n_moving_points = int(np.ceil(len(evo_training__evo_avg_happiness[method])/100))
y = running_mean(evo_training__evo_avg_happiness[method], n_moving_points)
x = range(len(y))
plt.plot(
x, y, label=method
, marker='', color=colors[cnt], linewidth=1, alpha=0.75
)
cnt += 1
plt.title('Happiness over time \n (smoothed over window size {})'.format(n_moving_points))
plt.ylim([-0.05, 1.05])
plt.xlabel('Episode \n '+ info_parameters)
plt.ylabel('Happiness (Smoothed)')
plt.legend(bbox_to_anchor=(0.5,-0.10), loc="lower center",
bbox_transform=fig.transFigure, ncol=4, fancybox=True, shadow=True, borderpad=1)
plt.grid(True)
plt.tight_layout()
plt.savefig('data/figures/Comparison__happiness.png', format='png', dpi=500)
plt.show()
# # Parametrisation
# +
# Defining the different parameters
init_epsilon = 1 # trade-off exploration/exploitation
init_alpha = 0.5 # learning rate
init_Q_type = 'ones'
# Episodes
n_episodes = 10000
nmax_steps = 60*24 # maximum steps per episode
# -
# # Training the learning agent - Monte-Carlo - every visit
# +
method = 'MC'
method_monte_carlo = 'every_visit' # every_visit or first_visit
method_name = method + '_' + method_monte_carlo
# -
# ## Initialisation
# Initializing the Q-matrix
Q = init_Q(n_actions, init_Q_type)
N = init_N(n_actions)
# Visualisation
(render_episode, render_training) = (False, False)
n_episodes_plot = int(np.ceil(n_episodes/100))
# Initializing the reward
evo_training = {
'evo_avg_reward_per_step': []
, 'evo_n_steps': []
, 'evo_avg_happiness': []
}
# ## Training
# Starting the SARSA learning
for episode in tqdm(range(n_episodes)):
(n_episode_steps, done) = (0, False)
evo_episode = {
'episode_sum_reward': 0
, 'evolution_sum_happiness': 0
}
# Update parameters
epsilon = get_epsilon(episode, init_epsilon)
alpha = get_alpha(episode, init_alpha)
# Get episode
steps_episode = []
state1 = env_reset()
evo_episode['evolution_sum_happiness'] += get_happiness(state1)
action1 = epsilon_greedy(Q, state1['state_id'], n_actions, init_epsilon)
while (not done) and (n_episode_steps < nmax_steps):
# Getting the next state
state2, reward1, done, info = env_step(state1, action1)
evo_episode['episode_sum_reward'] += reward1
evo_episode['evolution_sum_happiness'] += get_happiness(state2)
steps_episode.append({
'state': state1,
'action': action1,
'reward' : reward1})
# Choosing the next action
action2 = epsilon_greedy(Q, state2['state_id'], n_actions, epsilon)
# Updating the respective values
state1 = state2
action1 = action2
n_episode_steps += 1
# Add discounted reward
add_discounted_reward(steps_episode, gamma)
# Update N and Q
states_already_visited = []
for step_episode in steps_episode:
update_N_MC(N, step_episode, method_monte_carlo, states_already_visited)
update_Q_MC(Q, N, step_episode, method_monte_carlo, states_already_visited)
states_already_visited.append(step_episode['state']['state_id'])
# At the end of learning process
if render_episode:
print('Episode {0}, Score: {1}, Timesteps: {2}, Epsilon: {3}, Alpha: {4}'.format(
episode+1, episode_reward, n_episode_steps, epsilon, alpha))
evo_training['evo_avg_reward_per_step'].append(evo_episode['episode_sum_reward'] / n_episode_steps)
evo_training['evo_n_steps'].append(n_episode_steps)
evo_training['evo_avg_happiness'].append(evo_episode['evolution_sum_happiness'] / n_episode_steps)
if ((episode+1) % n_episodes_plot == 0):
with open('data/interim/{}__Q.pkl'.format(method_name), 'wb') as file:
dill.dump(Q, file)
with open('data/interim/{}__N.pkl'.format(method_name), 'wb') as file:
dill.dump(N, file)
with open('data/interim/{}__evo_training.pkl'.format(method_name), 'wb') as file:
dill.dump(evo_training, file)
#plot_evolution_reward(evo_training['evo_avg_reward_per_step'], method_name)
#plot_evolution_steps(evo_training['evo_n_steps'], method_name)
#plot_evolution_happiness(evo_training['evo_avg_happiness'], method_name)
plot_evolution_reward(evo_training['evo_avg_reward_per_step'], method_name)
plot_evolution_steps(evo_training['evo_n_steps'], method_name)
plot_evolution_happiness(evo_training['evo_avg_happiness'], method_name)
# # Training the learning agent - Monte-Carlo - first visit
# +
method = 'MC'
method_monte_carlo = 'first_visit' # every_visit or first_visit
method_name = method + '_' + method_monte_carlo
# -
# ## Initialisation
# Initializing the Q-matrix
Q = init_Q(n_actions, init_Q_type)
N = init_N(n_actions)
# Visualisation
(render_episode, render_training) = (False, False)
n_episodes_plot = int(np.ceil(n_episodes/100))
# Initializing the reward
evo_training = {
'evo_avg_reward_per_step': []
, 'evo_n_steps': []
, 'evo_avg_happiness': []
}
# ## Training
# Starting the SARSA learning
for episode in tqdm(range(n_episodes)):
(n_episode_steps, done) = (0, False)
evo_episode = {
'episode_sum_reward': 0
, 'evolution_sum_happiness': 0
}
# Update parameters
epsilon = get_epsilon(episode, init_epsilon)
alpha = get_alpha(episode, init_alpha)
# Get episode
steps_episode = []
state1 = env_reset()
evo_episode['evolution_sum_happiness'] += get_happiness(state1)
action1 = epsilon_greedy(Q, state1['state_id'], n_actions, init_epsilon)
while (not done) and (n_episode_steps < nmax_steps):
# Getting the next state
state2, reward1, done, info = env_step(state1, action1)
evo_episode['episode_sum_reward'] += reward1
evo_episode['evolution_sum_happiness'] += get_happiness(state2)
steps_episode.append({
'state': state1,
'action': action1,
'reward' : reward1})
# Choosing the next action
action2 = epsilon_greedy(Q, state2['state_id'], n_actions, epsilon)
# Updating the respective values
state1 = state2
action1 = action2
n_episode_steps += 1
# Add discounted reward
add_discounted_reward(steps_episode, gamma)
# Update N and Q
states_already_visited = []
for step_episode in steps_episode:
update_N_MC(N, step_episode, method_monte_carlo, states_already_visited)
update_Q_MC(Q, N, step_episode, method_monte_carlo, states_already_visited)
states_already_visited.append(step_episode['state']['state_id'])
# At the end of learning process
if render_episode:
print('Episode {0}, Score: {1}, Timesteps: {2}, Epsilon: {3}, Alpha: {4}'.format(
episode+1, episode_reward, n_episode_steps, epsilon, alpha))
evo_training['evo_avg_reward_per_step'].append(evo_episode['episode_sum_reward'] / n_episode_steps)
evo_training['evo_n_steps'].append(n_episode_steps)
evo_training['evo_avg_happiness'].append(evo_episode['evolution_sum_happiness'] / n_episode_steps)
if ((episode+1) % n_episodes_plot == 0):
with open('data/interim/{}__Q.pkl'.format(method_name), 'wb') as file:
dill.dump(Q, file)
with open('data/interim/{}__N.pkl'.format(method_name), 'wb') as file:
dill.dump(N, file)
with open('data/interim/{}__evo_training.pkl'.format(method_name), 'wb') as file:
dill.dump(evo_training, file)
#plot_evolution_reward(evo_training['evo_avg_reward_per_step'], method_name)
#plot_evolution_steps(evo_training['evo_n_steps'], method_name)
#plot_evolution_happiness(evo_training['evo_avg_happiness'], method_name)
plot_evolution_reward(evo_training['evo_avg_reward_per_step'], method_name)
plot_evolution_steps(evo_training['evo_n_steps'], method_name)
plot_evolution_happiness(evo_training['evo_avg_happiness'], method_name)
# # Training the learning agent - SARSA
method = 'SARSA'
method_name = method
# ## Initialisation
# Initializing the Q-matrix
Q = init_Q(n_actions, init_Q_type)
# Visualisation
(render_episode, render_training) = (False, False)
n_episodes_plot = int(np.ceil(n_episodes/100))
# Initializing the reward
evo_training = {
'evo_avg_reward_per_step': []
, 'evo_n_steps': []
, 'evo_avg_happiness': []
}
# ## Training
# Starting the SARSA learning
for episode in tqdm(range(n_episodes)):
(n_episode_steps, done) = (0, False)
evo_episode = {
'episode_sum_reward': 0
, 'evolution_sum_happiness': 0
}
# Update parameters
epsilon = get_epsilon(episode, init_epsilon)
alpha = get_alpha(episode, init_alpha)
# Get episode
state1 = env_reset()
evo_episode['evolution_sum_happiness'] += get_happiness(state1)
action1 = epsilon_greedy(Q, state1['state_id'], n_actions, init_epsilon)
while (not done) and (n_episode_steps < nmax_steps):
# Getting the next state
state2, reward1, done, info = env_step(state1, action1)
evo_episode['episode_sum_reward'] += reward1
evo_episode['evolution_sum_happiness'] += get_happiness(state2)
# Choosing the next action
action2 = epsilon_greedy(Q, state2['state_id'], n_actions, epsilon)
# Learning the Q-value
Q = update_Q_SARSA(Q,state1['state_id'], action1, reward1, state2['state_id'], action2)
# Updating the respective values
state1 = state2
action1 = action2
n_episode_steps += 1
# At the end of learning process
if render_episode:
print('Episode {0}, Score: {1}, Timesteps: {2}, Epsilon: {3}, Alpha: {4}'.format(
episode+1, episode_reward, n_episode_steps, epsilon, alpha))
evo_training['evo_avg_reward_per_step'].append(evo_episode['episode_sum_reward'] / n_episode_steps)
evo_training['evo_n_steps'].append(n_episode_steps)
evo_training['evo_avg_happiness'].append(evo_episode['evolution_sum_happiness'] / n_episode_steps)
if ((episode+1) % n_episodes_plot == 0):
with open('data/interim/{}__Q.pkl'.format(method_name), 'wb') as file:
dill.dump(Q, file)
with open('data/interim/{}__evo_training.pkl'.format(method_name), 'wb') as file:
dill.dump(evo_training, file)
#plot_evolution_reward(evo_training['evo_avg_reward_per_step'], method_name)
#plot_evolution_steps(evo_training['evo_n_steps'], method_name)
#plot_evolution_happiness(evo_training['evo_avg_happiness'], method_name)
plot_evolution_reward(evo_training['evo_avg_reward_per_step'], method_name)
plot_evolution_steps(evo_training['evo_n_steps'], method_name)
plot_evolution_happiness(evo_training['evo_avg_happiness'], method_name)
# # Training the learning agent - Q-learning
method = 'Q-Learning'
method_name = method
# ## Initialisation
# Initializing the Q-matrix
Q = init_Q(n_actions, init_Q_type)
# Visualisation
(render_episode, render_training) = (False, False)
n_episodes_plot = int(np.ceil(n_episodes/100))
# Initializing the reward
evo_training = {
'evo_avg_reward_per_step': []
, 'evo_n_steps': []
, 'evo_avg_happiness': []
}
# ## Training
# Starting the SARSA learning
for episode in tqdm(range(n_episodes)):
(n_episode_steps, done) = (0, False)
evo_episode = {
'episode_sum_reward': 0
, 'evolution_sum_happiness': 0
}
# Update parameters
epsilon = get_epsilon(episode, init_epsilon)
alpha = get_alpha(episode, init_alpha)
# Get episode
state1 = env_reset()
evo_episode['evolution_sum_happiness'] += get_happiness(state1)
while (not done) and (n_episode_steps < nmax_steps):
# Choose an action
action1 = epsilon_greedy(Q, state1['state_id'], n_actions, init_epsilon)
# Getting the next state
state2, reward1, done, info = env_step(state1, action1)
evo_episode['episode_sum_reward'] += reward1
evo_episode['evolution_sum_happiness'] += get_happiness(state2)
# Q-Learning
# Choosing the next action
action2 = select_best_action(Q[state2['state_id']])
# Learning the Q-value
Q = update_Q_Qlearning(Q, state1['state_id'], action1, reward1, state2['state_id'], action2)
# Updating the respective values
state1 = state2
n_episode_steps += 1
# At the end of learning process
if render_episode:
print('Episode {0}, Score: {1}, Timesteps: {2}, Epsilon: {3}, Alpha: {4}'.format(
episode+1, episode_reward, n_episode_steps, epsilon, alpha))
evo_training['evo_avg_reward_per_step'].append(evo_episode['episode_sum_reward'] / n_episode_steps)
evo_training['evo_n_steps'].append(n_episode_steps)
evo_training['evo_avg_happiness'].append(evo_episode['evolution_sum_happiness'] / n_episode_steps)
if ((episode+1) % n_episodes_plot == 0):
with open('data/interim/{}__Q.pkl'.format(method_name), 'wb') as file:
dill.dump(Q, file)
with open('data/interim/{}__evo_training.pkl'.format(method_name), 'wb') as file:
dill.dump(evo_training, file)
#plot_evolution_reward(evo_training['evo_avg_reward_per_step'], method_name)
#plot_evolution_steps(evo_training['evo_n_steps'], method_name)
#plot_evolution_happiness(evo_training['evo_avg_happiness'], method_name)
plot_evolution_reward(evo_training['evo_avg_reward_per_step'], method_name)
plot_evolution_steps(evo_training['evo_n_steps'], method_name)
plot_evolution_happiness(evo_training['evo_avg_happiness'], method_name)
# # Comparison
list_methods = ['MC_every_visit','MC_first_visit', 'SARSA','Q-Learning']
# +
evo_training__evo_avg_reward_per_step = {}
evo_training__evo_n_steps = {}
evo_training__evo_avg_happiness = {}
for method in list_methods:
with open("data/interim/{}__evo_training.pkl".format(method), "rb") as input_file:
evo_training = dill.load(input_file)
evo_training__evo_avg_reward_per_step[method] = evo_training['evo_avg_reward_per_step']
evo_training__evo_n_steps[method] = evo_training['evo_n_steps']
evo_training__evo_avg_happiness[method] = evo_training['evo_avg_happiness']
# -
plot_comparison_evolution_reward(evo_training__evo_avg_reward_per_step)
plot_comparison_evolution_steps(evo_training__evo_n_steps)
plot_comparison_evolution_happiness(evo_training__evo_avg_happiness)
|
notebooks/3.0-yp-Comparison.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:PythonData] *
# language: python
# name: conda-env-PythonData-py
# ---
from bs4 import BeautifulSoup
from splinter import Browser
import pandas as pd
from webdriver_manager.chrome import ChromeDriverManager
import requests
executable_path = {'executable_path': ChromeDriverManager().install()}
browser = Browser('chrome', **executable_path, headless=False)
url = "https://mars.nasa.gov/news/"
browser.visit(url)
html = browser.html
news_soup = BeautifulSoup(html, 'html.parser')
news_soup.find("div", class_="content_title")
news_title = news_soup.find("div", class_="content_title").get_text()
print(news_title)
news_p = news_soup.find("div", class_="article_teaser_body").get_text()
print(news_p)
browser.quit()
executable_path = {'executable_path': ChromeDriverManager().install()}
browser = Browser('chrome', **executable_path, headless=False)
url = "https://spaceimages-mars.com/"
browser.visit(url)
html = browser.html
image_soup = BeautifulSoup(html, "html.parser")
browser.links.find_by_partial_text('FULL IMAGE').click()
img_url = image_soup.find(class_="headerimage fade-in").get("src")
print(img_url)
featured_img_url = f"https://spaceimages-mars.com/{img_url}"
print(featured_img_url)
browser.quit()
mars_facts = pd.read_html("https://galaxyfacts-mars.com/")
print(mars_facts)
mars_facts_df = mars_facts[0]
mars_facts_df
mars_facts_df.reset_index(inplace=True)
mars_facts_df.columns=["ID", "Properties", "Mars", "Earth"]
mars_facts_df
executable_path = {'executable_path': ChromeDriverManager().install()}
browser = Browser('chrome', **executable_path, headless=False)
mhurl = 'https://astrogeology.usgs.gov/search/results?q=hemisphere+enhanced&k1=target&v1=Mars'
browser.visit(mhurl)
mhtml = browser.html
mh_soup = BeautifulSoup(mhtml,"html.parser")
results = mh_soup.find_all("div",class_='item')
hemisphere_image_urls = []
for result in results:
product_dict = {}
titles = result.find('h3').text
end_link = result.find("a")["href"]
image_link = "https://astrogeology.usgs.gov/" + end_link
browser.visit(image_link)
html = browser.html
soup= BeautifulSoup(html, "html.parser")
downloads = soup.find("div", class_="downloads")
image_url = downloads.find("a")["href"]
print(titles)
print(image_url)
product_dict['title']= titles
product_dict['image_url']= image_url
hemisphere_image_urls.append(product_dict)
hemisphere_image_urls
browser.quit()
|
Missions_to_Mars/mission_to_mars.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Ethan-Jeong/test_deeplearning/blob/master/reuter_LSTM.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="FeE9eLEB3I3j"
import tensorflow as tf
# + colab={"base_uri": "https://localhost:8080/"} id="w2hkoozD5b5o" outputId="0897dcb7-8853-425c-a6cd-e872670548d9"
(x_train,y_train),(x_test , y_test) = tf.keras.datasets.reuters.load_data(num_words=10000)
x_train.shape , y_train.shape , x_test.shape , y_test.shape
# + colab={"base_uri": "https://localhost:8080/"} id="eMHfoM376IfY" outputId="38f5a2c5-d382-415a-bd48-78073eb5a1f6"
print(y_train[50] , x_train[50])
# + colab={"base_uri": "https://localhost:8080/"} id="FVnWKvRJ_Wg2" outputId="9005b5b1-f12d-4e97-8891-a093d2b77e1c"
len(x_train[50]),len(x_train[100]),len(x_train[500]),len(x_train[1000])
# + colab={"base_uri": "https://localhost:8080/"} id="9lhHDRYoB2_U" outputId="562e5f72-27f6-4788-c0d2-99ad55ae621e"
pad_x_train = tf.keras.preprocessing.sequence.pad_sequences(x_train,maxlen=500)
len(pad_x_train)
# + id="QATyiTheEVdG"
import numpy as np
# + colab={"base_uri": "https://localhost:8080/"} id="UGhxpi2YEYnL" outputId="02396cb7-c35b-4ed0-f28b-60f75f9b607d"
np.unique(y_train) , len(np.unique(y_train))
# + [markdown] id="oTfFjoo07-Ap"
# # make model
# + id="aRDCsYPR7wkl"
model = tf.keras.models.Sequential()
# + id="mgv2K59J8CVX"
model.add( tf.keras.layers.Embedding(input_length=500,input_dim=10000, output_dim=24) ) # output_dim , 차원의 숫자를 뜻하며 임의로 숫자 설정할수 있음
model.add( tf.keras.layers.LSTM( 24 , return_sequences=True, activation='tanh')) # 차원의 숫자를 넣으면 됨
model.add( tf.keras.layers.LSTM( 12 , activation='tanh')) # 차원의 숫자를 넣으면 됨 , 임의의 숫자를 넣으면 됨
# model.add( tf.keras.layers.Flatten())
model.add( tf.keras.layers.Dense(46,activation='softmax'))
model.compile(optimizer='adam', loss='sparse_categorical_crossentropy' , metrics=['acc'])
# + colab={"base_uri": "https://localhost:8080/"} id="b1Mia8D9Hir5" outputId="217333d2-c9fc-46e8-9580-6af89afa25b7"
# hist = model.fit( pad_x_train , y_train , epochs=5 , validation_split=0.3 , batch_size=128)
hist = model.fit( pad_x_train , y_train , epochs=100 , validation_split=0.3 , batch_size=256)
# + [markdown] id="wCCIM5VoRpwP"
# # Evaluation
# + colab={"base_uri": "https://localhost:8080/"} id="7dEYJdvFPzPL" outputId="119ee1dd-de67-4706-f08e-cb7e9853ee6f"
model.evaluate( pad_x_train, y_train)
# + id="npRxRmMiUv1t"
pad_x_train = tf.keras.preprocessing.sequence.pad_sequences(x_train,maxlen=500)
pad_x_test = tf.keras.preprocessing.sequence.pad_sequences(x_test,maxlen=500)
# + id="7grhYoZoXKNO"
def pad_make(x_data):
pad_x = tf.keras.preprocessing.sequence.pad_sequences(x_data,maxlen=500)
return pad_x
# + id="ov9Vc2RMX7B5"
pad_make_x = pad_make(x_test)
# + colab={"base_uri": "https://localhost:8080/"} id="INte7KiHYOVC" outputId="def3f844-207e-468f-8dc4-77ba064b1b60"
model.evaluate(pad_make_x,y_test)
# + colab={"base_uri": "https://localhost:8080/"} id="5Aelb5YARwQ1" outputId="5617d3ce-0fda-4c2f-b7ce-8de2d2ac2ccc"
model.evaluate( pad_x_test , y_test)
# + id="Bql5wCZEVCwi"
import matplotlib.pyplot as plt
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="R_6GVOaaVo-s" outputId="7942a834-0a82-45d5-ade4-c53aaee4d646"
plt.plot(hist.history['loss'])
plt.plot(hist.history['val_loss'])
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 283} id="vOPtQbSTWFPK" outputId="d2a27b0e-42e8-47f0-a1db-4d01f19c3b48"
plt.plot(hist.history['acc'])
plt.plot(hist.history['val_acc'])
# + id="vOsypt8sWVhX"
from sklearn.metrics import classification_report
# + colab={"base_uri": "https://localhost:8080/"} id="54nSsPG2p7-Y" outputId="1ff4a0a5-c105-44d4-8b3c-80e5166b8147"
y_train_pred = model.predict(pad_x_train)
y_train_pred[0]
# + id="ug5vuXGwqNcP"
import numpy as np
# + colab={"base_uri": "https://localhost:8080/"} id="p5JV5y4QqYAN" outputId="db86b19e-9633-409e-959a-32da2d8b51b5"
y_pred = np.argmax(y_train_pred,axis=1)
y_pred.shape
# + id="_yhDRiAnqkj4"
print(classification_report(y_train,y_pred))
# + id="4CAQpjwOq231"
y_test_pred = model.predict(pad_x_test)
# + id="WguvVY1stAmz"
y_pred = np.argmax(y_test_pred,axis=1)
# + colab={"base_uri": "https://localhost:8080/"} id="is8m9oxJtN5S" outputId="dd657443-0874-4da5-fd48-463e4e322342"
print(classification_report(y_test,y_pred))
# + id="ZhxZuUmotkwH"
|
reuter_LSTM.ipynb
|
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .js
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Arc
// language: javascript
// name: arc
// ---
// %conf streaming=true
// ## Markdown cell
// A markdown cell to ensure the conversion doesn't break.
{
"type": "RateExtract",
"name": "create a streaming source",
"environments": [
"production",
"test"
]
}
// %sql
SELECT *
FROM stream
// + active=""
// a raw NBConvert cell
// -
// %configplugin
{
"type": "ai.tripl.arc.plugins.TestDynamicConfigurationPlugin",
"environments": ["test"],
"key": "testValue"
}
// %lifecycleplugin
{
"type": "ai.tripl.arc.plugins.TestLifecyclePlugin",
"name": "test",
"environments": ["test"],
"outputViewBefore": "before",
"outputViewAfter": "after",
"value": "testValue"
}
// %arc numRows=10
{
"type": "RateExtract",
"name": "create a streaming second source",
"environments": [
"production",
"test"
],
"outputView": "stream2"
}
|
src/test/resources/conf/error_job.ipynb
|
# -*- coding: utf-8 -*-
# # 边界检查
#
# 和许多其他现代编程语言一样,Julia 在访问数组元素的时候也要通过边界检查来确保程序安全。当循环次数很多,或者在其他性能敏感的场景下,你可能希望不进行边界检查以提高运行时性能。比如要使用矢量 (SIMD) 指令,循环体就不能有分支语句,因此无法进行边界检查。Julia 提供了一个宏 `@inbounds(...)` 来告诉编译器在指定语句块不进行边界检查。用户自定义的数组类型可以通过宏 `@boundscheck(...)` 来达到上下文敏感的代码选择目的。
#
# ## 移除边界检查
#
# 宏 `@boundscheck(...)` 把代码块标记为要执行边界检查。但当这些代码块被被宏 `@inbounds(...)` 标记的代码包裹时,它们可能会被编译器移除。仅当`@boundscheck(...)` 代码块被调用函数包裹时,编译器会移除它们。比如你可能这样写的 `sum` 方法:
# + attributes={"classes": ["julia"], "id": ""}
function sum(A::AbstractArray)
r = zero(eltype(A))
for i in eachindex(A)
@inbounds r += A[i]
end
return r
end
# -
# 使用自定义的类数组类型 `MyArray`,我们有:
# + attributes={"classes": ["julia"], "id": ""}
@inline getindex(A::MyArray, i::Real) = (@boundscheck checkbounds(A,i); A.data[to_index(i)])
# -
# 当 `getindex` 被 `sum` 包裹时,对 `checkbounds(A,i)` 的调用会被忽略。如果存在多层包裹,最多只有一个 `@boundscheck` 被忽略。这个规则用来防止将来代码被改变时潜在的多余忽略。
#
# ## Propagating inbounds
#
# There may be certain scenarios where for code-organization reasons you want more than one layer
# between the `@inbounds` and `@boundscheck` declarations. For instance, the default `getindex`
# methods have the chain `getindex(A::AbstractArray, i::Real)` calls `getindex(IndexStyle(A), A, i)`
# calls `_getindex(::IndexLinear, A, i)`.
#
# To override the "one layer of inlining" rule, a function may be marked with
# [`Base.@propagate_inbounds`](@ref) to propagate an inbounds context (or out of bounds
# context) through one additional layer of inlining.
#
# ## The bounds checking call hierarchy
#
# The overall hierarchy is:
#
# * `checkbounds(A, I...)` which calls
* `checkbounds(Bool, A, I...)` which calls
* `checkbounds_indices(Bool, axes(A), I)` which recursively calls
* `checkindex` for each dimension
# Here `A` is the array, and `I` contains the "requested" indices. `axes(A)` returns a tuple
# of "permitted" indices of `A`.
#
# `checkbounds(A, I...)` throws an error if the indices are invalid, whereas `checkbounds(Bool, A, I...)`
# returns `false` in that circumstance. `checkbounds_indices` discards any information about the
# array other than its `axes` tuple, and performs a pure indices-vs-indices comparison: this
# allows relatively few compiled methods to serve a huge variety of array types. Indices are specified
# as tuples, and are usually compared in a 1-1 fashion with individual dimensions handled by calling
# another important function, `checkindex`: typically,
# + attributes={"classes": ["julia"], "id": ""}
checkbounds_indices(Bool, (IA1, IA...), (I1, I...)) = checkindex(Bool, IA1, I1) &
checkbounds_indices(Bool, IA, I)
# -
# so `checkindex` checks a single dimension. All of these functions, including the unexported
# `checkbounds_indices` have docstrings accessible with `?` .
#
# If you have to customize bounds checking for a specific array type, you should specialize `checkbounds(Bool, A, I...)`.
# However, in most cases you should be able to rely on `checkbounds_indices` as long as you supply
# useful `axes` for your array type.
#
# If you have novel index types, first consider specializing `checkindex`, which handles a single
# index for a particular dimension of an array. If you have a custom multidimensional index type
# (similar to `CartesianIndex`), then you may have to consider specializing `checkbounds_indices`.
#
# Note this hierarchy has been designed to reduce the likelihood of method ambiguities. We try
# to make `checkbounds` the place to specialize on array type, and try to avoid specializations
# on index types; conversely, `checkindex` is intended to be specialized only on index type (especially,
# the last argument).
#
# ## Emit bounds checks
#
# Julia can be launched with `--check-bounds={yes|no|auto}` to emit bounds checks always, never, or respect @inbounds declarations.
|
zh_CN_Jupyter/doc/src/devdocs/boundscheck.md.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] toc=true
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#Numpy-Tutorial" data-toc-modified-id="Numpy-Tutorial-1"><span class="toc-item-num">1 </span>Numpy Tutorial</a></span><ul class="toc-item"><li><span><a href="#BASICS" data-toc-modified-id="BASICS-1.1"><span class="toc-item-num">1.1 </span>BASICS</a></span></li><li><span><a href="#Adding-and-removing-elements-from-numpy-arrays" data-toc-modified-id="Adding-and-removing-elements-from-numpy-arrays-1.2"><span class="toc-item-num">1.2 </span>Adding and removing elements from numpy arrays</a></span></li><li><span><a href="#Accessing/Changing-specific-elements,-rows,-columns,-etc" data-toc-modified-id="Accessing/Changing-specific-elements,-rows,-columns,-etc-1.3"><span class="toc-item-num">1.3 </span>Accessing/Changing specific elements, rows, columns, etc</a></span></li><li><span><a href="#Initializing-different-type-of-arrays" data-toc-modified-id="Initializing-different-type-of-arrays-1.4"><span class="toc-item-num">1.4 </span>Initializing different type of arrays</a></span></li><li><span><a href="#Mathematics" data-toc-modified-id="Mathematics-1.5"><span class="toc-item-num">1.5 </span>Mathematics</a></span><ul class="toc-item"><li><span><a href="#linear-algebra" data-toc-modified-id="linear-algebra-1.5.1"><span class="toc-item-num">1.5.1 </span>linear algebra</a></span></li><li><span><a href="#statistics" data-toc-modified-id="statistics-1.5.2"><span class="toc-item-num">1.5.2 </span>statistics</a></span></li></ul></li><li><span><a href="#Miscellaneous---loading-data" data-toc-modified-id="Miscellaneous---loading-data-1.6"><span class="toc-item-num">1.6 </span>Miscellaneous - loading data</a></span><ul class="toc-item"><li><span><a href="#load-data-from-file" data-toc-modified-id="load-data-from-file-1.6.1"><span class="toc-item-num">1.6.1 </span>load data from file</a></span></li><li><span><a href="#boolean-masking-and-advanced-indexing" data-toc-modified-id="boolean-masking-and-advanced-indexing-1.6.2"><span class="toc-item-num">1.6.2 </span>boolean masking and advanced indexing</a></span></li></ul></li><li><span><a href="#Creating-Structured-array" data-toc-modified-id="Creating-Structured-array-1.7"><span class="toc-item-num">1.7 </span>Creating Structured array</a></span></li><li><span><a href="#Rearranging-array-elements" data-toc-modified-id="Rearranging-array-elements-1.8"><span class="toc-item-num">1.8 </span>Rearranging array elements</a></span></li><li><span><a href="#Transpose-like-operations" data-toc-modified-id="Transpose-like-operations-1.9"><span class="toc-item-num">1.9 </span>Transpose-like operations</a></span></li><li><span><a href="#Universal-Functions" data-toc-modified-id="Universal-Functions-1.10"><span class="toc-item-num">1.10 </span>Universal Functions</a></span></li></ul></li></ul></div>
# -
# # Numpy Tutorial
#
# * links
# * [A Visual Intro to NumPy and Data Representation](http://jalammar.github.io/visual-numpy/)
# * [Python NumPy Tutorial for Beginners](https://www.youtube.com/watch?v=QUT1VHiLmmI)
# * [NumPy Data Science Essential Training With Python 3](https://www.youtube.com/playlist?list=PLZ7s-Z1aAtmIRpnGQGMTvV3AGdDK37d2b)
#
import numpy as np
# ## BASICS
# +
a = np.array([1, 2, 3])
b = np.array([
[1., 1.],
[1., 1.]
])
c = np.array([1,2, 3], dtype="int16")
# -
print(a)
print(b)
print(c)
print('-------')
print(a.ndim)
print(b.ndim)
print(c.ndim)
print('-------')
print(a.shape)
print(b.shape)
print(c.shape)
# * **Numpy Benefits**
#
# 1) Fixed Type
#
# 2) Contigious memory
#
# get type
print(a.dtype)
print(b.dtype)
print(c.dtype)
# get size - size of each item (B)
print(a.itemsize)
print(b.itemsize)
print(c.itemsize)
# get total size - total space used (B )
print(a.size * a.itemsize)
print(a.nbytes)
print(b.nbytes)
print(c.nbytes)
arr = [range(1, 5), range(5, 9)] # list of lists
e = np.array(arr) # 2d array
print(e)
x = e.tolist() # convert array back to list
print(type(x))
# ## Adding and removing elements from numpy arrays
a = np.array(np.arange(24)).reshape((2, 3, 4))
a
b = np.append(a, [5, 6, 7, 8])
b
b.shape
b.reshape((7, 4))
a
c = np.array(np.arange(24)).reshape((2, 3, 4)) * 10 + 3
c
np.append(a, c, axis=0)
np.append(a, c, axis=0).shape
np.append(a, c, axis=1)
np.append(a, c, axis=1).shape
np.append(a, c, axis=2)
np.append(a, c, axis=2).shape
after_insert_array = np.insert(c, 1, 444, axis=0)
after_insert_array
after_insert_array = np.insert(c, 1, 444, axis=1)
after_insert_array
after_insert_array = np.insert(c, 1, 444, axis=2)
after_insert_array
d = np.empty(c.shape)
np.copyto(d, c)
d
np.delete(d, 1, axis=0)
np.delete(d, 1, axis=1)
np.delete(d, 1, axis=2)
# ## Accessing/Changing specific elements, rows, columns, etc
a = np.array([[1,2, 3, 4, 5, 6, 7], [8, 9, 10, 11, 12, 13, 14]])
print(a)
print(a.shape)
# get a specific element [r, c]
print(a[1, 5])
print(a[1][5])
print(a[1][-2])
# get a specific row
a[0, :]
# get a specific col
a[:, 2]
# getting fancy [startindex: endindex: stepsize]
print(a[0, 1:6:2])
# changing element(s)
a [1, 5] = 20
print(a)
print('-----')
a[:, 2] = 5
print(a)
print('-----')
a[:, 2] = [1, 2]
print(a)
# +
# 3-d
a = np.array([[[1, 2], [3, 4]], [[5, 6], [7, 8]]])
print(a)
print('------')
print(a.shape)
# think of number of rows as original number of rows
# inside each box columns become rows and depth becomes columns
# think of it as [1, 2] at at position a[0, 0]
# NumPy’s order for printing n-dimensional arrays is that the last axis
# is looped over the fastest, while the first is the slowest
# +
print(a[0, 0])
print(a[0, 1])
print(a[1, 0])
print(a[1, 1])
# get specific element
print(a[0,1,1])
# +
# replace
a[:, 1, :] = [[10, 11], [12, 13]]
print(a)
print('-----')
a[0, 1, :] = [99, 98]
print(a)
# -
# ## Initializing different type of arrays
# all 0's matrix
np.zeros(shape=(2, 3))
# all 1's matrix
np.ones((4, 2, 2), dtype='int32')
# any other number
np.full((2, 2), 99)
# any other number (full_like)
a = np.array([[1,2, 3, 4, 5, 6, 7], [8, 9, 10, 11, 12, 13, 14]])
np.full_like(a, 8)
# random decimal numbers
np.random.rand(4, 2, 3)
np.random.random_sample(a.shape)
# random integer values
print(np.random.randint(7))
print(np.random.randint(7, size=(3, 3)))
# identity matrix
np.identity(3)
print(np.logspace(0, 3, 4)) # 10^0 to 10^3 (inclusive) with 4 points
# +
# repeat an array
arr = np.array([[1, 2, 3]])
r1 = np.repeat(arr, 3, axis=0)
print(r1)
print('-----')
r2 = np.repeat(arr, 3, axis=1)
print(r2)
# +
# creating a customized matrix
output = np.ones((5, 5))
print(output)
print('-----')
z = np.zeros((3, 3))
z[1, 1] = 9
print(z)
print('-----')
output[1:4, 1: 4] = z
print(output)
# +
# careful while copying
a = np.array([1, 2, 3])
b = a # just points
b [0]= 100
print(a)
print(b)
print('-----')
a = np.array([1, 2, 3])
b = a.copy()
b [0]= 100
print(a)
print(b)
# -
# adding an axis
a = np.array([0, 1])
a_col = a[:, np.newaxis]
print(a_col
)
a_col.T
# +
# flatten array
# always returns a flat copy of the original array
arr = np.arange(10, dtype=float).reshape((2, 5))
arr_flat = arr.flatten()
arr_flat[0] = 33
print(arr)
print(arr_flat)
# +
# Ravel: returns a view of the original array whenever possible.
arr_flt = arr.ravel()
arr_flt[0] = 33
print(arr)
print(arr_flt)
# +
# creating evenly spaced number over an specified interval
np.linspace(start = 0, stop=50, num=10)
np.linspace(5, 15, 10)
mylinspace = np.linspace(5, 15, 9, retstep=True)
mylinspace
# -
# ## Mathematics
a = np.array([1, 2, 3, 4])
print(a)
a + 2
a - 2
a / 2
b = np.array([1, 0, 1, 0])
a + b
np.sin(a)
# ### linear algebra
#
my_first_matrix = np.matrix([[3, 1, 4], [1, 5, 9], [2, 6, 5]])
my_first_matrix
my_first_matrix.T
my_first_matrix.I # inverse of matrix
# +
a = np.ones((2, 3))
print(a)
b = np.full((3, 2), 2)
print(b)
# -
c = np.identity(4) # or np.eye(5)
print(np.linalg.det(c))
# +
# Solve simultaneous linear equations
right_hand_side = np.matrix([[11],
[22],
[33]])
my_first_inverse = my_first_matrix.I
# -
solution = my_first_inverse * right_hand_side
solution
# more efficient for large matrices
from numpy.linalg import solve
solve(my_first_matrix, right_hand_side)
# compute the eigenvalues and eigenvectors
from numpy.linalg import eig
eig(my_first_matrix)
# ### statistics
stats = np.array([[1, 2, 3], [4, 5,6]])
stats
# +
print(np.min(stats, axis=0))
print(np.min(stats, axis=1))
print(np.max(stats, axis=0))
print(np.max(stats, axis=1))
# -
np.sum(stats, axis=0)
rnd = np.random.randn(4, 2) # random normals in 4x2 array
print(rnd.mean())
print(rnd.std())
print(rnd.argmin()) # index of minimum element
print(rnd.sum())
print(rnd.sum(axis=0)) # sum of columns
print(rnd.sum(axis=1)) # sum of rows
# ## Miscellaneous - loading data
# ### load data from file
#
filedata = np.genfromtxt('data.txt', delimiter=',')
filedata = filedata.astype('int32')
print(filedata)
# ### boolean masking and advanced indexing
filedata > 50
filedata[filedata > 50]
# we can index using a list
a = np.array([1, 2, 3, 4,5, 6, 7,8,9])
a[[1, 2, 8]]
np.any(filedata > 50, axis= 0)
np.all(filedata > 50, axis= 0)
((filedata > 50) & (filedata < 100))
(~(filedata > 50) & (filedata < 100))
filedata[((filedata > 50) & (filedata < 100))]
my_vector = np.array([-17, -4, 0, 2, 21, 37, 105])
zero_mod_7_mask = 0 == (my_vector % 7)
print(zero_mod_7_mask)
my_vector[zero_mod_7_mask]
mod_test = 0 == (my_vector % 7)
posmask = my_vector > 0
combined_mask = np.logical_and(mod_test, posmask)
# combined_mask
print(my_vector[combined_mask])
# example indexing
a = np.arange(1, 11).reshape((2, 5))
b = np.arange(11, 21).reshape((2, 5))
c = np.arange(21, 31).reshape((2, 5))
final = np.vstack([a, b, c])
print(final)
final[2:4, :2]
final[[0, 1, 2, 3], [1, 2, 3, 4]]
final[[0, 4, 5], 3:]
# ## Creating Structured array
#
# used for heterogeneous data while maintaining numpy's requirement that every element in an array use the same amount of memory space
person_data_def = [('name', '<U6'), ('height', 'f8'), ('weight', 'f8'), ('age', 'i8')]
person_data_def
people_array = np.zeros((4), dtype=person_data_def)
people_array
# +
# https://docs.scipy.org/doc/numpy/reference/arrays.dtypes.html
# https://jakevdp.github.io/PythonDataScienceHandbook/02.09-structured-data-numpy.html
# dt = np.dtype('i4') # 32-bit signed integer
# dt = np.dtype('f8') # 64-bit floating-point number
# dt = np.dtype('c16') # 128-bit complex floating-point number
# dt = np.dtype('a25') # 25-length zero-terminated bytes
# dt = np.dtype('U25') # 25-character string
# -
people_array
people_array[3] = ('delta', 73, 58, 28)
people_array[3]
people_array[1] = ('alpha', 83, 38, 48)
people_array[1]
people_array
people_array['age']
people_big_array = np.zeros((4, 3, 2), dtype=person_data_def)
people_big_array
people_big_array[3, 2, 1] = ('echo', 10, 20, 30)
people_big_array
# +
# creating Record arrays
# -
person_record_array = np.rec.array([('Delta', 73, 205, 34), ('alpha', 83, 38, 48)], dtype=person_data_def)
person_record_array
person_record_array[0].age
# using attributes instead of index
# ## Rearranging array elements
# +
before = np.array([[1, 2, 3 , 4], [5, 6, 7, 8]])
print(before)
print(before.reshape((8,1)))
# +
# vertically stacking arrays
v1 = np.array([1, 2, 3, 4])
v2 = np.array([11, 22, 33, 44])
np.vstack((v1, v2, v2))
# -
# horizontal stacking
h1 = np.ones((2, 4))
h2 = np.zeros((2, 2))
np.hstack([h1, h2])
my_start_array = np.array(np.arange(24))
my_3_8_array = my_start_array.reshape((3, 8))
my_2_3_4_array = my_start_array.reshape((2, 3, 4))
# +
# fliplr - flip left right
# -
my_3_8_array
np.fliplr(my_3_8_array)
my_2_3_4_array
np.fliplr(my_2_3_4_array) # flipping takes place over the last index
# flip upside down
np.flipud(my_3_8_array)
np.flipud(my_2_3_4_array)
my_start_array
# roll
np.roll(my_start_array, 5)
np.roll(my_start_array, -5)
np.roll(my_2_3_4_array, 2)
my_3_8_array
# +
# rotate 90 degree
np.rot90(my_3_8_array) # rotate in +ve direction (counter-clockwise)
# -
np.rot90(my_3_8_array, k=-1) # rotate in -ve direction (clockwise)
# ## Transpose-like operations
my_start_array = np.array(np.arange(24))
my_3_8_array = my_start_array.reshape((3, 8))
my_2_3_4_array = my_start_array.reshape((2, 3, 4))
print(my_start_array)
print('-----')
print(my_start_array.T)
# or
# print(np.transpose(my_start_array))
print(my_3_8_array)
print('-----')
print(my_3_8_array.T)
print(my_2_3_4_array)
print('-----')
print(np.transpose(my_2_3_4_array, axes=(0,2,1)))
# transpose over axes index by 2 and axes index by 1
# axes = By default, reverse the dimensions,
# otherwise permute the axes according to the values given.
# swapaxes(a, axis1, axis2) - interchange two axes of an array
print(my_2_3_4_array)
print('-----')
print(np.swapaxes(my_2_3_4_array, 1, 0) )
# np.rollaxis - roll the specified axis backwards, until it lies in a given position
print(my_2_3_4_array.shape)
print('-----')
print(np.rollaxis(my_2_3_4_array, axis=1, start=3).shape)
# axis 3 is not present but theoretically will be after axis 2 so axis
# 1 is rolled till it is behind axis 3
print(my_2_3_4_array.shape)
print('-----')
print(np.rollaxis(my_2_3_4_array, axis=1).shape)
print(np.rollaxis(my_2_3_4_array, axis=2, start=1).shape)
# * use np.transpose to permute all the axes at once
# * use np.swapaxes to swap any two axes
# * use np.rollaxis to "rotate" the axes
# +
# np.moveaxis(a, source, destination)
# Move axes of an array to new positions.
# Other axes remain in their original order.
print(my_2_3_4_array.shape)
print('-----')
print(np.moveaxis(my_2_3_4_array, 0, -1).shape)
print(np.moveaxis(my_2_3_4_array, -1, 0).shape)
# -
# ## Universal Functions
#
# * [Info](https://docs.scipy.org/doc/numpy/reference/ufuncs.html)
# truncated binomial: returns (x+1) ** 3 - (x) ** 3
def truncated_binomial(x):
return (x+1) ** 3 - (x) ** 3
np.testing.assert_equal(truncated_binomial(4), 61)
np.testing.assert_equal(truncated_binomial(4), 65)
my_numpy_function = np.frompyfunc(truncated_binomial, 1, 1)
my_numpy_function
test_array = np.arange(10)
my_numpy_function(test_array)
big_test_array = np.outer(test_array, test_array)
big_test_array
my_numpy_function(big_test_array)
# * pythogorean triplets
#
# $X^n + Y^n = Z ^n$
def is_integer(x):
return np.equal(np.mod(x, 1), 0)
numpy_is_integer = np.frompyfunc(is_integer, 1, 1)
# +
number_of_triangles = 9
base = np.arange(number_of_triangles) + 1
height = np.arange(number_of_triangles) + 1
# https://docs.scipy.org/doc/numpy/reference/generated/numpy.ufunc.outer.html
hypotenuse_squared = np.add.outer(base ** 2, height ** 2)
hypotenuse = np.sqrt(hypotenuse_squared)
numpy_is_integer(hypotenuse)
# -
# Another method
#
# for $m$ and $n$ $+ve$ integers, and m $\geq$ n:
# $X = m^2 - n^2; Y= 2mn; Z = m^2 + n^2$
|
numpy/Numpy basic tutorial.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python3
# name: python3
# ---
# # 17. Modern Regularization Techniques
# We are now going to look at modern regularization techniques such as drop out. Dropout is a technique developed by <NAME> and team. Up until we have worked with the following types of regularization:
#
# <br>
# ## 1.1 L1 and L2 Regularization
# **L1 regularization** encourages weights to be sparse, aka most weights being equal to 0. **L2 regularization** encourages most weights to be small, aka approximately equal to 0.
#
# $$L_{regularized} = L_{likelihood} + \lambda|\theta|^p$$
#
# <br>
# ## 1.2 Dropout
# In contrast to L1 and L2 regularization, **Dropout** does not add any penalty term to the cost. Instead, it works in a totally different way. Specifically, dropout works by dropping random nodes in the neural network during training. This has the effect of making it so that any hidden layer unit cannot just rely on 1 input feature, because at any time that node could be dropped.
#
# <img src="https://drive.google.com/uc?id=171gI21cZhPRGgA2q67nszI6gdx15Ikn3" width="500">
#
# We will see that dropout emulates an ensemble of neural networks. What exactly is meant by ensemble? Well, remember how we mentioned we would be dropping some nodes in the neural network. We can imagine that instead of just randomly dropped nodes during training, we could actually create several instances of neural networks with these different structures and train them all. Then, to calculate the final prediction, we could average the predictions of each individual neural network.
#
# <br>
# **Pseudocode**
# ```
# prediction1 = neuralNetwork1.predict(X)
# prediction2 = neuralNetwork2.predict(X)
# prediction3 = neuralNetwork3.predict(X)
#
# finalPrediction = mean(prediction1, prediction2, prediction3)
# ```
# ---
#
# <br>
# # 2. Dropout
# We are now going to dig further to see exactly how dropping nodes randomly in a neural network performs regularization, and how it emulates an ensemble of neural networks.
#
# <br>
# ### 2.0 Ensembles
# First, let's quickly discuss ensembles. The basic idea is that by using a group of prediction models that are different, and then taking the average or a majority vote we can end up with better accuracy than if we had just used 1 prediction model. So, what do we mean by different?
#
# <br>
# **Method 1**
# <br>
# One easy way is to just train on different subsets of random data. This is also good if your algorithm doesn't scale. As an example, say you have 1 million training points, but you train 10 different versions of a decision tree on only 100,000 points, then that would be an ensemble of decision trees. Then, to get a prediction you take a majority vote from these 10 different decision trees.
#
# <br>
# **Method 2**
# <br>
# Another method is to not use all of the features. So, if we have 100 features, maybe each of the 10 decision trees will only look at 10 different features each. The result is that instead of training 1 decision tree on a 1 million x 100 matrix X matrix, we train 10 decision trees on 100k x 10 matrices, which are all sampled from the original matrix. Miraculously, this results in better performance that just training 1 decision tree. Dropout is more like this method.
#
# ---
#
# <br>
# ## 2.1 Dropout
# So, how exactly does dropout work? Well, as we said, we are only going to use a subset of features. However, we are not only going to do this at the input layer, we are going to do this at every layer. At every layer, we are going to choose randomly which nodes to drop. We use a probability $p(drop)$ or $p(keep)$ to tell us the probability of dropping or keeping a node. Typical values of $p(keep)$ are 0.8 for the input layer, and 0.5 for the hidden layers.
#
# Note, we only drop layers during training. Also, notice that when we discussed ensembling there were 10 decision trees. However, with dropout there is still only 1 neural network. This is because we have only talked about training up until now. The other part is of course prediction.
#
# ### 2.1.1 Dropout - Prediction
# The way that we do prediction is that instead of dropping nodes, we multiply the output of a layer by its $p(keep)$. This effectively shrinks all of the values at that layer. In a way, that is similar to what L2 regularization does. It makes all of the weights smaller so that effectively all of the values are smaller.
#
# <br>
# **Pseudocode**
# <br>
# ```
# # Prediciton
# # If we have 1 hidden layer: X -> Y -> Z
# X_drop = p(keep | layer1) * X
# Z = f(X_drop.dot(W) + b)
# Z_drop = p(keep | layer2) * Z
# Y = softmax(Z_drop.dot(V) + c)
#
# # So we can see that we are multiplying by the p(keep) at that layer
# # This shrinks the value. L2 regularization also encourages weights to be small,
# # Leading to shrunken values
# ```
#
# Let's think about what this ensemble represents. If we have in total $N$ nodes in the neural network, that is just:
#
# $$N = number \; input \; nodes + number \; hidden \; nodes$$
#
# Each of these nodes can have 2 states: **On** or **Off**, **Drop** or **Keep**. So, that means in total we have:
#
# $$possible \; neural \; networks = 2^N$$
#
# Therefore, we are approximating an ensemble of $2^N$ different neural networks. Now, imagine the case where you were not doing an approximation. Let's take a very small neural network, only 100 nodes (N = 100). Keep in mind that is a very small neural net. For comparison, MNIST would have ~1000 nodes total; 700 for the input and 300 for the hidden layer. Anyways, if we only had 100 nodes, we would still have:
#
# $$2^{100} \approx 1.3 * 10^{30}$$
#
# Imagine training that many neural nets? It would clearly be infeasible. So, we can't actually do that ensemble, however, mutliplying by $p(keep)$ allows us to approximate it.
#
# ---
# <br>
# ## 2.2 Dropout - Implementation Theano
# The basic approach in theano to implement dropout, instead of actually dropping nodes out of the neural network, which would result in a different computational graph which theano wouldn't be able to handle, we are instead just going to multiply by 0. This has the same effect as dropping a node, because anything that comes after it will be mutliplied by 0 which is still 0. Since at each layer we are going to have an $N x D$ matrix, where N is equal to the batch size, we need to create an $N x D$ matrix of 0s and 1s to multiply that layer by. We call this matrix a **mask**.
#
# Now, recall that when you define the computational graph in theano you are not using real values. You are just specifying which nodes are connected to which other nodes. This means that we cannot multiply by a random numpy matrix in there, because that is effectively constant. If we randomly generate a 1 in the theano graph, and it is a numpy 1, then it is always going to be a 1 when we call the theano function to get the output of that graph. So, this would not work:
#
# <br>
# **Incorrect Pseudocode**
# <br>
# ```
# mask = np.random.binomial(...)
# Z = f((X * mask).dot(W) + b)
# ```
#
# Instead, what we want to do is have theano generate random numbers every time it passes through the graph. In order to do that we need an ojbect called random streams. Now, instead of passing an input directly to a hidden layer, we will first multiply it by a mask, which is equivalent to dropping some nodes. We then multiply by the weight and add the bias.
#
# <br>
# **Correct Pseudocode**
# <br>
# ```
# from theano.tensor.shared_randomstreams import RandomStreams
# rng = RandomStreams()
# mask = rng.binomial(n=1, p=p_keep, size=X.shape)
# X_drop = mask * X
# ```
#
# <br>
# ### 2.2.1 Dropout - Implementation Theano Code
# +
import numpy as np
import theano
import theano.tensor as T
import matplotlib.pyplot as plt
from theano.tensor.shared_randomstreams import RandomStreams # Import Random Streams
from util import get_normalized_data
from sklearn.utils import shuffle
"""
HiddenLayer class that knows nothing about dropout
"""
class HiddenLayer(object):
def __init__(self, M1, M2, an_id):
self.id = an_id
self.M1 = M1
self.M2 = M2
W = np.random.randn(M1, M2) * np.sqrt(2.0 / M1)
b = np.zeros(M2)
self.W = theano.shared(W, 'W_%s' % self.id)
self.b = theano.shared(b, 'b_%s' % self.id)
self.params = [self.W, self.b]
def forward(self, X):
return T.nnet.relu(X.dot(self.W) + self.b)
class ANN(object):
# Constructor now takes in p_keep - this is a list of probabilities since each layer
# can have its own probability of keeping
def __init__(self, hidden_layer_sizes, p_keep):
self.hidden_layer_sizes = hidden_layer_sizes
self.dropout_rates = p_keep
def fit(self, X, Y, learning_rate=1e-4, mu=0.9, decay=0.9, epochs=8,
batch_sz=100, show_fig=False):
# make a validation set
X, Y = shuffle(X, Y)
X = X.astype(np.float32)
Y = Y.astype(np.int32)
Xvalid, Yvalid = X[-1000:], Y[-1000:]
X, Y = X[:-1000], Y[:-1000]
self.rng = RandomStreams() # Creating object of type RandomStreams
# initialize hidden layers
N, D = X.shape
K = len(set(Y))
self.hidden_layers = []
M1 = D
count = 0
for M2 in self.hidden_layer_sizes:
h = HiddenLayer(M1, M2, count)
self.hidden_layers.append(h)
M1 = M2
count += 1
W = np.random.randn(M1, K) * np.sqrt(2.0 / M1)
b = np.zeros(K)
self.W = theano.shared(W, 'W_logreg')
self.b = theano.shared(b, 'b_logreg')
# collect params for later use
self.params = [self.W, self.b]
for h in self.hidden_layers:
self.params += h.params
# set up theano functions and variables
thX = T.matrix('X')
thY = T.ivector('Y')
pY_train = self.forward_train(thX) # Updates are defined using forward_train
# since dropping nodes occurs during training
# this cost is for training
cost = -T.mean(T.log(pY_train[T.arange(thY.shape[0]), thY]))
# gradients wrt each param
grads = T.grad(cost, self.params)
# for momentum
dparams = [theano.shared(np.zeros_like(p.get_value())) for p in self.params]
# for rmsprop
cache = [theano.shared(np.ones_like(p.get_value())) for p in self.params]
new_cache = [decay*c + (1-decay)*g*g for p, c, g in zip(self.params, cache, grads)]
new_dparams = [mu*dp - learning_rate*g/T.sqrt(new_c + 1e-10) for p, new_c, dp, g in zip(self.params, new_cache, dparams, grads)]
updates = [
(c, new_c) for c, new_c in zip(cache, new_cache)
] + [
(dp, new_dp) for dp, new_dp in zip(dparams, new_dparams)
] + [
(p, p + new_dp) for p, new_dp in zip(self.params, new_dparams)
]
# momentum only
# updates = [
# (p, p + mu*dp - learning_rate*T.grad(cost, p)) for p, dp in zip(self.params, dparams)
# ] + [
# (dp, mu*dp - learning_rate*T.grad(cost, p)) for p, dp in zip(self.params, dparams)
# ]
train_op = theano.function(
inputs=[thX, thY],
updates=updates
)
# for evaluation and prediction
pY_predict = self.forward_predict(thX)
cost_predict = -T.mean(T.log(pY_predict[T.arange(thY.shape[0]), thY]))
prediction = self.predict(thX)
cost_predict_op = theano.function(inputs=[thX, thY], outputs=[cost_predict, prediction])
n_batches = N // batch_sz
costs = []
for i in range(epochs):
X, Y = shuffle(X, Y)
for j in range(n_batches):
Xbatch = X[j*batch_sz:(j*batch_sz+batch_sz)]
Ybatch = Y[j*batch_sz:(j*batch_sz+batch_sz)]
train_op(Xbatch, Ybatch)
if j % 20 == 0:
c, p = cost_predict_op(Xvalid, Yvalid)
costs.append(c)
e = error_rate(Yvalid, p)
print("i:", i, "j:", j, "nb:", n_batches, "cost:", c, "error rate:", e)
if show_fig:
plt.plot(costs)
plt.show()
"""
Instead of forward function, we now have forward train function, and forward
predict function. This is because dropout acts differently during training and prediction.
During forward_train nodes are dropped. During forward_predict nodes are not dropped.
"""
def forward_train(self, X):
"""
We are generating a mask using the random streams object. We pass in n=1, meaning it
is like doing one coin flip, the probability of 1 is p_keep, and the size should be
the same size as what we are about to multiply by. This is done at every layer, and
then we do the softmax layer.
"""
Z = X
for h, p in zip(self.hidden_layers, self.dropout_rates[:-1]):
mask = self.rng.binomial(n=1, p=p, size=Z.shape)
Z = mask * Z
Z = h.forward(Z)
mask = self.rng.binomial(n=1, p=self.dropout_rates[-1], size=Z.shape)
Z = mask * Z
return T.nnet.softmax(Z.dot(self.W) + self.b)
def forward_predict(self, X):
"""
Here there is no mask. We just multiply each layer by its p_keep value. This is why
it is convenient to use the probability of keep instead of the probability of drop.
Since we can use p_keep directly in all cases.
"""
Z = X
for h, p in zip(self.hidden_layers, self.dropout_rates[:-1]):
Z = h.forward(p * Z)
return T.nnet.softmax((self.dropout_rates[-1] * Z).dot(self.W) + self.b)
def predict(self, X):
pY = self.forward_predict(X)
return T.argmax(pY, axis=1)
def error_rate(p, t):
return np.mean(p != t)
def relu(a):
return a * (a > 0)
def main():
# step 1: get the data and define all the usual variables
X, Y = get_normalized_data()
ann = ANN([500, 300], [0.8, 0.5, 0.5])
ann.fit(X, Y, show_fig=True)
if __name__ == '__main__':
main()
# -
# ---
#
# <br>
# # 3. Dropout - Implementation TensorFlow Code
# +
import numpy as np
import tensorflow as tf
import matplotlib.pyplot as plt
from util import get_normalized_data
from sklearn.utils import shuffle
"""HiddenLayer class that knows nothing about dropout"""
class HiddenLayer(object):
def __init__(self, M1, M2):
self.M1 = M1
self.M2 = M2
W = np.random.randn(M1, M2) * np.sqrt(2.0 / M1)
b = np.zeros(M2)
self.W = tf.Variable(W.astype(np.float32))
self.b = tf.Variable(b.astype(np.float32))
self.params = [self.W, self.b]
def forward(self, X):
return tf.nn.relu(tf.matmul(X, self.W) + self.b)
"""ANN class takes in p_keep. Everything proceeds as normal until we get to forward train"""
class ANN(object):
def __init__(self, hidden_layer_sizes, p_keep):
self.hidden_layer_sizes = hidden_layer_sizes
self.dropout_rates = p_keep
def fit(self, X, Y, lr=1e-4, mu=0.9, decay=0.9, epochs=15, batch_sz=100, split=True, print_every=20):
# make a validation set
X, Y = shuffle(X, Y)
X = X.astype(np.float32)
Y = Y.astype(np.int64)
if split:
Xvalid, Yvalid = X[-1000:], Y[-1000:]
X, Y = X[:-1000], Y[:-1000]
else:
Xvalid, Yvalid = X, Y
# initialize hidden layers
N, D = X.shape
K = len(set(Y))
self.hidden_layers = []
M1 = D
for M2 in self.hidden_layer_sizes:
h = HiddenLayer(M1, M2)
self.hidden_layers.append(h)
M1 = M2
W = np.random.randn(M1, K) * np.sqrt(2.0 / M1)
b = np.zeros(K)
self.W = tf.Variable(W.astype(np.float32))
self.b = tf.Variable(b.astype(np.float32))
# collect params for later use
self.params = [self.W, self.b]
for h in self.hidden_layers:
self.params += h.params
# set up theano functions and variables
inputs = tf.placeholder(tf.float32, shape=(None, D), name='inputs')
labels = tf.placeholder(tf.int64, shape=(None,), name='labels')
logits = self.forward(inputs)
"""Remember that tensorflow does softmax for us, so we only need to get logits"""
cost = tf.reduce_mean(
tf.nn.sparse_softmax_cross_entropy_with_logits(
logits=logits,
labels=labels
)
)
train_op = tf.train.RMSPropOptimizer(lr, decay=decay, momentum=mu).minimize(cost)
# train_op = tf.train.MomentumOptimizer(lr, momentum=mu).minimize(cost)
# train_op = tf.train.AdamOptimizer(lr).minimize(cost)
prediction = self.predict(inputs)
# validation cost will be calculated separately since nothing will be dropped
test_logits = self.forward_test(inputs)
test_cost = tf.reduce_mean(
tf.nn.sparse_softmax_cross_entropy_with_logits(
logits=test_logits,
labels=labels
)
)
n_batches = N // batch_sz
costs = []
init = tf.global_variables_initializer()
with tf.Session() as session:
session.run(init)
for i in range(epochs):
print("epoch:", i, "n_batches:", n_batches)
X, Y = shuffle(X, Y)
for j in range(n_batches):
Xbatch = X[j*batch_sz:(j*batch_sz+batch_sz)]
Ybatch = Y[j*batch_sz:(j*batch_sz+batch_sz)]
session.run(train_op, feed_dict={inputs: Xbatch, labels: Ybatch})
if j % print_every == 0:
c = session.run(test_cost, feed_dict={inputs: Xvalid, labels: Yvalid})
p = session.run(prediction, feed_dict={inputs: Xvalid})
costs.append(c)
e = error_rate(Yvalid, p)
print("i:", i, "j:", j, "nb:", n_batches, "cost:", c, "error rate:", e)
plt.plot(costs)
plt.show()
"""
Big difference compared to Theano. We utilize a built in function called
dropout, takes in input matrix and probability of keep
"""
def forward(self, X):
# tf.nn.dropout scales inputs by 1/p_keep
# therefore, during test time, we don't have to scale anything
Z = X
Z = tf.nn.dropout(Z, self.dropout_rates[0])
for h, p in zip(self.hidden_layers, self.dropout_rates[1:]):
Z = h.forward(Z)
Z = tf.nn.dropout(Z, p)
return tf.matmul(Z, self.W) + self.b
def forward_test(self, X):
Z = X
for h in self.hidden_layers:
Z = h.forward(Z)
return tf.matmul(Z, self.W) + self.b
def predict(self, X):
pY = self.forward_test(X)
return tf.argmax(pY, 1)
def error_rate(p, t):
return np.mean(p != t)
def relu(a):
return a * (a > 0)
def main():
# step 1: get the data and define all the usual variables
X, Y = get_normalized_data()
ann = ANN([500, 300], [0.8, 0.5, 0.5])
ann.fit(X, Y)
if __name__ == '__main__':
main()
# -
# # 4. Dropout Intuition
# We may ask the question at some point: "How does multiply by $p(keep)$ emulate an ensemble of neural networks where the probability of dropping a node is $1 - p(keep)$?"
#
# <br>
# Well, lets consider a portion of a neural network where there are 3 nodes on the left feeding into 1 node on the right. In other words we have 3 inputs and 1 output. Let's say that $p(keep)$ is 2/3 and all of the weights are 1. We will also say that all of the value sat the input are 1 as well, ($x_1 = x_2 = x_3 = 1$).
#
# <img src="https://drive.google.com/uc?id=1VC1nRR7NxkTVh0nWuX9O2Zvl94oJwSIT" width="300">
#
# The question is, how many possible configurations are there? Well, we have 3 nodes, each with a possible state of being kept or being dropped, hence have 8 different possible combinations:
#
# $$2^3 = 8$$
#
# We can see these possibilities below (the green means on and the white means off).
#
# <img src="https://drive.google.com/uc?id=1svJk9d_JcqpJ1G6WEUk7oRyNwoghOf61" width="700">
#
# What is important to realize here is that the probability of each of these configurations is not equivalent. In other words, each state does not have a probability of 1/8 of occuring. The probability of a node being on is 2/3, so the probability of a node being off is 1/3.
#
# $$p(drop) = 1 - p(keep) = \frac{1}{3}$$
#
# Therefore, if you have a configuration where all nodes are off, then it is:
#
# <br>
# $$p(drop)p(drop)p(drop) = \frac{1}{3}*\frac{1}{3}*\frac{1}{3} = \Big(\frac{1}{3}\Big)^3 = \frac{1}{27}$$
#
# If you have a configuration where 1 node is on and 2 nodes are off:
#
# <br>
# $$p(drop)p(keep)p(drop) = \frac{1}{3}*\frac{2}{3}*\frac{1}{3} = \frac{2}{27}$$
#
# We can see each combinations probability below:
#
# <img src="https://drive.google.com/uc?id=17lWuvNAjTJ6M1rnyETOfJelpsSX2k_ai" width="700">
#
# Now, for each of these configurations, it is easy to calculate the output value. Since we have assumed all of the inputs and weights are 1:
#
# $$0 \; nodes \; are \; on: 0$$
# $$1 \; node \; is \; on: 1 * 1 = 1$$
# $$2 \; nodes \; are \; on: 1 * 1 + 1*1 = 2$$
# $$3 \; nodes \; are \; on: 1 * 1 + 1*1 + 1*1 = 3$$
#
# <br>
# ### 4.1 Calculating Ensemble Output
# We now have all that we need to calculate the ensemble output. Let's "pretend" that we used 27 base models in our ensemble, and each configuration showed up the expected number of times. For example, since the probability of all 3 nodes being on is 8/27, then 8 of our 27 base models had all three nodes on. So, the final model is just the average of all the base model outputs:
#
# $$Output = \frac{\Big(0*1 + 1*2 + 1*2 + 1*2 +2*4 + 2*4 + 2*4 + 3*8\Big)}{27} = 2$$
#
# However, keep in mind that this would require us to actually enumerate all 27 different configurations and get the output from each of them. The only reason it seemed simple here is because we used 1 for all of the different values.
#
# <br>
# ### 4.2 Expected Value
# BUT, we can of course compute this as an expected value as well! We can do this by using the number of occurences divided by 27. Recall, the expected value of a random variable X is:
#
# $$E(X) = \sum x*p(x)$$
#
# In other words we are summing over all possible values of x, and multiplying each possible value by its corresponding probability of occuring. So, in the case of our **expected ensemble output**:
# $$0*\frac{1}{27} + 1*\frac{2}{27} +1*\frac{2}{27} +1*\frac{2}{27} +2*\frac{4}{27}+2*\frac{4}{27}+2*\frac{4}{27}+3*\frac{8}{27} = 2 $$
#
# Now, this was a very long process. But imagine if you had just multiplied the inputs by $p(keep)$.
#
# $$Output = (\frac{2}{3} * 1)*1 + (\frac{2}{3} * 1)*1 + (\frac{2}{3} * 1)*1 = 2$$
#
# We can clearly see here that multiplying by p(keep) at the input and using all the weights, gives us the same answer as determining all of the possible configurations, calculating all of the possible outputs, and then finding their expected average.
# ---
#
# <br>
# # 5. Noise Injection
# We will now discuss another regularization technique, called **noise injection**. Now, instead of getting into a deep mathematical discussion, which is definitely possible, we will instead look at an intuitive picture.
#
# The way that noise injection works is very simple. Instead of training on just the regular batches of $x$s and $y$s, we add some gaussian noise to $x$. Everytime we call the train function, we add randomly generated noise to $x$. In code that may look like:
#
# <br>
# **Without Noise Injection**
# <br>
# ```
# train(X, Y)
# ```
#
# <br>
# **With Noise Injection**
# <br>
# ```
# noise ~ N(O, small noise variance) # Amount of variance is a hyperparameter
# train(X + noise, Y)
# ```
#
# A question that comes up though is: "Why does this help?" Well, we know that underfitting (high bias) occurs because the decision boundary is too simple, and doesn't capture the complexity of the real pattern. Overfitting (high variance) occurs because the decision boundary is too complex, and it looks like it is capturing random noise variations in the data.
#
# If we add gaussian noise to these data points, we can see that the decision boundary is forced to balance between the two sides. To do this it must become simpler, and have less twists and turns. It brings the boundary away from the overfitting situation, and closer to the underfitting situation. To be clear, neural networks are much more susceptible to the overfitting situation, because they have many parameters and are very flexible in what they can represent.
#
# <img src="https://drive.google.com/uc?id=1esOEz4qRzjKeCvJjCAO-oMYoZvSRnSym" width="300">
#
# Another way to to go about noise injection is to add noise to the weights. This will make the neural network more robust to small changes in the weights.
#
# <br>
# ** Instead of adding the noise to the inputs**
# <br>
# ```
# Y = f(X + noise; W)
# ```
#
# <br>
# **We add noise to the weights**
# <br>
# ```
# Y = f(X; W + noise)
# ```
|
Deep_Learning/02-Modern_Deep_Learning-10-Modern-Regularization-Techniques.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# # Extreme Gradient Boosting Classification using DMatrix
# This Code template is for the Classification task using XGBoost along with the DMatrix. DMatrix is an internal data structure that gets used by the XGBoost framework. It is optimized for both memory efficiency and training speed.
#
# <img src="https://cdn.blobcity.com/assets/gpu_recommended.png" height="25" style="margin-bottom:-15px" />
# ## Coming Soon
|
Classification/XGBoost/XGBoostClassifier_DMatrix.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:matching-docs]
# language: python
# name: conda-env-matching-docs-py
# ---
# + [markdown] raw_mimetype="text/restructuredtext"
# # Finding a solution to a stable marriage game
#
# In this tutorial we will be setting up and solving an instance of SM with Matching. In particular, we will be using an example adapted from *Pride and Prejudice* [[Aus13]](../../reference/bibliography.rst) where four women (Charlotte, Elizabeth, Jane and Lydia) are being courted by four male suitors (Bingley, Collins, Darcy, Wickham).
#
# From here on out, we'll refer to the men and women as suitors and reviewers, respectively. If you'd like to know more about SM then head to its [discussion page](../../discussion/stable_marriage/index.rst).
#
# ---
#
# To begin, we create an instance of the `Player` class for each suitor and reviewer:
# +
from matching import Player
suitors = [
Player(name="Bingley"),
Player(name="Collins"),
Player(name="Darcy"),
Player(name="Wickham"),
]
reviewers = [
Player(name="Charlotte"),
Player(name="Elizabeth"),
Player(name="Jane"),
Player(name="Lydia"),
]
# -
# Now each player requires have its preferences set using the `Player.set_prefs` method. Each player's preferences must be a list of all the `Player` instances in the other party and be ordered according to their preference of that player.
#
# A nice way to do this is by unpacking `suitors` and `reviewers`:
# +
bingley, collins, darcy, wickham = suitors
charlotte, elizabeth, jane, lydia = reviewers
bingley.set_prefs([jane, elizabeth, lydia, charlotte])
collins.set_prefs([jane, elizabeth, lydia, charlotte])
darcy.set_prefs([elizabeth, jane, charlotte, lydia])
wickham.set_prefs([lydia, jane, elizabeth, charlotte])
charlotte.set_prefs([bingley, darcy, collins, wickham])
elizabeth.set_prefs([wickham, darcy, bingley, collins])
jane.set_prefs([bingley, wickham, darcy, collins])
lydia.set_prefs([bingley, wickham, darcy, collins])
# -
# With our now complete `Player` instances, we pass the lists of players to the `StableMarriage` class and find a suitor-optimal, stable matching using the `solve` method:
# +
from matching.games import StableMarriage
game = StableMarriage(suitors, reviewers)
game.solve()
|
docs/tutorials/stable_marriage/main.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import math
print(tf.__version__)
# # 1. Load and Explore the Data
# overall_summary = pd.read_csv('overall_summary.csv')
data = pd.read_csv('prediction_data340.csv')
data
data[["percent_volume"]].boxplot()
data[["lag_1_CR","lag_12_CR", "lag_24_CR", "lag_36_CR","lag_48_CR"]].boxplot()
data['percent_volume'].plot(figsize=(18,6));
data['lag_1_CR'].plot(figsize=(18,6));
data['lag_24_CR'].plot(figsize=(18,6));
data['lag_48_CR'].plot(figsize=(18,6));
# ### Decompose Time-Series to see Individual Components (trend + seasonality + noise)
#
# https://coderzcolumn.com/tutorials/data-science/how-to-remove-trend-and-seasonality-from-time-series-data-using-python-pandas
# +
from statsmodels.tsa.seasonal import seasonal_decompose
decompose_result4 = seasonal_decompose(data['percent_volume'], period = 24*90, model="additive")
trend4 = decompose_result4.trend
seasonal4 = decompose_result4.seasonal
residual4 = decompose_result4.resid
decompose_result5 = seasonal_decompose(data['lag_1_CR'][1:], period = 24*90, model="additive")
trend5 = decompose_result5.trend
seasonal5 = decompose_result5.seasonal
residual5 = decompose_result5.resid
decompose_result6 = seasonal_decompose(data['lag_24_CR'][24:], period = 24*90, model="additive")
trend6 = decompose_result6.trend
seasonal6 = decompose_result6.seasonal
residual6 = decompose_result6.resid
decompose_result7 = seasonal_decompose(data['lag_48_CR'][48:], period = 24*90, model="additive")
trend7 = decompose_result7.trend
seasonal7 = decompose_result7.seasonal
residual7 = decompose_result7.resid
decompose_result4.plot()
decompose_result5.plot()
decompose_result6.plot()
decompose_result7.plot();
# -
# ### **Dicky-Fuller Test for Stationarity**
#
# - `p-value > 0.05`: This implies that time-series is non-stationary.
# - `p-value <=0.05`: This implies that time-series is stationary.
#
#
# +
from statsmodels.tsa.stattools import adfuller
dftest = adfuller(data['lag_1_CR'][1:], autolag = "BIC" )
print("1. ADF : ",dftest[0])
print("2. P-Value : ", dftest[1])
print("3. Num Of Lags : ", dftest[2])
print("4. Method to use when automatically determining the lag length : ", 'BIC' )
print("5. Num Of Observations Used For ADF Regression and Critical Values Calculation :", dftest[3])
print("6. Critical Values :")
for key, val in dftest[4].items():
print("\t",key, ": ", val)
# +
from statsmodels.tsa.stattools import adfuller
dftest = adfuller(data['lag_24_CR'][24:], autolag = "BIC" )
print("1. ADF : ",dftest[0])
print("2. P-Value : ", dftest[1])
print("3. Num Of Lags : ", dftest[2])
print("4. Method to use when automatically determining the lag length : ", 'BIC' )
print("5. Num Of Observations Used For ADF Regression and Critical Values Calculation :", dftest[3])
print("6. Critical Values :")
for key, val in dftest[4].items():
print("\t",key, ": ", val)
# +
from statsmodels.tsa.stattools import adfuller
dftest = adfuller(data['lag_48_CR'][48:], autolag = "BIC" )
print("1. ADF : ",dftest[0])
print("2. P-Value : ", dftest[1])
print("3. Num Of Lags : ", dftest[2])
print("4. Method to use when automatically determining the lag length : ", 'BIC' )
print("5. Num Of Observations Used For ADF Regression and Critical Values Calculation :", dftest[3])
print("6. Critical Values :")
for key, val in dftest[4].items():
print("\t",key, ": ", val)
# -
# ### Preparation
#
# - Define functions for the creation of rolling windows and visualizations for final results.
# - The output of rolling window function include both predictors and response variable
def windowed_dataset(series, window_size, batch_size, shuffle_buffer_size):
dataset = tf.data.Dataset.from_tensor_slices(series)
dataset = dataset.window(window_size+1, shift=1, drop_remainder=True)
dataset = dataset.flat_map(lambda window: window.batch(window_size+1)) # Convert each window to numpy format
dataset = dataset.shuffle(shuffle_buffer_size, seed = 123).map(lambda window: (window[:-1], window[-1])) # Shuffle data and Retrieve features and response variable
dataset = dataset.batch(batch_size).prefetch(1) # Combines consecutive elements of this dataset into batches
return dataset
def plot_series(time, series, title, format="-", start=0, end=None):
plt.plot(time[start:end], series[start:end], format)
plt.xlabel("Time")
plt.ylabel("Value")
plt.title(title)
plt.grid(True)
# # 2. MLP Model --- lag_24_CR Prediction_FL340
#
# - split the data to training set, validate set, and final test set
# - define some variables
# +
tf.keras.backend.clear_session()
tf.random.set_seed(1234) # Set the global random seed.
np.random.seed(1234)
# Retrieve the data
lag_n = 24 # Only Change this
col_name = 'lag_'+str(lag_n)+'_CR'
time = data['datetime_id']
time = time.to_numpy()[lag_n:]
CR_series = data[col_name]
CR_series = CR_series.to_numpy()[lag_n:]
volume_series =data['percent_volume']
volume_series = volume_series.to_numpy()[lag_n:]
# Split dataset
split_time1 = 3000
split_time2 = 4000
## Traing set
time_train = time[:split_time1]
x_train = CR_series[:split_time1]
## Validatiin set
time_valid = time[split_time1:split_time2]
x_valid = CR_series[split_time1:split_time2]
## Final test set
time_test = time[split_time2:]
x_test = CR_series[split_time2:]
window_size = 20 #Hyperparameter
batch_size = 30
shuffle_buffer_size = 3000
# -
# - **Modeling**
# - Generate rolling windows
# - Train the model
# +
dataset = windowed_dataset(series = x_train,
window_size = window_size,
batch_size = batch_size,
shuffle_buffer_size = shuffle_buffer_size)
print(dataset)
# Three layers in a sequential. Totally 4 layers exits. Here, we have two hidden layer.
# The first has five neurons activated by tanh.
# The second is the same, and the third is a single dense would provide the predicted value.
model = tf.keras.models.Sequential([
tf.keras.layers.Dense(units = 5, input_shape=[window_size], activation="tanh"), #Input layer, and first hidden layer # The number of nodes
# tf.keras.layers.Dense(units = 5, activation="tanh"), # Second hidden layer
tf.keras.layers.Dense(units = 1),
tf.keras.layers.Lambda(lambda x: x * 100) # scale_layer
])
model.compile(loss="mae", optimizer=tf.keras.optimizers.SGD(lr=1e-6, momentum=0.9))
model.fit(dataset,epochs=100,verbose=0)
# -
# - **Make Prediction**
# - Make prediction for the entire timeline (from timestamp window_size+1 to timestamp 5496): **forecast**
# - Retrieve the predicted value for validation set: **results_valid**
# - Retrieve the predicted value for train set: **results_train**
# - Retrieve the predicted value for test set: **results_test**
# +
forecast = []
for timestamp in range(len(volume_series) - window_size):
forecast.append(model.predict(CR_series[timestamp:timestamp + window_size][np.newaxis]))
# predicted value on training set
forecast_train = forecast[:split_time1-window_size]
results_train = np.array(forecast_train)[:, 0, 0]
# predicted value on validate set
forecast_valid = forecast[split_time1-window_size:split_time2-window_size]
results_valid = np.array(forecast_valid)[:, 0, 0]
# predicted value on test set
forecast_test = forecast[split_time2-window_size:]
results_test = np.array(forecast_test)[:, 0, 0]
# -
# - **Model Performance**
print("Static-MAE(test set) =", tf.keras.metrics.mean_absolute_error(CR_series[split_time2:], CR_series[split_time2-1:-1]).numpy())
print("Static-RMSE(test set) =", math.sqrt(tf.keras.metrics.mean_squared_error(CR_series[split_time2:], CR_series[split_time2-1:-1]).numpy()))
print("MLP-MAE(training set) =", tf.keras.metrics.mean_absolute_error(x_train[window_size:], results_train).numpy())
print("MLP-RMSE(training set) =", math.sqrt(tf.keras.metrics.mean_squared_error(x_train[window_size:], results_train).numpy()))
print("MLP-MAE(valid set) =", tf.keras.metrics.mean_absolute_error(x_valid, results_valid).numpy())
print("MLP-RMSE(valid set) =", math.sqrt(tf.keras.metrics.mean_squared_error(x_valid, results_valid).numpy()))
print("MLP-MAE(test set) =", tf.keras.metrics.mean_absolute_error(x_test, results_test).numpy())
print("MLP-RMSE(test set) =", math.sqrt(tf.keras.metrics.mean_squared_error(x_test, results_test).numpy()))
# Parameter Tuning:
#
# |Flight Level|Window_size|# Hidden layer|MAE(valid set)|RMSE(valid set)|
# |----|-----------|---|--------------|---------------|
# |FL340|20|1|59.75264|87.40495954535417|
# |FL360|20|1|40.2134|57.680903858399056|
# |FL340|24|1|60.774666|87.81635334072179|
# |FL360|24|1|38.21996|55.41995212691906|
# |FL340|28|1|63.25586|90.454567649145|
# |FL360|28|1|41.93651|59.433916082144336|
# |FL340|20|2|64.293915|92.48989598531831|
# |FL360|20|2|40.31757|57.70645891397795|
# |FL340|24|2|61.33682|88.98613776787792|
# |FL360|24|2|39.572437|57.26746245346807|
# |FL340|28|2|61.43381|87.5719542317559|
# |FL360|28|2|40.537086|56.954342408113405|
# - **Visualization for predicted CR**
# +
plt.figure(figsize=(18, 6))
plot_series(time_test, x_test, title = "Plot for test set (Blue: observed; Orange: Predicted)")
plot_series(time_test, results_test, title = "Plot for test set (Blue: observed; Orange: Predicted)");
# Blue: x_valid (observed); Orange: results_valid(predicted)
plt.figure(figsize=(18, 6))
plot_series(time_train[window_size:], x_train[window_size:], title = "Plot for training set (Blue: observed; Orange: Predicted)")
plot_series(time_train[window_size:], results_train, title = "Plot for training set (Blue: observed; Orange: Predicted)");
# -
# - **Convert CR to percent_volume**
# +
lag_volume = volume_series[split_time2-lag_n:-lag_n]
pred_volume_test = lag_volume+np.exp(results_test*0.01)
print("Static-MAE(test set) =", tf.keras.metrics.mean_absolute_error(volume_series[split_time2:], volume_series[split_time2-lag_n:-lag_n]).numpy())
print("Static-RMSE(test set) =", math.sqrt(tf.keras.metrics.mean_squared_error(volume_series[split_time2:], volume_series[split_time2-lag_n:-lag_n]).numpy()))
print("MLP-MAE(test set) =", tf.keras.metrics.mean_absolute_error(volume_series[split_time2:], pred_volume_test).numpy())
print("MLP-RMSE(test set) =", math.sqrt(tf.keras.metrics.mean_squared_error(volume_series[split_time2:], pred_volume_test).numpy()))
plt.figure(figsize=(18, 6))
plot_series(time_test, volume_series[split_time2:], title = "Plot for test set (Blue: observed; Orange: Predicted)")
plot_series(time_test, pred_volume_test, title = "Plot for test set (Blue: observed; Orange: Predicted)");
# -
|
4 Prediction/4 fl340 prediction/MLP-Lag24-Prediction-FL340.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from bayes_tec.plotting.plot_datapack import animate_datapack, plot_data_vs_solution, plot_freq_vs_time
plot_freq_vs_time('../../scripts/data/killms_datapack.hdf5','../../scripts/data/killms_freq_time')
plot_freq_vs_time('../../scripts/data/killms_datapack_2.hdf5','../../scripts/data/killms_2_freq_time')
plot_freq_vs_time('../../scripts/data/killms_datapack_3.hdf5','../../scripts/data/killms_3_freq_time')
plot_freq_vs_time('../../scripts/data/killms_datapack_nchan4.hdf5','../../scripts/data/killms_nchan4_freq_time')
plot_freq_vs_time('../../scripts/data/ndppp_datapack.hdf5','../../scripts/data/ndppp_freq_time')
animate_datapack('../../scripts/data/killms_datapack_2.hdf5','killms_data_2d_18_chol_screen',32,labels_in_radec=True,plot_screen=True, phase_wrap=True,solset='screen_posterior_sol_18_chol',observable='tec', tec_eval_freq=140e6, pol_sel=slice(0,1,1),ant_sel='RS*',time_sel=slice(0,600,1),freq_sel=slice(25,26,1),plot_facet_idx=False)
plot_data_vs_solution('../../scripts/data/killms_datapack_2.hdf5',output_folder='killms_data_plots_ml_2',solution_solset='sol000',pol_sel=slice(0,1,1),ant_sel='RS210HBA',time_sel=slice(0,1300,1),freq_sel=slice(25,26,1))
import h5py
with h5py.File('ml_results.hdf5') as f:
tec = f['tec'][...]
sigma = f['sigma'][...]
import pylab as plt
import numpy as np
times = np.linspace(0,3600*8, 3600)
phase = np.angle(np.exp(1j*(tec[0,1,51,:]*-8.448e9/140e6)))
plt.plot(times,phase)
plt.fill_between(times,phase-sigma[0,1,51,:],phase+sigma[0,1,51,:],alpha=0.5)
plt.show()
tec.shape
# +
import numpy as np
def log_normal_solve_fwhm(a,b,D=0.5):
assert b > a
lower = np.log(a)
upper = np.log(b)
d = upper - lower #2 sqrt(2 sigma**2 ln2)
sigma2 = 0.5*(0.5*d)**2/np.log(1./D)
s = upper + lower #2 (mu - sigma**2)
mu = 0.5*s + sigma2
return mu, np.sqrt(sigma2)
# -
from bayes_tec.utils.stat_utils import log_normal_solve
log_normal_solve_fwhm(0.5,1.5,0.09)
log_normal_solve(1.,0.5)
import pylab as plt
# %matplotlib inline
plt.hist(np.random.lognormal(*log_normal_solve_fwhm(25,75,D=.1),size=10000),bins=100)
plt.show()
plt.hist(np.random.lognormal(*log_normal_solve(50,25),size=10000),bins=100)
plt.show()
|
notebooks/devel/plot.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + language="javascript"
# IPython.OutputArea.auto_scroll_threshold = 9999;
# -
import argparse
import os
from solver import Solver
import glob
from torchvision import transforms, utils
import torchvision.transforms as standard_transforms
from torch.utils.data import Dataset, DataLoader
from own_dataloader import Rescale
from own_dataloader import RescaleT
from own_dataloader import RandomCrop
from own_dataloader import CenterCrop
from own_dataloader import ToTensor
from own_dataloader import ToTensorLab
from own_dataloader import SalObjDataset
# +
#data_dir = 'C:/Users/tip/Documents/GitHub/Saliency_Dataset/dataset_test/DUTOMRON/'
#data_dir = 'C:/Users/tip/Documents/GitHub/Saliency_Dataset/dataset_test/DUTS-TE/'
#data_dir = 'C:/Users/tip/Documents/GitHub/Saliency_Dataset/dataset_test/ECSSD/'
data_dir = 'C:/Users/tip/Documents/GitHub/Saliency_Dataset/dataset_test/HKU-IS/'
#data_dir = 'C:/Users/tip/Documents/GitHub/Saliency_Dataset/dataset_test/PASCALS/'
#data_dir = 'C:/Users/tip/Documents/GitHub/Saliency_Dataset/dataset_test/SOD/'
#tra_image_dir = 'DUTS-TR/DUTS-TR-Image/'
#tra_label_dir = 'DUTS-TR/DUTS-TR-Mask/'
test_image_dir = 'Imgs/'
test_label_dir = 'gt/'
enableInpaintAug = False
batch_size_train=32
batch_size_val=32
image_ext = '.jpg'
label_ext = '.png'
vgg_path = 'D:/nonat project/Experiment 1.2/weights/vgg16_feat.pth'
trained_model='D:/nonat project/Experiment 1.2/weights/Experiment1.2.pth'
#test_folder='D:/nonat project/Experiment 1.2/weights/test/DUTOMRON TEST'
#test_folder='D:/nonat project/Experiment 1.2/weights/test/DUTS TEST'
#test_folder='D:/nonat project/Experiment 1.2/weights/test/ECCSD TEST'
test_folder='D:/nonat project/Experiment 1.2/weights/test/HKU-IS TEST'
#test_folder='D:/nonat project/Experiment 1.2/weights/test/PASCAL TEST'
#test_folder='D:/nonat project/Experiment 1.2/weights/test/SOD TEST'
#output_path='D:/nonat project/Experiment 1.2/Experiment 1.2 Predictions/DUTS OMRON Saliency Map Prediction/'
#output_path='D:/nonat project/Experiment 1.2/Experiment 1.2 Predictions/DUTS Saliency Map Prediction/'
#output_path='D:/nonat project/Experiment 1.2/Experiment 1.2 Predictions/ECCSD Saliency Map Prediction/'
output_path='D:/nonat project/Experiment 1.2/Experiment 1.2 Predictions/HKU-IS Saliency Map Prediction/'
#output_path='D:/nonat project/Experiment 1.2/Experiment 1.2 Predictions/PASCAL Saliency Map Prediction/'
#output_path='D:/nonat project/Experiment 1.2/Experiment 1.2 Predictions/SOD Saliency Map Prediction/'
# -
def main(config):
if config.mode == 'train':
tra_img_name_list = glob.glob(data_dir + tra_image_dir + '*' + image_ext)
print("data_dir + tra_image_dir + '*' + image_ext: ", data_dir + tra_image_dir + '*' + image_ext)
tra_lbl_name_list = []
for img_path in tra_img_name_list:
img_name = img_path.split("\\")[-1]
aaa = img_name.split(".")
bbb = aaa[0:-1]
imidx = bbb[0]
for i in range(1,len(bbb)):
imidx = imidx + "." + bbb[i]
tra_lbl_name_list.append(data_dir + tra_label_dir + imidx + label_ext)
print("---")
print("train images: ", len(tra_img_name_list))
print("train labels: ", len(tra_lbl_name_list))
print("---")
train_num = len(tra_img_name_list)
salobj_dataset = SalObjDataset(
img_name_list=tra_img_name_list,
lbl_name_list=tra_lbl_name_list,
transform=transforms.Compose([
RescaleT(256),
RandomCrop(224),
ToTensorLab(flag=0)]),
category="train",
enableInpaintAug=enableInpaintAug)
test_img_name_list = glob.glob(data_dir + test_image_dir + '*' + image_ext)
print("data_dir + test_image_dir + '*' + image_ext: ", data_dir + test_image_dir + '*' + image_ext)
test_lbl_name_list = []
for img_path in test_img_name_list:
img_name = img_path.split("\\")[-1]
aaa = img_name.split(".")
bbb = aaa[0:-1]
imidx = bbb[0]
for i in range(1,len(bbb)):
imidx = imidx + "." + bbb[i]
test_lbl_name_list.append(data_dir + test_label_dir + imidx + label_ext)
print("---")
print("test images: ", len(test_img_name_list))
print("test labels: ", len(test_lbl_name_list))
print("---")
test_num = len(test_img_name_list)
salobj_dataset_test = SalObjDataset(
img_name_list=test_img_name_list,
lbl_name_list=test_lbl_name_list,
transform=transforms.Compose([
RescaleT(256),
RandomCrop(224),
ToTensorLab(flag=0)]),
category="test",
enableInpaintAug=enableInpaintAug)
if config.mode == 'train':
train_loader = DataLoader(salobj_dataset, batch_size=config.batch_size, shuffle=True, num_workers=1)
if config.val:
val_loader = DataLoader(salobj_dataset_test, batch_size=config.batch_size_val, shuffle=True, num_workers=1)
run = 0
while os.path.exists("%s/run-%d" % (config.save_fold, run)): run += 1
os.mkdir("%s/run-%d" % (config.save_fold, run))
os.mkdir("%s/run-%d/logs" % (config.save_fold, run))
# os.mkdir("%s/run-%d/images" % (config.save_fold, run))
os.mkdir("%s/run-%d/models" % (config.save_fold, run))
config.save_fold = "%s/run-%d" % (config.save_fold, run)
if config.val:
train = Solver(train_loader, val_loader, None, config)
else:
train = Solver(train_loader, None, None, config)
train.train(100)
elif config.mode == 'test':
test_loader = DataLoader(salobj_dataset_test, batch_size=batch_size_val, shuffle=True, num_workers=1)
if not os.path.exists(config.test_fold): os.mkdir(config.test_fold)
test = Solver(None, None, test_loader, config)
test.test(100,output_path, use_crf=config.use_crf)
else:
raise IOError("illegal input!!!")
# +
if __name__ == '__main__':
data_root = os.path.join(os.path.expanduser('~'), 'data')
parser = argparse.ArgumentParser()
# Hyper-parameters
parser.add_argument('--n_color', type=int, default=3)
parser.add_argument('--img_size', type=int, default=256) # 256
parser.add_argument('--lr', type=float, default=1e-6)
parser.add_argument('--clip_gradient', type=float, default=1.0)
parser.add_argument('--cuda', type=bool, default=False)
# Training settings
parser.add_argument('--vgg', type=str, default=vgg_path)
parser.add_argument('--epoch', type=int, default=500)
parser.add_argument('--val', type=bool, default=True)
parser.add_argument('--num_thread', type=int, default=4)
parser.add_argument('--load', type=str, default='')
parser.add_argument('--save_fold', type=str, default='./results')
parser.add_argument('--epoch_val', type=int, default=10)
parser.add_argument('--batch_size', type=int, default=batch_size_train)
parser.add_argument('--batch_size_val', type=int, default=batch_size_val)
parser.add_argument('--epoch_save', type=int, default=10)
parser.add_argument('--epoch_show', type=int, default=1)
parser.add_argument('--pre_trained', type=str, default=None)
# Testing settings
parser.add_argument('--model', type=str, default=trained_model)
parser.add_argument('--test_fold', type=str, default='./results/test')
parser.add_argument('--use_crf', type=bool, default=False)
# Misc
parser.add_argument('--mode', type=str, default='train', choices=['train', 'test'])
parser.add_argument('--visdom', type=bool, default=False)
import easydict
config = easydict.EasyDict({
"n_color": 3,
"img_size":256,
"lr":1e-6,
"clip_gradient":1.0,
"cuda":True,
"vgg":vgg_path,
"epoch":500,
"val":True,
"num_thread":4,
"load":'',
"save_fold":'./results',
"epoch_val":10,
"batch_size":batch_size_train,
"batch_size_val":batch_size_val,
"epoch_save":10,
"epoch_show":1,
"pre_trained":None,
"model":trained_model,
"test_fold":'./results/test',
"use_crf":False,
"mode":'test',
"visdom":False,
})
if not os.path.exists(config.save_fold): os.mkdir(config.save_fold)
main(config)
|
Experiment2_HKU-IS.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="sblS7n3zWCWV"
# **Copyright 2019 The TensorFlow Authors.**
# + colab={} colab_type="code" id="0rvUzWmoWMH5"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="aCZBFzjClURz"
# # 텐서플로우 모델 생성 및 변환
# 이 노트북을 통해 TensorFlow 모델을 작성한 뒤 어떻게 TensorFlow Lite 모델로 변환하는지를 살펴볼 것입니다. 이 노트북에서 생성된 모델은 [TensorFlow Lite for Microcontrollers](https://www.tensorflow.org/lite/microcontrollers/overview)의 [hello_world](https://github.com/tensorflow/tensorflow/tree/master/tensorflow/lite/micro/examples/hello_world)샘플에서도 사용됩니다.
#
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/tensorflow/blob/master/tensorflow/lite/micro/examples/hello_world/create_sine_model.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/tensorflow/blob/master/tensorflow/lite/micro/examples/hello_world/create_sine_model.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# </table>
#
# + [markdown] colab_type="text" id="dh4AXGuHWeu1"
# ## 종속성 로드
# 가장 먼저 해야할 일은 우리가 필요한 종속성을 로드하는 것이다. 아래의 셀을 실행해보자.
# + colab={"base_uri": "https://localhost:8080/", "height": 479} colab_type="code" id="53PBJBv1jEtJ" outputId="9b035753-60e5-43db-a78d-284ea9de9513"
# TensorFlow는 오픈소스 라이브러리다.
# 참조: 아래의 코드는 텐서플로우 2버전을 사용함.
# !pip install tensorflow==2.0.0-beta0
import tensorflow as tf
# 넘파이(Numpy)는 수학 연산에 추가적으로 필요한 라이브러리이다.
import numpy as np
# Matplotlib은 그래프를 그리는데 필요한 라이브러리이다.
import matplotlib.pyplot as plt
# math는 파이썬의 수학 라이브러리이다.
import math
# + [markdown] colab_type="text" id="p-PuBEb6CMeo"
# ## 데이터 생성하기
# 딥러닝 네트워크는 데이터에서 패턴을 학습한다. 여기서는 [싸인함수](https://en.wikipedia.org/wiki/Sine)로 생성된 데이터를 모델링하기 위해 네트워크를 훈련한다. 훈련을 마치면 값`x`를 입력받아 `y`를 예측할 수있는 모델이 생성되는데, 사실 우리는 이미 싸인함수 공식을 알고 있기 때문에 필요하면 직접 계산할 수 있다. 그렇지만 지금은 모델 훈련을 위한 데이터가 어떤 패턴을 지니고 있는지 모르는 상태에서 모델을 훈련한다고 하자.
#
# TensorFlow Lite for Microcontrollers의 hello_world 샘플의 경우, 이 모델을 사용하여 LED가 순서대로 점등되도록 제어한다.
#
# 다음 셀의 코드는 임의의 `x`값 세트를 생성하고 사인 값을 계산하여 그래프에 표시한다:
# + colab={"base_uri": "https://localhost:8080/", "height": 269} colab_type="code" id="uKjg7QeMDsDx" outputId="b17a43c6-eba1-4cc7-8807-14fcf5918d01"
# 아래의 값만큼 데이터 샘플을 생성할 것이다.
SAMPLES = 1000
# 시드 값을 지정하여 이 노트북에서 실행할 때마다 다른 랜덤 값을 얻게 한다.
# 어떤 숫자든 사용할 수 있다.
np.random.seed(1337)
# 사인파 진폭의 범위인 0~2π 내에서 균일하게 분포된 난수 집합을 생성한다.
x_values = np.random.uniform(low=0, high=2*math.pi, size=SAMPLES)
# 값을 섞어서 생성된 값들이 순서를 따르지 않도록 한다.
np.random.shuffle(x_values)
# 해당 사인값을 계산한다
y_values = np.sin(x_values)
# 데이터를 그래프로 그린다. 'b.' 인수는 라이브러리에 점을 파란색으로 출력하도록 지시한다.
plt.plot(x_values, y_values, 'b.')
plt.show()
# + [markdown] colab_type="text" id="iWOlC7W_FYvA"
# ## 노이즈 추가
#
# 사인 함수에 의해 직접 생성되었으므로 데이터는 훌륭하고 부드러운 곡선으로 나타난다.
#
# 그러나 머신 러닝 모델은 보다 복잡한 실제 데이터에서 패턴을 알아낼 수 있다. 이를 위해 데이터에 약간의 노이즈를 추가하여 보다 실제와 비슷한 데이터를 만들어보자.
#
# 다음 셀에서는 각 값에 임의의 노이즈를 추가 한 다음 새 그래프를 그린다.
# + colab={"base_uri": "https://localhost:8080/", "height": 269} colab_type="code" id="i0FJe3Y-Gkac" outputId="60b19cdd-c69c-469e-9446-b738a79c1f51"
# 각 y 값에 임의의 작은 숫자를 추가한다.
y_values += 0.1 * np.random.randn(*y_values.shape)
# 그래프를 생성한다.
plt.plot(x_values, y_values, 'b.')
plt.show()
# + [markdown] colab_type="text" id="Up8Xk_pMH4Rt"
# ## 데이터 분할
# 이제 실제 데이터와 비슷한 노이즈가 추가된 데이터 세트가 생성되었다. 우리는 이 데이터를 모델을 훈련시키기 위해 사용할 것입니다.
#
# 우리가 훈련시키는 모델의 정확성을 평가하려면 예측을 실제 데이터와 비교하고 일치하는 정도를 확인해야한다. 이 평가는 훈련중(검증이라고 함)과 훈련후(테스트라고 함)발생한다. 두 경우 모두 모델 훈련에 사용되지 않은 새로운 데이터를 사용하는 것이 중요하다.
#
# 평가에 사용할 데이터를 확보하기 위해 훈련을 시작하기 전에 따로 분할하도록 하자. 검증셋에 20%, 테스트셋에 20%를 미리 할당해두자. 나머지 60%는 모델 학습에 사용된다. 이는 모델을 훈련 할 때 사용되는 일반적인 데이터 분할이다.
#
# 다음 코드는 데이터를 분할 한 다음 각 세트를 다른 색상으로 그래프에 나타낸다.
# + colab={"base_uri": "https://localhost:8080/", "height": 269} colab_type="code" id="nNYko5L1keqZ" outputId="b9f9c57b-b6aa-4817-8ab4-4a2201732b9a"
# 훈련에 60%, 테스트에 20%, 검증에 20%를 사용한다.
# 각 항목의 인덱스를 계산한다.
TRAIN_SPLIT = int(0.6 * SAMPLES)
TEST_SPLIT = int(0.2 * SAMPLES + TRAIN_SPLIT)
# np.split을 사용하여 데이터를 세 부분으로 자른다.
# np.split의 두 번째 인수는 데이터가 분할되는 인덱스 배열이며,
# 우리는 두 개의 인덱스를 제공하므로 데이터는 세 개의 덩어리로 나뉠 것이다.
x_train, x_test, x_validate = np.split(x_values, [TRAIN_SPLIT, TEST_SPLIT])
y_train, y_test, y_validate = np.split(y_values, [TRAIN_SPLIT, TEST_SPLIT])
# 분할한 데이터를 합쳤을 때 원래의 사이즈와 같은지 재확인한다.
assert (x_train.size + x_validate.size + x_test.size) == SAMPLES
# 분할된 각 데이터들을 다른 색상으로 그래프에 표시한다.
plt.plot(x_train, y_train, 'b.', label="Train")
plt.plot(x_test, y_test, 'r.', label="Test")
plt.plot(x_validate, y_validate, 'y.', label="Validate")
plt.legend()
plt.show()
# + [markdown] colab_type="text" id="t5McVnHmNiDw"
# ## 모델 설계
# 지금부터 입력값(`x`)을 받아 출력값(`x`의 사인)을 예측하는 모델을 만들 것이다. 이러한 유형의 문제를 회귀_regression_라고 한다.
#
# 이를 위해 간단한 신경망을 만들텐데, 뉴런_neurons_과 레이어_layers_를 사용하여 훈련 데이터의 기본적인 패턴을 학습을 통해 예측을 수행해보자.
#
# 먼저 두 개의 레이어를 정의하자. 첫 번째 레이어는 단일 입력(`x`값)을 가져와 16개의 뉴런을 통해 활성화된다. 이 입력에 따라 각 뉴런은 내부 상태 (가중치_weight_ 및 편향_bias_값)에 따라 어느정도까지 활성화_activated_된다. 뉴런의 활성화 정도는 숫자로 표현된다.
#
# 첫 번째 레이어의 활성화 정도는 두 번째 레이어에 입력으로 공급된다. 이 입력에 자체 가중치와 바이어스를 적용하고 활성화 정도를 계산하여 `y`값으로 출력한다.
#
# **참고:** 신경망의 기능에 대해 자세히 알아 보려면 [Learn TensorFlow](https://codelabs.developers.google.com/codelabs/tensorflow-lab1-helloworld) 코드 랩을 살펴보십시오.
#
# 다음 셀의 코드는 딥러닝 네트워크를 만들기위한 TensorFlow의 고수준 API인 [Keras](https://www.tensorflow.org/guide/keras)를 사용하여 모델을 정의한다. 일단 네트워크가 정의되면, 우리는 훈련 방법을 결정하는 파라미터를 지정하여 컴파일_compile_을 진행한다:
# + colab={} colab_type="code" id="gD60bE8cXQId"
# 간단한 모델 구조를 만들기 위해 케라스를 사용한다.
from tensorflow.keras import layers
model_1 = tf.keras.Sequential()
# 첫 번째 레이어는 16 개의 뉴런을 통해 스칼라Scalar 입력을 받아 다음 레이어에 전달한다.
# 뉴런은 'relu' 활성화 함수에 따라 값을 전달할지 말지를 결정하게 된다.
model_1.add(layers.Dense(16, activation='relu', input_shape=(1,)))
# 마지막 레이어는 하나의 뉴런인데, 원하는 결과값이 하나의 값이기 때문이다.
model_1.add(layers.Dense(1))
# 표준 옵티마이저와 손실을 사용하여 회귀 모델을 컴파일한다.
model_1.compile(optimizer='rmsprop', loss='mse', metrics=['mae'])
# + [markdown] colab_type="text" id="O0idLyRLQeGj"
# ## 모델 훈련하기
# 모델을 정의한 후에는 데이터를 사용하여 훈련_train_할 수 있다. 훈련은 신경망에 x값을 전달하고 네트워크의 출력이 예상되는 y값에서 얼마나 멀리 떨어져 있는지 확인하고 다음에 출력이 더 정확할 수 있도록 뉴런의 가중치와 바이어스를 조정한다.
#
# 훈련은 이 프로세스를 전체 데이터 세트에서 여러 번에 걸쳐 실행되는데, 해당 횟수를 에포크_epoch_라고 한다. 훈련중에 실행할 에포크 수는 우리가 설정할 수 있는 파라미터다.
#
# 각 에포크 동안 데이터는 여러 배치_batches_네트워크를 통해 실행된다. 네트워크의 가중치와 바이어스는 매 배치마다 한 번씩 조정되며, 배치크기_batch size_도 우리가 설정할 수 있는 파라미터이다.
#
# 다음 셀의 코드는 학습 데이터의 `x` 및`y` 값을 사용하여 모델을 학습시킨다. 1000 에포크_epochs_에 배치 사이즈는_batch_ 16이며 검증_validation_에 사용할 데이터도 전달된다. 셀을 실행하면 훈련을 완료하는 데 시간이 걸릴 수 있다.
#
#
# + colab={"base_uri": "https://localhost:8080/"} colab_type="code" id="p8hQKr4cVOdE" outputId="3f1a7904-ffcd-4bb7-8bbb-bcd85a132128"
# 모델 훈련
history_1 = model_1.fit(x_train, y_train, epochs=1000, batch_size=16,
validation_data=(x_validate, y_validate))
# + [markdown] colab_type="text" id="cRE8KpEqVfaS"
# ## 훈련 지표 확인
# 훈련 중에 모델의 성능은 훈련 데이터와 이전에 따로 설정한 유효성 검사 데이터에 대해 지속적으로 측정된다. 다음은 훈련 과정에서 모델의 성능이 어떻게 변했는지 알려주는 데이터 로그중 일부를 그래프로 표시한 것이다.
# + colab={"base_uri": "https://localhost:8080/", "height": 295} colab_type="code" id="CmvA-ksoln8r" outputId="1b834831-81e8-4548-dd8c-f5edf2c3ff43"
# 훈련과 검증 과정에서 예측값과 실제값 사이의 거리인 손실 그래프를 그린다.
loss = history_1.history['loss']
val_loss = history_1.history['val_loss']
epochs = range(1, len(loss) + 1)
plt.plot(epochs, loss, 'g.', label='Training loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
# + [markdown] colab_type="text" id="iOFBSbPcYCN4"
# ## 데이터 살펴보기
# 그래프는 각 에포크에 대한 손실값_loss_(또는 모델의 예측과 실제 데이터의 차이)를 보여준다. 손실을 계산하는 방법에는 여러 가지가 있으며, 우리가 사용한 방법은 평균제곱오차_mean squared error_이다. 훈련 및 검증 데이터에 대해 손실값을 확인할 수 있다.
#
# 보시다시, 손실량은 평평 해지기 전에 처음 25개 에포크에서 급속히 감소한다. 이는 모델이 보다 개선되어 정확한 예측을 수행하고 있음을 의미한다!
#
# 우리의 목표는 모델이 더 이상 개선되지 않거나 훈련 손실값_training loss_이 검증 손실값_validation loss_보다 작을 때 훈련을 중단하는 것이다. 이는 모델이 더 이상 새로운 데이터로 일반화 할 수 없도록 훈련 데이터를 잘 예측하는 법을 배웠음을 의미한다.
#
# 그래프의 평평한 부분을 보다 읽기 쉽게하기 위해 처음 50에포크를 제외해보자.
# + colab={"base_uri": "https://localhost:8080/", "height": 295} colab_type="code" id="Zo0RYroFZYIV" outputId="e6841332-0541-44bb-a186-ae5b46781e51"
# 그래프를 보기 쉽도록 처음 몇 에폭을 제외.
SKIP = 50
plt.plot(epochs[SKIP:], loss[SKIP:], 'g.', label='Training loss')
plt.plot(epochs[SKIP:], val_loss[SKIP:], 'b.', label='Validation loss')
plt.title('Training and validation loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
# + [markdown] colab_type="text" id="W4EQD-Bb8hLM"
# ## 추가 지표 활용
# 그래프에서 우리는 약 600 에폭까지 손실이 계속 감소하고 이 이후부터 안정적인 움직임을 보이는 것을 알 수 있다. 이는 600에폭이 넘어가면 네트워크를 훈련시킬 필요가 없음을 의미한다.
#
# 그러나 가장 낮은 손실 값이 여전히 약 0.155임을 알 수 있다. 이는 우리 네트워크 예측이 평균 ~15% 떨어져 있다는 것을 의미한다.
#
# 모델 성능에 대한 통찰력을 얻기 위해 더 많은 그래프를 그려볼 수 있는데, 이번에는 평균절대오차를 사용할 것이다. 이는 네트워크 예측이 실제 수치와 얼마나 멀리 떨어져 있는지 측정하는 또 다른 방법이다.
# + colab={"base_uri": "https://localhost:8080/", "height": 295} colab_type="code" id="Md9E_azmpkZU" outputId="39b97561-b01d-49f2-c35c-fbd8db663806"
plt.clf()
# 예측에서 오차의 양을 측정하는 또 다른 방법인 평균 절대 오차 그래프를 그린다.
mae = history_1.history['mae']
val_mae = history_1.history['val_mae']
plt.plot(epochs[SKIP:], mae[SKIP:], 'g.', label='Training MAE')
plt.plot(epochs[SKIP:], val_mae[SKIP:], 'b.', label='Validation MAE')
plt.title('Training and validation mean absolute error')
plt.xlabel('Epochs')
plt.ylabel('MAE')
plt.legend()
plt.show()
# + [markdown] colab_type="text" id="ctawd0CXAVEw"
# 평균절대오차_mean absolute error_ 그래프는 모델의 다른 측면을 알려준다. 훈련데이터에 비해 검증 데이터에서 보다 일관되게 오류가 높다는 것인데, 이는 네트워크에 과적합_overfit_이 있거나 훈련 데이터를 너무 학습하여 새로운 데이터에 대해 효과적으로 예측을 할 수 없음을 의미한다.
#
# 또한 평균 절대 오차 값은 ~0.305로 매우 높으며, 이는 일부 모델의 예측이 30% 이상 떨어져 있음을 의미한다. 30%오류는 사인파 함수를 정확하게 모델링하는 것과는 거리가 멀다는 것을 의미한다.
#
# 무슨일이 일어나고 있는지에 알아보기 위해 예측값을 그래프로 나타내보자.
# + colab={"base_uri": "https://localhost:8080/", "height": 281} colab_type="code" id="i13eVIT3B9Mj" outputId="afc103e2-0beb-4a26-fe18-c0cccc6d3d2a"
# 모델을 사용하여 검증 데이터로부터 예측값 생성
predictions = model_1.predict(x_train)
# 테스트 데이터와 함께 예측값을 그래프로 표현
plt.clf()
plt.title('Training data predicted vs actual values')
plt.plot(x_test, y_test, 'b.', label='Actual')
plt.plot(x_train, predictions, 'r.', label='Predicted')
plt.legend()
plt.show()
# + [markdown] colab_type="text" id="Wokallj1D21L"
# 이런! 그래프는 우리 모델이 사인 함수를 매우 제한된 방식으로만 예측한다는 걸 보여준다. 0<=x<=1.1에서 선은 대부분 적합하지만 나머지의 경우 부적절한 것으로 보인다.
#
# 왜 그럴까? 이 경우, 모델에 사인파 함수의 전체 복잡성을 학습할 수있는 충분한 Capacity가 없기 때문에 지나치게 단순한 방식으로만 예측을 수행한다. 모델을 더 크게 만들면 Capacity가 늘어나고 성능은 향상될 것이다.
#
# ## 모델 변경
#
# 모델을 더 크게 만들기 위해 레이어를 추가할 것이다. 다음 셀은 이전과 같은 방식으로 모델을 재정의하지만 중간에 16개의 뉴런이 추가로 있다.
# + colab={} colab_type="code" id="oW0xus6AF-4o"
model_2 = tf.keras.Sequential()
# 첫 번째 레이어는 스칼라 입력을 받아 16개의 뉴런을 통해 전달하고,
# 뉴런은 ‘relu’ 활성화 함수에 따라 활성화 여부를 결정한다.
model_2.add(layers.Dense(16, activation='relu', input_shape=(1,)))
# 새로운 두 번째 레이어는 네트워크가 더 복잡한 표현을 배우는 데 도움을 준다.
model_2.add(layers.Dense(16, activation='relu'))
# 단일 값을 출력해야 하기 때문에 최종 레이어는 단일 뉴런으로 구성된다.
model_2.add(layers.Dense(1))
# 표준 옵티마이저 및 손실 함수를 사용하여 회귀 모델을 컴파일
model_2.compile(optimizer='rmsprop', loss='mse', metrics=['mae'])
# + [markdown] colab_type="text" id="Dv2SC409Grap"
# 우리는 이제 새로운 모델을 훈련시킬 것이다. 시간을 절약하기 위해 600 에포크만 훈련하자.
# + colab={"base_uri": "https://localhost:8080/"} colab_type="code" id="DPAUrdkmGq1M" outputId="34ad91e0-229b-479c-bd65-12ad1ed1c660"
history_2 = model_2.fit(x_train, y_train, epochs=600, batch_size=16,
validation_data=(x_validate, y_validate))
# + [markdown] colab_type="text" id="Mc_CQu2_IvOP"
# ## 새 모델 평가하기
# 각 에포크 마다 모델은 훈련 및 검증셋에 대한 손실 및 평균 절대 오차를 출력한다. (정확한 숫자는 다를 수 있다).
#
# ```
# Epoch 600/600
# 600/600 [==============================] - 0s 109us/sample - loss: 0.0124 - mae: 0.0892 - val_loss: 0.0116 - val_mae: 0.0845
# ```
#
# 여기에서 모델이 크게 개선된 것을 확인할 수 있다. 검증 손실이 0.15에서 0.015로, 검증 MAE가 0.31에서 0.1로 떨어졌다.
#
# 다음 셀은 원래 모델을 평가하는 데 사용한 것과 동일한 그래프를 출력하지만 새로 훈련한 모델을 사용한 데이터를 보여줍니다.
# + colab={"base_uri": "https://localhost:8080/", "height": 851} colab_type="code" id="SYHGswAJJgrC" outputId="efcc51f6-f1f1-490a-ffba-ed283586f83e"
# 훈련과 검증 과정에서 예측값과 실제값 사이의 거리인 손실 그래프를 그린다.
loss = history_2.history['loss']
val_loss = history_2.history['val_loss']
epochs = range(1, len(loss) + 1)
plt.plot(epochs, loss, 'g.', label='Training loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
# 그래프를 보기 쉽도록 처음 몇 에폭을 제외.
SKIP = 100
plt.clf()
plt.plot(epochs[SKIP:], loss[SKIP:], 'g.', label='Training loss')
plt.plot(epochs[SKIP:], val_loss[SKIP:], 'b.', label='Validation loss')
plt.title('Training and validation loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
plt.clf()
# 예측에서 오차의 양을 측정하는 또 다른 방법인 평균 절대 오차 그래프를 그린다.
mae = history_2.history['mae']
val_mae = history_2.history['val_mae']
plt.plot(epochs[SKIP:], mae[SKIP:], 'g.', label='Training MAE')
plt.plot(epochs[SKIP:], val_mae[SKIP:], 'b.', label='Validation MAE')
plt.title('Training and validation mean absolute error')
plt.xlabel('Epochs')
plt.ylabel('MAE')
plt.legend()
plt.show()
# + [markdown] colab_type="text" id="f86dWOyZKmN9"
# 이 그래프에서 몇 가지 흥미로운 점을 살펴볼 수 있다.
#
# * 우리의 네트워크는 훨씬 더 빠르게 최고 정확도에 도달했다 (600 대신 200 에포크 이내).
# * 전체 손실과 MAE는 이전 네트워크보다 훨씬 개선되었다.
# * 전체적인 지표가 훈련보다 검증셋에 더 좋습니다. 즉, 네트워크가 과적합되지 않았다.
#
# 검증 지표가 훈련 지표보다 나은 이유는 검증 기간이 각 시대의 끝에서 계산되는 반면, 훈련 지표는 시대 전체에서 계산되므로 검증은 약간 더 오래 훈련된 모델에서 발생하기 때문이다.
#
# 모델을 좀 더 살펴보기 위해 앞에서 따로 설정한 테스트 데이터 세트와 비교하여 예측을 확인해보자.
# + colab={"base_uri": "https://localhost:8080/", "height": 298} colab_type="code" id="lZfztKKyhLxX" outputId="b792a12e-713d-4b07-9f8e-de0d059d5cdb"
# 테스트 데이터셋의 손실 계산 및 출력
loss = model_2.evaluate(x_test, y_test)
# 테스트 데이터셋 기반으로 예측
predictions = model_2.predict(x_test)
# 실제값에 대한 예측 그래프
plt.clf()
plt.title('Comparison of predictions and actual values')
plt.plot(x_test, y_test, 'b.', label='Actual')
plt.plot(x_test, predictions, 'r.', label='Predicted')
plt.legend()
plt.show()
# + [markdown] colab_type="text" id="3h7IcvuOOS4J"
# 훨씬 나아졌다. 우리가 출력한 평가 지표는 모델이 테스트 데이터에서 손실과 MAE가 낮고 예측이 데이터와 시각적으로 잘 일치함을 보여준다.
#
# 모델이 완벽하지는 않다. 매끄러운 사인 곡선을 형성하지 않기 때문인데, 예를 들어,`x`가 4.2와 5.2 사이일 때 선은 거의 직선이다. 더 개선하고 싶다면 모델의 용량을 더 늘리고, 추가적인 기법을 사용하여 과적합을 막을 수도 있을 것이다.
#
# 그러나 머신러닝의 중요한 부분은 언제 종료해야 하는지를 아는 것이다. 이 정도 모델이면 LED를 깜빡이는 예제로서는 충분하다.
#
# ## TensorFlow Lite로 변환
# 우리는 이제 원하는 모델을 얻었다. 그러나 모델을 마이크로 컨트롤러 용 TensorFlow Lite와 함께 사용하려면 올바른 형식으로 변환하여 파일로 다운로드 해야한다. 이를 위해 [TensorFlow Lite Converter](https://www.tensorflow.org/lite/convert)를 사용해보자. 변환기는 메모리 제한된 장치에서도 사용하기 위해 특수한 형식으로 출력한다.
#
# 이 모델은 마이크로 컨트롤러에 배포될 예정이므로 가능한 작은 크기로 만들어야 한다. 모델의 크기를 줄이는 한 가지 기술로 [양자화(quantization)] (https://www.tensorflow.org/lite/performance/post_training_quantization)가 있다. 이는 모델 가중치의 정밀도를 낮추어 정확도에 큰 영향을 미치지 않으면서 메모리를 절약할 수 있다. 양자화를 통해 모델 추론에 필요한 계산이 더 간단해지기 때문에 경량화 뿐만아니라 실행속도도 빨라진다.
#
# TensorFlow Lite 변환기는 모델을 변환하는 동안 양자화를 적용 할 수 있다. 다음 셀에서는 모델을 두 번 변환한다. 한 번은 양자화하고 진행하고 한 번은 하지 않고 변환한다:
# + colab={} colab_type="code" id="1muAoUm8lSXL"
# 양자화 없이 모델을 텐서플로우 라이트 형식으로 변환
converter = tf.lite.TFLiteConverter.from_keras_model(model_2)
tflite_model = converter.convert()
# 모델을 디스크에 저장
open("sine_model.tflite", "wb").write(tflite_model)
# 양자화하여 모델을 텐서플로우 라이트 형식으로 변환
converter = tf.lite.TFLiteConverter.from_keras_model(model_2)
converter.optimizations = [tf.lite.Optimize.OPTIMIZE_FOR_SIZE]
tflite_model = converter.convert()
# 모델을 디스크에 저장
open("sine_model_quantized.tflite", "wb").write(tflite_model)
# + [markdown] colab_type="text" id="L_vE-ZDkHVxe"
# ## 변환된 모델 테스트
# 모델이 양자화 변환 후에도 여전히 정확한지 입증하기 위해 두 모델을 모두 사용하여 예측하고 테스트 결과와 비교해보자.
# + colab={"base_uri": "https://localhost:8080/", "height": 281} colab_type="code" id="-J7IKlXiYVPz" outputId="0c10f56c-dbd7-4cc3-e332-30ad673769e5"
# 각 모델에 대한 인터프리터 인스턴스화.
sine_model = tf.lite.Interpreter('sine_model.tflite')
sine_model_quantized = tf.lite.Interpreter('sine_model_quantized.tflite')
# 각 모델에 대한 메모리 할당
sine_model.allocate_tensors()
sine_model_quantized.allocate_tensors()
# 입력과 결과 텐서에 대한 인덱스 가져오기
sine_model_input = sine_model.tensor(sine_model.get_input_details()[0]["index"])
sine_model_output = sine_model.tensor(sine_model.get_output_details()[0]["index"])
sine_model_quantized_input = sine_model_quantized.tensor(sine_model_quantized.get_input_details()[0]["index"])
sine_model_quantized_output = sine_model_quantized.tensor(sine_model_quantized.get_output_details()[0]["index"])
# 결과를 저장하기 위한 배열 생성
sine_model_predictions = np.empty(x_test.size)
sine_model_quantized_predictions = np.empty(x_test.size)
# 각 값에 대해 각 모델의 인터프리터를 실행하고 결과를 배열에 저장
for i in range(x_test.size):
sine_model_input().fill(x_test[i])
sine_model.invoke()
sine_model_predictions[i] = sine_model_output()[0]
sine_model_quantized_input().fill(x_test[i])
sine_model_quantized.invoke()
sine_model_quantized_predictions[i] = sine_model_quantized_output()[0]
# 데이터가 어떻게 정렬되는지 확인
plt.clf()
plt.title('Comparison of various models against actual values')
plt.plot(x_test, y_test, 'bo', label='Actual')
plt.plot(x_test, predictions, 'ro', label='Original predictions')
plt.plot(x_test, sine_model_predictions, 'bx', label='Lite predictions')
plt.plot(x_test, sine_model_quantized_predictions, 'gx', label='Lite quantized predictions')
plt.legend()
plt.show()
# + [markdown] colab_type="text" id="jWxvLGexKv0D"
# 그래프에서 원래 모델, 변환된 모델, 양자화 된 모델에 대한 예측이 모두 구별할 수 없을 정도로 차이가 없는 것을 확인할 수 있다. 이를통해 양자화된 모델을 사용하는 것이 큰 무리가 없음을 알 수 있다.
#
# 다음으로, 모델크기는 얼마나 차이나는지 알아보자.
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" id="6r42iBnULP4X" outputId="afe526c9-498d-498e-d768-1edfbf21e870"
import os
basic_model_size = os.path.getsize("sine_model.tflite")
print("Basic model is %d bytes" % basic_model_size)
quantized_model_size = os.path.getsize("sine_model_quantized.tflite")
print("Quantized model is %d bytes" % quantized_model_size)
difference = basic_model_size - quantized_model_size
print("Difference is %d bytes" % difference)
# + [markdown] colab_type="text" id="C2vpZE9ZshVH"
# 양자화된 모델은 원래 버전보다 16 바이트 작다. 약 2.6킬로 바이트 크기의 모델은 이미 너무 작아서 가중치가 전체 크기의 작은 부분만 차지하므로 양자화가 거의 영향을 미치지 않은 것이다.
#
# 더 복잡한 모델의 경우, 더 많은 가중치를 가지므로 양자화로 인한 공간 절약이 훨씬 높아져 최대 4배까지 모델 크기를 줄일 수 있다.
#
# 어쨌든, 양자화 된 모델은 원래 버전보다 실행시간이 덜 걸리며, 이는 마이크로 컨트롤러에서 중요한 부분이다.
#
# ## C 파일에 쓰기
# 마이크로 컨트롤러 용 TensorFlow Lite와 함께 사용할 모델을 준비하는 마지막 단계는 모델을 C 소스 파일로 변환하는 것이다. 이 형식의 예제는 [`hello_world / sine_model_data.cc`](https://github.com/tensorflow/tensorflow/blob/master/tensorflow/lite/micro/examples/hello_world/sine_model_data.cc)에서 확인할 수 있다.
#
# 이를 위해 [`xxd`](https://linux.die.net/man/1/xxd)라는 명령행 유틸리티를 사용할 수 있다. 다음 셀은 양자화된 모델에서 xxd를 실행하고 출력한다.
# + colab={"base_uri": "https://localhost:8080/", "height": 3808} colab_type="code" id="l4-WhtGpvb-E" outputId="f975721f-bdd1-440a-93af-55f13c4c8690"
# xxd를 사용할 수 없을 경우, 설치한다.
# !apt-get -qq install xxd
# 파일을 C 소스파일로 저장
# !xxd -i sine_model_quantized.tflite > sine_model_quantized.cc
# 소스파일을 출력
# !cat sine_model_quantized.cc
# + [markdown] colab_type="text" id="1sqrhBLXwILt"
# 이 출력값을 복사하여 프로젝트의 소스 코드에 붙여 넣거나이 Colab 왼쪽의 메뉴를 사용하여 파일을 다운로드 할 수 있다.
#
# -
|
tensorflow/lite/micro/examples/hello_world/create_sine_model_ko.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import ipyvolume as ipv
from forest3d.geometry import Tree
from scipy.spatial import cKDTree
from scipy.interpolate import RegularGridInterpolator
tree_x, tree_y, tree_z = Tree(species='Douglas-fir',
dbh=15, top_height=150,
stem_x=0, stem_y=0, stem_z=0,
crown_shapes=((2.0, 1.5, 1.7, 2.3), (1.3, 1.2, 1.0, 1.1)),
crown_radii=(5,6,8,7)
).get_crown()
tree_xs, tree_ys, tree_zs = tree_x.reshape((50,32)), tree_y.reshape((50,32)), tree_z.reshape((50,32))
grid_x = np.linspace(tree_x.min()-200, tree_x.max()+200,400)
grid_y = np.linspace(tree_y.min()-150, tree_y.max()+150,300)
grid_z = np.linspace(-10, tree_z.max()+10,100)
grid_xx, grid_yy, grid_zz = np.meshgrid(grid_x,grid_y,grid_z)
grid_xs = grid_xx.flatten()
grid_ys = grid_yy.flatten()
grid_zs = grid_zz.flatten()
ipv.figure(width=800)
ipv.scatter(grid_xs, grid_ys, grid_zs, marker="sphere", size=0.75, color='black')
# ipv.plot_surface(tree_xs, tree_ys, tree_zs, color="green")
ipv.plot_wireframe(tree_xs, tree_ys, tree_zs, color="green")
ipv.style.use('minimal')
ipv.show()
kdtree = cKDTree(data = np.stack((tree_x, tree_y, tree_z)).T)
grid_points = np.stack((grid_xs, grid_ys, grid_zs)).T
grid_distances = kdtree.query(grid_points)
interpolator = RegularGridInterpolator(points = (grid_x,grid_y,grid_z),
values = grid_distances[0].reshape(grid_x.shape[0], grid_y.shape[0], grid_z.shape[0]))
rand_x = (grid_x.max() - grid_x.min())*np.random.rand(50*32*5000) + grid_x.min()
rand_y = (grid_y.max() - grid_y.min())*np.random.rand(50*32*5000) + grid_y.min()
rand_z = (grid_z.max() - grid_z.min())*np.random.rand(50*32*5000) + grid_z.min()
ipv.scatter(rand_x, rand_y, rand_z, marker='sphere', color='blue', size=4)
# %%time
distances = interpolator((rand_x, rand_y, rand_z))
distance_metric = distances.sum()
print(distance_metric)
distances
|
notebooks/sandbox/Illustrating grid interpolation.ipynb
|