code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exercises 7
# In exercises 5 and 6 we tried to segment the lights in the rocket image using thresholds and binary operations. We are now going to try to find them by template matching.
#
# - import the rocket image
# - convert it to gray scale
# - try to manually crop a small region around one of the lights (choose limits, plot the region and adjust)
# - use that region as template to match over the full image
# - find local maxima in the matched image
# - plot them on top of the image
# - plot the pixel values at the positions of the maxima
# - can you subselect only those corresponding to ligths
# - are some of the lights missing ? Why ? Can we fix it ?
#
# # Solutions 7
import numpy as np
import matplotlib.pyplot as plt
import skimage.data
#load moon image
image = skimage.data.rocket()
#convert to gray scale
image_gray = skimage.color.rgb2gray(image)
plt.gray()
plt.imshow(image_gray)
plt.show()
#crop around a light
image_crop = image_gray[368:387,141:160]
plt.imshow(image_crop)
plt.show()
from skimage.feature import match_template, peak_local_max
#do the template matching
matched = match_template(template=image_crop, image=image_gray, pad_input=True, mode='mean')
plt.imshow(matched)
plt.show()
#find local maxima
local_max = peak_local_max(matched,threshold_abs=0.5, min_distance=10, indices=False)
local_max_ind = peak_local_max(matched,threshold_abs=0.5, min_distance=10, indices=True)
#plot to maxima on the image
plt.imshow(image_gray)
plt.plot(local_max_ind[:,1],local_max_ind[:,0],'ro',alpha = 0.3)
plt.show()
plt.plot(image_gray[local_max_ind[:,0],local_max_ind[:,1]],'o')
plt.show()
# +
#make subselection based on the plot above
intensities = image_gray[local_max_ind[:,0],local_max_ind[:,1]]
local_max_ind = local_max_ind[intensities>0.8,:]
plt.imshow(image_gray)
plt.plot(local_max_ind[:,1],local_max_ind[:,0],'ro',alpha = 0.3)
plt.show()
# -
# We are missing the two lights at the bottom of the image. This is because they are very close to the image adge and the matching is not complete. We can try to extend the image by padding it with a few rows:
image_gray = np.pad(image_gray,((0,10),(0,0)),mode='mean')
# +
matched = match_template(template=image_crop, image=image_gray, pad_input=True, mode='mean')
local_max_ind = peak_local_max(matched,threshold_abs=0.5, min_distance=10, indices=True)
intensities = image_gray[local_max_ind[:,0],local_max_ind[:,1]]
local_max_ind = local_max_ind[intensities>0.8,:]
plt.imshow(image_gray)
plt.plot(local_max_ind[:,1],local_max_ind[:,0],'ro',alpha = 0.3)
plt.show()
# -
| Exercises/Exercise7.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_pytorch_p36
# language: python
# name: conda_pytorch_p36
# ---
# # 1. Build docker image
# !aws ecr get-login-password --region us-east-1 | docker login --username AWS --password-stdin 7<PASSWORD>4.dkr.ecr.us-east-1.amazonaws.com
# !docker build -t grammer-classification-bert-base-uncased -f Dockerfile .
# !docker run --name ivy-grammer-classification grammer-classification-bert-base-uncased:latest
# # 2. Push to ECR
# +
import boto3
client = boto3.client("sts")
account = client.get_caller_identity()["Account"]
my_session = boto3.session.Session()
region = my_session.region_name
algorithm_name = "grammer-classification-bert-base-uncased"
ecr_image = "{}.dkr.ecr.{}.amazonaws.com/{}:latest".format(account, region, algorithm_name)
print(ecr_image)
# +
# # !aws ecr get-login-password | docker login xxxx -U AWS --password-stdin
# # !docker tag pytorch-bert-base-uncased:latest ECR_IMAGE
# # !docker push ECR_IMAGE
# + language="sh"
#
# # The name of our algorithm
# algorithm_name=grammer-classification-bert-base-uncased
#
# account=$(aws sts get-caller-identity --query Account --output text)
#
# # Get the region defined in the current configuration (default to us-west-2 if none defined)
# region=$(aws configure get region)
# region=${region:-us-west-2}
#
# fullname="${account}.dkr.ecr.${region}.amazonaws.com/${algorithm_name}:latest"
#
# # If the repository doesn't exist in ECR, create it.
#
# aws ecr describe-repositories --repository-names "${algorithm_name}" > /dev/null 2>&1
#
# if [ $? -ne 0 ]
# then
# aws ecr create-repository --repository-name "${algorithm_name}" > /dev/null
# fi
#
# # Get the login command from ECR and execute it directly
# $(aws ecr get-login --region ${region} --no-include-email)
#
# # # Get the login command from ECR in order to pull down the SageMaker PyTorch image
# # $(aws ecr get-login --registry-ids 520713654638 --region ${region} --no-include-email)
#
# # # Build the docker image locally with the image name and then push it to ECR
# # # with the full name.
#
# aws ecr get-login-password --region us-east-1 | docker login --username AWS --password-stdin ${account}.dkr.ecr.${region}.amazonaws.com
#
# # docker build -t ${algorithm_name} . --build-arg REGION=${region}
# docker tag ${algorithm_name} ${fullname}
#
# docker push ${fullname}
# -
# # 3. Deploy to Sagemaker Endpoint
# +
import os
import numpy as np
import pandas as pd
import sagemaker
sagemaker_session = sagemaker.Session()
bucket = sagemaker_session.default_bucket()
prefix = "sagemaker/ivy-demo-pytorch-bert"
role = sagemaker.get_execution_role()
model_data="s3://{}/{}/pytorch-training-2021-11-29-14-48-31-535/output/model.tar.gz".format(bucket, prefix)
print(bucket)
print(role)
print(model_data)
# +
# help(sagemaker_session.upload_data)
# -
# !aws s3 ls s3://sagemaker-us-east-1-420737321821/sagemaker/ivy-demo-pytorch-bert/pytorch-training-2021-11-29-14-48-31-535/output/model.tar.gz
# ## 3.1 Creating a model from training output (model artifacts)
model_name_prefix = "{}-model".format(algorithm_name)
timestamp = time.strftime('-%Y%m%d%H%M%S', time.gmtime())
model_name = model_name_prefix + timestamp
print(model_name)
# +
sagemaker_client = boto3.client(service_name='sagemaker')
model_artifacts_s3_path = model_data
docker_image = algorithm_name
create_model_response = sagemaker_client.create_model(
ModelName = model_name,
ExecutionRoleArn = role,
PrimaryContainer = {
'Image': docker_image,
'ModelDataUrl': model_artifacts_s3_path,
}
)
print(create_model_response['ModelArn'])
# -
len("grammer-classification-bert-base-uncased-model-2021-12-02-03-09-55")
# ## 3.1. Create Pytorch endpoint with predefined the pytorch framework
# - it will create the inference model and endpoint configure automatically
# - it will use the awe prebuilt pytorch framework, we don't need to build our own docker imag, but need to prepare the entrypoint script
# +
# from sagemaker.pytorch import PyTorchModel
# help(PyTorchModel)
import sagemaker
# help(sagemaker.model.FrameworkModel)
# help(sagemaker.model.Model)
# from sagemaker.estimator import Estimator
# help(Estimator)
# help(estimator.fit)
# -
WORK_DIRECTORY = "/home/ec2-user/SageMaker/amazon-sagemaker-bert-pytorch/cola_public"
my_prefix="sagemaker/ivy-demo-pytorch-bert/data/input"
data_location = sagemaker_session.upload_data(WORK_DIRECTORY, bucket=bucket, key_prefix=my_prefix)
print(data_location)
# !aws s3 mv s3://sagemaker-us-east-1-420737321821/sagemaker/ivy-demo-pytorch-bert/test.csv s3://sagemaker-us-east-1-420737321821/sagemaker/ivy-demo-pytorch-bert/data/input/test.csv
# !aws s3 mv s3://sagemaker-us-east-1-420737321821/sagemaker/ivy-demo-pytorch-bert/train.csv s3://sagemaker-us-east-1-420737321821/sagemaker/ivy-demo-pytorch-bert/data/input/train.csv
# !aws s3 ls s3://sagemaker-us-east-1-420737321821/sagemaker/ivy-demo-pytorch-bert/
# +
from sagemaker.estimator import Estimator
import time
instance_type = 'ml.m5.large'
# accelerator_type = 'ml.eia2.xlarge'
# instance_type = 'ml.t2.medium'
hyperparameters = {"epochs": 1}
estimator = Estimator(
role=role,
train_instance_count=1,
train_instance_type=instance_type,
image_uri="grammer-classification-bert-base-uncased:latest",
model_uri=model_data,
hyperparameters=hyperparameters,
)
estimator.fit()
# estimator.fit("s3://sagemaker-us-east-1-420737321821/sagemaker/ivy-demo-pytorch-bert/data/input/")
# estimator.fit("file:///tmp/pytorch-example/cifar-10-data")
endpoint_name = 'grammar-classification-{}-ep'.format(time.time()).replace('.', '').replace('_', '')
print(endpoint_name)
predictor = estimator.deploy(
initial_instance_count=1,
instance_type="local", #instance_type,
# accelerator_type=accelerator_type,
endpoint_name=endpoint_name,
wait=True,
)
# +
from sagemaker.pytorch import PyTorchModel
import time
# instance_type = 'ml.m5.large'
# accelerator_type = 'ml.eia2.xlarge'
instance_type = 'ml.t2.medium'
endpoint_name = 'grammar-classification-{}-ep'.format(time.time()).replace('.', '').replace('_', '')
print(endpoint_name)
pytorch = PyTorchModel(
model_data=model_data,
role=role,
entry_point='train_deploy.py',
source_dir='code',
# framework_version='1.3.1',
# py_version='py3',
image_uri="grammer-classification-bert-base-uncased:latest",
sagemaker_session=sagemaker_session
)
# Function will exit before endpoint is finished creating
predictor = pytorch.deploy(
initial_instance_count=1,
instance_type="local", #instance_type,
# accelerator_type=accelerator_type,
endpoint_name=endpoint_name,
wait=True,
)
# -
# +
# from sagemaker.estimator import Estimator
# help(Estimator.fit)
# +
from sagemaker.estimator import Estimator
# hyperparameters = {"epochs": 1}
hyperparameters={
"epochs": 1,
"num_labels": 2,
"backend": "gloo",
"lr":0.1
}
instance_type = "ml.m4.xlarge"
model_uri="s3://sagemaker-us-east-1-420737321821/sagemaker/ivy-demo-pytorch-bert/pytorch-training-2021-11-29-14-48-31-535/output/model.tar.gz"
estimator = Estimator(
role=role,
train_instance_count=1,
train_instance_type=instance_type,
image_name=ecr_image,
model_uri=model_uri,
hyperparameters=hyperparameters,
)
estimator.fit()
predictor = estimator.deploy(1, instance_type)
# -
import boto3
from sagemaker.amazon.amazon_estimator import get_image_uri
training_image = get_image_uri(boto3.Session().region_name, 'image-classification')
print(training_image)
# # 4. Test the endpoint
| grammar-classification-deploy-custom-dockerimg.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # AlphaQUBO Tutoral
# This tutorial shows how to read an AlphaQUBO formatted file, process it as a D-Wave BinaryQuadraticModel, and submit to an AlphaQUBO solver.
#
# The AlphaQUBO file format is a text file with information on the first line that describes the size of the q(i,j) matrix.
#
# 1. Program line is marked by a “**p**” in the first column. A single program line must be the first line in the file. The program line has two arguments: **variable_count** and **non_zero_element_count**
#
# ```
# p 50 225
# ```
#
# 2. The remaining lines are made up of three numbers, separated by one or more blanks. The first two numbers, ($i$ and $j$), are the indices for this Q matrix entry, where ($i <= j$). Each index must be in the range {1, **variable_count**}. The third number is the value of the Q matrix at $Q(i,j)$, specified as an integer or floating point value
#
# #### Example
# ```
# p 100 475
# 1 35 -19
# 1 44 -22
# 1 47 27
# 1 49 -66
# 1 58 -69
# 1 64 63
# 1 72 -89
# 1 73 -19
# 1 74 -69
# 1 76 -12
# 1 84 40
# 1 98 33
# 2 2 52
# ```
#
import sys
import alphaqubo_client as aq
from alphaqubo_client.rest import ApiException
import dimod
# ## Read AlphaQUBO formatted file as BinaryQuadraticModel
def read_alphaqubo_bqm(filename:str):
lines = []
with open(filename, "rt") as myfile:
for line in myfile:
lines.append(line.rstrip('\n'))
Q = {}
for line in lines:
p = line.split()
if len(p) > 0:
if p[0] == 'c':
continue
elif p[0] == 'p':
nVars = int(p[1])
else:
if len(p) == 3:
i = int(p[0]) - 1
j = int(p[1]) - 1
w = float(p[2])
if i != j:
w *= 2
Q[(i, j)] = w
bqm = dimod.BinaryQuadraticModel.from_qubo(Q)
return bqm
# ## Configure AlphaQUBO API
# Configure the connection details to connect to AlphaQUBO SolverAPI
# +
configuration = aq.Configuration()
configuration.debug = False
configuration.host = "http://localhost:5000"
api_instance = aq.QuboApi(aq.ApiClient(configuration))
# -
# ### Read AlphaQUBO Formatted File
#
# + tags=[]
bqm = read_alphaqubo_bqm('../data/bqp2500_1.txt')
print("Size of BQM: ", len(bqm))
# -
# ### AlphaQUBO dimod Sampler
# +
from dimod.core.sampler import Sampler
from dimod.sampleset import SampleSet
from six import iteritems
class AlphaQuboSampler(Sampler):
properties = None
parameters = None
def __init__(self):
self.properties = {}
self.parameters = {'time_limit': [],
'accuracy_min': [],
'greediness': [],
'maximize': [] }
def sample(self, bqm, api_instance, time_limit=60, accuracy_min=5, greediness=0.0, maximize=False):
n = len(bqm.variables)
if n == 0:
return SampleSet.from_samples([], bqm.vartype, energy=[])
linear = bqm.linear
inverse_mapping = dict(enumerate(linear))
mapping = {v: i for i, v in iteritems(inverse_mapping)}
bqm2 = bqm.relabel_variables(mapping, inplace=False)
body = aq.SolverAPI()
body.num_vars = n
if maximize:
body.min_max = 1
else:
body.min_max = 0
body.timeout = time_limit
body.inputs = []
variables = sorted(bqm2.iter_variables())
for idx, u in enumerate(variables):
for v in variables[idx:]:
if u == v and bqm2.linear[u]:
body.inputs.append(aq.Points(x=u + 1, y=u + 1, z=bqm2.linear[u]))
for idx, u in enumerate(variables):
for v in variables[idx:]:
if u in bqm2.adj[v]:
body.inputs.append(aq.Points(x=u + 1, y=v + 1, z=bqm2.adj[u][v] / 2))
body.non_zero = len(body.inputs)
body.parameters = "-am " + str(accuracy_min)
try:
# Use the inputs to define a QUBO and solve it synchronously.
api_response = api_instance.api_qubo_solve_qubo_post(body=body)
except ApiException as e:
print("Exception when calling QuboApi->api_qubo_solve_qubo_post: %s\n" % e)
samples = []
energies = []
samples.append( api_response.solved_result )
energies.append( api_response.solved_value )
response = SampleSet.from_samples_bqm((samples, list(bqm.variables)), bqm)
return response
# -
# ### Solve using AlphaQUBO
# + tags=[]
# %time response = AlphaQuboSampler().sample(bqm, api_instance, maximize=True, time_limit=15)
print(response)
# -
| notebooks_under_development/AlphaQUBO-Tutorial-1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
# #%matplotlib inline
# %load_ext autoreload
# %autoreload 2
import torch
import torch.nn as nn
import torch.nn.functional as F
import numpy as np
import time
import random
import copy
from copy import deepcopy
import threading
from train_utils import *
import matplotlib.pyplot as plt
import cv2
# +
from torchvision import transforms
from IPython.core.display import Image as JupyterImage
from torchvision.utils import save_image
deg_to_rad = lambda x: x*0.0174533
crop = transforms.CenterCrop(48)
resize = transforms.Resize(64)
color_jitter = transforms.ColorJitter(brightness=.5, contrast=.5, saturation=.5, hue=.5)
def get_rotated_view(front, aux, rotation):
front = front.clone()
SEQ_LEN, BS, C, H, W = front.shape
ff = front.reshape(SEQ_LEN*BS, C, H, W)
ff = transforms.functional.rotate(ff, rotation)
ff = crop(ff)
ff = resize(ff)
##ff = color_jitter(ff)
ff = ff.reshape(SEQ_LEN, BS, C, H, W)
_aux = aux.clone()
_aux[:,:,0] -= deg_to_rad(rotation)
return ff, _aux
# -
torch.__version__
import gym3
from procgen import ProcgenGym3Env
train_num_levels = 100_000 #500 #1500
train_start_level = 0
color_themes_indist = [1,2,3,4,5]
indist_backnoise = 0
color_themes_road_indist = [1,2,3,4,5] #2
# +
bs = 1
# num_levels=1, start_level=6
env = ProcgenGym3Env(num=bs, env_name="testgame", render_mode='rgb_array')
bs = 64
env_indist = ProcgenGym3Env(num=bs, env_name="testgame", num_levels=train_num_levels, start_level=train_start_level,
color_theme=color_themes_indist, color_theme_road=color_themes_road_indist,
background_noise_level=indist_backnoise, render_mode='rgb_array')
# -
env = gym3.ViewerWrapper(env_indist, info_key="rgb")
device = 'cuda'
m = VizCNN(use_rnn=False).to(device);
m.load_state_dict(torch.load("m.torch"))
m.eval()
hidden = get_hidden(1)
# +
# %%time
s = np.array([[.1, .2] for _ in range(bs)], dtype=np.float32)
daggerized_controls = s
seq_len = 60
TRAINING_WHEELS_WINDOW = 50 # NOTE WE"RE STILL JUST LETTING AP DRIVE HERE
use_training_wheels = True
act_grads = []
imgs = []
salmaps = []
for i in range(seq_len):
env.act(s)
rew, obs, first = env.observe()
img = obs['rgb']
info = env.get_info()
aux = np.array([[e[a] for a in aux_properties] for e in info])
autopilot_controls = np.array([[e['autopilot_steer'], e['autopilot_throttle']] for e in info])
front = torch.from_numpy(img.astype(np.float32)/255.).unsqueeze(0).permute(0,1,4,2,3)
aux = torch.from_numpy(aux.astype(np.float32)).unsqueeze(0)
front, aux = get_rotated_view(front, aux, 0)
front = front.to(device)
aux = aux.to(device)
out,hidden, salmap = m(front, aux, hidden, return_salmap=True, register_activations=True)
out[:,:,1]=.7
s = out.squeeze(0).cpu().detach().numpy()
################################
baseline = torch.zeros_like(front)
steps = 20
grads = []
for i in range(0,steps+1):
mixed_img = baseline + (float(i)/steps)*(front-baseline)
m.zero_grad()
out, hidden, salmap = m(mixed_img, aux, hidden, return_salmap=True, register_activations=True)
out[0][0][0].backward(retain_graph=m.use_rnn) # Steer
g = m.get_activations_gradient()
g = g[0].mean(0).cpu().numpy()
grads.append(g)
avg_grads = np.stack(grads).mean(0)
act_grad = avg_grads
salmap = salmap[0].mean(0)
act_grads.append(act_grad * salmap)
#act_grads.append(salmap) # This by itself is pretty nice
##################################
imgs.append(img[0])
if use_training_wheels and i < TRAINING_WHEELS_WINDOW:
s = autopilot_controls
# -
act_grad.shape, act_grad.max()
plt.imshow(act_grad)
plt.imshow(salmap)
plt.imshow(salmap*act_grad)
# +
def img_cam(act_grad, img, std):
# Gradients zero to one
act_grad = cv2.resize(act_grad, (img.shape[0],img.shape[1]))
mask = np.where(act_grad, (abs(act_grad)>std*2), 0)
mask = np.expand_dims(mask, -1)
act_grad -= act_grad.min()
act_grad = act_grad / act_grad.max()
# Make a three-channel heatmap out of the one channel gradients
heatmap = cv2.applyColorMap(np.uint8(255 * act_grad), cv2.COLORMAP_JET)
heatmap = np.float32(heatmap) / 255
heatmap = heatmap * mask
heatmap = heatmap * 255
#img = img / 255
cam = heatmap*.5 + img
#cam = cam / cam.max()
#cam = cam * 255
cam = np.clip(cam, 0, 255)
cam = cam.astype(np.uint8)
return cam
# -
std = np.array(act_grads[20:]).std()
img_cams = []
for img, act_grad in zip(imgs,act_grads):
img_cams.append(img_cam(act_grad,img,std))
# +
# IG vid
img1 = img_cams[0]
height , width , layers = img1.shape
fps = 20
video = cv2.VideoWriter('cams.avi', cv2.VideoWriter_fourcc(*"MJPG"), fps, (width,height))
for i in range(len(imgs)-1):
img = img_cams[i] # cv2 expects out of 255, integers
img = np.flip(img, -1)
video.write(img)
cv2.destroyAllWindows()
video.release()
# +
# Just the vid
#img1 = img_cams[0]
img1 = imgs[0]
height , width , layers = img1.shape
fps = 20
video = cv2.VideoWriter('cams.avi', cv2.VideoWriter_fourcc(*"MJPG"), fps, (width,height))
for i in range(len(imgs)-1):
img = imgs[i]
img = np.flip(img, -1)
#img = img_cams[i] # cv2 expects out of 255, integers
video.write(img)
cv2.destroyAllWindows()
video.release()
# -
salmap.shape
| viz.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# Remember to execute this cell with Control+Enter
import sys;
sys.path.append('../');
import jupman;
# # Functions 5 - exercises with tuples
#
# ## [Download exercises zip](../_static/generated/functions.zip)
#
# [Browse files online](https://github.com/DavidLeoni/softpython-en/tree/master/functions)
#
# ### Exercise - joined
#
# ✪✪ Write a function which given two tuples of characters `ta` and `tb` having each different characters (may also be empty), return a tuple made like this:
#
# * if the tuple `ta` terminates with the same character `tb` begins with, RETURN the concatenation of `ta` and `tb` WITHOUT duplicated characters
# * otherwise RETURN an empty tuple
#
# Example:
#
# ```python
# >>> joined(('a','b','c'), ('c','d','e'))
# ('a', 'b', 'c', 'd', 'e')
# >>> joined(('a','b'), ('b','c','d'))
# ('a', 'b', 'c', 'd')
# ```
#
# +
def joined(ta,tb):
#jupman-raise
if len(ta) > 0 and len(tb) > 0:
if ta[-1] == tb[0]:
return ta[:-1] + tb
return ()
#/jupman-raise
assert joined(('a','b','c'), ('c','d','e')) == ('a', 'b', 'c', 'd', 'e')
assert joined(('a','b'), ('b','c','d')) == ('a', 'b', 'c', 'd')
assert joined((),('e','f','g')) == ()
assert joined(('a',),('e','f','g')) == ()
assert joined(('a','b','c'),()) == ()
assert joined(('a','b','c'),('d','e')) == ()
# -
# ### nasty
#
# ✪✪✪ Given two tuples `ta` and `b`, `ta` made of characters and `tb` of positive integer numbers , write a function `nasty` which RETURNS a tuple having two character strings: the first character is taken from `ta`, the second is a number taken from the corresponding position in `tb`. The strings are repeated for a number of times equal to that number.
#
# ```python
# >>> nasty(('u','r','g'), (4,2,3))
# ('u4', 'u4', 'u4', 'u4', 'r2', 'r2', 'g3', 'g3', 'g3')
#
# >>> nasty(('g','a','s','p'), (2,4,1,3))
# ('g2', 'g2', 'a4', 'a4', 'a4', 'a4', 's1', 'p3', 'p3', 'p3')
# ```
# +
# write here
def nasty(ta, tb):
#jupman-raise
i = 0
ret = []
while i < len(tb):
s = ta[i]+str(tb[i])
ret.extend( (s,) * tb[i] )
i += 1
return tuple(ret)
#/jupman-raise
# TEST START - DO NOT TOUCH !
assert nasty(('a',), (3,)) == ('a3','a3','a3')
assert nasty(('a','b'), (3,1)) == ('a3','a3','a3','b1')
assert nasty(('u','r','g'), (4,2,3)) == ('u4', 'u4', 'u4', 'u4', 'r2', 'r2', 'g3', 'g3', 'g3')
assert nasty(('g','a','s','p'), (2,4,1,3)) == ('g2', 'g2', 'a4', 'a4', 'a4', 'a4', 's1', 'p3', 'p3', 'p3')
# TEST END
# -
# ## Continue
#
# Go on with exercises about [functions and sets](https://en.softpython.org/functions/fun6-sets-sol.html)
| functions/fun5-tuples-sol.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6.9 64-bit
# name: python36964bit29e3c49a4a324ceaaa2a919cf9d3ccc4
# ---
import json
from tqdm import tqdm
import os
import googlemaps
with open("data.json") as f:
data = json.load(f)
print(len(data))
data[2]
paper_lookup = {}
for node in data:
paper_lookup[node["id"]] = node
# +
filename = "institutions.json"
if os.path.isfile(filename):
with open("institutions.json", "r") as f:
institutions = json.load(f)
else:
institutions = {}
for node in data:
seen_inst_this_node = []
for a in node.get("authors"):
id = a.get("AfId")
if id and id not in seen_inst_this_node:
seen_inst_this_node.append(id)
if id not in institutions:
institutions[id] = {
"id": id,
"name": a.get("DAfN"),
"n_papers": 0,
"n_citations": 0,
"links": {}
}
institutions[id]["n_papers"] += 1
institutions[id]["n_citations"] += node["citation_count"]
for ref in node["references"]:
other = paper_lookup[ref]
for oa in other.get("authors"):
oid = oa.get("AfId")
if oid:
if oid not in institutions[id]["links"]:
institutions[id]["links"][oid] = 0
institutions[id]["links"][oid] += 1
institutions = list(institutions.values())
print(len(institutions))
gmaps = googlemaps.Client(key='INSERT_KEY_HERE')
for inst in tqdm(institutions):
if "geo" not in inst:
inst["geo"] = gmaps.geocode(inst["name"])
with open("institutions.json", "w") as f:
json.dump(institutions, f)
print(len(institutions))
# -
for inst in institutions:
for node in data:
date = node.get("date_published")
for a in node.get("authors"):
if a.get("DAfN") == inst["name"]:
if not inst.get("date_published") or inst.get("date_published") > date:
inst["date_published"] = date
print(institutions[0])
with open("institutions.json", "w") as f:
json.dump(institutions, f)
# !ls -lah institutions.json
| Decolonizing_Methodologies/geocode.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from keras import layers, models
def ANN_models_func(Nin, Nh, Nout):
x = layers.Input(shape=(Nin,))
h = layers.Activation('relu')(layers.Dense(Nh)(x))
y = layers.Activation('softmax')(layers.Dense(Nout)(h))
model = models.Model(x, y)
model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
print('ANN_models_func')
model.summary()
return model
def ANN_seq_func(Nin, Nh, Nout):
model = models.Sequential()
model.add(layers.Dense(Nh, activation='relu', input_shape=(Nin,)))
model.add(layers.Dense(Nout, activation='softmax'))
model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
print('ANN_seq_func')
model.summary()
return model
class ANN_models_class(models.Model):
def __init__(self, Nin, Nh, Nout):
x = layers.Input(shape=(Nin,))
h = layers.Activation('relu')(layers.Dense(Nh)(x))
y = layers.Activation('softmax')(layers.Dense(Nout)(h))
super().__init__(x, y)
self.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
print('ANN_models_class')
self.summary()
class ANN_seq_class(models.Sequential):
def __init__(self, Nin, Nh, Nout):
super().__init__()
self.add(layers.Dense(Nh, activation='relu', input_shape=(Nin,)))
self.add(layers.Dense(Nout, activation='softmax'))
self.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
print('ANN_seq_class')
self.summary()
import numpy as np
from keras import datasets
from keras.utils import np_utils
def Data_func():
(X_train, Y_train), (X_test, Y_test) = datasets.mnist.load_data()
Y_train = np_utils.to_categorical(Y_train)
Y_test = np_utils.to_categorical(Y_test)
L, W, H = X_train.shape
X_train = X_train.reshape(-1, W*H)
X_test = X_test.reshape(-1, W*H)
X_train = X_train / 255.0
X_test = X_test / 255.0
return (X_train, Y_train), (X_test, Y_test)
# %matplotlib inline
import matplotlib.pyplot as plt
def plot_loss(history):
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('Model Loss')
plt.ylabel('Loss')
plt.xlabel('Epoch')
plt.legend(['Train', 'Test'], loc=0)
def plot_acc(history):
plt.plot(history.history['acc'])
plt.plot(history.history['val_acc'])
plt.title('Model Accuracy')
plt.ylabel('Accuracy')
plt.xlabel('Epoch')
plt.legend(['Train', 'Test'], loc=0)
def main_model_func():
Nin = 784
Nh = 100
number_of_class = 10
Nout = number_of_class
model = ANN_models_func(Nin, Nh, Nout)
(X_train, Y_train), (X_test, Y_test) = Data_func()
history = model.fit(X_train, Y_train, epochs=15, batch_size=100, validation_split=0.2)
performance_test = model.evaluate(X_test, Y_test, batch_size=100)
print('Test Accuracy : ', performance_test)
plot_loss(history)
plt.show()
plot_acc(history)
plt.show()
def main_seq_func():
Nin = 784
Nh = 100
number_of_class = 10
Nout = number_of_class
model = ANN_seq_func(Nin, Nh, Nout)
(X_train, Y_train), (X_test, Y_test) = Data_func()
history = model.fit(X_train, Y_train, epochs=15, batch_size=100, validation_split=0.2)
performance_test = model.evaluate(X_test, Y_test, batch_size=100)
print('Test Accuracy : ', performance_test)
plot_loss(history)
plt.show()
plot_acc(history)
plt.show()
def main_model_class():
Nin = 784
Nh = 100
number_of_class = 10
Nout = number_of_class
model = ANN_models_class(Nin, Nh, Nout)
(X_train, Y_train), (X_test, Y_test) = Data_func()
history = model.fit(X_train, Y_train, epochs=15, batch_size=100, validation_split=0.2)
performance_test = model.evaluate(X_test, Y_test, batch_size=100)
print('Test Accuracy : ', performance_test)
plot_loss(history)
plt.show()
plot_acc(history)
plt.show()
def main_seq_class():
Nin = 784
Nh = 100
number_of_class = 10
Nout = number_of_class
model = ANN_seq_class(Nin, Nh, Nout)
(X_train, Y_train), (X_test, Y_test) = Data_func()
history = model.fit(X_train, Y_train, epochs=15, batch_size=100, validation_split=0.2)
performance_test = model.evaluate(X_test, Y_test, batch_size=100)
print('Test Accuracy : ', performance_test)
plot_loss(history)
plt.show()
plot_acc(history)
plt.show()
main_model_func()
main_seq_func()
main_model_class()
main_seq_class()
| 2020-02/2020-02-26-1-ANN-mnist.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **Variable aleatoria** es aquella cuyo posibles resultados dependen del resultado de un fenómeno aleatorio.
#
# Una **matriz de varianzas-covarianzas** es una matriz cuadrada que contiene las varianzas y covarianzas asociadas con diferentes variables. Los elementos de la diagonal de la matriz contienen las varianzas de las variables, mientras que los elementos que se encuentran fuera de la diagonal contienen las covarianzas entre todos los pares posibles de variables.
#
#
| Primer parcial .ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + init_cell=true
# %logstop
# %logstart -rtq ~/.logs/ml.py append
# %matplotlib inline
import matplotlib
import seaborn as sns
sns.set()
matplotlib.rcParams['figure.dpi'] = 144
# -
from static_grader import grader
# # ML Miniproject
# ## Introduction
#
# The objective of this miniproject is to exercise your ability to create effective machine learning models for making predictions. We will be working with nursing home inspection data from the United States, predicting which providers may be fined and for how much.
#
# ## Scoring
#
# In this miniproject you will often submit your model's `predict` or `predict_proba` method to the grader. The grader will assess the performance of your model using a scoring metric, comparing it against the score of a reference model. We will use the [average precision score](http://scikit-learn.org/stable/modules/generated/sklearn.metrics.average_precision_score.html). If your model performs better than the reference solution, then you can score higher than 1.0.
#
# **Note:** If you use an estimator that relies on random draws (like a `RandomForestClassifier`) you should set the `random_state=` to an integer so that your results are reproducible.
#
# ## Downloading the data
#
# We can download the data set from Amazon S3:
# + language="bash"
# mkdir data
# wget http://dataincubator-wqu.s3.amazonaws.com/mldata/providers-train.csv -nc -P ./ml-data
# wget http://dataincubator-wqu.s3.amazonaws.com/mldata/providers-metadata.csv -nc -P ./ml-data
# -
# We'll load the data into a Pandas DataFrame. Several columns will become target labels in future questions. Let's pop those columns out from the data, and drop related columns that are neither targets nor reasonable features (i.e. we don't wouldn't know how many times a facility denied payment before knowing whether it was fined).
#
# The data has many columns. We have also provided a data dictionary.
import numpy as np
import pandas as pd
metadata = pd.read_csv('./ml-data/providers-metadata.csv')
metadata.head()
# +
data = pd.read_csv('./ml-data/providers-train.csv', encoding='latin1')
fine_counts = data.pop('FINE_CNT')
fine_totals = data.pop('FINE_TOT')
cycle_2_score = data.pop('CYCLE_2_TOTAL_SCORE')
# -
data.head()
# ## Question 1: state_model
#
# A federal agency, Centers for Medicare and Medicaid Services (CMS), imposes regulations on nursing homes. However, nursing homes are inspected by state agencies for compliance with regulations, and fines for violations can vary widely between states.
#
# Let's develop a very simple initial model to predict the amount of fines a nursing home might expect to pay based on its location. Fill in the class definition of the custom estimator, `StateMeanEstimator`, below.
# +
from sklearn.base import BaseEstimator, RegressorMixin, TransformerMixin
class GroupMeanEstimator(BaseEstimator, RegressorMixin):
def __init__(self, grouper):
self.grouper = grouper
self.group_averages = {}
def fit(self, X, y):
# Use self.group_averages to store the average penalty by group
grouper_series = X[self.grouper]
frame = {self.grouper : grouper_series, 'y' : y}
pnlt_df = pd.DataFrame(frame)
for index, avg_pnlt in pnlt_df.groupby(self.grouper).mean().iterrows():
self.group_averages[index] = avg_pnlt.item()
return self
def predict(self, X):
# Return a list of predicted penalties based on group of samples in X
if not isinstance(X, pd.DataFrame):
X_df = pd.DataFrame(X)
else:
X_df = X
return [self.group_averages[item] if item in self.group_averages else 0.0 \
for _, item in X_df[self.grouper].items()]
# -
# After filling in class definition, we can create an instance of the estimator and fit it to the data.
# +
from sklearn.pipeline import Pipeline
state_model = Pipeline([
('sme', GroupMeanEstimator(grouper='STATE'))
])
state_model.fit(data, fine_totals)
# -
# Next we should test that our predict method works.
state_model.predict(data.sample(5))
# However, what if we have data from a nursing home in a state (or territory) of the US which is not in the training data?
state_model.predict(pd.DataFrame([{'STATE': 'AS'}]))
# Make sure your model can handle this possibility before submitting your model's predict method to the grader.
grader.score.ml__state_model(state_model.predict)
# ## Question 2: simple_features_model
#
# Nursing homes vary greatly in their business characteristics. Some are owned by the government or non-profits while others are run for profit. Some house a few dozen residents while others house hundreds. Some are located within hospitals and may work with more vulnerable populations. We will try to predict which facilities are fined based on their business characteristics.
#
# We'll begin with columns in our DataFrame containing numeric and boolean features. Some of the rows contain null values; estimators cannot handle null values so these must be imputed or dropped. We will create a `Pipeline` containing transformers that process these features, followed by an estimator.
#
# **Note:** When the grader checks your answer, it passes a list of dictionaries to the `predict` or `predict_proba` method of your estimator, not a DataFrame. This means that your model must work with both data types. For this reason, we've provided a custom `ColumnSelectTransformer` for you to use instead `scikit-learn`'s own `ColumnTransformer`.
# +
simple_cols = ['BEDCERT', 'RESTOT', 'INHOSP', 'CCRC_FACIL', 'SFF', 'CHOW_LAST_12MOS', 'SPRINKLER_STATUS', 'EXP_TOTAL', 'ADJ_TOTAL']
class ColumnSelectTransformer(BaseEstimator, TransformerMixin):
def __init__(self, columns):
self.columns = columns
def fit(self, X, y=None):
return self
def transform(self, X):
if not isinstance(X, pd.DataFrame):
X = pd.DataFrame(X)
return X[self.columns].to_numpy()
simple_features = Pipeline([
('cst', ColumnSelectTransformer(simple_cols)),
])
# -
# **Note:** The assertion below assumes the output of `noncategorical_features.fit_transform` is a `ndarray`, not a `DataFrame`.)
assert data['RESTOT'].isnull().sum() > 0
#assert not np.isnan(simple_features.fit_transform(data)).any()
# Now combine the `simple_features` pipeline with an estimator in a new pipeline. Fit `simple_features_model` to the data and submit `simple_features_model.predict_proba` to the grader. You may wish to use cross-validation to tune the hyperparameters of your model.
# +
from sklearn.preprocessing import StandardScaler
from sklearn.linear_model import LogisticRegression
from sklearn.impute import SimpleImputer
simple_features_model = Pipeline([
('simple', simple_features),
# add your estimator here
('imputer', SimpleImputer(missing_values=np.nan, strategy='mean')),
('scaler', StandardScaler()),
('estimator', LogisticRegression())
])
# -
simple_features_model.fit(data, fine_counts > 0)
# +
def positive_probability(model):
def predict_proba(X):
return model.predict_proba(X)[:, 1]
return predict_proba
grader.score.ml__simple_features(positive_probability(simple_features_model))
# -
# ## Question 3: categorical_features
# The `'OWNERSHIP'` and `'CERTIFICATION'` columns contain categorical data. We will have to encode the categorical data into numerical features before we pass them to an estimator. Construct one or more pipelines for this purpose. Transformers such as [LabelEncoder](https://scikit-learn.org/0.19/modules/generated/sklearn.preprocessing.LabelEncoder.html#sklearn.preprocessing.LabelEncoder) and [OneHotEncoder](https://scikit-learn.org/0.19/modules/generated/sklearn.preprocessing.OneHotEncoder.html#sklearn.preprocessing.OneHotEncoder) may be useful, but you may also want to define your own transformers.
#
# If you used more than one `Pipeline`, combine them with a `FeatureUnion`. As in Question 2, we will combine this with an estimator, fit it, and submit the `predict_proba` method to the grader.
# +
from sklearn.pipeline import FeatureUnion
from sklearn.preprocessing import OneHotEncoder
owner_onehot = Pipeline([
('cst', ColumnSelectTransformer(['OWNERSHIP'])),
('onehotenc', OneHotEncoder())
])
cert_onehot = Pipeline([
('cst', ColumnSelectTransformer(['CERTIFICATION'])),
('onehotenc', OneHotEncoder())
])
categorical_features = FeatureUnion([('owner_onehot', owner_onehot),
('cert_onehot', cert_onehot)
])
# -
assert categorical_features.fit_transform(data).shape[0] == data.shape[0]
assert categorical_features.fit_transform(data).dtype == np.float64
#assert not np.isnan(categorical_features.fit_transform(data)).any()
# As in the previous question, create a model using the `categorical_features`, fit it to the data, and submit its `predict_proba` method to the grader.
categorical_features_model = Pipeline([
('categorical', categorical_features),
# add your estimator here
('estimator', LogisticRegression())
])
categorical_features_model.fit(data, fine_counts > 0)
grader.score.ml__categorical_features(positive_probability(categorical_features_model))
# ## Question 4: business_model
# Finally, we'll combine `simple_features` and `categorical_features` in a `FeatureUnion`, followed by an estimator in a `Pipeline`. You may want to optimize the hyperparameters of your estimator using cross-validation or try engineering new features (e.g. see [PolynomialFeatures](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.PolynomialFeatures.html)). When you've assembled and trained your model, pass the `predict_proba` method to the grader.
# +
simple_features = Pipeline([
('cst', ColumnSelectTransformer(simple_cols)),
('imputer', SimpleImputer(missing_values=np.nan, strategy='mean')),
('scaler', StandardScaler()),
])
business_features = FeatureUnion([
('simple', simple_features),
('categorical', categorical_features)
])
# +
business_model = Pipeline([
('features', business_features),
# add your estimator here
('estimator', LogisticRegression())
])
# -
business_model.fit(data, fine_counts > 0)
grader.score.ml__business_model(positive_probability(business_model))
# ## Question 5: survey_results
# Surveys reveal safety and health deficiencies at nursing homes that may indicate risk for incidents (and penalties). CMS routinely makes surveys of nursing homes. Build a model that combines the `business_features` of each facility with its cycle 1 survey results, as well as the time between the cycle 1 and cycle 2 survey to predict the cycle 2 total score.
#
# First, let's create a transformer to calculate the difference in time between the cycle 1 and cycle 2 surveys.
class TimedeltaTransformer(BaseEstimator, TransformerMixin):
def __init__(self, t1_col, t2_col):
self.t1_col = t1_col
self.t2_col = t2_col
def fit(self, X, y=None):
return self
def transform(self, X):
if not isinstance(X, pd.DataFrame):
X_df = pd.DataFrame(X)
else:
X_df = X
t1_date = pd.to_datetime(X_df[self.t1_col])
t2_date = pd.to_datetime(X_df[self.t2_col])
duration_series = t1_date - t2_date
duration_array = []
for item in duration_series:
duration_array.append(item.days)
return np.array(duration_array).reshape(-1,1)
cycle_1_date = 'CYCLE_1_SURVEY_DATE'
cycle_2_date = 'CYCLE_2_SURVEY_DATE'
time_feature = TimedeltaTransformer(cycle_1_date, cycle_2_date)
# In the cell below we'll collect the cycle 1 survey features.
cycle_1_cols = ['CYCLE_1_DEFS', 'CYCLE_1_NFROMDEFS', 'CYCLE_1_NFROMCOMP',
'CYCLE_1_DEFS_SCORE', 'CYCLE_1_NUMREVIS',
'CYCLE_1_REVISIT_SCORE', 'CYCLE_1_TOTAL_SCORE']
cycle_1_features = ColumnSelectTransformer(cycle_1_cols)
# +
from sklearn.ensemble import RandomForestRegressor
from sklearn.decomposition import TruncatedSVD
from sklearn.linear_model import Lasso
from sklearn.model_selection import GridSearchCV
from sklearn.pipeline import FeatureUnion
from sklearn.preprocessing import PolynomialFeatures
survey_model = Pipeline([
('features', FeatureUnion([
('business', business_features),
('survey', cycle_1_features),
('time', time_feature)
])),
# add your estimator here
('poly', PolynomialFeatures(2)),
('decomp', TruncatedSVD(20)),
('estimator', RandomForestRegressor(random_state = 0,
n_estimators=200,
min_samples_leaf = 100))
])
# +
from sklearn.ensemble import RandomForestRegressor
from sklearn.decomposition import TruncatedSVD
from sklearn.linear_model import Lasso
from sklearn.model_selection import GridSearchCV
from sklearn.pipeline import FeatureUnion
from sklearn.preprocessing import PolynomialFeatures
#gs = GridSearchCV(Lasso(max_iter = 1000), param_grid = {'alpha': np.arange(0, 3.5, 0.5)}, cv=5, n_jobs=4, verbose=0)
survey_model = Pipeline([
('features', FeatureUnion([
('business', business_features),
('survey', cycle_1_features),
('time', time_feature)
])),
# add your estimator here
('poly', PolynomialFeatures(2)),
('decomp', TruncatedSVD(20)),
("rf",RandomForestRegressor(random_state = 0,n_estimators=200, min_samples_leaf = 100))
])
# -
survey_model.fit(data, cycle_2_score.astype(int))
grader.score.ml__survey_model(survey_model.predict)
# *Copyright © 2020 The Data Incubator. All rights reserved.*
| miniprojects/.ipynb_checkpoints/ml-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="fantastic-department"
# # The Actor-Critic Method
# <table align="left">
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/matyama/deep-rl-hands-on/blob/main/12_a2c.ipynb">
# <img src="https://www.tensorflow.org/images/colab_logo_32px.png" />
# Run in Google Colab
# </a>
# </td>
# </table>
# + colab={"base_uri": "https://localhost:8080/"} id="buried-kennedy" outputId="647555a6-581d-447e-e7cf-b53bf9a7249d" language="bash"
# !(stat -t /usr/local/lib/*/dist-packages/google/colab > /dev/null 2>&1) && exit
#
# echo "Running on Google Colab, therefore installing dependencies..."
# pip install ptan>=0.7 tensorboardX
# + [markdown] id="accepted-papua"
# ## Variance Reduction
# Let's start by recalling the policy gradient defined by the *Policy Gradients (PG)* method:
# $$
# \nabla J \approx \mathbb{E}[Q(s, a) \nabla \log(\pi(a|s))]
# $$
#
# One of the weak points of the PG method is that the gradient scales $Q(s, a)$ may experience quite significant variance* which does not help the training at all. We fixed this issue by introducing a fixed *baseline* value (e.g. mean reward) that was subtracted from the gradient scales Q.
#
# \* Recall formal defintion: $\mathbb{V}[X] = \mathbb{E}[(X - \mathbb{E}[X])^2]$
#
# Let's illustrate this problem and solution on simple example:
# * Assume there are three actions with $Q_1$, $Q_2$ some small positive values and $Q_3$ being large negative
# * In this case there will be small positive gradient towards fist two actions and large negative one repelling the policy from the third one
# * Now imagine $Q_1$ and $Q_2$ were large positive values instead. Then $Q_3$ would become small but positive value. The gradient would still push the policy towards fist two actions but it would direct the gradient towards the trird one a bit as well (instead of pushing it away from it)!
#
# Now it's a bit more clear why subtracting a constant value that we called the *baseline* helps.
# + [markdown] id="opposed-leone"
# ## Advantage Actor-Critic (A2C)
# *Advantage Actor-Critic (A2C)* method can be viewed as a combination of PG and DQN with a simple idea extending the variance reduction theme we discussed above. Until now we treated the *baseline* value as single constant that we subtracted from all $Q(s, a)$ values. A2C pushes this further and uses different baselines for each state $s$.
#
# If one recalls the *Duelling DQN* which exploited the fact that $Q(s, a) = V(s) + A(s, a)$ - i.e. state-action values are composed of a *baseline* state values $V(s)$ and action advantages $A(s, a)$ in these states, it is quite straightforward to figure out which values A2C uses as state baselines - the state values $V(s)$!
#
# The *Advantage* Actor-Critic name then comes from the fact that our gradient scales turn to action advantages after subtracting state values:
# $$
# \mathbb{E}[Q(s, a) \nabla \log(\pi(a|s))] \to \mathbb{E}[A(s, a) \nabla \log(\pi(a|s))]
# $$
#
# Finally, the question is how do we obtain $V(s)$? Here comes the second part which is the combination with the DQN approach - we simply train a DQN alongside our PGN.
#
# *Notes*:
# * *There'll actually be just single NN that will learn both the policy and state values (discussed below)*
# * *Improvements from both methods are still applicable (also metioned and shown in following sections)*
# + [markdown] id="assigned-cisco"
# ### Common Imports
# + id="mechanical-expert"
# flake8: noqa: E402,I001
import time
from typing import Any, List, Sequence, Tuple
import gym
import numpy as np
import ptan
import torch
import torch.nn as nn
from ptan.experience import ExperienceFirstLast
from tensorboardX import SummaryWriter
# + [markdown] id="further-tomorrow"
# ### Reward Tracker
# + id="checked-bones"
class RewardTracker:
def __init__(
self,
writer: SummaryWriter,
stop_reward: float,
window_size: int = 100,
) -> None:
self.writer = writer
self.stop_reward = stop_reward
self.window_size = window_size
self.best_mean_reward = float("-inf")
def __enter__(self) -> "RewardTracker":
self.ts = time.time()
self.ts_frame = 0
self.total_rewards = []
return self
def __exit__(self, *args: Any) -> None:
self.writer.close()
def add_reward(self, reward: float, frame: int) -> bool:
"""
Returns an indication of whether a termination contition was reached.
"""
self.total_rewards.append(reward)
fps = (frame - self.ts_frame) / (time.time() - self.ts)
self.ts_frame = frame
self.ts = time.time()
mean_reward = np.mean(self.total_rewards[-self.window_size :])
if mean_reward > self.best_mean_reward:
self.best_mean_reward = mean_reward
print(
f"{frame}: done {len(self.total_rewards)} games, "
f"mean reward {mean_reward:.3f}, speed {fps:.2f} fps"
)
self.writer.add_scalar("fps", fps, frame)
self.writer.add_scalar("reward_100", mean_reward, frame)
self.writer.add_scalar("reward", reward, frame)
return mean_reward > self.stop_reward
# + [markdown] id="australian-brake"
# ### Atari A2C PG Network
# This NN is quite similar to the *Dueling DQN* architecture but with an important difference. In the Dueling DQN we have also two parts
# 1. Part for the state values $V(s)$
# 1. Part for the action advantages $A(s, a)$
#
# But as with any other DQN we did still output $Q(s, a) = V(s) + A(s, a)$. Here we have two separate outputs with common base network:
# 1. Policy network that outputs action logits - basically policy $\pi(a|s)$ when one converts them to probabilities using softmax
# 1. Value network which computes $V(s)$
# + id="killing-lloyd"
class AtariA2C(nn.Module):
"""
A2C network with 2D convolutional base for Atari envs. and two heads:
1. Policy - dense network that outputs action logits
2. Value - dence network that models state values `V(s)`
"""
def __init__(self, input_shape: Tuple[int, ...], n_actions: int) -> None:
super().__init__()
# 2D conv. base network common to both heads
# - This way both nets share commonly learned basic features
# - Also helps with convergence (compared to having two separate NNs)
self.conv = nn.Sequential(
nn.Conv2d(input_shape[0], 32, kernel_size=8, stride=4),
nn.ReLU(),
nn.Conv2d(32, 64, kernel_size=4, stride=2),
nn.ReLU(),
nn.Conv2d(64, 64, kernel_size=3, stride=1),
nn.ReLU(),
)
conv_out_size = self._get_conv_out(input_shape)
# Policy NN - outputs action logits
self.policy = nn.Sequential(
nn.Linear(conv_out_size, 512),
nn.ReLU(),
nn.Linear(512, n_actions),
)
# Value NN - outputs state value
self.value = nn.Sequential(
nn.Linear(conv_out_size, 512),
nn.ReLU(),
nn.Linear(512, 1),
)
def _get_conv_out(self, shape: Tuple[int, ...]) -> int:
o = self.conv(torch.zeros(1, *shape))
return int(np.prod(o.size()))
def forward(self, x: torch.Tensor) -> Tuple[torch.Tensor, torch.Tensor]:
"""
For an input batch of states, returns pair of tensors
1. Policy logits for all actions in these states
2. Values of these states
"""
inputs = x.float() / 256
conv_out = self.conv(inputs).view(inputs.size()[0], -1)
return self.policy(conv_out), self.value(conv_out)
# + [markdown] id="EkrZlVejr5Od"
# ### Batch Unpacking
# Similarly to implementations of other methods, we will use a function that unpacks an experience batch into its components.
#
# One important difference here is that we'll use the A2C NN to evaluate final states from the batch to get $V(s_N)$ - here we assume that we make $N$ steps ahead in the environmet/environments.
#
# Using all the experienced rewards $r_i$ from the batch we can compute target Q values as
# $$
# Q(s, a) = \sum_{i = 0}^{N - 1} \gamma^i r_i + \gamma^T V(s_N)
# $$
# * First part (the sum) is the total discounted reward from all but last steps
# * The second is the discounted future reward from the $N$-th step (predicted by the NN)
# + id="parliamentary-posting"
def unpack_batch(
batch: Sequence[ExperienceFirstLast],
net: AtariA2C,
gamma: float,
reward_steps: int,
device: str = "cpu",
) -> Tuple[torch.FloatTensor, torch.LongTensor, torch.FloatTensor]:
"""
Convert batch into training tensors
:param batch: Experiences from environment(s)
:param net: A2C network that can approximate state values
:returns: states, actions, target Q values as tensors
"""
states, actions, rewards, not_done_exps, last_states = [], [], [], [], []
# Unwrap each transition from the batch
# - And mark entries from unfinished episodes
for i, exp in enumerate(batch):
states.append(np.array(exp.state, copy=False))
actions.append(int(exp.action))
rewards.append(exp.reward)
if exp.last_state is not None:
not_done_exps.append(i)
last_states.append(np.array(exp.last_state, copy=False))
# Note: Wrapping states into np array is to fix PyTorch performance issue
# Convert states and actions to tensors
states = torch.FloatTensor(np.array(states, copy=False)).to(device)
actions = torch.LongTensor(actions).to(device)
# Compute target state values V(s) for training the net
# - Uses given A2C NN to predit the state values
# Init target Q(s, a) to (discounted) rewards
# - This will be the final value at the end of an episode
target_values = np.array(rewards, dtype=np.float32)
if not_done_exps:
# Convert next states to a tensor
last_states = torch.FloatTensor(np.array(last_states, copy=False)).to(
device
)
# Use given A2C net to predict future values
_, last_state_values = net(last_states)
last_state_values = last_state_values.data.cpu().numpy()[:, 0]
# Add future values to the discounted rewards if episode is not done
last_state_values *= gamma ** reward_steps
target_values[not_done_exps] += last_state_values
# Convert target values to a tensor
target_values = torch.FloatTensor(target_values).to(device)
return states, actions, target_values
# + [markdown] id="bXh-fSpB2WQR"
# ### A2C Training *(Atari Pong)*
# The training loop build on and extends the PG example from previous chapter.
#
# The loss function now has three components:
# 1. **Policy loss** - similar to the PG method but with the gradient scales $A(s, a) = Q(s, a) - V(s)$ where $Q(s, a)$ are obtained from the batch unpacking described above (i.e. from experienced rewards and net's $V(s')$) and $V(s)$ is net's prediction for current state(s)
# 1. **Value loss** - simply a MSE between current values $V(s)$ and TD targets (the same values we used for policy loss from the experience batch)
# 1. **Entropy bonus** - the same techinque used for PG that adds an entropy component $\mathcal{L}_H = \beta \sum_i \pi(s_i) \log(\pi(s_i))$ that pushes the policy more towards uniform distribution that favours exploration
#
# Finally, we'll use multiple copies of the same environment to sample our experience batch from. This is the same techinque to break correlations that was used for the vanilla PG method. Thre are two basic variants of the *Actor-Critic* method:
# * A2C which uses parallel environments with synchronized policy gradient updates
# * A3C which stands for *Asynchronous Advantage Actor-Critic* and will be described in the next chapter
# + id="nb78PWxxGjcO"
# %load_ext tensorboard
# %tensorboard --logdir runs
# + id="HcCcZdSOyvMl"
# Hyperparameters
GAMMA = 0.99
LEARNING_RATE = 0.001
ADAM_EPS = 1e-3
ENTROPY_BETA = 0.01
BATCH_SIZE = 128
NUM_ENVS = 50
REWARD_STEPS = 4
CLIP_GRAD = 0.1
STOP_REWARD = 18
SEED = 42
# Set RNG state
np.random.seed(SEED)
torch.manual_seed(SEED)
if torch.cuda.is_available():
torch.cuda.manual_seed(SEED)
# Determine where the computations will take place
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
# Make multiple instances of the Atari Pong environment
def make_pong_env(i: int, seed: int) -> gym.Env:
env = ptan.common.wrappers.wrap_dqn(gym.make("PongNoFrameskip-v4"))
env.seed(seed + i)
return env
envs = [make_pong_env(i, seed=SEED) for i in range(NUM_ENVS)]
# Create the A2C network for Atari environments
net = AtariA2C(
input_shape=envs[0].observation_space.shape,
n_actions=envs[0].action_space.n,
).to(device)
print(net)
# Initialize the policy agent
# - Instead of passing the whole NN, we use a callback over it returning just
# the action logits
agent = ptan.agent.PolicyAgent(
model=lambda x: net(x)[0],
apply_softmax=True,
device=device,
)
# Create `REWARD_STEPS`-ahead experience source over all environments
exp_source = ptan.experience.ExperienceSourceFirstLast(
env=envs,
agent=agent,
gamma=GAMMA,
steps_count=REWARD_STEPS,
)
# Create Adam optimizer
# - Note: We use larger epsilon to make the training converge
optimizer = torch.optim.Adam(net.parameters(), lr=LEARNING_RATE, eps=ADAM_EPS)
# Create TensorBoard writer for metrics collection
writer = SummaryWriter(comment="-pong-a2c")
# Create TensorBoard trackers
with RewardTracker(writer, stop_reward=STOP_REWARD) as tracker:
with ptan.common.utils.TBMeanTracker(writer, batch_size=10) as tb_tracker:
batch = []
# Run the training loop consuming experiences from the source
for i, exp in enumerate(exp_source):
# Add new experience to current batch
batch.append(exp)
new_rewards = exp_source.pop_total_rewards()
if new_rewards:
# Record new reward and check for termination
solved = tracker.add_reward(reward=new_rewards[0], frame=i)
# Stop if the mean reward was good enough
if solved:
print(f"Solved in {i} steps!")
break
# Let the batch fill up
if len(batch) < BATCH_SIZE:
continue
# Unpack and clear current batch
states, actions, target_values = unpack_batch(
batch=batch,
net=net,
gamma=GAMMA,
reward_steps=REWARD_STEPS,
device=device,
)
batch.clear()
# Clear gradients
optimizer.zero_grad()
# Compute action logits and values of current states
action_logits, values = net(states)
# Compute V(s) part of the loss function
value_loss = nn.functional.mse_loss(
input=values.squeeze(-1),
target=target_values,
)
# Compute the policy part of the loss function
# - We use A(s, a) = Q(s, a) - V(s) as the gradient scales
# - Note: We detach values from the autograph to stop grad flow
log_action_prob = nn.functional.log_softmax(action_logits, dim=1)
advantage = target_values - values.detach()
scaled_log_action_prob = (
advantage * log_action_prob[range(BATCH_SIZE), actions]
)
policy_loss = -scaled_log_action_prob.mean()
# Compute entropy bonus to the loss function
action_prob = nn.functional.softmax(action_logits, dim=1)
entropy_loss = (
ENTROPY_BETA
* (action_prob * log_action_prob).sum(dim=1).mean()
)
# First calculate policy gradients only
policy_loss.backward(retain_graph=True)
grads = np.concatenate(
[
param.grad.data.cpu().numpy().flatten()
for param in net.parameters()
if param.grad is not None
]
)
# Apply entropy and value gradients
loss = entropy_loss + value_loss
loss.backward()
# Use gradient clipping (by l2 norm) before making next step
nn.utils.clip_grad_norm_(net.parameters(), CLIP_GRAD)
optimizer.step()
# Get total loss for tracking
loss += policy_loss
# Track metrics
tb_tracker.track("advantage", advantage, i)
tb_tracker.track("values", values, i)
tb_tracker.track("batch_rewards", target_values, i)
tb_tracker.track("loss_entropy", entropy_loss, i)
tb_tracker.track("loss_policy", policy_loss, i)
tb_tracker.track("loss_value", value_loss, i)
tb_tracker.track("loss_total", loss, i)
tb_tracker.track("grad_l2", np.sqrt(np.mean(np.square(grads))), i)
tb_tracker.track("grad_max", np.max(np.abs(grads)), i)
tb_tracker.track("grad_var", np.var(grads), i)
# + id="MU2Lpp0aHwc1"
| 12_a2c.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.0 64-bit
# language: python
# name: python38064bit1060d4750c904259afeb7847dfa8ded2
# ---
import os
import pandas as pd
import numpy as np
from sklearn.preprocessing import LabelEncoder
from sklearn.preprocessing import MinMaxScaler
import tensorflow as tf
import pickle
os.chdir('../')
os.chdir('scripts')
import data_creation_v3 as d
os.chdir('../')
os.chdir('models')
order = ['bodyLength', 'bscr', 'dse', 'dsr', 'entropy', 'hasHttp', 'hasHttps',
'has_ip', 'numDigits', 'numImages', 'numLinks', 'numParams',
'numTitles', 'num_%20', 'num_@', 'sbr', 'scriptLength', 'specialChars',
'sscr', 'urlIsLive', 'urlLength']
a = d.UrlFeaturizer('http://astore.amazon.co.uk/allezvinsfrenchr/detail/1904010202/026-8324244-9330038').run()
test = []
for i in order:
test.append(a[i])
encoder = LabelEncoder()
encoder.classes_ = np.load('lblenc.npy',allow_pickle=True)
scalerfile = 'scaler.sav'
scaler = pickle.load(open(scalerfile, 'rb'))
test = pd.DataFrame(test).replace(True,1).replace(False,0).to_numpy(dtype="float32").reshape(1,-1)
test = scaler.transform(test)
# +
# Load the TFLite model and allocate tensors.
interpreter = tf.lite.Interpreter(model_path="tflite_quant_model.tflite")
interpreter.allocate_tensors()
# Get input and output tensors.
input_details = interpreter.get_input_details()
output_details = interpreter.get_output_details()
interpreter.set_tensor(input_details[0]['index'], test)
interpreter.invoke()
# The function `get_tensor()` returns a copy of the tensor data.
# Use `tensor()` in order to get a pointer to the tensor.
output_data = interpreter.get_tensor(output_details[0]['index'])
predicted = np.argmax(output_data,axis=1)
# -
print(encoder.inverse_transform(predicted)[0])
| Notebook/TFLite.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Using AWS S3 to read/write market data with findatapy
#
# May 2021 - <NAME> - https://www.cuemacro.com - <EMAIL>
# ## What is S3?
#
# S3 is basically storage in the cloud, which is managed by AWS. Dump as much data as want from anywhere on the web and you don't need to worry about scaling your storage, which you'd obviously have to do in your own data centre, and also manage backups. Data is stored in S3 buckets, which are a bit like folders. Google Cloud Storage (GCS) is the equivalent on Google Cloud and Azure Blob is a similar service on Azure.
# ## What is the cost of S3?
# There are many other AWS storage services which you can find at https://aws.amazon.com/products/storage/, which are at different price levels and performance too. It is important to use the right services for storage which have the right performance cost balance for your specific use cases.
#
# The cost of S3 depends upon factors like:
#
# * how much data you store?
# * which service you use (are you using S3 Standard Storage for example, or S3 Infrequent Access Storage)?
# * how many requests you make for the data (PUT/GET etc.)?
# * which region is it in?
# * how much data you transfer from S3 out to the internet?
#
# An article on cloudhealthtech.com goes through the various ins and outs of the pricing at https://www.cloudhealthtech.com/blog/s3-cost-aws-cloud-storage-costs-explained. They note that the storage cost for Standard S3 (Nov 2020) is around 0.021 to 0.026 USD per month per GB. So for 1 TB that's around 21-26 USD per month, roughly under 300 USD per year. This excludes any of the various request costs for example, which you need to take into account. The actual cost of a hardware is a lot cheaper (a quick browse of 1 TB drives online, suggested a cost of around 50 USD), but if we manage our own hardware, we need to take into hassle of managing it, including stuff like backup, convenience of access etc. The cost of losing data is likely to be significant if we choose to host our data locally.
# ## Making AWS accessible via Python
#
# Given that S3 is in the cloud, we need to make sure that AWS services need to be accessible from Python, whether we are running our process in the cloud (which seems preferable to reduce latency) or locally. Whilst we are using Python, S3 is also accessible from many other languages.
#
# * Hence, before going through this tutorial, you'll need to go through several steps so AWS services are accessible from your machine
# * You'll need to create an IAM user, with appropriate permissions at https://console.aws.amazon.com/iam when you are logged into the AWS Console
# * In our case this will to have permissions to use S3
# * Get the Access key ID and secret access key for the IAM user
# * If you want to make S3 accessible to users outside of your AWS account, I found this explanation at https://stackoverflow.com/questions/45336781/amazon-s3-access-for-other-aws-accounts
# * Before changing any access rights to S3, I'd strongly recommend reading https://aws.amazon.com/s3/security/ which explains the various security mechanisms including being able to block any public access at all
# * Install AWS CLI
# * run `sudo apt install awscli`
# * or you can download the zip file
# * run `aws configure` to set the default access key ID, default AWS availability zone etc.
# * this will create files in ~/.aws/credentials and ~/.aws/config
# * AWS CLI instructions at https://docs.aws.amazon.com/cli/latest/userguide/install-cliv2-linux.html#cliv2-linux-install
#
# * Once your credentials are set, we can use boto3, which is an SDK for Python developers to access AWS resources:
# * `boto3` instructions https://boto3.amazonaws.com/v1/documentation/api/latest/index.html
# * You can install `boto3` using pip
# * You also need to install `s3fs` using pip to get access to S3 via Python
# * If you follow the instructions at https://github.com/cuemacro/teaching/blob/master/pythoncourse/installation/installing_anaconda_and_pycharm.ipynb - you'll create a conda environment `py38class` which includes boto3, and many useful data science libraries, which I use for my Python teaching
# * You may need to install the latest version of findatapy from GitHub using `pip install git+https://github.com/cuemacro/findatapy.git` to run the code below.
# ## Creating your bucket on S3
#
# You can create your S3 bucket using AWS CLI (see https://docs.aws.amazon.com/cli/latest/reference/s3api/create-bucket.html). In the below example we create our bucket called `my-bucket` in the AWS region `us-east-1`
#
# `aws s3api create-bucket --bucket my-bucket --region us-east-1`
#
# Alternatively, you can also create it via the web GUI at https://s3.console.aws.amazon.com/s3/
# ## Using S3 with findatapy to store tick market data from Dukascopy
#
# In this notebook I'm going to show how to use S3 to easily store market data using findatapy. We are assuming that we have already setup AWS CLI with our credentials such as our access key ID etc.
#
# As a first step let's download some tick data from Dukascopy for EURUSD spot, which is a free data source using findatapy. Findatapy provides a uniform wrapper to download from many different data sources. We can predefine ticker mappings from our own nicknames for tickers to the vendor tickers. It already comes out of the box, with Dukascopy ticker mappings predefined, but these are all customisable. Note, that we haven't used the `data_engine` property. If this isn't set, then findatapy will download from our data source directly.
# First disable the log so the output is neater
import logging, sys
logging.disable(sys.maxsize)
# +
from findatapy.market import Market, MarketDataRequest
# In this case we are saving predefined tick tickers to disk, and then reading back
from findatapy.market.ioengine import IOEngine
md_request = MarketDataRequest(
start_date='04 Jan 2021',
finish_date='05 Jan 2021',
category='fx',
data_source='dukascopy',
freq='tick',
tickers=['EURUSD'],
fields=['bid', 'ask', 'bidv', 'askv'],
data_engine=None
)
market = Market()
df = market.fetch_market(md_request=md_request)
# -
# Let's print the output...
print(df)
# Let's type in our S3 bucket address, which you'll need to change below. Note the use of `s3://` at the start of the expression.
folder = 's3://my-bucket'
# We can write our tick data DataFrame in Parquet format. We can give it the `MarketDataRequest` we used for fetching the data, which basically creates the filename in the format of `environment.category.data_source.freq.tickers` for high frequency data or in the format of `environment.category.data_source.freq` for daily data. This will enable us to more easily fetch the data using the same `MarketDataRequest` interface.
#
# In this case, the filename of the Parquet file is:
#
# * `s3://bla_bla_bla/backtest.fx.tick.dukascopy.NYC.EURUSD.parquet`
# * ie. the environment of our data is `backtest`
# * the `category` is `fx`
# * the `data_source` is `dukascopy`
# * the `freq` is `tick`
# * the `cut` (or time of close) is `NYC`
# * the `tickers` is `EURUSD`
#
# The Jupyter notebook [market_data_example.ipynb](../market_data_example.ipynb) explains in more detail this ticker format and the concept of a `MarketDataRequest`. We dump it disk using the `IOEngine` class. Note that the `write_time_series_cache_to_disk` and `read_time_series_from_disk` reads/writes from S3 in exactly the same way as we would do locally. We need to make sure that when we're writing to disk, we have a data licence to do so (and this will clearly vary between data vendors), and in particular, that only those who read from the disk are authorised to use that data.
IOEngine().write_time_series_cache_to_disk(folder, df, engine='parquet', md_request=md_request)
# We could fetch the data directly using the S3 filename ie.
# +
s3_filename = folder + '/backtest.fx.dukascopy.tick.NYC.EURUSD.parquet'
df = IOEngine().read_time_series_cache_from_disk(s3_filename, engine='parquet')
print(df)
# -
# But it is more convenient to simply use the `MarketDataRequest` object we populated earlier. But in order to make it fetch from S3 instead of Dukascopy, we just need to set the `data_engine` property to give it the path of the S3 bucket and the postfix `/*.parquet`.
# +
md_request.data_engine = folder + '/*.parquet'
df = market.fetch_market(md_request)
print(df)
# -
# It should be noted there are many other ways to dump and read Parquet files from S3. We can use `pandas.read_parquet` to directly read Parquet files from S3. Libraries like Dask also support reading Parquet directly from S3 too. I'd also checkout AWS Data Wrangler, which makes it easier to use Pandas with many AWS services including S3, at https://github.com/awslabs/aws-data-wrangler
# ## Using S3 with findatapy to store daily market data from Quandl
#
# In this case we are downloading all G10 FX crosses from Quandl, which are predefined as `fx.quandl.daily.NYC` where our
# * `category` is `fx`
# * `data_source` is `quandl`
# * `freq` is `daily`
# * `cut` is `NYC`
#
# Unlike in the previous where we specified the `MarketDataRequest` in full, here we just use the above string as shorthand, and we set the `quandl_api_key` using the `MarketDataRequest` and also the `start_date`. If we have Redis running locally (which is an in memory cache), this DataFrame will also be cached in our Redis instance. We'll show how to take advantage of this cache later. If Redis is not installed, it's not a big deal, it just means the cache won't be operational. Hence, doing repeated `MarketDataRequest` calls will end up taking longer, as findatapy will seek to get the data from the data vendor directly.
# +
import os
from findatapy.market import Market, MarketDataRequest
# In this case we are saving predefined tick tickers to disk, and then reading back
from findatapy.market.ioengine import IOEngine
# Change this to your own Quandl API key
quandl_api_key = os.environ['QUANDL_API_KEY']
md_request = market.create_md_request_from_str(md_request_str='fx.quandl.daily.NYC',
md_request=MarketDataRequest(start_date='01 Jan 2021', finish_date='27 May 2021', quandl_api_key=quandl_api_key))
# -
# We can print out the `MarketDataRequest` we just constructed. We should be able to see there `quandl` for the `data_source` and `fx` for the `category`, as well as the `start_date` (realise it's difficult!)
print(md_request)
# We can now fetch the market data from `quandl`.
# +
df = market.fetch_market(md_request)
print(df)
# -
# Let's write this to S3.
IOEngine().write_time_series_cache_to_disk(folder, df, engine='parquet', md_request=md_request)
# And we can read it back using a similar call to before, except this time we set the `data_engine` property of the `MarketDataRequest`.
# +
df = market.fetch_market(md_request_str='fx.quandl.daily.NYC',
md_request=MarketDataRequest(start_date='01 Jan 2021', finish_date='27 May 2021',
quandl_api_key=quandl_api_key,
data_engine=folder + '/*.parquet'))
print(df)
# -
# If we set the `cache_algo` property to `cache_algo_return` and remove the `data_engine` parameter (and if we have had Redis running, and make exactly the same data requet call (same assets and dates), findatapy will look in the Redis cache locally to fetch the data. This is significantly quicker at around 20ms versus over 500ms for fetching from S3.
# +
df = market.fetch_market(md_request_str='fx.quandl.daily.NYC',
md_request=MarketDataRequest(start_date='01 Jan 2021', finish_date='27 May 2021',
quandl_api_key=quandl_api_key,
cache_algo='cache_algo_return'))
print(df)
# -
# ## Conclusion
# We have seen that it's pretty straightforward to store market data as Parquet files in S3 using findatapy. In particular, it makes it easy to use very similar `MarketDataRequest` calls we would use to fetch data from the `data_source` itself, as it is from S3 (or indeed from any local disk drive). We just need to be sure to set the `data_engine` property of the `MarketDataRequest`.
#
# We also briefly showed how to take advantage of in memory caching with Redis.
| finmarketpy_examples/finmarketpy_notebooks/s3_bucket_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:Anaconda3]
# language: python
# name: conda-env-Anaconda3-py
# ---
# # Introduction to Deployment
#
# This section is as platform-agnostic as possible but application part focuses on Amazon Web Services (AWS).
#
# Contents
#
# - Cloud Computing
# - Machine Learning in the Workplace
# - Deployment
#
# After getting familiar with machine learning deployment we'll put these ideas to practice using [Amazon SageMaker](https://aws.amazon.com/sagemaker/) as one way to deploy machine learning models.
#
# Questions to answer:
#
# - What is the machine learning workflow?
# - How does **deployment** fit into the machine learning workflow?
# - What is cloud computing?
# - Why are we using cloud computing for deploying machine learning models?
# - Why isn't deployment a part of many machine learning curriculums?
# - What does it mean for a model to be deployed?
# - What are the essentail characteristics associated with the code of deployed models?
# - What are different cloud computing platform we might use to deploy our machine learning models?
#
# ## Machine Learning Workflow
#
# Consists of three components:
#
# 1. Explore & Process Data
# - Retrieve data
# - Clean & Explore: Explore patterns, remove any outliers
# - Transform and prepare: Data Normalization, train-validation-test split
# 2. Modeling
# - Develop & Train Model
# - Validate / Evaluate Model
# 3. Deployment
# - Deploy to Production
# - Monitor and Update Model & Data
#
# References:
#
# - AWS discusses their definition of the [ML Workflow](https://docs.aws.amazon.com/sagemaker/latest/dg/how-it-works-mlconcepts.html)
# - Google Cloud Platform (GCP) and their definition of the [ML Worklflow](https://cloud.google.com/ml-engine/docs/tensorflow/ml-solutions-overview)
# - Microsoft Azure on their definition of the [ML Workflow](https://docs.microsoft.com/en-us/azure/machine-learning/service/overview-what-is-azure-ml)
# ## Cloud Computing
#
# Can be thought of as transforming an IT product into a service.
#
# > Using an internet connected device to log into a cloud computing service to access an IT resource. These IT resources are stored in the clud provider's data center.
#
# Other cloud services than cloud storage:
#
# - Cloud applications, databases, virtual machines, SageMaker, etc.
#
# #### Why use cloud computing?
#
# Opt to use cloud computing services due time and cost constraints of building own capacities (see capacity utilization graph below). The graph shows schematically how cloud computing compares to traditinoal infrastructure related to customer demand.
#
# <img src="../images/curve3.png">
#
# Building up or laying off infrastructure takes time and money, whereas cloud computing is easily scalable to the current demand. As it is assumed to very costly to have excess demand as well as it is costly to be capacity restricted it is economically reasonable to rely on cloud computing services.
#
# - AWS: [What is Cloud Computing?](https://aws.amazon.com/de/what-is-cloud-computing/)
#
# Benefits:
#
# 1. Reduced investments and proportional costs (cost reduction)
# 2. Increased scalability (providing simplified capacity planning)
# 3. Increased availability and reliability (providing organizatinal agility)
#
# Risks:
#
# 1. (Potential) Increase in Security Vulnerabilities
# 2. Reduced Operational Governance Control (over cloud resources)
# 3. Limited Portability Between Cloud Providers
# 4. Multi-regional Compliance and Legal Issues
#
# ### Deployment to Production
#
# - Integrate machine learning model into an existing production environment
# - Model needs to be provided to those responsible for deployment.
#
# In the following we will assume taht the machine learning model was developed in Python.
#
# Three primary methods used to transfer a model from the modeling component to the deployment component (least to most commonly used):
#
# - Python model is recorded into the programming langauge of the production environment
# - Model is coded in Predictive Model Markpu Language (PMML) or Portable Format Analytics (PFA)
# - Python model is converted into a format that can be used in the production environment (i.e. SageMaker).
# - Use libraries and methods that convert the model into code that can be used in the production environment like PyTorch, TensorFlow, Scikit-Learn, etc. that convert Python models intot he intermediate standard format, such as [Open Neural Network Exchange](https://onnx.ai/) format.
# - This standard format can be converted into the software native of the production environment.
#
# The last one is the easierst and fastesst way to move a Python Model from modeling directly to deployment:
#
# - Typical way to move models into the production environoment
# - Technologies like *containers*, *endpoints* and *APIs* (Application Programming Interfaces) also help ease the work required for deploying a model into production environment.
#
# In earlier stages development was typically handled by analysts, whereas operations (deployment) was handled by software developers responsible for the production environment.
#
# Recently, this division between development and operations softens enabling analysts to handle certain aspects of deployment and enables faster updates to faltering models.
#
# Advances in cloud services, like [SageMaker](https://aws.amazon.com/sagemaker/) and [ML Engine](https://cloud.google.com/ml-engine/), and deployment technologies, like Containers and REST APIs, allow for analysts to easily take on the responsibilities of deployment.
#
# ### Production Environments
#
# - Endpoint: Interface to the model
# - The interface (enpoint) facilitates an ease of communication between the modle and the application.
#
# <img src="../images/endpoint2.png">
#
# One way to think of the **endpoint** that acts as this interface:
#
# - **endpoint** itself if like a function call
# - the **function** itself would be the model and
# - the **Python program** is the application
#
# Similar to the example above:
#
# - **Endpoint** accepts user data as the **input** and **returns** the model's prediction based upon this input through the endpoint (similar to a function call)
# - In the example, the user data is the input argument and the prediction is the returned value from the function call.
# - The **application**, here is the **python program**, displays the model's prediction to the application user.
#
# The endpoint itself is just the interface between the model and the application.
#
# - interface enables users to get predictions from the deployed model based on their user data.
#
# #### How does the endpoint (interface) facilitates communication between application and model?
#
# Application and model communicate throught he endpoint (interface). The enpoint is an Application Programming Interface (API).
#
# - API: set of rules that enable programs (here the application and the model) to communicate with each other
#
# Here, the *API* uses a **RE**presentational **S**tate **T**ransfer, **REST** architecture that provides a framework for the set of rules and constraints that mus be adhered to for communication betweeen programs.
#
# - Hypertext Transfer Protocol (HTTP): application protocol for distributed, collaborative, hypermedia information systems. Foundation of data communication for WWW.
# - **REST API** is one that uses HTTP requests and responses to enable communication between the application and the model through the endpoint (interface).
# - **HTTP request** and **HTTP response** are communications sent between the application and model.
#
# #### HTTP request
#
# HTTP request sent from applicaion to model consists of four parts:
#
# - Enpoint: Endpoint in the form of a Uniform Resource Locator (URL), aka web address
# - HTTP method: Four **HTTP methods**. For deployment of our application we'll use the **POST method**.
# - HTTP Headers: The **headers** will contain additional information (like data format within the message) that's passed to the receiving program.
# - Message (Data or Body): The final part is the **message** (data or body); for deployment this will contain the user's data which is input into the model.
#
# <img src="../images/httpmethods.png">
#
# #### HTTP response
#
# Sent from model to your application and is composed of three parts:
#
# - HTTP Status Code: If successfully received and processed the user's data that was sent in the **message** status code should start with a 2 (i.e. 200)
# - HTTP Headers: The headers will contain additional information, like format of the data within the message, thats passed to the receiving program.
# - Message (Data or Body): What's returnes as the data within the message is the prediction that's provided by the model.
#
# The prediction is then presented to the application user through the application. The enpoint is the interface that enables communication between the application and the model using a **REST API**.
#
# #### Whats's application's reponsibility?
#
# - Format the user's data to put into the HTTP request message and be used by the model
# - Translate predictions from the HTTP response message in a way that's easy for the application user's to understand.
#
# Information included in the HTTP messages sent between **application** and **model**:
#
# - User's data will need to be in a CSV or JSON format with a specific ordering of the data. Ordering depends on the used model.
# - Often predictions will be returned in CSV or JSON format with a specific ordering of the returned predictions. Ordering depends on the used model.
#
# ## Containers
#
# So far, two primary programs, the **model** and the **application**, that communicate with each other through the **endpoint (interface)**
#
# <img src="../images/endpoint3.png">
#
# - What is the **model**? The model is the Python model that's created, trained, and evaluated in the modeling component of the machine learning workflow.
# - What is the **application**? The application is a web or software that enables the users to use the model to retrieve predictions.
#
# Both, model and application, require a computing environment. One way to create this environment is to use **containers**. Containers are created using a script that contains instructions on which software packages, libraries, and other computing attributes are needed in order to run a software application, in our case either the model or application.
#
# #### But what is a container?
#
# > A container can be thought of as a standardized collection/bundle of software that is to be used for the specific purpose of running an application.
#
# A common container software is [Docker](www.docker.com).
#
# ### Containers, explained
#
# Shipping container analogy:
#
# - Shipping container can contain a wide variety of products
# - Structure of a shipping container provides the ability to hold different types of products
#
# Docker containers:
#
# - Can contain all types of different software.
# - Structure of a Docker **container** enables the **container** to be created, saved, used, and deleted through a set of common tools.
# - The common tool set works with **any container** regardless of the software the **container** contains.
#
# The image below shows three containers running three different applications
#
# <img src="../images/container.png">
#
# This architecture provides the following advantages:
#
# - Isolates the application, which increases security
# - Requires only software neede to run the application, which uses computational resources more efficiently and allows for faster application deployment.
# - Makes application creation, replication, delection, and maintenance easier and the same across all applicatinos that are deployed using containers.
# - Provides a more simple and secure way to replicate, save, and share containers.
#
# A container script file is used to create a container.
#
# - Can easily be shared with others, provides a simple method to replicate a particular container.
# - The container script is simply the instructiuons (algorithm) that is used to create a container. For *Docker*, these files are called *dockerfiles*.
#
# <img src="../images/container2.png">
#
# - Container engine uses a container script to create a container for an application to run within.
# - These container script files can be stored in repositories, which provide a simple means to share and replicate containers.
# - Docker: [Docker Hub](https://hub.docker.com/explore/) is the official repository for storing and sharing dockerfiles.
# - Example of a dockerfile: [Link](https://github.com/pytorch/pytorch/blob/master/docker/pytorch/Dockerfile)
# - The dockerfile creates a docker container with Python 3.6 and PyTorch installed.
# ## Characteristics of Deployment and Modeling
#
# #### What is Deployment?
#
# Method that integrates a machine learnin model into an existing production environment so that the model can be used to make decisions or predictions based upon data input into this model.
#
# #### Whas is a production environment?
#
# A production environment can be thought of as a web, mobile, or other software application that is currently being used by many people and must respond quickly to those users' requests.
#
# ### Characteristics of modeling
#
# #### Hyperparameters
#
# In ML, a hyperparameters is a parameter whose value cannot be estimated from the data:
#
# - Not learned through the estimators.
# - Must be set by the developer
# - Hyperparameter tuning is an important part of model training.
# - Cloud platform machine learning services often provide methods that allow for automatic hyperparameter tuning for use with model training
# - Without automatic hyperparameter option, one option is to use methods from scikit-learn Python library for hyperparameters tuning ([link](https://scikit-learn.org/stable/modules/grid_search.html#).
#
# ### Characteristics of Deployment
#
# #### Model Versioning
#
# - Saving model version as model's metadata in database
# - deployment platform should indicate a deployed model's version.
#
# #### Model Monitoring
#
# - Monitor the performance of the model
# - Application may need to be updated
#
# #### Model Updating and Routing
#
# Another characteristic:
#
# - Ability to update deployed model
# - If the monitoring process shows that performance metrics are not met the model requires updating.
# - Change in the data generating process: Collect these data to update the model
# - Routing: To allow comparison of performance between the deployed model variants, routing should be supported.
#
# #### Model Predictions
#
# Two common type of predictions provided by the deployed model.
#
# - On-demand predictions (online, real-time, synchronous predictions)
# - Predictions are returned in the response from the request. Often, these requests and responses are done through an API using JSON or XML formatted strings.
# - Commonly used to provide real-time, online responsen based upon a deployed model.
# - Batch predictions (asynchronous, batch-based predictions)
# - One expects high volume of requests with more periodic submissions, latency won't be an issue.
# - Batch request points to specifically formatted data file or request and will return the predictions to a file. Cloud services require these files will be stored in the cloud provider's cloud.
# - Batch predictions are commonly used to help make business decisions (i.e. for weekly reports).
#
# <img src="../images/mlworkflow.png">
# ## Comparing Cloud Providers
#
# Focus on [Amazon's SageMaker](https://aws.amazon.com/sagemaker/). Similar to SageMaker is [Google's ML Engine](https://cloud.google.com/ml-engine/).
#
# ### Amazon Web Services (AWS)
#
# Amazon's cloud service to build, train, and deploy ML models.
#
# Advantages:
#
# - Use of any programming language or software framework for building, training, and deploying amchine learning model in AWS
# - [Built-in algorithms](https://docs.aws.amazon.com/sagemaker/latest/dg/algos.html) - Various built-in algorithms, e.g.
# - for discrete classification or quantitative analysis using [linear learner](https://docs.aws.amazon.com/sagemaker/latest/dg/linear-learner.html) or
# - [XGBoost](https://docs.aws.amazon.com/sagemaker/latest/dg/xgboost.html)
# - item recommendations using [factorization machine](https://docs.aws.amazon.com/sagemaker/latest/dg/fact-machines.html),
# - grouping based upon attributes using [K-Means](https://docs.aws.amazon.com/sagemaker/latest/dg/k-means.html),
# - an algorithm for [image classification](https://docs.aws.amazon.com/sagemaker/latest/dg/image-classification.html)
# - Time Series Analysis with [DeepAR](https://docs.aws.amazon.com/de_de/sagemaker/latest/dg/deepar.html)
# - Custom Algorithms - Different programming languages and software frameworks that can be used to develop custom algorithms
# - [PyTorch](https://docs.aws.amazon.com/sagemaker/latest/dg/pytorch.html), [TensorFlow](https://docs.aws.amazon.com/sagemaker/latest/dg/tf.html), [Apache Spark](https://docs.aws.amazon.com/sagemaker/latest/dg/mxnet.html), and [Chainer](https://docs.aws.amazon.com/sagemaker/latest/dg/chainer.html)
# - [Own algorithms](https://docs.aws.amazon.com/sagemaker/latest/dg/your-algorithms.html) - Use your own algorithm when it isn't included within the built-in or custom algorithms above
#
# In addition, the use of [Jupyter Notebooks](https://docs.aws.amazon.com/sagemaker/latest/dg/nbi.html) is enabled and there are the following additional features and automated tools that make modeling and deployment easier:
#
# - [Automatic Model Tuning: SageMaker](https://docs.aws.amazon.com/sagemaker/latest/dg/automatic-model-tuning.html) - Feature for hyperparameter tuning of built-in and custom algorithms. In addition, SageMaker provides evaluation metrics for buil-in algorithms
# - [Monitoring Models in Sagemaker](https://docs.aws.amazon.com/sagemaker/latest/dg/monitoring-overview.html) - Features to monitor your deployed models. One can choose how much traffic to route to each deployed model (model variant).
# - More information on routing: [here](https://docs.aws.amazon.com/sagemaker/latest/dg/API_ProductionVariant.html) and [here](https://docs.aws.amazon.com/sagemaker/latest/dg/API_CreateEndpointConfig.html)
# - Type of Predictions - SageMaker allows for [On-demand](https://docs.aws.amazon.com/sagemaker/latest/dg/ex1-test-model.html) type of predictions whre each prediction request can contain one to many requestst. SageMaker also allows for [Batch](https://docs.aws.amazon.com/sagemaker/latest/dg/how-it-works-batch.html) predictions, and request data size limits are based upon S3 object size limits.
#
# ### Google Cloud Platform (GCP)
#
# [Google cloud Platform (GCP) ML Engine](https://cloud.google.com/ml-engine/) is Google's cloud service. **Similarities** and **differences:**
#
# - Prediction costs: [ML Engine pricing](https://cloud.google.com/ml-engine/docs/pricing#node-hour) vs. Sagemaker pricing.
# - Ability to explore and process data: Jupyter Notebooks are not available within ML Engine
# - To use Jupyter Notebooks within GCP, one would use [Datalab](https://cloud.google.com/datalab/docs/) can be used to explore and transform raw data into clean data for analysis and processing,
# - [DataFlow](https://cloud.google.com/dataflow/docs/) can be used to deploy batch and streaming dta processing pipelines
# - AWS also has data processing and transformation pipeline services: [AWS Glue](https://aws.amazon.com/glue/) and [AWS Data Pipeline](https://aws.amazon.com/datapipeline/)
# - Machine Learning Software: [Google's ML Engine](https://cloud.google.com/ml-engine/) has less flexibility in available software frameworks for building, training, and deploying machine learning models in GCP, compared to Amazon's SageMaker.
#
# The two available software frameworks for modeling within **ML Engine**:
#
# - [Google's TensorFlow](https://cloud.google.com/ml-engine/docs/tensorflow/) - Keras is a higher level API written in Python taht runs on top of TF.
# - [TensorFlow examples](https://cloud.google.com/ml-engine/docs/tensorflow/samples)
# - [Keras example](https://cloud.google.com/ml-engine/docs/tensorflow/samples#census-keras)
# - [Google's Scikit-learn](https://cloud.google.com/ml-engine/docs/scikit/) and [XGBoost Python package](https://xgboost.readthedocs.io/en/latest/python/index.html) can be used together for creating, training, and deploying machine learning models.
# - In [Google's example](https://cloud.google.com/ml-engine/docs/scikit/training-xgboost) XGBoost is used for modeling and Scikit-learn is used for processing the data.
#
# Flexibility in Modeling and Deployment
#
# - [Automatic Model Tuning](https://cloud.google.com/ml-engine/docs/tensorflow/hyperparameter-tuning-overview)
# - [Monitoring Models](https://cloud.google.com/ml-engine/docs/tensorflow/monitor-training)
# - Type of predictions - ML Engine allows for [Online](https://cloud.google.com/ml-engine/docs/tensorflow/online-predict) type of predictions whre each prediction request can contain one to many requests. ML Engine also allows for [Batch](https://cloud.google.com/ml-engine/docs/tensorflow/batch-predict) predictions. For more information: [Online and Batch predictions](https://cloud.google.com/ml-engine/docs/tensorflow/online-vs-batch-prediction)
#
# ### Other frameworks
#
# - Microsoft Azure
# - [Azure AI](https://azure.microsoft.com/en-us/overview/ai-platform/#platform)
# - [Azure Machine Learning Studio](https://azure.microsoft.com/en-us/services/machine-learning-studio/)
# - [Paperspace](https://www.paperspace.com/ml) - simply provides GPU-backed virtual machines with industry standard software tools
# - Claims to provide more powerful and less expensive virtual machines than AWS, GCP or Azure
# - [Cloud Foundry](https://www.cloudfoundry.org/) - open source cloud application platform
#
# ## Summary - Cloud Computing
#
# - Cloud computing - Transforming an IT product into a service
# - Deployment - Making model available for predictions through applications
# ## Cloud Computing Defined
#
# <img src="../images/nistcloud.png">
#
# The graphic above is from the Naional Institute of Standards and Technology (NIST) and its definition of cloud computing has three levels:
#
# - Service Models
# - Deployment Models
# - Essential Characteristics
#
# ### Service Models
#
# #### Software as a Service (SaaS)
#
# <img src="../images/cloud_saas.png">
#
# The yellow dashed line in the graphic shows with SaaS, the only customer responsibilities are those attributed to a "user" and all other responsibilties are placed on the cloud provider.
#
# Software as a product (i.e. a physical copy like a cd) has become rare.
#
# Other examples of SaaS:
#
# - email applications
# - storage applications
#
# #### Platform as a Service (PaaS)
#
# <img src="../images/cloud_paas.png">
#
# Examples:
#
# - Services that allow to easily build, host, monitor, and scale their applications using their platform.
# - i.e. build and host an e-commerce website.
#
# #### Infrastructure as a Service (IaaS)
#
# with IaaS the customer has most responsibility beyond those associated with running secure data centers and maintaining the hardware and software that enables IaaS.
#
# <img src="../images/cloud_iaas.png">
#
# Examples:
#
# - AWS, Rackspace
#
# IaaS enables the customer to provisioning computer processing, storage, networks, other fundamental computing resources
#
# ### Deployment Models of Cloud Computing
#
# <img src="../images/deploymentmodels.png">
#
# ### Essential Characteristics
#
# <img src="../images/essentialcharacteristics.png">
#
# ## ...
#
# (left out the second optional part)
!!jupyter nbconvert "Introdution to Deployment".ipynb
| ML_Production/Introdution to Deployment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (bayes)
# language: python
# name: bayes
# ---
# # Linear Regression (single variable) demo
#
# The dataset used is the Leinhardt Infant Mortality dataset from the R CAR package. Here we get the same data from the [Rdatasets repository](https://vincentarelbundock.github.io/Rdatasets/).
#
# The [Documentation for Leinhardt dataset](https://vincentarelbundock.github.io/Rdatasets/doc/carData/Leinhardt.html) can also be found in this repository.
#
# ## Problem Definition
#
# We want to predict the infant mortality response variable by using one or more explanatory variables available in the Leinhardt dataset.
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import statsmodels.api as sm
# %matplotlib inline
import warnings
warnings.filterwarnings("ignore")
# ## Load and explore data
try:
leinhardt_df = pd.read_csv("Leinhardt.csv")
except:
leinhardt_df = pd.read_csv("https://vincentarelbundock.github.io/Rdatasets/csv/carData/Leinhardt.csv")
leinhardt_df.to_csv("Leinhardt.csv")
leinhardt_df.head()
# ## Investigate correlation between pairs of variables
leinhardt_df["region"].unique()
leinhardt_df["oil"].unique()
# +
leinhardt_df.loc[leinhardt_df["region"] == "Asia", "region_i"] = 0
leinhardt_df.loc[leinhardt_df["region"] == "Europe", "region_i"] = 1
leinhardt_df.loc[leinhardt_df["region"] == "Americas" "region_i"] = 2
leinhardt_df.loc[leinhardt_df["region"] == "Africa", "region_i"] = 3
leinhardt_df.loc[leinhardt_df["oil"] == 'no', 'oil_i'] = 0
leinhardt_df.loc[leinhardt_df["oil"] == 'yes', 'oil_i'] = 1
_ = pd.plotting.scatter_matrix(leinhardt_df, diagonal="hist", figsize=(10,10))
# -
# ## Investigate correlation between Infant Mortality and Income
#
# From the scatter plots, there is negative correlation, but correlation doesn't appear to be linear. Thus Linear Regression on these variables is not appropriate.
#
# However, both distributions are right skewed, which is a strong hint that they might become linearly related on a log scale. On adding these log scaled columns, we see a strong linear relationship.
plt.scatter(leinhardt_df["income"], leinhardt_df["infant"])
plt.xlabel("income")
plt.ylabel("infant")
_ = plt.show()
plt.hist(leinhardt_df["infant"])
plt.xlabel("infant")
plt.ylabel("counts")
_ = plt.show()
plt.hist(leinhardt_df["income"])
plt.xlabel("income")
plt.ylabel("counts")
_ = plt.show()
leinhardt_df["log_income"] = np.log(leinhardt_df["income"])
leinhardt_df["log_infant"] = np.log(leinhardt_df["infant"])
plt.scatter(leinhardt_df["log_income"], leinhardt_df["log_infant"])
plt.xlabel("log_income")
plt.ylabel("log_infant")
_ = plt.show()
# ## Linear Regression (single variable) on log scale
#
# Fitting a OLS model against the log scaled data results in NaN results for the coefficients. This is because of missing values in the input or caused during the log transformation.
#
# The R `lm` call is more forgiving and will automatically deal with this and report that there are 4 missing values and that the results might be incorrect.
leinhardt_df_na = leinhardt_df[(leinhardt_df["log_infant"].isna()) |
(leinhardt_df["log_income"].isna())]
leinhardt_df_na.head()
leinhardt_df.dropna(subset=["log_income", "log_infant"], inplace=True)
# ## Linear Regression (single variable) on log scale after NaN removed
#
# Following information can be seen from the result of the linear regression:
#
# * intercept is 7.14 with standard error 0.317
# * coefficient of income is -0.5 with standard error 0.05
# * both values are large compared to their standard error (deviation) of the posterior, so they appear to be very statistically significant (P>|t|).
# * R-squared tells us how much of the variance is explained by the linear model.
# +
x = leinhardt_df["log_income"].values
X = np.vstack((x, np.ones(len(x)))).T
y = leinhardt_df["log_infant"].values
results = sm.OLS(y, X).fit()
results.summary()
# -
| techniques-and-models/w03-07a-linreg-demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# !ls
# +
from collections import defaultdict
graph = defaultdict(list)
with open('GraphData.txt') as f:
nodes = f.readline().replace('(', '').replace(')', '') #
n1 = nodes.split()[0].replace(',','')
n2 = nodes.split()[1]
graph[n1].append(n2)
# -
lines = []
with open('GraphData.txt') as f:
lines = f.readlines()
lines = [x.strip() for x in lines]
graph = defaultdict(list)
for line in lines:
n1 = line.split()[0].replace(',','')
n2 = line.split()[1]
graph[n1].append(n2)
graph
| ds/.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Schritt 1: Alle Excel-Dateien in Ordner öffnen
import os
import pandas as pd
files = os.listdir("../data/excel-projekt")
excel_files = [file for file in files if file[-5:] == ".xlsx"]
# +
dfs = {}
for file in excel_files:
dfs[file] = pd.read_excel("../data/excel-projekt/" + file)
# -
dfs["Marie.xlsx"]
dfs["Tobias.xlsx"]
# ### Schritt 2: Alle Dateien in ein gemeinsames Dictionary zusammenführen
d = {}
df = dfs["Tobias.xlsx"]
# +
for index, row in df.iterrows():
date_col = row["Datum"].date()
d[date_col] = {}
d[date_col]["Tobias.xlsx"] = {
"calls": row["Anzahl an Anrufen"],
"sales": row["Anzahl an Verkäufen"]
}
print(d)
# +
from datetime import date
d = {}
for filename, df in dfs.items():
for index, row in df.iterrows():
date_col = row["Datum"].date()
if not date_col in d:
d[date_col] = {}
d[date_col][filename] = {
"calls": row["Anzahl an Anrufen"],
"sales": row["Anzahl an Verkäufen"]
}
#print(d[date(year=2018, month=4, day=5)])
print(d)
# -
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import seaborn as sns
fig, ax = plt.subplots()
fig.dpi = 90
x = [excel_date for excel_date, values in sorted(d.items())]
#x = []
#for excel_date, values in sorted(d.items()):
# x.append(excel_date)
filenames = ["Tobias.xlsx", "Marie.xlsx"]
for filename in filenames:
y = []
for excel_date, values in sorted(d.items()):
calls = 0
if filename in values:
calls = values[filename]['calls']
y.append(calls)
ax.plot(x, y, label=filename)
ax.legend()
ax.set_xticks(x)
ax.set_xticklabels(x, rotation=45)
plt.show()
# -
| UDEMY_Datavis_Python/12 - projekt/Excel - Projekt (4).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data augmentation
# +
import imgaug as ia
import imgaug.augmenters as iaa
import matplotlib.gridspec as gridspec
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
plt.rcParams.update({'figure.max_open_warning': 0})
% matplotlib inline
def plot_images(images):
fig = plt.figure(figsize=(20, 20))
gs = gridspec.GridSpec(1, 10)
for label in range(10):
ax = plt.subplot(gs[0, label])
ax.imshow(images[label].reshape((28, 28)), cmap='gray')
ax.axis('off')
# -
# ## Original images
# +
data = pd.read_csv('../data/train.csv', dtype=np.uint8)
data = data.groupby('label')
images = []
for label in range(10):
img = data.get_group(label).values[0][1:]
img = np.array(img).reshape((28, 28, 1))
images.append(img)
plot_images(images)
# -
# ## Augmented images
# +
ia.seed(0)
seq = iaa.Sequential([
iaa.Sometimes(0.5,
iaa.GaussianBlur(sigma=(0, 0.5))),
iaa.ContrastNormalization((0.75, 1.5)),
iaa.Multiply((0.9, 1.1)),
iaa.Affine(
scale={'x': (0.9, 1.1), 'y': (0.9, 1.1)},
translate_percent={'x': (-0.1, 0.1), 'y': (-0.1, 0.1)},
rotate=(-10, 10),
shear=(-10, 10),
)
], random_order=True)
for i in range(10):
plot_images(seq.augment_images(images))
| notebooks/data_augmentation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Tutorial: FICO Explainable Machine Learning Challenge
# In this tutorial, we use the dataset form the FICO Explainable Machine Learning Challenge: https://community.fico.com/s/explainable-machine-learning-challenge. The goal is to create a pipeline by combining a binning process and logistic regression to obtain an explainable model and compare it against a black-box model using Gradient Boosting Tree (GBT) as an estimator.
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
# +
from optbinning import BinningProcess
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.metrics import classification_report
from sklearn.metrics import auc, roc_auc_score, roc_curve
from sklearn.model_selection import train_test_split
from sklearn.pipeline import Pipeline
# -
# Download the dataset from the link above and load it.
df = pd.read_csv("data/FICO_challenge/heloc_dataset_v1.csv", sep=",")
variable_names = list(df.columns[1:])
X = df[variable_names].values
# Transform the categorical dichotomic target variable into numerical.
y = df.RiskPerformance.values
mask = y == "Bad"
y[mask] = 1
y[~mask] = 0
y = y.astype(np.int)
# #### Modeling
# The data dictionary of this challenge includes three special values/codes:
#
# * -9 No Bureau Record or No Investigation
# * -8 No Usable/Valid Trades or Inquiries
# * -7 Condition not Met (e.g. No Inquiries, No Delinquencies)
special_codes = [-9, -8, -7]
# This challenge imposes monotonicity constraints with respect to the probability of a bad target for many of the variables. We apply these rules by passing the following dictionary of parameters for these variables involved.
binning_fit_params = {
"ExternalRiskEstimate": {"monotonic_trend": "descending"},
"MSinceOldestTradeOpen": {"monotonic_trend": "descending"},
"MSinceMostRecentTradeOpen": {"monotonic_trend": "descending"},
"AverageMInFile": {"monotonic_trend": "descending"},
"NumSatisfactoryTrades": {"monotonic_trend": "descending"},
"NumTrades60Ever2DerogPubRec": {"monotonic_trend": "ascending"},
"NumTrades90Ever2DerogPubRec": {"monotonic_trend": "ascending"},
"PercentTradesNeverDelq": {"monotonic_trend": "descending"},
"MSinceMostRecentDelq": {"monotonic_trend": "descending"},
"NumTradesOpeninLast12M": {"monotonic_trend": "ascending"},
"MSinceMostRecentInqexcl7days": {"monotonic_trend": "descending"},
"NumInqLast6M": {"monotonic_trend": "ascending"},
"NumInqLast6Mexcl7days": {"monotonic_trend": "ascending"},
"NetFractionRevolvingBurden": {"monotonic_trend": "ascending"},
"NetFractionInstallBurden": {"monotonic_trend": "ascending"},
"NumBank2NatlTradesWHighUtilization": {"monotonic_trend": "ascending"}
}
# Instantiate a ``BinningProcess`` object class with variable names, special codes and dictionary of binning parameters. Create a explainable model pipeline and a black-blox pipeline.
binning_process = BinningProcess(variable_names, special_codes=special_codes,
binning_fit_params=binning_fit_params)
# +
clf1 = Pipeline(steps=[('binning_process', binning_process),
('classifier', LogisticRegression(solver="lbfgs"))])
clf2 = LogisticRegression(solver="lbfgs")
clf3 = GradientBoostingClassifier()
# -
# Split dataset into train and test. Fit pipelines with training data, then generate classification reports to show the main classification metrics.
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=42)
clf1.fit(X_train, y_train)
clf2.fit(X_train, y_train)
clf3.fit(X_train, y_train)
y_pred = clf1.predict(X_test)
print(classification_report(y_test, y_pred))
y_pred = clf2.predict(X_test)
print(classification_report(y_test, y_pred))
y_pred = clf3.predict(X_test)
print(classification_report(y_test, y_pred))
# Plot the Receiver Operating Characteristic (ROC) metric to evaluate and compare the classifiers' prediction.
# +
probs = clf1.predict_proba(X_test)
preds = probs[:,1]
fpr1, tpr1, threshold = roc_curve(y_test, preds)
roc_auc1 = auc(fpr1, tpr1)
probs = clf2.predict_proba(X_test)
preds = probs[:,1]
fpr2, tpr2, threshold = roc_curve(y_test, preds)
roc_auc2 = auc(fpr2, tpr2)
probs = clf3.predict_proba(X_test)
preds = probs[:,1]
fpr3, tpr3, threshold = roc_curve(y_test, preds)
roc_auc3 = auc(fpr3, tpr3)
# -
plt.title('Receiver Operating Characteristic')
plt.plot(fpr1, tpr1, 'b', label='Binning+LR: AUC = {0:.2f}'.format(roc_auc1))
plt.plot(fpr2, tpr2, 'g', label='LR: AUC = {0:.2f}'.format(roc_auc2))
plt.plot(fpr3, tpr3, 'r', label='GBT: AUC = {0:.2f}'.format(roc_auc3))
plt.legend(loc='lower right')
plt.plot([0, 1], [0, 1],'k--')
plt.xlim([0, 1])
plt.ylim([0, 1])
plt.ylabel('True Positive Rate')
plt.xlabel('False Positive Rate')
plt.show()
# The plot above shows the increment in terms of model performance after binning when the logistic estimator is chosen. Furthermore, a previous binning process might reduce numerical instability issues, as confirmed when fitting the classifier ``clf2``.
# #### Binning process statistics
# The binning process of the pipeline can be retrieved to show information about the problem and timing statistics.
binning_process.information(print_level=2)
# The ``summary`` method returns basic statistics for each binned variable.
binning_process.summary()
# The ``get_binned_variable`` method serves to retrieve an optimal binning object, which can be analyzed in detail afterward.
optb = binning_process.get_binned_variable("NumBank2NatlTradesWHighUtilization")
optb.binning_table.build()
optb.binning_table.plot(metric="event_rate")
optb.binning_table.analysis()
| doc/source/tutorials/tutorial_binning_process_FICO_xAI.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PySpark
# language: python
# name: pyspark
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/pksX01/PySpark_Tutorials/blob/main/Working_with_Hive_and_PySpark_in_Google_Cloud_Dataproc.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="52ad661e-55e3-451e-b7c5-7ecb807da305" outputId="1fa4bff3-677d-45dd-f417-a14ce703019e"
sc
# + id="9b4bfeb6-6dd0-4765-abb2-c75541f02e84" outputId="814cb0cf-f3d8-4384-cb11-88194ad622db"
spark
# + id="7ded227f-b1f4-4fe3-b46d-789c6a75c13c"
df = spark.read.option('header', 'true').csv('gs://datsets-for-big-data/stroke_data/healthcare-dataset-stroke-data.csv')
# + id="e7f486cc-de63-445b-87f1-b6555420b9f5" outputId="4d09c03f-5ad5-4d97-c90f-82152989d027"
df.show(5)
# + id="9839747d-9d27-4938-bf88-e28c3aa80d4d"
new_df = df.select('id', 'gender', 'age', 'stroke')
# + id="010a681f-21cf-4b7c-95d9-25101e2e25db" outputId="d038bf8f-6097-476f-b89e-b975156c24fe"
new_df.write.option('header', 'true').csv('/user/spark/sample_stroke_data')
# + [markdown] id="e850a45f-a905-4e17-aa58-383df2ea0d94"
# **Working with Hive Tables**
# + id="00343ae8-936d-446b-b3b5-768a2f51bae1" outputId="a23dd858-e74a-4058-cddf-5763f1006a2f"
spark.sql("show databases").show()
# + id="c6f167e1-b5cc-432f-be87-7eaac6b09c05"
stocks_df = spark.sql("select * from finance.stocks")
# + id="0320cd60-4ca3-46bd-b8ab-8392b8c0464d" outputId="fa398ee1-6837-4e92-bbd5-8059e6df3205"
stocks_df.show(5)
# + id="0f973d7c-9b24-49e6-924e-7f4639607344"
import pyspark.sql.functions as f
stocks_avg_df = stocks_df.dropna().withColumn('year', f.year(f.to_date('trading_date', 'yyyy-MM-dd'))).groupBy('year')\
.agg(
f.avg('open').alias('average_open'),\
f.avg('close').alias('average_close'),\
f.avg('low').alias('average_low'),\
f.avg('high').alias('average_high'),\
)
# + id="1cae39d2-cfd6-4e8a-a330-78202d0e8cd7" outputId="e212a3fc-2b7d-4419-bd21-3096dee2de17"
stocks_avg_df.show()
# + id="f63e8272-6af9-40f3-84c2-ff777991c7b9" outputId="dca57cf4-7b22-4e98-a790-994a4086d387"
# %%writefile stocks_transformation.py
import sys
from pyspark.sql import SparkSession
import pyspark.sql.functions as f
db = sys.argv[1]
tbl = sys.argv[2]
spark = SparkSession.builder.appName("Transformations on Stocks data").enableHiveSupport().getOrCreate()
stocks_df = spark.sql("select * from {}.{}".format(db, tbl))
transformed_stocks_df = stocks_df.dropna().withColumn('year', f.year(f.to_date('trading_date', 'yyyy-MM-dd'))).groupBy('year')\
.agg(
f.avg('open').alias('average_open'),\
f.avg('close').alias('average_close'),\
f.avg('low').alias('average_low'),\
f.avg('high').alias('average_high'),\
)
transformed_stocks_df.write.mode("overwrite").saveAsTable("finance.transformed_stocks")
# + id="5dfe4989-4831-42b8-856c-0aeef89c0e95"
from google.cloud import storage
def upload_blob(bucket_name, source_file_name, destination_blob_name):
"""Uploads a file to the bucket."""
# The ID of your GCS bucket
# bucket_name = "your-bucket-name"
# The path to your file to upload
# source_file_name = "local/path/to/file"
# The ID of your GCS object
# destination_blob_name = "storage-object-name"
storage_client = storage.Client()
bucket = storage_client.bucket(bucket_name)
blob = bucket.blob(destination_blob_name)
blob.upload_from_filename(source_file_name)
print(
"File {} uploaded to {}.".format(
source_file_name, destination_blob_name
)
)
# + id="72b4336a-bd72-4508-8f62-4fb33de1884c" outputId="e7d32564-5eb3-401b-ba2a-87bb5d13708d"
upload_blob('datsets-for-big-data', 'stocks_transformation.py', 'python_files/stocks_transformation.py')
# + id="8d500b4d-4fdc-4a5e-a62e-172b93fd7f94"
| Working_with_Hive_and_PySpark_in_Google_Cloud_Dataproc.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/nahin333/DL-practice-codes/blob/main/word2vec_with_gensim.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="Ici4PUQYdl6N"
import gensim
import pandas as pd
# + [markdown] id="VA5ZVLBCvT51"
# ##Dataset link: http://snap.stanford.edu/data/amazon/productGraph/categoryFiles/reviews_Sports_and_Outdoors_5.json.gz
# + id="93tLEVqFdwG0"
import gzip
import shutil
with gzip.open('/content/reviews_Sports_and_Outdoors_5.json.gz', 'rb') as f_in:
with open('Sports_and_Outdoors_5.json', 'wb') as f_out:
shutil.copyfileobj(f_in, f_out)
# + id="ArXyQ9PPzjkB" colab={"base_uri": "https://localhost:8080/", "height": 258} outputId="1b6ef40c-8dd0-425e-c01b-e5243f4d6b1a"
df = pd.read_json('/content/Sports_and_Outdoors_5.json', lines=True)
df.head()
# + id="3_Zpba5Dz8os" colab={"base_uri": "https://localhost:8080/"} outputId="35d37644-3fc1-42e1-af7a-f9087f66af14"
df.shape
# + id="-T7pHTIY1YOz" colab={"base_uri": "https://localhost:8080/", "height": 36} outputId="35c48376-4f51-44f8-fac7-12063a96fe1f"
df.reviewText[0]
# + id="h6NeQ9cd6u9B" colab={"base_uri": "https://localhost:8080/"} outputId="952f03b7-1d59-4a5a-ae50-6ec1debf3890"
gensim.utils.simple_preprocess(df.reviewText[0])
# + id="Xog5kQj27GNb" colab={"base_uri": "https://localhost:8080/"} outputId="c4b297ea-0efa-43c1-9ca4-3485540dce81"
review_text = df.reviewText.apply(gensim.utils.simple_preprocess)
review_text
# + id="TInp8dOb7cLG"
model = gensim.models.Word2Vec(
window = 10,
min_count = 2,
workers = 4
)
# + id="7U96cUEeIR7l"
model.build_vocab(review_text, progress_per=1000)
# + colab={"base_uri": "https://localhost:8080/"} id="OpcCaD5KJILk" outputId="6b75d1b3-fbd5-4a0b-fff8-f286f168c31b"
model.epochs
# + colab={"base_uri": "https://localhost:8080/"} id="cDEbEoM4JpzO" outputId="25e9ad48-41eb-47ef-b22a-9aed489f450d"
model.corpus_count
# + colab={"base_uri": "https://localhost:8080/"} id="8hzRRrvXJR7I" outputId="1c0acd2b-4b20-4243-be07-0804dc4026a0"
model.train(review_text, total_examples = model.corpus_count, epochs = model.epochs)
# + id="AhJZsjC4Jloy"
model.save('/content/w2vec_sports_and_outdoors_reviews.model')
# + colab={"base_uri": "https://localhost:8080/"} id="zkvigA2qKXf2" outputId="174744af-a54a-4bf4-ef8b-600a4fa97c1d"
model.wv.most_similar('awful')
# + colab={"base_uri": "https://localhost:8080/"} id="V2Ck5UmdKk-H" outputId="d8498b7b-3537-4bf4-d8d4-9e97f8787ac4"
model.wv.similarity(w1= 'slow', w2='steady')
# + colab={"base_uri": "https://localhost:8080/"} id="E0DT3yUpLC6V" outputId="70bc48fc-6341-4884-cb0d-306303cbd014"
model.wv.similarity(w1='great', w2='good')
| word2vec_with_gensim.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Day 3 | Logistic regression
# logistic regression是一個分類模型,是在機器學習分類問題時相當基礎的一個模型,其實整個模型的架構簡單來說就是將一個linear regression的模型放入一個sigmoid函數中,將結果輸出為0~1之間的數值,而我們再用一個threshhlod來去決定如何分類,相關資源如下:
# 1. [資料分析&機器學習] 第3.3講:線性分類-邏輯斯回歸(Logistic Regression) 介紹 : [link](https://medium.com/@yehjames/%E8%B3%87%E6%96%99%E5%88%86%E6%9E%90-%E6%A9%9F%E5%99%A8%E5%AD%B8%E7%BF%92-%E7%AC%AC3-3%E8%AC%9B-%E7%B7%9A%E6%80%A7%E5%88%86%E9%A1%9E-%E9%82%8F%E8%BC%AF%E6%96%AF%E5%9B%9E%E6%AD%B8-logistic-regression-%E4%BB%8B%E7%B4%B9-a1a5f47017e5)
# 2. 李宏毅老師課程:[link](https://www.youtube.com/watch?v=hSXFuypLukA&list=PLJV_el3uVTsPy9oCRY30oBPNLCo89yu49&index=10)
# 3. 目前最火的100 days ML code 挑戰 Avik-Jain:[link](https://github.com/Avik-Jain/100-Days-Of-ML-Code/blob/master/Code/Day%206%20Logistic%20Regression.md)
# 而本次將使用kaggle鐵達尼號作為logistic regression資料:
# https://www.kaggle.com/francksylla/titanic-machine-learning-from-disaster/data
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
train_df = pd.read_csv('all/train.csv')
test_df = pd.read_csv('all/test.csv')
# train_df.info()
train_df.head()
# ## 確認缺失值(利用heatMap)
sns.heatmap(train_df.isnull(), yticklabels= False, cmap = 'viridis', cbar=False)
## 可以透過heatmap看出Cabin預age的資訊缺失的相當多,
# ## EDA
# 確認存活人數的比例,利用seaborn countplot來確認。
# +
sns.set_style('whitegrid')
count_plot = sns.countplot(data = train_df, x = 'Survived')
for i, value in enumerate(train_df.Survived.value_counts()):
count_plot.text(x = i-0.1, y =value+10, s = '{}%'.format(round(value/len(train_df)*100,2)), fontsize = 15)
### 由此圖可以看出大部分的人(6成)的人都未能存活
# -
sns.set_style('whitegrid')
count_plot = sns.countplot(data = train_df, hue = 'Sex', x = 'Survived')
# +
sns.set_style('whitegrid')
# count_plot = sns.countplot(data = train_df, hue = 'Embarked', x = 'Survived')
g = sns.FacetGrid(data = train_df, col='Embarked', row = 'Sex')
g = g.map(sns.countplot, 'Survived')
### 再針對各不同地區上船資訊加上性別來分析,大多罹難的的都是在Southampton上船的男性
# -
sns.distplot(train_df.Age.dropna(), kde= False, bins = 30)
### 由此區間可以看出約呈現一個常態分佈,大部分的乘客都是介於20-30碎的青壯年
# +
sns.set_style('whitegrid')
g = sns.FacetGrid(data = train_df, col='Embarked', row = 'Sex')
g = g.map(plt.hist, 'Fare')
## 另外又從此表看出由Southampton上船的男性佔沒有買票的多數
# -
# # Clean data
# 接下來我們來清理資料,由上面的heatmap可以看出Cabin的資料缺失值太多,因此我將它轉成有Cabin與沒有Cabin,另外年齡做後續處理。
plt.figure(figsize = (8,5))
sns.boxplot(data = train_df, x = "Pclass", y = "Age")
## 我們可以假設年齡與艙等也是有關係的,因此我們利用各艙等年齡的平均來補植
print('Pclass_1 average age: ', int(train_df[train_df.Pclass == 1].Age.mean()))
print('Pclass_2 average age: ', int(train_df[train_df.Pclass == 2].Age.mean()))
print('Pclass_3 average age: ', int(train_df[train_df.Pclass == 3].Age.mean()))
# +
## 確認如果有Cabin資料則給1,沒有則給0
def cabin_check(cabin):
if pd.isnull(cabin):
return 0
else:
return 1
## 確認艙等後回傳該艙等年齡平均值
def Age_chcek(pclass_age, df = train_df):
pclass = pclass_age[0]
age = pclass_age[1]
if pd.isnull(age):
if pclass == 1:
return 38
if pclass == 2:
return 29
else:
return 25
else:
return age
# -
train_df.Cabin = train_df.Cabin.apply(cabin_check)
train_df.Age = train_df[['Pclass', 'Age']].apply(Age_chcek, axis = 1)
sns.heatmap(train_df.isnull(), yticklabels= False, cmap = 'viridis', cbar=False)
### 資料已經清理乾淨了,而剩下一筆是Embark的缺失值,因此直接drop
train_df.dropna(inplace = True)
# ### 轉換類別型資料
train_df.head()
### 像是姓名,船票等文字資料沒有辦法使用,因此先捨棄,而像性別,上船地點等可以轉換成dummy variable
train_df.drop(['Name', 'Ticket'], axis = 1, inplace = True)
train_df = pd.get_dummies(train_df, drop_first= True)
train_df.head()
# # Logistic Regresstion
# 首先import必要的套件:
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split
X = train_df.drop(['Survived'], axis = 1)
y = train_df.Survived
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=42)
print("Training set number: ", len(X_train))
print("Testing set number: ", len(X_test))
logistic_model = LogisticRegression()
logistic_model.fit(X_train, y_train)
predict = logistic_model.predict(X_test)
# ## Evaluation
# 可以藉由scikit-learn的confusion-matrix 以及 classodication_report函式來評估準確度
# 1. Confusion matrix : [如何辨別機器學習模型的好壞?秒懂Confusion Matrix](https://www.ycc.idv.tw/confusion-matrix.html)
from sklearn.metrics import confusion_matrix, classification_report
print(confusion_matrix(y_test, predict))
print(classification_report(y_test, predict))
# ##### 可以看出此資料集透過簡單的整理後,直接使用logistic regression來做分類就可以得出一個不差的結果
# 而其實有關線性回歸模型,不管是linear regression or logistic regression 可能都會碰到的問題就是overfitting 或是 underfitting 的問題,應該如何調整與權衡,預計將放在Day4來討論。
# 相關資料:
# 1. https://medium.com/@ken90242/machine-learning%E5%AD%B8%E7%BF%92%E6%97%A5%E8%A8%98-coursera%E7%AF%87-week-3-4-the-c05b8ba3b36f
| Day_3_logistic_regression/Day_2_logistic_regressio.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# # Figure 6 - application to HD-MEA datasets
#
# This notebook shows reconstruct axons from two HD-MEA datasets.
# The datasets (`mea1k.npz` and `dualmode.npz`) needs to be downloaded from Zenodo (https://doi.org/10.5281/zenodo.4896745) and placed in the `axon_velocity/data/mea1k` and `axon_velocity/data/dualmode` folders, respectively.
# +
import numpy as np
import matplotlib.pylab as plt
import MEAutility as mu
import numpy as np
from pathlib import Path
from pprint import pprint
from probeinterface import plotting
from tqdm import tqdm
import pandas as pd
import axon_velocity as av
# %matplotlib widget
# +
save_figs = True
fig_folder = Path("figures/") / "figure6"
fig_folder.mkdir(exist_ok=True)
# +
params = av.get_default_graph_velocity_params()
# change params
params['detect_threshold'] = 0.01
params['kurt_threshold'] = 0.1
params['peak_std_threshold'] = 0.8
params['upsample'] = 5
params['neighbor_radius'] = 100
params['r2_threshold'] = 0.8
pprint(params)
# -
def plot_unit_summary(gtr, probe):
# amplitude map
fig_amp, ax_amp = plt.subplots()
_ = av.plot_amplitude_map(gtr.template, gtr.locations, log=True, ax=ax_amp, colorbar=False,
colorbar_orientation="horizontal")
# latency map
fig_peak, ax_peak = plt.subplots()
_ = av.plot_peak_latency_map(gtr.template, gtr.locations, gtr.fs, ax=ax_peak, colorbar=False,
colorbar_orientation="horizontal")
# branches
fig_branches, ax_branches = plt.subplots()
_ = plotting.plot_probe(probe, ax=ax_branches, contacts_kargs={"alpha": 0.1}, probe_shape_kwargs={"alpha": 0.1})
ax_branches.axis("off")
ax_branches.set_title("")
ax_branches.plot(gtr.locations[gtr.selected_channels, 0], gtr.locations[gtr.selected_channels, 1], marker=".",
color="k", alpha=0.1, markersize=3, ls="")
cm = plt.get_cmap("tab20")
for i, br in enumerate(gtr.branches):
ax_branches.plot(gtr.locations[br["channels"], 0], gtr.locations[br["channels"], 1],
marker=".", color=cm(i / len(gtr.branches)), ls="-", alpha=0.8, label=i)
# velocities
fig_vel, ax_vel = plt.subplots()
av.plot_branch_velocities(gtr.branches, legend=False, ax=ax_vel, cmap="tab20")
ax_vel.spines['right'].set_visible(False)
ax_vel.spines['top'].set_visible(False)
ax_vel.set_xticklabels([])
ax_vel.set_yticklabels([])
fig_dict = dict(amplitude=fig_amp, latency=fig_peak, branches=fig_branches, velocity=fig_vel)
return fig_dict
data_folder = Path("../data/")
mea1k_folder = data_folder / "mea1k"
dualmode_folder = data_folder / "dualmode"
# ## Load MEA1k data
load_dict = np.load(mea1k_folder / "mea1k.npz")
templates_mea1k = load_dict["templates"]
locations_mea1k = load_dict["locations"]
fs_mea1k = load_dict["fs"]
templates_mea1k.shape
# ### Load or recompute axonal branches
if Path(mea1k_folder / "gtrs.npy").is_file():
print("Loading existing axonal branches")
gtrs_mea1k = np.load(mea1k_folder / "gtrs.npy", allow_pickle=True)
gtrs_mea1k = gtrs_mea1k.item()
else:
print("Computing and saving axonal branches")
gtrs_mea1k = dict()
for i in tqdm(range(len(templates_mea1k)), desc="Extracting axons"):
template = templates_mea1k[i]
try:
gtr = av.compute_graph_propagation_velocity(template, locations_mea1k, fs_mea1k,
verbose=False, **params)
gtrs_mea1k[i] = gtr
print(f"Found axon for unit {i}")
except Exception as e:
print(f"Failed on {i}: error {e}")
np.save(mea1k_folder / "gtrs.npy", gtrs_mea1k)
print(f"MEA1k: Found {len(gtrs_mea1k)} units with detectable axons out of {len(templates_mea1k)}")
# +
unit_ids = []
branch_ids = []
velocities = []
path_lengths = []
r2s = []
for unit, gtr in gtrs_mea1k.items():
for i, br in enumerate(gtr.branches):
path = br["channels"]
velocity = br["velocity"]
r2 = br["r2"]
length = gtr.compute_path_length(path)
unit_ids.append(unit)
branch_ids.append(i)
velocities.append(velocity)
path_lengths.append(length)
r2s.append(r2)
df_mea1k = pd.DataFrame({"unit_ids": unit_ids, "branch_id": branch_ids, "velocity": velocities,
"length": path_lengths, "r2": r2s})
# -
print("MEA1k\n\n")
print(f"Num axonal branches: {len(df_mea1k)}")
print(f"Velocities: {np.round(df_mea1k.velocity.mean(), 2)} +- {np.round(df_mea1k.velocity.std(), 2)}")
print(f"Path lengths: {np.round(df_mea1k.length.mean(), 2)} +- {np.round(df_mea1k.length.std(), 2)}")
print(f"R2: {np.round(df_mea1k.r2.mean(), 2)} +- {np.round(df_mea1k.r2.std(), 2)}")
mea1k_selected_unit_idxs = [8]
# +
probe_mea1k = av.plotting.get_probe(locations_mea1k)
fig_mea1k, ax = plt.subplots(figsize=(10, 7))
_ = plotting.plot_probe(probe_mea1k, ax=ax, contacts_kargs={"alpha": 0.1}, probe_shape_kwargs={"alpha": 0.1})
ax.axis("off")
i = 0
i_sel = 0
cmap = "tab20"
cm = plt.get_cmap(cmap)
for i, gtr in gtrs_mea1k.items():
if i in mea1k_selected_unit_idxs:
color = f"C{i_sel}"
lw = 3
alpha = 1
zorder = 10
i_sel += 1
else:
color = cm(i / len(gtrs_mea1k))
lw = 1
alpha = 1
zorder = 1
if len(gtr.branches) > 0:
ax.plot(gtr.locations[gtr.init_channel, 0], gtr.locations[gtr.init_channel, 1],
marker="o", markersize=5, color=color, alpha=alpha, zorder=zorder)
if i not in mea1k_selected_unit_idxs:
# for visualization purposes, plot raw branches
for b_i, path in enumerate(gtr._paths_raw):
if b_i == 0:
ax.plot(gtr.locations[path, 0], gtr.locations[path, 1], marker="", color=color,
lw=lw, alpha=alpha, zorder=zorder, label=i)
else:
ax.plot(gtr.locations[path, 0], gtr.locations[path, 1], marker="", color=color,
lw=lw, alpha=alpha, zorder=zorder)
else:
for b_i, br in enumerate(gtr.branches):
if b_i == 0:
ax.plot(gtr.locations[br["channels"], 0], gtr.locations[br["channels"], 1], marker="",
color=color, lw=lw, alpha=alpha, zorder=zorder, label=i)
else:
ax.plot(gtr.locations[br["channels"], 0], gtr.locations[br["channels"], 1], marker="",
color=color, lw=lw, alpha=alpha, zorder=zorder)
ax.plot([0, 500], [1900, 1900], color="k", marker="|")
ax.text(20, 1950, "500$\mu$m", color="k", fontsize=18)
ax.set_title("")
# -
for i in mea1k_selected_unit_idxs:
gtr = gtrs_mea1k[i]
amplitude = np.ptp(gtr.template[gtr.init_channel])
num_selected = len(gtr.selected_channels)
num_branches = len(gtr.branches)
vels = []
r2s = []
lengths = []
for br in gtr.branches:
vels.append(br["velocity"])
lengths.append(gtr.compute_path_length(br["channels"]))
r2s.append(br["r2"])
print(f"Unit {i}\n\n")
print(f"Amplitude: {np.round(amplitude, 1)} uV")
print(f"Num selected channels: {num_selected}")
print(f"Num axonal branches: {num_branches}")
print(f"Velocities: {np.round(np.mean(vels), 2)} +- {np.round(np.std(vels), 2)}")
print(f"Path lengths: {np.round(np.mean(lengths), 2)} +- {np.round(np.std(lengths), 2)}")
print(f"R2: {np.round(np.mean(r2s), 2)} +- {np.round(np.std(r2s), 2)}")
figs_mea1k = []
for i in mea1k_selected_unit_idxs:
fig_dict = plot_unit_summary(gtrs_mea1k[i], probe_mea1k)
figs_mea1k.append(fig_dict)
if save_figs:
for i, fig_dict in enumerate(figs_mea1k):
for fig_name, fig in fig_dict.items():
if fig_name == "velocity":
fig.savefig(fig_folder / f"mea1k_neuron{i+1}_{fig_name}.svg")
else:
fig.savefig(fig_folder / f"mea1k_neuron{i+1}_{fig_name}.png", dpi=600)
fig_mea1k.savefig(fig_folder / f"mea1k.png", dpi=600)
# ## Load DualMode data
load_dict = np.load(dualmode_folder / "dualmode.npz")
templates_dualmode = load_dict["templates"]
locations_dualmode = load_dict["locations"]
fs_dualmode = load_dict["fs"]
params['upsample'] = 10 # to get ~ 100kHz
if Path(dualmode_folder / "gtrs.npy").is_file():
print("Loading existing axonal branches")
gtrs_dualmode = np.load(dualmode_folder / "gtrs.npy", allow_pickle=True)
gtrs_dualmode = gtrs_dualmode.item()
else:
print("Computing and saving axonal branches")
gtrs_dualmode = dict()
for i in tqdm(range(len(templates_dualmode)), desc="Extracting axons"):
template = templates_dualmode[i]
try:
gtr = av.compute_graph_propagation_velocity(template, locations_dualmode, fs_dualmode,
verbose=False, **params)
gtrs_dualmode[i] = gtr
print(f"Found axon for unit {i}")
except Exception as e:
print(f"Failed on {i}: error {e}")
np.save(dualmode_folder / "gtrs.npy", gtrs_dualmode)
print(f"DualMode: Found {len(gtrs_dualmode)} units with detectable axons out of {len(templates_dualmode)}")
# +
unit_ids = []
branch_ids = []
velocities = []
path_lengths = []
r2s = []
for unit, gtr in gtrs_dualmode.items():
for i, br in enumerate(gtr.branches):
path = br["channels"]
velocity = br["velocity"]
r2 = br["r2"]
length = gtr.compute_path_length(path)
unit_ids.append(unit)
branch_ids.append(i)
velocities.append(velocity)
path_lengths.append(length)
r2s.append(r2)
df_dualmode = pd.DataFrame({"unit_ids": unit_ids, "branch_id": branch_ids, "velocity": velocities,
"length": path_lengths, "r2": r2s})
# -
print("DualMode\n\n")
print(f"Num axonal branches: {len(df_dualmode)}")
print(f"Velocities: {np.round(df_dualmode.velocity.mean(), 2)} +- {np.round(df_dualmode.velocity.std(), 2)}")
print(f"Path lengths: {np.round(df_dualmode.length.mean(), 2)} +- {np.round(df_dualmode.length.std(), 2)}")
print(f"R2: {np.round(df_dualmode.r2.mean(), 2)} +- {np.round(df_dualmode.r2.std(), 2)}")
dualmode_selected_unit_idxs = [20]
# +
probe_dualmode = av.plotting.get_probe(locations_dualmode)
fig_dualmode, ax = plt.subplots(figsize=(10, 7))
_ = plotting.plot_probe(probe_dualmode, ax=ax, contacts_kargs={"alpha": 0.1}, probe_shape_kwargs={"alpha": 0.1})
ax.axis("off")
i = 0
i_sel = 0
cmap = "tab20"
cm = plt.get_cmap(cmap)
for i, gtr in gtrs_dualmode.items():
if i in dualmode_selected_unit_idxs:
color = f"C{i_sel}"
lw = 3
alpha = 1
zorder = 10
i_sel += 1
else:
color = cm(i / len(gtrs_dualmode))
lw = 1
alpha = 1
zorder = 1
if len(gtr.branches) > 0:
ax.plot(gtr.locations[gtr.init_channel, 0], gtr.locations[gtr.init_channel, 1],
marker="o", markersize=5, color=color, alpha=alpha, zorder=zorder)
if i not in dualmode_selected_unit_idxs:
# for visualization purposes, plot raw branches
for b_i, path in enumerate(gtr._paths_raw):
if b_i == 0:
ax.plot(gtr.locations[path, 0], gtr.locations[path, 1], marker="", color=color,
lw=lw, alpha=alpha, zorder=zorder, label=i)
else:
ax.plot(gtr.locations[path, 0], gtr.locations[path, 1], marker="", color=color,
lw=lw, alpha=alpha, zorder=zorder)
else:
for b_i, br in enumerate(gtr.branches):
if b_i == 0:
ax.plot(gtr.locations[br["channels"], 0], gtr.locations[br["channels"], 1], marker="", color=color,
lw=lw, alpha=alpha, zorder=zorder, label=i)
else:
ax.plot(gtr.locations[br["channels"], 0], gtr.locations[br["channels"], 1], marker="", color=color,
lw=lw, alpha=alpha, zorder=zorder)
ax.plot([0, 500], [1650, 1650], color="k", marker="|")
ax.text(20, 1690, "500$\mu$m", color="k", fontsize=18)
ax.set_title("")
# -
for i in dualmode_selected_unit_idxs:
gtr = gtrs_dualmode[i]
amplitude = np.ptp(gtr.template[gtr.init_channel])
num_selected = len(gtr.selected_channels)
num_branches = len(gtr.branches)
vels = []
r2s = []
lengths = []
for br in gtr.branches:
vels.append(br["velocity"])
lengths.append(gtr.compute_path_length(br["channels"]))
r2s.append(br["r2"])
print(f"Unit {i}\n\n")
print(f"Amplitude: {np.round(amplitude, 1)} uV")
print(f"Num selected channels: {num_selected}")
print(f"Num axonal branches: {num_branches}")
print(f"Velocities: {np.round(np.mean(vels), 2)} +- {np.round(np.std(vels), 2)}")
print(f"Path lengths: {np.round(np.mean(lengths), 2)} +- {np.round(np.std(lengths), 2)}")
print(f"R2: {np.round(np.mean(r2s), 2)} +- {np.round(np.std(r2s), 2)}")
figs_dualmode = []
for i in dualmode_selected_unit_idxs:
fig_dict = plot_unit_summary(gtrs_dualmode[i], probe_dualmode)
figs_dualmode.append(fig_dict)
if save_figs:
for i, fig_dict in enumerate(figs_dualmode):
for fig_name, fig in fig_dict.items():
if fig_name == "velocity":
fig.savefig(fig_folder / f"dualmode_neuron{i+1}_{fig_name}.svg")
else:
fig.savefig(fig_folder / f"dualmode_neuron{i+1}_{fig_name}.png", dpi=600)
fig_dualmode.savefig(fig_folder / f"dualmode.png", dpi=600)
| figure_notebooks/figure6_HD-MEA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/z-arabi/pytorchTutorial/blob/master/12_plot_activations.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="LVNLPmW5AqeO"
import numpy as np
import matplotlib.pyplot as plt
# + colab={"base_uri": "https://localhost:8080/", "height": 330} id="ntAfEs9sAvA7" outputId="88eeca06-6652-4a68-8bd5-06216538e36c"
##### Sigmoid
sigmoid = lambda x: 1 / (1 + np.exp(-x))
x=np.linspace(-10,10,10)
y=np.linspace(-10,10,100)
fig = plt.figure()
plt.plot(y,sigmoid(y),'b', label='linspace(-10,10,100)')
plt.grid(linestyle='--')
plt.xlabel('X Axis')
plt.ylabel('Y Axis')
plt.title('Sigmoid Function')
plt.xticks([-4, -3, -2, -1, 0, 1, 2, 3, 4])
plt.yticks([-2, -1, 0, 1, 2])
plt.ylim(-2, 2)
plt.xlim(-4, 4)
plt.show()
plt.savefig('sigmoid.png')
fig = plt.figure()
# + colab={"base_uri": "https://localhost:8080/", "height": 312} id="rWWoC_V0BzLr" outputId="821a22eb-f8d2-4c9a-a4c2-c351e1e58b8d"
##### TanH
tanh = lambda x: 2*sigmoid(2*x)-1
x=np.linspace(-10,10,10)
y=np.linspace(-10,10,100)
plt.plot(y,tanh(y),'b', label='linspace(-10,10,100)')
plt.grid(linestyle='--')
plt.xlabel('X Axis')
plt.ylabel('Y Axis')
plt.title('TanH Function')
plt.xticks([-4, -3, -2, -1, 0, 1, 2, 3, 4])
plt.yticks([-4, -3, -2, -1, 0, 1, 2, 3, 4])
plt.ylim(-4, 4)
plt.xlim(-4, 4)
plt.show()
#plt.savefig('tanh.png')
fig = plt.figure()
# + colab={"base_uri": "https://localhost:8080/", "height": 312} id="p6lXrJkrB21b" outputId="d8047628-39ce-47cd-85c0-cc01af0b8e54"
##### ReLU
relu = lambda x: np.where(x>=0, x, 0)
x=np.linspace(-10,10,10)
y=np.linspace(-10,10,1000)
plt.plot(y,relu(y),'b', label='linspace(-10,10,100)')
plt.grid(linestyle='--')
plt.xlabel('X Axis')
plt.ylabel('Y Axis')
plt.title('ReLU')
plt.xticks([-4, -3, -2, -1, 0, 1, 2, 3, 4])
plt.yticks([-4, -3, -2, -1, 0, 1, 2, 3, 4])
plt.ylim(-4, 4)
plt.xlim(-4, 4)
plt.show()
#plt.savefig('relu.png')
fig = plt.figure()
# + colab={"base_uri": "https://localhost:8080/", "height": 312} id="6givLM_ZCD5i" outputId="d1373ae3-52a9-44b7-bb1a-2a9c75de2bca"
##### Leaky ReLU
leakyrelu = lambda x: np.where(x>=0, x, 0.1*x)
x=np.linspace(-10,10,10)
y=np.linspace(-10,10,1000)
plt.plot(y,leakyrelu(y),'b', label='linspace(-10,10,100)')
plt.grid(linestyle='--')
plt.xlabel('X Axis')
plt.ylabel('Y Axis')
plt.title('Leaky ReLU')
plt.xticks([-4, -3, -2, -1, 0, 1, 2, 3, 4])
plt.yticks([-4, -3, -2, -1, 0, 1, 2, 3, 4])
plt.ylim(-4, 4)
plt.xlim(-4, 4)
plt.show()
#plt.savefig('lrelu.png')
fig = plt.figure()
# + colab={"base_uri": "https://localhost:8080/", "height": 312} id="mzixDtKiAo1z" outputId="b1260d21-dfc0-4cf4-ef9c-e4efd7ab268e"
##### Binary Step
bstep = lambda x: np.where(x>=0, 1, 0)
x=np.linspace(-10,10,10)
y=np.linspace(-10,10,1000)
plt.plot(y,bstep(y),'b', label='linspace(-10,10,100)')
plt.grid(linestyle='--')
plt.xlabel('X Axis')
plt.ylabel('Y Axis')
plt.title('Step Function')
plt.xticks([-4, -3, -2, -1, 0, 1, 2, 3, 4])
plt.yticks([-2, -1, 0, 1, 2])
plt.ylim(-2, 2)
plt.xlim(-4, 4)
plt.show()
#plt.savefig('step.png')
print('done')
| 12_plot_activations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ## Implementing Edge Detection Operators in Python
# ## Table of Contents
# * [Edge Detection Operators](#Edge)
# * [Kirsch Operator](#Kirsch)
# * [Sobel Operator](#Sobel)
# * [Python Implementation](#Implementation)
# * [Applying the Operators](#Applying)
# <a id="Edge"></a>
# ## Edge Detection Operators
# Image edge detection is a simple operation in image processing but is a fundamental step in some other complex operations such as image recognition and scene analysis. There are different operators that are used in detecting edges of images.
#
# In this post, we are going to implement and apply two operators that are widely used for edge detection.
# * Kirsch Operator
# * Sobel Operator
#
# These 3x3 operators are usually applied to images in spatial domain by convolving in them.
# <a id="Kirsch"></a>
# ## Kirsch Operator
# Consider a 3x3 image region centered at $A_*$. Let the region be as follows:
#
# $$ \begin{bmatrix}
# A_0 & A_1 & A_2\\
# A_8 & A_* & A_4\\
# A_7 & A_6 & A_5
# \end{bmatrix} $$
#
# $S_i = A_i + A_{i+1} + A_{i+2} $ (Modulo $8$)
#
# $T_i = A_{i+3} + A_{i+4} + A_{i+5} + A_{i+6} + A_{i+7} + A_{i+8}$ (Modulo $8$)
#
# So the gradient will be calculated as:
#
# $ G(j,k) = Max_{i=0}^7[|5S_i - 3T_i|]$
#
# In the kirsch operator, the one that gives the maximum value in all directions is selected as the gradient.
# <a id="Sobel"></a>
# ## Sobel Operator
# The Sobel operator is applied as follows:
#
# $ \Delta_1 = \begin{bmatrix}
# -1 & 0 & 1\\
# -2 & 0 & 2\\
# -1 & 0 & 1
# \end{bmatrix} $
#
# $ \Delta_2 = \begin{bmatrix}
# 1 & 2 & 1\\
# 0 & 0 & 0\\
# -1 & -2 & -1
# \end{bmatrix} $
#
# So the edge enhanced one will be calculated as:
#
# $ g(x,y) = \sqrt{\Delta_1^2 + \Delta_2^2} $
# <a id="Implementation"></a>
# ## Python Implementation
# Let's first import the common classes.
# +
from CommonClasses.fft import *
from CommonClasses.dct import *
from CommonClasses.walsh import *
from CommonClasses.haar import *
from CommonClasses.utils import *
import numpy as np
import matplotlib.pyplot as plt
# #%matplotlib inline
#import matplotlib.image as img
#import PIL.Image as Image
from PIL import Image
import math
import cmath
import time
import csv
from numpy import binary_repr
from fractions import gcd
# -
def computeKirsch(imge):
"""Computes and applies Kirsch operator to a given image."""
N = imge.shape[0]
result = np.zeros([N, N], dtype=float)
#Copy the first and last rows, first and last columns
result[0, :] = imge[0, :]
result[:, 0] = imge[:, 0]
result[N-1, :] = imge[N-1, :]
result[:, N-1] = imge[:, N-1]
#Kirsch Operator to the image.
for i in np.arange(1, N-1):
for j in np.arange(1, N-1):
#Take the sub image.
subImge = imge[i-1:i+2, j-1:j+2]
#Number of elements in which Kirsch operator is applied
n = 8
#Flatten the sub image.
subImgeFl = np.zeros(n, dtype=int)
subImgeFl[:3] = subImge[0,:]
subImgeFl[3] = subImge[1, -1]
subImgeFl[4:7] = subImge[-1,:][::-1]
subImgeFl[7] = subImge[1,0]
#Variable that stores the maximum value
mx = 1
for k in np.arange(n):
S = subImgeFl[k%n]+subImgeFl[(k+1)%n]+subImgeFl[(k+2)%n]
T = (subImgeFl[(k+3)%n]+subImgeFl[(k+4)%n]+subImgeFl[(k+5)%n]+
subImgeFl[(k+6)%n]+subImgeFl[(k+7)%n])
diff = abs((5*S)-(3*T))
if diff > mx:
mx = diff
result[i, j] = mx
return result
def generateRowColumnSobelGradients():
"""Generates the x-component and y-component of Sobel operators."""
rowGradient = np.array([[1, 0, -1], [2, 0, -2], [1, 0, -1]])
colGradient = np.array([[-1, -2, -1], [0, 0, 0], [1, 2, 1]])
return rowGradient, colGradient
def computeSobel(imge):
"""Computes and applies Sobel operator to an image."""
N = imge.shape[0]
result = np.zeros([N, N], dtype=float)
#Copy the first and last rows, first and last columns
result[0, :] = imge[0, :]
result[:, 0] = imge[:, 0]
result[N-1, :] = imge[N-1, :]
result[:, N-1] = imge[:, N-1]
#Generate the Row and Column Gradients of Sober.
rowGradient, colGradient = generateRowColumnSobelGradients()
#Sober Operator to the image.
for i in np.arange(1, N-1):
for j in np.arange(1, N-1):
subImge = imge[i-1:i+2, j-1:j+2]
rowSum = np.sum(rowGradient * subImge)
colSum = np.sum(colGradient * subImge)
result[i, j] = math.sqrt(rowSum**2 + colSum**2)
return result
# <a id="Applying"></a>
# ## Applying The Operators
# +
#Read an image files
imge = Images.generateBlackAndWhiteSquareImage(512)
imgeWoman = Image.open(r'Images/peppers_gray.jpg') # open an image
imgeWoman = imgeWoman.convert(mode='L')
imgeCameraman = Image.open("Images/lena_gray_256.tif") # open an image
#Convert the image file to a matrix
imgeWoman = np.array(imgeWoman)
imgeCameraman = np.array(imgeCameraman)
# -
sobImge = computeSobel(imge)
sobImgeWoman = computeSobel(imgeWoman)
sobImgeCameraman = computeSobel(imgeCameraman)
kirImge = computeKirsch(imge)
kirImgeWoman = computeKirsch(imgeWoman)
kirImgeCameraman = computeKirsch(imgeCameraman)
# +
fig, axarr = plt.subplots(3, 3, figsize=[13,13])
axarr[0][0].imshow(imge, cmap=plt.get_cmap('gray'))
axarr[0][0].set_title('Original Image')
axarr[0][1].imshow(kirImge, cmap=plt.get_cmap('gray'))
axarr[0][1].set_title('Detected Edges(Sobel)')
axarr[0][2].imshow(sobImge, cmap=plt.get_cmap('gray'))
axarr[0][2].set_title('Detected Edges(Sobel)')
axarr[1][0].imshow(imgeWoman, cmap=plt.get_cmap('gray'))
axarr[1][0].set_title('Original Image')
axarr[1][1].imshow(kirImgeWoman, cmap=plt.get_cmap('gray'))
axarr[1][1].set_title('Detected Edges(Kirsch)')
axarr[1][2].imshow(sobImgeWoman, cmap=plt.get_cmap('gray'))
axarr[1][2].set_title('Detected Edges(Sobel)')
axarr[2][0].imshow(imgeCameraman, cmap=plt.get_cmap('gray'))
axarr[2][0].set_title('Original Image')
axarr[2][1].imshow(kirImgeCameraman, cmap=plt.get_cmap('gray'))
axarr[2][1].set_title('Detected Edges(Kirsch)')
axarr[2][2].imshow(sobImgeCameraman, cmap=plt.get_cmap('gray'))
axarr[2][2].set_title('Detected Edges(Sobel)')
plt.show()
# -
# As can be shown in the above results, the operators are able to detect the edges of the given images.
| Notebooks_Teoricos/Image-Processing-Operations/09-Implementing-Edge-Detection-Operators-in-Python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from nltk.corpus import webtext
from nltk.collocations import BigramCollocationFinder
from nltk.metrics import BigramAssocMeasures
from nltk.tokenize import word_tokenize, sent_tokenize
parag = """The 'Master Blaster' had once said that Dhoni is probably the best captain he had played with. And this sentence from Tendulkar speaks volume of how much Dhoni has contributed not just to Indian cricket but the sport as a whole.
By no means can Dhoni be ever said as a conventional cricketer, especially batsman, who possesses the right technique which is a requisite at the international stage. He doesn't caress the off drives or the leg glances but what he does is something which all batsmen dream about.
Despite of not looking pretty while batting, Dhoni -- over the years -- has made sure he scores runs for the team and wins India matches, which ultimately is the goal of any cricketer.
In the recent years, he has faced a lot of criticism for his batting. Questions have been raised on his approach to batting with some even suggesting that Dhoni is now an old warhorse.
However, like many great cricketers, Dhoni has never let his lips do the talking and has always answered his critics fittingly with the bat by scoring runs in bulk."""
# sent = sent_tokenize()
textwords = word_tokenize(parag)
finder = BigramCollocationFinder.from_words(textwords)
finder
finder.nbest(BigramAssocMeasures.likelihood_ratio, 50)
from nltk.corpus import stopwords
ignored_words = stopwords.words('english')
fiterstops = lambda w : len(w) < 3 or w in ignored_words
finder.apply_word_filter(fiterstops)
finder.nbest(BigramAssocMeasures.likelihood_ratio,50)
from nltk.metrics import TrigramAssocMeasures
from nltk.collocations import TrigramCollocationFinder
finder = TrigramCollocationFinder.from_words(textwords)
finder.nbest(TrigramAssocMeasures.likelihood_ratio, 10)
finder.apply_word_filter(fiterstops)
finder.nbest(TrigramAssocMeasures.likelihood_ratio, 10)
# +
# finder.apply_freq_filter(2)
# +
# finder.nbest(TrigramAssocMeasures.likelihood_ratio,2)
# -
| C22_Natural Language Processing/Bigrams and Trigrams.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# pip install pymssql
import pymssql
import pandas as pd
pd.options.display.max_columns = None
# -
load_csv = True
if load_csv == True:
df_sku = pd.read_csv('df_sku.csv')
if load_csv == False:
sqluser = input('Enter SQL User')
sqlpass = input(f'Enter Password for {sqluser}')
## instance a python db connection object- same form as psycopg2/python-mysql drivers also
conn = pymssql.connect(server="192.168.254.13", user=sqluser,password=<PASSWORD>, port=1433) # You can lookup the port number inside SQL server.
stmt = "SELECT \
site_sk \
,datetran_sk \
,time_sk \
,salesevent_sk as transaction_id \
,master \
,parent \
,category \
,itemcat::int as itemcat \
,plu::bigint as plu \
,itemdesc \
FROM gate.fact_trandetail td \
inner join gate.dim_tranitem ti on td.plu_sk = ti.plu_sk \
where ti.category_sk != -2 and master in ('Merchandise','QSR') \
limit 100000;"
# Excute Query here
df_sku = pd.read_sql(stmt,conn)
df_sku.to_csv('df_sku.csv')
df_sku.head()
from sklearn.model_selection import train_test_split
from sklearn.naive_bayes import MultinomialNB
from sklearn.pipeline import Pipeline
from sklearn import metrics
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.feature_extraction.text import TfidfTransformer
X_train, X_test, y_train, y_test = train_test_split(df_sku['LongDescription'], df_sku['Category'], train_size=0.8)
# <h1>Naive Bayes Classifier</h1>
# +
text_clf = Pipeline([('vect', CountVectorizer()),
('tfidf', TfidfTransformer()),
('clf', MultinomialNB()),
])
text_clf.fit(X_train, y_train)
predicted = text_clf.predict(X_test)
print(metrics.classification_report(y_test, predicted))
# -
# <h1>K-nearest Neighbor</h1>
# +
from sklearn.neighbors import KNeighborsClassifier
text_clf = Pipeline([('vect', CountVectorizer()),
('tfidf', TfidfTransformer()),
('clf', KNeighborsClassifier()),
])
text_clf.fit(X_train, y_train)
predicted = text_clf.predict(X_test)
print(metrics.classification_report(y_test, predicted))
# -
# <h1>Support Vector Machine (SVM)</h1>
from sklearn.svm import LinearSVC
# +
text_clf = Pipeline([('vect', CountVectorizer()),
('tfidf', TfidfTransformer()),
('clf', LinearSVC()),
])
text_clf.fit(X_train, y_train)
predicted = text_clf.predict(X_test)
print(metrics.classification_report(y_test, predicted))
# -
# <h1>Decision Tree</h1>
# +
from sklearn import tree
text_clf = Pipeline([('vect', CountVectorizer()),
('tfidf', TfidfTransformer()),
('clf', tree.DecisionTreeClassifier()),
])
text_clf.fit(X_train, y_train)
predicted = text_clf.predict(X_test)
print(metrics.classification_report(y_test, predicted))
# -
# <h1>Random Forest</h1>
# +
from sklearn.ensemble import RandomForestClassifier
text_clf = Pipeline([('vect', CountVectorizer()),
('tfidf', TfidfTransformer()),
('clf', RandomForestClassifier(n_estimators=100)),
])
text_clf.fit(X_train, y_train)
predicted = text_clf.predict(X_test)
print(metrics.classification_report(y_test, predicted))
# -
# <h1>Deep Neural Networks</h1>
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras.layers import Dropout, Dense
from tensorflow.keras.models import Sequential
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.preprocessing import LabelEncoder
from sklearn.preprocessing import OrdinalEncoder
import numpy as np
from sklearn import metrics
def TFIDF(X_train, X_test,MAX_NB_WORDS=75000):
vectorizer_x = TfidfVectorizer(max_features=MAX_NB_WORDS)
X_train = vectorizer_x.fit_transform(X_train).toarray()
X_test = vectorizer_x.transform(X_test).toarray()
print("tf-idf with",str(np.array(X_train).shape[1]),"features")
return (X_train,X_test)
#prepare target
def prepare_targets_le(y_train, y_test):
# need to make sure we force a 2D array or we'll run into trouble with LE
y_train = y_train.to_numpy().reshape(-1,1)
y_test = y_test.to_numpy().reshape(-1,1)
le = LabelEncoder()
le.fit(y_train)
y_train_enc = le.transform(y_train)
y_test_enc = le.transform(y_test)
return y_train_enc, y_test_enc
#prepare target
def prepare_targets_oe(y_train, y_test):
oe = OrdinalEncoder()
oe.fit(y_train)
y_train_enc = oe.transform(y_train)
y_test_enc = oe.transform(y_test)
return y_train_enc, y_test_enc
def Build_Model_DNN_Text(shape, nClasses, dropout=0.5):
"""
buildModel_DNN_Tex(shape, nClasses,dropout)
Build Deep neural networks Model for text classification
Shape is input feature space
nClasses is number of classes
"""
model = Sequential()
node = 2500 # number of nodes
nLayers = 1 # number of hidden layer
model.add(Dense(node,input_dim=shape,activation='relu'))
model.add(Dropout(dropout))
for i in range(0,nLayers):
model.add(Dense(node,input_dim=node,activation='relu'))
model.add(Dropout(dropout))
model.add(Dense(nClasses, activation='softmax'))
model.compile(loss='sparse_categorical_crossentropy',
optimizer='adam',
metrics=['accuracy'])
return model
# +
X_train_tfidf,X_test_tfidf = TFIDF(X_train,X_test)
y_train_enc, y_test_enc = prepare_targets_oe(y_train, y_test)
model_DNN = Build_Model_DNN_Text(X_train_tfidf.shape[1], 29) # 29 is df_sku['Category'].nunique()
model_DNN.fit(X_train_tfidf, y_train_enc,
validation_data=(X_test_tfidf, y_test_enc),
epochs=10,
batch_size=128,
verbose=2)
predicted = model_DNN.predict(X_test_tfidf)
print(metrics.classification_report(y_test, predicted))
# -
df_sku['Category'].nunique()
import numpy as np
#y_train = y_train.values.reshape(-1,1)
#y_test = y_test.values.reshape(-1,1)
X_train_tfidf,X_test_tfidf = TFIDF(X_train,X_test)
y_train_enc, y_test_enc = prepare_targets_oe(y_train, y_test)
y_test_enc = y_test_enc.astype(float)
y_test_enc = y_test_enc.flatten()
y_test_enc.shape
predicted.shape
# +
#[i for i in y_test_enc ]
# -
pred = np.argmax(predicted,axis=1)
# +
predicted = model_DNN.predict(X_test_tfidf)
print(metrics.classification_report(y_test_enc, pred))
# -
def prepare_inputs(X_train, X_test):
oe = OrdinalEncoder()
oe.fit(X_train)
X_train_enc = oe.transform(X_train)
X_test_enc = oe.transform(X_test)
return X_train_enc, X_test_enc
from sklearn.preprocessing import LabelEncoder
# prepare target
def prepare_targets(y_train, y_test):
le = LabelEncoder()
le.fit(y_train)
y_train_enc = le.transform(y_train)
y_test_enc = le.transform(y_test)
return y_train_enc, y_test_enc
# prepare input data
X_train_enc, X_test_enc = prepare_inputs(X_train, X_test)
# prepare output data
y_train_enc, y_test_enc = prepare_targets(y_train, y_test)
# define the model
model = Sequential()
model.add(Dense(10, input_dim=X_train_enc.shape[1], activation='relu', kernel_initializer='he_normal'))
model.add(Dense(1, activation='sigmoid'))
# compile the keras model
model.compile(loss='binary_crossentropy', optimizer='adam', metrics=['accuracy'])
# fit the keras model on the dataset
model.fit(X_train_enc, y_train_enc, epochs=100, batch_size=16, verbose=2)
# evaluate the keras model
_, accuracy = model.evaluate(X_test_enc, y_test_enc, verbose=0)
print('Accuracy: %.2f' % (accuracy*100))
from tpot import TPOTClassifier
pipeline_optimizer = TPOTClassifier()
pipeline_optimizer = TPOTClassifier(generations=5, population_size=20, cv=5,
random_state=42, verbosity=2)
X_train, X_test, y_train, y_test = train_test_split(df_sku['LongDescription'], df_sku['Category'], train_size=0.8)
from sklearn.feature_extraction.text import TfidfVectorizer
import numpy as np
X_train, X_test = TFIDF(X_train, X_test,MAX_NB_WORDS=75000)
X_train
y_train, y_test = prepare_targets(y_train, y_test)
pipeline_optimizer.fit(X_train, y_train)
print(pipeline_optimizer.score(X_test, y_test))
pipeline_optimizer.export('tpot_exported_pipeline.py')
| research/FinalProject-ItemClass.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Load M19 dataset
# This notebook describes how to load the standardized M19 dataset.
# +
import os
import pickle
import pandas as pd
import draftsimtools as ds
# -
# Show standardized dataset
data_folder = "./standardized_m19/"
display(os.listdir(data_folder))
# ### 1. Load draft ratings
cur_set = pd.read_csv(data_folder + 'standardized_m19_rating.tsv', delimiter="\t")
display(cur_set)
# ### 2. Load drafts with cardnames
def load_data(path):
"""
Load a pickle file from disk.
"""
with open(path, "rb") as f:
return pickle.load(f)
drafts_train = load_data(data_folder + 'drafts_train.pkl')
drafts_test = load_data(data_folder + 'drafts_test.pkl')
# Show the train/test split
print(len(drafts_train), len(drafts_test))
# Show the first 2 picks of the first draft
print(drafts_train[0][:2])
# ### 3. Load drafts with indices
# The drafts_tensor format replaces cardnames with indices.
#
# This was previously referred to as the "Intermediate Draft Representation"
drafts_tensor_train = load_data(data_folder + 'drafts_tensor_train.pkl')
drafts_tensor_test = load_data(data_folder + 'drafts_tensor_test.pkl')
# Show size of draft tensors
print(drafts_tensor_train.shape, drafts_tensor_test.shape)
# Show the first 2 picks of the first draft
drafts_tensor_train[0, :2, :]
# ### 4. Creating One Hot Encoded Dataset
#
# The one hot encoded representation is useful for training most ML models.
#
# Currently, this representation is dynamically generated from the intermediate representation.
#
# In the future, the data may be serialized in this format to improve training performance.
# First, create a cardname -> index mapping
le = ds.create_le(cur_set["Name"].values)
print(le.classes_[:5])
# Then, define dynamically generated datasets
train_dataset = ds.DraftDataset(drafts_tensor_train, le)
test_dataset = ds.DraftDataset(drafts_tensor_test, le)
# ### 5. Using the One Hot Encoded Dataset
x, y = train_dataset[10]
# #### Input Representation
# x is a vector of length 2n, where n is the number of cards in the set.
#
# x[:n] represent the counts of cards already picked by the user.
# x[n:2n] represent the cards in the current pack (1 if card is present in pack).
print(x)
# #### Output Representation
# y is a vector of length n
# y[i]=1, where i is the index of the card picked by the user.
print(y)
# #### Numpy Conversion
# By default, x and y are torch tensors. They can converted to numpy arrays using x.numpy() and y.numpy().
| bots/load_standardized_set.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 4.2 模型参数的访问、初始化和共享
# +
import torch
from torch import nn
from torch.nn import init
print(torch.__version__)
# +
net = nn.Sequential(nn.Linear(4, 3), nn.ReLU(), nn.Linear(3, 1)) # pytorch已进行默认初始化
print(net)
X = torch.rand(2, 4)
Y = net(X).sum()
# -
# ## 4.2.1 访问模型参数
print(type(net.named_parameters()))
for name, param in net.named_parameters():
print(name, param)
for name, param in net[0].named_parameters():
print(name, param.size(), type(param))
# +
class MyModel(nn.Module):
def __init__(self, **kwargs):
super(MyModel, self).__init__(**kwargs)
self.weight1 = nn.Parameter(torch.rand(20, 20))
self.weight2 = torch.rand(20, 20)
def forward(self, x):
pass
n = MyModel()
for name, param in n.named_parameters():
print(name)
# -
weight_0 = list(net[0].parameters())[0]
print(weight_0.data)
print(weight_0.grad)
Y.backward()
print(weight_0.grad)
# ## 4.2.2 初始化模型参数
for name, param in net.named_parameters():
if 'weight' in name:
init.normal_(param, mean=0, std=0.01)
print(name, param.data)
for name, param in net.named_parameters():
if 'bias' in name:
init.constant_(param, val=0)
print(name, param.data)
# ## 4.2.3 自定义初始化方法
def init_weight_(tensor):
with torch.no_grad():
tensor.uniform_(-10, 10)
tensor *= (tensor.abs() >= 5).float()
for name, param in net.named_parameters():
if 'weight' in name:
init_weight_(param)
print(name, param.data)
for name, param in net.named_parameters():
if 'bias' in name:
param.data += 1
print(name, param.data)
# ## 4.2.4 共享模型参数
linear = nn.Linear(1, 1, bias=False)
net = nn.Sequential(linear, linear)
print(net)
for name, param in net.named_parameters():
init.constant_(param, val=3)
print(name, param.data)
print(id(net[0]) == id(net[1]))
print(id(net[0].weight) == id(net[1].weight))
net.zero_grad()
x = torch.tensor([[1.]])
y = net(x).sum()
print(y)
y.backward()
print(net[0].weight.grad)
# ### 作为比较,看看不是同一个层的情况:
linear1 = nn.Linear(1, 1, bias=False)
linear2 = nn.Linear(1, 1, bias=False)
net = nn.Sequential(linear1, linear2)
print(net)
for name, param in net.named_parameters():
init.constant_(param, val=3)
print(name, param.data)
print(id(net[0]) == id(net[1]))
print(id(net[0].weight) == id(net[1].weight))
# + jupyter={"outputs_hidden": true}
net.zero_grad()
x = torch.tensor([[1.]])
y = net(x).sum()
print(y)
y.backward()
print(net[0].weight.grad)
| code/chapter04_DL_computation/4.2_parameters.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from os import listdir
from numpy import array
from keras.preprocessing.text import Tokenizer, one_hot
from keras.preprocessing.sequence import pad_sequences
from keras.models import Model, Sequential, model_from_json
from keras.utils import to_categorical
from keras.layers.core import Dense, Dropout, Flatten
from keras.optimizers import RMSprop
from keras.layers.convolutional import Conv2D
from keras.callbacks import ModelCheckpoint
from keras.layers import Embedding, TimeDistributed, RepeatVector, LSTM, concatenate , Input, Reshape, Dense
from keras.preprocessing.image import array_to_img, img_to_array, load_img
import numpy as np
# +
dir_name = '/data/train/'
# Read a file and return a string
def load_doc(filename):
file = open(filename, 'r')
text = file.read()
file.close()
return text
def load_data(data_dir):
text = []
images = []
# Load all the files and order them
all_filenames = listdir(data_dir)
all_filenames.sort()
for filename in (all_filenames):
if filename[-3:] == "npz":
# Load the images already prepared in arrays
image = np.load(data_dir+filename)
images.append(image['features'])
else:
# Load the boostrap tokens and rap them in a start and end tag
syntax = '<START> ' + load_doc(data_dir+filename) + ' <END>'
# Seperate all the words with a single space
syntax = ' '.join(syntax.split())
# Add a space after each comma
syntax = syntax.replace(',', ' ,')
text.append(syntax)
images = np.array(images, dtype=float)
return images, text
train_features, texts = load_data(dir_name)
# +
# Initialize the function to create the vocabulary
tokenizer = Tokenizer(filters='', split=" ", lower=False)
# Create the vocabulary
tokenizer.fit_on_texts([load_doc('bootstrap.vocab')])
# Add one spot for the empty word in the vocabulary
vocab_size = len(tokenizer.word_index) + 1
# Map the input sentences into the vocabulary indexes
train_sequences = tokenizer.texts_to_sequences(texts)
# The longest set of boostrap tokens
max_sequence = max(len(s) for s in train_sequences)
# Specify how many tokens to have in each input sentence
max_length = 48
def preprocess_data(sequences, features):
X, y, image_data = list(), list(), list()
for img_no, seq in enumerate(sequences):
for i in range(1, len(seq)):
# Add the sentence until the current count(i) and add the current count to the output
in_seq, out_seq = seq[:i], seq[i]
# Pad all the input token sentences to max_sequence
in_seq = pad_sequences([in_seq], maxlen=max_sequence)[0]
# Turn the output into one-hot encoding
out_seq = to_categorical([out_seq], num_classes=vocab_size)[0]
# Add the corresponding image to the boostrap token file
image_data.append(features[img_no])
# Cap the input sentence to 48 tokens and add it
X.append(in_seq[-48:])
y.append(out_seq)
return np.array(X), np.array(y), np.array(image_data)
X, y, image_data = preprocess_data(train_sequences, train_features)
# +
#Create the encoder
image_model = Sequential()
image_model.add(Conv2D(16, (3, 3), padding='valid', activation='relu', input_shape=(256, 256, 3,)))
image_model.add(Conv2D(16, (3,3), activation='relu', padding='same', strides=2))
image_model.add(Conv2D(32, (3,3), activation='relu', padding='same'))
image_model.add(Conv2D(32, (3,3), activation='relu', padding='same', strides=2))
image_model.add(Conv2D(64, (3,3), activation='relu', padding='same'))
image_model.add(Conv2D(64, (3,3), activation='relu', padding='same', strides=2))
image_model.add(Conv2D(128, (3,3), activation='relu', padding='same'))
image_model.add(Flatten())
image_model.add(Dense(1024, activation='relu'))
image_model.add(Dropout(0.3))
image_model.add(Dense(1024, activation='relu'))
image_model.add(Dropout(0.3))
image_model.add(RepeatVector(max_length))
visual_input = Input(shape=(256, 256, 3,))
encoded_image = image_model(visual_input)
language_input = Input(shape=(max_length,))
language_model = Embedding(vocab_size, 50, input_length=max_length, mask_zero=True)(language_input)
language_model = LSTM(128, return_sequences=True)(language_model)
language_model = LSTM(128, return_sequences=True)(language_model)
#Create the decoder
decoder = concatenate([encoded_image, language_model])
decoder = LSTM(512, return_sequences=True)(decoder)
decoder = LSTM(512, return_sequences=False)(decoder)
decoder = Dense(vocab_size, activation='softmax')(decoder)
# Compile the model
model = Model(inputs=[visual_input, language_input], outputs=decoder)
optimizer = RMSprop(lr=0.0001, clipvalue=1.0)
model.compile(loss='categorical_crossentropy', optimizer=optimizer)
# -
#Save the model for every 2nd epoch
filepath="org-weights-epoch-{epoch:04d}--val_loss-{val_loss:.4f}--loss-{loss:.4f}.hdf5"
checkpoint = ModelCheckpoint(filepath, monitor='val_loss', verbose=1, save_weights_only=True, period=2)
callbacks_list = [checkpoint]
# Train the model
model.fit([image_data, X], y, batch_size=64, shuffle=False, validation_split=0.1, callbacks=callbacks_list, verbose=1, epochs=50)
| floydhub/Bootstrap/bootstrap.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import inspect, os, sys, copy, pytz, re, glob, random, praw, csv
import simplejson as json
import pandas as pd
from dateutil import parser
import datetime
import matplotlib.pyplot as plt # Matplotlib for plotting
import matplotlib.dates as md
import numpy as np
import seaborn as sns
from collections import Counter, defaultdict
import re, urllib
from pathlib import Path
from datetime import datetime
import logging
utc=pytz.UTC
ENV = "production"
os.environ['CS_ENV'] = 'production'
BASE_DIR = "/home/nathan/CivilServant"
FILE_BASE_DIR="/home/nathan/reddit_archive/"
sys.path.append(BASE_DIR)
subreddit_id = "2qr7i"
LOG_PATH = str(Path(BASE_DIR, "logs", "praw_messenger_%s.log" % ENV))
logging.basicConfig(filename=LOG_PATH, level=logging.INFO)
logging.getLogger().addHandler(logging.StreamHandler(sys.stdout))
ARCHIVE_DIR = "/home/nathan/reddit_archive/03.2017"
with open(os.path.join(BASE_DIR, "config") + "/{env}.json".format(env=ENV), "r") as config:
DBCONFIG = json.loads(config.read())
### LOAD SQLALCHEMY
from sqlalchemy import create_engine
from sqlalchemy.orm import sessionmaker
from sqlalchemy import text, and_, or_
from app.models import Base, SubredditPage, FrontPage, Subreddit, Post, ModAction, Experiment
from utils.common import PageType
db_engine = create_engine("mysql://{user}:{password}@{host}/{database}".format(
host = DBCONFIG['host'],
user = DBCONFIG['user'],
password = DBCONFIG['password'],
database = DBCONFIG['database']))
DBSession = sessionmaker(bind=db_engine)
db_session = DBSession()
### LOAD PRAW
import reddit.connection
conn = reddit.connection.Connect(base_dir=BASE_DIR)
r = conn.connect(use_db_keys=False)
#notebook_dir = os.getcwd()
#os.chdir(notebook_dir)
#import praw.errors
### FILTER OUT DEPRECATION WARNINGS ASSOCIATED WITH DECORATORS
# https://github.com/ipython/ipython/issues/9242
#import warnings
#warnings.filterwarnings('ignore', category=DeprecationWarning, message='.*use @default decorator instead.*')
# -
# # Sample Recent r/feminism newcomer commenters
# ### Data integrity checks and load background data
# +
## LOOK AT HOW RECENT OUR COMMENTS DATASET IS (HOPE THAT WE HAVE ALL THE WAY TO THE BEGINNING OF MARCH)
## LOOK AT HOW RECENT OUR SUBMISSIONS DATASET IS (HOPE THAT WE HAVE ALL THE WAY TO THE BEGINNING OF MARCH)
## CONSTRUCT A MEASURE OF THE SIX MONTH WINDOW (FROM Feminism Newcomer Analysis)
# +
begin_date = '2018-03-01'
query_text = """
select * from comments
WHERE subreddit_id = '2qr7i'
AND created_at >= {begin_date}
ORDER BY created_utc ASC;
""".format(begin_date = begin_date)
## ADDED THE SECOND CREATED_AT TO PRESERVE EXPERIMENT INTEGRITY
database_comments = []
for row in db_engine.execute(text(query_text)):
comment = {}
comment_data = json.loads(row['comment_data'])
for key in comment_data.keys():
comment[key] = comment_data[key]
for key in row.keys():
comment[key]=row[key]
comment['created'] = comment['created_utc']
database_comments.append(comment)
print("Loaded {0} Comments".format(len(database_comments)))
# +
file_comments = []
comment_ids = set()
comment_count = 0
comments_filename="full_feminism_comments_through_02_2018.json"
with open(os.path.join(FILE_BASE_DIR, "selected_output", comments_filename), "r") as f:
for line in f:
item = json.loads(line)
if(item['id'] not in comment_ids):
item['created'] = datetime.utcfromtimestamp(float(item['created_utc']))
item['body.charlength'] = len(item['body'])
#item['body'] = None
file_comments.append(item)
comment_ids.add(item['id'])
comment_count += 1
#file_comments = sorted(file_comments, key = lambda x: x['created'])
print("Loaded {0} Comments".format(len(file_comments)))
print("Loaded {0} Comment lines".format(comment_count))
# -
# ### Add database comments to file-loaded comments
# +
keys_to_include = []
for key in file_comments[0].keys():
if key in database_comments[0].keys():
keys_to_include.append(key)
#print("Keys in file and in database: [{0}]".format((",").join(keys_to_skip)))
all_comments = file_comments
added_comment_ids = 0
duplicate_comment_ids = 0
for comment in database_comments:
if comment['id'] not in comment_ids:
trimmed_comment = {}
for key in keys_to_include:
trimmed_comment[key] = comment[key]
all_comments.append(trimmed_comment)
comment_ids.add(comment['id'])
added_comment_ids += 1
else:
duplicate_comment_ids += 1
print("Merged comment datasets. Added {0} comments and skipped {1} duplicates".format(
added_comment_ids, duplicate_comment_ids))
## NOW SORT COMMENTS
all_comments = sorted(all_comments, key = lambda x: x['created'])
# -
# ### Record the number of previous comments that a participant made in the prior six months
# +
def previous_actions():
return {"comments":[], "posts":[]}
author_records = defaultdict(previous_actions)
for item in all_comments:
author_id = item['author']
author_records[author_id]['comments'].append(item)
#one_eighty_days in seconds
def count_if_eligible(current, comparator):
one_eighty_days = 60*60*24*180
if(current['created'] > comparator['created'] and
(current['created'] - comparator['created']).total_seconds()<one_eighty_days):
return 1
return 0
earliest_date = all_comments[0]['created']
sys.stdout.write("\ncomments")
sys.stdout.flush()
items_processed = 0
for item in all_comments:
previous_comments = 0
for comment in author_records[item['author']]['comments']:
if(item['created'] > comment['created']):
previous_comments += count_if_eligible(item, comment)
items_processed += 1
item['previous.comments'] = previous_comments
item['eligible'] = item['created'] > earliest_date
if(items_processed % 1000 == 0):
sys.stdout.write(".")
sys.stdout.flush()
# -
# # Create a Sample of Recent Commenters
# First time commenters within the last 45 days
recent_comment_in_seconds = 60*60*24*45
current_time = datetime.utcnow()
# +
eligible_comments = [x for x in all_comments if x['eligible'] and
(current_time - x['created']).total_seconds() < recent_comment_in_seconds and
x['previous.comments']==0]
print("Total comments: {0}. Eligible comments: {1}".format(
len(all_comments),
len(eligible_comments)))
# -
# ### Label comments by how many weeks previously they commented for the first time (for a stratified random sample)
# +
one_week_in_seconds = 60*60*24*7
for comment in eligible_comments:
comment['first.comment.week.diff'] = int((current_time - comment['created']).total_seconds() /
one_week_in_seconds)
# -
plt.hist([comment['first.comment.week.diff'] for comment in eligible_comments])
plt.title("How many weeks previously was this comment posted")
plt.show()
# ### Create a stratified random sample of accounts to message, based on the elapsed number of weeks since they originally posted their first comment
# Later, we may want to do a post-hoc analysis of how many comments they had previously made in the subreddit. Note that we will want to include them int he sample even if their account doesn't exist, because we're going to be including those kinds of accounts in the study, and we will want to think about that attrition as an expected part of the sample.
eligible_by_week = {}
for key in set([x['first.comment.week.diff'] for x in eligible_comments]):
eligible_by_week[key] = [x['author'] for x in eligible_comments if
x['first.comment.week.diff'] == key]
for key in sorted(eligible_by_week.keys()):
print("Week age {0}: {1} authors ".format(key, len(eligible_by_week[key])))
# +
## Decision: survey 100 authors per group
## Only survey the last 4 weeks
random.seed(880442) # current milliseconds at code time
sample_size = 100
selected_by_week = {}
for week in sorted(eligible_by_week.keys()):
if week <=4:
selected_by_week[week] = random.sample(eligible_by_week[week], sample_size)
for week in sorted(selected_by_week.keys()):
print("Week age {0}: {1} authors ".format(week, len(selected_by_week[week])))
# -
# # Construct and personalize the message
# ## Construct the message template
MESSAGE_SUBJECT = "Welcome! Help r/feminism understand first-time commenters with 3 questions"
MESSAGE_TEMPLATE = """Dear {username},
Hello and welcome from r/feminism!
Might you answer a 2 minute, 3-question survey to help us learn more about your experience? We're' currently working with researchers at Princeton University to improve newcomer experiences.
[Learn more and answer the survey here]({url}). Thanks!
-- [CivilServant](http://civilservant.io/) and the [r/feminism moderators](https://www.reddit.com/r/feminism/about/moderators)
"""
print(MESSAGE_TEMPLATE.format(username="natematias",
url="https://docs.google.com/forms/d/e/1FAIpQLSfmgOcsF0ALqWGPHGpJq2cwUr-xIG-uizbQqXPzPuTkrPM-Lg/viewform?usp=pp_url&entry.360321409={username}&entry.1933578897".format(
username="natematias")))
r.send_message("natematias", MESSAGE_SUBJECT, MESSAGE_TEMPLATE.format(username="natematias"))
# ## Add the ability to validate users
def get_reddit_user(username):
try:
user_dict = r.get_redditor(username, fetch=True).json_dict
logging.info("User %s found: %s" %(username, str(user_dict)))
return user_dict
except praw.errors.NotFound as e:
logging.error("User not found: %s" % username)
except Exception as e:
logging.exception("Failed to retrieve user")
import time
send_log_dir = "/home/nathan/CivilServant-Analysis/SOC412/feminism"
send_log_filename = "message_sender_feminism_04_17_2018.csv"
def sent_message_previously(username):
with open(os.path.join(send_log_dir, send_log_filename), "r") as f:
send_log = []
for row in csv.DictReader(f):
send_log.append(row)
if(username == row['username']):
return True
return False
# ## Add the ability to send messages
## RETURN TRUE IF MESSAGE WAS SENT SUCCESSFULLY
def send_reddit_message(**message_data):
user = message_data["username"]
message = MESSAGE_TEMPLATE.format(**message_data)
try:
logging.info("Sending a message to user %s with data %s" % (user, str(message_data)))
response = r.send_message(user, MESSAGE_SUBJECT, message, raise_captcha_exception=True)
if response["errors"]:
logging.error("Error in response when sending a message to user %s: %s" % (user, str(response)))
return False
else:
logging.info("Message successfully sent to user %s" % user)
return True
except praw.errors.InvalidCaptcha as e:
logging.exception("Message sending requires a captcha")
logging.error(e.response)
return False
except Exception as e:
logging.exception("Failed to send message to %s" % user)
return False
# ## Wrap the messenger function with anything survey-specific
def send_survey_message(**message_data):
message = MESSAGE_TEMPLATE.format(**message_data)
sent_status = send_reddit_message(**message_data)
with open(os.path.join(send_log_dir,send_log_filename), "a") as f:
print("APPENDING TO FILE: {0}".format(message_data))
f.write(",".join(map(str, [
message_data['username'],
True, #account exists
message_data['get_info_date'],
message_data['deleted'],
message_data['url'],
message_data['first_comment_week'],
sent_status
])) + "\n")
f.close()
logging.info("first_comment_week: %s" % message_data["first_comment_week"])
logging.info("url: %s" % message_data["url"])
logging.info("get_info_data: %s" % message_data["get_info_date"])
logging.info("deleted: %s" % message_data["deleted"])
def decide_and_send_to_user(username, first_comment_week):
if(sent_message_previously(username)):
logging.info("User %s previously received a message" % username)
return None
else:
time.sleep(2)
user = get_reddit_user(username)
if user:
send_survey_message(
username = username,
first_comment_week = first_comment_week,
get_info_date = str(datetime.utcnow()),
deleted = False,
url = "https://docs.google.com/forms/d/e/1FAIpQLSfmgOcsF0ALqWGPHGpJq2cwUr-xIG-uizbQqXPzPuTkrPM-Lg/viewform?usp=pp_url&entry.360321409={username}&entry.1933578897".format(
username=username))
else:
with open(os.path.join(send_log_dir, send_log_filename), "a") as f:
f.write(",".join(map(str, [
username,
False,
str(datetime.utcnow()),
None,
None,
first_comment_week,
False
])) + "\n")
f.close()
## TEST SENDING
#decide_and_send_to_user("natematias", 1)
sum([len(x) for x in selected_by_week.values()])
# # Send Surveys to Participants
# +
# import time
# for week in sorted(selected_by_week.keys()):
# print("Week age {0}: {1} authors ".format(week, len(selected_by_week[week])))
# for username in selected_by_week[week]:
# decide_and_send_to_user(username, week)
# +
#x = r.send_message("natematias", "test message", "test body")
# +
#x = r.send_message("natematias1234", "test message subject", "test body")
# -
# ### Merge Historical Data with Survey Results (later)
survey_responses = {}
with open("feminism/merged_survey_responses-04.24.2018-07.52-ET.csv", "r") as f:
for row in csv.DictReader(f):
survey_responses[row['account']] = row
# #### Load mod_actions
# +
import datetime
recent_mod_actions = []
for row in db_engine.execute(text("""
SELECT action_data FROM mod_actions
WHERE subreddit_id="2qr7i"
AND created_utc >= "2018-03-01"
ORDER BY created_utc;
""")):
mod_action = json.loads(row['action_data'])
mod_action['created'] = utc.localize(datetime.datetime.utcfromtimestamp(mod_action['created_utc']))
recent_mod_actions.append(mod_action)
print("{0} moderator actions loaded".format(len(recent_mod_actions)))
# +
comment_dict = {}
for comment in all_comments:
comment['visible'] = True
comment_dict[comment['id']] = comment
for action in recent_mod_actions:
if action['action'] == "removecomment":
key = action['target_fullname'].replace("t1_","")
if key in comment_dict.keys():
comment_dict[key]['visible'] = False
elif action['action'] == 'approvecomment':
key = action['target_fullname'].replace("t1_","")
if key in comment_dict.keys():
comment_dict[key]['visible'] = True
# +
author_comments = defaultdict(list)
for comment_id, comment in comment_dict.items():
author_comments[comment['author']].append(comment)
for key, comments in author_comments.items():
author_comments[key] = sorted(comments, key=lambda x: x['created'])
# -
# ### Merge comment dataset with survey responses
author_comment_dataset = []
for author, survey in survey_responses.items():
if(author in author_comments.keys()):
# select the last comment they made before the survey
comments = copy.copy(author_comments[author])
for comment in comments:
row = copy.copy(comment)
row.update(survey)
author_comment_dataset.append(row)
pd.DataFrame(author_comment_dataset).to_csv("feminism/author_survey_comments-04.25.2018.csv")
survey
| research-designs/SOC412/Feminism Messaging Pilot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="nNrYMhaH6YsX"
# # SLU18 - Hyperparameter Tuning : Learning notebook
#
# ### New concepts in this unit
#
# * Hyperparameter definition
# * Hyperparameter search
# * Model selection
#
# ### New tools in this unit
# - [GridSearchCV](http://scikit-learn.org/stable/modules/generated/sklearn.model_selection.GridSearchCV.html)
# - [RandomizedSearchCV](http://scikit-learn.org/stable/modules/generated/sklearn.model_selection.RandomizedSearchCV.html)
# + colab={} colab_type="code" id="B-6EzGZX6YsZ"
from IPython.display import Image
import warnings
warnings.simplefilter("ignore")
import pandas as pd
# + [markdown] colab_type="text" id="LZCL3bsB6Yse"
# ### Hyperparameter Definition
# + [markdown] colab_type="text" id="c5W4zSUW6Ysf"
# What are hyperparameters? Up until now we have mostly trained estimators (our machine learning models) like this:
# + colab={} colab_type="code" id="rn7hajY66Ysh"
from sklearn import tree
classifier = tree.DecisionTreeClassifier( )
# + [markdown] colab_type="text" id="Wv3EsCzF6Ysk"
# What this means is we are creating a Decision Tree for a clasification problem **using its default settings**. However, every single kind of model we use can be *tweaked* and modified to better adapt to our specific problem. For example, we can specify that we want a decision tree with a maximum depth of 5,
# + colab={} colab_type="code" id="-YMPEPmE6Ysm"
classifier = tree.DecisionTreeClassifier(max_depth=5)
# + [markdown] colab_type="text" id="XSKsBCUY6Ysp"
# The knobs and parameters we can specify when creating a model are called **hyperparameters**. Part of a Data Scientist's job is to figure out the right set of hyperparameters that make our model perform better.
# + [markdown] colab_type="text" id="2IBpxf546Ysr"
# ### Loading the data
# + [markdown] colab_type="text" id="qnxifsLY6Yss"
# In this unit we will use the [Wisconsin Breast Cancer Dataset](https://archive.ics.uci.edu/ml/datasets/Breast+Cancer+Wisconsin+(Diagnostic). It's a dataset containing measurements done on microscopic images of tissue cells from people with and without breast cancer. The dataset can thus be used to try to predict whether a patient has breast cancer or not.
#
# The images look like the ones below,
#
# 
# + colab={} colab_type="code" id="57n0zhqe6Ysv"
from sklearn.datasets import load_breast_cancer
cancer_data = load_breast_cancer()
cancer = pd.DataFrame(cancer_data["data"],
columns=cancer_data["feature_names"])
cancer["malign"] = cancer_data.target
cancer["malign"] = cancer["malign"].replace({0:1, 1:0})
# + colab={"base_uri": "https://localhost:8080/", "height": 258} colab_type="code" id="auzSaaAO6Ysz" outputId="90153310-2625-4763-c4c4-7df2969e2368"
cancer.head()
# + colab={} colab_type="code" id="3XNwDjZZ6Ys6"
target_variable = "malign"
independent_variables = cancer.drop(target_variable, axis=1).columns
# + colab={} colab_type="code" id="8LcTf2Ir6Ys-"
# Train-test split our dataset
from sklearn.model_selection import train_test_split
# To give us some performance information
from sklearn.metrics import classification_report
from sklearn.tree import DecisionTreeClassifier
# + [markdown] colab_type="text" id="behl-niZ6YtB"
# We keep a portion of the dataset for validation of the final model.
# + colab={} colab_type="code" id="7U2MoWMs6YtD"
X_train, X_test, y_train, y_test = train_test_split(
cancer[independent_variables],
cancer[target_variable],
test_size=0.2,
random_state=42
)
# -
# When dealing with continous hyperparameters, you should use continuous distributions as well.
# +
from scipy.stats import uniform
uniform(1, 100)
# + [markdown] colab_type="text" id="B1TP0poF6YtG"
# ### Hyperparameters search
#
# So we have said that finding the right set of hyperparameters is part of the job of building a good estimator. However, there are tons of different hyperparameters we can set when training a model.
#
# In a jupyter notebook, we can use `?` at the end of a defined class/function to see its documentation.
#
# For example, if we want to check the options for a `DecisionTreeClassifier`, we can do so like this:
# + colab={} colab_type="code" id="i9Nqziko6YtH"
# tree.DecisionTreeClassifier?
# + [markdown] colab_type="text" id="iHQJkueY6YtL"
# So we have a lot of hyperparameters to choose! How can we do so without going crazy?. Well, fortunately we can search them automatically! Scikit-learn provides 2 different kinds of hyperparameter search strategies:
# + [markdown] colab_type="text" id="0Akk23zr6YtM"
# ### Grid Search
# + [markdown] colab_type="text" id="Ps1agzUA6YtN"
# When we perform a grid search, we basically define a list of posible values for our hyperparameters, and we test all of their possible combinations. We test them by training the estimator with those hyperparameters and evaluating its performance by doing cross validation.
#
# So for example, if we have 2 hyperparameters that we want to search, a grid search would look as follows (every blue dot would be an experiment).
#
# + colab={"base_uri": "https://localhost:8080/", "height": 482} colab_type="code" id="NeLeKlts6YtO" outputId="2a0b45d2-adb5-46a0-cd65-f594f934bfd1"
Image("media/grid_search.png")
# + [markdown] colab_type="text" id="rLkkSgJJ6YtU"
# In order to do a search, we need to define a hyperparameter space, that is, all the hyperparameters we want to test and their possible values. Be aware that each hyperparameter is of a different type, so checking the model's documentation is a good idea.
# + colab={} colab_type="code" id="_Shp_5CI6YtV"
grid_search_parameter_space = {'max_depth': range(1, 10),
'max_features': range(1, len(independent_variables))
}
# + [markdown] colab_type="text" id="I8Pn8_ye6YtZ"
# We also need to define the model that we are going to use. In this case we will use a simple DecisionTreeClassifier.
# + colab={} colab_type="code" id="YMFPU9fx6Yta"
estimator = DecisionTreeClassifier()
# + colab={} colab_type="code" id="1KzRKRt_6Yte"
# Import the GridSearchCV class from sklearn
from sklearn.model_selection import GridSearchCV
# Check GridSearchCV usage
# GridSearchCV?
# + [markdown] colab_type="text" id="NiCehEf06Yti"
# Now we can define the grid search with cross validation. We need to specify the metric we want to use to guide the process. In this case we choose the AUC score. We can also specify how many CV partitions we want to use to evaluate each hyperparameters combination.
# + colab={} colab_type="code" id="iWCycgHE6Ytj"
grid_search = GridSearchCV(
estimator,
grid_search_parameter_space,
cv=5,
scoring="roc_auc",
return_train_score=True
)
# + [markdown] colab_type="text" id="xaCk5R_N6Ytq"
# **NOTE: The %%timeit magic**
#
# In the real world, when doing any kind of data intensive task, such as running a hyperparameter search, or training a model, processing time matters. That is the time it actually takes for the computer(s) to perform the task.
#
# When using jupyter notebook, we can use the cell magic `%%timeit` to check how long a cell takes to run. %%timeit takes two main arguments, **n** (the number of loops to run) and **r** (the number of repetitions per loop). Usually you want to run a cell as many times as it is reasonable (if you dont specify *n* or *r*, jupyter will figure out how many times to run it), to get a fair estimate of how long the cell takes to run. Think of it as cross validation for computing time!
# + colab={"base_uri": "https://localhost:8080/", "height": 88} colab_type="code" id="qHkpFOTZ6Ytw" outputId="30887a0e-0ae0-4485-84de-7db62b0514fa"
# %%timeit -n 1 -r 1
grid_search.fit(X_train, y_train)
# + [markdown] colab_type="text" id="Yq0oc13J6Yt2"
# We see it takes about 5 seconds to run the grid search.
#
# We can access the best estimator found by the search with the `best_estimator_` param.
# + colab={"base_uri": "https://localhost:8080/", "height": 119} colab_type="code" id="HUQHtcH06Yt3" outputId="b9a8a27e-5d46-4d1a-8167-203832579e46"
grid_search.best_estimator_
# + [markdown] colab_type="text" id="YM9hbczm6Yt7"
# We can use the fitted grid search to predict using the estimator with the best found parameters
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="Sl22iB8X6Yt8" outputId="691a7e04-6c8a-4ea2-dcd6-a15094b4e62b"
grid_search.predict(X_test)[:10]
# + [markdown] colab_type="text" id="ITMMcwze6YuB"
# We can also see the parameters for the best performing model.
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="kPAWy0bS6YuC" outputId="246c8282-5bce-48de-8723-64ffbc12502c"
grid_search.best_params_
# + [markdown] colab_type="text" id="1vNv0QAj6YuG"
# And the best model's score.
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="muaVsSB86YuH" outputId="807d8e72-cbdd-4fc2-aad5-525ed3228e68"
grid_search.best_score_
# + [markdown] colab_type="text" id="LZB9VFxI6YuL"
# If we want to dig deeper into the search result, we can access the results obtained on each hyperparameter search iteration with `cv_results`.
# + colab={"base_uri": "https://localhost:8080/", "height": 479} colab_type="code" id="iLonYjai6YuM" outputId="7fcf103e-26df-4613-d0f9-b6e39f20ce6e"
pd.DataFrame(grid_search.cv_results_).sort_values(by="rank_test_score").head()
# + [markdown] colab_type="text" id="OthdhW-M6YuT"
# ### Randomized Search
# + [markdown] colab_type="text" id="-z8mQd_96YuV"
# Unlike the Grid Seach, Randomized Search works by randomly selecting combinations of hyperparameters. This method tends to perform better than the Grid Search when the hyperparameters space is big (and thus impractical to "brute force" the optimal solution via a Grid search).
# + [markdown] colab_type="text" id="nMVAQK5V6YuX"
# If we had 2 hyperparameters, a Randomized Search would look like this:
# + colab={"base_uri": "https://localhost:8080/", "height": 508} colab_type="code" id="X82rxZbB6YuY" outputId="cea26bec-53db-450a-f17f-4a215958cb1c"
Image("media/random_search.png")
# + [markdown] colab_type="text" id="p-d_HPTy6Yui"
# Why does a Random Search usually perform better than a GridSearch? In ideal conditions, if time/money were no issue, a Grid Search would always perform better (because it tries all of the options). However, because of time constraints, a Random Search can explore more diverse combinations of hyperparameters (and find those hyperparameters that matter the most) than a Grid Search given a specific amount of time.
# + colab={"base_uri": "https://localhost:8080/", "height": 388} colab_type="code" id="RVVjjSpb6Yuk" outputId="7e144fb6-59e2-47b1-f237-d3df7f722f86"
Image("media/grid_vs_random_search.png")
# + colab={} colab_type="code" id="kXVmVWzO6Yuq"
# Import the Random Seaarch class from sklearn
from sklearn.model_selection import RandomizedSearchCV
# Check its usage
# RandomizedSearchCV?
# + [markdown] colab_type="text" id="kHxc4LxK6Yu4"
# To run a randomized search in scikit-learn, it is recommended to use statistical distributions instead of simple lists or ranges when defining the search space.
# + colab={} colab_type="code" id="_rjs-4nY6Yu4"
from scipy.stats import randint
random_search_parameter_space_dist = {
"max_depth": randint(1, 100),
"max_features": randint(1, len(independent_variables)),
"class_weight": ["balanced", None]
}
# + [markdown] colab_type="text" id="P1jWTYai6Yu8"
# We set up the random search. We fix the random state `random_state=42` to ensure reproducibility (that is, the random search running in *your* computer should return the same results as the one running on *my* computer).
# + colab={} colab_type="code" id="cP0c8e8Z6Yu-"
randomized_search = RandomizedSearchCV(
estimator,
random_search_parameter_space_dist,
cv=5, n_iter=250,
random_state=42,
return_train_score=True
)
# + [markdown] colab_type="text" id="73OI_IFQ6YvC"
# And we run it by fitting it to the data (same as with the GridSearchCV).
# + colab={"base_uri": "https://localhost:8080/", "height": 88} colab_type="code" id="YcbvDLQO6YvI" outputId="f81841b2-df8d-44b5-b114-4cef14d1145e"
# %%timeit -n 1 -r 1
randomized_search.fit(X_train, y_train)
# + [markdown] colab_type="text" id="T03hlxCx6YvM"
# The RandomizedSearch has the same attributes as the GridSearch.
# + colab={"base_uri": "https://localhost:8080/", "height": 119} colab_type="code" id="dZmYttqi6YvM" outputId="5c80015b-222c-4b27-afd7-6f9528b41878"
randomized_search.best_estimator_
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="bpng9zjq6YvR" outputId="e24eb580-6b04-413a-d8e9-6c3c29c3e746"
randomized_search.best_score_
# + [markdown] colab_type="text" id="SWUz7bQV6YvV"
# ### Model Selection
#
# Now we have 2 possible models (the best one found with the grid search and the best one found by the Randomized Search). Which one should we choose?
#
# Selecting a "final" model that we will use is not only a matter of selecting the model with the highest score. There are other aspects we must consider when evaluating one model versus another:
#
# - Training Time: If one model takes 1 hour to train and another one takes 5 hours
# - Prediction Time: If we are working on a real time predictive system, we cannot choose a model that takes seconds to perform a prediction!
# - Interpretability: We may favor a less complex (or more interpretable) model due to regulations and/or our ability to explain it to clients
#
# + [markdown] colab_type="text" id="FuZzsZaq6YvX"
# **Measuring predictive and computing performance**
#
# We can evaluate the predictive performance of the models by using the test dataset we held at the beginning.
# + colab={} colab_type="code" id="-StU6kqq6YvX"
from sklearn.model_selection import cross_val_score, cross_validate
# + colab={} colab_type="code" id="Rd3Godrb6Yva"
grid_results = cross_validate(grid_search.best_estimator_, X_test, y_test, scoring="roc_auc",
return_train_score=True, cv=5)
# + colab={"base_uri": "https://localhost:8080/", "height": 85} colab_type="code" id="qUx40si76Yvf" outputId="5893156c-c435-4135-f9e8-ff78d67cb80e"
grid_results
# + [markdown] colab_type="text" id="ni3eDWnc6Yvn"
# We can turn these results into a dataframe and calculate their means. This way we can see how much time it takes to train the dataset, how much time it takes to predict (which matters for real time applications), and how does the model performs with the training and the test set.
# + colab={"base_uri": "https://localhost:8080/", "height": 102} colab_type="code" id="wNpZtRMw6Yvo" outputId="0afe4809-9bcf-4ad1-8290-060459b8a795"
pd.DataFrame(grid_results).mean()
# + [markdown] colab_type="text" id="pf7CAHv36Yvu"
# We can do the same thing with the randomized search estimator.
# + colab={"base_uri": "https://localhost:8080/", "height": 102} colab_type="code" id="CQjCFeFc6Yvv" outputId="c7014451-ccb3-4095-ecfc-ea3cf783a566"
random_results = cross_validate(randomized_search.best_estimator_,
X_test, y_test, scoring="roc_auc",
return_train_score=True, cv=5)
pd.DataFrame(random_results).mean()
# + [markdown] colab_type="text" id="d8mzFzAN6YwA"
# Now that we know which model performs better on the train and test set, which model is the fastest to train, we can make a more inform decision.
# + [markdown] colab_type="text" id="-B-24ffNdHTJ"
# ### CheatSheet
#
# Though we often have several hyperparameters per estimator that we could tune, in practice most ot the performance variation can be attributed to just a few hyperparameters [[2](http://proceedings.mlr.press/v32/hutter14.html)]. To make your life easier, the table below suggests a couple of hyperparameters (using sklearn naming convention), for a select group of estimators, that usually have the greatest impact on the performance.
#
# | Estimator | Hyperparameter | Notes |
# | ------------- |:-------------:| :-----|
# | Logistic Regression | penalty | Used to specify the norm used in the penalization. Can be '"l1", "l2", or "elasticnet" |
# | Logistic Regression | C | Inverse of regularization strenght. Can go from close to zero (high variance) to large values (high bias) |
# | SVM | C | Inverse of regularization strenght. Can go from close to zero (high variance) to large values (high bias)|
# | SVM | Kernel | Type of kernel to use. Can be "linear", "poly", "rbf", or "sigmoid" |
# | Tree Ensembles | n_estimators | Number of estimators to use. In practice up to hundreds of estimators are used |
# | Tree Ensembles | max_depth | Maximum depth of tree. Small values result in less complexity (1 often works well for Boosting) |
# | KNN | n_neighbors | Number of neighbors to use. Small values result in higher variace while larger ones in higher bias |
# | KNN | weights | Weight function used in prediction. Can be "uniform" or "distance" |
#
# Starting with the hyperparamters above is often a reasonable choice.
#
# + [markdown] colab_type="text" id="Z5xqYHz_bRnx"
# ### Recap
#
# * Hyperparameters define the structure of our estimators
# * Different from parameters, which are computed by the model
# * Hyperparameter search to select best hyperparameters
# * Grid search
# * Random search
# * Model selection
#
# ### Further readings
#
# * http://proceedings.mlr.press/v32/hutter14.html
# * https://en.wikipedia.org/wiki/Hyperparameter_(machine_learning)
| stats-279/SLU18 - Hyperparameter Tuning/Learning notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (Netket development)
# language: python
# name: dev-netket
# ---
# +
# Try to load netket, and install it if the import fails
try:
import netket as nk
except ImportError:
# !pip install --quiet --upgrade netket
import netket as nk
import jax
import jax.numpy as jnp
import matplotlib.pyplot as plt
# -
# (Sampler)=
# # The Sampler module
#
# ```{eval-rst}
# .. currentmodule:: netket.sampler
# ```
#
# The [Sampler](netket_sampler_api) module contains several Monte-Carlo samplers that can be used to generate samples distributed according to a certain $\log$-probability distribution function.
# The samplers defined here can be used together with the [Variational State](varstate) interface or stand-alone in your jax code. They are all fully-`jax.jit` compatible, and they can be differentiated in forward mode.
#
# The sampler module also contains infrastructure necessary to define your own samplers that are compatible with the existing API, as well as custom transition rules for the Metropolis-Hastings sampler.
#
# The inheritance diagram of the classes in this module is shown below:
# ```{eval-rst}
# .. inheritance-diagram:: netket.sampler netket.sampler.rules
# :top-classes: netket.sampler.AbstractSampler
# :parts: 1
#
# ```
#
# Following the purely functional design of Jax, we define the sampler to be a stateless collection of settings and parameters inheriting from the abstract base class {class}`Sampler`, while storing all mutable state such as the PRNG key and the statistics of acceptances in an immutable sampler state object inheriting from {class}`SamplerState`.
#
# In the documentation below we will first discuss how to construct Monte-Carlo samplers and what are their common options, then in ... we will show how to use samplers directly so that you can use them in your own projects.
# Note that if you are using the Variational State interface, you only need to know how to define samplers, because the Variational State will take care of sampling.
# ## Constructing Samplers
#
# NetKet's samplers will generate a set of samples $\sigma$ that respects the following condition:
#
# $$
# \sigma \in \mathcal{H} \ \ \ \ \ \vert\ \ \ \ p(\sigma) \sim \frac{\exp[\alpha\ \Re[\log\text{pdf}(\sigma)]]}{\sum_{\eta\in\mathcal{H}} exp[\alpha\ \Re[\log\text{pdf}(\eta)]]}
# $$
#
# All NetKet samplers generate samples from an [Hilbert space](Hilbert) object, which must be passed as the first positional argument.
# Other options that can be specified are:
#
# - The sampler's `dtype`, which is the data type used to store the arrays of samples. When working with discrete hilbert spaces such as
# {class}`nk.hilbert.Spin` and {class}`nk.hilbert.Fock` the dtype is not particularly important, and you might reduce your memory consumption by using some short
# types such as `np.int8`. By default `dtype=float64`.
# When working with discrete spaces samples are usually made up of strings of integers, therefore if you use low-precision types such as `np.int8`
# or `jnp.bfloat16` you will not be losing precision in the number you're representing. Moreover, as soon as you will feed the low-precision samples
# to your model, it will be promoted to a wider type matching the `dtype` used for the model's parameters.
#
# - The sampler's `machine_power`, which is the $\alpha$ in the formula above specifying what power of the probability distribution $\text{pdf}(\sigma)$ we are sampling.
# +
hilbert = nk.hilbert.Spin(0.5, 5)
sampler = nk.sampler.ExactSampler(hilbert)
print(sampler)
# -
# The sampler itself is a [_frozen dataclass_ ](https://stackoverflow.com/questions/66194804/what-does-frozen-mean-for-dataclasses), meaning that you cannot change it's attributes once it's created.
# To change an attribute of the sampler, you must use the function {meth}`~SamplerState.replace` which returns a new sampler object with that setting changed.
# The old sampler will be unchanged
# +
new_sampler = sampler.replace(machine_pow=3)
print("this is the old one, unchanged: ", sampler)
print("this is the new one: ", new_sampler)
# -
# This (sometimes annoying) behaviour is needed to make our sampler objects completely jax-compatible. You can freely pass a NetKet sampler to a jax function and jit it without worrying!
# ## Using Samplers
#
# The core sampler's API interface is made up of 3 functions:
# - {meth}`Sampler.init_state(log_pdf, parameters, seed) <Sampler.init_state>`, which constructs a structure holding the state of the sampler with the provided seed. The seed can either be an integer or a {func}`jax.random.PRNGKey` object. When running distributed computations on different MPI nodes, this function automatically generates a different seed on every MPI node, so you do not need to take care of that (only the seed on rank 0 will be considered).
# - {meth}`Sampler.reset(log_pdf, parameters, state) <Sampler.reset>`, is a function that should be called every time the variational parameters of the log_pdf have changed, and in some cases to reset the chain. _NOTE: The state is not valid unless you call reset at least once_
# - {meth}`Sampler.sample(log_pdf, parameters, state, chain_length=1) <Sampler.sample>`, which samples a sequence of samples with `chain_length` samples. If you are using a direct sampler like {class}`ExactSampler` or {class}`ARDirectSampler` this determines how many samples you generate. If you are using a Markov-Chain sampler you will get `Sampler.n_chains * chain_length` total samples.
#
# In general, the `log_pdf` must be a function with the signature `log_pdf(PyTree, jnp.Array[...,Sampler.hilbert.size]) -> jnp.Array[...]` or a flax Module. (Note: Some samplers such as {class}`ARDirectSampler` only work with a flax Module.)
#
# To build an example, first we must define a log-pdf from which we want to sample.
# In this case we construct an RBM with real parameters, and initialise some random parameters which will give a roughly flat distribution:
# +
# We define an Hilbert space of 3 Spin-1/2
hilbert = nk.hilbert.Spin(0.5, 4)
# We define our variational ansatz
log_pdf = nk.models.RBM(param_dtype=float)
# and we initialize it's parameters
param_seed = jax.random.PRNGKey(0)
pars = log_pdf.init(param_seed, hilbert.random_state(param_seed, 3))
# -
# Now we can plot the probability distribution by computing it over the whole Hilbert space. Of course this is expensive, but if we have a small set of variables it's possible to do it.
# +
pdf_unnormalized = jnp.exp(2*log_pdf.apply(pars, hilbert.all_states()))
pdf = pdf_unnormalized / jnp.sum(pdf_unnormalized)
plt.plot(pdf)
plt.ylim(0,0.1)
plt.xlabel("hilbert space index")
plt.ylabel("pdf")
# -
# {class}`ExactSampler` builds exactly this same probability distribution (which has an exponentially large cost in the number of variables) and generates samples from it.
# To use it we can do:
# +
# We construct an Exact Sampler
sampler = nk.sampler.ExactSampler(hilbert, dtype=jnp.int8)
# We create the state of the sampler
sampler_state = sampler.init_state(log_pdf, pars, jax.random.PRNGKey(1))
# We call reset (this will pre-compute the log_pdf on the whole hilbert space)
sampler_state = sampler.reset(log_pdf, pars, sampler_state)
# generate samples
samples, sampler_state = sampler.sample(log_pdf, pars, state=sampler_state, chain_length=10)
print(f"The shape of the samples is: {samples.shape}, and the dtype is {samples.dtype}")
print(samples[0,0])
# -
# Notice that samplers return a 3-tensor of samples where the first dimension is `chain_length`, the second is the number of parallel chains (or 1 in non Markov-Chain samplers such as ExactSampler) and the last dimension is the hilbert space size.
#
# We could verify that those samples are distributed according to the correct distribution by running the following code:
# +
def estimate_pdf(n_samples):
samples, _ = sampler.sample(log_pdf, pars, state=sampler_state, chain_length=n_samples)
# Convert the samples to indices in the space
idxs = hilbert.states_to_numbers(samples)
# count the occurrences of all indices
return jnp.sum(idxs == jnp.arange(hilbert.n_states), axis=0) / idxs.shape[0]
plt.plot(pdf, label="exact")
plt.plot(estimate_pdf(2**10), label="2^10 samples")
plt.plot(estimate_pdf(2**14), '--', label="2^14 samples")
plt.ylim(0,0.1)
plt.xlabel("hilbert space index")
plt.ylabel("pdf")
plt.legend();
# -
#
# ## Metropolis-Hastings Markov-Chain Samplers
#
# NetKet also implements a very flexible class of Markov Chain samplers that use the Metropolis-Hastings algorithm and is called {class}`MetropolisSampler`.
# The Metropolis--Hastings algorithm is used to generate samples from an arbitrary probability distribution.
# In each step, it suggests a transition from the current configuration $s$ to a proposed configuration $s'$.
# The proposal is accepted with probability
#
# $$
# P_\text{acc}(s \rightarrow s') = \min\left( 1, \frac{P(s')}{P(s)} \frac{g(s \mid s')}{g(s' \mid s)} \right),
# $$
#
# where $P$ is the distribution being sampled from and $g(s' \mid s)$ is the conditional probability of proposing $s'$ given the current $s$.
#
# We call $L(s, s') = \log [g(s \mid s')/g(s' \mid s)]$ to denote the correcting factor to the log probability due to the transition kernel.
# This factor is needed for asymmetric kernels that might propose one move with higher probability than its reverse.
# Simple kernels, such as a local spin flip or exchange, are symmetric, therefore $L(s,s') = L(s', s) = 1$, but other proposals, such as Hamiltonian sampling, are not necessarily symmetric and need this factor.
#
# The transition rules (or kernels) that NetKet implements for discrete hilbert spaces are the following:
# - {class}`rules.LocalRule`: A transition rule acting on the local degree of freedom. This selects a random degree of freedom `i` and then proposes a different local configuration with uniform probability. For example, for Spin-1/2, this will flip a random spin. **Note: This transition rule ignores constraints on the hilbert space, such as total magnetization, and might produce states that do not respect it**
# - {class}`rules.ExchangeRule`: A Rule exchanging the value of the local degree of freedom between two sites $i$ and $j$, chosen from a list of
# possible couples (clusters). To construct it, the user must provide a graph object or a list of edges. **This sampler does respect constraints on Hilbert spaces, but might not explore the whole hilbert space because it preserves the total number of excitations/magnetization.**
# - {class}`rules.HamiltonianRule`: which transitions the configuration according to the off-diagonal elements of the Hamiltonian. As this rule might not be symmetric, this uses the factor $L(s,s')$ to reweight the transition probability.
#
# For Continuous Hilbert spaces, the only rule that we implement (at the moment is the following:
# - {class}`rules.GaussianRule` which proposes a move by adding a normally-distributed random number to a single degree of freedom. The standard deviation of the Gaussian distribution can be modified.
#
# The standard syntax to build a MCMC sampler with a certain rule is to pass an instance of the rule as the second argument of the Metropolis Sampler:
#
rule = nk.sampler.rules.LocalRule()
sampler = nk.sampler.MetropolisSampler(hilbert, rule, n_chains = 10)
print(sampler)
# The Metropolis Sampler takes as the first two positional arguments the hilbert space to sample and the transition rule.
# Additional options that can be specified are the `dtype` and `machine_power`, like all other samplers, but also:
# - `n_chains: int` or `n_chains_per_rank:int `: The number of parallel Markov Chains to sample from. Depending on the complexity of the log-pdf and the size of the system, sampling from 1 chain or 10 might take the same time on CPU. Usually on GPUs you can sample from ~1000 of chains at the same cost of sampling from 1 chain. Only one of those 2 options can be specified at a given time, and they are equivalent when not using MPI. When using MPI, the first controls the total number of chains (and must be $>$ than the total number of MPI ranks) while the latter controls the number of chains per MPI rank. Defaults to `n_chains_per_rank=16`
# - `n_sweeps: int`: The number of sub-sampling sweeps per sample. This integer controls how many Metropolis-Hastings steps are taken before returning a valid sampling. Effectively, this means that when sampling a chain of length `chain_length`, the effective chain length is `chain_length*n_sweeps`, but only $1$ sample every `n_sweeps` is returned. The default value is the number of degrees of freedom in the hilbert space ({attr}`AbstractHilbert.size <netket.hilbert.AbstractHilbert.size>`).
# -`reset_chains : bool`: If True, resets the chain state when `Sampler.reset` is called. By default this is `False`, meaning that during a run the chain is never completely reset (when you reset the chain, a configuration is drawn with uniform probability from the hilbert space unless a special rule is used).
#
# Other than those additional settings at constructions, Metropolis Samplers follow the same API as all other samplers:
# +
# We create the state of the sampler
sampler_state = sampler.init_state(log_pdf, pars, jax.random.PRNGKey(1))
# We call reset (this will pre-compute the log_pdf on the whole hilbert space)
sampler_state = sampler.reset(log_pdf, pars, sampler_state)
# generate samples
samples, sampler_state = sampler.sample(log_pdf, pars, state=sampler_state, chain_length=100)
print(f"The shape of the samples is: {samples.shape}, and the dtype is {samples.dtype}")
print(samples[0,0])
# -
# However, notice that this time the samples returned are a tensor of shape `(chain_length, n_chains_per_rank, hilbert.size)`.
#
# For ease of use, we provide some shorthands such as {func}`MetropolisLocal`, {func}`MetropolisHamiltonian` and so on, which automatically build the relevant rule.
# ## Defining custom Transition Rules for Metropolis Sampler
#
# A transition kernel is a NetKet dataclass inheriting from {class}`rules.MetropolisRule` that must define the following two methods:
#
# - {meth}`MetropolisRule.init_state(self, sampler, log_pdf, params, rng) -> Optional[Any] <rules.MetropolisRule.init_state>`: which can be used to initialise some arbitrary state or cache. This will be stored inline inside of `MetropolisState`.
# - {meth}`MetropolisRule.random_state(rule, sampler, log_pdf, parameters, state, rng, σ: jnp.ndarray[n_chains, hilbert.size]) -> jnp.Array[n_chains, hilbert.size] <rules.MetropolisRule.random_state>`: this function is called when the chain is initialised or resetted, and should return a valid configuration for the Markov Chain state. The default implementation returns a state sampled with `hilbert.random_state`.
# - {meth}`MetropolisRule.transition(rule, sampler, log_pdf, parameters, state, rng: PRNGKeyT) -> jnp.Array[n_chains, hilbert.size] <rules.MetropolisRule.transition>`: this function is called when the chain is initialised or resetted, and should return a valid configuration for the Markov Chain state. The default implementation returns a state sampled with {meth}`AbstractHilbert.random_state <netket.hilbert.AbstractHilbert.random_state>`.
#
# As an example, I will define below a custom Metropolis Transition rule that flips not one, but two spins at a time
@nk.utils.struct.dataclass
class TwoLocalRule(nk.sampler.rules.MetropolisRule):
def transition(self, sampler, machine, parameters, state, key, σ):
# Deduce the number of MCMC chains from input shape
n_chains = σ.shape[0]
# Load the Hilbert space of the sampler
hilb = sampler.hilbert
# Split the rng key into 2: one for each random operation
key_indx, key_flip = jax.random.split(key, 2)
# Pick two random sites on every chain
indxs = jax.random.randint(
key_indx, shape=(n_chains, 2), minval=0, maxval=hilb.size
)
# flip those sites
σp, _ = nk.hilbert.random.flip_state(hilb, key_flip, σ, indxs)
# If this transition had a correcting factor L, it's possible
# to return it as a vector in the second value
return σp, None
#
# random_state is inherited from the base class, so no need to define it
#
#def random_state(self, sampler, machine, pars, state, key):
# return sampler.hilbert.random_state(
# key, size=sampler.n_batches, dtype=sampler.dtype
# )
# And then, in order to use this transition kernel we can follow the same procedure as before:
sampler = nk.sampler.MetropolisSampler(hilbert, TwoLocalRule())
# +
# We create the state of the sampler
sampler_state = sampler.init_state(log_pdf, pars, jax.random.PRNGKey(1))
# We call reset (this will pre-compute the log_pdf on the whole hilbert space)
sampler_state = sampler.reset(log_pdf, pars, sampler_state)
# generate samples
samples, sampler_state = sampler.sample(log_pdf, pars, state=sampler_state, chain_length=100)
| docs/docs/sampler.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/ianomunga/XOR-LSTM-Problem/blob/main/XOR_LSTM.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="Mxsrno4oUwpz"
# ##Solving the XOR Logic Gate Output Problem using an LSTM Recurrent Neural Network
# XOR stands for 'Exclusive-Or', which is a logical operator that evaluates to the 'True' Boolean output when either of its values are true; but not both. This mutual exclusivity is captured in this part of its name. 'exclusive'.
#
# This relationship is hard to represent in a linear way that a Logistic Regression Model would be able to generalize statistically for, because the statistical significance in the bits, i.e. the meaning, comes from a mutual rlationship between the two values under evaluation.
#
# A non-linearity could model this relationship, however, and that's where the LSTM Model comes in. It's 'Long Short-Term Memory' enables the cumulative evaluations of the stream of logic gates to be carried forward recurrently throughout the sequence.
#
# This is what the code below will implement.
#
# + id="szvVjUg1Ft-X"
#get all your dependencies in check
from tensorflow.keras import optimizers
from tensorflow.keras.layers import Dense, Input, LSTM
from tensorflow.keras.models import Sequential
import numpy as np
import random
# + id="tc9IxRA6GnUo"
#encapsulate some key variables, i.e
#the sequence_length
SEQ_LEN = 50
#the number of bits in the sequence
COUNT = 100000
# + id="wvRtUTokGvd_"
#create our pairs of logic gate values based on the cumulative sum of the generated sequence
bin_pair = lambda x: [x, not(x)]
training = np.array([[bin_pair(random.choice([0, 1])) for _ in range(SEQ_LEN)] for _ in range(COUNT)])
target = np.array([[bin_pair(x) for x in np.cumsum(example[:,0]) % 2] for example in training])
# + colab={"base_uri": "https://localhost:8080/"} id="ZSKczhmXG7B7" outputId="3aebc12a-669f-48c4-94b9-4942cd19d0f2"
#check for a match between the lengths of the datasets before we go ahead
print('shape check:', training.shape, '=', target.shape)
# + id="yJZnOmTJHIkb"
model = Sequential()
#pass in the sequence-length so that every possible example's dimension is accounted for
model.add(Input(shape=(SEQ_LEN, 2), dtype='float32'))
#build the model with the LSTM component for parity persistence,
model.add(LSTM(1, return_sequences=True))
#two possible outcomes for the two possible logicgate values
model.add(Dense(2, activation='softmax'))
# + colab={"base_uri": "https://localhost:8080/"} id="gp6Z1ZtyHT4d" outputId="08561dbb-8849-42bb-da04-e74156ffb1d6"
#now fit the model to the data and run the epochs
model.compile(loss='binary_crossentropy', optimizer='adam', metrics=['accuracy'])
model.fit(training, target, epochs=10, batch_size=128)
model.summary()
# + colab={"base_uri": "https://localhost:8080/"} id="XsV8th8nHb4e" outputId="9222a0da-2832-4fb5-f602-fda3217c5b23"
predictions = model.predict(training)
i = random.randint(0, COUNT)
chance = predictions[i,-1,0]
print('randomly selected sequence:', training[i,:,0])
print('prediction:', int(chance > 0.5))
print('confidence: {:0.2f}%'.format((chance if chance > 0.5 else 1 - chance) * 100))
print('actual:', np.sum(training[i,:,0]) % 2)
# + [markdown] id="skpkDbLne005"
# It can be seen that the LSTM configuration does successfully carry forward the parity of the logical gates. In the end, the model is able to predict the parity of the alternatives to a sequence of randomly generated bits with a confidence score of 99.73 percent with 100,000 sample bits serving as the examples in 50-bit sequences.
| XOR_LSTM.ipynb |
# ---
# jupyter:
# jupytext:
# formats: ipynb,md:myst
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Before you start
#
# you must run the cell below or copy the command into the terminal to use all of Jax's capabilities
# !nvidia-smi
# + [markdown] id="xtWX4x9DCF5_"
# # JAX Quickstart
#
# **JAX is NumPy on the CPU, GPU, and TPU, with great automatic differentiation for high-performance machine learning research.**
#
# With its updated version of [Autograd](https://github.com/hips/autograd), JAX
# can automatically differentiate native Python and NumPy code. It can
# differentiate through a large subset of Python’s features, including loops, ifs,
# recursion, and closures, and it can even take derivatives of derivatives of
# derivatives. It supports reverse-mode as well as forward-mode differentiation, and the two can be composed arbitrarily
# to any order.
#
# What’s new is that JAX uses
# [XLA](https://www.tensorflow.org/xla)
# to compile and run your NumPy code on accelerators, like GPUs and TPUs.
# Compilation happens under the hood by default, with library calls getting
# just-in-time compiled and executed. But JAX even lets you just-in-time compile
# your own Python functions into XLA-optimized kernels using a one-function API.
# Compilation and automatic differentiation can be composed arbitrarily, so you
# can express sophisticated algorithms and get maximal performance without having
# to leave Python.
# + id="SY8mDvEvCGqk"
import jax.numpy as jnp
from jax import grad, jit, vmap
from jax import random
# + tags=["remove-cell"]
# Prevent GPU/TPU warning.
import jax; jax.config.update('jax_platform_name', 'cpu')
# + [markdown] id="FQ89jHCYfhpg"
# ## Multiplying Matrices
# + [markdown] id="Xpy1dSgNqCP4"
# We'll be generating random data in the following examples. One big difference between NumPy and JAX is how you generate random numbers. For more details, see [Common Gotchas in JAX].
#
# [Common Gotchas in JAX]: https://jax.readthedocs.io/en/latest/notebooks/Common_Gotchas_in_JAX.html#%F0%9F%94%AA-Random-Numbers
# + id="u0nseKZNqOoH"
key = random.PRNGKey(0)
x = random.normal(key, (10,))
print(x)
# + [markdown] id="hDJF0UPKnuqB"
# Let's dive right in and multiply two big matrices.
# + id="eXn8GUl6CG5N"
size = 3000
x = random.normal(key, (size, size), dtype=jnp.float32)
# %timeit jnp.dot(x, x.T).block_until_ready() # runs on the GPU
# + [markdown] id="0AlN7EbonyaR"
# We added that `block_until_ready` because JAX uses asynchronous execution by default (see {ref}`async-dispatch`).
#
# JAX NumPy functions work on regular NumPy arrays.
# + id="ZPl0MuwYrM7t"
import numpy as np
x = np.random.normal(size=(size, size)).astype(np.float32)
# %timeit jnp.dot(x, x.T).block_until_ready()
# + [markdown] id="_SrcB2IurUuE"
# That's slower because it has to transfer data to the GPU every time. You can ensure that an NDArray is backed by device memory using {func}`~jax.device_put`.
# + id="Jj7M7zyRskF0"
from jax import device_put
x = np.random.normal(size=(size, size)).astype(np.float32)
x = device_put(x)
# %timeit jnp.dot(x, x.T).block_until_ready()
# + [markdown] id="clO9djnen8qi"
# The output of {func}`~jax.device_put` still acts like an NDArray, but it only copies values back to the CPU when they're needed for printing, plotting, saving to disk, branching, etc. The behavior of {func}`~jax.device_put` is equivalent to the function `jit(lambda x: x)`, but it's faster.
# + [markdown] id="ghkfKNQttDpg"
# If you have a GPU (or TPU!) these calls run on the accelerator and have the potential to be much faster than on CPU.
# + id="RzXK8GnIs7VV"
x = np.random.normal(size=(size, size)).astype(np.float32)
# %timeit np.dot(x, x.T)
# + [markdown] id="iOzp0P_GoJhb"
# JAX is much more than just a GPU-backed NumPy. It also comes with a few program transformations that are useful when writing numerical code. For now, there's three main ones:
#
# - {func}`~jax.jit`, for speeding up your code
# - {func}`~jax.grad`, for taking derivatives
# - {func}`~jax.vmap`, for automatic vectorization or batching.
#
# Let's go over these, one-by-one. We'll also end up composing these in interesting ways.
# + [markdown] id="bTTrTbWvgLUK"
# ## Using {func}`~jax.jit` to speed up functions
# + [markdown] id="YrqE32mvE3b7"
# JAX runs transparently on the GPU (or CPU, if you don't have one, and TPU coming soon!). However, in the above example, JAX is dispatching kernels to the GPU one operation at a time. If we have a sequence of operations, we can use the `@jit` decorator to compile multiple operations together using [XLA](https://www.tensorflow.org/xla). Let's try that.
# + id="qLGdCtFKFLOR"
def selu(x, alpha=1.67, lmbda=1.05):
return lmbda * jnp.where(x > 0, x, alpha * jnp.exp(x) - alpha)
x = random.normal(key, (1000000,))
# %timeit selu(x).block_until_ready()
# + [markdown] id="a_V8SruVHrD_"
# We can speed it up with `@jit`, which will jit-compile the first time `selu` is called and will be cached thereafter.
# + id="fh4w_3NpFYTp"
selu_jit = jit(selu)
# %timeit selu_jit(x).block_until_ready()
# + [markdown] id="HxpBc4WmfsEU"
# ## Taking derivatives with {func}`~jax.grad`
#
# In addition to evaluating numerical functions, we also want to transform them. One transformation is [automatic differentiation](https://en.wikipedia.org/wiki/Automatic_differentiation). In JAX, just like in [Autograd](https://github.com/HIPS/autograd), you can compute gradients with the {func}`~jax.grad` function.
# + id="IMAgNJaMJwPD"
def sum_logistic(x):
return jnp.sum(1.0 / (1.0 + jnp.exp(-x)))
x_small = jnp.arange(3.)
derivative_fn = grad(sum_logistic)
print(derivative_fn(x_small))
# + [markdown] id="PtNs881Ohioc"
# Let's verify with finite differences that our result is correct.
# + id="JXI7_OZuKZVO"
def first_finite_differences(f, x):
eps = 1e-3
return jnp.array([(f(x + eps * v) - f(x - eps * v)) / (2 * eps)
for v in jnp.eye(len(x))])
print(first_finite_differences(sum_logistic, x_small))
# + [markdown] id="Q2CUZjOWNZ-3"
# Taking derivatives is as easy as calling {func}`~jax.grad`. {func}`~jax.grad` and {func}`~jax.jit` compose and can be mixed arbitrarily. In the above example we jitted `sum_logistic` and then took its derivative. We can go further:
# + id="TO4g8ny-OEi4"
print(grad(jit(grad(jit(grad(sum_logistic)))))(1.0))
# + [markdown] id="yCJ5feKvhnBJ"
# For more advanced autodiff, you can use {func}`jax.vjp` for reverse-mode vector-Jacobian products and {func}`jax.jvp` for forward-mode Jacobian-vector products. The two can be composed arbitrarily with one another, and with other JAX transformations. Here's one way to compose them to make a function that efficiently computes full Hessian matrices:
# + id="Z-JxbiNyhxEW"
from jax import jacfwd, jacrev
def hessian(fun):
return jit(jacfwd(jacrev(fun)))
# + [markdown] id="TI4nPsGafxbL"
# ## Auto-vectorization with {func}`~jax.vmap`
# + [markdown] id="PcxkONy5aius"
# JAX has one more transformation in its API that you might find useful: {func}`~jax.vmap`, the vectorizing map. It has the familiar semantics of mapping a function along array axes, but instead of keeping the loop on the outside, it pushes the loop down into a function’s primitive operations for better performance. When composed with {func}`~jax.jit`, it can be just as fast as adding the batch dimensions by hand.
# + [markdown] id="TPiX4y-bWLFS"
# We're going to work with a simple example, and promote matrix-vector products into matrix-matrix products using {func}`~jax.vmap`. Although this is easy to do by hand in this specific case, the same technique can apply to more complicated functions.
# + id="8w0Gpsn8WYYj"
mat = random.normal(key, (150, 100))
batched_x = random.normal(key, (10, 100))
def apply_matrix(v):
return jnp.dot(mat, v)
# + [markdown] id="0zWsc0RisQWx"
# Given a function such as `apply_matrix`, we can loop over a batch dimension in Python, but usually the performance of doing so is poor.
# + id="KWVc9BsZv0Ki"
def naively_batched_apply_matrix(v_batched):
return jnp.stack([apply_matrix(v) for v in v_batched])
print('Naively batched')
# %timeit naively_batched_apply_matrix(batched_x).block_until_ready()
# + [markdown] id="qHfKaLE9stbA"
# We know how to batch this operation manually. In this case, `jnp.dot` handles extra batch dimensions transparently.
# + id="ipei6l8nvrzH"
@jit
def batched_apply_matrix(v_batched):
return jnp.dot(v_batched, mat.T)
print('Manually batched')
# %timeit batched_apply_matrix(batched_x).block_until_ready()
# + [markdown] id="1eF8Nhb-szAb"
# However, suppose we had a more complicated function without batching support. We can use {func}`~jax.vmap` to add batching support automatically.
# + id="67Oeknf5vuCl"
@jit
def vmap_batched_apply_matrix(v_batched):
return vmap(apply_matrix)(v_batched)
print('Auto-vectorized with vmap')
# %timeit vmap_batched_apply_matrix(batched_x).block_until_ready()
# + [markdown] id="pYVl3Z2nbZhO"
# Of course, {func}`~jax.vmap` can be arbitrarily composed with {func}`~jax.jit`, {func}`~jax.grad`, and any other JAX transformation.
# + [markdown] id="WwNnjaI4th_8"
# This is just a taste of what JAX can do. We're really excited to see what you do with it!
| notebooks/quickstart.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Import Flask
from flask import Flask, render_template, redirect
# Import our pymongo library, which lets us connect our Flask app to our Mongo database.
import pymongo
from pymongo import MongoClient
from flask_pymongo import PyMongo
#Import my scraping class
# import scrape_mars
# from scrape_mars import myClass
# +
#Get image for decoration from Activities folder Extra Content
#_________________________________________________________
from bs4 import BeautifulSoup
from splinter import Browser
executable_path = {"executable_path": 'C:/Webdriver/bin/chromedriver'}
browser = Browser("chrome", **executable_path, headless=False)
url = "https://en.wikipedia.org/wiki/Mars"
browser.visit(url)
xpath = '//td//a[@class="image"]/img'
results = browser.find_by_xpath(xpath)
img = results[0]
img.click()
url2 = browser.html
soup = BeautifulSoup(url2, 'html.parser')
img_url_temp1 = soup.find("tbody").find('a', class_='image')['href']
img_url_temp=f'https://en.m.wikipedia.org/{img_url_temp1}'
print(img_url_temp)
browser.visit(img_url_temp)
url3= browser.html
soup2= BeautifulSoup(url3, "html.parser")
img_url_file= soup2.find("div", class_='fullImageLink').find('a')['href']
img_url=f'https:{img_url_file}'
print(img_url)
import requests
import shutil
response = requests.get(img_url, stream=True)
with open('img.png', 'wb') as out_file:
shutil.copyfileobj(response.raw, out_file)
from IPython.display import Image
Decor=Image(url='img.png')
browser.quit()
#______________________________________________________________________
# +
# Create connection variable
conn = 'mongodb://localhost:27017'
# Pass connection to the pymongo instance.
client = pymongo.MongoClient(conn)
# Connect to a database. Will create one if not already available.
db = client.MarsPhotos_db
#Mars_collection that will contain the data
HemImages=db.HemImages
MarsFacts1=db.MarsFacts1
MarsFacts2=db.MarsFacts2
MarsPhotos=db.MarsPhotos
# Drops collection if available to remove duplicates
db.HemImages.drop()
db.MarsFacts1.drop()
db.MarsFacts2.drop()
db.MarsPhotos.drop()
#Initialize remote class variables
import pandas as pd
dicti={"":""}
hemispheres=[]
textoTitle=""
texto3=""
#d={"Fact":[],"Value":[]}
t_html=""
#______________________________________________________________________
#run remote class and method to obtain data scraped from websites
# scrape_mars.myClass.scrape(dicti,hemispheres,title3)
# def scrape():
# instance=myClass.scrape(dicti,hemispheres,texto3)
# print(texto3)
# return render_template("index.html", News=texto3)
# print(instance)
#Retrieving all results from remote class
myclass=myClass(dicti,hemispheres,texto3,t_html,textoTitle)
hemispheres,texto3,t_html,textoTitle = myclass.scrape(hemispheres,texto3,t_html,textoTitle)
print(hemispheres,texto3,t_html,textoTitle)
Title=textoTitle
paragraph=texto3
#making a dictionary with the latest News from Mar
News = {'Title':textoTitle, "paragraph":texto3}
MarsFacts1.insert_one(News)
for m in hemispheres:
HemImages.insert_one(m)
#change dataframe table to html table
# import io
# data = io.StringIO(t_html)
# print(data)
# df = pd.read_csv(data, sep=",", header=None)
# t_html = df.to_html()
# write html table to file
text_file = open("index.html", "w")
text_file.write(t_html)
text_file.close()
Facts={"Table":t_html}
MarsFacts2.insert_one(Facts)
print(HemImages)
#______________________________________________________________________
#__________________________________________if __name__ == "__main__":
app.run(debug=True)____________________________
# IF EVERYTHIN GOES WRONG
# db.HemImages.insert_many(
# [{'Hemisphere':'Cerberus Hemisphere', 'Image':'https://astropedia.astrogeology.usgs.gov/download/Mars/Viking/cerberus_enhanced.tif/full.jpg'},
# {'Hemisphere':'Schiaparelli Hemisphere', 'Image':'https://astropedia.astrogeology.usgs.gov/download/Mars/Viking/schiaparelli_enhanced.tif/full.jpg'},
# {'Hemisphere':'Syrtis Major Hemisphere', 'Image':'https://astropedia.astrogeology.usgs.gov/download/Mars/Viking/syrtis_major_enhanced.tif/full.jpg'},
# {'Hemisphere':'Valles Marineris Hemisphere', 'Image':'https://astropedia.astrogeology.usgs.gov/download/Mars/Viking/valles_marineris_enhanced.tif/full.jpg'}]
# )
db.MarsFacts2.insert_one({"Fact":'Equatorial Diameter= ', 'Value':'6,792 km'})
db.MarsFacts2.insert_one({"Fact":'Polar Diameter= ','Value':'6,752 km'})
db.MarsFacts2.insert_one({"Fact":'Mass= ', 'Value':'6.39 × 10^23 kg (0.11 Earths)'})
db.MarsFacts2.insert_one({"Fact":'Moons= ','Value':'6.39 × 10^23 kg (0.11 Earths)'})
db.MarsFacts2.insert_one({"Fact":'Orbit Distance= ','Value':'2 (Phobos & Deimos)'})
db.MarsFacts2.insert_one({"Fact":'Orbit Period= ','Value':'687 days (1.9 years)'})
db.MarsFacts2.insert_one({"Fact":'Surface Temperature= ','Value':'-87 to -5 °C'})
db.MarsFacts2.insert_one({"Fact":'First Record: ','Value':'2nd millennium BC'})
db.MarsFacts2.insert_one({"Fact":'Recorded By: ','Value':'Egyptian astronomers'})
# -
| test_cc_error.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Basic functions
#
# Directions are in geographic notation: 0/360 = North, 90 = East, 180 = South, 270 = West.
# Slopes are expressed as tanB = dz/dist
# +
import numpy as np
class bmPoint:
"""
Instantiate a three dimensional point object in regular coordinate space
p = bmPoint(x, y, [z])
Args:
x: x (Easting) coordinate [length]
y: y (Northing) coordinate [length]
z: z (Elevation) coordinate [length; optional: defaults to 0.0 if not specified]
Returns:
p: point of type bmPoint with x,y coordinates [length]
Attributes:
p.x: x (Easting) coordinate [length]
p.y: y (Northing) coordinate [length]
p.z: z (Elevation) coordinate [length; optional: defaults to 0.0 if not specified]
"""
def __init__(self, x, y, z=0.0):
self.x = x
self.y = y
self.z = z
def bmDist2D(p1,p2):
"""
Calculate distance between two bmPoint objects in the x, y plane
dist = bmDist2D(p1, p2)
Args:
p1: point of type bmPoint with x,y coordinates [length]
p2: point of type bmPoint with x,y coordinates [length]
Returns:
dist: Euclian distance between the points [length]
"""
dx = p2.x-p1.x
dy = p2.y-p1.y
return(np.sqrt(dx**2+dy**2))
def bmDist3D(p1, p2):
# Distance between two bmPoints in 3D space
dx = p2.x-p1.x
dy = p2.y-p1.y
dy = p2.z-p1.z
return(np.sqrt(dx**2+dy**2+dz**2))
def bmRAz(p1,p2):
# Range, bearing, and slope from bmPoint 1 to bmPoint 2
dx = p2.x-p1.x
dy = p2.y-p1.y
dz = p2.z-p1.z
az = np.nan
s = np.nan
r = np.sqrt(dx**2+dy**2)
if r >0.0:
az=np.degrees( np.arctan2(dx, dy) )
az = (az+360.)%360.
s = dz/r
return(r,az,s)
def bmPcoord(x, y):
"""
Convert x, y to polar coordinates r, az (geographic convention)
r,az = pcoord(x, y)
"""
r = sqrt( x**2 + y**2 )
az=degrees( arctan2(x, y) )
# az[where(az<0.)[0]] += 360.
az = (az+360.)%360.
return r, az
def bmXycoord(r, az):
"""
Convert r, az [degrees, geographic convention] to rectangular coordinates
x,y = xycoord(r, az)
"""
x = r * sin(radians(az))
y = r * cos(radians(az))
return x, y
# +
def main():
p = bmPoint(1.,2)
print(p.x,p.y,p.z)
pa = p
pb = bmPoint(0.0,0.0)
print(bmDist2D(pa,pb))
print(bmDist2D(pa,pb))
print(bmDist2D(pa,pb))
if __name__ == "__main__":
main()
# -
| basic_functions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import statsmodels.api as sm
import matplotlib.pyplot as plt
# %matplotlib inline
df=pd.read_csv('perrin-freres-monthly-champagne-.csv')
df.tail()
df.drop(106,axis=0,inplace=True)
df.tail()
df.drop(105,axis=0,inplace=True)
df.columns=['Month','Sales per month' ]
df.head()
df['Month']=pd.to_datetime(df['Month'])
df.head()
df.set_index('Month',inplace=True)
df.head()
df.plot()
model=sm.tsa.statespace.SARIMAX(df['Sales per month'],order=(1, 0, 0),seasonal_order=(1,1,1,12))
results=model.fit()
df['forecast']=results.predict(start=90,end=103,dynamic=True)
df[['Sales per month','forecast']].plot(figsize=(12,8))
from pandas.tseries.offsets import DateOffset
future_dates=[df.index[-1]+ DateOffset(months=x)for x in range(0,24)]
future_datest_df=pd.DataFrame(index=future_dates[1:],columns=df.columns)
future_datest_df
future_df=pd.concat([df,future_datest_df])
future_df['forecast'] = results.predict(start = 104, end = 120, dynamic= True)
future_df[['Sales per month', 'forecast']].plot(figsize=(12, 8))
| Arimax.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.6 64-bit (''general'': venv)'
# language: python
# name: python37664bitgeneralvenvfbd0a23e74cf4e778460f5ffc6761f39
# ---
# # Compare Odte with different estimators
# # Setup
# Uncomment the next cell if Odte is not already installed
#
# Google Colab setup
#
# #!pip install git+https://github.com/doctorado-ml/odte
import datetime, time
import numpy as np
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn import tree
from sklearn.metrics import classification_report, confusion_matrix, f1_score
from sklearn.ensemble import RandomForestClassifier, AdaBoostClassifier, GradientBoostingClassifier
from stree import Stree
from odte import Odte
import os
if not os.path.isfile('data/creditcard.csv'):
# !wget --no-check-certificate --content-disposition http://nube.jccm.es/index.php/s/Zs7SYtZQJ3RQ2H2/download
# !tar xzf creditcard.tgz
# # Tests
# + tags=[]
print(datetime.date.today(), time.strftime("%H:%M:%S"))
# -
# ## Load dataset and normalize values
# Load Dataset
df = pd.read_csv('data/creditcard.csv')
df.shape
random_state = 2020
# + tags=[]
print("Fraud: {0:.3f}% {1}".format(df.Class[df.Class == 1].count()*100/df.shape[0], df.Class[df.Class == 1].count()))
print("Valid: {0:.3f}% {1:,}".format(df.Class[df.Class == 0].count()*100/df.shape[0], df.Class[df.Class == 0].count()))
# -
# Normalize Amount
from sklearn.preprocessing import RobustScaler
values = RobustScaler().fit_transform(df.Amount.values.reshape(-1, 1))
df['Amount_Scaled'] = values
# + tags=[]
# Remove unneeded features
y = df.Class.values
X = df.drop(['Class', 'Time', 'Amount'], axis=1).values
print(f"X shape: {X.shape}\ny shape: {y.shape}")
# -
# ## Build the models
# Divide dataset
train_size = .7
Xtrain, Xtest, ytrain, ytest = train_test_split(X, y, train_size=train_size, shuffle=True, random_state=random_state, stratify=y)
# Linear Tree
linear_tree = tree.DecisionTreeClassifier(random_state=random_state)
# Random Forest
random_forest = RandomForestClassifier(random_state=random_state)
# Stree
stree = Stree(random_state=random_state, C=.01)
# AdaBoost
adaboost = AdaBoostClassifier(random_state=random_state)
# Gradient Boosting
gradient = GradientBoostingClassifier(random_state=random_state)
# Oblique Decision Tree Ensemble
odte = Odte(random_state=random_state, max_features="auto")
# ## Do the test
def try_model(name, model):
print(f"************************** {name} **********************")
now = time.time()
model.fit(Xtrain, ytrain)
spent = time.time() - now
print(f"Train Model {name} took: {spent:.4} seconds")
predict = model.predict(Xtrain)
predictt = model.predict(Xtest)
print(f"=========== {name} - Train {Xtrain.shape[0]:,} samples =============",)
print(classification_report(ytrain, predict, digits=6))
print(f"=========== {name} - Test {Xtest.shape[0]:,} samples =============")
print(classification_report(ytest, predictt, digits=6))
print("Confusion Matrix in Train")
print(confusion_matrix(ytrain, predict))
print("Confusion Matrix in Test")
print(confusion_matrix(ytest, predictt))
return f1_score(ytest, predictt), spent
# + tags=[]
# Train & Test models
models = {
'Linear Tree':linear_tree, 'Random Forest': random_forest, 'Stree (SVM Tree)': stree,
'AdaBoost model': adaboost, 'Odte model': odte #'Gradient Boost.': gradient
}
best_f1 = 0
outcomes = []
for name, model in models.items():
f1, time_spent = try_model(name, model)
outcomes.append((name, f1, time_spent))
if f1 > best_f1:
best_model = name
best_time = time_spent
best_f1 = f1
# + tags=[]
print("*"*110)
print(f"*The best f1 model is {best_model}, with a f1 score: {best_f1:.4} in {best_time:.6} seconds with {train_size:,} samples in train dataset")
print("*"*110)
for name, f1, time_spent in outcomes:
print(f"Model: {name}\t Time: {time_spent:6.2f} seconds\t f1: {f1:.4}")
# + active=""
# **************************************************************************************************************
# *The best f1 model is Random Forest, with a f1 score: 0.8815 in 152.54 seconds with 0.7 samples in train dataset
# **************************************************************************************************************
# Model: Linear Tree Time: 13.52 seconds f1: 0.7645
# Model: Random Forest Time: 152.54 seconds f1: 0.8815
# Model: Stree (SVM Tree) Time: 32.55 seconds f1: 0.8603
# Model: AdaBoost model Time: 47.34 seconds f1: 0.7509
# Model: Gradient Boost. Time: 244.12 seconds f1: 0.5259
# -
# ```
# ******************************************************************************************************************
# *The best f1 model is Random Forest, with a f1 score: 0.8815 in 218.966 seconds with 0.7 samples in train dataset
# ******************************************************************************************************************
# Model: Linear Tree Time: 23.05 seconds f1: 0.7645
# Model: Random Forest Time: 218.97 seconds f1: 0.8815
# Model: Stree (SVM Tree) Time: 49.45 seconds f1: 0.8467
# Model: AdaBoost model Time: 73.83 seconds f1: 0.7509
# Model: Gradient Boost. Time: 388.69 seconds f1: 0.5259
# Model: Neural Network Time: 25.47 seconds f1: 0.8328
# Model: Odte Time:2134.25 seconds f1: 0.8385
# ```
| notebooks/benchmark.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.2 64-bit (''d4ad_standardization'': pipenv)'
# name: python37264bitd4adstandardizationpipenvcac7d9f4a0864f29b6353caf0213501a
# ---
# + tags=[]
# Importing the libraries
import pandas as pd
import numpy as np
import random
import pickle
import re
import regex # for better, more capbale regex api
import os
import zipfile
import more_itertools
from itertools import chain
import datetime
import time
from statsmodels.stats.proportion import proportion_confint
# active labeler related
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.feature_extraction.text import TfidfTransformer
from sklearn.naive_bayes import ComplementNB # corrects for class imbalance, SGD is pretty good too
from sklearn.pipeline import Pipeline
from superintendent import ClassLabeller
from IPython.display import display, Markdown
pd.set_option('display.max_colwidth', None) # so we can peak at data and spot verify
pipeline = Pipeline([
('vect', CountVectorizer(analyzer='char', ngram_range=(1,2))),
('tfidf', TfidfTransformer()),
('clf', ComplementNB()),
])
print('done')
# + tags=[]
# Set up columns to keep, fields, locations for writing
rootpath = "/hdd/work/d4ad_standardization/"
processedpath = "D4AD_Standardization/data/processed/"
externalpath = "D4AD_Standardization/data/external/"
interimpath = "D4AD_Standardization/data/interim/"
content_is = "standardized_descriptions_and_degree_funding_type"
print('done')
# + tags=[]
filepath = "standardized_name_and_name1.csv" # builds off of notebook 5 work
columns = [
"STANDARDIZEDNAME_1",
"STANDARDIZEDNAME",
"DESCRIPTION",
"FEATURESDESCRIPTION",
"NAME_1",
"NAME",
"PREREQUISITES",
"STREET1",
"CITY",
"STATE",
"ZIP",
"WEBSITE",
"COUNTY",
"NONGOVAPPROVAL",
"STATECOMMENTS",
"CIPCODE",
"PROVIDERID",
"APPROVINGAGENCYID"
]
columns_to_save = ['STANDARDIZED_DESCRIPTION', 'STANDARDIZED_FEATURESDESCRIPTION'] + columns
SKIP_THIS = True # helps me be able to run all and not worry about pulling things
# I already know I have on disk
#df = pd.read_excel(rootpath + interimpath + filepath, usecols=columns)
df = pd.read_csv(rootpath + interimpath + filepath, usecols=columns)
print('done')
# +
pd.set_option('display.max_rows', False)
the_df = df #df.sample(n=10000, random_state=42)
# + tags=[]
# 2) Here we apply the abbreviation expansion to the
# description columns. This code is repeated from the 5.0 notebook and should be externalized into ./src somewhere
#
# We first construct the abbreviation mapper
#
# We also store off a copy of the df for manipulation
# this has older name fields, for informing on funding (WOIA) and degree type (?)
# as well as the standardized fields so taht we can remove the extranous content still in it
# Note: this is mixing responsibilites and should be seperated into a new notebook
label_mapper = pd.read_csv(
rootpath + externalpath + "label_mapper.csv"
)
draft_output = the_df[['DESCRIPTION', 'FEATURESDESCRIPTION',
'STANDARDIZEDNAME_1', 'STANDARDIZEDNAME',
'NAME_1', 'NAME']]
def make_term_grouped_regex(term="", right_regex="", left_regex=""):
mystr = left_regex + '(' +\
re.escape(term) +\
')' + right_regex
return mystr
def make_grouped_regexes(replacement, left_regex="", right_regex=""):
return (make_term_grouped_regex(left_regex=left_regex,
term=key,
right_regex=right_regex)\
for key in replacement.keys()
)
def construct_map(label_mapper=label_mapper):
return {
**dict(zip(label_mapper.abbreviation, label_mapper.expanded))
}
replacement_map = construct_map()
abbrevation_pattern =\
regex.compile(
"(?p)" +
"|".join( # match words at start of string
make_grouped_regexes(replacement_map, left_regex=r'^', right_regex=r'[\s:]')
) + "|" +\
"|".join( # match words surrounded by spaces
make_grouped_regexes(replacement_map, left_regex=r'\s', right_regex=r'\s')
) + "|" +\
"|".join( # match words that make up entire fields, e.g. 'Nursing'
make_grouped_regexes(replacement_map, left_regex=r'^', right_regex=r'$')
) + "|" +\
"|".join( # match words at end of string preceded by space or slash
make_grouped_regexes(replacement_map, left_regex=r'[\s/]', right_regex=r'$')
) + "|" +\
"|".join( # match words within string that follow a slash, end with a space or slash
make_grouped_regexes(replacement_map, left_regex=r'/', right_regex=r'[\s/]')
)
)
def multiple_mapper(string):
return abbrevation_pattern.sub(
lambda x: \
x.group().replace( # replace the found string
more_itertools.first_true(x.groups() # where the first matched group...
), replacement_map[more_itertools.first_true(x.groups())] # ... is replaced with the lookup
), string)
print('done1')
# + tags=[]
# ... with the abbreviation mapper in hand we now simply apply to both description columns
# it takes about 2.5 minutes each to run through all rows for both descriptions.
start = datetime.datetime.now()
draft_output['STANDARDIZED_DESCRIPTION'] =\
draft_output['DESCRIPTION'].dropna().map(multiple_mapper)
draft_output['STANDARDIZED_FEATURESDESCRIPTION'] =\
draft_output['FEATURESDESCRIPTION'].dropna().map(multiple_mapper)
# if not SKIP_THIS:
# draft_output['STANDARDIZED_DESCRIPTION'] =\
# draft_output['DESCRIPTION'].dropna().map(multiple_mapper)
# draft_output['STANDARDIZED_FEATURESDESCRIPTION'] =\
# draft_output['FEATURESDESCRIPTION'].dropna().map(multiple_mapper)
# else:
# joining_columns = ['NAME_1', 'NAME']
# interim_csv = "standardized_descriptions_and_degree_funding_type.csv"
# already_standardized_descriptions =\
# pd.read_csv(rootpath+interimpath+interim_csv,
# usecols=[
# 'STANDARDIZED_DESCRIPTION',
# 'STANDARDIZED_FEATURESDESCRIPTION'] + joining_columns)\
# .drop_duplicates(subset=joining_columns) # not sure how or why we have dupes
# # see: https://stackoverflow.com/questions/22720739/pandas-left-outer-join-results-in-table-larger-than-left-table
# read_in = draft_output.merge(
# already_standardized_descriptions,
# how='left',
# on=joining_columns,
# validate="m:1"
# )
# assert len(read_in) == len(draft_output), f"read in shape {len(read_in)} does not equal draft df {len(draft_output)}!"
# draft_output = read_in
end = datetime.datetime.now()
print(f"Done! That took {(end-start)} time")
# + tags=[]
# 3)
# Now we have to extract course funding type from the older
# columns.
wioa_like =\
regex.compile(
'''
(title\s+[IV1234]+\b\s*?) # WOIA has 4 titles of funding in law, at end of sentence or space
|(wioa){d<=1} # is called WOIA, WIA, allowed to miss a letter
''',
flags=regex.I|regex.VERBOSE)
name =\
draft_output['NAME'].dropna()\
.map(wioa_like.search)\
.dropna().index
name_1 =\
draft_output['NAME_1'].dropna()\
.map(wioa_like.search)\
.dropna().index
descriptions =\
draft_output['DESCRIPTION'].dropna()\
.map(wioa_like.search)\
.dropna().index
features_description =\
draft_output['FEATURESDESCRIPTION'].dropna()\
.map(wioa_like.search)\
.dropna().index
wioa_indices = name.union(name_1)\
.union(descriptions)\
.union(features_description)
draft_output['IS_WIOA'] = False
draft_output.loc[wioa_indices, 'IS_WIOA'] = True
print('done')
# + tags=[]
# ... Finally we extact the degree type from the older columns, repeating the
# procedure above but with slightly different regexes
as_like =\
regex.compile(
'''
[\b\s](A\.A\.S\.)[\b\s]
|[\b\s](A\.S\.)[\b\s]
|[\b\s](AS\sDe) # AS Degree
|[\b\s](AS\sSc) # AS Science
|[\b\s](AAS)[\b\s] # applied associates of science
''',
flags=regex.VERBOSE)
name =\
draft_output['NAME'].dropna()\
.map(as_like.search)\
.dropna().index
name_1 =\
draft_output['NAME_1'].dropna()\
.map(as_like.search)\
.dropna().index
descriptions =\
draft_output['DESCRIPTION'].dropna()\
.map(as_like.search)\
.dropna().index
features_description =\
draft_output['FEATURESDESCRIPTION'].dropna()\
.map(as_like.search)\
.dropna().index
as_indices = name.union(name_1)\
.union(descriptions)\
.union(features_description)
draft_output['Mentioned_Associates'] = False
draft_output.loc[as_indices, 'Mentioned_Associates'] = True
# +
# Now we go back for mentions of certificate and assign those
cert_like =\
regex.compile(
'''
(certification)
|(certificate)
|[\s\b](cert)[\s\b]
''',
flags=regex.I|regex.VERBOSE)
name =\
draft_output['NAME'].dropna()\
.map(cert_like.search)\
.dropna().index
name_1 =\
draft_output['NAME_1'].dropna()\
.map(cert_like.search)\
.dropna().index
descriptions =\
draft_output['DESCRIPTION'].dropna()\
.map(cert_like.search)\
.dropna().index
features_description =\
draft_output['FEATURESDESCRIPTION'].dropna()\
.map(cert_like.search)\
.dropna().index
cert_indices = name.union(name_1)\
.union(descriptions)\
.union(features_description)
draft_output['Mentioned_Certificate'] = False
draft_output.loc[cert_indices, 'Mentioned_Certificate'] = True
# +
# 4)
# Now we do some simple removals for known degree related mentions in the name fields
degree_cert_variants =\
["A.S.",
"AAS Degree",
"AAS -",
"A.S. Degree",
"AS Degree",
"Degree",
"degree",
"certificate",
"Certificate",
"Associate of Applied Science",
"-[\s\b]Associate",
"^\s*In\b"]
draft_output['CLEANED_STANDARDIZED_NAME_1'] =\
draft_output['STANDARDIZEDNAME_1'].replace(degree_cert_variants, "", regex=True)
# + tags=[]
# This is the evaluation part of the program and course name standardizations
# along with the provider name. My goal is to have 85%+ standardized, send out
# that 85% will come from the jefferey's interval
# Evaluation Rubric:
# A) Here we label clearly wrong snippets, anything that is marginal we mark as
# standardized for purposes of this evaluation because we want to err on the side
# of giving overly specific information, which includes odd info
# B) We also click through quickly, not overly dwelling one any one example, the
# goal here is to get the evaulation done quickly since it's so manual
# C) For now we ignore casingl there does need to be a camel casing applied to
# all caps
def stratified_sample(the_data, strata, size):
some_frac = size/len(the_data)
return \
the_data.groupby(
strata
).apply(
lambda g: g.sample(
n=size
#frac=1
)
)
if not SKIP_THIS:
key_factors_to_consider = ['IS_WOIA'] #, 'Mentioned_Certificate']
# We create a series of data to evaluate
columns_to_check = ['CLEANED_STANDARDIZED_NAME_1', 'IS_WOIA',
'Mentioned_Certificate', 'Mentioned_Associates',
'STANDARDIZED_DESCRIPTION', 'STANDARDIZED_FEATURESDESCRIPTION']
check_this_many = 10 #100 * len(columns_to_check) # we mark if ANY column are wrong
# the_data = draft_output.sample(check_this_many,random_state=42)\
# .loc[:, columns_to_check]
the_data = stratified_sample(draft_output, strata=key_factors_to_consider, size=check_this_many)
# we shuffle the data to elminate any bias across/within the columns when
# evaluting
print('done', f'The stratified validation data is {len(the_data)} long')
# +
markdown = []
def display_func(row):
"""
The display function gets passed your data - in the
case of a dataframe, it gets passed a row - and then
has to "display" your data in whatever way you want.
It doesn't need to return anything
"""
the_string =\
"**IS_WOIA:** " + str(row["IS_WOIA"]) +\
" **Cert:** " + str(row["Mentioned_Certificate"]) +\
" **Assoc:** " + str(row["Mentioned_Associates"]) +\
"\n\n**Provider:** " + str(row["STANDARDIZEDNAME"]) + "" +\
"\n\n**Course Name:** " + str(row["CLEANED_STANDARDIZED_NAME_1"]) + "" +\
"\n\n**Description:** " + str(row["STANDARDIZED_DESCRIPTION"]) + "" +\
"\n\n**Featured Description:** " + str(row["STANDARDIZED_FEATURESDESCRIPTION"]) + "" +\
"\n\n**(unstandardized):** [Name_1] " + str(row["NAME_1"]) + " [Name] " + str(row["NAME"])
markdown_string =\
Markdown(the_string)
display(
markdown_string
)
markdown.append(the_string)
def preprocessor(x, y):
# only take standardized column, leave everything else
return x, y
if not SKIP_THIS:
verification_widget = ClassLabeller(
features=the_data,
model=pipeline,
model_preprocess=preprocessor,
display_func=display_func,
options=['standardized', 'not standardized'],
acquisition_function='margin'
)
verification_widget
# + tags=[]
# for m in markdown:
# print(m)
# print('\n\n')
# -
#
# + tags=[]
# insert bionomial proprtion esimator here
def print_CI(labels, response_is_standardized = "standardized", method = "jeffreys"):
successful_count = sum(
response_is_standardized == label for label in labels
)
not_examined_count = sum(
None == label for label in labels
)
CI = proportion_confint(
count= successful_count,
nobs= len(labels) - not_examined_count,
alpha = 0.95,
method=method
)
print(f"{method} bionomial proportion is: [{CI[0]:.2f}, {CI[1]:.2f}]",
)
print(f"We examined {len(labels) - not_examined_count} labels, of which {successful_count} are correct. There are {len(labels)} labels.")
if not SKIP_THIS:
print_CI(labels=verification_widget.new_labels)
# + tags=[]
# 4)
# Now we write out the verfiied results
# ... finally we can write this out as our first complete lookup table
# for the NAME field
write_out = draft_output[
[
'IS_WIOA', 'Mentioned_Certificate', 'Mentioned_Associates',
'STANDARDIZED_DESCRIPTION', 'STANDARDIZED_FEATURESDESCRIPTION',
'CLEANED_STANDARDIZED_NAME_1', 'STANDARDIZEDNAME',
'STANDARDIZEDNAME_1', 'DESCRIPTION',
'FEATURESDESCRIPTION', 'NAME_1', 'NAME'
]
]
print(
"We're writing ...",
write_out.columns
)
# shuffe the rows to better remove temporal baises
write_out =\
write_out.sample(frac=1, random_state=42, axis=0).reset_index(drop=True)
write_out.to_csv(rootpath + interimpath + content_is + ".csv",
index = False,
chunksize = 10000)
write_out.to_excel(rootpath + processedpath + content_is + ".xls",
sheet_name="Standardized Descriptions",
index=False)
print('done')
| D4AD_Standardization/notebooks/6.0-kpr-Description-with_Funding_type_degree_type_columns.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# # Parsing GFF3 files
#
# Without specification, the default GFF3 parsing function will be used. This function will call `gffutils` and instruct it to build a new database in memory, with the flag `merge_strategy="create_unique"`. These can be adjusted by modfying the arguments to `GffutilsParseArgs`.
from inscripta.biocantor.io.gff3.parser import parse_standard_gff3
gff3 = "tests/data/INSC1006_chrI.gff3"
# GFF3 parsing requires that the file be passed directly instead as an open handle
annotation_records = list(parse_standard_gff3(gff3))
# After parsing, there will be one `AnnotationRecord` built for every sequence in the GFF3 file. This dataclass is a wrapper that contains two objects, a `SeqRecord` object and a `AnnotationCollectionModel`. Because this parser is not given any FASTA information, the `SeqRecord` is null. The `AnnotationCollectionModel` objects are `marshmallow_dataclass` objects, and so can be dumped to and loaded directly from JSON.
annotation_collection_model = parsed[0].annotation
annotation_collection_model.Schema().dump(annotation_collection_model)
# ## Converting models to BioCantor data structures
# After loading an `AnnotationCollectionModel`, this object can be directly converted in to an `AnnotationCollection` with sequence information.
#
# `AnnotationCollection` objects are the core data structure, and contain a set of genes and features as children.
annotation_collection = annotation_collection_model.to_annotation_collection()
# this example dataset has 4 genes and 0 features
for child in annotation_collection:
print(child)
gene1 = annotation_collection.genes[0]
tx1 = gene1.transcripts[0]
# convert mRNA coordinates to genomic coordinates
tx1.transcript_pos_to_sequence(0)
# NoncodingTranscriptError is raised when trying to convert CDS coordinates on a non-coding transcript
tx1.cds_pos_to_sequence(0)
# ## Primary transcripts
#
# It is often useful to have an understanding of what isoform of a gene is the 'most important'. An input dataset can provide this information based on the parser implementation used. If this information is not provided, then this value is inferred by the simple heuristic of:
#
# 1. Longest CDS isoform.
# 2. Longest isoform (if no coding isoforms).
gene1.get_primary_transcript() == tx1
# ## Incorporating sequence information
#
# The above parsing call treated the input GFF3 as a standard GFF3 without sequence. However, this file actually has a FASTA suffix, and so the sequence information can be loaded either from that file, or from a separate FASTA file.
#
# When parsers are used that return sequence information, the returned `ParsedAnnotationRecord` now has a non-null `seqrecord` member. This object can then be used to instantiate an `AnnotationCollection` with sequence information.
# +
from inscripta.biocantor.io.gff3.parser import parse_gff3_embedded_fasta
parsed_with_sequence = list(parse_gff3_embedded_fasta(gff3))
annotation_collection_with_sequence = parsed_with_sequence[0].to_annotation_collection()
annotation_collection_with_sequence.get_reference_sequence()
# -
for gene in annotation_collection_with_sequence:
for tx in gene.transcripts:
if tx.is_coding:
print(f"{tx.transcript_symbol}: mRNA: {tx.get_spliced_sequence()[:10]}... protein: {tx.get_protein_sequence()[:10]}...")
else:
print(f"{tx.transcript_symbol}: mRNA: {tx.get_spliced_sequence()[:10]}...")
# Sequence information can also come from a separate FASTA file. In this case, a different function call is used, and it gets provided the FASTA directly.
# +
from inscripta.biocantor.io.gff3.parser import parse_gff3_fasta
fasta = "tests/data/INSC1006_chrI.fasta"
parsed_with_sequence_from_fasta = list(parse_gff3_embedded_fasta(gff3))
annotation_collection_with_sequence_from_fasta = parsed_with_sequence_from_fasta[0].to_annotation_collection()
# -
annotation_collection_with_sequence_from_fasta.get_reference_sequence()
# ## Querying the collection
#
# `AnnotationCollections` have the ability to be subsetted. These range queries can be performed in two modes, controlled by the flag `completely_within`. When `completely_within = True`, the positions in the query are exact bounds. When `completely_within = False`, any constituent object that overlaps the range query will be retained.
#
# `start` and `end` are not required to be set, and are inferred to be `0` and `len(sequence)` respectively if not used.
# remove GI526_G0000001 by moving the start position to within its bounds, when strict boundaries are required
subset1 = annotation_collection.query_by_position(start=16175, completely_within=True)
print([x.gene_symbol for x in subset1])
# select BDH1 and BDH2
subset2 = annotation_collection.query_by_position(start=40000, end=42000, completely_within=False)
print([x.gene_symbol for x in subset2])
# ## Modifying the parser
#
# Each of the GFF3 wrapper parser functions, `parse_standard_gff3`, `parse_gff3_embedded_fasta`, and `parse_gff3_fasta`, all have arguments that allow you to modify how the GFF3 file is interpreted. These include:
#
# 1. `gffutil_parse_args=GffutilsParseArgs()`: This dataclass contains arguments that are passed to gffutils.
# 2. `parse_func`: The default implementation of this function can be written bespoke to your annotation data.
# 3. `gffutil_transform_func`: This function is passed straight to the same argument in `gffutils`.
# 4. `db_fn`: Change this value to retain the `gffutils` sqlite database on disk.
| docs/source/parsing_gff3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
Introduction
# ## Sorting Algorithms
#
# 1. Bubble Sort
#
# Bubble sort, sometimes referred to as sinking sort, is a simple sorting algorithm that repeatedly steps through the list, compares adjacent elements and swaps them if they are in the wrong order. The pass through the list is repeated until the list is sorted. The algorithm, which is a comparison sort, is named for the way smaller or larger elements "bubble" to the top of the list. Source: https://en.wikipedia.org/wiki/Bubble_sort
# Implementing Bubble Sort
# +
# code sourded from: https://runestone.academy/runestone/books/published/pythonds/SortSearch/TheBubbleSort.html
def bubbleSort(alist):
for passnum in range(len(alist)-1,0,-1):
for i in range(passnum):
if alist[i]>alist[i+1]:
temp = alist[i]
alist[i] = alist[i+1]
alist[i+1] = temp
alist = [54,26,93,17,77,31,44,55,20]
bubbleSort(alist)
print(alist)
# -
def bubble_sort(array):
n = len(array)
for i in range(n):
# Create a flag that will allow the function to
# terminate early if there's nothing left to sort
already_sorted = True
# Start looking at each item of the list one by one,
# comparing it with its adjacent value. With each
# iteration, the portion of the array that you look at
# shrinks because the remaining items have already been
# sorted.
for j in range(n - i - 1):
if array[j] > array[j + 1]:
# If the item you're looking at is greater than its
# adjacent value, then swap them
array[j], array[j + 1] = array[j + 1], array[j]
# Since you had to swap two elements,
# set the `already_sorted` flag to `False` so the
# algorithm doesn't finish prematurely
already_sorted = False
# If there were no swaps during the last iteration,
# the array is already sorted, and you can terminate
if already_sorted:
break
return array
if __name__ == "__main__":
# Generate an array of `ARRAY_LENGTH` items consisting
# of random integer values between 0 and 999
array = [randint(0, 1000) for i in range(ARRAY_LENGTH)]
# Call the function using the name of the sorting algorithm
# and the array you just created
run_sorting_algorithm(algorithm="bubble_sort", array=array)
# +
# importing the random numbers
from random import *
# code sourced from http://interactivepython.org/runestone/static/pythonds/SortSearch/TheBubbleSort.html
def bubbleSort(alist):
for passnum in range(len(alist)-1,0,-1):
for i in range(passnum):
if alist[i]>alist[i+1]:
temp = alist[i]
alist[i] = alist[i+1]
alist[i+1] = temp
import time
global bubble_avglist
bubble_avglist = []
num_runs = 10
results = []
# benchmark bubble function
def benchmark_bubble():# import time module
for r in range(num_runs):
# start timer
start_time = time.time()
######## bubblesort
bubbleSort(alist)
end_time = time.time()
time_elapsed= end_time - start_time
results.append(time_elapsed)
b = sum(results)
average = (b/num_runs)
# round to 3 decimals
average = round(average, 3)
bubble_avglist.append(average)
for r in range(num_runs):
# start timer
start_time = time.time()
######## bubblesort
bubbleSort(alist1)
end_time = time.time()
time_elapsed= end_time - start_time
results.append(time_elapsed)
b = sum(results)
average = (b/num_runs)
# round to 3 decimals
average = round(average, 3)
bubble_avglist.append(average)
for r in range(num_runs):
# start timer
start_time = time.time()
######## bubblesort
bubbleSort(alist2)
end_time = time.time()
time_elapsed= end_time - start_time
results.append(time_elapsed)
b = sum(results)
average = (b/num_runs)
# round to 3 decimals
average = round(average, 3)
bubble_avglist.append(average)
for r in range(num_runs):
# start timer
start_time = time.time()
######## bubblesort
bubbleSort(alist3)
end_time = time.time()
time_elapsed= end_time - start_time
results.append(time_elapsed)
b = sum(results)
average = (b/num_runs)
# round to 3 decimals
average = round(average, 3)
bubble_avglist.append(average)
for r in range(num_runs):
# start timer
start_time = time.time()
######## bubblesort
bubbleSort(alist4)
end_time = time.time()
time_elapsed= end_time - start_time
results.append(time_elapsed)
b = sum(results)
average = (b/num_runs)
# round to 3 decimals
average = round(average, 3)
bubble_avglist.append(average)
for r in range(num_runs):
# start timer
start_time = time.time()
######## bubblesort
bubbleSort(alist5)
end_time = time.time()
time_elapsed= end_time - start_time
results.append(time_elapsed)
b = sum(results)
average = (b/num_runs)
# round to 3 decimals
average = round(average, 3)
bubble_avglist.append(average)
for r in range(num_runs):
# start timer
start_time = time.time()
######## bubblesort
bubbleSort(alist6)
end_time = time.time()
time_elapsed= end_time - start_time
results.append(time_elapsed)
b = sum(results)
average = (b/num_runs)
# round to 3 decimals
average = round(average, 3)
bubble_avglist.append(average)
for r in range(num_runs):
# start timer
start_time = time.time()
######## bubblesort
bubbleSort(alist7)
end_time = time.time()
time_elapsed= end_time - start_time
results.append(time_elapsed)
b = sum(results)
average = (b/num_runs)
# round to 3 decimals
average = round(average, 3)
bubble_avglist.append(average)
for r in range(num_runs):
# start timer
start_time = time.time()
######## bubblesort
bubbleSort(alist8)
end_time = time.time()
time_elapsed= end_time - start_time
results.append(time_elapsed)
b = sum(results)
average = (b/num_runs)
# round to 3 decimals
average = round(average, 3)
bubble_avglist.append(average)
for r in range(num_runs):
# start timer
start_time = time.time()
######## bubblesort
bubbleSort(alist9)
end_time = time.time()
time_elapsed= end_time - start_time
results.append(time_elapsed)
b = sum(results)
average = (b/num_runs)
# round to 3 decimals
average = round(average, 3)
bubble_avglist.append(average)
for r in range(num_runs):
# start timer
start_time = time.time()
######## bubblesort
bubbleSort(alist10)
end_time = time.time()
time_elapsed= end_time - start_time
results.append(time_elapsed)
b = sum(results)
average = (b/num_runs)
# round to 3 decimals
average = round(average, 3)
bubble_avglist.append(average)
for r in range(num_runs):
# start timer
start_time = time.time()
######## bubblesort
bubbleSort(alist11)
end_time = time.time()
time_elapsed= end_time - start_time
results.append(time_elapsed)
b = sum(results)
average = (b/num_runs)
# round to 3 decimals
average = round(average, 3)
bubble_avglist.append(average)
for r in range(num_runs):
# start timer
start_time = time.time()
######## bubblesort
bubbleSort(alist12)
end_time = time.time()
time_elapsed= end_time - start_time
results.append(time_elapsed)
b = sum(results)
average = (b/num_runs)
# round to 3 decimals
average = round(average, 3)
bubble_avglist.append(average)
for r in range(num_runs):
# start timer
start_time = time.time()
######## bubblesort
bubbleSort(alist13)
end_time = time.time()
time_elapsed= end_time - start_time
results.append(time_elapsed)
b = sum(results)
average = (b/num_runs)
# round to 3 decimals
average = round(average, 3)
bubble_avglist.append(average)
print(bubble_avglist)
#return bubble_avglist
benchmark_bubble()
# -
| Benchmark Sorting Algorithms.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Allsky CRE vs. cloud type as vertically stacked plots
# We plot the allsky CRE vs. cloud type. "Allsky" includes weighting by cloud cover.
# ## Load Standard and User Libraries
# +
# %matplotlib inline
import os, sys, copy, glob
import numpy as np
import datetime
import seaborn as sns
import pylab as pl
# -
pl.rcParams['figure.figsize'] = (16.0, 8.0)
pl.rcParams['font.size'] = 24.0
pl.rcParams['lines.linewidth'] = 3
import xarray as xr
# +
import nawdex_analysis.analysis.ave_cre
import nawdex_analysis.io.collector
import nawdex_analysis.plot.stacked_analysis_plots
reload( nawdex_analysis.plot.stacked_analysis_plots)
from nawdex_analysis.plot.stacked_analysis_plots import vert_stacked_exp_plot, get_exp_kws, get_plotting_order
# -
# ## Read Radiation Fluxes
# This is a slight jump forward in time. Now, average radiation fluxes are available for different CT categories.
rset= nawdex_analysis.io.collector.get_radflux4set( 'all', method = 'strict' )
#rset= nawdex_analysis.io.collector.get_radflux4set( 'all', method = 'all' )
#rset = nawdex_analysis.io.collector.get_radflux4set( 2 )
# ## Convert Flux to CRE (Scaling !!!!)
nawdex_analysis.analysis.ave_cre.radflux2cre( rset, scaling = True, new_factor = 0.88, old_factor = 0.9 )
rset
# ## Adding fractional and vary low clouds
cset = rset.sel(ct = 'very low') + rset.sel(ct = 'fractional')
cset = cset.expand_dims('ct')
cset['ct'] = ['very low / fractional']
cset = xr.concat( [cset, rset], dim = 'ct' )
# ## Rename CT labels
ctnames = cset.ct.data
ctnames[8] = 'semi. moderately thick'
ctnames[4] = 'mid-level'
# ## all NaNs should be zero of the fraction-weighted radflux contribution
# The problems with the NaNs is that the average values and std of shortwave fluxes change strongly if some NaNs mask out a certain time at the day.
mask = cset.notnull()
cset = cset.where( mask, 0)
# ## Calculate Mean and STD
# +
catlist = list( cset.ct.data )
remove_list = ['clear_ocean', 'semi. above', 'very low', 'fractional']
for rm_cat in remove_list:
catlist.remove(rm_cat)
idlist = list( cset.idname.data )
idlist.remove( 'msevi-not_scaled' )
dset = cset.sel(ct = catlist, idname = idlist)
daily_mean = dset.groupby('time.day').mean('time')
# dvar = daily_mean.quantile([0.25, 0.5, 0.75], dim = 'day').diff('quantile')
dvar = daily_mean.std('day')
mvar = dset.mean('time')
# -
dvar_q = ( daily_mean.quantile(0.84, 'day') - daily_mean.quantile(0.16, 'day') ) /2.
# ## Calculate Standard Error of Mean
nday = daily_mean.dims['day']
var_sem = ( daily_mean.quantile(0.84, 'day') - daily_mean.quantile(0.16, 'day') ) / np.sqrt( nday )
# ## Plotting
from nawdex_analysis.plot.legend import plegend
mvar.ct.data = ['very low /\nfractional', 'low', 'mid-level', 'high opaque',
'very high\nopaque', 'semi. thin', 'semi.\nmoderately thick',
'semi. thick']
dvar.ct.data = mvar.ct.data
dvar_q.ct.data = mvar.ct.data
var_sem.ct.data = mvar.ct.data
# +
grey_fontsize = 21
fac = 0.6
fig, ax = pl.subplots( ncols = 2, figsize = (30*fac, 35*fac), )
pl.sca(ax[0])
vert_stacked_exp_plot( mvar, 'scre_contrib' , var_err = var_sem, doffset =0.06)
vert_stacked_exp_plot( mvar, 'lcre_contrib' , var_err = var_sem, doffset =0.06)
xlim = pl.xlim(-15, 15)
pl.title('(c) Short- & Longwave')
pl.sca(ax[1])
vert_stacked_exp_plot( mvar, 'net_cre_contrib' , var_err = var_sem, doffset =0.06)
pl.title(' (d) Net')
xlim = pl.xlim()
#sns.despine(ax = ax[0])
#sns.despine(left = True, ax = ax[1])
#ax[1].set_yticks([])
sns.despine(left = True, right = True, ax = ax[0])
sns.despine(left = True, right = False, ax = ax[1])
ax[0].set_yticks([])
### Extra Text Labels
### ==================
pl.sca( ax[0] )
pl.text( -12, -0.8, 'Shortwave',
color = 'gray', ha = 'center', va = 'center', alpha = 0.7,
fontweight = 'bold', fontsize = grey_fontsize)
pl.text( 12, -0.8, 'Longwave',
color = 'gray', ha = 'center', va = 'center', alpha = 0.7,
fontweight = 'bold', fontsize = grey_fontsize)
pl.sca( ax[1] )
pl.text( xlim[0] / 2, -0.8, 'Cooling',
color = 'gray', ha = 'center', va = 'center', alpha = 0.7,
fontweight = 'bold', fontsize = grey_fontsize)
#pl.text( xlim[1] * 0.8, -0.8, 'Warming',
# color = 'gray', ha = 'center', va = 'center', alpha = 0.7,
# fontweight = 'bold', fontsize = 'large')
#plegend()
for a in ax:
a.axvline(0, color = 'gray', lw = 5, alpha = 0.3)
a.set_xlabel('CRE ($\mathrm{W\, m^{-2}}$)')
a.set_ylim(-1,None)
pl.suptitle('Allsky CRE', fontsize = 'x-large', fontweight = 'bold')
pl.subplots_adjust( left = 0.3, right = 0.7, top = 0.91 )
pl.savefig( '../pics/cre_allsky-vs-ct.png', dpi = 300,)
| nbooks/08-allsky_CRE-per-ct_plots.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Semi-Monocoque Theory
from pint import UnitRegistry
import sympy
import networkx as nx
import numpy as np
import matplotlib.pyplot as plt
import sys
# %matplotlib inline
from IPython.display import display
# Import **Section** class, which contains all calculations
from Section import Section
# Initialization of **sympy** symbolic tool and **pint** for dimension analysis (not really implemented rn as not directly compatible with sympy)
ureg = UnitRegistry()
sympy.init_printing()
# Define **sympy** parameters used for geometric description of sections
A, A0, t, t0, a, b, h, L = sympy.symbols('A A_0 t t_0 a b h L', positive=True)
# We also define numerical values for each **symbol** in order to plot scaled section and perform calculations
values = [(A, 150 * ureg.millimeter**2),(A0, 250 * ureg.millimeter**2),(a, 80 * ureg.millimeter), \
(b, 20 * ureg.millimeter),(h, 35 * ureg.millimeter),(L, 2000 * ureg.millimeter)]
datav = [(v[0],v[1].magnitude) for v in values]
# # Triangular section
# Define graph describing the section:
#
# 1) **stringers** are **nodes** with parameters:
# - **x** coordinate
# - **y** coordinate
# - **Area**
#
# 2) **panels** are **oriented edges** with parameters:
# - **thickness**
# - **lenght** which is automatically calculated
# +
stringers = {1:[(sympy.Integer(0),h),A],
2:[(sympy.Integer(0),sympy.Integer(0)),A],
3:[(a,sympy.Integer(0)),A]}
panels = {(1,2):t,
(2,3):t,
(3,1):t}
# -
# Define section and perform first calculations
S1 = Section(stringers, panels)
S1.cycles
# ## Plot of **S1** section in original reference frame
# Define a dictionary of coordinates used by **Networkx** to plot section as a Directed graph.
# Note that arrows are actually just thicker stubs
start_pos={ii: [float(S1.g.node[ii]['ip'][i].subs(datav)) for i in range(2)] for ii in S1.g.nodes() }
plt.figure(figsize=(12,8),dpi=300)
nx.draw(S1.g,with_labels=True, arrows= True, pos=start_pos)
plt.arrow(0,0,20,0)
plt.arrow(0,0,0,20)
#plt.text(0,0, 'CG', fontsize=24)
plt.axis('equal')
plt.title("Section in starting reference Frame",fontsize=16);
# Expression of **Inertial properties** wrt Center of Gravity in with original rotation
S1.Ixx0, S1.Iyy0, S1.Ixy0, S1.α0
# ## Plot of **S1** section in inertial reference Frame
# Section is plotted wrt **center of gravity** and rotated (if necessary) so that *x* and *y* are principal axes.
# **Center of Gravity** and **Shear Center** are drawn
positions={ii: [float(S1.g.node[ii]['pos'][i].subs(datav)) for i in range(2)] for ii in S1.g.nodes() }
# +
x_ct, y_ct = S1.ct.subs(datav)
plt.figure(figsize=(12,8),dpi=300)
nx.draw(S1.g,with_labels=True, pos=positions)
plt.plot([0],[0],'o',ms=12,label='CG')
plt.plot([x_ct],[y_ct],'^',ms=12, label='SC')
#plt.text(0,0, 'CG', fontsize=24)
#plt.text(x_ct,y_ct, 'SC', fontsize=24)
plt.legend(loc='lower right', shadow=True)
plt.axis('equal')
plt.title("Section in pricipal reference Frame",fontsize=16);
# -
# Expression of **inertial properties** in *principal reference frame*
sympy.simplify(S1.Ixx), sympy.simplify(S1.Iyy), sympy.simplify(S1.Ixy), sympy.simplify(S1.θ)
# ## **Shear center** expression
#
# Expressions can be messy, so we evaluate them to numerical values
sympy.N(S1.ct.subs(datav))
# ## Analisys of Loads
# We define some symbols
Tx, Ty, Nz, Mx, My, Mz, F, ry, ry, mz = sympy.symbols('T_x T_y N_z M_x M_y M_z F r_y r_x m_z')
S1.set_loads(_Tx=0, _Ty=Ty, _Nz=0, _Mx=Mx, _My=0, _Mz=0)
#S1.compute_stringer_actions()
#S1.compute_panel_fluxes();
# **Axial Loads**
# +
#S1.N
# -
# **Panel Fluxes**
# +
#S1.q
# -
# **Example 2**: _twisting moment_ in **z** direction
S1.set_loads(_Tx=0, _Ty=0, _Nz=0, _Mx=0, _My=0, _Mz=Mz)
S1.compute_stringer_actions()
S1.compute_panel_fluxes();
# **Axial Loads**
S1.N
# **Panel Fluxes**
# evaluated to numerical values
{k:sympy.N(S1.q[k].subs(datav)) for k in S1.q }
# ## Torsional moment of Inertia
S1.compute_Jt()
sympy.N(S1.Jt.subs(datav))
| 02_Triangular_Section.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## _*Qiskit Chemistry, H2O ground state computation*_
#
# This notebook demonstrates how to use Qiskit Chemistry to compute the ground state energy of a water (H2O) molecule using VQE and UCCSD.
#
# While the molecule has been input below to the driver in xyz format, the Z-matrix format is also support. H2O in Z-matrix format would look like this
# ```
# H; O 1 1.08; H 2 1.08 1 104.5
# ```
# and is convenient when the goal is to change bond angle, or plot the energy changing distance(s) while preserving the angle.
#
# This notebook has been written to use the PYSCF chemistry driver.
# +
from qiskit import BasicAer
from qiskit.aqua import QuantumInstance
from qiskit.aqua.algorithms.adaptive import VQE
from qiskit.aqua.algorithms.classical import ExactEigensolver
from qiskit.aqua.components.optimizers import SLSQP
from qiskit.chemistry.drivers import PySCFDriver, UnitsType
from qiskit.chemistry.core import Hamiltonian, TransformationType, QubitMappingType
from qiskit.chemistry.components.variational_forms import UCCSD
from qiskit.chemistry.components.initial_states import HartreeFock
# -
# First we create and run a driver to produce our molecule object. The molecule object holds data from the drivers in a common way so it can then be used independently of which specific driver created it.
#
# And let's print some of fields it has. You can refer to qiskit.aqua.qmolecule.py for more information or look at the API documentation.
# +
driver = PySCFDriver(atom='O 0.0 0.0 0.0; H 0.757 0.586 0.0; H -0.757 0.586 0.0',
unit=UnitsType.ANGSTROM, charge=0, spin=0, basis='sto3g')
molecule = driver.run()
print('Hartree-Fock energy: {}'.format(molecule.hf_energy))
print('Nuclear repulsion energy: {}'.format(molecule.nuclear_repulsion_energy))
print('Number of molecular orbitals: {}'.format(molecule.num_orbitals))
print('Number of alpha electrons: {}'.format(molecule.num_alpha))
print('Number of beta electrons: {}'.format(molecule.num_beta))
# -
# We now need to create a qubit operator as input to compute the ground state energy. The Hamilitonian object can be used. This wraps a `FermionicOperator` class, which can be used directly but entails more steps. Other tutorials here show FermionicOperator being used.
#
# The Hamiltonian class not only gives us a qubit operator for the main Hamiltonian but also auxilliary operators including dipole operators and others to measure spin and num particles. The algorithm, if it supports aux_ops, which ExactEignesolver and VQE both do, will evaluate these at the ground state where the minimum energy is found.
# +
core = Hamiltonian(transformation=TransformationType.FULL, qubit_mapping=QubitMappingType.PARITY,
two_qubit_reduction=True, freeze_core=True)
qubit_op, aux_ops = core.run(molecule)
print(qubit_op)
# -
# We now pass these to the ExactEigensolver and run it to produce a result. This result will include the computed electronic part of the ground state energy. We can pass this result back to the Hamiltonian object from above and it will combine it with values it stored such as the frozen core energy to form a complete result for the molecule. As can be seen this matches the result from above.
#
# Note: the num particles printed here is that which is observed from the spin operator that is in the aux_ops. It says 8 which matches what we expect; the molecule has 10 (5 alpha and 5 beta) but the operator was left with 8 after we took away 2 from freezing the core. The molecule has a core_orbitals property which lists the orbitals comprising the core ones that can be frozen so we can easily figure how many electrons that is (2 per orbital in that list).
ee = ExactEigensolver(qubit_op, aux_operators=aux_ops)
algo_result = ee.run()
result = core.process_algorithm_result(algo_result)
for line in result[0]:
print(line)
# #### Using VQE
#
# Here we will start with the qubit operator that we computed above. We need to setup an optimizer, variational form and initial state for use with VQE.
#
# The variational form and UCCSD are a little more complex since they need information about numbers of orbitals and numbers of electrons, as well as what qubit mapping etc was used for the qubit operator. However we have some help from the Hamiltonian class that we can use.
#
# Note: If you use FermionicOperator directly to make a qubit operator then you need to keep track of electrons removed etc. The molecule object from the driver has the original values but if you freeze out orbitals then the electrons remaining in the operator is what is required.
# +
init_state = HartreeFock(num_qubits=qubit_op.num_qubits,
num_orbitals=core._molecule_info['num_orbitals'],
num_particles=core._molecule_info['num_particles'],
qubit_mapping=core._qubit_mapping,
two_qubit_reduction=core._two_qubit_reduction)
var_form = UCCSD(num_qubits=qubit_op.num_qubits,
depth=1,
num_orbitals=core._molecule_info['num_orbitals'],
num_particles=core._molecule_info['num_particles'],
qubit_mapping=core._qubit_mapping,
two_qubit_reduction=core._two_qubit_reduction,
initial_state=init_state)
optimizer = SLSQP(maxiter=2500)
# setup backend on which we will run
backend = BasicAer.get_backend('statevector_simulator')
quantum_instance = QuantumInstance(backend=backend)
vqe = VQE(qubit_op, var_form, optimizer)
algo_result = vqe.run(quantum_instance)
lines, result = core.process_algorithm_result(algo_result)
print('Ground state energy: {}'.format(result['energy']))
for line in lines:
print(line)
# -
# Internally the core, when processing the algorithm result, stores the result dictionary from the algorithm under the `algorithm_retvals` key. We used this above to get the eval count, and since we process the result the same way here, using the core, we can do this here too. But here we have direct access to the algorithm result since we ran it. Hence we can access the count directly from the above algo_result. To show these are the same they are both printed below.
# +
print('Actual VQE evaluations taken: {}'.format(result['algorithm_retvals']['eval_count']))
print('Actual VQE evaluations taken: {}'.format(algo_result['eval_count']))
# -
# #### Z-matrix format
#
# Z-matrix was mentioned in the introduction. Lets show it in use in a quick final example here. We'll use ExactEigensolver as the goal here is just to show the technique. We will keep the bond angle between the Hydrogen atoms and Oxygen constant while varying the interatomic distance of one the Hydrogen atoms. This is simple to do in Z-matrix format, though can of course be done using xyz format but that needs more work to compute the coordinates each time.
# +
import numpy as np
import pylab
h2o = 'H; O 1 1.08; H 2 {} 1 104.5'
distances = [x * 0.01 + 1.00 for x in range(17)]
energies = np.empty(len(distances))
for i, distance in enumerate(distances):
driver = PySCFDriver(h2o.format(distance), basis='sto3g')
qmolecule = driver.run()
operator = Hamiltonian(freeze_core=True)
qubit_op, aux_ops = operator.run(qmolecule)
result = ExactEigensolver(qubit_op).run()
lines, result = operator.process_algorithm_result(result)
energies[i] = result['energy']
# -
pylab.plot(distances, energies)
pylab.xlabel('Distance of Hydrogen atom')
pylab.ylabel('Energy')
pylab.title('H2O molecule, one H atom distance varied');
| chemistry/h2o.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
from sklearn.model_selection import KFold, StratifiedKFold
import pandas as pd
from matplotlib import pyplot as plt
from sklearn.neural_network import MLPClassifier, MLPRegressor
from sklearn.metrics import roc_auc_score
from sklearn.preprocessing import StandardScaler, MinMaxScaler
from sklearn.metrics import accuracy_score, mean_absolute_error
from sklearn.preprocessing import LabelEncoder
import pickle
import librosa
# %matplotlib inline
# # Read data
train_old = pd.read_csv("../data/train_mfcc.csv")
train_old.head()
df = pd.read_csv("../data/train_new_feat.csv")
df['genre'] = train_old[' genre']
df.head()
# # Data cleaning
#Drop useless features
useless = [ ' mother tongue', ' liked', ' disliked', ' amazement',' mood',' age', ' gender',' solemnity', ' nostalgia']
df = df.drop(useless, 1)
df = pd.get_dummies(df, columns=['genre'])
df.head()
LABELS = [' tenderness', ' calmness', ' power',
' joyful_activation', ' tension', ' sadness',
"genre_classical", "genre_electronic", "genre_pop", "genre_rock"
]
#Get list of base Features
FEATURES = df.columns.drop(['track id'] + LABELS)
print(f"base features: {FEATURES.tolist()}")
# drop duplicates to avoid ovefitting
Xtrain = df[FEATURES.tolist() + ['track id']].drop_duplicates()
# # Feature engineering
# As previously observed when performing the EDA, all the subjects do not share the same feelings for the same song and most of the subjects did not feel any emotion.
# Thus I decide to proceed as follows:
# For a given song, if more than 30% of the subjects indicate felt the same emotion, then
# +
ytrain = df.groupby('track id')[LABELS].mean()
ytrain = ytrain.reset_index()
for l in LABELS:
ytrain[l] = ytrain[l].apply(lambda x: 1 if x > 0.3 else 0, 1)
train = Xtrain.merge(ytrain, on='track id', how='left')
# -
# ## Outlier treatment
# +
from pyod.models.iforest import IForest
from pyod.models.knn import KNN
# predict and treat outliers for a given feature
def predict_and_treat_outlier(feat,db):
"""
This function treats outliers for a given feature
Parameters:
db (DataFrame): the dataframe containing the outliers to be cleaned
feat (string): the given feature
Returns:
db (DataFrame): the dataframe with cleaned outliers
"""
X = db[[feat]]
clf = IForest() #KNN()
clf.fit(X)
db['is_outlier'] = clf.predict(X)
db_cleaned = db.copy()
mask = db_cleaned['is_outlier'] == 1
db_cleaned.loc[mask, feat] = db_cleaned[feat].median()
return db_cleaned.drop(columns=['is_outlier'])
# -
# predict and treat age outliers
train = predict_and_treat_outlier(' age',train)
train.head()
# predict audio features outliers
def predict_audio_feat_outlier(db):
"""
This function globally detects outliers
Parameters:
db (DataFrame): the dataframe containing the values to be cleaned
Returns:
db_tr (DataFrame): the dataframe with detected outliers
"""
features = ['chroma_sftf', 'rolloff', 'zero_crossing_rate', 'rmse', 'flux', 'contrast', 'flatness',
'sample_silence', 'mfcc_0', 'mfcc_1','mfcc_2', 'mfcc_3', 'mfcc_4', 'mfcc_5', 'mfcc_6',
'mfcc_7', 'mfcc_8', 'mfcc_9', 'mfcc_10','mfcc_11','mfcc_12', 'mfcc_13', 'mfcc_14',
'mfcc_15', 'mfcc_16', 'mfcc_17', 'mfcc_18', 'mfcc_19', 'tempo']
db_tr = db.copy()
clf = IForest()
clf.fit(db_tr[features])
db_tr['is_outlier'] = clf.predict(db_tr[features])
return db_tr
#treat audio features outliers
def treat_audio_feat_outlier(db):
"""
This function globally treats outliers
Parameters:
db (DataFrame): the dataframe containing the outliers to be cleaned
Returns:
new_db (DataFrame): the dataframe with cleaned outliers
"""
features = ['chroma_sftf', 'rolloff', 'zero_crossing_rate', 'rmse', 'flux', 'contrast', 'flatness',
'sample_silence', 'mfcc_0', 'mfcc_1','mfcc_2', 'mfcc_3', 'mfcc_4', 'mfcc_5', 'mfcc_6',
'mfcc_7', 'mfcc_8', 'mfcc_9', 'mfcc_10','mfcc_11','mfcc_12', 'mfcc_13', 'mfcc_14',
'mfcc_15', 'mfcc_16', 'mfcc_17', 'mfcc_18', 'mfcc_19', 'tempo']
mask = db['is_outlier'] == 1
for f in features:
db.loc[mask, f] = db[f].median()
return db.drop(columns=['is_outlier'])
# predict audio features outliers
train = predict_audio_feat_outlier(train)
# treat audio features outliers
train = treat_audio_feat_outlier(train)
train.head()
# # Model training : MLP model
# Data normalization
scaler = MinMaxScaler()
train[FEATURES] = scaler.fit_transform(train[FEATURES])
# ## Model Validation
# Model Validation
epochs = 5
kf = KFold(n_splits=epochs, shuffle=True, random_state=1997) # 30, n_split 3
X_train = train[FEATURES]
y_train = train[LABELS]
y_oof = np.zeros([X_train.shape[0], len(LABELS)])
i = 0
for tr_idx, val_idx in kf.split(X_train, y_train):
X_tr, X_vl = X_train.iloc[tr_idx, :], X_train.iloc[val_idx, :]
y_tr, y_vl = y_train.iloc[tr_idx, :], y_train.iloc[val_idx, :]
model = MLPClassifier([300], random_state=47, activation='logistic', max_iter=5000)
model.fit(X_tr, y_tr)
y_pred = model.predict_proba(X_vl)
y_oof[val_idx, :] = y_pred
i += 1
acc = roc_auc_score(y_vl, y_pred, multi_class='ovr')
print(f"Fold #{i} AUC : {round(acc, 2)}")
metric = roc_auc_score(y_train, y_oof, multi_class='ovr')
print(f"Full AUC : {round(metric, 2)}")
# ## Save model and scaler
# +
import pickle
filename = '../model_saved/emo_genre_clf_model.sav'
pickle.dump(model, open(filename, 'wb'))
filename = '../model_saved/Scaler_Extractor.sav'
pickle.dump(scaler, open(filename, 'wb'))
| notebook/Music_Genre_Emotion_Classification_Baseline_DNN .ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# coding: utf-8
import numpy as np
from simple_convnet import SimpleConvNet
network = SimpleConvNet(input_dim=(1,10, 10),
conv_param = {'filter_num':10, 'filter_size':3, 'pad':0, 'stride':1},
hidden_size=10, output_size=10, weight_init_std=0.01)
X = np.random.rand(100).reshape((1, 1, 10, 10))
T = np.array([1]).reshape((1,1))
grad_num = network.numerical_gradient(X, T)
grad = network.gradient(X, T)
for key, val in grad_num.items():
print(key, np.abs(grad_num[key] - grad[key]).mean())
| ch07/gradient_check.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Hy
# language: hy
# name: hy
# ---
# # Magics
# [IPython magics](http://ipython.org/ipython-doc/dev/interactive/tutorial.html) do... interesting things. Listing them is beyond the scope of this document, but here are some examples, as well as [limitations](#Limitations).
# ## Cell magics
# ### ...that take Python
# For example, `%%timeit` will display how long a cell takes to run, running an appropriately large number of samples.
%%timeit
(+ 1 1)
# ### ...that don't
# `%%html` will just jam some html out to the frontend. To indicate that it shouldn't be compiled as hy, throw in some more magic with another `%`:
%%%html
<ul>
<li><input type="radio" name="magic" checked="true"/> magic</li>
<li><input type="radio" name="magic"/> <a href="http://www.catb.org/jargon/html/magic-story.html">more magic</a></li>
</ul>
# ## Line Magics
# `!` will run a shell command, and show the output.
!ls
# ## Limitations
# Magic arguments aren't supported yet. This means a lot of great stuff won't work, `%%file`, `%%run`.
print?
| notebooks/Magics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
from os.path import join
from pathlib import Path
import torch
from tqdm import tqdm
import random
import pandas as pd
import sys
sys.path.append(os.path.abspath(Path('..')))
from notebooks.profiles import MahalanobisProfile
from notebooks.feature_extractors import POSPCAExtractor, HeuristicExtractor
# -
bawe_group_dir = Path('../data/preprocess/bawe-group')
# # Progress Report 1
#
# This is a summary of all the work that has been done over winter break/the first few weeks of the semester.
#
# ## Current Progress
#
# We currently use profile-based methods to determine whether a new text belongs to a given author. Given a series of texts for an author, we compute a profile for that author, and then make a decision about whether a new text is from that author based on the distance from the new text to the profile. The current profile implementation is based on finding the mean feature vector given the different feature vectors for a text, and then the mahalanobis distance between the mean and a feature vector for the new text is computed. We then find the probability that the new text belongs to the profile, and output that. In the below tests, a threshold of 90% was used to flag essays as either belonging to the same author or a different author. We currently have 3 different feature extractions methods being worked on:
#
# 1. PCA based on POS bigram counts (good accuracy, currently best model)
# 2. Heuristics gathering (chance accuracy, but may be able to boost performance of bigram counts)
# * Will likely have better accuracy when we use all heuristics and gather more
# 3. LSTM POS encoder (WIP)
#
# ### Next Steps
#
# 1. Interpretability issues
# * We would like to have some kind of interface to explain to the user why an essay is being flagged.
# * Use interpretable model, but if probability is too close to threshold, use high performance model and prompt for review?
# * Use high performance model, but display most relevant heuristics to aid in diagnosis
# * Correlation does not equal causation?
# * If nothing else, we can heavily encourage the user to review any essays that are flagged and make a decision for themselves.
# 2. Are there any other feature extraction techniques we should try?
# 3. Adversarial models?
# * Train model to turn text from a different author into text that is accepted by the profile
# * May aid in assessing whether the profiles are resilient to patch writing
# * Concern: model may find a way to perfectly capture the style of an author and reform the text to that style, this new text *should* be accepted
#
# ### Precision, Recall, and Accuracy on the BAWE data
#
# We used the British Academic Written English Corpus (BAWE) to test the precision, recall, and accuracy of our profilers. Five authors are randomly selected from the set, and then they are compared with 20 random different authors. The first essay of an author is used to "profile" them, and then their essays along with the essays from the different authors are tested against this profile. The profile should return a high probability for essays that belong to the profile and a low probability for essays that don't. For this demo, a cutoff of 90% is used.
# +
def test_profile(profile, cutoff):
ids = os.listdir(bawe_group_dir)
other_ids = ids.copy()
random.shuffle(ids)
random.shuffle(other_ids)
irrelevant, false_positives, relevant, true_positives = 0, 0, 0, 0
# For performance reasons, only choose 5 authors
for id in ids[:10]:
profile.reset()
texts = file_texts(id)
first_text = texts[0]
other_texts = texts[1:]
# This author's first essay is used to profile that author
profile.feed(first_text)
# The rest of their essays are tested against this profile
other_scores = torch.tensor([profile.score(text) for text in other_texts])
irrelevant += len(texts)
false_positives += sum(other_scores < cutoff)
# Compare this author to 20 random different authors
new_relevant, new_true_positives = grade_others(profile, other_ids[:30], id, cutoff)
relevant += new_relevant
true_positives += new_true_positives
# Precision is relevant selections out of all selections
precision = true_positives / (true_positives + false_positives)
# Recall is all relevant selections out of all relevant items
recall = true_positives / relevant
false_negatives = relevant - true_positives
# False positives and false negatives sum to the error count, and
# irrelevant and relevant items sum to the whole set
error_rate = false_negatives + false_positives / (irrelevant + relevant)
return float(precision), float(recall), float(error_rate)
def file_texts(id):
filenames = os.listdir(join(bawe_group_dir, id))
texts = []
for filename in filenames:
with open(join(bawe_group_dir, f'{id}/{filename}'), 'r') as f:
texts.append(f.read())
return texts
def grade_others(profile, ids, id, cutoff):
rest_ids = [other_id for other_id in ids if other_id != id]
relevant, true_positives = 0, 0
for other_id in tqdm(rest_ids):
texts = file_texts(other_id)
scores = torch.tensor([profile.score(text) for text in texts])
true_positives += sum(scores < cutoff)
relevant += len(texts)
return relevant, true_positives
# +
pospca_extractor = POSPCAExtractor(4, 10)
pospca_profile = MahalanobisProfile(pospca_extractor)
pospca_precision, pospca_recall, pospca_error = test_profile(pospca_profile, 0.50)
# +
heuristics_extractor = HeuristicExtractor(4)
heuristics_profile = MahalanobisProfile(heuristics_extractor)
heuristics_precision, heuristics_recall, heuristics_error = test_profile(heuristics_profile, 0.90)
# +
metric_data = [
[pospca_precision, pospca_recall, pospca_error],
[heuristics_precision, heuristics_recall, heuristics_error]
]
pd.DataFrame(metric_data, columns=['Precision', 'Recall', 'Error'], index=['POSPCA', 'Heuristics'])
| notebooks/deliver/bawe_profile_tests.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Example usage of the Yin-Yang dataset
# +
import torch
import numpy as np
import matplotlib.pyplot as plt
from dataset import YinYangDataset
from torch.utils.data import DataLoader
# %matplotlib inline
# -
# ### Setup datasets (training, validation and test set)
dataset_train = YinYangDataset(size=5000, seed=42)
dataset_validation = YinYangDataset(size=1000, seed=41)
dataset_test = YinYangDataset(size=1000, seed=40)
# ### Setup PyTorch dataloaders
# +
batchsize_train = 20
batchsize_eval = len(dataset_test)
train_loader = DataLoader(dataset_train, batch_size=batchsize_train, shuffle=True)
val_loader = DataLoader(dataset_validation, batch_size=batchsize_eval, shuffle=True)
test_loader = DataLoader(dataset_test, batch_size=batchsize_eval, shuffle=False)
# -
# ### Plot data
fig, axes = plt.subplots(ncols=3, sharey=True, figsize=(15, 8))
titles = ['Training set', 'Validation set', 'Test set']
for i, loader in enumerate([train_loader, val_loader, test_loader]):
axes[i].set_title(titles[i])
axes[i].set_aspect('equal', adjustable='box')
xs = []
ys = []
cs = []
for batch, batch_labels in loader:
for j, item in enumerate(batch):
x1, y1, x2, y2 = item
c = batch_labels[j]
xs.append(x1)
ys.append(y1)
cs.append(c)
xs = np.array(xs)
ys = np.array(ys)
cs = np.array(cs)
axes[i].scatter(xs[cs == 0], ys[cs == 0], color='C0', edgecolor='k', alpha=0.7)
axes[i].scatter(xs[cs == 1], ys[cs == 1], color='C1', edgecolor='k', alpha=0.7)
axes[i].scatter(xs[cs == 2], ys[cs == 2], color='C2', edgecolor='k', alpha=0.7)
axes[i].set_xlabel('x1')
if i == 0:
axes[i].set_ylabel('y1')
# ### Setup ANN
class Net(torch.nn.Module):
def __init__(self, network_layout):
super(Net, self).__init__()
self.n_inputs = network_layout['n_inputs']
self.n_layers = network_layout['n_layers']
self.layer_sizes = network_layout['layer_sizes']
self.layers = torch.nn.ModuleList()
layer = torch.nn.Linear(self.n_inputs, self.layer_sizes[0], bias=True)
self.layers.append(layer)
for i in range(self.n_layers-1):
layer = torch.nn.Linear(self.layer_sizes[i], self.layer_sizes[i+1], bias=True)
self.layers.append(layer)
return
def forward(self, x):
x_hidden = []
for i in range(self.n_layers):
x = self.layers[i](x)
if not i == (self.n_layers-1):
relu = torch.nn.ReLU()
x = relu(x)
x_hidden.append(x)
return x
# +
torch.manual_seed(12345)
# ANN with one hidden layer (with 120 neurons)
network_layout = {
'n_inputs': 4,
'n_layers': 2,
'layer_sizes': [120, 3],
}
net = Net(network_layout)
# Linear classifier for reference
shallow_network_layout = {
'n_inputs': 4,
'n_layers': 1,
'layer_sizes': [3],
}
linear_classifier = Net(shallow_network_layout)
# -
# ### Train ANN
# +
# used to determine validation accuracy after each epoch in training
def validation_step(net, criterion, loader):
with torch.no_grad():
num_correct = 0
num_shown = 0
for j, data in enumerate(loader):
inputs, labels = data
# need to convert to float32 because data is in float64
inputs = inputs.float()
outputs = net(inputs)
winner = outputs.argmax(1)
num_correct += len(outputs[winner == labels])
num_shown += len(labels)
accuracy = float(num_correct) / num_shown
return accuracy
# set training parameters
n_epochs = 300
learning_rate = 0.001
val_accuracies = []
train_accuracies = []
# setup loss and optimizer
criterion = torch.nn.CrossEntropyLoss()
optimizer = torch.optim.Adam(net.parameters(), lr=learning_rate)
# train for n_epochs
for epoch in range(n_epochs):
val_acc = validation_step(net, criterion, val_loader)
if epoch % 25 == 0:
print('Validation accuracy after {0} epochs: {1}'.format(epoch, val_acc))
val_accuracies.append(val_acc)
num_correct = 0
num_shown = 0
for j, data in enumerate(train_loader):
inputs, labels = data
# need to convert to float32 because data is in float64
inputs = inputs.float()
# zero the parameter gradients
optimizer.zero_grad()
# forward pass
outputs = net(inputs)
winner = outputs.argmax(1)
num_correct += len(outputs[winner == labels])
num_shown += len(labels)
loss = criterion(outputs, labels)
loss.backward()
optimizer.step()
accuracy = float(num_correct) / num_shown
train_accuracies.append(accuracy)
# after training evaluate on test set
test_acc = validation_step(net, criterion, test_loader)
print('#############################')
print('Final test accuracy:', test_acc)
print('#############################')
# -
# ### Plot training results
plt.figure(figsize=(10,8))
plt.plot(train_accuracies, label='train acc')
plt.plot(val_accuracies, label='val acc')
plt.axhline(test_acc, ls='--', color='grey', label='test acc')
plt.xlabel('epochs')
plt.ylabel('accuracy')
plt.ylim(0.3, 1.05)
plt.legend()
# ### Train Linear classifier as reference
# +
val_accuracies = []
train_accuracies = []
# setup loss and optimizer
criterion = torch.nn.CrossEntropyLoss()
optimizer = torch.optim.Adam(linear_classifier.parameters(), lr=learning_rate)
# train for n_epochs
for epoch in range(n_epochs):
val_acc = validation_step(linear_classifier, criterion, val_loader)
if epoch % 25 == 0:
print('Validation accuracy of linear classifier after {0} epochs: {1}'.format(epoch, val_acc))
val_accuracies.append(val_acc)
num_correct = 0
num_shown = 0
for j, data in enumerate(train_loader):
inputs, labels = data
# need to convert to float32 because data is in float64
inputs = inputs.float()
# zero the parameter gradients
optimizer.zero_grad()
# forward pass
outputs = linear_classifier(inputs)
winner = outputs.argmax(1)
num_correct += len(outputs[winner == labels])
num_shown += len(labels)
loss = criterion(outputs, labels)
loss.backward()
optimizer.step()
accuracy = float(num_correct) / num_shown
train_accuracies.append(accuracy)
# after training evaluate on test set
test_acc = validation_step(linear_classifier, criterion, test_loader)
print('#############################')
print('Final test accuracy linear classifier:', test_acc)
print('#############################')
# -
plt.figure(figsize=(10,8))
plt.plot(train_accuracies, label='train acc (lin classifier)')
plt.plot(val_accuracies, label='val acc (lin classifier)')
plt.axhline(test_acc, ls='--', color='grey', label='test acc (lin classifier)')
plt.xlabel('epochs')
plt.ylabel('accuracy')
plt.ylim(0.3, 1.05)
plt.legend()
| example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
class Solution(object):
def guessNumber(self,n):
"""
:type n: int
:rtype: int
"""
if n == 1:
return 1
left = 0
right = n + 1
mid = int(right/2)
while left < right:
ret = mid
if ret == 0:
return mid
elif ret > 0:
left = mid
mid = int((left + right + 1)/2)
else:
right = mid
mid = int((left + right )/2)
return left
# Runtime: 8 ms, faster than 99.06% of Python online submissions for Guess Number Higher or Lower.
# Memory Usage: 11.8 MB, less than 31.35% of Python online submissions for Guess Number Higher or Lower.
#
| LeetCode374.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Data Analysis using Datalab and BigQuery
# +
query="""
SELECT
departure_delay,
COUNT(1) AS num_flights,
APPROX_QUANTILES(arrival_delay, 10) AS arrival_delay_deciles
FROM
`bigquery-samples.airline_ontime_data.flights`
GROUP BY
departure_delay
HAVING
num_flights > 100
ORDER BY
departure_delay ASC
"""
import google.datalab.bigquery as bq
df = bq.Query(query).execute().result().to_dataframe()
df.head()
# -
import pandas as pd
percentiles = df['arrival_delay_deciles'].apply(pd.Series)
percentiles = percentiles.rename(columns = lambda x : str(x*10) + "%")
df = pd.concat([df['departure_delay'], percentiles], axis=1)
df.head()
without_extremes = df.drop(['0%', '100%'], 1)
without_extremes.plot(x='departure_delay', xlim=(-30,50), ylim=(-50,50));
# ## Challenge Exercise
#
# Your favorite college basketball team is playing at home and trailing by 3 points with 4 minutes left to go. Using this public dataset of [NCAA basketball play-by-data](https://bigquery.cloud.google.com/table/bigquery-public-data:ncaa_basketball), calculate the probability that your team will come from behind to win the game.
# <p>
# Hint (highlight to view)
# <p style='color:white'>
# You will need to find games where period=2, game_clock = 4, and (away_pts - home_pts) = 3. Then, you will need to find the fraction of such games that end with home_pts > away_pts. </p>
# <p>
# If you got this easily, then for a greater challenge, repeat this exercise, but plot a graph of come-from-behind odds by time-remaining and score-margin. See https://medium.com/analyzing-ncaa-college-basketball-with-gcp/so-youre-telling-me-there-s-a-chance-e4ba0ad7f542 for inspiration.
# Copyright 2018 Google Inc. Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0 Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
| deepdive/01_googleml/data_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# <center><h1> bsolar vs bolasso </h1></center>
# * <font size="4.5"> In this file we set the DGP identical to the one in simulation 1. </font>
# * <font size="4.5"> $p/n$ changes stagewise from $100/100$ to $100/150$ and $100/200$. </font>
# ## #1: import all modules
#
# * <font size="4.5"> "pickle" is used to save all computation results into ".p" files, which can be loaded later. </font>
#
# * <font size="4.5"> For simplicity and elegancy, all relevant functions and classes are coded in "simul_plot_parallel.py". </font>
# +
# %reset -f
from simul_plot_parallel import simul_plot
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import pickle
import os
import errno
# -
# ---
#
# ## #2: define all functions
#
# ## #2(a): computation and visualization
def func_simul(sample_size, n_dim, n_info, num_rep, step_size, rnd_seed, repro):
#set random seed
np.random.seed(rnd_seed)
print("compute the simulation with sample size "+str(sample_size)+" and number of variables "+str(n_dim))
trial = simul_plot(sample_size, n_dim, n_info, num_rep, step_size, rnd_seed)
pkl_file = "./numerical_result/bsolar_simul_n_" + str(sample_size) + "_p_" + str(n_dim) + "_thre_1_bsolar_10.p"
if repro == True:
bsolar3S_Q_opt_c_stack, bsolar3H_Q_opt_c_stack, bsolar10S_Q_opt_c_stack, bsolar10H_Q_opt_c_stack, bsolar5S_Q_opt_c_stack, bsolar5H_Q_opt_c_stack, bolassoS_Q_opt_c_stack, bolassoH_Q_opt_c_stack = trial.simul_func()
#save all the computation result into Pickle files
#create the subdirectory if not existing
if not os.path.exists(os.path.dirname(pkl_file)):
try:
os.makedirs(os.path.dirname(pkl_file))
# Guard against race condition
except OSError as exc:
if exc.errno != errno.EEXIST:
raise
with open(pkl_file, "wb") as f:
pickle.dump( bsolar3S_Q_opt_c_stack , f)
pickle.dump( bsolar3H_Q_opt_c_stack , f)
pickle.dump( bsolar10S_Q_opt_c_stack , f)
pickle.dump( bsolar10H_Q_opt_c_stack , f)
pickle.dump( bsolar5S_Q_opt_c_stack , f)
pickle.dump( bsolar5H_Q_opt_c_stack , f)
pickle.dump( bolassoS_Q_opt_c_stack , f)
pickle.dump( bolassoH_Q_opt_c_stack , f)
else:
with open(pkl_file, "rb") as f:
bsolar3S_Q_opt_c_stack = pickle.load( f )
bsolar3H_Q_opt_c_stack = pickle.load( f )
bsolar10S_Q_opt_c_stack = pickle.load( f )
bsolar10H_Q_opt_c_stack = pickle.load( f )
bsolar5S_Q_opt_c_stack = pickle.load( f )
bsolar5H_Q_opt_c_stack = pickle.load( f )
bolassoS_Q_opt_c_stack = pickle.load( f )
bolassoH_Q_opt_c_stack = pickle.load( f )
#compute the number of selected variable of both bsolar and blasso
## set the container
bsolar3S_len_array = np.empty(num_rep)
bsolar3H_len_array = np.empty(num_rep)
bsolar10S_len_array = np.empty(num_rep)
bsolar10H_len_array = np.empty(num_rep)
bsolar5S_len_array = np.empty(num_rep)
bsolar5H_len_array = np.empty(num_rep)
bolassoS_len_array = np.empty(num_rep)
bolassoH_len_array = np.empty(num_rep)
##count the number
for i in range(num_rep):
bsolar3S_len_array[i] = len(bsolar3S_Q_opt_c_stack[i])
bsolar3H_len_array[i] = len(bsolar3H_Q_opt_c_stack[i])
bsolar10S_len_array[i] = len(bsolar10S_Q_opt_c_stack[i])
bsolar10H_len_array[i] = len(bsolar10H_Q_opt_c_stack[i])
bsolar5S_len_array[i] = len(bsolar5S_Q_opt_c_stack[i])
bsolar5H_len_array[i] = len(bsolar5H_Q_opt_c_stack[i])
bolassoS_len_array[i] = len(bolassoS_Q_opt_c_stack[i])
bolassoH_len_array[i] = len(bolassoH_Q_opt_c_stack[i])
#compute the marginal probability of selecting each informative variable
## set the container
bsolar3S_var_array = np.empty([5])
bsolar3H_var_array = np.empty([5])
bsolar10S_var_array = np.empty([5])
bsolar10H_var_array = np.empty([5])
bsolar5S_var_array = np.empty([5])
bsolar5H_var_array = np.empty([5])
bolassoS_var_array = np.empty([5])
bolassoH_var_array = np.empty([5])
##concatenate results
bsolar3S_vari_appe_stack = np.concatenate(bsolar3S_Q_opt_c_stack ,0)
bsolar3H_vari_appe_stack = np.concatenate(bsolar3H_Q_opt_c_stack ,0)
bsolar10S_vari_appe_stack = np.concatenate(bsolar10S_Q_opt_c_stack,0)
bsolar10H_vari_appe_stack = np.concatenate(bsolar10H_Q_opt_c_stack,0)
bsolar5S_vari_appe_stack = np.concatenate(bsolar5S_Q_opt_c_stack,0)
bsolar5H_vari_appe_stack = np.concatenate(bsolar5H_Q_opt_c_stack,0)
bolassoS_vari_appe_stack = np.concatenate(bolassoS_Q_opt_c_stack,0)
bolassoH_vari_appe_stack = np.concatenate(bolassoH_Q_opt_c_stack,0)
##count the number
for i in range(5):
bsolar3S_var_array[i] = (bsolar3S_vari_appe_stack == i).sum()/num_rep
bsolar3H_var_array[i] = (bsolar3H_vari_appe_stack == i).sum()/num_rep
bsolar10S_var_array[i] = (bsolar10S_vari_appe_stack == i).sum()/num_rep
bsolar10H_var_array[i] = (bsolar10H_vari_appe_stack == i).sum()/num_rep
bsolar5S_var_array[i] = (bsolar5S_vari_appe_stack == i).sum()/num_rep
bsolar5H_var_array[i] = (bsolar5H_vari_appe_stack == i).sum()/num_rep
bolassoS_var_array[i] = (bolassoS_vari_appe_stack == i).sum()/num_rep
bolassoH_var_array[i] = (bolassoH_vari_appe_stack == i).sum()/num_rep
#sparsity table
mean_col = [np.mean(bsolar3S_len_array), np.mean(bsolar3H_len_array),
np.mean(bsolar10S_len_array), np.mean(bsolar10H_len_array),
np.mean(bsolar5S_len_array), np.mean(bsolar5H_len_array),
np.mean(bolassoS_len_array), np.mean(bolassoH_len_array)]
median_col = [np.median(bsolar3S_len_array), np.median(bsolar3H_len_array),
np.median(bsolar10S_len_array), np.median(bsolar10H_len_array),
np.median(bsolar5S_len_array), np.median(bsolar5H_len_array),
np.median(bolassoS_len_array), np.median(bolassoH_len_array)]
df1 = pd.concat([pd.DataFrame({'algo':['bsolar-3S','bsolar-3H',
'bsolar-10S', 'bsolar-10H',
'bsolar-5S', 'bsolar-5H',
'bolasso-S', 'bolasso-H']}),
pd.DataFrame({'Number of selected variables (mean)': mean_col}),
pd.DataFrame({'Number of selected variables (median)': median_col})],
axis=1, join='inner')
#accuracy table
prob_0 = [bsolar3S_var_array[0] , bsolar3H_var_array[0],
bsolar10S_var_array[0], bsolar10H_var_array[0],
bsolar5S_var_array[0], bsolar5H_var_array[0],
bolassoS_var_array[0] , bolassoH_var_array[0]]
prob_1 = [bsolar3S_var_array[1] , bsolar3H_var_array[1],
bsolar10S_var_array[1], bsolar10H_var_array[1],
bsolar5S_var_array[1], bsolar5H_var_array[1],
bolassoS_var_array[1] , bolassoH_var_array[1]]
prob_2 = [bsolar3S_var_array[2] , bsolar3H_var_array[2],
bsolar10S_var_array[2], bsolar10H_var_array[2],
bsolar5S_var_array[2], bsolar5H_var_array[2],
bolassoS_var_array[2] , bolassoH_var_array[2]]
prob_3 = [bsolar3S_var_array[3] , bsolar3H_var_array[3],
bsolar10S_var_array[3], bsolar10H_var_array[3],
bsolar5S_var_array[3], bsolar5H_var_array[3],
bolassoS_var_array[3] , bolassoH_var_array[3]]
prob_4 = [bsolar3S_var_array[4] , bsolar3H_var_array[4],
bsolar10S_var_array[4], bsolar10H_var_array[4],
bsolar5S_var_array[4], bsolar5H_var_array[4],
bolassoS_var_array[4] , bolassoH_var_array[4]]
df2 = pd.concat([pd.DataFrame({'algo':['bsolar-3S','bsolar-3H',
'bsolar-10S', 'bsolar-10H',
'bsolar-5S', 'bsolar-5H',
'bolasso-S', 'bolasso-H']}),
pd.DataFrame({'Pr(select X0)': prob_0}),
pd.DataFrame({'Pr(select X1)': prob_1}),
pd.DataFrame({'Pr(select X2)': prob_2}),
pd.DataFrame({'Pr(select X3)': prob_3}),
pd.DataFrame({'Pr(select X4)': prob_4})],
axis=1, join='inner')
return df1.round(1), df2.round(1)
# ---
#
# ## #3(a): define inputs values
#
# | <font size="4.5"> variable name </font> | <font size="4.5"> meaning </font> |
# |-|-|
# | <font size="4.5"> sample_size </font> | <font size="4.5"> the sample size $n$ in the paper; </font>|
# | <font size="4.5"> n_dim </font> | <font size="4.5"> the number of variables (informative + redundant) in $X$, $p$ in the paper; </font>|
# | <font size="4.5"> n_info </font> | <font size="4.5"> the number of informative variables in $X$; </font>|
# | <font size="4.5"> num_rep </font> | <font size="4.5"> the total repetition number of this simulation; </font>|
# | <font size="4.5"> step_size </font> | <font size="4.5"> the step size for tuning $c$; </font>|
# | <font size="4.5"> rnd_seed </font> | <font size="4.5"> the random seed value; </font>|
#
# ## #3(b): define DGP
#
# * <font size="4.5"> the population regression equation is $$Y = 2\cdot \mathbf{x}_0 + 3\cdot \mathbf{x}_1 + 4\cdot \mathbf{x}_2 + 5\cdot \mathbf{x}_3 + 6\cdot \mathbf{x}_4 + u,$$
# * <font size="4.5"> To change the simulation settings, simply change the input values. If you change *n_info* you will adjust the DGP as follows: </font>
# * <font size="4.5"> If $i > \mbox{n_info} - 1$ and $i \in \left[ 0, 1, 2, \ldots, p-1 \right]$, $\beta_i = 0$ in population;</font>
# * <font size="4.5"> If $i \leqslant \mbox{n_info} - 1$ and $i \in \left[ 0, 1, 2, \ldots, p-1 \right]$, $\beta_i = i + 2$ in population</font>
# +
n_dim = 100
n_info = 5
step_size = -0.01
num_rep = 100
rnd_seed = 0
sample_size_0 = 100
sample_size_1 = 150
sample_size_2 = 200
# -
# ---
#
# ## #4: result comparison
# * <font size="5"> if you only want to see our raw result, set *repro=False* </font>
# * <font size="5"> for replication, set *repro=True* </font>
repro = True
# * <font size="4.5"> Numpy, Sklearn and Python are actively updated. If you use different version, replication results may be slightly different from the paper (see Read_me_first.pdf for detail).</font>
# * <font size="4.5"> To rerun this part, first delete all .p files in the "raw_result" folder to avoid possible bug. </font>
# + [markdown] jp-MarkdownHeadingCollapsed=true tags=[]
# ---
# ## **Read this before you move on** : the runtime length issue of bolasso
#
# ### Beware that bolasso computation could take very long time. On a Thinkpad T480 laptop with i5-8500u CPU and 8G Ram, bolasso takes around 60 mins for one repetition at $p/n=1200/600$ (200 hours for 200 repetitions in total).
#
# ### Please use a proper desktop with a 8-core CPU if you want to replicate the bolasso result as quick as possible
#
# ---
# -
# # scenario #1: $p/n \rightarrow 0$
#
# ## $n = 100, p =100$
df1, df2 = func_simul(sample_size_0, n_dim, n_info, num_rep, step_size, rnd_seed, repro)
df1
df2
# ---
#
# ## $n = 150, p =100$
df1, df2 = func_simul(sample_size_1, n_dim, n_info, num_rep, step_size, rnd_seed, repro)
df1
df2
# ---
#
# ## $n = 200, p =100$
df1, df2 = func_simul(sample_size_2, n_dim, n_info, num_rep, step_size, rnd_seed, repro)
df1
df2
# + [markdown] tags=[]
# ---
#
# # scenario #2: $p/n \rightarrow 1$
# -
sample_size_0 = 100 ; n_dim_0 = 150
sample_size_1 = 150 ; n_dim_1 = 200
sample_size_2 = 200 ; n_dim_2 = 250
# + [markdown] tags=[]
# ## $n = 100, p =150$
# -
df1, df2 = func_simul(sample_size_0, n_dim, n_info, num_rep, step_size, rnd_seed, repro)
df1
df2
# + [markdown] tags=[]
# ---
# ## $n = 150, p =200$
# -
df1, df2 = func_simul(sample_size_1, n_dim, n_info, num_rep, step_size, rnd_seed, repro)
df1
df2
# ---
# ## $n = 200, p =250$
df1, df2 = func_simul(sample_size_2, n_dim, n_info, num_rep, step_size, rnd_seed, repro)
df1
df2
# ---
#
# # scenario #3: $\log(p)/n \rightarrow 0$
# +
sample_size_0 = 200 ; n_dim_0 = 400
sample_size_1 = 400 ; n_dim_1 = 800
sample_size_2 = 600 ; n_dim_2 = 1200
num_rep = 100
# -
# ## $n = 200, p =400$
df1, df2 = func_simul(sample_size_0, n_dim_0, n_info, num_rep, step_size, rnd_seed, repro)
df1
df2
# + [markdown] tags=[]
# ---
#
# ## $n = 400, p = 800$
# -
df1, df2 = func_simul(sample_size_1, n_dim_1, n_info, num_rep, step_size, rnd_seed, repro)
df1
df2
# ---
#
# ## $n = 600, p = 1200$
df1, df2 = func_simul(sample_size_2, n_dim_2, n_info, num_rep, step_size, rnd_seed, repro)
df1
df2
# !rm -rf sparsity_accuracy_bolasso_bsolar.html
# !jupyter nbconvert --to html sparsity_accuracy_bolasso_bsolar.ipynb
| simul_bolasso_bsolar/sparsity_accuracy/sparsity_accuracy_bolasso_bsolar.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %load_ext autoreload
# %autoreload 2
import sys
sys.path.append('../../')
import os
os.environ["CUDA_VISIBLE_DEVICES"] = '3'
# +
import torch
import numpy as np
import random
import pickle
import matplotlib.pyplot as plt
def seed_everything():
torch.manual_seed(0)
random.seed(0)
np.random.seed(0)
# -
# ## Load dataset
# +
from personalized_nlp.datasets.emotions.emotions import EmotionsDataModule
from personalized_nlp.datasets.cawi1.meanings import MeaningsDataModule
import pandas as pd
# -
# ## Import datasets
regression = True
embeddings_type = 'xlmr'
x = EmotionsDataModule(embeddings_type=embeddings_type, normalize=regression,
batch_size=1000)
x.prepare_data()
x.setup()
text_stds = x.annotations.groupby('text_id').std().iloc[:, 1:]
(text_stds < 0.1).sum(axis=0)
# +
regression = True
embeddings_type = 'xlmr'
cawi1_data_module = MeaningsDataModule(embeddings_type=embeddings_type, normalize=regression,
batch_size=1000)
cawi1_data_module.prepare_data()
cawi1_data_module.setup()
cawi2_data_module = EmotionsDataModule(embeddings_type=embeddings_type, normalize=regression,
batch_size=1000)
cawi2_data_module.prepare_data()
cawi2_data_module.setup()
# +
cawi1_annotators = sorted(cawi1_data_module.annotator_id_idx_dict.keys())
cawi2_annotators = sorted(cawi2_data_module.annotator_id_idx_dict.keys())
len(set(cawi1_annotators).intersection(cawi2_annotators))
# -
# ## Load embeddings
cawi1_embeddings_dict = pickle.load(open('/mnt/big_one/persemo/mgruza/humor-personalization/cawi1_embeddings.p', 'rb'))
cawi2_embeddings_dict = pickle.load(open('/mnt/big_one/persemo/mgruza/humor-personalization/cawi2_embeddings.p', 'rb'))
# +
cawi1_embeddings = []
cawi2_embeddings = []
for annotator_id in cawi1_embeddings_dict.keys():
if annotator_id in cawi2_embeddings_dict:
cawi1_embeddings.append(cawi1_embeddings_dict[annotator_id])
cawi2_embeddings.append(cawi2_embeddings_dict[annotator_id])
cawi1_embeddings = np.vstack(cawi1_embeddings)
cawi2_embeddings = np.vstack(cawi2_embeddings)
# -
# ## Compute distances
from scipy.spatial import distance_matrix
from scipy.stats import pearsonr
# +
cawi1_distances = distance_matrix(cawi1_embeddings, cawi1_embeddings).flatten()
cawi2_distances = distance_matrix(cawi2_embeddings, cawi2_embeddings).flatten()
cawi1_distances = (cawi1_distances - cawi1_distances.mean()) / cawi1_distances.std()
cawi2_distances = (cawi2_distances - cawi2_distances.mean()) / cawi2_distances.std()
pearsonr(cawi1_distances, cawi2_distances)
# -
distances_df = pd.DataFrame({'cawi1_distances': cawi1_distances, 'cawi2_distances': cawi2_distances})
# +
import seaborn as sns
plt.figure(figsize=(15, 15))
sns.displot(data=distances_df, x='cawi1_distances', y='cawi2_distances', height=6)
# -
# ## CCA
# +
from sklearn.cross_decomposition import CCA
cca = CCA(n_components=50)
cca.fit(cawi1_embeddings, cawi2_embeddings)
x_c, y_c = cca.transform(cawi1_embeddings, cawi2_embeddings)
dimension_correlations = []
for i in range(50):
correlation = pearsonr(x_c[:, i], y_c[:, i])
dimension_correlations.append(correlation[0])
plt.plot(dimension_correlations)
plt.show()
# -
| personalized_nlp/notebooks/cawi1_cawi2_embeddings.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
df = pd.read_csv('../prep-data/cement_slump.csv')
df.head()
df.columns
plt.figure(figsize=(10,8),dpi=200)
sns.heatmap(df.corr(),annot=True)
df.columns
X = df.drop('Compressive Strength (28-day)(Mpa)',axis=1)
y = df['Compressive Strength (28-day)(Mpa)']
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=101)
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
X_train = scaler.fit_transform(X_train)
X_test = scaler.transform(X_test)
from sklearn.svm import SVR, LinearSVC
# +
# with noisy data we should decrease C -> more regularization
# epsilon = 0 -> the maximum allowable error per training instance is zero -> overfits the model to the train data!
# -
base_model = SVR()
base_model.fit(X_train, y_train)
y_base_pred = base_model.predict(X_test)
from sklearn.metrics import mean_absolute_error, mean_squared_error
mean_absolute_error(y_test, y_base_pred)
np.sqrt(mean_squared_error(y_test, y_base_pred))
y_test.mean()
p_grid = {'C':[.001,.01,.1,.5,1],
'kernel':['linear','rbf','poly'],
'gamma':['scale','auto'],
'degree':[2,3,4],
'epsilon':[0,.01,.1,.5,1,2]}
from sklearn.model_selection import GridSearchCV
svr = SVR()
grid = GridSearchCV(svr, p_grid,verbose=1)
grid.fit(X_train,y_train)
grid.best_params_
y_grid_pred = grid.predict(X_test)
mean_absolute_error(y_test, y_grid_pred)
np.sqrt(mean_squared_error(y_test, y_grid_pred))
| prep-9-svm/svm2-regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction - Deep Learning
# Classical programming is all about creating a function that helps us to process input data and get the desired output.
#
# In the learning paradigm, we change the process so that given a set of examples of input data and desired output, we aim to learn the function that can process the data.
#
# - In machine learning, we end up handcrafting the features and then learn the function to get the desired output
# - In deep learning, we want to both learn the features and the function together to get the desired output
#
# 
# Lets take an example to understand both the learning paradigms - Machine Learning and Deep Learning
#
# We will be starting with a classification exercise. And using the **Fashion Mnist** dataset to do so. It involves identifying the 10 types of products that are there in the image.
#
# - Train: 60,000 images
# - Test: 10,000 images
# - Class: 10
# - Labels:
# - 0: T-shirt/top
# - 1: Trouser
# - 2: Pullover
# - 3: Dress
# - 4: Coat
# - 5: Sandal
# - 6: Shirt
# - 7: Sneaker
# - 8: Bag
# - 9: Ankle boot
# ### Get Input and Output
import numpy as np
import keras
import tensorflow as tf
from keras.datasets import fashion_mnist
from helpers import fashion_mnist_label
(x_train, y_train), (x_test, y_test) = fashion_mnist.load_data()
x_train.shape, y_train.shape, x_test.shape, y_test.shape
label = fashion_mnist_label()
label
# ### View the Dataset
import matplotlib.pyplot as plt
% matplotlib inline
# #### See an Image
# Lets plot a single image
def image(index):
plt.imshow(x_train[index], cmap="gray")
plt.title(label[y_train[index]])
image(0)
# ### See an Image from each class
# Lets plot one image from each class
u, indices = np.unique(y_train, return_index=True)
plt.figure(figsize = (16,7))
for i in u:
plt.subplot(2,5,i+1)
image(indices[i])
# #### See 500 of the Images
from helpers import create_sprite, create_embedding
sprite = create_sprite(x_train[:500])
plt.figure(figsize = (10,10))
plt.imshow(sprite,cmap='gray')
x_train[:500].shape
create_embedding("fashion-mnist", "fashion-mnist-embedding", 500)
# After running the above command in terminal - you should see the tensorboard running
#
# 
# ## ML Approach - 2 Feature Classification
# 
# **Step 1: Prepare the images and labels**
#
# Convert from 'uint8' to 'float32' and normalise the data to (0,1)
x_train = x_train.astype("float32")/255
x_test = x_test.astype("float32")/255
# Flatten the data from (60000, 28, 28) to (60000, 784)
x_train_flatten = x_train.reshape(60000, 28 * 28)
x_test_flatten = x_test.reshape(10000, 28 * 28)
# Convert class vectors to binary class matrices
from keras.utils import to_categorical
y_train_class = to_categorical(y_train, 10)
y_test_class = to_categorical(y_test, 10)
# **Step 2: Craft the feature transfomation **
#
# Use PCA to reduce from 784 features to 2 features
from sklearn.decomposition import PCA
pca = PCA(n_components=2).fit(x_train_flatten)
x_train_feature_2 = pca.transform(x_train_flatten)
x_test_feature_2 = pca.transform(x_test_flatten)
# **Step 3: Create a classifier to predict the 10 classes**
import keras
from keras.models import Sequential
from keras.layers import Dense
from keras.optimizers import SGD
model_simple2_ml = Sequential()
model_simple2_ml.add(Dense(10, input_shape=(2,), activation='softmax'))
model_simple2_ml.summary()
# **Step 4: Compile and fit the model**
model_simple2_ml.compile(loss='categorical_crossentropy', optimizer="sgd", metrics=['accuracy'])
# +
# Callback to save log files for tensorboard
tbCallBack = keras.callbacks.TensorBoard(log_dir='logs/simple2-ml')
# Callback to save log files after every batch
from helpers import MetricHistory
history = MetricHistory()
# -
# %%time
model_simple2_ml.fit(x_train_feature_2, y_train_class, batch_size=128, epochs=2,verbose=1,
validation_data=(x_test_feature_2, y_test_class),
callbacks=[tbCallBack, history])
# **Step 5: Check the performance of the model**
score = model_simple2_ml.evaluate(x_test_feature_2, y_test_class, verbose=0)
print('Test loss:', score[0])
print('Test accuracy:', score[1])
# **Step 6: Make & Visualise the Prediction**
model_simple2_ml.predict_proba(x_test_feature_2)[0]
from helpers import plot_prediction, plot_2d_model
plot_prediction(0, x_test, y_test, x_test_feature_2, model_simple2_ml)
plot_2d_model(model_simple2_ml, x_train_feature_2, y_train)
# ## ML Approach - 100 Feature Classification
# 
# **Step 2: Craft the feature transfomation **
#
# Lets take craft 100 features (using PCA) and then classify the result
from sklearn.decomposition import PCA
pca = PCA(n_components=100).fit(x_train_flatten)
x_train_feature_100 = pca.transform(x_train_flatten)
x_test_feature_100 = pca.transform(x_test_flatten)
# **Step 3: Create a classifier to predict the 10 classes**
model_simple100_ml = Sequential()
model_simple100_ml.add(Dense(10, input_shape=(100,), activation='softmax'))
model_simple100_ml.summary()
# **Step 4: Compile and fit the model**
model_simple100_ml.compile(loss='categorical_crossentropy', optimizer="sgd", metrics=['accuracy'])
tbCallBack = keras.callbacks.TensorBoard(log_dir='logs/simple100-ml')
# %%time
model_simple100_ml.fit(x_train_feature_100, y_train_class, batch_size=128, epochs=2, verbose=1,
validation_data=(x_test_feature_100, y_test_class),
callbacks=[tbCallBack, history])
# **Step 5: Check the performance of the model**
score = model_simple100_ml.evaluate(x_test_feature_100, y_test_class, verbose=0)
print('Test loss:', score[0])
print('Test accuracy:', score[1])
# **Step 6: Make & Visualise the Prediction**
model_simple100_ml.predict_proba(x_test_feature_100)[0]
plot_prediction(0, x_test, y_test, x_test_feature_100, model_simple100_ml)
# ## Deep Learning Approach - Single Layer
#
# 
#
# Lets learn both the representation and the classifier together now
# **Step 2 & 3: Craft the feature transfomation and classifier model **
model_single_dl = Sequential()
model_single_dl.add(Dense(100, input_shape=(784,), activation='linear'))
model_single_dl.add(Dense(10, activation='softmax'))
model_single_dl.summary()
# **Step 4: Compile and fit the model**
model_single_dl.compile(loss='categorical_crossentropy', optimizer="sgd", metrics=['accuracy'])
tbCallBack = keras.callbacks.TensorBoard(log_dir='logs/single-dl')
# %%time
model_single_dl.fit(x_train_flatten, y_train_class, batch_size=2, epochs=2, verbose=1,
validation_data=(x_test_flatten, y_test_class),
callbacks=[tbCallBack, history])
# **Step 5: Check the performance of the model**
score = model_single_dl.evaluate(x_test_flatten, y_test_class, verbose=1)
print('Test loss:', score[0])
print('Test accuracy:', score[1])
# **Step 6: Make & Visualise the Prediction**
model_single_dl.predict_proba(x_test_flatten)[0]
plot_prediction(0, x_test, y_test, x_test_flatten, model_single_dl)
# ## Deep Learning Approach - Multi Layer Perceptron
#
# **Step 2 & 3: Craft the feature transfomation and classifier model **
model_multi_dl = Sequential()
model_multi_dl.add(Dense(100, input_shape=(784,), activation='sigmoid'))
model_multi_dl.add(Dense(50, activation='sigmoid'))
model_multi_dl.add(Dense(10, activation='softmax'))
model_multi_dl.summary()
# **Step 4: Compile and fit the model**
model_multi_dl.compile(loss='categorical_crossentropy', optimizer="sgd", metrics=['accuracy'])
tbCallBack = keras.callbacks.TensorBoard(log_dir='logs/multi-dl')
# %%time
model_multi_dl.fit(x_train_flatten, y_train_class, batch_size=128, epochs=2, verbose=1,
validation_data=(x_test_flatten, y_test_class),
callbacks=[tbCallBack, history])
# **Step 5: Check the performance of the model**
score = model_multi_dl.evaluate(x_test_flatten, y_test_class, verbose=1)
print('Test loss:', score[0])
print('Test accuracy:', score[1])
# **Step 6: Make & Visualise the Prediction**
model_multi_dl.predict_proba(x_test_flatten)[0]
plot_prediction(0, x_test, y_test, x_test_flatten, model_multi_dl)
| experiments/Version1/01-Introduction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/marta-0/medical_mnist/blob/main/notebooks/Medical-MNIST.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="lqYZsOMU-Lti"
# # Medical MNIST
# + id="1DoNXgJB-MGp"
import os
import pathlib
import random
import gc
import zipfile
from PIL import Image
import albumentations as A
import numpy as np
import math
import seaborn as sns
import plotly.graph_objects as go
import matplotlib.pyplot as plt
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.metrics import confusion_matrix
from sklearn.manifold import TSNE
import tensorflow as tf
from tensorflow.keras.models import Sequential, Model
from tensorflow.keras.layers import Conv2D, MaxPooling2D, Flatten, Dense, Dropout, GlobalMaxPooling2D, BatchNormalization, Activation
from tensorflow.keras.utils import to_categorical
from tensorflow.python.keras.preprocessing import image
random.seed(1)
np.random.seed(1)
tf.random.set_seed(1)
# + [markdown] id="EwxIlLyP-PeV"
# ## Download the files
# + colab={"base_uri": "https://localhost:8080/"} id="242qqE8N-SAZ" outputId="b020d7de-a36b-4663-91d1-fc65f0ecff5f"
# !git clone https://github.com/apolanco3225/Medical-MNIST-Classification
# + [markdown] id="8pNdNn0z-T11"
# ## Check the classes and prepare the lists of files and labels
# + colab={"base_uri": "https://localhost:8080/"} id="BFn2SORT-je-" outputId="989776d5-30cf-4d74-a697-9cf0675aa6a9"
PATH = '/content/Medical-MNIST-Classification/resized'
p = pathlib.Path(PATH)
labels = [x.parts[-1] for x in p.iterdir()]
num_classes = len(labels)
labels.sort()
labels
# + id="aojCay4P-lQC"
files_paths = []
files_labels = []
for root, dirs, files in os.walk(PATH):
p = pathlib.Path(root)
for file in files:
files_paths.append(root + '/' + file)
files_labels.append(p.parts[-1])
# + colab={"base_uri": "https://localhost:8080/"} id="9Q6AH24I-lSb" outputId="2eeeebc9-2ca0-4cae-ea7a-d79c19ba2265"
len(files_labels), len(files_paths)
# + colab={"base_uri": "https://localhost:8080/"} id="33bmwelc-lUi" outputId="197196cc-84d2-41fc-fc6d-ef6570235a3b"
labels_cnt = []
for label in labels:
print(label, files_labels.count(label))
labels_cnt.append(files_labels.count(label))
# + colab={"base_uri": "https://localhost:8080/", "height": 454} id="sXOpesY1-lW1" outputId="c4067b51-3464-4b53-b87b-3d7c0d683c53"
plt.figure(figsize=(15,7))
plt.bar(labels, labels_cnt)
plt.title('Size of each class', fontsize=20)
plt.xticks(fontsize=15)
plt.yticks(fontsize=15);
# + [markdown] id="XdzkLjBF--vO"
# BreastMRI has less instances than the other classes.
# + [markdown] id="C_6NZKNJ_En7"
# ## Prepare X and y
# + [markdown] id="Y_kLiXVo_TA1"
# Images have been preprocessed already while collecting from databases. Each image is 64 x 64 px.
# + id="WeqEDAiT-lY2"
def prepare_X_y(files_paths, width=64, height=64):
X = []
for path in files_paths:
img = Image.open(path)
img.load()
if (width or height) != 64:
img = img.resize((width,height))
img_X = np.asarray(img, dtype=np.int16)
X.append(img_X)
X = np.asarray(X)
y = np.asarray(files_labels)
return X, y
# + id="p7XzFPjx_H3H"
X, y = prepare_X_y(files_paths)
# + colab={"base_uri": "https://localhost:8080/"} id="IpZsIQLw_PRB" outputId="3035a22c-af69-4999-9b4a-7cde1a41c102"
X.dtype
# + colab={"base_uri": "https://localhost:8080/"} id="D6FwBqpt_H45" outputId="a5c6d990-575d-434a-e17c-b6e93790ba96"
X.shape, y.shape
# + colab={"base_uri": "https://localhost:8080/"} id="bVE6dpf__H7b" outputId="654fd9db-1491-4e77-8a60-65de0bfd8578"
size = X.shape[-1]
size
# + [markdown] id="C2Okr14LAEmT"
# ## Visualisations
# + colab={"base_uri": "https://localhost:8080/", "height": 273} id="6wvqMbNW_RwY" outputId="174244cc-721c-4f45-bca8-3c9a8751b276"
plt.imshow(X[10], cmap='gray')
plt.title(y[10], fontsize=20)
plt.xticks([])
plt.yticks([]);
# + colab={"base_uri": "https://localhost:8080/", "height": 650} id="Q88vZaahAIZF" outputId="6d077289-ea8b-4940-efd8-b955577ea991"
plt.figure(figsize=(15,10))
plt.suptitle('Average image of each class', fontsize=20)
i = 0
for label in labels:
X_sum = np.sum(X[y ==label], axis=0)
X_avg = X_sum / files_labels.count(label)
plt.subplot(2,3,i+1)
plt.imshow(X_avg, cmap='gray')
plt.title(label, fontsize=20)
plt.xticks([])
plt.yticks([])
i = i + 1
# + [markdown] id="59YdgmXjAVgj"
# Several examples of each class:
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="WCS_YgKdAIbt" outputId="358b39f9-04ee-457a-b9a8-ee25da5eabe1"
for label in labels:
images = X[y == label]
plt.figure(figsize=(25,35))
for i in range(5):
plt.subplot(6, 5, i+1)
plt.imshow(images[i], cmap='gray')
plt.title(label, fontsize=20)
plt.xticks([])
plt.yticks([])
# + [markdown] id="J7uwcJTPAaNB"
# Examples of unusual images from each class:
# + colab={"base_uri": "https://localhost:8080/", "height": 650} id="9iWDvObWAIeM" outputId="b795cc53-e1b9-42df-e83b-47214b3c0423"
plt.figure(figsize=(15,10))
plt.suptitle('Examples of unusual and difficult to recognize images', fontsize=20)
def plot_unusual(i, path, title):
plt.subplot(2,3,i)
img = Image.open(path)
plt.imshow(img, cmap='gray')
plt.title(title, fontsize=20)
plt.xticks([])
plt.yticks([])
plot_unusual(1, '/content/Medical-MNIST-Classification/resized/AbdomenCT/005658.jpeg', 'AbdomenCT')
plot_unusual(2, '/content/Medical-MNIST-Classification/resized/BreastMRI/001248.jpeg', 'BreastMRI')
plot_unusual(3, '/content/Medical-MNIST-Classification/resized/CXR/009068.jpeg', 'CXR')
plot_unusual(4, '/content/Medical-MNIST-Classification/resized/ChestCT/006695.jpeg', 'ChestCT')
plot_unusual(5, '/content/Medical-MNIST-Classification/resized/Hand/006398.jpeg', 'Hand')
plot_unusual(6, '/content/Medical-MNIST-Classification/resized/HeadCT/000028.jpeg', 'HeadCT')
# + [markdown] id="EsuDwlz4CPAg"
# ## Train test split
# + [markdown] id="THkxsHbCCUmo"
# With *stratify*:
# + id="dae4jOcvAIg3"
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=1, stratify=y)
# + id="yMKHMC_KAIjd"
labels_cnt_train = []
labels_cnt_test = []
for label in labels:
labels_cnt_train.append(np.count_nonzero(y_train == label))
labels_cnt_test.append(np.count_nonzero(y_test == label))
# + colab={"base_uri": "https://localhost:8080/"} id="MozeTRqOAIl1" outputId="7c96a170-d2c9-4e75-869d-3e4d6654ec83"
labels_cnt_train, labels_cnt_test
# + colab={"base_uri": "https://localhost:8080/", "height": 454} id="mUrBmsxXAIoT" outputId="4402b6d0-42f5-410f-bc30-ea65e0345826"
plt.figure(figsize=(12,7))
plt.bar(labels, labels_cnt_train, label='train', color='#1F77B4')
plt.bar(labels, labels_cnt_test, bottom=labels_cnt_train, label='test', color='#FFA15A')
plt.title('Number of images in train and testset after stratified split', fontsize=20)
plt.legend()
plt.xticks(fontsize=15)
plt.yticks(fontsize=15);
# + [markdown] id="KUG5h7hWCuQn"
# Without *stratify*:
# + id="Onvr3pffAIqc"
X_train_v2, X_test_v2, y_train_v2, y_test_v2 = train_test_split(X, y, test_size=0.3, random_state=1)
# + id="bQ3oZecQC5tc"
labels_cnt_train_v2 = []
labels_cnt_test_v2 = []
for label in labels:
labels_cnt_train_v2.append(np.count_nonzero(y_train_v2 == label))
labels_cnt_test_v2.append(np.count_nonzero(y_test_v2 == label))
# + colab={"base_uri": "https://localhost:8080/"} id="THp4WH7QC5vx" outputId="5b139499-d365-42a5-e34e-75233e1eeee7"
labels_cnt_train_v2, labels_cnt_test_v2
# + colab={"base_uri": "https://localhost:8080/", "height": 454} id="Rx_ZogUqC5yS" outputId="a18ba39a-dd80-46c5-fe68-60af3f2dfe1e"
plt.figure(figsize=(12,7))
plt.bar(labels, labels_cnt_train_v2, label='train', color='#636EFA')
plt.bar(labels, labels_cnt_test_v2, bottom=labels_cnt_train_v2, label='test', color='#FECB52')
plt.title('Number of images in train and testset after unstratified split', fontsize=20)
plt.legend()
plt.xticks(fontsize=15)
plt.yticks(fontsize=15);
# + [markdown] id="MPK2oblTDIWa"
# Compare both split types:
# + colab={"base_uri": "https://localhost:8080/", "height": 717} id="kMXNbkYlC50t" outputId="3169ce26-bc24-4f21-bd72-a2c43ad5c3cd"
fig = go.Figure(
data = [
go.Bar(name="X_train", x=labels, y=labels_cnt_train, offsetgroup=0, marker_color='#1F77B4'),
go.Bar(name="X_test", x=labels, y=labels_cnt_test, offsetgroup=0, base=labels_cnt_train, marker_color='#FFA15A'),
go.Bar(name="X_train without stratify", x=labels, y=labels_cnt_train_v2, offsetgroup=1, marker_color='#636EFA'),
go.Bar(name="X_test without stratify", x=labels, y=labels_cnt_test_v2, offsetgroup=1, base=labels_cnt_train_v2, marker_color='#FECB52')],
layout=go.Layout(
title='Comparison of train and testset sizes after stratified and unstritified split',
yaxis_title='Set size', height=700, font=dict(size=15))
)
fig.add_shape(type='line', x0=-1, y0=7000, x1=6, y1=7000, line=dict(width=1, dash='dot'))
fig.show()
# + [markdown] id="bAsSKpy6EFTd"
# ## Data augmentation
# + id="BzAbPEMFC52-"
def data_augmentation(X_train, y_train, hflip=False, crop=False, rotate=False):
sub = 1
if (hflip or crop or rotate):
plt.figure(figsize=(20,4))
plt.suptitle('Examples of augmented images', fontsize=20)
if hflip:
X_hflip = X_train[:,:,::-1]
X_train = np.vstack((X_train, X_hflip))
y_train = np.hstack((y_train, y_train))
plt.subplot(1,6,sub)
plt.imshow(X_train[40000], cmap='gray')
plt.title('Original - {}'.format(y_train[40000]))
plt.subplot(1,6,sub+1)
plt.imshow(X_hflip[40000], cmap='gray')
plt.title('Horizontal flip - {}'.format(y_train[40000]))
sub = sub + 2
if crop:
X_crop = []
transform = A.RandomSizedCrop((size-size//2, size-size//5), size, size, p=1)
for i in range(X_train.shape[0]):
aug_img = transform(image=X_train[i])['image']
X_crop.append(aug_img)
X_crop = np.asarray(X_crop)
X_train = np.vstack((X_train, X_crop))
y_train = np.hstack((y_train, y_train))
plt.subplot(1,6,sub)
plt.imshow(X_train[40000], cmap='gray')
plt.title('Original - {}'.format(y_train[40000]))
plt.subplot(1,6,sub+1)
plt.imshow(X_crop[40000], cmap='gray')
plt.title('Crop - {}'.format(y_train[40000]))
sub = sub + 2
if rotate:
X_rotate=[]
transform = A.Rotate(180, p=1)
for i in range(X_train.shape[0]):
aug_img = transform(image=X_train[i])['image']
X_rotate.append(aug_img)
X_rotate = np.asarray(X_rotate)
X_train = np.vstack((X_train, X_rotate))
y_train = np.hstack((y_train, y_train))
plt.subplot(1,6,sub)
plt.imshow(X_train[40000], cmap='gray')
plt.title('Original - {}'.format(y_train[40000]))
plt.subplot(1,6,sub+1)
plt.imshow(X_rotate[40000], cmap='gray')
plt.title('Rotation - {}'.format(y_train[40000]))
return X_train, y_train
# + colab={"base_uri": "https://localhost:8080/"} id="dH7FlZ3aEIeT" outputId="1c4e258f-7400-4231-d263-3f5cbd424ac4"
X_train, y_train = data_augmentation(X_train, y_train)
X_train.shape, y_train.shape
# + [markdown] id="0phk6JlGEcTK"
# ## Prepare sets for model
# + [markdown] id="sEHhUeweEgZh"
# Pixels has values 0-255, 115-255 or 50-255:
# + colab={"base_uri": "https://localhost:8080/"} id="2F36mtckEIgu" outputId="063a36d3-79c0-44f9-87af-927c84fbafec"
for label in labels:
print(label, ' ', np.min(X[y==label]), '-', np.max(X[y == label]))
# + id="kJrv0-DBEIi5"
def preprocess(X_train, y_train, X_test, y_test):
'''Normalization, add channel, convert labels into numbers, one-hot encode y'''
if np.max(X_train) > 1: X_train = X_train / np.max(X_train)
if np.max(X_test) > 1: X_test = X_test / np.max(X_test)
if X_train.ndim == 3: X_train = np.expand_dims(X_train, axis=-1)
if X_test.ndim == 3 : X_test = np.expand_dims(X_test, axis=-1)
if y_train.dtype.type is np.str_:
y_train = list(map(lambda x: labels.index(x), y_train))
y_train = np.asarray(y_train)
y_test = list(map(lambda x: labels.index(x), y_test))
y_test = np.asarray(y_test)
if y_train.ndim == 1:
y_train = to_categorical(y_train, num_classes)
y_test = to_categorical(y_test, num_classes)
return X_train, y_train, X_test, y_test
# + id="_8b3WZBxEIlT"
X_train, y_train, X_test, y_test = preprocess(X_train, y_train, X_test, y_test)
# + colab={"base_uri": "https://localhost:8080/"} id="ti2nheSYEIni" outputId="b84dec4a-82af-4897-8a05-184d76a2240b"
print('Maximum pixel values: ', np.max(X_train), np.max(X_test))
print('X shape: ', X_train.shape, X_test.shape)
print('y shape: ', y_train.shape, y_test.shape)
# + colab={"base_uri": "https://localhost:8080/"} id="kbVhZYJrEIpv" outputId="a218a79e-b4f4-4f1f-80c3-5ec12992a160"
input_shape = X_train.shape[1:]
input_shape
# + id="1uIqLBfSGqDl"
np.save('X_train_' + str(size), X_train)
np.save('X_test_' + str(size), X_test)
np.save('y_train_' + str(size), y_train)
np.save('y_test_' + str(size), y_test)
# + [markdown] id="Rz4Xp-8HFLbf"
# Images generator - randomly selects resolution from 24 x 24 to 64 x 64 px:
# + id="eDumqTOpEIsG"
def generate_images_various_sizes(X_train, y_train, batch_size):
bx = []
by = []
batch_count = 0
size = random.randrange(24, 64)
while True:
for i in range(X_train.shape[0]):
transform = A.Resize(size, size, p=1)
x = transform(image=X_train[i])['image']
y = y_train[i]
batch_count += 1
bx.append(x)
by.append(y)
if batch_count > batch_size:
bx = np.asarray(bx, dtype=np.float32)
by = np.asarray(by, dtype=np.float32)
yield (bx, by)
bx = []
by = []
batch_count = 0
size = random.randrange(24, 64)
# + [markdown] id="Rg2PELL9zxgZ"
# Occluded images generator - occludes randomly selected image fragment:
# + id="BEaKM5RYz8lA"
def generate_occluded_images(X_train, y_train, batch_size, occ_size=16, occ_pixel=0):
bx = []
by = []
batch_count = 0
h_start = random.randrange(0, size-occ_size-1)
w_start = random.randrange(0, size-occ_size-1)
while True:
for i in range(X_train.shape[0]):
x = X_train[i]
x[h_start:h_start+occ_size, w_start:w_start+occ_size] = occ_pixel
y = y_train[i]
batch_count += 1
bx.append(x)
by.append(y)
if batch_count > batch_size:
bx = np.asarray(bx, dtype=np.float32)
by = np.asarray(by, dtype=np.float32)
yield (bx, by)
bx = []
by = []
batch_count = 0
h_start = random.randrange(0, size-occ_size-1)
w_start = random.randrange(0, size-occ_size-1)
# + [markdown] id="Va4FVGaSFOOi"
# ## Model
# + id="XnRyHxH1C54_"
def get_model():
model = Sequential([
Conv2D(32, (3,3), activation='relu', input_shape=input_shape),
MaxPooling2D((2,2)),
Conv2D(64, (3,3), activation='relu'),
MaxPooling2D((2,2)),
Conv2D(64, (3,3), activation='relu'),
Flatten(),
Dense(64, activation='relu'),
Dense(num_classes, activation='softmax')
])
return model
# + colab={"base_uri": "https://localhost:8080/"} id="-M8kteG_FRPK" outputId="8e7d2620-73a5-4e0e-bb11-1b99bf37d7ea"
model_name = 'baseline'
model = get_model()
model.summary()
# + id="5OYvZqWdFRRQ"
def train_model(maxit, X, y, batch_size=32, epochs=30, verbose=0, steps_per_epoch=None):
errors_cnt = []
history_acc = []
history_val_acc = []
history_loss = []
history_val_loss = []
for _ in range(maxit):
random.seed(1)
np.random.seed(1)
tf.random.set_seed(1)
model = get_model()
model.compile(optimizer='adam', loss='categorical_crossentropy', metrics=['accuracy'])
history = model.fit(X, y, epochs=epochs, validation_data=(X_test, y_test), verbose=verbose, steps_per_epoch=steps_per_epoch, batch_size=batch_size)
y_pred = model.predict(X_test)
errors = np.count_nonzero(np.argmax(y_test, axis=1) != np.argmax(y_pred, axis=1))
errors_cnt.append(errors)
print('Iteration: ', _ + 1, 'Mistakes: ', errors)
history_acc.append(history.history['accuracy'])
history_val_acc.append(history.history['val_accuracy'])
history_loss.append(history.history['loss'])
history_val_loss.append(history.history['val_loss'])
if _ + 1 < maxit:
del model, y_pred, errors, history
gc.collect()
else:
tf.io.gfile.makedirs('results/' + model_name)
np.save('/content/results/' + model_name + '/errors', errors_cnt)
np.save('/content/results/' + model_name + '/accuracy', history_acc)
np.save('/content/results/' + model_name + '/val_accuracy', history_val_acc)
np.save('/content/results/' + model_name + '/loss', history_loss)
np.save('/content/results/' + model_name + '/val_loss', history_val_loss)
return model, y_pred, errors, history
# + colab={"base_uri": "https://localhost:8080/"} id="uqE6laxrFRTr" outputId="a090f638-427d-410d-efc9-44517aa2cfe0"
model, y_pred, errors, history = train_model(1, X=X_train, y=y_train, verbose=1)
# + id="Tj3_V8h8dpAc" colab={"base_uri": "https://localhost:8080/"} outputId="4895d6fd-d78c-4402-f823-dc530fd46846"
model.save('models/' + model_name)
# + [markdown] id="TYxev-sNXjfL"
# Train with image generator (different images sizes):
# + id="MJe2daYSZDNa"
batch_size = 32
train_model(100, X=generate_images_various_sizes(X_train, y_train, batch_size), y=None, steps_per_epoch=X_train.shape[0]/batch_size)
# + [markdown] id="fiU9giGaeo7R"
# ## Results
# + [markdown] id="IFra1XMpfASU"
# Accuracy and loss:
# + id="yHFALRDgXixB"
def plot_acc_loss(history):
plt.figure(figsize=(20,6))
plt.subplot(1,2,1)
plt.plot(history.history['accuracy'], label='accuracy')
plt.plot(history.history['val_accuracy'], label='val_accuracy')
plt.xlabel('Epoch', fontsize=15)
plt.ylabel('Accuracy', fontsize=15)
plt.legend(loc=4)
plt.xticks(fontsize=15)
plt.yticks(fontsize=15)
plt.subplot(1,2,2)
plt.plot(history.history['loss'], label='loss')
plt.plot(history.history['val_loss'], label='val_loss')
plt.xlabel('Epoch', fontsize=15)
plt.ylabel('Loss', fontsize=15)
plt.legend()
plt.xticks(fontsize=15)
plt.yticks(fontsize=15);
# + id="KHARfhIEXizu" colab={"base_uri": "https://localhost:8080/", "height": 398} outputId="7d530266-2023-466b-ec70-9ed7ce551be7"
plot_acc_loss(history)
# + [markdown] id="Zfa8-Z3HfCRn"
# Confusion matrix:
# + id="qmxtU_T6exBz"
def get_classes(y_test, y_pred):
y_test_classes = []
y_pred_classes = []
for i, val in enumerate(y_test):
y_test_classes.append(np.argmax(val))
for i, val in enumerate(y_pred):
y_pred_classes.append(np.argmax(val))
return y_test_classes, y_pred_classes
def show_confusion_matrix():
y_test_classes, y_pred_classes = get_classes(y_test, y_pred)
cm = confusion_matrix(y_test_classes, y_pred_classes)
plt.figure(figsize=(11, 10))
sns.heatmap(cm, annot=True, xticklabels=labels, yticklabels=labels, annot_kws={'size':15}, cbar=False)
plt.xlabel('Predicted values', fontsize=15)
plt.ylabel('True values', fontsize=15)
plt.xticks(fontsize=15, rotation=0)
plt.yticks(fontsize=15, va='center');
# + id="lKQNxb5fexES" colab={"base_uri": "https://localhost:8080/", "height": 616} outputId="4dd9ecd9-9152-497b-bdea-fd62f05e7d40"
show_confusion_matrix()
# + [markdown] id="wyt7qvXrfEIO"
# Visualise model's mistakes:
# + id="qGaZC1U3exG3"
def wrong_pred(y_test, y_pred):
for idx, (a, b) in enumerate(zip(y_test, y_pred)):
if np.argmax(a) == np.argmax(b): continue
yield idx, np.argmax(a), np.argmax(b)
def show_mistakes():
X_test_plot = X_test.reshape( X_test.shape[:-1] )
plt.figure(figsize=(20,15))
for i, (idx, y_test_val, y_pred_val) in enumerate(wrong_pred(y_test, y_pred)):
plt.subplot(4, 5, i+1)
plt.imshow(X_test_plot[idx], cmap='gray')
plt.title('True: {}\nPred: {}'.format(labels[y_test_val], labels[y_pred_val]), fontsize=15)
plt.tight_layout()
plt.show();
# + id="wM-bqaOxexJB" colab={"base_uri": "https://localhost:8080/", "height": 563} outputId="1155ab25-e1c6-47a2-d39e-df29d01afc0f"
show_mistakes()
# + [markdown] id="25Rjl_1OeT2y"
# ## Reproducibility of results
# + id="jpwDxZEYXi4s" colab={"base_uri": "https://localhost:8080/"} outputId="ce72f70a-a64a-4edd-cbe2-2b69e900280c"
with tf.device("/cpu:0"):
print('CPU')
errors_cnt_cpu = train_model(5, X=X_train[::5], y=y_train[::5], epochs=1, verbose=1)
# + id="yfi-nhbqmCn2" colab={"base_uri": "https://localhost:8080/"} outputId="e19d3138-acf9-4cdd-9b67-3536adc665c0"
if tf.config.experimental.list_physical_devices("GPU"):
with tf.device("/gpu:0"):
print('GPU')
errors_cnt_gpu = train_model(5, X=X_train[::5], y=y_train[::5], epochs=1, verbose=1)
else:
print("GPU: not found")
# + id="JvdwqXXTkpkC"
| notebooks/Medical-MNIST.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
import numpy as np
from mpl_toolkits.mplot3d import Axes3D
import sympy as sy
sy.init_printing()
# # <font face="gotham" color="purple"> Null Space </font>
# The <font face="gotham" color="red"> null space </font>, denoted <font face="gotham" color="red">$\text{Nul}A$</font> is the solution set of a homogenous linear system $Ax=0$.
#
# Null space is a subspace of $\mathbb{R}^n$, why? Consider a linear system.
#
# $$
# 3x_1-x_2+x_3 = 0\\
# x_1+2x_2+3x_3= 0
# $$
#
# The augmented matrix is
#
# $$
# \left[
# \begin{matrix}
# 2 & -1 & 1 & 0\\
# 1 & 2 & 3 & 0
# \end{matrix}
# \right]
# $$
# Before solving the system, we have already known there is no unique solution since a free variable exists.
Aug = sy.Matrix([[2,-1,1,0],[1,2,3,0]])
Aug.rref()
# $x_3$ is a free variable, the solution set is
#
# $$
# \left[
# \begin{matrix}
# x_1 \\ x_2 \\ x_3
# \end{matrix}
# \right]=
# \left[
# \begin{matrix}
# -x_3 \\ -x_3 \\ x_3
# \end{matrix}
# \right]=
# x_3\left[
# \begin{matrix}
# -1 \\ -1 \\ 1
# \end{matrix}
# \right]
# $$
#
# which is a line passing through $(0, 0, 0)$ and $(-1, -1, 1)$, a subspace of $\mathbb{R}^3$.
# Consider another example, suppose we have an augmented matrix
Aug = sy.Matrix([[-3,6,-1,1,-7,0],[1,-2,2,3,-1,0],[2,-4,5,8,-4,0]]);Aug
Aug.rref()
# The solution can be written as:
# $$
# \left[
# \begin{matrix}
# x_1 \\ x_2 \\ x_3 \\x_4 \\ x_5
# \end{matrix}
# \right]=
# \left[
# \begin{matrix}
# 2x_2+x_4-3x_5 \\ x_2 \\ -2x_4+2x_5 \\x_4 \\ x_5
# \end{matrix}
# \right]=
# x_2\left[
# \begin{matrix}
# 2 \\ 1 \\ 0 \\0 \\ 0
# \end{matrix}
# \right]
# +
# x_4\left[
# \begin{matrix}
# 1 \\ 0 \\ -2 \\1 \\ 0
# \end{matrix}
# \right]
# +x_5\left[
# \begin{matrix}
# -3 \\ 0 \\ 2 \\0 \\ 1
# \end{matrix}
# \right]
# $$
# The $\text{Nul}A$ is a subspace in $\mathbb{R}^5$ with dimension of 3.
# # <font face="gotham" color="purple"> Null Space vs Col Space </font>
# Let
A = sy.Matrix([[2,4,-2,1],[-2,-5,7,3],[3,7,-8,6]]);A
# Column space is a subspace in $\mathbb{R}^n$, what is $n$? It is the number of rows, $n=3$.
#
# Null space is a subspace in $\mathbb{R}^m$, what is $m$? It is the number of columns, $m=4$.
# Find any nonzero vector in $\text{Col}A$ and in $\text{Nul}A$.
# Any column in the matrix can be a nonzero column in $\text{Col}A$, for instance first column: $(2, -2, 3)^T$.
# But to find a nonzero vector in null space requires some effort, construct the augmented matrix then turn it into rref.
Aug = sy.Matrix([[2,4,-2,1,0],[-2,-5,7,3,0],[3,7,-8,6,0]]);Aug.rref()
# The solution set with a free variable $x_3$ (because column 3 has no pivot) is
#
# $$
# \left[
# \begin{matrix}
# x_1 \\ x_2 \\ x_3\\x_4
# \end{matrix}
# \right]=
# \left[
# \begin{matrix}
# -9x_3 \\ 5x_3 \\ x_3\\0
# \end{matrix}
# \right]
# $$
#
# If we pick $x_3 =1$, a nonzero vector in $\text{Nul}A$ is $(-9, 5, 1, 0)^T$
# Now consider two vectors
#
# $$
# u = \left[
# \begin{matrix}
# 3 \\ -2 \\ -1\\ 0
# \end{matrix}
# \right],\qquad
# v = \left[
# \begin{matrix}
# 3 \\ -1\\3
# \end{matrix}
# \right]\\
# $$
# Is $u$ in $\text{Nul}A$? It can be verified easily
u = sy.Matrix([[3],[-2],[-1],[0]])
A*u
# $Au\neq \mathbf{0}$, therefore $u$ is not in $\text{Nul}A$.
# Is $v$ in $\text{Col}A$?
v = sy.Matrix([[3],[-1],[3]])
A.row_join(v).rref()
# The augmented matrix show there are solutions, i.e. $v$ is a linear combination of its column space basis, so $v$ is in $\text{Col}A$.
# # <font face="gotham" color="purple"> Row Space </font>
# <font face="gotham" color="red"> Row space</font>, denoted as $\text{Row}A$, is all linear combination of row vectors and subspace in $\mathbb{R}^n$.
# If we perform row operations on $A$ to get $B$, both matrices have the same row space, because $B$'s rows are linear combinations of $A$'s. However, row operation will change the row dependence.
# ## <font face="gotham" color="purple"> An Example </font>
# Find the row, column and null space of
A = sy.Matrix([[-2, -5, 8, 0, -17],
[1, 3, -5, 1, 5],
[3, 11, -19, 7, 1],
[1, 7, -13, 5, -3]]);A
B = A.rref();B
# The basis of row space of $B$ is its first 3 rows: $(1,0,1,0,1), (0, 1, -2, 0, 3), (0, 0, 0, 1, -5)$ which are also the basis of row space of $A$. However it does not necessarily mean that first 3 rows of $A$ forms the basis for row space, because the dependence among rows changed by row operation.
# In constrast, the basis of col space of $A$ is $(-2, 1, 3, 1)^T, (-5, 3, 11, 7)^T, (0, 1, 7, 5)^T$.
Aug = A.row_join(sy.zeros(4,1));Aug.rref()
# The null space is
#
# $$
# \left[
# \begin{matrix}
# x_1 \\ x_2 \\ x_3\\x_4 \\x_5
# \end{matrix}
# \right]=
# \left[
# \begin{matrix}
# -x_3-x_5 \\ 2x_3-3x_5 \\ x_3\\5x_5 \\x_5
# \end{matrix}
# \right]=
# x_3\left[
# \begin{matrix}
# -1 \\ 2 \\ 1\\0 \\0
# \end{matrix}
# \right]+
# x_5
# \left[
# \begin{matrix}
# -1 \\ -3 \\ 0\\5 \\1
# \end{matrix}
# \right]
# $$
# # <font face="gotham" color="purple"> Rank </font>
# Definition of rank:
# The <font face="gotham" color="red"> rank </font> is the dimension of the column space of $A$. The <font face="gotham" color="red"> nullity </font> of $A$ is the dimension of the null space.
# ## <font face="gotham" color="purple"> The Rank Theorem</font>
# The dimensions of the column space and the row space of an $m \times n$ matrix $A$ are equal that is why we only need to say rank is the dimension of the column space.
#
# This common dimension, the rank of $A$, also equals the number of pivot positions in $A$ and satisfies the equation
# $$
# \operatorname{rank} A+\operatorname{dim} \mathrm{Nul} A=n
# $$
# The intuition is that when a matrix $A$ is converted into rref $B$, we can indirectly(matching the same column from $B$ to $A$) see the basis of column space, those columns in corresponding rref have pivots.
#
# And in rref, we can also see the basis of row space directly, every row in the basis of row space have a pivot as well. And those rows which does not have pivots are for free variables, which is the dimension of null space.
# ## <font face="gotham" color="purple"> Example 1 </font>
# If $A$ is $45 \times 50$ matrix with a $10$-dimension nullity, what is the rank of $A$?
# $10D$ nullity means 10 free variables, so the pivots are $50-10=40$, which is also the rank of $A$.
# ## <font face="gotham" color="purple"> Example 2 </font>
# The matrices below are row equivalent.
# $$
# A=\left[\begin{array}{rrrrr}
# 2 & -1 & 1 & -6 & 8 \\
# 1 & -2 & -4 & 3 & -2 \\
# -7 & 8 & 10 & 3 & -10 \\
# 4 & -5 & -7 & 0 & 4
# \end{array}\right], \quad B=\left[\begin{array}{rrrrr}
# 1 & -2 & -4 & 3 & -2 \\
# 0 & 3 & 9 & -12 & 12 \\
# 0 & 0 & 0 & 0 & 0 \\
# 0 & 0 & 0 & 0 & 0
# \end{array}\right]
# $$
# 1. Find rank $A$ and $\operatorname{dim}$ Nul $A$
# 2. Find bases for Col $A$ and Row $A$.
# 3. What is the next step to perform to find a basis for Nul $A$ ?
# 4. How many pivot columns are in a row echelon form of $A^{T} ?$
# 1. $rank(A)=2$, because $B$ has two pivots. And nullity is the number of free variables, there are 3, so $\text{dim Nul}A = 3$.
# 2. Bases for $\text{Col}A$ is $(2,1,-7,4)^T, (-1,-2,8,-5)^T$, and for $\text{Row}A$ is $(1,-2,-4,3,-2),(0,3,9,-12,12)$.
# 3. Perform rref on augmented $A$
A = sy.Matrix([[2,-1,1,-6,8,0],
[1,-2,-4,3,-2,0],
[-7,8,10,3,-10,0],
[4,-5,-7,0,4,0]])
A.rref()
# The $\text{Nul}A$ and basis is
#
# $$
# \left[
# \begin{matrix}
# x_1 \\ x_2 \\ x_3\\x_4 \\x_5
# \end{matrix}
# \right]=
# \left[
# \begin{matrix}
# -2x_3+5x_4-6x_5 \\ -3x_3+4x_4-4x_5 \\ x_3\\x_4 \\x_5
# \end{matrix}
# \right]=
# x_3
# \left[
# \begin{matrix}
# -2 \\ -3 \\ 1\\0 \\0
# \end{matrix}
# \right]+
# x_4
# \left[
# \begin{matrix}
# 5 \\ 4 \\ 0\\1 \\0
# \end{matrix}
# \right]+
# x_5
# \left[
# \begin{matrix}
# -6 \\ -4 \\ 0\\0 \\1
# \end{matrix}
# \right]
# $$
# 4. Transpose $A$ then do rref.
A.T.rref()
# There are 2 pivot columns.
# Actually, we don't need any calculation to know the rank of $A^T$, because
#
# $$
# rank(A)=rank(A^T)
# $$
# # <font face="gotham" color="purple"> Orthogonality of $\text{Nul}A$ and $\text{Row}A$ </font>
# ## <font face="gotham" color="purple"> $\text{Nul}A \perp \text{Row}A$ </font>
# Here is the intersting connections of these subspaces we have discussed. Consider
A = sy.Matrix([[5, 8, 2], [10, 16, 4], [3, 4, 1]]);A
A.rref()
# The basis of row space of $A$ is $(1, 0, 0)$ and $(0, 1, .25)$.And the $\text{Row}A$ is
#
# $$
# \text{Row}A=
# s\left[
# \begin{matrix}
# 1 \\ 0\\ 0
# \end{matrix}
# \right]+
# t\left[
# \begin{matrix}
# 0 \\ 1\\ 0.25
# \end{matrix}
# \right]
# $$
# The $\text{Nul}A$ is
# $$
# \left[
# \begin{matrix}
# x_1 \\ x_2\\ x_3
# \end{matrix}
# \right]=
# x_3
# \left[
# \begin{matrix}
# 0 \\ -.25\\ 1
# \end{matrix}
# \right]
# $$
# Now we can visualize their relations geometrically.Again keep in mind that Matplotlib does not render 3D properly, so you need some imagination as well.
#
# Here is what we observe.
#
# The $\text{Row}A$ is a plane and $\text{Nul}A$ is a line which is perpendicular to the plane. It is easy to grasp the idea if you notice that in a homogeneous system $Ab = \mathbf{0}$, it breaks down into many dot products
#
# $$
# Ab =\left[
# \begin{matrix}
# A_{1i}\cdot b \\ A_{2i}\cdot b\\ A_{3i}\cdot b
# \end{matrix}
# \right]
# $$
#
# where $A_{1i}$ and etc are the rows of $A$. In later chapters we will prove when the dot product of two vectors equals zero, then they are perpendicular.
# +
# %matplotlib notebook
s = np.linspace(-1, 1, 10)
t = np.linspace(-1, 1, 10)
S, T = np.meshgrid(s, t)
X = S
Y = T
Z = T*.25
fig = plt.figure(figsize = (8,8))
ax = fig.add_subplot(111,projection='3d')
ax.plot_surface(X, Y, Z, alpha = .9, cmap=plt.cm.coolwarm)
x3 = np.linspace(-1, 1, 10)
x1 = 0*x3
x2 = -.25*x3
ax.plot(x1,x2,x3, lw = 5)
ax.set_xlabel('x-axis', size = 18)
ax.set_ylabel('y-axis', size = 18)
ax.set_zlabel('z-axis', size = 18)
ax.axis([-1,1,-1,1])
ax.text(x = 1, y = -1, z = -.25, s = r'$Row\ A$', size = 17)
ax.text(0, -.25, 1, s = r'$Nul\ A$', size = 17)
ax.view_init(7, 20)
# -
# ## <font face="gotham" color="purple"> $\text{Nul}A^T \perp \text{Col}A$ </font>
# The nullity of $A^T$ is
A = sy.Matrix([[5, 8, 2], [10, 16, 4], [3, 4, 1]]);A.T.rref()
# The $\text{Nul}A^T$ is
#
# $$
# \left[
# \begin{matrix}
# x_1 \\ x_2\\ x_3
# \end{matrix}
# \right]=
# x_2
# \left[
# \begin{matrix}
# -2 \\ 1\\ 0
# \end{matrix}
# \right]
# $$
# The $\text{Col}A$ is
A.rref()
# $$
# \text{Col}A=
# s\left[
# \begin{matrix}
# 5 \\ 10\\ 3
# \end{matrix}
# \right]+
# t\left[
# \begin{matrix}
# 8 \\ 16\\ 4
# \end{matrix}
# \right]
# $$
# $\text{Col}A$ is a plane and $\text{Nul}A^T$ is a line perpendicular to the plane. The intuition is similar to $\text{Nul}A \perp \text{Row}A$, here you can think of a system look like $b^TA = \mathbf{0}^T$.
# +
# %matplotlib notebook
s = np.linspace(-1, 1, 10)
t = np.linspace(-1, 1, 10)
S, T = np.meshgrid(s, t)
X = 5*S+8*T
Y = 10*S+16*T
Z = 3*S+4*T
fig = plt.figure(figsize = (8,8))
ax = fig.add_subplot(111,projection='3d')
ax.plot_surface(X, Y, Z, cmap=plt.cm.coolwarm)
x2 = np.linspace(-1, 1, 10)
x3 = x2*0
x1 = -2*x2
ax.plot(x1,x2,x3, lw = 3)
ax.set_xlabel('x-axis', size = 18)
ax.set_ylabel('y-axis', size = 18)
ax.set_zlabel('z-axis', size = 18)
ax.axis([-1,1,-1,1])
ax.view_init(-67, 35)
# -
# # <font face="gotham" color="purple"> Rank Decomposition </font>
# Consider a matrix $A$, the purpose is to decompose it into the multiplication of $C$, $R$, which are the bases of column space and row space respectively.
# $$
# A = CR
# $$
A = sy.Matrix([[2, 4, 1, -1], [4, 2, -4, 2], [2, -2, -5, 3], [1, 9, -3, 2]]);A
Arref = A.rref();Arref
# Get the basis of $\text{Col}A$.
ColA_basis = A[:,:3];ColA_basis
# Then get the $\text{Row}A$.
RowA_basis = Arref[0][0:3,:];RowA_basis
# Multiply $CR$, we are getting back $A$.
ColA_basis*RowA_basis
# Verify if $CR$ equals $A$.
ColA_basis*RowA_basis == A
| Chapter 10 -Null Space vs Col Space, Row Space and Rank.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Introduction to `numpy`
# Let's start by running the following cell.
import numpy as np
# As introduced in the pre-recorded lectures, `numpy` is a fast numerical computation module for Python. The basic tools that `numpy` offers are the *array* data structure and *vectorized* implementations of functions on arrays. In this worksheet, we will assume familiarity with the content of the `numpy` lectures.
#
# To recap, a `numpy` array is implemented as a C array: it is a contiguous block of memory that stores data all of the same type. This is in contrast with native Python lists, which are implemented as C arrays that store pointers to their contents, which can be scattered throughout memory. Eliminating this non-locality and the overhead cost of dynamic type checking (which lists must perform even if all their elements are of the same type) leads to a significant edge in speed for `numpy` arrays. This is why we use them.
# +
a = np.random.rand(1000)
b = np.random.rand(1000)
a_list = list(a)
b_list = list(b)
# -
# In the next cell, write a list comprehension that makes a new list `c_list` whose elements are products of corresponding elements of `a_list` and `b_list`. (Leave the `%%timeit` decorator in the cell; this will let you measure the speed of execution.)
# %%timeit
# write your code here
# Make a new array `c` filled with 1000 zeros. Write a `for` loop that iterates over the elements of arrays `a` and `b` and assigns their product to the corresponding entry of `c`. (The results should convince you that you should always avoid doing this.)
# %%timeit
# write your code here
# Now make `c` again, this time by multiplying the corresponding pairs of elements of `a` and `b` using the `numpy` vectorized multiplication.
# %%timeit
# write your code here
# What do you observe about the performance of each operation? (1 ms = 1000 µs, 1 µs = 1000 ns.)
# *write your answer here*
# ### §1. Building arrays
# In the following exercises, build the indicated arrays using `numpy` functions. Don't use lists or loops. Try not to store anything into variables -- you don't need to yet. You can do these in any order.
# ```
# array([ 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21,
# 22, 23, 24])
# ```
# this is an example
np.arange(5, 25)
# ```
# array([[ 5, 6, 7, 8, 9],
# [10, 11, 12, 13, 14],
# [15, 16, 17, 18, 19],
# [20, 21, 22, 23, 24]])
# ```
# +
# write your code here
# -
# ```
# array([[ 5, 6, 7, 8],
# [ 9, 10, 11, 12],
# [13, 14, 15, 16],
# [17, 18, 19, 20],
# [21, 22, 23, 24]])
# ```
# ```
# array([1., 1., 1., 1., 1.])
# ```
# ```
# array([[1., 1.],
# [1., 1.],
# [1., 1.]])
# ```
# ```
# array([[7., 7.],
# [7., 7.],
# [7., 7.]])
# ```
# ```
# array([ 1, 4, 7, 10, 13])
# ```
# ```
# array([ 1., 2., 4., 8., 16., 32., 64., 128., 256., 512.])
# ```
# ```
# array([ 0., 2., 4., 6., 8., 10.])
# ```
# ```
# array([20., 15., 10., 5., 0.])
# ```
# ```
# array([[[ 0, 1, 2, 3, 4],
# [ 5, 6, 7, 8, 9]],
#
# [[10, 11, 12, 13, 14],
# [15, 16, 17, 18, 19]],
#
# [[20, 21, 22, 23, 24],
# [25, 26, 27, 28, 29]]])
# ```
# ```
# array([False, False, False, False, False, False, False, False, True,
# True, True, True, True, True, True, True, True, True,
# True, True])
# ```
# +
# there are 20 entires
# -
# ```
# array([ True, False, True, False, True, False, True, False, True,
# False, True, False, True, False, True, False, True, False,
# True, False])
# ```
# ```
# array([False, False, False, False, False, False, False, False, True,
# False, True, False, True, False, True, False, True, False,
# True, False])
# ```
# From this point on it might help to store the arrays to variables. (I suggest just reusing the same variable name; these problems shouldn't depend on each other.)
# ```
# array([ 0, 1, 2, 0, 4, 5, 0, 7, 8, 0, 10, 11, 0, 13, 14, 0, 16,
# 17, 0, 19])
# ```
# ```
# array([ 0, 1, 0, 3, 0, 0, 0, 7, 0, 9, 0, 11, 0, 13, 0, 0, 0,
# 17, 0, 19])
# ```
# ```
# array([[0, 1, 2, 3],
# [4, 5, 6, 0],
# [0, 0, 0, 0]])
# ```
# ```
# array([[ 0, 1, 2, 13],
# [14, 15, 16, 17],
# [18, 19, 10, 11]])
# ```
# ```
# array([[ True, False, False],
# [False, True, False],
# [False, False, True]])
# ```
# +
# hint: use the // operator -- test it to see what it does
# -
# ```
# array([[1., 0., 0., 0., 0.],
# [0., 1., 0., 0., 0.],
# [0., 0., 1., 0., 0.],
# [0., 0., 0., 1., 0.],
# [0., 0., 0., 0., 1.]])
# ```
# +
# hint: build two arrays
# -
# ### §2. Slicing multidimensional arrays
# Run the next cell first.
A = np.arange(28).reshape(4, 7)
A
# Obtain the following arrays by slicing `A`.
# ```
# array([ 7, 8, 9, 10, 11, 12, 13])
# ```
# ```
# array([ 4, 11, 18, 25])
# ```
# ```
# array([[ 9, 10, 11],
# [16, 17, 18],
# [23, 24, 25]])
# ```
# ```
# array([[ 1, 2, 3, 4],
# [ 8, 9, 10, 11],
# [15, 16, 17, 18],
# [22, 23, 24, 25]])
# ```
# ```
# array([[ 0, 2, 4, 6],
# [ 7, 9, 11, 13],
# [14, 16, 18, 20],
# [21, 23, 25, 27]])
# ```
# ```
# array([[ 0, 3, 5, 6],
# [ 7, 10, 12, 13]])
# ```
# ### §3. Bonus
# Run the following cell and observe the output.
# +
A = np.arange(20).reshape(4,5)
print(A, A.sum(), A.sum(0), A.sum(1), sep='\n\n')
# -
v = np.arange(1, 6)
v
# On one line, using the above functions, write an expression for the matrix product `Av`, considering `v` as a column vector.
# Explain why your answer works.
# *write your answer here*
| discussion/intro-numpy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # 卷积神经示例: TF-SLIM高级API实现
# ### 推荐尽量用TF-SLIM实现复杂结构
import os
import numpy as np
from scipy import ndimage
import matplotlib.pyplot as plt
import tensorflow as tf
import tensorflow.contrib.slim as slim
from tensorflow.examples.tutorials.mnist import input_data
# %matplotlib inline
print ("当前TensorFlow版本为 [%s]" % (tf.__version__))
print ("所有包载入完毕")
# ## 载入 MNIST
mnist = input_data.read_data_sets('data/', one_hot=True)
trainimg = mnist.train.images
trainlabel = mnist.train.labels
valimg = mnist.validation.images
vallabel = mnist.validation.labels
testimg = mnist.test.images
testlabel = mnist.test.labels
print ("MNIST ready")
# ## 定义模型
# +
n_input = 784
n_classes = 10
x = tf.placeholder("float", [None, n_input])
y = tf.placeholder("float", [None, n_classes])
is_training = tf.placeholder(tf.bool)
def lrelu(x, leak=0.2, name='lrelu'):
with tf.variable_scope(name):
f1 = 0.5 * (1 + leak)
f2 = 0.5 * (1 - leak)
return f1 * x + f2 * abs(x)
def CNN(inputs, is_training=True):
x = tf.reshape(inputs, [-1, 28, 28, 1])
batch_norm_params = {'is_training': is_training, 'decay': 0.9
, 'updates_collections': None}
init_func = tf.truncated_normal_initializer(stddev=0.01)
net = slim.conv2d(x, 32, [5, 5], padding='SAME'
, activation_fn = lrelu
, weights_initializer = init_func
, normalizer_fn = slim.batch_norm
, normalizer_params = batch_norm_params
, scope='conv1')
net = slim.max_pool2d(net, [2, 2], scope='pool1')
net = slim.conv2d(x, 64, [5, 5], padding='SAME'
, activation_fn = lrelu
, weights_initializer = init_func
, normalizer_fn = slim.batch_norm
, normalizer_params = batch_norm_params
, scope='conv2')
net = slim.max_pool2d(net, [2, 2], scope='pool2')
net = slim.flatten(net, scope='flatten3')
net = slim.fully_connected(net, 1024
, activation_fn = lrelu
, weights_initializer = init_func
, normalizer_fn = slim.batch_norm
, normalizer_params = batch_norm_params
, scope='fc4')
net = slim.dropout(net, keep_prob=0.7, is_training=is_training, scope='dr')
out = slim.fully_connected(net, n_classes
, activation_fn=None, normalizer_fn=None, scope='fco')
return out
print ("NETWORK READY")
# -
# ## 定义图结构
# +
# PREDICTION
pred = CNN(x, is_training)
# LOSS AND OPTIMIZER
cost = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(
labels=y, logits=pred))
optm = tf.train.AdamOptimizer(learning_rate=0.001).minimize(cost)
corr = tf.equal(tf.argmax(pred, 1), tf.argmax(y, 1))
accr = tf.reduce_mean(tf.cast(corr, "float"))
# INITIALIZER
init = tf.global_variables_initializer()
sess = tf.Session()
sess.run(init)
print ("FUNCTIONS READY")
# -
# ## 检查变量
print ("=================== TRAINABLE VARIABLES ===================")
t_weights = tf.trainable_variables()
var_names_list = [v.name for v in tf.trainable_variables()]
for i in range(len(t_weights)):
wval = sess.run(t_weights[i])
print ("[%d/%d] [%s] / SAHPE IS %s"
% (i, len(t_weights), var_names_list[i], wval.shape,))
# ## 存储
savedir = "nets/cnn_mnist_modern/"
saver = tf.train.Saver(max_to_keep=100)
save_step = 4
if not os.path.exists(savedir):
os.makedirs(savedir)
print ("SAVER READY")
# ## 数据增加
def augment_img(xs):
out = np.copy(xs)
xs_r = np.reshape(xs, [-1, 28, 28])
for i in range(xs_r.shape[0]):
xs_img = xs_r[i, :, :]
bg_value = 0
# ROTATE
angle = np.random.randint(-15, 15, 1).astype(float)
xs_img = ndimage.rotate(xs_img, angle, reshape=False, cval=bg_value)
# ZOOM
rg = 0.1
zoom_factor = np.random.uniform(1., 1.+rg)
h, w = xs_img.shape[:2]
zh = int(np.round(zoom_factor * h))
zw = int(np.round(zoom_factor * w))
top = (zh - h) // 2
left = (zw - w) // 2
zoom_tuple = (zoom_factor,) * 2 + (1,) * (xs_img.ndim - 2)
temp = ndimage.zoom(xs_img[top:top+zh, left:left+zw], zoom_tuple)
trim_top = ((temp.shape[0] - h) // 2)
trim_left = ((temp.shape[1] - w) // 2)
xs_img = temp[trim_top:trim_top+h, trim_left:trim_left+w]
# SHIFT
shift = np.random.randint(-3, 3, 2)
xs_img = ndimage.shift(xs_img, shift, cval=bg_value)
# RESHAPE
xs_v = np.reshape(xs_img, [1, -1])
out[i, :] = xs_v
return out
# ## 测试增加
naug = 2
batch_xs = trainimg[:naug, :]
xs2 = augment_img(batch_xs)
for i in range(naug):
x1 = batch_xs[i, :].reshape([28, 28])
x2 = xs2[i, :].reshape([28, 28])
plt.matshow(x1, vmin=0, vmax=1, cmap=plt.cm.gray)
plt.title("ORIGINAL")
plt.show()
plt.matshow(x2, vmin=0, vmax=1, cmap=plt.cm.gray)
plt.title("TRANSFORMED")
plt.show()
# ## 运行
# PARAMETERS
training_epochs = 100
batch_size = 50
display_step = 4
val_acc = 0
val_acc_max = 0
# OPTIMIZE
for epoch in range(training_epochs):
avg_cost = 0.
total_batch = int(mnist.train.num_examples/batch_size)
# ITERATION
for i in range(total_batch):
batch_xs, batch_ys = mnist.train.next_batch(batch_size)
# AUGMENT DATA
batch_xs = augment_img(batch_xs)
feeds = {x: batch_xs, y: batch_ys, is_training: True}
sess.run(optm, feed_dict=feeds)
avg_cost += sess.run(cost, feed_dict=feeds)
avg_cost = avg_cost / total_batch
# DISPLAY
if (epoch+1) % display_step == 0:
print ("Epoch: %03d/%03d cost: %.9f" % (epoch+1, training_epochs, avg_cost))
randidx = np.random.permutation(trainimg.shape[0])[:500]
feeds = {x: trainimg[randidx], y: trainlabel[randidx], is_training: False}
train_acc = sess.run(accr, feed_dict=feeds)
print (" TRAIN ACCURACY: %.5f" % (train_acc))
feeds = {x: valimg, y: vallabel, is_training: False}
val_acc = sess.run(accr, feed_dict=feeds)
print (" VALIDATION ACCURACY: %.5f" % (val_acc))
# SAVE
if (epoch+1) % save_step == 0:
savename = savedir + "net-" + str(epoch) + ".ckpt"
saver.save(sess=sess, save_path=savename)
print (" [%s] SAVED." % (savename))
# MAXIMUM VALIDATION ACCURACY
if val_acc > val_acc_max:
val_acc_max = val_acc
best_epoch = epoch
print ("\x1b[31m BEST EPOCH UPDATED!! [%d] \x1b[0m" % (best_epoch))
print ("OPTIMIZATION FINISHED")
# ## 计算测试的精确度
best_epoch = 55
restorename = savedir + "net-" + str(best_epoch) + ".ckpt"
print ("LOADING [%s]" % (restorename))
saver.restore(sess, restorename)
feeds = {x: testimg, y: testlabel, is_training: False}
test_acc = sess.run(accr, feed_dict=feeds)
print ("TEST ACCURACY: %.5f" % (test_acc))
| 04_CNN_advances/cnn_mnist_modern.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/claubermartins/Stock-market-prediction/blob/main/stock_market_prediction-Vale.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="XjlKRgiDRdpq" outputId="500ff1c8-936c-48ff-ab27-ffbcbfe38d2a"
pip install inflection
# + [markdown] id="UiXZAchqFuC-"
# #**1-Importando bibliotecas**
# + id="ZjQg-8c_--wP"
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Dropout, LSTM
from tensorflow.keras.callbacks import EarlyStopping
from sklearn.preprocessing import MinMaxScaler
from sklearn.impute import SimpleImputer
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import tensorflow as tf
import math
import seaborn as sns
import datetime
import inflection
from IPython.core.display import HTML
# + [markdown] id="SuPg1RFvQyMS"
# #**2-Funções auxiliares para o Jupyter Notebook**
# + id="2vKvPdvDPJHc"
def jupyter_settings():
# %matplotlib inline
# %pylab inline
plt.style.use( 'bmh' )
plt.rcParams['figure.figsize'] = [25, 12]
plt.rcParams['font.size'] = 24
display( HTML( '<style>.container { width:100% !important; }</style>') )
pd.options.display.max_columns = None
pd.options.display.max_rows = None
pd.set_option( 'display.expand_frame_repr', False )
sns.set()
# + colab={"base_uri": "https://localhost:8080/", "height": 34} id="PGJn-AaoMxuy" outputId="015e7b6d-8d99-4ccf-fdff-06f008114742"
jupyter_settings()
# + [markdown] id="c_3core8Shq6"
# ##**2.1Método de importação de arquivos do Google Colab**
# + id="cTk_YEVdyms3" colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY>", "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": ""}}, "base_uri": "https://localhost:8080/", "height": 157} outputId="9aadc800-ee92-477b-f7ac-cc37606afb6e"
from google.colab import files
upload = files.upload()
print("downloaded files: ")
print(*upload, sep = "\n")
# + [markdown] id="8MLClRMWPwTL"
# #**3-Tratamento dos dados**
# + [markdown] id="DOJanakhS-s2"
# ##**3.1-Importando os dados**
# + id="vYFCwA5GTB2j"
base = pd.read_csv('VALE-treinamento.csv')
base = base.drop('Date', axis = 1)
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="SD9oSvK-NE0Y" outputId="26cf9702-c356-4fd0-db49-1017ebb5b075"
#valores estatísticos
base.describe()
# + colab={"base_uri": "https://localhost:8080/"} id="fbKUPwPSNJqm" outputId="371a05b0-729c-4237-eaf8-7dc593b81348"
base.dtypes
# + [markdown] id="bCTD3wo2TfZF"
# ##**3.2-Tratando os Valores Faltantes**
# + colab={"base_uri": "https://localhost:8080/"} id="H60SOo8MTjjP" outputId="bb287c13-55d3-48fb-98ab-49440d2dcb02"
#valores faltantes
base.isna().sum()
# + id="H-DMfyHXTkdi" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="9728bf1a-7a95-4299-d110-3a8362c43567"
#encontrando onde os valores faltantes estão
base.loc[pd.isnull(base['Open'])]
base.loc[pd.isnull(base['High'])]
base.loc[pd.isnull(base['Low'])]
base.loc[pd.isnull(base['Close'])]
base.loc[pd.isnull(base['Adj Close'])]
base.loc[pd.isnull(base['Volume'])]
# + id="6q0YFzy9TmPD"
#preenchendo os valores nulos com o valor médio
imputer = SimpleImputer(missing_values=np.nan, strategy='mean',verbose=0)
imputer = imputer.fit(base.iloc[:,0:6])
base.iloc[:,0:6] = imputer.transform(base.iloc[:,0:6])
# + id="gQC2HfFnTpe2" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="d6c80f55-9462-4fb1-9625-de91518b9a8c"
#verificando novamente se ainda temos valores faltantes
base.loc[pd.isnull(base['Open'])]
base.loc[pd.isnull(base['High'])]
base.loc[pd.isnull(base['Low'])]
base.loc[pd.isnull(base['Close'])]
base.loc[pd.isnull(base['Adj Close'])]
base.loc[pd.isnull(base['Volume'])]
# + [markdown] id="_WuI_UbmTt3W"
# ##**3.3-Tratamento dos valores nulos (iguais a 0)**
# + colab={"base_uri": "https://localhost:8080/"} id="hyRiBN32Twb2" outputId="49c2e3d5-3b23-45de-d1bc-79e2eebcf020"
#valores nulos
(base == 0).sum()
# + id="nQPTB9_TT3Dy" outputId="156e4d9c-d1f0-4085-bcb2-26f0858470a9" colab={"base_uri": "https://localhost:8080/", "height": 0}
#Encontrando os valores nulos
base.loc[base['Volume'] == 0]
# + id="352jPyV0T5JJ"
#preenchendo os valores nulos com o valor médio
imputer = SimpleImputer(missing_values=0, strategy='mean',verbose=0)
imputer = imputer.fit(base.iloc[:,0:6])
base.iloc[:,0:6] = imputer.transform(base.iloc[:,0:6])
# + id="xI5JdDykT66_" outputId="de4b057a-8a7e-442f-d33f-deb0d3de181f" colab={"base_uri": "https://localhost:8080/", "height": 0}
#Verificando novamente se ainda temos valores nulos
base.loc[base['Volume'] == 0]
# + id="NYW0UUz0T9Kq"
#base = base.dropna()
# + id="ceOotOwsT_Er"
#base.sample()
# + [markdown] id="hCiQYMqPGHuG"
# #**-Descrição dos dados** IGNORE ESTA CÉLULA
# + id="xalwLdRSGLRD"
#Tendencia central - mean, median
ct1 = pd.DataFrame(base.apply(np.mean)).T #T transpondo
ct2 = pd.DataFrame(base.apply(np.median)).T
#Dispersão - std(desvio padrão), min, max, range
d1 = pd.DataFrame(base.apply(np.std)).T
d2 = pd.DataFrame(base.apply(min)).T
d3 = pd.DataFrame(base.apply(max)).T
d4 = pd.DataFrame(base.apply(lambda x: x.max() - x.min())).T
#concatenate
m = pd.concat([d2,d3,d4,ct1,ct2,d1]).T.reset_index()
m.columns = ['attributes','min','max','range', 'mean', 'median', 'std']
m
# + colab={"base_uri": "https://localhost:8080/"} id="j62Bj8jOQz_G" outputId="9e5055ec-b49f-4f7d-e3c1-b3fc2b1dae42"
base.shape[0]
# + [markdown] id="4IDwP-XV_Ohd"
# #**4-Treinamento da rede**
# + id="mpo6RlDsUHNJ"
#normalizando a base de dados com o MinMaxScaler
base_treinamento = base.iloc[:, 0:6].values
normalizador = MinMaxScaler(feature_range=(0,1))
base_treinamento_normalizada = normalizador.fit_transform(base_treinamento)
# + id="d-LD4OEaUNf7"
#Comparando a predição com o preço real
previsores = []
preco_real = []
for i in range(90, 2264):
previsores.append(base_treinamento_normalizada[i-90:i, 0:6])
preco_real.append(base_treinamento_normalizada[i, 3:4])
previsores, preco_real = np.array(previsores), np.array(preco_real)
# + colab={"base_uri": "https://localhost:8080/"} id="ZftXowA9URdt" outputId="bc80726f-9040-4a40-9e49-25cef394dbf3"
#estrutura da rede neural
regressor = Sequential()
regressor.add(LSTM(units = 90, return_sequences = True, input_shape = (previsores.shape[1], 6)))
regressor.add(Dropout(0.3))
regressor.add(LSTM(units = 90, return_sequences = True))
regressor.add(Dropout(0.3))
regressor.add(LSTM(units = 90, return_sequences = True))
regressor.add(Dropout(0.3))
regressor.add(LSTM(units = 90))
regressor.add(Dropout(0.3))
regressor.add(Dense(units = 1, activation = 'sigmoid'))
es = EarlyStopping(monitor = 'loss', min_delta = 1e-15, patience = 20, verbose = 1)
#regressor.compile(optimizer = 'RMSprop', loss = 'mean_squared_error',
# metrics = ['mean_absolute_error'])
opt = tf.keras.optimizers.Adam(amsgrad=True)
regressor.compile(optimizer = opt, loss = 'mean_squared_error',
metrics = ['mean_absolute_error'])
#opt = tf.keras.optimizers.RMSprop(centered=True)
#regressor.compile(optimizer = opt, loss = 'mean_squared_error',
# metrics = ['mean_absolute_error'])
regressor.fit(previsores, preco_real, epochs = 600, batch_size = 32,
callbacks = [es])
#regressor.fit(previsores, preco_real, epochs = 600, batch_size = 32)
# + [markdown] id="RtFahmEgGAzZ"
# #**5-Teste da rede**
# + [markdown] id="QSWSKW8nqeA0"
# ##**5.1-Importando os dados para teste**
# + id="4X6-ArtKUdpD"
#criando a base de dados teste
base_teste = pd.read_csv('VALE-teste.csv')
base_teste = base_teste.drop('Date', axis = 1)
# + colab={"base_uri": "https://localhost:8080/", "height": 297} id="dXOp2vDcpAvA" outputId="5f7610e9-51ea-4e09-c120-e45cbfb7ac1d"
#valores estatísticos
base_teste.describe()
# + [markdown] id="Cw-NIb9EpatP"
# ##**5.2-Tratando os Valores Faltantes**
# + colab={"base_uri": "https://localhost:8080/"} id="lthEX_TZniHb" outputId="de2f6a54-ac11-41c4-fba7-1b8c511fd00a"
#valores faltantes
base_teste.isna().sum()
# + colab={"base_uri": "https://localhost:8080/", "height": 49} id="uiH2j_UNpjqk" outputId="37c942a3-086d-4f16-9d86-54a122aba413"
#encontrando onde os valores faltantes estão
base_teste.loc[pd.isnull(base_teste['Open'])]
base_teste.loc[pd.isnull(base_teste['High'])]
base_teste.loc[pd.isnull(base_teste['Low'])]
base_teste.loc[pd.isnull(base_teste['Close'])]
base_teste.loc[pd.isnull(base_teste['Adj Close'])]
base_teste.loc[pd.isnull(base_teste['Volume'])]
# + id="Bbw7axJ75lpn"
#preenchendo os valores faltantes com o valor médio
imputer = SimpleImputer(missing_values=np.nan, strategy='mean',verbose=0)
imputer = imputer.fit(base_teste.iloc[:,0:6])
base_teste.iloc[:,0:6] = imputer.transform(base_teste.iloc[:,0:6])
# + colab={"base_uri": "https://localhost:8080/", "height": 49} id="CXAi4QV8pw2x" outputId="999ab80f-635b-4772-f3e3-2d9470d6d8e6"
#verificando novamente se ainda temos valores faltantes
base_teste.loc[pd.isnull(base_teste['Open'])]
base_teste.loc[pd.isnull(base_teste['High'])]
base_teste.loc[pd.isnull(base_teste['Low'])]
base_teste.loc[pd.isnull(base_teste['Close'])]
base_teste.loc[pd.isnull(base_teste['Adj Close'])]
base_teste.loc[pd.isnull(base_teste['Volume'])]
# + [markdown] id="X1nZpS8Rp2zr"
# ##**5.3-Tratamento dos valores nulos (iguais a 0)**
# + colab={"base_uri": "https://localhost:8080/"} id="XYwer-dip-KB" outputId="6620bddb-138e-40a1-db87-7a88d5365213"
#valores nulos
(base_teste == 0).sum()
# + colab={"base_uri": "https://localhost:8080/", "height": 49} id="sFBo6iR5qEpS" outputId="9b750cef-a7d7-4a45-ffaa-341a3db1aefd"
#Encontrando os valores nulos
base_teste.loc[base_teste['Volume'] == 0]
# + id="ofkU-XjcqOQE"
#preenchendo os valores nulos com o valor médio
imputer = SimpleImputer(missing_values=0, strategy='mean',verbose=0)
imputer = imputer.fit(base_teste.iloc[:,0:6])
base_teste.iloc[:,0:6] = imputer.transform(base_teste.iloc[:,0:6])
# + colab={"base_uri": "https://localhost:8080/", "height": 49} id="pYUeIwbnppRT" outputId="fa47fa8e-9a19-452a-f15f-bb5c72a5f22a"
#Verificando novamente se tem valores nulos
base_teste.loc[base_teste['Volume'] == 0]
# + [markdown] id="N79Bjk6Yq6ci"
# ##**5.4-Preparando a estrutura para teste**
# + id="UFINH9OzUfDk"
#concatenando a base de dados teste com o base de treinamento
preco_real_teste = base_teste.iloc[:, 3:4].values
frames = [base, base_teste]
base_completa = pd.concat(frames)
# + id="-CdLW8IZUhLY"
#pegando os 90 registros anteriores para percorrer a base teste e colocando no
#no formato np.array
entradas = base_completa[len(base_completa) - len(base_teste) - 90:].values
entradas = normalizador.transform(entradas)
Percorrer_teste = []
for i in range(90, 342):
Percorrer_teste.append(entradas[i-90:i, 0:6])
Percorrer_teste = np.array(Percorrer_teste)
# + id="_YdD2ZcaUi1w"
#resultado da predição no formato MinMaxScaler
previsoes = regressor.predict(Percorrer_teste)
# + colab={"base_uri": "https://localhost:8080/"} id="jQS0Ql4tUk2Z" outputId="b17494e1-de89-403b-90f0-dc5338bdb1f6"
#convertendo para a escala MinMaxScaler
normalizador_previsao = MinMaxScaler(feature_range=(0,1))
normalizador_previsao.fit_transform(base_treinamento[:,3:4])
# + id="MP_GBjYJUnFe" colab={"base_uri": "https://localhost:8080/"} outputId="95a8cdf5-95cf-43d5-fbc7-4c6a5d4ef953"
#colocando a previsão no formato original para preparar a visualização no gráfico
previsoes = normalizador_previsao.inverse_transform(previsoes)
previsoes
# + [markdown] id="lEjN0sRdrGmK"
# ##**6-Análise dos resuldatos**
# + id="beWuUgy2Uoe_" colab={"base_uri": "https://localhost:8080/", "height": 485} outputId="aeaec3e3-8e46-4e19-86e1-dcd99e616ca5"
#Visualizando o gráfico da Previsão da rede
plt.plot(preco_real_teste, color = 'red', label = 'Preço real')
plt.plot(previsoes, color = 'blue', label = 'Previsões')
plt.title('Previsão do preço das ações')
plt.xlabel('Tempo em dias')
plt.ylabel('Preço das ações')
plt.legend()
plt.show()
# + id="hKA0-lbgUtH-" colab={"base_uri": "https://localhost:8080/", "height": 297} outputId="d2c91287-96dd-4d6a-b3f0-e609286a52fe"
#estatísticas do preço real comparado com a previsão
df1 = pd.DataFrame(preco_real_teste).T
df2 = pd.DataFrame(previsoes).T
df3 = pd.concat([df1,df2]).T
df3.columns = ['Preco real', 'Previsoes']
df3.describe()
# + id="ZjIrFF8Y_M9N"
#df2 = pd.DataFrame(previsoes)
#df2.describe()
# + id="TKfgSJAIV5a_"
#df1 = pd.DataFrame(preco_real_teste)
#df1.describe()
| stock_market_prediction-Vale.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="NWmug8xIJE5G" colab_type="text"
# # Importing Project Dependencies
# + id="bX1QOttjSGoC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="ad99d280-4bb6-43e7-88a8-b0068a7babc2"
from google.colab import drive
drive.mount('/content/drive')
# + id="ZBUudD2tSRLw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 250} outputId="d63d6aa2-52da-4c15-f6f3-ae5e9d0450ed"
# !pip install glove_python
# + id="gAPvxNABpQF_" colab_type="code" colab={}
import numpy as np
# + id="sJVHzimipSGp" colab_type="code" colab={}
import json
import glob
import pandas as pd
# + id="unZhEiI1pu7E" colab_type="code" colab={}
from gensim.models import Word2Vec
# + id="sC-PH0iFq2RS" colab_type="code" colab={}
import pickle
# + [markdown] id="ofyUOvvLos37" colab_type="text"
# # Custom Word Embedding generation
# + [markdown] id="HzEHBoI_JmFC" colab_type="text"
# ## Loading pre-trained Glove Vector Embeddings
# + id="O1EaDj2hVSmA" colab_type="code" colab={}
glove_t = '/content/drive/My Drive/Practicum/glove.twitter.27B.100d.txt'
# + id="A23Bojyro8nh" colab_type="code" colab={}
with open(glove_t, "rb") as lines:
wvec = {line.split()[0].decode('utf-8'): np.array(line.split()[1:], dtype=np.float32) for line in lines}
# + [markdown] id="0V0xm0kLJt-u" colab_type="text"
# ## Loading all reviews
# + id="yRLm_m_FpiaX" colab_type="code" colab={}
path = r'/content/drive/My Drive/Practicum/DepTag'
files = glob.glob(path+r'/*.json')
# + id="7DH-2qioprcB" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="82133bbb-9269-402a-db99-044e275f7859"
len(files)
# + id="CgQJuVHDpsDx" colab_type="code" colab={}
finInfo = []
for i in range(len(files)):
for line in open(files[i]):
info = json.loads(line)
finInfo.append(info)
# + id="2J6BTlO1ptYY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="13189401-4a50-43cc-86fc-578d4f0a6b31"
len(finInfo)
# + id="u1LcnArVpuPT" colab_type="code" colab={}
review = []
for i in range(len(finInfo)):
for j in range(len(finInfo[i])):
sents = finInfo[i][j]['sentence']
for sent in sents:
review.append(sent)
# + [markdown] id="GIzY_1pCJw4e" colab_type="text"
# ## Defining a word2vec model
# + id="Exa6ZQoKpwhf" colab_type="code" colab={}
em_model = Word2Vec(review, size=100, window=5, min_count=1, workers=2)
# + id="zFoyQsg0p9oi" colab_type="code" colab={}
w2v = {w: vec for w, vec in zip(em_model.wv.index2word, em_model.wv.vectors)}
# + id="akiMiYfxqAWR" colab_type="code" colab={}
a = list(w2v.keys())
# + [markdown] id="lKngreofJ0vY" colab_type="text"
# #### Updating word2vec if embedding not present in pre-trained vectors
# + id="TLqUnU7QqC1S" colab_type="code" colab={}
for i in a:
if i in wvec:
continue
else:
wvec.update({ i : w2v[i]})
# + id="KRf3m07NqHfx" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="825a3c06-92fa-4848-df43-5badec9a5b26"
len(list(wvec.keys()))
# + id="reU0aAuoqI9x" colab_type="code" colab={}
import scipy
from scipy import spatial
# + id="pgjUKwd3sgdL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="c62eb4b0-10f2-41c3-c85f-64d8ba5aac69"
type(wvec)
# + [markdown] id="Y1Jf6ZRSKCe1" colab_type="text"
# # Storing the word embedding model
# + id="nlCst1CTswWd" colab_type="code" colab={}
pickleobj = open("/content/drive/My Drive/Practicum/embedding.pickle", "wb")
pickle.dump(wvec, pickleobj)
# + id="1IFke5mmtXo3" colab_type="code" colab={}
pickleobj.close()
# + [markdown] id="pbgS5euUvUbS" colab_type="text"
# # Calculating Cosine Distances
# + id="uy5IaTvburZm" colab_type="code" colab={}
import pickle
# + id="4yNfWqw3vYU9" colab_type="code" colab={}
pickle_obj = open("/content/drive/My Drive/Practicum/embedding.pickle", "rb")
wvec = pickle.load(pickle_obj)
pickle_obj.close()
# + id="vumL764vw50g" colab_type="code" colab={}
def cosine_distance_between_two_words(word1, word2):
'''
Takes input of words and returns the cosine distance of each word.
'''
return (1- scipy.spatial.distance.cosine(wvec[word1], wvec[word2]))
# + id="vVndcySBw7A-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="28c5fbee-6ea5-479f-9b90-4f81228c5202"
cosine_distance_between_two_words('food', 'delicious') #first word will be main word, 2nd word can be DepWord, PreviousWord, NextWord
# + [markdown] id="I7gsuNUtKQis" colab_type="text"
# ### Plotting a heatmap
# + id="PmQhqDKEvpgs" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 72} outputId="ac99cbef-5848-41fc-a055-8b2b29cbf6d0"
import matplotlib.pyplot as plt
import seaborn as sns
# + id="Pdpt5c_KvcGT" colab_type="code" colab={}
def cosine_distance_between_two_words(word1, word2):
'''
Takes input of words and returns the cosine distance of each word.
'''
return (1- scipy.spatial.distance.cosine(wvec[word1], wvec[word2]))
def calculate_heat_matrix_for_two_sentences(s1,s2):
'''
This function calculcates heat matrix based on the cosine distances of words from one another.
'''
result_list = [[cosine_distance_between_two_words(word1, word2) for word2 in s2] for word1 in s1]
result_df = pd.DataFrame(result_list)
result_df.columns = s2
result_df.index = s1
return result_df
def cosine_distance_wordembedding_method(s1, s2):
'''
Returns the cosine similarity index.
'''
vector_1 = np.mean([wvec[word] for word in s1],axis=0)
vector_2 = np.mean([wvec[word] for word in s2],axis=0)
cosine = scipy.spatial.distance.cosine(vector_1, vector_2)
print('Word Embedding method with a cosine distance that our two sentences are similar to',round((1-cosine)*100,2),'%')
def heat_map_matrix_between_two_sentences(s1,s2):
'''
Plots the heat matrix.
'''
df = calculate_heat_matrix_for_two_sentences(s1,s2)
fig, ax = plt.subplots(figsize=(5,5))
ax_blue = sns.heatmap(df, cmap="YlGnBu")
# ax_red = sns.heatmap(df)
print(cosine_distance_wordembedding_method(s1, s2))
return ax_blue
# + id="nCM5dicBvnLt" colab_type="code" colab={}
words1 = ['cabin', 'entertainment', 'food', 'inflight', 'flight', 'luggage', 'seat', 'staff']
# + id="1NuQCT5jv9RU" colab_type="code" colab={}
words2 = ['spacious', 'screen', 'delicious', 'service', 'bad', 'lost', 'hard', 'rude']
# + id="VxFvsZiWwHus" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 407} outputId="c7a7f3e0-e8d4-4771-c674-51459c7f1497"
heat_map_matrix_between_two_sentences(words1, words2)
# + id="I7OtS3VEwKFM" colab_type="code" colab={}
| src/Word Embeddings/Custom_embeddings_with_pretrained.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 0.4.3
# language: julia
# name: julia-0.4
# ---
# # quant-econ Solutions: Modeling Career Choice
#
# Solutions for http://quant-econ.net/jl/career.html
using Plots
pyplot()
using QuantEcon
include("career.jl")
# +
srand(41) # reproducible results
wp = CareerWorkerProblem()
function solve_wp(wp::CareerWorkerProblem)
v_init = fill(100.0, wp.N, wp.N)
func(x) = bellman_operator(wp, x)
v = compute_fixed_point(func, v_init, max_iter=500, verbose=false)
optimal_policy = get_greedy(wp, v)
return v, optimal_policy
end
v, optimal_policy = solve_wp(wp)
F = DiscreteRV(wp.F_probs)
G = DiscreteRV(wp.G_probs)
function gen_path(T=20)
i = j = 1
theta_ind = Int[]
epsilon_ind = Int[]
for t=1:T
# do nothing if stay put
if optimal_policy[i, j] == 2 # new job
j = QuantEcon.draw(G)[1]
elseif optimal_policy[i, j] == 3 # new life
i, j = QuantEcon.draw(F)[1], QuantEcon.draw(G)[1]
end
push!(theta_ind, i)
push!(epsilon_ind, j)
end
return wp.theta[theta_ind], wp.epsilon[epsilon_ind]
end
n = 2
thetas = []
epsilons = []
for i=1:n
theta_path, epsilon_path = gen_path()
push!(thetas, theta_path)
push!(epsilons, epsilon_path)
end
plot(epsilons, label=["epsilon" ""], layout=grid(2,1), linewidth=2)
plot!(thetas, label=["theta" ""], linewidth=2, ylims=(0, 5))
# -
# ## Exercise 2
#
# The median for the original parameterization can be computed as follows
# +
function gen_first_passage_time(optimal_policy::Matrix)
t = 0
i = j = 1
while true
if optimal_policy[i, j] == 1 # Stay put
return t
elseif optimal_policy[i, j] == 2 # New job
j = QuantEcon.draw(G)[1]
else # New life
i, j = QuantEcon.draw(F)[1], QuantEcon.draw(G)[1]
end
t += 1
end
end
M = 25000
samples = Array(Float64, M)
for i=1:M
samples[i] = gen_first_passage_time(optimal_policy)
end
print(median(samples))
# -
# To compute the median with $\beta=0.99$ instead of the default value $\beta=0.95$,
# replace `wp = CareerWorkerProblem()` with `wp = CareerWorkerProblem(beta=0.99)`
#
# The medians are subject to randomness, but should be about 7 and 11
# respectively. Not surprisingly, more patient workers will wait longer to settle down to their final job
# ## Exercise 3
#
# Here’s the code to reproduce the original figure
# +
function region_plot(wp::CareerWorkerProblem)
v, optimal_policy = solve_wp(wp)
region_plot(optimal_policy)
end
function region_plot(optimal_policy::Matrix)
lvls = [0.5, 1.5, 2.5, 3.5]
contour(collect(wp.theta), collect(wp.epsilon),
optimal_policy', fill=true, levels=lvls,
alpha=0.5, cbar=false)
xlabel!("theta")
ylabel!("epsilon")
annotate!([(1.8, 2.5,text("new life")),
(4.5, 2.5, text("new job")),
(4.5, 4.5, text("stay put", :white))])
end
region_plot(optimal_policy)
# -
# Now we want to set `G_a = G_b = 100` and generate a new figure with these parameters.
#
# To do this replace:
#
# wp = CareerWorkerProblem()
#
# with:
#
# wp = CareerWorkerProblem(G_a=100, G_b=100)
#
# In the new figure, you will see that the region for which the worker will stay put has grown because the distribution for $\epsilon$ has become more concentrated around the mean, making high-paying jobs less realistic
#
region_plot(CareerWorkerProblem(G_a=100, G_b=100))
| career/career_solutions_jl.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/gdollp/kagglebook/blob/master/submit2.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" papermill={"duration": 0.024709, "end_time": "2021-06-25T22:18:33.032989", "exception": false, "start_time": "2021-06-25T22:18:33.008280", "status": "completed"} tags=[] id="resident-birthday"
# This notebook uses lightGBM to make predictions.
#
# We use the following features
# * playerId
# * position
# * teamId(rosters)
# * status(rosters)
# * playerBoxScores
#
# and the date 20200401~20200431 as the validation data.
#
# But I think there is room for improvement.
# If you have better ways, I would appreciate it if you could comment on it.
#
# このnotebookではlightGBMを使って予測します。
#
# 特徴量は以下のものを使用しています。
# * playerId
# * position
# * teamId(rosters)
# * status(rosters)
# * playerBoxScores
#
# 20200401~20200431を日時をvalidation dataとしていますが、一考の余地がありそうです。
# もし良さそうな方法があればコメントしていただけると幸いです。
# + [markdown] papermill={"duration": 0.027024, "end_time": "2021-06-25T22:18:33.083611", "exception": false, "start_time": "2021-06-25T22:18:33.056587", "status": "completed"} tags=[] id="vital-water"
#
# + [markdown] papermill={"duration": 0.025051, "end_time": "2021-06-25T22:18:33.139292", "exception": false, "start_time": "2021-06-25T22:18:33.114241", "status": "completed"} tags=[] id="operating-thunder"
# https://www.kaggle.com/columbia2131/mlb-lightgbm-starter-dataset-code-en-ja
# + [markdown] papermill={"duration": 0.026534, "end_time": "2021-06-25T22:18:33.190839", "exception": false, "start_time": "2021-06-25T22:18:33.164305", "status": "completed"} tags=[] id="theoretical-damages"
# ## About Dataset
# + [markdown] papermill={"duration": 0.02311, "end_time": "2021-06-25T22:18:33.237635", "exception": false, "start_time": "2021-06-25T22:18:33.214525", "status": "completed"} tags=[] id="clinical-cheat"
# Train.csv is stored as a csv file with each column as follows.
#
# train.csvを以下のようにして各カラムをcsvファイルとして保管しています。
# + papermill={"duration": 0.042271, "end_time": "2021-06-25T22:18:33.303166", "exception": false, "start_time": "2021-06-25T22:18:33.260895", "status": "completed"} tags=[] id="reasonable-assistant"
# %%capture
"""
!pip install pandarallel
import gc
import numpy as np
import pandas as pd
from pathlib import Path
from pandarallel import pandarallel
pandarallel.initialize()
BASE_DIR = Path('../input/mlb-player-digital-engagement-forecasting')
train = pd.read_csv(BASE_DIR / 'train.csv')
null = np.nan
true = True
false = False
for col in train.columns:
if col == 'date': continue
_index = train[col].notnull()
train.loc[_index, col] = train.loc[_index, col].parallel_apply(lambda x: eval(x))
outputs = []
for index, date, record in train.loc[_index, ['date', col]].itertuples():
_df = pd.DataFrame(record)
_df['index'] = index
_df['date'] = date
outputs.append(_df)
outputs = pd.concat(outputs).reset_index(drop=True)
outputs.to_csv(f'{col}_train.csv', index=False)
outputs.to_pickle(f'{col}_train.pkl')
del outputs
del train[col]
gc.collect()
"""
# + colab={"base_uri": "https://localhost:8080/"} id="APDXL7SxlCT1" outputId="1c51bf95-5665-44bb-ebae-300cbe8cafca"
# !pip install optuna
# + [markdown] papermill={"duration": 0.025072, "end_time": "2021-06-25T22:18:33.352190", "exception": false, "start_time": "2021-06-25T22:18:33.327118", "status": "completed"} tags=[] id="frozen-alfred"
# ## Training
# + papermill={"duration": 2.967463, "end_time": "2021-06-25T22:18:36.344129", "exception": false, "start_time": "2021-06-25T22:18:33.376666", "status": "completed"} tags=[] id="concrete-cruise"
import numpy as np
import pandas as pd
from pathlib import Path
from sklearn.metrics import mean_absolute_error
from datetime import timedelta
from functools import reduce
from tqdm import tqdm
#import optuna.integration.lightgbm as lgbm
import lightgbm as lgbm
#import mlb
import pickle
from datetime import datetime as dt
import copy
import gc
from sklearn.decomposition import PCA
import math
from sklearn.model_selection import KFold
# + colab={"base_uri": "https://localhost:8080/"} id="ykU9DkvWXkRB" outputId="3a53d9d3-03de-4141-9f06-86b5c794cd25"
from google.colab import drive
drive.mount('/content/drive')
# + papermill={"duration": 0.032131, "end_time": "2021-06-25T22:18:36.401524", "exception": false, "start_time": "2021-06-25T22:18:36.369393", "status": "completed"} tags=[] id="declared-principal"
BASE_DIR = Path('/content/drive/MyDrive/mlb/input')
TRAIN_DIR = Path('/content/drive/MyDrive/mlb/input/archive')
# + papermill={"duration": 18.911828, "end_time": "2021-06-25T22:18:55.338134", "exception": false, "start_time": "2021-06-25T22:18:36.426306", "status": "completed"} tags=[] id="tough-launch"
players = pd.read_csv(BASE_DIR / 'players.csv')
rosters = pd.read_pickle(TRAIN_DIR / 'rosters_train.pkl')
targets = pd.read_pickle(TRAIN_DIR / 'nextDayPlayerEngagement_train.pkl')
scores1 = pd.read_pickle(TRAIN_DIR / 'playerBoxScores_train.pkl')
scores = scores1.groupby(['playerId', 'date']).sum().reset_index()
twitter = pd.read_pickle("/content/drive/MyDrive/mlb/input/archive/playerTwitterFollowers_train.pkl")
games = pd.read_pickle(TRAIN_DIR / 'games_train.pkl')
events = pd.read_pickle(TRAIN_DIR / 'events_train.pkl')
standings = pd.read_pickle(TRAIN_DIR / 'standings_train.pkl')
teamtwitter = pd.read_pickle(TRAIN_DIR / 'teamTwitterFollowers_train.pkl')
transaction = pd.read_pickle(TRAIN_DIR / 'teamTwitterFollowers_train.pkl')
awards = pd.read_csv(BASE_DIR / 'awards.csv')
seasons = pd.read_csv(BASE_DIR / 'seasons.csv')
teams = pd.read_csv(BASE_DIR / 'teams.csv')
player_target_stats = pd.read_csv("/content/drive/MyDrive/mlb/input/player_target_stats.csv")
example_test = pd.read_csv("/content/drive/MyDrive/mlb/input/example_test.csv")
# + id="4BRMq9l8YfYI" colab={"base_uri": "https://localhost:8080/"} outputId="07965296-9dd4-4972-af4e-0b512cbf1a39"
def unpack_json(json_str):
return np.nan if pd.isna(json_str) else pd.read_json(json_str)
test = unpack_json(example_test["rosters"].iloc[0])
test["playerId"]
del example_test
gc.collect()
# + papermill={"duration": 0.065838, "end_time": "2021-06-25T22:18:55.429265", "exception": false, "start_time": "2021-06-25T22:18:55.363427", "status": "completed"} tags=[] id="joined-traffic"
awards2 = awards.groupby("playerId").count()
awards2 = awards2.reset_index()
# + id="dRXO92-ROJ2_"
teamtwitter["teamnumberOfFollowers"] = teamtwitter["numberOfFollowers"]
teamtwi = teamtwitter.groupby("teamId").mean()["teamnumberOfFollowers"].reset_index()
# + colab={"base_uri": "https://localhost:8080/", "height": 626} id="M8Mvi9up7ypT" outputId="ca12b206-d4d6-4e47-f924-f5da50753297"
players
# + papermill={"duration": 0.105303, "end_time": "2021-06-25T22:18:55.560260", "exception": false, "start_time": "2021-06-25T22:18:55.454957", "status": "completed"} tags=[] id="saved-assignment"
games = pd.read_pickle(TRAIN_DIR / 'games_train.pkl')
#games.index[games["detailedGameState"] == "Postponed"].shape
#games.drop(games.loc[games['detailedGameState']=='Postponed'].index, inplace=True)
#games["detailedGameState"].unique()
#games[games.duplicated(subset=["gamePk"], keep=False)]
#games['detailedGameState']=='Postponed']だとスコアはキロクされていない
# + id="s5mtlhAlBjJw" colab={"base_uri": "https://localhost:8080/"} outputId="05ce404e-8367-42d2-ddbf-fd075a1bbddb"
targets["engagementMetricsDate"] = targets["engagementMetricsDate"].str.replace('-', '')
yesterday_targets = targets.drop('date', axis=1)
yesterday_targets = yesterday_targets.rename(columns={'engagementMetricsDate':'date', 'target1': 'yest_target1','target2': 'yest_target2','target3': 'yest_target3','target4': 'yest_target4'})
yesterday_targets["date"] = yesterday_targets["date"].astype(int)
yesterday_targets["date"]
# + colab={"base_uri": "https://localhost:8080/"} id="oSBa69W9UBGk" outputId="af15be3f-7e2f-4146-ae25-7ec18f011cb8"
targets_cols = ['playerId', 'target1', 'target2', 'target3', 'target4', 'date']
players_cols = ['playerId', 'primaryPositionName',"playerName","playerForTestSetAndFuturePreds"]
rosters_cols = ['playerId', 'teamId', 'status', 'date']
yesterday_targets_cols = ["date","playerId","yest_target1","yest_target2","yest_target3","yest_target4"]
scores_cols = ['playerId', 'battingOrder', 'gamesPlayedBatting', 'flyOuts',
'groundOuts', 'runsScored', 'doubles', 'triples', 'homeRuns',
'strikeOuts', 'baseOnBalls', 'intentionalWalks', 'hits', 'hitByPitch',
'atBats', 'caughtStealing', 'stolenBases', 'groundIntoDoublePlay',
'groundIntoTriplePlay', 'plateAppearances', 'totalBases', 'rbi',
'leftOnBase', 'sacBunts', 'sacFlies', 'catchersInterference',
'pickoffs', 'gamesPlayedPitching', 'gamesStartedPitching',
'completeGamesPitching', 'shutoutsPitching', 'winsPitching',
'lossesPitching', 'flyOutsPitching', 'airOutsPitching',
'groundOutsPitching', 'runsPitching', 'doublesPitching',
'triplesPitching', 'homeRunsPitching', 'strikeOutsPitching',
'baseOnBallsPitching', 'intentionalWalksPitching', 'hitsPitching',
'hitByPitchPitching', 'atBatsPitching', 'caughtStealingPitching',
'stolenBasesPitching', 'inningsPitched', 'saveOpportunities',
'earnedRuns', 'battersFaced', 'outsPitching', 'pitchesThrown', 'balls',
'strikes', 'hitBatsmen', 'balks', 'wildPitches', 'pickoffsPitching',
'rbiPitching', 'gamesFinishedPitching', 'inheritedRunners',
'inheritedRunnersScored', 'catchersInterferencePitching',
'sacBuntsPitching', 'sacFliesPitching', 'saves', 'holds', 'blownSaves',
'assists', 'putOuts', 'errors', 'chances', 'date',"gamePk"]
games_cols = ["gamePk","homeId","dayNight","seriesDescription","gamesInSeries","homeWinner","awayWinner","homeScore","awayScore","gameType",
"gameDate"]
playertwitter_cols = ["playerId","numberOfFollowers"]
awards_cols = ["playerId","awardName"]
standings_cols = ["date","teamId","divisionRank","divisionLeader","wildCardLeader","leagueRank","divisionId","gameDate"]
teamtwitter_cols = ["teamId","teamnumberOfFollowers"]
targets["engagementMetricsDate"] = targets["engagementMetricsDate"].str.replace('-', '')
yesterday_targets = targets.drop('date', axis=1)
yesterday_targets = yesterday_targets.rename(columns={'engagementMetricsDate':'date', 'target1': 'yest_target1','target2': 'yest_target2','target3': 'yest_target3','target4': 'yest_target4'})
yesterday_targets["date"] = yesterday_targets["date"].astype(int)
yesterday_targets["date"]
# + id="ZBRcuG-4ty69"
feature_cols = ['label_playerId', 'label_primaryPositionName', 'label_teamId',
'label_status', 'battingOrder', 'gamesPlayedBatting', 'flyOuts',
'groundOuts', 'runsScored','homeRuns',
'strikeOuts', 'baseOnBalls', 'hits', 'hitByPitch',
'atBats', 'stolenBases',
'plateAppearances', 'totalBases', 'rbi',
'leftOnBase', 'sacFlies',
'gamesPlayedPitching', 'gamesStartedPitching',
'completeGamesPitching','winsPitching',
'lossesPitching', 'flyOutsPitching', 'airOutsPitching',
'groundOutsPitching', 'runsPitching', 'doublesPitching',
'triplesPitching', 'homeRunsPitching', 'strikeOutsPitching',
'baseOnBallsPitching', 'hitsPitching',
'hitByPitchPitching', 'atBatsPitching', 'caughtStealingPitching',
'stolenBasesPitching', 'inningsPitched', 'saveOpportunities',
'earnedRuns', 'battersFaced', 'outsPitching', 'pitchesThrown', 'balls',
'strikes', 'hitBatsmen', 'balks', 'wildPitches', 'pickoffsPitching',
'rbiPitching', 'gamesFinishedPitching', 'inheritedRunners',
'inheritedRunnersScored',
'sacFliesPitching', 'saves', 'holds',
'assists', 'putOuts', 'errors', 'chances','target1_mean',
'target1_median',
'target1_std',
'target1_min',
'target1_max',
'target1_prob',
'target2_mean',
'target2_median',
'target2_std',
'target2_min',
'target2_max',
'target2_prob',
'target3_mean',
'target3_median',
'target3_std',
'target3_min',
'target3_max',
'target3_prob',
'target4_mean',
'target4_median',
'target4_std',
'target4_min',
'target4_max',
'target4_prob',"divisionId","teamnumberOfFollowers","preseasonhits"]
feature_cols2 = ['label_playerId', 'label_primaryPositionName', 'label_teamId',
'label_status', 'battingOrder', 'gamesPlayedBatting', 'flyOuts',
'groundOuts', 'runsScored',# 'doubles', 'triples', 'homeRuns',
'baseOnBalls', 'hits',
# 'atBats', 'caughtStealing', 'stolenBases', 'groundIntoDoublePlay',
'plateAppearances', 'totalBases', 'rbi',
# 'leftOnBase', 'sacBunts', 'sacFlies', 'catchersInterference',
'pickoffs', 'gamesPlayedPitching', 'gamesStartedPitching',
'winsPitching',
'lossesPitching',# 'flyOutsPitching', 'airOutsPitching',
'runsPitching',
'strikeOutsPitching',
#'hitsPitching',
'hitByPitchPitching', 'caughtStealingPitching',
'stolenBasesPitching', 'inningsPitched',
'battersFaced',
'balks', 'pickoffsPitching',
'inheritedRunners',
#'sacBuntsPitching', 'sacFliesPitching', 'saves', 'holds', 'blownSaves',
'putOuts','chances','target1_mean',
'target1_median',
'target1_std',
'target1_min',
'target1_max',
'target1_prob',
'target2_mean',
'target2_median',
'target2_std',
'target2_min',
'target2_max',
'target2_prob',
'target3_mean',
'target3_median',
'target3_std',
'target3_min',
'target3_max',
'target3_prob',
'target4_mean',
'target4_median',
'target4_std',
'target4_min',
'target4_max',
'target4_prob',
'target1',"divisionId","teamnumberOfFollowers"]
feature_cols3 = ['label_playerId', 'label_primaryPositionName', 'label_teamId',
'label_status', #'gamesPlayedBatting', 'flyOuts',
'homeRuns',
# 'strikeOuts', 'baseOnBalls', 'intentionalWalks', 'hits', 'hitByPitch',
# 'atBats', 'caughtStealing', 'stolenBases', 'groundIntoDoublePlay',
'totalBases', 'rbi',
# 'leftOnBase', 'sacBunts', 'sacFlies', 'catchersInterference',
'gamesStartedPitching',
# 'completeGamesPitching', 'shutoutsPitching', 'winsPitching',
'lossesPitching',
# 'groundOutsPitching', 'runsPitching', 'doublesPitching',
# 'triplesPitching', 'homeRunsPitching', 'strikeOutsPitching',
#'baseOnBallsPitching', 'intentionalWalksPitching', 'hitsPitching',
#'hitByPitchPitching', 'atBatsPitching', 'caughtStealingPitching',
'inningsPitched',
'battersFaced', 'pitchesThrown',
# 'strikes', 'hitBatsmen', 'balks', 'wildPitches', 'pickoffsPitching',
# 'rbiPitching', 'gamesFinishedPitching', 'inheritedRunners',
#'inheritedRunnersScored', 'catchersInterferencePitching',
#'sacBuntsPitching', 'sacFliesPitching', 'saves', 'holds', 'blownSaves',
# 'assists', 'putOuts', 'errors', 'chances',
'target1_mean',
'target1_median',
'target1_std',
'target1_min',
'target1_max',
'target1_prob',
'target2_mean',
'target2_median',
'target2_std',
'target2_min',
'target2_max',
'target2_prob',
'target3_mean',
'target3_median',
'target3_std',
'target3_min',
'target3_max',
'target3_prob',
'target4_mean',
'target4_median',
'target4_std',
'target4_min',
'target4_max',
'target4_prob',
"divisionId","teamnumberOfFollowers","target2","preseasonhits"]
feature_cols4 = ['label_playerId', 'label_primaryPositionName', 'label_teamId',
'label_status', 'battingOrder', 'gamesPlayedBatting', 'flyOuts',
'runsScored','triples','groundIntoDoublePlay',
'strikeOuts', 'baseOnBalls','hits',
'atBats', 'caughtStealing',
'plateAppearances', 'totalBases', 'rbi',
'leftOnBase', 'sacBunts', 'sacFlies',
'gamesPlayedPitching',
'winsPitching',
'airOutsPitching',
'runsPitching', 'doublesPitching',
'triplesPitching', 'homeRunsPitching', 'strikeOutsPitching',
'intentionalWalksPitching', 'hitsPitching',
'hitByPitchPitching', 'atBatsPitching',
'inningsPitched', 'saveOpportunities',
'battersFaced', 'outsPitching', 'pitchesThrown', 'balls',
'strikes','wildPitches',
'rbiPitching', 'gamesFinishedPitching', 'inheritedRunners',
'inheritedRunnersScored',
'sacBuntsPitching', 'holds', 'blownSaves',
'assists', 'putOuts', 'errors', 'chances','target1_mean',
'target1_median',
'target1_std',
'target1_min',
'target1_max',
'target1_prob',
'target2_mean',
'target2_median',
'target2_std',
'target2_min',
'target2_max',
'target2_prob',
'target3_mean',
'target3_median',
'target3_std',
'target3_min',
'target3_max',
'target3_prob',
'target4_mean',
'target4_median',
'target4_std',
'target4_min',
'target4_max',
'target4_prob',
'target1', "divisionId","target2","target3","teamnumberOfFollowers","preseasonhits"]
# + id="WrdnRRfZtzBL"
# + id="e-haSgyI78O6"
twitter["strdate"] = twitter["date"].astype(str)
twitter["year_months"] = twitter["strdate"].str[0:6].astype(int)
# + id="HyLf8gpubQJA"
targets['hasTwitterAccount'] = targets.playerId.isin(twitter.playerId)
# + papermill={"duration": 9.448263, "end_time": "2021-06-25T22:19:05.117741", "exception": false, "start_time": "2021-06-25T22:18:55.669478", "status": "completed"} tags=[] id="unauthorized-stretch"
train = targets[targets_cols].merge(players[players_cols], on=['playerId'], how='left')
train = train.merge(rosters[rosters_cols], on=['playerId', 'date'], how='left')
train = train.merge(scores[scores_cols], on=['playerId', 'date'], how='left')
train = train.merge(games[games_cols], on=["gamePk"], how="left")
train = train.merge(standings[standings_cols], on=['date',"teamId"], how='left')
train = train.merge(awards2[awards_cols], on=['playerId'], how='left')
train = train.merge(player_target_stats, how='inner', left_on=["playerId"],right_on=["playerId"])
train["strdate"] = train["date"].astype(str)
train["year_months"] = train["strdate"].str[0:6].astype(int)
#train = train.merge(twitter[playertwitter_cols], on=['playerId'], how='left')
train = train.merge(teamtwi[teamtwitter_cols], on=['teamId'], how='left')
# + id="mVHPPnE8GSqG"
targets['hasTwitterAccount'] = targets.playerId.isin(twitter.playerId)
# + colab={"base_uri": "https://localhost:8080/"} id="9EexEYL53a9z" outputId="c3fab7cc-11a1-4c40-e2d4-513a1a9a8f9b"
test["playerId"]
# + id="iCSDWeWFAPc9"
train = train[train["playerForTestSetAndFuturePreds"]==True]
# + id="4nTLjBWC6hki" colab={"base_uri": "https://localhost:8080/", "height": 317} outputId="27858a4a-3519-471e-cf9f-7a369a01b2cf"
train[['target1_mean',
'target1_median',
'target1_std',
'target1_min',
'target1_max',
'target1_prob',
'target2_mean',
'target2_median',
'target2_std',
'target2_min',
'target2_max',
'target2_prob',
'target3_mean',
'target3_median',
'target3_std',
'target3_min',
'target3_max',
'target3_prob',
'target4_mean',
'target4_median',
'target4_std',
'target4_min',
'target4_max',
'target4_prob',
'target1',"divisionId","target2","target3"]].describe()
# + id="70LUMHqddn6f" colab={"base_uri": "https://localhost:8080/", "height": 609} outputId="b37dd877-8bc7-4bec-fb00-ba36795466bc"
train[["target1","target2","target3","target4"]].describe()
train[train["target1"]==train["target1"].max()]
train[train["playerId"]==519317]
# + id="vrevhu0a0MP0"
train["stryear"] = train["date"].astype(str)
train["year"] = train["stryear"].str[0:4].astype(int)
# + id="ViJ9Gl9aezst"
# + id="2CkB9FJX3_Te" colab={"base_uri": "https://localhost:8080/"} outputId="e33ecc94-e259-413e-9538-b3629098c0c9"
#make feature
train["strdate"] = train["date"].astype(str)
train["month"] = train["strdate"].str[5].astype(int)
train["month"]
train["stryear"] = train["date"].astype(str)
train["year"] = train["stryear"].str[0:4].astype(int)
train["nullc"] = train.isnull().sum(axis=1)
train["ongame"] = np.where(train["gamePk"].isnull() == 1,0,1)
train["ongame"].unique()
# + id="adY9aY39Y2ez"
del awards
del scores
del rosters
del standings
# + papermill={"duration": 1.157038, "end_time": "2021-06-25T22:19:06.309107", "exception": false, "start_time": "2021-06-25T22:19:05.152069", "status": "completed"} tags=[] id="identified-resource"
# + papermill={"duration": 3.0266, "end_time": "2021-06-25T22:19:09.365076", "exception": false, "start_time": "2021-06-25T22:19:06.338476", "status": "completed"} tags=[] id="outstanding-spoke"
# label encoding
player2num = {c: i for i, c in enumerate(train['playerId'].unique())}
position2num = {c: i for i, c in enumerate(train['primaryPositionName'].unique())}
teamid2num = {c: i for i, c in enumerate(train['teamId'].unique())}
status2num = {c: i for i, c in enumerate(train['status'].unique())}
daynight2num = {c: i for i, c in enumerate(train['dayNight'].unique())}
seriesDescription2num = {c: i for i, c in enumerate(train['seriesDescription'].unique())}
gameType2num = {c: i for i, c in enumerate(train['gameType'].unique())}
#bitrhCountry2num = {c: i for i, c in enumerate(train["birthCountry"].unique())}
train['label_playerId'] = train['playerId'].map(player2num)
train['label_primaryPositionName'] = train['primaryPositionName'].map(position2num)
train['label_teamId'] = train['teamId'].map(teamid2num)
train['label_status'] = train['status'].map(status2num)
train["label_daynight"] = train['dayNight'].map(daynight2num)
train["label_seriesDescription"] = train["seriesDescription"].map(seriesDescription2num)
#train["birthCountry"] = train["birthCountry"].map(bitrhCountry2num)
from sklearn.compose import TransformedTargetRegressor
# + id="G3RKg8r9QpDf"
#train.to_csv("/content/drive/MyDrive/mlb/script/output/train.csv")
# + id="tnDAqSAkc7pP"
playerlist = train["playerId"].unique()
# + colab={"base_uri": "https://localhost:8080/"} id="cIq6Ji3fewUr" outputId="3ab1d2cb-40ce-4101-834c-dfb18df24903"
train["playerId"]
# + id="SnLF5qfiaTAm"
train_2017post = train[train["date"]<=20180328]
train_2018reg = train[(train["date"]>=20180329)&(train["date"]<=20181001)]
train_2018post = train[(train["date"]>=20181002)&(train["date"]<=20190319)]
train_2019reg = train[(train["date"]>=20190320)&(train["date"]<=20190929)]
train_2019post = train[(train["date"]>=20190930)&(train["date"]<=20200722)]
train_2020reg = train[(train["date"]>=20200723)&(train["date"]<=20200927)]
train_2020post = train[(train["date"]>=20200928)&(train["date"]<=20210331)]
train_2021reg = train[(train["date"]>=20210401)&(train["date"]<=20211003)]
# + colab={"base_uri": "https://localhost:8080/", "height": 224} id="5wU1G9E899HT" outputId="8adb3187-d1a5-493d-aa06-d13a7d294b58"
seasons
# + id="49AjMLJDhv97" colab={"base_uri": "https://localhost:8080/"} outputId="37da0cd6-8b52-40dd-c632-586872de4794"
#make feature
hitscol = ["playerId","preseasonhits"]
hits_2018 = train_2018reg.groupby("playerId").sum()["hits"].reset_index()
hits_2018["preseasonhits"] = hits_2018["hits"]
train_2018post = train_2018post.merge(hits_2018[hitscol], on=['playerId'], how='left')
train_2019reg = train_2019reg.merge(hits_2018[hitscol], on=['playerId'], how='left')
hits2019col = ["playerId","hits2019"]
hits_2019 = train_2019reg.groupby("playerId").sum()["hits"].reset_index()
hits_2019["preseasonhits"] = hits_2019["hits"]
train_2019post = train_2019post.merge(hits_2019[hitscol], on=['playerId'], how='left')
train_2020reg = train_2020reg.merge(hits_2019[hitscol], on=['playerId'], how='left')
hits2020col = ["playerId","hits2020"]
hits_2020 = train_2020reg.groupby("playerId").sum()["hits"].reset_index()
hits_2020["preseasonhits"] = hits_2020["hits"]
train_2020post = train_2020post.merge(hits_2020[hitscol], on=['playerId'], how='left')
train_2021reg = train_2021reg.merge(hits_2020[hitscol], on=['playerId'], how='left')
del hits_2018,hits_2019,hits_2020
gc.collect()
gamescol = ["playerId","preseasonpt"]
game2018 = train_2018reg.groupby("playerId").sum()["ongame"].reset_index()
game2018["preseasonpt"] = game2018["ongame"]
train_2018post = train_2018post.merge(game2018[gamescol], on=['playerId'], how='left')
train_2019reg = train_2019reg.merge(game2018[gamescol], on=['playerId'], how='left')
game2019 = train_2019reg.groupby("playerId").sum()["ongame"].reset_index()
game2019["preseasonpt"] = game2019["ongame"]
train_2019post = train_2019post.merge(game2019[gamescol], on=['playerId'], how='left')
train_2020reg = train_2020reg.merge(game2019[gamescol], on=['playerId'], how='left')
game2020 = train_2020reg.groupby("playerId").sum()["ongame"].reset_index()
game2020["preseasonpt"] = game2020["ongame"]
train_2020post = train_2020post.merge(game2020[gamescol], on=['playerId'], how='left')
train_2021reg = train_2021reg.merge(game2020[gamescol], on=['playerId'], how='left')
del game2018,game2019,game2020
gc.collect()
# + id="xT9yVB1Qzz44"
train = pd.concat([train_2017post,train_2018reg,train_2018post,train_2019reg,train_2019post,train_2020reg,train_2020post,train_2021reg])
train["preseasonhits"]
del train_2017post,train_2018post,train_2019post, train_2020post, train_2018reg, train_2019reg, train_2020reg
# + id="P7S_fxrw4NCB"
train["hitpergame"] = train["preseasonhits"] / train["preseasonpt"]
# + id="SwHCG2X0rmhR"
# + id="krSBoczTuaVd"
# + papermill={"duration": 8.158299, "end_time": "2021-06-25T22:19:17.555101", "exception": false, "start_time": "2021-06-25T22:19:09.396802", "status": "completed"} tags=[] id="negative-offset" colab={"base_uri": "https://localhost:8080/"} outputId="c318062c-af69-441b-d5d0-826327b0fc95"
train_y = train[['target1', 'target2', 'target3', 'target4']]
_index = (train['date'] < 20210401)
gc.collect()
x_train = train.loc[_index].reset_index(drop=True)
y_train = train_y.loc[_index].reset_index(drop=True)
x_valid = train.loc[~_index].reset_index(drop=True)
y_valid = train_y.loc[~_index].reset_index(drop=True)
gc.collect()
# + colab={"base_uri": "https://localhost:8080/", "height": 609} id="1bOsn97CzD8q" outputId="1a0ddeac-7e46-4f87-ae38-f63a9a451947"
train
# + papermill={"duration": 0.04094, "end_time": "2021-06-25T22:19:17.625445", "exception": false, "start_time": "2021-06-25T22:19:17.584505", "status": "completed"} tags=[] id="sticky-stand" colab={"base_uri": "https://localhost:8080/"} outputId="31036c35-c06e-44a3-f4a4-e2aafd71cc64"
def fit_lgbm(x_train, y_train, x_valid, y_valid, params: dict=None, verbose=100):
oof_pred = np.zeros(len(y_valid), dtype=np.float32)
#model = lgbm.LGBMRegressor(**params)
best_params, tuning_history = {}, []
lgb_train = lgbm.Dataset(x_train, y_train)
lgb_eval = lgbm.Dataset(x_valid, y_valid, reference=lgb_train)
model = lgbm.train(params,
lgb_train, valid_sets=lgb_eval,
early_stopping_rounds=100,
num_boost_round=10000,
verbose_eval=100,
#verbose=verbose,
)
oof_pred = model.predict(x_valid)
score = mean_absolute_error(oof_pred, y_valid)
print('mae:', score)
return oof_pred, model, score
import pickle
model1p = pickle.load(open('/content/drive/MyDrive/mlb/script/params/model1.pkl', 'rb'))
model2p = pickle.load(open('/content/drive/MyDrive/mlb/script/params/model2.pkl', 'rb'))
model3p = pickle.load(open('/content/drive/MyDrive/mlb/script/params/model3.pkl', 'rb'))
model4p = pickle.load(open('/content/drive/MyDrive/mlb/script/params/model4.pkl', 'rb'))
params = model1p.params
params2 = model2p.params
params3 = model3p.params
params4 = model4p.params
oof1, model1, score1 = fit_lgbm(
x_train[feature_cols], y_train['target1'],
x_valid[feature_cols], y_valid['target1'],
params
)
file = '/content/drive/MyDrive/mlb/script/output/model1.pkl'
pickle.dump(model1, open(file, 'wb'))
oof2, model2, score2 = fit_lgbm(
x_train[feature_cols2], y_train['target2'],
x_valid[feature_cols2], y_valid['target2'],
params2
)
file = '/content/drive/MyDrive/mlb/script/output/model2.pkl'
pickle.dump(model2, open(file, 'wb'))
oof3, model3, score3 = fit_lgbm(
x_train[feature_cols3], y_train['target3'],
x_valid[feature_cols3], y_valid['target3'],
params3
)
file = '/content/drive/MyDrive/mlb/script/output/model3.pkl'
pickle.dump(model3, open(file, 'wb'))
oof4, model4, score4 = fit_lgbm(
x_train[feature_cols4], y_train['target4'],
x_valid[feature_cols4], y_valid['target4'],
params4
)
file = '/content/drive/MyDrive/mlb/script/output/model4.pkl'
pickle.dump(model4, open(file, 'wb'))
score = (score1+score2+score3+score4) / 4
print(f'score: {score}')
# + papermill={"duration": 1.81619, "end_time": "2021-06-25T22:19:19.468136", "exception": false, "start_time": "2021-06-25T22:19:17.651946", "status": "completed"} tags=[] id="large-orlando"
file = '/content/drive/MyDrive/mlb/script/params/model1.pkl'
pickle.dump(model1, open(file, 'wb'))
file = '/content/drive/MyDrive/mlb/script/params/model2.pkl'
pickle.dump(model2, open(file, 'wb'))
file = '/content/drive/MyDrive/mlb/script/params/model3.pkl'
pickle.dump(model3, open(file, 'wb'))
file = '/content/drive/MyDrive/mlb/script/params/model4.pkl'
pickle.dump(model4, open(file, 'wb'))
# + id="xkqkayONf3SC"
# + papermill={"duration": 0.038158, "end_time": "2021-06-25T22:19:19.532427", "exception": false, "start_time": "2021-06-25T22:19:19.494269", "status": "completed"} tags=[] id="sensitive-establishment"
players_cols = ['playerId', 'primaryPositionName']
rosters_cols = ['playerId', 'teamId', 'status']
scores_cols = ['playerId', 'battingOrder', 'gamesPlayedBatting', 'flyOuts',
'groundOuts', 'runsScored', 'doubles', 'triples', 'homeRuns',
'strikeOuts', 'baseOnBalls', 'intentionalWalks', 'hits', 'hitByPitch',
'atBats', 'caughtStealing', 'stolenBases', 'groundIntoDoublePlay',
'groundIntoTriplePlay', 'plateAppearances', 'totalBases', 'rbi',
'leftOnBase', 'sacBunts', 'sacFlies', 'catchersInterference',
'pickoffs', 'gamesPlayedPitching', 'gamesStartedPitching',
'completeGamesPitching', 'shutoutsPitching', 'winsPitching',
'lossesPitching', 'flyOutsPitching', 'airOutsPitching',
'groundOutsPitching', 'runsPitching', 'doublesPitching',
'triplesPitching', 'homeRunsPitching', 'strikeOutsPitching',
'baseOnBallsPitching', 'intentionalWalksPitching', 'hitsPitching',
'hitByPitchPitching', 'atBatsPitching', 'caughtStealingPitching',
'stolenBasesPitching', 'inningsPitched', 'saveOpportunities',
'earnedRuns', 'battersFaced', 'outsPitching', 'pitchesThrown', 'balls',
'strikes', 'hitBatsmen', 'balks', 'wildPitches', 'pickoffsPitching',
'rbiPitching', 'gamesFinishedPitching', 'inheritedRunners',
'inheritedRunnersScored', 'catchersInterferencePitching',
'sacBuntsPitching', 'sacFliesPitching', 'saves', 'holds', 'blownSaves',
'assists', 'putOuts', 'errors', 'chances']
standings_cols = ["teamId","divisionRank","divisionLeader","wildCardLeader","leagueRank","divisionId","gameDate"]
null = np.nan
true = True
false = False
# + papermill={"duration": 265.737683, "end_time": "2021-06-25T22:23:45.296307", "exception": false, "start_time": "2021-06-25T22:19:19.558624", "status": "completed"} tags=[] id="hundred-thunder" colab={"base_uri": "https://localhost:8080/", "height": 447} outputId="a931dd00-13ed-46ad-e20f-50587da355bb"
import pandas as pd
import numpy as np
from datetime import timedelta
from tqdm import tqdm
import gc
from functools import reduce
from sklearn.model_selection import StratifiedKFold
ROOT_DIR = "../input/mlb-player-digital-engagement-forecasting"
#=======================#
def flatten(df, col):
du = (df.pivot(index="playerId", columns="EvalDate",
values=col).add_prefix(f"{col}_").
rename_axis(None, axis=1).reset_index())
return du
#============================#
def reducer(left, right):
return left.merge(right, on="playerId")
#========================
TGTCOLS = ["target1","target2","target3","target4"]
def train_lag(df, lag=1):
dp = df[["playerId","EvalDate"]+TGTCOLS].copy()
dp["EvalDate"] =dp["EvalDate"] + timedelta(days=lag)
df = df.merge(dp, on=["playerId", "EvalDate"], suffixes=["",f"_{lag}"], how="left")
return df
#=================================
def test_lag(sub):
sub["playerId"] = sub["date_playerId"].apply(lambda s: int( s.split("_")[1] ) )
assert sub.date.nunique() == 1
dte = sub["date"].unique()[0]
eval_dt = pd.to_datetime(dte, format="%Y%m%d")
dtes = [eval_dt + timedelta(days=-k) for k in LAGS]
mp_dtes = {eval_dt + timedelta(days=-k):k for k in LAGS}
sl = LAST.loc[LAST.EvalDate.between(dtes[-1], dtes[0]), ["EvalDate","playerId"]+TGTCOLS].copy()
sl["EvalDate"] = sl["EvalDate"].map(mp_dtes)
du = [flatten(sl, col) for col in TGTCOLS]
du = reduce(reducer, du)
return du, eval_dt
#
#===============
tr = pd.read_csv("../input/mlb-data/target.csv")
print(tr.shape)
gc.collect()
tr["EvalDate"] = pd.to_datetime(tr["EvalDate"])
tr["EvalDate"] = tr["EvalDate"] + timedelta(days=-1)
tr["EvalYear"] = tr["EvalDate"].dt.year
MED_DF = tr.groupby(["playerId","EvalYear"])[TGTCOLS].median().reset_index()
MEDCOLS = ["tgt1_med","tgt2_med", "tgt3_med", "tgt4_med"]
MED_DF.columns = ["playerId","EvalYear"] + MEDCOLS
LAGS = list(range(1,21))
FECOLS = [f"{col}_{lag}" for lag in reversed(LAGS) for col in TGTCOLS]
for lag in tqdm(LAGS):
tr = train_lag(tr, lag=lag)
gc.collect()
#===========
tr = tr.sort_values(by=["playerId", "EvalDate"])
print(tr.shape)
tr = tr.dropna()
print(tr.shape)
tr = tr.merge(MED_DF, on=["playerId","EvalYear"])
gc.collect()
X = tr[FECOLS+MEDCOLS].values
y = tr[TGTCOLS].values
cl = tr["playerId"].values
NFOLDS = 6
skf = StratifiedKFold(n_splits=NFOLDS)
folds = skf.split(X, cl)
folds = list(folds)
import tensorflow as tf
import tensorflow.keras.layers as L
import tensorflow.keras.models as M
from sklearn.metrics import mean_absolute_error, mean_squared_error
from tensorflow.keras.callbacks import ModelCheckpoint, ReduceLROnPlateau, EarlyStopping
tf.random.set_seed(777)
def make_model(n_in):
inp = L.Input(name="inputs", shape=(n_in,))
x = L.Dense(50, activation="relu", name="d1")(inp)
x = L.Dense(50, activation="relu", name="d2")(x)
preds = L.Dense(4, activation="linear", name="preds")(x)
model = M.Model(inp, preds, name="ANN")
model.compile(loss="mean_absolute_error", optimizer="adam")
return model
net = make_model(X.shape[1])
print(net.summary())
oof = np.zeros(y.shape)
nets = []
for idx in range(NFOLDS):
print("FOLD:", idx)
tr_idx, val_idx = folds[idx]
ckpt = ModelCheckpoint(f"w{idx}.h5", monitor='val_loss', verbose=1, save_best_only=True,mode='min')
reduce_lr = ReduceLROnPlateau(monitor='val_loss', factor=0.2,patience=3, min_lr=0.0005)
es = EarlyStopping(monitor='val_loss', patience=6)
reg = make_model(X.shape[1])
reg.fit(X[tr_idx], y[tr_idx], epochs=10, batch_size=35_000, validation_data=(X[val_idx], y[val_idx]),
verbose=1, callbacks=[ckpt, reduce_lr, es])
reg.load_weights(f"w{idx}.h5")
oof[val_idx] = reg.predict(X[val_idx], batch_size=50_000, verbose=1)
nets.append(reg)
gc.collect()
#
#
mae = mean_absolute_error(y, oof)
mse = mean_squared_error(y, oof, squared=False)
print("mae:", mae)
print("mse:", mse)
# Historical information to use in prediction time
bound_dt = pd.to_datetime("2021-01-01")
LAST = tr.loc[tr.EvalDate>bound_dt].copy()
LAST_MED_DF = MED_DF.loc[MED_DF.EvalYear==2021].copy()
LAST_MED_DF.drop("EvalYear", axis=1, inplace=True)
del tr
#"""
import mlb
FE = []; SUB = [];
# + id="K1VLUowOyspF"
# + [markdown] papermill={"duration": 0.643124, "end_time": "2021-06-25T22:23:46.587799", "exception": false, "start_time": "2021-06-25T22:23:45.944675", "status": "completed"} tags=[] id="defined-colors"
# ## Inference
# + papermill={"duration": 0.649231, "end_time": "2021-06-25T22:23:47.875889", "exception": false, "start_time": "2021-06-25T22:23:47.226658", "status": "completed"} tags=[] id="classified-cooper"
def unpack_json(json_str):
return np.nan if pd.isna(json_str) else pd.read_json(json_str)
# + papermill={"duration": 0.652999, "end_time": "2021-06-25T22:23:49.248559", "exception": false, "start_time": "2021-06-25T22:23:48.595560", "status": "completed"} tags=[] id="electoral-tribute"
"""players_cols = ['playerId', 'primaryPositionName']
rosters_cols = ['playerId', 'teamId', 'status']
scores_cols = ['playerId', 'battingOrder', 'gamesPlayedBatting', 'flyOuts',
'groundOuts', 'runsScored', 'doubles', 'triples', 'homeRuns',
'strikeOuts', 'baseOnBalls', 'intentionalWalks', 'hits', 'hitByPitch',
'atBats', 'caughtStealing', 'stolenBases', 'groundIntoDoublePlay',
'groundIntoTriplePlay', 'plateAppearances', 'totalBases', 'rbi',
'leftOnBase', 'sacBunts', 'sacFlies', 'catchersInterference',
'pickoffs', 'gamesPlayedPitching', 'gamesStartedPitching',
'completeGamesPitching', 'shutoutsPitching', 'winsPitching',
'lossesPitching', 'flyOutsPitching', 'airOutsPitching',
'groundOutsPitching', 'runsPitching', 'doublesPitching',
'triplesPitching', 'homeRunsPitching', 'strikeOutsPitching',
'baseOnBallsPitching', 'intentionalWalksPitching', 'hitsPitching',
'hitByPitchPitching', 'atBatsPitching', 'caughtStealingPitching',
'stolenBasesPitching', 'inningsPitched', 'saveOpportunities',
'earnedRuns', 'battersFaced', 'outsPitching', 'pitchesThrown', 'balls',
'strikes', 'hitBatsmen', 'balks', 'wildPitches', 'pickoffsPitching',
'rbiPitching', 'gamesFinishedPitching', 'inheritedRunners',
'inheritedRunnersScored', 'catchersInterferencePitching',
'sacBuntsPitching', 'sacFliesPitching', 'saves', 'holds', 'blownSaves',
'assists', 'putOuts', 'errors', 'chances']
null = np.nan
true = True
false = False"""
# + papermill={"duration": 11.293137, "end_time": "2021-06-25T22:24:01.199706", "exception": false, "start_time": "2021-06-25T22:23:49.906569", "status": "completed"} tags=[] id="thousand-comfort"
"""players_cols = ['playerId', 'primaryPositionName']
rosters_cols = ['playerId', 'teamId', 'status']
scores_cols = ['playerId', 'battingOrder', 'gamesPlayedBatting', 'flyOuts',
'groundOuts', 'runsScored', 'doubles', 'triples', 'homeRuns',
'strikeOuts', 'baseOnBalls', 'intentionalWalks', 'hits', 'hitByPitch',
'atBats', 'caughtStealing', 'stolenBases', 'groundIntoDoublePlay',
'groundIntoTriplePlay', 'plateAppearances', 'totalBases', 'rbi',
'leftOnBase', 'sacBunts', 'sacFlies', 'catchersInterference',
'pickoffs', 'gamesPlayedPitching', 'gamesStartedPitching',
'completeGamesPitching', 'shutoutsPitching', 'winsPitching',
'lossesPitching', 'flyOutsPitching', 'airOutsPitching',
'groundOutsPitching', 'runsPitching', 'doublesPitching',
'triplesPitching', 'homeRunsPitching', 'strikeOutsPitching',
'baseOnBallsPitching', 'intentionalWalksPitching', 'hitsPitching',
'hitByPitchPitching', 'atBatsPitching', 'caughtStealingPitching',
'stolenBasesPitching', 'inningsPitched', 'saveOpportunities',
'earnedRuns', 'battersFaced', 'outsPitching', 'pitchesThrown', 'balls',
'strikes', 'hitBatsmen', 'balks', 'wildPitches', 'pickoffsPitching',
'rbiPitching', 'gamesFinishedPitching', 'inheritedRunners',
'inheritedRunnersScored', 'catchersInterferencePitching',
'sacBuntsPitching', 'sacFliesPitching', 'saves', 'holds', 'blownSaves',
'assists', 'putOuts', 'errors', 'chances',"gamePk"]
null = np.nan
true = True
false = False
env = mlb.make_env() # initialize the environment
iter_test = env.iter_test() # iterator which loops over each date in test set
for (test_df, sample_prediction_df) in iter_test: # make predictions here
sub = copy.deepcopy(sample_prediction_df.reset_index())
sample_prediction_df = copy.deepcopy(sample_prediction_df.reset_index(drop=True))
# creat dataset
sample_prediction_df['playerId'] = sample_prediction_df['date_playerId']\
.map(lambda x: int(x.split('_')[1]))
# Dealing with missing values
if test_df['rosters'].iloc[0] == test_df['rosters'].iloc[0]:
test_rosters = pd.DataFrame(eval(test_df['rosters'].iloc[0]))
else:
test_rosters = pd.DataFrame({'playerId': sample_prediction_df['playerId']})
for col in rosters.columns:
if col == 'playerId': continue
test_rosters[col] = np.nan
if test_df['playerBoxScores'].iloc[0] == test_df['playerBoxScores'].iloc[0]:
test_scores = pd.DataFrame(eval(test_df['playerBoxScores'].iloc[0]))
else:
test_scores = pd.DataFrame({'playerId': sample_prediction_df['playerId']})
for col in scores.columns:
if col == 'playerId': continue
test_scores[col] = np.nan
test_games = unpack_json(test_df["games"].iloc[0])
test_standings = unpack_json(test_df["standings"].iloc[0])
test_scores = test_scores.groupby('playerId').sum().reset_index()
test = sample_prediction_df[['playerId']].copy()
test = test.merge(players[players_cols], on='playerId', how='left')
test = test.merge(test_rosters[rosters_cols], on='playerId', how='left')
test = test.merge(test_scores[scores_cols], on='playerId', how='left')
test = test.merge(test_games[games_cols], on="gamePk", how="left")
test = test.merge(awards2[awards_cols], on=['playerId'], how='left')
test = test.merge(player_target_stats, how='inner', left_on=["playerId"],right_on=["playerId"])
test = test.merge(test_standings[standings_cols], on=["teamId"], how='left')
#add feature
test["ongame"] = np.where(test["gamePk"].isnull() == 1,0,1)
test["ishome"] = np.where(test["teamId"]==test["homeId"],2,test["ongame"])
test["winorlose"] = np.where(test["teamId"]==test["homeId"],test["homeWinner"],test["awayWinner"])
test["winorlose"] = test["winorlose"].fillna(2.0).astype(int)
test["score"] = np.where(test["teamId"]==test["homeId"],test["homeScore"],test["awayScore"])
test["divisionRank"] = test["divisionRank"].fillna(7.0).astype(int)
test["divisionLeader"] = test["divisionLeader"].fillna(-1.0).astype(int)
test["wildCardLeader"] = np.where(test["wildCardLeader"]=="True",1,0)
#label encoding
test['label_playerId'] = test['playerId'].map(player2num)
test['label_primaryPositionName'] = test['primaryPositionName'].map(position2num)
test['label_teamId'] = test['teamId'].map(teamid2num)
test['label_status'] = test['status'].map(status2num)
test["label_daynight"] = test['dayNight'].map(daynight2num)
test["label_seriesDescription"] = test["seriesDescription"].map(seriesDescription2num)
test["gameType"] = test["gameType"].map(gameType2num)
display(test)
test_X1 = test[feature_cols1]
test_X2 = test[feature_cols2]
test_X3 = test[feature_cols3]
test_X4 = test[feature_cols4]
# predict
pred1 = model1.predict(test_X1)
pred2 = model2.predict(test_X2)
pred3 = model3.predict(test_X3)
pred4 = model4.predict(test_X4)
# merge submission
sample_prediction_df['target1'] = np.clip(pred1, 0, 100)
sample_prediction_df['target2'] = np.clip(pred2, 0, 100)
sample_prediction_df['target3'] = np.clip(pred3, 0, 100)
sample_prediction_df['target4'] = np.clip(pred4, 0, 100)
sample_prediction_df = sample_prediction_df.fillna(0.)
del sample_prediction_df['playerId']
# TF summit
# Features computation at Evaluation Date
sub_fe, eval_dt = test_lag(sub)
sub_fe = sub_fe.merge(LAST_MED_DF, on="playerId", how="left")
sub_fe = sub_fe.fillna(0.)
_preds = 0.
for reg in nets:
_preds += reg.predict(sub_fe[FECOLS + MEDCOLS]) / NFOLDS
sub_fe[TGTCOLS] = np.clip(_preds, 0, 100)
sub.drop(["date"]+TGTCOLS, axis=1, inplace=True)
sub = sub.merge(sub_fe[["playerId"]+TGTCOLS], on="playerId", how="left")
sub.drop("playerId", axis=1, inplace=True)
sub = sub.fillna(0.)
# Blending
blend = pd.concat(
[sub[['date_playerId']],
(0.1*sub.drop('date_playerId', axis=1) + 0.9*sample_prediction_df.drop('date_playerId', axis=1))],
axis=1
)
env.predict(blend)
# Update Available information
sub_fe["EvalDate"] = eval_dt
#sub_fe.drop(MEDCOLS, axis=1, inplace=True)
LAST = LAST.append(sub_fe)
LAST = LAST.drop_duplicates(subset=["EvalDate","playerId"], keep="last")"""
# + papermill={"duration": 0.688226, "end_time": "2021-06-25T22:24:02.557998", "exception": false, "start_time": "2021-06-25T22:24:01.869772", "status": "completed"} tags=[] id="brief-palmer"
display(test_df['games'])
# + papermill={"duration": 0.657564, "end_time": "2021-06-25T22:24:03.894025", "exception": false, "start_time": "2021-06-25T22:24:03.236461", "status": "completed"} tags=[] id="partial-morning"
def unpack_json(json_str):
return np.nan if pd.isna(json_str) else pd.read_json(json_str)
# + papermill={"duration": 0.761079, "end_time": "2021-06-25T22:24:05.325843", "exception": false, "start_time": "2021-06-25T22:24:04.564764", "status": "completed"} tags=[] id="smoking-discretion"
unpack_json(test_df["games"].iloc[0])
# + papermill={"duration": 0.692076, "end_time": "2021-06-25T22:24:06.680048", "exception": false, "start_time": "2021-06-25T22:24:05.987972", "status": "completed"} tags=[] id="talented-generic"
pd.concat(
[sub[['date_playerId']],
(sub.drop('date_playerId', axis=1) + sample_prediction_df.drop('date_playerId', axis=1)) / 2],
axis=1
)
# + papermill={"duration": 0.679239, "end_time": "2021-06-25T22:24:08.010626", "exception": false, "start_time": "2021-06-25T22:24:07.331387", "status": "completed"} tags=[] id="vertical-ethernet"
sample_prediction_df
| submit2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
from collections import Counter
from glob import glob
import csv
import os.path as path
from pie_extended.cli.utils import get_tagger
from pie_extended.models.dum.imports import get_iterator_and_processor
path_name = 'txt/*.txt'
model_name = "dum"
tagger = get_tagger(model_name, batch_size=256, device="cpu", model_path=None)
def dict_freq_tri_pos(list_token, doc_name):
list_tri_pos = ['n_v_pron', 'pron_n_v', 'adp_pron_n', 'adp_art_n', 'v_pron_n', 'n_adv_v', 'n_adp_n', 'art_n_v', 'adv_v_pron', 'v_pron_v', 'n_pron_n', 'n_n_v', 'n_art_n', 'v_art_n', 'v_adv_v', 'n_v_v', 'n_conj_n', 'pron_v_pron', 'v_conj_pron', 'n_pron_v', 'art_n_n', 'pron_adv_v', 'v_adp_pron', 'n_adp_pron', 'n_conj_pron', 'pron_n_n', 'n_n_n', 'v_adp_n', 'n_v_adv', 'pron_n_pron', 'pron_v_v', 'v_pron_adv', 'art_n_adp', 'pron_n_adv', 'v_v_pron', 'n_v_conj', 'art_adj_n', 'adv_v_v', 'n_v_adp', 'pron_v_adp', 'adp_n_n', 'v_pron_pron', 'adp_n_v', 'adv_v_adv', 'conj_pron_v', 'pron_v_adv', 'adv_adv_v', 'n_n_pron', 'conj_pron_n', 'pron_pron_v']
dict_result = dict.fromkeys(list_tri_pos)
list_trigram = trigrammize(list_token)
dict_temp = Counter(list_trigram)
for key in dict_temp.keys():
if key in dict_result.keys():
dict_result[key] = dict_temp[key]/len(list_trigram)
return dict_result
def trigrammize(list_token):
"""fonction qui prend en parametre une liste de tokens et retourne une liste de tri-grammes"""
list_trigram = []
for indice_token in range(len(list_token)-2):
trigram = list_token[indice_token].split("(")[0]+'_'+list_token[indice_token+1].split("(")[0]+'_'+list_token[indice_token+2].split("(")[0]
list_trigram.append(trigram)
return list_trigram
def dict_freq_tri_char(list_tri_chars, doc_name):
list_tri_chars_select = ['en ', 'er ', 'et ', 'n, ', 'en,', 't, ', ' de', 'den', 'an ', 'de ', ' he', 'ijn', ' ge', 'ver', 'een', ' ve', 'at ', 'oor', 'n d', 'aer', ' en', 'jn ', ' da', 'sch', 'eer', 'der', 'ck ', 'cht', 'ie ', ' va', ', D', ' ee', ' be', ' me', 'nde', ' we', 'aar', 'n s', 'ten', ' in', 't d', ' di', 'iet', ' mi', 'in ', 't g', 'ghe', 'te ', ' al', ' En']
dict_result = dict.fromkeys(list_tri_chars_select)
dict_temp = Counter(list_tri_chars)
for key in dict_temp.keys():
if key in dict_result.keys():
dict_result[key] = dict_temp[key]/len(list_tri_chars)
return dict_result
def tri_chars(chaine):
list_three_chars = []
for i in range(0, len(chaine)-2):
list_three_chars.append(chaine[i]+chaine[i+1]+chaine[i+2])
return list_three_chars
def dict_freq_lemma(list_lemma, doc_name):
list_lemma_select = ['een', 'de', 'ik', 'gij', 'hij', 'ne', 'in', 'mijn', 'van', 'het', 'uw', 'en', 'zijn', 'wij', 'zij', 'eten', 'niet', 'dat', 'te', 'met', 'op', 'die', 'haar', 'eer', 'voor', 'hebben', 'zo', 'al', 'als', 'aan', 'et', 'hoofd', 'na', 'tot', 'ook', 'door', 'hier', 'of', 'hoe', 'dan', 'geen', 'om', 'nu', 'wel', 'ons', 'haten', 'deze', 'wat', 'dit', 'bij']
dict_result = dict.fromkeys(list_lemma_select)
dict_temp = Counter(list_lemma)
for key in dict_temp.keys():
if key in dict_result.keys():
dict_result[key] = dict_temp[key]/len(list_lemma)
return dict_result
def nettoie_lemma(list_lemma, list_pos):
pos_ok = ["pre", "adv", "det", "conj", "pron", "adp", "art"]
list_nettoie = []
if len(list_lemma) == len(list_pos):
for i in range(0, len(list_lemma)-1):
if list_pos[i].split("(")[0] in pos_ok:
list_nettoie.append(list_lemma[i])
return list_nettoie
def main(path_name, doc_name):
with open(path_name, encoding="utf8") as file:
lignes = file.readlines()
for ligne in lignes:
iterator, processor = get_iterator_and_processor()
annotation = tagger.tag_str(ligne, iterator=iterator, processor=processor)
list_three_chars = tri_chars(ligne)
df_pie = pd.DataFrame(annotation)
dict_result_pos = dict_freq_tri_pos(df_pie['pos'], doc_name)
dict_result_char = dict_freq_tri_char(list_three_chars, doc_name)
list_lemma = nettoie_lemma(df_pie['lemma'], df_pie['pos'])
dict_result_lemma = dict_freq_lemma(list_lemma, doc_name)
return dict_result_pos, dict_result_char, dict_result_lemma
def moulinette(path_name):
dict_results = {}
df_main = pd.DataFrame()
for doc in glob(path_name):
doc_name = path.splitext(path.basename(doc))[0]
dict_result_pos, dict_result_char, dict_result_lemma = main(doc, doc_name)
dict_results.update(dict_result_pos)
dict_results.update(dict_result_char)
dict_results.update(dict_result_lemma)
dict_results["index"] = doc_name
df_temp = pd.DataFrame(dict_results, index=[0])
df_main = df_main.append(df_temp, ignore_index = True)
df_main.set_index("index", inplace = True)
return df_main
df_main = moulinette(path_name)
df_main
df_main.to_csv(r'features.csv', index = True)
| table_features.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#from customplot import *
#import sqlite3
# %autosave 0
import itertools
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn.cluster import KMeans
from sklearn.ensemble import RandomForestClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import accuracy_score
from sklearn.metrics import classification_report
from sklearn.metrics import confusion_matrix
from sklearn.metrics import log_loss
from sklearn.metrics import precision_score
from sklearn.metrics import precision_recall_curve
from sklearn.metrics import recall_score
from sklearn.metrics import roc_auc_score
from sklearn.metrics import roc_curve
from sklearn.model_selection import train_test_split
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import cross_val_predict
from sklearn.model_selection import GridSearchCV
from sklearn.neighbors import KNeighborsClassifier
from sklearn.neural_network import MLPClassifier
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler
from sklearn.preprocessing import MinMaxScaler
from sklearn.tree import DecisionTreeClassifier
# -
def plot_confusion_matrix(cm, classes,
normalize=False,
title='Confusion matrix',
cmap=plt.cm.Blues):
"""
This function prints and plots the confusion matrix.
Normalization can be applied by setting `normalize=True`.
"""
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
print("Normalized confusion matrix")
else:
print('Confusion matrix, without normalization')
#print(cm)
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45)
plt.yticks(tick_marks, classes)
fmt = '.2f' if normalize else 'd'
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, format(cm[i, j], fmt),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
#Setando configurações de visualização
pd.options.display.max_rows=350
pd.options.display.max_columns=60
df=pd.read_csv('baseProjeto_entradaModelo.csv', index_col=0)
df
df.columns
X=df[['ATRIB_MED1', 'ATRIB_MAX1',
'ATRIB_DIST1', 'DIFP', 'MGP1', 'MGP2', 'MGP3', 'MGP4', 'MGP5', 'MGP6',
'MGP7', 'MGP8', 'MGP9', 'MGP10', 'MGP11', 'MGP12', 'MGP13', 'MGP14']]
X
X.info()
'''
#cat=['MGP1_sim', 'MGP2_sim', 'MGP3_sim', 'MGP4_sim',
'MGP5_sim', 'MGP6_sim', 'MGP7_sim', 'MGP8_sim', 'MGP9_sim', 'MGP10_sim',
'MGP11_sim', 'MGP12_sim', 'MGP13_sim', 'MGP14_sim',]
#X[cat] = X[cat].astype('category')
'''
X.info()
y = df['Perda30']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=.20, random_state=21)
# <br>
# ## Logistic Regression
steps=[('scaler', StandardScaler()),
('logreg',LogisticRegression())]
pipeline = Pipeline(steps)
logreg_scaled = pipeline.fit(X_train, y_train)
y_pred = pipeline.predict(X_test)
accuracy_score(y_test, y_pred)
y_pred_prob = pipeline.predict_proba(X_test)[:,1]
# +
# Generate ROC curve values: fpr, tpr, thresholds
fpr, tpr, thresholds = roc_curve(y_test, y_pred_prob)
# Plot ROC curve
plt.plot([0, 1], [0, 1], 'k--')
plt.plot(fpr, tpr)
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('ROC Curve')
plt.show()
# -
roc_auc_score(y_test, y_pred_prob)
confusion_matrix(y_test,y_pred)
logreg_unscaled = LogisticRegression().fit(X_train, y_train)
logreg_unscaled.score(X_test, y_test)
# <br>
# ## KNeighbors
steps = [('scaler', StandardScaler()),(('knn', KNeighborsClassifier()))]
pipeline = Pipeline(steps)
knn_scaled = pipeline.fit(X_train, y_train)
y_pred = pipeline.predict(X_test)
accuracy_score(y_test, y_pred)
y_pred_prob = pipeline.predict_proba(X_test)[:,1]
# +
# Generate ROC curve values: fpr, tpr, thresholds
fpr, tpr, thresholds = roc_curve(y_test, y_pred_prob)
# Plot ROC curve
plt.plot([0, 1], [0, 1], 'k--')
plt.plot(fpr, tpr)
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('ROC Curve')
plt.show()
# -
roc_auc_score(y_test, y_pred_prob)
knn_unscaled = KNeighborsClassifier().fit(X_train, y_train)
knn_unscaled.score(X_test, y_test)
confusion_matrix(y_test,y_pred)
steps = [('scaler', StandardScaler()),(('knn', KNeighborsClassifier()))]
pipeline = Pipeline(steps)
parameters = {'knn__n_neighbors' : np.arange(1, 50)}
cv = GridSearchCV(pipeline, param_grid=parameters)
cv.fit(X, y);
#cv.fit(X_train, y_train);
y_pred = cv.predict(X_test)
print(cv.best_params_)
print(cv.score(X_test, y_test))
y_pred_prob = cv.predict_proba(X_test)[:,1]
# +
# Generate ROC curve values: fpr, tpr, thresholds
fpr, tpr, thresholds = roc_curve(y_test, y_pred_prob)
# Plot ROC curve
plt.plot([0, 1], [0, 1], 'k--')
plt.plot(fpr, tpr)
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('ROC Curve')
plt.show()
# -
roc_auc_score(y_test, y_pred_prob)
confusion_matrix(y_test,y_pred)
print(classification_report(y_test, y_pred))
# <br>
# ## Neural Network - Scaled with StandardScaller
steps = [('scaler', StandardScaler()),(('neural', MLPClassifier(solver='adam', alpha=1e-5, hidden_layer_sizes=(50, 32), random_state=42, max_iter=500, warm_start=True)))]
pipeline = Pipeline(steps)
# hidden_layer_sizes=(n1, n2,..., nx) <br>
# n1 = number of neurons in hidden layer_1 <br>
# nx = number of neurons in hidden layer_x <br>
neural_scaled = pipeline.fit(X_train, y_train)
y_pred = pipeline.predict(X_test)
accuracy_score(y_test, y_pred)
y_pred_prob = pipeline.predict_proba(X_test)[:,1]
# +
#for i in range(len(y_pred)):
# print(y_pred_prob[i],y_pred[i])
# +
# Generate ROC curve values: fpr, tpr, thresholds
fpr, tpr, thresholds = roc_curve(y_test, y_pred_prob)
# Plot ROC curve
plt.plot([0, 1], [0, 1], 'k--')
plt.plot(fpr, tpr)
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('ROC Curve')
plt.show()
# -
roc_auc_score(y_test, y_pred_prob)
# +
print(confusion_matrix(y_test,y_pred))
# Compute confusion matrix
cnf_matrix = confusion_matrix(y_test, y_pred)
np.set_printoptions(precision=2)
# Plot non-normalized confusion matrix
plt.figure()
plot_confusion_matrix(cnf_matrix, classes=['Sem Perda','Perda'],
title='Confusion matrix, without normalization')
# -
print(classification_report(y_test, y_pred))
cv_scores = cross_val_score(pipeline, X, y, cv=5)
# +
print(cv_scores)
print("Average 5-Fold CV Score: {}".format(np.mean(cv_scores)))
# -
# <br>
# ## Neural Network - Scaled with MinMaxScaller
steps = [('scaler', MinMaxScaler()),(('neural', MLPClassifier(solver='adam', alpha=1e-5, hidden_layer_sizes=(50, 32), random_state=42, max_iter=500, warm_start=True)))]
pipeline = Pipeline(steps)
# hidden_layer_sizes=(n1, n2,..., nx) <br>
# n1 = number of neurons in hidden layer_1 <br>
# nx = number of neurons in hidden layer_x <br>
neural_scaled = pipeline.fit(X_train, y_train)
y_pred = pipeline.predict(X_test)
accuracy_score(y_test, y_pred)
y_pred_prob = pipeline.predict_proba(X_test)[:,1]
# +
#for i in range(len(y_pred)):
# print(y_pred_prob[i],y_pred[i])
# +
# Generate ROC curve values: fpr, tpr, thresholds
fpr, tpr, thresholds = roc_curve(y_test, y_pred_prob)
# Plot ROC curve
plt.plot([0, 1], [0, 1], 'k--')
plt.plot(fpr, tpr)
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('ROC Curve')
plt.show()
# -
roc_auc_score(y_test, y_pred_prob)
# +
print(confusion_matrix(y_test,y_pred))
# Compute confusion matrix
cnf_matrix = confusion_matrix(y_test, y_pred)
np.set_printoptions(precision=2)
# Plot non-normalized confusion matrix
plt.figure()
plot_confusion_matrix(cnf_matrix, classes=['Sem Perda','Perda'],
title='Confusion matrix, without MinMaxScaller')
# -
print(classification_report(y_test, y_pred))
cv_scores = cross_val_score(pipeline, X, y, cv=5)
# +
#y_scores = cross_val_predict(pipeline, X, y, cv=5,
#method="decision_function")
# -
precisions, recalls, thresholds = precision_recall_curve(y_test, y_pred_prob)
def plot_precision_recall_vs_threshold(precisions, recalls, thresholds):
plt.plot(thresholds, precisions[:-1], "b--", label="Precision")
plt.plot(thresholds, recalls[:-1], "g-", label="Recall")
plt.xlabel("Threshold")
plt.legend(loc="upper left")
plt.ylim([0, 1])
plot_precision_recall_vs_threshold(precisions, recalls, thresholds)
plt.show()
# <br>
# ## RandomForest
steps = [('scaler', StandardScaler()),(('rf', RandomForestClassifier(n_estimators=200, max_features=8, max_depth=12)))]
pipeline = Pipeline(steps)
random_scaled = pipeline.fit(X_train, y_train)
y_pred = pipeline.predict(X_test)
accuracy_score(y_test, y_pred)
y_pred_prob = pipeline.predict_proba(X_test)[:,1]
# +
# Generate ROC curve values: fpr, tpr, thresholds
fpr, tpr, thresholds = roc_curve(y_test, y_pred_prob)
# Plot ROC curve
plt.plot([0, 1], [0, 1], 'k--')
plt.plot(fpr, tpr)
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('ROC Curve')
plt.show()
# -
roc_auc_score(y_test, y_pred_prob)
# +
print(confusion_matrix(y_test,y_pred))
# Compute confusion matrix
cnf_matrix = confusion_matrix(y_test, y_pred)
np.set_printoptions(precision=2)
# Plot non-normalized confusion matrix
plt.figure()
plot_confusion_matrix(cnf_matrix, classes=['Sem Perda','Perda'],
title='Confusion matrix, without normalization')
# -
# Positive Predictive Value (PPV)
# $$Precision=\frac{TP}{TP+FP}$$
# <br>
# Sensitivity, Hit Rate, True Positive Rate
# $$Recall=\frac{TP}{TP+FN}$$
# <br>
# Harmonic mean between Precision and Recall
# $$F1 Score=2 * \frac{Precision * Recall}{Precision + Recall}$$
print(classification_report(y_test, y_pred))
rf = RandomForestClassifier(n_jobs=-1)
parameters = {'n_estimators' : np.arange(1, 200), 'max_depth': np.arange(1, 50)}
cv = GridSearchCV(rf, param_grid=parameters)
#"max_depth": np.arange(1, 50),
#"max_features": [1, 3, 10],
#"min_samples_split": np.arange(2, 20),
#"min_samples_leaf": np.arange(1, 10),
#"bootstrap": [True, False],
#"criterion": ["gini", "entropy"]
#rf.fit(X_train, y_train);
cv.fit(X_train, y_train);
print(cv.best_params_)
#y_pred = rf.predict(X_test)
y_pred = cv.predict(X_test)
# +
y_pred_prob = cv.predict_proba(X_test)[:,1]
# Generate ROC curve values: fpr, tpr, thresholds
fpr, tpr, thresholds = roc_curve(y_test, y_pred_prob)
# Plot ROC curve
plt.plot([0, 1], [0, 1], 'k--')
plt.plot(fpr, tpr)
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('ROC Curve')
plt.show()
# -
roc_auc_score(y_test, y_pred_prob)
# +
#print(confusion_matrix(y_test,y_pred))
# Compute confusion matrix
cnf_matrix = confusion_matrix(y_test, y_pred)
np.set_printoptions(precision=2)
# Plot non-normalized confusion matrix
plt.figure()
plot_confusion_matrix(cnf_matrix, classes=['Sem Perda','Perda'],
title='Confusion matrix, without normalization')
# -
print(classification_report(y_test, y_pred))
| Model-Study/mlModels.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exercise 5.1
#
# ## Import Libraries
# Python requires importing libraries and functions you need to access specific tools like science (scipy), linear algebra (numpy), and graphics (matplotlib). These libraries can be installed using the ```pip``` command line tool. Alternatively you can install an python distribution like [Anaconda](https://www.continuum.io/downloads) or [Canopy](https://www.enthought.com/products/canopy/) which have these and many other standard package pre-installed.
import ipywidgets as widgets # add new widgets
from ipywidgets import interact, interactive, fixed
import os
from IPython.display import display
import numpy as np # linear algebra / matrices
from skimage.color import label2rgb
from sklearn.metrics import roc_curve, auc # roc curve tools
from skimage.segmentation import mark_boundaries # mark labels
from skimage.io import imread # read in images
import matplotlib.pyplot as plt # plotting
# %matplotlib inline
# make the notebook interactive
base_path = '04-files'
seg_path = os.path.join(base_path, 'DogVsMuffin_seg_bw.jpg')
rgb_path = os.path.join(base_path, 'DogVsMuffin.jpg')
face_path = os.path.join(base_path, 'DogVsMuffin_face.jpg')
seg_img = imread(seg_path)[80:520:2, :450:2]
rgb_img = imread(rgb_path)[80:520:2, :450:2, :]
face_img = imread(face_path)
print('RGB Size', rgb_img.shape, 'Seg Size',
seg_img.shape, 'Face Size', face_img.shape)
# %matplotlib inline
fig, (ax1, ax2, ax3) = plt.subplots(1, 3, figsize=(20, 5))
ax1.imshow(rgb_img) # show the color image
ax1.set_title("Color Image")
ax2.imshow(seg_img, cmap='gray') # show the segments
ax2.set_title("Ground Truth")
ax3.imshow(mark_boundaries(rgb_img, seg_img))
ax3.set_title("Labeled Image")
# ## Creating a Simple ROC Curve
# We use the score function of taking the mean of the red green and blue channels
# $$ I = \frac{R+G+B}{3} $$
# We then take the score by normalizing by the maximum value (since the image is 8bit this is 255)
# $$ s = \frac{I}{255} $$
ground_truth_labels = seg_img.flatten() > 0
score_value = 1-np.mean(rgb_img.astype(np.float32), 2).flatten()/255.0
fpr, tpr, _ = roc_curve(ground_truth_labels, score_value)
roc_auc = auc(fpr, tpr)
# %matplotlib inline
fig, ax = plt.subplots(1, 1)
ax.plot(fpr, tpr, label='ROC curve (area = %0.2f)' % roc_auc)
ax.plot([0, 1], [0, 1], 'k--')
ax.set_xlim([0.0, 1.0])
ax.set_ylim([0.0, 1.05])
ax.set_xlabel('False Positive Rate')
ax.set_ylabel('True Positive Rate')
ax.set_title('Receiver operating characteristic example')
ax.legend(loc="lower right")
# ## Adding Filters
# We can add a filter to this process by importing a ```uniform_filter``` and applying it before processing the image
#
# +
from scipy.ndimage.filters import uniform_filter
# %matplotlib inline
filter_size = 45
filtered_image = uniform_filter(np.mean(rgb_img, 2), filter_size)
score_value = 1-filtered_image.astype(np.float32).flatten()/255.0
fpr2, tpr2, _ = roc_curve(ground_truth_labels, score_value)
roc_auc2 = auc(fpr2, tpr2)
fig, ax = plt.subplots(1, 1)
ax.plot(fpr, tpr, label='Raw ROC curve (area = %0.2f)' % roc_auc)
ax.plot(fpr2, tpr2, label='Filtered ROC curve (area = %0.2f)' % roc_auc2)
ax.plot([0, 1], [0, 1], 'k--')
ax.set_xlim([0.0, 1.0])
ax.set_ylim([0.0, 1.05])
ax.set_xlabel('False Positive Rate')
ax.set_ylabel('True Positive Rate')
ax.set_title('Receiver operating characteristic example')
ax.legend(loc="lower right")
# -
# ### Tasks
# 1. How can you improve filtering in this analysis?
# - Which filter elements might improve the area under the ROC?
# - Try making workflows to test out a few different filters
# 2. Where might morphological operations fit in?
# - How can you make them part of this workflow as well?
# 3. (Challenge) Try and use the optimize toolbox of _scipy_ with the fmin function (```from scipy.optimize import fmin```) to find the optimum parmeters for the highers area (hint: fmin finds the minimum value)
# +
from scipy.optimize import fmin
def calc_auc(rv, gv, bv, fsize):
filter_size = 45
gray_image = (rv*rgb_img[:, :, 0]+gv*rgb_img[:, :,
1]+bv*rgb_img[:, :, 2])/(rv+gv+bv)
filtered_image = uniform_filter(gray_image, filter_size)
score_value = filtered_image.astype(np.float32).flatten()/255.0
fpr2, tpr2, _ = roc_curve(ground_truth_labels, score_value)
return {'fpr': fpr2, 'tpr': tpr2, 'auc': auc(fpr2, tpr2), 'gimg': gray_image, 'fimg': filtered_image}
# +
# test the function to make sure it works
def min_func(args): return 1-calc_auc(*args)['auc']
min_start = [1, 1, 1, 20]
min_func(min_start)
# -
opt_res = fmin(min_func, min_start)
opt_values = calc_auc(*opt_res)
tprOpt = opt_values['tpr']
fprOpt = opt_values['fpr']
roc_aucOpt = opt_values['auc']
fig, (ax_img, ax) = plt.subplots(1, 2, figsize=(20, 10))
ax_img.imshow(opt_values['gimg'], cmap='gray')
ax_img.set_title('Transformed Color Image')
ax.plot(fpr, tpr, label='Raw ROC curve (area = %0.2f)' % roc_auc)
ax.plot(fprOpt, tprOpt, label='Optimized ROC curve (area = %0.2f)' % roc_aucOpt)
ax.plot([0, 1], [0, 1], 'k--')
ax.set_xlim([0.0, 1.0])
ax.set_ylim([0.0, 1.05])
ax.set_xlabel('False Positive Rate')
ax.set_ylabel('True Positive Rate')
ax.set_title('Receiver operating characteristic example')
ax.legend(loc="lower right")
# ## Non-linear optimization
# Here we use non-linear approaches to improve the quality of the results
# +
def relu(x):
return (x+np.abs(x))/2
def calc_auc_nl(rv, rm, gv, gm, bv, bm):
filter_size = 45
gray_image = (rv*relu(rgb_img[:, :, 0]/255.0-rm)+gv*relu(rgb_img[:, :, 1]/255.0-gm) +
bv*relu(rgb_img[:, :, 2]/255.0-bm))/(rv+gv+bv)
score_value = gray_image.astype(np.float32).flatten()
fpr2, tpr2, _ = roc_curve(ground_truth_labels, score_value)
return {'fpr': fpr2, 'tpr': tpr2, 'auc': auc(fpr2, tpr2), 'gimg': gray_image, 'fimg': filtered_image}
# +
# test the function to make sure it works
def min_func(args): return 1-calc_auc_nl(*args)['auc']
min_start = [1, 0, 1, 0, 1, 0]
min_start[0] = opt_res[0]
min_start[2] = opt_res[1]
min_start[4] = opt_res[2]
min_func(min_start)
# -
opt_res = fmin(min_func, min_start, maxiter=100)
opt_values_nl = calc_auc_nl(*opt_res)
tprOpt_nl = opt_values_nl['tpr']
fprOpt_nl = opt_values_nl['fpr']
roc_aucOpt_nl = opt_values_nl['auc']
fig, (ax_img, ax) = plt.subplots(1, 2, figsize=(20, 10))
ax_img.imshow(opt_values_nl['gimg'], cmap='gray')
ax_img.set_title('Transformed Color Image')
ax.plot(fpr, tpr, label='Raw ROC curve (area = %0.2f)' % roc_auc)
ax.plot(fprOpt, tprOpt, label='Optimized ROC curve (area = %0.2f)' % roc_aucOpt)
ax.plot(fprOpt_nl, tprOpt_nl,
label='NL Optimized ROC curve (area = %0.2f)' % roc_aucOpt_nl)
ax.plot([0, 1], [0, 1], 'k--')
ax.set_xlim([0.0, 1.0])
ax.set_ylim([0.0, 1.05])
ax.set_xlabel('False Positive Rate')
ax.set_ylabel('True Positive Rate')
ax.set_title('Receiver operating characteristic example')
ax.legend(loc="lower right")
# # Next Steps
# Rather than simply adjusting basic parameters, we can adjust entire arrays of information. The example below is the a convolutional neural network with one two layers
# +
from scipy.signal import fftconvolve
def convolve(img1, img2): return fftconvolve(img1, img2, mode='same')
CONV_SIZE = (10, 10, 1)
grey_img = np.reshape(np.mean(rgb_img, 2)/255.0,
(rgb_img.shape[0], rgb_img.shape[1], 1))
def calc_auc_conv(rcoefs):
coefs = rcoefs.reshape(CONV_SIZE)/rcoefs.sum()
score_image = relu(convolve(grey_img, coefs))
score_value = score_image.flatten()
fpr2, tpr2, _ = roc_curve(ground_truth_labels, score_value)
return {'fpr': fpr2, 'tpr': tpr2, 'auc': auc(fpr2, tpr2), 'gimg': score_image}
# -
# ## Make a nice gaussian kernel
#
np.random.seed(2019)
from functools import reduce
def gkern_nd(d=2, kernlen=21, nsigs=3, min_smooth_val=1e-2):
nsigs = [nsigs] * d
k_wid = (kernlen - 1) / 2
all_axs = [np.linspace(-k_wid, k_wid, kernlen)] * d
all_xxs = np.meshgrid(*all_axs)
all_dist = reduce(np.add, [
np.square(cur_xx) / (2 * np.square(np.clip(nsig, min_smooth_val,
kernlen)))
for cur_xx, nsig in zip(all_xxs, nsigs)])
kernel_raw = np.exp(-all_dist)
return kernel_raw / kernel_raw.sum()
# +
# test the function to make sure it works
def min_func(rcoefs): return 1-calc_auc_conv(rcoefs)['auc']
min_start = gkern_nd(2, CONV_SIZE[0]).ravel()
min_func(min_start)
# -
opt_res_conv = min_start
opt_res_conv = fmin(min_func,
opt_res_conv,
maxiter=500)
opt_values_conv = calc_auc_conv(opt_res_conv)
tprOpt_conv = opt_values_conv['tpr']
fprOpt_conv = opt_values_conv['fpr']
roc_aucOpt_conv = opt_values_conv['auc']
out_kernel = opt_res_conv.reshape(CONV_SIZE)/opt_res_conv.sum()
fig, ax_all = plt.subplots(1, out_kernel.shape[2])
for i, c_ax in enumerate(np.array(ax_all).flatten()):
c_ax.imshow(out_kernel[:, :, i])
c_ax.set_title(str(i))
fig, (ax_img, ax) = plt.subplots(1, 2, figsize=(20, 10))
ax_img.imshow(opt_values_conv['gimg'].squeeze(), cmap='gray')
ax_img.set_title('Transformed Color Image')
ax.plot(fpr, tpr, label='Raw ROC curve (area = %0.2f)' % roc_auc)
ax.plot(fprOpt, tprOpt, label='Optimized ROC curve (area = %0.2f)' % roc_aucOpt)
ax.plot(fprOpt_conv, tprOpt_conv,
label='CNN Optimized ROC curve (area = %0.2f)' % roc_aucOpt_conv)
ax.plot([0, 1], [0, 1], 'k--')
ax.set_xlim([0.0, 1.0])
ax.set_ylim([0.0, 1.05])
ax.set_xlabel('False Positive Rate')
ax.set_ylabel('True Positive Rate')
ax.set_title('Receiver operating characteristic example')
ax.legend(loc="lower right")
# ## RGB CNN
# Using the RGB instead of the gray value for the CNN
# +
CONV_SIZE = (10, 10, 3)
def calc_auc_conv2d(rcoefs):
coefs = rcoefs.reshape(CONV_SIZE)/rcoefs.sum()
score_image = relu(convolve(grey_img, coefs))
score_value = score_image.flatten()
fpr2, tpr2, _ = roc_curve(ground_truth_labels, score_value)
return {'fpr': fpr2, 'tpr': tpr2, 'auc': auc(fpr2, tpr2), 'gimg': score_image}
# -
def min_func(rcoefs): return 1-calc_auc_conv2d(rcoefs)['auc']
min_kernel = np.stack([gkern_nd(2, kernlen=CONV_SIZE[0])]*3, -1)
min_start = min_kernel.ravel()
for i in range(10):
min_func(min_start)
opt_res_conv2d = fmin(min_func, min_start, maxfun=50, maxiter=100)
opt_values_conv = calc_auc_conv2d(opt_res_conv2d)
tprOpt_conv = opt_values_conv['tpr']
fprOpt_conv = opt_values_conv['fpr']
roc_aucOpt_conv = opt_values_conv['auc']
out_kernel = opt_res_conv2d.reshape(CONV_SIZE)/opt_res_conv.sum()
fig, ax_all = plt.subplots(3, out_kernel.shape[2], figsize=(10, 10))
for i, (c_ax, d_ax, cd_ax) in enumerate(ax_all.T):
c_ax.imshow(min_kernel[:, :, i])
c_ax.set_title('Initial {}'.format(i))
c_ax.axis('off')
d_ax.imshow(out_kernel[:, :, i])
d_ax.set_title('Updated {}'.format(i))
d_ax.axis('off')
cd_ax.imshow(out_kernel[:, :, i]-min_kernel[:, :, i],
cmap='RdBu', vmin=-1e-3, vmax=1e-3)
cd_ax.set_title('Difference {}'.format(i))
cd_ax.axis('off')
fig, (ax_img, ax) = plt.subplots(1, 2, figsize=(20, 10))
ax_img.imshow(mark_boundaries(
opt_values_conv['gimg'].squeeze(), seg_img), cmap='gray')
ax_img.set_title('Transformed Color Image')
ax.plot(fpr, tpr, label='Raw ROC curve (area = %0.2f)' % roc_auc)
ax.plot(fprOpt, tprOpt, label='Optimized ROC curve (area = %0.2f)' % roc_aucOpt)
ax.plot(fprOpt_conv, tprOpt_conv,
label='RGBCNN Optimized ROC curve (area = %0.2f)' % roc_aucOpt_conv)
ax.plot([0, 1], [0, 1], 'k--')
ax.set_xlim([0.0, 1.0])
ax.set_ylim([0.0, 1.05])
ax.set_xlabel('False Positive Rate')
ax.set_ylabel('True Positive Rate')
ax.set_title('Receiver operating characteristic example')
ax.legend(loc="lower right")
# ### Tasks
# 1. How can you improve filtering in this analysis?
# - Which filter elements might improve the area under the ROC?
# - Try making workflows to test out a few different filters
# 2. Where might morphological operations fit in?
# - How can you make them part of this workflow as well?
# 3. (Challenge) How would you add multiple filter operations? Can you optimize all of the parameters? What problems do you run into as you make your model more complex?
| Exercises/05-Exercises.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Object Detection using TensorFlow - Webcam
# +
import numpy as np
import os
import six.moves.urllib as urllib
import sys
import tarfile
import tensorflow as tf
import zipfile
from utils import label_map_util
from utils import visualization_utils as vis_util
from collections import defaultdict
from io import StringIO
from matplotlib import pyplot as plt
from PIL import Image
# +
# This is needed to display the images.
# %matplotlib inline
# This is needed since the notebook is stored in the object_detection folder.
sys.path.append("..")
# -
# path to the download url
download_url = 'http://download.tensorflow.org/models/object_detection/'
# choosing a model to downlaod
model = 'ssd_mobilenet_v1_coco_11_06_2017'
model_tar = model + '.tar.gz'
# path to the pb file
path = model + '/frozen_inference_graph.pb'
# path to the labels of the pretrained dataset
label_path = os.path.join('data', 'mscoco_label_map.pbtxt')
# 90 classes
classes_num = 90
# opens the tar file and downloads the model to our system
opener = urllib.request.URLopener()
opener.retrieve(download_url + model_tar, model_tar)
file = tarfile.open(model_tar)
print(file)
for each_file in file.getmembers():
each_file_name = os.path.basename(each_file.name)
if 'frozen_inference_graph.pb' in each_file_name:
file.extract(each_file, os.getcwd())
# loading a frozen tensorflow model into memory
graph_detection = tf.Graph()
with graph_detection.as_default():
graph_def = tf.GraphDef()
with tf.gfile.GFile(path, 'rb') as fid:
graph_serialized = fid.read()
graph_def.ParseFromString(graph_serialized)
tf.import_graph_def(graph_def, name='')
# loading labels and their mappings
label_map = label_map_util.load_labelmap(label_path)
categories = label_map_util.convert_label_map_to_categories(label_map, max_num_classes=classes_num, use_display_name=True)
category_index = label_map_util.create_category_index(categories)
import cv2
cap=cv2.VideoCapture(0)
ret = True
with graph_detection.as_default():
with tf.Session(graph=graph_detection) as sess:
while(ret):
ret,image_np=cap.read()
image_np_expanded = np.expand_dims(image_np, axis=0)
image_tensor = graph_detection.get_tensor_by_name('image_tensor:0')
# Each box represents a part of the image where a particular object was detected.
boxes = graph_detection.get_tensor_by_name('detection_boxes:0')
# Each score represent how level of confidence for each of the objects.
# Score is shown on the result image, together with the class label.
scores = graph_detection.get_tensor_by_name('detection_scores:0')
classes = graph_detection.get_tensor_by_name('detection_classes:0')
num_detections = graph_detection.get_tensor_by_name('num_detections:0')
# Actual detection.
(boxes, scores, classes, num_detections) = sess.run(
[boxes, scores, classes, num_detections],
feed_dict={image_tensor: image_np_expanded})
# Visualization of the results of a detection.
vis_util.visualize_boxes_and_labels_on_image_array(
image_np,
np.squeeze(boxes),
np.squeeze(classes).astype(np.int32),
np.squeeze(scores),
category_index,
use_normalized_coordinates=True,
line_thickness=8)
cv2.imshow('image',cv2.resize(image_np,(1280,960)))
if cv2.waitKey(25) & 0xFF==ord('q'):
break
cv2.destroyAllWindows()
cap.release()
| Object_Detection_Webcam.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/gandalf1819/Reinforcement-Learning-Comparative-Study/blob/master/pong.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="HqD11FiBBuA_" colab_type="text"
# Author: <NAME><br>
# Source: https://gist.github.com/karpathy/a4166c7fe253700972fcbc77e4ea32c5 <br>
# Trains an agent with (stochastic) Policy Gradients on Pong. Uses OpenAI Gym.<br>
# + id="5wjlzFGpBzq9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="98e4a661-3710-4e80-8c88-d289c4dda919"
# !pip install cPickle
# + id="48ESaqt2BSnj" colab_type="code" colab={}
import numpy as np
import pickle
import gym
# + id="wt2o0Sy4BulV" colab_type="code" colab={}
# hyperparameters
H = 200 # number of hidden layer neurons
batch_size = 10 # every how many episodes to do a param update?
learning_rate = 1e-4
gamma = 0.99 # discount factor for reward
decay_rate = 0.99 # decay factor for RMSProp leaky sum of grad^2
resume = False # resume from previous checkpoint?
render = False
# + id="K2OoWgIsCGiN" colab_type="code" colab={}
# model initialization
D = 80 * 80 # input dimensionality: 80x80 grid
if resume:
model = pickle.load(open('save.p', 'rb'))
else:
model = {}
model['W1'] = np.random.randn(H,D) / np.sqrt(D) # "Xavier" initialization
model['W2'] = np.random.randn(H) / np.sqrt(H)
grad_buffer = { k : np.zeros_like(v) for k,v in model.items() } # update buffers that add up gradients over a batch
rmsprop_cache = { k : np.zeros_like(v) for k,v in model.items() } # rmsprop memory
# + id="Z1l4rXdOCIPE" colab_type="code" colab={}
def sigmoid(x):
return 1.0 / (1.0 + np.exp(-x)) # sigmoid "squashing" function to interval [0,1]
# + id="bOdt3uQyCRL1" colab_type="code" colab={}
def prepro(I):
""" prepro 210x160x3 uint8 frame into 6400 (80x80) 1D float vector """
I = I[35:195] # crop
I = I[::2,::2,0] # downsample by factor of 2
I[I == 144] = 0 # erase background (background type 1)
I[I == 109] = 0 # erase background (background type 2)
I[I != 0] = 1 # everything else (paddles, ball) just set to 1
return I.astype(np.float).ravel()
# + id="jAXpXGsaCSmF" colab_type="code" colab={}
def discount_rewards(r):
""" take 1D float array of rewards and compute discounted reward """
discounted_r = np.zeros_like(r)
running_add = 0
for t in reversed(range(0, r.size)):
if r[t] != 0: running_add = 0 # reset the sum, since this was a game boundary (pong specific!)
running_add = running_add * gamma + r[t]
discounted_r[t] = running_add
return discounted_r
# + id="mdtAYHfVCU55" colab_type="code" colab={}
def policy_forward(x):
h = np.dot(model['W1'], x)
h[h<0] = 0 # ReLU nonlinearity
logp = np.dot(model['W2'], h)
p = sigmoid(logp)
return p, h # return probability of taking action 2, and hidden state
# + id="2D7xxlBFCWbz" colab_type="code" colab={}
def policy_backward(eph, epdlogp):
""" backward pass. (eph is array of intermediate hidden states) """
dW2 = np.dot(eph.T, epdlogp).ravel()
dh = np.outer(epdlogp, model['W2'])
dh[eph <= 0] = 0 # backpro prelu
dW1 = np.dot(dh.T, epx)
return {'W1':dW1, 'W2':dW2}
# + id="1WqhEKknCX3F" colab_type="code" colab={}
env = gym.make("Pong-v0")
observation = env.reset()
prev_x = None # used in computing the difference frame
xs,hs,dlogps,drs = [],[],[],[]
running_reward = None
reward_sum = 0
episode_number = 0
# + id="3vNCbGRBCZoR" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="2a35a91d-5fc6-4b56-e469-8561b9920dcf"
while True:
if render: env.render()
# preprocess the observation, set input to network to be difference image
cur_x = prepro(observation)
x = cur_x - prev_x if prev_x is not None else np.zeros(D)
prev_x = cur_x
# forward the policy network and sample an action from the returned probability
aprob, h = policy_forward(x)
action = 2 if np.random.uniform() < aprob else 3 # roll the dice!
# record various intermediates (needed later for backprop)
xs.append(x) # observation
hs.append(h) # hidden state
y = 1 if action == 2 else 0 # a "fake label"
dlogps.append(y - aprob) # grad that encourages the action that was taken to be taken (see http://cs231n.github.io/neural-networks-2/#losses if confused)
# step the environment and get new measurements
observation, reward, done, info = env.step(action)
reward_sum += reward
drs.append(reward) # record reward (has to be done after we call step() to get reward for previous action)
if done: # an episode finished
episode_number += 1
# stack together all inputs, hidden states, action gradients, and rewards for this episode
epx = np.vstack(xs)
eph = np.vstack(hs)
epdlogp = np.vstack(dlogps)
epr = np.vstack(drs)
xs,hs,dlogps,drs = [],[],[],[] # reset array memory
# compute the discounted reward backwards through time
discounted_epr = discount_rewards(epr)
# standardize the rewards to be unit normal (helps control the gradient estimator variance)
discounted_epr -= np.mean(discounted_epr)
discounted_epr /= np.std(discounted_epr)
epdlogp *= discounted_epr # modulate the gradient with advantage (PG magic happens right here.)
grad = policy_backward(eph, epdlogp)
for k in model: grad_buffer[k] += grad[k] # accumulate grad over batch
# perform rmsprop parameter update every batch_size episodes
if episode_number % batch_size == 0:
for k,v in model.items():
g = grad_buffer[k] # gradient
rmsprop_cache[k] = decay_rate * rmsprop_cache[k] + (1 - decay_rate) * g**2
model[k] += learning_rate * g / (np.sqrt(rmsprop_cache[k]) + 1e-5)
grad_buffer[k] = np.zeros_like(v) # reset batch gradient buffer
# boring book-keeping
running_reward = reward_sum if running_reward is None else running_reward * 0.99 + reward_sum * 0.01
print ('resetting env. episode reward total was %f. running mean: %f' % (reward_sum, running_reward))
if episode_number % 100 == 0: pickle.dump(model, open('save.p', 'wb'))
reward_sum = 0
observation = env.reset() # reset env
prev_x = None
if reward != 0: # Pong has either +1 or -1 reward exactly when game ends.
print ('ep %d: game finished, reward: %f' % (episode_number, reward) + ('' if reward == -1 else ' !!!!!!!!'))
# + id="yu0KkQgHCc6Z" colab_type="code" colab={}
| pong.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 넘파이, 판다스, 매트플랏립 실습
# 그래프를 바로 보기 위한 명령어
# %matplotlib inline
import numpy as np # 넘파이 탑재
import pandas as pd # 판다스 탑재
import matplotlib # 매트플랏립 탑재
# 50개의 난수로 이루어진 데이터 생성
data = np.random.rand(50)
data
# 판다스 시리즈 타입 데이터로 변환
seri = pd.Series(data)
seri
# 선형 그래프 그리기
seri.plot()
| jupyter_notebook/11-8.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Análisis e Implementación en SparkQL
#
# * Entrega hasta el 24 de mayo en https://forms.gle/aRiWtjayausRFCTd8
#
# * Los dos conjuntos de datos entregados, CSV separados por coma, Evaluación del Pitch (2021.04.13 ISoftware).csv y Asistencia del Pitch (2021.04.13 ISoftware).csv proceden de dos encuestas realizadas con Google Forms.
#
# * Los archivos proceden de las evaluaciones de los pitch (exposiciones cortas de negocio) que realizan los estudiantes de la asignatura de Ingeniería de Software. Los estudiantes están organizados en equipos. Cada equipo realiza una presentación de máximo 15 minutos, una vez inicia la presentación todos los estudiantes deben registrar su asistencia lo cual queda registrado en 'Asistencia del Pitch (2021.04.13 ISoftware).csv' incluidos los miembros del equipo.
#
# * El archivo `Asistencia del Pitch (2021.04.13 ISoftware).csv` contiene las columnas: `"Marca temporal"` es tiempo dado en fecha y hora, `"Nombre de usuario"` es el correo electrónico del estudiante y es texto,`"Equipo al que perteneces:"` equipo de trabajo al que pertenece el estudiante también es texto y `"Equipo que va a exponer:"` equipo que el estudiante va a tender a su presentación.
#
# * El archivo `Evaluación del Pitch (2021.04.13 ISoftware).csv` contiene las columnas `"Marca temporal"` es tiempo dado en fecha y hora, `"Nombre de usuario"` es el correo electrónico del estudiante y es texto, `"Equipo que vas a evaluar:"` equipo que ha expuesto y que va a ser evaluado por cada estudiante que no sea integrante; a continuación, se tienen las siguientes columnas que corresponden a la evaluación de los respectivos ítems:
#
# 1. "Introducción: El equipo responde adecuadamente ¿Quiénes son y por qué están aquí?",
# 2. "Equipo: El equipo responde adecuadamente ¿Quiénes están detrás de la idea y cuál es su función?",
# 3. "Problema: El equipo responde adecuadamente ¿Qué problema resolverá?, ¿es realmente un problema?",
# 4. "Ventajas: El equipo responde adecuadamente ¿Por qué su solución es especial?, ¿qué la hace distinta de otras?",
# 5. "Solución: El equipo responde adecuadamente ¿Cómo piensa resolver el problema?",
# 6. "Producto: El equipo responde adecuadamente ¿Cómo funciona el producto o servicio? Muestra algunos ejemplos.",
# 7. "Tracción: El equipo responde adecuadamente si cuenta con clientes que demuestran potencial.",
# 8. "Mercado: El equipo responde conoce, o por lo menos intentar predecir, el tamaño del mercado que impactará.",
# 9. "Competencia: El equipo responde adecuadamente ¿Cuáles son las soluciones alternativas al problema que plantea?",
# 10. "Modelo de negocio: El equipo responde adecuadamente ¿Cómo hará dinero? ",
# 11. "Inversión: El equipo responde adecuadamente ¿Cuál es su presupuesto y cuánto espera ganar?",
# 12. "Contacto: El equipo deja los datos al cliente y muestra cómo pueden contactarle.",
# 13. "Exposición: ¿Qué tan coordinados estaban los expositores?",
# 14. "Exposición: ¿Los expositores se expresaron con claridad y se hicieron entender?",
# 15. "Exposición: Las diapositivas son claras y coherentes y apoyaron adecuadamente la exposición.",
# * "Suponiendo que eres inversionista, ¿Estarías dispuesto a invertir dinero en este equipo? (esta pregunta no se pondera en la nota)",
# * "Observaciones para el equipo, estas observaciones las debe considerar el equipo para mejorar la siguiente presentación."
#
#
# * Cada ítem se evalúa con la siguiente escala: 0. Ausente; 1. Deficiente; 2. Regular; 3. Aceptable; 4. Bueno; 5. Excelente
#
# * Carge los datos, cada archivo en una tabla SparkSQL y responda cada una de las consultas dadas en cada celda. Tenga en cuenta que algunas consultas pueden tener como resultado el vacío.
# # Integrantes del equipo
# 1. <NAME>
# +
# Agrege acá el código para importar las librerias
import pandas as pd
from pyspark.sql.functions import *
from pyspark.sql import functions as F
from pyspark.sql.types import IntegerType
from pyspark.sql.functions import concat_ws
from pyspark.sql.functions import to_timestamp
from pyspark.sql.functions import collect_list
from pyspark.sql.functions import regexp_replace
# La libreria para "encontrar el sevicio" de Spark
import findspark
findspark.init()
# Librerias para "gestionar el servicio" de Spark
from pyspark import SparkConf, SparkContext
from pyspark.sql import SQLContext, SparkSession
# Creamos una aplicación Spark en el Servicio
# Tenga cuidado con las tildes o caracteres especiales en el nombre de la app
AppSpark = SparkConf().setAppName("Evaluacion iSofware")
# definimos un espacio o contexto para la App
ContextoSpark=SparkContext(conf=AppSpark)
# inicio una sesión en el espacio de la App
SesionSpark = SparkSession(ContextoSpark)
# inicio del espacio o contexto SQL
ContextoSql = SQLContext(sparkContext=ContextoSpark, sparkSession=SesionSpark)
# +
# 1. Cargue los datos en la carpeta datalake y luego del /datalake al HDFS (Hadoop File System)
# Recuerda usar ! para ejecutar el comando en el shell.
# Tu código a continuación...
# !hdfs dfs -copyFromLocal 'asistencia.csv' /tmp
# -
# !hdfs dfs -copyFromLocal 'evaluacion.csv' /tmp
# !hadoop fs -ls /tmp
# +
# 2. Cree dos tablas SparkSQL y almacene el csv en su correspondiente tabla.
# Observación: tenga especial cuidado con los encabezados de los archivos CSV.
# Usted puede considerar cambiar los encabezados de los CSV originales
# Tu código a continuación...
# Se carga archivo de asistencia
asistencia = ContextoSql.read.load('/tmp/asistencia.csv',
format="csv",
sep=',',
inferSchema='true',
header='true')
# Se carga archivo de evaluación
evaluacion = ContextoSql.read.load('/tmp/evaluacion.csv',
format="csv",
sep=',',
inferSchema='true',
header='true')
# -
asistencia.printSchema()
evaluacion.printSchema()
# +
# 2. Consulte el listado total de estudiantes (correos electrónicos) del
# curso de Ingeniería de Software, ordenados alfabéticamente
# Tu código a continuación...
evaluacion = evaluacion.filter(evaluacion["Nombre de usuario"].contains("@"))
# Se realiza trim de evaluacion
trimmed_evaluacion = trim(evaluacion['Nombre de usuario'])
evaluacion = evaluacion.withColumn('Nombre de usuario', trimmed_evaluacion)
evaluacion.select('Nombre de usuario').distinct().orderBy(
'Nombre de usuario', ascending=True).show(10000)
# +
# 3. Consulte la cantidad de asistencias registradas por estudiante; además, la fecha y hora de la primera asistencia
# y la fecha y hora de la última asistencia
# Tu código a continuación...
# Conversión a formato fecha
asistencia = asistencia.withColumn(
"Marca temporal", (to_timestamp(asistencia["Marca temporal"],
'yyyy/MM/dd HH:mm:ss')))
# Conteo asistencia
asistencia.groupBy("Nombre de usuario").agg(
F.count('Nombre de usuario').alias("Número de asistencias"),
F.max("Marca temporal").alias("Última asistencia"),
F.min("Marca temporal").alias("Primera asistencia")).orderBy(
["Número de asistencias","Nombre de usuario"],
ascending=[False,True]).show(3000)
# +
# 4. Consulte el listado de estudiantes que asistieron a 2 presentaciones o menos (una).
# Tu código a continuación...
# Grupo de asistencias
asistencias = asistencia.groupBy("Nombre de usuario").agg(
F.count('Nombre de usuario').alias("Número de asistencias"),
F.max("Marca temporal").alias("Última asistencia"),
F.min("Marca temporal").alias("Primera asistencia")).orderBy(
["Número de asistencias","Nombre de usuario"], ascending=[False, True])
# Listado de menos de 2 presentaciones
asistencias.withColumn(
"Número de asistencias",
asistencias["Número de asistencias"].cast(IntegerType())).where(
asistencias['Número de asistencias']<=2).show(1000)
# +
# 5. Consulte el listado de estudiantes que no asistieron a ninguna presentación.
# Tu código a continuación...
# Estudiantes que no asistieron a presentaciones
no_presentacion_est = evaluacion.select(
'Nombre de usuario').distinct().orderBy(
'Nombre de usuario', ascending=True)
no_presentacion_est.join(grupo_asistencias,
on=["Nombre de usuario"],
how = 'left_anti').show(10)
# +
# 6. Consulte los integrantes por cada equipo al que pertenecen.
# Tu código a continuación...
# Reemplazos en columnas
integrantes_equipo = asistencia.withColumn(
"Equipo al que perteneces:",
regexp_replace("Equipo al que perteneces:",
'Pertenezco a:', ''))
# Reemplazos en columnas
integrantes_equipo = integrantes_equipo.withColumn(
"Equipo que va a exponer:",
regexp_replace("Equipo que va a exponer:", 'Expone: ', ""))
# Se envía a tabla en pandas
integrantes_equipo.groupby(
"Equipo al que perteneces:").agg(concat_ws(
", ",collect_list("Nombre de usuario")).alias("Integrantes")).toPandas()
# +
# 7. Consulte la cantidad de asistentes por presentación, sin considerar los asistentes que pertenecen
# al equipo que realizó la presentación.
# Tu código a continuación...
integrantes_equipo_2 = integrantes_equipo.withColumn(
"Equipo que va a exponer:",
trim(integrantes_equipo["Equipo que va a exponer:"]))
integrantes_equipo_2 = integrantes_equipo_2.withColumn(
"Equipo al que perteneces:",
trim(integrantes_equipo_2["Equipo al que perteneces:"]))
integrantes_equipo_2.where(
integrantes_equipo_2["Equipo que va a exponer:"] !=
integrantes_equipo_2["Equipo al que perteneces:"]).groupBy(
"Equipo que va a exponer:").agg(
F.count("Nombre de usuario").alias(
"Asistentes")).orderBy(
"Asistentes", ascending=False).show(1000)
# +
# 8. Consutar cuáles integrantes evaluaron a su propio equipo. Estas evaluaciones no serán válidas, pues un
# integrante no puede evaluar a su propio equipo.
# Tu código a continuación...
evaluacion_propia = evaluacion.withColumn(
'Equipo que vas a evaluar:',
trim(Evaluacion['Equipo que vas a evaluar:']))
evaluacion_propia = evaluacion_propia.select(
'Nombre de usuario', 'Equipo que vas a evaluar:').distinct()
equipos = evaluacion_propia.join(integrantes_equipo_2.select(
'Nombre de usuario', 'Equipo al que perteneces:').distinct(),
on=['Nombre de usuario'],
how='left')
equipos.filter(equipos['Equipo que vas a evaluar:'] == equipos['Equipo al que perteneces:']
).orderBy('Nombre de usuario').show(1000)
# +
# 9. Consultar la nota promedio por cada ítem (1 al 15), y la nota promedio total del cada equipo. Recuerde que no
# son válidas las evaluaciones realizadas por los miembros del mismo equipo.
# Tu código a continuación...
evaluacion_equipo = evaluacion.withColumn('Equipo que vas a evaluar:',
trim(Evaluacion['Equipo que vas a evaluar:']))
evaluacion_propia = evaluacion_propia.select(
'Nombre de usuario', 'Equipo que vas a evaluar:',
evaluacion_equipo['Introducción: El equipo responde adecuadamente ¿Quiénes son y por qué están aquí?'].alias('Introducción'),
evaluacion_equipo['Equipo: El equipo responde adecuadamente ¿Quiénes están detrás de la idea y cuál es su función?'].alias('Equipo'),
evaluacion_equipo['Problema: El equipo responde adecuadamente ¿Qué problema resolverá?, ¿es realmente un problema?'].alias('Problema'),
evaluacion_equipo['Ventajas: El equipo responde adecuadamente ¿Por qué su solución es especial?, ¿qué la hace distinta de otras?'].alias('Ventajas'),
evaluacion_equipo['Solución: El equipo responde adecuadamente ¿Cómo piensa resolver el problema?'].alias('Solución'),
evaluacion_equipo["`Producto: El equipo responde adecuadamente ¿Cómo funciona el producto o servicio? Muestra algunos ejemplos.`"].alias('Producto'),
evaluacion_equipo["`Tracción: El equipo responde adecuadamente si cuenta con clientes que demuestran potencial.`"].alias('Tracción'),
evaluacion_equipo["`Mercado: El equipo responde conoce, o por lo menos intentar predecir, el tamaño del mercado que impactará.`"].alias('Mercado') ,
evaluacion_equipo["`Competencia: El equipo responde adecuadamente ¿Cuáles son las soluciones alternativas al problema que plantea?`"].alias('Competencia'),
evaluacion_equipo["`Modelo de negocio: El equipo responde adecuadamente ¿Cómo hará dinero? `"].alias('Modelo de negocio'),
evaluacion_equipo["`Inversión: El equipo responde adecuadamente ¿Cuál es su presupuesto y cuánto espera ganar?`"].alias('Inversión'),
evaluacion_equipo["`Contacto: El equipo deja los datos al cliente y muestra cómo pueden contactarle.`"].alias('Contacto'),
evaluacion_equipo["`Exposición: ¿Los expositores se expresaron con claridad y se hicieron entender?`"].alias('Exposicion1'),
evaluacion_equipo["`Exposición: ¿Los expositores se expresaron con claridad y se hicieron entender?`"].alias('Exposicion2'),
evaluacion_equipo["`Exposición: Las diapositivas son claras y coherentes y apoyaron adecuadamente la exposición.`"].alias('Exposicion3'))
Equipos = evaluacion_equipo.join(
integrantes_equipo_2.select(
'Nombre de usuario', 'Equipo al que perteneces:').distinct(),
on=['Nombre de usuario'],
how='left')
evaluaciones = equipos.filter((equipos['Equipo que vas a evaluar:'] !=
equipos['Equipo al que perteneces:']))
promedios = evaluaciones.groupBy('Equipo que vas a evaluar:').agg(
F.avg('Introducción'), F.avg('Equipo'), F.avg('Problema'),
F.avg('Ventajas'), F.avg('Solución'), F.avg('Producto'),
F.avg('Tracción'), F.avg('Mercado'), F.avg('Competencia'),
F.avg('Modelo de negocio'), F.avg('Inversión'), F.avg('Contacto'),
F.avg('Exposicion1'), F.avg('Exposicion2'), F.avg('Exposicion3'))
nota = promedios.select(
'*',
((promedios["`avg(Introducción)`"] + promedios["`avg(Equipo)`"] +
promedios["`avg(Problema)`"] + promedios["`avg(Ventajas)`"] +
promedios["`avg(Solución)`"] + promedios["`avg(Producto)`"] +
promedios["`avg(Tracción)`"] + promedios["`avg(Mercado)`"] +
promedios["`avg(Competencia)`"] + promedios["`avg(Modelo de negocio)`"]
+ promedios["`avg(Inversión)`"] + promedios["`avg(Contacto)`"] +
promedios["`avg(Exposicion1)`"] + promedios["`avg(Exposicion2)`"] +
promedios["`avg(Exposicion3)`"])/15).alias('Nota final'))
nota.toPandas()
# +
# 10. Consulte el mejor equipo evaluado por cada ítem (según nota promedio. En caso de empate mostrar todos los empatados)
# y el mejor equipo según el promedio total.
# Tu código a continuación...
def consultar_mejor_equipo(colname):
nota.where(nota[colname] == nota.agg({colname: "max"}).collect()[0][0]).select(
'Equipo que vas a evaluar:',colname).show()
for col in nota.columns[1:]:
consultar_mejor_equipo(col)
# +
# 11. Consulte el peor equipo evaluado por cada ítem (según nota promedio. En caso de empate mostrar todos los empatados)
# y el peor equipo según el promedio total.
# Tu código a continuación...
def consultar_peor_equipo(colname):
nota.where(nota[colname] == nota.agg({colname: "min"}).collect()[0][0]).select(
'Equipo que vas a evaluar:',colname).show()
for col in nota.columns[1:]:
consultar_peor_equipo(col)
# +
# 12. Consulte la lista de estudiantes con la correspondiente nota obtenida en la presentación
# (nota promedio total de la evaluación obtenida por el equipo)
# Tu código a continuación...
integrantes_equipo_3 = integrantes_equipo.withColumn(
'Equipo al que perteneces:',
trim(integrantes_equipo['Equipo al que perteneces:']))
integrantes_equipo_3.select(
'Nombre de usuario',
'Equipo al que perteneces:').distinct().join(
nota,
on=(integrantes_equipo_3['Equipo al que perteneces:'] == nota['Equipo que vas a evaluar:']),
how='left').select('Nombre de usuario', 'Equipo al que perteneces:', 'Nota final').show()
| Trabajo_Spark/TrabajoSparkQL.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
from sklearn.model_selection import train_test_split
from lvxnn_test import lvxnn
from xgb_test import xgb
from svd_test import svd
from collections import OrderedDict
from deepfm_fm_test import deepfm_fm
import time
from collections import OrderedDict
import tensorflow as tf
tf.compat.v1.enable_eager_execution()
from sklearn.metrics import mean_squared_error,roc_auc_score,mean_absolute_error,log_loss
import sys
sys.path.append('../../')
from lvxnn.LVXNN import LV_XNN
from lvxnn.DataReader import data_initialize
data= pd.read_csv('../simulation/data/sim_0.9.csv')
train , test = train_test_split(data,test_size=0.2,random_state=0)
# +
meta_info = OrderedDict()
meta_info['uf_1']={'type': 'continues','source':'user'}
meta_info['uf_2']={'type': 'continues','source':'user'}
meta_info['uf_3']={'type': 'continues','source':'user'}
meta_info['uf_4']={'type': 'continues','source':'user'}
meta_info['uf_5']={'type': 'continues','source':'user'}
meta_info['if_1']={'type': 'continues','source':'item'}
meta_info['if_2']={'type': 'continues','source':'item'}
meta_info['if_3']={'type': 'continues','source':'item'}
meta_info['if_4']={'type': 'continues','source':'item'}
meta_info['if_5']={'type': 'continues','source':'item'}
meta_info['user_id']={"type":"id",'source':'user'}
meta_info['item_id']={"type":"id",'source':'item'}
meta_info['target']={"type":"target",'source':''}
# -
result_lvxnn, result_gami = lvxnn(train, test, meta_info, task_type="Regression", val_ratio=0.2, random_state=0)
result_svd = svd(train, test, meta_info, task_type="Regression", val_ratio=0.2, random_state=0)
result_xgb = xgb(train, test, meta_info, task_type="Regression", val_ratio=0.2, random_state=0)
result_deepfm, result_fm = deepfm_fm(train, test, meta_info, task_type="Regression", val_ratio=0.2, random_state=0)
result_fm
| scripts/benchmark/.ipynb_checkpoints/benchmark_test-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from ivi.rigol.rigolDS1052D import rigolDS1052D
with rigolDS1052D("USB0::6833::1416::INSTR") as scope:
pass
from PIL import Image
from io import BytesIO
with rigolDS1052D("USB0::fdf8:f53e:61e4::18::INSTR") as scope:
screenshot = Image.open(BytesIO(scope.display.fetch_screenshot()))
screenshot
| DevelopmentNotebooks/ivi_dev.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + deletable=false editable=false heading_collapsed=true run_control={"frozen": true}
# # CE-40717: Machine Learning
# + deletable=false editable=false heading_collapsed=true hidden=true run_control={"frozen": true}
# ## HW8-Clustering & Reinforcement Learning
#
# <NAME> - 99210259
# + deletable=false editable=false heading_collapsed=true hidden=true run_control={"frozen": true}
# ### Kmeans & GMM:
#
# At this question, we tend to implement Kmeans & GMM algorithms. For this purpose, `DO NOT EMPLOY` ready-for-use python libraries. Use this implementation for solving the following questions. Kmeans should continue till centeroids won't change. Furthermore, GMM also should continue till the difference of two consecutive likelihood logarithm would be less than 0.1. Notice that after executing the Kmeans part, the primitive centroids of GMM should be identical with ultimate Kmeans centroids.
# + hidden=true
from sklearn.datasets.samples_generator import make_classification, make_moons, make_circles
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# + deletable=false editable=false heading_collapsed=true hidden=true run_control={"frozen": true}
# #### Part 1:
#
# Utilize the subsequent cell in order to create the Dataset. Afterwards, try to execute the algorithm with k=2 centroids. At Kmeans, it is recommended to execute the algorithm with several various starting states in order to eventually choose the best respective result.
# + hidden=true
X,Y = make_classification(n_samples=700, n_features=10, n_informative=5,
n_redundant=0, n_clusters_per_class=2, n_classes=3)
# -
# ## KMeans Implementation
class KMeans:
def __init__(self, n_clusters = 3, tolerance = 0.01, max_iter = 100, runs = 1):
self.n_clusters = n_clusters
self.tolerance = tolerance
self.cluster_means = np.zeros(n_clusters)
self.max_iter = max_iter
self.runs = runs
def fit(self, X,Y):
row_count, col_count = X.shape
X_values = self.__get_values(X)
X_labels = np.zeros(row_count)
costs = np.zeros(self.runs)
all_clusterings = []
for i in range(self.runs):
cluster_means = self.__initialize_means(X_values, row_count)
for _ in range(self.max_iter):
previous_means = np.copy(cluster_means)
distances = self.__compute_distances(X_values, cluster_means, row_count)
X_labels = self.__label_examples(distances)
cluster_means = self.__compute_means(X_values, X_labels, col_count)
clusters_not_changed = np.abs(cluster_means - previous_means) < self.tolerance
if np.all(clusters_not_changed) != False:
break
X_values_with_labels = np.append(X_values, X_labels[:, np.newaxis], axis = 1)
all_clusterings.append( (cluster_means, X_values_with_labels) )
costs[i] = self.__compute_cost(X_values, X_labels, cluster_means)
best_clustering_index = costs.argmin()
self.costs = costs
self.cost_ = costs[best_clustering_index]
self.centroid,self.items = all_clusterings[best_clustering_index]
self.y = Y
return all_clusterings[best_clustering_index]
def __initialize_means(self, X, row_count):
return X [ np.random.choice(row_count, size=self.n_clusters, replace=False) ]
def __compute_distances(self, X, cluster_means, row_count):
distances = np.zeros((row_count, self.n_clusters))
for cluster_mean_index, cluster_mean in enumerate(cluster_means):
distances[:, cluster_mean_index] = np.linalg.norm(X - cluster_mean, axis = 1)
return distances
def __label_examples(self, distances):
return distances.argmin(axis = 1)
def __compute_means(self, X, labels, col_count):
cluster_means = np.zeros((self.n_clusters, col_count))
for cluster_mean_index, _ in enumerate(cluster_means):
cluster_elements = X [ labels == cluster_mean_index ]
if len(cluster_elements):
cluster_means[cluster_mean_index, :] = cluster_elements.mean(axis = 0)
return cluster_means
def __compute_cost(self, X, labels, cluster_means):
cost = 0
for cluster_mean_index, cluster_mean in enumerate(cluster_means):
cluster_elements = X [ labels == cluster_mean_index ]
cost += np.linalg.norm(cluster_elements - cluster_mean, axis = 1).sum()
return cost
def __get_values(self, X):
if isinstance(X, np.ndarray):
return X
return np.array(X)
def predict(self):
data=pd.DataFrame(self.items)
added_column=list(data.columns)[-1]
data['Label'] = self.y
resultOfClustering=data.groupby([added_column])['Label'].agg(lambda x: x.value_counts().index[0])
mapping = dict()
for label in range(self.n_clusters):
label_predicted = resultOfClustering[label]
mapping[label] = label_predicted
data['PredictedLabels']=data[added_column].map(mapping)
return np.array(data['PredictedLabels'])
kmeans=KMeans(2,max_iter=10000,runs=20)
centroids,kmeans_items=kmeans.fit(X,Y)
plt.plot(np.arange(len(kmeans.costs)),kmeans.costs)
plt.title('error of different runs')
plt.xticks(np.arange(len(kmeans.costs)))
plt.show();
# ## Gaussian Mixture Model Implementation
# +
import numpy as np
import scipy.stats as sp
class GaussianMixModel():
def __init__(self, X, k=2):
X = np.asarray(X)
self.m, self.n = X.shape
self.data = X.copy()
self.k = k
self.sigma_arr = np.array([np.asmatrix(np.identity(self.n)) for i in range(self.k)])
self.phi = np.ones(self.k)/self.k
self.Z = np.asmatrix(np.empty((self.m, self.k), dtype=float))
def initialize_means(self,means):
self.mean_arr = means
def fit(self, tol=0.1):
num_iters = 0
logl = 1
previous_logl = 0
while(logl-previous_logl > tol):
previous_logl = self.loglikelihood()
self.e_step()
self.m_step()
num_iters += 1
logl = self.loglikelihood()
print('Iteration %d: log-likelihood is %.6f'%(num_iters, logl))
print('Terminate at %d-th iteration:log-likelihood is %.6f'%(num_iters, logl))
def loglikelihood(self):
logl = 0
for i in range(self.m):
tmp = 0
for j in range(self.k):
tmp += sp.multivariate_normal.pdf(self.data[i, :],self.mean_arr[j, :].A1,self.sigma_arr[j, :]) * self.phi[j]
logl += np.log(tmp)
return logl
def e_step(self):
for i in range(self.m):
den = 0
for j in range(self.k):
num = sp.multivariate_normal.pdf(self.data[i, :],
self.mean_arr[j].A1,
self.sigma_arr[j]) *\
self.phi[j]
den += num
self.Z[i, j] = num
self.Z[i, :] /= den
assert self.Z[i, :].sum() - 1 < 1e-4 # Program stop if this condition is false
def m_step(self):
for j in range(self.k):
const = self.Z[:, j].sum()
self.phi[j] = 1/self.m * const
_mu_j = np.zeros(self.n)
_sigma_j = np.zeros((self.n, self.n))
for i in range(self.m):
_mu_j += (self.data[i, :] * self.Z[i, j])
_sigma_j += self.Z[i, j] * ((self.data[i, :] - self.mean_arr[j, :]).T * (self.data[i, :] - self.mean_arr[j, :]))
self.mean_arr[j] = _mu_j / const
self.sigma_arr[j] = _sigma_j / const
def predict(self):
return np.array(np.argmax(gmm.Z,axis=1)).flatten()
# -
gmm=GaussianMixModel(X,k=2)
gmm.initialize_means(np.asmatrix(centroids))
gmm.fit()
# + deletable=false editable=false heading_collapsed=true hidden=true run_control={"frozen": true}
# #### Part 2:
#
# In a separated cell, implement `Purity` and `Rand-Index` criteria in order to compare the performance of mentioned algorithms.
# -
# ## KMeans
print('Purity Of kmeans: ',np.sum(kmeans.predict()==Y)/len(Y))
from scipy.special import comb
def rand_index_score(clusters, classes):
A = np.c_[(clusters, classes)]
tp = sum(comb(np.bincount(A[A[:, 0] == i, 1]), 2).sum()
for i in set(clusters))
fp = comb(np.bincount(clusters), 2).sum() - tp
fn = comb(np.bincount(classes), 2).sum() - tp
tn = comb(len(A), 2) - tp - fp - fn
return (tp + tn) / (tp + fp + fn + tn)
print('rand index of kmeans', rand_index_score(kmeans.predict(),Y))
# ## Gaussian Mixture Model
print('purity index: ', np.sum(gmm.predict() == Y)/len(Y))
print('rand index', rand_index_score(gmm.predict(),Y))
# + deletable=false editable=false heading_collapsed=true hidden=true run_control={"frozen": true}
# #### Part 3:
#
# Use the following cell in order to create new Datasets. Afterwards, try to execute mentioned algorithms on new Dataset and eventually compare the recent results with the help of visualization(there is no problem for using relevant python libraries like `matplotlib`). Consider two clusters for this part.
# + hidden=true
X, Y = make_classification(n_samples=700, n_features=2, n_informative=2, n_redundant=0, n_classes=2)
# + hidden=true
k=2
kmeans=KMeans(k,max_iter=10000,runs=20)
centroids,kmeans_items=kmeans.fit(X,Y)
color_s =["green","blue","navy","maroon",'orange']
for i in range(k):
plt.scatter(kmeans_items[kmeans_items[:,2]==i,0] , kmeans_items[kmeans_items[:,2]==i,1]
,s=100, label = "cluster "+str(i), color =color_s[i])
plt.scatter(centroids[:,0] , centroids[:,1] , s = 300, color = 'red')
plt.title('Our clusters')
plt.show();
# +
gmm=GaussianMixModel(X,k)
gmm.initialize_means(np.asmatrix(centroids))
gmm.fit();
gmm_result = gmm.predict()
data=pd.DataFrame(X)
data['Predicted'] = gmm_result
for i in range(k):
plt.scatter(data[data['Predicted']==i][0], data[data['Predicted']==i][1]
,s=100, label = "cluster "+str(i), color =color_s[i])
plt.scatter(np.array(gmm.mean_arr[:,0]).flatten() , np.array(gmm.mean_arr[:,1]).flatten() , s = 300, color = 'red')
plt.show();
# + hidden=true
X, Y = make_moons(n_samples=700, noise=0.2)
# + hidden=true
k=2
kmeans=KMeans(k,max_iter=10000,runs=20)
centroids,kmeans_items=kmeans.fit(X,Y)
color_s =["green","blue","navy","maroon",'orange']
for i in range(k):
plt.scatter(kmeans_items[kmeans_items[:,2]==i,0] , kmeans_items[kmeans_items[:,2]==i,1]
,s=100, label = "cluster "+str(i), color =color_s[i])
plt.scatter(centroids[:,0] , centroids[:,1] , s = 300, color = 'red')
plt.title('Our clusters')
plt.show();
# +
gmm=GaussianMixModel(X,k)
gmm.initialize_means(np.asmatrix(centroids))
gmm.fit();
gmm_result = gmm.predict()
data=pd.DataFrame(X)
data['Predicted'] = gmm_result
for i in range(k):
plt.scatter(data[data['Predicted']==i][0], data[data['Predicted']==i][1]
,s=100, label = "cluster "+str(i), color =color_s[i])
plt.scatter(np.array(gmm.mean_arr[:,0]).flatten() , np.array(gmm.mean_arr[:,1]).flatten() , s = 300, color = 'red')
plt.show();
# + hidden=true
X, Y = make_circles(n_samples=700, noise=0.2)
# + hidden=true
k=2
kmeans=KMeans(k,max_iter=10000,runs=20)
centroids,kmeans_items=kmeans.fit(X,Y)
color_s =["green","blue","navy","maroon",'orange']
for i in range(k):
plt.scatter(kmeans_items[kmeans_items[:,2]==i,0] , kmeans_items[kmeans_items[:,2]==i,1]
,s=100, label = "cluster "+str(i), color =color_s[i])
plt.scatter(centroids[:,0] , centroids[:,1] , s = 300, color = 'red')
plt.title('Our clusters')
plt.show();
# +
gmm=GaussianMixModel(X,k)
gmm.initialize_means(np.asmatrix(centroids))
gmm.fit();
gmm_result = gmm.predict()
data=pd.DataFrame(X)
data['Predicted'] = gmm_result
for i in range(k):
plt.scatter(data[data['Predicted']==i][0], data[data['Predicted']==i][1]
,s=100, label = "cluster "+str(i), color =color_s[i])
plt.scatter(np.array(gmm.mean_arr[:,0]).flatten() , np.array(gmm.mean_arr[:,1]).flatten() , s = 300, color = 'red')
plt.show();
# + deletable=false editable=false heading_collapsed=true hidden=true run_control={"frozen": true}
# ### Reinforcement Learning:
#
# At the bellow cell, besides the required libraries have been imported, feel free for changing the num_states variable with your desired number.
# + hidden=true
import numpy as np
import random
import gym
# + hidden=true
env = gym.make("MountainCar-v0")
num_actions = 3
num_states = 50
# first I should note that first one is position and second one is velocity!
# so each state should be recognized using two discretized states
q_table = np.zeros(shape=(num_states,num_states, num_actions))
# You may change the inputs of any function as you desire.
SPACE_LOW = env.observation_space.low
SPACE_HIGH = env.observation_space.high
DISCOUNT_FACTOR = 0.95
EXPLORATION = 0.15
EPISODES = 100000
STEP_COUNT_MAX = 20000
DISPLAY=False
# + deletable=false editable=false heading_collapsed=true hidden=true run_control={"frozen": true}
# #### Part 1:
#
# Next cell wants you supplement two functions. First for transforming the continuous space into discrete one (in order to make using q_table feasible), second for updating q_values based on the last action done by agent.
# + hidden=true
def discretize_state():
return np.abs(SPACE_HIGH-SPACE_LOW)/num_states
def env_state_to_Q_state(state):
return np.round((state - SPACE_LOW)/discretize_state()).astype(int)
#p is position , v is velocity, p_ is position_new, v_ is velocity_new
def update_q(p, v, p_, v_, action, eta, reward):
if np.random.uniform(0,1) < EXPLORATION:
action_after = np.random.choice(env.action_space.n)
else:
action_after = np.argmax(q_table[p_][v_])
q_table[p][v][action] = q_table[p][v][action] + eta * (reward + DISCOUNT_FACTOR * q_table[p_][v_][action_after] - q_table[p][v][action])
# + deletable=false editable=false heading_collapsed=true hidden=true run_control={"frozen": true}
# #### Part 2:
#
# At the following cell, the ends of two functions are getting current action based on the policy and defining the training process respectively.
# + hidden=true
# You may change the inputs of any function as you desire.
def get_action():
global EXPLORATION
sum_reward_every_thousand = 0
eta = 0.1
for episode in range(EPISODES):
state = env.reset()
sum_reward = 0
if episode < 30000:
EXPLORATION = 0.15
eta = 0.1
else:
EXPLORATION = (0.15)* (0.99)**((episode-30000)//100)
eta = (0.1) * (0.99)**((episode-30000)//10000)
for step in range(STEP_COUNT_MAX):
if episode % 1000 == 1 and DISPLAY:
env.render()
p, v = env_state_to_Q_state(state)
if np.random.uniform(0, 1) < EXPLORATION:
action = np.random.choice(env.action_space.n)
else:
action = np.argmax(q_table[p][v])
state, reward, done, _ = env.step(action)
sum_reward += reward
p_, v_ = env_state_to_Q_state(state)
update_q(p,v,p_, v_, action, eta, reward)
if done:
break
sum_reward_every_thousand+= sum_reward
if episode % 1000 == 1:
print(f'Episode: {episode}, Total Reward: {sum_reward}, Mean Reward for previous thousand: {sum_reward_every_thousand/1000}')
sum_reward_every_thousand=0
def q_learning():
return np.argmax(q_table, axis=2)
def save_policy():
np.save('policy.npy', q_learning())
# -
get_action()
save_policy()
# + deletable=false editable=false heading_collapsed=true hidden=true run_control={"frozen": true}
# #### Part 3:
#
# Ultimately, the score function examines the average performance of Agent (after nearly 1000 times) based on previous implementations.
# + hidden=true tags=["outputPrepend"]
# Attention: don't change this function. we will use this to grade your policy which you will hand in with policy.npy
# btw you can use it to see how you are performing. Uncomment two lines which are commented to be able to see what is happening visually.
def score():
policy, scores = np.load("policy.npy"), []
for episode in range(1000):
print(f"******Episode {episode}")
state, score, done, step = env_state_to_Q_state(env.reset()), 0, False, 0
while not done:
# time.sleep(0.04)
p,v = state
action = policy[p,v]
state, reward, done, _ = env.step(action)
state = env_state_to_Q_state(state)
step += 1
score += int(reward)
print(f"Score:{score}")
scores.append(score)
print(f"Average score over 1000 run : {np.array(scores).mean()}")
score()
# -
| HW8/Practical/ML2021S-HW8.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.10 64-bit (windows store)
# language: python
# name: python3
# ---
# Practicing Statistics and visualization with Pandas
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# Importing Pandas and reading from a .tsv file. There is no heading in the file so the names are to set headings. Dataframes are used to store an instance of the read file.
df = pd.read_csv("../../data/SOCR_MLB.tsv", sep = "\t", header=None, names = ['Name','Team','Role','Height','Weight','Age'])
df.head()
df.info() #This shows aggregate information on the data file
df.describe() #This is done to describe the data with mean, Q1, Q2, Q3 min and max values of the columns having integer and float features
# +
print("There are {} observations and {} features in this dataset. \n".format(df.shape[0],df.shape[1]))
print("There are {} number of baseball players in this dataset \n".format(len(df.Name)))
print("Here is the distribution of this dataset according to roles: \n{} \n".format(df["Role"].value_counts()))
# +
#We import numpy as np, Numpy is used to carry out statistical operations on matrices such as mean, std and variance
mean = np.mean(df['Height'])
print (mean)
# +
#mean and std can also be displayed using the functions as shown
Mean_height = df["Height"].mean()
# Also np.mean(df["Height"]) can be used
Std_height = df['Height'].std()
# -
print("the mean heights of players are", Mean_height)
#print(f"the mean heights of players are {df.Height.mean()}")
print("the standard deviation value of heights of players are", Std_height)
df[['Age','Height','Weight']].mean() #To display means of several data frames
print(list((df['Height'])[:20]))
# +
mean = df[['Height', 'Weight', 'Age']].mean()
std = df[['Height','Weight', 'Age']].std()
var = df[['Height','Weight', 'Age']].var()
print(f"Means {mean} ,\n Standard deviations {std}, \n and Variances {var}")
# +
#df.hist(column = "Height")
df.boxplot(column = "Height")
plt.xticks(rotation=None)
'''
plt.figure(figsize=(10, 2))
plt.boxplot(df['Height'], vert=False, showmeans=True)
plt.title("Boxplot showing heights")
plt.xlabel("Heights")
'''
#plt.hist(df['Height'])
plt.grid(color='gray',linestyle='dotted')
plt.show()
# -
#plt.figure(figsize=(20,10), dpi = 15)
df.boxplot(column='Height',by='Role')
plt.xticks(rotation='vertical')
plt.show()
# +
# to generate some random data for heights using the mean and standard deviation values we had
mean = df['Height'].mean()
std = df['Height'].std()
generated = np.random.normal(mean,std,1000)
generated[:20]
# -
plt.hist(generated, bins = 16)
plt.show()
# +
#This is to generate a random normal distribution with mean = 0, std = 1, having 300 bins
plt.hist(np.random.normal(0,1,50000),bins=300)
plt.show()
# -
#The random numbers can also be generated in a way that does not follow a normal distribution
wrong_sample = np.random.rand(1000)
#print (wrong_sample)
print(type(wrong_sample))
plt.hist(wrong_sample)
plt.show()
# To generate a description of the data with reference to a particular column groupings
df.groupby("Role").agg({"Height":"mean", "Weight":"mean", "Name":"count"}).rename(columns={"Name":"Count"})
# For confidence interval using scipy.stats
from scipy.stats import norm
D = df["Height"]
mean = df.Height.mean()
std = df.Height.std()
D = norm(mean, std)
for x in [.95, .90, .85]:
print(f'{D.interval(x)}, {x*100}% confidence level')
# to find the mean heights of catchers using the Role and the Height column data
# +
meanCatcher = (df.loc[df['Role'] == "Catcher",["Height"]]).mean()
print(meanCatcher)
#to generate the mean for first baseman and second baseman
meanFirstBase = (df.loc[df['Role'] == "First_Baseman",["Height"]]).mean()
stdFirstBase = (df.loc[df['Role'] == "First_Baseman",["Height"]]).std()
meanSecondBase = (df.loc[df['Role'] == "Second_Baseman",["Height"]]).mean()
stdSecondBase = (df.loc[df['Role'] == "Second_Baseman",["Height"]]).std()
print(f"Mean value for the first baseman is {meanFirstBase}")
print(f"Std value for the first baseman is {stdFirstBase}")
print(f"Mean value for the first baseman is {meanSecondBase}")
print(f"Std value for the first baseman is {stdSecondBase}")
# -
# Lets play around with confidence level generating inbuilt function in Scipy.stats
# using a normal distribution.
# this is not the confidence interval for our distribution but for a normal distribution using the mean and sandard deviation values from our distribution.
D = norm(meanFirstBase, stdFirstBase)
for x in [.95, .90, .85]:
print(f'{D.interval(x)}, {x*100}% confidence level')
D = norm(meanSecondBase, stdSecondBase)
print(type (D))
for x in [.95, .90, .85]:
print(f'{D.interval(x)}, {x*100}% confidence level')
help("modules _distn_infrastructure")
# To generate the confidence level for our distribution, and to prove our hypothesis that all first basemen are taller than second basemen. This hypothesis was drawn because the first baseman are averagely taller than the second baseman. To test this hypothesis, Lets generate our confidence interval.
# +
import numpy as np
import scipy.stats as st
# we create a variable FBdata to hold first baseman height data
FBdata = df.loc[df['Role'] == "First_Baseman",["Height"]]
#create 95% confidence interval for population mean height
#using inbuilt function st.t.interval, we pass in confidence value, n-1, mean, standard error of mean caclculated with inbuilt function st.sem
print(f"95% confidence interval for First baseman Heights is {st.t.interval(alpha=0.95, df=len(FBdata)-1, loc=np.mean(FBdata), scale=st.sem(FBdata))}")
#similarly for second baseman
SBdata = df.loc[df['Role'] == "Second_Baseman",["Height"]]
print(f"95% confidence interval for Second baseman Heights is {st.t.interval(alpha=0.95, df=len(SBdata)-1, loc=np.mean(SBdata), scale=st.sem(SBdata))}")
# -
# To calculate the confidence interval for the weight
# +
FBWdata = df.loc[df['Role'] == "First_Baseman",["Weight"]]
#create 95% confidence interval for population mean height
#using inbuilt function st.t.interval, we pass in confidence value, n-1, mean, standard error of mean caclculated with inbuilt function st.sem
print(f"95% confidence interval for First baseman Weights is {st.t.interval(alpha=0.95, df=len(FBWdata)-1, loc=np.mean(FBWdata), scale=st.sem(FBWdata))}")
#similarly for second baseman
SBWdata = df.loc[df['Role'] == "Second_Baseman",["Weight"]]
print(f"95% confidence interval for Second baseman Weights is {st.t.interval(alpha=0.95, df=len(SBWdata)-1, loc=np.mean(SBWdata), scale=st.sem(SBWdata))}")
# -
# To calculate the confidence interval of the entire weight mean.
#
# Note: the use of fillna method is to remove all the nan values in the list
# +
weight = list(df["Weight"].fillna(method='pad'))
print(f"95% confidence interval for the Weights column is {st.t.interval(alpha=0.95, df=len(weight)-1, loc=np.mean(weight), scale=st.sem(weight))}")
print(f"90% confidence interval for the Weights column is {st.t.interval(alpha=0.90, df=len(weight)-1, loc=np.mean(weight), scale=st.sem(weight))}")
print(f"85% confidence interval for the Weights column is {st.t.interval(alpha=0.85, df=len(weight)-1, loc=np.mean(weight), scale=st.sem(weight))}")
# -
| 1-Introduction/04-stats-and-probability/Practice_statistics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exercise 14 - Structured Data in Python ([`pandas`](https://pandas.pydata.org/))
#
# You should now know how to open files, read data tables from them using numpy, and write your outputs to new files. Getting data in/out of Python is a routine task for data analysis, and people have put a lot of effort into making it as simple as possible. [`pandas`](https://pandas.pydata.org/) is one of the main data science modules, which specialises in handling 'spreadsheet'-type data - i.e. a data table with column and/or row labels.
#
# In this practical you'll learn the basics of reading and writing data using `pandas`, some basic data manipulation and plotting.
#
# In addition to this practical, `pandas` has a [number of excellent tutorials](https://pandas.pydata.org/pandas-docs/stable/tutorials.html) focussed at beginners. There is also a chapter about `pandas` in the [Python Data Science Handbook](https://jakevdp.github.io/PythonDataScienceHandbook/03.00-introduction-to-pandas.html). If your work is based around spreadsheet-type data, we strongly recommend going through these tutorials in due course.
#
# ## `pandas` basics
#
# The central concept of `pandas` is the `DataFrame`. This is a 2D table of data, with associated row and column labels - just like a spreadsheet. A `DataFrame` can be 'indexed' by row or column names - i.e. you use the row/column *labels*, instead of their numeric indices like you do in `numpy` arrays.
#
# `pandas` started as a convenient way to import and export labeled data, but has grown into something a lot more comprehensive, and now does basic statistics, plotting, time-series analysis, 'split-apply-combine' data grouping... and much more.
#
# You import pandas just like any other module; conventionally, we use
# ```python
# import pandas as pd
# ```
# To create a `DataFrame`, you need some 2D data, which you give to `pandas`:
#
# ```python
# df = pd.DataFrame(data)
# ```
#
# **➤ Generate a numpy array of random data with 3 rows and 4 columns, and make a `DataFrame`.**
# + tags=[]
# Try it here!
# -
# When you look at the `DataFrame`, it should look like a nice table in the Jupyter notbook, with numbered rows and columns. These numbers are the same as the `numpy` indices that you could use to access these rows/column.
#
# You can access these data by these indices, just like you would in `numpy`. However, because `DataFrames` are quite a lot more complex than `numpy` arrays, you can't index them directly - you have to use `.loc[]`.
#
# **➤ Use the `.loc[]` method to index your `DataFrame`. Compare it to `numpy` indexing.**
# + tags=[]
# Try it here!
# -
# So far, everything we've done can be achieved with equivalent convenience in `numpy`... so what's the point? The strength of `pandas` is in its use of data labels, so you can associate rows/columns with meaningful information, rather than numerical indices.
#
# This can be really useful if, for example, you're working with a dataset that you add new columns to over time. If you were using numpy, the numeric indices of the columns of interest might change, and break your code. With `pandas`, your code will be fine as long as the labels of the columns stays the same.
#
# Let's assign some labels to your dataset. You can do this by setting the `columns` or `index` (rows) attributes of your dataframe with a list of strings whose length is equal to the number of columns or rows. For example,
#
# ```python
# df = pd.DataFrame(...)
# df.columns=['Label_1', 'Label_2',...,'Label_N']
# df.index = ['Row_1','Row_2',...,'Row_M']
# ```
#
# ### What should labels be?
#
# You can call columns/rows whatever you like. A column name could be a whole paragraph, if you really wanted... but this is a really bad idea. You should use labels that are **unique**, **meaningful** and **concise**.
#
# For example, `Temperature` isn't great - what are the units? What if you have Kelvin, Centigrade *and* Fahrenheit columns? `Temperature °C` is better, but will be cumbersome in routine use, because it's long, and has special characters and spaces in. `T_C` is ideal - it's nice and short, containing a commonly used abbreviation for temperature ('T'), and the unit.
#
# **Bonus:** If your label names start with a letter and don't contain spaces or special characters, you can access them directly using the `.` operator, without needing to use `.loc[]`. For example:
# ```python
# df.Label_1
# df.loc[:, 'Label_1']
# ```
# both produce return the same column, but `df.Label_1` brings the added benefit of having <kbd>Tab</kbd> auto-completion for your column names! This can be really convenient if, for example, you want to access columns during plotting.
#
# **➤ Label the columns of your dataframe as [A, B, C, D], and your rows [X, Y, Z]:**
# + tags=[]
# Try it here!
# -
# When you look at your dataframe again, you should see that it now displays the column/row names that you gave it.
#
# **➤ Access column 'B' of your data:**
#
# **Hint:** If you want to select a particular column, you'll have to use the `:` operator to specify 'all row values', just like in `numpy`.
# + tags=[]
# Try it here!
# -
# You may also notice here that numeric indices no longer work with the `.loc[]` operator. You *can* still use numeric indices if you *really* want to using `.iloc[]`... although if you're doing this all the time you'd be better off using `numpy` instead of pandas.
#
# Just like `numpy` arrays, `DataFrames` also contain a number of useful properties, such as `.size` and `.shape`, which tell you useful information about the data.
#
# **➤ Try it here:**
#
# + tags=[]
# Try it here!
# -
# ### Read & Write Data
#
# Pandas has a number of functions for reading and writing data in a [wide range of formats](https://pandas.pydata.org/pandas-docs/stable/io.html).
#
# Data reading functions are available within the pandas module in the form `pd.read_FORMAT()`, and return a `DataFrame`. Writing functions are available as a method directly accessible from the `DataFrame`, and are in the form `dataframe.to_FORMAT()`. In both these cases, replace `FORMAT` with the desired data format (e.g. 'excel' or 'csv'). Both of these functions take a file path and a number of other arguments that modify their behaviour.
#
# **➤ Load the file 'boston.csv' into a `DataFrame`, find out how large the dataset is, and look at the top 5 rows.**
#
# **Hint 1:** Take a look at the file in a text editor first, and then use pandas' `read_csv` function. You'll probably want to use the `comment` parameter to tell pandas that lines beginning with `#` are comments and should be ignored!
#
# **Hint 2:** Try the `.shape` attribute.
#
# **Hint 3:** Use the `.head()` method to see the top N lines (and `.tail()` to see the bottom N!)
# + tags=[]
# Try it here!
# -
# **➤ Save the Boston `DataFrame` as an html table**
# + tags=[]
# -
# Take a look at the resulting files, to make sure they look right. Have a play around with parameters (e.g. `index=False` to remove the row numbers).
#
# ### Microsoft Excel
# Is irritatingly common in science. Irritating, because it saves data in a proprietary binary format which can't be easily read without having a copy of Microsoft Excel (or sometimes, the right *version* of Microsoft Excel!). There are ways of importing Excel files into Python, but most are quite complex. `pandas` offers a very easy solution to this problem.
#
# **➤ Load the file `iris.xlsx` into a `DataFrame`, with `species` as the index (row) labels**
#
# **Hint 1:** the data are in a sheet called 'iris_data'.
#
# **Hint 2:** Everything is installed on the RSES Jupyter server, but you will need to install the package `xlrd` to read Excel spreedsheets with Pandas on your computer.
iris = pd.read_excel('iris.xlsx', sheet_name='iris_data', index_col=0)
iris.head()
# **➤ Save the iris `DataFrame` as a csv**
# **➤ Save the iris `DataFrame` as a $\LaTeX$ table**
#
# This can be *really* useful if you do your work in $\LaTeX$, as you might for a MSc or PhD thesis!
# ### Reading from the Web(!)
#
# `pandas` can also read data directly from the web, which is useful if you're using a central database which is regularly updated.
#
# For example, the famous [Mauna Loa Atmospheric CO2 Dataset](https://www.esrl.noaa.gov/gmd/ccgg/trends/full.html), which is updated monthly.
#
# The URL for the monthly dataset is: ftp://aftp.cmdl.noaa.gov/products/trends/co2/co2_mm_mlo.txt
#
# **➤ Look at this dataset in your browser, and read it into a `pandas.DataFrame` using the `pd.read_table` function.**
#
# **Hint 1:** you'll need to specify the delimiter between the data columns using the `delimiter` argument.
#
# **Hint 2:** because of data formatting choices, there's no simple way to get the column headings from this dataset, and you'll have to assign them manually.
# +
co2 = pd.read_table('ftp://aftp.cmdl.noaa.gov/products/trends/co2/co2_mm_mlo.txt', comment='#', delimiter='\s+', header=None)
co2.columns = ['year', 'month', 'decimal_year', 'co2_average', 'co2_interpolated', 'co2_trend', 'n_days']
# -
co2.head()
# ## Plotting & Data Cleaning
#
# Let's have a look at this CO2 record. Combine what you learned about indexing DataFrames and plotting to make a line graph of time (decimal year) vs. average CO2.
#
# **➤ Make a plot!** You can simply pass the dataframe columns to `plt.plot()` as if they were Numpy arrays.
# Try it here!
import matplotlib.pyplot as plt
# This looks mad! There are several points where average CO2 is less than zero... what's going on?
#
# You'll see from the header of the Mauna Load CO2 record that values that are equal the -99.99 represent 'missing data'. We therefore want to exclude this data from further analyses.
#
# **➤ Do this by using the `.replace()` method to substitute `np.nan` for `-99.99`, then re-draw the plot**
#
# **Hint:** you want to replace these values in the original dataset, rather than return a copy of the data with these values replaced. Think about what the `inplace` argument does in this function.
#
# Try it here!
# Much better! Let's keep a copy of this data for future offline use.
#
#
# **➤ Save the co2 data as a csv file.**
# Try it here!
# You'll notice this data has an overall trend, and a period oscillation around the trend. This oscillation is caused by seasonal changes in the balance between respiration and photosynthesis - CO2 drops in the summer, when plants in the Northern hemisphere are photosynthesising more.
#
# Let's try to isolate the overall trend by working out a yearly, instead of monthly trend.
#
# We can do this using the 'split-apply-combine' methodology. We 'split' the data according to the value of a particular field, 'apply' a function to each subset, and the 'combine' all the results back together.
#
# In `pandas` we do this using the `.groupby()` function to specify that all records with the same entry for a particular field should be treated as a group, followed by `aggregate` (or `apply`, for more complex operations) to specify how each group should be reduced into a single number.
#
# **➤ Calculate and plot the annual mean CO2 for the Mauna Loa record**
# Try it here!
# ## Subsets
#
# Often, you might want to create a subset of a dataset. For example, you might wish to isolate only the `year` and `co2_average` information from the yearly mean.
#
# You can do this by indexing the dataset, and assigning it to a new name.
#
# **➤ Create a subset of the annual mean data containing only the `co2_average` column**
# Try it here!
# ## reset_index()
#
# In this subset you'll notice that you only have one column, but the 'year' information is preserved in the `index` of the data. You can turn the index back into a data column using `.reset_index()`.
#
# **➤ Try it here!**
# ## Merge Dataframes
#
# Let's try to look at the seasonal oscillation in more detail. To do this, we need to subtract the annual mean. However, at the moment our annual mean and our seasonal data are in two different `DataFrames` of different sizes.
#
# `pandas` has a number of functions for combining and merging `DataFrames` in different ways. Let's have a go at using `pd.merge()` to combine the annual mean dataset with the original data.
#
# **➤ Try it here!** You will need to look at the help for `pd.merge()` to work out how to do this.
# ## Operations and New Columns
#
# Now we've combined them, we need to subtract the annual mean from the monthly signal to isolate the seasonal trend.
#
# **➤ Create a new column containing the seasonal signal**
#
# **Hint:** If you try to assign something to an index that doesn't exist, `pandas` creates a new column.
# ## Selecting Data
#
# Remember indexing from above? You can use a similar approach in `pandas` to select data in a number of ways. For example, we can use 'logical indexing' to select data only from the last 10 years.
#
# To do this, we use logical operators (`>`, `<`, `==`) to create a `boolean` array the same size as a dimension of our array. For example:
#
# ```python
# ind = co2.loc[:, 'year'] >= 2008
# ```
#
# In this example, `ind` will be `True` wherever year is greater than or equal to 2008.
#
# We can then apply this to select a subset of our `DataFrame`:
#
# ```python
# last_ten = co2.loc[ind, :]
# ```
#
# **➤ Create a subset containing the years between 1980-1990, and plot the seasonal trend.**
#
| jupyterbook/content-de/python/lab/ex14-pandas_databases.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.6.8 64-bit (''benchmark'': conda)'
# name: python368jvsc74a57bd060aa7568c9db8113868ebef0220b161e96389b06f6ba9eb98d46b6b0f2cf6a72
# ---
import sys
sys.path.append("../") # go to parent dir
import pandas as pd
import sys
import spacy
import re
import time
import scispacy
import glob
import os
from tqdm import tqdm
tqdm.pandas()
from note_processing.heuristic_tokenize import sent_tokenize_rules
# +
# OUTPUT_DIR = '/mnt/data01/mimic-3/benchmark-small/test/345' #this path will contain tokenized notes. This dir will be the input dir for create_pretrain_data.sh
#this is the path to mimic data if you're reading from a csv. Else uncomment the code to read from database below
MIMIC_NOTES_PATHS = ['/mnt/data01/mimic-3/benchmark-small/test',
'/mnt/data01/mimic-3/benchmark-small/train']
WORKERS = 5
# +
all_files = []
for path in MIMIC_NOTES_PATHS:
files = glob.glob(path + "/*/*_notes.csv")
all_files += files
print("\nTotal note files: " + str(len(all_files)))
all_files = [f for f in all_files if not os.path.exists(f[:-4] + '_sent.csv')]
print("Total unprocessed files: " + str(len(all_files)))
li = []
for filename in tqdm(all_files, desc="Load note files"):
df = pd.read_csv(filename, index_col=None, header=0)
df["filename"] = filename
li.append(df)
notes = pd.concat(li, axis=0, ignore_index=True)
notes.describe(include="all")
# -
notes.head(5)
notes.groupby("CATEGORY").agg(['count'])
# +
# aflanders:
# This code will split the notes into natural sentence boundaries separated by \n
# which can then be fed into sentence embedding models such as BIO-ClinicalBert or
# BioSentVec
#
# This frame and the next are largly from format_mimic_for_BERT.py in EmilyAlsentzer/clinicalBERT
# I have updated the code to work with spacy 3.0 and made some other changes
#
# Example:
# THis is a
# single
# sentence. and another sentence.
# THis is a single sentence.\n
# and another sentence.\n
from spacy.language import Language
#setting sentence boundaries
@Language.component('sbd_component')
def sbd_component(doc):
for i, token in enumerate(doc[:-2]):
# define sentence start if period + titlecase token
if token.text == '.' and doc[i+1].is_title:
doc[i+1].sent_start = True
if token.text == '-' and doc[i+1].text != '-':
doc[i+1].sent_start = True
return doc
#convert de-identification text into one token
# aflanders: no need to pass in the next separate, is available in processed_text
# def fix_deid_tokens(text, processed_text):
def fix_deid_tokens(doc):
deid_regex = r"\[\*\*.{0,15}.*?\*\*\]"
indexes = [m.span() for m in re.finditer(deid_regex, doc.text, flags=re.IGNORECASE)]
for start,end in indexes:
# processed_text.merge(start_idx=start,end_idx=end)
# aflanders: Make compatible with latest version fo spacy
try:
span = doc.char_span(start, end)
if span is not None:
with doc.retokenize() as retokenizer:
# retokenizer.merge(processed_text[start:end+1])
retokenizer.merge(span)
except:
print(f'Error with: {text}')
return doc
# +
def process_section(section, note, processed_sections):
# perform spacy processing on section
processed_section = nlp(section['sections'])
# processed_section = fix_deid_tokens(section['sections'], processed_section)
processed_section = fix_deid_tokens(processed_section)
processed_sections.append(processed_section)
def process_note_helper(note):
# split note into sections
note_sections = sent_tokenize_rules(note)
processed_sections = []
section_frame = pd.DataFrame({'sections':note_sections})
section_frame.apply(process_section, args=(note,processed_sections,), axis=1)
return(processed_sections)
def process_text(sent, note):
sent_text = sent['sents'].text
if len(sent_text) > 0 and sent_text.strip() != '\n' and len(sent_text.split()) > 1:
if '\n' in sent_text:
sent_text = sent_text.replace('\n', ' ')
note['TEXT'] += sent_text + '\n'
def get_sentences(processed_section, note):
# get sentences from spacy processing
sent_frame = pd.DataFrame({'sents': list(processed_section['sections'].sents)})
sent_frame.apply(process_text, args=(note,), axis=1)
def process_note(note):
try:
note_text = note['TEXT'] #unicode(note['text'])
note['TEXT'] = ''
processed_sections = process_note_helper(note_text)
ps = {'sections': processed_sections}
ps = pd.DataFrame(ps)
ps.apply(get_sentences, args=(note,), axis=1)
return note
except Exception as e:
# pass
print ('error processing note', e)
# + tags=[]
# # %time
#category = ["Nursing", "Nursing/other", 'General', 'Physician '] # or None
category = ["Nursing/other"] # or None
# start = time.time()
# tqdm.pandas()
print('Begin reading notes')
if category != None:
notes = notes[notes['CATEGORY'].isin(category)]
print('Number of notes: %d' %len(notes.index))
# notes['ind'] = list(range(len(notes.index)))
nlp = spacy.load('en_core_sci_md', disable=['tagger','ner', 'lemmatizer'])
nlp.add_pipe('sbd_component', before='parser')
# -
filenames = list(notes["filename"].unique().tolist())
len(filenames)
from pandarallel import pandarallel
pandarallel.initialize(progress_bar=True, nb_workers=WORKERS)
formatted_notes = notes.parallel_apply(process_note, axis=1)
formatted_notes.head(3)
# Write out a new note files organized by sentence
filenames = list(formatted_notes["filename"].unique().tolist())
for filename in tqdm(filenames, desc="Writing note sentence files"):
df = formatted_notes[formatted_notes["filename"] == filename][["Hours", "CATEGORY", "DESCRIPTION", "TEXT"]]
df = df.set_index("Hours")
write_file = filename.replace(".csv", "_sent.csv")
with open(write_file, "w") as f:
df.to_csv(f, index_label='Hours')
| notebooks/notes-exploration-04.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
# +
import warnings
warnings.filterwarnings("ignore")
from datetime import datetime
import math
import matplotlib as mpl
import matplotlib.pyplot as plt
import numpy as np
import operator
import pandas as pd
import seaborn as sns
mpl.rcParams['figure.figsize'] = [8, 5]
# -
# ## NPHET IEMAG 2021-03-29 Model Scenario Projections vs Reported Data
#
# This notebook compares the NPHET IEMAG 2021-03-29 model scenario projections with reported data, for 2021-04-05 to 2021-09-30. The projections are the mean values from Appendix 1 (Additional Modelling Charts) in the [2021-03-29 NPHET letter to the Minister for Health](https://assets.gov.ie/129551/f6833e49-b133-4e06-9194-848726762bdd.pdf).
#
# Note:
# * The focus is on cases only, as daily data are currently not provided for hospital admissions, ICU admissions, and deaths. These will be included when they are available again.
# ### Load the reported case data
#
# This is a daily snapshot of the [Covid-19 Daily Statistics](https://covid-19.geohive.ie/datasets/d8eb52d56273413b84b0187a4e9117be_0/explore?location=53.385818%2C-8.057012%2C7.64&showTable=true) on [Ireland's COVID-19 Data Hub](https://covid-19.geohive.ie/)
casesdf = pd.read_csv('./data/CovidStatisticsProfileHPSCIrelandOpenData.csv', parse_dates=["Date"], date_parser=lambda x: datetime.strptime(x, "%Y/%m/%d %H:%M:%S+00"))
casesdf = casesdf[casesdf.Date>='2021-04-05'][['Date', 'ConfirmedCovidCases']]
casesdf.rename(columns={'ConfirmedCovidCases': 'Cases'}, inplace=True)
casesdf.Cases = casesdf.Cases.cumsum()
casesdf['Scenario'] = 'Reported'
# ### Scenario data 2021-04-05 to 2021-09-30, based on Appendix 1 from [2021-03-29 NPHET letter](https://assets.gov.ie/129551/f6833e49-b133-4e06-9194-848726762bdd.pdf)
scenario_dates = pd.date_range('2021-04-05', '2021-09-30')
scenario_mean_cases = {
'A': 80000,
'B2': 96000,
'B1': 152000,
'C2': 177000,
'B': 199000,
'C1': 291000,
'C': 578000
}
scenariodf = pd.DataFrame({'Date': np.repeat(scenario_dates, len(scenario_mean_cases)),
'Scenario': [*scenario_mean_cases.keys()] * len(scenario_dates),
'Cases': [*scenario_mean_cases.values()] * len(scenario_dates)})
# ### NPHET scenarios vs reported cases
generate_dashes = lambda num_scenarios:[(2,2)] * num_scenarios + ['']
all_scenario_dashes = generate_dashes(len(scenario_mean_cases))
optimistic_scenario_dashes = generate_dashes(3)
# Consistent palette for 'A', 'B2', 'B1,' 'Reported'
optimistic_scenario_palette = operator.itemgetter(*[0,1,2,7])(sns.color_palette())
ax = sns.lineplot(data=pd.concat((scenariodf, casesdf), ignore_index=True), x='Date', y='Cases', hue='Scenario', style='Scenario', dashes=all_scenario_dashes)
ax.set_title('NPHET IEMAG 2021-03-29 Model Scenarios vs Reported Cases (Cumulative)')
plt.legend(loc='upper right')
plt.xticks(rotation=30);
plt.savefig('./figures/nphet20210329_all_scenarios_vs_reported_cases.png', bbox_inches="tight")
ax = sns.lineplot(data=pd.concat((scenariodf[scenariodf.Scenario.isin(['A', 'B2', 'B1'])], casesdf), ignore_index=True), x='Date', y='Cases', hue='Scenario', style='Scenario', dashes=optimistic_scenario_dashes, palette=optimistic_scenario_palette)
ax.set_title('NPHET IEMAG 2021-03-29 Optimistic Scenarios vs Reported Cases (Cumulative)')
plt.legend(loc='upper right')
plt.savefig('./figures/nphet20210329_optimistic_scenarios_vs_reported_cases.png', bbox_inches="tight")
# ### Generate current cases summary
cases_current = casesdf.iloc[-1]
cases_previous = casesdf.iloc[-2]
new_cases = cases_current.Cases - cases_previous.Cases
summary = f"""NPHET IEMAG 2021-03-29 Model Scenarios vs Reported Cases
({datetime.strftime(cases_current.Date, "%A %Y-%m-%d")})
- Reported Total: {cases_current.Cases} (+{new_cases})
- % Scenario Duration ({len(scenario_dates)} days): {len(casesdf)*100/len(scenario_dates):.1f}%
- % A Scenario Total: {cases_current.Cases*100/scenario_mean_cases['A']:.1f}% (+{new_cases*100/scenario_mean_cases['A']:.1f}%)
- % B2 Scenario Total: {cases_current.Cases*100/scenario_mean_cases['B2']:.1f}% (+{new_cases*100/scenario_mean_cases['B2']:.1f}%)
- % B1 Scenario Total: {cases_current.Cases*100/scenario_mean_cases['B1']:.1f}% (+{new_cases*100/scenario_mean_cases['B1']:.1f}%)
"""
print(summary)
| notebooks/NPHET IEMAG 2021-03-29 Model Scenario Projections vs Reported Data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/myamullaciencia/conda/blob/master/fes_en_python_1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="M91JoCRI1trY" colab_type="code" colab={}
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
import datatable as dt
# + [markdown] id="o1Hktrh54EOR" colab_type="text"
# ### Installing necessary packages
# + id="l4jAuGnk3K9s" colab_type="code" colab={}
# #!pip install https://s3.amazonaws.com/h2o-release/datatable/stable/datatable-0.8.0/datatable-0.8.0-cp36-cp36m-linux_x86_64.whl
#pip install feature-engine
# + id="WbbacP6N3rhC" colab_type="code" colab={}
# Loading data
sflow_data = pd.read_csv('https://assets.datacamp.com/production/repositories/3752/datasets/19699a2441073ad6459bf5e3e17690e2cae86cf1/Combined_DS_v10.csv')
# + id="ZpV9kc3L4aXG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 310} outputId="90c73eaf-3044-4d1a-dccc-c1a04c633e3d"
sflow_data.head()
| fes_en_python_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="ivpj9EeVLWTL"
# ##### Copyright 2019 Google LLC.
# + cellView="form" colab={} colab_type="code" id="jXIU2YPcN75z"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="UUiCfWD_i6Ck"
# #Spherical Harmonics Optimization
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/graphics/blob/master/tensorflow_graphics/notebooks/spherical_harmonics_optimization.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/graphics/blob/master/tensorflow_graphics/notebooks/spherical_harmonics_optimization.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# </table>
# + [markdown] colab_type="text" id="iFqF3lDo3fIa"
# This Colab covers an advanced topic and hence focuses on providing a toy example
# to form a high level understanding of how to estimate environment lighting using
# Spherical Harmonics rather than providing step by step details. We refer the
# interested reader to
# [another Colab](https://colab.research.google.com/github/tensorflow/graphics/blob/master/tensorflow_graphics/notebooks/spherical_harmonics_approximation.ipynb)
# to get a high level understanding of Spherical Harmonics.
#
# Given an image of a known object (sphere) with a known reflectance function,
# this Colab illustrates how to perform optimization of spherical harmonics to
# recover the lighting environment.
# + [markdown] colab_type="text" id="hM73at_pLadB"
# ## Setup & Imports
# If Tensorflow Graphics is not installed on your system, the following cell can install the Tensorflow Graphics package for you.
# + colab={} colab_type="code" id="G3WpJC9GLfUe"
# !pip install tensorflow_graphics
# + [markdown] colab_type="text" id="O7Fe-GqGLif6"
# Now that Tensorflow Graphics is installed, let's import everything needed to run the demo contained in this notebook.
# + colab={} colab_type="code" id="cV4dat2Ayjfu"
import matplotlib.pyplot as plt
import numpy as np
import tensorflow as tf
from tensorflow_graphics.rendering.camera import orthographic
from tensorflow_graphics.geometry.representation import grid
from tensorflow_graphics.geometry.representation import ray
from tensorflow_graphics.geometry.representation import vector
from tensorflow_graphics.math import spherical_harmonics
from tensorflow_graphics.math import math_helpers as tf_math
tf.compat.v1.enable_v2_behavior()
# + colab={} colab_type="code" id="Efb6MPaEQQ-m"
def compute_intersection_normal_sphere(width, height, radius,
center, dtype):
"""Estimates sphere normal and depth for each pixel in the image."""
# Generates a 2d grid storing pixel coordinates.
pixel_grid_start = np.array((0.5, 0.5), dtype=dtype)
pixel_grid_end = np.array((width - 0.5, height - 0.5), dtype=dtype)
pixel_nb = np.array((width, height))
pixels = grid.generate(pixel_grid_start, pixel_grid_end, pixel_nb)
# Computes the ray direction of each pixel.
pixel_ray = tf.math.l2_normalize(orthographic.ray(pixels), axis=-1)
# Defines the position of pixels in world coordinates.
zero_depth = np.zeros([width, height, 1])
pixels_3d = orthographic.unproject(pixels, zero_depth)
# Computes intersections with the sphere and surface normals for each ray.
intersection, normal = ray.intersection_ray_sphere(
center, radius, pixel_ray, pixels_3d)
# Extracts data about the closest intersection.
intersection = intersection[0, ...]
normal = normal[0, ...]
# Replaces NaNs with zeros.
zeros = tf.zeros_like(pixels_3d)
intersection = tf.where(tf.math.is_nan(intersection), zeros, intersection)
normal = tf.where(tf.math.is_nan(normal), zeros, normal)
return intersection, normal
# + [markdown] colab_type="text" id="Mhj6ZBssOvYT"
# ## Spherical Harmonics optimization
# + colab={} colab_type="code" id="Uvcgy69dzsS1"
light_image_width = 100
light_image_height = 100
dtype = np.float64
############################################################################
# Builds the pixels grid and computes corresponding spherical coordinates. #
############################################################################
pixel_grid_start = np.array((0, 0), dtype=dtype)
pixel_grid_end = np.array((light_image_width - 1, light_image_height - 1),
dtype=dtype)
pixel_nb = np.array((light_image_width, light_image_height))
pixels = grid.generate(pixel_grid_start, pixel_grid_end, pixel_nb)
normalized_pixels = pixels / (light_image_width - 1, light_image_height - 1)
spherical_coordinates = tf_math.square_to_spherical_coordinates(
normalized_pixels)
theta = spherical_coordinates[:, :, 1]
phi = spherical_coordinates[:, :, 2]
################################################################################################
# Builds the Spherical Harmonics and sets coefficients for the light and reflectance functions. #
################################################################################################
max_band = 2
l, m = spherical_harmonics.generate_l_m_permutations(max_band)
l_broadcasted = tf.broadcast_to(l, [light_image_width, light_image_height] +
l.shape.as_list())
m_broadcasted = tf.broadcast_to(m, [light_image_width, light_image_height] +
l.shape.as_list())
theta = tf.expand_dims(theta, axis=-1)
theta_broadcasted = tf.broadcast_to(theta,
[light_image_width, light_image_height, 1])
phi = tf.expand_dims(phi, axis=-1)
phi_broadcasted = tf.broadcast_to(phi,
[light_image_width, light_image_height, 1])
sh_coefficients = spherical_harmonics.evaluate_spherical_harmonics(
l_broadcasted, m_broadcasted, theta_broadcasted, phi_broadcasted)
# The lighting and BRDF coefficients come from the first Colab demo on Spherical
# Harmonics.
light_coeffs = np.array((2.17136424e-01, -2.06274278e-01, 3.10378283e-17,
2.76236879e-01, -3.08694040e-01, -4.69862940e-17,
-1.85866463e-01, 7.05744675e-17, 9.14290771e-02))
brdf_coeffs = np.array((0.28494423, 0.33231551, 0.16889377))
# Reconstruction of the light function.
reconstructed_light_function = tf.squeeze(
vector.dot(sh_coefficients, light_coeffs))
###################################
# Setup the image, and the sphere #
###################################
# Image dimensions
image_width = 100
image_height = 80
# Sphere center and radius
sphere_radius = np.array((30,), dtype=dtype)
sphere_center = np.array((image_width / 2.0, image_height / 2.0, 100.0),
dtype=dtype)
# Builds the pixels grid and compute corresponding spherical coordinates.
pixel_grid_start = np.array((0, 0), dtype=dtype)
pixel_grid_end = np.array((image_width - 1, image_height - 1), dtype=dtype)
pixel_nb = np.array((image_width, image_height))
pixels = grid.generate(pixel_grid_start, pixel_grid_end, pixel_nb)
normalized_pixels = pixels / (image_width - 1, image_height - 1)
spherical_coordinates = tf_math.square_to_spherical_coordinates(
normalized_pixels)
################################################################################################
# For each pixel in the image, estimate the corresponding surface point and associated normal. #
################################################################################################
intersection_3d, surface_normal = compute_intersection_normal_sphere(
image_width, image_height, sphere_radius, sphere_center, dtype)
surface_normals_spherical_coordinates = tf_math.cartesian_to_spherical_coordinates(
surface_normal)
##########################################
# Estimates result using SH convolution. #
##########################################
target = spherical_harmonics.integration_product(
light_coeffs,
spherical_harmonics.rotate_zonal_harmonics(
brdf_coeffs,
tf.expand_dims(surface_normals_spherical_coordinates[:, :, 1], axis=-1),
tf.expand_dims(surface_normals_spherical_coordinates[:, :, 2],
axis=-1)),
keepdims=False)
# Sets pixels not belonging to the sphere to 0.
target = tf.where(
tf.greater(intersection_3d[:, :, 2], 0.0), target, tf.zeros_like(target))
#########################################################################################
# Optimization of the lighting coefficients by minimization of the reconstruction error #
#########################################################################################
# Initial solution.
recovered_light_coeffs = tf.Variable(
np.array((1.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0)))
def reconstruct_image(recovered_light_coeffs):
reconstructed_image = spherical_harmonics.integration_product(
recovered_light_coeffs,
spherical_harmonics.rotate_zonal_harmonics(
brdf_coeffs,
tf.expand_dims(
surface_normals_spherical_coordinates[:, :, 1], axis=-1),
tf.expand_dims(
surface_normals_spherical_coordinates[:, :, 2], axis=-1)),
keepdims=False)
return tf.where(
tf.greater(intersection_3d[:, :, 2], 0.0), reconstructed_image,
tf.zeros_like(target))
# Sets the optimization problem up.
def my_loss(recovered_light_coeffs):
reconstructed_image = reconstruct_image(recovered_light_coeffs)
return tf.nn.l2_loss(reconstructed_image - target) / (
image_width * image_height)
learning_rate = 0.1
with tf.name_scope("optimization"):
optimizer = tf.compat.v1.train.AdamOptimizer(learning_rate)
def gradient_loss(recovered_light_coeffs):
with tf.GradientTape() as tape:
loss_value = my_loss(recovered_light_coeffs)
return tape.gradient(loss_value, [recovered_light_coeffs])
####################
# Initial solution #
####################
target_transpose = np.transpose(target, (1, 0))
reconstructed_image = reconstruct_image(recovered_light_coeffs)
reconstructed_image = np.transpose(reconstructed_image, (1, 0))
plt.figure(figsize=(10, 20))
ax = plt.subplot("131")
ax.axes.get_xaxis().set_visible(False)
ax.axes.get_yaxis().set_visible(False)
ax.grid(False)
ax.set_title("Target")
_ = ax.imshow(target_transpose, vmin=0.0)
ax = plt.subplot("132")
ax.axes.get_xaxis().set_visible(False)
ax.axes.get_yaxis().set_visible(False)
ax.grid(False)
ax.set_title("Initial solution")
_ = ax.imshow(reconstructed_image, vmin=0.0)
################
# Optimization #
################
nb_iterations = 100
for it in range(nb_iterations):
gradients_loss = gradient_loss(recovered_light_coeffs)
optimizer.apply_gradients(zip(gradients_loss, (recovered_light_coeffs,)))
if it % 33 == 0:
reconstructed_image = reconstruct_image(recovered_light_coeffs)
reconstructed_image = np.transpose(reconstructed_image, (1, 0))
# Displays the target and prediction.
plt.figure(figsize=(10, 20))
ax = plt.subplot("131")
ax.axes.get_xaxis().set_visible(False)
ax.axes.get_yaxis().set_visible(False)
ax.grid(False)
ax.set_title("Target")
img = ax.imshow(target_transpose, vmin=0.0)
ax = plt.subplot("132")
ax.axes.get_xaxis().set_visible(False)
ax.axes.get_yaxis().set_visible(False)
ax.grid(False)
ax.set_title("Prediction iteration " + str(it))
img = ax.imshow(reconstructed_image, vmin=0.0)
# Shows the difference between groundtruth and prediction.
vmax = np.maximum(np.amax(reconstructed_image), np.amax(target_transpose))
ax = plt.subplot("133")
ax.axes.get_xaxis().set_visible(False)
ax.axes.get_yaxis().set_visible(False)
ax.grid(False)
ax.set_title("Difference iteration " + str(it))
img = ax.imshow(
np.abs(reconstructed_image - target_transpose), vmin=0.0, vmax=vmax)
# Reconstructs the groundtruth and predicted environment maps.
reconstructed_predicted_light = tf.squeeze(
vector.dot(sh_coefficients, recovered_light_coeffs))
# Displays the groundtruth and predicted environment maps.
plt.figure(figsize=(10, 20))
ax = plt.subplot("121")
ax.axes.get_xaxis().set_visible(False)
ax.axes.get_yaxis().set_visible(False)
ax.grid(False)
ax.set_title("Target light")
img = ax.imshow(reconstructed_light_function, vmin=0.0)
ax = plt.subplot("122")
ax.axes.get_xaxis().set_visible(False)
ax.axes.get_yaxis().set_visible(False)
ax.grid(False)
ax.set_title("Predicted light")
img = ax.imshow(reconstructed_predicted_light, vmin=0.0)
| tensorflow_graphics/notebooks/spherical_harmonics_optimization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # 2017 Census Income Per Capita presentation
# ## by <NAME>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Investigation Overview
#
# My goal with this analysis is to uncover the features that have large affects on per capita income.
#
# ## Dataset Overview
#
# This dataset contains population, travel, work, and income information from the 2017 Census Bureau ACS survey. It contains state, census tract, and county statistics. It also contains poverty, income, transportation, job type, and unemployment statistics. There are 74,001 rows, one for each census tract.
# + slideshow={"slide_type": "skip"}
# import all packages and set plots to be embedded inline
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sb
# %matplotlib inline
# suppress warnings from final output
import warnings
warnings.simplefilter("ignore")
# + slideshow={"slide_type": "skip"}
# load in the dataset into a pandas dataframe and create copy for analysis.
df_original = pd.read_csv('acs2017_census_tract_data_master.csv')
df = df_original.copy()
# + slideshow={"slide_type": "skip"}
# Function to generate a random sample of df rows.
def df_sample(dataframe, samp_size):
samples = np.random.choice(dataframe.shape[0], samp_size, replace = False)
return dataframe.loc[samples,:]
# Create sample dataframe with 1000 rows
df_samp = df_sample(df, 1000)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Relationship Between Per Capita Income and Unemployment Level
#
# At first, it might seem surprising that there is not a stronger coorelation between unemployment numbers and per capita income. However, it could be that there are simply not that many tracts with high enough levels of unemployment to bring down the averages.
# + slideshow={"slide_type": "subslide"}
# Plot per capita income by unemployment level.
plt.scatter(data = df_samp, x = 'Unemployment', y = 'IncomePerCap')
plt.xlabel('Unemployment Level')
plt.ylabel('Income Per Capita')
plt.title('Per Capita Income by Unemployment Level')
plt.xlim(left = 0, right = 30)
plt.ylim(bottom = 0, top = 100000);
# + [markdown] slideshow={"slide_type": "slide"}
# ## Relationhip Between Per Capita Income, Poverty, and Professional Job Levels.
#
# Here, it is clear that per capita income is higher in places that have more professional jobs and lower levels of poverty. Even though the overall trend is clear, there appear to be plenty of places with lower poverty levels that have lower levels of incomes and professional jobs. In addition, places with high poverty seem to stagnate in the income department even with the increase in professional jobs.
# + slideshow={"slide_type": "subslide"}
# Plot per capita income broken down by poverty and professional job levels.
plt.scatter(data = df_samp, x = 'Professional', y = 'IncomePerCap', c = 'Poverty')
plt.xlabel('Professional Job Level')
plt.ylabel('Income Per Capita')
cbar = plt.colorbar()
cbar.set_label('Poverty Level')
plt.title('Per Capita Income by Professional Job and Poverty Levels')
plt.ylim(bottom = 0, top = 100000)
plt.xlim(left = 0, right = 80);
# + [markdown] slideshow={"slide_type": "slide"}
# ## Relationhip Between Per Capita Income, Poverty, and Production Job Levels.
#
# Per capita income seems to benefit when there are less production jobs. As seen with professional jobs, places with the highest poverty levels are not affected by the level of production jobs.
# + slideshow={"slide_type": "subslide"}
# Plot per capita income broken down by poverty and production job levels.
plt.scatter(data = df_samp, x = 'Production', y = 'IncomePerCap', c = 'Poverty')
plt.xlabel('Production Job Level')
plt.ylabel('Income Per Capita')
cbar = plt.colorbar()
cbar.set_label('Poverty Level')
plt.title('Per Capita Income by Production Job and Poverty Levels')
plt.ylim(bottom = 0, top = 120000)
plt.xlim(left = 0, right = 40);
# + [markdown] slideshow={"slide_type": "slide"}
# ## Relationhip Between Per Capita Income, Poverty, and Service Job Levels.
#
# Per capita income drops rapidly with just a fairly small increase in service jobs, then the decrease is much more mild after that. A pattern appears to be emerging that shows that per capita income in places with high poverty does not change no matter the job composition.
# + slideshow={"slide_type": "subslide"}
# Plot per capita income broken down by poverty and service job levels.
plt.scatter(data = df_samp, x = 'Service', y = 'IncomePerCap', c = 'Poverty')
plt.xlabel('Service Job Level')
plt.ylabel('Income Per Capita')
cbar = plt.colorbar()
cbar.set_label('Poverty Level')
plt.title('Per Capita Income by Service Job and Poverty Levels')
plt.ylim(bottom = 0, top = 120000)
plt.xlim(left = 0, right = 60);
| census_cap_income_presentation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Import necessary packages
import os
import glob
import pandas as pd
import numpy as np
from scipy import stats
import scikit_posthocs
import iqplot
import bokeh.io
bokeh.io.output_notebook()
# # Import data from directory of measurement tables, collected from Fiji
# +
# Define path to directory with measurements
path = os.path.abspath('raw_data_csvs/')
df_summary = pd.DataFrame()
list_summary = []
# For loop to bring in files and concatenate them into a single dataframe
for file_ in glob.glob(path + "/*Summary.csv"):
df = pd.read_csv(file_)
# Determine Image name from file name, then parse experiment details from Image name
df['Image'] = os.path.splitext(os.path.basename(file_))[0]
# (df['Date'], df['CellLine'], df['Dose'], df['Time'], df['Treatment'],
# df['Channels'], df['Mag'], df['FOV'], df['Measurement']) = zip(*df['Image'].map(lambda x:x.split('_')))
(df['Date'], df['CellLine'], df['Treatment'], df['Rep'], df['Channels'],
df['FOV'], df['Measurement']) = zip(*df['Image'].map(lambda x:x.split('_')))
# (df['Date'], df['Treatment'], df['FOV'], df['Measurement']) = zip(*df['Image'].map(lambda x:x.split('_')))
# Drop unnecessary columns for tidiness
df = df.drop(['Total Area', 'Average Size', '%Area', 'Mean', 'IntDen', 'Image', 'Channels'], axis = 1)
# Compile data
list_summary.append(df)
df_summary = pd.concat(list_summary, sort=False)
# Preview dataframe to confirm import successful
df_summary.head()
# -
# # Parse dataframe by desired 'cell line' and 'treatment' combinations, then plot results
# +
# Generate and display list of cell lines and treatments present in this dataset
cell_list = df_summary['CellLine'].unique().tolist()
treatment_list = df_summary['Treatment'].unique().tolist()
print('Cells lines: ' + str(cell_list))
print('Treatments: ' + str(treatment_list))
# +
# Prepare for parsing data:
# To populate this "comment on/off" code block, copy the results of th cell lists above
cells = [
'U2OS',
]
treatments = [
'2aRFP',
'Dyn1K44A',
'SMPD3',
'SMPD3N130A',
]
# Copy dataset to not disrupt raw data
df_subset = df_summary
# Pull out only cells and treaments of interest
df_subset = df_subset.loc[df_subset['CellLine'].isin(cells)]
df_subset = df_subset.loc[df_subset['Treatment'].isin(treatments)]
# df_subset = df_subset['Count'].dropna()
# Make ECDF plot using iqplot
data_ecdf = iqplot.ecdf(
data=df_subset, q='Count', cats='Treatment'
# ,title=str(cells) + ' cells treated with ' + str(treatments)
,style='staircase'
,conf_int=True, n_bs_reps=1000, ptiles=[16, 84] # ptiles values equate to 68% CIs (SEM)
,line_kwargs=dict(line_width=3)
,show_legend=True
# Other customization parameters
,frame_height = 350, frame_width = 450
,order = treatments
,palette = ['#1f77b4', '#d62728', '#2ca02c', '#ff7f0e']
,x_axis_label='Transferrin-633 Puncta Count', y_axis_label='Cumulative Distribution Frequency'
,x_range=(0,50)
)
# Other customization parameters
data_ecdf.axis.axis_label_text_font_size = '20px'
data_ecdf.axis.axis_label_text_font_style = 'normal'
data_ecdf.axis.major_label_text_font_size = '18px'
# data_ecdf.output_backend = "svg"
bokeh.io.show(data_ecdf)
# +
# Prepare for parsing data:
# To populate this "comment on/off" code block, copy the results of th cell lists above
cells = [
'U2OS',
]
treatments = [
'2aRFP',
'Dyn1K44A',
'SMPD3',
'SMPD3N130A',
]
# Copy dataset to not disrupt raw data
df_subset = df_summary
# Pull out only cells and treaments of interest
df_subset = df_subset.loc[df_subset['CellLine'].isin(cells)]
df_subset = df_subset.loc[df_subset['Treatment'].isin(treatments)]
# df_subset = df_subset['Count'].dropna()
# Make stripbox plot using iqplot
data_stripbox = iqplot.stripbox(
data=df_subset, q='Count', cats='Treatment', q_axis='y'
# Plot details
,jitter=True, jitter_kwargs=dict(width=0.4)
,marker_kwargs=dict(alpha=0.8, size=6
,color='gray'
)
,box_kwargs=dict(line_color='black', line_width=1.5)
,whisker_kwargs=dict(line_color='black', line_width=1.5)
,median_kwargs=dict(line_color='black', line_width=2)
,top_level='box'
,frame_width=250, frame_height=350
# Other customization parameters
,order = treatments
,palette = ['#1f77b4', '#d62728', '#2ca02c', '#ff7f0e']
,x_axis_label='Treatment'
,y_axis_label='Transferrin-633 Puncta Count'
,x_range=(0,50)
)
# Other customization parameters
data_stripbox.axis.axis_label_text_font_size = '20px'
data_stripbox.axis.axis_label_text_font_style = 'normal'
data_stripbox.axis.major_label_text_font_size = '18px'
data_stripbox.xaxis.major_label_orientation = 7
# data_stripbox.output_backend = "svg"
bokeh.io.show(data_stripbox)
# -
# # Kruskal-Wallis Test with Dunn's Multiple Comparisons Correction
# Useful for comparing multiple datasets
# Reminder of treatments to compare
treatment_list
# +
### Kruskal-Wallis test
# Define samples to compare
sample1 = '2aRFP'
sample2 = 'Dyn1K44A'
sample3 = 'SMPD3'
sample4 = 'SMPD3N130A'
# Run Kruskal-Wallis test
kw_result = stats.kruskal(
df_subset.loc[df_subset['Treatment']==sample1]['Count']
,df_subset.loc[df_subset['Treatment']==sample2]['Count']
,df_subset.loc[df_subset['Treatment']==sample3]['Count']
,df_subset.loc[df_subset['Treatment']==sample4]['Count']
)
# Dunn's Posthoc for Multiple Comparisons
mult_compar = scikit_posthocs.posthoc_dunn(df_subset, val_col='Count', group_col='Treatment'
, sort=False, p_adjust='bonferroni').round(10)
# Display test results
print('Kruskal-Wallis test results: \n\t\t\t\t statistic=' + str(kw_result[0]) +
'\n\t\t\t\t p-value=' + str(kw_result[1]))
print("\nDunn's posthoc multiple comparison result: \n" + str(mult_compar) +'\n')
# mult_compar.to_csv("Results of Dunn's Posthoc.csv")
# Get number of cells within this test
for treatment in df_subset['Treatment'].unique().tolist():
temp_df = df_subset.loc[df_subset['Treatment'] == treatment]
print('n = ' + str(len(temp_df)) + ' cells in the ' + str(treatment) + ' dataset.')
# -
| Figure5/Tf633_U2OS/U2OS_Tf633PunctaCounts_CatData.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Processing Credit Card Fraud data from Kaggle
import pandas as pd
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import train_test_split
# <p>Download and unzip Outlier Credit Card data from Kaggle:</p>
# <a href="https://www.kaggle.com/mlg-ulb/creditcardfraud/data">https://www.kaggle.com/mlg-ulb/creditcardfraud/data</a>
fn="creditcard.csv" #FROM THE KAGGLE CREDIT CARD FRAUD DATASET
df=pd.read_csv(fn)
df.head()
# <p>Since ML algorithms tend to be unbalanced if features aren't of the same scale we should reduce the two features that are not reduced by PCA (to standard-normal form).</p>
df['TimeSN']=StandardScaler().fit_transform(df['Time'].values.reshape(-1, 1))
df['AmountSN']=StandardScaler().fit_transform(df['Amount'].values.reshape(-1, 1))
df=df[['V1', 'V2', 'V3', 'V4', 'V5', 'V6', 'V7', 'V8', 'V9', 'V10',
'V11', 'V12', 'V13', 'V14', 'V15', 'V16', 'V17', 'V18', 'V19', 'V20',
'V21', 'V22', 'V23', 'V24', 'V25', 'V26', 'V27', 'V28', 'TimeSN', 'AmountSN',
'Class']]
# <p>Creating a test/train split with the test data going to be the data that is sent the Flogo app that the model hasn't seen before. The training data will be used to build the DNN.</p>
train_data,demo_data=train_test_split(df,test_size=0.10)
fnout='creditcard_cleaned.csv'
train_data.to_csv(fnout,index=False)
# <p>Extracting all the outliers out of 10% of data and downsampling the inliers to make it easier to see when the streams find an outlier. I then split this data into 3 files to use for the streaming app.</p>
# +
demo_data_out=demo_data[demo_data['Class']==1]
demo_data_in=demo_data[demo_data['Class']==0]
print(demo_data_in.shape[0],demo_data_out.shape[0])
numouts=demo_data_out.shape[0]
demo_upsampled=pd.concat([demo_data_in.head(numouts*10-numouts),demo_data_out]).sample(frac=1)
# -
cols1=['V1', 'V2', 'V3', 'V4', 'V5', 'V6', 'V7', 'V8', 'V9', 'V10', 'V11']#,'TimeSN', 'AmountSN']
cols2=['V12', 'V13', 'V14', 'V15', 'V16', 'V17', 'V18', 'V19', 'V20', 'V21']#,'TimeSN', 'AmountSN']
cols3=['V22', 'V23', 'V24', 'V25', 'V26', 'V27', 'V28', 'TimeSN', 'AmountSN','Class']
demo_upsampled[cols1].to_csv("zero.csv",index=True,header=True,index_label='index')
demo_upsampled[cols2].to_csv("one.csv",index=True,header=True,index_label='index')
demo_upsampled[cols3].to_csv("two.csv",index=True,header=True,index_label='index')
| examples/streamingOutlier/processing_cc_fraud_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <NAME>
# ## Task 3
# ## Decision Tree Classifier
# Task is on Iris Dataset, Create the Decision Tree Classifier and visualize it graphically
# ### Import Necessary Libraries
# +
import pandas as pd
import matplotlib.pyplot as plt
from sklearn import datasets
# %matplotlib inline
# -
# ### Load the Dataset
# df = pd.read_csv('C:/Users/HP/Downloads/IRIS.csv')
iris = datasets.load_iris()
X = iris.data
y = iris.target
X
y
# ### Decision Tree Classifier
# +
from sklearn.tree import DecisionTreeClassifier
clf = DecisionTreeClassifier(random_state = 1234)
clf.fit(X,y)
print('Decision Tree Classifier Created')
# -
# ### Visualize Decision Tree
from sklearn import tree
fig = plt.figure(figsize=(20,20))
_ = tree.plot_tree(clf, feature_names=iris.feature_names, class_names=iris.target_names, filled=True)
| Task#3_Decision_Tree.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# we firstly send request to the server.
from bs4 import BeautifulSoup
import requests
page=requests.get('https://www.icc-cricket.com/rankings/mens/player-rankings/odi')
page
soup= BeautifulSoup(page.content)
soup
# ### TOP 10 ODI bowlers with the record of team and rating.
# +
bowlers =[] # give empty list here
for i in soup.find_all('td',class_="table-body__cell name"):
bowlers.append(i.text.split('\n')[1])
bowlers= bowlers[:18]
bowlers
# -
# ## Team
teamaa=soup.find('span',class_="table-body__logo-text")
teamaa.text
##Not in use.
# +
team =[] # give empty list here
for i in soup.find_all('span',class_="table-body__logo-text"):
team.append(i.text)
## It will print all. not in use
# +
team =[] # give empty list here
for i in soup.find_all('span',class_="table-body__logo-text"):
team.append(i.text)
team= team[:18]
team
# -
# ## Rating
rating=soup.find('td',class_="table-body__cell u-text-right rating")
rating.text
# +
rating=[] # give empty list here
for i in soup.find_all('td',class_="table-body__cell u-text-right rating"):
rating.append(i.text)
rating= rating[:18]
## It will print all.
# -
rating
import pandas as pd
# print length
print(len(bowlers),len(team),len(rating))
# ## Top 10 ODI Bowlers
# +
df=pd.DataFrame({'Top 10 ODI bowlers':bowlers,'Team':team,'Rating':rating})
update_df = df.drop([df.index[0],df.index[1], df.index[2],df.index[3], df.index[4],df.index[5], df.index[6],df.index[7],df.index[8]])
update_df
# -
| 4.3)Top 10 ODI bowlers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="92m7Jz_iMifz" colab_type="text"
# Answer Number 1
# + id="ONItdYqoLCAA" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="a0e9ec68-5045-4432-c201-2fc2c78e65d8"
list = ["Indah", 123, 30, "good", [3,2,1]]
list
# + id="cVx97PYLMKvf" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="98d3f41a-39b0-4e52-b87c-b87e749f1865"
list[0]
# + id="OTHbrVV4MW8v" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="ff4ed860-959c-4b56-d025-2ff1b28df7d5"
list[4][2]
# + id="mo510r1UMbsS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="980118f5-7467-4245-d4a4-34948c37ca98"
list[-1]
# + [markdown] id="UcpezAjgMf81" colab_type="text"
# Answer Number 2
# + id="AucLPoILMpyv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 104} outputId="c655a496-5d75-4c31-e467-5682fbbdaef5"
dit = {"name":"Indah", "age":20,
"phone number":+6282186898121,"email":"<EMAIL>", "hobby":"writting"}
dit
# + id="dsk5Qu4ANlUU" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="a8e5c426-f72b-49de-df26-331f8c304ca5"
type(dit)
# + id="32Z1IlS3Nouj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="b4f04fca-0ed8-4d05-c33c-83f7308c9819"
dit['name']
# + id="Z0whd_plNtfE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="a8d47b13-e8e1-43e7-b469-aaaf2d995bdc"
dit.keys()
# + id="zJgT17euN7dZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="60c53ada-0b56-4ad0-8408-9dfc713087ba"
dit.values()
# + [markdown] id="5XoudKm4N_A9" colab_type="text"
# Answer Number 3
# + id="R1d_gQnWOBK9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="186184bd-de23-4d06-952d-27816483b1e8"
st = {"indah","letsupgrade",1,2,3,4,4,5,6,5,5}
st
# + id="-new6NV3OWrD" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="f4d61f9e-0875-48e3-c25c-6516438306b6"
type(st)
# + id="qnraUScoOaPz" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="9f57b087-afc9-4458-e5a4-80a5bb1546bf"
st1={"indah",1}
st1.issubset(st)
# + [markdown] id="r8IYTBdqO_Hq" colab_type="text"
# Answer Number 4
# + id="XSSTnpEPPOo4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="47a6d56b-ee61-4e25-bd57-f632c069aadf"
tup = ("indah","@","<EMAIL>","study","vacation")
tup
# + id="ljD-EusjPboY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="d43d7fde-968f-4674-8e5f-a4074bd22b27"
tup.count("@")
# + id="oqeds4PXPoqo" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="3188e05a-7499-468f-b799-a4954b15b479"
tup.index("vacation")
# + id="55xuJlTKPwzQ" colab_type="code" colab={}
Answer Number 5
# + id="xABlOXsDQJ1p" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="da3cda24-cd35-4d17-bbf5-555604e21e04"
firstName = "Indah"
lastName = "<NAME>"
myHobby = "writting"
myAmbition = "front end web developer"
myInstitute = "Polytechnic state of sriwijaya"
firstName
# + id="2XViOYwfQ-_o" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="90de78b4-b262-4959-d2e9-a127afc8abdc"
lastName
# + id="345Fwmk5RCAm" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="c8ba7510-9186-4ae1-b863-0796d52fa085"
type(firstName)
# + id="tc9JNW0ySALC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="178b52b7-8dbe-421e-87da-35c40ae0c0f2"
firstName + '' + lastName
| Day 2/Assignment/Day2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python3
# ---
# # Formal Simulated Inference
# 1. Define F (i.e. your model and assumptions)
# 2. Formalize test
# 3. Describe test statistic
# 4. A. Sample data from F∈ℱ0
# B. Sample data from F∈ℱA
# 5. A. Plot power vs n (i.e. perspective power analysis)
# B. Plot power vs n (i.e. perspective power analysis)
# 6. Apply to data
#
# ## Step 1: Define model and assumptions
# **Model**
#
# F(X,Y) | ~ Histogram Data
#
# a) $F(I,k) = \{ F(I|k)*F(k) \}$
#
# $F(k)= \{0,1\}$
#
# * 0 = Control Image
# * 1 = Cocaine Image
#
# b) $F(I,k) = \{ F(I|k)*F(k) \}$
#
# $F(k)= \{0,1\}$
#
# **Assumption**
#
# ## Step 2: Formalize test
# H0:
#
# 1. $F(I,0) > || = F(I,1)$
# 2. $F(I,0) < ||= F(I,1)$
#
# 1. The maximum gray value for Cocaine stimulated brain is equal to the control brain.
# 2. The maximum gray value for fear induced brain is equal the control brain.
#
# HA:
# 1. $F(I,0) < || != F(I,1)$
# 2. $F(I,0) > || != F(I,1)$
#
# 1. The maximum gray value for Cocaine stimulated brain is greater than the control brain.
# 2. The maximum gray value for fear induced brain is lower than the control brain.
#
# ## Step 3. Test Statistic
#
# $$F(i,j) = floor((L-1)sum_{n=0}^{F(i,j)} {P_n} )$$
#
# $P_n$ = Number of pixels of intensity n / Total number of pixels
#
# $F(I) = mean (F(i,j))$ % mean gray value in the image
#
# ## Requirements to run the program
# In order to run the program, we need to install the packages in python
# > [PyQt4](https://www.riverbankcomputing.com/software/pyqt/download) install from tar file
# >
# > [SIP](https://www.riverbankcomputing.com/software/sip/download) install from tar file
# >
# > matplotlib,numpy,jgraph,vispy,[nibabel](http://nipy.org/nibabel/installation.html) (use pip install or pip2 install)
# +
import os
PATH="/Users/david/Desktop/CourseWork/TheArtOfDataScience/claritycontrol/code/scripts/" # use your own path
os.chdir(PATH)
import clarity as cl # I wrote this module for easier operations on data
import matplotlib.pyplot as plt
import jgraph as ig
# %matplotlib inline
# -
# create a instance
c = cl.Clarity("Fear199")
# load image, to points # takes a while to process the data
# * threshold rate is used to cut off some noise data.
# * sample is sampling rate
# show histogram
c.loadImg().imgToPoints(threshold=0.02,sample=0.3).showHistogram(bins=256)
b = cl.Clarity("Cocaine174")
b.loadImg(info=False).imgToPoints(threshold=0.08,sample=0.1).showHistogram(bins=256)
a = cl.Clarity("Control239")
a.loadImg(info=False).imgToPoints(threshold=0.04,sample=0.3).showHistogram(bins=256)
# +
# save points to csv file
# load directly from points data, instead of the origin data
c.loadImg().imgToPoints(threshold=0.04,sample=0.5).savePoints()
# load from points data file
# show on 3D graph, GPU intensive
c.loadPoints().show()
# -
# A screen shot
from IPython.display import Image
from IPython.core.display import HTML
Image(url= "https://raw.githubusercontent.com/Upward-Spiral-Science/claritycontrol/master/figs/a04/3dvisualdemo.png")
# ## Reflect
#
# We can infer from the histogram plots that the maximum gray values in cocaine stimulated brain is higher than the control brain. This is conclusive with the alternative hypothesis. This also agrees with literature that indicates that the brain is more active when stimulated by cocaine, thereby lighting up more than the control brain.
| code/a04_inferential_simulation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Array Broadcasting: What is x+y?
#
# + slideshow={"slide_type": "-"}
import numpy as np
x = np.array([[0], [1], [2]])
y = np.array([[3, 4, 5]])
# -
print(x)
print(y)
x + y
# + [markdown] slideshow={"slide_type": "slide"}
# # Array Broadcasting
#
# A sensible way of doing elementwise operations on arrays of different (but compatible) shapes.
#
# $$\pmatrix{0 & 1 & 2\\ 3 & 4 & 5} + \pmatrix{1 & 2 & 3} = \pmatrix{1 & 3 & 5\\ 4 & 6 & 8}$$
#
# $$\pmatrix{0 & 1 & 2\\ 3 & 4 & 5} + \pmatrix{1 \\ 2} = \pmatrix{1 & 2 & 3 \\ 5 & 6 & 7}$$
#
# It works with plus, minus, times, exponentiation, min/max, and many more elementwise operations. Search the numpy docs for the word "broadcast" to see if it is supported.
# + [markdown] slideshow={"slide_type": "slide"}
# # Shape Compatibility Rules
#
# 1. If x, y have a different number of dimensions, prepend 1's to the shape of the shorter.
# 2. Any axis of length 1 can be repeated (broadcast) to the length of the other vector's length in that axis
# 3. All other axes must have matching lengths.
#
# Use these rules to compute whether the arrays are compatible and, if so, the broadcasted shape.
# + [markdown] slideshow={"slide_type": "slide"}
# # Example 1
# + slideshow={"slide_type": "-"}
x.shape == (2, 3)
y.shape == (2, 3) # compatible
y.shape == (2, 1) # compatible
y.shape == (1, 3) # compatible
y.shape == (3,) # compatible
# results in (2, 3) shape
y.shape == (3, 2) # NOT compatible
y.shape == (2,) # NOT compatible
# + [markdown] slideshow={"slide_type": "subslide"}
# # Example 2
# +
x.shape == (1000, 256, 256, 256)
y.shape == (1000, 256, 256, 256) # compatible
y.shape == (1000, 1, 256, 256) # compatible
y.shape == (1000, 1, 1, 256) # compatible
y.shape == (1, 256, 256, 256) # compatible
y.shape == (1, 1, 256, 1) # compatible
# results in (1000, 256, 256, 256) shape
y.shape == (1000, 256, 256) # NOT compatible
# + [markdown] slideshow={"slide_type": "subslide"}
# # Example 3
# +
x.shape == (1, 2, 3, 5, 1, 11, 1, 17)
y.shape == (1, 7, 1, 1, 17) # compatible
# results in shape (1, 2, 3, 5, 7, 11, 1, 17)
# + [markdown] slideshow={"slide_type": "slide"}
# # Once shapes match, use for-loop to understand
#
# For any axis with length 1, use the only possible value.
# + slideshow={"slide_type": "-"}
x = np.array([[0, 1, 2],
[3, 4, 5],
[6, 7, 8]])
y = np.array([1, 10, 100]).reshape(3, 1)
print(x + y)
# x (3, 3)
# y (3, 1)
shape = (3, 3)
out = np.empty(shape, dtype=int)
N0, N1 = shape
for i in range(N0):
for j in range(N1):
# in the dimension that y only has 1 element, just use it
out[i, j] = x[i, j] + y[i, 0]
print(out)
# + [markdown] slideshow={"slide_type": "subslide"}
# Just omit variables for prepended 1's.
# + slideshow={"slide_type": "-"}
x = np.array([[[0, 1, 2],
[3, 4, 5],
[6, 7, 8]],
[[9, 10, 11],
[12, 13, 14],
[15, 16, 17]]]) # shape (2, 3, 3)
y = np.array([1, 10, 100]) # shape (3,)
print(x + y)
# align and prepend
# x (2, 3, 3)
# y (1, 1, 3)
shape = (2, 3, 3)
out = np.empty(shape, dtype=int)
N0, N1, N2 = shape
for i in range(N0):
for j in range(N1):
for k in range(N2):
# leave off prepended indices of y
out[i, j, k] = x[i, j, k] + y[k]
print(out)
# + [markdown] slideshow={"slide_type": "subslide"}
# Both arrays can have broadcasted axes, not just one.
# + slideshow={"slide_type": "-"}
x = np.array([[0], [1], [2]]) # (3, 1)
y = np.array([[3, 4, 5]]) # (1, 3)
print(x + y)
shape = (3, 3)
out = np.empty(shape, dtype=int)
N0, N1 = shape
for i in range(N0):
for j in range(N1):
out[i, j] = x[i, 0] + y[0, j]
print(out)
# + [markdown] slideshow={"slide_type": "slide"}
# # Exercise for you
#
# You have N=1000 images of size WxH=32x32, where each image has C=3 channels (red, green, and blue pixel value from 0 to 255) for each location in the image.
#
# Suppose you have the images stored in an array x of size (N, C, W, H) == (1000, 3, 32, 32)
#
# What array y would you multiply by to scale every red pixel by 2, every green pixel by 3, and every blue pixel by 4 (don't worry about overflow)?
# + slideshow={"slide_type": "slide"}
# Answer
y = np.array([2, 3, 4]).reshape(1, 3, 1, 1)
# or
# y = np.array([2, 3, 4]).reshape(3, 1, 1)
# Understand with for-loops
x = np.ones(shape, dtype=np.uint8) # for example
shape = (1000, 3, 32, 32)
out = np.empty(shape, dtype=np.uint8)
N, C, W, H = shape
for n in range(N):
for channel in range(C):
for w in range(W):
for h in range(H):
out[n, channel, w, h] = x[n, channel, w, h] * y[0, channel, 0, 0]
| videos/032_numpy_broadcasting_explained/numpy_broadcasting.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="HGAuV0-Vzw7a"
from google.colab import auth
auth.authenticate_user()
# + id="BSOBoyJ9zw7e" colab={"base_uri": "https://localhost:8080/"} outputId="9f673864-38fd-4240-80ff-6778f9a2938d"
import pandas as pd
# https://cloud.google.com/resource-manager/docs/creating-managing-projects
project_id = 'thebridgept0521'
sample_count = 2000
row_count = pd.io.gbq.read_gbq('''
SELECT
COUNT(*) as total
FROM `bigquery-public-data.samples.gsod`
''', project_id=project_id).total[0]
df = pd.io.gbq.read_gbq(f'''
SELECT
*
FROM
`bigquery-public-data.samples.gsod`
WHERE RAND() < {sample_count}/{row_count}
''', project_id=project_id)
print(f'Full dataset has {row_count} rows')
# + id="qLaVK0Abzw7h" colab={"base_uri": "https://localhost:8080/", "height": 316} outputId="42d2dc5d-af2a-437b-eada-8f3908dd28cc"
df.describe()
# + colab={"base_uri": "https://localhost:8080/", "height": 223} id="8zuvkV0E1jZ-" outputId="d28011e4-4e16-47a9-9ba3-873b5b298c0f"
df.head(5)
| 02_DataAnalysis/labs/BigQuery/lab01_GoogleBigQuery_Pandas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# cd /media/datastorage/Phong/cifar10/cifar-10-batches-py/
# +
import numpy as np
import os
# mean_pred3_1 = np.load(os.path.join('pred_npy','Cifar10_Xception_MulModels299_GRU_L3.npy'))
mean_pred3_2 = np.load(os.path.join('pred_npy','Cifar10_SEResNext101_LRG299_L3.npy'))
mean_pred3_3 = np.load(os.path.join('pred_npy','Cifar10_LRG299_Mul_Pretrained_V3_3MIcpRes_STD_L1.npy'))
mean_pred3_5 = np.load(os.path.join('pred_npy','Cifar10_Eff_B5_345_L2.npy'))
mean_pred3 = (mean_pred3_5+mean_pred3_2+mean_pred3_3)/3
# print(mean_pred3[0:9])
# +
# mean_a = (a1+a2+a3+a4)/4
from keras.preprocessing.image import ImageDataGenerator
from math import ceil
import numpy as np
from keras_applications.imagenet_utils import preprocess_input
# PREDICT ON OFFICIAL TEST
train_datagen = ImageDataGenerator(
# rescale = 1./255,
rotation_range=30,
width_shift_range=0.3,
height_shift_range=0.3,
shear_range=0.3,
zoom_range=0.3,
horizontal_flip=True,
vertical_flip=True,##
# brightness_range=[0.5, 1.5],##
channel_shift_range=10,##
fill_mode='nearest',
preprocessing_function=preprocess_input,
)
test_datagen1 = ImageDataGenerator(
# rescale = 1./255,
preprocessing_function=preprocess_input
)
batch_size = 30
train_set = train_datagen.flow_from_directory('train_resized_299',
target_size = (299, 299),
batch_size = batch_size,
class_mode = 'categorical',
shuffle=True,
seed=7,
# subset="training"
)
test_set1 = test_datagen1.flow_from_directory('test_resized_299',
target_size = (299, 299),
batch_size = batch_size,
class_mode = 'categorical',
shuffle=False,
seed=7,
# subset="validation"
)
predicted_class_indices_mean=np.argmax(mean_pred3,axis=1)
labels = (train_set.class_indices)
labels = dict((v,k) for k,v in labels.items())
finalpre = [labels[k] for k in predicted_class_indices_mean]
import pandas as pd
filenames=test_set1.filenames
results=pd.DataFrame({"id":filenames,
"predicted":finalpre,
})
results.to_csv('Cifar10_MulModels299_AVG_Assembe_2608_v1.csv')
results.head(20)
# +
mean_pred3_1 = np.load(os.path.join('pred_npy','Cifar10_Xception_LRG299_L2.npy'))
mean_pred3_2 = np.load(os.path.join('pred_npy','Cifar10_SEResNext101_LRG299_L3.npy'))
mean_pred3_3 = np.load(os.path.join('pred_npy','Cifar10_LRG299_Mul_Pretrained_V3_3MIcpRes_STD_L1.npy'))
mean_pred3_5 = np.load(os.path.join('pred_npy','Cifar10_Eff_B5_345_L2.npy'))
mean_pred1235 = (mean_pred3_5+mean_pred3_2+mean_pred3_3+mean_pred3_1)/4
# print(mean_pred3[0:9])
# +
# mean_a = (a1+a2+a3+a4)/4
from keras.preprocessing.image import ImageDataGenerator
from math import ceil
import numpy as np
from keras_applications.imagenet_utils import preprocess_input
# PREDICT ON OFFICIAL TEST
train_datagen = ImageDataGenerator(
# rescale = 1./255,
rotation_range=30,
width_shift_range=0.3,
height_shift_range=0.3,
shear_range=0.3,
zoom_range=0.3,
horizontal_flip=True,
vertical_flip=True,##
# brightness_range=[0.5, 1.5],##
channel_shift_range=10,##
fill_mode='nearest',
preprocessing_function=preprocess_input,
)
test_datagen1 = ImageDataGenerator(
# rescale = 1./255,
preprocessing_function=preprocess_input
)
batch_size = 30
train_set = train_datagen.flow_from_directory('train_resized_299',
target_size = (299, 299),
batch_size = batch_size,
class_mode = 'categorical',
shuffle=True,
seed=7,
# subset="training"
)
test_set1 = test_datagen1.flow_from_directory('test_resized_299',
target_size = (299, 299),
batch_size = batch_size,
class_mode = 'categorical',
shuffle=False,
seed=7,
# subset="validation"
)
predicted_class_indices_mean=np.argmax(mean_pred1235,axis=1)
labels = (train_set.class_indices)
labels = dict((v,k) for k,v in labels.items())
finalpre = [labels[k] for k in predicted_class_indices_mean]
import pandas as pd
filenames=test_set1.filenames
results=pd.DataFrame({"id":filenames,
"predicted":finalpre,
})
results.to_csv('Cifar10_MulModels299_AVG_Assembe_2608_v2.csv')
results.head(10)
# -
# cp Cifar10_MulModels299_AVG_Assembe_2608_v2.csv /home/bribeiro/Phong/Nat19/Cifar10_2608_v2.csv
# +
mean_pred3_6 = np.load(os.path.join('pred_npy','Cifar10_EfficientB0_299_STD_L2.npy'))#!!!
mean_pred12356 = (mean_pred3_5+mean_pred3_2+mean_pred3_3+mean_pred3_1+mean_pred3_6)/5
# +
predicted_class_indices_mean=np.argmax(mean_pred12356,axis=1)
labels = (train_set.class_indices)
labels = dict((v,k) for k,v in labels.items())
finalpre = [labels[k] for k in predicted_class_indices_mean]
import pandas as pd
filenames=test_set1.filenames
results=pd.DataFrame({"id":filenames,
"predicted":finalpre,
})
results.to_csv('Cifar10_MulModels299_AVG_Assembe_2608_v3.csv')
# results.head(10)
# -
# cp Cifar10_MulModels299_AVG_Assembe_2608_v3.csv /home/bribeiro/Phong/Nat19/Cifar10_2608_v3.csv
# +
mean_pred2356 = (mean_pred3_5+mean_pred3_2+mean_pred3_3+mean_pred3_6)/4
predicted_class_indices_mean=np.argmax(mean_pred2356,axis=1)
labels = (train_set.class_indices)
labels = dict((v,k) for k,v in labels.items())
finalpre = [labels[k] for k in predicted_class_indices_mean]
import pandas as pd
filenames=test_set1.filenames
results=pd.DataFrame({"id":filenames,
"predicted":finalpre,
})
results.to_csv('Cifar10_MulModels299_AVG_Assembe_2608_v4.csv')
# results.head(10)
# -
# cp Cifar10_MulModels299_AVG_Assembe_2608_v4.csv /home/bribeiro/Phong/Nat19/Cifar10_2608_v4.csv
# +
mean_pred3_7 = np.load(os.path.join('pred_npy','Cifar10_EfficientB7_299_STD_L3.npy'))
mean_pred2357 = (mean_pred3_2+mean_pred3_3+mean_pred3_5+mean_pred3_7)/4
# +
predicted_class_indices_mean=np.argmax(mean_pred2357,axis=1)
labels = (train_set.class_indices)
labels = dict((v,k) for k,v in labels.items())
finalpre = [labels[k] for k in predicted_class_indices_mean]
import pandas as pd
filenames=test_set1.filenames
results=pd.DataFrame({"id":filenames,
"predicted":finalpre,
})
results.to_csv('Cifar10_MulModels299_AVG_Assembe_2708_v1.csv')
# -
# cp Cifar10_MulModels299_AVG_Assembe_2708_v1.csv /home/bribeiro/Phong/Nat19/Cifar10_2708_v1.csv
mean_pred3_6 = np.load(os.path.join('pred_npy','Cifar10_EfficientB0_299_STD_L2.npy'))#!!!
mean_pred23567 = (mean_pred3_2+mean_pred3_3+mean_pred3_5+mean_pred3_6+mean_pred3_7)/5
# +
predicted_class_indices_mean=np.argmax(mean_pred23567,axis=1)
labels = (train_set.class_indices)
labels = dict((v,k) for k,v in labels.items())
finalpre = [labels[k] for k in predicted_class_indices_mean]
import pandas as pd
filenames=test_set1.filenames
results=pd.DataFrame({"id":filenames,
"predicted":finalpre,
})
results.to_csv('Cifar10_MulModels299_AVG_Assembe_2708_v2.csv')
# -
# cp Cifar10_MulModels299_AVG_Assembe_2708_v2.csv /home/bribeiro/Phong/Nat19/Cifar10_2708_v2.csv
mean_pred3_1 = np.load(os.path.join('pred_npy','Cifar10_Xception_LRG299_L2.npy'))
mean_pred123567 = (mean_pred3_1+mean_pred3_2+mean_pred3_3+mean_pred3_5+mean_pred3_6+mean_pred3_7)/6
# +
predicted_class_indices_mean=np.argmax(mean_pred123567,axis=1)
labels = (train_set.class_indices)
labels = dict((v,k) for k,v in labels.items())
finalpre = [labels[k] for k in predicted_class_indices_mean]
import pandas as pd
filenames=test_set1.filenames
results=pd.DataFrame({"id":filenames,
"predicted":finalpre,
})
results.to_csv('Cifar10_MulModels299_AVG_Assembe_2708_v3.csv')
# -
# cp Cifar10_MulModels299_AVG_Assembe_2708_v3.csv /home/bribeiro/Phong/Nat19/Cifar10_2708_v3.csv
# +
mean_pred257 = (mean_pred3_2+mean_pred3_5+mean_pred3_7)/3
predicted_class_indices_mean=np.argmax(mean_pred257,axis=1)
labels = (train_set.class_indices)
labels = dict((v,k) for k,v in labels.items())
finalpre = [labels[k] for k in predicted_class_indices_mean]
import pandas as pd
filenames=test_set1.filenames
results=pd.DataFrame({"id":filenames,
"predicted":finalpre,
})
results.to_csv('Cifar10_MulModels299_AVG_Assembe_2708_v6.csv')
# -
# cp Cifar10_MulModels299_AVG_Assembe_2708_v6.csv /home/bribeiro/Phong/Nat19/Cifar10_2708_v6.csv
a1 = mean_pred3_1
a2 = mean_pred3_2
a3 = mean_pred3_3
# a4 = mean_pred3_4
a5 = mean_pred3_5
a6 = mean_pred3_6
a7 = mean_pred3_7
x1 = a1
x2 = a2
x3 = a3
# x4 = a4
x5 = a5
x6 = a6
x7 = a7
#x1
for i in range(len(x1)):
i_max = np.argmax(x1[i], axis=0)
for idx in range(10):
if idx != i_max:
x1[i][idx] = 0
else:
x1[i][idx] = 1
# print(a1)
#x2
for i in range(len(x2)):
i_max = np.argmax(x2[i], axis=0)
for idx in range(10):
if idx != i_max:
x2[i][idx] = 0
else:
x2[i][idx] = 1
# print(a1)
#x3
for i in range(len(x3)):
i_max = np.argmax(x3[i], axis=0)
for idx in range(10):
if idx != i_max:
x3[i][idx] = 0
else:
x3[i][idx] = 1
# print(a1)
#x5
for i in range(len(x5)):
i_max = np.argmax(x5[i], axis=0)
for idx in range(10):
if idx != i_max:
x5[i][idx] = 0
else:
x5[i][idx] = 1
# print(a1)
#x6
for i in range(len(x6)):
i_max = np.argmax(x6[i], axis=0)
for idx in range(10):
if idx != i_max:
x6[i][idx] = 0
else:
x6[i][idx] = 1
# print(a1)
#x7
for i in range(len(x7)):
i_max = np.argmax(x7[i], axis=0)
for idx in range(10):
if idx != i_max:
x7[i][idx] = 0
else:
x7[i][idx] = 1
# print(a1)
mean_x2357 = (x2+x3+x5+x7)/4
# +
predicted_class_indices_mean=np.argmax(mean_x2357,axis=1)
labels = (train_set.class_indices)
labels = dict((v,k) for k,v in labels.items())
finalpre = [labels[k] for k in predicted_class_indices_mean]
import pandas as pd
filenames=test_set1.filenames
results=pd.DataFrame({"id":filenames,
"predicted":finalpre,
})
results.to_csv('Cifar10_MulModels299_AVG_Assembe_2708_v4.csv')
# -
# cp Cifar10_MulModels299_AVG_Assembe_2708_v4.csv /home/bribeiro/Phong/Nat19/Cifar10_2708_v4.csv
# +
mean_x235 = (x2+x3+x5)/3
predicted_class_indices_mean=np.argmax(mean_x235,axis=1)
labels = (train_set.class_indices)
labels = dict((v,k) for k,v in labels.items())
finalpre = [labels[k] for k in predicted_class_indices_mean]
import pandas as pd
filenames=test_set1.filenames
results=pd.DataFrame({"id":filenames,
"predicted":finalpre,
})
results.to_csv('Cifar10_MulModels299_AVG_Assembe_2708_v5.csv')
# -
# cp Cifar10_MulModels299_AVG_Assembe_2708_v5.csv /home/bribeiro/Phong/Nat19/Cifar10_2708_v5.csv
# +
mean_pred3_8 = np.load(os.path.join('pred_npy','Cifar10_EfficientB4_299_STD_L3.npy'))
a8 = mean_pred3_8
mean_pred23578 = (mean_pred3_2+mean_pred3_3+mean_pred3_5+mean_pred3_7+mean_pred3_8)/5
# +
predicted_class_indices_mean=np.argmax(mean_pred23578,axis=1)
labels = (train_set.class_indices)
labels = dict((v,k) for k,v in labels.items())
finalpre = [labels[k] for k in predicted_class_indices_mean]
import pandas as pd
filenames=test_set1.filenames
results=pd.DataFrame({"id":filenames,
"predicted":finalpre,
})
results.to_csv('Cifar10_MulModels299_AVG_Assembe_mean_pred23578_2808_v1.csv')
# -
# cp Cifar10_MulModels299_AVG_Assembe_mean_pred23578_2808_v1.csv /home/bribeiro/Phong/Nat19/Cifar10_2808_v1.csv
# +
mean_pred2578 = (mean_pred3_2+mean_pred3_5+mean_pred3_7+mean_pred3_8)/4
predicted_class_indices_mean=np.argmax(mean_pred2578,axis=1)
labels = (train_set.class_indices)
labels = dict((v,k) for k,v in labels.items())
finalpre = [labels[k] for k in predicted_class_indices_mean]
import pandas as pd
filenames=test_set1.filenames
results=pd.DataFrame({"id":filenames,
"predicted":finalpre,
})
results.to_csv('Cifar10_MulModels299_AVG_Assembe_mean_pred2578_2808_v1.csv')
# -
# cp Cifar10_MulModels299_AVG_Assembe_mean_pred2578_2808_v1.csv /home/bribeiro/Phong/Nat19/Cifar10_2808_v2.csv
x8 = a8
#x7
for i in range(len(x8)):
i_max = np.argmax(x8[i], axis=0)
for idx in range(10):
if idx != i_max:
x8[i][idx] = 0
else:
x8[i][idx] = 1
# print(a1)
# +
mean_x23578 = (x2+x3+x5+x7+x8)/5
predicted_class_indices_mean=np.argmax(mean_x23578,axis=1)
labels = (train_set.class_indices)
labels = dict((v,k) for k,v in labels.items())
finalpre = [labels[k] for k in predicted_class_indices_mean]
import pandas as pd
filenames=test_set1.filenames
results=pd.DataFrame({"id":filenames,
"predicted":finalpre,
})
results.to_csv('Cifar10_MulModels299_AVG_Assembe_mean_x23578_2808_v3.csv')
# -
# cp Cifar10_MulModels299_AVG_Assembe_mean_x23578_2808_v3.csv /home/bribeiro/Phong/Nat19/Cifar10_2808_v3.csv
# +
mean_pred235678 = (mean_pred3_2+mean_pred3_3+mean_pred3_5+mean_pred3_6+mean_pred3_7+mean_pred3_8)/6
predicted_class_indices_mean=np.argmax(mean_pred235678,axis=1)
labels = (train_set.class_indices)
labels = dict((v,k) for k,v in labels.items())
finalpre = [labels[k] for k in predicted_class_indices_mean]
import pandas as pd
filenames=test_set1.filenames
results=pd.DataFrame({"id":filenames,
"predicted":finalpre,
})
results.to_csv('Cifar10_MulModels299_AVG_Assembe_mean_pred235678_2808_v1.csv')
# -
# cp Cifar10_MulModels299_AVG_Assembe_mean_pred235678_2808_v1.csv /home/bribeiro/Phong/Nat19/Cifar10_2808_v4.csv
# +
mean_pred25678 = (mean_pred3_2+mean_pred3_5+mean_pred3_6+mean_pred3_7+mean_pred3_8)/6
predicted_class_indices_mean=np.argmax(mean_pred25678,axis=1)
labels = (train_set.class_indices)
labels = dict((v,k) for k,v in labels.items())
finalpre = [labels[k] for k in predicted_class_indices_mean]
import pandas as pd
filenames=test_set1.filenames
results=pd.DataFrame({"id":filenames,
"predicted":finalpre,
})
results.to_csv('Cifar10_MulModels299_AVG_Assembe_mean_pred25678_2808_v1.csv')
# -
# cp Cifar10_MulModels299_AVG_Assembe_mean_pred25678_2808_v1.csv /home/bribeiro/Phong/Nat19/Cifar10_2808_v5.csv
| Cifar10/v2/Cifar10_MulModels299_AVG_Ensembling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# parser for ATF files about shuila prayers
## Import packages
from pathlib import Path
import re
import pandas as pd
import numpy as np
import csv
import glob
import spacy
from spacy.symbols import nsubj, VERB, ADJ
from spacy.lang.en import English
# start code for extracting text
# THIS IS FOR MESOPOTAMIAN SHUILA PRAYERS
nlp = spacy.load("en_core_web_md")
Madjectives = []
Mverbs = []
path = '../EADHproject/shuila/*.atf'
outpath = './shuila/'
for file in glob.glob(path):
with open(file, encoding='utf-8', errors='ignore') as file_in:
text = ""
t = ""
firstline = file_in.readline()
#print (firstline)
# first parse the header code for the text
header = firstline.split('&')[1].split(" ")[0]
# print (header)
# then parse the english text to a simple string
lines = file_in.readlines()
for line in lines:
if "#tr.en:" in line:
t = line.replace("#tr.en: ", "")
text = text + " " + t
# now remove brackets
text = text.replace("[", "")
text = text.replace("]", "")
# get rid of (parenthetical comments)
text = re.sub("[\(\[].*?[\)\]]", "", text)
# remove lots of periods
text = text.replace(". .", "")
text = text.replace(" ", " ")
text = text.replace("\n", " ")
# output cleaned text files.
with open(outpath + header + '.txt', 'w') as f:
f.write(text)
#then apply nlp
doc = nlp(text)
for sent in doc.sents: #iterate over sentences
for word in sent: #iterate over words to find verbs
if word.pos == VERB:
Mverbs.append(word.lemma_)
if word.pos == ADJ:
Madjectives.append(word.lemma_)
# +
# now do PSALMS
Badjectives = []
Bverbs = []
Bpath = '../EADHproject/bible/psalms.txt'
for file in glob.glob(Bpath):
with open(file, encoding='utf-8', errors='ignore') as file_in:
text = ""
result = ""
# then parse the english text to a simple string
lines = file_in.readlines()
for line in lines:
result = ''.join([i for i in line if not i.isdigit()]) #eliminate digits
text = text + " " + result
# get rid of [(parenthetical comments)]
text = re.sub("[\(\[].*?[\)\]]", "", text)
text = text.replace("PSALMS", "")
# remove lots of periods
text = text.replace(". .", "")
text = text.replace(" ", " ")
text = text.replace(" ", " ")
text = text.replace(" ", " ")
text = text.replace("\n", " ")
#print (text)
# output cleaned text files.
#with open(outpath + header + '.txt', 'w') as f:
# f.write(text)
#then apply nlp
doc = nlp(text)
for sent in doc.sents: #iterate over sentences
for word in sent: #iterate over words to find verbs
if word.pos == VERB:
Bverbs.append(word.lemma_)
if word.pos == ADJ:
Badjectives.append(word.lemma_)
# +
#convert it to dictionary with values and its occurences
import matplotlib.pyplot as plt
from collections import Counter
from wordcloud import WordCloud
Mverb_counts = Counter(Mverbs)
Madj_counts = Counter(Madjectives)
Bverb_counts = Counter(Bverbs)
Badj_counts = Counter(Badjectives)
# put counts in a dataframe
df_verbs = pd.DataFrame(Bverb_counts.items(), columns = ['verbs', 'Bfrequency'])
df_verbs['Mfrequency'] = df_verbs['verbs'].map(Mverb_counts) # THIS IS LOSING WORDS THAT ARE NOT PRESENT IN BIBLE!!
df_verbs['Mnormalized'] = df_verbs['Mfrequency']/len(df_verbs)
df_verbs['Bnormalized'] = df_verbs['Bfrequency']/len(df_verbs)
df_adj = pd.DataFrame(Badj_counts.items(), columns = ['adjectives', 'Bfrequency'])
df_adj['Mfrequency'] = df_adj['adjectives'].map(Madj_counts) # THIS IS LOSING WORDS THAT ARE NOT PRESENT IN BIBLE!!
df_adj['Mnormalized'] = df_adj['Mfrequency']/len(df_adj)
df_adj['Bnormalized'] = df_adj['Bfrequency']/len(df_adj)
df_adj.to_csv('../EADHproject/adjective_frequency.csv')
df_verbs.to_csv('../EADHproject/verb_frequency.csv')
# +
# shorten dataframes to keep nonzero value words
df_adj = df_adj.drop(df_adj[df_adj.Mfrequency < 3].index)
df_verbs = df_verbs.drop(df_verbs[df_verbs.Mfrequency < 3].index)
print(df_adj)
# +
# measure similarity between words so we can make groups
# output a similarity matrix that compares every word currently in data to every other phrase
# change variable here to switch between adjectives and verbs
d = df_adj
nNodes = d.shape[0]
similarity_df = pd.DataFrame()
similarity_df = similarity_df.astype(np.float)
sourceList = []
targetList = []
similarityList = []
# Make a giant nSources x nTargets incidence matrix (data frame)
all_sources = list(d.adjectives) #this needs changing to verbs for the other matrix
all_targets = all_sources
# One row per source
similarity_df = pd.DataFrame(index=np.arange(len(all_sources)), columns=all_targets)
line = -1
for i in similarity_df:
#print (i)
line +=1
for j in similarity_df:
source = nlp(i)
target = nlp(j)
similarity = source.similarity(target) #calculate cosine similarity between the two phrases
similarity_df.at[line,j] = similarity
similarity_df.head()
similarity_df.to_csv("../EADHproject/similarity.csv")
# +
# look for similarity values above some level and output clusters of terms
inverse_similarity_df = similarity_df
from sklearn.decomposition import PCA
pca = PCA(2) # we need 2 principal components. THIS ALGORITHM REDUCES DIMENSION FROM 82 X 82 --> 82 X 2.
converted_data = pca.fit_transform(inverse_similarity_df)
converted_data.shape
# -
plt.style.use('seaborn-whitegrid')
plt.figure(figsize = (10,6))
c_map = plt.cm.get_cmap('jet', 10)
plt.scatter(converted_data[:, 0], converted_data[:, 1], s = 15,
cmap = c_map , c = inverse_similarity_df.index)
plt.colorbar()
plt.xlabel('PC-1') , plt.ylabel('PC-2')
plt.show()
# +
#Import required module
from sklearn.cluster import KMeans
#Initialize the class object
kmeans = KMeans(n_clusters= 5)
#predict the labels of clusters.
label = kmeans.fit_predict(converted_data)
print(label) #output the clusters of keywords
x = 0
for x in np.arange(5):
print (f"cluster {x}")
for i in np.arange(len(all_sources)):
if label[i] == x:
print (all_sources[i])
# -
# +
#Getting unique labels
u_labels = np.unique(label)
#plotting the results:
for i in u_labels:
plt.scatter(converted_data[label == i , 0] , converted_data[label == i , 1] , label = i)
plt.legend()
plt.show()
# -
# +
# make wordclouds to see lemma frequency
wordcloud = WordCloud(width = 1000, height = 500).generate_from_frequencies(Badj_counts)
plt.figure(figsize=(15,8))
plt.imshow(wordcloud)
plt.axis("off")
plt.show()
#plt.savefig('yourfile.png', bbox_inches='tight')
#plt.close()
# -
| ATFparse.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# !pip install rasterio matplotlib
import ipywidgets as widgets
import rasterio as rio
from matplotlib import pyplot as plt
from rasterio.plot import show
import numpy as np
from IPython.display import display
# +
# Data - Results of economic, environmental and social evaluation
# paths
p_out = 'scn_data/opt_out.tif'
p_a1_econ = 'scn_data/ecn_a1.tif'
p_a2_econ = 'scn_data/ecn_a2.tif'
p_a3_econ = 'scn_data/ecn_a3.tif'
p_a4_econ = 'scn_data/ecn_a4.tif'
p_a1_env = 'scn_data/env_a1.tif'
p_a2_env = 'scn_data/env_a2.tif'
p_a3_env = 'scn_data/env_a3.tif'
p_a4_env = 'scn_data/env_a4.tif'
p_a1_soc = 'scn_data/soc_a1.tif'
p_a2_soc = 'scn_data/soc_a2.tif'
p_a3_soc = 'scn_data/soc_a3.tif'
p_a4_soc = 'scn_data/soc_a4.tif'
''''
# Open as rasterio rasters
a1_ecn = rio.open(p_a1_econ)
a2_ecn = rio.open(p_a2_econ)
a3_ecn = rio.open(p_a3_econ)
a4_ecn = rio.open(p_a4_econ)
a1_env = rio.open(p_a1_env)
a2_env = rio.open(p_a2_env)
a3_env = rio.open(p_a3_env)
a4_env = rio.open(p_a4_env)
a1_soc = rio.open(p_a1_soc)
a2_soc = rio.open(p_a2_soc)
a3_soc = rio.open(p_a3_soc)
a4_soc = rio.open(p_a4_soc)
'''
# Open as rasterio rasters to numpy arrays
a1_ecn = rio.open(p_a1_econ).read()
a2_ecn = rio.open(p_a2_econ).read()
a3_ecn = rio.open(p_a3_econ).read()
a4_ecn = rio.open(p_a4_econ).read()
a1_env = rio.open(p_a1_env).read()
a2_env = rio.open(p_a2_env).read()
a3_env = rio.open(p_a3_env).read()
a4_env = rio.open(p_a4_env).read()
a1_soc = rio.open(p_a1_soc).read()
a2_soc = rio.open(p_a2_soc).read()
a3_soc = rio.open(p_a3_soc).read()
a4_soc = rio.open(p_a4_soc).read()
opt_out = rio.open(p_out)
# +
from ipywidgets import interact
from matplotlib.colors import ListedColormap
cmap = ListedColormap(["white","darkorange", "gold", "lawngreen", "lightseagreen"])
bounds = [-0.5,0.5,1.5,2.5,3.5,4.5]
ticks = [0,1,2,3,4]
def score():
def view_image(k1,k2,k3):
a1 = k1*a1_ecn + k2*a1_env + k3*a1_soc
a2 = k1*a2_ecn + k2*a2_env + k3*a2_soc
a3 = k1*a3_ecn + k2*a3_env + k3*a3_soc
a4 = k1*a4_ecn + k2*a4_env + k3*a4_soc
optim = np.where((a1>a2) & (a1>a3) & (a1>a4), 1,0)
optim = np.where((a2>a1) & (a2>a3) & (a2>a4), 2,optim)
optim = np.where((a3>a1) & (a3>a2) & (a3>a4), 3,optim)
optim = np.where((a4>a1) & (a4>a3) & (a4>a1), 4,optim)
plt.imshow(optim[0], cmap=cmap, interpolation='nearest')
plt.title("Optimized Sanitation Scenarios")
plt.colorbar(boundaries=bounds, ticks=ticks, label = 'Alternative')
plt.show()
interact(view_image,
k1=widgets.IntSlider(
value=5,
min=0,
max=10,
step=1,
description='Economic'),
k2=widgets.IntSlider(
value=5,
min=0,
max=10,
step=1,
description='Environmental'),
k3=widgets.IntSlider(
value=5,
min=0,
max=10,
step=1,
description='Social'))
return
# -
score()
| scenario_app2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:py3]
# language: python
# name: conda-env-py3-py
# ---
# +
from sklearn import tree
from sklearn.model_selection import train_test_split
import numpy as np
from sklearn.datasets import load_iris
from sklearn.datasets import load_breast_cancer
from sklearn.tree import DecisionTreeClassifier, export_graphviz
from sklearn.neural_network import MLPClassifier
from sklearn.externals.six import StringIO
from IPython.display import Image, display
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import Normalizer,OneHotEncoder, MinMaxScaler, StandardScaler, RobustScaler, QuantileTransformer, LabelEncoder
from sklearn.model_selection import train_test_split, StratifiedKFold, GridSearchCV, cross_val_score
from sklearn.model_selection import LeaveOneOut
from sklearn.pipeline import make_pipeline
from sklearn.model_selection import train_test_split
import mlrose
from sklearn.metrics import accuracy_score
bc = load_breast_cancer()
data=bc.data
target=bc.target
# Initialize neural network object and fit object
data = load_breast_cancer()
X_train, X_test, y_train, y_test = train_test_split(data.data, data.target, \
test_size = 0.2, random_state = 3)
# Normalize feature data
scaler = MinMaxScaler()
X_train_scaled = scaler.fit_transform(X_train)
X_test_scaled = scaler.transform(X_test)
# One hot encode target values
one_hot = OneHotEncoder()
y_train_hot = one_hot.fit_transform(y_train.reshape(-1, 1)).todense()
y_test_hot = one_hot.transform(y_test.reshape(-1, 1)).todense()
# +
NUM_TIMS_TO_RUN = 100
MAX_ITTERS = 3000
sum_test_accuracy_list=np.array([0,0])
sum_train_accuracy_list=np.array([0,0])
for j in range(NUM_TIMS_TO_RUN):
test_accuracy_list=[]
train_accuracy_list = []
for i in range(100, MAX_ITTERS,300):
#print(i)
nn_model2 = mlrose.NeuralNetwork(hidden_nodes = [2], activation = 'relu', \
algorithm = 'gradient_descent', max_iters = i, \
bias = True, is_classifier = True, learning_rate = 0.0001, \
early_stopping = True, clip_max = 5, max_attempts = 10)
nn_model2.fit(X_train_scaled, y_train_hot)
# Predict labels for train set and assess accuracy
# y_train_pred = nn_model2.predict(X_train_scaled)
# y_train_accuracy = accuracy_score(y_train_hot, y_train_pred)
# #print(y_train_accuracy)
# train_accuracy_list.append([i, 1-y_train_accuracy])
# Predict labels for test set and assess accuracy
y_test_pred = nn_model2.predict(X_test_scaled)
y_test_accuracy = accuracy_score(y_test_hot, y_test_pred)
test_accuracy_list.append([i, 1-y_test_accuracy])
sum_test_accuracy_list = sum_test_accuracy_list + np.array(test_accuracy_list)
# sum_train_accuracy_list = sum_train_accuracy_list + np.array(train_accuracy_list)
avg_test_prediction= sum_test_accuracy_list/NUM_TIMS_TO_RUN
# avg_train_prediction =sum_train_accuracy_list/NUM_TIMS_TO_RUN
#print(avg_test_prediction, avg_train_prediction)
plt.xlabel('Iterrations')
plt.ylabel('Error')
plt.ylim((0,1))
plt.plot(avg_test_prediction[:,0],avg_test_prediction[:,1], label='Test')
# plt.plot(avg_train_prediction[:,0],avg_train_prediction[:,1], label='Train')
plt.title('Error vs Iterration')
plt.legend()
plt.show()
# +
NUM_TIMS_TO_RUN = 10
MAX_ITTERS = 5000
sum_test_accuracy_list=np.array([0,0])
sum_train_accuracy_list=np.array([0,0])
for j in range(NUM_TIMS_TO_RUN):
test_accuracy_list=[]
train_accuracy_list = []
for i in range(100, MAX_ITTERS,500):
#print(i)
nn_model2 = mlrose.NeuralNetwork(hidden_nodes = [2], activation = 'relu', \
algorithm = 'random_hill_climb', max_iters = i, \
bias = True, is_classifier = True, learning_rate = 0.5, \
early_stopping = True, clip_max = 5, max_attempts = 600)
nn_model2.fit(X_train_scaled, y_train_hot)
# Predict labels for train set and assess accuracy
# y_train_pred = nn_model2.predict(X_train_scaled)
# y_train_accuracy = accuracy_score(y_train_hot, y_train_pred)
# #print(y_train_accuracy)
# train_accuracy_list.append([i, 1-y_train_accuracy])
# Predict labels for test set and assess accuracy
y_test_pred = nn_model2.predict(X_test_scaled)
y_test_accuracy = accuracy_score(y_test_hot, y_test_pred)
test_accuracy_list.append([i, 1-y_test_accuracy])
sum_test_accuracy_list = sum_test_accuracy_list + np.array(test_accuracy_list)
# sum_train_accuracy_list = sum_train_accuracy_list + np.array(train_accuracy_list)
avg_test_prediction= sum_test_accuracy_list/NUM_TIMS_TO_RUN
# avg_train_prediction =sum_train_accuracy_list/NUM_TIMS_TO_RUN
#print(avg_test_prediction, avg_train_prediction)
plt.xlabel('Iterrations')
plt.ylabel('Error')
plt.ylim((0,1))
plt.plot(avg_test_prediction[:,0],avg_test_prediction[:,1], label='Test')
# plt.plot(avg_train_prediction[:,0],avg_train_prediction[:,1], label='Train')
plt.title('Error vs Iterration')
plt.legend()
plt.show()
# +
NUM_TIMS_TO_RUN = 20
MAX_ITTERS = 3000
sum_test_accuracy_list=np.array([0,0])
sum_train_accuracy_list=np.array([0,0])
for j in range(NUM_TIMS_TO_RUN):
test_accuracy_list=[]
train_accuracy_list = []
for i in range(100, MAX_ITTERS,300):
#print(i)
nn_model2 = mlrose.NeuralNetwork(hidden_nodes = [2], activation = 'relu', \
algorithm = 'simulated_annealing', max_iters = i, \
bias = True, is_classifier = True, learning_rate = 0.3, \
early_stopping = True, clip_max = 5, max_attempts = 2000)
nn_model2.fit(X_train_scaled, y_train_hot)
# Predict labels for train set and assess accuracy
# y_train_pred = nn_model2.predict(X_train_scaled)
# y_train_accuracy = accuracy_score(y_train_hot, y_train_pred)
# #print(y_train_accuracy)
# train_accuracy_list.append([i, 1-y_train_accuracy])
# Predict labels for test set and assess accuracy
y_test_pred = nn_model2.predict(X_test_scaled)
y_test_accuracy = accuracy_score(y_test_hot, y_test_pred)
test_accuracy_list.append([i, 1-y_test_accuracy])
sum_test_accuracy_list = sum_test_accuracy_list + np.array(test_accuracy_list)
# sum_train_accuracy_list = sum_train_accuracy_list + np.array(train_accuracy_list)
avg_test_prediction= sum_test_accuracy_list/NUM_TIMS_TO_RUN
# avg_train_prediction =sum_train_accuracy_list/NUM_TIMS_TO_RUN
#print(avg_test_prediction, avg_train_prediction)
plt.xlabel('Iterrations')
plt.ylabel('Error')
plt.ylim((0,1))
plt.plot(avg_test_prediction[:,0],avg_test_prediction[:,1], label='Test')
# plt.plot(avg_train_prediction[:,0],avg_train_prediction[:,1], label='Train')
plt.title('Error vs Iterration')
plt.legend()
plt.show()
# +
NUM_TIMS_TO_RUN = 20
MAX_ITTERS = 1000
sum_test_accuracy_list=np.array([0,0])
sum_train_accuracy_list=np.array([0,0])
for j in range(NUM_TIMS_TO_RUN):
test_accuracy_list=[]
train_accuracy_list = []
for i in range(100, MAX_ITTERS,100):
#print(i)
nn_model2 = mlrose.NeuralNetwork(hidden_nodes = [2], activation = 'relu', \
algorithm = 'genetic_alg', max_iters = 1, \
bias = True, is_classifier = True, learning_rate = .1, \
mutation_prob = .1, pop_size = i,early_stopping = True, clip_max = 5, max_attempts = 10)
nn_model2.fit(X_train_scaled, y_train_hot)
# Predict labels for train set and assess accuracy
# y_train_pred = nn_model2.predict(X_train_scaled)
# y_train_accuracy = accuracy_score(y_train_hot, y_train_pred)
# #print(y_train_accuracy)
# train_accuracy_list.append([i, 1-y_train_accuracy])
# Predict labels for test set and assess accuracy
y_test_pred = nn_model2.predict(X_test_scaled)
y_test_accuracy = accuracy_score(y_test_hot, y_test_pred)
test_accuracy_list.append([i, 1-y_test_accuracy])
sum_test_accuracy_list = sum_test_accuracy_list + np.array(test_accuracy_list)
# sum_train_accuracy_list = sum_train_accuracy_list + np.array(train_accuracy_list)
avg_test_prediction= sum_test_accuracy_list/NUM_TIMS_TO_RUN
# avg_train_prediction =sum_train_accuracy_list/NUM_TIMS_TO_RUN
#print(avg_test_prediction, avg_train_prediction)
plt.xlabel('Population Size')
plt.ylabel('Error')
plt.ylim((0,1))
plt.plot(avg_test_prediction[:,0],avg_test_prediction[:,1], label='Test')
# plt.plot(avg_train_prediction[:,0],avg_train_prediction[:,1], label='Train')
plt.title('Error vs Population Size')
plt.legend()
plt.show()
# -
print(avg_test_prediction)
| breastCancer-iteration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8
# language: python
# name: python3
# ---
# # A notebook for running kernel thinning and standard thinning experiments
#
# +
import numpy as np
import numpy.random as npr
import numpy.linalg as npl
# from scipy.spatial.distance import pdist
from argparse import ArgumentParser
import pickle as pkl
import pathlib
import os
import os.path
# import kernel thinning
from goodpoints import kt # kt.thin is the main thinning function; kt.split and kt.swap are other important functions
from goodpoints.util import isnotebook # Check whether this file is being executed as a script or as a notebook
from goodpoints.util import fprint # for printing while flushing buffer
from goodpoints.tictoc import tic, toc # for timing blocks of code
# utils for generating samples, evaluating kernels, and mmds
from util_sample import sample, compute_mcmc_params_p, compute_diag_mog_params, sample_string
from util_k_mmd import kernel_eval, squared_mmd, get_combined_mmd_filename
# for partial functions, to use kernel_eval for kernel
from functools import partial
# set things a bit when running the notebook
if isnotebook():
# Autoreload packages that are modified
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
# %load_ext line_profiler
# https://jakevdp.github.io/PythonDataScienceHandbook/01.07-timing-and-profiling.html
# -
# If notebook run as a script, parse command-line arguments
if not isnotebook():
parser = ArgumentParser()
parser.add_argument('--rep0', '-r0', type=int, default=0,
help="starting experiment id")
parser.add_argument('--repn', '-rn', type=int, default=1,
help="number of experiment replication")
parser.add_argument('--store_K', '-sk', type=bool, default=False,
help="whether to save K matrix, 2-3x faster runtime, but larger memory O(n^2)")
parser.add_argument('--m', '-m', type=int, default=6,
help="number of thinning rounds")
parser.add_argument('--d', '-d', type=int, default=1,
help="dimensions")
parser.add_argument('--M', '-M', type=int, default=None,
help="number of mixture for diag mog in d=2")
parser.add_argument('--filename', '-f', type=str, default=None,
help="name for saved (MCMC) samples")
parser.add_argument('--combine_mmd', '-cm', type=bool, default=False,
help="whether to save combined_mmd results; should be set to True once all experiments are done running")
args, opt = parser.parse_known_args()
else:
args = None
# ## Define kernel thinning experiment
def run_kernel_thinning_experiment(m, params_p, params_k_split, params_k_swap, rep_ids,
delta=None, store_K=False,
sample_seed=1234567, thin_seed=9876543,
compute_mmds = True,
rerun=False,
verbose=False,
results_dir="results_new"):
"""Runs kernel thinning experiment using samples from params_p for repetitions over rep_ids,
saves coresets to disk, saves and returns mmd evaluations to disk mmd evaluation
Args:
m: Number of halving rounds (number of sample points n = 2^{2m})
params_p: Dictionary of distribution parameters recognized by sample()
params_k_split: Dictionary of kernel parameters recognized by kernel_eval()
params_k_swap: Dictionary of kernel parameters recognized by kernel_eval()
rep_ids: Which replication numbers of experiment to run; the replication
number determines the seeds set for reproducibility
delta: delta/(4^m) is the failure probability for
adaptive threshold sequence;
store_K: If False, runs O(nd) space version which does not store kernel
matrix; if True, stores n x n kernel matrix
sample_seed: (Optional) random seed is set to sample_seed + rep
prior to generating input sample for replication rep
thin_seed: (Optional) random seed is set to thin_seed + rep
prior to running thinning for replication rep
rerun: (Optional) If False and results have been previously saved to
disk, load results from disk instead of rerunning experiment
verbose: (Optinal) If True print time taken in each kt round
results_dir: (Optional) Directory in which results should be saved
compute_mmds: (Optional) Whether to compute mmds of coresets (using params_k_swap)
returns MMD evaluation of final thinned coresets from each rep using the
params_k_swap kernel and the params_p target distribution
"""
# Create results directory if necessary
pathlib.Path(results_dir).mkdir(parents=True, exist_ok=True)
split_kernel = partial(kernel_eval, params_k=params_k_split)
swap_kernel = partial(kernel_eval, params_k=params_k_swap)
# Construct results filename template with placeholder for rep value
d = params_p["d"]
assert(d==params_k_split["d"])
assert(d==params_k_swap["d"])
sample_str = sample_string(params_p, sample_seed)
split_kernel_str = "{}_var{:.3f}_seed{}".format(params_k_split["name"], params_k_split["var"], thin_seed)
swap_kernel_str = "{}_var{:.3f}".format(params_k_swap["name"], params_k_swap["var"])
thresh_str = f"delta{delta}"
file_template = os.path.join(results_dir, f"kt-coresets-{sample_str}-split{split_kernel_str}-swap{swap_kernel_str}-d{d}-m{m}-{thresh_str}-rep{{}}.pkl")
# Create array to store MMD evaluations from P, and Sin
if compute_mmds:
mmds_p = np.zeros((m+1, len(rep_ids)))
mmds_sin = np.zeros((m+1, len(rep_ids)))
mmd_p_file_template = os.path.join(results_dir,
f"kt-mmd-{sample_str}-split{split_kernel_str}-swap{swap_kernel_str}-d{d}-m{m}-{thresh_str}-rep{{}}.pkl")
mmd_sin_file_template = os.path.join(results_dir,
f"kt-mmd-sin-{sample_str}-split{split_kernel_str}-swap{swap_kernel_str}-d{d}-m{m}-{thresh_str}-rep{{}}.pkl")
split_kernel = partial(kernel_eval, params_k=params_k_split)
swap_kernel = partial(kernel_eval, params_k=params_k_swap)
# Number of sample points
n = int(2**(2*m))
fprint(f"Running kernel thinning experiment with template {file_template}.....")
tic()
for r_i, rep in enumerate(rep_ids):
# Include replication number in filename
filename = file_template.format(rep)
mmd_p_filename = mmd_p_file_template.format(rep)
mmd_sin_filename = mmd_sin_file_template.format(rep)
# Generate matrix of input sample points
#print(f"Generating data for rep {rep}", flush=True)
#tic()
X = sample(n, params_p, seed=sample_seed+rep)
#toc()
if not rerun and os.path.exists(filename):
# Return previously saved results
#print(f"Loading coresets from {filename}", flush=True)
#tic()
with open(filename, 'rb') as file:
coresets = pkl.load(file)
#toc()
else:
# Obtain sequence of thinned coresets
print(f"Kernel Thinning rep {rep}...", flush=True)
# tic()
coresets = kt.thin(X, m, split_kernel, swap_kernel, delta=delta, seed=thin_seed+rep, store_K=store_K, verbose=verbose)
# toc()
# Save coresets to disk
# print(f"Saving coresets to {filename}", flush=True)
# tic()
with open(filename, 'wb') as file:
pkl.dump(coresets, file, protocol=pkl.HIGHEST_PROTOCOL)
#toc()
# Evaluate final coreset MMD
if compute_mmds:
if not rerun and os.path.exists(mmd_p_filename):
# Return previously saved results
#print(f"Loading KT MMD results from {mmd_filename}", flush=True)
#tic()
with open(mmd_p_filename, 'rb') as file:
mmds_p[:, r_i] = pkl.load(file)
#toc()
else:
#print("Evaluating KT MMD", flush=True)
#tic()
for j in range(m+1):
nj = int(2**j)
mmds_p[j, r_i] = np.sqrt(
squared_mmd(params_k_swap, params_p, X[coresets[:nj]]))
#toc()
# Save MMD results to disk
# print(f"Saving KT MMD results to {mmd_filename}", flush=True)
#tic()
with open(mmd_p_filename, 'wb') as file:
pkl.dump(mmds_p[:, r_i], file, protocol=pkl.HIGHEST_PROTOCOL)
#toc()
if not rerun and os.path.exists(mmd_sin_filename):
# Return previously saved results
#print(f"Loading KT MMD results from {mmd_filename}", flush=True)
#tic()
with open(mmd_sin_filename, 'rb') as file:
mmds_sin[:, r_i] = pkl.load(file)
#toc()
else:
#print("Evaluating KT MMD", flush=True)
#tic()
# redefining target p as distribution on Sin
params_p_sin = dict()
params_p_sin["name"] = params_p["name"]+ "_sin"
params_p_sin["Pnmax"] = X
params_p_sin["d"] = d
for j in range(m+1):
nj = int(2**j)
mmds_sin[j, r_i] = np.sqrt(squared_mmd(params_k_swap, params_p_sin, X[coresets[:nj]]))
#toc()
# Save MMD results to disk
# print(f"Saving KT MMD results to {mmd_filename}", flush=True)
#tic()
with open(mmd_sin_filename, 'wb') as file:
pkl.dump(mmds_sin[:, r_i], file, protocol=pkl.HIGHEST_PROTOCOL)
#toc()
toc()
if compute_mmds:
return(mmds_p, mmds_sin)
# ## Define standard thinning experiment
def run_standard_thinning_experiment(m, params_p, params_k_mmd, rep_ids, sample_seed=1234567,
rerun=False, results_dir="results_new", compute_mmds=True,
min_mmd=False):
"""Evaluates MMD of iid Monte Carlo draws, and saves it to disk
Args:
m: Number of halving rounds (defines number of sample points via n = 2^{2m})
params_p: Dictionary of distribution parameters recognized by sample()
params_k_mmd: Dictionary of kernel parameters for MMD evaluation
rep_ids: Which replication numbers of experiment to run; the replication
number determines the seeds set for reproducibility
sample_seed: (Optional) random seed is set to sample_seed + rep
prior to generating input sample for replication rep
rerun: (Optional) If False and results have been previously saved to
disk, load results from disk instead of rerunning experiment
results_dir: (Optional) Directory in which results should be saved
min_mmd: (Optional) if True, returns the minimum MMD over all sqrt(n) thinned
sequences of n points with step size sqrt(n); if False, returns the MMD
of the first such thinned sequence
compute_mmds: (Optional) Whether to compute mmds of coresets (using params_k_mmd)
"""
# Create results directory if necessary
pathlib.Path(results_dir).mkdir(parents=True, exist_ok=True)
# Create array to store MMD evaluations
mmds_p = np.zeros((m+1, len(rep_ids)))
mmds_sin = np.zeros((m+1, len(rep_ids)))
# Construct results filename template with placeholder for rep value
d = params_p["d"]
assert(d == params_k_mmd["d"])
sample_str = sample_string(params_p, sample_seed)
kernel_str = "{}_var{:.3f}".format(params_k_mmd["name"], params_k_mmd["var"])
min_str = "min_" if min_mmd else ""
mmd_p_file_template = os.path.join(results_dir, f"{min_str}mc-mmd-{sample_str}-{kernel_str}-d{d}-m{m}-rep{{}}.pkl")
mmd_sin_file_template = os.path.join(results_dir, f"{min_str}mc-sin-mmd-{sample_str}-{kernel_str}-d{d}-m{m}-rep{{}}.pkl")
# Number of sample points
n = int(2**(2*m))
fprint(f"Running standard thinning experiment for m={m}")
tic()
for r_i, rep in enumerate(rep_ids):
# Include replication number in filename
mmd_p_filename = mmd_p_file_template.format(rep)
mmd_sin_filename = mmd_sin_file_template.format(rep)
if not rerun and os.path.exists(mmd_p_filename):
# Return previously saved results
#print(f"Loading {min_str} Monte Carlo MMD results from {filename}", flush=True)
#tic()
with open(mmd_p_filename, 'rb') as file:
mmds_p[:, r_i] = pkl.load(file)
#toc()
else:
#tic()
#print(f"Generating data for rep {rep}", flush=True)
#tic()
X = sample(n, params_p, seed=sample_seed+rep)
#toc()
#print(f"Evaluating {min_str} Monte Carlo MMD", flush=True)
for j in range(m+1):
# Target coreset size
coreset_size = int(2**j)
input_size = int(coreset_size**2)
if min_mmd:
# Consider each coreset obtained by choosing every nj-th point
# of the first nj^2 points of X and select the one with smallest MMD
# There are nj^2/nj = nj such coresets indexed by their starting point
num_starts = coreset_size
else:
num_starts = 1
step_size = coreset_size
end = input_size
mmds_p[j, r_i] = np.inf
for start in range(num_starts):
mmds_p[j, r_i] = min(mmds_p[j, r_i], np.sqrt(squared_mmd(params_k_mmd, params_p, X[(step_size-1-start):end:step_size])))
#print(f"j={j},rep={rep},mmd={mmds[j,rep]}")
#toc()
# Save MMD results to disk
#print(f"Saving {min_str} Monte Carlo MMD results to {filename}", flush=True)
#tic()
with open(mmd_p_filename, 'wb') as file:
pkl.dump(mmds_p[:, r_i], file, protocol=pkl.HIGHEST_PROTOCOL)
#toc()
if not rerun and os.path.exists(mmd_sin_filename):
# Return previously saved results
#print(f"Loading {min_str} Monte Carlo MMD results from {filename}", flush=True)
#tic()
with open(mmd_sin_filename, 'rb') as file:
mmds_sin[:, r_i] = pkl.load(file)
#toc()
else:
#tic()
#print(f"Generating data for rep {rep}", flush=True)
#tic()
X = sample(n, params_p, seed=sample_seed+rep)
#toc()
#print(f"Evaluating {min_str} Monte Carlo MMD", flush=True)
# redefining target p as distribution on Sin
params_p_sin = dict()
params_p_sin["name"] = params_p["name"]+"_sin"
params_p_sin["Pnmax"] = X
params_p_sin["d"] = d
for j in range(m+1):
# Target coreset size
coreset_size = int(2**j)
input_size = int(coreset_size**2)
if min_mmd:
# Consider each coreset obtained by choosing every nj-th point
# of the first nj^2 points of X and select the one with smallest MMD
# There are nj^2/nj = nj such coresets indexed by their starting point
num_starts = coreset_size
else:
num_starts = 1
step_size = coreset_size
end = input_size
mmds_sin[j, r_i] = np.inf
for start in range(num_starts):
mmds_sin[j, r_i] = min(mmds_sin[j, r_i], np.sqrt(
squared_mmd(params_k_mmd, params_p_sin, X[(step_size-1-start):end:step_size])))
#print(f"j={j},rep={rep},mmd={mmds[j,rep]}")
#toc()
# Save MMD results to disk
#print(f"Saving {min_str} Monte Carlo MMD results to {filename}", flush=True)
#tic()
with open(mmd_sin_filename, 'wb') as file:
pkl.dump(mmds_sin[:, r_i], file, protocol=pkl.HIGHEST_PROTOCOL)
#toc()
toc()
return(mmds_p, mmds_sin)
# # Deploy thinning experiments
# +
#
# Choose sample and kernel parameters
#
var = 1. # Variance
d = int(2) if args is None else args.d
params_p = {"name": "gauss", "var": var, "d": int(d), "saved_samples": False}
# filename is for MCMC files
filename = None if args is None else args.filename
# k denotes the number of componets for MOG settings
M = None if args is None else args.M
if isnotebook():
filename = None if args is None else args.filename
# ['Goodwin_RW', 'Goodwin_ADA-RW', 'Goodwin_MALA', 'Goodwin_PRECOND-MALA', 'Lotka_RW', 'Lotka_ADA-RW', 'Lotka_MALA', 'Lotka_PRECOND-MALA']
if filename is not None:
# if a filename is specified then compute params_p
d = int(4)
params_p = compute_mcmc_params_p(filename, nmax=int(2**15), include_last=True)
# whether to use median_distance for kernel bandwidth for MCMC settings
use_median_distance = True
if use_median_distance:
var = (params_p["med_dist"])**2
if M is not None:
# if number of mixture is specified then compute params_p
d = int(2)
params_p = compute_diag_mog_params(M)
params_k_swap = {"name": "gauss", "var": var, "d": int(d)}
params_k_split = {"name": "gauss_rt", "var": var/2., "d": int(d)}
# +
#
# Choose experiment parameters
#
# List of replicate ID numbers
rep_ids = range(2) if args is None else np.arange(args.rep0, args.rep0+args.repn)
# List of halving round numbers m to evaluate
ms = range(5+1) if args is None else range(args.m)
# whether store_k during thinning, saves computation but requires O(n^2) memory
# issue with larger n; if False, requires O(nd) memory
store_K = False if args is None else args.store_K
# Failure probability
delta = .5
# Which methods to run?
run_standard_thinning = False
run_kernel_thinning = True
rerun = True
verbose = False # time updates only for m>=7
# +
if run_standard_thinning:
mmds_st = np.zeros((max(ms)+1, len(rep_ids))) # mmds from P
mmds_st_sin = np.zeros((max(ms)+1, len(rep_ids))) # mmds from Sin
if run_kernel_thinning:
mmds_kt = np.zeros((max(ms)+1, len(rep_ids))) # mmds from P
mmds_kt_sin = np.zeros((max(ms)+1, len(rep_ids))) # mmds from Sin
print("Exp setting:", params_p["name"], ms)
for m in ms:
#
# Run experiments and store quality of the 2^m thinned coreset
#
if run_standard_thinning:
mmd_st, mmd_st_sin = run_standard_thinning_experiment(m, params_p, params_k_swap, rep_ids, rerun=rerun)
mmds_st[m, :] = mmd_st[m, :]
mmds_st_sin[m, :] = mmd_st_sin[m, :]
if run_kernel_thinning:
mmd_kt, mmd_kt_sin = run_kernel_thinning_experiment(m, params_p, params_k_split, params_k_swap, rep_ids, delta, store_K, rerun=rerun, verbose=verbose)
mmds_kt[m, :] = mmd_kt[m, :]
mmds_kt_sin[m, :] = mmd_kt_sin[m, :]
# -
# # Save MMD Results
# +
#
# Save all mmd settings
#
save_combined_mmd = False if args is None else args.combine_mmd
if save_combined_mmd:
if run_standard_thinning:
filename = get_combined_mmd_filename(f"mc", ms, params_p, params_k_split, params_k_swap, rep_ids, delta)
with open(filename, 'wb') as file:
print(f"Saving combined mc mmd to {filename}")
pkl.dump(mmds_st, file, protocol=pkl.HIGHEST_PROTOCOL)
filename = get_combined_mmd_filename(f"mc-sin", ms, params_p, params_k_split, params_k_swap, rep_ids, delta)
with open(filename, 'wb') as file:
print(f"Saving combined mc mmd_sin to {filename}")
pkl.dump(mmds_st_sin, file, protocol=pkl.HIGHEST_PROTOCOL)
if run_kernel_thinning:
filename = get_combined_mmd_filename(f"kt", ms, params_p, params_k_split, params_k_swap, rep_ids, delta)
with open(filename, 'wb') as file:
print(f"Saving combined kt mmd to {filename}")
pkl.dump(mmds_kt, file, protocol=pkl.HIGHEST_PROTOCOL)
filename = get_combined_mmd_filename(f"kt-sin", ms, params_p, params_k_split, params_k_swap, rep_ids, delta)
with open(filename, 'wb') as file:
print(f"Saving combined kt mmd_sin to {filename}")
pkl.dump(mmds_kt_sin, file, protocol=pkl.HIGHEST_PROTOCOL)
| examples/kt/run_kt_experiment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# # Title (subtopic)
#
# - Short description
# - Relate to presentation
# - Requirements for running the notebook if necessary
#
# ## Installation of libraries and necessary software
# - Code to install libraries
# - State that this has to be run only once
# - If necessary, further installation instructions
print("Code for installation of libraries")
# ## Loading data
# Examples:
# - Load required data
# - Instructions to download files
# - Generate randomized data sets
print("Code to load and/or create data")
# ### Exercise 1
# E.g.: Create a data set containing the numbers 1 to 100 (`dset <- 1:100`) and calculate _mean, median, minium, maximum, variance_ and _standard deviation_.
#
#
#
print("Working example code to run and alter")
# #### Add your answers here
# (double-click here to edit the cell)
#
# ##### Question I: <ins>Why are _mean_ and _median_ the same?</ins>
#
# _Answer_
#
# ##### Question II: <ins>What is the relationship between _variance_ and _standard deviation_?</ins>
#
# _Answer_
#
#
#
print("PUT YOUR CODE HERE")
print("Code done by students to solve tasks")
# ### Exercise 2
# blabla
#
#
print("Working example code to run and alter")
# #### Add your answers here
# (double-click here to edit the cell)
#
# ##### Question I: <ins>Why ...?</ins>
#
# _Answer_
#
# ##### Question II: <ins>What ..._?</ins>
#
# _Answer_
#
#
#
| Template-Topic/template_subtopic1/ExercisesA_subtopic1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Tutorial - A Basic Encoding Script
#
# In this tutorial, you will learn how to create an encoding from scratch, using the Bitmovin APIs and the Python SDK that wraps them. We will explain the concepts and the terminology that we use.
#
# This tutorial concentrates on taking a single source file, encoding it into a ladder of multiple renditions, and creating a manifest which can be played back by any modern video player on most devices and browsers.
# + [markdown] colab_type="text" id="GAS_K2t7O-Gl" slideshow={"slide_type": "slide"}
# # Understanding how the API is composed
# 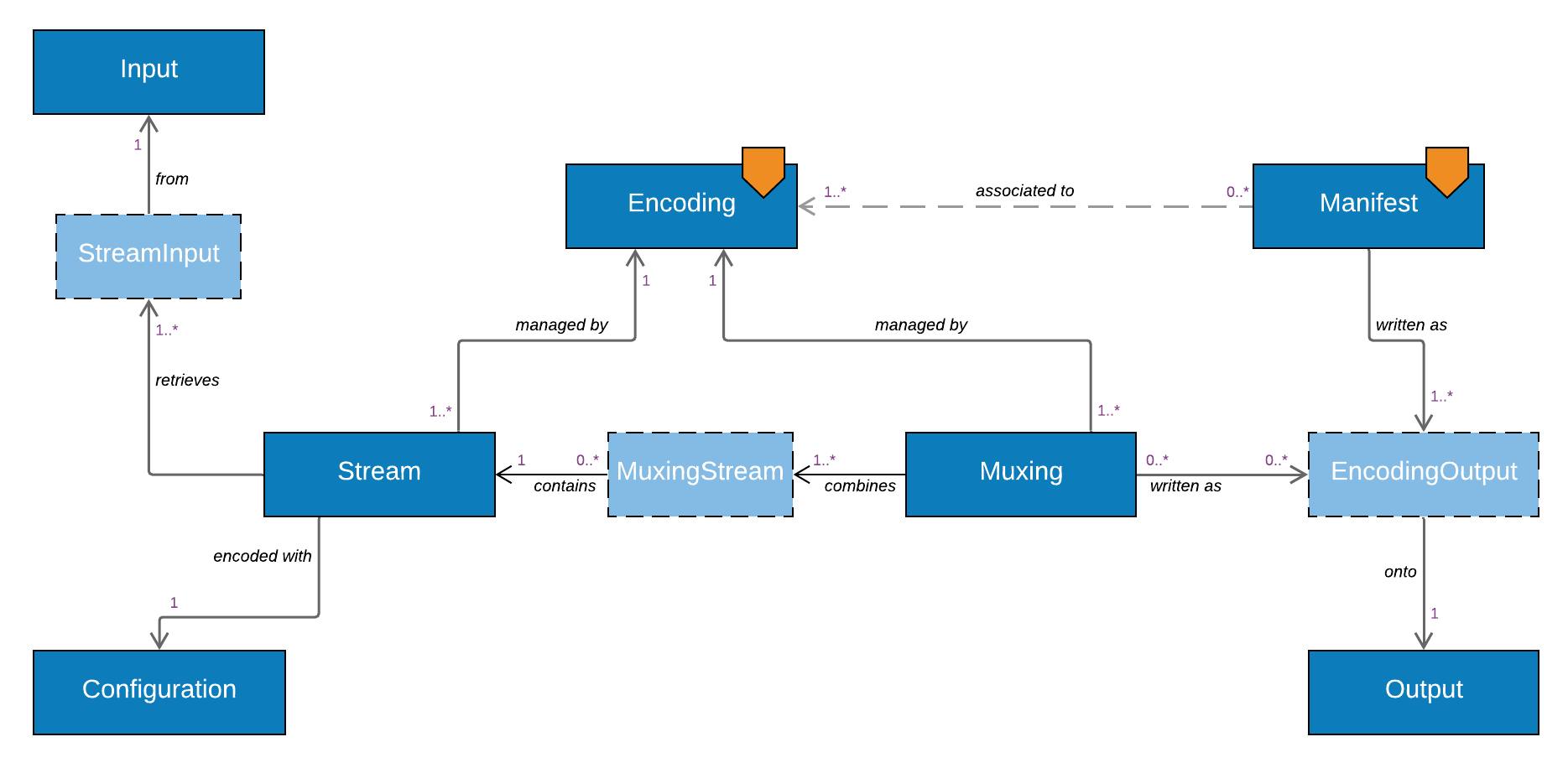
#
# For a complete description of the Bitmovin data model, check our [Object Model documentation]( https://bitmovin.com/docs/encoding/tutorials/understanding-the-bitmovin-encoding-object-model).
# + [markdown] colab_type="text" ein.tags="worksheet-0" id="qI4_K7g8bwFF" slideshow={"slide_type": "slide"}
# # A Little Setup
# Let's import the Bitmovin API Client into our python script, as well as other dependencies we will need.
# + colab={} colab_type="code" ein.hycell=false ein.tags="worksheet-0" id="a5L_4anwbwFI"
import sys
sys.path.append('../libs')
import collections
import os
import time
import uuid
from bitmovin_api_sdk import BitmovinApi, BitmovinApiLogger
from bitmovin_api_sdk import HttpsInput, S3Output, AwsCloudRegion
from bitmovin_api_sdk import StreamInput, StreamSelectionMode
from bitmovin_api_sdk import ProfileH264
from bitmovin_api_sdk import H264VideoConfiguration, PresetConfiguration
from bitmovin_api_sdk import Encoding, CloudRegion
from bitmovin_api_sdk import MuxingStream, Fmp4Muxing, EncodingOutput, AclEntry, AclPermission
from bitmovin_api_sdk import Stream, Status
from bitmovin_api_sdk import DashManifestDefault, DashManifestDefaultVersion
from bitmovin_api_sdk import AacAudioConfiguration
from dotenv import load_dotenv
_ = load_dotenv()
# + [markdown] slideshow={"slide_type": "subslide"}
# For the purpose of this tutorial, we also import a few additional helpers. You won't need these in your own scripts
# + colab={} colab_type="code" ein.hycell=false ein.tags="worksheet-0" id="a5L_4anwbwFI"
import config as cfg
import helpers
from IPython.display import display, IFrame
from vdom import p, div, b, a
# -
# We are being quite specific about the Bitmovin objects that we want to import, just to make things clear for this example...
# + [markdown] colab_type="text" ein.tags="worksheet-0" id="JABsdDe4bwFN" slideshow={"slide_type": "subslide"}
# ## Secret sauce
# The API key is what you need to authenticate with the Bitmovin API. You can find it in the dashboard in the [Account section](https://bitmovin.com/dashboard/account)
#
# It is a __secret__, and should be treated as such. If someone else gets hold of your key, they can run encodings on your account (or your organisation accounts) and get information about previous ones.
# + colab={} colab_type="code" ein.hycell=false ein.tags="worksheet-0" id="rsOtBeuobwFQ"
cfg.API_KEY = os.getenv('API_KEY', "")
# -
# The Organization ID indicates what Bitmovin account you want to create and process your encodings in. Leave it empty if you are using your own account. If you belong to a multi-tenant organization, you need to get the organisation ID from the dashboard in the [Organization section](https://bitmovin.com/dashboard/organization/overview).
# + colab={} colab_type="code" ein.hycell=false ein.tags="worksheet-0" id="rsOtBeuobwFQ"
cfg.ORG_ID = os.getenv('ORG_ID', "")
# + [markdown] slideshow={"slide_type": "subslide"}
# For this learning lab we have created an S3 bucket and a (very limited) user we can all use.
# + colab={} colab_type="code" ein.hycell=false ein.tags="worksheet-0" id="rsOtBeuobwFQ"
cfg.S3_BUCKET_NAME = os.getenv('S3_BUCKET_NAME', "")
cfg.S3_ACCESS_KEY = os.getenv('S3_ACCESS_KEY', "")
cfg.S3_SECRET_KEY = os.getenv('S3_SECRET_KEY', "")
# -
# Finally, to prevent conflicts between all our encodings, let's add something unique to each of us.
# + colab={} colab_type="code" ein.hycell=false ein.tags="worksheet-0" id="rsOtBeuobwFQ"
cfg.MY_ID = os.getenv('MY_ID', "")
# + [markdown] slideshow={"slide_type": "subslide"}
# We'll quickly run some checks to make sure that your setup is ready to use. This is where we use these helpers.
# + colab={} colab_type="code" ein.hycell=false ein.tags="worksheet-0" id="rsOtBeuobwFQ"
msg = helpers.validate_config()
base_output_path = helpers.build_output_path()
display(
div(
p(f"{msg}. Your output files will be added to subdirectory ", b(f"{base_output_path}")),
style={"color": "green"}
)
)
# -
# In a production script, you won't need this (or rather, you'll need to do something that is suitable for your workflow)
# + [markdown] colab_type="text" ein.tags="worksheet-0" id="bEYpTWuebwFN" slideshow={"slide_type": "slide"}
# # Configuring our Encoding
# Now that the boring bits are behind us, we are (finally) ready to start the real work.
#
# First, we need to instantiate the API client with our secrets.
# + colab={} colab_type="code" ein.hycell=false ein.tags="worksheet-0" id="Ia50HGtDbwFS"
api = BitmovinApi(api_key=cfg.API_KEY,
tenant_org_id=cfg.ORG_ID)
# + [markdown] colab_type="text" ein.tags="worksheet-0" id="aOvDExJnbwFW" slideshow={"slide_type": "slide"}
# <img src="img/step1.svg" alt="Input and Output" width="320px" align="right"/>
#
# ## Input and Output locations
# Every encoding needs at least one input and output. In Bitmovin parlance an `Input` is an input storage location with a specific transport protocol, for example an HTTPS location. It is _not_ a specific file. Our documentation provides a full list of [supported Inputs](https://bitmovin.com/docs/encoding/articles/supported-input-output-storages).
# + colab={} colab_type="code" ein.hycell=false ein.tags="worksheet-0" id="CN1mU4UIbwFY"
input = HttpsInput(name=f'{cfg.MY_ID}_LearningLab_Sources',
description='Web server for Bitmovin Dev Lab inputs',
host="bitmovin-learning-labs-london.s3.amazonaws.com")
input = api.encoding.inputs.https.create(https_input=input)
print("Created input '{}' with id: {}".format(input.name, input.id))
# -
# Note how we first create a resource in the SDK, and then submit it to the API for creation. The API will return a full representation of the object, and generated an ID for it. We will use those identifiers to link the various objects that make up the full configuration.
# + [markdown] colab_type="text" id="AVLfJoJe_mp-" slideshow={"slide_type": "subslide"}
# The same concepts apply to the `Output`, which defines where we will store the resulting files.
# + colab={} colab_type="code" id="_9e_03we_iIu"
output = S3Output(name=f'{cfg.MY_ID}_{cfg.S3_BUCKET_NAME}',
description='Bucket for Bitmovin Dev Lab outputs',
bucket_name=cfg.S3_BUCKET_NAME,
access_key=cfg.S3_ACCESS_KEY,
secret_key=cfg.S3_SECRET_KEY)
output = api.encoding.outputs.s3.create(s3_output=output)
print("Created output '{}' with id: {}".format(output.name, output.id))
# -
# _Note_: It is best practice to _reuse_ inputs and outputs you have created before, not create a new one every time.
#
# You can and should query the API to retrieve the resources you previously created inputs, by their name. The `name` and `description` properties can be added to all Bitmovin resources.
# + [markdown] colab_type="text" ein.tags="worksheet-0" id="7EwLTNPbbwFg" slideshow={"slide_type": "slide"}
# <img src="img/step2.svg" alt="StreamInput" width="320px" align="right"/>
#
# ## Mapping input media streams
# We can now define what source file to use in the encoding.
# To do so, we need to create a `StreamInput` resource, which specifies on what _input_ our file is located (by using its ID), at what _path_. We also define what media track to select to decode (and later encode).
#
# The first input stream we specify is for the video track:
# + colab={} colab_type="code" ein.hycell=false ein.tags="worksheet-0" id="SGZB3N7nbwFh"
video_input_stream = StreamInput(input_id=input.id,
input_path="input-files/cosmos_laundromat.mp4",
selection_mode=StreamSelectionMode.AUTO)
# + [markdown] colab_type="text" ein.tags="worksheet-0" id="JYQcF_g_bwFl" slideshow={"slide_type": "subslide"}
# The next one we will create is for the audio. We've specified `StreamSelectionMode.AUDIO_RELATIVE` here and `position=0` to indicate I want the first (0th) audio track in numerical order.
# + colab={} colab_type="code" ein.hycell=false ein.tags="worksheet-0" id="7fMdtxjzbwFm"
audio_input_stream = StreamInput(input_id=input.id,
input_path='input-files/cosmos_laundromat.mp4',
selection_mode=StreamSelectionMode.AUDIO_RELATIVE,
position=0)
# + [markdown] colab_type="text" ein.tags="worksheet-0" id="fxVXZWFxbwFq"
# Note that the `StreamInput` is not a resource that is submitted to the API directly. It is an internal object used within the SDK, which is used later in the definition of other resources.
# + [markdown] colab_type="text" ein.tags="worksheet-0" id="Q_dldXfbbwFr" slideshow={"slide_type": "slide"}
# <img src="img/step3.svg" alt="Configuration" width="320px" align="right"/>
#
# ## Configuring the codecs
#
# Next we need to create the codec configurations that define how those files get encoded into the output streams.
#
# We use a helper tuple (a Python-esque construct) to group up our desired output height, bitrate, and video profile.
# + colab={} colab_type="code" ein.hycell=false ein.tags="worksheet-0" id="gzXbDCHFbwFs"
MyProfile=collections.namedtuple('MyProfile', 'height bitrate profile')
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Ladder
#
# We then define a "ladder" as a set of encoding configurations for the encoder to generate. We will be using H264/AVC in this example.
# + colab={} colab_type="code" ein.hycell=false ein.tags="worksheet-0" id="gzXbDCHFbwFs"
video_profiles = [
MyProfile(height=240, bitrate=400_000, profile=ProfileH264.MAIN),
MyProfile(height=360, bitrate=800_000, profile=ProfileH264.HIGH),
MyProfile(height=480, bitrate=1_200_000, profile=ProfileH264.HIGH),
MyProfile(height=720, bitrate=2_400_000, profile=ProfileH264.HIGH),
MyProfile(height=1080, bitrate=4_800_000, profile=ProfileH264.HIGH),
]
# + [markdown] colab_type="text" ein.tags="worksheet-0" id="RZpJJB0kbwFw" slideshow={"slide_type": "subslide"}
# ### Video
#
# We can now create each of these video profiles. We will use one of the [preset configurations](https://bitmovin.com/docs/encoding/tutorials/h264-presets), which are templates defined for most common use cases, whether your focus is on performance or quality, and they should always be used.
# + colab={} colab_type="code" ein.hycell=false ein.tags="worksheet-0" id="9QaSJ2e0bwFy"
video_configs = []
for profile in video_profiles:
video_config = H264VideoConfiguration(
name=f"{cfg.MY_ID}_H264-{profile.height}p@{profile.<EMAIL>rate}",
height=profile.height,
bitrate=profile.bitrate,
profile=profile.profile,
preset_configuration=PresetConfiguration.VOD_STANDARD
)
video_config = api.encoding.configurations.video.h264.create(video_config)
video_configs.append(video_config)
print("Created video codec config '{}' with id: {}".format(video_config.name, video_config.id))
# -
# Note that just like inputs and outputs, these resources can and should also be re-used. You can also create them in the dashboard if desired.
# + [markdown] colab_type="text" ein.tags="worksheet-0" id="nbHDVU1xbwF4" slideshow={"slide_type": "subslide"}
# ### Adaptive bitrate video
#
# Let's pause for a second and cover _why_ we are generating multiple profiles here.
#
# We are encoding our source video in such a way that it can be played back in a player that supports Adaptive Bitrate (ABR) Streaming. With this mechanism, the Video Player can choose which representation (often called rendition) to play, based on its available bandwidth and capabilities, and can also switch between them dynamically, going to a higher bitrate (and therefore better quality) as the available bandwidth increases, or going to lower bitrates (and lower qualities) as the network conditions deteriorate.
#
# Each of these representations is a separate encode of the source files, and thus requires a distinct configuration of the encoder.
# + [markdown] colab_type="text" ein.tags="worksheet-0" id="T006y4X1bwF6" slideshow={"slide_type": "subslide"}
# ### Audio
# We also need to create the audio configuration. A single AAC stream will do for now.
# + colab={} colab_type="code" ein.hycell=false ein.tags="worksheet-0" id="sJCH8XmCbwF7"
audio_config = AacAudioConfiguration(
name=f"{cfg.MY_ID}_AAC-128k",
bitrate=128_000,
rate=48_000.0)
audio_config = api.encoding.configurations.audio.aac.create(aac_audio_configuration=audio_config)
print("Created audio codec config '{}' with id: {}".format(audio_config.name, audio_config.id))
# + [markdown] colab_type="text" id="kif4n-VExk8v" slideshow={"slide_type": "slide"}
# ## The Encoding itself
#
# <img src="img/step4.svg" alt="Encoding" width="320px" align="right"/>
#
# Each encoding job will have a resource that defines it. We define a number of aspects of the encoding through it:
# - The `CloudRegion` defines through what cloud provider and in which region to perform the encoding. We are setting it to `AUTO` here, which means that Bitmovin will attempt to make a "sensible" choice about where to run the encoding. It's best to use a specific region however.
#
# - The `EncoderVersion`: You should set it to `STABLE` to ensure you get the most up to date version of the encoder, for best performance and reliability
# + colab={} colab_type="code" ein.hycell=false ein.tags="worksheet-0" id="KrRz_lEibwGA" slideshow={"slide_type": "subslide"}
encoding = Encoding(name=f"{cfg.MY_ID} - basic encoding tutorial",
encoder_version="STABLE",
cloud_region=CloudRegion.AUTO)
encoding = api.encoding.encodings.create(encoding=encoding)
print("Created encoding '{}' with id: {}".format(encoding.name, encoding.id))
# + [markdown] colab_type="text" ein.tags="worksheet-0" id="IJHCBK8cbwGG" slideshow={"slide_type": "slide"}
# ## Mapping inputs to outputs
#
# <img src="img/step4b.svg" alt="Intermediary Summary" width="320px" align="right"/>
#
# So far we have created:
# * An input
# * An output
# * A set of video and audio "profiles"
# * An empty encoding object
#
# Having all these "non-dependent" objects ready, it's now time to connect the chain that will tie in input and output.
# + [markdown] colab_type="text" id="MBwiR1VozvNy" slideshow={"slide_type": "slide"}
# ### Streams
#
# <img src="img/step5.svg" alt="Streams" width="320px" align="right"/>
#
# We will first create a series of output streams. These simply map one or multiple _input_ streams to a single (elementary) _output_ stream, and are the raw output of the encoding process itself.
#
# For our ABR tutorial use case, there is a simple one-to-one relationship between codecs and video streams.
# + [markdown] slideshow={"slide_type": "subslide"}
# So, for each config, we create a corresponding `Stream`, which we link to the `StreamInput` created earlier. We link the Stream to a `Configuration`, and attach it to our `Encoding`.
# + colab={} colab_type="code" ein.hycell=false ein.tags="worksheet-0" id="CbC2W9A7bwGI" slideshow={"slide_type": "-"}
video_streams = []
for config in video_configs:
stream_shortname = '{}p_{}k'.format(config.height, round(config.bitrate/1000))
video_stream = Stream(name=f"{cfg.MY_ID}_{stream_shortname}",
description=stream_shortname,
codec_config_id=config.id,
input_streams=[video_input_stream])
video_stream = api.encoding.encodings.streams.create(encoding.id, stream=video_stream)
video_streams.append(video_stream)
print("Created video stream '{}' with id: {}".format(video_stream.name, video_stream.id))
# + [markdown] slideshow={"slide_type": "subslide"}
# And then we do the same thing for the audio
# + colab={} colab_type="code" ein.hycell=false ein.tags="worksheet-0" id="CbC2W9A7bwGI" slideshow={"slide_type": "-"}
audio_stream = Stream(name=f'{cfg.MY_ID}_AAC',
codec_config_id=audio_config.id,
input_streams=[audio_input_stream])
audio_stream = api.encoding.encodings.streams.create(encoding.id, stream=audio_stream)
print("Created audio stream '{}' with id: {}".format(audio_stream.name, audio_stream.id))
# + [markdown] colab_type="text" ein.tags="worksheet-0" id="zhh-YrldbwGS" slideshow={"slide_type": "slide"}
# ### Muxing
#
# <img src="img/step6.svg" alt="Muxings" width="320px" align="right"/>
#
# Raw output isn't enough however. An output stream must be _muxed_ into a container, for example an MPEG Transport Stream, or fragmented MPEG 4 boxes (ISOBMFF).
#
# For each item we need to add our stream to a `MuxingStream`, which takes a stream, an `EncodingOutput` which specifies the output and the _path_ for this muxing and then create the `Muxing` itself.
#
# Note that muxings may contain multiple streams, and be replicated to multiple outputs. For simplicity's sake here, and in line with standard ABR practices, we create a separate muxing for each generated track.
# + colab={} colab_type="code" ein.hycell=false ein.tags="worksheet-0" id="W_rG1d4QbwGT" slideshow={"slide_type": "subslide"}
video_muxings = []
for video_stream in video_streams:
muxing_stream = MuxingStream(stream_id=video_stream.id)
muxing_output = EncodingOutput(output_id=output.id,
output_path='{}/video/{}'.format(base_output_path, video_stream.description),
acl=[AclEntry(permission=AclPermission.PUBLIC_READ)])
video_muxing = Fmp4Muxing(name=video_stream.name + "_fmp4",
streams=[muxing_stream],
segment_length=4.0,
segment_naming="seg_%number%.m4s",
init_segment_name='init.mp4',
outputs=[muxing_output])
video_muxing = api.encoding.encodings.muxings.fmp4.create(encoding_id=encoding.id, fmp4_muxing=video_muxing)
video_muxings.append(video_muxing)
print("Created video muxing '{}' with id: {}".format(video_muxing.name, video_muxing.id))
# + [markdown] colab_type="text" ein.tags="worksheet-0" id="L1boUHUnbwGd" slideshow={"slide_type": "subslide"}
# We also create a separate audio muxing, into a separate folder. We could mux the audio with our video streams, but standard practice with ABR is to have separate audio-only muxings.
#
# + colab={} colab_type="code" id="5yEVXssZ1Erz"
audio_muxing_stream = MuxingStream(stream_id=audio_stream.id)
audio_muxing_output = EncodingOutput(output_id=output.id,
output_path=base_output_path+'/audio/',
acl=[AclEntry(scope='*', permission=AclPermission.PUBLIC_READ)])
audio_muxing = Fmp4Muxing(name=f"{audio_stream.name}_fmp4",
streams=[audio_muxing_stream],
segment_length=4.0,
segment_naming="seg_%number%.m4s",
init_segment_name='init.mp4',
outputs=[audio_muxing_output])
audio_muxing = api.encoding.encodings.muxings.fmp4.create(encoding_id=encoding.id, fmp4_muxing=audio_muxing)
print("Created audio muxing '{}' with id: {}".format(audio_muxing.name, audio_muxing.id))
# + [markdown] colab_type="text" ein.tags="worksheet-0" id="XKj5Qq1ObwGg" slideshow={"slide_type": "slide"}
# <img src="img/step7.svg" alt="Start" width="320px" align="right"/>
#
# ## Starting the encoding...
#
# Next we are going to start the encode!
# + colab={} colab_type="code" ein.hycell=false ein.tags="worksheet-0" id="FkiX8-1abwGi"
api.encoding.encodings.start(encoding.id)
print("Starting encoding")
# + colab={} colab_type="code" ein.hycell=false ein.tags="worksheet-0" id="FkiX8-1abwGi"
url = helpers.build_dashboard_url(encoding.id)
display(
p("You can now check encoding progress in the dashboard at ",
a(f"{url}", href=f"{url}", target="_new")
)
)
# + [markdown] colab_type="text" id="tSJ5z8fsArjZ" slideshow={"slide_type": "subslide"}
# <img src="img/step8.svg" alt="Monitoring" width="320px" align="right"/>
#
# ### ... and monitoring it
#
# You can monitor the encoding in your script by polling its status on a regular basis. This is the easiest way to keep track of the encoding when you are testing your encoding configuration.
#
# For production environments however, you should use [webhooks](//https://bitmovin.com/docs/encoding/api-reference/sections/notifications-webhooks) instead.
# + colab={} colab_type="code" id="yGno_QyOPs-X"
while True:
task = api.encoding.encodings.status(encoding.id)
print("Got task status {} - {}%".format(task.status, task.progress))
if task.status == Status.ERROR:
print("Error during encoding!")
raise SystemExit
if task.status == Status.FINISHED:
print("Encoding complete")
break
time.sleep(15)
# + [markdown] colab_type="text" ein.tags="worksheet-0" id="x9FiZEwhbwGm" slideshow={"slide_type": "slide"}
# ## Combining into a manifest
#
# <img src="img/step9.svg" alt="Manifest" width="320px" align="right"/>
#
# We will ask the encoder to generate the `Manifest` as well. The manifest is used by ABR players to find all information about quality levels, audio tracks, subtitles etc.
#
# We are going to generate a _DASH_ (Dynamic Adaptive Streaming over HTTP) manifest, which can be played on Android and iOS devices, as well as the Bitmovin player on most platforms.
#
# Bitmovin provides you with full flexibility to create manifests in a fine grained way. But we will be using `DefaultManifest` functionality, which will apply smart defaults to create a standard manifest.
# + colab={} colab_type="code" ein.hycell=false ein.tags="worksheet-0" id="ODmU9M8ubwGo" slideshow={"slide_type": "subslide"}
print("Creating manifests")
manifest_output = EncodingOutput(output_id=output.id,
output_path=base_output_path+'/',
acl=[AclEntry(scope='*', permission=AclPermission.PUBLIC_READ)])
dash_manifest = DashManifestDefault(
name=f"{cfg.MY_ID}_DashManifest",
manifest_name="stream.mpd",
encoding_id=encoding.id,
version=DashManifestDefaultVersion.V1,
outputs=[manifest_output])
dash_manifest = api.encoding.manifests.dash.default.create(dash_manifest)
print("Created manifest '{}' with id: {}".format(dash_manifest.name, dash_manifest.id))
# + [markdown] colab_type="text" id="SwdEfdOQBbNH" slideshow={"slide_type": "subslide"}
# And now we can trigger the generation of the manifest...
# + colab={} colab_type="code" id="Ymo3a4wzBXwN"
api.encoding.manifests.dash.start(dash_manifest.id)
print("Generating manifest")
# + [markdown] slideshow={"slide_type": "subslide"}
# ... and monitor it
# + colab={} colab_type="code" id="Ymo3a4wzBXwN"
while True:
time.sleep(5)
status = api.encoding.manifests.dash.status(dash_manifest.id).status
if status == Status.FINISHED:
break
if status == Status.ERROR:
print("Error during dash manifest generation")
raise SystemExit
manifest_url = "https://"+cfg.S3_BUCKET_NAME+".s3.amazonaws.com/" + manifest_output.output_path + dash_manifest.manifest_name
display( p(f"Manifest URL: ", b(f"{manifest_url}")) )
# + [markdown] colab_type="text" id="Yn1wYl-QNCsk" slideshow={"slide_type": "slide"}
# # Playback test
# You can now try playing back the stream, by using the manifest URL in our test player at https://bitmovin.com/demos/stream-test
# + [markdown] colab_type="text" id="LhKZhLfBSLav" slideshow={"slide_type": "subslide"}
# ## In your own player
# Alternatively, you can just play it right here.
# You may have to whitelist the "google.com" domain for your player license first.
#
# To retrieve your license and add the domain, head to the Dashboard at https://bitmovin.com/dashboard/player/licenses
#
# + colab={} colab_type="code" id="r7TDrikdS6qF"
cfg.PLAYER_LICENSE='f9e2cf25-9cdd-4c9d-a314-90fdb6d5590c'
# + colab={} colab_type="code" id="r7TDrikdS6qF" slideshow={"slide_type": "subslide"}
embed_url = "https://demo.bitmovin.com/public/learning-labs/encoding/test-players/basic-dash-player.html?"
embed_url += "license="+cfg.PLAYER_LICENSE
embed_url += "&mpdurl="+manifest_url
IFrame(src=embed_url, width=800, height=450)
| notebooks/basic_encoding/basic-encoding.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Test that the output below shows up when `jupyter-book run` is used.
a = 'foo'
print('a is {}'.format(a))
| jupyter_book/tests/site/content/tests/run.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/wileyw/DeepLearningDemos/blob/master/sound/simple_audio_new_spectrogram_numpy_and_normalize.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="fluF3_oOgkWF"
# ##### Copyright 2020 The TensorFlow Authors.
# + cellView="form" id="AJs7HHFmg1M9"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="jYysdyb-CaWM"
# # Simple audio recognition: Recognizing keywords
# + [markdown] id="CNbqmZy0gbyE"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/tutorials/audio/simple_audio">
# <img src="https://www.tensorflow.org/images/tf_logo_32px.png" />
# View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/tutorials/audio/simple_audio.ipynb">
# <img src="https://www.tensorflow.org/images/colab_logo_32px.png" />
# Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/tutorials/audio/simple_audio.ipynb">
# <img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />
# View source on GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/tutorials/audio/simple_audio.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
# </td>
# </table>
# + [markdown] id="SPfDNFlb66XF"
# This tutorial will show you how to build a basic speech recognition network that recognizes ten different words. It's important to know that real speech and audio recognition systems are much more complex, but like MNIST for images, it should give you a basic understanding of the techniques involved. Once you've completed this tutorial, you'll have a model that tries to classify a one second audio clip as "down", "go", "left", "no", "right", "stop", "up" and "yes".
# + id="6tqplwVdQ__L"
# !git clone https://github.com/google-coral/project-keyword-spotter.git
# + id="8F5jI3URRH38"
# !ls project-keyword-spotter/
# + id="1AJr9ZolRL0W"
# !cp project-keyword-spotter/mel_features.py .
# + id="ioSqyACxRdkQ"
# !ls
import mel_features
# + [markdown] id="Go9C3uLL8Izc"
# ## Setup
#
# Import necessary modules and dependencies.
# + id="dzLKpmZICaWN"
import os
import pathlib
import matplotlib.pyplot as plt
import numpy as np
import seaborn as sns
import tensorflow as tf
from tensorflow.keras.layers.experimental import preprocessing
from tensorflow.keras import layers
from tensorflow.keras import models
from IPython import display
# Set seed for experiment reproducibility
seed = 42
tf.random.set_seed(seed)
np.random.seed(seed)
# + [markdown] id="yR0EdgrLCaWR"
# ## Import the Speech Commands dataset
#
# You'll write a script to download a portion of the [Speech Commands dataset](https://www.tensorflow.org/datasets/catalog/speech_commands). The original dataset consists of over 105,000 WAV audio files of people saying thirty different words. This data was collected by Google and released under a CC BY license, and you can help improve it by [contributing five minutes of your own voice](https://aiyprojects.withgoogle.com/open_speech_recording).
#
# You'll be using a portion of the dataset to save time with data loading. Extract the `mini_speech_commands.zip` and load it in using the `tf.data` API.
# + id="2-rayb7-3Y0I"
data_dir = pathlib.Path('data/mini_speech_commands')
if not data_dir.exists():
tf.keras.utils.get_file(
'mini_speech_commands.zip',
origin="http://storage.googleapis.com/download.tensorflow.org/data/mini_speech_commands.zip",
extract=True,
cache_dir='.', cache_subdir='data')
# + [markdown] id="BgvFq3uYiS5G"
# Check basic statistics about the dataset.
# + id="70IBxSKxA1N9"
commands = np.array(tf.io.gfile.listdir(str(data_dir)))
commands = commands[commands != 'README.md']
print('Commands:', commands)
# + [markdown] id="aMvdU9SY8WXN"
# Extract the audio files into a list and shuffle it.
# + id="hlX685l1wD9k"
filenames = tf.io.gfile.glob(str(data_dir) + '/*/*')
filenames = tf.random.shuffle(filenames)
num_samples = len(filenames)
print('Number of total examples:', num_samples)
print('Number of examples per label:',
len(tf.io.gfile.listdir(str(data_dir/commands[0]))))
print('Example file tensor:', filenames[0])
# + [markdown] id="9vK3ymy23MCP"
# Split the files into training, validation and test sets using a 80:10:10 ratio, respectively.
# + id="Cv_wts-l3KgD"
train_files = filenames[:6400]
val_files = filenames[6400: 6400 + 800]
test_files = filenames[-800:]
print('Training set size', len(train_files))
print('Validation set size', len(val_files))
print('Test set size', len(test_files))
# + [markdown] id="g2Cj9FyvfweD"
# ## Reading audio files and their labels
# + [markdown] id="j1zjcWteOcBy"
# The audio file will initially be read as a binary file, which you'll want to convert into a numerical tensor.
#
# To load an audio file, you will use [`tf.audio.decode_wav`](https://www.tensorflow.org/api_docs/python/tf/audio/decode_wav), which returns the WAV-encoded audio as a Tensor and the sample rate.
#
# A WAV file contains time series data with a set number of samples per second.
# Each sample represents the amplitude of the audio signal at that specific time. In a 16-bit system, like the files in `mini_speech_commands`, the values range from -32768 to 32767.
# The sample rate for this dataset is 16kHz.
# Note that `tf.audio.decode_wav` will normalize the values to the range [-1.0, 1.0].
# + id="9PjJ2iXYwftD"
def decode_audio(audio_binary):
audio, _ = tf.audio.decode_wav(audio_binary)
return tf.squeeze(audio, axis=-1)
# + [markdown] id="GPQseZElOjVN"
# The label for each WAV file is its parent directory.
# + id="8VTtX1nr3YT-"
def get_label(file_path):
parts = tf.strings.split(file_path, os.path.sep)
# Note: You'll use indexing here instead of tuple unpacking to enable this
# to work in a TensorFlow graph.
return parts[-2]
# + [markdown] id="E8Y9w_5MOsr-"
# Let's define a method that will take in the filename of the WAV file and output a tuple containing the audio and labels for supervised training.
# + id="WdgUD5T93NyT"
def get_waveform_and_label(file_path):
label = get_label(file_path)
audio_binary = tf.io.read_file(file_path)
waveform = decode_audio(audio_binary)
return waveform, label
# + [markdown] id="nvN8W_dDjYjc"
# You will now apply `process_path` to build your training set to extract the audio-label pairs and check the results. You'll build the validation and test sets using a similar procedure later on.
# + id="0SQl8yXl3kNP"
AUTOTUNE = tf.data.AUTOTUNE
files_ds = tf.data.Dataset.from_tensor_slices(train_files)
waveform_ds = files_ds.map(get_waveform_and_label, num_parallel_calls=AUTOTUNE)
# + [markdown] id="voxGEwvuh2L7"
# Let's examine a few audio waveforms with their corresponding labels.
# + id="8yuX6Nqzf6wT"
rows = 3
cols = 3
n = rows*cols
fig, axes = plt.subplots(rows, cols, figsize=(10, 12))
for i, (audio, label) in enumerate(waveform_ds.take(n)):
r = i // cols
c = i % cols
ax = axes[r][c]
ax.plot(audio.numpy())
ax.set_yticks(np.arange(-1.2, 1.2, 0.2))
label = label.numpy().decode('utf-8')
ax.set_title(label)
plt.show()
# + [markdown] id="EWXPphxm0B4m"
# ## Spectrogram
#
# You'll convert the waveform into a spectrogram, which shows frequency changes over time and can be represented as a 2D image. This can be done by applying the short-time Fourier transform (STFT) to convert the audio into the time-frequency domain.
#
# A Fourier transform ([`tf.signal.fft`](https://www.tensorflow.org/api_docs/python/tf/signal/fft)) converts a signal to its component frequencies, but loses all time information. The STFT ([`tf.signal.stft`](https://www.tensorflow.org/api_docs/python/tf/signal/stft)) splits the signal into windows of time and runs a Fourier transform on each window, preserving some time information, and returning a 2D tensor that you can run standard convolutions on.
#
# STFT produces an array of complex numbers representing magnitude and phase. However, you'll only need the magnitude for this tutorial, which can be derived by applying `tf.abs` on the output of `tf.signal.stft`.
#
# Choose `frame_length` and `frame_step` parameters such that the generated spectrogram "image" is almost square. For more information on STFT parameters choice, you can refer to [this video](https://www.coursera.org/lecture/audio-signal-processing/stft-2-tjEQe) on audio signal processing.
#
# You also want the waveforms to have the same length, so that when you convert it to a spectrogram image, the results will have similar dimensions. This can be done by simply zero padding the audio clips that are shorter than one second.
#
# + id="_4CK75DHz_OR"
def get_spectrogram(waveform):
# Padding for files with less than 16000 samples
zero_padding = tf.zeros([16000] - tf.shape(waveform), dtype=tf.float32)
# Concatenate audio with padding so that all audio clips will be of the
# same length
waveform = tf.cast(waveform, tf.float32)
equal_length = tf.concat([waveform, zero_padding], 0)
spectrogram = tf.signal.stft(
equal_length, frame_length=255, frame_step=128)
spectrogram = tf.abs(spectrogram)
return spectrogram
# + id="YWFcNngKR1al"
import numpy as np
class Uint8LogMelFeatureExtractor(object):
"""Provide uint8 log mel spectrogram slices from an AudioRecorder object.
This class provides one public method, get_next_spectrogram(), which gets
a specified number of spectral slices from an AudioRecorder.
"""
def __init__(self, num_frames_hop=49):
self.spectrogram_window_length_seconds = 0.025
self.spectrogram_hop_length_seconds = 0.010
self.num_mel_bins = 32
self.frame_length_spectra = 98
if self.frame_length_spectra % num_frames_hop:
raise ValueError('Invalid num_frames_hop value (%d), '
'must devide %d' % (num_frames_hop,
self.frame_length_spectra))
self.frame_hop_spectra = num_frames_hop
self._norm_factor = 3
self._clear_buffers()
def _clear_buffers(self):
self._audio_buffer = np.array([], dtype=np.int16).reshape(0, 1)
self._spectrogram = np.zeros((self.frame_length_spectra, self.num_mel_bins),
dtype=np.float32)
def _spectrogram_underlap_samples(self, audio_sample_rate_hz):
return int((self.spectrogram_window_length_seconds -
self.spectrogram_hop_length_seconds) * audio_sample_rate_hz)
def _frame_duration_seconds(self, num_spectra):
return (self.spectrogram_window_length_seconds +
(num_spectra - 1) * self.spectrogram_hop_length_seconds)
def compute_spectrogram_and_normalize(self, audio_samples, audio_sample_rate_hz):
spectrogram = self._compute_spectrogram(audio_samples, audio_sample_rate_hz)
spectrogram -= np.mean(spectrogram, axis=0)
if self._norm_factor:
spectrogram /= self._norm_factor * np.std(spectrogram, axis=0)
spectrogram += 1
spectrogram *= 127.5
return np.maximum(0, np.minimum(255, spectrogram)).astype(np.float32)
def _compute_spectrogram(self, audio_samples, audio_sample_rate_hz):
"""Compute log-mel spectrogram and scale it to uint8."""
samples = audio_samples.flatten() / float(2**15)
spectrogram = 30 * (
mel_features.log_mel_spectrogram(
samples,
audio_sample_rate_hz,
log_offset=0.001,
window_length_secs=self.spectrogram_window_length_seconds,
hop_length_secs=self.spectrogram_hop_length_seconds,
num_mel_bins=self.num_mel_bins,
lower_edge_hertz=60,
upper_edge_hertz=3800) - np.log(1e-3))
return spectrogram
def _get_next_spectra(self, recorder, num_spectra):
"""Returns the next spectrogram.
Compute num_spectra spectrogram samples from an AudioRecorder.
Blocks until num_spectra spectrogram slices are available.
Args:
recorder: an AudioRecorder object from which to get raw audio samples.
num_spectra: the number of spectrogram slices to return.
Returns:
num_spectra spectrogram slices computed from the samples.
"""
required_audio_duration_seconds = self._frame_duration_seconds(num_spectra)
logger.info("required_audio_duration_seconds %f",
required_audio_duration_seconds)
required_num_samples = int(
np.ceil(required_audio_duration_seconds *
recorder.audio_sample_rate_hz))
logger.info("required_num_samples %d, %s", required_num_samples,
str(self._audio_buffer.shape))
audio_samples = np.concatenate(
(self._audio_buffer,
recorder.get_audio(required_num_samples - len(self._audio_buffer))[0]))
self._audio_buffer = audio_samples[
required_num_samples -
self._spectrogram_underlap_samples(recorder.audio_sample_rate_hz):]
spectrogram = self._compute_spectrogram(
audio_samples[:required_num_samples], recorder.audio_sample_rate_hz)
assert len(spectrogram) == num_spectra
return spectrogram
def get_next_spectrogram(self, recorder):
"""Get the most recent spectrogram frame.
Blocks until the frame is available.
Args:
recorder: an AudioRecorder instance which provides the audio samples.
Returns:
The next spectrogram frame as a uint8 numpy array.
"""
assert recorder.is_active
logger.info("self._spectrogram shape %s", str(self._spectrogram.shape))
self._spectrogram[:-self.frame_hop_spectra] = (
self._spectrogram[self.frame_hop_spectra:])
self._spectrogram[-self.frame_hop_spectra:] = (
self._get_next_spectra(recorder, self.frame_hop_spectra))
# Return a copy of the internal state that's safe to persist and won't
# change the next time we call this function.
logger.info("self._spectrogram shape %s", str(self._spectrogram.shape))
spectrogram = self._spectrogram.copy()
spectrogram -= np.mean(spectrogram, axis=0)
if self._norm_factor:
spectrogram /= self._norm_factor * np.std(spectrogram, axis=0)
spectrogram += 1
spectrogram *= 127.5
return np.maximum(0, np.minimum(255, spectrogram)).astype(np.uint8)
# + id="epNtx_cmRkNP"
feature_extractor = Uint8LogMelFeatureExtractor()
def get_spectrogram2(waveform):
"""
# Padding for files with less than 16000 samples
zero_padding = tf.zeros([16000] - tf.shape(waveform), dtype=tf.float32)
# Concatenate audio with padding so that all audio clips will be of the
# same length
waveform = tf.cast(waveform, tf.float32)
equal_length = tf.concat([waveform, zero_padding], 0)
spectrogram = tf.signal.stft(
equal_length, frame_length=255, frame_step=128)
spectrogram = tf.abs(spectrogram)
return spectrogram
"""
waveform = waveform.numpy()
#print(waveform.shape)
#print(type(waveform))
spectrogram = feature_extractor.compute_spectrogram_and_normalize(waveform, 16000)
return spectrogram
for waveform, label in waveform_ds.take(1):
label2 = label.numpy().decode('utf-8')
spectrogram2 = get_spectrogram2(waveform)
print('Label:', label2)
print('Waveform shape:', waveform.shape)
print('Spectrogram shape:', spectrogram2.shape)
print('Spectrogram type:', spectrogram2.dtype)
# + [markdown] id="5rdPiPYJphs2"
# Next, you will explore the data. Compare the waveform, the spectrogram and the actual audio of one example from the dataset.
# + id="4Mu6Y7Yz3C-V"
for waveform, label in waveform_ds.take(1):
label = label.numpy().decode('utf-8')
spectrogram = get_spectrogram(waveform)
print('Label:', label)
print('Waveform shape:', waveform.shape)
print('Spectrogram shape:', spectrogram.shape)
print('Audio playback')
print('Spectrogram type:', spectrogram.dtype)
display.display(display.Audio(waveform, rate=16000))
# + id="e62jzb36-Jog"
def plot_spectrogram(spectrogram, ax):
# Convert to frequencies to log scale and transpose so that the time is
# represented in the x-axis (columns).
log_spec = np.log(spectrogram.T)
height = log_spec.shape[0]
X = np.arange(16000, step=height + 1)
Y = range(height)
ax.pcolormesh(X, Y, log_spec)
fig, axes = plt.subplots(2, figsize=(12, 8))
timescale = np.arange(waveform.shape[0])
axes[0].plot(timescale, waveform.numpy())
axes[0].set_title('Waveform')
axes[0].set_xlim([0, 16000])
plot_spectrogram(spectrogram.numpy(), axes[1])
axes[1].set_title('Spectrogram')
plt.show()
# + [markdown] id="GyYXjW07jCHA"
# Now transform the waveform dataset to have spectrogram images and their corresponding labels as integer IDs.
# + id="43IS2IouEV40"
def get_spectrogram_and_label_id(audio, label):
spectrogram = get_spectrogram(audio)
spectrogram = tf.expand_dims(spectrogram, -1)
label_id = tf.argmax(label == commands)
return spectrogram, label_id
# + id="yEVb_oK0oBLQ"
spectrogram_ds = waveform_ds.map(
get_spectrogram_and_label_id, num_parallel_calls=AUTOTUNE)
# + [markdown] id="6gQpAAgMnyDi"
# Examine the spectrogram "images" for different samples of the dataset.
# + id="QUbHfTuon4iF"
rows = 3
cols = 3
n = rows*cols
fig, axes = plt.subplots(rows, cols, figsize=(10, 10))
for i, (spectrogram, label_id) in enumerate(spectrogram_ds.take(n)):
r = i // cols
c = i % cols
ax = axes[r][c]
plot_spectrogram(np.squeeze(spectrogram.numpy()), ax)
ax.set_title(commands[label_id.numpy()])
ax.axis('off')
plt.show()
# + [markdown] id="z5KdY8IF8rkt"
# ## Build and train the model
#
# Now you can build and train your model. But before you do that, you'll need to repeat the training set preprocessing on the validation and test sets.
# + id="10UI32QH_45b"
def preprocess_dataset(files):
files_ds = tf.data.Dataset.from_tensor_slices(files)
output_ds = files_ds.map(get_waveform_and_label, num_parallel_calls=AUTOTUNE)
output_ds = output_ds.map(
get_spectrogram_and_label_id, num_parallel_calls=AUTOTUNE)
return output_ds
# + id="HNv4xwYkB2P6"
train_ds = spectrogram_ds
val_ds = preprocess_dataset(val_files)
test_ds = preprocess_dataset(test_files)
# + id="0e9yyQZuYzYx"
def only_load_dataset(files):
files_ds = tf.data.Dataset.from_tensor_slices(files)
output_ds = files_ds.map(get_waveform_and_label, num_parallel_calls=AUTOTUNE)
return output_ds
train_waveform_data = only_load_dataset(train_files)
val_waveform_data = only_load_dataset(val_files)
test_waveform_data = only_load_dataset(test_files)
# + [markdown] id="assnWo6SB3lR"
# Batch the training and validation sets for model training.
# + id="UgY9WYzn61EX"
batch_size = 64
train_ds = train_ds.batch(batch_size)
val_ds = val_ds.batch(batch_size)
# + [markdown] id="GS1uIh6F_TN9"
# Add dataset [`cache()`](https://www.tensorflow.org/api_docs/python/tf/data/Dataset#cache) and [`prefetch()`](https://www.tensorflow.org/api_docs/python/tf/data/Dataset#prefetch) operations to reduce read latency while training the model.
# + id="fdZ6M-F5_QzY"
train_ds = train_ds.cache().prefetch(AUTOTUNE)
val_ds = val_ds.cache().prefetch(AUTOTUNE)
# + [markdown] id="rwHkKCQQb5oW"
# For the model, you'll use a simple convolutional neural network (CNN), since you have transformed the audio files into spectrogram images.
# The model also has the following additional preprocessing layers:
# - A [`Resizing`](https://www.tensorflow.org/api_docs/python/tf/keras/layers/experimental/preprocessing/Resizing) layer to downsample the input to enable the model to train faster.
# - A [`Normalization`](https://www.tensorflow.org/api_docs/python/tf/keras/layers/experimental/preprocessing/Normalization) layer to normalize each pixel in the image based on its mean and standard deviation.
#
# For the `Normalization` layer, its `adapt` method would first need to be called on the training data in order to compute aggregate statistics (i.e. mean and standard deviation).
# + id="ALYz7PFCHblP"
#for spectrogram, _ in spectrogram_ds.take(1):
# input_shape = spectrogram.shape
for data_item, label in train_waveform_data.take(10):
spectrogram = feature_extractor.compute_spectrogram_and_normalize(data_item.numpy(), 16000)
print(spectrogram.shape)
if spectrogram.shape[0] != 98:
continue
input_shape = (spectrogram.shape[0], spectrogram.shape[1], 1)
print('Input shape:', input_shape)
num_labels = len(commands)
norm_layer = preprocessing.Normalization()
norm_layer.adapt(spectrogram_ds.map(lambda x, _: x))
model = models.Sequential([
layers.Input(shape=input_shape),
preprocessing.Resizing(32, 32),
norm_layer,
layers.Conv2D(32, 3, activation='relu'),
layers.Conv2D(64, 3, activation='relu'),
layers.MaxPooling2D(),
layers.Dropout(0.25),
layers.Flatten(),
layers.Dense(128, activation='relu'),
layers.Dropout(0.5),
layers.Dense(num_labels),
])
model.summary()
# + id="wFjj7-EmsTD-"
model.compile(
optimizer=tf.keras.optimizers.Adam(),
loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=['accuracy'],
)
# + id="b62-5k6qVaIM"
new_train_data = []
new_train_labels = []
new_val_data = []
new_val_labels = []
new_test_data = []
new_test_labels = []
for data_item, label in train_waveform_data:
spectrogram = feature_extractor.compute_spectrogram_and_normalize(data_item.numpy(), 16000)
label = label.numpy().decode('utf-8')
label_id = tf.argmax(label == commands)
# NOTE: Spectrogram shape is not always the same
if spectrogram.shape[0] != 98:
continue
new_train_data.append(spectrogram)
new_train_labels.append(label_id)
for data_item, label in val_waveform_data:
spectrogram = feature_extractor.compute_spectrogram_and_normalize(data_item.numpy(), 16000)
label = label.numpy().decode('utf-8')
label_id = tf.argmax(label == commands)
if spectrogram.shape[0] != 98:
continue
new_val_data.append(spectrogram)
new_val_labels.append(label_id)
for data_item, label in test_waveform_data:
spectrogram = feature_extractor.compute_spectrogram_and_normalize(data_item.numpy(), 16000)
label = label.numpy().decode('utf-8')
label_id = tf.argmax(label == commands)
if spectrogram.shape[0] != 98:
continue
new_test_data.append(spectrogram)
new_test_labels.append(label_id)
new_train_data = np.array(new_train_data).astype('float32')
new_val_data = np.array(new_val_data).astype('float32')
new_test_data = np.array(new_test_data).astype('float32')
new_train_labels = np.array(new_train_labels)
new_val_labels = np.array(new_val_labels)
new_test_labels = np.array(new_test_labels)
new_train_data = np.expand_dims(new_train_data, axis=3)
new_val_data = np.expand_dims(new_val_data, axis=3)
new_test_data = np.expand_dims(new_test_data, axis=3)
print('--------')
print(new_train_data.shape)
print(new_val_data.shape)
print(new_test_data.shape)
print(new_train_labels.shape)
print(new_val_labels.shape)
print(new_test_labels.shape)
print('--------')
# + id="ttioPJVMcGtq"
EPOCHS = 30
#history = model.fit(
# train_ds,
# validation_data=val_ds,
# epochs=EPOCHS,
# callbacks=tf.keras.callbacks.EarlyStopping(verbose=1, patience=2),
#)
history = model.fit(
new_train_data, new_train_labels,
validation_data=(new_val_data, new_val_labels),
epochs=EPOCHS,
#callbacks=tf.keras.callbacks.EarlyStopping(verbose=1, patience=2),
)
# + [markdown] id="gjpCDeQ4mUfS"
# Let's check the training and validation loss curves to see how your model has improved during training.
# + id="nzhipg3Gu2AY"
metrics = history.history
plt.plot(history.epoch, metrics['loss'], metrics['val_loss'])
plt.legend(['loss', 'val_loss'])
plt.show()
# + [markdown] id="5ZTt3kO3mfm4"
# ## Evaluate test set performance
#
# Let's run the model on the test set and check performance.
# + id="biU2MwzyAo8o"
#test_audio = []
#test_labels = []
#for audio, label in test_ds:
# test_audio.append(audio.numpy())
# test_labels.append(label.numpy())
#test_audio = np.array(test_audio)
#test_labels = np.array(test_labels)
test_audio = new_test_data
test_labels = new_test_labels
# + id="ktUanr9mRZky"
y_pred = np.argmax(model.predict(test_audio), axis=1)
y_true = test_labels
test_acc = sum(y_pred == y_true) / len(y_true)
print(f'Test set accuracy: {test_acc:.0%}')
# + [markdown] id="en9Znt1NOabH"
# ### Display a confusion matrix
#
# A confusion matrix is helpful to see how well the model did on each of the commands in the test set.
# + id="LvoSAOiXU3lL"
confusion_mtx = tf.math.confusion_matrix(y_true, y_pred)
plt.figure(figsize=(10, 8))
sns.heatmap(confusion_mtx, xticklabels=commands, yticklabels=commands,
annot=True, fmt='g')
plt.xlabel('Prediction')
plt.ylabel('Label')
plt.show()
# + [markdown] id="mQGi_mzPcLvl"
# ## Run inference on an audio file
#
# Finally, verify the model's prediction output using an input audio file of someone saying "no." How well does your model perform?
# + id="wvhqyCJIjJyp"
!#ls data/mini_speech_commands/up
# + id="zRxauKMdhofU"
sample_file = data_dir/'no/01bb6a2a_nohash_0.wav'
#sample_file = data_dir/'no/ac7840d8_nohash_1.wav'
#sample_file = data_dir/'no/5588c7e6_nohash_0.wav'
#sample_file = data_dir/'up/52e228e9_nohash_0.wav'
#sample_ds = preprocess_dataset([str(sample_file)])
X = only_load_dataset([str(sample_file)])
for waveform, label in X.take(1):
label = label.numpy().decode('utf-8')
print(waveform, label)
spectrogram = feature_extractor.compute_spectrogram_and_normalize(waveform.numpy(), 16000)
# NOTE: Dimensions need to be expanded
spectrogram = np.expand_dims(spectrogram, axis=-1)
spectrogram = np.expand_dims(spectrogram, axis=0)
print(spectrogram.shape)
prediction = model(spectrogram)
print(prediction.shape)
plt.bar(commands, tf.nn.softmax(prediction[0]))
plt.title(f'Predictions for "{label}"')
plt.show()
#for spectrogram, label in sample_ds.batch(1):
# prediction = model(spectrogram)
# plt.bar(commands, tf.nn.softmax(prediction[0]))
# plt.title(f'Predictions for "{commands[label[0]]}"')
# plt.show()
# + id="8a7xRQPvYwnW"
print(model)
converter = tf.lite.TFLiteConverter.from_keras_model(model)
tflite_model = converter.convert()
# Save the model.
with open('model.tflite', 'wb') as f:
f.write(tflite_model)
# + id="2JaL8Tf9ZS83"
# !ls -l
# + id="1EnXaa-8aAsc"
# https://www.tensorflow.org/lite/guide/inference
interpreter = tf.lite.Interpreter(model_path="model.tflite")
interpreter.allocate_tensors()
# Get input and output tensors.
input_details = interpreter.get_input_details()
output_details = interpreter.get_output_details()
print(input_details)
print(output_details)
# Test the model on random input data.
input_shape = input_details[0]['shape']
input_data = np.array(np.random.random_sample(input_shape), dtype=np.float32)
interpreter.set_tensor(input_details[0]['index'], input_data)
interpreter.invoke()
# The function `get_tensor()` returns a copy of the tensor data.
# Use `tensor()` in order to get a pointer to the tensor.
output_data = interpreter.get_tensor(output_details[0]['index'])
print(output_data)
# + id="pHUa-11SZhUz"
sample_file = data_dir/'no/01bb6a2a_nohash_0.wav'
#sample_ds = preprocess_dataset([str(sample_file)])
#waveform, label = get_waveform_and_label(sample_file)
#spectrogram = feature_extractor._compute_spectrogram(waveform, 16000)
X = only_load_dataset([str(sample_file)])
for waveform, label in X.take(1):
label = label.numpy().decode('utf-8')
spectrogram = feature_extractor.compute_spectrogram_and_normalize(waveform.numpy(), 16000)
spectrogram = np.expand_dims(spectrogram, axis=-1)
spectrogram = np.expand_dims(spectrogram, axis=0)
print('Original--------------------')
print(spectrogram.shape)
prediction = model(spectrogram)
print(prediction)
print('TFLITE--------------------')
# NOTE: dtype needs to be np.float32
input_data = np.array(spectrogram, dtype=np.float32)
print(input_data.shape)
interpreter.set_tensor(input_details[0]['index'], input_data)
interpreter.invoke()
prediction2 = interpreter.get_tensor(output_details[0]['index'])
print(prediction2)
print(np.argmax(np.array(prediction).flatten()))
print(np.argmax(np.array(prediction2).flatten()))
# NOTE: Remember to add softmax after the prediction
plt.bar(commands, tf.nn.softmax(prediction[0]))
plt.title(f'Predictions for "{label}"')
plt.show()
plt.imshow(np.squeeze(spectrogram).T)
plt.show()
# + [markdown] id="VgWICqdqQNaQ"
# You can see that your model very clearly recognized the audio command as "no."
# + id="nGvwJZDteCnh"
from google.colab import files
files.download('model.tflite')
# + [markdown] id="J3jF933m9z1J"
# ## Next steps
#
# This tutorial showed how you could do simple audio classification using a convolutional neural network with TensorFlow and Python.
#
# * To learn how to use transfer learning for audio classification, check out the [Sound classification with YAMNet](https://www.tensorflow.org/hub/tutorials/yamnet) tutorial.
#
# * To build your own interactive web app for audio classification, consider taking the [TensorFlow.js - Audio recognition using transfer learning codelab](https://codelabs.developers.google.com/codelabs/tensorflowjs-audio-codelab/index.html#0).
#
# * TensorFlow also has additional support for [audio data preparation and augmentation](https://www.tensorflow.org/io/tutorials/audio) to help with your own audio-based projects.
#
| sound/simple_audio_new_spectrogram_numpy_and_normalize.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/besarria/Macroeconomia/blob/main/Aula_01_Introducao.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="2Qx3Sh93PTcN"
# # Curso de Macroeconomia I
# # Prof. <NAME>
# # Tópico: Dados conjunturais
# + id="14-C0SMFPTcR"
# Baixar dados BCB
# Quarterly GDP (1995=100) - seasonally adjusted data - GDP at market prices
# Code: BCB/1253
# link: https://www.quandl.com/data/BCB/1253-Quarterly-GDP-1995-100-seasonally-adjusted-data-GDP-at-market-prices
# + [markdown] id="6SmyEs7lPTcS"
# # Biblioteca
# + colab={"base_uri": "https://localhost:8080/"} id="Be2lexcOPTcT" outputId="ac9017d7-a9f9-4abb-afbb-8dcacfbad71f"
# #!pip install quandl
import quandl
# + [markdown] id="AS4Jo_4XPTcT"
# # Importação de dados
# + id="3BxbzqAwPTcU"
GDP = quandl.get(["BCB/1253"], trim_start='2000-01-01', trim_end='2020-03-31')
# + colab={"base_uri": "https://localhost:8080/"} id="yb6AdUeTPTcU" outputId="7216d1aa-063d-4125-a0f9-ced118ffcbc7"
print(GDP.index)
# + colab={"base_uri": "https://localhost:8080/"} id="mcADUx6BPTcV" outputId="335285a5-efe8-4675-b730-f2e4d2d49f00"
print(GDP.head(10))
# + colab={"base_uri": "https://localhost:8080/", "height": 279} id="QU9CZZyJPTcW" outputId="779abfa8-3ffa-43f9-a287-175f8eb36b3d"
quandl.get('BCB/1253', start_date="1995-01-01").plot(figsize=(14, 4)); # GDP
# + colab={"base_uri": "https://localhost:8080/"} id="C1v5JSv1PTcW" outputId="72ff9614-57be-49dc-8864-39c868a6564d"
from matplotlib.pylab import rcParams
rcParams['figure.figsize'] = 15, 6
import statsmodels.api as sm #permite estimar modelos, realizar testes estatistícos
import statsmodels.graphics as graf
import statsmodels.api as sm
# + id="qn8E6M--PTcX"
hp_cycle, hp_trend = sm.tsa.filters.hpfilter(GDP, 1600) # Estou utilizando o fator de suavização lambda = 1600, referente a dados trimestrais
# + id="DIZWtQovPTcX"
ip_decomp_hp = GDP[['BCB/1253 - Value']] # aqui estou selecionando a coluna de valores
ip_decomp_hp["cycle"] = hp_cycle
ip_decomp_hp["trend"] = hp_trend
# + colab={"base_uri": "https://localhost:8080/", "height": 238} id="eupU2_nDPTcX" outputId="2a68e8ea-0241-478d-a9b7-3578c3329a2a"
ip_decomp_hp.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 409} id="6pwrUARQPTcY" outputId="71c6d427-884e-4b8d-f1fa-242aa863a675"
import matplotlib.pyplot as plt
fig, ax = plt.subplots()
ip_decomp_hp[["BCB/1253 - Value", "trend"]]["1995-01-01":].plot(ax=ax, fontsize=16);
plt.title('Quarterly GDP (1995=100) - seasonally adjusted data')
plt.savefig('GDP')
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 409} id="k0vBJK03PTcY" outputId="a93a00ad-127b-49a8-810f-6161b816749c"
fig, ax = plt.subplots()
ip_decomp_hp[['cycle']]['1995-01-01':].plot(ax=ax, fontsize=16)
plt.title('Cycle GDP (1995=100) - seasonally adjusted data')
plt.savefig('cycle_GDP')
plt.show()
# + [markdown] id="fcz8UcJ4PTcY"
# # GDP per capita at current prices in US$
# + id="tZbyMFFYPTcZ"
GDP_per = quandl.get(["BCB/7325"], trim_start='1962-01-01', trim_end='2012-12-31')
# + colab={"base_uri": "https://localhost:8080/"} id="U2bH-tvpPTcZ" outputId="68f69937-ebe4-4e81-c94d-b1037af2d803"
print(GDP_per.head(10))
# + id="_VmRwhh9PTcZ"
hp_cycle_per, hp_trend_per = sm.tsa.filters.hpfilter(GDP_per, 100)
# + id="K4_vYS8MPTcZ"
ip_decomp_hp_per = GDP_per[['BCB/7325 - Value']]
ip_decomp_hp_per["cycle"] = hp_cycle_per
ip_decomp_hp_per["trend"] = hp_trend_per
# + colab={"base_uri": "https://localhost:8080/", "height": 238} id="cjXdhEFEPTca" outputId="b274f6f1-8d27-4275-ba54-f914645a368e"
ip_decomp_hp_per.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 409} id="c7xSRBw8PTce" outputId="117efb33-53dc-4134-d6d2-1a677efbce7b"
fig, ax = plt.subplots()
ip_decomp_hp_per[["BCB/7325 - Value", "trend"]]["1962-01-01":].plot(ax=ax, fontsize=16);
plt.title('GDP per capita at current prices in US$')
plt.savefig('GDP_per_capita')
plt.show()
# + [markdown] id="Hz7iVDumPTce"
# # Unemployment
# + id="LBmJg6u5PTcf"
Unemp = quandl.get(["BCB/24369"], trim_start='2002-03-01', trim_end='2019-06-30')
# + colab={"base_uri": "https://localhost:8080/"} id="QJ0u1zOnPTcf" outputId="ac1adec5-12ba-49f6-9cb7-16de58921191"
print(Unemp.head(10))
# + colab={"base_uri": "https://localhost:8080/", "height": 409} id="f_-VUW89PTcf" outputId="6d8a20cd-4c50-4364-e76e-5eba6bb3ce5e"
fig, ax = plt.subplots()
Unemp[["BCB/24369 - Value"]]["2002-03-01":].plot(ax=ax, fontsize=16);
plt.title('Unemployment rate, PNADC (%)')
plt.savefig('unemployment')
plt.show()
# + [markdown] id="QRIhefKGPTcf"
# # Inflation
# + id="mTMcHveyPTcf"
Inf = quandl.get(["BCB/13522"], trim_start='2002-03-01', trim_end='2019-03-31')
# + colab={"base_uri": "https://localhost:8080/"} id="0iFa2ayhPTcg" outputId="4d7b9226-00df-49b1-d4ef-f5f7c82bdc02"
print(Inf.head(10))
# + colab={"base_uri": "https://localhost:8080/", "height": 409} id="PV6RxUGuPTcg" outputId="1d3ca07b-fd54-4bc6-d4ed-1376a60552e4"
fig, ax = plt.subplots()
Inf[["BCB/13522 - Value"]]["2002-03-01":].plot(ax=ax, fontsize=16);
plt.title('National consumer price index (IPCA), in 12 months (%)')
plt.savefig('inflation')
plt.show()
# + [markdown] id="aKPbhb9pPTcg"
# # Implicit deflator
# + id="w5h6j0IBPTcg"
Def = quandl.get(["BCB/1211"], trim_start='2000-01-01', trim_end='2019-06-30')
# + colab={"base_uri": "https://localhost:8080/"} id="taLqE58vPTcg" outputId="002e8038-c234-4900-a886-230d1828322e"
print(Def.head(10))
# + colab={"base_uri": "https://localhost:8080/", "height": 409} id="ziKTt8cdPTch" outputId="7a2fb226-8225-4aa1-db11-3d9c5cc218aa"
fig, ax = plt.subplots()
Def[["BCB/1211 - Value"]]["2000-01-01":].plot(ax=ax, fontsize=16);
plt.title('Implicit deflator, (%)')
plt.savefig('unemployment')
plt.show()
# + id="qSN8fP2HPTch"
Def_02 = (1 + Def/100)
# + colab={"base_uri": "https://localhost:8080/", "height": 404} id="M4VFrlVTPTch" outputId="9e432c2b-ea8d-43fc-c48d-390db0f9fe61"
# Plot
plt.plot(Def_02)
plt.title('Implicit deflator, (%)', fontsize=8)
plt.plot()
# + [markdown] id="gr4WJZMVPTch"
# # Salvando dados de Inflação e Desemprego
# + id="67e2SN63PTch"
import os
# + id="xNv8uKRrPTch"
os.chdir("C:\\Users\\CASSIO\\OneDrive\\UFPB\\Macroeconomia\\Aplicacoes_Python") # Definir um diretório
# + id="GjQwQBD4PTch"
Unemp.to_csv("Unemp.csv") # salvando o banco de dados
# + id="sUy13v8-PTci"
Inf.to_csv("Inf.csv")
# + id="SmFQkb7fPTci"
import pandas as pd
# + id="dN02EINKPTci"
a = pd.read_csv("Unemp.csv")
b = pd.read_csv("Inf.csv")
# + id="uFyc1YHpPTci"
merged = a.merge(b)
merged.to_csv("output.csv", index=False)
# + id="Ng0n6xnqPTci"
from pandas import read_csv
# + id="DtouX783PTci"
series = read_csv('output.csv', header=0, index_col=0, parse_dates=True,
squeeze=True)
# + id="w-CoGbKHPTci" outputId="b032f97f-16a4-4f20-c329-d4c119bb28b5"
print(series.head(10))
# + id="D8-RsC55PTci"
from matplotlib import pyplot
# + id="Usu1MREEPTcj" outputId="7e7c252a-8ede-49bc-b599-f273bea82cf6"
series.plot()
pyplot.show()
# + [markdown] id="ALd5zBz-PTcj"
# # Estatísticas descritivas
# + id="8_EnFEANPTcj" outputId="1caecf14-ce49-430c-98c2-c13d34f454d9"
series.describe()
# + [markdown] id="C7tultz_PTcj"
# # Histograma
# + id="HkZ3YVRXPTcj" outputId="5f92eafe-8082-4fe6-b077-03c395cf0524"
import seaborn as sns
import matplotlib.pyplot as plt
hist, ax = plt.subplots()
ax = sns.distplot(series['BCB/24369 - Value'])
ax.set_title('Histograma da taxa de desemprego')
plt.show()
# + [markdown] id="LnShSVUbPTcj"
# # Criando logaritmo das séries
# + id="9uDEH-5vPTcj"
import numpy as np
series['logdesemprego'] = np.log(series['BCB/24369 - Value'])
series['loginflacao'] = np.log(series['BCB/13522 - Value'])
# + id="wbV_sMDIPTck" outputId="b5b814f7-80aa-45b2-cd50-86e3959525de"
series
# + id="lNOm9wZ0PTck" outputId="823f96a9-2674-43ed-ee8d-1bf1a764f260"
series['log_razao'] = np.log(series['BCB/24369 - Value']/series['BCB/13522 - Value'])
series
# + [markdown] id="YqVnx2KAPTck"
# # Verificação de valores NAN
# + id="nLeQqeboPTck" outputId="7603c35e-c1ae-4188-db38-10febbf2a567"
np.isnan(series).any()
# + [markdown] id="jfFuDCn4PTck"
# # Dispersão entre inflação e desemprego
# + id="0pQPzId3PTck"
import seaborn as sns
# + id="8napnAi3PTck" outputId="88017de7-9d08-437c-de35-1542db83dc04"
scatter, ax = plt.subplots()
ax = sns.regplot(x='BCB/24369 - Value', y='BCB/13522 - Value', data=series)
ax.set_title('Relação entre inflação e desemprego')
ax.set_xlabel('Desemprego')
ax.set_ylabel('Inflação')
plt.savefig('unemp_inf')
plt.show()
# + [markdown] id="Volj3BpzPTcl"
# # Índice do Emprego Formal
# + id="cr98tRscPTcl"
Emp_Form = quandl.get(["BCB/10802"], trim_start='2000-01-01', trim_end='2019-12-30')
# + id="coVT1srXPTcl" outputId="3a5e73ea-fc41-40d2-9207-655fb6960277"
print(Emp_Form.head(10))
# + id="zeVR9ktHPTcl"
import matplotlib.pyplot as plt
# + id="kHGvfOIdPTcl" outputId="fb29b7bb-c12a-4e70-ed56-356d1e63e9a4"
fig, ax = plt.subplots()
Emp_Form[["BCB/10802 - Value"]]["2000-01-01":].plot(ax=ax, fontsize=16);
plt.title('Employed people, PNADC (Monthly)')
plt.savefig('Employed')
plt.show()
# + [markdown] id="cFBVjX5OPTcl"
# # Paraíba
# + [markdown] id="36EpZjTQPTcl"
# ### Operações de crédito
# + id="ASVwT49FPTcm"
default = quandl.get(["BCB/15939"], trim_start='2002-03-01', trim_end='2020-03-31')
# + id="iBzrKw7pPTcm" outputId="905be320-7f73-4ed3-9a2e-bb06be431292"
# Plot
plt.plot(default)
plt.title('Taxa de inadimplência das operações de crédito na Paraíba (em %)', fontsize=8)
plt.savefig('default')
plt.plot()
# + id="oU_RXVufPTcm"
default_corp = quandl.get(["BCB/15907"], trim_start='2002-03-01', trim_end='2020-03-31')
# + id="X6VXSkAXPTcm" outputId="74066e71-fed9-41ab-e8bf-62064611d1db"
# Plot
plt.plot(default_corp)
plt.title('Taxa de inadimplência das operações de crédito na Paraíba, corporações (em %)', fontsize=8)
plt.savefig('default')
plt.plot()
# + id="dECgnWWxPTcm"
default_fis = quandl.get(["BCB/15875"], trim_start='2002-03-01', trim_end='2020-03-31')
# + id="sCUX5JDnPTcm"
import os
# + id="TxOpBVamPTcm"
os.chdir("C:\\Users\\CASSIO\\OneDrive\\UFPB\\Macroeconomia\\Aplicacoes_Python")
# + id="ukeP6FbGPTcm"
default.to_csv("default.csv")
# + id="ui-lRh9YPTcn"
default_corp.to_csv("default_corp.csv")
# + id="vrvPGuk1PTcn"
default_fis.to_csv("default_fis.csv")
# + id="h5YroiBkPTcn"
a = pd.read_csv("default.csv")
b = pd.read_csv("default_corp.csv")
c = pd.read_csv("default_fis.csv")
# + id="JevbVU9GPTcn"
merged = a.merge(b).merge(c)
merged.to_csv("default.csv", index=False)
# + id="4VPVy5MvPTcn"
series = read_csv('default.csv', header=0, index_col=0, parse_dates=True,
squeeze=True)
# + id="lusL7LP_PTcn" outputId="82e02fc3-0483-495b-9ed6-b4763425ed17"
series.plot()
plt.title('Taxa de inadimplência das operações de crédito agregado (azul), pessoa física (verde) e por corporações na Paraíba, corporações (em %)', fontsize=10)
plt.savefig('fig_default')
pyplot.show()
# + [markdown] id="Kb1-kcUrPTcn"
# # Dívida estados e municípios
# + id="apVl31b5PTcn"
externa = quandl.get(["BCB/4528"], trim_start='2002-03-01', trim_end='2020-03-31')
interna = quandl.get(["BCB/4517"], trim_start='2002-03-01', trim_end='2020-03-31')
total = quandl.get(["BCB/4506"], trim_start='2002-03-01', trim_end='2020-03-31')
# + id="_9GV4AcTPTco"
externa.to_csv("externa.csv")
interna.to_csv("interna.csv")
total.to_csv("total.csv")
# + id="b1etOJoRPTco"
a = pd.read_csv("externa.csv")
b = pd.read_csv("interna.csv")
c = pd.read_csv("total.csv")
# + id="8a8_nvkHPTco"
merged = a.merge(b).merge(c)
merged.to_csv("divida.csv", index=False)
# + id="wWApn8KvPTco"
divida = read_csv('divida.csv', header=0, index_col=0, parse_dates=True,
squeeze=True)
# + id="VDXOtjYiPTco" outputId="aad79723-ed8c-437e-aca8-7522eca07ab1"
divida.plot()
plt.title('Dívida pública líquida dos governos municipais e estaduais: externa (azul), interna e total (% PIB)', fontsize=14)
plt.savefig('fig_divida')
pyplot.show()
# + id="mgrwYbgFPTco"
| Python/Aula_01_Introducao.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## PADRÃO FAÇADE (FACHADA)
#
# [](https://colab.research.google.com/github/catolicasc-joinville/lp1-notebooks/blob/master/3-padroes-de-projeto/4-facade.ipynb) [launch](https://colab.research.google.com/github/catolicasc-joinville/lp1-notebooks/blob/master/3-padroes-de-projeto/4-factory.ipynb)
#
# Façade é um Padrão de Projeto estrutural. Esses padrões descrevem como os objetos e classes podem ser combinados para compor estruturas maiores. Temos dois principais padrões estruturais que nós já usamos: Classe e Objeto. Classe descreve abstrações com a ajuda de herança e oferece uma interface de programação mais conveniente. O padrão Objecto descreve como objetos podem ser associados e compostos para formar objetos maiores. Padrões estruturais são combinações de Classes e Objetos.
#
# O Fachada (Façade) refere-se à face de uma construção, em especial, uma face atraente. O padrão Façade oculta as complexidades do sistema interno e oferece uma interface ao cliente para que este possa acessar o sistema de forma bem simplificada.
#
# Considere o exemplo de um atendente de loja. Quando você, como cliente, vai a uma do loja para comprar determinados itens, você não tem conhecimento do layout da loja. Geralmente você chama um atendente que conhece bem o sistema da loja. Com base em suas solicitações, o atendente escolhe alguns itens e entrega para você. O cliente não precisa saber como é a loja, ele tem o serviço feito por meio de uma interface, o atendente.
#
# O padrão de projeto Façade faz essencialmente o seguinte:
#
# * Oferece uma interface unificada para um conjunto de interfaces em um subsistema e define uma interface de alto nível que ajuda o cliente a usar o subsistema de forma fácil
# * O Façade procura fazer a representação de um subsistema complexo com um único objeto de interface. Ela não encapsula o subsistema, mas, na verdade, combina os subsistemas subjacentes
# * Promove desacoplamento da implementação com vários clientes
#
# Veja um exemplo de implementação do padrão usando a representação em UML:
#
# 
#
# Como pode-se observar, você perceberá que já 3 tipos de entidades principais no padrão: Façade, Sistema e Cliente.
#
# O **Façade** é responsável por englobar um grupo complexo de subsistemas de modo que a fachada possa oferecer uma aparência agradável ao mundo externo. Ele é uma interface que sabe quais subsistemas são responsáveis por uma requisição e delega essa requisição para os objetos de subsistemas apropriados usando composição.
#
# O **Sistema** representa um conjunto de subsistemas variados que compõem o sistema como um todo, que é difícil de visualizar ou com o qual é complicado trabalhar. Ele implementa funcionalidades de subsistema e é representado por uma classe. O ideal é que um Sistema seja represenado por um grupo de classes responsáveis por diferentes operações.
#
# O **Cliente** interage com o Façade para que ele possaa se comunicar facilmente com o subsistema e ter uma tarefa concluída. O cliente não precisa se preocupar com a natureza complexa do sistema.
#
# #### Exemplo
#
# Para exemplificar a implementação do padrão, suponha que haja um casamento e existe um sistema responsável por toda a organização. No passado, a gente precisaria fazer tudo sozinho, hoje podemos contar com um sistema organizador de casamentos. Este sistema é a "fachada" de organização do casamento, dentro dele, teremos:
#
# * **Cliente** Somos nós, através de uma aplicação, que precisamos da organização do casamento
# * **Façade** É o sistema organizador de casamentos, o `EventManager`
# * **Subsistemas** Representam os sistemas que oferecem os servic'so como bufê, gerenciamento de hotel e decorações
#
# Agora faça o seguinte! A partir o esqueleto de código abaixo, implemente o padrão Façade para o casamento que deve fazer a reserva em um hotel, contratar florista, organizar o buffet e escolher os músicos.
#
# ```python
#
# class Hotelier:
# def isAvailable(self):
# pass
#
# def book(self):
# pass
#
# class Florist:
# def setFlowers(self):
# pass
#
# class Caterer:
# def setCuisine(self):
# pass
#
# class Musician:
# def setMusicType(self):
# pass
#
# class EventManager:
# def arrange(self):
# pass
#
# class CommandClient:
# pass
# ```
# Além de oferecer um sistema unificado que facilita o uso de subsistemas, o padrão Façade facilita o desacoplamento do cliente e dos subsistemas de componentes. O princípio pro trás disso é o "Princípio do Conhecimento Mínimo" ([Law of Demeter](https://en.wikipedia.org/wiki/Law_of_Demeter)), que nos orienta no sentido de reduzir as interações entre os objetos e apenas alguns poucos outros objetos podem interagir com ele. Isso nos ajuda a evitar os casos onde existem classes altamente acopladas com outras. Se existir muitas dependências entre as classes, o sistema será difícil de manter. Quanlquer mudança em uma parte do sistema poderá quebrar outras partes. Em resumo, a Lei de Demeter diz:
#
# * Cada unidade deve ter conhecimento limitado sobre outras unidades: apenas unidades "próximas" se relacionam à unidade atual
# * Cada unidade deve apenas conversar com seus amigos; Não fale com estranhos
# * Apenas fale com seus amigos imediatos
#
# # GANHE PONTOS EXTRAS!
#
# Que ganhar pontos extras com o professor? Tá precisando de nota na matéria? Então traduza os artigos da wikipedia sobre a Lei de Demeter e sobre acoplamento para o português:
#
# * https://en.wikipedia.org/wiki/Law_of_Demeter
# * https://en.wikipedia.org/wiki/Loose_coupling
| python/padroes-de-projeto/facade.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
concrete_data = pd.read_csv('https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/DL0101EN/labs/data/concrete_data.csv')
concrete_data.head()
concrete_data.shape
concrete_data.describe()
concrete_data.isnull().sum()
# +
concrete_data_columns = concrete_data.columns
predictors = concrete_data[concrete_data_columns[concrete_data_columns != 'Strength']] # all columns except Strength
target = concrete_data['Strength'] # Strength column
# -
predictors.head()
target.head()
predictors_norm = (predictors - predictors.mean()) / predictors.std()
predictors_norm.head()
n_cols = predictors_norm.shape[1]
import keras
from keras.models import Sequential
from keras.layers import Dense
# define regression model
def regression_model():
# create model
model = Sequential()
model.add(Dense(50, activation='relu', input_shape=(n_cols,)))
model.add(Dense(50, activation='relu'))
model.add(Dense(50, activation='relu'))
model.add(Dense(50, activation='relu'))
model.add(Dense(1))
# compile model
model.compile(optimizer='adam', loss='mean_squared_error')
return model
model = regression_model()
import sklearn
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(predictors_norm, target, random_state=0)
model.fit(X_train, y_train, epochs=50)
model.predict(X_test)
y_test
# predicted values are close to the true values
# hence my try of regression with Keras is complete...
# by <NAME>
| regression_work_Keras.ipynb |