code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Importing all the dependencies
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.model_selection import train_test_split
from sklearn import metrics
from xgboost import XGBRegressor
from sklearn.preprocessing import LabelEncoder
# # Data collection and analysis
df = pd.read_csv('C:/Users/Hemant/jupyter_codes/ML Project 1/Big mart sales prediction/train.csv')
#print the fisrt 5 rows of the dataset
'''
FD = food
DR = drink
NC = non consumable
'''
df.head()
# print the last 5 rows of the dataset
df.tail()
# shape of the dataset
df.shape
# getting some info about the dataset
df.info()
#checking for any missing values
df.isnull().sum()
# stastical measure of the dataset
df.describe()
#checking for categorical data in diff object type columns
objlist = df.select_dtypes('object').columns
for i in objlist:
print(f'\n{i}')
print(df[i].value_counts(), end = '\n')
# Handling the missing values
#
# Mean ---> Average value
# Mode ---> Most repeated value
# mean value of 'Item weight' collumn
mean_value = df['Item_Weight'].mean()
# filling the missing value with mean in 'item weight' column
df['Item_Weight'].fillna(mean_value, inplace = True)
#checking for missing values
df.isnull().sum()
# replacing the missing value with mode in 'Outlet Size' column
mode_value = df.pivot_table(values = 'Outlet_Size', columns = 'Outlet_Type', aggfunc = (lambda x : x.mode()[0]))
print(mode_value)
missing_values = df['Outlet_Size'].isnull()
df.loc[missing_values, 'Outlet_Size'] = df.loc[missing_values, 'Outlet_Type'].apply(lambda x : mode_value[x])
#checking for missing values
df.isnull().sum()
# Data analysis
# stastical measure of the data
df.describe()
# Numerical features
sns.set_style(style = 'darkgrid')
#item weight distribution
plt.figure(figsize = (6,6))
sns.displot(df['Item_Weight'], kde= True)
plt.show()
#item visibility distribution
plt.figure(figsize = (6,6))
sns.displot(df['Item_Visibility'], kde= True)
plt.show()
#item MRP distribution
plt.figure(figsize = (6,6))
sns.displot(df['Item_MRP'], kde= True)
plt.show()
#Item_Outlet_Sales distribution
plt.figure(figsize = (6,6))
sns.displot(df['Item_Outlet_Sales'], kde= True)
plt.show()
#Outlet_Establishment_Year distribution
plt.figure(figsize = (6,6))
sns.countplot(x= 'Outlet_Establishment_Year', data = df)
plt.show()
# Categoruical features
#Item_Fat_Content distribution
plt.figure(figsize = (6,6))
sns.countplot(x= 'Item_Fat_Content', data = df)
plt.show()
# Item_Type distribution
plt.figure(figsize = (30,6))
sns.countplot(x= 'Item_Type', data = df)
plt.show()
# Outlet location type distribution
plt.figure(figsize = (6,6))
sns.countplot(x = 'Outlet_Location_Type', data = df)
plt.show()
# # Data preprocessing
df.head()
df['Item_Fat_Content'].value_counts()
df.replace({'Item_Fat_Content' : {'low fat' : 'Low Fat', 'LF' : 'Low Fat', 'reg' : 'Regular'}}, inplace = True)
df['Item_Fat_Content'].value_counts()
# Label Encoding
# +
encoder = LabelEncoder()
objlist = df.select_dtypes('object').columns
for i in objlist:
df[i] = encoder.fit_transform(df[i])
# -
df.head()
correlation = df.corr()
plt.figure(figsize = (20,20))
sns.heatmap(correlation , cbar = True, cmap = 'Blues',square = True, annot = True, fmt = '.1f', annot_kws = {'size' : 8})
# Splitting features and targets
X = df.drop(columns = 'Item_Outlet_Sales' ,axis = 1)
Y = df['Item_Outlet_Sales']
# # Splitting the data into training and testing data
x_train, x_test, y_train, y_test = train_test_split(X, Y, test_size = .2, random_state = 6)
print(x_train.shape, x_test.shape)
print(y_train.shape, y_test.shape)
# # Machine learning model
model = XGBRegressor()
model.fit(x_train, y_train)
# Model evaluatuion on training data
# +
train_prediction = model.predict(x_train)
accuracy_training = metrics.r2_score(y_train, train_prediction)
print('R SQUARED ERROR OF TRAINING DATA :', accuracy_training)
# -
# Model evaluatuion on testing data
# +
test_prediction = model.predict(x_test)
accuracy_testing = metrics.r2_score(y_test, test_prediction)
print('R SQUARED ERROR OF TESTING DATA :', accuracy_testing)
# -
| Sales prediction model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <NAME> 1806554 (T&T lab) Test
import collections
with open("wordanagram.txt", "r") as file:
allopt = file.read()
choices = list(map(str, allopt.split()))
r = []
m = len(choices)
for i in choices:
res = ''.join(sorted(i))
r.append(res)
res = ''
y = set(r)
n = len(y)
occurrences = collections.Counter(r)
print(occurrences)
print("no of anagrams",": ",m-n)
# # 2
# +
# x = ["ram","sham","cool","folo","empty","hut"]
# words = ["ram"]
# y = list(lambda x: [' '.join(w for w in x.split() if w not in words)])
# print(y)
#x = lambda x: [x for x in x if x not in ["name", "is", "m"]]
# -
def isSortedlist(nums, k=lambda x: x):
for i, e in enumerate(nums[1:]):
if k(e) < k(nums[i]):
return False
return True
n1 = [1,2,4,6,8,10,12,14,16,17]
print(n1)
print(isSortedlist(n1))
# +
# -
import numpy as np
import pandas as pd
df = pd.read_csv('TestData.csv')
df.head()
import pandas as pd
import numpy as np
df = pd.read_csv('TestData.csv', names=['a', 'b', 'c', 'd'])
df['x'] = np.sqrt(df['a']*2 + df['b']*2)
df['y'] = df['a'] + df['b'] + df['c'] + df['d']
df['y'] = df['y'].mean()
df.drop(['a', 'b', 'c', 'd'], axis=1, inplace=True)
df
| T&T/python scripts/eval_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Best Practices for Preprocessing Natural Language Data
# In this notebook, we improve the quality of our Project Gutenberg word vectors by adopting best-practices for preprocessing natural language data.
#
# **N.B.:** Some, all or none of these preprocessing steps may be helpful to a given downstream application.
# #### Load dependencies
# +
import nltk
from nltk import word_tokenize, sent_tokenize
from nltk.corpus import stopwords
from nltk.stem.porter import *
nltk.download('gutenberg')
nltk.download('punkt')
nltk.download('stopwords')
import string
import gensim
from gensim.models.phrases import Phraser, Phrases
from gensim.models.word2vec import Word2Vec
from sklearn.manifold import TSNE
import pandas as pd
from bokeh.io import output_notebook, output_file
from bokeh.plotting import show, figure
# %matplotlib inline
# -
# #### Load Data
from nltk.corpus import gutenberg
len(gutenberg.fileids())
gutenberg.fileids()
len(gutenberg.words())
gberg_sent_tokens = sent_tokenize(gutenberg.raw())
gberg_sent_tokens[0:6]
gberg_sent_tokens[1]
word_tokenize(gberg_sent_tokens[1])
word_tokenize(gberg_sent_tokens[1])[14]
# a convenient method that handles newlines, as well as tokenizing sentences and words in one shot
gberg_sents = gutenberg.sents()
gberg_sents[0:6]
gberg_sents[4][14]
# #### Iteratively preprocess a sentence
# ##### a tokenized sentence:
gberg_sents[4]
# ##### to lowercase:
[w.lower() for w in gberg_sents[4]]
# ##### remove stopwords and punctuation:
stpwrds = stopwords.words('english') + list(string.punctuation)
stpwrds
[w.lower() for w in gberg_sents[4] if w.lower() not in stpwrds]
# ##### stem words:
stemmer = PorterStemmer()
[stemmer.stem(w.lower()) for w in gberg_sents[4]
if w.lower() not in stpwrds]
# ##### handle bigram collocations:
phrases = Phrases(gberg_sents) # train detector
bigram = Phraser(phrases) # create a more efficient Phraser object for transforming sentences
bigram.phrasegrams # output count and score of each bigram
tokenized_sentence = "Jon lives in New York City".split()
tokenized_sentence
bigram[tokenized_sentence]
# #### Preprocess the corpus
# as in Maas et al. (2001):
# - leave in stop words ("indicative of sentiment")
# - no stemming ("model learns similar representations of words of the same stem when data suggests it")
lower_sents = []
for s in gberg_sents:
lower_sents.append([w.lower() for w in s if w.lower()
not in list(string.punctuation)])
lower_sents[0:5]
lower_bigram = Phraser(Phrases(lower_sents))
lower_bigram.phrasegrams # miss taylor, mr woodhouse, mr weston
lower_bigram["jon lives in new york city".split()]
lower_bigram = Phraser(Phrases(lower_sents,
min_count=32, threshold=64))
lower_bigram.phrasegrams
clean_sents = []
for s in lower_sents:
clean_sents.append(lower_bigram[s])
clean_sents[0:9]
clean_sents[6]
# #### Run word2vec
# +
# max_vocab_size can be used instead of min_count (which has increased here)
# model = Word2Vec(sentences=clean_sents, size=64,
# sg=1, window=10, iter=5,
# min_count=10, workers=4)
# model.save('clean_gutenberg_model.w2v')
# -
# #### Explore model
# skip re-training the model with the next line:
model = gensim.models.Word2Vec.load('clean_gutenberg_model.w2v')
len(model.wv.vocab) # would be 17k if we carried out no preprocessing
model.wv['dog']
len(model.wv['dog'])
model.wv.most_similar('dog', topn=3)
model.wv.most_similar('eat', topn=3)
model.wv.most_similar('day', topn=3)
model.wv.most_similar('father', topn=3)
model.wv.most_similar('ma_am', topn=3)
model.wv.doesnt_match("mother father sister brother dog".split())
model.wv.similarity('father', 'dog')
model.wv.most_similar(positive=['father', 'woman'], negative=['man'])
model.wv.most_similar(positive=['husband', 'woman'], negative=['man'])
# #### Reduce word vector dimensionality with t-SNE
# +
# tsne = TSNE(n_components=2, n_iter=1000)
# +
# X_2d = tsne.fit_transform(model.wv[model.wv.vocab])
# +
# coords_df = pd.DataFrame(X_2d, columns=['x','y'])
# coords_df['token'] = model.wv.vocab.keys()
# +
# coords_df.head()
# +
# coords_df.to_csv('clean_gutenberg_tsne.csv', index=False)
# -
# #### Visualise
coords_df = pd.read_csv('clean_gutenberg_tsne.csv')
_ = coords_df.plot.scatter('x', 'y', figsize=(12,12),
marker='.', s=10, alpha=0.2)
output_notebook()
subset_df = coords_df.sample(n=5000)
p = figure(plot_width=800, plot_height=800)
_ = p.text(x=subset_df.x, y=subset_df.y, text=subset_df.token)
show(p)
# +
# output_file() here
| notebooks/natural_language_preprocessing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from rdkit import Chem
lines = open('data/small.smi').readlines()
mols = [Chem.MolFromSmiles(x) for x in lines]
r_info = mols[0].GetRingInfo()
from __future__ import print_function
from rdkit import Chem
from rdkit.Chem.Draw import IPythonConsole
from IPython.display import SVG
import time
print(time.asctime())
from rdkit.Chem import rdDepictor
from rdkit.Chem.Draw import rdMolDraw2D
m = mols[0]
m
# +
import itertools
# NOTE(LESWING) this version does not handle bridged rings correctly see version later
flatten = lambda l: [item for sublist in l for item in sublist]
r_info = m.GetRingInfo()
v1 = set()
bond_rings = set(flatten(r_info.BondRings()))
all_bonds = [x.GetIdx() for x in m.GetBonds()]
non_ring_bonds = set(all_bonds) - bond_rings
v1 = set([tuple(sorted(
[m.GetBonds()[x].GetBeginAtom().GetIdx(), m.GetBonds()[x].GetEndAtom().GetIdx()]))
for x in non_ring_bonds])
v2 = set()
v2.update([tuple(sorted((x))) for x in r_info.AtomRings()])
to_merge = set()
for r1, r2 in itertools.product(v2, repeat=2):
if r1 >= r2:
continue
intersection = set(r1).intersection(set(r2))
if len(intersection) >= 3:
to_merge.add((r1,r2))
for r1, r2 in to_merge:
v2.remove(r1)
v2.remove(r2)
s1 = set()
s1.update(r1)
s1.update(r2)
v2.add(s1)
all_clusters = set()
all_clusters.update(v1)
all_clusters.update(v2)
v0 = set()
for atom in m.GetAtoms():
atom_id = atom.GetIdx()
count = sum([
atom_id in x for x in all_clusters
])
if count >= 3:
v0.add(atom_id)
# -
def draw_cluster(m, atom_ids):
colors = {}
for atom in atom_ids:
colors[atom] = (1, 0, 0)
drawer = rdMolDraw2D.MolDraw2DSVG(400,200)
drawer.DrawMolecule(m,highlightAtoms=atom_ids,highlightAtomColors=colors)
drawer.FinishDrawing()
svg = drawer.GetDrawingText().replace('svg:','')
return SVG(svg)
from rdkit.Chem import AllChem
AllChem.Compute2DCoords(m)
draw_cluster(m, list(all_clusters)[0])
draw_cluster(m, list(all_clusters)[1])
draw_cluster(m, list(all_clusters)[2])
draw_cluster(m, list(all_clusters)[3])
draw_cluster(m, list(all_clusters)[4])
draw_cluster(m, list(all_clusters)[5])
draw_cluster(m, list(all_clusters)[6])
draw_cluster(m, list(all_clusters)[7])
draw_cluster(m, list(all_clusters)[8])
draw_cluster(m, list(all_clusters)[9])
draw_cluster(m, list(all_clusters)[10])
draw_cluster(m, list(all_clusters)[11])
draw_cluster(m, list(all_clusters)[12])
from rdkit.Chem import EditableMol
def create_substructure(m, atom_ids):
emol = EditableMol(m)
to_remove_ids = [x.GetIdx() for x in m.GetAtoms()]
for atom_id in atom_ids:
to_remove_ids.remove(atom_id)
to_remove_ids.reverse()
for atom_id in to_remove_ids:
emol.RemoveAtom(atom_id)
return emol.GetMol()
frag = create_substructure(m, list(all_clusters)[10])
Chem.MolToSmiles(frag)
m = Chem.MolFromSmiles('Cc1c(Br)c(C(=O)NC(C)C23CC4CC(CC(C4)C2)C3)nn1C')
# +
import itertools
import networkx as nx
flatten = lambda l: [item for sublist in l for item in sublist]
r_info = m.GetRingInfo()
v1 = set()
bond_rings = set(flatten(r_info.BondRings()))
all_bonds = [x.GetIdx() for x in m.GetBonds()]
non_ring_bonds = set(all_bonds) - bond_rings
v1 = set([tuple(sorted(
[m.GetBonds()[x].GetBeginAtom().GetIdx(), m.GetBonds()[x].GetEndAtom().GetIdx()]))
for x in non_ring_bonds])
v2 = set()
v2.update([tuple(sorted((x))) for x in r_info.AtomRings()])
to_merge = set()
for r1, r2 in itertools.product(v2, repeat=2):
if r1 >= r2:
continue
intersection = set(r1).intersection(set(r2))
if len(intersection) >= 3:
to_merge.add((r1,r2))
g = nx.Graph()
for f, t in to_merge:
g.add_edge(f, t)
graphs = list(nx.connected_component_subgraphs(g))
to_merge = [list(x.nodes) for x in graphs]
for merge_keys in to_merge:
s1 = set()
for merge_key in merge_keys:
v2.remove(merge_key)
s1.update(merge_key)
s1 = tuple(sorted(list(s1)))
v2.add(s1)
all_clusters = set()
all_clusters.update(v1)
all_clusters.update(v2)
v0 = set()
for atom in m.GetAtoms():
atom_id = atom.GetIdx()
count = sum([
atom_id in x for x in all_clusters
])
if count >= 3:
v0.add(atom_id)
# -
AllChem.Compute2DCoords(m)
m
draw_cluster(m, list(all_clusters)[0])
draw_cluster(m, list(all_clusters)[1])
draw_cluster(m, list(all_clusters)[2])
draw_cluster(m, list(all_clusters)[3])
draw_cluster(m, list(all_clusters)[4])
draw_cluster(m, list(all_clusters)[5])
draw_cluster(m, list(all_clusters)[6])
draw_cluster(m, list(all_clusters)[7])
draw_cluster(m, list(all_clusters)[8])
draw_cluster(m, list(all_clusters)[9])
list(g.nodes())
import json
fragments = json.load(open('data/fragments.txt'))
# +
mols = [Chem.MolFromSmiles(x, sanitize=False) for x in fragments if x is not None]
neutral_smiles = set()
neutral_mols = []
for mol in mols:
if mol is None:
continue
for atom in mol.GetAtoms():
atom.SetFormalCharge(0)
neutral_mols.append(mol)
neutral_smiles.add(Chem.MolToSmiles(mol))
# -
neutral_mols
mols
len(neutral_smile780s)
with open('data/to_ld_fragments.csv', 'w') as fout:
fout.write('smiles\n')
for frag in fragments:
fout.write("%s\n" % frag)
m1 = Chem.MolFromSmiles(fragments[0])
atom = m1.GetAtoms()[0]
atom.SetFormalCharge(0)
from __future__ import print_function
from rdkit import Chem
from rdkit.Chem.Draw import IPythonConsole
from IPython.display import SVG
import time
print(time.asctime())
from rdkit.Chem import rdDepictor
from rdkit.Chem.Draw import rdMolDraw2D
import json
from rdkit import Chem
m = json.loads(open('data/frag_with_aromatic.json').read())
mols = [Chem.MolFromSmiles(x, sanitize=False) for x in m]
mols[11]
my_mol = open('data/250k_rndm_zinc_drugs_clean.smi').readlines()[0]
my_mol = Chem.MolFromSmiles(my_mol)
my_mol = Chem.AddHs(my_mol)
my_mol
fragments = json.loads(open('data/frag_with_aromatic.json').read())
fragments = [Chem.MolFromSmiles(x, sanitize=False) for x in fragments]
fragments[0]
fragments[1]
fragments[2]
fragments[3]
fragments[4]
fragments[5]
fragments[6]
fragments[7]
fragments[8]
fragments[9]
fragments[10]
fragments[11]
fragments = json.loads(open('data/frag_with_aromatic.json').read())
fragments = [Chem.MolFromSmiles(x, sanitize=False) for x in fragments]
from rdkit.Chem import Descriptors
for mol in fragments:
try:
Chem.SanitizeMol(mol)
except:
pass
weights = [Descriptors.MolWt(x) for x in fragments]
print(len(set(weights)))
weights = sorted(zip(weights, fragments), key=lambda x: x[0])
weights[:10]
weights[0][1]
weights[1][1]
weights[2][1]
weights[3][1]
print(Chem.MolToSmiles(weights[4][1]))
weights[4][1]
weights[5][1]
weights[6][1]
weights[7][1]
weights[8][1]
weights[9][1]
weights[10][1]
weights[11][1]
weights[12][1]
weights[-300][1]
weights[-299][1]
weights[-298][1]
len(set([x[0] for x in weights]))
len(fragments)
def has_ring(x):
r_info = x.GetRingInfo()
return len(r_info.BondRings()) > 0
has_ring(fragments[-1])
has_ring(fragments[0])
i = 0
while has_ring(fragments[i]):
i+=1
fragments[i]
# +
from rdkit.Chem import EditableMol
def remove_any_atom_bonds(m):
if not has_ring(m):
return m
emol = EditableMol(Chem.MolFromSmiles(""))
atom_map = {}
for atom in m.GetAtoms():
if atom.GetAtomicNum() == 0:
continue
new_idx = emol.AddAtom(Chem.Atom(atom.GetAtomicNum()))
atom_map[atom.GetIdx()] = new_idx
for bond in m.GetBonds():
start, end = bond.GetBeginAtom(), bond.GetEndAtom()
if start.GetAtomicNum() == 0:
continue
if end.GetAtomicNum() == 0:
continue
start, end = atom_map[start.GetIdx()], atom_map[end.GetIdx()]
emol.AddBond(start, end, bond.GetBondType())
return emol.GetMol()
# -
rings_only = [remove_any_atom_bonds(x) for x in fragments]
rings_only_smiles = [Chem.MolToSmiles(x) for x in rings_only]
rings_only = [Chem.MolFromSmiles(x, sanitize=False) for x in rings_only_smiles]
len(set(rings_only_smiles))
def karl_weight(m):
total = 0
for atom in m.GetAtoms():
total += atom.GetAtomicNum()
return total
weights = [karl_weight(m) for m in rings_only]
l = sorted(zip(weights, rings_only), key=lambda x: x[0])
l_1 = list(filter(lambda x: x[0] == 32, l))
l_1[0][1]
l_1[1][1]
l_1[10][1]
l_1[11][1]
substructure_smiles = rings_only_smiles
with open('data/frag_reduced.json', 'w') as fout:
fout.write(json.dumps(list(substructure_smiles)))
with open('data/frag_reduced.csv', 'w') as fout:
fout.write('smiles\n')
for line in substructure_smiles:
fout.write("%s\n" % line)
| dev_tree_decomp.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.7 64-bit (''hse'': conda)'
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import scipy.sparse as sparse
train = pd.read_parquet('data/train.par')
test = pd.read_parquet('data/test.par')
items = pd.read_parquet('data/items.par')
train
items
items.drop_duplicates(subset=['item_id'], inplace=True)
# ## Top-Popular Model
# +
def top_popular(interactions: pd.DataFrame, k=10):
item_popularity = interactions.groupby('item_id').size().reset_index(name='popularity')
top_popular = item_popularity.sort_values('popularity', ascending=False).head(k).item_id.values
prediction = interactions[['user_id']].drop_duplicates(ignore_index=True)
prediction['item_id'] = prediction.user_id.apply(lambda x: top_popular)
return prediction
toppop_prediction = top_popular(train)
# -
import my_metrics
my_metrics.compute(toppop_prediction, test)
# ## ALS
n_users = train.user_id.max() + 1
n_items = train.item_id.max() + 1
n_users, n_items
# +
train_ratings = train \
.groupby(['item_id', 'user_id'], as_index=False) \
.size() \
.rename(columns={'size': 'rating'})
user_sum_rating = train_ratings.groupby('user_id').rating.sum()
train_ratings = train_ratings.join(user_sum_rating, on='user_id', rsuffix='_sum')
train_ratings['rating_normal'] = train_ratings['rating'] / train_ratings['rating_sum']
# -
train_ratings
# +
confidence = 1.0 + train_ratings.rating_normal.values * 40.0
rating_matrix = sparse.csr_matrix(
(
confidence,
(
train_ratings.item_id.values,
train_ratings.user_id.values
)
),
shape=(n_items, n_users)
)
rating_matrix_T = sparse.csr_matrix(
(
np.full(rating_matrix.nnz, 1),
(
train_ratings.user_id.values,
train_ratings.item_id.values
)
),
shape=(n_users, n_items)
)
# -
rating_matrix.nnz / (n_items * n_users) * 100
# +
import implicit
als = implicit.als.AlternatingLeastSquares(factors=128,
calculate_training_loss=True,
iterations=100)
als.fit(rating_matrix)
# +
import joblib
def predict_als_for_user(user_id):
recommendations = als.recommend(user_id, rating_matrix_T, N=10)
recommended_items = [x for x, _ in recommendations]
recommended_scores = [x for _, x in recommendations]
return user_id, recommended_items, recommended_scores
als_prediction_raw = joblib.Parallel(backend='multiprocessing', verbose=1, n_jobs=32)(
joblib.delayed(predict_als_for_user)(u) for u in train.user_id.unique()
)
als_prediction = pd.DataFrame(als_prediction_raw, columns=['user_id', 'item_id', 'score'])
# -
my_metrics.compute(als_prediction, test)
| 1_als.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from BCAPI import BCAPI
bcapi = BCAPI()
# ## Options of the query
# #### Academic Units
bcapi.academic_unit_options()
# #### Categories
bcapi.category_options()
# #### Campuses
bcapi.campus_options()
# #### Semesters
bcapi.semester_options()
# ## Queries
# BCAPI has 2 methods that perform a requests the first one returns the response as the original html and the second as json
bcapi.search_html(semester="2019-2", name="diseño")
# The json method returns the response and a boolean that is True if the courses returned are NOT all the courses that meet the search params
res, incomplete = bcapi.search_json(semester="2019-2", name="diseño")
res
incomplete
import json
res = json.loads(res)
res[0]
| example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Lecture 03: Optimize, print and plot
# [Download on GitHub](https://github.com/NumEconCopenhagen/lectures-2022)
#
# [<img src="https://mybinder.org/badge_logo.svg">](https://mybinder.org/v2/gh/NumEconCopenhagen/lectures-2022/master?urlpath=lab/tree/03/Optimize_print_and_plot.ipynb)
# 1. [The consumer problem](#The-consumer-problem)
# 2. [Numerical python (numpy)](#Numerical-python-(numpy))
# 3. [Utility function](#Utility-function)
# 4. [Algorithm 1: Simple loops](#Algorithm-1:-Simple-loops)
# 5. [Algorithm 2: Use monotonicity](#Algorithm-2:-Use-monotonicity)
# 6. [Algorithm 3: Call a solver](#Algorithm-3:-Call-a-solver)
# 7. [Indifference curves](#Indifference-curves)
# 8. [A classy solution](#A-classy-solution)
# 9. [Summary](#Summary)
#
# You will learn how to work with numerical data (**numpy**) and solve simple numerical optimization problems (**scipy.optimize**) and report the results both in text (**print**) and in figures (**matplotlib**).
# **Links:**:
#
# - **print**: [examples](https://www.python-course.eu/python3_formatted_output.php) (very detailed)
# - **numpy**: [detailed tutorial](https://www.python-course.eu/numpy.php)
# - **matplotlib**: [examples](https://matplotlib.org/tutorials/introductory/sample_plots.html#sphx-glr-tutorials-introductory-sample-plots-py), [documentation](https://matplotlib.org/users/index.html), [styles](https://matplotlib.org/3.1.0/gallery/style_sheets/style_sheets_reference.html)
# - **scipy-optimize**: [documentation](https://docs.scipy.org/doc/scipy/reference/optimize.html)
# <a id="The-consumer-problem"></a>
#
# # 1. The consumer problem
# Consider the following 2-good consumer problem with
#
# * utility function $u(x_1,x_2):\mathbb{R}^2_{+}\rightarrow\mathbb{R}$,
# * exogenous income $I$, and
# * price-vector $(p_1,p_2)$,
# given by
#
# $$
# \begin{aligned}
# V(p_{1},p_{2},I) & = \max_{x_{1},x_{2}}u(x_{1},x_{2})\\
# \text{s.t.}\\
# p_{1}x_{1}+p_{2}x_{2} & \leq I,\,\,\,p_{1},p_{2},I>0\\
# x_{1},x_{2} & \geq 0
# \end{aligned}
# $$
# **Specific example:** Let the utility function be Cobb-Douglas,
#
# $$
# u(x_1,x_2) = x_1^{\alpha}x_2^{1-\alpha}
# $$
#
# We then know the solution is given by
#
# $$
# \begin{aligned}
# x_1^{\ast} &= \alpha \frac{I}{p_1} \\
# x_2^{\ast} &= (1-\alpha) \frac{I}{p_2}
# \end{aligned}
# $$
#
# which implies that $\alpha$ is the budget share of the first good and $1-\alpha$ is the budget share of the second good.
# <a id="Numerical-python-(numpy)"></a>
#
# # 2. Numerical python (numpy)
import numpy as np # import the numpy module
# A **numpy array** is like a list, but with two important differences:
#
# 1. Elements must be of **one homogenous type**
# 2. A **slice returns a view** rather than extract content
# ## 2.1 Basics
# Numpy arrays can be **created from lists** and can be **multi-dimensional**:
# +
A = np.array([0, 1, 2, 3, 4, 5, 6, 7, 8, 9]) # one dimension
B = np.array([[3.4, 8.7, 9.9],
[1.1, -7.8, -0.7],
[4.1, 12.3, 4.8]]) # two dimensions
print(type(A),type(B)) # type
print(A.dtype,B.dtype) # data type
print(A.ndim,B.ndim) # dimensions
print(A.shape,B.shape) # shape (1d: (columns,), 2d: (row,columns))
print(A.size,B.size) # size
# -
# **Slicing** a numpy array returns a **view**:
A = np.array([0, 1, 2, 3, 4, 5, 6, 7, 8, 9])
B = A.copy() # a copy of A
C = A[2:6] # a view into A
C[0] = 0
C[1] = 0
print(A) # changed
print(B) # not changed
# Numpy array can also be created using numpy functions:
print(np.ones((2,3)))
print(np.zeros((4,2)))
print(np.linspace(0,1,6)) # linear spacing
#np.linspace()
# **Tip 1:** Try pressing <kbd>Shift</kbd>+<kbd>Tab</kbd> inside a function.<br>
#
# **Tip 2:** Try to write `?np.linspace` in a cell
# ?np.linspace
# ## 2.2 Math
# Standard **mathematical operations** can be applied:
# +
A = np.array([[1,0],[0,1]])
B = np.array([[2,2],[2,2]])
print(A+B,'\n')
print(A-B,'\n')
print(A*B,'\n') # element-by-element product
print(A/B,'\n') # element-by-element division
print(A@B,'\n') # matrix product
# -
# If arrays does not fit together **broadcasting** is applied. Here is an example with multiplication:
# +
A = np.array([ [10, 20, 30], [40, 50, 60] ]) # shape = (2,3)
B = np.array([1, 2, 3]) # shape = (3,) = (1,3)
C = np.array([[1],[2]]) # shape = (2,1)
print(A, A.shape, '\n')
print(B, B.shape, '\n') # Notice the shape 'transformation' column vector!
print(C, C.shape, '\n')
print(A*B,'\n') # every row is multiplied by B
print(A*C,'\n') # every column is multiplied by C
# -
# If you want to e.g. add arrays where broadcasting is not possible consider **np.newaxis**:
# +
A = np.array([1, 2, 3]) # Is only 1D, shape = (3,)
B = np.array([1,2]) # Is only 1D, shape = (2,)
# You cannot broadcast B on A, because neither have 2 dimensions.
# Therefore, use newaxis
print(A[:,np.newaxis], A[:,np.newaxis].shape, '\n') # Is now (3,1)
print(B[np.newaxis,:], B[np.newaxis,:].shape, '\n') # Is now (1,2)
print(A[:,np.newaxis]*B[np.newaxis,:], '\n') # A is column vector, B is row vector
print(A[np.newaxis,:]*B[:,np.newaxis]) # A is row vector, B is column vector
# -
# **General rule:** Numpy arrays can be added/substracted/multiplied/divided if they in all dimensions have the same size or one of them has a size of one. If the numpy arrays differ in number of dimensions, this only has to be true for the (inner) dimensions they share.
# **More on broadcasting:** [Documentation](https://docs.scipy.org/doc/numpy-1.13.0/user/basics.broadcasting.html).
# A lot of **mathematical procedures** can easily be performed on numpy arrays.
A = np.array([3.1, 2.3, 9.1, -2.5, 12.1])
print(np.min(A)) # find minimum
print(np.argmin(A)) # find index for minimum
print(np.mean(A)) # calculate mean
print(np.sort(A)) # sort (ascending)
# **Note:** Sometimes a method can be used instead of a function, e.g. ``A.mean()``. Personally, I typically stick to functions because that always works.
# ## 2.3 Indexing
# **Multi-dimensional** indexing is done as:
X = np.array([ [11, 12, 13], [21, 22, 23] ])
print(X)
print(X[0,0]) # first row, first column
print(X[0,1]) # first row, second column
print(X[1,2]) # second row, third column
X[0] # first row
# Indexes can be **logical**. Logical 'and' is `&` and logical 'or' is `|`.
# +
A = np.array([1,2,3,4,1,2,3,4])
B = np.array([3,3,3,3,2,3,2,2])
I = (A < 3) & (B == 3) # note & instead of 'and'
print(I)
print(A[I],'\n')
# Two ways of getting indices of the elements == True
print(np.where(I)) # A 'where' clause normally asks for where the True elements are.
print(I.nonzero()) # Because a True boolean is a 1 while a False is a 0.
# -
I = (A < 3) | (B == 3) # note | instead of 'or'
print(A[I])
# ## 2.4 List of good things to know
# **Attributes and methods** to know:
#
# - size / ndim / shape
# - ravel / reshape / sort
# - copy
# **Functions** to know:
#
# - array / empty / zeros / ones / linspace
# - mean / median / std / var / sum / percentile
# - min/max, argmin/argmax / fmin / fmax / sort / clip
# - meshgrid / hstack / vstack / concatenate / tile / insert
# - allclose / isnan / isinf / isfinite / any / all
# **Concepts** to know:
#
# - view vs. copy
# - broadcasting
# - logical indexing
# **Quizz:** Follow this [link](https://forms.office.com/Pages/ResponsePage.aspx?id=kX-So6HNlkaviYyfHO_6kckJrnVYqJlJgGf8Jm3FvY9UQlM0OUs0UkJGV0tYMzhTVU81VkFMMUdCMCQlQCN0PWcu) to take a quizz on numpy arrays.
#
# **Additional question:** Consider the following code:
A = np.array([1,2,3,4,5])
B = A[3:]
B[:] = 0
# What is `np.sum(A)` equal to?
# - **A:** 15
# - **B:** 10
# - **C:** 6
# - **D:** 0
# - **E:** Don't know
# ## 2.5 Extra: Memory
# Memory is structured in **rows**:
A = np.array([[3.1,4.2],[5.7,9.3]])
B = A.ravel() # one-dimensional view of A
print(A.shape,A[0,:])
print(B.shape,B)
# <a id="Utility-function"></a>
#
# # 3. Utility function
# Define the utility function:
# +
def u_func(x1,x2,alpha=0.50):
return x1**alpha*x2**(1-alpha)
# x1,x2 are positional arguments
# alpha is a keyword argument with default value 0.50
# -
# ## 3.1 Print to screen
# Print a **single evaluation** of the utility function.
# +
x1 = 1
x2 = 3
u = u_func(x1,x2)
# f'text' is called a "formatted string"
# {x1:.3f} prints variable x1 as floating point number with 3 decimals
print(f'x1 = {x1:.3f}, x2 = {x2:.3f} -> u = {u:.3f}')
# -
# Print **multiple evaluations** of the utility function.
# +
x1_list = [2,4,6,8,10,12]
x2 = 3
for x1 in x1_list: # loop through each element in x1_list
u = u_func(x1,x2,alpha=0.25)
print(f'x1 = {x1:.3f}, x2 = {x2:.3f} -> u = {u:.3f}')
# -
# And a little nicer...
# +
for i,x1 in enumerate(x1_list): # i is a counter
u = u_func(x1,x2,alpha=0.25)
print(f'{i:2d}: x1 = {x1:<6.3f} x2 = {x2:<6.3f} -> u = {u:<6.3f}')
# {i:2d}: integer a width of 2 (right-aligned)
# {x1:<6.3f}: float width of 6 and 3 decimals (<, left-aligned)
# -
# See also [this source](https://www.geeksforgeeks.org/python-output-formatting/) for more info on output formatting.
# **Task**: Write a loop printing the results shown in the answer below.
# +
# write your code here
# -
# **Answer:**
for i,x1 in enumerate(x1_list): # i is a counter
u = u_func(x1,x2,alpha=0.25)
print(f'{i:2d}: u({x1:.2f},{x2:.2f}) = {u:.4f}')
# **More formatting options?** See these [examples](https://www.python-course.eu/python3_formatted_output.php).
# ## 3.2 Print to file
# Open a text-file as a handle and write lines in it:
# +
with open('somefile.txt', 'w') as the_file: # 'w' is for 'write'
for i, x1 in enumerate(x1_list):
u = u_func(x1,x2,alpha=0.25)
text = f'{i+10:2d}: x1 = {x1:<6.3f} x2 = {x2:<6.3f} -> u = {u:<6.3f}'
the_file.write(text + '\n') # \n gives a lineshift
# note: the with clause ensures that the file is properly closed afterwards
# -
# Open a text-file and read the lines in it and then print them:
with open('somefile.txt', 'r') as the_file: # 'r' is for 'read'
lines = the_file.readlines()
for line in lines:
print(line,end='') # end='' removes the extra lineshift print creates
# > **Note:** You could also write tables in LaTeX format and the import them in your LaTeX document.
# ## 3.3 Calculate the utility function on a grid
# **Calculate the utility function** on a 2-dimensional grid with $N$ elements in each dimension:
# +
# a. settings
N = 100 # number of elements
x_max = 10 # maximum value
# b. allocate numpy arrays
shape_tuple = (N,N)
x1_values = np.empty(shape_tuple) # allocate 2d numpy array with shape=(N,N)
x2_values = np.empty(shape_tuple)
u_values = np.empty(shape_tuple)
# Note: x1_values and x2_values are 2d. This is not strictly necessary in the present case. 1d arrays would suffice below in the nested loop and filling out of u_values.
# However, it makes them isomorphic with the mesh grids used for countour plots, which often need 2d mesh grids.
# c. fill numpy arrays
for i in range(N): # 0,1,...,N-1
for j in range(N): # 0,1,...,N-1
x1_values[i,j] = (i/(N-1))*x_max # in [0,x_max]
x2_values[i,j] = (j/(N-1))*x_max # in [0,x_max]
u_values[i,j] = u_func(x1_values[i,j],x2_values[i,j],alpha=0.25)
# -
# **Alternatively:** Use internal numpy functions:
x_vec = np.linspace(0,x_max,N)
x1_values_alt,x2_values_alt = np.meshgrid(x_vec,x_vec,indexing='ij')
print('Dimension of grid over x1 (and x2): ', x1_values_alt.shape) # Note that the grid is 2d. u_func needs 2d to calculate element by element.
u_values_alt = u_func(x1_values_alt, x2_values_alt, alpha=0.25)
# **Mesh grids** are a little tricky to understand, but important for creating surface plots. You can read more at:
# [GeeksforGeeks](https://www.geeksforgeeks.org/numpy-meshgrid-function/), [stack overflow](https://stackoverflow.com/questions/36013063/what-is-the-purpose-of-meshgrid-in-python-numpy) and the [numpy doc](https://numpy.org/doc/stable/reference/generated/numpy.meshgrid.html)
# Test whether the results are the same:
# +
# a. maximum absolute difference
max_abs_diff = np.max(np.abs(u_values-u_values_alt))
print(max_abs_diff) # very close to zero
# b. test if all values are "close"
print(np.allclose(u_values,u_values_alt))
# -
# **Note:** The results are not exactly the same due to floating point arithmetics.
# ## 3.4 Plot the utility function
# Import modules and state that the figures should be inlined:
# %matplotlib inline
import matplotlib.pyplot as plt # baseline modul
from mpl_toolkits.mplot3d import Axes3D # for 3d figures
plt.style.use('seaborn-whitegrid') # whitegrid nice with 3d
# Construct the actual plot:
# +
fig = plt.figure() # create the figure
ax = fig.add_subplot(1,1,1,projection='3d') # create a 3d axis in the figure
ax.plot_surface(x1_values,x2_values,u_values); # create surface plot in the axis
# note: fig.add_subplot(a,b,c) creates the c'th subplot in a grid of a times b plots
# -
# Make the figure **zoomable** and **panable** using a widget:
# %matplotlib widget
fig = plt.figure() # create the figure
ax = fig.add_subplot(1,1,1,projection='3d') # create a 3d axis in the figure
ax.plot_surface(x1_values,x2_values,u_values); # create surface plot in the axis
# Turn back to normal inlining:
# %matplotlib inline
# **Extensions**: Use a colormap, make it pretier, and save to disk.
# +
from matplotlib import cm # for colormaps
# a. actual plot
fig = plt.figure()
ax = fig.add_subplot(1,1,1,projection='3d')
ax.plot_surface(x1_values,x2_values,u_values,cmap=cm.jet)
# b. add labels
ax.set_xlabel('$x_1$')
ax.set_ylabel('$x_2$')
ax.set_zlabel('$u$')
# c. invert xaxis to bring Origin in center front
ax.invert_xaxis()
# d. save
fig.tight_layout()
fig.savefig('someplot.pdf') # or e.g. .png
# -
# **More formatting options?** See these [examples](https://matplotlib.org/tutorials/introductory/sample_plots.html#sphx-glr-tutorials-introductory-sample-plots-py).
# **Quizz:** follow [this link](https://forms.office.com/Pages/ResponsePage.aspx?id=kX-So6HNlkaviYyfHO_6kckJrnVYqJlJgGf8Jm3FvY9UREZRUU1BNDdJOEZDSTNMTzNTSVg1UlZNRSQlQCN0PWcu) to take a quizz on 3d plotting.
# **Task**: Construct the following plot:
# 
# **Answer:**
# +
# write your code here
# +
# a. actual plot
fig = plt.figure()
ax = fig.add_subplot(1,1,1,projection='3d')
ax.plot_wireframe(x1_values,x2_values,u_values,edgecolor='black')
# b. add labels
ax.set_xlabel('$x_1$')
ax.set_ylabel('$x_2$')
ax.set_zlabel('$u$')
# c. invert xaxis
ax.invert_xaxis()
# e. save
fig.tight_layout()
fig.savefig('someplot_wireframe.png')
fig.savefig('someplot_wireframe.pdf')
# -
# ## 3.5 Summary
# We have talked about:
#
# 1. Print (to screen and file)
# 2. Figures (matplotlib)
# **Other plotting libraries:** [seaborn](https://seaborn.pydata.org/) and [bokeh](https://bokeh.pydata.org/en/latest/).
# <a id="Algorithm-1:-Simple-loops"></a>
#
# # 4. Algorithm 1: Simple loops
# Remember the problem we wanted to solve:
# $$
# \begin{aligned}
# V(p_{1},p_{2},I) & = \max_{x_{1},x_{2}}u(x_{1},x_{2})\\
# & \text{s.t.}\\
# p_{1}x_{1}+p_{2}x_{2} & \leq I,\,\,\,p_{1},p_{2},I>0\\
# x_{1},x_{2} & \geq 0
# \end{aligned}
# $$
# **Idea:** Loop through a grid of $N_1 \times N_2$ possible solutions. This is the same as solving:
#
# $$
# \begin{aligned}
# V(p_{1},p_{2},I) & = \max_{x_{1}\in X_1,x_{2} \in X_2} x_1^{\alpha}x_2^{1-\alpha}\\
# & \text{s.t.}\\
# X_1 & = \left\{0,\frac{1}{N_1-1}\frac{I}{p_1},\frac{2}{N_1-1}\frac{I}{p_1},\dots,\frac{I}{p_1}\right\} \\
# X_2 & = \left\{0,\frac{1}{N_2-1}\frac{I}{p_2},\frac{2}{N_2-1}\frac{ I}{p_2},\dots,\frac{ I}{p_2}\right\} \\
# p_{1}x_{1}+p_{2}x_{2} & \leq I\\
# \end{aligned}
# $$
# Function doing just this:
# +
def find_best_choice(alpha,I,p1,p2,N1,N2,do_print=True):
# a. allocate numpy arrays
shape_tuple = (N1,N2)
x1_values = np.empty(shape_tuple)
x2_values = np.empty(shape_tuple)
u_values = np.empty(shape_tuple)
# b. start from guess of x1=x2=0
x1_best = 0
x2_best = 0
u_best = u_func(0,0,alpha=alpha)
# c. loop through all possibilities
for i in range(N1):
for j in range(N2):
# i. x1 and x2 (chained assignment)
x1_values[i,j] = x1 = (i/(N1-1))*I/p1
x2_values[i,j] = x2 = (j/(N2-1))*I/p2
# ii. utility
if p1*x1 + p2*x2 <= I: # u(x1,x2) if expenditures <= income
u_values[i,j] = u_func(x1,x2,alpha=alpha)
else: # u(0,0) if expenditures > income
u_values[i,j] = u_func(0,0,alpha=alpha)
# iii. check if best sofar
if u_values[i,j] > u_best:
x1_best = x1_values[i,j]
x2_best = x2_values[i,j]
u_best = u_values[i,j]
# d. print
if do_print:
print_solution(x1_best,x2_best,u_best,I,p1,p2)
return x1_best,x2_best,u_best,x1_values,x2_values,u_values
# function for printing the solution
def print_solution(x1,x2,u,I,p1,p2):
print(f'x1 = {x1:.8f}')
print(f'x2 = {x2:.8f}')
print(f'u = {u:.8f}')
print(f'I-p1*x1-p2*x2 = {I-p1*x1-p2*x2:.8f}')
# -
# Call the function:
sol = find_best_choice(alpha=0.25,I=10,p1=1,p2=2,N1=500,N2=400)
# Plot the solution:
# +
# %matplotlib widget
# a. unpack solution
x1_best,x2_best,u_best,x1_values,x2_values,u_values = sol
# b. setup figure
fig = plt.figure(dpi=100,num='')
ax = fig.add_subplot(1,1,1,projection='3d')
# c. plot 3d surface of utility values for different choices
ax.plot_surface(x1_values,x2_values,u_values,cmap=cm.jet)
ax.invert_xaxis()
# d. plot optimal choice
ax.scatter(x1_best,x2_best,u_best,s=50,color='black');
# -
# **Quizz:** take a quick [quizz](https://forms.office.com/Pages/ResponsePage.aspx?id=kX-So6HNlkaviYyfHO_6kckJrnVYqJlJgGf8Jm3FvY9UOFVXRE5YSEQwVjdETDY4MDVCODlTMk5UQiQlQCN0PWcu) on plotting the simple loop.
# %matplotlib inline
# **Task**: Can you find a better solution with higher utility and lower left-over income, $I-p_1 x_1-p_2 x_2$?
# +
# write your code here
# sol = find_best_choice()
# -
# **Answer:**
sol = find_best_choice(alpha=0.25,I=10,p1=1,p2=2,N1=1000,N2=1000)
# <a id="Algorithm-2:-Use-monotonicity"></a>
#
# # 5. Algorithm 2: Use monotonicity
# **Idea:** Loop through a grid of $N$ possible solutions for $x_1$ and assume the remainder is spent on $x_2$. This is the same as solving:
#
# $$
# \begin{aligned}
# V(p_{1},p_{2},I) & = \max_{x_{1}\in X_1} x_1^{\alpha}x_2^{1-\alpha}\\
# \text{s.t.}\\
# X_1 & = \left\{0,\frac{1}{N-1}\frac{}{p_1},\frac{2}{N-1}\frac{I}{p_1},\dots,\frac{I}{p_1}\right\} \\
# x_{2} & = \frac{I-p_{1}x_{1}}{p_2}\\
# \end{aligned}
# $$
# Function doing just this:
def find_best_choice_monotone(alpha,I,p1,p2,N,do_print=True):
# a. allocate numpy arrays
shape_tuple = (N)
x1_values = np.empty(shape_tuple)
x2_values = np.empty(shape_tuple)
u_values = np.empty(shape_tuple)
# b. start from guess of x1=x2=0
x1_best = 0
x2_best = 0
u_best = u_func(0,0,alpha)
# c. loop through all possibilities
for i in range(N):
# i. x1
x1_values[i] = x1 = i/(N-1)*I/p1
# ii. implied x2
x2_values[i] = x2 = (I-p1*x1)/p2
# iii. utility
u_values[i] = u_func(x1,x2,alpha)
if u_values[i] >= u_best:
x1_best = x1_values[i]
x2_best = x2_values[i]
u_best = u_values[i]
# d. print
if do_print:
print_solution(x1_best,x2_best,u_best,I,p1,p2)
return x1_best,x2_best,u_best,x1_values,x2_values,u_values
sol_monotone = find_best_choice_monotone(alpha=0.25,I=10,p1=1,p2=2,N=1000)
# Plot the solution:
# +
plt.style.use("seaborn-whitegrid")
# a. create the figure
fig = plt.figure(figsize=(10,4))# figsize is in inches...
# b. unpack solution
x1_best,x2_best,u_best,x1_values,x2_values,u_values = sol_monotone
# c. left plot
ax_left = fig.add_subplot(1,2,1)
ax_left.plot(x1_values,u_values)
ax_left.scatter(x1_best,u_best) # Add the solution as a dot
ax_left.set_title('value of choice, $u(x_1,x_2)$')
ax_left.set_xlabel('$x_1$')
ax_left.set_ylabel('$u(x_1,(I-p_1 x_1)/p_2)$')
ax_left.grid(True)
# c. right plot
ax_right = fig.add_subplot(1,2,2)
ax_right.plot(x1_values,x2_values)
ax_right.scatter(x1_best,x2_best)
ax_right.set_title('implied $x_2$')
ax_right.set_xlabel('$x_1$')
ax_right.set_ylabel('$x_2$')
ax_right.grid(True)
# -
# <a id="Algorithm-3:-Call-a-solver"></a>
#
# # 6. Algorithm 3: Call a solver
#
from scipy import optimize
# Choose paramters:
alpha = 0.25 # preference parameter
I = 10 # income
p1 = 1 # price 1
p2 = 2 # price 2
# **Case 1**: Scalar solver using monotonicity.
# +
# a. objective funciton (to minimize)
def value_of_choice(x1,alpha,I,p1,p2):
x2 = (I-p1*x1)/p2
return -u_func(x1,x2,alpha)
# b. call solver
sol_case1 = optimize.minimize_scalar(
value_of_choice,method='bounded',
bounds=(0,I/p1),args=(alpha,I,p1,p2))
# c. unpack solution
x1 = sol_case1.x
x2 = (I-p1*x1)/p2
u = u_func(x1,x2,alpha)
print_solution(x1,x2,u,I,p1,p2)
# -
# **Case 2**: Multi-dimensional constrained solver.
# +
# a. objective function (to minimize)
def value_of_choice(x,alpha,I,p1,p2):
# note: x is a vector
x1 = x[0]
x2 = x[1]
return -u_func(x1,x2,alpha)
# b. constraints (violated if negative) and bounds
constraints = ({'type': 'ineq', 'fun': lambda x: I-p1*x[0]-p2*x[1]})
bounds = ((0,I/p1),(0,I/p2))
# c. call solver
initial_guess = [I/p1/2,I/p2/2]
sol_case2 = optimize.minimize(
value_of_choice,initial_guess,args=(alpha,I,p1,p2),
method='SLSQP',bounds=bounds,constraints=constraints)
# d. unpack solution
x1 = sol_case2.x[0]
x2 = sol_case2.x[1]
u = u_func(x1,x2,alpha)
print_solution(x1,x2,u,I,p1,p2)
# -
# **Case 3**: Multi-dimensional unconstrained solver with constrains implemented via penalties.
# +
# a. objective function (to minimize)
def value_of_choice(x,alpha,I,p1,p2):
# i. unpack
x1 = x[0]
x2 = x[1]
# ii. penalty
penalty = 0
E = p1*x1+p2*x2 # total expenses
if E > I: # expenses > income -> not allowed
fac = I/E
penalty += 1000*(E-I) # calculate penalty
x1 *= fac # force E = I
x2 *= fac # force E = I
return -u_func(x1,x2,alpha)
# b. call solver
initial_guess = [I/p1/2,I/p2/2]
sol_case3 = optimize.minimize(
value_of_choice,initial_guess,method='Nelder-Mead',
args=(alpha,I,p1,p2))
# c. unpack solution
x1 = sol_case3.x[0]
x2 = sol_case3.x[1]
u = u_func(x1,x2,alpha)
print_solution(x1,x2,u,I,p1,p2)
# -
# **Task:** Find the <font color='red'>**error**</font> in the code in the previous cell.
# +
# write your code here
# -
# **Answer:**
# +
# a. objective function (to minimize)
def value_of_choice(x,alpha,I,p1,p2):
# i. unpack
x1 = x[0]
x2 = x[1]
# ii. penalty
penalty = 0
E = p1*x1+p2*x2 # total expenses
if E > I: # expenses > income -> not allowed
fac = I/E
penalty += 1000*(E-I) # calculate penalty
x1 *= fac # force E = I
x2 *= fac # force E = I
return -u_func(x1,x2,alpha) + penalty # the error
# b. call solver
initial_guess = [I/p1/2,I/p2/2]
sol_case3 = optimize.minimize(
value_of_choice,initial_guess,method='Nelder-Mead',
args=(alpha,I,p1,p2))
# c. unpack solution
x1 = sol_case3.x[0]
x2 = sol_case3.x[1]
u = u_func(x1,x2,alpha)
print_solution(x1,x2,u,I,p1,p2)
# -
# <a id="Indifference-curves"></a>
#
# # 7. Indifference curves
# Remember that the indifference curve through the point $(y_1,y_2)$ is given by
#
# $$
# \big\{(x_1,x_2) \in \mathbb{R}^2_+ \,|\, u(x_1,x_2) = u(y_1,y_2)\big\}
# $$
#
# To find the indifference curve, we can fix a grid for $x_2$, and then find the corresponding $x_1$ which solves $u(x_1,x_2) = u(y_1,y_2)$ for each value of $x_2$.
# +
def objective(x1,x2,alpha,u):
return u_func(x1,x2,alpha)-u
# = 0 then on indifference curve with utility = u
def find_indifference_curve(y1,y2,alpha,N,x2_max):
# a. utility in (y1,y2)
u_y1y2 = u_func(y1,y2,alpha)
# b. allocate numpy arrays
x1_vec = np.empty(N)
x2_vec = np.linspace(1e-8,x2_max,N)
# c. loop through x2
for i,x2 in enumerate(x2_vec):
x1_guess = 0 # initial guess
sol = optimize.root(objective, x1_guess, args=(x2,alpha,u_y1y2))
# optimize.root -> solve objective = 0 starting from x1 = x1_guess
x1_vec[i] = sol.x[0]
return x1_vec,x2_vec
# -
# Find and plot an inddifference curve:
# +
# a. find indifference curve through (4,4) for x2 in [0,10]
x2_max = 10
x1_vec,x2_vec = find_indifference_curve(y1=4,y2=4,alpha=0.25,N=100,x2_max=x2_max)
# b. plot inddifference curve
fig = plt.figure(figsize=(6,6))
ax = fig.add_subplot(1,1,1)
ax.plot(x1_vec,x2_vec)
ax.set_xlabel('$x_1$')
ax.set_ylabel('$x_2$')
ax.set_xlim([0,x2_max])
ax.set_ylim([0,x2_max])
ax.grid(True)
# -
# **Task:** Find the indifference curve through $x_1 = 15$ and $x_2 = 3$ with $\alpha = 0.5$.
# + code_folding=[] hidden=true
# write your code here
# +
x2_max = 20
x1_vec,x2_vec = find_indifference_curve(y1=15,y2=3,alpha=0.5,N=100,x2_max=x2_max)
fig = plt.figure(figsize=(6,6))
ax = fig.add_subplot(1,1,1)
ax.plot(x1_vec,x2_vec)
ax.set_xlabel('$x_1$')
ax.set_ylabel('$x_2$')
ax.set_xlim([0,x2_max])
ax.set_ylim([0,x2_max])
ax.grid(True)
# -
# <a id="A-classy-solution"></a>
#
# # 8. A classy solution
# > **Note:** This section is advanced due to the use of a module with a class. It is, however, a good example of how to structure code for solving and illustrating a model.
# **Load module** I have written (consumer_module.py in the same folder as this notebook).
from consumer_module import consumer
# ## 8.1 Jeppe
# Give birth to a consumer called **jeppe**:
jeppe = consumer() # create an instance of the consumer class called jeppe
print(jeppe)
# Solve **jeppe**'s problem.
jeppe.solve()
print(jeppe)
# ## 8.2 Mette
# Create a new consumer, called Mette, and solve her problem.
mette = consumer(alpha=0.25)
mette.solve()
mette.find_indifference_curves()
print(mette)
# Make an illustration of Mette's problem and it's solution:
# +
fig = plt.figure(figsize=(6,6))
ax = fig.add_subplot(1,1,1)
mette.plot_indifference_curves(ax)
mette.plot_budgetset(ax)
mette.plot_solution(ax)
mette.plot_details(ax)
# -
# **Advanced note:** Looking at the code behind the consumer class, you'll notice the 'self' argument a lot. This argument links the functions in the class definition, which holds all the instructions for behavior, and the specific object. And exactly because 'self' is the first argument in class functions, one can actually call the general class and provide it with the object one wants to evaluate a function on.
# +
# Example:
christian = consumer()
# Calling the consumer class function and providing the object christian :
consumer.solve(christian)
print('call to consumer class: \n',christian)
# is the same as the call to the christian object directly
christian.solve()
print('call to the object christian directly: \n',christian)
# -
# <a id="Summary"></a>
#
# # 9. Summary
# **This lecture:** We have talked about:
#
# 1. Numpy (view vs. copy, indexing, broadcasting, functions, methods)
# 2. Print (to screen and file)
# 3. Figures (matplotlib)
# 4. Optimization (using loops or scipy.optimize)
# 5. Advanced: Consumer class
# Most economic models contain optimizing agents solving a constrained optimization problem. The tools applied in this lecture is not specific to the consumer problem in anyway.
# **Your work:** Before solving Problem Set 1 read through this notebook and play around with the code. To solve the problem set, you only need to modify the code used here slightly.
# **Next lecture:** Random numbers and simulation.
| web/03/Optimize_print_and_plot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="MjBeTMLJQJuQ"
# # Object Detection with Region Based Convolutional Neural Network.
# + [markdown] colab_type="text" id="-uYOhP5GR2L0"
# You should have seen in our previous posts that Convolutional Neural Network is the state of the art for any computer vision task like
#
# - [Image classification](https://www.learnopencv.com/pytorch-for-beginners-image-classification-using-pre-trained-models/)
#
#
# - [Semantic Segmentation](https://www.learnopencv.com/pytorch-for-beginners-semantic-segmentation-using-torchvision/)
#
# In this notebook we will look at another computer vision application called object detection
# + [markdown] colab_type="text" id="pNatz6V4Z3JZ"
# # Object Localization
#
# In addition to classify an image, we want to know where in the image the object is. So we want to classify the image as cat/dog/..etc and we want to draw a bounding box around the region of the object in the image.
#
#
# But Object localization is pretty simple, it classifies an image like image classiifcation and draw a bounding box around the classified image. So if more that 1 class is present in the image, it ignores the other class and classifies image as a single class and draw box around it.
#
# Now given an image, the model will return the probability of the content of the image belonging to different known classes and the model will also return 4 floating values of the coordinates of the bounding box.
#
#
# + [markdown] colab_type="text" id="5SOkRRaEl40c"
# # Object Detection
#
# Object Detection is the next step of Object Localization where the task remains the same(classification and bounding box) but obejct detection input images may have multiple known class objects in a single image and the task is to draw a bounding box around all the objects and classify each object.
#
# The main challenge here is that there might be a varying number of objects in every input image.
#
#
# - ## Sliding Window Approach
# Sliding window is one of the oldest approach in object detection where the input image is split into multiple crops and each crop of the image is classified and if the crop contains a class, then the crop is decided as the bounding box. But this approach is never used in practice as each input image may have 1000s of such crops and each crop passing through the network for classification may take time.
#
# - ## Region Proposal (RCNN)
# Image processing techniques are used to make list of proposed regions in the input image which are then sent through the network for classification. But this is computationally more efficient than sliding window approach as only fewer potential crops which may contain the object is classified by the network.
#
# 
# Image Source : [<NAME> et al](https://arxiv.org/pdf/1311.2524.pdf)
#
# RCNN is better than sliding window, but its still computationally expensive as the network has to classify all the region proposals. It takes around 30-40s for inference of a single image.
#
#
# - ## Fast Region Proposal (Fast RCNN)
# In fast RCNN, rather than getting region proposals and classifying each region proposals, the input image is sent into the CNN network which gives a feature map of the image. Again some region proposals are used but now we get the region proposals from the feature map of the image and these feature maps are classified. This reduces the computation as some of the CNN layers are common for the whole image.
#
# 
# Image Source : [<NAME>](https://arxiv.org/pdf/1504.08083.pdf)
#
#
# - ## Faster R-CNN
# The idea of Faster R-CNN is to use CNNs to propose potential region of interest and the network is called Region Proposal Network. After getting the region proposals , its just like Fast RCNN, we use every regions for classification.
# 
# + [markdown] colab_type="text" id="3joSB5Ta7t5j"
# # Object Detection, Instance Segmentation and Person Keypoint Detection in PyTorch
#
# It is fine if you don't understand every detail of the models discussed above. If you want to learn more about all of these models and many more application and concepts of Deep Learning and Computer Vision indetail, check out the official [Deep Learning and Computer Vision courses](https://opencv.org/courses/) by OpenCV.org.
#
# Now we will use pre trained models in PyTorch for
# - Object Detection
# - Instance Segmentation
# - Person Keypoint Detection
#
#
# All the pretrained models in pytorch can be found in [torchvision.models](https://pytorch.org/docs/stable/torchvision/models.html)
#
# + [markdown] colab_type="text" id="dNWsNDZQ8HF3"
# # Object Detection with PyTorch
#
# The pretrained Faster-RCNN ResNet-50 model we are going to use expects the input image tensor to be in the form ```[n, c, h, w]```
# where
# - n is the number of images
# - c is the number of channels , for RGB images its 3
# - h is the height of the image
# - w is the widht of the image
#
# The model will return
# - Bounding boxes [x0, y0, x1, y1] all all predicted classes of shape (N,4) where N is the number of classes predicted by the model to be present in the image.
# - Labels of all predicted classes.
# - Scores of each predicted label.
#
#
# + colab={} colab_type="code" id="qWOd_NvOYY-P"
# import necessary libraries
from PIL import Image
import matplotlib.pyplot as plt
import torch
import torchvision.transforms as T
import torchvision
import torch
import numpy as np
import cv2
# get the pretrained model from torchvision.models
# Note: pretrained=True will get the pretrained weights for the model.
# model.eval() to use the model for inference
model = torchvision.models.detection.fasterrcnn_resnet50_fpn(pretrained=True)
model.eval()
# Class labels from official PyTorch documentation for the pretrained model
# Note that there are some N/A's
# for complete list check https://tech.amikelive.com/node-718/what-object-categories-labels-are-in-coco-dataset/
# we will use the same list for this notebook
COCO_INSTANCE_CATEGORY_NAMES = [
'__background__', 'person', 'bicycle', 'car', 'motorcycle', 'airplane', 'bus',
'train', 'truck', 'boat', 'traffic light', 'fire hydrant', 'N/A', 'stop sign',
'parking meter', 'bench', 'bird', 'cat', 'dog', 'horse', 'sheep', 'cow',
'elephant', 'bear', 'zebra', 'giraffe', 'N/A', 'backpack', 'umbrella', 'N/A', 'N/A',
'handbag', 'tie', 'suitcase', 'frisbee', 'skis', 'snowboard', 'sports ball',
'kite', 'baseball bat', 'baseball glove', 'skateboard', 'surfboard', 'tennis racket',
'bottle', 'N/A', 'wine glass', 'cup', 'fork', 'knife', 'spoon', 'bowl',
'banana', 'apple', 'sandwich', 'orange', 'broccoli', 'carrot', 'hot dog', 'pizza',
'donut', 'cake', 'chair', 'couch', 'potted plant', 'bed', 'N/A', 'dining table',
'N/A', 'N/A', 'toilet', 'N/A', 'tv', 'laptop', 'mouse', 'remote', 'keyboard', 'cell phone',
'microwave', 'oven', 'toaster', 'sink', 'refrigerator', 'N/A', 'book',
'clock', 'vase', 'scissors', 'teddy bear', 'hair drier', 'toothbrush'
]
def get_prediction(img_path, threshold):
"""
get_prediction
parameters:
- img_path - path of the input image
- threshold - threshold value for prediction score
method:
- Image is obtained from the image path
- the image is converted to image tensor using PyTorch's Transforms
- image is passed through the model to get the predictions
- class, box coordinates are obtained, but only prediction score > threshold
are chosen.
"""
img = Image.open(img_path)
transform = T.Compose([T.ToTensor()])
img = transform(img)
pred = model([img])
pred_class = [COCO_INSTANCE_CATEGORY_NAMES[i] for i in list(pred[0]['labels'].numpy())]
pred_boxes = [[(i[0], i[1]), (i[2], i[3])] for i in list(pred[0]['boxes'].detach().numpy())]
pred_score = list(pred[0]['scores'].detach().numpy())
pred_t = [pred_score.index(x) for x in pred_score if x>threshold][-1]
pred_boxes = pred_boxes[:pred_t+1]
pred_class = pred_class[:pred_t+1]
return pred_boxes, pred_class
def object_detection_api(img_path, threshold=0.5, rect_th=3, text_size=3, text_th=3):
"""
object_detection_api
parameters:
- img_path - path of the input image
- threshold - threshold value for prediction score
- rect_th - thickness of bounding box
- text_size - size of the class label text
- text_th - thichness of the text
method:
- prediction is obtained from get_prediction method
- for each prediction, bounding box is drawn and text is written
with opencv
- the final image is displayed
"""
boxes, pred_cls = get_prediction(img_path, threshold)
img = cv2.imread(img_path)
img = cv2.cvtColor(img, cv2.COLOR_BGR2RGB)
for i in range(len(boxes)):
cv2.rectangle(img, boxes[i][0], boxes[i][1],color=(0, 255, 0), thickness=rect_th)
cv2.putText(img,pred_cls[i], boxes[i][0], cv2.FONT_HERSHEY_SIMPLEX, text_size, (0,255,0),thickness=text_th)
plt.figure(figsize=(20,30))
plt.imshow(img)
plt.xticks([])
plt.yticks([])
plt.show()
# + [markdown] colab_type="text" id="v2Mo500ZJi1i"
# Let's try one more complex example
# ```
# # # !wget https://cdn.pixabay.com/photo/2013/07/05/01/08/traffic-143391_960_720.jpg -O traffic.jpg
#
# object_detection_api('/content/traffic.jpg', rect_th=2, text_th=1, text_size=1)
# ```
#
# + colab={"base_uri": "https://localhost:8080/", "height": 969} colab_type="code" id="R8Q4sr3sYZD3" outputId="c370d6d4-e8e5-47b4-f406-7aa1f0b72fdf"
# download an image for inference
# # !wget https://www.wsha.org/wp-content/uploads/banner-diverse-group-of-people-2.jpg -O people.jpg
# use the api pipeline for object detection
# the threshold is set manually, the model sometimes predict
# random structures as some object, so we set a threshold to filter
# better prediction scores.
object_detection_api('./people.jpg', threshold=0.8)
# + colab={"base_uri": "https://localhost:8080/", "height": 907} colab_type="code" id="Qwlq1NJKS4dS" outputId="9d4fd60e-4626-4591-fbdc-80c439793fb8"
# # !wget https://hips.hearstapps.com/hmg-prod.s3.amazonaws.com/images/10best-cars-group-cropped-1542126037.jpg -O cars.jpg
object_detection_api('./cars.jpg', rect_th=6, text_th=5, text_size=5)
# + colab={"base_uri": "https://localhost:8080/", "height": 969} colab_type="code" id="2W86i2oxWyH2" outputId="7037d4ab-424f-4fd2-f5c0-9c82fca415b7"
# # !wget https://cdn.pixabay.com/photo/2013/07/05/01/08/traffic-143391_960_720.jpg -O traffic_scene.jpg
object_detection_api('./traffic_scene.jpg', rect_th=2, text_th=1, text_size=1)
# + colab={"base_uri": "https://localhost:8080/", "height": 969} colab_type="code" id="tApZDCJGVcxQ" outputId="84a60471-174b-4814-e632-e9fc0d12dcd8"
# # !wget https://images.unsplash.com/photo-1458169495136-854e4c39548a -O traffic_scene2.jpg
object_detection_api('./traffic_scene2.jpg', rect_th=15, text_th=7, text_size=5, threshold=0.8)
# + [markdown] colab_type="text" id="8st2TuBIQkSJ"
# # Comparing the inference time of model in CPU & GPU
#
#
# + colab={} colab_type="code" id="9OYNHMIih4up"
import time
def check_inference_time(image_path, gpu=False):
model = torchvision.models.detection.fasterrcnn_resnet50_fpn(pretrained=True)
model.eval()
img = Image.open(image_path)
transform = T.Compose([T.ToTensor()])
img = transform(img)
if gpu:
model.cuda()
img = img.cuda()
else:
model.cpu()
img = img.cpu()
start_time = time.time()
pred = model([img])
end_time = time.time()
return end_time-start_time
# + [markdown] colab_type="text" id="wxkL9LRPXa5X"
# ## Inference time for Object Detection
# + colab={"base_uri": "https://localhost:8080/", "height": 85} colab_type="code" id="ptbJ57cUSfut" outputId="7849439d-f7a8-41f6-e2cc-ed3436261e86"
cpu_time = sum([check_inference_time('./girl_cars.jpg', gpu=False) for _ in range(10)])/10.0
# gpu_time = sum([check_inference_time('./girl_cars.jpg', gpu=True) for _ in range(10)])/10.0
print('\nAverage Time take by the model with CPU = {}s'.format(cpu_time))
# print('\n\nAverage Time take by the model with GPU = {}s\nAverage Time take by the model with CPU = {}s'.format(gpu_time, cpu_time))
| faster-RCNN/.ipynb_checkpoints/faster_RCNN-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="_ZYxOI1mGNV1" colab_type="text"
# #Step 01. Install All Dependencies
#
# This installs Apache Spark 2.3.3, Java 8, Findspark library that makes it easy for Python to work on the given Big Data.
# + id="MG5jn29qF91X" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="8f203828-514e-4d55-95af-4ba46648c82d"
import os
#OpenJDK Dependencies for Spark
os.system('apt-get install openjdk-8-jdk-headless -qq > /dev/null')
#Download Apache Spark
os.system('wget -q http://apache.osuosl.org/spark/spark-2.3.3/spark-2.3.3-bin-hadoop2.7.tgz')
#Apache Spark and Hadoop Unzip
os.system('tar xf spark-2.3.3-bin-hadoop2.7.tgz')
#FindSpark Install
os.system('pip install -q findspark')
# + [markdown] id="uE_C9VOSHOKy" colab_type="text"
# # Step 02. Set Environment Variables
# Set the locations where Spark and Java are installed based on your installation configuration. Double check before you proceed.
# + id="qrOQoyMmHRPL" colab_type="code" colab={}
import os
os.environ["JAVA_HOME"] = "/usr/lib/jvm/java-8-openjdk-amd64"
os.environ["SPARK_HOME"] = "/content/spark-2.3.3-bin-hadoop2.7"
# + [markdown] id="qY1GD4JslzUH" colab_type="text"
# # Step 03. ELT - Load the Data: Mega Cloud Access
# This is an alternative approach to load datasets from already stored in [**Mega Cloud**](https://mega.nz) cloud repository. You need to install the necessary packages and put the link URL of cloud to load the file from cloud directly.
# + id="LCqsmO2fl_9Y" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="72ff9155-a67d-42fb-ada6-e2ddc1d943af"
import os
os.system('git clone https://github.com/jeroenmeulenaar/python3-mega.git')
os.chdir('python3-mega')
os.system('pip install -r requirements.txt')
# + [markdown] id="BAYD5dmomHgL" colab_type="text"
# # Step 04. ELT - Load the Data: Read Uploaded Dataset
# In this approach you can directly load the uploaded dataset downloaded fro Mega Cloud Infrastructure
# + id="CCWFVfBsmM03" colab_type="code" colab={}
from mega import Mega
os.chdir('../')
m_revenue = Mega.from_ephemeral()
m_revenue.download_from_url('https://mega.nz/#!1lJH3Q4K!N94-KRSyn22FPb0yxiVJgndjxUStdlfC2_prWDYI2f0')
# + [markdown] id="9iRVypEMHiDe" colab_type="text"
# # Step 05. Start a Spark Session
# This basic code will prepare to start a Spark session.
# + id="vrW7H-rmHmFm" colab_type="code" colab={}
import findspark
findspark.init()
from pyspark.sql import SparkSession
spark = SparkSession.builder.appName('ml-datathon19-easy01').master("local[*]").getOrCreate()
# + [markdown] id="vyv5qDPL2RCp" colab_type="text"
# # Step 06. Exploration - Data Schema View
# Now let's load the DataFrame and see the schema view of the Spark dataset
# + id="KPXZWq4X2RMV" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 102} outputId="8cb4db1b-a768-4051-ef15-a51346c47921"
df = spark.read.csv('revenue.csv', header = True, inferSchema = True)
df.printSchema()
# + [markdown] id="xChkf1Gf5DGC" colab_type="text"
# # Step 07. Exploration - Data Type Overview
# Now let's see data types of all available fields of the Spark dataset
# + id="8kIzQHQ35Cf1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="c2a3e14b-9367-4f79-92a8-9d1d2acb5ab6"
df.dtypes
# + [markdown] id="0QIDzJSb-4nh" colab_type="text"
# # Step 08. Exploration - More Statistical Insights from Data
# Now we'll grab total number of entries and other statistical analysis of the Spark dataset to have an overview of data
# + id="gSA4uWpd9-jL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 187} outputId="feaf8150-cecb-47c8-a2ba-23e5977c59d1"
df.describe().show()
# + [markdown] id="LJWm-tno9gq0" colab_type="text"
# # Step 09. Exploration - Total Unique Row Count
# Now we'll grab total number of unique entries or unique row count of the Spark dataset to have an overview of duplicate data
# + id="2qF1rbIN7XTo" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="322c208f-833f-46eb-ea50-d18ec492d5ff"
print("Unique Rows: ")
df.distinct().count()
# + [markdown] id="PZfgybnr95iA" colab_type="text"
# # Step 10. Implementation - Run the SQL Command
# Now since we got the idea that there is no NULL values and Duplicate rows, we can straightly go for executing SQL command to get the desired outcome. As a part of optimisation, we can drop of the column week_number as this is not relevant to this problem.
# + id="EA037xxSLEfF" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="b5f9d1f2-7597-49f3-b491-d0e7c6dc2653"
from pyspark.sql.functions import desc
df2 = df.drop(df.week_number)
Easy01 = df2.groupBy('msisdn').agg({'revenue_usd':'sum'}).sort(desc("sum(revenue_usd)"))
Easy01 = Easy01.withColumnRenamed("sum(revenue_usd)", "TotalRevenue_USD")
Easy01 = Easy01.withColumnRenamed("msisdn", "User")
Easy01.show(n=5, truncate=False)
| robi_datathon_19_easy_01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/codename-602/UseKoBERT/blob/main/KorQuad1_0%EA%B8%B0%EA%B3%84%EB%8F%85%ED%95%B4.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="0c6n1T3MgwtD"
# ## 환경 준비
# + id="-R-Ea18ygwtJ" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="75078c8e-eb63-4491-9bdd-9378c9ed05a0"
# !wget https://raw.githubusercontent.com/NLP-kr/tensorflow-ml-nlp-tf2/master/requirements.txt -O requirements.txt
# !pip install -r requirements.txt
# !pip install tensorflow==2.2.0
# + id="B9WLyWEWgdDR"
import os
import re
import json
import string
import numpy as np
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import layers
from tokenizers import BertWordPieceTokenizer
from transformers import BertTokenizer, TFBertModel
from tensorflow.keras.callbacks import EarlyStopping, ModelCheckpoint
import matplotlib.pyplot as plt
import urllib
MAX_LEN = 384
EPOCHS = 3
VERBOSE = 2
BATCH_SIZE = 16
# + id="68HVB3dYgi0w"
DATA_OUT_PATH = './data_out/KOR'
# + id="zvoswBdyglTQ"
def plot_graphs(history, string, string_1, string_2):
# loss
plt.plot(history.history[string])
plt.plot(history.history[string_1])
plt.plot(history.history[string_2])
plt.xlabel("Epochs")
plt.ylabel(string)
plt.legend([string, string_1, string_2])
plt.show()
# + id="4p79S-U1gwtK"
SEED_NUM = 1234
tf.random.set_seed(SEED_NUM)
np.random.seed(SEED_NUM)
# + id="HDI_cm3sgm6N" colab={"base_uri": "https://localhost:8080/", "height": 67, "referenced_widgets": ["1f3c34aa4d814a9490e0e803fc9a502b", "1a1d22f38e154302951fde1dad54248d", "7ecd4f87ed7446e2812e956abd9dc62d", "318901396a05480eb576aab1f80210cc", "d708fb1b7a664afea546a5fb32d5d4b3", "94ccedafdbe249b1979d36d2e2a23f11", "ae5692760b54401688024fa0cf527121", "7c0255d1a3de4bf5a51f8183d8a7c65b"]} outputId="92a89a05-9183-4008-c484-2a6f769f98fe"
# Save the slow pretrained tokenizer
slow_tokenizer = BertTokenizer.from_pretrained("bert-base-multilingual-cased", lowercase=False)
save_path = "bert-base-multilingual-cased/"
if not os.path.exists(save_path):
os.makedirs(save_path)
slow_tokenizer.save_pretrained(save_path)
# Load the fast tokenizer from saved file
tokenizer = BertWordPieceTokenizer("bert-base-multilingual-cased/vocab.txt", lowercase=False)
# + id="an5cGi-GgpG4" colab={"base_uri": "https://localhost:8080/"} outputId="1e5cebb7-fd0e-4410-d609-f0ffdd8a289e"
train_data_url = "https://korquad.github.io/dataset/KorQuAD_v1.0_train.json"
train_path = keras.utils.get_file("train.json", train_data_url)
eval_data_url = "https://korquad.github.io/dataset/KorQuAD_v1.0_dev.json"
eval_path = keras.utils.get_file("eval.json", eval_data_url)
# + id="iEgXeM2kgwtL" colab={"base_uri": "https://localhost:8080/"} outputId="a8f96315-1d83-4d08-b46b-cddf82eb76b2"
# !wget -P ./bert-base-multilingual-cased/ https://s3.amazonaws.com/models.huggingface.co/bert/bert-base-multilingual-cased-config.json
# + id="WWkLhEvtgwtL"
# !mv ./bert-base-multilingual-cased/bert-base-multilingual-cased-config.json ./bert-base-multilingual-cased/config.json
# + id="bp-0a9TbgwtM" colab={"base_uri": "https://localhost:8080/"} outputId="04c063db-d902-4471-e056-bda3c0b690e6"
# !wget -P ./bert-base-multilingual-cased/ https://s3.amazonaws.com/models.huggingface.co/bert/bert-base-multilingual-cased-tf_model.h5
# + id="XesV3eH7gwtM"
# !mv ./bert-base-multilingual-cased/bert-base-multilingual-cased-tf_model.h5 ./bert-base-multilingual-cased/tf_model.h5
# + id="PkuK7N_ngrMd"
class SquadExample:
def __init__(self, question, context, start_char_idx, answer_text):
self.question = question
self.context = context
self.start_char_idx = start_char_idx
self.answer_text = answer_text
self.skip = False
def preprocess(self):
context = self.context
question = self.question
answer_text = self.answer_text
start_char_idx = self.start_char_idx
# Clean context, answer and question
context = " ".join(str(context).split())
question = " ".join(str(question).split())
answer = " ".join(str(answer_text).split())
# Find end character index of answer in context
end_char_idx = start_char_idx + len(answer)
if end_char_idx >= len(context):
self.skip = True
return
# Mark the character indexes in context that are in answer
is_char_in_ans = [0] * len(context)
for idx in range(start_char_idx, end_char_idx):
is_char_in_ans[idx] = 1
# Tokenize context
tokenized_context = tokenizer.encode(context)
# Find tokens that were created from answer characters
ans_token_idx = []
for idx, (start, end) in enumerate(tokenized_context.offsets):
if sum(is_char_in_ans[start:end]) > 0:
ans_token_idx.append(idx)
if len(ans_token_idx) == 0:
self.skip = True
return
# Find start and end token index for tokens from answer
start_token_idx = ans_token_idx[0]
end_token_idx = ans_token_idx[-1]
# Tokenize question
tokenized_question = tokenizer.encode(question)
# Create inputs
input_ids = tokenized_context.ids + tokenized_question.ids[1:]
token_type_ids = [0] * len(tokenized_context.ids) + [1] * len(
tokenized_question.ids[1:]
)
attention_mask = [1] * len(input_ids)
# Pad and create attention masks.
# Skip if truncation is needed
padding_length = MAX_LEN - len(input_ids)
if padding_length > 0: # pad
input_ids = input_ids + ([0] * padding_length)
attention_mask = attention_mask + ([0] * padding_length)
token_type_ids = token_type_ids + ([0] * padding_length)
elif padding_length < 0: # skip
self.skip = True
return
self.input_ids = input_ids
self.token_type_ids = token_type_ids
self.attention_mask = attention_mask
self.start_token_idx = start_token_idx
self.end_token_idx = end_token_idx
self.context_token_to_char = tokenized_context.offsets
def create_squad_examples(raw_data):
squad_examples = []
for item in raw_data["data"]:
for para in item["paragraphs"]:
context = para["context"]
for qa in para["qas"]:
question = qa["question"]
answer_text = qa["answers"][0]["text"]
start_char_idx = qa["answers"][0]["answer_start"]
squad_eg = SquadExample(
question, context, start_char_idx, answer_text
)
squad_eg.preprocess()
squad_examples.append(squad_eg)
return squad_examples
def create_inputs_targets(squad_examples):
dataset_dict = {
"input_ids": [],
"token_type_ids": [],
"attention_mask": [],
"start_token_idx": [],
"end_token_idx": [],
}
for item in squad_examples:
if item.skip == False:
for key in dataset_dict:
dataset_dict[key].append(getattr(item, key))
for key in dataset_dict:
dataset_dict[key] = np.array(dataset_dict[key])
x = [
dataset_dict["input_ids"],
dataset_dict["token_type_ids"],
dataset_dict["attention_mask"],
]
y = [dataset_dict["start_token_idx"], dataset_dict["end_token_idx"]]
return x, y
# + id="R6PaDmHbgwtO" colab={"base_uri": "https://localhost:8080/"} outputId="146c250e-35d7-48d8-b183-8aad9c6b9df5"
with open(train_path) as f:
raw_train_data = json.load(f)
with open(eval_path) as f:
raw_eval_data = json.load(f)
train_squad_examples = create_squad_examples(raw_train_data)
x_train, y_train = create_inputs_targets(train_squad_examples)
print(f"{len(train_squad_examples)} training points created.")
eval_squad_examples = create_squad_examples(raw_eval_data)
x_eval, y_eval = create_inputs_targets(eval_squad_examples)
print(f"{len(eval_squad_examples)} evaluation points created.")
# + id="mIjk3_XeguBj"
class TFBERTQuestionAnswering(tf.keras.Model):
def __init__(self, model_name, dir_path, num_class):
super(TFBERTQuestionAnswering, self).__init__()
self.encoder = TFBertModel.from_pretrained(model_name, cache_dir=dir_path)
self.start_logit = tf.keras.layers.Dense(num_class, name="start_logit", use_bias=False)
self.end_logit = tf.keras.layers.Dense(num_class, name="end_logit", use_bias=False)
self.flatten = tf.keras.layers.Flatten()
self.softmax = tf.keras.layers.Activation(tf.keras.activations.softmax)
def call(self, inputs):
input_ids, token_type_ids, attention_mask = inputs
embedding = self.encoder(input_ids, token_type_ids=token_type_ids, attention_mask=attention_mask)[0]
start_logits = self.start_logit(embedding)
start_logits = self.flatten(start_logits)
end_logits = self.end_logit(embedding)
end_logits = self.flatten(end_logits)
start_probs = self.softmax(start_logits)
end_probs = self.softmax(end_logits)
return start_probs, end_probs
# + id="k4t_2T7vgwOu" colab={"base_uri": "https://localhost:8080/"} outputId="5189f27b-f64b-442c-fa9e-6760e22b6d34"
korquad_model = TFBERTQuestionAnswering(model_name='./bert-base-multilingual-cased/',dir_path='bert_ckpt', num_class=1)
optimizer = tf.keras.optimizers.Adam(learning_rate=5e-5)
loss = keras.losses.SparseCategoricalCrossentropy(from_logits=False)
# + id="YZtVFA3PgyL0"
def normalized_answer(s):
def remove_(text):
''' 불필요한 기호 제거 '''
text = re.sub("'", " ", text)
text = re.sub('"', " ", text)
text = re.sub('《', " ", text)
text = re.sub('》', " ", text)
text = re.sub('<', " ", text)
text = re.sub('>', " ", text)
text = re.sub('〈', " ", text)
text = re.sub('〉', " ", text)
text = re.sub("\(", " ", text)
text = re.sub("\)", " ", text)
text = re.sub("‘", " ", text)
text = re.sub("’", " ", text)
return text
def white_space_fix(text):
return ' '.join(text.split())
def remove_punc(text):
exclude = set(string.punctuation)
return ''.join(ch for ch in text if ch not in exclude)
def lower(text):
return text.lower()
return white_space_fix(remove_punc(lower(remove_(s))))
# + id="rVTh1qKng1p8"
class ExactMatch(keras.callbacks.Callback):
def __init__(self, x_eval, y_eval):
self.x_eval = x_eval
self.y_eval = y_eval
def on_epoch_end(self, epoch, logs=None):
pred_start, pred_end = self.model.predict(self.x_eval)
count = 0
eval_examples_no_skip = [_ for _ in eval_squad_examples if _.skip == False]
for idx, (start, end) in enumerate(zip(pred_start, pred_end)):
squad_eg = eval_examples_no_skip[idx]
offsets = squad_eg.context_token_to_char
start = np.argmax(start)
end = np.argmax(end)
if start >= len(offsets):
continue
pred_char_start = offsets[start][0]
if end < len(offsets):
pred_char_end = offsets[end][1]
pred_ans = squad_eg.context[pred_char_start:pred_char_end]
else:
pred_ans = squad_eg.context[pred_char_start:]
normalized_pred_ans = normalized_answer(pred_ans)
normalized_true_ans = normalized_answer(squad_eg.answer_text)
if normalized_pred_ans in normalized_true_ans:
count += 1
acc = count / len(self.y_eval[0])
print(f"\nepoch={epoch+1}, exact match score={acc:.2f}")
# + id="sTgvtk0og4Ow"
exact_match_callback = ExactMatch(x_eval, y_eval)
# + id="7EuBYS58g6QZ"
korquad_model.compile(optimizer=optimizer, loss=[loss, loss])
# + id="ZehxFPSrg8Q2" colab={"base_uri": "https://localhost:8080/"} outputId="0ecdb400-1b7a-4980-d888-ea56ec89d793"
model_name = "tf2_bert_korquad"
checkpoint_path = os.path.join(DATA_OUT_PATH, model_name, 'weights.h5')
checkpoint_dir = os.path.dirname(checkpoint_path)
# Create path if exists
if os.path.exists(checkpoint_dir):
print("{} -- Folder already exists \n".format(checkpoint_dir))
else:
os.makedirs(checkpoint_dir, exist_ok=True)
print("{} -- Folder create complete \n".format(checkpoint_dir))
cp_callback = ModelCheckpoint(
checkpoint_path, verbose=1, save_best_only=True, save_weights_only=True)
# + id="2ljuajCLmyws" colab={"base_uri": "https://localhost:8080/"} outputId="855a91ee-39a6-4bc6-b236-a65ee30eff83"
history = korquad_model.fit(
x_train,
y_train,
epochs=EPOCHS, # For demonstration, 3 epochs are recommended
verbose=VERBOSE,
batch_size=BATCH_SIZE,
callbacks=[exact_match_callback, cp_callback]
)
# + id="XKakfI_TgwtQ" colab={"base_uri": "https://localhost:8080/"} outputId="a6b8a51d-2350-4b80-b6b7-bc9ab1dd8e8b"
print(history.history)
# + id="QxaigHy2m4JB" colab={"base_uri": "https://localhost:8080/", "height": 279} outputId="8e5a8788-e3ac-4a7f-9775-eb49c77e5d7f"
plot_graphs(history, 'loss', 'output_1_loss', 'output_2_loss')
# + id="_RQUXGwlgwtR"
| KorQuad1_0기계독해.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Deep Learning for Audio Part 2a - Pre-process UrbanSound Datset
# ## Introduction
# In this jupyter notebook, we will process the audio files and extract the useful features that will be fed into a Convolutional Neural Network.
#
#
#
# We will train and predict on [UrbanSound8K](https://serv.cusp.nyu.edu/projects/urbansounddataset/download-urbansound8k.html) dataset. There are a few published benchmarks, which are mentioned in the papers below:
#
# - [Environmental sound classification with convolutional neural networks](http://karol.piczak.com/papers/Piczak2015-ESC-ConvNet.pdf) by <NAME>.
# - [Deep convolutional neural networks and data augmentation for environmental sound classification](https://arxiv.org/abs/1608.04363) by <NAME> and <NAME>
# - [Learning from Between-class Examples for Deep Sound Recognition](https://arxiv.org/abs/1711.10282) by <NAME>, <NAME>, <NAME>
#
#
# The state-of-art result is from the last paper by Tokozume et al., where the best error rate achieved is 21.7%. In this tutorial we will show you how to build a neural network that can achieve the state-of-art performance using Azure.
#
#
# This jupyter notebook borrows some of the pre-processing code on the Github Repo here: http://aqibsaeed.github.io/2016-09-24-urban-sound-classification-part-2/, but with a lot of modifications. It is tested with **Python3.5**, **Keras 2.1.2** and **Tensorflow 1.4.0**.
# ## Setup
# We will use librosa as our audio processing library. For more details on librosa, please refer to the librosa documenent [here](https://librosa.github.io/librosa/tutorial.html). We also need to install a bunch of libraries. Most of them are python packages, but you still may need to install a few audio processing libraries using apt-get:
#
# `sudo apt-get install -y --no-install-recommends \
# openmpi-bin \
# build-essential \
# autoconf \
# libtool \
# libav-tools \
# pkg-config`
#
#
# We also need to install librosa and a few other deep learning libraries in pip:
#
# `pip install librosa pydot graphviz keras tensorflow-gpu`
#
# ## Download dataset
# Due to licensing issues, we cannot download the data directly. Please go to the [UrbanSound8K Download](https://serv.cusp.nyu.edu/projects/urbansounddataset/download-urbansound8k.html) site, fill in the related information, download from there, and put it in the right place. You need to update the `parent_path` and `save_dir` below. In this particular case, we don't need the label file, as the labels are already reflected in the file names. We will parse the labels directly from the file names.
# ## Import libraries and initialize global varaibles
# +
import glob
import os
import librosa
import numpy as np
from joblib import Parallel, delayed
# used to featurize the dataset
from scipy import signal
# how many classes do we have; for one-hot encoding and parallel processing purpose
num_total_classes = 10
# Where you have saved the UrbanSound8K data set. Need to be absolute path.
parent_dir = "/mnt/UrbanSound8K/audio"
# specify bands that you want to use. This is also the "height" of the spectrogram image
n_bands = 150
# specify frames that you want to use. This is also the "width" of the spectrogram image
n_frames = 150
# sample rate of the target files
sample_rate = 22050
# update this part to produce different images
save_dir = "/mnt/us8k-" + str(n_bands) + "bands-" + str(n_frames) + "frames-3channel"
# -
# ## Preprocessing the Data
# The choice of the length of the sliding window used to featurize the data into a mel spectrogram is empirical – based on [Environmental sound classification with convolutional neural networks](http://karol.piczak.com/papers/Piczak2015-ESC-ConvNet.pdf) paper by Piczak, longer windows seems to perform better than shorter windows. In this blog, we will use a sliding window with a length of 2s with a 1 second overlapping; this will also determines the width of our spectrogram.
#
# +
# Read wav helper method to force audio resampling
# duration is set for a 4 second clip
def read_audio(audio_path, target_fs=None, duration=4):
(audio, fs) = librosa.load(audio_path, sr=None, duration=duration)
# if this is not a mono sounds file
if audio.ndim > 1:
audio = np.mean(audio, axis=1)
if target_fs is not None and fs != target_fs:
audio = librosa.resample(audio, orig_sr=fs, target_sr=target_fs)
fs = target_fs
return audio, fs
def pad_trunc_seq_rewrite(x, max_len):
"""Pad or truncate a sequence data to a fixed length.
Args:
x: ndarray, input sequence data.
max_len: integer, length of sequence to be padded or truncated.
Returns:
ndarray, Padded or truncated input sequence data.
"""
if x.shape[1] < max_len:
pad_shape = (x.shape[0], max_len - x.shape[1])
pad = np.ones(pad_shape) * np.log(1e-8)
#x_new = np.concatenate((x, pad), axis=1)
x_new = np.hstack((x, pad))
# no pad necessary - truncate
else:
x_new = x[:, 0:max_len]
return x_new
# +
def extract_features(parent_dir, sub_dirs, bands, frames, file_ext="*.wav"):
# 4 second clip with 50% window overlap with small offset to guarantee frames
n_window = int(sample_rate * 4. / frames * 2) - 4 * 2
# 50% overlap
n_overlap = int(n_window / 2.)
# Mel filter bank
melW = librosa.filters.mel(sr=sample_rate, n_fft=n_window, n_mels=bands, fmin=0., fmax=8000.)
# Hamming window
ham_win = np.hamming(n_window)
log_specgrams_list = []
labels = []
for l, sub_dir in enumerate(sub_dirs):
for fn in glob.glob(os.path.join(parent_dir, sub_dir, file_ext)):
# print("processing", fn)
sound_clip, fn_fs = read_audio(fn, target_fs=sample_rate)
assert (int(fn_fs) == sample_rate)
if sound_clip.shape[0] < n_window:
print("File %s is shorter than window size - DISCARDING - look into making the window larger." % fn)
continue
label = fn.split('fold')[1].split('-')[1]
# Skip corrupted wavs
if sound_clip.shape[0] == 0:
print("File %s is corrupted!" % fn)
continue
# raise NameError("Check filename - it's an empty sound clip.")
# Compute spectrogram
[f, t, x] = signal.spectral.spectrogram(
x=sound_clip,
window=ham_win,
nperseg=n_window,
noverlap=n_overlap,
detrend=False,
return_onesided=True,
mode='magnitude')
x = np.dot(x.T, melW.T)
x = np.log(x + 1e-8)
x = x.astype(np.float32).T
x = pad_trunc_seq_rewrite(x, frames)
log_specgrams_list.append(x)
labels.append(label)
log_specgrams = np.asarray(log_specgrams_list).reshape(len(log_specgrams_list), bands, frames, 1)
features = np.concatenate((log_specgrams, np.zeros(np.shape(log_specgrams))), axis=3)
features = np.concatenate((features, np.zeros(np.shape(log_specgrams))), axis=3)
for i in range(len(features)):
# first order difference, computed over 9-step window
features[i, :, :, 1] = librosa.feature.delta(features[i, :, :, 0])
# for using 3 dimensional array to use ResNet and other frameworks
features[i, :, :, 2] = librosa.feature.delta(features[i, :, :, 1])
return np.array(features), np.array(labels, dtype=np.int)
# convert labels to one-hot encoding
def one_hot_encode(labels):
n_labels = len(labels)
n_unique_labels = num_total_classes
one_hot_encode = np.zeros((n_labels, n_unique_labels))
one_hot_encode[np.arange(n_labels), labels] = 1
return one_hot_encode
# -
# ## Saving Extracted Features
# The code in the cell below can convert the raw audio files into features using multi-processing to fully utilize the CPU. The processed data are stored as numpy arrays and will be loaded during training time.
#
# It takes around 10 mins to complete - the time will vary depending on your CPU.
# +
# %%time
# use this to process the audio files into numpy arrays
def save_folds(data_dir, k, bands, frames):
fold_name = 'fold' + str(k)
print("Saving " + fold_name)
features, labels = extract_features(parent_dir, [fold_name], bands=bands, frames=frames)
labels = one_hot_encode(labels)
print("Features of", fold_name, " = ", features.shape)
print("Labels of", fold_name, " = ", labels.shape)
feature_file = os.path.join(data_dir, fold_name + '_x.npy')
labels_file = os.path.join(data_dir, fold_name + '_y.npy')
np.save(feature_file, features)
print("Saved " + feature_file)
np.save(labels_file, labels)
print("Saved " + labels_file)
def assure_path_exists(path):
mydir = os.path.join(os.getcwd(), path)
if not os.path.exists(mydir):
os.makedirs(mydir)
assure_path_exists(save_dir)
Parallel(n_jobs=num_total_classes)(delayed(save_folds)(save_dir, k, bands=n_bands, frames=n_frames) for k in range(1, 11))
| Tutorials/DeepLearningForAudio/Deep Learning for Audio Part 2a - Pre-process UrbanSound Datset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
__author__ = '<NAME> <<EMAIL>>, <NAME> <<EMAIL>>'
__version__ = '20190102' # yyyymmdd; version datestamp of this notebook
__datasets__ = ['des_dr1']
__keywords__ = ['dwarf galaxies','convolution','WCS','SIA']
# # Dwarf galaxies in DES DR1
# *<NAME>, <NAME> & NOAO Data Lab Team*
# ### Table of contents
# * [Imports & setup](#import)
# * [Coordinate resolution and querying for photometry](#photometry)
# * [The dwarf filter](#dwarffilter)
# * [Retrieve images via SIA service](#sia)
# * [Resources and references](#resources)
# # Goals
# * Resolve spatial coordinates of known dwarf companions of the Milky Way
# * Query DES DR1 for stars around these positions
# * Construct a WCS for each query and plot the spatial distribution of stars
# * Apply spatial filtering techniques to detect stellar overdensities
# * Retrieve and display image cutouts
# # Summary
# In this notebook, we demonstrate the discovery of faint Milky Way dwarf companions in DES DR1. We query the database around the positions of known dwarfs and apply filtering techniques to reveal the dwarfs as spatial overdensities of filtered sources.
#
# **Background**
# Ultrafaint dwarf galaxies are crucial to understanding many aspects of the universe. For instance, they are dominated by dark matter; their localization in space can thus trace the large-scale structure of the dark matter distribution. Furthermore, dwarf galaxies are suspected to host intermediate-mass black holes (IMBH), which so far have eluded efforts to find them. IMBHs will naturally bridge the gap between the solar-mass black hole and super-massive blackholes that reside at the center of virtually every large galaxy.
#
# **Data retrieval**
# We will retrieve stars and images around the locations of seven dwarf galaxies found in the DES DR1 catalog.
#
# **Detection**
# We will convolve the spatial distribution of our dataset with a pair of Gaussian kernels and subtract the results, as done in e.g. [Stanford et al. (2005, ApJ, 634, 2, L129)](http://adsabs.harvard.edu/abs/2005ApJ...634L.129S) (galaxy clusters), or [Koposov et al. (2008, ApJ, 686, 279)](http://adsabs.harvard.edu/abs/2008ApJ...686..279K) (MW satellites). This has the effect of convolving the spatial distribution with a Mexican hat filter, which is useful for detecting objects at a desired spatial scale.
# # Disclaimer & attribution
# If you use this notebook for your published science, please acknowledge the following:
#
# * Data Lab concept paper: Fitzpatrick et al., "The NOAO Data Laboratory: a conceptual overview", SPIE, 9149, 2014, http://dx.doi.org/10.1117/12.2057445
#
# * Data Lab disclaimer: http://datalab.noao.edu/known-issues.php
# <a class="anchor" id="import"></a>
# # Imports and initialization
# Import relevant modules and set up URL for SIA image retrieval.
# +
# std lib
from collections import OrderedDict
# 3rd party
import numpy as np
import pylab as plt
from astropy import utils, io, convolution, wcs
from astropy.visualization import make_lupton_rgb
from astropy.coordinates import name_resolve
from pyvo.dal import sia
# %matplotlib inline
# Data Lab
from dl import queryClient as qc
from dl.helpers.utils import convert
# set up Simple Image Access (SIA) service
DEF_ACCESS_URL = "http://datalab.noao.edu/sia/des_dr1"
svc = sia.SIAService(DEF_ACCESS_URL)
# -
# <a class="anchor" id="photometry"></a>
# # Coordinates and photometry data around dwarfs
# We resolve the positions of seven dwarf galaxies and query the database for stars around them.
# +
# Dwarf names from Bechtol et al. (2015)
names = ['Ret II','Eri II','Tuc II','Hor I','Pic I','Phe II','Eri III']
radius = 1. # search radius in degrees
# columns to query for
columns = '''ra,dec,mag_auto_g,mag_auto_i,mag_auto_r,mag_auto_z,flags_g,flags_i,flags_r,flags_z,
flux_auto_g,fluxerr_auto_g,flux_auto_i,fluxerr_auto_i,spread_model_g,spread_model_i,
spread_model_r,spread_model_z,class_star_g,class_star_i,class_star_r,class_star_z,
kron_radius,tilename'''
# +
# a function to retrieve data around a point in the sky
def getData(ra,dec,radius=1.0,columns='*'):
query_template =\
"""SELECT {0} FROM des_dr1.main
WHERE q3c_radial_query(ra,dec,{1},{2},{3})"""
query = query_template.format(columns,ra,dec,radius)
try:
result = qc.query(sql=query) # by default the result is a CSV formatted string
except Exception as e:
print(e.message)
df = convert(result,'pandas')
return df
# gets coordinates of a named source
def resolve_coordinates(name):
try:
coords = name_resolve.get_icrs_coordinates(name)
except Exception as e:
raise
ra = coords.ra.to('deg').value
dec = coords.dec.to('deg').value
return coords, ra, dec
# +
dgs = OrderedDict([(name,dict()) for name in names]) # empty dictionary of dictionaries
# loop over dwarfs, resolving coordinates and querying for photometry data
for j,name in enumerate(dgs.keys()):
print('{:s}: resolving coordinates and querying for data'.format(name))
coords, ra0, dec0 = resolve_coordinates(name)
dgs[name]['ra0'] = ra0
dgs[name]['dec0'] = dec0
df = getData(ra0,dec0,radius=radius,columns=columns)
dgs[name]['df'] = df
print("Done.")
# -
# <a class="anchor" id="dwarffilter"></a>
# # Filtering and plotting the dwarfs
# We filter the photometry to include objects without SE flags, with S/N>10, that are point-like (using SPREAD_MODEL, CLASS_STAR, and KRON_RADIUS--see the notebook StarGalQsoDESDR1 for details), and that are relatively blue (*g-i* < 1.0). We convolve the result with the spatial filter (defined below).
#
# Some helper functions first:
# +
# create a proper WCS object
def get_wcs(ra,dec,image,fov=1.,unit='deg',projection=("RA---TAN","DEC--TAN")):
npix = image.shape[0]
crpix = npix/2 + 1
cdelt = fov/float(npix)
w = wcs.WCS(naxis=2)
w.wcs.cunit = (unit,unit)
w.wcs.crpix = (crpix,crpix)
w.wcs.cdelt = np.array((-cdelt,cdelt))
w.wcs.ctype = projection
w.wcs.crval = (ra,dec) #coords.ra.to(unit).value, coords.dec.to(unit).value)
return w
# a spatial convolution filter
def dwarf_filter (ra,dec,fwhm_small=2.0,fwhm_big=20):
"""Differential convolution with 2D Gaussian kernels.
Based on Koposov et al. (2008).
Code by <NAME> and <NAME>.
Minor edits by RN.
Parameters
----------
ra, dec : float or array
RA & Dec in degrees.
fwhm_small, fwhm_big : float
Full-width half maximum sizes of the small and big Gaussian kernels
to use in convolution, in arcminutes.
"""
x, y = ra, dec
print("Computing differential convolution")
# Information about declination (y) [degrees]
ymean = (y.min() + y.max()) / 2.0
ydiff_arcmin = (y.max() - y.min()) * 60.0 # convert from degrees to arcmin
# Information about right ascension (x) [degrees in time]:
xdiff = x.max() - x.min() # angular separation [degrees (time)]
xmean = (x.min() + x.max()) / 2.0
# convert from degrees in time to separation in angular degrees:
xdiff_angular = (x.max() - x.min()) * np.cos(ymean*(np.pi/180.0))
# convert from degress to arcmin
xdiff_angular_arcmin = xdiff_angular * 60.0
# Get the number of one-arcmin pixels in the X and Y directions:
nx = np.rint(xdiff_angular_arcmin).astype('int')
ny = np.rint(ydiff_arcmin).astype('int')
# Create a two-dimensional histogram of the raw counts:
Counts, xedges, yedges = np.histogram2d (x, y, (nx,ny) )
extent = [xedges[0], xedges[-1], yedges[0], yedges[-1]]
raw_hist = np.rot90(Counts).copy() # hack around Pythonic weirdness
# Make the small and big Gaussian kernels with a standard deviation
# of the given FWHM in arcmin^2 pixels.
kernel_small = convolution.Gaussian2DKernel(fwhm_small/2.35,factor=1)
kernel_big = convolution.Gaussian2DKernel(fwhm_big/2.35,factor=1)
# Compute the differential convolution kernels.
conv_big = convolution.convolve(raw_hist, kernel_big)
conv_small = convolution.convolve(raw_hist, kernel_small)
conv_delta = conv_small - conv_big
delta = conv_delta.copy()
# Compute statistics and the floor
mean = np.mean(delta, dtype='float64')
sigma = np.std(delta, dtype='float64')
sigmaRaw = np.std(raw_hist,dtype='float64')
median = np.median(delta) # not used
floor = mean
clipped = delta.copy()
clipped[delta < floor] = floor
# Return the computed fields.
return raw_hist, extent, delta, clipped, sigma
# -
# Now let's run the dwarf filter on every retrieved field and plot the resulting convolved spatial distributions:
# +
nrow, ncol = 2, 4 # figure layout
# common constraints
maginvalid = 90
spread = 0.01
# set up figure
fig = plt.figure(figsize=(6*ncol,6*nrow))
# loop over dwarfs
for j,name in enumerate(dgs.keys()):
print("{:>10s}: ".format(name),end='')
dwarf = dgs[name]
df = dwarf['df'] # the Pandas dataframe object of current dwarf
# define constraining criteria
# (valid magnitudes, no flags, SNR>10, color range, likelihood(star) high-ish, stars fainter that gmag=18)
keep = (df['mag_auto_g']<maginvalid) & (df['mag_auto_i']<maginvalid) &\
(df['flags_g']==0) & (df['flags_i']==0) &\
((df['flux_auto_g']/df['fluxerr_auto_g'])>10) &\
((df['mag_auto_g']-df['mag_auto_r'])>-0.5) & ((df['mag_auto_g']-df['mag_auto_r'])<1.0) &\
(np.abs(df['spread_model_g'])<spread) & (np.abs(df['spread_model_i'])<spread) &\
(df['class_star_g']>0.1) & (df['kron_radius']<6) &\
(df['mag_auto_g']>18)
# apply constraints, and run the dwarf filter
raw_hist, extent, delta, clipped, sigma = dwarf_filter(df['ra'][keep],df['dec'][keep])
# construct a WCS
w = get_wcs(dwarf['ra0'],dwarf['dec0'],clipped,fov=1.)
# plot the clipped 2d histogram
ax = fig.add_subplot(nrow,ncol,j+1,projection=w)
im = plt.imshow(clipped)
ax.set_title(name)
print("Done.")
# -
# <a class="anchor" id="sia"></a>
# # Retrieve images (SIA service)
# We demonstrate how to retrieve images though the SIA service for one of the dwarfs. First, define two useful helper functions: one to download the deepest available image in a given band, and one to easily plot images.
# +
# download the deepest stacked images
def download_deepest_images(ra,dec,fov=0.1,bands=list('gri')):
imgTable = svc.search((ra,dec), (fov/np.cos(dec*np.pi/180), fov), verbosity=2).to_table() #.votable #.get_first_table()
print("The full image list contains {:d} entries.".format(len(imgTable)))
sel0 = (imgTable['proctype'] == b'Stack') & (imgTable['prodtype']==b'image') # basic selection
images = []
for band in bands:
print("Band {:s}: ".format(band), end='')
sel = sel0 & (imgTable['obs_bandpass'] == band.encode()) # add 'band' to selection
Table = imgTable[sel] # select
row = Table[np.argmax(Table['exptime'].data.data.astype('float'))] # pick image with longest exposure time
url = row['access_url'] # get the download URL
print('downloading deepest stacked image...')
img = io.fits.getdata(utils.data.download_file(url.decode(),cache=True,show_progress=False,timeout=120)) # .decode() b/c in Python 3 url is of "byte" type and getdata() expects "string" type
images.append(img)
print("Downloaded {:d} images.".format(len(images)))
return images
# multi panel image plotter
def plot_images(images,geo=None,panelsize=5,titles=list('gri'),cmap=plt.cm.gray_r):
if geo is None:
geo = (len(images),1) # ncols, nrows
fig = plt.figure(figsize=(geo[0]*panelsize,geo[1]*panelsize))
for j,img in enumerate(images):
ax = fig.add_subplot(geo[1],geo[0],j+1)
ax.imshow(img,origin='lower',interpolation='none',cmap=cmap,norm=plt.mpl.colors.PowerNorm(0.1))
ax.set_title('{:s}'.format(titles[j]))
plt.axis('off')
fig.subplots_adjust(wspace=0.05)
# -
# Let's download the deepest available image per band
name = '<NAME>'
fov = 0.25 # degrees
bands = list('gri')
images = download_deepest_images(dgs[name]['ra0'],dgs[name]['dec0'],fov=0.25,bands=bands)
# We also compute a false-color 3-band image, and plot all images
# %%capture --no-display
images_ = [im-np.median(im) for im in images]
images_ += [make_lupton_rgb(*images[::-1],Q=3,stretch=30)] # the func expects the images in red,green,blue order
plot_images(images_,geo=(4,1),titles=bands+['False-color 3-band image'])
# <a class="anchor" id="resources"></a>
# ## Resources and references
#
# <NAME>., et al. (2015, ApJ, 807, 50) "Eight New Milky Way Companions Discovered in First-year Dark Energy Survey Data":
# http://adsabs.harvard.edu/abs/2015ApJ...807...50B
#
# Koposov et al. (2008, ApJ, 686, 279) "The Luminosity Function of the Milky Way Satellites": http://adsabs.harvard.edu/abs/2008ApJ...686..279K
#
| 03_ScienceExamples/DwarfGalaxies/DwarfGalaxiesInDesDr1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import re
import pandas as pd
import csv
class Appearance:
"""
Represents the appearance of a term in a given document, along with the
frequency of appearances in the same one.
"""
def __init__(self, docId, frequency):
self.docId = docId
self.frequency = frequency
def __repr__(self):
"""
String representation of the Appearance object
"""
return str(self.__dict__)
# -
class Database:
"""
In memory database representing the already indexed documents.
"""
def __init__(self):
self.db = dict()
def __repr__(self):
"""
String representation of the Database object
"""
return str(self.__dict__)
def get(self, id):
return self.db.get(id, None)
def add(self, document):
"""
Adds a document to the DB.
"""
return self.db.update({document['id']: document})
def remove(self, document):
"""
Removes document from DB.
"""
return self.db.pop(document['id'], None)
class InvertedIndex:
"""
Inverted Index class.
"""
def __init__(self, db):
self.index = dict()
self.db = db
def returnIndex(self):
return self.index
def exportDictToCSV(self):
with open('mycsvfile.csv','wb') as f:
w = csv.writer(f)
w.writerows(self.index.items())
def __repr__(self):
"""
String representation of the Database object
"""
return str(self.index)
def index_document(self, document):
"""
Process a given document, save it to the DB and update the index.
"""
# Remove punctuation from the text.
clean_text = re.sub(r'[^\w\s]','', document['text'])
terms = clean_text.split(' ')
appearances_dict = dict()
# Dictionary with each term and the frequency it appears in the text.
for term in terms:
term_frequency = appearances_dict[term].frequency if term in appearances_dict else 0
appearances_dict[term] = Appearance(document['id'], term_frequency + 1)
# Update the inverted index
update_dict = { key: [appearance]
if key not in self.index
else self.index[key] + [appearance]
for (key, appearance) in appearances_dict.items() }
self.index.update(update_dict)
# Add the document into the database
self.db.add(document)
return document
def lookup_query(self, query):
"""
Returns the dictionary of terms with their correspondent Appearances.
This is a very naive search since it will just split the terms and show
the documents where they appear.
"""
return { term: self.index[term] for term in query.split(' ') if term in self.index }
import pandas as pd
data = pd.read_csv('../Data/ProcessedTweets.csv');
#data.columns = ['id','content']
#data = data[:]
# +
def highlight_term(id, term, text):
replaced_text = text.replace(term, "\033[1;32;40m {term} \033[0;0m".format(term=term))
return "--- document {id}: {replaced}".format(id=id, replaced=replaced_text)
db = Database()
index = InvertedIndex(db)
for i in range(len(data)):
text = data['content'][i]
if(pd.isnull(text)):
continue
document = {
'id': data['id'][i],
'text': data['content'][i]
}
index.index_document(document)
# -
import json
search_term = "colombia"
result = index.lookup_query(search_term)
print(result)
dic = index.returnIndex().__repr__
print(len(index.returnIndex().keys()))
with open('index.json', 'w') as json_file:
json.dump(dic(), json_file)
for term in result.keys():
for appearance in result[term]:
# Belgium: { docId: 1, frequency: 1}
document = db.get(appearance.docId)
print(highlight_term(appearance.docId, term, document['text']))
print(data.iloc[appearance.docId]['id_str'])
print()
print("-----------------------------")
| IRComponent/IRTweetsScript.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# G1953496892-LPCLOUD
# - BROWSE https://lpdaac.earthdata.nasa.gov/lp-prod-public/HLSS30.015/HLS.S30.T19TCL.2020288T154231.v1.5.jpg
right={'zone':'19', 'id':'G1953496892-LPCLOUD',
'browse':'https://lpdaac.earthdata.nasa.gov/lp-prod-public/HLSS30.015/HLS.S30.T19TCL.2020288T154231.v1.5.jpg'}
right
# G1951510836-LPCLOUD
# - BROWSE https://lpdaac.earthdata.nasa.gov/lp-prod-public/HLSS30.015/HLS.S30.T18TYR.2020283T154059.v1.5.jpg
#
#
left={'zone':'18', 'id':'G1951510836-LPCLOUD',
'browse':'https://lpdaac.earthdata.nasa.gov/lp-prod-public/HLSS30.015/HLS.S30.T18TYR.2020283T154059.v1.5.jpg'}
left
# +
from IPython.display import Image
display(Image(left['browse']))
# -
display(Image(right['browse']))
# +
import urllib
import json
#def return_assets(id):
id=left['id']
json_file = f'https://cmr.earthdata.nasa.gov/search/concepts/{id}.umm_json'
with urllib.request.urlopen(json_file) as url:
my_meta = json.loads(url.read().decode())
# -
my_meta
# 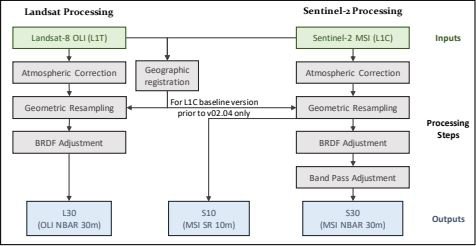
# ```
# band02 B02 2 2 0.45 – 0.51 Blue
# band03 B03 3 3 0.53 – 0.59 Green
# band04 B04 4 4 0.64 – 0.67 Red
# ```
my_meta["RelatedUrls"]
url_list = []
for ref in my_meta["RelatedUrls"]:
print(ref['URL'])
url_list.append(ref['URL'])
def return_band_url(band_name, url_list):
for ref in url_list:
if band_name in ref:
return(ref)
band_name = 'B04'
red = return_band_url(band_name, url_list)
# +
# #! mkdir -p ~/hls
# -
red
# ! pip install --user wget
# +
import sys
sys.path.insert(0, '.')
from hls_functions import urs_authenticate
urs_authenticate()
# -
# # next cell fails with unauthorized huh?
# +
# import wget
# import os
# home = os.getenv("HOME")
# output_directory = home + '/hls'
# filename = wget.download(red, out=output_directory)
# filename
# +
# #! wget https://lpdaac.earthdata.nasa.gov/lp-prod-protected/HLSS30.015/HLS.S30.T18TYR.2020283T154059.v1.5.B04.tif
# +
# #! mv HLS.S30.T18TYR.2020283T154059.v1.5.B04.tif ~/hls
# +
#os.system('wget https://lpdaac.earthdata.nasa.gov/lp-prod-protected/HLSS30.015/HLS.S30.T18TYR.2020283T154059.v1.5.B04.tif')
# +
# #!ls
# -
import urllib
import json
def return_assets(id):
json_file = f'https://cmr.earthdata.nasa.gov/search/concepts/{id}.umm_json'
with urllib.request.urlopen(json_file) as url:
my_meta = json.loads(url.read().decode())
url_list = []
for ref in my_meta["RelatedUrls"]:
print(ref['URL'])
url_list.append(ref['URL'])
return(url_list)
# +
assets = return_assets(id)
red = return_band_url(band_name, assets)
red
# -
import os
def wget_fetch(bref_url):
home=os.getenv('HOME')
filename = bref_url.split('/')[-1]
cmd = f'wget {bref_url} -O {home}/hls/{filename}'
print(cmd)
os.system(cmd)
# +
bands=['B04', 'B03', 'B02']
for band in bands:
bref = return_band_url(band, assets)
print(bref)
wget_fetch(bref)
# -
assets = return_assets(right['id'])
# +
bands=['B04', 'B03', 'B02']
for band in bands:
bref = return_band_url(band, assets)
print(bref)
wget_fetch(bref)
# -
# ! ls ~/hls/
# +
# #!ls
# +
# #! rm G*umm*
# +
# #!ls
# +
# #! rm H*.tif*
# +
# #!ls
# -
| 00-notebooks/e_display_the_browse_side_by_side_and_xarray_merge.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Computes and plots geoid undulations from a gravity model
#
# #### PYTHON DEPENDENCIES:
# - [numpy: Scientific Computing Tools For Python](https://numpy.org)
# - [matplotlib: Python 2D plotting library](http://matplotlib.org/)
# - [cartopy: Python package designed for geospatial data processing](https://scitools.org.uk/cartopy)
# - [lxml: processing XML and HTML in Python](https://pypi.python.org/pypi/lxml)
#
# #### PROGRAM DEPENDENCIES:
# - utilities.py: download and management utilities for syncing files
# - geoid_undulation.py: geoidal undulation at a given latitude and longitude
# - read_ICGEM_harmonics.py: reads the coefficients for a given gravity model file
# - calculate_tidal_offset.py: calculates the C20 offset for a tidal system
# - real_potential.py: real potential at a latitude and height for gravity model
# - norm_potential.py: normal potential of an ellipsoid at a latitude and height
# - norm_gravity.py: normal gravity of an ellipsoid at a latitude and height
# - ref_ellipsoid.py: Computes parameters for a reference ellipsoid
# - gauss_weights.py: Computes Gaussian weights as a function of degree
import numpy as np
import matplotlib
matplotlib.rcParams['axes.linewidth'] = 2.0
matplotlib.rcParams['font.family'] = 'sans-serif'
matplotlib.rcParams['font.sans-serif'] = ['Helvetica']
matplotlib.rcParams['mathtext.default'] = 'regular'
import matplotlib.pyplot as plt
import matplotlib.cm as cm
import matplotlib.colors as colors
import matplotlib.ticker as ticker
import cartopy.crs as ccrs
import ipywidgets as widgets
import geoid_toolkit.utilities
from geoid_toolkit.read_ICGEM_harmonics import read_ICGEM_harmonics
from geoid_toolkit.geoid_undulation import geoid_undulation
# #### Choose gravity model
#-- list gfc models from GFZ ICGEM
MODELS = geoid_toolkit.utilities.icgem_list()
modelDropdown = widgets.Dropdown(
options=sorted(MODELS.keys()),
value='GGM05C',
description='Model:',
disabled=False,
)
display(modelDropdown)
# #### Read gravity model coefficients
#-- gfc file with spherical harmonic coefficients
MODEL = MODELS[modelDropdown.value]
GRAVITY = geoid_toolkit.utilities.get_data_path(['data',MODEL[-1]])
MD5 = geoid_toolkit.utilities.get_hash(GRAVITY)
#-- Download coefficients from GFZ ICGEM server
geoid_toolkit.utilities.from_http(['http://icgem.gfz-potsdam.de',*MODEL],
local=GRAVITY, hash=MD5, verbose=True)
#-- use maximum degree and order of model
LMAX = None
#-- use original tide system
TIDE = None
#-- read gravity model Ylms and change tide if specified
Ylms = read_ICGEM_harmonics(GRAVITY,LMAX=LMAX,TIDE=TIDE)
#-- extract parameters
R = np.float64(Ylms['radius'])
GM = np.float64(Ylms['earth_gravity_constant'])
LMAX = np.int64(Ylms['max_degree']) if not LMAX else LMAX
# #### Calculate map of geoid height
#-- PURPOSE: calculate geoid heights at a set of latitudes and longitudes
dlon,dlat = (1.0,1.0)
lon = np.arange(-180+dlon/2.0,180+dlon/2.0,dlon)
lat = np.arange(-90+dlat/2.0,90+dlat/2.0,dlat)
nlon = len(lon)
nlat = len(lat)
#-- reference to WGS84 ellipsoid
REFERENCE = 'WGS84'
#-- Gaussian Smoothing Radius in km (default is no filtering)
#-- no gaussian smoothing
GAUSS = 0
#-- calculate geoid at coordinates
N = np.zeros((nlat,nlon))
for i in range(nlat):
N[i,:] = geoid_undulation(np.ones((nlon))*lat[i], lon, REFERENCE,
Ylms['clm'], Ylms['slm'], LMAX, R, GM, GAUSS=GAUSS)
# #### Create output plot
# +
#-- setup Plate Carree projection
fig, ax1 = plt.subplots(num=1, nrows=1, ncols=1, figsize=(10.375,6.625),
subplot_kw=dict(projection=ccrs.PlateCarree()))
#-- contours
PRANGE = (-80,80,20)
levels = np.arange(PRANGE[0],PRANGE[1]+PRANGE[2],PRANGE[2])
norm = colors.Normalize(vmin=PRANGE[0],vmax=PRANGE[1])
#-- plot image with transparency using normalization
im = ax1.imshow(N, interpolation='nearest', cmap=cm.viridis_r,
extent=(lon.min(),lon.max(),lat.min(),lat.max()),
norm=norm, alpha=1.0, transform=ccrs.PlateCarree())
#-- add generic coastlines
ax1.coastlines()
#-- draw lat/lon grid lines
GRID = [15,15]
grid_meridians = np.arange(0,360+GRID[0],GRID[0])
grid_parallels = np.arange(-90,90+GRID[1],GRID[1])
gl = ax1.gridlines(crs=ccrs.PlateCarree(), draw_labels=False,
linewidth=0.1, color='0.25', linestyle='-')
gl.xlocator = ticker.FixedLocator(grid_meridians)
gl.ylocator = ticker.FixedLocator(grid_parallels)
#-- Add horizontal colorbar and adjust size
#-- extend = add extension triangles to upper and lower bounds
#-- options: neither, both, min, max
#-- pad = distance from main plot axis
#-- shrink = percent size of colorbar
#-- aspect = lengthXwidth aspect of colorbar
cbar = plt.colorbar(im, ax=ax1, extend='both', extendfrac=0.0375,
orientation='horizontal', pad=0.025, shrink=0.90, aspect=22,
drawedges=False)
#-- rasterized colorbar to remove lines
cbar.solids.set_rasterized(True)
#-- Add label to the colorbar
cbar.ax.set_xlabel('Geoidal Undulation', labelpad=10, fontsize=20)
cbar.ax.set_ylabel('m', fontsize=20, rotation=0)
cbar.ax.yaxis.set_label_coords(1.04, 0.15)
#-- Set the tick levels for the colorbar
cbar.set_ticks(levels)
cbar.set_ticklabels(['{0:d}'.format(ct) for ct in levels])
#-- ticks lines all the way across
cbar.ax.tick_params(which='both', width=1, length=27, labelsize=20,
direction='in')
#-- axis = equal
ax1.set_aspect('equal', adjustable='box')
#-- no ticks on the x and y axes
ax1.get_xaxis().set_ticks([])
ax1.get_yaxis().set_ticks([])
#-- add main title
ax1.set_title(modelDropdown.value, fontsize=24)
ax1.title.set_y(1.01)
#-- stronger linewidth on frame
ax1.outline_patch.set_linewidth(2.0)
ax1.outline_patch.set_capstyle('projecting')
#-- output to file
fig.subplots_adjust(left=0.04,right=0.96,bottom=0.05,top=0.96)
plt.show()
# -
| notebooks/Calculate-Geoidal-Undulation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: py3.7
# language: python
# name: py3.7
# ---
# # Exercise 2.01
#
# ## Scalars, Vectors, Matrices and Tensors
#
# Let's start with some basic definitions:
#
# <em>Difference between a scalar, a vector, a matrix and a tensor</em>
#
# - A scalar is a single number
# - A vector is a one-dimensional array of numbers
# - A matrix is a two-dimensional array of numbers
# - A tensor is a $n$-dimensional array of numbers with $n>2$
#
# We will start by creating a vector. This is just a 1-dimensional array of numbers, we can use the numpy library since it includes many functions for working with and processing vectors and matrices.
import numpy as np
vec1 = np.array([1, 2, 3, 4, 5, 6, 7, 8, 9, 10])
vec1
# We can also create matrices with the same `array` function, here we can pass an array of arrays.
mat1 = np.array([[1, 2, 3], [4, 5, 6], [7, 8, 9], [10, 11, 12]])
mat1
# Another way we can create a matrix is by using the `matrix` function. We provide the function the same array of arrays to initialize the matrix.
mat2 = np.matrix([[1, 2, 3], [4, 5, 6], [7, 8, 9], [10, 11, 12]])
mat2
# The difference between creating matrices with the two format are the methods that the `array` and `matrix` class have.
#
# We can also create 3-dimensional array, or tensor, by passing an array of arrays of arrays to the `array` function.
ten1 = np.array([[[1, 2, 3], [4, 5, 6]], [[7, 8, 9], [10, 11, 12]]])
ten1
# ### Shape
#
# The shape of an vector or matrix is determined by the number of values in the vector or matrix in each dimension. For a vector the shape is determined by the total number of elements in the vector, since it is 1-dimensional. For a matrix it will give you the number of rows and columns in the matrix (in general the rows are provided first).
#
# We can determine the shape of our vectors and matrices as follows:
vec1.shape
# The value is simply 10.
#
# And for the matrices:
mat1.shape
# We can see the matrix has 4 rows and 3 columns.
#
# Finally, for tensors:
ten1.shape
# Note that we can also use the `len()` function to determine the first element of the `shape` method. For vectors this is equivalent to the shape, and for matrices this is equivalent to the number of rows.
# ## Addition
#
# Matrices can be added if they have the same shape.
# This is achieved by adding the corresponding cell from each matrix together. The resulting matric will have the same shape as the two input matrices.
#
# $$A_{i,j} + B_{i,j} = C_{i,j}$$
#
# $i$ is the row index and $j$ the column index.
#
# $$
# C=
# \begin{bmatrix}
# A_{1,1} & A_{1,2} \\\\
# A_{2,1} & A_{2,2} \\\\
# A_{3,1} & A_{3,2}
# \end{bmatrix}+
# \begin{bmatrix}
# B_{1,1} & B_{1,2} \\\\
# B_{2,1} & B_{2,2} \\\\
# B_{3,1} & B_{3,2}
# \end{bmatrix}=
# \begin{bmatrix}
# A_{1,1} + B_{1,1} & A_{1,2} + B_{1,2} \\\\
# A_{2,1} + B_{2,1} & A_{2,2} + B_{2,2} \\\\
# A_{3,1} + B_{3,1} & A_{3,2} + B_{3,2}
# \end{bmatrix}
# $$
#
# With Numpy you can add matrices just as you would add vectors or scalars.
mat1 = np.matrix([[1, 2, 3], [4, 5, 6], [7, 8, 9], [10, 11, 12]])
mat1
mat2 = np.matrix([[2, 1, 4], [4, 1, 7], [4, 2, 9], [5, 21, 1]])
mat2
# Add matrices mat1 and mat2
mat3 = mat1 + mat2
mat3
# Scalar values can also be added to matrices. This is achieved by adding the scalar value to each cell of the matrix.
#
# $$
# \alpha+ \begin{bmatrix}
# A_{1,1} & A_{1,2} \\\\
# A_{2,1} & A_{2,2} \\\\
# A_{3,1} & A_{3,2}
# \end{bmatrix}=
# \begin{bmatrix}
# \alpha + A_{1,1} & \alpha + A_{1,2} \\\\
# \alpha + A_{2,1} & \alpha + A_{2,2} \\\\
# \alpha + A_{3,1} & \alpha + A_{3,2}
# \end{bmatrix}
# $$
mat1 = np.matrix([[1, 2, 3], [4, 5, 6], [7, 8, 9], [10, 11, 12]])
mat1
mat1 + 4
| Chapter02/Exercise2.01/Exercise2_01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.12 ('COVID')
# language: python
# name: python3
# ---
# En este cuaderno tiene como producto final un gráfico de barras de la sintomatología más común de los casos pediátricos presentados en Sonora en la base de COVID-19 la cual dividimos en dos grupos: de [0,12) y [12,18].
import pandas as pd
import seaborn as sns
from matplotlib import pyplot as plt
#leemos los datos
df = pd.read_csv("PediatricoSonora.csv",parse_dates=['FECHA_ACTUALIZACION'])
#revisamos las columnas
df.columns
#damos una revisada solo a las columnas que nos interesan para ver los datos
df[["FECHA_ACTUALIZACION","SEXO","EDAD","MUNICIPIO_RES","FECHA_SINTOMAS","FECHA_DEF","EMBARAZO","DIABETES","ASMA","INMUSUPR","HIPERTENSION",'OTRO_CASO',"RENAL_CRONICA","TABAQUISMO"]]
#dataframe con los niños de [0,12) años
dfninos=df[(df["EDAD"]>=0) & (df["EDAD"]<12)]
dfninos
#data frame con los adolescentes de [12,18] años
dfteen=df[df["EDAD"]>=12]
dfteen
#damos una revisada a la tabla de niños
dfninos.columns
#como se realizará una gráfica con frecuencias, creamos una tabla "niños frecuencia" que incluya solamente las tablas que nos interesan
#segun el diccionario de la secretaría de salud 1-si, 2-no, 97-no aplica, 98-se ignora, 99-no especificado
dfninosFreq=pd.DataFrame(columns=["NEUMONIA","DIABETES","EPOC","OBESIDAD","RENAL_CRONICA","TABAQUISMO","OTRO_CASO","EMBARAZO"],index=[1,2,97,98,99])
dfninosFreq
#vamos a llenar esta tabla usando un for sobre una lista de strings, así que primero definimos la lista
Factores=["NEUMONIA","DIABETES","EPOC","OBESIDAD","TABAQUISMO","OTRO_CASO","EMBARAZO","RENAL_CRONICA"]
for factor in Factores:
dfninos.groupby([factor]).count().transpose().loc["FECHA_ACTUALIZACION"]
dfninosFreq[factor]=dfninos.groupby([factor]).count().transpose().loc["FECHA_ACTUALIZACION"]
dfninosFreq
#en este caso, los NAN son cero, por lo que los llenamos con cero
dfninosFreq.fillna(0)
#solo me interesa graficar el primer renglón, que es el de SI (si tienen la sintomatologia de la columna)
row = dfninosFreq.iloc[0]
row
plt.rcParams["figure.dpi"] = 100
ax = row.plot(kind="bar")
for p in ax.patches:
ax.annotate(str(p.get_height()), (p.get_x() * 1.005, p.get_height() * 1.005))
# +
#repetimos lo anterior pero más rápido
dfteenFreq=pd.DataFrame(columns=["NEUMONIA","DIABETES","EPOC","OBESIDAD","RENAL_CRONICA","TABAQUISMO","OTRO_CASO","EMBARAZO"],index=[1,2,97,98,99])
for factor in Factores:
dfteen.groupby([factor]).count().transpose().loc["FECHA_ACTUALIZACION"]
dfteenFreq[factor]=dfteen.groupby([factor]).count().transpose().loc["FECHA_ACTUALIZACION"]
dfteenFreq.fillna(0)
rowteen = dfteenFreq.iloc[0]
plt.rcParams["figure.dpi"] = 100
ax = rowteen.plot(kind="bar")
for p in ax.patches:
ax.annotate(str(p.get_height()), (p.get_x() * 1.005, p.get_height() * 1.005))
| AnalisisPediatrico.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Unit Testing `GiRaFFE_NRPy`: $A_k$ to $B^i$
#
# ### Author: <NAME>
#
# This notebook validates our A-to-B solver for use in `GiRaFFE_NRPy`. Because the original `GiRaFFE` used staggered grids and we do not, we can not trivially do a direct comparison to the old code. Instead, we will compare the numerical results with the expected analytic results.
#
# **Module Status:** <font color=red><b> In-Progress </b></font>
#
# **Validation Notes:** This module will validate the routines in [Tutorial-GiRaFFE_HO_C_code_library-A2B](../Tutorial-GiRaFFE_HO_C_code_library-A2B.ipynb).
#
# It is, in general, good coding practice to unit test functions individually to verify that they produce the expected and intended output. Here, we expect our functions to produce the correct cross product in an arbitrary spacetime. To that end, we will choose functions that are easy to differentiate, but lack the symmetries that would trivialize the finite-difference algorithm. Higher-order polynomials are one such type of function.
#
# We will start with the simplest case - testing the second-order solver. In second-order finite-differencing, we use a three-point stencil that can exactly differentiate polynomials up to quadratic. So, we will use cubic functions three variables. For instance,
#
# \begin{align}
# A_x &= ax^3 + by^3 + cz^3 + dy^2 + ez^2 + f \\
# A_y &= gx^3 + hy^3 + lz^3 + mx^2 + nz^2 + p \\
# A_z &= px^3 + qy^3 + rz^3 + sx^2 + ty^2 + u. \\
# \end{align}
#
# It will be much simpler to let NRPy+ handle most of this work. So, we will import the core functionality of NRPy+, build the expressions, and then output them using `outputC()`.
# +
import shutil, os, sys # Standard Python modules for multiplatform OS-level functions
# First, we'll add the parent directory to the list of directories Python will check for modules.
nrpy_dir_path = os.path.join("..")
if nrpy_dir_path not in sys.path:
sys.path.append(nrpy_dir_path)
from outputC import * # NRPy+: Core C code output module
import finite_difference as fin # NRPy+: Finite difference C code generation module
import NRPy_param_funcs as par # NRPy+: Parameter interface
import grid as gri # NRPy+: Functions having to do with numerical grids
import loop as lp # NRPy+: Generate C code loops
import indexedexp as ixp # NRPy+: Symbolic indexed expression (e.g., tensors, vectors, etc.) support
import reference_metric as rfm # NRPy+: Reference metric support
import cmdline_helper as cmd # NRPy+: Multi-platform Python command-line interface
out_dir = "Validation/"
cmd.mkdir(out_dir)
thismodule = "Unit_Test_GiRaFFE_NRPy_Ccode_library_A2B"
a,b,c,d,e,f,g,h,l,m,n,o,p,q,r,s,t,u = par.Cparameters("REAL",thismodule,["a","b","c","d","e","f","g","h","l","m","n","o","p","q","r","s","t","u"],10.0)
gammadet = gri.register_gridfunctions("AUXEVOL","gammadet")
DIM = 3
par.set_parval_from_str("grid::DIM",DIM)
par.set_parval_from_str("reference_metric::CoordSystem","Cartesian")
rfm.reference_metric()
x = rfm.xxCart[0]
y = rfm.xxCart[1]
z = rfm.xxCart[2]
AD = ixp.register_gridfunctions_for_single_rank1("EVOL","AD")
AD[0] = a*x**3 + b*y**3 + c*z**3 + d*y**2 + e*z**2 + f
AD[1] = g*x**3 + h*y**3 + l*z**3 + m*x**2 + n*z**2 + o
AD[2] = p*x**3 + q*y**3 + r*z**3 + s*x**2 + t*y**2 + u
# -
# Next, we'll let NRPy+ compute derivatives analytically according to $$B^i = \frac{[ijk]}{\sqrt{\gamma}} \partial_j A_k.$$ Then we can carry out two separate tests to verify the numerical derivatives. First, we will verify that when we let the cubic terms be zero, the two calculations of $B^i$ agree to roundoff error. Second, we will verify that when we set the cubic terms, our error is dominated by trunction error that converges to zero at the expected rate.
# +
import WeylScal4NRPy.WeylScalars_Cartesian as weyl
LeviCivitaDDD = weyl.define_LeviCivitaSymbol_rank3()
LeviCivitaUUU = ixp.zerorank3()
for i in range(DIM):
for j in range(DIM):
for k in range(DIM):
LeviCivitaUUU[i][j][k] = LeviCivitaDDD[i][j][k] / sp.sqrt(gammadet)
B_analyticU = ixp.register_gridfunctions_for_single_rank1("AUXEVOL","B_analyticU")
for i in range(DIM):
B_analyticU[i] = 0
for j in range(DIM):
for k in range(DIM):
B_analyticU[i] += LeviCivitaUUU[i][j][k] * sp.diff(AD[k],rfm.xxCart[j])
# -
# Now that we have our vector potential and analytic magnetic field to compare against, we will start writing our unit test. We'll also import common C functionality, define `REAL`, the number of ghost zones, and the faces, and set the standard macros for NRPy+ style memory access.
out_string = """
// These are common packages that we are likely to need.
#include "stdio.h"
#include "stdlib.h"
#include "math.h"
#include "string.h" // Needed for strncmp, etc.
#include "stdint.h" // Needed for Windows GCC 6.x compatibility
#include <time.h> // Needed to set a random seed.
# define REAL double
const int MAXFACE = -1;
const int NUL = +0;
const int MINFACE = +1;
const int NGHOSTS = 1;
// Standard NRPy+ memory access:
#define IDX4(g,i,j,k) \
( (i) + Nxx_plus_2NGHOSTS[0] * ( (j) + Nxx_plus_2NGHOSTS[1] * ( (k) + Nxx_plus_2NGHOSTS[2] * (g) ) ) )
#define IDX3(i,j,k) ( (i) + Nxx_plus_2NGHOSTS[0] * ( (j) + Nxx_plus_2NGHOSTS[1] * (k) ) )
// Assuming idx = IDX3(i,j,k). Much faster if idx can be reused over and over:
#define IDX4pt(g,idx) ( (idx) + (Nxx_plus_2NGHOSTS[0]*Nxx_plus_2NGHOSTS[1]*Nxx_plus_2NGHOSTS[2]) * (g) )
"""
# We'll now define the gridfunction names.
out_string += """
// Let's also #define the NRPy+ gridfunctions
#define AD0GF 0
#define AD1GF 1
#define AD2GF 2
#define NUM_EVOL_GFS 3
#define GAMMADETGF 0
#define B_ANALYTICU0GF 1
#define B_ANALYTICU1GF 2
#define B_ANALYTICU2GF 3
#define BU0GF 4
#define BU1GF 5
#define BU2GF 6
#define NUM_AUXEVOL_GFS 7
"""
# Now, we'll handle the different A2B codes. There are several things to do here. First, we'll add `#include`s to the C code so that we have access to the functions we want to test. We must also create a directory and copy the files to that directory. We will choose to do this in the subfolder `A2B` relative to this tutorial.
# +
out_string += """
#include "../A2B/driver_AtoB.c" // This file contains both functions we need.
"""
cmd.mkdir(os.path.join("A2B/"))
shutil.copy(os.path.join("../GiRaFFE_HO/GiRaFFE_Ccode_library/A2B/driver_AtoB.c"),os.path.join("A2B/"))
# -
# We also should write a function that will use the analytic formulae for $B^i$. Then, we'll need to call the function from the module `GiRaFFE_HO_A2B` to generate the different header files. Also, we will declare the parameters for the vector potential functions.
# +
out_string += """
REAL a,b,c,d,e,f,g,h,l,m,n,o,p,q,r,s,t,u;
void calculate_exact_BU(const int Nxx_plus_2NGHOSTS[3],REAL *xx[3],double *auxevol_gfs) {
for(int i2=0;i2<Nxx_plus_2NGHOSTS[2];i2++) for(int i1=0;i1<Nxx_plus_2NGHOSTS[1];i1++) for(int i0=0;i0<Nxx_plus_2NGHOSTS[0];i0++) {
REAL xx0 = xx[0][i0];
REAL xx1 = xx[1][i1];
REAL xx2 = xx[2][i2];
"""
B_analyticU_to_print = [\
lhrh(lhs=gri.gfaccess("out_gfs","B_analyticU0"),rhs=B_analyticU[0]),\
lhrh(lhs=gri.gfaccess("out_gfs","B_analyticU1"),rhs=B_analyticU[1]),\
lhrh(lhs=gri.gfaccess("out_gfs","B_analyticU2"),rhs=B_analyticU[2]),\
]
B_analyticU_kernel = fin.FD_outputC("returnstring",B_analyticU_to_print,params="outCverbose=False")
out_string += B_analyticU_kernel
out_string += """
}
}
"""
gri.glb_gridfcs_list = []
import GiRaFFE_HO.GiRaFFE_HO_A2B as A2B
# We'll generate these into the A2B subdirectory since that's where the functions
# we're testing expect them to be.
A2B.GiRaFFE_HO_A2B("A2B/")
# -
# We'll now write a function to set the vector potential $A_k$. This simply uses NRPy+ to generte most of the code from the expressions we wrote at the beginning.
# +
out_string += """
void calculate_AD(const int Nxx_plus_2NGHOSTS[3],REAL *xx[3],double *out_gfs) {
for(int i2=0;i2<Nxx_plus_2NGHOSTS[2];i2++) for(int i1=0;i1<Nxx_plus_2NGHOSTS[1];i1++) for(int i0=0;i0<Nxx_plus_2NGHOSTS[0];i0++) {
REAL xx0 = xx[0][i0];
REAL xx1 = xx[1][i1];
REAL xx2 = xx[2][i2];
"""
AD_to_print = [\
lhrh(lhs=gri.gfaccess("out_gfs","AD0"),rhs=AD[0]),\
lhrh(lhs=gri.gfaccess("out_gfs","AD1"),rhs=AD[1]),\
lhrh(lhs=gri.gfaccess("out_gfs","AD2"),rhs=AD[2]),\
]
AD_kernel = fin.FD_outputC("returnstring",AD_to_print,params="outCverbose=False")
out_string += AD_kernel
out_string += """
}
}
"""
# -
# We will define the extent of our grid here.
out_string += """
const REAL xmin = -0.01,xmax=0.01;
const REAL ymin = -0.01,ymax=0.01;
const REAL zmin = -0.01,zmax=0.01;
"""
# Now, we'll write the main method. First, we'll set up the grid. In this test, we cannot use only one point. As we are testing a three-point stencil, we can get away with a minimal $3 \times 3 \times 3$ grid. Then, we'll write the A fields. After that, we'll calculate the magnetic field two ways.
out_string += """
main(int argc, const char *argv[]) {
// Let the first argument be the test we're doing. 1 = coarser grid, 0 = finer grid.
int do_quadratic_test = atoi(argv[4]);
// We'll use this grid. It has one point and one ghost zone.
const int Nxx[3] = {atoi(argv[1]),atoi(argv[2]),atoi(argv[3])};
int Nxx_plus_2NGHOSTS[3];
for (int i=0;i<3;i++) Nxx_plus_2NGHOSTS[i] = Nxx[i]+2*NGHOSTS;
const REAL xxmin[3] = {xmin,ymin,zmin};
const REAL xxmax[3] = {xmax,ymax,zmax};
REAL dxx[3];
for(int i=0;i<3;i++) dxx[i] = (xxmax[i] - xxmin[i]) / ((REAL)Nxx[i]+1.0);
// We'll define our grid slightly different from how we normally would. We let our outermost
// ghostzones coincide with xxmin and xxmax instead of the interior of the grid. This means
// that the ghostzone points will have identical positions so we can do convergence tests of them.
// Step 0d.ii: Set up uniform coordinate grids
REAL *xx[3];
for(int i=0;i<3;i++) {
xx[i] = (REAL *)malloc(sizeof(REAL)*Nxx_plus_2NGHOSTS[i]);
for(int j=0;j<Nxx_plus_2NGHOSTS[i];j++) {
xx[i][j] = xxmin[i] + ((REAL)(j))*dxx[i]; // Face-centered grid.
}
}
//for(int i=0;i<Nxx_plus_2NGHOSTS[0];i++) printf("xx[0][%d] = %.15e\\n",i,xx[0][i]);
// This is the array to which we'll write the NRPy+ variables.
REAL *auxevol_gfs = (REAL *)malloc(sizeof(REAL) * NUM_AUXEVOL_GFS * Nxx_plus_2NGHOSTS[2] * Nxx_plus_2NGHOSTS[1] * Nxx_plus_2NGHOSTS[0]);
REAL *evol_gfs = (REAL *)malloc(sizeof(REAL) * NUM_EVOL_GFS * Nxx_plus_2NGHOSTS[2] * Nxx_plus_2NGHOSTS[1] * Nxx_plus_2NGHOSTS[0]);
for(int i2=0;i2<Nxx_plus_2NGHOSTS[2];i2++) for(int i1=0;i1<Nxx_plus_2NGHOSTS[1];i1++) for(int i0=0;i0<Nxx_plus_2NGHOSTS[0];i0++) {
//auxevol_gfs[IDX4(GAMMADETGF,i0,i1,i2)] = 1.0; // Flat Space
auxevol_gfs[IDX4(GAMMADETGF,i0,i1,i2)] = 1.0 - 1.0/(2.0+xx[0][i0]*xx[0][i0]+xx[1][i1]*xx[1][i1]+xx[2][i2]*xx[2][i2]);
}
// We now want to set up the vector potential. First, we must set the coefficients.
// We will use random integers between -10 and 10. For the first test, we let the
// Cubic coefficients remain zero. Those are a,b,c,g,h,l,p,q, and r.
d = (double)(rand()%20-10);
e = (double)(rand()%20-10);
f = (double)(rand()%20-10);
m = (double)(rand()%20-10);
n = (double)(rand()%20-10);
o = (double)(rand()%20-10);
s = (double)(rand()%20-10);
t = (double)(rand()%20-10);
u = (double)(rand()%20-10);
if(do_quadratic_test) {
calculate_AD(Nxx_plus_2NGHOSTS,xx,evol_gfs);
// We'll also calculate the exact solution for B^i
calculate_exact_BU(Nxx_plus_2NGHOSTS,xx,auxevol_gfs);
// And now for the numerical derivatives:
driver_A_to_B(Nxx,Nxx_plus_2NGHOSTS,dxx,evol_gfs,auxevol_gfs);
printf("This test uses quadratic vector potentials, so the magnetic fields should agree to roundoff error.\\n");
printf("Below, each row represents one point. Each column represents a component of the magnetic field.\\n");
printf("Shown is the number of Significant Digits of Agreement, at least 13 is good, higher is better:\\n\\n");
//Two variables for inside the loop:
int ghost_zone_overlap;int indices[3];
for(int i2=0;i2<Nxx_plus_2NGHOSTS[2];i2++) for(int i1=0;i1<Nxx_plus_2NGHOSTS[1];i1++) for(int i0=0;i0<Nxx_plus_2NGHOSTS[0];i0++) {
// Are we on an edge/vertex? This algorithm can probably be improved.
ghost_zone_overlap = 0;
indices[0] = i0;
indices[1] = i1;
indices[2] = i2;
for(int dim=0;dim<3;dim++) {
if(indices[dim]%(Nxx[dim]+NGHOSTS)<NGHOSTS) {
ghost_zone_overlap++;
}
}
if (ghost_zone_overlap < 2) {
// Don't print if we're on an edge or vertex
printf("SDA: %.3f, %.3f, %.3f\\n",
1.0-log10(2.0*fabs(auxevol_gfs[IDX4(B_ANALYTICU0GF,i0,i1,i2)]-auxevol_gfs[IDX4(BU0GF,i0,i1,i2)])/(fabs(auxevol_gfs[IDX4(B_ANALYTICU0GF,i0,i1,i2)])+fabs(auxevol_gfs[IDX4(BU0GF,i0,i1,i2)])+1.e-15)),
1.0-log10(2.0*fabs(auxevol_gfs[IDX4(B_ANALYTICU1GF,i0,i1,i2)]-auxevol_gfs[IDX4(BU1GF,i0,i1,i2)])/(fabs(auxevol_gfs[IDX4(B_ANALYTICU1GF,i0,i1,i2)])+fabs(auxevol_gfs[IDX4(BU1GF,i0,i1,i2)])+1.e-15)),
1.0-log10(2.0*fabs(auxevol_gfs[IDX4(B_ANALYTICU2GF,i0,i1,i2)]-auxevol_gfs[IDX4(BU2GF,i0,i1,i2)])/(fabs(auxevol_gfs[IDX4(B_ANALYTICU2GF,i0,i1,i2)])+fabs(auxevol_gfs[IDX4(BU2GF,i0,i1,i2)])+1.e-15))
);
}
}
}
// Now, we'll set the cubic coefficients:
a = (double)(rand()%20-10);
b = (double)(rand()%20-10);
c = (double)(rand()%20-10);
g = (double)(rand()%20-10);
h = (double)(rand()%20-10);
l = (double)(rand()%20-10);
p = (double)(rand()%20-10);
q = (double)(rand()%20-10);
r = (double)(rand()%20-10);
// And recalculate on our initial grid:
calculate_AD(Nxx_plus_2NGHOSTS,xx,evol_gfs);
// We'll also calculate the exact solution for B^i
calculate_exact_BU(Nxx_plus_2NGHOSTS,xx,auxevol_gfs);
// And now for the numerical derivatives:
driver_A_to_B(Nxx,Nxx_plus_2NGHOSTS,dxx,evol_gfs,auxevol_gfs);
// Some variables needed for the loop:
int ghost_zone_overlap; int indices[3];
char filename[100];
sprintf(filename,"out%d-numer.txt",Nxx[0]);
FILE *out2D = fopen(filename, "w");
if(do_quadratic_test) {
for(int i2=0;i2<Nxx_plus_2NGHOSTS[2];i2++) for(int i1=0;i1<Nxx_plus_2NGHOSTS[1];i1++) for(int i0=0;i0<Nxx_plus_2NGHOSTS[0];i0++) {
ghost_zone_overlap = 0;
indices[0] = i0;
indices[1] = i1;
indices[2] = i2;
for(int dim=0;dim<3;dim++) {
if(indices[dim]%(Nxx[dim]+NGHOSTS)<NGHOSTS) {
ghost_zone_overlap++;
}
}
if (ghost_zone_overlap < 2) {
// We print the difference between approximate and exact numbers.
fprintf(out2D,"%.16e\t%.16e\t%.16e\\n",
auxevol_gfs[IDX4(B_ANALYTICU0GF,i0,i1,i2)]-auxevol_gfs[IDX4(BU0GF,i0,i1,i2)],
auxevol_gfs[IDX4(B_ANALYTICU1GF,i0,i1,i2)]-auxevol_gfs[IDX4(BU1GF,i0,i1,i2)],
auxevol_gfs[IDX4(B_ANALYTICU2GF,i0,i1,i2)]-auxevol_gfs[IDX4(BU2GF,i0,i1,i2)]
);
}
}
}
else {
for(int i2=0;i2<Nxx_plus_2NGHOSTS[2];i2++) for(int i1=0;i1<Nxx_plus_2NGHOSTS[1];i1++) for(int i0=0;i0<Nxx_plus_2NGHOSTS[0];i0++) {
ghost_zone_overlap = 0;
indices[0] = i0;
indices[1] = i1;
indices[2] = i2;
for(int dim=0;dim<3;dim++) {
if(indices[dim]%(Nxx[dim]+NGHOSTS)<NGHOSTS) {
ghost_zone_overlap++;
}
}
// Don't print on the edges or corners
if (ghost_zone_overlap < 2) {
// Only print points shared between the grids
if (i0%2==0 && i1%2==0 && i2%2==0) {
// We print the difference between approximate and exact numbers.
fprintf(out2D,"%.16e\t%.16e\t%.16e\\n",
auxevol_gfs[IDX4(B_ANALYTICU0GF,i0,i1,i2)]-auxevol_gfs[IDX4(BU0GF,i0,i1,i2)],
auxevol_gfs[IDX4(B_ANALYTICU1GF,i0,i1,i2)]-auxevol_gfs[IDX4(BU1GF,i0,i1,i2)],
auxevol_gfs[IDX4(B_ANALYTICU2GF,i0,i1,i2)]-auxevol_gfs[IDX4(BU2GF,i0,i1,i2)]
);
}
}
}
}
fclose(out2D);
}
"""
# Now, we must write out the code to a `.C` file.
with open(os.path.join(out_dir,"A2B_unit_test.C"),"w") as file:
file.write(out_string)
# Now that we have our file, we can compile it and run the executable.
# +
import time
print("Now compiling, should take ~2 seconds...\n")
start = time.time()
cmd.C_compile(os.path.join(out_dir,"A2B_unit_test.C"), os.path.join(out_dir,"A2B_unit_test"))
end = time.time()
print("Finished in "+str(end-start)+" seconds.\n\n")
# os.chdir(out_dir)
print("Now running...\n")
start = time.time()
# cmd.Execute(os.path.join("Stilde_flux_unit_test"))
# !./Validation/A2B_unit_test 1 1 1 1
# To do a convergence test, we'll also need a second grid with twice the resolution.
# !./Validation/A2B_unit_test 3 3 3 0
end = time.time()
print("Finished in "+str(end-start)+" seconds.\n\n")
# os.chdir(os.path.join("../"))
# -
# Now that we have shown that when we use a quadratic vector potential, we get roundoff-level agreement (which is to be expected, since the finite-differencing used approximates the underlying function with a quadratic), we will use do a convergence test to show that when we can't exactly model the function, the truncation error dominates and converges to zero at the expected rate. For this, we use cubic functions for the vector potential. In the code above, we output the difference beteween the numeric and exact magnetic fields at the overlapping, non-edge, non-vertex points of two separate grids. Here, we import that data and calculate the convergence in the usual way,
# $$
# k = \log_2 \left( \frac{F - F_1}{F - F_2} \right),
# $$
# where $k$ is the convergence order, $F$ is the exact solution, $F_1$ is the approximate solution on the coarser grid with resolution $\Delta x$, and $F_2$ is the approximate solution on the finer grid with resolution $\Delta x/2$.
# +
import numpy as np
import matplotlib.pyplot as plt
Data1 = np.loadtxt("out1-numer.txt")
Data2 = np.loadtxt("out3-numer.txt")
convergence = np.log(np.divide(np.abs(Data1),np.abs(Data2)))/np.log(2)
print("Convergence test: All should be approximately 2\n")
print(convergence)
| GiRaFFE_standalone_Ccodes/Tutorial-Unit_Test-GiRaFFE_NRPy_Ccode_library-A2B.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# # Nearest Neighbors regression
#
#
# Demonstrate the resolution of a regression problem
# using a k-Nearest Neighbor and the interpolation of the
# target using both barycenter and constant weights.
#
# +
print(__doc__)
# Author: <NAME> <<EMAIL>>
# <NAME> <<EMAIL>>
#
# License: BSD 3 clause (C) INRIA
# #############################################################################
# Generate sample data
import numpy as np
import matplotlib.pyplot as plt
from sklearn import neighbors
np.random.seed(0)
X = np.sort(5 * np.random.rand(40, 1), axis=0)
T = np.linspace(0, 5, 500)[:, np.newaxis]
y = np.sin(X).ravel()
# Add noise to targets
y[::5] += 1 * (0.5 - np.random.rand(8))
# #############################################################################
# Fit regression model
n_neighbors = 5
for i, weights in enumerate(['uniform', 'distance']):
knn = neighbors.KNeighborsRegressor(n_neighbors, weights=weights)
y_ = knn.fit(X, y).predict(T)
plt.subplot(2, 1, i + 1)
plt.scatter(X, y, color='darkorange', label='data')
plt.plot(T, y_, color='navy', label='prediction')
plt.axis('tight')
plt.legend()
plt.title("KNeighborsRegressor (k = %i, weights = '%s')" % (n_neighbors,
weights))
plt.tight_layout()
plt.show()
| sklearn/sklearn learning/demonstration/auto_examples_jupyter/neighbors/plot_regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # 2014 NFIRS FireCARES data statistics and loading
#
# ## Overview
#
# The purpose of this collection of scripts is to load the NFIRS yearly data into a dump of the existing Firecares Database. The individual 18 incident types and ancillary tables (fireincident, hazchem, hazmat, etc) are loaded into temporary tables and then appended to the master "fireincident", etc tables. The bulk of this script has to do w/ matching addresses in geocoding results to the new incidents in the "incidentaddress_2014" and appending that to the "incidentaddress" table. The incident_address_2014_aa/ab/ac/ad/ae tables contain the geocoding information from shapefiles and will be used to augment the incidentaddress_2014 table's records with geometries.
#
# ## Assumptions
#
# * You have jupyter>=1.0.0 and psycopg2 and dependencies installed
# * See append_nfirs_yearly_data.sh for more information / prereqs and initial data loading
# * Database has been restored as "nfirs_2014"
# * All of the `Incident_address_2014_a*.shp` data has been loaded into the database as "incident_address_2014_aa/ab/ac/ad/ae" tables
# * Dates have been converted to a Postgres-parseable date (MM/DD/YYYY) in:
# * `fireincident.txt`
# * `incidentaddress.txt`
# * `arsonjuvsub.txt`
# * `basicincident.txt`
# * `civiliancasualty.txt`
# * `ffcasualty.txt`
# * Null (`\000`) characters have been stripped from `incidentaddress.txt` and `fdheader.txt`
# * `RISK_FACT1` codes in `arsonjuvsub.txt` have been replaced with their single-character equivalent (eg. "U", "1", etc)
# * The 18 ancillary tables have been loaded as [FILENAME w/o extension]_2014 and include data from:
# * `fireincident.txt` => maps to `fireincident_2014`
# * `hazchem.txt`
# * `hazmat.txt`
# * `hazmatequipinvolved.txt`
# * `hazmobprop.txt`
# * `incidentaddress.txt`
# * `wildlands.txt`
# * `arson.txt`
# * `arsonagencyreferal.txt`
# * `arsonjuvsub.txt`
# * `basicaid.txt`
# * `basicincident.txt`
# * `civiliancasualty.txt`
# * `codelookup.txt`
# * `ems.txt`
# * `fdheader.txt`
# * `ffcasualty.txt`
# * `ffequipfail.txt`
# * An `id` serial primary key has been added to the "incidentaddress_2014" table
# * "incidentaddress" table has `source` column added
#
# The rest continues from (as of this writing) line 101 of `append_nfirs_yearly_data.sh`
# +
import pandas as pd
import psycopg2
conn = psycopg2.connect("dbname=nfirs_2014")
# +
# Counts of the geolocated incident addresses
pd.read_sql_query("""
select 'AA' as table, count(1) from incident_address_2014_aa union
select 'AB' as table, count(1) from incident_address_2014_ab union -- ab's .dbf was corrupted at record 494357
select 'AC' as table, count(1) from incident_address_2014_ac union
select 'AD' as table, count(1) from incident_address_2014_ad union
select 'AE' as table, count(1) from incident_address_2014_ae""", conn)
# +
# Determine consistency between aa/ab/ac/ad/ae tables
def same_cols():
aa = pd.read_sql_query("select column_name, data_type from information_schema.columns where table_name='incident_address_2014_aa'", conn)
ab = pd.read_sql_query("select column_name, data_type from information_schema.columns where table_name='incident_address_2014_ab'", conn)
ac = pd.read_sql_query("select column_name, data_type from information_schema.columns where table_name='incident_address_2014_ac'", conn)
ad = pd.read_sql_query("select column_name, data_type from information_schema.columns where table_name='incident_address_2014_ad'", conn)
ae = pd.read_sql_query("select column_name, data_type from information_schema.columns where table_name='incident_address_2014_ae'", conn)
dest = pd.read_sql_query("select column_name, data_type from information_schema.columns where table_name='incidentaddress_2014'", conn)
final = pd.read_sql_query("select column_name, data_type from information_schema.columns where table_name='incidentaddress'", conn)
aa_cols = set([(x.items()[1][1], x.items()[0][1]) for x in aa.to_dict('records')])
ab_cols = set([(x.items()[1][1], x.items()[0][1]) for x in ab.to_dict('records')])
ac_cols = set([(x.items()[1][1], x.items()[0][1]) for x in ac.to_dict('records')])
ad_cols = set([(x.items()[1][1], x.items()[0][1]) for x in ad.to_dict('records')])
ae_cols = set([(x.items()[1][1], x.items()[0][1]) for x in ae.to_dict('records')])
dest_cols = set([(x.items()[1][1], x.items()[0][1]) for x in dest.to_dict('records')])
final_cols = set([(x.items()[1][1], x.items()[0][1]) for x in final.to_dict('records')])
union = aa_cols | ab_cols | ac_cols | ad_cols | ae_cols
print 'AA col diffs: '
print union - aa_cols
print 'AB col diffs: '
print union - ab_cols
print 'AC col diffs: '
print union - ac_cols
print 'AD col diffs: '
print union - ad_cols
print 'AE col diffs: '
print union - ae_cols
dest_union = dest_cols | final_cols
print dest_union - dest_cols
print 'Dest (incidentaddress_2014) col diffs: ' # Should only have "id" as a differing column
print dest_union - final_cols
same_cols()
# aa was numeric before running this
# -
# Change num_mile in aa/ac/ad to be a string vs number
for table in ['aa', 'ac', 'ad']:
with conn.cursor() as cursor:
cursor.execute("alter table incident_address_2014_%s alter column num_mile type character varying(8);" % table)
# Per 2013 data, the zip5 was unreliable
pd.read_sql_query("""
select 'AA' as table, count(1) from incident_address_2014_aa where zip5 = 0 union
select 'AB' as table, count(1) from incident_address_2014_ab where zip5 = 0 union
select 'AC' as table, count(1) from incident_address_2014_ac where zip5 = 0 union
select 'AD' as table, count(1) from incident_address_2014_ad where zip5 = 0 union
select 'AE' as table, count(1) from incident_address_2014_ae where zip5 = 0""", conn)
# and it looks to be the same in 2014's data, lots of zeros
# "Postal" might be a better option
pd.read_sql_query("""
select 'AA' as table, count(1) from incident_address_2014_aa where postal is null or length(postal) != 5 union
select 'AB' as table, count(1) from incident_address_2014_ab where postal is null or length(postal) != 5 union
select 'AC' as table, count(1) from incident_address_2014_ac where postal is null or length(postal) != 5 union
select 'AD' as table, count(1) from incident_address_2014_ad where postal is null or length(postal) != 5 union
select 'AE' as table, count(1) from incident_address_2014_ae where postal is null or length(postal) != 5""", conn)
# About the same...overall
# +
# Ok, so use "postal" for zip5 IF zip5 is zero
with conn.cursor() as cursor:
cursor.execute("""update incident_address_2014_aa set zip5 = to_number(postal, '99999') where zip5 = 0;""")
print cursor.rowcount
cursor.execute("""update incident_address_2014_ab set zip5 = to_number(postal, '99999') where zip5 = 0;""")
print cursor.rowcount
cursor.execute("""update incident_address_2014_ac set zip5 = to_number(postal, '99999') where zip5 = 0;""")
print cursor.rowcount
cursor.execute("""update incident_address_2014_ad set zip5 = to_number(postal, '99999') where zip5 = 0;""")
print cursor.rowcount
cursor.execute("""update incident_address_2014_ae set zip5 = to_number(postal, '99999') where zip5 = 0;""")
print cursor.rowcount
# Already ran aa/ab, counts for ab was 228, missed aa
# -
# Even-out some of the nullable fields
for table in ['aa', 'ab', 'ac', 'ad', 'ae']:
with conn.cursor() as cursor:
print 'Updating %s' % table
cursor.execute("update incident_address_2014_%s set street_pre = '' where street_pre is null;" % table)
print cursor.rowcount
cursor.execute("update incident_address_2014_%s set streettype = '' where streettype is null;" % table)
print cursor.rowcount
cursor.execute("update incident_address_2014_%s set streetsuf = '' where streetsuf is null;" % table)
print cursor.rowcount
cursor.execute("update incident_address_2014_%s set streetname = '' where streetname is null;" % table)
print cursor.rowcount
cursor.execute("update incident_address_2014_%s set num_mile = '' where num_mile = '0';" % table)
print cursor.rowcount
cursor.execute("update incident_address_2014_%s set apt_no = '' where apt_no = '0';" % table)
print cursor.rowcount
cursor.execute("update incident_address_2014_%s set city_1 = '' where city_1 is null;" % table)
print cursor.rowcount
cursor.execute("update incident_address_2014_%s set state_id = '' where state_id is null;" % table)
print cursor.rowcount
cursor.execute("update incident_address_2014_%s set streetname = upper(streetname);" % table)
print cursor.rowcount
cursor.execute("update incident_address_2014_%s set streettype = upper(streettype);" % table)
print cursor.rowcount
cursor.execute("update incident_address_2014_%s set streetsuf = upper(streetsuf);" % table)
print cursor.rowcount
cursor.execute("update incident_address_2014_%s set street_pre = upper(street_pre);" % table)
print cursor.rowcount
conn.commit()
# +
# Looks like we're good on the loc_type being the same set across all of the tables
import numpy as np
aa = pd.read_sql_query("select distinct loc_type from incident_address_2014_aa;", conn)
ab = pd.read_sql_query("select distinct loc_type from incident_address_2014_ab;", conn)
ac = pd.read_sql_query("select distinct loc_type from incident_address_2014_ac;", conn)
ad = pd.read_sql_query("select distinct loc_type from incident_address_2014_ad;", conn)
ae = pd.read_sql_query("select distinct loc_type from incident_address_2014_ae;", conn)
(np.sort(aa.transpose()) == np.sort(ab.transpose())).all() and \
(np.sort(aa.transpose()) == np.sort(ac.transpose())).all() and \
(np.sort(aa.transpose()) == np.sort(ad.transpose())).all() and \
(np.sort(aa.transpose()) == np.sort(ae.transpose())).all()
# +
# Ugh, num_mile's type is inconsistent across the these tables
tables = ['aa', 'ab', 'ac', 'ad', 'ae']
for table in tables:
with conn.cursor() as cursor:
print "Updating %s" % table
cursor.execute("alter table incident_address_2014_%s alter column num_mile type character varying(254);" % table)
conn.commit()
# -
# So we know the exact source of the geocoded address after merging these tables together
for table in tables:
with conn.cursor() as cursor:
print "Updating %s" % table
cursor.execute("alter table incident_address_2014_%s add column source character varying (8) default '%s';" % (table, table))
conn.commit()
same_cols()
# +
# Make aa/ab/ac/ad/ae consistent
with conn.cursor() as cursor:
cursor.execute("alter table incident_address_2014_aa drop column x, drop column y, drop column street_add, drop column inc_date, drop column zip4")
cursor.execute("alter table incident_address_2014_ab drop column full_stree, drop column inc_date, drop column zip4")
cursor.execute("alter table incident_address_2014_ac drop column full_stree, drop column inc_date")
cursor.execute("alter table incident_address_2014_ad drop column full_stree, drop column inc_date")
cursor.execute("alter table incident_address_2014_ae drop column full_stree, drop column inc_date, drop column zip4")
conn.commit()
# -
same_cols()
# +
# Create and stage consolidated table w/ indexes
with conn.cursor() as cursor:
cursor.execute("create table address_to_geo_2014 as table incident_address_2014_aa with no data;")
cursor.execute("create index on address_to_geo_2014 (num_mile, upper(apt_no), upper(city_1), zip5, upper(street_pre), loc_type, upper(streetname), upper(streettype), upper(streetsuf), upper(state_id));")
cursor.execute("create index on address_to_geo_2014 (state_id);")
cursor.execute("create index on address_to_geo_2014 (source);")
cursor.execute("create index on address_to_geo_2014 (ogc_fid)")
conn.commit()
# +
# Dump and load, specifying ALL of the cols in the source and dest is tedious, this is a workaround
for table in tables:
with conn.cursor() as cursor:
print "Dumping %s" % table
cursor.copy_to(open("/tmp/%s.dump" % table, "w"), "incident_address_2014_%s" % table)
for table in tables:
with conn.cursor() as cursor:
print "Loading %s" % table
cursor.copy_from(open("/tmp/%s.dump" % table), "address_to_geo_2014")
conn.commit()
# +
# Redundant, but a habit
tot = 0
for table in tables:
with conn.cursor() as cursor:
cursor.execute("select count(1) from incident_address_2014_%s" % table)
tot = tot + cursor.fetchone()[0]
with conn.cursor() as cursor:
cursor.execute("select count(1) from address_to_geo_2014")
assert tot == cursor.fetchone()[0]
# -
# Need to change the zip5 datatype to match incidentaddress_2014
with conn.cursor() as cursor:
cursor.execute("alter table address_to_geo_2014 alter column zip5 type character varying(5);")
cursor.execute("update address_to_geo_2014 set zip5 = lpad(zip5, 5, '0')")
print cursor.rowcount
conn.commit()
# Need to change the loc_type datatype to match incidentaddress_2014
with conn.cursor() as cursor:
cursor.execute("alter table address_to_geo_2014 alter column loc_type type character varying(1);")
conn.commit()
# Ugh, forgot to update the apt_no
with conn.cursor() as cursor:
cursor.execute("update address_to_geo_2014 set apt_no = '' where apt_no is null")
print cursor.rowcount
conn.commit()
# Just so we can track the source shapefile for the geocoding in case of the potential issues
with conn.cursor() as cursor:
cursor.execute("alter table incidentaddress_2014 add column source character varying (8);")
conn.commit()
# Lets see how many incidents there are in each state for 2014
from pretty import pprint
with conn.cursor() as cursor:
cursor.execute("select state_id, count(state_id) from incidentaddress_2014 group by state_id order by state_id;")
pprint(cursor.fetchall())
conn.commit()
# +
# Create some indexes so this query doesn't take years to complete
with conn.cursor() as cursor:
cursor.execute("""
create index on incidentaddress_2014 (num_mile, upper(apt_no), upper(city), zip5, upper(street_pre),
loc_type, upper(streetname), upper(streettype), upper(streetsuf), upper(state_id));""")
cursor.execute("""
create index on address_to_geo_2014 (num_mile, upper(apt_no), upper(city_1), zip5, upper(street_pre), loc_type, upper(streetname), upper(streettype), upper(streetsuf), upper(state_id));
create index on address_to_geo_2014 (state_id);
create index on address_to_geo_2014 (source);""")
conn.commit()
# +
# Execute the matching of the geocoded addresses to incident address for 2014, updating the geometry associated w/ each incident
with conn.cursor() as cursor:
cursor.execute("""
update incidentaddress_2014 as ia set geom = res.wkb_geometry, source = res.source
from (
select ia.id, aa.wkb_geometry, aa.source from address_to_geo_2014 aa inner join incidentaddress_2014 ia on (
aa.num_mile = ia.num_mile and
upper(aa.apt_no) = upper(ia.apt_no) and
upper(aa.city_1) = upper(ia.city) and
aa.zip5 = ia.zip5 and
upper(aa.street_pre) = upper(ia.street_pre) and
aa.loc_type = ia.loc_type and
upper(aa.streetname) = upper(ia.streetname) and
upper(aa.streettype) = upper(ia.streettype) and
upper(aa.streetsuf) = upper(ia.streetsuf) and
upper(aa.state_id) = upper(ia.state_id)
) where aa.num_mile != '' and score != 0) as res
where ia.id = res.id
""")
print cursor.rowcount
conn.commit()
# +
# Now lets see how many incidents there are in each state for 2014 that are geocoded vs not geocoded
from decimal import *
from pretty import pprint
with conn.cursor() as cursor:
cursor.execute("""
select (100.0 * sum(case when geom is null then 0 else 1 end) / count(1)) as percent_with_geom,
sum(case when geom is null then 0 else 1 end) as with_geom, state_id,
count(state_id) as total_incidents
from incidentaddress_2014 group by state_id order by percent_with_geom desc;
""")
print '% | Has geom | State | Total rows'
for row in cursor.fetchall():
print '{} | {} | {} | {}'.format(row[0], row[1], row[2], row[3])
cursor.execute("""
select sum(case when geom is null then 0 else 1 end) as matches,
(100.0 * sum(case when geom is null then 0 else 1 end) / count(1)) as percent,
count(1) as total
from incidentaddress_2014 order by percent desc;""")
row = cursor.fetchone()
print '# geocode matches: {}, Percent geocoded: {}, Total {}'.format(row[0], row[1], row[2])
conn.commit()
# +
# Just for sanity sake, make sure that the column types are the same between the 2014 and base data
addl_tables = ['arson', 'arsonagencyreferal', 'arsonjuvsub', 'basicaid', 'basicincident', 'civiliancasualty', 'codelookup', 'ems', 'fdheader', 'ffcasualty', 'ffequipfail', 'fireincident', 'hazchem', 'hazmat', 'hazmatequipinvolved', 'hazmobprop', 'incidentaddress', 'wildlands']
for table in addl_tables:
t = pd.read_sql_query("select column_name, data_type from information_schema.columns where table_name='%s'" % table, conn)
t_2014 = pd.read_sql_query("select column_name, data_type from information_schema.columns where table_name='%s_2014'" % table, conn)
t_cols = set([(x.items()[1][1], x.items()[0][1]) for x in arson.to_dict('records')])
t2014_cols = set([(x.items()[1][1], x.items()[0][1]) for x in arson2014.to_dict('records')])
print 'Table: {}, col diffs: {}'.format(table, t2014_cols - t_cols)
# +
# get some pre counts
with conn.cursor() as cursor:
for t in addl_tables:
cursor.execute("select '{table}', count(1) from {table};".format(table=t))
print cursor.fetchone()
conn.commit()
# +
# Update the fdheader table first
with conn.cursor() as cursor:
cursor.execute("select count(1) from fdheader_2014;")
print 'fdheader_2014 count: %s' % cursor.fetchone()[0]
cursor.execute("select count(1) from fdheader;")
pre = cursor.fetchone()[0]
cursor.execute("""INSERT INTO fdheader(
state, fdid, fd_name, fd_str_no, fd_str_pre, fd_street, fd_str_typ,
fd_str_suf, fd_city, fd_zip, fd_phone, fd_fax, fd_email, fd_fip_cty,
no_station, no_pd_ff, no_vol_ff, no_vol_pdc)
(SELECT distinct on (state, fdid) state, fdid, fd_name, fd_str_no, fd_str_pre, fd_street, fd_str_typ,
fd_str_suf, fd_city, fd_zip, fd_phone, fd_fax, fd_email, fd_fip_cty,
no_station, no_pd_ff, no_vol_ff, no_vol_pdc
FROM fdheader_2014 where (state, fdid) not in (select state, fdid from fdheader));""")
inserted = cursor.rowcount
cursor.execute("select count(1) from fdheader;")
post = cursor.fetchone()[0]
print 'FDHeader Pre: {}, Post: {}, Insert count: {}, Post - pre: {}'.format(pre, post, inserted, post - pre)
conn.commit()
# -
# Update the "codeloop" (appears to be a static table) w/ the 2014 data (truncate and load)
with conn.cursor() as cursor:
cursor.execute("delete from codelookup;")
print 'Deleted: %s' % cursor.rowcount
cursor.execute("insert into codelookup select * from codelookup_2014;")
print 'Inserted: %s' % cursor.rowcount
conn.commit()
# Insert the 2014 data into the foundation data
def load_2014(table, commit=True):
with conn.cursor() as cursor:
cursor.execute("select count(1) from %s;" % table)
pre = cursor.fetchone()[0]
cursor.execute("insert into %s select * from %s_2014;" % (table, table))
inserted = cursor.rowcount
cursor.execute("select count(1) from %s;" % table)
post = cursor.fetchone()[0]
print 'Table: {}, Pre: {}, Post: {}, Insert count: {}, Post - pre: {}'.format(table, pre, post, inserted, post - pre)
if commit:
conn.commit()
print 'COMMITTED'
else:
conn.rollback()
print 'ROLLBACK'
# Load EVERYTHING except for basicincident, fireincident, incidentaddress
for table in ['arson', 'arsonagencyreferal', 'arsonjuvsub', 'basicaid', 'civiliancasualty', 'ems', 'ffcasualty',
'ffequipfail', 'hazchem', 'hazmat', 'hazmatequipinvolved', 'hazmobprop', 'wildlands']:
load_2014(table)
# +
# There must have been an error with the MO 06101 entry in "fdheader_2014", since there are 101 references to it in "basicincident_2014"
with conn.cursor() as cursor:
cursor.execute("""select count(state), state, fdid from basicincident_2014 bi
where (state, fdid) not in (select state, fdid from fdheader)
and (state, fdid) not in (select state, fdid from fdheader_2014)
group by state, fdid""")
print cursor.fetchall()
cursor.execute("select count(1) from basicincident where state = 'MO' and fdid = '6101'")
print cursor.fetchall()
cursor.execute("select count(1) from basicincident_2014 where state = 'MO' and fdid = '6101'")
print cursor.fetchall()
# -
# Going to do a one-off update of (MO,6101) => (MO,06101) since that's the only offender
with conn.cursor() as cursor:
cursor.execute("update basicincident_2014 set fdid = '6101' where state = 'MO' and fdid = '06101'")
print cursor.rowcount
conn.commit()
with conn.cursor() as cursor:
cursor.execute("""select 'fireincident_2014', count(1) from fireincident_2014
union select 'basicincident_2014', count(1) from basicincident_2014
union select 'incidentaddress_2014', count(1) from incidentaddress_2014""")
print cursor.fetchall()
load_2014('basicincident', commit=True)
# +
# Looks like fireincident_2014 has the same issue w/ that MO 6101 fire department
with conn.cursor() as cursor:
cursor.execute("""select count(state), state, fdid from fireincident_2014 bi
where (state, fdid) not in (select state, fdid from fdheader)
and (state, fdid) not in (select state, fdid from fdheader_2014)
group by state, fdid""")
print cursor.fetchall()
cursor.execute("select count(1) from fireincident where state = 'MO' and fdid = '6101'")
print cursor.fetchall()
cursor.execute("select count(1) from fireincident_2014 where state = 'MO' and fdid = '6101'")
print cursor.fetchall()
# -
# Going to do a one-off update of (MO,6101) => (MO,06101) since that's the only offender
with conn.cursor() as cursor:
cursor.execute("update fireincident_2014 set fdid = '6101' where state = 'MO' and fdid = '06101'")
print cursor.rowcount
conn.commit()
load_2014('fireincident', commit=True)
# Loading all of the 2014 incident addresses into the master "incidentaddress" table
with conn.cursor() as cursor:
cursor.execute("select count(1) from incidentaddress_2014;")
print 'incidentaddress_2014 count: %s' % cursor.fetchone()[0]
cursor.execute("select count(1) from incidentaddress;")
pre = cursor.fetchone()[0]
cursor.execute("""INSERT INTO incidentaddress(
state, fdid, inc_date, inc_no, exp_no, loc_type, num_mile, street_pre,
streetname, streettype, streetsuf, apt_no, city, state_id, zip5,
zip4, x_street, addid, addid_try, geom, bkgpidfp00, bkgpidfp10)
(SELECT state, fdid, inc_date, inc_no, exp_no, loc_type, num_mile, street_pre,
streetname, streettype, streetsuf, apt_no, city, state_id, zip5,
zip4, x_street, addid, addid_try, geom, bkgpidfp00, bkgpidfp10
FROM incidentaddress_2014);""")
inserted = cursor.rowcount
cursor.execute("select count(1) from incidentaddress;")
post = cursor.fetchone()[0]
print 'Incident address Pre: {}, Post: {}, Insert count: {}, Post - pre: {}'.format(pre, post, inserted, post - pre)
conn.commit()
# Start moving things over to a separate schema
with conn.cursor() as cursor:
cursor.execute("create schema if not exists geocoding_2014;")
conn.commit()
with conn.cursor() as cursor:
cursor.execute("alter table address_to_geo_2014 set schema geocoding_2014;")
conn.commit()
tables = ["incident_address_2014_aa", "incident_address_2014_ab", "incident_address_2014_ac", "incident_address_2014_ad",
"incident_address_2014_ae"]
with conn.cursor() as cursor:
for table in tables:
cursor.execute("alter table %s set schema geocoding_2014;" % table)
conn.commit()
# +
# We should be able to drop all of the 2014 tables now
tables_2014 = ['arson_2014', 'arsonagencyreferal_2014', 'arsonjuvsub_2014', 'basicaid_2014', 'basicincident_2014',
'civiliancasualty_2014', 'codelookup_2014', 'ems_2014', 'fdheader_2014', 'ffcasualty_2014',
'ffequipfail_2014', 'fireincident_2014', 'hazchem_2014', 'hazmat_2014', 'hazmatequipinvolved_2014',
'hazmobprop_2014', 'incidentaddress_2014', 'wildlands_2014']
with conn.cursor() as cursor:
for table in addl_tables_2014:
cursor.execute("drop table %s;" % table)
conn.commit()
# -
# [x] - arson_2014 -> arson
# [x] - arsonagencyreferal_2014 -> arsonagencyreferal
# [x] - arsonjuvsub_2014 -> arsonjuvsub
# [x] - basicaid_2014 -> basicaid
# [x] - basicincident_2014 -> basicincident
# [x] - civiliancasualty_2014 -> civiliancasualty
# [x] - codelookup_2014 -> codelookup
# [x] - ems_2014 -> ems
# [x] - fdheader_2014 -> fdheader
# [x] - ffcasualty_2014 -> ffcasualty
# [x] - ffequipfail_2014 -> ffequipfail
# [x] - fireincident_2014 -> fireincident
# [x] - hazchem_2014 -> hazchem
# [x] - hazmat_2014 - > hazmat
# [x] - hazmatequipinvolved_2014 -> hazmatequipinvolved
# [x] - hazmobprop_2014 -> hazmobprop
# [x] - incidentaddress_2014 -> incidentaddress
# [x] - wildlands_2014 -> wildlands
# ### Dumping the db / compressing
#
# ```bash
# pg_dump -O -x nfirs_2014 > nfirs_2014.sql
# # mv nfirs_2014.sql nfirs_2014_addresses_matched.sql
# gzip nfirs_2014_addresses_matched.sql
# ```
| sources/nfirs/scripts/NFIRS 2014 load.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:notebook] *
# language: python
# name: conda-env-notebook-py
# ---
# ## Basic example of how to bin along-track ICESat-2 sea ice data, as requeated during the hackweek.
#
# * Notebook author: <NAME>
# * Description: Notebook showing (one way) of binning along-track data to the NSIDC grid
# * Input requirements: Demo ATL10 data file
# * Date: July 2020
#
# **Please note that this notebook will not run in Binder without first uploading the required input data files.**
import warnings
warnings.filterwarnings('ignore')
import numpy as np
import pandas as pd
import datetime as dt
import matplotlib.pyplot as plt
import matplotlib
matplotlib.use('Agg')
import cartopy.crs as ccrs
import h5py
import scipy
from astropy.time import Time
import utils as ut
import readers as rd
import pyproj
import os
#Magic function to enable interactive plotting in Jupyter notebook
# %matplotlib inline
# +
data_loc='./data/' # update to where you downloaded the data
for file in os.listdir(data_loc):
if file.startswith('ATL10-01_20181115003141_07240101') and file.endswith('.h5'):
fname = file
fileT= data_loc+fname
# -
beamStr='gt1r'
dF10 = rd.getATL10data(fileT, beam=beamStr)
dF10.head()
# +
def create_grid_nsidc(epsg_string='3411', nx=304, ny=448, leftx=-3837500, dxRes=25000, uppery=5837500, dyRes=25000):
""" Use pyproj to generate the NSIDC North Polar Stereographic grid covering the given domain (defined by the projection and the corner lat/lons)"""
crs = pyproj.CRS.from_string("epsg:"+epsg_string)
p=pyproj.Proj(crs)
print(dxRes, dyRes)
x=leftx+dxRes*np.indices((ny,nx),np.float32)[1]
y=uppery-dxRes*np.indices((ny,nx),np.float32)[0]
lons, lats = p(x, y, inverse=True)
return x, y, lats, lons, p
def create_grid_og(epsg_string='3413', dxRes=25000., lllat=36, llon=-90, urlat=36, urlon=90):
""" Use pyproj to generate a grid covering the given domain (defined by the projection and the corner lat/lons)"""
crs = pyproj.CRS.from_string("epsg:"+epsg_string)
p=pyproj.Proj(crs)
llcrn=p(llon, lllat)
urcrn=p(urlon, urlat)
print(llcrn)
print(urcrn)
nx = int((urcrn[0]-llcrn[0])/dxRes)+1
ny = int((urcrn[1]-llcrn[1])/dxRes)+1
print(nx, ny)
x = llcrn[0]+dxRes*np.indices((ny,nx),np.float32)[0] # 1=column indicdes
y = llcrn[1]+dxRes*np.indices((ny,nx),np.float32)[1] # 0=row indices
lons, lats = p(x, y, inverse=True)
return x, y, lats, lons, p
# -
xptsG, yptsG, latG, lonG, proj = create_grid_nsidc()
# +
def bin_data(xpts, ypts, var, xptsG, yptsG, dx):
""" Bin data using numpy histogram2d
Adapted for the NSIDC grid which has its orgin in the top left corner.
"""
# Need to flip the arrays because the origin is in the top left but the histogram2d function needs x/y increasing.
xptsG2=np.flipud(xptsG)
yptsG2=np.flipud(yptsG)
# get bin edges by subtracting half a grid-width and adding on another bin in both directions
xbins=xptsG2[0]-(dx/2)
ybins=yptsG2[:, 0]-(dx/2)
xbins2=np.append(xbins, xbins[-1]+dx)
ybins2=np.append(ybins, ybins[-1]+dx)
print('binning..')
#print(xbins2.shape)
#print(ybins2.shape)
counts, xedges, yedges = np.histogram2d(xpts, ypts,bins=(xbins2, ybins2))
z, _, _ = np.histogram2d(xpts, ypts,bins=(xbins2, ybins2), weights=var)
varG = z / counts
# Need to re-flip the arrays then transpose because of how histogram2d works across columns then rows.
varG=np.flipud(varG.T)
counts=np.flipud(counts.T)
return varG
def bin_data_og(xpts, ypts, var, xptsG, yptsG, dx):
""" Bin data using numpy histogram 2d
You can use this one when using the og grid"""
xbins=xptsG[:,0]-(dx/2)
ybins=yptsG[0, :]-(dx/2)
xbins=np.append(xbins, xbins[-1]+dx)
ybins=np.append(ybins, ybins[-1]+dx)
counts, xedges, yedges = np.histogram2d(xpts, ypts,bins=(xbins, ybins))
z, _, _ = np.histogram2d(xpts, ypts,bins=(xbins, ybins), weights=var)
varG = z / counts
return varG
# +
# Define a projection
mapProj = pyproj.Proj("+init=EPSG:3411")
x10, y10=mapProj(dF10.lon.values, dF10.lat.values)
varG=bin_data(x10, y10, dF10.freeboard.values, xptsG, yptsG, dx)
print(varG.shape)
# -
varG
# +
# Plot this gridded data
# Note that we're not using the exact same projection here but a similar built-in North Polar Stereographic projection, just for plotting.
fig=plt.figure(figsize=(5, 6))
# Use the in-built northpolarstereo to vizualize (should somehow use the actual projection)
ax = plt.axes(projection =ccrs.NorthPolarStereo(central_longitude=-45))
cs=ax.pcolormesh(lonG, latG, varG, vmin=0, vmax=0.5,transform=ccrs.PlateCarree(), zorder=2)
ax.coastlines(zorder=3)
ax.gridlines(draw_labels=True,
linewidth=0.22, color='gray', alpha=0.5, linestyle='--')
ax.set_extent([-179, 179, 50, 90], ccrs.PlateCarree())
# +
# As a sanity check let's plot the raw data too...
# +
# Plot this gridded data
fig=plt.figure(figsize=(5, 6))
# Use the in-built northpolarstereo to vizualize (should somehow use the actual projection)
ax = plt.axes(projection =ccrs.NorthPolarStereo(central_longitude=-45))
cs=ax.scatter(dF10.lon.values, dF10.lat.values, c=dF10.freeboard.values,vmin=0, vmax=0.5,transform=ccrs.PlateCarree(), zorder=2)
ax.coastlines(zorder=3)
ax.gridlines(draw_labels=True,
linewidth=0.22, color='gray', alpha=0.5, linestyle='--')
ax.set_extent([-179, 179, 50, 90], ccrs.PlateCarree())
# -
| 10.Sea_Ice_Applications/GriddingDemo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import pandas as pd
from Player.Players import Players
from Regression.Reg_Model import Reg_Model
#Creates list of players during 2015-2016 season
Player_List = Players('2015-16').players
#prepare model for subset of players
p_subset_list=[]
index=0
for key in Player_List:
p=Player_List[key]
stats = p.get_stats('GAME_DATE', 'Game_ID', 'PTS', ['PTS', 'MIN', 'FGM', 'FGA', 'FG_PCT'], 5)
if len(stats>0):
stats=stats[stats['n_games']>=5]
y=stats['PTS']
x=stats[['PTS_avg_5', 'MIN_avg_5', 'FGM_avg_5', 'FGA_avg_5', 'FG_PCT_avg_5']]
if len(y)>=5:
Model = Reg_Model()
Model.set_training(x,y)
Model.calc_model()
p.set_model('PTS', Model)
p_subset_list.append(key)
index += 1
if index>50:
break
#average MSE for players
import numpy as np
print np.mean([Player_List[a].model_list['PTS'].mse for a in p_subset_list])
# +
#Try to see how data looks like by position, season_exp
exp = [0,4,8,12]
positions = set([Player_List[x].desc['POSITION'][0] for x in Player_List])
mse_by_seg = {}
for pos in positions:
for ex in exp:
x =pd.DataFrame()
y =pd.DataFrame()
for key in Player_List:
if Player_List[key].desc['POSITION'][0] == pos:
if Player_List[key].desc['SEASON_EXP'][0]>=ex and Player_List[key].desc['SEASON_EXP'][0]<(ex+4):
stats = Player_List[key].get_stats('GAME_DATE', 'Game_ID', 'PTS', ['PTS', 'MIN', 'FGM', 'FGA', 'FG_PCT'], 5)
if len(stats>0):
stats=stats[stats['n_games']>=5]
y = y.append(stats[['PTS', 'PTS_avg_5']], ignore_index=True).reset_index(drop=True)
x = x.append(stats[['PTS_avg_5', 'MIN_avg_5', 'FGM_avg_5', 'FGA_avg_5', 'FG_PCT_avg_5']], ignore_index=True).reset_index(drop=True)
y=y['PTS']
Model = Reg_Model()
Model.set_training(x,y)
Model.calc_model()
segment_name = pos + '-' + str(ex)
mse_by_seg[segment_name] = Model.mse
# -
mse_by_seg
# +
#Repeat with 10 games
exp = [0,4,8,12]
positions = set([Player_List[x].desc['POSITION'][0] for x in Player_List])
mse_by_seg = {}
for pos in positions:
for ex in exp:
x =pd.DataFrame()
y =pd.DataFrame()
for key in Player_List:
if Player_List[key].desc['POSITION'][0] == pos:
if Player_List[key].desc['SEASON_EXP'][0]>=ex and Player_List[key].desc['SEASON_EXP'][0]<(ex+4):
stats = Player_List[key].get_stats('GAME_DATE', 'Game_ID', 'PTS', ['PTS', 'MIN', 'FGM', 'FGA', 'FG_PCT'], 10)
if len(stats>0):
stats=stats[stats['n_games']>=5]
y = y.append(stats[['PTS', 'PTS_avg_10']], ignore_index=True).reset_index(drop=True)
x = x.append(stats[['PTS_avg_10', 'MIN_avg_10', 'FGM_avg_10', 'FGA_avg_10', 'FG_PCT_avg_10']], ignore_index=True).reset_index(drop=True)
y=y['PTS']
Model = Reg_Model()
Model.set_training(x,y)
Model.calc_model()
segment_name = pos + '-' + str(ex)
mse_by_seg[segment_name] = Model.mse
# -
mse_by_seg
#Now i want to see how i can add other variables to the model. probably want a model that takes into account: opposing team, current team
from Schedule.Schedule import Schedule
from Schedule.Stats import Stats
sched_2015 = Schedule(b_dt = '10/1/2015')
sched_2015.add_four_factors()
# Create last n statistics
games = sched_2015.get_games()
stats = Stats(games, 'avg', 'GAME_DATE', 'Home Team', 'Away Team', 'Pts_diff', ['Game_ID'])
stats_10 = stats.get_lastn_stats(10)
# +
print len(stats_10)
stats_10 = stats_10[stats_10['H_10_games']==10]
print len(stats_10)
stats_10 = stats_10[stats_10['A_10_games']==10]
print len(stats_10)
# +
#Create model based on team data
exp = [0,4,8,12]
positions = set([Player_List[x].desc['POSITION'][0] for x in Player_List])
mse_by_seg = {}
for pos in positions:
for ex in exp:
x =pd.DataFrame()
y =pd.DataFrame()
for key in Player_List:
if Player_List[key].desc['POSITION'][0] == pos:
if Player_List[key].desc['SEASON_EXP'][0]>=ex and Player_List[key].desc['SEASON_EXP'][0]<(ex+4):
stats = Player_List[key].get_stats('GAME_DATE', 'Game_ID', 'PTS', ['PTS', 'MIN', 'FGM', 'FGA', 'FG_PCT'], 10)
if len(stats>0):
data = Player_List[key].game_logs
for index, game in data.iterrows():
splits = game['MATCHUP'].split(' ')
if splits[1] == '@':
data.set_value(index, 'Home', 0)
else:
data.set_value(index, 'Home', 1)
data = data[['Game_ID', 'Home']]
stats = pd.merge(stats, data, on='Game_ID')
stats=stats[stats['n_games']>=5]
col_list = {}
for col in h_games.columns.values:
if col[0:2]=='H_':
new_col = 'Y_' + col[2:]
elif col[0:2]=='A_':
new_col = 'M_' + col[2:]
else:
new_col=col
col_list[col]=new_col
h_games.rename(columns=col_list,inplace=True)
col_list = {}
for col in a_games.columns.values:
if col[0:2]=='H_':
new_col = 'M_' + col[2:]
elif col[0:2]=='A_':
new_col = 'Y_' + col[2:]
else:
new_col=col
col_list[col]=new_col
a_games.rename(columns=col_list,inplace=True)
stats_all = h_games.append(a_games, ignore_index=True).reset_index(drop=True)
stats_all = stats_all[stats_all['n_games']>=5]
y=stats_all[['PTS', 'PTS_avg_10']]
x=stats_all[['PTS_avg_10', 'MIN_avg_10', 'FGM_avg_10', 'FGA_avg_10', 'FG_PCT_avg_10', 'M_BTB',
'M_FF_EFG_10', 'M_FF_FTFGA_10', 'M_FF_ORB_10',
'M_FF_TOV_10', 'M_O_FF_EFG_10',
'M_O_FF_FTFGA_10', 'M_O_FF_ORB_10',
'M_O_FF_TOV_10', 'M_O_PTS_10','M_O_WL_10',
'M_PTS_10', 'M_WL_10',
'Y_FF_EFG_10', 'Y_FF_FTFGA_10', 'Y_FF_ORB_10',
'Y_FF_TOV_10', 'Y_O_FF_EFG_10',
'Y_O_FF_FTFGA_10', 'Y_O_FF_ORB_10',
'Y_O_FF_TOV_10', 'Y_O_PTS_10', 'Y_O_WL_10',
'Y_PTS_10', 'Y_WL_10']]
y=y['PTS']
Model = Reg_Model()
Model.set_training(x,y)
Model.calc_model()
segment_name = pos + '-' + str(ex)
mse_by_seg[segment_name] = Model.mse
#Gets Player stats data and adds home indicator
# -
mse_by_seg
stats_10.columns
#try to see what variables i have on tap
# +
#join player stats and game stats for 1 player
from Player.Player import Player
lebron = Player(f_name='Lebron', l_name='James')
data = lebron.game_logs
for index, game in data.iterrows():
splits = game['MATCHUP'].split(' ')
if splits[1] == '@':
data.set_value(index, 'Home', 0)
else:
data.set_value(index, 'Home', 1)
data = data[['Game_ID', 'Home']]
stats = lebron.get_stats('GAME_DATE', 'Game_ID', 'PTS', ['PTS', 'MIN', 'FGM', 'FGA', 'FG_PCT'], 10)
stats_v2 = pd.merge(stats, data, on='Game_ID')
print len(stats_v2)
stats_v3 = stats_v2[stats_v2['n_games']>=10]
print len(stats_v3)
# +
h_games = stats_v3[stats_v3['Home']==1]
print len(h_games)
a_games = stats_v3[stats_v3['Home']==0]
print len(a_games)
h_games = pd.merge(h_games, stats_10, on='Game_ID')
print len(h_games)
a_games = pd.merge(a_games, stats_10, on='Game_ID')
print len(a_games)
# +
print len(h_games.columns)
col_list = {}
for col in h_games.columns.values:
if col[0:2]=='H_':
new_col = 'Y_' + col[2:]
elif col[0:2]=='A_':
new_col = 'M_' + col[2:]
else:
new_col=col
col_list[col]=new_col
h_games.rename(columns=col_list,inplace=True)
print len(h_games.columns)
print len(a_games.columns)
col_list = {}
for col in a_games.columns.values:
if col[0:2]=='H_':
new_col = 'M_' + col[2:]
elif col[0:2]=='A_':
new_col = 'Y_' + col[2:]
else:
new_col=col
col_list[col]=new_col
a_games.rename(columns=col_list,inplace=True)
print len(a_games.columns)
# -
stats_all = h_games.append(a_games, ignore_index=True).reset_index(drop=True)
print len(stats_all)
print len(a_games.columns)
print len(a_games.columns)
print len(stats_all.columns)
stats_all.columns
stats_all = stats_all[stats_all['n_games']>4]
y=stats_all['PTS']
x=stats_all[['PTS_avg_10', 'MIN_avg_10', 'FGM_avg_10', 'FGA_avg_10', 'FG_PCT_avg_10', 'M_BTB']]
stats_all = stats_all[stats_all['n_games']>4]
y=stats_all['PTS']
x=stats_all[['PTS_avg_10', 'MIN_avg_10', 'FGM_avg_10', 'FGA_avg_10', 'FG_PCT_avg_10', 'M_BTB',
'M_FF_EFG_10', 'M_FF_FTFGA_10', 'M_FF_ORB_10',
'M_FF_TOV_10', 'M_O_FF_EFG_10',
'M_O_FF_FTFGA_10', 'M_O_FF_ORB_10',
'M_O_FF_TOV_10', 'M_O_PTS_10','M_O_WL_10',
'M_PTS_10', 'M_WL_10',
'Y_FF_EFG_10', 'Y_FF_FTFGA_10', 'Y_FF_ORB_10',
'Y_FF_TOV_10', 'Y_O_FF_EFG_10',
'Y_O_FF_FTFGA_10', 'Y_O_FF_ORB_10',
'Y_O_FF_TOV_10', 'Y_O_PTS_10', 'Y_O_WL_10',
'Y_PTS_10', 'Y_WL_10']]
# +
for index, row in x.iterrows():
for col in x.columns.values:
print type(x.ix[index, col]), col
break
# -
x
Model = Reg_Model()
Model.set_training(x,y)
Model.calc_model()
Model.mse
Model.model_type
| .ipynb_checkpoints/Player Analysis-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
pd.options.display.max_columns = 500
df.to_csv('FE_fights', index=False)
# # Feature Egineering
# ### - Create time from last fight✅
# ### - ordinal encode Max_round ✅
# ### - HomeTown will be logitude latitude
df = pd.read_csv('FE_fights')
df.tail()
# ## Ordinal Encoding ✅
# Ordinal encoded max rounds as 3_round, 4_round, 5_round
dummies = pd.get_dummies(df.Max_round)
ex = dummies.rename(columns={3:'3_round', 4:'4_round', 5:'5_round'})
cdf = df.copy()
df = cdf.join(ex)
# +
# Destroy max_round column and BStreak because Red does not have that feature
# df = df.drop(columns=['Max_round'])
# df = df.drop(columns=['BStreak'])
# -
df.head()
# ## Create time since last fight ✅
# - group each fighter (need to group so that both corners are looked at for time line)
# - order based on date
# - use an array the size of the data set
# - replace none values to the index of that fighter
# - do this for each corner
date = pd.to_datetime(df.Date)
df.Date = date
date.iloc[0]
R_group = df.groupby('R_ID')
# len(R_group.groups)
R_group.groups
B_group = df.groupby('B_ID')
B_group.groups
129 in R_group.groups
R_i = R_group.indices[129].tolist()
B_i = B_group.indices[129].tolist()
R_i + B_i
df.iloc[R_group.indices[129]]
R_delta = np.empty(df.shape[0], dtype=pd.Timestamp)
B_delta = np.empty(df.shape[0], dtype=pd.Timestamp)
# - iterate through all fighter IDs
# - Use ID to get the Red corner and Blue corners into variables
# - Create a data frame for each fighter containing the date of the fight and whether which corner they were in and the index and
# - Iterate through the dataframe and create a time delta from last match column
# - Add that to corresponding index and corresponding array
# Create list of all IDs
IDs = pd.concat([df.R_ID, df.B_ID])
IDs = IDs.drop_duplicates()
# Create column names for empty DataFrame
columns = ['corner', 'Date', 'diff', 'index'] #, 'index']
# +
for ID in IDs:
temp_df = pd.DataFrame(columns=columns)
if ID in R_group.groups:
r_corner = ['r'] * len(R_group.groups[ID])
r_date = df.iloc[R_group.groups[ID]].Date.tolist()
r_index = R_group.indices[ID].tolist()
if ID in B_group.groups:
b_corner = ['b'] * len(B_group.groups[ID])
b_date = df.iloc[B_group.groups[ID]].Date.tolist()
b_index = B_group.indices[ID].tolist()
temp_df['corner'] = r_corner + b_corner
temp_df['Date'] = r_date + b_date
temp_df['index'] = r_index + b_index
temp_df = temp_df.sort_values(by=['Date'])
temp_df['diff'] = temp_df['Date'].diff().fillna(pd.to_timedelta(0))
# print(temp_df.head())
for _, row in temp_df.iterrows():
if row.corner == 'r':
R_delta[row['index']] = row['diff']
elif row['corner'] == 'b':
B_delta[row['index']] = row['diff']
# -
type([x.days for x in B_delta][1])
df['B_diff'] = [x.days for x in B_delta]
df['R_diff'] = [x.days for x in R_delta]
pd.Series(sorted(df.iloc[[1, 20, 50]].Date)).diff()
c = ['r'] * 10
c2 = ['b'] * 3
c3 = c + c2
# dfdf = pd.DataFrame({'corner':c3})
# dfdf
c3
df.R_diff.dtype
df.B_diff.hist();
df.R_diff.hist()
# ## Vectorize names with word to vec
# - I don't think this will work because there's already an ID
# +
from nltk.tokenize import sent_tokenize, word_tokenize
import warnings
warnings.filterwarnings(action = 'ignore')
import gensim
from gensim.models import Word2Vec
# -
names = df.head().B_Name.values
names
tokens = [word_tokenize(x) for x in names]
tokens
[Word2Vec(x) for x in tokens]
# ## Use Geopy to add longitude and latitude based off HomeTown
from geopy.geocoders import Nominatim
geolocator = Nominatim(user_agent='pwalis', timeout=3)
location = geolocator.geocode("Brazil", addressdetails=True)
one = location
import time
start = time.process_time()
[geolocator.geocode(x) for x in df.R_HomeTown.head(30)]
print(time.process_time() - start)
R_geo = []
i = 0
for x in df.R_HomeTown:
i+=1
print(i)
geo_info = geolocator.geocode(x)
R_geo.append(geo_info)
# honestly I just want a green square
# A lot of places aren't getting recognized as a location I should look into how the api expects the input
# Research how to handle different errors
# if statements based on the error, If it can't get the location info add an entry into the dict that has the hometown string
from geopy.exc import GeocoderTimedOut
i = 0
R_geo = dict()
for x in df.R_HomeTown:
i+=1
try:
location = geolocator.geocode(x).raw
R_geo.update({i:location})
except Exception as e:
print("Error: geocode failed on input %s with message %s"%(x, e))
len(R_geo)
i=0
while i<1162:
i+=1
if len(R_geo[i])<10:
print(R_geo[i])
R_geo.keys()
| Feature_engineering.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## MURA dataset pre-processing for Machine Learning
# This notebook does the following things:
#
# * **Read** the images from the dataset.
# * **Match** the data and the label.(Along with human part, patient number, study number, etc)
# * **Transfer** the images into gray (RGB channels to 1 channel).
# * **Reshape** the images to a standard size (here, $128\times 128$) and then do the **standardization** and **edge detection** (sobel operation) and **histogram equalization**.
# * The final image vector will be shaped as: $1\times(128*128)$
# * Perform **PCA**
#
# <h1 style="text-align:right">$\mathcal{ZLF}$ </h1>
# ## 1. Read the images
# +
# import packages
from __future__ import print_function
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
import os
import cv2
from skimage.transform import resize
from PIL import Image
from skimage.filters import sobel
from scipy import ndimage
# %matplotlib inline
# -
# get the data paths (and labels)
train_img_path = pd.read_csv("train_image_paths.csv", header=None)
# train_img_label = pd.read_csv("train_labeled_studies.csv", header=None)
valid_img_path = pd.read_csv("valid_image_paths.csv", header=None)
# valid_img_label = pd.read_csv("valid_labeled_studies.csv", header=None)
# show the data
train_img_path.head()
# ## 2. Add labels to the data
# +
# add labels to the data sets
redundant_prefix = 'MURA-v1.1/' ##preparing labels
len_prefix = len(redundant_prefix) # len_prefix = 10
train_img_path['Path2Img'] = train_img_path[0].apply(lambda x: x[len_prefix:])
valid_img_path['Path2Img'] = valid_img_path[0].apply(lambda x: x[len_prefix:])
def extract_label(string, d):
"""
extract labels "positive/negative" from file path
"""
try:
pre_str = os.path.splitext(string)[0]
label = pre_str.split("/")[-2].split("_")[-1]
if np.isin(label, list(d.keys())):
return(d[label])
else:
return('No corresponding key value')
except IOError:
return(np.nan)
d = {'positive':1, 'negative':0}
train_img_path['label'] = train_img_path[0].apply(lambda x: extract_label(x, d=d))
valid_img_path['label'] = valid_img_path[0].apply(lambda x: extract_label(x, d=d))
# -
train_img_path.head()
train_img_path.info()
valid_img_path.head()
valid_img_path.info()
# ## 3. Transfer to gray & Resize & Standardization & Edge detection & Histogram Equalization
# +
#From here, prepare images to 1-D arrays
img_size = (128, 128) ##size of images, can be modified
# transfer to gray: In fact, we just use cv2.imread(path, 0) function to get grayscale img instead of this function
def identify_tranfer2gray(raw_image_tensor, ratio = [0.299, 0.587, 0.114]):##ratio can be modified, but be sure the sum is 1.!
"""
Identify images' color status, if it is gray-valued, no action, otherwise transform colorful images to gray-valued images.
---------
Input:
param path: A tensor, from desired image
param ratio: A list, ratio of merging gray value image from a RGB image
---------
Output: A matrix, of size height*width from raw image
"""
if len(raw_image_tensor.shape)==3: # if the input is 3 dimensional tensor
return(np.dot(raw_image_tensor[...,:3], ratio))
else:
return(raw_image_tensor)
# Resize and standardization and edge detection:
def read_resize(path, rescale = img_size, sobel_op = False, hist = False, *args, **kwargs):
"""
Dealing gray-value images(i.e. no color)
Read images from file path, and rescale it according to provided image size
---------
Input:
param path: A string, path to image
param rescale: A tuple, new size of image
sobel_op: If true, perform Sobel Operation
hist: If true, perform Histogram Equalization
---------
Output:
row vector of size (1, new_height*new_width)
"""
raw_img_tensor = cv2.imread(path, 0) # load img as grayscale
if hist:
img_tensor = cv2.equalizeHist(raw_img_tensor)
else:
img_tensor = raw_img_tensor
if sobel_op:
img_tensor = sobel(img_tensor)
else:
img_tensor = img_tensor
# Notice that "resize" function do the standardization(divided by 255) automatically
rescale_image_tensor = resize(img_tensor, output_shape=rescale, mode = 'constant')
try:
rescale_image_vector = np.reshape(rescale_image_tensor, newshape=(1, rescale[0]*rescale[1]))
return(np.squeeze(rescale_image_vector))
except IOError:
return(np.nan)
# -
train_img_path['img_vector'] = train_img_path['Path2Img'].apply(lambda x: read_resize(x)) #images have been normalized
train_img_path.head()
valid_img_path['img_vector'] = valid_img_path['Path2Img'].apply(lambda x: read_resize(x)) #images have been normalized
valid_img_path.head()
# ## 4. Other info
# +
###Prepare other labels that may be helpful
train_img_path.drop(labels=0, axis=1,inplace=True) #drop redundant column
valid_img_path.drop(labels=0, axis=1,inplace=True) #drop redundant column
def human_part(path):
expected_output = path.split('/')[1].split("_")[1]
return(expected_output)
def patient_number(path):
expected_output = path.split('/')[2][-5:]
try:
expected_output = np.int(expected_output)
return(expected_output)
except IOError:
return(np.nan)
def study_number(path):
expected_output = path.split('/')[3].split("_")[0][-1]
try:
expected_output = np.int(expected_output)
return(expected_output)
except IOError:
return(np.nan)
train_img_path['human_part'] = train_img_path['Path2Img'].apply(lambda x: human_part(x))
train_img_path['patient_number'] = train_img_path['Path2Img'].apply(lambda x: patient_number(x))
train_img_path['study_number'] = train_img_path['Path2Img'].apply(lambda x: study_number(x))
valid_img_path['human_part'] = valid_img_path['Path2Img'].apply(lambda x: human_part(x))
valid_img_path['patient_number'] = valid_img_path['Path2Img'].apply(lambda x: patient_number(x))
valid_img_path['study_number'] = valid_img_path['Path2Img'].apply(lambda x: study_number(x))
# -
# show final data structure
train_img_path.head()
# show final data structure
valid_img_path.tail()
###Showing an image, paint it as you like(by modifying ratio!!!!)
plt.imshow(train_img_path['img_vector'][77].reshape(img_size[0], img_size[1]));
# ## 5. Obtain the whole dataset
# concat the train and valid data
data = pd.concat([train_img_path, valid_img_path], ignore_index=True)
data.info()
# split the data
elbow = data[data['human_part'] == 'ELBOW']
finger = data[data['human_part'] == 'FINGER']
forearm = data[data['human_part'] == 'FOREARM']
hand = data[data['human_part'] == 'HAND']
humerus = data[data['human_part'] == 'HUMERUS']
shoulder = data[data['human_part'] == 'SHOULDER']
wrist = data[data['human_part'] == 'WRIST']
# ## 6.1 Split and PCA for specific dataset
# +
from sklearn.decomposition import PCA
from sklearn.model_selection import train_test_split
def my_split_PCA(data, img_size, num_of_components):
# obtain the data
quene = list()
for j in range(len(data)):
for i in data['img_vector'].values[j]:
quene.append(i)
X = np.array(quene).reshape(len(data), img_size[0]*img_size[1])
y = data['label']
# split the data
X_train, X_test, y_train, y_test = train_test_split(X,
y,
test_size = 0.1,
random_state = 123,
shuffle = True,
stratify = y)
# perform PCA on train set
pca = PCA(n_components=num_of_components)
# train the PCA
pca_result_train = pca.fit_transform(X_train)
pca_feature_cols = ['pca' + str(i) for i in range(1,pca_result_train.shape[1]+1)]
pca_train_data = pd.DataFrame(pca_result_train, columns=pca_feature_cols)
pca_train_data['label'] = y_train.values
# perform the same PCA on the test set
pca_result_test = pca.transform(X_test)
pca_test_data = pd.DataFrame(pca_result_test, columns=pca_feature_cols)
pca_test_data['label'] = y_test.values
# print PCA exlained variance ratio
print('PCA exlained variance ratio:', pca.explained_variance_ratio_)
print('Total PCA exlained variance ratio:', np.sum(pca.explained_variance_ratio_))
return pca_train_data, pca_test_data
# -
# perform PCA and save the data
elbow_train, elbow_test = my_split_PCA(elbow, img_size = (128, 128), num_of_components = 0.8)
elbow_train.to_csv('elbow_train.csv')
elbow_test.to_csv('elbow_test.csv')
finger_train, finger_test = my_split_PCA(finger, img_size = (128, 128), num_of_components = 0.8)
finger_train.to_csv('finger_train.csv')
finger_test.to_csv('finger_test.csv')
forearm_train, forearm_test = my_split_PCA(forearm, img_size = (128, 128), num_of_components = 0.8)
forearm_train.to_csv('forearm_train.csv')
forearm_test.to_csv('forearm_test.csv')
hand_train, hand_test = my_split_PCA(hand, img_size = (128, 128), num_of_components = 0.8)
hand_train.to_csv('hand_train.csv')
hand_test.to_csv('hand_test.csv')
humerus_train, humerus_test = my_split_PCA(humerus, img_size = (128, 128), num_of_components = 0.8)
humerus_train.to_csv('humerus_train.csv')
humerus_test.to_csv('humerus_test.csv')
shoulder_train, shoulder_test = my_split_PCA(shoulder, img_size = (128, 128), num_of_components = 0.8)
shoulder_train.to_csv('shoulder_train.csv')
shoulder_test.to_csv('shoulder_test.csv')
wrist_train, wrist_test = my_split_PCA(wrist, img_size = (128, 128), num_of_components = 0.8)
wrist_train.to_csv('wrist_train.csv')
wrist_test.to_csv('wrist_test.csv')
# ## 6.2 Split and PCA for the whole dataset
# split the data into train and test
from sklearn.model_selection import train_test_split
def my_split(data, img_size, test_size):
# obtain the data
quene = list()
for j in range(len(data)):
for i in data['img_vector'].values[j]:
quene.append(i)
X = np.array(quene).reshape(len(data), img_size[0]*img_size[1])
y = data['label']
# split the data
X_train, X_test, y_train, y_test = train_test_split(X,
y,
test_size = test_size,
random_state = 123,
shuffle = True,
stratify = y)
return X_train, X_test, y_train, y_test
from sklearn.decomposition import PCA
def my_PCA(X_train, X_test, y_train, y_test, num_of_components):
# perform PCA on train set
pca = PCA(n_components=num_of_components)
# train the PCA
pca_result_train = pca.fit_transform(X_train)
pca_feature_cols = ['pca' + str(i) for i in range(1,pca_result_train.shape[1]+1)]
pca_train_data = pd.DataFrame(pca_result_train, columns=pca_feature_cols)
pca_train_data['label'] = y_train.values
# perform the same PCA on the test set
pca_result_test = pca.transform(X_test)
pca_test_data = pd.DataFrame(pca_result_test, columns=pca_feature_cols)
pca_test_data['label'] = y_test.values
# print PCA exlained variance ratio
print('PCA exlained variance ratio:', pca.explained_variance_ratio_)
print('Total PCA exlained variance ratio:', np.sum(pca.explained_variance_ratio_))
return pca_train_data, pca_test_data
# since the data is too big, we take the data apart
data1 = data.iloc[0:10000,:]
data2 = data.iloc[10000:20000,:]
data3 = data.iloc[20000:30000,:]
data4 = data.iloc[30000:40005,:]
X_train1, X_test1, y_train1,y_test1 = my_split(data1, img_size=(128,128), test_size=0.1)
X_train2, X_test2, y_train2,y_test2 = my_split(data2, img_size=(128,128), test_size=0.1)
X_train3, X_test3, y_train3,y_test3 = my_split(data3, img_size=(128,128), test_size=0.1)
X_train4, X_test4, y_train4,y_test4 = my_split(data4, img_size=(128,128), test_size=0.1)
X_train = np.concatenate((X_train1, X_train2, X_train3, X_train4))
X_test = np.concatenate((X_test1, X_test2, X_test3, X_test4))
y_train = pd.concat((y_train1, y_train2, y_train3, y_train4))
y_test = pd.concat((y_test1, y_test2, y_test3, y_test4))
# perform PCA
whole_train, whole_test = my_PCA(X_train, X_test, y_train, y_test, num_of_components=0.8)
# save the data
whole_train.to_csv('whole_train.csv')
whole_test.to_csv('whole_test.csv')
# ## <center>$\mathcal{FIN}$</center>
| 1_Data_pre_processing_for_ML.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Resnet Supervised Learning
# This file trains a supervised learning model (Resnet50) using transfer learning to train on an unknown dataset. And the results give us a baseline accuracy to compare with the semi-supervised model (Mixmatch).
from keras.utils import np_utils
import numpy as np
from resnet_lib import create_dataset, create_model, test_callback, tune_model, visualize_model
# ## Train on streetview_v3 dataset
# +
# Global constants
NUM_CLASSES = 2
IMAGE_SIZE = 64
BATCH_SIZE = 64
EPOCHS = 300
# Directories
INPUT_TF_TRAIN = '../Mixmatch/ML_DATA/streetview_v3_64-train.tfrecord'
INPUT_TF_TEST = '../Mixmatch/ML_DATA/streetview_v3_64-test.tfrecord'
# -
# Construct the datsets
X_train, y_train = create_dataset(INPUT_TF_TRAIN)
X_test, y_test = create_dataset(INPUT_TF_TEST)
y_train = np_utils.to_categorical(y_train, NUM_CLASSES)
y_test = np_utils.to_categorical(y_test, NUM_CLASSES)
# +
# Build model and fit the top layers
model = create_model((IMAGE_SIZE, IMAGE_SIZE, 3), NUM_CLASSES, lr=0.0001)
model.fit(x=X_train, y=y_train, epochs=EPOCHS, batch_size=BATCH_SIZE, verbose=1, validation_split = 0.005, validation_freq=1, callbacks=[test_callback((X_test, y_test))])
# -
# Visualize the model and determine which block will be trained in the tuning phase
visualize_model(model)
# Fine tune the model
EPOCHS = 30
model = tune_model(model, trainable_layer=0, lr=0.0001)
model.fit(x=X_train, y=y_train, epochs=EPOCHS, batch_size=BATCH_SIZE, verbose=1, validation_split = 0.1, validation_freq=1, callbacks=[test_callback((X_test, y_test))])
# Output the predictions of a model to csv files for further analysis
pred = model.predict(X_test.astype(np.float32))
np.savetxt("results.csv", np.concatenate((y_test, pred), axis=1), delimiter=",")
# ## Train on streetview_v4 dataset
# +
# Global constants
NUM_CLASSES = 2
IMAGE_SIZE = 64
BATCH_SIZE = 64
EPOCHS = 300
# Directories
INPUT_TF_TRAIN = '../Mixmatch/ML_DATA/streetview_v4_64-train.tfrecord'
INPUT_TF_TEST = '../Mixmatch/ML_DATA/streetview_v4_64-test.tfrecord'
# -
# Construct the datsets
X_train, y_train = create_dataset(INPUT_TF_TRAIN, word_embeddings=True)
X_test, y_test = create_dataset(INPUT_TF_TEST, word_embeddings=True)
y_train = np_utils.to_categorical(y_train, NUM_CLASSES)
y_test = np_utils.to_categorical(y_test, NUM_CLASSES)
# Build model and fit the top layers
model = create_model((IMAGE_SIZE, IMAGE_SIZE, 4), NUM_CLASSES, lr=0.0001)
model.fit(x=X_train, y=y_train, epochs=EPOCHS, batch_size=BATCH_SIZE, verbose=1, validation_split = 0.005, validation_freq=1, callbacks=[test_callback((X_test, y_test))])
| Supervised_learning/resnet_main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import re,pickle,nltk.data,os,progressbar,time,gensim
import pandas as pd
from nltk.stem import WordNetLemmatizer
from nltk.corpus import stopwords
from gensim.models import word2vec
from nltk.corpus import wordnet
from gensim.models import Phrases
from gensim.models.phrases import Phraser
import os
# ##### <b> 1. Load Data And Write Texts To Individual .txt Files </b>
# +
# # Read data from files
# train = pd.read_csv("data/train.csv", header=0)
# test = pd.read_csv( "data/test.csv", header=0)
# print("Read %d labeled train reviews and %d unlabelled test reviews" % (len(train),len(test)))
# train_comments = train['comment_text'].fillna("_na_").tolist()
# test_comments = test['comment_text'].fillna("_na_").tolist()
# all_comments = train['comment_text'].fillna("_na_").tolist() + test['comment_text'].fillna("_na_").tolist()
# def comments_to_txtfile(comment_list,output_file):
# with open(output_file, "w+") as output:
# for comment in comment_list:
# comment = " ".join(comment.split()).replace('"',"").replace('|',"").replace('{',"").replace('}',"")
# comment = " ".join(comment.split()).replace('"',"")
# comment = re.sub("[^a-zA-Z0-9 ]","",str(comment)) #Remove unneeded whitespace and punctuation and move to lowercase
# output.write("%s\n" % str(comment))
# return print("%s Written" % output_file)
# comments_to_txtfile(train_comments,'data/raw_comments/train_comments.csv')
# comments_to_txtfile(test_comments,'data/raw_comments/test_comments.csv')
# comments_to_txtfile(all_comments,'data/raw_comments/all_comments.csv')
# -
def nlp_utility(input_file,
embeddings_file ='',
fasttext_file = '',
remove_stops = True,
lemmatize = True, #Must lemmatize to be able to remove proper nouns - can be fixed in future
remove_proper_nouns = True,
n_grams = 0,
fasttext_spellcheck = True,
filter_embeddings_file = False,
max_vocabulary = 0
):
import os,shutil,smart_open,gensim,sys,time,re
from gensim.models import Word2Vec
from sys import platform
from nltk.corpus import stopwords
from nltk.stem import WordNetLemmatizer
from nltk.corpus import wordnet
from nltk import pos_tag
from gensim.models import FastText
stops = set(stopwords.words("english"))
unfiltered_glove_path = 'w2v_models/unfiltered_models/' + embeddings_file
filtered_glove_path = 'w2v_models/filtered_models/' + 'vocab=' + str(max_vocabulary) + '.remove_pn=' + str(remove_proper_nouns) + '.remove_stops=' + str(remove_stops) + '.lemmatize=' + str(lemmatize) + '.spellcheck=' + str(fasttext_spellcheck) + '.' + embeddings_file
gensim_filtered_glove_path = 'w2v_models/gensim_filtered_models/' + 'vocab=' + str(max_vocabulary) + '.remove_pn=' + str(remove_proper_nouns) + '.remove_stops=' + str(remove_stops) + '.lemmatize=' + str(lemmatize) + '.spellcheck=' + str(fasttext_spellcheck) + '.' + embeddings_file
output_file = 'data/tokenized_comments/' + 'vocab=' + str(max_vocabulary) + '.remove_pn=' + str(remove_proper_nouns) + '.remove_stops=' + str(remove_stops) + '.lemmatize=' + str(lemmatize) + '.spellcheck=' + str(fasttext_spellcheck) + '.' + embeddings_file + '.' + input_file.split('/')[2]
print('Result File = %s' % output_file)
#Create a generator function to run through the input file one line at a time:
class FileToComments(object):
def __init__(self, filename):
self.filename = filename
def __iter__(self):
for line in open(self.filename, 'r'):
yield line
#Function used in Step 2 to match treebank tags to pos tags
def get_wordnet_pos(treebank_tag):
if treebank_tag.startswith('J'):
return wordnet.ADJ
elif treebank_tag.startswith('V'):
return wordnet.VERB
elif treebank_tag.startswith('N'):
return wordnet.NOUN
elif treebank_tag.startswith('R'):
return wordnet.ADV
else:
return ''
def filter_vectors_gensim(unfiltered_glove_file,comment_file):
unfiltered_glove_path = 'w2v_models/unfiltered_models/' + unfiltered_glove_file
filtered_glove_path = 'w2v_models/filtered_models/' + 'vocab=' + str(max_vocabulary) + '.remove_pn=' + str(remove_proper_nouns) + '.remove_stops=' + str(remove_stops) + '.lemmatize=' + str(lemmatize) + '.spellcheck=' + str(fasttext_spellcheck) + '.' + unfiltered_glove_file
gensim_filtered_glove_path = 'w2v_models/gensim_filtered_models/' +'vocab=' + str(max_vocabulary) + '.remove_pn=' + str(remove_proper_nouns) + '.remove_stops=' + str(remove_stops) + '.lemmatize=' + str(lemmatize) + '.spellcheck=' + str(fasttext_spellcheck) + '.' + unfiltered_glove_file
comment_streamer = FileToComments(comment_file)
#Create a Vocabulary From Our Existing Words
from collections import defaultdict
import codecs, gensim
vocab = defaultdict(int)
for sentence in comment_streamer:
sentence = sentence.split()
for word in sentence:
vocab[word] += 1
vocab = set(vocab)
print('Vocabulary set, starting to filter word embeddings')
nread = 0
nwrote = 0
with codecs.open(unfiltered_glove_path, encoding='utf-8') as f:
with codecs.open(filtered_glove_path, 'w', encoding='utf-8') as out:
for line in f:
nread += 1
line = line.strip()
if not line: continue
if line.split(u' ', 1)[0] in vocab:
out.write(line + '\n')
nwrote += 1
if (nwrote >= max_vocabulary) & (max_vocabulary > 0):
sys.stdout.write("\r\n")
print('read %s lines, wrote %s' % (nread, nwrote))
print('Prepending Gensim W2V Line Count and Embedding Size')
gensim.scripts.glove2word2vec.glove2word2vec(filtered_glove_path,gensim_filtered_glove_path)
return gensim_filtered_glove_path
else:
sys.stdout.write("Read: %d lines. Wrote %d To Word Embedding File \r" % (nread,nwrote))
sys.stdout.flush()
sys.stdout.write("\r\n")
print('read %s lines, wrote %s' % (nread, nwrote))
print('Prepending Gensim W2V Line Count and Embedding Size')
gensim.scripts.glove2word2vec.glove2word2vec(filtered_glove_path,gensim_filtered_glove_path)
return gensim_filtered_glove_path
#Step 1 - Split Words And Make Lowercase
comment_streamer = FileToComments(input_file)
counter = 0
entities_removed = 0
punctuation = re.compile(r'[►*/-@.?!~,":#$;\'()=+|0-9]') #[punctuation and numbers to remove]
with open('intermediate_comments_preproc.txt', "w+") as output:
for comment in comment_streamer:
comment =punctuation.sub("", comment)
comment = comment.lower().split()
#Step 1 - Remove Stopwords
if remove_stops:
comment = [word for word in comment if not word in stops]
#Step 2 - Lemmatize
if lemmatize:
tagged_words = pos_tag(comment)
tokenized_words = []
for word in tagged_words:
if (remove_proper_nouns) & ((word[1] == "NNP") | (word[1] == "NNPS")):
entities_removed += 1
else:
thisTag = get_wordnet_pos(word[1])
if thisTag == '':
tokenized_words.append(word[0])
else:
thisLemma = WordNetLemmatizer().lemmatize(word[0],thisTag)
tokenized_words.append(thisLemma)
output.write("%s\n" % ' '.join(tokenized_words))
else:
output.write("%s\n" % ' '.join(comment))
counter += 1
sys.stdout.write("Cleaned %d comments from %s. Removed %d entities. \r" % (counter,input_file,entities_removed))
sys.stdout.flush()
sys.stdout.write("\r\n")
# Step 4 - Filter Word Embedding File (For Speed In Training)
# We now have our commenst rewritten in a clean format with stop words removed and lemmatization in place
# Now we need to take our chosen vector_model and sort it for words found in our sentences
if filter_embeddings_file:
filtered_vectors_path = filter_vectors_gensim(embeddings_file,comment_file = 'intermediate_comments_preproc.txt')
print('Embedding file filtered and cleaned for corpora. Location : %s' % gensim_filtered_glove_path)
return
else:
filtered_vectors_path = gensim_filtered_glove_path
#Step 5- Use Fasttext Model For Spelling Correction
if fasttext_spellcheck:
#Load Glove Model
glove_word_vectors = gensim.models.KeyedVectors.load_word2vec_format(filtered_vectors_path)
print("Filtered Gensim Model Loaded")
#Load Fasttext Model
fasttext_vectors = FastText.load_fasttext_format(fasttext_file)
print("Fasttext Model Loaded")
intermediate_token_streamer = FileToComments('intermediate_comments_preproc.txt')
model_vocabulary = set(glove_word_vectors.wv.vocab)
word_subs = 0
counter = 0
print("Substituting mispelled words using Fasttext Model")
with open(output_file, "w+") as output:
for wordlist in intermediate_token_streamer:
wordlist = wordlist.split()
output_words = []
for word in wordlist:
if word in model_vocabulary:
output_words.append(word)
else:
try:
for pairing in fasttext_vectors.wv.most_similar(word,topn=3):
if pairing[0] in model_vocabulary:
output_words.append(pairing[0])
word_subs += 1
except:
next
output.write("%s\n" % ' '.join(output_words))
counter += 1
sys.stdout.write("Substituted %d Mispelled Words from %d comments\r" % (word_subs,counter))
sys.stdout.flush()
print("Substituted %s Words. Printed to %s" % (word_subs,output_file))
else:
intermediate_token_streamer = FileToComments('intermediate_comments_preproc.txt')
print("Saving to Result File")
with open(output_file, "w+") as output:
for wordlist in intermediate_token_streamer:
wordlist = wordlist.split()
output.write("%s\n" % ' '.join(wordlist))
print("Printed to %s" % (output_file))
return
print('Available Models:')
os.listdir('w2v_models/unfiltered_models/')
# Set Params For Preprocessing
remove_stops = True
lemmatize = True
fasttext_spellcheck = False
remove_proper_nouns = True
embeddings_file = 'glove.840B.300d.txt'
fasttext_file = 'fasttext-training/word_vector_models/cbow_fasttext_model.bin'
max_vocab = 100000
#Filter Vectors
nlp_utility(input_file = 'data/raw_comments/all_comments.csv',
embeddings_file =embeddings_file,
fasttext_file = fasttext_file,
remove_stops = remove_stops,
remove_proper_nouns = remove_proper_nouns,
lemmatize = lemmatize,
fasttext_spellcheck = fasttext_spellcheck,
filter_embeddings_file = True,
max_vocabulary = max_vocab
)
nlp_utility(input_file = 'data/raw_comments/train_comments.csv',
embeddings_file = embeddings_file,
fasttext_file = fasttext_file,
remove_stops = remove_stops,
lemmatize = lemmatize,
remove_proper_nouns = remove_proper_nouns,
fasttext_spellcheck = fasttext_spellcheck,
max_vocabulary = max_vocab
)
nlp_utility(input_file = 'data/raw_comments/test_comments.csv',
embeddings_file = embeddings_file,
fasttext_file = fasttext_file,
remove_stops = remove_stops,
lemmatize = lemmatize,
remove_proper_nouns = remove_proper_nouns,
fasttext_spellcheck = fasttext_spellcheck,
max_vocabulary = max_vocab
)
| advanced-nlp-prepoc-r2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="6l-i8H-gQ40U"
#
# Right click and open the link below in new tab to open the notebook in google colab.
#
# [](https://colab.research.google.com/github/KenR22/twitter_spark_nlp/blob/master/bds_project.ipynb)
# + [markdown] id="rmzeHHvQ9_SV"
# # Download the Data
# + [markdown] id="ifD8Nhf_FW3W"
# Upload your Kaggle.json file here if you want to download the data directly from Kaggle. To avoid downloading data from Kaggle, upload the csv file under the data folder in github repo and skip the kaggle download section:
# + colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY>", "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": ""}}, "base_uri": "https://localhost:8080/", "height": 73} id="Q0YkMAMz9Vtn" outputId="49e4e1fd-43a1-4d3f-940d-4ca7767d1589"
from google.colab import files
uploaded = files.upload()
# + [markdown] id="Y1x1DhcqRLbk"
# # Kaggle Download
# + colab={"base_uri": "https://localhost:8080/"} id="tISy0-IRF9hO" outputId="80996b7e-815e-487a-979d-1bd8fc5e7b8c"
# !kaggle
# !mv kaggle.json ../root/.kaggle/kaggle.json
# !kaggle datasets download -d ayushggarg/all-trumps-twitter-insults-20152021
# !unzip all-trumps-twitter-insults-20152021.zip
# + [markdown] id="oTNLrsWA98iV"
# # Imports
#
# + colab={"base_uri": "https://localhost:8080/"} id="nNVmzwWKmOIx" outputId="c92c480d-5ed4-41b5-d0fe-b00d7176b7eb"
# !wget http://setup.johnsnowlabs.com/colab.sh -O - | bash
# + id="Wia9EJgLobJj"
import sparknlp
from sparknlp.pretrained import PretrainedPipeline
spark = sparknlp.start()
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
import pyspark
from pyspark.sql import *
from pyspark.sql.functions import *
from pyspark import SparkContext, SparkConf
from pyspark.sql.functions import to_date,month, year
import plotly.graph_objs as go
import pandas as pd
import requests
# + id="wWnirnXeoeqG"
# create the session
conf = SparkConf().set("spark.ui.port", "4050")
# create the context
spark = SparkSession.builder.getOrCreate()
# + [markdown] id="7B0ZeHoskrfL"
# ## Load the Data
# If the download from Kaggle worked then the csv file should be in working directory. The csv is loaded as RDD below. There is one date column that was converted to date format from string manually.
# + id="hDOM31wgorgv" colab={"base_uri": "https://localhost:8080/"} outputId="3f94c4a3-fe47-480b-8a3f-390240eee5ba"
df = spark.read.format("csv").load("trump_insult_tweets_2014_to_2021.csv",
header='true',
inferSchema='false')
df = df.select(to_date(df.date, 'yyyy-MM-dd').alias('date'),"target","insult","tweet")
df.printSchema()
# + [markdown] id="qam5iwb8lHcN"
# Show the first 5 rows of the RDD
# + colab={"base_uri": "https://localhost:8080/"} id="uM1MJZyDpGXW" outputId="25f4e342-afd7-48af-f72f-c835ba6d02d5"
df.show(5)
# + colab={"base_uri": "https://localhost:8080/"} id="3desoWg3wEQi" outputId="4861ad63-6389-490e-bb77-262a51dd7209"
print("Total Number of Tweets: {0} ".format(df.count()))
# + [markdown] id="tafwTWXXle5D"
# That is a lot of insult in 5 years!
#
# + [markdown] id="7W6NjvLbjz4J"
#
# + [markdown] id="p2FKBZIhyhHi"
# # Targets
#
# How many targets did he have in his tweets? Were they all the similar?
# + [markdown] id="e5kwpsLgl9WR"
# Let's group by the target column and get the counts.
# + id="FAdB_c9TwMJH"
count_target=df.groupBy("target").agg(count("*").alias("Number of Tweet")).sort(desc("Number of Tweet")).filter(col("Number of Tweet")>120)
# + colab={"base_uri": "https://localhost:8080/"} id="TsyjIfO74HgE" outputId="92fb0077-e43a-4025-9c05-835b5470d1a2"
count_target.show()
# + id="5w93LCpG3FKM"
import plotly.express as px
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="NI0VuybPyeUh" outputId="a8134d23-180c-45cb-d264-23735989ebf4"
fig = px.bar(count_target.toPandas(), x='target', y='Number of Tweet')
fig.show()
# + [markdown] id="UqO4D1fNGRN0"
# The biggest target of Trump's tweet is the media. Other top targets are his political rivals like <NAME> and <NAME>. Events like impeachment inquiry and 2020 election can also be seen on his top targets.
# + [markdown] id="Jhdt9sIzGqrc"
# ## Insult Types
# There are different types of instult type. How many uniqe insults?
# + colab={"base_uri": "https://localhost:8080/"} id="8z3usHQmITaB" outputId="96852fe6-6ff3-4690-907e-22240fb89a9d"
print("Unique type of insults: ",len(df.select('insult').distinct().collect()))
# + [markdown] id="WwgiXcBOJAcM"
# However there are in total 10492 tweets. So lets take a look a at top recurring insult types.
# + id="0aUjC0xIK3BZ"
insult_count=df.groupBy("insult").agg(count("*").alias("Occurence")).sort(desc("Occurence")).filter(col("Occurence")>25)
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="QtH9aYp7LUPp" outputId="ea70a5ac-582f-4e7c-ce3f-68895af806b0"
fig = px.bar(insult_count.toPandas(), x='insult', y='Occurence')
fig.show()
# + [markdown] id="NAQCPEAtLfTp"
# Wow his insults towards media is very high! Calling names like, "sleepy" and "crooked" pops up too.
# + [markdown] id="3-jI4eEmLvqT"
# # Tweet Date
# Let's take a look at how his tweeting has evolved over time.
# + id="enEyOFqDL5Q1"
date_agg=df \
.groupBy(year("date").alias("year"),month("date").alias("month"))\
.agg(count("*").alias("Tweets"))
# + id="gRumZeO3o85t"
date_agg=date_agg.withColumn(
"date",
expr("make_date(year, month, 1)")
).sort(desc("date"))
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="_Bq-ExIxp87B" outputId="81437677-efc5-469c-8a20-2eebbf0bf3be"
fig = px.line(date_agg.toPandas(), x='date', y="Tweets")
fig.show()
# + [markdown] id="iUm9quOlqokm"
# He has been consistent in insulting people over the years. There are some sudden peaks that can be related with
# + [markdown] id="-YtQKLRCe1Gf"
# ## Sentiment Analysis
#
# These tweets are all insults. However could there be tweets that are positive? Since the tweets do not have any specific labels, a pretrained model for sentiment analysis will be deployed for sentiment analysis.
# + id="nJYL-AQdgpqr"
import sparknlp
from sparknlp.pretrained import PretrainedPipeline
# + colab={"base_uri": "https://localhost:8080/"} id="de8TU6Kug0RS" outputId="193320a3-e8ec-4a32-8e5e-4341bcc66e9f"
pipeline = PretrainedPipeline("analyze_sentimentdl_use_twitter", lang="en")
# + id="-wjGaTR2qUdx"
df_result = \
pipeline.transform(df.withColumnRenamed("tweet", "text"))
# + colab={"base_uri": "https://localhost:8080/"} id="WLyO7aF_uarJ" outputId="7753c5c8-d144-463e-f5d0-bc6e8cd61cee"
df_result.show()
# + colab={"base_uri": "https://localhost:8080/"} id="fef9OLuFrDqs" outputId="6893b560-3d30-4d99-aa7c-544c4da8ed6b"
df_result.printSchema()
# + id="etRqC09AyGz1"
sentiment_df=df_result.select("*",df_result.sentiment.result.alias('output'))
# + id="T9CQbUC2AV6m"
sentiment_count=sentiment_df.groupBy("output").agg(count("*").alias("Number of Tweet")).sort(desc("Number of Tweet"))
# + colab={"base_uri": "https://localhost:8080/"} id="tIzj1agHB_db" outputId="de463039-8340-41c4-d4b8-90b8bd8c2140"
sentiment_count.show()
# + [markdown] id="hdPgxWlMuQN4"
# Interesting! The sentiments should all be negative as this dataset contains insults only. Let's take some of the Tweets that were predicted as positive.
# + colab={"base_uri": "https://localhost:8080/"} id="gqUPAXJGvzD-" outputId="3ce1be05-2055-47d5-ee6d-7abe6e6de325"
explore_sentiment=sentiment_df.selectExpr('output[0] as output',"text")
explore_sentiment.filter(explore_sentiment.output == "positive").dropDuplicates().show(truncate=False)
# + [markdown] id="UzPaNKqrCi-7"
# ## Finding Entities
#
# Using a similar pretrained pipeline, entities can be detected too. This section will cover that method.
# + colab={"base_uri": "https://localhost:8080/"} id="wPOpw0rm3Gg2" outputId="5b1d5a74-c716-47c0-80ce-9df33b1cd0e0"
pipeline = PretrainedPipeline('onto_recognize_entities_bert_tiny')
# + id="_pV5yVq84rKA"
df_entity = \
pipeline.transform(df.withColumnRenamed("tweet", "text"))
# + colab={"base_uri": "https://localhost:8080/"} id="zbCvrGYK5b6n" outputId="f4d66879-f859-4c8e-9e3f-ce50666c28e9"
df_entity.show()
# + colab={"base_uri": "https://localhost:8080/"} id="iUHNXDyf5nJ2" outputId="d573eb1e-8a26-4921-a603-0c6b497940b7"
df_entity.printSchema()
# + id="H2YMd_SDWuEw"
# + id="JPo-fTPw7YV9"
NER_df=df_entity.selectExpr('entities.result as entities',"text","NER.result as tokens").dropDuplicates()
# + colab={"base_uri": "https://localhost:8080/"} id="Sa3Nxk0wWvBp" outputId="2181fce9-4e63-4c44-eb25-f209ffb11b5f"
NER_df.show(10,truncate=False)
# + id="vsYSC7t2f4pc"
NER_count=NER_df.select("entities").withColumn("entities" ,explode('entities')).groupBy("entities").agg(count("*").alias("Occurence")).sort(desc("Occurence")).filter(col("Occurence")>20)
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="csZ9W3TsiO83" outputId="85902b68-080b-45b0-9436-636a9ccad27b"
fig = px.bar(NER_count.toPandas(), x='entities', y='Occurence')
fig.show()
# + [markdown] id="eCAtr9ozGfRK"
# Unlike the targets columns the most occuring entity is democrats. This might be due to the fact that he mentioned democrats when tweeting against Hilary clinton and <NAME>. Intestingly 2016 and 2020 came up as years. This makes sense because these two years are the ones when Trump was involved in the presidential elections.
# + id="watoJZDqi4gS"
NER_type_count=NER_df.select("tokens").withColumn("tokens" ,explode('tokens')).groupBy("tokens").agg(count("*").alias("Occurence")).sort(desc("Occurence")).filter(col("Occurence")<5000)
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="iqvlv0TkkMOy" outputId="077e34ac-4d46-4281-8b22-286ac826654e"
fig = px.bar(NER_type_count.toPandas(), x='tokens', y='Occurence')
fig.show()
# + [markdown] id="DtSrcTUaGiW1"
# The plot above shows his insults mentined more persons than organizations. B-Person represents person entity. B-GPE is geopolitical entity. This may be due to the fact that he mentions U.S. a lot in his tweets.
| bds_project.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python2
# ---
import gpflow
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
import sys
sys.path.append('../')
from GPHetero import hetero_kernels, hetero_likelihoods, hetero_gpmc
from pyDOE import *
import os
from scipy.stats import norm
class Ex5Func(object):
def __init__(self, sigma=lambda x: 0.5):
self.sigma = sigma
def __call__(self, x):
"""
Dette et. al. function.
<NAME>, and <NAME>. "Generalized Latin hypercube design for computer experiments." Technometrics 52, no. 4 (2010): 421-429.
"""
y = 4 * ((x[0] - 2 + 8 * x[1] - 8 * (x[1] ** 2)) ** 2) + (3 - 4 * x[1]) ** 2 + 16 * np.sqrt(x[2] + 1) * ((2 * x[2] - 1)**2)
return (y - 50) / 50.
# +
dim = 3
n = 50
noise=0
sigma = eval('lambda x: ' + str(noise))
objective = Ex5Func(sigma=sigma)
X = lhs(dim, n , criterion='center')
Xnorm = (X - 0.5) /0.5
Y = np.array([objective(x) for x in X])[:, None]
# -
kerns_list = [gpflow.kernels.RBF(1), gpflow.kernels.RBF(1), gpflow.kernels.RBF(1)]
mean_funcs_list = [gpflow.mean_functions.Constant(0), gpflow.mean_functions.Constant(0),
gpflow.mean_functions.Constant(0)]
nonstat = hetero_kernels.NonStationaryLengthscaleRBF()
m = hetero_gpmc.GPMCAdaptiveLengthscaleMultDimDev(Xnorm, Y, kerns_list, nonstat, mean_funcs_list)
# +
for i in xrange(dim):
m.kerns_list[i].lengthscales.prior = gpflow.priors.Gamma(1., 1.)
m.kerns_list[i].variance.prior = gpflow.priors.Gamma(1., 1.)
m.nonstat.signal_variance.prior = gpflow.priors.Gamma(1., 2.)
m.likelihood.variance = 1e-6
m.likelihood.variance.fixed = True
# +
#m.mean_funcs_list[0].c = 0.
#m.mean_funcs_list[0].c.fixed = True
#m.mean_funcs_list[1].c = 0.
#m.mean_funcs_list[1].c.fixed = True
#m.mean_funcs_list[2].c = 0.
#m.mean_funcs_list[2].c.fixed = True
# -
m.optimize(maxiter=5000) # start near MAP
samples = m.sample(500, verbose=True, epsilon=0.00005, thin = 2, burn = 500, Lmax = 20)
plt.figure(figsize=(16, 4))
plt.plot(samples[:,10:80])
X_test = lhs(dim, 100 , criterion='center')
X_test_norm = (X_test - 0.5) /0.5
Y_test = np.array([objective(x) for x in X_test])[:, None]
sample_df = m.get_samples_df(samples)
mean_f_mat = np.zeros(shape=(sample_df.shape[0], X_test_norm.shape[0]))
var_f_mat = np.zeros(shape=(sample_df.shape[0], X_test_norm.shape[0]))
for i, s in sample_df.iterrows():
m.set_parameter_dict(s)
mean_f, var_f = m.predict(X_test_norm)
mean_f_mat[i, :] = mean_f[:,0]
var_f_mat[i, :] = np.diag(var_f)
plt.scatter(mean_f_mat[1,:], Y_test)
mean_f_mat.shape
Y_test.shape
| notebooks/.ipynb_checkpoints/gpflow3d_nonstat_new_class_march7-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: SageMath 9.0
# language: sage
# name: sagemath
# ---
# # Groups
# Here we dive into groups in SageMath. There are many different types of groups that we can use in SageMath, including:
#
# * Symmetric Groups
# * Dihedral Groups
# * Alternating Groups
# * The Klein Four Group
# * The Quaternion Group
# * And many others
#
# More information about what named groups are available can be found in your local sage distribution, for example, on my computer, opening the file
#
# > /usr/lib/python3/dist-packages/sage/groups/perm-gps/pergroup_named.py
#
# provides much information about other types of groups that can use defined. Most of these we will not use in this series of documents, but they are there for you to use if would like.
#
# We begin with the Dihedral Group. Considering the symmetries of a triangle, we can define the Dihedral Group of order 3 by writting the following:
T = DihedralGroup(3)
T
# To see all of the elements in this group, we can use the `list` command.
T.list()
# In the above list, we see the six elements listed of the Dihedral Group of order six. SageMath uses a notation that we may not have seen before if we are still being introduced to Groups in Abstract Algebra. It is refered to as cycle notation, and if you are in an Undergraduate Abstract Algebra class, you will probably be introduced to cycle notation soon. A cycle $ (1,3,2) $ in sage denotes a permutation that sends $1\rightarrow 3$, $3\rightarrow 2$ and $2\rightarrow 1$, they are used extensively in Group theory with permutations. So in the Dihedral group above, we see the two rotations of a triangle, and the three reflections, as well as the identity $()$.
#
# We can specify elements of this group by writing the following:
sigma = T('(1,3,2)')
tau = T('(2,3)')
# Now that we have some elements from this group, we can perform operations with them, like the following.
print(sigma*tau)
print(tau*sigma)
print(sigma^3)
print(tau^(-1))
# Note that in SageMath, permutation multiplication is done from LEFT to RIGHT.
#
# Some other groups, we can define look like this:
S5 = SymmetricGroup(5)
print(f"The order of S5 is {S5.order()}")
print(f"Is S5 abelian? {S5.is_abelian()}")
# Another great function with groups in SageMath is the `cayley_table()` function. This is used as follows:
T.cayley_table()
# Note that this will be quite large for most groups, usually it will be much too large to be usefull, for example:
S5.cayley_table()
# Another great use of SageMath for groups is the `cayley_graph()` function. It provides a similar operation as the function `cayley_table()` but provides a graphic to visualize the structure of the group.
T.cayley_graph()
# The other types of groups are declared by using the following syntax:
#
# * A = AlternatingGroup(n)
# * Q = QuaternionGroup()
# * K = KleinFourGroup()
| Groups/Groups.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # README
#
# ---
#
# VCP SDKを用いてクラウド上にCoursewareHub環境を構築します。
# ## 変更履歴
#
# ### [20.04.0](https://github.com/nii-gakunin-cloud/ocs-templates/releases/tag/release%2F20.04.0)からの変更点
#
#
# 主に[NII-cloud-operation/CoursewareHub-LC_platform](https://github.com/NII-cloud-operation/CoursewareHub-LC_platform)の [#14](https://github.com/NII-cloud-operation/CoursewareHub-LC_platform/pull/14)から[#25](https://github.com/NII-cloud-operation/CoursewareHub-LC_platform/pull/25)までの変更内容を取り込んでいます。
#
# 追加された主要な機能を以下に記します。
#
# * 学認連携に基づく認証
# * LMS(Learning Management System)との認証連携
# * managerノードとNFSサーバを別ノードとして構成することを選択できるようになった
# * single-userコンテナに割り当てるリソース量をグループや認可タイプ毎に設定できるようになった
# ## はじめに
#
# このアプリケーションテンプレートではVCPで作成したノードに[CoursewareHub](https://github.com/NII-cloud-operation/CoursewareHub-LC_jupyterhub-deploy)環境を構築します。
# ### CoursewareHubのユーザ認証について
#
# CoursewareHubではユーザの認証機能として以下に示す三つの方式に対応しています。
#
# * ローカルユーザ認証
# * CoursewareHubのローカルユーザデータベースを用いてユーザ管理を行う
# * 学認連携に基づく認証
# * [学認](https://www.gakunin.jp/)のSPとして登録し、認証連携を行う
# * CoursewareHubを直接SPとしては登録せずに、プロキシ(IdPプロキシ)を経由して連携することも可能
# * LMS(Learning Management System)との認証連携
# * [LTI 1.3](http://www.imsglobal.org/spec/lti/v1p3/)による認証連携を行う
# * このテンプレートでは連携するLMSとして[Moodle](https://moodle.org/)を想定している
#
# それぞれの認証機能は共存することが可能になっています。ただし、学認連携認証を用いる場合はコンテナの構成や設定手順が異なります。そのため、それに応じた異なる構築手順を用意しています。
#
# 一方 LMSとの認証連携を行う場合は、まずローカルユーザ認証、あるいは学認連携認証の手順でCoursewareHubを構築してください。その後にLMSとの認証連携の設定を追加する手順となっています。
# ### コンテナの構成について
#
# ローカルユーザ認証のみを用いる場合と、学認連携認証を利用する場合とではコンテナの構成が異なります。ここでは、それぞれのコンテナ構成について記します。
#
# CoursewareHubでは、学認連携の有無、あるいは連携方法の違いにより以下に示す方式を選択することができます。
# * ローカルユーザ認証のみを利用する
# * 学認フェデレーションに参加し、学認のIdPを利用して認証を行う
# - IdP-proxyをSPとして学認に登録し、複数のCoursewareHubがIdP-proxyを通して学認のIdPを利用する
# - CoursewareHubを直接SPとして学認に登録する
#
# それぞれの方式に対応する構成図を以下に示します。
#
# #### ローカルユーザ認証のみを利用する場合
#
# 
#
# #### IdP-proxyを利用する場合
#
# 
#
# #### CoursewareHubを直接SPとして登録する場合
#
# 
# ### ノード構成
#
# CoursewareHubのノードは役割に応じて以下のものに分類されます
#
# * manager
# - JupyterHub, auth-proxy, PostgreSQLなどのSystemコンテナを実行するノード
# - Docker Swarmのmanagerノードとなる
# * worker
# - single-user Jupyter notebook serverを実行するノード
# - Docker Swarm の workerノードとなる
#
# CoursewareHubではデータやNotebookなどをノード間で共有するためにNFSを利用します。NFSサーバの配置により以下の3つパターン構成が可能となっています。
#
# 1. 構成1
# - managerノードにNFSサーバを配置する
# 1. 構成2
# - managerノードとNFSサーバを別のノードとして構成する
# 1. 構成3
# - 構成2のNFSサーバに、新たなCoursewareHub環境を追加する構成
# #### 構成1
#
# managerノードでNFSサーバを実行します。
#
# 
# #### 構成2
#
# managerノードとNFSサーバを分離し別々のノードとして構築します。
#
# 
# #### 構成3
#
# 構成2のNFSサーバに、新たなCoursewareHub環境を追加します。NFSサーバは複数のCoursewareHub環境で共有されます。
#
# 
# ### 収容設計について
# #### managerノード
# * システム用コンテナが実行される
# - auth-proxyコンテナ
# - JupyterHubコンテナ
# - PostgreSQLコンテナ
# * ユーザが利用する single-userサーバコンテナは実行されない
# * NFSサーバをmanagerノードに共存させる場合(構成1)はディスク容量を適切な値に設定する
# #### workerノード
# * ユーザが利用するsingle-userコンテナが実行される
# * single-userコンテナのリソース量として以下の設定を行っている
# - 最大CPU利用数
# - 最大メモリ量(GB)
# - 保証される最小割当てメモリ量(GB)
# * システム全体で必要となるリソース量を見積もるには
# - (コンテナに割り当てるリソース量)×(最大同時使用人数)+(システムが利用するリソース量)×(ノード数)
# #### 運用例
# * 最大同時使用人数
# - 400 人
# * コンテナに割り当てるリソース量
# - メモリ最小値保証
# - 1GB
# - メモリ最大値制限
# - 2GB(swap 4GB)
# - CPU最大値制限
# - 200% (2cores)
#
# 上記の条件で運用を行った際の実績値を示します。
# * managerノード
# - vCPU
# - 10
# - memory
# - 16GB
# - HDD
# - 800GB
# * workerノード
# - ノードA
# - ノード数
# - 4
# - vCPU
# - 30
# - memory
# - 100GB
# - HDD
# - 300GB
# - ノードB
# - ノード数
# - 1
# - vCPU
# - 20
# - memory
# - 80GB
# - HDD
# - 300GB
#
# > workerノードはリソース量の異なるノードAとノードBで構成されていた。
#
# workerノードのメモリ総量は480GB(=100×4+80)となっていますが、これは以下のように見積もっています。
# ```
# (コンテナのメモリ最小値保証)×(最大同時使用人数)+(システム利用分)
# = 1GB × 400人 + 80GB
# ```
# ## 事前に準備が必要となるものについて
#
# このアプリケーションテンプレートを実行するにあたって事前に準備が必要となるものについて記します。
# ### VCノード
#
# ノードを作成するとき必要となるものについて記します。
# * VCCアクセストークン
# - VCP SDKを利用してクラウド環境にノード作成などを行うために必要となります
# - VCCアクセストークンがない場合はVC管理者に発行を依頼してください
# * SSH公開鍵ペア
# - VCノードに登録するSSHの公開鍵
# - このNotebook環境内で新たに作成するか、事前に作成したものをこの環境にアップロードしておいてください
# * VCノードに割り当てるアドレス
# - ノードのネットワークインタフェースに以下に示す何れかのアドレスを指定することができます
# - IPアドレス
# - MACアドレス
# * NTPの設定
# - 学認フェデレーションに参加し SAML 認証を利用する場合、正しい時刻設定が必要となります
# - VCノードのNTPサービスを有効にするためには、事前にVCコントローラへの設定が必要となります
# - VCコントローラへの設定にはOCS運用担当者への申請が必要となります
# ### CoursewareHub
#
# CoursewareHub環境を構築する際に必要となるものについて記します。
# * CoursewareHubのサーバ証明書
# - CoursewareHubではHTTPSでサーバを公開するため、サーバ証明書とその秘密鍵が必要となります
# - 必要に応じて、サーバ証明書の中間CA証明書を準備してください
# - サーバ証明書に記載されているホスト名のDNS登録も必要となります
#
# また事前の段階では不要ですが、学認のIdPを認証に利用する場合は構築手順の過程で
# 学認フェデレーションに参加の申請を行う必要があります。
# ### IdP-proxy
#
# IdP-proxy を構築する際に必要となるものについて記します。
# * IdP-proxyのサーバ証明書
# - IdP-proxyではHTTPSでサーバを公開するため、サーバ証明書とその秘密鍵が必要となります
# - 必要に応じて、サーバ証明書の中間CA証明書を準備してください
# - サーバ証明書に記載されているホスト名のDNS登録も必要となります
#
# また事前の段階では不要ですが、学認のIdPを認証に利用する場合は構築手順の過程で
# 学認フェデレーションに参加の申請を行う必要があります。
# ## Notebookの一覧
#
# テンプレートのNotebook一覧を示します。
# **注意**:
#
# この節ではテンプレートのNotebookへのリンクを示す箇所がありますが、リンク先のNotebookは参照用となっていて**そのままでは実行できません**。
#
# > Notebook自体は実行できてしまいますが、パスなどが想定しているものと異なるため正しく処理できずエラーとなります。
#
# 次のどちらかの手順で作業用Notebookを作成する必要があります。
#
# * 次節の「作業用Notebookの作成」で作業用のNotebookを作成する。
# * テンプレートのNotebookを配置してある `notebooks/` から、この`000-README.ipynb`と同じディレクトリにNotebookをコピーする。
# ### 各Notebookの関連について
#
# 各Notebookの実行順序などの関係性を示す図を表示します。
# 次のセルを実行すると、各Notebookの関連を示す図を表示します。
#
# > 図が表示されずに `<IPython.core.display.SVG object>` と表示されている場合は、次のセルを `unfreeze` した後に再実行してください。
#
# 図に表示される1つのブロックが1つのNotebookに対応しており、ブロックのタイトル部分にNotebookへのリンクが埋め込まれています。
# #### ローカルユーザ認証のみを利用する場合
from IPython.display import SVG
# %run scripts/nb_utils.py
setup_diag()
SVG(filename=generate_svg_diag(diag='images/notebooks-a.diag'))
# #### IdP-proxyを利用する場合
from IPython.display import SVG
# %run scripts/nb_utils.py
setup_diag()
SVG(filename=generate_svg_diag(diag='images/notebooks-b.diag'))
# #### CoursewareHubを直接SPとして登録する場合
#
from IPython.display import SVG
# %run scripts/nb_utils.py
setup_diag()
SVG(filename=generate_svg_diag(diag='images/notebooks-c.diag'))
# ### 各Notebookの目次
# 次のセルを実行すると、各Notebookの目次が表示されます。
#
# > 目次が表示されずに `<IPython.core.display.Markdown object>` と表示されている場合は、次のセルを `unfreeze` した後に再実行してください。
#
# リンクが表示されている項目が一つのNotebookに対応しており、そのサブ項目が各Notebook内の目次になっています。
from IPython.display import Markdown
# %run scripts/nb_utils.py
Markdown(notebooks_toc())
# ## 作業用Notebookの作成
#
# この節のセルを実行することで、テンプレートのNotebookから作業用Notebookを作成することができます。
# まず、作業用Notebookを配置するディレクトリを指定してください。
WORK_DIR = 'work'
# 以下のセルを実行すると、Notebook名のドロップダウンリストと「作業開始」ボタンが現れます。
# 「作業開始」ボタンを押すと、テンプレートのNotebookをコピーし、そのNotebookを自動的にブラウザで開きます。
# Notebookの説明を確認しながら実行、適宜修正しながら実行していってください。
#
# > このNotebookを Shutdown した後に再度開いた場合、次のセルに既に表示されている「作用開始」ボタンが正しく動作しません。この節のセルをいったん unfreeze した後、セルを再実行してから「作業開始」ボタンをクリックして下さい。
#
# 認証方式毎に実行するNotebookが異なるので、対応する節のセルを実行してください。
# ### ローカルユーザ認証のみを利用する場合
from IPython.core.display import HTML
# %run scripts/nb_utils.py
setup_nb_workdir(WORK_DIR)
HTML(generate_html_work_nbs(WORK_DIR, nb_group='group-a'))
# ### IdP-proxyを利用する場合
# #### IdP-proxyの構築
from IPython.core.display import HTML
# %run scripts/nb_utils.py
setup_nb_workdir(WORK_DIR)
HTML(generate_html_work_nbs(WORK_DIR, nb_group='group-b0'))
# #### CoursewareHubの構築
from IPython.core.display import HTML
# %run scripts/nb_utils.py
setup_nb_workdir(WORK_DIR)
HTML(generate_html_work_nbs(WORK_DIR, nb_group='group-b1'))
# ### CoursewareHubを直接SPとして登録する場合
from IPython.core.display import HTML
# %run scripts/nb_utils.py
setup_nb_workdir(WORK_DIR)
HTML(generate_html_work_nbs(WORK_DIR, nb_group='group-c'))
| CoursewareHub/000-README.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Regularization
#
# One of the goals of a machine learning project is to make models which are highly predictive.
# If the model fails to generalize to unseen data then the model is bad
#
#
# ## Review of Earlier concepts
#
# ### Overfitting vs Underfitting
# 1. Underfit Models fail to capture all of the information in the data
# 1. Ex. the mean leaves a lot of information on the table
# 2. Overfit models fit to the noise in the data and fail to generalize
# 3. How would we know if our model is over or underfit?
# 1. Train test split
# 2. Look at the testing error
# 3. As model complexity increases so does the possibility for overfitting
#
# ### Bias variance tradeoff
# 1. High bias
# 1. Systematic error in predictions (ie the average)
# 2. Bias is about the strength of assumptions the model makes
# 3. Underfit models tend to be high bias
# 2. High variance
# 1. The model is highly sensitive to changes in the data
# 2. Overfit models tend to be low bias
# ### Regularization - a way to prevent over fitting
# 1. Types of regularization
# 1. Reducing the amount of features
# 2. Increase the amount of data
# 3. Ridge, lasso, elastic net
#
# Again, complex models are very flexible in the patterns that they can model but this also means that they can easily find patterns that are simply statistical flukes of one particular dataset rather than patterns reflective of the underlying data generating process.
# ## Ridge and Lasso Regression
from sklearn.preprocessing import StandardScaler, PolynomialFeatures
from sklearn.linear_model import Ridge, Lasso, ElasticNet, LinearRegression
from sklearn.model_selection import train_test_split
import pandas as pd
from sklearn.metrics import mean_squared_error
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
ads = pd.read_csv('Advertising.csv', index_col=0)
ads.head()
X_train, X_test, y_train, y_test = train_test_split(ads[['TV',
'radio',
'newspaper']],
ads['sales'])
# ### First simple model
# +
lr1 = LinearRegression()
lr1.fit(X_train, y_train)
ads_preds = lr1.predict(X_test)
np.sqrt(mean_squared_error(ads_preds, y_test))
# -
sns.pairplot(ads);
# ### Polynomial Features
# +
# Try and create your own polynomial features with degree 2 in sklearn and create a linear model
# Answer is at the bottom
# -
# ### Ridge Regression
# +
ss = StandardScaler()
pf = PolynomialFeatures()
X_train_processed = pf.fit_transform(ss.fit_transform(X_train))
X_test_processed = pf.transform(ss.transform(X_test))
#alpha == lambda since lamda is a key word in python
rr = Ridge(alpha=1)
rr.fit(X_train_processed, y_train)
ridge_preds = rr.predict(X_test_processed)
np.sqrt(mean_squared_error(ridge_preds, y_test))
# -
# ### Why did I pick an alpha of 1? Is there a principled way to choose this hyperparamter?
#
# Create many models on the training set and see which one does best on unseen data
# ## Crossvalidation
#
# How can we know what values of lamda to choose for ridge or lasso regression? Crossvalidation!
#
# We have already discussed the train test split and that is the basis for all other kinds of crossvalidation. We can extend this
# +
from sklearn.model_selection import cross_val_score
cross_val_score()
# -
# ## Try to find the optimal value for alpha/lamda for lasso regression
# ## Appendix
# +
pf = PolynomialFeatures()
X_poly_train = pf.fit_transform(X_train[['TV', 'radio', 'newspaper']])
pf_sales = pd.DataFrame(X_poly, columns=pf.get_feature_names(['TV', 'radio', 'newspaper']))
# -
pf_y = pd.concat([pf_sales[['TV', 'radio', 'newspaper', 'TV radio',
'TV newspaper']],
ads['sales']], axis=1)
# +
poly_lr = LinearRegression()
poly_lr.fit(X_poly_train, y_train)
X_poly_test = pf.transform(X_test[['TV', 'radio', 'newspaper']])
poly_preds = poly_lr.predict(X_poly_test)
np.sqrt(mean_squared_error(poly_preds, y_test))
# -
list(zip(pf_sales.columns, poly_lr.coef_))
sns.pairplot(pf_y);
| regularization_crossvalidation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] _uuid="281275d20b3c5f659236d880031e3da4b7977e06"
# # Import Data
# + _uuid="ebb7ec8cdfe9b840f5b5ba1524f4e892994e46cc"
import pandas as pd
import numpy as np
import os
import gc
# + _uuid="713132d2f045501c82e42290550d356ae0727c35"
DATA_ROOT = '../input/'
ORIGINAL_DATA_FOLDER = os.path.join(DATA_ROOT, 'movie-review-sentiment-analysis-kernels-only')
TMP_DATA_FOLDER = os.path.join(DATA_ROOT, 'kaggle_review_sentiment_tmp_data')
# + _uuid="98fd884ce5265326d434498bda1f5bb813f24e95"
train_data_path = os.path.join(ORIGINAL_DATA_FOLDER, 'train.tsv')
test_data_path = os.path.join(ORIGINAL_DATA_FOLDER, 'test.tsv')
sub_data_path = os.path.join(ORIGINAL_DATA_FOLDER, 'sampleSubmission.csv')
train_df = pd.read_csv(train_data_path, sep="\t")
test_df = pd.read_csv(test_data_path, sep="\t")
sub_df = pd.read_csv(sub_data_path, sep=",")
# + [markdown] _uuid="ee07dfaceeb2d800b4c95430a5f44d7386fed119"
# # EDA
# + _uuid="abc33cf6801467c6c8ccd5266961d5e74a70b030"
import seaborn as sns
# + _uuid="33485dab7ac4678cc77fed13b2c8e500d341f72f"
train_df.head()
# + _uuid="c2e0e484aacbb21ebb2807c79beca5aa054c3830"
test_df.head()
# + _uuid="641a74568482c8ad42d1ac1c8982a44b84641166"
sub_df.head()
# + [markdown] _uuid="c4d3c60c638b8ed36a90102e2f5d0fcb034b9ac4"
# ## Find Overlapped Phrases Between Train and Test Data
# + _uuid="0a31536ca50b348303c4980a7535fdea69716d6e"
overlapped = pd.merge(train_df[["Phrase", "Sentiment"]], test_df, on="Phrase", how="inner")
overlap_boolean_mask_test = test_df['Phrase'].isin(overlapped['Phrase'])
# + [markdown] _uuid="2de53b6baefcd985866f4e3789cb923e7c8d57c8"
# ## Histogram of phrase length
# + _uuid="25a101f200f79abef0c91b2fdce721a22ad66b1d"
print("training data phrase length distribution")
sns.distplot(train_df['Phrase'].map(lambda ele: len(ele)), kde_kws={"label": "train"})
print("testing data phrase length distribution")
sns.distplot(test_df[~overlap_boolean_mask_test]['Phrase'].map(lambda ele: len(ele)), kde_kws={"label": "test"})
# + [markdown] _uuid="22c203439a2310d54f0283dc4e117307a1afa82d"
# ## Explore Sentence Id
# + _uuid="c957e257b2865c258a6d9ce23b9ed15c316b90b8"
print("training and testing data sentences hist:")
sns.distplot(train_df['SentenceId'], kde_kws={"label": "train"})
sns.distplot(test_df['SentenceId'], kde_kws={"label": "test"})
# + _uuid="6ae6308dd0bb586f7cc6e75619e228f7984116ab"
print("The number of overlapped SentenceId between training and testing data:")
train_overlapped_sentence_id_df = train_df[train_df['SentenceId'].isin(test_df['SentenceId'])]
print(train_overlapped_sentence_id_df.shape[0])
del train_overlapped_sentence_id_df
gc.collect()
# + _kg_hide-output=true _uuid="030ab4a34391188a30160731259e21d4611762a2"
pd.options.display.max_colwidth = 250
print("Example of sentence and phrases: ")
sample_sentence_id = train_df.sample(1)['SentenceId'].values[0]
sample_sentence_group_df = train_df[train_df['SentenceId'] == sample_sentence_id]
sample_sentence_group_df
# + [markdown] _uuid="<KEY>"
# 1. There are overlapped phrase texts between training and testing data, which should assign training data labels directly instead of getting from prediction.
# 2. Max text length should be set around 60.
# 3. There is no overlapped sentence between training and testing data. Within each sentence group, the phraseId order is the pre-order tanversal over the dependency parsing tree of the sentence text. (This might be a very important information as we can utilized the composition as powerful predictive information).
#
#
# + [markdown] _uuid="09c94783b2b9f99c5d62ea251391642329ec46a6"
# # Data Preprocessing
# + _uuid="f15786c0c632835ae7a008114319badd92dce4c8"
from keras.preprocessing import text
from keras.preprocessing import sequence
import gensim
from sklearn import preprocessing as skp
# + _uuid="b50c845cca35a18a4f2986d03b37eb826c17e109"
max_len = 50
embed_size = 300
max_features = 30000
pretrained_w2v_path = os.path.join(DATA_ROOT, "nlpword2vecembeddingspretrained/GoogleNews-vectors-negative300.bin")
# + [markdown] _uuid="69e929bce14433c86cda53d35abc91a3b3fd4e72"
# ### Tokenize Text
# + _uuid="f05a0bcf40751a8e33814fd3314b7f7e1d5149e5"
full_text = list(train_df['Phrase'].values) + list(test_df[~overlap_boolean_mask_test]['Phrase'].values)
tk = text.Tokenizer(lower = True, filters='')
tk.fit_on_texts(full_text)
train_tokenized = tk.texts_to_sequences(train_df['Phrase'])
test_tokenized = tk.texts_to_sequences(test_df[~overlap_boolean_mask_test]['Phrase'])
X_train = sequence.pad_sequences(train_tokenized, maxlen = max_len)
X_test = sequence.pad_sequences(test_tokenized, maxlen = max_len)
# + [markdown] _uuid="d42d7f7ad3c09838e1648beb5743dbf4e313a543"
# ### Build embedding matrix
# + _uuid="3467eb40c92d5490a26629f6ab6c7505b8439979"
w2v = gensim.models.KeyedVectors.load_word2vec_format(pretrained_w2v_path, binary=True).wv
word_index = tk.word_index
nb_words = min(max_features, len(word_index))
embedding_matrix = np.zeros((nb_words + 1, embed_size))
for word, i in word_index.items():
if i >= max_features: continue
embedding_vector = None
if word in w2v:
embedding_vector = w2v[word]
if embedding_vector is not None: embedding_matrix[i] = embedding_vector
del w2v
gc.collect()
# + [markdown] _uuid="26e06e8aeabd698b0fe2fb208f2c4ec2855babcd"
# ### Encode labels
# + _uuid="57a0df2d2e98b1b1a8edc5ef87dd26f4f1f5d63c"
y_train = train_df['Sentiment']
led = skp.LabelEncoder()
led.fit(y_train.values)
y_train = led.transform(y_train.values)
# + [markdown] _uuid="4675184968f46ed87fbd703696978dc3535ce89e"
# # Define Keras Model
# + _uuid="391b187f5f79d60fc329dd8a805708ac71937b5b"
import tensorflow as tf
from keras import callbacks as kc
from keras import optimizers as ko
from keras import initializers, regularizers, constraints
from keras.engine import Layer
import keras.backend as K
# + [markdown] _uuid="413a0fcd52787baf4cb702cc1074b8086675b1ba"
# ## Define Attention Layer
# + _uuid="e59457ba4f41bd4082188ccacff7f1cf8601545c"
def _dot_product(x, kernel):
"""
Wrapper for dot product operation, in order to be compatible with both
Theano and Tensorflow
Args:
x (): input
kernel (): weights
Returns:
"""
if K.backend() == 'tensorflow':
# todo: check that this is correct
return K.squeeze(K.dot(x, K.expand_dims(kernel)), axis=-1)
else:
return K.dot(x, kernel)
class AttentionWeight(Layer):
"""
This code is a modified version of cbaziotis implementation: GithubGist cbaziotis/AttentionWithContext.py
Attention operation, with a context/query vector, for temporal data.
Supports Masking.
Follows the work of Yang et al. [https://www.cs.cmu.edu/~diyiy/docs/naacl16.pdf]
"Hierarchical Attention Networks for Document Classification"
by using a context vector to assist the attention
# Input shape
3D tensor with shape: `(samples, steps, features)`.
# Output shape
2D tensor with shape: `(samples, steps)`.
:param kwargs:
Just put it on top of an RNN Layer (GRU/LSTM/SimpleRNN) with return_sequences=True.
The dimensions are inferred based on the output shape of the RNN.
Example:
model.add(LSTM(64, return_sequences=True))
model.add(AttentionWeight())
"""
def __init__(self,
W_regularizer=None, u_regularizer=None, b_regularizer=None,
W_constraint=None, u_constraint=None, b_constraint=None,
bias=True, **kwargs):
self.supports_masking = True
self.init = initializers.get('glorot_uniform')
self.W_regularizer = regularizers.get(W_regularizer)
self.u_regularizer = regularizers.get(u_regularizer)
self.b_regularizer = regularizers.get(b_regularizer)
self.W_constraint = constraints.get(W_constraint)
self.u_constraint = constraints.get(u_constraint)
self.b_constraint = constraints.get(b_constraint)
self.bias = bias
super(AttentionWeight, self).__init__(**kwargs)
def build(self, input_shape):
assert len(input_shape) == 3
self.W = self.add_weight((input_shape[-1], input_shape[-1],),
initializer=self.init,
name='{}_W'.format(self.name),
regularizer=self.W_regularizer,
constraint=self.W_constraint)
if self.bias:
self.b = self.add_weight((input_shape[-1],),
initializer='zero',
name='{}_b'.format(self.name),
regularizer=self.b_regularizer,
constraint=self.b_constraint)
self.u = self.add_weight((input_shape[-1],),
initializer=self.init,
name='{}_u'.format(self.name),
regularizer=self.u_regularizer,
constraint=self.u_constraint)
super(AttentionWeight, self).build(input_shape)
def compute_mask(self, input, input_mask=None):
# do not pass the mask to the next layers
return None
def call(self, x, mask=None):
uit = _dot_product(x, self.W)
if self.bias:
uit += self.b
uit = K.tanh(uit)
ait = _dot_product(uit, self.u)
a = K.exp(ait)
# apply mask after the exp. will be re-normalized next
if mask is not None:
# Cast the mask to floatX to avoid float64 upcasting in theano
a *= K.cast(mask, K.floatx())
# in some cases especially in the early stages of training the sum may be almost zero
# and this results in NaN's. A workaround is to add a very small positive number ε to the sum.
# a /= K.cast(K.sum(a, axis=1, keepdims=True), K.floatx())
a /= K.cast(K.sum(a, axis=1, keepdims=True) + K.epsilon(), K.floatx())
return a
def compute_output_shape(self, input_shape):
return input_shape[0], input_shape[1]
def get_config(self):
config = {
'W_regularizer': regularizers.serialize(self.W_regularizer),
'u_regularizer': regularizers.serialize(self.u_regularizer),
'b_regularizer': regularizers.serialize(self.b_regularizer),
'W_constraint': constraints.serialize(self.W_constraint),
'u_constraint': constraints.serialize(self.u_constraint),
'b_constraint': constraints.serialize(self.b_constraint),
'bias': self.bias
}
base_config = super(AttentionWeight, self).get_config()
return dict(list(base_config.items()) + list(config.items()))
# + [markdown] _uuid="c3e4ae86f2e94275e3bc3de33a5eb9b1f8327caa"
# ## Define Models
# + _uuid="ab6fd1ea52e5cdeb110d5ef79e0cde12b195c586"
def is_integer(val):
return isinstance(val, (int, np.int_))
def predict(keras_model, x, learning_phase=0):
if isinstance(keras_model.input, list):
f = backend.function(
keras_model.input + [backend.learning_phase()],
[keras_model.output, ]
)
y = f(tuple(x) + (learning_phase,))[0]
else:
f = backend.function(
[keras_model.input, backend.learning_phase()],
[keras_model.output, ]
)
y = f((x, learning_phase))[0]
return y
def build_birnn_attention_model(
voca_dim, time_steps, output_dim, rnn_dim, mlp_dim,
item_embedding=None, rnn_depth=1, mlp_depth=1, num_att_channel=1,
drop_out=0.5, rnn_drop_out=0., rnn_state_drop_out=0.,
trainable_embedding=False, gpu=False, return_customized_layers=False):
"""
Create A Bidirectional Attention Model.
:param voca_dim: vocabulary dimension size.
:param time_steps: the length of input
:param output_dim: the output dimension size
:param rnn_dim: rrn dimension size
:param mlp_dim: the dimension size of fully connected layer
:param item_embedding: integer, numpy 2D array, or None (default=None)
If item_embedding is a integer, connect a randomly initialized embedding matrix to the input tensor.
If item_embedding is a matrix, this matrix will be used as the embedding matrix.
If item_embedding is None, then connect input tensor to RNN layer directly.
:param rnn_depth: rnn depth
:param mlp_depth: the depth of fully connected layers
:param num_att_channel: the number of attention channels, this can be used to mimic multi-head attention mechanism
:param drop_out: dropout rate of fully connected layers
:param rnn_drop_out: dropout rate of rnn layers
:param rnn_state_drop_out: dropout rate of rnn state tensor
:param trainable_embedding: boolean
:param gpu: boolean, default=False
If True, CuDNNLSTM is used instead of LSTM for RNN layer.
:param return_customized_layers: boolean, default=False
If True, return model and customized object dictionary, otherwise return model only
:return: keras model
"""
if item_embedding is not None:
inputs = models.Input(shape=(time_steps,), dtype='int32', name='input0')
x = inputs
# item embedding
if isinstance(item_embedding, np.ndarray):
assert voca_dim == item_embedding.shape[0]
x = layers.Embedding(
voca_dim, item_embedding.shape[1], input_length=time_steps,
weights=[item_embedding, ], trainable=trainable_embedding,
mask_zero=False, name='embedding_layer0'
)(x)
elif utils.is_integer(item_embedding):
x = layers.Embedding(
voca_dim, item_embedding, input_length=time_steps,
trainable=trainable_embedding,
mask_zero=False, name='embedding_layer0'
)(x)
else:
raise ValueError("item_embedding must be either integer or numpy matrix")
else:
inputs = models.Input(shape=(time_steps, voca_dim), dtype='float32', name='input0')
x = inputs
x = layers.SpatialDropout1D(rnn_drop_out, name='rnn_spatial_droutout_layer')(x)
if gpu:
# rnn encoding
for i in range(rnn_depth):
x = layers.Bidirectional(
layers.CuDNNLSTM(rnn_dim, return_sequences=True),
name='bi_lstm_layer' + str(i))(x)
x = layers.BatchNormalization(name='rnn_batch_norm_layer' + str(i))(x)
x = layers.Dropout(rate=rnn_drop_out, name="rnn_dropout_layer" + str(i))(x)
else:
# rnn encoding
for i in range(rnn_depth):
x = layers.Bidirectional(
layers.LSTM(rnn_dim, return_sequences=True, dropout=rnn_drop_out, recurrent_dropout=rnn_state_drop_out),
name='bi_lstm_layer' + str(i))(x)
x = layers.BatchNormalization(name='rnn_batch_norm_layer' + str(i))(x)
# attention
attention_heads = []
x_per = layers.Permute((2, 1), name='permuted_attention_x')(x)
for h in range(max(1, num_att_channel)):
attention = AttentionWeight(name="attention_weights_layer" + str(h))(x)
xx = layers.Dot([2, 1], name='focus_head' + str(h) + '_layer0')([x_per, attention])
attention_heads.append(xx)
if num_att_channel > 1:
x = layers.Concatenate(name='focus_layer0')(attention_heads)
else:
x = attention_heads[0]
x = layers.BatchNormalization(name='focused_batch_norm_layer')(x)
x = layers.Dropout(rate=rnn_drop_out, name="focused_dropout_layer")(x)
# MLP Layers
for i in range(mlp_depth - 1):
x = layers.Dense(mlp_dim, activation='selu', kernel_initializer='lecun_normal', name='selu_layer' + str(i))(x)
x = layers.AlphaDropout(drop_out, name='alpha_layer' + str(i))(x)
outputs = layers.Dense(output_dim, activation="softmax", name="softmax_layer0")(x)
model = models.Model(inputs, outputs)
if return_customized_layers:
return model, {'AttentionWeight': AttentionWeight}
return model
def build_cnn_model(
voca_dim, time_steps, output_dim, mlp_dim, num_filters, filter_sizes,
item_embedding=None, mlp_depth=1,
drop_out=0.5, cnn_drop_out=0.5, pooling='max', padding='valid',
trainable_embedding=False, return_customized_layers=False):
"""
Create A CNN Model.
:param voca_dim: vocabulary dimension size.
:param time_steps: the length of input
:param output_dim: the output dimension size
:param num_filters: list of integers
The number of filters.
:param filter_sizes: list of integers
The kernel size.
:param mlp_dim: the dimension size of fully connected layer
:param item_embedding: integer, numpy 2D array, or None (default=None)
If item_embedding is a integer, connect a randomly initialized embedding matrix to the input tensor.
If item_embedding is a matrix, this matrix will be used as the embedding matrix.
If item_embedding is None, then connect input tensor to RNN layer directly.
:param mlp_depth: the depth of fully connected layers
:param drop_out: dropout rate of fully connected layers
:param cnn_drop_out: dropout rate of between cnn layer and fully connected layers
:param pooling: str, either 'max' or 'average'
Pooling method.
:param padding: One of "valid", "causal" or "same" (case-insensitive).
Padding method.
:param trainable_embedding: boolean
:param return_customized_layers: boolean, default=False
If True, return model and customized object dictionary, otherwise return model only
:return: keras model
"""
if item_embedding is not None:
inputs = models.Input(shape=(time_steps,), dtype='int32', name='input0')
x = inputs
# item embedding
if isinstance(item_embedding, np.ndarray):
assert voca_dim == item_embedding.shape[0]
x = layers.Embedding(
voca_dim, item_embedding.shape[1], input_length=time_steps,
weights=[item_embedding, ], trainable=trainable_embedding,
mask_zero=False, name='embedding_layer0'
)(x)
elif utils.is_integer(item_embedding):
x = layers.Embedding(
voca_dim, item_embedding, input_length=time_steps,
trainable=trainable_embedding,
mask_zero=False, name='embedding_layer0'
)(x)
else:
raise ValueError("item_embedding must be either integer or numpy matrix")
else:
inputs = models.Input(shape=(time_steps, voca_dim), dtype='float32', name='input0')
x = inputs
x = layers.SpatialDropout1D(cnn_drop_out, name='cnn_spatial_droutout_layer')(x)
pooled_outputs = []
for i in range(len(filter_sizes)):
conv = layers.Conv1D(num_filters[i], kernel_size=filter_sizes[i], padding=padding, activation='relu')(x)
if pooling == 'max':
conv = layers.GlobalMaxPooling1D(name='global_pooling_layer' + str(i))(conv)
else:
conv = layers.GlobalAveragePooling1D(name='global_pooling_layer' + str(i))(conv)
pooled_outputs.append(conv)
x = layers.Concatenate(name='concated_layer')(pooled_outputs)
x = layers.Dropout(cnn_drop_out, name='conv_dropout_layer')(x)
x = layers.BatchNormalization(name="batch_norm_layer")(x)
# MLP Layers
for i in range(mlp_depth - 1):
x = layers.Dense(mlp_dim, activation='selu', kernel_initializer='lecun_normal', name='selu_layer' + str(i))(x)
x = layers.AlphaDropout(drop_out, name='alpha_layer' + str(i))(x)
outputs = layers.Dense(output_dim, activation="softmax", name="softmax_layer0")(x)
model = models.Model(inputs, outputs)
if return_customized_layers:
return model, dict()
return model
def build_birnn_cnn_model(
voca_dim, time_steps, output_dim, rnn_dim, mlp_dim, num_filters, filter_sizes,
item_embedding=None, rnn_depth=1, mlp_depth=1,
drop_out=0.5, rnn_drop_out=0.5, rnn_state_drop_out=0.5, cnn_drop_out=0.5, pooling='max', padding='valid',
trainable_embedding=False, gpu=False, return_customized_layers=False):
"""
Create A Bidirectional CNN Model.
:param voca_dim: vocabulary dimension size.
:param time_steps: the length of input
:param output_dim: the output dimension size
:param rnn_dim: rrn dimension size
:param num_filters: list of integers
The number of filters.
:param filter_sizes: list of integers
The kernel size.
:param mlp_dim: the dimension size of fully connected layer
:param item_embedding: integer, numpy 2D array, or None (default=None)
If item_embedding is a integer, connect a randomly initialized embedding matrix to the input tensor.
If item_embedding is a matrix, this matrix will be used as the embedding matrix.
If item_embedding is None, then connect input tensor to RNN layer directly.
:param rnn_depth: rnn depth
:param mlp_depth: the depth of fully connected layers
:param num_att_channel: the number of attention channels, this can be used to mimic multi-head attention mechanism
:param drop_out: dropout rate of fully connected layers
:param rnn_drop_out: dropout rate of rnn layers
:param rnn_state_drop_out: dropout rate of rnn state tensor
:param cnn_drop_out: dropout rate of between cnn layer and fully connected layers
:param pooling: str, either 'max' or 'average'
Pooling method.
:param padding: One of "valid", "causal" or "same" (case-insensitive).
Padding method.
:param trainable_embedding: boolean
:param gpu: boolean, default=False
If True, CuDNNLSTM is used instead of LSTM for RNN layer.
:param return_customized_layers: boolean, default=False
If True, return model and customized object dictionary, otherwise return model only
:return: keras model
"""
if item_embedding is not None:
inputs = models.Input(shape=(time_steps,), dtype='int32', name='input0')
x = inputs
# item embedding
if isinstance(item_embedding, np.ndarray):
assert voca_dim == item_embedding.shape[0]
x = layers.Embedding(
voca_dim, item_embedding.shape[1], input_length=time_steps,
weights=[item_embedding, ], trainable=trainable_embedding,
mask_zero=False, name='embedding_layer0'
)(x)
elif utils.is_integer(item_embedding):
x = layers.Embedding(
voca_dim, item_embedding, input_length=time_steps,
trainable=trainable_embedding,
mask_zero=False, name='embedding_layer0'
)(x)
else:
raise ValueError("item_embedding must be either integer or numpy matrix")
else:
inputs = models.Input(shape=(time_steps, voca_dim), dtype='float32', name='input0')
x = inputs
x = layers.SpatialDropout1D(rnn_drop_out, name='rnn_spatial_droutout_layer')(x)
if gpu:
# rnn encoding
for i in range(rnn_depth):
x = layers.Bidirectional(
layers.CuDNNLSTM(rnn_dim, return_sequences=True),
name='bi_lstm_layer' + str(i))(x)
x = layers.BatchNormalization(name='rnn_batch_norm_layer' + str(i))(x)
x = layers.Dropout(rate=rnn_drop_out, name="rnn_dropout_layer" + str(i))(x)
else:
# rnn encoding
for i in range(rnn_depth):
x = layers.Bidirectional(
layers.LSTM(rnn_dim, return_sequences=True, dropout=rnn_drop_out, recurrent_dropout=rnn_state_drop_out),
name='bi_lstm_layer' + str(i))(x)
x = layers.BatchNormalization(name='rnn_batch_norm_layer' + str(i))(x)
pooled_outputs = []
for i in range(len(filter_sizes)):
conv = layers.Conv1D(num_filters[i], kernel_size=filter_sizes[i], padding=padding, activation='relu')(x)
if pooling == 'max':
conv = layers.GlobalMaxPooling1D(name='global_pooling_layer' + str(i))(conv)
else:
conv = layers.GlobalAveragePooling1D(name='global_pooling_layer' + str(i))(conv)
pooled_outputs.append(conv)
x = layers.Concatenate(name='concated_layer')(pooled_outputs)
x = layers.BatchNormalization(name="batch_norm_layer")(x)
x = layers.Dropout(cnn_drop_out, name='conv_dropout_layer')(x)
# MLP Layers
for i in range(mlp_depth - 1):
x = layers.Dense(mlp_dim, activation='selu', kernel_initializer='lecun_normal', name='selu_layer' + str(i))(x)
x = layers.AlphaDropout(drop_out, name='alpha_layer' + str(i))(x)
outputs = layers.Dense(output_dim, activation="softmax", name="softmax_layer0")(x)
model = models.Model(inputs, outputs)
if return_customized_layers:
return model, dict()
return model
def build_birnn_hierarchy_cnn_model(
voca_dim, time_steps, output_dim, rnn_dim, mlp_dim, num_filters, filter_sizes,
dilation_rates=1, strides=1,
item_embedding=None, rnn_depth=1, mlp_depth=1,
drop_out=0.5, rnn_drop_out=0.5, rnn_state_drop_out=0.5, cnn_drop_out=0.5, pooling='max', padding='valid',
trainable_embedding=False, gpu=False, return_customized_layers=False):
"""
Create A Bidirectional CNN Model.
:param voca_dim: vocabulary dimension size.
:param time_steps: the length of input
:param output_dim: the output dimension size
:param rnn_dim: rrn dimension size
:param num_filters: list of integers
The number of filters.
:param filter_sizes: list of integers
The kernel size.
:param mlp_dim: the dimension size of fully connected layer
:param item_embedding: integer, numpy 2D array, or None (default=None)
If item_embedding is a integer, connect a randomly initialized embedding matrix to the input tensor.
If item_embedding is a matrix, this matrix will be used as the embedding matrix.
If item_embedding is None, then connect input tensor to RNN layer directly.
:param rnn_depth: rnn depth
:param mlp_depth: the depth of fully connected layers
:param num_att_channel: the number of attention channels, this can be used to mimic multi-head attention mechanism
:param drop_out: dropout rate of fully connected layers
:param rnn_drop_out: dropout rate of rnn layers
:param rnn_state_drop_out: dropout rate of rnn state tensor
:param cnn_drop_out: dropout rate of between cnn layer and fully connected layers
:param pooling: str, either 'max' or 'average'
Pooling method.
:param padding: One of "valid", "causal" or "same" (case-insensitive).
Padding method.
:param trainable_embedding: boolean
:param gpu: boolean, default=False
If True, CuDNNLSTM is used instead of LSTM for RNN layer.
:param return_customized_layers: boolean, default=False
If True, return model and customized object dictionary, otherwise return model only
:return: keras model
"""
if item_embedding is not None:
inputs = models.Input(shape=(time_steps,), dtype='int32', name='input0')
x = inputs
# item embedding
if isinstance(item_embedding, np.ndarray):
assert voca_dim == item_embedding.shape[0]
x = layers.Embedding(
voca_dim, item_embedding.shape[1], input_length=time_steps,
weights=[item_embedding, ], trainable=trainable_embedding,
mask_zero=False, name='embedding_layer0'
)(x)
elif utils.is_integer(item_embedding):
x = layers.Embedding(
voca_dim, item_embedding, input_length=time_steps,
trainable=trainable_embedding,
mask_zero=False, name='embedding_layer0'
)(x)
else:
raise ValueError("item_embedding must be either integer or numpy matrix")
else:
inputs = models.Input(shape=(time_steps, voca_dim), dtype='float32', name='input0')
x = inputs
x = layers.SpatialDropout1D(rnn_drop_out, name='rnn_spatial_droutout_layer')(x)
if gpu:
# rnn encoding
for i in range(rnn_depth):
x = layers.Bidirectional(
layers.CuDNNLSTM(rnn_dim, return_sequences=True),
name='bi_lstm_layer' + str(i))(x)
x = layers.BatchNormalization(name='rnn_batch_norm_layer' + str(i))(x)
x = layers.Dropout(rate=rnn_drop_out, name="rnn_dropout_layer" + str(i))(x)
else:
# rnn encoding
for i in range(rnn_depth):
x = layers.Bidirectional(
layers.LSTM(rnn_dim, return_sequences=True, dropout=rnn_drop_out, recurrent_dropout=rnn_state_drop_out),
name='bi_lstm_layer' + str(i))(x)
x = layers.BatchNormalization(name='rnn_batch_norm_layer' + str(i))(x)
for i in range(len(filter_sizes)):
if is_integer(dilation_rates):
di_rate = dilation_rates
else:
di_rate = dilation_rates[i]
if is_integer(strides):
std = strides
else:
std = strides[i]
x = layers.Conv1D(num_filters[i], kernel_size=filter_sizes[i], padding=padding, activation='relu', dilation_rate=di_rate, strides=std)(x)
if pooling == 'max':
x = layers.GlobalMaxPooling1D(name='global_pooling_layer')(x)
else:
x = layers.GlobalAveragePooling1D(name='global_pooling_layer')(x)
x = layers.BatchNormalization(name="batch_norm_layer")(x)
x = layers.Dropout(cnn_drop_out, name='conv_dropout_layer')(x)
# MLP Layers
for i in range(mlp_depth - 1):
x = layers.Dense(mlp_dim, activation='selu', kernel_initializer='lecun_normal', name='selu_layer' + str(i))(x)
x = layers.AlphaDropout(drop_out, name='alpha_layer' + str(i))(x)
outputs = layers.Dense(output_dim, activation="softmax", name="softmax_layer0")(x)
model = models.Model(inputs, outputs)
if return_customized_layers:
return model, dict()
return model
# + [markdown] _uuid="0f8a54b45eb4838d7459e6db57e9f0a95bfe7e60"
# # Build and Train Models
# + _uuid="9dd9a276408dd3543d44e4e82cd8086024f56b3f"
from keras.utils import model_to_dot
from keras import models
from keras import layers
import matplotlib.pyplot as plt
from IPython.display import SVG
# + _uuid="83515918a1861d9aa3ef7751aaf05ca1635d9620"
histories = list()
iterations = list()
model_builders = list()
# + [markdown] _uuid="d514af1525ce6dd02614bfdc4c1c707617eceade"
# ## CNN Model
# + _uuid="9fcdab9f1bfb6d88b2e3d110dea35f87c71dbf51"
def build_model1():
voca_dim = embedding_matrix.shape[0]
time_steps = max_len
output_dim = led.classes_.shape[0]
mlp_dim = 50
num_filters = [128, 128, 128]
filter_sizes = [1, 3, 5]
item_embedding = embedding_matrix
mlp_depth = 2
cnn_drop_out = 0.2
mlp_drop_out = 0.2
padding = 'causal'
return build_cnn_model(
voca_dim, time_steps, output_dim, mlp_dim, num_filters, filter_sizes,
item_embedding=item_embedding, mlp_depth=2, cnn_drop_out=cnn_drop_out,
padding=padding,
return_customized_layers=True
)
model_builders.append(build_model1)
# + _uuid="f5e58b1f129140130e4559b1610c307d50f15f8d"
model, cnn_cl = build_model1()
print(model.summary())
# + _uuid="82b16272ecfe86ee0dc3e5731d4e33aa016842c3"
adam = ko.Nadam()
model.compile(adam, loss="sparse_categorical_crossentropy", metrics=["sparse_categorical_accuracy",])
file_path = "best_cnn_model.hdf5"
check_point = kc.ModelCheckpoint(file_path, monitor = "val_sparse_categorical_accuracy", verbose = 1, save_best_only = True, mode = "max")
early_stop = kc.EarlyStopping(monitor = "val_sparse_categorical_accuracy", mode = "max", patience=3)
history = model.fit(X_train, y_train, batch_size=500, epochs=20, validation_split=0.1, callbacks = [check_point, early_stop])
histories.append(np.max(np.asarray(history.history['val_sparse_categorical_accuracy'])))
iterations.append(np.argmax(np.asarray(history.history['val_sparse_categorical_accuracy'])))
del model, history
gc.collect()
# + [markdown] _uuid="db9ac105bd787c06a1371eaf61936b1159c1645c"
# ## Attention RNN Model
# + _uuid="46d854b13ad591f5d349af3007a0b17559f72535"
def build_model2():
voca_dim = embedding_matrix.shape[0]
time_steps = max_len
output_dim = led.classes_.shape[0]
rnn_dim = 100
mlp_dim = 50
item_embedding = embedding_matrix
rnn_depth=1
mlp_depth = 2
rnn_drop_out = 0.3
rnn_state_drop_out = 0.3
mlp_drop_out = 0.2
num_att_channel = 1
gpu=True
return build_birnn_attention_model(
voca_dim, time_steps, output_dim, rnn_dim, mlp_dim,
item_embedding=item_embedding, rnn_depth=rnn_depth, mlp_depth=mlp_depth, num_att_channel=num_att_channel,
rnn_drop_out=rnn_drop_out, rnn_state_drop_out=rnn_state_drop_out,
gpu=gpu, return_customized_layers=True
)
model_builders.append(build_model2)
# + _uuid="2a264d510a763478c67b572c14366ae2144ce3f8"
model, rnn_cl = build_model2()
print(model.summary())
# + _uuid="d40857a43116f8587f28ba89baef17b519c58d45"
adam = ko.Nadam(clipnorm=2.0)
model.compile(adam, loss="sparse_categorical_crossentropy", metrics=["sparse_categorical_accuracy",])
file_path = "best_birnn_attention_model.hdf5"
check_point = kc.ModelCheckpoint(file_path, monitor = "val_sparse_categorical_accuracy", verbose = 1, save_best_only = True, mode = "max")
early_stop = kc.EarlyStopping(monitor = "val_sparse_categorical_accuracy", mode = "max", patience=3)
history = model.fit(X_train, y_train, batch_size=500, epochs=20, validation_split=0.1, callbacks = [check_point, early_stop])
histories.append(np.max(np.asarray(history.history['val_sparse_categorical_accuracy'])))
iterations.append(np.argmax(np.asarray(history.history['val_sparse_categorical_accuracy'])))
del model, history
gc.collect()
# + [markdown] _uuid="e53b77f6a8507bc64a2de060a1718366ee24200f"
# ## RNN-CNN Model
# + _uuid="1a3294e306d5ac35c0bf0fa039fc17099a83f367"
def build_model3():
voca_dim = embedding_matrix.shape[0]
time_steps = max_len
output_dim = led.classes_.shape[0]
rnn_dim = 100
mlp_dim = 50
item_embedding = embedding_matrix
rnn_depth=1
mlp_depth = 2
num_filters = [128, 128, 128]
filter_sizes = [1, 3, 5]
cnn_drop_out = 0.2
rnn_drop_out = 0.3
rnn_state_drop_out = 0.3
mlp_drop_out = 0.2
padding = 'causal'
gpu=True
return build_birnn_cnn_model(
voca_dim, time_steps, output_dim, rnn_dim, mlp_dim, num_filters, filter_sizes,
item_embedding=item_embedding, rnn_depth=rnn_depth, mlp_depth=mlp_depth,
rnn_drop_out=rnn_drop_out, rnn_state_drop_out=rnn_state_drop_out, cnn_drop_out=cnn_drop_out,
padding=padding,
gpu=gpu, return_customized_layers=True
)
model_builders.append(build_model3)
# + _uuid="c9e49dcf31d32f1b4717a3ccf665aacad1a1e165"
model, rc_cl = build_model3()
print(model.summary())
# + _uuid="39f08fded4a3019dfd07504da932064425f8a16b"
adam = ko.Nadam(clipnorm=2.0)
model.compile(adam, loss="sparse_categorical_crossentropy", metrics=["sparse_categorical_accuracy",])
file_path = "best_birnn_cnn_model.hdf5"
check_point = kc.ModelCheckpoint(file_path, monitor = "val_sparse_categorical_accuracy", verbose = 1, save_best_only = True, mode = "max")
early_stop = kc.EarlyStopping(monitor = "val_sparse_categorical_accuracy", mode = "max", patience=3)
history = model.fit(X_train, y_train, batch_size=500, epochs=20, validation_split=0.1, callbacks = [check_point, early_stop])
histories.append(np.max(np.asarray(history.history['val_sparse_categorical_accuracy'])))
iterations.append(np.argmax(np.asarray(history.history['val_sparse_categorical_accuracy'])))
del model, history
gc.collect()
# + [markdown] _uuid="b4165b5741a971bb4a651a9d513ee2e1d3c19aa5"
# ## RNN-HierarchyCNN
# + _uuid="0ee00f133652fa813b55f4f223b87beddb588bcf"
def build_model4():
voca_dim = embedding_matrix.shape[0]
time_steps = max_len
output_dim = led.classes_.shape[0]
rnn_dim = 100
mlp_dim = 50
item_embedding = embedding_matrix
rnn_depth=1
mlp_depth = 2
num_filters = [128, 256, 512]
filter_sizes = [1, 3, 5]
dilation_rates = [1, 2, 4]
strides=1
cnn_drop_out = 0.2
rnn_drop_out = 0.3
rnn_state_drop_out = 0.3
mlp_drop_out = 0.2
padding = 'causal'
gpu=True
return build_birnn_hierarchy_cnn_model(
voca_dim, time_steps, output_dim, rnn_dim, mlp_dim, num_filters, filter_sizes,
dilation_rates=dilation_rates, strides=strides,
item_embedding=item_embedding, rnn_depth=rnn_depth, mlp_depth=mlp_depth,
rnn_drop_out=rnn_drop_out, rnn_state_drop_out=rnn_state_drop_out, cnn_drop_out=cnn_drop_out,
padding=padding,
gpu=gpu, return_customized_layers=True
)
model_builders.append(build_model4)
# + _uuid="d85a1e00009b635853e8928c83b3733a32f37d48"
model, rhc_cl = build_model4()
print(model.summary())
# + _uuid="276d7133a37091457cdead0efb311ce01e20f689"
adam = ko.Nadam(clipnorm=2.0)
model.compile(adam, loss="sparse_categorical_crossentropy", metrics=["sparse_categorical_accuracy",])
file_path = "best_birnn_hierarchy_cnn_model.hdf5"
check_point = kc.ModelCheckpoint(file_path, monitor = "val_sparse_categorical_accuracy", verbose = 1, save_best_only = True, mode = "max")
early_stop = kc.EarlyStopping(monitor = "val_sparse_categorical_accuracy", mode = "max", patience=3)
history = model.fit(X_train, y_train, batch_size=500, epochs=20, validation_split=0.1, callbacks = [check_point, early_stop])
histories.append(np.max(np.asarray(history.history['val_sparse_categorical_accuracy'])))
iterations.append(np.argmax(np.asarray(history.history['val_sparse_categorical_accuracy'])))
del model, history
gc.collect()
# + [markdown] _uuid="db9a896a94311439b6eda66aa247e828fb57eca2"
# # Make Prediction
# + _uuid="428880b633b909db6d5056d4751ce3dabc9fd1a5"
histories = np.asarray(histories)
model_paths = [
"best_cnn_model.hdf5",
"best_birnn_attention_model.hdf5",
"best_birnn_cnn_model.hdf5",
"best_birnn_hierarchy_cnn_model.hdf5"
]
cls =[
cnn_cl, rnn_cl, rc_cl, rhc_cl
]
pred = list()
for idx in range(len(model_paths)):
model = models.load_model(model_paths[idx], cls[idx])
pred_tmp = model.predict(X_test, batch_size = 1024, verbose = 1)
pred.append(np.round(np.argmax(pred_tmp, axis=1)).astype(int))
# + _uuid="3ef083a0a29b808560344a0d52d1fb5a77a3a2da"
pred = [
np.asarray([1, 2, 3, 4]),
np.asarray([0, 3, 5, 5]),
np.asarray([1, 2, 2, 4])
]
def majority_vote(preds_data_point):
unique, counts = np.unique(preds_data_point, return_counts=True)
idx = np.argmax(counts)
return unique[idx]
pred = np.asarray(pred)
predictions = list()
for i in range(pred.shape[1]):
predictions.append(majority_vote(pred[:, i]))
predictions = np.asarray(predictions)
test_not_overlap_df = test_df[~overlap_boolean_mask_test]
test_not_overlap_df['Sentiment'] = predictions
res_df = pd.concat([overlapped, test_not_overlap_df], sort=True)[sub_df.columns.values.tolist()]
assert sub_df.shape[0] == res_df.shape[0]
assert sub_df.shape[1] == res_df.shape[1]
res_df.to_csv("submission.csv", index=False)
| question_classification_v0/main_notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
#
#
# (c) <NAME> <br />
# Team Neighborhood Change <br />
# Last Updated 9/15/2016 <br />
#
# # NOTEBOOK: DBSCAN with PCA and interpolate features
# * This is to chose the best feature selecton method and model selection method
#
# +
####################
## DEPENDENCIES ####
####################
import pandas as pd
import os
import csv
import xlrd
import numpy as np
import matplotlib
import seaborn as sns
import matplotlib.pyplot as plt
from pandas.tools.plotting import scatter_matrix
from __future__ import print_function
from sklearn.decomposition import PCA
from sklearn.feature_selection import SelectFromModel
from sklearn.linear_model import Ridge, Lasso, ElasticNet
from sklearn.decomposition import PCA
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis as LDA
from sklearn.preprocessing import Imputer
from sklearn import preprocessing
matplotlib.style.use('ggplot')
from sklearn.cluster import KMeans
from sklearn.cluster import MiniBatchKMeans
## http://stats.stackexchange.com/questions/89809/is-it-important-to-scale-data-before-clustering
# above says that scaling makes for better
from time import time
import numpy as np
import matplotlib.pyplot as plt
import numpy as np
from sklearn.cluster import DBSCAN
from sklearn import metrics
from sklearn.datasets.samples_generator import make_blobs
from sklearn.preprocessing import StandardScaler
from sklearn import metrics
from sklearn.cluster import KMeans
from sklearn.datasets import load_digits
from sklearn.decomposition import PCA
from sklearn.preprocessing import scale
from sklearn.preprocessing import StandardScaler, MinMaxScaler
# %matplotlib inline
# -
df = pd.read_csv('indicators.csv')
indicators = df[['cbsa', 'msa_name', 'violent_crime_rate',
'murder_manslaughter',
'rape', 'robbery',
'aggravated_assault',
'property_crime_rate',
'burglary', 'larceny_theft',
'motor_vehicle_theft',
'total_crime_rate',
'median_gross_rent',
'median_monthly_mortgage',
'rent_burden', 'mortgage_burden',
'income_change_2012_to_2014',
'median_age_of_men', 'median_age_of_women',
'median_age', 'median_household_income',
'single_men_population',
'single_women_population',
'ratio_of_single_men_to_single_women',
'population_percent_of_single_men',
'population_percent_of_single_women',
'population', 'edu_average_scale_score',
'pct_laccess_pop10', 'pct_laccess_lowi10',
'pct_laccess_child10', 'pct_laccess_seniors10',
'pct_laccess_hhnv10', 'event_mpmt',
'fatalities_mpmt', 'injuries_mpmt',
'walk_score', 'transit_score', 'bike_score',
'unemploymentrate', 'employment', 'laborforce']]
# +
## all Nans to white space
indicators = indicators.replace(np.nan,' ', regex=True)
indicators = indicators.replace(np.nan,'NaN', regex=True)
# convert to all floats except cbsa and msa_name
indicators = indicators.convert_objects(convert_numeric=True)
##imputer
# -
indicators.columns = ['cbsa', 'msa_name', 'violent_crime_rate',
'murder_manslaughter',
'rape', 'robbery',
'aggravated_assault',
'property_crime_rate',
'burglary', 'larceny_theft',
'motor_vehicle_theft',
'total_crime_rate',
'median_gross_rent',
'median_monthly_mortgage',
'rent_burden', 'mortgage_burden',
'income_change_2012_to_2014',
'median_age_of_men', 'median_age_of_women',
'median_age', 'median_household_income',
'single_men_population',
'single_women_population',
'ratio_of_single_men_to_single_women',
'population_percent_of_single_men',
'population_percent_of_single_women',
'population', 'edu_average_scale_score',
'pct_laccess_pop10', 'pct_laccess_lowi10',
'pct_laccess_child10', 'pct_laccess_seniors10',
'pct_laccess_hhnv10', 'event_mpmt',
'fatalities_mpmt', 'injuries_mpmt',
'walk_score', 'transit_score', 'bike_score',
'unemploymentrate', 'employment',
'laborforce']
# +
featuresx = df[['violent_crime_rate',
'murder_manslaughter',
'rape', 'robbery',
'aggravated_assault',
'property_crime_rate',
'burglary', 'larceny_theft',
'motor_vehicle_theft',
'total_crime_rate',
'median_gross_rent',
'median_monthly_mortgage',
'rent_burden', 'mortgage_burden',
'income_change_2012_to_2014',
'median_age_of_men', 'median_age_of_women',
'median_age', 'median_household_income',
'single_men_population',
'single_women_population',
'ratio_of_single_men_to_single_women',
'population_percent_of_single_men',
'population_percent_of_single_women',
'population', 'edu_average_scale_score',
'pct_laccess_pop10', 'pct_laccess_lowi10',
'pct_laccess_child10', 'pct_laccess_seniors10',
'pct_laccess_hhnv10', 'event_mpmt',
'fatalities_mpmt', 'injuries_mpmt',
'walk_score', 'transit_score', 'bike_score',
'unemploymentrate', 'employment', 'laborforce']]
labelsx = df["cbsa"]
# -
# Dealing with Missing Data
## all Nans to white space
featuresx = featuresx.replace(np.nan,' ', regex=True)
featuresx = featuresx.replace(np.nan,'NaN', regex=True)
##convert to all floats
featuresx = featuresx.convert_objects(convert_numeric=True)
##imputer
featuresx = featuresx.interpolate()
featuresx.columns = ['violen t_crime_rate', 'murder_manslaughter', 'rape', 'robbery', 'aggravated_assault', 'property_crime_rate', 'burglary', 'larceny_theft', 'motor_vehicle_theft', 'total_crime_rate', 'median_gross_rent', 'median_monthly_mortgage', 'rent_burden', 'mortgage_burden', 'income_change_2012_to_2014', 'median_age_of_men', 'median_age_of_women', 'median_age', 'median_household_income', 'single_men_population', 'single_women_population', 'ratio_of_single_men_to_single_women', 'population_percent_of_single_men', 'population_percent_of_single_women', 'population', 'edu_average_scale_score', 'pct_laccess_pop10', 'pct_laccess_lowi10', 'pct_laccess_child10', 'pct_laccess_seniors10', 'pct_laccess_hhnv10', 'event_mpmt', 'fatalities_mpmt', 'injuries_mpmt', 'walk_score', 'transit_score', 'bike_score', 'unemploymentrate', 'employment', 'laborforce']
# #### DBSCAN
# Trial
# +
# Compute DBSCAN
stscaler = StandardScaler().fit(featuresx)
featuresx = stscaler.transform(featuresx)
featuresx = pd.DataFrame(featuresx)
pca = PCA(n_components = 23)
pca_features = pca.fit(featuresx).transform(featuresx)
pca_features = pd.DataFrame(pca_features)
features = featuresx.values
#DBSCAN
db = DBSCAN(eps = 3.4, min_samples = 10).fit(features)
core_samples_mask = np.zeros_like(db.labels_, dtype=bool)
core_samples_mask[db.core_sample_indices_] = True
labels = db.labels_
# Number of clusters in labels, ignoring noise if present.
n_clusters_ = len(set(labels)) - (1 if -1 in labels else 0)
print('Estimated number of clusters: %d' % n_clusters_)
print("Homogeneity: %0.3f" % metrics.homogeneity_score(labelsx, labels))
print("Completeness: %0.3f" % metrics.completeness_score(labelsx, labels))
print("V-measure: %0.3f" % metrics.v_measure_score(labelsx, labels))
print("Adjusted Rand Index: %0.3f"
% metrics.adjusted_rand_score(labelsx, labels))
print("Adjusted Mutual Information: %0.3f"
% metrics.adjusted_mutual_info_score(labelsx, labels))
print("Silhouette Coefficient: %0.3f"
% metrics.silhouette_score(features, labels))
##############################################################################
# Plot result
import matplotlib.pyplot as plt
# Black removed and is used for noise instead.
unique_labels = set(labels)
colors = plt.cm.Spectral(np.linspace(0, 1, len(unique_labels)))
for k, col in zip(unique_labels, colors):
if k == -1:
# Black used for noise.
col = 'k'
class_member_mask = (labels == k)
xy = features[class_member_mask & core_samples_mask]
plt.plot(xy[:, 0], xy[:, 1], 'o', markerfacecolor=col,
markeredgecolor='k', markersize=14)
xy = features[class_member_mask & ~core_samples_mask]
plt.plot(xy[:, 0], xy[:, 1], 'o', markerfacecolor=col,
markeredgecolor='k', markersize=6)
plt.title('Estimated number of clusters: %d' % n_clusters_)
plt.show()
# -
# # LOOP to try different eps & min_samples
# +
# Compute DBSCAN
import numpy as np
pca = PCA(n_components = 23)
pca_features = pca.fit(featuresx).transform(featuresx)
pca_features = pd.DataFrame(pca_features)
features = featuresx.values
#DBSCAN
range_of_m_samples= range(5,40)
for m in range_of_m_samples:
range_of_eps = np.arange(5.1, 20.5, 0.5)
for e in range_of_eps:
print ('min samples = %d' % (m))
print ('eps : %0.1f' % (e))
db = DBSCAN(eps = e, min_samples = m).fit(features)
core_samples_mask = np.zeros_like(db.labels_, dtype=bool)
core_samples_mask[db.core_sample_indices_] = True
labels = db.labels_
# Number of clusters in labels, ignoring noise if present.
n_clusters_ = len(set(labels)) - (1 if -1 in labels else 0)
try:
print('Estimated number of clusters: %d' % n_clusters_)
print("Homogeneity: %0.3f" % metrics.homogeneity_score(labelsx, labels))
print("Completeness: %0.3f" % metrics.completeness_score(labelsx, labels))
print("V-measure: %0.3f" % metrics.v_measure_score(labelsx, labels))
print("Adjusted Rand Index: %0.3f"
% metrics.adjusted_rand_score(labelsx, labels))
print("Adjusted Mutual Information: %0.3f"
% metrics.adjusted_mutual_info_score(labelsx, labels))
print("Silhouette Coefficient: %0.3f" % metrics.silhouette_score(features, labels))
##############################################################################
# Plot result
import matplotlib.pyplot as plt
# Black removed and is used for noise instead.
unique_labels = set(labels)
colors = plt.cm.Spectral(np.linspace(0, 1, len(unique_labels)))
for k, col in zip(unique_labels, colors):
if k == -1:
# Black used for noise.
col = 'k'
class_member_mask = (labels == k)
xy = features[class_member_mask & core_samples_mask]
plt.plot(xy[:, 0], xy[:, 1], 'o', markerfacecolor=col,
markeredgecolor='k', markersize=14)
xy = features[class_member_mask & ~core_samples_mask]
plt.plot(xy[:, 0], xy[:, 1], 'o', markerfacecolor=col,
markeredgecolor='k', markersize=6)
plt.title('Estimated number of clusters: %d' % n_clusters_)
plt.show()
except ValueError:
pass
# -
| ml-application/ml-unsupervised-part2c-dbscan.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="Y9LyDe0e9n-r" colab_type="code" outputId="94854b84-8fdf-454b-a5b6-2e69f1b2564e" colab={"base_uri": "https://localhost:8080/", "height": 373}
# %matplotlib inline
# !pip install category_encoders
from google.colab import drive
import os
import time
import pandas as pd
import numpy as np
from matplotlib import pyplot as plt
import pickle
drive.mount('/content/drive')
os.chdir("/content/drive/My Drive/Colab Notebooks/AML")
pd.set_option('display.max_rows', 20)
# + id="sWBKK_WT9ucL" colab_type="code" colab={}
import tensorflow as tf
from sklearn.datasets import load_digits
from sklearn.model_selection import train_test_split, StratifiedShuffleSplit
from sklearn.model_selection import GridSearchCV
from keras.wrappers.scikit_learn import KerasClassifier
from keras.regularizers import l2
from keras.utils import to_categorical
from keras.layers import Dense, BatchNormalization, Dropout, Activation
from keras.models import Sequential
from keras.datasets import fashion_mnist
# + id="FwLO9DRb9w75" colab_type="code" outputId="c398285c-f15a-4f83-daa1-b021d017b6fe" colab={"base_uri": "https://localhost:8080/", "height": 151}
tmp=fashion_mnist.load_data()
# + id="o3p5Xp_tXMab" colab_type="code" colab={}
(X_train, y_train) = tmp[0]
# + id="63PspHJ_Xk7q" colab_type="code" colab={}
(X_test, y_test) = tmp[1]
# + id="gsGsg2rPXonW" colab_type="code" outputId="9a8114b1-09b4-4f99-c605-148bcf7f32f5" colab={"base_uri": "https://localhost:8080/", "height": 34}
y_train.shape
# + id="fk9ZdlzeYE2P" colab_type="code" colab={}
y_test_n=to_categorical(y_test, cls)
y_train_n=to_categorical(y_train, cls)
# + id="b4zNto1jaeeh" colab_type="code" colab={}
X_train=X_train.reshape(X_train.shape[0], X_train.shape[1]*X_train.shape[2])
X_train=X_train.astype('float64')
X_train=X_train/255
# + id="6lKrw7KmamkS" colab_type="code" colab={}
X_test=X_test.reshape(X_test.shape[0], X_test.shape[1]*X_test.shape[2])
X_test=X_test.astype('float64')
X_test=X_test/255
# + id="UP29cIGkapJw" colab_type="code" outputId="afc36b76-d4e0-47bb-f56d-034ee989474e" colab={"base_uri": "https://localhost:8080/", "height": 34}
cls=len(np.unique(y_train))
print(cls)
# + id="A1zWPZ5XaxRJ" colab_type="code" outputId="ebf365ae-2307-4c9d-a97e-cbc4ce656705" colab={"base_uri": "https://localhost:8080/", "height": 34}
sss = StratifiedShuffleSplit(n_splits=1, test_size=10000, random_state=1)
sss.get_n_splits(X_train, y_train_n)
# + id="6gVn0tGtbSTL" colab_type="code" colab={}
for train_index, test_index in sss.split(X_train, y_train_n):
X_train_new, X_val_new = X_train[train_index], X_train[test_index]
y_train_new_n, y_val_new_n = y_train_n[train_index], y_train_n[test_index]
# + id="iD9csQivch3Q" colab_type="code" colab={}
inp=X_train_new[0].shape
# + [markdown] id="z_RJ16O_FXv0" colab_type="text"
# Baseline dense model
# + id="RxGOFiGdcUlP" colab_type="code" colab={}
def build_model(neurons=32, l2_val=0.1):
inp=X_train[0].shape
model=Sequential([Dense(neurons,activation='relu',kernel_regularizer=l2(l=l2_val), input_shape=inp),Dense(neurons,activation='relu',kernel_regularizer=l2(l=l2_val)), Dense(cls,activation='softmax',kernel_regularizer=l2(l=l2_val)) ])
model.compile(optimizer='adam',loss='categorical_crossentropy', metrics=['accuracy'])
return model
# + id="dNaQMEy6ccoD" colab_type="code" colab={}
model_mlp1=build_model(neurons=64,l2_val= 0.001)
# + id="dAHMfdATfb0l" colab_type="code" outputId="2069fd12-106a-4bc0-d8b5-c511f9b10e2e" colab={"base_uri": "https://localhost:8080/", "height": 1000}
l_curve=model_mlp1.fit(X_train_new, y_train_new_n, epochs=50, validation_data=(X_val_new, y_val_new_n))
# + id="ULnhk8vLgZ5w" colab_type="code" colab={}
arr=l_curve.history
df=pd.DataFrame(arr)
# + id="EmNelahGirTB" colab_type="code" outputId="6d6dda35-703e-4948-d8cf-f37d002a0f79" colab={"base_uri": "https://localhost:8080/", "height": 195}
df.head()
# + id="cElDq0kpjqUZ" colab_type="code" outputId="98b8629c-efa1-4e5e-a145-8ec5300b1e7f" colab={"base_uri": "https://localhost:8080/", "height": 312}
plt.plot(df['val_accuracy'], label='validation accuracy')
plt.plot(df['accuracy'], label='accuracy')
plt.ylabel('accuracy score')
plt.xlabel('no of epochs')
plt.title('Accuracy scores vs epochs')
plt.legend(loc="lower right")
plt.figure(figsize=(60,10))
plt.show()
# + [markdown] id="0OLEbSshFR7w" colab_type="text"
# Dense model with only dropout
# + id="PUvyI46wk_vI" colab_type="code" colab={}
def build_model2(neurons=32, l2_val=0.1):
inp=X_train[0].shape
model=Sequential([Dense(neurons,activation='relu',kernel_regularizer=l2(l=l2_val), input_shape=inp),Dropout(0.4), Dense(neurons,activation='relu',kernel_regularizer=l2(l=l2_val)), Dropout(0.4), Dense(cls,activation='softmax',kernel_regularizer=l2(l=l2_val)) ])
model.compile(optimizer='adam',loss='categorical_crossentropy', metrics=['accuracy'])
return model
# + id="nM9mYCAYtCp-" colab_type="code" colab={}
model_mlp2=build_model2(neurons=64,l2_val= 0.001)
# + id="ZxVxN05XtJt-" colab_type="code" outputId="8ad5d9a8-6ad2-4d3b-e97f-003c8b0e9de4" colab={"base_uri": "https://localhost:8080/", "height": 1000}
l_curve2=model_mlp2.fit(X_train_new, y_train_new_n, epochs=50, validation_data=(X_val_new, y_val_new_n))
# + id="GhSQJFT9uRdc" colab_type="code" outputId="6da9b6aa-dd1c-4674-9ddb-75c5c0e05241" colab={"base_uri": "https://localhost:8080/", "height": 50}
model_mlp2.evaluate(X_val_new, y_val_new_n)
# + id="JLrjU-YHur4H" colab_type="code" colab={}
arr=l_curve2.history
df=pd.DataFrame(arr)
# + id="igoLILqcuzhA" colab_type="code" outputId="64cde78c-b276-42a1-a661-782e9847bdaa" colab={"base_uri": "https://localhost:8080/", "height": 312}
plt.plot(df['val_accuracy'], label='validation accuracy')
plt.plot(df['accuracy'], label='accuracy')
plt.ylabel('accuracy score')
plt.xlabel('no of epochs')
plt.title('Accuracy scores vs epochs')
plt.legend(loc="lower right")
plt.figure(figsize=(60,10))
plt.show()
# + [markdown] id="8O8GI5UoFJKY" colab_type="text"
# Trying another network architecture with Batch Norm and dropout
# + id="DLBtKiEyu7rC" colab_type="code" colab={}
def build_model3(neurons=32, l2_val=0.1):
inp=X_train[0].shape
model=Sequential([Dense(neurons,activation='relu',kernel_regularizer=l2(l=l2_val), input_shape=inp),Dropout(0.4), Dense(neurons,activation='relu',kernel_regularizer=l2(l=l2_val)), Dropout(0.4), Dense(neurons,activation='relu',kernel_regularizer=l2(l=l2_val)), Dropout(0.4),Dense(cls,activation='softmax',kernel_regularizer=l2(l=l2_val)) ])
model.compile(optimizer='adam',loss='categorical_crossentropy', metrics=['accuracy'])
return model
# + id="FgKOKrjuvH0g" colab_type="code" colab={}
model_mlp3=build_model3(neurons=512,l2_val= 0.001)
# + id="nD3ZsjJXvOp5" colab_type="code" outputId="f6452d38-9b85-4cff-bcb1-778721c7c415" colab={"base_uri": "https://localhost:8080/", "height": 386}
model_mlp3.summary()
# + id="r1PHRCh2vjKQ" colab_type="code" outputId="8644a96e-737b-4939-f65f-8ae34317edf3" colab={"base_uri": "https://localhost:8080/", "height": 1000}
l_curve2=model_mlp3.fit(X_train_new, y_train_new_n, epochs=50, validation_data=(X_val_new, y_val_new_n))
# + id="AtNjFthevvIR" colab_type="code" outputId="c69369e8-1c6c-4607-9284-b53c67ae2061" colab={"base_uri": "https://localhost:8080/", "height": 50}
model_mlp3.evaluate(X_val_new, y_val_new_n)
# + id="6b6XFLqzvnTl" colab_type="code" outputId="539d30e0-2284-4818-e8a7-68a4689bdac5" colab={"base_uri": "https://localhost:8080/", "height": 312}
arr=l_curve2.history
df=pd.DataFrame(arr)
plt.plot(df['val_accuracy'], label='validation accuracy')
plt.plot(df['accuracy'], label='accuracy')
plt.ylabel('accuracy score')
plt.xlabel('no of epochs')
plt.title('Accuracy scores vs epochs')
plt.legend(loc="lower right")
plt.figure(figsize=(60,10))
plt.show()
# + [markdown] id="g8KdaYaIE6by" colab_type="text"
# Using Batch normalization, dropout layers on the neural net
# + id="hurFsCV_v3Qy" colab_type="code" colab={}
def build_modelb1(neurons=32, l2_val=0.1):
inp=X_train[0].shape
model=Sequential([Dense(neurons,kernel_regularizer=l2(l=l2_val), input_shape=inp),BatchNormalization(), Activation('relu') ,Dropout(0.4), Dense(neurons,kernel_regularizer=l2(l=l2_val)), BatchNormalization(), Activation('relu'), Dropout(0.4), Dense(cls,activation='softmax',kernel_regularizer=l2(l=l2_val)) ])
model.compile(optimizer='adam',loss='categorical_crossentropy', metrics=['accuracy'])
return model
# + id="TzihRdnfyUIf" colab_type="code" colab={}
model_mlp4=build_modelb1(neurons=64,l2_val= 0.001)
# + id="_OXBmzngyZ6x" colab_type="code" outputId="d11823ac-103c-414c-f2e1-f8b8eb5f9db4" colab={"base_uri": "https://localhost:8080/", "height": 1000}
l_curve2=model_mlp4.fit(X_train_new, y_train_new_n, epochs=50, validation_data=(X_val_new, y_val_new_n))
# + id="nZA7b9Sty0We" colab_type="code" outputId="7aed953d-0251-4c13-bfa5-99b98fc9f6a5" colab={"base_uri": "https://localhost:8080/", "height": 50}
model_mlp4.evaluate(X_val_new, y_val_new_n)
# + id="Zhz-GTqMy32V" colab_type="code" outputId="a7799980-33c0-4a94-ea3f-666ff9fde744" colab={"base_uri": "https://localhost:8080/", "height": 312}
arr=l_curve2.history
df=pd.DataFrame(arr)
plt.plot(df['val_accuracy'], label='validation accuracy')
plt.plot(df['accuracy'], label='accuracy')
plt.ylabel('accuracy score')
plt.xlabel('no of epochs')
plt.title('Accuracy scores vs epochs')
plt.legend(loc="lower right")
plt.figure(figsize=(60,10))
plt.show()
# + [markdown] id="imFQVZ87EvR2" colab_type="text"
# Applying Batch Normalization and Dropout on a bigger network
# + id="h7NYrRkOzPJP" colab_type="code" colab={}
def build_modelb2(neurons=32, l2_val=0.1):
inp=X_train[0].shape
model=Sequential([Dense(neurons,kernel_regularizer=l2(l=l2_val), input_shape=inp),BatchNormalization(), Activation('relu') ,Dropout(0.4), Dense(neurons,kernel_regularizer=l2(l=l2_val)), BatchNormalization(), Activation('relu'), Dropout(0.4), Dense(neurons,kernel_regularizer=l2(l=l2_val)), BatchNormalization(), Activation('relu'), Dropout(0.4), Dense(cls,activation='softmax',kernel_regularizer=l2(l=l2_val)) ])
model.compile(optimizer='adam',loss='categorical_crossentropy', metrics=['accuracy'])
return model
# + id="vat2weku0sBc" colab_type="code" colab={}
model_mlp5=build_modelb2(neurons=512,l2_val= 0.001)
# + id="_vk4tP0S0xWg" colab_type="code" outputId="917b722d-a3e8-4fee-f9e1-9e5383dbf8cf" colab={"base_uri": "https://localhost:8080/", "height": 1000}
l_curve2=model_mlp5.fit(X_train_new, y_train_new_n, epochs=100, validation_data=(X_val_new, y_val_new_n))
# + id="suJHqqB800w1" colab_type="code" outputId="8dcc3766-1122-40db-b659-5de176752c27" colab={"base_uri": "https://localhost:8080/", "height": 50}
model_mlp5.evaluate(X_val_new, y_val_new_n)
# + id="wsck9_k605DE" colab_type="code" outputId="a7bfdcff-c596-4530-eeab-dbb4c7e0558e" colab={"base_uri": "https://localhost:8080/", "height": 312}
arr=l_curve2.history
df=pd.DataFrame(arr)
plt.plot(df['val_accuracy'], label='validation accuracy')
plt.plot(df['accuracy'], label='accuracy')
plt.ylabel('accuracy score')
plt.xlabel('no of epochs')
plt.title('Accuracy scores vs epochs')
plt.legend(loc="lower right")
plt.figure(figsize=(60,10))
plt.show()
# + [markdown] id="CBMO-SYYEMGS" colab_type="text"
# On the basis of the learning curves and evaluation scores on the validation set, we see that all the models provide ~ 0.85 on the validation set, so we can pick the bigger network with dropout and batch normalization as our model of choice, as it would be more robust to generalizing to other datasets.
| Keras Neural Networks/Deep Neural Net with Batch Norm and Dropout.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Шаблоны. Продвинутый материал
# [Modern Template Techniques - <NAME> - Meeting C++ 2019](https://youtu.be/MLV4IVc4SwI)
# <br />
# ##### Повторение основ
# __Вопрос__: зачем нужны шаблоны?
# __Вопрос__: что может быть параметром шаблона?
# Код для повторения:
#
# ```c++
# template<typename T>
# void my_swap(T& x, T& y)
# {
# T t = std::move(x);
# x = std::move(y);
# y = std::move(t);
# }
#
# template<>
# void my_swap<string>(string& x, string& y)
# {
# x.swap(y);
# }
# ```
# <br />
# Шаблонный класс:
#
# ```c++
# template<typename T>
# struct Point3
# {
# T x;
# T y;
# T z;
# };
#
# using Point3F = Point3<float>;
# using Point3D = Point3<double>;
# ```
# Или так:
#
# ```c++
# template<typename T, int N>
# struct PointN
# {
# T data[N];
# };
#
# using Point3F = PointN<float, 3>;
# using Point3D = PointN<double, 3>;
# ```
# __Вопрос__: что такое частичная специализация? К чему её можно применять?
# <br />
# __Вопрос__: скомпилируется ли этот код?
#
# ```c++
# template<typename T>
# class Person
# {
# private:
# T id;
#
# public:
# void set_id(T i_id) { id = std::move(i_id); }
#
# const T& get_id() const { return id; }
#
# void clear_id() { id.clear(); }
# };
#
# int main()
# {
# Person<int> p;
# p.set_id(15);
# std::cout << p.get_id() << std::endl;
# }
# ```
# <br />
# __Вопрос__: как устроена компиляция и линковка шаблонов?
# <br />
# ##### SFINAE
# https://en.cppreference.com/w/cpp/language/sfinae
#
# https://en.cppreference.com/w/cpp/language/overload_resolution
#
# https://jguegant.github.io/blogs/tech/sfinae-introduction.html
#
# https://www.bfilipek.com/2016/02/notes-on-c-sfinae.html
# Рассмотрим пример, как через SFINAE реализовать `my_swap` так, чтобы версия с методом была вызвана для всех типов, у которых есть соответствующий метод, а не только для тех, для которых не забыли сделать специализацию:
#
# ```c++
# template<typename T>
# void my_swap(T& x, T& y)
# {
# T t = std::move(x);
# x = std::move(y);
# y = std::move(t);
# }
#
# template<>
# void my_swap<string>(string& x, string& y)
# {
# x.swap(y);
# }
# ```
# SFINAE = Substitution Failure Is Not An Error
#
# SFINAE - ovreloads resolution правило при наличии шаблонов: если не получается провести подстановку / сделать вывод шаблонных параметров, не генерировать ошибку компиляции, а продолжить искать другую подходящую перегрузку
# Если речь идёт про SFINAE, то должны участвовать:
# * шаблоны
# * перегрузки
# * поиск нужной перегрузки
# Необходимые отступления:
#
# * _substitution_ происходит в аргументах функции, возвращаемом типе и параметрах шаблона
#
# ```c++
# template<HERE>
# HERE my_function(HERE x, HERE y)
# { ...; }
# ```
#
# * _substitution failure_ - ситуация, когда в результате подстановки в HERE получается ill-formed выражение
# * если ill-formed случился не в местах подстановки, а, например, в теле функции после подстановки, то это не случай SFINAE, а обычный hard error
# * если ill-formed случился как side-effect подстановки, то это тоже hard error (что считать side-effect-ом, что точно является sfinae error - подробнее читайте документацию):
#
# ```c++
# struct Point1 {
# float x, y, z;
# };
#
# struct Point2 {
# float x, y, z;
# using type = float;
# };
#
#
# // ex. 1
# template <class T, class = typename T::type> // SFINAE failure if T has no member type
# void foo(T);
#
# // ex. 2
# template <typename T>
# struct B {
# using type = typename T::type;
# };
#
# template <class T, class = typename T::type> // SFINAE failure if T has no member type
# void foo(T);
#
# template <class T, class = typename S<T>::type> // hard error if B<T> has no member type
# void foo (T);
# ```
# Демонстрация sfinae с cppreference (разобрать подробно):
#
# ```c++
# #include <iostream>
#
# // this overload is always in the set of overloads
# // ellipsis parameter has the lowest ranking for overload resolution
# void test(...)
# {
# std::cout << "test(...)\n";
# }
#
# // this overload is added to the set of overloads if
# // C is a reference-to-class type and F is a pointer to member function of C
# template <class C, class F>
# auto test(C c, F f) -> decltype((void)(c.*f)(), void())
# {
# std::cout << "test(object)\n";
# }
#
# // this overload is added to the set of overloads if
# // C is a pointer-to-class type and F is a pointer to member function of C
# template <class C, class F>
# auto test(C c, F f) -> decltype((void)((c->*f)()), void())
# {
# std::cout << "test(pointer)\n";
# }
#
# struct X { void f() {} };
#
# int main(){
# X x;
# test( x, &X::f);
# test(&x, &X::f);
# test(42, 1337);
# }
# ```
# Стандартный трюк для определения какого-нибудь свойства типа (подробно и нудно разобрать каждую строчку и почему оно работает):
#
# ```c++
# // trait helper:
# // is_class<T>::value is true iff T is a class
# //
# // is_class<int>::value == false
# // is_class<string>::value == true
# template<typename T>
# class is_class {
# typedef char yes[1];
# typedef char no [2];
# template<typename C> static yes& test(int C::*); // selected if C is a class type
# template<typename C> static no& test(...); // selected otherwise
# public:
# static bool const value = (sizeof(test<T>(nullptr)) == sizeof(yes));
# };
# ```
# <br />
# Теперь мы можем написать идеальный `swap`.
#
# ```c++
# template<typename T>
# class has_swap_method
# {
# ...
# };
#
# ...
# ```
# __Шаг 1__: напишем `class has_swap_method` по аналогии с `class is_class`, который будет отвечать на вопрос, есть ли у шаблонного параметра метод `swap` (подробно объяснить)
# ```c++
# template<typename T>
# class has_swap_method
# {
# typedef char yes[1];
# typedef char no [2];
# template<typename U, void (U::*)(U&)> class S {};
# template<typename U> static yes& test(S<U, &U::swap>*);
# template<typename U> static no& test(...);
#
# public:
# static constexpr bool value = sizeof(test<T>(nullptr)) == sizeof(yes);
# };
# ```
# Протестируем:
#
# ```c++
# #include "has_swap_method_trait.h"
#
# #include <iostream>
# #include <string>
# #include <vector>
#
# template<typename T>
# void test_has_swap(const char* descr)
# {
# std::cout << descr
# << " has swap method: "
# << has_swap_method<T>::value
# << std::endl;
# }
#
# struct Point
# {
# float x;
# float y;
# };
#
# int main()
# {
# test_has_swap<int> ("int ");
# test_has_swap<float> ("float ");
# test_has_swap<Point> ("Point ");
# test_has_swap<std::vector<int>>("std::vector<int>");
# test_has_swap<std::string> ("std::string ");
#
# return 0;
# }
# ```
# Вывод:
#
# ```sh
# int has swap method: 0
# float has swap method: 0
# Point has swap method: 0
# std::vector<int> has swap method: 1
# std::string has swap method: 1
# ```
# __Шаг 2__: реализация `my_swap` (почти идеально; с использованием простенького ):
#
# ```c++
# template<typename T>
# void my_swap(T& x, T& y, std::false_type)
# {
# std::cout << "default swap\n";
#
# T t = std::move(x);
# x = std::move(y);
# y = std::move(t);
# }
#
# template<typename T>
# void my_swap(T& x, T& y, std::true_type)
# {
# std::cout << "optimized swap\n";
#
# x.swap(y);
# }
#
# template<typename T>
# void my_swap(T& x, T& y)
# {
# my_swap(x, y,
# std::integral_constant<bool, has_swap_method<T>::value>());
# }
# ```
# **Вопрос на понимание**: почему нельзя так?
#
# ```c++
# template<typename T>
# void my_swap(T& x, T& y)
# {
# if (has_swap_method<T>::value)
# {
# x.swap(y);
# }
# else
# {
# T t = std::move(x);
# x = std::move(y);
# y = std::move(t);
# }
# }
# ```
#
# **Замечание**: С С++17 есть `if constexpr` (показать на примере, как его написать, чтобы заработало)
#
# https://en.cppreference.com/w/cpp/language/if
# Протестируем:
#
# ```c++
# int main()
# {
# std::cout << "int: ";
# int i1 = 1, i2 = 2;
# my_swap(i1, i2);
#
# std::cout << "string: ";
# std::string s1 = "abc", s2 = "def";
# my_swap(s1, s2);
#
# return 0;
# }
# ```
# Вывод:
#
# ```sh
# int: default swap
# string: optimized swap
# ```
# <br />
# __Шаг 2__ через `std::enable_if` - лучше (объяснить):
#
# ```c++
# template<typename T>
# typename std::enable_if<has_swap_method<T>::value, void>::type
# my_swap(T& x, T& y)
# {
# std::cout << "optimized swap\n";
# x.swap(y);
# }
#
# template<typename T>
# typename std::enable_if<!has_swap_method<T>::value, void>::type
# my_swap(T& x, T& y)
# {
# std::cout << "default swap\n";
# T t = std::move(x);
# x = std::move(y);
# y = std::move(t);
# }
# ```
# Протестируем:
#
# ```c++
# int main()
# {
#
# std::cout << "int: ";
# int i1 = 1, i2 = 2;
# my_swap(i1, i2);
#
# std::cout << "string: ";
# std::string s1 = "abc", s2 = "def";
# my_swap(s1, s2);
#
# return 0;
# }
# ```
# Вывод:
#
# ```sh
# int: default swap
# string: optimized swap
# ```
# <br />
# __Вопрос на понимание:__ почему такой вариант не скомпилируется?
#
# ```c++
# template<typename T,
# void (T::*)(T&) = &T::swap>
# void my_swap(T& x, T& y)
# {
# std::cout << "optimized swap\n";
# x.swap(y);
# }
#
# template<typename T>
# void my_swap(T& x, T& y)
# {
# std::cout << "default swap\n";
# T t = std::move(x);
# x = std::move(y);
# y = std::move(t);
# }
#
# int main()
# {
#
# std::cout << "int: ";
# int i1 = 1, i2 = 2;
# my_swap(i1, i2);
#
# std::cout << "string: ";
# std::string s1 = "abc", s2 = "def";
# my_swap(s1, s2);
#
# return 0;
# }
# ```
#
# <details>
# <summary>ответ</summary>
# <p>
#
# для `std::string` оба варианта перегрузки подходят и имеют одинаковый приоритет в механизме overloading resolution, поэтому компилятор не может сделать выбор и генерирует ошибку.
#
# </p>
# </details>
# <br />
# Альтернативный способ реализации `has_method_swap` с использованием `decltype`:
#
# ```c++
# template<typename T>
# class has_swap_method
# {
# template<typename U, void (U::*)(U&)> class S {};
#
# template <typename C>
# static constexpr decltype(S<C, &C::swap>(), bool()) test(int /* unused */)
# {
# return true;
# }
#
# template <typename C>
# static constexpr bool test(...)
# {
# return false;
# }
#
# public:
# static constexpr bool value = test<T>(int());
# };
# ```
# <br />
# ##### variadic templates (C++11)
# https://en.cppreference.com/w/cpp/language/parameter_pack
# *variadic templates* - фича стандарта С++11, позволяющая писать шаблоны с переменным числом аргументов.
#
# Начнём сразу с демонстрационного примера: шаблонная функция записывает строку в csv-файл
#
# ```c++
# template<typename T>
# void printCSVLine(std::ostream& os, const T& v)
# {
# os << v << std::endl;
# }
#
# template<typename T1, typename T2, typename ...Args>
# void printCSVLine(std::ostream& os, const T1& v1, const T2& v2, Args&&... args)
# {
# os << v1 << ',';
# printCSVLine(os, v2, args...);
# }
# ```
#
# Использование:
#
# ```c++
# printCSVLine(std::cout, "name", "surname", "age", "gender");
# printCSVLine(std::cout, "Добрыня", "Никитич", 42, 'm');
# printCSVLine(std::cout, "Илья"s, "Муромец"sv, 33, 'm');
# printCSVLine(std::cout, "Василиса", "Премудрая", 35, 'f');
# ```
# <br />
# Пример форматированного вывода с cppreference:
#
# ```c++
# #include <iostream>
#
# void tprintf(const char* format) // base function
# {
# std::cout << format;
# }
#
# template<typename T, typename... Targs>
# void tprintf(const char* format, const T& value, const Targs&... Fargs) // recursive variadic function
# {
# for ( ; *format != '\0'; format++ ) {
# if ( *format == '%' ) {
# std::cout << value;
# tprintf(format + 1, Fargs...); // recursive call
# return;
# }
# std::cout << *format;
# }
# }
#
# int main()
# {
# tprintf("% world% %\n","Hello",'!',123); // Hello world! 123
# }
# ```
# <br />
# Как работает `...`?
#
# Простое объяснение: выражение слева от `...` раскрывается для каждого аргумента. В зависимости от контекста между аргументами может быть автоматически поставлена запятая (надо смотреть правила).
#
# ```c++
# f(&args...); // f(&E1, &E2, &E3)
# f(n, ++args...); // f(n, ++E1, ++E2, ++E3);
# f(++args..., n); // f(++E1, ++E2, ++E3, n);
# f(h(args...) + args...); // f(h(E1,E2,E3) + E1, h(E1,E2,E3) + E2, h(E1,E2,E3) + E3)
# ```
# <br />
# ##### fold expression (C++17)
# https://en.cppreference.com/w/cpp/language/fold
# *fold expressions* - фича стандарта С++17 - добавляет различные способы раскрытия `...` у variadic template.
# Общий вид добавленных правил:
#
# ```c++
# ( pack op ... )
# ( ... op pack )
# ( pack op ... op init )
# ( init op ... op pack )
# ```
#
# где op - бинарная/унарная правая/левая операция.
#
# Соответственно, 4 правила развёртки:
#
# * Unary right fold $(E op ...) -> (E_1 op (... op (E_{N-1} op E_N)))$
# * Unary left fold $(... op E) -> (((E_1 op E_2) op ...) op E_N)$
# * Binary right fold $(E op ... op I) -> (E_1 op (... op (E_{N−1} op (E_N op I))))$
# * Binary left fold $(I op ... op E) -> ((((I op E_1) op E_2) op ...) op E_N)$
# Примеры их применения:
# ```c++
# template<typename T, typename ...Args>
# auto sum(const T& v, const Args&... args) {
# return v + ... + args;
# }
#
# sum(1, 2, 3, 4, 5, 6, 7); // 28
# sum("abc"s, "def", "ghi"); // ?
# ```
# ```c++
# template<typename ...Args>
# void printer(Args&&... args) {
# (std::cout << ... << args) << '\n';
# }
#
# printer("hello", "world"); // helloworld
# ```
# <br />
# ##### sizeof...(args) (C++11)
# Есть особый оператор `sizeof...(args)`, который возвращает число аргументов.
#
# Рассмотрим его на примере добавления нескольких элементов в `std::vector`:
#
# ```c++
# template<typename T, typename... Args>
# void add_to_vector(std::vector<T>& v, Args&&... args)
# {
# v.reserve(v.size() + sizeof...(args));
# (v.push_back(std::forward<Args>(args)), ...);
# }
#
# // usage:
# std::vector<std::string> v;
# add_to_vector(v, "Добрыня", "Илюша"s, "Алёша"sv);
# add_to_vector(v, "Василиса", "Настасья"s);
# ```
# <br />
# **Замечание**: почти все примеры на variadic templates были ученическими, но они являют собой достаточно мощный механизм.
#
# Классический пример "боевого" применения - форматирование строк `std::format`, варианты inplace конструирования `make_unique`/`make_shared`/`vector::emplace`/`optional::emplace` и т.д:
#
# ```c++
# auto x = std::make_unique<std::vector<std::string>>("abc", 10);
# ```
#
# > template< class T, class... Args >
# unique_ptr<T> make_unique( Args&&... args );
#
# ```c++
# std::vector<std::string> v;
# v.emplace_back("a", 10);
# ```
#
# > template< class... Args >
# reference emplace_back( Args&&... args );
#
# **Вопрос**: что такое и почему && ?
# <br />
# ##### tag dispatching && type traits
# *tag dispatching* - техника выбора поведения/реализации, основыванная на типе одного из параметров (тэге) и механизме перегрузки. Обычно, этот параметр - пустая структурка.
# С примером простенького tag dispatching мы уже познакомились на примере `my_swap`:
#
# ```c++
# template<typename T>
# void my_swap_dispatched(T& x, T& y, std::false_type)
# {
# T t = std::move(x);
# x = std::move(y);
# y = std::move(t);
# }
#
# template<typename T>
# void my_swap_dispatched(T& x, T& y, std::true_type)
# {
# x.swap(y);
# }
#
# template<typename T>
# void my_swap(T& x, T& y)
# {
# my_swap_dispatched(x, y, std::integral_constant<bool, has_swap_method<T>::value>());
# }
# ```
# <br />
# Традииционный пример использования tag dispatching вместе с type triants: `std::advance`
#
# Что такое `std::advance`:
#
# ```c++
# // интерфейс
# template< class InputIt, class Distance >
# void advance( InputIt& it, Distance n );
#
# // использование:
# std::vector<int> v = { ... };
# auto vit = v.begin();
# std::advance(vit, 5); // один "прыжок"
#
# std::list<int> l = { ... };
# auto lit = l.end();
# std::advance(lit, -5); // 5 "прыжков" назад
#
# std::forward_list<int> f = { ... };
# auto fit = f.begin();
# std::advance(fit, 5); // 5 "прыжков" вперёд
# ```
#
# Как его можно реализовать:
#
# ```c++
# namespace std {
# namespace detail {
# template <class It, class D>
# void advance_dispatch(It& i, D n, input_iterator_tag) {
# while (n--) ++i;
# }
#
# template <class It, class D>
# void advance_dispatch(It& i, D n, bidirectional_iterator_tag) {
# if (n >= 0)
# while (n--) ++i;
# else
# while (n++) --i;
# }
#
# template <class It, class D>
# void advance_dispatch(It& i, D n, random_access_iterator_tag) {
# i += n;
# }
# }
#
# template <class It, class Distance>
# void advance(It& i, Distance n) {
# typename iterator_traits<It>::iterator_category category;
# detail::advance_dispatch(i, n, category());
# }
# }
# ```
# Нюанс в определении типа `iterator_traits<It>::iterator_category`
# Делается это примерно так:
#
# ```c++
# struct input_iterator_tag { };
# struct bidirectional_iterator_tag { };
# struct random_access_iterator_tag { };
#
# template<typename It>
# struct iterator_traits
# {
# using iterator_category = input_iterator_tag;
# ...
# };
#
# template<typename T>
# struct iterator_traits<std::vector<T>::iterator>
# {
# using iterator_category = random_access_iterator_tag;
# ...
# };
# ```
#
# После того как определён type trait для итераторов, любой алгоритм, желающий использовать преимущества random_access_iterator, может сделать это через tag dispatching.
#
# Минус подхода в том, что для такой реализации нужно для каждого итераторане забыть прописать iterator_traits (или он свалится в самый слабый вариант traits)
# <br />
# ##### стандартные type traits
# https://en.cppreference.com/w/cpp/types
#
# В стандартную библиотеку добавлено большое кол-во trait-ов, чтобы программисты не мучились и не писали свои велосипеды. Полный список смотрите по ссылке, а вот некоторые из них:
#
# ```c++
# std::is_integral<T>
# std::is_floating_point<T>
# std::is_array<T>
# std::is_trivial<T>
# std::is_same<T, U>
# ```
# <br />
# ##### deduction guides (C++17)
# https://en.cppreference.com/w/cpp/language/class_template_argument_deduction
# *class template argument deduction* - нововведение С++17: при использовании шаблона класса программист не обязан указывать параметры шаблона, если компилятор может вывести их из контекста (и программиста устроит то, что вывел компилятор).
#
# ```c++
# std::vector v = {1, 2, 3, 4, 5}; // std::vector<int> автоматически
# std::pair p = {42, 3.14}; // std::pair<int, double> автоматически
# std::list l = {"hello", "world" }; // ?
# ```
#
#
#
#
# Именно поэтому работают примеры из лекции по многопотоности:
#
# ```c++
# std::mutex mtx;
# std::lock_guard guard(mtx); // std::lock_guard<std::mutex> guard(mtx);
# ```
# <br />
# Рассмотрим снова пример
#
# ```c++
# std::vector v = {1, 2, 3, 4, 5};
# ```
#
# Правило, по которому компилятор понимает, что при создании `std::vector<T>` из `std::initializer_list<int>` нужно взять `T = int`, называется *deduction guides*.
# Какие-то правила прописаны по умолчанию и неявно применяются для всех классов (implicitly-generated deduction guides), какие они - читайте стандарт по ссылке.
# Иногда неявных правил не хватает, и программисту хочется добавить свои. Такие правила называются пользовательскими (user defined deduction guides)
# Для имеющихся контейнеров уже всё готово, поэтому предположим, у нас есть свой личный контейнер:
#
# ```c++
# template<typename T>
# class flat_set
# {
# public:
# flat_set(std::initializer_list<T> items);
#
# template<typename It>
# flat_set(It begin, It end);
# };
# ```
#
# Напишем deduction guides для нашего типа:
#
# ```c++
# template<class It>
# flat_set(It begin, It end) -> flat_set<typename std::iterator_traits<It>::value_type>;
# ```
#
# Использование:
#
# ```c++
# flat_set x = {1, 2, 3, 4, 5}; // flat_set<int> по неявным правилам вывода
#
# std::vector v = {1, 2, 3, 4, 5};
# flat_set x{v.begin(), v.end()}; // flat_set<int> по нашему правилу
# ```
# <br />
# **Резюме**:
#
# * `SFINAE` - техника выбора более оптимизированной реализации на этапе компиляции по типу, пример с `my_swap` (шаблоны + перегрузки)
# * variadic templates && fold expressions - шаблоны переменного числа аргументов, необходимый механизм для функций `make_unique`, `make_shared`, `emplace_back`.
# * deduction guides - способ автоматического вывода типов у шаблонов классов через выражение справа.
# <br />
# **Замечания после лекции**:
# * разбавить примерами, плохо понимают sfinae
# * int C::* - записать объяснение что это за такое
# <br />
| 2020/sem2/lecture_5_templates_adv/lecture.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: tf1
# language: python
# name: tf1
# ---
# # Week 3 - Notes
#
#
#
# These are some concise notes representing the most important ideas discussed.
#
# --------
#
# ## MDP: RL Approach
# ### Batch Reinforcement Learning
#
# - Unlike DP, RL does NOT assume any knowledge of the world dynamics. It instead relies on samples to find optimal policy.
# - The **Conventional Approach** to Option Pricing requires building a model of the world by designing a stochastic process and then caliberating it to aution and stock pricing data.
# - **Model Calibration**: is the idea of formulating the law of dynamics by estimating model parameters. This amounts to minimization of some loss function between what's observed and model outputs.
#
#
# - **RL** on the other hand focuses on the original taks of finding an optimal price and hedge by relying on data samples instead of a model.
#
# - **Batch RL** is an off-line RL, where agent mainly relies on some histoically collected data.
#
#
# - **<NAME>** formulated the principle: "one should avoid solving more difficult intermediate problems when solving a target problem."
#
#
#
# - Unlike DP, RL does NOT need to know reward and transition probability functions, as it relies on samples.
#
# - The information set for RL includes a tuplet $(X_t^{(n)}, a_t^{(n)}, R_t^{(n)}, X_t+1^{(n)}$ for each step.
#
#
# ### Stochastic Approximations
#
#
# - Again, we try to approximately solve Bellman Optimality Eq: $$Q_t^*(x,a) = \mathbb{E}[R_t + \gamma \max_{a\in A} Q_{t+1}^*(X_{t+1}|x,a)]$$
# - Optimal Q-fn value is equal to the expected optimal rewards, plus the discounted expected value of the next step optimal Q-fn.
# - Expectation here is one-step expectation, involving next step quantities being conditional on the information available at time T.
#
#
# - For a *discrete* state model, we simply sum up over all possible next states with the corresponding transition probabilities as weights in this sum.
# - For a *continuous* state model, the calculation of expectation involves integrals instead of sums.
# - We replace expectations by their empirical averages.
#
#
# - We already know how the conventional DP would work in such setting. Yet with RL we observe stock prices in the rewards, and mainly rely on empirical means to estimate the Right Hand of Bellman Optimality. Hence, estimating optimal Q-values.
#
# - The **Robbins-Monro** Alg estimates the mean without directly summing the samples, but instead adding data points one by one, and iteratively updating the running estimation of the mean $\hat{x}_k$ (where $k$ is the number of iterations, or the number of data points in a dataset): $$\hat{x}_{k+1}=(1-\alpha_k)\hat{x}_k + \alpha_k x_k$$
# - The advantage of such approach is it actually does converge to a true mean with probability of one under certain conditions.
#
#
#
# - Robbins-Monro Alg can be used for both on-line and batch-mode setting.
# - The optimal choice of learning rate in the Robbins-Monroe Alg is NOT universal but depends on the problem.
# ### Q-Learning
#
#
# - As per Watkins, Q-Learning works only in a setting called discrete states and discrete actions. We say Q-fn is in tabular form when using one value of Q-fn for each combination of states and actions.
#
# - Q-Learning converges to the true optimal action-value fn with probability of one, given enough data.
#
# - The optimal Q-fn is learnt in Q-Learning iteratively, where each step (Q-Iteration) implements one iteration of the Rob<NAME>.
#
# - Q-Learning is obtained by using the Robbins-Monro Stochastic Approximation to estimate the unknown expectation in Bellman Optimality.
#
#
# $$Q_{t,k+1}^*(X_t,a_t)=(1-\alpha_k)Q_{tk}^*(X_t,a_t)+\alpha_k[R_t(X_t,a_t,X_{t+1}+\gamma \max_{a\in A}Q{t+1,k}^*(X_{t+1},a) )]$$
# ### Fitted Q-Iteration
#
#
# - For Monte Carlo path, historical path of the stock was used simultaneously when calculating an optimal policy. The problem however was that the classical Q-Learning will too long to converge. We aim for something faster.
#
# - Most popular extension of of Q-Learning to Batch RL setting is what's called **Fitted Q Iteration**. It was mainly developed for time-stationary problems, where Q-fn does NOT depend on time. Yet it can be also applied for both discrete and continuous state-action spaces.
#
# - FQI works by using all Monte Carlo paths for the replication portfolio simultaneously. We use the same set of basis functions ${\Phi_n(x)}$ as we did with DP.
# - Optimal Q-fn $Q_t^*(X_t,a_t)$ is a quadratic function of $a_t$. It can be written as an expansion in basis functions, with time-dependent coefficient matrix $\mathbf W_t$:
#
#
# $$Q_t^*(X_t,a_t)= A_t^T \mathbf W_t \Phi(X_t)$$
#
# ### Fitted Q-Iteration: the $\epsilon$-basis
#
# - Here we just want to convert matrix $W_t$ to a vector. So we are going to rearrange the previous equation, converting it into a product of a parameter vector and a vector that depends on both the state and the action:
#
#
# $$Q_t^*(x,a)=A_t^T \mathbf W_t \Phi (X)$$
#
#
# $$ = \vec{\mathbf W}_t \vec{\Psi} (X_t,a_t) $$
#
#
# - $\vec{W}_t$ is obtained by concatenating columns of matrix $\mathbf W_t$ while vec $ vec \left( {\bf \Psi} \left(X_t,a_t \right) \right) =
# vec \, \left( {\bf A}_t \otimes {\bf \Phi}^T(X) \right) $ stands for
# a vector obtained by concatenating columns of the outer product of vectors $ {\bf A}_t $ and $ {\bf \Phi}(X) $.
#
#
# - For **RL**:
# - both parameter vec $vec{\mathbf W}_t$ and state-action basis $vec{\Psi}_t$ have 3M components.
# - number of data records per time step is 3N $(X_t,a_t,R_t)$.
#
# - For **DP**:
# - 2M parameters.
# - only N values of $X_t$ as input data.
#
# - Counting by the number of parameters to learn, the RL setting has more unknowns, but also a higher dimensionality of data (more data per observation) than DP setting.
# ### Fitted Q-Iteration at Work
#
#
#
#
# - Coefficients $\mathbf W_t$ shall be computed recursively backward in time as before.
# - One step **Bellman Iteration** can be interpreted as Regression: $$R_t(X_t,a_t,x_{t+1}) + \gamma \max Q_{a \in A}^*(X_{t+1},a)$$
#
# $$ = \vec{\mathbf W}_t \vec{\mathbf \Psi}(X_t,a_t) + \epsilon_t $$
#
#
# - For **DP**:
# - Q-fn is expanded in the state basis $\Phi_t$.
# - applies only at optimal Q-fn for optimal action $a_t^*$
# - rewards $R_t$ are computed
#
# - For **RL**:
# - Q-fn is expanded in the state-action basis $\vec{\Psi}_t$
# - works for any action $a_t$, not just optimal actions
# - both rewards $R_t$ and actions $a_t$ are observed
#
#
#
#
# - In the DP and RL solutions, Q-fn is an expansion in a set of state-action basis functions.
# ### RL Solution: Discussion and Examples
| Finance/ML & RL in Finance - NYU/3. RL in Finance/3_week_3_notes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/NeuromatchAcademy/course-content-dl/blob/main/tutorials/W2D5_GenerativeModels/student/W2D5_Tutorial2.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# -
# # Tutorial 2: Introduction to GANs and Density Ratio Estimation Perspective of GANs
#
# **Week 2, Day 5: Generative Models**
#
# **By Neuromatch Academy**
#
# __Content creators:__ <NAME>, <NAME>, <NAME>
#
# __Content reviewers:__ <NAME>, <NAME>, <NAME>, <NAME>, <NAME>
#
# __Content editors:__ <NAME>, <NAME>
#
# __Production editors:__ <NAME>, <NAME>
# **Our 2021 Sponsors, including Presenting Sponsor Facebook Reality Labs**
#
# <p align='center'><img src='https://github.com/NeuromatchAcademy/widgets/blob/master/sponsors.png?raw=True'/></p>
# ---
#
# ## Tutorial Objectives
#
# The goal of this tutorial is two-fold; first you will be introduced to GANs training, and you will be able to understand how GANs are connected to other generative models that we have been before.
#
# By the end of the first part of this tutorial you will be able to:
# - Understand, at a high level, how GANs are implemented.
# - Understand the training dynamics of GANs.
# - Know about a few failure modes of GAN training.
# - Understand density ratio estimation using a binary classifier
# - Understand the connection between GANs and other generative models.
# - Implement a GAN.
# + cellView="form"
# @title Tutorial slides
# @markdown These are the slides for the videos in this tutorial
from IPython.display import IFrame
IFrame(src=f"https://mfr.ca-1.osf.io/render?url=https://osf.io/dftym/?direct%26mode=render%26action=download%26mode=render", width=854, height=480)
# -
# ---
# # Setup
# +
# Imports
import torch
import numpy as np
import torch.nn as nn
import torch.optim as optim
import matplotlib.pyplot as plt
from time import time
# + cellView="form"
# @title Figure settings
import ipywidgets as widgets # interactive display
# %config InlineBackend.figure_format = 'retina'
plt.style.use("https://raw.githubusercontent.com/NeuromatchAcademy/content-creation/main/nma.mplstyle")
# + cellView="form"
# @title Plotting functions
ld_true = [-7.0066e-01, -2.6368e-01, -2.4250e+00, -2.0247e+00, -1.1795e+00,
-4.5558e-01, -7.1316e-01, -1.0932e-01, -7.8608e-01, -4.5838e-01,
-1.0530e+00, -9.1201e-01, -3.8020e+00, -1.7787e+00, -1.2246e+00,
-6.5677e-01, -3.6001e-01, -2.2313e-01, -1.8262e+00, -1.2649e+00,
-3.8330e-01, -8.8619e-02, -9.2357e-01, -1.3450e-01, -8.6891e-01,
-5.9257e-01, -4.8415e-02, -3.3197e+00, -1.6862e+00, -9.8506e-01,
-1.1871e+00, -7.0422e-02, -1.7378e+00, -1.3099e+00, -1.8926e+00,
-3.4508e+00, -1.5696e+00, -7.2787e-02, -3.2420e-01, -2.9795e-01,
-6.4189e-01, -1.4120e+00, -5.3684e-01, -3.4066e+00, -1.9753e+00,
-1.4178e+00, -2.0399e-01, -2.3173e-01, -1.2792e+00, -7.2990e-01,
-1.9872e-01, -2.9378e-03, -3.5890e-01, -5.6643e-01, -1.8003e-01,
-1.5818e+00, -5.2227e-01, -2.1862e+00, -1.8743e+00, -1.4200e+00,
-3.1988e-01, -3.5513e-01, -1.5905e+00, -4.2916e-01, -2.5556e-01,
-8.2807e-01, -6.5568e-01, -4.8475e-01, -2.1049e-01, -2.0104e-02,
-2.1655e+00, -1.1496e+00, -3.6168e-01, -8.9624e-02, -6.7098e-02,
-6.0623e-02, -5.1165e-01, -2.7302e+00, -6.0514e-01, -1.6756e+00,
-3.3807e+00, -5.7368e-02, -1.2763e-01, -6.6959e+00, -5.2157e-01,
-8.7762e-01, -8.7295e-01, -1.3052e+00, -3.6777e-01, -1.5904e+00,
-3.8083e-01, -2.8388e-01, -1.5323e-01, -3.7549e-01, -5.2722e+00,
-1.7393e+00, -2.8814e-01, -5.0310e-01, -2.2077e+00, -1.5507e+00,
-6.8569e-01, -1.4620e+00, -9.2639e-02, -1.4160e-01, -3.6734e-01,
-1.0053e+00, -6.7353e-01, -2.2676e+00, -6.0812e-01, -1.0005e+00,
-4.2908e-01, -5.1369e-01, -2.2579e-02, -1.8496e-01, -3.4798e-01,
-7.3089e-01, -1.1962e+00, -1.6095e+00, -1.7558e-01, -3.3166e-01,
-1.1445e+00, -2.4674e+00, -5.0600e-01, -2.0727e+00, -5.4371e-01,
-8.0499e-01, -3.0521e+00, -3.6835e-02, -2.0485e-01, -4.6747e-01,
-3.6399e-01, -2.6883e+00, -1.9348e-01, -3.1448e-01, -1.6332e-01,
-3.2233e-02, -2.3336e-01, -2.6564e+00, -1.2841e+00, -1.3561e+00,
-7.4717e-01, -2.7926e-01, -8.7849e-01, -3.3715e-02, -1.4933e-01,
-2.7738e-01, -1.6899e+00, -1.5758e+00, -3.2608e-01, -6.5770e-01,
-1.7136e+00, -5.8316e+00, -1.1988e+00, -8.3828e-01, -1.8033e+00,
-2.3017e-01, -8.9936e-01, -1.1917e-01, -1.6659e-01, -2.7669e-01,
-1.2955e+00, -1.2076e+00, -2.2793e-01, -1.0528e+00, -1.4894e+00,
-5.7428e-01, -7.3208e-01, -9.5673e-01, -1.6617e+00, -3.9169e+00,
-1.2182e-01, -3.8092e-01, -1.1924e+00, -2.4566e+00, -2.7350e+00,
-2.8332e+00, -9.1506e-01, -6.7432e-02, -7.8965e-01, -2.0727e-01,
-3.4615e-02, -2.8868e+00, -2.1218e+00, -1.2368e-03, -9.0038e-01,
-5.3746e-01, -5.4080e-01, -3.1625e-01, -1.1786e+00, -2.2797e-01,
-1.1498e+00, -1.3978e+00, -1.9515e+00, -1.1614e+00, -5.1456e-03,
-1.9316e-01, -1.3849e+00, -9.2799e-01, -1.1649e-01, -2.3837e-01]
def plotting_ld(ld, true=ld_true):
fig, ax = plt.subplots(figsize=(7, 7))
ax.plot([-6, 1], [-6, 1], label="Ground Truth")
ax.scatter(true, ld, marker="x",
label="Your implementation")
ax.set_xlabel("Loss from oracle implementation")
ax.set_ylabel("Loss from your implementation")
ax.legend()
ax.set_title("Discriminator Loss")
lg_true = [-7.0066e-01, -2.6368e-01, -2.4250e+00, -2.0247e+00, -1.1795e+00,
-4.5558e-01, -7.1316e-01, -1.0932e-01, -7.8608e-01, -4.5838e-01,
-1.0530e+00, -9.1201e-01, -3.8020e+00, -1.7787e+00, -1.2246e+00,
-6.5677e-01, -3.6001e-01, -2.2313e-01, -1.8262e+00, -1.2649e+00,
-3.8330e-01, -8.8619e-02, -9.2357e-01, -1.3450e-01, -8.6891e-01,
-5.9257e-01, -4.8415e-02, -3.3197e+00, -1.6862e+00, -9.8506e-01,
-1.1871e+00, -7.0422e-02, -1.7378e+00, -1.3099e+00, -1.8926e+00,
-3.4508e+00, -1.5696e+00, -7.2787e-02, -3.2420e-01, -2.9795e-01,
-6.4189e-01, -1.4120e+00, -5.3684e-01, -3.4066e+00, -1.9753e+00,
-1.4178e+00, -2.0399e-01, -2.3173e-01, -1.2792e+00, -7.2990e-01,
-1.9872e-01, -2.9378e-03, -3.5890e-01, -5.6643e-01, -1.8003e-01,
-1.5818e+00, -5.2227e-01, -2.1862e+00, -1.8743e+00, -1.4200e+00,
-3.1988e-01, -3.5513e-01, -1.5905e+00, -4.2916e-01, -2.5556e-01,
-8.2807e-01, -6.5568e-01, -4.8475e-01, -2.1049e-01, -2.0104e-02,
-2.1655e+00, -1.1496e+00, -3.6168e-01, -8.9624e-02, -6.7098e-02,
-6.0623e-02, -5.1165e-01, -2.7302e+00, -6.0514e-01, -1.6756e+00,
-3.3807e+00, -5.7368e-02, -1.2763e-01, -6.6959e+00, -5.2157e-01,
-8.7762e-01, -8.7295e-01, -1.3052e+00, -3.6777e-01, -1.5904e+00,
-3.8083e-01, -2.8388e-01, -1.5323e-01, -3.7549e-01, -5.2722e+00,
-1.7393e+00, -2.8814e-01, -5.0310e-01, -2.2077e+00, -1.5507e+00]
def plotting_lg(lg, true=lg_true):
fig, ax = plt.subplots(figsize=(7, 7))
ax.plot([-6, 1], [-6, 1], label="Ground Truth")
ax.scatter(true, lg, marker="x",
label="Your implementation")
ax.set_xlabel("Loss from oracle implementation")
ax.set_ylabel("Loss from your implementation")
ax.legend()
ax.set_title("Generator loss")
def plotting_ratio_impl(ax, x_real, x_fake, ratio, yscale="linear"):
dist_p = torch.distributions.normal.Normal(loc=0, scale=1)
dist_q = torch.distributions.normal.Normal(loc=-2, scale=1)
x = torch.linspace(-3, 5, 100)
prob_p = torch.exp(dist_p.log_prob(x))
prob_q = torch.exp(dist_q.log_prob(x))
trueRatio = prob_p / prob_q
ax.plot(x, trueRatio, label="True tatio")
x = torch.cat([x_real, x_fake])
ax.scatter(x[:,0][::10], ratio[:,0][::10], marker="x",
label="Ratio from discriminator")
ax.hist(x_real[:,0], density=True, bins=50, histtype="step", label="Real")
ax.hist(x_fake[:,0], density=True, bins=50, histtype="step", label="Fake")
ax.set_yscale(yscale)
title = "Densities and the ratio from discriminator"
if yscale == "log":
title += " in log scale"
ax.set_title(title)
ax.legend()
def plotting_ratio(x_real, x_fake, ratio):
fig, axes = plt.subplots(1, 2, figsize=(2 * 7, 7))
plotting_ratio_impl(axes[0], x_real, x_fake, ratio, yscale="linear")
plotting_ratio_impl(axes[1], x_real, x_fake, ratio, yscale="log")
class Interactive:
def display_widgets(self):
for widget in self.widgets:
display(widget)
def __init__(self, widgets, handler):
def handler_with_extra_steps(b):
clear_output(wait=True)
handler(*map(lambda w: w.value, widgets))
self.display_widgets()
self.widgets = widgets
for widget in self.widgets:
widget.observe(handler_with_extra_steps,
names=['value'])
handler(*map(lambda w: w.value, widgets))
self.display_widgets()
# Using Interactive
# All widgets: https://ipywidgets.readthedocs.io/en/latest/examples/Widget%20List.html
def make_plot(xmax, title, ftype):
fig, ax = plt.subplots()
x = np.linspace(-2, xmax, 100)
if ftype == "sin":
y = np.sin(x)
if ftype == "cos":
y = np.cos(x)
if ftype == "tanh":
y = np.tanh(x)
ax.scatter(x, y)
ax.set_xlim(-2.1, 2.1)
ax.set_ylim(-2, 2)
if title:
ax.set_title(f"Range from -1 to {xmax}")
return fig
# + cellView="form"
# @title Set random seed
# @markdown Executing `set_seed(seed=seed)` you are setting the seed
# for DL its critical to set the random seed so that students can have a
# baseline to compare their results to expected results.
# Read more here: https://pytorch.org/docs/stable/notes/randomness.html
# Call `set_seed` function in the exercises to ensure reproducibility.
import random
import torch
def set_seed(seed=None, seed_torch=True):
if seed is None:
seed = np.random.choice(2 ** 32)
random.seed(seed)
np.random.seed(seed)
if seed_torch:
torch.manual_seed(seed)
torch.cuda.manual_seed_all(seed)
torch.cuda.manual_seed(seed)
torch.backends.cudnn.benchmark = False
torch.backends.cudnn.deterministic = True
print(f'Random seed {seed} has been set.')
# In case that `DataLoader` is used
def seed_worker(worker_id):
worker_seed = torch.initial_seed() % 2**32
np.random.seed(worker_seed)
random.seed(worker_seed)
# + cellView="form"
# @title Set device (GPU or CPU). Execute `set_device()`
# especially if torch modules used.
# inform the user if the notebook uses GPU or CPU.
def set_device():
device = "cuda" if torch.cuda.is_available() else "cpu"
if device != "cuda":
print("WARNING: For this notebook to perform best, "
"if possible, in the menu under `Runtime` -> "
"`Change runtime type.` select `GPU` ")
else:
print("GPU is enabled in this notebook.")
return device
# -
SEED = 2021
set_seed(seed=SEED)
DEVICE = set_device()
# ---
# # Section 1: How to train GANs.
#
# + cellView="form"
# @title Video 1: Generative Adversarial Networks
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = "https://player.bilibili.com/player.html?bvid={0}&page={1}".format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id=f"BV19V411H72M", width=854, height=480, fs=1)
print("Video available at https://www.bilibili.com/video/{0}".format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id=f"fCGnJPW2RKM", width=854, height=480, fs=1, rel=0)
print("Video available at https://youtube.com/watch?v=" + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
# -
# GANs consist two networks: A critic or discriminator (`disc`) and a generator (`gen`) that are trained by alternating between the following two steps:
# - In step 1, we update the parameters (`disc.params`) of the discriminator by backpropagating through the discriminator loss (BCE loss) `disc.loss`.
# - In step 2, we update the parameters (`gen.params`) of the generator by backpropagating through the generator loss, `gen.loss` (-1 * BCE loss).
#
# We will now implement a simple GAN training loop!
# ## Coding Exercise 1: The GAN training loop
#
# To get you started we have implemented a simple GAN in pseudocode. All you have to do is to implement the training loop.
#
# __Your goal__ is to arrange the functions given below in the correct order in the `train_gan_iter` function
# - `disc.loss(x_real, x_fake)`: Discriminator loss
# - `disc.classify(x)`: Classify `x` as real or fake
# - `gen.loss(x_fake, disc_fn)`: Generator loss
# - `disc_fn(x)` is a function to check `x` is real or fake.
# - `gen.sample(num_samples)`: Generate samples from the generator
# - `backprop(loss, model)`: Compute gradient of `loss` wrt `model`
# - `model` is either `disc` or `gen`
#
# We have already taken care of most of these functions. So you only have to figure out the placement of `disc.loss` and `gen.loss` functions.
#
# __We highly recommend studying `train_gan_iter` function to understand how the GAN training loop is structured.__
# + cellView="form"
# @markdown *Execute this cell to enable helper functions*
def get_data():
return "get_data"
class Disc:
def loss(self, x_real, x_fake):
assert x_real == "get_data" and x_fake == "gen.sample","Inputs to disc.loss is wrong"
def classify(self, x):
return "disc.classify"
class Gen:
def loss(self, x_fake, disc_fn):
assert x_fake == "gen.sample" and disc_fn(None) == "disc.classify", "Inputs to gen.loss is wrong"
def sample(self, num_samples):
return "gen.sample"
def backprop(loss, model):
pass
def update(model, grad):
pass
# +
def train_gan_iter(data, disc, gen):
"""Update the discriminator (`disc`) and the generator (`gen`) using `data`
Args:
data (ndarray): An array of shape (N,) that contains the data
disc (Disc): The discriminator
gen (Gen): The generator
Returns:
"""
#################################################
# Intructions for students: #
# Fill out ... in the function and remove below #
#################################################
# Number of samples in the data batch
num_samples = 200
# The data is the real samples
x_real = data
## Discriminator training
# Ask the generator to generate some fake samples
x_fake = gen.sample(num_samples)
#################################################
## TODO for students: details of what they should do ##
# Fill out function and remove
raise NotImplementedError("Student exercise: Write code to compute disc_loss")
#################################################
# Compute the discriminator loss
disc_loss = ...
# Compute the gradient for discriminator
disc_grad = backprop(disc_loss, disc)
# Update the discriminator
update(disc, disc_grad)
## Generator training
# Ask the generator to generate some fake samples
x_fake = gen.sample(num_samples)
#################################################
## TODO for students: details of what they should do ##
# Fill out function and remove
raise NotImplementedError("Student exercise: Write code to compute gen_loss")
#################################################
# Compute the generator loss
gen_loss = ...
# Compute the gradient for generator
gen_grad = backprop(gen_loss, gen)
# Update the generator
update(gen, gen_grad)
print("Your implementation passes the check!")
data = get_data()
disc = Disc()
gen = Gen()
## Uncomment below to check your function
# train_gan_iter(data, disc, gen)
# + [markdown] colab_type="text"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content-dl/tree/main//tutorials/W2D5_GenerativeModels/solutions/W2D5_Tutorial2_Solution_dc5b38a9.py)
#
#
# -
# ---
# # Section 2: The difficulty of GAN training.
#
# In this section we will develop an intuition for the training dynamics of GANs.
# ## Interactive Demo 2: Failure modes of GAN training
#
# GAN training is notoriously difficult because
# it is very sensitive to hyper-parameters such as learning rate and model architecture. To help you develop a sense of this here is a very simple GAN training demo that we have borrowed from [<NAME>'s website](https://cs.stanford.edu/people/karpathy/gan/).
#
# The generator $G$, pictured in red, takes inputs sampled from a uniform distribution, $z$. It attempts to transform these to match a data distribution, shown below in blue. Meanwhile, the discriminator $D$ attempts to determine whether a sample is from the data distribution or the generating distribution. In the demo, the green curve represents the output of the discriminator. Its value is high where the discriminator is more confident that a sample with that value is drawn from the data distribution.
#
# Even though the GAN in this demo is very simple and operates in either 1D or 2D spaces, it is however very sensitive to the learning rate. Try it for yourself!
# + cellView="form"
# @title GAN training demo
# @markdown Make sure you execute this cell to enable the widget!
from IPython.display import IFrame
IFrame(src='https://xukai92.github.io/gan_demo/index.html', width=900, height=600)
# -
# ## Exercise 2: What makes GANs hard to train?
#
# You have played with the demo and it's time to think about a few questions
#
# 1. Which target is more stable to train, 1D or 2D?
# 2. If you keep increasing the learning rate, what happens? Does it happen in both the cases, i.e., 1D/2D targets?
# 3. Can you think of some drawbacks of using small learning rates?
# + [markdown] colab_type="text"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content-dl/tree/main//tutorials/W2D5_GenerativeModels/solutions/W2D5_Tutorial2_Solution_8febc070.py)
#
#
# -
# ---
# # Section 3: GAN Training Objective
# The training objective of GANs consists of the losses for generators and discriminators respectively. In this section we will be implementing these objectives.
# + cellView="form"
# @title Video 2: Principles of GANs
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = "https://player.bilibili.com/player.html?bvid={0}&page={1}".format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id=f"BV1Fq4y1s7pf", width=854, height=480, fs=1)
print("Video available at https://www.bilibili.com/video/{0}".format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id=f"bTJfZro6A9Q", width=854, height=480, fs=1, rel=0)
print("Video available at https://youtube.com/watch?v=" + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
# -
# ## Section 3.1: Discriminator Loss
#
# The critic or the discriminator in a vanilla GAN is trained as a binary classifier using the BCE criteria. In this section, we will implement the training objective for the discriminator.
#
# \begin{equation}
# \text{BCE}_\omega = \mathbb{E}_{x \sim p}[\log(\sigma(D_\omega(x)))] + \mathbb{E}_{x \sim q}[\log(1 - \sigma(D_\omega(x)))]
# \end{equation}
#
# Here, $p$ is the data distribution and $q$ is the generator distribution. $D_\omega$ is the logit, which represents $\log \frac{p}{q}$. $\sigma$ is the sigmoid function and therfore, $\sigma(D_\omega)$ represents $\frac{p}{p+q}$.
# ### Coding Exercise 3.1: Implement Discriminator Loss
#
# To get you started we have implemented a simple GAN in pseudocode and partially implemented the discriminator training objective.
#
# **Your goal** is to complete the missing part in the training objective of the discriminator in the function `loss_disc`.
#
# `loss_disc` also allows you evaluate the loss function on some random samples.
# If your implementation is correct, you will see a plot where the loss values from your implementation will match the ground truth loss values.
#
# In practice, given $N$ samples, we estimate BCE as
#
# \begin{equation}
# \text{BCE}_\omega = -\frac{1}{N} \sum_{i=1}^N y_i \log(\sigma(D_\omega(x_i)) + (1-y_i) \log(1-\sigma(D_\omega(x_i))).
# \end{equation}
#
# Here, $y$ is the label. $y=1$ when $x \sim p$ (real data) and $y=0$ when $x \sim q$ (i.e., fake data).
#
# Please note, `disc.classify` = $\sigma(D_\omega)$ in `loss_disc`.
# + cellView="form"
# @markdown *Execute this cell to enable helper functions*
def get_data(num_samples=100, seed=0):
set_seed(seed)
return torch.randn([num_samples, 1])
class DummyGen:
def sample(self, num_samples=100, seed=1):
set_seed(seed)
return torch.randn([num_samples, 1]) + 2
class DummyDisc:
def classify(self, x, seed=0):
set_seed(seed)
return torch.rand([x.shape[0], ])
# +
def loss_disc(disc, x_real, x_fake):
"""Compute the discriminator loss for `x_real` and `x_fake` given `disc`
Args:
disc: The discriminator
x_real (ndarray): An array of shape (N,) that contains the real samples
x_fake (ndarray): An array of shape (N,) that contains the fake samples
Returns:
ndarray: The discriminator loss
"""
label_real = 1
#################################################
# TODO for students: Loss for real data
raise NotImplementedError("Student exercise: Implement loss for real samples")
#################################################
loss_real = label_real * ...
label_fake = 0
#################################################
# TODO for students: Loss for fake data
raise NotImplementedError("Student exercise: Implement loss for fake samples")
#################################################
loss_fake = ... * torch.log(1 - disc.classify(x_fake))
return torch.cat([loss_real, loss_fake])
disc = DummyDisc()
gen = DummyGen()
x_real = get_data()
x_fake = gen.sample()
## Uncomment to check your function
# ld = loss_disc(disc, x_real, x_fake)
# plotting_ld(ld)
# + [markdown] colab_type="text"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content-dl/tree/main//tutorials/W2D5_GenerativeModels/solutions/W2D5_Tutorial2_Solution_393f89f3.py)
#
# *Example output:*
#
# <img alt='Solution hint' align='left' width=971.0 height=972.0 src=https://raw.githubusercontent.com/NeuromatchAcademy/course-content-dl/main/tutorials/W2D5_GenerativeModels/static/W2D5_Tutorial2_Solution_393f89f3_1.png>
#
#
# -
# **A note on numerical stability**
#
# It is common that functions like $\log$ throw a numerical error.
# For $\log$, it happens when $x$ in $\log(x)$ is very close to 0.
# The most common practice is to always add some very small value $\epsilon$ to $x$, i.e. use $\log(x + \epsilon)$ instead.
# Most build-in functions in modern DL frameworks like TensorFlow or PyTorch handle such things in their build-in loss already, e.g., `torch.nn.BCE`, which is equivalent to the loss you implemented above.
# ## Section 3.2: Density ratio estimation and the optimal discriminator
# As explained in the lecture, the critic in GAN that trains as a binary classifier, upon training becomes a *density ratio estimator* of $\frac{p}{q}$.
#
# The estimated density ratio allows for quantifying how far the real and generator distributions are from each other using $f$-divergence. The generator minimizes this estimated $f$-divergence during training.
#
# We will now train a discriminator to see how it estimates the density ratio between two distributions.
# ### Coding Exercise 3.2: Estimating the density ratio by the discriminator
#
# We have provided an implementation of a binary critic in the class `OptimalDisc`.
#
# __Your goal__ is to complete the implementation of the function `ratio_disc` below using the function `classify` from `OptimalCritic` class.
#
# Upon correct implementation, you should see that the plot of the ratios from the optimal discriminator align to the true density ratio.
#
# __Remember, $\frac{p}{p + q} = \sigma(D_\omega(x))$__
# + cellView="form"
# @markdown *Execute this cell to enable the optimal discriminator: `OptimalDisc`*
class OptimalDisc:
def classify(self, x):
dist_p = torch.distributions.normal.Normal(loc=0, scale=1)
dist_q = torch.distributions.normal.Normal(loc=-2, scale=1)
prob_p = torch.exp(dist_p.log_prob(x))
prob_q = torch.exp(dist_q.log_prob(x))
return prob_p / (prob_p + prob_q)
# +
def ratio_disc(disc, x_real, x_fake):
"""Compute the density ratio between real distribution and fake distribution for `x`
Args:
disc: The discriminator
x (ndarray): An array of shape (N,) that contains the samples to evaluate
Returns:
ndarray: The density ratios
"""
# Put samples together
x = torch.cat([x_real, x_fake])
#################################################
# TODO for students: Compute p / (p + q) i.e. p(D=1|x)
raise NotImplementedError("Student exercise: Implement p_over_pplusq")
#################################################
p_over_pplusq = ...
#################################################
# TODO for students: Compute q / (p + q) i.e. 1 - p(D=1|x)
raise NotImplementedError("Student exercise: Implement q_over_pplusq")
#################################################
q_over_pplusq = ...
# Compute p / q
p_over_q = p_over_pplusq / q_over_pplusq
return p_over_q
disc = OptimalDisc()
gen = DummyGen()
x_real = get_data(1_000)
x_fake = gen.sample(1_000)
## Uncomment below to check your function
# ratio = ratio_disc(disc, x_real, x_fake)
# plotting_ratio(x_real, x_fake, ratio)
# + [markdown] colab_type="text"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content-dl/tree/main//tutorials/W2D5_GenerativeModels/solutions/W2D5_Tutorial2_Solution_007bde00.py)
#
# *Example output:*
#
# <img alt='Solution hint' align='left' width=1979.0 height=972.0 src=https://raw.githubusercontent.com/NeuromatchAcademy/course-content-dl/main/tutorials/W2D5_GenerativeModels/static/W2D5_Tutorial2_Solution_007bde00_1.png>
#
#
# -
# ## Section 3.3: The generator loss
#
# Now that we have a trained critic, lets see how to train the generator using it.
# ### Coding Exercise 3.3: The generator loss
#
# We will now implement the generator loss function and evaluate it on some fixed points.
#
# **Your goal** is to complete the implementation of the function `loss_gen` using the optimal critic from above.
#
# Upon correct implementation, you shall see a plot where the loss values from generator samples align with the "Correct" values.
#
# **HINT:** You simply need to change the labels.
# +
def loss_gen(disc, x_fake):
"""Compute the generator loss for `x_fake` given `disc`
Args:
disc: The generator
x_fake (ndarray): An array of shape (N,) that contains the fake samples
Returns:
ndarray: The generator loss
"""
#################################################
# TODO for students: Loss for fake data
raise NotImplementedError("Student exercise: Implement loss for fake data")
#################################################
label_fake = ...
loss_fake = label_fake * ...
return loss_fake
disc = DummyDisc()
gen = DummyGen()
x_fake = gen.sample()
## Uncomment below to check your function
# lg = loss_gen(disc, x_fake)
# plotting_lg(lg)
# + [markdown] colab_type="text"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content-dl/tree/main//tutorials/W2D5_GenerativeModels/solutions/W2D5_Tutorial2_Solution_04a08f8f.py)
#
# *Example output:*
#
# <img alt='Solution hint' align='left' width=971.0 height=972.0 src=https://raw.githubusercontent.com/NeuromatchAcademy/course-content-dl/main/tutorials/W2D5_GenerativeModels/static/W2D5_Tutorial2_Solution_04a08f8f_1.png>
#
#
# -
# **Did you notice?**
#
# The loss you implemented for generator is essentially the part for real data in `loss_disc`, i.e., it is saying, "*the data I am feeding to you is real and not fake*".
# ---
# # Section 4: GAN training in action!
#
# In this section we will be playing with a complete implementation of GAN.
# ## Interactive Demo 4: GAN training in action
#
# + cellView="form"
# @markdown *Execute this cell to enable the implemented GAN*
# @markdown
# @markdown You are encouraged to take a look at the implementation as well.
from IPython.display import clear_output
class Generator(nn.Module):
def __init__(self, latent_dim, layers, output_activation=None):
super(Generator, self).__init__()
self.latent_dim = latent_dim
self.output_activation = output_activation
self._init_layers(layers)
def _init_layers(self, layers):
"""Initialize the layers and store as self.module_list."""
self.module_list = nn.ModuleList()
last_layer = self.latent_dim
for index, width in enumerate(layers):
self.module_list.append(nn.Linear(last_layer, width))
last_layer = width
if index + 1 != len(layers):
self.module_list.append(nn.LeakyReLU())
else:
if self.output_activation is not None:
self.module_list.append(self.output_activation())
def forward(self, input_tensor):
"""Forward pass; map latent vectors to samples."""
intermediate = input_tensor
for layer in self.module_list:
intermediate = layer(intermediate)
return intermediate
class Discriminator(nn.Module):
def __init__(self, input_dim, layers):
super(Discriminator, self).__init__()
self.input_dim = input_dim
self._init_layers(layers)
def _init_layers(self, layers):
"""Initialize the layers and store as self.module_list."""
self.module_list = nn.ModuleList()
last_layer = self.input_dim
for index, width in enumerate(layers):
self.module_list.append(nn.Linear(last_layer, width))
last_layer = width
if index + 1 != len(layers):
self.module_list.append(nn.LeakyReLU())
else:
self.module_list.append(nn.Sigmoid())
def forward(self, input_tensor):
"""Forward pass; map samples to confidence they are real [0, 1]."""
intermediate = input_tensor
for layer in self.module_list:
intermediate = layer(intermediate)
return intermediate
class VanillaGAN():
def __init__(self, generator, discriminator, noise_fn, data_fn,
batch_size=100, device='cpu', lr_d=1e-3, lr_g=2e-4):
self.generator = generator
self.generator = self.generator.to(device)
self.discriminator = discriminator
self.discriminator = self.discriminator.to(device)
self.noise_fn = noise_fn
self.data_fn = data_fn
self.batch_size = batch_size
self.device = device
self.criterion = nn.BCELoss()
self.optim_d = optim.Adam(discriminator.parameters(),
lr=lr_d, betas=(0.5, 0.999))
self.optim_g = optim.Adam(generator.parameters(),
lr=lr_g, betas=(0.5, 0.999))
self.target_ones = torch.ones((batch_size, 1)).to(device)
self.target_zeros = torch.zeros((batch_size, 1)).to(device)
def generate_samples(self, latent_vec=None, num=None):
"""Sample from the generator.
"""
num = self.batch_size if num is None else num
latent_vec = self.noise_fn(num) if latent_vec is None else latent_vec
with torch.no_grad():
samples = self.generator(latent_vec)
return samples
def real_data(self, num=None):
"""Real Data
"""
num = self.batch_size if num is None else num
with torch.no_grad():
samples = self.data_fn(num)
return samples
def train_step_generator(self):
"""Train the generator one step and return the loss."""
self.generator.zero_grad()
latent_vec = self.noise_fn(self.batch_size)
generated = self.generator(latent_vec)
classifications = self.discriminator(generated)
loss = self.criterion(classifications,
self.target_ones)
loss.backward()
self.optim_g.step()
return loss.item()
def train_step_discriminator(self):
"""Train the discriminator one step and return the losses."""
self.discriminator.zero_grad()
# real samples
real_samples = self.data_fn(self.batch_size)
pred_real = self.discriminator(real_samples)
loss_real = self.criterion(pred_real,
self.target_ones)
# generated samples
latent_vec = self.noise_fn(self.batch_size)
with torch.no_grad():
fake_samples = self.generator(latent_vec)
pred_fake = self.discriminator(fake_samples)
loss_fake = self.criterion(pred_fake,
self.target_zeros)
# combine
loss = (loss_real + loss_fake) / 2
loss.backward()
self.optim_d.step()
return loss_real.item(), loss_fake.item()
def train_step(self):
"""Train both networks and return the losses."""
loss_d = self.train_step_discriminator()
loss_g = self.train_step_generator()
return loss_g, loss_d
def train(mean=0., sigma=.2, device='cpu'):
epochs = 30
batches = 100
generator = Generator(2, [64, 32, 2])
discriminator = Discriminator(2, [64, 32, 1])
noise_fn = lambda x: torch.rand((x, 2), device=device)
data_fn = lambda x: mean + sigma*torch.randn((x, 2), device=device)
gan = VanillaGAN(generator,
discriminator,
noise_fn,
data_fn,
lr_d=1e-3,
lr_g=2e-4,
device=device)
loss_g, loss_d_real, loss_d_fake = [], [], []
start = time()
fig, ax = plt.subplots(1, 1,figsize=(5,5))
x, y = np.random.random((2, 200))
data_scat = ax.scatter(x, y,
label='Data',
alpha=0.9,
s=10.,
c='r')
gan_scat = ax.scatter(x,y,
label='GAN',
alpha=0.9,
s=10.,
c='b')
ax.legend()
ax.set_xlim(-2, 2)
ax.set_ylim(-2, 2)
for epoch in range(epochs):
loss_g_running = 0
loss_d_real_running = 0
loss_d_fake_running = 0
for batch in range(batches):
lg_, (ldr_, ldf_) = gan.train_step()
loss_g_running += lg_
loss_d_real_running += ldr_
loss_d_fake_running += ldf_
loss_g.append(loss_g_running / batches)
loss_d_real.append(loss_d_real_running / batches)
loss_d_fake.append(loss_d_fake_running / batches)
print(f"Epoch {epoch+1}/{epochs} ({int(time() - start)}s):"
f" G={loss_g[-1]:.3f},"
f" Dr={loss_d_real[-1]:.3f},"
f" Df={loss_d_fake[-1]:.3f}")
gan_scat.set_offsets(gan.generate_samples())
data_scat.set_offsets(gan.real_data())
clear_output(wait=True)
display(fig)
plt.close(1)
# -
# Notice in the implementaiton above, we use `torch.nn.BCELoss` to implement the discriminator and generator loss with proper "labels".
# They are actually equivalent to your implementation - do you want to varify this fact?
# + cellView="form"
# @title GAN demo
# @markdown Make sure you execute this cell to enable the widget!
import ipywidgets
# https://ipywidgets.readthedocs.io/en/latest/examples/Widget%20List.html
slider_mean = ipywidgets.FloatSlider(value=0., min=-1, max=1,
step=.1, readout_format='.5f',
description="Set Target Mean",
style={'description_width': 'initial'})
slider_sigma = ipywidgets.FloatSlider(value=.2, min=.2, max=1.,
step=.1, readout_format='.5f',
description="Set Target Sigma",
style={'description_width': 'initial'})
button = ipywidgets.Button(description="Start Training!")
output = ipywidgets.Output()
def on_button_clicked(b):
# Display the message within the output widget.
with output:
train(slider_mean.value, slider_sigma.value)
button.on_click(on_button_clicked)
display(slider_mean, slider_sigma, button, output)
# -
# ---
# # Summary
#
# Through this tutorial, we have learned
#
# - How to implement the training loop of GANs.
# - Developed an intuition about the training dynamics of GANs.
# - How to implement the training objectives for the generator and discriminator of GANs.
# - How are GANs connected to density ratio estimation.
#
# Next tutorial will cover conditional GANs and ethical issues of DL.
| tutorials/W2D5_GenerativeModels/student/W2D5_Tutorial2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import sys
sys.path.append("../tests")
sys.path.append("../implicit")
from implicit.als import AlternatingLeastSquares
from implicit.approximate_als import (AnnoyAlternatingLeastSquares, FaissAlternatingLeastSquares,
NMSLibAlternatingLeastSquares)
from implicit.bpr import BayesianPersonalizedRanking
from implicit.nearest_neighbours import (BM25Recommender, CosineRecommender,
TFIDFRecommender, bm25_weight)
from implicit.evaluation import precision_at_k, train_test_split, mean_average_precision_at_k
from twitter import get_twitter, read_data
from implicit.datasets.lastfm import get_lastfm
from recommender_base import RandomRecommender
from recommender_base_test import TestRecommenderBaseMixin
# %load_ext autoreload
# %autoreload 2
# +
# maps command line model argument to class name
MODELS = {"als": AlternatingLeastSquares,
"nmslib_als": NMSLibAlternatingLeastSquares,
"annoy_als": AnnoyAlternatingLeastSquares,
"faiss_als": FaissAlternatingLeastSquares,
"tfidf": TFIDFRecommender,
"cosine": CosineRecommender,
"bpr": BayesianPersonalizedRanking,
"bm25": BM25Recommender}
def get_model(model_name):
model_class = MODELS.get(model_name)
if not model_class:
raise ValueError("Unknown Model '%s'" % model_name)
# some default params
if issubclass(model_class, AlternatingLeastSquares):
params = {'factors': 64, 'dtype': np.float32}
elif model_name == "bm25":
params = {'K1': 100, 'B': 0.5}
elif model_name == "bpr":
params = {'factors': 63}
else:
params = {}
return model_class(**params)
# -
def evaluate_model(model_name="als", dataset='twitter'):
"""evaluate the model by cross-validation"""
# train the model based off input params
if dataset is 'twitter':
artists, users, plays = get_twitter()
if dataset is 'lastfm':
artists, users, plays = get_lastfm()
# create a model from the input data
model = CosineRecommender()
# split data_set to train set and testing set
train, test = train_test_split(plays)
print(train.shape)
print(test.shape)
# evaluation
p = precision_at_k(model, train.T.tocsr(), test.T.tocsr(), K=20, num_threads=4)
print('precision@k = ', p)
# +
artists, users, ratings = get_twitter()
train, test = train_test_split(ratings, train_percentage=0.8)
model_names = ['als',
# 'nmslib_als', 'annoy_als', 'faiss_als',
'tfidf', 'cosine', 'bpr', 'bm25']
twitter_dict_mean_pk = dict()
for model_name in model_names:
model = get_model(model_name)
# model = TFIDFRecommender()
model.fit(train)
p = mean_average_precision_at_k(model, train.T.tocsr(), test.T.tocsr(), K=20, num_threads=4)
twitter_dict_mean_pk[model_name] = p
print('model: ', model_name, 'p@k', p)
# -
twitter_dict_pk
twitter_dict_mean_pk
lastfm_dict
# ## Test RandomRecommender
artists, users, plays = get_twitter()
model = RandomRecommender()
train, test = train_test_split(plays, train_percentage=0.8)
model.fit(train)
mean_average_precision_at_k(model, train.T.tocsr(), test.T.tocsr(), K=20, num_threads=4)
| evalutation_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="XZgrRx-yUjRq"
# # IA Heart Disease Dataset from UCI
#
# + colab={"base_uri": "https://localhost:8080/", "height": 399} colab_type="code" id="caCoDFCpUjR3" outputId="8f551aee-1f07-4f62-c274-93b06fe2ac3f"
# !pip install pydotplus
# !pip install graphviz
# !pip install matplotlib
# !pip install scipy
# !pip install sklearn
# !pip install pandas
# !pip install numpy
# + [markdown] colab_type="text" id="aH8rc3UeUjSV"
# link al conjunto de datos https://www.kaggle.com/ronitf/heart-disease-uci
# + [markdown] colab_type="text" id="O_BVh0hXUjS9"
# ## Información provista sobre el conjunto de datos
#
# Citado de el link provisto por el profesor
#
# This database contains 76 attributes, but all published experiments refer to using a subset of 14 of them. In particular, the Cleveland database is the only one that has been used by ML researchers to this date. The "goal" field refers to the presence of heart disease in the patient. It is integer valued from 0 (no presence) to 4. Experiments with the Cleveland database have concentrated on simply attempting to distinguish presence (values 1,2,3,4) from absence (value 0).
#
# Complete attribute documentation:
#
# 1. age: age in years
# 2. sex: sex (1 = male; 0 = female)
# 3. cp: chest pain type <br>
# -- Value 1: typical angina<br>
# -- Value 2: atypical angina<br>
# -- Value 3: non-anginal pain<br>
# -- Value 4: asymptomatic<br>
# 4. trestbps: resting blood pressure (in mm Hg on admission to the hospital)
# 5. chol: serum cholestoral in mg/dl Y
# 6. fbs: (fasting blood sugar > 120 mg/dl) (1 = true; 0 = false)
# 7. restecg: resting electrocardiographic results<br>
# -- Value 0: normal<br>
# -- Value 1: having ST-T wave abnormality (T wave inversions and/or ST elevation or depression of > 0.05 mV)<br>
# -- Value 2: showing probable or definite left ventricular hypertrophy by Estes' criteria<br>
# 8. thalach: maximum heart rate achieved
# 9. exang: exercise induced angina (1 = yes; 0 = no)
# 10. oldpeak = ST depression induced by exercise relative to rest
# 11. slope: the slope of the peak exercise ST segment<br>
# -- Value 1: upsloping<br>
# -- Value 2: flat<br>
# -- Value 3: downsloping
# 12. ca: number of major vessels (0-3) colored by flourosopy
# 13. thal:
# -- 3 = normal
# -- 6 = fixed defect
# -- 7 = reversable defect
# 14. num: diagnosis of heart disease (angiographic disease status)
# -- Value 0: < 50% diameter narrowing
# -- Value 1: > 50% diameter narrowing (in any major vessel: attributes
# + colab={} colab_type="code" id="vJ9YIroHUjSa"
import matplotlib
from matplotlib import pyplot as plt
import pandas as pd
import numpy as np
import argparse
import glob
from pandas import DataFrame
import sklearn
from sklearn import tree, metrics
from sklearn.model_selection import train_test_split
from sklearn.metrics import confusion_matrix
from sklearn.naive_bayes import GaussianNB
#Necesario para graficar el arbol de decision
from sklearn.externals.six import StringIO
from IPython.display import Image
from sklearn.tree import export_graphviz
import pydotplus
# + [markdown] colab_type="text" id="4UJIzm6TUjSq"
# importando los datos
# + colab={} colab_type="code" id="QT0iBUTwUjSw"
dataset=pd.read_csv("heart.csv")
# -
dataset.head(10)
# + [markdown] colab_type="text" id="IdgSR0y0UjTS"
# Graficaremos las variables numericas y categorias
# + [markdown] colab_type="text" id="fYyH8-oqUjTW"
# ## Variables Categoricas
#
# Las variables categóricas también se denominan variables cualitativas o variables de atributos. Los valores de una variable categórica son categorías o grupos mutuamente excluyentes. Los datos categóricos pueden tener o no tener un orden lógico.
#
# ## Variables numericas
#
# Las variables numericas también se denominan variables cuantitativas, los datos numericos describen una caracteristica en terminos de un valor numerico.
#
# ## Clasificando las variables
#
# Dadas las descripciones dadas de los datos del dataset y la definición de lo que es una variable categorica podemos decir que los siguientes atributos son de tipo categorico
# - sex
# - cp
# - fbs
# - restecg
# - exang
# - slope
# - thal
# <br>y las siguientes son de tipo numerico
# - age
# - trestbps
# - chol
# - thalach
# - oldpeak
# - ca
# + colab={"base_uri": "https://localhost:8080/", "height": 322} colab_type="code" id="V_2UKMFuUjTd" outputId="6a28bb5a-4bcb-4906-e61a-808f1d4be6b6"
fig, axs = plt.subplots(2, 7, figsize = (15,5))
#numeric
#age
axs[0, 0].hist(dataset.age)
axs[0, 0].set_title('age')
#trestbps
axs[0, 1].hist(dataset.trestbps)
axs[0, 1].set_title('trestbps')
#chol
axs[0, 2].hist(dataset.chol)
axs[0, 2].set_title('chol')
#thalach
axs[0, 3].hist(dataset.thalach)
axs[0, 3].set_title('thalach')
#oldpeak
axs[0, 4].hist(dataset.oldpeak)
axs[0, 4].set_title('oldpeak')
# #cat
axs[0, 5].hist(dataset.ca)
axs[0, 5].set_title('cat')
#categoric
#sex
datasex = [dataset.sex.sum(), dataset.sex.shape[0] - dataset.sex.sum()]
axs[1, 0].pie(datasex, labels = ['m','f'])
axs[1, 0].axis('equal')
axs[1, 0].set_title('sex')
# #cp
datacp = []
datacp.append((dataset['cp'] == 0).sum())
datacp.append((dataset['cp'] == 1).sum())
datacp.append((dataset['cp'] == 2).sum())
datacp.append((dataset['cp'] == 3).sum())
axs[1, 1].pie(datacp, labels =['0','1','2','3'])
axs[1, 1].axis('equal')
axs[1, 1].set_title('cp')
#fbs
datafbs = [dataset.fbs.sum(), dataset.fbs.shape[0] - dataset.fbs.sum()]
axs[1, 2].pie(datafbs, labels = ['1', '0'])
axs[1, 2].axis('equal')
axs[1, 2].set_title('fbs')
#restecg
datarestecg = []
datarestecg.append((dataset['restecg'] == 0).sum())
datarestecg.append((dataset['restecg'] == 1).sum())
datarestecg.append((dataset['restecg'] == 2).sum())
axs[1, 3].pie(datarestecg, labels = ['0', '1', '2'])
axs[1, 3].axis('equal')
axs[1, 3].set_title('restecg')
#exang
dataexang = [dataset.exang.sum(), dataset.exang.shape[0] - dataset.exang.sum()]
axs[1, 4].pie(dataexang, labels = ['1','0'])
axs[1, 4].axis('equal')
axs[1, 4].set_title('exang')
#slope
dataslope = []
dataslope.append((dataset['slope'] == 0).sum())
dataslope.append((dataset['slope'] == 1).sum())
dataslope.append((dataset['slope'] == 2).sum())
axs[1, 5].pie(dataslope, labels=['0','1','2'])
axs[1, 5].axis('equal')
axs[1, 5].set_title('slope')
#thal
datathal = []
datathal.append((dataset['thal'] == 0).sum())
datathal.append((dataset['thal'] == 1).sum())
datathal.append((dataset['thal'] == 2).sum())
datathal.append((dataset['thal'] == 3).sum())
axs[1, 6].pie(datathal, labels = ['0','1','2','3'])
axs[1, 6].axis('equal')
axs[1, 6].set_title('thal')
y = ['numeric','categoric']
i = 0
for ax in axs.flat:
if (i < 6):
ax.set(xlabel = '', ylabel=y[0])
else:
ax.set(xlabel = '', ylabel=y[1])
i += 1
# Hide x labels and tick labels for top plots and y ticks for right plots.
for ax in axs.flat:
ax.label_outer()
fig.delaxes(axs.flatten()[6])
# + [markdown] colab_type="text" id="fkKABNlqUjTq"
# ## Elementos faltantes
#
# Veremos si hay datos nulos en el conjunto de datos
# -
print(dataset.isnull().sum().sum())
# Como no hay datos nulos entonces no hay necesidad de llenarlo u obviarlos de los siguientes pasos.
# + [markdown] colab_type="text" id="WiNJX6BmUjTu"
# ## Partimos los datos en entrenamiento y test
# 80% para entrenamiento y 20% para test
# + colab={"base_uri": "https://localhost:8080/", "height": 52} colab_type="code" id="OXc3nhwWUjT0" outputId="fad773fe-af25-4fae-e148-6bddcb8a81bb"
columnsTitles = ["age","sex","cp","trestbps","chol","fbs","restecg","thalach","exang","oldpeak","slope","ca","thal"]
features = dataset[columnsTitles].values
target = dataset["target"].values
# Split the data into train and test
trainX, testX, trainY, testY = train_test_split(features, target, test_size=0.2)
print("Conjunto de entrenamiento: ")
print("cantidad de registros: ", trainX.shape[0])
print("cantidad de columnas: ", trainX.shape[1])
print("cantidad de registro target: ", trainY.shape[0])
print("\nConjunto de prueba: ")
print("cantidad de registros: ", testX.shape[0])
print("cantidad de columnas: ", testX.shape[1])
print("cantidad de registros target: ", testY.shape[0])
# + [markdown] colab_type="text" id="Hu6B70a5UjUB"
# ## Arbol de decision y matriz de confusión
#
# entrenamos el arbol de decisión.
# + colab={"base_uri": "https://localhost:8080/", "height": 89} colab_type="code" id="HKOBl88IUjUF" outputId="e58481f6-7803-4751-e022-3d38e33ed227"
#Control overfitting by setting "max_depth" to 10 and "min_samples_split" to 5 : my_tree_two
modelDesicionTree = tree.DecisionTreeClassifier(max_depth=10, min_samples_split = 5, random_state = 42)
modelDesicionTree = modelDesicionTree.fit(trainX, trainY)
#Print the score on the train data
print("training set: ", modelDesicionTree.score(trainX, trainY)*100, "%")
#Print the score on the test data
print("test set: ", modelDesicionTree.score(testX, testY)*100, "%")
matrixDecisionTree = confusion_matrix(testY, modelDesicionTree.predict(testX))
print("confusion matrix: ")
print(matrixDecisionTree)
#graph of the confusion matrix
label = [0,1]
_X = np.arange(len(label))
x = [matrixDecisionTree[0][0], matrixDecisionTree[1][0]]
y = [matrixDecisionTree[0][1], matrixDecisionTree[1][1]]
training = plt.bar(_X-0.1,x, width=0.2, align='center')
test = plt.bar(_X+0.1,y, width=0.2, align='center')
plt.xticks(np.arange(2), ['no enfermos','enfermos'])
def autolabel(rects, bars):
for rect,bar in zip(rects,bars):
height = rect.get_height()
plt.text(rect.get_x() + rect.get_width() / 2, height + 0.1, bar,ha='center', va='bottom')
autolabel(training,["bien","mal"])
autolabel(test,["mal", "bien"])
plt.title('Matriz de Confusión')
# -
# ## Graficamos el arbol de decision
dot_data = StringIO()
export_graphviz(modelDesicionTree,
out_file=dot_data,
filled=True,
rounded=True,
feature_names = list(dataset.columns[:13]),
class_names = ['no disease','disease'],
special_characters=True)
graph = pydotplus.graph_from_dot_data(dot_data.getvalue())
Image(graph.create_png())
# + [markdown] colab_type="text" id="70Uf5IrLUjUn"
# # Interpretación del modelo obtenido
# -
# ## Entendiendo los contenidos de los nodos
# Para poder comprender nuestro arbol de decision, debemos analizar detalladamente los elementos presentes en cada nodo
#
# ### comparacion (Feature):
# En el primer elemento de arriba a abajo de cada nodo, se encuentra una operacion logica donde se evalua el valor de algun feature en especifico. Esta comparacion es basicamente una pregunta que se realiza a cada muestra (sample) presente en el nodo, cuando un sample cumple con la condicion dada este es enviado al nodo hijo izquierdo, en caso contrario es enviado al nodo hijo derecho
#
# ### gini
# El coeficiente de gini es una metrica que cuantifica la pureza de un nodo/hoja. Una coeficiente de gini mayor que cero implica que hay muestras contenidas en el nodo que pertenencen a clases diferentes. Un coeficiente de gini igual a cero significa que el nodo es puro, esto es que en un nodo existen muestras de una sola clase.
#
# ### samples
# Este valor es solo una suma de todas las muestras de cada clase contenidas en el nodo
#
# ### values
# Values es una lista que indica cuantas muestras en un nodo dado caen en cada categoria. En nuestro caso tenemos en la posicion 0 la categoria 'no disease/no enfermo' y en la 1 la categoria 'disease/enfermo'
# ## Divisiones del arbol
# Para determinar que feature usar para realizar la primer division (crear el nodo raiz) el algoritmo toma un feature y lo divide. Luego mira los subconjuntos y mide su impureza usando el coeficiente de gini. Esto es realizado para diferentes umbrales y luego determina que la mejor division para un feature dado es aquel que produce los subconjuntos mas puros. Este proceso es repetido para todos los features en el conjunto de entrenamiento. Por ultimo, el nodo raiz es determinado por el feature que produce una division con los subconjuntos mas puros. Una vez que el nodo raiz es decidido, el arbol crece a una profundidad de uno. Este proceso es repetido para los otros nodos en el arbol.
# ### Interpretacion de resultados
# En el arbol de decision se obtuvieron valores no muy precisos para el conjunto de entrenamiento y el de prueba, podemos afirmar que este modelo no resulto ser muy preciso al momento de predecir el target. Para el arbol de decisiones se probaron distintas configuraciones de las cuales gran parte no influenciaron los resultados del modelo. Sin embargo, se tenian casos donde los resultados parecian prometedores, la configuracion de dichos casos fue donde la profundidad del arbol era menor que 5, dicho resultado se lograba estableciendo el parametro ```max_depth``` directamente a valores menores que 5 o estableciendo ```min_samples_split``` a valores muy altos (mayores a 25). Para ambos casos la profundidad del arbol resultaba ser pequeña y el porcentaje de precision tanto del conjunto de prueba como el conjunto de entrenamiento se acercaban mucho. Sin embargo, observando el arbol generado por ambos casos, podemos afirmar las predicciones hechas por el modelo pueden resultar erradas esto debido a la poca cantidad de caracteristicas verificadas y a la alta impureza de los nodos hoja.
# ### Resultados para baja profundidad de arbol (max_depth)
# 
# 
# ### Resultados para muestras minimas por nodo
#
# 
# 
#
# + [markdown] colab_type="text" id="5u9fW7ApUjUs"
# ## Modelo de Ingenuo de Bayes
#
# Entrenamos modelo de bayes ingenuo
#
# + colab={"base_uri": "https://localhost:8080/", "height": 52} colab_type="code" id="bJEJ2_6XUjUw" outputId="92cce08d-fa10-4a4f-897d-158b46359bd9"
gnb = GaussianNB()
gaussianModel = gnb.fit(trainX, trainY)
#Print the score on the train data
print("training set: ", gaussianModel.score(trainX, trainY) * 100, "%")
#Print the score on the test data
print("test set: ", gaussianModel.score(testX, testY)*100, "%")
matrixGaussian = confusion_matrix(testY, gaussianModel.predict(testX))
print("confusion matrix: ")
print(matrixGaussian)
label = [0,1]
_X = np.arange(len(label))
x = [matrixGaussian[0][0], matrixGaussian[1][0]]
y = [matrixGaussian[0][1], matrixGaussian[1][1]]
training = plt.bar(_X-0.1,x, width=0.2, align='center')
test = plt.bar(_X+0.1,y, width=0.2, align='center')
plt.xticks(np.arange(2), ['no enfermos','enfermos'])
def autolabel(rects, bars):
for rect,bar in zip(rects,bars):
height = rect.get_height()
plt.text(rect.get_x() + rect.get_width() / 2, height + 0.1, bar,ha='center', va='bottom')
autolabel(training,["bien","mal"])
autolabel(test,["mal", "bien"])
plt.title('Matriz de Confusión')
# -
# ## Interpretación del modelo Ingenuo de Bayes
# La inferencia bayesiana es un método estadístico, en el que el teorema de Bayes se usa para actualizar la probabilidad de una hipótesis a medida que hay más evidencia o información disponible.
#
# En este caso, los datos de entrenamiento corresponden a una cantidad de datos considerables de la muestra. El algoritmo consiste en tomar los datos de entrenamiento y calcular la probabilidad de que el atributo target sea 1 ó 0 dados unos valores anteriores.
#
# Este metodo no tiene parametros modificables dada la naturaleza de su estrategia para determinar si hay o no enfermedades en el grupo de datos de prueba.
#
# Para las pruebas ejecutadas, el exito de las mismas, se encuentra alrededor del 80%, porcentaje cercano al relacionado con el entrenamiento, lo que podria ser interpretado como una consistencia a lo largo del conjunto de datos en la muestra, siendo razonable su probabilidad de acierto dados los datos anteriores para determinar la presencia de enfermedades cardiovasculares.
# + [markdown] colab_type="text" id="tGY_p6N0UjU5"
# ## Red Neuronal
# + colab={"base_uri": "https://localhost:8080/", "height": 173} colab_type="code" id="11pgdFfLUjU9" outputId="7e2788ce-8e0f-4d82-9a18-232cef93f758"
from sklearn.neural_network import MLPClassifier
from sklearn.preprocessing import StandardScaler
# NN is sensitive to data scale. We must normilize
scaler = StandardScaler()
trainXX = trainX.copy()
testXX = testX.copy()
# Don't cheat - fit only on training data
scaler.fit(trainX)
trainXX = scaler.transform(trainXX)
# apply same transformation to test data
testXX = scaler.transform(testXX)
modelNeuralNetwork = MLPClassifier(solver='lbfgs', alpha=1e-3, activation = 'relu', max_iter=5000,
hidden_layer_sizes = (12,10,8,6,4,2), random_state=1, verbose = True)
modelNeuralNetwork.fit(trainXX, trainY)
#Print the score on the train data
print("On training: ", modelNeuralNetwork.score(trainXX, trainY)*100, "%")
#matrix = confusion_matrix(model3.predict(trainXX), trainY)
#Print the score on the test data
print("\nOn test: ", modelNeuralNetwork.score(testXX, testY)*100, "%")
matrixNeuralNetwork = confusion_matrix( testY, modelNeuralNetwork.predict(testXX))
print("confusion matrix: ")
print(matrixNeuralNetwork)
label = [0,1]
_X = np.arange(len(label))
x = [matrixNeuralNetwork[0][0], matrixNeuralNetwork[0][1]]
y = [matrixNeuralNetwork[1][0], matrixNeuralNetwork[1][1]]
training = plt.bar(_X-0.1,x, width=0.2, align='center')
test = plt.bar(_X+0.1,y, width=0.2, align='center')
plt.xticks(np.arange(2), ['no enfermos','enfermos'])
def autolabel(rects, bars):
for rect,bar in zip(rects,bars):
height = rect.get_height()
plt.text(rect.get_x() + rect.get_width() / 2, height + 0.1, bar,ha='center', va='bottom')
autolabel(training,["bien","mal"])
autolabel(test,["mal", "bien"])
plt.title('Matriz de Confusión')
# + [markdown] colab={} colab_type="code" id="vci5KcqVUjVN"
# ## Interpretación del modelo de Redes Neuronales
# -
# <img src="mlp_network.png" alt="Figura 1: Perceptron multicapa" style="width: 400px;"/>
# Un perceptron multi capa es un algoritmo de aprendizaje supervizado que aprende una funcion $f(\cdot ):R^{m}\rightarrow R^{o}$ entrenandose con un conjunto de datos (dataset). donde $m$ es el numero de dimensiones de entrada y $o$ es el numero de dimensiones de salida. Dado un conjunto de catacteristicas $X=x_1,x_2,...,x_m$ y un target $y$, la red puede aprender un aproximador de funcion no lineal para clasificacion o regresion. Entre la capa de entrada y la de salida, puede haber una o más capas no lineales, llamadas capas ocultas. La Figura 1 muestra un MLP de una capa oculta con salida escalar.
# La capa más a la izquierda, conocida como capa de entrada, consiste en un conjunto de neuronas $\left \{ x_i|x_1, x_2, ..., x_m \right \}$ que representan los features de entrada. Cada neurona en la capa escondida transforma los valores de una capa anterior con una suma lineal ponderada $w_1x_1 +w_2x_2+...+w_mx_m$, seguida de una funcion de activacion no lineal - como la funcion tangente hiperbolica. La función de activación se encarga de devolver una salida a partir de un valor de entrada, normalmente el conjunto de valores de salida en un rango determinado como $(0,1)$ o $(-1,1)$. Se buscan funciones que las derivadas sean simples, para minimizar con ello el coste computacional.
# ## MLPClassifier
# La clase MLPclassifier implementa un algoritmo de perceptron multicapa que entrena usando retropropagacion
# ### Backpropagation
#
# Una vez que se ha aplicado un patrón a la entrada de la red como estímulo, este se propaga desde la primera capa a través de las capas siguientes de la red, hasta la ultima capa de la red donde se genera una salida. La señal de salida se compara con la salida deseada y se calcula una señal de error para cada una de las salidas.
#
# Esta señal de error se propagan hacia atrás, partiendo de la capa de salida, hacia todas las neuronas de la capa oculta que contribuyen directamente a la salida. Sin embargo las neuronas de la capa oculta solo reciben una fracción de la señal total del error, basándose aproximadamente en la contribución relativa que haya aportado cada neurona a la salida original. Este proceso se repite, capa por capa, hasta que todas las neuronas de la red hayan recibido una señal de error que describa su contribución relativa al error total.
#
# La importancia de este proceso consiste en que, a medida que se entrena la red, las neuronas de las capas intermedias se organizan a sí mismas de tal modo que las distintas neuronas aprenden a reconocer distintas características del espacio total de entrada.
# ## Comparación de los 3 modelos
#
# Compare los resultados de los 3 modelos usados en términos de la precisión, la estabilidad y la interpretabilidad de los resultados.
#
# ## Precisión
#
# Cuando predice positivo, porcentaje clasificado correctamente ${vp} / {prediccion positivos}$, se busca un valor cercano a 1
#
print("precision arbol de decision",matrixDecisionTree[0][0] / (matrixDecisionTree[0][0] + matrixDecisionTree[1][0]))
print("precision modelo de bayes",matrixGaussian[0][0]/(matrixGaussian[0][0]+ matrixGaussian[1][0]))
print("precision red neuronal",matrixNeuralNetwork[0][0] / (matrixNeuralNetwork[0][0] + matrixNeuralNetwork[1][0]))
# ## Estabilidad
#
# Diferencia entre el puntaje del conjunto de entrenamiento y el conjunto de prueba, se busca que sea pequeña
print("estabilidad arbol de decision",abs(modelDesicionTree.score(trainX, trainY)*100 - modelDesicionTree.score(testX, testY)*100))
print("estabilidad modelo de bayes",abs(gaussianModel.score(trainX, trainY) * 100 - gaussianModel.score(testX, testY) * 100))
print("estabilidad red neuronal",abs(modelNeuralNetwork.score(trainXX, trainY)*100 - modelNeuralNetwork.score(testXX, testY)*100))
# ## Interpretabilidad
#
# En cuanto a la interpretabilidad de las estrategias para la predicción de la enfermedad con el conjunto de datos dado, podemos notar por un lado la cantidad de variables que se presentan con el árbol de decisión y más aún con las redes neuronales, donde las variaciones generan mejoras o desaciertos en los resultados deseados, siendo muy dificil la tarea de ajustar correctamente cada uno de estos dos modelos. Por otro lado, el modelo ingenuo de Bayes, donde no es necesaria la configuración del modelo, debido a que su estrategia funciona acorde al calculo de probabilidades de un evento, dada la ocurrencia de otros más, parece ir acorde al problema planteado, su interpretabilidad es muy natural y sus resultados son satisfactorios en comparación a los demás.
# ## Comparación
#
# Primero se busco cuales son los factores que pueden provocar enfermedades del corazon y si alguna esta presente entre los datos proporcionados por el conjunto de datos, entre ellos encontramos:
#
# - hipertension - presente en el conjunto de datos
# - nivel de colesterol alto - presente en el conjunto de datos
# - diabetes - presente en el conjunto de datos
# - obesidad y sobrepeso
# - tabaquismo
# - inactividad fisica
# - sexo masculino - presente en el conjunto de datos
# - hereditario
# - edad avanzada - presente en el conjunto de datos
#
# entre otros
#
# [fuente](https://www.texasheart.org/heart-health/heart-information-center/topics/factores-de-riesgo-cardiovascular/)
#
# Algunos de los datos en el conjunto de datos son factores de riesgo reconocidos hoy en dia para el diagnostico.
#
# Revisando los resultados obtenidos de precision y estabilidad, el arbol de decision tanto en precision como en estabilidad presenta resultados no cercanos al deseado, y tanto los del modelo ingenuo de bayes como redes neuronales presentan resultados en estos aceptables.
#
# Respecto al modelo ingenuo de bayes ya que es un metodo estadistico, y entre los datos habia factores de riesgo, era un metodo apropiado y tenia los factores para funcionar correctamente.
| IA (copia).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="AHtp6jeDvVjb" outputId="0787acbf-e9d3-46d9-9b64-600b5c62aa6a"
# !pip install selenium
# !apt-get update
# !apt install chromium-chromedriver
# + colab={"base_uri": "https://localhost:8080/"} id="utBGU9H91uUD" outputId="9a5c45b3-34a2-4cfa-abcb-09e9ba1ad180"
from selenium import webdriver
from selenium.webdriver.support import expected_conditions as EC
import os
chrome_options = webdriver.ChromeOptions()
chrome_options.add_argument('--headless')
chrome_options.add_argument('--no-sandbox')
chrome_options.add_argument('--disable-dev-shm-usage')
wd = webdriver.Chrome('chromedriver',chrome_options=chrome_options)
driver = webdriver.Chrome('chromedriver',chrome_options=chrome_options)
# + id="W2Q2jwkKt5k_" colab={"base_uri": "https://localhost:8080/"} outputId="7e263dc8-76ef-4f09-9efe-261c1a840f14"
folder = "/content/screen/"
urls = get_urls()
for url in urls:
file_name = url.split("//")[1] + ".png"
file_name = file_name.replace("/","-")
try:
driver.get(url)
except Exception:
continue
el = driver.find_element_by_tag_name('html')
width = 1920
height = 1080
driver.set_window_size(width, height)
path = folder + file_name
driver.implicitly_wait(10)
try:
driver.save_screenshot(path)
print("Saved png for " + url)
except Exception:
height = 1080
driver.set_window_size(width, height)
driver.save_screenshot(path)
try:
make_elements(driver, height, file_name)
except Exception:
print("Removing " + path)
os.remove(path)
# + id="TgavuhLxMylx"
def make_elements(driver, height, file_name):
cords = []
buttons = driver.find_elements_by_tag_name("button")
inputs = driver.find_elements_by_tag_name("input")
ejs = driver.find_elements_by_tag_name("a")
elements = buttons + inputs + ejs
for element in elements:
if element.rect["y"] + element.rect["height"] < height and element.rect["height"] > 0 and element.rect["width"] > 0 and element.rect["x"] + element.rect["width"] > 0 and element.is_displayed():
cords.append(make_cords(element, height))
cords_to_file(cords, file_name)
print("Saved cords for " + file_name)
# + id="ZV_gJsPelfwt"
def make_cords(element, height):
x = (element.rect["x"] + element.rect["x"] + element.rect["width"])/2
y = (element.rect["y"] + element.rect["y"] + element.rect["height"])/2
cords = []
cords.append(0)
cords.append(x/1920)
cords.append(y/height)
cords.append(element.rect["width"]/1920)
cords.append(element.rect["height"]/height)
return cords
# + id="fSB2El7Tn0Dx"
def cords_to_file(cords, file_name):
file_real_name = file_name.split(".png")[0]
path = folder + file_real_name + ".txt"
f = open(path, "w")
for cord in cords:
text = str(cord[0]) + " " + str(cord[1]) + " " + str(cord[2]) + " " + str(cord[3]) + " " + str(cord[4]) + "\n"
f.write(text)
f.close()
# + id="sK1iEcnUhaVx"
def get_urls():
path = "/content/url.txt"
file = open(path, "r")
urls = [line.rstrip('\n') for line in file]
return urls
# + id="ObzVJTyZxUJd"
driver.quit()
# + colab={"base_uri": "https://localhost:8080/"} id="3vV15vbna3CJ" outputId="baef6a33-22d7-4000-ad00-4f9b3f4ff822"
# !zip -r /content/file.zip /content/screen
# + [markdown] id="5GWrfsjexDBu"
#
| Screen_full_webpageV2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python
# language: python
# name: conda-env-python-py
# ---
# <center>
# <img src="https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBMDeveloperSkillsNetwork-PY0101EN-SkillsNetwork/IDSNlogo.png" width="300" alt="cognitiveclass.ai logo" />
# </center>
#
# # Reading Files Python
#
# Estimated time needed: **40** minutes
#
# ## Objectives
#
# After completing this lab you will be able to:
#
# * Read text files using Python libraries
#
# <h2>Table of Contents</h2>
# <div class="alert alert-block alert-info" style="margin-top: 20px">
# <ul>
# <li><a href="https://download/?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDeveloperSkillsNetworkPY0101ENSkillsNetwork19487395-2021-01-01">Download Data</a></li>
# <li><a href="https://read/?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDeveloperSkillsNetworkPY0101ENSkillsNetwork19487395-2021-01-01">Reading Text Files</a></li>
# <li><a href="https://better/?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDeveloperSkillsNetworkPY0101ENSkillsNetwork19487395-2021-01-01">A Better Way to Open a File</a></li>
# </ul>
#
# </div>
#
# <hr>
#
# <h2 id="download">Download Data</h2>
#
import urllib.request
url = 'https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBMDeveloperSkillsNetwork-PY0101EN-SkillsNetwork/labs/Module%204/data/example1.txt'
filename = 'Example1.txt'
urllib.request.urlretrieve(url, filename)
# +
# Download Example file
# !wget -O /resources/data/Example1.txt https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBMDeveloperSkillsNetwork-PY0101EN-SkillsNetwork/labs/Module%204/data/example1.txt
# -
# <hr>
#
# <h2 id="read">Reading Text Files</h2>
#
# One way to read or write a file in Python is to use the built-in <code>open</code> function. The <code>open</code> function provides a **File object** that contains the methods and attributes you need in order to read, save, and manipulate the file. In this notebook, we will only cover **.txt** files. The first parameter you need is the file path and the file name. An example is shown as follow:
#
# <img src="https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBMDeveloperSkillsNetwork-PY0101EN-SkillsNetwork/labs/Module%204/images/ReadOpen.png" width="500" />
#
# The mode argument is optional and the default value is **r**. In this notebook we only cover two modes:
#
# <ul>
# <li>**r**: Read mode for reading files </li>
# <li>**w**: Write mode for writing files</li>
# </ul>
#
# For the next example, we will use the text file **Example1.txt**. The file is shown as follows:
#
# <img src="https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBMDeveloperSkillsNetwork-PY0101EN-SkillsNetwork/labs/Module%204/images/ReadFile.png" width="100" />
#
# We read the file:
#
# +
# Read the Example1.txt
example1 = "Example1.txt"
file1 = open(example1, "r")
# -
# We can view the attributes of the file.
#
# The name of the file:
#
# +
# Print the path of file
file1.name
# -
# The mode the file object is in:
#
# +
# Print the mode of file, either 'r' or 'w'
file1.mode
# -
# We can read the file and assign it to a variable :
#
# +
# Read the file
FileContent = file1.read()
FileContent
# -
# The **/n** means that there is a new line.
#
# We can print the file:
#
# +
# Print the file with '\n' as a new line
print(FileContent)
# -
# The file is of type string:
#
# +
# Type of file content
type(FileContent)
# -
# It is very important that the file is closed in the end. This frees up resources and ensures consistency across different python versions.
#
# +
# Close file after finish
file1.close()
# -
# <hr>
#
# <h2 id="better">A Better Way to Open a File</h2>
#
# Using the <code>with</code> statement is better practice, it automatically closes the file even if the code encounters an exception. The code will run everything in the indent block then close the file object.
#
# +
# Open file using with
with open(example1, "r") as file1:
FileContent = file1.read()
print(FileContent)
# -
# The file object is closed, you can verify it by running the following cell:
#
# +
# Verify if the file is closed
file1.closed
# -
# We can see the info in the file:
#
# +
# See the content of file
print(FileContent)
# -
# The syntax is a little confusing as the file object is after the <code>as</code> statement. We also don’t explicitly close the file. Therefore we summarize the steps in a figure:
#
# <img src="https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBMDeveloperSkillsNetwork-PY0101EN-SkillsNetwork/labs/Module%204/images/ReadWith.png" width="500" />
#
# We don’t have to read the entire file, for example, we can read the first 4 characters by entering three as a parameter to the method **.read()**:
#
# +
# Read first four characters
with open(example1, "r") as file1:
print(file1.read(4))
# -
# Once the method <code>.read(4)</code> is called the first 4 characters are called. If we call the method again, the next 4 characters are called. The output for the following cell will demonstrate the process for different inputs to the method <code>read()</code>:
#
# +
# Read certain amount of characters
with open(example1, "r") as file1:
print(file1.read(4))
print(file1.read(4))
print(file1.read(7))
print(file1.read(15))
# -
# The process is illustrated in the below figure, and each color represents the part of the file read after the method <code>read()</code> is called:
#
# <img src="https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBMDeveloperSkillsNetwork-PY0101EN-SkillsNetwork/labs/Module%204/images/read.png" width="500" />
#
# Here is an example using the same file, but instead we read 16, 5, and then 9 characters at a time:
#
# +
# Read certain amount of characters
with open(example1, "r") as file1:
print(file1.read(16))
print(file1.read(5))
print(file1.read(9))
# -
# We can also read one line of the file at a time using the method <code>readline()</code>:
#
# +
# Read one line
with open(example1, "r") as file1:
print("first line: " + file1.readline())
# -
# We can also pass an argument to <code> readline() </code> to specify the number of charecters we want to read. However, unlike <code> read()</code>, <code> readline()</code> can only read one line at most.
#
with open(example1, "r") as file1:
print(file1.readline(20)) # does not read past the end of line
print(file1.read(20)) # Returns the next 20 chars
# We can use a loop to iterate through each line:
#
# +
# Iterate through the lines
with open(example1,"r") as file1:
i = 0;
for line in file1:
print("Iteration", str(i), ": ", line)
i = i + 1
# -
# We can use the method <code>readlines()</code> to save the text file to a list:
#
# +
# Read all lines and save as a list
with open(example1, "r") as file1:
FileasList = file1.readlines()
# -
# Each element of the list corresponds to a line of text:
#
# +
# Print the first line
FileasList[0]
# -
# # Print the second line
#
# FileasList\[1]
#
# +
# Print the third line
FileasList[2]
# -
# <hr>
# <h2>The last exercise!</h2>
# <p>Congratulations, you have completed your first lesson and hands-on lab in Python.
# <hr>
#
# ## Author
#
# <a href="https://www.linkedin.com/in/joseph-s-50398b136/?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDeveloperSkillsNetworkPY0101ENSkillsNetwork19487395-2021-01-01" target="_blank"><NAME></a>
#
# ## Other contributors
#
# <a href="https://www.linkedin.com/in/jiahui-mavis-zhou-a4537814a?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDeveloperSkillsNetworkPY0101ENSkillsNetwork19487395-2021-01-01"><NAME></a>
#
# ## Change Log
#
# | Date (YYYY-MM-DD) | Version | Changed By | Change Description |
# | ----------------- | ------- | ------------- | --------------------------------------------------------- |
# | 2022-01-10 | 2.1 | Malika | Removed the readme for GitShare |
# | 2020-09-30 | 1.3 | Malika | Deleted exericse "Weather Data" |
# | 2020-09-30 | 1.2 | <NAME> | Weather Data dataset link added |
# | 2020-09-30 | 1.1 | <NAME> | Added exericse "Weather Data" |
# | 2020-09-30 | 1.0 | <NAME> | Added blurbs about closing files and read() vs readline() |
# | 2020-08-26 | 0.2 | Lavanya | Moved lab to course repo in GitLab |
# | | | | |
# | | | | |
#
# <hr/>
#
# ## <h3 align="center"> © IBM Corporation 2020. All rights reserved. <h3/>
#
| PY0101EN-4-1-ReadFile.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
3
12
1 1 2 2 3 2 3 6 6 6 7
# +
from collections import Counter
k=int(input())
n=int(input())
a=list(map(int,input().split()))
result = [item for items, c in Counter(a).most_common()
for item in [items] * c]
c=0
b=result.copy()
while len(b)>=k:
m=b[0]
while any(m==i for i in b) and len(b)>=k:
b.remove(m)
c+=1
if any(i in a for i in b):
c-=1
print(c)
# -
k=int(input())
n=int(input())
a=list(map(int,input().split()))
from collections import Counter
result = [item for items, c in Counter(a).most_common()
for item in [items] * c]
print(str(result))
a
c=0
b=result.copy()
while len(b)>=k:
m=b[0]
while any(m==i for i in b) and len(b)>=k:
b.remove(m)
c+=1
c
if any(i in a for i in b):
c-=1
c
b
result
while any(6==i for i in a ):
a.remove(6)
a
a
a.count(1)
s=list(set(a))
s
l=[a.count(i) for i in s]
l
max(l)
s[l.index(l.pop(max(l)))]
l
c=0
while len(a)>=k:
m=s[l.index(max(l))]
while any(m==i for i in a ):
a.remove(m)
c+=1
print(c)
import sys
sys.setrecursionlimit(100000)
k=input()
n=input()
l=[]
#a=map(int,raw_input().split())
if n==1:
print 0
elif k==1:
a=map(int,raw_input().split())
print n
else:
a=map(int,raw_input().split())
G={ i:[] for i in xrange(1,n+1)}
mark=[-1]*(n+1)
for i in xrange(2,len(a)+2):
G[a[i-2]].append(i)
G[i].append(a[i-2])
res=0
def dfs(x):
global res
s=1
mark[x]=0
for i in G[x]:
if mark[i]==-1:
s+=dfs(i)
mark[i]=0
if s>=k:
res+=1
s=0
return s
dfs(1)
print res
# +
import sys
sys.setrecursionlimit(100000)
k=input()
n=input()
l=[]
if n==1:
print 0
elif k==1:
a=map(int,raw_input().split())
print n
else:
a=map(int,raw_input().split())
G={ i:[] for i in xrange(1,n+1)}
mark=[-1]*(n+1)
for i in xrange(2,len(a)+2):
G[a[i-2]].append(i)
G[i].append(a[i-2])
res=0
def dfs(x):
global res
s=1
mark[x]=0
for i in G[x]:
if mark[i]==-1:
s+=dfs(i)
mark[i]=0
if s>=k:
res+=1
s=0
return s
dfs(1)
print(res)
# -
# # pass by
# ###
# - mutable --> ***pass by refrance*** ( List, Dictionary)
# - immutable --> ***pass by value*** ( Primitive Data types,int,float,string,tuple,boolean)
# refrence
def li(l):
# l*=2
# l=l*2
l.append(10)
print(l)
l=[1,2,3,4,5,6,7,8]
print(l)
li(l)
print(l)
t=1,2,3,4
def tup(t):
t*=2
print(t)
t=[1,2,3,4]
print(t)
tup(t)
print(t)
l*2
l1=[1,2,3,4]
# # Exceptional handling
try:
# l1[4]//0
# l1[2]//0
print(l1[4])
except ZeroDivisionError:
print('zero division error')
# except IndexError:
# print('Index error')
except:#default except should be in last
print('not specific error')
try:
print(23/32)
except Exception as e:
print(e)
except:
print('kesh')
else:
print('if there is no exception then this excuted')
print('it always be placed after all except block not before and after try,finally block')
finally:
print('it always excuted')
# # Regular Expression
import re
l='i am <NAME> and i am python developer'
re.search('python',l)
k=re.search('python',l).span()
k[1]-k[0]
if re.search('python',l):
print('pattern matched')
else:
print('pattern not matched')
if re.search('^i.*developer',l):
print('pattern matched')
else:
print('pattern not matched')
if re.search('developer$',l):
print('pattern matched')
else:
print('pattern not matched')
re.sub('developer','lover',l)
# # class
# ### Static
# - share data among different object of my class
# #### Static variable
class a:
x=23
def pass1(self):
a.x=28
a.x
a().pass1()
a.x
# +
class mobile:
discount=50
def __init__(self,price,brand):
self.price=price
self.brand=brand
def purchase(self):
print(self.brand,' mobile with price ',self.price,' is availabe after discount is : ',round((100-mobile.discount)*self.price/100,2))
def enable_discount():
mobile.discount=50
def disable_discount():
mobile.discount=0
mob1=mobile(2000,'Apple')
mob2=mobile(4000,'Samsung')
mob3=mobile(5000,'Nokia')
mob1.purchase()
disable_discount()
mob2.purchase()
enable_discount()
mob3.purchase()
# -
# #### Static method
# +
class mobile:
__discount=50
def __init__(self,price,brand):
self.price=price
self.brand=brand
def purchase(self):
print(self.brand,' mobile with price ',self.price,' is availabe after discount is : ',round((100-mobile.discount)*self.price/100,2))
@staticmethod
def get_discount():
return mobile.__discount
@staticmethod
def set_discount(discount):
mobile.__discount=discount
print(mobile.get_discount())
mobile.set_discount(48)
print(mobile.get_discount())
# -
# # Relationship
# - Inheritence => IS-A
# - Aggretation => HAS-A
# - Association => USES-A \
# ***AGGRETATION***
# +
class Customer:
def __init__(self, name, age, phone_no, address):
self.name = name
self.age = age
self.phone_no = phone_no
self.address = address
def view_details(self):
print (self.name, self.age, self.phone_no)
print (self.address.door_no, self.address.street, self.address.pincode)
def update_details(self,add):
self.address = add
class Address:
def __init__(self, door_no, street, pincode):
self.door_no = door_no
self.street = street
self.pincode = pincode
def update_address(self):
pass
add1=Address(123, "5th Lane", 56001)
add2=Address(567, "6th Lane", 82006)
cus1=Customer("Jack", 24, 1234, add1)
cus1.view_details()
cus1.update_details(add2)
cus1.view_details()
# -
# ##### weaker relationship
# +
class Customer:
def __init__(self, name, age, phone_no):
self.name = name
self.age = age
self.phone_no = phone_no
def purchase(self, payment):
if payment.type == "card":
print ("Paying by card")
elif payment.type == "e-wallet":
print ("Paying by wallet")
else:
print ("Paying by cash")
class Payment:
def __init__(self, type):
self.type = type
payment1=Payment("card")
c=Customer("Jack",23,1234)
c.purchase(payment1)
# -
# # Inheritence
# ***ex1***
class A:
def __init__(self):
print('A')
class B(A):
def __init__(self):
print('B',type(self))
super().__init__()
print(type(self))
B()
# ***ex - 2***
class A:
def __init__(self):
print('A')
print(type(self))
class B(A):
def __init__(self):
print('B')
super().__init__()
print(type(self))
B()
A()
class c:
__x=9
@staticmethod
def x1():
return c.__x
c.x1()
# ***ex-3***
class A:
def __init__(self):
print('A')
super().__init__()
class B(A):
def __init__(self):
print('B')
super().__init__()
class C(A):
def __init__(self):
print('C')
super().__init__()
class D(B):
def __init__(self):
print('D')
super().__init__()
class E(B,C):
def __init__(self):
print('E')
super().__init__()
class F(E,D):
def __init__(self):
print('F')
super().__init__()
class G(E):
def __init__(self):
print('G')
super().__init__()
class H(F,G):
def __init__(self):
print('H')
super().__init__()
H()
# ### Method overiding in python
# - In Python method overriding occurs simply defining in the child class a method with the same name of a method in the parent class.
# +
class Parent(object):
def __init__(self):
self.value = 5
def get_value(self):
return self.value
class Child(Parent):
def get_value(self):
return self.value + 1
c = Child()
c.get_value()
# -
# ### Abstract method
# - An abstract class should not be instantiated.
# - An abstract class may contain 0 or many abstract methods
from abc import ABCMeta,abstractmethod
# ***ex-1***
# - Even if one method is abstract, then we will get an error if we try to instantiate the class.
class a(metaclass=ABCMeta):
@abstractmethod
def return_policy(self):
pass
class b(a):
pass
b()
# ***ex-2***
# - If a method is abstract, then the Subclass must override the abstract method. Else we cannot instantiate the subclass also.
class a(metaclass=ABCMeta):
@abstractmethod
def return_policy(self):
pass
class b(a):
def return_policy(self):
print('Keshav')
b().return_policy()
# ***ex-3***
# - If a child class overrides the abstract method, then its own child classes need not override the abstract method
class a(metaclass=ABCMeta):
@abstractmethod
def return_policy(self):
pass
class b(a):
def return_policy(self):
print('Keshav')
class c(b):
pass
c()
# #### most important
# ***ex-4***
class a(metaclass=ABCMeta):
@abstractmethod
def return_policy(self):
pass
class b(a):
pass
class c(b):
pass
class d(c):
def return_policy(self):
print('Keshav')
c()
# ***quick summary***
# - Usually the parent class is an abstract class.
# - Abstract classes should not be instantiated.
# - If a class has an abstract method, then the class cannot be instantiated.
# - Abstract classes are meant to be inherited.
# - The child class must implement/override all the abstract methods of the parent class. Else the child class cannot be instantiated.
# ***ex-4***
import math
math.ceil(4.6)
math.floor(4.6)
round(2/3,2)
| Infosys/python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
x = list()
dir(x)
x.append('Book')
x.append('99')
print(x)
sum = [10, 13, 14]
10 in sum
12 not in sum
nums = [3, 41, 12, 9, 74, 15]
print(len(nums))
print(max(nums))
| Untitled1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] tags=["pdf-title"]
# # Network Visualization (TensorFlow)
#
# In this notebook we will explore the use of *image gradients* for generating new images.
#
# When training a model, we define a loss function which measures our current unhappiness with the model's performance; we then use backpropagation to compute the gradient of the loss with respect to the model parameters, and perform gradient descent on the model parameters to minimize the loss.
#
# Here we will do something slightly different. We will start from a convolutional neural network model which has been pretrained to perform image classification on the ImageNet dataset. We will use this model to define a loss function which quantifies our current unhappiness with our image, then use backpropagation to compute the gradient of this loss with respect to the pixels of the image. We will then keep the model fixed, and perform gradient descent *on the image* to synthesize a new image which minimizes the loss.
#
# In this notebook we will explore three techniques for image generation:
#
# 1. **Saliency Maps**: Saliency maps are a quick way to tell which part of the image influenced the classification decision made by the network.
# 2. **Fooling Images**: We can perturb an input image so that it appears the same to humans, but will be misclassified by the pretrained network.
# 3. **Class Visualization**: We can synthesize an image to maximize the classification score of a particular class; this can give us some sense of what the network is looking for when it classifies images of that class.
#
# This notebook uses **TensorFlow**; we have provided another notebook which explores the same concepts in PyTorch. You only need to complete one of these two notebooks.
# + tags=["pdf-ignore"]
# As usual, a bit of setup
import sys
sys.path.append('../')
import time, os, json
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
from cs231n.classifiers.squeezenet import SqueezeNet
from cs231n.data_utils import load_tiny_imagenet
from cs231n.image_utils import preprocess_image, deprocess_image
from cs231n.image_utils import SQUEEZENET_MEAN, SQUEEZENET_STD
# %matplotlib inline
plt.rcParams['figure.figsize'] = (10.0, 8.0) # set default size of plots
plt.rcParams['image.interpolation'] = 'nearest'
plt.rcParams['image.cmap'] = 'gray'
# for auto-reloading external modules
# see http://stackoverflow.com/questions/1907993/autoreload-of-modules-in-ipython
# %load_ext autoreload
# %autoreload 2
# + [markdown] tags=["pdf-ignore"]
# # Pretrained Model
#
# For all of our image generation experiments, we will start with a convolutional neural network which was pretrained to perform image classification on ImageNet. We can use any model here, but for the purposes of this assignment we will use SqueezeNet [1], which achieves accuracies comparable to AlexNet but with a significantly reduced parameter count and computational complexity.
#
# Using SqueezeNet rather than AlexNet or VGG or ResNet means that we can easily perform all image generation experiments on CPU.
#
# We have ported the PyTorch SqueezeNet model to TensorFlow; see: `cs231n/classifiers/squeezenet.py` for the model architecture.
#
# To use SqueezeNet, you will need to first **download the weights** by descending into the `cs231n/datasets` directory and running `get_squeezenet_tf.sh`. Note that if you ran `get_assignment3_data.sh` then SqueezeNet will already be downloaded.
#
# Once you've downloaded the Squeezenet model, we can load it into a new TensorFlow session:
#
# [1] Iandola et al, "SqueezeNet: AlexNet-level accuracy with 50x fewer parameters and < 0.5MB model size", arXiv 2016
#
# **NOTE:** Ignore Tensorflow warnings from the cell below
# + tags=["pdf-ignore"]
SAVE_PATH = None
# Local
#SAVE_PATH = 'cs231n/datasets/squeezenet.ckpt'
# Colab
#SAVE_PATH = '/content/drive/My Drive/{}/{}'.format(FOLDERNAME, 'cs231n/datasets/squeezenet.ckpt')
assert SAVE_PATH is not None, "[!] Choose path to squeezenet.ckpt"
if not os.path.exists(SAVE_PATH + ".index"):
raise ValueError("You need to download SqueezeNet!")
model = SqueezeNet()
status = model.load_weights(SAVE_PATH)
model.trainable = False
# -
# ## Load some ImageNet images
# We have provided a few example images from the validation set of the ImageNet ILSVRC 2012 Classification dataset. To download these images, descend into `cs231n/datasets/` and run `get_imagenet_val.sh`.
#
# Since they come from the validation set, our pretrained model did not see these images during training.
#
# Run the following cell to visualize some of these images, along with their ground-truth labels.
# +
from cs231n.data_utils import load_imagenet_val
X_raw, y, class_names = load_imagenet_val(num=5)
plt.figure(figsize=(12, 6))
for i in range(5):
plt.subplot(1, 5, i + 1)
plt.imshow(X_raw[i])
plt.title(class_names[y[i]])
plt.axis('off')
plt.gcf().tight_layout()
# -
# ## Preprocess images
# The input to the pretrained model is expected to be normalized, so we first preprocess the images by subtracting the pixelwise mean and dividing by the pixelwise standard deviation.
X = np.array([preprocess_image(img) for img in X_raw])
# # Saliency Maps
# Using this pretrained model, we will compute class saliency maps as described in Section 3.1 of [2].
#
# A **saliency map** tells us the degree to which each pixel in the image affects the classification score for that image. To compute it, we compute the gradient of the unnormalized score corresponding to the correct class (which is a scalar) with respect to the pixels of the image. If the image has shape `(H, W, 3)` then this gradient will also have shape `(H, W, 3)`; for each pixel in the image, this gradient tells us the amount by which the classification score will change if the pixel changes by a small amount. To compute the saliency map, we take the absolute value of this gradient, then take the maximum value over the 3 input channels; the final saliency map thus has shape `(H, W)` and all entries are nonnegative.
#
# Open the file `cs231n/classifiers/squeezenet.py` and read the code to make sure you understand how to use the model. You will have to use [`tf.GradientTape()`](https://www.tensorflow.org/versions/r2.0/api_docs/python/tf/GradientTape) to compute gradients with respect to the pixels of the image. In particular, it will be very useful to read this [section](https://www.tensorflow.org/alpha/tutorials/eager/automatic_differentiation#gradient_tapes) for better understanding.
#
# [2] <NAME>, <NAME>, and <NAME>. "Deep Inside Convolutional Networks: Visualising
# Image Classification Models and Saliency Maps", ICLR Workshop 2014.
#
# Implement ```compute_saliency_maps``` function inside ```cs231n/net_visualization_tensorflow.py```
# ### Hint: Tensorflow `gather_nd` method
# Recall in Assignment 1 you needed to select one element from each row of a matrix; if `s` is an numpy array of shape `(N, C)` and `y` is a numpy array of shape `(N,`) containing integers `0 <= y[i] < C`, then `s[np.arange(N), y]` is a numpy array of shape `(N,)` which selects one element from each element in `s` using the indices in `y`.
#
# In Tensorflow you can perform the same operation using the `gather_nd()` method. If `s` is a Tensor of shape `(N, C)` and `y` is a Tensor of shape `(N,)` containing longs in the range `0 <= y[i] < C`, then
#
# `tf.gather_nd(s, tf.stack((tf.range(N), y), axis=1))`
#
# will be a Tensor of shape `(N,)` containing one entry from each row of `s`, selected according to the indices in `y`.
#
# You can also read the documentation for the [gather_nd method](https://www.tensorflow.org/versions/r2.0/api_docs/python/tf/gather_nd).
# Once you have completed the implementation above, run the following to visualize some class saliency maps on our example images from the ImageNet validation set:
# + tags=["pdf-ignore-input"]
from cs231n.net_visualization_tensorflow import compute_saliency_maps
def show_saliency_maps(X, y, mask):
mask = np.asarray(mask)
Xm = X[mask]
ym = y[mask]
saliency = compute_saliency_maps(Xm, ym, model)
for i in range(mask.size):
plt.subplot(2, mask.size, i + 1)
plt.imshow(deprocess_image(Xm[i]))
plt.axis('off')
plt.title(class_names[ym[i]])
plt.subplot(2, mask.size, mask.size + i + 1)
plt.title(mask[i])
plt.imshow(saliency[i], cmap=plt.cm.hot)
plt.axis('off')
plt.gcf().set_size_inches(10, 4)
plt.show()
mask = np.arange(5)
show_saliency_maps(X, y, mask)
# + [markdown] tags=["pdf-inline"]
# # INLINE QUESTION
#
# A friend of yours suggests that in order to find an image that maximizes the correct score, we can perform gradient ascent on the input image, but instead of the gradient we can actually use the saliency map in each step to update the image. Is this assertion true? Why or why not?
#
# **Your Answer:**
#
#
# -
# # Fooling Images
# We can also use image gradients to generate "fooling images" as discussed in [3]. Given an image and a target class, we can perform gradient **ascent** over the image to maximize the target class, stopping when the network classifies the image as the target class. Implement the following function to generate fooling images.
#
# [3] Szegedy et al, "Intriguing properties of neural networks", ICLR 2014
#
# Implement ```make_fooling_image``` function inside ```cs231n/net_visualization_tensforflow.py```
#
# Run the following to generate a fooling image. You should ideally see at first glance no major difference between the original and fooling images, and the network should now make an incorrect prediction on the fooling one. However you should see a bit of random noise if you look at the 10x magnified difference between the original and fooling images. Feel free to change the `idx` variable to explore other images.
# + tags=["pdf-ignore-input"]
from cs231n.net_visualization_tensorflow import make_fooling_image
idx = 0
Xi = X[idx][None]
target_y = 6
X_fooling = make_fooling_image(Xi, target_y, model)
# Make sure that X_fooling is classified as y_target
scores = model(X_fooling)
assert tf.math.argmax(scores[0]).numpy() == target_y, 'The network is not fooled!'
# Show original image, fooling image, and difference
orig_img = deprocess_image(Xi[0])
fool_img = deprocess_image(X_fooling[0])
plt.figure(figsize=(12, 6))
# Rescale
plt.subplot(1, 4, 1)
plt.imshow(orig_img)
plt.axis('off')
plt.title(class_names[y[idx]])
plt.subplot(1, 4, 2)
plt.imshow(fool_img)
plt.title(class_names[target_y])
plt.axis('off')
plt.subplot(1, 4, 3)
plt.title('Difference')
plt.imshow(deprocess_image((Xi-X_fooling)[0]))
plt.axis('off')
plt.subplot(1, 4, 4)
plt.title('Magnified difference (10x)')
plt.imshow(deprocess_image(10 * (Xi-X_fooling)[0]))
plt.axis('off')
plt.gcf().tight_layout()
# -
# # Class visualization
# By starting with a random noise image and performing gradient ascent on a target class, we can generate an image that the network will recognize as the target class. This idea was first presented in [2]; [3] extended this idea by suggesting several regularization techniques that can improve the quality of the generated image.
#
# Concretely, let $I$ be an image and let $y$ be a target class. Let $s_y(I)$ be the score that a convolutional network assigns to the image $I$ for class $y$; note that these are raw unnormalized scores, not class probabilities. We wish to generate an image $I^*$ that achieves a high score for the class $y$ by solving the problem
#
# $$
# I^* = {\arg\max}_I (s_y(I) - R(I))
# $$
#
# where $R$ is a (possibly implicit) regularizer (note the sign of $R(I)$ in the argmax: we want to minimize this regularization term). We can solve this optimization problem using gradient ascent, computing gradients with respect to the generated image. We will use (explicit) L2 regularization of the form
#
# $$
# R(I) = \lambda \|I\|_2^2
# $$
#
# **and** implicit regularization as suggested by [3] by periodically blurring the generated image. We can solve this problem using gradient ascent on the generated image.
#
# [2] <NAME>, <NAME>, and <NAME>. "Deep Inside Convolutional Networks: Visualising
# Image Classification Models and Saliency Maps", ICLR Workshop 2014.
#
# [3] Yosinski et al, "Understanding Neural Networks Through Deep Visualization", ICML 2015 Deep Learning Workshop
#
#
# In `cs231n/net_visualization_tensorflow.py` complete the implementation of the `image_visualization_update_step` used in the `create_class_visualization` function below.
# Once you have completed that implementation, run the following cells to generate an image of a Tarantula:
# +
from cs231n.net_visualization_tensorflow import class_visualization_update_step, jitter, blur_image
def create_class_visualization(target_y, model, **kwargs):
"""
Generate an image to maximize the score of target_y under a pretrained model.
Inputs:
- target_y: Integer in the range [0, 1000) giving the index of the class
- model: A pretrained CNN that will be used to generate the image
Keyword arguments:
- l2_reg: Strength of L2 regularization on the image
- learning_rate: How big of a step to take
- num_iterations: How many iterations to use
- blur_every: How often to blur the image as an implicit regularizer
- max_jitter: How much to jitter the image as an implicit regularizer
- show_every: How often to show the intermediate result
"""
l2_reg = kwargs.pop('l2_reg', 1e-3)
learning_rate = kwargs.pop('learning_rate', 25)
num_iterations = kwargs.pop('num_iterations', 100)
blur_every = kwargs.pop('blur_every', 10)
max_jitter = kwargs.pop('max_jitter', 16)
show_every = kwargs.pop('show_every', 25)
# We use a single image of random noise as a starting point
X = 255 * np.random.rand(224, 224, 3)
X = preprocess_image(X)[None]
loss = None # scalar loss
grad = None # gradient of loss with respect to model.image, same size as model.image
X = tf.Variable(X)
for t in range(num_iterations):
# Randomly jitter the image a bit; this gives slightly nicer results
ox, oy = np.random.randint(0, max_jitter, 2)
X = jitter(X, ox, oy)
X = class_visualization_update_step(X, model, target_y, l2_reg, learning_rate)
# Undo the jitter
X = jitter(X, -ox, -oy)
# As a regularizer, clip and periodically blur
X = tf.clip_by_value(X, -SQUEEZENET_MEAN/SQUEEZENET_STD, (1.0 - SQUEEZENET_MEAN)/SQUEEZENET_STD)
if t % blur_every == 0:
X = blur_image(X, sigma=0.5)
# Periodically show the image
if t == 0 or (t + 1) % show_every == 0 or t == num_iterations - 1:
plt.imshow(deprocess_image(X[0]))
class_name = class_names[target_y]
plt.title('%s\nIteration %d / %d' % (class_name, t + 1, num_iterations))
plt.gcf().set_size_inches(4, 4)
plt.axis('off')
plt.show()
return X
# -
# Once you have completed the implementation in the cell above, run the following cell to generate an image of Tarantula:
target_y = 76 # Tarantula
out = create_class_visualization(target_y, model)
# Try out your class visualization on other classes! You should also feel free to play with various hyperparameters to try and improve the quality of the generated image, but this is not required.
target_y = np.random.randint(1000)
# target_y = 78 # Tick
# target_y = 187 # Yorkshire Terrier
# target_y = 683 # Oboe
# target_y = 366 # Gorilla
# target_y = 604 # Hourglass
print(class_names[target_y])
X = create_class_visualization(target_y, model)
| assignment3/NetworkVisualization-TensorFlow.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import os
#Read data
raw_data_path =os.path.join(os.path.pardir,os.path.pardir,'data','raw', 'Dataset10')
train_file_path = os.path.join(raw_data_path,'data.csv')
df = pd.read_csv(train_file_path)
df =df.drop(['Names','Company','Location','Onboard_date'], axis=1)
#Write data
proccessed_data_path =os.path.join(os.path.pardir,os.path.pardir,'data','processed')
write_train_path = os.path.join(proccessed_data_path,'dataset10.csv')
df.to_csv(write_train_path)
# -
df.Churn.value_counts().plot(kind= 'pie', title='Churned proportion of customers', explode = [0,0.1],autopct='%1.1f%%', shadow=True);
| project/notebooks/Dataset10/Dataset10 - EDA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Censored Data Models
# +
from copy import copy
import arviz as az
import matplotlib.pyplot as plt
import numpy as np
import pymc3 as pm
import seaborn as sns
from numpy.random import default_rng
# -
# %config InlineBackend.figure_format = 'retina'
rng = default_rng(1234)
az.style.use("arviz-darkgrid")
# [This example notebook on Bayesian survival
# analysis](http://docs.pymc.io/notebooks/survival_analysis.html) touches on the
# point of censored data. _Censoring_ is a form of missing-data problem, in which
# observations greater than a certain threshold are clipped down to that
# threshold, or observations less than a certain threshold are clipped up to that
# threshold, or both. These are called right, left and interval censoring,
# respectively. In this example notebook we consider interval censoring.
#
# Censored data arises in many modelling problems. Two common examples are:
#
# 1. _Survival analysis:_ when studying the effect of a certain medical treatment
# on survival times, it is impossible to prolong the study until all subjects
# have died. At the end of the study, the only data collected for many patients
# is that they were still alive for a time period $T$ after the treatment was
# administered: in reality, their true survival times are greater than $T$.
#
# 2. _Sensor saturation:_ a sensor might have a limited range and the upper and
# lower limits would simply be the highest and lowest values a sensor can
# report. For instance, many mercury thermometers only report a very narrow
# range of temperatures.
#
# This example notebook presents two different ways of dealing with censored data
# in PyMC3:
#
# 1. An imputed censored model, which represents censored data as parameters and
# makes up plausible values for all censored values. As a result of this
# imputation, this model is capable of generating plausible sets of made-up
# values that would have been censored. Each censored element introduces a
# random variable.
#
# 2. An unimputed censored model, where the censored data are integrated out and
# accounted for only through the log-likelihood. This method deals more
# adequately with large amounts of censored data and converges more quickly.
#
# To establish a baseline we compare to an uncensored model of the uncensored
# data.
# +
# Produce normally distributed samples
size = 500
true_mu = 13.0
true_sigma = 5.0
samples = rng.normal(true_mu, true_sigma, size)
# Set censoring limits
low = 3.0
high = 16.0
def censor(x, low, high):
x = copy(x)
x[x <= low] = low
x[x >= high] = high
return x
# Censor samples
censored = censor(samples, low, high)
# -
# Visualize uncensored and censored data
_, ax = plt.subplots(figsize=(10, 3))
edges = np.linspace(-5, 35, 30)
ax.hist(samples, bins=edges, density=True, histtype="stepfilled", alpha=0.2, label="Uncensored")
ax.hist(censored, bins=edges, density=True, histtype="stepfilled", alpha=0.2, label="Censored")
[ax.axvline(x=x, c="k", ls="--") for x in [low, high]]
ax.legend();
# ## Uncensored Model
def uncensored_model(data):
with pm.Model() as model:
mu = pm.Normal("mu", mu=((high - low) / 2) + low, sigma=(high - low))
sigma = pm.HalfNormal("sigma", sigma=(high - low) / 2.0)
observed = pm.Normal("observed", mu=mu, sigma=sigma, observed=data)
return model
# We should predict that running the uncensored model on uncensored data, we will get reasonable estimates of the mean and variance.
uncensored_model_1 = uncensored_model(samples)
with uncensored_model_1:
trace = pm.sample(tune=1000, return_inferencedata=True)
az.plot_posterior(trace, ref_val=[true_mu, true_sigma], round_to=3);
# And that is exactly what we find.
#
# The problem however, is that in censored data contexts, we do not have access to the true values. If we were to use the same uncensored model on the censored data, we would anticipate that our parameter estimates will be biased. If we calculate point estimates for the mean and std, then we can see that we are likely to underestimate the mean and std for this particular dataset and censor bounds.
np.mean(censored), np.std(censored)
uncensored_model_2 = uncensored_model(censored)
with uncensored_model_2:
trace = pm.sample(tune=1000, return_inferencedata=True)
az.plot_posterior(trace, ref_val=[true_mu, true_sigma], round_to=3);
# The figure above confirms this.
#
# ## Censored data models
#
# The models below show 2 approaches to dealing with censored data. First, we need to do a bit of data pre-processing to count the number of observations that are left or right censored. We also also need to extract just the non-censored data that we observe.
n_right_censored = sum(censored >= high)
n_left_censored = sum(censored <= low)
n_observed = len(censored) - n_right_censored - n_left_censored
uncensored = censored[(censored > low) & (censored < high)]
assert len(uncensored) == n_observed
# ### Model 1 - Imputed Censored Model of Censored Data
#
# In this model, we impute the censored values from the same distribution as the uncensored data. Sampling from the posterior generates possible uncensored data sets.
#
# This model makes use of [PyMC3's bounded variables](https://docs.pymc.io/api/bounds.html).
with pm.Model() as imputed_censored_model:
mu = pm.Normal("mu", mu=((high - low) / 2) + low, sigma=(high - low))
sigma = pm.HalfNormal("sigma", sigma=(high - low) / 2.0)
right_censored = pm.Bound(pm.Normal, lower=high)(
"right_censored", mu=mu, sigma=sigma, shape=n_right_censored
)
left_censored = pm.Bound(pm.Normal, upper=low)(
"left_censored", mu=mu, sigma=sigma, shape=n_left_censored
)
observed = pm.Normal("observed", mu=mu, sigma=sigma, observed=uncensored, shape=n_observed)
with imputed_censored_model:
trace = pm.sample(return_inferencedata=True)
az.plot_posterior(trace, var_names=["mu", "sigma"], ref_val=[true_mu, true_sigma], round_to=3);
# We can see that the bias in the estimates of the mean and variance (present in the uncensored model) have been largely removed.
# ### Model 2 - Unimputed Censored Model of Censored Data
#
# In this model, we do not impute censored data, but instead integrate them out through the likelihood.
#
# The implementations of the likelihoods are non-trivial. See the [Stan manual](https://github.com/stan-dev/stan/releases/download/v2.14.0/stan-reference-2.14.0.pdf) (section 11.3 on censored data) and the [original PyMC3 issue on GitHub](https://github.com/pymc-devs/pymc3/issues/1833) for more information.
#
# This model makes use of [PyMC3's `Potential`](https://docs.pymc.io/api/model.html#pymc3.model.Potential).
# +
# Import the log cdf and log complementary cdf of the normal Distribution from PyMC3
from pymc3.distributions.dist_math import normal_lccdf, normal_lcdf
# Helper functions for unimputed censored model
def left_censored_likelihood(mu, sigma, n_left_censored, lower_bound):
""" Likelihood of left-censored data. """
return n_left_censored * normal_lcdf(mu, sigma, lower_bound)
def right_censored_likelihood(mu, sigma, n_right_censored, upper_bound):
""" Likelihood of right-censored data. """
return n_right_censored * normal_lccdf(mu, sigma, upper_bound)
# -
with pm.Model() as unimputed_censored_model:
mu = pm.Normal("mu", mu=0.0, sigma=(high - low) / 2.0)
sigma = pm.HalfNormal("sigma", sigma=(high - low) / 2.0)
observed = pm.Normal(
"observed",
mu=mu,
sigma=sigma,
observed=uncensored,
)
left_censored = pm.Potential(
"left_censored", left_censored_likelihood(mu, sigma, n_left_censored, low)
)
right_censored = pm.Potential(
"right_censored", right_censored_likelihood(mu, sigma, n_right_censored, high)
)
# Sampling
with unimputed_censored_model:
trace = pm.sample(tune=1000, return_inferencedata=True)
az.plot_posterior(trace, var_names=["mu", "sigma"], ref_val=[true_mu, true_sigma], round_to=3);
# Again, the bias in the estimates of the mean and variance (present in the uncensored model) have been largely removed.
# ## Discussion
#
# As we can see, both censored models appear to capture the mean and variance of the underlying distribution as well as the uncensored model! In addition, the imputed censored model is capable of generating data sets of censored values (sample from the posteriors of `left_censored` and `right_censored` to generate them), while the unimputed censored model scales much better with more censored data, and converges faster.
# ## Authors
#
# - Originally authored by [<NAME>](https://github.com/domenzain) on Mar 7, 2017.
# - Updated by [<NAME>](https://github.com/eigenfoo) on Jul 14, 2018.
# - Updated by [<NAME>](https://github.com/drbenvincent) in May 2021.
# %load_ext watermark
# %watermark -n -u -v -iv -w -p theano,xarray
| examples/survival_analysis/censored_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # MOSFiT Module Analysis
# Using 1 component rprocess model as input, specific modules are computed. We want to know what each module does and what each component does.
#
# r-process Model Modules:
# - parameter : ebv
# - parameter : nhhost
# - parameter : texplosion
# - parameter : temperature
# - parameter : kappa
# - parameter : kappagamma
# - parameter : frad
# - parameter : mejecta
# - parameter : vejecta
# - module : engine : rprocess
# - module : transform : diffusion
# - module : photosphere : temperature_floor
# - module : sed : blackbody
# - module : sed : losextinction
# - module : observable : photometry
#
# There are other modules that need to be called in order to compute the modules in the model (listed above). Namely, `resttimes` and `densetimes` need to be called before everything else.
#
# We choose to look at the parameters of mejecta and vejecta and see how the physics changes, but this should be easy enough to modify to analyze any other parameter (e.g. if you were interested in kappa, make an array of kappas and iterate over kappa in plotting scripts).
#
# Without further ado...
#
# ## Instantiate + Import
import numpy as np
import mosfit
import matplotlib.pyplot as plt
import matplotlib as mpl
import json
from collections import OrderedDict
# %matplotlib inline
# Analyzing 1-Component Kilonova Model
my_model = mosfit.model.Model(model='rprocess')
# +
# Parameters
kappa = 1 # opacity
kappa_gamma = 1000 # opacity for high-energy photons
#red, middle, blue from Villar et al. 2017
frad = 0.999 # Fraction radioactive material
mejectas = np.array([.01, .015, .02])
vejectas = np.array([.137,.201,.266])*(mosfit.constants.C_CGS /mosfit.constants.KM_CGS)
# Values for GW170817
# These are calculated in ../notMOSFiT/mosfit/
redshift = 0.009727
lumdist = 39.5
# -
# ### ....an interlude for an explanation of the observables parameters
# The way the observables parameters are organizes is as one big table:
#
# time | band | telescope | instrument | mode | measure | system | freq | u_freq | zeropt | bandset
# --- |--- |--- |--- |--- |--- |--- |--- |--- |--- |--- |--- |
# 0.0 | u | Blanco | DECam | - | - | DECam | - | - | - | DECam
# 0.0 | g | Blanco | DECam | - | - | DECam | - | - | - | DECam
# 0.1 | u | Blanco | DECam | - | - | DECam | - | - | - | DECam
# 0.1 | g | Blanco | DECam | - | - | DECam | - | - | - | DECam
# etc. | - | - | - | - | - | - | - | - | - |
#
# And the way MOSFiT stores this data is each column is an array. So we're gonna fake it and generate these arrays for each time in np.linspace(0,10,101) and for ugrizY.
#
# Note: I actually couldn'y get this to work right unless I used only one band. So the above explanation could be straight up wrong.
# +
# Actual Setting of Values
texplosion = 0
times = np.linspace(0,10,101)
bands = ['u']
# Creating MOSFiT-happy structures
all_bands = [j for i in times for j in bands]
all_times = [i for i in times for j in bands]
modes = ['' for i in times for j in bands]
measures = ['' for i in times for j in bands]
systems = ['DECam' for i in times for j in bands]
telescopes = ['Blanco' for i in times for j in bands]
instruments = ['DECam' for i in times for j in bands]
frequencies = [None for i in times for j in bands]
u_frequencies = [None for i in times for j in bands]
zeropoints = [None for i in times for j in bands]
bandsets = ['' for i in times for j in bands]
obs_types = ['magnitude' for i in times for j in bands]
band_indices = [32 for i in times for j in bands ] # Set the only band to be DECam 'u'
sample_wavs = [np.linspace(300,1000, 100) for i in range(0,1415)]
# -
# ## AllTimes
# [alltimes.py](./mosfit/modules/arrays/alltimes.py)
#
# AllTimes is supposed to calculate a list of times associated with real observations, but in reality, I need it to calculate band indices correctly.
time_converter = mosfit.modules.AllTimes(name='time_convert',
model=my_model)
# ## Obvservables : Photometry
# [photometry.py](./mosfit/modules/observables/photometry.py)
#
# The photometry class calculates observables from the luminosity, given the bands. It also does some internal book-keeping to keep track of the telescope bands as indices.
#
#
#
# We need to import this class because it is called by time_converter. Then, we can run time converter an get the main data structure we want: band_indices
# +
my_photometry = mosfit.modules.Photometry(name='photometry', model=my_model,
bands=bands, telescope='Blanco',
instrument='DECam',mode='', bandset='DECam')
time_converter._photometry = my_photometry
times = time_converter.process(times=all_times,
bands=all_bands,
telescopes=telescopes,
modes=modes,
measures=measures,
systems=systems,
frequencies=frequencies,
u_frequencies=u_frequencies,
zeropoints=zeropoints,
bandsets=bandsets,
instruments=instruments)
my_photometry.load_bands(times['all_band_indices'])
# -
# ## Rest Times
# [resttimes.py](./mosfit/modules/arrays/resttimes.py)
#
# Calculates the times in the rest frame of the event with:
#
# $$ t_{rest} = \frac{t}{1+z} $$
#
# Inputs:
#
# Key | Explanation | Units
# --- | --- | ---
# all_times | Times of observation | list of floats (MDJs)
# texplosion | Time of explosion | float (MJD)
# redshift | Redshift of event | float
#
# Outputs:
#
# Key | Explanation | Units
# --- | --- | ---
# rest_times | Times in rest frame of the event | list of floats (MDJs)
# resttexplosion | Time of explosion in rest frame | float (MJD)
#
rest_time_converter = mosfit.modules.RestTimes(name='resttimes', model=my_model)
rest_times = rest_time_converter.process(all_times=times['all_times'],
redshift=redshift,
texplosion=texplosion)
# ## Dense Times
# [densetimes.py](./mosfit/modules/arrays/densetimes.py)
#
# Generates an array of evenly-spaced times for use in calculations.
#
# Inputs:
#
# Key | Explanation | Units
# --- | --- | ---
# rest_times | Times in rest frame of the event | array of floats (MDJs)
# resttexplosion | Time of explosion in rest frame | float (MJD)
#
#
# Key | Explanation | Units
# --- | --- | ---
# dense_times | Times in rest frame of the event for which calculations will be done| array of floats (MDJs)
# dense_indices | Actually I don't know what this is | array of ints
#
#
dense_converter = mosfit.modules.DenseTimes(name='densetimes', model=my_model)
dense_times = dense_converter.process(rest_times=rest_times['rest_times'],
resttexplosion=rest_times['resttexplosion'])
# ## Engine: r-process
# [rprocess.py](./mosfit/modules/engines/rprocess.py)
#
# This script calculates the luminosity based on the equation for heating rate in
# [Metzger 2017](https://ui.adsabs.harvard.edu/#abs/2017LRR....20....3M/abstract), Eqn. 22.
#
# $$ e_r = 4 x 10^{18} \epsilon_{th,v} ( 0.5 - \pi^{-1} arctan[(t-t_0)/\sigma])^{1.3} erg \ s^{-1} g^{-1} $$
#
# where $ \epsilon_{th,v}$ is
#
# $$ \epsilon_{th,v}(t) = 0.36 \Bigg[ \exp (-a_vt_{day}) + \frac{\ln\big(1 + 2b_vt^{d_v}_{day}\big)}{2b_vt^{d_v}_{day}} \Bigg] $$
#
# Inputs:
#
# Key | Explanation | Units
# --- | --- | ---
# dense_times | Array of times to evaluate | MJDs
# mejecta | Mass of ejected material | $M_{\odot}$
# resttexplosion | Time of explosion at rest | MJD
# vejecta | Velocity of ejected mass | km/s
#
#
# Outputs:
# A dictionary with the sole key being luminosities.
# +
my_engine = mosfit.modules.engines.rprocess.RProcess(name='my_engine', model=my_model)
fig, ax = plt.subplots(3, 3, sharex='col', sharey='row', figsize=(12,12))
for i,k in zip(mejectas, range(3)):
for j, n in zip(vejectas, range(3)):
luminosities = my_engine.process( dense_times=dense_times['dense_times'],
mejecta=i,
resttexplosion=rest_times['resttexplosion'],
vejecta=j )
ax[k,n].plot(dense_times['dense_times'], luminosities['luminosities'])
ax[k,n].set_title('M = ' + str(i) + ', v = ' + str(j/ (mosfit.constants.C_CGS /mosfit.constants.KM_CGS)))
ax[k,n].set_ylim(-.01e50, 1.5e50)
ax[k,n].set_xlim(-.01, .01)
fig.text(0.5, .08, 'Time (MJDs)', ha='center')
fig.text(0.08, 0.5, 'Luminosity (erg/s)', va='center', rotation='vertical')
# -
# ## Transform: Diffusion
# [diffusion.py](./mosfit/modules/transforms/diffusion.py)
#
# Given luminosities from the engine, it modifies them to model diffusion of photons through matter, as described in [Arnett 1982](https://ui.adsabs.harvard.edu/#abs/1982ApJ...253..785A/abstract). Specifically, it returns a time scale (Eqn. 18):
#
# $$ \tau_m^2 = 2\tau_0\tau_h $$
#
#
# or, as given in [Nicholl et al. 2017](https://ui.adsabs.harvard.edu/#abs/2017ApJ...850...55N/abstract), Eqn. 6,
#
# $$ t_{diff} = \Big( \frac{2\kappa M_{ej}}{\beta c \nu_{ej}} \Big) $$
#
# and the leakage paramater (or trap_coeff), which controls the fraction of the magnetar input energy that is thermalized in the ejecta
#
# $$ A = \frac{3\kappa_{\gamma}M_{ej}}{4\pi \nu_{ej}^2} $$
#
# Combining these, the output luminosity is given as Eqn.5:
#
# $$ L_{out}(t) = \exp(t/t_{diff})^2 (1 - e^{-At^{-2}}) \int_{0}^{t} 2 F_{in}(t') \frac{t'}{t_{diff}} \exp(t' / t_{diff})^2 \frac{dt'}{t_{diff}} $$
#
# Inputs:
#
# Key | Explanation | Units
# --- | --- | ---
# dense_times | Array of times to evaluate | MJDs
# rest_times | Array of times to evaluate (in rest frame) - use same array | MJDs
# times | rest_times (see transform.py) | MJDs
# mejecta | Mass of ejected material | $M_{\odot}$
# resttexplosion | Time of explosion at rest | MJD
# vejecta | Velocity of ejected mass | km/s
# kappa | Opacity of ejected material to optical | $cm^2 g^{-1}$
# kappa_gamma | Opacity of ejected material to gamma rays | $cm^2 g^{-1}$
#
#
# Outputs:
#
# Key | Explanation
# --- | ---
# tau_diffusion | Characteristic time
# luminosities | Luminosity after diffusion
# +
my_transform = mosfit.modules.transforms.diffusion.Diffusion(name='diffusion', model=my_model)
fig, ax = plt.subplots(3, 3, sharex='col', sharey='row', figsize=(12,12))
for i,k in zip(mejectas, range(3)):
for j, n in zip(vejectas, range(3)):
luminosities = my_engine.process( dense_times=dense_times['dense_times'],
mejecta=i,
resttexplosion=rest_times['resttexplosion'],
vejecta=j )
new_lums = my_transform.process(dense_times=dense_times['dense_times'],
mejecta=i,
resttexplosion=rest_times['resttexplosion'],
dense_luminosities=luminosities['luminosities'],
kappa=kappa, kappagamma=kappa_gamma,
rest_times=rest_times['rest_times'], vejecta=j,
times=rest_times['rest_times'])
ax[k,n].plot(rest_times['rest_times'], new_lums['luminosities'], )
ax[k,n].set_title('M = ' + str(i) + ', v = ' + str(j/ (mosfit.constants.C_CGS /mosfit.constants.KM_CGS)))
# ax[k,n].set_ylim(-.01e37, 1.e37)
fig.text(0.5, .08, 'Time (MJDs)', ha='center')
fig.text(0.08, 0.5, 'Luminosity (erg/s)', va='center', rotation='vertical')
# -
# ## Photosphere: Temperature Floor
# [temperature_floor.py](./mosfit/modules/photospheres/temperature_floor.py)
#
# This module returns the temepearture and radius of the photosphere as a function of time. The temeprature is from the Stefan-Boltzmann law, "until the ejecta cools to some critical tempearture." The radius is calculated as $R = vt$, where $v$ is $v_{phot} \approx v_{ejecta}$. This method is outlined in [Nicholl et al 2017](https://ui.adsabs.harvard.edu/#abs/2017ApJ...850...55N/abstract), section 3.6.
#
# Inputs:
#
# Key | Explanation | Units
# --- | --- | ---
# luminosities | Luminosity as a function of time | $ erg \ s^{-1}$
# rest_times | Array of times to evaluate (in rest frame) - use same array | MJDs
# temperature | A final temperature | K
# times | Array of times to evaluate - use same array | MJDs
# mejecta | Mass of ejected material | $M_{\odot}$
# resttexplosion | Time of explosion at rest | MJD
# vejecta | Velocity of ejected mass | km/s
# kappa | Opacity of ejected material to optical | $cm^2 g^{-1}$
#
# Outputs:
#
# Key | Explanation | Units
# --- | --- | ---
# radiusphot | Time-dependent radius of the photosphere | km
# temperaturephot | Time-dependent tempearature of the photosphere | K
#
# ### Below, I set temperature floor to 0.
# +
my_photosphere = mosfit.modules.photospheres.temperature_floor.TemperatureFloor(name='tempfloor', model=my_model)
fig1, ax1 = plt.subplots(3, 3, sharex='col', sharey='row', figsize=(12,12))
fig2, ax2 = plt.subplots(3, 3, sharex='col', sharey='row', figsize=(12,12))
for i,k in zip(mejectas, range(3)):
for j, n in zip(vejectas, range(3)):
luminosities = my_engine.process( dense_times=dense_times['dense_times'],
mejecta=i,
resttexplosion=rest_times['resttexplosion'],
vejecta=j )
new_lums = my_transform.process(dense_times=dense_times['dense_times'],
mejecta=i,
resttexplosion=rest_times['resttexplosion'],
dense_luminosities=luminosities['luminosities'],
kappa=kappa, kappagamma=kappa_gamma,
rest_times=rest_times['rest_times'], vejecta=j,
times=rest_times['rest_times'])
photosphere_output = my_photosphere.process(
mejecta=i,
resttexplosion=rest_times['resttexplosion'],
luminosities=new_lums['luminosities'],
temperature=0, kappa=kappa,
vejecta=j,
rest_times=rest_times['rest_times'])
ax1[k,n].plot(rest_times['rest_times'], photosphere_output['radiusphot'])
ax[k,n].set_title('M = ' + str(i) + ', v = ' + str(j/ (mosfit.constants.C_CGS /mosfit.constants.KM_CGS)))
ax1[k,n].set_ylim(-.1e15, 7e15)
ax2[k,n].plot(rest_times['rest_times'], photosphere_output['temperaturephot'])
ax[k,n].set_title('M = ' + str(i) + ', v = ' + str(j/ (mosfit.constants.C_CGS /mosfit.constants.KM_CGS)))
ax2[k,n].set_ylim(-400, 16000)
fig1.text(0.5, .08, 'Time (MJDs)', ha='center')
fig1.text(0.08, 0.5, 'Radius Photosphere (km)', va='center', rotation='vertical')
fig2.text(0.5, .08, 'Time (MJDs)', ha='center')
fig2.text(0.03, 0.5, 'Temperature Photosphere (K)', va='center', rotation='vertical')
# -
# ## SED: Blackbody
# [blackbody.py](./mosfit/modules/seds/blackbody.py)
#
# The blackbody module returns the spectrum of the object as a function of time. The blackbody class inherits from the SED class. The spectra is calculated from the Planck formula. More information can be found in [Nicholl et al 2017](https://ui.adsabs.harvard.edu/#abs/2017ApJ...850...55N/abstract), section 3.7.
#
# Inputs:
#
# Key | Explanation | Units
# --- | --- | ---
# luminosities | Luminosity as a function of time | $$ erg \ s^{-1} $$
# all_bands | Bands to return observations in | list of strings
# all_band_indices | An internal band-matched index corresponding to all bands | list of ints
# all_frequencies | For count-based observations (ignore) | -
# radiusphot | Time-dependent radius of the photosphere | km?
# redshift | Redshift of event | z
# sample_wavelengths | Wavelengths for which to evaluate SED | nm
#
# Outputs:
#
# Key | Explanation | Units
# --- | --- | ---
# sample_wavelengths | Wavelengths of the SED | nm
# seds | A list of np.arrays corresponding to an SED for each time that luminosities was evaluated in | erg, probably
# Cheating gently to get the output of the SED
my_blackbody = mosfit.modules.seds.blackbody.Blackbody(name='bb', model=my_model)
my_blackbody._sample_wavelengths = sample_wavs
# +
fig, ax = plt.subplots(3, 3, sharex='col', sharey='row', figsize=(14,12))
c = rest_times['rest_times'][0:20]
norm = mpl.colors.Normalize(vmin=c.min(), vmax=c.max())
cmap = mpl.cm.ScalarMappable(norm=norm, cmap=mpl.cm.jet)
cmap.set_array([])
for i,k in zip(mejectas, range(3)):
for j, n in zip(vejectas, range(3)):
luminosities = my_engine.process( dense_times=dense_times['dense_times'],
mejecta=i,
resttexplosion=rest_times['resttexplosion'],
vejecta=j )
new_lums = my_transform.process(dense_times=dense_times['dense_times'],
mejecta=i,
resttexplosion=rest_times['resttexplosion'],
dense_luminosities=luminosities['luminosities'],
kappa=kappa, kappagamma=kappa_gamma,
rest_times=rest_times['rest_times'], vejecta=j,
times=rest_times['rest_times'])
photosphere_output = my_photosphere.process(
mejecta=i,
resttexplosion=rest_times['resttexplosion'],
luminosities=new_lums['luminosities'],
temperature=0, kappa=kappa,
vejecta=j,
rest_times=rest_times['rest_times'])
bb_output = my_blackbody.process(luminosities=new_lums['luminosities'],
all_bands = times['all_bands'],
all_band_indices=band_indices,
all_frequencies=frequencies,
radiusphot=photosphere_output['radiusphot'],
temperaturephot=photosphere_output['temperaturephot'],
redshift=redshift)
for m, color in enumerate(c):
ax[k,n].plot(bb_output['sample_wavelengths'][m], bb_output['seds'][m], c=cmap.to_rgba(color))
ax[k,n].set_title('M = ' + str(i) + ', v = ' + str(j/ (mosfit.constants.C_CGS /mosfit.constants.KM_CGS)))
cbar_ax = fig.add_axes([.93, 0.15, 0.03, 0.7])
fig.colorbar(cmap, ticks=c, cax=cbar_ax)
fig.text(0.5, .08, 'Wavelength (nm)', ha='center')
fig.text(0.08, 0.5, 'Energy (erg)', va='center', rotation='vertical')
# -
# This is the point at which I would attempt to try to plot actual observables, but to be perfectly honest? It works when you run mosfit from the command line, and that's good enough for me.
| module_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="jk5SHfqIc6Ey"
# ### **INITIALIZATION:**
# - I use these three lines of code on top of my each notebooks because it will help to prevent any problems while reloading the same project. And the third line of code helps to make visualization within the notebook.
# + id="z9K2einKadKr"
#@ INITIALIZATION:
# %reload_ext autoreload
# %autoreload 2
# %matplotlib inline
# + [markdown] id="RTFZaPENdHTs"
# **LIBRARIES AND DEPENDENCIES:**
# - I have downloaded all the libraries and dependencies required for the project in one particular cell.
# + id="GZDkvIrydDtY"
#@ INSTALLING DEPENDENCIES: UNCOMMENT BELOW:
# # !pip install -Uqq fastbook
# import fastbook
# fastbook.setup_book()
# + id="qTIdUUtEdOah"
#@ DOWNLOADING LIBRARIES AND DEPENDENCIES:
from fastbook import * # Getting all the Libraries.
from fastai.callback.fp16 import *
from fastai.text.all import * # Getting all the Libraries.
from IPython.display import display, HTML
# + [markdown] id="PAY-kcjCd-Tz"
# ### **GETTING THE DATASET:**
# - I will get the **IMDB Dataset** here.
# + colab={"base_uri": "https://localhost:8080/", "height": 54} id="iv-s-9D1dsia" outputId="e45ff349-4494-4c2d-a21a-6184d08452b7"
#@ GETTING THE DATASET:
path = untar_data(URLs.IMDB) # Getting Path to the Dataset.
path.ls() # Inspecting the Path.
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="dBavrB_6e_Of" outputId="87fe6ba0-d4e2-4921-8f5c-09d3aa9d3877"
#@ GETTING TEXT FILES:
files = get_text_files(path, folders=["train", "test", "unsup"]) # Getting Text Files.
txt = files[0].open().read() # Getting a Text.
txt[:75] # Inspecting Text.
# + [markdown] id="kG832kuRgXeo"
# ### **WORD TOKENIZATION:**
# - **Word Tokenization** splits a sentence on spaces as well as applying language specific rules to try to separate parts of meaning even when there are no spaces. Generally punctuation marks are also split into separate tokens. **Token** is a element of a list created by the **Tokenization** process which could be a word, a part of a word or subword or a single character.
# + colab={"base_uri": "https://localhost:8080/"} id="yx1lsBcYgQ9K" outputId="a97b8cdd-70b7-473b-e322-aefacf89acfc"
#@ INITIALIZING WORD TOKENIZATION:
spacy = WordTokenizer() # Initializing Tokenizer.
toks = first(spacy([txt])) # Getting Tokens of Words.
print(coll_repr(toks, 30)) # Inspecting Tokens.
# + colab={"base_uri": "https://localhost:8080/"} id="ti9Hi_uriKxe" outputId="3ab387c8-7c32-4966-bed7-fd8a701d8f3e"
#@ INSPECTING TOKENIZATION: EXAMPLE:
first(spacy(['The U.S. dollar $1 is $1.00.'])) # Inspecting Tokens.
# + colab={"base_uri": "https://localhost:8080/"} id="liWRboFJi84p" outputId="dee03077-aa0c-4f98-9f81-44bf1904ebee"
#@ INITIALIZING WORD TOKENIZATION WITH FASTAI:
tkn = Tokenizer(spacy) # Initializing Tokenizer.
print(coll_repr(tkn(txt), 31)) # Inspecting Tokens.
# + [markdown] id="lsVt5tA8P1hU"
# **Note:**
# - **xxbos** : Indicates the beginning of a text.
# - **xxmaj** : Indicates the next word begins with a capital.
# - **xxunk** : Indicates the next word is unknown.
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="X-GgI67Ajr-K" outputId="d74da75e-bd5e-4308-c48f-4e2353f7c647"
#@ INSPECTING TOKENIZATION: EXAMPLE:
coll_repr(tkn('© Fast.ai www.fast.ai/INDEX'), 30) # Inspecting Tokens.
# + [markdown] id="Z8clYuvALmBg"
# ### **SUBWORD TOKENIZATION:**
# - **Word Tokenization** relies on an assumption that spaces provide a useful separation of components of meaning in a sentence which is not always appropriate. Languages such as Chinese and Japanese don't use spaces and in such cases **Subword Tokenization** generally plays the best role. **Subword Tokenization** splits words into smaller parts based on the most commonly occurring sub strings.
# + id="AO-mr3aLm2QB" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="3bc61f99-1e67-4d46-a21e-cbc7c46f0933"
#@ INITIALIZING SUBWORD TOKENIZATION: EXAMPLE:
txts = L(o.open().read() for o in files[:2000]) # Getting List of Reviews.
#@ INITIALIZING SUBWORD TOKENIZER:
def subword(sz): # Defining Function.
sp = SubwordTokenizer(vocab_sz=sz) # Initializing Subword Tokenizer.
sp.setup(txts) # Getting Sequence of Characters.
return " ".join(first(sp([txt]))[:40]) # Inspecting the Vocab.
#@ IMPLEMENTATION:
subword(1000) # Inspecting Subword Tokenization.
# + [markdown] id="KWSi_eQzQcxL"
# **Notes:**
# - Here **setup** is a special fastai method that is called automatically in usual data processing pipelines which reads the documents and find the common sequences of characters to create the vocab. Similarly [**L**](https://fastcore.fast.ai/#L) is also referred as superpowered list. The special character '_' represents a space character in the original text.
# + colab={"base_uri": "https://localhost:8080/", "height": 52} id="MpSPYDjIQbVP" outputId="75fbfe62-3b0e-48a7-d503-67e2a52b6c82"
#@ IMPLEMENTATION OF SUBWORD TOKENIZATION:
subword(200) # Inspecting Vocab.
subword(10000) # Inspecting Vocab.
# + [markdown] id="xTxPsa_RT-oU"
# **Note:**
# - A larger vocab means fewer tokens per sentence which means faster training, less memory, and less state for the model to remember but it means larger embedding matrices and require more data to learn. **Subword Tokenization** provides a way to easily scale between character tokenization i.e. using a small subword vocab and word tokenization i.e using a large subword vocab and handles every human language without needing language specific algorithms to be developed.
# + [markdown] id="9scF0s-DVQjf"
# ### **NUMERICALIZATION:**
# - **Numericalization** is the process of mapping tokens to integers. It involves making a list of all possible levels of that categorical variable or the vocab and replacing each level with its index in the vocab.
# + colab={"base_uri": "https://localhost:8080/"} id="vV8wXsyFTb42" outputId="256b3df0-a2aa-4826-ba1a-74454dfd901d"
#@ INITIALIZING TOKENS:
toks = tkn(txt) # Getting Tokens.
print(coll_repr(tkn(txt), 31)) # Inspecting Tokens.
# + colab={"base_uri": "https://localhost:8080/"} id="5GORafHQXscv" outputId="47f5ece8-fad8-4402-8181-a6a8e763a528"
#@ INITIALIZING TOKENS:
toks200 = txts[:200].map(tkn) # Getting Tokens.
toks200[0] # Inspecting Tokens.
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="ta9px3-xZfNs" outputId="b1dc6542-a1a8-4591-a774-54743f09096b"
#@ NUMERICALIZATION USING FASTAI:
num = Numericalize() # Initializing Numericalization.
num.setup(toks200) # Getting Integers.
coll_repr(num.vocab, 20) # Inspecting Vocabulary.
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="tt8fU-YNfROJ" outputId="ac4ee7a6-e7aa-4c61-f910-7c939dd7bcdb"
#@ INITIALIZING NUMERICALIZATION:
nums = num(toks)[:20]; nums # Inspection.
" ".join(num.vocab[o] for o in nums) # Getting Original Text.
# + [markdown] id="F_ZJYk8jVowc"
# ### **CREATING BATCHES FOR LANGUAGE MODEL:**
# - At every epoch I will shuffle the collection of documents and concatenate them into a stream of tokens and cut that stream into a batch of fixedsize consecutive ministreams. The model will then read the ministreams in order.
# + colab={"base_uri": "https://localhost:8080/"} id="thsG0lr0Xq1w" outputId="56362e99-1d77-4877-eee9-043d2ae0c806"
#@ CREATING BATCHES FOR LANGUAGE MODEL:
nums200 = toks200.map(num) # Initializing Numericalization.
dl = LMDataLoader(nums200) # Creating Language Model Data Loaders.
#@ INSPECTING FIRST BATCH:
x, y = first(dl) # Getting First Batch of Data.
x.shape, y.shape # Inspecting Shape of Data.
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="nYkjwoIMZEIQ" outputId="73ab7655-5530-4077-c6ee-8f8e1d4be81b"
#@ INSPECTING THE DATA:
" ".join(num.vocab[o] for o in x[0][:20]) # Inspecting Independent Variable.
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="jnoIOJ2oZsNJ" outputId="179fe7c0-6cda-491d-8d4f-0d9c8c1cc08d"
#@ INSPECTING THE DATA:
" ".join(num.vocab[o] for o in y[0][:20]) # Inspecting Dependent Variable.
# + [markdown] id="KYAR0Mx9a9DY"
# ### **TRAINING A TEXT CLASSIFIER:**
# + [markdown] id="uDYi_GrzfvqP"
# **LANGUAGE MODEL USING DATABLOCK:**
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="NjRlvq8KbbXF" outputId="5b9453c8-24e0-4fa8-f199-4d9ed3deb96a"
#@ CREATING LANGUAGE MODEL USING DATABLOCK:
get_imdb = partial(get_text_files, folders=["train", "test", "unsup"]) # Getting Text Files.
db = DataBlock(blocks=TextBlock.from_folder(path, is_lm=True), # Initializing TextBlock.
get_items=get_imdb, splitter=RandomSplitter(0.1)) # Initializing DataBlock.
#@ CREATING LANGUAGE MODEL DATALOADERS:
dls_lm = db.dataloaders(path, path=path, bs=128, seq_len=80) # Initializing Data Loaders.
# + colab={"base_uri": "https://localhost:8080/", "height": 247} id="t8zab_Z5fBgA" outputId="a75a06d5-5dde-4af7-ace8-c33647941da3"
#@ INSPECTING THE BATCHES OF DATA:
dls_lm.show_batch(max_n=2)
# + [markdown] id="Oa7FZgFegs1r"
# **FINETUNING THE LANGAUGE MODEL:**
# - I will use **Embeddings** to convert the integer word indices into activations that can be used for the neural networks. These embeddings are feed into **Recurrent Neural Network** using and architecture called **AWD-LSTM**.
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="K3Qda2V4f9O_" outputId="a47f5c8e-cdbc-4e47-c295-a0186abad3e4"
#@ INITIALIZING LANGUAGE MODEL LEARNER:
learn = language_model_learner(dls_lm, AWD_LSTM, drop_mult=0.3, # Using AWD LSTM Architecture.
metrics=[accuracy, Perplexity()]).to_fp16() # Initializing LM Learner.
# + colab={"base_uri": "https://localhost:8080/", "height": 80} id="DwMl0Jl5mZuA" outputId="859de7e5-7d5f-4dff-cf34-dcc1890894bb"
#@ TRAINING EMBEDDINGS WITH RANDOM INITIALIZATION:
learn.fit_one_cycle(1, 2e-2)
# + [markdown] id="PlBstN_DoYVY"
# **SAVING AND LOADING MODELS:**
# + colab={"base_uri": "https://localhost:8080/"} id="KJP-rFdEnDH_" outputId="e85809a6-da32-49e1-82e8-2b4eec8aaa7e"
#@ SAVING MODELS:
learn.save("/content/gdrive/MyDrive/1Epoch") # Saving the Model.
# + id="UweY10WKotCV"
#@ LOADING MODELS:
learn = learn.load("/content/gdrive/MyDrive/1Epoch") # Loading the Model.
# + colab={"base_uri": "https://localhost:8080/", "height": 359} id="cqgqbPgQxn7o" outputId="707075b6-6ad5-40fb-e3cf-a4c1d1ef1887"
#@ FINETUNING THE LANGUAGE MODEL:
learn.unfreeze() # Unfreezing the Layers.
learn.fit_one_cycle(10, 2e-3) # Training the Model.
# + [markdown] id="bKYtCD3ozJ8f"
# **ENCODER**
# - **Encoder** is defined as the model which doesn't contain task specific final layers. The term **Encoder** means much the same thing as body when applied to vision **CNN** but **Encoder** tends to be more used for NLP and generative models.
# + id="YmDWEDIfy3Pw"
#@ SAVING MODELS:
learn.save_encoder("/content/gdrive/MyDrive/FineTuned") # Saving the Model.
# + [markdown] id="SmrreawCleIT"
# ### **TEXT GENERATION:**
# - I will use the model to write new reviews.
# + id="r0qs7j0P1Hio" colab={"base_uri": "https://localhost:8080/", "height": 88} outputId="c17e773c-5e03-4745-81b6-148b7cbb66ae"
#@ INITIALIZING TEXT GENERATION:
TEXT = "I hate the movie because" # Text Example.
N_WORDS = 40
N_SENTENCES = 2
preds = [learn.predict(TEXT, N_WORDS, temperature=0.75)
for _ in range(N_SENTENCES)] # Initializing Text Generation.
print(" ".join(preds)) # Inspecting Generated Reviews.
# + [markdown] id="4wtpZHuJn966"
# ### **TEXT CLASSIFICATION:**
# - Here, I am moving towards **Classifier** fine tuning rather than **Language Model** fine tuning as mentioned above. A **Language Model** predicts the next word of a document so it doesn't require any external labels. A **Classifier** predicts an external labels.
# + colab={"base_uri": "https://localhost:8080/", "height": 247} id="xpjJuTaUo65Z" outputId="87c20cf4-06fb-4585-bd7b-31ef88eeefea"
#@ INITIALIZING DATALOADERS:
db_clas = DataBlock(blocks=(TextBlock.from_folder(path, vocab=dls_lm.vocab), # Initializing Text Blocks.
CategoryBlock), # Initializing Category Block.
get_y=parent_label, # Getting Target.
get_items=partial(get_text_files,folders=["train","test"]), # Getting Text Files.
splitter=GrandparentSplitter(valid_name="test")) # Splitting the Data.
dls_clas = db_clas.dataloaders(path, path=path, bs=128, seq_len=72) # Initializing DataLoaders.
#@ INSPECTING THE BATCHES:
dls_clas.show_batch(max_n=2) # Inspection.
# + id="m3zGfwJ119qn"
#@ CREATING MODEL FOR TEXT CLASSIFICATION:
learn = text_classifier_learner(dls_clas, AWD_LSTM, drop_mult=0.5,
metrics=accuracy).to_fp16() # Initializing Text Classifier Learner.
learn = learn.load_encoder("/content/gdrive/MyDrive/FineTuned") # Loading the Encoder.
# + [markdown] id="Fj-j8-CZ4yK0"
# **FINETUNING THE CLASSIFIER:**
# - I will train the **Classifier** with discriminative learning rates and gradaul unfreezing. In computer vision unfreezing the model at once is common approach but for **NLP Classifier** unfreezing a few layers at a time will make a real difference.
# + colab={"base_uri": "https://localhost:8080/", "height": 80} id="I998L26w2riy" outputId="a625671a-e120-433f-9b98-645b4ce01644"
#@ TRAINING THE CLASSIFIERS:
learn.fit_one_cycle(1, 2e-2) # Freezing.
# + colab={"base_uri": "https://localhost:8080/", "height": 80} id="eYa6uUIA2tyR" outputId="ee40e601-2b10-4787-8c3d-ed37812c4175"
#@ TRAINING THE CLASSIFIERS: UNFREEZING LAYERS:
learn.freeze_to(-2) # Unfreezing.
learn.fit_one_cycle(1, slice(1e-2/(2.6**4), 1e-2)) # Training the Classifier.
# + colab={"base_uri": "https://localhost:8080/", "height": 80} id="IDYKJTwf7X5B" outputId="c8e5159c-4663-4ed7-d752-49fce5473d07"
#@ TRAINING THE CLASSIFIERS: UNFREEZING LAYERS:
learn.freeze_to(-3) # Unfreezing.
learn.fit_one_cycle(1, slice(5e-3/(2.6**4), 5e-3)) # Training the Classifier.
# + colab={"base_uri": "https://localhost:8080/", "height": 111} id="yN2WgnOR7r1P" outputId="e427cd5b-82e8-4f90-ac4b-55fecc19a505"
#@ TRAINING THE CLASSIFIERS: UNFREEZING LAYERS:
learn.unfreeze() # Unfreezing.
learn.fit_one_cycle(2, slice(1e-3/(2.6**4), 1e-3)) # Training the Classifier.
| 09. Natural Language Processing/NLP.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Creating a Regression Model
#
# In this exercise, you will implement a regression model that uses features of a flight to predict how late or early it will arrive.
#
# ### Import Spark SQL and Spark ML Libraries
#
# First, import the libraries you will need:
# +
from pyspark.sql.types import *
from pyspark.sql.functions import *
from pyspark.ml.regression import LinearRegression
from pyspark.ml.feature import VectorAssembler
# -
# ### Load Source Data
# The data for this exercise is provided as a CSV file containing details of flights. The data includes specific characteristics (or *features*) for each flight, as well as a *label* column indicating how many minutes late or early the flight arrived.
#
# You will load this data into a DataFrame and display it.
wkdir ='file:///mnt/c/Users/Adura/Google Drive/Projects/Jupyter/SparkMs/data/'
csv = spark.read.csv(wkdir + 'flights.csv', inferSchema=True, header=True)
csv.show()
# ### Prepare the Data
# Most modeling begins with exhaustive exploration and preparation of the data. In this example, you will simply select a subset of columns to use as *features* as well as the **ArrDelay** column, which will be the *label* your model will predict.
data = csv.select("DayofMonth", "DayOfWeek", "OriginAirportID", "DestAirportID", "DepDelay", "ArrDelay")
data.show()
# ### Split the Data
# It is common practice when building supervised machine learning models to split the source data, using some of it to train the model and reserving some to test the trained model. In this exercise, you will use 70% of the data for training, and reserve 30% for testing.
splits = data.randomSplit([0.7, 0.3])
train = splits[0]
test = splits[1]
train_rows = train.count()
test_rows = test.count()
print("Training Rows:", train_rows, " Testing Rows:", test_rows)
# ### Prepare the Training Data
# To train the regression model, you need a training data set that includes a vector of numeric features, and a label column. In this exercise, you will use the **VectorAssembler** class to transform the feature columns into a vector, and then rename the **ArrDelay** column to **label**.
assembler = VectorAssembler(inputCols = ["DayofMonth", "DayOfWeek", "OriginAirportID", "DestAirportID", "DepDelay"], outputCol="features")
training = assembler.transform(train).select(col("features"), (col("ArrDelay").cast("Int").alias("label")))
training.show()
# ### Train a Regression Model
# Next, you need to train a regression model using the training data. To do this, create an instance of the regression algorithm you want to use and use its **fit** method to train a model based on the training DataFrame. In this exercise, you will use a *Linear Regression* algorithm - though you can use the same technique for any of the regression algorithms supported in the spark.ml API.
lr = LinearRegression(labelCol="label",featuresCol="features", maxIter=10, regParam=0.3)
model = lr.fit(training)
print("Model trained!")
# ### Prepare the Testing Data
# Now that you have a trained model, you can test it using the testing data you reserved previously. First, you need to prepare the testing data in the same way as you did the training data by transforming the feature columns into a vector. This time you'll rename the **ArrDelay** column to **trueLabel**.
testing = assembler.transform(test).select(col("features"), (col("ArrDelay")).cast("Int").alias("trueLabel"))
testing.show()
# ### Test the Model
# Now you're ready to use the **transform** method of the model to generate some predictions. You can use this approach to predict arrival delay for flights where the label is unknown; but in this case you are using the test data which includes a known true label value, so you can compare the predicted number of minutes late or early to the actual arrival delay.
prediction = model.transform(testing)
predicted = prediction.select("features", "prediction", "trueLabel")
predicted.show()
# Looking at the result, the **prediction** column contains the predicted value for the label, and the **trueLabel** column contains the actual known value from the testing data. It looks like there is some variance between the predictions and the actual values (the individual differences are referred to as *residuals*)- later in this course you'll learn how to measure the accuracy of a model.
| DataAnalyticsWithSpark/Supervised/Python Regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Scraping monsters and anticipating ACs to estimate critical threat probabilities
#
# In d&d 3.5e, rolling a "critical" is an exciting event. You get to do extra damage to the monsters you are fighting and everyone always is rather excited to roll high when it happens. While the traditional critical happens when you roll 20 on the 20 sided die, some weapons have a larger range. Calculating the probability of rolling a number in this range is fairly straightforward:
#
# | critical threat range | probability |
# |-----------------------|-------------|
# | 20 | 0.05 |
# | 19-20 | 0.10 |
# | 18-20 | 0.15 |
#
# Basically, the more sides of a die, the more increments of 0.05 you get in probability. While this step is straightforward, actually succeeding and thus causing critical damage is more complex.
#
# After your first successful critical roll you must again roll against the [Armor Class (AC)](http://www.d20srd.org/srd/combat/combatStatistics.htm) of the creature you are attacking. This produces an unknown since we typically do not know the AC of the creature we are attacking (the information is hidden by the dungeon master). Thus it becomes difficult to antipate the probability of successfully checking your critical. This notebook stands to provide some insight into these probabilities.
#
# The goal of this notebook is to do the following:
#
# 1. scrape all the stats of monsters on the [d&d 3.5e SRD](https://d20srd.org)
# 2. Calculate the average/distribution of ACs of monsters that sum to appropriate [encounter levels](http://www.d20srd.org/srd/monsters/intro.htm#challengeRating) for parties level 1-20
# 3. Calculate the probabilities of critical threats based on different critical threat ranges for each party level based on the established distribution of ACs
#
# ## What are Encounter Levels?
#
# Encounter levels are an inaccurate measurement of how difficult a battle will be for a party of level X. A 4th level party (i.e., all the party members are level 4 characters) fighting a battle of Encounter level 4 should have a reasonable amount of difficulty in the encounter. Greater than 4 its more difficult, less than 4 it is more easy. I say this is "inaccurate" because ultimately there are many things that go into whether an encounter is difficult or easy (feats and abilities, magic items the party possesses, etc.). But basically, it's all we got to generate a list of monsters suitable to fight a specific party. To generate an encounter level we sum the challenge ratings of the monsters the party is fighting. So if a zombie has a Challenge rating of 1/2 and we have a level 4 party, they should fight 8 zombies to provide an adequate challenge.
#
# ## Caveats
#
# 1. Currently, if there is more than one table describing a monster in the monster description, only the first table is included, this will probably bias the monster statistics negatively since it typically shows the least powerful monster first
# +
# %matplotlib inline
import bs4 as bs
import urllib.request
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
# -
# This pulls the list of links from the srd website for all the monsters. They are formatted in 'columns' right now the code beneath only looks at the first column (`monster_cols[0]`) but this can be changed with a simple for loop
source = urllib.request.urlopen('http://www.d20srd.org/indexes/monsters.htm').read()
soup = bs.BeautifulSoup(source,'lxml')
monster_cols = soup.findAll('ul', {'class':'column'})
# Some of the links here have sub links, thus we split on the '#' and then remove the duplicates with the list/set combo. We are probably losing some of the creatures since there is sometimes creatures that are coming in different strengths and weaknesses but thats fine for now.
# +
parsed_links = []
for col in monster_cols:
links = col.findAll('a', href=True)
for link in links:
beginning = 'http://www.d20srd.org'
link = beginning + link['href'].split('#')[0]
parsed_links.append(link)
parsed_links = list(set(parsed_links))
print('number of monsters:', len(parsed_links))
# -
# Here we take out all of the data in the statBlocks and turn it into data that will fit into a pandas dataframe. Note the comment in the try/except block. There will need to be more parsing of data done before its in a format that lends itself to statistics, but getting all the data into a tabulated format is important first, then we can tease apart the rest.
# +
from IPython.display import clear_output
errors = []
data_length = []
for i, link in enumerate(parsed_links):
if i % 3:
clear_output()
source = urllib.request.urlopen(link).read()
soup = bs.BeautifulSoup(source, 'lxml')
table = soup.findAll('table', {'class': 'statBlock'})
try:
table_rows = table[0].findAll('tr')
rows = []
for tr in table_rows:
rows.append(tr.text)
data = []
data.append(['Link', link])
for row in rows:
data.append(row.split('\n')[1:3])
if 'df' not in locals():
print('df created . . .')
df = pd.DataFrame()
df = pd.concat([df, pd.DataFrame(dict(data), index=(i,))])
else:
df = pd.concat([df, pd.DataFrame(dict(data), index=(i,))], sort=False)
print('df row {n} appended . . .'.format(n=i))
errors.append(0)
data_length.append(len(data))
except IndexError:
"""
The exceptions here are okay because they are excepting out data
that isn't appearing in the format. That is there is no statBlock
data in the links being searched so they are excepted out.
"""
errors.append(1)
data_length.append(len(data))
error_analysis = pd.DataFrame({'errors':errors, 'data_length':data_length, 'links':parsed_links})
# +
"""
all of these are monster pages that do not have any monster statistics blocks
in them
"""
error_analysis[error_analysis.errors > 0]
# +
"""
There may be some duplicates here, lets get rid of them
Also, since we care so much about AC, rows with NaN ACs are
useless. drop them too.
TODO: inspect the input data so there is no duplicates anymore
"""
df.drop_duplicates(inplace=True)
df = df[~df['Armor Class:'].isnull()].copy()
# -
df['Title'] = df.Link.apply(lambda row: row.split('/')[-1].split('.')[0])
df.shape
# Thus the raw data is parsed and we now have stats for 240 monsters
#
df.columns
# +
"""
Converting AC into integers helps with statistics
This ignores touch AC or flat footed AC
"""
df['AC(int)'] = df['Armor Class:'].apply(lambda row: row.split(' ')[0]).astype(int)
# +
"""
Challenge ratings can be fractional (e.g., goblins are weak but often
there are a lot of them). Thus we need a function that will convert the
fractional strings into floats so we can do statistics.
Additionally, there is some string handling that is done because sometimes
there are
"""
def get_float_from_fraction_str(fraction):
fraction = fraction.split('/')
numerator = float(fraction[0])
denominator = float(fraction[1])
return numerator/denominator
def convert_challenge_rating(CR):
CR_len = len(CR.split('/'))
if '/' in CR:
return get_float_from_fraction_str(CR)
elif ' ' in CR:
return CR.split(' ')[0]
else:
try:
return float(CR)
except ValueError:
import unicodedata
return unicodedata.numeric(CR)
# -
df['CR(float)'] = df['Challenge Rating:'].apply(convert_challenge_rating).astype(float)
df[['AC(int)', 'CR(float)']].describe()
# +
fig, (ax, ax2)= plt.subplots(1, 2, figsize=(10, 6), sharey=True)
df['AC(int)'].hist(bins=np.arange(0, 41, 1), grid=False, ax=ax, color='green')
df['CR(float)'].hist(bins=np.arange(0, 22, 1), grid=False, ax=ax2, color='Purple')
ax.set_xlabel('Armor Class', fontsize=15)
ax2.set_xlabel('Challenge Rating', fontsize=15)
fig, ax3 = plt.subplots(figsize=(10, 7))
ax3.scatter(x=df['CR(float)'].values, y=df['AC(int)'].values, alpha=0.5)
ax3.set_xlabel('Challenge Rating', fontsize=15)
ax3.set_ylabel('Armor Class', fontsize=15)
ax4 = fig.add_axes([0.6, 0.2, 0.25, 0.3])
ax4.scatter(x=df['CR(float)'].values, y=df['AC(int)'].values, alpha=0.5)
ax4.set_xlim(0, 5)
ax4.set_ylim(10, 20)
ax4.set_ylabel('Armor Class', fontsize=12)
ax4.set_xlabel('Challenge Rating', fontsize=12)
p = np.poly1d(np.polyfit(x=df['CR(float)'].values, y=df['AC(int)'].values, deg=2))
x = np.arange(0, 30, 0.1)
y = p(x)
ax3.plot(x, y, color='red', linewidth=3, alpha=0.5)
ax4.plot(x, y, color='red', linewidth=3, alpha=0.5)
"""
As I expected, there is a somewhat linear relation between challenge rating
and armor class, that is, as challenge rating goes up, so does armor class.
Armor class seems to be roughly normally distributed, whereas challenge rating
seems to be rougly poisson distributed.
"""
''
# -
# Calculating the average AC for encounters that the party will encounter
#
# 1. for each level party, assume that there are 4 players built exclusively
# from the SRD.
#
# 2. build a sub database of monsters of the CR equal to the party and lower
#
# 3. Bootstrap encounters from the sub database for the appropriate ECL
#
# 4. Calculate average AC
#
# 5. Calculate probability to crit successfully for attack modifiers +1 to +10
#
AC_df = df[['Link', 'Title', 'AC(int)', 'CR(float)']].copy()
# +
# AC_df[AC_df['CR(float)'] <= 4].sample(n=1)['Title'].values[0]
# +
def get_monster(monster_db):
monster = monster_db.sample(n=1)
monster_ac = monster['AC(int)'].values[0]
monster_cr = monster['CR(float)'].values[0]
monster_name = monster['Title'].values[0]
return {'name':monster_name, 'cr':monster_cr, 'ac':monster_ac}
def get_party_at_ECL(ECL, monster_db):
monster_db = monster_db[monster_db['CR(float)'] < ECL].copy()
cr = 0
monsters = []
while cr < ECL:
monster = get_monster(monster_db)
# print(monster)
cr += monster['cr']
monsters.append(monster)
# print('cr:', cr)
return monsters
get_party_at_ECL(ECL=9, monster_db=df)
# +
"""
Here we create the randomly sampled database of encounters per level
"""
party_levels = np.arange(1, 21, 1)
level_dfs = []
for level in party_levels:
level_df = pd.DataFrame()
for i in np.arange(1000):
level_df = pd.concat([level_df, pd.DataFrame(get_party_at_ECL(ECL=level, monster_db=df))])
level_dfs.append(level_df)
# +
avg_ac = np.array([level['ac'].mean() for level in level_dfs])
sem_ac = np.array([level['ac'].sem() for level in level_dfs])
fig, ax = plt.subplots(figsize=(9, 7))
ax.errorbar(x=party_levels, y=avg_ac, yerr=sem_ac*5, alpha=0.5, color='Purple', capsize=5)
ax.set_xlim(0, 21)
ax.set_xticks(party_levels)
ax.set_ylim(13, 18)
ax.set_ylabel('Average Armor Class', fontsize=15)
ax.set_xlabel('Party Level', fontsize=15)
"""
This plot was created by randomly sampling 1000 encounters per party level to produce
a sub-database of Armor Classes. These ACs are then average and the 5 sigma SEM is
calculated (thus producing the error bars).
"""
''
# -
# # Probability to beat the AC
#
# To calculate the probability to beat an AC we need the following information:
#
# 1. The [attack modifier the player will use](http://www.d20srd.org/srd/combat/combatStatistics.htm#attackBonus)
# 2. The average AC that the player will encounter
#
# For a single attack a player rolls a 20-sided die and adds the attack modifier, if its equal or higher to the AC, you hit. If the number on the die is equal to or higher than the minimum threat range, you have have threatened a critical. A subsequent role will determine if the critical threat succeeds. For a 20 sided die, to roll any one side the probability is 1/20. Thus for a longsword whose critical threat range is 19-20 then the probability to threaten is 1/10. If the player has an attack modifier of +6 and the monster they attack has an AC of 12 than the probability of successfully critically damaging is:
#
# $$P(crit success) = P(roll 19 or 20) * P(roll >= AC)$$
#
# Where
#
# $$P(roll >= AC) = 1 - (AC - modifier)/20$$
#
# Thus
#
# $$P(crit success) = 2/20 * [1 - (12 - 6)/20] = 2/20 * (1 - 6/20) = 0.07$$
#
# Doing this for the range of attack modifiers between +1 and +20 produces the below plot for a single attack.
#
# Assuming your base attack bonus is at least +6, you get an extra attacks for full round attacks. Thus the probability for these full round actions of a single successsful critical occurs changes. Because each attack is independent from each other attack we will combine the probabilities. This will produce the probability per round (or density) of a successful critical occurring.
#
# $$P(crit success) = P(roll_1 19 or 20) * P(roll_1 >= AC) * P(roll_2 19 or 20) * P(roll_2 >= AC)$$
#
# Where
#
# $$P(roll_1 >= AC) = 1 -(AC - modifier_1)/20 = 1 - (12 - 6)/20 = 0.7$$
#
# and
#
# $$P(roll_2 >= AC) = 1 - (AC - modifier_2)/20 = 1 - (12 - 1)/20 = 0.45$$
#
# Thus
#
# $$P(crit success) = 2/20 * 0.7 + 2/20 * 0.45 = 0.1145$$
#
# As we can see, the more you attack the more liekly you are to have a critical success.
# +
probability_of_threats = (np.abs(np.arange(15, 21, 1)-20)+1)/20 # for threat ranges of 15-20 to 20
average_modifier = np.arange(0, 20, 1) # assuming a stat of 10 or 11 for STR or DEX
probability_to_beat_ac = 1 - (avg_ac - average_modifier)/20
# +
fig, ax = plt.subplots(figsize=(9, 7))
ax.plot(party_levels, probability_to_beat_ac*0.3, label='15-20', linewidth=3)
ax.fill_between(x=party_levels, y1=probability_to_beat_ac*0.3-sem_ac, y2=probability_to_beat_ac*0.3+sem_ac, label=None, linewidth=3, alpha=0.3)
# ax.plot(party_levels, probability_to_beat_ac*0.25, label='16-20', linewidth=3, linestyle='--')
ax.plot(party_levels, probability_to_beat_ac*0.2, label='17-20', linewidth=3)
ax.fill_between(x=party_levels, y1=probability_to_beat_ac*0.2-sem_ac, y2=probability_to_beat_ac*0.2+sem_ac, label=None, linewidth=3, alpha=0.3)
# ax.plot(party_levels, probability_to_beat_ac*0.15, label='18-20', linewidth=3, linestyle='--')
ax.plot(party_levels, probability_to_beat_ac*0.1, label='19-20', linewidth=3)
ax.fill_between(x=party_levels, y1=probability_to_beat_ac*0.1-sem_ac, y2=probability_to_beat_ac*0.1+sem_ac, label=None, linewidth=3, alpha=0.3)
ax.plot(party_levels, probability_to_beat_ac*0.05, label='20 (default)', linewidth=3, linestyle='--', color='black', zorder=0)
ax.fill_between(x=party_levels, y1=probability_to_beat_ac*0.05-sem_ac, y2=probability_to_beat_ac*0.05+sem_ac, label=None, linewidth=3, alpha=0.3, color='black', zorder=0)
leg = ax.legend(fontsize=15)
leg.set_title('Critical Threat Range',prop={'size':15})
ax.set_ylabel('Joint probability to roll a critical\nand successfully check', fontsize=15)
ax.set_xlabel('Attack Ability Modifier', fontsize=15)
ax.set_xticks(party_levels)
ax.set_ylim(0, 0.4)
ax.set_title('A single attack at full attack', fontsize=15)
# +
fig, ax = plt.subplots(1, 2, figsize=(10, 5), sharey=True, sharex=True)
ax[0].set_title('single attack')
ax[1].set_title('full round attack')
ax[0].plot(party_levels, probability_to_beat_ac*0.3, label='15-20', linewidth=3)
ax[0].fill_between(x=party_levels, y1=probability_to_beat_ac*0.3-sem_ac, y2=probability_to_beat_ac*0.3+sem_ac, label=None, linewidth=3, alpha=0.3)
ax[0].plot(party_levels, probability_to_beat_ac*0.05, label='20 (default)', linewidth=3, linestyle='--', color='black', zorder=0)
ax[0].fill_between(x=party_levels, y1=probability_to_beat_ac*0.05-sem_ac, y2=probability_to_beat_ac*0.05+sem_ac, label=None, linewidth=3, alpha=0.3, color='black', zorder=0)
def full_round_attack(attack_modifier, threat_range):
if attack_modifier < 6:
return probability_to_beat_ac[attack_modifier]*threat_range + probability_to_beat_ac[attack_modifier]*threat_range
elif attack_modifier >= 6 and attack_modifier < 11:
return probability_to_beat_ac[attack_modifier]*threat_range + probability_to_beat_ac[attack_modifier]*threat_range + probability_to_beat_ac[attack_modifier-5]*threat_range
elif attack_modifier >= 11 and attack_modifier < 16:
return probability_to_beat_ac[attack_modifier]*threat_range + probability_to_beat_ac[attack_modifier]*threat_range + probability_to_beat_ac[attack_modifier-5]*threat_range + probability_to_beat_ac[attack_modifier-10]*threat_range
else:
return probability_to_beat_ac[attack_modifier]*threat_range + probability_to_beat_ac[attack_modifier]*threat_range + probability_to_beat_ac[attack_modifier-5]*threat_range+ probability_to_beat_ac[attack_modifier-10]*threat_range+ probability_to_beat_ac[attack_modifier-15]*threat_range
full_round_1520 = [full_round_attack(a, 0.3) for a in np.arange(0, 20, 1)]
full_round_20 = [full_round_attack(a, 0.05) for a in np.arange(0, 20, 1)]
ax[1].plot(party_levels, full_round_1520, label='15-20', linewidth=3)
ax[1].fill_between(party_levels, full_round_1520-sem_ac, full_round_1520+sem_ac, alpha=0.5)
ax[1].plot(party_levels, full_round_20, label='20', linewidth=3, color='black', linestyle='--')
ax[1].fill_between(party_levels, full_round_20-sem_ac, full_round_20+sem_ac, alpha=0.5, color='black')
leg = ax[0].legend(fontsize=15)
leg.set_title('Critical Threat Range',prop={'size':15})
ax[0].set_ylabel('Probability of successful critical')
fig.text(s='Attack Bonus', x=0.45, y=-0.02, fontsize=15)
ax[0].set_xticks(np.arange(0, 21, 1))
fig.tight_layout()
# +
p = probability_to_beat_ac[13-2]*0.1 + probability_to_beat_ac[13-2]*0.1 + probability_to_beat_ac[7-2]*0.1 + probability_to_beat_ac[7-5]*0.1
print('probability for ulrik von bek to have one successful critical strike in one round:', p)
# +
p = probability_to_beat_ac[13-2]*0.15 + probability_to_beat_ac[13-2]*0.15 + probability_to_beat_ac[7-2]*0.15 + probability_to_beat_ac[7-5]*0.15
print('probability for ulrik von bek to have one successful critical strike in one round:', p)
# -
# # How was this final plot created from start to finish?
#
# The process to create this plot is hopefully evident from the code itself, however, I will break it down here.
#
# 1. Data from [d20SRD monster database](http://www.d20srd.org/indexes/monsters.htm) was scraped and processed to produce a collection of armor classes
# 2. Using the challenge ratings of the creatures, we calculuate encounter levels of 1000 separate encounters and put them into a dataframe for encounter levels 1 through 20
# 3. This allows us to produce average ACs to calculate the second probability of success
# 4. Calculating the joint probability of successfully rolling a critical ( P(roll within critical threat range) * P(succeeding against average AC) )
# 5. Plot is produced
#
# # What results can we derive from this analysis?
#
# Obviously, the best way to increase your chances of rolling a successful critical is by increasing the critical threat range of your weapon. It's also apparent that successfully rolling a critical strike is much smaller than the probability of simply rolling in the critical threat range and the only way to increase this is to increase the attack ability modifier. To increase the critical threat range you have several options.
#
# You can do this by using the [keen spell](http://www.d20srd.org/srd/spells/keenEdge.htm), by taking the feat [Improved Critical](http://www.d20srd.org/srd/feats.htm#improvedCritical), or by using a magical weapon with the [keen ability](http://www.d20srd.org/srd/magicItems/magicWeapons.htm#keen). Unfortunately none of these stack so all things being equal, the best choice is to take Improved Critical (which requires a base attack bonus of +8).
#
# Beyond using weapons with magical damage, increasing your ability to successfully roll critical threats is good way to deal more damage. Additionally, the more times you get to roll, the higher the likelihood that you will successfully roll a critical threat.
#
# Additionally, this shows that [vorpal weapons](http://www.d20srd.org/srd/magicItems/magicWeapons.htm#vorpal) are basically useless because they require a natural 20 thus the maximum joint probability can never be much more than 5% (see the black line in the plot above). Thus the likelihood of a vorpal weapon striking true (and thus instantly killing an enemy by severing their head) is less than 1 in 20 attacks. This makes increasing your critical threat range even more enticing (compared to a vorpal sword) because of the [massive damage rules](http://www.d20srd.org/srd/combat/injuryandDeath.htm#massiveDamage). By maximizing critical damage you make it more likely to deal at least 50 points of damage in a single hit. I will add later the probability of doing 50 points of damage or more later.
#
#
| ipython-notebooks/dnd critical threat probabilities.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# +
# Import SQL Alchemy
from sqlalchemy import create_engine
import psycopg2
#Import password from passwords python file
from passwords import password
# -
#Create a connection to database
engine = create_engine(f'postgresql://Administrator:{password}@localhost:5432/SQL_Homework')
connection = engine.connect()
| Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="PRveIVUVGZUu" colab_type="code" outputId="07bf3e2a-81e5-4c09-9a88-45ec8f6ce1cc" colab={"base_uri": "https://localhost:8080/", "height": 34}
from google.colab import drive
drive.mount('/content/drive')
# + id="ORiofjZgaf7H" colab_type="code" outputId="0de1fa97-fdd9-498f-8922-7f125f4685d7" colab={"base_uri": "https://localhost:8080/", "height": 102}
# !pip install scikit-surprise
# + [markdown] id="nOrfjgotqJvA" colab_type="text"
# **Preprocessing Data**
#
# - Read data
# - Filter and clean data
# + id="VIawGSqlWhVp" colab_type="code" colab={}
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# + id="m8kSv0nTW2Oz" colab_type="code" colab={}
ratings = pd.read_csv('/content/drive/My Drive/Colab Notebooks/MovieLens/ml-latest-small/ratings.csv')
movies = pd.read_csv('/content/drive/My Drive/Colab Notebooks/MovieLens/ml-latest-small/movies.csv')
tags = pd.read_csv('/content/drive/My Drive/Colab Notebooks/MovieLens/ml-latest-small/tags.csv')
# + id="b6T7BfkJqQ0Q" colab_type="code" colab={}
movies['genres'] = movies['genres'].str.replace('|',' ')
# + id="FQLAsZMJqf8l" colab_type="code" colab={}
ratings_f = ratings.groupby('userId').filter(lambda x: len(x) >= 55)
movie_list_rating = ratings_f.movieId.unique().tolist()
# + id="muT7cm1Hqt4v" colab_type="code" outputId="5ef1a82a-0890-441b-8669-aa6ca3e410ee" colab={"base_uri": "https://localhost:8080/", "height": 34}
len(ratings_f.movieId.unique())/len(movies.movieId.unique()) * 100
# + id="U9g6jPijqzLN" colab_type="code" outputId="240f602f-9eeb-42a5-984f-2a7c55078093" colab={"base_uri": "https://localhost:8080/", "height": 34}
len(ratings_f.userId.unique())/len(ratings.userId.unique()) * 100
# + id="RpHpqwOiq-mA" colab_type="code" colab={}
movies = movies[movies.movieId.isin(movie_list_rating)]
# + id="y3vnMZuArDyD" colab_type="code" outputId="494b420f-d9b1-418c-99db-4f3bef7af160" colab={"base_uri": "https://localhost:8080/", "height": 197}
movies.head()
# + id="hpVWboYkrlVK" colab_type="code" colab={}
# map movie to id:
Mapping_file = dict(zip(movies.title.tolist(), movies.movieId.tolist()))
# + id="bjSko2WIrqCH" colab_type="code" outputId="d897ed15-2eb0-4845-a908-28ed2193bfb5" colab={"base_uri": "https://localhost:8080/", "height": 102}
tags.drop(['timestamp'],1, inplace=True)
ratings_f.drop(['timestamp'],1, inplace=True)
# + id="tGn8JcS-sHpy" colab_type="code" outputId="512a0623-e4c6-4bc3-9675-41726d8a82ac" colab={"base_uri": "https://localhost:8080/", "height": 197}
mixed = pd.merge(movies, tags, on='movieId', how='left')
mixed.head()
# + id="h5YO_2XxsJ_b" colab_type="code" outputId="f6a1ce04-0b9c-4caf-ef14-a73a80141cf6" colab={"base_uri": "https://localhost:8080/", "height": 197}
# create metadata from tags and genres
mixed.fillna("", inplace=True)
mixed = pd.DataFrame(mixed.groupby('movieId')['tag'].apply(lambda x: "%s" % ' '.join(x)))
Final = pd.merge(movies, mixed, on='movieId', how='left')
Final ['metadata'] = Final[['tag', 'genres']].apply(lambda x: ' '.join(x), axis = 1)
Final[['movieId','title','metadata']].head()
# + [markdown] id="pv5Y1GXbmnEN" colab_type="text"
# **Memory based Collaborative Filtering**
#
# - K-nearest Neighbors on user rating matrix
# + id="n7E8YMnEmy5e" colab_type="code" outputId="901dfb15-b70e-4502-bb5f-f94d1eb5ddd3" colab={"base_uri": "https://localhost:8080/", "height": 197}
ratings_f.head()
# + id="h7Nfjl86xl5c" colab_type="code" outputId="508a2a3a-47f9-4724-91ed-71ce3e098693" colab={"base_uri": "https://localhost:8080/", "height": 216}
ratings_f1 = pd.merge(movies[['movieId']], ratings_f, on="movieId", how="right")
ratings_f2 = ratings_f1.pivot(index = 'movieId', columns ='userId', values = 'rating').fillna(0)
ratings_f2.head(3)
# + id="AYLb9zRm2Tcj" colab_type="code" outputId="ddbf4642-5891-45a8-f012-7791a74ab2eb" colab={"base_uri": "https://localhost:8080/", "height": 187}
from sklearn.neighbors import NearestNeighbors
from sklearn.model_selection import KFold
model_knn = NearestNeighbors(metric='cosine', algorithm='brute', n_neighbors=20, n_jobs=-1)
kf = KFold(n_splits=10)
for train_index, test_index in kf.split(ratings_f2):
model_knn.fit(ratings_f2.iloc[train_index])
distances, indices = model_knn.kneighbors(ratings_f2.iloc[test_index], n_neighbors= 1)
print('Average Distance : %lf'%(np.sum(distances)/test_index.shape[0]))
# + [markdown] id="YSo8lQO00C9C" colab_type="text"
# **Memory based Collaborative Filtering**
#
# - K-nearest Neighbors on user rating latent matrix
# + id="W8Au_fZMx8xC" colab_type="code" outputId="2c93b681-9284-49ea-ecd3-613fea960742" colab={"base_uri": "https://localhost:8080/", "height": 254}
from sklearn.decomposition import TruncatedSVD
svd = TruncatedSVD(n_components=200)
latent_matrix_ratings = svd.fit_transform(ratings_f2)
latent_matrix_ratings_df = pd.DataFrame(latent_matrix_ratings, index=Final.title.tolist())
latent_matrix_ratings_df.head(3)
# + id="_lu_FzZByGvq" colab_type="code" outputId="64a29306-9765-44aa-eb10-3f6a20938127" colab={"base_uri": "https://localhost:8080/", "height": 281}
# plot variance expalined to see what latent dimensions to use
explained = svd.explained_variance_ratio_.cumsum()
plt.plot(explained, '.-', ms = 16, color='red')
plt.xlabel('Singular value components', fontsize= 12)
plt.ylabel('Cumulative percent of variance', fontsize=12)
plt.show()
# + id="k3dZAxWb0mUO" colab_type="code" outputId="1b6e4055-9f00-4098-a5be-dc172dfc06ff" colab={"base_uri": "https://localhost:8080/", "height": 187}
from sklearn.neighbors import NearestNeighbors
from sklearn.model_selection import KFold
model_knn = NearestNeighbors(metric='cosine', algorithm='brute', n_neighbors=20, n_jobs=-1)
kf = KFold(n_splits=10)
for train_index, test_index in kf.split(latent_matrix_ratings_df):
model_knn.fit(latent_matrix_ratings_df.iloc[train_index])
distances, indices = model_knn.kneighbors(latent_matrix_ratings_df.iloc[test_index], n_neighbors= 1)
print('Average Distance : %lf'%(np.sum(distances)/test_index.shape[0]))
# + [markdown] id="fHUWUQzZnM9l" colab_type="text"
# **Memory based Content Filtering**
#
# - TF-idf matrix with K-nearest Neighbors
# + id="c7f0uukQnznx" colab_type="code" outputId="173ae601-5b2e-406d-a9b0-4084c61ab701" colab={"base_uri": "https://localhost:8080/", "height": 137}
Final[['movieId','title','metadata']].head(3)
# + id="rho3FSc-oGEw" colab_type="code" outputId="c83c0f4c-571d-4b4a-c59d-53d9a990575b" colab={"base_uri": "https://localhost:8080/", "height": 186}
from sklearn.feature_extraction.text import TfidfVectorizer
tfidf = TfidfVectorizer(stop_words='english')
tfidf_matrix = tfidf.fit_transform(Final['metadata'])
tfidf_df = pd.DataFrame(tfidf_matrix.toarray(), index=Final.index.tolist())
tfidf_df.head(3)
# + id="3wlfIvdAoVbx" colab_type="code" outputId="f9a9ea91-ce09-4a84-88da-c8258667da20" colab={"base_uri": "https://localhost:8080/", "height": 285}
svd = TruncatedSVD(n_components=310)
latent_matrix_content_tfidf = svd.fit_transform(tfidf_df)
explained = svd.explained_variance_ratio_.cumsum()
plt.plot(explained, '.-', ms = 16, color='red')
plt.xlabel('Singular value components', fontsize= 12)
plt.ylabel('Cumulative percent of variance', fontsize=12)
plt.show()
# + id="XNFeu_SPESmf" colab_type="code" outputId="5d8dfac1-bedb-4739-dbf0-c5267a31cf6d" colab={"base_uri": "https://localhost:8080/", "height": 187}
from sklearn.neighbors import NearestNeighbors
from sklearn.model_selection import KFold
model_knn = NearestNeighbors(metric='cosine', algorithm='brute', n_neighbors=20, n_jobs=-1)
kf = KFold(n_splits=10)
for train_index, test_index in kf.split(latent_matrix_content_tfidf):
model_knn.fit(latent_matrix_content_tfidf[train_index])
distances, indices = model_knn.kneighbors(latent_matrix_content_tfidf[test_index], n_neighbors= 1)
print('Average Distance : %lf'%(np.sum(distances)/test_index.shape[0]))
# + [markdown] id="ytGeagwtlOa5" colab_type="text"
# **Model based Collaborative Filtering**
#
# - Singular value decomposition (SVD)
# - Non-negative matrix factorization (NMF)
# + id="hq9fCWY0wc8D" colab_type="code" colab={}
from surprise import SVD, NMF, KNNBasic
from surprise.model_selection import KFold
from surprise import accuracy
from surprise import Reader, Dataset
# + id="go_LZ_PxXGIp" colab_type="code" colab={}
ratings_dict = {'movieID': list(ratings.movieId), 'userID': list(ratings.userId), 'rating': list(ratings.rating)}
df = pd.DataFrame(ratings_dict)
reader = Reader(rating_scale=(0.5, 5.0))
data = Dataset.load_from_df(df[['userID', 'movieID', 'rating']], reader)
# + id="dx4At0LA1Q8r" colab_type="code" outputId="1b6d92f4-f38c-47ea-a1cc-a1aa4442423f" colab={"base_uri": "https://localhost:8080/", "height": 405}
data.df
# + id="grSqSrejXfa2" colab_type="code" outputId="6c384d76-e2e8-4d93-8417-b51c29e7e7af" colab={"base_uri": "https://localhost:8080/", "height": 187}
kf = KFold(n_splits=10)
algo = SVD()
for trainset, testset in kf.split(data):
algo.fit(trainset)
predictions = algo.test(testset)
accuracy.rmse(predictions, verbose=True)
# + colab_type="code" outputId="84acc299-cc16-49ce-909c-c02830813718" id="elkV-5nR6CdQ" colab={"base_uri": "https://localhost:8080/", "height": 187}
algo = NMF()
for trainset, testset in kf.split(data):
algo.fit(trainset)
predictions = algo.test(testset)
accuracy.rmse(predictions, verbose=True)
# + id="4RFpNBKIhVfA" colab_type="code" outputId="4b341315-7b57-4a68-db3c-430b137cc1cd" colab={"base_uri": "https://localhost:8080/", "height": 529}
algo = KNNBasic()
for trainset, testset in kf.split(data):
algo.fit(trainset)
predictions = algo.test(testset)
accuracy.rmse(predictions, verbose=True)
# + id="pMaO_XWQiMQJ" colab_type="code" colab={}
| Movie_Recommendation_System_.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python2
# ---
import pandas as pd
import sklearn as sk
import seaborn as sb
import nltk
from nltk.corpus import gutenberg
ds = pd.read_csv("B:/Data_Mining_2/train_data.csv")
ds
# +
# ds = ds.drop(ds.columns[[1,2,4,5,6,7,8]], axis=1)
# -
df = pd.DataFrame(columns=['video_id','Entertainment','Music', 'Comedy', 'People & Blogs',
'Travel & Places', 'Film & Animation', 'Sports', 'News & Politics',
'Gadgets & Games', 'Howto & DIY', 'Pets & Animals',
'Autos & Vehicles', ' UNA ','nan'])
i=0
for index,rows in ds.iterrows():
for label in range(0,20):
if(label == 0) :
toquery = rows.video_id
else :
toquery = rows[str(label)]
if(str(toquery)!='nan' and str(toquery)!='#NAME?'):
repeat_row = df[df.video_id==toquery]
if(repeat_row.size == 0):
new_row = {'video_id' : str(toquery),'Entertainment': 0,'Music': 0, 'Comedy': 0, 'People & Blogs': 0,
'Travel & Places': 0, 'Film & Animation': 0, 'Sports': 0, 'News & Politics': 0,
'Gadgets & Games': 0, 'Howto & DIY': 0, 'Pets & Animals': 0,
'Autos & Vehicles': 0, ' UNA ': 0,'nan' : 0}
s = pd.Series(new_row)
s[str(rows.category)] += 1
df.loc[i] = s
i+=1
else :
repeat_row[str(rows.category)] += 1
df.update(repeat_row)
copied = df.copy()
copied
del df['video_id']
df
answer = pd.DataFrame(columns=['video_id','category'])
i=0
for index,rows in df.iterrows():
new_row = {'category' : rows.idxmax()}
s = pd.Series(new_row)
answer.loc[i] = s
i+=1
answer['video_id'] = copied.video_id
answer
answer.to_csv('answer.csv',index=False)
test_data = pd.read_csv("B:/Data_Mining_2/test_data.csv")
test_data.insert(3,'edge_present',0)
test_data
for index,rows in test_data.iterrows():
toquery1 = rows.source
repeat_row1 = answer[answer.video_id==toquery1]
toquery2 = rows.target
repeat_row2 = answer[answer.video_id==toquery2]
if(repeat_row1.size != 0 and repeat_row2.size != 0) :
if(str(repeat_row1.category.values[0]) == str(repeat_row2.category.values[0])):
rows.edge_present = 1
test_data.loc[index] = rows
test_data
del test_data['source']
del test_data['target']
test_data
test_data.to_csv('submission2.csv',index=False)
| Youtube-Link-Recommendation/link-recommendation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/csaybar/EarthEngineMasterGIS/blob/master/module02/02_estructuradedatos.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="5sHvEhH37__p" colab_type="text"
# <!--COURSE_INFORMATION-->
# <img align="left" style="padding-right:10px;" src="https://user-images.githubusercontent.com/16768318/73986808-75b3ca00-4936-11ea-90f1-3a6c352766ce.png" width=10% >
# <img align="right" style="padding-left:10px;" src="https://user-images.githubusercontent.com/16768318/73986811-764c6080-4936-11ea-9653-a3eacc47caed.png" width=10% >
#
# **Bienvenidos!** Este *colab notebook* es parte del curso [**Introduccion a Google Earth Engine con Python**](https://github.com/csaybar/EarthEngineMasterGIS) desarrollado por el equipo [**MasterGIS**](https://www.mastergis.com/). Obten mas informacion del curso en este [**enlace**](https://www.mastergis.com/product/google-earth-engine/). El contenido del curso esta disponible en [**GitHub**](https://github.com/csaybar/EarthEngineMasterGIS) bajo licencia [**MIT**](https://opensource.org/licenses/MIT).
# + [markdown] id="XOqUVkFKBByo" colab_type="text"
# # **MASTERGIS: Sintaxis de minima de Python para GEE**
#
# En esta lectura, usted aprendera lo esencial acerca de las **estructuras de datos mas utilizadas**.
#
# Veremos los siguientes topicos:
#
# 1) listas.
# 2) diccionarios.
#
# ### **Listas**
#
# Las listas son el concepto mas basico de estructura de datos en python. A diferencia de los _string_, las **listas** pueden cambiar de elementos, es decir, son **mutables**.
#
# Para crear listas en python debemos usar [] y comas separando cada elemento.
#
# + id="6JuUJ_cQBCbM" colab_type="code" colab={}
# Asignacion de una lista
my_list = [1,2,3]
# + id="--6MQHO6BSOI" colab_type="code" colab={}
# No solamente podemos crear una lista con elementos del mismo tipo de datos.
my_list = ['A string',23,100.232,'o']
# + id="6l6c91__BXFP" colab_type="code" colab={}
# ¿Como saber la longitud de una lista?
len(my_list)
# + [markdown] id="EASVfdCDBYNe" colab_type="text"
# #### **_indexing y slicing_**
# _indexing and slicing_ funcionan de igual manera que con los _strings_. Hagamos unos ejercicios para refrescar lo aprendido.!
#
# + id="j0CMujNVBaj9" colab_type="code" colab={}
my_list = ['one','two','three',4,5]
# + id="BfVkALEqBd9i" colab_type="code" outputId="25768eac-8b22-44eb-f6aa-4a9ce9640af4" colab={"base_uri": "https://localhost:8080/", "height": 34}
# Obtengamos el primer elemento (Recurda que en Python3 la identación empieza en cero!!)
my_list[0]
# + id="CBRYk7b3Bekr" colab_type="code" outputId="c8119fac-009c-4718-cbba-03e2d7a93bb8" colab={"base_uri": "https://localhost:8080/", "height": 34}
# Obten el segundo elemento hasta el ultimo
my_list[1:]
# + id="UHC6H0uCBfLe" colab_type="code" outputId="2188f09c-93f9-4711-8127-7f5cc140e010" colab={"base_uri": "https://localhost:8080/", "height": 34}
# Obten del tercer elemento al primero
my_list[:3]
# + id="ArvC8Tf7BfzM" colab_type="code" outputId="76307ee7-b622-4ec3-c682-44d31ceafa9c" colab={"base_uri": "https://localhost:8080/", "height": 34}
#Agregemos un elemento a una lista!
my_list + ['hola']
# + id="0GAQZ-QoBhAp" colab_type="code" outputId="66ef80bd-61ce-4463-ba9f-4d0145d3bdec" colab={"base_uri": "https://localhost:8080/", "height": 34}
#Los cambios no fueron guardados.
my_list
# + id="cYuI19yQBj4m" colab_type="code" outputId="3618542c-3408-4601-c58f-6d947ae62dff" colab={"base_uri": "https://localhost:8080/", "height": 34}
#Guardando los cambios!
my_list = my_list + ['hola']
print(my_list)
# + id="i_6_Q0nDBksH" colab_type="code" outputId="eb760ed1-245c-4887-8e3a-80d6b783aaec" colab={"base_uri": "https://localhost:8080/", "height": 34}
# Duplicar mi lista
my_list * 2
# + [markdown] id="eH9-w7KXBlSS" colab_type="text"
# #### **Metodos Basicos en una lista**
#
# Si esta familiarizado con otros lenguaje de programacion (eg.C), usted podria pensar que los mismo conceptos referente a las listas son aplicables en python3. Sin embargo, las listas en python3 tienden a ser mas flexibles en comparacion a
# otros lenguajes, sobretodo porque aqui **Las listas no tienen un tamaño fijo!!**. Dicho esto,exploremos cuales son los metodos mas importantes en las listas:
# + id="xzArnKtGBptY" colab_type="code" colab={}
# Lista nueva
list1 = [1,2,3]
# + [markdown] id="vR7gbr3yBs5C" colab_type="text"
# Usamos __*append*__ para agregar un elemento permanentemente
# + id="FOJqT5j9BttY" colab_type="code" colab={}
# Append
list1.append('append me!')
# + id="ELGTsoxcBuVf" colab_type="code" outputId="a62340e1-7b58-44af-c2a0-077447d0733d" colab={"base_uri": "https://localhost:8080/", "height": 34}
print(list1)
# + [markdown] id="VZKL1xdBBu3v" colab_type="text"
# Usamos __*reverse*__ para revertir el orden de los elementos
# + id="OmKDHer2B1SR" colab_type="code" colab={}
new_list = ['a','e','x','b','c']
# + id="I45077hIB2Qe" colab_type="code" outputId="20b8520b-55d5-4b8f-f020-f67aeb135221" colab={"base_uri": "https://localhost:8080/", "height": 34}
new_list.reverse()
print(new_list)
# + [markdown] id="YQXzvCzZB4rk" colab_type="text"
# Usamos __*sort*__ para ordenar los elementos.
# + id="q_tVmODaB7uH" colab_type="code" outputId="dcd3686d-e96a-4d15-f35b-4ad5571e0b6f" colab={"base_uri": "https://localhost:8080/", "height": 34}
new_list.sort()
print(new_list)
# + [markdown] id="ZvOttTYLB8xg" colab_type="text"
# #### **Listas Anidadas**
# Las listas no solo pueden contener numeros y string, tambien pueden contener otras listas!. Veamos algunos ejemplos.
# + id="b9hFr-kwB_VC" colab_type="code" colab={}
# Creemos algunas listas de ejemplo
lst_1=[1,2,3]
lst_2=[4,5,6]
lst_3=[7,8,9]
# Creemos una lista de lista
matrix = [lst_1,lst_2,lst_3]
# + id="trUulMUACBzr" colab_type="code" outputId="36d3144b-0715-4111-d29c-f495bc40f151" colab={"base_uri": "https://localhost:8080/", "height": 34}
matrix[2][2]
# + [markdown] id="E5k0iA85CCiW" colab_type="text"
# ### **Diccionarios**
#
# Ya hemos aprendido acerca de listas, ahora aprenderemos sobre *diccionarios* en python3. Nuevamente si estas familiarizado con otros lenguajes de programacion, piensa en los diccionarios como tablas hash. Los diccionarios al igual que las listas son estructuras **mutables** , solamente que cambiamos los [] por {}.
#
# Empezos a construir diccionarios en python3!
# + id="EzHzJm3GCJ6A" colab_type="code" colab={}
# Diccionarios son una estructura dual {KEY:VALUE}
my_dict = {'key1':'value1','key2':'value2'}
# + id="WIX0VlKzCLie" colab_type="code" outputId="aaf276de-b69d-4c4f-dd65-9b04ba5c68fa" colab={"base_uri": "https://localhost:8080/", "height": 34}
#Al igual que los strings y listas podemos llamar a los elementos mediante []
my_dict['key2']
# + [markdown] id="6aG4KDz5CM4G" colab_type="text"
# Es importante tener en cuenta que los diccionarios son muy flexibles en los tipos de datos que pueden contener. Por ejemplo:
# + id="abeWUriZCRzm" colab_type="code" colab={}
my_dict = {'key1':123,'key2':[12,23,33],'key3':['item0','item1','item2']}
# + id="DWXb0EmpCSmX" colab_type="code" outputId="a2a4871e-ee4e-426c-89a6-4ceb8cc44c20" colab={"base_uri": "https://localhost:8080/", "height": 34}
# Llamemos a otro item del diccionario
my_dict['key3']
# + [markdown] id="HzZLPPrxCTN8" colab_type="text"
# ¿Recuerdas que dijimos que los diccionarios eran estructuras __mutables__?
# + id="vhBTfMnxCaK1" colab_type="code" outputId="6d6e3539-4207-414e-a81c-c74f66eed18b" colab={"base_uri": "https://localhost:8080/", "height": 51}
print(my_dict)
my_dict['key1'] = my_dict['key1'] - 10
print(my_dict)
# + [markdown] id="oG5LDcr5CazK" colab_type="text"
# #### **Metodos Basicos en un diccionario**
#
# Hay algunos metodos que podemos invocar en un diccionario. Veamos una breve introducción a algunos de ellos:
# + id="i6ZzVAxpCegM" colab_type="code" colab={}
d = {'key1':1,'key2':2,'key3':3}
# + id="QzAGeoeyCfq-" colab_type="code" outputId="665c94e9-21bc-4c99-8add-eeb1a7a13d70" colab={"base_uri": "https://localhost:8080/", "height": 34}
# retorna los key del diccionario
d.keys()
# + id="nrGDAMqWCgrq" colab_type="code" outputId="6ca8e58b-c08c-4653-b3c2-e4ceaa4c00d0" colab={"base_uri": "https://localhost:8080/", "height": 34}
# retorna los values del diccionario
d.values()
# + id="z3QwqUycChzl" colab_type="code" outputId="c77c0937-dfdb-42cd-ddb5-20562af36617" colab={"base_uri": "https://localhost:8080/", "height": 34}
# Retorna todos los elementos como una tupla (lo veremos en breve!!)
d.items()
# + [markdown] id="4x_uCsOECjRK" colab_type="text"
# #### **Diccionarios Anidados**
#
# Es de esperar que comiences a ver cuan poderoso es Python con su flexibilidad de anidar objetos y metodos de invocacion sobre ellos. Veamos como anidar diccionarios!
# + id="sJuCOIVUCnuJ" colab_type="code" colab={}
d = {'key1':{'key2':{'key3':'value1'}}}
# + id="gXDkvxlCCosb" colab_type="code" outputId="d2239456-29c3-4c69-a0f9-08cb2c85bfef" colab={"base_uri": "https://localhost:8080/", "height": 34}
# Como extraemos el valor de value1
d['key1']['key2']['key3']
# + [markdown] id="a-Kgq6o6FDWg" colab_type="text"
# ### **¿Dudas con este Jupyer-Notebook?**
#
# Estaremos felices de ayudarte!. Create una cuenta Github si es que no la tienes, luego detalla tu problema ampliamente en: https://github.com/csaybar/EarthEngineMasterGIS/issues
#
# **Tienes que dar clic en el boton verde!**
#
# <center>
# <img src="https://user-images.githubusercontent.com/16768318/79680748-d5511000-81d8-11ea-9f89-44bd010adf69.png" width = 70%>
# </center>
| module02/02_estructuradedatos.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="Tce3stUlHN0L"
# ##### Copyright 2020 The TensorFlow Authors.
# + cellView="form" id="tuOe1ymfHZPu"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="8yo62ffS5TF5"
# # Using text and neural network features
#
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/decision_forests/tutorials/intermediate_colab"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/decision-forests/blob/main/documentation/tutorials/intermediate_colab.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/decision-forests/blob/main/documentation/tutorials/intermediate_colab.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View on GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/decision-forests/documentation/tutorials/intermediate_colab.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
# </td>
# <td>
# <a href="https://tfhub.dev/google/universal-sentence-encoder/4"><img src="https://www.tensorflow.org/images/hub_logo_32px.png" />See TF Hub model</a>
# </td>
# </table>
#
# + [markdown] id="zrCwCCxhiAL7"
# Welcome to the **Intermediate Colab** for **TensorFlow Decision Forests** (**TF-DF**).
# In this colab, you will learn about some more advanced capabilities of **TF-DF**, including how to deal with natural language features.
#
# This colab assumes you are familiar with the concepts presented the [Beginner colab](beginner_colab.ipynb), notably about the installation about TF-DF.
#
# In this colab, you will:
#
# 1. Train a Random Forest that consumes text features natively as categorical sets.
#
# 1. Train a Random Forest that consumes text features using a [TensorFlow Hub](https://www.tensorflow.org/hub) module. In this setting (transfer learning), the module is already pre-trained on a large text corpus.
#
# 1. Train a Gradient Boosted Decision Trees (GBDT) and a Neural Network together. The GBDT will consume the output of the Neural Network.
# + [markdown] id="Rzskapxq7gdo"
# ## Setup
# + id="mZiInVYfffAb"
# Install TensorFlow Dececision Forests
# !pip install tensorflow_decision_forests
# + [markdown] id="2EFndCFdoJM5"
# Install [Wurlitzer](https://pypi.org/project/wurlitzer/). It can be used to show
# the detailed training logs. This is only needed in colabs.
# + id="L06XWRdSoLj5"
# !pip install wurlitzer
# + [markdown] id="i7PlfbnxYcPf"
# Import the necessary libraries.
# + id="RsCV2oAS7gC_"
import tensorflow_decision_forests as tfdf
import os
import numpy as np
import pandas as pd
import tensorflow as tf
import math
try:
from wurlitzer import sys_pipes
except:
from colabtools.googlelog import CaptureLog as sys_pipes
from IPython.core.magic import register_line_magic
from IPython.display import Javascript
# + [markdown] id="w2fsI0y5x5i5"
# The hidden code cell limits the output height in colab.
# + cellView="form" id="jZXB4o6Tlu0i"
#@title
# Some of the model training logs can cover the full
# screen if not compressed to a smaller viewport.
# This magic allows setting a max height for a cell.
@register_line_magic
def set_cell_height(size):
display(
Javascript("google.colab.output.setIframeHeight(0, true, {maxHeight: " +
str(size) + "})"))
# + [markdown] id="M_D4Ft4o65XT"
# ## Use raw text as features
#
# TF-DF can consume [categorical-set](https://arxiv.org/pdf/2009.09991.pdf) features natively. Categorical-sets represent text features as bags of words (or n-grams).
#
# For example: `"The little blue dog" ` → `{"the", "little", "blue", "dog"}`
#
# In this example, you'll will train a Random Forest on the [Stanford Sentiment Treebank](https://nlp.stanford.edu/sentiment/index.html) (SST) dataset. The objective of this dataset is to classify sentences as carrying a *positive* or *negative* sentiment. You'll will use the binary classification version of the dataset curated in [TensorFlow Datasets](https://www.tensorflow.org/datasets/catalog/glue#gluesst2).
#
# **Note:** Categorical-set features can be expensive to train. In this colab, we will train a small Random Forest with 20 trees.
# + id="SgEiFy23j14S"
# Install the nighly TensorFlow Datasets package
# TODO: Remove when the release package is fixed.
# !pip install tfds-nightly -U --quiet
# + id="uVN-j0E4Q1T3"
# Load the dataset
import tensorflow_datasets as tfds
all_ds = tfds.load("glue/sst2")
# Display the first 3 examples of the test fold.
for example in all_ds["test"].take(3):
print({attr_name: attr_tensor.numpy() for attr_name, attr_tensor in example.items()})
# + [markdown] id="UHiQUWE2XDYN"
# The dataset is modified as follows:
#
# 1. The raw labels are integers in `{-1, 1}`, but the learning algorithm expects positive integer labels e.g. `{0, 1}`. Therefore, the labels are transformed as follows: `new_labels = (original_labels + 1) / 2`.
# 1. A batch-size of 64 is applied to make reading the dataset more efficient.
# 1. The `sentence` attribute needs to be tokenized, i.e. `"hello world" -> ["hello", "world"]`.
#
#
# **Note:** This example doesn't use the `test` split of the dataset as it does not have labels. If `test` split had labels, you could concatenate the `validation` fold into the `train` one (e.g. `all_ds["train"].concatenate(all_ds["validation"])`).
#
# **Details:** Some decision forest learning algorithms do not need a validation dataset (e.g. Random Forests) while others do (e.g. Gradient Boosted Trees in some cases). Since each learning algorithm under TF-DF can use validation data differently, TF-DF handles train/validation splits internally. As a result, when you have a training and validation sets, they can always be concatenated as input to the learning algorithm.
# + id="yqYDKTKdSPYw"
def prepare_dataset(example):
label = (example["label"] + 1) // 2
return {"sentence" : tf.strings.split(example["sentence"])}, label
train_ds = all_ds["train"].batch(64).map(prepare_dataset)
test_ds = all_ds["validation"].batch(64).map(prepare_dataset)
# + [markdown] id="YYkIjROI9w43"
# Finaly, train and evaluate the model as usual. TF-DF automatically detects multi-valued categorical features as categorical-set.
#
# + id="mpxTtYo39wYZ"
# %set_cell_height 300
# Specify the model.
model_1 = tfdf.keras.RandomForestModel(num_trees=30)
# Optionally, add evaluation metrics.
model_1.compile(metrics=["accuracy"])
# Train the model.
with sys_pipes():
model_1.fit(x=train_ds)
# + [markdown] id="D9FMFGzwiHCt"
# In the previous logs, note that `sentence` is a `CATEGORICAL_SET` feature.
#
# The model is evaluated as usual:
# + id="cpf-wHl094S1"
evaluation = model_1.evaluate(test_ds)
print(f"BinaryCrossentropyloss: {evaluation[0]}")
print(f"Accuracy: {evaluation[1]}")
# + [markdown] id="YliBX4GtjncQ"
# The training logs looks are follow:
# + id="OnTTtBNmjpo7"
import matplotlib.pyplot as plt
logs = model_1.make_inspector().training_logs()
plt.plot([log.num_trees for log in logs], [log.evaluation.accuracy for log in logs])
plt.xlabel("Number of trees")
plt.ylabel("Out-of-bag accuracy")
pass
# + [markdown] id="d4qJ0ig3kgic"
# More trees would probably be beneficial (I am sure of it because I tried :p).
# + [markdown] id="Iil_oyOhCNx6"
# ## Use a pretrained text embedding
#
# The previous example trained a Random Forest using raw text features. This example will use a pre-trained TF-Hub embedding to convert text features into a dense embedding, and then train a Random Forest on top of it. In this situation, the Random Forest will only "see" the numerical output of the embedding (i.e. it will not see the raw text).
#
# In this experiment, will use the [Universal-Sentence-Encoder](https://tfhub.dev/google/universal-sentence-encoder/4). Different pre-trained embeddings might be suited for different types of text (e.g. different language, different task) but also for other type of structured features (e.g. images).
#
# **Note:** This embedding is large (1GB) and therefore the final model will be slow to run (compared to classical decision tree inference).
#
# The embedding module can be applied in one of two places:
#
# 1. During the dataset preparation.
# 2. In the pre-processing stage of the model.
#
# The second option is often preferable: Packaging the embedding in the model makes the model easier to use (and harder to misuse).
#
# First install TF-Hub:
# + id="QfYGXim_DskC"
# !pip install --upgrade tensorflow-hub
# + [markdown] id="kNSEhJgjEXww"
# Unlike before, you don't need to tokenize the text.
# + id="pS5SYqoScbOc"
def prepare_dataset(example):
label = (example["label"] + 1) // 2
return {"sentence" : example["sentence"]}, label
train_ds = all_ds["train"].batch(64).map(prepare_dataset)
test_ds = all_ds["validation"].batch(64).map(prepare_dataset)
# + id="zHEsd8q_ESpC"
# %set_cell_height 300
import tensorflow_hub as hub
# NNLM (https://tfhub.dev/google/nnlm-en-dim128/2) is also a good choice.
hub_url = "http://tfhub.dev/google/universal-sentence-encoder/4"
embedding = hub.KerasLayer(hub_url)
sentence = tf.keras.layers.Input(shape=(), name="sentence", dtype=tf.string)
embedded_sentence = embedding(sentence)
raw_inputs = {"sentence": sentence}
processed_inputs = {"embedded_sentence": embedded_sentence}
preprocessor = tf.keras.Model(inputs=raw_inputs, outputs=processed_inputs)
model_2 = tfdf.keras.RandomForestModel(
preprocessing=preprocessor,
num_trees=100)
model_2.compile(metrics=["accuracy"])
with sys_pipes():
model_2.fit(x=train_ds)
# + id="xPLoDqiFKY18"
evaluation = model_2.evaluate(test_ds)
print(f"BinaryCrossentropyloss: {evaluation[0]}")
print(f"Accuracy: {evaluation[1]}")
# + [markdown] id="WPsD3LyaMLHm"
# Note that categorical sets represent text differently from a dense embedding, so it may be useful to use both strategies jointly.
# + [markdown] id="37AGJamzboZQ"
# ## Train a decision tree and neural network together
#
# The previous example used a pre-trained Neural Network (NN) to
# process the text features before passing them to the Random Forest. This example will train both the Neural Network and the Random Forest from scratch.
#
# + [markdown] id="YJIxGwwzMkFl"
# TF-DF's Decision Forests do not back-propagate gradients ([although this is the subject of ongoing research](https://arxiv.org/abs/2007.14761)). Therefore, the training happens in two stages:
#
# 1. Train the neural-network as a standard classification task:
#
# ```
# example → [Normalize] → [Neural Network*] → [classification head] → prediction
# *: Training.
# ```
#
# 2. Replace the Neural Network's head (the last layer and the soft-max) with a Random Forest. Train the Random Forest as usual:
#
# ```
# example → [Normalize] → [Neural Network] → [Random Forest*] → prediction
# *: Training.
# ```
#
#
# + [markdown] id="YSIvuAhzbjWO"
# ### Prepare the dataset
#
# This example uses the [Palmer's Penguins](https://allisonhorst.github.io/palmerpenguins/articles/intro.html) dataset. See the [Beginner colab](beginner_colab.ipynb) for details.
# + [markdown] id="InUot_K2b3Mz"
# First, download the raw data:
# + id="rNyaeCx0b1be"
# !wget -q https://storage.googleapis.com/download.tensorflow.org/data/palmer_penguins/penguins.csv -O /tmp/penguins.csv
# + [markdown] id="pNPZzQekb9z_"
# Load a dataset into a Pandas Dataframe.
# + id="9lA3peQ4sa9a"
dataset_df = pd.read_csv("/tmp/penguins.csv")
# Display the first 3 examples.
dataset_df.head(3)
# + [markdown] id="v-_SZpRWcAoX"
#
# Prepare the dataset for training.
# + id="rtyi8UoqtzhM"
label = "species"
# Replaces numerical NaN (representing missing values in Pandas Dataframe) with 0s.
# ...Neural Nets don't work well with numerical NaNs.
for col in dataset_df.columns:
if dataset_df[col].dtype not in [str, object]:
dataset_df[col] = dataset_df[col].fillna(0)
# + id="GKrW5Yfjso0k"
# Split the dataset into a training and testing dataset.
def split_dataset(dataset, test_ratio=0.30):
"""Splits a panda dataframe in two."""
test_indices = np.random.rand(len(dataset)) < test_ratio
return dataset[~test_indices], dataset[test_indices]
train_ds_pd, test_ds_pd = split_dataset(dataset_df)
print("{} examples in training, {} examples for testing.".format(
len(train_ds_pd), len(test_ds_pd)))
# Convert the datasets into tensorflow datasets
train_ds = tfdf.keras.pd_dataframe_to_tf_dataset(train_ds_pd, label=label)
test_ds = tfdf.keras.pd_dataframe_to_tf_dataset(test_ds_pd, label=label)
# + [markdown] id="ore7f6tgcOMh"
# ### Build the models
#
# Next create the neural network model using [Keras' functional style](https://www.tensorflow.org/guide/keras/functional).
#
# To keep the example simple this model only uses two inputs.
# + id="S1Jfe4YteBqY"
input_1 = tf.keras.Input(shape=(1,), name="bill_length_mm", dtype="float")
input_2 = tf.keras.Input(shape=(1,), name="island", dtype="string")
nn_raw_inputs = [input_1, input_2]
# + [markdown] id="ZjlvAUNGeDM8"
# Use [`experimental.preprocessing` layers](https://www.tensorflow.org/guide/keras/preprocessing_layers) to convert the raw inputs to inputs apropriate for the neural netrwork.
# + id="9Q09Nkp6ei21"
# Normalization.
Normalization = tf.keras.layers.experimental.preprocessing.Normalization
CategoryEncoding = tf.keras.layers.experimental.preprocessing.CategoryEncoding
StringLookup = tf.keras.layers.experimental.preprocessing.StringLookup
values = train_ds_pd["bill_length_mm"].values[:, tf.newaxis]
input_1_normalizer = Normalization()
input_1_normalizer.adapt(values)
values = train_ds_pd["island"].values
input_2_indexer = StringLookup(max_tokens=32)
input_2_indexer.adapt(values)
input_2_onehot = CategoryEncoding(output_mode="binary", max_tokens=32)
normalized_input_1 = input_1_normalizer(input_1)
normalized_input_2 = input_2_onehot(input_2_indexer(input_2))
nn_processed_inputs = [normalized_input_1, normalized_input_2]
# + [markdown] id="ZCoQljyhelau"
# Build the body of the neural network:
# + id="KzocgbYNsH6y"
y = tf.keras.layers.Concatenate()(nn_processed_inputs)
y = tf.keras.layers.Dense(16, activation=tf.nn.relu6)(y)
last_layer = tf.keras.layers.Dense(8, activation=tf.nn.relu, name="last")(y)
# "3" for the three label classes. If it were a binary classification, the
# output dim would be 1.
classification_output = tf.keras.layers.Dense(3)(y)
nn_model = tf.keras.models.Model(nn_raw_inputs, classification_output)
# + [markdown] id="zPbRKf1CfIrj"
# This `nn_model` directly produces classification logits.
#
# Next create a decision forest model. This will operate on the high level features that the neural network extracts in the last layer before that classification head.
# + id="7fnpGNyTuXvH"
# To reduce the risk of mistakes, group both the decision forest and the
# neural network in a single keras model.
nn_without_head = tf.keras.models.Model(inputs=nn_model.inputs, outputs=last_layer)
df_and_nn_model = tfdf.keras.RandomForestModel(preprocessing=nn_without_head)
# + [markdown] id="trq07lvMudlz"
# ### Train and evaluate the models
#
# The model will be trained in two stages. First train the neural network with its own classification head:
# + id="h4OyUWKiupuF"
# %set_cell_height 300
nn_model.compile(
optimizer=tf.keras.optimizers.Adam(),
loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=["accuracy"])
nn_model.fit(x=train_ds, validation_data=test_ds, epochs=10)
nn_model.summary()
# + [markdown] id="N2mgMZOpgMQp"
# The neural network layers are shared between the two models. So now that the neural network is trained the decision forest model will be fit to the trained output of the neural network layers:
# + id="JAc9niXqud7V"
# %set_cell_height 300
df_and_nn_model.compile(metrics=["accuracy"])
with sys_pipes():
df_and_nn_model.fit(x=train_ds)
# + [markdown] id="HF8Ru2HSv1a5"
# Now evaluate the composed model:
# + id="EPMlcObzuw89"
print("Evaluation:", df_and_nn_model.evaluate(test_ds))
# + [markdown] id="awiHEznlv5sI"
# Compare it to the Neural Network alone:
# + id="--ompWYTvxM-"
print("Evaluation :", nn_model.evaluate(test_ds))
| documentation/tutorials/intermediate_colab.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## <center> SURP CTA200H Project: Understanding the distribution of galaxies in the <br> low-to-mid-redshift universe
# ### <center> <NAME>
# <center> May 14, 2021
# ### Part I: Generating Matter Power Spectra with $\verb+camb+$
# #### a) Installing $\verb+camb+$ via conda install -c conda-forge camb:
#
# +
#Importing necessary libraries and functions:
import matplotlib
from matplotlib import pyplot as plt
import matplotlib.lines as mlines
import numpy as np
import numpy.fft
import camb
from camb import model, initialpower
#Defining Plotting Style:
fsize = 15
tsize = 12
tdir = 'in'
major = 5.0
minor = 3.0
tdir = "in"
alwidth = 0.5
lwidth = 1.0
lhandle = 1.0
dpi = 100
plt.style.use('default')
plt.rcParams['text.usetex'] = True
plt.rcParams['font.size'] = fsize
plt.rcParams['xtick.major.size'] = major
plt.rcParams['xtick.minor.size'] = minor
plt.rcParams['xtick.direction'] = tdir
plt.rcParams['ytick.direction'] = tdir
plt.rcParams['axes.linewidth'] = alwidth
plt.rcParams['lines.linewidth'] = lwidth
plt.rcParams['legend.fontsize'] = tsize
plt.rcParams['legend.handlelength'] = lhandle
plt.rcParams['savefig.dpi'] = dpi
plt.rcParams['figure.figsize'] = (8,8)
# -
# #### b) Generating both linear and non-linear matter power spectra for for DES cosmology for 0.1 < z < 2.0:
#
# Set cosmological parameters from the [Dark Energy Survey Year I Results 2019](https://arxiv.org/pdf/1708.01530.pdf):
#
# $H_0$: Hubble Constant $\rm{[km\ s^{-1}\ {Mpc}^{-1}]}$\
# $h = \rm{\frac{H_0}{100\ km\ s^{-1}\ {Mpc}^{-1}}}$: Reduced Hubble Constant\
# ${\Omega_b}h^2$: Physical baryon density parameter\
# ${\Omega_c}h^2$: Physical cold dark matter density parameter\
# $n_s$: Scalar spectral index
# +
#Cosmological parameters:
h = 0.685 #[H/100 km/s/Mpc]
pars = camb.CAMBparams()
pars.set_cosmology(H0=68.5, ombh2=0.0479*h**2, omch2=0.298*h**2)
#Primordial power spectrum in the standard power law expansion:
pars.InitPower.set_params(ns=0.973)
#Parameters for calculating matter power spectra & #transfer functions:
pars.set_matter_power(redshifts=np.arange(0.5,2.25,0.5),kmax = 2.0)
#Linear Spectra:
pars.NonLinear = model.NonLinear_none
results = camb.get_results(pars)
kh, z, pk = results.get_matter_power_spectrum(minkh=1e-4, maxkh=pars.Transfer.kmax/h, npoints = 200)
s8 = np.array(results.get_sigma8())
r = results.comoving_radial_distance(z = np.arange(0.5,2.25,0.5))
#Non-linear Spectra:
pars.NonLinear = model.NonLinear_both
results.calc_power_spectra(pars)
kh_nonlin, z_nonlin, pk_nonlin = results.get_matter_power_spectrum(minkh=1e-4,\
maxkh=pars.Transfer.kmax/h, \
npoints = 200)
s8_nonlin = np.array(results.get_sigma8())
r_nonlin = results.comoving_radial_distance(z = np.arange(0.5,2.25,0.5))
# -
# #### c) Plotting the power spectra:
#
# +
#Plotting:
handles = [mlines.Line2D([], [], color='black', ls='-', label='Linear'),
mlines.Line2D([], [], color='black', ls='--', label='Non-linear')]
for i, (redshift, color) in enumerate(zip(z,["darkblue","black", "g","m"])):
plt.loglog(kh, pk[i,:], color=color, ls = "-", lw = 1.0)
plt.loglog(kh_nonlin, pk_nonlin[i,:], color=color, ls = "--", lw = 1.0)
handles.append(mlines.Line2D([],[],color=color, ls = '-', label = "z={0}".format(z[i])))
plt.xlabel(r'Wavenumber $\rm{k\ [h^{-1}\ {Mpc}^{-1}]}$')
plt.ylabel(r'Power Spectrum $\rm{P(k)\ [{(h^{-1}\ Mpc)}^3]}$')
plt.legend(handles=handles, edgecolor = "black", loc = "best")
plt.title('Linear and Non-linear Matter Power Spectra at 0.5 $\leq z \leq$ 2.0', size = 16)
plt.savefig('PowerSpectra.pdf')
# -
# #### d) Obtaining the 2-point correlation functions:
#
# The following steps were attempted to obtain the correlation function at each z, using the power spectrum, by making use of the np.fft.ifftn function. At the time of writing, the attempts were unfortunately unsuccessful, but will hopefully be resolved in the near future after additional tries.
# +
#Relating r to the wavenumber:
r = (2*np.pi)/(kh*h) #[Mpc]
r
# +
#Try for the z = 2.0 linear matter power spectrum:
#Direct application of the the ifftn function doesn't quite yield the function expected.
CR_z2 = np.real(np.fft.ifftn(pk[0,:]))
plt.plot(r, CR_z2)
plt.show() #Doesn't quite work..
| Project/CTA200Project_Khan_PartI.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.3 ('base')
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import seaborn as sns
s = pd.Series()
arr = np.array([1,2,3,4,5])
s_from_arr = pd.Series(arr)
s_from_arr
s_from_arr = pd.Series(arr, index = ['a','b','c','d','e'])
s_from_arr
dict_ = {'a':0, 'b':1, 'c': 2}
s_from_dict = pd.Series(dict_)
s_from_dict
s_from_dict[2]
s_from_dict[1:]
df = pd.DataFrame()
type(df)
d_from_arr = pd.DataFrame(arr)
d_from_arr
d_from_arr = pd.DataFrame(arr, columns=['Elements'])
d_from_arr
data = {'col_1': [3, 2, 1, 0], 'col_2': ['a', 'b', 'c', 'd']}
d_from_dict = pd.DataFrame.from_dict(data)
d_from_dict
d_from_dict.columns
d_from_dict.columns = ['A', 'B']
d_from_dict
data = [['Alex',10, 2009],['Bob',12, 2011],['Clarke',13, 2012]]
df = pd.DataFrame(data,columns=['Name','Age', 'Year'])
df
data = [{'a': 1, 'b': 2},{'a': 5, 'b': 10, 'c': 20}]
df_with_cols_rows = pd.DataFrame(data, index=['first', 'second'], columns=['a', 'b', 'c'])
df_with_cols_rows
df_with_cols_rows['a']
df_with_cols_rows[['b']]
df=pd.read_csv(r'D:\Uni\2курс(2021-2022)\Прикладная эконометрика\Econometrics1\rankmd.csv',
delimiter=';', encoding='ISO-8859-2')
df.head()
df.columns
df.info()
df.describe()
df.ndim
df.size
df.shape
df['premature_deathDeaths'].max() #sum(), mean(), min()
df.isnull()
df.isnull().sum()
df['State'].nunique()
df[df['State'] == 'Alabama']
len(df[df['State'] == 'Alabama'])
df[(df['State'] == 'Alabama') | (df['State'] == 'Wyoming')]
df['State'] = ...
sns.distplot(df['poor-or_fair_health_% Fair or Poor Health'])
| pandaslesson.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### Programming for Data Analysis - Project.
#
# #### Author: <NAME>
#
# ****
# #### Problem statement
#
# Create a data set by simulating a real-world phenomenon of your choosing. Then, rather than collect data related to the phenomenon, you should model and synthesise such data using Python [1].
#
# ****
#
# #### Project Background
#
# In a Manufacturing enviornment process classification and performance are vital to both company performance and customer satisfaction. A measure of this performance is OEE (Overall Equipment Effectiveness). The OEE concept is based off the following:
#
# 1. How often is the machine available to run?
# 2. How fast does it run when its running?
# 3. How many acceptable parts were produced?
#
# The above formula is calculated as : **** Availability x Performance X Quality (A x P x Q).**** [2].
#
# Quality is a key area of focus for Continuous Improvement (C.I) in order to improve OEE performance.
#
# In order to establish a starting point for C.I projects it is necessary to determine current process capabilities. For the purpose of this project a rough guide is used to determine the ***Process Classification***.
#
# The term ***Six sigma*** is loosely coined in this project to order process classification.
#
# Six Sigma classifies process under 7 categories: [3].
#
# 1. Level 1 - % Yield of 31% & % defective 69%.
# 2. Level 2 - % Yield of 69% & % defective 31%.
# 3. Level 3 - % Yield of 93.3% & % defective 6.7%.
# 4. Level 4 - % Yield of 99.38% & % defective 0.62%.
# 5. Level 5 - % Yield of 99.977% & % defective 0.023%.
# 6. Level 6 - % Yield of 99.99966% & % defective 0.000034%.
#
# The relationship between machine age **"Years in Production"** and defective product **"Units Scrapped"** produced from the machine is used in this instance to determine the six sigma classification. It is generally accepted that the **"Scrap Rate"** will increase as machine age.
#
# Any scrapped units are expected to negatively impact units shipped to customer.
#
# The above relationships will determine the direction used to assess and begin CI activites with the company across the 100 machines the Data is simulated for.
#
# Note:
#
# * Pandas documentation was reffered to for all DataFrame construction - https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.html
# * Seaborn & Matplotlib Documentation was reffered to for all plotting information - https://seaborn.pydata.org/ & https://matplotlib.org/
#
#
# #### References:
# * [1] Project - Programming for Data Analysis, GMIT Project assignment.
# * [2] OEE, Six Sigma Material, https://www.six-sigma-material.com/OEE.html
# * [3] Six Sigma, Wikipedia, https://en.wikipedia.org/wiki/Six_Sigma#
# * [4] 5 ways to apply an IF condition in Pandas DataFrame, data to fish, https://datatofish.com/if-condition-in-pandas-dataframe/
# * [5] Python If ... Else, w3schools,https://www.w3schools.com/python/python_conditions.asp
# * [6] cmdlinetips.com, How To Make Histogram in Python with Pandas and Seaborn?, https://cmdlinetips.com/2019/02/how-to-make-histogram-in-python-with-pandas-and-seaborn/
# importing the required libraries
import pandas as pd
import numpy as np
import random
import matplotlib.pyplot as plt
import seaborn as sns
# +
N = 280 # Batch Size for machines in Question
rng = np.random.default_rng() # initiate random number generator
Machine = pd.DataFrame({"Years in Production": rng.integers(0,10, size=100), "Batches Produced": rng.integers(0, 10, size =100),}) # Used to create Data Frame
# -
Machine ['Units Planned'] = Machine ['Batches Produced'] * N
# +
n1 = random.uniform(0, 0.0001) # Used to generate random values between 0 & 0.0001
n2 = random.uniform(0, 0.05) # Used to generate random values between 0 & 0.05
n3 = random.uniform(0, 0.1) # Used to generate random values between 0 & 0.1
# Determine Scrap Rate dependant on machine age
Machine.loc[Machine["Years in Production"] <= 2, 'Scrap Rate'] =n1
Machine.loc[Machine["Years in Production"] > 2, 'Scrap Rate'] = n2
Machine.loc[Machine["Years in Production"] >= 7, 'Scrap Rate'] = n3
# -
Machine['Units Scrapped'] = Machine ['Units Planned'] * Machine ['Scrap Rate'] # Calculate Units Scrapped
Machine ['Units Shipped'] = Machine ['Units Planned'] - Machine ['Units Scrapped'] # Calculate Units Shipped
Machine ['Percentage Yield'] = Machine ["Units Shipped"] / Machine ["Units Planned"] # Calculate Percentage Yield
# +
# Define machine classifications
Machine.loc[Machine['Units Shipped'] == 0, 'Process Classification'] = "Out of Service"
Machine.loc[Machine['Percentage Yield'] >= 0.31, 'Process Classification'] = "1 Sigma"
Machine.loc[Machine['Percentage Yield'] >= 0.69, 'Process Classification'] = "2 Sigma"
Machine.loc[Machine['Percentage Yield'] >= 0.933, 'Process Classification'] = "3 Sigma"
Machine.loc[Machine['Percentage Yield'] >= 0.9938, 'Process Classification'] = "4 Sigma"
Machine.loc[Machine['Percentage Yield'] >= 0.99977, 'Process Classification'] = "5 Sigma"
Machine.loc[Machine['Percentage Yield'] >= 0.9999966, 'Process Classification'] = "6 Sigma"
# -
Machine.round({'Units Scrapped':0,'Scrap Rate':7, 'Units Shipped':0,'Percentage Yield':7}) # Round values to required values and display DataFrame
print(Machine["Process Classification"].value_counts()) # Print Count of machine classifications
# Ref [6] Display total machine classifications.
Machine["Process Classification"].hist(bins=10, grid=False, xlabelsize=12, ylabelsize=12)
plt.xlabel("Process Classification", fontsize=12)
plt.ylabel("Count",fontsize=12)
# Ref Ianmcloughlin lecture notes
coeffs = np.polyfit(Machine["Years in Production"], Machine["Units Scrapped"], 1)
coeffs
# +
# Ref Ianmcloughlin lecture notes
plt.plot(Machine["Years in Production"], Machine["Units Scrapped"], '.', label="Units Scrapped")
plt.plot(Machine["Years in Production"], coeffs[0] * Machine["Years in Production"] + coeffs[1], '-', label='Trend-line')
plt.legend();
# -
sns.lineplot(data=Machine, x="Years in Production", y="Percentage Yield")
# #### Conculsion
#
# It is possible to conclude the following:
# 1. The total number of machine in a specific classification. This information will help guide the starting point of the process improvements.
# 2. Confirms the generally accepted theory that the defective units produced is mainly dependant on the number of years a machine is in production.
# #### End
| Programming for Data Analysis - Project.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # ALE simulation study set-up
#
# First we need to install `treeducken` and then figure out our starting parameters, write the settings files, and then
#
# + pycharm={"name": "#%% \n"}
import subprocess
import os
check_install = "../bin/treeducken/"
git_repo = "http://github.com/wadedismukes/treeducken.git"
install_cmd = ["make", "install"]
if not os.path.exists(check_install):
subprocess.call(['git', 'clone', '--depth=1', git_repo, check_install])
os.chdir(check_install + "src/")
install_process = subprocess.Popen(install_cmd, stdout=subprocess.PIPE, stderr=subprocess.PIPE)
out, err = install_process.communicate()
errcode = install_process.returncode
else:
os.chdir(check_install + "src/")
install_process = subprocess.Popen(install_cmd, stdout=subprocess.PIPE, stderr=subprocess.PIPE)
out, err = install_process.communicate()
errcode = install_process.returncode
os.chdir("../../../notebooks")
| notebooks/sim_setup.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
from pandas import read_csv
from pandas import datetime
from pandas import DataFrame
from statsmodels.tsa.arima_model import ARIMA
from matplotlib import pyplot
series = read_csv('./datasets/japan/non_seasonal_split/train.csv', parse_dates=["datetime"], index_col="datetime", squeeze=True).iloc[:,0]
# fit model
model = ARIMA(series, order=(5,1,0))
model_fit = model.fit(disp=0)
print(model_fit.summary())
# plot residual errors
residuals = DataFrame(model_fit.resid)
residuals.plot()
pyplot.show()
residuals.plot(kind='kde')
pyplot.show()
print(residuals.describe())
# -
| .ipynb_checkpoints/ARIMA-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import tensorflow as tf
# ### 1. Naći indeks najmanje i najveće vrednost u datim tenzorima. Probati sa različitim osama tenzora.
# ### 2. Naći prosek u tenzoru. Probati sa različitim osama tenzora.
# ### 3. Naći sumu tenzora. Probati sa različitim osama tenzora.
# ### 4. Naći prosek random tenzora oblika (4, 3) koji se dobija iz normalne raspodele sa prosekom $\mu=5$ i standardnom devijacijom $\sigma=7$. Obratiti pažnju da je neophodno raditi proseke po kolonama.
# ### 5. Napisati program koji rešava dati sistem linearni jednačina.
# $$
# \begin{array}{c}
# a_{11}x + a_{12}y = b_{11} \\
# a_{21}x + a_{22}y = b_{21} \\
# \end{array}
# $$
# ### 6. Napisati program koji nalazi rešenja kvadratne jednačine oblika $ax^{2} + bx + c$, za proizvoljne $a, b, c$ koeficijente, pri čemu $a\ne 0$.
# ### 7. Naći minimum funkcije $y=x^2$ korišćenjem [Gradijentalnog spusta](https://en.wikipedia.org/wiki/Gradient_descent), iterativnog algoritma za optimizaciju.
# Hint : tf.train.GradientDescentOptimizer(learning_rate).minimize(function)
| source/01_TensorFlow/Homework.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="copyright"
# Copyright 2022 Google LLC
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="title"
# # Vertex AI SDK: AutoML tabular forecasting model for batch prediction
#
# <table align="left">
# <td>
# <a href="https://colab.research.google.com/github/GoogleCloudPlatform/vertex-ai-samples/blob/main/notebooks/official/automl/sdk_automl_tabular_forecasting_batch.ipynb">
# <img src="https://cloud.google.com/ml-engine/images/colab-logo-32px.png" alt="Colab logo"> Run in Colab
# </a>
# </td>
# <td>
# <a href="https://github.com/GoogleCloudPlatform/vertex-ai-samples/blob/main/notebooks/official/automl/sdk_automl_tabular_forecasting_batch.ipynb">
# <img src="https://cloud.google.com/ml-engine/images/github-logo-32px.png" alt="GitHub logo">
# View on GitHub
# </a>
# </td>
# <td>
# <a href="https://console.cloud.google.com/vertex-ai/notebooks/deploy-notebook?download_url=https://github.com/GoogleCloudPlatform/vertex-ai-samples/blob/main/notebooks/official/automl/sdk_automl_tabular_forecasting_batch.ipynb">
# <img src="https://lh3.googleusercontent.com/UiNooY4LUgW_oTvpsNhPpQzsstV5W8F7rYgxgGBD85cWJoLmrOzhVs_ksK_vgx40SHs7jCqkTkCk=e14-rj-sc0xffffff-h130-w32" alt="Vertex AI logo">
# Open in Vertex AI Workbench
# </a>
# </td>
# </table>
# <br/><br/><br/>
# + [markdown] id="overview:automl"
# ## Overview
#
#
# This tutorial demonstrates how to use the Vertex AI SDK to create tabular forecasting models and do batch prediction using a Google Cloud [AutoML](https://cloud.google.com/vertex-ai/docs/start/automl-users) model.
# + [markdown] id="dataset:covid,forecast"
# ### Dataset
#
# The dataset used for this tutorial a time series dataset containing samples drawn from the Iowa Liquor Retail Sales dataset. Data were made available by the Iowa Department of Commerce. It is provided under the Creative Commons Zero v1.0 Universal license. For more details, see: https://console.cloud.google.com/marketplace/product/iowa-department-of-commerce/iowa-liquor-sales. This dataset does not require any feature engineering. The version of the dataset you will use in this tutorial is stored in BigQuery.
# + [markdown] id="objective:automl,training,batch_prediction"
# ### Objective
#
# In this tutorial, you create an `AutoML` tabular forecasting model from a Python script, and then do a batch prediction using the Vertex AI SDK. You can alternatively create and deploy models using the `gcloud` command-line tool or online using the Cloud Console.
#
# This tutorial uses the following Google Cloud ML services:
#
# - `AutoML Training`
# - `Vertex AI Batch Prediction`
# - `Vertex AI Model` resource
#
# The steps performed include:
#
# - Create a `Vertex AI Dataset` resource.
# - Train an `AutoML` tabular forecasting `Model` resource.
# - Obtain the evaluation metrics for the `Model` resource.
# - Make a batch prediction.
#
# + [markdown] id="costs"
# ### Costs
#
# This tutorial uses billable components of Google Cloud:
#
# * Vertex AI
# * Cloud Storage
#
# Learn about [Vertex AI
# pricing](https://cloud.google.com/vertex-ai/pricing) and [Cloud Storage
# pricing](https://cloud.google.com/storage/pricing), and use the [Pricing
# Calculator](https://cloud.google.com/products/calculator/)
# to generate a cost estimate based on your projected usage.
# + [markdown] id="setup_local"
# ### Set up your local development environment
#
# If you are using Colab or Workbench AI Notebooks, your environment already meets all the requirements to run this notebook. You can skip this step.
#
# Otherwise, make sure your environment meets this notebook's requirements. You need the following:
#
# - The Cloud Storage SDK
# - Git
# - Python 3
# - virtualenv
# - Jupyter notebook running in a virtual environment with Python 3
#
# The Cloud Storage guide to [Setting up a Python development environment](https://cloud.google.com/python/setup) and the [Jupyter installation guide](https://jupyter.org/install) provide detailed instructions for meeting these requirements. The following steps provide a condensed set of instructions:
#
# 1. [Install and initialize the SDK](https://cloud.google.com/sdk/docs/).
#
# 2. [Install Python 3](https://cloud.google.com/python/setup#installing_python).
#
# 3. [Install virtualenv](https://cloud.google.com/python/setup#installing_and_using_virtualenv) and create a virtual environment that uses Python 3. Activate the virtual environment.
#
# 4. To install Jupyter, run `pip3 install jupyter` on the command-line in a terminal shell.
#
# 5. To launch Jupyter, run `jupyter notebook` on the command-line in a terminal shell.
#
# 6. Open this notebook in the Jupyter Notebook Dashboard.
#
# + [markdown] id="install_aip:mbsdk"
# ## Installation
#
# Install the latest version of Vertex AI SDK for Python.
# + id="install_aip:mbsdk"
import os
# The Vertex AI Workbench Notebook product has specific requirements
IS_WORKBENCH_NOTEBOOK = os.getenv("DL_ANACONDA_HOME") and not os.getenv("VIRTUAL_ENV")
IS_USER_MANAGED_WORKBENCH_NOTEBOOK = os.path.exists(
"/opt/deeplearning/metadata/env_version"
)
# Vertex AI Notebook requires dependencies to be installed with '--user'
USER_FLAG = ""
if IS_WORKBENCH_NOTEBOOK:
USER_FLAG = "--user"
# ! pip3 install --upgrade google-cloud-aiplatform $USER_FLAG -q
# ! pip3 install --upgrade tensorflow $USER_FLAG -q
# + [markdown] id="restart"
# ### Restart the kernel
#
# Once you've installed the additional packages, you need to restart the notebook kernel so it can find the packages.
# + id="restart"
import os
if not os.getenv("IS_TESTING"):
# Automatically restart kernel after installs
import IPython
app = IPython.Application.instance()
app.kernel.do_shutdown(True)
# + [markdown] id="before_you_begin:nogpu"
# ## Before you begin
#
# ### GPU runtime
#
# This tutorial does not require a GPU runtime.
#
# ### Set up your Google Cloud project
#
# **The following steps are required, regardless of your notebook environment.**
#
# 1. [Select or create a Google Cloud project](https://console.cloud.google.com/cloud-resource-manager). When you first create an account, you get a $300 free credit towards your compute/storage costs.
#
# 2. [Make sure that billing is enabled for your project.](https://cloud.google.com/billing/docs/how-to/modify-project)
#
# 3. [Enable the following APIs: Vertex AI APIs, Compute Engine APIs, and Cloud Storage.](https://console.cloud.google.com/flows/enableapi?apiid=ml.googleapis.com,compute_component,storage-component.googleapis.com)
#
# 4. If you are running this notebook locally, you will need to install the [Cloud SDK]((https://cloud.google.com/sdk)).
#
# 5. Enter your project ID in the cell below. Then run the cell to make sure the
# Cloud SDK uses the right project for all the commands in this notebook.
#
# **Note**: Jupyter runs lines prefixed with `!` as shell commands, and it interpolates Python variables prefixed with `$`.
# + id="set_project_id"
PROJECT_ID = "[your-project-id]" # @param {type:"string"}
# + id="autoset_project_id"
if PROJECT_ID == "" or PROJECT_ID is None or PROJECT_ID == "[your-project-id]":
# Get your GCP project id from gcloud
shell_output = ! gcloud config list --format 'value(core.project)' 2>/dev/null
PROJECT_ID = shell_output[0]
print("Project ID:", PROJECT_ID)
# + id="set_gcloud_project_id"
# ! gcloud config set project $PROJECT_ID
# + [markdown] id="region"
# #### Region
#
# You can also change the `REGION` variable, which is used for operations
# throughout the rest of this notebook. Below are regions supported for Vertex AI. We recommend that you choose the region closest to you.
#
# - Americas: `us-central1`
# - Europe: `europe-west4`
# - Asia Pacific: `asia-east1`
#
# You may not use a multi-regional bucket for training with Vertex AI. Not all regions provide support for all Vertex AI services.
#
# Learn more about [Vertex AI regions](https://cloud.google.com/vertex-ai/docs/general/locations)
# + id="region"
REGION = "[your-region]" # @param {type: "string"}
if REGION == "[your-region]":
REGION = "us-central1"
# + [markdown] id="timestamp"
# #### Timestamp
#
# If you are in a live tutorial session, you might be using a shared test account or project. To avoid name collisions between users on resources created, you create a timestamp for each instance session, and append the timestamp onto the name of resources you create in this tutorial.
# + id="timestamp"
from datetime import datetime
TIMESTAMP = datetime.now().strftime("%Y%m%d%H%M%S")
# + [markdown] id="gcp_authenticate"
# ### Authenticate your Google Cloud account
#
# **If you are using Workbench AI Notebooks**, your environment is already authenticated. Skip this step.
#
# **If you are using Colab**, run the cell below and follow the instructions when prompted to authenticate your account via oAuth.
#
# **Otherwise**, follow these steps:
#
# In the Cloud Console, go to the [Create service account key](https://console.cloud.google.com/apis/credentials/serviceaccountkey) page.
#
# **Click Create service account**.
#
# In the **Service account name** field, enter a name, and click **Create**.
#
# In the **Grant this service account access to project** section, click the Role drop-down list. Type "Vertex" into the filter box, and select **Vertex Administrator**. Type "Storage Object Admin" into the filter box, and select **Storage Object Admin**.
#
# Click Create. A JSON file that contains your key downloads to your local environment.
#
# Enter the path to your service account key as the GOOGLE_APPLICATION_CREDENTIALS variable in the cell below and run the cell.
# + id="gcp_authenticate"
# If you are running this notebook in Colab, run this cell and follow the
# instructions to authenticate your GCP account. This provides access to your
# Cloud Storage bucket and lets you submit training jobs and prediction
# requests.
import os
import sys
# If on Vertex AI Workbench, then don't execute this code
IS_COLAB = "google.colab" in sys.modules
if not os.path.exists("/opt/deeplearning/metadata/env_version") and not os.getenv(
"DL_ANACONDA_HOME"
):
if "google.colab" in sys.modules:
from google.colab import auth as google_auth
google_auth.authenticate_user()
# If you are running this notebook locally, replace the string below with the
# path to your service account key and run this cell to authenticate your GCP
# account.
elif not os.getenv("IS_TESTING"):
# %env GOOGLE_APPLICATION_CREDENTIALS ''
# + [markdown] id="bucket:mbsdk"
# ### Create a Cloud Storage bucket
#
# **The following steps are required, regardless of your notebook environment.**
#
# When you initialize the Vertex SDK for Python, you specify a Cloud Storage staging bucket. The staging bucket is where all the data associated with your dataset and model resources are retained across sessions.
#
# Set the name of your Cloud Storage bucket below. Bucket names must be globally unique across all Google Cloud projects, including those outside of your organization.
# + id="bucket"
BUCKET_NAME = "[your-bucket-name]" # @param {type:"string"}
BUCKET_URI = f"gs://{BUCKET_NAME}"
# + id="autoset_bucket"
if BUCKET_URI == "" or BUCKET_URI is None or BUCKET_URI == "gs://[your-bucket-name]":
BUCKET_NAME = PROJECT_ID + "aip-" + TIMESTAMP
BUCKET_URI = "gs://" + BUCKET_NAME
# + [markdown] id="create_bucket"
# **Only if your bucket doesn't already exist**: Run the following cell to create your Cloud Storage bucket.
# + id="create_bucket"
# ! gsutil mb -l $REGION $BUCKET_URI
# + [markdown] id="validate_bucket"
# Finally, validate access to your Cloud Storage bucket by examining its contents:
# + id="validate_bucket"
# ! gsutil ls -al $BUCKET_URI
# + [markdown] id="setup_vars"
# ### Set up variables
#
# Next, set up some variables used throughout the tutorial.
# ### Import libraries and define constants
# + id="import_aip:mbsdk"
import google.cloud.aiplatform as aiplatform
# + [markdown] id="init_aip:mbsdk"
# ## Initialize Vertex AI SDK for Python
#
# Initialize the Vertex AI SDK for Python for your project and corresponding bucket.
# + id="init_aip:mbsdk"
aiplatform.init(project=PROJECT_ID, staging_bucket=BUCKET_URI)
# + [markdown] id="tutorial_start:automl"
# # Tutorial
#
# Now you are ready to start creating your own AutoML tabular forecasting model.
# + [markdown] id="import_file:u_dataset,csv"
# #### Location of BigQuery training data.
#
# Now set the variable `TRAINING_DATASET_BQ_PATH` to the location of the BigQuery table.
# + id="import_file:covid,csv,forecast"
TRAINING_DATASET_BQ_PATH = (
"bq://bigquery-public-data:iowa_liquor_sales_forecasting.2020_sales_train"
)
# + [markdown] id="create_dataset:tabular,forecast"
# ### Create the Dataset
#
# Next, create the `Dataset` resource using the `create` method for the `TimeSeriesDataset` class, which takes the following parameters:
#
# - `display_name`: The human readable name for the `Dataset` resource.
# - `gcs_source`: A list of one or more dataset index files to import the data items into the `Dataset` resource.
# - `bq_source`: Alternatively, import data items from a BigQuery table into the `Dataset` resource.
#
# This operation may take several minutes.
# + id="create_dataset:tabular,forecast"
dataset = aiplatform.TimeSeriesDataset.create(
display_name="iowa_liquor_sales_train" + "_" + TIMESTAMP,
bq_source=[TRAINING_DATASET_BQ_PATH],
)
time_column = "date"
time_series_identifier_column = "store_name"
target_column = "sale_dollars"
print(dataset.resource_name)
# + id="set_transformations:covid"
COLUMN_SPECS = {
time_column: "timestamp",
target_column: "numeric",
"city": "categorical",
"zip_code": "categorical",
"county": "categorical",
}
# + [markdown] id="create_automl_pipeline:tabular,forecast"
# ### Create and run training job
#
# To train an AutoML model, you perform two steps: 1) create a training job, and 2) run the job.
#
# #### Create training job
#
# An AutoML training job is created with the `AutoMLForecastingTrainingJob` class, with the following parameters:
#
# - `display_name`: The human readable name for the `TrainingJob` resource.
# - `column_transformations`: (Optional): Transformations to apply to the input columns
# - `optimization_objective`: The optimization objective to minimize or maximize.
# - `minimize-rmse`
# - `minimize-mae`
# - `minimize-rmsle`
#
# The instantiated object is the job for the training pipeline.
# + id="create_automl_pipeline:tabular,forecast"
MODEL_DISPLAY_NAME = f"iowa-liquor-sales-forecast-model_{TIMESTAMP}"
training_job = aiplatform.AutoMLForecastingTrainingJob(
display_name=MODEL_DISPLAY_NAME,
optimization_objective="minimize-rmse",
column_specs=COLUMN_SPECS,
)
# + [markdown] id="run_automl_pipeline:forecast"
# #### Run the training pipeline
#
# Next, you start the training job by invoking the method `run`, with the following parameters:
#
# - `dataset`: The `Dataset` resource to train the model.
# - `model_display_name`: The human readable name for the trained model.
# - `training_fraction_split`: The percentage of the dataset to use for training.
# - `test_fraction_split`: The percentage of the dataset to use for test (holdout data).
# - `target_column`: The name of the column to train as the label.
# - `budget_milli_node_hours`: (optional) Maximum training time specified in unit of millihours (1000 = hour).
# - `time_column`: Time-series column for the forecast model.
# - `time_series_identifier_column`: ID column for the time-series column.
#
# The `run` method when completed returns the `Model` resource.
#
# The execution of the training pipeline will take up to one hour.
# + id="run_automl_pipeline:forecast"
model = training_job.run(
dataset=dataset,
target_column=target_column,
time_column=time_column,
time_series_identifier_column=time_series_identifier_column,
available_at_forecast_columns=[time_column],
unavailable_at_forecast_columns=[target_column],
time_series_attribute_columns=["city", "zip_code", "county"],
forecast_horizon=30,
context_window=30,
data_granularity_unit="day",
data_granularity_count=1,
weight_column=None,
budget_milli_node_hours=1000,
model_display_name=MODEL_DISPLAY_NAME,
predefined_split_column_name=None,
)
# + [markdown] id="evaluate_the_model:mbsdk"
# ## Review model evaluation scores
#
# After your model training has finished, you can review the evaluation scores for
# + id="evaluate_the_model:mbsdk"
model_evaluations = model.list_model_evaluations()
for model_evaluation in model_evaluations:
print(model_evaluation.to_dict())
# + [markdown] id="make_prediction"
# ## Send a batch prediction request
#
# Send a batch prediction to your deployed model.
# + [markdown] id="batch_request:mbsdk,both_csv"
# ### Make the batch prediction request
#
# Now that your Model resource is trained, you can make a batch prediction by invoking the batch_predict() method using a BigQuery source and destination, with the following parameters:
#
# - `job_display_name`: The human readable name for the batch prediction job.
# - `bigquery_source`: BigQuery URI to a table, up to 2000 characters long. For example: `bq://projectId.bqDatasetId.bqTableId`
# - `bigquery_destination_prefix`: The BigQuery dataset or table for storing the batch prediction resuls.
# - `instances_format`: The format for the input instances. Since a BigQuery source is used here, this should be set to `bigquery`.
# - `predictions_format`: The format for the output predictions, `bigquery` is used here to output to a BigQuery table.
# - `generate_explanations`: Set to `True` to generate explanations.
# - `sync`: If set to True, the call will block while waiting for the asynchronous batch job to complete.
# + id="2c9d935c6ab9"
import os
from google.cloud import bigquery
batch_predict_bq_output_dataset_name = f"iowa_liquor_sales_predictions_{TIMESTAMP}"
batch_predict_bq_output_dataset_path = "{}.{}".format(
PROJECT_ID, batch_predict_bq_output_dataset_name
)
batch_predict_bq_output_uri_prefix = "bq://{}.{}".format(
PROJECT_ID, batch_predict_bq_output_dataset_name
)
# Must be the same region as batch_predict_bq_input_uri
client = bigquery.Client(project=PROJECT_ID)
bq_dataset = bigquery.Dataset(batch_predict_bq_output_dataset_path)
dataset_region = "US" # @param {type : "string"}
bq_dataset.location = dataset_region
bq_dataset = client.create_dataset(bq_dataset)
print(
"Created bigquery dataset {} in {}".format(
batch_predict_bq_output_dataset_path, dataset_region
)
)
# + [markdown] id="99b7a9287ba6"
# For AutoML models, manual scaling can be adjusted by setting both min and max nodes i.e., `starting_replica_count` and `max_replica_count` as the same value(in this example, set to 1). The node count can be increased or decreased as required by load.
#
# `batch_predict` can export predictions either to BigQuery or GCS. The BigQuery options are commented out below and the predictions will be exported to the BUCKET_URI.
# + id="batch_request:mbsdk,both_csv"
PREDICTION_DATASET_BQ_PATH = (
"bq://bigquery-public-data:iowa_liquor_sales_forecasting.2021_sales_predict"
)
batch_prediction_job = model.batch_predict(
job_display_name=f"iowa_liquor_sales_forecasting_predictions_{TIMESTAMP}",
bigquery_source=PREDICTION_DATASET_BQ_PATH,
instances_format="bigquery",
bigquery_destination_prefix=batch_predict_bq_output_uri_prefix,
predictions_format="bigquery",
generate_explanation=True,
sync=False,
)
print(batch_prediction_job)
# + [markdown] id="batch_request_wait:mbsdk"
# ### Wait for completion of batch prediction job
#
# Next, wait for the batch job to complete. Alternatively, you can set the parameter `sync` to `True` in the `batch_predict()` method to block until the batch prediction job is completed.
# + id="batch_request_wait:mbsdk"
batch_prediction_job.wait()
# + [markdown] id="get_batch_prediction:mbsdk,forecast"
# ### Get the predictions and explanations
#
# Next, get the results from the completed batch prediction job and print them out. Each result row will include the prediction and explanation.
# + id="get_batch_prediction:mbsdk,forecast"
for row in batch_prediction_job.iter_outputs():
print(row)
# + [markdown] id="78080aa9088e"
# ### Visualize the forecasts
#
# Lastly, follow the given link to visualize the generated forecasts in [Data Studio](https://support.google.com/datastudio/answer/6283323?hl=en).
# The code block included in this section dynamically generates a Data Studio link that specifies the template, the location of the forecasts, and the query to generate the chart. The data is populated from the forecasts generated using BigQuery options where the destination dataset is `batch_predict_bq_output_dataset_path`.
#
# You can inspect the used template at https://datastudio.google.com/c/u/0/reporting/067f70d2-8cd6-4a4c-a099-292acd1053e8. This was created by Google specifically to view forecasting predictions.
#
# **Note:** The Data Studio dashboard can only show the charts properly when the `batch_predict` job is run successfully using the BigQuery options.
# + id="f82f00be2160"
import urllib
tables = client.list_tables(batch_predict_bq_output_dataset_path)
prediction_table_id = ""
for table in tables:
if (
table.table_id.startswith("predictions_")
and table.table_id > prediction_table_id
):
prediction_table_id = table.table_id
batch_predict_bq_output_uri = "{}.{}".format(
batch_predict_bq_output_dataset_path, prediction_table_id
)
def _sanitize_bq_uri(bq_uri):
if bq_uri.startswith("bq://"):
bq_uri = bq_uri[5:]
return bq_uri.replace(":", ".")
def get_data_studio_link(
batch_prediction_bq_input_uri,
batch_prediction_bq_output_uri,
time_column,
time_series_identifier_column,
target_column,
):
batch_prediction_bq_input_uri = _sanitize_bq_uri(batch_prediction_bq_input_uri)
batch_prediction_bq_output_uri = _sanitize_bq_uri(batch_prediction_bq_output_uri)
base_url = "https://datastudio.google.com/c/u/0/reporting"
query = (
"SELECT \\n"
" CAST(input.{} as DATETIME) timestamp_col,\\n"
" CAST(input.{} as STRING) time_series_identifier_col,\\n"
" CAST(input.{} as NUMERIC) historical_values,\\n"
" CAST(predicted_{}.value as NUMERIC) predicted_values,\\n"
" * \\n"
"FROM `{}` input\\n"
"LEFT JOIN `{}` output\\n"
"ON\\n"
"CAST(input.{} as DATETIME) = CAST(output.{} as DATETIME)\\n"
"AND CAST(input.{} as STRING) = CAST(output.{} as STRING)"
)
query = query.format(
time_column,
time_series_identifier_column,
target_column,
target_column,
batch_prediction_bq_input_uri,
batch_prediction_bq_output_uri,
time_column,
time_column,
time_series_identifier_column,
time_series_identifier_column,
)
params = {
"templateId": "067f70d2-8cd6-4a4c-a099-292acd1053e8",
"ds0.connector": "BIG_QUERY",
"ds0.projectId": PROJECT_ID,
"ds0.billingProjectId": PROJECT_ID,
"ds0.type": "CUSTOM_QUERY",
"ds0.sql": query,
}
params_str_parts = []
for k, v in params.items():
params_str_parts.append('"{}":"{}"'.format(k, v))
params_str = "".join(["{", ",".join(params_str_parts), "}"])
return "{}?{}".format(base_url, urllib.parse.urlencode({"params": params_str}))
print(
get_data_studio_link(
PREDICTION_DATASET_BQ_PATH,
batch_predict_bq_output_uri,
time_column,
time_series_identifier_column,
target_column,
)
)
# + [markdown] id="cleanup:mbsdk"
# # Cleaning up
#
# To clean up all Google Cloud resources used in this project, you can [delete the Google Cloud
# project](https://cloud.google.com/resource-manager/docs/creating-managing-projects#shutting_down_projects) you used for the tutorial.
#
# Otherwise, you can delete the individual resources you created in this tutorial:
#
# - Dataset
# - AutoML Training Job
# - Model
# - Batch Prediction Job
# - Cloud Storage Bucket
# + id="cleanup:mbsdk"
# Set this to true only if you'd like to delete your bucket
delete_bucket = False
# Delete dataset
dataset.delete()
# Training job
training_job.delete()
# Delete model
model.delete()
# Delete batch prediction job
batch_prediction_job.delete()
if delete_bucket or os.getenv("IS_TESTING"):
# ! gsutil rm -r $BUCKET_URI
| notebooks/official/automl/sdk_automl_tabular_forecasting_batch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Emotion Classification
# **Module 3: Data acquisition**
# * Author: [<NAME>](https://github.com/andresmitre), [Center for Research in Mathematics (CIMAT)](http://www.cimat.mx/en) in Zacatecas, México.
#
# Module |Title
# ---------|--------------
# Module 1 |[Introduction](https://github.com/andresmitre/Emotion_Classification/blob/master/introduction.ipynb)
# Module 2 |[Haar Cascade Algorithm](https://github.com/andresmitre/Emotion_Classification/blob/master/Haar_Feature_based_Cascade_Classifiers.ipynb)
# Module 3 |[Data acquisition](https://github.com/andresmitre/Emotion_Classification/blob/master/data_acquisition.ipynb)
# Module 4 |[Convolutional Neural Network](https://github.com/andresmitre/Emotion_Classification/blob/master/CNN.ipynb)
#
# # Goals
#
# * Capture RAW data from FER and GSR.
# * Export data to a CSV file.
#
# # Raw Data
#
# The data collection, starts with a FER(Facial Expression Recognition) using [haarcascades](https://github.com/opencv/opencv), with a professional camera/webcam. Images were taken while 23 participants watched a series of films related with emotions. The images were saved into separate emotions folders. GSR lectures were recorded as well since stress and boredom emotions stimulates the activity of sweet glands. the GSR were captured with the [Grove - GSR sensor](https://www.seeedstudio.com/Grove-GSR-sensor-p-1614.html) and saved into a CSV file.
#
# **Galvanic Skin Response (GSR)**
#
# The Galvanic Skin Response (GSR) is defined as a change in the electrical properties of the skin. The signal can be used for capturing the autonomic nerve responses as a parameter of the sweat gland function. The measurement is relatively simple, and has a good repeatability. Therefore the GSR measurement can be considered to be a simple and useful tool for examination of the autonomous nervous system function, and especially the peripheral sympathetic system.
#
# For detailed information I highly reccomend to check [The complete pocket guide by IMOTIONS](https://imotions.com/guides/).
#
# * Materials required:
# * **Camera** - [EOS Rebel T3 – Canon Profesional.](https://www.amazon.com/Canon-Digital-18-55mm-discontinued-manufacturer/dp/B004J3Y9U6).
# * **Monitor** - [LCD 22 in, TFT FPD2275W-MX](https://www.cnet.com/products/gateway-fpd2275w/specs/).
# * **Keyboard** - DELL SK-8115.
# * **Speakers** - Subwoofer DELL A525.
# * **GSR Sensor** - [Grove - GSR Sensor](https://www.seeedstudio.com/Grove-GSR-sensor-p-1614.html).
# * **Computer** - [Dell Inspiron 13-7359 Signature Edition](http://www.dell.com/en-us/shop/dell-laptops/new-inspiron-13-7000-series-2-in-1-laptop/spd/inspiron-13-7359-laptop).
# * Films Stimulants:
#
# Emotion |Film |Director
# ---------|---------------------|--------
# Neutral |The Lover |<NAME>
# Happy |When Harry Met Sally |<NAME>
# Stress |Irreversible |Gaspar Noé
# Boredom |Amateur film |[<NAME>. and <NAME>.](https://www.youtube.com/watch?v=s34zGmq3rXQ)
#
# # Procedure
#
# The emotions from the above table were validated by more than 100 participants in the documentation project [[contact Author]](https://github.com/andresmitre). the collection of raw data takes place in a lapse of 30 seconds in every emotion. The Raw data was collected in the next sequence:
#
# * Sequence:
# * **Neutral > Stress > Neutral > Boredom > Neutral > Happy**.
#
# The photos were taken at 10FPS. a total of 1380 images were selected as the training data set. where 10 images per emotion of every participant were captured. The images were selected according to the biggest difference between the emotion and the neutral.
#
# 
#
# Participant during the experiment.
#
# ##CSV Files
#
# The raw data was collected into a CSV file. Each participant had their each CSV file for every emotion. Time, ID, Emotion and Picture number were the fields collected into de CSV, to identify the corresponding image to the specific time. The next figure represents an example of the data collected coresponding to the participant 3 (S3).
#
#
# ##FER Dataset
# The Facial Expression Recognition images were saved into seperate directories (named with the emotion) every single image were recorded in the next format:
#
# * **Emotion + ID of the participant + Picture Number**.
#
# 
#
# 
#
#
# ##Source Code
# The main code is posted [here](https://github.com/andresmitre/Emotion_Classification/blob/master/esponteanea.py)
#
#
| data_acquisition.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import tensorflow as tf
import numpy as np
import matplotlib
import os
tf.set_random_seed(777)
if "DISPLAY" not in os.environ:
# remove Travis CI Error
matplotlib.use('Agg')
import matplotlib.pyplot as plt
def MinMaxScalar(data):
''' Min Max Normalization
References
[1] http://sebastianraschka.com/Articles/2014_about_feature_scaling.html'''
numerator = data - np.min(data, 0)
denominator = np.max(data, 0) - np.min(data, 0)
return numerator / (denominator + 1e-7)
#Train Parameter
seq_length = 7
data_dim = 5
hidden_dim = 10
output_dim = 1
learning_rate = 0.01
iterations = 500
# Open, High, Low, Volume, Close
xy = np.loadtxt('data-02-stock_daily.csv', delimiter=',')
xy = xy[::-1] # reverse order (chronically ordered)
xy = MinMaxScalar(xy)
x = xy
y = xy[:, [-1]] # Close as label
#Bulid a dataset
dataX = []
dataY = []
for i in range (0,len(y) - seq_length):
_x = x[i:i + seq_length]
_y = y[i + seq_length] #Next close price
#print(_x, "->", _y)
dataX.append(_x)
dataY.append(_y)
#train/test split
train_size = int(len(dataY) * 0.7)
test_size = len(dataY) - train_size
trainX, testX = np.array(dataX[0:train_size]), np.array(dataX[train_size:len(dataX)])
trainY, testY = np.array(dataY[0:train_size]), np.array(dataY[train_size:len(dataY)])
#input placeholder
X = tf.placeholder(tf.float32, [None, seq_length, data_dim])
Y = tf.placeholder(tf.float32, [None,1])
#build a LSTM network
cell = tf.contrib.rnn.BasicLSTMCell(num_units = hidden_dim, state_is_tuple = True, activation = tf.tanh)
outputs, _states = tf.nn.dynamic_rnn(cell, X, dtype=tf.float32)
Y_pred = tf.contrib.layers.fully_connected(
outputs[:, -1], output_dim, activation_fn = None) #We use the last cell's output
# +
#loss function
loss = tf.reduce_sum(tf.square(Y_pred - Y)) #sum of the squares
#optimizer
optimizer = tf.train.AdamOptimizer(learning_rate)
train = optimizer.minimize(loss)
# -
#RMSE
targets = tf.placeholder(tf.float32, [None, 1])
predictions = tf.placeholder(tf.float32, [None, 1])
rmse = tf.sqrt(tf.reduce_mean(tf.square(targets - predictions)))
with tf.Session() as sess:
init = tf.global_variables_initializer()
sess.run(init)
#Training step
for i in range(iterations):
_, step_loss = sess.run([train, loss], feed_dict ={
X: trainX, Y: trainY
})
#Test step
test_predict = sess.run(Y_pred, feed_dict = {X: testX})
rmse_val = sess.run(rmse, feed_dict ={
targets: testY, predictions : test_predict
})
print("RMSE: {}".format(rmse_val))
#Plot predictions
plt.plot(testY)
plt.plot(test_predict)
plt.xlabel("Time Period")
plt.ylabel("Stock Price")
plt.show()
| Lab_12_Rnn_Stock_Prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.5 64-bit
# name: python3
# ---
os.add_dll_directory(os.path.join(os.environ['CUDA_PATH'], 'bin'))
import cv2
import mediapipe as mp
import time
def fancyDraw(img, x, y, w, h, l=30, t=5, rt= 1):
x1, y1 = x + w, y + h
# Top Left x,y
cv2.line(img, (x, y), (x + l, y), (255, 0, 255), t)
cv2.line(img, (x, y), (x, y+l), (255, 0, 255), t)
# Top Right x1,y
cv2.line(img, (x1, y), (x1 - l, y), (255, 0, 255), t)
cv2.line(img, (x1, y), (x1, y+l), (255, 0, 255), t)
# Bottom Left x,y1
cv2.line(img, (x, y1), (x + l, y1), (255, 0, 255), t)
cv2.line(img, (x, y1), (x, y1 - l), (255, 0, 255), t)
# Bottom Right x1,y1
cv2.line(img, (x1, y1), (x1 - l, y1), (255, 0, 255), t)
cv2.line(img, (x1, y1), (x1, y1 - l), (255, 0, 255), t)
return img
# +
cap = cv2.VideoCapture(0)
cap.set(cv2.CAP_PROP_FRAME_WIDTH, 1280);
cap.set(cv2.CAP_PROP_FRAME_HEIGHT, 720);
#cap.set(cv2.CAP_PROP_FOURCC, cv2.VideoWriter_fourcc('M', 'J', 'P', 'G'));
#cap.set(cv2.CAP_PROP_FOURCC, cv2.VideoWriter_fourcc('A', 'V', 'C', '1'));
cap.set(cv2.CAP_PROP_FOURCC, cv2.VideoWriter_fourcc('H', '2', '6', '5')); #Daha iyi sıkıştırma sağlıyor.
cap.set(cv2.CAP_PROP_FPS, 60);
pTime = 0
mpFaceDetection = mp.solutions.face_detection
mpDraw = mp.solutions.drawing_utils
faceDetection = mpFaceDetection.FaceDetection(0.5)
while True:
success, img = cap.read()
imgRGB = cv2.cvtColor(img, cv2.COLOR_BGR2RGB)
results = faceDetection.process(imgRGB)
if results.detections:
for id, detection in enumerate(results.detections):
mpDraw.draw_detection(img, detection)
bboxC = detection.location_data.relative_bounding_box
ih, iw, ic = img.shape
bbox = [int(bboxC.xmin * iw), int(bboxC.ymin * ih), int(bboxC.width * iw), int(bboxC.height * ih)]
img = fancyDraw(img, int(bboxC.xmin * iw), int(bboxC.ymin * ih), int(bboxC.width * iw), int(bboxC.height * ih), l=30, t=5, rt=1)
cv2.putText(img, f'{int(detection.score[0] * 100)}%',(bbox[0], bbox[1] - 20),cv2.FONT_HERSHEY_PLAIN,2,(255, 0, 255), 2)
cTime = time.time()
fps = 1 / (cTime - pTime)
pTime = cTime
cv2.putText(img, f'FPS: {int(fps)}', (20, 70), cv2.FONT_HERSHEY_PLAIN, 3, (0, 255, 0), 2)
# Oluşan çerçeveyi ekrana yansıt
cv2.imshow('Video', img)
# Çıkış için 'q'
if cv2.waitKey(1) & 0xFF == ord('q'):
break
#end
cv2.destroyAllWindows();
cap.release();
# -
cv2.destroyAllWindows();
cap.release();
ih, iw, ic = img.shape
cv2.imshow('Video', imgRGB[bbox[1]:bbox[1]+bbox[2], bbox[0]:bbox[0]+bbox[3]])
cv2.waitKey(0)
| Face/mediapipe_face/faceD__mediapipe_BlazeFace(SSD-MobileNet1-2).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.10 64-bit (''base'': conda)'
# name: python3
# ---
# # Figure 4. Individual case study
# ## Import libraries
import pandas as pd
import altair as alt
from helper import RawData, InteractivePlot
# ## Import part II data (Expanded control parameter range)
raw = RawData("../../data/data_part2_1750.csv")
df = raw.get(epoch_less_than=0.3)
# ## Plot figure 4
fig4 = InteractivePlot(df=df) # see helper.py for details
fig4.plot()
# Figure 4 is just four selected setting in the interactive plot:
# 1. Baseline: P-Noise=0; Hidden=100; Epsilon=.006
# 2. Phonological dyslexia: **P-Noise=8**; Hidden=100; Epsilon=.006
# 3. Surface dyslexia: P-Noise=0; **Hidden=25**; Epsilon=.006
# 4. Developmental delay: P-Noise=0; Hidden=100; **Epsilon=.002**
| code/python/figure4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import os
import numpy as np
import matplotlib.pyplot as plt
from matplotlib import colors
bool_cmap = colors.ListedColormap([(1, 1, 1, 0), 'black'])
from scipy.constants import centi, milli
from fastadjust.io import h5read
# SIMION array
fil = os.path.join(r"../data", "quad.h5")
fa = h5read(fil)
# move xy origin to center of the grid
fa.x0 = - (fa.nx - 1) * fa.dx / 2
fa.y0 = - (fa.ny - 1) * fa.dy / 2
# ## electrode geometry
# +
# z grid position
zg = 75
el = fa.electrode
# electrodes
fig, ax = plt.subplots(figsize=(9, 4))
c0 = ax.imshow(el[:, :, zg].T, origin='lower', extent=fa.extent[:4] / milli, cmap=bool_cmap, vmin=0, vmax=1)
cbar = fig.colorbar(c0, label='electrode')
ax.set_aspect('equal')
ax.set_xlabel('x (mm)')
ax.set_ylabel('y (mm)')
plt.show()
# -
# ## electric potential
# +
voltages = np.array([2.5, 2.5, -2.5, -2.5], dtype='float64')
phi = fa.potential(voltages)
# potential
fig, ax = plt.subplots(figsize=(9, 4))
c0 = ax.imshow(phi[:, :, zg].T, origin='lower', extent=fa.extent[:4] / milli, cmap='RdBu')
cbar = fig.colorbar(c0, label='potential (V)')
ax.set_aspect('equal')
ax.set_xlabel('x (mm)')
ax.set_ylabel('y (mm)')
plt.show()
# +
X, Y, Z = fa.grid()
# potential
fig, ax = plt.subplots(figsize=(9,4))
c0 = ax.contourf(X[:, :, zg] * 1e3, Y[:, :, zg] * 1e3, phi[:, :, zg], cmap='RdBu')
cbar = fig.colorbar(c0, label='potential (V)')
ax.set_aspect('equal')
ax.set_xlabel('x (mm)')
ax.set_ylabel('y (mm)')
plt.show()
# -
# ## electric field
# +
# subset
xmin, ymin, zmin = np.round(fa.grid_r((-0.003, -0.002, 0)))
xmax, ymax, zmax = np.round(fa.grid_r((0.0032, 0.0022, 0)))
subset = (slice(int(xmin), int(xmax), 2), slice(int(ymin), int(ymax), 2), zg)
# field
ex, ey, ez = fa.field(voltages)
fig, ax = plt.subplots(figsize=(6, 6))
ax.quiver(X[subset] * 1e3, Y[subset] * 1e3, ex[subset], ey[subset], angles='xy')
ax.set_aspect('equal')
ax.set_xlabel('x (mm)')
ax.set_ylabel('y (mm)')
plt.show()
# -
# ## field amplitude
# +
famp = fa.amp_field(voltages)
fig, ax = plt.subplots(figsize=(9, 4))
c0 = ax.imshow(famp[:, :, zg].T * centi, origin='lower', extent=fa.extent[:4] / milli, cmap='viridis')
cbar = fig.colorbar(c0, label='electric field (V / cm)')
ax.set_aspect('equal')
ax.set_xlabel('x (mm)')
plt.show()
# -
fig, ax = plt.subplots(figsize=(9,4))
c0 = ax.contourf(X[:, :, zg] * 1e3, Y[:, :, zg] * 1e3, famp[:, :, zg] * centi, 12, cmap='viridis')
cbar = fig.colorbar(c0, label='electric field (V / cm)')
ax.set_aspect('equal')
ax.set_xlabel('x (mm)')
plt.show()
# ## field gradient
# +
gx, gy, gz = fa.grad_field(voltages)
fig, ax = plt.subplots(figsize=(6, 6))
ax.quiver(X[subset] * 1e3, Y[subset] * 1e3, gx[subset], gy[subset], angles='xy')
ax.set_aspect('equal')
ax.set_xlabel('x (mm)')
ax.set_ylabel('y (mm)')
plt.show()
# -
# ## point calculations
# +
xvals = np.arange(-10, 10, .1) * 1e-3
fvals = [fa.amp_field_r((x, 0, 0.075), voltages) * centi for x in xvals]
fig, ax = plt.subplots(figsize=(8,4))
ax.plot(xvals / milli, fvals)
ax.set_ylabel('electric field (V / cm)')
ax.set_xlabel('x (mm)')
plt.show()
# +
gx, gy, gz = np.array([np.array(fa.grad_field_r((x, 0, 0.075), voltages)) * centi**2.0 for x in xvals]).T
fig, ax = plt.subplots(figsize=(8,4))
ax.plot(xvals / milli, gx)
ax.plot(xvals / milli, gy)
ax.plot(xvals / milli, gz)
ax.set_ylabel('grad |F| (V / cm^2)')
ax.set_xlabel('x (mm)')
plt.show()
# +
yvals = np.arange(-8, 8, .1) * 1e-3
fvals = [fa.amp_field_r((0, y, 0.075), voltages) * centi for y in yvals]
fig, ax = plt.subplots(figsize=(8,4))
ax.plot(yvals / milli, fvals)
ax.set_ylabel('electric field (V / cm)')
ax.set_xlabel('y (mm)')
plt.show()
# +
gx, gy, gz = np.array([np.array(fa.grad_field_r((0, y, 0.075), voltages)) * centi**2.0 for y in yvals]).T
fig, ax = plt.subplots(figsize=(8,4))
ax.plot(yvals / milli, gx)
ax.plot(yvals / milli, gy)
ax.plot(yvals / milli, gz)
ax.set_ylabel('grad |F| (V / cm^2)')
ax.set_xlabel('y (mm)')
plt.show()
# -
| notebooks/fastadjust.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/LeoVogiatzis/medical_data_analysis/blob/main/Eda.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="M4U1ulXRG4DD" outputId="fd5dc702-ae9b-4e63-8918-c5d915049457"
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# !pip install pandas-profiling
from pandas_profiling import ProfileReport
import seaborn as sns
# %matplotlib notebook
# %matplotlib inline
# + colab={"base_uri": "https://localhost:8080/", "height": 391} id="ZVQrgCFjH8KW" outputId="6a660db4-ae40-4e49-fd79-7f9169a0f96e"
data = pd.read_csv('/content/Thyroid_Sick.csv')
data.head(10)
# + colab={"base_uri": "https://localhost:8080/", "height": 297} id="D9-DxRtbIuxE" outputId="e644d184-5c51-47c8-cdc1-27de96ba6223"
data.describe()
# + id="8poRGb8BJ2kw"
#data.describe(include=[object])
# data.describe(include=[data])
data['referral source'].unique()
data.drop(columns=['referral source'], inplace=True)
# + colab={"base_uri": "https://localhost:8080/", "height": 473} id="oN5acFd0Zi4C" outputId="29807e80-9481-4c8c-a04a-9e0855866c07"
data.loc[:, ~data.columns.str.endswith("measured")]
# + id="yjduBNJnm0UZ"
data.replace(['f', 't'], [0, 1], inplace=True)
data.replace(['F', 'M'], [0, 1], inplace=True)
data.replace(['negative', 'sick'], [0, 1], inplace=True)
# + colab={"base_uri": "https://localhost:8080/", "height": 473} id="T8aAiV6Hm7MM" outputId="50bbc521-3a3e-4489-9bdd-e6f4e34e9d06"
data
# + id="pDc3MeKdo-2S"
target = pd.DataFrame(data['Class'])
data.drop(columns=['Class'], inplace=True)
# + colab={"base_uri": "https://localhost:8080/", "height": 425} id="BY-eKTVXfIyP" outputId="f900bea2-38f1-4d20-b08b-45f037b37e79"
from sklearn.impute import KNNImputer
# nan = np.nan
imputer = KNNImputer(n_neighbors=5, weights="uniform")
data = pd.DataFrame(imputer.fit_transform(data), columns=data.columns)
# + id="TMJkFSkafR7N"
# + colab={"base_uri": "https://localhost:8080/", "height": 374} id="9u7gF5WBI0_N" outputId="88818567-7378-40fd-fc9e-ca11d89e6d13"
# # !pip install sweetviz
import sweetviz as sv
#EDA using Autoviz
sweet_report = sv.analyze(data)
#Saving results to HTML file
sweet_report.show_html('sweet_report.html')
# + id="08et_f1frrVp"
corr_df = data.copy()
corr_df.drop(columns=['TBG'], axis=True, inplace=True)
plt.figure(figsize=(8,6))
sns.heatmap(corr_df.corr(),cmap=plt.cm.Reds,annot=True)
plt.title('Heatmap displaying the relationship between\nthe features of the data',
fontsize=13)
plt.show()
# + id="41dp18o1x95D"
corr_df.hist(figsize=(12,6),bins=100)
plt.show()
# + id="_Qb-G1A6yZ_C"
f, axes = plt.subplots(1, 7, figsize=(15, 5), sharex=False,sharey=True)
sns.countplot(x='sick',data=data,ax=axes[0],palette="Set2")
sns.countplot(x='sex',data=data,ax=axes[1],palette="Set2")
sns.countplot(x='pregnant',data=data,ax=axes[2],palette="Set2")
sns.countplot(x='lithium',data=data,ax=axes[3],palette="Set2")
sns.countplot(x='tumor',data=data,ax=axes[4],palette="Set2")
sns.countplot(x='Class',data=data,ax=axes[5],palette="Set2")
sns.countplot(x='psych',data=data,ax=axes[6],palette="Set2")
plt.title('Categorical Bar charts', fontsize=20, loc='left')
data.columns
# + id="jP4in85m9tdG"
#data.dropna(inplace=True)
data.isna
data.isnull
# numerics = ['int16', 'int32', 'int64', 'float16', 'float32', 'float64']
# newdf = data.select_dtypes(include=numerics)
#sns.pairplot(data.drop("TBG", axis=1), hue="Class", size=3)
#data
# example of a ordinal encoding
# + id="Y_HoaHCeHeWt"
# + id="GUxWY4nz4pLw"
lk# sns.set()
cols = ['age', 'sex', 'query on thyroxine','on antithyroid medication', 'pregnant','Class']
sns.pairplot(data[cols], height = 2.5)
plt.show()
| notebooks/Eda.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import findspark
findspark.init()
from pyspark.sql import SparkSession
from pyspark.sql.types import *
spark = SparkSession.builder.appName('Stream File').getOrCreate()
spark.sparkContext.setLogLevel('ERROR')
mySchema = StructType([
StructField('Date', TimestampType(),True),
StructField('Message', StringType(),True)
])
streamDF = spark.readStream.option('delimiter','|').schema(mySchema).csv(path='logs/')
streamDF.createOrReplaceTempView('SDF')
outputDF = spark.sql(sqlQuery='select * from SDF')
outputDF.writeStream.format(source='console').outputMode(outputMode='update').start().awaitTermination() # can be append on complete
| sparkStreaming.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + active=""
# tune cascade classifier:
# detectMultiscale(img,scale factor, min ngbr)
# HSV colorspace
# +
# part 1
# import libraries
import cv2
import numpy as np
# parameters for shi-tomasi corner detection
st_params = dict(maxCorners=30,
qualityLevel=0.2,
minDistance=2,
blockSize=7)
# parameters for Lucas-Kande optical flow
lk_params = dict(winSize= (15,15),
maxLevel=2,
criteria=(cv2.TERM_CRITERIA_EPS | cv2.TERM_CRITERIA_COUNT , 10, 1))
# Video Capture
cap = cv2.VideoCapture(0)
# color for optical flow
# part 2
color= (0,255,0)
# read the capture and get the first frame
_,frame_t = cap.read()
# convert to grayscale
prev_gray=cv2.cvtColor(frame_t,cv2.COLOR_BGR2GRAY)
# find the strongest corner in the first frame
prev = cv2.goodFeaturesToTrack(prev_gray,mask=None,**st_params)
# create an image with same the dimensions for the later drawing purposes
mask = np.zeros_like(frame_t)
# while loop
while (cap.isOpened()):
_,frame=cap.read()
frame = cv2.flip(frame,1)
gray=cv2.cvtColor(frame,cv2.COLOR_BGR2GRAY)
# callulate optical flow by lucas-kanade
next,status,error = cv2.calcOpticalFlowPyrLK(prev_gray,gray,prev, None, **lk_params)
# select a good feature for previous position
good_old = prev[status ==1]
# select a good feature for next position
good_new = next[status ==1]
# draw the optical flow track
for i ,(new,old) in enumerate (zip(good_new,good_old)):
# return coordinates for a new point & old point
a,b = new.ravel()
c,d = old.ravel()
# draw a line between new and old points.. which is green
mask =cv2.line(mask,(a,b),(c,d),color,2)
# draw filled circle
frame = cv2.circle(frame,(a,b),3,color,-1)
# overlay optical on origional frame
output = cv2.add(frame,mask)
# update previous pad was acting as a toggle for insert mode.frame
prev_gray = gray.copy()
# update previous good feature points
prev = good_new.reshape(-1,1,2)
# open new window and display output
cv2.imshow("Optical Flow",output)
# close the frame
if cv2.waitKey(300) & 0xFF == ord("q"):
break
# release and destroy
cap.release()
cv2.destroyAllWindows()
# -
| optical flow.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pyqtgraph as pg
import sys
from PyQt5.QtWidgets import QWidget,QApplication,QFrame,QGridLayout,QVBoxLayout
class Example(QWidget):
def __init__(self):
super(Example, self).__init__()
self.initUI()
self.generate_image()
def initUI(self):
self.setGeometry(200,200,1000,800)
self.setWindowTitle("实时刷新正余弦波形图")
self.gridLayout = QGridLayout(self)
self.frame = QFrame(self)
self.frame.setFrameShape(QFrame.Panel)
self.frame.setFrameShadow(QFrame.Plain)
self.frame.setLineWidth(2)
self.frame.setStyleSheet("background-color:rgb(0,255,255);")
self.gridLayout.addWidget(self.frame,0,0,1,2)
self.setLayout(self.gridLayout)
def generate_image(self):
verticalLayout = QVBoxLayout(self.frame)
win = pg.GraphicsLayoutWidget(self.frame)
verticalLayout.addWidget(win)
p = win.addPlot(title="动态波形图")
p.showGrid(x=True,y=True)
p.setLabel(axis="left",text="Amplitude / V")
p.setLabel(axis="bottom",text="t / s")
p.setTitle("y = sin(x)")
p.addLegend()
# mkpen('y', width=3, style=QtCore.Qt.DashLine)
self.curve1 = p.plot(pen=pg.mkPen("r",width=2),name="y1") #设置pen 格式
self.Fs = 1024.0 #采样频率
self.N = 1024 #采样点数
self.f0 = 4.0 #信号频率
self.pha = 0 #初始相位
self.t = np.arange(self.N) / self.Fs #时间向量 1*1024的矩阵
self.curve1.setData(self.t, np.sin(8 * np.pi * self.t + self.pha * np.pi / 180.0))
if __name__ == '__main__':
app = QApplication(sys.argv)
ex = Example()
ex.show()
sys.exit(app.exec_())
# -
| ref/test_graph.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Unit 7 - Distinguishing Sentiments
# The first plot will be and/or feature the following:
#
# * Be a scatter plot of sentiments of the last 100 tweets sent out by each news organization, ranging from -1.0 to 1.0, where a score of 0 expresses a neutral sentiment, -1 the most negative sentiment possible, and +1 the most positive sentiment possible.
# * Each plot point will reflect the compound sentiment of a tweet.
# * Sort each plot point by its relative timestamp.
# * The second plot will be a bar plot visualizing the overall sentiments of the last 100 tweets from each organization. For this plot, you will again aggregate the compound sentiments analyzed by VADER.
#
# The tools of the trade you will need for your task as a data analyst include the following: tweepy, pandas, matplotlib, and VADER.
# Your final Jupyter notebook must:
#
# * Pull last 100 tweets from each outlet.
# * Perform a sentiment analysis with the compound, positive, neutral, and negative scoring for each tweet.
# * Pull into a DataFrame the tweet's source acount, its text, its date, and its compound, positive, neutral, and negative sentiment scores.
# * Export the data in the DataFrame into a CSV file.
# * Save PNG images for each plot.
#
# As final considerations:
#
# * Use the Matplotlib libraries.
# * Include a written description of three observable trends based on the data.
# * Include proper labeling of your plots, including plot titles (with date of analysis) and axes labels.
# * Include an exported markdown version of your Notebook called README.md in your GitHub repository.
# ## 3 Overall Observations
# * The sentiment analysis for each media outlet fluctuates frequently making it difficult to truly understand the overall sentiment.
# * Generally speaking, most outlets are biased in what is tweeted out. For example, the positive tweets from FOX News are very right leaning whereas the tweets with a negative sentiment are focused on left leaning politics.
# * Additionally, FOX News tends to tweet out more negatively focused messaging.
# +
# Import dependencies
import tweepy
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
from vaderSentiment.vaderSentiment import SentimentIntensityAnalyzer
analyzer = SentimentIntensityAnalyzer()
from datetime import datetime
from config import (consumer_key, consumer_secret, access_token, access_token_secret)
# -
# Setup Tweepy API authentication
auth = tweepy.OAuthHandler(consumer_key, consumer_secret)
auth.set_access_token(access_token, access_token_secret)
api = tweepy.API(auth, parser=tweepy.parsers.JSONParser())
# Target Twitter Accounts
target_terms = ("@BBC", "@CBS", "@CNN", "@FOXNEWS", "@NYTIMES")
# Filter out non-human activity
min_tweets = 5
max_tweets = 10000
max_followers = 2500
max_following = 2500
lang = "en"
# +
# Create array to hold sentiment
sentiments = []
counter = 1
# Loop through target accounts
for target in target_terms:
oldest_tweet = None
compound_list = []
positive_list = []
neutral_list = []
negative_list = []
# Loop through range to obtain the last 100 tweets by each account
for loop in range(4):
public_tweets = api.search(target, count=100, result_type="recent", max_id=oldest_tweet)
for tweet in public_tweets["statuses"]:
# Utilize the non-human filters
if (tweet["user"]["followers_count"] < max_followers and
tweet["user"]["statuses_count"] > min_tweets and
tweet["user"]["statuses_count"] < max_tweets and
tweet["user"]["friends_count"] < max_following and
tweet["user"]["lang"] == lang):
#Run VADER Analysis on each tweet
results = analyzer.polarity_scores(tweet["text"])
compound = results["compound"]
pos = results["pos"]
neu = results["neu"]
neg = results["neg"]
tweets_ago = counter
# Push to each array
compound_list.append(compound)
positive_list.append(pos)
neutral_list.append(neu)
negative_list.append(neg)
# Set oldest tweet value
oldest_tweet = int(tweet["id_str"]) - 1
# Create a sentiments dictionary
sentiments.append({"User": target,
"Tweet": tweet["text"],
"Date": tweet["created_at"],
"Compound": compound,
"Positive": pos,
"Neutral": neu,
"Negative": neg,
"Tweets Ago": counter})
# Add to counter
counter = counter+1
# -
# Convert sentiments to dataframe
sentiments_pd = pd.DataFrame.from_dict(sentiments)
sentiments_pd.head()
# Split the sentiments_pd dataframe into smaller dataframes for each news outlet
bbc_df = sentiments_pd.loc[sentiments_pd['User']=='@BBC', :].reset_index(drop=True).head(100)
cbs_df = sentiments_pd.loc[sentiments_pd['User']=='@CBS', :].reset_index(drop=True).head(100)
cnn_df = sentiments_pd.loc[sentiments_pd['User']=='@CNN', :].reset_index(drop=True) .head(100)
fox_df = sentiments_pd.loc[sentiments_pd['User']=='@FOXNEWS', :].reset_index(drop=True).head(100)
nyt_df = sentiments_pd.loc[sentiments_pd['User']=='@NYTIMES', :].reset_index(drop=True).head(100)
# +
# Create scatter plot for BBC sentiment analysis
plt.plot(np.arange(len(bbc_df["Compound"])),
bbc_df["Compound"], marker="o", linewidth=0.5,
alpha=0.65, color='b', linestyle=':')
plt.title("Sentiment Analysis of BBC Tweets (03/16/2018)")
plt.ylabel("Tweet Polarity")
plt.xlabel("Tweets Ago")
plt.grid(linestyle='dotted')
plt.savefig("BBC_Sentiment.png")
plt.show()
# +
# Create scatter plot for CBS sentiment analysis
plt.plot(np.arange(len(cbs_df["Compound"])),
cbs_df["Compound"], marker="o", linewidth=0.5,
alpha=0.65, color='r', linestyle=':')
plt.title("Sentiment Analysis of CBS Tweets (03/16/2018)")
plt.ylabel("Tweet Polarity")
plt.xlabel("Tweets Ago")
plt.grid(linestyle='dotted')
plt.savefig("CBS_Sentiment.png")
plt.show()
# +
# Create scatter plot for CNN sentiment analysis
plt.plot(np.arange(len(cnn_df["Compound"])),
cnn_df["Compound"], marker="o", linewidth=0.5,
alpha=0.65, color='g', linestyle=':')
plt.title("Sentiment Analysis of CNN Tweets (03/16/2018)")
plt.ylabel("Tweet Polarity")
plt.xlabel("Tweets Ago")
plt.grid(linestyle='dotted')
plt.savefig("CNN_Sentiment.png")
plt.show()
# +
# Create scatter plot for Fox sentiment analysis
plt.plot(np.arange(len(fox_df["Compound"])),
fox_df["Compound"], marker="o", linewidth=0.5,
alpha=0.65, color='y', linestyle=':')
plt.title("Sentiment Analysis of Fox News Tweets (03/16/2018)")
plt.ylabel("Tweet Polarity")
plt.xlabel("Tweets Ago")
plt.grid(linestyle='dotted')
plt.savefig("FOX_Sentiment.png")
plt.show()
# +
# Create scatter plot for NYT sentiment analysis
plt.plot(np.arange(len(nyt_df["Compound"])),
cnn_df["Compound"], marker="o", linewidth=0.5,
alpha=0.65, color='orange', linestyle=':')
plt.title("Sentiment Analysis of New York Times Tweets (03/16/2018)")
plt.ylabel("Tweet Polarity")
plt.xlabel("Tweets Ago")
plt.grid(linestyle='dotted')
plt.savefig("NYT_Sentiment.png")
plt.show()
# +
# Combo scatter plot
bbc_plt = plt.plot(np.arange(len(bbc_df["Compound"])),bbc_df["Compound"], marker="o", linewidth=0.5,alpha=0.65, color='b', linestyle=':')
cbs_plt = plt.plot(np.arange(len(cbs_df["Compound"])),cbs_df["Compound"], marker="o", linewidth=0.5,alpha=0.65, color='r', linestyle=':')
cnn_plt = plt.plot(np.arange(len(cnn_df["Compound"])),cnn_df["Compound"], marker="o", linewidth=0.5,alpha=0.65, color='g', linestyle=':')
fox_plt = plt.plot(np.arange(len(fox_df["Compound"])),fox_df["Compound"], marker="o", linewidth=0.5,alpha=0.65, color='y', linestyle=':')
nyt_plt = plt.plot(np.arange(len(nyt_df["Compound"])),cnn_df["Compound"], marker="o", linewidth=0.5,alpha=0.65, color='orange', linestyle=':')
plt.grid(linestyle='dotted')
plt.title("Sentiment Analysis of Media Tweets (03/16/2018)")
plt.ylabel("Tweet Polarity")
plt.xlabel("Tweets Ago")
plt.legend(('BBC', 'CBS', 'CNN', 'FOX', 'NYT'),scatterpoints=1,loc='upper left',bbox_to_anchor=(1.0, 1.035),ncol=1,\
fontsize=8, markerscale=0.75,title='Media Outlets',edgecolor='none',framealpha=1.00)
plt.savefig("Media_Aggregate_Analysis.png")
plt.show()
# -
# aggregate overall media sentiment for each news outlet
bbc_agg = bbc_df['Compound'].mean()
cbs_agg = cbs_df['Compound'].mean()
cnn_agg = cnn_df['Compound'].mean()
fox_agg = fox_df['Compound'].mean()
nyt_agg = nyt_df['Compound'].mean()
# +
# Combo bar plot
media_agg = ['BBC', 'CBS', 'CNN', 'FOX', 'NYT']
sentiment = [-0.033, 0.090, -0.095, -0.199, 0.123]
x_axis = np.arange(len(sentiment))
colors = ['r', 'b', 'g', 'y', 'orange']
plt.bar(x_axis, sentiment, color=colors, alpha=0.65, align='edge')
tick_locations = [value+0.4 for value in x_axis]
plt.xticks(tick_locations, ["BBC", "CBS", "CNN", "FOX", "NYT"])
plt.grid(linestyle='dotted')
plt.title("Overall Media Sentiment (03/16/2018)")
plt.xlabel("Media")
plt.ylabel("Twitter Polarity")
plt.savefig("Media_Mean_Sentiment.png")
plt.show()
# -
| SocialPy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from pathlib import Path
from tqdm.auto import tqdm
import torch
import torch.nn as nn
import torchvision
from dataloading.nvidia import NvidiaDataset, NvidiaSpringTrainDataset, NvidiaTrainDataset, NvidiaValidationDataset
from network import PilotNet
from metrics.metrics import calculate_closed_loop_metrics, calculate_lateral_errors, calculate_interventions
# %load_ext autoreload
# %autoreload 2
# -
def calculate_metrics(driving_ds, expert_ds):
metrics_df = pd.DataFrame(columns=["model", "mae", "rmse", "max", "failure_rate",
"interventions", "whiteness", "expert_whiteness"])
for name, ds in driving_ds.items():
print(f"Calculating metrics for {name}")
metrics = calculate_closed_loop_metrics(ds.frames, expert_ds.frames)
metrics['model'] = name
metrics_df = metrics_df.append(metrics, ignore_index=True)
return metrics_df
def draw_error_plot_ax(ax, model_ds, expert_ds, title=None):
lat_errors = calculate_lateral_errors(model_ds.frames[:-1000], expert_ds.frames, only_autonomous=True)
positions_df = model_ds.frames[:-1000]
autonomous_df = positions_df[positions_df.autonomous].reset_index(drop=True)
ax.scatter(autonomous_df["position_x"], autonomous_df["position_y"],
s=5,
c=lat_errors, cmap=plt.cm.coolwarm)
manual_df = positions_df[positions_df.autonomous == False].reset_index(drop=True)
ax.scatter(manual_df["position_x"], manual_df["position_y"],
s=5,
c="#2BFA00")
positions_df['autonomous_next'] = positions_df.shift(-1)['autonomous']
interventions_df = positions_df[positions_df.autonomous & (positions_df.autonomous_next == False)]
ax.scatter(interventions_df["position_x"], interventions_df["position_y"],
s=150,
marker='x',
c="red")
if title:
ax.set_title(title)
def draw_error_plot(model_ds, expert_ds, title=None):
fig, ax = plt.subplots(1, 1, figsize=(10, 10))
draw_error_plot_ax(ax, model_ds, expert_ds, title)
root_path = Path("/media/romet/data2/datasets/rally-estonia/dataset")
expert_ds = NvidiaDataset([root_path / '2021-10-26-10-49-06_e2e_rec_ss20_elva'])
expert_back_ds = NvidiaDataset([root_path / '2021-10-26-11-08-59_e2e_rec_ss20_elva_back'])
autumn3_ds = NvidiaDataset([root_path / '2021-11-25-12-09-43_e2e_rec_elva-nvidia-v1-0.8'])
print(calculate_closed_loop_metrics(autumn3_ds.frames, expert_ds.frames))
draw_error_plot(autumn3_ds, expert_ds, "2021-11-25-12-09-43_e2e_rec_elva-nvidia-v1-0.8")
nvidia_1_backward = NvidiaDataset([root_path / '2021-11-25-12-09-43_e2e_rec_elva-nvidia-v1-0.8']).frames
backward_metrics = calculate_closed_loop_metrics(nvidia_1_backward, expert_ds.frames)
backward_metrics['interventions'] = backward_metrics['interventions'] - 1
backward_metrics
nvidia_1_forward = NvidiaDataset([root_path / '2021-11-25-12-21-17_e2e_rec_elva-nvidia-v1-0.8-forward']).frames[:-1000]
forward_metrics = calculate_closed_loop_metrics(nvidia_1_forward, expert_ds.frames)
forward_metrics['interventions'] = forward_metrics['interventions'] - 1
forward_metrics
nvidia_1 = pd.concat([nvidia_1_forward, nvidia_1_backward])
total_metrics = calculate_closed_loop_metrics(nvidia_1, expert_ds.frames)
total_metrics['distance'] = forward_metrics['distance'] + backward_metrics['distance']
total_metrics['interventions'] = forward_metrics['interventions'] + backward_metrics['interventions']
total_metrics['distance_per_intervention'] = total_metrics['distance'] / total_metrics['interventions']
total_metrics
print(total_metrics)
x1 = frames['position_x']
y1 = frames['position_y']
x2 = frames.shift(-1)['position_x']
y2 = frames.shift(-1)['position_y']
frames['distance'] = np.sqrt((x2-x1)**2 + (y2-y1)**2)
frames[['position_x', 'position_y', 'distance']]
autumn3_ds = NvidiaDataset([root_path / '2021-11-25-12-37-01_e2e_rec_elva-lidar-v1-0.8-back'])
print(calculate_closed_loop_metrics(autumn3_ds.frames, expert_ds.frames))
draw_error_plot(autumn3_ds, expert_ds, "2021-11-25-12-37-01_e2e_rec_elva-lidar-v1-0.8-back")
autumn3_ds = NvidiaDataset([root_path / '2021-11-25-12-45-35_e2e_rec_elva-lidar-v1-0.8-back'])
print(calculate_closed_loop_metrics(autumn3_ds.frames, expert_ds.frames))
draw_error_plot(autumn3_ds, expert_ds, "2021-11-25-12-45-35_e2e_rec_elva-lidar-v1-0.8-back")
frames = autumn3_ds.frames
frames['autonomous_next'] = frames.shift(-1)['autonomous']
interventions = frames[frames.autonomous & (frames.autonomous_next == False)]
autumn3_ds = NvidiaDataset([root_path / '2021-11-25-12-21-17_e2e_rec_elva-nvidia-v1-0.8-forward'])
print(calculate_closed_loop_metrics(autumn3_ds.frames, expert_ds.frames))
draw_error_plot(autumn3_ds, expert_ds, "nvidia-v1-0.8-forward")
datasets = {
'autumn-v3': NvidiaDataset([root_path / '2021-11-03-12-35-19_e2e_rec_elva_autumn-v3']),
'wide-v2': NvidiaDataset([root_path / '2021-11-03-13-13-16_e2e_rec_elva_wide-v2']),
'autumn-v1': NvidiaDataset([root_path / '2021-11-03-13-51-53_e2e_rec_elva_autumn-v1',
root_path / '2021-11-03-14-02-07_e2e_rec_elva_autumn-v1_continue']),
'autumn-v3-overfit': NvidiaDataset([root_path / '2021-11-03-14-31-14_e2e_rec_elva_autumn-v3-last']),
'autumn-v3-drive-2': NvidiaDataset([root_path / '2021-11-03-15-07-12_e2e_rec_elva_autumn-v3-drive2'])
}
metrics = calculate_metrics(datasets, expert_ds)
metrics
# +
# intervention_adjustments = {
# 'autumn-v3': 0,
# 'wide-v2': 0,
# 'autumn-v1': 0,
# 'autumn-v3-overfit': 0,
# 'autumn-v3-drive-2': 0,
# }
# for name, adj in intervention_adjustments.items():
# metrics[metrics.model == name] = metrics[metrics.model == name].interventions - adj
# -
datasets_backwards = {
'autumn-v3': NvidiaDataset([root_path / '2021-11-03-12-53-38_e2e_rec_elva_back_autumn-v3']),
'wide-v2': NvidiaDataset([root_path / '2021-11-03-13-30-48_e2e_rec_elva_back_wide-v2']),
'autumn-v1': NvidiaDataset([root_path / '2021-11-03-14-12-10_e2e_rec_elva_back_autumn-v1']),
'autumn-v3-overfit': NvidiaDataset([root_path / '2021-11-03-14-48-21_e2e_rec_elva_back_autumn-v3-last']),
'autumn-v3-drive-2': NvidiaDataset([root_path / '2021-11-03-15-25-36_e2e_rec_elva_back_autumn-v3-drive2'])
}
metrics_backwards = calculate_metrics(datasets_backwards, expert_back_ds)
metrics_backwards
# +
fig, ax = plt.subplots(5, 2, figsize=(15, 30))
for i, (name, ds) in enumerate(datasets.items()):
print(i, name)
draw_error_plot_ax(ax[i][0], ds, expert_ds, f"{name} elva forward")
for i, (name, ds) in enumerate(datasets_backwards.items()):
print(i, name)
draw_error_plot_ax(ax[i][1], ds, expert_back_ds, f"{name} Elva backward")
# -
fig.savefig("elva-2021-11-03.png", facecolor="white")
| metrics/closed_loop.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
# data manipulation
import numpy as np
import pandas as pd
#file processing / requests
from pathlib import Path
from tqdm import tqdm
import json
import urllib
#computer vision
import cv2
import torch
import torchvision
import PIL
#ML
from sklearn.model_selection import train_test_split
#display
from IPython.display import display
from pylab import rcParams
#visualization
import seaborn as sns
import matplotlib.pyplot as plt
from matplotlib import rc
import PIL.Image as Image
#utils2.py package can be found under https://github.com/philippe-heitzmann/Object_Detection
import sys
sys.path.append('/Users/Administrator/DS/DeepRoad/Object_Detection')
sys.path.append('/Users/phil0/DS/DeepRoad/Object_Detection')
import utils2
from importlib import reload
reload(utils2)
from utils2 import *
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:100% !important; }</style>"))
pd.set_option('display.max_columns', None)
pd.set_option('display.float_format', lambda x: '%.2f' % x)
# -
# !pip freeze
# +
# from https://github.com/CSAILVision/places365/blob/master/run_placesCNN_basic.py
# PlacesCNN for scene classification
#
# by <NAME>
# last modified by <NAME>, Dec.27, 2017 with latest pytorch and torchvision (upgrade your torchvision please if there is trn.Resize error)
import torch
from torch.autograd import Variable as V
import torchvision.models as models
from torchvision import transforms as trn
from torch.nn import functional as F
import os
from PIL import Image
# the architecture to use
arch = 'resnet18'
# load the pre-trained weights
model_file = '%s_places365.pth.tar' % arch
if not os.access(model_file, os.W_OK):
weight_url = 'http://places2.csail.mit.edu/models_places365/' + model_file
os.system('wget ' + weight_url)
# +
model = models.__dict__[arch](num_classes=365)
checkpoint = torch.load(model_file, map_location=lambda storage, loc: storage)
state_dict = {str.replace(k,'module.',''): v for k,v in checkpoint['state_dict'].items()}
print(state_dict)
model.load_state_dict(state_dict)
model.eval()
# -
model.eval()
# +
# load the image transformer
#trn = torchvision transforms
centre_crop = trn.Compose([
trn.Resize((256,256)),
trn.CenterCrop(224),
trn.ToTensor(),
trn.Normalize([0.485, 0.456, 0.406], [0.229, 0.224, 0.225])
])
# load the class label
file_name = 'categories_places365.txt'
if not os.access(file_name, os.W_OK):
synset_url = 'https://raw.githubusercontent.com/csailvision/places365/master/categories_places365.txt'
os.system('wget ' + synset_url)
classes = list()
with open(file_name) as class_file:
for line in class_file:
classes.append(line.strip().split(' ')[0][3:])
classes = tuple(classes)
print(classes)
# -
# load the test image
img_name = '12.jpg'
if not os.access(img_name, os.W_OK):
print('Fetching image')
img_url = 'http://places.csail.mit.edu/demo/' + img_name
os.system('wget ' + img_url)
img = Image.open(img_name)
input_img = V(centre_crop(img).unsqueeze(0))
input_img
print(input_img.size())
# +
img = Image.open(img_name)
input_img = V(centre_crop(img).unsqueeze(0))
# forward pass
logit = model.forward(input_img)
h_x = F.softmax(logit, 1).data.squeeze()
probs, idx = h_x.sort(0, True)
print('{} prediction on {}'.format(arch,img_name))
# output the prediction
for i in range(0, 5):
print('{:.3f} -> {}'.format(probs[i], classes[idx[i]]))
# -
# !python run_placesCNN_basic.py
# !python run_placesCNN_basic.py --model_type 'resnet18' --folder '/Users/Administrator/DS/places365/images'
# !python run_placesCNN_basic.py --model_type 'resnet18' --folder '/Users/Administrator/DS/places365/images'
# +
### Randomly sampling n images from folders
# -
# !pip install spacy nltk gensim textblob bokeh imutils pytesseract plotly hdbscan statsmodels
# +
#utils2.py package can be found under https://github.com/philippe-heitzmann/Object_Detection
import sys
sys.path.append('/Users/Administrator/DS/DeepRoad/Object_Detection')
sys.path.append('/Users/phil0/DS/DeepRoad/Object_Detection')
import utils2
from importlib import reload
reload(utils2)
from utils2 import *
results1 = get_files_in_dir('jpg', 'png', path = '/Users/Administrator/DS/places365/images', fullpath = True)
results1 = np.random.choice(results1, size=1, replace=False) #, p=[0.3, 0.5, 0.0, 0.2]
print(results1)
# +
allfiles = getListOfFiles('jpg', dirName = '/Users/Administrator/DS/places365/images')
print(allfiles, len(allfiles))
# -
| Running_Places365_Scene_Classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/jana0601/Summer_school_archiv-/blob/main/nb4_neuralnetworks_interactive.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="hllkE2OzS_eu"
import torch
import torch.nn as nn
import numpy as np
# + [markdown] id="qANeo4lx2ILD"
# First, we define the Neural Network architecture we use for regression. It is a Feedforward NN with 1 hidden layer and ReLu activation function.
# + id="GK8dBoV_Q1cZ"
class Feedforward(torch.nn.Module):
def __init__(self, input_size, hidden_size,hidden_size2):
super(Feedforward, self).__init__()
self.input_size = input_size
self.hidden_size = hidden_size
self.hidden_size2 = hidden_size2
self.hl1 = torch.nn.Linear(self.input_size, self.hidden_size)
self.hl2 = torch.nn.Linear(self.hidden_size, self.hidden_size2)
self.hl3 = torch.nn.Linear(self.hidden_size2, self.hidden_size2)
self.relu = torch.nn.ReLU()
self.output_layer = torch.nn.Linear(self.hidden_size2, 1)
def forward(self, x):
hidden = self.hl1(x)
relu = self.relu(hidden)
hidden2 = self.hl2(relu)
relu2 = self.relu(hidden2)
hidden3 = self.hl3(relu2)
relu3 = self.relu(hidden3)
output = self.output_layer(relu3)
return output
# + [markdown] id="uMneqgI72MBa"
# Next, we create our artificial data. We create data tuples $(x_i,y_i)$, $i=1,...,n$ where the targets $y_i = f(x_i) + \varepsilon$ are noisy realizations of some function $f$ of $x$. Our goal is to recover $f$ with the Neural Network.
#
# Feel free to use any of the provided functions or to define your own.
# + id="EcCkqnJpTF_X"
def create_toy_data(func, n=1000, std=.1, domain=[0., 1.]):
# x = torch.linspace(domain[0], domain[1], n)
x = (domain[1]-domain[0]) * torch.rand((n,)) + domain[0]
t = func(x) + torch.normal(mean=torch.zeros_like(x),std=std*torch.ones_like(x))
return x, t
def sinusoidal(x):
return torch.sin(2 * np.pi * x)
def square(x):
return torch.square(x)
def abs(x):
return torch.abs(x)
def heavyside(x):
return 0.5 * (torch.sign(x) + 1)
def exp(x):
return torch.exp(x)
function = sinusoidal
domain = [-1., 1.]
n = 50
# Define data for training
data, target = create_toy_data(function, n=n, std=.1, domain=domain)
data = data[:,None]
# Define separate data set for testing the model once training is finished
test_data, test_target = create_toy_data(function, n=n, std=.0, domain=domain)
test_data = test_data[:,None]
# + [markdown] id="yxje5i3r5ctY"
# With real data, $f$ is of course unknown and usualy not a "nice" mathematical function like the sine. These examples serve as a showcase for the suitability as well as the limitations of simple Neural Networks.
# + [markdown] id="TNkc5pE-2dLN"
# We initialize the model, the loss to minimize, and the optimizer (we use stochastic gradient descend).
#
# The mean square error (MSE) loss we minimize is given by
# $$E(\mathbf{w}) = \frac{1}{n}\sum_{i=1}^{n} |y_i - NN(\mathbf{w})(x_i)|^2,$$
# where $NN(\mathbf{w})$ is the forward pass of the Neural Network with parameters $\mathbf{w}$.
# + id="7il2wHzfTJf_"
model = Feedforward(1,2000,300)
criterion = torch.nn.MSELoss()
optimizer = torch.optim.SGD(model.parameters(), lr = 0.01)
# + [markdown] id="5lVUzyvl9qsG"
# Evaluate the model on the test data set before training
# + colab={"base_uri": "https://localhost:8080/"} id="AHTgMoFqXzTA" outputId="c4770741-98f3-4909-f2d9-dd1302df30d6"
model.eval()
pred = model(test_data)
before_train = criterion(pred.squeeze(), test_target)
print('Test loss before training' , before_train.item())
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="1MuY3tkfaurQ" outputId="164be24c-e5a2-40d1-a4d6-087ad3d5c806"
from matplotlib import pyplot as plt
x = torch.linspace(domain[0],domain[1],n)[:,None]
y = model(x).detach().numpy()
plt.scatter(data, target, facecolor="none", edgecolor="b", color="blue", label="training data")
plt.plot(x, function(x), color="g", label="true function")
plt.plot(x, y, color="r", label="NN prediction")
plt.legend()
plt.show()
# + [markdown] id="E1ezy99T95mF"
# Execute the training loop.
# + id="FXj9qQZvZMJG"
model.train()
epoch = 5000
for epoch in range(epoch):
optimizer.zero_grad()
pred = model(data) # get predictions NN(w)(x_i) of the current model on the data
loss = criterion(pred.squeeze(), target) # get MSE loss sum_i^n |NN(w)(x_i) - y_i|^2/n
# print('Epoch {}: train loss: {}'.format(epoch, loss.item()))
loss.backward() # compute gradients
optimizer.step() # do a gradient descend step
# + [markdown] id="osMr_2ha-01j"
# And evauate the model again after training
# + colab={"base_uri": "https://localhost:8080/"} id="ryPM-AD6qJZM" outputId="d4e9e75e-cdde-42e4-f49b-6a4969cb0521"
model.eval()
pred = model(test_data)
after_train = criterion(pred.squeeze(), test_target)
print('Test loss after training' , after_train.item())
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="jf3I_v5_aCTM" outputId="fbf2dbd3-86e6-441e-8aae-a8bdbdd90467"
from matplotlib import pyplot as plt
x = torch.linspace(domain[0],domain[1],n)[:,None]
y = model(x).detach().numpy()
plt.scatter(data, target, facecolor="none", edgecolor="b", color="blue", label="training data")
plt.plot(x, function(x), color="g", label="true function")
plt.plot(x, y, color="r", label="NN prediction")
plt.legend()
plt.show()
# + [markdown] id="NazG_Hn3ACIX"
# The result is obviously not convincing. Feel free to change the architecture of the NN (maybe the net is too shallow?) and the parameters of the training loop to improve accuracy.
| nb4_neuralnetworks_interactive.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:vgcharm] *
# language: python
# name: conda-env-vgcharm-py
# ---
# + pycharm={"name": "#%%\n"}
import sys
from autograph.play.maze_nn_aut_adv_team_share import init_game
from autograph.lib.util.test_helper_team import doenvsteps, get_and_plot_policies
#config_file = 'autograph/play/config/LiorBrett/seeBothInfs.json'
# use a limited quick MCTS search
config_file = '../../autograph/play/config/guardInf/positiveRewards/seeBothClean500.json'
# Checkpoint path
checkpoint_base_path = '../../checkpoints/local500/%s'
# If you don't want to use a checkpoint, use this
#checkpoint_base_path = ''
run_name = 'seeBothClean_500_positive_loc'
postfix = ''
envs, get_pi_cnn, do_mcts_batch, get_tree_and_play_policies = init_game(config_file, checkpoint_base_path,run_name,postfix, "cpu")
# + pycharm={"name": "#%% Inf_0 breaks wall, is seen by guards\n"}
actionlist = [[0,0,2,0,0,1,0,2,0,0,0,0,1,3,0],
[4,4,4,4,4,4,4,4,4,4,4,4,4,4],
[4,4,4,2,0,0,0,0,0,1,0,2,3],
[1,0,4,4,4,4,4,4,4,4,4,4,4,4]]
stepnum, done, lastplayer, lastrew, lastautstate, finishedrun = doenvsteps(envs, actionlist, do_mcts_batch=do_mcts_batch)
assert(finishedrun)
print("")
envs[0].render()
get_and_plot_policies(envs, do_mcts_batch=do_mcts_batch, get_pi_cnn=get_pi_cnn, get_tree_and_play_policies=get_tree_and_play_policies)
# + pycharm={"name": "#%% Guard_1 moves so they have Inf_2 in sight. System out of sight\n"}
actionlist = [[0],
[0,1,0],
[1,0],
[]]
stepnum, done, lastplayer, lastrew, lastautstate, finishedrun = doenvsteps(envs, actionlist, do_mcts_batch=do_mcts_batch)
assert(finishedrun)
print("")
envs[0].render()
get_and_plot_policies(envs, do_mcts_batch=do_mcts_batch, get_pi_cnn=get_pi_cnn, get_tree_and_play_policies=get_tree_and_play_policies)
# + pycharm={"name": "#%% Inf_2 reveals system to guard team. I thought this would make zero potentially turn\n"}
# Inf_0 saw the system as they entered though, not sure how memory of past moves works.
actionlist = [[],
[],
[2,0,1,0,3],
[]]
stepnum, done, lastplayer, lastrew, lastautstate, finishedrun = doenvsteps(envs, actionlist, do_mcts_batch=do_mcts_batch)
assert(finishedrun)
print("")
envs[0].render()
get_and_plot_policies(envs, do_mcts_batch=do_mcts_batch, get_pi_cnn=get_pi_cnn, get_tree_and_play_policies=get_tree_and_play_policies)
| tests/test_aut_adv_team_share/.ipynb_checkpoints/test_seeBothClean_guard_troubleshoot-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Parameterisation
#
# In this notebook, we show how to find which parameters are needed in a model and define them.
#
# For other notebooks about parameterization, see:
#
# - The API documentation of [Parameters](https://pybamm.readthedocs.io/en/latest/source/parameters/index.html) can be found at [pybamm.readthedocs.io](https://pybamm.readthedocs.io/)
# - [Setting parameter values](https://github.com/pybamm-team/PyBaMM/blob/develop/examples/notebooks/Getting%20Started/Tutorial%204%20-%20Setting%20parameter%20values.ipynb) can be found at `pybamm/examples/notebooks/Getting Started/Tutorial 4 - Setting parameter values.ipynb`. This explains the basics of how to set the parameters of a model (in less detail than here).
# - [parameter-management.ipynb](https://github.com/pybamm-team/PyBaMM/blob/develop/examples/notebooks/parameterization/parameter-management.ipynb) can be found at `pybamm/examples/notebooks/parameterization/parameter-management.ipynb`. This explains how to add and edit parameters in the `pybamm/input` directories
# - [parameter-values.ipynb](https://github.com/pybamm-team/PyBaMM/blob/develop/examples/notebooks/parameterization/parameter-values.ipynb) can be found at `pybamm/examples/notebooks/parameterization/parameter-values.ipynb`. This explains the basics of the `ParameterValues` class.
#
# ## Adding your own parameter sets (using a dictionary)
#
# We will be using the model defined and explained in more detail in [3-negative-particle-problem.ipynb](https://github.com/pybamm-team/PyBaMM/blob/develop/examples/notebooks/Creating%20Models/3-negative-particle-problem.ipynb) example notebook. We begin by importing the required libraries
# %pip install pybamm -q # install PyBaMM if it is not installed
import pybamm
import numpy as np
import matplotlib.pyplot as plt
# ## Setting up the model
#
# We define all the parameters and variables using `pybamm.Parameter` and `pybamm.Variable` respectively.
# +
c = pybamm.Variable("Concentration [mol.m-3]", domain="negative particle")
R = pybamm.Parameter("Particle radius [m]")
D = pybamm.FunctionParameter("Diffusion coefficient [m2.s-1]", {"Concentration [mol.m-3]": c})
j = pybamm.InputParameter("Interfacial current density [A.m-2]")
c0 = pybamm.Parameter("Initial concentration [mol.m-3]")
c_e = pybamm.Parameter("Electrolyte concentration [mol.m-3]")
# -
# Now we define our model equations, boundary and initial conditions. We also add the variables required using the dictionary `model.variables`
# +
model = pybamm.BaseModel()
# governing equations
N = -D * pybamm.grad(c) # flux
dcdt = -pybamm.div(N)
model.rhs = {c: dcdt}
# boundary conditions
lbc = pybamm.Scalar(0)
rbc = -j
model.boundary_conditions = {c: {"left": (lbc, "Neumann"), "right": (rbc, "Neumann")}}
# initial conditions
model.initial_conditions = {c: c0}
model.variables = {
"Concentration [mol.m-3]": c,
"Surface concentration [mol.m-3]": pybamm.surf(c),
"Flux [mol.m-2.s-1]": N,
}
model.length_scales = {"negative particle": pybamm.Scalar(1)}
# -
# We also define the geometry, since there are parameters in the geometry too
r = pybamm.SpatialVariable("r", domain=["negative particle"], coord_sys="spherical polar")
geometry = pybamm.Geometry({"negative particle": {r: {"min": pybamm.Scalar(0), "max": R}}})
# ## Finding the parameters required
# To know what parameters are required by the model and geometry, we can do
model.print_parameter_info()
geometry.print_parameter_info()
# This tells us that we need to provide parameter values for the initial concentration and Faraday constant, an `InputParameter` at solve time for the interfacial current density, and diffusivity as a function of concentration. Since the electrolyte concentration does not appear anywhere in the model, there is no need to provide a value for it.
# ## Adding the parameters
#
# Now we can proceed to the step where we add the `parameter` values using a dictionary. We set up a dictionary with parameter names as the dictionary keys and their respective values as the dictionary values.
# +
def D_fun(c):
return 3.9 #* pybamm.exp(-c)
values = {
"Particle radius [m]": 2,
"Diffusion coefficient [m2.s-1]": D_fun,
"Initial concentration [mol.m-3]": 2.5,
}
# -
# Now we can pass this dictionary in `pybamm.ParameterValues` class which accepts a dictionary of parameter names and values. We can then print `param` to check if it was initialised.
# +
param = pybamm.ParameterValues(values)
param
# -
# ## Updating the parameter values
#
# The parameter values or `param` can be further updated by using the `update` function of `ParameterValues` class. The `update` function takes a dictionary with keys being the parameters to be updated and their respective values being the updated values. Here we update the `"Particle radius [m]"` parameter's value. Additionally, a function can also be passed as a `parameter`'s value which we will see ahead, and a new `parameter` can also be added by passing `check_already_exists=False` in the `update` function.
param.update({"Initial concentration [mol.m-3]": 1.5})
param
# ## Solving the model
# ## Finding the parameters in a model
#
# The `parameter` function of the `BaseModel` class can be used to obtain the parameters of a model.
parameters = model.parameters
parameters
# As explained in the [3-negative-particle-problem.ipynb](https://github.com/pybamm-team/PyBaMM/blob/develop/examples/notebooks/Creating%20Models/3-negative-particle-problem.ipynb) example, we first process both the `model` and the `geometry`.
param.process_model(model)
param.process_geometry(geometry)
# We can now set up our mesh, choose a spatial method, and discretise our model
# +
submesh_types = {"negative particle": pybamm.MeshGenerator(pybamm.Uniform1DSubMesh)}
var_pts = {r: 20}
mesh = pybamm.Mesh(geometry, submesh_types, var_pts)
spatial_methods = {"negative particle": pybamm.FiniteVolume()}
disc = pybamm.Discretisation(mesh, spatial_methods)
disc.process_model(model);
# -
# We choose a solver and times at which we want the solution returned, and solve the model. Here we give a value for the current density `j`.
# +
# solve
solver = pybamm.ScipySolver()
t = np.linspace(0, 3600, 600)
solution = solver.solve(model, t, inputs={"Interfacial current density [A.m-2]": 1.4})
# post-process, so that the solution can be called at any time t or space r
# (using interpolation)
c = solution["Concentration [mol.m-3]"]
c_surf = solution["Surface concentration [mol.m-3]"]
# plot
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(13, 4))
ax1.plot(solution.t, c_surf(solution.t))
ax1.set_xlabel("Time [s]")
ax1.set_ylabel("Surface concentration [mol.m-3]")
rsol = mesh["negative particle"].nodes # radial position
time = 1000 # time in seconds
ax2.plot(rsol * 1e6, c(t=time, r=rsol), label="t={}[s]".format(time))
ax2.set_xlabel("Particle radius [microns]")
ax2.set_ylabel("Concentration [mol.m-3]")
ax2.legend()
plt.tight_layout()
plt.show()
# -
# # Using pre-defined models in `PyBaMM`
#
# In the next few steps, we will be showing the same workflow with the Single Particle Model (`SPM`). We will also see how you can pass a function as a `parameter`'s value and how to plot such `parameter functions`.
#
# We start by initializing our model
spm = pybamm.lithium_ion.SPM()
# ## Finding the parameters in a model
#
# We can print the `parameters` of a model by using the `get_parameters_info` function.
spm.print_parameter_info()
# Note that there are no `InputParameter` objects in the default SPM. Also, note that if a `FunctionParameter` is expected, it is ok to provide a scalar (parameter) instead. However, if a `Parameter` is expected, you cannot provide a function instead.
# Another way to view what parameters are needed is to print the default parameter values. This can also be used to get some good defaults (but care must be taken when combining parameters across datasets and chemistries)
{k: v for k,v in spm.default_parameter_values.items() if k in spm._parameter_info}
# We now define a dictionary of values for `ParameterValues` as before (here, a subset of the `Chen2020` parameters)
# +
def graphite_mcmb2528_diffusivity_Dualfoil1998(sto, T):
D_ref = 3.9 * 10 ** (-14)
E_D_s = 42770
arrhenius = exp(E_D_s / constants.R * (1 / 298.15 - 1 / T))
return D_ref * arrhenius
neg_ocp = np.array([[0. , 1.81772748],
[0.03129623, 1.0828807 ],
[0.03499902, 0.99593794],
[0.0387018 , 0.90023398],
[0.04240458, 0.79649431],
[0.04610736, 0.73354429],
[0.04981015, 0.66664314],
[0.05351292, 0.64137149],
[0.05721568, 0.59813869],
[0.06091845, 0.5670836 ],
[0.06462122, 0.54746181],
[0.06832399, 0.53068399],
[0.07202675, 0.51304734],
[0.07572951, 0.49394092],
[0.07943227, 0.47926274],
[0.08313503, 0.46065259],
[0.08683779, 0.45992726],
[0.09054054, 0.43801501],
[0.09424331, 0.42438665],
[0.09794607, 0.41150269],
[0.10164883, 0.40033659],
[0.10535158, 0.38957134],
[0.10905434, 0.37756538],
[0.1127571 , 0.36292541],
[0.11645985, 0.34357086],
[0.12016261, 0.3406314 ],
[0.12386536, 0.32299468],
[0.12756811, 0.31379458],
[0.13127086, 0.30795386],
[0.13497362, 0.29207319],
[0.13867638, 0.28697687],
[0.14237913, 0.27405477],
[0.14608189, 0.2670497 ],
[0.14978465, 0.25857493],
[0.15348741, 0.25265783],
[0.15719018, 0.24826777],
[0.16089294, 0.2414345 ],
[0.1645957 , 0.23362778],
[0.16829847, 0.22956218],
[0.17200122, 0.22370236],
[0.17570399, 0.22181271],
[0.17940674, 0.22089651],
[0.1831095 , 0.2194268 ],
[0.18681229, 0.21830064],
[0.19051504, 0.21845333],
[0.1942178 , 0.21753715],
[0.19792056, 0.21719357],
[0.20162334, 0.21635373],
[0.2053261 , 0.21667822],
[0.20902886, 0.21738444],
[0.21273164, 0.21469313],
[0.2164344 , 0.21541846],
[0.22013716, 0.21465495],
[0.22383993, 0.2135479 ],
[0.2275427 , 0.21392964],
[0.23124547, 0.21074206],
[0.23494825, 0.20873788],
[0.23865101, 0.20465319],
[0.24235377, 0.20205732],
[0.24605653, 0.19774358],
[0.2497593 , 0.19444147],
[0.25346208, 0.19190285],
[0.25716486, 0.18850531],
[0.26086762, 0.18581399],
[0.26457039, 0.18327537],
[0.26827314, 0.18157659],
[0.2719759 , 0.17814088],
[0.27567867, 0.17529686],
[0.27938144, 0.1719375 ],
[0.28308421, 0.16934161],
[0.28678698, 0.16756649],
[0.29048974, 0.16609676],
[0.29419251, 0.16414985],
[0.29789529, 0.16260378],
[0.30159806, 0.16224113],
[0.30530083, 0.160027 ],
[0.30900361, 0.15827096],
[0.31270637, 0.1588054 ],
[0.31640913, 0.15552238],
[0.32011189, 0.15580869],
[0.32381466, 0.15220118],
[0.32751744, 0.1511132 ],
[0.33122021, 0.14987253],
[0.33492297, 0.14874637],
[0.33862575, 0.14678037],
[0.34232853, 0.14620776],
[0.34603131, 0.14555879],
[0.34973408, 0.14389819],
[0.35343685, 0.14359279],
[0.35713963, 0.14242846],
[0.36084241, 0.14038612],
[0.36454517, 0.13882096],
[0.36824795, 0.13954628],
[0.37195071, 0.13946992],
[0.37565348, 0.13780934],
[0.37935626, 0.13973714],
[0.38305904, 0.13698858],
[0.38676182, 0.13523254],
[0.3904646 , 0.13441178],
[0.39416737, 0.1352898 ],
[0.39787015, 0.13507985],
[0.40157291, 0.13647321],
[0.40527567, 0.13601512],
[0.40897844, 0.13435452],
[0.41268121, 0.1334765 ],
[0.41638398, 0.1348317 ],
[0.42008676, 0.13275118],
[0.42378953, 0.13286571],
[0.4274923 , 0.13263667],
[0.43119506, 0.13456447],
[0.43489784, 0.13471718],
[0.43860061, 0.13395369],
[0.44230338, 0.13448814],
[0.44600615, 0.1334765 ],
[0.44970893, 0.13298023],
[0.45341168, 0.13259849],
[0.45711444, 0.13338107],
[0.46081719, 0.13309476],
[0.46451994, 0.13275118],
[0.46822269, 0.13443087],
[0.47192545, 0.13315202],
[0.47562821, 0.132713 ],
[0.47933098, 0.1330184 ],
[0.48303375, 0.13278936],
[0.48673651, 0.13225491],
[0.49043926, 0.13317111],
[0.49414203, 0.13263667],
[0.49784482, 0.13187316],
[0.50154759, 0.13265574],
[0.50525036, 0.13250305],
[0.50895311, 0.13324745],
[0.51265586, 0.13204496],
[0.51635861, 0.13242669],
[0.52006139, 0.13233127],
[0.52376415, 0.13198769],
[0.52746692, 0.13254122],
[0.53116969, 0.13145325],
[0.53487245, 0.13298023],
[0.53857521, 0.13168229],
[0.54227797, 0.1313578 ],
[0.54598074, 0.13235036],
[0.5496835 , 0.13120511],
[0.55338627, 0.13089971],
[0.55708902, 0.13109058],
[0.56079178, 0.13082336],
[0.56449454, 0.13011713],
[0.5681973 , 0.129869 ],
[0.57190006, 0.12992626],
[0.57560282, 0.12942998],
[0.57930558, 0.12796026],
[0.58300835, 0.12862831],
[0.58671112, 0.12656689],
[0.59041389, 0.12734947],
[0.59411664, 0.12509716],
[0.59781941, 0.12110791],
[0.60152218, 0.11839751],
[0.60522496, 0.11244226],
[0.60892772, 0.11307214],
[0.61263048, 0.1092165 ],
[0.61633325, 0.10683058],
[0.62003603, 0.10433014],
[0.6237388 , 0.10530359],
[0.62744156, 0.10056993],
[0.63114433, 0.09950104],
[0.63484711, 0.09854668],
[0.63854988, 0.09921473],
[0.64225265, 0.09541635],
[0.64595543, 0.09980643],
[0.64965823, 0.0986612 ],
[0.653361 , 0.09560722],
[0.65706377, 0.09755413],
[0.66076656, 0.09612258],
[0.66446934, 0.09430929],
[0.66817212, 0.09661885],
[0.67187489, 0.09366032],
[0.67557767, 0.09522548],
[0.67928044, 0.09535909],
[0.68298322, 0.09316404],
[0.686686 , 0.09450016],
[0.69038878, 0.0930877 ],
[0.69409156, 0.09343126],
[0.69779433, 0.0932404 ],
[0.70149709, 0.09350762],
[0.70519988, 0.09339309],
[0.70890264, 0.09291591],
[0.7126054 , 0.09303043],
[0.71630818, 0.0926296 ],
[0.72001095, 0.0932404 ],
[0.72371371, 0.09261052],
[0.72741648, 0.09249599],
[0.73111925, 0.09240055],
[0.73482204, 0.09253416],
[0.7385248 , 0.09209515],
[0.74222757, 0.09234329],
[0.74593034, 0.09366032],
[0.74963312, 0.09333583],
[0.75333589, 0.09322131],
[0.75703868, 0.09264868],
[0.76074146, 0.09253416],
[0.76444422, 0.09243873],
[0.76814698, 0.09230512],
[0.77184976, 0.09310678],
[0.77555253, 0.09165615],
[0.77925531, 0.09159888],
[0.78295807, 0.09207606],
[0.78666085, 0.09175158],
[0.79036364, 0.09177067],
[0.79406641, 0.09236237],
[0.79776918, 0.09241964],
[0.80147197, 0.09320222],
[0.80517474, 0.09199972],
[0.80887751, 0.09167523],
[0.81258028, 0.09322131],
[0.81628304, 0.09190428],
[0.81998581, 0.09167523],
[0.82368858, 0.09285865],
[0.82739136, 0.09180884],
[0.83109411, 0.09150345],
[0.83479688, 0.09186611],
[0.83849965, 0.0920188 ],
[0.84220242, 0.09320222],
[0.84590519, 0.09131257],
[0.84960797, 0.09117896],
[0.85331075, 0.09133166],
[0.85701353, 0.09089265],
[0.86071631, 0.09058725],
[0.86441907, 0.09051091],
[0.86812186, 0.09033912],
[0.87182464, 0.09041547],
[0.87552742, 0.0911217 ],
[0.87923019, 0.0894611 ],
[0.88293296, 0.08999555],
[0.88663573, 0.08921297],
[0.89033849, 0.08881213],
[0.89404126, 0.08797229],
[0.89774404, 0.08709427],
[0.9014468 , 0.08503284],
[1. , 0.07601531]])
pos_ocp = np.array([[0.24879728, 4.4 ],
[0.26614516, 4.2935653 ],
[0.26886763, 4.2768621 ],
[0.27159011, 4.2647018 ],
[0.27431258, 4.2540312 ],
[0.27703505, 4.2449446 ],
[0.27975753, 4.2364879 ],
[0.28248 , 4.2302647 ],
[0.28520247, 4.2225528 ],
[0.28792495, 4.2182574 ],
[0.29064743, 4.213294 ],
[0.29336992, 4.2090373 ],
[0.29609239, 4.2051239 ],
[0.29881487, 4.2012677 ],
[0.30153735, 4.1981564 ],
[0.30425983, 4.1955218 ],
[0.30698231, 4.1931167 ],
[0.30970478, 4.1889744 ],
[0.31242725, 4.1881533 ],
[0.31514973, 4.1865883 ],
[0.3178722 , 4.1850228 ],
[0.32059466, 4.1832285 ],
[0.32331714, 4.1808805 ],
[0.32603962, 4.1805749 ],
[0.32876209, 4.1789522 ],
[0.33148456, 4.1768146 ],
[0.33420703, 4.1768146 ],
[0.3369295 , 4.1752872 ],
[0.33965197, 4.173111 ],
[0.34237446, 4.1726718 ],
[0.34509694, 4.1710877 ],
[0.34781941, 4.1702285 ],
[0.3505419 , 4.168797 ],
[0.35326438, 4.1669831 ],
[0.35598685, 4.1655135 ],
[0.35870932, 4.1634517 ],
[0.3614318 , 4.1598248 ],
[0.36415428, 4.1571712 ],
[0.36687674, 4.154079 ],
[0.36959921, 4.1504135 ],
[0.37232169, 4.1466532 ],
[0.37504418, 4.1423388 ],
[0.37776665, 4.1382346 ],
[0.38048913, 4.1338248 ],
[0.38321161, 4.1305799 ],
[0.38593408, 4.1272392 ],
[0.38865655, 4.1228104 ],
[0.39137903, 4.1186109 ],
[0.39410151, 4.114182 ],
[0.39682398, 4.1096005 ],
[0.39954645, 4.1046948 ],
[0.40226892, 4.1004758 ],
[0.4049914 , 4.0956464 ],
[0.40771387, 4.0909696 ],
[0.41043634, 4.0864644 ],
[0.41315882, 4.0818448 ],
[0.41588129, 4.077683 ],
[0.41860377, 4.0733309 ],
[0.42132624, 4.0690737 ],
[0.42404872, 4.0647216 ],
[0.4267712 , 4.0608654 ],
[0.42949368, 4.0564747 ],
[0.43221616, 4.0527525 ],
[0.43493864, 4.0492401 ],
[0.43766111, 4.0450211 ],
[0.44038359, 4.041986 ],
[0.44310607, 4.0384736 ],
[0.44582856, 4.035171 ],
[0.44855103, 4.0320406 ],
[0.45127351, 4.0289288 ],
[0.453996 , 4.02597 ],
[0.45671848, 4.0227437 ],
[0.45944095, 4.0199757 ],
[0.46216343, 4.0175133 ],
[0.46488592, 4.0149746 ],
[0.46760838, 4.0122066 ],
[0.47033085, 4.009954 ],
[0.47305333, 4.0075679 ],
[0.47577581, 4.0050669 ],
[0.47849828, 4.0023184 ],
[0.48122074, 3.9995501 ],
[0.48394321, 3.9969349 ],
[0.48666569, 3.9926589 ],
[0.48938816, 3.9889555 ],
[0.49211064, 3.9834003 ],
[0.4948331 , 3.9783037 ],
[0.49755557, 3.9755929 ],
[0.50027804, 3.9707632 ],
[0.50300052, 3.9681098 ],
[0.50572298, 3.9635665 ],
[0.50844545, 3.9594433 ],
[0.51116792, 3.9556634 ],
[0.51389038, 3.9521511 ],
[0.51661284, 3.9479132 ],
[0.51933531, 3.9438281 ],
[0.52205777, 3.9400866 ],
[0.52478024, 3.9362304 ],
[0.52750271, 3.9314201 ],
[0.53022518, 3.9283848 ],
[0.53294765, 3.9242232 ],
[0.53567012, 3.9192028 ],
[0.53839258, 3.9166257 ],
[0.54111506, 3.9117961 ],
[0.54383753, 3.90815 ],
[0.54656 , 3.9038739 ],
[0.54928247, 3.8995597 ],
[0.55200494, 3.8959136 ],
[0.5547274 , 3.8909314 ],
[0.55744986, 3.8872662 ],
[0.56017233, 3.8831048 ],
[0.5628948 , 3.8793442 ],
[0.56561729, 3.8747628 ],
[0.56833976, 3.8702576 ],
[0.57106222, 3.8666878 ],
[0.57378469, 3.8623927 ],
[0.57650716, 3.8581741 ],
[0.57922963, 3.854146 ],
[0.5819521 , 3.8499846 ],
[0.58467456, 3.8450022 ],
[0.58739702, 3.8422534 ],
[0.59011948, 3.8380919 ],
[0.59284194, 3.8341596 ],
[0.5955644 , 3.8309333 ],
[0.59828687, 3.8272109 ],
[0.60100935, 3.823164 ],
[0.60373182, 3.8192315 ],
[0.60645429, 3.8159864 ],
[0.60917677, 3.8123021 ],
[0.61189925, 3.8090379 ],
[0.61462172, 3.8071671 ],
[0.61734419, 3.8040555 ],
[0.62006666, 3.8013639 ],
[0.62278914, 3.7970879 ],
[0.62551162, 3.7953317 ],
[0.62823408, 3.7920673 ],
[0.63095656, 3.788383 ],
[0.63367903, 3.7855389 ],
[0.6364015 , 3.7838206 ],
[0.63912397, 3.78111 ],
[0.64184645, 3.7794874 ],
[0.64456893, 3.7769294 ],
[0.6472914 , 3.773608 ],
[0.65001389, 3.7695992 ],
[0.65273637, 3.7690265 ],
[0.65545884, 3.7662776 ],
[0.65818131, 3.7642922 ],
[0.66090379, 3.7626889 ],
[0.66362625, 3.7603791 ],
[0.66634874, 3.7575538 ],
[0.66907121, 3.7552056 ],
[0.67179369, 3.7533159 ],
[0.67451616, 3.7507198 ],
[0.67723865, 3.7487535 ],
[0.67996113, 3.7471499 ],
[0.68268361, 3.7442865 ],
[0.68540608, 3.7423012 ],
[0.68812855, 3.7400677 ],
[0.69085103, 3.7385788 ],
[0.6935735 , 3.7345319 ],
[0.69629597, 3.7339211 ],
[0.69901843, 3.7301605 ],
[0.7017409 , 3.7301033 ],
[0.70446338, 3.7278316 ],
[0.70718585, 3.7251589 ],
[0.70990833, 3.723861 ],
[0.71263081, 3.7215703 ],
[0.71535328, 3.7191267 ],
[0.71807574, 3.7172751 ],
[0.72079822, 3.7157097 ],
[0.72352069, 3.7130945 ],
[0.72624317, 3.7099447 ],
[0.72896564, 3.7071004 ],
[0.7316881 , 3.7045615 ],
[0.73441057, 3.703588 ],
[0.73713303, 3.70208 ],
[0.73985551, 3.7002664 ],
[0.74257799, 3.6972122 ],
[0.74530047, 3.6952841 ],
[0.74802293, 3.6929362 ],
[0.7507454 , 3.6898055 ],
[0.75346787, 3.6890991 ],
[0.75619034, 3.686522 ],
[0.75891281, 3.6849759 ],
[0.76163529, 3.6821697 ],
[0.76435776, 3.6808143 ],
[0.76708024, 3.6786573 ],
[0.7698027 , 3.6761947 ],
[0.77252517, 3.674763 ],
[0.77524765, 3.6712887 ],
[0.77797012, 3.6697233 ],
[0.78069258, 3.6678908 ],
[0.78341506, 3.6652565 ],
[0.78613753, 3.6630611 ],
[0.78885999, 3.660274 ],
[0.79158246, 3.6583652 ],
[0.79430494, 3.6554828 ],
[0.79702741, 3.6522949 ],
[0.79974987, 3.6499848 ],
[0.80247234, 3.6470451 ],
[0.8051948 , 3.6405547 ],
[0.80791727, 3.6383405 ],
[0.81063974, 3.635076 ],
[0.81336221, 3.633549 ],
[0.81608468, 3.6322317 ],
[0.81880714, 3.6306856 ],
[0.82152961, 3.6283948 ],
[0.82425208, 3.6268487 ],
[0.82697453, 3.6243098 ],
[0.829697 , 3.6223626 ],
[0.83241946, 3.6193655 ],
[0.83514192, 3.6177621 ],
[0.83786439, 3.6158531 ],
[0.84058684, 3.6128371 ],
[0.84330931, 3.6118062 ],
[0.84603177, 3.6094582 ],
[0.84875424, 3.6072438 ],
[0.8514767 , 3.6049912 ],
[0.85419916, 3.6030822 ],
[0.85692162, 3.6012688 ],
[0.85964409, 3.5995889 ],
[0.86236656, 3.5976417 ],
[0.86508902, 3.5951984 ],
[0.86781149, 3.593843 ],
[0.87053395, 3.5916286 ],
[0.87325642, 3.5894907 ],
[0.87597888, 3.587429 ],
[0.87870135, 3.5852909 ],
[0.88142383, 3.5834775 ],
[0.8841463 , 3.5817785 ],
[0.88686877, 3.5801177 ],
[0.88959124, 3.5778842 ],
[0.89231371, 3.5763381 ],
[0.8950362 , 3.5737801 ],
[0.89775868, 3.5721002 ],
[0.90048116, 3.5702102 ],
[0.90320364, 3.5684922 ],
[0.90592613, 3.5672133 ],
[1. , 3.52302167]])
from pybamm import exp, constants, Parameter
def graphite_LGM50_electrolyte_exchange_current_density_Chen2020(c_e, c_s_surf, T):
m_ref = 6.48e-7 # (A/m2)(mol/m3)**1.5 - includes ref concentrations
E_r = 35000
arrhenius = exp(E_r / constants.R * (1 / 298.15 - 1 / T))
c_n_max = Parameter("Maximum concentration in negative electrode [mol.m-3]")
return (
m_ref * arrhenius * c_e ** 0.5 * c_s_surf ** 0.5 * (c_n_max - c_s_surf) ** 0.5
)
def nmc_LGM50_electrolyte_exchange_current_density_Chen2020(c_e, c_s_surf, T):
m_ref = 3.42e-6 # (A/m2)(mol/m3)**1.5 - includes ref concentrations
E_r = 17800
arrhenius = exp(E_r / constants.R * (1 / 298.15 - 1 / T))
c_p_max = Parameter("Maximum concentration in positive electrode [mol.m-3]")
return (
m_ref * arrhenius * c_e ** 0.5 * c_s_surf ** 0.5 * (c_p_max - c_s_surf) ** 0.5
)
values = {
'Negative electrode thickness [m]': 8.52e-05,
'Separator thickness [m]': 1.2e-05,
'Positive electrode thickness [m]': 7.56e-05,
'Electrode height [m]': 0.065,
'Electrode width [m]': 1.58,
'Nominal cell capacity [A.h]': 5.0,
'Typical current [A]': 5.0,
'Current function [A]': 5.0,
'Maximum concentration in negative electrode [mol.m-3]': 33133.0,
'Negative electrode diffusivity [m2.s-1]': 3.3e-14,
'Negative electrode OCP [V]': ('graphite_LGM50_ocp_Chen2020', neg_ocp),
'Negative electrode porosity': 0.25,
'Negative electrode active material volume fraction': 0.75,
'Negative particle radius [m]': 5.86e-06,
'Negative electrode Bruggeman coefficient (electrolyte)': 1.5,
'Negative electrode electrons in reaction': 1.0,
'Negative electrode exchange-current density [A.m-2]': graphite_LGM50_electrolyte_exchange_current_density_Chen2020,
'Negative electrode OCP entropic change [V.K-1]': 0.0,
'Maximum concentration in positive electrode [mol.m-3]': 63104.0,
'Positive electrode diffusivity [m2.s-1]': 4e-15,
'Positive electrode OCP [V]': ('nmc_LGM50_ocp_Chen2020', pos_ocp),
'Positive electrode porosity': 0.335,
'Positive electrode active material volume fraction': 0.665,
'Positive particle radius [m]': 5.22e-06,
'Positive electrode Bruggeman coefficient (electrolyte)': 1.5,
'Positive electrode electrons in reaction': 1.0,
'Positive electrode exchange-current density [A.m-2]': nmc_LGM50_electrolyte_exchange_current_density_Chen2020,
'Positive electrode OCP entropic change [V.K-1]': 0.0,
'Separator porosity': 0.47,
'Separator Bruggeman coefficient (electrolyte)': 1.5,
'Typical electrolyte concentration [mol.m-3]': 1000.0,
'Reference temperature [K]': 298.15,
'Ambient temperature [K]': 298.15,
'Number of electrodes connected in parallel to make a cell': 1.0,
'Number of cells connected in series to make a battery': 1.0,
'Lower voltage cut-off [V]': 2.5,
'Upper voltage cut-off [V]': 4.4,
'Initial concentration in negative electrode [mol.m-3]': 29866.0,
'Initial concentration in positive electrode [mol.m-3]': 17038.0,
'Initial temperature [K]': 298.15
}
param = pybamm.ParameterValues(values)
param
# -
# Here we would have got the same result by doing
chemistry = pybamm.parameter_sets.Chen2020
param_same = pybamm.ParameterValues(chemistry=chemistry)
{k: v for k,v in param_same.items() if k in spm._parameter_info}
# ## Updating a specific parameter
# Once a parameter set has been defined (either via a dictionary or a pre-built set), single parameters can be updated
# ### Using a constant value:
# +
param.search("Current function [A]")
param.update(
{
"Current function [A]": 4.0
}
)
param["Current function [A]"]
# -
# ### Using a function:
# +
def curren_func(time):
return 1 + pybamm.sin(2 * np.pi * time / 60)
param.update(
{
"Current function [A]": curren_func
}
)
param["Current function [A]"]
# -
# ## Plotting parameter functions
#
# As seen above, functions can be passed as parameter values. These parameter values can then be plotted by using `pybamm.plot`
# ### Plotting "Current function \[A]"
currentfunc = param["Current function [A]"]
time = pybamm.linspace(0, 120, 60)
evaluated = param.evaluate(currentfunc(time))
evaluated = pybamm.Array(evaluated)
pybamm.plot(time, evaluated)
# Taking another such example:
#
# ### Plotting "Negative electrode exchange-current density \[A.m-2]"
negative_electrode_exchange_current_density = param["Negative electrode exchange-current density [A.m-2]"]
x = pybamm.linspace(3000,6000,100)
evaluated = param.evaluate(negative_electrode_exchange_current_density(1000,x,300))
evaluated = pybamm.Array(evaluated)
pybamm.plot(x, evaluated)
# ## Simulating and solving the model
#
# Finally we can simulate the model and solve it using `pybamm.Simulation` and `solve` respectively.
sim = pybamm.Simulation(spm, parameter_values=param)
t_eval = np.arange(0, 3600, 1)
sim.solve(t_eval=t_eval)
sim.plot()
# ## References
# The relevant papers for this notebook are:
pybamm.print_citations()
| examples/notebooks/parameterization/parameterization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="pzFdeoQzhAUu"
import pandas as pd
from sklearn.datasets import load_iris
from sklearn.cluster import KMeans
import matplotlib.pyplot as plt
# + id="n7nlEFMIhPiZ"
# importing UCI iris dataset, included with sklearn
iris = load_iris()
X = pd.DataFrame(iris.data, columns=iris['feature_names'])
data = X[['sepal length (cm)', 'sepal width (cm)', 'petal length (cm)']]
# + colab={"base_uri": "https://localhost:8080/"} id="lIgslEx3hSWV" outputId="f51be5c6-7ae6-4762-bb40-d7705bd5f482"
sse = {}
# trying k values of 1-5
for k in range(1, 6):
print("K VALUE: {}".format(k))
kmeans = KMeans(n_clusters=k, max_iter=100).fit(data)
data["clusters"] = kmeans.labels_
#sklearn inertia - Sum of squared distances of samples to their closest cluster center.
sse[k] = kmeans.inertia_
print("SSE: {}".format(sse[k]))
print("")
# + colab={"base_uri": "https://localhost:8080/", "height": 279} id="K7fzW-1_hVeb" outputId="f279fe4d-3a55-4c9b-8fab-dbbe8d22424f"
# plotting elbow method chart, shows ideal k value
plt.figure()
plt.plot(list(sse.keys()), list(sse.values()))
plt.xlabel("Cluster Number K")
plt.ylabel("SSE[K]")
plt.show()
| Clustering.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# #### New to Plotly?
# Plotly's Python library is free and open source! [Get started](https://plotly.com/python/getting-started/) by downloading the client and [reading the primer](https://plotly.com/python/getting-started/).
# <br>You can set up Plotly to work in [online](https://plotly.com/python/getting-started/#initialization-for-online-plotting) or [offline](https://plotly.com/python/getting-started/#initialization-for-offline-plotting) mode, or in [jupyter notebooks](https://plotly.com/python/getting-started/#start-plotting-online).
# <br>We also have a quick-reference [cheatsheet](https://images.plot.ly/plotly-documentation/images/python_cheat_sheet.pdf) (new!) to help you get started!
#
# ### Horizontal Legend
# +
import plotly.plotly as py
import plotly.graph_objs as go
import numpy as np
trace1 = go.Scatter(
x=np.random.randn(75),
mode='markers',
name="Plot1",
marker=dict(
size=16,
color='rgba(152, 0, 0, .8)'
))
trace2 = go.Scatter(
x=np.random.randn(75),
mode='markers',
name="Plot2",
marker=dict(
size=16,
color='rgba(0, 152, 0, .8)'
))
trace3 = go.Scatter(
x=np.random.randn(75),
mode='markers',
name="Plot3",
marker=dict(
size=16,
color='rgba(0, 0, 152, .8)'
))
data = [trace1, trace2, trace3]
layout = go.Layout(
legend=dict(
orientation="h")
)
figure=go.Figure(data=data, layout=layout)
py.iplot(figure)
# -
# #### Reference
# See https://plotly.com/python/reference/#layout-legend-orientation for more information and chart attribute options!
# +
from IPython.display import display, HTML
display(HTML('<link href="//fonts.googleapis.com/css?family=Open+Sans:600,400,300,200|Inconsolata|Ubuntu+Mono:400,700rel="stylesheet" type="text/css" />'))
display(HTML('<link rel="stylesheet" type="text/csshref="http://help.plot.ly/documentation/all_static/css/ipython-notebook-custom.css">'))
# ! pip install git+https://github.com/plotly/publisher.git --upgrade
import publisher
publisher.publish(
'horizontal-legends.ipynb', 'python/horizontal-legend/', 'Horizontal legend | plotly',
'How to add images to charts as background images or logos.',
title = 'Horizontal legend | plotly',
name = 'Horizontal Legends',
has_thumbnail='false', thumbnail='thumbnail/your-tutorial-chart.jpg',
language='python', page_type='example_index',
display_as='style_opt', order=12,
ipynb= '~notebook_demo/94')
# -
| _posts/python-v3/advanced/horizontal-legend/horizontal-legends.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.11 64-bit (''otopy38'': conda)'
# name: python3
# ---
from pydantic_wrap import SimplePageFuncPydanticWithOutput, pydantic_model_from_type
# +
def ff(x:int)->int:
return x
mytype = ff.__annotations__["return"]
mod = pydantic_model_from_type(mytype)
# -
mod.__fields__
# # flat dispatch
# +
from streamlitfront import dispatch_funcs
from front.scrap.pydantic_wrap import SimplePageFuncPydanticWithOutput
from typing import Optional
from streamlit_pydantic.types import FileContent
from pydantic import Field, BaseModel
from dol import Store
# +
def func(salary: float, n_months=12):
return salary * n_months
salary_store = {
'sylvain': 10000,
'christian': 2000,
'thor': 50000
}
def store_wrapped_func(salary: str, n_months=12):
salary = salary_store[salary]
return func(salary, n_months)
assert store_wrapped_func('sylvain', 6) == 60000
# +
from i2.wrapper import wrap
def ingress_salary(salary_str:str, n_months:int):
return (salary_store[salary_str], n_months), dict()
wrapped = wrap(func, ingress = ingress_salary)
assert wrapped('sylvain', 12)==120000
# +
"""A simple app to demo the use of Mappings to handle complex type"""
from typing import Mapping
ComplexType = float # just pretend it's complex!
def func(salary: ComplexType, n_months: int = 12):
return salary * n_months
SalaryKey = str # or some type that will resolve in store-fed key selector
SalaryMapping = Mapping[SalaryKey, ComplexType]
salary_store: SalaryMapping
salary_store = {"sylvain": 10000, "christian": 2000, "thor": 50000}
def store_wrapped_func(salary: SalaryKey, n_months: int = 12):
salary = salary_store[salary]
return func(salary, n_months)
assert store_wrapped_func("sylvain", 6) == 60000
if __name__ == "__main__":
from streamlitfront.base import dispatch_funcs
app = dispatch_funcs([func, store_wrapped_func])
app()
# +
"""A simple app to demo the use of Mappings to handle complex type"""
from typing import Mapping
from enum import Enum
from i2.wrapper import wrap
ComplexType = float # just pretend it's complex!
def func(salary: ComplexType, n_months: int = 12):
return salary * n_months
SalaryKey = str # or some type that will resolve in store-fed key selector
SalaryMapping = Mapping[SalaryKey, ComplexType]
salary_store: SalaryMapping
salary_store = {"sylvain": 10000, "christian": 2000, "thor": 50000}
def choose_key(choice):
pass
def ingress_salary(salary_str:SalaryKey, n_months:int):
return (salary_store[salary_str], n_months), dict()
store_wrapped_func = wrap(func, ingress = ingress_salary)
assert store_wrapped_func("sylvain", 6) == 60000
class SelectionValue(str, Enum):
FOO = "foo"
BAR = "bar"
class ExampleModel(BaseModel):
single_selection: SelectionValue = Field(
..., description="Only select a single item from a set."
)
# +
class SelectionValue(str, Enum):
FOO = "foo"
BAR = "bar"
class ExampleModel(BaseModel):
single_selection: SelectionValue = Field(
..., description="Only select a single item from a set."
)
# +
from dataclasses import dataclass, field, make_dataclass
from typing import List
store = {'sly':20000, 'chris':4000}
def mk_choices():
return list(store.keys())
@dataclass
class C:
mylist: List[str] = field(default_factory=mk_choices)
c = C()
c.mylist += [1, 2, 3]
# -
DFLT_CHOICES = ['sly','chris']
C = Enum("Choices", {'sly':'sly', 'chris':'chris'})
C
# # Scrap
# +
"""
Same as take_04_model_run, but where the dispatch is not as manual.
"""
# This is what we want our "dispatchable" wrapper to look like
# There should be real physical stores for those types (FVs, FittedModel) that need them
from typing import Any, Mapping, Tuple
import numpy as np
from sklearn.preprocessing import MinMaxScaler
FVs = Any
FittedModel = Any
# ---------------------------------------------------------------------------------------
# The function(ality) we want to dispatch:
def apply_model(fitted_model: FittedModel, fvs: FVs, method="transform"):
method_func = getattr(fitted_model, method)
return method_func(list(fvs))
# ---------------------------------------------------------------------------------------
# The stores that will be used -- here, all stores are just dictionaries, but the
# contract is with the typing.Mapping (read-only here) interface.
# As we grow up, we'll use other mappings, such as:
# - server side RAM (as done here, simply)
# - server side persistence (files or any DB or file system thanks to the dol package)
# - computation (when you want the request for a key to actually launch a process that
# will generate the value for you (some say you should be obvious to that detail))
# - client side RAM (when we figure that out)
mall = dict(
fvs=dict(
train_fvs_1=np.array([[1], [2], [3], [5], [4], [2], [1], [4], [3]]),
train_fvs_2=np.array([[1], [10], [5], [3], [4]]),
test_fvs=np.array([[1], [5], [3], [10], [-5]]),
),
fitted_model=dict(
fitted_model_1=MinMaxScaler().fit(
[[1], [2], [3], [5], [4], [2], [1], [4], [3]]
),
fitted_model_2=MinMaxScaler().fit([[1], [10], [5], [3], [4]]),
),
model_results=dict(),
)
# ---------------------------------------------------------------------------------------
# dispatchable function:
from streamlitfront.examples.crude.crude_util import auto_key
from i2 import Sig
from i2.wrapper import Ingress, wrap
from inspect import Parameter
def prepare_for_crude_dispatch(func, store_for_param=None, output_store_name=None):
"""Wrap func into something that is ready for CRUDE dispatch."""
ingress = None
if store_for_param is not None:
sig = Sig(func)
print('sig:', sig)
crude_params = [x for x in sig.names if x in store_for_param]
print(f'crude params: {crude_params}')
def kwargs_trans(outer_kw):
def gen():
for store_name in crude_params:
store_key = outer_kw[store_name]
yield store_name, store_for_param[store_name][store_key]
return dict(gen())
save_name_param = Parameter(
name="save_name",
kind=Parameter.KEYWORD_ONLY,
default="",
annotation=str,
)
print(save_name_param)
outer_sig=(
sig.ch_annotations(**{name: str for name in crude_params})
+ [save_name_param]
)
print(outer_sig)
ingress = Ingress(
inner_sig=sig,
kwargs_trans=kwargs_trans,
outer_sig=(
sig.ch_annotations(**{name: str for name in crude_params})
+ [save_name_param]
),
)
egress = None
if output_store_name:
def egress(func_output):
print(f"{list(store_for_param[output_store_name])=}")
store_for_param[output_store_name] = func_output
print(f"{list(store_for_param[output_store_name])=}")
return func_output
wrapped_f = wrap(func, ingress, egress)
return wrapped_f
def prepare_for_crude_dispatch2(func, store_for_param=None, output_store_name=None):
"""Wrap func into something that is ready for CRUDE dispatch."""
ingress = None
if store_for_param is not None:
sig = Sig(func)
crude_params = [x for x in sig.names if x in store_for_param]
def kwargs_trans(outer_kw):
def gen():
for store_name in crude_params:
store_key = outer_kw[store_name]
yield store_name, store_for_param[store_name][store_key]
return dict(gen())
save_name_param = Parameter(
name="save_name",
kind=Parameter.KEYWORD_ONLY,
default="",
annotation=str,
)
outer_sig=(
sig.ch_annotations(**{name: str for name in crude_params})
+ [save_name_param]
)
print(outer_sig)
ingress = Ingress(
inner_sig=sig,
kwargs_trans=kwargs_trans,
outer_sig=(
sig.ch_annotations(**{name: str for name in crude_params})
+ [save_name_param]
),
)
egress = None
if output_store_name:
def egress(func_output):
print(f"{list(store_for_param[output_store_name])=}")
store_for_param[output_store_name] = func_output
print(f"{list(store_for_param[output_store_name])=}")
return func_output
wrapped_f = wrap(func, ingress, egress)
return wrapped_f
f = prepare_for_crude_dispatch(apply_model, mall)
assert all(
f("fitted_model_1", "test_fvs") == np.array([[0.0], [1.0], [0.5], [2.25], [-1.5]])
)
# -
from dataclasses import make_dataclass
from pydantic import BaseModel
CC = make_dataclass('Y', fields=[('s', str), ('x', int)], bases=(BaseModel,))
CC.__fields__
issubclass(CC, BaseModel)
func = apply_model
sig = Sig(func)
sig.names
dict(sig)
from inspect import signature
sg = signature(func)
sg.parameters['fvs'].annotation
sig.annotations.items()
# +
from typing import List
mall = {'x':{'xs_1':3, 'xs_2':5}, 'fvs':{'fv_1':[1,2,3], 'fv_2':[4,5,6]}}
def f(x:int, fvs:List[int]):
return [x*item for item in fvs]
sig_list = Sig(f).annotations.items()
CC = make_dataclass('Input', fields=sig_list, bases=(BaseModel,))
# +
import streamlit_pydantic as sp
import streamlit as st
from pydantic import BaseModel, Field, ValidationError, parse_obj_as
class InputFromStore:
pass
from enum import Enum
from dataclasses import field
CC = make_dataclass('Input', fields=[('FOO',str, field(init='foo')),('BAR', str, field(init='bar'))], bases=(str,Enum,))
class SelectionValue(str, Enum):
FOO = "foo"
BAR = "bar"
def mk(store, key):
choices = store[key].keys()
class ExampleModel(BaseModel):
single_selection: SelectionValue= Field(
..., description="Only select a single item from a set."
)
data = sp.pydantic_form(key="my_form", model=ExampleModel)
if data:
st.json(data.json())
# -
CC.__annotations__
Input = make_dataclass(
"Input",
fields=[("FOO", str, field(init="foo")), ("BAR", str, field(init="bar"))],
bases=(
str,
Enum,
),
)
dir(Input)
from dataclasses import dataclass
make_dataclass('Input',fields=[('x',int)])
# +
mall = {"x": {"xs_1": 3, "xs_2": 5}, "fvs": {"fv_1": [1, 2, 3], "fv_2": [4, 5, 6]}}
def mk_Enum_from_store(store, key):
choices = store[key].keys()
choice = Enum(f"DynamicEnum_{key}", {item: item for item in choices})
return choice
class ExampleModel2(BaseModel):
fvs: mk_Enum_from_store(mall, key="x")
# +
from pydantic import BaseModel, create_model
DynamicFoobarModel = create_model('ExampleModel3', fvs=(mk_Enum_from_store(mall, key="x"), ...))
# -
from typing_extensions import TypedDict
Movie = TypedDict('Movie', {'name': str, 'year': int})
from pydantic import create_model_from_typeddict
UserM = create_model_from_typeddict(Movie)
UserM
from dol import Files
| front/scrap/notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import os, sys
sys.path.append(os.environ['HOME'] + '/src/models/')
from deeplearning_models import DLTextClassifier
from sklearn.model_selection import train_test_split
# ### Experiment 1
#
# Train: C3 Train
# Test: C3 Test
def run_dl_experiment(C3_train_df,
C3_test_df,
results_csv_path = os.environ['HOME'] + 'models/CNN_predictions.csv',
model = 'cnn'):
"""
"""
X_train = C3_train_df['pp_comment_text'].astype(str)
y_train = C3_train_df['constructive_binary']
X_test = C3_test_df['pp_comment_text'].astype(str)
y_test = C3_test_df['constructive_binary']
dlclf = DLTextClassifier(X_train, y_train)
if model.endswith('lstm'):
dlclf.build_bilstm()
elif model.endswith('cnn'):
dlclf.build_cnn()
dlclf.train(X_train, y_train)
print('\nTrain results: \n\n')
dlclf.evaluate(X_train, y_train)
print('\nTest results: \n\n')
dlclf.evaluate(X_test, y_test)
results_df = dlclf.write_model_scores_df(C3_test_df, results_csv_path)
# ### Experiment 1
#
# - Train: C3 Train (80%)
# - Test: C3 Test (20%)
C3_train_df = pd.read_csv(os.environ['C3_TRAIN'])
C3_test_df = pd.read_csv(os.environ['C3_TEST'])
C3_test_df.shape
# +
#run_dl_experiment(C3_train_df,
# C3_test_df,
# results_csv_path = os.environ['HOME'] + 'models/biLSTM_C3_test_predictions.csv',
# model = 'lstm')
# -
run_dl_experiment(C3_train_df, C3_test_df, model = 'cnn')
# ### Experiment 2
#
# - Train: C3_MINUS_LB + C3-LB train (80%)
# - Test: C3 Test
C3_train_df = pd.read_csv(os.environ['C3_MINUS_LB'])
C3_test_df = pd.read_csv(os.environ['C3_LB'])
C3_train_df.columns
feats = ['comment_counter', 'pp_comment_text', 'constructive']
LB_X_train, LB_X_test, LB_y_train, LB_y_test = train_test_split(C3_test_df[feats], C3_test_df['constructive_binary'], train_size = 0.80, random_state=1)
inter_df = pd.concat([LB_X_train, LB_y_train], axis = 1)
inter_df.head()
C3_train_df.head()
train_df = pd.concat([C3_train_df, inter_df])
train_df.shape
test_df = pd.concat([LB_X_test, LB_y_test], axis = 1)
run_dl_experiment(train_df, test_df, model = 'cnn')
run_dl_experiment(train_df, test_df, model = 'lstm')
# ### Experiment 3
#
# - Train: LB Train (80%)
# - Test: LB test (20%)
LB_train_df = pd.concat([LB_X_train, LB_y_train], axis = 1)
LB_test_df = pd.concat([LB_X_test, LB_y_test], axis = 1)
run_dl_experiment(LB_train_df, LB_test_df, model = 'cnn')
run_dl_experiment(LB_train_df, LB_test_df, model = 'lstm')
# ### Experiment 4
#
# - Train: LB Train (80%)
# - Test: C3 test (20%)
run_dl_experiment(LB_train_df, C3_test_df, model = 'cnn')
run_dl_experiment(LB_train_df, C3_test_df, model = 'lstm')
| notebooks/3.0-kvarada-length-experiments.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + slideshow={"slide_type": "skip"}
# %matplotlib inline
# + [markdown] slideshow={"slide_type": "slide"}
# # Fetching data from the IOOS CSW catalog
#
# This notebook shows a typical workflow to query a [Catalog Service for the Web (CSW)](https://en.wikipedia.org/wiki/Catalog_Service_for_the_Web) and create a request for data endpoints that are suitable for download.
#
# The first step is to create the data filter based on the **geographical** region bounding box, the **time** span, and the CF **variable** standard name.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Region: west coast
# + slideshow={"slide_type": "-"}
min_lon, max_lon = -123, -121
min_lat, max_lat = 36, 40
bbox = [min_lon, min_lat, max_lon, max_lat]
crs = "urn:ogc:def:crs:OGC:1.3:CRS84"
# + [markdown] slideshow={"slide_type": "slide"}
# ### Temporal range: past week
# + slideshow={"slide_type": "-"}
from datetime import datetime, timedelta
now = datetime.utcnow()
start, stop = now - timedelta(days=(14)), now - timedelta(days=(7))
# + [markdown] slideshow={"slide_type": "slide"}
# ### Surface velocity CF names
# + slideshow={"slide_type": "-"}
cf_names = [
"surface_northward_sea_water_velocity",
"surface_eastward_sea_water_velocity",
]
# + slideshow={"slide_type": "slide"}
print(f"""
*standard_names*: {cf_names}
*start and stop dates*: {start} to {stop}
*bounding box*:{bbox}
*crs*: {crs}""")
# + [markdown] slideshow={"slide_type": "slide"}
# Now it is possible to assemble a [OGC Filter Encoding (FE)](http://www.opengeospatial.org/standards/filter) for the search using `owslib.fes`\*. Note that the final result is only a list with all the filtering conditions.
#
# \* OWSLib is a Python package for client programming with Open Geospatial Consortium (OGC) web service (hence OWS) interface standards, and their related content models.
# + slideshow={"slide_type": "slide"}
from owslib import fes
def fes_date_filter(start, stop, constraint="overlaps"):
start = start.strftime("%Y-%m-%d %H:00")
stop = stop.strftime("%Y-%m-%d %H:00")
if constraint == "overlaps":
propertyname = "apiso:TempExtent_begin"
begin = fes.PropertyIsLessThanOrEqualTo(
propertyname=propertyname, literal=stop)
propertyname = "apiso:TempExtent_end"
end = fes.PropertyIsGreaterThanOrEqualTo(
propertyname=propertyname, literal=start)
elif constraint == "within":
propertyname = "apiso:TempExtent_begin"
begin = fes.PropertyIsGreaterThanOrEqualTo(
propertyname=propertyname, literal=start)
propertyname = "apiso:TempExtent_end"
end = fes.PropertyIsLessThanOrEqualTo(
propertyname=propertyname, literal=stop)
else:
raise NameError(f"Unrecognized constraint {constraint}")
return begin, end
# + [markdown] slideshow={"slide_type": "slide"}
# ### Create the filter
# + slideshow={"slide_type": "-"}
kw = dict(wildCard='*', escapeChar='\\',
singleChar='?', propertyname='apiso:AnyText')
or_filt = fes.Or([fes.PropertyIsLike(literal=('*%s*' % val), **kw)
for val in cf_names])
# Exclude GNOME returns.
not_filt = fes.Not([fes.PropertyIsLike(literal='*GNOME*', **kw)])
begin, end = fes_date_filter(start, stop)
bbox_crs = fes.BBox(bbox, crs=crs)
filter_list = [fes.And([bbox_crs, begin, end, or_filt, not_filt])]
# + slideshow={"slide_type": "slide"}
def get_csw_records(csw, filter_list, pagesize=10,
maxrecords=1000):
"""Iterate maxrecords/pagesize times until the requested
value in maxrecords is reached."""
from owslib.fes import SortBy, SortProperty
sortby = SortBy([SortProperty("dc:title", "ASC")])
csw_records = {}
startposition = 0
nextrecord = getattr(csw, "results", 1)
while nextrecord != 0:
csw.getrecords2(constraints=filter_list, startposition=startposition,
maxrecords=pagesize, sortby=sortby)
csw_records.update(csw.records)
if csw.results["nextrecord"] == 0:
break
startposition += pagesize + 1 # Last one is included.
if startposition >= maxrecords:
break
csw.records.update(csw_records)
# + slideshow={"slide_type": "slide"}
from owslib.csw import CatalogueServiceWeb
endpoint = "https://data.ioos.us/csw"
csw = CatalogueServiceWeb(endpoint, timeout=60)
get_csw_records(csw, filter_list, pagesize=10, maxrecords=1000)
records = "\n".join(csw.records.keys())
# + slideshow={"slide_type": "slide"}
print(f"Found {len(csw.records.keys())} records.\n")
for key, value in list(csw.records.items()):
print(f"[{value.title}]: {key}")
# + [markdown] slideshow={"slide_type": "slide"}
# Let us check the 6 km resolution metadata record we found above.
# + slideshow={"slide_type": "-"}
value = csw.records[
"HFR/USWC/6km/hourly/RTV/"
"HFRADAR_US_West_Coast_6km_Resolution_Hourly_RTV_best.ncd"
]
print(value.abstract)
# + slideshow={"slide_type": "slide"}
attrs = [attr for attr in dir(value) if not attr.startswith("_")]
nonzero = [attr for attr in attrs if getattr(value, attr)]
nonzero
# + [markdown] slideshow={"slide_type": "slide"}
# ### What is in there?
# + slideshow={"slide_type": "-"}
value.subjects
# + [markdown] slideshow={"slide_type": "slide"}
# ### Is it up-to-date?
# + slideshow={"slide_type": "-"}
value.modified
# + [markdown] slideshow={"slide_type": "slide"}
# ### The actual bounding box of the data
# + slideshow={"slide_type": "-"}
bbox = (
value.bbox.minx, value.bbox.miny,
value.bbox.maxx, value.bbox.maxy
)
bbox
# + [markdown] slideshow={"slide_type": "slide"}
# ### Now we can "sniff" the URLs with *geolinks*
# + slideshow={"slide_type": "-"}
from geolinks import sniff_link
for ref in value.references:
if ref["scheme"] == "OPeNDAP:OPeNDAP":
url = ref["url"] # save the opendap for later
geolink = sniff_link(ref["url"])
msg = (f"geolink: {geolink}\nscheme: {ref['scheme']}\n"
f"URL: {ref['url']}\n")
print(msg)
# + [markdown] slideshow={"slide_type": "slide"}
# For a detailed description of what those `geolink` results mean check the [lookup](https://github.com/OSGeo/Cat-Interop/blob/master/LinkPropertyLookupTable.csv) table.
# There are Web Coverage Service (WCS), Web Map Service (WMS),
# direct links, and OPeNDAP services available.
#
# We can use any of those to obtain the data but the easiest one to explore interactively is the open OPeNDAP endpoint.
# + slideshow={"slide_type": "slide"}
import xarray as xr
ds = xr.open_dataset(url)
ds
# + [markdown] slideshow={"slide_type": "slide"}
# ### Select "yesterday" data
# + slideshow={"slide_type": "-"}
from datetime import date, timedelta
yesterday = date.today() - timedelta(days=1)
ds = ds.sel(time=yesterday)
# + [markdown] slideshow={"slide_type": "slide"}
# ### Compute the speed while masking invalid values
# + slideshow={"slide_type": "-"}
import numpy.ma as ma
u = ds["u"].data
v = ds["v"].data
lon = ds.coords["lon"].data
lat = ds.coords["lat"].data
time = ds.coords["time"].data
u = ma.masked_invalid(u)
v = ma.masked_invalid(v)
# + [markdown] slideshow={"slide_type": "slide"}
# This cell is only a trick to show all quiver arrows with the same length,
# for visualization purposes,
# and indicate the vector magnitude with colors instead.
# + slideshow={"slide_type": "fragment"}
import numpy as np
from oceans.ocfis import uv2spdir, spdir2uv
angle, speed = uv2spdir(u, v)
us, vs = spdir2uv(np.ones_like(speed), angle, deg=True)
# + [markdown] slideshow={"slide_type": "slide"}
# ### And now we are ready to create the plot
# + slideshow={"slide_type": "-"}
import cartopy.crs as ccrs
import matplotlib.pyplot as plt
from cartopy import feature
from cartopy.mpl.gridliner import LONGITUDE_FORMATTER, LATITUDE_FORMATTER
LAND = feature.NaturalEarthFeature(
"physical", "land", "10m",
edgecolor="face",
facecolor="lightgray",
)
sub = 2
dx = dy = 0.5
center = -122.416667, 37.783333 # San Francisco.
bbox = lon.min(), lon.max(), lat.min(), lat.max()
# + slideshow={"slide_type": "skip"}
def plot_hfradar():
fig, (ax0, ax1) = plt.subplots(
ncols=2,
figsize=(20, 20),
subplot_kw=dict(projection=ccrs.PlateCarree())
)
ax0.set_extent(bbox)
cs = ax0.pcolormesh(lon, lat, ma.masked_invalid(speed))
gl = ax0.gridlines(draw_labels=True)
gl.xlabels_top = gl.ylabels_right = False
gl.xformatter = LONGITUDE_FORMATTER
gl.yformatter = LATITUDE_FORMATTER
cbar = fig.colorbar(cs, ax=ax0, shrink=0.45, extend="both")
cbar.ax.set_title(r"speed m s$^{-1}$", loc="left")
ax0.add_feature(LAND, zorder=0, edgecolor="black")
ax0.set_title(f"{value.title}\n{ds['time'].values}")
ax1.set_extent([center[0]-dx-dx, center[0]+dx, center[1]-dy, center[1]+dy])
q = ax1.quiver(lon[::sub], lat[::sub],
us[::sub, ::sub], vs[::sub, ::sub],
speed[::sub, ::sub], scale=30)
ax1.quiverkey(q, 0.5, 0.85, 1, r"1 m s$^{-1}$", coordinates="axes")
gl = ax1.gridlines(draw_labels=True)
gl.xlabels_top = gl.ylabels_right = False
gl.xformatter = LONGITUDE_FORMATTER
gl.yformatter = LATITUDE_FORMATTER
ax1.add_feature(LAND, zorder=0, edgecolor="black")
ax1.plot(ds["site_lon"], ds["site_lat"], marker="o", linestyle="none", color="darkorange")
ax1.set_title("San Francisco Bay area");
# + slideshow={"slide_type": "slide"}
plot_hfradar();
# + [markdown] slideshow={"slide_type": "slide"}
# ### Try to create your own filter and enjoy data!
#
# ```python
# filter_list = [
# fes.And(
# [
# bbox_crs,
# begin, end,
# or_filt, not_filt,
# ]
# )
# ]
# ```
| 00-IOOS-CSW-HFRadar.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # "Etiquetado de variables y valores en las encuestas de INEGI usando Python"
# > "Para entender los datos de encuestas es necesario contar con un diccionario de variables y un catálogo de valores para las variables que se analizan. Organizar esta información puede ser un reto cuando estos metadatos vienen en archivos planos como `.csv`. Aquí muestro una aplicación usando datos de la ENSU de INEGI."
# - toc: true
# - badges: true
# - comments: true
# - categories: [pandas, encuestas, inegi]
# - permalink: /etiquetas-encuestas-inegi/
# > Elaborado por <NAME> ([@jjsantoso](https://twitter.com/jjsantoso))
# ## Introducción
# Hace poco me tocó trabajar con los datos de una encuesta de INEGI y usé Python para hacer el análisis descriptivo. Quienes trabajamos con datos del INEGI hemos visto que es usual que los archivos de datos abiertos vengan en varias carpetas que contienen tanto los datos como los metadatos e información adicional sobre la encuesta. Por ejemplo, si descargamos los datos para la [Encuesta Nacional de Seguridad Pública Urbana (ENSU)](https://www.inegi.org.mx/programas/ensu/#Datos_abiertos) veremos que vienen 3 carpetas, cada una corresponde a una sección de la encuesta:
#
# > Note: Los datos se pueden descargar directamente de la página de INEGI, pero aquí dejo una copia de los que yo usé para este tutorial. [Descargar datos](http://jjsantoso.com/blog/datos/conjunto_de_datos_ENSU_2020_4t_csv.zip)
#
#
# * `conjunto_de_datos_VIV_ENSU_12_2020`: Cuestionario sociodemográfico sección I y II
# * `conjunto_de_datos_CS_ENSU_12_2020`: Cuestionario sociodemográfico sección III
# * `conjunto_de_datos_CB_ENSU_12_2020`: Cuestionario principal de la encuesta sección I, II, III y IV
#
# Si vemos al interior de uno de estos módulos, la estructura incluye las siguientes carpetas:
#
# * **Catálogos**: tiene los catálogos para cada variable en el cuestionario
# * **Conjunto de datos**: tiene los datos principales de la encuesta
# * **Diccionario de datos**: el nombre e información de cada variable
# * **Metadatos**: tiene información de la encuesta.
# * **Modelo entidad relación**: es un diagrama que muestra cómo se relacionan los diferentes conjuntos de datos.
# 
# Si echamos un vistazo rápido a los datos en Excel (`conjunto_de_datos/conjunto_de_datos_CB_ENSU_12_2020.csv`) veremos que la mayor parte de las variables viene codificada. Solo con este archivo no podemos saber qué es cada columna y cuáles es el significado de sus valores. Nos hace falta el diccionario de variables y los catálogos para poder interpretarlas.
# 
# En el archivo `diccionario_de_datos/diccionario_de_datos_CB_ENSU_12_2020.csv` tenemos cuál es el texto de cada pregunta. De ahí sabemos que, por ejemplo, la pregunta `BP1_1` es "Percepción de seguridad en la ciudad". Estos valores se conocen como etiquetas de las variables.
#
# Por otro lado, dentro de la carpeta `catalogos` viene un archivo `csv` por cada variable del conjunto de datos.
# 
# Este archivo nos dice cómo debemos transformar los valores numéricos de las categorías por sus valores de texto. Por ejemplo, si abrimos el archivo "BP1_1.csv" su contenido nos muestra que para la variable `BP1_1` debemos interpretar que un 1 corresponde a la categorías "seguro?", el 2 corresponde a "inseguro?" y el 9 a "No sabe/No responde". Estos valores se conocen como etiquetas de los valores.
#
# > Note: Las etiquetas de valores tiene sentido para variables categóricas, es decir las que tienen pocos valores nominales. Para variables numéricas o puramente de texto no es necesario usar etiquetas de valores.
# 
# Es evidente que para manejar una encuesta es fundamental conocer las etiquetas de las variables y sus valores. Sería mucho más fácil si estas etiquetas estuvieran incluidas en el mismo archivo junto con los datos, pero como están en formato `.csv` no es posible guadar esa información en un solo archivo y por tanto, termina repartida en muchos. Entonces, nuestro objetivo es integrar el diccionario y el catálogo a los datos para que sea más fácil hacer nuestro análisis. Queremos que en las tablas o gráficas que hagamos, las variables categóricas aparezcan como texto, en lugar de los valores numéricos que asignó INEGI. De igual forma, nos gustaría que en lugar de aparecer el nombre de la variable como en la base de datos, aparezca su descripción. Para lograr esto usaremos objetos tipo diccionario nativos de Python y dataframes de Pandas.
#
# > Tip: Otros formatos de datos, como por ejemplo los archivos `.dta` de Stata o `.sav` de SPSS sí permiten guardar esas etiquetas junto con los datos, sin embargo, esos no son formatos de datos abiertos que sean fácilmente accesible. En algunos casos, como en la [ENOE](https://www.inegi.org.mx/programas/enoe/15ymas/#Microdatos), INEGI también publica archivos `.dta` y `.sav`.
#
# ## Datos
#
# Primero, vamos a importar las librerías necesarias.
# +
import glob
import sys
import pandas as pd
print('Python', sys.version)
print(pd.__name__, pd.__version__)
# -
# Para ilustrar, vamos a seleccionar los datos de la carpeta `conjunto_de_datos_CB_ENSU_12_2020` (Cuestionario principal de la encuesta sección I, II, III y IV).
datos = pd.read_csv('conjunto_de_datos_CB_ENSU_12_2020/conjunto_de_datos/conjunto_de_datos_CB_ENSU_12_2020.csv')
datos.head()
datos.info()
# Necesitamos el diccionario de variables y los catálogos. Para el diccionario de variables cargamos el archivo `conjunto_de_datos_CB_ENSU_12_2020/diccionario_de_datos/diccionario_de_datos_CB_ENSU_12_2020.csv`
preguntas = pd.read_csv('conjunto_de_datos_CB_ENSU_12_2020/diccionario_de_datos/diccionario_de_datos_CB_ENSU_12_2020.csv', encoding='latin1')
preguntas.head(10)
# Este nos muestra cómo aparece cada variable en el archivo de datos ("NEMONICO") y cuál es su descripción ("NOMBRE_CAMPO"), además contiene el tipo y rango de valores que puede tener cada variable. Por esta razón, el nombre de cada variable puede estar repetido varias veces. Lo que haremos es eliminar los duplicados y quedarnos con los valores únicos, para después crear un objeto diccionario que tenga como llaves el "NEMONICO" y como valores el "NOMBRE_CAMPO". A continuación se ve el proceso y el resultado:
dicc_preguntas = preguntas.drop_duplicates(subset=['NEMONICO']).set_index('NEMONICO')['NOMBRE_CAMPO'].to_dict()
print(str(dicc_preguntas)[:500])
# Para el catálogo tenemos que trabajar un poco más porque la información está en muchos archivos. Primero vamos a generar una lista de todos los archivos en la carpeta `catalogo`.
dir_catalogos = 'conjunto_de_datos_CB_ENSU_12_2020/catalogos/'
archivos_catalogo_respuestas = glob.glob1(dir_catalogos, '*.csv')
archivos_catalogo_respuestas[:10]
# A continuación vamos a leer cada uno de los archivos individuales y generar un diccionario por comprensión que contenga como llaves el "NEMONICO" y los valores serán otro diccionario que tiene la relación de los valores numéricos y de texto para los valores de cada variable.
dicc_respuestas = {
f[:-4]: pd.read_csv(f'{dir_catalogos}/{f}', encoding='latin1', index_col=f[:-4])['descrip'].to_dict() for f in archivos_catalogo_respuestas
}
print(str(dicc_respuestas)[:500])
# Ahora tenemos dos diccionarios, uno con las etiquetas de las variables (`dicc_preguntas`) y otro con las etiquetas de los valores de cada pregunta `dicc_respuestas`. En ambos diccionarios la llave principal es el némonico de la pregunta, por tanto si queremos saber cuáles son las etiquetas de variables y valores solo tenemos que indexar los diccionarios con el nemónico correspondiente, por ejemplo para "SEX" y "BP1_1" obtenemos:
print('Etiqueta de variable:', dicc_preguntas['SEX'], '\nEtiqueta de valores',dicc_respuestas['SEX'])
print('Etiqueta de variable:', dicc_preguntas['BP1_1'], '\nEtiqueta de valores',dicc_respuestas['BP1_1'])
# ## Uso de las etiquetas
#
# Ya que tenemos las etiquetas, veamos cómo podemos usarlas para interpretar mejor en nuestros análisis. Hagamos una tabulación cruzada para ver la distribución de respuestas de las variables "SEX" y "BP1_1"
#
# > Warning: En estos ejemplos no estamos considerando que cada observación tiene una ponderación distinta (variable `FAC_SEL`) por tanto los resultados no son estimaciones válidas. En una próxima entrada veremos cómo integrar las ponderaciones.
pd.crosstab(datos['SEX'], datos['BP1_1'])
# El índice del dataframe y los nombres de las columnas contienen los valores numéricos de las categorías. Vamos a reemplazarlos por sus valores de texto renombrándolo con el método `.rename()`
pd.crosstab(datos['SEX'], datos['BP1_1'])\
.rename(index=dicc_respuestas['SEX'],
columns=dicc_respuestas['BP1_1'])
# Ahora es mucho más fácil de entender esta tabla.
#
# Vamos a hacer otra tabla similar a la anterior, pero en este caso desagregando por más variables y usando el método groupby:
vars_by = ['SEX', 'BP1_1', 'BP1_5_1']
grouped = datos.groupby(vars_by).agg(N=('ID_PER', 'count'))
grouped.head(15)
# Nuevamente reemplazamos los valores de las categorías usando el método `.rename()`. Hacemos un loop para reemplazar las categorías de cada variable. Como tenemos varios niveles en el índice, especificamos la opción `level` para que use solo el catálogo con la variable que le corresponde.
# +
for v in vars_by:
grouped.rename(index=dicc_respuestas[v], level=v, inplace=True)
grouped
# -
# Para que sea más fácil de ver, reestructuramos la tabla usando `.unstack()`.
grouped.unstack('BP1_1')
# Ya integramps las etiquetas de los valores, ahora faltan las etiquetas de las variables. Para eso usaremos el método `.rename_axis()`
# +
cuadro = grouped.unstack('BP1_1')\
.rename_axis(index=dicc_preguntas, columns=dicc_preguntas)
cuadro
# -
# Nuestro cuadro ya es entendible, podemos exportarlo a Excel y verificar que tenemos las etiquetas:
cuadro.to_excel('reporte.xlsx')
# 
# ## Si usas Stata...(o incluso si no)
#
# Como dije antes, el formato `.dta` permite guardar las etiquetas de variables y valores junto con los datos. Los DataFrames de Pandas traen de forma nativa el método [`.to_stata()`](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.to_stata.html) para exportar la información a este formato. Para que Stata reconozca correctamnete las etiquetas de valores es necesario que en pandas las variables sean de [tipo categoria](https://pandas.pydata.org/pandas-docs/stable/user_guide/categorical.html). A continuación, vamos a seleccionar un subconjunto de variables, reemplazaremos sus valores numéricos por las categorías de texto y convertiremos la variable en tipo categórica:
vars_export = ['SEX', 'BP1_1', 'BP1_2_01', 'BP1_2_02', 'BP1_2_03', 'BP1_2_04', 'BP1_2_05', 'BP1_2_06', 'BP1_2_07', 'BP1_2_08', 'BP1_2_09', 'BP1_2_10', 'BP1_2_11', 'BP1_2_12']
datos_stata = datos[vars_export].apply(lambda s: s.map(dicc_respuestas[s.name])).astype('category')
datos_stata
# Este dataframe lo exportaremos a Stata, junto con el diccionario que contiene las etiquetas de variables, especificando en la opción `variable_labels`.
datos_stata.to_stata('datos_stata.dta', write_index=False, variable_labels=dicc_preguntas)
# Al abrir el arcivo en Stata podemos ver que efectivamente se guardaron las etiquetas de los valores y de las variables:
# 
# Hasta donde entiendo, esta sería una forma más fácil de etiquetar datos provenientes de INEGI para usuarios de Stata que haciendo el procedimiento entero en Stata. Igualmente, para los que no son usuarios de Stata, pero sí de Python o R y quieren conservar las variables categóricas etiquetadas este es un buen formato.
datos_2 = pd.read_stata('datos_stata.dta')
datos_2.dtypes
| _notebooks/2021-03-08-etiquetas en encuestas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="406c3a58-001" colab_type="text"
# #1. Install Dependencies
# First install the libraries needed to execute recipes, this only needs to be done once, then click play.
#
# + id="406c3a58-002" colab_type="code"
# !pip install git+https://github.com/google/starthinker
# + [markdown] id="406c3a58-003" colab_type="text"
# #2. Get Cloud Project ID
# To run this recipe [requires a Google Cloud Project](https://github.com/google/starthinker/blob/master/tutorials/cloud_project.md), this only needs to be done once, then click play.
#
# + id="406c3a58-004" colab_type="code"
CLOUD_PROJECT = 'PASTE PROJECT ID HERE'
print("Cloud Project Set To: %s" % CLOUD_PROJECT)
# + [markdown] id="406c3a58-005" colab_type="text"
# #3. Get Client Credentials
# To read and write to various endpoints requires [downloading client credentials](https://github.com/google/starthinker/blob/master/tutorials/cloud_client_installed.md), this only needs to be done once, then click play.
#
# + id="406c3a58-006" colab_type="code"
CLIENT_CREDENTIALS = 'PASTE CREDENTIALS HERE'
print("Client Credentials Set To: %s" % CLIENT_CREDENTIALS)
# + [markdown] id="406c3a58-007" colab_type="text"
# #4. Enter DV360 Entity Read Files To BigQuery Parameters
# Import public and private <a href='https://developers.google.com/bid-manager/guides/entity-read/format-v2' target='_blank'>Entity Read Files</a> into a BigQuery dataset.<br/>CAUTION: PARTNER ONLY, ADVERTISER FILTER IS NOT APPLIED.
# 1. Entity Read Files ONLY work at the partner level.
# 1. Advertiser filter is NOT APPLIED.
# 1. Specify one or more partners to be moved into the dataset.
# Modify the values below for your use case, can be done multiple times, then click play.
#
# + id="406c3a58-008" colab_type="code"
FIELDS = {
'auth_write': 'service', # Credentials used for writing data.
'auth_read': 'user', # Credentials used for reading data.
'partners': '[]', # Comma sparated list of DV360 partners.
'dataset': '', # BigQuery dataset to write tables for each entity.
}
print("Parameters Set To: %s" % FIELDS)
# + [markdown] id="406c3a58-009" colab_type="text"
# #5. Execute DV360 Entity Read Files To BigQuery
# This does NOT need to be modified unless you are changing the recipe, click play.
#
# + id="406c3a58-010" colab_type="code"
from starthinker.util.configuration import Configuration
from starthinker.util.configuration import commandline_parser
from starthinker.util.configuration import execute
from starthinker.util.recipe import json_set_fields
USER_CREDENTIALS = '/content/user.json'
TASKS = [
{
'dataset': {
'auth': 'user',
'dataset': {'field': {'name': 'dataset','kind': 'string','order': 3,'default': '','description': 'BigQuery dataset to write tables for each entity.'}}
}
},
{
'entity': {
'auth': 'user',
'prefix': 'Entity',
'entities': [
'Campaign',
'LineItem',
'Creative',
'UserList',
'Partner',
'Advertiser',
'InsertionOrder',
'Pixel',
'InventorySource',
'CustomAffinity',
'UniversalChannel',
'UniversalSite',
'SupportedExchange',
'DataPartner',
'GeoLocation',
'Language',
'DeviceCriteria',
'Browser',
'Isp'
],
'partners': {
'single_cell': True,
'values': {'field': {'name': 'partners','kind': 'integer_list','order': 1,'default': '[]','description': 'Comma sparated list of DV360 partners.'}}
},
'out': {
'bigquery': {
'auth': 'user',
'dataset': {'field': {'name': 'dataset','kind': 'string','order': 3,'default': '','description': 'BigQuery dataset to write tables for each entity.'}}
}
}
}
}
]
json_set_fields(TASKS, FIELDS)
execute(Configuration(project=CLOUD_PROJECT, client=CLIENT_CREDENTIALS, user=USER_CREDENTIALS, verbose=True), TASKS, force=True)
| colabs/entity.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6.4 64-bit
# metadata:
# interpreter:
# hash: 5c4d2f1fdcd3716c7a5eea90ad07be30193490dd4e63617705244f5fd89ea793
# name: python3
# ---
# ### What is GridSearch?
# GridSearch is an optimization tool that we use when tuning hyperparameters. We define the grid of parameters that we want to search through, and we select the best combination of parameters for our data.
#
# https://towardsdatascience.com/gridsearch-the-ultimate-machine-learning-tool-6cd5fb93d07
# ### The “Grid” in GridSearch
#
# 
# # 1: One way
# +
import warnings
warnings.filterwarnings("ignore", category=DeprecationWarning)
# + tags=[]
from sklearn import svm, datasets
from sklearn.model_selection import GridSearchCV
iris = datasets.load_iris()
# 'kernel':('linear', 'poly', 'rbf', 'sigmoid'),
parameters = {
'kernel': ('linear', 'rbf', 'sigmoid'),
'C':[0.0001,0.1, 0.5, 1, 5, 10, 100],
'degree': [1,2,3,4,5,6,7,8,9],
'coef0': [-10.,-1., 0., 0.1, 0.5, 1, 10, 100],
'gamma': ('scale', 'auto')
}
svc = svm.SVC()
clf = GridSearchCV(estimator=svc, param_grid=parameters, n_jobs=-1, cv=10)
X = iris.data
y = iris.target
clf.fit(X, y)
print("clf.best_stimator_", clf.best_estimator_)
print("clf.best_params_", clf.best_params_)
# Mean cross-validated score of the best_estimator
print("clf.best_score", clf.best_score_)
# -
# # 2: Almost-Pro way
#
# La forma pro es la que hace esto mismo y va recogiendo los errores de entrenamiento, de validación y tiene la capacidad de parar el proceso cuando se requiera además de guardar el modelo en local una vez terminado si es mejor que el que había anteriormente y de cargar el modelo anterior y seguir reentrenando.
import pickle
# Load libraries
import numpy as np
from sklearn import datasets
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import GridSearchCV
from sklearn.pipeline import Pipeline
from sklearn.model_selection import RepeatedKFold
from sklearn.model_selection import train_test_split
# Set random seed
np.random.seed(0)
# Load data
iris = datasets.load_iris()
X = iris.data
y = iris.target
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=2)
to_test = np.arange(1, 10)
# +
# Create a pipeline
# Le podemos poner cualquier clasificador. Irá cambiando según va probando pero necesita 1.
pipe = Pipeline(steps=[('classifier', RandomForestClassifier())])
logistic_params = {
'classifier': [LogisticRegression()],
'classifier__penalty': ['l1', 'l2'],
'classifier__C': np.logspace(0, 4, 10)
}
random_forest_params = {
'classifier': [RandomForestClassifier()],
'classifier__n_estimators': [10, 100, 1000],
'classifier__max_features': [1, 2, 3]
}
svm_params = {
'classifier': [svm.SVC()],
'classifier__kernel':('linear', 'rbf', 'sigmoid'),
'classifier__C':[0.2, 0.3, 0.4, 0.5, 0.6, 0.7, 0.8, 0.9],
'classifier__degree': to_test,
'classifier__coef0': [-10.,-1., 0., 0.1, 0.5, 1, 10, 100],
'classifier__gamma': ('scale', 'auto')
}
# hypertuning
# Create space of candidate learning algorithms and their hyperparameters
search_space = [
logistic_params,
random_forest_params,
svm_params
]
# + tags=[]
# %%time
cv = RepeatedKFold(n_splits=10, n_repeats=1, random_state=1)
# Create grid search
clf = GridSearchCV(estimator=pipe, param_grid=search_space, cv=cv, verbose=0, n_jobs=-1)
# Fit grid search
best_model = clf.fit(X_train, y_train)
# View best model
separator = "\n############################\n"
print(separator)
print("best estimator:", best_model.best_estimator_.get_params()['classifier'])
print(separator)
print("clf.best_params_", clf.best_params_)
print(separator)
# Mean cross-validated score of the best_estimator
print("clf.best_score", clf.best_score_)
#SAVE MODEL
# save the model to disk
filename = 'finished_model.sav'
pickle.dump(best_model, open(filename, 'wb'))
# -
path = os.getcwd() + os.sep
full_file_name = path + "finished_model.sav"
loaded_model = pickle.load(open(full_file_name, 'rb'))
# Predict target vector
best_model.score(X_test, y_test) * 100
type(loaded_model)
# # 3 Another way - No pro
# + tags=[]
from sklearn import datasets
from sklearn.model_selection import train_test_split
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import classification_report
from sklearn.svm import SVC
# Loading the Digits dataset
digits = datasets.load_digits()
# To apply an classifier on this data, we need to flatten the image, to
# turn the data in a (samples, feature) matrix:
n_samples = len(digits.images)
X = digits.images.reshape((n_samples, -1))
y = digits.target
# -
0 0
0 0
# +
# Split the dataset in two equal parts
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.5, random_state=0)
# Set the parameters by cross-validation
tuned_parameters = [{'kernel': ['rbf'],
'gamma': [1e-3, 1e-4],
'C': [1, 10, 100, 1000]},
{'kernel': ['linear'],
'C': [1, 10, 100, 1000]}]
scores = ['precision', 'recall']
# + tags=[]
bar = "######################################"
for score in scores:
print(bar + "\n########## SCORE " + score + " ########### \n" + bar)
print("# Tuning hyper-parameters for %s" % score)
print()
clf = GridSearchCV(estimator=SVC(), param_grid=tuned_parameters, scoring=str(score)+'_macro')
clf.fit(X_train, y_train)
print("Best parameters set found on development set:")
print()
print(clf.best_params_)
print()
print("Grid scores on development set:")
print()
means = clf.cv_results_['mean_test_score']
stds = clf.cv_results_['std_test_score']
for mean, std, params in zip(means, stds, clf.cv_results_['params']):
print("%0.3f (+/-%0.03f) for %r"
% (mean, std * 2, params))
print("Detailed classification report:")
print()
print("The model is trained on the full development set.")
print("The scores are computed on the full evaluation set.")
print()
y_true, y_pred = y_test, clf.predict(X_test)
print(classification_report(y_true, y_pred))
print()
# -
| week9_ML_svm_poly_norm/day4_grid_save_model/theory/grid_search.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Jmol Jsmol applets working in Jupyter notebooks
#
# This notebook shows how to run Jmol/ Jsmol applets in Jupyter notebook a couple of ways.
#
# For the second one, using jupyter-jsmol, to work you have to run this notebook in sessions launched from [my repo with jupyter-jsmol listed](https://github.com/fomightez/jupyter-jsmol-binder) or [the jupyter-jsmol extension development repo](https://github.com/fekad/jupyter-jsmol).
#
# If not sorely using the jupyter-jsmol method:
# First, separately as another notebook on the same MyBinder session, I ran the first several cells of `Jmol served via a MyBinder remote session.ipynb` to get Jmol and place a directory `jsmol` in the root directory of the running session. **This step is important to put in place the jsmol directory with the javascript that the applet needs.**
# +
Old(?) versions of these steps are listed below for convenience.
```bash
# !curl -L -o Jmol-14.31.20-binary.tar.gz https://sourceforge.net/projects/jmol/files/Jmol/Version%2014.31/Jmol%2014.31.20/Jmol-14.31.20-binary.tar.gz/download
# !tar xzf Jmol-14.31.20-binary.tar.gz
# !unzip jmol-14.31.20/jsmol.zip
```
# -
# -----
#
# ### Applet and Jmol/Jsmol examples in Jupyter notebook cells
# I was hoping to get it to work in a notebook and found the original source of this code by searching 'simple jmol page' and seeing the 3rd one listed was on sourcrforge and had text saying 'HTML5', clicking on it took me to http://jmol.sourceforge.net/ which listed a link to demos under 'Samples' that took me to [Demonstration page](http://jmol.sourceforge.net/demo/) that had a link to 'Very simple page, with source code and explanations' at http://jmol.sourceforge.net/demo/jssample0.html ,but it didn't seem to work even when I edited it like below to get javascript from other places (but see next cell for more about applets encoded in notebook cell attempt that lead me to try this again):
# **(NEXT CELL DOESN'T WORK, ONES BELOW DO!! But detailing process so if something breaks can more easily troubleshoot)**
# %%HTML
<script src="http://jmol.sourceforge.net/jmol/JSmol.min.js" type="text/javascript"></script>
<script type="text/javascript">
var myJmol = 'myJmol';
var JmolInfo = {
width:'100%',
height:'100%',
color:'#E2F4F5',
serverURL:'https://chemapps.stolaf.edu/jmol/jsmol/php/jsmol.php',
use:'html5'
}
$(document).ready(function(){
$('#JmolDiv').html( Jmol.getAppletHtml(myJmol, JmolInfo) );
Jmol.script(myJmol, 'load $caffeine');
});
</script>
<div id="JmolDiv" style="width:30vmin; height:30vmin;"></div>
# Note the unedited (original source) version of that from http://jmol.sourceforge.net/demo/jssample0.html was:
# ```javascript
# <head>
# <script type="text/javascript">
# var myJmol = 'myJmol';
# var JmolInfo = {
# width:'100%',
# height:'100%',
# color:'#E2F4F5',
# j2sPath:'../jmol/j2s',
# serverURL:'../jmol/php/jsmol.php',
# use:'html5'
# }
# $(document).ready(function(){
# $('#JmolDiv').html( Jmol.getAppletHtml(myJmol, JmolInfo) );
# Jmol.script(myJmol, 'load ../model/caffeine.xyz');
# });
# </script>
# </head>
# <body>
# <div id="JmolDiv" style="width:30vmin; height:30vmin;"></div>
# </body>
# ```
# So after that failure I showed following procedure and code in `trying Jmol applet in a Jupyter notebook.ipynb` that I could at least get Jsmol code to run inside jupyter's `%%HTML` magics. It had issues though that it took over the page and oddly didn't load what it was supposed to and when it did it was zoomed out invisible and couldn't be moved by mouse.
#
# So rexamined the code that I used `trying Jmol applet in a Jupyter notebook.ipynb` (from https://stackoverflow.com/a/44995828/8508004 ) in comparison to the simple code I was trying in the notebook **because I knew I got a white space** and because of effort with `trying Jmol applet in a Jupyter notebook.ipynb`, I sort of wondered if molecule was there but too zoomed out to see. Maybe the javascript set up that at least causes something to work could inform me and I could get the code pasted in working. Worth a shot. So I edited above again, this time tried following and it worked with the adpatations based on code from https://stackoverflow.com/a/44995828/8508004 that almost worked but took over page:
# %%HTML
<script type="text/javascript" src="jsmol/JSmol.min.js"></script>
<script type="text/javascript">
var myJmol = 'myJmol';
var JmolInfo = {
width:'100%',
height:'100%',
color:'#E2F4F5',
j2sPath: "jsmol/j2s",
use:'html5'
}
$(document).ready(function(){
$('#JmolDiv').html( Jmol.getAppletHtml(myJmol, JmolInfo) );
Jmol.script(myJmol, 'load $caffeine');
});
</script>
<div id="JmolDiv" style="width:30vmin; height:30vmin;"></div>
# I don't honestly even know where it is getting caffeine molecule from because I don't see cafffein listed in `jsmol` directory, but very pleased to see **IT WORKS** and can spin caeffeine molecule with mouse in notebook cell.
# (Note that given https://chemapps.stolaf.edu/jmol/jsmol/jsmolgl.htm I think it means the load with dollar sign defaults to getting structure details from NCI and so this would mean this applet running in the cell can load structures from other places on web, too.)
#
# I'm noticing though that I cannot fully use console it seems. For example, I was trying to type in `set antialiasDisplay` and as I started typing with the `s` it kept triggering a notebook save step and not prinitng the `s` but prinint just `et`. Still major progress that this simple code works in the notebook.
# Note that WebGL can be used, too, based on https://chemapps.stolaf.edu/jmol/jsmol/jsmolgl.htm to get **better lighting**:
# %%HTML
<script type="text/javascript" src="jsmol/JSmol.min.js"></script>
<script type="text/javascript" src="jsmol/JSmol.GLmol.min.js"></script>
<script type="text/javascript">
var myJmol = 'myJmol';
var JmolInfo = {
width:'100%',
height:'100%',
color:'#E2F4F5',
j2sPath: "jsmol/j2s",
use: "WEBGL HTML5",
}
$(document).ready(function(){
$('#JmolDiv').html( Jmol.getAppletHtml(myJmol, JmolInfo) );
Jmol.script(myJmol, 'load $caffeine;set antialiasDisplay;');
});
</script>
<div id="JmolDiv" style="width:30vmin; height:30vmin;"></div>
# Note WebGL doesn't render the background color.
#
# Can use `=` in front of PDB id to to fetch a structure from PDB/RCSB online as illustrated [here](https://chemapps.stolaf.edu/jmol/jsmol/jsmolgl.htm). Trying that with some other variations. (Do I need to clear output first or can multiple run on page?)
# %%HTML
<script type="text/javascript" src="jsmol/JSmol.min.js"></script>
<script type="text/javascript">
var myJmol = 'myJmol';
var JmolInfo = {
width:'100%',
height:'100%',
color:'#E2F4F5',
j2sPath: "jsmol/j2s",
use: "html5",
}
$(document).ready(function(){
$('#JmolDivProtein').html( Jmol.getAppletHtml(myJmol, JmolInfo) );
Jmol.script(myJmol, 'load =1ehz; spacefill off; wireframe off; backbone off; cartoon on; color structure; set antialiasDisplay; moveto 0.0 { -667 -744 -36 75.21} 100.0 0.0 0.0 {60.993 51.431 25.1785} 51.865986946675356 {0 0 0} 0 0 0 3.0 0.0 0.0;');
});
</script>
<div id="JmolDivProtein" style="width:400px; height:400px;"></div>
# First try moving script location to be like source at https://chemapps.stolaf.edu/jmol/jsmol/jsmolgl.htm because I think script farther on left will be easier to edit.
# %%HTML
<script type="text/javascript" src="jsmol/JSmol.min.js"></script>
<script type="text/javascript">
var myJmol = 'myJmol';
var JmolInfo = {
width:'100%',
height:'100%',
color:'#E2F4F5',
j2sPath: "jsmol/j2s",
use: "html5",
script: 'load =1ehz; spacefill off; wireframe off; backbone off; cartoon on; color structure; set antialiasDisplay; moveto 0.0 { -667 -744 -36 75.21} 100.0 0.0 0.0 {60.993 51.431 25.1785} 51.865986946675356 {0 0 0} 0 0 0 3.0 0.0 0.0;',
}
$(document).ready(function(){
$('#JmolDivProtein').html( Jmol.getAppletHtml(myJmol, JmolInfo) );
});
</script>
<div id="JmolDivProtein" style="width:400px; height:400px;"></div>
# Can WebGL be used with proteins, too?
# %%HTML
<script type="text/javascript" src="jsmol/JSmol.min.js"></script>
<script type="text/javascript" src="jsmol/JSmol.GLmol.min.js"></script>
<script type="text/javascript">
var myJmol = 'myJmol';
var JmolInfo = {
width:'100%',
height:'100%',
color:'#E2F4F5',
j2sPath: "jsmol/j2s",
use: "WEBGL HTML5",
script: 'load =1ehz; spacefill off; wireframe off; backbone off; cartoon on; color structure; set antialiasDisplay; moveto 0.0 { -667 -744 -36 75.21} 100.0 0.0 0.0 {60.993 51.431 25.1785} 51.865986946675356 {0 0 0} 0 0 0 3.0 0.0 0.0;',
}
$(document).ready(function(){
$('#JmolDivProteinGL').html( Jmol.getAppletHtml(myJmol, JmolInfo) );
});
</script>
<div id="JmolDivProteinGL" style="width:400px; height:400px;"></div>
# ### Amazing jupyter-jsmol
#
# (I had a whole folder on my computer wondering how to do this and I had even contact <NAME> to floar the idea. I need to update my own documents in light of finding this.)
#
# Searching 'jsmol jupyter notebook' to hopefully find source of simple applet I had tried in notebook, I saw:
#
# https://pypi.org/project/jupyter-jsmol/
#
# Under Project Links on the right, is a link to the 'Homepage'
# which takes you to:
# https://github.com/fekad/jupyter-jsmol
#
# And it has a `launch binder` badge that launched where it seems to work!!! Fairly recent commits from 4 months ago when I found it January 2021.
# +
# #%pip install ipywidgets
# #%pip install jupyter_jsmol
# -
from jupyter_jsmol import JsmolView
from ipywidgets import Layout, widgets, interact
# !curl -OL https://raw.githubusercontent.com/fekad/jupyter-jsmol/master/examples/data/c2h410.xyz
view1 = JsmolView.from_file("c2h410.xyz")
# view1 = JsmolView.from_file("data/c2h410.xyz", inline=False)
view1
view1.close()
view2 = JsmolView.from_file("c2h410.xyz", inline=True)
view2
view2.close()
view5.close_all()
# +
import time
def executeSomething():
#code here
print ('.')
time.sleep(480) #60 seconds times 8 minutes
while True:
executeSomething()
# -
| Jmol Jsmol applets working in Jupyter notebooks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.5 64-bit (''base'': conda)'
# language: python
# name: python3
# ---
import pandas as pd
# +
df1 = pd.read_csv("./outputs/dumbbell_improvement/exp/acc.csv")
df1['NonIID'] = df1['Bucketing'].apply(lambda x: True)
df2 = pd.read_csv("./outputs/dumbbell/exp/acc.csv")
df2['Bucketing'] = df2['Agg'].apply(lambda x: False)
df2['RandomEdge'] = df2['Agg'].apply(lambda x: False)
df2 = df2[~df2['NonIID']]
df = pd.concat([df1, df2])
df = df[df['Group'] == 'clique 1']
# +
import matplotlib.pyplot as plt
import seaborn as sns
plt.rcParams["font.family"] = "Times New Roman"
plt.rcParams["font.size"] = 2
plt.rcParams["legend.columnspacing"] = 0.5
plt.rcParams["legend.handlelength"] = 1
plt.rcParams["legend.borderaxespad"] = 0
plt.rcParams["legend.labelspacing"] = 0.1
plt.rcParams["legend.frameon"] = False
sns.set()
aggregators = ['SCClip', 'GM', 'MOZI', 'TM', 'Gossip']
colors = sns.color_palette()[:len(aggregators)]
fig, axes = plt.subplots(nrows=1, ncols=5, figsize=(12, 2), sharey=True)
def plot_iid(df, ax):
for agg, c in zip(aggregators, colors):
_df = df[df['Agg'] == agg]
ax.plot(_df['Iterations'], _df['Accuracy (%)'], color=c, label=agg)
ax.legend(loc='lower left', bbox_to_anchor=[0.1, 0.])
ax.set_xlim(0, 750)
ax.set_xlabel("Iterations")
ax.set_title("IID")
plot_iid(df[~df['NonIID']], axes[0])
def plot_noniid(df, ax):
for agg, c in zip(aggregators, colors):
_df = df[df['Agg'] == agg]
ax.plot(_df['Iterations'], _df['Accuracy (%)'], color=c, label=agg)
ax.set_xlim(0, 750)
ax.set_xlabel("Iterations")
ax.set_title("NonIID")
plot_noniid(df[(df['NonIID']) & (~df['Bucketing']) & (~df['RandomEdge'])], axes[1])
def plot_bucketing(df, ax):
for agg, c in zip(aggregators, colors):
_df = df[df['Agg'] == agg]
ax.plot(_df['Iterations'], _df['Accuracy (%)'], color=c, label=agg)
ax.set_xlim(0, 750)
ax.set_xlabel("Iterations")
ax.set_title("NonIID + B.")
plot_bucketing(df[(df['NonIID']) & (df['Bucketing']) & (~df['RandomEdge'])], axes[2])
def plot_edge(df, ax):
for agg, c in zip(aggregators, colors):
_df = df[df['Agg'] == agg]
ax.plot(_df['Iterations'], _df['Accuracy (%)'], color=c, label=agg)
ax.set_xlim(0, 750)
ax.set_xlabel("Iterations")
ax.set_title("NonIID + R.")
plot_edge(df[(df['NonIID']) & (~df['Bucketing']) & (df['RandomEdge'])], axes[3])
def plot_both(df, ax):
for agg, c in zip(aggregators, colors):
_df = df[df['Agg'] == agg]
ax.plot(_df['Iterations'], _df['Accuracy (%)'], color=c, label=agg)
ax.set_xlim(0, 750)
ax.set_xlabel("Iterations")
ax.set_title("NonIID + B. + R.")
plot_both(df[(df['NonIID']) & (df['Bucketing']) & (df['RandomEdge'])], axes[4])
axes[0].set_ylim(45, 100)
axes[0].set_yticks([50, 60, 70, 80, 90, 100])
axes[0].set_ylabel('Accuracy (%)', labelpad=-2)
for i in range(5):
axes[i].tick_params(axis='y', which='major', pad=-4)
axes[i].tick_params(axis='x', which='major', pad=-2)
fig.subplots_adjust(wspace=0.093)
fig.savefig("./dumbbell_acc.pdf", bbox_inches="tight", dpi=720)
# -
| dumbbell_plot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
import pandas as pd
import matplotlib.pyplot as plt
from functools import reduce
import os
plt.rcParams['figure.figsize'] = (10.0, 8.0)
plt.style.use("ggplot")
# %matplotlib inline
d_f = pd.read_csv("out_data/chicago_features.csv")
'''
define function to make plot easier, for different purpose just modify the according lines
'''
def plot_func(names, picname):
loc_df_ = d_f[[names, "Primary Type"]].groupby("Primary Type")
dicts = {}
def func(d, data):
d.setdefault(data[0], data[1][0])
return d
for crimes in loc_df["Primary Type"].unique():
temp_df = loc_df_.get_group(crimes).groupby(names).count().sort("Primary Type", ascending=False)
tol = temp_df.sum().values[0]
most_name = temp_df.index.values # moidfy this line and related lines to get the full value
most_num = temp_df.values/tol*100
maplist = zip(most_name, most_num)
dicts.setdefault(crimes, {})
t_dict = dict(reduce(func, maplist, {}))
dicts[crimes] = t_dict
loc_ = pd.DataFrame.from_dict(dicts, orient="columns") # also change the orient for different purpose
ax = loc_.plot(figsize=(15,10), rot=-45) ## modify this line to get the different kind of plot
ax.set_xlabel("day of week")
ax.set_ylabel("percentage(%)")
ax.set_xticklabels(["Mon", "Tue", "Wen", "Thu", "Fri", "Sat", "Sun"]) #keep or remove this line
plt.savefig(os.path.join("picture/", picname, ".png"))
plot_func("Month", "month_plot")
| Descriptive_plot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.10 64-bit
# name: python3
# ---
# sorting algorithms
from sort_me import sort
# function to measure time
from time import time
# graph plotting library
from matplotlib import pyplot as plt
# random number generation
from random import choices
# CSV files handling
import csv
# Data analysis
import pandas as pd
# CSV files handling
import csv
# Path handling
from os import path
import sys
sys.setrecursionlimit(1000000)
LIST_SIZES = [1e1, 1e2, 1e3, 1e4]
with open('sorting_data.csv', 'w') as csv_file:
field_names = ['algorithm', 'entries', 'time']
csv_writer = csv.DictWriter(csv_file, fieldnames=field_names)
csv_writer.writeheader()
for size in LIST_SIZES:
random_list = choices(range(1000), k=int(size))
time_bubble = time()
_ = sort.bubble(random_list)
time_bubble = time() - time_bubble
time_merge = time()
_ = sort.merge(random_list)
time_merge = time() - time_merge
time_quick = time()
_ = sort.quick(random_list)
time_quick = time() - time_quick
csv_writer.writerow({'algorithm': 'bubble', 'entries': int(size), 'time': time_bubble})
csv_writer.writerow({'algorithm': 'merge', 'entries': int(size), 'time': time_merge})
csv_writer.writerow({'algorithm': 'quick', 'entries': int(size), 'time': time_quick})
# +
data = pd.read_csv(path.join(path.curdir, "sorting_data.csv"))
bubble = data.loc[data['algorithm'] == 'bubble', ['entries', 'time']].set_index('entries').rename(columns={'time':'Bubble sort'})
merge = data.loc[data['algorithm'] == 'merge', ['entries', 'time']].set_index('entries').rename(columns={'time':'Merge sort'})
quick = data.loc[data['algorithm'] == 'quick', ['entries', 'time']].set_index('entries').rename(columns={'time':'Quick sort'})
data = pd.concat([bubble, merge, quick], axis=1)
# -
plt.style.use('seaborn')
data.plot(
style="o-",
loglog=True,
x_compat=True,
xlabel="Quantidade de elementos",
ylabel="Tempo(s)",
title="Algoritmos de ordenação")
| main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# ### Exmaple of Testing Difference Between Two Samples
#
# * Are the hospital inpatient charge data similar between HI And DC
# * we will compare the "Average Total Payments"
# * the numberr of providers is simialr between both state
#
# * Complete dataset can be obtained here:
# https://data.cms.gov/Medicare-Inpatient/Inpatient-Prospective-Payment-System-IPPS-Provider/97k6-zzx3
# + slideshow={"slide_type": "slide"}
import pandas as pd
data = pd.read_csv("data/hospital_charge_data.csv")
data.head()
# + slideshow={"slide_type": "slide"}
payment_data = data[['Provider State', ' Average Total Payments ']]
payment_data.columns = ['state', 'avg_payments']
payment_data = payment_data[(payment_data["state"] == "DC") | (payment_data["state"] == "HI")]
payment_data.sample(10)
# + slideshow={"slide_type": "slide"}
payment_data.groupby("state").mean()
# -
plt.hist(payment_data[payment_data["state"] == "HI"]["avg_payments"])
plt.hist(payment_data[payment_data["state"] == "DC"]["avg_payments"])
# ### Hypothesis Testing
#
# * How similar are the hospital inpatient charges in Hawaii and DC?
#
# * Null Hypothesis ($H_0$)
# * The mean values across both state are the same and any difference is due to sampling differences
# * The values we would observe if we were to sample HI and compare it to itself would be different
#
#
# * Alternative Hypothesis ($H_A$)
# * The mean values across both state are different
# * The observed difference is not due to chance
#
#
#
| morea/parameter_estimation/resources/.ipynb_checkpoints/Hospital_Payments_Hawaii_DC-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %pwd
# %load_ext autoreload
# %autoreload 2
import pandas as pd
from datetime import datetime
from radiosonde.loader.igra2.igra2_loader import Igra2SondeLoader
start_date = datetime(2020, 9, 12, 0, 0, 0)
end_date = datetime(2018, 9, 14)
site_id_list = ['CAM00071802']
dates = pd.date_range("2020-06-20", "2020-10-20", freq='12H').to_pydatetime()
loader = Igra2SondeLoader()
sondes = loader.load_many(dates)
| notebooks/igra2-test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Algorithms before feature selection
## Import the required libraries
import pandas as pd
import numpy as np
import imblearn
from imblearn.pipeline import make_pipeline as make_pipeline_imbfinal
from imblearn.over_sampling import SMOTE
from imblearn.metrics import classification_report_imbalanced
from sklearn.metrics import precision_score, recall_score, f1_score, roc_auc_score, accuracy_score, classification_report
from sklearn.cross_validation import train_test_split
from sklearn.ensemble import RandomForestClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.naive_bayes import BernoulliNB
#from sklearn import svm
from sklearn.metrics import *
import pickle
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
df = pd.read_csv('ckd.csv')
df.head()
df_train,df_test = train_test_split(df,train_size=0.7,random_state=42)
drop_col_list=['RBC_normal','PCC_present','BA_present','CAD_yes','Ane_yes' , 'PC_normal' , 'Appet_good' ,
'age' , 'pot' , 'wc' , 'PE_yes' , 'bp' , 'su' ]
x_train=df_train.iloc[:,:24]
x_train.drop(drop_col_list,axis=1,inplace=True)
y_train=df_train['Classification_ckd']
scaler.fit(x_train)
x_train_sc=scaler.transform(x_train)
x_test=df_test.iloc[:,:24]
x_test.drop(drop_col_list,axis=1,inplace=True)
y_test=df_test['Classification_ckd']
scaler.fit(x_test)
x_test_sc=scaler.transform(x_test)
accuracy =[]
model_name =[]
dataset=[]
f1score = []
precision = []
recall = []
true_positive =[]
false_positive =[]
true_negative =[]
false_negative =[]
# # Logistic Regression
logreg = LogisticRegression()
## fitiing the model
logreg.fit(x_train_sc, y_train)
filename = 'logReg_model.sav'
pickle.dump(logreg,open(filename,'wb'))
logreg
# Training
prediction = logreg.predict(x_train_sc)
f1 = f1_score(y_train, prediction)
p = precision_score(y_train, prediction)
r = recall_score(y_train, prediction)
a = accuracy_score(y_train, prediction)
cm = confusion_matrix(y_train, prediction)
tp = cm[0][0]
fp = cm[0][1]
fn = cm[1][0]
tn = cm[1][1]
dataset.append('Training')
model_name.append('Logistic Regression')
f1score.append(f1)
precision.append(p)
recall.append(r)
accuracy.append(a)
true_positive.append(tp)
false_positive.append(fp)
true_negative.append(tn)
false_negative.append(fn)
cm
# Testing
prediction = logreg.predict(x_test_sc)
f1 = f1_score(y_test, prediction)
p = precision_score(y_test, prediction)
r = recall_score(y_test, prediction)
a = accuracy_score(y_test, prediction)
cm = confusion_matrix(y_test, prediction)
tp = cm[0][0]
fp = cm[0][1]
fn = cm[1][0]
tn = cm[1][1]
model_name.append('Logistc Regression')
dataset.append('Testing')
f1score.append(f1)
precision.append(p)
recall.append(r)
accuracy.append(a)
true_positive.append(tp)
false_positive.append(fp)
true_negative.append(tn)
false_negative.append(fn)
cm
# # Bernoulli Naive Bayes
NB = BernoulliNB()
## fitiing the model
NB.fit(x_train_sc, y_train)
filename = 'BernoulliNB_model.sav'
pickle.dump(NB,open(filename,'wb'))
NB
# Training
prediction = NB.predict(x_train_sc)
f1 = f1_score(y_train, prediction)
p = precision_score(y_train, prediction)
r = recall_score(y_train, prediction)
a = accuracy_score(y_train, prediction)
cm = confusion_matrix(y_train, prediction)
tp = cm[0][0]
fp = cm[0][1]
fn = cm[1][0]
tn = cm[1][1]
model_name.append('Bernoulli Naive Bayes')
dataset.append('Training')
f1score.append(f1)
precision.append(p)
recall.append(r)
accuracy.append(a)
true_positive.append(tp)
false_positive.append(fp)
true_negative.append(tn)
false_negative.append(fn)
cm
# Testing
prediction = NB.predict(x_test_sc)
f1 = f1_score(y_test, prediction)
p = precision_score(y_test, prediction)
r = recall_score(y_test, prediction)
a = accuracy_score(y_test, prediction)
cm = confusion_matrix(y_test, prediction)
tp = cm[0][0]
fp = cm[0][1]
fn = cm[1][0]
tn = cm[1][1]
model_name.append('Bernoulli Naive Bayes')
dataset.append('Testing')
f1score.append(f1)
precision.append(p)
recall.append(r)
accuracy.append(a)
true_positive.append(tp)
false_positive.append(fp)
true_negative.append(tn)
false_negative.append(fn)
cm
# # Random Forest Classifier
rfc = RandomForestClassifier(n_estimators=50,random_state=0)
## fitiing the model
rfc.fit(x_train_sc, y_train)
filename = 'RFC_model.sav'
pickle.dump(rfc,open(filename,'wb'))
rfc
# Training
prediction = rfc.predict(x_train_sc)
f1 = f1_score(y_train, prediction)
p = precision_score(y_train, prediction)
r = recall_score(y_train, prediction)
a = accuracy_score(y_train, prediction)
cm = confusion_matrix(y_train, prediction)
tp = cm[0][0]
fp = cm[0][1]
fn = cm[1][0]
tn = cm[1][1]
model_name.append('Random Forest Classifier')
dataset.append('Training')
f1score.append(f1)
precision.append(p)
recall.append(r)
accuracy.append(a)
true_positive.append(tp)
false_positive.append(fp)
true_negative.append(tn)
false_negative.append(fn)
cm
# Testing
prediction = rfc.predict(x_test_sc)
f1 = f1_score(y_test, prediction)
p = precision_score(y_test, prediction)
r = recall_score(y_test, prediction)
a = accuracy_score(y_test, prediction)
cm = confusion_matrix(y_test, prediction)
tp = cm[0][0]
fp = cm[0][1]
fn = cm[1][0]
tn = cm[1][1]
model_name.append('Random Forest Classifier')
dataset.append('Testing')
f1score.append(f1)
precision.append(p)
recall.append(r)
accuracy.append(a)
true_positive.append(tp)
false_positive.append(fp)
true_negative.append(tn)
false_negative.append(fn)
cm
# # k Nearest Neighbour
from sklearn.neighbors import KNeighborsClassifier
knn = KNeighborsClassifier(n_neighbors = 2)
knn.fit(x_train_sc, y_train)
filename = 'KNN_model.sav'
pickle.dump(knn,open(filename,'wb'))
knn
# Training
prediction = knn.predict(x_train_sc)
f1 = f1_score(y_train, prediction)
p = precision_score(y_train, prediction)
r = recall_score(y_train, prediction)
a = accuracy_score(y_train, prediction)
cm = confusion_matrix(y_train, prediction)
tp = cm[0][0]
fp = cm[0][1]
fn = cm[1][0]
tn = cm[1][1]
model_name.append('k nearest Neighbour')
dataset.append('Training')
f1score.append(f1)
precision.append(p)
recall.append(r)
accuracy.append(a)
true_positive.append(tp)
false_positive.append(fp)
true_negative.append(tn)
false_negative.append(fn)
cm
# Testing
prediction = knn.predict(x_test_sc)
f1 = f1_score(y_test, prediction)
p = precision_score(y_test, prediction)
r = recall_score(y_test, prediction)
a = accuracy_score(y_test, prediction)
cm = confusion_matrix(y_test, prediction)
tp = cm[0][0]
fp = cm[0][1]
fn = cm[1][0]
tn = cm[1][1]
model_name.append('k nearest Neighbour')
dataset.append('Testing')
f1score.append(f1)
precision.append(p)
recall.append(r)
accuracy.append(a)
true_positive.append(tp)
false_positive.append(fp)
true_negative.append(tn)
false_negative.append(fn)
cm
# # Support Vector Classifier
from sklearn.svm import SVC
svc = SVC (kernel = 'linear' , C = 0.025 , random_state = 42)
svc.fit(x_train_sc, y_train)
filename = 'SVC_model.sav'
pickle.dump(svc,open(filename,'wb'))
svc
# Training
prediction = svc.predict(x_train_sc)
f1 = f1_score(y_train, prediction)
p = precision_score(y_train, prediction)
r = recall_score(y_train, prediction)
a = accuracy_score(y_train, prediction)
cm = confusion_matrix(y_train, prediction)
tp = cm[0][0]
fp = cm[0][1]
fn = cm[1][0]
tn = cm[1][1]
model_name.append('Support Vector Classifier')
dataset.append('Training')
f1score.append(f1)
precision.append(p)
recall.append(r)
accuracy.append(a)
true_positive.append(tp)
false_positive.append(fp)
true_negative.append(tn)
false_negative.append(fn)
cm
# Testing
prediction = svc.predict(x_test_sc)
f1 = f1_score(y_test, prediction)
p = precision_score(y_test, prediction)
r = recall_score(y_test, prediction)
a = accuracy_score(y_test, prediction)
cm = confusion_matrix(y_test, prediction)
tp = cm[0][0]
fp = cm[0][1]
fn = cm[1][0]
tn = cm[1][1]
model_name.append('Support Vector Classifier')
dataset.append('Testing')
f1score.append(f1)
precision.append(p)
recall.append(r)
accuracy.append(a)
true_positive.append(tp)
false_positive.append(fp)
true_negative.append(tn)
false_negative.append(fn)
cm
# # Stochastic Gradient Decent
from sklearn.linear_model import SGDClassifier
sgd = SGDClassifier (loss = 'modified_huber' , shuffle = True , random_state = 42)
sgd.fit(x_train_sc, y_train)
filename = 'SGD_model.sav'
pickle.dump(sgd,open(filename,'wb'))
sgd
# Training
prediction = sgd.predict(x_train_sc)
f1 = f1_score(y_train, prediction)
p = precision_score(y_train, prediction)
r = recall_score(y_train, prediction)
a = accuracy_score(y_train, prediction)
cm = confusion_matrix(y_train, prediction)
tp = cm[0][0]
fp = cm[0][1]
fn = cm[1][0]
tn = cm[1][1]
model_name.append('Stochastic Gradient Decent')
dataset.append('Training')
f1score.append(f1)
precision.append(p)
recall.append(r)
accuracy.append(a)
true_positive.append(tp)
false_positive.append(fp)
true_negative.append(tn)
false_negative.append(fn)
cm
# Testing
prediction = sgd.predict(x_test_sc)
f1 = f1_score(y_test, prediction)
p = precision_score(y_test, prediction)
r = recall_score(y_test, prediction)
a = accuracy_score(y_test, prediction)
cm = confusion_matrix(y_test, prediction)
tp = cm[0][0]
fp = cm[0][1]
fn = cm[1][0]
tn = cm[1][1]
model_name.append('Stochastic Gradient Decent')
dataset.append('Testing')
f1score.append(f1)
precision.append(p)
recall.append(r)
accuracy.append(a)
true_positive.append(tp)
false_positive.append(fp)
true_negative.append(tn)
false_negative.append(fn)
cm
# # Gradient Naive Bayes
from sklearn.naive_bayes import GaussianNB
gnb = GaussianNB()
gnb.fit(x_train_sc, y_train)
filename = 'GNB_model.sav'
pickle.dump(gnb,open(filename,'wb'))
gnb
# Training
prediction = gnb.predict(x_train_sc)
f1 = f1_score(y_train, prediction)
p = precision_score(y_train, prediction)
r = recall_score(y_train, prediction)
a = accuracy_score(y_train, prediction)
cm = confusion_matrix(y_train, prediction)
tp = cm[0][0]
fp = cm[0][1]
fn = cm[1][0]
tn = cm[1][1]
model_name.append('Gradient Naive Bayes')
dataset.append('Training')
f1score.append(f1)
precision.append(p)
recall.append(r)
accuracy.append(a)
true_positive.append(tp)
false_positive.append(fp)
true_negative.append(tn)
false_negative.append(fn)
cm
# Testing
prediction = gnb.predict(x_test_sc)
f1 = f1_score(y_test, prediction)
p = precision_score(y_test, prediction)
r = recall_score(y_test, prediction)
a = accuracy_score(y_test, prediction)
cm = confusion_matrix(y_test, prediction)
tp = cm[0][0]
fp = cm[0][1]
fn = cm[1][0]
tn = cm[1][1]
model_name.append('Gradient Naive Bayes')
dataset.append('Testing')
f1score.append(f1)
precision.append(p)
recall.append(r)
accuracy.append(a)
true_positive.append(tp)
false_positive.append(fp)
true_negative.append(tn)
false_negative.append(fn)
cm
# # Writing summary metrics
Summary = model_name,dataset,f1score,precision,recall,accuracy,true_positive,false_positive,true_negative,false_negative
#Summary
## Making a dataframe of the accuracy and error metrics
describe1 = pd.DataFrame(Summary[0],columns = {"Model_Name "})
describe2 = pd.DataFrame(Summary[1],columns = {"Dataset"})
describe3 = pd.DataFrame(Summary[2],columns = {"F1_score"})
describe4 = pd.DataFrame(Summary[3],columns = {"Precision_score"})
describe5 = pd.DataFrame(Summary[4],columns = {"Recall_score"})
describe6 = pd.DataFrame(Summary[5], columns ={"Accuracy_score"})
describe7 = pd.DataFrame(Summary[6], columns ={"True_Positive"})
describe8 = pd.DataFrame(Summary[7], columns ={"False_Positive"})
describe9 = pd.DataFrame(Summary[8], columns ={"True_Negative"})
describe10 = pd.DataFrame(Summary[9], columns ={"False_Negative"})
des = describe1.merge(describe2, left_index=True, right_index=True, how='inner')
des = des.merge(describe3,left_index=True, right_index=True, how='inner')
des = des.merge(describe4,left_index=True, right_index=True, how='inner')
des = des.merge(describe5,left_index=True, right_index=True, how='inner')
des = des.merge(describe6,left_index=True, right_index=True, how='inner')
des = des.merge(describe7,left_index=True, right_index=True, how='inner')
des = des.merge(describe8,left_index=True, right_index=True, how='inner')
des = des.merge(describe9,left_index=True, right_index=True, how='inner')
des = des.merge(describe10,left_index=True, right_index=True, how='inner')
#des = des.merge(describe9,left_index=True, right_index=True, how='inner')
Summary_csv = des.sort_values(ascending=True,by="False_Negative").reset_index(drop = True)
Summary_csv
Summary_csv.to_csv('Summary.csv')
| Kidney Disease/Algorithms after feature selection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Creating colorful cars
#
# Now, the `car.py` file has been modified so that `__init__` takes in an optional color parameter!
#
# Your tasks for this notebook are:
# 1. Create multiple cars of different colors
# 2. Move them around and display the result
# 3. (Optional) Add another variable to __init__ like maximum_speed or a boolean true or false depending on if the car has good speakers. It's up to you!
#
# Your options for color values include:
# * b: blue
# * g: green
# * r: red
# * c: cyan
# * m: magenta
# * y: yellow
# * k: black
# * w: white
#
# More color info can be found, [here](https://matplotlib.org/api/colors_api.html).
# +
import numpy as np
import car
# %matplotlib inline
# Auto-reload function so that this notebook keeps up with
# changes in the class file
# %load_ext autoreload
# %autoreload 2
# -
# ### Define some initial variables
# +
# Create a 2D world of 0's
height = 4
width = 6
world = np.zeros((height, width))
# Define the initial car state
initial_position = [0, 0] # [y, x] (top-left corner)
velocity = [0, 1] # [vy, vx] (moving to the right)
# -
## TODO: Create two cars of different colors and display their different worlds
car1 = car.Car(initial_position, velocity, world)
car2 = car.Car(initial_position, velocity, world, 'y')
car1.move()
car2.move()
car2.move()
car1.display_world()
car2.display_world()
# You can also check out one potential solution to this in the solution notebook, which can be found by clicking on "Jupyter" in the top left.
| CVND_Exercises/3_5_State_and_Motions/2. Multiple, Colorful Cars.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
import numpy as np
import os
from scipy.linalg import schur
import tensorflow as tf
from backend.attention_network import Model
import backend as be
from backend.simulation_tools import Simulator
from backend.weight_initializer import weight_initializer
from tasks import color_matching as cm
import matplotlib.pyplot as plt
import time
import pickle
# %matplotlib inline
# +
def softmax(data):
return np.exp(data).T/np.sum(np.exp(data),axis=1)
def relu(x):
return np.maximum(x,0)
def state_to_output(s,w):
o = np.zeros([s.shape[0],s.shape[1],w['b_out'].shape[0]])
for ii in range(s.shape[1]):
o[:,ii,:] = relu(s[:,ii,:]).dot(w['W_out'].T) + w['b_out']
return o
def get_rts(out,thresh=.4):
n_trials = out.shape[1]
n_steps = out.shape[0]
rts = -np.ones(n_trials).astype(int)
choice = np.zeros(n_trials)
for ii in range(n_trials):
cross = np.where(np.abs(out[:,ii,0]-out[:,ii,1])>thresh)
if len(cross[0]) != 0:
rts[ii] = np.min(cross[0])
choice[ii] = np.argmax(out[rts[ii],ii,:])
return rts,choice
def ols(X,Y,reg=0.):
X = np.hstack([X,np.ones([X.shape[0],1])])
w = np.linalg.inv(X.T.dot(X) + reg*np.eye(X.shape[1])).dot(X.T).dot(Y)
return w,X.dot(w)
def run_ols(X,w):
X = np.hstack([X,np.ones([X.shape[0],1])])
return X.dot(w)
# +
n_hidden = 250
tau = 100.0 #As double
dt = 20.0 #As double
dale_ratio = None
rec_noise = 0.1
stim_noise = 0.1
batch_size = 64
cohs = [.1,.2,.5,.7,1.]
rt_version = False
reward_version = False
#train params
learning_rate = .0001
training_iters = 200000
display_step = 20
'''curriculum trained:
cm_att_curr1 - with attention (400 ms stimuli epoch)
cm_att_rew_curr0 - with attention and rewards (400 ms stimuli epoch)
cm_att_curr2 - with attention (stimulus stays until end of trial)
cm_att_l2_curr0 - same as cm_att_curr2 with l2 penalty on attention
'''
tag = 'cm_att_l2_curr0'
weights_path = '../weights/'+tag+'.npz'
params_path = '../weights/'+tag+'_params.p'
params = cm.set_params(coherences=cohs,
stim_noise = stim_noise, rec_noise = rec_noise, L1_rec = 0,
L2_firing_rate = .2, sample_size = 128, epochs = 100, N_rec = n_hidden,
dale_ratio=dale_ratio, tau=tau, dt = dt, task='n_back',rt_version=rt_version,reward_version=reward_version)
generator = cm.generate_train_trials(params)
# -
#Attention Weight specific params
params['L2_attention'] = .002 #attention regularizer
params['tau_fw'] = 50. #initial time constant of attention update
params['gamma_train'] = True #trainable gamma (dt/tu_fw)
params['phi_train'] = False #trainable phi (sensitivity to fws)
params['global_phi'] = False #Single global or synapse specific plasticity
# +
trial = cm.build_train_trials(params)
for ii in range(4):
plt.subplot(4,1,ii+1)
plt.plot(trial[0][ii,:,:])
plt.plot(trial[1][ii,:,:])
plt.plot(trial[2][ii,:,:])
plt.show()
# -
trial[2].shape
# +
# output_weights_path = weights_path
# params['init_type'] = 'gaussian'
# 'external weight intializer class'
# autapses = True
# w_initializer = weight_initializer(params,output_weights_path[:-4] + '_init',autapses=autapses)
# input_weights_path = w_initializer.gen_weight_dict()
# params['load_weights_path'] = input_weights_path + '.npz'
# w_init = np.load(input_weights_path + '.npz')
# plt.imshow(w_init['W_rec'],interpolation='none')
# plt.figure()
# plt.plot(w_init['W_in'])
# plt.show()
# -
#regular training
if False:
tf.reset_default_graph()
model = Model(params)
sess = tf.Session()
t,att,phi,gamma,s,trial_data = model.train(sess, generator, learning_rate = learning_rate, training_iters = training_iters,
save_weights_path = weights_path, display_step=display_step,batch_size=batch_size)
sess.close()
#curriculum learning
if False:
coh_factors = np.array([1.,.8,.5,.3,.1]) #np.arange(1.,0,-.2)
params['load_weights_path'] = None
params['N_batch'] = params['sample_size'] = 128
training_iters = 300000
#train loop
for coh_factor in coh_factors:
print coh_factor
tf.reset_default_graph()
params['coherences'] = coh_factor*np.array(cohs)
if coh_factor == coh_factors[-1]:
training_iters = 300000
generator = cm.generate_train_trials(params)
model = Model(params)
sess = tf.Session()
t,att,phi,gamma,s,trial_data = model.train(sess, generator, learning_rate = learning_rate, training_iters = training_iters,
save_weights_path = weights_path, display_step=display_step,batch_size=batch_size)
sess.close()
params['load_weights_path'] = weights_path
training_iters = 100000
#save train params
pickle.dump( params, open( params_path, "wb" ) )
# +
#run null trials
tf.reset_default_graph()
params['coherences'] = np.array([0.])
params['N_batch'] = params['sample_size'] = 10000
params['load_weights_path'] = weights_path
generator = cm.generate_train_trials(params)
model = Model(params)
null_trials = generator.next()
sess = tf.Session()
sess.run(tf.global_variables_initializer())
att_null,s_null = sess.run([model.attention,model.states], feed_dict={model.x: null_trials[0]})
sess.close()
# +
w = np.load(weights_path)
att_null = np.asarray(att_null)
s_null = np.asarray(s_null)
o_null = state_to_output(s_null,w)
soft_att_null = np.zeros(att_null.shape)
for ii in range(soft_att_null.shape[1]):
soft_att_null[:,ii,:] = softmax(att_null[:,ii,:]).T
rts,choice = get_rts(o_null,thresh=.6)
plt.subplot(3,1,1)
plt.plot(soft_att_null[:,:,0])
plt.axvline(50,c='k',linestyle='--')
plt.axvline(90,c='k',linestyle='--')
plt.axvline(130,c='k',linestyle='--')
plt.subplot(3,1,2)
plt.plot(o_null[:,:,0])
plt.axvline(50,c='k',linestyle='--')
plt.axvline(90,c='k',linestyle='--')
plt.axvline(130,c='k',linestyle='--')
plt.subplot(3,1,3)
plt.hist(rts[rts>0],30)
plt.xlim([0,250])
plt.axvline(50,c='k',linestyle='--')
plt.axvline(90,c='k',linestyle='--')
plt.axvline(130,c='k',linestyle='--')
plt.show()
# +
# plt.hist(rts[np.logical_and(rts>0,rts<130)],30)
plt.figure(figsize=(10,6))
for ii in range(15):
plt.subplot(5,3,ii+1)
plt.imshow(relu(s_null[:,:,20+ii]).T,aspect='auto',interpolation='none',cmap='RdBu_r')
# plt.axvline(50,c='r',linestyle='--')
# plt.axvline(90,c='r',linestyle='--')
# plt.axvline(130,c='r',linestyle='--')
plt.tight_layout()
plt.show()
# +
#psychophysical kernel
plt.plot(np.mean(null_trials[0][choice==0],axis=0)[:,0])
plt.plot(np.mean(null_trials[0][choice==1],axis=0)[:,0])
plt.axvline(50,c='k',linestyle='--')
plt.axvline(90,c='k',linestyle='--')
plt.axvline(130,c='k',linestyle='--')
plt.show()
# +
M = np.zeros([10000,4])
M[:,0] = rts
M[:,1] = 0. #correct.astype('int')
M[:,2] = 0
M[:,3] = 1.
np.save('rnn_null_data',M)
# +
#run regular trials
tf.reset_default_graph()
params['coherences'] = np.array([.005,.01,.02,.03,.05,.08,.1])
params['N_batch'] = params['sample_size'] = 1000
params['load_weights_path'] = weights_path
generator = cm.generate_train_trials(params)
model = Model(params)
reg_trials = generator.next()
sess = tf.Session()
sess.run(tf.global_variables_initializer())
att,s = sess.run([model.attention,model.states], feed_dict={model.x: reg_trials[0]})
sess.close()
# +
# t,att,phi,gamma,s,trial_data = t
w = np.load(weights_path)
# print np.asarray(att).shape
# print t
# print s[0].shape
# print trial_data[0].shape
att_ = np.asarray(att)
s_ = np.asarray(s)
o_ = state_to_output(s_,w)
rts,choice = get_rts(o_,thresh=.8)
# rule = np.argmax(trial_data[0][:,50,:2],axis=1)
target = np.argmax(reg_trials[1][:,199,:],axis=1)
# choice = np.argmax(o_[-1,:,:],axis=1)
correct = choice == target
# coh = np.zeros(len(rule))
# for ii in range(len(rule)):
# coh[ii] = np.mean(trial_data[0][ii,100:180,rule[ii]+2],axis=0)
print('Accuracy: {}'.format(np.mean(correct)))
trial = 1
plt.subplot(3,1,1)
# plt.plot(trial_data[0][trial,:,:])
plt.subplot(3,1,2)
plt.plot(s_[:,trial,:])
plt.subplot(3,1,3)
plt.imshow(softmax(att_[:,trial,:]),aspect='auto',interpolation='none',cmap='Blues')
# plt.imshow((np.exp(att_[:,trial,:]).T/np.sum(np.exp(att_[:,trial,:]),axis=1)),aspect='auto',interpolation='none',cmap='Blues')
# plt.colorbar()
plt.legend(range(4),frameon=False,fontsize=8)
plt.show()
# +
def get_timing(trials):
times = np.zeros(trials[1].shape[0])
times[np.sum(trials[1][:,131,:],axis=1)>.3] = 130.
times[np.sum(trials[1][:,91,:],axis=1)>.3] = 90.
times[np.sum(trials[1][:,51,:],axis=1)>.3] = 50.
return times
times = get_timing(reg_trials).astype('int')
coh = np.zeros(len(times))
for ii in range(len(times)):
coh[ii] = np.mean(reg_trials[0][ii,times[ii]:times[ii]+40,0])
plt.hist(coh)
plt.show()
# +
M = np.zeros([1000,4])
M[:,0] = rts
M[:,1] = correct.astype('int')
M[:,2] = coh
M[:,3] = 1.
np.save('rnn_rdm_data',M)
# from ddm import Sample
# conditions = ["coh", "monkey", "trgchoice"]
# rnn_sample = Sample.from_numpy_array(M, conditions)
# # correct.astype('int')
# +
# colors = ['b','g','r','c']
# soft_att_0 = softmax(np.mean(att_[:,rule==0,:],axis=1)).T
# soft_att_1 = softmax(np.mean(att_[:,rule==1,:],axis=1)).T
# for ii in range(4):
# plt.plot(soft_att_0[:,ii],colors[ii])
# plt.plot(soft_att_1[:,ii],colors[ii],linestyle='--',label='_nolegend_')
# plt.legend(['rule0','rule1','in0','in1'],frameon=False)
# plt.show()
# +
soft_att = np.zeros(att_.shape)
for ii in range(soft_att.shape[1]):
soft_att[:,ii,:] = softmax(att_[:,ii,:]).T
# plt.plot(soft_att[:,:,0])
# plt.show()
print soft_att[:,times==50,0].shape
plt.subplot(3,1,1)
plt.plot(soft_att[:,times==50,0],'c',alpha=.05)
plt.ylim([-.1,1.1])
plt.axvline(50,c='k',linestyle='--')
plt.axvline(90,c='k',linestyle='--')
plt.axvline(130,c='k',linestyle='--')
plt.subplot(3,1,2)
plt.plot(soft_att[:,times==90,0],'m',alpha=.05)
plt.ylim([-.1,1.1])
plt.axvline(50,c='k',linestyle='--')
plt.axvline(90,c='k',linestyle='--')
plt.axvline(130,c='k',linestyle='--')
plt.subplot(3,1,3)
plt.plot(soft_att[:,times==130,0],'b',alpha=.05)
plt.ylim([-.1,1.1])
plt.axvline(50,c='k',linestyle='--')
plt.axvline(90,c='k',linestyle='--')
plt.axvline(130,c='k',linestyle='--')
plt.tight_layout()
plt.show()
# +
plt.subplot(3,1,1)
plt.plot(np.mean(relu(s_[:,times==50,:]),axis=1),'c',alpha=.5)
# plt.plot(np.mean(np.mean(relu(s_[:,times==90,:]),axis=1),axis=1),'m',alpha=1.)
# plt.plot(np.mean(np.mean(relu(s_[:,times==130,:]),axis=1),axis=1),'b',alpha=1.)
# plt.ylim([-.1,1.1])
plt.axvline(50,c='k',linestyle='--')
plt.axvline(90,c='k',linestyle='--')
plt.axvline(130,c='k',linestyle='--')
plt.subplot(3,1,2)
plt.plot(np.mean(relu(s_[:,times==90,:]),axis=1),'m',alpha=.5)
# plt.ylim([-.1,1.1])
plt.axvline(50,c='k',linestyle='--')
plt.axvline(90,c='k',linestyle='--')
plt.axvline(130,c='k',linestyle='--')
plt.subplot(3,1,3)
plt.plot(np.mean(relu(s_[:,times==130,:]),axis=1),'b',alpha=.5)
# plt.ylim([-.1,1.1])
plt.axvline(50,c='k',linestyle='--')
plt.axvline(90,c='k',linestyle='--')
plt.axvline(130,c='k',linestyle='--')
plt.tight_layout()
plt.show()
# +
time_idx = [50,90,130]
coh_bins = [[0,.01],[.01,.5],[.05,1.]]
plt.figure(figsize=(8,6))
count=1
for ii in range(len(time_idx)):
for jj in range(len(coh_bins)):
plt.subplot(3,3,count)
coh_idx = np.logical_and(np.abs(coh)>coh_bins[jj][0],np.abs(coh)<coh_bins[jj][1])
plt.hist(rts[np.logical_and(times==time_idx[ii],coh_idx)],bins=np.linspace(0,250,30))
plt.xlim([0,250])
plt.axvline(50,c='k',linestyle='--')
plt.axvline(90,c='k',linestyle='--')
plt.axvline(130,c='k',linestyle='--')
count+=1
plt.tight_layout()
plt.show()
# -
for jj in range(len(coh_bins)):
coh_idx = np.logical_and(np.abs(coh)>coh_bins[jj][0],np.abs(coh)<coh_bins[jj][1])
print np.mean(correct[coh_idx])
| notebooks/color_matching_training_attention.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Trigger code in Python from Minecraft
# To find out what is happening in the Minecraft world, Python can ask via the API. The `pollBlockHits()` method will return data about each block hit with a sword since the last time it was called. **Note: You must use a sword, and you must Right-Click. This is the only action detected by this function.**
# Like in other exercises, we need to bring in some libraries of code that enable Python to speak to Minecraft.
import mcpi.minecraft as minecraft
import mcpi.block as block
import time
# Connect to the Minecraft server, store the connection in a variable named `world`:
world = minecraft.Minecraft.create()
# This script works best with blocks that change appearance when their block data changes. One good example is wool, so let's create a small field of wool for you to hit with the sword:
[x,y,z] = world.player.getPos()
world.setBlocks(x,y-1,z,x+10,y-1,z+10, block.WOOL)
# Start looping forever. This is an example of a game loop, because it keeps on processsing until it is interrupted. If you want to stop the loop, press the "stop" button, or use the **Kernel -> Interrupt** option in the notebook menubar.
while True:
hits = world.events.pollBlockHits()
for hit in hits:
print "Have hit {h.x},{h.y},{h.z}".format(h=hit.pos)
block = world.getBlockWithData(hit.pos.x, hit.pos.y, hit.pos.z)
block.data = (block.data + 1) & 0xf ## The & 0xf keeps the block data value below 16 (0xf in hexadecimal)
world.setBlock(hit.pos.x, hit.pos.y, hit.pos.z, block.id, block.data)
world.postToChat("Block data is now " + str(block.data))
time.sleep(1)
# The block of code above is more interesting with blocks that change based on their `data` value, such as wool, flowers, and wood. Wool changes color with the different data values.
# ## Advanced Exercises and Questions
# 1. What other block types change their appearance when you hit them with a sword while this script is running?
# 1. Can you draw a creeper with just this code and a sword?
| classroom-code/exercises/Exercise 5 -- Minecraft changes trigger activity in Python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Early Stopping Example
# In this notebook, we will train an Multi-Layer Perceptron (MLP) to classify images from the [MNIST database](http://yann.lecun.com/exdb/mnist/) hand-written digit database, and use early stopping to stop the training when the model starts to overfit to the training data.
# import libraries
import torch
import numpy as np
import matplotlib.pyplot as plt
# ### Load and Batch the Data
# +
from torchvision import datasets
import torchvision.transforms as transforms
from torch.utils.data.sampler import SubsetRandomSampler
def create_datasets(batch_size):
# percentage of training set to use as validation
valid_size = 0.2
# convert data to torch.FloatTensor
transform = transforms.ToTensor()
# choose the training and test datasets
train_data = datasets.MNIST(root='data',
train=True,
download=True,
transform=transform)
test_data = datasets.MNIST(root='data',
train=False,
download=True,
transform=transform)
# obtain training indices that will be used for validation
num_train = len(train_data)
indices = list(range(num_train))
np.random.shuffle(indices)
split = int(np.floor(valid_size * num_train))
train_idx, valid_idx = indices[split:], indices[:split]
# define samplers for obtaining training and validation batches
train_sampler = SubsetRandomSampler(train_idx)
valid_sampler = SubsetRandomSampler(valid_idx)
# load training data in batches
train_loader = torch.utils.data.DataLoader(train_data,
batch_size=batch_size,
sampler=train_sampler,
num_workers=0)
# load validation data in batches
valid_loader = torch.utils.data.DataLoader(train_data,
batch_size=batch_size,
sampler=valid_sampler,
num_workers=0)
# load test data in batches
test_loader = torch.utils.data.DataLoader(test_data,
batch_size=batch_size,
num_workers=0)
return train_loader, test_loader, valid_loader
# -
# ### Define the Network
# Defining a simple MLP model.
# +
import torch.nn as nn
import torch.nn.functional as F
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
self.fc1 = nn.Linear(28 * 28, 128)
self.fc2 = nn.Linear(128, 128)
self.fc3 = nn.Linear(128, 10)
self.dropout = nn.Dropout(0.5)
def forward(self, x):
# flatten image input
x = x.view(-1, 28 * 28)
# add hidden layer, with relu activation function
x = F.relu(self.fc1(x))
x = self.dropout(x)
# add hidden layer, with relu activation function
x = F.relu(self.fc2(x))
x = self.dropout(x)
# add output layer
x = self.fc3(x)
return x
# initialize the NN
model = Net()
print(model)
# -
# ### Specify Loss Function and Optimizer
# +
# specify loss function
criterion = nn.CrossEntropyLoss()
# specify optimizer
optimizer = torch.optim.Adam(model.parameters())
# -
# ### Import the Early Stopping Class
# import EarlyStopping
from pytorchtools import EarlyStopping
# ### Train the Model using Early Stopping
def train_model(model, batch_size, patience, n_epochs):
# to track the training loss as the model trains
train_losses = []
# to track the validation loss as the model trains
valid_losses = []
# to track the average training loss per epoch as the model trains
avg_train_losses = []
# to track the average validation loss per epoch as the model trains
avg_valid_losses = []
# initialize the early_stopping object
early_stopping = EarlyStopping(patience=patience, verbose=True)
for epoch in range(1, n_epochs + 1):
###################
# train the model #
###################
model.train() # prep model for training
for batch, (data, target) in enumerate(train_loader, 1):
# clear the gradients of all optimized variables
optimizer.zero_grad()
# forward pass: compute predicted outputs by passing inputs to the model
output = model(data)
# calculate the loss
loss = criterion(output, target)
# backward pass: compute gradient of the loss with respect to model parameters
loss.backward()
# perform a single optimization step (parameter update)
optimizer.step()
# record training loss
train_losses.append(loss.item())
######################
# validate the model #
######################
model.eval() # prep model for evaluation
for data, target in valid_loader:
# forward pass: compute predicted outputs by passing inputs to the model
output = model(data)
# calculate the loss
loss = criterion(output, target)
# record validation loss
valid_losses.append(loss.item())
# print training/validation statistics
# calculate average loss over an epoch
train_loss = np.average(train_losses)
valid_loss = np.average(valid_losses)
avg_train_losses.append(train_loss)
avg_valid_losses.append(valid_loss)
epoch_len = len(str(n_epochs))
print_msg = (f'[{epoch:>{epoch_len}}/{n_epochs:>{epoch_len}}] ' +
f'train_loss: {train_loss:.5f} ' +
f'valid_loss: {valid_loss:.5f}')
print(print_msg)
# clear lists to track next epoch
train_losses = []
valid_losses = []
# early_stopping needs the validation loss to check if it has decresed,
# and if it has, it will make a checkpoint of the current model
early_stopping(valid_loss, model)
if early_stopping.early_stop:
print("Early stopping")
break
# load the last checkpoint with the best model
model.load_state_dict(torch.load('checkpoint.pt'))
return model, avg_train_losses, avg_valid_losses
# +
batch_size = 256
n_epochs = 100
train_loader, test_loader, valid_loader = create_datasets(batch_size)
# early stopping patience; how long to wait after last time validation loss improved.
patience = 20
model, train_loss, valid_loss = train_model(model, batch_size, patience, n_epochs)
# -
# ### Visualizing the Loss and the Early Stopping Checkpoint
# From the plot we can see that the last Early Stopping Checkpoint was saved right before the model started to overfit.
# +
# visualize the loss as the network trained
fig = plt.figure(figsize=(10,8))
plt.plot(range(1,len(train_loss)+1),train_loss, label='Training Loss')
plt.plot(range(1,len(valid_loss)+1),valid_loss,label='Validation Loss')
# find position of lowest validation loss
minposs = valid_loss.index(min(valid_loss))+1
plt.axvline(minposs, linestyle='--', color='r',label='Early Stopping Checkpoint')
plt.xlabel('epochs')
plt.ylabel('loss')
plt.ylim(0, 0.5) # consistent scale
plt.xlim(0, len(train_loss)+1) # consistent scale
plt.grid(True)
plt.legend()
plt.tight_layout()
plt.show()
fig.savefig('loss_plot.png', bbox_inches='tight')
# -
# ### Test the Trained Network
# +
# initialize lists to monitor test loss and accuracy
test_loss = 0.0
class_correct = list(0. for i in range(10))
class_total = list(0. for i in range(10))
model.eval() # prep model for evaluation
for data, target in test_loader:
if len(target.data) != batch_size:
break
# forward pass: compute predicted outputs by passing inputs to the model
output = model(data)
# calculate the loss
loss = criterion(output, target)
# update test loss
test_loss += loss.item()*data.size(0)
# convert output probabilities to predicted class
_, pred = torch.max(output, 1)
# compare predictions to true label
correct = np.squeeze(pred.eq(target.data.view_as(pred)))
# calculate test accuracy for each object class
for i in range(batch_size):
label = target.data[i]
class_correct[label] += correct[i].item()
class_total[label] += 1
# calculate and print avg test loss
test_loss = test_loss/len(test_loader.dataset)
print('Test Loss: {:.6f}\n'.format(test_loss))
for i in range(10):
if class_total[i] > 0:
print('Test Accuracy of %5s: %2d%% (%2d/%2d)' % (
str(i), 100 * class_correct[i] / class_total[i],
np.sum(class_correct[i]), np.sum(class_total[i])))
else:
print('Test Accuracy of %5s: N/A (no training examples)' % (classes[i]))
print('\nTest Accuracy (Overall): %2d%% (%2d/%2d)' % (
100. * np.sum(class_correct) / np.sum(class_total),
np.sum(class_correct), np.sum(class_total)))
# -
# ### Visualize Sample Test Results
# +
# obtain one batch of test images
dataiter = iter(test_loader)
images, labels = dataiter.next()
# get sample outputs
output = model(images)
# convert output probabilities to predicted class
_, preds = torch.max(output, 1)
# prep images for display
images = images.numpy()
# plot the images in the batch, along with predicted and true labels
fig = plt.figure(figsize=(25, 4))
for idx in np.arange(20):
ax = fig.add_subplot(2, 20/2, idx+1, xticks=[], yticks=[])
ax.imshow(np.squeeze(images[idx]), cmap='gray')
ax.set_title("{} ({})".format(str(preds[idx].item()), str(labels[idx].item())),
color=("green" if preds[idx]==labels[idx] else "red"))
# -
# ## References
# The MNIST training example code is mainly taken from the [mnist_mlp_solution_with_validation.ipynb](https://github.com/udacity/deep-learning-v2-pytorch/blob/master/convolutional-neural-networks/mnist-mlp/mnist_mlp_solution_with_validation.ipynb) notebook from the [deep-learning-v2-pytorch repository](https://github.com/udacity/deep-learning-v2-pytorch) made by [Udacity](https://www.udacity.com/), and has been fitted with my early stopping code.
| octis/models/early_stopping/MNIST_Early_Stopping_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/tjwei/GAN_Tutorial/blob/master/ACGAN_intro_for_MNIST.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# +
# for tf 2.0
# #!pip install -U tensorflow-gpu
# + colab={} colab_type="code" id="ckOHCuiArv5c"
import tensorflow as tf
import matplotlib.pyplot as plt
import numpy as np
import os
import PIL
import time
from skimage.io import imshow
from IPython.display import display
tf.__version__
# + colab={} colab_type="code" id="1V2PJHrDrxf2"
(train_images, train_labels), (test_images, test_labels) = tf.keras.datasets.mnist.load_data()
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="L0iNo21Arykj" outputId="05458ce2-9340-4569-f2f6-2712f80e1b1c"
train_images.dtype, train_images.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 314} colab_type="code" id="uAcCsSinus1d" outputId="6c5b8499-8d08-415c-c2e3-36520c5c4a30"
imshow(train_images[0])
# + colab={} colab_type="code" id="uSDHJmuwtxT4"
def img_to_float(img):
return (np.float32(img)[..., None]-127.5)/127.5
def img_to_uint8(img):
return np.uint8(img*127.5+128).clip(0, 255)[...,0]
# + colab={"base_uri": "https://localhost:8080/", "height": 314} colab_type="code" id="9yGVvMP9vUn0" outputId="e930d29b-1f49-471f-bce8-353c729eddca"
train_img_f32 = img_to_float(train_images)
imshow(img_to_uint8(train_img_f32[0]))
# + colab={} colab_type="code" id="oSrY5b6ev0kO"
BUFFER_SIZE = train_img_f32.shape[0]
BATCH_SIZE = 32
train_dataset_y = tf.data.Dataset.from_tensor_slices(train_labels).map(lambda y: tf.one_hot(y, 10))
train_dataset_x = tf.data.Dataset.from_tensor_slices(train_img_f32)
train_dataset = tf.data.Dataset.zip((train_dataset_x, train_dataset_y)).shuffle(BUFFER_SIZE).batch(BATCH_SIZE)
# + colab={} colab_type="code" id="ZaJFsMs2v1R1"
from tensorflow.keras.layers import Dense, BatchNormalization, LeakyReLU, Reshape, Conv2DTranspose, Input, concatenate
from tensorflow.keras import Model
latent_dim = 100
_0 = Input((latent_dim,))
_1 = Input((10,))
_ = concatenate([_0, _1])
_ = Dense(7*7*256, use_bias=False, input_shape=(latent_dim,))(_)
_ = BatchNormalization()(_)
_ = LeakyReLU()(_)
_ = Reshape((7, 7, 256))(_)
_ = Conv2DTranspose(128, (5, 5), strides=(1, 1), padding='same', use_bias=False)(_)
_ = BatchNormalization()(_)
_ = LeakyReLU()(_)
_ = Conv2DTranspose(64, (5, 5), strides=(2, 2), padding='same', use_bias=False)(_)
_ = BatchNormalization()(_)
_ = LeakyReLU()(_)
_ = Conv2DTranspose(1, (5, 5), strides=(2, 2), padding='same', use_bias=False, activation='tanh')(_)
generator = Model(inputs=[_0,_1], outputs=_)
# + colab={} colab_type="code" id="kqdl08rxvWiS"
from tensorflow.keras.layers import Conv2D, Dropout, Flatten
_i = Input((28,28, 1))
_ = Conv2D(64, (5, 5), strides=(2, 2), padding='same')(_i)
_ = LeakyReLU()(_)
_ = Dropout(0.3)(_)
_ = Conv2D(128, (5, 5), strides=(2, 2), padding='same')(_)
_ = LeakyReLU()(_)
_ = Dropout(0.3)(_)
_ = Flatten()(_)
_0 = Dense(1)(_)
_1 = Dense(10)(_)
discriminator = Model(inputs=_i, outputs=[_0, _1])
# + colab={} colab_type="code" id="nd8zcnhjzKWq"
BCE = tf.keras.losses.BinaryCrossentropy(from_logits=True)
CCE = tf.keras.losses.CategoricalCrossentropy(from_logits=True)
def generator_loss(generated_output, labels):
out_d, out_c = generated_output
loss_d = BCE(tf.ones_like(out_d), out_d)
loss_c = CCE(labels, out_c)
return loss_d+loss_c
# + colab={} colab_type="code" id="fgWHoC6FzbUA"
def discriminator_loss(real_output, generated_output, labels):
# [1,1,...,1] with real output since it is true and we want our generated examples to look like it
real_out_d, real_out_c = real_output
real_loss_d = BCE(tf.ones_like(real_out_d), real_out_d)
real_loss_c = CCE(labels, real_out_c)
real_loss = real_loss_d + real_loss_c
# [0,0,...,0] with generated images since they are fake
generated_out_d, generated_out_c = generated_output
generated_loss = BCE(tf.zeros_like(generated_out_d), generated_out_d)
total_loss = real_loss + generated_loss
return total_loss
# + colab={} colab_type="code" id="KCYwQqxgz8o_"
generator_optimizer = tf.keras.optimizers.Adam(1e-4)
discriminator_optimizer = tf.keras.optimizers.Adam(1e-4)
# + colab={} colab_type="code" id="Rb2-KU6K1J0I"
EPOCHS = 50
num_examples_to_generate = 20
# We'll re-use this random vector used to seed the generator so
# it will be easier to see the improvement over time.
random_vector_for_generation = tf.random.normal([num_examples_to_generate,
latent_dim])
condition_vector_generation = tf.one_hot(list(range(10))+list(range(10)), 10)
# + colab={} colab_type="code" id="Xnu88uGR1etc"
@tf.function
def train_step(images, labels):
# generating noise from a normal distribution
noise = tf.random.normal([BATCH_SIZE, latent_dim])
with tf.GradientTape() as gen_tape, tf.GradientTape() as disc_tape:
generated_images= generator([noise, labels], training=True)
real_output = discriminator(images, training=True)
generated_output = discriminator(generated_images, training=True)
gen_loss = generator_loss(generated_output, labels)
disc_loss = discriminator_loss(real_output, generated_output, labels)
gradients_of_generator = gen_tape.gradient(gen_loss, generator.trainable_variables)
gradients_of_discriminator = disc_tape.gradient(disc_loss, discriminator.trainable_variables)
generator_optimizer.apply_gradients(zip(gradients_of_generator, generator.trainable_variables))
discriminator_optimizer.apply_gradients(zip(gradients_of_discriminator, discriminator.trainable_variables))
# + colab={"base_uri": "https://localhost:8080/", "height": 874} colab_type="code" id="1DPQkOVx2yon" outputId="1e45a090-fff8-4f8c-dd0d-af96f55f7db3"
for epoch in range(15):
start_time = time.time()
for images, labels in train_dataset:
train_step(images, labels)
fake = generator([random_vector_for_generation, condition_vector_generation], training=False)
fake_concat = np.transpose(img_to_uint8(fake), [1,0,2]).reshape((28,-1))
print(epoch, time.time()-start_time)
display(PIL.Image.fromarray(fake_concat))
| ACGAN_intro_for_MNIST.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
### http://www.gisdeveloper.co.kr/?p=6862
### https://docs.opencv.org/3.4/d2/d55/group__bgsegm.html
# +
import numpy as np
import cv2
cap = cv2.VideoCapture("B3-Parking.avi")
fgbg = cv2.createBackgroundSubtractorKNN(history=100, dist2Threshold = 12.0, detectShadows=False)
while(1):
ret, frame = cap.read()
fgmask = fgbg.apply(frame)
cv2.imshow('frame',fgmask)
k = cv2.waitKey(30) & 0xff
if k == 27:
break
cap.release()
cv2.destroyAllWindows()
| BackgroundSubtractor.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 1. Datatypes
# ---
# **What are comments?**
#
# **Why are comments useful?**
#
# *Comments are ignored by Python Interpreter *
# +
# this is comment
# +
# this is
# multiline comment
# +
# -*- coding: utf-8 -*-
# -
# ## Python Datatypes
# ---
#
# - Integer
# - Float
# - Boolean
# - None ( not exactly a type )
# - Complex
# - String
# - Bytes
# ### Integers
# ---
12
-72
# *Integers are Negative numbers and whole numbers such as -1, -2 ... or 0 or 1, 2 ...*
# ### Floats
# ---
1.5
0.9
-.8
# *Floats are decimal numbers, or in mathematical terms they are Rational/Irrational numbers*
# ### Boolean
# ---
True
False
# *Denotes a expression to be either true or false*
# ### None
# ---
None
# *Defines emptiness or non existence of value*
# ### Complex Number
# ---
#
# see definition of complex number [here](https://en.wikipedia.org/wiki/Complex_number)
1 + 3j
# *In Algebra you would write above as $1 + i3$*
#
# - 1 is real part
# - 3 is imaginary part
# ### String
# ---
'a'
# *__'a'__ is called __char__ in programming term, in Python it is string with length 1.*
'this is string'
"this is also string"
# *Either single quoted '' or doubled quoted "", there is not real difference between them*
"this is
also string"
# *__Note__: single/doubled quoted strings must written in single line, ie newline is not allowed*
# #### Tripple quoted string
""" This is new string
with new line that
preserves spaces
"""
''' This is same as above,
only use of type of quote
differs'''
# *While triple quoted __'''__ or __"""__, preserves newlines and spaces.*
# ## Mathematical Operations
# ---
# ### Addition ( + operator )
# ---
# $int + int = int$
1 + 2
# *Python automatically converts similar type, also known as implicit conversion.*
# $float + int = float + float(int) = float$
1.5 + 2
# $int + bool = int + int(bool) = int$
1 + True
# $float + bool = float + float(bool) = float$
1.5 + True
# $int + string \neq int + int(string)$
1 + '2'
# *You have to do it yourself. This is called explicit conversion.*
1 + int('2')
# *We need to explicity change a string to integer ourselves, in this case.*
# $Concatenation$
'a' + 'bc'
# *This is concatenation ie joining of two strings together.*
1.5 + (1 + 4j)
(1 + 2j) + (3 + 5j)
b'a' + b'bc'
# ### Substraction ( - ), Multiplication ( * ), Division ( / or // )
# ---
# **H/W:** *Try all above operations with substraction (__-__)*
# #### Real Division
3 / 2
4.8 / 4
2 / 5.9
# #### Integer Division
3 // 2
4.8 // 4
2 // 5.9
# *Note: Here some of the above answers are return as __float__ this is beacuse one of the operand is __float__, in that case it will try to cast result into __float__.*
#
# *Notice, that result return by __real__ division and __integer__ division are not same, for same operands.*
# #### Multiplication
3 * 2
99.1 * 88
# #### Repitition
'a' * 3
'hello ' * 5
# *Note, that when __$*$__ is used with string and integer it repeats the string with given times.*
# **H/W try all the combinations**
# #### Power ( ** )
# $2**3 \rightarrow 2^3$
2 ** 3
# ## Names/Variables
# ---
#
# We give _name_ to data, so that they can be identified later on. Such names are also known as _variables_.
pi = 3.1415
# *Much like above names must be meaningful*
#
# *for example, if we need to use $\pi$ later on we can just use __pi__ instead of using __3.1415__ each time.*
# There is a rule for defining a _variable_ name or just _name_ ( which we will call from now on. )
#
# - It must start with either a _character_ or a _underscore_
# - while it can have a _character_, a _underscore_ or a _number_ after that
a = 1
_ = 5
a1 = 6
# *valid names*
2bc = 77
# *Invalid name*
# ## Quick Task
# ---
#
# ### Sum of two numbers
# lets add two numbers 72 and 67
a = 72
b = 67
sum_of_addition = a + b
print(sum_of_addition)
# *Review ( line by line )*
#
# ```python
# 1. # lets add two numbers 72 and 67
# 2. a = 72
# 3. b = 67
# 4. sum_of_addition = a + b
# 5. print(sum_of_addition)
# ```
#
# 1. __comment__ starts with __#__ symbol,
# - comments are part of code which are ignored by python, but is there for us to put notes/explanation of code.
# 2. we assign __72__ to __a__.
# - __a__ is _name_ we assign __72__ to, which is generally called as _variable_
# 3. similarly we assign __67__ to __b__
# 4. we add __a__ and __b__, and assign result to __sum\_of\_addition__
# 5. finally we are displaying result using __print__ function
# - [reference to print function](http://nbviewer.jupyter.org/github/lfapython/pybasics/blob/master/Python/Beginner/unit-01-datatypes-reference.ipynb#print)
#
# Finally we get result as __139__, But what happens when we change __72__ to __72.8__ and __67__ to __67.2__?
72.8 + 67.2
# *There are two types of __numbers__ in python, __integer__ and __float__.*
#
# - __integers__ are Negative numbers, 0 and Positive numbers.
# - __floats__ are decimal numbers, the ones with decimal point in it.
1.8 + 6
# *While using either integer or float, we get following result*
#
# - Adding __int__ ( short for integer ) and __int__, we get result as __int__
# - Adding __float__ and __float__, we get result as __float__
# - Adding __int__ and __float__, we get result as __float__
# - This is because __float__ are stored in larger container than __int__.
# - And it is easier to put smaller values into larger container than to put larger values in smaller container without chopping data.
# ### Difference of two numbers
#
# Try using __-__ operator instead of __+__ with different numbers.
89.4 - 12
# ## Other Strings
# ---
# ### Unicode string
'नेपालि'
# *Unicode is builtin to Python 3 Strings.*
#
# *Strings are sequence of unicodes.*
# ### Bytes
b'This is byte string'
# *Bytes are sequence of bytes ie number from 0 to 255*
# ## Builtin Helpful Functions
# ---
#
# Builtin functions are those given by python, which gives us certain information or does some tasks.
#
# functions are generally **names** with opening and closing parenthesis **()**.
#
# between parenthesis we pass in some value,
# ### print
print("Hello world")
print(2)
print(5 ** 4)
# *__print()__ function prints/displays anything into screen.*
#
# *Note: while anything that is last on notebook cell (like this) are displayed directly, which is not the case in application run in command prompt or terminal.*
# ### input
input("Enter a number:")
inp = input("Enter your name: ")
inp
# *__input()__ is exact opposite of __print__, in that it takes data from user*
#
# *Note: notice we used __inp__ name to store data obtain from input.*
#
# *Note: the data obtain from user is always string type.*
# ### type
type(1)
type(2.5)
type('Nepal')
type(2 + 3j)
type(b'This is what?')
# *__type()__ functions returns/gives, what is datatype of given data/object.*
# ### len
len('This is some string')
len(b'This')
len('नेपाली')
# *__len()__ functions returns/gives the length of given string or bytes*
#
# *It doesn't work on other datatypes above*
# ### help
help(dir)
# *__help()__ shows help text for given data/functions*
# ## Changing from one type to another (Type Casting)
# ---
#
# Changing from one type to another is called type casting
# ### int
int(1.2)
int('4')
int(True)
# *We can change, a float, a boolean or a string into integer using __int()__ function*
int(2 + 3j)
# *We cannot change a complex number to a integer*
int('a')
# *While we must remember only numbers represented as string (or numbers in quotes) can be converted to integer*
int('1')
# *but not*
int('1a')
# ### float
float(1)
float('5.5')
float(False)
float('p')
float(3 + 7j)
# *Similar to __int()__, __float()__ functions changes a integer, string or boolean to a float*
# ### str
str(1)
str(4.5)
str(True)
str(5 + 9j)
# *__str()__ function converts all other types to string.*
# ### complex
complex(1)
complex(4.5)
complex('4')
complex(True)
complex('p')
# *__complex()__ function converts other datatypes to complex number.*
#
# *__Note:__ Remember the difference between string number and string character.*
#
#
height = 8848
# *__height__ is given name for __8848__*
height.real
height.imag
# *Above are attributes for integer i.e __height__*
cmplx = 7 + 5j
cmplx.real
cmplx.imag
cmplx.conjugate()
# *conjugate is __method__ associated with a complex number*
#
# *__Note:__ Remember the __dot__ it follows.*
# Resources
#
# - http://learnpython.org
#
# - http://www.diveintopython3.net/
#
# - http://www.pythontutor.com
| References/1.Datatypes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# importing necessary libraries
import pandas as pd
import numpy as np
from datetime import datetime
# To plot pretty figures
# %matplotlib inline
import matplotlib
import matplotlib.pyplot as plt
import seaborn as sns
# +
# import training data
train_df = pd.read_csv('sales_train_v2.csv')
# import information on shops
shops_df = pd.read_csv('shops.csv')
# import information on items
items_df = pd.read_csv('items.csv')
# import information on categories
categories_df = pd.read_csv('item_categories.csv')
# -
# looking at description of training data
train_df.describe()
# checking missing values in training data
train_df.isnull().sum()
train_df.head()
# count number of unique items
train_df['item_id'].nunique()
# count the number of times each item was sold OR returned
number_of_items = train_df.groupby('item_id').size()#.sort_values(ascending=False)
number_of_items_df = pd.DataFrame([])
number_of_items_df['item_num'] = list(number_of_items)
number_of_items_df.head()
fig = plt.hist(number_of_items_df['item_num'], bins=101)
# items that have been sold at least 50 times give 94% of all sales
number_of_items_df[number_of_items_df['item_num'] > 50].sum()/2.935849e+06
# items sorted by their price
fig = plt.hist(train_df['item_price'], bins=101)
# items under 300 Rubles give 94% of all sales
train_df[train_df['item_price'] < 3000].shape[0]/2.935849e+06
train_df.head()
items_df.head()
shops_df.head()
categories_df.head()
train_df = train_df.merge(items_df, on = 'item_id',how='left')
train_df = train_df.merge(shops_df, on = 'shop_id',how='left')
train_df = train_df.merge(categories_df, on = 'item_category_id',how='left')
train_df.head()
# check if the item prices are always the same
gb = train_df[['item_id','item_price']].groupby(['item_id']).agg(['min', 'max'], axis="columns")
gb['item_price']['min'] - gb['item_price']['max']
train_df.head()
gr = train_df[['date_block_num','shop_id','item_id','item_price','item_cnt_day']].groupby(['date_block_num','shop_id','item_id'], as_index = False)
gr.agg({"item_price": "mean","item_cnt_day": "sum"}).head()
train_df[(train_df['date_block_num'] == 0) & (train_df['shop_id'] == 0) & (train_df['item_id'] == 32)]\
[['date_block_num','shop_id','item_id','item_price','item_cnt_day']]
| Predict_Sales_v1.0.ipynb |