code
stringlengths 38
801k
| repo_path
stringlengths 6
263
|
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # CNTK 202: Language Understanding with Recurrent Networks
#
# This tutorial shows how to implement a recurrent network to process text,
# for the [Air Travel Information Services](https://catalog.ldc.upenn.edu/LDC95S26)
# (ATIS) task of slot tagging (tag individual words to their respective classes,
# where the classes are provided as labels in the training data set).
#
# There are 2 parts to this tutorial:
# - Part 1: We will tag each word in a sequence to their corresponding label
# - Part 2: We will classify a sequence to its corresponding intent.
#
# We will start with a straight-forward (linear) embedding of the words followed by a recurrent LSTM to label each word in a sequence to the corresponding class. We will show how to classify each word token in a sequence to the corresponding class. This will then be extended to include neighboring words and run bidirectionally.
#
# We will take the last state of the sequence and train a model that classifies the entire sequence to the corresponding class label (in this case the intent associated with the sequence).
#
# The techniques you will practice are:
# * model description by composing layer blocks, a convenient way to compose
# networks/models without requiring the need to write formulas,
# * creating your own layer block
# * variables with different sequence lengths in the same network
# * training the network
#
# We assume that you are familiar with basics of deep learning, and these specific concepts:
# * recurrent networks ([Wikipedia page](https://en.wikipedia.org/wiki/Recurrent_neural_network))
# * text embedding ([Wikipedia page](https://en.wikipedia.org/wiki/Word_embedding))
#
# ## Prerequisites
#
# We assume that you have already [installed CNTK](https://docs.microsoft.com/en-us/cognitive-toolkit/Setup-CNTK-on-your-machine).
# This tutorial requires CNTK V2. We strongly recommend to run this tutorial on a machine with
# a capable CUDA-compatible GPU. Deep learning without GPUs is not fun.
# ## Data download
#
# In this tutorial, we are going to use a (lightly preprocessed) version of the ATIS dataset. You can download the data automatically by running the cells below or by executing the manual instructions.
#
# **Fallback manual instructions**
# Download the ATIS [training](https://github.com/Microsoft/CNTK/blob/release/2.5/Tutorials/SLUHandsOn/atis.train.ctf)
# and [test](https://github.com/Microsoft/CNTK/blob/release/2.5/Tutorials/SLUHandsOn/atis.test.ctf)
# files and put them at the same folder as this notebook. If you want to see how the model is
# predicting on new sentences you will also need the vocabulary files for
# [queries](https://github.com/Microsoft/CNTK/blob/release/2.5/Examples/LanguageUnderstanding/ATIS/BrainScript/query.wl) and
# [slots](https://github.com/Microsoft/CNTK/blob/release/2.5/Examples/LanguageUnderstanding/ATIS/BrainScript/slots.wl)
# +
from __future__ import print_function # Use a function definition from future version (say 3.x from 2.7 interpreter)
import requests
import os
def download(url, filename):
""" utility function to download a file """
response = requests.get(url, stream=True)
with open(filename, "wb") as handle:
for data in response.iter_content():
handle.write(data)
locations = ['Tutorials/SLUHandsOn', 'Examples/LanguageUnderstanding/ATIS/BrainScript']
data = {
'train': { 'file': 'atis.train.ctf', 'location': 0 },
'test': { 'file': 'atis.test.ctf', 'location': 0 },
'query': { 'file': 'query.wl', 'location': 1 },
'slots': { 'file': 'slots.wl', 'location': 1 },
'intent': { 'file': 'intent.wl', 'location': 1 }
}
for item in data.values():
location = locations[item['location']]
path = os.path.join('..', location, item['file'])
if os.path.exists(path):
print("Reusing locally cached:", item['file'])
# Update path
item['file'] = path
elif os.path.exists(item['file']):
print("Reusing locally cached:", item['file'])
else:
print("Starting download:", item['file'])
url = "https://github.com/Microsoft/CNTK/blob/release/2.5/%s/%s?raw=true"%(location, item['file'])
download(url, item['file'])
print("Download completed")
# -
# **Importing libraries**: CNTK, math and numpy
#
# CNTK's Python module contains several submodules like `io`, `learner`, and `layers`. We also use NumPy in some cases since the results returned by CNTK work like NumPy arrays.
# +
import math
import numpy as np
import cntk as C
import cntk.tests.test_utils
cntk.tests.test_utils.set_device_from_pytest_env() # (only needed for our build system)
C.cntk_py.set_fixed_random_seed(1) # fix a random seed for CNTK components
# -
# ## Task overview: Slot tagging
#
# The task we want to approach in this tutorial is slot tagging.
# We use the [ATIS corpus](https://catalog.ldc.upenn.edu/LDC95S26).
# ATIS contains human-computer queries from the domain of Air Travel Information Services,
# and our task will be to annotate (tag) each word of a query whether it belongs to a
# specific item of information (slot), and which one.
#
# The data in your working folder has already been converted into the "CNTK Text Format."
# Let us look at an example from the test-set file `atis.test.ctf`:
#
# 19 |S0 178:1 |# BOS |S1 14:1 |# flight |S2 128:1 |# O
# 19 |S0 770:1 |# show |S2 128:1 |# O
# 19 |S0 429:1 |# flights |S2 128:1 |# O
# 19 |S0 444:1 |# from |S2 128:1 |# O
# 19 |S0 272:1 |# burbank |S2 48:1 |# B-fromloc.city_name
# 19 |S0 851:1 |# to |S2 128:1 |# O
# 19 |S0 789:1 |# st. |S2 78:1 |# B-toloc.city_name
# 19 |S0 564:1 |# louis |S2 125:1 |# I-toloc.city_name
# 19 |S0 654:1 |# on |S2 128:1 |# O
# 19 |S0 601:1 |# monday |S2 26:1 |# B-depart_date.day_name
# 19 |S0 179:1 |# EOS |S2 128:1 |# O
#
# This file has 7 columns:
#
# * a sequence id (19). There are 11 entries with this sequence id. This means that sequence 19 consists
# of 11 tokens;
# * column `S0`, which contains numeric word indices; the input data is encoded in one-hot vectors. There are 943 words in the vocabulary, so each word is a 943 element vector of all 0 with a 1 at a vector index chosen to represent that word. For example the word "from" is represented with a 1 at index 444 and zero everywhere else in the vector. The word "monday" is represented with a 1 at index 601 and zero everywhere else in the vector.
# * a comment column denoted by `#`, to allow a human reader to know what the numeric word index stands for;
# Comment columns are ignored by the system. `BOS` and `EOS` are special words
# to denote beginning and end of sentence, respectively;
# * column `S1` is an intent label, which we will use in the second part of the tutorial;
# * another comment column that shows the human-readable label of the numeric intent index;
# * column `S2` is the slot label, represented as a numeric index; and
# * another comment column that shows the human-readable label of the numeric label index.
#
# The task of the neural network is to look at the query (column `S0`) and predict the
# slot label (column `S2`).
# As you can see, each word in the input gets assigned either an empty label `O`
# or a slot label that begins with `B-` for the first word, and with `I-` for any
# additional consecutive word that belongs to the same slot.
#
# ### Model Creation
#
# The model we will use is a recurrent model consisting of an embedding layer,
# a recurrent LSTM cell, and a dense layer to compute the posterior probabilities:
#
#
# slot label "O" "O" "O" "O" "B-fromloc.city_name"
# ^ ^ ^ ^ ^
# | | | | |
# +-------+ +-------+ +-------+ +-------+ +-------+
# | Dense | | Dense | | Dense | | Dense | | Dense | ...
# +-------+ +-------+ +-------+ +-------+ +-------+
# ^ ^ ^ ^ ^
# | | | | |
# +------+ +------+ +------+ +------+ +------+
# 0 -->| LSTM |-->| LSTM |-->| LSTM |-->| LSTM |-->| LSTM |-->...
# +------+ +------+ +------+ +------+ +------+
# ^ ^ ^ ^ ^
# | | | | |
# +-------+ +-------+ +-------+ +-------+ +-------+
# | Embed | | Embed | | Embed | | Embed | | Embed | ...
# +-------+ +-------+ +-------+ +-------+ +-------+
# ^ ^ ^ ^ ^
# | | | | |
# w ------>+--------->+--------->+--------->+--------->+------...
# BOS "show" "flights" "from" "burbank"
#
# Or, as a CNTK network description. Please have a quick look and match it with the description above:
# (descriptions of these functions can be found at: [the layers reference](http://cntk.ai/pythondocs/layerref.html))
#
# +
# number of words in vocab, slot labels, and intent labels
vocab_size = 943 ; num_labels = 129 ; num_intents = 26
# model dimensions
input_dim = vocab_size
label_dim = num_labels
emb_dim = 150
hidden_dim = 300
# Create the containers for input feature (x) and the label (y)
x = C.sequence.input_variable(vocab_size)
y = C.sequence.input_variable(num_labels)
def create_model():
with C.layers.default_options(initial_state=0.1):
return C.layers.Sequential([
C.layers.Embedding(emb_dim, name='embed'),
C.layers.Recurrence(C.layers.LSTM(hidden_dim), go_backwards=False),
C.layers.Dense(num_labels, name='classify')
])
# -
# Now we are ready to create a model and inspect it.
#
# The model attributes are fully accessible from Python. The first layer named `embed` is an Embedding layer. Here we use the CNTK default, which is linear embedding. It is a simple matrix with dimension (input word encoding x output projected dimension). You can access its parameter `E` (where the embeddings are stored) like any other attribute of a Python object. Its shape contains a `-1` which indicates that this parameter (with input dimension) is not fully specified yet, while the output dimension is set to `emb_dim` ( = 150 in this tutorial).
#
# Additionally, we also inspect the value of the bias vector in the `Dense` layer named `classify`. The `Dense` layer is a fundamental compositional unit of a Multi-Layer Perceptron (as introduced in CNTK 103C tutorial). The `Dense` layer has both `weight` and `bias` parameters, one each per `Dense` layer. Bias terms are by default initialized to 0 (but there is a way to change that if you need). As you create the model, one should name the layer component and then access the parameters as shown here.
#
# **Suggested task**: What should be the expected dimension of the `weight` matrix from the layer named `classify`? Try printing the weight matrix of the `classify` layer? Does it match with your expected size?
# peek
z = create_model()
print(z.embed.E.shape)
print(z.classify.b.value)
# In our case we have input as one-hot encoded vector of length 943 and the output dimension `emb_dim` is set to 150. In the code below we pass the input variable `x` to our model `z`. This binds the model with input data of known shape. In this case, the input shape will be the size of the input vocabulary. With this modification, the parameter returned by the embed layer is completely specified (943, 150). **Note**: You can initialize the Embedding matrix with pre-computed vectors using [Word2Vec](https://en.wikipedia.org/wiki/Word2vec) or [GloVe](https://en.wikipedia.org/wiki/GloVe_%28machine_learning%29).
# Pass an input and check the dimension
z = create_model()
print(z(x).embed.E.shape)
# To train and test a model in CNTK, we need to create a model and specify how to read data and perform training and testing.
#
# In order to train we need to specify:
#
# * how to read the data
# * the model function, its inputs, and outputs
# * hyper-parameters for the learner such as the learning rate
#
# [comment]: <> (For testing ...)
#
# ## Data Reading
#
# We already looked at the data.
# But how do you generate this format?
# For reading text, this tutorial uses the `CNTKTextFormatReader`. It expects the input data to be
# in a specific format, as described [here](https://docs.microsoft.com/en-us/cognitive-toolkit/Brainscript-CNTKTextFormat-Reader).
#
# For this tutorial, we created the corpora by two steps:
# * convert the raw data into a plain text file that contains of TAB-separated columns of space-separated text. For example:
#
# ```
# BOS show flights from burbank to st. louis on monday EOS (TAB) flight (TAB) O O O O B-fromloc.city_name O B-toloc.city_name I-toloc.city_name O B-depart_date.day_name O
# ```
#
# This is meant to be compatible with the output of the `paste` command.
# * convert it to CNTK Text Format (CTF) with the following command:
#
# ```
# python [CNTK root]/Scripts/txt2ctf.py --map query.wl intent.wl slots.wl --annotated True --input atis.test.txt --output atis.test.ctf
# ```
# where the three `.wl` files give the vocabulary as plain text files, one word per line.
#
# In these CTF files, our columns are labeled `S0`, `S1`, and `S2`.
# These are connected to the actual network inputs by the corresponding lines in the reader definition:
def create_reader(path, is_training):
return C.io.MinibatchSource(C.io.CTFDeserializer(path, C.io.StreamDefs(
query = C.io.StreamDef(field='S0', shape=vocab_size, is_sparse=True),
intent = C.io.StreamDef(field='S1', shape=num_intents, is_sparse=True),
slot_labels = C.io.StreamDef(field='S2', shape=num_labels, is_sparse=True)
)), randomize=is_training, max_sweeps = C.io.INFINITELY_REPEAT if is_training else 1)
# peek
reader = create_reader(data['train']['file'], is_training=True)
reader.streams.keys()
# ## Training
#
# We also must define the training criterion (loss function), and also an error metric to track. In most tutorials, we know the input dimensions and the corresponding labels. We directly create the loss and the error functions. In this tutorial we will do the same. However, we take a brief detour and learn about placeholders. This concept would be useful for Task 3.
#
# **Learning note**: Introduction to `placeholder`: Remember that the code we have been writing is not actually executing any heavy computation it is just specifying the function we want to compute on data during training/testing. And in the same way that it is convenient to have names for arguments when you write a regular function in a programming language, it is convenient to have placeholders that refer to arguments (or local computations that need to be reused). Eventually, some other code will replace these placeholders with other known quantities in the same way that in a programming language the function will be called with concrete values bound to its arguments.
#
# Specifically, the input variables you have created above `x = C.sequence.input_variable(vocab_size)` holds data pre-defined by `vocab_size`. In the case where such instantiations are challenging or not possible, using `placeholder` is a logical choice. Having the `placeholder` only allows you to defer the specification of the argument at a later time when you may have the data.
#
# Here is an example below that illustrates the use of `placeholder`.
# +
def create_criterion_function(model):
labels = C.placeholder(name='labels')
ce = C.cross_entropy_with_softmax(model, labels)
errs = C.classification_error (model, labels)
return C.combine ([ce, errs]) # (features, labels) -> (loss, metric)
criterion = create_criterion_function(create_model())
criterion.replace_placeholders({criterion.placeholders[0]: C.sequence.input_variable(num_labels)})
# -
# While the cell above works well when one has input parameters defined at network creation, it compromises readability. Hence we prefer creating functions as shown below
def create_criterion_function_preferred(model, labels):
ce = C.cross_entropy_with_softmax(model, labels)
errs = C.classification_error (model, labels)
return ce, errs # (model, labels) -> (loss, error metric)
def train(reader, model_func, max_epochs=10, task='slot_tagging'):
# Instantiate the model function; x is the input (feature) variable
model = model_func(x)
# Instantiate the loss and error function
loss, label_error = create_criterion_function_preferred(model, y)
# training config
epoch_size = 18000 # 18000 samples is half the dataset size
minibatch_size = 70
# LR schedule over epochs
# In CNTK, an epoch is how often we get out of the minibatch loop to
# do other stuff (e.g. checkpointing, adjust learning rate, etc.)
lr_per_sample = [3e-4]*4+[1.5e-4]
lr_per_minibatch = [lr * minibatch_size for lr in lr_per_sample]
lr_schedule = C.learning_parameter_schedule(lr_per_minibatch, epoch_size=epoch_size)
# Momentum schedule
momentums = C.momentum_schedule(0.9048374180359595, minibatch_size=minibatch_size)
# We use a the Adam optimizer which is known to work well on this dataset
# Feel free to try other optimizers from
# https://www.cntk.ai/pythondocs/cntk.learner.html#module-cntk.learner
learner = C.adam(parameters=model.parameters,
lr=lr_schedule,
momentum=momentums,
gradient_clipping_threshold_per_sample=15,
gradient_clipping_with_truncation=True)
# Setup the progress updater
progress_printer = C.logging.ProgressPrinter(tag='Training', num_epochs=max_epochs)
# Uncomment below for more detailed logging
#progress_printer = ProgressPrinter(freq=100, first=10, tag='Training', num_epochs=max_epochs)
# Instantiate the trainer
trainer = C.Trainer(model, (loss, label_error), learner, progress_printer)
# process minibatches and perform model training
C.logging.log_number_of_parameters(model)
# Assign the data fields to be read from the input
if task == 'slot_tagging':
data_map={x: reader.streams.query, y: reader.streams.slot_labels}
else:
data_map={x: reader.streams.query, y: reader.streams.intent}
t = 0
for epoch in range(max_epochs): # loop over epochs
epoch_end = (epoch+1) * epoch_size
while t < epoch_end: # loop over minibatches on the epoch
data = reader.next_minibatch(minibatch_size, input_map= data_map) # fetch minibatch
trainer.train_minibatch(data) # update model with it
t += data[y].num_samples # samples so far
trainer.summarize_training_progress()
# **Run the trainer**
#
# You can find the complete recipe below.
def do_train():
global z
z = create_model()
reader = create_reader(data['train']['file'], is_training=True)
train(reader, z)
do_train()
# This shows how learning proceeds over epochs (passes through the data).
# For example, after four epochs, the loss, which is the cross-entropy criterion,
# has reduced significantly as measured on the ~18000 samples of this epoch,
# and the same with the error rate on those same 18000 training samples.
#
# The epoch size is the number of samples--counted as *word tokens*, not sentences--to
# process between model checkpoints.
#
# Once the training has completed (a little less than 2 minutes on a Titan-X or a Surface Book),
# you will see an output like this
# ```
# Finished Epoch[10 of 10]: [Training] loss = 0.157653 * 18039, metric = 3.41% * 18039
# ```
# which is the loss (cross entropy) and the metric (classification error) averaged over the final epoch.
#
# On a CPU-only machine, it can be 4 or more times slower. You can try setting
# ```python
# emb_dim = 50
# hidden_dim = 100
# ```
# to reduce the time it takes to run on a CPU, but the model will not fit as well as when the
# hidden and embedding dimension are larger.
# ### Evaluating the model
#
# Like the train() function, we also define a function to measure accuracy on a test set by computing the error over multiple minibatches of test data. For evaluating on a small sample read from a file, you can set a minibatch size reflecting the sample size and run the test_minibatch on that instance of data. To see how to evaluate a single sequence, we provide an instance later in the tutorial.
def evaluate(reader, model_func, task='slot_tagging'):
# Instantiate the model function; x is the input (feature) variable
model = model_func(x)
# Create the loss and error functions
loss, label_error = create_criterion_function_preferred(model, y)
# process minibatches and perform evaluation
progress_printer = C.logging.ProgressPrinter(tag='Evaluation', num_epochs=0)
# Assign the data fields to be read from the input
if task == 'slot_tagging':
data_map={x: reader.streams.query, y: reader.streams.slot_labels}
else:
data_map={x: reader.streams.query, y: reader.streams.intent}
while True:
minibatch_size = 500
data = reader.next_minibatch(minibatch_size, input_map= data_map) # fetch minibatch
if not data: # until we hit the end
break
evaluator = C.eval.Evaluator(loss, progress_printer)
evaluator.test_minibatch(data)
evaluator.summarize_test_progress()
# Now we can measure the model accuracy by going through all the examples in the test set and using the ``C.eval.Evaluator`` method.
def do_test():
reader = create_reader(data['test']['file'], is_training=False)
evaluate(reader, z)
do_test()
z.classify.b.value
# The following block of code illustrates how to evaluate a single sequence. Additionally we show how one can pass in the information using NumPy arrays.
# +
# load dictionaries
query_wl = [line.rstrip('\n') for line in open(data['query']['file'])]
slots_wl = [line.rstrip('\n') for line in open(data['slots']['file'])]
query_dict = {query_wl[i]:i for i in range(len(query_wl))}
slots_dict = {slots_wl[i]:i for i in range(len(slots_wl))}
# let's run a sequence through
seq = 'BOS flights from new york to seattle EOS'
w = [query_dict[w] for w in seq.split()] # convert to word indices
print(w)
onehot = np.zeros([len(w),len(query_dict)], np.float32)
for t in range(len(w)):
onehot[t,w[t]] = 1
#x = C.sequence.input_variable(vocab_size)
pred = z(x).eval({x:[onehot]})[0]
print(pred.shape)
best = np.argmax(pred,axis=1)
print(best)
list(zip(seq.split(),[slots_wl[s] for s in best]))
# -
# ## Modifying the Model
#
# In the following, you will be given tasks to practice modifying CNTK configurations.
# The solutions are given at the end of this document... but please try without!
#
# **A Word About [`Sequential()`](https://www.cntk.ai/pythondocs/layerref.html#sequential)**
#
# Before jumping to the tasks, let's have a look again at the model we just ran.
# The model is described in what we call *function-composition style*.
# ```python
# Sequential([
# Embedding(emb_dim),
# Recurrence(LSTM(hidden_dim), go_backwards=False),
# Dense(num_labels)
# ])
# ```
# You may be familiar with the "sequential" notation from other neural-network toolkits.
# If not, [`Sequential()`](https://www.cntk.ai/pythondocs/layerref.html#sequential) is a powerful operation that,
# in a nutshell, allows to compactly express a very common situation in neural networks
# where an input is processed by propagating it through a progression of layers.
# `Sequential()` takes an list of functions as its argument,
# and returns a *new* function that invokes these functions in order,
# each time passing the output of one to the next.
# For example,
# ```python
# FGH = Sequential ([F,G,H])
# y = FGH (x)
# ```
# means the same as
# ```
# y = H(G(F(x)))
# ```
# This is known as ["function composition"](https://en.wikipedia.org/wiki/Function_composition),
# and is especially convenient for expressing neural networks, which often have this form:
#
# +-------+ +-------+ +-------+
# x -->| F |-->| G |-->| H |--> y
# +-------+ +-------+ +-------+
#
# Coming back to our model at hand, the `Sequential` expression simply
# says that our model has this form:
#
# +-----------+ +----------------+ +------------+
# x -->| Embedding |-->| Recurrent LSTM |-->| DenseLayer |--> y
# +-----------+ +----------------+ +------------+
# ### Task 1: Add Batch Normalization
#
# We now want to add new layers to the model, specifically batch normalization.
#
# Batch normalization is a popular technique for speeding up convergence.
# It is often used for image-processing setups. But could it work for recurrent models, too?
#
# > Note: training with Batch Normalization is currently only supported on GPU.
#
# So your task will be to insert batch-normalization layers before and after the recurrent LSTM layer.
# If you have completed the [hands-on labs on image processing](https://github.com/Microsoft/CNTK/blob/release/2.5/Tutorials/CNTK_201B_CIFAR-10_ImageHandsOn.ipynb),
# you may remember that the [batch-normalization layer](https://www.cntk.ai/pythondocs/layerref.html#batchnormalization-layernormalization-stabilizer) has this form:
# ```
# BatchNormalization()
# ```
# So please go ahead and modify the configuration and see what happens.
#
# If everything went right, you will notice improved convergence speed (`loss` and `metric`)
# compared to the previous configuration.
# +
# Your task: Add batch normalization
def create_model():
with C.layers.default_options(initial_state=0.1):
return C.layers.Sequential([
C.layers.Embedding(emb_dim),
C.layers.Recurrence(C.layers.LSTM(hidden_dim), go_backwards=False),
C.layers.Dense(num_labels)
])
# Enable these when done:
#do_train()
#do_test()
# -
# ### Task 2: Add a Lookahead
#
# Our recurrent model suffers from a structural deficit:
# Since the recurrence runs from left to right, the decision for a slot label
# has no information about upcoming words. The model is a bit lopsided.
# Your task will be to modify the model such that
# the input to the recurrence consists not only of the current word, but also of the next one
# (lookahead).
#
# Your solution should be in function-composition style.
# Hence, you will need to write a Python function that does the following:
#
# * takes no input arguments
# * creates a placeholder (sequence) variable
# * computes the "next value" in this sequence using the `sequence.future_value()` operation and
# * concatenates the current and the next value into a vector of twice the embedding dimension using `splice()`
#
# and then insert this function into `Sequential()`'s list right after the embedding layer.
# +
# Your task: Add lookahead
def create_model():
with C.layers.default_options(initial_state=0.1):
return C.layers.Sequential([
C.layers.Embedding(emb_dim),
C.layers.Recurrence(C.layers.LSTM(hidden_dim), go_backwards=False),
C.layers.Dense(num_labels)
])
# Enable these when done:
#do_train()
#do_test()
# -
# ### Task 3: Bidirectional Recurrent Model
#
# Aha, knowledge of future words help. So instead of a one-word lookahead,
# why not look ahead until all the way to the end of the sentence, through a backward recurrence?
# Let us create a bidirectional model!
#
# Your task is to implement a new layer that
# performs both a forward and a backward recursion over the data, and
# concatenates the output vectors.
#
# Note, however, that this differs from the previous task in that
# the bidirectional layer contains learnable model parameters.
# In function-composition style,
# the pattern to implement a layer with model parameters is to write a *factory function*
# that creates a *function object*.
#
# A function object, also known as [*functor*](https://en.wikipedia.org/wiki/Function_object), is an object that is both a function and an object.
# Which means nothing else that it contains data yet still can be invoked as if it was a function.
#
# For example, `Dense(outDim)` is a factory function that returns a function object that contains
# a weight matrix `W`, a bias `b`, and another function to compute
# `input @ W + b.` (This is using
# [Python 3.5 notation for matrix multiplication](https://docs.python.org/3/whatsnew/3.5.html#whatsnew-pep-465).
# In Numpy syntax it is `input.dot(W) + b`).
# E.g. saying `Dense(1024)` will create this function object, which can then be used
# like any other function, also immediately: `Dense(1024)(x)`.
#
# Let's look at an example for further clarity: Let us implement a new layer that combines
# a linear layer with a subsequent batch normalization.
# To allow function composition, the layer needs to be realized as a factory function,
# which could look like this:
#
# ```python
# def DenseLayerWithBN(dim):
# F = Dense(dim)
# G = BatchNormalization()
# x = placeholder()
# apply_x = G(F(x))
# return apply_x
# ```
#
# Invoking this factory function will create `F`, `G`, `x`, and `apply_x`. In this example, `F` and `G` are function objects themselves, and `apply_x` is the function to be applied to the data.
# Thus, e.g. calling `DenseLayerWithBN(1024)` will
# create an object containing a linear-layer function object called `F`, a batch-normalization function object `G`,
# and `apply_x` which is the function that implements the actual operation of this layer
# using `F` and `G`. It will then return `apply_x`. To the outside, `apply_x` looks and behaves
# like a function. Under the hood, however, `apply_x` retains access to its specific instances of `F` and `G`.
#
# Now back to our task at hand. You will now need to create a factory function,
# very much like the example above.
# You shall create a factory function
# that creates two recurrent layer instances (one forward, one backward), and then defines an `apply_x` function
# which applies both layer instances to the same `x` and concatenate the two results.
#
# Alright, give it a try! To know how to realize a backward recursion in CNTK,
# please take a hint from how the forward recursion is done.
# Please also do the following:
# * remove the one-word lookahead you added in the previous task, which we aim to replace; and
# * make sure each LSTM is using `hidden_dim//2` outputs to keep the total number of model parameters limited.
# +
# Your task: Add bidirectional recurrence
def create_model():
with C.layers.default_options(initial_state=0.1):
return C.layers.Sequential([
C.layers.Embedding(emb_dim),
C.layers.Recurrence(C.layers.LSTM(hidden_dim), go_backwards=False),
C.layers.Dense(num_labels)
])
# Enable these when done:
#do_train()
#do_test()
# -
# The bidirectional model has 40% less parameters than the lookahead one. However, if you go back and look closely
# you may find that the lookahead one trained about 30% faster.
# This is because the lookahead model has both less horizontal dependencies (one instead of two
# recurrences) and larger matrix products, and can thus achieve higher parallelism.
# **Solution 1: Adding Batch Normalization**
# +
def create_model():
with C.layers.default_options(initial_state=0.1):
return C.layers.Sequential([
C.layers.Embedding(emb_dim),
#C.layers.BatchNormalization(), #Remove this comment if running on GPU
C.layers.Recurrence(C.layers.LSTM(hidden_dim), go_backwards=False),
#C.layers.BatchNormalization(), #Remove this comment if running on GPU
C.layers.Dense(num_labels)
])
do_train()
do_test()
# -
# **Solution 2: Add a Lookahead**
# +
def OneWordLookahead():
x = C.placeholder()
apply_x = C.splice(x, C.sequence.future_value(x))
return apply_x
def create_model():
with C.layers.default_options(initial_state=0.1):
return C.layers.Sequential([
C.layers.Embedding(emb_dim),
OneWordLookahead(),
C.layers.Recurrence(C.layers.LSTM(hidden_dim), go_backwards=False),
C.layers.Dense(num_labels)
])
do_train()
do_test()
# -
# **Solution 3: Bidirectional Recurrent Model**
# +
def BiRecurrence(fwd, bwd):
F = C.layers.Recurrence(fwd)
G = C.layers.Recurrence(bwd, go_backwards=True)
x = C.placeholder()
apply_x = C.splice(F(x), G(x))
return apply_x
def create_model():
with C.layers.default_options(initial_state=0.1):
return C.layers.Sequential([
C.layers.Embedding(emb_dim),
BiRecurrence(C.layers.LSTM(hidden_dim//2),
C.layers.LSTM(hidden_dim//2)),
C.layers.Dense(num_labels)
])
do_train()
do_test()
# -
# ## Task overview: Sequence classification
#
# We will reuse the same data for this task. We revisit a sample again:
#
# 19 |S0 178:1 |# BOS |S1 14:1 |# flight |S2 128:1 |# O
# 19 |S0 770:1 |# show |S2 128:1 |# O
# 19 |S0 429:1 |# flights |S2 128:1 |# O
# 19 |S0 444:1 |# from |S2 128:1 |# O
# 19 |S0 272:1 |# burbank |S2 48:1 |# B-fromloc.city_name
# 19 |S0 851:1 |# to |S2 128:1 |# O
# 19 |S0 789:1 |# st. |S2 78:1 |# B-toloc.city_name
# 19 |S0 564:1 |# louis |S2 125:1 |# I-toloc.city_name
# 19 |S0 654:1 |# on |S2 128:1 |# O
# 19 |S0 601:1 |# monday |S2 26:1 |# B-depart_date.day_name
# 19 |S0 179:1 |# EOS |S2 128:1 |# O
#
# The task of the neural network is to look at the query (column `S0`) and predict the
# intent of the sequence (column `S1`). We will ignore the slot-tags (column `S2`) this time.
#
#
# ### Model Creation
#
# The model we will use is a recurrent model consisting of an embedding layer,
# a recurrent LSTM cell, and a dense layer to compute the posterior probabilities. Though very similar to the slot tagging model in this case we look only at the embedding from the last layer:
#
#
# intent "flight"
# ^
# |
# +-------+
# | Dense | ...
# +-------+
# ^
# |
# +------+ +------+ +------+ +------+ +------+
# 0 -->| LSTM |-->| LSTM |-->| LSTM |-->| LSTM |-->| LSTM |-->...
# +------+ +------+ +------+ +------+ +------+
# ^ ^ ^ ^ ^
# | | | | |
# +-------+ +-------+ +-------+ +-------+ +-------+
# | Embed | | Embed | | Embed | | Embed | | Embed | ...
# +-------+ +-------+ +-------+ +-------+ +-------+
# ^ ^ ^ ^ ^
# | | | | |
# w ------>+--------->+--------->+--------->+--------->+------...
# BOS "show" "flights" "from" "burbank"
#
# Or, as a CNTK network description. Please have a quick look and match it with the description above:
# (descriptions of these functions can be found at: [the layers reference](http://cntk.ai/pythondocs/layerref.html))
#
# #### Points to note:
# - The first difference between this model with the previous one is with regards to the specification of the label `y`. Since there is only one label per sequence, we use `C.input_variable`.
# - The second difference is the use of [Stabilizer](http://ieeexplore.ieee.org/document/7472719/). We stabilize the embedded output. The stabilizer adds an additional scalar parameter to the learning that can help our network converge more quickly during training.
# - The third difference is the use of a layer function called `Fold`. As shown in the model above we want the model to have LSTM recurrence except the final one, we set up an LSTM recurrence. The final recurrence will be a Fold operation where we pick the hidden state from the last LSTM block and use it for classification of the entire sequence.
#
# +
# number of words in vocab, slot labels, and intent labels
vocab_size = 943 ; num_intents = 26
# model dimensions
emb_dim = 150
hidden_dim = 300
# Create the containers for input feature (x) and the label (y)
x = C.sequence.input_variable(vocab_size)
y = C.input_variable(num_intents)
def create_model():
with C.layers.default_options(initial_state=0.1):
return C.layers.Sequential([
C.layers.Embedding(emb_dim, name='embed'),
C.layers.Stabilizer(),
C.layers.Fold(C.layers.LSTM(hidden_dim), go_backwards=False),
C.layers.Dense(num_intents, name='classify')
])
# -
# We create the `criterion` function with the new model and correspondingly update the placeholder.
criterion = create_criterion_function(create_model())
criterion.replace_placeholders({criterion.placeholders[0]: C.sequence.input_variable(num_intents)})
# The same train code can be used except for the fact that in this case we provide the `intent` tags as the labels.
def do_train():
global z
z = create_model()
reader = create_reader(data['train']['file'], is_training=True)
train(reader, z, 5, 'intent')
do_train()
# Now we can measure the model accuracy by going through all the examples in the test set and using the C.eval.Evaluator method.
def do_test():
reader = create_reader(data['test']['file'], is_training=False)
evaluate(reader, z, 'intent')
do_test()
z.classify.b.value
# The following block of code illustrates how to evaluate a single sequence. Additionally, we show how one can pass in the information using NumPy arrays.
# +
# load dictionaries
query_wl = [line.rstrip('\n') for line in open(data['query']['file'])]
intent_wl = [line.rstrip('\n') for line in open(data['intent']['file'])]
query_dict = {query_wl[i]:i for i in range(len(query_wl))}
intent_dict = {intent_wl[i]:i for i in range(len(intent_wl))}
# let's run a sequence through
seq = 'BOS flights from new york to seattle EOS'
w = [query_dict[w] for w in seq.split()] # convert to word indices
onehot = np.zeros([len(w),len(query_dict)], np.float32)
for t in range(len(w)):
onehot[t,w[t]] = 1
pred = z(x).eval({x:[onehot]})[0]
best = np.argmax(pred)
print(best)
print(seq, ":", intent_wl[best])
# -
# **Task 4: Use all hidden states for sequence classification**
#
# In the last model, we looked at the output of the last LSTM block. There is another way to model, where we aggregate the output from all the LSTM blocks and use the aggregated output to the final `Dense` layer.
#
# So your task will be to replace the `C.layers.Fold` with `C.layers.Recurrence` layer function. This is the explicit way of setting up recurrence. You will aggregate all the intermediate outputs from LSTM blocks using `C.sequence.reduce_sum`. Note: this is different from last model where we looked only at the output of the last LSTM block.
#
# So please go ahead and modify the configuration and see what happens.
#
# If everything went right, you will notice improved accuracy (`metric`). The solution is presented right after, but we suggest you refrain from looking at the solution.
# Replace the line with Fold operation
def create_model():
with C.layers.default_options(initial_state=0.1):
return C.layers.Sequential([
C.layers.Embedding(emb_dim, name='embed'),
C.layers.Stabilizer(),
C.layers.Fold(C.layers.LSTM(hidden_dim), go_backwards=False),
C.layers.Dense(num_intents, name='classify')
])
# Enable these when done:
#do_train()
#do_test()
# Aggregating all the intermediate states improves the accuracy for the same number of iterations without any significant increase in the computation time.
# **Solution 4: Use all hidden states for sequence classification**
# +
def create_model():
with C.layers.default_options(initial_state=0.1):
return C.layers.Sequential([
C.layers.Embedding(emb_dim, name='embed'),
C.layers.Stabilizer(),
C.sequence.reduce_sum(C.layers.Recurrence(C.layers.LSTM(hidden_dim), go_backwards=False)),
C.layers.Dense(num_intents, name='classify')
])
do_train()
do_test()
|
Tutorials/CNTK_202_Language_Understanding.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
"""
@author: Ajay
"""
# +
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
import torch
import torch.nn as nn
import torch.optim as optim
from torch.utils.data import Dataset, DataLoader
from sklearn.preprocessing import StandardScaler, MinMaxScaler
from sklearn.model_selection import train_test_split
from sklearn.metrics import confusion_matrix, classification_report
from sklearn.datasets import make_classification
from pyNM.cf_matrix import make_confusion_matrix
from pyNM.spiking_binary_classifier import *
from plot_metric.functions import MultiClassClassification, BinaryClassification
# +
#fixing random state
random_state=1234
# Generate 2 class dataset
X, Y = make_classification(n_samples=10000, n_classes=2, weights=[1,1], random_state=1)
# split into train/test sets
#X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.5, random_state=2)
# Load dataset (we just selected 4 classes of digits)
#X, Y = load_digits(n_class=4, return_X_y=True)
print(f'Predictors: {X}')
print(f'Outcome: {Y}')
print(f'Distribution of target:')
print(pd.value_counts(Y))
# Add noisy features to make the problem more harder
#random_state = np.random.RandomState(123)
#n_samples, n_features = X.shape
#X = np.c_[X, random_state.randn(n_samples, 1000 * n_features)]
## Spliting data into train and test sets.
X, X_test, y, y_test = train_test_split(X, Y, test_size=0.4,
random_state=123)
## Spliting train data into training and validation sets.
X_train, X_valid, y_train, y_valid = train_test_split(X, y, test_size=0.2,
random_state=1)
print('Data shape:')
print('X_train: %s, X_valid: %s, X_test: %s \n' %(X_train.shape, X_valid.shape,
X_test.shape))
# +
# Scale data to have mean '0' and variance '1'
# which is importance for convergence of the neural network
scaler = StandardScaler()
X_train = scaler.fit_transform(X_train)
X_valid = scaler.transform(X_valid)
X_test = scaler.transform(X_test)
X_train, y_train = np.array(X_train), np.array(y_train)
X_valid, y_valid = np.array(X_valid), np.array(y_valid)
X_test, y_test = np.array(X_test), np.array(y_test)
# -
EPOCHS = 50
BATCH_SIZE = 64
LEARNING_RATE = 0.001
# +
## train data
class trainData(Dataset):
def __init__(self, X_data, y_data):
self.X_data = X_data
self.y_data = y_data
def __getitem__(self, index):
return self.X_data[index], self.y_data[index]
def __len__ (self):
return len(self.X_data)
train_data = trainData(torch.FloatTensor(X_train),
torch.FloatTensor(y_train))
## validation data
class valData(Dataset):
def __init__(self, X_data, y_data):
self.X_data = X_data
self.y_data = y_data
def __getitem__(self, index):
return self.X_data[index], self.y_data[index]
def __len__ (self):
return len(self.X_data)
val_data = valData(torch.FloatTensor(X_valid),
torch.FloatTensor(y_valid))
## test data
class testData(Dataset):
def __init__(self, X_data):
self.X_data = X_data
def __getitem__(self, index):
return self.X_data[index]
def __len__ (self):
return len(self.X_data)
test_data = testData(torch.FloatTensor(X_test))
# -
train_loader = DataLoader(dataset=train_data, batch_size=BATCH_SIZE, shuffle=True)
val_loader = DataLoader(dataset=val_data, batch_size=BATCH_SIZE, shuffle=True)
test_loader = DataLoader(dataset=test_data, batch_size=1)
class binaryClassification(nn.Module):
def __init__(self):
super(binaryClassification, self).__init__()
# Number of input features is 12.
self.layer_1 = nn.Linear(20, 64)
#self.layer_2 = nn.Linear(100, 1)
self.layer_out = nn.Linear(64, 1)
self.relu = nn.ReLU()
self.dropout = nn.Dropout(p=0.1)
self.batchnorm1 = nn.BatchNorm1d(64)
#self.batchnorm2 = nn.BatchNorm1d(64)
def forward(self, inputs):
x = self.relu(self.layer_1(inputs))
x = self.batchnorm1(x)
#x = self.relu(self.layer_2(x))
#x = self.batchnorm2(x)
x = self.dropout(x)
x = self.layer_out(x)
return x
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
print(device)
#model = binaryClassification()
#model = SpikingNeuralNetwork(device, X_train.shape[1], hidden_dim_l1=64, hidden_dim_l2=64, n_time_steps=64, begin_eval=0)
model = SpikingNeuralNetwork(device, X_train.shape[1], n_time_steps=500, begin_eval=0)
model.to(device)
print(model)
criterion = nn.BCEWithLogitsLoss()
optimizer = optim.Adam(model.parameters(), lr=LEARNING_RATE)
def binary_acc(y_pred, y_test):
y_pred_tag = torch.round(torch.sigmoid(y_pred))
correct_results_sum = (y_pred_tag == y_test).sum().float()
acc = correct_results_sum/y_test.shape[0]
acc = torch.round(acc * 100)
return acc
model.train()
for e in range(1,EPOCHS+1):
epoch_loss = 0
epoch_acc = 0
for X_batch, y_batch in train_loader:
X_batch, y_batch = X_batch.to(device), y_batch.to(device)
optimizer.zero_grad()
y_pred = model(X_batch)
loss = criterion(y_pred, y_batch.unsqueeze(1))
acc = binary_acc(y_pred, y_batch.unsqueeze(1))
loss.backward()
optimizer.step()
epoch_loss += loss.item()
epoch_acc += acc.item()
print(f'Epoch {e+0:03}: | Loss: {epoch_loss/len(train_loader):.5f} | Acc: {epoch_acc/len(train_loader):.3f}')
# +
y_pred_list = []
model.eval()
with torch.no_grad():
for X_batch in test_loader:
X_batch = X_batch.to(device)
y_test_pred = model(X_batch)
y_test_pred = torch.sigmoid(y_test_pred)
y_pred_tag = torch.round(y_test_pred)
y_pred_list.append(y_pred_tag.cpu().numpy())
y_pred_list = [a.squeeze().tolist() for a in y_pred_list]
# -
#Get the confusion matrix
cf_matrix = confusion_matrix(y_test, y_pred_list)
print(cf_matrix)
make_confusion_matrix(cf_matrix, figsize=(8,6), cbar=False, title='CF Matrix')
'''
[[1978 24]
[ 622 1376]]
[[1376 626]
[ 11 1987]]
[[1972 30]
[ 623 1375]]
'''
# NON-SPIKING NN
'''
[[1963 39]
[ 47 1951]]
[[1975 27]
[ 36 1962]]
'''
print(classification_report(y_test, y_pred_list))
# +
# report
# Visualisation of plots
bc = BinaryClassification(y_test, y_pred_list, labels=[0, 1])
# Figures
plt.figure(figsize=(15,10))
plt.subplot2grid(shape=(2,6), loc=(0,0), colspan=2)
bc.plot_roc_curve()
plt.subplot2grid((2,6), (0,2), colspan=2)
bc.plot_precision_recall_curve()
plt.subplot2grid((2,6), (0,4), colspan=2)
bc.plot_class_distribution()
plt.subplot2grid((2,6), (1,1), colspan=2)
bc.plot_confusion_matrix()
plt.subplot2grid((2,6), (1,3), colspan=2)
bc.plot_confusion_matrix(normalize=True)
# Save figure
plt.savefig('../figures/images/example_binary_classification.png')
# Display Figure
plt.show()
plt.close()
# Full report of the classification
bc.print_report()
# Example custom param using dictionnary
param_pr_plot = {
'c_pr_curve':'blue',
'c_mean_prec':'cyan',
'c_thresh_lines':'red',
'c_f1_iso':'green',
'beta': 2,
}
plt.figure(figsize=(6,6))
bc.plot_precision_recall_curve(**param_pr_plot)
# Save figure
plt.savefig('../figures/images/example_binary_class_PRCurve_custom.png')
# Display Figure
plt.show()
plt.close()
# -
|
pyNM/spiking-binary-classifier-model.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="zwNCA2HzDVI7"
from sklearn.preprocessing import LabelEncoder
from sklearn.preprocessing import MinMaxScaler
import numpy as np
import cv2
import os
import h5py
# + colab={"base_uri": "https://localhost:8080/"} id="Lqo7yiTGENTT" outputId="a73166a0-1aeb-4819-dadf-722842a4334e"
# !pip install mahotas # Fast Computer Vision Algorithms Package using C++
# + id="04a8HPj_DVG5"
import mahotas
# + id="xGNRtBn_DVEZ"
images_per_class = 800
fixed_size = tuple((500, 500))
train_path = "/content/drive/MyDrive/dataset/train"
h5_train_data = '/content/drive/MyDrive/dataset/output/train_data.h5'
h5_train_labels = '/content/drive/MyDrive/dataset/output/train_labels.h5'
bins = 8
# + id="NCh9QG1tDVCa"
# Converting each image to RGB from BGR format
def rgb_bgr(image):
rgb_img = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
return rgb_img
# + id="l8_-FeStDVAJ"
# Conversion to HSV image format from RGB
def bgr_hsv(rgb_img):
hsv_img = cv2.cvtColor(rgb_img, cv2.COLOR_RGB2HSV)
return hsv_img
# + id="x1ltHr50DU92"
# Image Segmentation for Extraction of Green and Brown color
def img_segmentation(rgb_img,hsv_img):
lower_green = np.array([25,0,20])
upper_green = np.array([100,255,255])
healthy_mask = cv2.inRange(hsv_img, lower_green, upper_green)
result = cv2.bitwise_and(rgb_img,rgb_img, mask=healthy_mask)
lower_brown = np.array([10,0,10])
upper_brown = np.array([30,255,255])
disease_mask = cv2.inRange(hsv_img, lower_brown, upper_brown)
disease_result = cv2.bitwise_and(rgb_img, rgb_img, mask=disease_mask)
final_mask = healthy_mask + disease_mask
final_result = cv2.bitwise_and(rgb_img, rgb_img, mask=final_mask)
return final_result
# + id="SenmV-NpE1UQ"
# feature-descriptor-1: Hu Moments
def fd_hu_moments(image):
image = cv2.cvtColor(image, cv2.COLOR_BGR2GRAY)
feature = cv2.HuMoments(cv2.moments(image)).flatten()
return feature
# + id="YaYiIjq_E1SA"
# feature-descriptor-2: Haralick Texture
def fd_haralick(image):
gray = cv2.cvtColor(image, cv2.COLOR_BGR2GRAY)
haralick = mahotas.features.haralick(gray).mean(axis=0)
return haralick
# + id="Dd2VOM1QE1Px"
# feature-descriptor-3: Color Histogram
def fd_histogram(image, mask=None):
image = cv2.cvtColor(image, cv2.COLOR_BGR2HSV)
hist = cv2.calcHist([image], [0, 1, 2], None, [bins, bins, bins], [0, 256, 0, 256, 0, 256])
cv2.normalize(hist, hist)
return hist.flatten()
# + colab={"base_uri": "https://localhost:8080/"} id="fZotsDdHE1NR" outputId="14fe231d-377f-4836-8a65-fe514d616ba8"
# get the training labels
train_labels = os.listdir(train_path)
# sort the training labels
train_labels.sort()
print(train_labels)
# empty lists to hold feature vectors and labels
global_features = []
labels = []
# + colab={"base_uri": "https://localhost:8080/", "height": 298} id="VSXLwo7ySqhq" outputId="a1263775-4a02-44f7-884b-33a9092cdfa6"
import matplotlib.image as mpimg
image_diseased = mpimg.imread("/content/drive/MyDrive/dataset/train/diseased/3.jpg")
plt.title("Diseased Leaf Example")
plt.imshow(image_diseased)
# + colab={"base_uri": "https://localhost:8080/", "height": 298} id="whMRU8hTUnbL" outputId="79db429a-1163-4d0b-d5c2-d18736fb2e3a"
image_healthy = mpimg.imread("/content/drive/MyDrive/dataset/train/healthy/3.jpg")
plt.title("Healthy Leaf Example")
plt.imshow(image_healthy)
# + colab={"base_uri": "https://localhost:8080/"} id="t_SLhDrzE1Kw" outputId="acfcf905-257f-4d28-e751-2f3a141914f2"
# loop over the training data sub-folders
for training_name in train_labels:
# join the training data path and each species training folder
dir = os.path.join(train_path, training_name)
# get the current training label
current_label = training_name
# loop over the images in each sub-folder
for x in range(1,images_per_class+1):
# get the image file name
file = dir + "/" + str(x) + ".jpg"
# read the image and resize it to a fixed-size
image = cv2.imread(file)
image = cv2.resize(image, fixed_size)
# Running Function Bit By Bit
RGB_BGR = rgb_bgr(image)
BGR_HSV = bgr_hsv(RGB_BGR)
IMG_SEGMENT = img_segmentation(RGB_BGR,BGR_HSV)
# Call for Global Feature Descriptors
fv_hu_moments = fd_hu_moments(IMG_SEGMENT)
fv_haralick = fd_haralick(IMG_SEGMENT)
fv_histogram = fd_histogram(IMG_SEGMENT)
# Concatenate
global_feature = np.hstack([fv_histogram, fv_haralick, fv_hu_moments])
# update the list of labels and feature vectors
labels.append(current_label)
global_features.append(global_feature)
print("[STATUS] processed folder: {}".format(current_label))
print("[STATUS] completed Global Feature Extraction...")
# + colab={"base_uri": "https://localhost:8080/"} id="VMw48xjGE1IA" outputId="824803a2-e86c-465f-c4c3-94a20d838e70"
# Encode The Target labels
targetNames = np.unique(labels)
le = LabelEncoder()
target = le.fit_transform(labels)
print("training labels encoded...!")
# + colab={"base_uri": "https://localhost:8080/"} id="OGmNEK3dIdiS" outputId="f02be0c0-6c0e-494f-bc9b-15e6cdfde748"
# Scale features in the range (0-1)
from sklearn.preprocessing import MinMaxScaler # Could use Standard Scaler too!
scaler = MinMaxScaler(feature_range=(0, 1))
rescaled_features = scaler.fit_transform(global_features)
print("feature vector normalized...")
# + colab={"base_uri": "https://localhost:8080/"} id="mAwCCzkFR2w9" outputId="7d5a4a41-7468-4113-c7ab-abfbe2354dd0"
print(global_features)
# + colab={"base_uri": "https://localhost:8080/"} id="phCqpWPZIdf2" outputId="d1099196-4896-43f0-afba-39d41251b300"
# Save the feature vectors using HDF5
h5f_data = h5py.File(h5_train_data, 'w')
h5f_data.create_dataset('dataset_1', data=np.array(rescaled_features))
# + colab={"base_uri": "https://localhost:8080/"} id="7Le__LvDIdcz" outputId="740ec0a7-f3a4-4086-f3c4-bff569a33891"
h5f_label = h5py.File(h5_train_labels, 'w')
h5f_label.create_dataset('dataset_1', data=np.array(target))
# + id="2dERIogxIdXp"
h5f_data.close()
h5f_label.close()
# + id="vPeS2Ny_MpvU" colab={"base_uri": "https://localhost:8080/"} outputId="4cef6d39-0177-488c-cee8-6157f8211ce1"
# TRAINING OUR MODEL
#-----------------------------------
import h5py
import numpy as np
import os
import glob
import cv2
import warnings
from matplotlib import pyplot
from sklearn.model_selection import train_test_split, cross_val_score
from sklearn.model_selection import KFold, StratifiedKFold
from sklearn.metrics import confusion_matrix, accuracy_score, classification_report
from sklearn.linear_model import LogisticRegression
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.neighbors import KNeighborsClassifier
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
from sklearn.naive_bayes import GaussianNB
from sklearn.svm import SVC
from sklearn.externals import joblib
warnings.filterwarnings('ignore')
#--------------------
# tunable-parameters
#--------------------
num_trees = 100
test_size = 0.20
seed = 9
train_path = "/content/drive/MyDrive/dataset/train"
test_path = "/content/drive/MyDrive/dataset/test"
h5_train_data = '/content/drive/MyDrive/dataset/output/train_data.h5'
h5_train_labels = '/content/drive/MyDrive/dataset/output/train_labels.h5'
scoring = "accuracy"
# get the training labels
train_labels = os.listdir(train_path)
# sort the training labels
train_labels.sort()
if not os.path.exists(test_path):
os.makedirs(test_path)
# create all the machine learning models
models = []
models.append(('LR', LogisticRegression(random_state=seed)))
models.append(('LDA', LinearDiscriminantAnalysis()))
models.append(('KNN', KNeighborsClassifier()))
models.append(('CART', DecisionTreeClassifier(random_state=seed)))
models.append(('RF', RandomForestClassifier(n_estimators=num_trees, random_state=seed)))
models.append(('NB', GaussianNB()))
models.append(('SVM', SVC(random_state=seed)))
# variables to hold the results and names
results = []
names = []
# import the feature vector and trained labels
h5f_data = h5py.File(h5_train_data, 'r')
h5f_label = h5py.File(h5_train_labels, 'r')
global_features_string = h5f_data['dataset_1']
global_labels_string = h5f_label['dataset_1']
global_features = np.array(global_features_string)
global_labels = np.array(global_labels_string)
h5f_data.close()
h5f_label.close()
# verify the shape of the feature vector and labels
print("Features shape: {}".format(global_features.shape))
print("Labels shape: {}".format(global_labels.shape))
print("Training Started...")
# + colab={"base_uri": "https://localhost:8080/"} id="QMUHNXdlNb66" outputId="2333db58-40dd-4de2-ccf8-30ad919b054a"
# split the training and testing data
(trainDataGlobal, testDataGlobal, trainLabelsGlobal, testLabelsGlobal) = train_test_split(np.array(global_features),
np.array(global_labels),
test_size=test_size,
random_state=seed)
print("[STATUS] splitted train and test data...")
print("Train data : {}".format(trainDataGlobal.shape))
print("Test data : {}".format(testDataGlobal.shape))
# + colab={"base_uri": "https://localhost:8080/"} id="MpA1MCVMNfSh" outputId="d08d6288-c329-4c8a-a86c-23d0b4ecc37f"
trainDataGlobal
# + colab={"base_uri": "https://localhost:8080/", "height": 412} id="PCmj2EqYNfQm" outputId="8908696e-fbad-4a5a-be22-9a8a47395597"
import matplotlib.pyplot as plt
# 10-fold cross validation
for name, model in models:
kfold = KFold(n_splits=10, random_state=seed)
cv_results = cross_val_score(model, trainDataGlobal, trainLabelsGlobal, cv=kfold, scoring=scoring)
results.append(cv_results)
names.append(name)
msg = "%s: %f (%f)" % (name, cv_results.mean(), cv_results.std())
print(msg)
# boxplot algorithm comparison
fig = plt.figure()
fig.suptitle('Comparing All Algorithms Used')
ax = fig.add_subplot(111)
plt.boxplot(results)
ax.set_xticklabels(names)
plt.show()
# + id="2E4WpH-zOB1y"
clf = RandomForestClassifier(n_estimators=num_trees, random_state=seed)
# + colab={"base_uri": "https://localhost:8080/"} id="JG7uo5QWNfOh" outputId="2a90f479-4b21-4c90-8afa-ff8a92ead833"
clf.fit(trainDataGlobal, trainLabelsGlobal)
# + id="MBS9sSSCNfMI"
y_predict=clf.predict(testDataGlobal)
# + colab={"base_uri": "https://localhost:8080/"} id="NgJfB5H4NfJn" outputId="988e4d63-0949-410b-d965-de27c0723231"
y_predict
# + id="fbAj_TyuOONB"
cm = confusion_matrix(testLabelsGlobal,y_predict)
# + colab={"base_uri": "https://localhost:8080/", "height": 286} id="L9fvLI7WOOFM" outputId="7a9dca12-5309-4728-aec6-5c949e057ff7"
import seaborn as sns
sns.heatmap(cm ,annot=True)
# + id="PnQ666TKOTnv"
from sklearn.metrics import accuracy_score
# + colab={"base_uri": "https://localhost:8080/"} id="aBRgfVJYOTpz" outputId="fbf81ba8-8440-426f-c2bc-2038fd0846ef"
accuracy_score(testLabelsGlobal, y_predict)
# + id="_nheGIcLSqfY"
# + id="07FoyeigSqc6"
|
Plant Disease Detection Using Machine Learning.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia
# language: julia
# name: julia-1.5
# ---
# # Convergence of RK methods
#
# We consider the IVP $u'=\sin[(u+t)^2]$ over $0\le t \le 4$, with $u(0)=-1$.
using FundamentalsNumericalComputation
# +
f = (u,p,t) -> sin((t+u)^2)
tspan = (0.0,4.0)
u0 = -1.0
ivp = ODEProblem(f,u0,tspan)
# -
# We use a `DifferentialEquations` solver to construct an accurate approximation to the exact solution.
u_exact = solve(ivp,Tsit5(),reltol=1e-14,abstol=1e-14);
# Now we perform a convergence study of our two Runge--Kutta implementations.
# +
n = @. 50*2^(0:5)
err_IE2 = zeros(size(n))
err_RK4 = zeros(size(n))
for (j,n) = enumerate(n)
t,u = FNC.ie2(ivp,n)
err_IE2[j] = maximum( @.abs(u_exact(t)-u) )
t,u = FNC.rk4(ivp,n)
err_RK4[j] = maximum( @.abs(u_exact(t)-u) )
end
pretty_table((n=n,e2=err_IE2,e4=err_RK4),["n","error in IE2","error in RK4"],backend=:html)
# -
# The amount of computational work at each time step is assumed to be proportional to the number of stages. Let's compare on an apples-to-apples basis by using the number of $f$-evaluations on the horizontal axis.
# +
using Plots
plot([2n 4n],[err_IE2 err_RK4],m=:o,label=["IE2" "RK4"],
xaxis=(:log10,"f-evaluations"),yaxis=(:log10,"inf-norm error"),
title="Convergence of RK methods",leg=:bottomleft)
plot!(2n,0.01*(n/n[1]).^(-2),l=:dash,label="2nd order")
plot!(4n,1e-6*(n/n[1]).^(-4),l=:dash,label="4th order")
# -
# The fourth-order variant is more efficient in this problem over a wide range of accuracy.
|
book/ivp/demos/rk-converge.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (TensorFlow 2.6 Python 3.8 CPU Optimized)
# language: python
# name: python3__SAGEMAKER_INTERNAL__arn:aws:sagemaker:us-east-1:081325390199:image/tensorflow-2.6-cpu-py38-ubuntu20.04-v1
# ---
# # Training Job
# #### In this notebook we will focus on reading the data that we pre-processed previouslly and then start training
# +
import boto3
import sagemaker
from sagemaker import get_execution_role
region = boto3.session.Session().region_name
role = get_execution_role()
# -
# ## 1. Reading Data
# +
import argparse
import os
import warnings
import pandas as pd
import pathlib
import numpy as np
import tensorflow as tf
from tensorflow.keras import layers
from tensorflow.keras import models
from IPython import display
# -
# # Writing the Entrypoint script
# #### For starting a training job we need a script which will be used by the training job
# +
# %%writefile utils/train.py
import argparse
import os
import warnings
import itertools
import io
import pandas as pd
import matplotlib.pyplot as plt
import pathlib
import numpy as np
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import layers
from tensorflow.keras import models
from IPython import display
class EvaluationCallback(keras.callbacks.Callback):
def on_test_end(self, logs=None):
self.log_confusion_matrix()
def plot_confusion_matrix(self, cm, class_names):
"""
Returns a matplotlib figure containing the plotted confusion matrix.
Args:
cm (array, shape = [n, n]): a confusion matrix of integer classes
class_names (array, shape = [n]): String names of the integer classes
"""
figure = plt.figure(figsize=(8, 8))
plt.imshow(cm, interpolation='nearest', cmap=plt.cm.Blues)
plt.title("Confusion matrix")
plt.colorbar()
tick_marks = np.arange(len(class_names))
plt.xticks(tick_marks, class_names, rotation=45)
plt.yticks(tick_marks, class_names)
# Normalize the confusion matrix.
cm = cm.numpy()
cm = np.around(cm.astype('float') / cm.sum(axis=1)[:, np.newaxis], decimals=2)
# Use white text if squares are dark; otherwise black.
threshold = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
color = "white" if cm[i, j] > threshold else "black"
plt.text(j, i, cm[i, j], horizontalalignment="center", color=color)
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
# plt.show()
return figure
def plot_to_image(self, figure):
"""
Converts the matplotlib plot specified by 'figure' to a PNG image and
returns it. The supplied figure is closed and inaccessible after this call.
"""
buf = io.BytesIO()
# Use plt.savefig to save the plot to a PNG in memory.
plt.savefig(buf, format='png')
# Closing the figure prevents it from being displayed directly inside
# the notebook.
plt.close(figure)
buf.seek(0)
# Use tf.image.decode_png to convert the PNG buffer
# to a TF image. Make sure you use 4 channels.
image = tf.image.decode_image(buf.getvalue(), channels=4)
# print(base64.b64decode((buf.getvalue())))
# Use tf.expand_dims to add the batch dimension
image = tf.expand_dims(image, 0)
return image
def log_confusion_matrix(self):
# Use the model to predict the values from the test_images.
test_audio = []
test_labels = []
for audio, label in test_ds:
print(audio.shape)
break
for audio, label in test_ds:
test_audio.append(audio.numpy())
test_labels.append(label.numpy())
test_audio = np.array(test_audio, dtype=object)
test_labels = np.array(test_labels, dtype=object)
print("test_audio shape : ", test_audio.shape)
y_pred = np.argmax(model.predict(test_audio), axis=1)
y_true = test_labels
# Calculate the confusion matrix using sklearn.metrics
cm = tf.math.confusion_matrix(y_true, y_pred)
print(cm)
figure = self.plot_confusion_matrix(cm, class_names=['yes', 'no'])
cm_image = self.plot_to_image(figure)
# print(cm_image)
# Log the confusion matrix as an image summary.
with file_writer_cm.as_default():
tf.summary.image("Confusion Matrix", cm_image, step=0)
if __name__ =='__main__':
parser = argparse.ArgumentParser()
# hyperparameters sent by the client are passed as command-line arguments to the script.
parser.add_argument('--epochs', type=int, default=10)
parser.add_argument('--batch_size', type=int, default=64)
# input data and model directories
parser.add_argument('--model_dir', type=str)
parser.add_argument('--train', type=str, default=os.environ.get('SM_CHANNEL_TRAIN'))
parser.add_argument('--val', type=str, default=os.environ.get('SM_CHANNEL_VAL'))
parser.add_argument('--test', type=str, default=os.environ.get('SM_CHANNEL_TEST'))
parser.add_argument('--labels', type=str, default=os.environ.get('SM_CHANNEL_LABELS'))
args, _ = parser.parse_known_args()
spectrogram_ds = tf.data.experimental.load(args.train)
train_ds = spectrogram_ds
val_ds = tf.data.experimental.load(args.val)
test_ds = tf.data.experimental.load(args.test)
commands = np.load(args.labels + "/commands.npy")
batch_size = args.batch_size
train_ds = train_ds.batch(batch_size)
val_ds = val_ds.batch(batch_size)
test_ds_batch = test_ds.batch(batch_size)
#Add Dataset.cache and Dataset.prefetch operations to reduce read latency while training the model:
AUTOTUNE = tf.data.AUTOTUNE
train_ds = train_ds.cache().prefetch(AUTOTUNE)
val_ds = val_ds.cache().prefetch(AUTOTUNE)
#For the model, you'll use a simple convolutional neural network (CNN), since you have transformed the audio files into spectrogram images.
# Your tf.keras.Sequential model will use the following Keras preprocessing layers:
# - tf.keras.layers.Resizing: to downsample the input to enable the model to train faster.
# - tf.keras.layers.Normalization: to normalize each pixel in the image based on its mean and standard deviation.
# - For the Normalization layer, its adapt method would first need to be called on the training data in order to compute aggregate statistics (that is, the mean and the standard deviation).
for spectrogram, _ in spectrogram_ds.take(1):
input_shape = spectrogram.shape
print('Input shape:', input_shape)
num_labels = len(commands)
print(num_labels)
for spectrogram, _ in val_ds.take(1):
tmp = spectrogram.shape
print('val shape:', tmp)
# Instantiate the `tf.keras.layers.Normalization` layer.
norm_layer = layers.Normalization()
# Fit the state of the layer to the spectrograms
# with `Normalization.adapt`.
norm_layer.adapt(data=spectrogram_ds.map(map_func=lambda spec, label: spec))
model = models.Sequential([
layers.Input(shape=input_shape),
# Downsample the input.
layers.Resizing(32, 32),
# Normalize.
norm_layer,
layers.Conv2D(32, 3, activation='relu'),
layers.Conv2D(64, 3, activation='relu'),
layers.MaxPooling2D(),
layers.Dropout(0.25),
layers.Flatten(),
layers.Dense(128, activation='relu'),
layers.Dropout(0.5),
layers.Dense(num_labels),
])
print(model.summary())
# Configure the Keras model with the Adam optimizer and the cross-entropy loss:
model.compile(
optimizer=tf.keras.optimizers.Adam(),
loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=['accuracy'],
)
LOG_DIR = "/opt/ml/output/tensorboard"
tensorboard_callback = tf.keras.callbacks.TensorBoard(log_dir=LOG_DIR, histogram_freq=1)
file_writer_cm = tf.summary.create_file_writer(LOG_DIR + '/cm')
EPOCHS = args.epochs
history = model.fit(
train_ds,
validation_data=val_ds,
epochs=EPOCHS,
callbacks=[tf.keras.callbacks.EarlyStopping(verbose=1, patience=2), tensorboard_callback],
)
# Creating Confusion matrix using test_ds
# Define the per-epoch callback.
result = model.evaluate(
test_ds_batch,
callbacks=[tensorboard_callback, EvaluationCallback()]
)
print("test set results:", dict(zip(model.metrics_names, result)))
# Save model to the SM_MODEL_DIR path
model_path = os.environ.get('SM_MODEL_DIR') + "/1"
print("Saving model to {}".format(model_path))
model.save(model_path)
# -
# # Creating the Training Job
# ! pip install -U sagemaker
# + jupyter={"outputs_hidden": true}
from sagemaker.tensorflow import TensorFlow
from sagemaker.debugger import TensorBoardOutputConfig
tensorboard_output_config = TensorBoardOutputConfig(
s3_output_path = 's3://sagemaker-studio-062044820001-7qbctb3w94p/Training/tensorboard_log_folder/'
)
tf_estimator = TensorFlow(
entry_point="utils/train.py",
role=role,
instance_count=1,
instance_type="ml.m5.large",
framework_version="2.6",
py_version="py38",
tensorboard_output_config=tensorboard_output_config,
output_path = "s3://sagemaker-studio-062044820001-7qbctb3w94p/Training/models/1"
)
data_path = "s3://sagemaker-studio-062044820001-7qbctb3w94p/Datasets/mini-speech-commands/pre-processed/"
tf_estimator.fit({'train': data_path + "train/",
'val': data_path + "val/",
'test': data_path + "test/",
'labels': data_path + "commands/"})
# -
tensorboard_s3_output_path = tf_estimator.latest_job_tensorboard_artifacts_path()
tensorboard_s3_output_path
# +
# To check the plots and evulation on the validation set run the following commands in the system termnal of the notebook
# pip install tensorboard
# tensorboard --logdir '<tensorboard_s3_output_path>'
# +
instance_type = "ml.t2.medium"
predictor = tf_estimator.deploy(
initial_instance_count=1,
instance_type=instance_type,
)
# -
sample = []
for spectrogram, _ in test_ds.take(1):
sample = spectrogram
sample.shapeb
result = predictor.predict(np.array(sample))
|
.ipynb_checkpoints/training-job-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os.path
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.linear_model import LinearRegression
from traitlets import traitlets
from IPython.display import display
from ipywidgets import HBox, VBox, BoundedFloatText, BoundedIntText, Text, Layout, Button
# # Processing demographic data
# We have retrieved demographic data from the data portal of the **National Centre for Statistics & Information** of the *Sultanate of Oman* (https://data.gov.om/)
#
# Data are formatted in a two-columns file, where<br>
# * The __first__ column contains _years_.<br>
# * The __second__ column, which must be named *Population*, contains integer numbers and reports the total population in the Muscat region per year [2010-2019]
# +
if os.path.exists("./playgrounds/data/JNNP_2020/Epidemiology_Oman.txt"):
epi_file = "./playgrounds/data/JNNP_2020/Epidemiology_Oman.txt"
elif not os.path.exists("./data/JNNP_2020/Epidemiology_Oman.txt"):
# !git clone https://github.com/mazzalab/playgrounds.git
epi_file = "./playgrounds/data/JNNP_2020/Epidemiology_Oman.txt"
else:
epi_file = "./data/JNNP_2020/Epidemiology_Oman.txt"
df = pd.read_csv(epi_file, sep='\t', index_col=0)
print(df)
# -
# ## Predicting Muscat population growth
# The observed period of time starts in **2014** and ends in the year specified below.
# +
style = {'description_width': 'initial'}
simulation_end_text = BoundedIntText(
min=2015,
max=2100,
step=1,
value=2050,
description='Simulate from 2014 until:', style=style)
class GenerateTimeButton(Button):
def __init__(self, value=None, *args, **kwargs):
super(GenerateTimeButton, self).__init__(*args, **kwargs)
# Create the value attribute.
self.add_traits(value=traitlets.Any(value))
# Generate time period (list of years) to be simulated
def on_generate_time_button_clicked(button):
button.value = np.arange(2014, simulation_end_text.value+1, 1).reshape((-1, 1))
print("Simulation time points generated")
generate_time_button = GenerateTimeButton(
description="Generate",
button_style='info',
tooltip='Generate simulation time points'
)
generate_time_button.value=np.array([])
generate_time_button.on_click(on_generate_time_button_clicked)
hbox_time = HBox([simulation_end_text, generate_time_button])
display(hbox_time)
# -
# Linear regression analysis is conducted on demographic data using the *LinearRegression* module from the Python **sklearn** package. The typical linear regression equation: \begin{align}y & = mx + b\end{align} is fitted and *coefficient of determination ($r^2$)*, *intercept* ($b$) and *slope* ($m$) are inferred.
# +
if not generate_time_button.value.any():
simulation_end_text.value=2050
generate_time_button.click()
x_new = generate_time_button.value
#######################################
# %matplotlib inline
x = df.index.values.reshape((-1, 1))
y = df.Population
model = LinearRegression().fit(x, y)
r_sq = model.score(x, y)
print(" ")
print('coefficient of determination:', np.round(r_sq, 3))
print('intercept (b):', np.round(model.intercept_, 3))
print('slope (m):', np.round(model.coef_, 3))
print('')
# Predict response of known data
y_pred = model.predict(x)
print('time:', np.transpose(x)[0], sep='\t\t\t\t')
print('predicted response:', np.around(y_pred), sep='\t\t')
# Plot outputs
axis_scale = 1e5
plt.scatter(x, y/axis_scale, color='black', label="act. population")
plt.plot(x, y_pred/axis_scale, color='blue',
linewidth=3, label="Interpolation")
plt.title('Actual vs Predicted')
plt.xlabel('years')
plt.ylabel('population growth (1e5)')
plt.legend(loc='upper left', frameon=False)
plt.savefig('linear_regression.svg', format='svg', dpi=600)
# Predict response of future data [2018-2020]
y_pred = np.round(model.predict(x_new))
print('predicted response [2018]', y_pred[4], sep='\t')
print('actual data [Dec. 2018]:\t{}'.format(df.iloc[8]['Population']))
print('predicted response [2019]', y_pred[5], sep='\t')
print('actual data [Dec. 2019]:\t{}'.format(567851))
print('predicted response [2020]', y_pred[6], sep='\t')
print('actual data [Feb. 2020]:\t{}'.format(570196))
# -
# ### Theoretical response with plateau and inflection point at 2030, 2040 and 2050
# Population growth was simulated considering a plateau in 2030, 2040 and in 2050 through the exponential function: $y = Y_M-(Y_M-Y_0)\cdot(e^{-ax})$, where $Y_M$ is the maximum at which the plateau ends up, $Y_0$ is the starting population and $a$ is a rate constant that governs how fast it gets there. Here, $Y_0 = 5.44 \cdot 1e5$ (actual population in 2018), while $Y_M$ is equals to $7.57$, $9.35$ and $11.13$ per $100000$ individuals, as inferred by linear regression respectively for 2030, 2040 and 2050.
# +
import math
axis_scale = 1e5
y_pred = model.predict(x_new)/axis_scale
fig = plt.figure(figsize=(3,3))
# Plot outputs
plt.scatter(x, y/axis_scale, color='black', label="actual", s=10)
plt.plot(x_new, y_pred, color='#984ea3', linewidth=3, label="linear")
def plateau(x, M, M0, a):
return M - (M-M0)*math.exp(-a*x)
y_pl = np.vectorize(plateau)
M_2018 = y_pred[np.where(x_new==2018)[0][0]]
M_2030 = y_pred[np.where(x_new==2030)[0][0]]
M_2040 = y_pred[np.where(x_new==2040)[0][0]]
M_2050 = y_pred[np.where(x_new==2050)[0][0]]
y_2030= y_pl(x=np.arange(0,33), M=M_2030, M0=M_2018, a=.08)
y_2040= y_pl(x=np.arange(0,33), M=M_2040, M0=M_2018, a=.045)
y_2050= y_pl(x=np.arange(0,33), M=M_2050, M0=M_2018, a=.035)
plt.plot(x_new[4:], y_2030, color='#377eb8',
linewidth=3, label="2030")
plt.plot(x_new[4:], y_2040, color='#ff7f00',
linewidth=3, label="2040")
plt.plot(x_new[4:], y_2050, color='#4daf4a',
linewidth=3, label="2050")
plt.axvline(x=2030, linewidth=.5, linestyle='dashdot', color="#377eb8", ymax=.40)
plt.axvline(x=2040, linewidth=.5, linestyle='dashdot', color="#ff7f00", ymax=.55)
plt.axvline(x=2050, linewidth=.5, linestyle='dashdot', color="#4daf4a", ymax=.72)
plt.xlabel('years')
plt.ylabel('population growth (1e5)')
plt.legend(loc='upper left', frameon=False)
plt.rcParams.update({'font.size': 11})
plt.savefig('fitted_vs_predicted.svg', format='svg', dpi=1200)
# -
# # Markov Chain design
# A **discrete-time Markov chain (DTMC)** is designed to model the *death* event of HD patients and *onset* of the disease among inhabitants of Muscat. To do that, *birth/death* events were inferred from observational data collected from 2013 to 2019 in the Muscat population.
# ## Set the transition rates
# The modeled process is stochastic and *memoryless*, in that it allows to make predictions based solely on its present state. The process passes through three states: **Healthy**, **HD Alive** and **HD Dead** as a result of *birth* and *death* events, driven in turn by: <b>I</b> = incidence rate of the disease and <b>D</b> = death rate because of the disease.<br/>
# <img src="data/JNNP_2020/MC_states.png" align="middle" width="300">
#
# where:<br/>
# * $I_{avg}$:  the average HD incidence rate in the world population (min=**0.38** per 100,000 per year, [https://doi.org/10.1002/mds.25075]; max=**0.9** per 100,000 per year, [https://doi.org/10.1186/s12883-019-1556-3])
# * $I_{Muscat}$: the actual HD incidence rate in Muscat (**0.56** per 100,000 per year [2013-2019])
# * $D_{avg}$:  the HD death rate (min=**1.55** per million population registered in England-Wales in 1960-1973, [https://pubmed.ncbi.nlm.nih.gov/6233902/]; max=**2.27** per million population registered in USA in 1971-1978, [https://doi.org/10.1212/wnl.38.5.769])
# * $D_{Muscat}$: the actual HD death rate in Muscat (**1.82** per million population [2013-2019])
# +
# estimated incidence rate world-wide
## min = 0.38/year per 100,000 [https://doi.org/10.1002/mds.25075]
## max = 0.9/year per 100,000 [https://doi.org/10.1186/s12883-019-1556-3]
est_inc_rates = [0.38, 0.56, 0.7, 0.9]
# actual incidence rate (0.56 per 100,000 per year) ~ 3 new HD patients in Muscat in 2018 [https://data.gov.om/OMPOP2016/population?indicator=1000140®ion=1000020-muscat&nationality=1000010-omani]
act_inc_rate = 0.56
# estimated death rate world-wide
## min = 1.55 per million (England-Wales in 1960-1973, https://www.ncbi.nlm.nih.gov/pubmed/6233902)
## max = 2.27 per million (United States in 1971-1978, https://doi.org/10.1212/wnl.38.5.769)
est_death_rates = [1.55, 1.819, 2.27]
# actual death rate ~ 1 patient in 2018 per 100.000 per year in Muscat
act_death_rate = 1.819
# starting HD individuals in Muscat in 2018 (32)
hd_2018 = 32
# -
# ### Set the number of simulation traces
# In order to approximate the posterior distribution of the HD prevalence, one performs a *Monte Carlo* simulation with $1000$ independent simulation runs from where calculating the *average number of incident cases* per year.
simulation_traces = BoundedIntText(
min=1,
max=5000,
step=1,
value=1000,
description='Simulation traces:', style=style)
display(simulation_traces)
# ### Initialize state vectors
# For each simulation step, $6$ vectors are updated:<br/>
# * ***est_inc*** stores the variation of incidence over time according to $I_{avg}$
# * ***act_inc*** stores the variation of incidence over time according to $I_{Muscat}$
# * ***est_death*** stores the numbers of deaths over time according to $D_{avg}$
# * ***act_death*** stores the numbers of deaths over time according to $D_{Muscat}$
# * ***est_alive_HD*** stores the number of alive HD patients over time, based on ***est_inc*** and ***est_death***
# * ***act_alive_HD*** astores the number of alive HD patients over time, based on ***act_inc*** and ***act_death***
# +
# record the number of simulation steps
sim_time = len(x_new[4:])
# get the number of simulation traces from the textfield above
num_traces = simulation_traces.value
# get number of possible Iavg and Davg
num_iavg = len(est_inc_rates)
num_davg = len(est_death_rates)
est_inc = np.zeros((num_iavg, num_davg, num_traces, sim_time), dtype=int)
act_inc = np.zeros((num_traces, sim_time), dtype=int)
est_death = np.zeros((num_iavg, num_davg, num_traces, sim_time), dtype=int)
act_death = np.zeros((num_traces, sim_time), dtype=int)
est_alive_HD = np.zeros((num_iavg, num_davg, num_traces, sim_time), dtype=int)
est_alive_HD[:, :, :, 0] = hd_2018
act_alive_HD = np.zeros((num_traces, sim_time), dtype=int)
act_alive_HD[:, 0] = hd_2018
# -
# ### Trigger the Monte Carlo simulation
# All possible estimates of $I_{avg}$ and $D_{avg}$ are shuffled here and, for each combination, 1000 independent simulations will be launched. For each simulation and for each time step, the number of alive HD patients will be calculated as the sum of the number of *currently alive* HD patients and that of *new* HD patients, from which the number of currently *deceased* patients is subtracted. This computation is performed for:
# #### Linear estimate (regression)
for iavg_idx, iavg_value in enumerate(est_inc_rates):
for davg_idx, davg_value in enumerate(est_death_rates):
print("Evaluating Iavg={} and Davg={}".format(iavg_value, davg_value))
for rep in range(num_traces):
for t in range(sim_time-1):
curr_pop = y_pred[t]*100000 # <-------
if(act_inc[rep, t] == 0):
act_inc[rep, t] = np.random.poisson((act_inc_rate * curr_pop)/100000)
act_death[rep, t] = np.random.poisson((act_death_rate * curr_pop)/1000000)
act_alive_HD[rep, t+1] = act_alive_HD[rep, t] - act_death[rep, t] + act_inc[rep, t]
est_inc[iavg_idx, davg_idx, rep, t] = np.random.poisson(
(iavg_value * curr_pop)/100000)
est_death[iavg_idx, davg_idx, rep, t] = np.random.poisson(
(davg_value * curr_pop)/1000000)
est_alive_HD[iavg_idx, davg_idx, rep, t+1] = est_alive_HD[iavg_idx, davg_idx, rep, t] - \
est_death[iavg_idx, davg_idx, rep, t] + est_inc[iavg_idx, davg_idx, rep, t]
# ##### Plot HD population predicted to be alive
# N.b., __est. HD population__ accounts for the population that is predicted to be alive every year according to *incidence* and *death* rates that are the world average; __act. HD population__ refers instead to the actual rates calculated stright on the actual Muscat population records, as previously described.
#
# A ranom plot (i.e., a random trace among 1000) will be generaed for each combination of $I_{avg}$ and $D_{avg}$.
# + code_folding=[]
# Select a random trace
rtrace = np.random.randint(0,num_traces)
# Plot a random trace of est/act. alive HD cases in the Muscat region
# for each combination of Iavg and Davg
fig, ax = plt.subplots(len(est_inc_rates), len(est_death_rates), figsize=(12, 9))
for iavg_idx, iavg_value in enumerate(est_inc_rates):
for davg_idx, davg_value in enumerate(est_death_rates):
ax[iavg_idx, davg_idx].plot(x_new[4:], est_alive_HD[iavg_idx, davg_idx, rtrace, :],
lw=2, label='est. {}, {}'.format(iavg_value, davg_value))
ax[iavg_idx, davg_idx].plot(x_new[4:], act_alive_HD[rtrace, :],
lw=2, label='act. {}, {}'.format(act_inc_rate, str(round(act_death_rate, 2))))
ax[iavg_idx, davg_idx].legend(loc='upper left', frameon=False)
plt.savefig('alive_population.svg', format='svg', dpi=1200)
# -
# #### Inflation at 2030
# +
# record the number of simulation steps
sim_time_2030 = len(y_2030)
# get the number of simulation traces from the textfield above
num_traces_2030 = simulation_traces.value
# get number of possible Iavg and Davg
num_iavg_2030 = len(est_inc_rates)
num_davg_2030 = len(est_death_rates)
est_inc_2030 = np.zeros((num_iavg_2030, num_davg_2030, num_traces_2030, sim_time_2030), dtype=int)
act_inc_2030 = np.zeros((num_traces_2030, sim_time_2030), dtype=int)
est_death_2030 = np.zeros((num_iavg_2030, num_davg_2030, num_traces_2030, sim_time_2030), dtype=int)
act_death_2030 = np.zeros((num_traces_2030, sim_time_2030), dtype=int)
est_alive_HD_2030 = np.zeros((num_iavg_2030, num_davg_2030, num_traces_2030, sim_time_2030), dtype=int)
est_alive_HD_2030[:, :, :, 0] = hd_2018
act_alive_HD_2030 = np.zeros((num_traces_2030, sim_time_2030), dtype=int)
act_alive_HD_2030[:, 0] = hd_2018
# -
for iavg_idx, iavg_value in enumerate(est_inc_rates):
for davg_idx, davg_value in enumerate(est_death_rates):
print("Evaluating Iavg={} and Davg={}".format(iavg_value, davg_value))
for rep in range(num_traces_2030):
for t in range(sim_time_2030-1):
curr_pop = y_2030[t]*100000 # <-------
if(act_inc_2030[rep, t] == 0):
act_inc_2030[rep, t] = np.random.poisson((act_inc_rate * curr_pop)/100000)
act_death_2030[rep, t] = np.random.poisson((act_death_rate * curr_pop)/1000000)
act_alive_HD_2030[rep, t+1] = act_alive_HD_2030[rep, t] - act_death_2030[rep, t] + act_inc_2030[rep, t]
est_inc_2030[iavg_idx, davg_idx, rep, t] = np.random.poisson(
(iavg_value * curr_pop)/100000)
est_death_2030[iavg_idx, davg_idx, rep, t] = np.random.poisson(
(davg_value * curr_pop)/1000000)
est_alive_HD_2030[iavg_idx, davg_idx, rep, t+1] = est_alive_HD_2030[iavg_idx, davg_idx, rep, t] - \
est_death_2030[iavg_idx, davg_idx, rep, t] + est_inc_2030[iavg_idx, davg_idx, rep, t]
# ##### Plot HD population predicted to be alive
# +
# Select a random trace
rtrace = np.random.randint(0,num_traces_2030)
# Plot a random trace of est/act. alive HD cases in the Muscat region
# for each combination of Iavg and Davg
fig, ax = plt.subplots(len(est_inc_rates), len(est_death_rates), figsize=(12, 9))
for iavg_idx, iavg_value in enumerate(est_inc_rates):
for davg_idx, davg_value in enumerate(est_death_rates):
ax[iavg_idx, davg_idx].plot(x_new[4:], est_alive_HD_2030[iavg_idx, davg_idx, rtrace, :],
lw=2, label='est. {}, {}'.format(iavg_value, davg_value))
ax[iavg_idx, davg_idx].plot(x_new[4:], act_alive_HD_2030[rtrace, :],
lw=2, label='act. {}, {}'.format(act_inc_rate, str(round(act_death_rate, 2))))
ax[iavg_idx, davg_idx].legend(loc='upper left', frameon=False)
plt.savefig('alive_population_y2030.svg', format='svg', dpi=1200)
# -
# #### Inflation at 2040
# +
# record the number of simulation steps
sim_time_2040 = len(y_2040)
# get the number of simulation traces from the textfield above
num_traces_2040 = simulation_traces.value
# get number of possible Iavg and Davg
num_iavg_2040 = len(est_inc_rates)
num_davg_2040 = len(est_death_rates)
est_inc_2040 = np.zeros((num_iavg_2040, num_davg_2040, num_traces_2040, sim_time_2040), dtype=int)
act_inc_2040 = np.zeros((num_traces_2040, sim_time_2040), dtype=int)
est_death_2040 = np.zeros((num_iavg_2040, num_davg_2040, num_traces_2040, sim_time_2040), dtype=int)
act_death_2040 = np.zeros((num_traces_2040, sim_time_2040), dtype=int)
est_alive_HD_2040 = np.zeros((num_iavg_2040, num_davg_2040, num_traces_2040, sim_time_2040), dtype=int)
est_alive_HD_2040[:, :, :, 0] = hd_2018
act_alive_HD_2040 = np.zeros((num_traces_2040, sim_time_2040), dtype=int)
act_alive_HD_2040[:, 0] = hd_2018
# -
for iavg_idx, iavg_value in enumerate(est_inc_rates):
for davg_idx, davg_value in enumerate(est_death_rates):
print("Evaluating Iavg={} and Davg={}".format(iavg_value, davg_value))
for rep in range(num_traces_2040):
for t in range(sim_time_2040-1):
curr_pop = y_2040[t]*100000 # <-------
if(act_inc_2040[rep, t] == 0):
act_inc_2040[rep, t] = np.random.poisson((act_inc_rate * curr_pop)/100000)
act_death_2040[rep, t] = np.random.poisson((act_death_rate * curr_pop)/1000000)
act_alive_HD_2040[rep, t+1] = act_alive_HD_2040[rep, t] - act_death_2040[rep, t] + act_inc_2040[rep, t]
est_inc_2040[iavg_idx, davg_idx, rep, t] = np.random.poisson(
(iavg_value * curr_pop)/100000)
est_death_2040[iavg_idx, davg_idx, rep, t] = np.random.poisson(
(davg_value * curr_pop)/1000000)
est_alive_HD_2040[iavg_idx, davg_idx, rep, t+1] = est_alive_HD_2040[iavg_idx, davg_idx, rep, t] - \
est_death_2040[iavg_idx, davg_idx, rep, t] + est_inc_2040[iavg_idx, davg_idx, rep, t]
# ##### Plot HD population predicted to be alive
# +
# Select a random trace
rtrace = np.random.randint(0,num_traces_2040)
# Plot a random trace of est/act. alive HD cases in the Muscat region
# for each combination of Iavg and Davg
fig, ax = plt.subplots(len(est_inc_rates), len(est_death_rates), figsize=(12, 9))
for iavg_idx, iavg_value in enumerate(est_inc_rates):
for davg_idx, davg_value in enumerate(est_death_rates):
ax[iavg_idx, davg_idx].plot(x_new[4:], est_alive_HD_2040[iavg_idx, davg_idx, rtrace, :],
lw=2, label='est. {}, {}'.format(iavg_value, davg_value))
ax[iavg_idx, davg_idx].plot(x_new[4:], act_alive_HD_2040[rtrace, :],
lw=2, label='act. {}, {}'.format(act_inc_rate, str(round(act_death_rate, 2))))
ax[iavg_idx, davg_idx].legend(loc='upper left', frameon=False)
plt.savefig('alive_population_y2040.svg', format='svg', dpi=1200)
# -
# #### Inflation at 2050
# +
# record the number of simulation steps
sim_time_2050 = len(y_2050)
# get the number of simulation traces from the textfield above
num_traces_2050 = simulation_traces.value
# get number of possible Iavg and Davg
num_iavg_2050 = len(est_inc_rates)
num_davg_2050 = len(est_death_rates)
est_inc_2050 = np.zeros((num_iavg_2050, num_davg_2050, num_traces_2050, sim_time_2050), dtype=int)
act_inc_2050 = np.zeros((num_traces_2050, sim_time_2050), dtype=int)
est_death_2050 = np.zeros((num_iavg_2050, num_davg_2050, num_traces_2050, sim_time_2050), dtype=int)
act_death_2050 = np.zeros((num_traces_2050, sim_time_2050), dtype=int)
est_alive_HD_2050 = np.zeros((num_iavg_2050, num_davg_2050, num_traces_2050, sim_time_2050), dtype=int)
est_alive_HD_2050[:, :, :, 0] = hd_2018
act_alive_HD_2050 = np.zeros((num_traces_2050, sim_time_2050), dtype=int)
act_alive_HD_2050[:, 0] = hd_2018
# -
for iavg_idx, iavg_value in enumerate(est_inc_rates):
for davg_idx, davg_value in enumerate(est_death_rates):
print("Evaluating Iavg={} and Davg={}".format(iavg_value, davg_value))
for rep in range(num_traces_2050):
for t in range(sim_time_2050-1):
curr_pop = y_2050[t]*100000 # <-------
if(act_inc_2050[rep, t] == 0):
act_inc_2050[rep, t] = np.random.poisson((act_inc_rate * curr_pop)/100000)
act_death_2050[rep, t] = np.random.poisson((act_death_rate * curr_pop)/1000000)
act_alive_HD_2050[rep, t+1] = act_alive_HD_2050[rep, t] - act_death_2050[rep, t] + act_inc_2050[rep, t]
est_inc_2050[iavg_idx, davg_idx, rep, t] = np.random.poisson(
(iavg_value * curr_pop)/100000)
est_death_2050[iavg_idx, davg_idx, rep, t] = np.random.poisson(
(davg_value * curr_pop)/1000000)
est_alive_HD_2050[iavg_idx, davg_idx, rep, t+1] = est_alive_HD_2050[iavg_idx, davg_idx, rep, t] - \
est_death_2050[iavg_idx, davg_idx, rep, t] + est_inc_2050[iavg_idx, davg_idx, rep, t]
# ##### Plot HD population predicted to be alive
# +
# Select a random trace
rtrace = np.random.randint(0,num_traces_2050)
# Plot a random trace of est/act. alive HD cases in the Muscat region
# for each combination of Iavg and Davg
fig, ax = plt.subplots(len(est_inc_rates), len(est_death_rates), figsize=(12, 9))
for iavg_idx, iavg_value in enumerate(est_inc_rates):
for davg_idx, davg_value in enumerate(est_death_rates):
ax[iavg_idx, davg_idx].plot(x_new[4:], est_alive_HD_2050[iavg_idx, davg_idx, rtrace, :],
lw=2, label='est. {}, {}'.format(iavg_value, davg_value))
ax[iavg_idx, davg_idx].plot(x_new[4:], act_alive_HD_2050[rtrace, :],
lw=2, label='act. {}, {}'.format(act_inc_rate, str(round(act_death_rate, 2))))
ax[iavg_idx, davg_idx].legend(loc='upper left', frameon=False)
plt.savefig('alive_population_y2050.svg', format='svg', dpi=1200)
# -
# # Calculate average estimates of prevalence over traces
# The **prevalence** of HD is calculated year by year until the end of simulation and for each simulation trace.
# +
# act_alive_HD_mu = np.around(np.mean(act_alive_HD, axis=0) / y_pred, 2)
# act_alive_HD_sigma = np.around(np.std(act_alive_HD, axis=0) / y_pred, 2)
# act_alive_HD_2030_mu = np.around(np.mean(act_alive_HD_2030 / y_2030, axis=0), 2)
# act_alive_HD_2030_sigma = np.around(np.std(act_alive_HD_2030 / y_2030, axis=0), 2)
# act_alive_HD_2040_mu = np.around(np.mean(act_alive_HD_2040 / y_2040, axis=0), 2)
# act_alive_HD_2040_sigma = np.around(np.std(act_alive_HD_2040 / y_2040, axis=0), 2)
# act_alive_HD_2050_mu = np.around(np.mean(act_alive_HD_2050 / y_2050, axis=0), 2)
# act_alive_HD_2050_sigma = np.around(np.std(act_alive_HD_2050 / y_2050, axis=0), 2)
# Iavg = [0.38, 0.56, 0.7, 0.9]
# Davg = [1.55, 1.819, 2.27]
# Select simulations with Iavg=0.56 (index=1) and Davg=2.27 (index=2) estimates
est_alive_HD_mu = np.around(np.mean(est_alive_HD[1,2,:,:], axis=0) / y_pred[4:], 2)
est_alive_HD_sigma = np.around(np.std(est_alive_HD[1,2,:,:], axis=0) / y_pred[4:], 2)
est_alive_HD_2030_mu = np.around(np.mean(est_alive_HD_2030[1,2,:,:]/ y_2030, axis=0), 2)
est_alive_HD_2030_sigma = np.around(np.std(est_alive_HD_2030[1,2,:,:]/ y_2030, axis=0), 2)
est_alive_HD_2040_mu = np.around(np.mean(est_alive_HD_2040[1,2,:,:]/ y_2040, axis=0), 2)
est_alive_HD_2040_sigma = np.around(np.std(est_alive_HD_2040[1,2,:,:]/ y_2040, axis=0), 2)
est_alive_HD_2050_mu = np.around(np.mean(est_alive_HD_2050[1,2,:,:]/ y_2050, axis=0), 2)
est_alive_HD_2050_sigma = np.around(np.std(est_alive_HD_2050[1,2,:,:]/ y_2050, axis=0), 2)
print("Linear regression (est.): {}".format(est_alive_HD_mu))
# print("Linear regression (act.): {}\n".format(act_alive_HD_mu))
print("flection at 2030 (est.): {}".format(est_alive_HD_2030_mu))
# print("flection at 2030 (act.): {}\n".format(act_alive_HD_2030_mu))
print("flection at 2040 (est.): {}".format(est_alive_HD_2040_mu))
# print("flection at 2040 (act.): {}\n".format(act_alive_HD_2040_mu))
print("flection at 2050 (est.): {}".format(est_alive_HD_2050_mu))
# print("flection at 2050 (act.): {}".format(act_alive_HD_2050_mu))
# -
# ### Plot prevalence estimates
# Calculate the **average prevalence** values ($\mu$) for each year and over all simulation traces, together with the **standard deviation** ($\sigma$) values.<br/>
# Make a line plot with bands, with *years* on the X-axis and the *prevalence* values on the Y-axis.
# + code_folding=[0, 40]
def plot_double_prevalence(splot_idx: int, ax, x_vector: list, y1_vector_mu: list, y1_vector_sigma:
list, y2_vector_mu: list, y2_vector_sigma: list,
enable_y_axis_label: bool = False):
markers_on = [len(y1_vector_mu)-1]
ax[splot_idx].fill_between(x_vector.flatten(), y1_vector_mu+y1_vector_sigma, y1_vector_mu-y1_vector_sigma,
facecolor='#377eb8', alpha=0.1)
ax[splot_idx].fill_between(x_vector.flatten(), y2_vector_mu+y2_vector_sigma, y2_vector_mu-y2_vector_sigma,
facecolor='#ff7f00', alpha=0.1)
ax[splot_idx].plot(x_vector, y1_vector_mu, '-gD', lw=2,
label=r'$I_{avg}=0.56$, $D_{avg}=1.82$', markevery=markers_on, color='#377eb8')
ax[splot_idx].set_xticks([2014, 2015, 2025, 2035, 2045], minor=True)
ax[splot_idx].plot(x_vector, y2_vector_mu, '-gD', lw=2,
label=r'$I_{avg}=0.56$, $D_{avg}=2.27$', markevery=markers_on, color="#ff7f00")
if enable_y_axis_label:
ax[splot_idx].set_ylabel('Est. prevalence')
ax[splot_idx].set_xticks([2014, 2015, 2025, 2035, 2045], minor=True)
ax[splot_idx].spines['top'].set_visible(False)
ax[splot_idx].spines['right'].set_visible(False)
ax[splot_idx].spines['bottom'].set_visible(True)
ax[splot_idx].spines['left'].set_visible(True)
ax[splot_idx].annotate(y1_vector_mu[-1],
(2050, y1_vector_mu[-1]),
textcoords="offset points",
xytext=(-30, 10),
ha='left')
ax[splot_idx].annotate(y2_vector_mu[-1],
(2050, y2_vector_mu[-1]),
textcoords="offset points",
xytext=(-30, -20),
ha='left')
def plot_prevalence(ax, x_vector: list, y1_vector_mu: list, y1_vector_sigma: list,
y2_vector_mu: list, y2_vector_sigma: list,
y3_vector_mu: list, y3_vector_sigma: list,
y4_vector_mu: list, y4_vector_sigma: list):
markers_on = [len(y1_vector_mu)-1]
ax.fill_between(x_vector.flatten(), y1_vector_mu+y1_vector_sigma,
y1_vector_mu-y1_vector_sigma, facecolor='#377eb8', alpha=0.1)
ax.plot(x_vector, y1_vector_mu, '-gD', lw=2, label=r'2030',
markevery=markers_on, color="#377eb8")
ax.fill_between(x_vector.flatten(), y2_vector_mu+y2_vector_sigma,
y2_vector_mu-y2_vector_sigma, facecolor='#ff7f00', alpha=0.1)
ax.plot(x_vector, y2_vector_mu, '-gD', lw=2, label=r'2040',
markevery=markers_on, color="#ff7f00")
ax.fill_between(x_vector.flatten(), y3_vector_mu+y3_vector_sigma,
y3_vector_mu-y3_vector_sigma, facecolor='#4daf4a', alpha=0.1)
ax.plot(x_vector, y3_vector_mu, '-gD', lw=2, label=r'2050',
markevery=markers_on, color="#4daf4a")
ax.fill_between(x_vector.flatten(), y4_vector_mu+y4_vector_sigma,
y4_vector_mu-y4_vector_sigma, facecolor='#984ea3', alpha=0.1)
ax.plot(x_vector, y4_vector_mu, '-gD', lw=2, label=r'linear',
markevery=markers_on, color="#984ea3")
ax.set_ylabel('Est. prevalence')
ax.set_xticks([2014, 2015, 2025, 2035, 2045], minor=True)
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
ax.spines['bottom'].set_visible(True)
ax.spines['left'].set_visible(True)
ax.annotate(y1_vector_mu[-1],
(2050, y1_vector_mu[-1]),
textcoords="offset points",
xytext=(10, 0),
ha='left')
ax.annotate(y2_vector_mu[-1],
(2050, y2_vector_mu[-1]),
textcoords="offset points",
xytext=(10, 0),
ha='left')
ax.annotate(y3_vector_mu[-1],
(2050, y3_vector_mu[-1]),
textcoords="offset points",
xytext=(10, -3),
ha='left')
ax.annotate(y4_vector_mu[-1],
(2050, y4_vector_mu[-1]),
textcoords="offset points",
xytext=(10, -5),
ha='left')
fig, ax = plt.subplots(1, 1, figsize=(2.5, 3))
plot_prevalence(ax, x_new[4:], est_alive_HD_2030_mu, est_alive_HD_2030_sigma, est_alive_HD_2040_mu, est_alive_HD_2040_sigma,
est_alive_HD_2050_mu, est_alive_HD_2050_sigma, est_alive_HD_mu, est_alive_HD_sigma)
lines, labels = fig.axes[-1].get_legend_handles_labels()
fig.legend(lines, labels, bbox_to_anchor=(
0.2, 0.9), loc='upper left', frameon=False)
fig.savefig('prevalence_est_I056_D227.svg', format='svg', dpi=1200)
# -
# ### Generate prevalence tables
prev_df = pd.DataFrame({'Years':x_new[4:].flatten(),
'Linear (avg)':est_alive_HD_mu, 'Linear (std)':est_alive_HD_sigma,
'2030 (avg)':est_alive_HD_2030_mu, '2030 (std)':est_alive_HD_2030_sigma,
'2040 (avg)':est_alive_HD_2040_mu, '2040 (std)':est_alive_HD_2040_sigma,
'2050 (avg)':est_alive_HD_2050_mu, '2050 (std)':est_alive_HD_2050_sigma
})
prev_df.to_excel("prevalence_estimates.xlsx", index=False)
print(prev_df.head())
# ### Miscellaneous plots
# Create two plots of **frequency** of *adults*, *youngs* HD and *at-risk* subjects.
# +
bars_file = "./data/JNNP_2020/Bar_plots.xlsx"
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(6, 2.5), gridspec_kw={'width_ratios': [2, 1]})
######## Panel A ########
dfA = pd.read_excel(bars_file, sheet_name="Figure A") #index_col=0,
print(dfA)
print("")
labels = dfA.Patients
x = np.arange(len(labels))
adult_onset = dfA.iloc[:,1]
juvenile_onset = dfA.iloc[:,2]
width = 0.35*2 # the width of the bars
rects1 = ax1.bar(x, adult_onset, width, label='Adult onset',
color='white', edgecolor='black')
rects2 = ax1.bar(x, juvenile_onset, width, label='Juvenile onset',
color='lightgray', edgecolor='black', hatch="//////")
ax1.set_ylabel('HD subjects')
ax1.set_xticks(x)
ax1.set_yticks([10, 30, 50], minor=True)
ax1.set_xticklabels(labels)
ax1.legend(loc='upper center', bbox_to_anchor=(0.5, 1.25),ncol=2, prop={"size":8}, frameon=False)
ax1.spines['top'].set_visible(False)
ax1.spines['right'].set_visible(False)
ax1.spines['bottom'].set_visible(True)
ax1.spines['left'].set_visible(True)
######## Panel B ########
dfB = pd.read_excel(bars_file, sheet_name="Figure B")
print(dfB)
print("")
labels = dfB.Patients
more50 = dfB.iloc[:,1]
less50 = dfB.iloc[:,2]
x = np.arange(len(labels))
width = 0.35*2
rects1 = ax2.bar(x, more50, width, label='>50% risk',
color='white', edgecolor='black')
rects2 = ax2.bar(x, less50, width, label='≤50% risk',
color='lightgray', edgecolor='black', hatch="xXX")
ax2.set_ylabel('At-risk subjects')
ax2.set_xticks(x)
ax2.set_xticklabels(labels)
ax2.legend(loc='upper center', bbox_to_anchor=(0.5, 1.25),ncol=2, prop={"size":8}, frameon=False)
ax2.spines['top'].set_visible(False)
ax2.spines['right'].set_visible(False)
ax2.spines['bottom'].set_visible(True)
ax2.spines['left'].set_visible(True)
##### Plotting #####
fig.tight_layout()
plt.rcParams.update({'font.size': 10})
plt.show()
def on_save_misc_button(but):
fig.savefig('misc_plots.svg', format='svg', dpi=1200)
print('Figure saved')
save_misc_button = Button(
description="Save SVG",
button_style='info',
tooltip='Save to SVG file'
)
save_misc_button.on_click(on_save_misc_button)
display(save_misc_button)
# -
# # Print system and required packages information
# +
# %load_ext watermark
# %watermark -v -m -p numpy,pandas,matplotlib,sklearn,traitlets,IPython,ipywidgets
# date
print(" ")
# %watermark -u -n -t -z
|
HD_prevalence_JNNP_2020.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
my_list_1 = [1,2,3,4,"a","a"]
my_list_1.append("a")
my_list_1
a = [1,2]
b = [3,4]
a.append(b)
a
# append() içine seçtiğimiz herhangi bir şeyi eklememize yarar ayrıca iki listeyi birbirine de eklememizi sağlar
my_list_1.clear()
my_list_1
# clear() listenin içini siler
my_list_2 = [1,2,3,"a","a"]
x = my_list_2.copy()
x
# copy() başka bi değişkene kopyalamamızı sağlar
my_list_3 = [1,2,3,"a","a"]
my_list_3.count("a")
# count() içine girdiğimiz ifadeden kaç tane var ise onun sayısını verir
my_list_4 = ["bmv" , "apple"]
my_list_4.extend(my_list_3)
my_list_4
# yukarıdaki gibi my_list_4 ün içine my_list_3 ü ekledi
my_list_5 = [1,2,34,"a"]
my_list_5.index(34)
my_list_5.index("a")
# +
## yukarıdaki gibi index() komutu içine yazdığımız ifadenin indexini veriyor bize
# -
|
05-ListMethods.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from __future__ import division, absolute_import
import sys
import random
import pickle
from sklearn import metrics
import pandas as pd
import numpy as np
import h5py
from plotnine import *
#root
absPath = '/home/angela3/imbalance_pcm_benchmark/'
sys.path.insert(0, absPath)
from src.Target import Target
from src.imbalance_functions import *
np.random.seed(8)
random.seed(8)
# +
protein_type = "GPCRs" #"kinases"
nfolds = 10
group = '/activity'
table = "prot_comp"
#Loading maximum lengths of proteins and compounds
with open("".join((absPath, 'data/prot_max_len.pickle')), "rb") as input_file:
max_len_prot = pickle.load(input_file)
#Defining protein dictionary
instarget = Target("AAA")
prot_dict = instarget.predefining_dict()
with open("".join((absPath, "data/", protein_type, "/", protein_type, "_prots.pickle")), 'rb') as handle:
unique_prots = pickle.load(handle)
# -
protein_info = "".join((absPath, "data/", protein_type, "/prots_df.csv"))
prot_df = pd.read_csv(protein_info)
prot_df.drop(["Unnamed: 0"],axis=1, inplace=True)
prot_df.head()
prot_df["len_seq"] = prot_df.apply(len_seq, axis=1)
ratios_per_fold = []
for fold in range(nfolds):
f_train = h5py.File("".join((absPath, "data/", protein_type, "/resampling_after_clustering/",
str(fold), "/compounds_activity_training.h5")), "r")
f_test = h5py.File("".join((absPath, "data/", protein_type, "/resampling_after_clustering/",
str(fold), "/compounds_activity_test.h5")), "r")
sample_indices_train = range(len(f_train[group][table]))
sample_indices_test = range(len(f_test[group][table]))
training_ratios = creating_ratios_list("training", fold, f_train, group, table, sample_indices_train,
unique_prots)
test_ratios = creating_ratios_list("test", fold, f_test, group, table, sample_indices_test,
unique_prots)
pred_test_path = "".join((absPath, "data/", protein_type, "/resampling_after_clustering/predictions/",
str(fold), "/test.csv"))
pred_test = pd.read_csv(pred_test_path)
pred_test_ratios, pred_test_ = predictions_ratios_list(fold, pred_test, unique_prots)
list_metrics = []
for prot in unique_prots:
dict_prot = computing_metrics_per_prot(prot, pred_test_)
list_metrics.append(dict_prot)
filt_list_metrics = []
for val in list_metrics:
if val != None:
filt_list_metrics.append(val)
ratios_df = converting_ratios_to_df(training_ratios, test_ratios, pred_test_ratios, filt_list_metrics,
"resampling_after_clustering", prot_df)
ratios_df["fold"] = str(fold)
ratios_per_fold.append(ratios_df)
ratios_complete_df = pd.concat(ratios_per_fold)
ratios_complete_df.info()
ratios_seq = pd.merge(ratios_complete_df, prot_df, "left", on="DeepAffinity Protein ID")
print(ratios_seq.info())
print(ratios_seq.head())
ratios_seq.to_csv("".join((absPath, "data/", protein_type, "/resampling_after_clustering/results/ratios_df.csv")))
|
scripts/resampling_after_clustering/02_computing_ratios.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Color Wheel
#
# For this example we'll be making a simple color picker that's powered by python.
import math
import purly
# To set things up we'll need to stand up our model server. By default the server has a max refresh rate of `25` hertz. However for this example to be convincingly smooth we'll want to bump that up to about `60`. If you want to unlock the refresh rate set `refresh=None`. We'll then hook up a layout object to a model resource (see the introductory example or [read the docs](https://github.com/rmorshea/purly#purly) if this doesn't make sense).
# +
from example_utils import localhost
# increase model server refresh cap to 60 hertz.
purly.state.Machine(refresh=60).run()
# name the layout resource "color-wheel" and connect to the update stream
websocket_url = localhost('ws', 8000) + '/model/color-wheel/stream'
layout = purly.Layout(websocket_url)
# -
# To do this we'll need a simple function that will create an HSL (hue, saturation, and lightness) color string. The function will accept the radius of the color picker wheel, and the x-y position of the mouse in order to select a color based on the angle around the circle, and distance from the center.
def hsl(radius, x, y):
"""Return an HSL color string."""
x -= radius
y -= radius
unit_radius = int((x ** 2 + y **2) ** 0.5) / radius
degrees = int(math.atan2(x, y) * 180 / math.pi)
return "hsl(%s, 100%%, %s%%)" % (degrees, unit_radius * 100)
# Next we'll need style up the the color wheel, and a selection indicator.
radius = 50
wheel = layout.html('div')
wheel.style.update(
height="%spx" % (radius * 2),
width="%spx" % (radius * 2),
backgroundColor="hsl(120, 100%, 50%)",
borderRadius="50%",
)
selection = layout.html('div')
selection.style.update(
height="20px",
width="20px",
backgroundColor="hsl(120, 100%, 50%)",
)
layout.children.append(wheel)
layout.children.append(selection)
layout.sync()
layout
# You should now see the color wheel and selector indicator in the output above!
#
# However when you mouse over the wheel nothing happens, so we'll need to hook into some mouse events to make the display animate. To do that requires the `onMouseMove` and `onClick` events which can be captures via the `on` decorator of the `wheel` element. The underlying logic of each is actually pretty simple:
#
# 1. `onMouseMove`: if the mouse moves over the color wheel, then set the color wheel to the HSL color that corresponds to its x-y position.
# 2. `onClick`: if the mouse clicks on the wheel, set the selection indicator to the corresponding HSL color.
# +
@wheel.on('MouseMove')
def cause_color_change(offsetX, offsetY):
wheel.style["backgroundColor"] = hsl(50, offsetX, offsetY)
@wheel.on("Click")
def cause_color_select(offsetX, offsetY):
selection.style["backgroundColor"] = hsl(50, offsetX, offsetY)
# -
# Finally we'll need to serve up our event handlers in order to animate.
layout.serve()
# Now mouse over the wheel and try selecting a color!
|
examples/colors.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Loading and Displaying Well Log Data from LAS
# **Created by:** <NAME>
#
# This notebook illustrates how to load data in from a LAS file and carry out a basic QC of the data before plotting it on a log plot.
# ## Loading and Checking Data
# The first step is to import the required libraries: pandas, matplotlib and LASIO.
# lasio is a library that has been developed to handle and work with LAS files. More info on the library can be found at: https://lasio.readthedocs.io/en/latest/
import os
import pandas as pd
import matplotlib.pyplot as plt
import lasio
# To load our file in, we can use the read() method from LASIO like so:
root = '/users/kai/desktop/data_science/data/dongara'
well_name = 'dongara_24'
file_format = '.las'
las = lasio.read(os.path.join(root,well_name+file_format))
# Now that our file has been loaded, we can start investigating it's contents.
# To find information out about where the file originated from, such as the well name, location and what the depth range of the file covers, we can create a simple for loop to go over each header item. Using Python's f-string we can join the items together.
for item in las.well:
print(f"{item.descr} ({item.mnemonic}): {item.value}")
# If we just want to extract the Well Name, we can simply call it by:
las.well.WELL.value
# To quickly see what curves are present within the las file we can loop through `las.curves`
for curve in las.curves:
print(curve.mnemonic)
# To see what curves are present within the las file, we can repeat the process with the CurveItem object and call upon the `unit` and `descr` functions to get info on the units and the curve's description.
# The enumerate function allows us to keep a count of the number of curves that are present within the file. As enumerate returns a 0 on the first loop, we need to 1 to it if we want to include the depth curve.
for count, curve in enumerate(las.curves):
print(f"Curve: {curve.mnemonic}, Units: {curve.unit}, Description: {curve.descr}")
print(f"There are a total of: {count+1} curves present within this file")
# ## Creating a Pandas Dataframe
# Data loaded in using LASIO can be converted to a pandas dataframe using the .df() function. This allows us to easily plot data and pass it into one of the many machine learning algorithms.
well = las.df()
# The `.head()` function generates a table view of the header and the first 5 rows within the dataframe.
well.head(10000)
# To find out more information about data, we can call upon the `.info()` and `.describe()` functions.
#
# The `.info()` function provides information about the data types and how many non-null values are present within each curve.
# The `.describe()` function, provides statistical information about each curve and can be a useful QC for each curve.
well.describe()
well.info()
# ## Visualising Data Extent
# Instead of the summary provided by the pandas describe() function, we can create a visualisation using matplotlib. Firstly, we need to work out where we have nulls (nan values). We can do this by creating a second dataframe and calling .notnull() on our well dataframe.
#
# As this returns a boolean (True or False) for each depth, we need to multiply by 1 to convert the values from True and False to 1 and 0 respectively.
well_nan = well.notnull() * 1
well_nan.head()
# We can now create a summary plot of the missing data
# +
fig = plt.subplots(figsize=(7,10))
#Set up the plot axes
ax1 = plt.subplot2grid((1,7), (0,0), rowspan=1, colspan = 1)
ax2 = plt.subplot2grid((1,7), (0,1), rowspan=1, colspan = 1)
ax3 = plt.subplot2grid((1,7), (0,2), rowspan=1, colspan = 1)
ax4 = plt.subplot2grid((1,7), (0,3), rowspan=1, colspan = 1)
ax5 = plt.subplot2grid((1,7), (0,4), rowspan=1, colspan = 1)
ax6 = plt.subplot2grid((1,7), (0,5), rowspan=1, colspan = 1)
ax7 = plt.subplot2grid((1,7), (0,6), rowspan=1, colspan = 1)
columns = well_nan.columns
axes = [ax1, ax2, ax3, ax4, ax5, ax6, ax7]
for i, ax in enumerate(axes):
ax.plot(well_nan.iloc[:,i], well_nan.index, lw=0)
ax.set_ylim(5000, 0)
ax.set_xlim(0, 1)
ax.set_title(columns[i])
ax.set_facecolor('whitesmoke')
ax.fill_betweenx(well_nan.index, 0, well_nan.iloc[:,i], facecolor='red')
# Remove tick labels from each subplot
if i > 0:
plt.setp(ax.get_yticklabels(), visible = False)
plt.setp(ax.get_xticklabels(), visible = False)
ax1.set_ylabel('Depth', fontsize=14)
plt.subplots_adjust(wspace=0)
plt.show()
# -
# ## Plotting Log Data
# Finally, we can plot our data using the code below. Essentially, the code is building up a series of subplots and plotting the data on the relevant tracks.
#
# When we add curves to the tracks, we need to set the curve's properties, including the limits, colour and labels. We can also specify the shading between curves. An example has been added to the caliper curve to show shading between a bitsize value (8.5") and the CALI curve.
#
# If there are a number of features that are common between the plots, we can iterate over them using a for loop.
# +
fig, ax = plt.subplots(figsize=(15,10))
#Set up the plot axes
ax1 = plt.subplot2grid((1,6), (0,0), rowspan=1, colspan = 1)
ax2 = plt.subplot2grid((1,6), (0,1), rowspan=1, colspan = 1, sharey = ax1)
ax3 = plt.subplot2grid((1,6), (0,2), rowspan=1, colspan = 1, sharey = ax1)
ax4 = plt.subplot2grid((1,6), (0,3), rowspan=1, colspan = 1, sharey = ax1)
ax5 = ax3.twiny() #Twins the y-axis for the density track with the neutron track
ax6 = plt.subplot2grid((1,6), (0,4), rowspan=1, colspan = 1, sharey = ax1)
ax7 = ax2.twiny()
# As our curve scales will be detached from the top of the track,
# this code adds the top border back in without dealing with splines
ax10 = ax1.twiny()
ax10.xaxis.set_visible(False)
ax11 = ax2.twiny()
ax11.xaxis.set_visible(False)
ax12 = ax3.twiny()
ax12.xaxis.set_visible(False)
ax13 = ax4.twiny()
ax13.xaxis.set_visible(False)
ax14 = ax6.twiny()
ax14.xaxis.set_visible(False)
# Gamma Ray track
ax1.plot(well["GR"], well.index, color = "green", linewidth = 0.5)
ax1.set_xlabel("Gamma")
ax1.xaxis.label.set_color("green")
ax1.set_xlim(0, 200)
ax1.set_ylabel("Depth (m)")
ax1.tick_params(axis='x', colors="green")
ax1.spines["top"].set_edgecolor("green")
ax1.title.set_color('green')
ax1.set_xticks([0, 50, 100, 150, 200])
# Resistivity track
# ax2.plot(well["RDEP"], well.index, color = "red", linewidth = 0.5)
# ax2.set_xlabel("Resistivity - Deep")
# ax2.set_xlim(0.2, 2000)
# ax2.xaxis.label.set_color("red")
# ax2.tick_params(axis='x', colors="red")
# ax2.spines["top"].set_edgecolor("red")
# ax2.set_xticks([0.1, 1, 10, 100, 1000])
# ax2.semilogx()
# Resistivity track
ax2.plot(well["LLD"], well.index, color = "red", linewidth = 0.5)
ax2.set_xlabel("SHALLOW RESISTIVITY")
ax2.set_xlim(0.2, 2000)
ax2.xaxis.label.set_color("red")
ax2.tick_params(axis='x', colors="red")
ax2.spines["top"].set_edgecolor("red")
ax2.set_xticks([0.1, 1, 10, 100, 1000])
ax2.semilogx()
# Density track
ax3.plot(well["RHOB"], well.index, color = "red", linewidth = 0.5)
ax3.set_xlabel("Bulk Density")
ax3.set_xlim(1.95, 2.95)
ax3.xaxis.label.set_color("red")
ax3.tick_params(axis='x', colors="red")
ax3.spines["top"].set_edgecolor("red")
ax3.set_xticks([1.95, 2.45, 2.95])
# Sonic track
ax4.plot(well["DTLN"], well.index, color = "purple", linewidth = 0.5)
ax4.set_xlabel("Delta-T Long Spacing Near")
ax4.set_xlim(140, 40)
ax4.xaxis.label.set_color("purple")
ax4.tick_params(axis='x', colors="purple")
ax4.spines["top"].set_edgecolor("purple")
# Neutron track placed ontop of density track
ax5.plot(well["NRHO"], well.index, color = "blue", linewidth = 0.5)
ax5.set_xlabel('Neutron')
ax5.xaxis.label.set_color("blue")
ax5.set_xlim(45, -15)
ax5.set_ylim(4150, 3500)
ax5.tick_params(axis='x', colors="blue")
ax5.spines["top"].set_position(("axes", 1.08))
ax5.spines["top"].set_visible(True)
ax5.spines["top"].set_edgecolor("blue")
ax5.set_xticks([45, 15, -15])
# Caliper track
ax6.plot(well["CALI"], well.index, color = "black", linewidth = 0.5)
ax6.set_xlabel("Caliper")
ax6.set_xlim(6, 16)
ax6.xaxis.label.set_color("black")
ax6.tick_params(axis='x', colors="black")
ax6.spines["top"].set_edgecolor("black")
ax6.fill_betweenx(well_nan.index, 8.5, well["CALI"], facecolor='yellow')
ax6.set_xticks([6, 11, 16])
# Resistivity track - Curve 2
ax7.plot(well["LLS"], well.index, color = "green", linewidth = 0.5)
ax7.set_xlabel("SHALLOW RESISTIVITY")
ax7.set_xlim(0.2, 2000)
ax7.xaxis.label.set_color("green")
ax7.spines["top"].set_position(("axes", 1.08))
ax7.spines["top"].set_visible(True)
ax7.tick_params(axis='x', colors="green")
ax7.spines["top"].set_edgecolor("green")
ax7.set_xticks([0.1, 1, 10, 100, 1000])
ax7.semilogx()
# Common functions for setting up the plot can be extracted into
# a for loop. This saves repeating code.
for ax in [ax1, ax2, ax3, ax4, ax6]:
ax.set_ylim(4500, 3500)
ax.grid(which='major', color='lightgrey', linestyle='-')
ax.xaxis.set_ticks_position("top")
ax.xaxis.set_label_position("top")
ax.spines["top"].set_position(("axes", 1.02))
for ax in [ax2, ax3, ax4, ax6]:
plt.setp(ax.get_yticklabels(), visible = False)
plt.tight_layout()
fig.subplots_adjust(wspace = 0.15)
# -
|
07 - Working With LASIO.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: tensorflow
# language: python
# name: tensorflow
# ---
# # T81-558: Applications of Deep Neural Networks
# **Class 7: Kaggle Data Sets.**
# * Instructor: [<NAME>](https://sites.wustl.edu/jeffheaton/), School of Engineering and Applied Science, [Washington University in St. Louis](https://engineering.wustl.edu/Programs/Pages/default.aspx)
# * For more information visit the [class website](https://sites.wustl.edu/jeffheaton/t81-558/).
# # Helpful Functions from Before
# +
from sklearn import preprocessing
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import shutil
import os
# Encode text values to dummy variables(i.e. [1,0,0],[0,1,0],[0,0,1] for red,green,blue)
def encode_text_dummy(df, name):
dummies = pd.get_dummies(df[name])
for x in dummies.columns:
dummy_name = "{}-{}".format(name, x)
df[dummy_name] = dummies[x]
df.drop(name, axis=1, inplace=True)
# Encode text values to a single dummy variable. The new columns (which do not replace the old) will have a 1
# at every location where the original column (name) matches each of the target_values. One column is added for
# each target value.
def encode_text_single_dummy(df, name, target_values):
for tv in target_values:
l = list(df[name].astype(str))
l = [1 if str(x) == str(tv) else 0 for x in l]
name2 = "{}-{}".format(name, tv)
df[name2] = l
# Encode text values to indexes(i.e. [1],[2],[3] for red,green,blue).
def encode_text_index(df, name):
le = preprocessing.LabelEncoder()
df[name] = le.fit_transform(df[name])
return le.classes_
# Encode a numeric column as zscores
def encode_numeric_zscore(df, name, mean=None, sd=None):
if mean is None:
mean = df[name].mean()
if sd is None:
sd = df[name].std()
df[name] = (df[name] - mean) / sd
# Convert all missing values in the specified column to the median
def missing_median(df, name):
med = df[name].median()
df[name] = df[name].fillna(med)
# Convert all missing values in the specified column to the default
def missing_default(df, name, default_value):
df[name] = df[name].fillna(default_value)
# Convert a Pandas dataframe to the x,y inputs that TensorFlow needs
def to_xy(df, target):
result = []
for x in df.columns:
if x != target:
result.append(x)
# find out the type of the target column. Is it really this hard? :(
target_type = df[target].dtypes
target_type = target_type[0] if hasattr(target_type, '__iter__') else target_type
# Encode to int for classification, float otherwise. TensorFlow likes 32 bits.
if target_type in (np.int64, np.int32):
# Classification
return df.as_matrix(result).astype(np.float32), df.as_matrix([target]).astype(np.int32)
else:
# Regression
return df.as_matrix(result).astype(np.float32), df.as_matrix([target]).astype(np.float32)
# Nicely formatted time string
def hms_string(sec_elapsed):
h = int(sec_elapsed / (60 * 60))
m = int((sec_elapsed % (60 * 60)) / 60)
s = sec_elapsed % 60
return "{}:{:>02}:{:>05.2f}".format(h, m, s)
# Regression chart, we will see more of this chart in the next class.
def chart_regression(pred, y):
t = pd.DataFrame({'pred': pred, 'y': y.flatten()})
t.sort_values(by=['y'], inplace=True)
a = plt.plot(t['y'].tolist(), label='expected')
b = plt.plot(t['pred'].tolist(), label='prediction')
plt.ylabel('output')
plt.legend()
plt.show()
# Get a new directory to hold checkpoints from a neural network. This allows the neural network to be
# loaded later. If the erase param is set to true, the contents of the directory will be cleared.
def get_model_dir(name, erase):
base_path = os.path.join(".", "dnn")
model_dir = os.path.join(base_path, name)
os.makedirs(model_dir, exist_ok=True)
if erase and len(model_dir) > 4 and os.path.isdir(model_dir):
shutil.rmtree(model_dir, ignore_errors=True) # be careful, this deletes everything below the specified path
return model_dir
# Remove all rows where the specified column is +/- sd standard deviations
def remove_outliers(df, name, sd):
drop_rows = df.index[(np.abs(df[name] - df[name].mean()) >= (sd * df[name].std()))]
df.drop(drop_rows, axis=0, inplace=True)
# Encode a column to a range between normalized_low and normalized_high.
def encode_numeric_range(df, name, normalized_low=-1, normalized_high=1,
data_low=None, data_high=None):
if data_low is None:
data_low = min(df[name])
data_high = max(df[name])
df[name] = ((df[name] - data_low) / (data_high - data_low)) \
* (normalized_high - normalized_low) + normalized_low
# -
# # What is Kaggle?
#
# [Kaggle](http://www.kaggle.com) runs competitions in which data scientists compete in order to provide the best model to fit the data. The capstone project of this chapter features Kaggle’s [Titanic data set](https://www.kaggle.com/c/titanic-gettingStarted). Before we get started with the Titanic example, it’s important to be aware of some Kaggle guidelines. First, most competitions end on a specific date. Website organizers have currently scheduled the Titanic competition to end on December 31, 2016. However, they have already extended the deadline several times, and an extension beyond 2014 is also possible. Second, the Titanic data set is considered a tutorial data set. In other words, there is no prize, and your score in the competition does not count towards becoming a Kaggle Master.
# # Kaggle Ranks
#
# Kaggle ranks are achieved by earning gold, silver and bronze medals.
#
# * [Kaggle Top Users](https://www.kaggle.com/rankings)
# * [Current Top Kaggle User's Profile Page](https://www.kaggle.com/stasg7)
# * [<NAME>'s (your instructor) Kaggle Profile](https://www.kaggle.com/jeffheaton)
# * [Current Kaggle Ranking System](https://www.kaggle.com/progression)
# # Typical Kaggle Competition
#
# A typical Kaggle competition will have several components. Consider the Titanic tutorial:
#
# * [Competition Summary Page](https://www.kaggle.com/c/titanic)
# * [Data Page](https://www.kaggle.com/c/titanic/data)
# * [Evaluation Description Page](https://www.kaggle.com/c/titanic/details/evaluation)
# * [Leaderboard](https://www.kaggle.com/c/titanic/leaderboard)
#
# ## How Kaggle Competitions are Scored
#
# Kaggle is provided with a data set by the competition sponsor. This data set is divided up as follows:
#
# * **Complete Data Set** - This is the complete data set.
# * **Training Data Set** - You are provided both the inputs and the outcomes for the training portion of the data set.
# * **Test Data Set** - You are provided the complete test data set; however, you are not given the outcomes. Your submission is your predicted outcomes for this data set.
# * **Public Leaderboard** - You are not told what part of the test data set contributes to the public leaderboard. Your public score is calculated based on this part of the data set.
# * **Private Leaderboard** - You are not told what part of the test data set contributes to the public leaderboard. Your final score/rank is calculated based on this part. You do not see your private leaderboard score until the end.
#
# 
#
# ## Preparing a Kaggle Submission
#
# Code need not be submitted to Kaggle. For competitions, you are scored entirely on the accuracy of your sbmission file. A Kaggle submission file is always a CSV file that contains the **Id** of the row you are predicting and the answer. For the titanic competition, a submission file looks something like this:
#
# ```
# PassengerId,Survived
# 892,0
# 893,1
# 894,1
# 895,0
# 896,0
# 897,1
# ...
# ```
#
# The above file states the prediction for each of various passengers. You should only predict on ID's that are in the test file. Likewise, you should render a prediction for every row in the test file. Some competitions will have different formats for their answers. For example, a multi-classification will usually have a column for each class and your predictions for each class.
# # Select Kaggle Competitions
#
# There have been many interesting competitions on Kaggle, these are some of my favorites.
#
# ## Predictive Modeling
#
# * [Otto Group Product Classification Challenge](https://www.kaggle.com/c/otto-group-product-classification-challenge)
# * [Galaxy Zoo - The Galaxy Challenge](https://www.kaggle.com/c/galaxy-zoo-the-galaxy-challenge)
# * [Practice Fusion Diabetes Classification](https://www.kaggle.com/c/pf2012-diabetes)
# * [Predicting a Biological Response](https://www.kaggle.com/c/bioresponse)
#
# ## Computer Vision
#
# * [Diabetic Retinopathy Detection](https://www.kaggle.com/c/diabetic-retinopathy-detection)
# * [Cats vs Dogs](https://www.kaggle.com/c/dogs-vs-cats)
# * [State Farm Distracted Driver Detection](https://www.kaggle.com/c/state-farm-distracted-driver-detection)
#
# ## Time Series
#
# * [The Marinexplore and Cornell University Whale Detection Challenge](https://www.kaggle.com/c/whale-detection-challenge)
#
# ## Other
#
# * [Helping Santa's Helpers](https://www.kaggle.com/c/helping-santas-helpers)
#
# # Iris as a Kaggle Competition
#
# If the Iris data were used as a Kaggle, you would be given the following three files:
#
# * [kaggle_iris_test.csv](https://github.com/jeffheaton/t81_558_deep_learning/blob/master/data/kaggle_iris_test.csv) - The data that Kaggle will evaluate you on. Contains only input, you must provide answers. (contains x)
# * [kaggle_iris_train.csv](https://github.com/jeffheaton/t81_558_deep_learning/blob/master/data/kaggle_iris_train.csv) - The data that you will use to train. (contains x and y)
# * [kaggle_iris_sample.csv](https://github.com/jeffheaton/t81_558_deep_learning/blob/master/data/kaggle_iris_sample.csv) - A sample submission for Kaggle. (contains x and y)
#
# Important features of the Kaggle iris files (that differ from how we've previously seen files):
#
# * The iris species is already index encoded.
# * Your training data is in a separate file.
# * You will load the test data to generate a submission file.
#
# The following program generates a submission file for "Iris Kaggle". You can use it as a starting point for assignment 3.
# +
import os
import pandas as pd
from sklearn.model_selection import train_test_split
import tensorflow as tf
import tensorflow.contrib.learn as learn
import numpy as np
from tensorflow.contrib.learn.python.learn.metric_spec import MetricSpec
# Set the desired TensorFlow output level for this example
tf.logging.set_verbosity(tf.logging.FATAL)
path = "./data/"
filename_train = os.path.join(path,"kaggle_iris_train.csv")
filename_test = os.path.join(path,"kaggle_iris_test.csv")
filename_submit = os.path.join(path,"kaggle_iris_submit.csv")
df_train = pd.read_csv(filename_train,na_values=['NA','?'])
# Encode feature vector
encode_numeric_zscore(df_train,'petal_w')
encode_numeric_zscore(df_train,'petal_l')
encode_numeric_zscore(df_train,'sepal_w')
encode_numeric_zscore(df_train,'sepal_l')
df_train.drop('id', axis=1, inplace=True)
num_classes = len(df_train.groupby('species').species.nunique())
print("Number of classes: {}".format(num_classes))
# Create x & y for training
# Create the x-side (feature vectors) of the training
x, y = to_xy(df_train,'species')
# Split into train/test
x_train, x_validate, y_train, y_validate = train_test_split(
x, y, test_size=0.25, random_state=42)
# Get/clear a directory to store the neural network to
model_dir = get_model_dir('iris-kaggle',True)
# Create a deep neural network with 3 hidden layers of 30, 20, 5
feature_columns = [tf.contrib.layers.real_valued_column("", dimension=x.shape[1])]
classifier = learn.DNNClassifier(
model_dir= model_dir,
config=tf.contrib.learn.RunConfig(save_checkpoints_secs=1),
hidden_units=[30, 20, 5], n_classes=num_classes, feature_columns=feature_columns)
# Might be needed in future versions of "TensorFlow Learn"
#classifier = learn.SKCompat(classifier) # For Sklearn compatibility
# Early stopping
validation_monitor = tf.contrib.learn.monitors.ValidationMonitor(
x_validate,
y_validate,
every_n_steps=500,
early_stopping_metric="loss",
early_stopping_metric_minimize=True,
early_stopping_rounds=50)
# Fit/train neural network
classifier.fit(x_train, y_train,monitors=[validation_monitor],steps=10000)
# +
from sklearn import metrics
# Calculate multi log loss error
pred = list(classifier.predict_proba(x_validate, as_iterable=True))
score = metrics.log_loss(y_validate, pred)
print("Log loss score: {}".format(score))
# +
# Generate Kaggle submit file
# Encode feature vector
df_test = pd.read_csv(filename_test,na_values=['NA','?'])
encode_numeric_zscore(df_test,'petal_w')
encode_numeric_zscore(df_test,'petal_l')
encode_numeric_zscore(df_test,'sepal_w')
encode_numeric_zscore(df_test,'sepal_l')
ids = df_test['id']
df_test.drop('id', axis=1, inplace=True)
x = df_test.as_matrix().astype(np.float32)
# Generate predictions
pred = list(classifier.predict_proba(x, as_iterable=True))
#pred
# Create submission data set
df_submit = pd.DataFrame(pred)
df_submit.insert(0,'id',ids)
df_submit.columns = ['id','species-0','species-1','species-2']
df_submit.to_csv(filename_submit, index=False)
print(df_submit)
# -
# # Programming Assignment 3
#
# Kaggke competition site for current semester (Spring 2017):
# * [Spring 2017 Kaggle Assignment](https://inclass.kaggle.com/c/applications-of-deep-learning-wustl-spring-2017)
# * [Assignment File](https://raw.githubusercontent.com/jeffheaton/t81_558_deep_learning/master/pdf/t81_559_program_3.pdf)
#
# Output from generation program.
#
# ```
# Total pages: 17,303,347
# Template pages: 580,572
# Article pages: 8,852,289
# Redirect pages: 7,870,486
# Elapsed time: 0:56:07.48
# ```
#
# Previous Kaggle competition sites for this class (for your reference, do not use):
# * [Fall 2016 Kaggle Assignment](https://inclass.kaggle.com/c/wustl-t81-558-washu-deep-learning-fall-2016)
#
#
#
#
|
t81_558_class7_kaggle.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/aarjunsrinivasan/CatsvsDogs-Classification/blob/master/catdog.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="XlXW6JfcPe1D" colab_type="text"
# # Cats vs Dogs Images
# + [markdown] id="SH6ly97PPe1H" colab_type="text"
# ## Imports
# + id="P0EpqO0qPe1L" colab_type="code" colab={}
import time
import os
import numpy as np
import torch
import torch.nn.functional as F
import torch.nn as nn
from torch.utils.data import Dataset
from torch.utils.data import DataLoader
from torchvision import transforms
from PIL import Image
import matplotlib.pyplot as plt
if torch.cuda.is_available():
torch.backends.cudnn.deterministic = True
# + id="IzBaTw9jPe1o" colab_type="code" colab={}
# %matplotlib inline
# + [markdown] id="m0qwSPJ9Pe2D" colab_type="text"
# ## Settings
# + id="DO4ef2EOPe2H" colab_type="code" outputId="20aaa747-3b03-4250-d276-e627cf19c143" colab={"base_uri": "https://localhost:8080/", "height": 35}
##########################
### SETTINGS
##########################
# Devic
DEVICE = torch.device("cuda" if torch.cuda.is_available() else "cpu")
# Hyperparameters
RANDOM_SEED = 1
LEARNING_RATE = 0.00001
NUM_EPOCHS = 10
BATCH_SIZE = 500
# Architecture
NUM_CLASSES = 2
DEVICE
# + id="R5aWmB5dW66R" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 128} outputId="ecfd5126-9656-4c68-8c95-122b9c87125a"
from google.colab import drive
drive.mount('/content/drive')
# + id="_iQYBDyYPe2S" colab_type="code" outputId="b3cd8dac-67b1-4e10-c15a-9d1e1b5940e5" colab={"base_uri": "https://localhost:8080/", "height": 54}
import os
num_train_cats = len([i for i in os.listdir('/content/drive/My Drive/train/')
if i.endswith('.jpg') and i.startswith('cat')])
num_train_dogs = len([i for i in os.listdir('/content/drive/My Drive/train/')
if i.endswith('.jpg') and i.startswith('dog')])
print(f'Training set cats: {num_train_cats}')
print(f'Training set dogs: {num_train_dogs}')
# + [markdown] id="U0Ry21oVPe2l" colab_type="text"
# The naming scheme within each of these subfolders is `<class>.<imagenumber>.jpg`.
# + id="XNsRblZsPe2p" colab_type="code" outputId="33f1d68f-0a34-4e89-ae87-b795885e6c12" colab={}
img = Image.open(os.path.join('/home/arjun/Downloads/dogs-vs-cats','train', 'cat.59.jpg'))
print(np.asarray(img, dtype=np.uint8).shape)
plt.imshow(img);
# + [markdown] id="a6fbbLfIPe2-" colab_type="text"
# ### Creating Validation and Test Subsets
# + [markdown] id="N5ufUjGoPe3A" colab_type="text"
# - Move 2500 images from the training folder into a test set folder
# - Move 2500 images from the training folder into a validation set folder
# + id="1vgKO2cBPe3C" colab_type="code" colab={}
if not os.path.exists(os.path.join('/content/drive/My Drive/', 'test')):
os.mkdir(os.path.join('/content/drive/My Drive/', 'test'))
if not os.path.exists(os.path.join('/content/drive/My Drive/', 'valid')):
os.mkdir(os.path.join('/content/drive/My Drive/', 'valid'))
# + id="RU-5gCXnPe3L" colab_type="code" colab={}
for fname in os.listdir(os.path.join('/content/drive/My Drive/', 'train')):
if not fname.endswith('.jpg'):
continue
_, img_num, _ = fname.split('.')
filepath = os.path.join('/content/drive/My Drive/', 'train', fname)
img_num = int(img_num)
if img_num > 11249:
os.rename(filepath, filepath.replace('train', 'test'))
elif img_num > 9999:
os.rename(filepath, filepath.replace('train', 'valid'))
# + [markdown] id="q9txCTUXPe3R" colab_type="text"
# ### Standardizing Images
# + [markdown] id="V4_3Q_KnPe3T" colab_type="text"
# Getting mean and standard devation for normalizing images via z-score normalization. For details, see the related notebook [./cnn-standardized.ipynb](cnn-standardized.ipynb).
# + id="aC74MC1oPe3V" colab_type="code" outputId="187d4583-ad60-4497-d400-8f91333b29b2" colab={"base_uri": "https://localhost:8080/", "height": 54}
class CatsDogsDataset(Dataset):
"""Custom Dataset for loading CelebA face images"""
def __init__(self, img_dir, transform=None):
self.img_dir = img_dir
self.img_names = [i for i in
os.listdir(img_dir)
if i.endswith('.jpg')]
self.y = []
for i in self.img_names:
if i.split('.')[0] == 'cat':
self.y.append(0)
else:
self.y.append(1)
self.transform = transform
def __getitem__(self, index):
img = Image.open(os.path.join(self.img_dir,
self.img_names[index]))
if self.transform is not None:
img = self.transform(img)
label = self.y[index]
return img, label
def __len__(self):
return len(self.y)
custom_transform1 = transforms.Compose([transforms.Resize([64, 64]),
transforms.ToTensor()])
train_dataset = CatsDogsDataset(img_dir=os.path.join('/content/drive/My Drive/', 'train'),
transform=custom_transform1)
train_loader = DataLoader(dataset=train_dataset,
batch_size=500,
shuffle=False)
train_mean = []
train_std = []
for i, image in enumerate(train_loader, 0):
numpy_image = image[0].numpy()
batch_mean = np.mean(numpy_image, axis=(0, 2, 3))
batch_std = np.std(numpy_image, axis=(0, 2, 3))
train_mean.append(batch_mean)
train_std.append(batch_std)
break
train_mean = torch.tensor(np.mean(train_mean, axis=0))
train_std = torch.tensor(np.mean(train_std, axis=0))
print('Mean:', train_mean)
print('Std Dev:', train_std)
# + [markdown] id="SlsloRy5Pe3k" colab_type="text"
# ### Dataloaders
# + id="Sy4WOQVtPe3o" colab_type="code" colab={}
data_transforms = {
'train': transforms.Compose([
transforms.RandomRotation(5),
transforms.RandomHorizontalFlip(),
transforms.RandomResizedCrop(64, scale=(0.96, 1.0), ratio=(0.95, 1.05)),
transforms.ToTensor(),
transforms.Normalize(train_mean, train_std)
]),
'valid': transforms.Compose([
transforms.Resize([64, 64]),
transforms.ToTensor(),
transforms.Normalize(train_mean, train_std)
]),
}
train_dataset = CatsDogsDataset(img_dir=os.path.join('/content/drive/My Drive/', 'train'),
transform=data_transforms['train'])
train_loader = DataLoader(dataset=train_dataset,
batch_size=BATCH_SIZE,
drop_last=True,
shuffle=True)
valid_dataset = CatsDogsDataset(img_dir=os.path.join('/content/drive/My Drive/', 'valid'),
transform=data_transforms['valid'])
valid_loader = DataLoader(dataset=valid_dataset,
batch_size=BATCH_SIZE,
shuffle=False)
test_dataset = CatsDogsDataset(img_dir=os.path.join('/content/drive/My Drive/', 'test'),
transform=data_transforms['valid'])
test_loader = DataLoader(dataset=test_dataset,
batch_size=BATCH_SIZE,
shuffle=False)
# + [markdown] id="zxcRkcfoPe33" colab_type="text"
# ## Model
# + id="wysj3unVPe35" colab_type="code" colab={}
##########################
### MODEL
##########################
class mymodel(torch.nn.Module):
def __init__(self, num_classes):
super(mymodel, self).__init__()
# calculate same padding:
# (w - k + 2*p)/s + 1 = o
# => p = (s(o-1) - w + k)/2
self.block_1 = nn.Sequential(
nn.Conv2d(in_channels=3,
out_channels=64,
kernel_size=(3, 3),
stride=(1, 1),
# (1(32-1)- 32 + 3)/2 = 1
padding=1),
nn.ReLU(),
nn.Conv2d(in_channels=64,
out_channels=64,
kernel_size=(3, 3),
stride=(1, 1),
padding=1),
nn.ReLU(),
nn.MaxPool2d(kernel_size=(2, 2),
stride=(2, 2))
)
self.block_2 = nn.Sequential(
nn.Conv2d(in_channels=64,
out_channels=128,
kernel_size=(3, 3),
stride=(1, 1),
padding=1),
nn.ReLU(),nn.Dropout(p=0.2),
nn.Conv2d(in_channels=128,
out_channels=128,
kernel_size=(3, 3),
stride=(1, 1),
padding=1),
nn.ReLU(),
nn.MaxPool2d(kernel_size=(2, 2),
stride=(2, 2)),nn.Dropout(p=0.2)
)
self.block_3 = nn.Sequential(
nn.Conv2d(in_channels=128,
out_channels=256,
kernel_size=(3, 3),
stride=(1, 1),
padding=1),
nn.ReLU(),nn.Dropout(p=0.2),
nn.MaxPool2d(kernel_size=(2, 2),
stride=(2, 2))
)
self.block_4 = nn.Sequential(
nn.Conv2d(in_channels=256,
out_channels=512,
kernel_size=(3, 3),
stride=(1, 1),
padding=1),
nn.ReLU(),nn.Dropout(p=0.2),
nn.Conv2d(in_channels=512,
out_channels=512,
kernel_size=(3, 3),
stride=(1, 1),
padding=1),
nn.ReLU(),
nn.MaxPool2d(kernel_size=(2, 2),
stride=(2, 2))
)
self.block_5 = nn.Sequential(
nn.Conv2d(in_channels=512,
out_channels=512,
kernel_size=(3, 3),
stride=(1, 1),
padding=1),
nn.ReLU(),nn.Dropout(p=0.2),
nn.MaxPool2d(kernel_size=(2, 2),
stride=(2, 2))
)
self.classifier = nn.Sequential(
nn.Linear(512*2*2, 4096),
nn.ReLU(),
nn.Linear(4096, 4096),
nn.ReLU(),
nn.Linear(4096, num_classes)
)
for m in self.modules():
if isinstance(m, torch.nn.Conv2d):
#n = m.kernel_size[0] * m.kernel_size[1] * m.out_channels
#m.weight.data.normal_(0, np.sqrt(2. / n))
m.weight.detach().normal_(0, 0.05)
if m.bias is not None:
m.bias.detach().zero_()
elif isinstance(m, torch.nn.Linear):
m.weight.detach().normal_(0, 0.05)
m.bias.detach().detach().zero_()
def forward(self, x):
x = self.block_1(x)
x = self.block_2(x)
x = self.block_3(x)
x = self.block_4(x)
x = self.block_5(x)
logits = self.classifier(x.view(-1, 512*2*2))
probas = F.softmax(logits, dim=1)
return logits, probas
# + id="ksmgvrYWPe4C" colab_type="code" colab={}
torch.manual_seed(RANDOM_SEED)
model = mymodel(num_classes=NUM_CLASSES)
model = model.to(DEVICE)
optimizer = torch.optim.Adam(model.parameters(), lr=LEARNING_RATE)
# + id="seGPUKM5ZwMU" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 840} outputId="4debf18a-9ceb-4d19-85d8-ee9ebe64c773"
model
# + [markdown] id="Ni0gbJynPe4O" colab_type="text"
# ## Training
# + id="bYZOqEeJPe4R" colab_type="code" outputId="a13362e8-63d8-4b94-d561-970fb9b2aa17" colab={"base_uri": "https://localhost:8080/", "height": 35}
def compute_accuracy_and_loss(model, data_loader, device):
correct_pred, num_examples = 0, 0
cross_entropy = 0.
for i, (features, targets) in enumerate(data_loader):
features = features.to(device)
targets = targets.to(device)
logits, probas = model(features)
cross_entropy += F.cross_entropy(logits, targets).item()
_, predicted_labels = torch.max(probas, 1)
num_examples += targets.size(0)
correct_pred += (predicted_labels == targets).sum()
return correct_pred.float()/num_examples * 100, cross_entropy/num_examples
start_time = time.time()
train_acc_lst, valid_acc_lst = [], []
train_loss_lst, valid_loss_lst = [], []
for epoch in range(NUM_EPOCHS):
model.train()
for batch_idx, (features, targets) in enumerate(train_loader):
### PREPARE MINIBATCH
features = features.to(DEVICE)
targets = targets.to(DEVICE)
### FORWARD AND BACK PROP
logits, probas = model(features)
cost = F.cross_entropy(logits, targets)
optimizer.zero_grad()
cost.backward()
### UPDATE MODEL PARAMETERS
optimizer.step()
### LOGGING
if not batch_idx % 20:
print (f'Epoch: {epoch+1:03d}/{NUM_EPOCHS:03d} | '
f'Batch {batch_idx:03d}/{len(train_loader):03d} |'
f' Cost: {cost:.4f}')
# no need to build the computation graph for backprop when computing accuracy
model.eval()
with torch.set_grad_enabled(False):
train_acc, train_loss = compute_accuracy_and_loss(model, train_loader, device=DEVICE)
valid_acc, valid_loss = compute_accuracy_and_loss(model, valid_loader, device=DEVICE)
train_acc_lst.append(train_acc)
valid_acc_lst.append(valid_acc)
train_loss_lst.append(train_loss)
valid_loss_lst.append(valid_loss)
print(f'Epoch: {epoch+1:03d}/{NUM_EPOCHS:03d} Train Acc.: {train_acc:.2f}%'
f' | Validation Acc.: {valid_acc:.2f}%')
elapsed = (time.time() - start_time)/60
print(f'Time elapsed: {elapsed:.2f} min')
elapsed = (time.time() - start_time)/60
print(f'Total Training Time: {elapsed:.2f} min')
# + id="nbck-T6gPe4p" colab_type="code" outputId="b97a8d3e-e399-4141-9b35-4e6446576913" colab={}
plt.plot(range(1, NUM_EPOCHS+1), train_loss_lst, label='Training loss')
plt.plot(range(1, NUM_EPOCHS+1), valid_loss_lst, label='Validation loss')
plt.legend(loc='upper right')
plt.ylabel('Cross entropy')
plt.xlabel('Epoch')
plt.show()
# + id="zZGOUgisPe49" colab_type="code" outputId="083c9c68-43ee-462f-ac46-ed69af8226c3" colab={}
plt.plot(range(1, NUM_EPOCHS+1), train_acc_lst, label='Training accuracy')
plt.plot(range(1, NUM_EPOCHS+1), valid_acc_lst, label='Validation accuracy')
plt.legend(loc='upper left')
plt.ylabel('Cross entropy')
plt.xlabel('Epoch')
plt.show()
# + [markdown] id="WBEf5bw1Pe5L" colab_type="text"
# ## Evaluation
# + id="_VWAc3z0Pe5N" colab_type="code" outputId="8afb7b80-0245-4ebf-d335-c6492464d54e" colab={}
model.eval()
with torch.set_grad_enabled(False): # save memory during inference
test_acc, test_loss = compute_accuracy_and_loss(model, test_loader, DEVICE)
print(f'Test accuracy: {test_acc:.2f}%')
# + id="J7Vbo2HTPe5W" colab_type="code" colab={}
class UnNormalize(object):
def __init__(self, mean, std):
self.mean = mean
self.std = std
def __call__(self, tensor):
"""
Args:
tensor (Tensor): Tensor image of size (C, H, W) to be normalized.
Returns:
Tensor: Normalized image.
"""
for t, m, s in zip(tensor, self.mean, self.std):
t.mul_(s).add_(m)
# The normalize code -> t.sub_(m).div_(s)
return tensor
unorm = UnNormalize(mean=train_mean, std=train_std)
# + id="PIQnH8OTPe5q" colab_type="code" outputId="aa467176-1060-4d05-d0d2-fcff4ca6989f" colab={}
test_loader = DataLoader(dataset=train_dataset,
batch_size=BATCH_SIZE,
shuffle=True)
for features, targets in test_loader:
break
_, predictions = model.forward(features[:8].to(DEVICE))
predictions = torch.argmax(predictions, dim=1)
d = {0: 'cat',
1: 'dog'}
fig, ax = plt.subplots(1, 8, figsize=(20, 10))
for i in range(8):
img = unorm(features[i])
ax[i].imshow(np.transpose(img, (1, 2, 0)))
ax[i].set_xlabel(d[predictions[i].item()])
plt.show()
|
Code/model_no_gpu.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/SamH3pn3r/DS-Unit-1-Sprint-1-Dealing-With-Data/blob/master/Copy_of_DS_Unit_1_Sprint_Challenge_1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="NooAiTdnafkz" colab_type="text"
# # Data Science Unit 1 Sprint Challenge 1
#
# ## Loading, cleaning, visualizing, and analyzing data
#
# In this sprint challenge you will look at a dataset of the survival of patients who underwent surgery for breast cancer.
#
# http://archive.ics.uci.edu/ml/datasets/Haberman%27s+Survival
#
# Data Set Information:
# The dataset contains cases from a study that was conducted between 1958 and 1970 at the University of Chicago's Billings Hospital on the survival of patients who had undergone surgery for breast cancer.
#
# Attribute Information:
# 1. Age of patient at time of operation (numerical)
# 2. Patient's year of operation (year - 1900, numerical)
# 3. Number of positive axillary nodes detected (numerical)
# 4. Survival status (class attribute)
# -- 1 = the patient survived 5 years or longer
# -- 2 = the patient died within 5 year
#
# Sprint challenges are evaluated based on satisfactory completion of each part. It is suggested you work through it in order, getting each aspect reasonably working, before trying to deeply explore, iterate, or refine any given step. Once you get to the end, if you want to go back and improve things, go for it!
# + [markdown] id="5wch6ksCbJtZ" colab_type="text"
# ## Part 1 - Load and validate the data
#
# - Load the data as a `pandas` data frame.
# - Validate that it has the appropriate number of observations (you can check the raw file, and also read the dataset description from UCI).
# - Validate that you have no missing values.
# - Add informative names to the features.
# - The survival variable is encoded as 1 for surviving >5 years and 2 for not - change this to be 0 for not surviving and 1 for surviving >5 years (0/1 is a more traditional encoding of binary variables)
#
# At the end, print the first five rows of the dataset to demonstrate the above.
# + id="CYZTMVbt_Yv2" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 367} outputId="b6539e01-fcfd-46e9-99d6-629c0018c83a"
# !pip install pandas==0.23.4
# + id="287TpoGKFRVK" colab_type="code" colab={}
# TODO
import pandas as pd
hab_data = pd.read_csv('haberman.data')
# + id="dBYV_7Lrt7xK" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 300} outputId="7d343f6b-c300-4497-cd01-6270995bbf56"
hab_data.describe()
#Missing a row of data
# + id="_Q_55VqeuWq5" colab_type="code" colab={}
fixed_hab_data = pd.read_csv('haberman.data', header = None)
# + id="_vpIfWAwu2oH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 300} outputId="90741322-4e91-44e6-c0c6-5ca2f9ddd338"
fixed_hab_data.describe()
#we now have the num of instances that we want
# + id="07XFHBwGvB21" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 109} outputId="8eb6662c-b00d-436b-d834-b846fc4d5955"
fixed_hab_data.isnull().sum()
#No missing values which matches the .names file
# + id="ZUNVW308vTHh" colab_type="code" colab={}
column_headers = ['Age','Year','Num Nodes','Survival Status']
# + id="gmb_EFnwvl9i" colab_type="code" colab={}
fixed_hab = pd.read_csv('haberman.data', names = column_headers, header = None)
# + id="ND4LEABjv0NY" colab_type="code" colab={}
new_stat = {1:1,
2:0}
fixed_hab['Survival Status'] = fixed_hab['Survival Status'].map(new_stat)
# + id="beorwi9uw4gd" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 206} outputId="7a3bac36-090d-435f-de29-2c562693d17a"
fixed_hab.head()
# + [markdown] id="G7rLytbrO38L" colab_type="text"
# ## Part 2 - Examine the distribution and relationships of the features
#
# Explore the data - create at least *2* tables (can be summary statistics or crosstabulations) and *2* plots illustrating the nature of the data.
#
# This is open-ended, so to remind - first *complete* this task as a baseline, then go on to the remaining sections, and *then* as time allows revisit and explore further.
#
# Hint - you may need to bin some variables depending on your chosen tables/plots.
# + id="IAkllgCIFVj0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 300} outputId="425d184e-4e08-43d2-d179-67224162b8d6"
# TODO
fixed_hab.describe()
# + id="RmHGjfcR0c_W" colab_type="code" colab={}
age_bins = pd.cut(fixed_hab['Age'], 6)
year_bins = pd.cut(fixed_hab['Year'], 6)
node_bins = pd.cut(fixed_hab['Num Nodes'], 6)
# + id="sLdCF7jpzm-o" colab_type="code" colab={}
As = pd.crosstab(age_bins, fixed_hab['Survival Status'])
Ys = pd.crosstab(year_bins, fixed_hab['Survival Status'])
Ns = pd.crosstab(node_bins, fixed_hab['Survival Status'])
# + id="-cg1XVmG0PFt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 269} outputId="a20b1690-7919-4443-a6b2-38e83337e7a1"
As
# + id="rxvhgc3A3ENU" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 269} outputId="7cc385f4-b8a4-45d9-9dea-87217edaed20"
Ys
# + id="T96qZtzS4uKj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 269} outputId="6d270ff3-9bfe-4b2c-bd22-56c21dbf3e73"
Ns
# + id="2jh9A1P31uc0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 785} outputId="e54d6d61-0e75-4b67-d01f-d16f874ee2e9"
As.plot();
Ys.plot();
Ns.plot();
# + id="Vr59bgbY1u05" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="ad1a51b4-c8cc-45a8-b6d4-8ade1476b85d"
As.plot(kind ='bar');
Ys.plot(kind='bar');
Ns.plot(kind='bar');
# + [markdown] id="ZM8JckA2bgnp" colab_type="text"
# ## Part 3 - Analysis and Interpretation
#
# Now that you've looked at the data, answer the following questions:
#
# - What is at least one feature that looks to have a positive relationship with survival?
# - What is at least one feature that looks to have a negative relationship with survival?
# - How are those two features related with each other, and what might that mean?
#
# Answer with text, but feel free to intersperse example code/results or refer to it from earlier.
# + [markdown] id="HJjHGpYJ3rvc" colab_type="text"
#
#
# * Age and Num Nodes looks to have a positive relationship with survival.
# * Year looks to have a negative relationship with survival.
# * Age is related to year but Num Nodes doesn't seem related to either. It could be that as you get older, you're more likely to get breast cancer and the more positive nodes, the more likely you are to die. Overall, if you get breast cancer at an older age, the more likely you are to die.
#
#
#
#
|
Copy_of_DS_Unit_1_Sprint_Challenge_1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ## Dict 和 Set
#
# ### dict和集合的基础知识
# * 字典是一系列由键(key)和值(value)配对组成的元素的集合: d = { 'name': 'Tony', 'job': 'DevOps'}
# * 在Python3.6+之后,字典是有序的, 键值对A先插入字典,键值对B后插入字典,但是当你打印字典的Keys列表时,你会发现B可能在A的前面。但是从Python 3.6开始,字典是变成有顺序的了。你先插入键值对A,后插入键值对B,那么当你打印Keys列表的时候,你就会发现B在A的后面。
# * 相比list和tuple,字典的性能更优,特别是对于增删改查,字典的操作复杂度是O(1)
# * 集合没有键和值的配对,是一系列无序的、唯一的元素组合。s = {'tony', 'python', 3, 2}
# * 虽然集合看起来很像list或者tuple,但是集合并不支持索引操作,因为集合本质上是一个哈希表,和列表不一样。
# +
s = [1, 2, 3, 4]
s[2]
s = {1, 2, 3, 4}
s[2]
# -
# ### dict和集合的创建
# dict和set的创建通常来说有以下几种方式
# ```bash
# d1 = {'name': 'jason', 'age': 20, 'gender': 'male'}
# d2 = dict({'name': 'jason', 'age': 20, 'gender': 'male'})
# d3 = dict([('name', 'jason'), ('age', 20), ('gender', 'male')])
# d4 = dict(name='jason', age=20, gender='male')
# d1 == d2 == d3 ==d4
# True
#
# s1 = {1, 2, 3}
# s2 = set([1, 2, 3])
# s1 == s2
# True
# ```
# ### dict的访问
# * dict可以直接通过索引键访问,如果不存在索引键,那么就会抛出异常
# ```bash
# d = {'name': 'jason', 'age': 20}
# d['name']
# 'jason'
# d['location']
# ```
# * 你也可以用get()函数来进行访问
# ```bash
# d = {'name': 'jason', 'age': 20}
# d.get('name')
# 'jason'
# d.get('location', 'null')
# 'null'
# ```
#
# ### set的访问
# * 集合并不支持索引操作,因为集合本质上是一个哈希表,和列表不一样。大家看看下面这个例子,觉得返回值是多少呢?
# ```bash
# s = {1, 2, 3}
# s[0]
# ```
# * 考考大家,如果我想从一个set里面获取第二个元素,有什么办法呢?
#
# ### dict和set的基本操作
# * 判断是否存在
# ```bash
#
# s = {1, 2, 3}
# 1 in s
# True
# 10 in s
# False
#
# d = {'name': 'jason', 'age': 20}
# 'name' in d
# True
# 'location' in d
# False
# ```
# * 增加元素
# ```bash
# d = {'name': 'jason', 'age': 20}
# d['gender'] = 'male' # 增加元素对'gender': 'male'
# d['dob'] = '1999-02-01' # 增加元素对'dob': '1999-02-01'
# s = {1, 2, 3}
# s.add(4) # 增加元素4到集合
# ```
# * 删除元素
# ```bash
# d.pop('dob') # 删除键为'dob'的元素对
# '1998-01-01'
#
# s = {1, 2, 3, 4}
# s.remove(4)
# ```
#
s = {1, 2, 3}
1 in s
# ### dict和set的排序
# 很多情况下,我们需要对字典或集合进行排序,比如,取出值最大的 50 对。而对于dict,我们通常会根据键或值,进行升序或降序排序
# ```bash
# d = {'b': 1, 'a': 2, 'c': 10}
# d_sorted_by_key = sorted(d.items(), key=lambda x: x[0]) # 根据字典键的升序排序
# d_sorted_by_value = sorted(d.items(), key=lambda x: x[1]) # 根据字典值的升序排序
# d_sorted_by_key
# [('a', 2), ('b', 1), ('c', 10)]
# d_sorted_by_value
# [('b', 1), ('a', 2), ('c', 10)]
# ```
# set的排序比较简单,直接sort就可以
# ```bash
# s = {3, 4, 2, 1}
# sorted(s) # 对集合的元素进行升序排序
# [1, 2, 3, 4]
# ```
#
# ### 字典和集合的工作原理
# 每次向字典或集合插入一个元素时,Python 会首先计算键的哈希值(hash(key)),再和 mask = PyDicMinSize - 1 做与操作,计算这个元素应该插入哈希表的位置 index = hash(key) & mask。如果哈希表中此位置是空的,那么这个元素就会被插入其中。
#
# 而如果此位置已被占用,Python 便会比较两个元素的哈希值和键是否相等。
# * 若两者都相等,则表明这个元素已经存在,如果值不同,则更新值。
# * 若两者中有一个不相等,这种情况我们通常称为哈希冲突(hash collision),意思是两个元素的键不相等,但是哈希值相等。这种情况下,Python 便会继续寻找表中空余的位置,直到找到位置为止。
#
# 插入、查找和删除的时间复杂度为 O(1)
#
# ### 应用举例
# 比如电商企业的后台,存储了每件产品的 ID、名称和价格。现在的需求是,给定某件商品的 ID,我们要找出其价格。
#
# 如果我们用列表来存储这些数据结构,并进行查找,相应的代码如下:
# ```bash
#
# def find_product_price(products, product_id):
# for id, price in products:
# if id == product_id:
# return price
# return None
#
# products = [
# (143121312, 100),
# (432314553, 30),
# (32421912367, 150)
# ]
#
# print('The price of product 432314553 is {}'.format(find_product_price(products, 432314553)))
#
# # 输出
# The price of product 432314553 is 30
# ```
#
# 用字典的代码
# ```bash
#
# products = {
# 143121312: 100,
# 432314553: 30,
# 32421912367: 150
# }
# print('The price of product 432314553 is {}'.format(products[432314553]))
#
# # 输出
# The price of product 432314553 is 30
# ```
# ### 思考
# 既然我们有了list和tuple?我们为什么需要集合?
# +
l1 = [1, 2, 3, 4, 5, 5, 5, 6]
temp_l1 = []
for x in l1:
if x not in temp_l1:
temp_l1.append(x)
print(temp_l1)
temp_l2 = list(set(l1))
print(temp_l2)
# -
|
module1/jupyter/dict-and-set.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
my_set=set()
my_set
my_set.add(1)
my_set.add(2)
my_set.add(2)
my_set
my_set={1,2,3,4,5,6,6,6,6,6,6,7,7,7}
my_set
|
trial scripts/sets.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: webAutomation
# language: python
# name: webautomation
# ---
# +
from single_bet_type_analyzer import SingleBetTypeAnalyzer
from dask.distributed import LocalCluster
from scrapers.betflag import BetflagScraper
from scrapers.betfair import BetfairScraper
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
import ipywidgets as widgets
from IPython.display import display, clear_output
import winsound
import time
def play_beep():
for i in range(3):
for j in range(2):
winsound.Beep(1000, 100)
time.sleep(0.05)
time.sleep(0.7)
cluster = LocalCluster(processes=False)
cluster
# +
sport = widgets.Dropdown(
options=['calcio', 'tennis', 'basket'],
value='calcio',
description='Sport:',
disabled=False,
)
bet_type = widgets.Dropdown(
options=['1x2', '12', 'uo1.5', 'uo2.5', 'uo3.5', 'uo4.5'],
value='1x2',
description='Bet Type:',
disabled=False,
)
offline = widgets.Checkbox(
value=True,
description='Non-Live',
disabled=False
)
live = widgets.Checkbox(
value=True,
description='Live',
disabled=False
)
headless = widgets.Checkbox(
value=True,
description='Headless',
disabled=False
)
widgets.VBox([sport, bet_type, offline, live, headless])
# -
# %%time
try:
if (sport.value != 'calcio' and bet_type.value != '12') or (sport.value == 'calcio' and bet_type.value == '12'):
print('Error: coppia sport-bet_type non accettata!')
analyzer = None
else:
analyzer = SingleBetTypeAnalyzer(sport=sport.value, bet_type=bet_type.value, cluster=cluster,
offline=offline.value, live=live.value, headless=headless.value)
# analyzer = SingleBetTypeAnalyzer('basket', '12', cluster=cluster)
except:
analyzer.close()
finally:
print('Fine')
# %%time
df = analyzer.analyze_bets()
while True:
df = analyzer.analyze_bets()
clear_output()
display(df)
if len(df) > 0:
play_beep()
time.sleep(50)
time.sleep(10)
analyzer.close()
cluster.close()
|
analyzer_notebook.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/oferbaharav/Data-Science/blob/master/airbnb_berlin_notebook_(Ofer_update_Fri_28_Feb).ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="92O-J9K9I-13" colab_type="code" colab={}
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from scipy import stats
# In order to see all of the columns of the dataset we need to set the display options
# from the Pandas package to at least 100 (the dataset has 96 columns) and, for the rows,
# I set it to at least 100 which will help when I check for null values and dtypes.
pd.set_option('display.max_columns', 100)
pd.set_option('display.max_rows', 100)
# + id="KWMuEo-UI-1-" colab_type="code" colab={}
# Importing the CSV 'listings_summary.csv' from the Kaggle dataset found at this
# URL: https://www.kaggle.com/brittabettendorf/berlin-airbnb-data
listings_summary = pd.read_csv('https://raw.githubusercontent.com/BuildWeekAirbnbOptimal2/Datascience/master/Berlin.csv')
# + id="uF5N9hUFI-2D" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="9d4bde3f-b926-4404-a270-1f7637549f17"
# As stated above, there are 96 columns and over 20,000 observations
listings_summary.shape
# + id="Y7B0JP7_I-2I" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="2a8bcefd-fef1-4c68-c21f-e29982317e67"
# Checking the dtypes of the dataset...
# The goal of this project is to find the optimal price for an AirBnB in Belin, Germany so,
# the target variable will be the 'price' which is currently an object and therefore, will
# have to be dealt with appropriately.
listings_summary.dtypes
# + id="pO0Bo1oDI-2M" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="d9728322-0f30-4b63-cfed-fe7375a49eef"
# Next we will check for the null values within the dataset - there are quite a few...
listings_summary.isna().sum()
# + id="LnhJr1aPI-2Q" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 202} outputId="e1a13d86-920b-430e-a4f8-aefe0462b2dc"
# Calling the head of the dataset to visualize what the first row of observations looks like
listings_summary.head(1)
# + id="qLjDA1dEI-2U" colab_type="code" colab={}
# We can already tell later on we will have to drop a few columns where the cardinality for some
# object features, while finite, will be very high epecially in the case of URLs, names, reviews,
# descriptions, etc. so we will remove a few of them now and possibly later.
# + id="j-LG5uTeI-2Y" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="0af02fd8-ea45-4ec3-cfc8-67783e79edac"
# First, we will use a for loop to check the number of unique values in each column. This is acheived
# by taking the length of the value_counts of a column.
for col in listings_summary:
print(f'There are/is {len(listings_summary[col].value_counts())} unique value(s) for column: {col}') if listings_summary[col].dtypes=='O' else print(None)
# + id="fYrC8g-QI-2b" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 202} outputId="1c9e649b-f581-4fdc-907c-9149fb0779f0"
listings_summary.head(1)
# + id="ND4mo69tI-2e" colab_type="code" colab={}
# The first thing we will do is remove the object columns with high cardinality and features that are probably
# redundant like 'city' since this is the Berlin AirBnB dataset - 'zipcode' may be useful but neighbourhood could
# cover that.
high_cardin = ['listing_url', 'name', 'summary', 'space', 'description', 'experiences_offered', 'neighborhood_overview',
'notes', 'transit', 'access', 'interaction', 'house_rules', 'thumbnail_url', 'medium_url',
'picture_url', 'xl_picture_url', 'host_url', 'host_name', 'host_about', 'host_thumbnail_url',
'host_picture_url', 'host_verifications', 'street', 'city', 'state', 'zipcode', 'market',
'smart_location', 'country_code', 'country', 'bed_type', 'amenities', 'weekly_price', 'monthly_price',
'has_availability', 'calendar_last_scraped', 'requires_license', 'license', 'is_business_travel_ready',
'require_guest_profile_picture', 'require_guest_phone_verification']
# + id="CRV8gcp9I-2i" colab_type="code" colab={}
listings_df = listings_summary.drop(columns=high_cardin)
# + id="A_43oL4II-2m" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 985} outputId="4449481f-e1d1-4f21-aef2-6776a2dd1dbe"
listings_df.isna().sum()
# + id="Lw0Hx1ZKI-2p" colab_type="code" colab={}
# We will also remove columns that have many NaN values
high_na = ['host_response_time', 'host_response_rate', 'host_acceptance_rate', 'square_feet', 'jurisdiction_names']
Berlin_airbnb = listings_df.drop(columns=high_na)
# + id="sXWJOOiDI-2u" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 898} outputId="584fa378-919e-4bc4-b9bb-48971237005e"
Berlin_airbnb.dtypes
# + id="yBMvU63lI-2x" colab_type="code" colab={}
# Next we will engineer some features based on the data
# + id="8vhYXw5bI-21" colab_type="code" colab={}
# Originally, the 'security_deposit' column would've been kept and replaced NaN values with the mean but,
# Since there are many NaN values we will make a binary feature stating '1' if they require a security deposit
# and '0' if the do not require one.
# TODO: drop Berlin_airbnb['security_deposit']
has_security_dep = []
for i in Berlin_airbnb['security_deposit']:
if i==np.NaN:
has_security_dep.append(0)
else:
has_security_dep.append(1)
Berlin_airbnb['require_security_deposit'] = np.array(has_security_dep).astype(int)
# + id="xrIaWPgGI-24" colab_type="code" colab={}
# We will do the same with cleaning fee and call it 'has_cleaning_service'...
# TODO: drop Berlin_airbnb['cleaning_fee']
has_cleaning = []
for i in Berlin_airbnb['cleaning_fee']:
if i==np.NaN:
has_cleaning.append(0)
else:
has_cleaning.append(1)
Berlin_airbnb['has_cleaning_service'] = np.array(has_cleaning).astype(int)
# + id="SgHBQcbBI-27" colab_type="code" colab={}
# Possible columns to impute or use for feature engineering
# review_scores_rating - mode = 100.00 (46 unique values between 50.00 and 100.00)
# review_scores_accuracy - mode = 10.0 (more than 50% of the data)
# review_scores_cleanliness - mode = 10.0
# review_scores_checkin - mode = 10.0 (more than 50% of the data)
# review_scores_communication - mode = 10.0 (more than 50% of the data)
# review_scores_location - mode = 10.0
# review_scores_value - mode = 10.0
# + id="K-8mgCyoI-2-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="bd37db60-ae97-4d8a-bd9e-c4add8eb27d3"
# Next, we will get rid of the dollar signs and any commas that may be contained in the 'price'
# and 'extra_people' column by making a function that will strip the dollar sign ('$') from the
# array, remove the redundant '.00', and then remove commas for amounts 1000 or larger
def dollar_to_int(row):
return row.strip('$')[:-3]
def no_comma(row):
return row.replace(',','')
# To show it works...
amount = dollar_to_int('$1,300.00')
print(no_comma(amount))
# + id="twMMCI5_I-3A" colab_type="code" colab={}
# Applying them to the dataset...
Berlin_airbnb['price'] = Berlin_airbnb['price'].apply(dollar_to_int).apply(no_comma).astype(int)
Berlin_airbnb['extra_people'] = Berlin_airbnb['extra_people'].apply(dollar_to_int).apply(no_comma).astype(int)
# + id="0ZIyggKuI-3D" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="1460981c-8c87-4274-95d2-9b6f4a167515"
Berlin_airbnb.shape
# + id="PM7Wcuo8I-3H" colab_type="code" colab={}
Berlin_airbnb = Berlin_airbnb.drop(columns=['security_deposit', 'cleaning_fee'])
# + id="vZKt5jcKI-3L" colab_type="code" colab={}
# 'property_type', 'room_type', 'accommodates','bathrooms', 'bedrooms', 'beds', 'bed_type','price','number_of_reviews',('review_scores_value '),'instant_bookable','cancellation_policy','neighbourhood','host_identity_verified'
# + id="cOLXPpfvI-3P" colab_type="code" colab={}
# Possibly useful: - Predicting 'PRICE'
# 1. neighbourhood
# 2. property type
# 3. room type
# 4. accommodates
# 5. bathrooms
# 6. bedrooms
# 7. beds
# 8. reviews_scores_value
# 9. instant_bookable
# 10. cancellation_policy
# 10. has_cleaning_service
### Columns we may go with
# 'property_type', 'room_type', 'accommodates','bathrooms', 'bedrooms', 'beds', 'bed_type','price','number_of_reviews',('review_scores_value '),'instant_bookable','cancellation_policy','neighbourhood','host_identity_verified'
# + id="SLc7ZmZVI-3S" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 305} outputId="6c8bdc44-c3df-4925-c1de-08b98570a102"
Berlin_subset = Berlin_airbnb[['property_type', 'room_type', 'accommodates', 'bathrooms', 'bedrooms', 'beds',
'price', 'number_of_reviews', 'review_scores_value', 'instant_bookable',
'cancellation_policy', 'neighbourhood', 'host_identity_verified']]
Berlin_subset.head()
# + id="r95hu8ugI-3V" colab_type="code" colab={}
###### We need to include why we are using these columns!! ######
# i.e. Why we chose to condense 'accommodates'
# + id="HtGGD_-NI-3Z" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 259} outputId="8ec09559-ce02-4551-af60-ff493056f3cd"
Berlin_subset.dtypes
# + id="d2INBQWzI-3b" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 311} outputId="3c939244-d65e-4772-dcb6-e995aad7b615"
Berlin_subset['accommodates'].value_counts()
# + id="wD4od-onI-3e" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="bb1891fb-5667-4a56-beef-f81d664b4648"
# Minimizing the values for the accommodates column
# We will make them objects from 1-6 and then 7+
accommodate = []
for int in Berlin_subset['accommodates']:
if int==1:
accommodate.append('1')
elif int==2:
accommodate.append('2')
elif int==3:
accommodate.append('3')
elif int==4:
accommodate.append('4')
elif int==5:
accommodate.append('5')
elif int==6:
accommodate.append('6')
elif int>=7:
accommodate.append('7+')
else:
accommodate.append('')
set(accommodate)
# + id="K5UalDf3I-3h" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="b3d4a8b3-a82c-4491-b913-16c6b5ec0197"
len(Berlin_subset['accommodates'])==len(accommodate)
# + id="RtHBEzDbI-3l" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 121} outputId="b973ce75-0315-40c2-d6af-b80dcfb35a00"
Berlin_subset['can_accommodate'] = np.array(accommodate)
# + id="mq5Np0tNI-3o" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="821d408a-98e1-4106-ec05-7e8305e9f70a"
bedrooms = []
for bed in Berlin_subset['bedrooms']:
if bed==1.0:
bedrooms.append('1')
else:
bedrooms.append('2+')
set(bedrooms)
# + id="Xty0gVezI-3r" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 121} outputId="89487da7-1365-44d1-a6bc-0dc896c3c815"
Berlin_subset['n_bedrooms'] = np.array(bedrooms)
# + id="WVMEGMfrI-3u" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="0a5e1256-3b57-43f1-84ba-c652efa154f0"
bathrooms = []
for bath in Berlin_subset['bathrooms']:
if bath==1.0:
bathrooms.append('1')
else:
bathrooms.append('2+')
set(bathrooms)
# + id="eCdAtQHfI-3x" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 121} outputId="6c08b3bc-d58a-4701-daed-201aa97bedfb"
Berlin_subset['n_bathrooms'] = np.array(bathrooms)
# + id="9_LR1D2JI-30" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="54d2f893-ef36-46c7-f4e0-3e25b6e32668"
beds = []
for bed in Berlin_subset['beds']:
if bed==1.0:
beds.append('1')
else:
beds.append('2+')
set(beds)
# + id="D-BZUxb4I-33" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 121} outputId="944656e6-3aa4-4cb6-8649-6c175b099148"
Berlin_subset['n_beds'] = np.array(beds)
# + id="ZO8GdBJTI-36" colab_type="code" colab={}
def to_nbool(array):
for i in array:
if i=='t':
return 1
else:
return 0
# + id="FEe6nJFtI-38" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 121} outputId="174c4fc1-9191-4082-f926-4903335f4c49"
Berlin_subset['host_identity_verified'] = Berlin_subset['host_identity_verified'].dropna().apply(to_nbool)
# + id="jeZHJ2arI-3_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 121} outputId="9100a80d-19c0-41c8-eec3-b08ec66d1506"
Berlin_subset['instant_bookable'] = Berlin_subset['instant_bookable'].dropna().apply(to_nbool)
# + id="l-Kcuj0EI-4C" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 121} outputId="31c2be74-3828-4201-c926-9f0ac362172e"
Berlin_subset['review_scores_value'] = Berlin_subset['review_scores_value'].replace(np.NaN, 0)
# + id="I3Kp1--AI-4F" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="c4b42051-6868-4a03-8bb4-37119ff6b624"
scores = []
for rating in Berlin_subset['review_scores_value']:
if rating>=7.0:
scores.append(rating)
else:
scores.append(0.0)
set(scores)
# + id="pIkZmXVdI-4I" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 121} outputId="0c0364f9-17d4-434a-f798-b9f3133e1d90"
Berlin_subset['review_score'] = scores
# + id="Z1ODuJ56I-4L" colab_type="code" colab={}
Berlin = Berlin_subset.drop(columns=['accommodates', 'bathrooms', 'bedrooms',
'beds', 'review_scores_value'])
# + id="6tMFTWcMI-4N" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="9d9a3159-6a38-44ac-ef60-51392be8897a"
Berlin.shape
# + id="vYoWIuPfI-4R" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 305} outputId="23541419-2dcb-4120-b3da-9c39209e13a3"
Berlin.head()
# + id="c3g74IFVa0G7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="0405c2e0-86c4-4b7d-da55-f6b246beb630"
len(Berlin.columns)
# + id="ChDrWEX_ctUL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 259} outputId="df2595b0-1825-4d91-b4c1-31142d413144"
Berlin.isnull().sum()
# + id="TOPPNgOzdVjZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="d6bd49e5-d1d3-4f41-cb66-6f2e0c3de513"
#stripping NaN values
berlin_na_stripped = Berlin[Berlin['neighbourhood'].notna() & Berlin['host_identity_verified'].notna()]
berlin_na_stripped.shape
# + id="7okBlQ0nevLG" colab_type="code" colab={}
Berlin = berlin_na_stripped
# + id="Oxq-TNo5I-4V" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="3a2ab771-f78b-405d-cb53-0df70d7a9205"
#Ofer starts here, continues on above work by James
#Create Train/Test split:
import pandas as pd
from sklearn import datasets, linear_model
from sklearn.model_selection import train_test_split
from matplotlib import pyplot as plt
# create training and testing vars
X = Berlin.drop(columns='price')
y = Berlin.price
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2)
print(X_train.shape, y_train.shape)
print(X_test.shape, y_test.shape)
# + id="zstAoQ7gYzme" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 365} outputId="3d3e563a-73fc-47cd-8166-f5206530c427"
# Arrange data into X features matrix and y target vector
target = 'price'
# !pip install --upgrade category_encoders
import category_encoders as ce
from sklearn.pipeline import make_pipeline
from sklearn.impute import SimpleImputer
from sklearn.ensemble import RandomForestRegressor
from sklearn.metrics import mean_absolute_error
pipeline = make_pipeline(
ce.OrdinalEncoder(),
# SimpleImputer(strategy='median'),
RandomForestRegressor(n_estimators=250, random_state=42, n_jobs=-1)
)
# Fit on train, score on test
pipeline.fit(X_train, y_train)
y_pred_train = pipeline.predict(X_train)
y_pred_test = pipeline.predict(X_test)
rf = pipeline.named_steps['randomforestregressor']
encoder = pipeline.named_steps['ordinalencoder']
# Print Results
print('Training R^2', pipeline.score(X_train, y_train))
print(f'Training MAE: {mean_absolute_error(y_train, y_pred_train)} dollars')
print('Validation R^2', pipeline.score(X_test, y_test))
print(f'Validation MAE: {mean_absolute_error(y_test, y_pred_test)} dollars')
# + id="TB_j4WcNfeqd" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 390} outputId="91ceece2-a9dc-4348-e7df-7f984277bc8b"
# Get feature importances
rf = pipeline.named_steps['randomforestregressor']
importances = pd.Series(rf.feature_importances_, X_train.columns)
# Plot feature importances
# %matplotlib inline
import matplotlib.pyplot as plt
plt.figure(figsize=(8,6))
plt.title('Feature Importance')
importances.sort_values().plot.barh(color='grey');
# + id="aVMNUVL9hslE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 542} outputId="254c6def-1010-465c-aa43-3d085d43a960"
#try to graph it out
import plotly.express as px
px.scatter(Berlin, x='neighbourhood', y= target)
#this shows pricey neighbourhoods from left to right
# + id="XwV2xl5Gixzm" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 542} outputId="2a869559-3e78-415b-8c8a-50ba2732d6b5"
#try to graph it out
import plotly.express as px
px.scatter(Berlin, x='number_of_reviews', y= target)
#this shows the less reviews, the higher the price (this probably suggests that highly priced properties don't get booked much)
# + id="IJlbL-tPieVP" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 296} outputId="c3665026-a153-4ea1-9a85-dcf227e50e4c"
#try to graph it out
import seaborn as sns
sns.boxplot(y_train)
# + id="aXJXrZfklKyI" colab_type="code" colab={}
#throw some shapley values
# # !pip install shap
# import shap
# X_train_encoded = encoder.transform(X_train)
# row = X_train_encoded
# explainer = shap.TreeExplainer(rf)
# shap_values = explainer.shap_values(row)
# shap.initjs()
# shap.force_plot(
# # shap.summary_plot(
# base_value=explainer.expected_value,
# shap_values=shap_values,
# features=row
# )
# + id="c9yHXjqOp56x" colab_type="code" colab={}
# # Feature Scaling
# from sklearn.preprocessing import StandardScaler
# sc = StandardScaler()
# X_train = sc.fit_transform(X_train)
# X_test = sc.transform(X_test)
# + id="WXpTqjA4t88K" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 257} outputId="04bf5ae5-9a0a-4a5d-8235-b8939bb99fba"
X_test.head(4)
# + id="S2H58mlqnEE3" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="3bfb76f1-fb0b-4beb-bbaa-2242c9052751"
#I think this is predicting the first 4 rows prices? so, $32, $31, $113.9, $67 ?
#Can someone verify this? :)
y_pred = pipeline.predict(X_test[:3])
y_pred
# + id="SN3SD9q_wMQQ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 104} outputId="fb9842f7-9f80-4f1c-bf64-0359538476ec"
#looking up the first 4 prices of y
y_test.head(4)
#So wait - I thought this was a 1 dimensional array
#what is the number on the left? on the right it's the price correct?
# + id="5fYzedfTA5xX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 210} outputId="453898f2-7d09-4687-885c-69f90bc2d4a1"
X_test.head(3)
# + id="XOV-jOcfw3-B" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 242} outputId="6c19ba74-d0b5-40dc-94aa-df6851371f7f"
X_test.dtypes
# + id="XuKRlBHeAOiV" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="37643e74-ea2d-4dcd-c72c-40bc14fac78f"
X_test.n_bathrooms.unique()
# + id="4qrQmoCmAt3n" colab_type="code" colab={}
|
airbnb_berlin_notebook_(Ofer_update_Fri_28_Feb).ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# # Slider Demo
#
#
# Using the slider widget to control visual properties of your plot.
#
# In this example, a slider is used to choose the frequency of a sine
# wave. You can control many continuously-varying properties of your plot in
# this way.
#
#
# +
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.widgets import Slider, Button, RadioButtons
fig, ax = plt.subplots()
plt.subplots_adjust(left=0.25, bottom=0.25)
t = np.arange(0.0, 1.0, 0.001)
a0 = 5
f0 = 3
delta_f = 5.0
s = a0 * np.sin(2 * np.pi * f0 * t)
l, = plt.plot(t, s, lw=2)
ax.margins(x=0)
axcolor = 'lightgoldenrodyellow'
axfreq = plt.axes([0.25, 0.1, 0.65, 0.03], facecolor=axcolor)
axamp = plt.axes([0.25, 0.15, 0.65, 0.03], facecolor=axcolor)
sfreq = Slider(axfreq, 'Freq', 0.1, 30.0, valinit=f0, valstep=delta_f)
samp = Slider(axamp, 'Amp', 0.1, 10.0, valinit=a0)
def update(val):
amp = samp.val
freq = sfreq.val
l.set_ydata(amp*np.sin(2*np.pi*freq*t))
fig.canvas.draw_idle()
sfreq.on_changed(update)
samp.on_changed(update)
resetax = plt.axes([0.8, 0.025, 0.1, 0.04])
button = Button(resetax, 'Reset', color=axcolor, hovercolor='0.975')
def reset(event):
sfreq.reset()
samp.reset()
button.on_clicked(reset)
rax = plt.axes([0.025, 0.5, 0.15, 0.15], facecolor=axcolor)
radio = RadioButtons(rax, ('red', 'blue', 'green'), active=0)
def colorfunc(label):
l.set_color(label)
fig.canvas.draw_idle()
radio.on_clicked(colorfunc)
plt.show()
|
slider_demo.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <table border="0">
# <tr>
# <td>
# <img src="https://ictd2016.files.wordpress.com/2016/04/microsoft-research-logo-copy.jpg" style="width 30px;" />
# </td>
# <td>
# <img src="https://www.microsoft.com/en-us/research/wp-content/uploads/2016/12/MSR-ALICE-HeaderGraphic-1920x720_1-800x550.jpg" style="width 100px;"/></td>
# </tr>
# </table>
# # Metalearners Estimators: Use Cases and Examples
# Metalearners are binary treatment CATE estimators that model the response surfaces, $Y(0)$ and $Y(1)$, separately. To account for a heterogeneous propensity of treatment $P(T\mid X)$, the two modeled responses $E(Y(0)\mid X)$ and $E(Y(1)\mid X)$ are weighted in different ways in the final CATE estimation. For a detailed overview of these methods, see [this paper](https://arxiv.org/abs/1706.03461).
#
# The EconML SDK implements the following `metalearners`:
#
# * T-Learner
#
# * S-Learner
#
# * X-Learner
#
# * DomainAdaptation-Learner
#
# * DoublyRobust-Learner
#
# In this notebook, we compare the performance of these four CATE estimatiors on synthetic data and semi-synthetic data.
#
# **Notebook contents:**
#
# 1. Example usage with synthetic data
#
# 2. Example usage with semi-synthetic data
# +
# Main imports
from econml.metalearners import TLearner, SLearner, XLearner, DomainAdaptationLearner
# Helper imports
import numpy as np
from numpy.random import binomial, multivariate_normal, normal, uniform
from sklearn.linear_model import LinearRegression
from sklearn.ensemble import RandomForestRegressor, RandomForestClassifier, GradientBoostingRegressor
import matplotlib.pyplot as plt
# %matplotlib inline
# -
# ## 1. Example Usage with Synthetic Data
# ### 1.1. DGP
# We use the data generating process (DGP) from [Kunzel et al.](https://arxiv.org/abs/1706.03461). The DGP is described by the following equations:
#
# $
# Y = \mu_1(x) \cdot T + \mu_0(x) \cdot (1-T) + \epsilon \\
# T \sim Bern(e(x)), \; e(x) = P(T=1|X=x)
# $
#
# where
#
# $
# \mu_0(x) = x^T\beta,\; with \;\beta\sim Unif([-3, 3]^d),\; X_i \sim N(0, \Sigma)\\
# \mu_1(x) = \mu_0(x) + 8 \mathbb{I}(x_2>0.1) => CATE(x) = 8 \mathbb{I}(x_2>0.1)
# $
#
# Define DGP
def generate_data(n, d, controls_outcome, treatment_effect, propensity):
"""Generates population data for given untreated_outcome, treatment_effect and propensity functions.
Parameters
----------
n (int): population size
d (int): number of covariates
controls_outcome (func): untreated outcome conditional on covariates
treatment_effect (func): treatment effect conditional on covariates
propensity (func): probability of treatment conditional on covariates
"""
# Generate covariates
X = multivariate_normal(np.zeros(d), np.diag(np.ones(d)), n)
# Generate treatment
T = np.apply_along_axis(lambda x: binomial(1, propensity(x), 1)[0], 1, X)
# Calculate outcome
Y0 = np.apply_along_axis(lambda x: controls_outcome(x), 1, X)
treat_effect = np.apply_along_axis(lambda x: treatment_effect(x), 1, X)
Y = Y0 + treat_effect * T
return (Y, T, X)
# controls outcome, treatment effect, propensity definitions
def generate_controls_outcome(d):
beta = uniform(-3, 3, d)
return lambda x: np.dot(x, beta) + normal(0, 1)
treatment_effect = lambda x: (1 if x[1] > 0.1 else 0)*8
propensity = lambda x: (0.8 if (x[2]>-0.5 and x[2]<0.5) else 0.2)
# DGP constants and test data
d = 5
n = 1000
n_test = 250
controls_outcome = generate_controls_outcome(d)
X_test = multivariate_normal(np.zeros(d), np.diag(np.ones(d)), n_test)
delta = 6/n_test
X_test[:, 1] = np.arange(-3, 3, delta)
Y, T, X = generate_data(n, d, controls_outcome, treatment_effect, propensity)
# ### 1.2. Train Estimators
# Instantiate T learner
models = GradientBoostingRegressor(n_estimators=100, max_depth=6, min_samples_leaf=int(n/100))
T_learner = TLearner(models=models)
# Train T_learner
T_learner.fit(Y, T, X=X)
# Estimate treatment effects on test data
T_te = T_learner.effect(X_test)
# Instantiate S learner
overall_model = GradientBoostingRegressor(n_estimators=100, max_depth=6, min_samples_leaf=int(n/100))
S_learner = SLearner(overall_model=overall_model)
# Train S_learner
S_learner.fit(Y, T, X=X)
# Estimate treatment effects on test data
S_te = S_learner.effect(X_test)
# Instantiate X learner
models = GradientBoostingRegressor(n_estimators=100, max_depth=6, min_samples_leaf=int(n/100))
propensity_model = RandomForestClassifier(n_estimators=100, max_depth=6,
min_samples_leaf=int(n/100))
X_learner = XLearner(models=models, propensity_model=propensity_model)
# Train X_learner
X_learner.fit(Y, T, X=X)
# Estimate treatment effects on test data
X_te = X_learner.effect(X_test)
# Instantiate Domain Adaptation learner
models = GradientBoostingRegressor(n_estimators=100, max_depth=6, min_samples_leaf=int(n/100))
final_models = GradientBoostingRegressor(n_estimators=100, max_depth=6, min_samples_leaf=int(n/100))
propensity_model = RandomForestClassifier(n_estimators=100, max_depth=6,
min_samples_leaf=int(n/100))
DA_learner = DomainAdaptationLearner(models=models,
final_models=final_models,
propensity_model=propensity_model)
# Train DA_learner
DA_learner.fit(Y, T, X=X)
# Estimate treatment effects on test data
DA_te = DA_learner.effect(X_test)
# +
# Instantiate Doubly Robust Learner
from econml.dr import DRLearner
outcome_model = GradientBoostingRegressor(n_estimators=100, max_depth=6, min_samples_leaf=int(n/100))
pseudo_treatment_model = GradientBoostingRegressor(n_estimators=100, max_depth=6, min_samples_leaf=int(n/100))
propensity_model = RandomForestClassifier(n_estimators=100, max_depth=6,
min_samples_leaf=int(n/100))
DR_learner = DRLearner(model_regression=outcome_model, model_propensity=propensity_model,
model_final=pseudo_treatment_model, cv=5)
# Train DR_learner
DR_learner.fit(Y, T, X=X)
# Estimate treatment effects on test data
DR_te = DR_learner.effect(X_test)
# -
# ### 1.3. Visual Comparisons
### Comparison plot of the different learners
plt.figure(figsize=(7, 5))
plt.plot(X_test[:, 1], np.apply_along_axis(treatment_effect, 1, X_test), color='black', ls='--', label='Baseline')
plt.scatter(X_test[:, 1], T_te, label="T-learner")
plt.scatter(X_test[:, 1], S_te, label="S-learner")
plt.scatter(X_test[:, 1], DA_te, label="DA-learner")
plt.scatter(X_test[:, 1], X_te, label="X-learner")
plt.scatter(X_test[:, 1], DR_te, label="DR-learner")
plt.xlabel('$x_1$')
plt.ylabel('Treatment Effect')
plt.legend()
plt.show()
# Visualization of bias distribution
expected_te = np.apply_along_axis(treatment_effect, 1, X_test)
plt.violinplot([np.abs(T_te - expected_te),
np.abs(S_te - expected_te),
np.abs(DA_te - expected_te),
np.abs(X_te - expected_te),
np.abs(DR_te - expected_te)
], showmeans=True)
plt.ylabel("Bias distribution")
plt.xticks([1, 2, 3, 4, 5], ['T-learner', 'S-learner', 'DA-learner', 'X-learner', 'DR-learner'])
plt.show()
# ## 2. Example Usage with Semi-synthetic Data
# ### 2.1. DGP
#
# We use the Response Surface B from [Hill (2011)](https://www.tandfonline.com/doi/pdf/10.1198/jcgs.2010.08162) to generate sythetic outcome surfaces from real-world covariates and treatment assignments (Infant Health Development Program data). Since the original data was part of a randomized trial, a subset of the treated infants (those with non-white mothers) has been removed from the data in order to mimic the observational data setting. For more details, see [Hill (2011)](https://www.tandfonline.com/doi/pdf/10.1198/jcgs.2010.08162).
#
#
# The DGP is described by the following equations:
#
# $
# Y(0) = e^{(X+W)\beta} + \epsilon_0, \;\epsilon_0 \sim N(0, 1)\\
# Y(1) = X\beta - \omega + \epsilon_1, \;\epsilon_1 \sim N(0, 1)\\
# $
#
# where $X$ is a covariate matrix, $W$ is a constant matrix with entries equal to $0.5$ and $w$ is a constant calculated such that the CATT equals $4$.
from econml.data.dgps import ihdp_surface_B
Y, T, X, expected_te = ihdp_surface_B()
# ### 2.2. Train Estimators
# T-learner
T_learner.fit(Y, T, X=X)
T_te = T_learner.effect(X)
# S-learner
S_learner.fit(Y, T, X=X)
S_te = S_learner.effect(X)
# X-learner
X_learner.fit(Y, T, X=X)
X_te = X_learner.effect(X)
# Domain adaptation learner
DA_learner.fit(Y, T, X=X)
DA_te = DA_learner.effect(X)
# Doubly robust learner
DR_learner.fit(Y, T, X=X)
DR_te = DR_learner.effect(X)
# ### 2.3. Visual Comparisons
# Visualization of bias distribution
plt.violinplot([np.abs(T_te - expected_te),
np.abs(S_te - expected_te),
np.abs(DA_te - expected_te),
np.abs(X_te - expected_te),
np.abs(DR_te - expected_te)
], showmeans=True)
plt.ylabel("Bias distribution")
plt.xticks([1, 2, 3, 4, 5], ['T-learner', 'S-learner', 'DA-learner', 'X-learner', 'DR-learner'])
plt.show()
|
notebooks/Metalearners Examples.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Непараметрические криетрии
# Критерий | Одновыборочный | Двухвыборочный | Двухвыборочный (связанные выборки)
# ------------- | -------------|
# **Знаков** | $\times$ | | $\times$
# **Ранговый** | $\times$ | $\times$ | $\times$
# **Перестановочный** | $\times$ | $\times$ | $\times$
# ## Терапия при анорексии
# В исследовании оценивается эффективность поведенческой терапии для лечения анорексии. Для 50 пациентов известен вес до начала терапии и по её окончании. Была ли терапия эффективной?
# +
import numpy as np
import pandas as pd
import itertools
from scipy import stats
from statsmodels.stats.descriptivestats import sign_test
from statsmodels.stats.weightstats import zconfint
# -
# %pylab inline
# ### Загрузка данных
weight_data = pd.read_csv('weight.txt', sep = '\t', header = 0)
weight_data.head()
# +
pylab.figure(figsize=(12,4))
pylab.subplot(1,2,1)
pylab.grid()
pylab.hist(weight_data.Before, color = 'r')
pylab.xlabel('Before')
pylab.subplot(1,2,2)
pylab.grid()
pylab.hist(weight_data.After, color = 'b')
pylab.xlabel('After')
pylab.show()
# -
weight_data.describe()
# ## Двухвыборочные критерии для связных выборок
# $H_0\colon$ медианы веса до и после терапии совпадает
#
# $H_1\colon$ медианы веса до и после тепрапии отличаются
print '95%% confidence interval for mean weight before therapy: [%f, %f]' % zconfint(weight_data.Before)
print '95%% confidence interval for mean weight after therapy: [%f, %f]' % zconfint(weight_data.After)
pylab.hist(weight_data.After - weight_data.Before)
pylab.show()
# ### Критерий знаков
# $H_0\colon P\left(X_1>X_2\right)=\frac1{2},$
#
# $H_1\colon P\left(X_1>X_2\right)\neq\frac1{2}$
print "M: %d, p-value: %f" % sign_test(weight_data.After - weight_data.Before)
# ### Критерий знаковых рангов Уилкоксона
# $H_0\colon med\left(X_1-X_2\right)=0,$
#
# $H_1\colon med\left(X_1-X_2\right)\neq0$
stats.wilcoxon(weight_data.After, weight_data.Before)
stats.wilcoxon(weight_data.After - weight_data.Before)
# ### Перестановочный критерий
# $H_0\colon \mathbb{E}(X_1 - X_2) = 0$
#
# $H_1\colon \mathbb{E}(X_1 - X_2) \neq 0$
def permutation_t_stat_1sample(sample, mean):
t_stat = sum(map(lambda x: x - mean, sample))
return t_stat
def permutation_zero_distr_1sample(sample, mean, max_permutations = None):
centered_sample = map(lambda x: x - mean, sample)
if max_permutations:
signs_array = set([tuple(x) for x in 2 * np.random.randint(2, size = (max_permutations,
len(sample))) - 1 ])
else:
signs_array = itertools.product([-1, 1], repeat = len(sample))
distr = [sum(centered_sample * np.array(signs)) for signs in signs_array]
return distr
pylab.hist(permutation_zero_distr_1sample(weight_data.After - weight_data.Before, 0.,
max_permutations = 10000))
pylab.show()
def permutation_test(sample, mean, max_permutations = None, alternative = 'two-sided'):
if alternative not in ('two-sided', 'less', 'greater'):
raise ValueError("alternative not recognized\n"
"should be 'two-sided', 'less' or 'greater'")
t_stat = permutation_t_stat_1sample(sample, mean)
zero_distr = permutation_zero_distr_1sample(sample, mean, max_permutations)
if alternative == 'two-sided':
return sum([1. if abs(x) >= abs(t_stat) else 0. for x in zero_distr]) / len(zero_distr)
if alternative == 'less':
return sum([1. if x <= t_stat else 0. for x in zero_distr]) / len(zero_distr)
if alternative == 'greater':
return sum([1. if x >= t_stat else 0. for x in zero_distr]) / len(zero_distr)
print "p-value: %f" % permutation_test(weight_data.After - weight_data.Before, 0.,
max_permutations = 1000)
print "p-value: %f" % permutation_test(weight_data.After - weight_data.Before, 0.,
max_permutations = 50000)
|
4 Stats for data analysis/Lectures notebooks/10 non-parametric tests rel/stat.non_parametric_tests_rel.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # SQL translators
# The purpose of this vignette is to walk through how expressions like `_.id.mean()` are converted into SQL.
#
# This process involves 3 parts
#
# 1. SQL translation functions, e.g. taking column "id" and producing the SQL "ROUND(id)".
# 2. SQL translation from a symbolic call
# - Converting method calls like `_.id.round(2)` to `round(_.id, 2)`
# - Looking up SQL translators (e.g. for "mean" function call)
# 3. Handling SQL partitions, like in OVER clauses
#
# ### Using sqlalchemy select statment for convenience
#
# Throughout this vignette, we'll use a select statement object from sqlalchemy,
# so we can conveniently access its columns as needed.
# +
from sqlalchemy import sql
col_names = ['id', 'x', 'y']
sel = sql.select([sql.column(x) for x in col_names])
print(sel)
print(type(sel.columns))
print(sel.columns)
# -
# ## Translator functions
#
# A SQL translator function takes...
#
# * a first argument that is a sqlalchemy Column
# * (optional) additional arguments for the translation
# ### A simple translator
# +
f_simple_round = lambda col, n: sql.func.round(col, n)
sql_expr = f_simple_round(sel.columns.x, 2)
print(sql_expr)
# -
# The function above is essentially what most translator functions are.
#
# For example, here is the round function defined for postgresql.
# One key difference is that it casts the column to a numeric beforehand.
# +
from siuba.sql.dialects.postgresql import funcs
f_round = funcs['scalar']['round']
sql_expr = f_round(sel.columns.x, 2)
print(sql_expr)
# -
# ### Handling windows with custom Over clauses
# +
f_win_mean = funcs['window']['mean']
sql_over_expr = f_win_mean(sel.columns.x)
print(type(sql_over_expr))
print(sql_over_expr)
# -
# Notice that this window expression has an empty over clause. This clause needs to be able to include any variables we've grouped the data by.
#
# Siuba handles this by implementing a `set_over` method on these custom sqlalchemy Over clauses, which takes grouping and ordering variables as arguments.
group_by_clause = sql.elements.ClauseList(sel.columns.x, sel.columns.y)
print(sql_over_expr.set_over(group_by_clause))
# ## Call shaping
#
# The section above discusses how SQL translators are functions that take a sqlalchemy column, and return a SQL expression. However, when using siuba we often have expressions like...
#
# ```
# mutate(data, x = _.y.round(2))
# ```
#
# In this case, before we can even use a SQL translator, we need to...
#
# * find the name and arguments of the method being called
# * find the column it is being called on
#
# This is done by using the `CallTreeLocal` class to analyze the tree of operations for each expression.
# +
from siuba.siu import Lazy, CallTreeLocal, Call, strip_symbolic
from siuba import _
_.y.round(2)
# -
# ### Example of translation with CallTreeLocal
# +
from siuba.sql.dialects.postgresql import funcs
local_funcs = {**funcs['scalar'], **funcs['window']}
call_shaper = CallTreeLocal(
local_funcs,
call_sub_attr = ('dt',)
)
# -
symbol = _.id.mean()
call = strip_symbolic(symbol)
print(call)
func_call = call_shaper.enter(call)
print(func_call(sel.columns))
# This is the same result as when we called the SQL translator for `mean` manually!
# In that section we also showed that we can set group information, so that it takes
# an average within each group.
#
# In this case it's easy to set group information to the Over clause.
# However, an additional challenge is when it's part of a larger expression...
# +
call2 = strip_symbolic(_.id.mean() + 1)
func_call2 = call_shaper.enter(call2)
func_call2(sel.columns)
# -
# ## Handling partitions
#
# While the first section showed how siuba's custom Over clauses can add grouping info to a translation, it is missing one key detail: expressions that generate Over clauses, like `_.id.mean()`, can be part of larger expressions. For example `_.id.mean() + 1`.
#
# In this case, if we look at the call tree for that expression, the top operation is the addition...
_.id.mean() + 1
# How can we create the appropriate expression...
#
# ```
# avg(some_col) OVER (PARTITION BY x, y) + 1
# ```
#
# when the piece that needs grouping info is not easily accessible? The answer is by using a tree visitor, which steps down every black rectangle in the call tree shown above, from top to bottom.
#
# ### Full example
#
# Below, we copy the code from the call shaping section..
# +
from siuba.sql.verbs import track_call_windows
from siuba import _
from siuba.sql.dialects.postgresql import funcs
local_funcs = {**funcs['scalar'], **funcs['window']}
call_shaper = CallTreeLocal(
local_funcs,
call_sub_attr = ('dt',)
)
symbol3 = _.id.mean() + 1
call3 = strip_symbolic(symbol3)
func_call3 = call_shaper.enter(call3)
# -
# Finally, we pass the shaped call...
# +
col, windows, window_cte = track_call_windows(
func_call3,
sel.columns,
group_by = ['x', 'y'],
order_by = [],
# note that this is optional, and results in window_cte being a
# copy of this select that contains the window clauses
window_cte = sel.select()
)
print(col)
print(windows)
print(window_cte)
|
docs/developer/sql-translators.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from tpot import TPOTClassifier
import pandas as pd
from sklearn.model_selection import train_test_split
import pandas as pd
from sklearn import metrics
import scikitplot as skplt
acidoCEST_ML = pd.read_csv('acido_CEST_MRI_MegaBox_01_to_08_clean.csv')
acidoCEST_ML = acidoCEST_ML.drop(['Unnamed: 0','ApproT1(sec)','Temp','FILE','Conc(mM)','ExpT1(ms)', 'ExpT2(ms)', 'ExpB1(percent)', 'ExpB0(ppm)',
'ExpB0(Hz)', 'SatPower(uT)', 'SatTime(ms)'], axis = 1)
# acidoCEST_ML.head(2)
acidoCEST_ML.columns
# ## pH > 7.0
X_train, X_test, y_train, y_test = train_test_split( acidoCEST_ML.drop('pH',axis=1) , 1*(acidoCEST_ML.pH > 7.0 ), test_size=0.30, random_state=42)
tpot = TPOTClassifier(generations=10, population_size=10, verbosity=2, n_jobs= 3 , cv = 3,
template='PCA-Selector-Classifier',early_stop=3,max_time_mins=60
,scoring ='precision_weighted')
tpot.fit(X_train,y_train)
skplt.metrics.plot_confusion_matrix(y_test, tpot.predict(X_test), normalize=False)
print( metrics.classification_report(y_test, tpot.predict(X_test)) )
# ### Next
# - Do we need the data at (power= 0.5 and 1.0 uT, sat time = 0.50 or 1.0 seconds) (heat map of error 2d)
# - How much does measuring other things beside the Z spectra help?
# - Are the regression outlier due to Power and T1
# - Effect of noise
#
# - Do these later
# 2. Can we start with fewer frequencies ?
# 3. What frequencies are needed ?
# 5. Max 27 or 25 frequencies
# 6. pH higher than 6.8
#
#
#
# ## pH > 6.5
X_train, X_test, y_train, y_test = train_test_split( acidoCEST_ML.drop('pH',axis=1) , 1*(acidoCEST_ML.pH > 6.5 ), test_size=0.30, random_state=42)
tpot = TPOTClassifier(generations=10, population_size=10, verbosity=2, n_jobs= 3 , cv = 3,
template='PCA-Selector-Classifier',early_stop=3,max_time_mins=60
,scoring ='precision_weighted')
tpot.fit(X_train,y_train)
skplt.metrics.plot_confusion_matrix(y_test, tpot.predict(X_test), normalize=False)
print( metrics.classification_report(y_train, tpot.predict(X_train)) )
tpot.export('acidoCEST_ML_tpot_pH6p5_classifier.py')
acidoCEST_ML.head()
metrics.classification_report( 1*(acidoCEST_ML.pH > 6.5), acidoCEST_ML.drop('pH', axis = 1))
|
2-TPOT classifier-Copy1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python3
# ---
# <h1 style="color:blue">Predicting Car Prices</h1>
# <p>In this project I will try to predict the prices of some cars with the help K-Nearest Neighbor algorithm</p>
#importing libraries
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.neighbors import KNeighborsRegressor
from sklearn.metrics import mean_squared_error
from sklearn.metrics import r2_score
from sklearn.model_selection import cross_val_score,KFold
# +
# The code was removed by Watson Studio for sharing.
# +
# selecting only values that are continuous
continuous_values_cols = ['normalized-losses', 'wheel-base', 'length', 'width', 'height', 'curb-weight', 'bore', 'stroke', 'compression-rate', 'horsepower', 'peak-rpm', 'city-mpg', 'highway-mpg', 'price']
numeric_cars = df_cars[continuous_values_cols]
numeric_cars.head()
# -
# <h2>Data Cleaning</h2>
numeric_cars=numeric_cars.replace("?",np.nan)
numeric_cars.head()
#converting datatype of float
numeric_cars=numeric_cars.astype('float')
numeric_cars.isnull().sum()
# Because `price` is the column we want to predict, let's remove any rows with missing `price` values.
numeric_cars.dropna(subset=["price"],inplace=True)
numeric_cars.isnull().sum()
# Replace missing values in other columns using column means.
numeric_cars=numeric_cars.fillna(numeric_cars.mean())
numeric_cars.isnull().sum()
# Normalizing the values
price_col = numeric_cars['price']
numeric_cars = (numeric_cars - numeric_cars.min())/(numeric_cars.max() - numeric_cars.min())
numeric_cars['price'] = price_col
# <h1>Training and testing the data</h1>
x = numeric_cars[['normalized-losses', 'wheel-base', 'length', 'width', 'height', 'curb-weight', 'bore', 'stroke', 'compression-rate', 'horsepower', 'peak-rpm', 'city-mpg', 'highway-mpg']]
x.head()
y=numeric_cars["price"]
y.head()
rmsesavg=[] # for root mean square error average
rmsestd=[] # for root mean square error standard deviation
for i in range(4,32,4):
kf=KFold(i,shuffle=True,random_state=1)
knn=KNeighborsRegressor()
mses=cross_val_score(knn,x,y,scoring='neg_mean_squared_error',cv=kf)
rmsesavg.append(np.mean(np.sqrt(np.absolute(mses))))
rmsestd.append(np.std(np.sqrt(np.absolute(mses))))
plt.scatter([4,8,12,16,20,24,28],rmsesavg)
plt.show()
plt.scatter([4,8,12,16,20,24,28],rmsestd)
plt.show()
print("From the above plots it seems that for our case the value k = 16 is near to ideal")
# <h1>Splitting Data</h1>
p=np.random.rand(len(numeric_cars))<0.8
Train=numeric_cars[p]
Test=numeric_cars[~p]
len(Train)
Train.columns
# +
Train_X = Train[['normalized-losses', 'wheel-base', 'length', 'width', 'height', 'curb-weight', 'bore', 'stroke', 'compression-rate', 'horsepower', 'peak-rpm', 'city-mpg', 'highway-mpg']]
Train_Y = Train['price']
Test_X = Test[['normalized-losses', 'wheel-base', 'length', 'width', 'height', 'curb-weight', 'bore', 'stroke', 'compression-rate', 'horsepower', 'peak-rpm', 'city-mpg', 'highway-mpg']]
Test_Y = Test['price']
# -
# <h1>Prediction</h1>
# +
knn = KNeighborsRegressor()
knn.fit(Train_X,Train_Y)
pred = knn.predict(Test_X)
mse = mean_squared_error(Test_Y,pred)
rmse = np.sqrt(np.absolute(mse))
print(rmse)
# -
r2score = r2_score(Test_Y,pred)
print(r2score)
|
Python/Machine Learning/Predicting_car_prices/Predicting car prices.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python2
# ---
from articledata import *
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from nltk.corpus import names
from nltk.tag import StanfordNERTagger
import os
from nltk.tokenize import word_tokenize
data = pd.read_pickle('/Users/teresaborcuch/capstone_project/notebooks/ss_entity_data.pkl')
data.head(1)
labeled_names = ([(name, 'male') for name in names.words("male.txt")] + [(name, 'female') for name in names.words('female.txt')])
male_names = names.words("male.txt")
female_names = names.words("female.txt")
male_names[:5]
people, _ = evaluate_entities(data = data, section = 'politics', source = 'Fox')
# +
count = 0
for person in people:
if person in male_names:
count +=1
print count
# +
count = 0
for person in people:
if person in female_names:
count +=1
print count
# -
male_counts = []
female_counts = []
os.environ['CLASSPATH'] = "/Users/teresaborcuch/stanford-ner-2013-11-12/stanford-ner.jar"
os.environ['STANFORD_MODELS'] = '/Users/teresaborcuch/stanford-ner-2013-11-12/classifiers'
st = StanfordNERTagger('english.all.3class.distsim.crf.ser.gz')
for x in data['body']:
m_count = 0
f_count = 0
tokens = word_tokenize(x)
tags = st.tag(tokens)
for pair in tags:
if pair[1] == 'PERSON':
if pair[0] in male_names:
m_count += 1
elif pair[0] in female_names:
f_count += 1
else:
continue
male_counts.append(m_count)
female_counts.append(f_count)
data['f_count'] = female_counts
data['m_count'] = male_counts
data.to_pickle('/Users/teresaborcuch/capstone_project/notebooks/ss_entity_data.pkl')
data.head(1)
data.pivot_table(index = ['condensed_section'], values = ['f_count', 'm_count']).sort_values('f_count', ascending = False)
|
notebooks/ner_mf_count.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="LI3aEhtENm6t" colab_type="code" outputId="82102fe1-42fd-4f13-c68b-dd98d066d709" executionInfo={"status": "ok", "timestamp": 1583456406491, "user_tz": -60, "elapsed": 11307, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13241675469300863486"}} colab={"base_uri": "https://localhost:8080/", "height": 302}
# !pip install --upgrade tables
# !pip install eli5
# !pip install xgboost
# + id="stssTYi_OfgN" colab_type="code" colab={}
import pandas as pd
import numpy as np
from sklearn.dummy import DummyRegressor
from sklearn.tree import DecisionTreeRegressor
from sklearn.ensemble import RandomForestRegressor
import xgboost as xgb
from sklearn.metrics import mean_absolute_error as mae
from sklearn.model_selection import cross_val_score, KFold
import eli5
from eli5.sklearn import PermutationImportance
# + id="wAZBT_ONQKwj" colab_type="code" outputId="4be15058-d8a2-40da-c0e3-6a44990bd09b" executionInfo={"status": "ok", "timestamp": 1583456411613, "user_tz": -60, "elapsed": 713, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13241675469300863486"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# cd "/content/drive/My Drive/Colab Notebooks/matrix/matrix_two/dw_matrix_car"
# + id="FEQVAQN8Q7wx" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 128} outputId="00186aed-3e96-458e-a428-02a0ece33f55" executionInfo={"status": "error", "timestamp": 1583456413510, "user_tz": -60, "elapsed": 803, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13241675469300863486"}}
Wczytywanie danych
# + id="i3zFSyWfQIgK" colab_type="code" outputId="a6ea1a8e-7979-4aa0-e4b3-79230b78cb10" executionInfo={"status": "ok", "timestamp": 1583456417685, "user_tz": -60, "elapsed": 3220, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13241675469300863486"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
df = pd.read_hdf('data/car.h5')
df.shape
# + [markdown] id="FA5FJwP5VgCN" colab_type="text"
# **Feature Engineering**
# + id="LEZVROyEWAae" colab_type="code" colab={}
SUFFIX_CAT = '_cat'
for feat in df.columns:
if isinstance(df[feat][0], list): continue
factorized_values = df[feat].factorize()[0]
if SUFFIX_CAT in feat:
df[feat] = factorized_values
else:
df[feat + SUFFIX_CAT] = factorized_values
# + id="DWazfHsEYEx3" colab_type="code" outputId="66158d2d-e7c6-4ae4-def1-c237d120fd0d" executionInfo={"status": "ok", "timestamp": 1583456424511, "user_tz": -60, "elapsed": 888, "user": {"displayName": "Sl G", "photoUrl": "", "userId": "13241675469300863486"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
cat_feats = [x for x in df.columns if SUFFIX_CAT in x]
cat_feats = [x for x in cat_feats if 'price' not in x]
len(cat_feats)
# + id="8pO2kIlFaJYY" colab_type="code" colab={}
def run_model(model, feats):
X = df[feats].values
y = df['price_value'].values
scores = cross_val_score(model, X, y, cv=3, scoring='neg_mean_absolute_error')
return np.mean(scores), np.std(scores)
# + id="Vxa4enTlcCXu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="a99d1318-a0ba-4ae0-e0d1-3e2bf5cf0532" executionInfo={"status": "ok", "timestamp": 1583456436885, "user_tz": -60, "elapsed": 5354, "user": {"displayName": "Sl G", "photoUrl": "", "userId": "13241675469300863486"}}
run_model(DecisionTreeRegressor(max_depth=5), cat_feats)
# + [markdown] id="JENx8YxtnSo0" colab_type="text"
# **Random Forest**
# + id="vXsww_3viVLH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="b7380413-7828-48c0-a27a-feb39fabc690" executionInfo={"status": "ok", "timestamp": 1583456557847, "user_tz": -60, "elapsed": 111272, "user": {"displayName": "Sl G", "photoUrl": "", "userId": "13241675469300863486"}}
model = RandomForestRegressor(max_depth=5, n_estimators=50, random_state=0)
run_model(model, cat_feats)
# + id="Id49vG_qmd9v" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 84} outputId="99f7126d-f5d0-4afb-a8ec-5146f863a87f" executionInfo={"status": "ok", "timestamp": 1583456684854, "user_tz": -60, "elapsed": 59367, "user": {"displayName": "Sl G", "photoUrl": "", "userId": "13241675469300863486"}}
xgb_params = {
'max_depth': 5,
'n_estimators': 50,
'learning_rate': 0.1,
'seed': 0
}
run_model(xgb.XGBRegressor(**xgb_params), cat_feats)
# + id="YmZpv-keoYBI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 403} outputId="f066cbb2-919c-4890-8b7d-07d1efd4571d" executionInfo={"status": "ok", "timestamp": 1583457073248, "user_tz": -60, "elapsed": 354301, "user": {"displayName": "Sl G", "photoUrl": "", "userId": "13241675469300863486"}}
m = xgb.XGBRegressor(max_depth=5, n_estimators=50, learning_rate=0.1, seed=0)
m.fit(X, y)
imp = PermutationImportance(m, random_state=0).fit(X, y)
eli5.show_weights(imp, feature_names=cat_feats)
# + id="IYJoVb-ZwrPZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="98dee69d-b9e4-4da3-e4e8-539f94fc1ae0" executionInfo={"status": "ok", "timestamp": 1583457105355, "user_tz": -60, "elapsed": 991, "user": {"displayName": "Sl G", "photoUrl": "", "userId": "13241675469300863486"}}
len(cat_feats)
# + id="ewUPm0uAw2Ao" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 84} outputId="3187cfcf-21ec-4c57-e5d9-1106ae227cd1" executionInfo={"status": "ok", "timestamp": 1583457120118, "user_tz": -60, "elapsed": 13629, "user": {"displayName": "Sl G", "photoUrl": "", "userId": "13241675469300863486"}}
feats = ['param_napęd_cat', 'param_rok-produkcji_cat', 'param_stan_cat', 'param_skrzynia-biegów_cat', 'param_faktura-vat_cat', 'param_moc_cat', 'param_marka-pojazdu_cat', 'feature_kamera-cofania_cat', 'param_typ_cat', 'param_pojemność-skokowa_cat', 'seller_name_cat', 'feature_wspomaganie-kierownicy_cat', 'param_model-pojazdu_cat', 'param_wersja_cat', 'param_kod-silnika_cat', 'feature_system-start-stop_cat', 'feature_asystent-pasa-ruchu_cat', 'feature_czujniki-parkowania-przednie_cat', 'feature_łopatki-zmiany-biegów_cat', 'feature_regulowane-zawieszenie_cat']
run_model(xgb.XGBRegressor(**xgb_params), feats)
# + id="Y_1cwn2zy4Ea" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 84} outputId="ffd69f68-d87d-4753-d195-d09dee4a7ce7" executionInfo={"status": "ok", "timestamp": 1583457159116, "user_tz": -60, "elapsed": 14164, "user": {"displayName": "Sl G", "photoUrl": "", "userId": "13241675469300863486"}}
df['param_rok-produkcji'] = df['param_rok-produkcji'].map(lambda x: -1 if str(x) == 'None' else int (x))
feats = ['param_napęd_cat', 'param_rok-produkcji', 'param_stan_cat', 'param_skrzynia-biegów_cat', 'param_faktura-vat_cat', 'param_moc_cat', 'param_marka-pojazdu_cat', 'feature_kamera-cofania_cat', 'param_typ_cat', 'param_pojemność-skokowa_cat', 'seller_name_cat', 'feature_wspomaganie-kierownicy_cat', 'param_model-pojazdu_cat', 'param_wersja_cat', 'param_kod-silnika_cat', 'feature_system-start-stop_cat', 'feature_asystent-pasa-ruchu_cat', 'feature_czujniki-parkowania-przednie_cat', 'feature_łopatki-zmiany-biegów_cat', 'feature_regulowane-zawieszenie_cat']
run_model(xgb.XGBRegressor(**xgb_params), feats)
# + id="KbXUcKvS0Kuc" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 218} outputId="2fc42e73-5624-4bd4-b773-c7b84fb327d0" executionInfo={"status": "ok", "timestamp": 1583457233194, "user_tz": -60, "elapsed": 987, "user": {"displayName": "Sl G", "photoUrl": "", "userId": "13241675469300863486"}}
df['param_moc'].map(lambda x: -1 if str(x) == 'None' else int(x.split(' ')[0]))
# + id="zGs7Ib2P1hSs" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 84} outputId="fb67e938-e986-49c6-9336-c27f76d088eb" executionInfo={"status": "ok", "timestamp": 1583457250193, "user_tz": -60, "elapsed": 13802, "user": {"displayName": "Sl G", "photoUrl": "", "userId": "13241675469300863486"}}
df['param_moc'] = df['param_moc'].map(lambda x: -1 if str(x) == 'None' else int(x.split(' ')[0]) )
df['param_rok-produkcji'] = df['param_rok-produkcji'].map(lambda x: -1 if str(x) == 'None' else int (x))
feats = ['param_napęd_cat', 'param_rok-produkcji', 'param_stan_cat', 'param_skrzynia-biegów_cat', 'param_faktura-vat_cat', 'param_moc', 'param_marka-pojazdu_cat', 'feature_kamera-cofania_cat', 'param_typ_cat', 'param_pojemność-skokowa_cat', 'seller_name_cat', 'feature_wspomaganie-kierownicy_cat', 'param_model-pojazdu_cat', 'param_wersja_cat', 'param_kod-silnika_cat', 'feature_system-start-stop_cat', 'feature_asystent-pasa-ruchu_cat', 'feature_czujniki-parkowania-przednie_cat', 'feature_łopatki-zmiany-biegów_cat', 'feature_regulowane-zawieszenie_cat']
run_model(xgb.XGBRegressor(**xgb_params), feats)
# + id="fphZprbz2UHX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 84} outputId="5035fa7c-b4cb-450c-870d-08691fd8667b" executionInfo={"status": "ok", "timestamp": 1583457385350, "user_tz": -60, "elapsed": 14033, "user": {"displayName": "Sl G", "photoUrl": "", "userId": "13241675469300863486"}}
df['param_pojemność-skokowa'] = df['param_pojemność-skokowa'].map(lambda x: -1 if str(x) == 'None' else int (str(x).split('cm')[0].replace(' ', '')) )
feats = ['param_napęd_cat', 'param_rok-produkcji', 'param_stan_cat', 'param_skrzynia-biegów_cat', 'param_faktura-vat_cat', 'param_moc', 'param_marka-pojazdu_cat', 'feature_kamera-cofania_cat', 'param_typ_cat', 'param_pojemność-skokowa', 'seller_name_cat', 'feature_wspomaganie-kierownicy_cat', 'param_model-pojazdu_cat', 'param_wersja_cat', 'param_kod-silnika_cat', 'feature_system-start-stop_cat', 'feature_asystent-pasa-ruchu_cat', 'feature_czujniki-parkowania-przednie_cat', 'feature_łopatki-zmiany-biegów_cat', 'feature_regulowane-zawieszenie_cat']
run_model(xgb.XGBRegressor(**xgb_params), feats)
# + id="rZtiogH1PEAW" colab_type="code" colab={}
|
day4.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import trimesh
import numpy as np
from shapely.geometry import LineString
# %pylab inline
# load the mesh from filename
# file objects are also supported
mesh = trimesh.load_mesh('../models/featuretype.STL')
# get a single cross section of the mesh
slice = mesh.section(plane_origin=mesh.centroid,
plane_normal=[0,0,1])
# the section will be in the original mesh frame
slice.show()
# we can move the 3D curve to a Path2D object easily
slice_2D, to_3D = slice.to_planar()
slice_2D.show()
# if we wanted to take a bunch of parallel slices, like for a 3D printer
# we can do that easily with the section_multiplane method
# we're going to slice the mesh into evenly spaced chunks along z
# this takes the (2,3) bounding box and slices it into [minz, maxz]
z_extents = mesh.bounds[:,2]
# slice every .125 model units (eg, inches)
z_levels = np.arange(*z_extents, step=.125)
# find a bunch of parallel cross sections
sections = mesh.section_multiplane(plane_origin=mesh.bounds[0],
plane_normal=[0,0,1],
heights=z_levels)
sections
# summing the array of Path2D objects will put all of the curves
# into one Path2D object, which we can plot easily
combined = np.sum(sections)
combined.show()
# if we want to intersect a line with this 2D polygon, we can use shapely methods
polygon = slice_2D.polygons_full[0]
# intersect line with one of the polygons
hits = polygon.intersection(LineString([[-4,-1], [3,0]]))
# check what class the intersection returned
hits.__class__
# we can plot the intersection (red) and our original geometry(black and green)
for h in hits:
plt.plot(*h.xy, color='r')
slice_2D.show()
# the medial axis is available for closed Path2D objects
(slice_2D + slice_2D.medial_axis()).show()
|
examples/section.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
star_wars = pd.read_csv('star_wars.csv', encoding='ISO-8859-1')
star_wars.head(10)
# -
star_wars.columns
star_wars['Which of the following Star Wars films have you seen? Please select all that apply.'].value_counts()
# +
yn_tf = {
'Yes': True,
'No': False
}
star_wars['Have you seen any of the 6 films in the Star Wars franchise?'] = star_wars['Have you seen any of the 6 films in the Star Wars franchise?'].map(yn_tf)
star_wars['Do you consider yourself to be a fan of the Star Wars film franchise?'] = star_wars['Do you consider yourself to be a fan of the Star Wars film franchise?'].map(yn_tf)
# -
star_wars.loc[:, ['Have you seen any of the 6 films in the Star Wars franchise?', 'Do you consider yourself to be a fan of the Star Wars film franchise?']]
# +
import numpy as np
seen_cols = ['Which of the following Star Wars films have you seen? Please select all that apply.',
'Unnamed: 4',
'Unnamed: 5',
'Unnamed: 6',
'Unnamed: 7',
'Unnamed: 8']
movies = ['Star Wars: Episode I The Phantom Menace',
'Star Wars: Episode II Attack of the Clones',
'Star Wars: Episode III Revenge of the Sith',
'Star Wars: Episode IV A New Hope',
'Star Wars: Episode V The Empire Strikes Back',
'Star Wars: Episode VI Return of the Jedi']
for i in range(0, 6):
star_wars[seen_cols[i]] = star_wars[seen_cols[i]].map({movies[i] : True, np.NaN : False})
star_wars.rename(columns={seen_cols[i] : 'seen_' + str(i+1)}, inplace=True)
star_wars.iloc[:,3:9]
# -
star_wars.iloc[:,9:15]
star_wars.iloc[:,9:15].astype(float)
star_wars.iloc[:,9:15].info()
star_wars.rename(columns={'Please rank the Star Wars films in order of preference with 1 being your favorite film in the franchise and 6 being your least favorite film.' : 'Star Wars: Episode I The Phantom Menace',
'Unnamed: 10' : 'Star Wars: Episode II Attack of the Clones',
'Unnamed: 11' : 'Star Wars: Episode III Revenge of the Sith',
'Unnamed: 12' : 'Star Wars: Episode IV A New Hope',
'Unnamed: 13' : 'Star Wars: Episode V The Empire Strikes Back',
'Unnamed: 14' : 'Star Wars: Episode VI Return of the Jedi'
}, inplace=True)
star_wars.iloc[:, 9:15]
# +
import matplotlib.pyplot as plt
# %matplotlib inline
star_wars.iloc[:, 9:15].mean().plot.bar()
plt.show()
# -
# ### Observations
# Above is a bar plot of the average rankings per Star Wars movie where the six movies were ranked from 1-6 where 1 is the most favorite and 6 the least favorite. Because lower numbers rank higher, lower means indicate higher ranked movies.
#
# According to the surveys, the movies rank from most favorite to least favorite:
# 1. Star Wars: Episode V The Empire Strikes Back
# 2. Star Wars: Episode VI Return of the Jedi
# 3. Star Wars: Episode IV A New Hope
# 4. Star Wars: Episode I The Phantom Menace
# 5. Star Wars: Episode II Attack of the Clones
# 6. Star Wars: Episode III Revenge of the Sith
#
# The original Star Wars movies (Episodes 4-6) rank higher than the prequels (Episodes 1-3). It seems that the original episodes are favored more than newer movies among these surveyees.
#
# The highest-ranked Star Wars movie according to the FiveThirtyEight surveys appears to be "Star Wars: Episode V The Empire Strikes Back".
plt.bar(['Star Wars ' + str(i) for i in range(1,7)], star_wars.iloc[:,3:9].sum())
plt.xticks(rotation=90)
# ### Observations
# The original Star Wars movies (Episodes 4-6) have a higher view count than the newer movies, as per the earlier observation of the rankings.
#
# We've found that Episode 5 was the highest ranking Star Wars movies based on the survey answers, and it is also the most viewed movie in the survey. It makes sense that the highest ranking movie would also have the highest view count. For one, you wouldn't rank a movie you haven't watched.
sw_fan_t = star_wars[star_wars['Do you consider yourself to be a fan of the Star Wars film franchise?'] == True]
sw_fan_f = star_wars[star_wars['Do you consider yourself to be a fan of the Star Wars film franchise?'] == False]
sw_fan_t
# ## Star Wars fans most viewed and highest ranked movies
# +
fig, ax = plt.subplots(nrows=3, ncols=2, figsize=(10,10))
fig.tight_layout(pad=3)
ax[0,0].bar(['Ep. ' + str(i) for i in range(1,7)], sw_fan_t.iloc[:, 9:15].mean())
ax[0,0].text(2,5,'Rankings (Fan)')
ax[0,1].bar(['Ep. ' + str(i) for i in range(1,7)], sw_fan_t.iloc[:, 3:9].sum())
ax[0,1].text(2,600,'Views (Fan)')
ax[1,0].bar(['Ep. ' + str(i) for i in range(1,7)], sw_fan_f.iloc[:, 9:15].mean())
ax[1,0].text(2,4.75,'Rankings (Not Fan)')
ax[1,1].bar(['Ep. ' + str(i) for i in range(1,7)], sw_fan_f.iloc[:, 3:9].sum())
ax[1,1].text(2,250,'Views (Not Fan)')
ax[2,0].bar(['Ep. ' + str(i) for i in range(1,7)], star_wars.iloc[:, 9:15].mean())
ax[2,0].text(2,4.75,'Rankings (All)')
ax[2,1].bar(['Ep. ' + str(i) for i in range(1,7)], star_wars.iloc[:, 3:9].sum())
ax[2,1].text(2,850,'Views (All)')
plt.show()
# -
print(sw_fan_t.shape[0]/star_wars.shape[0])
print(sw_fan_f.shape[0]/star_wars.shape[0])
sw_fan_f.iloc[:, 9:15].mean().sort_values()
# ### Observations
# Displayed above are the rankings (left column) and view counts (right column) for those indicated as fans, not fans, and either (all surveyees- fans or not fans). The graphs in the third row indicated by (All) are the same rankings and view count graphs we displayed earlier, for comparison between the fan and not fan groups.
#
# The majority of those who took the surveys are fans- they make up 46.5% of the survey responses as opposed to non-fans being 23.9% and those undeclared being 29.6%.
#
# Among declared fans, the original episodes (episodes 4-6) are ranked much higher than the newer prequels (episodes 1-3) compared to non-fans. The distribution of views among the 6 movies are much more even compared to non-fans and the overall views plot- this makes sense as declared fans of the series will likely watch most, if not all, movies of their franchise compared to a non-fan.
#
# Among declared non-fans, the original episodes did not necessarily rank higher than the newer ones, though Episode V is still their highest ranked movie, as per fans and overall rankings. The ranking in terms of view count is the same compared to fans and overall view counts, but are significantly lesser for Episodes 2-4. The most viewed movie among non-fans is still Episode V. This is likely due to non-fans primarily watching higher-ranked and more popular episodes. The larger differences in view counts between the movies are likely due to non-fans cherry picking which to watch and likely not having an urgency to watch all 6 movies.
#
# The rankings and view counts for declared fans are very similar in trend to the overall rankings and view counts, although slightly more evenly distributed, likely due to fans being the majority of the surveyees. The rankings among non-fans differ from the overall ranking and have no clear bias towards originals vs. newer movies. The view counts, however, follow the same pattern as the overall view count. Both the rankings and view counts among non-fans have a more uneven distribution than overall and fan rankings and view counts.
#
# However, among fans, non-fans, and either fans or non-fans, Star Wars Episode V reigns the highest ranking and most viewed episode, with Episode III being the lower ranking and lowest viewed among all groups.
|
Star Wars Survey.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
from openpyxl import load_workbook
from bs4 import BeautifulSoup
from selenium import webdriver
from time import sleep
import csv
from random import randint
import json, io
from selenium.webdriver.common.by import By
from selenium.webdriver.common.keys import Keys
from selenium.webdriver.support.ui import Select
from selenium.webdriver.support.ui import WebDriverWait
from selenium.common.exceptions import TimeoutException
from selenium.webdriver.support import expected_conditions as EC
from selenium.common.exceptions import NoSuchElementException
from selenium.common.exceptions import NoAlertPresentException
from selenium.webdriver.common.action_chains import ActionChains
import urllib
import urllib3
import requests
import json, io
from bs4 import BeautifulSoup
urllib3.disable_warnings()
header = {'User-Agent':'Mozilla/6.0'}
chrome_options = webdriver.ChromeOptions()
chrome_options.add_argument('--user-agent="Mozilla/6.0')
chrome_options.add_argument("user-data-dir=selenium")
driver = webdriver.Chrome(chrome_options=chrome_options, executable_path=r'chromedriver.exe')
# cookies = json.load(open('cookiesdict.txt'))
# for cookie in cookies:
# driver.add_cookie(cookie)
# -
driver.get('https://webcache.gmc-uk.org/gmccgs_enu/start.swe')
driver.find_elements_by_tag_name('nobr')
driver.find_elements_by_name('s_1_1_0_0')
so1.find_all('tbody')
so1=BeautifulSoup(driver.page_source, 'lxml')
so1
|
gmc-uk/gmc.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Measure Watson Assistant Performance
# 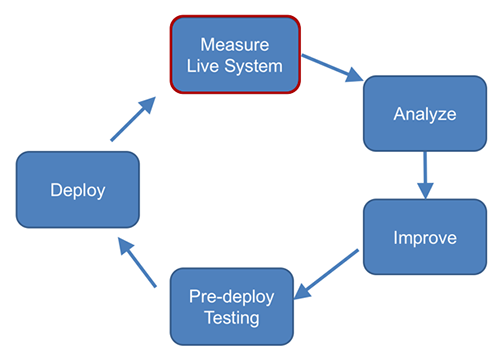
#
# ## Introduction
#
# This notebook demonstrates how to setup automated metrics that help you measure, monitor, and understand the behavior of your Watson Assistant system.As described in <a href="https://github.com/watson-developer-cloud/assistant-improve-recommendations-notebook/raw/master/notebook/IBM%20Watson%20Assistant%20Continuous%20Improvement%20Best%20Practices.pdf" target="_blank" rel="noopener noreferrer">Watson Assistant Continuous Improvement Best Practices</a>, this is the first step of your continuous improvement process. The goal of this step is to understand where your assistant is doing well vs where it isn’t and to potentially focus your improvement effort to one of the problem areas identified. We define two measures to achieve this goal: **Coverage** and **Effectiveness**.
#
# - **Coverage** is the portion of total user messages your assistant is attempting to respond to.
#
# - **Effectiveness** refers to how well your assistant is handling the conversations it is attempting to respond to.
#
# The pre-requisite for running this notebook is Watson Assistant (formerly Watson Conversation). This notebook assumes familiarity with Watson Assistant and concepts such as workspaces, intents and training examples.
#
# ### Programming language and environment
# Some familiarity with Python is recommended. This notebook runs on Python 3.5 with Default Python 3.5 XS environment.
# ***
# ## Table of contents
# 1. [Configuration and setup](#setup)<br>
# 1.1 [Import and apply global CSS styles](#css)<br>
# 1.2 [Install required Python libraries](#python)<br>
# 1.3 [Import functions used in the notebook](#function)<br>
# 2. [Load and format data](#load)<br>
# 2.1 [Option one: from a Watson Assistant instance](#load_remote)<br>
# 2.2 [Option two: from JSON files](#load_local)<br>
# 2.3 [Format the log data](#format_data)<br>
# 3. [Define coverage and effectiveness metrics](#set_metrics)<br>
# 3.1 [Customize coverage](#set_coverage)<br>
# 3.2 [Customize effectiveness](#set_effectiveness)<br>
# 4. [Calculate overall coverage and effectiveness](#overall)<br>
# 4.1 [Calculate overall metrics](#overall1)<br>
# 4.2 [Display overall results](#overall2)<br>
# 5. [Analyze coverage](#msg_analysis)<br>
# 5.1 [Display overall coverage](#msg_analysis1)<br>
# 5.2 [Calculate coverage over time](#msg_analysis2)<br>
# 6. [Analyze effectiveness](#conv_analysis)<br>
# 6.1 [Generate excel file and upload to our project](#conv_analysis1)<br>
# 6.2 [Plot breakdown by effectiveness graph](#conv_analysis2)<br>
# 7. [Root cause analysis of non coverage](#root_cause)<br>
# 8. [Summary and next steps](#summary)<br>
# <a id="setup"></a>
# ## 1. Configuration and Setup
#
# In this section, we add data and workspace access credentials, import required libraries and functions.
# ### <a id="css"></a> 1.1 Import and apply global CSS styles
from IPython.display import HTML
# !curl -O https://raw.githubusercontent.com/watson-developer-cloud/assistant-improve-recommendations-notebook/master/src/main/css/custom.css
HTML(open('custom.css', 'r').read())
# ### <a id="python"></a> 1.2 Install required Python libraries
# +
# install watson-developer-cloud python SDK
# After running this cell once, comment out the following code. Packages only need to be installed once.
# !pip3 install --upgrade "watson-developer-cloud>=1.0,<2.0";
# Import required libraries
import pandas as pd
import matplotlib.pyplot as plt
import json
from pandas.io.json import json_normalize
from watson_developer_cloud import AssistantV1
import matplotlib.dates as mdates
import re
from IPython.display import display
# -
# ### <a id="function"></a> 1.3 Import functions used in the notebook
# +
# Import function module files
# !curl -O https://raw.githubusercontent.com/watson-developer-cloud/assistant-improve-recommendations-notebook/master/src/main/python/cos_op.py
# !curl -O https://raw.githubusercontent.com/watson-developer-cloud/assistant-improve-recommendations-notebook/master/src/main/python/watson_assistant_func.py
# !curl -O https://raw.githubusercontent.com/watson-developer-cloud/assistant-improve-recommendations-notebook/master/src/main/python/visualize_func.py
# !curl -O https://raw.githubusercontent.com/watson-developer-cloud/assistant-improve-recommendations-notebook/master/src/main/python/computation_func.py
# Import the visualization related functions
from visualize_func import make_pie
from visualize_func import coverage_barh
from visualize_func import width_bar
# Import Cloud Object Storage related functions
from cos_op import generate_link
from cos_op import generate_excel_measure
# Import Watson Assistant related functions
from watson_assistant_func import get_logs_jupyter
# Import Dataframe computation related functions
from computation_func import get_effective_df
from computation_func import get_coverage_df
from computation_func import chk_is_valid_node
from computation_func import format_data
# -
# ## <a id="load"></a> 2. Load and format data
# ### <a id="load_remote"></a> 2.1 Option one: from a Watson Assistant instance
#
# #### 2.1.1 Add Watson Assistant configuration
#
# Provide your Watson Assistant credentials and the workspace id that you want to fetch data from.
#
# - For more information about obtaining Watson Assistant credentials, see [Service credentials for Watson services](https://console.bluemix.net/docs/services/watson/getting-started-credentials.html#creating-credentials).
# - API requests require a version parameter that takes a date in the format version=YYYY-MM-DD. For more information about version, see [Versioning](https://www.ibm.com/watson/developercloud/assistant/api/v1/curl.html?curl#versioning).
#
# +
# Provide credentials to connect to assistant
creds = {'username':'YOUR_USERNAME',
'password':'<PASSWORD>',
'version':'2018-05-03'}
# Connect to Watson Assistant
conversation = AssistantV1(username=creds['username'],
password=<PASSWORD>['password'],
version=creds['version'])
# -
# #### 2.1.2 Fetch and load a workspace
#
# Fetch the workspace for the workspace id given in `workspace_id` variable.
# +
# Provide the workspace id you want to analyze
workspace_id = ''
if len(workspace_id) > 0:
# Fetch the worksapce info. for the input workspace id
workspace = conversation.get_workspace(workspace_id = workspace_id, export=True)
# Store the workspace details in a dataframe
df_workspace = json_normalize(workspace)
# Get all intents present in current version of workspace
workspace_intents= [intent['intent'] for intent in df_workspace['intents'].values[0]]
# Get all dialog nodes present in current version of workspace
workspace_nodes= pd.DataFrame(df_workspace['dialog_nodes'].values[0])
# Mark the workspace loaded
workspace_loaded = True
else:
workspace_loaded = False
# -
# #### 2.1.3 Fetch and load workspace logs
#
# Fetch the logs for the workspace id given in `workspace_id` variable. Any necessary filter can be specified in the `filter` variable.<br>
# Note that if the logs were already fetched in a previous run, it will be read from the a cache file.
if len(workspace_id) > 0:
# Filter to be applied while fetching logs, e.g., removing empty input 'meta.summary.input_text_length_i>0', 'response_timestamp>=2018-09-18'
filter = 'meta.summary.input_text_length_i>0'
# Send this info into the get_logs function
workspace_creds={'sdk_object':conversation, 'ws_id':workspace['workspace_id'], 'ws_name':workspace['name']}
# Fetch the logs for the workspace
df = get_logs_jupyter(num_logs=10000, log_list=[], workspace_creds=workspace_creds, log_filter=filter)
# Mark the logs loaded
logs_loaded = True
else:
logs_loaded = False
# ### <a id="load_local"></a> 2.2 Option two: from JSON files
#
# #### 2.2.1 Load a workspace JSON file
# +
if not workspace_loaded:
# The following code is for using demo workspace
import requests
print('Loading workspace data from Watson developer cloud Github repo ... ', end='')
workspace_data = requests.get("https://raw.githubusercontent.com/watson-developer-cloud/assistant-improve-recommendations-notebook/master/notebook/data/workspace.json").text
print('completed!')
df_workspace = json_normalize(json.loads(workspace_data))
# # Specify a workspace JSON file
# workspace_file = 'workspace.json'
# # Store the workspace details in a dataframe
# df_workspace = json_normalize(json.load(open(workspace_file)))
# Get all intents present in current version of workspace
workspace_intents = [intent['intent'] for intent in df_workspace['intents'].values[0]]
# Get all dialog nodes present in current version of workspace
workspace_nodes = pd.DataFrame(df_workspace['dialog_nodes'].values[0])
# -
# #### 2.2.2 Load a log JSON file
# +
if not logs_loaded:
# The following code is for using demo logs
import requests
print('Loading demo log data from Watson developer cloud Github repo ... ', end='')
log_raw_data = requests.get("https://raw.githubusercontent.com/watson-developer-cloud/assistant-improve-recommendations-notebook/master/notebook/data/sample_logs.json").text
print('completed!')
df = pd.DataFrame.from_records(json.loads(log_raw_data))
# # Specify a log JSON file
# log_file = 'sample_logs.json'
# # Create a dataframe for logs
# df = pd.DataFrame.from_records(json.load(open(log_file)))
# -
# ### <a id="format_data"></a> 2.3 Format the log data
# Format the logs data from the workspace
df_formated = format_data(df)
# <a id="set_metrics"></a>
# ## 3. Define effectiveness and coverage metrics
# As described in Watson Assistant Continuous Improvement Best Practices, **Effectiveness** and **Coverage** are the two measures that provide a reliable understanding of your assistant’s overall performance. Both of the two measures are customizable based on your preferences. In this section, we provide a guideline for setting each of them.
# ### <a id="set_coverage"></a> 3.1 Customize coverage
#
# Coverage measures your Watson Assistant system at the utterance level. You may include automated metrics that help identify utterences that your service is not answering. Example metrics include:
#
# - Confidence threshold
# - Dialog information
#
# For Confidence threshold, you can set a threshold to include utterances with confidence values below this threshold. For more information regarding Confidence, see [Absolute scoring](https://console.bluemix.net/docs/services/conversation/intents.html#absolute-scoring-and-mark-as-irrelevant).
#
# For Dialog information, you can specify what the notebook should look for in your logs to determine that a message is not covered by your assistant.
#
# - Use the node_ids list to include the identifiers of any dialog nodes you've used to model that a message is out of scope.
# - Similarly, use the node_names list to include any dialog nodes.
# - Use node_conditions for dialog conditions that indicate a message is out of scope.
#
# Note that these lists are treated as "OR" conditions - any occurrence of any of them will signify that a message is not covered.
#
# __Where to find node id, node name, and node condition__?
#
# You can find the values of these variables from your workspace JSON file based on following mappings.
#
# - node id: `dialog_node`
# - node name: `title`
# - node condition: `conditions`
#
# You can also find node name, and node condition in your workspace dialog editor. For more information, see [Dialog Nodes].(https://console.bluemix.net/docs/services/conversation/dialog-overview.html#dialog-nodes)
#
# Below we provide example code for identifying coverage based on confidence and dialog node.
# +
#Specify the confidence threhold you want to look for in the logs
confidence_threshold = .20
# Add coverage node ids, if any, to list
node_ids = ['node_1_1467910920863', 'node_1_1467919680248']
# Add coverage node names, if any, to list
node_names = []
# Add coverage node conditions, if any, to list
node_conditions = ['#out_of_scope || #off_topic', 'anything_else']
# Check if the dialog nodes are present in the current version of workspace
df_coverage_nodes = chk_is_valid_node(node_ids, node_names, node_conditions, workspace_nodes)
df_coverage_nodes
# -
# ### <a id="set_effectiveness"></a> 3.2 Customize effectiveness
#
# Effectiveness measures your Watson Assistant system at the conversation level. You may include automated metrics that help identify problematic conversations. Example metrics include:
#
# - Escalations to live agent: conversations escalated to a human agent for quality reasons.
# - Poor NPS: conversations that received a poor NPS (Net Promoter Score), or other explicit user feedback.
# - Task not completed: conversations failed to complete the task the user was attempting.
# - Implicit feedback: conversations containing implicit feedback that suggests failure, such as links provided not being clicked.
#
# Below we provide example code for identifying escalation based on intents and dialog information.
# #### <a id="set_escalation_intent"></a> 3.2.1 Specify intents to identify escalations
# If you have specific intents that point to escalation or any other effectiveness measure, specify those in `chk_effective_intents` lists below. <br>
# **Note:** If you don't have specific intents to capture effectiveness, leave chk_effective_intents list empty.
# +
# Add your escalation intents to the list
chk_effective_intents=['connect_to_agent']
# Store the intents in a dataframe
df_chk_effective_intents = pd.DataFrame(chk_effective_intents, columns = ['Intent'])
# Add a 'valid' flag to the dataframe
df_chk_effective_intents['Valid']= True
# Checking the validity of the specified intents. Look out for the `valid` column in the table displayed below.
for intent in chk_effective_intents:
# Check if intent is present in workspace
if intent not in workspace_intents:
# If not present, mark it as 'not valid'
df_chk_effective_intents.loc[df_chk_effective_intents['Intent']==intent,['Valid']] = False
# Remove intent from the chk_effective_intents list
chk_effective_intents.remove(intent)
# Display intents and validity
df_chk_effective_intents
# -
# #### <a id="set_escalation_dialog"></a> 3.2.2 Specify dialog nodes to identify escalations
# If you have specific dialog nodes that point to escalation or any other effectiveness measure, you can automated capture them based on three variables: node id, node name, and node condition.
#
# - Use the node_ids list to include the identifiers of any dialog nodes you've used to model that a message indicates an escalation.
# - Similarly, use the node_names list to include dialog nodes.
# - Use node_conditions for dialog conditions that indicate a message is out of scope.
#
# Note that these lists are treated as "OR" conditions - any occurrence of any of them will signify that a message is not covered.
#
# __Where to find node id, node name, and node condition__?
#
# You can find the values of these variables from your workspace JSON file based on following mappings.
#
# - node id: `dialog_node`
# - node name: `title`
# - node condition: `conditions`
#
# You can also find node name, and node condition in your workspace dialog editor. For more information, see [Dialog Nodes](https://console.bluemix.net/docs/services/conversation/dialog-overview.html#dialog-nodes).
#
# **Note:** If your assistant does not incorporate escalations and you do not have any other automated conversation-level quality metrics to identify problematic conversations (e.g., poor NPS, task not completed), you can simply track coverage and average confidence over a recent sample of your entire production logs. Leave an empty list for node_ids, node_names and node_conditions.
# +
# Add effectiveness node ids, if any, to list
node_ids = []
# Add effectiveness node names, if any, to list
node_names = ['not_trained']
# Add effectiveness node conditions, if any, to list
node_conditions = ['#connect_to_agent', '#answer_not_helpful']
# If your assistant does not incorporate escalations and you do not have any other automated conversation-level quality metrics, uncomment lines below
# node_ids = []
# node_names = []
# node_conditions = []
# Check if the dialog nodes are present in the current version of workspace
df_chk_effective_nodes = chk_is_valid_node(node_ids, node_names, node_conditions, workspace_nodes)
df_chk_effective_nodes
# -
# ## 4. Calculate overall coverage and effectiveness<a id="overall"></a>
# The combination of effectiveness and coverage is very powerful for diagnostics.
# If your effectiveness and coverage metrics are high, it means that your assistant is responding to most inquiries and responding well. If either effectiveness or coverage are low, the metrics provide you with the information you need to start improving your assistant.
#
# ### 4.1 Calculate overall metrics<a id="overall1"></a>
# +
df_formated_copy = df_formated.copy(deep = True)
# Mark if a message is covered and store results in df_coverage dataframe
df_coverage = get_coverage_df(df_formated_copy , df_coverage_nodes, confidence_threshold)
# Mark if a conversation is effective and store results in df_coverage dataframe
df_effective = get_effective_df(df_formated_copy, chk_effective_intents, df_chk_effective_nodes, filter_non_intent_node=True, workspace_nodes=workspace_nodes)
# Calculate average confidence
avg_conf = float("{0:.2f}".format(df_coverage[df_coverage['Covered']==True]['response.top_intent_confidence'].mean()*100))
# Calculate coverage
coverage = float("{0:.2f}".format((df_coverage['Covered'].value_counts().to_frame()['Covered'][True]/df_coverage['Covered'].value_counts().sum())*100))
# Calculate effectiveness
effective_perc = float("{0:.2f}".format((df_effective.loc[df_effective['Escalated_conversation']==False]['response.context.conversation_id'].nunique()/df_effective['response.context.conversation_id'].nunique())*100))
# Plot pie graphs for coverage and effectiveness
coverage_pie = make_pie(coverage, "Percent of total messages covered")
effective_pie = make_pie(effective_perc, 'Percent of non-escalated conversations')
# -
# Messages to be displayed with coverage and effectiveness metrics are given below
# +
# Messages to be displayed with effectiveness and coverage
coverage_msg = '<h2>Coverage</h2></br>A message that is not covered would either be a \
message your assistant responded to with some form \
of “I’m not trained” or that it immediately handed over \
to a human agent without attempting to respond'
effectiveness_msg = '<h2>Effectiveness</h2></br>This notebook provides a list of metrics customers \
can use to assess how effective their assistant is at \
responding to conversation and metrics '
# -
# ### 4.2 Display overall results<a id="overall2"></a>
#
# Display the coverage and effectiveness pie charts
HTML('<tr><th colspan="4"><div align="center"><h2>Coverage and Effectiveness<hr/></h2></div></th></tr>\
<tr>\
<td style="width:500px">{c_pie}</td>\
<td style="width:450px"><div align="left"> {c_msg} </div></td>\
<td style="width:500px">{e_pie}</td>\
<td style="width:450px"><div align="left"> {e_msg} </div></td>\
</tr>'
.format(c_pie=coverage_pie, c_msg = coverage_msg, e_pie = effective_pie, e_msg = effectiveness_msg))
# Here, we can see our assistant's coverage and effectiveness. We will have to take a deeper look at both of these metrics to understand the nuances and decide where we should focus next.
#
# Note that the distinction between a user message and a conversation. A conversation in Watson Assistant represents a session of one or more messages from a user and the associated responses returned to the user from the assistant. A conversation includes a Conversation id for the purposes of grouping a sequence of messages and responses.
# <a id="msg_analysis"></a>
# ## 5. Analyze coverage
#
# Here, we take a deeper look at the Coverage of our Watson Assistant.
#
# ### 5.1 Display overall coverage<a id="msg_analysis1"></a>
#
# +
# %matplotlib inline
# Compute the number of conversations in the log
convs = df_coverage['response.context.conversation_id'].nunique()
# Compute the number of messages in the log
msgs = df_coverage['response.context.conversation_id'].size
#Display the results
print('Overall messages\n', "=" * len('Overall messages'), '\nTotal Conversations: ', convs, '\nTotal Messages: ', msgs, '\n\n', sep = '')
#Display the coverage bar chart
display(coverage_barh(coverage, avg_conf, 'Coverage & Average confidence', False))
# -
# Here, we see the percentage of messages covered and their average confidence. Now, let us take a look at the coverage over time.
#
# ### 5.2 Calculate coverage over time<a id="msg_analysis2"></a>
#
# +
# Make a copy of df_coverage dataframe
df_Tbot_raw1 = df_coverage.copy(deep=True)
# Group by date and covered and compute the count
covered_counts = df_Tbot_raw1[['Date','Covered']].groupby(['Date','Covered']).agg({'Covered': 'count'})
# Convert numbers to percentage
coverage_grp = covered_counts.groupby(level=0).apply(lambda x:round(100 * x / float(x.sum()),2)).rename(columns = {'Covered':'Coverage'}).reset_index()
# Get only covered messages
coverage_time = coverage_grp[coverage_grp['Covered']==True].reset_index(drop = True)
# Determine the number of xticks required
xticks = [d for d in coverage_time['Date']]
# +
# Plot the coverage over time graph
fig, ax = plt.subplots(figsize=(30,8))
# Format the date on x-axis
ax.xaxis.set_major_formatter(mdates.DateFormatter('%Y-%m-%d'))
ax.xaxis_date()
ax.set_xticks(xticks)
# Plot a line plot if there are more data points
if len(coverage_time) >1:
ax.plot_date(coverage_time['Date'], coverage_time['Coverage'], fmt = '-', color = '#4fa8f6', linewidth=6)
# Plot a scatter plot if there is only one date on x-axis
else:
ax.plot_date(coverage_time['Date'], coverage_time['Coverage'], color = '#4fa8f6', linewidth=6)
# Set axis labels and title
ax.set_xlabel("Time", fontsize=20, fontweight='bold')
ax.set_ylabel("Coverage %", fontsize=20, fontweight='bold')
ax.set_title('Coverage over time', fontsize=25, fontweight = 'bold')
ax.tick_params(axis='both', labelsize=15)
# Hide the right and top spines
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
# -
# **Note:** Compare the coverage over time with any major updates to your assistant, to see if the changes affected the performance.
# <a id="conv_analysis"></a>
# ## 6. Analyze effectiveness
# Here, we will take a deeper look at effectiveness of the assistant
# +
# Get the escalated conversations
df_effective_true = df_effective.loc[df_effective['Escalated_conversation']==True]
# Get the non-escalated conversations
df_not_effective = df_effective.loc[df_effective['Escalated_conversation']==False]
# Calculate percentage of escalated conversations
ef_escalated = float("{0:.2f}".format(100-effective_perc))
# Calculate coverage and non-coverage in escalated conversations
if len(df_effective_true) > 0:
escalated_covered = float("{0:.2f}".format((df_effective_true['Covered'].value_counts().to_frame()['Covered'][True]/df_effective_true['Covered'].value_counts().sum())*100))
escalated_not_covered = float("{0:.2f}".format(100- escalated_covered))
else:
escalated_covered = 0
escalated_not_covered = 0
# Calculate coverage and non-coverage in non-escalated conversations
if len(df_not_effective) > 0:
not_escalated_covered = float("{0:.2f}".format((df_not_effective['Covered'].value_counts().to_frame()['Covered'][True]/df_not_effective['Covered'].value_counts().sum())*100))
not_escalated_not_covered = float("{0:.2f}".format(100 - not_escalated_covered))
else:
not_escalated_covered = 0
not_escalated_not_covered = 0
# Calculate average confidence of escalated conversations
if len(df_effective_true) > 0:
esc_avg_conf = float("{0:.2f}".format(df_effective_true[df_effective_true['Covered']==True]['response.top_intent_confidence'].mean()*100))
else:
esc_avg_conf = 0
# Calculate average confidence of non-escalated conversations
if len(df_not_effective) > 0:
not_esc_avg_conf = float("{0:.2f}".format(df_not_effective[df_not_effective['Covered']==True]['response.top_intent_confidence'].mean()*100))
else:
not_esc_avg_conf = 0
# -
# ### 6.1 Generate excel file and upload to our project<a id="conv_analysis1"></a>
# +
# Copy the effective dataframe
df_excel = df_effective.copy(deep=True)
# Rename columns to generate excel
df_excel = df_excel.rename(columns={'log_id':'Log ID', 'response.context.conversation_id':'Conversation ID',
'response.timestamp':'Response Timestamp',
'request_input':'Utterance Text',
'response_text':'Response Text', 'response.top_intent_intent':'Detected top intent',
'response.top_intent_confidence':'Detected top intent confidence',
'Intent 2 intent': 'Intent 2', 'Intent 2 confidence':'Intent 2 Confidence',
'Intent 3 intent': 'Intent 3', 'Intent 3 confidence':'Intent 3 Confidence',
'response_entities':'Detected Entities', 'Escalated_conversation':'Escalated conversation?',
'Covered':'Covered?', 'Not Covered cause':'Not covered - cause',
'response.output.nodes_visited_s':'Dialog Flow', 'response_dialog_stack':'Dialog stack',
'response_dialog_request_counter':'Dialog request counter', 'response_dialog_turn_counter':'Dialog turn counter'
})
existing_columns = ['Log ID', 'Conversation ID', 'Response Timestamp', 'Customer ID (must retain for delete)',
'Utterance Text', 'Response Text', 'Detected top intent', 'Detected top intent confidence',
'Intent 2', 'Intent 2 Confidence', 'Confidence gap (between 1 and 2)', 'Intent 3', 'Intent 3 Confidence',
'Detected Entities', 'Escalated conversation?', 'Covered?', 'Not covered - cause',
'Dialog Flow', 'Dialog stack', 'Dialog request counter', 'Dialog turn counter']
# Add new columns for annotating problematic logs
new_columns_excel = ['Response Correct (Y/N)?', 'Response Helpful (Y/N)?', 'Root cause (Problem with Intent, entity, dialog)',
'Wrong intent? If yes, put the correct intent. Otherwise leave it blank', 'New intent needed? (A new intent. Otherwise leave blank)',
'Add Utterance to Training data (Y/N)', 'Entity missed? If yes, put the missed entity value. Otherwise leave it blank', 'New entity needed? If yes, put the entity name',
'New entity value? If yes, put the entity value', 'New dialog logic needed?', 'Wrong dialog node? If yes, put the node name. Otherwise leave it blank','No dialog node triggered']
# Add the new columns to the dataframe
df_excel = df_excel.reindex(columns=[*existing_columns, *new_columns_excel], fill_value='')
# +
# Set maximum sampling size
SAMPLE_SIZE = 200
# Set output filename
all_file = 'All.xlsx'
escalated_sample_file = 'Escalated_sample.xlsx'
non_escalated_sample_file = 'NotEscalated_sample.xlsx'
# Generate all covered sample file
df_covered = df_excel[df_excel['Covered?']==True].reset_index(drop=True)
# Generate all not covered sample file
df_not_covered = df_excel[df_excel['Covered?']==False].reset_index(drop=True)
# Convert to Excel format and upload to COS
generate_excel_measure([df_covered,df_not_covered], ['Covered', 'Not_Covered'], filename=all_file, project_io=None)
# Generate escalated and covered sample file
df_escalated_true = df_excel.loc[df_excel['Escalated conversation?']==True]
df_escalated_covered = df_escalated_true[df_escalated_true['Covered?']==True]
if len(df_escalated_covered) > 0:
df_escalated_covered = df_escalated_covered.sample(n=min(len(df_escalated_covered), SAMPLE_SIZE), random_state=1).reset_index(drop=True)
# Generate escalated but not covered sample file
df_escalated_not_covered = df_escalated_true[df_escalated_true['Covered?']==False]
if len(df_escalated_not_covered) > 0:
df_escalated_not_covered = df_escalated_not_covered.sample(n=min(len(df_escalated_not_covered), SAMPLE_SIZE), random_state=1).reset_index(drop=True)
# Covert to Excel format and upload to COS
generate_excel_measure([df_escalated_covered,df_escalated_not_covered], ['Covered', 'Not_Covered'], filename=escalated_sample_file, project_io=None)
# Generate not escalated but covered sample file
df_not_escalated = df_excel.loc[df_excel['Escalated conversation?']==False]
df_not_escalated_covered = df_not_escalated[df_not_escalated['Covered?']==True]
if len(df_not_escalated_covered) > 0:
df_not_escalated_covered = df_not_escalated_covered.sample(n=min(len(df_not_escalated_covered), SAMPLE_SIZE), random_state=1).reset_index(drop=True)
# Generate not escalated and not covered sample file
df_not_escalated_not_covered = df_not_escalated[df_not_escalated['Covered?']==False]
if len(df_not_escalated_not_covered) > 0:
df_not_escalated_not_covered = df_not_escalated_covered.sample(n=min(len(df_not_escalated_not_covered), SAMPLE_SIZE), random_state=1).reset_index(drop=True)
# Covert to Excel format and upload to COS
generate_excel_measure([df_not_escalated_covered,df_not_escalated_not_covered], ['Covered', 'Not_Covered'], filename=non_escalated_sample_file, project_io=None)
# -
# ### 6.2 Plot breakdown by effectiveness graph<a id="conv_analysis2"></a>
# +
#### Get the links to the excels
all_html_link = '<a href={} target="_blank">All.xlsx</a>'.format(all_file)
escalated_html_link = '<a href={} target="_blank">Escalated_sample.xlsx</a>'.format(escalated_sample_file)
not_escalated_html_link = '<a href={} target="_blank">NotEscalated_sample.xlsx</a>'.format(non_escalated_sample_file)
# Embed the links in HTML table format
link_html = '<tr><th colspan="4"><div align="left"><a id="file_list"></a>View the lists here: {} {} {}</div></th></tr>'.format(all_html_link, escalated_html_link, not_escalated_html_link)
if 100-effective_perc > 0:
escalated_bar = coverage_barh(escalated_covered, esc_avg_conf, '', True, 15, width_bar(100-effective_perc))
else:
escalated_bar = ''
if effective_perc > 0:
non_escalated_bar = coverage_barh(not_escalated_covered, not_esc_avg_conf, '' , True , 15,width_bar(effective_perc))
else:
non_escalated_bar = ''
# Plot the results
HTML('<tr><th colspan="4"><div align="left"><h2>Breakdown by effectiveness<hr/></h2></div></th></tr>\
'+ link_html + '<tr><td style= "border-right: 1px solid black; border-bottom: 1px solid black; width : 400px"><div align="left"><strong>Effectiveness (Escalated) </br>\
<font size="5">{ef_escalated}%</strong></font size></br></div></td>\
<td style="width:1000px; height=100;">{one}</td></tr>\
<tr><td style= "border-right: 1px solid black; border-bottom: 1px solid black; width : 400px;"><div align="left"><strong>Effectiveness (Not escalated) </br>\
<font size="5">{effective_perc}%</strong></font size></br></div></td>\
<td style="width:1000px; height=100;border-bottom: 1px solid black;">{two}</td>\
</tr>'.format(ef_escalated= ef_escalated,
one = escalated_bar,
effective_perc = effective_perc,
two = non_escalated_bar))
# -
# You can download all the analyzed data from `All.xlsx`. A sample of escalated and non-escalated conversations are available in `Escalated_sample.xlsx` and `NotEscalated_sample.xlsx` respectively.
#
# <a id="root_cause"></a>
# ## 7. Root cause analysis of non coverage
# Lets us take a look at the reasons for non-coverage of messages
# Count the causes for non-coverage and store results in dataframe
not_covered = pd.DataFrame(df_coverage['Not Covered cause'].value_counts().reset_index())
# Name the columns in the dataframe
not_covered.columns = ['Messages', 'Total']
not_covered
# <a id="summary"></a>
# ## 8. Summary and next steps
#
# The metrics described above help you narrow your immediate focus of improvement. We suggest the following two strategies:
#
# - **Toward improving Effectiveness**
#
# We suggest focusing on a group of problematic conversations, e.g., escalated conversations, then performing a deeper analysis on these conversation as follows. <br>
# 1. Choose to download either the complete conversations ([All.xlsx](#file_list)), or sampled escalated conversations [Escalated_sample.xlsx](#file_list), or non-escalated conversations [NotEscalated_sample.xlsx](#file_list).<br>
# 2. Perform a manual assessment of these conversations.<br>
# 3. Analyze the results using our __Analyze Watson Assistant Effectiveness__ Jupyter Notebook.
#
#
# - **Toward improving Coverage**
#
# For utterances where an intent was found but no response was given. We suggest performing a deeper analysis to identify root causes, e.g., missing entities or lacking of dialog logic.
#
# For utterances where no intent was found, we suggest expanding intent coverage as follows.
#
# 1. Examine utterances from the production log, especially focus on the utterances that are below the confidence (0.2 by default).
# 2. If you set a confidence threshold significantly higher than 0.2, we suggest looking at utterances that are below but close to the threshold.
# 3. Once you select a collection of utterances, intent expansion, you can focus on intent expansion by two methods:
# - One-by-One: examine each utterance to either change to an existing intent or add a new intent.
# - Unsupervised Learning: perform semantic clustering to generate utterance clusters; examine each cluster to decide (1) adding utterances of an existing intent or (2) creating a new intent.
#
# For more information, please check <a href="https://github.com/watson-developer-cloud/assistant-improve-recommendations-notebook/raw/master/notebook/IBM%20Watson%20Assistant%20Continuous%20Improvement%20Best%20Practices.pdf" target="_blank" rel="noopener noreferrer">Watson Assistant Continuous Improvement Best Practices</a>.
# ### <a id="authors"></a>Authors
#
# **<NAME>**, Ph.D. in Computer Science, is an Advisory Software Engineer for IBM Watson AI. Zhe has a research background in Natural Language Processing, Sentiment Analysis, Text Mining, and Machine Learning. His research has been published at leading conferences and journals including ACL and EMNLP.
#
# **<NAME>** is a Data Scientist for IBM Watson AI. Sherin has her graduate degree in Business Intelligence and Data Analytics and has experience in Data Analysis, Warehousing and Machine Learning.
# ### <a id="acknowledgement"></a> Acknowledgement
#
# The authors would like to thank the following members of the IBM Research and Watson Assistant teams for their contributions and reviews of the notebook: <NAME>, <NAME>, <NAME>, <NAME>.
# Copyright © 2018 IBM. This notebook and its source code are released under the terms of the MIT License.
|
notebook/Measure Notebook.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: kgtk-env
# language: python
# name: kgtk-env
# ---
# # Computes Graph and Text Embeddings, Elasticsearch Ready KGTK File and RDF Triples for Blazegraph
#
# This notebook computes the following:
#
# - `complEx` graph embeddings
# - `transE` graph embeddings
# - `BERT` text embeddings
# - `elasticsearch` ready KGTK edge for [KGTK Search](https://kgtk.isi.edu/search/)
# - `elasticsearch` ready KGTK edge file for Table Linker
# - `RDF Triples` to be loaded into blazegraph
#
# Inputs:
#
# - `item_file`: the subset of the `claims_file` consistin of edges for property of data type `wikibase-item`
# - `label_file`, `alias_file` and `description_file` containing labels, aliases and descriptions. It is assume that these files contain the labels, aliases and descriptions of all nodes appearing in the claims file. Users may provide these files for specific languages only.
#
# ### Batch Invocation
# Example batch command. The second argument is a notebook where the output will be stored. You can load it to see progress.
#
# ```
# papermill Embeddings-Elasticsearch-&-Triples.ipynb Embeddings-Elasticsearch-&-Triples.out.ipynb \
# -p claims_file /Volumes/GoogleDrive/Shared\ drives/KGTK-public-graphs/wikidata-20200803-v4/all.tsv.gz \
# -p label_file /Volumes/GoogleDrive/Shared\ drives/KGTK-public-graphs/wikidata-20200803-v4/part.label.en.tsv.gz \
# -p item_file /Volumes/GoogleDrive/Shared\ drives/KGTK-public-graphs/wikidata-20200803-v4/part.wikibase-item.tsv.gz \
# -p property_item_file = /Volumes/GoogleDrive/Shared\ drives/KGTK-public-graphs/wikidata-20200803-v4/part.property.wikibase-item.tsv.gz \
# -p output_path <local folder> \
# -p output_folder useful_files_v4 \
# -p temp_folder temp.useful_files_v4 \
# -p delete_database no
# -p languages es,ru,zh-cn
# ```
# + tags=["parameters"]
# Parameters
wikidata_root_folder = "/data/amandeep/wikidata-20210215-dwd-v2"
items_file = "claims.wikibase-item.tsv.gz"
all_sorted_file = "all.sorted.tsv.gz"
en_labels_file = "labels.en.tsv.gz"
temp_folder = "temp.elasticsearch.embeddings"
# -
import os
os.environ['OUT'] = f"{wikidata_root_folder}"
os.environ['kgtk'] = "kgtk --debug"
os.environ['ITEMS'] = f"{wikidata_root_folder}/{items_file}"
os.environ['ALL'] = f"{wikidata_root_folder}/{all_sorted_file}"
os.environ['LABELS_EN'] = f"{wikidata_root_folder}/{en_labels_file}"
# ## Graph Embeddings
# ### complEx
complex_temp_folder = f"{wikidata_root_folder}/temp.graph-embeddings.complex"
# !mkdir -p {complex_temp_folder}
os.environ['TEMP_COMPLEX'] = complex_temp_folder
# !kgtk graph-embeddings --verbose -i "$ITEMS" \
# -o $OUT/wikidatadwd.complEx.graph-embeddings.txt \
# --retain_temporary_data True \
# --operator ComplEx \
# --workers 24 \
# --log $TEMP_COMPLEX/ge.complex.log \
# -T $TEMP_COMPLEX \
# -ot w2v \
# -e 600
# ### transE
transe_temp_folder = f"{wikidata_root_folder}/temp.graph-embeddings.transe"
# !mkdir -p {transe_temp_folder}
os.environ['TEMP_TRANSE'] = transe_temp_folder
# !$kgtk graph-embeddings --verbose -i "$ITEMS" \
# -o $OUT/wikidatadwd.transE.graph-embeddings.txt \
# --retain_temporary_data True \
# --operator TransE \
# --workers 24 \
# --log $TEMP_TRANSE/ge.transE.log \
# -T $TEMP_TRANSE \
# -ot w2v \
# -e 600
# ### BERT Embeddings
# !$kgtk text-embedding -i $ALL \
# --model roberta-large-nli-mean-tokens \
# --property-labels-file $LABELS_EN \
# --isa-properties P31 P279 P106 P39 P1382 P373 P452 \
# --save-embedding-sentence > $OUT/wikidatadwd-text-embeddings-all.tsv
# ### Build KGTK edge file for KGTK Search
# !$kgtk cat -i $OUT/all.sorted.tsv.gz \
# -i $OUT/derived.isastar.tsv.gz \
# -i $OUT/metadata.property.datatypes.tsv.gz \
# -i $OUT/metadata.pagerank.undirected.tsv.gz \
# -i $OUT/metadata.pagerank.directed.tsv.gz \
# -o $OUT/wikidata.dwd.all.kgtk.search.unsorted.tsv.gz
# !$kgtk sort -i $OUT/wikidata.dwd.all.kgtk.search.unsorted.tsv.gz \
# --columns node1 \
# --extra '--parallel 24 --buffer-size 30% --temporary-directory ' + temp_folder_path \
# -o $OUT/wikidata.dwd.all.kgtk.search.sorted.tsv.gz
# + active=""
# tl build-elasticsearch-input --input-file wikidata.dwd.all.kgtk.search.sorted.tsv.gz \
# --output-file wikidata.dwd.all.kgtk.search.sorted.jl \
# --label-properties label \
# --alias-properties alias \
# --description-properties description \
# --pagerank-properties directed_pagerank \
# --mapping-file wikidata_os_mapping_es_ver7_v2.json \
# --copy-to-properties labels,aliases --es-version 7
# -
# The cell above and below are same except `Pdirected_pagerank` instead of `directed_pagerank`
tl build-elasticsearch-input --input-file wikidata.dwd.all.kgtk.search.sorted.tsv.gz \
--output-file wikidata.dwd.all.kgtk.search.sorted.jl \
--label-properties label \
--alias-properties alias \
--description-properties description \
--pagerank-properties Pdirected_pagerank \
--mapping-file wikidata_os_mapping_es_ver7_v2.json \
--copy-to-properties labels,aliases --es-version 7
# ### Build KGTK edge file for Triple generation
#
# !$kgtk cat \
# -i $OUT/wikidata.dwd.all.kgtk.search.sorted.tsv.gz \
# -i $OUT/derived.isa.tsv.gz \
# -i $OUT/derived.P279star.tsv.gz \
# -i $OUT/metadata.in_degree.tsv.gz \
# -i $OUT/metadata.out_degree.tsv.gz \
# -o $TEMP/wikidata.dwd.all.kgtk.triples.1.tsv.gz
# !$kgtk add-id -i $TEMP/wikidata.dwd.all.kgtk.triples.1.tsv.gz \
# --id-style wikidata \
# -o $TEMP/wikidata.dwd.all.kgtk.triples.2.tsv.gz
# !$kgtk sort -i $TEMP/wikidata.dwd.all.kgtk.triples.2.tsv.gz \
# --columns node1 \
# --extra '--parallel 24 --buffer-size 30% --temporary-directory ' + temp_folder_path \
# -o $OUT/wikidata.dwd.all.kgtk.triples.sorted.tsv.gz
# Split the triples file to parallelize triple generation
# !mkdir -p $OUT/kgtk_triples_split
# !$kgtk split -i $OUT/wikidata.dwd.all.kgtk.triples.sorted.tsv.gz \
# --output-path $OUT/kgtk_triples_split \
# --gzipped-output --lines 10000000 \
# --file-prefix kgtk_triples
# !curl https://raw.githubusercontent.com/usc-isi-i2/kgtk/dev/kgtk-properties/kgtk.properties.tsv -o $TEMP/kgtk-properties.tsv
# !$kgtk filter -p ";data_type;" -i $TEMP/kgtk-properties.tsv -o $TEMP/kgtk-properties.datatype.tsv.gz
# !$kgtk cat -i $TEMP/kgtk-properties.datatype.tsv.gz $OUT/metadata.property.datatypes.tsv.gz -o $OUT/metadata.property.datatypes.augmented.tsv.gz
# +
# ls $OUT/kgtk_triples_split/*.tsv.gz | parallel -j 18 'kgtk --debug generate-wikidata-triples -lp label -ap alias -dp description -pf $OUT/metadata.property.datatypes.augmented.tsv.gz --output-n-lines 100000 --generate-truthy --warning --use-id --log-path $TEMP/generate_triples_log.txt --error-action log -i {} -o {.}.ttl'
# -
# ### Build KGTK edge file for Table Linker Index
# The augmentation files can be downloaded from Google Drive,
# https://drive.google.com/drive/u/1/folders/1qbbgjo7pddMdDvQzOSeSaL6lYwj_f5gi
def convert_w2v_kgtk(input_path, output_path, label):
f = open(input_path)
o = gzip.open(output_path, 'wt')
o.write("node1\tlabel\tnode2\n")
i = 0
for line in f:
if i % 1000000 == 0:
print(i)
i += 1
vals = line.strip().split(" ")
if len(vals) == 2: # first line
continue
qnode = vals[0]
vector = ",".join(vals[1:])
o.write(f"{qnode}\t{label}\t{vector}\n")
print("Done!")
convert_w2v_kgtk("$OUT/wikidatadwd.complEx.graph-embeddings.txt", "$OUT/wikidatadwd.complEx.graph-embeddings.tsv.gz", "graph_embeddings_complEx")
convert_w2v_kgtk('$OUT/wikidatadwd.transE.graph-embeddings.txt', '$OUT/wikidatadwd.transE.graph-embeddings.tsv.gz', 'graph_embeddings_transE')
# !$kgtk cat -i $OUT/wikidata.dwd.all.kgtk.search.sorted.tsv.gz \
# /data/amandeep/wikidata_augmented_files_for_table_linker_index/augmentation.wikipedia.anchors.tsv.gz \
# /data/amandeep/wikidata_augmented_files_for_table_linker_index/augmentation.wikipedia.tables.anchors.tsv.gz \
# /data/amandeep/wikidata_augmented_files_for_table_linker_index/augmentation.wikipedia.redirect.tsv.gz \
# $OUT/item.property.count.compact.tsv.gz \
# $OUT/dwd_isa_class_count.compact.tsv.gz \
# $OUT/wikidatadwd.complEx.graph-embeddings.tsv.gz \
# $OUT/wikidatadwd.transE.graph-embeddings.tsv.gz \
# $OUT/text-embeddings-concatenated.tsv.gz \
# $OUT/derived.dwd_isa.tsv.gz \
# $OUT/table_linker.qnode.property.values.tsv.gz \
# $OUT/derived.isastar.tsv.gz \
# -o $TEMP/wikidata.dwd.table-linker.index.tsv.gz
# !$kgtk sort -i $TEMP/wikidata.dwd.table-linker.index.tsv.gz \
# --columns node1 \
# --extra '--parallel 24 --buffer-size 30% --temporary-directory ' + temp_folder_path \
# -o $OUT/wikidata.dwd.table-linker.index.sorted.tsv.gz
# !$kgtk build-kgtk-search-input --input-file $OUT/wikidata.dwd.table-linker.index.sorted.tsv.gz \
# --output-file $OUT/wikidata.dwd.table-linker.index.sorted.jl \
# --label-properties label \
# --alias-properties alias \
# --extra-alias-properties P1448,P1705,P1477,P1810,P742,P1449 \
# --description-properties description \
# --pagerank-properties directed_pagerank \
# --mapping-file $OUT/wikidata_dwd.v2.table-linker.json \
# --property-datatype-file $OUT/metadata.property.datatypes.augmented.tsv.gz
# Use above or below, depending on what the pagerank property is.
# !$kgtk build-kgtk-search-input --input-file $OUT/wikidata.dwd.table-linker.index.sorted.tsv.gz \
# --output-file $OUT/wikidata.dwd.table-linker.index.sorted.jl \
# --label-properties label \
# --alias-properties alias \
# --extra-alias-properties P1448,P1705,P1477,P1810,P742,P1449 \
# --description-properties description \
# --pagerank-properties Pdirected_pagerank \
# --mapping-file $OUT/wikidata_dwd.v2.table-linker.json \
# --property-datatype-file $OUT/metadata.property.datatypes.augmented.tsv.gz
|
use-cases/Embeddings-Elasticsearch-&-Triples.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# <font size="+5">#05 | Cluster Analysis con k-Means</font>
# - Subscribe to my [Blog ↗](https://blog.pythonassembly.com/)
# - Let's keep in touch on [LinkedIn ↗](www.linkedin.com/in/jsulopz) 😄
# # Discipline to Search Solutions in Google
# > Apply the following steps when **looking for solutions in Google**:
# >
# > 1. **Necesity**: How to load an Excel in Python?
# > 2. **Search in Google**: by keywords
# > - `load excel python`
# > - ~~how to load excel in python~~
# > 3. **Solution**: What's the `function()` that loads an Excel in Python?
# > - A Function to Programming is what the Atom to Phisics.
# > - Every time you want to do something in programming
# > - **You will need a `function()`** to make it
# > - Theferore, you must **detect parenthesis `()`**
# > - Out of all the words that you see in a website
# > - Because they indicate the presence of a `function()`.
# # Load the Data
# > - Simply execute the following lines of code to load the data
# > - This dataset contains **statistics** (columns)
# > - About **Car Models** (rows)
import seaborn as sns
import pandas as pd
import numpy as np
df = sns.load_dataset(name='mpg', index_col='name')
df.sample(10)
df
# # Data `preprocessing`
#
# > - Do you need to *transform* the data
# > - To get a **truthful insight** of the model?
from sklearn.preprocessing import MinMaxScaler
scaler = MinMaxScaler(feature_range=0,1)
scaler = MinMaxScaler(feature_range=(0, 1))
scaler.fit(X=df)
df = pd.get_dummies(df, drop_first=True)
df.head(5)
scaler.fit(df)
df
scaler = MinMaxScaler()
scaler.__dict__
scaler.fit(X=df)
scaler.__dict__
dfsel = df[['...']]
data = scaler.transform(X=df)
dfnorm = pd.DataFrame(data,columns=df.columns, index=df.index)
dfnorm
df.
df
df.head()
df.corr().style.background_gradient()
dfnorm.head()
dfnorm.corr().style.background_gradient()
# # `KMeans()` Model in Python
# ## Build the Model
# > 1. **Necesity**: Build Model
# > 2. **Google**: How do you search for the solution?
# > 3. **Solution**: Find the `function()` that makes it happen
# ## Code Thinking
#
# > Which function computes the Model?
# > - `fit()`
# >
# > How could can you **import the function in Python**?
# 1. creo que tengo que hacer esto...
# 2. he apilcado la intuicion importando sklearn
# 3. no he visto nada qu emayude
# 4. me voy a google y busco esto " ...... "
# 5. me he metido en esta pagina y creo que este codigo esta bien
# 6. copio y pego
from sklearn.cluster import KMeans
model = KMeans(n_clusters=3)
# + [markdown] tags=[]
# ### Separate Variables for the Model
#
# > Regarding their role:
# > 1. **Target Variable `y`**
# >
# > - [ ] What would you like **to predict**?
# >
# > The group to which each row belongs to
# >
# > 2. **Explanatory Variable `X`**
# >
# > - [ ] Which variable will you use **to explain** the target?
# >
# > - [ ] Is something extrange in the `instructions manual` of the function?
# -
model.fit(X=df)
# ### Data Visualization to Analyize Patterns
# > - Visualize the 2 variables with a `scatterplot()`
# > - And decide *how many `clusters`* you'd like to calculate
df.head()
sns.scatterplot(x='mpg', y='weight', data=df)
# ### Finally `fit()` the Model
model.fit(X=dfnorm[['mpg', 'weight']])
# ## `predict()` the *cluster* for every row
# > - `model.` + `↹`
grupos = model.predict(dfnorm[['mpg', 'weight']])
grupos
# > - Create a `dfsel` DataFrame
# > - That contains the **columns you used for the model**
dfsel = df[['mpg', 'weight']].copy()
dfsel.head()
# > - Add a **new column**
# > - That **contains the `cluster` prediction** for every USA State
dfsel['grupos'] = grupos
# ## Model Visualization
# > - You may `hue=` the points with the `cluster` column
df['grupos']
sns.scatterplot(x='mpg', y='weight', data=dfsel, hue='grupos')
sns.scatterplot(x='mpg', y='weight', data=df, hue='grupos')
sns.scatterplot(x='mpg', y='weight', data=df, hue=grupos)
# ## Model Interpretation
# +
# %%HTML
<iframe width="560" height="315" src="https://www.youtube.com/embed/4b5d3muPQmA" title="YouTube video player" frameborder="0" allow="accelerometer; autoplay; clipboard-write; encrypted-media; gyroscope; picture-in-picture" allowfullscreen></iframe>
# -
# > - Can you put a **`nickname` to each group**?
# > - Observe the `centroids` within `model.` + `↹`
# ## Model Visualization with Centroids
#
# > - I want to see the `centroid`
# > - with a **big `marker="X"`** in the plot
# # Achieved Goals
# _Double click on **this cell** and place an `X` inside the square brackets (i.e., [X]) if you think you understand the goal:_
#
# - [ ] Understand how the **machine optimizes a model**
# - No more than to find the best numbers for a mathematical equation
# - [ ] **Residual Sum of Squares (RSS)** as a fundamental measure for the **error**. We see it on ↓
# - [Neural Networks](https://youtu.be/IHZwWFHWa-w?t=211)
# - Linear Regression
# - Variance
# - [ ] Understand the necessity to **Scale** the Data
# - For all algorithms that involves **distance calculation**.
# - [ ] Understand that programming is not an end itself, but a tool to achieve the end
# - We need to understand the problem and design the solution before coding
# - But we won't need how to design the solution if we don't know how to code first
# - Solution? Apply the discipline
# - [ ] There is **not a unique way to group data**. The same way it is not a unique way ↓
# - To predict a number **Regression Mathematical Equations**
# - To predict a category **Classification Mathematical Equations**
|
II Machine Learning & Deep Learning/05_Cluster Analysis con k-Means/05practice_cluster-kmeans.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Childhood pneumonia detection
#
# * Student name: <NAME>
# * Student pace: Part Time Online
# * Scheduled project review date/time: June 27, 2020 2:00 PM PDT
# * Instructor name: <NAME>
# * Blog post URL: https://github.com/sn95033/pneumonia-detection
#
#
# * **Data Source:**
# - References: <br>https://www.kaggle.com/paultimothymooney/chest-xray-pneumonia
# <br> http://www.cell.com/cell/fulltext/S0092-8674(18)30154-5 <br> <br>
#
# <div>
# <img src= "childrens_pneumonia.png"
# width=400"/>
# </div
#
# <br> Illustrative Examples of Chest X-Rays in Patients with Pneumonia.
# The normal chest X-ray (left panel) depicts clear lungs without any areas of abnormal opacification in the image. Bacterial pneumonia (middle) typically exhibits a focal lobar consolidation, in this case in the right upper lobe (white arrows), whereas viral pneumonia (right) manifests with a more diffuse ‘‘interstitial’’ pattern in both lungs.
# http://www.cell.com/cell/fulltext/S0092-8674(18)30154-5
#
# The dataset is organized into 3 folders (train, test, val) and contains subfolders for each image category (Pneumonia/Normal). There are 5,863 X-Ray images (JPEG) and 2 categories (Pneumonia/Normal).
#
# Chest X-ray images (anterior-posterior) were selected from retrospective cohorts of pediatric patients of one to five years old from Guangzhou Women and Children’s Medical Center, Guangzhou. All chest X-ray imaging was performed as part of patients’ routine clinical care.
#
# For the analysis of chest x-ray images, all chest radiographs were initially screened for quality control by removing all low quality or unreadable scans. The diagnoses for the images were then graded by two expert physicians before being cleared for training the AI system. In order to account for any grading errors, the evaluation set was also checked by a third expert.
#
# Acknowledgements
# Data: https://data.mendeley.com/datasets/rscbjbr9sj/2
#
# License: CC BY 4.0
#
# Citation: http://www.cell.com/cell/fulltext/S0092-8674(18)30154-5
#
# # Outline of Process
#
# 1. **Import required libraries**
# 2. **Define Functions for Analysing Results**
# 3. ** Optional Setup Colab integration**
# - **Setting up Google Drive for data**
# 4. **Loading in the data and creating a file structure**
# 5. **Inspecting the data and Decisions for Next Steps**
# 6. **Image Process for initial CNN model development**
# 7. **Validation and Training Data**
# - **Compacting the image data**
# - **Normalization of images**
# - **Form data into Numpy Arrays for Keras**
# - **Use of Image Augmentation for Class Balance on the Training Dataset** <br><br>
# 8. **Creation of Initial Model**
# 9. **Creating Callbacks for early stopping, saving model**
# 10. **Loading of Pre-trained Weights - Transfer Learning**
# 11. **Optional - Starting Colab Pro GPU and RAM**
# 12. **Running model**
#
#
#
# # 1. Importing Libraries
# +
# # !pip install -U imageio
# # !pip install -U scikit-image
# # !pip install pillow
# # !pip install opencv-contrib-python
# # !pip install -U fsds
# # !pip install -U tensorflow
# # !pip install -U keras
# # !pip install -U pandas
# # !pip install -U pandas-profiling
# # # %conda update matploltib
# # !pip install -U matplotlib
# # !pip install imgaug
# # !pip install mlxtend
# -
import os
import glob
import h5py
import shutil
import imgaug as aug
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
import seaborn as sns
import matplotlib.pyplot as plt
import matplotlib.image as mimg
import imgaug.augmenters as iaa
from os import listdir, makedirs, getcwd, remove
from os.path import isfile, join, abspath, exists, isdir, expanduser
from PIL import Image
from pathlib import Path
from skimage.io import imread
from skimage.transform import resize
from keras.models import Sequential, Model
from keras.applications.vgg16 import VGG16, preprocess_input
from keras.preprocessing.image import ImageDataGenerator,load_img, img_to_array
from keras.models import Sequential
from keras.layers import Conv2D, MaxPooling2D, Dense, Dropout, Input, Flatten, SeparableConv2D
from keras.layers import GlobalMaxPooling2D
from keras.layers.normalization import BatchNormalization
from keras.layers.merge import Concatenate
from keras.models import Model
from keras.optimizers import Adam, SGD, RMSprop
from keras.callbacks import ModelCheckpoint, Callback, EarlyStopping
from keras.utils import to_categorical
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
from mlxtend.plotting import plot_confusion_matrix
from sklearn.metrics import confusion_matrix
import cv2
from keras import backend as K
color = sns.color_palette()
# %matplotlib inline
# #!pip install -U fsds_100719
from fsds_100719.imports import *
# # 2. Functions Created for Analysis
# +
import sklearn.metrics as metrics
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
def plot_keras_history(history,figsize=(10,4),subplot_kws={}):
if hasattr(history,'history'):
history=history.history
try:
acc_keys = list(filter(lambda x: 'acc' in x,history.keys()))
except:
print('No acc keys found')
pass
try:
loss_keys = list(filter(lambda x: 'loss' in x,history.keys()))
except:
print('No loss keys found')
pass
plot_me = pd.DataFrame(history)
fig,axes=plt.subplots(ncols=2,figsize=figsize,**subplot_kws)
axes = axes.flatten()
y_labels= ['Accuracy','Loss']
for a, metric in enumerate([acc_keys,loss_keys]):
for i in range(len(metric)):
ax = pd.Series(history[metric[i]],
name=metric[i]).plot(ax=axes[a],label=metric[i])
[ax.legend() for ax in axes]
[ax.xaxis.set_major_locator(mpl.ticker.MaxNLocator(integer=True)) for ax in axes]
[ax.set(xlabel='Epochs') for ax in axes]
plt.suptitle('Model Training Results',y=1.01)
plt.tight_layout()
plt.show()
return plt.gcf()
def plot_confusion_matrix(conf_matrix, classes = None, normalize=True,
title='Confusion Matrix', cmap="Blues",
print_raw_matrix=False,
fig_size=(4,4)):
"""Check if Normalization Option is Set to True.
If so, normalize the raw confusion matrix before visualizing
#Other code should be equivalent to your previous function.
Note: Taken from bs_ds and modified
- Can pass a tuple of (y_true,y_pred) instead of conf matrix.
"""
import itertools
import numpy as np
import matplotlib.pyplot as plt
import sklearn.metrics as metrics
## make confusion matrix if given tuple of y_true,y_pred
if isinstance(conf_matrix, tuple):
y_true = conf_matrix[0].copy()
y_pred = conf_matrix[1].copy()
if y_true.ndim>1:
y_true = y_true.argmax(axis=1)
if y_pred.ndim>1:
y_pred = y_pred.argmax(axis=1)
cm = metrics.confusion_matrix(y_true,y_pred)
else:
cm = conf_matrix
## Generate integer labels for classes
if classes is None:
classes = list(range(len(cm)))
## Normalize data
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
fmt='.2f'
else:
fmt= 'd'
fontDict = {
'title':{
'fontsize':16,
'fontweight':'semibold',
'ha':'center',
},
'xlabel':{
'fontsize':14,
'fontweight':'normal',
},
'ylabel':{
'fontsize':14,
'fontweight':'normal',
},
'xtick_labels':{
'fontsize':10,
'fontweight':'normal',
# 'rotation':45,
'ha':'right',
},
'ytick_labels':{
'fontsize':10,
'fontweight':'normal',
'rotation':0,
'ha':'right',
},
'data_labels':{
'ha':'center',
'fontweight':'semibold',
}
}
# Create plot
fig,ax = plt.subplots(figsize=fig_size)
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title,**fontDict['title'])
plt.colorbar()
tick_marks = classes#np.arange(len(classes))
plt.xticks(tick_marks, classes, **fontDict['xtick_labels'])
plt.yticks(tick_marks, classes,**fontDict['ytick_labels'])
# Determine threshold for b/w text
thresh = cm.max() / 2.
# fig,ax = plt.subplots()
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, format(cm[i, j], fmt),
color='darkgray',**fontDict['data_labels']) #color="white" if cm[i, j] > thresh else "black"
plt.tight_layout()
plt.ylabel('True label',**fontDict['ylabel'])
plt.xlabel('Predicted label',**fontDict['xlabel'])
if print_raw_matrix:
print_title = 'Raw Confusion Matrix Counts:'
print('\n',print_title)
print(conf_matrix)
fig = plt.gcf()
return fig
class Timer():
## def init
def __init__(self,format_="%m/%d/%y - %I:%M %p"):
import tzlocal
self.tz = tzlocal.get_localzone()
self.fmt = format_
self.created_at = self.get_time()# get time
## def get time method
def get_time(self):
import datetime as dt
return dt.datetime.now(self.tz)
## def start
def start(self):
time = self.get_time()
self.start = time
print(f"[i] Timer started at{self.start.strftime(self.fmt)}")
## def stop
def stop(self):
time = self.get_time()
self.end = time
print(f"[i] Timer ended at {self.end.strftime(self.fmt)}")
print(f"- Total time = {self.end-self.start}")
# -
# # 3. Option - Colaboratory Integration
# ## A. Mounting the Google Drive so it can be accessed in Colab
#
# ## B. Defining the Folder where data is located and finding the zip file
#
# ## C. Reading the zip file
#
# ## D. Creating a local Colab file structure to store images
#
# + code_folding=[0]
# Mount the Google Drive so you can access it (where your data is stored) from your Colab Account:
from google.colab import drive
drive.mount('/gdrive',force_remount=True)
# # %cd /gdrive/My\ Drive
# %cd ~
# %cd ..
# -
# #### You will see a pop-up as shown below, which will ask you to get a permission key from Google drive. Just Click the link, login to your Google account, get the key (authorization code), and paste it into the Colab Notebook entry
#
# Go to this URL in a browser: https://accounts.google.com/o/oauth2/auth?client_id=947318989803-6bn6qk8qdgf4n4g3pfee6491hc0brc4i.apps.googleusercontent.com&redirect_uri=urn%3aietf%3awg%3aoauth%3a2.0%3aoob&response_type=code&scope=email%20https%3a%2f%2fwww.googleapis.com%2fauth%2fdocs.test%20https%3a%2f%2fwww.googleapis.com%2fauth%2fdrive%20https%3a%2f%2fwww.googleapis.com%2fauth%2fdrive.photos.readonly%20https%3a%2f%2fwww.googleapis.com%2fauth%2fpeopleapi.readonly
#
# Enter your authorization code:
# ··········
# Mounted at /gdrive
# /root
# /
# +
# import os,glob
# os is a utility which allows use of Unix commands to access the file structure in Google Drive.
# glob is a utility so that you don't have to specify a specific file path, you can just type "glob" as a wildcard meaning, any directory
print(os.path.abspath(os.curdir))
source_folder = r'/gdrive/My Drive/Datasets/' #setup the source_folder to read from the directory where the data is
# since the data is stored in a zip file, the filename has a .zip at the end of it
# Below we assign variable file to look for any director, with the source_folder path, and ends with .zip
# Note this means that there should only be 1 zip file inside that folder.
# Recursive = true enables multiple folders to be opened
# [0] means that the file is set to the top level folder
file = glob.glob(source_folder+'*.zip',recursive=True)[0]
file # Print the file structure where the zip file is loaded
# target_folder = r'/content/'
# os.listdir(source_folder), os.listdir(target_folder)
# +
# Assign a zip_path where the files in the zip file will be stored
zip_path = file
# !cp "{zip_path}" . # Create the zip path in the root director (.)
# !unzip -q 17810_23812_bundle_archive.zip # Unzip the files
print(os.path.abspath(os.curdir))
os.listdir()
# os.chdir('My Drive/')
# +
# Input data files are available in the "../input/" directory.
# For example, running this (by clicking run or pressing Shift+Enter) will list the files in the input directory
print(os.listdir("chest_xray"))
# Any results you write to the current directory are saved as output.
# -
# # 4. Local File Structure, Data Loading, and Data Inspection
# +
# Place the unzipped data files into a folder called Datasets directory.
# The zipped data can be obtained from kaggle: https://www.kaggle.com/paultimothymooney/chest-xray-pneumonia
# Note: As of 6/1/2020, the Kaggle zipped file contains a directory called chest_xray, and a duplicate directory called chest_xray
# Both chest_xray directories contain folders named "train", "val", and "test"
# Obviously, only one chest_xray with its folders is needed to be placed into the input directory
# List the contents of the chest_xray folder
#print(os.listdir("Datasets/chest_xray"))
# +
# Define path to the data directory
data_dir = Path('Datasets/chest_xray')
# Path to train directory (Fancy pathlib...no more os.path!!)
train_dir = data_dir / 'train'
# Path to validation directory
val_dir = data_dir / 'val'
# Path to test directory
test_dir = data_dir / 'test'
# +
# Get the path to the normal and pneumonia sub-directories
normal_cases_dir = train_dir / 'NORMAL'
pneumonia_cases_dir = train_dir / 'PNEUMONIA'
# Get the list of all the images
normal_cases = normal_cases_dir.glob('*.jpeg')
pneumonia_cases = pneumonia_cases_dir.glob('*.jpeg')
# An empty list. We will insert the data into this list in (img_path, label) format
train_data = []
# Go through all the normal cases. The label for these cases will be 0
for img in normal_cases:
train_data.append((img,0))
# Go through all the pneumonia cases. The label for these cases will be 1
for img in pneumonia_cases:
train_data.append((img, 1))
# Get a pandas dataframe from the data we have in our list
train_data = pd.DataFrame(train_data, columns=['image', 'label'],index=None)
# Shuffle the data
train_data = train_data.sample(frac=1.).reset_index(drop=True)
# How the dataframe looks like?
train_data.head()
# +
# Get the # of each class
case_count = train_data['label'].value_counts()
print(case_count)
# Plot the results
plt.figure(figsize=(10,8))
sns.barplot(x=case_count.index, y= case_count.values)
plt.title('Number of Pneumonia Images vs. Normal Images', fontsize=14)
plt.xlabel('Case Type', fontsize=12)
plt.ylabel('Count', fontsize=12)
plt.xticks(range(len(case_count.index)), ['Normal(0)', 'Pneumonia(1)'])
plt.show();
# +
# Get few samples for both the classes
pneumonia_samples = (train_data[train_data['label']==1]['image'].iloc[:25:5]).tolist()
normal_samples = (train_data[train_data['label']==0]['image'].iloc[:25:5]).tolist()
# Concat the data in a single list and del the above two list
samples = pneumonia_samples + normal_samples
del pneumonia_samples, normal_samples
# Plot the data
f, ax = plt.subplots(2,5, figsize=(30,10))
for i in range(10):
img = imread(samples[i])
ax[i//5, i%5].imshow(img, cmap='gray')
if i<5:
ax[i//5, i%5].set_title("Pneumonia")
else:
ax[i//5, i%5].set_title("Normal")
ax[i//5, i%5].axis('off')
ax[i//5, i%5].set_aspect('auto')
plt.show()
# -
plt.savefig("Typical images of pneumonia and normal cases.png")
# # 5. Discussion on Data Inspection and Next Steps
# ## Conclusions
#
# * The classes are imbalanced -- there are 3X more pneumonia images than normal images for training.**
#
# * Inspecting the images shows there are some pretty subtle differences between pneumonia and normal images. Having a classifier with high accuracy to prevent false negatives, is probably more important than false positives.**
#
# * Class imbalance should be corrected either by down-sampling the pneumonia case or by: <br> <br>
#
# - [x] Down-sampling the "Pneumonia" image class <br>
# - [x] Augmenting the "Normal" image class <br><br>
#
# * 4. Next Step:
#
# - [x] We will start by down-sampling the "Pneumonia" image class, to get a preliminary model.
# - [x] Then improve the model by using the full dataset, and augmenting the "Normal" class, to provide class balance
#
# +
# change dataset_folder to match where you stored the files
#dataset_folder = "Datasets/chest_xray/"
#os.makedirs(dataset_folder,exist_ok=True)
# os.listdir(dataset_folder)
#base_folder = dataset_folder
#base_folder
# +
import shutil
import cv2,glob,os
#train_base_dir = base_folder+'train/'
#test_base_dir =base_folder+'test/'
#train_sick = train_base_dir+'PNEUMONIA/'
#train_well = train_base_dir+'NORMAL/'
#test_sick = test_base_dir+'PNEUMONIA/'
#test_well = test_base_dir+'NORMAL/'
# Define path to the data directory
data_dir = Path('Datasets/chest_xray')
# Path to train directory
train_dir = data_dir / 'train'
# Path to validation directory
val_dir = data_dir / 'val'
# Path to test directory
test_dir = data_dir / 'test'
# Get the path to the normal and pneumonia sub-directories
well_train_dir = train_dir / 'NORMAL'
sick_train_dir = train_dir / 'PNEUMONIA'
# Get the list of all the images
well_train_files = well_train_dir.glob('*.jpeg')
sick_train_files = sick_train_dir.glob('*.jpeg')
# Get the path to the normal and pneumonia sub-directories
well_test_dir = test_dir / 'NORMAL'
sick_test_dir = test_dir / 'PNEUMONIA'
# Get the list of all the images
well_test_files = well_test_dir.glob('*.jpeg')
sick_test_files = sick_test_dir.glob('*.jpeg')
# An empty list. We will insert the data into this list in (img_path, label) format
#train_data = []
# Go through all the normal cases. The label for these cases will be 0
#for img in normal_cases:
# train_data.append((img,0))
# Go through all the pneumonia cases. The label for these cases will be 1
#for img in pneumonia_cases:
# train_data.append((img, 1))
# Get a pandas dataframe from the data we have in our list
#train_data = pd.DataFrame(train_data, columns=['image', 'label'],index=None)
# Shuffle the data
#train_data = train_data.sample(frac=1.).reset_index(drop=True)
# How the dataframe looks like?
#train_data.head()
# +
import cv2,glob,os
#sick_train_files = glob.glob(train_sick+'*.jpg')
#well_train_files = glob.glob(train_well+'*.jpg')
#all_train_files = [*sick_train_files,*well_train_files]
#sick_test_files = glob.glob(test_sick+'*.jpg')
#well_test_files = glob.glob(test_well+'*.jpg')
#all_test_files = [*sick_test_files,*well_test_files]
# print(len(img_filenames))
# img_filenames[:10]
#all_filename_vars = [*all_train_files, *all_test_files]
#print(all_test_files, all_filename_vars)
# -
def load_image_cv2(filename, RGB=True):
"""Loads image using cv2 and converts to either matplotlib-RGB (default)
or grayscale."""
import cv2
IMG = cv2.imread(filename)
if RGB: cmap = cv2.COLOR_BGR2RGB
else: cmap=cv2.COLOR_BGR2GRAY
return cv2.cvtColor(IMG,cmap)
# +
from PIL import Image
from keras.preprocessing import image
from imageio import imread
from skimage.transform import resize
import cv2
from tqdm import tqdm
from keras.utils import to_categorical
# defining a function to read images
def read_img(img_path,target_size=(32,32,3)):#(64, 64, 3)):
img = image.load_img(img_path, target_size=target_size)
img = image.img_to_array(img)
return img
def load_train_test_val(sick_training_filenames, well_training_filenames,
sick_test_filenames,well_test_filenames,
img_size=(32,32,3),val_size=0.1):
"""Reads in training and test filenames to produce X and y data splits.
The validation set is intended to be used during training and is created
from the training images using train test split.
ylabels are encoded as 0=cat, 1=dog
Returns: X_train, X_test, X_val, y_train, y_test,y_val"""
#
display('[i] LOADING IMAGES')
train_img = []
train_label = []
for img_path in tqdm(sick_training_filenames):
train_img.append(read_img(img_path,target_size=img_size))
train_label.append(1)
for img_path in tqdm(well_training_filenames):
train_img.append(read_img(img_path,target_size=img_size))
train_label.append(0)
test_img = []
test_label = []
for img_path in tqdm(sick_test_files):
test_img.append(read_img(img_path,target_size=img_size))
test_label.append(1)
for img_path in tqdm(well_test_files):
test_img.append(read_img(img_path,target_size=img_size))
test_label.append(0)
# print('\n',pd.Series(train_label).value_counts())
from sklearn.model_selection import train_test_split
X = np.array(train_img, np.float32)
y = np.array(train_label)
y = to_categorical(y)
X_test = np.array(test_img, np.float32)
y_test = to_categorical(np.array(test_label))
X_train,X_val,y_train,y_val = train_test_split(X,y,test_size=0.1)
print('\n[i] Length of Splits:')
print(f"X_train={len(X_train)}, X_test={len(X_test)}, X_val={len(X_val)}")
return X_train, X_test, X_val, y_train, y_test,y_val
def train_test_val_datagens(X_train,X_test,X_val,y_train,y_test,y_val,
BATCH_SIZE = 32, train_datagen_kws= dict(
shear_range = 0.2,
zoom_range = 0.2,
horizontal_flip = True)):
"""Creates ImageDataGenerators for train,test,val data.
Returns: training_set,test_set,val_set"""
## Create training and test data
from keras.preprocessing.image import ImageDataGenerator
train_datagen = ImageDataGenerator(rescale = 1./255,**train_datagen_kws)
test_datagen = ImageDataGenerator(rescale = 1./255)
val_datagen = ImageDataGenerator(rescale = 1./255)
training_set = train_datagen.flow(X_train,y=y_train,batch_size=BATCH_SIZE)
test_set = test_datagen.flow(X_test,y=y_test,batch_size=BATCH_SIZE)
val_set = val_datagen.flow(X_val,y=y_val,batch_size=BATCH_SIZE)
return training_set,test_set,val_set
def get_shapes_dict(training_set,verbose=False):
shapes = ["Batchsize", "img_width","img_height","img_dim"]
SHAPES = dict(zip(shapes, training_set[0][0].shape))
if verbose:
print('SHAPES DICT:')
print(SHAPES)
print(training_set[0][0].shape)
print('\n[i] Labels for batch (0=cat,1=dog)')
print(training_set[0][1])
return SHAPES
# +
## USING FUNCTIONS TO LOAD IN IMAGES
X_train,X_test,X_val,y_train,y_test,y_val = load_train_test_val(sick_train_files, well_train_files,
sick_test_files,well_test_files,
val_size=0.1,img_size=(32,32,3))#(64,64,3))
train_test_val_vars = [X_train,X_test,X_val,y_train,y_test,y_val ]
# -
y_train
ex_img = X_train[0]
ex_img.shape
ex_img.shape[0] *ex_img.shape[1]*ex_img.shape[2]
print(ex_img.shape)
ex_img = X_train[0]
num_images = X_train.shape[0]
num_images_test = X_test.shape[0]
len_unrowed = ex_img.shape[0] *ex_img.shape[1]*ex_img.shape[2]
num_images,num_images_test, len_unrowed
X_train.shape
X_train[0].shape, 32*32*3
X_train_img = X_train.reshape(num_images,len_unrowed).astype('float32')/255
X_train_img.shape
X_val_img = X_val.reshape(X_val.shape[0],len_unrowed).astype('float32')/255
X_val_img.shape
X_test_img = X_test.reshape(num_images_test,len_unrowed).astype('float32')/255
X_test_img.shape
# + active=""
#
# +
from PIL import Image
from keras.preprocessing import image
# Check one image
i = np.random.choice(range(len(y_train)))
display(y_train[i])
display(image.array_to_img(X_train_img[i].reshape(32,32,3)))
# -
# Part 1 - Building the CNN
# Importing the Keras libraries and packages
from keras.models import Sequential
# from keras.layers import Conv2D
# from keras.layers import MaxPooling2D
# from keras.layers import Flatten
from keras.layers import Dense
X_train_img[0].shape
X_train_img.shape, y_train.shape
X_train_img[0].shape
def make_model():
model = Sequential()
model.add(Dense(64,activation='relu',input_shape=(X_train_img.shape[1],)))
model.add(Dense(2, activation='softmax'))
model.compile(loss='binary_crossentropy', optimizer='sgd', metrics=['accuracy'])
return model
model = make_model()
model.summary()
# +
def evaluate_model(y_true, y_pred,history=None):
from sklearn import metrics
if y_true.ndim>1:
y_true = y_true.argmax(axis=1)
if y_pred.ndim>1:
y_pred = y_pred.argmax(axis=1)
try:
if history is not None:
plot_keras_history(history)
except:
pass
num_dashes=20
print('\n')
print('---'*num_dashes)
print('\tCLASSIFICATION REPORT:')
print('---'*num_dashes)
try:
print(metrics.classification_report(y_true,y_pred))
fig = plot_confusion_matrix((y_true,y_pred))
plt.show()
except Exception as e:
print(f"[!] Error during model evaluation:\n\t{e}")
# -
timer = Timer()
model = make_model()
model.summary()
timer.start()
history = model.fit(X_train_img, y_train, epochs=100, batch_size=64,
validation_data=(X_val_img, y_val))
timer.stop()
# +
y_hat_test = model.predict(X_test_img)
evaluate_model(y_test,y_hat_test,history)
# -
# # Discussion and Next Steps
#
# * The plain vanilla model has 99% True positives (1 = well, 0 = sick), e.g. the model is overfitted.
#
# * Additionally, there are 68% false positives, i.e. those images showing pneumonia were predicted to be "well". Leaving a false negative of 32%. This is not what we would hope to have as the best result of the model.
#
# * We now can go ahead with the first approach mentioned earlier
#
# * [x] Augmentation to balance the classes (increase the number of "well" images to increase the data
# * [x] Decreasing the compression of the images
# * [x] Creating more layers in the model
# * [x] Using a pre-trained model
#
#
# +
# Get the path to the sub-directories
train_normal_cases_dir = train_dir / 'NORMAL'
train_pneumonia_cases_dir = train_dir / 'PNEUMONIA'
# Get the list of all the images
train_normal_cases = train_normal_cases_dir.glob('*.jpeg')
train_pneumonia_cases = train_pneumonia_cases_dir.glob('*.jpeg')
# List that are going to contain validation images data and the corresponding labels
train_data = []
train_labels = []
image_size = (32,32)
# Some images are in grayscale while majority of them contains 3 channels. So, if the image is grayscale, we will convert into a image with 3 channels.
# We will normalize the pixel values and resizing all the images to 224x224
# Normal cases
for img in train_normal_cases:
img = cv2.imread(str(img))
img = cv2.resize(img, image_size)
if img.shape[2] ==1:
img = np.dstack([img, img, img])
img = cv2.cvtColor(img, cv2.COLOR_BGR2RGB)
img = img.astype(np.float32)/255.
label = to_categorical(0, num_classes=2)
train_data.append(img)
train_labels.append(label)
aug_img1 = seq.augment_image(img)
aug_img2 = seq.augment_image(img)
aug_img1 = cv2.cvtColor(aug_img1, cv2.COLOR_BGR2RGB)
aug_img2 = cv2.cvtColor(aug_img2, cv2.COLOR_BGR2RGB)
aug_img1 = aug_img1.astype(np.float32)/255.
aug_img2 = aug_img2.astype(np.float32)/255.
train_data.append(aug_img1)
train_labels.append(label)
train_data.append(aug_img2)
train_labels.append(label)
check_train_data = np.array(train_data)
check_train_labels = np.array(train_labels)
print("Total number of Normal Train examples: ", check_train_data.shape)
print("Total number of Normal Train lables: ", check_train_labels.shape)
# Pneumonia cases
for img in train_pneumonia_cases:
img = cv2.imread(str(img))
img = cv2.resize(img, (image_size))
if img.shape[2] ==1:
img = np.dstack([img, img, img])
img = cv2.cvtColor(img, cv2.COLOR_BGR2RGB)
img = img.astype(np.float32)/255.
label = to_categorical(1, num_classes=2)
train_data.append(img)
train_labels.append(label)
# Convert the list into numpy arrays
train_data = np.array(train_data)
train_labels = np.array(train_labels)
print("Total number of train examples: ", train_data.shape)
print("Total number of train labels:", train_labels.shape)
# -
# # 7.
# # Basics of Building a Neural Network with Keras:
#
#
# 1. **Import required modules**
# - **For general neural network**
# - `from keras import models, layers,optimizers`
# - **For text:**
# - `from keras.preprocessing.text import Tokenizer`
# - `from keras.utils import to_categorical`
# - **For images:**
# - `from keras.preprocessing.image import ImageDataGenerator, img_to_array, array_to_img`
# - **For relocating image files:**
# - `import os, shutil`
#
# 2. **Decide on a network architecture (have only discussed sequential thus far)**
# - `model = models.Sequential()`
#
# 3. **Adding layers - specifying layer type, number of neurons, activation functions, and, optionally, the input shape.**
# - `model.add(layers.Dense(units, activation='relu', input_shape))`
# - `model.add(layers.Dense(units, activation='relu')`
# - **3B. Final layer choice:**
# - Want to have as many neurons as classes you are trying to predict
# - Final activation function:
# - For binary classificaiton, use `activation='sigmoid'`
# - For multi classificaiton, use `activation='softmax'`
# - For regression tasks, have a single final neuron.
#
# 5. **Training the model**
# - `model.fit(X_train, y_train, epochs=20,batch_size=512,validation_data=(x_val,y_val))`
# - Note: if using images with ImageDataGenerator, use `model.fit_generator()`
#
# - **batches:**
# - a set of N samples, processed independently in parallel
# - a batch determines how many samples are fed through before back-propagation.
# - model only updates after a batch is complete.
# - ideally have as large of a batch as your hardware can handle without going out of memory.
# - larger batches usually run faster than smaller ones for evaluation/prediction.
# - **epoch:**
# - arbitrary cutoff / "one pass over the entire dataset", useful for logging and periodic evaluation
# - when using kera's `model.fit` parameters `validation_data` or `validation_split`, these evaluations run at the end of every epoch.
# - Within Keras can add callbacksto be run at the end of an epoch. Examples of these are learning rate changes and model checkpointing (saving).
#
#
# 6. **Evaluation / Predictions**
# - To get predicted results:
# - `y_hat_test = model.predict(test)`
# - To get evaluation metrics:
# - `results_test = model.evaluate(test, label_test)`
#
#
# 7. **Visualization** <br>
# - **`history = model.fit()` creates history object with .history attribute.**
# - `history.history()` returns a dictionary of metrics from each epoch.
# - `history.history['loss']` and `history.history['acc']` <br>
#
# 8. **Data Augmentation (not covered in class)** <br><br>
# - Simplest way to reduce overfitting is to increase the size of the training data.
# - Difficult to do with large datasets, but can be implemented with images as shown below:
# - **For augmenting image data:**
# - Can alter the images already present in the training data by shifting, shearing, scaling, rotating.<br><br> <img src ="https://www.dropbox.com/s/9i1hl3quwo294jr/data_augmentation_example.png?raw=1" width=300>
# - This usually provides a big leap in improving the accuracy of the model. It can be considered as a mandatory trick in order to improve our predictions.
#
# - **In Keras:**
# - `ImageDataGenerator` contains several augmentations available.
# - Example below:
#
# ```python
# from keras.preprocessing.image import ImageDataGenerator
# datagen = ImageDataGenerator(horizontal flip=True)
# datagen.fit(train)
#
# +
# !pip install -U imageio
# !pip install -U scikit-image
# !pip install pillow
# !pip install opencv-contrib-python
# !pip install -U fsds
# !pip install -U tensorflow
# !pip install -U keras
# !pip install -U pandas
# !pip install -U pandas-profiling
# %conda update matploltib
# %conda update scikit-learn
# !pip install -U matplotlib
# -
# !pip install -U fsds_100719
from fsds_100719.imports import *
# +
from PIL import Image
from keras.preprocessing import image
from keras.preprocessing.image import ImageDataGenerator, array_to_img, img_to_array, load_img
from keras.utils import to_categorical
from imageio import imread
from skimage.transform import resize
import cv2
from tqdm import tqdm
# -
import os # Do we need this?
# +
# Directory path
train_data_dir = 'chest_xray/train'
test_data_dir = 'chest_xray/test'
val_data_dir = 'chest_xray/val'
# get all the data from the test_data_dir and reshape them
test_generator = ImageDataGenerator().flow_from_directory(
test_data_dir,
target_size=(64, 64), batch_size=132)
# Get all the data in train_data_dir and
train_generator = ImageDataGenerator().flow_from_directory(
train_data_dir,
target_size=(64, 64), batch_size=32)
# Create the datasets
train_images, train_labels = next(train_generator)
test_images, test_labels = next(test_generator)
#Found 132 images belonging to 2 classes.
#Found 790 images belonging to 2 classes.
# -
train_labels
# change dataset_folder to match where you stored the files
dataset_folder = "chest_xray"
base_folder = dataset_folder
base_folder
# +
import shutil,glob
# ## DOG VS CAT
train_base_dir = base_folder+'training_set/'
test_base_dir =base_folder+'test_set/'
train_dogs = train_base_dir+'dogs/'
train_cats = train_base_dir+'cats/'
test_dogs = test_base_dir+'dogs/'
test_cats = test_base_dir+'cats/'
dog_train_files = glob.glob(train_dogs+'*.jpg')
cat_train_files = glob.glob(train_cats+'*.jpg')
all_train_files = [*dog_train_files,*cat_train_files]
dog_test_files = glob.glob(test_dogs+'*.jpg')
cat_test_files = glob.glob(test_cats+'*.jpg')
all_test_files = [*dog_test_files,*cat_test_files]
# print(len(img_filenames))
# img_filenames[:10]
all_filename_vars = [dog_train_files, cat_train_files,
dog_test_files,cat_test_files]
# -
|
.ipynb_checkpoints/Pneumonia_detection_v2-Colab-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/nsriniva/DS-Unit-2-Applied-Modeling/blob/master/module4-model-interpretation/LS_DS_234_assignment.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="oHW-SqodnJxH"
#
# Lambda School Data Science
#
# *Unit 2, Sprint 3, Module 4*
#
# ---
# + [markdown] id="y9XKLobPnJxM"
# # Model Interpretation
#
# You will use your portfolio project dataset for all assignments this sprint.
#
# ## Assignment
#
# Complete these tasks for your project, and document your work.
#
# - [ ] Continue to iterate on your project: data cleaning, exploratory visualization, feature engineering, modeling.
# - [ ] Make at least 1 partial dependence plot to explain your model.
# - [ ] Make at least 1 Shapley force plot to explain an individual prediction.
# - [ ] **Share at least 1 visualization (of any type) on Slack!**
#
# If you aren't ready to make these plots with your own dataset, you can practice these objectives with any dataset you've worked with previously. Example solutions are available for Partial Dependence Plots with the Tanzania Waterpumps dataset, and Shapley force plots with the Titanic dataset. (These datasets are available in the data directory of this repository.)
#
# Please be aware that **multi-class classification** will result in multiple Partial Dependence Plots (one for each class), and multiple sets of Shapley Values (one for each class).
# + [markdown] id="WEGJYivXnJxN"
# ## Stretch Goals
#
# #### Partial Dependence Plots
# - [ ] Make multiple PDPs with 1 feature in isolation.
# - [ ] Make multiple PDPs with 2 features in interaction.
# - [ ] Use Plotly to make a 3D PDP.
# - [ ] Make PDPs with categorical feature(s). Use Ordinal Encoder, outside of a pipeline, to encode your data first. If there is a natural ordering, then take the time to encode it that way, instead of random integers. Then use the encoded data with pdpbox. Get readable category names on your plot, instead of integer category codes.
#
# #### Shap Values
# - [ ] Make Shapley force plots to explain at least 4 individual predictions.
# - If your project is Binary Classification, you can do a True Positive, True Negative, False Positive, False Negative.
# - If your project is Regression, you can do a high prediction with low error, a low prediction with low error, a high prediction with high error, and a low prediction with high error.
# - [ ] Use Shapley values to display verbal explanations of individual predictions.
# - [ ] Use the SHAP library for other visualization types.
#
# The [SHAP repo](https://github.com/slundberg/shap) has examples for many visualization types, including:
#
# - Force Plot, individual predictions
# - Force Plot, multiple predictions
# - Dependence Plot
# - Summary Plot
# - Summary Plot, Bar
# - Interaction Values
# - Decision Plots
#
# We just did the first type during the lesson. The [Kaggle microcourse](https://www.kaggle.com/dansbecker/advanced-uses-of-shap-values) shows two more. Experiment and see what you can learn!
# + [markdown] id="PIbYtRjOnJxN"
# ### Links
#
# #### Partial Dependence Plots
# - [Kaggle / <NAME>: Machine Learning Explainability — Partial Dependence Plots](https://www.kaggle.com/dansbecker/partial-plots)
# - [<NAME>: Interpretable Machine Learning — Partial Dependence Plots](https://christophm.github.io/interpretable-ml-book/pdp.html) + [animated explanation](https://twitter.com/ChristophMolnar/status/1066398522608635904)
# - [pdpbox repo](https://github.com/SauceCat/PDPbox) & [docs](https://pdpbox.readthedocs.io/en/latest/)
# - [Plotly: 3D PDP example](https://plot.ly/scikit-learn/plot-partial-dependence/#partial-dependence-of-house-value-on-median-age-and-average-occupancy)
#
# #### Shapley Values
# - [Kaggle / <NAME>: Machine Learning Explainability — SHAP Values](https://www.kaggle.com/learn/machine-learning-explainability)
# - [<NAME>: Interpretable Machine Learning — Shapley Values](https://christophm.github.io/interpretable-ml-book/shapley.html)
# - [SHAP repo](https://github.com/slundberg/shap) & [docs](https://shap.readthedocs.io/en/latest/)
# + id="Aph78LJWnJxN"
# %%capture
import sys
# If you're on Colab:
if 'google.colab' in sys.modules:
DATA_PATH = 'https://raw.githubusercontent.com/LambdaSchool/DS-Unit-2-Applied-Modeling/master/data/'
# !pip install category_encoders==2.*
# !pip install eli5
# !pip install pdpbox
# !pip install shap
# If you're working locally:
else:
DATA_PATH = '../data/'
from collections import OrderedDict
from math import isclose
import zipfile
from urllib.request import urlopen
import io
import requests
from bs4 import BeautifulSoup
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from scipy import stats
from scipy.stats import chi2_contingency
from sklearn.preprocessing import StandardScaler
from sklearn.decomposition import PCA
from sklearn.cluster import KMeans
from sklearn.linear_model import LinearRegression
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
from category_encoders import OrdinalEncoder, OneHotEncoder
from sklearn.impute import SimpleImputer
from sklearn.preprocessing import StandardScaler
from sklearn.pipeline import make_pipeline
from sklearn.tree import DecisionTreeClassifier
from xgboost import XGBClassifier
from sklearn.metrics import accuracy_score, roc_auc_score
import eli5
from eli5.sklearn import PermutationImportance
import shap
from pdpbox.pdp import pdp_isolate, pdp_plot, pdp_interact, pdp_interact_plot
# For details about the data cleanup, please see
# https://github.com/nsriniva/DS-Unit-2-Applied-Modeling/blob/master/CleanupOnlineNewsPopularity.ipynb
# and 'The Dataset' section of
# https://nsriniva.github.io/2020-10-23-DSPT9-Unit1-BuildProject/
# Cleaned up and uploaded csv data file from
# https://archive.ics.uci.edu/ml/machine-learning-databases/00332/OnlineNewsPopularity.zip
# in
# https://archive.ics.uci.edu/ml/datasets/Online+News+Popularity
# to my github repo as
# https://github.com/nsriniva/DS-Unit-2-Applied-Modeling/blob/master/OnlineNewsPopularity.csv.zip?raw=true
# The associated names file is available at
# https://raw.githubusercontent.com/nsriniva/DS-Unit-2-Applied-Modeling/master/OnlineNewsPopularity.names
onp_url = 'https://github.com/nsriniva/DS-Unit-2-Applied-Modeling/blob/master/OnlineNewsPopularity.csv.zip?raw=true'
onp_df = pd.read_csv(onp_url, compression='zip')
null_values = onp_df.isna().sum().sum()
print(f"There are {['','no'][int(null_values==0)]} invalid values in the dataset!")
# The zscore() method from the scipy.stats package is used to compute z scores
# for the shares values. These z scores is compared against the specified
# sigma value to generate a boolean filter array that could be used to
# paritition the dataset based on whether the zscore is greater than the
# specified sigma.
def get_sigma_filter(df, sigma=0.5):
z = np.abs(stats.zscore(df.shares))
return np.where(z>sigma)[0]
# Use the boolean filter array provided by get_sigma_filter() to
# ignore entries with zscore greater than 0.5 and compute the
# median and max 'shares' values for the remaining entries.
def classification_marks(df):
shares_info = df.drop(get_sigma_filter(df)).shares
max = shares_info.max()
median = shares_info.median()
return median, max
shares_median = onp_df.shares.median()
print(shares_median)
# Use the medium(median) value to classify articles into
# unpopular(0) and popular(1)
onp_df['popularity'] = onp_df.shares.apply(lambda x: 0 if x < shares_median else 1)
display(onp_df.shape)
# Remove outliers
def remove_outliers(df, sigma=0.5):
df = df.copy()
return df.drop(get_sigma_filter(df, sigma))
onp_no_df = onp_df.copy()
#onp_no_df = remove_outliers(onp_no_df, 0.25)
shares_median = onp_no_df.shares.median()
print(shares_median)
# Use the medium(median) value to classify articles into
# unpopular(0) and popular(1)
onp_no_df['popularity'] = onp_no_df.shares.apply(lambda x: 0 if x < shares_median else 1)
display(onp_no_df.shape)
# The baseline accuracy or the value we'd get by just guessing that that the
# value is always the majority class
target = 'popularity'
baseline_accuracy = onp_no_df[target].value_counts(normalize=True).max()
print(f'baseline_accuracy = {baseline_accuracy:0.4f}')
# Drop the 'shares' column used to derive 'popularity' along
# with the non predictive 'url' and 'timedelta' columns.
drop_cols = ['shares', 'url', 'timedelta']
onp_no_df = onp_no_df.drop(columns=drop_cols)
# Will use a random split of 64% Training, 16% Validation and 20% Test
X = onp_no_df.drop(columns=target)
y = onp_no_df[target]
X_train_val, X_test, y_train_val, y_test = train_test_split(X,y,train_size=0.8, random_state=42)
X_train, X_val, y_train, y_val = train_test_split(X_train_val, y_train_val, train_size=0.8, random_state=42)
display(X_train.shape, X_val.shape, X_test.shape, y_train.shape, y_val.shape, y_test.shape)
display(y_train.value_counts(normalize=True))
baseline_accuracy = y_train.value_counts(normalize=True).max()
print(f'baseline_accuracy = {baseline_accuracy:0.4f}')
# Simple model, with OrdinalEncoder for the data_channel and weekday categorical
# columns and a DecisionTreeClassifier with default parameter values.
model = make_pipeline(
OrdinalEncoder(),
DecisionTreeClassifier()
)
model.fit(X_train, y_train)
display(y_train.value_counts(normalize=True))
display(y_val.value_counts(normalize=True))
training_bl = y_train.value_counts(normalize=True).max()
validation_bl = y_val.value_counts(normalize=True).max()
training_acc = model.score(X_train, y_train)
validation_acc = model.score(X_val, y_val)
print(f'Training Accuracy:{training_acc:0.4f}/{training_bl:0.4f}')
print(f'Validation Accuracy:{validation_acc:0.4f}/{validation_bl:0.4f}')
transformers = make_pipeline(
OrdinalEncoder(),
SimpleImputer(strategy='median')
)
X_train_transformed = transformers.fit_transform(X_train)
X_val_transformed = transformers.transform(X_val)
model = RandomForestClassifier(n_estimators=103, random_state=42, n_jobs=-1, max_depth=25, min_samples_leaf=3, max_features=0.3)
model.fit(X_train_transformed, y_train)
permuter = PermutationImportance(
model,
scoring='accuracy',
n_iter=5,
random_state=42
)
permuter.fit(X_val_transformed, y_val)
# + colab={"base_uri": "https://localhost:8080/", "height": 833} id="mMGL_S0_OGvb" outputId="5e64c964-1c00-4c99-c059-c0346215f4ce"
feature_names = X_val.columns.tolist()
eli5.show_weights(
permuter,
top=None, # No limit: show permutation importances for all features
feature_names=feature_names # must be a list
)
# + colab={"base_uri": "https://localhost:8080/"} id="p-TamhqFC8Cv" outputId="2060b5b4-038d-4773-e15e-c26ab9804097"
print('Shape before removing', X_train.shape)
minimum_importance = 0
mask = permuter.feature_importances_ > minimum_importance
features = X_train.columns[mask]
X_train = X_train[features]
print('Shape after removing ', X_train.shape)
X_val = X_val[features]
X_test = X_test[features]
# + colab={"base_uri": "https://localhost:8080/", "height": 153} id="r6o0TRJkqKJe" outputId="d3da60ff-1b90-4b0c-d5ae-d944d4f1ec80"
model = make_pipeline(
OrdinalEncoder(),
DecisionTreeClassifier(max_depth=7,random_state=42, min_samples_leaf=3)
)
model.fit(X_train, y_train)
display(y_train.value_counts(normalize=True))
display(y_val.value_counts(normalize=True))
training_bl = y_train.value_counts(normalize=True).max()
validation_bl = y_val.value_counts(normalize=True).max()
training_acc = model.score(X_train, y_train)
validation_acc = model.score(X_val, y_val)
print(f'Training Accuracy:{training_acc:0.4f}/{training_bl:0.4f}')
print(f'Validation Accuracy:{validation_acc:0.4f}/{validation_bl:0.4f}')
# + colab={"base_uri": "https://localhost:8080/"} id="hGorSrqgo5jE" outputId="c1dc7ea4-824e-4a34-abc2-d446b296e8ac"
pipe_elems = (
OrdinalEncoder(),
SimpleImputer(strategy='median'),
RandomForestClassifier(n_estimators=103, random_state=42, n_jobs=-1, max_depth=25, min_samples_leaf=3, max_features=0.3)
)
pipe = make_pipeline(
*pipe_elems
)
# Fit on train, score on val
pipe.fit(X_train, y_train)
print('Validation Accuracy', pipe.score(X_val, y_val))
print('Test Accuracy', pipe.score(X_test, y_test))
# + id="qtygtjfdnJxO" colab={"base_uri": "https://localhost:8080/"} outputId="883bc982-cfcd-411e-a470-d4286738004a"
encoder = OrdinalEncoder()
X_train_encoded = encoder.fit_transform(X_train)
X_val_encoded = encoder.transform(X_val)
X_test_encoded = encoder.transform(X_test)
eval_set = [(X_train_encoded, y_train),
(X_val_encoded, y_val)]
model = XGBClassifier(n_estimators=1000, random_state=42, n_jobs=-1, max_depth=6, learning_rate=0.5)
eval_metric = 'error'
model.fit(X_train_encoded, y_train,
eval_set=eval_set,
eval_metric=eval_metric,
early_stopping_rounds=500)
y_pred = model.predict(X_val_encoded)
print('Validation Accuracy', accuracy_score(y_val, y_pred))
# + colab={"base_uri": "https://localhost:8080/", "height": 296} id="-ulMUDYCydEw" outputId="dfa89588-8966-4012-8fba-74720dd811c3"
results = model.evals_result()
train_error = results['validation_0'][eval_metric]
val_error = results['validation_1'][eval_metric]
epoch = list(range(1, len(train_error)+1))
plt.plot(epoch, train_error, label='Train')
plt.plot(epoch, val_error, label='Validation')
plt.ylabel(f'Classification {eval_metric.capitalize()}')
plt.xlabel('Model Complexity (n_estimators)')
plt.title('Validation Curve for this XGBoost model')
#plt.ylim((0.18, 0.22)) # Zoom in
plt.legend();
# + colab={"base_uri": "https://localhost:8080/"} id="r_Is5nvJYhu2" outputId="3499f270-168a-492b-b11c-1f8de809a4dc"
class_index = 1
y_pred_proba = model.predict_proba(X_test_encoded)[:, class_index]
print(f'Test ROC AUC for class {class_index}:')
print(roc_auc_score(y_test, y_pred_proba)) # Ranges from 0-1, higher is better
# + colab={"base_uri": "https://localhost:8080/"} id="QxwQseqJ9aLb" outputId="1af0dc55-50b5-4623-85b8-a1b792f9cba8"
features = ['is_weekend', 'kw_avg_avg']
print(X_train_encoded.shape, X_val_encoded.shape)
isolated = []
for feature in features:
isolated.append(
pdp_isolate(
model=model,
dataset=X_val_encoded,
model_features=X_val_encoded.columns,
feature=feature
)
)
interaction = pdp_interact(
model=model,
dataset=X_val_encoded,
model_features=X_val_encoded.columns,
features=features
)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="NgflqC9aeaKJ" outputId="99ea50b7-31cd-4258-f238-28cc487144b1"
for idx,elem in enumerate(isolated):
pdp_plot(elem, feature_name=features[idx]);
if features[idx] == 'is_weekend':
# Manually change the xticks labels
plt.xticks([0, 1], ['False', 'True']);
pdp = interaction.pdp.pivot_table(
values='preds',
columns=features[0], # First feature on x axis
index=features[1] # Next feature on y axis
)[::-1] # Reverse the index order so y axis is ascending
pdp = pdp.rename(columns={0:'False', 1:'True'})
plt.figure(figsize=(10,8))
sns.heatmap(pdp, annot=True, fmt='.3f', cmap='viridis')
plt.title('Partial Dependence of Article Popularity on is_weekend & kw_avg_avg');
# + colab={"base_uri": "https://localhost:8080/", "height": 287} id="vjGLK7WfcWrg" outputId="7e938830-924c-4bc5-f7b6-927b91453ec9"
display(X_test_encoded.head())
row = X_test_encoded.loc[[198]]
row
# + colab={"base_uri": "https://localhost:8080/", "height": 193} id="Wz1Avu_wxWbR" outputId="e935987e-80c7-46b9-b9a8-aa5646ef5d7a"
explainer = shap.TreeExplainer(model)
shap_values = explainer.shap_values(row)
shap.initjs()
shap.force_plot(
base_value=explainer.expected_value,
shap_values=shap_values,
features=row,
#matplotlib=True, # This does not work if link is set to 'logit'
link='logit' # For classification, this shows predicted probabilities
)
|
module4-model-interpretation/LS_DS_234_assignment.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# FIXME : 以下の関数は定義されたファイルの形式に依存するので、utilsに記載できない。
def is_env_notebook():
"""Determine wheather is the environment Jupyter Notebook"""
if 'get_ipython' not in globals():
# Python shell
return False
env_name = get_ipython().__class__.__name__
if env_name == 'TerminalInteractiveShell':
# IPython shell
return False
# Jupyter Notebook
return True
# +
import argparse
from itertools import islice
import json
from pathlib import Path
import shutil
import warnings
from typing import Dict
import os
import numpy as np
import pandas as pd
from sklearn.metrics import fbeta_score
from sklearn.exceptions import UndefinedMetricWarning
import torch
from torch import nn, cuda
from torch.optim import Adam
import tqdm
import torch.nn.functional as F
from sklearn.metrics import cohen_kappa_score
from IPython.core.debugger import Pdb
# +
ON_KAGGLE: bool = 'KAGGLE_WORKING_DIR' in os.environ
if ON_KAGGLE:
from . import models
from .dataset import TrainDataset, TTADataset, get_ids, N_CLASSES, DATA_ROOT
from .transforms import train_transform, test_transform
from .utils import (
write_event, load_model, mean_df, ThreadingDataLoader as DataLoader,
ON_KAGGLE)
else:
import models
from dataset import TrainDataset, TTADataset, get_ids, N_CLASSES, DATA_ROOT
from transforms import train_transform, test_transform
from utils import (
write_event, load_model, mean_df, ThreadingDataLoader as DataLoader,
ON_KAGGLE)
# +
def main(*args):
#def main():
# print("do main")
parser = argparse.ArgumentParser()
arg = parser.add_argument
# TODO : "--modeにGradCAMを追加"
arg('--mode', choices=['train', 'validate', 'predict_valid', 'predict_test'])
arg('--run_root')
arg('--model', default='resnet50')
arg('--loss',default="focalloss")
arg('--pretrained', type=int, default=1)
arg('--batch-size', type=int, default=64)
arg('--step', type=int, default=1)
arg('--workers', type=int, default=4 if ON_KAGGLE else 4)
arg('--lr', type=float, default=1e-4)
arg('--patience', type=int, default=4)
arg('--clean', action='store_true')
arg('--n-epochs', type=int, default=100)
arg('--epoch-size', type=int)
arg('--tta', type=int, default=4)
arg('--use-sample', action='store_true', help='use a sample of the dataset')
arg('--debug', action='store_true')
arg('--limit', type=int)
arg('--fold', type=int, default=0)
arg('--regression',type=int,default=0)
arg('--finetuning',type=int,default=1)
# TODO : classificationかregressionかをオプションで追加できるようにする。
# from IPython.core.debugger import Pdb; Pdb().set_trace()
if is_env_notebook():
args = parser.parse_args(args=args[0])
else:
args = parser.parse_args()
# from IPython.core.debugger import Pdb; Pdb().set_trace()
run_root = Path(args.run_root)
folds = pd.read_csv('folds.csv')
train_root = DATA_ROOT / ('train_sample' if args.use_sample else 'train_images')
if args.use_sample:
folds = folds[folds['Id'].isin(set(get_ids(train_root)))]
# Pdb().set_trace()
# -1 はleakデータ
train_fold = folds[folds['fold'] != args.fold]
leak_fold = folds[folds['fold'] == -1]
train_fold = pd.concat([train_fold,leak_fold])
valid_fold = folds[folds['fold'] == args.fold]
if args.limit:
train_fold = train_fold[:args.limit]
valid_fold = valid_fold[:args.limit]
def make_loader(df: pd.DataFrame, image_transform,regression=args.regression,shuffle=False,balanced=True) -> DataLoader:
return DataLoader(
TrainDataset(train_root, df, image_transform, debug=args.debug,regression=regression,balanced=balanced),
shuffle=shuffle,
batch_size=args.batch_size,
num_workers=args.workers,
)
## TODO : regressionようにモデルを書き換え
if args.regression:
criterion = nn.MSELoss()
# TODO : 回帰モデルへ変更
model = getattr(models, args.model)(
num_classes=1, pretrained=args.pretrained)
else:
# 分類モデル
criterion = FocalLoss()#nn.BCEWithLogitsLoss(reduction='none')
model = getattr(models, args.model)(
num_classes=N_CLASSES, pretrained=args.pretrained)
# Pdb().set_trace()
use_cuda = cuda.is_available()
fresh_params = list(model.fresh_params())
all_params = list(model.parameters())
if use_cuda:
model = model.cuda()
if args.mode == 'train':
if run_root.exists() and args.clean:
shutil.rmtree(run_root)
run_root.mkdir(exist_ok=True, parents=True)
(run_root / 'params.json').write_text(
json.dumps(vars(args), indent=4, sort_keys=True))
train_loader = make_loader(train_fold, train_transform,regression=args.regression,balanced=True)
valid_loader = make_loader(valid_fold, test_transform,regression=args.regression,balanced=False)
print(f'{len(train_loader.dataset):,} items in train, '
f'{len(valid_loader.dataset):,} in valid')
train_kwargs = dict(
args=args,
model=model,
criterion=criterion,
train_loader=train_loader,
valid_loader=valid_loader,
patience=args.patience,
init_optimizer=lambda params, lr: Adam(params, lr),
use_cuda=use_cuda,
)
# from IPython.core.debugger import Pdb; Pdb().set_trace()
if args.pretrained:
if train(params=fresh_params, n_epochs=1, **train_kwargs):
train(params=all_params, **train_kwargs)
else:
train(params=all_params, **train_kwargs)
# fine-tunig after balanced learning
if args.finetuning:
print("Start Fine-tuning")
TUNING_EPOCH = 5
train_loader = make_loader(train_fold, train_transform,regression=args.regression,balanced=False)
# 学習率を小さくする
args.lr = args.lr / 5
train_kwargs["train_loader"] = train_loader
train(params=all_params,n_epochs=args.n_epochs+TUNING_EPOCH,**train_kwargs,finetuning=args.finetuning)
elif args.mode == 'validate':
valid_loader = make_loader(valid_fold, test_transform)
load_model(model, run_root / 'model.pt')
validation(model, criterion, tqdm.tqdm(valid_loader, desc='Validation',valid_fold=valid_fold),
use_cuda=use_cuda)
elif args.mode.startswith('predict'):
print("load model predict")
load_model(model, run_root / 'best-model.pt')
predict_kwargs = dict(
batch_size=args.batch_size,
tta=args.tta,
use_cuda=use_cuda,
workers=args.workers,
)
if args.mode == 'predict_valid':
#predict(model, df=valid_fold, root=train_root,
# out_path=run_root / 'val.h5',
# **predict_kwargs)
valid_loader = make_loader(valid_fold, test_transform,shuffle=False,balanced=False)
#model: nn.Module, criterion, valid_loader, use_cuda,valid_predict:bool=False
# TODO : valid foldに予測結果をくっ付ける操作を追加
validation(model,criterion,valid_loader,use_cuda,valid_fold=valid_fold,valid_predict=True,save_path=run_root)
elif args.mode == 'predict_test':
test_root = DATA_ROOT / (
'test_sample' if args.use_sample else 'test_images')
ss = pd.read_csv(DATA_ROOT / 'sample_submission.csv')
if args.use_sample:
ss = ss[ss['id'].isin(set(get_ids(test_root)))]
if args.limit:
ss = ss[:args.limit]
predict(model, df=ss, root=test_root,
out_path=run_root / 'test.h5',
**predict_kwargs)
def predict(model, root: Path, df: pd.DataFrame, out_path: Path,
batch_size: int, tta: int, workers: int, use_cuda: bool):
loader = DataLoader(
dataset=TTADataset(root, df, test_transform, tta=tta),
shuffle=False,
batch_size=batch_size,
num_workers=workers,
)
model.eval()
all_outputs, all_ids = [], []
with torch.no_grad():
for inputs, ids in tqdm.tqdm(loader, desc='Predict'):
if use_cuda:
inputs = inputs.cuda()
outputs = torch.sigmoid(model(inputs))
all_outputs.append(outputs.data.cpu().numpy())
all_ids.extend(ids)
df = pd.DataFrame(
data=np.concatenate(all_outputs),
index=all_ids,
columns=map(str, range(N_CLASSES)))
df = mean_df(df)
df.to_hdf(out_path, 'prob', index_label='id')
print(f'Saved predictions to {out_path}')
def train(args, model: nn.Module, criterion, *, params,
train_loader, valid_loader, init_optimizer, use_cuda,
n_epochs=None, patience=2, max_lr_changes=2,finetuning=False) -> bool:
lr = args.lr
n_epochs = n_epochs or args.n_epochs
params = list(params)
optimizer = init_optimizer(params, lr)
run_root = Path(args.run_root)
model_path = run_root / 'model.pt'
best_model_path = run_root / 'best-model.pt'
if model_path.exists():
state = load_model(model, model_path)
epoch = state['epoch']
step = state['step']
best_valid_loss = state['best_valid_loss']
if best_model_path.exists() and finetuning:
state = load_model(model,best_model_path)
epoch = 1
step = 0
# epoch = state['epoch']
# step = state['step']
best_valid_loss = state['best_valid_loss']
else:
epoch = 1
step = 0
best_valid_loss = float('inf')
lr_changes = 0
save = lambda ep: torch.save({
'model': model.state_dict(),
'epoch': ep,
'step': step,
'best_valid_loss': best_valid_loss
}, str(model_path))
report_each = 10
log = run_root.joinpath('train.log').open('at', encoding='utf8')
valid_losses = []
lr_reset_epoch = epoch
for epoch in range(epoch, n_epochs + 1):
model.train()
tq = tqdm.tqdm(total=(args.epoch_size or
len(train_loader) * args.batch_size))
tq.set_description(f'Epoch {epoch}, lr {lr}')
losses = []
tl = train_loader
# from IPython.core.debugger import Pdb; Pdb().set_trace()
if args.epoch_size:
tl = islice(tl, args.epoch_size // args.batch_size)
try:
mean_loss = 0
# Pdb().set_trace()
for i, (inputs, targets) in enumerate(tl):
if use_cuda:
inputs, targets = inputs.cuda(), targets.cuda()
outputs = model(inputs)
loss = criterion(outputs, targets)#_reduce_loss(criterion(outputs, targets))
batch_size = inputs.size(0)
(batch_size * loss).backward()
if (i + 1) % args.step == 0:
optimizer.step()
optimizer.zero_grad()
step += 1
tq.update(batch_size)
losses.append(loss.item())
mean_loss = np.mean(losses[-report_each:])
tq.set_postfix(loss=f'{mean_loss:.3f}')
if i and i % report_each == 0:
write_event(log, step, loss=mean_loss)
# Pdb().set_trace()
write_event(log, step, loss=mean_loss)
tq.close()
save(epoch + 1)
valid_metrics = validation(model, criterion, valid_loader, use_cuda)
# Pdb().set_trace()
write_event(log, step, **valid_metrics)
valid_loss = valid_metrics['valid_loss']
valid_losses.append(valid_loss)
# from IPython.core.debugger import Pdb; Pdb().set_trace()
if valid_loss < best_valid_loss:
best_valid_loss = valid_loss
shutil.copy(str(model_path), str(best_model_path))
elif (patience and epoch - lr_reset_epoch > patience and
min(valid_losses[-patience:]) > best_valid_loss):
# "patience" epochs without improvement
lr_changes +=1
if lr_changes > max_lr_changes:
break
lr /= 5
print(f'lr updated to {lr}')
lr_reset_epoch = epoch
optimizer = init_optimizer(params, lr)
except KeyboardInterrupt:
tq.close()
print('Ctrl+C, saving snapshot')
save(epoch)
print('done.')
return False
return True
def validation(
model: nn.Module, criterion, valid_loader, use_cuda,valid_fold=None,valid_predict:bool=False,save_path:Path=""
) -> Dict[str, float]:
model.eval()
all_losses, all_predictions, all_targets = [], [], []
with torch.no_grad():
for inputs, targets in valid_loader:
all_targets.append(targets.numpy().copy())
if use_cuda:
inputs, targets = inputs.cuda(), targets.cuda()
outputs = model(inputs)
loss = criterion(outputs, targets)
# all_losses.append(_reduce_loss(loss).item())
all_losses.append(loss.item())
predictions = torch.sigmoid(outputs)
all_predictions.append(predictions.cpu().numpy())
all_predictions = np.concatenate(all_predictions)
all_targets = np.concatenate(all_targets)
# Pdb().set_trace()
def get_score(y_pred):
with warnings.catch_warnings():
warnings.simplefilter('ignore', category=UndefinedMetricWarning)
# return fbeta_score(
# all_targets, y_pred, beta=2, average='samples')
return qk(y_pred,all_targets)
metrics = {}
#argsorted = all_predictions.argsort(axis=1)
# Pdb().set_trace()
#for threshold in [0.05, 0.10, 0.15, 0.20]:
# metrics[f'valid_f2_th_{threshold:.2f}'] = get_score(
# binarize_prediction(all_predictions, threshold, argsorted))
#metrics = get_score(all_predictions)
if valid_predict:
# Pdb().set_trace()
# TOOD : 予測結果をpd データフレーム形式で保存
valid_fold["prediction"] = all_predictions.argmax(axis=1)
valid_fold.to_csv(save_path / "valid_prediction.csv",index=False)
# run_root = Path(args.run_root)
with open(save_path / "best_score.txt",mode="w") as f:
f.write("best valid kapa : {score}".format(score=get_score(all_predictions)))
f.write("best valid loss : {loss}".format(loss=np.mean(all_losses)))
# from IPython.core.debugger import Pdb; Pdb().set_trace()
metrics['valid_kapa'] = get_score(all_predictions)
metrics['valid_loss'] = np.mean(all_losses)
#print(' | '.join(f'{k} {v:.3f}' for k, v in sorted(
# metrics.items(), key=lambda kv: -kv[1])))
print(metrics)
return metrics
def visualization():
"""
GRAD-CAMによるNNが判断の根拠としている領域の可視化
"""
pass
def binarize_prediction(probabilities, threshold: float, argsorted=None,
min_labels=1, max_labels=10):
""" Return matrix of 0/1 predictions, same shape as probabilities.
"""
assert probabilities.shape[1] == N_CLASSES
if argsorted is None:
argsorted = probabilities.argsort(axis=1)
max_mask = _make_mask(argsorted, max_labels)
min_mask = _make_mask(argsorted, min_labels)
prob_mask = probabilities > threshold
return (max_mask & prob_mask) | min_mask
def _make_mask(argsorted, top_n: int):
mask = np.zeros_like(argsorted, dtype=np.uint8)
col_indices = argsorted[:, -top_n:].reshape(-1)
row_indices = [i // top_n for i in range(len(col_indices))]
mask[row_indices, col_indices] = 1
return mask
def _reduce_loss(loss):
return loss.sum() / loss.shape[0]
class FocalLoss(nn.Module):
def __init__(self, gamma=2):
super().__init__()
self.gamma = gamma
def forward(self, logit, target):
target = target.float()
max_val = (-logit).clamp(min=0)
loss = logit - logit * target + max_val + \
((-max_val).exp() + (-logit - max_val).exp()).log()
invprobs = F.logsigmoid(-logit * (target * 2.0 - 1.0))
loss = (invprobs * self.gamma).exp() * loss
if len(loss.size())==2:
loss = loss.sum(dim=1)
return loss.mean()
def qk(y_pred, y):
## Pdb().set_trace()
#y_pred = torch.from_numpy(y_pred)
y_pred = np.argmax(y_pred,axis=1)
y = np.argmax(y,axis=1)
#y = torch.argmax(y,dim=1)
# Pdb().set_trace()
return cohen_kappa_score(y_pred, y, weights='quadratic')
#return torch.tensor(cohen_kappa_score(torch.round(y_pred), y, weights='quadratic'), device='cuda:0')
# -
if __name__ == '__main__' and ON_KAGGLE:
main()
if __name__ == '__main__' and not(ON_KAGGLE):
import gc
###########################################################
# FOLDを修正
folds = [0,1,2,3]
#1,2,3]#[0,1,
# N_EPOCH = 25
# model_name = "02_brightness_cotrast"
#model_name = "HueSaturationValue"
#model_name = "RandomGamma"
# model_name = "regression"
# model_name = "RandomSizeCrop_validation"
# model_name = "CentorCrop_Oneof"
#model_name = "CentorCrop_Rotation_Val"
# model_name = "Rockman_aug_nonCircleCrop"
# model_name = "Rockman_aug_CircleCrop"
# model_name = "10_test"
# model_name = "11_No_Crop_balanced_finetuning"
# model_name = "12_add_11_scale_and_gamma"
# model_name = "13_nodup"
# model_name = "14_nodup_refine"
model_name = "15_nodup_dup_leak"
N_EPOCH = 10
# limit変更
for fold in folds:
# 学習
# jupyter-notebookの場合、ここで引数を選択しないといけない。
train_args = ["--mode","train",
"--run_root","{model_name}_{fold}".format(model_name=model_name,fold=fold),
# "--limit","100", # TODO : 適宜変更
"--fold","{fold}".format(fold=fold),
"--n-epochs","{epoch}".format(epoch=N_EPOCH),
'--workers',"16",
'--patience',"2",
"--finetuning","1"
# "--regression","1"
]
main(train_args)
# validation
val_args = ["--mode","predict_valid",
"--run_root","{model_name}_{fold}".format(model_name=model_name,fold=fold),
# "--limit","100"
]
main(val_args)
gc.collect()
# break
#print(N_CLASSES)
if __name__ == '__main__' and not(ON_KAGGLE):
model_name = "CentorCrop_Rotation_Val/CentorCrop_Rotation_Val_0"
# jupyter-notebookの場合、ここで引数を選択しないといけない。
arg_list = ["--mode","predict_test",
"--run_root",model_name,
# "--limit","100",
"--tta","4"
]
main(arg_list)
#print(N_CLASSES)
if __name__ == '__main__' and not(ON_KAGGLE):
# jupyter-notebookの場合、ここで引数を選択しないといけない。
model_name = model_name + "_0"
arg_list = ["--mode","predict_valid",
"--run_root",model_name,
"--limit","100"]
main(arg_list)
|
apotos/main.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] _cell_guid="6a09d4fb-60c5-4f45-b844-8c788a50c543" _uuid="8e892e637f005dd61ec7dcb95865e52f3de2a77f" id="qZH6mu82AbET"
# # Titanic: Machine Learning from Disaster
# ### Predict survival on the Titanic
# - Defining the problem statement
# - Collecting the data
# - Exploratory data analysis
# - Feature engineering
# - Modelling
# - Testing
# + [markdown] _cell_guid="4af5e83d-7fd8-4a61-bf26-9583cb6d3476" _uuid="65d04d276a8983f62a49261f6e94a02b281dbcc9" id="fP9ReT8gAbEW"
# ## 1. Defining the problem statement
# Complete the analysis of what sorts of people were likely to survive.
# In particular, we ask you to apply the tools of machine learning to predict which passengers survived the Titanic tragedy.
# + colab={"base_uri": "https://localhost:8080/", "height": 643} id="HzV4wiu8AbEW" outputId="f5451050-8f6a-4209-ca67-ccd2f777f09a"
from IPython.display import Image
Image(url= "https://static1.squarespace.com/static/5006453fe4b09ef2252ba068/5095eabce4b06cb305058603/5095eabce4b02d37bef4c24c/1352002236895/100_anniversary_titanic_sinking_by_esai8mellows-d4xbme8.jpg")
# + [markdown] _cell_guid="3f529075-7f9b-40ff-a79a-f3a11a7d8cbe" _uuid="64ca0f815766e3e8074b0e04f53947930cb061aa" id="V50LyJS4AbEX"
# ## 2. Collecting the data
#
# training data set and testing data set are given by Kaggle
# you can download from
# my github [https://github.com/minsuk-heo/kaggle-titanic/tree/master](https://github.com/minsuk-heo/kaggle-titanic)
# or you can download from kaggle directly [kaggle](https://www.kaggle.com/c/titanic/data)
#
# ### load train, test dataset using Pandas
# + _cell_guid="e58a3f06-4c2a-4b87-90de-f8b09039fd4e" _uuid="46f0b12d7bf66712642e9a9b807f5ef398426b83" id="zyh-_vfJAbEY"
import pandas as pd
train = pd.read_csv('https://raw.githubusercontent.com/lab-ml-itba/Arboles-de-decision/master/data/train.csv')
# +
# import wget
# url='https://raw.githubusercontent.com/lab-ml-itba/Arboles-de-decision/master/data/train.csv'
# wget.download(url)
# +
# import pandas as pd
# train = pd.read_csv('/train.csv')
# + [markdown] _cell_guid="836a454f-17bc-41a2-be69-cd86c6f3b584" _uuid="1ed3ad39ead93977b8936d9c96e6f6f806a8f9b3" id="WuaoR0nFAbEY"
# ## 3. Exploratory data analysis
# Printing first 5 rows of the train dataset.
# + _cell_guid="749a3d70-394c-4d2c-999a-4d0567e39232" _uuid="b9fdb3b19d7a8f30cd0bb69ae434e04121ecba93" colab={"base_uri": "https://localhost:8080/", "height": 205} id="QfDgmgqUAbEZ" outputId="c6e2ffad-dd99-4bf8-91f1-11dd4ea3a7f8"
train.head()
# -
train.describe()
# + [markdown] id="Orr9rEHAAbEZ"
# ### Data Dictionary
# - Survived: 0 = No, 1 = Yes
# - pclass: Ticket class 1 = 1st, 2 = 2nd, 3 = 3rd
# - sibsp: # of siblings / spouses aboard the Titanic
# - parch: # of parents / children aboard the Titanic
# - ticket: Ticket number
# - cabin: Cabin number
# - embarked: Port of Embarkation C = Cherbourg, Q = Queenstown, S = Southampton
# + [markdown] _cell_guid="5ebc1e0e-2b5a-4d92-98e0-defa019d4439" _uuid="1892fbb34b26d775d1c428fdb7b6254449286b28" id="r3fBZINrAbEa"
# **Total rows and columns**
#
# We can see that there are 891 rows and 12 columns in our training dataset.
# + _cell_guid="ed1e7849-d1b6-490d-b86b-9ca71dfafc7d" _uuid="5a641beccf0e555dfd7b9a53a17188ea6edef95b" colab={"base_uri": "https://localhost:8080/"} id="9KrORSwaAbEa" outputId="7dfa22be-d32f-4001-f45d-1c6570c42066"
train.shape
# + _cell_guid="418b8a69-f2aa-442d-8f45-fa8887190938" _uuid="4ee2591110660a4a16b3da7a7530f0945e121b46" colab={"base_uri": "https://localhost:8080/"} id="8TqLfnuoAbEb" outputId="4cd8a6fd-91e8-4941-a264-dc9c65634fcc"
train.info()
# + [markdown] _cell_guid="abc3c4fc-6419-405f-927a-4214d2c73eec" _uuid="622d4d4b2ba8f77cc537af97fc343d4cd6de26b2" id="sdq4Nn6zAbEb"
# We can see that *Age* value is missing for many rows.
#
# Out of 891 rows, the *Age* value is present only in 714 rows.
#
# Similarly, *Cabin* values are also missing in many rows. Only 204 out of 891 rows have *Cabin* values.
# + _cell_guid="0663e2bb-dc27-4187-94b1-ff4ff78b68bc" _uuid="3bf74de7f2483d622e41608f6017f2945639e4df" colab={"base_uri": "https://localhost:8080/"} id="ICiCH5iTAbEb" outputId="e74dd840-ba8e-49a5-aacd-4aad9ef3d4b2"
train.isnull().sum()
# + [markdown] _cell_guid="176aa52d-fde8-42e6-a3ee-db31f8b0ca49" _uuid="b48a9feff6004d783960aa1b32fdfde902d87e21" id="m1fypBsgAbEc"
# There are 177 rows with missing *Age*, 687 rows with missing *Cabin* and 2 rows with missing *Embarked* information.
# + [markdown] _cell_guid="c8553d48-c5e0-4947-bd13-1b38509c850c" _uuid="1a28e607e9ed63cefe0f35a4e4d72f2f36299323" id="q55oew6WAbEd"
# ### import python lib for visualization
# + _cell_guid="b1d8a6d2-c22d-435c-8c98-973e8f41b138" _uuid="26411c710f69b29939c815d5f5ab01d9177df7d0" id="099TZgCFAbEd"
import matplotlib.pyplot as plt
# %matplotlib inline
import seaborn as sns
sns.set() # setting seaborn default for plots
# + [markdown] id="-iBDgB6DAbEd"
# ### Bar Chart for Categorical Features
# - Pclass
# - Sex
# - SibSp ( # of siblings and spouse)
# - Parch ( # of parents and children)
# - Embarked
# - Cabin
# + id="8DcYXusHAbEd"
def bar_chart(feature):
survived = train[train['Survived']==1][feature].value_counts()
dead = train[train['Survived']==0][feature].value_counts()
df = pd.DataFrame([survived,dead])
df.index = ['Survived','Dead']
df.plot(kind='bar',stacked=True, figsize=(10,5))
# + [markdown] id="0QUJh_lmLRqt"
# # 1) Seleccionar las correctas:
# La mayoría de los hombres sobrevivieron.
#
# La mayoría de las mujeres sobrevivieron. ``True.``
#
# La mayoría de los sobrevivientes fueron mujeres. ``True.``
#
# La mayoría de los sobrevivientes fueron hombres.
# + colab={"base_uri": "https://localhost:8080/", "height": 361} id="Wwl0cpsyAbEd" outputId="51aa2885-e078-4c59-e10b-27abf1b41562"
bar_chart('Sex')
# + colab={"base_uri": "https://localhost:8080/"} id="Y67oAVUD344Y" outputId="fe1d3e8f-a105-4137-c67a-61c2fbe6eba7"
# Cuantos sobrevivieron y cuantos murieron:
print(train['Survived'].value_counts())
# De los que sobrevivieron, cuantos eran mujeres y cuantos hombres:
print(train[train['Survived']==1]["Sex"].value_counts())
# + [markdown] id="9jnYskRZAbEe"
# # 2) ¿Qué clase sobrevivió mas?
# ``Clase 1``
#
# # 3) ¿Qué clase falleció mas?
# ``Clase 3``
# + colab={"base_uri": "https://localhost:8080/", "height": 361} id="SaQsIT7hGICO" outputId="49746b19-203b-42ee-a842-641ac638f751"
bar_chart('Pclass')
# + colab={"base_uri": "https://localhost:8080/"} id="w7UTKR3SO2GZ" outputId="e9c35d99-a58a-4300-ea84-f206b94fbbf4"
#Las clases que más sobrevivieron:
train[train['Survived']==1]["Pclass"].value_counts()
# + colab={"base_uri": "https://localhost:8080/"} id="CWHfIPu4QPeP" outputId="24864287-47c9-4398-8c0a-19f7923cf058"
#Las clases que más fallecierón:
train[train['Survived']==0]["Pclass"].value_counts()
# + [markdown] id="7cS0JQVUAbEe"
# # 4) Seleccione las verdaderas:
#
# Las personas pertenecientes a familias numerosas en general fallecieron.
#
# Las personas solas en general fallecieron. ``True``
#
# Las personas solas en general sobrevivieron.
#
# Las personas pertenecientes a familias numerosas en general Sobrevivieron.``True``
# + colab={"base_uri": "https://localhost:8080/", "height": 361} id="YGB3SrGTAbEe" outputId="6b3f2318-4de9-4002-b9d8-0626a93cec65"
# Realizar un gráfico de barras para para la variable "Pchart"
bar_chart('Parch')
# + colab={"base_uri": "https://localhost:8080/"} id="tfGXG5cLZyFH" outputId="777c38a4-e874-43f3-e449-c39b4d70d469"
# los que mas sobreviveron:
train[train['Survived']==1]["Parch"].value_counts()
# + colab={"base_uri": "https://localhost:8080/"} id="GFwwMPcUZ6iE" outputId="ba19d12e-127f-49b2-ebf2-bd74ca213056"
# los que mas fallecieron:
train[train['Survived']==0]["Parch"].value_counts()
# + colab={"base_uri": "https://localhost:8080/", "height": 361} id="uWkU0b3EAbEe" outputId="73a31d53-7e97-42b9-b685-998af46cd3c2"
# Realizar un gráfico de barras para la variable "SibSp"
bar_chart('SibSp')
# + colab={"base_uri": "https://localhost:8080/"} id="aE_g1RlCbBF5" outputId="e0473e55-c495-4cca-c12e-4818134917af"
# los que mas sobreviveron:
train[train['Survived']==1]["SibSp"].value_counts()
# + colab={"base_uri": "https://localhost:8080/"} id="hMV6r8ITbFNf" outputId="86a9c3fa-a496-4900-affc-9f99b75dc038"
# los que mas fallecieron:
train[train['Survived']==0]["SibSp"].value_counts()
# + [markdown] id="Y1_viZ0mAbEf"
# The Chart confirms **a person aboarded with more than 2 parents or children** more likely survived
# The Chart confirms ** a person aboarded alone** more likely dead
# + colab={"base_uri": "https://localhost:8080/", "height": 361} id="PznJFTfWAbEf" outputId="8f85d0d8-f865-4f7a-a16b-e16a53622bd4"
bar_chart('Embarked')
# + [markdown] id="q151pH-4AbEf"
# The Chart confirms **a person aboarded from C** slightly more likely survived
# The Chart confirms **a person aboarded from Q** more likely dead
# The Chart confirms **a person aboarded from S** more likely dead
# + [markdown] _cell_guid="810cd964-24eb-44fb-9e7b-18bbddd4900f" _uuid="fd86ccdf2d1248b79c68365444e96e46a50f3f5a" id="gzxWz0yKAbEf"
# ## 4. Feature engineering
#
# Feature engineering is the process of using domain knowledge of the data
# to create features (**feature vectors**) that make machine learning algorithms work.
#
# feature vector is an n-dimensional vector of numerical features that represent some object.
# Many algorithms in machine learning require a numerical representation of objects,
# since such representations facilitate processing and statistical analysis.
# + colab={"base_uri": "https://localhost:8080/", "height": 205} id="3M0uz91sAbEg" outputId="60695882-03cb-40e6-aa56-a4036c402ee0"
train.head()
# + [markdown] id="yjTfWnr5AbEg"
#
#
# ### 4.1 how titanic sank?
# sank from the bow of the ship where third class rooms located
# conclusion, Pclass is key feature for classifier
# + colab={"base_uri": "https://localhost:8080/", "height": 810} id="ceUlbVyqAbEg" outputId="a6df1133-3b1f-4a22-9766-19878555d93d"
Image(url= "https://static1.squarespace.com/static/5006453fe4b09ef2252ba068/t/5090b249e4b047ba54dfd258/1351660113175/TItanic-Survival-Infographic.jpg?format=1500w")
# + colab={"base_uri": "https://localhost:8080/", "height": 396} id="iEiUbQWHAbEg" outputId="32cebfb9-6804-474f-a147-d32c3b37bbc6"
train.head(10)
# + [markdown] id="gQacmMFBAbEh"
# ### 4.2 Name
# + id="CAY-3OkJAbEh"
# Se puede utilizar para obtener el título de una persona: Mr
train_test_data = [train] # combining train and test dataset
for dataset in train_test_data:
dataset['Title'] = dataset['Name'].str.extract(' ([A-Za-z]+)\.', expand=False)
# + colab={"base_uri": "https://localhost:8080/"} id="_S3dLyLjAbEh" outputId="18f07a45-c114-4659-c90e-1d4b2d30265f"
# Escribir una línea de código que muestre cuántas veces apareció cada título (value_counts)
train['Title'].value_counts()
# + [markdown] id="OlRE8SIEAbEh"
# # 5) Cuantas veces apareció:
#
# Mr ``517``
# Miss ``182``
# Mrs ``125``
#
# + [markdown] id="C5DANxZvAbEh"
# #### Title map
# Mr : 0
# Miss : 1
# Mrs: 2
# Others: 3
#
# + id="sYUsW1LFAbEh"
#we convert the categorical Title values into a numeric form.
title_mapping = {"Mr": 0, "Miss": 1, "Mrs": 2,
"Master": 3, "Dr": 3, "Rev": 3, "Col": 3, "Major": 3, "Mlle": 3,"Countess": 3,
"Ms": 3, "Lady": 3, "Jonkheer": 3, "Don": 3, "Dona" : 3, "Mme": 3,"Capt": 3,"Sir": 3 }
for dataset in train_test_data:
dataset['Title'] = dataset['Title'].map(title_mapping)
# + colab={"base_uri": "https://localhost:8080/"} id="F15mnTzOGRhu" outputId="2f77432c-d999-4f3f-cba3-cf8d8489fc3e"
dataset['Title'].value_counts()
# + colab={"base_uri": "https://localhost:8080/", "height": 257} id="GhD3y2QDAbEi" outputId="fcda3974-15c0-4282-d2e4-ea7ecbd6f871"
train.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 361} id="Wboeqj98AbEi" outputId="e597d4f2-198f-4ecd-b0b6-fcb3a4052bb1"
bar_chart('Title')
# + id="5En641u9AbEi"
# Borrar la columna Name del dataset
train.drop('Name', axis=1, inplace=True)
# + colab={"base_uri": "https://localhost:8080/", "height": 205} id="QgzRAYBsAbEi" outputId="feb62656-0c5b-48d1-81ae-b497b0e08ead"
train.head()
# + [markdown] id="n_sAeLhLAbEi"
# ### 4.3 Sex
#
# male: 0
# female: 1
# + id="Jm-l05D0AbEi"
# we convert the categorical Sex values into a numeric form.
sex_mapping = {"male": 0, "female": 1}
for dataset in train_test_data:
dataset['Sex'] = dataset['Sex'].map(sex_mapping)
# + colab={"base_uri": "https://localhost:8080/", "height": 361} id="_axH3iZhAbEi" outputId="29540d9e-d158-47e1-ea03-4b1e4f82ae65"
bar_chart('Sex')
# + [markdown] id="5TC9scgjAbEj"
# ### 4.4 Age
# + [markdown] id="i5gKJCqoAbEj"
# #### 4.4.1 some age is missing
#
# Como algunas edades no figuran, hay que hacer data imputation
# + colab={"base_uri": "https://localhost:8080/", "height": 422} id="ChiuxD31AbEj" outputId="c933fa1c-a2a8-4b68-aae5-409a7451e9a7"
train.head(100)
# + id="Bu2ZI66LAbEj"
# Esta es la línea que se utiliza para hacer la imputación de la edad
train["Age"].fillna(train.groupby("Title")["Age"].transform("median"), inplace=True)
# + colab={"base_uri": "https://localhost:8080/"} id="vLVGR5h3SBIm" outputId="bf05151a-104f-429c-eba0-b77fbbf972d1"
train.isnull().sum() # Age now is without NaN
# + [markdown] id="evq1lQJdAbEj"
# # 6) La imputación de la edad se hace según:
# La media de la columna.
# La media de la columna condicionada por el título de la persona.
# La mediana de la columna.
# La mediana de la columna condicionada por el título de la persona.``True.``
# + colab={"base_uri": "https://localhost:8080/"} id="DPnVQ5FJAbEj" outputId="8cb253bc-6438-4194-f2ec-8b3966a9778f"
train.head()
train.groupby("Title")["Age"].transform("median")
# + colab={"base_uri": "https://localhost:8080/", "height": 221} id="M-2DTz9PAbEl" outputId="85c2aa03-f476-452c-c763-68d28295a4f2"
facet = sns.FacetGrid(train, hue="Survived",aspect=4)
facet.map(sns.kdeplot,'Age',shade= True)
facet.set(xlim=(0, train['Age'].max()))
facet.add_legend()
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 239} id="cvqH6lfbAbEl" outputId="4f7802a6-d5df-497d-946d-26c4813770b4"
facet = sns.FacetGrid(train, hue="Survived",aspect=4)
facet.map(sns.kdeplot,'Age',shade= True)
facet.set(xlim=(0, train['Age'].max()))
facet.add_legend()
plt.xlim(0, 20)
# + colab={"base_uri": "https://localhost:8080/", "height": 239} id="AOCfZ0JoAbEl" outputId="144a16c4-b31c-435f-ffea-d6dedd2b246d"
facet = sns.FacetGrid(train, hue="Survived",aspect=4)
facet.map(sns.kdeplot,'Age',shade= True)
facet.set(xlim=(0, train['Age'].max()))
facet.add_legend()
plt.xlim(20, 30)
# + colab={"base_uri": "https://localhost:8080/", "height": 239} id="iaKLfLnUAbEl" outputId="ef316a28-ca08-4569-c701-d6dcba905f54"
facet = sns.FacetGrid(train, hue="Survived",aspect=4)
facet.map(sns.kdeplot,'Age',shade= True)
facet.set(xlim=(0, train['Age'].max()))
facet.add_legend()
plt.xlim(30, 40)
# + colab={"base_uri": "https://localhost:8080/", "height": 239} id="IEiRnf5IAbEl" outputId="ef4d994b-82fb-453f-cc3b-b109842ebc00"
facet = sns.FacetGrid(train, hue="Survived",aspect=4)
facet.map(sns.kdeplot,'Age',shade= True)
facet.set(xlim=(0, train['Age'].max()))
facet.add_legend()
plt.xlim(40, 60)
# + colab={"base_uri": "https://localhost:8080/", "height": 239} id="rcrWjLFDAbEl" outputId="a9e91646-55f9-48f8-dbf0-b4d0907137fa"
facet = sns.FacetGrid(train, hue="Survived",aspect=4)
facet.map(sns.kdeplot,'Age',shade= True)
facet.set(xlim=(0, train['Age'].max()))
facet.add_legend()
plt.xlim(40, 60)
# + colab={"base_uri": "https://localhost:8080/", "height": 239} id="dbfygOoRAbEl" outputId="7a949ece-25a3-4104-ca85-feb0ffc59e56"
facet = sns.FacetGrid(train, hue="Survived",aspect=4)
facet.map(sns.kdeplot,'Age',shade= True)
facet.set(xlim=(0, train['Age'].max()))
facet.add_legend()
plt.xlim(60)
# + colab={"base_uri": "https://localhost:8080/"} id="OpZWbSWjAbEm" outputId="0138d3f6-4185-4331-abd7-f36793632fea"
train.info()
# + [markdown] id="RXDQ3tQAAbEm"
# #### 4.4.2 Binning
#
#
# Se convierte la edad numérica en variables categóricas.
#
# feature vector map:
# child: 0
# young: 1
# adult: 2
# mid-age: 3
# senior: 4
# +
# We divide Age in 4 parts and change it from categorical variable to a numeric form
# for dataset in train_test_data:
# dataset['Age'] = dataset['Age'].astype(int)
# train['AgeBand'] = pd.cut(train['Age'], 5)
# print (train[['AgeBand', 'Survived']].groupby(['AgeBand'], as_index=False).mean())
# +
for dataset in train_test_data:
dataset['Age'] = dataset['Age'].astype(int)
bins = pd.IntervalIndex.from_tuples([(0, 16), (17, 26), (27, 36), (37, 62), (63, 99)])
train['AgeBand'] = pd.cut(train['Age'], bins)
print(train[['AgeBand', 'Survived']].groupby(['AgeBand'], as_index=False).mean())
# -
for dataset in train_test_data:
dataset.loc[ dataset['Age'] <= 16, 'Age'] = 0
dataset.loc[(dataset['Age'] > 16) & (dataset['Age'] <= 26), 'Age'] = 1
dataset.loc[(dataset['Age'] > 26) & (dataset['Age'] <= 36), 'Age'] = 2
dataset.loc[(dataset['Age'] > 36) & (dataset['Age'] <= 62), 'Age'] = 3
dataset.loc[ dataset['Age'] > 62, 'Age'] = 4
dataset['Age'] = dataset['Age'].astype(int)
# + id="27Bz3pfwAbEm"
train.head()
# + id="PLiM472pAbEm"
bar_chart('Age')
# + [markdown] id="8vCnyNp4AbEm"
# ### 4.5 Embarked
# + [markdown] id="_BJRdHn_AbEm"
# #### 4.5.1 filling missing values
# + id="NHnJpSbrAbEm"
Pclass1 = train[train['Pclass']==1]['Embarked'].value_counts()
Pclass2 = train[train['Pclass']==2]['Embarked'].value_counts()
Pclass3 = train[train['Pclass']==3]['Embarked'].value_counts()
df = pd.DataFrame([Pclass1, Pclass2, Pclass3])
df.index = ['1st class','2nd class', '3rd class']
df.plot(kind='bar',stacked=True, figsize=(10,5))
# + [markdown] id="kiSiPLMMAbEn"
# # 7) Con qué criterio rellenaría el puerto de embarque?
#
# S para todas las clases ``True``
# C para primera clase, S para el resto
# Q para tercera clase, S para el resto
# + id="FOwPvRhCAbEn"
for dataset in train_test_data:
dataset['Embarked'] = dataset['Embarked'].fillna('S')
# + id="RANxqt39AbEn"
train.head()
# + id="Y0CTG5TnAbEn"
embarked_mapping = {"S": 0, "C": 1, "Q": 2}
for dataset in train_test_data:
dataset['Embarked'] = dataset['Embarked'].map(embarked_mapping)
# + [markdown] id="KKCV0WBQAbEn"
# ### 4.6 Fare
#
# Rellenar la tarifa con la mediana de la tarifa condicionada por cada clase
#
# + id="WDAALXL_AbEn"
# fill missing Fare with median fare for each Pclass
# Escribir el código acá
train["Fare"].fillna(train.groupby("Pclass")["Fare"].transform("median"), inplace=True)
train.head(50)
# + id="g9CWwly6AbEn"
facet = sns.FacetGrid(train, hue="Survived",aspect=4)
facet.map(sns.kdeplot,'Fare',shade= True)
facet.set(xlim=(0, train['Fare'].max()))
facet.add_legend()
plt.show()
# + id="V2a-f5PIAbEo"
facet = sns.FacetGrid(train, hue="Survived",aspect=4)
facet.map(sns.kdeplot,'Fare',shade= True)
facet.set(xlim=(0, train['Fare'].max()))
facet.add_legend()
plt.xlim(0, 20)
# + id="n3zOkF8AAbEo"
facet = sns.FacetGrid(train, hue="Survived",aspect=4)
facet.map(sns.kdeplot,'Fare',shade= True)
facet.set(xlim=(0, train['Fare'].max()))
facet.add_legend()
plt.xlim(0, 30)
# + id="VuvXzCIkAbEo"
facet = sns.FacetGrid(train, hue="Survived",aspect=4)
facet.map(sns.kdeplot,'Fare',shade= True)
facet.set(xlim=(0, train['Fare'].max()))
facet.add_legend()
plt.xlim(0)
# +
# We divide Fare in 4 parts and change it from categorical variable to a numeric form
# +
for dataset in train_test_data:
dataset['Fare'] = dataset['Fare'].astype(int)
bins = pd.IntervalIndex.from_tuples([(0, 17), (18, 30), (31, 100), (101, 513)])
train['FareBand'] = pd.cut(train['Fare'], bins)
print(train[['FareBand', 'Survived']].groupby(['FareBand'], as_index=False).mean())
for dataset in train_test_data:
dataset.loc[ dataset['Fare'] <= 17, 'Fare'] = 0
dataset.loc[(dataset['Fare'] > 17) & (dataset['Fare'] <= 30), 'Fare'] = 1
dataset.loc[(dataset['Fare'] > 30) & (dataset['Fare'] <= 100), 'Fare'] = 2
dataset.loc[ dataset['Fare'] > 100, 'Fare'] = 3
dataset['Fare'] = dataset['Fare'].astype(int)
# + id="Gr16z4cPAbEo"
train.head()
# + [markdown] id="O2dn3SbGAbEo"
# ### 4.7 Cabin
# + id="BJTkExiAAbEp"
train.Cabin.value_counts()
# + id="wbptNfQLAbEp"
for dataset in train_test_data:
dataset['Cabin'] = dataset['Cabin'].str[:1]
# + id="XRAViBjUAbEp"
Pclass1 = train[train['Pclass']==1]['Cabin'].value_counts()
Pclass2 = train[train['Pclass']==2]['Cabin'].value_counts()
Pclass3 = train[train['Pclass']==3]['Cabin'].value_counts()
df = pd.DataFrame([Pclass1, Pclass2, Pclass3])
df.index = ['1st class','2nd class', '3rd class']
df.plot(kind='bar',stacked=True, figsize=(10,5))
# + id="fjSiehIHAbEp"
cabin_mapping = {"A": 0, "B": 0.4, "C": 0.8, "D": 1.2, "E": 1.6, "F": 2, "G": 2.4, "T": 2.8}
for dataset in train_test_data:
dataset['Cabin'] = dataset['Cabin'].map(cabin_mapping)
# + id="Imytn2sfAbEp"
# fill missing Cabin with median Cabin for each Pclass
train["Cabin"].fillna(train.groupby("Pclass")["Cabin"].transform("median"), inplace=True)
# + [markdown] id="q-Ogs9DhAbEp"
# ### 4.8 FamilySize
# + id="N4HjcUYhAbEp"
train["FamilySize"] = train["SibSp"] + train["Parch"] + 1
# + id="QqAtWDJzAbEq"
facet = sns.FacetGrid(train, hue="Survived",aspect=4)
facet.map(sns.kdeplot,'FamilySize',shade= True)
facet.set(xlim=(0, train['FamilySize'].max()))
facet.add_legend()
plt.xlim(0)
# + id="4XgXxcQoAbEq"
family_mapping = {1: 0, 2: 0.4, 3: 0.8, 4: 1.2, 5: 1.6, 6: 2, 7: 2.4, 8: 2.8, 9: 3.2, 10: 3.6, 11: 4}
for dataset in train_test_data:
dataset['FamilySize'] = dataset['FamilySize'].map(family_mapping)
# + id="NIFhgCBDAbEq"
train.head()
# + id="Upj0DRPBAbEq"
train.head()
# + id="02wQV5G8AbEq"
#Now we drop useless features
features_drop = ['Ticket', 'SibSp', 'Parch', 'AgeBand', 'FareBand']
train = train.drop(features_drop, axis=1)
train = train.drop(['PassengerId'], axis=1)
# + id="-n7UKwZLAbEq"
train_data = train.drop('Survived', axis=1)
target = train['Survived']
train_data.shape, target.shape
# + id="-DAqyw05AbEq"
train_data.head(10)
# + [markdown] id="fGJ8S0RUAbEq"
# ## 5. Modelling
# + id="ybixBlSNAbEs"
# Importing Classifier Modules
from sklearn.neighbors import KNeighborsClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.naive_bayes import GaussianNB
from sklearn.svm import SVC
import numpy as np
# + id="mSoF61QwAbEs"
train.info()
# + [markdown] id="8k2lkGlYAbEs"
# ### 6.2 Cross Validation (K-fold)
# + id="CoAUvjFkAbEs"
from sklearn.model_selection import KFold
from sklearn.model_selection import cross_val_score
k_fold = KFold(n_splits=5, shuffle=True, random_state=0)
# + [markdown] id="-ihBx4anAbEs"
# ### 6.2.1 kNN
# + id="Nal7TUZ9AbEs"
clf = KNeighborsClassifier(n_neighbors = 13)
scoring = 'accuracy'
score = cross_val_score(clf, train_data, target, cv=k_fold, n_jobs=1, scoring=scoring)
print(score)
# + id="_K8NGvnsAbEt"
# kNN Score
round(np.mean(score)*100, 2)
# + [markdown] id="sCmt2PF9AbEt"
# ### 6.2.2 Decision Tree
# + id="1k-Z5w3fAbEt"
clf = DecisionTreeClassifier()
scoring = 'accuracy'
score = cross_val_score(clf, train_data, target, cv=k_fold, n_jobs=1, scoring=scoring)
print(score)
# + id="sJqMemMEAbEt"
# decision tree Score
round(np.mean(score)*100, 2)
# + [markdown] id="ghnU18kCAbEt"
# ### 6.2.3 Ramdom Forest
# + id="tV6SGGEGAbEt"
clf = RandomForestClassifier(n_estimators=13)
scoring = 'accuracy'
score = cross_val_score(clf, train_data, target, cv=k_fold, n_jobs=1, scoring=scoring)
print(score)
# + id="LcI0qQN8AbEu"
# Random Forest Score
round(np.mean(score)*100, 2)
# -
# ### 6.2.4 ExtraTree
from sklearn.ensemble import ExtraTreesClassifier
clf = ExtraTreesClassifier()
scoring = 'accuracy'
score = cross_val_score(clf, train_data, target, cv=k_fold, n_jobs=1, scoring=scoring)
print(score)
# Extra Tree Score
round(np.mean(score)*100, 2)
# + [markdown] id="-dyRapxMAbEu"
# ### 6.2.5 Naive Bayes
# + id="aPNe16cHAbEu"
clf = GaussianNB()
scoring = 'accuracy'
score = cross_val_score(clf, train_data, target, cv=k_fold, n_jobs=1, scoring=scoring)
print(score)
# + id="5-iRrPOdAbEv"
# Naive Bayes Score
round(np.mean(score)*100, 2)
# + [markdown] id="i-2BgmneAbEv"
# # 8)
# Realizar una búsqueda de hiperparámetros para un modelo con Decision Tree, Random Forest y Extra Tree.
# Informar el error de validación cruzada usando k-folding con k=5 para cada uno de los mejores modelos.
#
# Decision Tree: ``80.14``
#
# Random Forest: ``80.7``
#
# ExtRa Tree: ``80.25``
# + [markdown] id="NSNBogBCAbEv"
# ## References
#
# This notebook is created by learning from the following notebooks:
#
# - [M<NAME>againTitanic Solution: A Beginner's Guide](https://www.kaggle.com/chapagain/titanic-solution-a-beginner-s-guide?scriptVersionId=1473689)
# - [How to score 0.8134 in Titanic Kaggle Challenge](http://ahmedbesbes.com/how-to-score-08134-in-titanic-kaggle-challenge.html)
# - [Titanic: factors to survive](https://olegleyz.github.io/titanic_factors.html)
# - [Titanic Survivors Dataset and Data Wrangling](http://www.codeastar.com/data-wrangling/)
#
|
ML/Laboratorio5/Laboratorio_5.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Descriptors
# As we discussed earlier, Python instance properties (using `@property` for example) are based on Python descriptors which are simply classes that implement the **descriptor protocol**.
#
# The protocol is comprised of the following special methods - not all are required.
# - `__get__`: used to retrieve the property value
# - `__set__`: used to store the property value (we'll see where we can do this in a bit)
# - `__del__`: delete a property from the instance
# - `__set_name__`: new to Python 3.6, we can use this to capture the property name as it is being defined in the owner class (the class where the property is defined).
#
# There are two types of descriptors we need to distingush as I explain in the video lecture:
# - non-data descriptors: these are descriptors that only implement `__get__` (and optionally `__set_name__`)
# - data descriptors: these implement the `__set__` method, and normally, also the `__get__` method.
#
# As we'll see in a bit, functions in Python actually implement the (non-data) descriptor protocol as well - indeed that is how instance methods work!
# Let's start with a quick example first:
# +
from datetime import datetime
class TimeUTC:
def __get__(self, instance, owner_class):
return datetime.utcnow().isoformat()
# -
# So `TimeUTC` is a **non-data descriptor** since it only implements `__get__`.
class Logger:
current_time = TimeUTC()
l = Logger()
l.current_time
# As we discussed in the lecture, the `__get__` method will know what instance (if any) was used to call it, as well as the class that owns the instance of `TimeUTC` (the descriptor instance).
#
# This information is passed to `__get__` when it gets called:
class TimeUTC:
def __get__(self, instance, owner_class):
print('__get__ called', instance, owner_class)
return datetime.utcnow().isoformat()
class Logger:
current_time = TimeUTC()
# When accessing the `current_time` attribute from the class:
Logger.current_time
# and as you can see, `instance` was `None`, while `owner_class` is the `Logger` class defined in our global scope (not an instance of `Logger`, but the class itself)
# Now let's create an instance of the `Logger` class:
l = Logger()
hex(id(l))
# and call `current_time` from the instance:
l.current_time
# and as you can see here, `instance` was the instance `l`, and `owner_class` is still the same `Logger` class.
# Often we choose to return the descriptor instance when called from the class, like so:
class TimeUTC:
def __get__(self, instance, owner_class):
if not instance:
return self
return datetime.utcnow().isoformat()
class Logger:
current_time = TimeUTC()
# So now we have:
Logger.current_time
# the instance of the `TimeUTC` class, but when called from an instance:
l = Logger()
l.current_time
# we get the time string returned instead.
# Looking at data descriptors now we're going to implement a `__set__` method
# We'll need to decide where to store the attribute value - naively we'll store it directly in the descriptor itself (spoiler alert - that won't work the way we probably want it to work):
class IntegerValue:
def __set__(self, instance, value):
self.value = int(value)
def __get__(self, instance, owner_class):
if not instance:
return self
return self.value
class Point2D:
x = IntegerValue()
y = IntegerValue()
p1 = Point2D()
p1.x = 100.1
p1.x
# Ok, that works...
p1.y = 200.2
p1.y
# That seems to work too...
# But now let's create a second point:
p2 = Point2D()
p2.x = 1.1
p2.x
p1.x
# So, although we were aiming to modify the `x` value on `p2` we ended up modifying it on `p1` as well - this is because both `p1` and `p2` share the same **class level** instance of `IntegerValue`.
# As you can see, when we "store" data we need to be mindful of the **instance** we are storing the data for - otherwise if we just store the data in the descriptor instance, insce all instances of our class (`Point2D`) share the same instance of the descriptor, we are essentially working with a **class** level property, not an instance property (which is how the `@property` descriptor works - it creates **instance** properties).
#
# Since we know the instance we are dealing with in both the `__get__` and `__set__` methods, we could easily use the instance dictionary to store the attribute value:
class IntegerValue:
def __set__(self, instance, value):
instance.value = int(value)
def __get__(self, instance, owner_class):
if not instance:
return self
return instance.value
class Point2D:
x = IntegerValue()
y = IntegerValue()
p1 = Point2D()
p2 = Point2D()
p1.x = 1.1
p2.x = 10.1
p1.x, p2.x
# And let's see what's in our instance dictionaries:
p1.__dict__
# As you can see we used `value` to store the value for `x` in that instance.
#
# Where's the value for `y` going to get stored?
#
# Yeah, in `value` as well!
p1 = Point2D()
p1.x = 10.1
p1.__dict__
p1.y = 20.2
p1.__dict__
p1.y
# that looks good, but:
p1.x
# That's not so good - we overwrote the value for `x`.
#
# Ok, no big deal, we just have to give a different name to the dictionary key...
#
# How do we modify this code so that every instance of `IntegerValue` uses a different symbol for storage??
class IntegerValue:
def __set__(self, instance, value):
instance.value = int(value)
def __get__(self, instance, owner_class):
if not instance:
return self
return instance.value
# Even if we figure out a way to do this, or if we only used a single `IntegerValue` in our class:
class Point1D:
x = IntegerValue()
p1 = Point1D()
p1.x = 100.1
p1.x
p2 = Point1D()
p2.x = 200.2
p2.x
p1.x
# That works, but what if we had defined our `Point1D` class using slots?
class Point1D:
__slots__ = 'origin',
x = IntegerValue()
def __init__(self, origin):
self.origin = origin
p1 = Point1D(0)
p1.x = 100.1
# Right, we cannot assign to `value` because it is not defined in `__slots__` and we don't have a `__dict__` available.
#
# We could solve this in one of two ways: add `value` to slots, or add `__dict__` to slots:
class Point1D:
__slots__ = 'origin', 'value'
x = IntegerValue()
def __init__(self, origin):
self.origin = origin
p = Point1D(0)
p.x = 100.1
p.x
# + active=""
# or, using `__dict__` instead:
# -
class Point1D:
__slots__ = 'origin', '__dict__'
x = IntegerValue()
def __init__(self, origin):
self.origin = origin
p1 = Point1D(0)
p1.x = 100.1
p1.x
# But this is really not a very user-friendly thing to do to users of our data descriptor!
#
# There are some other ways of doing this, where we do not use the instances to store the data but instead store the values somewhere else (and retrieve them from that same place).
# We could try to use a dictionary for that - assuming our instances are hashable:
class IntegerValue:
def __init__(self):
self.data = {}
def __set__(self, instance, value):
self.data[instance] = int(value)
def __get__(self, instance, owner_class):
if not instance:
return self
return self.data.get(instance)
class Point2D:
x = IntegerValue()
y = IntegerValue()
p1 = Point2D()
p2 = Point2D()
p1.x = 10.1
p1.y = 20.2
p2.x = 100.1
p2.y = 200.2
p1.x, p1.y
p2.x, p2.y
# But we have a potential for memory leaks!
#
# Let's look at this closer.
p = Point2D()
p.x = 12345
# So we have a number of objects at play here:
#
# - the `p` instance
# - the instance of `IntegerValue` we are using for the `x` property
p_id = id(p)
x_attrib_id = id(Point2D.x)
hex(p_id), hex(x_attrib_id)
# Now, let's delete `p` - this should remove all the references to that object, so it should get garbage collected (through reference counting)
del p
# `p` is no longer in our global namespace:
'p' in globals()
# But what about the data descriptor instance? It **still** has a reference to that object as a **key** in it's dictionary!
x_data_descriptor = Point2D.x
x_data_descriptor.data
# As you can see we have an entry for our "deleted" point still in that dictionary! The object still exists, and we have a memory leak.
# Let's make sure the object actually still exists:
point = list(x_data_descriptor.data.keys())[-1]
point.x
# and in fact, the `id` will match our original `id`:
id(point), p_id
# ### Weak References
# We know that with reference counting, objects are garbage collected only when all references to them are gone. This is the problem we just saw.
#
# What we need is a way to hold a reference to an object without "affecting" the reference count - or at least letting Python know that although we have a referfence to an object, we don't want our reference to "count" in it's reference counting.
#
# This is called a **weak reference** - as opposed to a normal or **strong** reference.
class Person:
def __init__(self, name):
self.name = name
p = Person('Guido')
# Now we can read and write the `name` property of `p`:
p.name
p.name = 'Alex'
p.name
# We can set up a second (strong) reference to our object `p`:
p2 = p
id(p2), id(p)
# And now, even though we delete `p`:
del p
'p' in globals()
# The object still exists, as it is referenced by `p2`:
p2.name
# Let's try creating a weak reference instead:
# +
import weakref
p = Person('Raymond')
p2 = weakref.ref(p)
# -
hex(id(p)), hex(id(p2))
# Ah, not the same id!
#
# In fact, that weakref does have a (weak) reference to `p`, but it is pointing (indirectly) to the same object:
p2
# We can get the original object back by calling the value returned from `ref`:
hex(id(p2()))
p2().name
# So now we have two references to the `Person` instance.
#
# What's the reference count?
# +
import ctypes
ctypes.c_long.from_address(id(p)).value
# -
# We can get the number of weak references, using the `getweakrefcount` from the `weakref` module:
weakref.getweakrefcount(p)
# Now let's delete our original strong reference (`p`) - that should leave us with only a single weak reference:
del p
'p' in globals()
# What happened to our weak reference?
p2, p2()
# As you can see, the original object is `dead` - concise terminology I guess :-), and calling `p2()` returns `None`.
# Using the strong reference count is not accurate since we don't know what's stored at that memory address anymore. But the weak reference tells us that the original object is now gone (note that `dead` really means the object is no longer usable - but because of the non-deterministic nature of the garbage collector, the actual may actually hang around for a while until it is actually destroyed. From our viewpoint though, the object is gone, and the memory it was holding on to, will, eventually, be released.
# This can be really useful for avoiding memory leaks in our data descriptors - instead of using a dictionary that contains strong references to our instances as the keys, we can use weak references instead - that way if the original object goes away (the instance that contains the property value), we can be assured that the object will be garbage collected - so no memory leak.
# Not every object in Python supports wesk references. Dictionaries do not for example, neither do tuple or ints. But custom classes do, and that's really all we're interested in here. Just be aware of that, and read up the Python docs on `weakref` if you want more info.
# There is a lot of functionality in the `weakref` module, more than we really need for this course, but there is one more I want to discuss: the `WeakKeyDictionary`.
#
# It works like a standard dictionary, but all the keys are stored as weak references to the key object, instead of strong references we would have with a standard dictionary.
p1 = Person('Guido')
p2 = Person('Raymond')
p3 = Person('Mark')
p4 = Person('Alex')
# +
d = weakref.WeakKeyDictionary()
d[p1] = 'Guido'
d[p2] = 'Raymond'
d[p3] = 'Mark'
d[p4] = 'Alex'
id_p1 = id(p1)
print(hex(id_p1))
# -
ctypes.c_long.from_address(id_p1).value
weakref.getweakrefcount(p1)
list(d.keyrefs())
hex(id(p1))
# Now, what happens if we destroy `p1` for example, by remoiving the only strong reference we have to it:
del p1
list(d.keyrefs())
# As you can see, `p1` has been removed from the `WeakKeyDictionary` automatically. That's handy too.
# We can access elements in the `WeakKeyDictionary` using the original objects:
d[p2]
# In summary, it is very difficult to do low-level operations like reference counting and so on in Python - it was not really built to expose these inner workings to us, the Python developer. Just understand the difference between weak and strong references, and trust Python to do its memory management correctly - of course you need to be aware of the memory leak traps like the one we encountered with data descriptors.
# ### Data Descriptors and Weak References
# Now that we understand weak references and WeakKeyDictionaries, let's go back to our data descriptor example, and use weak references to insure no memory leaks.
# +
from weakref import WeakKeyDictionary
class IntegerValue:
def __init__(self):
self.data = WeakKeyDictionary()
def __set__(self, instance, value):
self.data[instance] = int(value)
def __get__(self, instance, owner_class):
if not instance:
return self
return self.data.get(instance)
# -
class Point2D:
x = IntegerValue()
y = IntegerValue()
# +
p1 = Point2D()
p2 = Point2D()
p1.x = 10.1
p1.y = 20.2
p2.x = 100.1
p2.y = 200.2
# -
p1.x, p1.y
p2.x, p2.y
list(Point2D.x.data.keyrefs())
list(Point2D.y.data.keyrefs())
# And now, if we delete one of our points:
del p1
list(Point2D.x.data.keyrefs())
list(Point2D.y.data.keyrefs())
# As you can observe our weak dict no longer has weak references to the point `p1` - because the object `p1` was referencing was garbage collected, and this was picked up by the weak dict.
# So, this technique will work well, but there is still one slight issue.
#
# Our `Point2D` class was hashable - but what happens if it is not?
class Point2D:
x = IntegerValue()
y = IntegerValue()
def __eq__(self, other):
if isinstance(other, Point2D):
return self.x == other.x and self.y == other.y
return False
p1 = Point2D()
p1.x = 10.1
# The problem is that we are trying to make a non-hashable object a key in a dictionary - and that obviously cannot work, not even with weak key dictionaries!
# So, we'll need some kind of work-around.
#
# Let's consider what we need to do to keep the attribute values for each point instances: we need a way to look up a value for a specific object (instance).
#
# But two things:
# 1. we need to keep weak references to the object only
# 2. we cannot use the object directly as a key in a dictionary
#
# How about just using the `id` of the object? We don't even need a weak reference if we do that...
class IntegerValue:
def __init__(self):
self.data = {}
def __set__(self, instance, value):
self.data[id(instance)] = int(value)
def __get__(self, instance, owner_class):
if not instance:
return self
return self.data.get(id(instance))
class Point2D:
x = IntegerValue()
y = IntegerValue()
# +
p1 = Point2D()
p1.x = 10.1
p1.y = 20.2
p2 = Point2D()
p2.x = 100.1
p2.y = 200.2
# -
p1.x, p1.y, p2.x, p2.y
# So that seems to work just fine. But let's look at the dictionaries used to store the values in the data descriptors before and after we finalize the objects:
Point2D.x.data, Point2D.y.data
del p1
del p2
Point2D.x.data, Point2D.y.data
# As you can see the dictionary does not get cleaned up.
# That's probably not going to be a problem in practice, but there is always a chance of a new object getting the same id as an old one still present in the dictionary. Although it won't affect the setter (it will just replace the old value), we could end up with a value for a getter for a property that has not been set yet.
#
# Although unlikely, it would not be good practice to allow for that possibility.
#
# Somehow we need to clean up the dictionary when the objects are finalized.
#
# There are a number of ways we could do this, but remember how `weakref.ref` returns a callable that itself returns `None` if the object it was pointing to has been finalized?
#
# We can use that to our advantage - instead of just storing the value in our dictionary, we are going to store both the weakref and the value. Whenever a value is requested, we'll first make sure the weakref is not `None`.
class IntegerValue:
def __init__(self):
self.data = {}
def __set__(self, instance, value):
key = id(instance)
weak_ref = weakref.ref(instance)
self.data[key] = weak_ref, int(value)
def __get__(self, instance, owner_class):
if not instance:
return self
key = id(instance)
weak_ref, value = self.data.get(key, (None, None))
if value is not None and weak_ref is not None:
# key present in dictionary
# but object has ben garbage collected
# remove it from the dictionary
del self.data[key]
return self.data.get(key)
class Point2D:
x = IntegerValue()
y = IntegerValue()
p1 = Point2D()
p1.x = 10.1
Point2D.x.data
del p1
Point2D.x.data
# So, if we ever get another point object with the same memory address we should be fine.
# However, we are still leaving "dead" entries behind in the dictionary.
#
# This lazy deletion could be a problem in certain circumstances.
#
# We could do this another way yet, using a feature the `weakref.ref` class has: a callback feature for when the weakly referenced object is being finalized.
#
# Let's see how that callback works first:
p = Point2D()
# Create the callback function:
def finalize_callback(weak_ref):
print('object being finalized...')
print(hex(id(weak_ref)), weak_ref())
# Create the weak reference, specifying the callback function:
p_weak = weakref.ref(p, finalize_callback)
del p
# Nice!! As you can see our callback unfction was called once the object `p` was finalized. So let's use that technique instead:
class IntegerValue:
def __init__(self):
self.data = {}
def __set__(self, instance, value):
key = id(instance)
weak_ref = weakref.ref(instance, self.instance_finalizer)
self.data[key] = weak_ref, int(value)
def __get__(self, instance, owner_class):
if not instance:
return self
return self.data.get(id(instance))
def instance_finalizer(self, weak_ref):
# now we need to find the key corresponding to that weak_ref
# unfortunately we do not have the object being finalized
# so we have to do a reverse lookup from the weak refs
# stored in the data dictionary
reverse_lookup = [key for key, value in self.data.items()
if value[0] is weak_ref]
if reverse_lookup:
# key found
print('Cleaning up weak ref entry!')
key = reverse_lookup[0]
del self.data[key]
class Point2D:
x = IntegerValue()
y = IntegerValue()
p = Point2D()
p.x = 10.1
del p
Point2D.x.data
# So now, (finally!) we have a way to use data descriptors for instance properties that handles objects with no `__dict__` and potentially unhashable too.
# ### `__set_name__` Descriptor Method
# Let's look at thet `__set_name__` method in more detail now. New to Python 3.6, it is very handy.
# Let's first see how it works:
class Descriptor:
def __set_name__(self, owner_class, name):
print(f'receiving property name: {name}')
self.name = name
def __set__(self, instance, value):
print(f'set called for property {self.name}')
def __get__(self, instance, owner_class):
print(f'get called for property {self.name}')
class MyClass:
my_attrib = Descriptor()
# See how the `__set_name__` method was called immediately after the descriptor was created?
# This method is called only once, when the descriptor is being instantiated, and we can now store the property name (as used in the owner class) directly in the descriptor instance.
obj = MyClass()
obj.my_attrib
obj.my_attrib = 100
# ### Property Value Lookup Resolution
# Remember that I mentioned earlier that distinguishing between data and non-data descriptors was important?
#
# Let's look into that now.
class DataDescriptor:
def __get__(self, instance, owner_class):
print('using __get__')
def __set__(self, instance, value):
print('using __set__')
class MyClass:
prop = DataDescriptor()
m = MyClass()
m.prop
m.prop = 100
m.__dict__
m.__dict__['prop'] = 'Some value'
m.__dict__
m.prop
m.prop = 100
# So, with **data** descriptors, the `__get__` and `__set__` methods are always called (by default anyways - that can be overrideen if desired)
# Now let's see how it works with **non-data** descriptors:
class NonDataDescriptor:
def __get__(self, instance, owner_class):
print('using __get__')
class MyClass:
prop = NonDataDescriptor()
m = MyClass()
m.prop
m.__dict__['prop'] = 100
m.__dict__
m.prop
# Aha, that returns the value stored in the instance dictionary - did not call the `__get__` method!
# What about setting via dotted notation:
m.prop = 200
m.__dict__
# Ok, so that went straight to the instance dictionary too - but that should be expected, our descriptor class does not implement a `__set__` method.
# How about getting the property using the `getattr` function?
getattr(m, 'prop')
# That also uses the dictionary.
# One last variation, what happens if we have a non-data descriptor and we try to set the same property name using dotted notation instead of using `__dict__`:
m = MyClass()
m.__dict__
m.prop = 100
m.__dict__
# And now of course, when we try to retrieve `prop` the instance dictionary will take precendence:
m.prop
# So the basics here is:
#
# ##### Data Descriptors
# - always uses the descriptor instance `__set__` and `__get__` methods
#
# ##### Non-Data Descriptors
# - looks in the instance dictionary first
# - falls back to the descriptor instance
# ### Validation using Descriptors
# So now here is a typical example of how we can leverage data descriptors in a re-useable fashion:
# +
from numbers import Integral
class IntegerRange:
def __init__(self, min_value, max_value):
self.min_value = min_value
self.max_value = max_value
def __set_name__(self, owner_class, name):
self.name = name
def __get__(self, instance, owner_class):
return instance.__dict__[self.name]
def __set__(self, instance, value):
if not isinstance(value, Integral):
raise ValueError(f'{self.name}: must be an integer.')
value = int(value)
if self.min_value <= value <= self.max_value:
instance.__dict__[self.name] = value
else:
raise ValueError(f'{self.name}: must be between '
f'{self.min_value} and {self.max_value}.')
# -
class Person:
age = IntegerRange(0, 150)
p = Person()
p.age = "10"
p.age = 200
p.age = 18
p.age
p.age = -10
# ### Exercise: property and Descriptors
# Now that we understand descriptors, let's go back to that `property` and the related decorators we used for quick and simple property definitions in our classes:
# +
from numbers import Integral
class Person:
@property
def age(self):
return self._age
@age.setter
def age(self, value):
if not isinstance(value, Integral):
raise ValueError('age: must be an integer')
value = int(value)
if value < 0:
raise ValueError('age: must be a non-negative integer.')
self._age = value
# -
p = Person()
p.age = -10
p.age = 10
p.age
# What the property (and the decorators) do is essentially create all the (in this case data) descriptor for us.
# Recall how the non-decorator version worked:
class Person:
def get_age(self):
return self._age
def set_age(self, value):
if not isinstance(value, Integral):
raise ValueError('age: must be an integer')
value = int(value)
if value < 0:
raise ValueError('age: must be a non-negative integer.')
self._age = value
age = property(fget=get_age, fset=set_age)
p = Person()
p.age = 10
p.age
p.age = -10
# Let's see how we might create our own version of `property` - it's a good exercise and will help solidy our understanding of data descriptors. For simplicity we'll omit support for `__del__` - but iot works the same way.
# First let's deal with the non-decorator version of `property`:
class MakeProperty:
def __init__(self, fget=None, fset=None):
self.fget = fget
self.fset = fset
def __set_name__(self, owner_class, name):
self.name = name
def __get__(self, instance, owner_class):
if not instance:
return self
if not self.fget:
raise AttributeError(f'{self.name} is not readable.')
return self.fget(instance)
def __set__(self, instance, value):
if not self.fset:
raise AttributeError(f'{self.name} is not writable.')
self.fset(instance, value)
# This is enough to make a generic property creator:
class Person:
def get_name(self):
return self._name
def set_name(self, value):
self._name = value
name = MakeProperty(fget=get_name, fset=set_name)
p = Person()
p.name = 'Alex'
p.name
# Now, let's handle the decorators.
#
# Remember how the `property` decorator works - the `@property` decorator is used to specify the getter, and whatever returns from that decorator must have a `setter` attribute we can use to decorate the setter method.
# + active=""
# First thing we are going to do is add a few methods to the `MakeProperty` descriptor:
# -
class MakeProperty:
def __init__(self, fget=None, fset=None):
self.fget = fget
self.fset = fset
def __get__(self, instance, owner_class):
if not instance:
return self
if not self.fget:
raise AttributeError('attribute is not readable.')
return self.fget(instance)
def __set__(self, instance, value):
if not self.fset:
raise AttributeError('attribute is not writable.')
self.fset(instance, value)
def getter(self, fget):
self.fset = fset
return self
def setter(self, fset):
self.fset = fset
return self
class Person:
@MakeProperty
def name(self):
return self._name
@name.setter
def name(self, value):
self._name = value
p = Person()
p.name = 'Alex'
p.name
# Of course our implementation is simplistic and omits things, like the `__doc__` attribute iof the original function for example, for example, or the `__delete__`, but this is the basic idea.
# ### Functions are Non-Data Descriptors
# Functions as we know are objects. In fact, they implement the non-data descriptor protocol!
#
# Yes, they have a `__get__` method. That's how functions defined in a class, actually become instance methods when called from class instances.
# Let's see this:
class Person:
def __init__(self, name):
self.name = name
def say_hello(self):
return f'{self.name} says hello'
say_hello_func = Person.__dict__['say_hello']
say_hello_func
# So, as we can see, the `Person` class contains an attribute in its instance dictionary for `say_hello` which is just a plain ordinary function.
#
# A function is an object, and it has attributes too:
dir(say_hello_func)
# Notice the `__get__` attribute?
# We know what the `__get__` method looks like when it is called:
# ```
# def __get__(self, instance, owner_class)
# ```
# What's `self` in this case? The function itself (it is the descriptor)
#
# What will `instance` be? The instance we are calling the function from.
#
# What is the `owner_class`? It is the `Person` class in this case.
#
# So let's call the **function** as if it were being called using a dotted notation.
# First we create an instance of `Person`:
p = Person('Alex')
# Now we can call the method this way:
p.say_hello()
# But this is the same as doing this:
say_hello_func.__get__(p, Person)()
# (Note that we do not specify `self` ourselves since we are calling the `__get__` function as a method already bound to `say_hello_func` when we write `say_hello_func.__get__`)
# And that's how functions become "automatically" bound to the instance when calling them using dotted notation.
#
# Remember how we programmed our own descriptors `__get__` when the instance was `None`? We just returned the descriptor itself.
#
# In this case, the descriptor is the function `say_hello` defined in the `Person` class - so we can recover the descriptor from the class using dotted notation too:
Person.say_hello
# This is the same as doing this:
Person.__dict__['say_hello'].__get__(None, Person)
# In fact, just out of curiosity, let's try writing our own "function" class that is both callable and implements a non-data descriptor so we can bind it to instances.
# +
from functools import partial
class CustomFunc:
def __call__(self, instance, *args, **kwargs):
# define the body of the "function" here...
print('instance:', instance)
print('args:', args)
print('kwargs:', kwargs)
return f'{instance.name} says hello!'
def __get__(self, instance, owner_class):
if instance is None:
return self
return partial(self.__call__, instance)
# -
class Person:
def __init__(self, name):
self.name = name
say_hello = CustomFunc()
Person.say_hello
p = Person('Alex')
p.say_hello(1, 2, 3, a=1, b=2)
# Now don't write code like this!! Python provides us functions that implement the descriptor protocol, but I just wanted to show you how we could roughly approximate the same functionality using custom classes.
#
# Nice to gain a better understanding of descriptors, but not at all practical!
# ### Exercise
# Create two data descriptors to handle
# - an integer-only field, named `IntegerField`, with a min and max value (just like we did before)
# - a string-only field, named `CharField`, with a min and max length
#
# For simplicity assume this will only be used for objects (class instances) that have an available `__dict__` - in other words you can use it for instance storage.
#
# After you have done that, use inheritance to create a base descriptor that can factor out the repetitive code from the two descriptors above.
#
# Finally, as a small enhancement, make the `IntegerField` such that `min` and `max` can be unlimited.
# For the `CharField` make it such that values can be assigned without a maximum length.
# #### Solution
# First let's write each data descriptor completely distinct from each other, then we'll determine what we could factor out into a base class.
# +
from numbers import Integral
class IntegerField:
def __init__(self, minimum, maximum):
self.minimum = minimum
self.maximum = maximum
def __set_name__(self, owner_class, name):
self.name = name
def __set__(self, instance, value):
if not isinstance(value, Integral):
raise ValueError(f'{self.name} must be an integer number.')
if value < self.minimum or value > self.maximum:
raise ValueError(f'{self.name} must be between {self.minimum} and {self.maximum}')
instance.__dict__[self.name] = value
def __get__(self, instance, owner_class):
if instance is None:
return self
return instance.__dict__[self.name]
# -
class Person:
age = IntegerField(0, 150)
# Let's test this to make sure it works:
p = Person()
p.age = 10
print(p.age)
try:
p.age = -100
except ValueError as ex:
print(ex)
try:
p.age = 'Unknown'
except ValueError as ex:
print(ex)
# OK, so now let's write the `CharField` descriptor:
class CharField:
def __init__(self, min_length, max_length):
self.min_length = min_length
self.max_length = max_length
def __set_name__(self, owner_class, name):
self.name = name
def __set__(self, instance, value):
if not isinstance(value, str):
raise ValueError(f'{self.name} must be a string.')
if value is None or len(value) < self.min_length or len(value) > self.max_length:
raise ValueError(f'{self.name} length must be between '
f'{self.min_length} and {self.max_length} chars long.')
instance.__dict__[self.name] = value
def __get__(self, instance, owner_class):
if instance is None:
return self
return instance.__dict__[self.name]
class Person:
age = IntegerField(0, 200)
name = CharField(1, 20)
def __init__(self, name, age):
self.name = name
self.age = age
def __repr__(self):
return f'Person(name={self.name}, age={self.age})'
p = Person('Alex', 18)
print(p)
try:
p.name = ''
except ValueError as ex:
print(ex)
try:
p.name = 'a' * 50
except ValueError as ex:
print(ex)
# OK, so now let's see what code appears to be common to these two validators (and would likely be common with other validators too):
class IntegerField:
def __init__(self, minimum, maximum):
self.minimum = minimum
self.maximum = maximum
def __set_name__(self, owner_class, name):
self.name = name
def __set__(self, instance, value):
if not isinstance(value, Integral):
raise ValueError(f'{self.name} must be an integer number.')
if value < self.minimum or value > self.maximum:
raise ValueError(f'{self.name} must be between {self.minimum} and {self.maximum}')
instance.__dict__[self.name] = value
def __get__(self, instance, owner_class):
if instance is None:
return self
return instance.__dict__[self.name]
class CharField:
def __init__(self, min_length, max_length):
self.min_length = min_length
self.max_length = max_length
def __set_name__(self, owner_class, name):
self.name = name
def __set__(self, instance, value):
if not isinstance(value, str):
raise ValueError(f'{self.name} must be a string.')
if len(value) < self.min_length or len(value) > self.max_length:
raise ValueError(f'{self.name} length must be between '
f'{self.min_length} and {self.max_length} chars long.')
instance.__dict__[self.name] = value
def __get__(self, instance, owner_class):
if instance is None:
return self
return instance.__dict__[self.name]
# As we can see we have commonalities in:
# - `__set_name__` (exactly the same)
# - `__get__` (exactly the same)
# - `__set__` (validation tests are different, but storage mechanism is the same)
class ValidatorField:
def __set_name__(self, owner_class, name):
self.name = name
def __get__(self, instance, owner_class):
if instance is None:
return self
return instance.__dict__[self.name]
def __set__(self, instance, value):
instance.__dict__[self.name] = value
# And we can now re-write our validators inheriting from this base `ValidatorField`:
class IntegerField(ValidatorField):
def __init__(self, minimum, maximum):
self.minimum = minimum
self.maximum = maximum
def __set__(self, instance, value):
if not isinstance(value, Integral):
raise ValueError(f'{self.name} must be an integer number.')
if value < self.minimum or value > self.maximum:
raise ValueError(f'{self.name} must be between {self.minimum} and {self.maximum}')
super().__set__(instance, value)
class CharField(ValidatorField):
def __init__(self, min_length, max_length):
self.min_length = min_length
self.max_length = max_length
def __set__(self, instance, value):
if not isinstance(value, str):
raise ValueError(f'{self.name} must be a string.')
if len(value) < self.min_length or len(value) > self.max_length:
raise ValueError(f'{self.name} length must be between '
f'{self.min_length} and {self.max_length} chars long.')
super().__set__(instance, value)
class Person:
age = IntegerField(0, 200)
name = CharField(1, 20)
def __init__(self, name, age):
self.name = name
self.age = age
def __repr__(self):
return f'Person(name={self.name}, age={self.age})'
p = Person('Alex', 18)
p
try:
p.age = -10
except ValueError as ex:
print(ex)
try:
p.name = ''
except ValueError as ex:
print(ex)
# And of course, we can re-use these validators anywhere we need them!
# Now let's make a few enhancements to make our validator a bit more useful.
#
# We'll start with `CharField` and allow unlimited max length:
class CharField(ValidatorField):
def __init__(self, min_length, max_length=None):
self.min_length = min_length
self.max_length = max_length
def __set__(self, instance, value):
if not isinstance(value, str):
raise ValueError(f'{self.name} must be a string.')
if (len(value) < self.min_length
or (self.max_length is not None and len(value) > self.max_length)):
raise ValueError(f'{self.name} length must be between '
f'{self.min_length} and {self.max_length} chars long.')
super().__set__(instance, value)
class Person:
name = CharField(1)
p = Person()
p.name = 'a'
p.name = 'a'*10_000
try:
p.name = ''
except ValueError as ex:
print(ex)
# And now let's do something similar for `IntegerField`:
class IntegerField(ValidatorField):
def __init__(self, minimum=None, maximum=None):
self.minimum = minimum
self.maximum = maximum
def __set__(self, instance, value):
if not isinstance(value, Integral):
raise ValueError(f'{self.name} must be an integer number.')
if ((self.minimum is not None and value < self.minimum) or
(self.maximum is not None and value > self.maximum)):
raise ValueError(f'{self.name} out of bounds.')
super().__set__(instance, value)
class Point2D:
x = IntegerField(0)
y = IntegerField(maximum=10)
p = Point2D()
p.x = 1_000_000
p.y = -1_000_000
try:
p.x = -10
except ValueError as ex:
print(ex)
try:
p.y = 12
except ValueError as ex:
print(ex)
|
python-descriptors.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import selenium
from selenium import webdriver
import bs4
from bs4 import BeautifulSoup
import re
import time
import nltk
from gensim.models import Word2Vec
# +
driver = webdriver.Chrome()
url = 'https://en.wikipedia.org/wiki/Artificial_intelligence'
driver.get(url)
time.sleep(3)
html_doc = driver.page_source
soup = BeautifulSoup(html_doc, 'html.parser')
paragraphs = soup.find_all('p')
article_text = ""
for p in paragraphs:
article_text += p.text
# -
print(article_text)
# Cleans the text
processed_article = article_text.lower()
processed_article = re.sub('[^a-zA-Z]', ' ', processed_article )
processed_article = re.sub(r'\s+', ' ', processed_article)
processed_article
# +
# Converts the article into sentences
all_sentences = nltk.sent_tokenize(processed_article)
# Converts sentences into words
all_words = [nltk.word_tokenize(sent) for sent in all_sentences]
from nltk.corpus import stopwords
for i in range(len(all_words)):
all_words[i] = [w for w in all_words[i] if w not in stopwords.words('english')]
print(all_words)
# +
# min_count=2 specifies to only include words in the word2vec model that appear at least twice in the corpus
word2vec = Word2Vec(all_words, min_count=2)
vocabulary = word2vec.wv.vocab
print(vocabulary)
# -
sim_words = word2vec.wv.most_similar('intelligence')
print(sim_words)
pat = re.compile('artificial|intelligence')
string = 'Artificial intelligence is the'
m = pat.match(string)
|
scripts/Word2Vec_tutorial.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # solve ODE using euler method
from matplotlib import pyplot as plt
import numpy as np
# %matplotlib inline
#euler method
x_init,y_init=0,1
x,y=x_init,y_init
f=lambda x,y:x+y
phi=lambda x,y:f(x,y)
h=0.1
maxiterator=100
xs=[]
ys=[]
for i in range(maxiterator):
xs.append(x)
ys.append(y)
y+=h*phi(x,y)
x+=h
fig,ax=plt.subplots()
ax.plot(xs,ys,'bo')
f_true=lambda x:-x-1+2*np.exp(x)
f_true=np.vectorize(f_true)
ys_true=f_true(xs)
ax.plot(xs,ys_true)
# # solve ODE using heun method
#heun method
x_init,y_init=0,1
x,y=x_init,y_init
f=lambda x,y:x+y
phi=lambda x,y:(f(x,y)+f(x+h,y+h*f(x,y)))/2
h=0.1
maxiterator=100
xs=[]
ys=[]
for i in range(maxiterator):
xs.append(x)
ys.append(y)
y+=h*phi(x,y)
x+=h
fig,ax=plt.subplots()
ax.plot(xs,ys,'bo')
xs=np.array(xs)
f_true=lambda x:-x-1+2*np.exp(x)
ys_true=f_true(xs)
ax.plot(xs,ys_true,'g-')
# # modified euler method
#euler method
x_init,y_init=0,1
x,y=x_init,y_init
f=lambda x,y:x+y
phi=lambda x,y:f(x+h/2.0,y+h*f(x,y)/2.0)
h=0.1
maxiterator=100
xs=[]
ys=[]
for i in range(maxiterator):
xs.append(x)
ys.append(y)
y+=h*phi(x,y)
x+=h
fig,ax=plt.subplots()
ax.plot(xs,ys,'bo')
f_true=lambda x:-x-1+2*np.exp(x)
f_true=np.vectorize(f_true)
ys_true=f_true(xs)
ax.plot(xs,ys_true)
# # solve ODE using Runge-Kutta method
#heun method
x_init,y_init=0,1
x,y=x_init,y_init
f=lambda x,y:x+y
k_1=lambda x,y:f(x,y)
k_2=lambda x,y:f(x+h/2.0,y+h*k_1(x,y)/2.0)
k_3=lambda x,y:f(x+h/2.0,y+h*k_2(x,y)/2.0)
k_4=lambda x,y:f(x+h/2.0,y+h*k_3(x,y))
phi=lambda x,y:(k_1(x,y)+2*k_2(x,y)+2*k_3(x,y)+k_4(x,y))/6.0
h=0.1
maxiterator=100
xs=[]
ys=[]
for i in range(maxiterator):
xs.append(x)
ys.append(y)
y+=h*phi(x,y)
x+=h
fig,ax=plt.subplots()
ax.plot(xs,ys,'bo')
xs=np.array(xs)
f_true=lambda x:-x-1+2*np.exp(x)
ys_true=f_true(xs)
ax.plot(xs,ys_true,'g-')
# # solve higher order ODE
# $$y^{(n)}=f(x,y,y',\dots,y^{(n)}) \quad y^{(k)}(x)\mid_{x=a}\;=y_k$$
# $z_1(x) = y(x), z_2(x)=y'(x), \dots , z_k=y^{(k-1)}, \dots, z_n(x) = y^{(n-1)}(x)$ と置くと,
# $z_1'(x)=z_2(x),\dots , z'_{n-1}(x)=z_n(x),z'_n(x)=f(x,z_1,z_2,\dots,z_n)$
# solve higher order $y''=-y, y(0)=0, y'(0)=-1$
x_init=0.0
y0_init,y1_init=1.0,0.0
x=x_init
y=np.array([y0_init,y1_init])
f0=lambda x,y : y[1]
f1=lambda x,y : -y[0]
f=lambda x,y:np.array([f0(x,y),f1(x,y)])
phi=lambda x,y : (f(x,y)+ f(x+h,y+h*f(x,y)))/2.0
maxiterator=1000
h=0.1
xs=[]
ys=[]
for i in range(maxiterator):
xs.append(x)
ys.append(y.copy())
x+=h
y+=h*phi(x,y)
yplt=np.array(ys).transpose()[1]
plt.plot(xs,yplt)
# solve ODE $y''=F(x,y,y')=-y \quad y(0)=0.0, y'(0)=-1.0$
# +
#solve ODE y^{(maxdeg)}=F(x,y',\dots,y^{(maxdeg-1)})
maxdeg=2
x_init=0.0
#y(0)=0,y'(0)=-1.0
y0_init,y1_init=0.0,-1.0
#F=-y
F =lambda x,y : -y[0]
#set initial value
x=x_init
y=np.array([y0_init,y1_init])
#create tmp array to define fv
fs=[lambda x,y : y[k+1] for k in range(maxdeg-1)]
fs.append(lambda x,y : F(x,y))
fv=lambda x,y:np.array([fs[k](x,y) for k in range(len(fs))])
#apply Heun method
phi=lambda x,y : (fv(x,y)+ fv(x+h,y+h*fv(x,y)))/2.0
maxiterator=1000
h=0.1
xplt,ys=[],[]
for i in range(maxiterator):
xplt.append(x)
ys.append(y.copy())
x+=h
y+=h*phi(x,y)
yplt=np.array(ys).transpose()[0]
plt.plot(xplt,yplt)
# -
|
odeExercise/solveODE.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from nyisotoolkit import NYISOData
from nyisotoolkit.nyisostat.nyisostat import table_load_weighted_price
import matplotlib.pyplot as plt
import seaborn as sns
import pandas as pd
# -
# # Covariance Risk of Renewables
# +
def covariance(year, sources, rt):
lw_lbmp = table_load_weighted_price(year=year, rt=rt).iloc[:,0] #convert to series
fuel_mix = NYISOData(dataset="fuel_mix_5m", year=year).df.tz_convert('US/Eastern') # MW
fuel_mix = (fuel_mix[sources].sum(axis="columns")*1/12) #MW->MWh
if not rt:
fuel_mix = fuel_mix.resample("H").sum()
assert lw_lbmp.index.isin(fuel_mix.index).any(), "Indices are not the same"
assert len(lw_lbmp.index)==len(fuel_mix.index), "Indices are not the same size"
return fuel_mix, lw_lbmp
def renewable_covariance(year, rt):
scatter_kwargs = {"x":None, "y":None,
"hue":None, "style":None,
"size":None, "data":None,
"palette":None,
"hue_order":None, "hue_norm":None,
"sizes":None, "size_order":None, "size_norm":None,
"markers":True, "style_order":None,
"x_bins":None, "y_bins":None, "units":None,
"estimator":None, "ci":95, "n_boot":1000,
"alpha":None, "x_jitter":None, "y_jitter":None,
"legend":'auto',
"ax":None}
for source in ["Wind", "Other Renewables"]:
da_rt = "RT" if rt else "DA"
ax_kwargs = {"title":f'Covariance Risk of {source} ({year})',
"xlabel":f"{source} [MWh]","ylabel":f"State-wide Average {da_rt} Price [$/MWh]",
"xlim":None, "ylim":None}
fuel_mix, lw_lbmp = covariance(year=year, sources=[source], rt=rt)
scatter_kwargs.update({"x": fuel_mix.values,
"y": lw_lbmp.values})
fig, ax = plt.subplots(figsize=(6,3), dpi=300)
sns.scatterplot(**scatter_kwargs)
ax.set(**ax_kwargs)
for rtorda in [False, True]:
renewable_covariance(year=2019, rt=rtorda)
|
docs/ideas/wind_and_solar_covariance_risk.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import json
from web3 import Web3
import web3 as w3
import numpy as np
import pandas as pd
ganache_URL = 'HTTP://127.0.0.1:7545'
web3 = Web3(Web3.HTTPProvider(ganache_URL))
# Set the default account
web3.eth.defaultAccount = web3.eth.accounts[0]
# Create a dataset with all citizens
# Suppose 5 citizens, 2 trucks and 2 plants
assert len(web3.eth.accounts) == 10
# + active=""
# # Get a block
# web3.eth.getBlock('latest')
# + active=""
# abiRemix = json.loads("""""")
# addressRemix = web3.toChecksumAddress("")
# contract = web3.eth.contract(address=addressRemix, abi=abiRemix)
# print(contract.functions.retrieve().call())
# -
json_file = ''.join([i.strip() for i in open('abi.txt', 'r')])
abiRemix = json.loads(json_file)
addressRemix = web3.toChecksumAddress("0x8649045F12c67ab756cA5180310cA062953b9db4")
contract = web3.eth.contract(address=addressRemix, abi=abiRemix)
contract.all_functions()
print(contract.functions.owner().call())
print(contract.functions.numberC().call())
print(contract.functions.numberT().call())
print(contract.functions.numberS().call())
print(contract.functions.citizens(web3.eth.accounts[0]).call())
# Upload data from Municipality
data = pd.read_excel('citizen_data.xlsx', engine="openpyxl")
data['address'] = web3.eth.accounts
assert contract.functions.owner().call() == data.loc[0,'address']
data
# +
# Create CITIZENS - (address payable _address, string memory _name, uint _family, uint _house, uint256 _w)
citizensL = [[r.address, " ".join([r.name, r.surname]), int(r.family), int(r.mq), int(r.weight)]
for r in data.itertuples() if r.role == 'citizen']
# Create TRUCKS - (address _address, bool _recycle)
trucksL = [[r.address, r.recycle] for r in data.itertuples() if r.role == 'truck']
# Create STATIONS - (address _address, bool _recycle, int _lat, int _long)
stationsL = [[r.address, r.recycle, int(r.lat*(10**13)), int(r.long*(10**13))] for r in data.itertuples() if r.role == 'station']
# + active=""
# # Returns the hash of a transaction
# contract.functions.createCitizen(web3.eth.accounts[1], '<NAME>', 4, 70, 0).transact()
# -
def create(c, t, s):
for i in range(len(c)):
contract.functions.createCitizen(c[i][0], c[i][1], c[i][2], c[i][3], c[i][4]).transact()
for j in range(len(t)):
contract.functions.createTruck(t[j][0], t[j][1]).transact()
for k in range(len(s)):
contract.functions.createStation(s[k][0], s[k][1], s[k][2], s[k][3]).transact()
print('Done')
create(citizensL, trucksL, stationsL)
print(contract.functions.numberC().call())
print(contract.functions.numberT().call())
print(contract.functions.numberS().call())
|
old_code/web3_oldVersion.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 数据准备
import numpy as np
import pandas as pd
import random
import matplotlib.pyplot as plt
# %matplotlib inline
from matplotlib.pylab import rcParams
rcParams['figure.figsize'] = 8, 6
pd.options.display.float_format = '{:,.2}'.format
# +
np.random.seed(42)
x = np.array([i * np.pi / 180 for i in range(-180, 60, 5)])
y = np.cos(x) + np.random.normal(0, 0.15, len(x))
data = pd.DataFrame(np.column_stack([x, y]), columns=['x', 'y'])
pow_max = 13
# 构造不同幂的x
for i in range(2, pow_max):
colname = 'x_%d' % i
data[colname] = data['x']**i
# -
data.shape
# 原始x散点图
plt.plot(data['x'],data['y'],'.')
# # 普通线性回归
# +
from sklearn.linear_model import LinearRegression
def myplot(x, y, y_pred, sub, title):
plt.subplot(sub)
plt.tight_layout()
plt.plot(x, y_pred)
plt.plot(x, y, '.')
plt.title(title)
def summary(y, y_pred, intercept_, coef_):
rss = sum((y_pred - y)**2)
ret = [rss]
ret.extend([intercept_])
ret.extend(coef_)
return ret
def linear_regression(data, power, models_to_plot):
#设置预测变量x,x_2,x_3...
predictors = ['x']
if power >= 2:
predictors.extend(['x_%d' % i for i in range(2, power + 1)])
#线性拟合
linreg = LinearRegression(normalize=True)
linreg.fit(data[predictors], data['y'])
y_pred = linreg.predict(data[predictors])
# 绘制指定的幂的图形
if power in models_to_plot:
myplot(data['x'], data['y'], y_pred, \
models_to_plot[power], 'power=%d' % power)
# 记录模型拟合效果rss、截距和系数
return summary(data['y'], y_pred, linreg.intercept_, linreg.coef_)
# +
col = ['rss', 'intercept'] + ['coef_x_%d' % i for i in range(1, pow_max)]
ind = ['pow_%d' % i for i in range(1, pow_max)]
coef_matrix_linear = pd.DataFrame(index=ind, columns=col)
# 设置显示4个图形,分别幂为,1,4,8,12
models_to_plot = {1: 221, 4: 222, 8: 223, 12: 224}
# 拟合所有幂的变量
for i in range(1, pow_max):
coef_matrix_linear.iloc[i - 1, 0:i + 2] = linear_regression(
data, power=i, models_to_plot=models_to_plot)
# -
# 系数矩阵为:
coef_matrix_linear
# # Lasso回归
# +
from sklearn.linear_model import Lasso
def lasso_regression(data, predictors, alpha, models_to_plot):
lassoreg = Lasso(alpha=alpha, normalize=True, max_iter=1e6)
lassoreg.fit(data[predictors], data['y'])
y_pred = lassoreg.predict(data[predictors])
# 绘制指定的alpha的图形
if alpha in models_to_plot:
myplot(data['x'], data['y'], y_pred, \
models_to_plot[alpha], 'alpha=%.3g' % alpha)
# 记录模型拟合效果rss、截距和系数
return summary(data['y'], y_pred, lassoreg.intercept_, lassoreg.coef_)
# +
# 拟合了所有的 x
predictors = ['x']
predictors.extend(['x_%d' % i for i in range(2, pow_max)])
# 设置正则系数
alpha_lasso = [
1e-15, 1e-10, 1e-8, 1e-4, 1e-3, 1e-2, 1e-1, 1, 5, 10, 20, 50
]
# -
ind = ['alpha_%.2g' % alpha_lasso[i] for i in range(0, len(alpha_lasso))]
coef_matrix_lasso = pd.DataFrame(index=ind, columns=col)
# +
models_to_plot = {1e-15: 221, 1e-3: 222, 1e-2: 223, 1e-1: 224}
for i in range(len(alpha_lasso)):
coef_matrix_lasso.iloc[i, ] = lasso_regression(data, predictors,
alpha_lasso[i],
models_to_plot)
# +
pd.options.display.float_format = '{:,.2g}'.format
coef_matrix_lasso
# -
# # ridge回归
# +
from sklearn.linear_model import Ridge
def ridge_regression(data, predictors, alpha, models_to_plot):
ridgereg = Ridge(alpha=alpha, normalize=True)
ridgereg.fit(data[predictors], data['y'])
y_pred = ridgereg.predict(data[predictors])
# 绘制指定的alpha的图形
if alpha in models_to_plot:
myplot(data['x'], data['y'], y_pred, \
models_to_plot[alpha], 'alpha=%.3g' % alpha)
# 记录模型拟合效果rss、截距和系数
return summary(data['y'], y_pred, ridgereg.intercept_, ridgereg.coef_)
# +
# 拟合了所有的 x
predictors = ['x']
predictors.extend(['x_%d' % i for i in range(2, pow_max)])
# 设置正则系数
alpha_ridge = [1e-15, 1e-10, 1e-8, 1e-4, 1e-3, 1e-2, 1e-1, 1, 5, 10, 20, 50]
# +
ind = ['alpha_%.2g' % alpha_ridge[i] for i in range(0, len(alpha_ridge))]
coef_matrix_ridge = pd.DataFrame(index=ind, columns=col)
models_to_plot = {1e-15: 221, 1e-3: 222, 1: 223, 50: 224}
for i in range(len(alpha_ridge)):
coef_matrix_ridge.iloc[i, ] = ridge_regression(data, predictors,
alpha_ridge[i],
models_to_plot)
# -
pd.options.display.float_format = '{:,.2g}'.format
coef_matrix_ridge
|
ch09-linear_model/src/linear_model.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # LAB 04.01 - Cleaning Data
# +
# !wget --no-cache -O init.py -q https://raw.githubusercontent.com/rramosp/20201.xai4eng/master/content/init.py
import init; init.init(force_download=False); init.get_weblink()
init.endpoint
# -
from local.lib.rlxmoocapi import submit, session
student = session.Session(init.endpoint).login( course_id=init.course_id,
session_id="UDEA",
lab_id="L04.01" )
init.get_weblink()
# observe the following synthetic example with missing data
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
from IPython.display import Image
import numpy as np
import seaborn as sns
n = 20
place = np.r_[["Medellin", "Bogota", "Madrid"]][(np.random.randint(3, size=n))]
age = np.random.randint(50, size=n)+10
children = np.r_[[(np.random.randint(2) if i<30 else (np.random.randint(4))) for i in age]]
risk = np.r_[[np.random.random()*(.2 if i=="Medellin" else .8) for i in place]].round(3)
risk[np.random.permutation(len(risk))[:5]]=np.nan
# +
d01 = pd.DataFrame([age, risk, children, place], index=["age", "risk", "children", "place"]).T
d01.to_csv("risk.csv", index=False)
d01
# -
# observe, in particular, that risk in Medellín is usually lower than in Bogotá, so we will try to fix missing data using this fact.
# +
k = d01[d01.place=="Bogota"]["risk"].dropna()
plt.scatter(k, [0]*len(k), label="Bogota")
k = d01[d01.place=="Medellin"]["risk"].dropna()
plt.scatter(k, [1]*len(k), label="Medellin")
k = d01[d01.place=="Madrid"]["risk"].dropna()
plt.scatter(k, [2]*len(k), label="Madrid")
plt.grid();
plt.xlabel("risk level")
plt.ylabel("city")
plt.legend()
# -
# ## Task 1. FillNA in `risk` with corresponding city average
#
# Observe that the above dataframe has been stored in the file `risk.csv`. You will have to fill in the missing values in the **risk** column with the related city mean in the following way:
#
# 1. Download the file `risk.csv`
# 1. Compute the mean risk **per city**
# 1. Substitute any missing value in the risk column by the corresponding city mean
# 1. Create a new csv file named `risk_fixed.csv`, with the **exact** same structure but with the missing values replaced
# 1. Upload your `risk_fixed.csv` file to the notebook environment
# 1. Run the evaluation cell below
#
# ### Use the tool of your choice
# (Excel, Orange, your programming language, or even this notebook if you can program python)
#
# **For Python**, you do not have to download and upload anything, just use Pandas and store the resulting dataset in the variable `r01`
#
# **use three decimal places for precision**
#
# ### Example
# for the following data
Image("local/imgs/cities.png", width=200)
# you must create a file with the following content
Image("local/imgs/cities-riskfree.png", width=200)
# **your solution**
r01 = pd.read_csv("risk_fixed.csv")
r01
# #### submit your answer
student.submit_task(globals(), task_id="task_01");
# ## Task 2. Standardize `age` so that min=0, max=1
#
# Standardizing values is, in certain cases, a necesity for ML models, providing stability and increased performance.
#
# In this task you will have to standardize the column `age` so that all values stay in the [0,1] interval. Given any value $x_i$, its corresponsing stardardized value $s_i$ will be:
#
# $$s_i = \frac{s_i-min}{max-min}$$
#
# where $min$, $max$ is the minimum and maximum ages respectively
#
# You must use again the file `risk.csv` and create and upload a file named `age_minmax.csv` with your answer. You should **only modify** the `age` column, leaving the rest as you find them in the csv file.
#
# **For Python**, you do not have to download and upload anything, just use Pandas and store the resulting dataset in the variable `r02`
#
# For the previous example, the correct answer would be
Image("local/imgs/cities-ageminmax.png")
# **load your file**
r02 = pd.read_csv("age_minmax.csv")
r02
# #### submit your answer
student.submit_task(globals(), task_id="task_02");
# ## Task 3. Standardize `age` so that $\mu=0$ and $\sigma=1$
#
#
# In this task you will have to standardize the column `age` so that all values stay have zero mean and standard deviation of 1. Given any value $x_i$, its corresponsing stardardized value $s_i$ will be:
#
# $$s_i = \frac{s_i-\mu}{\sigma}$$
#
# where $\mu$ is the mean of all age values, and $\sigma$ is the standard deviation.
#
# You must use again the file `risk.csv` and create and upload a file named `age_meanstd.csv` with your answer. You should **only modify** the `age` column, leaving the rest as you find them in the csv file.
#
# **For Python**, you do not have to download and upload anything, just use Pandas and store the resulting dataset in the variable `r03`
#
# For the previous example, the correct answer would be
Image("local/imgs/cities-agemeanstd.png")
# **load your file**
r03 = pd.read_csv("age_meanstd.csv")
r03
# #### submit your answer
student.submit_task(globals(), task_id="task_03");
# ## Task 4. Create a one-hot encoding for `place`
#
# substitute the column `place` for three new columns with **onehot** encoding. You must use again the file `risk.csv` and create and upload a file named `place_onehot.csv` with your answer.
#
# **For Python**, you do not have to download and upload anything, just use Pandas and store the resulting dataset in the variable `r04`
#
# The solution for the example above should look like this. Observe that **you must name the columns** as shown here:
Image("local/imgs/cities_onehot.png")
# **load your file**
r04 = pd.read_csv("place_onehot.csv")
r04
# #### submit your answer
student.submit_task(globals(), task_id="task_04");
|
content/LAB 04.01 - CLEANING DATA.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
from sklearn.preprocessing import LabelEncoder, StandardScaler, MinMaxScaler
from sklearn.pipeline import make_pipeline
from sklearn.compose import make_column_transformer
from sklearn.model_selection import train_test_split, cross_val_score
from sklearn import svm
from sklearn.tree import DecisionTreeClassifier
import matplotlib.pyplot as plt
from imblearn.over_sampling import SMOTE
from helper import get_performance
EPOCHS = 700
BATCH_SIZE = 2048
ACTIVATION = 'swish'
LEARNING_RATE = 0.0007
FOLDS = 5
# +
# Reading the dataset
data = pd.read_csv("dataset/Hotel_Booking/hotel_bookings.csv")
data = data.sample(frac=0.2, replace=True, random_state=1).reset_index(drop=True)
data = data.drop(['company'], axis = 1)
data['children'] = data['children'].fillna(0)
data['hotel'] = data['hotel'].map({'Resort Hotel':0, 'City Hotel':1})
data['arrival_date_month'] = data['arrival_date_month'].map({'January':1, 'February': 2, 'March':3, 'April':4, 'May':5, 'June':6, 'July':7,
'August':8, 'September':9, 'October':10, 'November':11, 'December':12})
def family(data):
if ((data['adults'] > 0) & (data['children'] > 0)):
val = 1
elif ((data['adults'] > 0) & (data['babies'] > 0)):
val = 1
else:
val = 0
return val
def deposit(data):
if ((data['deposit_type'] == 'No Deposit') | (data['deposit_type'] == 'Refundable')):
return 0
else:
return 1
def feature(data):
data["is_family"] = data.apply(family, axis = 1)
data["total_customer"] = data["adults"] + data["children"] + data["babies"]
data["deposit_given"] = data.apply(deposit, axis=1)
data["total_nights"] = data["stays_in_weekend_nights"]+ data["stays_in_week_nights"]
return data
data = feature(data)
# Information of these columns is also inside of new features, so it is better to drop them.
# I did not drop stays_nights features, I can't decide which feature is more important there.
data = data.drop(columns = ['adults', 'babies', 'children', 'deposit_type', 'reservation_status_date'])
indices = data.loc[pd.isna(data["country"]), :].index
data = data.drop(data.index[indices])
data = data.drop(columns = ['arrival_date_week_number', 'stays_in_weekend_nights', 'arrival_date_month', 'agent'], axis = 1)
df1 = data.copy()
#one-hot-encoding
df1 = pd.get_dummies(data = df1, columns = ['meal', 'market_segment', 'distribution_channel',
'reserved_room_type', 'assigned_room_type', 'customer_type', 'reservation_status'])
le = LabelEncoder()
df1['country'] = le.fit_transform(df1['country'])
# There are more than 300 classes, so I wanted to use label encoder on this feature.
df2 = df1.drop(columns = ['reservation_status_Canceled', 'reservation_status_Check-Out', 'reservation_status_No-Show'], axis = 1)
df2.rename(columns={'market_segment_Offline TA/TO' : 'market_segment_Offline_TA_TO',
'market_segment_Online TA' : 'market_segment_Online_TA',
'distribution_channel_TA/TO' : 'distribution_channel_TA_TO',
'customer_type_Transient-Party' : 'customer_type_Transient_Party'}, inplace=True)
y = df2["is_canceled"]
X = df2.drop(["is_canceled"], axis=1)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.30, random_state = 42)
print("Train data: ", X_train.shape)
print("Test data: ", X_test.shape)
# -
y_train.value_counts()
y_test.value_counts()
# # Default Model
model_default = svm.SVC()
scores_default = cross_val_score(model_default, X=X_train, y=y_train, cv = FOLDS)
model_default.fit(X_train, y_train)
y_pred_default = model_default.predict(X_test)
get_performance(X_test, y_test, y_pred_default)
pd.DataFrame(y_pred_default).value_counts()
import time
import sys
sys.path.insert(1, './mmd')
from mmd import diagnoser
from scipy import stats as st
import numpy
#notebook's library
# %matplotlib inline
from helper import get_top_f1_rules, get_relevent_attributs_target, get_MMD_results, get_biased_features, get_BGMD_results
from helper import generateTrain_data_Weights
default_result = pd.concat([X_test, y_test], axis=1, join='inner')
default_result.loc[:,"pred"] = y_pred_default
def mispredict_label(row):
if row['is_canceled'] == row['pred']:
return False
return True
default_result_copy = default_result.copy()
X_test_copy = X_test.copy()
X_test_copy['mispredict'] = default_result_copy.apply(lambda row: mispredict_label(row), axis=1)
# +
settings = diagnoser.Settings
settings.all_rules = True
# Get relevent attributes and target
relevant_attributes, Target = get_relevent_attributs_target(X_test_copy)
# Generate MMD rules and correspodning information
MMD_rules, MMD_time, MMD_Features = get_MMD_results(X_test_copy, relevant_attributes, Target)
#Get biased attributes this time
biased_attributes = get_biased_features(X_test_copy, relevant_attributes)
BGMD_rules, BGMD_time, BGMD_Features = get_BGMD_results(X_test_copy, biased_attributes, Target)
print('MMD Spent:', MMD_time, 'BGMD Spent:', BGMD_time)
MMD_rules, BGMD_rules
# -
# # Decision Tree
model_default = DecisionTreeClassifier()
scores_default = cross_val_score(model_default, X=X_train, y=y_train, cv = FOLDS)
model_default.fit(X_train, y_train)
y_pred_default = model_default.predict(X_test)
get_performance(X_test, y_test, y_pred_default)
# +
default_result = pd.concat([X_test, y_test], axis=1, join='inner')
default_result.loc[:,"pred"] = y_pred_default
default_result_copy = default_result.copy()
X_test_copy = X_test.copy()
X_test_copy['mispredict'] = default_result_copy.apply(lambda row: mispredict_label(row), axis=1)
settings = diagnoser.Settings
settings.all_rules = True
# Get relevent attributes and target
relevant_attributes, Target = get_relevent_attributs_target(X_test_copy)
# Generate MMD rules and correspodning information
MMD_rules, MMD_time, MMD_Features = get_MMD_results(X_test_copy, relevant_attributes, Target)
#Get biased attributes this time
biased_attributes = get_biased_features(X_test_copy, relevant_attributes)
BGMD_rules, BGMD_time, BGMD_Features = get_BGMD_results(X_test_copy, biased_attributes, Target)
print('MMD Spent:', MMD_time, 'BGMD Spent:', BGMD_time)
MMD_rules, BGMD_rules
|
BGMD/RQ1_Hotel_Bookings.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Granero0011/AB-Demo/blob/master/DS_Unit_2_Sprint_Challenge_3_Classification_Validation.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="PC9RfopIWrc9"
# _Lambda School Data Science Unit 2_
#
# # Classification & Validation Sprint Challenge
# + [markdown] colab_type="text" id="UV7ArLFQN84W"
# Follow the instructions for each numbered part to earn a score of 2. See the bottom of the notebook for a list of ways you can earn a score of 3.
# + [markdown] colab_type="text" id="bAZcbTtiUlkI"
# #### For this Sprint Challenge, you'll predict whether a person's income exceeds $50k/yr, based on census data.
#
# You can read more about the Adult Census Income dataset at the UCI Machine Learning Repository: https://archive.ics.uci.edu/ml/datasets/adult
# + [markdown] id="YHMuDE2ERkXt" colab_type="text"
# #### Run this cell to load the data:
# + colab_type="code" id="gvV9VORbxyvu" colab={}
import pandas as pd
columns = ['age',
'workclass',
'fnlwgt',
'education',
'education-num',
'marital-status',
'occupation',
'relationship',
'race',
'sex',
'capital-gain',
'capital-loss',
'hours-per-week',
'native-country',
'income']
df = pd.read_csv('https://archive.ics.uci.edu/ml/machine-learning-databases/adult/adult.data',
header=None, names=columns)
df['income'] = df['income'].str.strip()
# + [markdown] id="L4fXtSxURkXy" colab_type="text"
# ## Part 1 — Begin with baselines
#
# Split the data into an **X matrix** (all the features) and **y vector** (the target).
#
# (You _don't_ need to split the data into train and test sets here. You'll be asked to do that at the _end_ of Part 1.)
# + id="Wip-pD3eRkXz" colab_type="code" colab={}
#Let's split the data
X=df.drop('income', axis=1);
y= df['income']
# + [markdown] colab_type="text" id="IxKfgx4ycb3c"
# What **accuracy score** would you get here with a **"majority class baseline"?**
#
# (You can answer this question either with a scikit-learn function or with a pandas function.)
# + id="8AS22eIDWy1X" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="6abba8a1-3ce2-4906-9d44-e6a284ed7e3f"
#baseline
y.value_counts(normalize=True)
# + id="gjMgSFdkXuLn" colab_type="code" colab={}
majority_class = y.mode()[0]
y_pred = [majority_class] * len(y)
# + id="_7S23QFbXxjA" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="f1e1b2f6-fd8e-4797-b578-e26fa4b21e22"
#accuracu score
from sklearn.metrics import accuracy_score
accuracy_score(y, y_pred)
# + [markdown] colab_type="text" id="_KdxE1TrcriI"
# What **ROC AUC score** would you get here with a **majority class baseline?**
#
# (You can answer this question either with a scikit-learn function or with no code, just your understanding of ROC AUC.)
# + colab_type="code" id="ILS0fN0Cctyc" colab={"base_uri": "https://localhost:8080/", "height": 343} outputId="6d54c0f0-9d5f-4de8-9542-6332ccda809e"
#ROC_AUC score
from sklearn.metrics import roc_auc_score
roc_auc_score(y, y_pred)
# + [markdown] colab_type="text" id="QqYNDtwKYhji"
# In this Sprint Challenge, you will use **"Cross-Validation with Independent Test Set"** for your model validaton method.
#
# First, **split the data into `X_train, X_test, y_train, y_test`**. You can include 80% of the data in the train set, and hold out 20% for the test set.
# + colab_type="code" id="mPKf86yDYf0t" colab={}
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.2)
# + [markdown] id="VgpdQXR1RkYG" colab_type="text"
# ## Part 2 — Modeling with Logistic Regression!
# + [markdown] colab_type="text" id="E_ATNJdqTCuZ"
# - You may do exploratory data analysis and visualization, but it is not required.
# - You may **use all the features, or select any features** of your choice, as long as you select at least one numeric feature and one categorical feature.
# - **Scale your numeric features**, using any scikit-learn [Scaler](https://scikit-learn.org/stable/modules/classes.html#module-sklearn.preprocessing) of your choice.
# - **Encode your categorical features**. You may use any encoding (One-Hot, Ordinal, etc) and any library (category_encoders, scikit-learn, pandas, etc) of your choice.
# - You may choose to use a pipeline, but it is not required.
# - Use a **Logistic Regression** model.
# - Use scikit-learn's [**cross_val_score**](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.cross_val_score.html) function. For [scoring](https://scikit-learn.org/stable/modules/model_evaluation.html#the-scoring-parameter-defining-model-evaluation-rules), use **accuracy**.
# - **Print your model's cross-validation accuracy score.**
# + id="3ZgowKurRkYI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 343} outputId="3c3473ec-46c7-4bb9-c5ba-fdf9e63cea0a"
#Let's check this out
X.head(5)
# + id="aqn95jpFb2z4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 275} outputId="283674fa-bb1d-48bc-b28c-0cee4e98134e"
# !pip install category_encoders
# + id="9WUwRy7HZTI2" colab_type="code" colab={}
import category_encoders as ce
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import StandardScaler
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import cross_val_score
# + id="zI_-4SgyjvW2" colab_type="code" colab={}
import warnings
from sklearn.exceptions import DataConversionWarning
warnings.filterwarnings(action='ignore', category=DataConversionWarning)
# + id="FGcjf6Ajb9DE" colab_type="code" colab={}
#features and target
features= ['age','education', 'education-num','hours-per-week']
target='income'
X= df[features]
y= df[target]
# + id="UmKcWlYbh-lH" colab_type="code" colab={}
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.2)
# + id="GBZILea2gI6Q" colab_type="code" colab={}
# let's create a pipeline
pipeline = make_pipeline(
ce.OneHotEncoder(),
StandardScaler(),
LogisticRegression(solver='lbfgs')
)
# + id="OK6FR1GJiPWd" colab_type="code" colab={}
#Let's calculate the cross validation scores
scores = cross_val_score(pipeline, X_train, y_train, scoring='accuracy', cv=10)
# + id="7-vhx8Ioj076" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="f069412e-346f-4e4d-bb41-85835c274dc7"
#Let's print it
print('Cross-Validation Accuracy scores:', scores)
print('Average:', scores.mean())
# + [markdown] id="7GHivozyRkYM" colab_type="text"
# ## Part 3 — Modeling with Tree Ensembles!
#
# Part 3 is the same as Part 2, except this time, use a **Random Forest** or **Gradient Boosting** classifier. You may use scikit-learn, xgboost, or any other library. Then, print your model's cross-validation accuracy score.
# + id="h7Ck8rVBlD4b" colab_type="code" colab={}
from sklearn.ensemble import RandomForestClassifier
# + id="AitOGFsunzk4" colab_type="code" colab={}
#Let's create a new pipeline
pipeline = make_pipeline(
ce.OneHotEncoder(),
StandardScaler(),
RandomForestClassifier(max_depth=3, n_estimators=100, n_jobs=-1, random_state=42))
# + id="rbiIsG7fnHET" colab_type="code" colab={}
scores = cross_val_score(pipeline,X_train, y_train, scoring='accuracy', cv=10)
# + id="yxHH4ZgNoZUt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="624ca443-96a0-4e96-e38c-4e8a4eeb233f"
#Let's print all this
print('Cross-Validation Accuracy scores:', scores)
print('Average:', scores.mean())
# + [markdown] colab_type="text" id="jkyHoRIbEgRR"
# ## Part 4 — Calculate classification metrics from a confusion matrix
#
# Suppose this is the confusion matrix for your binary classification model:
#
# <table>
# <tr>
# <td colspan="2" rowspan="2"></td>
# <td colspan="2">Predicted</td>
# </tr>
# <tr>
# <td>Negative</td>
# <td>Positive</td>
# </tr>
# <tr>
# <td rowspan="2">Actual</td>
# <td>Negative</td>
# <td style="border: solid">85</td>
# <td style="border: solid">58</td>
# </tr>
# <tr>
# <td>Positive</td>
# <td style="border: solid">8</td>
# <td style="border: solid"> 36</td>
# </tr>
# </table>
# + [markdown] colab_type="text" id="LhyMM5H-JpVB"
# Calculate accuracy
# + colab_type="code" id="TZPwqdh2KUcB" colab={}
# accuracy = (36+85)/(85+58+8+36) =0.647
# + [markdown] colab_type="text" id="BRWLfGcGKeQw"
# Calculate precision
# + colab_type="code" id="A-FEZ4i_Kf_n" colab={}
# precision =(36)/36+58=0.383
# + [markdown] colab_type="text" id="h_mH2NYDKi2C"
# Calculate recall
# + colab_type="code" id="U4_wJGyjKkXJ" colab={}
#recall = 36/(36+8)=0.81
# + [markdown] colab_type="text" id="9KEaWsk5Kk9W"
# ## BONUS — How you can earn a score of 3
#
# ### Part 1
# Do feature engineering, to try improving your cross-validation score.
#
# ### Part 2
# Experiment with feature selection, preprocessing, categorical encoding, and hyperparameter optimization, to try improving your cross-validation score.
#
# ### Part 3
# Which model had the best cross-validation score? Refit this model on the train set and do a final evaluation on the held out test set — what is the test score?
#
# ### Part 4
# Calculate F1 score and False Positive Rate.
|
DS_Unit_2_Sprint_Challenge_3_Classification_Validation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from collections import Counter
import pandas as pd
import numpy as np
# +
clinical = pd.read_csv('data/clinical.tsv', sep='\t', index_col='ID')
genefpkm = pd.read_csv('data/gene_count.tsv', sep='\t', index_col='ID')
pd.DataFrame({'Rows': [genefpkm.shape[0], clinical.shape[0]],
'Columns': [genefpkm.shape[1], clinical.shape[1]]}, index=['GENEFPKM', 'CLINICAL'])
# +
joined = clinical.join(genefpkm, how='inner')
pd.DataFrame({'Rows': [joined.shape[0]],
'Columns': [joined.shape[1]]}, index=['JOINED'])
# -
pd.DataFrame(dict(Counter(joined['therapy_first_line_class'])), index=['Count'])
pd.DataFrame(Counter(joined['response_best_response_first_line'].replace(np.nan, 'nan')), index=['Count'])
pd.DataFrame(Counter(joined['response_days_to_disease_progression'].replace(np.nan, 'nan')), index=['Count'])
# +
df_dtdp = clinical.copy()
del df_dtdp['response_best_response_first_line']
df_dtdp = df_dtdp.loc[~df_dtdp['response_days_to_disease_progression'].isnull()]
df_dtdp.to_csv('data/clinical_dtdp.tsv', sep='\t', index=True)
pd.DataFrame(Counter(df_dtdp['response_days_to_disease_progression'].replace(np.nan, 'nan')), index=['Count'])
# +
df_brfl = clinical.copy()
del df_brfl['response_days_to_disease_progression']
df_brfl = df_brfl.loc[~df_brfl['response_best_response_first_line'].isnull()]
df_brfl.to_csv('data/clinical_brfl.tsv', sep='\t', index=True)
pd.DataFrame(Counter(df_brfl['response_best_response_first_line'].replace(np.nan, 'nan')), index=['Count'])
# -
pd.DataFrame(Counter(df_brfl['therapy_first_line_class']), index=['Count'])
pd.DataFrame(Counter(df_dtdp['therapy_first_line_class']), index=['Count'])
|
00_03_clean_per_response.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="ZAmqzg-o_CSz"
# ##### Copyright 2020 Google LLC.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# + executionInfo={"elapsed": 187, "status": "ok", "timestamp": 1616167471036, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="XSfSUHE6_QHV"
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="vYCGvyyWrtPb"
# # Meta-Dataset leaderboard
#
# This notebook computes leaderboard tables for different models on Meta-Dataset.
#
# Results for each model in each training setting (ImageNet-only or all datasets) are defined in a different DataFrame. This script aggregates the data in one DataFrame, ranks the models in each setting (using a statistical test for equality), and produces the final tables.
# + executionInfo={"elapsed": 160, "status": "ok", "timestamp": 1616167471208, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="LLfLWaldshP_"
import re
import textwrap
import numpy as np
import pandas as pd
from IPython import display
# + executionInfo={"elapsed": 120, "status": "ok", "timestamp": 1616167471340, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="SduvTxvOsmFO"
# Explicit list of evaluation datasets.
# ILSVRC (valid) is included for completeness, but does not have to be reported.
datasets = [
"ILSVRC (valid)",
"ILSVRC (test)",
"Omniglot",
"Aircraft",
"Birds",
"Textures",
"QuickDraw",
"Fungi",
"VGG Flower",
"Traffic signs",
"MSCOCO"
]
# Explicit list of articles and references, filled in throughout the notebook.
references = []
# + [markdown] id="G1Ooh3dZr3z2"
# ## Results from Triantafillou et al. (2020)
# + executionInfo={"elapsed": 114, "status": "ok", "timestamp": 1616167471469, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="o4fTwLvbnB7A"
ref = ("Triantafillou et al. (2020)",
"<NAME>, <NAME>, <NAME>, <NAME>, "
"<NAME>, <NAME>, <NAME>, <NAME>, <NAME>, "
"<NAME>, <NAME>; "
"[_Meta-Dataset: A Dataset of Datasets for Learning to Learn from Few "
"Examples_](https://arxiv.org/abs/1903.03096); ICLR 2020.")
references.append(ref)
# display.display(display.Markdown(ref[1]))
# + [markdown] id="iWhbg1_Qr_K-"
# ### k-NN (`baseline`)
# + executionInfo={"elapsed": 153, "status": "ok", "timestamp": 1616167471631, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="hDidqhwSr-fS"
baseline_imagenet_df = pd.DataFrame(
columns=['mean (%)', '95% CI', '# episodes'],
index=datasets
)
baseline_imagenet_df['# episodes'] = 600
# + executionInfo={"elapsed": 154, "status": "ok", "timestamp": 1616167471798, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="90ULGCOyrlK4"
baseline_imagenet_df.loc[datasets[1:], ['mean (%)', '95% CI']] = [
[41.03, 1.01],
[37.07, 1.15],
[46.81, 0.89],
[50.13, 1.00],
[66.36, 0.75],
[32.06, 1.08],
[36.16, 1.02],
[83.10, 0.68],
[44.59, 1.19],
[30.38, 0.99]
]
# + colab={"height": 381} executionInfo={"elapsed": 213, "status": "ok", "timestamp": 1616167472026, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="6bIjgidkwZu3" outputId="890678b4-a3d4-4b6b-d5ba-cda682a3dbe8"
baseline_imagenet_df
# + colab={"height": 381} executionInfo={"elapsed": 189, "status": "ok", "timestamp": 1616167472404, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="M4XqRjuBwbvO" outputId="97c98b30-22aa-4155-b1de-44e74092b16c"
baseline_all_df = pd.DataFrame(
columns=['mean (%)', '95% CI', '# episodes'],
index=datasets
)
baseline_all_df['# episodes'] = 600
baseline_all_df.loc[datasets[1:], ['mean (%)', '95% CI']] = [
[38.55, 0.94],
[74.60, 1.08],
[64.98, 0.82],
[66.35, 0.92],
[63.58, 0.79],
[44.88, 1.05],
[37.12, 1.06],
[83.47, 0.61],
[40.11, 1.10],
[29.55, 0.96]
]
baseline_all_df
# + [markdown] id="b19z3rD21o9b"
# ### Finetune (`baselinefinetune`)
# + colab={"height": 381} executionInfo={"elapsed": 188, "status": "ok", "timestamp": 1616167472752, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="MSgDpmn92bHk" outputId="5d005b35-f852-4fdb-b8ed-272f2eb3a94d"
baselineft_imagenet_df = pd.DataFrame(
columns=['mean (%)', '95% CI', '# episodes'],
index=datasets
)
baselineft_imagenet_df['# episodes'] = 600
baselineft_imagenet_df.loc[datasets[1:], ['mean (%)', '95% CI']] = [
[45.78, 1.10],
[60.85, 1.58],
[68.69, 1.26],
[57.31, 1.26],
[69.05, 0.90],
[42.60, 1.17],
[38.20, 1.02],
[85.51, 0.68],
[66.79, 1.31],
[34.86, 0.97]
]
baselineft_imagenet_df
# + colab={"height": 381} executionInfo={"elapsed": 232, "status": "ok", "timestamp": 1616167473137, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="D_aP8Bhf2a4m" outputId="3a2e299b-6b24-4664-cb5e-effada0177aa"
baselineft_all_df = pd.DataFrame(
columns=['mean (%)', '95% CI', '# episodes'],
index=datasets
)
baselineft_all_df['# episodes'] = 600
baselineft_all_df.loc[datasets[1:], ['mean (%)', '95% CI']] = [
[43.08, 1.08],
[71.11, 1.37],
[72.03, 1.07],
[59.82, 1.15 ],
[69.14, 0.85],
[47.05, 1.16 ],
[38.16, 1.04],
[85.28, 0.69],
[66.74, 1.23],
[35.17, 1.08]
]
baselineft_all_df
# + [markdown] id="eQyNDVQ3guwR"
# ### MatchingNet (`matching`)
# + colab={"height": 381} executionInfo={"elapsed": 166, "status": "ok", "timestamp": 1616167473459, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="tjrqS-YzhL51" outputId="25ba4fe8-2f21-4cac-b20b-df0718936f14"
matching_imagenet_df = pd.DataFrame(
columns=['mean (%)', '95% CI', '# episodes'],
index=datasets
)
matching_imagenet_df['# episodes'] = 600
matching_imagenet_df.loc[datasets[1:], ['mean (%)', '95% CI']] = [
[45.00, 1.10],
[52.27, 1.28],
[48.97, 0.93],
[62.21, 0.95],
[64.15, 0.85],
[42.87, 1.09],
[33.97, 1.00],
[80.13, 0.71],
[47.80, 1.14],
[34.99, 1.00]
]
matching_imagenet_df
# + colab={"height": 381} executionInfo={"elapsed": 180, "status": "ok", "timestamp": 1616167473790, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="iJC5oVQXhPgA" outputId="5e6f247c-f8c5-4504-95e6-379f95a7bdce"
matching_all_df = pd.DataFrame(
columns=['mean (%)', '95% CI', '# episodes'],
index=datasets
)
matching_all_df['# episodes'] = 600
matching_all_df.loc[datasets[1:], ['mean (%)', '95% CI']] = [
[36.08, 1.00],
[78.25, 1.01],
[69.17, 0.96],
[56.40, 1.00],
[61.80, 0.74],
[60.81, 1.03],
[33.70, 1.04],
[81.90, 0.72],
[55.57, 1.08],
[28.79, 0.96]
]
matching_all_df
# + [markdown] id="yN18Y280jgUl"
# ### ProtoNet (`prototypical`)
# + colab={"height": 381} executionInfo={"elapsed": 229, "status": "ok", "timestamp": 1616167474173, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="G81j2wbrjnYZ" outputId="e86426e5-2322-4809-d79b-34825ec08c94"
prototypical_imagenet_df = pd.DataFrame(
columns=['mean (%)', '95% CI', '# episodes'],
index=datasets
)
prototypical_imagenet_df['# episodes'] = 600
prototypical_imagenet_df.loc[datasets[1:], ['mean (%)', '95% CI']] = [
[50.50, 1.08],
[59.98, 1.35],
[53.10, 1.00],
[68.79, 1.01],
[66.56, 0.83],
[48.96, 1.08],
[39.71, 1.11],
[85.27, 0.77],
[47.12, 1.10],
[41.00, 1.10]
]
prototypical_imagenet_df
# + colab={"height": 381} executionInfo={"elapsed": 282, "status": "ok", "timestamp": 1616167474621, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="ZpH3yku4jptm" outputId="0e37f6af-1263-4f70-f82f-2283a979c98d"
prototypical_all_df = pd.DataFrame(
columns=['mean (%)', '95% CI', '# episodes'],
index=datasets
)
prototypical_all_df['# episodes'] = 600
prototypical_all_df.loc[datasets[1:], ['mean (%)', '95% CI']] = [
[44.50, 1.05],
[79.56, 1.12],
[71.14, 0.86],
[67.01, 1.02],
[65.18, 0.84],
[64.88, 0.89],
[40.26, 1.13],
[86.85, 0.71],
[46.48, 1.00],
[39.87, 1.06]
]
prototypical_all_df
# + [markdown] id="4qiZg0G0k2X1"
# ### fo-MAML (`maml`)
# + colab={"height": 381} executionInfo={"elapsed": 172, "status": "ok", "timestamp": 1616167474948, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="SIgkFZLelCDR" outputId="3bca5c87-6426-4dbe-eb71-ef47b80bc516"
maml_imagenet_df = pd.DataFrame(
columns=['mean (%)', '95% CI', '# episodes'],
index=datasets
)
maml_imagenet_df['# episodes'] = 600
maml_imagenet_df.loc[datasets[1:], ['mean (%)', '95% CI']] = [
[45.51, 1.11],
[55.55, 1.54],
[56.24, 1.11],
[63.61, 1.06],
[68.04, 0.81],
[43.96, 1.29],
[32.10, 1.10],
[81.74, 0.83],
[50.93, 1.51],
[35.30, 1.23]
]
maml_imagenet_df
# + colab={"height": 381} executionInfo={"elapsed": 301, "status": "ok", "timestamp": 1616167475470, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="fb_hmcotlEYp" outputId="4395e46b-0622-46bf-e5b7-481e0a35b70e"
maml_all_df = pd.DataFrame(
columns=['mean (%)', '95% CI', '# episodes'],
index=datasets
)
maml_all_df['# episodes'] = 600
maml_all_df.loc[datasets[1:], ['mean (%)', '95% CI']] = [
[37.83, 1.01],
[83.92, 0.95],
[76.41, 0.69],
[62.43, 1.08],
[64.16, 0.83],
[59.73, 1.10],
[33.54, 1.11],
[79.94, 0.84],
[42.91, 1.31],
[29.37, 1.08]
]
maml_all_df
# + [markdown] id="cvHMHPNpmWQx"
# ### RelationNet (`relationnet`)
# + colab={"height": 381} executionInfo={"elapsed": 162, "status": "ok", "timestamp": 1616167475766, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="_Rg78TVbrgVH" outputId="c80011bb-9db1-4478-c2a7-715e81c147de"
relationnet_imagenet_df = pd.DataFrame(
columns=['mean (%)', '95% CI', '# episodes'],
index=datasets
)
relationnet_imagenet_df['# episodes'] = 600
relationnet_imagenet_df.loc[datasets[1:], ['mean (%)', '95% CI']] = [
[34.69, 1.01],
[45.35, 1.36],
[40.73, 0.83],
[49.51, 1.05],
[52.97, 0.69],
[43.30, 1.08],
[30.55, 1.04],
[68.76, 0.83],
[33.67, 1.05],
[29.15, 1.01]
]
relationnet_imagenet_df
# + colab={"height": 381} executionInfo={"elapsed": 192, "status": "ok", "timestamp": 1616167476105, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="UHo_yRvIriQJ" outputId="23882b9f-2947-4345-eb44-64a99032c5cd"
relationnet_all_df = pd.DataFrame(
columns=['mean (%)', '95% CI', '# episodes'],
index=datasets
)
relationnet_all_df['# episodes'] = 600
relationnet_all_df.loc[datasets[1:], ['mean (%)', '95% CI']] = [
[30.89, 0.93],
[86.57, 0.79],
[69.71, 0.83],
[54.14, 0.99],
[56.56, 0.73],
[61.75, 0.97],
[32.56, 1.08],
[76.08, 0.76],
[37.48, 0.93],
[27.41, 0.89]
]
relationnet_all_df
# + [markdown] id="uHUCxdbP2KRk"
# ### fo-Proto-MAML (`maml_init_with_proto`)
# + colab={"height": 381} executionInfo={"elapsed": 264, "status": "ok", "timestamp": 1616167476538, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="AsqraIRJ1ZmF" outputId="6c0b19f4-3609-4828-b939-17f00f365a82"
protomaml_imagenet_df = pd.DataFrame(
columns=['mean (%)', '95% CI', '# episodes'],
index=datasets
)
protomaml_imagenet_df['# episodes'] = 600
protomaml_imagenet_df.loc[datasets[1:], ['mean (%)', '95% CI']] = [
[49.53, 1.05],
[63.37, 1.33],
[55.95, 0.99],
[68.66, 0.96],
[66.49, 0.83],
[51.52, 1.00],
[39.96, 1.14],
[87.15, 0.69],
[48.83, 1.09],
[43.74, 1.12],
]
protomaml_imagenet_df
# + colab={"height": 381} executionInfo={"elapsed": 166, "status": "ok", "timestamp": 1616167476819, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="EQ4Ginc41xSj" outputId="659b17f5-0b73-4f5e-f376-c8dbc26bdb24"
protomaml_all_df = pd.DataFrame(
columns=['mean (%)', '95% CI', '# episodes'],
index=datasets
)
protomaml_all_df['# episodes'] = 600
protomaml_all_df.loc[datasets[1:], ['mean (%)', '95% CI']] = [
[46.52, 1.05],
[82.69, 0.97],
[75.23, 0.76],
[69.88, 1.02],
[68.25, 0.81],
[66.84, 0.94],
[41.99, 1.17],
[88.72, 0.67],
[52.42, 1.08],
[41.74, 1.13]
]
protomaml_all_df
# + [markdown] id="cZM7wrVvuZBn"
# ## Results from Requeima et al. (2019)
# + executionInfo={"elapsed": 123, "status": "ok", "timestamp": 1616167477079, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="tsrQ52955gvn"
ref = ("Requeima et al. (2019)",
"<NAME>, <NAME>, <NAME>, <NAME>, "
"<NAME>; "
"[_Fast and Flexible Multi-Task Classification Using Conditional Neural "
"Adaptive Processes_](https://arxiv.org/abs/1906.07697); "
"NeurIPS 2019.")
references.append(ref)
# display.display(display.Markdown(ref[1]))
# + [markdown] id="pVAsKCYQvmZF"
# ### CNAPs (`cnaps`)
# + colab={"height": 381} executionInfo={"elapsed": 217, "status": "ok", "timestamp": 1616167477327, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="E_TQ8a8xuYXl" outputId="ee2e441b-1365-4079-d6a4-56af6c610d04"
cnaps_all_df = pd.DataFrame(
columns=['mean (%)', '95% CI', '# episodes'],
index=datasets
)
cnaps_all_df['# episodes'] = 600
cnaps_all_df.loc[datasets[1:], ['mean (%)', '95% CI']] = [
[50.8, 1.1],
[91.7, 0.5],
[83.7, 0.6],
[73.6, 0.9],
[59.5, 0.7],
[74.7, 0.8],
[50.2, 1.1],
[88.9, 0.5],
[56.5, 1.1],
[39.4, 1.0]
]
cnaps_all_df
# + [markdown] id="ndtJdlVg7YXm"
# ## Results from Baik et al. (2020)
# + executionInfo={"elapsed": 107, "status": "ok", "timestamp": 1616167477650, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="7QbfK6gO6F-z"
ref = ("Baik et al. (2020)",
"<NAME>, <NAME>, <NAME>, <NAME>, <NAME>; "
"[_Meta-Learning with Adaptive Hyperparameters_]"
"(https://papers.nips.cc/paper/2020/hash/ee89223a2b625b5152132ed77abbcc79-Abstract.html); "
"NeurIPS 2020.")
references.append(ref)
# display.display(display.Markdown(ref[1]))
# + [markdown] id="FZNhlOGJ8GPJ"
# ### ALFA + fo-Proto-MAML (`alfa_protomaml`)
# + colab={"height": 381} executionInfo={"elapsed": 316, "status": "ok", "timestamp": 1616167477976, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="KzyAsGiB7Xm3" outputId="9a4309c4-5e2d-4140-ccc7-379a93c6af80"
alfa_protomaml_imagenet_df = pd.DataFrame(
columns=['mean (%)', '95% CI', '# episodes'],
index=datasets
)
alfa_protomaml_imagenet_df['# episodes'] = 600
alfa_protomaml_imagenet_df.loc[datasets[1:], ['mean (%)', '95% CI']] = [
[52.80, 1.11],
[61.87, 1.51],
[63.43, 1.10],
[69.75, 1.05],
[70.78, 0.88],
[59.17, 1.16],
[41.49, 1.17],
[85.96, 0.77],
[60.78, 1.29],
[48.11, 1.14]
]
alfa_protomaml_imagenet_df
# + [markdown] id="xMXch9G3Cxf4"
# ### ALFA + fo-MAML (`alfa_maml`)
# Not included in the global table as it performs worse than ALFA + fo-Proto-MAML overall, but provided here for reference.
# + colab={"height": 381} executionInfo={"elapsed": 169, "status": "ok", "timestamp": 1616167478304, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="nPlXwPMkCwt9" outputId="78641b40-903c-4cf8-812c-8c834a9564bb"
alfa_maml_imagenet_df = pd.DataFrame(
columns=['mean (%)', '95% CI', '# episodes'],
index=datasets
)
alfa_maml_imagenet_df['# episodes'] = 600
alfa_maml_imagenet_df.loc[datasets[1:], ['mean (%)', '95% CI']] = [
[51.09, 1.17],
[67.89, 1.43],
[66.34, 1.17],
[67.67, 1.06],
[65.34, 0.95],
[60.53, 1.13],
[37.41, 1.00],
[84.28, 0.97],
[60.86, 1.43],
[40.05, 1.14]
]
alfa_maml_imagenet_df
# + [markdown] id="zOfPE8JAG9N9"
# ## Results from Doersch et al. (2020)
# <NAME>, <NAME>, <NAME>,
# _CrossTransformers: spatially-aware few-shot transfer_,
# NeurIPS 2020
# + executionInfo={"elapsed": 114, "status": "ok", "timestamp": 1616167478527, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="nAOiHySt615d"
ref = ("Doersch et al. (2020)",
"<NAME>, <NAME>, <NAME>; "
"[_CrossTransformers: spatially-aware few-shot transfer_]"
"(https://arxiv.org/abs/2007.11498); "
"NeurIPS 2020.")
references.append(ref)
# display.display(display.Markdown(ref[1]))
# + [markdown] id="gGepAWQ1NEOI"
# ### ProtoNet large (`protonet_large`)
# Larger-scale prototypical networks, including:
# - 224x224 input size
# - ResNet-34 backbone
# - SimCLR Episodes
# + colab={"height": 381} executionInfo={"elapsed": 230, "status": "ok", "timestamp": 1616167478790, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="XRZXBgmdPuGM" outputId="494e9828-9fa8-4a24-d0f3-fddab660571b"
protonet_large_imagenet_df = pd.DataFrame(
columns=['mean (%)', '95% CI', '# episodes'],
index=datasets
)
protonet_large_imagenet_df['# episodes'] = 600
protonet_large_imagenet_df.loc[datasets[1:], ['mean (%)', '95% CI']] = [
[53.69, 1.07],
[68.50, 1.27],
[58.04, 0.96],
[74.07, 0.92],
[68.76, 0.77],
[53.30, 1.06],
[40.73, 1.15],
[86.96, 0.73],
[58.11, 1.05],
[41.70, 1.08],
]
protonet_large_imagenet_df
# + [markdown] id="TgRONFqnRSCj"
# ### CrossTransformers (`ctx`)
#
# CrossTransformers network with:
# - 224x224 input size
# - ResNet-34 backbone
# - SimCLR episodes
# - 14x14 feature grid
# - BOHB-inspired data augmentation
# + colab={"height": 381} executionInfo={"elapsed": 182, "status": "ok", "timestamp": 1616167479104, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="3gaPGfZpXmcp" outputId="27672efc-9dfe-43aa-efee-30e83353720b"
ctx_imagenet_df = pd.DataFrame(
columns=['mean (%)', '95% CI', '# episodes'],
index=datasets
)
ctx_imagenet_df['# episodes'] = 600
ctx_imagenet_df.loc[datasets[1:], ['mean (%)', '95% CI']] = [
[62.76, 0.99],
[82.21, 1.00],
[79.49, 0.89],
[80.63, 0.88],
[75.57, 0.64],
[72.68, 0.82],
[51.58, 1.11],
[95.34, 0.37],
[82.65, 0.76],
[59.90, 1.02],
]
ctx_imagenet_df
# + [markdown] id="QHxQ9x9QxbCn"
# ## Results from Saikia et al. (2020)
# <NAME>, <NAME>, <NAME>, _Optimized Generic Feature Learning for Few-shot Classification across Domains_, arXiv 2020
# + executionInfo={"elapsed": 120, "status": "ok", "timestamp": 1616167479321, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="8qmBpCR57UrS"
ref = ("Saikia et al. (2020)",
"Tonmo<NAME>, <NAME>, <NAME>; "
"[_Optimized Generic Feature Learning for Few-shot Classification "
"across Domains_]"
"(https://arxiv.org/abs/2001.07926); "
"arXiv 2020.")
references.append(ref)
# display.display(display.Markdown(ref[1]))
# + [markdown] id="IjqU7WWIxtg0"
# ### BOHB (`bohb`)
# Validated on _S1_ (ImageNet) only, nearest-centroid classifier (NC).
# + colab={"height": 381} executionInfo={"elapsed": 182, "status": "ok", "timestamp": 1616167479522, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="jz3QtXuwxyev" outputId="50ae1135-862e-4983-9290-e0b41a0a885b"
bohb_imagenet_df = pd.DataFrame(
columns=['mean (%)', '95% CI', '# episodes'],
index=datasets
)
bohb_imagenet_df['# episodes'] = 600
bohb_imagenet_df.loc[datasets[1:], ['mean (%)', '95% CI']] = [
[51.92, 1.05],
[67.57, 1.21],
[54.12, 0.90],
[70.69, 0.90],
[68.34, 0.76],
[50.33, 1.04],
[41.38, 1.12],
[87.34, 0.59],
[51.80, 1.04],
[48.03, 0.99],
]
bohb_imagenet_df
# + [markdown] id="2gRbl98l2dTw"
# ## Results from Dvornik et al. (2020)
#
# + executionInfo={"elapsed": 166, "status": "ok", "timestamp": 1616167479860, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="3KJN0qML2qhJ"
ref = ("Dvornik et al. (2020)",
"<NAME>, <NAME>, <NAME>; "
"[_Selecting Relevant Features from a Multi-domain Representation for "
"Few-shot Classification_](https://arxiv.org/abs/2003.09338); "
"ECCV 2020.")
references.append(ref)
# display.display(display.Markdown(ref[1]))
# + [markdown] id="ozJPWXZp3IsG"
# ### SUR (`sur`)
# + colab={"height": 381} executionInfo={"elapsed": 207, "status": "ok", "timestamp": 1616167480096, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="W6PG4luB3IKS" outputId="50028bd1-1fdd-426b-b514-4862cabc4004"
sur_all_df = pd.DataFrame(
columns=['mean (%)', '95% CI', '# episodes'],
index=datasets
)
sur_all_df['# episodes'] = 600
sur_all_df.loc[datasets[1:], ['mean (%)', '95% CI']] = [
[56.1, 1.1],
[93.1, 0.5],
[84.6, 0.7],
[70.6, 1.0],
[71.0, 0.8],
[81.3, 0.6],
[64.2, 1.1],
[82.8, 0.8],
[53.4, 1.0],
[50.1, 1.0],
]
sur_all_df
# + [markdown] id="B3mfXh9C5rFM"
# ### SUR-pnf (`sur_pnf`)
# SUR with parametric network family, also referred as "SUR-pf".
# + colab={"height": 381} executionInfo={"elapsed": 158, "status": "ok", "timestamp": 1616167480389, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="P_ud7YYt5nsc" outputId="c8fe2b7c-6167-4bae-d654-3d9d0d5bf00b"
sur_pnf_all_df = pd.DataFrame(
columns=['mean (%)', '95% CI', '# episodes'],
index=datasets
)
sur_pnf_all_df['# episodes'] = 600
sur_pnf_all_df.loc[datasets[1:], ['mean (%)', '95% CI']] = [
[56.0, 1.1],
[90.0, 0.6],
[79.7, 0.8],
[75.9, 0.9],
[72.5, 0.7],
[76.7, 0.7],
[49.8, 1.1],
[90.0, 0.6],
[52.2, 0.8],
[50.2, 1.1],
]
sur_pnf_all_df
# + [markdown] id="fJFVMyQf0IZg"
# ## Results from Liu et al. (2021)
#
# <NAME>, <NAME>, <NAME>, <NAME>, <NAME>,
# _A Universal Representation Transformer Layer for Few-Shot Image Classification_, ICLR 2021
#
#
# + executionInfo={"elapsed": 170, "status": "ok", "timestamp": 1616167480673, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="xHrrosbq0idZ"
ref = ("Liu et al. (2021)",
"<NAME>, <NAME>, <NAME>, <NAME>, <NAME>; "
"[_Universal Representation Transformer Layer for Few-Shot Image "
"Classification_](https://arxiv.org/abs/2006.11702); "
"ICLR 2021.")
references.append(ref)
# display.display(display.Markdown(ref[1]))
# + [markdown] id="8An2wNp52o5C"
# ### URT (`urt`)
# + colab={"height": 381} executionInfo={"elapsed": 176, "status": "ok", "timestamp": 1616167480863, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="2hDG7v9A2hOQ" outputId="e1443d90-f2c8-4b0f-9389-b902dcfb53c0"
urt_all_df = pd.DataFrame(
columns=['mean (%)', '95% CI', '# episodes'],
index=datasets
)
urt_all_df['# episodes'] = 600
urt_all_df.loc[datasets[1:], ['mean (%)', '95% CI']] = [
[55.7, 1.0],
[94.4, 0.4],
[85.8, 0.6],
[76.3, 0.8],
[71.8, 0.7],
[82.5, 0.6],
[63.5, 1.0],
[88.2, 0.6],
[65.9, 1.3],
[52.2, 1.1],
]
urt_all_df
# + [markdown] id="wgWP-4Pa1fsW"
# ## Template to add a new paper
#
# ```
# ref = ("Author et al. (year)",
# "First Author, Second Author, Last Author; "
# "[_Title of Paper_](https://paper.url/); "
# "Venue year.")
# references.append(ref)
# # display.display(display.Markdown(ref[1]))
# ```
# + [markdown] id="zgukmFo7zoO3"
# ### Template to add a new model
#
# ```
# <model_name>_<train_source>_df = pd.DataFrame(
# columns=['mean (%)', '95% CI', '# episodes'],
# index=datasets
# )
# <model_name>_<train_source>_df['# episodes'] = ...
# <model_name>_<train_source>_df.loc[datasets[1:], ['mean (%)', '95% CI']] = [...]
# ```
# + [markdown] id="12ACpi1B3P_4"
# ## Aggregate in table
# + executionInfo={"elapsed": 134, "status": "ok", "timestamp": 1616167481109, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="bU1CMkCk3nP6"
imagenet_dfs = {
'k-NN': baseline_imagenet_df,
'Finetune': baselineft_imagenet_df,
'MatchingNet': matching_imagenet_df,
'ProtoNet': prototypical_imagenet_df,
'fo-MAML': maml_imagenet_df,
'RelationNet': relationnet_imagenet_df,
'fo-Proto-MAML': protomaml_imagenet_df,
'ALFA+fo-Proto-MAML': alfa_protomaml_imagenet_df,
'ProtoNet (large)': protonet_large_imagenet_df,
'CTX': ctx_imagenet_df,
'BOHB': bohb_imagenet_df,
}
# + colab={"height": 515} executionInfo={"elapsed": 234, "status": "ok", "timestamp": 1616167481367, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="j6qo-9t44hDf" outputId="dfa01188-18e7-4763-d8f6-4036b2a12336"
imagenet_df = pd.concat(
imagenet_dfs.values(),
axis=1,
keys=imagenet_dfs.keys())
imagenet_df
# + colab={"height": 515} executionInfo={"elapsed": 219, "status": "ok", "timestamp": 1616167481737, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="AHwV57i_4YDr" outputId="d1d76676-46e9-4f92-ae5e-27529635940d"
all_dfs = {
'k-NN': baseline_all_df,
'Finetune': baselineft_all_df,
'MatchingNet': matching_all_df,
'ProtoNet': prototypical_all_df,
'fo-MAML': maml_all_df,
'RelationNet': relationnet_all_df,
'fo-Proto-MAML': protomaml_all_df,
'CNAPs': cnaps_all_df,
'SUR': sur_all_df,
'SUR-pnf': sur_pnf_all_df,
'URT': urt_all_df,
}
all_df = pd.concat(
all_dfs.values(),
axis=1,
keys=all_dfs.keys())
all_df
# + colab={"height": 502} executionInfo={"elapsed": 167, "status": "ok", "timestamp": 1616167482049, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="Bs_5hsod7tcb" outputId="2cf83ec0-8e96-4552-a56b-dd2a092fbdd9"
models_df = pd.DataFrame.from_dict(
orient='index',
columns=["ref"],
data={
'k-NN': 'Triantafillou et al. (2020)',
'Finetune': 'Triantafillou et al. (2020)',
'MatchingNet': 'Triantafillou et al. (2020)',
'ProtoNet': 'Triantafillou et al. (2020)',
'fo-MAML': 'Triantafillou et al. (2020)',
'RelationNet': 'Triantafillou et al. (2020)',
'fo-Proto-MAML': 'Triantafillou et al. (2020)',
'CNAPs': 'Requeima et al. (2019)',
'ALFA+fo-Proto-MAML': 'Baik et al. (2020)',
'ProtoNet (large)': 'Doersch et al. (2020)',
'CTX': 'Doersch et al. (2020)',
'BOHB': 'Saikia et al. (2020)',
'SUR': 'Dvornik et al. (2020)',
'SUR-pnf': 'Dvornik et al. (2020)',
'URT': 'Liu et al. (2021)',
})
models_df
# + [markdown] id="CSOMsz4pPS2n"
# ### Add stddev
# + executionInfo={"elapsed": 165, "status": "ok", "timestamp": 1616167482352, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="E6sHKzIvPiYD"
def add_stddev(df):
# Extract original order of labels
datasets = df.index
models = df.columns.levels[0]
# Have only one result (mean, CI, ...) per row
stacked_df = df.stack(0)
# Add 'stddev' as column
stacked_df['stddev'] = stacked_df['95% CI'] * np.sqrt(stacked_df['# episodes']) / 1.96
# Reshape and put back in original order
new_df = stacked_df.unstack().swaplevel(0, 1, axis=1)
new_df = new_df.loc[datasets][models]
return new_df
# + colab={"height": 515} executionInfo={"elapsed": 285, "status": "ok", "timestamp": 1616167482655, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="DWgzJ8PRSxQ7" outputId="7a0ad97a-81a7-48dd-e80a-ed5773b42e05"
imagenet_df = add_stddev(imagenet_df)
imagenet_df
# + colab={"height": 515} executionInfo={"elapsed": 269, "status": "ok", "timestamp": 1616167483134, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="rOZxblRL42fi" outputId="456febc7-1e84-4d75-b0de-15322e8632f5"
all_df = add_stddev(all_df)
all_df
# + [markdown] id="ggXNVVn8zjhM"
# ### Add rankings
# + executionInfo={"elapsed": 152, "status": "ok", "timestamp": 1616167483390, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="u0k0DqdXt07o"
def is_difference_significant(best_stats, candidate_stats):
# compute a 95% confidence for the difference of means.
ci = 1.96 * np.sqrt((best_stats['stddev'] ** 2) / best_stats['# episodes'] +
(candidate_stats['stddev'] ** 2) / candidate_stats['# episodes'])
diff_of_means = best_stats['mean (%)'] - candidate_stats['mean (%)']
return np.abs(diff_of_means) > ci
# + executionInfo={"elapsed": 166, "status": "ok", "timestamp": 1616167483580, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="0vn3fGqHnzJe"
def compute_ranks(dataset_series):
dataset_df = dataset_series.unstack()
n_models = len(dataset_df.index)
remaining_models = list(dataset_df.index)
next_available_rank = 1
ranks = {}
# Iteratively pick the best models, then all the ones statistically equivalent
while remaining_models:
accuracies = dataset_df.loc[remaining_models]['mean (%)'].astype('d')
best_model = accuracies.idxmax(axis=1)
best_stats = dataset_df.loc[best_model]
tied_models = [best_model]
potential_tied_models = [model for model in remaining_models
if model != best_model]
for candidate in potential_tied_models:
candidate_stats = dataset_df.loc[candidate]
if not is_difference_significant(best_stats, candidate_stats):
tied_models.append(candidate)
n_ties = len(tied_models)
# All tied models share the same rank, which is the average of the next
# `n_ties` available ranks (the ranks they would have without the ties), or
# next_available_rank + (1 + ... + (n_ties - 1)) / n_ties, which gives:
shared_rank = next_available_rank + (n_ties - 1) / 2
next_available_rank += n_ties
for model in tied_models:
ranks[model] = shared_rank
# Remove picked models for next iteration
remaining_models = [model for model in remaining_models
if model not in tied_models]
return pd.Series(ranks, name='rank')
# + executionInfo={"elapsed": 180, "status": "ok", "timestamp": 1616167483770, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="0Ra3dxcIwYE4"
def add_ranks(df):
# Get ranks as a data frame (ignore "ILSVRC (valid)")
ranks = df[1:].apply(compute_ranks, axis=1)
# Set the columns as (model, 'rank') Multi-index
ranks = pd.concat([ranks], axis=1, keys=['rank']).swaplevel(0, 1, axis=1)
# Concatenate with the original dataframe and defrag columns
new_df = pd.concat([df, ranks], axis=1)[df.columns.levels[0]]
return new_df
# + colab={"height": 515} executionInfo={"elapsed": 521, "status": "ok", "timestamp": 1616167484305, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="w2mbMuBZxrE2" outputId="cdc061ea-5b7d-4303-84dd-24ded3b2a23d"
imagenet_df = add_ranks(imagenet_df)
imagenet_df
# + colab={"height": 515} executionInfo={"elapsed": 502, "status": "ok", "timestamp": 1616167484929, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="ZWkM2g3Ayf5o" outputId="734bcf75-54a0-47d9-edc5-9b74ce9c3507"
all_df = add_ranks(all_df)
all_df
# + executionInfo={"elapsed": 196, "status": "ok", "timestamp": 1616167485241, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="UD-EU6P_9Moc" outputId="4ec590cd-ed6c-4cfa-95fc-cf90f57b5b06"
imagenet_df.xs('rank', axis=1, level=1).mean()
# + executionInfo={"elapsed": 190, "status": "ok", "timestamp": 1616167485470, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="B7GabELTBq-o" outputId="0d7ef929-6f7e-4174-9ac7-4e5e196efff7"
all_df.xs('rank', axis=1, level=1).mean()
# + [markdown] id="tn0hBqntzpG9"
# ### Display in HTML
# This section uses the DataFrame's "styler" object, which renders nicely within the notebook.
#
# Unfortunately, the HTML it outputs is not compatible with GitHub's markdown (as it relies on the `<style>` tag).
# + executionInfo={"elapsed": 210, "status": "ok", "timestamp": 1616167485718, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="u0S8Uno9yem3"
def str_summary(series):
# Summarize each (episode, model) by a single cell
# Non-breaking space to keep things on the same line
nbsp = '\u00A0'
string = '%(acc)s±%(ci)s%(nbsp)s(%(rank)g)' % {
'acc': series['mean (%)'],
'ci': series['95% CI'],
'rank': series['rank'],
'nbsp': nbsp
}
return string
# + executionInfo={"elapsed": 170, "status": "ok", "timestamp": 1616167485900, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="_uFqczyza6XZ"
def display_table(df, models=None):
accuracies_df = df.stack(0).apply(str_summary, axis=1).unstack(0)[df.index[1:]]
rank_df = df.xs('rank', axis=1, level=1).loc[df.index[1:]]
avg_rank_df = pd.DataFrame(rank_df.mean(), columns=['Avg rank'])
display_df = pd.concat([avg_rank_df, accuracies_df], axis=1)
if models:
# Try and force a particular order of models
display_df = display_df.loc[models]
# Bold cells corresponding to the best rank
best_acc_mask = rank_df.T == rank_df.min(axis=1)
best_avg_mask = avg_rank_df == avg_rank_df.min()
best_mask = pd.concat([best_avg_mask, best_acc_mask], axis=1)
if models:
best_mask = best_mask.loc[models]
bold_mask = best_mask.applymap(lambda v: 'font-weight: bold' if v else '')
display_style = display_df.style.apply(lambda f: bold_mask, axis=None)
display_style = display_style.format({'Avg rank': '{:g}'})
return display_style
# + colab={"height": 468} executionInfo={"elapsed": 257, "status": "ok", "timestamp": 1616167486170, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="Ab_CB6wcgtJ9" outputId="6e51ca36-d24f-4c92-b138-d7dc63f35708"
imagenet_display = display_table(imagenet_df, models=imagenet_dfs.keys())
imagenet_display
# + executionInfo={"elapsed": 154, "status": "ok", "timestamp": 1616167486479, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="dMjhdJkiimQx"
# print(imagenet_display.render())
# + colab={"height": 434} executionInfo={"elapsed": 241, "status": "ok", "timestamp": 1616167486754, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="UvuDF4GbeZpQ" outputId="85a795d2-b686-4bdb-d791-3675c0f5b403"
all_display = display_table(all_df, models=all_dfs.keys())
all_display
# + executionInfo={"elapsed": 158, "status": "ok", "timestamp": 1616167487047, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="ASEngzGKifwU"
# print(all_display.render())
# + [markdown] id="nLxs4WA4GPDu"
# ### Display in MarkDown
# At least, in GitHub-flavored MarkDown.
# + executionInfo={"elapsed": 277, "status": "ok", "timestamp": 1616167487331, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="cB_SuLA4GNQ7"
def md_render(series):
# Summarize each (episode, model) by a single cell containing MarkDown
nbsp = ' '
md_string = '%(bold)s%(acc)5.2f%(bold)s±%(ci)4.2f%(nbsp)s(%(rank)g)' % {
'acc': series['mean (%)'],
'ci': series['95% CI'],
'rank': series['rank'],
'bold': '**' if series['best_rank'] else '',
'nbsp': nbsp
}
return md_string
# + executionInfo={"elapsed": 181, "status": "ok", "timestamp": 1616167487522, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="2xe8elpLLFkH"
def md_table(df, models=None):
# Whether a model has the best rank on a given dataset
rank_df = df.xs('rank', axis=1, level=1).loc[df.index[1:]]
best_rank = pd.concat([rank_df.T == rank_df.min(axis=1)], axis=1,
keys=['best_rank']).swaplevel(0, 1, axis=1)
accuracies_df = df[1:].T.unstack(1)
accuracies_df = pd.concat([accuracies_df, best_rank], axis=1)
accuracies_md = accuracies_df.stack(0).apply(md_render, axis=1).unstack(1)
# Average rank (and whether it's the best)
avg_rank_df = pd.DataFrame(rank_df.mean(), columns=['Avg rank'])
best_avg_rank = (avg_rank_df == avg_rank_df.min()).rename(
columns={'Avg rank': 'best_rank'})
avg_rank_md = pd.concat([avg_rank_df, best_avg_rank], axis=1).apply(
lambda s: '%(bold)s%(avg_rank)g%(bold)s' % {
'avg_rank': s['Avg rank'],
'bold': '**' if s['best_rank'] else ''
},
axis=1).rename('Avg rank')
# Display method name with a pointer to the reference, defined later.
ref_to_link = {ref[0]: "[[%i]]" % i for i, ref in enumerate(references, 1)}
method_md = models_df.apply(lambda r: ref_to_link[r['ref']], axis='columns')
display_md = pd.concat([avg_rank_md, accuracies_md[df.index[1:]]], axis=1)
if models:
# Try and force a particular order of models
display_md = display_md.loc[list(models)]
# Pad all cells so they align well, 27 chars should be enough
header_str = '|'.join(['%-27s' % c
for c in ['Method'] + list(display_md.columns)])
sep_str = '|'.join(['-' * 27 for _ in [''] + list(display_md.columns)])
rows = [
'|'.join(['%-27s' % c for c in ([' '.join((i, method_md.loc[i]))] +
list(display_md.loc[i]))])
for i in display_md.index
]
return '\n'.join([header_str, sep_str] + rows)
# + executionInfo={"elapsed": 177, "status": "ok", "timestamp": 1616167487714, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="l77g7JRGTY_a" outputId="e9b6abdb-7581-4872-c604-ea2094c86af4"
print(md_table(imagenet_df, models=imagenet_dfs.keys()))
# + executionInfo={"elapsed": 161, "status": "ok", "timestamp": 1616167487889, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="LvUgO5y5qZfx" outputId="bdc8a0d7-840e-4a1d-904b-1bb2fc08b578"
print(md_table(all_df, models=all_dfs.keys()))
# + [markdown] id="7vzf_wCyZSoB"
# ## Export to MarkDown
# + [markdown] id="fLCbben1USd0"
# ### Reference list
# + executionInfo={"elapsed": 109, "status": "ok", "timestamp": 1616167488067, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="dc5MOCw1n02N"
def sanitize_anchor(string):
# Try to mimic the MarkDown function that transforms a section title into an
# html link anchor, that is:
# - put it in lower case
# - remove everything that is not a text character ("\w", which includes "_"),
# a space ("\s") or dash ("-")
# - replace spaces and "_" by "-" (and deduplicate)
anchor = string.lower()
anchor = re.sub('[^\w\s-]', '', anchor)
anchor = re.sub('[\s_-]+', '-', anchor)
return anchor
# + executionInfo={"elapsed": 192, "status": "ok", "timestamp": 1616167488267, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="76-TdhVTZfdZ"
def ref_list():
# Define links from [i] to the reference section
links = []
for i, ref in enumerate(references, 1):
links.append('[%(i)i]: #%(i)i-%(r)s' % dict(
i=i,
r=sanitize_anchor(ref[0])))
references_md = []
# Content of the reference section
for i, ref in enumerate(references, 1):
references_md.append(textwrap.dedent(r'''
###### \[%(i)i\] %(shortref)s
%(fullref)s
''') % dict(i=i, shortref=ref[0], fullref=ref[1]))
return '\n'.join(links + references_md)
# + executionInfo={"elapsed": 115, "status": "ok", "timestamp": 1616167488397, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="GzAvwnl4j4iB" outputId="6313e0ba-b700-4419-c279-6a9df15f5624"
print(ref_list())
# + [markdown] id="y-lfGUt6Uc2F"
# ### Full section
# + executionInfo={"elapsed": 184, "status": "ok", "timestamp": 1616167488592, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="aOAZQlucj59E"
def export_md():
begin_line = '<!-- Beginning of content generated by `Leaderboard.ipynb` -->'
end_line = '<!-- End of content generated by `Leaderboard.ipynb` -->'
parts = [
begin_line,
'## Training on ImageNet only',
md_table(imagenet_df, models=imagenet_dfs.keys()),
'## Training on all datasets',
md_table(all_df, models=all_dfs.keys()),
'## References',
ref_list(),
end_line
]
return '\n\n'.join(parts)
# + executionInfo={"elapsed": 233, "status": "ok", "timestamp": 1616167488839, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="_tAgydyHX_R_" outputId="8ef2c2c5-3ad2-485a-985e-bb8507e6c941"
print(export_md())
# + executionInfo={"elapsed": 192, "status": "ok", "timestamp": 1616167489094, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="Rjrd9raOBqbf"
|
meta-dataset/Leaderboard.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] deletable=false editable=false nbgrader={"checksum": "3d7e0bf7bf37a14b2f4cc8896af5d808", "grade": false, "grade_id": "cell-c9904c1c46f57746", "locked": true, "schema_version": 1, "solution": false}
# # Assignment 1: Bandits and Exploration/Exploitation
# + [markdown] deletable=false editable=false nbgrader={"checksum": "c8f4f6a23a8695f62a4e02738e204550", "grade": false, "grade_id": "cell-6ef89310dd46c266", "locked": true, "schema_version": 1, "solution": false}
# Welcome to Assignment 1. This notebook will:
# - Help you create your first bandit algorithm
# - Help you understand the effect of epsilon on exploration and learn about the exploration/exploitation tradeoff
# - Introduce you to some of the reinforcement learning software we are going to use for this specialization
#
# This class uses RL-Glue to implement most of our experiments. It was originally designed by <NAME>, <NAME>, and <NAME>. This library will give you a solid framework to understand how reinforcement learning experiments work and how to run your own. If it feels a little confusing at first, don't worry - we are going to walk you through it slowly and introduce you to more and more parts as you progress through the specialization.
#
# We are assuming that you have used a Jupyter notebook before. But if not, it is quite simple. Simply press the run button, or shift+enter to run each of the cells. The places in the code that you need to fill in will be clearly marked for you.
# + [markdown] deletable=false editable=false nbgrader={"checksum": "770bc9fe226cb87b7ad8ca34de163650", "grade": false, "grade_id": "cell-180a16562fb72a2d", "locked": true, "schema_version": 1, "solution": false}
# ## Section 0: Preliminaries
# + deletable=false editable=false nbgrader={"checksum": "f301a0a9497887bcc0a8652dd598de26", "grade": false, "grade_id": "cell-b1f350f6be960eea", "locked": true, "schema_version": 1, "solution": false}
# Import necessary libraries
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
from rl_glue import RLGlue
import main_agent
import ten_arm_env
import test_env
from tqdm import tqdm
import time
# + [markdown] deletable=false editable=false nbgrader={"checksum": "392a2cda80f785dd798d7155ea7eb2b7", "grade": false, "grade_id": "cell-e2a306e4cfd3e433", "locked": true, "schema_version": 1, "solution": false}
# In the above cell, we import the libraries we need for this assignment. We use numpy throughout the course and occasionally provide hints for which methods to use in numpy. Other than that we mostly use vanilla python and the occasional other library, such as matplotlib for making plots.
#
# You might have noticed that we import ten_arm_env. This is the __10-armed Testbed__ introduced in [section 2.3](http://www.incompleteideas.net/book/RLbook2018.pdf) of the textbook. We use this throughout this notebook to test our bandit agents. It has 10 arms, which are the actions the agent can take. Pulling an arm generates a stochastic reward from a Gaussian distribution with unit-variance. For each action, the expected value of that action is randomly sampled from a normal distribution, at the start of each run. If you are unfamiliar with the 10-armed Testbed please review it in the textbook before continuing.
#
# DO NOT IMPORT OTHER LIBRARIES as this will break the autograder.
# + [markdown] deletable=false editable=false nbgrader={"checksum": "6dfc1a07738ba7ef428ad0d6045b194d", "grade": false, "grade_id": "cell-753cb03c956b611e", "locked": true, "schema_version": 1, "solution": false}
# ## Section 1: Greedy Agent
# + [markdown] deletable=false editable=false nbgrader={"checksum": "26fc8f97320909c8ac7e8c66f5b73fca", "grade": false, "grade_id": "cell-8e7576e85bbe82fc", "locked": true, "schema_version": 1, "solution": false}
# We want to create an agent that will find the action with the highest expected reward. One way an agent could operate is to always choose the action with the highest value based on the agent’s current estimates. This is called a greedy agent as it greedily chooses the action that it thinks has the highest value. Let's look at what happens in this case.
#
# First we are going to implement the argmax function, which takes in a list of action values and returns an action with the highest value. Why are we implementing our own instead of using the argmax function that numpy uses? Numpy's argmax function returns the first instance of the highest value. We do not want that to happen as it biases the agent to choose a specific action in the case of ties. Instead we want to break ties between the highest values randomly. So we are going to implement our own argmax function. You may want to look at [np.random.choice](https://docs.scipy.org/doc/numpy/reference/generated/numpy.random.choice.html) to randomly select from a list of values.
# + deletable=false nbgrader={"checksum": "7da87f1e2652ce6f916164cbc8527ef4", "grade": false, "grade_id": "cell-00a70af9534c45cb", "locked": false, "schema_version": 1, "solution": true}
# [Graded]
def argmax(q_values):
"""
Takes in a list of q_values and returns the index
of the item with the highest value. Breaks ties randomly.
returns: int - the index of the highest value in q_values
"""
top = float("-inf")
ties = []
for i in range(len(q_values)):
# if a value in q_values is greater than the highest value, then update top and reset ties to zero
# if a value is equal to top value, then add the index to ties (hint: do this no matter what)
# Note: You do not have to follow this exact solution. You can choose to do your own implementation.
### START CODE HERE ###
if top < q_values[i]:
top = q_values[i]
ties = []
ties.append(i)
elif top == q_values[i]:
ties.append(i)
### END CODE HERE ###
# return a random selection from ties. (hint: look at np.random.choice)
### START CODE HERE ###
action = np.random.choice(ties)
### END CODE HERE ###
return action # change this
# + deletable=false editable=false nbgrader={"checksum": "a59aa0c5fb1fb6ff3925280d345b6fb6", "grade": true, "grade_id": "cell-f227246db2235e96", "locked": true, "points": 0, "schema_version": 1, "solution": false}
# Test argmax implentation
test_array = [0, 0, 0, 0, 0, 0, 0, 0, 1, 0]
assert argmax(test_array) == 8, "Check your argmax implementation returns the index of the largest value"
test_array = [1, 0, 0, 1]
total = 0
for i in range(100):
total += argmax(test_array)
np.save("argmax_test", total)
assert total > 0, "Make sure your argmax implementation randomly choooses among the largest values."
assert total != 300, "Make sure your argmax implementation randomly choooses among the largest values."
# + [markdown] deletable=false editable=false nbgrader={"checksum": "60f1a63e2e8eadfa949c0c7e5641c6c3", "grade": false, "grade_id": "cell-80dca165281ba2f3", "locked": true, "schema_version": 1, "solution": false}
# Now we introduce the first part of an RL-Glue agent that you will implement. Here we are going to create a GreedyAgent and implement the agent_step method. This method gets called each time the agent takes a step. The method has to return the action selected by the agent. This method also ensures the agent’s estimates are updated based on the signals it gets from the environment.
#
# Fill in the code below to implement a greedy agent.
# + deletable=false nbgrader={"checksum": "b65c3822e9e411db5f265be3b2d4fb96", "grade": false, "grade_id": "cell-582d9e7f86d07eb6", "locked": false, "schema_version": 1, "solution": true}
# Greedy agent here [Graded]
class GreedyAgent(main_agent.Agent):
def agent_step(self, reward, observation):
"""
Takes one step for the agent. It takes in a reward and observation and
returns the action the agent chooses at that time step.
Arguments:
reward -- float, the reward the agent received from the environment after taking the last action.
observation -- float, the observed state the agent is in. Do not worry about this for this assignment
as you will not use it until future lessons.
Returns:
current_action -- int, the action chosen by the agent at the current time step.
"""
### Useful Class Variables ###
# self.q_values : An array with the agent’s value estimates for each action.
# self.arm_count : An array with a count of the number of times each arm has been pulled.
# self.last_action : The action that the agent took on the previous time step.
#######################
# Update action values. Hint: Look at the algorithm in section 2.4 of the textbook.
# Increment the counter in self.arm_count for the action from the previous time step
# Update the step size using self.arm_count
# Update self.q_values for the action from the previous time step
# (~3-5 lines)
### START CODE HERE ###
self.arm_count[self.last_action] += 1
step_size = 1 / (self.arm_count[self.last_action])
self.q_values[self.last_action] += step_size * (reward - self.q_values[self.last_action])
### END CODE HERE ###
# current action = ? # Use the argmax function you created above
# (~2 lines)
### START CODE HERE ###
current_action = argmax(self.q_values)
### END CODE HERE ###
self.last_action = current_action
return current_action
# + deletable=false editable=false nbgrader={"checksum": "6a40d37a378dc173aa2654f9fdb0ff7f", "grade": true, "grade_id": "cell-08fc9e17dec07fd5", "locked": true, "points": 0, "schema_version": 1, "solution": false}
# Do not modify this cell
# Test for Greedy Agent Code
greedy_agent = GreedyAgent()
greedy_agent.q_values = [0, 0, 1.0, 0, 0]
greedy_agent.arm_count = [0, 1, 0, 0, 0]
greedy_agent.last_action = 1
action = greedy_agent.agent_step(1, 0)
print(greedy_agent.q_values)
np.save("greedy_test", greedy_agent.q_values)
print("Output:")
print(greedy_agent.q_values)
print("Expected Output:")
print([0, 0.5, 1.0, 0, 0])
assert action == 2, "Check that you are using argmax to choose the action with the highest value."
assert greedy_agent.q_values == [0, 0.5, 1.0, 0, 0], "Check that you are updating q_values correctly."
# + [markdown] deletable=false editable=false nbgrader={"checksum": "90a42fd6968847f33177fb88b0154707", "grade": false, "grade_id": "cell-0edf7d5d440cdc40", "locked": true, "schema_version": 1, "solution": false}
# Let's visualize the result. Here we run an experiment using RL-Glue to test our agent. For now, we will set up the experiment code; in future lessons, we will walk you through running experiments so that you can create your own.
# + deletable=false editable=false nbgrader={"checksum": "c2bbd4838419e16d407d08f2a7c2fb66", "grade": false, "grade_id": "cell-13bf4a5ec5402a22", "locked": true, "schema_version": 1, "solution": false}
# Plot Greedy Result
num_runs = 200 # The number of times we run the experiment
num_steps = 1000 # The number of steps each experiment is run for
env = ten_arm_env.Environment # We the environment to use
agent = GreedyAgent # We choose what agent we want to use
agent_info = {"num_actions": 10} # Pass the agent the information it needs;
# here it just needs the number of actions (number of arms).
env_info = {} # Pass the environment the information it needs; in this case, it is nothing.
all_averages = []
for i in tqdm(range(num_runs)): # tqdm is what creates the progress bar below once the code is run
rl_glue = RLGlue(env, agent) # Creates a new RLGlue experiment with the env and agent we chose above
rl_glue.rl_init(agent_info, env_info) # Pass RLGlue what it needs to initialize the agent and environment
rl_glue.rl_start() # Start the experiment
scores = [0]
averages = []
for i in range(num_steps):
reward, _, action, _ = rl_glue.rl_step() # The environment and agent take a step and return
# the reward, and action taken.
scores.append(scores[-1] + reward)
averages.append(scores[-1] / (i + 1))
all_averages.append(averages)
plt.figure(figsize=(15, 5), dpi= 80, facecolor='w', edgecolor='k')
plt.plot([1.55 for _ in range(num_steps)], linestyle="--")
plt.plot(np.mean(all_averages, axis=0))
plt.legend(["Best Possible", "Greedy"])
plt.title("Average Reward of Greedy Agent")
plt.xlabel("Steps")
plt.ylabel("Average reward")
plt.show()
greedy_scores = np.mean(all_averages, axis=0)
np.save("greedy_scores", greedy_scores)
# + [markdown] deletable=false editable=false nbgrader={"checksum": "a853027b4891a856372c381b05461a4f", "grade": false, "grade_id": "cell-04a8bd103b7af798", "locked": true, "schema_version": 1, "solution": false}
# How did our agent do? Is it possible for it to do better?
# -
# ## Section 2: Epsilon-Greedy Agent
# We learned about [another way for an agent to operate](https://www.coursera.org/learn/fundamentals-of-reinforcement-learning/lecture/tHDck/what-is-the-trade-off), where it does not always take the greedy action. Instead, sometimes it takes an exploratory action. It does this so that it can find out what the best action really is. If we always choose what we think is the current best action is, we may miss out on taking the true best action, because we haven't explored enough times to find that best action.
#
# Implement an epsilon-greedy agent below. Hint: we are implementing the algorithm from [section 2.4](http://www.incompleteideas.net/book/RLbook2018.pdf#page=52) of the textbook. You may want to use your greedy code from above and look at [np.random.random](https://docs.scipy.org/doc/numpy/reference/generated/numpy.random.random.html), as well as [np.random.randint](https://docs.scipy.org/doc/numpy/reference/generated/numpy.random.randint.html), to help you select random actions.
# + deletable=false nbgrader={"checksum": "00712574db774e4553878a18665e2751", "grade": false, "grade_id": "cell-6862cb5ef5702d22", "locked": false, "schema_version": 1, "solution": true}
# Epsilon Greedy Agent here [Graded]
class EpsilonGreedyAgent(main_agent.Agent):
def agent_step(self, reward, observation):
"""
Takes one step for the agent. It takes in a reward and observation and
returns the action the agent chooses at that time step.
Arguments:
reward -- float, the reward the agent received from the environment after taking the last action.
observation -- float, the observed state the agent is in. Do not worry about this for this assignment
as you will not use it until future lessons.
Returns:
current_action -- int, the action chosen by the agent at the current time step.
"""
### Useful Class Variables ###
# self.q_values : An array with the agent’s value estimates for each action.
# self.arm_count : An array with a count of the number of times each arm has been pulled.
# self.last_action : The action that the agent took on the previous time step.
# self.epsilon : The probability an epsilon greedy agent will explore (ranges between 0 and 1)
#######################
# Update action-values - this should be the same update as your greedy agent above
# (~3-5 lines)
### START CODE HERE ###
self.arm_count[self.last_action] += 1
step_size = 1 / (self.arm_count[self.last_action])
self.q_values[self.last_action] += step_size * (reward - self.q_values[self.last_action])
### END CODE HERE ###
# Choose action using epsilon greedy
# Randomly choose a number between 0 and 1 and see if it is less than self.epsilon
# (Hint: look at np.random.random()). If it is, set current_action to a random action.
# Otherwise choose current_action greedily as you did above.
# (~4 lines)
### START CODE HERE ###
prob = np.random.random()
if prob > self.epsilon:
current_action = argmax(self.q_values)
else:
current_action = np.random.choice(len(self.q_values))
### END CODE HERE ###
self.last_action = current_action
return current_action
# + deletable=false editable=false nbgrader={"checksum": "e530a16b139b2a9966b1a65e25086f70", "grade": true, "grade_id": "cell-3099aff70dfd2e61", "locked": true, "points": 0, "schema_version": 1, "solution": false}
# Do not modify this cell
# Test Code for Epsilon Greedy Agent
e_greedy_agent = EpsilonGreedyAgent()
e_greedy_agent.q_values = [0, 0, 1.0, 0, 0]
e_greedy_agent.arm_count = [0, 1, 0, 0, 0]
e_greedy_agent.num_actions = 5
e_greedy_agent.last_action = 1
e_greedy_agent.epsilon = 0.5
action = e_greedy_agent.agent_step(1, 0)
print("Output:")
print(e_greedy_agent.q_values)
print("Expected Output:")
print([0, 0.5, 1.0, 0, 0])
# assert action == 2, "Check that you are using argmax to choose the action with the highest value."
assert e_greedy_agent.q_values == [0, 0.5, 1.0, 0, 0], "Check that you are updating q_values correctly."
# + [markdown] deletable=false editable=false nbgrader={"checksum": "5488f20b68110a856dad3a003f51db32", "grade": false, "grade_id": "cell-762b0b3997c2300f", "locked": true, "schema_version": 1, "solution": false}
# Now that we have our epsilon greedy agent created. Let's compare it against the greedy agent with epsilon of 0.1.
# + deletable=false editable=false nbgrader={"checksum": "9a2b47a106185d21f2e00f81871f7476", "grade": false, "grade_id": "cell-2f6cef9d3ecdace7", "locked": true, "schema_version": 1, "solution": false}
# Plot Epsilon greedy results and greedy results
num_runs = 200
num_steps = 1000
epsilon = 0.1
agent = EpsilonGreedyAgent
env = ten_arm_env.Environment
agent_info = {"num_actions": 10, "epsilon": epsilon}
env_info = {}
all_averages = []
for i in tqdm(range(num_runs)):
rl_glue = RLGlue(env, agent)
rl_glue.rl_init(agent_info, env_info)
rl_glue.rl_start()
scores = [0]
averages = []
for i in range(num_steps):
reward, _, action, _ = rl_glue.rl_step() # The environment and agent take a step and return
# the reward, and action taken.
scores.append(scores[-1] + reward)
averages.append(scores[-1] / (i + 1))
all_averages.append(averages)
plt.figure(figsize=(15, 5), dpi= 80, facecolor='w', edgecolor='k')
plt.plot([1.55 for _ in range(num_steps)], linestyle="--")
plt.plot(greedy_scores)
plt.title("Average Reward of Greedy Agent vs. Epsilon-Greedy Agent")
plt.plot(np.mean(all_averages, axis=0))
plt.legend(("Best Possible", "Greedy", "Epsilon Greedy: Epsilon = 0.1"))
plt.xlabel("Steps")
plt.ylabel("Average reward")
plt.show()
np.save("e-greedy", all_averages)
# + [markdown] deletable=false editable=false nbgrader={"checksum": "ed0fa5039cf69237a1caf29b273b2942", "grade": false, "grade_id": "cell-23cf04f952075345", "locked": true, "schema_version": 1, "solution": false}
# Notice how much better the epsilon-greedy agent did. Because we occasionally choose a random action we were able to find a better long term policy. By acting greedily before our value estimates are accurate, we risk settling on a suboptimal action.
# + [markdown] deletable=false editable=false nbgrader={"checksum": "7acf4c8b67b66a59bd737bbb29d3d9f7", "grade": false, "grade_id": "cell-edb9184608392c62", "locked": true, "schema_version": 1, "solution": false}
# ## 1.2 Averaging Multiple Runs
# + [markdown] deletable=false editable=false nbgrader={"checksum": "7c51be606d9d6554fdd916078b0bda57", "grade": false, "grade_id": "cell-1b55f263f08b1389", "locked": true, "schema_version": 1, "solution": false}
# Did you notice that we averaged over 2000 runs? Why did we do that?
#
# To get some insight, let's look at the results of two individual runs by the same agent.
# + deletable=false editable=false nbgrader={"checksum": "6a98c68aa5e4e1270799ea04bed4721f", "grade": false, "grade_id": "cell-69d62e83fc1d91bc", "locked": true, "schema_version": 1, "solution": false}
# Plot runs of e-greedy agent
agent = EpsilonGreedyAgent
agent_info = {"num_actions": 10, "epsilon": 0.1}
env_info = {}
all_averages = []
plt.figure(figsize=(15, 5), dpi= 80, facecolor='w', edgecolor='k')
num_steps = 1000
for run in (0, 1):
np.random.seed(run) # Here we set the seed so that we can compare two different runs
averages = []
rl_glue = RLGlue(env, agent)
rl_glue.rl_init(agent_info, env_info)
rl_glue.rl_start()
scores = [0]
for i in range(num_steps):
reward, state, action, is_terminal = rl_glue.rl_step()
scores.append(scores[-1] + reward)
averages.append(scores[-1] / (i + 1))
# all_averages.append(averages)
plt.plot(averages)
# plt.plot(greedy_scores)
plt.title("Comparing two runs of the same agent")
plt.xlabel("Steps")
plt.ylabel("Average reward")
# plt.plot(np.mean(all_averages, axis=0))
# plt.legend(("Greedy", "Epsilon: 0.1"))
plt.show()
# + [markdown] deletable=false editable=false nbgrader={"checksum": "9c6b5d4ea841a388245eb1fdc732a3ed", "grade": false, "grade_id": "cell-cbabc6468847faab", "locked": true, "schema_version": 1, "solution": false}
# Notice how the two runs were different? But, if this is the exact same algorithm, why does it behave differently in these two runs?
#
# The answer is that it is due to randomness in the environment and in the agent. Depending on what action the agent randomly starts with, or when it randomly chooses to explore, it can change the results of the runs. And even if the agent chooses the same action, the reward from the environment is randomly sampled from a Gaussian. The agent could get lucky, and see larger rewards for the best action early on and so settle on the best action faster. Or, it could get unlucky and see smaller rewards for best action early on and so take longer to recognize that it is in fact the best action.
#
# To be more concrete, let’s look at how many times an exploratory action is taken, for different seeds.
# + deletable=false editable=false nbgrader={"checksum": "9c0105d5ee8f26ec966704dad4d013f0", "grade": false, "grade_id": "cell-a6e9ef699d799240", "locked": true, "schema_version": 1, "solution": false}
print("Random Seed 1")
np.random.seed(1)
for _ in range(15):
if np.random.random() < 0.1:
print("Exploratory Action")
print()
print()
print("Random Seed 2")
np.random.seed(2)
for _ in range(15):
if np.random.random() < 0.1:
print("Exploratory Action")
# + [markdown] deletable=false editable=false nbgrader={"checksum": "bc8ff22ac82750f9eb3e0f901d5f4166", "grade": false, "grade_id": "cell-42f5c9cb11fffbb0", "locked": true, "schema_version": 1, "solution": false}
# With the first seed, we take an exploratory action three times out of 15, but with the second, we only take an exploratory action once. This can significantly affect the performance of our agent because the amount of exploration has changed significantly.
#
# To compare algorithms, we therefore report performance averaged across many runs. We do this to ensure that we are not simply reporting a result that is due to stochasticity, as explained [in the lectures](https://www.coursera.org/learn/fundamentals-of-reinforcement-learning/lecture/PtVBs/sequential-decision-making-with-evaluative-feedback). Rather, we want statistically significant outcomes. We will not use statistical significance tests in this course. Instead, because we have access to simulators for our experiments, we use the simpler strategy of running for a large number of runs and ensuring that the confidence intervals do not overlap.
# + [markdown] deletable=false editable=false nbgrader={"checksum": "65cc408096713cec77d263be0fd90b0d", "grade": false, "grade_id": "cell-1d4132f4b28f4881", "locked": true, "schema_version": 1, "solution": false}
# ## Section 3: Comparing values of epsilon
# + [markdown] deletable=false editable=false nbgrader={"checksum": "81b41ca2616b4d370e19c911cf4ab88e", "grade": false, "grade_id": "cell-f62fa977aac5da68", "locked": true, "schema_version": 1, "solution": false}
# Can we do better than an epsilon of 0.1? Let's try several different values for epsilon and see how they perform. We try different settings of key performance parameters to understand how the agent might perform under different conditions.
#
# Below we run an experiment where we sweep over different values for epsilon:
# + deletable=false editable=false nbgrader={"checksum": "2d9ab9563f8699e0b8aabcb1087f454b", "grade": false, "grade_id": "cell-4c9881740ba46656", "locked": true, "schema_version": 1, "solution": false}
# Experiment code for epsilon-greedy with different values of epsilon
epsilons = [0.0, 0.01, 0.1, 0.4]
plt.figure(figsize=(15, 5), dpi= 80, facecolor='w', edgecolor='k')
plt.plot([1.55 for _ in range(num_steps)], linestyle="--")
n_q_values = []
n_averages = []
n_best_actions = []
num_runs = 200
for epsilon in epsilons:
all_averages = []
for run in tqdm(range(num_runs)):
agent = EpsilonGreedyAgent
agent_info = {"num_actions": 10, "epsilon": epsilon}
env_info = {"random_seed": run}
rl_glue = RLGlue(env, agent)
rl_glue.rl_init(agent_info, env_info)
rl_glue.rl_start()
best_arm = np.argmax(rl_glue.environment.arms)
scores = [0]
averages = []
best_action_chosen = []
for i in range(num_steps):
reward, state, action, is_terminal = rl_glue.rl_step()
scores.append(scores[-1] + reward)
averages.append(scores[-1] / (i + 1))
if action == best_arm:
best_action_chosen.append(1)
else:
best_action_chosen.append(0)
if epsilon == 0.1 and run == 0:
n_q_values.append(np.copy(rl_glue.agent.q_values))
if epsilon == 0.1:
n_averages.append(averages)
n_best_actions.append(best_action_chosen)
all_averages.append(averages)
plt.plot(np.mean(all_averages, axis=0))
plt.legend(["Best Possible"] + epsilons)
plt.xlabel("Steps")
plt.ylabel("Average reward")
plt.show()
# + [markdown] deletable=false editable=false nbgrader={"checksum": "621e4edf3ee0456e562f8f61899fafd8", "grade": false, "grade_id": "cell-1763c2a2a2863158", "locked": true, "schema_version": 1, "solution": false}
# Why did 0.1 perform better than 0.01?
#
# If exploration helps why did 0.4 perform worse that 0.0 (the greedy agent)?
#
# Think about these and how you would answer these questions. They are questions in the practice quiz. If you still have questions about it, retake the practice quiz.
# + [markdown] deletable=false editable=false nbgrader={"checksum": "4107b76e0b504556e7760f38c7c603b2", "grade": false, "grade_id": "cell-7f65b4e031a22732", "locked": true, "schema_version": 1, "solution": false}
# ## Section 4: The Effect of Step Size
# + [markdown] deletable=false editable=false nbgrader={"checksum": "dacfdaab4566f744379cf2b63aa38125", "grade": false, "grade_id": "cell-a12e885539decec6", "locked": true, "schema_version": 1, "solution": false}
# In Section 1 of this assignment, we decayed the step size over time based on action-selection counts. The step-size was 1/N(A), where N(A) is the number of times action A was selected. This is the same as computing a sample average. We could also set the step size to be a constant value, such as 0.1. What would be the effect of doing that? And is it better to use a constant or the sample average method?
#
# To investigate this question, let’s start by creating a new agent that has a constant step size. This will be nearly identical to the agent created above. You will use the same code to select the epsilon-greedy action. You will change the update to have a constant step size instead of using the 1/N(A) update.
# + deletable=false nbgrader={"checksum": "0c375e4a3d46bfed1a8403807bf447ab", "grade": false, "grade_id": "cell-fe26903228ef0c50", "locked": false, "schema_version": 1, "solution": true}
# Constant Step Size Agent Here [Graded]
# Greedy agent here
class EpsilonGreedyAgentConstantStepsize(main_agent.Agent):
def agent_step(self, reward, observation):
"""
Takes one step for the agent. It takes in a reward and observation and
returns the action the agent chooses at that time step.
Arguments:
reward -- float, the reward the agent received from the environment after taking the last action.
observation -- float, the observed state the agent is in. Do not worry about this for this assignment
as you will not use it until future lessons.
Returns:
current_action -- int, the action chosen by the agent at the current time step.
"""
### Useful Class Variables ###
# self.q_values : An array with the agent’s value estimates for each action.
# self.arm_count : An array with a count of the number of times each arm has been pulled.
# self.last_action : The action that the agent took on the previous time step.
# self.step_size : A float which is the current step size for the agent.
# self.epsilon : The probability an epsilon greedy agent will explore (ranges between 0 and 1)
#######################
# Update q_values for action taken at previous time step
# using self.step_size intead of using self.arm_count
# (~1-2 lines)
### START CODE HERE ###
self.arm_count[self.last_action] += 1
self.q_values[self.last_action] += self.step_size * (reward - self.q_values[self.last_action])
### END CODE HERE ###
# Choose action using epsilon greedy. This is the same as you implemented above.
# (~4 lines)
### START CODE HERE ###
prob = np.random.random()
if prob > self.epsilon:
current_action = argmax(self.q_values)
else:
current_action = np.random.choice(len(self.q_values))
### END CODE HERE ###
self.last_action = current_action
return current_action
# + deletable=false editable=false nbgrader={"checksum": "fb2ba0590f0266b9aac4956d8f3c5489", "grade": true, "grade_id": "cell-ba6bdf28928e3042", "locked": true, "points": 0, "schema_version": 1, "solution": false}
# Do not modify this cell
# Test Code for Epsilon Greedy with Different Constant Stepsizes
for step_size in [0.01, 0.1, 0.5, 1.0]:
e_greedy_agent = EpsilonGreedyAgentConstantStepsize()
e_greedy_agent.q_values = [0, 0, 1.0, 0, 0]
# e_greedy_agent.arm_count = [0, 1, 0, 0, 0]
e_greedy_agent.num_actions = 5
e_greedy_agent.last_action = 1
e_greedy_agent.epsilon = 0.0
e_greedy_agent.step_size = step_size
action = e_greedy_agent.agent_step(1, 0)
print("Output for step size: {}".format(step_size))
print(e_greedy_agent.q_values)
print("Expected Output:")
print([0, step_size, 1.0, 0, 0])
assert e_greedy_agent.q_values == [0, step_size, 1.0, 0, 0], "Check that you are updating q_values correctly using the stepsize."
# + deletable=false editable=false nbgrader={"checksum": "005b2c64dcbef3a81bee85d79af688a4", "grade": false, "grade_id": "cell-a5d327f4d52578e6", "locked": true, "schema_version": 1, "solution": false}
# Experiment code for different step sizes [graded]
step_sizes = [0.01, 0.1, 0.5, 1.0]
epsilon = 0.1
num_steps = 1000
num_runs = 200
fig, ax = plt.subplots(figsize=(15, 5), dpi= 80, facecolor='w', edgecolor='k')
q_values = {step_size: [] for step_size in step_sizes}
true_values = {step_size: None for step_size in step_sizes}
best_actions = {step_size: [] for step_size in step_sizes}
for step_size in step_sizes:
all_averages = []
for run in tqdm(range(num_runs)):
agent = EpsilonGreedyAgentConstantStepsize
agent_info = {"num_actions": 10, "epsilon": epsilon, "step_size": step_size, "initial_value": 0.0}
env_info = {"random_seed": run}
rl_glue = RLGlue(env, agent)
rl_glue.rl_init(agent_info, env_info)
rl_glue.rl_start()
best_arm = np.argmax(rl_glue.environment.arms)
scores = [0]
averages = []
if run == 0:
true_values[step_size] = np.copy(rl_glue.environment.arms)
best_action_chosen = []
for i in range(num_steps):
reward, state, action, is_terminal = rl_glue.rl_step()
scores.append(scores[-1] + reward)
averages.append(scores[-1] / (i + 1))
if action == best_arm:
best_action_chosen.append(1)
else:
best_action_chosen.append(0)
if run == 0:
q_values[step_size].append(np.copy(rl_glue.agent.q_values))
best_actions[step_size].append(best_action_chosen)
ax.plot(np.mean(best_actions[step_size], axis=0))
if step_size == 0.01:
np.save("step_size", best_actions[step_size])
ax.plot(np.mean(n_best_actions, axis=0))
fig.legend(step_sizes + ["1/N(A)"])
plt.title("% Best Action Taken")
plt.xlabel("Steps")
plt.ylabel("% Best Action Taken")
vals = ax.get_yticks()
ax.set_yticklabels(['{:,.2%}'.format(x) for x in vals])
plt.show()
# + [markdown] deletable=false editable=false nbgrader={"checksum": "4490c3113b9b460e79a0f92ae6fb2433", "grade": false, "grade_id": "cell-6704fdb6f4f612fb", "locked": true, "schema_version": 1, "solution": false}
# Notice first that we are now plotting the amount of time that the best action is taken rather than the average reward. To better understand the performance of an agent, it can be useful to measure specific behaviors, beyond just how much reward is accumulated. This measure indicates how close the agent’s behaviour is to optimal.
#
# It seems as though 1/N(A) performed better than the others, in that it reaches a solution where it takes the best action most frequently. Now why might this be? Why did a step size of 0.5 start out better but end up performing worse? Why did a step size of 0.01 perform so poorly?
#
# Let's dig into this further below. Let’s plot how well each agent tracks the true value, where each agent has a different step size method. You do not have to enter any code here, just follow along.
# + deletable=false editable=false nbgrader={"checksum": "6c6c60dcf228a25e321b78ea59e86c71", "grade": false, "grade_id": "cell-49e29a510956e277", "locked": true, "schema_version": 1, "solution": false}
# Plot various step sizes and estimates
largest = 0
num_steps = 1000
for step_size in step_sizes:
plt.figure(figsize=(15, 5), dpi= 80, facecolor='w', edgecolor='k')
largest = np.argmax(true_values[step_size])
plt.plot([true_values[step_size][largest] for _ in range(num_steps)], linestyle="--")
plt.title("Step Size: {}".format(step_size))
plt.plot(np.array(q_values[step_size])[:, largest])
plt.legend(["True Expected Value", "Estimated Value"])
plt.xlabel("Steps")
plt.ylabel("Value")
plt.show()
plt.figure(figsize=(15, 5), dpi= 80, facecolor='w', edgecolor='k')
plt.title("Step Size: 1/N(A)")
plt.plot([true_values[step_size][largest] for _ in range(num_steps)], linestyle="--")
plt.plot(np.array(n_q_values)[:, largest])
plt.legend(["True Expected Value", "Estimated Value"])
plt.xlabel("Steps")
plt.ylabel("Value")
plt.show()
# + [markdown] deletable=false editable=false nbgrader={"checksum": "f0ecb1029a80a609aa9e18b280572f82", "grade": false, "grade_id": "cell-a0948edb96aacc70", "locked": true, "schema_version": 1, "solution": false}
# These plots help clarify the performance differences between the different step sizes. A step size of 0.01 makes such small updates that the agent’s value estimate of the best action does not get close to the actual value. Step sizes of 0.5 and 1.0 both get close to the true value quickly, but are very susceptible to stochasticity in the rewards. The updates overcorrect too much towards recent rewards, and so oscillate around the true value. This means that on many steps, the action that pulls the best arm may seem worse than it actually is. A step size of 0.1 updates fairly quickly to the true value, and does not oscillate as widely around the true values as 0.5 and 1.0. This is one of the reasons that 0.1 performs quite well. Finally we see why 1/N(A) performed well. Early on while the step size is still reasonably high it moves quickly to the true expected value, but as it gets pulled more its step size is reduced which makes it less susceptible to the stochasticity of the rewards.
#
# Does this mean that 1/N(A) is always the best? When might it not be? One possible setting where it might not be as effective is in non-stationary problems. You learned about non-stationarity in the lessons. Non-stationarity means that the environment may change over time. This could manifest itself as continual change over time of the environment, or a sudden change in the environment.
#
# Let's look at how a sudden change in the reward distributions affects a step size like 1/N(A). This time we will run the environment for 2000 steps, and after 1000 steps we will randomly change the expected value of all of the arms. We compare two agents, both using epsilon-greedy with epsilon = 0.1. One uses a constant step size of 0.1, the other a step size of 1/N(A) that reduces over time.
# + deletable=false editable=false nbgrader={"checksum": "d04001916905f0fb420618c4014cc638", "grade": false, "grade_id": "cell-55536f4ac923ab96", "locked": true, "schema_version": 1, "solution": false}
epsilon = 0.1
num_steps = 2000
num_runs = 200
step_size = 0.1
plt.figure(figsize=(15, 5), dpi= 80, facecolor='w', edgecolor='k')
plt.plot([1.55 for _ in range(num_steps)], linestyle="--")
for agent in [EpsilonGreedyAgent, EpsilonGreedyAgentConstantStepsize]:
all_averages = []
for run in tqdm(range(num_runs)):
agent_info = {"num_actions": 10, "epsilon": epsilon, "step_size": step_size}
env_info = {"random_seed": run}
rl_glue = RLGlue(env, agent)
rl_glue.rl_init(agent_info, env_info)
rl_glue.rl_start()
scores = [0]
averages = []
for i in range(num_steps):
reward, state, action, is_terminal = rl_glue.rl_step()
scores.append(scores[-1] + reward)
averages.append(scores[-1] / (i + 1))
if i == 1000:
rl_glue.environment.arms = np.random.randn(10)
all_averages.append(averages)
plt.plot(np.mean(all_averages, axis=0))
plt.legend(["Best Possible", "1/N(A)", "0.1"])
plt.xlabel("Steps")
plt.ylabel("Average reward")
plt.show()
# + [markdown] deletable=false editable=false nbgrader={"checksum": "714c6cad23e3d8fe31496ffbe8674620", "grade": false, "grade_id": "cell-c4a8be88bbcc9b38", "locked": true, "schema_version": 1, "solution": false}
# Now the agent with a step size of 1/N(A) performed better at the start but then performed worse when the environment changed! What happened?
#
# Think about what the step size would be after 1000 steps. Let's say the best action gets chosen 500 times. That means the step size for that action is 1/500 or 0.002. At each step when we update the value of the action and the value is going to move only 0.002 * the error. That is a very tiny adjustment and it will take a long time for it to get to the true value.
#
# The agent with step size 0.1, however, will always update in 1/10th of the direction of the error. This means that on average it will take ten steps for it to update its value to the sample mean.
#
# These are the types of tradeoffs we have to think about in reinforcement learning. A larger step size moves us more quickly toward the true value, but can make our estimated values oscillate around the expected value. A step size that reduces over time can converge to close to the expected value, without oscillating. On the other hand, such a decaying stepsize is not able to adapt to changes in the environment. Nonstationarity---and the related concept of partial observability---is a common feature of reinforcement learning problems and when learning online.
# -
# ## Section 5: Conclusion
# + [markdown] deletable=false editable=false nbgrader={"checksum": "3c335943267e235b3001c228e4bf9ba3", "grade": false, "grade_id": "cell-3c25a546a3d44e22", "locked": true, "schema_version": 1, "solution": false}
# Great work! You have:
# - Implemented your first agent
# - Learned about the effect of epsilon, an exploration parameter, on the performance of an agent
# - Learned about the effect of step size on the performance of the agent
# - Learned about a good experiment practice of averaging across multiple runs
# -
|
Fundamentals of Reinforcement Learning/Week 1/Notebook_ Bandits and Exploration-Exploitation/C1M1-Assignment1-v9.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: mlops_test
# language: python
# name: mlops_test
# ---
# +
# Sample Code to demonstrate how to use the Batch Prediction API to score data living in an Azure Blob Storage container and output the results back to the same container
# -
import datarobot as dr
# +
# Set DataRobot connection info here
API_KEY ='YOUR API KEY'
BATCH_PREDICTIONS_URL = "https://app.datarobot.com/api/v2"
DEPLOYMENT_ID = 'YOUR DEPLOYMENT ID'
# Set Azure connection blob info here
AZURE_STORAGE_ACCOUNT = "YOUR AZURE STORAGE ACCOUNT NAME"
AZURE_STORAGE_CONTAINER = "YOUR AZURE STORAGE ACCOUNT CONTAINER"
# you can get the storage access key by going to "Access keys" in the left menu of the storage account and use either key
AZURE_STORAGE_ACCESS_KEY = "AZURE STORAGE ACCOUNT ACCESS KEY"
# Set Azure input and output file names
AZURE_INPUT_SCORING_FILE = "YOUR INPUT SCORING FILE NAME"
AZURE_OUTPUT_RESULTS_FILE = "YOUR OUTPUT RESULTS FILE NAME"
# Set name for azure credential in DataRobot
DR_CREDENTIAL_NAME = "Azure_{}".format(AZURE_STORAGE_ACCOUNT)
# -
dr.Client(token=API_KEY,endpoint=BATCH_PREDICTIONS_URL)
# +
# Create an Azure-specific Credential
# Connection String format: DefaultEndpointsProtocol=https;AccountName=****;AccountKey=****
# The connection string is also found below the access key in Azure if you want to copy that directly.
credential = dr.Credential.create_azure(
name=DR_CREDENTIAL_NAME,
azure_connection_string="DefaultEndpointsProtocol=https;AccountName={};AccountKey={};".format(AZURE_STORAGE_ACCOUNT, AZURE_STORAGE_ACCESS_KEY)
)
credential
# +
# Use this code to look up the ID of the credential object created.
credential_id = None
for cred in dr.Credential.list():
if cred.name == DR_CREDENTIAL_NAME:
credential_id = cred.credential_id
print(credential_id)
# +
# Set up our batch prediction job
# Input: Azure Blob Storage
# Output: Azure Blob Storage
job = dr.BatchPredictionJob.score(
deployment=DEPLOYMENT_ID,
intake_settings={
'type': 'azure',
'url': "https://{}.blob.core.windows.net/{}/{}".format(AZURE_STORAGE_ACCOUNT, AZURE_STORAGE_CONTAINER,AZURE_INPUT_SCORING_FILE),
"credential_id": credential_id
},
output_settings={
'type': 'azure',
'url': "https://{}.blob.core.windows.net/{}/{}".format(AZURE_STORAGE_ACCOUNT, AZURE_STORAGE_CONTAINER,AZURE_OUTPUT_RESULTS_FILE),
"credential_id": credential_id
},
# If explanations are required, uncomment the line below
max_explanations=5,
# If passthrough columns are required, use this line
passthrough_columns=['column1','column2']
)
# -
job.wait_for_completion()
job.get_status()
|
batch_predictions/azure_batch_prediction.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Universidade Federal do Rio Grande do Sul (UFRGS)
# Programa de Pós-Graduação em Engenharia Civil (PPGEC)
#
# # Efeito de segunda ordem na frequência fundamental de torres esbeltas
#
# [1. Introdução](#section_1)
# [2. Soluções analíticas exatas e aproximadas](#section_2)
# [3. Solução numérica com matrix de rigidez geométrica](#section_3)
# [4. Modelo experimental](#section_4)
# [5. Comparação de resultados](#section_5)
# [6. Conclusões](#section_6)
#
# ---
# _Prof. <NAME>, Dr.techn._ [(ORCID)](https://orcid.org/0000-0001-5640-1020)
# _Porto Alegre, RS, Brazil_
#
# +
# Importing Python modules required for this notebook
# (this cell must be executed with "shift+enter" before any other Python cell)
import numpy as np
import matplotlib.pyplot as plt
import pickle as pk
# Module for system matrices calculation
from FEM_column import *
# -
# __Resumo:__ A consideração dos efeitos de segunda ordem é imprescindível para o projeto adequado de torres esbeltas com concentração de massa no topo. Neste contexto, torres de telecomunicações devem ser dimensionadas para resistirem ao peso das antenas de transmissão. A carga compressiva produzida pelo conjunto de antenas pode tornar relevante o efeito de segunda ordem nas propriedades dinâmicas da torre, reduzindo as frequências naturais de vibração livre, causando assim um aumento na resposta ressonante frente à ação dinâmica do vento. Nesse contexto, o presente trabalho possui como objetivo avaliar experimentalmente, por meio de modelos reduzidos, a influência dos efeitos de segunda ordem na frequência fundamental de torres de telecomunicações, considerando que esta frequência é função direta da rigidez efetiva da estrutura. Para isto foram confeccionados modelos reduzidos com os quais foram realizadas uma série de medições de frequências naturais mediante o aumento progressivo da carga axial em seu topo, até a configuração de flambagem global. Por meio da comparação com os resultados provenientes do cálculo de frequência teórico, sem consideração de efeitos de segunda ordem, verificam-se valores experimentais para a frequência fundamental cada vez menores com o aumento da carga axial, quantificando-se assim a relevância destes efeitos. Finalmente, os resultados experimentais são comparados com um modelo teórico mais elaborado, no qual a matriz de rigidez da torre é corrigida por uma matriz geométrica linearizada.
#
# ## 1. Introdução <a name="section_1"></a>
#
# 1. A principal carga em torres é o vento.
# 2. Torres esbeltas tem resposta ressonante importante.
# 3. A resposta ressonante depende da frequência fundamental da torre.
# 4. Algumas torres tem grande massa adicionada ao topo (mirantes, antenas, etc.)
# 5. Massas adicionais causam um efeito de segunda ordem na resposta em vibração livre.
# 6. Neste caso resposta dinâmica à ação do vento deve considerar uma frequência reduzida.
#
# ## 2. Soluções analíticas aproximadas <a name="section_2"></a>
#
# ### 2.1. Solução exata para a barra de seção constante <a name="section_21"></a>
#
# Para torres esbeltas de seção constante é conhecida a solução analítica para a
# frequência natural de vibração livre associada ao modo fundamental:
#
# $$ f_{\rm n} = \frac{1}{2\pi} \left( \frac{1.8751}{H} \right)^2 \sqrt{\frac{EI}{\mu}} $$
#
# onde H é a altura da torre, $EI$ é a rigidez à flexão, e $\mu$ é a massa por unidade
# de comprimento. A constante 1.8751 é aproximada e consiste na menor raiz não nula e
# positiva, $a$, da equação característica:
#
# $$ \cos(a) \cosh(a) + 1 = 0 $$
#
# As demais soluções positivas estão associadas à modos superiores que não serão
# considerados neste trabalho.
#
# <img src="resources/tower.png" alt="Tower model" width="280px"/></td>
#
# A frequência fundamental pode ser calculada com esta fórmula para as três hastes
# de alumínio com seção transversal 2cm $\times$ 0,5mm e com comprimentos 22, 30 e 38cm,
# respectivamente, que são utilizadas na parte experimental deste trabalho. Assim temos:
#
# +
# Seção transversal em alumínio
EI = 7.2e10*(0.020*0.0005**3)/12
mu = 2.7e03*(0.020*0.0005)
print('Rigidez da seção transversal à flexão: {0:6.4f}Nm²'.format(EI))
print('Massa por unidade de comprimento: {0:6.4f}kg/m'.format(mu))
# -
# Aplicando-se então a fórmula para os três comprimentos de haste a serem estudados tem-se:
#
# +
H22 = 0.22
H30 = 0.30
H38 = 0.38
f0 = np.sqrt(EI/mu)/(2*np.pi)
f22 = ((1.875/H22)**2)*f0
f30 = ((1.875/H30)**2)*f0
f38 = ((1.875/H38)**2)*f0
print('Frequência fundamental para a haste de 22cm: {0:5.2f}Hz'.format(f22))
print('Frequência fundamental para a haste de 30cm: {0:5.2f}Hz'.format(f30))
print('Frequência fundamental para a haste de 38cm: {0:5.2f}Hz'.format(f38))
# -
# É importante lembrar que o resultado acima desconsidera o efeito de segunda ordem
# decorrente do peso próprio da torre ou do acréscimo de qualquer massa adicional.
#
# Como este trabalho tem como objetivo justamente a quantificação do efeito
# de segunda ordem na frequência fundamental, determina-se a seguir as cargas críticas
# de flambagem elástica (carga de Euler) para os três comprimentos de haste utilizados
# na parte experimental.
# Estas cargas correspondem ao peso da maior massa que pode ser acrescentada ao topo
# das hastes e serão posteriormente utilizadas como parâmetro de adimensionalização.
#
# +
r0 = 0.0005/np.sqrt(12) # raio de giração
P0 = (np.pi**2)*7.2E10*(0.020*0.0005) # numerador da fórmula de Euler
P22 = P0/(2*H22/r0)**2
P30 = P0/(2*H30/r0)**2
P38 = P0/(2*H38/r0)**2
print('Carga crítica para a haste de 22cm: {0:5.3f}N ({1:4.1f}g)'.format(P22, 1000*P22/9.81))
print('Carga crítica para a haste de 30cm: {0:5.3f}N ({1:4.1f}g)'.format(P30, 1000*P30/9.81))
print('Carga crítica para a haste de 38cm: {0:5.3f}N ({1:4.1f}g)'.format(P38, 1000*P38/9.81))
# -
# ### 2.2. Solução aproximada por quociente de Rayleigh <a name="section_22"></a>
#
# Para o cálculo analítico da frequência fundamental com o acréscimo de uma massa no topo
# da torre pode ser utilizado o método do quociente de Rayleigh, que representa um
# estimador da frequência natural de vibração livre e é dado pela relação entre a
# energia potencial elástica, $V$, e a energia cinética de referência, $T_{\rm ref}$:
#
# $$ f_{\rm n} \leq \frac{1}{2\pi} \sqrt{\frac{V}{T_{\rm ref}}} $$
#
# O cálculo destas energias requer a definição de uma função de interpolação para
# a linha elástica, tão próxima quanto possível da forma modal. Por exemplo, a solução
# da linha elástica para uma carga horizontal no topo é muito semelhante ao primeiro
# modo de vibração livre:
#
# $$ \varphi(\xi) = \frac{1}{2}\left(3\xi^2 - \xi^3\right)$$
#
# com $\xi = z/H$ sendo a coordenada vertical adimensionalizada. Note-se que esta
# função de interpolação está normalizada para deslocamento unitário no topo.
# Uma vez escolhida a função $\varphi(\xi)$, as energias $V$ e $T_{\rm ref}$ são
# calculadas como:
#
# \begin{align*}
# V &= \frac{1}{2} \int_0^H { EI \left[ \varphi^{\prime\prime}(z) \right] ^2 \, dz} \\
# T_{\rm ref} &= \frac{1}{2} \int_0^H {\mu \left[ \varphi(z) \right] ^2 \, dz}
# \end{align*}
#
# Para as três hastes utilizadas na parte experimental deste trabalho as frequências
# obtidas por este método estão apresentadas abaixo:
#
# +
phi = lambda z: (3*(z/H)**2 - (z/H)**3)/2 # função de interpolação
ph1 = lambda z: (6*(z/H) - 3*(z/H)**2)/2/H # primeira derivada = rotação
ph2 = lambda z: (6 - 6*(z/H))/2/(H**2) # segunda derivada = curvatura
n = 100 # número de segmentos de discretização
H = H22
zi = np.linspace(0, H22, n)
V = np.trapz(ph2(zi)**2, dx=H22/n)*(EI/2)
Tr = np.trapz(phi(zi)**2, dx=H22/n)*(mu/2)
f22r = np.sqrt(V/Tr)/(2*np.pi)
er22 = (f22r - f22)*100/f22
H = H30
zi = np.linspace(0, H30, n)
V = np.trapz(ph2(zi)**2, dx=H30/n)*(EI/2)
Tr = np.trapz(phi(zi)**2, dx=H30/n)*(mu/2)
f30r = np.sqrt(V/Tr)/(2*np.pi)
er30 = (f30r - f30)*100/f30
H = H38
zi = np.linspace(0, H38, n)
V = np.trapz(ph2(zi)**2, dx=H38/n)*(EI/2)
Tr = np.trapz(phi(zi)**2, dx=H38/n)*(mu/2)
f38r = np.sqrt(V/Tr)/(2*np.pi)
er38 = (f38r - f38)*100/f38
print('Frequência fundamental para a haste de 22cm: {0:6.2f}Hz'.format(f22r))
print('Erro de aproximação: {0:6.2f}% '.format(er22))
print('')
print('Frequência fundamental para a haste de 30cm: {0:6.2f}Hz'.format(f30r))
print('Erro de aproximação: {0:6.2f}% '.format(er30))
print('')
print('Frequência fundamental para a haste de 38cm: {0:6.2f}Hz'.format(f38r))
print('Erro de aproximação: {0:6.2f}% '.format(er38))
# -
# ### 2.3. Quociente de Rayleigh com massas adicionais <a name="section_23"></a>
#
# Portanto, o erro é de apenas 1,5% para a função de interpolação proposta, a qual
# intencionalmente difere da linha elástica associada ao modo fundamental.
# Contudo, na medida em que se acrescente massa no topo este erro tende a diminuir,
# já que as hastes neste caso serão de fato submetida a uma carga inercial
# concentrada no topo, cuja linha elástica tenderá então à função proposta.
#
# A grande vantagem do uso do quociente de Rayleigh é a fácil introdução de massas
# adicionais no denominador, as quais devem ser multiplicadas pelos respectivos valores
# da função de interpolação nas posições em que se encontram. Por exemplo, uma massa
# adicional no topo, $M_1$, deve ser multiplicada pelo valor $\left[\varphi(H/H)\right]^2 = 1^2$
# e neste caso a energia cinética de referência fica:
#
# $$ T_{\rm ref} = \frac{1}{2} \left( \int_0^H {\mu \left[ \varphi(z) \right] ^2 \, dz}
# + M_1 \cdot 1^2 \right) $$
#
# A título de exemplo, vamos aplicar a equação acima às hastes utilizadas na parte
# experimental deste trabalho. Em cada caso admite-se que a massa adicionada no topo
# corresponde à 50% da carga crítica de flambagem:
#
# +
H = H22
zi = np.linspace(0, H22, n)
V = np.trapz(ph2(zi)**2, dx=H22/n)*(EI/2)
Tr = np.trapz(phi(zi)**2, dx=H22/n)*(mu/2) + (0.5*P22/9.81)/2
f22M = np.sqrt(V/Tr)/(2*np.pi)
H = H30
zi = np.linspace(0, H30, n)
V = np.trapz(ph2(zi)**2, dx=H30/n)*(EI/2)
Tr = np.trapz(phi(zi)**2, dx=H30/n)*(mu/2) + (0.5*P30/9.81)/2
f30M = np.sqrt(V/Tr)/(2*np.pi)
H = H38
zi = np.linspace(0, H38, n)
V = np.trapz(ph2(zi)**2, dx=H38/n)*(EI/2)
Tr = np.trapz(phi(zi)**2, dx=H38/n)*(mu/2) + (0.5*P38/9.81)/2
f38M = np.sqrt(V/Tr)/(2*np.pi)
print('Frequência para a haste de 22cm com massa no topo: {0:6.2f}Hz'.format(f22M))
print('Frequência para a haste de 30cm com massa no topo: {0:6.2f}Hz'.format(f30M))
print('Frequência para a haste de 38cm com massa no topo: {0:6.2f}Hz'.format(f38M))
# -
# ### 2.4. Quociente de Rayleigh com efeito de segunda ordem
#
# Embora o cálculo acima considere, com excelente precisão, a massa adicional presente
# no topo de uma torre, ele ainda não inclui o efeito de segunda ordem devido à compressão
# decorrente de uma carga compressiva elevada. Para isso é necessário incluir-se também
# uma modificação no cálculo da energia potencial elástica do sistema, de modo a se considerar
# a deformação axial adicional:
#
# $$ V = \frac{1}{2} \left( \int_0^H { EI \left[ \varphi^{\prime\prime}(z) \right] ^2 \, dz}
# - \int_0^H { P \left[ \varphi^{\prime}(z) \right] ^2 \, dz} \right) $$
#
# onde a segunda integral corresponde ao trabalho realizado pela carga compressiva pelo
# encurtamento vertical da torre. Note-se que o sinal negativo (compressão) implica em
# uma redução da frequência natural, sendo que esta tenderá a zero na medida em que a
# carga $P$ se aproxima da carga crítica de flambagem.
#
# Aplicando-se esta nova equação às hastes sujeitas à 50% da carga de crítca
# obtem-se os seguintes resultados:
#
# +
H = H22
zi = np.linspace(0, H22, n)
V = np.trapz(ph2(zi)**2, dx=H22/n)*(EI/2)
V -= np.trapz(ph1(zi)**2, dx=H22/n)*(0.5*P22/2)
Tr = np.trapz(phi(zi)**2, dx=H22/n)*(mu/2) + (0.5*P22/9.81)/2
f22P = np.sqrt(V/Tr)/(2*np.pi)
ef22 = (f22M - f22P)*100/f22P
H = H30
zi = np.linspace(0, H30, n)
V = np.trapz(ph2(zi)**2, dx=H30/n)*(EI/2)
V -= np.trapz(ph1(zi)**2, dx=H30/n)*(0.5*P30/2)
Tr = np.trapz(phi(zi)**2, dx=H30/n)*(mu/2) + (0.5*P30/9.81)/2
f30P = np.sqrt(V/Tr)/(2*np.pi)
ef30 = (f30M - f30P)*100/f30P
H = H38
zi = np.linspace(0, H38, n)
V = np.trapz(ph2(zi)**2, dx=H38/n)*(EI/2)
V -= np.trapz(ph1(zi)**2, dx=H38/n)*(0.5*P38/2)
Tr = np.trapz(phi(zi)**2, dx=H38/n)*(mu/2) + (0.5*P38/9.81)/2
f38P = np.sqrt(V/Tr)/(2*np.pi)
ef38 = (f38M - f38P)*100/f38P
print('Frequência para a haste de 22cm com efeito de 2a ordem: {0:6.2f}Hz'.format(f22P))
print('Erro pelo efeito de 2a. ordem: {0:6.2f}% '.format(ef22))
print('')
print('Frequência para a haste de 30cm com efeito de 2a ordem: {0:6.2f}Hz'.format(f30P))
print('Erro pelo efeito de 2a. ordem: {0:6.2f}% '.format(ef30))
print('')
print('Frequência para a haste de 38cm com efeito de 2a ordem: {0:6.2f}Hz'.format(f38P))
print('Erro pelo efeito de 2a. ordem: {0:6.2f}% '.format(ef30))
# -
# Portanto, a não consideração do efeito de segunda ordem para uma carga compressiva
# correspondente a apenas 50% da carga crítica implica que a frequência natural das hastes
# de seção constante são superestimadas em aproximadamente 40%.
# Uma diferença desta magnitude tem severas implicações nos cálculos das amplitudes de
# resposta dinâmica à ação do vento.
#
# ## 3. Solução numérica com matriz de rigidez geométrica <a name="section_3"></a>
#
# ### 3.1. Matriz de rigidez elástica <a name="section_31"></a>
#
# <table>
# <tr>
# <td><img src="resources/discretization.png" alt="Discretization" width="280px"/></td>
# <td><img src="resources/element.png" alt="Finite element" width="280px"/></td>
# </tr>
# </table>
#
#
# $$ \mathbf{K} = \frac{EI}{L^3} \;
# \left[ \begin{array}{cccc}
# 12 & 6L & -12 & 6L \\
# 6L & 4L^2 & -6L & 2L^2 \\
# -12 & -6L & 12 & -6L \\
# 6L & 2L^2 & -6L & 4L^2
# \end{array} \right] $$
#
# +
# Discretiza lâminas de alumínio
L22 = 0.01*np.ones(22)
L30 = 0.01*np.ones(30)
L38 = 0.01*np.ones(38)
# Matrizes de rigidez elásticas in N/m
KE22 = stiffness(L22, EI, P=0)
KE30 = stiffness(L30, EI, P=0)
KE38 = stiffness(L38, EI, P=0)
# Visualização das matrizes
fig1, ax = plt.subplots(1, 3, figsize=(12,4))
plt.suptitle('Stiffness Matrices', fontweight='bold', fontsize=16)
hax0 = ax[0].imshow(KE22); tax0 = ax[0].title.set_text("K for 22cm blade")
hax1 = ax[1].imshow(KE30); tax1 = ax[1].title.set_text("K for 30cm blade")
hax2 = ax[2].imshow(KE38); tax2 = ax[2].title.set_text("K for 38cm blade")
# -
# ### 3.2. Matriz de rigidez geométrica <a name="section_32"></a>
#
# $$ \mathbf{K_{\rm G}} = \frac{P}{30L} \;
# \left[ \begin{array}{cccc}
# 36 & 3L & -36 & 3L \\
# 3L & 4L^2 & -3L & -L^2 \\
# -36 & -3L & 36 & -3L \\
# 3L & -L^2 & -3L & 4L^2
# \end{array} \right] $$
#
# +
# Matrizes de rigidez geométricas in N/m
KG22 = stiffness(L22, EI=0, P=-P22/2)
KG30 = stiffness(L30, EI=0, P=-P30/2)
KG38 = stiffness(L38, EI=0, P=-P38/2)
# Visualização das matrizes
fig2, ax = plt.subplots(1, 3, figsize=(12,4))
plt.suptitle('Geometric Matrices', fontweight='bold', fontsize=16)
hax0 = ax[0].imshow(KG22); tax0 = ax[0].title.set_text("K for 22cm blade")
hax1 = ax[1].imshow(KG30); tax1 = ax[1].title.set_text("K for 30cm blade")
hax2 = ax[2].imshow(KG38); tax2 = ax[2].title.set_text("K for 38cm blade")
# -
# Sobrepondo as matrizes elástica e geométrica tem-se a matriz de rigidez com
# efeito de segunda ordem:
#
# +
# Matrizes de rigidez geométricas in N/m
K22 = stiffness(L22, EI=EI, P=-P22/2)
K30 = stiffness(L30, EI=EI, P=-P30/2)
K38 = stiffness(L38, EI=EI, P=-P38/2)
# Visualização das matrizes
fig3, ax = plt.subplots(1, 3, figsize=(12,4))
plt.suptitle('2nd Order Matrices', fontweight='bold', fontsize=16)
hax0 = ax[0].imshow(K22); tax0 = ax[0].title.set_text("K for 22cm blade")
hax1 = ax[1].imshow(K30); tax1 = ax[1].title.set_text("K for 30cm blade")
hax2 = ax[2].imshow(K38); tax2 = ax[2].title.set_text("K for 38cm blade")
# -
# ### 3.3. Matriz de massa consistente <a name="section_33"></a>
#
# $$ \mathbf{M} = \frac{\mu L}{420} \;
# \left[ \begin{array}{cccc}
# 156 & 22L & 54 & -13L \\
# 22L & 4L^2 & 13L & -3L^2 \\
# 54 & 13L & 156 & -22L \\
# -13L & -3L^2 & -22L & 4L^2
# \end{array} \right] $$
#
# +
# Consistent masses in kg
M22 = consistMass(L22, mu)
M30 = consistMass(L30, mu)
M38 = consistMass(L38, mu)
# Visualização das matrizes
fig4, ax = plt.subplots(1, 3, figsize=(12,4))
plt.suptitle('Mass Matrices', fontweight='bold', fontsize=16)
hax0 = ax[0].imshow(M22); tax0 = ax[0].title.set_text("M for 22cm blade")
hax1 = ax[1].imshow(M30); tax1 = ax[1].title.set_text("M for 30cm blade")
hax2 = ax[2].imshow(M38); tax2 = ax[2].title.set_text("M for 38cm blade")
# -
# ### 3.4. Estimativa de frequências naturais <a name="section_34"></a>
#
#
#
# +
import scipy.linalg as sc
# For L = 22cm
Z22 = L22.cumsum()[::-1]
KT22 = K22[:-2,:-2]
MT22 = M22[:-2,:-2]
MT22[0,0] += 0.5*P22/9.81 # massa adicional no topo é 50% da massa de flambagem
w22, Ph22 = sc.eig(KT22, MT22)
iw = w22.argsort()
w22 = w22[iw]
Ph22 = Ph22[:,iw]
wk22 = np.sqrt(np.real(w22))
fk22 = wk22/2/np.pi
fig5, ax = plt.subplots(1, 3, figsize=(12,6))
plt.suptitle('Vibration Modes', fontweight='bold', fontsize=16)
for k in range(3):
pk = Ph22[0:-1:2,k] # retem somente a translação
pm = np.max(np.abs(pk)) # normaliza máxima amplitude unitária
ha = ax[k].plot(pk/pm, Z22);
tx = ax[k].title.set_text('Modo {0} :: fk = {1:4.2f}Hz'.format(k+1,fk22[k]))
ax[k].axis([-1.5, 1.5, 0, Z22[0]])
ax[k].grid(True)
# +
H = 0.22
P1 = P22/2
n = 22
fn = analyseCase(H, EI, mu, P1, n)
print(fn)
# -
# ## 4. Modelo experimental <a name="section_4"></a>
#
#
# ## 5. Comparação de resultados <a name="section_5"></a>
#
#
#
|
.ipynb_checkpoints/Jornadas_2020-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Validation & Novel Data Generation
#
# As all the 3 models we created were trained with a seemingly high accuracy, we then proceeded to evaluate novel data points using those models. For the same, we generated and evaluated ~750K Novel Points using QGIS. The step by step guide is mentioned in this notebook.
#
# Also, you can find all the files that we create in this notebook [here](https://drive.google.com/drive/folders/1BRlfZGzUpyTjIrAThURLgLkGEeEuDDiu?usp=sharing).
#
#
# So without any further ado, let's start!
#
# 1. In QGIS, we opened the smallest raster by size - `SA_TMI` - to get the boundary.
# 2. Then, we plotted the ~415,000 known data-points as Delimitted Text File over the raster. (`unsampled.csv` file - the one used for training of model 2). _Note - The CRS of the ratser file was changed to EPSG 4326 by clicking on the Bottom right button to match that of the CSV file.
# 
# 3. Then, by using the **Buffer Tool**, we create a circular shape arround all points. This will create a Shape file for a polygon from which we will then generate the novel points.
# 
# 
# _Note: Don't Change the default values in the dialogue box till neccesary_
# 4. Then by using the `Polygon Vector Tool`, we created a polygon encapsulating the raster's full boundries.
#
# 
# 5. Then, by using the `Difference Tool`, we created a polygon for the Area that didnt had any plotted mineral locations. The resulting polygon was saved as `AreaUnexplored.gpkg`
# 
# 
# 
# 6. Then, by using the `Random Point inside polygon Tool`, we created a vector Layer with 1 Million point lying inside the `AreaUnexplored` polygon.
#
# 7. We then saved this layer as a Delimitted Text File - `validation_unsampled.csv` for further sampling and validation. You can find the sa
# 
# 
# *Note: The generated points seem to overlap but when you zoom in, you'll find that they are spaced apart.*
# 
# Now that we have obtained the `validation_unsampled.csv`, we will now have to sample these points via QGIS to obtain the `validation_sampled.csv`. This csv will contain 1 million points, from which after dropping NA's, we obtain a final csv file with `~723K points`. This file was renamed to `Validation_Model_1.csv`.
#
# This csv file is then used to obtain predictions from the first model. The predictions are saved as `Model_1_Predictions.csv`.
# In the next notebook, we'll have a look about how the predictions of Model 2 & Model 3 are obtained.
|
validation/0_Validation_Data_Preparation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Week 3 - Functions
#
# The main objective for this part of the lecture is to develop an understanding of:
#
# * What is a function?
# * How can functions be used?
# * Why is using functions a good idea?
# ## Build-In Functions
help(pow)
pow(3, 3)
id(pow)
id(3)
# ## Creating our own functions
def BMI ( weight_kg, height_cm ):
height_m = height_cm / 100
bmi = weight_kg / (height_m ** 2)
return bmi
power=2
BMI (90.7, 182)
height_m = 98
height_m
# ## Adding documentation
def BMI ( weight_kg, height_cm ):
""" (float, float) -> float
Returns the BMI calculated by dividing weight in kilograms by the square of the height in meters squared.
>>> BMI (90.7, 182)
27.38195870064002
"""
height_m = height_cm / 100
bmi = weight_kg / (height_m ** 2)
return bmi
help(BMI)
def doSometing(a):
""" (string) -> string
This function does something great!
>>> doSomething(a)
Magic!!!
"""
return 'Magic!!!'
# ## Another function
# Let's right a new function that calculates the drip rate for an IV drip as in the following example:
#
# Imagine that you have a 1,000 mL IV bag and need to infuse 125 mg of ABC at a rate of 5 mg per hour. Assuming you use entire bag to deliver the dose of ABC, what should the drip rate be (mL per hour) in order to deliver the entire IV bag.
def drip_rate ( med_dose, med_per_time, bag_size ):
""" (int, int, int) -> float
Returns the drip rate using the units of volume from bag_size and the unit of time from the med_per_time parameter.
>>> drip_rate ( 10, 1, 10 )
1.0
"""
time = med_dose / med_per_time
drip_rate = bag_size / time
return drip_rate
help(drip_rate)
drip_rate ( 125, 5, 1000 )
drip_rate ( 10, 1, 10 )
drip_rate ( med_per_time = 5, med_dose = 125, bag_size = 1000 )
l = [[125,5,1000],[10,1,10]]
for item in l:
print(drip_rate(item[0],item[1],item[2]))
|
week03/week03-functions.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
from sklearn.datasets import load_breast_cancer
cancer = load_breast_cancer()
type(cancer)
cancer.keys()
print(cancer['DESCR'])
data = pd.DataFrame(cancer['data'],columns=cancer['feature_names'])
data.head()
cancer['target']
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
scaler.fit(data)
scaled = scaler.transform(data)
# PCA
from sklearn.decomposition import PCA
pca = PCA(n_components=2)
pca.fit(scaled)
x_pca = pca.transform(scaled)
x_pca.shape
plt.figure(figsize=(8,6))
plt.scatter(x_pca[:,0],x_pca[:,1],c=cancer['target'],cmap='plasma')
plt.xlabel('First Principal Component')
plt.ylabel('Second Principal Component')
pca.components_
data_com = pd.DataFrame(pca.components_,columns=cancer['feature_names'])
plt.figure(figsize=(12,6))
sns.heatmap(data_com,cmap='plasma')
|
Principal Component Analysis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Example Notebook
#
# Jupyter notebooks are a great tool for data analysis and prototyping. Each engineer will have their own folder under `notebooks/{username}/` to store Jupyter notebooks.
#
# This notebook shows some examples.
#
# Every notebook will start with this code cell. It moves up two directories from where the notebook is, so that we can execute code from the root directory, `transithealth/`. This allows us to import code from across project as well as read/write files using paths relative to the root directory.
import os
os.chdir("../../")
# ## Importing Code and Accessing Files
#
# Now we can import code from the backend API and run it!
from api.metrics.rideshare import RideshareMetrics
from api.utils.testing import create_test_db
# We can also refer to scripts from the offline pipeline using their relative paths from the root.
# +
con, cur = create_test_db(
scripts=[
"./pipeline/load/rideshare.sql"
],
tables={
"rideshare": [
{ "n_trips": 7 },
{ "n_trips": 14 },
{ "n_trips": 3 }
]
}
)
metric = RideshareMetrics(con)
actual = metric.get_max_trips()
expected = { "max_trips": 14 }
assert actual == expected
print(f"Actual: {actual}")
print(f"Expected: {expected}")
# -
# ## Writing SQL
#
# There are two ways to write SQL in a Jupyter notebooks:
#
# 1. With the `sqlite3` module built into Python.
# 2. With the SQL extension for Jupyter notebooks.
#
# Here is an example using Python. The default fetched response does not include the columns, so we have a method in `api.utils.database` to help. We can also use Pandas to load the result into a DataFrame.
# +
import sqlite3
con = sqlite3.connect("./pipeline/database.db")
cur = con.cursor()
rows = cur.execute("""
SELECT *
FROM rideshare
LIMIT 5
""").fetchall()
rows
# +
import pandas as pd
from api.utils.database import rows_to_dicts
pd.DataFrame(rows_to_dicts(cur, rows))
# -
# Jupyter also supports notebook extensions. This extension allows us to declare a cell as a SQL cell with `%%sql` on the first line of the cell, write queries directly in the cell body, and then view the result as a table.
# %%capture
# %load_ext sql
# %sql sqlite:///pipeline/database.db
# + language="sql"
# SELECT *
# FROM rideshare
# LIMIT 5
# -
# ## Querying a Socrata Data Portal
#
# Socrata SQL (SoQL) is a special dialect of SQL that we can use to access datasets from the City of Chicago data portal, as well as other data portals hosted on Socrata.
#
# Below is an example code cell, which uses the `request` module to send a query and get the response as well as Pandas to display the result.
#
# It can take a while to get a response, because you are sending a request to a remote server that will run your SoQL query against the entire dataset. That is why we write scripts in our offline pipeline to aggregate and download data before applying transformations locally.
# +
import pandas as pd
import requests
dataset_json_url = "https://data.cityofchicago.org/resource/m6dm-c72p.json"
query = """
SELECT
pickup_community_area,
trip_seconds / 60 as trip_minutes
LIMIT 5
"""
r = requests.get(dataset_json_url, params={"$query": query})
pd.DataFrame(r.json())
# -
|
notebooks/mgarcia18/My Example Notebook.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from CausalLSTM.data import make_forcing_colm_fluxnet
ROOT = '/hard/lilu/Fluxnet/HH/'
PATH = ROOT + 'FLX_US-Blo_FLUXNET2015_FULLSET_HH_1997-2007_1-3.csv'
make_forcing_colm_fluxnet(
PATH=PATH,
forcing_params=[
'SW_IN_F_MDS', 'LW_IN_F_MDS',
'P_F', 'TA_F_MDS', 'WS_F',
'PA_F', 'RH'],
site_name='US-Blo')
# -
|
make_forcing_colm_fluxnet.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: py3-gpu-jupyter-ds
# language: python
# name: py3-gpu-jupyter-ds
# ---
# # CNN Modeling Experiments
# ## fine tuning gesture digital
#
# QLQ Oct, 2020
#
# This notebook will contain modeling, fit and validation. and by fine tuning based on Resnet50 model
# work flowing
# - Data import
# - Split data
# - Data generator
# - Modeling
# - Compile
# - Fit
# - Validation
# - show training result chart
# - prediction and show result
#
# 下面的例子参考
# https://pchun.work/resnet%e3%82%92fine-tuning%e3%81%97%e3%81%a6%e8%87%aa%e5%88%86%e3%81%8c%e7%94%a8%e6%84%8f%e3%81%97%e3%81%9f%e7%94%bb%e5%83%8f%e3%82%92%e5%ad%a6%e7%bf%92%e3%81%95%e3%81%9b%e3%82%8b/
import tensorflow
from PIL import Image
import numpy as np
import pylab
import pandas as pd
from keras.preprocessing import image
import matplotlib.pyplot as plt
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from tensorflow.keras.layers import Input, Dense, Flatten, Dropout
from tensorflow.keras.models import Model, Sequential
from tensorflow.keras.applications.resnet50 import ResNet50
from tensorflow.keras.callbacks import EarlyStopping
from tensorflow.keras import optimizers
# ### Import data
# Changing directory because all modules are in root directory
import os
pic_path = os.path.join('../../workspace', 'python')
if os.getcwd().split('/')[-1] == 'experiment':
os.chdir(pic_path)
os.getcwd()
# +
# Global constants
base_dir = 'hand_sign_digit_data'
train_path = os.path.join(base_dir, "train")
validation_path = os.path.join(base_dir, "validation")
test_path = os.path.join(base_dir, "test")
model_name = 'CNN-fine-tuning-gesture-digital.h5'
model_path = os.path.join('../../workspace', 'model', model_name)
csv_name = 'CNN-fine-tuning-gesture-digital_result.csv'
csv_path = os.path.join('../../workspace', 'model', csv_name)
# categorical constants
# 这里定义的类的名字要和分类文件夹的名字一样,否则数据生成的时候无法找到文件
classes = ['0', '1', '2', '3', '4', '5', '6', '7', '8', '9']
classes_num = len(classes)
# other constands
img_width = 100
img_height = 100
# -
train_path,validation_path,test_path
pic=Image.open(os.path.join(train_path,'0','0_2.jpg'))
pic_array=np.asarray(pic)
plt.imshow(pic_array)
plt.show()
pic_array.shape
# ### Data generator
# +
# https://keras.io/ja/preprocessing/image/
train_datagenerator = ImageDataGenerator(
rescale=1.0 / 255,
shear_range=0.2,
zoom_range=0.2,
horizontal_flip=True,
rotation_range=30.,
width_shift_range=0.2,
height_shift_range=0.2
)
validation_datagenerator = ImageDataGenerator(rescale=1.0 / 255)
# +
train_data = train_datagenerator.flow_from_directory(train_path,
target_size=(img_width,
img_height),
color_mode='rgb',
classes=classes,
class_mode='categorical',
batch_size=32,
shuffle=True)
validation_data = validation_datagenerator.flow_from_directory(
validation_path,
target_size=(img_width, img_height),
color_mode='rgb',
classes=classes,
class_mode='categorical',
batch_size=32,
shuffle=True)
# -
# ### modeling
input_tensor = Input(shape=(img_width, img_height, 3))
base_model = ResNet50(weights='imagenet',
input_tensor=input_tensor,
input_shape=(img_width, img_height, 3),
include_top=False
# pooling='avg'
)
len(base_model.layers)
base_model.trainable = False
model = Sequential([
base_model,
# Dropout(0.25),
Flatten(),
Dense(512, activation='relu'),
# Dropout(0.5),
Dense(classes_num, activation='softmax')
])
model.summary()
# ### compile
model.compile(loss='categorical_crossentropy',
optimizer=optimizers.Adam(),
metrics=['accuracy'])
# ### fit
# 如果不重启jupyter,重复执行fit的话,每次都会继续上一次的训练继续训练
history = model.fit_generator(
train_data,
# steps_per_epoch=52,
epochs=50,
validation_data=validation_data
# callbacks=[EarlyStopping()]
# validation_steps=14
)
# ### Training result chart show with plt
# +
acc = history.history['accuracy']
val_acc = history.history['val_accuracy']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(len(acc))
plt.plot(epochs, acc, 'bo', label='Training acc')
plt.plot(epochs, val_acc, 'b', label='Validation acc')
plt.title('Training and validation accuracy')
plt.legend()
plt.figure()
plt.plot(epochs, loss, 'bo', label='Training loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.legend()
plt.show()
# -
# ### Save model
model_path='../../workspace/model/CNN-fine-tuning-gesture-digital_200.h5'
model_path
model.save(model_path)
# ### Read save
path=os.path.join('../../workspace/model/','sign_language_vgg16_1.h5')
model=tensorflow.keras.models.load_model(path)
model.summary()
# ### prediction data
test_pic=Image.open(os.path.join(test_path,'1.jpg'))
test_pic_array=np.asarray(test_pic)
plt.imshow(test_pic_array)
print(test_pic_array.shape)
plt.show()
# 导入图片的时候就进行图片尺寸更改,效果显示,基本没有失真,但是图片尺寸改变了
file = 'hand_sign_digit_data/test/1.jpg'
pre_pic = image.load_img(file, target_size=(img_width, img_height))
pre_pic_array = image.img_to_array(pre_pic)
pylab.imshow(pre_pic)
pylab.show()
def pre_processing(file):
pre_pic = image.load_img(file, target_size=(img_width, img_height))
pre_pic_array = image.img_to_array(pre_pic)
pre_pic_array = np.expand_dims(pre_pic_array, 0) / 255.0
print(pre_pic_array.shape, pre_pic_array.dtype)
return pre_pic_array
# +
#os.walk方法获取当前路径下的root(所有路径)、dirs(所有子文件夹)、files(所有文件)
files_name_list = []
def get_files_name(pic_path):
for root, dirs, files in os.walk(pic_path):
for file in files:
#print file.decode('gbk') #文件名中有中文字符时转码
if os.path.splitext(file)[1] == '.jpg':
# print(os.path.join(root, file))
files_name_list.append(os.path.join(root, file))
return files_name_list
# -
pd_res=pd.DataFrame()
path = os.path.join(test_path)
for file in get_files_name(path):
print(file)
result = model.predict(pre_processing(file))[0]
print(result)
print("prediction result TOP 5:")
# 先降序排列然后取前5个值
res = result.argsort()[::-1][:5]
res_ = [(classes[i], result[i]) for i in res]
pd_res_new=pd.DataFrame(res_,columns=['Digital_Result','Score'])
step=np.array(['200']*5)
file_name=file.split('/')[-1]
file_name=np.array([file_name]*5)
step=pd.DataFrame(step,columns=['step'])
file_name=pd.DataFrame(file_name,columns=['file_name'])
pd_res_new=pd.concat([pd_res_new,step,file_name],axis=1)
pd_res=pd.concat([pd_res,pd_res_new],axis=1)
for value in res_:
print(value)
pd_res
pd_res.to_csv(csv_path)
|
experiment/.ipynb_checkpoints/CNN-fine-tuning-gesture-digital-2020-11-08-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="wIEdlxy8eGwy" colab_type="code" colab={} cellView="form"
#@title Copyright 2020 AIEDX. Double-click here for license information.
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="GK3nFc09esyY" colab_type="text"
# # <font color='Orange' size="33" type="Arial Black">ML-</font><font color='blue' font-backgroun="red" size='33'>O</Font><font color='Orange' size='33'>ne</font> <font color="grey" size='3'> Powered by AIEdX</font>
#
#
# <img src="https://storage.googleapis.com/aiedx-public/AIEDX_logo%20-Favicon.jpg" height="80" width="80">
#
#
# ## Tier 1 Basic
#
# ###Lesson 3: Chocolate Chip Muffin
#
# <img src="https://storage.googleapis.com/aiedx-public/muffin-Art.png" height="60" width="60">
# + [markdown] id="e0iVtAHYjqN0" colab_type="text"
# ## Standard Imports
# + id="ynX8guERjbRt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 452} outputId="9dc47d7e-4661-466b-f1b8-9f90e6f43191"
# !pip install pixiedust
import pixiedust
# + [markdown] id="5H4AeVrYg1oD" colab_type="text"
# ##Clone a Git Hub Repository directly into Colabratory
#
#
# + id="jmLb00EiTNOL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 138} outputId="0957154e-40fb-47b3-998c-a962c2e734d8"
# !git clone https://github.com/AIEdX/MLOne-Basic.git
# + id="YKynoZM8P64L" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="1229ad5b-40b2-4891-db6f-970dbc4121cc"
# !dir
# + [markdown] id="yYucsz_ohL7U" colab_type="text"
#
# + [markdown] colab_type="text" id="Ux-VlINHhMQ8"
# ## Mount Google Drive on Colabratory
#
# + id="zSt943nGUHEB" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 124} outputId="694db7f7-de84-4278-ece3-ae3ba1f16e86"
from google.colab import drive
drive.mount('/content/drive')
# + [markdown] id="JrjN4FxuhZ9m" colab_type="text"
# ## Ensure that .py file is in pythonpath
# + id="bO1GxFg1Tq3j" colab_type="code" colab={}
import sys
sys.path.append('/content/MLOne-Basic/Python Exercise')
# + id="jdpR3z52aB2J" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 190} outputId="4d72a792-67ff-4988-9086-2c178ef1eb36"
sys.path
# + [markdown] id="eIQ_DGJMhj8N" colab_type="text"
# ##Debugging using PDB
# + id="KBns5S0jbS4U" colab_type="code" colab={}
import PythonBasics as pb
# + id="WwkriP-PU3kV" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="514cf48b-bf90-4cca-9021-699a56fe36c2"
# %pdb on
# + id="d50jUkT7WWUU" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 363} outputId="38ac2a72-ce4b-4466-95b9-e1bd683fa8e2"
# %xmode Plain
# %pdb on
#import pdb; pdb.set_trace()
# %debug Z=pb.add1(3,5)
Z
# + [markdown] id="V0RpUoVCgeKd" colab_type="text"
# ##Using Pixie Debugger(Interactive Debugger based on PDB)
#
# https://medium.com/codait/the-visual-python-debugger-for-jupyter-notebooks-youve-always-wanted-761713babc62
# + id="61-RlZcQiiEH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="87184b2c-bfd9-4c05-9e17-102b0d506c38"
# %pdb off
# + id="awvkokf6iypC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="a8bd4deb-10c4-4b35-d1c2-6d327a8cbca3"
# %pdb on
# + id="JPo8QSsugbdr" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 17} outputId="a0f12dbb-2d04-425b-c2dc-291d5c9c4344"
# %%pixie_debugger
Z=pb.add1(3,5)
Z
# + id="kuU9OuT8SYZB" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="60582fe9-2e79-4602-9b8e-3c3a733fa0f3"
# %cd 'MLOne-Bash
# + id="stckqNzKSfwh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="31888770-e568-4ddf-d4ac-b0def878accf"
# !git add "/content/drive/My Drive/Colab Notebooks/Untitled.ipynb"
# + id="0IR2S88NRpSr" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="9901901d-3466-4dc7-c04d-5<PASSWORD>"
# !git commit -m 'add a new GithubNClonefile'
# + id="Y-90Aa96SU3V" colab_type="code" colab={}
|
GithubNClone.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
from sound_animator import animate_sound
from util import load_wav
x = load_wav('beethoven.wav')
animate_sound(x,rate=22050,fps=20)
|
animate_sound_demo.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import mxnet as mx
from mxnet import nd, autograd
from mxnet import gluon
import numpy as np
import time
import matplotlib.pyplot as plt
data_ctx = mx.cpu()
model_ctx = mx.cpu()
# model_ctx = mx.gpu()
batch_size = 64
num_inputs = 784
num_outputs = 10
num_examples = 60000
def transform(data, label):
return data.astype(np.float32)/255, label.astype(np.float32)
train_data = mx.gluon.data.DataLoader(mx.gluon.data.vision.MNIST(train=True, transform=transform),
batch_size, shuffle=True)
test_data = mx.gluon.data.DataLoader(mx.gluon.data.vision.MNIST(train=False, transform=transform),
batch_size, shuffle=False)
net = gluon.nn.Sequential()
with net.name_scope():
net.add(gluon.nn.Dense(10))
net.collect_params().initialize(mx.init.Xavier(magnitude=2.24), ctx=model_ctx)
loss = gluon.loss.SoftmaxCrossEntropyLoss()
trainer = gluon.Trainer(net.collect_params(), 'sgd', {'learning_rate': 0.1})
# +
# define evaluation function
def evaluate_accuracy(data_iterator, net, loss_fun):
acc = mx.metric.Accuracy()
loss_avg = 0.
for i, (data, label) in enumerate(data_iterator):
data = data.as_in_context(model_ctx).reshape((-1,784))
label = label.as_in_context(model_ctx)
output = net(data)
loss = loss_fun(output, label)
predictions = nd.argmax(output, axis=1)
acc.update(preds=predictions, labels=label)
loss_avg = loss_avg*i/(i+1) + nd.mean(loss).asscalar()/(i+1)
return acc.get()[1], loss_avg
def plot_learningcurves(loss_tr,loss_ts, acc_tr,acc_ts):
xs = list(range(len(loss_tr)))
f = plt.figure(figsize=(12,6))
fg1 = f.add_subplot(121)
fg2 = f.add_subplot(122)
fg1.set_xlabel('epoch',fontsize=14)
fg1.set_title('Comparing loss functions')
fg1.semilogy(xs, loss_tr)
fg1.semilogy(xs, loss_ts)
fg1.grid(True,which="both")
fg1.legend(['training loss', 'testing loss'],fontsize=14)
fg2.set_title('Comparing accuracy')
fg1.set_xlabel('epoch',fontsize=14)
fg2.plot(xs, acc_tr)
fg2.plot(xs, acc_ts)
fg2.grid(True,which="both")
fg2.legend(['training accuracy', 'testing accuracy'],fontsize=14)
# +
epochs = 10
moving_loss = 0.
niter=0
loss_seq_train = []
loss_seq_test = []
acc_seq_train = []
acc_seq_test = []
for e in range(epochs):
start = time.time()
for i, (data, label) in enumerate(train_data):
data = data.as_in_context(model_ctx).reshape((-1,784))
label = label.as_in_context(model_ctx)
with autograd.record():
output = net(data)
cross_entropy = loss(output, label)
cross_entropy.backward()
trainer.step(data.shape[0])
##########################
# Keep a moving average of the losses
##########################
niter +=1
moving_loss = .99 * moving_loss + .01 * nd.mean(cross_entropy).asscalar()
est_loss = moving_loss/(1-0.99**niter)
end = time.time()
test_accuracy, test_loss = evaluate_accuracy(test_data, net, loss)
train_accuracy, train_loss = evaluate_accuracy(train_data, net, loss)
# save them for later
loss_seq_train.append(train_loss)
loss_seq_test.append(test_loss)
acc_seq_train.append(train_accuracy)
acc_seq_test.append(test_accuracy)
if e % 2 == 0:
print("Completed epoch %s. Train Loss: %s, Test Loss %s, Train_acc %s, Test_acc %s, Take %s s" %
(e+1, train_loss, test_loss, train_accuracy, test_accuracy, (end-start)))
## Plotting the learning curves
plot_learningcurves(loss_seq_train,loss_seq_test,acc_seq_train,acc_seq_test)
# +
net.collect_params().initialize(mx.init.Xavier(magnitude=2.24), ctx=model_ctx, force_reinit=True)
trainer = gluon.Trainer(net.collect_params(), 'sgd', {'learning_rate': 0.01, 'wd': 0.001})
moving_loss = 0.
niter=0
loss_seq_train = []
loss_seq_test = []
acc_seq_train = []
acc_seq_test = []
for e in range(epochs):
start = time.time()
for i, (data, label) in enumerate(train_data):
data = data.as_in_context(model_ctx).reshape((-1,784))
label = label.as_in_context(model_ctx)
with autograd.record():
output = net(data)
cross_entropy = loss(output, label)
cross_entropy.backward()
trainer.step(data.shape[0])
##########################
# Keep a moving average of the losses
##########################
niter +=1
moving_loss = .99 * moving_loss + .01 * nd.mean(cross_entropy).asscalar()
est_loss = moving_loss/(1-0.99**niter)
end = time.time()
test_accuracy, test_loss = evaluate_accuracy(test_data, net,loss)
train_accuracy, train_loss = evaluate_accuracy(train_data, net, loss)
# save them for later
loss_seq_train.append(train_loss)
loss_seq_test.append(test_loss)
acc_seq_train.append(train_accuracy)
acc_seq_test.append(test_accuracy)
if e % 2 == 0:
print("Completed epoch %s. Train Loss: %s, Test Loss %s, Train_acc %s, Test_acc %s, Take %s s" %
(e+1, train_loss, test_loss, train_accuracy, test_accuracy, (end-start)))
## Plotting the learning curves
plot_learningcurves(loss_seq_train,loss_seq_test,acc_seq_train,acc_seq_test)
# -
|
DS502-1704/MXNet-week1-part1/gluon-MNIST/MNIST-Gluon.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Examples of all decoders (except Kalman Filter)
#
# In this example notebook, we:
# 1. Import the necessary packages
# 2. Load a data file (spike trains and outputs we are predicting)
# 3. Preprocess the data for use in all decoders
# 4. Run some example decoders and print the goodness of fit
# 5. Plot example decoded outputs
#
# See "Examples_kf_decoder_hc" for a Kalman filter example. <br>
# Because the Kalman filter utilizes different preprocessing, we don't include an example here (to keep this notebook more understandable)
# ## 1. Import Packages
#
# Below, we import both standard packages, and functions from the accompanying .py files
#
# Note that you may need to specify the path below
# + jupyter={"outputs_hidden": false}
#Import standard packages
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
from scipy import io
from scipy import stats
import pickle
import sys
#Import function to get the covariate matrix that includes spike history from previous bins
from Neural_Decoding.preprocessing_funcs import get_spikes_with_history
#Import metrics
from Neural_Decoding.metrics import get_R2
from Neural_Decoding.metrics import get_rho
#Import decoder functions
from Neural_Decoding.decoders import WienerCascadeDecoder
from Neural_Decoding.decoders import WienerFilterDecoder
from Neural_Decoding.decoders import DenseNNDecoder
from Neural_Decoding.decoders import SimpleRNNDecoder
from Neural_Decoding.decoders import GRUDecoder
from Neural_Decoding.decoders import LSTMDecoder
from Neural_Decoding.decoders import XGBoostDecoder
from Neural_Decoding.decoders import SVRDecoder
# -
# ## 2. Load Data
# The data for this example can be downloaded at this [link](https://www.dropbox.com/s/e9mul73ur9omu5f/example_data_hc.pickle?dl=0).
#
# It is the hc-2 dataset from [crcns](https://crcns.org/data-sets/hc/hc-2). Specifically, we use the dataset "ec014.333"
#
#
# The data that we load is in the format described below. We have another example notebook, "Example_format_data_hc", that may be helpful towards putting the data in this format.
#
# Neural data should be a matrix of size "number of time bins" x "number of neurons", where each entry is the firing rate of a given neuron in a given time bin
#
# The output you are decoding should be a matrix of size "number of time bins" x "number of features you are decoding"
#
#
# + jupyter={"outputs_hidden": false}
folder='' #ENTER THE FOLDER THAT YOUR DATA IS IN
# folder='/home/jglaser/Data/DecData/'
# folder='/Users/jig289/Dropbox/Public/Decoding_Data/'
with open(folder+'example_data_hc.pickle','rb') as f:
# neural_data,pos_binned=pickle.load(f,encoding='latin1') #If using python 3
neural_data,pos_binned=pickle.load(f) #If using python 2
# +
import h5py
folder='E:/Users/samsoon.inayat/OneDrive - University of Lethbridge/Data/Neural_Decoding/' #ENTER THE FOLDER THAT YOUR DATA IS IN
filename = folder + 'NB_decoding.mat'
arrays = {}
fm = h5py.File(filename)
an = 1
num_points = 10000
aXs_C = fm['aXs_C'][an][0]
aXs_C1 = np.array(fm[fm[aXs_C][0][0]])
# for ii in range(0,aXs_C1.shape[1]):
# aXs_C1[:,ii] = aXs_C1[:,ii]/4
aYs_C = fm['aYs_C'][an][0]
aYs_C1p = np.array(fm[fm[aYs_C][0][0]])
aYs_C1 = np.zeros([aYs_C1p.shape[0],2])
aYs_C1[:,0] = aYs_C1p[:,0]
# aYs_C1[:,1] = aYs_C1p[:,0]
# plt.figure(figsize=(8, 4))
# plt.plot(aXs_C1[:,1])
# plt.xlim([0,10000])
# plt.figure(figsize=(8, 4))
# plt.plot(neural_data[:,0])
# plt.xlim([0,10000])
neural_data = aXs_C1[:num_points,:]
pos_binned = aYs_C1[:num_points,:]
fm.close()
# -
# ## 3. Preprocess Data
# ### 3A. User Inputs
# The user can define what time period to use spikes from (with respect to the output).
bins_before=4 #How many bins of neural data prior to the output are used for decoding
bins_current=1 #Whether to use concurrent time bin of neural data
bins_after=5 #How many bins of neural data after the output are used for decoding
# ### 3B. Format Covariates
# #### Format Input Covariates
#Remove neurons with too few spikes in HC dataset
nd_sum=np.nansum(neural_data,axis=0) #Total number of spikes of each neuron
rmv_nrn=np.where(nd_sum<100) #Find neurons who have less than 100 spikes total
neural_data=np.delete(neural_data,rmv_nrn,1) #Remove those neurons
# +
# Format for recurrent neural networks (SimpleRNN, GRU, LSTM)
# Function to get the covariate matrix that includes spike history from previous bins
X=get_spikes_with_history(neural_data,bins_before,bins_after,bins_current)
# Format for Wiener Filter, Wiener Cascade, XGBoost, and Dense Neural Network
#Put in "flat" format, so each "neuron / time" is a single feature
X_flat=X.reshape(X.shape[0],(X.shape[1]*X.shape[2]))
# -
# #### Format Output Covariates
# + jupyter={"outputs_hidden": false}
#Set decoding output
y=pos_binned
# + jupyter={"outputs_hidden": false}
#Remove time bins with no output (y value)
rmv_time=np.where(np.isnan(y[:,0]) | np.isnan(y[:,1])) #Find time bins with no output
X=np.delete(X,rmv_time,0) #Remove those time bins from X
X_flat=np.delete(X_flat,rmv_time,0) #Remove those time bins from X_flat
y=np.delete(y,rmv_time,0) #Remove those time bins from y
# -
# ### 3C. Split into training / testing / validation sets
# Note that hyperparameters should be determined using a separate validation set.
# Then, the goodness of fit should be be tested on a testing set (separate from the training and validation sets).
# #### User Options
#Set what part of data should be part of the training/testing/validation sets
#Note that there was a long period of no movement after about 80% of recording, so I did not use this data.
training_range=[0, 0.5]
valid_range=[0.5,0.65]
testing_range=[0.65, 0.8]
# #### Split Data
# + jupyter={"outputs_hidden": false}
num_examples=X.shape[0]
#Note that each range has a buffer of"bins_before" bins at the beginning, and "bins_after" bins at the end
#This makes it so that the different sets don't include overlapping neural data
training_set=np.arange(np.int(np.round(training_range[0]*num_examples))+bins_before,np.int(np.round(training_range[1]*num_examples))-bins_after)
testing_set=np.arange(np.int(np.round(testing_range[0]*num_examples))+bins_before,np.int(np.round(testing_range[1]*num_examples))-bins_after)
valid_set=np.arange(np.int(np.round(valid_range[0]*num_examples))+bins_before,np.int(np.round(valid_range[1]*num_examples))-bins_after)
#Get training data
X_train=X[training_set,:,:]
X_flat_train=X_flat[training_set,:]
y_train=y[training_set,:]
#Get testing data
X_test=X[testing_set,:,:]
X_flat_test=X_flat[testing_set,:]
y_test=y[testing_set,:]
#Get validation data
X_valid=X[valid_set,:,:]
X_flat_valid=X_flat[valid_set,:]
y_valid=y[valid_set,:]
# -
# ### 3D. Process Covariates
# We normalize (z_score) the inputs and zero-center the outputs.
# Parameters for z-scoring (mean/std.) should be determined on the training set only, and then these z-scoring parameters are also used on the testing and validation sets.
# + jupyter={"outputs_hidden": false}
#Z-score "X" inputs.
X_train_mean=np.nanmean(X_train,axis=0)
X_train_std=np.nanstd(X_train,axis=0)
X_train=(X_train-X_train_mean)/X_train_std
X_test=(X_test-X_train_mean)/X_train_std
X_valid=(X_valid-X_train_mean)/X_train_std
#Z-score "X_flat" inputs.
X_flat_train_mean=np.nanmean(X_flat_train,axis=0)
X_flat_train_std=np.nanstd(X_flat_train,axis=0)
X_flat_train=(X_flat_train-X_flat_train_mean)/X_flat_train_std
X_flat_test=(X_flat_test-X_flat_train_mean)/X_flat_train_std
X_flat_valid=(X_flat_valid-X_flat_train_mean)/X_flat_train_std
#Zero-center outputs
y_train_mean=np.mean(y_train,axis=0)
y_train=y_train-y_train_mean
y_test=y_test-y_train_mean
y_valid=y_valid-y_train_mean
# -
# ## 4. Run Decoders
# In this example, we are evaluating the model fit on the validation set
#
# **In this file, I only include some of the decoders. For examples of all the decoders, see the main example file (used with S1 data).**
# ### 4A. Wiener Filter (Linear Regression)
# + jupyter={"outputs_hidden": false}
#Declare model
model_wf=WienerFilterDecoder()
#Fit model
model_wf.fit(X_flat_train,y_train)
#Get predictions
y_valid_predicted_wf=model_wf.predict(X_flat_valid)
#Get metric of fit
R2s_wf=get_R2(y_valid,y_valid_predicted_wf)
print('R2s:', R2s_wf)
# -
# ### 4B. Wiener Cascade (Linear Nonlinear Model)
# + jupyter={"outputs_hidden": false}
#Declare model
model_wc=WienerCascadeDecoder(degree=2)
#Fit model
model_wc.fit(X_flat_train,y_train)
#Get predictions
y_valid_predicted_wc=model_wc.predict(X_flat_valid)
#Get metric of fit
R2s_wc=get_R2(y_valid,y_valid_predicted_wc)
print('R2s:', R2s_wc)
# -
# ### 4C. Dense (Feedfoward) Neural Network
# + jupyter={"outputs_hidden": false}
#Declare model
model_dnn=DenseNNDecoder(units=100,dropout=0.25,num_epochs=10)
#Fit model
model_dnn.fit(X_flat_train,y_train)
#Get predictions
y_valid_predicted_dnn=model_dnn.predict(X_flat_valid)
#Get metric of fit
R2s_dnn=get_R2(y_valid,y_valid_predicted_dnn)
print('R2s:', R2s_dnn)
# -
# ### 4D. LSTM
# + jupyter={"outputs_hidden": false}
#Declare model
model_lstm=LSTMDecoder(units=100,dropout=.25,num_epochs=10)
#Fit model
model_lstm.fit(X_train,y_train)
#Get predictions
y_valid_predicted_lstm=model_lstm.predict(X_valid)
#Get metric of fit
R2s_lstm=get_R2(y_valid,y_valid_predicted_lstm)
print('R2s:', R2s_lstm)
# -
# ## 5. Make Plots
# + jupyter={"outputs_hidden": false}
#As an example, I plot an example 3000 values of the x position (column index 0), both true and predicted with the Feedfoward neural network
#Note that I add back in the mean value, so that both true and predicted values are in the original coordinates
fig_x_dnn=plt.figure()
plt.plot(y_valid[:,0]+y_train_mean[0],'b')
plt.plot(y_valid_predicted_wf[:,0]+y_train_mean[0],'r')
#Save figure
# fig_x_dnn.savefig('x_position_decoding.eps')
# -
|
Examples_hippocampus/.ipynb_checkpoints/Examples_decoders_hc-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Movie Recommendation with Content-Based and Collaborative Filtering
# “*What movie should I watch this evening?*”
#
# Have you ever had to answer this question at least once when you came home from work? As for me — yes, and more than once. From Netflix to Hulu, the need to build robust movie recommendation systems is extremely important given the huge demand for personalized content of modern consumers.
#
# An example of recommendation system is such as this:
# * User A watches **Game of Thrones** and **Breaking Bad**.
# * User B does search on **Game of Thrones**, then the system suggests **Breaking Bad** from data collected about user A.
#
# Recommendation systems are used not only for movies, but on multiple other products and services like Amazon (Books, Items), Pandora/Spotify (Music), Google (News, Search), YouTube (Videos) etc.
#
# 
#
# Two most ubiquitous types of personalized recommendation systems are **Content-Based** and **Collaborative Filtering**. Collaborative filtering produces recommendations based on the knowledge of users’ attitude to items, that is it uses the “wisdom of the crowd” to recommend items. In contrast, content-based recommendation systems focus on the attributes of the items and give you recommendations based on the similarity between them.
#
# In this notebook, I will attempt at implementing these two systems to recommend movies and evaluate them to see which one performs better.
#
# After reading this post you will know:
#
# * About the MovieLens dataset problem for recommender system.
# * How to load and process the data.
# * How to do exploratory data analysis.
# * The 2 different types of recommendation engines.
# * How to develop a content-based recommendation model based on movie genres.
# * How to develop a collaborative filtering model based on user ratings.
# * Alternative approach to improve existing models.
#
# Let’s get started.
# ## The MovieLens Dataset
# One of the most common datasets that is available on the internet for building a Recommender System is the [MovieLens DataSet](https://grouplens.org/datasets/movielens/). This version of the dataset that I'm working with ([1M](https://grouplens.org/datasets/movielens/1m/)) contains 1,000,209 anonymous ratings of approximately 3,900 movies made by 6,040 MovieLens users who joined MovieLens in 2000.
#
# The data was collected by GroupLens researchers over various periods of time, depending on the size of the set. This 1M version was released on February 2003. Users were selected at random for inclusion. All users selected had rated at least 20 movies. Each user is represented by an id, and no other information is provided.
#
# The original data are contained in three files, [movies.dat](https://github.com/khanhnamle1994/movielens/blob/master/dat/movies.dat), [ratings.dat](https://github.com/khanhnamle1994/movielens/blob/master/dat/ratings.dat) and [users.dat](https://github.com/khanhnamle1994/movielens/blob/master/dat/users.dat). To make it easier to work with the data, I converted them into csv files. The process can be viewed in my [Data Processing Notebook](https://github.com/khanhnamle1994/movielens/blob/master/Data_Processing.ipynb).
#
# 
# ## Data Preparation
# Let's load this data into Python. I will load the dataset with Pandas onto Dataframes **ratings**, **users**, and **movies**. Before that, I'll also pass in column names for each CSV and read them using pandas (the column names are available in the [Readme](https://github.com/khanhnamle1994/movielens/blob/master/README.md) file).
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# Reading ratings file
# Ignore the timestamp column
ratings = pd.read_csv('ratings.csv', sep='\t', encoding='latin-1', usecols=['user_id', 'movie_id', 'rating'])
# Reading users file
users = pd.read_csv('users.csv', sep='\t', encoding='latin-1', usecols=['user_id', 'gender', 'zipcode', 'age_desc', 'occ_desc'])
# Reading movies file
movies = pd.read_csv('movies.csv', sep='\t', encoding='latin-1', usecols=['movie_id', 'title', 'genres'])
# -
# Now lets take a peak into the content of each file to understand them better.
#
# ### Ratings Dataset
# Check the top 5 rows
print(ratings.head())
# Check the file info
print(ratings.info())
# This confirms that there are 1M ratings for different user and movie combinations.
#
# ### Users Dataset
# Check the top 5 rows
print(users.head())
# Check the file info
print(users.info())
# This confirms that there are 6040 users and we have 5 features for each (unique user ID, gender, age, occupation and the zip code they are living in).
#
# ### Movies Dataset
# Check the top 5 rows
print(movies.head())
# Check the file info
print(movies.info())
# This dataset contains attributes of the 3883 movies. There are 3 columns including the movie ID, their titles, and their genres. Genres are pipe-separated and are selected from 18 genres (Action, Adventure, Animation, Children's, Comedy, Crime, Documentary, Drama, Fantasy, Film-Noir, Horror, Musical, Mystery, Romance, Sci-Fi, Thriller, War, Western).
# ## Data Exploration
# ### Titles
# Are there certain words that feature more often in Movie Titles? I'll attempt to figure this out using a word-cloud visualization.
# +
# Import new libraries
# %matplotlib inline
import wordcloud
from wordcloud import WordCloud, STOPWORDS
# Create a wordcloud of the movie titles
movies['title'] = movies['title'].fillna("").astype('str')
title_corpus = ' '.join(movies['title'])
title_wordcloud = WordCloud(stopwords=STOPWORDS, background_color='black', height=2000, width=4000).generate(title_corpus)
# Plot the wordcloud
plt.figure(figsize=(16,8))
plt.imshow(title_wordcloud)
plt.axis('off')
plt.show()
# -
# Beautiful, isn't it? I can recognize that there are a lot of movie franchises in this dataset, as evidenced by words like *II* and *III*... In addition to that, *Day*, *Love*, *Life*, *Time*, *Night*, *Man*, *Dead*, *American* are among the most commonly occuring words.
#
# ### Ratings
# Next I want to examine the **rating** further. Let's take a look at its summary statistics and distribution.
# Get summary statistics of rating
ratings['rating'].describe()
# +
# Import seaborn library
import seaborn as sns
sns.set_style('whitegrid')
sns.set(font_scale=1.5)
# %matplotlib inline
# Display distribution of rating
sns.distplot(ratings['rating'].fillna(ratings['rating'].median()))
# -
# It appears that users are quite generous in their ratings. The mean rating is 3.58 on a scale of 5. Half the movies have a rating of 4 and 5. I personally think that a 5-level rating skill wasn’t a good indicator as people could have different rating styles (i.e. person A could always use 4 for an average movie, whereas person B only gives 4 out for their favorites). Each user rated at least 20 movies, so I doubt the distribution could be caused just by chance variance in the quality of movies.
#
# Let's also take a look at a subset of 20 movies with the highest rating.
# Join all 3 files into one dataframe
dataset = pd.merge(pd.merge(movies, ratings),users)
# Display 20 movies with highest ratings
dataset[['title','genres','rating']].sort_values('rating', ascending=False).head(20)
# ### Genres
# The genres variable will surely be important while building the recommendation engines since it describes the content of the film (i.e. Animation, Horror, Sci-Fi). A basic assumption is that films in the same genre should have similar contents. I'll attempt to see exactly which genres are the most popular.
# +
# Make a census of the genre keywords
genre_labels = set()
for s in movies['genres'].str.split('|').values:
genre_labels = genre_labels.union(set(s))
# Function that counts the number of times each of the genre keywords appear
def count_word(dataset, ref_col, census):
keyword_count = dict()
for s in census:
keyword_count[s] = 0
for census_keywords in dataset[ref_col].str.split('|'):
if type(census_keywords) == float and pd.isnull(census_keywords):
continue
for s in [s for s in census_keywords if s in census]:
if pd.notnull(s):
keyword_count[s] += 1
#______________________________________________________________________
# convert the dictionary in a list to sort the keywords by frequency
keyword_occurences = []
for k,v in keyword_count.items():
keyword_occurences.append([k,v])
keyword_occurences.sort(key = lambda x:x[1], reverse = True)
return keyword_occurences, keyword_count
# Calling this function gives access to a list of genre keywords which are sorted by decreasing frequency
keyword_occurences, dum = count_word(movies, 'genres', genre_labels)
keyword_occurences[:5]
# -
# The top 5 genres are, in that respect order: Drama, Comedy, Action, Thriller, and Romance. I'll show this on a wordcloud too in order to make it more visually appealing.
# +
# Define the dictionary used to produce the genre wordcloud
genres = dict()
trunc_occurences = keyword_occurences[0:18]
for s in trunc_occurences:
genres[s[0]] = s[1]
# Create the wordcloud
genre_wordcloud = WordCloud(width=1000,height=400, background_color='white')
genre_wordcloud.generate_from_frequencies(genres)
# Plot the wordcloud
f, ax = plt.subplots(figsize=(16, 8))
plt.imshow(genre_wordcloud, interpolation="bilinear")
plt.axis('off')
plt.show()
# -
# ## Types of Recommendation Engines
#
# ### 1. Content-Based
# The Content-Based Recommender relies on the similarity of the items being recommended. The basic idea is that if you like an item, then you will also like a “similar” item. It generally works well when it's easy to determine the context/properties of each item.
#
# A content based recommender works with data that the user provides, either explicitly movie ratings for the MovieLens dataset. Based on that data, a user profile is generated, which is then used to make suggestions to the user. As the user provides more inputs or takes actions on the recommendations, the engine becomes more and more accurate.
#
# ### 2. Collaborative Filtering
# The Collaborative Filtering Recommender is entirely based on the past behavior and not on the context. More specifically, it is based on the similarity in preferences, tastes and choices of two users. It analyses how similar the tastes of one user is to another and makes recommendations on the basis of that.
#
# For instance, if user A likes movies 1, 2, 3 and user B likes movies 2,3,4, then they have similar interests and A should like movie 4 and B should like movie 1. This makes it one of the most commonly used algorithm as it is not dependent on any additional information.
#
# In general, collaborative filtering is the workhorse of recommender engines. The algorithm has a very interesting property of being able to do feature learning on its own, which means that it can start to learn for itself what features to use. It can be divided into **Memory-Based Collaborative Filtering** and **Model-Based Collaborative filtering**. In this post, I'll only focus on the Memory-Based Collaborative Filtering technique.
#
# 
# ## Content-Based Recommendation Model
# ### Theory
# The concepts of **Term Frequency (TF)** and **Inverse Document Frequency (IDF)** are used in information retrieval systems and also content based filtering mechanisms (such as a content based recommender). They are used to determine the relative importance of a document / article / news item / movie etc.
#
# TF is simply the frequency of a word in a document. IDF is the inverse of the document frequency among the whole corpus of documents. TF-IDF is used mainly because of two reasons: Suppose we search for “**the results of latest European Socccer games**” on Google. It is certain that “**the**” will occur more frequently than “**soccer games**” but the relative importance of **soccer games** is higher than the search query point of view. In such cases, TF-IDF weighting negates the effect of high frequency words in determining the importance of an item (document).
#
# Below is the equation to calculate the TF-IDF score:
# 
#
# After calculating TF-IDF scores, how do we determine which items are closer to each other, rather closer to the user profile? This is accomplished using the **Vector Space Model** which computes the proximity based on the angle between the vectors. In this model, each item is stored as a vector of its attributes (which are also vectors) in an **n-dimensional space** and the angles between the vectors are calculated to **determine the similarity between the vectors**. Next, the user profile vectors are also created based on his actions on previous attributes of items and the similarity between an item and a user is also determined in a similar way.
#
# 
#
# Sentence 2 is more likely to be using Term 2 than using Term 1. Vice-versa for Sentence 1. The method of calculating this relative measure is calculated by taking the cosine of the angle between the sentences and the terms. The ultimate reason behind using cosine is that the **value of cosine will increase with decreasing value of the angle** between which signifies more similarity. The vectors are length normalized after which they become vectors of length 1 and then the cosine calculation is simply the sum-product of vectors.
#
# ### Implementation
# With all that theory in mind, I am going to build a Content-Based Recommendation Engine that computes similarity between movies based on movie genres. It will suggest movies that are most similar to a particular movie based on its genre. To do so, I will make use of the file **movies.csv**.
# Break up the big genre string into a string array
movies['genres'] = movies['genres'].str.split('|')
# Convert genres to string value
movies['genres'] = movies['genres'].fillna("").astype('str')
# I do not have a quantitative metric to judge our machine's performance so this will have to be done qualitatively. In order to do so, I'll use **TfidfVectorizer** function from **scikit-learn**, which transforms text to feature vectors that can be used as input to estimator.
from sklearn.feature_extraction.text import TfidfVectorizer
tf = TfidfVectorizer(analyzer='word',ngram_range=(1, 2),min_df=0, stop_words='english')
tfidf_matrix = tf.fit_transform(movies['genres'])
tfidf_matrix.shape
# I will be using the **[Cosine Similarity](https://masongallo.github.io/machine/learning,/python/2016/07/29/cosine-similarity.html)** to calculate a numeric quantity that denotes the similarity between two movies. Since we have used the TF-IDF Vectorizer, calculating the Dot Product will directly give us the Cosine Similarity Score. Therefore, we will use sklearn's **linear_kernel** instead of cosine_similarities since it is much faster.
from sklearn.metrics.pairwise import linear_kernel
cosine_sim = linear_kernel(tfidf_matrix, tfidf_matrix)
cosine_sim[:4, :4]
# I now have a pairwise cosine similarity matrix for all the movies in the dataset. The next step is to write a function that returns the 20 most similar movies based on the cosine similarity score.
# +
# Build a 1-dimensional array with movie titles
titles = movies['title']
indices = pd.Series(movies.index, index=movies['title'])
# Function that get movie recommendations based on the cosine similarity score of movie genres
def genre_recommendations(title):
idx = indices[title]
sim_scores = list(enumerate(cosine_sim[idx]))
sim_scores = sorted(sim_scores, key=lambda x: x[1], reverse=True)
sim_scores = sim_scores[1:21]
movie_indices = [i[0] for i in sim_scores]
return titles.iloc[movie_indices]
# -
# Let's try and get the top recommendations for a few movies and see how good the recommendations are.
genre_recommendations('Good Will Hunting (1997)').head(20)
genre_recommendations('Toy Story (1995)').head(20)
genre_recommendations('Saving Private Ryan (1998)').head(20)
# As you can see, I have quite a decent list of recommendation for **Good Will Hunting** (Drama), **Toy Story** (Animation, Children's, Comedy), and **Saving Private Ryan** (Action, Thriller, War).
#
# Overall, here are the pros of using content-based recommendation:
# * No need for data on other users, thus no cold-start or sparsity problems.
# * Can recommend to users with unique tastes.
# * Can recommend new & unpopular items.
# * Can provide explanations for recommended items by listing content-features that caused an item to be recommended (in this case, movie genres)
#
# However, there are some cons of using this approach:
# * Finding the appropriate features is hard.
# * Does not recommend items outside a user's content profile.
# * Unable to exploit quality judgments of other users.
# ## Collaborative Filtering Recommendation Model
# The content based engine suffers from some severe limitations. It is only capable of suggesting movies which are close to a certain movie. That is, it is not capable of capturing tastes and providing recommendations across genres.
#
# Also, the engine that we built is not really personal in that it doesn't capture the personal tastes and biases of a user. Anyone querying our engine for recommendations based on a movie will receive the same recommendations for that movie, regardless of who she/he is.
#
# Therefore, in this section, I will use Memory-Based Collaborative Filtering to make recommendations to movie users. The technique is based on the idea that users similar to a me can be used to predict how much I will like a particular product or service those users have used/experienced but I have not.
#
# ### Theory
# There are 2 main types of memory-based collaborative filtering algorithms:
# 1. **User-User Collaborative Filtering**: Here we find look alike users based on similarity and recommend movies which first user’s look-alike has chosen in past. This algorithm is very effective but takes a lot of time and resources. It requires to compute every user pair information which takes time. Therefore, for big base platforms, this algorithm is hard to implement without a very strong parallelizable system.
# 2. **Item-Item Collaborative Filtering**: It is quite similar to previous algorithm, but instead of finding user's look-alike, we try finding movie's look-alike. Once we have movie's look-alike matrix, we can easily recommend alike movies to user who have rated any movie from the dataset. This algorithm is far less resource consuming than user-user collaborative filtering. Hence, for a new user, the algorithm takes far lesser time than user-user collaborate as we don’t need all similarity scores between users. And with fixed number of movies, movie-movie look alike matrix is fixed over time.
#
# 
#
# In either scenario, we builds a similarity matrix. For user-user collaborative filtering, the **user-similarity matrix** will consist of some distance metrics that measure the similarity between any two pairs of users. Likewise, the **item-similarity matrix** will measure the similarity between any two pairs of items.
#
# There are 3 distance similarity metrics that are usually used in collaborative filtering:
# 1. **Jaccard Similarity**:
# * Similarity is based on the number of users which have rated item A and B divided by the number of users who have rated either A or B
# * It is typically used where we don’t have a numeric rating but just a boolean value like a product being bought or an add being clicked
#
# 2. **Cosine Similarity**: (as in the Content-Based system)
# * Similarity is the cosine of the angle between the 2 vectors of the item vectors of A and B
# * Closer the vectors, smaller will be the angle and larger the cosine
#
# 3. **Pearson Similarity**:
# * Similarity is the pearson coefficient between the two vectors.
#
# For the purpose of diversity, I will use **Pearson Similarity** in this implementation.
#
# ### Implementation
# I will use the file **ratings.csv** first as it contains User ID, Movie IDs and Ratings. These three elements are all I need for determining the similarity of the users based on their ratings for a particular movie.
#
# First I do some quick data processing:
# +
# Fill NaN values in user_id and movie_id column with 0
ratings['user_id'] = ratings['user_id'].fillna(0)
ratings['movie_id'] = ratings['movie_id'].fillna(0)
# Replace NaN values in rating column with average of all values
ratings['rating'] = ratings['rating'].fillna(ratings['rating'].mean())
# -
# Due to the limited computing power in my laptop, I will build the recommender system using only a subset of the ratings. In particular, I will take a random sample of 20,000 ratings (2%) from the 1M ratings.
# Randomly sample 1% of the ratings dataset
small_data = ratings.sample(frac=0.02)
# Check the sample info
print(small_data.info())
# Now I use the **scikit-learn library** to split the dataset into testing and training. **Cross_validation.train_test_split** shuffles and splits the data into two datasets according to the percentage of test examples, which in this case is 0.2.
from sklearn import cross_validation as cv
train_data, test_data = cv.train_test_split(small_data, test_size=0.2)
# Now I need to create a user-item matrix. Since I have splitted the data into testing and training, I need to create two matrices. The training matrix contains 80% of the ratings and the testing matrix contains 20% of the ratings.
# +
# Create two user-item matrices, one for training and another for testing
train_data_matrix = train_data.as_matrix(columns = ['user_id', 'movie_id', 'rating'])
test_data_matrix = test_data.as_matrix(columns = ['user_id', 'movie_id', 'rating'])
# Check their shape
print(train_data_matrix.shape)
print(test_data_matrix.shape)
# -
# Now I use the **pairwise_distances** function from sklearn to calculate the [Pearson Correlation Coefficient](https://stackoverflow.com/questions/1838806/euclidean-distance-vs-pearson-correlation-vs-cosine-similarity). This method provides a safe way to take a distance matrix as input, while preserving compatibility with many other algorithms that take a vector array.
# +
from sklearn.metrics.pairwise import pairwise_distances
# User Similarity Matrix
user_correlation = 1 - pairwise_distances(train_data, metric='correlation')
user_correlation[np.isnan(user_correlation)] = 0
print(user_correlation[:4, :4])
# -
# Item Similarity Matrix
item_correlation = 1 - pairwise_distances(train_data_matrix.T, metric='correlation')
item_correlation[np.isnan(item_correlation)] = 0
print(item_correlation[:4, :4])
# With the similarity matrix in hand, I can now predict the ratings that were not included with the data. Using these predictions, I can then compare them with the test data to attempt to validate the quality of our recommender model.
#
# For the user-user CF case, I will look at the similarity between 2 users (A and B, for example) as weights that are multiplied by the ratings of a similar user B (corrected for the average rating of that user). I also need to normalize it so that the ratings stay between 1 and 5 and, as a final step, sum the average ratings for the user that I am trying to predict. The idea here is that some users may tend always to give high or low ratings to all movies. The relative difference in the ratings that these users give is more important than the absolute values.
# Function to predict ratings
def predict(ratings, similarity, type='user'):
if type == 'user':
mean_user_rating = ratings.mean(axis=1)
# Use np.newaxis so that mean_user_rating has same format as ratings
ratings_diff = (ratings - mean_user_rating[:, np.newaxis])
pred = mean_user_rating[:, np.newaxis] + similarity.dot(ratings_diff) / np.array([np.abs(similarity).sum(axis=1)]).T
elif type == 'item':
pred = ratings.dot(similarity) / np.array([np.abs(similarity).sum(axis=1)])
return pred
# ### Evaluation
# There are many evaluation metrics but one of the most popular metric used to evaluate accuracy of predicted ratings is **Root Mean Squared Error (RMSE)**. I will use the **mean_square_error (MSE)** function from sklearn, where the RMSE is just the square root of MSE.
#
# $$\mathit{RMSE} =\sqrt{\frac{1}{N} \sum (x_i -\hat{x_i})^2}$$
#
# I'll use the scikit-learn's **mean squared error** function as my validation metric. Comparing user- and item-based collaborative filtering, it looks like user-based collaborative filtering gives a better result.
# +
from sklearn.metrics import mean_squared_error
from math import sqrt
# Function to calculate RMSE
def rmse(pred, actual):
# Ignore nonzero terms.
pred = pred[actual.nonzero()].flatten()
actual = actual[actual.nonzero()].flatten()
return sqrt(mean_squared_error(pred, actual))
# +
# Predict ratings on the training data with both similarity score
user_prediction = predict(train_data_matrix, user_correlation, type='user')
item_prediction = predict(train_data_matrix, item_correlation, type='item')
# RMSE on the test data
print('User-based CF RMSE: ' + str(rmse(user_prediction, test_data_matrix)))
print('Item-based CF RMSE: ' + str(rmse(item_prediction, test_data_matrix)))
# -
# RMSE on the train data
print('User-based CF RMSE: ' + str(rmse(user_prediction, train_data_matrix)))
print('Item-based CF RMSE: ' + str(rmse(item_prediction, train_data_matrix)))
# RMSE of training of model is a metric which measure how much the signal and the noise is explained by the model. I noticed that my RMSE is quite big. I suppose I might have overfitted the training data.
#
# Overall, Memory-based Collaborative Filtering is easy to implement and produce reasonable prediction quality. However, there are some drawback of this approach:
#
# * It doesn't address the well-known cold-start problem, that is when new user or new item enters the system.
# * It can't deal with sparse data, meaning it's hard to find users that have rated the same items.
# * It suffers when new users or items that don't have any ratings enter the system.
# * It tends to recommend popular items.
# ## Alternative Approach
# As I mentioned above, it looks like my Collaborative Filtering model suffers from overfitting problem as I only train it on a small sample dataset (2% of the actual 1M ratings). In order to deal with this, I need to apply dimensionality reduction techniques to capture more signals from the big dataset. Thus comes the use of **low-dimensional factor models (aka, Model-Based Collaborative Filtering)**. I won't be able to implement this approach in this notebook due to computing limit, however, I want to introduce it here to give you a general sense of its advantages.
#
# In this approach, CF models are developed using machine learning algorithms to predict user’s rating of unrated items. It has been shown that Model-based Collaborative Filtering has received greater exposure in industry research, mainly as an unsupervised learning method for latent variable decomposition and dimensionality reduction. An example is the competition to win the [Netflix Prize](https://en.wikipedia.org/wiki/Netflix_Prize), which used the best collaborative filtering algorithm to predict user ratings for films, based on previous ratings without any other information about the users or films.
#
# Matrix factorization is widely used for recommender systems where it can deal better with scalability and sparsity than Memory-based CF. The goal of MF is to learn the latent preferences of users and the latent attributes of items from known ratings (learn features that describe the characteristics of ratings) to then predict the unknown ratings through the dot product of the latent features of users and items. As per my understanding, the algorithms in this approach can further be broken down into 3 sub-types:
#
# * **Matrix Factorization (MF)**: The idea behind such models is that attitudes or preferences of a user can be determined by a small number of hidden latent factors. These factors are also called **Embeddings**, which represent different characteristics for users and items. Matrix factorization can be done by various methods including Support Vecot Decomposition (SVD), Probabilistic Matrix Factorization (PMF), and Non-Negative Matrix Factorization (NMF).
#
# * **Clustering based algorithm (KNN)**: The idea of clustering is same as that of memory-based recommendation systems. In memory-based algorithms, we use the similarities between users and/or items and use them as weights to predict a rating for a user and an item. The difference is that the similarities in this approach are calculated based on an unsupervised learning model, rather than Pearson correlation or cosine similarity.
#
# * **Neural Nets / Deep Learning**: The idea of using Neural Nets is similar to that of Model-Based Matrix Factorization. In matrix factorizaion, we decompose our original sparse matrix into product of 2 low rank orthogonal matrices. For neural net implementation, we don’t need them to be orthogonal, we want our model to learn the values of embedding matrix itself. The user latent features and movie latent features are looked up from the embedding matrices for specific movie-user combination. These are the input values for further linear and non-linear layers. We can pass this input to multiple relu, linear or sigmoid layers and learn the corresponding weights by any optimization algorithm (Adam, SGD, etc.).
#
# 
# ## Summary
# In this post, I introduced the Movie Lens dataset for building movie recommendation system.
#
# Specifically, I have developed recommendation models including:
#
# * How to load and review the data.
# * How to develop a content-based recommendation model based on movie genres.
# * How to develop a memory-based collaborative filtering model based on user ratings.
# * A glimpse at model-based collaborative filtering models as alternative options.
|
.ipynb_checkpoints/Content_Based_and_Collaborative_Filtering_Models-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # IIR Filter -Zero Pole Placement Method for Notch Filter
#
# A **notch filter** rejects a narrow frequency band and leaves the rest of the spectrum little changed. The most common example is 60-Hz noise from power lines. Another is low-frequency ground roll. Such filters can easily be made using a slight variation on the all-pass filter. In the all-pass filter, the pole and zero have equal (logarithmic) relative distances from the unit circle. All we need to do is **put the zero closer to the circle**. Indeed, there is no reason why we should not put the zero right on the circle: then the frequency at which the zero is located is exactly canceled from the spectrum of input data.
#
# Narrow-band filters and sharp cutoff filters should be used with caution. An ever-present penalty for using such filters is that they do not decay rapidly in time. Although this may not present problems in some applications, it will certainly do so in others.
#
# 
# ## Algorithm for Designing a Notch Filter
#
# In order to design a notch filter using the zero pole placement method we perform the following steps:
# 1. Set a zero near the unit circle at the frequency of interest.
# 2. Find the complex conjugate of the zero obtained in step 1.
# 3. Set a pole and it's conjugate near the unit circle at the frequency of interest. To maintain stability, the zero should be closest to the unit circle than the pole's location.
# 4. Merge zeros and poles in a transfer function.
# 5. Adjust the zeros and poles to your needs.
import sys
sys.path.insert(0, '../')
# +
import matplotlib.pyplot as plt
import numpy as np
from Common import common_plots
from Common import fourier_inverse_transform
cplots = common_plots.Plot()
from Common import digital_filter
# -
# ## Implementation:
# #### Step 1: Set a zero near the unit circle at the frequency of interest.
#
# In this case we will define a transfer function $H_1(z)$ with a zero
# $$ H_1(z)=1-b z^{-1} $$
#
# where $b$ is a complex constant with a radius $r$ and phase angle $\phi$ radians:
# $$ b=r e^{j \phi}$$
#
#
# To implement this in our code, we need to change the transfer function $H_1(z)$ into our coding style. Remember that the format our transfer function has is the following:
#
# $$H[z] = \frac{c_0 + c_1z + c_2z^{2} + c_3z^{3} + \cdot \cdot \cdot}{d_0 + d_1z + d_2z^{2} + d_3z^{3}+ \cdot \cdot \cdot} $$
#
# Therefore, rewriting our transfer function $H_1(z)$ becomes:
#
# $$ H_1(z) = \frac{-b+z}{z}$$
#
# We now can implement a transfer function $H_1(z)$ with a radius close to the unit circle, i.e. $r=0.9$ and a value of $\phi = \pi/4$ radians.
r_zero = 0.9
phi = np.pi/4
b = r_zero*np.exp(1j*phi)
c = np.array([-b,1])
d = np.array([1])
w = np.arange(0, 2*np.pi, 0.01)
H_w = digital_filter.filter_frequency_response(c,d,w)
idft = fourier_inverse_transform.FourierInverseTransform(H_w, np.zeros(len(H_w)))
z, p, g = digital_filter.zeros_poles_gain(c, d)
# +
plt.rcParams["figure.figsize"] = (15,5)
plt.subplot(1, 3, 1)
plt.plot(w/(max(w)), np.log(np.absolute(H_w)), '.-')
plt.title('Frequency Response')
plt.xlabel('frequency')
plt.ylabel('dB')
plt.grid('on')
plt.subplot(1, 3, 2)
plt.stem(np.real(idft.synth[0:25]), use_line_collection=True)
plt.title('Impulse Response')
plt.xlabel('sample')
plt.grid('on')
cplots.plot_zeros_poles(z, p)
plt.title('Zeros and Poles')
plt.xlabel('sample');
# -
# #### Step 2: Find the complex conjugate of the zero obtained in step 1.
# Now we will expand our transfer function, and obtain $H_2(z)$. This transfer function has two zeros, one is the previous one obtained in step 1 and the other is it's conjugate.
#
# $$H_2(z) = (1-b z^{-1})(1-b^* z^{-1})$$
#
# To be able to use it, we perform some manipulations to get $H_2(z)$ into our coding style, therefore we get:
#
# $$H_2(z) = \frac{bb^*-(b+b^*)z+z^2}{z^2}$$
#
#
r_zero = 0.9
phi = np.pi/4
b = r_zero*np.exp(1j*phi)
c = np.array([b*np.conj(b),-(b+np.conj(b)), 1])
d = np.array([1])
w = np.arange(0, 2*np.pi, 0.01)
H_w = digital_filter.filter_frequency_response(c,d,w)
idft = fourier_inverse_transform.FourierInverseTransform(H_w, np.zeros(len(H_w)))
z, p, g = digital_filter.zeros_poles_gain(c, d)
# +
plt.rcParams["figure.figsize"] = (15,5)
plt.subplot(1, 3, 1)
plt.plot(w/(max(w)), np.log(np.absolute(H_w)), '.-')
plt.title('Frequency Response')
plt.xlabel('frequency')
plt.ylabel('dB')
plt.grid('on')
plt.subplot(1, 3, 2)
plt.stem(np.real(idft.synth[0:25]), use_line_collection=True)
plt.title('Impulse Response')
plt.xlabel('sample')
plt.grid('on')
cplots.plot_zeros_poles(z, p)
plt.title('Zeros and Poles')
plt.xlabel('sample');
# -
# #### Step 3: Set a pole and it's conjugate near the unit circle at the frequency of interest.
# We now proceed to create a transfer function with a pole near the unit circle. To avoid instability, we make the radius of the pole smaller than the zero.
#
# The transfer function is
# $$ H_3(z) = \frac{1}{(1-az^{-1})(1-a^*z^{-1})}$$
#
# <br>
# Again we perform some manipulations to get $H_3(z)$ into our coding style, and we get:
#
#
# $$ H_3(z) = \frac{z^2}{aa^*-(a+a^*)z+z^2}$$
r_pole = 0.85
phi = np.pi/4
a = r_pole*np.exp(1j*phi)
c = np.array([1])
d = np.array([a*np.conj(a),-(a+np.conj(a)), 1])
w = np.arange(0, 2*np.pi, 0.01)
H_w = digital_filter.filter_frequency_response(c,d,w)
idft = fourier_inverse_transform.FourierInverseTransform(H_w, np.zeros(len(H_w)))
z, p, g = digital_filter.zeros_poles_gain(c, d)
# +
plt.rcParams["figure.figsize"] = (15,5)
plt.subplot(1, 3, 1)
plt.plot(w/(max(w)), np.log(np.absolute(H_w)), '.-')
plt.title('Frequency Response')
plt.xlabel('frequency')
plt.ylabel('dB')
plt.grid('on')
plt.subplot(1, 3, 2)
plt.stem(np.real(idft.synth[0:25]), use_line_collection=True)
plt.title('Impulse Response')
plt.xlabel('sample')
plt.grid('on')
cplots.plot_zeros_poles(z, p)
plt.title('Zeros and Poles')
plt.xlabel('sample');
# -
# #### Step 4: Merge zeros and poles in a transfer function.
# Finally you can combine transfer functions $H_2(z)$ and $H_3(z)$ in a single transfer function
#
# $$H(z) = H_2(z)H_3(z)$$
#
# $$H(z) = \left(\frac{bb^*-(b+b^*)z+z^2}{z^2} \right) \left(\frac{z^2}{aa^*-(a+a^*)z+z^2}\right)$$
#
# Therefore we get
# $$H(z) = \frac{bb^*-(b+b^*)z+z^2}{aa^*-(a+a^*)z+z^2}$$
#
# which is the desired form for our transfer function.
# +
r_zero = 0.9
r_pole = 0.85
phi = np.pi/4
b = r_zero*np.exp(1j*phi)
a = r_pole*np.exp(1j*phi)
# -
c = np.array([b*np.conj(b),-(b+np.conj(b)), 1])
d = np.array([a*np.conj(a),-(a+np.conj(a)), 1])
w = np.arange(0, 2*np.pi, 0.01)
H_w = digital_filter.filter_frequency_response(c,d,w)
idft = fourier_inverse_transform.FourierInverseTransform(H_w, np.zeros(len(H_w)))
z, p, g = digital_filter.zeros_poles_gain(c, d)
# +
plt.rcParams["figure.figsize"] = (15,5)
plt.subplot(1, 3, 1)
plt.plot(w/(max(w)), np.log(np.absolute(H_w)), '.-')
plt.title('Frequency Response')
plt.xlabel('frequency')
plt.ylabel('dB')
plt.grid('on')
plt.subplot(1, 3, 2)
plt.stem(np.real(idft.synth[0:25]), use_line_collection=True)
plt.title('Impulse Response')
plt.xlabel('sample')
plt.grid('on')
cplots.plot_zeros_poles(z, p)
plt.title('Zeros and Poles')
plt.xlabel('sample');
# -
# ### Reference
# [1] https://dsp.stackexchange.com/questions/41642/filter-design-with-zero-pole-placement-method <br>
# [2] http://sepwww.stanford.edu/sep/prof/pvi/zp/paper_html/node30.html
|
11. Digital Filter Design/.ipynb_checkpoints/IIR Filter -Zero Pole Placement Method_Solution-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Assignment 2 - Elementary Probability and Information Theory
# # Boise State University DL - Dr. Kennington
#
# ### Instructions and Hints:
#
# * This notebook loads some data into a `pandas` dataframe, then does a small amount of preprocessing. Make sure your data can load by stepping through all of the cells up until question 1.
# * Most of the questions require you to write some code. In many cases, you will write some kind of probability function like we did in class using the data.
# * Some of the questions only require you to write answers, so be sure to change the cell type to markdown or raw text
# * Don't worry about normalizing the text this time (e.g., lowercase, etc.). Just focus on probabilies.
# * Most questions can be answered in a single cell, but you can make as many additional cells as you need.
# * When complete, please export as HTML. Follow the instructions on the corresponding assignment Trello card for submitting your assignment.
# +
import pandas as pd
data = pd.read_csv('pnp-train.txt',delimiter='\t',encoding='latin-1', # utf8 encoding didn't work for this
names=['type','name']) # supply the column names for the dataframe
# this next line creates a new column with the lower-cased first word
data['first_word'] = data['name'].map(lambda x: x.lower().split()[0])
# -
data[:10]
data.describe()
# ## 1. Write a probability function/distribution $P(T)$ over the types.
#
# Hints:
#
# * The Counter library might be useful: `from collections import Counter`
# * Write a function `def P(T='')` that returns the probability of the specific value for T
# * You can access the types from the dataframe by calling `data['type']`
# ## 2. What is `P(T='movie')` ?
# ## 3. Show that your probability distribution sums to one.
# ## 4. Write a joint distribution using the type and the first word of the name
#
# Hints:
#
# * The function is $P2(T,W_1)$
# * You will need to count up types AND the first words, for example: ('person','bill)
# * Using the zip function was useful for me here
# ## 5. What is P2(T='person', W1='bill')? What about P2(T='movie',W1='the')?
# ## 6. Show that your probability distribution P(T,W1) sums to one.
# ## 7. Make a new function Q(T) from marginalizing over P(T,W1) and make sure that Q(T) sums to one.
#
# Hints:
#
# * Your Q function will call P(T,W1)
# * Your check for the sum to one should be the same answer as Question 3, only it calls Q instead of P.
# ## 8. What is the KL Divergence of your Q function and your P function for Question 1?
#
# * Even if you know the answer, you still need to write code that computes it.
# ## 9. Convert from P(T,W1) to P(W1|T)
#
# Hints:
#
# * Just write a comment cell, no code this time.
# * Note that $P(T,W1) = P(W1,T)$
# ## $P(W_1|T) = \frac{P(W_1,T)}{P(T)}$
# ## 10. Write a function `Pwt` (that calls the functions you already have) to compute $P(W_1|T)$.
#
# * This will be something like the multiplication rule, but you may need to change something
# ## 11. What is P(W1='the'|T='movie')?
# ## 12. Use Baye's rule to convert from P(W1|T) to P(T|W1). Write a function Ptw to reflect this.
#
# Hints:
#
# * Call your other functions.
# * You may need to write a function for P(W1) and you may need a new counter for `data['first_word']`
# ## 13
# ### What is P(T='movie'|W1='the')?
# ### What about P(T='person'|W1='the')?
# ### What about P(T='drug'|W1='the')?
# ### What about P(T='place'|W1='the')
# ### What about P(T='company'|W1='the')
# ## 14 Given this, if the word 'the' is found in a name, what is the most likely type?
# ## 15. Is Ptw(T='movie'|W1='the') the same as Pwt(W1='the'|T='movie') the same? Why or why not?
#
# ## 16. Do you think modeling Ptw(T|W1) would be better with a continuous function like a Gaussian? Why or why not?
#
#
# * Load the `rivers.csv` file and print out a histogram
# * Which disitrbutions do you think would fit the data for the `x` column? Here is a list: https://en.wikipedia.org/wiki/Category:Continuous_distributions
# * Hint: look at some of the exponential distributions, and maybe the Pareto, you can always use Gaussian
# * Use the principle of Maximum Entropy to detemrine which of the two distributions would best fit your data (you can sum over all values in your data)
# * You will need to estimate the parameters needed for each distribution (you may need to write maximum likelihood estimation functions for estimating your parameters)
# * You can use built-in python functions to model the (pdf) distribution, if they exist. Otherwise, you may need to write your own
# * Show the Maximum Entropy calculations.
# * Calculate the KL divergence on the two distributions (in both directions)
# * make nice markdown commments so I know where everything is
# <body>
# <section style="background-color:White; font-family:Georgia;text-align:center">
# <h1 style="font-family:Garamond; color:tomato">Excercise rivers.csv</h1>
# <h2 style="font-family:Garamond; color:solid #229954">Step 1/9</h2>
# <h3 style="font-family:Garamond;">Load the rivers.csv and printout a histogram</h3>
# <hr/>
# </section>
# </body>
# +
# %matplotlib inline
import matplotlib.pyplot as plt
# -
data = pd.read_csv('rivers.csv')
data.describe()
data.x.hist()
|
introds/introds/assignment1/gc1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Predicting the position of player in Football Manager 2020
# ## Importing the required libraries and Loading Dataset
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.linear_model import LinearRegression
from sklearn.model_selection import train_test_split
from sklearn.metrics import r2_score, mean_squared_error
#Random Forest
from sklearn.ensemble import RandomForestRegressor
# -
df = pd.read_csv(r'C:\Users\Nilay\Desktop\Ashay\dataset.csv')
# ## Understanding the data
df.head()
df.shape
df.columns
df.describe()
df = df.dropna()
df['PositionsDesc']
df['PositionsDesc'].value_counts()
df = df.set_index('UID')
# ## **Removing Unwanted Columns**
#
# Since, we want to predict the position of the player from his attributes, we will have to filter the columns which we need such as Born, IntCaps and U21 details,etc.
df = df.drop(['NationID','Born','IntCaps','IntGoals','U21Caps','U21Goals'],axis=1)
position = df[['Goalkeeper', 'Sweeper', 'Striker',
'AttackingMidCentral', 'AttackingMidLeft', 'AttackingMidRight',
'DefenderCentral', 'DefenderLeft', 'DefenderRight',
'DefensiveMidfielder', 'MidfielderCentral', 'MidfielderLeft',
'MidfielderRight', 'WingBackLeft', 'WingBackRight']]
# This is what we want to predict but we can create a simpler single column to predict
df['Actual_Position']=position.idxmax(axis=1)
df = df.replace({'Goalkeeper':'GK', 'DefenderCentral':'CB','DefenderLeft':'LB', 'DefenderRight':'RB', 'DefensiveMidfielder':'DM','MidfielderCentral':'CM','Sweeper':'SW', 'Striker':'ST',
'AttackingMidCentral':'AMC', 'AttackingMidLeft':'AML', 'AttackingMidRight':'AMR','MidfielderLeft':'LM','MidfielderRight':'RM', 'WingBackLeft':'LWB', 'WingBackRight':'RWB'})
e = df.groupby(['Actual_Position'])[['Height']].mean()
e.plot()
df
# Having successfully obtained a **single coloumn corresponding to a player's position**, we can now dropo the individual position attributes columns.
df = df.drop(['Goalkeeper', 'Sweeper', 'Striker',
'AttackingMidCentral', 'AttackingMidLeft', 'AttackingMidRight',
'DefenderCentral', 'DefenderLeft', 'DefenderRight',
'DefensiveMidfielder', 'MidfielderCentral', 'MidfielderLeft',
'MidfielderRight', 'WingBackLeft', 'WingBackRight'],axis=1)
df=df.drop(['PositionsDesc'],axis=1)
df
((df.isnull().sum() / df.shape[0]) * 100).sort_values(ascending = False)
# ## Identifying Causation and Correlation in Variables
# **Which variables are most likely to affect the Position of a player?**
#
# First, we remove variables such as **Name, Controversy, etc.** which are likely to not cause any effect on position
#
# But before that let us convert position from a categorical variable to numeric variable. We choose values such that Defenders, Midfielders and Attackers can be differentiated with ease. Here, we must apply some basic knowledge of football. You can even arbitrarily assign values, but I think it would be better to keep distance between opposite positions.
x=df['Actual_Position']
df = df.replace({'GK':100,'CB':2000,'LB':2800, 'RB':2700, 'DM':2500,'CM':3300,'SW':1900, 'ST':4600,
'AMC': 3800, 'AML':3900, 'AMR':4000,'LM':3400,'RM':3500, 'LWB':2400, 'RWB':2700})
df
l = df.groupby(['Controversy'])['Actual_Position'].mean()
l.plot()
# We will use heatmaps and correlation coefficient.
#
# The standard cutoff is that the magnitude of Correlation coefficient must be
# * between 0.3-0.7 - Moderately significant variable
# * above 0.7 - Highly significant
# * below 0.3 - not significant / can be discarded
r = df[[ 'Versatility', 'Adaptability', 'Ambition', 'Loyalty',
'Pressure', 'Professional', 'Sportsmanship', 'Temperament',
'Controversy','Actual_Position']]
sns.heatmap(r.corr(),annot=True,fmt=".1f")
# So, the above attributes have no correlation with the position
#
# Therefore, we drop the columns.
df = df.drop([ 'Versatility', 'Adaptability', 'Ambition', 'Loyalty',
'Pressure', 'Professional', 'Sportsmanship', 'Temperament',
'Controversy'],axis=1)
r = df[['Consistency', 'Dirtiness', 'ImportantMatches',
'InjuryProness','Actual_Position']]
sns.heatmap(r.corr(),annot=True)
df = df.drop(['Consistency', 'Dirtiness', 'ImportantMatches',
'InjuryProness'],axis=1)
df.columns
d1 = df[['Name', 'Age', 'Height', 'Weight','Actual_Position']]
sns.heatmap(d1.corr(),annot=True,cmap='flag')
# Therefore,height has a good correlation with the position.
df=df.drop(['Name', 'Age', 'Weight'],axis=1)
d2 = df[['AerialAbility', 'CommandOfArea',
'Communication', 'Eccentricity', 'Handling','Actual_Position']]
sns.heatmap(d2.corr(),annot=True,cmap='CMRmap')
# So,all these attributes have a very good correlation with actual_position and with each other. (except maybe eccentricity)
d3 = df[['Kicking', 'OneOnOnes',
'Reflexes', 'RushingOut', 'TendencyToPunch', 'Throwing','Actual_Position']]
sns.heatmap(d3.corr(),annot=True)
# The above two correlation plots suggest that the variables arew closely related with each other(All have similar and strongcorrelation with each other). So let us see if we can combine the variables into a single one and study its correlation with the Position variable.
# This is expected as if you have some basic football knowledge, you would easily guess that the above two variable sets are Goalkeepeing attributes
d2['gk_attributes1']= d2.iloc[:,0:5].mean(axis=1)
sns.heatmap(d2.corr(),annot=True,cmap='cividis')
d2 = d2.drop(['AerialAbility', 'CommandOfArea',
'Communication', 'Eccentricity', 'Handling'],axis=1)
# So, we can replace this in actual dataframe
df['gk_attributes1'] = d2['gk_attributes1']
# Similarly for the other Goalkeeping attributes
d3['gk_attributes2']= d3.iloc[:,0:6].mean(axis=1)
sns.heatmap(d3.corr(),annot=True,cmap='ocean')
df['gk_attributes2'] = d3['gk_attributes2']
df = df.drop(['AerialAbility', 'CommandOfArea',
'Communication', 'Eccentricity', 'Handling', 'Kicking', 'OneOnOnes',
'Reflexes', 'RushingOut', 'TendencyToPunch', 'Throwing'],axis=1)
d4 = df[['Crossing', 'Dribbling', 'Finishing', 'FirstTouch','Heading', 'LongShots','Actual_Position']]
sns.heatmap(d4.corr(),annot=True,cmap='inferno')
# So, heading and crossing are not a conclusive ability in determining position.Lets drop heading and see how the average of other abilities fares
d4 = df[['Dribbling', 'Finishing', 'FirstTouch', 'LongShots','Actual_Position']]
d4['Attacking_abilities'] = d4.iloc[:,0:4].mean(axis=1)
sns.heatmap(d4.corr(),annot=True,cmap='prism')
df['Attacking_abilities'] = d4['Attacking_abilities']
df = df.drop(['Crossing', 'Dribbling', 'Finishing', 'FirstTouch','Heading', 'LongShots'],axis=1)
df
d5 = df[['Corners','Freekicks','PenaltyTaking','Longthrows','Actual_Position']]
sns.heatmap(d5.corr(),annot=True,cmap='gnuplot')
# So, none of these abilities has a definite correlation with the position. So, we can drop these columns.
df=df.drop(['Corners','Freekicks','PenaltyTaking','Longthrows'],axis=1)
df.columns
d6 = df[['Marking','Tackling','Strength','Decisions','Positioning','Actual_Position']]
sns.heatmap(d6.corr(),annot=True,cmap='gnuplot2')
# So, we can drop all the columns except positioning
df = df.drop(['Marking','Tackling','Strength','Decisions'],axis=1)
d7 = df[['Vision','Technique','Teamwork','Flair','Passing','Actual_Position']]
sns.heatmap(d7.corr(),annot=True)
# So, only technique,passing and flair are deterministic to position
df = df.drop(['Vision','Teamwork'],axis=1)
df.columns
d8 = df[['Aggression', 'Anticipation',
'Bravery', 'Composure', 'Concentration', 'Determination','Actual_Position']]
sns.heatmap(d8.corr(),annot=True,cmap='brg')
df = df.drop(['Aggression', 'Anticipation','Composure', 'Concentration', 'Determination'],axis=1)
df.columns
d9 = df[['Leadership','OffTheBall','Workrate','Actual_Position']]
sns.heatmap(d9.corr(),annot=True)
df = df.drop(['Leadership','Workrate',],axis=1)
d10 = df[['Acceleration', 'Agility',
'Balance', 'Jumping', 'NaturalFitness', 'Pace',
'Stamina','Actual_Position']]
sns.heatmap(d10.corr(),annot=True)
# None of the attributes is prominent, so all can be dropped
df = df.drop(['Acceleration', 'Agility',
'Balance', 'Jumping', 'NaturalFitness', 'Pace',
'Stamina'],axis=1)
d11 = df[['LeftFoot','RightFoot','Actual_Position']]
sns.heatmap(d11.corr(),annot=True,cmap='viridis')
df = df.drop(['LeftFoot','RightFoot'],axis=1)
df
# ## Model - Predict, Fit, Deploy, Score
X = df.drop(['Actual_Position'],axis=1)
y = df['Actual_Position']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.25, random_state=42)
ForestRegressor = RandomForestRegressor(n_estimators=500)
ForestRegressor.fit(X_train, y_train)
y_test_preds = ForestRegressor.predict(X_test)
print(r2_score(y_test, y_test_preds))
print(mean_squared_error(y_test, y_test_preds))
# +
from sklearn.linear_model import LinearRegression
lm_model = LinearRegression(normalize=True) # Instantiate
lm_model.fit(X_train, y_train) #Fit
y_test_preds2 = lm_model.predict(X_test)
"The r-squared score for your model was {} on {} values.".format(r2_score(y_test, y_test_preds2), len(y_test))
# -
y_test_pred = (y_test_preds/ 100).astype(int) *100
y_test_pred2 = (y_test_preds2/ 100).astype(int) *100
y_test_pred
print(r2_score(y_test, y_test_pred))
print(r2_score(y_test, y_test_pred2))
# ## Conclusion
# We tested 2 models
# 1. Linear Regression Technique - R square score = 84.2%
# 2. Random Forest Regression Technique - R square score = 90.48%
#
# The above R2 scores that our model has been successful in predicting the positions of the players based on the input variable attributes.
#
# In this project, we have answered the question that yes, using certain attributes we can be successful in predicting the position of players in FM2020 upto a certain degree of accuracy. We also identified the key attributesin determining the position were:
# * Goalkeeping Attributes = AerialAbility, CommandOfArea,Communication,Handling,Kicking,OneOnOnes,Reflexes,RushingOut, TendencyToPunch,Throwing
# * Height
# * Passing
# * Technique
# * Bravery
# * Flair
# * Positioning
# * Attacking attributes = Dribbling, Finishing, FirstTouch, LongShots
# ### Limitations
# However, the above implementation of the model in this code was a Regression and not a Classification.
#
# You might be wondering, "Why so?" and rightly because the aim of this project was to classify a player and assign him one of the available positions.
#
# But, in the practical world of football, it is not that a player can play in a single position. He can play in multiple positions and it would become hard for a Classifier to classify the players into a single position.
#
# Instead, what we did was assign a value to each position based on the inter-dependence and the relationship between each position. For instance, the central defenders will have similar defensive abilities(and so closer positional index) to defensive midfielders but their attacking prowess will be low. ALso, Attaacking Midfield should have similar positional index to Striker(upper bound) and centra mifield(lower bound).
# To quantify these indices, we need basic football knowledge and help from online resources of FM like http://footballmanagerblog.org//.
#
# So, the predicting model might predict a position close to each other. Like Central Attacking midfielder might be predicted as Left Attacking Midfielder. For this reason, we preferred a regression model over classification.
|
FMproject.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/violigon/Ocean_Python_03_11_2020/blob/main/Ocean_Python_03_11_2020.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="K-VduN8X8tjX" outputId="f2e3b57d-b8c1-44aa-c28f-75d716dac3c5" colab={"base_uri": "https://localhost:8080/"}
# Posso colocar qualquer texto que ele não será interpretado pela linguagem
print(1)
print("Paulo")
print(1.5)
# + id="e77bskZY9OsV" outputId="1f74835f-86a1-4acc-a903-c8b2f17921d7" colab={"base_uri": "https://localhost:8080/"}
# Variável -> Guardar uma informação para usar depois
# Armazena uma informação na memória RAM do computador
nome = "Paulo"
print(nome)
# + id="J77kaC7M-o_7" outputId="125f7a65-b497-414f-bf98-3cbf4e5823ab" colab={"base_uri": "https://localhost:8080/", "height": 251}
# Erros
nome = "Paulo"
print(nome)
print(20)
print(Teste)
print(55)
# + id="MqUbFp7O_AAj"
# Números e textos
# Número -> int (inteiros) e float (decimais)
# Texto -> string ou str
# Verdadeiro/Falso -> True ou False (bool) -> Booleano
# + id="c6c3qH9L_n8f" outputId="1457133a-06ad-4a22-bd67-37266e0e1f39" colab={"base_uri": "https://localhost:8080/"}
numero_int = 5
numero_float = 5.5
texto_str = "Paulo"
verdadeiro_bool = True
falso_bool = False
print(numero_int)
print(numero_float)
print(texto_str)
print(verdadeiro_bool)
print(falso_bool)
# + id="mlCf5nZfABN7" outputId="66b9c3c8-9553-4c49-dcae-7771f1785e9a" colab={"base_uri": "https://localhost:8080/"}
# Função type para checagem de tipos
idade = 25
nome = "João"
nivel_interesse_programacao = 9.4
usa_oculos = True
print(idade)
print(type(idade))
print(nome)
print(type(nome))
print(nivel_interesse_programacao)
print(type(nivel_interesse_programacao))
print(usa_oculos)
print(type(usa_oculos))
# + id="OdI9N4txBhW-"
idade = 25
# + id="TP4fyPayD6g4"
# Comentário de UMA linha
'''
Comentário
de
várias
linhas
'''
"""
Comentário
com
aspas
duplas
"""
"""
Estou escrevendo um texto que vai me ajudar
a estudar a programação, ou um texto que fala
um pouco sobre o que esse trecho de código faz
"""
# + id="ScjawTwjEfQS" outputId="1ffb1ed5-9401-42f4-98b8-17a8bb6b7daa" colab={"base_uri": "https://localhost:8080/"}
print(5 + 2)
print(10 + 12)
# + id="6ude8ozuFh2B" outputId="b77735a9-3048-460c-dd14-57383d7d88f1" colab={"base_uri": "https://localhost:8080/"}
soma = 1 + 2
subtracao = 3 - 1
multiplicacao = 3 * 2
divisao = 5 / 3
divisao_int = 5 // 3
print(soma)
print(type(soma))
print(subtracao)
print(type(subtracao))
print(multiplicacao)
print(type(multiplicacao))
print(divisao)
print(type(divisao))
print(divisao_int)
print(type(divisao_int))
# + [markdown] id="JK9pkakxHhKV"
# # Exercício 1 - "E os 10% do garçom?"
#
# - Defina uma variável para o valor de uma refeição que custou `R$ 42,54`;
# - Defina uma variável para o valor da taxa de serviço que é de `10%`;
# - Defina uma variável que calcula o valor total da conta e exiba-o no console (sem formatação, por enquanto, apenas o valor do resultado).
#
# + id="_iKRRVbIHz85" outputId="cc2cf91f-c8a8-4f5d-8e3c-1e3e8c671edc" colab={"base_uri": "https://localhost:8080/"}
valor_refeicao = 42.54
taxa_servico = 10
total_conta = valor_refeicao + valor_refeicao * 10 / 100
print(total_conta)
# Próximo passo: colocar R$ e formatar com apenas 2 casas decimais
# + id="BYD5SytANgXb" outputId="e2e99626-0272-4dbb-c5c7-6ea9dba2a0c2" colab={"base_uri": "https://localhost:8080/"}
# Concatenação -> União de informações, porém, feita de maneira lógica, ou seja, que o PC entenda
valor_refeicao = 42.54
taxa_servico = 10
total_conta = valor_refeicao + valor_refeicao * 10 / 100
# print("R$ " + str(total_conta))
# Formatação
print(f"Sua refeição custou R$ {valor_refeicao}, mais a taxa de serviço de {taxa_servico}%.")
print(f"O valor total ficou de R$ {total_conta:.2f}.")
# + id="jxYbrTD9P4Mo" outputId="8abcdbb6-e5c2-477f-eab8-71ff90a40ede" colab={"base_uri": "https://localhost:8080/"}
# Curiosidade, substituindo ponto de decimal por vírgula
total_com_virgula = f"R$ {total_conta:.2f}".replace(".", ",")
print(total_com_virgula)
# + id="Xk5qcfNMQtTu" outputId="c1df6913-502e-458c-8a52-7cf928963d99" colab={"base_uri": "https://localhost:8080/"}
# Indentação e formatação dos blocos
def teste():
print(1)
print(2)
if True:
print(3)
else:
print(5)
print(3)
teste()
if True:
print(4)
# + id="nBFV10AgTLRC" outputId="180ebdd6-60fe-4e0a-d9f9-70a182971abe" colab={"base_uri": "https://localhost:8080/"}
print("paulo".upper())
print("NOME EM MINÚSCULO".lower())
frase = "essa é uma frase".capitalize()
print(frase)
nome = "paulo".capitalize()
sobrenome = "salvatore".capitalize()
qtde_caracteres = len(nome) + len(sobrenome) + 1
print(f"{nome} {sobrenome} possui {qtde_caracteres} caracteres.")
frase_titulo = "texto qualquer frase".title()
print(frase_titulo)
# + id="NLOJ2cgdVTkF" outputId="2b52fd71-6604-4b37-e3d8-74c1147a6a75" colab={"base_uri": "https://localhost:8080/"}
numero = 5.555555
print(f"R$ {numero:.2f}")
# + id="1-PsEKXOVxJk" outputId="a9a5d357-04ad-43c3-9c0e-4bc88d19b0d3" colab={"base_uri": "https://localhost:8080/"}
# Para pegar a quantidade de caracteres, utilizamos o len()
# Capitalize -> Apenas a primeira letra da string maiúscula
# "paulo salvatore" vira "Paulo salvatore"
# "salvatore" vira "Salvatore"
# Title -> Todas as primeiras letras de cada palavra em uma frase viram maiúsculas
# "<NAME>" vira "<NAME>"
nome = "paulo".capitalize()
sobrenome = "salvatore".capitalize()
print(nome, len(nome))
print(sobrenome, len(sobrenome))
nome_completo = nome + " " + sobrenome
total = len(nome_completo)
print(nome_completo, total)
# + id="L-1OZqO0W6RL" outputId="35bc6abe-6cca-4e6c-a82b-476725c201af" colab={"base_uri": "https://localhost:8080/"}
nome = input("Digite o seu nome: ")
print(f"Olá, {nome}")
# + id="8HG0immSXQH5" outputId="5d057944-5440-41ea-a6b1-6428acc24c19" colab={"base_uri": "https://localhost:8080/"}
idade = int(input("Quantos anos você tem? "))
print(idade)
print(type(idade))
print(f"Daqui 5 anos você terá {idade + 5}")
# + id="wbvHgy9dXtPi" outputId="832fcbae-b6ac-4a63-fd91-a05869cce295" colab={"base_uri": "https://localhost:8080/"}
texto = int('5')
print(texto, type(texto))
# + id="GgjQ-mVnY1EN"
"""
Exercício 2
Nome: Input de Informações
Objetivo: Receber dados do usuário, trabalhar com os valores e exibir para o usuário.
Dificuldade: Principiante
1 - Crie um programa que receba do usuário seu nome, idade e gênero;
2 - Exiba na tela seguinte mensagem: "Olá, {nome}, você possui {idade} anos de idade e é do gênero {genero}.";
3 - Exiba também: "Já pensou no que você fará no seu aniversário de {idade + 1} anos?.".
Adicionando uma pimentinha extra: "Se o usuário digitar idade 1, exiba apenas "ano" em vez de "anos".
"""
|
Ocean_Python_03_11_2020.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
# -*- coding: utf-8 -*-
# This work is part of the Core Imaging Library (CIL) developed by CCPi
# (Collaborative Computational Project in Tomographic Imaging), with
# substantial contributions by UKRI-STFC and University of Manchester.
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
# http://www.apache.org/licenses/LICENSE-2.0
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# Copyright 2021 UKRI-STFC, Technical University of Denmark
# Authored by: <NAME> (DTU)
# <NAME> (UKRI-STFC)
# <NAME> (UKRI-STFC)
# <NAME> (UKRI-STFC)
# -
# # Key points demonstrated in this notebook:
#
#
#
# - ### Use CIL data readers to read in data
#
# - ### Use CIL Processors to manipulate, reduce and preprocess projection data
#
# - ### Use CIL Plugins for ASTRA or TIGRE toolbox for forward and backprojection
#
# - ### Use FBP for filtered back-projection reconstruction
#
# - ### Use CIL display tools show2D and islicer to visualise data and reconstructions
#
# - ### Use iterative algorithms such as SIRT as alternative if bad data
#
# - ### Modify image geometry to reduce reconstruction volume to save memory and time
# First import all modules we will need:
# +
import numpy as np
import os
import matplotlib.pyplot as plt
from cil.io import TXRMDataReader, TIFFWriter
from cil.processors import TransmissionAbsorptionConverter, CentreOfRotationCorrector, Slicer
from cil.framework import AcquisitionData
from cil.plugins.astra import FBP
from cil.utilities.display import show2D, show_geometry
from cil.utilities.jupyter import islicer, link_islicer
# -
# Load the 3D cone-beam projection data of a walnut:
# +
#base_dir = os.path.abspath("/mnt/materials/SIRF/Fully3D/CIL/")
#data_name = "Usb"
filename = "/mnt/materials/SIRF/Fully3D/CIL/Usb/gruppe 4_2014-03-20_1404_12/tomo-A/gruppe 4_tomo-A.txrm"
data = TXRMDataReader(file_name=filename).read()
# -
# The data is loaded in as a CIL `AcquisitionData` object:
type(data)
# We can call `print` for the data to get some basic information:
print(data)
# Note how labels refer to the different dimensions. We infer that this data set contains 801 projections each size 1024x1024 pixels.
# In addition to the data itself, `AcquisionData` contains geometric metadata in an `AcquisitionGeometry` object in the `geometry` field, which can be printed for more detailed information:
print(data.geometry)
# CIL can illustrate the scan setup visually from the AcquisitionData geometry:
show_geometry(data.geometry)
# We can use the dimension labels to extract and display 2D slices of data, such as a single projection:
show2D(data, slice_list=('angle',220))
# From the background value of 1.0 we infer that the data is transmission data (it is known to be already centered and flat field corrected) so we just need to convert to absorption/apply the negative logarithm, which can be done using a CIL processor, which will handle small/large outliers:
data = TransmissionAbsorptionConverter()(data)
# We again take a look at a slice of the data, now a vertical one to see the central slice sinogram after negative logarithm:
show2D(data, slice_list=('vertical',512))
# ## Crop data by 200 pixels on both sides to save memory and computation time
data = Slicer(roi={'horizontal':(200,-200)})(data)
show2D(data, slice_list=('vertical',512))
# CIL supports different back-ends for which data order conventions may differ. Here we use the FBP algorithm from the ASTRA Toolbox, which requires us to permute the data array into the right order:
data.dimension_labels
data.reorder(order='astra')
data.dimension_labels
# The data is now ready for reconstruction. To set up the FBP algorithm we must specify the size/geometry of the reconstruction volume. Here we use the default one:
ig = data.geometry.get_ImageGeometry()
# We can then create the FBP algorithm (really FDK since 3D cone-beam) from ASTRA running on the GPU and reconstruct the data:
fbp = FBP(ig, data.geometry, "gpu")
recon = fbp(data)
show2D(recon, slice_list=[('vertical',512), ('horizontal_x', 325)], fix_range=(-0.1,1))
# ## Offset initial angle to align reconstruction
data.geometry.set_angles(data.geometry.angles, initial_angle=-11.5)
fbp = FBP(ig, data.geometry, "gpu")
recon = fbp(data)
show2D(recon, slice_list=[('vertical',512), ('horizontal_x', 325)], fix_range=(-0.1,1))
# ## Use interactive islicer to flick through slices
islicer(recon,direction='vertical',size=10, minmax=(0.1,1))
islicer(recon,direction='horizontal_x',size=10, minmax=(0.1,1))
# ## Extract and reconstruct only central 2D slice
data2d = data.get_slice(vertical='centre')
data2d.dimension_labels
show2D(data2d)
ig2d = data2d.geometry.get_ImageGeometry()
recon2d = FBP(ig2d,data2d.geometry)(data2d)
show2D(recon2d,fix_range=(0,1))
# ## Simulate limited angle scenario with few projections
# +
idx = [*range(50,400,10)] + [*range(450,800,10)]
# A number of other projection index ranges tried
# idx = [*range(50,350,10)] + [*range(450,750,10)] #+ [*range(400,500)] + [*range(600,700)]
# idx = [*range(50,150,10)] + [*range(200,350,10)] + [*range(450,550,10)] + [*range(600,750,10)]
# idx = [*range(25,375,10)] + [*range(425,775,10)]
# idx = [*range(0,125,10)] + [*range(275,525,10)] + [*range(675,800,10)]
# idx = [*range(50,200,5)] + [*range(300,450,10)] + [*range(550,800,20)]
# idx = [*range(100,350,10)] + [*range(500,750,10)]
# idx = [*range(0,100,20)] + [*range(350,500,20)] + [*range(750,800,20)]
# +
plt.figure(figsize=(20,10))
plt.subplot(1,2,1)
plt.plot(np.cos(np.deg2rad(data2d.geometry.angles)), np.sin(np.deg2rad(data2d.geometry.angles)),'.')
plt.axis('equal')
plt.title('All angles',fontsize=20)
plt.subplot(1,2,2)
plt.plot(np.cos(np.deg2rad(data2d.geometry.angles[idx]+90-11.5)), np.sin(np.deg2rad(data2d.geometry.angles[idx]+90-11.5)),'.')
plt.axis('equal')
plt.title('Limited and few angles',fontsize=20)
# -
# ## Manually extract numpy array with selected projections only
data_array = data2d.as_array()[idx,:]
data_array.shape
data2d.as_array().shape
# ## Create updated geometry with selected angles only
ag_reduced = data2d.geometry.copy()
ag_reduced.set_angles(ag_reduced.angles[idx], initial_angle=-11.5)
# ## Combine to new `AcquisitionData` with selected data only
data2d_reduced = AcquisitionData(data_array, geometry=ag_reduced)
# ## Reconstruct by FBP
recon2d_reduced = FBP(ig2d,ag_reduced)(data2d_reduced)
show2D(recon2d_reduced, fix_range=(-0.1,1))
# ## Try iterative SIRT reconstruction
# Now set up the discrete linear inverse problem `Ax = b` and solve weighted least-squares problem using the SIRT algorithm:
from cil.plugins.astra.operators import ProjectionOperator
from cil.optimisation.algorithms import SIRT
A = ProjectionOperator(ig2d, ag_reduced, device="gpu")
# ## Specify initial guess and initialise algorithm
x0 = ig2d.allocate(0.0)
mysirt = SIRT(initial=x0, operator=A, data=data2d_reduced, max_iteration=1000)
# ## Run a low number of iterations and inspect intermediate result
mysirt.run(10, verbose=1)
show2D(mysirt.solution, fix_range=(-0.1, 1))
# ## Run more iterations and inspect
mysirt.run(90, verbose=1)
show2D(mysirt.solution, fix_range=(-0.1, 1))
# ## Run even more iterations for final SIRT reconstruction
mysirt.run(900, verbose=1)
show2D(mysirt.solution, fix_range=(-0.1, 1))
# ## Add non-negativity constraint using input `lower=0.0`
mysirt_lower0 = SIRT(initial=x0, operator=A, data=data2d_reduced, max_iteration=1000, lower=0.0, update_objective_interval=50)
mysirt_lower0.run(1000, verbose=1)
show2D(mysirt_lower0.solution, fix_range=(-0.1, 1))
# ## Compare all reduced data reconstructions in tighter colour range
show2D([recon2d_reduced, mysirt.solution, mysirt_lower0.solution], title=["FBP","SIRT","SIRT nonneg"], num_cols=3, fix_range=(-0.3,0.5))
# ## Compare horizontal line profiles
# +
linenumy = 258
plt.figure(figsize=(20,10))
plt.plot(recon2d_reduced.get_slice(horizontal_y=linenumy).as_array(),':k',label="fbp")
plt.plot(mysirt.solution.get_slice(horizontal_y=linenumy).as_array(),label="unconstrained")
plt.plot(mysirt_lower0.solution.get_slice(horizontal_y=linenumy).as_array(), label="constrained")
plt.legend(fontsize=20)
# -
# ## Go back to full data FBP reconstruction, adjust reconstruction geometry to save time and memory
show2D(recon2d,fix_range=(-0.1,1))
print(ig2d)
# ## Reduce the number of voxels
ig2d.voxel_num_x = 200
ig2d.voxel_num_y = 500
print(ig2d)
recon2d = FBP(ig2d,data2d.geometry)(data2d)
show2D(recon2d,fix_range=(0,1))
# ## Centre the reconstruction volume around the sample:
ig2d.center_x = 30*ig2d.voxel_size_x
ig2d.center_y = -40*ig2d.voxel_size_y
print(ig2d)
recon2d = FBP(ig2d,data2d.geometry)(data2d)
show2D(recon2d,fix_range=(0,1))
# ## Further reduce the reconstruction volume
ig2d.voxel_num_x = 100
ig2d.voxel_num_y = 400
print(ig2d)
recon2d = FBP(ig2d,data2d.geometry)(data2d)
show2D(recon2d,fix_range=(0,1))
print(ig2d)
# ## Increase voxel size by a factor
ig2d.voxel_size_x = 4*ig2d.voxel_size_x
ig2d.voxel_size_y = 4*ig2d.voxel_size_y
print(ig2d)
recon2d = FBP(ig2d,data2d.geometry)(data2d)
show2D(recon2d,fix_range=(0,1))
# ## Reduce number of voxels by same factor
ig2d.voxel_num_x = 25
ig2d.voxel_num_y = 100
recon2d = FBP(ig2d,data2d.geometry)(data2d)
show2D(recon2d,fix_range=(0,1))
|
examples/1_Introduction/05_usb_limited_angle_fbp_sirt.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Dask can read table from multiple files:
import dask.dataframe as dd
df = dd.read_csv('./data/*.csv.gz', compression = 'gzip')
df.head()
type(df)
|
notebooks/dask/dask_read_distributed_data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # CT-LTI: Figure 4.
# These are the plots found in Figures 4a, 4b and 4c and containing training metrics over epochs.
#
# Please make sure that the required data folder is available at the paths used by the script.
# You may generate the required data by running the python script
# ```nodec_experiments/ct_lti/gen_parameters.py```.
#
# Please also make sure that a training and an evaluation proceedures has produced results in the corresponding paths used below.
# Running ```nodec_experiments/ct_lti/single_sample/train.ipynb``` and
# ```nodec_experiments/ct_lti/single_sample/figure_4_evaluate.ipynb```
# with default paths is expected to generate at the required location.
#
# As neural network intialization is stochastic, please make sure that appropriate seeds are used or expect some variance to paper results.
#
# ## Imports
# +
# # %load_ext autoreload
# # %autoreload 2
# +
import os
os.sys.path.append('../../../')
import torch
import numpy as np
import pandas as pd
from tqdm.cli import tqdm
import plotly
import plotly.express as px
from nnc.controllers.baselines.ct_lti.dynamics import ContinuousTimeInvariantDynamics
from nnc.controllers.baselines.ct_lti.optimal_controllers import ControllabiltyGrammianController
from nnc.helpers.torch_utils.graphs import adjacency_tensor, drivers_to_tensor
from nnc.helpers.graph_helper import load_graph
from nnc.helpers.torch_utils.evaluators import FixedInteractionEvaluator
from nnc.helpers.torch_utils.losses import FinalStepMSE
from nnc.helpers.torch_utils.trainers import NODECTrainer
from nnc.helpers.torch_utils.file_helpers import read_tensor_from_collection
from nnc.controllers.neural_network.nnc_controllers import NNCDynamics
from nnc.helpers.torch_utils.nn_architectures.fully_connected import StackedDenseTimeControl
from nnc.helpers.plot_helper import base_layout, sci_notation, ColorRegistry
# -
# ## Load sample and parameters
# Here we load the sample that we trained NODEC on as well as the parameters for the dynamics.
# +
device = 'cpu'
graph='lattice'
# load graph data
experiment_data_folder = '../../../../data/parameters/ct_lti/'
results_data_folder = '../../../../data/results/ct_lti/single_sample/'
graph_folder = experiment_data_folder+graph+'/'
adj_matrix = torch.load(graph_folder+'adjacency.pt').to(dtype=torch.float, device=device)
n_nodes = adj_matrix.shape[0]
drivers = torch.load(graph_folder + 'drivers.pt')
n_drivers = len(drivers)
pos = pd.read_csv(graph_folder + 'pos.csv').set_index('index').values
driver_matrix = drivers_to_tensor(n_nodes, drivers).to(device)
target_states = torch.load(graph_folder+'target_states.pt').to(device)
initial_states = torch.load(experiment_data_folder+'init_states.pt').to(device)
current_sample_id = 24
x0 = initial_states[current_sample_id].unsqueeze(0)
xstar = target_states[current_sample_id].unsqueeze(0)
# total time for control
total_time=0.5
# select dynamics type and initial-target states
dyn = ContinuousTimeInvariantDynamics(adj_matrix, driver_matrix)
# Below is a helper function that loads parameters from a specific epoch and uses them to evaluate.
def check_for_params(params, n_interactions, logdir=None, epoch=0):
nn = StackedDenseTimeControl(n_nodes,
n_drivers,
n_hidden=0,
hidden_size=15,
activation=torch.nn.functional.elu,
use_bias=True
).to(x0.device)
nndyn = NNCDynamics(dyn, nn).to(x0.device)
nndyn.nnc.load_state_dict(params)
loss_fn = FinalStepMSE(xstar, total_time=total_time)
nn_evaluator = FixedInteractionEvaluator(
'early_eval_nn_sample_ninter_' + str(n_interactions),
log_dir=logdir,
n_interactions=n_interactions,
loss_fn=loss_fn,
ode_solver=None,
ode_solver_kwargs={'method' : 'dopri5'},
preserve_intermediate_states=False,
preserve_intermediate_controls=False,
preserve_intermediate_times=False,
preserve_intermediate_energies=False,
preserve_intermediate_losses=False,
preserve_params=False,
)
nn_res = nn_evaluator.evaluate(dyn, nndyn.nnc, x0, total_time, epoch=epoch)
return nn_evaluator, nn_res
all_epochs = pd.read_csv(results_data_folder + 'nn_sample_train/epoch_metadata.csv')['epoch']
# -
# ## Generating Figure 4a.
# For this figure we first need to load all stored parameters per epoch and evaluate them for all 3 different interaction intervals $10^{-2}, 10^{-3}, 10^{-4}$. Since this is a costly operation, we can also choose to reload an existing file if there is one.
#
# ### Getting the data
losses_df = pd.read_csv(results_data_folder + 'nn_sample_train/losses_interactions_training.csv',
engine='python')
for i, column in enumerate(losses_df.columns):
if i >0:
losses_df.columns.values[i] = int(column)
# Please check here if columns need to be string or int, different pandas version return different outcomes
losses_melted = losses_df.reset_index().melt(id_vars='Epoch', value_vars=[50,500,5000],
var_name='Interaction Interval',
value_name = 'Total Loss')
# From total interactions to interval
losses_melted['Interaction Interval'] = total_time/losses_melted['Interaction Interval'].astype(float)
losses_melted['Interaction Interval'] = losses_melted['Interaction Interval'].map(lambda x: sci_notation(x))
# ## Plotting the figure
# +
train_file = results_data_folder + 'nn_sample_train/'
evaluation_files = dict(oc_50 = results_data_folder + 'oc_sample_ninter_50/',
oc_500 = results_data_folder + 'oc_sample_ninter_500/',
oc_5000 = results_data_folder + 'oc_sample_ninter_5000/',
nodec_50 = results_data_folder + 'eval_nn_sample_ninter_50/',
nodec_500 = results_data_folder + 'eval_nn_sample_ninter_500/',
nodec_5000 = results_data_folder + 'eval_nn_sample_ninter_5000/',
)
oc_500_df = pd.read_csv(evaluation_files['oc_500'] + 'epoch_metadata.csv')
nn_500_df = pd.read_csv(evaluation_files['nodec_500'] + 'epoch_metadata.csv')
oc_500_loss_val = oc_500_df['final_loss'].values[0]
oc_500_energy_val = oc_500_df['total_energy'].values[0]
df_training = pd.read_csv(train_file+'/epoch_metadata.csv')
epoch_range = [df_training['epoch'].min(), df_training['epoch'].max()]
nodec_500_loss = px.line(df_training[['total_energy', 'epoch', 'final_loss']], x='epoch', y='final_loss').data[0]
nodec_500_loss.name = 'NODEC Loss'
nodec_500_loss = px.line(df_training[['epoch', 'final_loss']], x='epoch', y='final_loss').data[0]
nodec_500_loss.line.color = ColorRegistry.nodec
nodec_500_loss.name = 'NODEC Loss'
nodec_500_loss.showlegend = True
oc_500_loss = px.line(x=epoch_range, y=[oc_500_loss_val, oc_500_loss_val]).data[0]
oc_500_loss.name = 'OC Loss'
oc_500_loss.line.color = ColorRegistry.oc
oc_500_loss.showlegend = True
nodec_500_energy = px.line(df_training[['total_energy', 'epoch', 'final_loss']], x='epoch', y='total_energy').data[0]
nodec_500_energy.line.color = ColorRegistry.nodec
nodec_500_energy.name = 'NODEC Energy'
nodec_500_energy.showlegend = True
nodec_500_energy.line.dash = 'dot'
oc_500_energy = px.line(x=epoch_range, y=[oc_500_energy_val, oc_500_energy_val]).data[0]
oc_500_energy.name = 'OC Energy'
oc_500_energy.line.dash = 'dot'
oc_500_energy.line.color = ColorRegistry.oc
oc_500_energy.showlegend = True
fig_epoch_comparison = plotly.subplots.make_subplots(1,1, specs=[[{"secondary_y": True}]])
fig_epoch_comparison.add_trace(nodec_500_energy, secondary_y=True)
fig_epoch_comparison.add_trace(oc_500_energy, secondary_y=True)
fig_epoch_comparison.add_trace(nodec_500_loss)
fig_epoch_comparison.add_trace(oc_500_loss)
fig_epoch_comparison.update_layout(base_layout)
fig_epoch_comparison.update_yaxes(type='log', exponentformat='power', showgrid=False)
fig_epoch_comparison.update_layout(#width=240,
#height=180,
width = 210,
height=210,
margin = dict(t=50,b=0,l=0,r=20),
legend=dict(
orientation="h",
x=0.0,
y=1.4,
bgcolor="rgba(0,0,0,0)",
bordercolor="Black",
borderwidth=0
)
)
fig_epoch_comparison.layout.yaxis.title = 'Final Loss'
fig_epoch_comparison.layout.yaxis2.title = 'Total Energy'
fig_epoch_comparison.layout.xaxis.title = 'Epoch'
fig_epoch_comparison.layout.yaxis.exponentformat = 'SI'
fig_epoch_comparison.layout.yaxis.nticks = 7
fig_epoch_comparison.update_layout(width=400, height=300)
fig_epoch_comparison
# -
# ## Generating Figure 4b
# For this figure we collect all loss and energy values for $10^{-3}$ interaction interval time per epoch.
param_squared_norms = []
for epoch in tqdm(all_epochs):
params = read_tensor_from_collection(results_data_folder + 'nn_sample_train/' + 'epochs', 'nodec_params/ep_'+str(epoch)+'.pt')
squared_norm = sum([(param**2).sum().item() for param in params.values()])
param_squared_norms.append(squared_norm)
# +
vcolors = np.array(plotly.colors.qualitative.Dark24)
vcolors = [col.replace('rgb', 'rgba').replace(')', ',0.3)') for col in vcolors]
fig = px.line(y=param_squared_norms, x=losses_df.index)
fig.data[0].line.color = vcolors[0]
fig.layout.xaxis.title = 'Epoch'
fig.layout.yaxis.title = r'$||w||_2^2$'
fig.update_layout(base_layout)
fig.update_layout(width=160, height=160, margin = dict(t=0,b=20,l=20,r=0),
legend=dict(
orientation="h",
font = dict(size=8),
x=0,
y=1.35,
bgcolor="rgba(0,0,0,0)",
bordercolor="Black",
borderwidth=0,
title = dict(side = 'top')
)
)
fig.update_xaxes(tickangle=45)
fig.update_layout(width=400, height=300)
fig
# -
# ## Generating Figure 4c
# The figure that shows loss values per interaction interval for NODEC.
# +
fig = px.line(losses_melted, x='Epoch', y='Total Loss', color='Interaction Interval', log_y=True,
color_discrete_sequence=vcolors, render_mode='svg')
fig.update_layout(base_layout)
#fig.data[0].line.dash = 'dot'
#fig.data[1].line.dash = 'dot'
fig.data[2].line.dash = 'dot'
fig.update_yaxes(exponentformat='power')
fig.update_layout(width=160, height=195, margin = dict(t=35,b=0,l=0,r=0),
legend=dict(
orientation="h",
font = dict(size=8),
x=0,
y=1.45,
bgcolor="rgba(0,0,0,0)",
bordercolor="Black",
borderwidth=0,
title = dict(side = 'top')
)
)
fig.layout.yaxis.tickfont = dict(size=9)
fig.layout.yaxis.nticks = 7
fig.layout.yaxis.tickmode = 'auto'
fig.layout.yaxis.exponentformat = 'SI'
fig.update_layout(width=400, height=300)
fig
|
nodec_experiments/ct_lti/single_sample/figure_4.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # ULMFit
# Fine-tuning a forward and backward langauge model to get to 95.4% accuracy on the IMDB movie reviews dataset. This tutorial is done with fastai v1.0.53.
from fastai.text import *
# ## From a language model...
# ### Data collection
# This was run on a Titan RTX (24 GB of RAM) so you will probably need to adjust the batch size accordinly. If you divide it by 2, don't forget to divide the learning rate by 2 as well in the following cells. You can also reduce a little bit the bptt to gain a bit of memory.
bs,bptt=256,80
# This will download and untar the file containing the IMDB dataset, returning a `Pathlib` object pointing to the directory it's in (default is ~/.fastai/data/imdb0). You can specify another folder with the `dest` argument.
path = untar_data(URLs.IMDB)
# We then gather the data we will use to fine-tune the language model using the [data block API](https://fastai1.fast.ai/data_block.html). For this step, we want all the texts available (even the ones that don't have lables in the unsup folder) and we won't use the IMDB validation set (we will do this for the classification part later only). Instead, we set aside a random 10% of all the texts to build our validation set.
#
# The fastai library will automatically launch the tokenization process with the [spacy tokenizer](https://spacy.io/api/tokenizer/) and a few [default rules](https://fastai1.fast.ai/text.transform.html#Rules) for pre and post-processing before numericalizing the tokens, with a vocab of maximum size 60,000. Tokens are sorted by their frequency and only the 60,000 most commom are kept, the other ones being replace by an unkown token. This cell takes a few minutes to run, so we save the result.
data_lm = (TextList.from_folder(path)
#Inputs: all the text files in path
.filter_by_folder(include=['train', 'test', 'unsup'])
#We may have other temp folders that contain text files so we only keep what's in train, test and unsup
.split_by_rand_pct(0.1)
#We randomly split and keep 10% (10,000 reviews) for validation
.label_for_lm()
#We want to do a language model so we label accordingly
.databunch(bs=bs, bptt=bptt))
data_lm.save('data_lm.pkl')
# When restarting the notebook, as long as the previous cell was executed once, you can skip it and directly load your data again with the following.
data_lm = load_data(path, 'data_lm.pkl', bs=bs, bptt=bptt)
# Since we are training a language model, all the texts are concatenated together (with a random shuffle between them at each new epoch). The model is trained to guess what the next word in the sentence is.
data_lm.show_batch()
# For a backward model, the only difference is we'll have to pqss the flag `backwards=True`.
data_bwd = load_data(path, 'data_lm.pkl', bs=bs, bptt=bptt, backwards=True)
data_bwd.show_batch()
# ### Fine-tuning the forward language model
# The idea behind the [ULMFit paper](https://arxiv.org/abs/1801.06146) is to use transfer learning for this classification task. Our language model isn't randomly initialized but with the weights of a model pretrained on a larger corpus, [Wikitext 103](https://blog.einstein.ai/the-wikitext-long-term-dependency-language-modeling-dataset/). The vocabulary of the two datasets are slightly different, so when loading the weights, we take care to put the embedding weights at the right place, and we rando;ly initiliaze the embeddings for words in the IMDB vocabulary that weren't in the wikitext-103 vocabulary of our pretrained model.
#
# This is all done by the first line of code that will download the pretrained model for you at the first use. The second line is to use [Mixed Precision Training](), which enables us to use a higher batch size by training part of our model in FP16 precision, and also speeds up training by a factor 2 to 3 on modern GPUs.
learn = language_model_learner(data_lm, AWD_LSTM)
learn = learn.to_fp16(clip=0.1)
# The `Learner` object we get is frozen by default, which means we only train the embeddings at first (since some of them are random).
learn.fit_one_cycle(1, 2e-2, moms=(0.8,0.7), wd=0.1)
# Then we unfreeze the model and fine-tune the whole thing.
learn.unfreeze()
learn.fit_one_cycle(10, 2e-3, moms=(0.8,0.7), wd=0.1)
# Once done, we jsut save the encoder of the model (everything except the last linear layer that was decoding our final hidden states to words) because this is what we will use for the classifier.
learn.save_encoder('fwd_enc')
# ### The same but backwards
# You can't directly train a bidirectional RNN for language modeling, but you can always enseble a forward and backward model. fastai provides a pretrained forward and backawrd model, so we can repeat the previous step to fine-tune the pretrained backward model. The command `language_model_learner` checks the `data` object you pass to automatically decide if it should use the pretrained forward or backward model.
learn = language_model_learner(data_bwd, AWD_LSTM)
learn = learn.to_fp16(clip=0.1)
# Then the training is the same:
learn.fit_one_cycle(1, 2e-2, moms=(0.8,0.7), wd=0.1)
learn.unfreeze()
learn.fit_one_cycle(10, 2e-3, moms=(0.8,0.7), wd=0.1)
learn.save_encoder('bwd_enc')
# ## ... to a classifier
# ### Data Collection
# The classifier is a model that is a bit heavier, so we have lower the batch size.
path = untar_data(URLs.IMDB)
bs = 128
# We use the data block API again to gather all the texts for classification. This time, we only keep the ones in the trainind and validation folderm and label then by the folder they are in. Since this step takes a bit of time, we save the result.
# +
data_clas = (TextList.from_folder(path, vocab=data_lm.vocab)
#grab all the text files in path
.split_by_folder(valid='test')
#split by train and valid folder (that only keeps 'train' and 'test' so no need to filter)
.label_from_folder(classes=['neg', 'pos'])
#label them all with their folders
.databunch(bs=bs))
data_clas.save('data_clas.pkl')
# -
# As long as the previous cell was executed once, you can skip it and directly do this.
data_clas = load_data(path, 'data_clas.pkl', bs=bs)
data_clas.show_batch()
# Like before, you only have to add `backwards=True` to load the data for a backward model.
data_clas_bwd = load_data(path, 'data_clas.pkl', bs=bs, backwards=True)
data_clas_bwd.show_batch()
# ### Fine-tuning the forward classifier
# The classifier needs a little less dropout, so we pass `drop_mult=0.5` to multiply all the dropouts by this amount (it's easier than adjusting all the five different values manually). We don't load the pretrained model, but instead our fine-tuned encoder from the previous section.
learn = text_classifier_learner(data_clas, AWD_LSTM, drop_mult=0.5, pretrained=False)
learn.load_encoder('fwd_enc')
# Then we train the model using gradual unfreezing (partially training the model from everything but the classification head frozen to the whole model trianing by unfreezing one layer at a time) and differential learning rate (deeper layer gets a lower learning rate).
lr = 1e-1
learn.fit_one_cycle(1, lr, moms=(0.8,0.7), wd=0.1)
learn.freeze_to(-2)
lr /= 2
learn.fit_one_cycle(1, slice(lr/(2.6**4),lr), moms=(0.8,0.7), wd=0.1)
learn.freeze_to(-3)
lr /= 2
learn.fit_one_cycle(1, slice(lr/(2.6**4),lr), moms=(0.8,0.7), wd=0.1)
learn.unfreeze()
lr /= 5
learn.fit_one_cycle(2, slice(lr/(2.6**4),lr), moms=(0.8,0.7), wd=0.1)
learn.save('fwd_clas')
# ### The same but backwards
# Then we do the same thing for the backward model, the only thigns to adjust are the names of the data object and the fine-tuned encoder we load.
learn_bwd = text_classifier_learner(data_clas_bwd, AWD_LSTM, drop_mult=0.5, pretrained=False)
learn_bwd.load_encoder('bwd_enc')
learn_bwd.fit_one_cycle(1, lr, moms=(0.8,0.7), wd=0.1)
learn_bwd.freeze_to(-2)
lr /= 2
learn_bwd.fit_one_cycle(1, slice(lr/(2.6**4),lr), moms=(0.8,0.7), wd=0.1)
learn_bwd.freeze_to(-3)
lr /= 2
learn_bwd.fit_one_cycle(1, slice(lr/(2.6**4),lr), moms=(0.8,0.7), wd=0.1)
learn_bwd.unfreeze()
lr /= 5
learn_bwd.fit_one_cycle(2, slice(lr/(2.6**4),lr), moms=(0.8,0.7), wd=0.1)
learn_bwd.save('bwd_clas')
# ### Ensembling the two models
# For our final results, we'll take the average of the predictions of the forward and the backward models. SInce the samples are sorted by text lengths for batching, we pass the argument `ordered=True` to get the predictions in the order of the texts.
pred_fwd,lbl_fwd = learn.get_preds(ordered=True)
pred_bwd,lbl_bwd = learn_bwd.get_preds(ordered=True)
final_pred = (pred_fwd+pred_bwd)/2
accuracy(pred, lbl_fwd)
# And we get the 95.4% accuracy reported in the paper!
|
examples/ULMFit.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
# encoding: UTF-8
"""
本文件中包含了自动交易系统中的一些基础设置、类和常量等。
"""
from __future__ import division
import hashlib
import os
from datetime import datetime, timedelta, time
import pandas as pd
import numpy as np
from pymongo import MongoClient
from pymongo.errors import ConnectionFailure
from collections import OrderedDict
import copy
from vnpy.trader.app.ctaStrategy.ctaBase import loadContractDetail
#定义一些常量
DIRECTION_LONG = u'多'
#----------------------------------------
def get_file_md5(file_path):
"""计算文件MD5值"""
f = open(file_path, 'rb')
md5_obj = hashlib.md5()
while True:
d = f.read(8096)
if not d:
break
md5_obj.update(d)
hashcode = md5_obj.hexdigest()
f.close()
return hashcode
##############################################################################
class VtTradeData(object):
"""成交数据类"""
def __init__(self, series):
self.gatewayName = series.loc['gatewayName']
self.rawData = series.loc['rawData']
#代码编号相关
self.symbol = series.loc['symbol']
self.exchange = series.loc['exchange']
self.vtSymbol = series.loc['vtSymbol']
self.tradeID = series.loc['tradeID']
self.vtTradeID = series.loc['vtTradeID']
self.orderID = series.loc['orderID']
self.vtOrderID = series.loc['vtOrderID']
#成交相关
self.direction = series.loc['direction']
self.offset = series.loc['offset']
self.price = series.loc['price']
self.volume = series.loc['volume']
self.tradeTime = series.loc['tradeTime']
self.datetime = series.loc['datetime']
self.dt = series.loc['dt']
self.trade_date = series.loc['trade_date']
####################################################################################
class TradingResult(object):
"""每笔交易的结果"""
def __init__(self, entryPrice, entryDt, exitPrice,
exitDt, volume, rate, slippage, size):
"""Constructor"""
self.entryPrice = entryPrice # 开仓价格
self.exitPrice = exitPrice # 平仓价格
self.entryDt = entryDt # 开仓时间datetime
self.exitDt = exitDt # 平仓时间
self.volume = volume # 交易数量(+/-代表方向)
self.turnover = (self.entryPrice+self.exitPrice)*size*abs(volume) # 成交金额
self.commission = self.turnover*rate # 手续费成本
self.slippage = slippage*2*size*abs(volume) # 滑点成本
self.pnl = ((self.exitPrice - self.entryPrice) * volume * size
- self.commission - self.slippage) # 净盈亏
########################################################################################
class StrategyDBProcessor(object):
"""从数据库中获取策略的不同信息"""
def __init__(self):
self.host = 'localhost'
self.port = 27017
#------------------------------------------------------------------------------------
def get_data(self, db_name, tb_name, order):
"""依据order从数据库提取数据"""
#连接数据库
try:
db_client = MongoClient(self.host, self.port,
connectTimeoutMS=500)
#调用server_info查询服务器状态,防止服务器异常并未连接成功
db_client.server_info()
except:
print(u'Mongodb连接失败')
return
db = db_client[db_name]
collection = db[tb_name]
cursor = collection.find(order)
data = list(cursor)
result = pd.DataFrame(data)
if not result.empty:
del result['_id']
#断开数据库连接
db_client.close()
return result
#--------------------------------------------------------------------------------------
def date_format_trans(self, start_date, end_date):
"""将str格式转化为datetime格式"""
start_date = pd.Timestamp(start_date)
if end_date is None:
end_date = start_date
else:
end_date = pd.Timestamp(end_date)
#为了获取的数据包括end_date这一天
if end_date.time() == time(0, 0):
end_date = end_date + timedelta(hours = 23) + timedelta(minutes=59)
return start_date, end_date
#----------------------------------------------------------------------------------------
def get_strategy_information(self, strategy_name, start_date, end_date=None):
"""从mongodb中获取策略在一段时间内的账户详情"""
#调整日期格式
start_date, end_date = self.date_format_trans(start_date, end_date)
d = {'account_datetime':{'$gte': start_date, '$lte': end_date}}
db_name = 'Realtime_Strategy_Information'
result = self.get_data(db_name, strategy_name, d)
#如果是空列表返回一个空表
if result.empty:
return result
result.set_index('account_datetime', inplace=True)
result.sort_index(ascending=True, inplace=True)
result['deposit_ratio'] = result['deposit'] / result['capital'] * 100
result['pnl'] = result['capital'] - result['original_capital']
result['pnl_ratio'] = result['pnl'] / result['original_capital'] * 100
#记录开平仓时间点,0:仓位无变化;1:开多;2:开空;3:平多;4:平空
pos_change_signal = [0]
index = result.index
for i in range(1, len(index)):
if result.loc[index[i], 'pos'] == result.loc[index[i-1], 'pos']:
pos_change_signal.append(0)
elif (result.loc[index[i], 'pos'] > result.loc[index[i-1], 'pos'] and
result.loc[index[i], 'pos_long'] > result.loc[index[i-1], 'pos_long']):
pos_change_signal.append(1)
elif (result.loc[index[i], 'pos'] < result.loc[index[i-1], 'pos'] and
result.loc[index[i], 'pos_short'] > result.loc[index[i-1], 'pos_short']):
pos_change_signal.append(2)
elif (result.loc[index[i], 'pos'] < result.loc[index[i-1], 'pos'] and
result.loc[index[i], 'pos_long'] < result.loc[index[i-1], 'pos_long']):
pos_change_signal.append(3)
elif (result.loc[index[i], 'pos'] > result.loc[index[i-1], 'pos'] and
result.loc[index[i], 'pos_short'] < result.loc[index[i-1], 'pos_short']):
pos_change_signal.append(4)
result['pos_change_signal'] = pos_change_signal
pos_change_signal_dict = {0: 'not', 1: 'long',
2: u'short', 3: u'sell', 4: u'cover'}
trade_signal = [pos_change_signal_dict[i] for i in pos_change_signal]
result['trade_signal'] = trade_signal
return result
#----------------------------------------------------------------------------------
def get_trade_result(self, strategy_name, start_date, end_date):
"""统计一段时间策略的交易表现"""
start_date, end_date = self.date_format_trans(start_date, end_date)
db_name = 'TradeRecord'
d = {'datetime':{'$gte': start_date, '$lte':end_date}}
result = self.get_data(db_name, strategy_name, d)
#如果是空列表返回一个空表
if result.empty:
return result
result.sort_values(by='datetime', ascending=True, inplace=True)
trade_dict = OrderedDict()
for i in range(len(result)):
trade_dict[i] = VtTradeData(result.iloc[i,:])
#获取最后交易日价格
strategy_data = self.get_strategy_information(strategy_name, start_date,
end_date)
#如果是空列表返回一个空表
if strategy_data.empty:
return strategy_data
last_price = strategy_data.iloc[-1,:].loc['last_price']
last_date = strategy_data.index[-1]
trade_result = self.calculate_trade(trade_dict, last_price, last_date)
#计算夏普率
date = [i.date() for i in strategy_data.index]
strategy_data['trade_date'] = date
new_date = pd.Series(date).drop_duplicates()
daily_pnl = pd.Series([])
#每日盈亏额
for i in range(len(new_date)):
d_2 = new_date.iloc[i]
if i == 0:
sd_1 = strategy_data[strategy_data['trade_date']==d_2].iloc[0,:].loc['pnl']
sd_2 = strategy_data[strategy_data['trade_date']==d_2].iloc[-1,:].loc['pnl']
else:
d_1 = new_date.iloc[i-1]
sd_1 = strategy_data[strategy_data['trade_date']==d_1].iloc[-1,:].loc['pnl']
sd_2 = strategy_data[strategy_data['trade_date']==d_2].iloc[-1,:].loc['pnl']
daily_pnl[d_2] = sd_2 - sd_1
trade_result['daily_pnl'] = daily_pnl
pnl_std = daily_pnl.std()
pnl_mean = daily_pnl.mean()
sharp_ratio = pnl_mean / pnl_std * np.sqrt(240)
trade_result['sharp_ratop'] = sharp_ratio
trade_reault['name'] = strategy_name
return trade_result
#----------------------------------------------------------------------------------------
def calculate_trade(self, trade_dict, last_price, last_date):
"""
计算回测结果
"""
# 检查成交记录
if not trade_dict:
print(u'成交记录为空,无法计算回测结果')
return {}
#载入合约参数
symbol = trade_dict[0].symbol
con_d = loadContractDetail(symbol)
size = con_d['trade_size']
price_tick = con_d['price_tick']
slip = 0
slippage = slip * price_tick #默认滑点为0个跳价
rate = 0.3/10000 #默认手续费万分之0.3
# 首先基于回测后的成交记录,计算每笔交易的盈亏
resultList = [] # 交易结果列表
longTrade = [] # 未平仓的多头交易
shortTrade = [] # 未平仓的空头交易
tradeTimeList = [] # 每笔成交时间戳
posList = [0] # 每笔成交后的持仓情况
for trade in trade_dict.values():
# 复制成交对象,因为下面的开平仓交易配对涉及到对成交数量的修改
# 若不进行复制直接操作,则计算完后所有成交的数量会变成0
trade = copy.copy(trade)
# 多头交易
if trade.direction == DIRECTION_LONG:
# 如果尚无空头交易
if not shortTrade:
longTrade.append(trade)
# 当前多头交易为平空
else:
while True:
entryTrade = shortTrade[0]
exitTrade = trade
# 清算开平仓交易
closedVolume = min(exitTrade.volume, entryTrade.volume)
result = TradingResult(entryTrade.price, entryTrade.dt,
exitTrade.price, exitTrade.dt,
-closedVolume, rate, slippage, size)
resultList.append(result)
posList.extend([-1,0])
tradeTimeList.extend([result.entryDt, result.exitDt])
# 计算未清算部分
entryTrade.volume -= closedVolume
exitTrade.volume -= closedVolume
# 如果开仓交易已经全部清算,则从列表中移除
if not entryTrade.volume:
shortTrade.pop(0)
# 如果平仓交易已经全部清算,则退出循环
if not exitTrade.volume:
break
# 如果平仓交易未全部清算,
if exitTrade.volume:
# 且开仓交易已经全部清算完,则平仓交易剩余的部分
# 等于新的反向开仓交易,添加到队列中
if not shortTrade:
longTrade.append(exitTrade)
break
# 如果开仓交易还有剩余,则进入下一轮循环
else:
pass
# 空头交易
else:
# 如果尚无多头交易
if not longTrade:
shortTrade.append(trade)
# 当前空头交易为平多
else:
while True:
entryTrade = longTrade[0]
exitTrade = trade
# 清算开平仓交易
closedVolume = min(exitTrade.volume, entryTrade.volume)
result = TradingResult(entryTrade.price, entryTrade.dt,
exitTrade.price, exitTrade.dt,
closedVolume, rate, slippage, size)
resultList.append(result)
posList.extend([1,0])
tradeTimeList.extend([result.entryDt, result.exitDt])
# 计算未清算部分
entryTrade.volume -= closedVolume
exitTrade.volume -= closedVolume
# 如果开仓交易已经全部清算,则从列表中移除
if not entryTrade.volume:
longTrade.pop(0)
# 如果平仓交易已经全部清算,则退出循环
if not exitTrade.volume:
break
# 如果平仓交易未全部清算,
if exitTrade.volume:
# 且开仓交易已经全部清算完,则平仓交易剩余的部分
# 等于新的反向开仓交易,添加到队列中
if not longTrade:
shortTrade.append(exitTrade)
break
# 如果开仓交易还有剩余,则进入下一轮循环
else:
pass
# 到最后交易日尚未平仓的交易,则以最后价格平仓
endPrice = last_price
for trade in longTrade:
result = TradingResult(trade.price, trade.dt, endPrice, last_date,
trade.volume, rate, slippage, size)
resultList.append(result)
for trade in shortTrade:
result = TradingResult(trade.price, trade.dt, endPrice, last_date,
-trade.volume, rate, slippage, size)
resultList.append(result)
# 检查是否有交易
if not resultList:
print(u'无交易结果')
return {}
# 然后基于每笔交易的结果,我们可以计算具体的盈亏曲线和最大回撤等
capital = 0 # 资金
maxCapital = 0 # 资金最高净值
drawdown = 0 # 回撤
totalResult = 0 # 总成交数量
totalTurnover = 0 # 总成交金额(合约面值)
totalCommission = 0 # 总手续费
totalSlippage = 0 # 总滑点
timeList = [] # 时间序列
pnlList = [] # 每笔盈亏序列
capitalList = [] # 盈亏汇总的时间序列
drawdownList = [] # 回撤的时间序列
winningResult = 0 # 盈利次数
losingResult = 0 # 亏损次数
totalWinning = 0 # 总盈利金额
totalLosing = 0 # 总亏损金额
for result in resultList:
capital += result.pnl
maxCapital = max(capital, maxCapital)
drawdown = capital - maxCapital
pnlList.append(result.pnl)
timeList.append(result.exitDt) # 交易的时间戳使用平仓时间
capitalList.append(capital)
drawdownList.append(drawdown)
totalResult += 1
totalTurnover += result.turnover
totalCommission += result.commission
totalSlippage += result.slippage
if result.pnl >= 0:
winningResult += 1
totalWinning += result.pnl
else:
losingResult += 1
totalLosing += result.pnl
# 计算盈亏相关数据
winningRate = winningResult/totalResult*100 # 胜率
averageWinning = 0 # 这里把数据都初始化为0
averageLosing = 0
profitLossRatio = 0
if winningResult:
averageWinning = totalWinning/winningResult # 平均每笔盈利
if losingResult:
averageLosing = totalLosing/losingResult # 平均每笔亏损
if averageLosing:
profitLossRatio = -averageWinning/averageLosing # 盈亏比
# 返回回测结果
d = {}
d['capital'] = capital
d['maxCapital'] = maxCapital
d['drawdown'] = drawdown
d['totalResult'] = totalResult
d['totalTurnover'] = totalTurnover
d['totalCommission'] = totalCommission
d['totalSlippage'] = totalSlippage
d['timeList'] = timeList
d['pnlList'] = pnlList
d['capitalList'] = capitalList
d['drawdownList'] = drawdownList
d['winningRate'] = winningRate
d['averageWinning'] = averageWinning
d['averageLosing'] = averageLosing
d['profitLossRatio'] = profitLossRatio
d['posList'] = posList
d['tradeTimeList'] = tradeTimeList
d['resultList'] = resultList
trade_result = pd.Series([])
trade_result[u'first_deal'] = d['timeList'][0]
trade_result[u'last_deal'] = d['timeList'][-1]
trade_result[u'number_of_deals'] = d['totalResult']
trade_result[u'total_pnl'] = d['capital']
trade_result[u'maximun_withdrawal'] =min(d['drawdownList'])
trade_result[u'average_profit_per_deal'] = d['capital']/d['totalResult']
trade_result[u'average_slippage_per_deal'] = d['totalSlippage']/d['totalResult']
trade_result[u'average_commission_per_deal'] = d['totalCommission']/d['totalResult']
trade_result[u'win_rate'] = d['winningRate']
trade_result[u'average_winning'] = d['averageWinning']
trade_result[u'average_losing'] = d['averageLosing']
trade_result[u'profit_loss_ratio'] = d['profitLossRatio']
return trade_result
# +
import sys
sys.path.append('/home/freeman/Desktop/auto_trade_system/lib')
from ats_base import StrategyDBProcessor
import time
start = time.time()
sdbp = StrategyDBProcessor()
dat = sdbp.get_strategy_information('TrendTunnelStrategy_FG_60min', '20190211', '20190219')
data = sdbp.get_trade_result('TrendTunnelStrategy_FG_60min', '20190211', '20190219')
print(time.time() - start)
# -
dict(dat)
print dict(data)
|
auto_trade_system/lib/ats_base.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.5 64-bit (''origin'': conda)'
# language: python
# name: python395jvsc74a57bd04e62c5f72907caf2f8e50c4e1b95548d48fade11eacc296cd4faa51b1d77304e
# ---
# +
import sys
import numpy as np
import matplotlib.pyplot as plt
import empymod
sys.path.append('../')
import emulatte.forward as fwd
# +
# VMD
emsrc_name = 'VMD'
# 層厚
thicks = [100] #空気層+2層
# 比抵抗
res = [2e14, 10, 100] #空気層+2層
# 送信座標
sc = [0, 0, 0]
# 受信座標
rc = [50, 0, 0]
# 測定周波数
freqs = np.logspace(-2, 5, 500)
# 地下の物性値
props = {'res' : res}
# 物理空間モデルの呼び出し
model = fwd.model(thicks)
# 物性値の割り当て
model.set_properties(**props)
# 送信ソースの呼び出しと設定
emsrc = fwd.transmitter(emsrc_name, freqs, moment=1)
# 送信ソースの設置と受信座標の設定
model.locate(emsrc, sc, rc)
# 電磁応答の取得
EMF = model.emulate(hankel_filter='werthmuller201')
# 成分抽出
hz = EMF['h_z']
re = hz.real
im = hz.imag
# 描画
fig = plt.figure(figsize=(8,5), facecolor='w')
ax = fig.add_subplot(111)
ax.plot(freqs, re, "C0-", label='real')
ax.plot(freqs, -re, "C0--")
ax.plot(freqs, im, "C1-", label='imaginary')
ax.plot(freqs, -im, "C1--")
ax.grid(which='major', c='#ccc')
ax.grid(which='minor', c='#eee')
plt.tick_params(which='both', direction='in')
ax.set_yscale('log')
ax.set_xscale('log')
ax.set_xlabel('Frequency [Hz]')
ax.set_ylabel('$H_z$ [A/m]')
ax.set_xlim(1e-2, 1e5)
ax.set_title(emsrc_name+' Frequency Domain')
ax.legend()
plt.savefig(emsrc_name+'_FD.png')
# +
# VMD
emsrc_name = 'HMDx'
# 層厚
thicks = [100] #空気層+2層
# 比抵抗
res = [2e14, 10, 100] #空気層+2層
# 送信座標
sc = [0, 0, 0]
# 受信座標
rc = [50, 0, 0]
# 測定周波数
freqs = np.logspace(-2, 5, 500)
# 地下の物性値
props = {'res' : res}
# 物理空間モデルの呼び出し
model = fwd.model(thicks)
# 物性値の割り当て
model.set_properties(**props)
# 送信ソースの呼び出しと設定
emsrc = fwd.transmitter(emsrc_name, freqs, moment=1)
# 送信ソースの設置と受信座標の設定
model.locate(emsrc, sc, rc)
# 電磁応答の取得
EMF = model.emulate(hankel_filter='werthmuller201')
# 成分抽出
hz = EMF['h_z']
re = hz.real
im = hz.imag
# 描画
fig = plt.figure(figsize=(8,5), facecolor='w')
ax = fig.add_subplot(111)
ax.plot(freqs, re, "C0-", label='real')
ax.plot(freqs, -re, "C0--")
ax.plot(freqs, im, "C1-", label='imaginary')
ax.plot(freqs, -im, "C1--")
ax.grid(which='major', c='#ccc')
ax.grid(which='minor', c='#eee')
plt.tick_params(which='both', direction='in')
ax.set_yscale('log')
ax.set_xscale('log')
ax.set_xlabel('Frequency [Hz]')
ax.set_ylabel('$H_z$ [A/m]')
ax.set_xlim(1e-2, 1e5)
ax.set_title(emsrc_name+' Frequency Domain')
ax.legend()
plt.savefig(emsrc_name+'_FD.png')
# -
|
test/FDALL.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: is310_env
# language: python
# name: is310_env
# ---
# How do we read in our film scripts dataset? How would we sort the values in the dataset? How could we remove all the null values from the dataset? How we group our data by year?
import pandas as pd
film_scripts = pd.read_csv('pudding_film_scripts.csv')
film_scripts[0:1] #indexing rows
film_scripts.sort_values(by=['title'])
film_scripts = film_scripts.rename(columns={'gross (inflation-adjusted)': 'gross_ia'})
# +
# for index, row in film_scripts[0:10].iterrows():
# print(row['gross_ia'])
# if row['gross_ia'] >= 0:
# print('is number')
# -
len(film_scripts)
subset_film_scripts = film_scripts[film_scripts.gross_ia.notna()]
films_year = subset_film_scripts.groupby('year')
films_year.get_group(1931)
subset_film_scripts.nlargest(columns=['gross_ia'], n=5)
subset_film_scripts[['gross_ia']].plot()
film_scripts.fillna(0, inplace=True)
film_scripts.groupby('year').size()
films_year = film_scripts.groupby('year').size().reset_index(name='counts')
films_year
film_scripts_cleaned = film_scripts
film_scripts_cleaned.groupby('year')['gross_ia'].max().reset_index()
film_scripts[film_scripts.year == 2015]['gross_ia'].sum()
film_scripts.duplicated().any()
film_scripts
test_df = pd.DataFrame({'gender': ['m','Male','fem.','FemalE','Femle']})
test_df
test_df['gender'].map({
'm':'male', 'Male':'male', 'fem.':'female'
})
import re
test_df.gender[test_df['gender'].str.match(r"m", flags=re.IGNORECASE)] = 'male'
test_df.gender[test_df['gender'].str.match(r"f", flags=re.IGNORECASE)] = 'female'
test_df
film_scripts_cleaned
films_gross_sum = film_scripts_cleaned.groupby('year')['gross_ia'].sum().reset_index()
films_gross_sum
films_gross_sum.shape, film_scripts_cleaned.shape
films_gross_sum = films_gross_sum.rename(columns={'gross_ia': 'gross_sum'})
films_gross_sum.columns, film_scripts_cleaned.columns
merged_films = subset_film_scripts.merge(films_gross_sum, how='outer', on=['year'])
merged_films
merged_films.plot.scatter(x='gross_ia', y='gross_sum')
|
assets/files/Intro_EDA.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="eI3CZoyA45oh" outputId="67c6c1de-81f8-42b5-b32a-752c91495225"
# !pip install datasets transformers
# !pip install onnxruntime onnxruntime_tools
# + id="24Vd0apE5eUR"
from transformers import AutoTokenizer, AutoModelForQuestionAnswering
import torch
import numpy as np
from onnxruntime import ExecutionMode, InferenceSession, SessionOptions
tokenizer = AutoTokenizer.from_pretrained("distilbert-base-cased-distilled-squad")
model = AutoModelForQuestionAnswering.from_pretrained("distilbert-base-cased-distilled-squad")
# + id="FajWYrKDejN6"
question = "When did <NAME> died??"
context = """On 17 December 2011, <NAME> died from a heart attack. His youngest son <NAME> was announced as his successor.[92] In the face of international condemnation, North Korea continued to develop its nuclear arsenal, possibly including a hydrogen bomb and a missile capable of reaching the United States.[93]
Throughout 2017, following Donald Trump's assumption of the US presidency, tensions between the United States and North Korea increased, and there was heightened rhetoric between the two, with Trump threatening "fire and fury"[94] and North Korea threatening to test missiles that would land near Guam.[95] The tensions substantially decreased in 2018, and a détente developed.[96] A series of summits took place between Kim Jong-un of North Korea, President Moon Jae-in of South Korea, and President Trump.[97] It has been 3 years, 3 months since North Korea's last ICBM test."""
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="BORUmEiq5h93" outputId="753a2606-2655-43a5-a120-f4111a156d94"
inputs = tokenizer.encode_plus(question, context, return_tensors="pt")
answer = model(**inputs)
answer_start = torch.argmax(answer[0])
answer_end = torch.argmax(answer[1]) + 1
output = tokenizer.convert_tokens_to_string(tokenizer.convert_ids_to_tokens(inputs["input_ids"][0][answer_start:answer_end]))
output
# + id="kERVNHB0bSCj"
ids = inputs["input_ids"]
mask = inputs["attention_mask"]
# + colab={"base_uri": "https://localhost:8080/"} id="aAVbAcki5pCe" outputId="9cfefe8d-899e-40ff-8024-7daa85f9e2da"
torch.onnx.export(model,
(ids, mask),
"john_model.onnx",
input_names=["input_ids", "attention_mask"],
output_names=["output"],
dynamic_axes = {
"input_ids": {0: "batch_size"},
"attention_mask": {0: "batch_size"},
"output": {0: "batch_size"}
}
)
# Handles all the above steps for you
# + id="bBTagD-r5s8G" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="a790eef0-5568-42f9-c6e1-bd01c9ae552a"
session = InferenceSession("john_model.onnx")
tokens = tokenizer.encode_plus(question, context, return_tensors="np")
inputs_onnx = {"input_ids":tokens["input_ids"], "attention_mask":tokens["attention_mask"]}
output = session.run(None, inputs_onnx)
answer_start = np.argmax(output[0]) # get the most likely beginning of answer with the argmax of the score
answer_end = np.argmax(output[1]) + 1
output = tokenizer.convert_tokens_to_string(tokenizer.convert_ids_to_tokens(inputs["input_ids"][0][answer_start:answer_end]))
output
|
John/prepare_john.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# Basic Interactor Demo
# ---------------------
#
# This demo shows off an interactive visualization using [Bokeh](https://bokeh.pydata.org) for plotting, and Ipython interactors for widgets. The demo runs entirely inside the Ipython notebook, with no Bokeh server required.
#
# The dropdown offers a choice of trig functions to plot, and the sliders control the frequency, amplitude, and phase.
#
# To run, click on, `Cell->Run All` in the top menu, then scroll to the bottom and move the sliders.
# +
from ipywidgets import interact
import numpy as np
from bokeh.io import push_notebook, show, output_notebook
from bokeh.plotting import figure
output_notebook()
# -
x = np.linspace(0, 2*np.pi, 2000)
y = np.sin(x)
p = figure(title="simple line example", plot_height=300, plot_width=600, y_range=(-5,5))
r = p.line(x, y, color="#2222aa", line_width=3)
def update(f, w=1, A=1, phi=0):
if f == "sin": func = np.sin
elif f == "cos": func = np.cos
elif f == "tan": func = np.tan
r.data_source.data['y'] = A * func(w * x + phi)
push_notebook()
show(p, notebook_handle=True)
interact(update, f=["sin", "cos", "tan"], w=(0,100), A=(1,5), phi=(0, 20, 0.1))
|
_posts/JupyterInteractors.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
from scipy import stats
import matplotlib.pyplot as plt
import arviz as az
np.random.seed(42)
# # Visualization
#
# Data visualization is a very broad area with graphical representations targeting very particular audiences ranging from a scientific paper in some not-over-hyped subject to newspapers with million readers. We will focus on scientific visualizations and in particular visualizations useful in a Bayesian setting.
#
# As humans are generally good at visualizing data, data visualization is both a powerful tool for analyzing data and models and is also a powerful tool to convey information to our target audience. Using words, tables and single numbers are generally less effective ways to communicate information. At the same time our visual system can be fooled, as you may have experienced by being tricked by visual illusions. The reason is that our visual system is tuned to process information in useful ways and this generally means not just seeing the information, but *interpreting* it as well. Put less formally, our brains _guess stuff_ and don't just _reproduce the outside world_. Effective data visualization requires that we recognize the abilities and limitations of our own visual systems.
# ## Plot elements
#
# To convey visual information we generally use shapes, including lines, circles, squares etc. These elements have properties associated to them like, position, shape and color.
#
# ArviZ is build on top of matplotlib, thus is a good idea to get familiar with the names of the elements that are used by matplotlib to create a plot.
#
# ![]()
#
# <a href="https://matplotlib.org/3.1.1/gallery/showcase/anatomy.html"><img src="https://matplotlib.org/_images/anatomy.png"></a>
# ## Colors
#
# Matplotlib allows user to easily switch between plotting styles by defining style sheets. ArviZ is delivered with a few additional styles that can be applied globally by writing `az.style.use(nameofstyle)` or locally using a `with` statement as in the following example:
# +
x = np.linspace(0, 1, 100)
dist = stats.beta(2, 5).pdf(x)
fig = plt.figure()
with az.style.context('arviz-colors'):
for i in range(10):
plt.plot(x, dist - i, f'C{i}', label=f'C{i}')
plt.xlabel('x')
plt.ylabel('f(x)', rotation=0, labelpad=15);
# -
# `az.style` is just an alias of `matplotlib.pyplot.style`, so everything you can do with one of them you can do with the other.
#
# All styles included with ArviZ use the same color-blind friendly palette. This palette was designed using https://colorcyclepicker.mpetroff.net/. If you need to do plots in grey-scale we recommend to restrict yourself to the first 3 colors of the arviz default palette ('C0', 'C1' and 'C2'), otherwise you may need to use different [line styles](https://matplotlib.org/api/_as_gen/matplotlib.lines.Line2D.html#matplotlib.lines.Line2D.set_linestyle) or [different markers](https://matplotlib.org/api/markers_api.html#module-matplotlib.markers)
from matplotlib import lines
print(lines.lineStyles)
from matplotlib import markers
print(markers.MarkerStyle.markers)
# ## Continuous and discrete distributions
#
# A discrete distribution represents variables which can only take a countable number of values. Some examples of discrete random variables are the number of coins in your pocket, spots on a giraffe, red cars in a city, people with flu etc. As we generally use integers to represent discrete variables, when ArviZ is asked to plot integer data it will use [histograms](https://en.wikipedia.org/wiki/Histogram) to represent them. ArviZ always tries to associate the binned data with discrete values. For example in the following plot each _bar_ is associated with an integer in the interval [0, 9].
d_values = stats.poisson(3).rvs(500)
az.plot_dist(d_values);
# A continuous distribution represents variables taking an uncountable number of values. Some examples of continuous random variables are the temperature during summer, the blood pressure of a patience, the time needed to finish a task, etc. By default ArviZ uses kernel density estimation (KDE) to represent continuous distributions.
c_values = stats.gamma(2, 3).rvs(500)
az.plot_dist(c_values);
# Kernel density estimation (KDE) is a non-parametric way to estimate the probability density function of a random variable.
#
# Conceptually you place a _kernel function_ like a gaussian _on top_ of a data-point, then you sum all the gaussians, generally evaluated over a grid and not over the data points. Results are normalized so the total area under the curve is one.
#
# The following block of code shows a very simple example of a KDE.
# +
_, ax = plt.subplots(figsize=(12, 4))
bw = 0.4
np.random.seed(19)
datapoints = 7
y = np.random.normal(7, size=datapoints)
x = np.linspace(y.min() - bw * 3, y.max() + bw * 3, 100)
kernels = np.transpose([stats.norm.pdf(x, i, bw) for i in y])
kernels *= 1/datapoints # normalize the results
ax.plot(x, kernels, 'k--', alpha=0.5)
ax.plot(y, np.zeros(len(y)), 'C1o')
ax.plot(x, kernels.sum(1))
ax.set_xticks([])
ax.set_yticks([]);
# -
# Compared to other KDEs in the Python ecosystem, the KDE implemented in ArviZ automatically handles the boundaries of a distribution. Basically, ArviZ will assign a density of zero to any point outside the range of the data. Another nice feature of ArviZ's KDE is the method it uses to estimate the _bandwith_. The bandwidth of a kernel density estimator is a parameter that controls its degree of smoothness. ArviZ's method works well for a wide range of distributions including multimodal ones. The following plot compares the KDEs for ArviZ (on the left) and SciPy (on the right). The blue line is the theoretical distribution, the light blue bars give a histogram computed from samples drawn from the distribution, and the orange lines are the kernel density estimations.
# +
def scipykdeplot(data, ax, **kwargs):
x = np.linspace(data.min(), data.max(), len(data))
kde = stats.gaussian_kde(data)
density = kde.evaluate(x)
ax.plot(x, density, **kwargs)
size = 1000
bw = 4.5 # ArviZ's default value
_, ax = plt.subplots(5, 2, figsize=(15, 10), constrained_layout=True)
a_dist = stats.vonmises(loc=np.pi, kappa=20)
b_dist = stats.beta(a=2, b=5)
c_dist = [stats.norm(-8, 0.75), stats.norm(8, 1)]
d_dist = stats.norm(0, 1)
e_dist = stats.uniform(-1, 1)
a = a_dist.rvs(size)
a = np.arctan2(np.sin(a), np.cos(a))
b = b_dist.rvs(size)
c = np.concatenate((c_dist[0].rvs(7000), c_dist[1].rvs(3000)))
d = d_dist.rvs(size)
e = e_dist.rvs(size)
ax[0, 0].set_title('ArviZ')
ax[0, 1].set_title('Scipy')
for idx, (i, dist) in enumerate(zip([d, a, c, b, e], [d_dist, a_dist, c_dist, b_dist, e_dist] )):
x = np.linspace(i.min()+0.01, i.max()-0.01, 200)
if idx == 2:
x_dist = np.concatenate((dist[0].pdf(x[:100]) * 0.7, dist[1].pdf(x[100:]) * 0.3))
else:
x_dist = dist.pdf(x)
ax[idx, 0].plot(x, x_dist, 'C0', lw=2)
az.plot_kde(i, ax=ax[idx, 0], bw=bw, textsize=11, plot_kwargs={'color':'C1', 'linewidth':2})
ax[idx, 0].set_yticks([])
ax[idx, 0].hist(i, bins='auto', alpha=0.2, density=True)
ax[idx, 1].plot(x, x_dist, 'C0', lw=2)
scipykdeplot(i, ax=ax[idx, 1], color='C1', lw=2)
ax[idx, 1].set_yticks([])
ax[idx, 1].hist(i, bins='auto', alpha=0.2, density=True)
|
content/Section_01/Visualization.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import json
import pandas as pd
import numpy as np
from pathlib import Path
data_classes = ['yelp', 'reddit', 'stackexchange']
# +
numberOfUsers = []
numberOfReviews = []
for data_class in data_classes:
if data_class == 'yelp':
data_dir = Path.cwd() / 'datasets/yelp/'
elif data_class == 'reddit':
data_dir = Path.cwd() / 'datasets/reddit/'
elif data_class == 'stackexchange':
data_dir = Path.cwd() / 'datasets/stackexchange/'
else:
print('Class not found.')
disclosed_gender_df = pd.read_csv(data_dir / 'disclosed_dataset.csv')
undisclosed_gender_df = pd.read_csv(data_dir / 'undisclosed_dataset.csv')
if data_class == 'yelp':
numberOfUsers.append(disclosed_gender_df.user_id.unique().shape[0] + undisclosed_gender_df.user_id.unique().shape[0])
numberOfReviews.append(disclosed_gender_df.shape[0] + undisclosed_gender_df.shape[0])
elif data_class == 'reddit':
numberOfUsers.append(disclosed_gender_df.UserName.unique().shape[0] + undisclosed_gender_df.UserName.unique().shape[0])
numberOfReviews.append(disclosed_gender_df.shape[0] + undisclosed_gender_df.shape[0])
elif data_class == 'stackexchange':
numberOfUsers.append(disclosed_gender_df.UserId.unique().shape[0] + undisclosed_gender_df.UserId.unique().shape[0])
numberOfReviews.append(disclosed_gender_df.shape[0] + undisclosed_gender_df.shape[0])
else:
print('Class not found.')
# +
import numpy as np
import matplotlib.pyplot as plt
N = 3
numberOfUsers = (yelp_data[0],reddit_data[0], stack_data[0])
# men_std = (2, 3, 4, 1, 2)
ind = np.arange(N) # the x locations for the groups
width = 0.35 # the width of the bars
fig, ax = plt.subplots()
rects1 = ax.bar(ind, numberOfUsers, width, color='g')
numberOfReviews = (yelp_data[1],reddit_data[1], stack_data[1])
# women_std = (3, 5, 2, 3, 3)
rects2 = ax.bar(ind + width, numberOfReviews, width, color='y')
# add some text for labels, title and axes ticks
ax.set_ylabel('Amount')
ax.set_title('The number of Users and Reviews')
ax.set_xticks(ind + width / 2)
ax.set_xticklabels(('Yelp', 'Reddit', 'StackOverflow'))
ax.legend((rects1[0], rects2[0]), ('#Users', '#Reviews'))
def autolabel(rects):
"""
Attach a text label above each bar displaying its height
"""
for rect in rects:
height = rect.get_height()
ax.text(rect.get_x() + rect.get_width()/2., height,
'%d' % int(height),
ha='center', va='bottom')
autolabel(rects1)
autolabel(rects2)
plt.show()
# plt.savefig("dataset_statistic.png", dpi=300)
# -
|
StatisticUsersReviews.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Heat Equation
# ## The Differential Equation
# $$ \frac{\partial u}{\partial t} = -\frac{1}{16}\frac{\partial^2 u}{\partial x^2}$$
# ## Initial Condition
# $$ u(x,0)=1-x-\frac{1}{\pi}\sin(2\pi x) $$
#
# ## Boundary Condition
# $$ u(0,t)=1, u(1,t)=0 $$
#
# ## The Difference Equation
# $$ w[k+1,i] = w[k,i] + \frac{-1}{16}\frac{k}{h^2}(w[k,i+1]-2w[k,i]+w[k,i-1])$$
#
# +
# LIBRARY
# vector manipulation
import numpy as np
# math functions
import math
# THIS IS FOR PLOTTING
# %matplotlib inline
import matplotlib.pyplot as plt # side-stepping mpl backend
import warnings
warnings.filterwarnings("ignore")
# +
N=40
Nt=15
h=1/N
ht=1/10000
time=np.arange(0,1.0001,ht)
x=np.arange(0,1.0001,h)
w=np.zeros((Nt+1,N+1))
Solution=np.zeros((Nt+1,N+1))
A=np.zeros((N-1,N-1))
c=np.zeros(N-1)
b=np.zeros(N-1)
b[0]=1
# Initial Condition
for i in range (1,N):
w[0,i]=1-x[i]-1/np.pi*np.sin(2*np.pi*x[i])
Solution[0,i]=1-x[i]-1/np.pi*np.sin(2*np.pi*x[i])
# Boundary Condition
for k in range (0,Nt+1):
w[k,0]=1
w[k,N]=0
Solution[k,0]=1
Solution[k,N]=0
for i in range (0,N-1):
A[i,i]=-2
for i in range (0,N-2):
A[i+1,i]=1
A[i,i+1]=1
A=np.eye(N-1)+ht/(h*h)*(A)
fig = plt.figure(figsize=(8,4))
plt.matshow(A)
print(Solution.shape)
print(w.shape)
for k in range (1,Nt+1):
w[k,1:(N)]=np.dot(A,w[k-1,1:(N)])+ht/(h*h)*b
Solution[k,1:(N)]=1-x[1:N]-1/np.pi*np.sin(2*np.pi*x[1:N])
#print(np.dot(A,c))
fig = plt.figure(figsize=(8,4))
plt.matshow(w)
plt.matshow(Solution)
# -
|
Chapter 08 - Heat Equations/ParabolicEquation/Parabolic - ForwardTimeCenteredSpace Example 3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.3 64-bit (''base'': conda)'
# metadata:
# interpreter:
# hash: ead85409a3df2736fa4852a03cf3afe7e5b1dbdc0e40d7d22bbcbf8cf1adf5fa
# name: python3
# ---
# # Python in Action
# ## Part 1: Python Fundamentals
# ### 09 — User input
# > accepting user input in Python
#
# Python is well-known for being extremely useful as a scripting language.
#
# You can display information on the standard output using `print()` and you can get information from the user using `input()`:
print('About to delete resource')
print('Are you sure you want to continue?')
user_input = input()
print('You typed ' + user_input)
|
01-python-basics/09-python-user-input.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19"
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load in
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
# Input data files are available in the "../input/" directory.
# For example, running this (by clicking run or pressing Shift+Enter) will list the files in the input directory
import os
print(os.listdir("../input"))
# Any results you write to the current directory are saved as output.
# + _uuid="5363c3301fe77aeac413f57975e1759c5a6ae125"
import numpy as np
import lightgbm as lgb
import pandas as pd
from kaggle.competitions import twosigmanews
import matplotlib.pyplot as plt
import random
from datetime import datetime, date
from xgboost import XGBClassifier
from sklearn import model_selection
from sklearn.metrics import mean_squared_error
import time
# + _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a"
# official way to get the data
from kaggle.competitions import twosigmanews
env = twosigmanews.make_env()
print('Done!')
# + _uuid="079b1d2474f911283fc1b04e552f421af763fdad"
(market_train_df, news_train_df) = env.get_training_data()
# + _uuid="0a5f2fbd071df4c2a86b408fd1e8a719757ae4e6"
market_train_df['time'] = market_train_df['time'].dt.date
market_train_df = market_train_df.loc[market_train_df['time']>=date(2010, 1, 1)]
# + _uuid="ae10d6a84619658ee14c3ed8c0ed25717e55f0f5"
from multiprocessing import Pool
def create_lag(df_code,n_lag=[3,7,14,],shift_size=1):
code = df_code['assetCode'].unique()
for col in return_features:
for window in n_lag:
rolled = df_code[col].shift(shift_size).rolling(window=window)
lag_mean = rolled.mean()
lag_max = rolled.max()
lag_min = rolled.min()
lag_std = rolled.std()
df_code['%s_lag_%s_mean'%(col,window)] = lag_mean
df_code['%s_lag_%s_max'%(col,window)] = lag_max
df_code['%s_lag_%s_min'%(col,window)] = lag_min
# df_code['%s_lag_%s_std'%(col,window)] = lag_std
return df_code.fillna(-1)
def generate_lag_features(df,n_lag = [3,7,14]):
features = ['time', 'assetCode', 'assetName', 'volume', 'close', 'open',
'returnsClosePrevRaw1', 'returnsOpenPrevRaw1',
'returnsClosePrevMktres1', 'returnsOpenPrevMktres1',
'returnsClosePrevRaw10', 'returnsOpenPrevRaw10',
'returnsClosePrevMktres10', 'returnsOpenPrevMktres10',
'returnsOpenNextMktres10', 'universe']
assetCodes = df['assetCode'].unique()
print(assetCodes)
all_df = []
df_codes = df.groupby('assetCode')
df_codes = [df_code[1][['time','assetCode']+return_features] for df_code in df_codes]
print('total %s df'%len(df_codes))
pool = Pool(4)
all_df = pool.map(create_lag, df_codes)
new_df = pd.concat(all_df)
new_df.drop(return_features,axis=1,inplace=True)
pool.close()
return new_df
# + _uuid="8bd4c3ddba90f4b23e324eefabb98272689ac0ce"
# return_features = ['close']
# new_df = generate_lag_features(market_train_df,n_lag = 5)
# market_train_df = pd.merge(market_train_df,new_df,how='left',on=['time','assetCode'])
# + _uuid="d80bb484853e216761cf2e7e08c0e9b093c08505"
return_features = ['returnsClosePrevMktres10','returnsClosePrevRaw10','open','close']
n_lag = [3,7,14]
new_df = generate_lag_features(market_train_df,n_lag=n_lag)
market_train_df = pd.merge(market_train_df,new_df,how='left',on=['time','assetCode'])
# + _uuid="8b67955bcf92b534e101be7b8b6aa697869fb7db"
print(market_train_df.columns)
# + _uuid="cc53bf017273aa8c59a0f2dabf80c70eeb20108b"
# return_features = ['open']
# new_df = generate_lag_features(market_train_df,n_lag=[3,7,14])
# market_train_df = pd.merge(market_train_df,new_df,how='left',on=['time','assetCode'])
# + _uuid="4681d35d83bf94c0b9e95904124902ef2153e3bc"
def mis_impute(data):
for i in data.columns:
if data[i].dtype == "object":
data[i] = data[i].fillna("other")
elif (data[i].dtype == "int64" or data[i].dtype == "float64"):
data[i] = data[i].fillna(data[i].mean())
else:
pass
return data
market_train_df = mis_impute(market_train_df)
# + _uuid="3b9aa2e280d60a66c006af63f78700861ef90698"
def data_prep(market_train):
lbl = {k: v for v, k in enumerate(market_train['assetCode'].unique())}
market_train['assetCodeT'] = market_train['assetCode'].map(lbl)
market_train = market_train.dropna(axis=0)
return market_train
market_train_df = data_prep(market_train_df)
# # check the shape
print(market_train_df.shape)
# + _uuid="f947d36b6b6c986fcba9a7ccb4a71fd5e58021d0"
from sklearn.preprocessing import LabelEncoder
up = market_train_df['returnsOpenNextMktres10'] >= 0
universe = market_train_df['universe'].values
d = market_train_df['time']
fcol = [c for c in market_train_df if c not in ['assetCode', 'assetCodes', 'assetCodesLen', 'assetName', 'audiences',
'firstCreated', 'headline', 'headlineTag', 'marketCommentary', 'provider',
'returnsOpenNextMktres10', 'sourceId', 'subjects', 'time', 'time_x', 'universe','sourceTimestamp']]
X = market_train_df[fcol].values
up = up.values
r = market_train_df.returnsOpenNextMktres10.values
# Scaling of X values
# It is good to keep these scaling values for later
mins = np.min(X, axis=0)
maxs = np.max(X, axis=0)
rng = maxs - mins
X = 1 - ((maxs - X) / rng)
# Sanity check
assert X.shape[0] == up.shape[0] == r.shape[0]
from xgboost import XGBClassifier
from sklearn import model_selection
from sklearn.metrics import mean_squared_error
import time
# X_train, X_test, up_train, up_test, r_train, r_test,u_train,u_test,d_train,d_test = model_selection.train_test_split(X, up, r,universe,d, test_size=0.25, random_state=99)
te = market_train_df['time']>date(2015, 1, 1)
tt = 0
for tt,i in enumerate(te.values):
if i:
idx = tt
print(i,tt)
break
print(idx)
# for ind_tr, ind_te in tscv.split(X):
# print(ind_tr)
X_train, X_test = X[:idx],X[idx:]
up_train, up_test = up[:idx],up[idx:]
r_train, r_test = r[:idx],r[idx:]
u_train,u_test = universe[:idx],universe[idx:]
d_train,d_test = d[:idx],d[idx:]
train_data = lgb.Dataset(X_train, label=up_train.astype(int))
# train_data = lgb.Dataset(X, label=up.astype(int))
test_data = lgb.Dataset(X_test, label=up_test.astype(int))
# + _uuid="d0f9657c977e8556be91af02d04b84afd4fb460f"
# these are tuned params I found
x_1 = [0.19000424246380565, 2452, 212, 328, 202]
x_2 = [0.19016805202090095, 2583, 213, 312, 220]
print(up_train)
def exp_loss(p,y):
y = y.get_label()
# p = p.get_label()
grad = -y*(1.0-1.0/(1.0+np.exp(-y*p)))
hess = -(np.exp(y*p)*(y*p-1)-1)/((np.exp(y*p)+1)**2)
return grad,hess
params_1 = {
'task': 'train',
'boosting_type': 'gbdt',
'objective': 'binary',
# 'objective': 'regression',
'learning_rate': x_1[0],
'num_leaves': x_1[1],
'min_data_in_leaf': x_1[2],
# 'num_iteration': x_1[3],
'num_iteration': 239,
'max_bin': x_1[4],
'verbose': 1
}
params_2 = {
'task': 'train',
'boosting_type': 'gbdt',
'objective': 'binary',
# 'objective': 'regression',
'learning_rate': x_2[0],
'num_leaves': x_2[1],
'min_data_in_leaf': x_2[2],
# 'num_iteration': x_2[3],
'num_iteration': 172,
'max_bin': x_2[4],
'verbose': 1
}
gbm_1 = lgb.train(params_1,
train_data,
num_boost_round=100,
valid_sets=test_data,
early_stopping_rounds=5,
# fobj=exp_loss,
)
gbm_2 = lgb.train(params_2,
train_data,
num_boost_round=100,
valid_sets=test_data,
early_stopping_rounds=5,
# fobj=exp_loss,
)
# + _uuid="57f90489c498f068d2dc0a0bd7e8e8c138899ae0"
confidence_test = (gbm_1.predict(X_test) + gbm_2.predict(X_test))/2
confidence_test = (confidence_test-confidence_test.min())/(confidence_test.max()-confidence_test.min())
confidence_test = confidence_test*2-1
print(max(confidence_test),min(confidence_test))
# calculation of actual metric that is used to calculate final score
r_test = r_test.clip(-1,1) # get rid of outliers. Where do they come from??
x_t_i = confidence_test * r_test * u_test
data = {'day' : d_test, 'x_t_i' : x_t_i}
df = pd.DataFrame(data)
x_t = df.groupby('day').sum().values.flatten()
mean = np.mean(x_t)
std = np.std(x_t)
score_test = mean / std
print(score_test)
# + _uuid="4cd0e0e8c5724f660e3095c6e2a3e8459afc7d6c"
import gc
del X_train,X_test
gc.collect()
# + _uuid="8ba5794894665dfc9ee492c60268bf073ca18c05"
# #prediction
# days = env.get_prediction_days()
# n_days = 0
# prep_time = 0
# prediction_time = 0
# packaging_time = 0
# total_market_obs_df = []
# for (market_obs_df, news_obs_df, predictions_template_df) in days:
# n_days +=1
# if (n_days%50==0):
# print(n_days,end=' ')
# t = time.time()
# market_obs_df['time'] = market_obs_df['time'].dt.date
# return_features = ['returnsClosePrevMktres10','returnsClosePrevRaw10','open','close']
# total_market_obs_df.append(market_obs_df)
# if len(total_market_obs_df)==1:
# history_df = total_market_obs_df[0]
# else:
# history_df = pd.concat(total_market_obs_df[-(np.max(n_lag)+1):])
# print(history_df)
# new_df = generate_lag_features(history_df,n_lag=[3,7,14])
# market_obs_df = pd.merge(market_obs_df,new_df,how='left',on=['time','assetCode'])
# # return_features = ['open']
# # new_df = generate_lag_features(market_obs_df,n_lag=[3,7,14])
# # market_obs_df = pd.merge(market_obs_df,new_df,how='left',on=['time','assetCode'])
# market_obs_df = mis_impute(market_obs_df)
# market_obs_df = data_prep(market_obs_df)
# # market_obs_df = market_obs_df[market_obs_df.assetCode.isin(predictions_template_df.assetCode)]
# X_live = market_obs_df[fcol].values
# X_live = 1 - ((maxs - X_live) / rng)
# prep_time += time.time() - t
# t = time.time()
# lp = (gbm_1.predict(X_live) + gbm_2.predict(X_live))/2
# prediction_time += time.time() -t
# t = time.time()
# confidence = lp
# confidence = (confidence-confidence.min())/(confidence.max()-confidence.min())
# confidence = confidence * 2 - 1
# preds = pd.DataFrame({'assetCode':market_obs_df['assetCode'],'confidence':confidence})
# predictions_template_df = predictions_template_df.merge(preds,how='left').drop('confidenceValue',axis=1).fillna(0).rename(columns={'confidence':'confidenceValue'})
# env.predict(predictions_template_df)
# packaging_time += time.time() - t
# env.write_submission_file()
# sub = pd.read_csv("submission.csv")
|
3 mu sigma using news to predict news/eda-script-67.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python2
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/gurher/Pandas/blob/main/Pandas_Practice/4_Students_Alcohol_Consumption_Exercises.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="TGTEqeAYHSzi"
# # Student Alcohol Consumption
# + [markdown] id="IefCDkHWHSzl"
# ### Introduction:
#
# This time you will download a dataset from the UCI.
#
# ### Step 1. Import the necessary libraries
# + id="4b75r1jXHSzl"
import pandas as pd
# + [markdown] id="hG1OvXCfHSzm"
# ### Step 2. Import the dataset from this [address](https://raw.githubusercontent.com/guipsamora/pandas_exercises/master/04_Apply/Students_Alcohol_Consumption/student-mat.csv).
# + [markdown] id="sc-gzwCOHSzm"
# ### Step 3. Assign it to a variable called df.
# + colab={"base_uri": "https://localhost:8080/"} id="QViYx1VrHSzm" outputId="5a472c3a-05dc-4d9f-c467-5abc56a2b3d3"
url = 'https://raw.githubusercontent.com/guipsamora/pandas_exercises/master/04_Apply/Students_Alcohol_Consumption/student-mat.csv'
df = pd.read_csv(url)
df.head()
df.columns.tolist()
# + [markdown] id="t7F_CQ4GHSzn"
# ### Step 4. For the purpose of this exercise slice the dataframe from 'school' until the 'guardian' column
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="NDgIKXfqHSzn" outputId="888056b7-52c3-435f-ad0b-8b1ef95896d8"
df.loc[:,'school':'guardian'].head()
# + [markdown] id="F65DH4DEHSzn"
# ### Step 5. Create a lambda function that capitalize strings.
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="6NILrLPFHSzn" outputId="72151dea-f098-4650-b018-e8461ca00c34"
capitalizer = lambda x: x.capitalize() # capitalize only very first letter
capitalize = lambda x: x.upper()
# + [markdown] id="ymRfeivPHSzn"
# ### Step 6. Capitalize both Mjob and Fjob
# + id="qzEOz7o7HSzo"
# df['Mjob'].apply(capitalize)
# df['Fjob'].apply(capitalize)
df['Fjob'].str.upper()
df['Mjob'].str.capitalize()
# + [markdown] id="nB16o_XjHSzo"
# ### Step 7. Print the last elements of the data set.
# + colab={"base_uri": "https://localhost:8080/", "height": 233} id="K0-JvIobHSzo" outputId="aeb93e5c-362f-403e-991a-ff34b94e81b7"
df.tail()
# + [markdown] id="D0FyCCTuHSzo"
# ### Step 8. Did you notice the original dataframe is still lowercase? Why is that? Fix it and capitalize Mjob and Fjob.
# + colab={"base_uri": "https://localhost:8080/", "height": 233} id="Cx3JBmGFHSzo" outputId="c318a247-6d1a-4e11-eec6-dbc0452f231a"
# df['Mjob'].apply(capitalize)
# df['Fjob'].apply(capitalize)
df['Fjob'] = df['Fjob'].str.upper()
df['Mjob'] = df['Mjob'].str.capitalize()
df.head()
# + [markdown] id="JjrHJPuzHSzp"
# ### Step 9. Create a function called majority that return a boolean value to a new column called legal_drinker (Consider majority as older than 17 years old)
# + id="PP3TzVX4HSzp"
def majority(age):
if age>17:
return True
else:
return False
# + id="xUV8E2apHSzp"
df['legal_drinker'] = df.age.apply(majority)
df.legal_drinker
# + [markdown] id="PoEDo3UoHSzp"
# ### Step 10. Multiply every number of the dataset by 10.
# ##### I know this makes no sense, don't forget it is just an exercise
# + colab={"base_uri": "https://localhost:8080/", "height": 233} id="YgQRsMgnHSzp" outputId="d36db2f5-69e0-4c4c-9662-978a7b311403"
df.dtypes
df.dtypes[df.dtypes=='int64'].keys().tolist()
integer = df.dtypes[df.dtypes=='int64'].index.tolist()
df.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="6LNci8QWHSzq" outputId="7a6f71c5-5407-4d7f-bcca-2b56046d51db"
df[integer] * 10
df
|
Python/Pandas_Practice/4_Students_Alcohol_Consumption_Exercises.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.6.10 64-bit (''Project8'': conda)'
# language: python
# name: python3
# ---
import os, csv
filepath = os.path.join("Resources","budget_data.csv")
total_months = []
net_total = []
monthly_change = []
with open(filepath, newline="", encoding='utf-8') as datafile:
csvreader = csv.reader(datafile, delimiter=",")
header = next(csvreader)
#Iterate through rows and save to lists
for row in csvreader:
total_months.append(row[0])
net_total.append(int(row[1]))
for i in range(len(net_total)-1):
monthly_change.append(net_total[i+1]-net_total[i])
#Find maxes
max_increase = max(monthly_change)
max_decrease = min(monthly_change)
#Month of max increase of profit and max decrease of profit
max_increase_month = total_months[monthly_change.index(max_increase) + 1]
max_decrease_month = total_months[monthly_change.index(max_decrease) + 1]
#Print Statement
print("Financial Analysis")
print("----------------------------")
print(f"Total Months: {len(total_months)}")
print(f"Total: ${sum(net_total)}")
print(f"Average Change: {round(sum(monthly_change)/len(monthly_change),2)}")
print(f"Greatest Increase in Profits: {max_increase_month} (${max_increase})")
print(f"Greatest Decrease in Profits: {max_decrease_month} (${max_decrease})")
# +
filepath = os.path.join("Resources", "financial_analysis_summary.txt")
with open(filepath, "w") as file:
file.write("Financial Analysis")
file.write("\n")
file.write("----------------------------")
file.write("\n")
file.write(f"Total Months: {len(total_months)}")
file.write("\n")
file.write(f"Total: ${sum(net_total)}")
file.write("\n")
file.write(f"Average Change: {round(sum(monthly_change)/len(monthly_change),2)}")
file.write("\n")
file.write(f"Greatest Increase in Profits: {max_increase_month} (${max_increase})")
file.write("\n")
file.write(f"Greatest Decrease in Profits: {max_decrease_month} (${max_decrease})")
# -
|
PyBank/main.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Using Pivoted DataFrames with Perspective
#
# Perspective tries to infer as much information as possible from already-pivoted dataframes:
import pandas as pd
import numpy as np
from perspective import Table, PerspectiveWidget
# For the dataset, we'll use `superstore.arrow` which is used in various Perspective demos:
# +
import requests
# Download the arrow
url = "https://unpkg.com/@jpmorganchase/perspective-examples@0.2.0-beta.2/build/superstore.arrow"
req = requests.get(url)
arrow = req.content
# -
# Create a table
table = Table(arrow)
view = table.view()
df = view.to_df()
display(df.shape, df.dtypes)
# ### Row Pivots
#
# Create a row pivoted dataframe, and pass it into `PerspectiveWidget`:
df_pivoted = df.set_index(['Country', 'Region'])
df_pivoted.head()
# Pivots will be read from the df and applied
PerspectiveWidget(df_pivoted)
# ### Column Pivots
#
# The same is true with column pivots:
# +
arrays = [np.array(['bar', 'bar', 'bar', 'bar', 'baz', 'baz', 'baz', 'baz', 'foo', 'foo', 'foo', 'foo', 'qux', 'qux', 'qux', 'qux']),
np.array(['one', 'one', 'two', 'two', 'one', 'one', 'two', 'two', 'one', 'one', 'two', 'two', 'one', 'one', 'two', 'two']),
np.array(['X', 'Y', 'X', 'Y', 'X', 'Y', 'X', 'Y', 'X', 'Y', 'X', 'Y', 'X', 'Y', 'X', 'Y'])]
tuples = list(zip(*arrays))
index = pd.MultiIndex.from_tuples(tuples, names=['first', 'second', 'third',])
df_col = pd.DataFrame(np.random.randn(3, 16), index=['A', 'B', 'C'], columns=index)
df_col
# -
# Pivots again automatically applied
PerspectiveWidget(df_col)
# ### Pivot Table (Row and Column Pivots)
pt = pd.pivot_table(df, values = 'Discount', index=['Country','Region'], columns = ['Category', 'Segment'])
pt
PerspectiveWidget(pt)
# ### More pivot examples
# +
arrays = {'A':['bar', 'bar', 'bar', 'bar', 'baz', 'baz', 'baz', 'baz', 'foo', 'foo', 'foo', 'foo', 'qux', 'qux', 'qux', 'qux'],
'B':['one', 'one', 'two', 'two', 'one', 'one', 'two', 'two', 'one', 'one', 'two', 'two', 'one', 'one', 'two', 'two'],
'C':['X', 'Y', 'X', 'Y', 'X', 'Y', 'X', 'Y', 'X', 'Y', 'X', 'Y', 'X', 'Y', 'X', 'Y'],
'D':np.arange(16)}
df2 = pd.DataFrame(arrays)
df2
# -
df2_pivot = df2.pivot_table(values=['D'], index=['A'], columns=['B','C'], aggfunc={'D':'count'})
df2_pivot
PerspectiveWidget(df2_pivot)
pt = pd.pivot_table(df, values = ['Discount','Sales'], index=['Country','Region'],aggfunc={'Discount':'count','Sales':'sum'},columns=["State","Quantity"])
pt
PerspectiveWidget(pt)
|
examples/jupyter-notebooks/pandas_pivot.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Calculation of Principal Moment of Inertia
#
# <hr>
# <NAME>.; <NAME>. Molecular Shape Diversity of Combinatorial Libraries: A Prerequisite for Broad Bioactivity. J. Chem. Inf. Comput. Sci. 2003, 43 (3), 987–1003. https://doi.org/10.1021/ci025599w.
#
# + init_cell=true
# %reload_ext autoreload
# %autoreload 2
# def warn(*args, **kwargs):
# warn pass # to silence scikit-learn warnings
import warnings
warnings.filterwarnings('ignore')
# warnings.warn = warn
# Global Imports
# from collections import Counter
# import glob
from pathlib import Path
import sys
import pandas as pd
import numpy as np
# import seaborn as sns
# from matplotlib import pyplot as plt
from rdkit import Chem
from rdkit.Chem import AllChem as Chem
from rdkit.Chem import Descriptors as Desc
from rdkit.Chem import rdMolDescriptors as rdMolDesc
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
# Project-local Imports
PROJECT_DIR = list(Path("..").absolute().parents)[1]
sys.path.append(str(PROJECT_DIR))
import plt_style
import utils as u
from utils import lp
# +
def gen_3d(mol):
mh = Chem.AddHs(mol)
Chem.EmbedMolecule(mh, Chem.ETKDG())
res = 10
ntries = -1
iters = [100, 300, 1000]
while res > 0 and ntries < 3:
ntries += 1
res = Chem.UFFOptimizeMolecule(mh, maxIters=iters[ntries])
return mh, res
def calc_pmi(inp, source, avg=3):
source = source.lower()
did_not_converge = 0
pmi1 = []
pmi2 = []
if isinstance(inp, str):
inp = [inp]
for i in inp:
mol = Chem.MolFromSmiles(i)
pmi1_avg = []
pmi2_avg = []
for _ in range(avg):
mol, res = gen_3d(mol)
did_not_converge += res
pmis = sorted([rdMolDesc.CalcPMI1(mol), rdMolDesc.CalcPMI2(mol), rdMolDesc.CalcPMI3(mol)])
pmi1_avg.append(pmis[0] / pmis[2])
pmi2_avg.append(pmis[1] / pmis[2])
pmi1.append(np.median(pmi1_avg))
pmi2.append(np.median(pmi2_avg))
print("* {} minimizations did not converge.".format(did_not_converge))
return pmi1, pmi2 # pmi1, pmi2 are lists
# -
# +
df = u.read_tsv("../Input Data/pmi_input.tsv")
smiles = list(df['SMILES'])
PMIk, PMIl = calc_pmi(smiles, 'smiles')
df['PMIx'] = PMIk
df['PMIy'] = PMIl
print(df.keys())
# -
u.write_tsv(df, "results/pmi_results.tsv")
|
6. PMI/.ipynb_checkpoints/calculate_pmi_data-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %reload_ext autoreload
# %autoreload 2
# %matplotlib inline
import kerr
import numpy as np
import matplotlib.pyplot as plt
# +
U = np.arange(-6, 6, 1)
delta = 28*U
gam = 0.1*U
eta = 0.1*U
f = 1/(U - eta*(1j))
g = 1/(U - eta*(1j))
c = (delta + (1j*gam)/2.)/(U - eta*(1j))
# -
f, g, c
system = kerr.Kerr(f[0], g[0], c[0])
print("Normalization", system.normalization())
x = y = np.arange(-6, 6, 0.1)
X, Y = np.meshgrid(x, y)
Z = system.wigner(X+(1j)*Y)
print("Wigner plot for f = {} \n g = {} \n c = {}".format(f[0], g[0], c[0]))
plt.contour(X, Y, Z)
plt.title("Wigner plot")
plt.ylabel("Real(z)")
plt.xlabel("Imag(z)")
plt.grid()
plt.colorbar()
plt.show()
|
doc/notebooks/wigner.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <font color='blue'>Data Science Academy</font>
# # <font color='blue'>Big Data Real-Time Analytics com Python e Spark</font>
#
# # <font color='blue'>Capítulo 7</font>
# ### *********** Atenção: ***********
# Utilize Java JDK 11 e Apache Spark 2.4.2
# *Caso receba mensagem de erro "name 'sc' is not defined", interrompa o pyspark e apague o diretório metastore_db no mesmo diretório onde está este Jupyter notebook*
# Acesse http://localhost:4040 sempre que quiser acompanhar a execução dos jobs
# # Transformações
# Criando uma lista em Python
lista1 = [124, 901, 652, 102, 397]
type(lista1)
# Carregando dados de uma coleção
lstRDD = sc.parallelize(lista1)
type(lstRDD)
lstRDD.collect()
lstRDD.count()
# Carregando um arquivo e criando um RDD.
autoDataRDD = sc.textFile("data/carros.csv")
type(autoDataRDD)
# Operação de Ação.
autoDataRDD.first()
autoDataRDD.take(5)
# Cada ação gera um novo processo de computação dos dados.
# Mas podemos persistir os dados em cache para que ele possa ser usado por outras ações, sem a necessidade
# de nova computação.
autoDataRDD.cache()
for line in autoDataRDD.collect():
print(line)
# Map() e criação de um novo RDD - Transformação - Lazy Evaluation
tsvData = autoDataRDD.map(lambda x : x.replace(",","\t"))
tsvData.take(5)
autoDataRDD.first()
# Filter() e criação de um novo RDD - Transformação - Lazy Evaluation
toyotaData = autoDataRDD.filter(lambda x: "toyota" in x)
# Ação
toyotaData.count()
# Ação
toyotaData.take(20)
# Pode salvar o conjunto de dados, o RDD.
# Nesse caso, o Spark solicita os dados ao processo Master e então gera um arquivo de saída.
savedRDD = open("data/carros_v2.csv","w")
savedRDD.write("\n".join(autoDataRDD.collect()))
savedRDD.close()
# ## Operações Set
# Set operations
palavras1 = sc.parallelize(["Big Data","Data Science","Analytics","Visualization"])
palavras2 = sc.parallelize(["Big Data","R","Python","Scala"])
# União
for unions in palavras1.union(palavras2).distinct().collect():
print(unions)
# Interseção
for intersects in palavras1.intersection(palavras2).collect():
print(intersects)
rdd01 = sc.parallelize(range(1,10))
rdd02 = sc.parallelize(range(10,21))
rdd01.union(rdd02).collect()
rdd01 = sc.parallelize(range(1,10))
rdd02 = sc.parallelize(range(5,15))
rdd01.intersection(rdd02).collect()
# ## Left/Right Outer Join
names1 = sc.parallelize(("banana", "uva", "laranja")).map(lambda a: (a, 1))
names2 = sc.parallelize(("laranja", "abacaxi", "manga")).map(lambda a: (a, 1))
names1.join(names2).collect()
names1.leftOuterJoin(names2).collect()
names1.rightOuterJoin(names2).collect()
# ## Distinct
# Distinct
lista1 = [124, 901, 652, 102, 397, 124, 901, 652]
lstRDD = sc.parallelize(lista1)
for numbData in lstRDD.distinct().collect():
print(numbData)
# ## Transformação e Limpeza
# RDD Original
autoDataRDD.collect()
# Transformação e Limpeza
def LimpaRDD(autoStr) :
# Verifica a indexação
if isinstance(autoStr, int) :
return autoStr
# Separa cada índice pela vírgula (separador de colunas)
attList = autoStr.split(",")
# Converte o número de portas para um num
if attList[3] == "two" :
attList[3] = "2"
elif attList[3] == "four":
attList[3] = "4"
# Convert o modelo do carro para uppercase
attList[5] = attList[4].upper()
return ",".join(attList)
# Transformação
RDD_limpo = autoDataRDD.map(LimpaRDD)
print(RDD_limpo)
# Ação
RDD_limpo.collect()
# ## Ações
# reduce() - soma de valores
lista2 = [124, 901, 652, 102, 397, 124, 901, 652]
lstRDD = sc.parallelize(lista2)
lstRDD.collect()
lstRDD.reduce(lambda x,y: x + y)
# Encontrando a linha com menor número de caracteres
autoDataRDD.reduce(lambda x,y: x if len(x) < len(y) else y)
# Criando uma função para redução
def getMPG(autoStr) :
if isinstance(autoStr, int) :
return autoStr
attList = autoStr.split(",")
if attList[9].isdigit() :
return int(attList[9])
else:
return 0
# Encontrando a média de MPG para todos os carros
media_mpg = round(autoDataRDD.reduce(lambda x,y : getMPG(x) + getMPG(y)) / (autoDataRDD.count() -1), 2)
print(media_mpg)
times = sc.parallelize(("Flamengo", "Vasco", "Botafogo", "Fluminense", "Palmeiras", "Bahia"))
times.takeSample(True, 3)
times = sc.parallelize(("Flamengo", "Vasco", "Botafogo", "Fluminense", "Palmeiras", "Bahia", "Bahia", "Flamengo"))
times.map(lambda k: (k,1)).countByKey().items()
autoDataRDD.saveAsTextFile("data/autoDataRDD.txt")
# # Fim
# ### Obrigado - Data Science Academy - <a href="http://facebook.com/dsacademybr">facebook.com/dsacademybr</a>
|
Apache Spark/03-Cap07-Spark-Operacoes.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Clase 1: Introducción al curso
#
# - IE0417: Diseño de Software para Ingeniería
# + [markdown] cell_style="split" slideshow={"slide_type": "slide"}
# ## Introducción Profesor
# - <NAME>
# - [LinkedIn](https://www.linkedin.com/in/esteban-zamora-a54484102/)
#
# #### Bach Ing. Eléctrica UCR (2014-2018)
# - Énfasis en Compus y Redes
# - PRIS-Lab: High Performance Computing (HPC)
#
# + [markdown] cell_style="split" slideshow={"slide_type": "fragment"}
# #### Embedded Software Engineer (2018+)
# - HPE Aruba Switching R&D, ASIC-SDK
# - *Senior Engineer & Tech Lead* (Costa Rica + Roseville, CA)
# - Kernel drivers y aplicaciones en Linux y RTOS (C/C++)
# - Herramientas de testing y observabilidad (Python, Go)
#
# + [markdown] cell_style="center" slideshow={"slide_type": "slide"}
# #### Introducción Estudiantes
# - Presentación
# - Nombre, Énfasis, etc
# - Qué sienten que pueden hacer con lo que saben de programación?
# - Interés y expectativas sobre el curso
#
# + [markdown] slideshow={"slide_type": "slide"}
# #### Profesor <-> Estudiantes
# - Comunicación asertiva
# - Retroalimentación continua
# + [markdown] cell_style="split" slideshow={"slide_type": "slide"}
# ### Qué Es / No Es?
# #### Es
# - Enfoque en sistemas embebidos
# - EIE > ECCI
# - IoT, Web-Backend, Edge-to-Cloud
# - Introducción al proceso y herramientas de desarrollo de software
# - Programación avanzada en C/C++ y Python (...Go?)
# - Spanglish :P
# + [markdown] cell_style="split" slideshow={"slide_type": "fragment"}
# <p> </p>
#
# #### No Es
# - Un laboratorio de programación libre
# - Enfoque en programación Web-Frontend o Movil
# - Un manual de *reglas de oro* de como desarrollar software
# + [markdown] slideshow={"slide_type": "slide"}
# #### Motivación
# - El software es un pilar del mundo moderno
# - Las vidas de las personas dependen del software
# - Buenos procesos de desarrollo y calidad son críticos => Ingeniería
#
# + [markdown] slideshow={"slide_type": "slide"}
# #### Que vamos a hacer en el curso?
# - Tomarnos el tiempo para aprender a hacer software para la vida real
# - No más correr para entregar un "*main*"
# - No más "*En mi compu sí servía*"
# - No más "*trabajo_final_v7.2_xyz-ahora_si-en_serio_x2.zip*"
# - Qué me hubiera servido conocer al empezar como Ing de Software Embebido?
# - Campo laboral para ingenieros con background de Ing Eléctrica (HW+SW)
# + [markdown] cell_style="split" slideshow={"slide_type": "slide"}
# ##### CI-0202, IE-0117, IE-0217
# 
# + [markdown] cell_style="split" slideshow={"slide_type": "slide"}
# ##### IE-0417
# 
# + [markdown] cell_style="split" slideshow={"slide_type": "fragment"}
# 
# + [markdown] cell_style="split" slideshow={"slide_type": "slide"}
# #### Fallas del hardware
# 
# + [markdown] cell_style="split" slideshow={"slide_type": "fragment"}
# #### Fallas del software
# 
# + [markdown] cell_style="split" slideshow={"slide_type": "slide"}
# #### Qué pasa cuando no se aplica bien la Ing. de software?
# 
# *Software rot*
# + [markdown] cell_style="split" slideshow={"slide_type": "fragment"}
# <p> </p>
# <p> </p>
#
# 
# + [markdown] slideshow={"slide_type": "slide"}
# #### Pero... El software no lo es todo
# - Proceso >> Código
# - Flujo de actividades
# - Dominio de aplicación
# - Personas >> Programas
# - Interacciones humanas
# - Cultura organizacional
# + [markdown] cell_style="split" slideshow={"slide_type": "slide"}
# 
# + [markdown] cell_style="split" slideshow={"slide_type": "fragment"}
# [Boeing 737 Max disaster](https://spectrum.ieee.org/how-the-boeing-737-max-disaster-looks-to-a-software-developer)
# + [markdown] cell_style="split" slideshow={"slide_type": "slide"}
#
# #### Detalles del curso
# - [Mediación Virtual](https://mv1.mediacionvirtual.ucr.ac.cr/course/view.php?id=26366)
# - [Carta al Estudiante](https://mv1.mediacionvirtual.ucr.ac.cr/pluginfile.php/2070064/mod_resource/content/1/Carta%20al%20estudiante.pdf)
# - [IE-0417 - GitHub](https://github.com/ezamoraa/ie0417)
# - [IE-0417 - Read The Docs](https://ie0417.readthedocs.io/es/latest/index.html)
#
# + [markdown] cell_style="split" slideshow={"slide_type": "fragment"}
# *Disclaimer*: Es la primera vez que se da el curso
#
# #### Desventajas
# - No hay feedback de semestres anteriores
# - Parte del material está en desarrollo
#
# #### Ventajas
# - Hay posibilidad de influir en detalles
# - Aprender algo que otras generaciones de EIE no han visto
# - Vivir al límite :)
# + [markdown] slideshow={"slide_type": "slide"}
# #### Reglas del curso
# - Se vale "copiar" código ... siempre y cuando se respeten las licencias de software
# - Los labs y el proyecto deben
# - Aplicar correctamente una licencia de software
# - Estar documentados con Sphinx, Doxygen para C/C++
# - Tabajarse en git + [github.com](https://github.com)
# - Demostrar que se desarrolló con git y branches, no solo subir "commits de entrega"
# + [markdown] slideshow={"slide_type": "slide"}
# #### Licencias de software libre: Por qué son importantes?
# - El software open source (OSS) está en todas partes
# - Tecnologías que damos por sentado
# - Software moderno se apalanca de OSS y da valor agregado
# - Son útiles para aprender a colaborar en proyectos open source
# + [markdown] slideshow={"slide_type": "slide"}
# 
# + [markdown] slideshow={"slide_type": "slide"}
# 
# + [markdown] slideshow={"slide_type": "slide"}
# #### Licencias de software: Qué son?
# - Instrumentos legales que definen como usar y distribuir el software
# - Otorgan permiso por adelantado sobre el uso del software
# - Empoderan al autor para decidir los detalles de como hacerlo
# - Reducen la ambiguedad al trabajar con software externo
#
# + [markdown] slideshow={"slide_type": "slide"}
# #### Copyright
# - Mecanismos de protección sobre los derechos de autoría sobre trabajo creativo
# - Derechos exclusivos para copiar, modificar, crear trabajos derivados y distribuir
#
# #### Copyright holder
# - Titular de derechos de autor
# - Persona o organización que tiene los derechos exclusivos sobre el uso de su trabajo y distribución
#
# + [markdown] slideshow={"slide_type": "slide"}
# #### Licencia Open Source
# - Cumplir con definición de OSI: [OSD](https://opensource.org/osd-annotated)
# - Permisivas vs Copyleft
# - Imponen más o menos requisitos
#
# #### Licencia no Open Source (propietaria)
# - Autor se reserva la mayoría de los derechos
# - Se imponen restricciones sobre el uso
# - Propietario != comercial (`$`)
#
# + [markdown] slideshow={"slide_type": "slide"}
# #### Open Source vs Software Libre
# - *Open Source* se refiere a las implicaciones prácticas de las licencias
# - Colaboración efectiva para el desarrollo de software
# - *Software Libre* se refiere a un conjunto de valores basado en 4 libertades
# - Ejecutar el programa como se desee, con cualquier propósito
# - Estudiar cómo funciona el programa, y cambiarlo para que haga lo que se desee
# - Redistribuir copias para ayudar a otros
# - Distribuir copias de sus versiones modificadas a terceros
# + [markdown] slideshow={"slide_type": "slide"}
#
# #### SPDX: Software Package Data Exchange
# - [Estandar](https://spdx.dev/specifications/) abierto para SBOM (Software Bill of Materials)
# - [Identificadores cortos de licencias](https://spdx.org/licenses/)
# ```c
# /* SPDX-License-Identifier: GPL-3.0-or-later */
# ```
#
# + [markdown] slideshow={"slide_type": "slide"}
# #### Consideraciones sobre licencias OSS
# - Si un proyecto puede ser descargado libremente pero no tiene licencia
# - No se puede asumir que es "open source"
# - Licencias open source vs patentes
# - Una patente le da al dueño derechos exclusivos sobre:
# - Construcción, el uso y la venta de una invención
# - Las licencias OSS pueden entrar en conflicto con las patentes
# - Las licencias OSS definen clausulas de patentes
# + [markdown] slideshow={"slide_type": "slide"}
# #### Recursos extra sobre licencias OSS
# - [Curso gratuito Linux Foundation](https://training.linuxfoundation.org/training/open-source-licensing-basics-for-software-developers/)
# - [Elige una licencia Open Source](https://choosealicense.com/)
# + [markdown] cell_style="center" slideshow={"slide_type": "slide"}
# ##### Quiz 1: Licencias OSS
# - Presentación el Lunes 4 de Abril
# - En grupos (2 o 3 personas)
# - Ver [licencias Open Source](https://opensource.org/licenses/category)
# - Ejemplos: Apache-2.0, MIT, BSD-3-Clause, GPL-2, GPL-3, MPL-2.0, CC licenses
#
# + [markdown] cell_style="center" slideshow={"slide_type": "slide"}
# ##### Quiz 1: Licencias OSS
# - Presentación debe responder las siguientes preguntas:
# - Cual es el tipo de licencia (permisiva vs copyleft)?
# - Cuales son los derechos y restricciones para uso, modificación y distribución?
# - Énfasis en cuales son consideradas las ventajas y casos de aplicación
# - Diferencias con licencias relacionadas (ejemplo: GPL-2 vs GPL-3)
# - Como aplicar esta licencia en un proyecto propio?
# - Archivo LICENSE, Copyright notice, etc
# - Como modificar un archivo existente en un proyecto que usa esta licencia?
# - Cual es un ejemplo de un proyecto conocido que la usa?
# - Impacto o implicaciones específicas de la licencia sobre el proyecto
# - Casos legales históricos relacionados a la licencia
|
presentations/001_intro_curso/slides.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Computer Vision Nanodegree
#
# ## Project: Image Captioning
#
# ---
#
# In this notebook, you will learn how to load and pre-process data from the [COCO dataset](http://cocodataset.org/#home). You will also design a CNN-RNN model for automatically generating image captions.
#
# Note that **any amendments that you make to this notebook will not be graded**. However, you will use the instructions provided in **Step 3** and **Step 4** to implement your own CNN encoder and RNN decoder by making amendments to the **models.py** file provided as part of this project. Your **models.py** file **will be graded**.
#
# Feel free to use the links below to navigate the notebook:
# - [Step 1](#step1): Explore the Data Loader
# - [Step 2](#step2): Use the Data Loader to Obtain Batches
# - [Step 3](#step3): Experiment with the CNN Encoder
# - [Step 4](#step4): Implement the RNN Decoder
# <a id='step1'></a>
# ## Step 1: Explore the Data Loader
#
# We have already written a [data loader](http://pytorch.org/docs/master/data.html#torch.utils.data.DataLoader) that you can use to load the COCO dataset in batches.
#
# In the code cell below, you will initialize the data loader by using the `get_loader` function in **data_loader.py**.
#
# > For this project, you are not permitted to change the **data_loader.py** file, which must be used as-is.
#
# The `get_loader` function takes as input a number of arguments that can be explored in **data_loader.py**. Take the time to explore these arguments now by opening **data_loader.py** in a new window. Most of the arguments must be left at their default values, and you are only allowed to amend the values of the arguments below:
# 1. **`transform`** - an [image transform](http://pytorch.org/docs/master/torchvision/transforms.html) specifying how to pre-process the images and convert them to PyTorch tensors before using them as input to the CNN encoder. For now, you are encouraged to keep the transform as provided in `transform_train`. You will have the opportunity later to choose your own image transform to pre-process the COCO images.
# 2. **`mode`** - one of `'train'` (loads the training data in batches) or `'test'` (for the test data). We will say that the data loader is in training or test mode, respectively. While following the instructions in this notebook, please keep the data loader in training mode by setting `mode='train'`.
# 3. **`batch_size`** - determines the batch size. When training the model, this is number of image-caption pairs used to amend the model weights in each training step.
# 4. **`vocab_threshold`** - the total number of times that a word must appear in the in the training captions before it is used as part of the vocabulary. Words that have fewer than `vocab_threshold` occurrences in the training captions are considered unknown words.
# 5. **`vocab_from_file`** - a Boolean that decides whether to load the vocabulary from file.
#
# We will describe the `vocab_threshold` and `vocab_from_file` arguments in more detail soon. For now, run the code cell below. Be patient - it may take a couple of minutes to run!
# +
import sys
sys.path.append('/opt/cocoapi/PythonAPI')
from pycocotools.coco import COCO
# !pip install nltk
import nltk
nltk.download('punkt')
from data_loader import get_loader
from torchvision import transforms
# Define a transform to pre-process the training images.
transform_train = transforms.Compose([
transforms.Resize(256), # smaller edge of image resized to 256
transforms.RandomCrop(224), # get 224x224 crop from random location
transforms.RandomHorizontalFlip(), # horizontally flip image with probability=0.5
transforms.ToTensor(), # convert the PIL Image to a tensor
transforms.Normalize((0.485, 0.456, 0.406), # normalize image for pre-trained model
(0.229, 0.224, 0.225))])
# Set the minimum word count threshold.
vocab_threshold = 5
# Specify the batch size.
batch_size = 10
# Obtain the data loader.
data_loader = get_loader(transform=transform_train,
mode='train',
batch_size=batch_size,
vocab_threshold=vocab_threshold,
vocab_from_file=False)
# -
# When you ran the code cell above, the data loader was stored in the variable `data_loader`.
#
# You can access the corresponding dataset as `data_loader.dataset`. This dataset is an instance of the `CoCoDataset` class in **data_loader.py**. If you are unfamiliar with data loaders and datasets, you are encouraged to review [this PyTorch tutorial](http://pytorch.org/tutorials/beginner/data_loading_tutorial.html).
#
# ### Exploring the `__getitem__` Method
#
# The `__getitem__` method in the `CoCoDataset` class determines how an image-caption pair is pre-processed before being incorporated into a batch. This is true for all `Dataset` classes in PyTorch; if this is unfamiliar to you, please review [the tutorial linked above](http://pytorch.org/tutorials/beginner/data_loading_tutorial.html).
#
# When the data loader is in training mode, this method begins by first obtaining the filename (`path`) of a training image and its corresponding caption (`caption`).
#
# #### Image Pre-Processing
#
# Image pre-processing is relatively straightforward (from the `__getitem__` method in the `CoCoDataset` class):
# ```python
# # Convert image to tensor and pre-process using transform
# image = Image.open(os.path.join(self.img_folder, path)).convert('RGB')
# image = self.transform(image)
# ```
# After loading the image in the training folder with name `path`, the image is pre-processed using the same transform (`transform_train`) that was supplied when instantiating the data loader.
#
# #### Caption Pre-Processing
#
# The captions also need to be pre-processed and prepped for training. In this example, for generating captions, we are aiming to create a model that predicts the next token of a sentence from previous tokens, so we turn the caption associated with any image into a list of tokenized words, before casting it to a PyTorch tensor that we can use to train the network.
#
# To understand in more detail how COCO captions are pre-processed, we'll first need to take a look at the `vocab` instance variable of the `CoCoDataset` class. The code snippet below is pulled from the `__init__` method of the `CoCoDataset` class:
# ```python
# def __init__(self, transform, mode, batch_size, vocab_threshold, vocab_file, start_word,
# end_word, unk_word, annotations_file, vocab_from_file, img_folder):
# ...
# self.vocab = Vocabulary(vocab_threshold, vocab_file, start_word,
# end_word, unk_word, annotations_file, vocab_from_file)
# ...
# ```
# From the code snippet above, you can see that `data_loader.dataset.vocab` is an instance of the `Vocabulary` class from **vocabulary.py**. Take the time now to verify this for yourself by looking at the full code in **data_loader.py**.
#
# We use this instance to pre-process the COCO captions (from the `__getitem__` method in the `CoCoDataset` class):
#
# ```python
# # Convert caption to tensor of word ids.
# tokens = nltk.tokenize.word_tokenize(str(caption).lower()) # line 1
# caption = [] # line 2
# caption.append(self.vocab(self.vocab.start_word)) # line 3
# caption.extend([self.vocab(token) for token in tokens]) # line 4
# caption.append(self.vocab(self.vocab.end_word)) # line 5
# caption = torch.Tensor(caption).long() # line 6
# ```
#
# As you will see soon, this code converts any string-valued caption to a list of integers, before casting it to a PyTorch tensor. To see how this code works, we'll apply it to the sample caption in the next code cell.
sample_caption = 'A person doing a trick on a rail while riding a skateboard.'
# In **`line 1`** of the code snippet, every letter in the caption is converted to lowercase, and the [`nltk.tokenize.word_tokenize`](http://www.nltk.org/) function is used to obtain a list of string-valued tokens. Run the next code cell to visualize the effect on `sample_caption`.
# +
import nltk
sample_tokens = nltk.tokenize.word_tokenize(str(sample_caption).lower())
print(sample_tokens)
# -
# In **`line 2`** and **`line 3`** we initialize an empty list and append an integer to mark the start of a caption. The [paper](https://arxiv.org/pdf/1411.4555.pdf) that you are encouraged to implement uses a special start word (and a special end word, which we'll examine below) to mark the beginning (and end) of a caption.
#
# This special start word (`"<start>"`) is decided when instantiating the data loader and is passed as a parameter (`start_word`). You are **required** to keep this parameter at its default value (`start_word="<start>"`).
#
# As you will see below, the integer `0` is always used to mark the start of a caption.
# +
sample_caption = []
start_word = data_loader.dataset.vocab.start_word
print('Special start word:', start_word)
sample_caption.append(data_loader.dataset.vocab(start_word))
print(sample_caption)
# -
# In **`line 4`**, we continue the list by adding integers that correspond to each of the tokens in the caption.
sample_caption.extend([data_loader.dataset.vocab(token) for token in sample_tokens])
print(sample_caption)
# In **`line 5`**, we append a final integer to mark the end of the caption.
#
# Identical to the case of the special start word (above), the special end word (`"<end>"`) is decided when instantiating the data loader and is passed as a parameter (`end_word`). You are **required** to keep this parameter at its default value (`end_word="<end>"`).
#
# As you will see below, the integer `1` is always used to mark the end of a caption.
# +
end_word = data_loader.dataset.vocab.end_word
print('Special end word:', end_word)
sample_caption.append(data_loader.dataset.vocab(end_word))
print(sample_caption)
# -
# Finally, in **`line 6`**, we convert the list of integers to a PyTorch tensor and cast it to [long type](http://pytorch.org/docs/master/tensors.html#torch.Tensor.long). You can read more about the different types of PyTorch tensors on the [website](http://pytorch.org/docs/master/tensors.html).
# +
import torch
sample_caption = torch.Tensor(sample_caption).long()
print(sample_caption)
# -
# And that's it! In summary, any caption is converted to a list of tokens, with _special_ start and end tokens marking the beginning and end of the sentence:
# ```
# [<start>, 'a', 'person', 'doing', 'a', 'trick', 'while', 'riding', 'a', 'skateboard', '.', <end>]
# ```
# This list of tokens is then turned into a list of integers, where every distinct word in the vocabulary has an associated integer value:
# ```
# [0, 3, 98, 754, 3, 396, 207, 139, 3, 753, 18, 1]
# ```
# Finally, this list is converted to a PyTorch tensor. All of the captions in the COCO dataset are pre-processed using this same procedure from **`lines 1-6`** described above.
#
# As you saw, in order to convert a token to its corresponding integer, we call `data_loader.dataset.vocab` as a function. The details of how this call works can be explored in the `__call__` method in the `Vocabulary` class in **vocabulary.py**.
#
# ```python
# def __call__(self, word):
# if not word in self.word2idx:
# return self.word2idx[self.unk_word]
# return self.word2idx[word]
# ```
#
# The `word2idx` instance variable is a Python [dictionary](https://docs.python.org/3/tutorial/datastructures.html#dictionaries) that is indexed by string-valued keys (mostly tokens obtained from training captions). For each key, the corresponding value is the integer that the token is mapped to in the pre-processing step.
#
# Use the code cell below to view a subset of this dictionary.
# Preview the word2idx dictionary.
dict(list(data_loader.dataset.vocab.word2idx.items())[:10])
# We also print the total number of keys.
# Print the total number of keys in the word2idx dictionary.
print('Total number of tokens in vocabulary:', len(data_loader.dataset.vocab))
# As you will see if you examine the code in **vocabulary.py**, the `word2idx` dictionary is created by looping over the captions in the training dataset. If a token appears no less than `vocab_threshold` times in the training set, then it is added as a key to the dictionary and assigned a corresponding unique integer. You will have the option later to amend the `vocab_threshold` argument when instantiating your data loader. Note that in general, **smaller** values for `vocab_threshold` yield a **larger** number of tokens in the vocabulary. You are encouraged to check this for yourself in the next code cell by decreasing the value of `vocab_threshold` before creating a new data loader.
# +
# Modify the minimum word count threshold.
vocab_threshold = 4
# Obtain the data loader.
data_loader = get_loader(transform=transform_train,
mode='train',
batch_size=batch_size,
vocab_threshold=vocab_threshold,
vocab_from_file=False)
# -
# Print the total number of keys in the word2idx dictionary.
print('Total number of tokens in vocabulary:', len(data_loader.dataset.vocab))
# There are also a few special keys in the `word2idx` dictionary. You are already familiar with the special start word (`"<start>"`) and special end word (`"<end>"`). There is one more special token, corresponding to unknown words (`"<unk>"`). All tokens that don't appear anywhere in the `word2idx` dictionary are considered unknown words. In the pre-processing step, any unknown tokens are mapped to the integer `2`.
# +
unk_word = data_loader.dataset.vocab.unk_word
print('Special unknown word:', unk_word)
print('All unknown words are mapped to this integer:', data_loader.dataset.vocab(unk_word))
# -
# Check this for yourself below, by pre-processing the provided nonsense words that never appear in the training captions.
print(data_loader.dataset.vocab('jfkafejw'))
print(data_loader.dataset.vocab('ieowoqjf'))
# The final thing to mention is the `vocab_from_file` argument that is supplied when creating a data loader. To understand this argument, note that when you create a new data loader, the vocabulary (`data_loader.dataset.vocab`) is saved as a [pickle](https://docs.python.org/3/library/pickle.html) file in the project folder, with filename `vocab.pkl`.
#
# If you are still tweaking the value of the `vocab_threshold` argument, you **must** set `vocab_from_file=False` to have your changes take effect.
#
# But once you are happy with the value that you have chosen for the `vocab_threshold` argument, you need only run the data loader *one more time* with your chosen `vocab_threshold` to save the new vocabulary to file. Then, you can henceforth set `vocab_from_file=True` to load the vocabulary from file and speed the instantiation of the data loader. Note that building the vocabulary from scratch is the most time-consuming part of instantiating the data loader, and so you are strongly encouraged to set `vocab_from_file=True` as soon as you are able.
#
# Note that if `vocab_from_file=True`, then any supplied argument for `vocab_threshold` when instantiating the data loader is completely ignored.
# Obtain the data loader (from file). Note that it runs much faster than before!
data_loader = get_loader(transform=transform_train,
mode='train',
batch_size=batch_size,
vocab_from_file=True)
# In the next section, you will learn how to use the data loader to obtain batches of training data.
# <a id='step2'></a>
# ## Step 2: Use the Data Loader to Obtain Batches
#
# The captions in the dataset vary greatly in length. You can see this by examining `data_loader.dataset.caption_lengths`, a Python list with one entry for each training caption (where the value stores the length of the corresponding caption).
#
# In the code cell below, we use this list to print the total number of captions in the training data with each length. As you will see below, the majority of captions have length 10. Likewise, very short and very long captions are quite rare.
# +
from collections import Counter
# Tally the total number of training captions with each length.
counter = Counter(data_loader.dataset.caption_lengths)
lengths = sorted(counter.items(), key=lambda pair: pair[1], reverse=True)
for value, count in lengths:
print('value: %2d --- count: %5d' % (value, count))
# -
# To generate batches of training data, we begin by first sampling a caption length (where the probability that any length is drawn is proportional to the number of captions with that length in the dataset). Then, we retrieve a batch of size `batch_size` of image-caption pairs, where all captions have the sampled length. This approach for assembling batches matches the procedure in [this paper](https://arxiv.org/pdf/1502.03044.pdf) and has been shown to be computationally efficient without degrading performance.
#
# Run the code cell below to generate a batch. The `get_train_indices` method in the `CoCoDataset` class first samples a caption length, and then samples `batch_size` indices corresponding to training data points with captions of that length. These indices are stored below in `indices`.
#
# These indices are supplied to the data loader, which then is used to retrieve the corresponding data points. The pre-processed images and captions in the batch are stored in `images` and `captions`.
# +
import numpy as np
import torch.utils.data as data
# Randomly sample a caption length, and sample indices with that length.
indices = data_loader.dataset.get_train_indices()
print('sampled indices:', indices)
# Create and assign a batch sampler to retrieve a batch with the sampled indices.
new_sampler = data.sampler.SubsetRandomSampler(indices=indices)
data_loader.batch_sampler.sampler = new_sampler
# Obtain the batch.
images, captions = next(iter(data_loader))
print('images.shape:', images.shape)
print('captions.shape:', captions.shape)
# (Optional) Uncomment the lines of code below to print the pre-processed images and captions.
# print('images:', images)
# print('captions:', captions)
# -
# Each time you run the code cell above, a different caption length is sampled, and a different batch of training data is returned. Run the code cell multiple times to check this out!
#
# You will train your model in the next notebook in this sequence (**2_Training.ipynb**). This code for generating training batches will be provided to you.
#
# > Before moving to the next notebook in the sequence (**2_Training.ipynb**), you are strongly encouraged to take the time to become very familiar with the code in **data_loader.py** and **vocabulary.py**. **Step 1** and **Step 2** of this notebook are designed to help facilitate a basic introduction and guide your understanding. However, our description is not exhaustive, and it is up to you (as part of the project) to learn how to best utilize these files to complete the project. __You should NOT amend any of the code in either *data_loader.py* or *vocabulary.py*.__
#
# In the next steps, we focus on learning how to specify a CNN-RNN architecture in PyTorch, towards the goal of image captioning.
# <a id='step3'></a>
# ## Step 3: Experiment with the CNN Encoder
#
# Run the code cell below to import `EncoderCNN` and `DecoderRNN` from **model.py**.
# +
# Watch for any changes in model.py, and re-load it automatically.
% load_ext autoreload
% autoreload 2
# Import EncoderCNN and DecoderRNN.
from model import EncoderCNN, DecoderRNN
# -
# In the next code cell we define a `device` that you will use move PyTorch tensors to GPU (if CUDA is available). Run this code cell before continuing.
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
# Run the code cell below to instantiate the CNN encoder in `encoder`.
#
# The pre-processed images from the batch in **Step 2** of this notebook are then passed through the encoder, and the output is stored in `features`.
# +
# Specify the dimensionality of the image embedding.
embed_size = 256
#-#-#-# Do NOT modify the code below this line. #-#-#-#
# Initialize the encoder. (Optional: Add additional arguments if necessary.)
encoder = EncoderCNN(embed_size)
# Move the encoder to GPU if CUDA is available.
encoder.to(device)
# Move last batch of images (from Step 2) to GPU if CUDA is available.
images = images.to(device)
# Pass the images through the encoder.
features = encoder(images)
print('type(features):', type(features))
print('features.shape:', features.shape)
# Check that your encoder satisfies some requirements of the project! :D
assert type(features)==torch.Tensor, "Encoder output needs to be a PyTorch Tensor."
assert (features.shape[0]==batch_size) & (features.shape[1]==embed_size), "The shape of the encoder output is incorrect."
# -
# The encoder that we provide to you uses the pre-trained ResNet-50 architecture (with the final fully-connected layer removed) to extract features from a batch of pre-processed images. The output is then flattened to a vector, before being passed through a `Linear` layer to transform the feature vector to have the same size as the word embedding.
#
# 
#
# You are welcome (and encouraged) to amend the encoder in **model.py**, to experiment with other architectures. In particular, consider using a [different pre-trained model architecture](http://pytorch.org/docs/master/torchvision/models.html). You may also like to [add batch normalization](http://pytorch.org/docs/master/nn.html#normalization-layers).
#
# > You are **not** required to change anything about the encoder.
#
# For this project, you **must** incorporate a pre-trained CNN into your encoder. Your `EncoderCNN` class must take `embed_size` as an input argument, which will also correspond to the dimensionality of the input to the RNN decoder that you will implement in Step 4. When you train your model in the next notebook in this sequence (**2_Training.ipynb**), you are welcome to tweak the value of `embed_size`.
#
# If you decide to modify the `EncoderCNN` class, save **model.py** and re-execute the code cell above. If the code cell returns an assertion error, then please follow the instructions to modify your code before proceeding. The assert statements ensure that `features` is a PyTorch tensor with shape `[batch_size, embed_size]`.
# <a id='step4'></a>
# ## Step 4: Implement the RNN Decoder
#
# Before executing the next code cell, you must write `__init__` and `forward` methods in the `DecoderRNN` class in **model.py**. (Do **not** write the `sample` method yet - you will work with this method when you reach **3_Inference.ipynb**.)
#
# > The `__init__` and `forward` methods in the `DecoderRNN` class are the only things that you **need** to modify as part of this notebook. You will write more implementations in the notebooks that appear later in the sequence.
#
# Your decoder will be an instance of the `DecoderRNN` class and must accept as input:
# - the PyTorch tensor `features` containing the embedded image features (outputted in Step 3, when the last batch of images from Step 2 was passed through `encoder`), along with
# - a PyTorch tensor corresponding to the last batch of captions (`captions`) from Step 2.
#
# Note that the way we have written the data loader should simplify your code a bit. In particular, every training batch will contain pre-processed captions where all have the same length (`captions.shape[1]`), so **you do not need to worry about padding**.
# > While you are encouraged to implement the decoder described in [this paper](https://arxiv.org/pdf/1411.4555.pdf), you are welcome to implement any architecture of your choosing, as long as it uses at least one RNN layer, with hidden dimension `hidden_size`.
#
# Although you will test the decoder using the last batch that is currently stored in the notebook, your decoder should be written to accept an arbitrary batch (of embedded image features and pre-processed captions [where all captions have the same length]) as input.
#
# 
#
# In the code cell below, `outputs` should be a PyTorch tensor with size `[batch_size, captions.shape[1], vocab_size]`. Your output should be designed such that `outputs[i,j,k]` contains the model's predicted score, indicating how likely the `j`-th token in the `i`-th caption in the batch is the `k`-th token in the vocabulary. In the next notebook of the sequence (**2_Training.ipynb**), we provide code to supply these scores to the [`torch.nn.CrossEntropyLoss`](http://pytorch.org/docs/master/nn.html#torch.nn.CrossEntropyLoss) optimizer in PyTorch.
# +
# Specify the number of features in the hidden state of the RNN decoder.
hidden_size = 512
#-#-#-# Do NOT modify the code below this line. #-#-#-#
# Store the size of the vocabulary.
vocab_size = len(data_loader.dataset.vocab)
# Initialize the decoder.
decoder = DecoderRNN(embed_size, hidden_size, vocab_size)
# Move the decoder to GPU if CUDA is available.
decoder.to(device)
# Move last batch of captions (from Step 1) to GPU if CUDA is available
captions = captions.to(device)
# Pass the encoder output and captions through the decoder.
outputs = decoder(features, captions)
print('type(outputs):', type(outputs))
print('outputs.shape:', outputs.shape)
# Check that your decoder satisfies some requirements of the project! :D
assert type(outputs)==torch.Tensor, "Decoder output needs to be a PyTorch Tensor."
assert (outputs.shape[0]==batch_size) & (outputs.shape[1]==captions.shape[1]) & (outputs.shape[2]==vocab_size), "The shape of the decoder output is incorrect."
# -
# When you train your model in the next notebook in this sequence (**2_Training.ipynb**), you are welcome to tweak the value of `hidden_size`.
|
1_Preliminaries.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # [Multi-class classification with focal loss for imbalanced datasets](https://www.dlology.com/blog/multi-class-classification-with-focal-loss-for-imbalanced-datasets/)
# ## focal loss model
# + _cell_guid="7b4a9f11-c3ee-3525-8c28-c32b1ac0c586"
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
from tensorflow import keras
np.random.seed(42)
# + _cell_guid="31bf5b85-e43e-2e03-7c86-a7ce42a978bf"
# create data frame containing your data, each column can be accessed # by df['column name']
dataset = pd.read_csv('../input/PS_20174392719_1491204439457_log.csv')
del dataset['nameDest']
del dataset['nameOrig']
del dataset['type']
dataset.head()
# -
dataset['isFraud'].value_counts()
def feature_normalize(dataset):
mu = np.mean(dataset, axis=0)
sigma = np.std(dataset, axis=0)
return (dataset - mu) / sigma
# + _cell_guid="22768c2d-9bd9-3ad6-3cf7-cb124e755981"
#Splitting the Training/Test Data
from sklearn.model_selection import train_test_split
X, y = dataset.iloc[:,:-2], dataset.iloc[:, -2]
y = keras.utils.to_categorical(y, num_classes=2)
X = feature_normalize(X.as_matrix())
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=1)
# + _cell_guid="c54a1358-8290-efad-7f3a-39df646850cf"
from tensorflow.keras.models import Sequential
import tensorflow as tf
model = Sequential()
from tensorflow.keras.layers import Dense
input_dim = X_train.shape[1]
nb_classes = y_train.shape[1]
model.add(Dense(10, input_dim=input_dim, activation='relu', name='input'))
model.add(Dense(20, activation='relu', name='fc1'))
model.add(Dense(10, activation='relu', name='fc2'))
model.add(Dense(nb_classes, activation='softmax', name='output'))
# +
def focal_loss(gamma=2., alpha=4.):
gamma = float(gamma)
alpha = float(alpha)
def focal_loss_fixed(y_true, y_pred):
"""Focal loss for multi-classification
FL(p_t)=-alpha(1-p_t)^{gamma}ln(p_t)
Notice: y_pred is probability after softmax
gradient is d(Fl)/d(p_t) not d(Fl)/d(x) as described in paper
d(Fl)/d(p_t) * [p_t(1-p_t)] = d(Fl)/d(x)
Focal Loss for Dense Object Detection
https://arxiv.org/abs/1708.02002
Arguments:
y_true {tensor} -- ground truth labels, shape of [batch_size, num_cls]
y_pred {tensor} -- model's output, shape of [batch_size, num_cls]
Keyword Arguments:
gamma {float} -- (default: {2.0})
alpha {float} -- (default: {4.0})
Returns:
[tensor] -- loss.
"""
epsilon = 1.e-9
y_true = tf.convert_to_tensor(y_true, tf.float32)
y_pred = tf.convert_to_tensor(y_pred, tf.float32)
model_out = tf.add(y_pred, epsilon)
ce = tf.multiply(y_true, -tf.log(model_out))
weight = tf.multiply(y_true, tf.pow(tf.subtract(1., model_out), gamma))
fl = tf.multiply(alpha, tf.multiply(weight, ce))
reduced_fl = tf.reduce_max(fl, axis=1)
return tf.reduce_mean(reduced_fl)
return focal_loss_fixed
model.compile(loss=focal_loss(alpha=1),
optimizer='nadam',
metrics=['accuracy'])
# -
model.summary()
# + _cell_guid="9153fa38-a6c0-834b-3f90-74067157abb0"
# model.compile(loss='binary_crossentropy',
# optimizer='nadam',
# metrics=['accuracy'])
# +
# class_weight = {0 : 1.,
# 1: 20.}
# -
y_train.shape
# + _cell_guid="9698ed89-e4e9-33a3-abdc-cb44fa032f8d"
model.fit(X_train, y_train, epochs=3, batch_size=1000)
# + _cell_guid="ef09528a-24db-d9b1-5d05-85cac75d5d27"
score = model.evaluate(X_test, y_test, batch_size=1000)
score
# +
# %matplotlib inline
from sklearn import metrics
import matplotlib.pyplot as plt
import seaborn as sns
sns.set(font_scale=2)
predictions = model.predict(X_test, batch_size=1000)
LABELS = ['Normal','Fraud']
max_test = np.argmax(y_test, axis=1)
max_predictions = np.argmax(predictions, axis=1)
confusion_matrix = metrics.confusion_matrix(max_test, max_predictions)
plt.figure(figsize=(5, 5))
sns.heatmap(confusion_matrix, xticklabels=LABELS, yticklabels=LABELS, annot=True, fmt="d", annot_kws={"size": 20});
plt.title("Confusion matrix", fontsize=20)
plt.ylabel('True label', fontsize=20)
plt.xlabel('Predicted label', fontsize=20)
plt.show()
# -
# ### Total miss-classified labels
values = confusion_matrix.view()
error_count = values.sum() - np.trace(values)
error_count
|
src/keras_focal_loss.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Yfyangd/BMIR/blob/master/HW2_Optional.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="D1M38Y1jmeXT" colab_type="text"
# # Import Library
# + id="Le7kh1AEmiQB" colab_type="code" colab={}
from tensorflow import keras
import tensorflow as tf
from tensorflow.keras import datasets, layers, models
# + [markdown] id="ihIusDZOmi2_" colab_type="text"
# # Import Fashion Minst Datasets
# + id="Th7p6DukmCbI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 151} outputId="d758c52d-1a5c-4d2f-e3f5-dfd487a4f864"
fashion_mnist = keras.datasets.fashion_mnist
(train_images, train_labels), (test_images, test_labels) = fashion_mnist.load_data()
class_names = ['T-shirt/top', 'Trouser', 'Pullover', 'Dress', 'Coat','Sandal', 'Shirt', 'Sneaker', 'Bag', 'Ankle boot']
train_images = train_images.astype('float32') / 255
train_images = train_images.reshape(train_images.shape[0], 28, 28, 1)
test_images = test_images.astype('float32') / 255
test_images = test_images.reshape(test_images.shape[0], 28, 28, 1)
train_labels = train_labels.reshape(train_labels.shape[0], 1)
test_labels = test_labels.reshape(test_labels.shape[0], 1)
# + id="XOm8Ug4nnM-J" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="31245d05-9e17-4902-f919-0731b096aaa2"
train_images.shape
# + [markdown] id="MTzDmwkXqDEH" colab_type="text"
# # Traditional neural network of HW1
# Test accuracy is **0.8827**
# + id="vPEKzrwBpZdU" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 403} outputId="fda2bd8b-0e02-40da-ae99-d8a3540b1f71"
model = keras.Sequential([
keras.layers.Flatten(input_shape=(28, 28)),
keras.layers.Dense(128, activation='relu'),
keras.layers.Dense(10)
])
model.compile(optimizer='adam',
loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=['accuracy'])
model.fit(train_images, train_labels, epochs=10)
test_loss, test_acc = model.evaluate(test_images, test_labels, verbose=2)
print('\nTest accuracy:', test_acc)
# + [markdown] id="yGjsHc8MnA93" colab_type="text"
# # Model: LeNet-5
# + id="nDHgd7rQnHHp" colab_type="code" colab={}
model = keras.Sequential()
model.add(layers.Conv2D(filters=6, kernel_size=(3, 3), activation='relu', input_shape=(28,28,1)))
model.add(layers.AveragePooling2D())
model.add(layers.Conv2D(filters=16, kernel_size=(3, 3), activation='relu'))
model.add(layers.AveragePooling2D())
model.add(layers.Flatten())
model.add(layers.Dense(units=128, activation='relu'))
model.add(layers.Dense(units=64, activation='relu'))
model.add(layers.Dense(units=10, activation = 'softmax'))
# + [markdown] id="EHjKfLitnXAq" colab_type="text"
# # Model Training
# + id="cWky-QV7nZnZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 353} outputId="d30ed68c-2d5d-4413-a373-2c139346ec33"
model.compile(optimizer='adam',
loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=['accuracy'])
history = model.fit(train_images, train_labels, epochs=10,
validation_data=(test_images, test_labels))
# + id="zQHT4O-1npzx" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 50} outputId="bdb89cdc-806f-40fd-eddc-ece495e70783"
test_loss, test_acc = model.evaluate(test_images, test_labels, verbose=2)
print("Part_3_fashion_mist: test accuracy: ", test_acc)
# + [markdown] id="jQflpgyVnnNi" colab_type="text"
# # Result
# * Compare to the traditional neural network of HW1 (accuracy: **0.8791**), the test accuracy of LeNet-5 is **0.8724**.
# * 理論上Convolution Network performance應該要比傳統的Neural Network要高, 但是在這邊LeNet-5 準確度 與 HW-1傳統model 相差不多的原因可能如下:
# > * LeNet-5 convolution 只有2層, 特徵提取不太足夠. 在我的Homework2中, 使用3層convolution 的準確度為0.9074, 高於LetNet-5 與 傳統的Nerual network
# > * 對於解析度比較低的圖片, 傳統的Neural network全部平坦化的節點數不多, 沒有convolution提取特徵的缺點比較不顯著
# > * LeNet-5 使用的是Average Pooling, 在homework-2使用的是Max Pooling, 這可能也是影響performance的原因
|
HW2_Optional.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # NTDS'18 tutorial 2: build a graph from features
# [<NAME>](https://people.epfl.ch/benjamin.ricaud), [EPFL LTS2](https://lts2.epfl.ch), with contributions from [<NAME>](http://deff.ch) and [<NAME>](https://lts4.epfl.ch/simou).
#
# * Dataset: [Iris](https://archive.ics.uci.edu/ml/datasets/Iris)
# * Tools: [pandas](https://pandas.pydata.org), [numpy](http://www.numpy.org), [scipy](https://www.scipy.org), [matplotlib](https://matplotlib.org), [networkx](https://networkx.github.io), [gephi](https://gephi.org/)
# ## Tools
# The below line is a [magic command](https://ipython.readthedocs.io/en/stable/interactive/magics.html) that allows plots to appear in the notebook.
# %matplotlib inline
# The first thing is always to import the packages we'll use.
import pandas as pd
import numpy as np
from scipy.spatial.distance import pdist, squareform
from matplotlib import pyplot as plt
import networkx as nx
# Tutorials on pandas can be found at:
# * <https://pandas.pydata.org/pandas-docs/stable/10min.html>
# * <https://pandas.pydata.org/pandas-docs/stable/tutorials.html>
#
# Tutorials on numpy can be found at:
# * <https://docs.scipy.org/doc/numpy/user/quickstart.html>
# * <http://www.scipy-lectures.org/intro/numpy/index.html>
# * <http://www.scipy-lectures.org/advanced/advanced_numpy/index.html>
#
# A tutorial on networkx can be found at:
# * <https://networkx.github.io/documentation/stable/tutorial.html>
# ## Import and explore the data
#
# We will play with the famous Iris dataset. This dataset can be found in many places on the net and was first released at <https://archive.ics.uci.edu/ml/index.php>. For example it is stored on [Kaggle](https://www.kaggle.com/uciml/iris/), with many demos and Jupyter notebooks you can test (have a look at the "kernels" tab).
#
# 
iris = pd.read_csv('data/iris.csv')
iris.head()
# The description of the entries is given here:
# https://www.kaggle.com/uciml/iris/home
iris['Species'].unique()
iris.describe()
# ## Build a graph from the features
#
# We are going to build a graph from this data. The idea is to represent iris samples (rows of the table) as nodes, with connections depending on their physical similarity.
#
# The main question is how to define the notion of similarity between the flowers. For that, we need to introduce a measure of similarity. It should use the properties of the flowers and provide a positive real value for each pair of samples. The value should be larger for more similar samples.
#
# Let us separate the data into two parts: physical properties and labels.
features = iris.loc[:, ['SepalLengthCm', 'SepalWidthCm', 'PetalLengthCm', 'PetalWidthCm']]
species = iris.loc[:, 'Species']
features.head()
species.head()
# ### Similarity, distance and edge weight
# You can define many similarity measures. One of the most intuitive and perhaps the easiest to program relies on the notion of distance. If a distance between samples is defined, we can compute the weight accordingly: if the distance is short, which means the nodes are similar, we want a strong edge (large weight).
# #### Different distances
# The cosine distance is a good candidate for high-dimensional data. It is defined as follows:
# $$d(u,v) = 1 - \frac{u \cdot v} {\|u\|_2 \|v\|_2},$$
# where $u$ and $v$ are two feature vectors.
#
# The distance is proportional to the angle formed by the two vectors (0 if colinear, 1 if orthogonal, 2 if opposed direction).
#
# Alternatives are the [$p$-norms](https://en.wikipedia.org/wiki/Norm_%28mathematics%29#p-norm) (or $\ell_p$-norms), defined as
# $$d(u,v) = \|u - v\|_p,$$
# of which the Euclidean distance is a special case with $p=2$.
# The `pdist` function from `scipy` computes the pairwise distance. By default it is the Euclidian distance. `features.values` is a numpy array extracted from the Pandas dataframe. Very handy.
# +
#from scipy.spatial.distance import pdist, squareform
# pdist?
# -
distances = pdist(features.values, metric='euclidean')
# other metrics: 'cosine', 'cityblock', 'minkowski'
# Now that we have a distance, we can compute the weights.
# #### Distance to weights
# A common function used to turn distances into edge weights is the Gaussian function:
# $$\mathbf{W}(u,v) = \exp \left( \frac{-d^2(u, v)}{\sigma^2} \right),$$
# where $\sigma$ is the parameter which controls the width of the Gaussian.
#
# The function giving the weights should be positive and monotonically decreasing with respect to the distance. It should take its maximum value when the distance is zero, and tend to zero when the distance increases. Note that distances are non-negative by definition. So any funtion $f : \mathbb{R}^+ \rightarrow [0,C]$ that verifies $f(0)=C$ and $\lim_{x \rightarrow +\infty}f(x)=0$ and is *strictly* decreasing should be adapted. The choice of the function depends on the data.
#
# Some examples:
# * A simple linear function $\mathbf{W}(u,v) = \frac{d_{max} - d(u, v)}{d_{max} - d_{min}}$. As the cosine distance is bounded by $[0,2]$, a suitable linear function for it would be $\mathbf{W}(u,v) = 1 - d(u,v)/2$.
# * A triangular kernel: a straight line between the points $(0,1)$ and $(t_0,0)$, and equal to 0 after this point.
# * The logistic kernel $\left(e^{d(u,v)} + 2 + e^{-d(u,v)} \right)^{-1}$.
# * An inverse function $(\epsilon+d(u,v))^{-n}$, with $n \in \mathbb{N}^{+*}$ and $\epsilon \in \mathbb{R}^+$.
# * You can find some more [here](https://en.wikipedia.org/wiki/Kernel_%28statistics%29).
#
# Let us use the Gaussian function
kernel_width = distances.mean()
weights = np.exp(-distances**2 / kernel_width**2)
# Turn the list of weights into a matrix.
adjacency = squareform(weights)
# Sometimes, you may need to compute additional features before processing them with some machine learning or some other data processing step. With Pandas, it is as simple as that:
# Compute a new column using the existing ones.
features['SepalLengthSquared'] = features['SepalLengthCm']**2
features.head()
# Coming back to the weight matrix, we have obtained a full matrix but we may not need all the connections (reducing the number of connections saves some space and computations!). We can sparsify the graph by removing the values (edges) below some fixed threshold. Let us see what kind of threshold we could use:
plt.hist(weights)
plt.title('Distribution of weights')
plt.show()
# Let us choose a threshold of 0.6.
# Too high, we will have disconnected components
# Too low, the graph will have too many connections
adjacency[adjacency < 0.6] = 0
# #### Remark: The distances presented here do not work well for categorical data.
# ## Graph visualization
#
# To conclude, let us visualize the graph. We will use the python module networkx.
# A simple command to create the graph from the adjacency matrix.
graph = nx.from_numpy_array(adjacency)
# Let us try some direct visualizations using networkx.
nx.draw_spectral(graph)
# Oh! It seems to be separated in 3 parts! Are they related to the 3 different species of iris?
#
# Let us try another [layout algorithm](https://en.wikipedia.org/wiki/Graph_drawing#Layout_methods), where the edges are modeled as springs.
nx.draw_spring(graph)
# Save the graph to disk in the `gexf` format, readable by gephi and other tools that manipulate graphs. You may now explore the graph using gephi and compare the visualizations.
nx.write_gexf(graph,'iris.gexf')
|
tutorials/02b_graph_from_features.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import sys
import scipy.io
import scipy.misc
import imageio
import matplotlib.pyplot as plt
from matplotlib.pyplot import imshow
from PIL import Image
from nst_utils import *
import numpy as np
import tensorflow as tf
# %matplotlib inline
# -
model = load_vgg_model("pretrained-model/imagenet-vgg-verydeep-19.mat")
print(model)
content_image = imageio.imread("/Users/chukaezema/Documents/style_transfer/images/uOttawa-building1.jpg")
imshow(content_image)
def compute_content_cost(a_C, a_G):
"""
Computes the content cost
Arguments:
a_C -- tensor of dimension (1, n_H, n_W, n_C), hidden layer activations representing content of the image C
a_G -- tensor of dimension (1, n_H, n_W, n_C), hidden layer activations representing content of the image G
Returns:
J_content -- scalar that you compute using equation 1 above.
"""
# Retrieving the dimensions from a_G
m, n_H, n_W, n_C = a_G.get_shape().as_list()
# Reshaping a_C and a_G
a_C_unrolled = tf.transpose(
tf.reshape(a_C,(1, n_H, n_W, n_C)),
name='transpose'
)
a_G_unrolled = tf.transpose(
tf.reshape(a_G,(1, n_H, n_W, n_C)),
name='transpose'
)
# computing the cost with tensorflow
J_content = (1/(4*n_H*n_W*n_C))*tf.reduce_sum(tf.square(tf.subtract(a_C_unrolled, a_G_unrolled)))
return J_content
# +
tf.reset_default_graph()
with tf.Session() as test:
tf.set_random_seed(1)
a_C = tf.random_normal([1, 4, 4, 3], mean=1, stddev=4)
a_G = tf.random_normal([1, 4, 4, 3], mean=1, stddev=4)
J_content = compute_content_cost(a_C, a_G)
print("J_content = " + str(J_content.eval()))
# -
style_image = imageio.imread("/Users/chukaezema/Documents/style_transfer/images/web_IMG_3091.jpg")
imshow(style_image)
def gram_matrix(A):
"""
Argument:
A -- matrix of shape (n_C, n_H*n_W)
Returns:
GA -- Gram matrix of A, of shape (n_C, n_C)
"""
GA = tf.matmul(A, tf.transpose(A))
return GA
# +
tf.reset_default_graph()
with tf.Session() as test:
tf.set_random_seed(1)
A = tf.random_normal([3, 2*1], mean=1, stddev=4)
GA = gram_matrix(A)
print("GA = " + str(GA.eval()))
# -
def compute_layer_style_cost(a_S, a_G):
"""
Arguments:
a_S -- tensor of dimension (1, n_H, n_W, n_C), hidden layer activations representing style of the image S
a_G -- tensor of dimension (1, n_H, n_W, n_C), hidden layer activations representing style of the image G
Returns:
J_style_layer -- tensor representing a scalar value, style cost defined above by equation (2)
"""
# Retrieving dimensions from a_G
m, n_H, n_W, n_C = a_G.get_shape().as_list()
# Reshaping the images to have shape (n_C, n_H*n_W)
a_S = tf.transpose(tf.reshape(a_S, [n_H*n_W, n_C]))
a_G = tf.transpose(tf.reshape(a_G, [n_H*n_W, n_C]))
# Computing gram_matrices for both images S and G
GS = gram_matrix(a_S)
GG = gram_matrix(a_G)
# Computing the loss (≈1 line)
J_style_layer = (1/(4*np.square(n_C)*np.square(n_H*n_W))) * tf.reduce_sum(tf.square(tf.subtract(GS,GG)))
return J_style_layer
# +
tf.reset_default_graph()
with tf.Session() as test:
tf.set_random_seed(1)
a_S = tf.random_normal([1, 4, 4, 3], mean=1, stddev=4)
a_G = tf.random_normal([1, 4, 4, 3], mean=1, stddev=4)
J_style_layer = compute_layer_style_cost(a_S, a_G)
print("J_style_layer = " + str(J_style_layer.eval()))
# -
STYLE_LAYERS = [
('conv1_1', 0.2),
('conv2_1', 0.2),
('conv3_1', 0.2),
('conv4_1', 0.2),
('conv5_1', 0.2)]
def compute_style_cost(model, STYLE_LAYERS):
# initializing the overall style cost
J_style = 0
for layer_name, coeff in STYLE_LAYERS:
# Selecting the output tensor of the currently selected layer
out = model[layer_name]
# Setting a_S to be the hidden layer activation from the layer we have selected, by running the session on out
a_S = sess.run(out)
# Setting a_G to be the hidden layer activation from same layer. Here, a_G references model[layer_name]
a_G = out
# Computing style_cost for the current layer
J_style_layer = compute_layer_style_cost(a_S, a_G)
# Adding coeff * J_style_layer of this layer to overall style cost
J_style += coeff * J_style_layer
return J_style
def total_cost(J_content, J_style, alpha = 10, beta = 40):
J = (alpha*J_content)+(beta*J_style)
return J
# +
tf.reset_default_graph()
with tf.Session() as test:
np.random.seed(3)
J_content = np.random.randn()
J_style = np.random.randn()
J = total_cost(J_content, J_style)
print("J = " + str(J))
# +
# Resetting the graph
tf.reset_default_graph()
# Starting interactive session
sess = tf.InteractiveSession()
# -
content_image = imageio.imread("/Users/chukaezema/Documents/style_transfer/images/uOttawa-building1.jpg")
content_image = reshape_and_normalize_image(content_image)
style_image = imageio.imread("/Users/chukaezema/Documents/style_transfer/images/web_IMG_3091.jpg")
style_image = reshape_and_normalize_image(style_image)
generated_image = generate_noise_image(content_image)
imshow(generated_image[0])
model = load_vgg_model("pretrained-model/imagenet-vgg-verydeep-19.mat")
# +
# Assigning the content image to be the input of the VGG model.
sess.run(model['input'].assign(content_image))
# Selecting the output tensor of layer conv4_2
out = model['conv4_2']
# Setting a_C to be the hidden layer activation from the layer we have selected
a_C = sess.run(out)
# Setting a_G to be the hidden layer activation from same layer. Here, a_G references model['conv4_2']
a_G = out
# Computing the content cost
J_content = compute_content_cost(a_C, a_G)
# +
# Assigning the input of the model to be the "style" image
sess.run(model['input'].assign(style_image))
# Computing the style cost
J_style = compute_style_cost(model, STYLE_LAYERS)
# -
J = total_cost(J_content, J_style, alpha = 10, beta = 40)
# +
# defining optimizer
optimizer = tf.train.AdamOptimizer(2.0)
# defining train_step
train_step = optimizer.minimize(J)
# -
def model_nn(sess, input_image, num_iterations = 200):
# Initializing global variables (you need to run the session on the initializer)
sess.run(tf.global_variables_initializer())
# Running the noisy input image (initial generated image) through the model. Use assign().
sess.run(model['input'].assign(input_image))
for i in range(num_iterations):
# Running the session on the train_step to minimize the total cost
_ = sess.run(train_step)
### END CODE HERE ###
# Computing the generated image by running the session on the current model['input']
generated_image = sess.run(model['input'])
# Printing every 20 iteration.
if i%20 == 0:
Jt, Jc, Js = sess.run([J, J_content, J_style])
print("Iteration " + str(i) + " :")
print("total cost = " + str(Jt))
print("content cost = " + str(Jc))
print("style cost = " + str(Js))
# save current generated image in the "/output" directory
save_image("output/" + str(i) + ".png", generated_image)
# saving last generated image
save_image('output/generated_image.jpg', generated_image)
return generated_image
model_nn(sess, generated_image)
|
Neural_Style_Transfer.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Build a sklearn Pipeline for a to ML contest submission
# In the ML_coruse_train notebook we at first analyzed the housing dataset to gain statistical insights and then e.g. features added new,
# replaced missing values and scaled the colums using pandas dataset methods.
# In the following we will use sklearn [Pipelines](https://scikit-learn.org/stable/modules/generated/sklearn.pipeline.Pipeline.html) to integrate all these steps into one final *estimator*. The resulting pipeline can be used for saving an ML estimator to a file and use it later for production.
#
# *Optional:*
# If you want, you can save your estimator as explained in the last cell at the bottom of this notebook.
# Based on a hidden dataset, it's performance will then be ranked against all other submissions.
# +
# read housing data again
import pandas as pd
import numpy as np
housing = pd.read_csv("datasets/housing/housing.csv")
# Try to get header information of the dataframe:
housing.head()
# -
# One remark: sklearn transformers do **not** act on pandas dataframes. Instead, they use numpy arrays.
# Now try to [convert](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.to_numpy.html) a dataframe to a numpy array:
housing.head().to_numpy()
housing.head()
# As you can see, the column names are lost now.
# In a numpy array, columns indexed using integers and no more by their names.
# ### Add extra feature columns
# At first, we again add some extra columns (e.g. `rooms_per_household, population_per_household, bedrooms_per_household`) which might correlate better with the predicted parameter `median_house_value`.
# For modifying the dataset, we now use a [FunctionTransformer](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.FunctionTransformer.html), which we later can put into a pipeline.
# Hints:
# * For finding the index number of a given column name, you can use the method [get_loc()](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.Index.get_loc.html)
# * For concatenating the new columns with the given array, you can use numpy method [c_](https://docs.scipy.org/doc/numpy/reference/generated/numpy.c_.html)
# +
from sklearn.preprocessing import FunctionTransformer
# At first, get the indexes as integers from the column names:
rooms_ix = housing.columns.get_loc("total_rooms")
bedrooms_ix = housing.columns.get_loc("total_bedrooms")
population_ix = housing.columns.get_loc("population")
household_ix = housing.columns.get_loc("households")
# Now implement a function which takes a numpy array a argument and adds the new feature columns
def add_extra_features(X):
rooms_per_household = X[:, rooms_ix] / X[:, household_ix]
population_per_household = X[:, population_ix] / X[:, household_ix]
bedrooms_per_household = X[:, bedrooms_ix] / X[:, household_ix]
# Concatenate the original array X with the new columns
return np.c_[X, rooms_per_household, population_per_household, bedrooms_per_household]
attr_adder = FunctionTransformer(add_extra_features, validate = False)
housing_extra_attribs = attr_adder.fit_transform(housing.values)
assert housing_extra_attribs.shape == (16512, 13)
housing_extra_attribs
# -
# ### Imputing missing elements
# For replacing nan values in the dataset with the mean or median of the column they are in, you can also use a [SimpleImputer](https://scikit-learn.org/stable/modules/generated/sklearn.impute.SimpleImputer.html) :
# +
from sklearn.impute import SimpleImputer
# Drop the categorial column ocean_proximity
housing_num = housing.drop('ocean_proximity', axis=1)
print("We have %d nan elements in the numerical columns" %np.count_nonzero(np.isnan(housing_num.values)))
imp_mean = SimpleImputer(missing_values=np.nan, strategy='mean')
housing_num_cleaned = imp_mean.fit_transform(housing_num)
assert np.count_nonzero(np.isnan(housing_num_cleaned)) == 0
housing_num_cleaned[1,:]
# -
# ### Column scaling
# For scaling the columns, you can use a [StandardScaler](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.StandardScaler.html)
# +
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
scaled = scaler.fit_transform(housing_num_cleaned)
print("mean of the columns is: " , np.mean(scaled, axis=0))
print("standard deviation of the columns is: " , np.std(scaled, axis=0))
# -
# ### Putting all preprocessing steps together
# Now let's build a pipeline for preprocessing the **numerical** attributes.
# The pipeline shall process the data in the following steps:
# * [Impute](https://scikit-learn.org/stable/modules/generated/sklearn.impute.SimpleImputer.html) median or mean values for elements which are NaN
# * Add attributes using the FunctionTransformer with the function add_extra_features().
# * Scale the numerical values using the [StandardScaler()](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.StandardScaler.html)
# +
from sklearn.pipeline import Pipeline
num_pipeline = Pipeline([
('imputer', SimpleImputer(strategy="median")),
('attribs_adder', FunctionTransformer(add_extra_features, validate=False)),
('std_scaler', StandardScaler()),
])
# Now test the pipeline on housing_num
num_pipeline.fit_transform(housing_num)
# -
# Now we have a pipeline for the numerical columns.
# But we still have a categorical column:
housing['ocean_proximity'].head()
# We need one more pipeline for the categorical column. Instead of the "Dummy encoding" we used before, we now use the [OneHotEncoder](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.OneHotEncoder.html) from sklearn.
from sklearn.preprocessing import OneHotEncoder
housing_cat = housing[['ocean_proximity']]
cat_encoder = OneHotEncoder(sparse = False)
housing_cat_1hot = cat_encoder.fit_transform(housing_cat)
housing_cat_1hot
# We have everything we need for building a preprocessing pipeline which transforms the columns including all the steps before.
# Since we have columns where different transformations should be applied, we use the class [ColumnTransformer](https://scikit-learn.org/stable/modules/generated/sklearn.compose.ColumnTransformer.html)
# +
from sklearn.compose import ColumnTransformer
num_attribs = ["longitude","latitude","housing_median_age","total_rooms", "total_bedrooms",
"population","households", "median_income"]
cat_attribs = ["ocean_proximity"]
full_prep_pipeline = ColumnTransformer([
("num", num_pipeline, num_attribs),
("cat", OneHotEncoder(), cat_attribs),
])
full_prep_pipeline.fit_transform(housing)
# -
# ### Train an estimator
# Include `full_prep_pipeline` into a further pipeline where it is followed by an RandomForestRegressor.
# This way, at first our data is prepared using `full_prep_pipeline` and then the RandomForestRegressor is trained on it.
# +
from sklearn.ensemble import RandomForestRegressor
from sklearn.model_selection import train_test_split
full_pipeline_with_predictor = Pipeline([
("preparation", full_prep_pipeline),
("forest", RandomForestRegressor())
])
# -
# At first, seperate the label colum (`median_house_value`) and feature columns (all other columns).
# Split the data into a training and testing dataset using train_test_split.
# +
# Create two dataframes, one for the labels one for the features
housing_features = housing.drop("median_house_value", axis = 1)
housing_labels = housing["median_house_value"]
# Split the two dataframes into a training and a test dataset
X_train, X_test, y_train, y_test = train_test_split(housing_features, housing_labels, test_size = 0.20)
# Now train the full_pipeline_with_predictor on the training dataset
full_pipeline_with_predictor.fit(X_train, y_train)
# -
# As usual, calculate some score metrics:
# +
from sklearn.metrics import mean_squared_error
y_pred = full_pipeline_with_predictor.predict(X_test)
tree_mse = mean_squared_error(y_pred, y_test)
tree_rmse = np.sqrt(tree_mse)
tree_rmse
# +
from sklearn.metrics import r2_score
r2_score(y_pred, y_test)
# -
# Use the [pickle serializer](https://docs.python.org/3/library/pickle.html) to save your estimator to a file for contest participation.
# +
import pickle
import getpass
from sklearn.utils.validation import check_is_fitted
your_regressor = full_pipeline_with_predictor
assert isinstance(your_regressor, Pipeline)
pickle.dump(your_regressor, open(getpass.getuser() + "s_model.p", "wb" ) )
# -
housing_valid = pd.read_csv("datasets/housing/housing_valid.csv")
housing_valid_features = housing_valid.drop("median_house_value", axis = 1)
housing_valid_labels = housing_valid["median_house_value"]
housing_valid_labels
with open('chrus_model.p', 'rb') as handle:
contestModel = pickle.load(handle)
y_pred = contestModel.predict(housing_valid_features)
tree_mse = mean_squared_error(y_pred, housing_valid_labels.to_numpy())
tree_rmse = np.sqrt(tree_mse)
print(tree_rmse)
# For generating hidden test set
housing_orig = pd.read_csv("datasets/housing/housing_orig.csv")
course, valid = train_test_split(housing_orig, test_size=0.2, random_state = 26)
valid.to_csv("datasets/housing/housing_valid.csv", index = False)
course.to_csv("datasets/housing/housing.csv", index = False)
course.head(5)
valid.head(5)
|
ML_course/ML_Contest.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Grove Light Sensor example
# ----
# * [Introduction](#Introduction)
# * [Set up the board](#Set-up-the-board)
# * [Setup and read from the sensor](#Setup-and-read-from-the-sensor)
# * [Display a graph](#Display-a-graph)
#
# ----
# ## Introduction
#
# This example shows how to use the [Grove Light Sensor v1.1](http://wiki.seeedstudio.com/Grove-Light_Sensor/), and how to plot a graph using matplotlib.
#
# The Grove Light Sensor produces an analog signal which requires an ADC.
#
# The Grove Light Sensor, Pynq Grove Adapter, and Grove ADC are used for this example.
#
# The Grove ADC is an example of an I2C peripheral.
#
#
# When the ambient light intensity increases, the resistance of the LDR or Photoresistor will decrease. This means that the output signal from this module will be HIGH in bright light, and LOW in the dark. Values for the sensor ranges from ~5.0 (bright) to >100.0 (dark). The sensor is more sensitive to darkness, and saturates in ambient light.
# ----
# ## Set up the board
#
# Start by loading the Base Overlay.
from pynq.overlays.base import BaseOverlay
base = BaseOverlay("base.bit")
# #### Connect the peripheral to the board.
# 1. Connect the Pynq Grove Adapter to ***PMODB.***
# 2. Connect ***Grove ADC*** port ***J1*** (SCL, SDA, VCC, GND) to port ***G4*** of the Pynq Grove Adapter.
# 3. Connect the ***Grove ALS*** to port ***J2*** of the ***Grove ADC*** (GND, VCC, NC, SIG)
#
# ----
#
# ## Setup and read from the sensor
# Grove ADC provides a raw sample which is converted into resistance first and then converted into temperature.
# This example shows on how to read from the Grove Light sensor which is connected to the ADC.
# +
from pynq.lib.pmod import Grove_Light
from pynq.lib.pmod import PMOD_GROVE_G4 # Import constants
# Grove2pmod is connected to PMODB (2)
# Grove ADC is connected to G4 (pins [6,2])
lgt = Grove_Light(base.PMODB, PMOD_GROVE_G4)
# -
sensor_val = lgt.read()
print(sensor_val)
# The peripheral can be connected to other ports by specifying the interface and the port when the sensor is declared above.
#
# ### Start logging once every 100ms for 10 seconds
#
# Executing the next cell will start logging the sensor values every 100ms, and will run for 10s. You can try covering and uncovering the light sensor to vary the measured signal.
#
# You can vary the logging interval and the duration by changing the values 100 and 10 in the cell below.
# +
import time
lgt.set_log_interval_ms(100)
lgt.start_log()
time.sleep(10) # Change input during this time
r_log = lgt.get_log()
print("Logged {} samples".format(len(r_log)))
# -
# ----
# ## Display a graph
#
# This example reads multiple values over a 10 second period.
#
# To change the light intensity, cover and uncover the light sensor. In typical ambient light, there is no need to provide an external light source, as the sensor is already reading at full scale.
# +
# %matplotlib inline
import matplotlib.pyplot as plt
plt.plot(range(len(r_log)), r_log, 'ro')
plt.title('Grove Light Sensor Plot')
plt.xlabel('Sample')
plt.ylabel('Darkness')
min_r_log = min(r_log)
max_r_log = max(r_log)
plt.axis([0, len(r_log), min_r_log, max_r_log])
plt.show()
|
zynq-old/PYNQ_Workshop/Session_2/4_pmod_grove_light.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Jupyter notebooks are widely used for data exploration, enabling analysts to write documents that contain software code, computational output, formatted text and data visualizations. In fact, this paper is written entirely in a Jupyter Notebook, which can be run by the interested reader in order to interact with the visualizations and explore the source code in more detail. The notebook is available on the paper GitHub page[^1]. Visualization is an essential component of the data exploration process, and can be frequently found in Jupyter Notebooks. For example, a recent study of public GitHub repositories found that *matplotlib* was the second most imported package in the notebook environment \cite{pimentel2019large}.
#
# [^1]: https://github.com/VIDA-NYU/Interactive-Visualization-Jupyter-Notebooks
#
# # Interactive Visualization in Jupyter Notebooks
#
# When datasets are too large or too complex, interactive visualization becomes a useful tool in exploratory data analysis. Interactive visualizations can enable, among many others, the display of information at multiple levels of detail, the exploration of data using coordinated views, and the dynamic change of the charts to focus on the user's interests. While notebooks have traditionally been used with static visualizations, it is possible to
# embed sophisticated interactive visualizations and support advanced visual analysis as well.
#
# In this paper, we present three simple and powerful approaches in which data scientists can create interactive visualizations in Jupyter Notebooks: *matplotlib* callbacks, visualization toolkits and custom HTML embedding. These approaches offer a number of benefits and drawbacks that need to be considered by the developer so that they can make an informed decision about their visualization task. By the end of this paper, the reader will have a good understanding of the three methods, and will be able to select an implementation approach depending on the level of interaction, customization and data flow desired.
# **Matplotlib Callbacks.** The *matplotlib* library \cite{hunter2007matplotlib} is the most popular general purpose visualization package for Jupyter Notebooks \cite{pimentel2019large}. This tool enables the creation of static, animated, and interactive visualizations, that can be rendered directly as the output of notebook cells. However, the available user interactions are limited: there is support for click and keypress events, but drag-and-drop, tooltips, and cross-filtering, frequently supported in visualization tools, are not directly provided. To expand the possible user interactions, [*ipywidgets*](https://ipywidgets.readthedocs.io) can be used. *ipywidgets* is a library that provides HTML form inputs in the Jupyter interface, including drop down menus, text boxes, and sliders.
#
# **Visualization Toolkits.** In order to enable the creation of more interactive visualizations in Python and Jupyter Notebooks, many open source visualization toolkits have been developed. Among those, Perkel et al. \cite{perkel2018data} highlight [*Plotly*](https://plotly.com/), [*Bokeh*](https://bokeh.org/) and [*Altair*](https://altair-viz.github.io/). These libraries are built on top of web technologies, and create visualizations that can be seen in web browsers. Syntax-wise, *Plotly* and *Bokeh* are very similar to *matplotlib*. However, both libraries have been developed with a focus on user interaction, enabling the creation of web-based dashboards that combine interactive widgets and charts, and support multiple user inputs, including click, drag-and-drop, tooltips, selection, crossfilter, and bidirectional communication with Python via callbacks. *Altair* \cite{vanderplas2018altair} differs from the aforementioned libraries in the way visualizations are defined: it uses a declarative specification that ports VEGA-Lite \cite{satyanarayan2016vega}, a data visualization grammar, to Python. A wide range of interactive visualizations can be expressed using a small number of Altair primitives, making this library very flexible. However, the produced visualizations cannot communicate with Python and therefore the results of user interactions cannot be used in further computations.
#
# **Custom HTML Embedding.** There might be cases when a visualization cannot be created using any off-the-shelf Python libraries. When this happens, the developer has the option to code the visualization using a web framework and embed it in the notebook. This option offers the most flexibility, as the visualization can be fully customized and interactions can be scripted on demand. JavaScript libraries such as [React](https://reactjs.org/) and [D3](https://d3js.org/) can be used to facilitate the implementation of custom visualizations.
#
# Table 1 summarizes the different approaches to add interactive visualizations in Jupyter Notebooks. The approaches are classified in terms of interaction, type of output, level of customization, support for dashboards, and data flow. When creating a new visualization, we believe these properties should be taken into consideration.
# **Table 1**: Summary of Interactive Visualization Approaches in Jupyter Notebook
#
# | Library | Interaction | Output | Customization | Dashboard | Data Flow |
# |:---------------|:--------------|:---------|:----------------|:------------|:--------------------------|
# | matplotlib | Low | Flexible | Low | Limited | Bidirectional |
# | Plotly | High | HTML | Low | Yes | Bidirectional |
# | Bokeh | High | HTML | Low | Yes | Bidirectional |
# | Altair | High | HTML | Low | Yes | Python → JavaScript |
# | HTML Embedding | High | HTML | High | Yes | Bidirectional |
# # Interactive Visualizations in Action
#
# In this section, we will show how to create interactive visualizations in Jupyter notebooks using three approaches discussed in the previous section: *matplotlib* charts, *Altair* specifications, and custom HTML visualizations. Since the syntax of *Plotly* and *Bokeh* are very similar to *matplotlib*, we will not cover them in this paper. We refer the interested reader to their online documentations.
# ## Matplotlib with Callbacks
#
# In order to enable interactive *matplotlib* charts in the notebook environment, users need to activate this option using the *"%matplotlib notebook"* magic command [^2]. The produced charts will natively support pan and zoom operations, but can be configured to receive other types of user input, such as mouse click and key press, which can trigger the run of user-defined callback functions [^3].
#
# After a chart is created, for example, using *pyplot.scatter*, the user events can be captured by setting callback functions on the *canvas* using the method *mpl_connect*. Multiple events are available, including *button_press_event*, *button_release_event*, *key_press_event* and *key_release_event*.
#
# [^2]: https://matplotlib.org/3.3.3/users/interactive.html
# [^3]: https://matplotlib.org/3.3.3/users/event_handling.html
#
# We show a minimal example below, where the visualization draws points on top of the user clicks. The resulting visualization in shown in Figure 1.
# + tags=["remove_output"]
# %matplotlib notebook
import matplotlib.pyplot as plt
fig, ax = plt.subplots(); # Creating an empty chart
plt.xlim([0, 10]); plt.ylim([0, 10]) # Setting X and Y axis limits
def onclick(event): # Callback function
ax.scatter(event.xdata, event.ydata, color='steelblue') # Draw a point on top of the user click position.
cid = fig.canvas.mpl_connect('button_press_event', onclick) # Callback setup
# -
# 
#
# **Figure 1**: Interactive Matplotlib chart, where the user can click on the canvas in order to add a point at that position. The interactive chart also enables pan and zoom operations by default.
# This approach can add *click* interactions to a chart with a few lines of code. However, we are limited to the types of charts and interactions supported by matplotlib. When these options are not enough, the developer might need to consider other libraries, such as *Altair*, or creating their own visualization in HTML/Javascript.
# ## Altair Specification
#
# Altair enables the creation of interactive visualizations by using a pythonic port of the Vega-Lite specification \cite{vanderplas2018altair}. Altair uses a declarative visualization paradigm: instead of telling the library every step of how to draw a chart, the programmer specifies the data and the visual encodings, and the library takes care of the rest.
#
# In order to create a chart, the developer needs to have a *Pandas DataFrame* containing the data to be visualized. An *Altair.Chart* object needs to be created, with the corresponding *DataFrame* passed as a parameter. Next, an *encoding* and a *mark* needs to be selected. *Encodings* tell *Altair* how the *DataFrame* columns should be mapped to visual attributes. Meanwhile, *marks* specify how the attributes should be represented on the plot (for example, as a circle, line, area chart, etc.).
#
# We show a basic example of an Altair scatter plot with the Iris dataset (Figure 2). The dataset contains information regarding 150 Iris flowers, with measurements of length and width of the plant, as well as the flower species. Data points can be hovered to show additional information as a tooltip (notice that this was not possible in *matplotlib*). In the code below, *mark_circle* is used to indicate the type of chart desired (scatter plot with circles) and the *encode* function specify the chart encoding, in this case, what columns are mapped to the *x* and *y* positions, *color* of the circle, and *tooltip* on hover.
# + tags=["remove_output"]
import altair as alt
from vega_datasets import data
df = data.iris()
alt.Chart(df).mark_circle().encode(
x='petalLength',
y='petalWidth',
color='species',
tooltip=['sepalLength', 'sepalWidth', 'petalLength', 'petalWidth', 'species']
).interactive()
# -
# 
#
# **Figure 2**: Interactive Altair scatter plot of the Iris dataset. The chart displays a tooltip with flower information on mouse hover. The library also enables pan and zoom.
# For more complex examples, please see the Altair documentation. There are many chart possibilities, and graphics can be combined to create interactive dashboards with multiple views. For example, Figure 3 (1) shows an Altair dashboard that visualizes a flight dataset (example taken from the online documentation[^4]). (2) The user can select flights based on delay (in hours) and see how delay correlates with the other variables (distance and time).
#
# 
#
# **Figure 3**: Altair Dashboard showing a flight dataset. (1) Histograms for flight distance, delay and time. (2) The user selected a range of delay values and the system automatically updates the other views.
#
# One disadvantage of *Altair* is that we cannot have access to data generated by the user in Python. For example, we would not be able to receive data points selected in Altair in the next Jupyter cell. Such capability exists in *matplotlib* and in custom JavaScript visualizations, because we can set up callbacks between JavaScript and Python.
#
# [^4]: https://altair-viz.github.io/gallery/interactive_layered_crossfilter.html
# + tags=["remove_cell"]
# Altair Dashboard with crossfilter
# Example taken from the Altair documentation
# https://altair-viz.github.io/gallery/interactive_layered_crossfilter.html
import altair as alt
from vega_datasets import data
source = alt.UrlData(
data.flights_2k.url,
format={'parse': {'date': 'date'}}
)
brush = alt.selection(type='interval', encodings=['x'])
# Define the base chart, with the common parts of the
# background and highlights
base = alt.Chart().mark_bar().encode(
x=alt.X(alt.repeat('column'), type='quantitative', bin=alt.Bin(maxbins=20)),
y='count()'
).properties(
width=160,
height=130
)
# gray background with selection
background = base.encode(
color=alt.value('#ddd')
).add_selection(brush)
# blue highlights on the transformed data
highlight = base.transform_filter(brush)
# layer the two charts & repeat
alt.layer(
background,
highlight,
data=source
).transform_calculate(
"time",
"hours(datum.date)"
).repeat(column=["distance", "delay", "time"])
# -
# ## HTML Embedding
#
# Displaying custom visualizations in a Jupyter Notebook can be done in a few lines of code using
# the package [*Ipython.display*](https://ipython.readthedocs.io/en/stable/api/generated/IPython.display.html), which embeds HTML code in notebook cells. The HTML may contain both CSS and JavaScript, which affords flexible, interactive and customizable visualizations to be created.
#
# In order to embed the visualization in a cell, one needs to create a string variable containing all the HTML, JavaScript, data and CSS code needed for the visualization. Since writing everything in a Jupyter cell can be too cumbersome, one can write the visualization in a code editor and then load the document in Python. JavaScript Bundlers, such as [Webpack](https://webpack.js.org/), can convert multiple HTML, JavaScript and CSS files into a single file, facilitating this process.
#
# In the following, we show an example of HTML embedding in a Jupyter cell. The code adds a single button to the page, which when clicked displays an alert box with the message "Hello World".
# + tags=["remove_output"]
from IPython.display import display, HTML
html_string = """<button onclick="alert('Hello World')">Hello World</button>"""
display(HTML(html_string))
# -
# Formatting methods can be used to create the HTML string. For example, a base string may contain the container *div* where the visualization is going to be inserted, a *script* tag where the bundled code is going to be added, and a function call to plot the visualization with the provided data in JSON format. The *string.format* function can be used to add the remaining information to the string, filling in the placeholders.
#
# The following code snippet shows how to embed a JavaScript library and CSV data in the HTML string. This example visualization shows an interactive chart that displays baseball game trajectories (Figure 4). The user can control the progress of the play using a slider. Furthermore, the user can select a player or the ball to edit its trajectory (either clicking on the field, or the button "Clear trajectory"). This visualization is an adaptation of the Baseball annotation system HistoryTracker \cite{ono2019historytracker}.
# + tags=["remove_output"]
from IPython.display import display, HTML
import pandas as pd
with open("./BaseballVisualizer/build/baseballvisualizer.js", "r") as f:
bundled_code = f.read()
play = pd.read_csv("./BaseballVisualizer/play_annotated.csv")
data = {'tracking': play.to_json(orient="records")}
html = """
<html>
<body>
<div id="container"/>
<script type="application/javascript">
{bundled_code}
baseballvisualizer.renderBaseballAnnotator("#container", {data});
</script>
</body>
</html>
""".format(bundled_code=bundled_code, data=data)
display(HTML(html))
# -
# 
#
# **Figure 4**: Custom JavaScript visualization of Baseball plays. The user can: (1) animate the play using the slider. Select a position to edit (in the picture, the BALL is selected) and (2) clear the trajectory. Annotate the positions of the ball when it is thrown (3) and hit to the center field (4).
# Callbacks can be set up in both JavaScript and Python using the *comm* API[^5] to send data from one to the other. For example, if a sports analyst is interested in modifying a Baseball trajectory and running some further analysis in Python, he might set up this bidirectional communication.
#
# A minor change needs to be made to both the JavaScript and to the Python code. In JavaScript, a new *comm* object needs to be created with an identifier (in this example, *submit_trajectory*). Then, the *comm* object is used to *send* a message to Python, containing the edited trajectory data. Finally, when Python acknowledges the message, we display an alert.
#
# ```javascript
# function submitTrajectoryToServer(trajectory){
# let comm = window.Jupyter.notebook.kernel.comm_manager.new_comm('submit_trajectory')
# // Send trajectory to Python
# comm.send({'trajectory': trajectory})
#
# // Receive message from Python
# comm.on_msg(function(msg) {
# alert("Trajectory received by Jupyter Notebook.")
# });
# }
# ```
#
# The Python code needs to expect a message from JavaScript. In order to set this up, we use the *register_target* function, passing to it the communication identifier and the Python callback function. In the following code snippet, this callback will store the trajectory in the variable *received_trajectory*.
#
# [^5]: https://jupyter-notebook.readthedocs.io/en/stable/comms.html
# + tags=["remove_output"]
received_trajectory = []
def receive_trajectory(comm, open_msg):
# comm is the kernel Comm instance
# Register handler for future messages
@comm.on_msg
def _recv(msg):
global received_trajectory
# Use msg['content']['data'] for the data in the message
received_trajectory = msg['content']['data']['trajectory']
print(received_trajectory)
comm.send({'received': True})
get_ipython().kernel.comm_manager.register_target('submit_trajectory', receive_trajectory)
# -
# Finally, after the user clicks the "Submit" button, the trajectory can be retrieved in the Jupyter notebook, analyzed and saved to disk.
# + tags=["remove_output"]
received_trajectory_df = pd.DataFrame(received_trajectory)
received_trajectory_df.to_csv("edited_trajectory.csv", index=False)
# -
# # To Look Further
#
# There are many domain-specific visualization libraries for Jupyter notebook which use the techniques described in this paper. Figure 5 shows examples in three different domains, which illustrate how diverse and flexible these visualizations can be. The examples belong to the fields of scientific visualization \cite{breddels2020ipygany}, sports analytics \cite{lage2016statcast} and machine learning \cite{nori2019interpretml, ono2020pipelineprofiler}. 1) *ipygany* \cite{breddels2020ipygany} enables the visualization of 3D meshes in Jupyter notebooks. Users can zoom, rotate and apply effects to 3D meshes interactively using this library. 2) *StatCast Dashboard* \cite{lage2016statcast} supports the interactive query, filter, and visualization of spatiotemporal baseball trajectories and statistics. The library communicates with a baseball play database in order to execute complex queries involving player, teams, game dates and events. 3) *InterpretML* \cite{nori2019interpretml} is a Python package that contains a collection of algorithms for explaining and visualizing Machine Learning (ML) models, including LIME, SHAP and Partial Dependency Plots. Finally, 4) *PipelineProfiler* \cite{ono2020pipelineprofiler} contains visualizations that enable the exploration and comparison of ML pipelines produced by Automatic Machine Learning systems.
#
# One of the advantages of Jupyter Notebooks is that they support reproducibility \cite{pimentel2019large}. However, when interactive visualizations are used in computational notebooks, additional mechanisms are needed to afford reproducible results. We refer the interested reader to Fekete and Freire \cite{fekete2020exploring} for a survey of reproducibility challenges faced by the visualization community and how those challenges can be addressed.
#
# In this paper, we have presented three ways to create interactive visualizations in Jupyter Notebooks: *matplotlib* charts, *Altair* specifications and custom HTML visualizations. We hope that this document will help developers to create their own interactive charts.
# 
#
# **Figure 5**: Domain-specific visualization libraries for Jupyter Notebook. 1) *ipygany*: visualization of 3D meshes. 2) *StatCast Dashboard*: visualization of Baseball trajectories and game statistics. 3) *InterpretML*: visualization of machine learning model explanations. 4) *PipelineProfiler*: visualization of machine learning pipelines produced by AutoML systems.
# # Acknowledgements
#
# We thank <NAME> for his thoughtful comments that helped improve our paper. This work was partially supported by the DARPA D3M program, NASA, Adobe, and NSF awards CNS-1229185, CCF-1533564, CNS-1544753, CNS-1730396, and CNS-1828576. Any opinions, findings, and conclusions or recommendations expressed in this material are those of the authors and do not necessarily reflect the views of NSF and DARPA.
|
Paper.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
data <- c(rnorm(20, mean=69, sd = 12))
summary(data)
sum(data)/20
# # Histograms with R
options(repr.plot.width = 6, repr.plot.height=4)
hist(data)
dataset <- c(91,49,76,112,97,42,70, 100, 8, 112, 95, 90, 78, 62,
56, 94, 65, 58, 109, 70, 109, 91, 71, 76, 68, 62, 134, 57, 83, 66)
hist(dataset, freq = F, xlab = "DataPoints", ylab = "Density")
hist(dataset, freq = F, xlab = "DataPoints", ylab = "Density", col="orange")
hist(dataset, freq = F, xlab = "DataPoints", ylab = "Density", col="orange")
lines(density(dataset))
hist(dataset, freq = F, xlab = "DataPoints", ylab = "Density", col="green")
lines(density(dataset), col = "red", lwd = 3)
hist(dataset, freq = F, xlab = "DataPoints", ylab = "Density", col="green", breaks = 30)
lines(density(dataset), col = "red", lwd = 3)
# # Histograms with GGplot2
library(ggplot2)
library(MASS)
df <- as.data.frame(dataset)
colnames(df) <- "x"
head(df)
ggplot(df) +
geom_histogram(aes(x=x), bins = 40) +
xlab("Data") +
ylab("Frequency") +
ggtitle("Histogram of My Data with Frequency") +
theme(plot.title = element_text(hjust = 0.5))
ggplot(df) +
geom_histogram(aes(x = x, y=..density..), bins = 40) +
geom_density(aes(x = x, y=..density..), size=0.6, fill="green", alpha=0.2, col="black") +
ggtitle("Histogram of My Data with Density") +
labs(xlabel = "Data", ylabel = "Frequency") +
theme(plot.title = element_text(hjust = 0.5))
ggplot(df, aes(x=x)) +
geom_histogram(aes(y=..density.., fill = factor(..count..)), col = "black", bins = 40) +
geom_density(aes(y=..density..), size=0.6, fill="gray", alpha=0.6, col="black") +
xlab(label = "Data") +
ylab(label = "Frequency") +
ggtitle("Histogram of My Data with Density") +
theme(plot.title = element_text(hjust = 0.5))
# # ScatterPlots in R
# +
set.seed(2016) # There is a typo in the video (set.seed=2016)
test1 <- c(round(rnorm(30, 78, 30)), round(rnorm(50, 15, 20)))
test2 <- c(round(rnorm(30, 78, 30)), round(rnorm(50, 15, 20)))
df <- as.data.frame(cbind(test1, test2))
# -
head(df)
options(repr.plot.width = 4, repr.plot.height=4)
plot(df$test1, df$test2, xlab = "Test 1", ylab = "Test 2", main="Scatter Plot")
plot(df$test1, df$test2, xlab = "Test 1", ylab = "Test 2", main="Scatter Plot", col="blue")
# # ScatterPlots in R
# +
options(repr.plot.width = 7, repr.plot.height=5)
ggplot(df, aes(x=test1, y=test2)) +
geom_jitter()
# -
ggplot(df, aes(x=test1, y=test2, z=sin(sqrt(test1^2 + test2^2)))) +
geom_jitter() +
geom_density_2d(col="blue", size=0.5)
ggplot(df, aes(x=test1, y=test2, z=sin(sqrt(test1^2 + test2^2)))) +
geom_jitter() +
stat_density_2d(aes(fill = factor(stat(level))), geom = "polygon")
ggplot(df, aes(x=test1, y=test2, z=sin(sqrt(test1^2 + test2^2)))) +
geom_jitter() +
geom_density_2d(aes(col = factor(..level..)), size = 0.7)
ggplot(df, aes(x=test1, y=test2, z=sin(sqrt(test1^2 + test2^2)))) +
geom_jitter() +
stat_density_2d(geom = "raster", aes(fill = stat(density)), contour = FALSE)
ggplot(df, aes(x=test1, y=test2, z=sin(sqrt(test1^2 + test2^2)))) +
geom_jitter() +
stat_density_2d(geom = "point", aes(size = stat(density)), n = 20, contour = FALSE)
|
Week1/R Basics.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import pytorch_lightning as pl
from drugexr.data_structs.vocabulary import Vocabulary
from drugexr.models.generator import Generator
from drugexr.config import constants as c
from torch.utils.data import DataLoader
import torch
import numpy as np
import pandas as pd
from rdkit import Chem
from rdkit.Chem.Draw import IPythonConsole
from rdkit.Chem import Draw
IPythonConsole.ipython_useSVG=True
from drugexr.utils import tensor_ops
from rdkit import RDLogger
RDLogger.DisableLog('rdApp.*')
# +
vocabulary = Vocabulary(vocabulary_path=c.PROC_DATA_PATH / "chembl_voc.txt")
# generator = Generator(vocabulary=vocabulary)
generator = Generator.load_from_checkpoint("../models/output/rnn/fine_tuned_lstm.ckpt", vocabulary=vocabulary)
# encoded_samples = generator.sample(1024)
# decoded_samples = [vocabulary.decode(sample) for sample in encoded_samples]
# -
sequences = generator.sample(1024)
indices = tensor_ops.unique(sequences)
sequences = sequences[indices]
smiles, valids = vocabulary.check_smiles(sequences)
frac_valid = sum(valids) / len(sequences)
frac_valid
valid_mask = np.array(valids, dtype='bool')
decoded_samples = [vocabulary.decode(sample) for sample in sequences]
decoded_samples = np.array(decoded_samples)
valid_decode_samples = decoded_samples[valid_mask]
valid_decode_samples
Draw.MolsToGridImage([Chem.MolFromSmiles(smi) for smi in valid_decode_samples[:100]])
|
notebooks/08-sample-checking.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction
#
# This notebook provides a tutorial on how to use pymatgen's basic functionality, and how to interact with the Materials API through pymatgen's high-level interface.
# Let's start with some basic imports of the classes/methods we need.
import warnings
warnings.filterwarnings('ignore')
from pymatgen import MPRester, Structure, Lattice, Element
from pprint import pprint
# ## Basic functionality of pymatgen
#
# Now, let us create the SrTiO3 cubic perovskite manually from a spacegroup.
perovskite = Structure.from_spacegroup(
"Pm-3m", # CsCl has the primitive cubic spacegroup Pm-3m
Lattice.cubic(3), # We use the convenient constructor to generate a cubic lattice with lattice parameter 3.
["Sr", "Ti", "O"], # These are the two unique species.
[[0,0,0], [0.5, 0.5, 0.5], [0.5, 0.5, 0]] # These are the fractional coordinates.
)
# Let us now see the structure.
print(perovskite)
# The Structure object provides methods to export the structure to a Crystallographic Information File (CIF) and other formats that can be opened by a wide variety of materials science software.
perovskite.to(filename="SrTiO3.cif")
# Similarly, we can read a Structure from many common file formats.
perov_from_file = Structure.from_file("SrTiO3.cif")
print(perov_from_file) # Just to confirm that we have the same structure.
# Very often, we want to manipulate an existing to create a new structure. Here, we will apply a strain in the c direction followed by a substitution of Sr with Ba to create a tetragonal form of BaTiO3.
new_structure = perovskite.copy()
new_structure.apply_strain([0, 0, 0.1])
new_structure["Sr"] = "Ba"
print(new_structure)
# We can determine the spacegroup of this new structure. We get a tetragonal space group as expected.
print(new_structure.get_space_group_info())
# Pymatgen comes with many powerful analysis tools. There are too many to go through. Here, we will present a few examples.
# Analysis 1: Plotting the XRD pattern
# %matplotlib inline
from pymatgen.analysis.diffraction.xrd import XRDCalculator
c = XRDCalculator()
c.show_plot(perovskite)
# Analysis example 2: Generating surface structures
#
# This code was used extensively in the creating of [Crystalium](http://crystalium.materialsvirtuallab.org).
from pymatgen.core.surface import generate_all_slabs
slabs = generate_all_slabs(perovskite, 2, 10, 10)
for slab in slabs:
print(slab.miller_index)
# ## Using the Materials API
#
# pymatgen provides a high-level interface to the Materials Project API called MPRester. Using MPRester, you can easily grab almost any piece of data from the Materials Project easily. Here we demonstrate some of the high level functions. To use the Materials API, you need to already have an account and an API key from the Materials Project. Get it at https://www.materialsproject.org/dashboard.
mpr = MPRester()
entries = mpr.get_entries("CsCl", inc_structure=True)
for e in entries:
print(e.structure)
print(e.structure.get_space_group_info())
print(e.energy_per_atom)
# ## Simple machine learning with Materials Project data
#
# Here, I will demonstrate a simple machine learning exercise using data from the Materials Project using pymatgen combined with numpy. Pymatgen's Element object has all the elemental melting points available. Materials Project has computed the elastic constants of all the elements. Is there a relationship between the bulk modulus of an element and its melting point?
# For the purposes of this exercise, we will be using the powerful `query` method in MPRester, which essentially allows a user to submit any type of data request to the Materials Project. We will demonstrate the basic principles, and to find out more, please go to http://bit.ly/materialsapi for more information on how to do more sophisticated queries.
# +
criteria = {
"nelements": 1, # This specifies we are getting data on materials with only one species, i.e., elements
"elasticity": {"$exists": True} # We only want elements with elasticity data.
}
properties = [
"elements",
"task_id",
"final_energy_per_atom",
"elasticity.K_VRH"
]
data = mpr.query(criteria=criteria, properties=properties)
print(len(data))
print(data[0])
# -
melting_points = []
bulk_moduli = []
for d in data:
melting_points.append(Element(d["elements"][0]).melting_point)
bulk_moduli.append(d["elasticity.K_VRH"])
# +
import seaborn as sns
import numpy as np
import matplotlib.pyplot as plt
sns.set(style="darkgrid", font='sans-serif', font_scale=2)
g = sns.jointplot(melting_points, bulk_moduli, kind="reg",
xlim=(0, 3500), ylim=(0, 400), color="m", height=8)
label = plt.xlabel("Melting point (K)")
label = plt.ylabel("K_VRH (GPa)")
# -
# We will use scipy to get the linear equation line.
from scipy import stats
# get coeffs of linear fit
slope, intercept, r_value, p_value, std_err = stats.linregress(melting_points, bulk_moduli)
print("K_VRH = {0:.1f} MP {1:.1f}".format(slope,intercept))
print("R = %.4f" % r_value)
# The problem with the above analysis is that it includes all polymorphs of the same element. Obviously, the melting point refers to the standard state of the element. We will use the ground state (lowest energy per atom) of each element as the representative data instead.
import itertools
data = sorted(data, key=lambda d: d["elements"][0])
melting_points_gs = []
bulk_moduli_gs = []
for k, g in itertools.groupby(data, key=lambda d: d["elements"][0]):
gs_data = min(g, key=lambda d: d["final_energy_per_atom"])
melting_points_gs.append(Element(k).melting_point)
bulk_moduli_gs.append(gs_data["elasticity.K_VRH"])
g = sns.jointplot(melting_points_gs, bulk_moduli_gs, kind="reg",
xlim=(0, 3500), ylim=(0, 400), color="m", height=8)
import matplotlib.pyplot as plt
label = plt.xlabel("Melting point (K)")
label = plt.ylabel("K_VRH (GPa)")
slope, intercept, r_value, p_value, std_err = stats.linregress(melting_points_gs, bulk_moduli_gs)
print("K_VRH = {0:.1f} Melting_Point {1:.1f}".format(slope,intercept))
print("R = %.4f" % r_value)
# ## Multiple linear regression
#
# Instead of assuming that the bulk modulus can be predicted from just the melting point, we will now use multiple variables, including the melting point, boiling point, atomic number, electronegativity and atomic radius.
parameters_gs = []
bulk_moduli_gs = []
for k, g in itertools.groupby(data, key=lambda d: d["elements"][0]):
gs_data = min(g, key=lambda d: d["final_energy_per_atom"])
el = Element(k)
if (not el.is_noble_gas) and el.boiling_point:
parameters_gs.append([el.melting_point, el.boiling_point, el.Z, el.X, el.atomic_radius])
bulk_moduli_gs.append(gs_data["elasticity.K_VRH"])
# +
import statsmodels.api as sm
n = len(bulk_moduli_gs)
model = sm.OLS(bulk_moduli_gs, np.concatenate((np.array(parameters_gs), np.ones((n, 1))), axis=1)).fit()
print("K_VRH = {0:.3f} Melting_Point {1:+.3f} Boiling_Point {2:+.3f}Z {3:+.3f}X {4:+.3f}r {5:+.3f}".format(
*model.params))
print("R = %.4f" % np.sqrt(model.rsquared))
# -
# The R-value improves from previous 0.7721 to the current 0.8148. However, the selection of five variables is arbitrary. Do we really need five variables?
#
import pandas as pd
f, ax = plt.subplots(figsize=(8, 6))
df = pd.DataFrame(parameters_gs, columns=["MP", "BP", "Z", "X", "r"])
sns.heatmap(df.corr(), cmap="coolwarm", vmin=-1, vmax=1, ax=ax)
ax.set_title('Variable correlation')
# Note that the **melting point (MP)** and **boiling point (BP)** are positively correlated, and **electronegativity X** and **atomic radius r** are negatively correlated.
#
# How do we automatically remove those redundant variables?
# ## Least absolute shrinkage and selection operator (LASSO)
#
# Let us now use a more sophisticated machine learning approach - LASSO.
#
# Linear models using the ordinary least squares (OLS) method finds the coefficients $
# \vec{\beta}$'s by minimizing the following objective function.
#
# $s = \frac{1}{N}\sum_i^N({y_i - \beta_0-\vec{x}_i^T\vec{\beta}})^2$
#
# where $N$ is the total number of data points $(\vec{x}, y)$.
#
# In the LASSO approach, the $L_1$ norm of $\vec{\beta}$ is also minimized and the objective function is the following:
#
# $s = \frac{1}{N}\sum_i^N({y_i - \beta_0-\vec{x}_i^T\vec{\beta}})^2 + \alpha \sum_j^M|\beta_j|$
#
# Therefore when $\alpha$ is large, some $\beta_j$'s will be reduced to zeros. We will use the LASSO implementation in the scikit-learn package.
from sklearn import linear_model
reg = linear_model.Lasso(alpha=0.3, max_iter=10000)
reg.fit(parameters_gs, bulk_moduli_gs)
print(reg.coef_)
predicted_bulk_moduli = reg.predict(parameters_gs)
from pymatgen.util.plotting import pretty_plot
plt = pretty_plot(12, 8)
plt.plot(predicted_bulk_moduli, bulk_moduli_gs, "x")
plt.xlabel("%.2f MP + %.2f BP %.2f Z + %.2f $\chi$ + %.2f $r_{atomic}$ %+.2f" % tuple((*reg.coef_, reg.intercept_)))
plt.ylabel("Computed K_VRH (GPa)")
# As expected, the atomic radius becomes zero since it is correlated with electronegativity. MP and BP remain because their absolute values are much greater than the rest variables.
|
notebooks/Using pymatgen and the Materials API.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _kg_hide-input=true
import os
import cv2
import math
import warnings
import numpy as np
import pandas as pd
import seaborn as sns
import tensorflow as tf
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
from sklearn.metrics import confusion_matrix, fbeta_score
from keras import optimizers
from keras import backend as K
from keras.models import Sequential, Model
from keras import applications
from keras.preprocessing.image import ImageDataGenerator
from keras.callbacks import LearningRateScheduler, EarlyStopping
from keras.layers import Dense, Dropout, Flatten, Conv2D, MaxPool2D, Activation, BatchNormalization, GlobalAveragePooling2D, Input
# Set seeds to make the experiment more reproducible.
from tensorflow import set_random_seed
from numpy.random import seed
set_random_seed(0)
seed(0)
# %matplotlib inline
sns.set(style="whitegrid")
warnings.filterwarnings("ignore")
# + _kg_hide-input=true
train = pd.read_csv('../input/imet-2019-fgvc6/train.csv')
labels = pd.read_csv('../input/imet-2019-fgvc6/labels.csv')
test = pd.read_csv('../input/imet-2019-fgvc6/sample_submission.csv')
train["attribute_ids"] = train["attribute_ids"].apply(lambda x:x.split(" "))
train["id"] = train["id"].apply(lambda x: x + ".png")
test["id"] = test["id"].apply(lambda x: x + ".png")
print('Number of train samples: ', train.shape[0])
print('Number of test samples: ', test.shape[0])
print('Number of labels: ', labels.shape[0])
display(train.head())
display(labels.head())
# -
# ### Model parameters
# Model parameters
BATCH_SIZE = 64
EPOCHS = 30
LEARNING_RATE = 0.0001
HEIGHT = 64
WIDTH = 64
CANAL = 3
N_CLASSES = labels.shape[0]
ES_PATIENCE = 15
DECAY_DROP = 0.5
DECAY_EPOCHS = 10
classes = list(map(str, range(N_CLASSES)))
# + _kg_hide-input=true
def f2_score_thr(threshold=0.5):
def f2_score(y_true, y_pred):
beta = 2
y_pred = K.cast(K.greater(K.clip(y_pred, 0, 1), threshold), K.floatx())
true_positives = K.sum(K.clip(y_true * y_pred, 0, 1), axis=1)
predicted_positives = K.sum(K.clip(y_pred, 0, 1), axis=1)
possible_positives = K.sum(K.clip(y_true, 0, 1), axis=1)
precision = true_positives / (predicted_positives + K.epsilon())
recall = true_positives / (possible_positives + K.epsilon())
return K.mean(((1+beta**2)*precision*recall) / ((beta**2)*precision+recall+K.epsilon()))
return f2_score
def custom_f2(y_true, y_pred):
beta = 2
tp = np.sum((y_true == 1) & (y_pred == 1))
tn = np.sum((y_true == 0) & (y_pred == 0))
fp = np.sum((y_true == 0) & (y_pred == 1))
fn = np.sum((y_true == 1) & (y_pred == 0))
p = tp / (tp + fp + K.epsilon())
r = tp / (tp + fn + K.epsilon())
f2 = (1+beta**2)*p*r / (p*beta**2 + r + 1e-15)
return f2
def step_decay(epoch):
initial_lrate = LEARNING_RATE
drop = DECAY_DROP
epochs_drop = DECAY_EPOCHS
lrate = initial_lrate * math.pow(drop, math.floor((1+epoch)/epochs_drop))
return lrate
# + _kg_hide-input=true
train_datagen=ImageDataGenerator(rescale=1./255, validation_split=0.25)
train_generator=train_datagen.flow_from_dataframe(
dataframe=train,
directory="../input/imet-2019-fgvc6/train",
x_col="id",
y_col="attribute_ids",
batch_size=BATCH_SIZE,
shuffle=True,
class_mode="categorical",
classes=classes,
target_size=(HEIGHT, WIDTH),
subset='training')
valid_generator=train_datagen.flow_from_dataframe(
dataframe=train,
directory="../input/imet-2019-fgvc6/train",
x_col="id",
y_col="attribute_ids",
batch_size=BATCH_SIZE,
shuffle=True,
class_mode="categorical",
classes=classes,
target_size=(HEIGHT, WIDTH),
subset='validation')
test_datagen = ImageDataGenerator(rescale=1./255)
test_generator = test_datagen.flow_from_dataframe(
dataframe=test,
directory = "../input/imet-2019-fgvc6/test",
x_col="id",
target_size=(HEIGHT, WIDTH),
batch_size=1,
shuffle=False,
class_mode=None)
# -
# ### Model
def create_model(input_shape, n_out):
input_tensor = Input(shape=input_shape)
base_model = applications.Xception(weights=None, include_top=False,
input_tensor=input_tensor)
base_model.load_weights('../input/xception/xception_weights_tf_dim_ordering_tf_kernels_notop.h5')
x = GlobalAveragePooling2D()(base_model.output)
x = Dropout(0.5)(x)
x = Dense(1024, activation='relu')(x)
x = Dropout(0.5)(x)
final_output = Dense(n_out, activation='sigmoid', name='final_output')(x)
model = Model(input_tensor, final_output)
return model
# + _kg_hide-output=true
# warm up model
# first: train only the top layers (which were randomly initialized)
model = create_model(input_shape=(HEIGHT, WIDTH, CANAL), n_out=N_CLASSES)
for layer in model.layers:
layer.trainable = False
for i in range(-5,0):
model.layers[i].trainable = True
optimizer = optimizers.Adam(lr=LEARNING_RATE)
metrics = ["accuracy", "categorical_accuracy"]
es = EarlyStopping(monitor='val_loss', mode='min', verbose=1, patience=ES_PATIENCE)
callbacks = [es]
model.compile(optimizer=optimizer, loss="binary_crossentropy", metrics=metrics)
model.summary()
# -
# #### Train top layers
# + _kg_hide-input=true _kg_hide-output=true
STEP_SIZE_TRAIN = train_generator.n//train_generator.batch_size
STEP_SIZE_VALID = valid_generator.n//valid_generator.batch_size
history = model.fit_generator(generator=train_generator,
steps_per_epoch=STEP_SIZE_TRAIN,
validation_data=valid_generator,
validation_steps=STEP_SIZE_VALID,
epochs=EPOCHS,
callbacks=callbacks,
verbose=2,
max_queue_size=16, workers=3, use_multiprocessing=True)
# -
# #### Fine-tune the complete model
# + _kg_hide-output=true
for i in range(-17, 0):
model.layers[i].trainable = True
metrics = ["accuracy", "categorical_accuracy"]
lrate = LearningRateScheduler(step_decay)
es = EarlyStopping(monitor='val_loss', mode='min', verbose=1, patience=(ES_PATIENCE))
callbacks = [es, lrate]
optimizer = optimizers.SGD(lr=0.0001, momentum=0.9)
model.compile(optimizer=optimizer, loss="binary_crossentropy", metrics=metrics)
model.summary()
# + _kg_hide-input=true _kg_hide-output=true
STEP_SIZE_TRAIN = train_generator.n//train_generator.batch_size
STEP_SIZE_VALID = valid_generator.n//valid_generator.batch_size
history = model.fit_generator(generator=train_generator,
steps_per_epoch=STEP_SIZE_TRAIN,
validation_data=valid_generator,
validation_steps=STEP_SIZE_VALID,
epochs=EPOCHS,
callbacks=callbacks,
verbose=2,
max_queue_size=16, workers=3, use_multiprocessing=True)
# -
# ### Complete model graph loss
# + _kg_hide-input=true
sns.set_style("whitegrid")
fig, (ax1, ax2, ax3) = plt.subplots(1, 3, sharex='col', figsize=(20,7))
ax1.plot(history.history['loss'], label='Train loss')
ax1.plot(history.history['val_loss'], label='Validation loss')
ax1.legend(loc='best')
ax1.set_title('Loss')
ax2.plot(history.history['acc'], label='Train Accuracy')
ax2.plot(history.history['val_acc'], label='Validation accuracy')
ax2.legend(loc='best')
ax2.set_title('Accuracy')
ax3.plot(history.history['categorical_accuracy'], label='Train Cat Accuracy')
ax3.plot(history.history['val_categorical_accuracy'], label='Validation Cat Accuracy')
ax3.legend(loc='best')
ax3.set_title('Cat Accuracy')
plt.xlabel('Epochs')
sns.despine()
plt.show()
# -
# ### Find best threshold value
# + _kg_hide-input=false
lastFullValPred = np.empty((0, N_CLASSES))
lastFullValLabels = np.empty((0, N_CLASSES))
for i in range(STEP_SIZE_VALID+1):
im, lbl = next(valid_generator)
scores = model.predict(im, batch_size=valid_generator.batch_size)
lastFullValPred = np.append(lastFullValPred, scores, axis=0)
lastFullValLabels = np.append(lastFullValLabels, lbl, axis=0)
print(lastFullValPred.shape, lastFullValLabels.shape)
# + _kg_hide-input=true
def find_best_fixed_threshold(preds, targs, do_plot=True):
score = []
thrs = np.arange(0, 0.5, 0.01)
for thr in thrs:
score.append(custom_f2(targs, (preds > thr).astype(int)))
score = np.array(score)
pm = score.argmax()
best_thr, best_score = thrs[pm], score[pm].item()
print(f'thr={best_thr:.3f}', f'F2={best_score:.3f}')
if do_plot:
plt.plot(thrs, score)
plt.vlines(x=best_thr, ymin=score.min(), ymax=score.max())
plt.text(best_thr+0.03, best_score-0.01, f'$F_{2}=${best_score:.3f}', fontsize=14);
plt.show()
return best_thr, best_score
threshold, best_score = find_best_fixed_threshold(lastFullValPred, lastFullValLabels, do_plot=True)
# -
# ### Apply model to test set and output predictions
# + _kg_hide-input=true
test_generator.reset()
STEP_SIZE_TEST = test_generator.n//test_generator.batch_size
preds = model.predict_generator(test_generator, steps=STEP_SIZE_TEST)
# + _kg_hide-input=true
predictions = []
for pred_ar in preds:
valid = []
for idx, pred in enumerate(pred_ar):
if pred > threshold:
valid.append(idx)
if len(valid) == 0:
valid.append(np.argmax(pred_ar))
predictions.append(valid)
# + _kg_hide-input=true
filenames = test_generator.filenames
label_map = {valid_generator.class_indices[k] : k for k in valid_generator.class_indices}
results = pd.DataFrame({'id':filenames, 'attribute_ids':predictions})
results['id'] = results['id'].map(lambda x: str(x)[:-4])
results['attribute_ids'] = results['attribute_ids'].apply(lambda x: list(map(label_map.get, x)))
results["attribute_ids"] = results["attribute_ids"].apply(lambda x: ' '.join(x))
results.to_csv('submission.csv',index=False)
results.head(10)
|
Model backlog/Deep Learning/Xception/[73th] - Fine-tune - Xception - Top 2 conv2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
plt.style.use('fivethirtyeight')
import numpy as np
import scipy.special as ss
x = np.arange(0,0.5, 0.001)
y = []
for alpha in x:
sum_ = 0
for j in range(17, 33):
sum_ += ss.binom(32, j) * np.power(alpha, j+2) * np.power(1-alpha, 32-j)
y.append(sum_)
f, ax = plt.subplots(figsize=(7,4))
ax.plot(x, y)
ax.set_facecolor('white')
f.set_facecolor('white')
ax.set_xlabel(r'$\alpha$')
ax.set_ylabel(r'probability of attack')
plt.show()
x = np.arange(0,0.3, 0.001)
y = []
for alpha in x:
sum_ = 0
for j in range(17, 33):
sum_ += ss.binom(32, j) * np.power(alpha, j+2) * np.power(1-alpha, 32-j)
y.append(sum_)
f, ax = plt.subplots(figsize=(7,4))
ax.plot(x, y)
ax.set_facecolor('white')
f.set_facecolor('white')
ax.set_xlabel(r'$\alpha$')
ax.set_ylabel(r'probability of attack')
plt.show()
x = np.arange(0,0.5, 0.001)
y = []
for alpha in x:
sum_ = 0
for j in range(17, 33):
sum_ += ss.binom(32, j) * np.power(alpha, j+2) * np.power(1-alpha, 32-j)
y.append(sum_ * 525600)
sim_x = np.arange(0,0.5, 0.025)
sim_y = [0, 0, 0, 0, 0, 0, 0, 0, 1, 3, 17, 76, 256, 697, 1739, 3914, 7634, 14382, 24078, 38327]
f, ax = plt.subplots(figsize=(7,4))
ax.plot(x, y, 'g--', linewidth=1, alpha=0.5, label='theoretical')
ax.set_facecolor('white')
f.set_facecolor('white')
ax.set_xlabel(r'$\alpha$')
ax.set_ylabel(r'extra blocks (one year)')
ax.plot(sim_x, sim_y, 'rx', label='simulated')
plt.legend(facecolor='white', framealpha=1)
plt.show()
0.4*525600
np.arange(0, 0.5, 0.025)
|
stake/theoretical.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
df = pd.read_csv('Dataset.csv')
df.head()
# +
df['All usual residents'] = pd.to_numeric(df['All usual residents'].str.replace(',', ''))
df['Passports held: No passport'] = pd.to_numeric(df['Passports held: No passport'].str.replace(',', ''))
df['Passports held: United Kingdom only'] = pd.to_numeric(df['Passports held: United Kingdom only'].str.replace(',', ''))
df['Passports held: Ireland only'] = pd.to_numeric(df['Passports held: Ireland only'].str.replace(',', ''))
df['Passports held: United Kingdom and Ireland only'] = pd.to_numeric(df['Passports held: United Kingdom and Ireland only'].str.replace(',', ''))
df['Passports held: United Kingdom and other (not Ireland)'] = pd.to_numeric(df['Passports held: United Kingdom and other (not Ireland)'].str.replace(',', ''))
df['Passports held: Ireland and other (not United Kingdom)'] = pd.to_numeric(df['Passports held: Ireland and other (not United Kingdom)'].str.replace(',', ''))
df['Passports held: EU/EEA (not United Kingdom or Ireland)'] = pd.to_numeric(df['Passports held: EU/EEA (not United Kingdom or Ireland)'].str.replace(',', ''))
df['Passports held: Other'] = pd.to_numeric(df['Passports held: Other'].str.replace(',', ''))
df.head()
# -
df.head()
df
|
Passports.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1><center>Non Linear Regression Analysis</center></h1>
# If the data shows a curvy trend, then linear regression will not produce very accurate results when compared to a non-linear regression because, as the name implies, linear regression presumes that the data is linear.
# Let's learn about non linear regressions and apply an example on python. In this notebook, we fit a non-linear model to the datapoints corrensponding to China's GDP from 1960 to 2014.
# <h2 id="importing_libraries">Importing required libraries</h2>
# + jupyter={"outputs_hidden": false}
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# -
# Though Linear regression is very good to solve many problems, it cannot be used for all datasets. First recall how linear regression, could model a dataset. It models a linear relation between a dependent variable y and independent variable x. It had a simple equation, of degree 1, for example y = $2x$ + 3.
# +
x = np.arange(-5.0, 5.0, 0.1)
##You can adjust the slope and intercept to verify the changes in the graph
y = 2*(x) + 3
y_noise = 2 * np.random.normal(size=x.size)
ydata = y + y_noise
#plt.figure(figsize=(8,6))
plt.plot(x, ydata, 'bo')
plt.plot(x,y, 'r')
plt.ylabel('Dependent Variable')
plt.xlabel('Indepdendent Variable')
plt.show()
# -
# Non-linear regressions are a relationship between independent variables $x$ and a dependent variable $y$ which result in a non-linear function modeled data. Essentially any relationship that is not linear can be termed as non-linear, and is usually represented by the polynomial of $k$ degrees (maximum power of $x$).
#
# $$ \ y = a x^3 + b x^2 + c x + d \ $$
#
# Non-linear functions can have elements like exponentials, logarithms, fractions, and others. For example: $$ y = \log(x)$$
#
# Or even, more complicated such as :
# $$ y = \log(a x^3 + b x^2 + c x + d)$$
# Let's take a look at a cubic function's graph.
# + jupyter={"outputs_hidden": false}
x = np.arange(-5.0, 5.0, 0.1)
##You can adjust the slope and intercept to verify the changes in the graph
y = 1*(x**3) + 1*(x**2) + 1*x + 3
y_noise = 20 * np.random.normal(size=x.size)
ydata = y + y_noise
plt.plot(x, ydata, 'bo')
plt.plot(x,y, 'r')
plt.ylabel('Dependent Variable')
plt.xlabel('Indepdendent Variable')
plt.show()
# -
# As you can see, this function has $x^3$ and $x^2$ as independent variables. Also, the graphic of this function is not a straight line over the 2D plane. So this is a non-linear function.
# Some other types of non-linear functions are:
# ### Quadratic
# $$ Y = X^2 $$
# + jupyter={"outputs_hidden": false}
x = np.arange(-5.0, 5.0, 0.1)
##You can adjust the slope and intercept to verify the changes in the graph
y = np.power(x,2)
y_noise = 2 * np.random.normal(size=x.size)
ydata = y + y_noise
plt.plot(x, ydata, 'bo')
plt.plot(x,y, 'r')
plt.ylabel('Dependent Variable')
plt.xlabel('Indepdendent Variable')
plt.show()
# -
# ### Exponential
# An exponential function with base c is defined by $$ Y = a + b c^X$$ where b ≠0, c > 0 , c ≠1, and x is any real number. The base, c, is constant and the exponent, x, is a variable.
#
#
# + jupyter={"outputs_hidden": false}
X = np.arange(-5.0, 5.0, 0.1)
##You can adjust the slope and intercept to verify the changes in the graph
Y= np.exp(X)
plt.plot(X,Y)
plt.ylabel('Dependent Variable')
plt.xlabel('Indepdendent Variable')
plt.show()
# -
# ### Logarithmic
#
# The response $y$ is a results of applying logarithmic map from input $x$'s to output variable $y$. It is one of the simplest form of __log()__: i.e. $$ y = \log(x)$$
#
# Please consider that instead of $x$, we can use $X$, which can be polynomial representation of the $x$'s. In general form it would be written as
# \begin{equation}
# y = \log(X)
# \end{equation}
# + jupyter={"outputs_hidden": false}
X = np.arange(-5.0, 5.0, 0.1)
Y = np.log(X)
plt.plot(X,Y)
plt.ylabel('Dependent Variable')
plt.xlabel('Indepdendent Variable')
plt.show()
# -
# ### Sigmoidal/Logistic
# $$ Y = a + \frac{b}{1+ c^{(X-d)}}$$
# +
X = np.arange(-5.0, 5.0, 0.1)
Y = 1-4/(1+np.power(3, X-2))
plt.plot(X,Y)
plt.ylabel('Dependent Variable')
plt.xlabel('Indepdendent Variable')
plt.show()
# -
# <a id="ref2"></a>
# # Non-Linear Regression example
# For an example, we're going to try and fit a non-linear model to the datapoints corresponding to China's GDP from 1960 to 2014. We download a dataset with two columns, the first, a year between 1960 and 2014, the second, China's corresponding annual gross domestic income in US dollars for that year.
# + jupyter={"outputs_hidden": false}
import numpy as np
import pandas as pd
#downloading dataset
# !wget -nv -O china_gdp.csv https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/ML0101ENv3/labs/china_gdp.csv
df = pd.read_csv("china_gdp.csv")
df.head(10)
# -
# __Did you know?__ When it comes to Machine Learning, you will likely be working with large datasets. As a business, where can you host your data? IBM is offering a unique opportunity for businesses, with 10 Tb of IBM Cloud Object Storage: [Sign up now for free](http://cocl.us/ML0101EN-IBM-Offer-CC)
# ### Plotting the Dataset ###
# This is what the datapoints look like. It kind of looks like an either logistic or exponential function. The growth starts off slow, then from 2005 on forward, the growth is very significant. And finally, it decelerate slightly in the 2010s.
# + jupyter={"outputs_hidden": false}
plt.figure(figsize=(8,5))
x_data, y_data = (df["Year"].values, df["Value"].values)
plt.plot(x_data, y_data, 'ro')
plt.ylabel('GDP')
plt.xlabel('Year')
plt.show()
# -
# ### Choosing a model ###
#
# From an initial look at the plot, we determine that the logistic function could be a good approximation,
# since it has the property of starting with a slow growth, increasing growth in the middle, and then decreasing again at the end; as illustrated below:
# + jupyter={"outputs_hidden": false}
X = np.arange(-5.0, 5.0, 0.1)
Y = 1.0 / (1.0 + np.exp(-X))
plt.plot(X,Y)
plt.ylabel('Dependent Variable')
plt.xlabel('Indepdendent Variable')
plt.show()
# -
#
#
# The formula for the logistic function is the following:
#
# $$ \hat{Y} = \frac1{1+e^{\beta_1(X-\beta_2)}}$$
#
# $\beta_1$: Controls the curve's steepness,
#
# $\beta_2$: Slides the curve on the x-axis.
# ### Building The Model ###
# Now, let's build our regression model and initialize its parameters.
def sigmoid(x, Beta_1, Beta_2):
y = 1 / (1 + np.exp(-Beta_1*(x-Beta_2)))
return y
# Lets look at a sample sigmoid line that might fit with the data:
# + jupyter={"outputs_hidden": false}
beta_1 = 0.10
beta_2 = 1990.0
#logistic function
Y_pred = sigmoid(x_data, beta_1 , beta_2)
#plot initial prediction against datapoints
plt.plot(x_data, Y_pred*15000000000000.)
plt.plot(x_data, y_data, 'ro')
# -
# Our task here is to find the best parameters for our model. Lets first normalize our x and y:
# Lets normalize our data
xdata =x_data/max(x_data)
ydata =y_data/max(y_data)
# #### How we find the best parameters for our fit line?
# we can use __curve_fit__ which uses non-linear least squares to fit our sigmoid function, to data. Optimal values for the parameters so that the sum of the squared residuals of sigmoid(xdata, *popt) - ydata is minimized.
#
# popt are our optimized parameters.
from scipy.optimize import curve_fit
popt, pcov = curve_fit(sigmoid, xdata, ydata)
#print the final parameters
print(" beta_1 = %f, beta_2 = %f" % (popt[0], popt[1]))
# Now we plot our resulting regression model.
x = np.linspace(1960, 2015, 55)
x = x/max(x)
plt.figure(figsize=(8,5))
y = sigmoid(x, *popt)
plt.plot(xdata, ydata, 'ro', label='data')
plt.plot(x,y, linewidth=3.0, label='fit')
plt.legend(loc='best')
plt.ylabel('GDP')
plt.xlabel('Year')
plt.show()
# ## Practice
# Can you calculate what is the accuracy of our model?
# +
# write your code here
# split data into train/test
msk = np.random.rand(len(df)) < 0.8
train_x = xdata[msk]
test_x = xdata[~msk]
train_y = ydata[msk]
test_y = ydata[~msk]
# build the model using train set
popt, pcov = curve_fit(sigmoid, train_x, train_y)
# predict using test set
y_hat = sigmoid(test_x, *popt)
# evaluation
print("Mean absolute error: %.2f" % np.mean(np.absolute(y_hat - test_y)))
print("Residual sum of squares (MSE): %.2f" % np.mean((y_hat - test_y) ** 2))
from sklearn.metrics import r2_score
print("R2-score: %.2f" % r2_score(y_hat , test_y) )
|
Machine learning with Python/Week2/NoneLinearRegression.ipynb
|