code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Import packages
# +
# %matplotlib inline
import matplotlib
import matplotlib.pyplot as plt
import os
import sys
import dill
import yaml
import numpy as np
import pandas as pd
import ast
import collections
import seaborn as sns
import matplotlib.ticker as mtick
sns.set(style='ticks')
# -
# ### Import submodular-optimization packages
sys.path.insert(0, "/Users/smnikolakaki/GitHub/submodular-linear-cost-maximization/submodular_optimization/")
# ### Visualizations directory
VIZ_DIR = os.path.abspath("/Users/smnikolakaki/GitHub/submodular-linear-cost-maximization/submodular_optimization/viz/")
# ### Plotting utilities
def set_style():
# This sets reasonable defaults for font size for a paper
sns.set_context("paper")
# Set the font to be serif
sns.set(font='serif')#, rc={'text.usetex' : True})
# Make the background white, and specify the specific font family
sns.set_style("white", {
"font.family": "serif",
"font.serif": ["Times", "Palatino", "serif"]
})
# Set tick size for axes
sns.set_style("ticks", {"xtick.major.size": 6, "ytick.major.size": 6})
def set_size(fig, width=6, height=4):
fig.set_size_inches(width, height)
plt.tight_layout()
def save_fig(fig, filename):
fig.savefig(os.path.join(VIZ_DIR, filename), dpi=600, format='pdf', bbox_inches='tight')
# #### Details
# Original marginal gain: $$g(e|S) = f(e|S) - w(e)$$
# Scaled marginal gain: $$\tilde{g}(e|S) = f(e|S) - 2w(e)$$
#
# #### Algorithms:
# 1. Greedy: The algorithm performs iterations i = 0,...,n-1. In each iteration the algorithm selects the element that maximizes the original marginal gain. It adds the element to the solution if the original marginal gain of the element is >= 0. It updates the set of valid elements by keeping only those elements that when added to the solution preserve that it belongs to the independent set. The running time is O($n^2$).
#
# 2. Cost Greedy (Algorithm 1 from arXiv): The algorithm performs iterations i = 0,...,n-1. In each iteration the algorithm selects the element that maximizes the scaled marginal gain. It adds the element to the solution if the original marginal gain of the element is >= 0. It updates the set of valid elements by keeping only those elements that when added to the solution preserve that it belongs to the independent set. The running time is O($n^2$).
#
#
# 3. Cost Lazy Greedy (Algorithm 1 from arXiv with lazy eval): The algorithm first initializes a max heap with all the elements. The key of each element is its scaled marginal gain and the value is the element id. If the original marginal gain of an element is < 0 the algorithm discards the element and never inserts it in the heap. Then, for 0,...,n-1 iterations the algorithm does the following: (i) pops the top element from the heap and computes its new scaled marginal gain, (ii) It checks the old scaled marginal gain of the next element in the heap, (iii) if the popped element's new scaled marginal gain is >= the next elements's old gain we return the popped element, otherwise if its new original marginal gain is >= 0 we reinsert the element to the heap and repeat step iii. If the returned element's original marginal gain is >= 0 we add it to the solution and update the set of valid elements. The algorithm returns a solution S: f(S) - w(S) >= (1/2)f(OPT) - w(OPT). The running time in the worst case is O($n^2$).
#
# 4. Cost Lazy Scaled Greedy: Same as above. The only thing that changes is step (iii) where we reinsert an element to the heap if its scaled marginal gain is >= 0 (instead of the original gain).
#
# +
legends = {
"partition_matroid_greedy":"Greedy",
"cost_scaled_partition_matroid_lazy_greedy":"MCSLG",
"baseline_topk_matroid": 'Top-k-Experts-Matroid'
}
legends = collections.OrderedDict(sorted(legends.items()))
line_styles = {'partition_matroid_greedy':"--",
'cost_scaled_partition_matroid_lazy_greedy':'-',
'baseline_topk_matroid':'--'}
line_styles = collections.OrderedDict(sorted(line_styles.items()))
marker_style = {'partition_matroid_greedy':"h",
'cost_scaled_partition_matroid_lazy_greedy':'x',
'baseline_topk_matroid':'d'}
marker_style = collections.OrderedDict(sorted(marker_style.items()))
marker_size = {'partition_matroid_greedy':30,
'cost_scaled_partition_matroid_lazy_greedy':30,
'baseline_topk_matroid':22}
marker_size = collections.OrderedDict(sorted(marker_size.items()))
marker_edge_width = {'partition_matroid_greedy':6,
'cost_scaled_partition_matroid_lazy_greedy':10,
'baseline_topk_matroid':6}
marker_edge_width = collections.OrderedDict(sorted(marker_edge_width.items()))
line_width = {'partition_matroid_greedy':5,
'cost_scaled_partition_matroid_lazy_greedy':5,
'baseline_topk_matroid':5}
line_width = collections.OrderedDict(sorted(line_width.items()))
name_objective = "Combined objective (g)"
fontsize = 53
legendsize = 42
labelsize = 53
x_size = 20
y_size = 16
# -
# #### Performance comparison for salary partitions
# The experimental setting for the salary partitions is the following:
# 1. Get a sample of users
# 2. Create the ordering of the users' unique salary values. Divide the sorted salaries into 20 partitions.
# 3. Assign each user to her corresponding cost partition range.
# 3. From each partition the solution can select only one user in this setting.
def plot_performance_comparison(df):
palette = sns.color_palette(['#b30000','#dd8452', '#ccb974', '#4c72b0', '#8172b3',
'#55a868',
'#8172b3', '#937860', '#da8bc3', '#8c8c8c',
'#ccb974', '#64b5cd'],3)
ax = sns.lineplot(x='cardinality_constraint', y='val', data=df,
hue='Algorithm', ci='sd',
mfc='none',palette=palette)
i = 0
for key, val in line_styles.items():
ax.lines[i].set_linestyle(val)
# ax.lines[i].set_color(colors[key])
ax.lines[i].set_linewidth(line_width[key])
ax.lines[i].set_marker(marker_style[key])
ax.lines[i].set_markersize(marker_size[key])
ax.lines[i].set_markeredgewidth(marker_edge_width[key])
ax.lines[i].set_markeredgecolor(None)
i += 1
plt.yticks(np.arange(0, 45000, 5000))
# plt.xticks(np.arange(0, 1.1, 0.1))
plt.xlabel('Constraint (k)', fontsize=fontsize)
plt.ylabel(name_objective, fontsize=fontsize)
# plt.title('Performance comparison')
fig = plt.gcf()
figlegend = plt.legend([val for key,val in legends.items()],loc=3, bbox_to_anchor=(0., 1.02, 1., .102),
ncol=2, mode="expand", borderaxespad=0., frameon=False,prop={'size': legendsize})
ax = plt.gca()
plt.gca().tick_params(axis='y', labelsize=labelsize)
plt.gca().tick_params(axis='x', labelsize=labelsize)
return fig, ax
# +
df = pd.read_csv("/Users/smnikolakaki/GitHub/submodular-linear-cost-maximization/jupyter/experiment_04_guru_salary_pop08_rare01_final.csv",
header=0,
index_col=False)
df.columns = ['Algorithm', 'sol', 'val', 'submodular_val', 'cost', 'runtime', 'lazy_epsilon',
'sample_epsilon','user_sample_ratio','scaling_factor','num_rare_skills','num_common_skills',
'num_popular_skills','num_sampled_skills','seed','k','cardinality_constraint','num_of_partitions']
df = df[(df.Algorithm == 'partition_matroid_greedy')
|(df.Algorithm == 'cost_scaled_partition_matroid_lazy_greedy')
|(df.Algorithm == 'baseline_topk_matroid')
]
df0 = df[(df['num_of_partitions'] == 5)]
df0.sort_values(by ='Algorithm',inplace=True)
set_style()
fig, axes = plot_performance_comparison(df0)
set_size(fig, x_size, y_size)
save_fig(fig,'score_partition_guru_salary_pop08_rare01.pdf')
# -
# #### Running time comparison for salary partitions
# +
legends = {
"partition_matroid_greedy":"Greedy",
"cost_scaled_partition_matroid_lazy_greedy":"MCSLG",
"baseline_topk_matroid": 'Top-k-Experts-Matroid',
"cost_scaled_partition_matroid_greedy":"MCSG"
}
legends = collections.OrderedDict(sorted(legends.items()))
line_styles = {'partition_matroid_greedy':':',
'cost_scaled_partition_matroid_lazy_greedy':'-',
'baseline_topk_matroid':'--',
'cost_scaled_partition_matroid_greedy':"-"}
line_styles = collections.OrderedDict(sorted(line_styles.items()))
marker_style = {'partition_matroid_greedy':'h',
'cost_scaled_partition_matroid_lazy_greedy':'x',
'baseline_topk_matroid':'d',
"cost_scaled_partition_matroid_greedy":"x"}
marker_style = collections.OrderedDict(sorted(marker_style.items()))
marker_size = {'partition_matroid_greedy':25,
'cost_scaled_partition_matroid_lazy_greedy':30,
'baseline_topk_matroid':22,
"cost_scaled_partition_matroid_greedy":30}
marker_size = collections.OrderedDict(sorted(marker_size.items()))
marker_edge_width = {'partition_matroid_greedy':6,
'cost_scaled_partition_matroid_lazy_greedy':10,
'baseline_topk_matroid':6,
"cost_scaled_partition_matroid_greedy":10}
marker_edge_width = collections.OrderedDict(sorted(marker_edge_width.items()))
line_width = {'partition_matroid_greedy':5,
'cost_scaled_partition_matroid_lazy_greedy':5,
'baseline_topk_matroid':5,
"cost_scaled_partition_matroid_greedy":5}
line_width = collections.OrderedDict(sorted(line_width.items()))
name_objective = "Combined objective (g)"
fontsize = 53
legendsize = 42
labelsize = 53
x_size = 20
y_size = 16
# -
def plot_performance_comparison(df):
palette = sns.color_palette(['#b30000','#937860','#dd8452','#ccb974','#4c72b0'
'#55a868',
'#8172b3', '#937860', '#da8bc3', '#8c8c8c',
'#ccb974', '#64b5cd'],4)
ax = sns.lineplot(x='cardinality_constraint', y='runtime', data=df,
hue='Algorithm', ci='sd',
mfc='none',palette=palette)
i = 0
for key, val in line_styles.items():
ax.lines[i].set_linestyle(val)
# ax.lines[i].set_color(colors[key])
ax.lines[i].set_linewidth(line_width[key])
ax.lines[i].set_marker(marker_style[key])
ax.lines[i].set_markersize(marker_size[key])
ax.lines[i].set_markeredgewidth(marker_edge_width[key])
ax.lines[i].set_markeredgecolor(None)
i += 1
# plt.yticks(np.arange(0, 45000, 5000))
# plt.xticks(np.arange(0, 1.1, 0.1))
plt.ylabel('Time (sec)', fontsize=fontsize)
plt.xlabel('Constraint (k)', fontsize=fontsize)
# plt.title('Performance comparison')
fig = plt.gcf()
figlegend = plt.legend([val for key,val in legends.items()],loc=3, bbox_to_anchor=(0., 1.02, 1., .102),
ncol=2, mode="expand", borderaxespad=0., frameon=False,prop={'size': legendsize})
plt.gca().tick_params(axis='y', labelsize=labelsize)
plt.gca().tick_params(axis='x', labelsize=labelsize)
a = plt.axes([.17, .53, .35, .3])
ax2 = sns.lineplot(x='cardinality_constraint', y='runtime', data=df,
hue='Algorithm', legend=False,
mfc='none',palette=palette,label=False)
i = 0
for key, val in line_styles.items():
ax2.lines[i].set_linestyle(val)
# ax.lines[i].set_color(colors[key])
ax2.lines[i].set_linewidth(2)
ax2.lines[i].set_marker(marker_style[key])
ax2.lines[i].set_markersize(12)
ax2.lines[i].set_markeredgewidth(3)
ax2.lines[i].set_markeredgecolor(None)
i += 1
ax2.set(ylim=(0, 3))
ax2.set(xlim=(0, 10.5))
ax2.set_ylabel('')
ax2.set_xlabel('')
# plt.gca().xaxis.set_major_formatter(mtick.FormatStrFormatter('%.1e'))
# plt.gca().yaxis.set_major_formatter(mtick.FormatStrFormatter('%.1e'))
plt.gca().tick_params(axis='x', labelsize=22)
plt.gca().tick_params(axis='y', labelsize=22)
plt.tight_layout()
return fig, ax
# +
df = pd.read_csv("/Users/smnikolakaki/GitHub/submodular-linear-cost-maximization/jupyter/experiment_04_guru_salary_pop08_rare01_final.csv",
header=0,
index_col=False)
df.columns = ['Algorithm', 'sol', 'val', 'submodular_val', 'cost', 'runtime', 'lazy_epsilon',
'sample_epsilon','user_sample_ratio','scaling_factor','num_rare_skills','num_common_skills',
'num_popular_skills','num_sampled_skills','seed','k','cardinality_constraint','num_of_partitions']
df = df[(df.Algorithm == 'partition_matroid_greedy')
|(df.Algorithm == 'cost_scaled_partition_matroid_lazy_greedy')
|(df.Algorithm == 'baseline_topk_matroid')
|(df.Algorithm == 'cost_scaled_partition_matroid_greedy')
]
df0 = df[(df['num_of_partitions'] == 5)]
df0.sort_values(by ='Algorithm',inplace=True)
set_style()
fig, axes = plot_performance_comparison(df0)
set_size(fig, x_size, y_size)
save_fig(fig,'time_partition_guru_salary_pop08_rare01.pdf')
# -
| jupyter/.ipynb_checkpoints/Experiment_partition_matroid_problem_guru_pop08_rare01-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
# # Unsupervised Learning
#
# Many instances of unsupervised learning, such as dimensionality reduction, manifold learning and feature extraction, find a new representation of the input data without any additional input.
#
# <img src="figures/unsupervised_workflow.svg" width="100%">
#
# The most simple example of this, which can barely be called learning, is rescaling the data to have zero mean and unit variance. This is a helpful preprocessing step for many machine learning models.
#
# Applying such a preprocessing has a very similar interface to the supervised learning algorithms we saw so far.
# Let's load the iris dataset and rescale it:
# +
from sklearn.datasets import load_iris
iris = load_iris()
X, y = iris.data, iris.target
print(X.shape)
# -
# The iris dataset is not "centered" that is it has non-zero mean and the standard deviation is different for each component:
#
print("mean : %s " % X.mean(axis=0))
print("standard deviation : %s " % X.std(axis=0))
# To use a preprocessing method, we first import the estimator, here StandardScaler and instantiate it:
#
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
# As with the classification and regression algorithms, we call ``fit`` to learn the model from the data. As this is an unsupervised model, we only pass ``X``, not ``y``. This simply estimates mean and standard deviation.
scaler.fit(X)
# Now we can rescale our data by applying the ``transform`` (not ``predict``) method:
X_scaled = scaler.transform(X)
# ``X_scaled`` has the same number of samples and features, but the mean was subtracted and all features were scaled to have unit standard deviation:
print(X_scaled.shape)
print("mean : %s " % X_scaled.mean(axis=0))
print("standard deviation : %s " % X_scaled.std(axis=0))
# Principal Component Analysis
# ============================
# An unsupervised transformation that is somewhat more interesting is Principle Component Analysis (PCA).
# It is a technique to reduce the dimensionality of the data, by creating a linear projection.
# That is, we find new features to represent the data that are a linear combination of the old data (i.e. we rotate it).
#
# The way PCA finds these new directions is by looking for the directions of maximum variance.
# Usually only few components that explain most of the variance in the data are kept. To illustrate how a rotation might look like, we first show it on two dimensional data and keep both principal components.
#
# We create a Gaussian blob that is rotated:
rnd = np.random.RandomState(5)
X_ = rnd.normal(size=(300, 2))
X_blob = np.dot(X_, rnd.normal(size=(2, 2))) + rnd.normal(size=2)
y = X_[:, 0] > 0
plt.scatter(X_blob[:, 0], X_blob[:, 1], c=y, linewidths=0, s=30)
plt.xlabel("feature 1")
plt.ylabel("feature 2")
# As always, we instantiate our PCA model. By default all directions are kept.
from sklearn.decomposition import PCA
pca = PCA()
# Then we fit the PCA model with our data. As PCA is an unsupervised algorithm, there is no output ``y``.
pca.fit(X_blob)
# Then we can transform the data, projected on the principal components:
# +
X_pca = pca.transform(X_blob)
plt.scatter(X_pca[:, 0], X_pca[:, 1], c=y, linewidths=0, s=30)
plt.xlabel("first principal component")
plt.ylabel("second principal component")
# -
# On the left of the plot you can see the four points that were on the top right before. PCA found fit first component to be along the diagonal, and the second to be perpendicular to it. As PCA finds a rotation, the principal components are always at right angles to each other.
# Dimensionality Reduction for Visualization with PCA
# -------------------------------------------------------------
# Consider the digits dataset. It cannot be visualized in a single 2D plot, as it has 64 features. We are going to extract 2 dimensions to visualize it in, using the example from the sklearn examples [here](http://scikit-learn.org/stable/auto_examples/manifold/plot_lle_digits.html)
# +
from figures.plot_digits_datasets import digits_plot
digits_plot()
# -
# Note that this projection was determined *without* any information about the
# labels (represented by the colors): this is the sense in which the learning
# is **unsupervised**. Nevertheless, we see that the projection gives us insight
# into the distribution of the different digits in parameter space.
# ## Manifold Learning
#
# One weakness of PCA is that it cannot detect non-linear features. A set
# of algorithms known as *Manifold Learning* have been developed to address
# this deficiency. A canonical dataset used in Manifold learning is the
# *S-curve*, which we briefly saw in an earlier section:
# +
from sklearn.datasets import make_s_curve
X, y = make_s_curve(n_samples=1000)
from mpl_toolkits.mplot3d import Axes3D
ax = plt.axes(projection='3d')
ax.scatter3D(X[:, 0], X[:, 1], X[:, 2], c=y)
ax.view_init(10, -60)
# -
# This is a 2-dimensional dataset embedded in three dimensions, but it is embedded
# in such a way that PCA cannot discover the underlying data orientation:
X_pca = PCA(n_components=2).fit_transform(X)
plt.scatter(X_pca[:, 0], X_pca[:, 1], c=y)
# Manifold learning algorithms, however, available in the ``sklearn.manifold``
# submodule, are able to recover the underlying 2-dimensional manifold:
# +
from sklearn.manifold import Isomap
iso = Isomap(n_neighbors=15, n_components=2)
X_iso = iso.fit_transform(X)
plt.scatter(X_iso[:, 0], X_iso[:, 1], c=y)
# -
# ##Exercise
# Compare the results of Isomap and PCA on a 5-class subset of the digits dataset (``load_digits(5)``).
#
# __Bonus__: Also compare to TSNE, another popular manifold learning technique.
# +
from sklearn.datasets import load_digits
digits = load_digits(5)
X = digits.data
# ...
| notebooks/02.3 Unsupervised Learning - Transformations and Dimensionality Reduction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import numpy as np
from environment import environment
env = environment(base=3, random_state=211)
#env.create_environment()
env.print_board(env.current_state)
print(env.current_state)
print(env.next_state)
# -
env.single_play((0,1,'O'), env.next_state)
env.print_board(env.current_state)
print(' ')
env.print_board(env.next_state)
print(env.next_state)
env.reset()
# +
from agent_monte_carlo import agent
env = environment(known_num=40, random_state=211)
env.create_environment()
for i in range(300):
player = agent(environment=env, random_state=None)
r = player.play_game()
player.environment.print_board(env.next_state)
# +
from agent_monte_carlo import agent
#env = environment(known_num=40, random_state=211)
#env.create_environment()
gamma = 1
#for i in range(3000):
# player = agent(environment=env, random_state=None)
# player.play_game()
total_reward = []
for i in range(50000):
count = 0
boolean = True
env = environment(known_num=40, random_state=211)
env.create_environment()
sr = 0
k = 0
while boolean:
player = agent(environment=env, random_state=None)
r = player.play_game()
if player.game_over or player.win:
boolean = False
count += 1
sr += (gamma ** k) * r
k += 1
total_reward.append(sr)
#print(sr)
if player.win:
print(count, player.win)
#player.environment.print_board(env.next_state)
#print(count, sr, player.win, player.game_over)
player.environment.print_board(env.next_state)
# +
import matplotlib.pyplot as plt
plt.plot(total_reward,'.')
# -
len(set([1,1,1]))
| TicTacToe/Sudoku.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/fxnnxc/Movie_Sentiment_Classification/blob/master/Vader_sentianalysis.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="3TSyIipPHgON" colab_type="code" outputId="84b09573-b646-4089-d79d-334a0cacd4d1" colab={"base_uri": "https://localhost:8080/", "height": 35}
import nltk
nltk.downloader.download('vader_lexicon')
import re
from nltk.sentiment.vader import SentimentIntensityAnalyzer
import numpy as np
def make_sent_list(trans_list):
sent_list = []
senti_analyzer = SentimentIntensityAnalyzer()
for i in trans_list:
score=senti_analyzer.polarity_scores(i)['compound']
sent_list.append(score)
return sent_list
# + id="xXN7qNTiHlDw" colab_type="code" colab={}
sentences ="""
Do you always have dinner at a different time, too?
No. I almost always eat dinner around 6 o'clock. How about you? When do you eat dinner?
My dinner time varies a lot just like my breakfast. So maybe I eat dinner later than most people. I think I would have dinner at 7:00 PM at the earliest. But sometimes, I will have dinner as late as 9 o'clock at night.
Really?
Yeah. I don't mind it. I really like cooking so, if I'm cooking a short thing, I'll have dinner earlier. But I don't mind if it takes two hours to cook something really good, I'll eat later at night.
Wow. Can you tell me about your work routine? What time do you usually go to work?
See, my work schedule is different everyday. That's why I wake up at a different time everyday. Maybe on – for example, some weeks on Monday, Wednesday and Thursday, I start work at 7:00. But on Thursday and Friday, I don't start work until 11:30.
Ah.
How about you? Do you have a different work time sometimes?
Well, right now, I'm on maternity leave. So I stay home and take care of my new baby.
Oh, congratulations.
Thank you. I try to work at home a little bit everyday though. If the kids are sleeping or they're playing quietly, I try to do some work. Maybe around 2 o'clock, I can usually get some work done because the kids are sleeping.
Oh, if you don't start getting some work done until 2 o'clock in the afternoon, you must be very busy every morning.
Yeah. I usually go to the grocery store. Sometimes I take the kids to the park around 10:00 in the morning. I often do the laundry or wash the dishes, and then as soon as the dishes are washed, it's time to make lunch. So I'm busy all day.
I see. So that's why your lunch time can change so much.
Yeah.
Oh, it sounds like a busy day.
You are so bad
killing is not good
I love you
""".strip().split('\n')
# + id="EIBIVdA3IiRi" colab_type="code" outputId="c43a2a6f-fa03-46be-98f2-6b08b220f429" colab={"base_uri": "https://localhost:8080/", "height": 669}
import pandas as pd
pd.set_option("display.max_colwidth", 100)
pd.DataFrame({'sentence':sentences, 'score':make_sent_list(sentences)})
# + id="G9PPYaXVJtf0" colab_type="code" colab={}
| colab scripts/Vader_sentianalysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5"
import json
import os
from pathlib import Path
import time
import copy
import numpy as np
import pandas as pd
import torch
from torch import nn, optim
from torch.utils.data import Dataset, DataLoader
from torchvision import models
from fastai.dataset import open_image
import json
from PIL import ImageDraw, ImageFont
import matplotlib.pyplot as plt
from matplotlib import patches, patheffects
import cv2
from tqdm import tqdm
# + _uuid="118bf972c981d93118fe3b203b95d9094b85643e"
SIZE = 224
IMAGES = 'images'
ANNOTATIONS = 'annotations'
CATEGORIES = 'categories'
ID = 'id'
NAME = 'name'
IMAGE_ID = 'image_id'
BBOX = 'bbox'
CATEGORY_ID = 'category_id'
FILE_NAME = 'file_name'
# -
# !pwd
# !ls $HOME/data/pascal
# + _uuid="8c132fd434c2843a1ad894aa36665272901d74f1"
# !ls ../input/pascal/pascal
# + _uuid="3f0c759a56e66938c6739e49811c39275f5e1d3c"
PATH = Path('/home/paperspace/data/pascal')
list(PATH.iterdir())
# + _uuid="73c706f2cf75e5859ba28933ca3cb5b6ca21d9a8"
train_data = json.load((PATH/'pascal_train2007.json').open())
val_data = json.load((PATH/'pascal_val2007.json').open())
test_data = json.load((PATH/'pascal_test2007.json').open())
print('train:', train_data.keys())
print('val:', val_data.keys())
print('test:', test_data.keys())
# + _uuid="ef583f5efd5e4adf8093ca1f3dcf96febc4a2adf"
train_data[ANNOTATIONS][:1]
# + _uuid="9a314f39827bd1e068ee355679491feb828137f1"
train_data[IMAGES][:2]
# + _uuid="f4a3cdc3f414c19e16cf66aabdbd43aca7fc3fda"
len(train_data[CATEGORIES])
# + _uuid="3f18ad0f37bdc398a5b1c56a7ab8fe8e93dc5393"
next(iter(train_data[CATEGORIES]))
# -
# ## Categories - 1th indexed
# + _uuid="d14164141afed91ff611899627d1c52f0e6fddc5"
categories = {c[ID]:c[NAME] for c in train_data[CATEGORIES]}
categories
# + _uuid="35f1eab3f886a09f36e1658dddceb0f20d1f2d00"
len(categories)
# + _uuid="6820463abe2e61a1bc7abf40c74d43b3084e3d63"
IMAGE_PATH = Path(PATH/'JPEGImages/')
list(IMAGE_PATH.iterdir())[:2]
# + _uuid="cf9938b6fb1ca4d717531172025ef3068a571e89"
train_filenames = {o[ID]:o[FILE_NAME] for o in train_data[IMAGES]}
print('length:', len(train_filenames))
image1_id, image1_fn = next(iter(train_filenames.items()))
image1_id, image1_fn
# + _uuid="7c7958c1b7c228d3558308641306516cd4560ab4"
train_image_ids = [o[ID] for o in train_data[IMAGES]]
print('length:', len(train_image_ids))
train_image_ids[:5]
# + _uuid="d8a744ba6d5455454d8c0ed5537e868c8f1262b8"
IMAGE_PATH
# + _uuid="fcf7690daaf38bb77739606b77b9f0b0fe291b87"
image1_path = IMAGE_PATH/image1_fn
image1_path
# + _uuid="79c3b1f68813ba869ad489c2806bb4dd1f1411d4"
str(image1_path)
# + _uuid="76de823201eae04a02faf97c98b3084401d4469e"
im = open_image(str(IMAGE_PATH/image1_fn))
print(type(im))
# + _uuid="2d5c0a08b490fb6a5985349972e260f362643bf1"
im.shape
# + _uuid="f465a259b4afa143f445c89e3c4b80c009c31224"
len(train_data[ANNOTATIONS])
# + _uuid="1fd225ea5f55b210b885cfbc549d010ce9a64f2f"
# get the biggest object label per image
# + _uuid="786f2c45d422151696a9ab2c344cd3b778e8a36c"
train_data[ANNOTATIONS][0]
# + _uuid="098571a72d31ba5b09c0b9edfd858171b1568eb0"
bbox = train_data[ANNOTATIONS][0][BBOX]
bbox
# + _uuid="08656dd9054e753b304690af1727afecf3a1cfd9"
def fastai_bb(bb):
return np.array([bb[1], bb[0], bb[3]+bb[1]-1, bb[2]+bb[0]-1])
print(bbox)
print(fastai_bb(bbox))
# + _uuid="b8cdddedeca3f539403aa1e28ebb8b9a81a735e7"
fbb = fastai_bb(bbox)
fbb
# + _uuid="1ce11890ef052df4eddc2b2ae7c68ffa69151881"
def fastai_bb_hw(bb):
h= bb[3]-bb[1]+1
w = bb[2]-bb[0]+1
return [h,w]
fastai_bb_hw(fbb)
# + _uuid="c0452e7a5f625fb0b5947f0fea21cf6ae6e00d19"
def pascal_bb_hw(bb):
return bb[2:]
pascal_bb_hw(bbox)
# + _uuid="2fd4a8a9d6cfbd964501993d1812ab1a7e0c600f"
train_image_w_area = {i:None for i in train_image_ids}
print(image1_id, train_image_w_area[image1_id])
# + _uuid="177ed24133426328f89e9e1a49150a0181a6358c"
for x in train_data[ANNOTATIONS]:
bbox = x[BBOX]
new_category_id = x[CATEGORY_ID]
image_id = x[IMAGE_ID]
h, w = pascal_bb_hw(bbox)
new_area = h*w
cat_id_area = train_image_w_area[image_id]
if not cat_id_area:
train_image_w_area[image_id] = (new_category_id, new_area)
else:
category_id, area = cat_id_area
if new_area > area:
train_image_w_area[image_id] = (new_category_id, new_area)
# + _uuid="7e2446560a85a0548759818de474a0df8b6f3349"
train_image_w_area[image1_id]
# + _uuid="58f7d2e4f6a1a751d09ac798baba6ac1349f90ea"
plt.imshow(im)
# + _uuid="116b99671bd3cef3900ae72cda80af98172281dd"
def show_img(im, figsize=None, ax=None):
if not ax:
fig,ax = plt.subplots(figsize=figsize)
ax.imshow(im)
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
return ax
show_img(im)
# + _uuid="b8080309513f671e708f1e4ea8cba25bc10a6745"
# show_img(im)
# b = bb_hw(im0_a[0])
# draw_rect(ax, b)
image1_fn
# + _uuid="c161a064474370841fed827793c2717c33ee7447"
def draw_rect(ax, b):
patch = ax.add_patch(patches.Rectangle(b[:2], *b[-2:], fill=False, edgecolor='white', lw=2))
draw_outline(patch, 4)
# + _uuid="cf021e31ea6e290ec39b41b034e66b6f207407a3"
image1_id
# + _uuid="262d753f2c11c7dac7533b05025d2ec19c15f37f"
image1_path
# + _uuid="ec08f6416f18375f8a7fad243a69b24a2a3f3782"
plt.imshow(open_image(str(image1_path)))
# + _uuid="24d41b670341994c3fff5463e34fd8a79c33b1ec"
train_data[ANNOTATIONS][0]
# + _uuid="296d6fd07221ebca30bd4be49febeb48fc75fdee"
im = open_image(str(image1_path))
ax = show_img(im)
# + _uuid="534a61df556b06d86dbf374c762950db3b0ded42"
def draw_outline(o, lw):
o.set_path_effects([patheffects.Stroke(
linewidth=lw, foreground='black'), patheffects.Normal()])
# + _uuid="c70c53d8b2c4b475af25026f623367aad118ee9f"
image1_ann = train_data[ANNOTATIONS][0]
b = fastai_bb(image1_ann[BBOX])
b
# + _uuid="30f7096c698701199700f19b27eaab4fc2b779a2"
def draw_text(ax, xy, txt, sz=14):
text = ax.text(*xy, txt,
verticalalignment='top', color='white', fontsize=sz, weight='bold')
draw_outline(text, 1)
# + _uuid="58f8b738c13533067884c56fa48c4a0d5d21430b"
ax = show_img(im)
b = image1_ann[BBOX]
print(b)
draw_rect(ax, b)
draw_text(ax, b[:2], categories[image1_ann[CATEGORY_ID]])
# + _uuid="db13646d8655ff32ec8ee684fd01fddb3cac5023"
# create a Pandas dataframe for: image_id, filename, category
BIGGEST_OBJECT_CSV = '../input/pascal/pascal/tmp/biggest-object.csv'
IMAGE = 'image'
CATEGORY = 'category'
train_df = pd.DataFrame({
IMAGE_ID: image_id,
IMAGE: str(IMAGE_PATH/image_fn),
CATEGORY: train_image_w_area[image_id][0]
} for image_id, image_fn in train_filenames.items())
train_df.head()
# + _uuid="066b6c2995117c09300a4621711646c103b20e3c"
# NOTE: won't work in Kaggle Kernal b/c read-only file system
# train_df.to_csv(BIGGEST_OBJECT_CSV, index=False)
# + _uuid="61250190d6ec20d07ec17784e8cd1ad00f0075ed"
train_df.iloc[0]
# + _uuid="e48d246e8d361e484f451a05d20bd52ef7c1e50f"
len(train_df)
# + _uuid="eb55adfbedb20416d4f3e07dcfb0536118fb8b07"
class BiggestObjectDataset(Dataset):
def __init__(self, df):
self.df = df
def __len__(self):
return len(self.df)
def __getitem__(self, idx):
im = open_image(self.df.iloc[idx][IMAGE]) # HW
resized_image = cv2.resize(im, (SIZE, SIZE)) # HW
image = np.transpose(resized_image, (2, 0, 1)) # CHW
category = self.df.iloc[idx][CATEGORY]
return image, category
dataset = BiggestObjectDataset(train_df)
inputs, label = dataset[0]
# + _uuid="9f8da61548d0a2761086ff6d5c435c59458e01d8"
label
# + _uuid="2a0dc536d68e374045cd5cd0bfc0946320b0d018"
inputs.shape
# + _uuid="d33bd294f773946be7ade97f64385e28708897c3"
hwc_image = np.transpose(inputs, (1, 2, 0))
plt.imshow(hwc_image)
# + [markdown] _uuid="d75f6c02fc275340aeaf75465da92ec2f12bca8e"
# # DataLoader
# + _uuid="4832dc8acf67d3a16edaca67aabb76f0f0f5c2cc"
BATCH_SIZE = 64
NUM_WORKERS = 4
dataloader = DataLoader(dataset, batch_size=BATCH_SIZE,
shuffle=True, num_workers=NUM_WORKERS)
batch_inputs, batch_labels = next(iter(dataloader))
# + _uuid="e927c3a2a5d6ebf5dcacad899471fa396118a0ee"
batch_inputs.size()
# + _uuid="82a9ba8db25bfb14ea785e12b1aaa76f0bad8a18"
batch_labels
# + _uuid="9d7898260ad5613212d5c8fa13c1328108a8511f"
np_batch_inputs = batch_inputs.numpy()
i = np.random.randint(0,20)
print(categories[batch_labels[i].item()])
chw_image = np_batch_inputs[i]
print(chw_image.shape)
hwc_image = np.transpose(chw_image, (1, 2, 0))
plt.imshow(hwc_image)
# + _uuid="cd0f59df9865ed0bb5a58bf904f899f13948662f"
NUM_CATEGORIES = len(categories)
NUM_CATEGORIES
# -
# ## train the model
# + _uuid="9046792f1d8e1e7adc00d3ad1dfc9d13fbbd66ef"
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
# device = torch.device('cpu')
print('device:', device)
model_ft = models.resnet18(pretrained=True)
# freeze pretrained model
for layer in model_ft.parameters():
layer.requires_grad = False
num_ftrs = model_ft.fc.in_features
print('final layer in/out:', num_ftrs, NUM_CATEGORIES)
model_ft.fc = nn.Linear(num_ftrs, NUM_CATEGORIES)
model_ft = model_ft.to(device)
criterion = nn.CrossEntropyLoss()
# Observe that all parameters are being optimized
optimizer = optim.SGD(model_ft.parameters(), lr=0.01, momentum=0.9)
# + _uuid="a03afc1a50a6ca92ed8602552c9e527665e60928"
EPOCHS = 10
epoch_losses = []
epoch_accuracies = []
for epoch in range(EPOCHS):
print('epoch:', epoch)
running_loss = 0.0
running_correct = 0
for inputs, labels in tqdm(dataloader):
inputs = inputs.to(device)
labels = labels.to(device)
# clear gradients
optimizer.zero_grad()
# forward pass
outputs = model_ft(inputs)
_, preds = torch.max(outputs, dim=1)
labels_0_indexed = labels-1
loss = criterion(outputs, labels_0_indexed)
# backwards pass
loss.backward()
optimizer.step()
# step stats
running_loss += loss.item() * inputs.size(0)
running_correct += torch.sum(preds == labels_0_indexed)
# epoch stats
epoch_loss = running_loss / len(dataset)
epoch_acc = running_correct.double().item() / len(dataset)
epoch_losses.append(epoch_loss)
epoch_accuracies.append(epoch_acc)
print('loss:', epoch_loss, 'acc:', epoch_acc)
# + _uuid="f64c8664a6f39865d8d4e5f9ee81131beea09d18"
epoch_losses
# + _uuid="688e093bf7e97c6de853d23934c0cff927f99ee3"
epoch_accuracies
# + _uuid="180a093a6d69e456d65124a424c0013e3168de24"
plt.plot(epoch_losses)
# + _uuid="b116d1d203132fa94e75e0fab51e7c0796b45cee"
plt.plot(epoch_accuracies)
# + _uuid="42bddd466cfef27b0c74cc8f7bdbe7fe58029c40"
| notebooks/pascal-fastai-object-classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # dataprep.py example
#
# This notebook will show how the functions contained within the `dataprep.py` module are used to generate hdf5 files for storing raw data and peak fitting results. This module is also the primary way in which the functions contained within spectrafit.py are utilized.
import os
import h5py
import matplotlib.pyplot as plt
from ramandecompy import dataprep
from ramandecompy import datavis
# ### dataprep.new_hdf5
#
# The first function in the module simply generates a new `.hdf5` file in your active directory. The only input required is the desired name of the file. Typically, a user will want to generate two files: a `calibration.hdf5` and an `experiment,hdf5`. If a .hdf5 file with that name already exists, an error will be thrown.
dataprep.new_hdf5('dataprep_calibration')
# ### dataprep.view_hdf5
#
# The module contains a function (`dataprep.view_hdf5`) that will help display the groups and dataset contained within the `.hdf5` file. At this point, `dataprep_calibration.hdf5` is empty so only the filename is output. `dataprep.view_hdf5` displays groups in **bold** and datasets in a standard font.
dataprep.view_hdf5('dataprep_calibration.hdf5')
# ### dataprep. add_calibration
#
# There are two functions for adding data to a .hdf5 file. The first is `dataprep.add_calibration` and is used to add calibration data to a `calibration.hdf5`
dataprep.add_calibration('dataprep_calibration.hdf5',
'../ramandecompy/tests/test_files/Hydrogen_Baseline_Calibration.xlsx',
label='Hydrogen')
dataprep.view_hdf5('dataprep_calibration.hdf5')
# Now using `dataprep.view_hdf5` we can see that our `dataprep_calibration_hdf5` file now contains one 1st order group named with our assigned label. This group contains 6 datasets. The first four datasets consist of of the six fit parameters defining the pseudo-Voigt profiles of each detected peak, along with a 7th value coresponding to the area under the curve of the pseudo-Voigt profile. The last two datasets are the raw x (wavenumber) and y (counts) values from the calibration spectra.
#
# Next we will add one more set of calibration data to `dataprep_calibration.hdf5`.
dataprep.add_calibration('dataprep_calibration.hdf5',
'../ramandecompy/tests/test_files/Methane_Baseline_Calibration.xlsx',
label='Methane')
dataprep.view_hdf5('dataprep_calibration.hdf5')
# Using `dataprep.view_hdf5` we can now see that both **Hydrogen** and **Methane** groups are contained within the file. The detected and fitted peak profiles are saved under each group along with the raw data. In this way, we see how multiple calibration data can be stored within a single `calibration.hdf5`.
#
# ### dataprep.add_experiment
#
# Next we will see how the slighly different function `dataprep.add_experiment` operates and how it stores experimental data under groups that specify the temperature and residence time for each experiment added. First we will make a new `experiment.hdf5` file to store the experimental data. Importing this file will take longer than the earlier examples since this spectra contains a larger number of peaks that need to be fit.
dataprep.new_hdf5('dataprep_experiment')
dataprep.add_experiment('dataprep_experiment.hdf5', '../ramandecompy/tests/test_files/FA_3.6wt%_300C_25s.csv')
dataprep.add_experiment('dataprep_experiment.hdf5', '../ramandecompy/tests/test_files/FA_3.6wt%_300C_35s.csv')
dataprep.add_experiment('dataprep_experiment.hdf5', '../ramandecompy/tests/test_files/FA_3.6wt%_300C_45s.csv')
dataprep.view_hdf5('dataprep_experiment.hdf5')
# ### dataprep.adjust_peaks
#
# In order to give the user the ability to apply their expert knowledge, this function allows peaks to be added and subtracted from the automatically generated fit. In this way, more difficult to parse compound peaks can be fit more accurately, which will be critical for future chemical yield calculations. The `dataplot.plot_fit()` function is used below to give an idea of how the automatic fit compares to the expert adjusted fit.
# plot of the original automatic fit from dataprep.add_experiment()
fig, ax1, ax2 = datavis.plot_fit('dataprep_experiment.hdf5', '300C/25s')
# +
# the add_list argument consists of a list of integer wavenumbers where the peak should approximately be
# the function allows the fit to adjust the center of the peak +/- 10 cm^-1
add_list = [1270, 1350, 1385]
# the drop_list argument consists of the string labels of the datasets/labels shown in the hdf5 file and
# the plot produced using dataprep.plot_fit
drop_list = ['Peak_01']
dataprep.adjust_peaks('dataprep_experiment.hdf5', '300C/25s', add_list, drop_list, plot_fits=True)
# -
# Running the dataprep.plot_fit function again we see that Peak_01 has been removed and Peak_09, Peak_10, and Peak_11 have been added to fit fit. These three added peaks contain a `*` in their label to identify them as manually added to the fit.
fig, ax1, ax2 = datavis.plot_fit('dataprep_experiment.hdf5', '300C/25s')
# ## Remove the file so that there are no errors - this is done in the basic .py file
# In order to keep the file system clean, and to avoid errors associated with running this notebook multiple times, we lastly will delete the two `.hdf5` files generated by this notebook. Comment out the final cell if you wish you explore these files further.
os.remove('dataprep_calibration.hdf5')
os.remove('dataprep_experiment.hdf5')
| examples/dataprep_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # Deep learning for hyperspectral image processing: Multi-layer perceptron networks
# This notebook demonstrates application of Multi-Layer Perceptron (MLP) networks to land use classification. Two seperate notebooks are also available describing the applications of 2-Dimensional Convolutional Neural Network [(2-D CNN)](deep_learning_2D_CNN.ipynb) and 3-Dimenaional Convolutional Neural Network [(3-D CNN)](deep_learning_3D_CNN.ipynb) models to landuse classification.
#
# ## Module imports
# Below is the list of libraries and modules that are required in this notebook. The 'keras' package provides the building blocks for model configuration and training. The 'img_util' contains a set of useful functions for pre-processing of raw data to create input and output data containers, compatible to the 'keras' package. In addition, it provides a set of functions for post-processing of results and visualization of prediction maps.
# The 'sio' and 'os' module were used for working with external files. The plotting of data and generation of prediction maps were achieved using plotting functionalities of 'matplotlib'.
import numpy as np
from keras import models, layers, optimizers, metrics, losses, regularizers
import img_util as util
from scipy import io as sio
import os
from matplotlib import pyplot as plt
# ## Hyperspectral dataset
# A set of publically-available hyperspectral imageray datasets can be downloaded form [this](http://www.ehu.eus/ccwintco/index.php/Hyperspectral_Remote_Sensing_Scenes) website. The Indian Pine dataset was downloaded and used in this notebook. The dataset consists of 150*150 pixels with 200 refelactance bands. The ground truth data for the dataset consists of 16 different classes. A summary of landuse types and their corresponding number of samples can be found in the following table:
# | ID | Class | Samples |
# |----------|------------------------------|---------|
# | 0 | Unlabeled | 10776 |
# | 1 | Alfalfa | 46 |
# | 2 | Corn-notill | 1428 |
# | 3 | Corn-mintill | 830 |
# | 4 | Corn | 237 |
# | 5 | Grass-pasture | 483 |
# | 6 | Grass-trees | 730 |
# | 7 | Grass-pasture-mowed | 28 |
# | 8 | Hay-windrowed | 478 |
# | 9 | Oats | 20 |
# | 10 | Soybean-notill | 972 |
# | 11 | Soybean-mintill | 2455 |
# | 12 | Soybean-clean | 593 |
# | 13 | Wheat | 205 |
# | 14 | Woods | 1265 |
# | 15 | Buildings-Grass-Trees-Drives | 386 |
# | 16 | Stone-Steel-Towers | 93 |
# The image data and class labels are available in two separate Matlab files with .mat extension. Therefore, the data were loaded into Python using the 'loadmat' function, available in the 'io' module of Scipy.
# +
data_folder = 'Datasets'
data_file= 'Indian_pines_corrected'
gt_file = 'Indian_pines_gt'
data_set = sio.loadmat(os.path.join(data_folder, data_file)).get('indian_pines_corrected')
gt = sio.loadmat(os.path.join(data_folder, gt_file)).get('indian_pines_gt')
# Checking the shape of data_set (containing image data) and gt (containing ground truth data) Numpy arrays.
print(data_set.shape ,gt.shape)
# -
# ## Training and test data
# The 'data_split' function was used for splitting the data into training and test sets using 0.85 as the split ratio (85% of labeled pixels were used for training). This function ensures that all classes are represented in the training dataset (see function documentation for available split methods). In addition, it allows users to focus their analysis on certain classes and remove those pixels that are not labeled. For example, the unlabeled data are represented by 0 in the gourd truth data file. Therefore, 0 was included in 'rem_classes' list, indicating its removal from the dataset.
# +
train_fraction = 0.85
rem_classes = [0]
(train_rows, train_cols), (test_rows, test_cols) = util.data_split(gt,
train_fraction=train_fraction,
rem_classes=rem_classes)
print('Number of training samples = {}.\nNumber of test samples = {}.'.format(len(train_rows), len(test_rows)))
# -
# A portion of training data can optionally be set aside for validation.
val_fraction = 0.05
(train_rows_sub, train_cols_sub), (val_rows, val_cols) = util.val_split(
train_rows, train_cols, gt, val_fraction=val_fraction)
# ## Dimensionality reduction
# The spectral dimension of an image dataset can be reduced using Principle Component Analysis (PCA). Although, this step is not necessary, it could significantly reduce the spectral dimension without losing important information. The 'reduce_dim' function takes the numpy array containing image data as its first argument and the number of reduced dimensions (i.e., an integer) or the minimum variance captured by PCA dimensions (i.e., a float) as the second argument.
data_set = util.reduce_dim(img_data=data_set, n_components=.999)
data_set.shape
# Using a value of 0.999 for the percentage of captured variance, The spectral dimension was reduced from 200 to 69 bands. The new dimensions were sorted according to their contribution to the dataset variance. The top 10 dimensions of transformed data are illustrated below.
# +
fig, axes = plt.subplots(2,5, True, True, figsize=(15,7))
for numb, axe in enumerate(axes.flat):
axe.imshow(data_set[:,:,numb])
axe.set_title('dim='+' '+str(numb))
fig.subplots_adjust(wspace=0, hspace=.2)
plt.show()
# -
# ## Rescaling data
# The 'rescale_data' function provides four methods for rescaling data at each spectral dimension. In this notebook, the'standard' method which transforms the data to have zero mean a standard deviation of 1 was used for rescaling data.
data_set = util.rescale_data(data_set)
# ## Creating input and target tensors
# The input and target tensors should be compatible with the type of neural network model that is used for classification. The 'create_patch' function can create inputs, compatible to both pixel inputs for [MLP](deep_learning_MLP.ipynb) models as well as patch inputs for [2-D CNN](deep_learning_2D_CNN.ipynb) and [3-D CNN](deep_learning_3D_CNN.ipynb) models.
# In this notebook, an MLP model was used for classification. Each pixel in the training dataset would constitute an input to the neural network model, therefore the value of 'patch_size' parameter should be set to 1.
# +
patch_size=1
train_pixel_indices_sub = (train_rows_sub, train_cols_sub)
val_pixel_indices = (val_rows, val_cols)
test_pixel_indices = (test_rows, test_cols)
catg_labels = np.unique([int(gt[idx[0],idx[1]]) for idx in zip(train_rows, train_cols)])
int_to_vector_dict = util.label_2_one_hot(catg_labels)
train_input_sub, y_train_sub = util.create_patch(
data_set=data_set,
gt=gt,
pixel_indices=train_pixel_indices_sub,
patch_size=patch_size,
label_vect_dict=int_to_vector_dict)
val_input, y_val = util.create_patch(
data_set=data_set,
gt=gt,
pixel_indices=val_pixel_indices,
patch_size=patch_size,
label_vect_dict=int_to_vector_dict)
test_input, y_test = util.create_patch(
data_set=data_set,
gt=gt,
pixel_indices=test_pixel_indices,
patch_size=patch_size,
label_vect_dict=int_to_vector_dict)
# -
# ## Creating an MLP neural network model
# The network architecture consists of an input layer whose neurons correspond to the dimension of inputs (i.e., the number of spectral bands). The input layer is followed by a Flatten layer which merely reshape the outputs of the input layer. The third layer is a 'dense' layer and contains the hidden neurons. A Dropout layer is placed after the hidden layer which randomly sets to zero the outputs of the hidden layer during the training phase. The last layer is the output layer whose dimension depends on the number of classes.
# +
units_1 = 2**8
drop_rate =0.35
num_catg = len(catg_labels)
input_shape = (patch_size, patch_size, data_set.shape[-1])
# Building a MLP network model
nn_model = models.Sequential()
#
# dense_input
nn_model.add(layer=layers.Dense(units=data_set.shape[2], activation='relu',
input_shape=input_shape))
# flatten_1, changes input shape from (1,1,num_band) to (num_band,)
nn_model.add(layer=layers.Flatten())
# dense_1
nn_model.add(layer=layers.Dense(units=units_1, activation='relu'))
# dropout_1
nn_model.add(layer=layers.Dropout(drop_rate))
# dense_output
nn_model.add(layer=layers.Dense(units=num_catg, activation='softmax'))
nn_model.summary()
# -
# ## Training model and plotting training history
# The model was compiled and trained using the training, validation and test [data.](#Creating-input-and-target-tensors)
# +
lr = 1e-4
batch_size = 2**3
# Compiling the modele
nn_model.compile(optimizer=optimizers.RMSprop(lr=lr),
loss=losses.categorical_crossentropy,
metrics=[metrics.categorical_accuracy])
# Training the model
history = nn_model.fit(x=train_input_sub, y=y_train_sub, batch_size=batch_size,
epochs=50, validation_data=(val_input, y_val), verbose=False)
# Plotting history
epoches = np.arange(1,len(history.history.get('loss'))+1)
fig, (ax1, ax2) = plt.subplots(1, 2, True, figsize=(15,7))
ax1.plot(epoches, history.history.get('loss'), 'b',label='Loss')
ax1.plot(epoches, history.history.get('val_loss'),'bo', label='Validation loss')
ax1.set_title('Training and validation loss')
ax1.legend()
ax2.plot(epoches, history.history.get('categorical_accuracy'), 'b',label='Accuracy')
ax2.plot(epoches, history.history.get('val_categorical_accuracy'),'bo', label='Validation accuracy')
ax2.set_title('Training and validation accuracy')
ax2.legend()
plt.show()
# -
# ## Model performance evaluation
# Overall loss and accuracy of the model was calculated using the 'evaluate' method. The loss and accuracy for each class was also calculated using the 'calc_metrics' function of the 'img_util' module.
# +
overall_loss, overal_accu = nn_model.evaluate(test_input, y_test, verbose=False)
print('Overall loss = {}'.format(overall_loss))
print('Overall accuracy = {}\n'.format(overal_accu))
# Calculating accuracy for each class
model_metrics = util.calc_metrics(nn_model, test_input,
y_test, int_to_vector_dict, verbose=False)
#Printing accuracy per class
print('{}{:>13}\n{}'.format('Class ID','Accuracy', 30*'_'))
for key, val in model_metrics.items():
print(('{:>2d}{:>18.4f}\n'+'{}').format(key, val[0][1], 30*'_'))
# -
# ## Making predictions using using test data
# The trained model was used for label predictions using the training, validation, and test datasets. It was also used to make label prediction for the entire dataset including unlabeled pixels.
# +
# Plotting predicted results
concat_rows = np.concatenate((train_rows_sub, val_rows, test_rows))
concat_cols = np.concatenate((train_cols_sub, val_cols, test_cols))
concat_input = np.concatenate((train_input_sub, val_input, test_input))
concat_y = np.concatenate((y_train_sub, y_val, y_test))
pixel_indices = (concat_rows, concat_cols)
partial_map = util.plot_partial_map(nn_model, gt, pixel_indices, concat_input,
concat_y, int_to_vector_dict, plo=False)
full_map = util.plot_full_map(nn_model, data_set, gt, int_to_vector_dict, patch_size, plo=False)
fig, (ax1, ax2) = plt.subplots(1,2,True, True, figsize=(15,7))
ax1.imshow(partial_map)
ax1.set_title('Prediction map for labeled data', fontweight="bold", fontsize='14')
ax2.imshow(full_map)
ax2.set_title('Prediction map for all data', fontweight="bold", fontsize='14')
plt.show()
# -
# The prediction map may be further improved using an appropriate filter (e.g. median filter) for removing the salt-and-pepper noise from the predicted pixels. Alternatively, CNN models which are less prone to producing a noisy prediction map could be used for landuse classification.
# See also:
# ### [Deep learning for hyperspectral image processing: Multi-layer perceptron networks](deep_learning_MLP.ipynb)
# ### [Deep learning for hyperspectral image processing: 2-D convolutional neural networks](deep_learning_2D_CNN.ipynb)
# ### [Deep learning for hyperspectral image processing: 3-D convolutional neural networks](deep_learning_3D_CNN.ipynb)
| deep_learning_MLP.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] id="cac470df-29e7-4148-9bbd-d8b9a32fa570" tags=[]
# # (시도) Node Classification with Graph Neural Networks
# >
#
# - toc:true
# - branch: master
# - badges: true
# - comments: false
# - author: 최서연
# - categories: [GNN]
# -
# ### Import
import os
import pandas as pd
import numpy as np
import networkx as nx
import matplotlib.pyplot as plt
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import layers
# ### 데이터 구성
# ---
# ```python
# p = 0.3
# papers = pd.concat([pd.DataFrame(np.array([[p]*1500,[1-p]*1500]).reshape(1000,3),columns=['X1','X2','X3']),
# pd.DataFrame(np.array([[1-p]*1500,[p]*1500]).reshape(1000,3),columns=['X4','X5','X6']),
# pd.DataFrame(np.array([['Deep learning']*500,['Reinforcement learning']*500]).reshape(1000,1))],axis=1).reset_index().rename(columns={'index':'paper_id',0:'subject'})
# papers['paper_id'] = papers['paper_id']+1
# #시도1: 2500 행 모두 target/source 상관없이 1~1000 임의 부여
# citations = pd.DataFrame(np.array([[np.random.choice(range(1,1001),size=(2500,1))],
# [np.random.choice(range(1,1001),size=(2500,1))]]).reshape(2500,2)).rename(columns = {0:'target',1:'source'})
# #시도2: 나머지 1250 행 reinforcement learning 행 500~1000를 target, source에 임의 부여
# citations = pd.concat([pd.DataFrame(np.array([np.random.choice(range(1,501),size=(1250,1)),np.random.choice(range(1,501),size=(1250,1))]).reshape(1250,2)),
# pd.DataFrame(np.array([np.random.choice(range(501,1001),size=(1250,1)),np.random.choice(range(501,1001),size=(1250,1))]).reshape(1250,2))],ignore_index=True).rename(columns={0:'target',1:'source'})
# ```
# ---
(x_train, y_train), (x_test, y_test) = tf.keras.datasets.mnist.load_data()
x_train.shape, y_train.shape, x_test.shape, y_test.shape
x_train, x_test = x_train[..., np.newaxis]/255.0, x_test[..., np.newaxis]/255.0
# def filter_36(x, y):
# keep = (y == 3) | (y == 7)
# x, y = x[keep], y[keep]
# y = y == 3
# return x,y
# x_train, y_train = filter_36(x_train, y_train)
# x_test, y_test = filter_36(x_test, y_test)
X= x_train.reshape(-1,784)/255
y = y_train
#y = list(map(lambda x: 0 if x == True else 1,y_train))
#XX = x_test.reshape(-1,784)/255
#yy = list(map(lambda x: 0 if x == True else 1,y_test))
# y가 3이면 0
#
# y가 7이면 1로
add_list = []
add_list.append(X)
papers = pd.concat([pd.DataFrame(np.array(add_list).reshape(-1,784)),pd.DataFrame(np.array(y).reshape(-1,1))],axis=1).reset_index().iloc[:4999]
column_names = ["paper_id"] + [f"X_{idx}" for idx in range(1,785)] + ["subject"]
papers.columns = column_names
papers['paper_id'] = papers['paper_id'] + 1
papers
_a = []
for i in range(1,len(papers)+1):
for j in range(1,len(papers)+1):
_a.append([i])
_a.append([j])
citations = pd.DataFrame(np.array(_a).reshape(-1,2)).rename(columns = {0:'target',1:'source'})
citations
# + [markdown] tags=[]
# ### 그래프 표현
# +
class_values = sorted(papers["subject"].unique())
class_idx = {name: id for id, name in enumerate(class_values)}
paper_idx = {name: idx for idx, name in enumerate(sorted(papers["paper_id"].unique()))}
papers["paper_id"] = papers["paper_id"].apply(lambda name: paper_idx[name])
citations["source"] = citations["source"].apply(lambda name: paper_idx[name])
citations["target"] = citations["target"].apply(lambda name: paper_idx[name])
papers["subject"] = papers["subject"].apply(lambda value: class_idx[value])
# -
plt.figure(figsize=(10, 10))
colors = papers["subject"].tolist()
cora_graph = nx.from_pandas_edgelist(citations.sample(n=800))
subjects = list(papers[papers["paper_id"].isin(list(cora_graph.nodes))]["subject"])
nx.draw_spring(cora_graph, node_size=15, node_color=subjects)
# ### Test vs Train
# +
train_data, test_data = [], []
for _, group_data in papers.groupby("subject"):
# Select around 50% of the dataset for training.
random_selection = np.random.rand(len(group_data.index)) <= 0.5
train_data.append(group_data[random_selection])
test_data.append(group_data[~random_selection])
train_data = pd.concat(train_data).sample(frac=1)
test_data = pd.concat(test_data).sample(frac=1)
print("Train data shape:", train_data.shape)
print("Test data shape:", test_data.shape)
# -
hidden_units = [32,32]
learning_rate = 0.01
dropout_rate = 0.5
num_epochs = 300
batch_size = 256
def run_experiment(model, x_train, y_train):
# Compile the model.
model.compile(
optimizer=keras.optimizers.Adam(learning_rate),
loss=keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=[keras.metrics.SparseCategoricalAccuracy(name="acc")],
)
# Create an early stopping callback.
early_stopping = keras.callbacks.EarlyStopping(
monitor="val_acc", patience=50, restore_best_weights=True
)
# Fit the model.
history = model.fit(
x=x_train,
y=y_train,
epochs=num_epochs,
batch_size=batch_size,
validation_split=0.15,
callbacks=[early_stopping],
)
return history
def display_learning_curves(history):
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(15, 5))
ax1.plot(history.history["loss"])
ax1.plot(history.history["val_loss"])
ax1.legend(["train", "test"], loc="upper right")
ax1.set_xlabel("Epochs")
ax1.set_ylabel("Loss")
ax2.plot(history.history["acc"])
ax2.plot(history.history["val_acc"])
ax2.legend(["train", "test"], loc="upper right")
ax2.set_xlabel("Epochs")
ax2.set_ylabel("Accuracy")
plt.show()
def create_ffn(hidden_units, dropout_rate, name=None):
fnn_layers = []
for units in hidden_units:
fnn_layers.append(layers.BatchNormalization())
fnn_layers.append(layers.Dropout(dropout_rate))
fnn_layers.append(layers.Dense(units, activation=tf.nn.gelu))
return keras.Sequential(fnn_layers, name=name)
# +
feature_names = set(papers.columns) - {"paper_id", "subject"}
num_features = len(feature_names)
num_classes = len(class_idx)
# Create train and test features as a numpy array.
x_train = train_data[feature_names].to_numpy()
x_test = test_data[feature_names].to_numpy()
# Create train and test targets as a numpy array.
y_train = train_data["subject"]
y_test = test_data["subject"]
# + tags=[]
def create_baseline_model(hidden_units, num_classes, dropout_rate=0.2):
inputs = layers.Input(shape=(num_features,), name="input_features")
x = create_ffn(hidden_units, dropout_rate, name=f"ffn_block1")(inputs)
for block_idx in range(4):
# Create an FFN block.
x1 = create_ffn(hidden_units, dropout_rate, name=f"ffn_block{block_idx + 2}")(x)
# Add skip connection.
x = layers.Add(name=f"skip_connection{block_idx + 2}")([x, x1])
# Compute logits.
logits = layers.Dense(num_classes, name="logits")(x)
# Create the model.
return keras.Model(inputs=inputs, outputs=logits, name="baseline")
baseline_model = create_baseline_model(hidden_units, num_classes, dropout_rate)
baseline_model.summary()
# + tags=[]
history = run_experiment(baseline_model, x_train, y_train)
# -
display_learning_curves(history)
_, test_accuracy = baseline_model.evaluate(x=x_test, y=y_test, verbose=0)
print(f"Test accuracy: {round(test_accuracy * 100, 2)}%")
# + [markdown] tags=[]
# ### baseline 모델 예측
# +
def generate_random_instances(num_instances):
token_probability = x_train.mean(axis=0)
instances = []
for _ in range(num_instances):
probabilities = np.random.uniform(size=len(token_probability))
instance = (probabilities <= token_probability).astype(int)
instances.append(instance)
return np.array(instances)
def display_class_probabilities(probabilities):
for instance_idx, probs in enumerate(probabilities):
print(f"Instance {instance_idx + 1}:")
for class_idx, prob in enumerate(probs):
print(f"- {class_values[class_idx]}: {round(prob * 100, 2)}%")
# -
new_instances = generate_random_instances(num_classes)
logits = baseline_model.predict(new_instances)
probabilities = keras.activations.softmax(tf.convert_to_tensor(logits)).numpy()
display_class_probabilities(probabilities)
theta = 5000
edge_weights = []
for i in range(len(citations)):
edge_weights.append(np.exp(-((citations.target[i] - citations.source[i])**2/theta).sum()))
# +
# Create an edges array (sparse adjacency matrix) of shape [2, num_edges].
edges = citations[["source", "target"]].to_numpy().T
# Create an edge weights array of ones.
#edge_weights = tf.ones(shape=edges.shape[1])
edge_weights = tf.constant(edge_weights,shape=edges.shape[1])
# Create a node features array of shape [num_nodes, num_features].
node_features = tf.cast(
papers.sort_values("paper_id")[feature_names].to_numpy(), dtype=tf.dtypes.float32
)
# Create graph info tuple with node_features, edges, and edge_weights.
graph_info = (node_features, edges, edge_weights)
print("Edges shape:", edges.shape)
print("Edge weight shape:", edge_weights.shape)
print("Nodes shape:", node_features.shape)
# -
class GraphConvLayer(layers.Layer):
def __init__(
self,
hidden_units,
dropout_rate=0.2,
aggregation_type="mean",
combination_type="concat",
normalize=False,
*args,
**kwargs,
):
super(GraphConvLayer, self).__init__(*args, **kwargs)
self.aggregation_type = aggregation_type
self.combination_type = combination_type
self.normalize = normalize
self.ffn_prepare = create_ffn(hidden_units, dropout_rate)
if self.combination_type == "gated":
self.update_fn = layers.GRU(
units=hidden_units,
activation="tanh",
recurrent_activation="sigmoid",
dropout=dropout_rate,
return_state=True,
recurrent_dropout=dropout_rate,
)
else:
self.update_fn = create_ffn(hidden_units, dropout_rate)
def prepare(self, node_repesentations, weights=None):
# node_repesentations shape is [num_edges, embedding_dim].
messages = self.ffn_prepare(node_repesentations)
if weights is not None:
messages = messages * tf.expand_dims(weights, -1)
return messages
def aggregate(self, node_indices, neighbour_messages):
# node_indices shape is [num_edges].
# neighbour_messages shape: [num_edges, representation_dim].
num_nodes = tf.math.reduce_max(node_indices) + 1
if self.aggregation_type == "sum":
aggregated_message = tf.math.unsorted_segment_sum(
neighbour_messages, node_indices, num_segments=num_nodes
)
elif self.aggregation_type == "mean":
aggregated_message = tf.math.unsorted_segment_mean(
neighbour_messages, node_indices, num_segments=num_nodes
)
elif self.aggregation_type == "max":
aggregated_message = tf.math.unsorted_segment_max(
neighbour_messages, node_indices, num_segments=num_nodes
)
else:
raise ValueError(f"Invalid aggregation type: {self.aggregation_type}.")
return aggregated_message
def update(self, node_repesentations, aggregated_messages):
# node_repesentations shape is [num_nodes, representation_dim].
# aggregated_messages shape is [num_nodes, representation_dim].
if self.combination_type == "gru":
# Create a sequence of two elements for the GRU layer.
h = tf.stack([node_repesentations, aggregated_messages], axis=1)
elif self.combination_type == "concat":
# Concatenate the node_repesentations and aggregated_messages.
h = tf.concat([node_repesentations, aggregated_messages], axis=1)
elif self.combination_type == "add":
# Add node_repesentations and aggregated_messages.
h = node_repesentations + aggregated_messages
else:
raise ValueError(f"Invalid combination type: {self.combination_type}.")
# Apply the processing function.
node_embeddings = self.update_fn(h)
if self.combination_type == "gru":
node_embeddings = tf.unstack(node_embeddings, axis=1)[-1]
if self.normalize:
node_embeddings = tf.nn.l2_normalize(node_embeddings, axis=-1)
return node_embeddings
def call(self, inputs):
"""Process the inputs to produce the node_embeddings.
inputs: a tuple of three elements: node_repesentations, edges, edge_weights.
Returns: node_embeddings of shape [num_nodes, representation_dim].
"""
node_repesentations, edges, edge_weights = inputs
# Get node_indices (source) and neighbour_indices (target) from edges.
node_indices, neighbour_indices = edges[0], edges[1]
# neighbour_repesentations shape is [num_edges, representation_dim].
neighbour_repesentations = tf.gather(node_repesentations, neighbour_indices)
# Prepare the messages of the neighbours.
neighbour_messages = self.prepare(neighbour_repesentations, edge_weights)
# Aggregate the neighbour messages.
aggregated_messages = self.aggregate(node_indices, neighbour_messages)
# Update the node embedding with the neighbour messages.
return self.update(node_repesentations, aggregated_messages)
class GNNNodeClassifier(tf.keras.Model):
def __init__(
self,
graph_info,
num_classes,
hidden_units,
aggregation_type="sum",
combination_type="concat",
dropout_rate=0.2,
normalize=True,
*args,
**kwargs,
):
super(GNNNodeClassifier, self).__init__(*args, **kwargs)
# Unpack graph_info to three elements: node_features, edges, and edge_weight.
node_features, edges, edge_weights = graph_info
self.node_features = node_features
self.edges = edges
self.edge_weights = edge_weights
# Set edge_weights to ones if not provided.
if self.edge_weights is None:
self.edge_weights = tf.ones(shape=edges.shape[1])
# Scale edge_weights to sum to 1.
self.edge_weights = self.edge_weights / tf.math.reduce_sum(self.edge_weights)
# Create a process layer.
self.preprocess = create_ffn(hidden_units, dropout_rate, name="preprocess")
# Create the first GraphConv layer.
self.conv1 = GraphConvLayer(
hidden_units,
dropout_rate,
aggregation_type,
combination_type,
normalize,
name="graph_conv1",
)
# Create the second GraphConv layer.
self.conv2 = GraphConvLayer(
hidden_units,
dropout_rate,
aggregation_type,
combination_type,
normalize,
name="graph_conv2",
)
# Create a postprocess layer.
self.postprocess = create_ffn(hidden_units, dropout_rate, name="postprocess")
# Create a compute logits layer.
self.compute_logits = layers.Dense(units=num_classes, name="logits")
def call(self, input_node_indices):
# Preprocess the node_features to produce node representations.
x = self.preprocess(self.node_features)
# Apply the first graph conv layer.
x1 = self.conv1((x, self.edges, self.edge_weights))
# Skip connection.
x = x1 + x
# Apply the second graph conv layer.
x2 = self.conv2((x, self.edges, self.edge_weights))
# Skip connection.
x = x2 + x
# Postprocess node embedding.
x = self.postprocess(x)
# Fetch node embeddings for the input node_indices.
node_embeddings = tf.gather(x, input_node_indices)
# Compute logits
return self.compute_logits(node_embeddings)
# +
gnn_model = GNNNodeClassifier(
graph_info=graph_info,
num_classes=num_classes,
hidden_units=hidden_units,
dropout_rate=dropout_rate,
name="gnn_model",
)
print("GNN output shape:", gnn_model([1, 10, 100]))
gnn_model.summary()
# + tags=[]
x_train = train_data.paper_id.to_numpy()
history = run_experiment(gnn_model, x_train, y_train)
# -
display_learning_curves(history)
x_test = test_data.paper_id.to_numpy()
_, test_accuracy = gnn_model.evaluate(x=x_test, y=y_test, verbose=0)
print(f"Test accuracy: {round(test_accuracy * 100, 2)}%")
# + tags=[]
# First we add the N new_instances as nodes to the graph
# by appending the new_instance to node_features.
num_nodes = node_features.shape[0]
new_node_features = np.concatenate([node_features, new_instances])
# Second we add the M edges (citations) from each new node to a set
# of existing nodes in a particular subject
new_node_indices = [i + num_nodes for i in range(num_classes)]
new_citations = []
for subject_idx, group in papers.groupby("subject"):
subject_papers = list(group.paper_id)
# Select random x papers specific subject.
selected_paper_indices1 = np.random.choice(subject_papers, 5)
# Select random y papers from any subject (where y < x).
selected_paper_indices2 = np.random.choice(list(papers.paper_id), 2)
# Merge the selected paper indices.
selected_paper_indices = np.concatenate(
[selected_paper_indices1, selected_paper_indices2], axis=0
)
# Create edges between a citing paper idx and the selected cited papers.
citing_paper_indx = new_node_indices[subject_idx]
for cited_paper_idx in selected_paper_indices:
new_citations.append([citing_paper_indx, cited_paper_idx])
new_citations = np.array(new_citations).T
new_edges = np.concatenate([edges, new_citations], axis=1)
# +
print("Original node_features shape:", gnn_model.node_features.shape)
print("Original edges shape:", gnn_model.edges.shape)
gnn_model.node_features = new_node_features
gnn_model.edges = new_edges
gnn_model.edge_weights = tf.ones(shape=new_edges.shape[1])
print("New node_features shape:", gnn_model.node_features.shape)
print("New edges shape:", gnn_model.edges.shape)
logits = gnn_model.predict(tf.convert_to_tensor(new_node_indices))
probabilities = keras.activations.softmax(tf.convert_to_tensor(logits)).numpy()
display_class_probabilities(probabilities)
# -
| _notebooks/2022-06-08-GNN-practice.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Coding Exercise #0401
# ### 1. K-Means clustering with simulated data:
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import os
from sklearn.cluster import KMeans
from sklearn.datasets import make_blobs
# %matplotlib inline
# #### 1.1. Generate simulated data and visualize:
# Dataset #1.
X1, label1 = make_blobs(n_samples=100, n_features=2, centers=2, cluster_std = 5, random_state=123)
plt.scatter(X1[:,0],X1[:,1], c= label1, alpha=0.7 )
plt.title('Dataset #1 : Original')
plt.show()
# Dataset #2
X2, label2 = make_blobs(n_samples=100, n_features=2, centers=3, cluster_std = 1, random_state=321)
plt.scatter(X2[:,0],X2[:,1], c= label2, alpha=0.7 )
plt.title('Dataset #2 : Original')
plt.show()
# #### 1.2. Apply k-means clustering and visualize:
# Dataset #1 and two clusters.
kmeans = KMeans(n_clusters=2,random_state=123) # kmeans object for 2 clusters. radom_state=123 means deterministic initialization.
kmeans.fit(X1) # Unsupervised learning => Only X1.
myColors = {0:'red',1:'green', 2:'blue'} # Define a color palette: 0~2.
plt.scatter(X1[:,0],X1[:,1], c= pd.Series(kmeans.labels_).apply(lambda x: myColors[x]), alpha=0.7 )
plt.title('Dataset #1 : K-Means')
plt.show()
# Dataset #1 and three clusters.
kmeans = KMeans(n_clusters=3,random_state=123) # kmeans object for 3 clusters. radom_state=123 means deterministic initialization.
kmeans.fit(X1) # Unsupervised learning => Only X1.
plt.scatter(X1[:,0],X1[:,1], c= pd.Series(kmeans.labels_).apply(lambda x: myColors[x]), alpha=0.7 )
plt.title('Dataset #1 : K-Means')
plt.show()
# Dataset #2 and two clusters.
kmeans = KMeans(n_clusters=2,random_state=123) # kmeans object for 2 clusters. radom_state=123 means deterministic initialization.
kmeans.fit(X2) # Unsupervised learning => Only X2.
plt.scatter(X2[:,0],X2[:,1], c= pd.Series(kmeans.labels_).apply(lambda x: myColors[x]), alpha=0.7 )
plt.title('Dataset #2 : K-Means')
plt.show()
# Dataset #2 and three clusters.
kmeans = KMeans(n_clusters=3, random_state=123) # kmeans object for 3 clusters. radom_state=123 means deterministic initialization.
kmeans.fit(X2) # Unsupervised learning => Only X2.
plt.scatter(X2[:,0],X2[:,1], c= pd.Series(kmeans.labels_).apply(lambda x: myColors[x]), alpha=0.7 )
plt.title('Dataset #2 : K-Means')
plt.show()
# NOTE: k means clusters have linear boundaries. The clustering result may reveal the existing structure, but this is not always guaranteed.
| SIC_AI_Coding_Exercises/SIC_AI_Chapter_05_Coding_Exercises/ex_0401.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Work
# 1. 請將 Epoch 加到 500 個,並觀察 learning curve 的走勢
# 2. 請將 Optimizer 換成 SGD,並觀察 learning curve 的走勢
# +
import os
import keras
# 本作業可以不需使用 GPU, 將 GPU 設定為 "無" (若有 GPU 且想開啟,可設為 "0")
os.environ["CUDA_VISIBLE_DEVICES"] = "0"
# -
# 從 Keras 的內建功能中,取得 train 與 test 資料集
train, test = keras.datasets.cifar10.load_data()
# +
# 將 X 與 Y 獨立放進變數
x_train, y_train = train
x_test, y_test = test
# 資料前處理 - 標準化
x_train = x_train / 255.
x_test = x_test / 255.
# 將資料從圖形 (RGB) 轉為向量 (Single Vector)
x_train = x_train.reshape((len(x_train), -1))
x_test = x_test.reshape((len(x_test), -1))
# 將目標轉為 one-hot encoding
y_train = keras.utils.to_categorical(y_train, num_classes=10)
y_test = keras.utils.to_categorical(y_test, num_classes=10)
# -
def build_mlp():
"""Code Here
建立你的神經網路
"""
input_layer = keras.layers.Input([x_train.shape[-1]])
x = keras.layers.Dense(units=512, activation="relu")(input_layer)
x = keras.layers.Dense(units=256, activation="relu")(x)
x = keras.layers.Dense(units=128, activation="relu")(x)
out = keras.layers.Dense(units=10, activation="softmax")(x)
model = keras.models.Model(inputs=[input_layer], outputs=[out])
return model
model = build_mlp()
# +
"""
Compile 模型
"""
model = build_mlp()
# 用 Keras 內建方法檢視模型各層參數量
model.summary()
optimizer = keras.optimizers.Adam(lr=0.001)
model.compile(loss="categorical_crossentropy", metrics=["accuracy"], optimizer=optimizer)
# -
"""
設定要訓練的 Epoch 數
"""
model.fit(x_train, y_train,
epochs=500,
batch_size=256,
validation_data=(x_test, y_test),
shuffle=True)
# +
import matplotlib.pyplot as plt
# 以視覺畫方式檢視訓練過程
train_loss = model.history.history["loss"]
valid_loss = model.history.history["val_loss"]
train_acc = model.history.history["acc"]
valid_acc = model.history.history["val_acc"]
plt.plot(range(len(train_loss)), train_loss, label="train loss")
plt.plot(range(len(valid_loss)), valid_loss, label="valid loss")
plt.legend()
plt.title("Loss")
plt.show()
plt.plot(range(len(train_acc)), train_acc, label="train accuracy")
plt.plot(range(len(valid_acc)), valid_acc, label="valid accuracy")
plt.legend()
plt.title("Accuracy")
plt.show()
| homeworks/D077/Day077_HW.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Reproducing graphs and measurements from the VoteAgain paper
#
# This notebook reproduces the performance graphs and numbers used in the paper.
#
# ## Setup
# We first input some packages
# +
import copy
import csv
import math
import numpy
import statistics
import matplotlib.pyplot as plt
# -
# And set up matplotlib.
plt.rc('text', usetex=True)
plt.rc('font', size=12)
#plt.rc('figure', figsize=(5.5,5))
# ## Simple measurements
#
# We benchmarked the cost of a single P256 group operation on the same machine that was used to run the other experiments.
GROUP_OPERATION_TIME = 47.556254e-6
# ## Utility functions
# We define two utility functions to parse the CSV files output by the various scripts, and to compute statistics on them
#
# **TODO**: documentation of `compute_stats` is not completely up to date
def parse_csv(filename):
"""
WARNING - WARNING -> See below
The content of the file as:
Foo, Bar, Baz
1, 3, 5
2, 4, 6
becomes:
{'Foo': [1,2], 'Bar': [3,4], 'Baz': [5,6]}
:param filename: Name of the CSV file.
:return: The content of the CSV file as a dict.
"""
content = {}
with open(filename, newline='') as f:
reader = csv.reader(f, delimiter=',')
# Retrieve the titles and set them as keys.
titles = reader.__next__()
for title in titles:
content[title] = []
for row in reader:
# Only the lines which does not contains the titles are considered.
if row[0] not in content.keys():
for index, item in enumerate(row):
content[titles[index]].append(float(item))
return content
def compute_stats(parsed_csv, x_vars):
"""
Compute statistics for buckets of data.
in:
{
'Foo': [1, 1, 2, 2],
'Bar': [1, 2, 3, 4]
}
out:
{
'measurements': [1, 2],
'Bar':
{
'means': [1.5, 3.5],
'sems': [0.5, 0.5]
}
}
"""
# Convenience function, convert x_vars into list if it is a string
if isinstance(x_vars, str):
x_vars = [x_vars]
# The indexes for each set of values.
index_same_values = {}
for index, value in enumerate(zip(*[parsed_csv[x_var] for x_var in x_vars])):
index_same_values.setdefault(value, []).append(index)
fields_to_parse = set(parsed_csv.keys()) - set(x_vars)
output = {"measurements": list(index_same_values.keys())}
for key in fields_to_parse:
raw_values = parsed_csv[key]
means = []
sems = []
for indexes in index_same_values.values():
bucket = [raw_values[i] for i in indexes]
mean = statistics.mean(bucket)
sem = statistics.stdev(bucket) / math.sqrt(len(bucket))
means.append(mean)
sems.append(sem)
output[key] = {'means': means, 'sems': sems}
return output
# The following function creates a quick plot of the imported data.
def show_plot(values, titles, x_var):
"""
Display a nice plot for a set of data.
:param values: values to be plotted as a dict.
:param titles: list of titles in order.
:param x_val: values to use as the x.
"""
fig, ax = plt.subplots()
colors = ('blue', 'red', 'green', 'cyan', 'magenta', 'yellow')
col = iter(colors)
x = values["measurements"]
for title in titles:
y = values[title]['means']
yerr = values[title]['sems']
ax.errorbar(x, y, yerr=yerr, label=title.replace('_', ' '), color=next(col), fmt='.-')
ax.set_ylabel('Time [s]')
ax.set_xscale('log')
ax.set_yscale('log')
ax.set_xlabel(x_var.replace('_', ' '))
ax.legend(loc=0)
# ax.xlim(1e4, 1e6)
plt.show()
# ## Plotting performance of VoteAgain
#
# We first import the data from the experiments.
# +
full_filter_raw = parse_csv('../data/filter/full_filter.csv')
full_filter_ver_raw = parse_csv('../data/filter/full_filter_ver.csv')
full_filter = compute_stats(full_filter_raw, 'Nr_Voters')
full_filter_ver = compute_stats(full_filter_ver_raw, 'Nr_Voters')
# -
# We plot a summary of the results. These figures are not included in the paper.
titles = ['Shuffle', 'Decrypt' , 'Reencrypt' , 'FinalShuffle', 'FullFilter']
show_plot(full_filter, ["Dummies"] + titles , 'Nr_Voters')
show_plot(full_filter_ver, titles + ['FinalOpen'], 'Nr_Voters')
# We compute an lowerbound for Achenbach et al.'s protocol based on a count of the number of group operations and a measurement of the time it takes to perform such a group operations.
filter_achenbach = []
for voters in full_filter["measurements"]:
# voters is tuple with only a single element
voters = voters[0]
square = voters * voters
filter_achenbach.append(29 * square * GROUP_OPERATION_TIME)
# We plot the Filter and VerifyFilter times for VoteAgain, together with the estimated lower-bounds for Achenbach et al.'s protocol.
# +
fig, ax = plt.subplots(figsize=(4,2.5))
colors = ('blue', 'red', 'green', 'cyan', 'magenta', 'yellow')
col = iter(colors)
x = [val[0] for val in full_filter["measurements"]]
filter_time = full_filter["FullFilter"]['means']
filter_time_err = full_filter["FullFilter"]['sems']
ax.errorbar(x, filter_time, yerr=filter_time_err, label="Filter", color="blue", fmt='o-')
filter_time_ver = full_filter_ver["FullFilter"]['means']
filter_time_ver_err = full_filter_ver["FullFilter"]['sems']
ax.errorbar(x, filter_time_ver, yerr=filter_time_ver_err, label="VerifyFilter", color="red", fmt='^-')
ax.errorbar(x, filter_achenbach, label="Achenbach Filter", color="green", fmt='--')
ax.set_ylabel('Time (s)')
ax.set_xscale('log')
ax.set_yscale('log')
ax.set_xlabel("\#voters")
ax.legend(loc=0)
# ax.xlim(1e4, 1e6)
plt.savefig("../figures/measured-cost.pdf", bbox_inches="tight")
# -
# ## Estimating cost on Xeon machine
# We use the above measured running times to estimate the cost on a 8x28 core Xeon machine, assuming a scaling factor of 170.
scale = 170
# We first fit the measured data so we can use it in our computations.
fit = numpy.polyfit(x, filter_time, 1)
filter_time_fn = numpy.poly1d(fit)
fit = numpy.polyfit(x, filter_time_ver, 1)
filter_time_ver_fn = numpy.poly1d(fit)
# And we plot the estimates in a graph for the paper.
# +
x = numpy.logspace(5, 9, num = 100)
fig, ax = plt.subplots(figsize=(4,2.5))
ax.errorbar(x, filter_time_fn(x) / scale, label="Filter", color="blue", fmt='--')
ax.errorbar(x, filter_time_ver_fn(x) / scale, label="VerifyFilter", color="red", fmt='--')
ax.set_ylabel('Estimated time (s)')
ax.set_xscale('log')
ax.set_yscale('log')
ax.set_xlabel("\#voters")
ax.legend(loc=0)
plt.savefig("../figures/estimated-cost.pdf", bbox_inches="tight")
# -
# ## Graphs for ballot distribution
#
# We start out by reading the data gathered, and parsing it
# +
full_distr_raw = parse_csv('../data/distr/full_filter.csv')
full_distr_ver_raw = parse_csv('../data/distr/full_filter_ver.csv')
stats_raw = compute_stats(full_distr_raw, ["Nr_Voters","Total_Revotes","Extra_Padding"])
stats_ver_raw = compute_stats(full_distr_ver_raw, ["Nr_Voters","Total_Revotes","Extra_Padding"])
# -
# We now process the data we gathered to compute processing time per ballot. The experiments do not output the total number of balots, but instead output the number of boters (who each cast one ballot), the sum of revotes (by all voters together) and the extra ballots added by the TS (extra padding). We add these to get the total number of ballots.
# +
total_ballots = numpy.array([ nv + tr + ep for (nv, tr, ep) in stats_raw['measurements']])
ballots_per_voter = total_ballots / numpy.array([nv for (nv, _, _) in stats_raw['measurements']])
total_time = (numpy.array(stats_raw['FullFilter']['means']) - numpy.array(stats_raw['Dummies']['means'])) / total_ballots
total_time_sem = numpy.array(stats_raw['FullFilter']['sems']) / total_ballots
total_time_ver = numpy.array(stats_ver_raw['FullFilter']['means']) / total_ballots
total_time_ver_sem = numpy.array(stats_raw['FullFilter']['sems']) / total_ballots
# -
# We now plot the data to produce the figure for the paper.
# +
fig, ax = plt.subplots(figsize=(4,2.5))
colors = ('blue', 'red', 'green', 'cyan', 'magenta', 'yellow')
col = iter(colors)
ax.errorbar(ballots_per_voter, total_time, yerr=total_time_sem, label="Filter", color=next(col), fmt='.-')
ax.errorbar(ballots_per_voter, total_time_ver, yerr=total_time_ver_sem, label="VerifyFilter", color=next(col), fmt='.-')
ax.set_ylabel('Time per ballot (s)')
ax.set_xlabel('Ballots per voter')
ax.set_xscale('log')
ax.set_ylim(bottom=0)
ax.legend(loc=0)
plt.savefig("../figures/distr.pdf", bbox_inches="tight")
# -
# ## Computing numbers used in the paper
# We estimate computation time in several countries. We did not use these numbers.
# +
print("Some data for specific countries:")
countries = {"France": 67372000, "Russia": 146877088, "Brazil": 204519000, "Brazil (registered)": 147305825}
for country, n in countries.items():
t = filter_time_fn(n) / scale
print("{}: {} s or {} min or {} hours".format(country, t, t / 60, t / 3600))
# -
# ### Computing cost for Iowa Caucasus
# Used in the introduction.
# +
def report_computation_time(number_ballots):
number_operations = (number_ballots ** 2) * 29
achenbach_time_seconds = GROUP_OPERATION_TIME * number_operations
print("Achenbach time (days): ", achenbach_time_seconds / 86400)
print("Achenbach time (months): ", achenbach_time_seconds / (86400 * 30))
print("Achenbach time (years): ", achenbach_time_seconds / 31536000)
voteagain_time_seconds = filter_time_fn(number_ballots)
print("\nVoteAgain time (seconds)", voteagain_time_seconds)
print("VoteAgain time (minutes)", voteagain_time_seconds / 60)
iowa_size = 176_574
print("## Iowa Caucus ({} voters) ##".format(iowa_size))
report_computation_time(iowa_size)
small_town = 100_000
print("\n\n## Small Town ({} voters) ##".format(small_town))
report_computation_time(small_town)
# -
# ### Computing cost for a large City
# +
def report_computation_time_server(number_ballots):
number_operations = (number_ballots ** 2) * 29
achenbach_time_seconds = (GROUP_OPERATION_TIME * number_operations) / scale
print("Achenbach time (days): ", achenbach_time_seconds / 86400)
print("Achenbach time (months): ", achenbach_time_seconds / (86400 * 30))
print("Achenbach time (years): ", achenbach_time_seconds / 31536000)
voteagain_time_seconds = filter_time_fn(number_ballots) / scale
print("\nVoteAgain time (seconds)", voteagain_time_seconds)
print("VoteAgain time (minutes)", voteagain_time_seconds / 60)
big_town = 1_000_000
print("## Large City ({} voters) ##".format(big_town))
report_computation_time_server(big_town)
# -
# ### Time for Brazilian election
# We first compute the time to filter the Brazilian election assuming no revotes, and 147M voters.
number_voters = 147_000_000
time = filter_time_fn(number_voters) / scale
print("Brazilian elections (147M voters, no revoting): {} minutes".format(time / 60))
# Next, we assume revoting, with a maximum of 50% (i.e., at most 147M/2 extra ballots) and a maximum rate of 1/10 seconds.
number_ballots = number_voters * 10 # Using the x10 numbers from the graphs
time = filter_time_fn(number_ballots) / scale
print("Brazilian elections (147M voters, no revoting): {} hours".format(time / 3600))
# # Mix and decrypt times
# First, we read the data gathered and parsing it.
mix_raw = parse_csv('../data/mix/mix_and_decrypt.csv')
mix_stats_raw = compute_stats(mix_raw, ["NumberCiphertexts"])
# The experiment separately measures the time to do a verifiable proof and to do a verifiable decryption. Therefore we add the corresponding numbers. For convenience, we plot the result. We do not use this plot in the paper.
# +
nr_ciphertexts = [v[0] for v in mix_stats_raw["measurements"]]
mix_dec_time = numpy.array(mix_stats_raw["ShuffleAndProofTime"]["means"]) + numpy.array(mix_stats_raw["DecryptAndProofTime"]["means"])
fig, ax = plt.subplots(figsize=(4,2.5))
colors = ('blue', 'red', 'green', 'cyan', 'magenta', 'yellow')
col = iter(colors)
ax.errorbar(nr_ciphertexts, mix_dec_time, label="MixAndDecrypt", color=next(col), fmt='.-')
ax.set_ylabel('Time (s)')
ax.set_xlabel('Nr. of ciphertexts')
ax.set_xscale('log')
ax.set_yscale('log')
ax.set_ylim(bottom=0.05)
ax.legend(loc=0)
# -
# We use the measurements to fit an estimate. We use this estimate to compute the numbers used in the paper.
# +
# Fit MixAndDecrypt time
fit = numpy.polyfit(nr_ciphertexts, mix_dec_time, 1)
mix_dec_time_fn = numpy.poly1d(fit)
print("Time to mix+decrypt 100,000 ballots is {} seconds".format(mix_dec_time_fn(100_000)))
time1M = mix_dec_time_fn( 1_000_000 ) / scale
print("Time to mix+decrypt 1M ballots on server is {} seconds".format(time1M))
| analysis/analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# # SELECT names
#
# ## Pattern Matching Strings
# This tutorial uses the **LIKE** operator to check names. We will be using the SELECT command on the table world:
library(tidyverse)
library(DBI)
library(getPass)
drv <- switch(Sys.info()['sysname'],
Windows="PostgreSQL Unicode(x64)",
Darwin="/usr/local/lib/psqlodbcw.so",
Linux="PostgreSQL")
con <- dbConnect(
odbc::odbc(),
driver = drv,
Server = "localhost",
Database = "sqlzoo",
UID = "postgres",
PWD = getPass("Password?"),
Port = 5432
)
options(repr.matrix.max.rows=20)
# ## 1.
#
# You can use `WHERE name LIKE 'B%'` to find the countries that start with "B".
#
# The % is a _wild-card_ it can match any characters
#
# **Find the country that start with Y**
world <- dbReadTable(con, 'world')
world %>%
filter(str_starts(name, '[Yy]')) %>%
select(name)
# ## 2.
#
# **Find the countries that end with y**
world %>%
filter(str_ends(name, '[Yy]')) %>%
select(name)
# ## 3.
#
# Luxembourg has an **x** - so does one other country. List them both.
#
# **Find the countries that contain the letter x**
world %>%
filter(str_detect(name, '[Xx]')) %>%
select(name)
# ## 4.
#
# Iceland, Switzerland end with **land** - but are there others?
#
# **Find the countries that end with land**
world %>%
filter(str_ends(name, 'land')) %>%
select(name)
# ## 5.
#
# Columbia starts with a **C** and ends with **ia** - there are two more like this.
#
# **Find the countries that start with C and end with ia**
world %>%
filter(str_detect(name, '^[Cc].*ia$')) %>%
select(name)
# ## 6.
# Greece has a double **e** - who has **a** double **o**?
#
# **Find the country that has oo in the name**
world %>%
filter(str_detect(name, 'oo')) %>%
select(name)
# ## 7.
#
# Bahamas has three **a** - who else?
#
# **Find the countries that have three or more a in the name**
world %>%
filter(str_detect(name, 'a.*a.*a')) %>%
select(name)
# ## 8.
#
# India and Angola have an **n** as the second character. You can use the underscore as a single character wildcard.
#
# ```sql
# SELECT name FROM world
# WHERE name LIKE '_n%'
# ORDER BY name
# ```
#
# **Find the countries that have "t" as the second character.**
world %>%
filter(str_detect(name, '^.{1}t')) %>%
select(name)
# ## 9.
#
# Lesotho and Moldova both have two o characters separated by two other characters.
#
# **Find the countries that have two "o" characters separated by two others.**
world %>%
filter(str_detect(name, 'o.{2}o')) %>%
select(name)
# ## 10.
#
# Cuba and Togo have four characters names.
#
# **Find the countries that have exactly four characters.**
world %>%
filter(str_detect(name, '^.{4}$')) %>%
select(name)
# ## 11.
#
# The capital of **Luxembourg** is **Luxembourg**. Show all the countries where the capital is the same as the name of the country
#
# **Find the country where the name is the capital city.**
world %>%
filter(name==capital) %>%
select(name)
# ## 12.
#
# The capital of **Mexico** is **Mexico City**. Show all the countries where the capital has the country together with the word "City".
#
# **Find the country where the capital is the country plus "City".**
#
# > _The concat function_
# > The function concat is short for concatenate - you can use it to combine two or more strings.
world %>%
filter(capital==paste(name, 'City')) %>%
select(name)
# ## 13.
#
# **Find the capital and the name where the capital includes the name of the country.**
world %>%
filter(str_detect(capital, name)) %>%
select(capital, name)
# ## 14.
#
# **Find the capital and the name where the capital is an extension of name of the country.**
#
# You _should_ include **Mexico City** as it is longer than **Mexico**. You _should not_ include **Luxembourg** as the capital is the same as the country.
world %>%
filter(str_detect(capital, name) & capital != name) %>%
select(capital, name)
# ## 15.
#
# For **Monaco-Ville** the name is **Monaco** and the extension is **-Ville**.
#
# **Show the name and the extension where the capital is an extension of name of the country.**
#
# You can use the SQL function [REPLACE](https://sqlzoo.net/wiki/REPLACE).
world %>%
filter(str_detect(capital, paste('^', name, '.+$', sep=''))) %>%
mutate(extension=str_replace(capital, name, '')) %>%
select(name, extension)
dbDisconnect(con)
| R/01 SELECT name.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:100% !important; }</style>"))
# # Make Segmentation Figures
# ### Tiles
# +
import cv2
import numpy
import os, sys, glob
import numpy as np
# %matplotlib inline
from matplotlib import pyplot as plt
# +
imglist = sorted(glob.glob('/Users/nathaning/Dropbox/projects/semantic_pca/results/xval_set_0_val/jpg/*jpg'))
gtlist = sorted(glob.glob('/Users/nathaning/Dropbox/projects/semantic_pca/results/xval_set_0_val/mask/*png'))
fcn_512_list = sorted(glob.glob('/Users/nathaning/Dropbox/projects/semantic_pca/results/analysis_fcn/xval_set_0_1024/*png'))
segnet_512_list = sorted(glob.glob('/Users/nathaning/Dropbox/projects/semantic_pca/results/analysis_segnet/xval_set_0_1024/*png'))
segnet_basic_512_list = sorted(glob.glob('/Users/nathaning/Dropbox/projects/semantic_pca/results/analysis_segnet_basic/xval_set_0_1024/*png'))
print len(imglist), len(gtlist), len(fcn_512_list), len(segnet_512_list), len(segnet_basic_512_list)
# -
def colorize(mask):
colors={
0: [30, 30, 234],
1: [63, 210, 68],
2: [245, 25, 70],
3: [210, 210, 210]
}
img = np.zeros_like(mask, dtype=np.uint8)
img = np.dstack([img, img, img])
for u in np.unique(mask):
img[(mask == u), :] = colors[u]
return img
# +
outpath = '/Users/nathaning/Dropbox/_Papers/pca segmentation/Figs/tile_segmentations/xval_0_1024'
for imgp, gtp, fcnp, segnetp, segnetbp in zip(
imglist, gtlist, fcn_512_list, segnet_512_list, segnet_basic_512_list):
# imgp = imglist[0]
# gtp = gtlist[0]
# fcnp = fcn_512_list[0]
# segnetp = segnet_512_list[0]
# segnetbp = segnet_basic_512_list[0]
img_base = os.path.basename(imgp)
outfile = os.path.join(outpath, img_base.replace('.jpg', '.png'))
img = cv2.imread(imgp)[:,:,::-1]
gt = cv2.imread(gtp, -1)
fcn = cv2.imread(fcnp, -1)
segnet = cv2.imread(segnetp, -1)
segnetb = cv2.imread(segnetbp, -1)
size = 300
img = cv2.resize(img, dsize=(size,size))
gt = cv2.resize(gt, dsize=(size,size), interpolation=cv2.INTER_NEAREST)
fcn = cv2.resize(fcn, dsize=(size,size), interpolation=cv2.INTER_NEAREST)
segnet = cv2.resize(segnet, dsize=(size,size), interpolation=cv2.INTER_NEAREST)
segnetb = cv2.resize(segnetb, dsize=(size,size), interpolation=cv2.INTER_NEAREST)
gt = colorize(gt)
fcn = colorize(fcn)
segnet = colorize(segnet)
segnetb = colorize(segnetb)
img_= cv2.copyMakeBorder(img,10,10,10,10,cv2.BORDER_CONSTANT,value=(255,255,255))
gt_ = cv2.copyMakeBorder(gt,10,10,10,10,cv2.BORDER_CONSTANT,value=(255,255,255))
fcn_ = cv2.copyMakeBorder(fcn,10,10,10,10,cv2.BORDER_CONSTANT,value=(255,255,255))
segnet_ = cv2.copyMakeBorder(segnet,10,10,10,10,cv2.BORDER_CONSTANT,value=(255,255,255))
segnetb_ = cv2.copyMakeBorder(segnetb,10,10,10,10,cv2.BORDER_CONSTANT,value=(255,255,255))
imgout = np.vstack([img_, gt_, fcn_, segnet_, segnetb_])
cv2.imwrite(outfile, imgout[:,:,::-1])
# -
plt.imshow(np.vstack([img_, gt_, fcn_, segnet_, segnetb_]))
# # WSI
annot_list = sorted(glob.glob('/Users/nathaning/Dropbox/projects/semantic_pca/data/annotations/wsi_annotation/*.jpg'))
fcn_list = sorted(glob.glob('/Users/nathaning/Dropbox/projects/semantic_pca/results/fcn8s/*RGB.png'))
full_list = sorted(glob.glob('/Users/nathaning/Dropbox/projects/semantic_pca/results/segnet_full/*RGB.png'))
basic_list = sorted(glob.glob('/Users/nathaning/Dropbox/projects/semantic_pca/results/segnet_basic/*RGB.png'))
# +
outpath = '/Users/nathaning/Dropbox/_Papers/pca segmentation/Figs/wsi/figs'
for annotp, fcnp, fullp, basicp in zip(
annot_list, fcn_list, full_list, basic_list):
# annotp = annot_list[0]
# fcnp = fcn_list[0]
# fullp = full_list[0]
# basicp = basic_list[0]
outname = os.path.basename(annotp)
annot = cv2.imread(annotp)
fcn = cv2.imread(fcnp)
full = cv2.imread(fullp)
basic = cv2.imread(basicp)
targ_y = 600.
x,y = annot.shape[:2]
r = targ_y / y
annot = cv2.resize(annot, dsize=(0,0), fx=r, fy=r)
fcn = cv2.resize(fcn, dsize=(0,0), fx=r, fy=r, interpolation=cv2.INTER_NEAREST)
full = cv2.resize(full, dsize=(0,0), fx=r, fy=r, interpolation=cv2.INTER_NEAREST)
basic = cv2.resize(basic, dsize=(0,0), fx=r, fy=r, interpolation=cv2.INTER_NEAREST)
annot_ = cv2.copyMakeBorder(annot,10,10,10,10,cv2.BORDER_CONSTANT,value=(255,255,255))
fcn_ = cv2.copyMakeBorder(fcn,10,10,10,10,cv2.BORDER_CONSTANT,value=(255,255,255))
full_ = cv2.copyMakeBorder(full,10,10,10,10,cv2.BORDER_CONSTANT,value=(255,255,255))
basic_ = cv2.copyMakeBorder(basic,10,10,10,10,cv2.BORDER_CONSTANT,value=(255,255,255))
outimg = np.vstack([annot_, fcn_, full_, basic_])
outname = os.path.join(outpath, outname)
cv2.imwrite(outname, outimg)
# plt.imshow(outimg)
# -
y
y_ratio
print y, x
| notebooks/make_segmentation_figures.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Section 3.1: Linear regression, part 1
#
# Thus far in this track we have largely investigated just machine-learning (ML) classification techniques. With linear regression, we will look at one of the most important predictive algorithms in ML.
#
# The essence of linear regression is arguably the simplest form of ML: drawing a line through points. You might have done a simple form of this in your high school physics class: plot the results of a series of experiments on graph paper and then draw a line through as many points as possible (and include as many points below the line as above the line where the points don't fall on the line). That is a very form of linear regression. We will build on that conceptual foundation to address more complex situations, such as having points in more than two dimensions or even points whose relationship seems non-linear.
#
# Formally, linear regression is used to predict a quantitative *response* (the values on a Y axis) that is dependent on one or more *predictors* (values on one or more axes that are orthogonal to $Y$, commonly just thought of collectively as $X$). The working assumption is that the relationship between the predictors and the response is more or less linear. The goal of linear regression is to fit a straight line in the best possible way to minimize the deviation between our observed responses in the dataset and the responses predicted by our line, the linear approximation.
#
# How do we tell that we have the best fit possible for our line? The most common means of assessing the error between the fit of our line -- our model -- and the data is called the [***least squares method***](https://en.wikipedia.org/wiki/Least_squares). This method consists of minimizing the number you get when you square the differences between your predicted values (the line) and the actual values (the data) and add up all of those squared differences for your entire dataset.
#
# > **Learning goal:** By the end of this section, you should be comfortable fitting linear-regression models, and you should have some familiarity with interpreting their output.
#
# ## Load the data
#
# In this section and the next (3.2), we will use national statistics gathered by the United Nations (UN) from 2009-2011 (accessed from the United Nations Statistics Division's [Social indicators page](https://unstats.un.org/unsd/demographic/products/socind/) on April 23, 2012). The data includes national health and welfare statistics for 199 countries and territories; these locations are mostly UN members, but the list also includes other areas that are not independent countries (such as Hong Kong).
#
# The dataset includes 199 observations with the following features:
# - **region:** Region of the world
# - **group:** A factor (or categorical variable) with the following levels:
# - **oecd:** Countries that were members of the [Organisation for Economic Co-operation and Development](www.oecd.org) (OECD) as of May 25, 2012
# - **africa:** Countries on the continent of Africa (note: no OECD countries are located in Africa)
# - **other:** For all other countries
# - **fertility:** The total number of children born or likely to be born to a woman in her life time if she were subject to the prevailing rate of age-specific fertility in the population
# - **ppgdp:** Per-capita gross domestic product (GDP) in 2012 US dollars (measure of a country's economic output that accounts for its number of people.)
# - **lifeExpF:** Female life expectancy in years
# - **pctUrban:** Percent of the population urbanized (living in urban areas)
#
# We will need to load several modules for this section to handle the ML and visualizations.
import pandas as pd
import numpy as np
from sklearn.linear_model import LinearRegression
import matplotlib.pyplot as plt
# %matplotlib inline
# We'll then load the data.
df = pd.read_csv('UN11.csv')
df.head()
df['pctUrban'].min()
# ## Simple linear regression
#
# Let's plot out the data to see the relationship between per-capita GDP and female life expectancy.
plt.scatter(df['ppgdp'], df['lifeExpF'], alpha=0.3);
# > **Technical note:** The `alpha` parameter we supplied in the matplotlib `scatter` function; it makes the points semi-transparent so that we can where data points bunch up. Also note the semicolon at the end of the code snippet above; it silences the matplotlib memory-path output for cleaner inline graphing (without additional output above the graph, such as `<matplotlib.collections.PathCollection at 0x7f2c54737f28>`).
#
# Let's now plot a line and see what we get.
# +
model = LinearRegression(fit_intercept=True)
X = df['ppgdp'][:, np.newaxis]
y = df['lifeExpF']
model.fit(X, y)
x_plot = np.linspace(0, 100000)
y_plot = model.predict(x_plot[:, np.newaxis])
plt.scatter(df['ppgdp'], df['lifeExpF'], alpha=0.3)
plt.plot(x_plot, y_plot, c='orange');
# -
# > **Technical note:** Notice in the code cell above that we did not fit the model using `model.fit(df['ppgdp'], df['lifeExpF'])`. Instead, we had to use `df['ppgdp'][:, np.newaxis]` for our predictors rather than just `df['ppgdp']`. The addition of `[:, np.newaxis]` changes `df['ppgdp']` from a pandas `Series` to an array in matrix format. (If you're unsure what that looks like, create a new code cell below this cell using **Insert > Insert Cell Below** and then run `df['ppgdp']` and then `df['ppgdp'][:, np.newaxis]` in order to see the difference.
#
# Just how poor is this initial model? Let's calculate the $R^2$ score for it to see. The $R^2$ score (also called the [coefficient of determination](https://en.wikipedia.org/wiki/Coefficient_of_determination)) represents the proportion of the variance in our response that is predictable from the predictors -- so 0 is the worst (a model explains none of the variance) and 1 is the best (a model explains all of it).
# +
from sklearn.metrics import r2_score
model = LinearRegression(fit_intercept=True)
model.fit(df['ppgdp'][:, np.newaxis], df['lifeExpF'])
predictions = model.predict(df['ppgdp'][:, np.newaxis])
r2_score(df['lifeExpF'], predictions)
# -
# This first model accounts for only 30 percent of the variability in `lifeExpF` and is really not a very good representation of the relationship between economic activity and female life expectancy.
#
# These results are not good, which stems from the fact that there is no linear relationship between per-capita GDP and female life expectancy. Instead, the relationship has an elbow-like curve to it. When countries are very poor, the data suggests that even modest increases to GDP per capita can dramatically increase female life expectancy, but only up to a point; once countries hit about USD 10,000 per head, additional gains correlated to increases in wealth are much smaller. This suggests a logarithmic relationship between these factors: female life expectancy being not related to GPD, but to its logarithm.
#
# Let's create a new column that contains the logarithm of per-capita GDP by country. Note that, because we are dealing with powers of 10 in the GDP column, we will use the base-10 logarithm rather than the natural logarithm in order to make interpretation easier.
df['log_ppgdp'] = np.log10(df['ppgdp'])
df.head()
# Let's now plot our new `log_ppgdp` column against `lifeExpF` to see if there is a more linear relationship.
# +
model = LinearRegression(fit_intercept=True)
X = df['log_ppgdp'][:, np.newaxis]
y = df['lifeExpF']
model.fit(X, y)
x_min = df['log_ppgdp'].min()
x_max = df['log_ppgdp'].max()
x_plot = np.linspace(x_min, x_max)
y_plot = model.predict(x_plot[:, np.newaxis])
plt.scatter(df['log_ppgdp'], df['lifeExpF'], alpha=0.3)
plt.plot(x_plot, y_plot, c='orange');
# -
# This is much better, but it is still far from perfect. The shape of the data seems to have a curve to it and we will examine how to deal with that shortly. However, let's first interpret the model have right here to see what it tells us. How much better is it than the first model? Let's look at the $R^2$ score.
# +
model.fit(df['log_ppgdp'][:, np.newaxis], df['lifeExpF'])
predictions = model.predict(df['log_ppgdp'][:, np.newaxis])
r2_score(df['lifeExpF'], predictions)
# -
# Using `log_ppgdp` rather than `ppgdp` in the model roughly doubles the amount of variance in `lifeExpF` that we can account for with this model (to 60 percent). But what does our model actually mean?
print("Model slope: ", model.coef_[0])
print("Model intercept:", model.intercept_)
# Remember that in high school algebra lines were generally defined by an equation of the form
#
# $$
# y = ax + b
# $$
#
# where $a$ is the *slope* and $b$ is the *intercept*. That samer terminology applies in linear regression. The slope refers to our model's predicted change in units of female life expectancy (years) for each unit of the base-10 logarithm of per-capita GDP. In other words, our model predicts that, on average, women's life expectancies increase by 11.6 years every time per-capita GDP increases 10 fold.
#
# The intercept is a little more abstract because it is not directly tied to any data point. It shows the value of the $y$-axis at the point where our line crossed that axis (where $x=0$). If we were still modeling `ppgdp` versus `lifeExpF`, we might interpret the intercept as representing women's baseline life expectancy in a hypothetical country with a per-capita GDP of USD 0: 29.8 years. However, we are modeling `log_ppgdp` versus `lifeExpF`, and the logarithm of 0 is undefined. Therefore, it can be easiest to accept the intercept in our model as a mathematical abstraction necessary to making other parts of our model work. Our model can be stated as:
#
# $$
# {\rm lifeExpF} = 11.6 \times {\rm log\_ppgdp} + 29.8
# $$
# ## Polynomial regression
#
# We can generalize the line equation above in the form favored by statisticians:
#
# $$
# y = β_0 + β_1x + \epsilon
# $$
#
# where $\epsilon$ is an unobserved random error that we generally fold into $β_0$. Nothing says that we can have only one $x$ term, however. We can define a linear model for our data of the form
#
# $$
# y = β_0 + β_1x + β_2x^2 + \epsilon
# $$
#
# This is still a linear relationship because none of our $\beta$s ever multiply or divide each other. In fact, we can generalize linear models to the form
#
# $$
# y = β_0 + β_1x + β_2x^2 + β_3x^3 + \cdots + β_nx^n + \epsilon
# $$
#
# The linearity of our models depend on the linearity of $β_n$, not $x_n$. We will use this fact to use linear regression to model data that does not follow a straight line. Let's apply this to our model of `log_ppgdp` and `lifeExpF`.
# +
from sklearn.preprocessing import PolynomialFeatures
poly = PolynomialFeatures(degree=2)
X = df['log_ppgdp'][:, np.newaxis]
y = df['lifeExpF']
x_min = df['log_ppgdp'].min()
x_max = df['log_ppgdp'].max()
x_plot = np.linspace(x_min, x_max)
x_fit = poly.fit_transform(x_plot[:, np.newaxis])
X_ = poly.fit_transform(X)
poly_model = LinearRegression(fit_intercept=True)
poly_model.fit(X_, y)
x_fit[:, np.newaxis]
y_fit = poly_model.predict(x_fit)
plt.scatter(df['log_ppgdp'], df['lifeExpF'], alpha=0.3)
plt.plot(x_fit[:,1], y_fit, c='orange');
# -
# Adding the polynomial term provides us with a much more intuitive fit of the data! The `degree=2` parameter that we supply to the `PolynomialFeatures` function dictates that our model takes the form of
#
# $$
# y = β_0 + β_1x + β_2x^2
# $$
#
# Let’s see what the coefficients for our model are.
print("Model slope: ", poly_model.coef_)
print("Model intercept:", poly_model.intercept_)
#
# We can state our polynomial model as
#
# $$
# {\rm lifeExpF} = -6.5 + 32.1 \times {\rm log\_ppgdp} - 2.8 \times {\rm log\_ppgdp}^2
# $$
#
# Using the polynomial model improves predictive power, but it comes at the cost of interpretability. What is the intuitive relationship between `lifeExpF` and `log_ppgdp` now?
#
# > **Technical note:** Fitting the polynomial-regression model above has a lot of steps in it, and performing these transformations (transforming the features for polynomial regression and fitting the regression model) manually can quickly become tedious and error prone. To streamline this type of processing, scikit-learn provides the `Pipeline` object, which you can use to encapsulate several transformations into one step. Let's run this model again using scikit-learn `make_pipeline()`.
# +
from sklearn.pipeline import make_pipeline
poly_model = make_pipeline(PolynomialFeatures(2),
LinearRegression())
X = df['log_ppgdp'][:, np.newaxis]
y = df['lifeExpF']
poly_model.fit(X, y)
x_min = df['log_ppgdp'].min()
x_max = df['log_ppgdp'].max()
x_plot = np.linspace(x_min, x_max)
y_plot = poly_model.predict(x_plot[:, np.newaxis])
plt.scatter(df['log_ppgdp'], df['lifeExpF'], alpha=0.3)
plt.plot(x_plot, y_plot, c='orange');
# -
# That was much simpler to code! But how much did going through the work doing the polynomial regression help our model?
# +
poly_model = make_pipeline(PolynomialFeatures(2),
LinearRegression())
poly_model.fit(df['log_ppgdp'][:, np.newaxis], df['lifeExpF'])
predictions = poly_model.predict(df['log_ppgdp'][:, np.newaxis])
r2_score(df['lifeExpF'], predictions)
# -
# Our improved, polynomial model now accounts for 61.4 percent of the variance in `lifeExpF`. Clearly an improvement, but a modest one.
#
# > **Exercise**
# >
# > Go to the code cell in which we fitted the polynomial model using `make_pipeline()` and try different values (>2) in `PolynomialFeatures` to see what using higher-degree polynomials does for our model.
#
# > **Exercise solution**
# Here is a comparison of the outputs for models using three-degree, four-degree, and five-degree polynomials.
# +
colors = ['teal', 'yellowgreen', 'gold']
x_min = df['log_ppgdp'].min()
x_max = df['log_ppgdp'].max()
x_plot = np.linspace(x_min, x_max)
plt.scatter(df['log_ppgdp'], df['lifeExpF'], alpha=0.3, c='gray')
for count, degree in enumerate([3, 4, 5]):
model = make_pipeline(PolynomialFeatures(degree),
LinearRegression())
X = df['log_ppgdp'][:, np.newaxis]
y = df['lifeExpF']
model.fit(X, y)
y_plot = model.predict(x_plot[:, np.newaxis])
plt.plot(x_plot, y_plot, color=colors[count],
linewidth=2, label="Degree %d" % degree)
plt.legend(loc='lower right')
plt.show();
# -
# Let’s see what the $R^2$ scores for the different-degree polynomial models are.
for count, degree in enumerate([3, 4, 5]):
model = make_pipeline(PolynomialFeatures(degree),
LinearRegression())
X = df['log_ppgdp'][:, np.newaxis]
y = df['lifeExpF']
model.fit(X, y)
predictions = model.predict(X)
print("Degree %d" % degree, "r-squared score:",
r2_score(df['lifeExpF'], predictions))
# Each additional polynomial degree improves the fit of our model (as demonstrated by the incremental improvements to the r-squared scores). However, adding more degrees to the polynomial regressions opens us to the risk of [overfitting](https://en.wikipedia.org/wiki/Overfitting), a process by which our models come to fit the training data too closely and are thus less useful in predicting more generalized data.
#
# Higher-degree polynomials also bring back the [curse of dimensionality](https://en.wikipedia.org/wiki/Curse_of_dimensionality). Simple linear models need only $N + 1$ sample points to fit, where $N$ is the number of dimensions (2 points in 1 dimension, 3 in 2 dimensions, 4 in three dimensions, and so on). However, each additional polynomial degree increases the number of sample points required for a given dimensionality much faster. Particularly if certain data points are difficult or expensive to come by, you might run out of data in order to fit a high-degree polynomial model.
# ## Multiple regression
#
# Ultimately, no matter how complex the model we construct between `log_ppgdp` and `lifeExpF`, we will only be able to explain so much of variability because factors other than per-capita GDP affect female life expectancy. Using more than one predictor in our regression model helps us capture more of this richness of detail.
#
# Let's start by plotting the relationship between log per-capita GDP, urbanization, and female life expectancy in three dimensions.
# +
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
# %matplotlib inline
fig = plt.figure(figsize=(6,6))
ax = fig.add_subplot(111, projection='3d')
ax.scatter(df['log_ppgdp'], df['pctUrban'], df['lifeExpF'])
ax.set_xlabel('GDP per capita (log)')
ax.set_ylabel('Percent urbanized')
ax.set_zlabel('Life expectancy (years)')
plt.show();
# +
model = LinearRegression(fit_intercept=True)
X = df[['log_ppgdp', 'pctUrban']]
y = df['lifeExpF']
model.fit(X, y)
x1_plot = np.linspace(df['log_ppgdp'].min(), df['log_ppgdp'].max())
x2_plot = np.linspace(df['pctUrban'].min(), df['pctUrban'].max())
X1_plot, X2_plot = np.meshgrid(x1_plot, x2_plot)
y_plot = model.predict(np.c_[X1_plot.ravel(), X2_plot.ravel()])
fig = plt.figure(figsize=(6,6))
ax = fig.add_subplot(111, projection='3d')
ax.scatter(df['log_ppgdp'], df['pctUrban'], df['lifeExpF'])
ax.plot_surface(X1_plot, X2_plot, y_plot.reshape(X1_plot.shape), cmap='viridis');
ax.set_xlabel('GDP per capita (log)')
ax.set_ylabel('Percent urbanized')
ax.set_zlabel('Life expectancy (years)')
plt.show();
# -
# How accurate is our multiple-regression model?
# +
model = LinearRegression(fit_intercept=True)
X = df[['log_ppgdp', 'pctUrban']]
y = df['lifeExpF']
model.fit(X, y)
predictions = model.predict(X)
r2_score(df['lifeExpF'], predictions)
# -
# This model explains 59.8 percent of the variance in `lifeExpF`: better than our initial simple linear model ($R^2=$ 0.596), but not spectacularly so.
#
# What does this new model mean?
print("Model slopes: ", model.coef_)
print("Model intercept:", model.intercept_)
# Our model now has two predictors in it, so it takes the generalized form:
#
# $$
# y = β_0 + β_1x_1 + β_2x_2
# $$
#
# Specifically, our model is:
#
# $$
# {\rm lifeExpF} = 30.7 + 11 \times {\rm log\_ppgdp} + 0.023 \times {\rm pctUrban}
# $$
#
# Multiple regression is a little trickier to interpret than simple regression, but not enormously so. Our model says that if we were to hold all other factors equal, then increasing the per-capita GDP of a country 10 fold will (on average) add 11 years to women's life expectancy. It also says that if we keep everything else the same, then increasing the urbanization of a country by 1 percent will increase women's life expectancy by 0.023 years. (Remember that we can’t think of the intercept as representing a hypothetical baseline country with USD0 GDP and 0 urbanization, because the logarithm of 0 is undefined.) This is another way of showing that adding `pctUrban` to our model provides some additional predictive power to our simple model, but not much. But does it do anything if we add it to a polynomial model?
# +
poly_model = make_pipeline(PolynomialFeatures(2),
LinearRegression())
X = df[['log_ppgdp', 'pctUrban']]
y = df['lifeExpF']
poly_model.fit(X, y)
x1_plot = np.linspace(df['log_ppgdp'].min(), df['log_ppgdp'].max())
x2_plot = np.linspace(df['pctUrban'].min(), df['pctUrban'].max())
X1_plot, X2_plot = np.meshgrid(x1_plot, x2_plot)
y_plot = poly_model.predict(np.c_[X1_plot.ravel(), X2_plot.ravel()])
fig = plt.figure(figsize=(6,6))
ax = fig.add_subplot(111, projection='3d')
ax.scatter(df['log_ppgdp'], df['pctUrban'], df['lifeExpF'])
ax.plot_surface(X1_plot, X2_plot, y_plot.reshape(X1_plot.shape), cmap='viridis');
ax.set_xlabel('GDP per capita (log)')
ax.set_ylabel('Percent urbanized')
ax.set_zlabel('Life expectancy (years)')
plt.show();
# -
# Let’s take a look at the $R^2$ for this model.
# +
poly_model = make_pipeline(PolynomialFeatures(2),
LinearRegression())
X = df[['log_ppgdp', 'pctUrban']]
y = df['lifeExpF']
poly_model.fit(X, y)
predictions = poly_model.predict(X)
r2_score(df['lifeExpF'], predictions)
# -
# In the polynomial regression, adding `pctUrban` to our model provides a decent improvement to our model's predictive power (for example, this model's $R^2$ score is higher than those that we got with our two-degree, three-degree, or four-degree models using just `log_ppgdp`).
#
# More than just boosting the $R^2$ score, fitting the multiple polynomial regression provides additional insights from the visualization. If you rotate the visualization above 180 degrees about the $z$-axis, you will notice that while our model predicts increased female life expectancy at high incomes, in poor countries, our model actually shows a *decrease* in female life expectancy in poor countries correlated with increased urbanization.
#
# All of these conclusions come from a model that treats all of the data as coming from a rather monolithic whole. We have other types of data that we can also use in our modeling to try and arrive at different insights.
# ## Categorical data
#
# Our dataset has two categorical features (also known as *factors* in the statistical world): `region` and `group`. There are multiple ways of handling data like this in linear regression; here, we will handle it by building sub-models for it.
#
# To begin moving in that analytical direction, let's start by color coding our 3D scatterplot points by `group`.
# +
groups = df.groupby('group')
fig = plt.figure(figsize=(6,6))
ax = fig.add_subplot(111, projection='3d')
for name, group in groups:
ax.scatter(group['log_ppgdp'],
group['pctUrban'],
group['lifeExpF'], label=name)
ax.legend()
ax.set_xlabel('GDP per capita (log)')
ax.set_ylabel('Percent urbanized')
ax.set_zlabel('Life expectancy (years)')
ax.legend()
plt.show();
# -
# Unsurprisingly, OECD-member countries cluster at the high end of the income scale and, sadly, African countries lag at the poorer end. Countries from the `other` group include countries ranging from poor ones in Southeast Asia to oil-rich Middle Eastern countries, and thus countries from that group are scattered across the income spectrum.
#
# Now that we have the data points detailed in color, let's fit three separate, simple linear models for each group of countries.
# +
model = LinearRegression(fit_intercept=True)
fig = plt.figure(figsize=(6,6))
ax = fig.add_subplot(111, projection='3d')
groups = df.groupby('group')
cmaps = ['Blues', 'Oranges', 'Greens']
for name, group in groups:
X = group[['log_ppgdp', 'pctUrban']]
y = group['lifeExpF']
model.fit(X, y)
x1_plot = np.linspace(group['log_ppgdp'].min(), group['log_ppgdp'].max())
x2_plot = np.linspace(group['pctUrban'].min(), group['pctUrban'].max())
X1_plot, X2_plot = np.meshgrid(x1_plot, x2_plot)
y_plot = model.predict(np.c_[X1_plot.ravel(), X2_plot.ravel()])
ax.scatter(group['log_ppgdp'], group['pctUrban'], group['lifeExpF'], label=name)
cmap_index = sorted(df['group'].unique().tolist()).index(name)
cmap = cmaps[cmap_index]
ax.plot_surface(X1_plot, X2_plot, y_plot.reshape(X1_plot.shape), cmap=cmap);
ax.set_xlabel('GDP per capita (log)')
ax.set_ylabel('Percent urbanized')
ax.set_zlabel('Life expectancy (years)')
plt.show();
# -
# > **Technical note:** The blue plane models the `africa` group, the orange one `oecd`, and `other` is green.
#
# The opacity of the various planes can make it hard to pick out the details, but if you rotate the graph, you should be able to see that while the `other` and `oecd` models behave similarly, the `africa` sub-model exhibits different behavior and responds more dramatically to increases in per-capita GDP and urbanization for increasing women's lifespans.
#
# > **Exercise**
# >
# > Changing color map for 3D plots like these can sometime help make different details clearer. Locate the `cmap` parameter in the `plot_surface()` function and change it to `cmap='viridis'`.
#
# > **Exercise solution**
# +
model = LinearRegression(fit_intercept=True)
fig = plt.figure(figsize=(6,6))
ax = fig.add_subplot(111, projection='3d')
groups = df.groupby('group')
cmaps = ['Blues', 'Oranges', 'Greens']
for name, group in groups:
X = group[['log_ppgdp', 'pctUrban']]
y = group['lifeExpF']
model.fit(X, y)
x1_plot = np.linspace(group['log_ppgdp'].min(), group['log_ppgdp'].max())
x2_plot = np.linspace(group['pctUrban'].min(), group['pctUrban'].max())
X1_plot, X2_plot = np.meshgrid(x1_plot, x2_plot)
y_plot = model.predict(np.c_[X1_plot.ravel(), X2_plot.ravel()])
ax.scatter(group['log_ppgdp'], group['pctUrban'], group['lifeExpF'], label=name)
ax.plot_surface(X1_plot, X2_plot, y_plot.reshape(X1_plot.shape), cmap='viridis');
ax.set_xlabel('GDP per capita (log)')
ax.set_ylabel('Percent urbanized')
ax.set_zlabel('Life expectancy (years)')
plt.show();
# -
# How good are these models?
# +
groups = df.groupby('group')
for name, group in groups:
X = group[['log_ppgdp', 'pctUrban']]
y = group['lifeExpF']
model.fit(X, y)
predictions = model.predict(X)
print(name, "r-squared score:",
r2_score(group['lifeExpF'],
predictions))
# -
# These models are not great. But we will see if they are improved by using polynomial regression later on.
# +
model = LinearRegression(fit_intercept=True)
groups = df.groupby('group')
for name, group in groups:
X = group[['log_ppgdp', 'pctUrban']]
y = group['lifeExpF']
model.fit(X, y)
print(name, "slopes: ", model.coef_)
print(name, "intercept:", model.intercept_)
# -
# What do these plots based on group tell us? The slopes for `log_ppgdp` are similar between the `africa` and `oecd` groups, but the slope is different for the `other` group. This suggests that there might be some interaction between `group` and `log_ppgdp` in explaining `lifeExpF`. The `pctUrban` slope for the `oecd` group has a different sign than for `africa` or `other`, which might indicate another interaction, but we are on shaky statistical ground here because we have done no testing to see if these differences in slope are statistically significant (for `pctUrban`, the differences—and the numbers—are small).
#
# What do we mean by interactions between groups? Recall that we generalized a linear model with two features as:
#
# $$
# y=β_0+β_1 x_1+β_2 x_2
# $$
#
# However, if $x_1$ and $x_2$ interact—if different values of $x_1$, for example, change the influence of $x_2$ on $y$ — we need to include that in the model like so:
#
# $$
# y=β_0+β_1 x_1+β_2 x_2+β_3x_1x_2
# $$
#
# Our model involves three predictors (`log_ppgdp`, `pctUrban`, and `group`) and has this form:
#
# $$
# y=β_0+β_1 x_1+β_2 x_2+β_3 u_2+β_4 u_3+β_5x_1x_2+β_6 x_1 u_2+β_7 x_1 u_3+β_8 x_2 u_2+β_9 x_2 u_3+β_{10} x_1x_2u_2+β_{11}x_1x_2 u_3
# $$
#
# Think of this as another aspect of the curse of dimensionality: as we add features (especially categorical ones), the number of potential interactions between features that we have to account for increases even faster.
#
# > **Technical note:** Statisticians often use the variable u for categorical features, a convention that we have used here. Also note that we only included $u_2$ and $u_3$ in the generalized equation for the model, even though we have three groups in the categorical feature group. This is not a mistake; it is because one group from the categorical feature gets included in the intercept.
#
#
# > **Question**
# >
# > Do you see where these numbers come from? What do you think the intercepts indicate for each of these groups?
#
# Let's now see what happens when we plot polynomial models for each of these groups.
# +
poly_model = make_pipeline(PolynomialFeatures(2),
LinearRegression())
fig = plt.figure(figsize=(6,6))
ax = fig.add_subplot(111, projection='3d')
groups = df.groupby('group')
cmaps = ['Blues_r', 'Oranges_r', 'Greens_r']
for name, group in groups:
X = group[['log_ppgdp', 'pctUrban']]
y = group['lifeExpF']
poly_model.fit(X, y)
x1_plot = np.linspace(group['log_ppgdp'].min(), group['log_ppgdp'].max())
x2_plot = np.linspace(group['pctUrban'].min(), group['pctUrban'].max())
X1_plot, X2_plot = np.meshgrid(x1_plot, x2_plot)
y_plot = poly_model.predict(np.c_[X1_plot.ravel(), X2_plot.ravel()])
ax.scatter(group['log_ppgdp'], group['pctUrban'], group['lifeExpF'], label=name)
cmap_index = sorted(df['group'].unique().tolist()).index(name)
cmap = cmaps[cmap_index]
ax.plot_surface(X1_plot, X2_plot, y_plot.reshape(X1_plot.shape), cmap=cmap);
ax.set_xlabel('GDP per capita (log)')
ax.set_ylabel('Percent urbanized')
ax.set_zlabel('Life expectancy (years)')
plt.show();
# -
# The differences in shapes for these surfaces suggest interaction between `log_ppgdp`, `pctUrban`, and `group`. However, insightful as these plots can be, the nature of 3D visualization can make them hard to see. Another way to present the data is by breaking each model into its own subplot.
# +
poly_model = make_pipeline(PolynomialFeatures(2),
LinearRegression())
fig = plt.figure(figsize=(18, 6))
groups = df.groupby('group')
cmaps = ['Blues_r', 'Oranges_r', 'Greens_r']
colors = ['blue', 'orange', 'green']
for name, group in groups:
X = group[['log_ppgdp', 'pctUrban']]
y = group['lifeExpF']
poly_model.fit(X, y)
x1_plot = np.linspace(group['log_ppgdp'].min(), group['log_ppgdp'].max())
x2_plot = np.linspace(group['pctUrban'].min(), group['pctUrban'].max())
X1_plot, X2_plot = np.meshgrid(x1_plot, x2_plot)
y_plot = poly_model.predict(np.c_[X1_plot.ravel(), X2_plot.ravel()])
index = sorted(df['group'].unique().tolist()).index(name)
ax = fig.add_subplot(1, 3, index + 1, projection='3d')
color = colors[index]
ax.scatter(group['log_ppgdp'], group['pctUrban'], group['lifeExpF'],
label=name, c=color)
cmap = cmaps[index]
ax.plot_surface(X1_plot, X2_plot, y_plot.reshape(X1_plot.shape), cmap=cmap);
ax.set_title(name)
ax.set_xlabel('GDP per capita (log)')
ax.set_ylabel('Percent urbanized')
ax.set_zlabel('Life expectancy (years)')
plt.show();
# -
# How useful are these models for prediction? Let's look at the $R^2$ scores.
# +
poly_model = make_pipeline(PolynomialFeatures(2),
LinearRegression())
groups = df.groupby('group')
for name, group in groups:
X = group[['log_ppgdp', 'pctUrban']]
y = group['lifeExpF']
poly_model.fit(X, y)
predictions = poly_model.predict(X)
print(name, "r-squared score:",
r2_score(group['lifeExpF'],
predictions))
# -
# Not uniformly good. Adding polynomial regression improved the model for the `oecd` group, but worsened it for `africa` and `other`. Let's see if increasing the degrees of the polynomials helps.
#
# > **Exercise**
# >
# > Try re-running the $R^2$ scoring code cell above using different polynomial degrees in the `PolynomialFeatures()` function until you get better-fitting models.
#
# > **Possible exercise solution**
# +
poly_model = make_pipeline(PolynomialFeatures(5),
LinearRegression())
groups = df.groupby('group')
for name, group in groups:
X = group[['log_ppgdp', 'pctUrban']]
y = group['lifeExpF']
poly_model.fit(X, y)
predictions = poly_model.predict(X)
print(name, "r-squared score:",
r2_score(group['lifeExpF'],
predictions))
# -
# > **Exercise**
# >
# > Now that you have a better polynomial degree to use in the models, re-run the code to plot them to see what they look like.
#
# > **Exercise solution**
# +
poly_model = make_pipeline(PolynomialFeatures(5),
LinearRegression())
fig = plt.figure(figsize=(18, 6))
groups = df.groupby('group')
cmaps = ['Blues_r', 'Oranges_r', 'Greens_r']
colors = ['blue', 'orange', 'green']
for name, group in groups:
X = group[['log_ppgdp', 'pctUrban']]
y = group['lifeExpF']
poly_model.fit(X, y)
x1_plot = np.linspace(group['log_ppgdp'].min(), group['log_ppgdp'].max())
x2_plot = np.linspace(group['pctUrban'].min(), group['pctUrban'].max())
X1_plot, X2_plot = np.meshgrid(x1_plot, x2_plot)
y_plot = poly_model.predict(np.c_[X1_plot.ravel(), X2_plot.ravel()])
index = sorted(df['group'].unique().tolist()).index(name)
ax = fig.add_subplot(1, 3, index + 1, projection='3d')
color = colors[index]
ax.scatter(group['log_ppgdp'], group['pctUrban'], group['lifeExpF'],
label=name, c=color)
cmap = cmaps[index]
ax.plot_surface(X1_plot, X2_plot, y_plot.reshape(X1_plot.shape), cmap=cmap);
ax.set_title(name)
ax.set_xlabel('GDP per capita (log)')
ax.set_ylabel('Percent urbanized')
ax.set_zlabel('Life expectancy (years)')
plt.show();
# -
# The differences in the shapes of these surfaces suggest interactions between `log_ppgdp`, `pctUrban`, and `group`, but we would have to do additional tests to establish what the specifics of those interactions are.
#
# > **Question**
# >
# > What do these plots tell you about the dangers of extrapolating too much from a model? Is overfitting a possible concern with these tightly fit models?
#
# > **Takeaway**
# >
# > Linear regression can be a flexible tool for modeling the relationships between features in our data, particularly with polynomial regression. However, this flexibility comes with dangers, particularly the hazard of overfitting our models to our data. With multiple regression, we can produce richer models that include more relationships, but interpretation becomes murkier with each additional feature, particularly when categorical features are included.
| online-event-resources/data-science-and-machine-learning/regression/Regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="mMbSCJGsDSKu"
# [][#gh-colab-quickstart]
# [][#gh-binder-quickstart]
# [][#gh-sagemaker-studiolab-quickstart]
# [][#gh-deepnote-quickstart]
# [][#gh-kaggle-quickstart]
#
# [#gh-colab-quickstart]: https://colab.research.google.com/github/sugatoray/genespeak/blob/master/notebooks/quickstart_genespeak.ipynb
#
# [#gh-binder-quickstart]: https://mybinder.org/v2/gh/sugatoray/genespeak/master?labpath=notebooks%2Fquickstart_genespeak.ipynb
#
# [#gh-sagemaker-studiolab-quickstart]: https://studiolab.sagemaker.aws/import/github/sugatoray/genespeak/blob/master/notebooks/quickstart_genespeak.ipynb
#
# [#gh-deepnote-quickstart]: https://deepnote.com/viewer/github/sugatoray/genespeak/blob/master/notebooks/quickstart_genespeak.ipynb
#
# [#gh-kaggle-quickstart]: https://kaggle.com/kernels/welcome?src=https://github.com/sugatoray/genespeak/blob/master/notebooks/quickstart_genespeak.ipynb
# + [markdown] id="YNsaOUzthiaP"
# ## Install `genespeak`
# + id="aNrOIBDAbzMh"
# ! pip install -Uqq genespeak
# + [markdown] id="bQaOCl1uhnsq"
# ## Import `genespeak` and check version
# + colab={"base_uri": "https://localhost:8080/"} id="bLyHCLGpcVs2" outputId="9fadb71a-22c7-41cf-b66b-26c2aa699251"
import genespeak as gp
print(f'{gp.__name__} version: {gp.__version__}')
# + [markdown] id="g3mUn9LWj0az"
# ## Usage
# + [markdown] id="dcvQ8IFPhIX9"
# ### A. Handle `ascii` text
#
# If your text does not contain non-english text or emojis, it is possible to use any strategy (`utf-8` or `ascii`) to properly convert the text into synthetic dna-single-strand. In case, the text is *ascii-only*, it is preferrable to use `ascii` strategy. In the following example, we have used `schema = "ATCG"`. The same *schema* and *strategy* pair is necessary for **text-to-dna** or **dna-to-text** conversion.
# + colab={"base_uri": "https://localhost:8080/"} id="WS3nPmj-cBw4" outputId="14b0d9e2-fa08-4176-fefc-171765432c73"
schema = "ATCG" # (1)
strategy = "ascii" # (2)
text = "Hello World!"
dna = gp.text_to_dna(text, schema=schema)
text_from_dna = gp.dna_to_text(dna, schema=schema)
print(f'Text: {text}\nEncoded DNA: {dna}\nDecoded Text: {text_from_dna}\nSuccess: {text == text_from_dna}')
# + [markdown] id="m5Z7Ayw-dKoi"
# ### B. Handle `utf-8` text
#
# If your text contains non-english text or emojis, it is necessary to use `strategy = "utf-8"` to properly convert the text into synthetic dna-single-strand. In the following example, we have used `schema = "ACGT"`. Same *schema* and *strategy* pair is necessary for **text-to-dna** or **dna-to-text** conversion.
# + colab={"base_uri": "https://localhost:8080/"} id="81b2UwuWch0x" outputId="4714cb55-a648-48c0-8575-96ab97d1efe9"
schema = "ACGT" # (1)
strategy = "utf-8" # (2)
text = "Hello World 😀!"
dna = gp.text_to_dna(text, schema=schema, strategy=strategy)
text_from_dna = gp.dna_to_text(dna, schema=schema, binary_string_length=24, strategy=strategy)
print(f'Text: {text}\nEncoded DNA: {dna}\nDecoded Text: {text_from_dna}\nSuccess: {text == text_from_dna}')
# + [markdown] id="nyPbHPUUjZIw"
# ## Have fun playing with `genespeak` ! 🚀 🤗
# + id="tVW4QCULgPRs"
| notebooks/quickstart_genespeak.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# default_exp direction
# -
# # Direction
#
# > A cyclic direction class to make rotations easier.
#hide
from nbdev.showdoc import *
#export
class Direction:
"""
A cyclic direction class to make rotation easier.
The directions are written in a specific order of North, East, South, West
instead of North, South, East, West to make rotation simpler.
To rotate the direction clockwise, this class simply updates the `idx` to `idx+1`.
To rotate the direction clockwise, this class simply updates the `idx` to `idx-1`.
Returns:
An instance of itself with the updated direction on calling either `left` or `right`.
"""
def __init__(self, directions=['NORTH', 'EAST', 'SOUTH', 'WEST'], idx=None):
# get random direction if not passed
if idx is None:
idx = np.random.choice(np.arange(4))
self.directions = directions
self.max_idx = len(directions)-1
self.f = self.directions[idx]
def get_idx(self):
"return the current direction index from `self.directions`."
return self.directions.index(self.f)
def left(self):
"first get current direction index, then subtract 1 to rotate left."
idx = self.get_idx()
new_idx = idx-1 if idx>=1 else self.max_idx
return self.__class__(idx=new_idx)
def right(self):
"first get current direction index, then add 1 to rotate right."
idx = self.get_idx()
new_idx = idx+1 if idx<=2 else 0
return self.__class__(idx=new_idx)
def __repr__(self):
return self.f
d = Direction(idx=0)
#currently facing north
assert d.get_idx()==0
assert repr(d.f) == "'NORTH'"
#calling left changes idx to 3 and str representation to 'WEST'
assert d.left().get_idx()==3
assert repr(d.left().f) == "'WEST'"
#can do multiple lefts or left and right combination
assert d.left().left().get_idx()==2
assert d.left().right().get_idx()==0
from nbdev.export import *
notebook2script()
| 01_direction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:anaconda3]
# language: python
# name: conda-env-anaconda3-py
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
import glob
import h5py
from astropy.io import fits
from PIL import Image
from scipy.misc import toimage
import pandas as pd
import seaborn; seaborn.set() # set plot styles
import sys
sys.path.insert(0,'../rficnn/')
import rficnn as rfc
# %matplotlib inline
# -
sim_files = glob.glob('/home/anke/HIDE_simulations/hide_sims_train/calib_1year/*.fits') + glob.glob('/home/anke/HIDE_simulations/hide_sims_train/calib_1year/*.h5')
l = len(sim_files)
file_set = sim_files[:20]
l
nx,ny = 276, 14400
dp = rfc.read_part_chunck_sdfits(file_set[0],label_tag='RFI')
dp = np.array(dp)
data, rfi = dp
def ecdf(file):
ext = file.split('.')[-1]
if ext == 'fits':
data, rfi = rfc.read_part_chunck_sdfits(file, label_tag='RFI')
elif ext == 'h5':
data, rfi = rfc.read_part_chunck_hdf5(filename, label_tag)
rfi[np.logical_and(rfi<0,data<0)] = 0
data[np.logical_and(rfi<0,data<0)] = 0
rfi[rfi<0] = 0
rfi[data<0] = 0
data[data<0] = 0
rfi = np.array(rfi)
rfi = np.reshape(rfi,-1)
count = len(rfi)
rfi_sorted = np.sort(rfi)
y = np.arange(1, len(rfi_sorted)+1)/count # fraction of the array
return rfi_sorted,y
# +
# find 2 threshold
rfi[np.logical_and(rfi<0,data<0)] = 0
data[np.logical_and(rfi<0,data<0)] = 0
rfi[rfi<0] = 0
rfi[data<0] = 0
data[data<0] = 0
rfi = np.array(rfi)
rfi = np.reshape(rfi,-1)
count = len(rfi)
rfi_sorted = np.sort(rfi)
y = np.arange(1, len(rfi_sorted)+1)/count
# -
def find_nearest(array, value):
array = np.asarray(array)
idx = (np.abs(array - value)).argmin()
return idx
# +
def find_threshold(rfi_sorted, y, values):
thresholds = []
for i in values:
thresholds.append(rfi_sorted[find_nearest(y, i)])
return thresholds
rfi_sorted[find_nearest(y, 0.333)]
# -
find_threshold(rfi_sorted, y, [0.333,0.666])
def plot_rfi_ecdf(file):
'''Plots the Empirical Cumulative Distribution Function for all the RFI
Able to read off the thresholds'''
ext = file.split('.')[-1]
if ext == 'fits':
rfi = rfc.read_part_chunck_sdfits(file, label_tag='RFI')[1]
elif ext == 'h5':
rfi = rfc.read_part_chunck_hdf5(filename, label_tag)[1]
rfi[np.logical_and(rfi<0,data<0)] = 0
data[np.logical_and(rfi<0,data<0)] = 0
rfi[rfi<0] = 0
rfi[data<0] = 0
data[data<0] = 0
rfi = np.array(rfi)
rfi = np.reshape(rfi,-1)
count = len(rfi) # amount of data in array that have value bigger than zero
rfi_sorted = np.sort(rfi)
y = np.arange(1, len(rfi_sorted)+1)/count
import matplotlib.pyplot as plt
#plt.figure(figsize=(8,6))
plt.title('ECDF')
plt.xlabel('RFI amplitude (Jy)')
plt.ylabel('Fraction of dataset')
plt.xscale('log')
plt.margins(0.02)
plt.plot(rfi_sorted, y, marker='.', linestyle=None)
plt.savefig('ECDF.png', format='png')
for file in file_set[0:3]:
plot_rfi_ecdf(file)
hist, bin_edges = np.histogram(rfi_sorted[~rfi_sorted.mask],bins='auto')
plt.bar(bin_edges[:-1], hist, width = 1)
#plt.xlim(min(bin_edges), max(bin_edges))
# ---
# +
# Alireza code
bins = 100
min_ratio= 1e-4
max_ratio=np.inf
rfi_ratio=abs(100*rfi/data) # percentage rfi
rfi_accept = (min_ratio<rfi_ratio) & (rfi_ratio<max_ratio) #
print(rfi_accept)
'''Return a copy of the array collapsed into one dimension.'''
r2d = rfi_ratio[rfi_accept].flatten() #np.clip(.reshape(-1),0,200)
r=[1,200]
hist, bin_edges = np.histogram(r2d,bins=bins,range=r)
print( 'percentage of pixels in range: ',r,'is', 100.0*hist.sum()/float(len(r2d)) )
bin_cents = [0.5*(bin_edges[i]+bin_edges[i+1]) for i in range(bins)]
plt.xlabel('Percentage RFI')
plt.ylabel('Cumulative Sum')
plt.plot(bin_cents,np.cumsum(hist))
plt.xscale('log')
plt.yscale('log')
# +
def plot_rfi_cdf(file,min_ratio=1e-4, max_ratio=np.inf, bins=100, r=[1,2]):
'''Plots the Cumulative Distribution function of RFI percentage'''
h = read_chunck_sdfits_modified(file, label_tag='RFI')
h = np.array(h)
rfi = h[1]
data = h[0]
per_rfi = abs(100*rfi/data)
#per_rfi = np.reshape(per_rfi,(-1))
per_rfi_sorted = np.sort(per_rfi)
rfi_accept = (min_ratio<per_rfi) & (per_rfi<max_ratio)
r2d = rfi_ratio[rfi_accept].flatten() #np.clip(.reshape(-1),0,200)
hist, bin_edges = np.histogram(r2d,bins=bins,range=r)
print( 'percentage of pixels in range: ',r,'is', 100.0*hist.sum()/float(len(r2d)) )
bin_cents = [0.5*(bin_edges[i]+bin_edges[i+1]) for i in range(bins)]
plt.xlabel('Percentage RFI')
plt.ylabel('Cumulative Sum')
plt.plot(bin_cents,np.cumsum(hist))
plt.xscale('log')
plt.yscale('log')
# -
# ---
| notebooks/backup/cdf_plots.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import geopandas as gpd
from shapely import wkt
from matplotlib import pyplot
import descartes
import seaborn as sns
import datetime as dt
from sklearn.preprocessing import MultiLabelBinarizer
from statsmodels.stats.outliers_influence import variance_inflation_factor as vif
airbnb = pd.read_csv('../data/preprocessing/airbnb.csv')
airbnb.info()
targets= ['price_person','middle_estimate/month', 'lowest_estimate/month','price']
#this removes features that I won't use for modeling. I'll keep them in a separate dataframe incase I want to use
#spacial information or NLP in the future.
reserved_features = ['the_geom', 'geometry','OBJECTID', 'geoid', 'index_right','id', 'latitude', 'longitude', 'name', 'description', 'neighborhood_overview', 'host_name', 'host_location', 'host_about']
reserved_df = airbnb[reserved_features]
airbnb = airbnb.drop(columns = reserved_features)
# # Dummy Variables and One Hot Encoding
#
# The first thing I want to do is make a dataframe for linear regression and one for tree based ensembles. For linear regression, the categorical columns will need to be turned into multiple columns for each category. One Hot Encoding does this by turning each category into its own column and Get Dummies does this by making a column for each category - 1. For categories that are mutually exclusive I should use Dummy Variables and for categories that aren't, I'll use one hot encoding.
# ## One Hot Encoding for Amenities
#
# I'll use multiplabelbinarizer from sklearn preprocessing to make a column for each amenity listed and fill in 1s for airbnbs with that amenity and fill in 0s for aibnbs without. Since each instance can contain any combination of amenities, this will be like one hot encoding.
airbnb.amenities.head()
#prepping amenities colulumn to work with multilabelbinarizer
#binarizer method works by separating the words in one string into categories that do not repeat.
#so I have to remove:
#the brakets
airbnb['amenities']=airbnb['amenities'].map(lambda x: x.lstrip('[').rstrip(']'))
#the quotation marks
airbnb['amenities']=airbnb['amenities'].map(lambda x: x.replace('"',''))
#and leading whitespaces
airbnb['amenities']=airbnb['amenities'].map(lambda x: x.replace(', ',','))
print(airbnb.amenities[:5])
#creates a new data frame of amenities as separate columns with boolean values
mlb = MultiLabelBinarizer()
amenities_df = pd.DataFrame(mlb.fit_transform(airbnb['amenities'].str.split(',')),columns=mlb.classes_)
print(amenities_df.info())
#checks to see if any amenities are used by each airbnb and therefore have no variance
col_list=[]
for col in amenities_df.columns:
if amenities_df[col].nunique()<2:
col_list.append(col)
print("There are " +str(len(col_list)) + ' columns with fewer than 2 unique values')
#joins main dataframe with amenities dataframe with a prefix.
airbnb = airbnb.reset_index()
airbnb = airbnb.join(amenities_df.add_prefix('amen_'))
#
# ## Finding and Transforming Other Categorical Features
#
# +
#first change boolean columns to integers
boolean = airbnb.select_dtypes(bool).columns
for col in airbnb[boolean]:
airbnb[col]=np.multiply(airbnb[col], 1)
#then make a dataframe to look at columns that need to be encoded by removing numeric
numeric = airbnb.select_dtypes(np.number).columns
other = airbnb.drop(columns = numeric)
other.dtypes
# -
#dropping amenities since it already has been encoded.
other.drop(columns = 'amenities', inplace = True)
#making dummy columns with prefixes
dummy_columns = other.columns
prefixes = ['resp_tim_','nbhd_','prop']
prefixes = dict(zip(dummy_columns, prefixes))
other = pd.get_dummies(data = other, prefix = prefixes, drop_first = True)
linear_data = airbnb[numeric].join(other)
linear_data.drop(columns = 'index', inplace = True)
print(linear_data.shape)
linear_data.head()
print(airbnb.shape)
airbnb.head()
# ## Data Split for Decision Tree or Linear Regression
# The linear_data dataset is now completely encoded for linear regression.
# The airbnb dataset can be used for decision tree type regressions such as the random forest regressor. The amenities are encoded, but the other categorical and boolean features are kept as they are.
#
#
#
#
#
#
#
#
#
# # Feature Selection
#
# Encoding created a lot of features. The following will try to address feature selection. I would like to select features rather than extract with PCA. I've chosen homeowners as my target audience and I want to stick with methods that easily show what aspects of a rental affect its price.
# ## Colinearity
#
# Linear regression will be negatively affected by co and multilinearity in independant features. The following section will address this issue.
# +
corr=linear_data.drop(columns = targets).corr().abs()
corr = corr.unstack().sort_values().reset_index()
corr = corr[corr['level_0']!=corr['level_1']]#removes columns correlated with themselves from analysis
#takes the correlated data and removes duplicates. otherwise data frame will have feature 1 - feature 2 and feature 2 - feature 1
corr['ordered-cols'] = corr.apply(lambda x: '-'.join(sorted([x['level_0'],x['level_1']])),axis=1)
corr = corr.drop_duplicates(['ordered-cols'])
corr.drop(['ordered-cols'], axis=1, inplace=True)
# -
print(corr.shape)
print(corr.tail(5))
#finds correlation price_person has with each feature for comparison
price_0 = []
price_1 = []#lists for storing results
for i, r in corr.iterrows():
column0 = corr.level_0[i]
column1 = corr.level_1[i]
price_0.append(abs(linear_data['price_person'].corr(linear_data[column0])))
price_1.append(abs(linear_data['price_person'].corr(linear_data[column1])))
#makes a new columns to compare price correlation with feature 0 with price correlation with feature 1.
corr = corr.reset_index()
corr['price_0'] = pd.Series(price_0)
corr['price_1'] = pd.Series(price_1)
# +
corr[corr[0]>.7]#filters rows with high colinearity
remove = []#starts a list for storing values
for i, row in corr[corr[0]>.7].iterrows():#loop that appends the feature with the smaller correlation to price
if corr.price_0[i]<corr.price_1[i]:
remove.append(corr.level_0[i])
else:
remove.append(corr.level_1[i])
remove = set(remove)#makes list unique
print(remove)
print(len(remove))
# +
lin_reg_data = linear_data.drop(columns = remove)#makes a new data frame that removes colinear features.
price_corr = {}#dictionary for storing
for column in lin_reg_data.drop(columns = targets).columns:#iterates over columns other than targets
price_corr[column] = abs(lin_reg_data['price_person'].corr(lin_reg_data[column]))#finds correlation between column and price.
price_corr = pd.DataFrame(price_corr.items())#stores dict to dataframe for easier filtering
# -
price_corr = price_corr.sort_values(1, ascending = False)#sort values by descending correlation to price
highest_20 = list(price_corr[0][:20])#list to keep 20 most correlated values
lin_reg_data = lin_reg_data[highest_20]
# ## Conclusion
# There should now be appropriate features for decision tree regressions, standard linear regression and lasso regressions.
airbnb.price_person
linear_data[remove].to_csv(r'../data/modeling/co_linear.csv')
linear_data.to_csv(r'../data/modeling/airbnb.csv')
airbnb[targets].to_csv(r'../data/modeling/targets.csv')
lin_reg_data.to_csv(r'../data/modeling/linear_data.csv')
linear_data.head()
airbnb.head()
airbnb.price.hist()
np.log(airbnb.price).hist()
| notebooks/Preprocessing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ### 数据操作
import torch
torch.manual_seed(0)
torch.__version__
# ### 2.2.1 创建Tensor
#
# > **创建一个5x3的未初始化的Tensor**
#
# - torch.empty(5, 3)
# - torch.rand(5, 3)
# - torch.zeros(5, 3)
# - torch.tensor([5, 3])
# - x.new_ones(5, 3, dtype=torch.float) # 在现有的tensor上穿件新的tensor
# - torch.rand_like(x, dtype=torch.float) # 根据x维度来创建随机tensor
#
x = torch.empty(5, 3)
x
x = torch.rand(5, 3)
x
x = torch.zeros(5, 3, dtype=torch.long)
x
x = torch.tensor([5.5, 3])
x
# 通过现有的tensor来创建
x = x.new_ones(5, 3, dtype=torch.float64)
x
x = torch.rand_like(x, dtype=torch.float)
x
# 返回结果是tuple
print(x.shape)
print(x.size())
# ### 2.2.2 操作
#
# #### 算术操作
#
# > 1. **加法形式一: x + y**
# > 2. **加法形式二: torch.add(x, y)**
# > 3. **加法形式三: x.add_(y), 此时为 x = x + y**
y = torch.rand(5, 3)
x + y
torch.add(x, y)
x.add_(y)
# ### 索引操作
#
# > **torch的索引操作类似于numpy, 索引出来的结果与原数据共享内存**
z = x[0, :]
z
z.add_(1)
z
x[0, :]
# ### 改变形状
#
# > 1. **通过tensor.view()改变形状**
# > 2. **tensor.view()返回的新tensor与原tensor共享内存**
# > 3. **通过tensor.clone()创造一个副本来进行修改**
# > 4. **通过tensor.item()将一个标量tensor转化为python number**
y = x.view(15)
y.shape
a = x.view(5, -1)
a.shape
x_cp = x.clone().view(15)
x_cp
x -= 1
x_cp
x
# ### 2.2.3 广播机制
#
# > 1. **每个张量具有至少一个维度**
# > 2. **在遍历维度大小时, 从尾部维度开始遍历, 两者维度必须"相同", 此处"相同"三种含义: 两者相等;其中一个是1;其中一个不存在**
x = torch.arange(1, 3).view(1, 2)
print(x)
y = torch.arange(1, 4).view(3, 1)
z = x+y
print(y)
print(z)
z - y
# ### 2.2.4 运算的内存开销
x = torch.tensor([1, 2])
y = torch.tensor([3, 4])
id_before = id(y)
y = y + x
id(y) == id_before
x = torch.tensor([1, 2])
y = torch.tensor([3, 4])
id_before = id(y)
y[:] = y + x
id(y) == id_before
x = torch.tensor([1, 2])
y = torch.tensor([3, 4])
id_before = id(y)
torch.add(x, y, out=y) # 等价于 y += x; y.add_(x)
id(y) == id_before
# ### 2.2.5 Tensor和Numpy相互转换
#
# > 1. **tenfor -> numpy : tensor.numpy()**
# > 1. **numpy -> tenfor : torch.from_numpy()**
# > 1. **numpy()和from_numpy()产生的tensor和 numpy array使用的相同的内存**
# > 4. **torch.tensor()会进行数据拷贝,返回的tensor不会和原来的数据共享存**
a = torch.ones(5)
b = a.numpy()
print(a)
print(b)
a += 1
print(a)
print(b)
b += 1
print(a)
print(b)
import numpy as np
a = np.ones(5)
b = torch.from_numpy(a)
print(a, b)
a += 1
print(a, b)
b += 1
print(a, b)
c = torch.tensor(a)
c += 1
print(a, c)
# ### 2.2.6 Tensor On GPU
if torch.cuda.is_available():
device = torch.device('cuda') # GPU
y = torch.ones_like(x, device=device) # 创建一个在GPU上的tensor
x = x.to(device) # 等价于 .to("cuda")
z = x + y
z.to("cpu", torch.double)
| dl/dive-into-dl/chapter02-prerequisite/2.1_tensor.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] pycharm={"name": "#%% md\n"}
#
# # Motor Imagery Left/Right Classification Using Machine Learning
# ## BCI-4-ALS, Ben Gurion University, 2020
# ## <NAME>, <NAME>
# + pycharm={"name": "#%%\n"}
import numpy as np
import pyxdf
import matplotlib.pyplot as plt
f = r"C:\Users\noam\Documents\CurrentStudy\sub-P001\ses-S001\eeg\sub-P001_ses-S001_task-Default_run-001_eeg_old3.xdf"
f
# + pycharm={"name": "#%%\n"}
data, header = pyxdf.load_xdf(f)
# + pycharm={"name": "#%%\n"}
data
# + pycharm={"name": "#%%\n"}
header
# + pycharm={"name": "#%%\n"}
| examples/.ipynb_checkpoints/explore_recording-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3-azureml
# kernelspec:
# display_name: Python 3.6 - AzureML
# language: python
# name: python3-azureml
# ---
# %load_ext autoreload
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
# %autoreload 2
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}} gather={"logged": 1616755186841}
import sys
sys.path.append('/home/azureuser/cloudfiles/code/Users/src/')
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}} gather={"logged": 1616755316320}
import ee
# Trigger the authentication flow.
ee.Authenticate()
# Initialize the library.
ee.Initialize()
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}} gather={"logged": 1616755196759}
"""
Detect Methane hotspots
------------------------------
Functions to load and detect methane hotspots
"""
import pandas as pd
import numpy as np
import geopandas as gpd
import shapely
import ee
# Load infra
from infrastructure import plants_as_gdf, pipelines_as_gdf
df_plants = plants_as_gdf()
df_pipelines = pipelines_as_gdf()
df_pipelines = df_pipelines[df_pipelines.geometry.notnull()]
def hotspots_as_gdf(hotspots_gpd, start_date, end_date):
"""
Merge hotspots with infrastructure data and return list of most critical
methane events linked to fossil fuel production site
:param hotspots_gpd: a geopandas dataframe of detected methane events
:return: a geopandas dataframe with most critical events linked to fossil fule infrastructure
"""
return (hotspots_gpd
.assign(min_dist_plant = lambda _df: _df.apply(lambda _e: df_plants.geometry.distance(_e.geometry).min(), axis=1))
.assign(min_dist_pipeline = lambda _df: _df.apply(lambda _e: df_pipelines.geometry.distance(_e.geometry).min(), axis=1))
.assign(min_dist_infra = lambda _df: _df[["min_distance_to_plant", "min_distance_to_pipeline"]].min(axis=1))
.assign(area_m2 = lambda _df: _df.geometry.area)
# 0 criticality if more than 20km, linear in-between
.assign(infra_dist_score = lambda _df: (20 - _df.min_distance_to_infra.clip(0, 20) / 20) )
# log of size
.assign(criticality = lambda _df: np.log(_df.area_m2+1) / np.log(1.01) / _df.infrastructure_distance_score)
.sort_values(by="criticality", ascending=False)
.assign(start_date=start_date)
.assign(end_date=end_date)
)
# Taken from: https://github.com/rutgerhofste/eeconvert/blob/master/eeconvert/__init__.py
def fcToGdf(fc, crs = {'init' :'epsg:4326'}):
"""converts a featurecollection to a geoPandas GeoDataFrame. Use this function only if all features have a geometry.
caveats:
Currently only supports non-geodesic (planar) geometries because of limitations in geoPandas and Leaflet. Geodesic geometries are simply interpreted as planar geometries.
FeatureCollections larger than memory are currently not supported. Consider splitting data and merging (server side) afterwards.
:param fc (ee.FeatureCollection) : the earth engine feature collection to convert.
:param crs (dictionary, optional) : the coordinate reference system in geopandas format. Defaults to {'init' :'epsg:4326'}
:return: gdf (geoPandas.GeoDataFrame or pandas.DataFrame) : the corresponding (geo)dataframe.
"""
crs = {'init' :'epsg:4326'}
features = fc.getInfo()['features']
dictarr = []
print("Got features")
for f in features:
#geodesic = ee.Feature(f).geometry().edgesAreGeodesics()
#if geodesic:
attr = f['properties']
attr['geometry'] = f['geometry']
attr['geometry']
dictarr.append(attr)
gdf = gpd.GeoDataFrame(dictarr)
gdf['geometry'] = list(map(lambda s: shapely.geometry.shape(s), gdf.geometry))
gdf.crs = crs
return gdf
def methane_hotspots(start_date, end_date):
"""
Return detected methane leaks over period of interest
:param start_date: Initial date of interest (str: 'YYYY-MM-dd')
:param start_date: inal date of interest (str: 'YYYY-MM-dd')
:return: ee.FeatureCollection with leaks detected
"""
methane_volume = 'CH4_column_volume_mixing_ratio_dry_air'
imageCollection = ee.ImageCollection('COPERNICUS/S5P/OFFL/L3_CH4')
#Import a Landsat 8 image, subset the thermal band, and clip to the
# area of interest.
image = (
imageCollection
.select([methane_volume])
.filterDate(start_date, end_date)
.mean()
.rename('ch4')
)
uniform_kernel = ee.Kernel.square(20, 'pixels')
image_smooth = image.reduceNeighborhood(ee.Reducer.median(),uniform_kernel)
diff_w_smooth = image.subtract(image_smooth)
# Threshold the thermal band to set hot pixels as value 1, mask all else.
hotspots_diff = (
diff_w_smooth
.gt(70)
.selfMask()
.rename('hotspots_diff')
)
# Start forming components from filtered areas
objects = hotspots_diff.connectedComponents(connectedness=ee.Kernel.plus(10),maxSize=128)
# Filter on size of objects
objectSize = (
objects
.select('labels')
.connectedPixelCount(128, False)
)
# Get a pixel area image.
pixelArea = ee.Image.pixelArea()
# Multiply pixel area by the number of pixels in an object to calculate
# the object area. The result is an image where each pixel
# of an object relates the area of the object in m^2.
objectArea = objectSize.multiply(pixelArea)
areaMask = objectArea.gte(10)
# Update the mask of the `objects` layer defined previously using the
# minimum area mask just defined.
objects = objects.updateMask(areaMask)
# Whole world extracted
aoi = ee.Geometry.Polygon(
[[[-179.0, 78.0], [-179.0, -58.0], [179.0, -58.0], [179.0, 78.0]]], None,
False)
toVectors = objects.reduceToVectors(
geometry=aoi,
crs=image.projection(),
scale=2000,
geometryType='polygon',
eightConnected=False,
labelProperty='hotspots',
reducer=ee.Reducer.mean(),
maxPixels=10e8,
)
return toVectors
def save_methane_hotspots(start_date, end_date):
"""
Return detected methane leaks over period of interest as geopandas dataframe and save file
:param start_date: Initial date of interest (str: 'YYYY-MM-dd')
:param start_date: inal date of interest (str: 'YYYY-MM-dd')
:return:
"""
methane_hotspots_vectors = methane_hotspots(start_date, end_date)
gpd = fcToGdf(methane_hotspots_vectors)
gpd.to_file(f'methane_hotspots_start_date={start_date}_end_date={end_date}.geojson', driver='GeoJSON')
def run(start_date, end_date, fdir='/mounted/'):
"""
Return detected methane leaks over period of interest as geopandas dataframe and save file
:param start_date: Initial date of interest (str: 'YYYY-MM-dd')
:param start_date: inal date of interest (str: 'YYYY-MM-dd')
:return:
"""
# From GEE
methane_hotspots_vectors = methane_hotspots(start_date, end_date)
# Transform to geopandas
hotspots_gpd = fcToGdf(methane_hotspots_vectors)
# Link with infra
hotspot_w_infra = hotspots_as_gdf(hotspots_gpd, start_date, end_date)
# write to disk
gpd.to_file(f'{fdir}/methane_hotspots_start_date={start_date}_end_date={end_date}.geojson', driver='GeoJSON')
return "Success"
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
run("2021-03-14", "2021-03-21", fdir='/mounted/')
run("2021-03-07", "2021-03-14", fdir='/mounted/')
run("2021-02-28", "2021-03-07", fdir='/mounted/')
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
run("2021-02-21", "2021-02-28", fdir='/mounted/')
run("2021-02-14", "2021-02-21", fdir='/mounted/')
run("2021-02-07", "2021-02-21", fdir='/mounted/')
run("2021-01-31", "2021-02-07", fdir='/mounted/')
# + [markdown] nteract={"transient": {"deleting": false}}
# # Time serie
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}} gather={"logged": 1616752773026}
import pandas as pd
def ee_array_to_df(arr, list_of_bands):
"""Transforms client-side ee.Image.getRegion array to pandas.DataFrame."""
df = pd.DataFrame(arr)
# Rearrange the header.
headers = df.iloc[0]
df = pd.DataFrame(df.values[1:], columns=headers)
# Remove rows without data inside.
df = df[['longitude', 'latitude', 'time', *list_of_bands]].dropna()
# Convert the data to numeric values.
for band in list_of_bands:
df[band] = pd.to_numeric(df[band], errors='coerce')
# Convert the time field into a datetime.
df['datetime'] = pd.to_datetime(df['time'], unit='ms')
# Keep the columns of interest.
df = df[['time','datetime', *list_of_bands]]
return df
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}} gather={"logged": 1616753119794}
import geopandas as gpd
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}} gather={"logged": 1616753139573}
fpath_hotspots = 'methane_hotspots_start_date=2021-03-01_end_date=2021-03-04.geojson'
hotspots = gpd.read_file(fpath_hotspots)
# + [markdown] nteract={"transient": {"deleting": false}}
# # Timeseries
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}} gather={"logged": 1616754429146}
start_date="2021-02-01"
end_date = "2021-04-01"
roi = ee.Geometry.Polygon(list(hotspots.iloc[1].geometry.exterior.coords))
im_coll = (ee.ImageCollection('COPERNICUS/S5P/OFFL/L3_CH4')
.filterBounds(roi)
.filterDate(ee.Date(start_date),ee.Date(end_date))
.map(lambda img: img.set('date', ee.Date(img.date()).format('YYYYMMdd')))
.sort('date'))
region_im = im_coll.getRegion(roi, 2000).getInfo()
timestamplist = (im_coll.aggregate_array('date')
.map(lambda d: ee.String('T').cat(ee.String(d)))
.getInfo())
#timestamplist
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}} gather={"logged": 1616754429400}
df_mth = ee_array_to_df(region_im, ['CH4_column_volume_mixing_ratio_dry_air'])
ax = df_mth.plot(x="datetime", y="CH4_column_volume_mixing_ratio_dry_air")
ax.set_ylim(1700, 2000)
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
| notebooks/extract_hotspots.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Model Selection
# +
goog_drive=False
if goog_drive:
from google.colab import drive
drive.mount('/content/drive')
goog_dir = '/content/drive/My Drive/lending_club_project/'
else:
goog_dir = ''
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import os
import pickle
from utils import chunk_loader
from sklearn.metrics import classification_report, roc_curve, precision_recall_curve, roc_auc_score
from sklearn.naive_bayes import MultinomialNB
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import GradientBoostingClassifier, RandomForestClassifier
from sklearn.neighbors import KNeighborsClassifier
from sklearn.svm import SVC
import xgboost as xgb
# %matplotlib inline
# -
#directory to save files
save_path = os.path.join(goog_dir, 'models')
# # 1.0 Download the Data
# +
#get directory
df_train_path = os.path.join(goog_dir, 'data','df_train_scaled.csv')
df_test_path = os.path.join(goog_dir,'data','df_test_scaled.csv')
#download in chunks
df_train = pd.read_csv(df_train_path, index_col=0, compression='zip')
df_test = pd.read_csv(df_test_path, index_col=0, compression='zip')
# -
df_train.shape
df_test.shape
# +
target_col = 'loan_status'
#feature space
X_train = df_train.drop(columns=[target_col])
X_test = df_test.drop(columns=[target_col])
#target variable
y_train = df_train[target_col]
y_test = df_test[target_col]
# -
# # 2.0 Defining Our Metrics
# The dataset is heaviliy imbalanced in favor of the positive class (pays back loan). We will take into account:
# - Precision
# - Recall
# - F1 Score
# - AUC
# +
#ROC dict for TPR and FPR values
roc_dict = {}
#precision recall curve
prc_dict = {}
# +
def modify_roc_dict(clf, y_test, X_test, name ,dict_modify):
"""
modifies a dictionary in place and adds keys for ROC metrics
"""
fpr, tpr, thresholds = roc_curve(y_test,
clf.predict_proba(X_test)[:,1],
pos_label=1)
clf_roc_dict = {'fpr':fpr, 'tpr': tpr, 'thresholds': thresholds}
dict_modify[name] = clf_roc_dict
# -
def modify_prc_dict(clf, y_test, X_test, name ,dict_modify):
"""
modifies a dictionary in place and adds keys for precision recall metrics
"""
precision, recall, thresholds = precision_recall_curve(y_test,
clf.predict_proba(X_test)[:,1],
pos_label=1)
clf_prc_dict = {'precision':precision, 'recall': recall, 'thresholds': thresholds}
dict_modify[name] = clf_prc_dict
# # 3.0 Fit & Score
def fit_score_clf(clf, X_train, y_train, X_test, y_test, name='clf'):
"""
fits a classifer from sklearn and returns a dataframe
clf(sklearn): classifier
X_train(numpy): train features
y_train(numpy): predictions row vector
X_test(numpy): test features
y_test(numpy): ground truth row vector
name(string): name of classifier
"""
#fit to train data
clf.fit(X_train, y_train)
#make predictions
y_pred = clf.predict(X_test)
#make dataframe from report
clf_report = pd.DataFrame(classification_report(y_test, y_pred, output_dict=True))
#add clf name as column
clf_report['clf'] = name
return clf_report
# ## 3.1 Naive Bayes
# Worth mentioning but not used since in our scaling of the data produces negative values. Naive Bayes does not accept negative values, for reference see article below:
#
# https://stats.stackexchange.com/questions/169400/naive-bayes-questions-continus-data-negative-data-and-multinomialnb-in-scikit
#
# ## 3.2 Logistic Regression
# +
#instantiate with default params
lr = LogisticRegression(penalty='l2',
C=1.0,
solver = 'lbfgs',
max_iter=400,
n_jobs=-1)
lr_report = fit_score_clf(lr, X_train, y_train, X_test, y_test, name='lr')
lr_report.head()
# -
#metrics for lr
modify_roc_dict(clf=lr,
y_test=y_test,
X_test=X_test,
name='Logistic Regression',
dict_modify=roc_dict)
modify_prc_dict(clf=lr,
y_test=y_test,
X_test=X_test,
name='Logistic Regression',
dict_modify=prc_dict)
# ## 3.3 Gradient Boosted Tree
# +
gbtree = GradientBoostingClassifier(learning_rate=0.1,
n_estimators = 100)
gbtree_report = fit_score_clf(gbtree, X_train, y_train, X_test, y_test, name='gbtree')
gbtree_report.head()
# -
#metrics for gbtree
modify_roc_dict(gbtree, y_test, X_test, name='GBTree',dict_modify=roc_dict)
modify_prc_dict(clf=gbtree,
y_test=y_test,
X_test=X_test,
name='GBTree',
dict_modify=prc_dict)
# ## 3.4 KNN
# +
knn = KNeighborsClassifier(n_neighbors =5, n_jobs=-1)
knn_report = fit_score_clf(knn, X_train, y_train, X_test, y_test, name='KNN')
knn_report.head()
# -
#metrics for knn
modify_roc_dict(knn, y_test, X_test, name='KNN',dict_modify=roc_dict)
modify_prc_dict(clf=knn,
y_test=y_test,
X_test=X_test,
name='knn',
dict_modify=prc_dict)
# ## 3.5 SVM
# +
#put hard limit on iterations to save time in exploration phase
svc = SVC(C=1.0, kernel='rbf',
gamma='auto',
probability=True,
max_iter=400)
svc_report = fit_score_clf(svc, X_train, y_train, X_test, y_test, name='svc')
svc_report.head()
# -
#metrics for svc
modify_roc_dict(svc, y_test, X_test, name='SVC',dict_modify=roc_dict)
modify_prc_dict(clf=svc,
y_test=y_test,
X_test=X_test,
name='SVC',
dict_modify=prc_dict)
# ## 3.6 Random Forest
# +
rf = RandomForestClassifier(n_estimators=100, n_jobs=-1)
rf_report = fit_score_clf(rf, X_train, y_train, X_test, y_test, name='rf')
rf_report.head()
# -
#metrics for random forest
modify_roc_dict(rf, y_test, X_test, name='RandomForest',dict_modify=roc_dict)
modify_prc_dict(clf=rf,
y_test=y_test,
X_test=X_test,
name='RandomForest',
dict_modify=prc_dict)
# ## 3.7 XGBoost
# +
xgb_clf = xgb.XGBClassifier(n_estimators=300,
subsample=0.9,
n_jobs=-1)
xgb_report = fit_score_clf(xgb_clf, X_train, y_train, X_test, y_test, name='xgb')
xgb_report.head()
# -
#metrics for svc
modify_roc_dict(xgb_clf, y_test, X_test, name='XGBoost',dict_modify=roc_dict)
modify_prc_dict(clf=xgb_clf,
y_test=y_test,
X_test=X_test,
name='XGBoost',
dict_modify=prc_dict)
# # 4.0 Comparing Metrics
# After training some classifiers with default paramters, we can get a first impression of performance. With the ROC metric, the best model is closest to the "North West" corver of the axes, where as for the PRC the best location is "North East".
#
# We will show the y=x line in the ROC plot in order to provide a baseline. Any classifer at this line makes completely random predictions, a classifer below it makes worse predictions than a random guess. Should this be the case, it is possible that our features were not approriately set up for that classifier or that the initial default parmaeters were very poorly choosen.
#see keys we have
roc_dict.keys()
for key in roc_dict:
clf = roc_dict[key]
plt.plot(clf['fpr'], clf['tpr'], label=key)
plt.xlabel("FPR")
plt.ylabel("TPR")
plt.title("ROC")
plt.plot([0,1], [0,1], label='No Discrimination', linestyle='-', dashes=(5, 5))
plt.legend()
plt.savefig(os.path.join(goog_dir, 'plots/roc_curve_all_models.png'))
plt.show()
# +
for key in prc_dict:
clf = prc_dict[key]
plt.plot(clf['precision'], clf['recall'], label=key)
plt.xlabel("Precision")
plt.ylabel("Recall")
plt.title("Precision Recall Curve")
plt.legend()
plt.savefig(os.path.join(goog_dir, 'plots/prc_curve_all_models.png'))
plt.show()
# -
# # 5.0 Picking a Model
# Looking at the overall performance, XGBoost seems to be the most viable option. In the next notebook we will iterate on the parameters to finetune performance.
# +
#define location to save trained model
save_model_dir = os.path.join(save_path, 'xgb_default_params.pickle')
print("Saving model at: {}".format(save_model_dir))
#save the model
with open(save_model_dir, 'wb') as handle:
pickle.dump(xgb_clf,
handle,
protocol=pickle.HIGHEST_PROTOCOL)
# -
| notebooks/03_Model_Selection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.10 ('s2search397')
# language: python
# name: python3
# ---
# ### Init
# +
sys.path.insert(1, '../../')
import numpy as np, sys, os, pandas as pd,warnings
from getting_data import read_conf
import matplotlib.pyplot as plt
from s2search_score_pdp import pdp_based_importance, apply_order
warnings.filterwarnings("ignore")
old_list_idx_map = dict(
title=0,
abstract=1,
venue=2,
authors=3,
year=4,
n_citations=5,
)
old_list = [
'title',
'abstract',
'venue',
'authors',
'year',
'n_citations'
]
def get_Z_by_flist(f_list):
print()
print(f_list)
print()
Z = np.zeros([len(f_list), len(f_list)])
# load interaction from 2 way ale
for i in range(len(f_list)):
f1_name = f_list[i]
for j in range(i + 1, len(f_list)):
f2_name = f_list[j]
file = os.path.join('.', 'scores', f'{sample_name}_hs_metrix.npz')
# hs_metrix = np.load(file)['hs_sqrt_metrix']
hs_metrix = np.load(file)['hs_metrix']
# hs_metrix = hs_metrix / np.linalg.norm(hs_metrix)
Z[i][j] = hs_metrix[old_list_idx_map[f1_name]][old_list_idx_map[f2_name]]
Z[j][i] = hs_metrix[old_list_idx_map[f1_name]][old_list_idx_map[f2_name]]
return Z
# -
# ### Loading data
# +
pd.set_option('display.expand_frame_repr', False)
sample_name = 'cslg-rand-1000'
Z = get_Z_by_flist(old_list)
for a in Z:
print(a)
def get_wi_order(z):
order = []
col = 0
for row in z:
vi = row[col]
other_sij = [*row[0:col], *row[col+1:]]
sij = np.max(other_sij)
order.append({
'feature_name': old_list[col],
'wi': vi + sij
})
col += 1
order.sort(key=lambda x: x['wi'], reverse=True)
return [x['feature_name'] for x in order]
sorted_f_list = get_wi_order(Z / np.linalg.norm(Z))
Z = get_Z_by_flist(sorted_f_list)
for a in Z:
print(a)
sorted_f_list = [x.title() for x in sorted_f_list]
# -
# ### Heatmap Plot
# +
mask= np.array([
[1,0,0,0,0,0],
[0,1,0,0,0,0],
[0,0,1,0,0,0],
[0,0,0,1,0,0],
[0,0,0,0,1,0],
[0,0,0,0,0,1]
]).astype(bool)
z1 = np.copy(Z)
z1[mask] = np.nan
z2 = np.copy(Z)
z2[~mask] = np.nan
z1 = np.flipud(z1)
z2 = np.flipud(z2)
fig, ax = plt.subplots(figsize=(14, 8), dpi=200, constrained_layout=True)
ax.xaxis.tick_top()
ax.set_xticklabels(sorted_f_list, fontsize=16)
ax.set_yticklabels(np.flip(sorted_f_list), fontsize=14)
map1 = ax.pcolormesh(sorted_f_list, np.flip(sorted_f_list), z1, cmap='Purples', alpha=0.9)
# map2 = ax.pcolormesh(sorted_f_list, np.flip(sorted_f_list), z2, cmap='Greens', alpha=0.9)
map1cb = fig.colorbar(map1, ax=ax, shrink=0.6, pad=0.03)
map1cb.ax.set_title('FInt', size=16)
# map2cb = fig.colorbar(map2, ax=ax, shrink=0.4, pad=0.03, anchor=(-0.9, 0.1))
# map2cb.ax.set_title('FImp', size=16)
for (i, j), z in np.ndenumerate(np.flipud(Z)):
ax.text(j, i, '{:0.4f}'.format(z), ha='center', va='center', bbox=dict(boxstyle='round', facecolor='white', edgecolor='0.8'), size=14)
plt.savefig(os.path.join('.', 'plot', f'{sample_name}-heatmap.png'), facecolor='white', transparent=False, bbox_inches='tight')
# -
# ### Network Plot
# +
import matplotlib as mpl
import matplotlib.pyplot as plt
import networkx as nx
import math
def plot_net_work(f_list, z1, z2, sl=0, ignore_edges=[], plot_last_name=''):
f_list = f_list[sl:]
z1 = np.flipud(z1)
z2 = np.flipud(z2)
z1 = z1[sl: , sl:]
z2 = z2[sl: , sl:]
plt.figure(figsize=(14,8), dpi=200, facecolor='white', tight_layout=True)
G = nx.Graph()
pos_cor = [
(2,0),
(1,math.sqrt(3)),
(-1,math.sqrt(3)),
(-2,0),
(-1,-math.sqrt(3)),
(1,-math.sqrt(3))
][sl:]
pos_cor_2 = [
(2,-0.2),
(1,math.sqrt(3) + 0.2),
(-1,math.sqrt(3) + 0.2),
(-2.1,0.2),
(-1,-math.sqrt(3) - 0.15),
(1,-math.sqrt(3) - 0.15)
][sl:]
pos = {}
pos_2 = {}
for i in range(len(f_list)):
feature_name = f_list[i]
G.add_nodes_from([(i, {"feature_name": feature_name, 'imp': z2[i][i]})])
pos[i] = np.array(pos_cor[i])
pos_2[i] = np.array(pos_cor_2[i])
edge_widths = []
edge_colors = []
for i in range(len(f_list)):
f1_name = f_list[i]
for j in range(i + 1, len(f_list)):
f2_name = f_list[j]
if f'{f1_name}-{f2_name}'.lower() not in ignore_edges and f'{f2_name}-{f1_name}'.lower() not in ignore_edges:
G.add_edges_from([(i, j, {'int': (z1)[i][j]})])
edge_widths.append((z1)[i][j] + (1.2 if (z1)[i][j] < 1 else 0))
edge_colors.append((z1)[i][j] + (0.8 if (z1)[i][j] < 1 and (len(ignore_edges) == 0 and sl == 0) else 0))
node_sizes = [100 for x in range(len(f_list))]
node_colors = [nx.get_node_attributes(G, 'imp').get(x) for x in range(len(f_list))]
nodes = nx.draw_networkx_nodes(G, pos, node_size=node_sizes, node_color=node_colors, cmap='Greens', edgecolors='#ddd')
edges = nx.draw_networkx_edges(
G,
pos,
node_size=node_sizes,
arrowstyle="-",
edge_color=edge_colors,
edge_cmap=plt.cm.get_cmap('Purples'),
width=edge_widths,
)
map2cb = plt.colorbar(edges, shrink=0.6)
map2cb.ax.set_title('FInt', size=14)
nodes = nx.draw_networkx_labels(G, pos_2, labels=nx.get_node_attributes(G, 'feature_name'))
ax = plt.gca()
ax.set_axis_off()
plt.savefig(os.path.join('.', 'plot', f'{sample_name}-network_graph_{plot_last_name}.png'), facecolor='white', transparent=False, bbox_inches='tight')
plot_net_work(sorted_f_list, z1, z2)
# -
plot_net_work(sorted_f_list, z1, z2, sl=1, plot_last_name='remove_top_node')
| pipelining/pdp-hs-exp1/visualizing_cslg-rand-1000.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Workbook to generate an example of a difficult to analyze spreadsheet (csv file)
# Starting with a well formated and normalized workbook
# and introducing errors into it
# -
import pandas as pd
import math
df_clean = pd.read_csv('data/10-20_carbonhealth_and_braidhealth.csv')
pd.set_option('display.max_columns', None)
df_clean.head(10)
len(df_clean)
from pandasql import sqldf
pysqldf = lambda q: sqldf(q, globals())
pysqldf("SELECT * FROM df_clean").head(5)
df_clean.columns
#pysqldf("SELECT high_risk_interactions FROM df_clean WHERE high_risk_interactions IS NULL")
len(df_clean)
df_clean.columns
# +
symptoms = ['labored_respiration', 'rhonchi', 'wheezes', 'cough', 'cough_severity',
'fever', 'sob', 'sob_severity', 'diarrhea', 'fatigue', 'headache',
'loss_of_smell', 'loss_of_taste', 'runny_nose', 'muscle_sore', 'sore_throat']
vitals = ['pulse', 'sys', 'dia']
numerical = ['temperature', 'rr', 'sats',
'ctab', 'days_since_symptom_onset']
comorbidities = ['diabetes', 'chd', 'htn', 'cancer', 'asthma', 'copd', 'autoimmune_dis', 'smoker']
cxrs = ['cxr_findings', 'cxr_impression', 'cxr_label', 'cxr_link']
risks = ['high_risk_exposure_occupation', 'high_risk_interactions']
tests = ['test_name', 'swab_type']
test_types = ['covid19_test_results', 'rapid_flu_results', 'rapid_strep_results']
cs = []
cc = []
cr = []
ct = []
cv = []
cpt = []
cnt = []
for index, row in df_clean.iterrows():
rs = []
rc = []
rr = []
rt = []
rv = []
rpt = []
rnt = []
#print(row)
for s in symptoms:
if row[s] == 1:
rs.append(s)
for c in comorbidities:
if row[c] == 1:
rc.append(c)
for r in risks:
if row[r] == 1:
rr.append(r)
for r in tests:
rt.append(row[r])
for r in vitals:
if not math.isnan(float(row[r])):
rv.append(str(row[r]))
for r in test_types:
if row[r] == 'Positive':
rpt.append(r[:-8])
if row[r] == 'Negative':
rnt.append(r[:-8])
cs.append(', '.join(rs))
cc.append(', '.join(rc))
cr.append(', '.join(rr))
ct.append(', '.join(rt))
cv.append(', '.join(rv))
cpt.append(', '.join(rpt))
cnt.append(', '.join(rnt))
# -
print(len(df_clean['batch_date']))
print(len(ct))
df_ugh = pd.DataFrame({'Date': df_clean['batch_date'], 'Type': ct, 'Co-Morbidities': cc,
'Symptom(s)': cs, 'Risk(s)': cr, 'Age': df_clean['age'],
'Vitals': cv, 'Pos Tests': cpt, 'Neg Tests': cnt})
for n in numerical:
df_ugh[n] = df_clean[n]
df_ugh = df_ugh.replace(math.nan, '')
# keeping the output short for github
# you'll need to view more of these to see ssome of the the denormalized fields (symptoms, tests, vitals)
df_ugh.head(500)
# create a smaller set with most of the data fields populated
df_minimal = pysqldf("""
SELECT * FROM df_ugh
WHERE `Risk(s)` <> ''
AND `Co-Morbidities` <> ''
AND `Vitals` IS NOT NULL
ORDER BY LENGTH(`Neg Tests`) DESC
""")
len(df_minimal)
pd.set_option('display.max_rows', None)
df_minimal.head(100)
df_minimal.head(100).to_csv('denormalized_minimal_dataset.csv', index=False)
df_ugh.to_csv('denormalized_dataset.csv', index=False)
# +
# other things to demonstrate
# different types of date format (text, dashes, forward slash, different arrangements, some only month and year)
# negative values, null, outliers (like -999 would throw off age or a vital)
# (highlight a column, other purely visual indicator)
# condensed version (a few hundred, so that you can see the errors without long scrolling)
# inconsistent colums (test_result_1, test_result_2, test_result_3)
# late Jan/Feb for workshop
# -
| Denormalize-Dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Installing Dependencies
# + _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19"
# !pip install tensorflow-gpu==2.0.0-beta1
# Load the TensorBoard notebook extension
# %load_ext tensorboard
# -
# ## Importing Dependencies**
# + _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a"
import os
import math
import shutil
import datetime
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import tensorflow as tf
from tensorflow import keras as krs
from tensorflow.keras.preprocessing.image import ImageDataGenerator
# -
from PIL import Image
from PIL import ImageDraw
from glob import glob
from tqdm import tqdm
from skimage.io import imread
from IPython.display import SVG
from sklearn.utils import shuffle
from sklearn.metrics import classification_report
from sklearn.model_selection import train_test_split
from sklearn.metrics import confusion_matrix, roc_curve, auc
tf.__version__
# !pip install livelossplot
from livelossplot.keras import PlotLossesCallback
# ## Data Preparation**
# +
TRAINING_LOGS_FILE = "training_logs.csv"
MODEL_SUMMARY_FILE = "model_summary.txt"
MODEL_FILE = "histopathologic_cancer_detector.h5"
TRAINING_PLOT_FILE = "training.png"
VALIDATION_PLOT_FILE = "validation.png"
#ROC_PLOT_FILE = "roc.png"
INPUT_DIR = '../input/'
SAMPLE_COUNT = 60000
TESTING_BATCH_SIZE = 5000
# +
training_dir = INPUT_DIR + 'train/'
df = pd.DataFrame({'path': glob(os.path.join(training_dir,'*.tif'))})
df['id'] = df.path.map(lambda x: x.split('/')[3].split('.')[0])
labels = pd.read_csv(INPUT_DIR + 'train_labels.csv')
df = df.merge(labels,on='id')
negative_values = df[df.label == 0].sample(SAMPLE_COUNT)
positive_values = df[df.label == 1].sample(SAMPLE_COUNT)
df = pd.concat([negative_values,positive_values]).reset_index()
df = df[['path','id','label']]
df['image'] = df['path'].map(imread)
# +
train_path = '../training'
val_path = '../validation'
for directory in [train_path,val_path]:
for sub_directory in ['0','1']:
path = os.path.join(directory,sub_directory)
os.makedirs(path,exist_ok=True)
train,val = train_test_split(df,train_size=0.8,stratify=df['label'])
df.set_index('id',inplace=True)
for images_paths in [(train,train_path),(val,val_path)]:
images = images_paths[0]
path = images_paths[1]
for image in images['id'].values:
file_name = image + '.tif'
label = str(df.loc[image,'label'])
destination = os.path.join(path,label,file_name)
if not os.path.exists(destination):
source = os.path.join(INPUT_DIR + 'train',file_name)
shutil.copyfile(source,destination)
# -
# ## Data Generators and Data Augmentation**
#
# **Note: ** Take note of the input shape of Data Generated Image.
# +
train_datagen = ImageDataGenerator(rescale=1/255,
horizontal_flip=True,
vertical_flip=True,
rotation_range=30,
zoom_range=0.2,
width_shift_range=0.2,
height_shift_range=0.2,
shear_range=0.05,
channel_shift_range=0.1)
train_generator = train_datagen.flow_from_directory(train_path,
target_size=(96,96),
batch_size=10,
class_mode='binary')
validation_datagen = ImageDataGenerator(rescale=1/255)
validation_generator = validation_datagen.flow_from_directory(val_path,
target_size=(96,96),
batch_size=10,
class_mode='binary')
# -
# ## Pre-Train Network (Transfer Learning) / Creating the Model
#
# **Note: ** Take note of the input shape of pre-trained network.
base_model = krs.applications.VGG19(include_top=False,
input_shape = (96,96,3),
weights = 'imagenet')
# + _kg_hide-output=true
# Checking the Layers of the Model
'''for layer in base_model.layers[:-15]:
layer.trainable = False'''
for layer in base_model.layers:
print(layer,layer.trainable)
# -
# ## Creating the Model
# +
def create_model(base_model, summary=False):
model = krs.Sequential(name='VGG19 + FC')
model.add(base_model)
model.add(krs.layers.Flatten())
model.add(krs.layers.Dense(1024,activation='relu'))
model.add(krs.layers.Dropout(0.5))
model.add(krs.layers.Dense(1,activation='sigmoid'))
if summary==True:
model.summary()
return model
# tf.keras.layers.Add
# tf.keras.layers.Input
# -
model = create_model(base_model, True)
# ## Checking the Entire Model**
# +
# Checking the Layers of the Model
'''for layer in base_model.layers[:-15]:
layer.trainable = False'''
for layer in base_model.layers:
print(layer,layer.trainable)
# +
# Checking the Layers of the Model
'''for layer in model.layers[:-15]:
layer.trainable = False'''
for layer in model.layers:
print(layer,layer.trainable)
# -
# ## Callbacks**
#
# **Note: ** Although functioning in code, tensorboard is not displaying on kaggle's kernel.
# Load the TensorBoard notebook extension.
# %load_ext tensorboard
# %reload_ext tensorboard
# +
from keras.callbacks import ModelCheckpoint, CSVLogger
log_dir="logs/fit" + datetime.datetime.now().strftime("%Y%m%d-%H%M%S")
checkpoint = ModelCheckpoint(
'./base.model',
monitor='val_loss',
verbose=1,
save_best_only=True,
mode='min',
save_weights_only=False,
period=1
)
csvlogger = CSVLogger(
filename= "training_csv.log",
separator = ",",
append = False
)
tensorboard_callback = tf.keras.callbacks.TensorBoard(log_dir=log_dir, histogram_freq=1)
# -
# Clear any logs from previous runs
# !rm -rf ./logs/
callbacks = [tensorboard_callback]
# ## Tuning, Compiling and Training the Model **
# +
# Hyperparameters
total_data = 9600
lr = 1e-4
n_epochs = 50
steps_epoch = total_data / 4
verbosity = 1
# Compile the Model, Loss and Optimizer
model.compile(loss='binary_crossentropy',
optimizer=krs.optimizers.Adam(lr=lr),
metrics=['accuracy'])
# -
training = model.fit_generator(train_generator,
#batch_size=steps_epoch,
steps_per_epoch=steps_epoch,
epochs=n_epochs,
validation_data=validation_generator,
validation_steps=steps_epoch,
verbose=verbosity,
callbacks=callbacks)
# ## Showing the Results**
#
# **Note: ** Although functioning in code, tensorboard is not displaying on kaggle's kernel.
#----Custom function to visualize the training of the model------#
def show_final_history(score):
fig, ax = plt.subplots(1, 2, figsize=(16,5))
ax[0].plot(score.epoch, score.history["loss"], label="Train loss")
ax[0].plot(score.epoch, score.history["val_loss"], label="Validation loss")
ax[0].ylabel('Loss')
ax[0].xlabel('# of Episode')
ax[0].grid(which="major", alpha=0.30)
ax[1].plot(score.epoch, score.history["acc"], label="Train acc")
ax[1].plot(score.epoch, score.history["val_acc"], label="Validation acc")
ax[1].ylabel('Accuracy')
ax[1].xlabel('# of Episode')
ax[1].grid(which="major", alpha=0.30)
ax[0].legend()
ax[1].legend()
show_final_history(history)
print("Validation Accuracy: " + str(history.history['val_acc'][-1:]))
# Tensorboard Not Working
# Starting Tensorboard
# %tensorboard --logdir logs/fit
# ## Saving and Loading the Results
## Saving and Loading the WEIGHTS
model.save_weights('my_model_weights.h5')
model.load_weights('my_model_weights.h5')
# +
# Saving and Loading the whole model (ARCHITECTURE + WEIGHTS + OPTIMIZER STATE)
model.save('my_model.h5') # creates a HDF5 file 'my_model.h5'
del model # deletes the existing model
# returns a compiled model
# identical to the previous one
model = load_model('my_model.h5')
# -
# ## ROC & AUC
#
# - **ROC Curve**
# - Curve of probability
# - In a ROC curve the true positive rate (Sensitivity) is plotted in function of the false positive rate (100-Specificity) for different cut-off points of a parameter. Each point on the ROC curve represents a sensitivity/specificity pair corresponding to a particular decision threshold.
#
#
# - **AUD or Area under the Curve**
# - is a measure of how well a parameter can distinguish between two diagnostic groups (diseased/normal).
# ## Acknowledgement
# - https://towardsdatascience.com/understanding-auc-roc-curve-68b2303cc9c5
# - https://www.medcalc.org/manual/roc-curves.php
| Histopathologic-Cancer/Cancer Detection TF2_initial.ipynb |
/ ---
/ jupyter:
/ jupytext:
/ text_representation:
/ extension: .q
/ format_name: light
/ format_version: '1.5'
/ jupytext_version: 1.14.4
/ kernelspec:
/ display_name: Q (kdb+)
/ language: q
/ name: qpk
/ ---
/ ## Outline
/
/ Often ingestion tasks are written to simply dump data to disk. A second job then has to perform a disk sort.
/ This is slow.
/
/ This notebook shows how to perform a distributed ingest while also sorting.
/ ## Create sample table
\l ../../qparquet.q
p)import numpy as np
p)times=[np.datetime64('2012-06-30T21:00:00.000000000-0400')] * 4
p)table=pandas.DataFrame(columns=['time','sym','price','size'])
p)table['time'] = times
p)table['sym'] = ['a','b','a','b']
p)table['price'] = [4.0,3.0,2.0,1.0]
p)table['size'] = [100,200,300,400]
p)print(table)
p)parquet.write_table(pyarrow.Table.from_pandas(table), 'example.parquet')
/ ## Sorting
/
/ The important change in this example is that we extract the columns we wish to sort on in the master process.
/
/ Using these the correct sort index for the data is create.
/
/ This is then sent to all slave processes which will use it to correctly save each column in the same sort order
/
/ ```q
/ sortTab:.qparquet.getColumnsCustom[file;2#columns;pyConversions;qConversions];
/ sortTab:update ind:i from sortTab;
/ sortTab:update `p#sym from `sym`time xasc sortTab;
/ .Q.dd[destination;`] set .Q.en[`.] delete ind from sortTab;
/
/ -25!(.z.pd;({sortInd::x};sortTab`ind));
/ ```
/ ## Running the example
/ Start your worker processes
/
/ ```bash
/ q ../../qparquet.q -p 5001 &
/ q ../../qparquet.q -p 5002 &
/ q ../../qparquet.q -p 5003 &
/ ```
/
/ Run the master process to distribute the work
/
/ ```bash
/ q convert.q -s -3 -slaves 5001 5002 5003
/ ```
/ The output shows that the qparquet data is now successfully a q splayed table with corrct sort and attributes
/
/ ```q
/ `:splayed/price`:splayed/size
/ `:splayed/.d
/ "Took 0D00:00:00.074172000"
/ `splayed
/ time sym price size
/ --------------------------------------------
/ 2012.07.01D01:00:00.000000000 a 4 100
/ 2012.07.01D01:00:00.000000000 a 2 300
/ 2012.07.01D01:00:00.000000000 b 3 200
/ 2012.07.01D01:00:00.000000000 b 1 400
/ c | t f a
/ -----| -----
/ time | p
/ sym | s p
/ price| f
/ size | j
/ ```
/ ### convert.q
/
/ This script coordinates distributing the work of converting the parquet file across multiple processes
/
/ ```q
/ //Load needed functions
/ \l ../../qparquet.q
/
/ //Open handles to worker processes
/ .z.pd:`u#asc hopen each"J"$(.Q.opt .z.X)`slaves
/
/ file:`:example.parquet;
/
/ columns:.qparquet.getColumnNames[file];
/
/ destination:`:splayed
/
/ pyConversions:enlist[`time]!enlist "table[\"time\"]=pandas.to_numeric(table[\"time\"])";
/ qConversions:`time`sym!({`timestamp$x-`long$2000.01.01D-1970.01.01D};`$);
/
/ start:.z.p;
/
/ sortTab:.qparquet.getColumnsCustom[file;2#columns;pyConversions;qConversions];
/ sortTab:update ind:i from sortTab;
/ sortTab:update `p#sym from `sym`time xasc sortTab;
/ .Q.dd[destination;`] set .Q.en[`.] delete ind from sortTab;
/
/ -25!(.z.pd;({sortInd::x};sortTab`ind));
/ {neg[x][]} each .z.pd;
/
/ //Distribute tasks to workers
/ //Each worker reads a column at a time
/ {[f;d;p;q;c]
/ show string[.z.p]," ",string c;
/ .Q.dd[d;c] set @[;sortInd] .qparquet.getColumnsCustom[f;enlist c;p;q]c
/ }[file;destination;pyConversions;qConversions] peach columns except `time`sym
/
/ //Add a .d file to the destination to inform q of the order of columns
/ .Q.dd[destination;`.d] set columns
/
/ end:.z.p;
/
/ show "Took ",string end-start;
/
/ //Load the converted table
/ \l splayed
/
/ //Query the q table
/ show select from splayed
/
/ show meta splayed
/ ```
| examples/6. scaling with sort/qparquet - scaling with sort.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # NoneType
#
# In Python, a variable can have nothing in it. It's a `NoneType` and evaluates as `None`.
sports = None
type(sports)
type(None)
bool(None)
if sports is None:
pass
d = {
'thing': 'thing2'
}
d
d.get('thing')
a = d.get('sometihng_else', None)
if a is None:
print("Did not find something else")
type(a)
| 220_NoneType.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.5 64-bit (''base'': conda)'
# name: python3
# ---
# + [markdown] id="ico4BTYhNl0m"
# # GradientBoostingRegressor with Normalize
#
#
# -
# This Code template is for regression analysis using a simple GradientBoostingRegressor based on the Gradient Boosting Ensemble Learning Technique using Normalize for Feature Rescaling.
# ### Required Packages
# + id="WqDGnvsjNl0n"
import warnings
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as se
from sklearn.preprocessing import Normalizer
from sklearn.model_selection import train_test_split
from sklearn.ensemble import GradientBoostingRegressor
from sklearn.metrics import r2_score, mean_absolute_error, mean_squared_error
warnings.filterwarnings('ignore')
# + [markdown] id="_KGh5ftRNl0o"
# ### Initialization
#
# Filepath of CSV file
# + id="MwMmFVILNl0o"
#filepath
file_path = ""
# + [markdown] id="z8Rf1JjwNl0p"
# List of features which are required for model training .
# + id="K0hTzBsMNl0p"
#x_values
features=[]
# + [markdown] id="83jstykTNl0p"
# Target feature for prediction.
# + id="WgnTeW7LNl0q"
#y_value
target = ''
# + [markdown] id="U1mFO8sVNl0q"
# ### Data Fetching
#
# Pandas is an open-source, BSD-licensed library providing high-performance, easy-to-use data manipulation and data analysis tools.
#
# We will use panda's library to read the CSV file using its storage path.And we use the head function to display the initial row or entry.
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="C8Qr3n_5Nl0r" outputId="2ed124c5-e010-48ea-c512-5622c11a8dbb"
df=pd.read_csv(file_path)
df.head()
# + [markdown] id="QigUBczdNl0r"
# ### Feature Selections
#
# It is the process of reducing the number of input variables when developing a predictive model. Used to reduce the number of input variables to both reduce the computational cost of modelling and, in some cases, to improve the performance of the model.
#
# We will assign all the required input features to X and target/outcome to Y.
# + id="GS_Kj423Nl0s"
X = df[features]
Y = df[target]
# + [markdown] id="BVcFxYCBNl0s"
# ### Data Preprocessing
#
# Since the majority of the machine learning models in the Sklearn library doesn't handle string category data and Null value, we have to explicitly remove or replace null values. The below snippet have functions, which removes the null value if any exists. And convert the string classes data in the datasets by encoding them to integer classes.
#
# + id="DLHaBrsXNl0s"
def NullClearner(df):
if(isinstance(df, pd.Series) and (df.dtype in ["float64","int64"])):
df.fillna(df.mean(),inplace=True)
return df
elif(isinstance(df, pd.Series)):
df.fillna(df.mode()[0],inplace=True)
return df
else:return df
def EncodeX(df):
return pd.get_dummies(df)
# + [markdown] id="sBWSTsJnNl0s"
# Calling preprocessing functions on the feature and target set.
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="3w19akyQNl0t" outputId="5917a5b5-f988-44ab-eb8e-aa0185295cab"
x=X.columns.to_list()
for i in x:
X[i]=NullClearner(X[i])
X=EncodeX(X)
Y=NullClearner(Y)
X.head()
# + [markdown] id="StjCRw6BNl0t"
# #### Correlation Map
#
# In order to check the correlation between the features, we will plot a correlation matrix. It is effective in summarizing a large amount of data where the goal is to see patterns.
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="MRFjRNM6Nl0t" outputId="edc91b60-e752-4748-8f44-f3ea52bef662"
f,ax = plt.subplots(figsize=(18, 18))
matrix = np.triu(X.corr())
se.heatmap(X.corr(), annot=True, linewidths=.5, fmt= '.1f',ax=ax, mask=matrix)
plt.show()
# + [markdown] id="GXTu5ne0Nl0u"
# ### Data Splitting
#
# The train-test split is a procedure for evaluating the performance of an algorithm. The procedure involves taking a dataset and dividing it into two subsets. The first subset is utilized to fit/train the model. The second subset is used for prediction. The main motive is to estimate the performance of the model on new data.
# + id="rfpdZN1HNl0u"
X_train, X_test, y_train, y_test = train_test_split(X, Y, test_size = 0.2, random_state = 123)#performing datasplitting
# + [markdown] id="t8YH4hv8QAa9"
# ## Data Rescaling
# Normalizer normalizes samples (rows) individually to unit norm.
#
# Each sample with at least one non zero component is rescaled independently of other samples so that its norm (l1, l2 or inf) equals one.
#
# We will fit an object of Normalizer to train data then transform the same data via fit_transform(X_train) method, following which we will transform test data via transform(X_test) method.
# + id="iV0VOry9QQUp"
normalizer = Normalizer()
X_train = normalizer.fit_transform(X_train)
X_test = normalizer.transform(X_test)
# + [markdown] id="3vagL5mMNl0u"
# ### Model
#
# Gradient Boosting builds an additive model in a forward stage-wise fashion; it allows for the optimization of arbitrary differentiable loss functions. In each stage a regression tree is fit on the negative gradient of the given loss function.
#
# #### Model Tuning Parameters
#
# 1. loss : {‘ls’, ‘lad’, ‘huber’, ‘quantile’}, default=’ls’
# > Loss function to be optimized. ‘ls’ refers to least squares regression. ‘lad’ (least absolute deviation) is a highly robust loss function solely based on order information of the input variables. ‘huber’ is a combination of the two. ‘quantile’ allows quantile regression (use `alpha` to specify the quantile).
#
# 2. learning_ratefloat, default=0.1
# > Learning rate shrinks the contribution of each tree by learning_rate. There is a trade-off between learning_rate and n_estimators.
#
# 3. n_estimators : int, default=100
# > The number of trees in the forest.
#
# 4. criterion : {‘friedman_mse’, ‘mse’, ‘mae’}, default=’friedman_mse’
# > The function to measure the quality of a split. Supported criteria are ‘friedman_mse’ for the mean squared error with improvement score by Friedman, ‘mse’ for mean squared error, and ‘mae’ for the mean absolute error. The default value of ‘friedman_mse’ is generally the best as it can provide a better approximation in some cases.
#
# 5. max_depth : int, default=3
# > The maximum depth of the individual regression estimators. The maximum depth limits the number of nodes in the tree. Tune this parameter for best performance; the best value depends on the interaction of the input variables.
#
# 6. max_features : {‘auto’, ‘sqrt’, ‘log2’}, int or float, default=None
# > The number of features to consider when looking for the best split:
#
# 7. random_state : int, RandomState instance or None, default=None
# > Controls both the randomness of the bootstrapping of the samples used when building trees (if <code>bootstrap=True</code>) and the sampling of the features to consider when looking for the best split at each node (if `max_features < n_features`).
#
# 8. verbose : int, default=0
# > Controls the verbosity when fitting and predicting.
#
# 9. n_iter_no_change : int, default=None
# > <code>n_iter_no_change</code> is used to decide if early stopping will be used to terminate training when validation score is not improving. By default it is set to None to disable early stopping. If set to a number, it will set aside <code>validation_fraction</code> size of the training data as validation and terminate training when validation score is not improving in all of the previous <code>n_iter_no_change</code> numbers of iterations. The split is stratified.
#
# 10. tol : float, default=1e-4
# > Tolerance for the early stopping. When the loss is not improving by at least tol for <code>n_iter_no_change</code> iterations (if set to a number), the training stops.
# + colab={"base_uri": "https://localhost:8080/"} id="lgKlnakENl0w" outputId="7b23f30b-f1a1-42a1-f6d5-c3e06b8844a8"
# Build Model here
model = GradientBoostingRegressor(random_state = 123)
model.fit(X_train, y_train)
# + [markdown] id="UMZ3j6CuNl0w"
# #### Model Accuracy
#
# We will use the trained model to make a prediction on the test set.Then use the predicted value for measuring the accuracy of our model.
#
# > **score**: The **score** function returns the coefficient of determination <code>R<sup>2</sup></code> of the prediction.
# + colab={"base_uri": "https://localhost:8080/"} id="mygbtfiJNl0w" outputId="5f58ba18-a747-46c4-c3bd-e760b1086d80"
print("Accuracy score {:.2f} %\n".format(model.score(X_test,y_test)*100))
# + [markdown] id="nJpFRQM5Nl0x"
# > **r2_score**: The **r2_score** function computes the percentage variablility explained by our model, either the fraction or the count of correct predictions.
#
# > **mae**: The **mean abosolute error** function calculates the amount of total error(absolute average distance between the real data and the predicted data) by our model.
#
# > **mse**: The **mean squared error** function squares the error(penalizes the model for large errors) by our model.
# + colab={"base_uri": "https://localhost:8080/"} id="vhc99DYcNl0x" outputId="3519d79a-5a3b-45fe-fbc4-b21896578312"
y_pred=model.predict(X_test)
print("R2 Score: {:.2f} %".format(r2_score(y_test,y_pred)*100))
print("Mean Absolute Error {:.2f}".format(mean_absolute_error(y_test,y_pred)))
print("Mean Squared Error {:.2f}".format(mean_squared_error(y_test,y_pred)))
# + [markdown] id="GKehBDRnNl0x"
# #### Feature Importances
# The Feature importance refers to techniques that assign a score to features based on how useful they are for making the prediction.
# + colab={"base_uri": "https://localhost:8080/", "height": 405} id="yHWw4iSENl0y" outputId="30085893-bc9c-464f-a184-679b41714d80"
plt.figure(figsize=(8,6))
n_features = len(X.columns)
plt.barh(range(n_features), model.feature_importances_, align='center')
plt.yticks(np.arange(n_features), X.columns)
plt.xlabel("Feature importance")
plt.ylabel("Feature")
plt.ylim(-1, n_features)
# + [markdown] id="j5_BJeh1Nl0y"
# #### Prediction Plot
#
# First, we make use of a plot to plot the actual observations, with x_train on the x-axis and y_train on the y-axis.
# For the regression line, we will use x_train on the x-axis and then the predictions of the x_train observations on the y-axis.
# + colab={"base_uri": "https://localhost:8080/", "height": 621} id="uF7R3c4XNl0y" outputId="b30c4591-f1bd-4908-a039-f472c86d8231"
plt.figure(figsize=(14,10))
plt.plot(range(20),y_test[0:20], color = "blue")
plt.plot(range(20),model.predict(X_test[0:20]), color = "red")
plt.legend(["Actual","prediction"])
plt.title("Predicted vs True Value")
plt.xlabel("Record number")
plt.ylabel(target)
plt.show()
# + [markdown] id="QV-JG9jRNl0y"
# #### Creator: <NAME> , Github: [Profile](https://github.com/Shikiz)
| Regression/Gradient Boosting Machine/GradientBoostingRegressor_Normalize.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="vwJqYAAFse3g" colab_type="text"
# # import Libraries
# + id="wRYtO0B7skuB" colab_type="code" colab={}
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# + [markdown] id="SCRN3wA1suww" colab_type="text"
# # Import dataset
# + id="oTWfUhhAsy8Z" colab_type="code" colab={}
dataset=pd.read_csv('Social_Network_Ads.csv');
x=dataset.iloc[:,:-1].values
y=dataset.iloc[:,-1].values
# + [markdown] id="iR5Va-76tkL3" colab_type="text"
# # Split the dataset into training set and test set
# + id="26seN15FtojN" colab_type="code" colab={}
from sklearn.model_selection import train_test_split
x_train,x_test,y_train,y_test=train_test_split(x,y,test_size=0.2, random_state=0)
# + [markdown] id="bfBx5XQwuYyf" colab_type="text"
# # Apply feature scalling to the training model
# + id="u-z97tFuuhlM" colab_type="code" colab={}
from sklearn.preprocessing import StandardScaler
sc=StandardScaler()
x_train=sc.fit_transform(x_train)
x_test=sc.transform(x_test)
# + id="XiEmWA1qu9zh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} executionInfo={"status": "ok", "timestamp": 1591718683515, "user_tz": -60, "elapsed": 937, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "01062919807050385714"}} outputId="a4a2ae71-a513-44e9-f9f5-92aab82f4e34"
print(x_train)
# + id="ybmPpLXhvW_w" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} executionInfo={"status": "ok", "timestamp": 1591718683517, "user_tz": -60, "elapsed": 907, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "01062919807050385714"}} outputId="cdd974c8-3061-4608-84b4-637262952910"
print(x_test)
# + [markdown] id="oodv2nUswVl6" colab_type="text"
# # Apply Kernel SVM
# + id="eQxOYAOTwYf0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"status": "ok", "timestamp": 1591718780364, "user_tz": -60, "elapsed": 973, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "01062919807050385714"}} outputId="686d1aeb-d3e0-4d49-cbdc-d64c7f5d2636"
from sklearn.naive_bayes import GaussianNB
clf=GaussianNB()
clf.fit(x_train,y_train)
# + [markdown] id="HqilgqNWx2_0" colab_type="text"
# # Predecting the test result
# + id="wz-cYYztx6pm" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} executionInfo={"status": "ok", "timestamp": 1591718786886, "user_tz": -60, "elapsed": 979, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "01062919807050385714"}} outputId="8f026ede-c471-40ba-b206-31b8e8384398"
y_pred = clf.predict(x_test)
print(np.concatenate((y_pred.reshape(len(y_pred),1), y_test.reshape(len(y_test),1)),1))
# + [markdown] id="hfXsVBubyHoB" colab_type="text"
# # Making the confusion metrices
# + id="-f8Q-NttyQiC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 67} executionInfo={"status": "ok", "timestamp": 1591718794501, "user_tz": -60, "elapsed": 1413, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "01062919807050385714"}} outputId="842aaaf6-606c-49f3-e2f6-40dbdd55d6c2"
from sklearn.metrics import confusion_matrix, accuracy_score
cm = confusion_matrix(y_test, y_pred)
print(cm)
accuracy_score(y_test, y_pred)
# + [markdown] id="uyiVAW7Cyv8l" colab_type="text"
# # Visualizing the training set
# + id="Db6vLPPcyzL6" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 349} executionInfo={"status": "ok", "timestamp": 1591718831687, "user_tz": -60, "elapsed": 2960, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "01062919807050385714"}} outputId="27d12305-1508-4458-9202-4c9f174a7647"
from matplotlib.colors import ListedColormap
X_set, y_set = sc.inverse_transform(x_train), y_train
X1, X2 = np.meshgrid(np.arange(start = X_set[:, 0].min() - 10, stop = X_set[:, 0].max() + 10, step = 1),
np.arange(start = X_set[:, 1].min() - 1000, stop = X_set[:, 1].max() + 1000, step = 1))
plt.contourf(X1, X2, clf.predict(sc.transform(np.array([X1.ravel(), X2.ravel()]).T)).reshape(X1.shape),
alpha = 0.75, cmap = ListedColormap(('red', 'green')))
plt.xlim(X1.min(), X1.max())
plt.ylim(X2.min(), X2.max())
for i, j in enumerate(np.unique(y_set)):
plt.scatter(X_set[y_set == j, 0], X_set[y_set == j, 1], c = ListedColormap(('red', 'green'))(i), label = j)
plt.title('K-NN (Training set)')
plt.xlabel('Age')
plt.ylabel('Estimated Salary')
plt.legend()
plt.show()
# + [markdown] id="ydBK3MVzy37X" colab_type="text"
# # Visualizing the test set
# + id="quCXcRCSy8ki" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 349} executionInfo={"status": "ok", "timestamp": 1591718840740, "user_tz": -60, "elapsed": 2855, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "01062919807050385714"}} outputId="0c9c5bdd-29a8-44f2-86b3-8a7533be99df"
from matplotlib.colors import ListedColormap
X_set, y_set = sc.inverse_transform(x_test), y_test
X1, X2 = np.meshgrid(np.arange(start = X_set[:, 0].min() - 10, stop = X_set[:, 0].max() + 10, step = 1),
np.arange(start = X_set[:, 1].min() - 1000, stop = X_set[:, 1].max() + 1000, step = 1))
plt.contourf(X1, X2, clf.predict(sc.transform(np.array([X1.ravel(), X2.ravel()]).T)).reshape(X1.shape),
alpha = 0.75, cmap = ListedColormap(('red', 'green')))
plt.xlim(X1.min(), X1.max())
plt.ylim(X2.min(), X2.max())
for i, j in enumerate(np.unique(y_set)):
plt.scatter(X_set[y_set == j, 0], X_set[y_set == j, 1], c = ListedColormap(('red', 'green'))(i), label = j)
plt.title('K-NN (Test set)')
plt.xlabel('Age')
plt.ylabel('Estimated Salary')
plt.legend()
plt.show()
| Classification/Naive_Bayes_Classifier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import seaborn as sns
import pandas as pd
import matplotlib.pyplot as plt
import datetime
# #%matplotlib inline
output_folder = '/drone/src/output/'
abundances = pd.read_csv(output_folder + 'abundance.tsv', delimiter='\t')
# -
# Figure 1
datetime_str = datetime.datetime.now().strftime("%Y-%m-%d %H:%M")
generated_text = 'Figures generated at: ' + datetime_str
g = sns.distplot(abundances['eff_length'], kde=False, color="b")
g.figure.suptitle(generated_text, fontsize=18, fontweight='bold')
g.figure.savefig(output_folder + 'fig1.png', bbox_inches='tight')
# Figure 2
sns.set(style="darkgrid", color_codes=True)
g = sns.jointplot("length", "est_counts", data=abundances, kind="reg",
xlim=(800, 2400), ylim=(0, 1000), color="r", size=7)
g.savefig(output_folder + 'fig2.png', bbox_inches='tight')
# +
# Figure 3
f, ax = plt.subplots(figsize=(8, 10))
sns.set_color_codes("pastel")
sns.barplot(x="length", y="target_id", data=abundances,
label="Length", color="b")
# Plot the crashes where alcohol was involved
sns.set_color_codes("muted")
sns.barplot(x="eff_length", y="target_id", data=abundances,
label="Effective Length", color="b")
ax.legend(ncol=2, loc="lower right", frameon=True)
ax.set(ylabel="",
xlabel="Length and Effective Length")
sns.despine(left=True, bottom=True)
f.savefig(output_folder + 'fig3.png', bbox_inches='tight')
# +
# Figure 4
f, ax = plt.subplots(figsize=(8, 10))
sns.set_color_codes("pastel")
sns.barplot(x="tpm", y="target_id", data=abundances,
label="TPM", color="b")
# Plot the crashes where alcohol was involved
sns.set_color_codes("muted")
sns.barplot(x="est_counts", y="target_id", data=abundances,
label="Estimated Counts", color="b")
ax.legend(ncol=2, loc="lower right", frameon=True)
ax.set(ylabel="",
xlabel="Counts and Transcripts Per Million")
sns.despine(left=True, bottom=True)
f.savefig(output_folder + 'fig4.png', bbox_inches='tight')
| Drone_Plotting.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Symbol
import mxnet as mx
a = mx.sym.Variable('a')
b = mx.sym.Variable('b')
c = a + b
(a, b, c)
# # Let's see what we have done in graph
mx.viz.plot_network(symbol=c)
# # Scalar, Vector, Matrix???????
shape = {'a':(1,1),'b':(1,1)} #follow NCHW
mx.viz.plot_network(symbol=c,shape=shape)
# # Numerical Results
a_val = mx.nd.ones((10,10),ctx=mx.cpu()) #use mx.gpu(#) if you are tuhao or you use AWS
b_val = mx.nd.ones((10,10),ctx=mx.cpu())
exe = c.bind(ctx=mx.cpu(0),args={'a':a_val, 'b':b_val})
exe
f = exe.forward()
f
f[0].asnumpy()
# # Let's do the same thing in NDArray section, with better big picture
# +
data = mx.sym.Variable('data')
label = mx.sym.Variable('softmax_label')
def cnn(x):
w_c = mx.sym.Variable('w_c')
conv = mx.sym.Convolution(data=data, weight=w_c, num_filter=32, kernel=(3,3), stride=(2,2), pad=(1,1), name='first_conv', no_bias=True)
act = mx.sym.Activation(data=conv, act_type='relu', name='first_relu')
pool = mx.sym.Pooling(data=act,stride=(2,2),kernel=(2,2),pool_type='max', name='first_pool')
w_fc = mx.sym.Variable('w_fc')
fc = mx.sym.FullyConnected(data=pool, weight=w_fc, num_hidden=10, name='first_fc',no_bias=True)
logit = fc
return logit
def cross_entropy(logit,y):
return mx.sym.softmax_cross_entropy(logit,y)
logit = cnn(data)
loss = cross_entropy(logit,label)
loss
# -
mx.viz.plot_network(loss)
# +
import matplotlib.pyplot as plt
import numpy as np
img_numpy = plt.imread('Lenna.png')
plt.imshow(img_numpy) # asnumpy is the magic communicating numpy and mxnet NDarray
plt.show()
img_T = img_numpy.transpose(1,2,0).reshape((1,3,512,512))
img_mx = mx.nd.array(img_T, ctx=mx.cpu())
label_mx = mx.nd.array([1], ctx=mx.cpu())
w_c_init_mx = mx.nd.array(np.ones((32,3,3,3)),ctx=mx.cpu())
w_fc_init_mx = mx.nd.array(np.ones((10,524288)),ctx=mx.cpu())
exe = loss.bind(ctx=mx.cpu(0),args={'data':img_mx,
'softmax_label':label_mx,
'w_c':w_c_init_mx,
'w_fc':w_fc_init_mx})
# -
y = exe.forward()
y[0].asnumpy()
| mxnet-week1/MXNet Sym.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import precip as pc
import pandas as pd
import matplotlib.pyplot as plt
from matplotlib import dates as mdates
plt.style.use('ggplot')
df = pd.read_csv('flow.csv', index_col=0, header=0, parse_dates=True)
precip = pd.read_csv('precip.csv', index_col=0, header=0, parse_dates=True)
precip = precip.resample('12H').sum()
# +
# pdf = pc.get_precip('ATC01_MADIS', '2015-04-22 00:00', '2015-5-31 23:00')
# del pdf['Snow Melt']
# pdf.columns = ['rainfall [in]']
# pdf.to_csv('precip.csv')
# pdf
# +
fig, axs = plt.subplots(3, 1, figsize=(8.5,5.5), sharex=True)
axs2 = [ax.twinx() for ax in axs]
for ax, ax2, column in zip(axs, axs2, df):
ax.plot(df[column], color='r')
ax2.bar(precip.index, precip['rainfall [in]'], color='#5287A7', alpha=0.35, width=0.5, align='edge')
ax.set_title(column)
ax.set_ylabel('Flow [gpm]', color='#BA3723')
ax2.set_ylabel('Rainfall [in]', color='#3a7998')
ax2.grid(b=False)
for tick_label in ax.get_yticklabels():
tick_label.set_color('#BA3723')
for tick_label in ax2.get_yticklabels():
tick_label.set_color('#5287A7')
dateFmt = mdates.DateFormatter('%b %d')
ax.xaxis.set_major_locator(mdates.DayLocator(interval=7))
ax.xaxis.set_major_formatter(dateFmt)
ax.xaxis.set_minor_locator(mdates.DayLocator(interval=1))
ax.grid(b=True, which='minor')
ax2.grid(b=False)
# -
plt.show()
# +
#fig.savefig('output')
# -
fig.savefig('output.png', dpi=600)
| assets/hydrographs_in_python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: credit_risk
# language: python
# name: credit_risk
# ---
# +
import os
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import missingno as msno
import pickle
sns.set()
# -
def distplot(series, **kwargs):
"""Create a figure with two subplots.
The lower part of the figure is distplot and the upper part display
a box plot for the same sample.
:arg:
series (pd.Series): The sample you want to plot.
kwargs : all keyword argument accepted by seaborn.distplot.
"""
# Cut the window in 2 parts
kwrgs = {"height_ratios": (.15, .85)}
f, (ax_box, ax_hist) = plt.subplots(2, sharex=True, figsize=(8, 8),
gridspec_kw=kwrgs)
# Add a graph in each part
sns.boxplot(series, ax=ax_box)
sns.distplot(series, ax=ax_hist, **kwargs)
# Remove x axis name for the boxplot
ax_box.set(xlabel='')
data = dict()
for dirname, _, filenames in os.walk('../data/raw'):
for filename in filenames:
print(filename)
# On recharge les données intégralement
app_train = pd.read_csv('../data/raw/application_train.csv')\
.sort_values('SK_ID_CURR')\
.reset_index(drop=True)
app_test = pd.read_csv('../data/raw/application_test.csv')\
.sort_values('SK_ID_CURR')\
.reset_index(drop=True)
bureau = pd.read_csv('../data/raw/bureau.csv')\
.sort_values(['SK_ID_CURR', 'SK_ID_BUREAU'])\
.reset_index(drop=True)
bureau_balance = pd.read_csv('../data/raw/bureau_balance.csv')\
.sort_values('SK_ID_BUREAU')\
.reset_index(drop=True)
cash = pd.read_csv('../data/raw/POS_CASH_balance.csv')\
.sort_values(['SK_ID_CURR', 'SK_ID_PREV'])\
.reset_index(drop=True)
credit = pd.read_csv('../data/raw/credit_card_balance.csv')\
.sort_values(['SK_ID_CURR', 'SK_ID_PREV'])\
.reset_index(drop=True)
previous = pd.read_csv('../data/raw/previous_application.csv')\
.sort_values(['SK_ID_CURR', 'SK_ID_PREV'])\
.reset_index(drop=True)
installments = pd.read_csv('../data/raw/installments_payments.csv')\
.sort_values(['SK_ID_CURR', 'SK_ID_PREV'])\
.reset_index(drop=True)
# ## Manual feature engineering
#
# On reprend les quelques variables qui ont le plus de sens. Voir Notebook '1.0-tg-initial-EDA'
# +
features = app_train[['SK_ID_CURR',
'DAYS_BIRTH', # Age
#'CODE_GENDER', # Sex
'OCCUPATION_TYPE', # Job
'AMT_INCOME_TOTAL', # Revenues
'AMT_CREDIT', # Credit amount
'NAME_CONTRACT_TYPE', # Contract type, Cash/Revolving
'AMT_ANNUITY', # Annuity amount
'EXT_SOURCE_1',
'EXT_SOURCE_2',
'EXT_SOURCE_3',
]]
features_test = app_test[['SK_ID_CURR',
'DAYS_BIRTH', # Age
#'CODE_GENDER', # Sex
'OCCUPATION_TYPE', # Job
'AMT_INCOME_TOTAL', # Revenues
'AMT_CREDIT', # Credit amount
'NAME_CONTRACT_TYPE', # Contract type, Cash/Revolving
'AMT_ANNUITY', # Annuity amount
'EXT_SOURCE_1',
'EXT_SOURCE_2',
'EXT_SOURCE_3',
]]
# -
features = pd.concat([features, features_test], axis=0)
features.shape
features
features['DAYS_BIRTH'] = features['DAYS_BIRTH'] / -365
features
# ### payment default
bureau
bureau = bureau.set_index('SK_ID_BUREAU').join(pd.get_dummies(bureau.set_index('SK_ID_BUREAU')['CREDIT_ACTIVE'], prefix='CREDIT_ACTIVE'))
bureau[bureau['SK_ID_CURR'] == 456116]
bureau_balance['STATUS'].replace('C', 0, inplace=True)
bureau_balance['STATUS'].replace('X', 0, inplace=True)
bureau_balance['STATUS'] = bureau_balance['STATUS'].astype('int')
count_late = bureau_balance.groupby('SK_ID_BUREAU')['STATUS'].sum()
count_late.describe()
bureau = pd.merge(bureau, count_late, left_on='SK_ID_BUREAU', right_on='SK_ID_BUREAU')
bureau.rename(columns={"STATUS": 'REPORTED_DPD'}, inplace=True)
bureau
bureau[[x for x in bureau.columns if x.startswith('CREDIT_ACTIVE_')] + ['SK_ID_CURR', 'REPORTED_DPD']].groupby('SK_ID_CURR').sum()
bureau_history = bureau[[x for x in bureau.columns if x.startswith('CREDIT_ACTIVE_')] + ['SK_ID_CURR', 'REPORTED_DPD']].groupby('SK_ID_CURR').sum()
features = features.set_index('SK_ID_CURR').join(bureau_history).reset_index()
features
features.columns
# No significant features.
# From feature importance with LightGBM
features.drop(columns=['CREDIT_ACTIVE_Bad debt', 'CREDIT_ACTIVE_Sold'], inplace=True)
features.join(app_train['TARGET'])
# ### payment default from HC
previous
cash
cash['SK_DPD'].describe()
installments
installments['BAD_PAYMENT_HC'] = installments['AMT_INSTALMENT'] != installments['AMT_PAYMENT']
installments['BAD_PAYMENT_HC'].describe()
bad_payment_hc_history = installments[['SK_ID_CURR', 'BAD_PAYMENT_HC']].groupby('SK_ID_CURR').count()
features = features.set_index('SK_ID_CURR').join(bad_payment_hc_history).sort_index()
previous
previous['CNT_PAYMENT']
previous['DAYS_TERMINATION'].describe()
previous['IS_ACTIVE'] = previous['DAYS_TERMINATION'].apply(lambda x: 1 if x > 0 else 0)
previous['IS_ACTIVE'].describe()
active_cred_hc = previous[['SK_ID_CURR', 'IS_ACTIVE']].groupby('SK_ID_CURR').sum()
features = features.join(active_cred_hc)
features.rename(columns={'IS_ACTIVE': 'ACTIVE_CRED_HC'}, inplace=True)
features
features.index
features['TOTAL_PREV_HC'] = previous[['SK_ID_CURR', 'IS_ACTIVE']].groupby('SK_ID_CURR').count()
features['TOTAL_PREV_HC'].describe()
features.describe()
features
features = features.join(app_train.set_index('SK_ID_CURR')['TARGET'])
features.to_csv('../data/processed/features.csv')
new_features = ["CREDIT_ACTIVE_Active",
"CREDIT_ACTIVE_Closed",
"REPORTED_DPD",
"BAD_PAYMENT_HC",
"ACTIVE_CRED_HC",
"TOTAL_PREV_HC"]
for feature in new_features:
distplot(features[feature])
# ### Baseline
# Add TARGET and make a logistic reg.
# +
from sklearn.preprocessing import LabelEncoder
le_ = 0
categ_var = list()
label_encoders = dict()
for col in features.columns:
if features[col].dtype.name == 'object':
if len(features[col].unique()) <= 2:
# label encoder
le = LabelEncoder()
print('Encoding %s' % col)
le.fit(features[col])
features[col] = le.transform(features[col])
le_ += 1
categ_var.append(col)
label_encoders[col] = le
print(f"{le_} columns encoded")
# -
for col in features.columns:
if col not in app_train.columns:
features[col].fillna(0, inplace=True)
features_ohe = pd.get_dummies(features)
# +
features_le = features.copy()
le_ = 0
for col in features.columns:
if features[col].dtype.name == 'object':
if len(features[col].unique()) >= 2:
le = LabelEncoder()
# label encoder
print('Encoding %s' % col)
le.fit(features[col].astype(str))
features_le[col] = le.transform(features[col].astype(str))
le_ += 1
categ_var.append(col)
label_encoders[col] = le
print(f"{le_} columns encoded")
# -
# #### Baseline Logistic reg
# +
from sklearn.impute import SimpleImputer
from sklearn.preprocessing import MinMaxScaler
train = features_ohe[features_ohe['TARGET'].notna()].copy()
test = features_ohe[features_ohe['TARGET'].isna()].copy()
target = train['TARGET']
train.drop(columns=['TARGET'], inplace=True)
test.drop(columns=['TARGET'], inplace=True)
imputer = SimpleImputer(strategy='constant', fill_value=0)
scaler = MinMaxScaler(feature_range=(0, 1))
imputer.fit(train)
scaler.fit(train)
train = imputer.transform(train)
test = imputer.transform(test)
train = scaler.transform(train)
test = scaler.transform(test)
features_names = list(features_ohe.drop(columns=['TARGET']).columns)
print(f'train set shape : {train.shape}')
print(f'test set shape : {test.shape}')
# +
from sklearn.linear_model import LogisticRegression
reg = LogisticRegression(C=1e-4)
reg.fit(train, target)
# -
baseline_results = app_test[['SK_ID_CURR']].copy()
baseline_results['TARGET'] = reg.predict_proba(test)[:, 1]
path = os.path.join(os.path.abspath('../reports/'), 'logistic_reg_features_engineering_baseline.csv')
baseline_results.to_csv(path, index=False)
features.to_csv('../data/interim/features.csv')
# #### Résultats:
#
# **ROC_AUC = 0.68867**
# ### LightGBM
import lightgbm as lgb
features_le
# +
from sklearn.impute import SimpleImputer
from sklearn.preprocessing import MinMaxScaler
train = features_le[features_le['TARGET'].notna()].copy()
test = features_le[features_le['TARGET'].isna()].copy()
target = train['TARGET']
train.drop(columns=['TARGET'], inplace=True)
test.drop(columns=['TARGET'], inplace=True)
imputer = SimpleImputer(strategy='constant', fill_value=0)
scaler = MinMaxScaler(feature_range=(0, 1))
imputer.fit(train)
scaler.fit(train)
train = imputer.transform(train)
test = imputer.transform(test)
train = scaler.transform(train)
test = scaler.transform(test)
features_names = list(features_le.drop(columns=['TARGET']).columns)
print(f'train set shape : {train.shape}')
print(f'test set shape : {test.shape}')
# -
features_le
pd.DataFrame(train, columns=features_names, index=app_train['SK_ID_CURR'])\
.join(app_train[['SK_ID_CURR', 'TARGET']].set_index("SK_ID_CURR"))
# +
train_data = lgb.Dataset(train, label=target,
feature_name=features_names,
categorical_feature=categ_var)
param = {'num_leaves': 31, 'objective': 'binary'}
param['metric'] = 'auc'
num_round = 100
# -
bst = lgb.train(param, train_data, num_round)
baseline_results = app_test[['SK_ID_CURR']].copy()
baseline_results['TARGET'] = bst.predict(test)
path = os.path.join(os.path.abspath('../reports/'), 'lgbm.csv')
baseline_results.to_csv(path, index=False)
# #### résultats:
#
# ROC_AUC = 0.72253
fig, ax = plt.subplots(1, figsize=(12, 8))
lgb.plot_importance(bst, ignore_zero=False, ax=ax)
# ### LightGBM optimization
# +
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(train, target)
# -
train_data = lgb.Dataset(X_train, label=y_train, feature_name=features_names)
test_data = lgb.Dataset(X_test, label=y_test, feature_name=features_names, reference=train_data)
evals_results = dict()
# +
param = {'num_leaves': 31, 'objective': 'binary'}
param['metric'] = 'auc'
num_round = 1000
# -
bst = lgb.train(param, train_data, num_boost_round=num_round,
verbose_eval=1, evals_result=evals_results,
valid_sets=[train_data, test_data],
early_stopping_rounds=30)
fig, ax = plt.subplots(1, figsize=(12, 8))
sns.lineplot(x=range(len(evals_results['training']['auc'])),
y=evals_results['training']['auc'], label='train')
sns.lineplot(x=range(len(evals_results['valid_1']['auc'])),
y=evals_results['valid_1']['auc'], label='test')
plt.legend()
plt.show()
evals_results.keys()
baseline_results = app_test[['SK_ID_CURR']].copy()
baseline_results['TARGET'] = bst.predict(test)
path = os.path.join(os.path.abspath('../reports/'), 'lgbm_early_stop.csv')
baseline_results.to_csv(path, index=False)
# ### Résultats LightGBM
#
# **ROC_AUC = 0.73572**
# ### Interprétabilité du modèle
# +
import shap
shap.initjs()
# -
train = features_le[features_le['TARGET'].notna()].copy()
sample_data = pd.DataFrame(train.drop(columns=['TARGET']),
columns=features_names).sample(100, random_state=100)
index = sample_data.index
sample_data = MinMaxScaler(feature_range=[0, 1]).fit_transform(sample_data)
sample_data = SimpleImputer(strategy='constant', fill_value=0).fit_transform(sample_data)
sample_data = pd.DataFrame(sample_data, columns=features_names, index=index)
explainer = shap.TreeExplainer(bst, data=sample_data, model_output='probability')
shap_values = explainer.shap_values(sample_data)
sample_data
shap_values.shape
sample_data.shape
shap.summary_plot(shap_values, features=sample_data, feature_names=features_names)
explainer.expected_value
plot = shap.force_plot(explainer.expected_value,
shap_values[0,:], features=sample_data.iloc[0, :])
shap.save_html('../reports/figures/explainer_single.html', plot)
plot = shap.force_plot(explainer.expected_value,
shap_values, features=sample_data)
shap.save_html('../reports/figures/explainer_multi.html', plot)
features_le.to_csv('../data/processed/features.csv')
| notebooks/2.0-feature-engineering.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] nbgrader={"grade": false, "grade_id": "q3_prompt", "locked": true, "schema_version": 1, "solution": false}
# # Q3
#
# ### THIS QUESTION IS OPTIONAL. You do not need to install Python on your machine! Please still fill out the question box below in A.
#
# To repeat: installing Python on your computer is *optional*. You are more than welcome to use JupyterHub for all your Python needs.
#
# But if you want, it's pretty easy to install Python. Important point: if you install Python on your machine, **do NOT install the version on python.org**. It's difficult to configure, and doesn't include any additional functionality that we'll be using in this class.
#
# Instead, we'll use a pre-made Python distribution: **Anaconda**. Anaconda is basically a fully-functional, prepackaged Python distribution that is ready-built for any of the three major operating systems: Windows, macOS, and Linux. Another benefit is that Anaconda is self-contained, so if you accidentally mess up the install, you can simply delete it and start over.
# + [markdown] nbgrader={"grade": false, "grade_id": "q3a_prompt", "locked": true, "schema_version": 1, "solution": false}
# ### Part A
#
# Go to the Anaconda downloads page: [https://www.continuum.io/downloads](https://www.continuum.io/downloads) . Click on the link for your operating system, and download the graphical installer (you'll most likely want 64-bit).
#
# **MAKE SURE YOU DOWNLOAD ANACONDA FOR PYTHON 3**.
#
# Once the install is finished, let's test that it worked: Fire up a command prompt / Terminal window. Type `python`.
#
# If all is well, you should see exactly what we saw in the JupyterHub terminal.
#
# 
#
# ### In the box below, inform me whether or not you were able to successfully complete A. If you did not attempt A, please mention that as well, and if you're comfortable with it, your reasons why.
# + [markdown] nbgrader={"grade": true, "grade_id": "q3a", "locked": false, "points": 25, "schema_version": 1, "solution": true}
#
| assignments/A0/A0_Q3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
dataset = pd.read_csv('C:/Users/Gray/Desktop/dataset/kc_house_data.csv')
#dataset.head(10)
#X = dataset.iloc[:, [5,6,12,13,19,20]]
#X = dataset.iloc[:, [3, 4, 5, 6, 7, 8, 9, 10, 11, 14, 15, 19, 20]]
X = dataset.iloc[:, 3:]
Y = dataset.iloc[:, [2]]
X
# +
# Normalisation
meu = np.mean(X)
std = np.std(X)
X = (X-meu)/std
# -
X
# +
#meu
# +
#std
# -
X_train = X.iloc[:17290, :]
X_test = X.iloc[17289:, :]
Y_train = Y.iloc[:17290, :]
Y_test = Y.iloc[17289:, :]
def biasing_X(X):
N = X.shape
arr = np.ones((N[0], 1))
X = np.hstack((arr, X))
w = np.zeros((N[1]+1, 1))
return X, w
X_bias, theta = biasing_X(X_train)
# +
X_bias = np.array(X_bias)
X_bias = X_bias.T
print(X_bias.shape)
# -
theta = np.array(theta)
theta.shape
def hypo(X, W):
return np.dot(X.T, W)
def cost(X, Y, W):
m = X.shape[1]
#print(m)
n = X.shape[0]
sum = 0
Temp = hypo(X, W) - Y
J = (np.dot(Temp.T, Temp))/(2*m)
#print(J)
return J
def gradient_descent(X, W, Y, Alpha, Step):
N = X.shape[1]
print(N)
J = []
initial = cost(X, Y, W)
J.append(initial)
#np.dot(x.T, (y_hat-y)))
print(J)
for i in range(Step):
Te = hypo(X, W)
Te = Te - Y
K = X@Te
print(i)
#print(K.shape)
#print(K.T)
W = W - (Alpha/N)*K
Temp = cost(X, Y, W)
J.append(Temp)
print(Temp)
return J, W
J, W = gradient_descent(X_bias, theta, Y_train, 0.05, 1000)
print(W) # value from Gradient Descent
theta1 = np.linalg.pinv(X_bias@X_bias.T)@(X_bias@Y_train)
theta1 #value from formula
def prediction(W1):
X_b1, _ = biasing_X(X_test)
Y_pred = X_b1@W1
return Y_pred
y_bar = np.mean(Y_test)
y_bar
def error(Y_pp):
R_error = ((Y_pp - y_bar).T@(Y_pp - y_bar)) /((Y_test - y_bar).T@(Y_test- y_bar))
return R_error
Y_pred1 = prediction(W)
error(Y_pred1) # predicted by descent
# +
Y_pred = prediction(theta1)
error(Y_pred) # predicted by formula
# -
Y_pred1
Y_pred
Y_pred = np.array(Y_pred)
Y_pred1 = np.array(Y_pred1)
mse = (np.sum(Y_pred - Y_test)**2)/(Y_test.shape[0])
mse1 = (np.sum(Y_pred1 - Y_test)**2)/(Y_test.shape[0])
np.sqrt(mse)
np.sqrt(mse1)
from sklearn.linear_model import LinearRegression
regressor = LinearRegression()
regressor.fit(X_train, Y_train)
print('Variance score: %.2f' % regressor.score(X_test, Y_test))
# +
from sklearn.model_selection import cross_val_score
ols2 = LinearRegression()
# X_T_1 = np.reshape(,1)
print(X_train.shape)
ols_cv_mse = cross_val_score(ols2, X_train, Y_train, scoring='v_measure_score', cv=10)
ols_cv_mse.mean()
# -
import sklearn
sorted(sklearn.metrics.SCORERS.keys())
| Edureka_MicroCourse/multivariate_gradientDescent.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="NGGrt9EYlCqY"
# <img align="left" src="https://lever-client-logos.s3.amazonaws.com/864372b1-534c-480e-acd5-9711f850815c-1524247202159.png" width=200>
# <br></br>
#
# # Backpropagation Practice
#
# ## *Data Science Unit 4 Sprint 2 Assignment 2*
#
# Implement a 3 input, 4 node hidden-layer, 1 output node Multilayer Perceptron on the following dataset:
#
# | x1 | x2 | x3 | y |
# |----|----|----|---|
# | 0 | 0 | 1 | 0 |
# | 0 | 1 | 1 | 1 |
# | 1 | 0 | 1 | 1 |
# | 0 | 1 | 0 | 1 |
# | 1 | 0 | 0 | 1 |
# | 1 | 1 | 1 | 0 |
# | 0 | 0 | 0 | 0 |
#
# If you look at the data you'll notice that the first two columns behave like an XOR gate while the last column is mostly just noise. Remember that creating an XOR gate was what the perceptron was criticized for not being able to learn.
# + colab_type="code" id="nEREYT-3wI1f" colab={"base_uri": "https://localhost:8080/", "height": 289} outputId="a69aeb33-0a81-4e20-f46d-8d52f2473573"
import numpy as np
np.random.seed(42)
# 3 input nodes
X = np.array(([0,0,1], [0,1,1], [1,0,1], [0,1,0], [1,0,0], [1,1,1], [0,0,0]), dtype=float)
# 1 output node
y = np.array(([0], [1], [1], [1], [1], [0], [0]), dtype=float)
print("3 inputs \n", X)
print("1 output \n", y)
# + id="7ewIvPV89bia" colab_type="code" colab={}
class NeuralNetwork:
def __init__(self):
# Set up Architecture of Neural Network
self.inputs = 3
self.hiddenNodes = 4
self.outputNodes = 1
# Initial Weights
# 3x4 Matrix Array for the First Layer
self.weights1 = np.random.randn(self.inputs, self.hiddenNodes)
# 4x1 Matrix Array for Hidden to Output
self.weights2 = np.random.rand(self.hiddenNodes, self.outputNodes)
def sigmoid(self, s):
return 1 / (1 + np.exp(-s))
def sigmoidPrime(self, s):
return s * (1 - s)
def feed_forward(self, X):
"""
Calculate the NN inference using feed forward.
aka "predict"
"""
# Weighted sum of inputs => hidden layer
self.hidden_sum = np.dot(X, self.weights1)
# Activations of weighted sum
self.activated_hidden = self.sigmoid(self.hidden_sum)
# Weight sum between hidden and output
self.output_sum = np.dot(self.activated_hidden, self.weights2)
# Final activation of output
self.activated_output = self.sigmoid(self.output_sum)
return self.activated_output
def backward(self, X, y, o):
"""
Backward propagate through the network
"""
# Error in output
self.o_error = y - o
# Apply derivative of sigmoid to error
# How far off are we in relation to the Sigmoid f(x) of the output
# ^- hidden => output
self.o_delta = self.o_error * self.sigmoidPrime(o)
# z2 error
self.z2_error = self.o_delta.dot(self.weights2.T)
# How much of that "far off" can be explained by the inputs => hidden layer
self.z2_delta = self.z2_error * self.sigmoidPrime(self.activated_hidden)
# Adjustment to first set of weights (input => hidden)
self.weights1 += X.T.dot(self.z2_delta)
# Adjustment to second set of weights (hidden => output)
self.weights2 += self.activated_hidden.T.dot(self.o_delta)
def train(self, X, y):
o = self.feed_forward(X)
self.backward(X, y, o)
# + id="rK8cBzJ-9kjI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 986} outputId="e538571f-0fec-4020-b2f0-452e7cc17f7b"
# Train the NN
nn = NeuralNetwork()
# number of epochs / iterations
for i in range(1000):
if (i+1 in [1,2,3]) or ((i+1) % 50 ==0):
print('+' + '---' * 3 + f'EPOCH {i+1}' + '---'*3 + '+')
# print('Input: \n', X)
# print('Actual Output: \n', y)
# print('Predicted Output: \n', str(nn.feed_forward(X)))
loss = np.mean(np.square(y - nn.feed_forward(X)))
if loss < .01:
print('Input: \n', X)
print('Actual Output: \n', y)
print('Predicted Output: \n', str(nn.feed_forward(X)))
print("Loss: \n", str(loss))
break
print("Loss: \n", str(loss))
nn.train(X,y)
# + [markdown] colab_type="text" id="8b-r70o8p2Dm"
# ## Try building/training a more complex MLP on a bigger dataset.
#
# Use the [MNIST dataset](http://yann.lecun.com/exdb/mnist/) to build the cannonical handwriting digit recognizer and see what kind of accuracy you can achieve.
#
# If you need inspiration, the internet is chalk-full of tutorials, but I want you to see how far you can get on your own first. I've linked to the original MNIST dataset above but it will probably be easier to download data through a neural network library. If you reference outside resources make sure you understand every line of code that you're using from other sources, and share with your fellow students helpful resources that you find.
#
#
# ### Parts
# 1. Gathering & Transforming the Data
# 2. Making MNIST a Binary Problem
# 3. Estimating your Neural Network (the part you focus on)
# + [markdown] id="-jTGR8LJ9HGE" colab_type="text"
# ### Gathering the Data
#
# `keras` has a handy method to pull the mnist dataset for you. You'll notice that each observation is a 28x28 arrary which represents an image. Although most Neural Network frameworks can handle higher dimensional data, that is more overhead than necessary for us. We need to flatten the image to one long row which will be 784 values (28X28). Basically, you will be appending each row to one another to make on really long row.
# + id="bfFf-_qp9HGF" colab_type="code" colab={}
import numpy as np
from tensorflow.keras.datasets import mnist
from tensorflow.keras.utils import to_categorical
# + id="n_iuc91W9HGI" colab_type="code" colab={}
# input image dimensions
img_rows, img_cols = 28, 28
# + id="XNYmBUx69HGN" colab_type="code" colab={}
# the data, split between train and test sets
(X_train, y_train), (X_test, y_test) = mnist.load_data()
# + id="s9cJCHci9HGR" colab_type="code" colab={}
X_train = X_train.reshape(x_train.shape[0], img_rows * img_cols)
X_test = X_test.reshape(x_test.shape[0], img_rows * img_cols)
# Normalize Our Data
X_train = x_train / 255
X_test = x_test / 255
# + id="H3W8IoTN9HGU" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="17ab4267-3974-438b-c742-5ddbb217d915"
# Now the data should be in a format you're more familiar with
X_train.shape
# + [markdown] id="G6KUSDve9HGY" colab_type="text"
# ### Making MNIST a Binary Problem
# MNIST is multiclass classification problem; however we haven't covered all the necessary techniques to handle this yet. You would need to one-hot encode the target, use a different loss metric, and use softmax activations for the last layer. This is all stuff we'll cover later this week, but let us simply the problem for now: Zero or all else.
# + id="N00wf8PM9HGZ" colab_type="code" colab={}
import numpy as np
y_temp = np.zeros(y_train.shape)
y_temp[np.where(y_train == 0.0)[0]] = 1
y_train = y_temp
y_temp = np.zeros(y_test.shape)
y_temp[np.where(y_test == 0.0)[0]] = 1
y_test = y_temp
# + id="hHT4iQdA9HGc" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="5f324bfc-1239-43cd-f612-933bab92c3ce"
# A Nice Binary target for ya to work with
y_train.shape
# + [markdown] id="kDCOKJUj9HGf" colab_type="text"
# ### Estimating Your `net
# + colab_type="code" id="5MOPtYdk1HgA" colab={"base_uri": "https://localhost:8080/", "height": 476} outputId="38060726-995e-4533-b9e3-810740940674"
for i in range(2000):
if (i+1 in [1,2,3]) or ((i+1) % 50 ==0):
print('+' + '---' * 3 + f'EPOCH {i+1}' + '---'*3 + '+')
# print('Input: \n', X)
# print('Actual Output: \n', y)
# print('Predicted Output: \n', str(nn.feed_forward(X)))
loss = np.mean(np.square(y - nn.feed_forward(X)))
if loss < .001:
print('Input: \n', X)
print('Actual Output: \n', y)
print('Predicted Output: \n', str(nn.feed_forward(X)))
print("Loss: \n", str(loss))
break
print("Loss: \n", str(loss))
nn.train(X_train,y_train)
# + [markdown] colab_type="text" id="FwlRJSfBlCvy"
# ## Stretch Goals:
#
# - Make MNIST a multiclass problem using cross entropy & soft-max
# - Implement Cross Validation model evaluation on your MNIST implementation
# - Research different [Gradient Descent Based Optimizers](https://keras.io/optimizers/)
# - [<NAME> the evolution of gradient descent](https://www.youtube.com/watch?v=nhqo0u1a6fw)
# - Build a housing price estimation model using a neural network. How does its accuracy compare with the regression models that we fit earlier on in class?
| curriculum/unit-4-machine-learning/sprint-2-intro-nn/module2-backpropagation/LS_DS_422_Backprop_Assignment.ipynb |
# ---
# title: "Random Forest Regression"
# author: "<NAME>"
# date: 2017-12-20T11:53:49-07:00
# description: "Training a random forest regressor in scikit-learn."
# type: technical_note
# draft: false
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Preliminaries
# Load libraries
from sklearn.ensemble import RandomForestRegressor
from sklearn import datasets
# ## Load Boston Housing Data
# Load data with only two features
boston = datasets.load_boston()
X = boston.data[:,0:2]
y = boston.target
# ## Create Random Forest Regressor
# Create decision tree classifer object
regr = RandomForestRegressor(random_state=0, n_jobs=-1)
# ## Train Random Forest Regressor
# Train model
model = regr.fit(X, y)
| docs/machine_learning/trees_and_forests/random_forest_regressor.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
# write raw rbnf source code.
from rbnf.easy import build_parser, Language, build_language
from typing import NamedTuple, List
my_lisp_definition = """
ignore [space]
space := R'\s'
term := R'[^\(\)\s]'
sexpr ::= '(' [sexpr as head sexpr* as tail] ')' | term as atom
rewrite
if atom:
return atom
if not head:
return ()
return SExpr(head, tail)
"""
class SExpr(NamedTuple):
head: 'SExpr'
tail: List['SExpr']
lisp = Language('lisp')
lisp.namespace['SExpr'] = SExpr
build_language(my_lisp_definition, lisp, '<tutorials>')
parse = build_parser(lisp)
parse("(+ 1 (* 2 3))").result
# +
# Use Python to write rbnf
from rbnf.easy import Parser, Lexer, Language, auto_context, C, build_parser
from rbnf.core.State import State
from rbnf.core.Tokenizer import Tokenizer
from typing import NamedTuple, List, Optional
lisp = Language('lisp')
lisp.ignore('space')
@lisp
class space(Lexer):
@staticmethod
def regex():
return '\s'
@lisp
class term(Lexer):
@staticmethod
def regex():
return '[^\(\)\s]'
@lisp
class sexpr(Parser):
@staticmethod
def bnf():
return C('(') + (sexpr @ "head" + sexpr.unlimited @ "tail").optional + C(")") | term @ "term"
@staticmethod
@auto_context
def rewrite(state: State):
head: Optional[SExpr]
tail: Optional[List[SExpr]]
term: Optional[Tokenizer]
if term:
return term
return SExpr(head, tail)
class SExpr(NamedTuple):
head: 'SExpr'
tail: List['SExpr']
lisp.build()
parse = build_parser(lisp, use_parser='sexpr')
parse("(+ 1 (* 2 3))").result
# +
# About literal parsers
from rbnf.core.Tokenizer import Tokenizer
from rbnf.core.CachingPool import ConstStrPool
from rbnf.core.State import State
from rbnf.easy import N, NC, C, R, V
const_value = ConstStrPool.cast_to_const('<value>')
name = ConstStrPool.cast_to_const('<name>')
tokens0 = [Tokenizer('<name>x'[:-1], const_value, lineno=0, colno=0)]
tokens1 = [Tokenizer(name, '<value>x'[:-1], lineno=0, colno=0)]
tokens2 = [Tokenizer(name, const_value, lineno=0, colno=0)]
make_state = lambda : State({}, '<playground>')
# +
name_parser = N('<name>')
print('parse tokens0: ', name_parser.match(tokens0, make_state()))
print('parse tokens1:', name_parser.match(tokens1, make_state()))
print('parse tokens2:', name_parser.match(tokens2, make_state()))
# +
value_parser = V('<value>')
print('parse tokens0: ', value_parser.match(tokens0, make_state()))
print('parse tokens1:', value_parser.match(tokens1, make_state()))
print('parse tokens2:', value_parser.match(tokens2, make_state()))
# +
value_parser = C('<value>')
print('parse tokens0: ', value_parser.match(tokens0, make_state()))
print('parse tokens1:', value_parser.match(tokens1, make_state()))
print('parse tokens2:', value_parser.match(tokens2, make_state()))
# -
regex_parser = R("\S+")
# Not recommend to use regex parser when parsing.
# Lexers use regex to tokenize raw input into tokenizers
# with constant names which could be compared by memory address
print('parse tokens0: ', regex_parser.match(tokens0, make_state()))
print('parse tokens1:', regex_parser.match(tokens1, make_state()))
print('parse tokens2:', regex_parser.match(tokens2, make_state()))
# +
name_and_value = NC(name, const_value)
print('parse tokens0: ', name_and_value.match(tokens0, make_state()))
print('parse tokens1:', name_and_value.match(tokens1, make_state()))
print('parse tokens2:', name_and_value.match(tokens2, make_state()))
# +
# TO BE CONTINUE
| tutorials.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="SpGU-Qe7dRMy"
# ```
# // Copyright 2020 Twitter, Inc.
# // SPDX-License-Identifier: Apache-2.0
# ```
#
# # Multilingual Alignment of mBERT via Translation Pair Prediction
#
# This notebook can be used to add the Translation Pair Pre-training task to multilingual models. We demonstrate our usecase using mBERT.
#
# + [markdown] id="FiBMR4ZMtWql"
# ## Setup libraries
#
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 126558, "status": "ok", "timestamp": 1631941385505, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GijxQCX_py83nArX6F4S58845Caf6w8Apxd0EpS=s64", "userId": "02687645879366791042"}, "user_tz": 240} id="zj19wTW-2Eha" outputId="10d9b4a3-3a9a-4fcf-8823-22e6c2ff27d8"
# %pip install transformers==3.5.1 datasets==1.1.2 torch==1.4.0 seqeval==1.2.2 gensim==3.8.1
# + [markdown] id="C6t__yBzfcn7"
# ## Define parameters
#
# + id="GE1uhBC-cqxK"
EN_ONLY_MODEL = False
WIKIDATA_MODEL = True
# Options for WIKIDATA_PREFIX: en_ja, en_ar, en_hi, en_hi_ja_ar_equal en_hi_ja_ar
WIKIDATA_PREFIX = "en_ja"
HOMEDIR = "./"
DATADIR = f"{HOMEDIR}/tatoeba"
pre_trained_model_path = "bert-base-multilingual-uncased"
# + [markdown] id="BRkive45crz2"
# ## Setup Helpers
#
# + executionInfo={"elapsed": 3537, "status": "ok", "timestamp": 1631941389038, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GijxQCX_py83nArX6F4S58845Caf6w8Apxd0EpS=s64", "userId": "02687645879366791042"}, "user_tz": 240} id="tWjzYQzH2m-Y"
import json
import random
import re
from dataclasses import dataclass
from itertools import combinations
from pathlib import Path
from typing import Any, Callable, Dict, List, NewType, Optional, Tuple, Union
import numpy as np
import torch
from sklearn.metrics import classification_report
from torch.nn import BCEWithLogitsLoss, CosineEmbeddingLoss, CrossEntropyLoss, MSELoss
from torch.nn.utils.rnn import pad_sequence
from torch.utils.data import IterableDataset
from tqdm import tqdm, trange
from transformers import (
BertForNextSentencePrediction,
BertTokenizerFast,
DataCollatorWithPadding,
EvalPrediction,
RobertaTokenizerFast,
Trainer,
TrainingArguments,
)
from transformers.modeling_bert import BertModel, BertOnlyNSPHead, BertPreTrainedModel
from transformers.modeling_outputs import NextSentencePredictorOutput
from transformers.tokenization_utils_base import (
BatchEncoding,
PaddingStrategy,
PreTrainedTokenizerBase,
)
# + executionInfo={"elapsed": 148, "status": "ok", "timestamp": 1631941389184, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GijxQCX_py83nArX6F4S58845Caf6w8Apxd0EpS=s64", "userId": "02687645879366791042"}, "user_tz": 240} id="k6Z0Fo-D2rwn"
class BertTwoTowerHead(torch.nn.Module):
def __init__(self, config):
super().__init__()
self.projection = torch.nn.Linear(config.hidden_size, config.hidden_size)
def forward(self, pooled_output):
projection = self.projection(pooled_output)
return projection
class BertTwoTowerLoss(torch.nn.Module):
def __init__(self, loss_type, margin=0.0):
super().__init__()
if loss_type == "bce":
self.loss_compute = self._bce_loss()
elif loss_type == "cosine":
self.loss_compute = self._cosine_loss(margin)
else:
raise NotImplementedError(f"loss_type={loss_type} not implemented")
def _bce_loss(self):
loss_fn = BCEWithLogitsLoss()
def _loss_compute(x1, x2, labels):
seq_relationship_scores = (x1 * x2).sum(axis=-1).view(-1)
labels = labels.view(-1) * 1.0
loss = loss_fn(seq_relationship_scores, labels)
return loss
return _loss_compute
def _cosine_loss(self, margin):
loss_fn = CosineEmbeddingLoss(margin=margin)
def _loss_compute(x1, x2, labels):
x1 = x1.view(-1, x1.shape[-1])
x2 = x2.view(-1, x2.shape[-1])
labels = labels.view(-1)
neg_label = torch.tensor(-1, device=labels.device)
labels = torch.where(labels > 0, labels, neg_label) # Cosine requires this
loss = loss_fn(x1, x2, labels)
return loss
return _loss_compute
def forward(self, x1, x2, labels):
loss = self.loss_compute(x1, x2, labels)
return loss
class BertForTwoTowerPrediction(BertPreTrainedModel):
def __init__(self, config, loss_type="bce", margin=0.0):
super().__init__(config)
self.bert = BertModel(config)
self.cls = BertTwoTowerHead(config)
self.loss_fn = BertTwoTowerLoss(loss_type, margin)
self.init_weights()
def forward(
self,
input_ids=None,
attention_mask=None,
token_type_ids=None,
position_ids=None,
head_mask=None,
inputs_embeds=None,
labels=None,
output_attentions=None,
output_hidden_states=None,
return_dict=None,
**kwargs,
):
r"""
labels (:obj:`torch.LongTensor` of shape :obj:`(batch_size,)`, `optional`):
Labels for computing the next sequence prediction (classification) loss. Input should be a sequence pair
(see ``input_ids`` docstring). Indices should be in ``[0, 1]``:
- 0 indicates sequence B is a continuation of sequence A,
- 1 indicates sequence B is a random sequence.
Returns:
Example::
>>> from transformers import BertTokenizer, BertForNextSentencePrediction
>>> import torch
>>> tokenizer = BertTokenizer.from_pretrained('bert-base-uncased')
>>> model = BertForNextSentencePrediction.from_pretrained('bert-base-uncased')
>>> prompt = "In Italy, pizza served in formal settings, such as at a restaurant, is presented unsliced."
>>> next_sentence = "The sky is blue due to the shorter wavelength of blue light."
>>> encoding = tokenizer(prompt, next_sentence, return_tensors='pt')
>>> outputs = model(**encoding, labels=torch.LongTensor([1]))
>>> logits = outputs.logits
>>> assert logits[0, 0] < logits[0, 1] # next sentence was random
"""
if "next_sentence_label" in kwargs:
warnings.warn(
"The `next_sentence_label` argument is deprecated and will be removed in a future version, use `labels` instead.",
FutureWarning,
)
labels = kwargs.pop("next_sentence_label")
return_dict = (
return_dict if return_dict is not None else self.config.use_return_dict
)
outputs = self.bert(
input_ids,
attention_mask=attention_mask,
token_type_ids=token_type_ids,
position_ids=position_ids,
head_mask=head_mask,
inputs_embeds=inputs_embeds,
output_attentions=output_attentions,
output_hidden_states=output_hidden_states,
return_dict=return_dict,
)
pooled_output = self.cls(outputs[1])
pooled_output_pair = pooled_output.reshape(-1, 2, *pooled_output.shape[1:])
# Dot product
x1 = pooled_output_pair[:, 0]
x2 = pooled_output_pair[:, 1]
seq_relationship_scores = (x1 * x2).sum(axis=-1)
# Only take the label of the first pair, cast to float
labels = labels.reshape(-1, 2, *labels.shape[1:])[:, 0] # * 1.0
next_sentence_loss = None
if labels is not None:
next_sentence_loss = self.loss_fn(x1, x2, labels)
if not return_dict:
output = (seq_relationship_scores,) + outputs[2:]
return (
((next_sentence_loss,) + output)
if next_sentence_loss is not None
else output
)
return NextSentencePredictorOutput(
loss=next_sentence_loss,
logits=seq_relationship_scores,
hidden_states=outputs.hidden_states,
attentions=outputs.attentions,
)
# + executionInfo={"elapsed": 115, "status": "ok", "timestamp": 1631941389297, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GijxQCX_py83nArX6F4S58845Caf6w8Apxd0EpS=s64", "userId": "02687645879366791042"}, "user_tz": 240} id="JxXbdzu72wjZ"
URL_REGEX = re.compile(r"http[s]?://[^ ]+")
def clean_text(text):
text = text.replace("\n", " [LF] ")
return URL_REGEX.sub("[URL]", text)
def read_file(file_path):
with open(file_path) as fp:
for line in fp:
line = line.strip()
if not line:
continue
line = json.loads(line)
num_samples = 5
line = [clean_text(t) for t in line["unique_label_desc"]]
line = np.random.permutation(line)[:num_samples].tolist()
n_tweets = len(line)
if n_tweets < 1:
continue
yield line, n_tweets
def get_tweet_pairs(tweet_list):
np.random.shuffle(tweet_list)
for t1, t2 in combinations(tweet_list, 2):
yield [t1, t2]
def get_all_pairs(url_tweets, n_url_tweets, num_negatives=1):
"""
file_data: list of list of tweets related to a URL
"""
n_urls = len(url_tweets)
n_url_tweets = np.array(n_url_tweets)
max_n_tweets = max(n_url_tweets)
for i, tweet_list in enumerate(url_tweets):
n_tweets = n_url_tweets[i]
n_pairs = n_tweets * (n_tweets - 1) // 2
neg_url_indexes = np.concatenate(
[np.arange(i), np.arange(i + 1, n_urls)]
) # Skip this URL
neg_url_samples = np.random.choice(neg_url_indexes, (n_pairs, num_negatives))
neg_url_positions = np.random.randint(
2, size=(n_pairs, num_negatives)
) # Positions
neg_tweet_indexes = np.random.randint(
max_n_tweets, size=(n_pairs, num_negatives)
)
neg_tweet_indexes = neg_tweet_indexes % n_url_tweets[neg_url_samples]
for j, pair in enumerate(get_tweet_pairs(tweet_list)):
yield pair, [
1,
1,
] # Pos data <-- This needs to be made 0 for compatibility with API
for nj in range(num_negatives):
neg_pair_position = neg_url_positions[j][nj]
neg_url_idx = neg_url_samples[j][nj]
neg_tweet_idx = neg_tweet_indexes[j][nj] # % n_url_tweets[neg_url_idx]
neg_tweet = url_tweets[neg_url_idx][neg_tweet_idx]
neg_pair = (
[pair[0], neg_tweet]
if neg_pair_position == 1
else [neg_tweet, pair[1]]
)
yield neg_pair, [
0,
0,
] # Neg data <-- This needs to be made 1 for compatibility with API
def read_sentencepair_data(file_paths):
for file_path in file_paths:
url_tweets, n_url_tweets = zip(*read_file(file_path))
yield from get_all_pairs(url_tweets, n_url_tweets)
# + executionInfo={"elapsed": 2, "status": "ok", "timestamp": 1631941389297, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GijxQCX_py83nArX6F4S58845Caf6w8Apxd0EpS=s64", "userId": "02687645879366791042"}, "user_tz": 240} id="ftpRQ-pF22hk"
label2id = {False: 0, True: 1}
id2label = {v: k for k, v in label2id.items()}
class SentencePairDataset(IterableDataset):
def __init__(self, file_paths, tokenizer, test_mode=False):
self.file_paths = file_paths
self.tokenizer = tokenizer
self.test_mode = test_mode
self._setup()
def _setup(self):
dataset_length = 0
for file_path in tqdm(self.file_paths):
url_tweets, n_url_tweets = zip(*read_file(file_path))
n_url_tweets = np.array(n_url_tweets)
n_pairs = (
n_url_tweets * (n_url_tweets - 1)
).sum() # Don't divide by 2 as for each positive there is a negative
dataset_length += n_pairs
self.dataset_length = dataset_length
def prepare_tokenized_examples(self, batch_size=100000):
chunk_pairs = []
chunk_labels = []
def _process_chunk(chunk_pairs, chunk_labels):
encodings = self.tokenizer(chunk_pairs, max_length=512, truncation=True)
for i, label in enumerate(chunk_labels):
item = {key: torch.tensor(val[i]) for key, val in encodings.items()}
item["label"] = torch.tensor(label)
yield item
if self.test_mode:
np.random.seed(1337)
for i, (pair, label) in enumerate(read_sentencepair_data(self.file_paths)):
if self.test_mode:
if i > 10000:
break
if len(chunk_pairs) > batch_size:
yield from _process_chunk(chunk_pairs, chunk_labels)
chunk_labels = []
chunk_pairs = []
chunk_pairs.extend(pair)
chunk_labels.extend(label)
if chunk_pairs:
yield from _process_chunk(chunk_pairs, chunk_labels)
def __iter__(self):
for item in self.prepare_tokenized_examples(batch_size=100000):
yield item
def __len__(self):
return self.dataset_length
def compute_metrics(p: EvalPrediction):
preds = p.predictions[0] if isinstance(p.predictions, tuple) else p.predictions
preds = np.argmax(preds, axis=1)
result = classification_report(p.label_ids, preds)
return result
# + [markdown] id="_ezRYNW-3AQL"
# # Setup Datasets
#
# + colab={"base_uri": "https://localhost:8080/", "height": 149, "referenced_widgets": ["e985737fb27949fe9f604f043f9815b2", "82e101337c10435e930a3b67b69d5f42", "8e78275792f74ff088754297a204cae6", "467f4b4ed30545c8a19b46638750bd0e", "f38f180edce94cdd920494cce5fe9d18", "f644d155131a4ed39226dd4150fb9507", "7610f5ff055743c2b2013c17f847c587", "96cfc5b54dfb4ab68fcb06ea754b9ba0", "49e3ff206b2d44aba9326e9f01e83185", "5828c23fe29d4da8879d1b9051c6de88", "ede8bd135caa493f9ccafa88a670d711", "b0a0f96d891a4dcd85b0daf451f50966", "abfb0af9a84948fabd95612a52729bb6", "<KEY>", "c08ef40fd1ec4813a4f579ae8c9eb6e6", "e50b1f6ad7cd4b0cb007edf5e9613aeb"]} executionInfo={"elapsed": 1347, "status": "ok", "timestamp": 1631941390642, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GijxQCX_py83nArX6F4S58845Caf6w8Apxd0EpS=s64", "userId": "02687645879366791042"}, "user_tz": 240} id="XQe6JZKn25q_" outputId="9fac863f-df63-4168-9bd1-fc4578f307e7"
file_paths = list(Path(DATADIR).expanduser().glob(f"./{WIKIDATA_PREFIX}*.json"))
model_dir = pre_trained_model_path
tokenizer = BertTokenizerFast.from_pretrained(
str(model_dir), max_len=512, truncation=True, padding=True
)
if WIKIDATA_MODEL:
train_file_paths = file_paths
val_file_paths = []
test_file_paths = []
else:
train_file_paths = file_paths[:-2]
val_file_paths = file_paths[-2:-1] # Single file
test_file_paths = file_paths[-1:] # Single file
# langs = {"en"}
# Set test_mode to false to run the model on full data
test_mode = False
train_dataset = SentencePairDataset(train_file_paths, tokenizer, test_mode=test_mode)
val_dataset = SentencePairDataset(val_file_paths, tokenizer, test_mode=test_mode)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 6, "status": "ok", "timestamp": 1631941390642, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GijxQCX_py83nArX6F4S58845Caf6w8Apxd0EpS=s64", "userId": "02687645879366791042"}, "user_tz": 240} id="SiuSyyEP3KxK" outputId="e0543a6d-b2aa-4f99-9e13-caf03e2bea78"
data = []
for i, (pair, label) in enumerate(read_sentencepair_data(file_paths[:1])):
print(i, label, pair)
data.append(pair)
if i > 10:
break
print("***" * 10)
for i, pair in enumerate(data):
print(i, pair)
# + [markdown] id="RBe6yiMPOB6h"
# # Setup Model
#
# + executionInfo={"elapsed": 5, "status": "ok", "timestamp": 1631941390643, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GijxQCX_py83nArX6F4S58845Caf6w8Apxd0EpS=s64", "userId": "02687645879366791042"}, "user_tz": 240} id="Y6Z9C7qm3M4i"
loss_type = "bce"
loss_margin = 0 # 0, 0.5, -0.5, -1
model_prefix = f"{loss_type}_{loss_margin}" if loss_type == "cosine" else f"{loss_type}"
extra_model_prefix = "from_tt_wm" # "from_tt_wd" # None #"from_tt" # "from_wm"
if extra_model_prefix:
model_prefix = f"{model_prefix}_{extra_model_prefix}"
multi_type = "wiki" # "curated" # "wikimat" # "wiki" # "tatoeba"
model_prefix = f"{WIKIDATA_PREFIX}_{model_prefix}"
finetuned_model_dir = str(
Path(f"{HOMEDIR}/multi_{multi_type}_2t_{model_prefix}_model").expanduser()
)
logging_dir = str(
Path(f"{HOMEDIR}/multi_{multi_type}_2t_{model_prefix}_logs").expanduser()
)
data_collator = DataCollatorWithPadding(tokenizer, padding=True, max_length=514)
num_epochs = 1
if test_mode:
num_epochs = 1000
elif WIKIDATA_MODEL and WIKIDATA_PREFIX.startswith("en_"):
num_epochs = 3
training_args = TrainingArguments(
output_dir=str(finetuned_model_dir), # output directory
num_train_epochs=num_epochs, # total number of training epochs
per_device_train_batch_size=32, # batch size per device during training # Ensure this is multiple of 4
warmup_steps=500, # number of warmup steps for learning rate scheduler
logging_dir=str(logging_dir), # directory for storing logs
logging_steps=10,
save_total_limit=2,
prediction_loss_only=True,
learning_rate=1e-6,
label_names=[id2label[i] for i in range(len(id2label))],
)
# + executionInfo={"elapsed": 4, "status": "ok", "timestamp": 1631941390643, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GijxQCX_py83nArX6F4S58845Caf6w8Apxd0EpS=s64", "userId": "02687645879366791042"}, "user_tz": 240} id="c0S6OU6cOSAU"
def train_model():
model = BertForTwoTowerPrediction.from_pretrained(
str(model_dir),
num_labels=len(label2id),
id2label=id2label,
label2id=label2id,
loss_type=loss_type,
margin=loss_margin,
)
# Patch model token_type_embedding as it is of wrong size during pre-training
# model.config.type_vocab_size += 1 # Increase it for next sentence prediction task
# token_type_embeddings = torch.nn.Embedding(model.config.type_vocab_size, model.config.hidden_size)
# token_type_embeddings.weight[0] = model.bert.embeddings.token_type_embeddings.weight[0]
# model.bert.embeddings.token_type_embeddings = token_type_embeddings # Re-assign new embedding
# Ready to train
trainer = Trainer(
model=model, # the instantiated 🤗 Transformers model to be trained
args=training_args, # training arguments, defined above
train_dataset=train_dataset, # training dataset
tokenizer=tokenizer,
data_collator=data_collator,
compute_metrics=compute_metrics,
)
trainer.train()
trainer.save_model(finetuned_model_dir)
tokenizer.save_pretrained(finetuned_model_dir)
return model, trainer
# + colab={"base_uri": "https://localhost:8080/", "height": 312, "referenced_widgets": ["5a316dd79d2c48c09ea58bc38a9c1f2f", "d70494a82cac4233a1f8ce7a07e56693", "54c0aa205fe543f8b85ad95a05269e48", "91664779084249f7bc45ee320723c5a1", "715d0a59bc814029a35a28f8279ad356", "<KEY>", "2627279826ec481e97fcc3cb5e44b8b1", "<KEY>", "<KEY>", "17165c85e134426a952429b45c0c155a", "f7c801c4ee1f4325aa721f0c940b72a1", "22e0f1f2f8c0450a86b78ac1cfa877c3", "2425521a0f7c4d8d8b754bfde86eaf11", "b374658337494c74bdf857697c4fb2f1", "<KEY>", "11287abb454b476e90ee111813e12e23"]} executionInfo={"elapsed": 35174, "status": "ok", "timestamp": 1631941425813, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GijxQCX_py83nArX6F4S58845Caf6w8Apxd0EpS=s64", "userId": "02687645879366791042"}, "user_tz": 240} id="xusuhI5OOVMY" outputId="016d9b9e-caf0-4765-d116-3b36c51b011a"
# %%time
model, trainer = train_model()
# + executionInfo={"elapsed": 5, "status": "ok", "timestamp": 1631941425813, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GijxQCX_py83nArX6F4S58845Caf6w8Apxd0EpS=s64", "userId": "02687645879366791042"}, "user_tz": 240} id="-cG2lin5OXMu"
# # ! rm -rf multi_wiki_2t_en_ja_bce_from_tt_wm_*
# + executionInfo={"elapsed": 5, "status": "ok", "timestamp": 1631941425814, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GijxQCX_py83nArX6F4S58845Caf6w8Apxd0EpS=s64", "userId": "02687645879366791042"}, "user_tz": 240} id="SNWmxGTWOwmT"
| notebooks/Pretrain_TranslationPairPrediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import re
def _parse_line(line):
"""
Do a regex search against all defined regexes and
return the key and match result of the first matching regex
"""
for key, rx in rx_dict.items():
match = rx.search(line)
if match:
return key, match
# if there are no matches
return None, None
def parse_file(filepath, name):
"""
Parse text at given filepath
Parameters
----------
filepath : str
Filepath for file_object to be parsed
Returns
-------
data : pd.DataFrame
Parsed data
"""
data = [] # create an empty list to collect the data
# open the file and read through it line by line
count = 0
count_or = 0
rnd='no round'
with open(filepath, 'r') as file_object:
line = file_object.readline()
while line:
# at each line check for a match with a regex
key, match = _parse_line(line)
# extract round
if key == 'round':
count += 1
rnd_raw = match.group(1)
opr = re.compile(r'Operating Round ')
if opr.match(rnd_raw):
count_or+=1
rnd = opr.sub('OR', rnd_raw)
item = [count,count_or,name,None,None,None, rnd]
data.append(item)
# extract train buying
if key == 'buytrain':
count += 1
company = match.group('company')
train = match.group('train')
source = match.group('source')
item = [count,count_or,name,company,train,source, rnd]
data.append(item)
line = file_object.readline()
df = pd.DataFrame(data, columns = ['Count', 'Count_OR', 'Game', 'Company', 'Train', 'Source', 'Round'])
return df
rx_dict = {
'round': re.compile(r'== Start of (.*) =='),
'buytrain': re.compile(r'(?P<company>.*) buys a (?P<train>.*-train) from (?P<source>.*) for'),
'name_score': re.compile(r'(?P<name_score>Name|Score)'),
}
rx_dict
# +
import os
logdir="logfiles"
directory = os.fsencode(logdir)
dflist=[]
for file in os.listdir(directory):
filename = os.fsdecode(file)
filepath = os.path.join(logdir, filename)
name = os.path.splitext(filename)[0]
dff = parse_file(filepath, name)
dflist.append(dff)
df=pd.concat(dflist)
# -
df_ipo = df[df['Source']=='IPO']
df
import matplotlib.pyplot as plt
# +
for game in df_ipo['Game'].unique():
opr=re.compile(r'OR')
select_all = df[df['Game']==game]
unique=select_all.Round.unique()
select_or=[el for el in unique if opr.match(el)]
fig, ax = plt.subplots()
select_ipo = df_ipo[df_ipo['Game']==game]
x=select_ipo['Count_OR']
y=select_ipo['Train']
plt.title(game)
ind = np.arange(len(select_or))
plt.xticks(ind, select_or)
plt.scatter(x,y)
# -
select1 = df[df['Game']=="R01"]
select1.Round.unique()
opr=re.compile(r'OR')
sel1=select1.Round.unique()
selectedlist=[el for el in sel1 if opr.match(el)]
selectedlist
len(selectedlist)
df_ipo.groupby(['Game', 'Train']).last()
game='R09'
if True:
opr=re.compile(r'OR')
select_all = df[df['Game']==game]
unique=select_all.Round.unique()
select_or=[el for el in unique if opr.match(el)]
fig, ax = plt.subplots()
select_ipo = df_ipo[df_ipo['Game']==game]
x=select_ipo['Count_OR']
y=select_ipo['Train']
plt.title(game)
ind = np.arange(len(select_or))
plt.xticks(ind, select_or)
plt.scatter(x,y)
select_or
| .ipynb_checkpoints/parse-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: python-sentdex-ml-tiD-lXqz
# language: python
# name: python-sentdex-ml-tid-lxqz
# ---
import pandas as pd
import numpy as np
college = pd.read_csv('data/college.csv')
columns = college.columns
columns
columns.values
columns[5]
columns[[1,8,10]]
columns[-7:-4]
columns.min(), columns.max(), columns.isnull().sum()
columns + '_A'
columns > 'G'
# +
# columns[1] = 'city' # doesn't work because indexes are immutable
# -
c1 = columns[:4]
c1
c2 = columns[2:6]
c2
c1.union(c2) # or c1 | c2
c1.symmetric_difference(c2) # or c1 ^ c2
s1 = pd.Series(index=list('aaab'), data=np.arange(4))
s1
s2 = pd.Series(index=list('cababb'), data=np.arange(6))
s2
s1 + s2
s1 = pd.Series(index=list('aaabb'), data=np.arange(5))
s2 = pd.Series(index=list('aaabb'), data=np.arange(5))
s1, s2
s1 + s2
s1 = pd.Series(index=list('aaabb'), data=np.arange(5))
s2 = pd.Series(index=list('bbaaa'), data=np.arange(5))
s1, s2
s1+s2
employee = pd.read_csv('data/employee.csv', index_col='RACE')
employee.head()
salary1 = employee['BASE_SALARY']
salary2 = employee['BASE_SALARY']
salary1 is salary2
salary1 = employee['BASE_SALARY'].copy()
salary2 = employee['BASE_SALARY'].copy()
salary1 is salary2
salary1 = salary1.sort_index()
salary1.head()
salary2.head()
salary_add = salary1 + salary2
salary_add.head()
len(salary_add)
salary_add1 = salary1 + salary1
len(salary1), len(salary2), len(salary_add), len(salary_add1)
index_vc = salary1.index.value_counts(dropna=False)
index_vc
index_vc.pow(2).sum()
baseball_14 = pd.read_csv('data/baseball14.csv', index_col='playerID')
baseball_15 = pd.read_csv('data/baseball15.csv', index_col='playerID')
baseball_16 = pd.read_csv('data/baseball16.csv', index_col='playerID')
baseball_14.head()
baseball_14.index.difference(baseball_15.index)
baseball_14.index.difference(baseball_16.index)
hits_14 = baseball_14['H']
hits_15 = baseball_15['H']
hits_16 = baseball_16['H']
hits_14.head()
(hits_14 + hits_15).head()
hits_14.add(hits_15, fill_value=0).head()
hits_total = hits_14.add(hits_15, fill_value=0).add(hits_16, fill_value=0)
hits_total.head()
hits_total.hasnans
df_14 = baseball_14[['G', 'AB', 'R', 'H']]
df_14.head()
df_15 = baseball_15[['AB', 'R', 'H', 'HR']]
df_15.head()
(df_14 + df_15).head(10).style.highlight_null('yellow')
df_14.add(df_15, fill_value=0).head(10).style.highlight_null('yellow')
employee = pd.read_csv('data/employee.csv')
dept_sal = employee[['DEPARTMENT', 'BASE_SALARY']]
dept_sal = dept_sal.sort_values(['DEPARTMENT', 'BASE_SALARY'], ascending=[True, False])
max_dept_sal = dept_sal.drop_duplicates(subset='DEPARTMENT')
max_dept_sal.head()
max_dept_sal = max_dept_sal.set_index('DEPARTMENT')
employee = employee.set_index('DEPARTMENT')
employee['MAX_DEPT_SALARY'] = max_dept_sal['BASE_SALARY']
employee.head()
employee.query('BASE_SALARY > MAX_DEPT_SALARY')
employee['MAX_SALARY2'] = max_dept_sal['BASE_SALARY'].head(3)
employee.MAX_SALARY2.value_counts()
employee.MAX_SALARY2.isnull().mean()
college = pd.read_csv('data/college.csv', index_col='INSTNM')
college.dtypes
college.MD_EARN_WNE_P10.iloc[0]
college.GRAD_DEBT_MDN_SUPP.iloc[0]
college.MD_EARN_WNE_P10.sort_values(ascending=False).head()
college.GRAD_DEBT_MDN_SUPP.sort_values(ascending=False).head()
cols = ['MD_EARN_WNE_P10', 'GRAD_DEBT_MDN_SUPP']
for col in cols:
college[col] = pd.to_numeric(college[col], errors='coerce')
college.dtypes.loc[cols]
college_n = college.select_dtypes(include=[np.number])
college_n.head()
criteria = college_n.nunique() == 2
criteria.head()
binary_cols = college_n.columns[criteria].tolist()
binary_cols
college_n2 = college_n.drop(labels=binary_cols, axis='columns')
college_n2.head()
max_cols = college_n2.idxmax()
max_cols
unique_max_cols = max_cols.unique()
unique_max_cols[:5]
college_n2.loc[unique_max_cols].style.highlight_max()
college = pd.read_csv('data/college.csv', index_col='INSTNM')
college_ugds = college.filter(like='UGDS_').head(5)
college_ugds
college_ugds.style.highlight_max(axis='columns')
college = pd.read_csv('data/college.csv', index_col='INSTNM')
cols = ['MD_EARN_WNE_P10', 'GRAD_DEBT_MDN_SUPP']
for col in cols:
college[col] = pd.to_numeric(college[col], errors='coerce')
college_n = college.select_dtypes(include=[np.number])
criteria = college_n.nunique() == 2
binary_cols = college_n.columns[criteria].tolist()
college_n = college_n.drop(labels=binary_cols, axis='columns')
college_n.max().head()
college_n.eq(college_n.max()).head()
has_row_max = college_n.eq(college_n.max()).any(axis='columns')
has_row_max.head()
college_n.shape
has_row_max.sum()
college_n.eq(college_n.max()).cumsum()
college_n.eq(college_n.max()).cumsum().cumsum()
has_row_max2 = college_n.eq(college_n.max()).cumsum().cumsum().eq(1).any(axis='columns')
has_row_max2.head()
has_row_max2.sum()
idxmax_cols = has_row_max2[has_row_max2].index
idxmax_cols
set(college_n.idxmax().unique()) == set(idxmax_cols)
# %timeit college_n.idxmax().values
# %timeit college_n.eq(college_n.max()) \
# .cumsum() \
# .cumsum() \
# .eq(1) \
# .any(axis='columns') \
# [lambda x: x].index
college = pd.read_csv('data/college.csv', index_col='INSTNM')
college_ugds = college.filter(like='UGDS_')
college_ugds.head()
highest_percentage_race = college_ugds.idxmax(axis='columns')
highest_percentage_race.head()
highest_percentage_race.value_counts(normalize=True)
college_black = college_ugds[highest_percentage_race == 'UGDS_BLACK']
college_black = college_black.drop('UGDS_BLACK', axis='columns')
college_black.idxmax(axis='columns').value_counts(normalize=True)
| Index alignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import networkx as nx
import pandas as pd
import numpy as np
import random
import re
import os
data_file=os.listdir('./CG_graph')
for filename in data_file:
if filename.endswith('.txt'):
print(filename)
data=pd.read_csv('./final_file/'+filename,sep='\t',header=None)
n=len(data)
g = nx.Graph()
for (u,v) in data.values:
g.add_edge(u,v)
degree_dict=dict(g.degree)
degree=list(set(degree_dict.values()))
degree.sort()
print('the length of degree is {}'.format(len(degree)))
degree_size=int(len(degree)/16)
degree_one=degree[degree_size*2]
degree_two=degree[degree_size*4]
degree_three=degree[degree_size*6]
degree_four=degree[degree_size*8]
node_low=[k for k,v in degree_dict.items() if v < degree_one]
node_middle=[k for k,v in degree_dict.items() if v >= degree_two and v < degree_three]
node_high=[k for k,v in degree_dict.items() if v >= degree_four]
print(len(node_low),len(node_middle),len(node_high))
print(n)
for batch_size in [500,1000,2000,5000]:
print(batch_size)
for way in ['add','minus']:
if way=='add':
print(way)
situation_dic={'low':node_low,'middle':node_middle,'high':node_high}
for degree_situation in ['low','middle','high']:
node_=situation_dic[degree_situation]
node_choise = list(np.random.choice(node_, size=batch_size, replace=True))
data_final=pd.DataFrame()
sign=['+']*(n+batch_size)
in_=list(data[0].values)
out_=list(data[1].values)
for u in node_choise:
v=random.sample(g.nodes(),1)[0]
while g.has_edge(u,v) and u==v:
v=random.sample(g.nodes(),1)[0]
g.add_edge(u,v)
in_.append(u)
out_.append(v)
data_final['sign']=sign
data_final['in']=in_
data_final['out']=out_
name=filename.strip('.txt')
data_final.to_csv('./degree/add/'+name+'_'+'batchsize_'+str(batch_size)+'_'+degree_situation+'_add_degree.txt',sep='\t',index=None,header=None)
else:
print(way)
situation_dic={'low':node_low,'middle':node_middle,'high':node_high}
for degree_situation in ['low','middle','high']:
node_=set(situation_dic[degree_situation])
#random.sample(node_,1)[0]
#node_choise=random.sample(node_,batch_size)
i=0
data_final=pd.DataFrame()
sign=['+']*n
sign.extend(['-']*batch_size)
in_=list(data[0].values)
out_=list(data[1].values)
while i<batch_size:
u,v=random.sample(g.edges(),1)[0]
if u in node_ or v in node_:
g.remove_edge(u,v)
in_.append(u)
out_.append(v)
i=i+1
data_final['sign']=sign
data_final['in']=in_
data_final['out']=out_
name=filename.strip('.txt')
data_final.to_csv('./degree/minus/'+name+'_'+'batchsize_'+str(batch_size)+'_'+degree_situation+'_minus_degree.txt',sep='\t',index=None,header=None)
data=pd.read_csv('./PLG_graph/'+file_name,sep='\t',header=None)
# + jupyter={"source_hidden": true}
import os
file=os.listdir('../max_clique/clean')
print(file)
# -
file=['Email-EuAll.txt']
for file_name in file:
if file_name.endswith('.txt'):
data=pd.read_csv(file_name,sep='\t',header=None)
a=['+']*len(data)
a.extend(['-']*len(data))
a=np.array(a)
data_end=pd.DataFrame()
data_end[0]=a
in_=data[0].values
out_=data[1].values
data_end[1]=np.hstack((in_,in_[::-1]))
data_end[2]=np.hstack((out_,out_[::-1]))
name=file_name.strip('.txt')
print(name)
data_end.to_csv('./'+name+'_add_minus.txt',sep='\t',index=None,header=None)
# +
filename='./real_data/Email-EuAll.txt'
G = nx.DiGraph()
i=0
with open(filename) as file:
for line in file:
if i<4:
i+=1
continue
head, tail= [int(x) for x in re.split('[ ,\t]',line)] #如果节点使用数组表示的可以将str(x)改为int(x)
if head!=tail:
if head>tail:
head,tail=tail,head
G.add_edge(head,tail)
DVweight = G.degree()
max_degree = max(span for n, span in DVweight) #节点最大度数
ver_num=len(G.nodes())
edge_num=len(G.edges())
print(filename,ver_num,edge_num,(2*edge_num)/(ver_num*ver_num),max_degree)
data_list=[[edge[0],edge[1]] for edge in G.edges()]
data=pd.DataFrame(data=data_list,columns=['in','out'])
print(len(data))
name=filename.split('/')[-1]
data.to_csv('./'+name,sep='\t',index=None,header=None)
# -
filename='Email-EuAll.txt'
data=pd.read_csv(filename,sep='\t',header=None)
data=data.sort_values(axis=0, ascending=True, by=1).reset_index(drop=True)
print(data.head(10))
data.to_csv('./Email-EuAll.txt',sep='\t',index=None,header=None)
| DATA/generate data corresponding with degree.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
print('Siapa namamu?')
name = input()
print('Menarik. Namaku juga', name)
print('Pernahkah kamu memikirkan tentang lubang hitam hari ini?')
yes_no = input()
print('Saya sangat senang kamu berkata', yes_no, '. Aku berpikir hal yang sama.')
print('Kita adalah roh yang sama,', name, '. Nanti ngobrol lagi.')
print('Seberapa cerdas kamu? 0 sangat bodoh. Dan 10 adalah jenius')
smarts = input()
smarts = int(smarts)
if smarts <= 3:
print('Aku gak percaya kamu.')
print('Seberapa buruk hari yang kamu alami? 0 adalah yang terburuk, dan 10 adalah yang terbaik.')
day = input()
day = int(day)
if day <= 5:
print('Jika saya manusia, saya akan memelukmu.')
else:
print('Mungkin aku harus mencoba mendekatimu.')
elif smarts <= 6:
print('Saya pikir kamu sebenarnya lebih pintar.')
print('Berapa banyak waktu yang kamu habiskan untuk online? 0 tidak ada dan 10 adalah 24 jam sehari.')
hours = input()
hours = int(hours)
if hours <= 4:
print('Wah gawat bahaya.')
else:
print('Kupikir cuma aku.')
elif smarts <= 8:
print('Apakah kamu manusia betulan? Tunggu. Jangan jawab itu.')
print('Seberapa manusia anda? 0 tidak sama sekali dan 10 adalah manusia sepenuhnya.')
human = input()
human = int(human)
if human <= 5:
print('Aku tahu itu.')
else:
print('Aku pikir hubungan ini sudah berakhir.')
else:
print('Begitu ... Berapa banyak sistem operasi yang kamu jalankan?')
os = input()
os = int(os)
if os <= 2:
print('Untung anda mengikuti pelajaran ini.')
else:
print('Apa ini? Sebuah kompetisi?')
| Sesi-1-matematika-string-kondisi-perulangan/code/Kegiatan5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:torch]
# language: python
# name: conda-env-torch-py
# ---
import sys
[sys.path.append(i) for i in ['.', '..']]
import torch
import numpy as np
# import sigpy as sp
# from sigpy.mri.app import L1WaveletRecon
from prox import Prox
from alg.eig import power_iter
from alg.cg import CG
from linear import LinearMap, FFTCn, NuSense, Sense, FFTCn, Identity, Diff2d
import matplotlib.pyplot as plt
import copy
import h5py
import torchkbnufft as tkbn
# FFT example
device0 = torch.device('cuda:0')
from skimage.data import shepp_logan_phantom
I_shepp = torch.tensor(shepp_logan_phantom()).to(device0)
Fop = FFTCn((400, 400), (400, 400), (0,1), norm = 'ortho')
k_shepp = Fop*I_shepp
plt.imshow(torch.log(torch.abs(k_shepp)).cpu().data.numpy())
plt.colorbar()
# MRI parallel imaging example (SENSE)
ex_multi = np.load('AXT2_210_6001736_layer11.npz')
device0 = torch.device('cpu')
k_c = torch.tensor(ex_multi['k_r'] + 1j*ex_multi['k_i']).to(device0)/32767.0
s_c = torch.tensor(ex_multi['s_r'] + 1j*ex_multi['s_i']).to(device0)/32767.0
(nc, nh, nw) = s_c.shape
Fop = FFTCn((nc, nh, nw), (nc, nh, nw), (1,2), norm = 'ortho')
I1 = Fop.H*k_c
I1 = torch.sqrt(I1.abs().pow(2).sum(dim=0))
mask = torch.zeros(nh,nw)
mask[:,0:nw:8]=1
Sop = Sense((nh, nw), (nc, nh, nw), (0,1), s_c, mask, batchmode = False)
I0 = Sop.H*k_c
plt.figure(figsize=(20,10))
plt.imshow(torch.abs(I0).data.numpy())
plt.colorbar()
T = Diff2d(Sop.size_in, dim = (0,1))
CG_tik = CG(Sop.H*Sop+0.02*T.H*T, max_iter = 20)
I = Identity(Sop.size_in, Sop.size_in)
CG_tik = CG(Sop.H*Sop+0.02*I, max_iter = 20)
I0.requires_grad_()
I_tik = CG_tik.run(I0, I0)
torch.sum(torch.pow(torch.abs(I_tik),2)).backward()
I0.grad
plt.figure(figsize=(20,10))
plt.imshow(torch.abs(I0.grad).cpu().data.numpy())
plt.colorbar()
plt.figure(figsize=(20,10))
plt.imshow(torch.abs(I_tik).cpu().data.numpy())
plt.colorbar()
hf = h5py.File('./b0.h5', 'r')
nx = 320;
ny = 320;
ktraj = hf['ktraj'][()]
ktraj = np.remainder(ktraj + np.pi, 2*np.pi)-np.pi
# ktraj = np.transpose(ktraj)/np.pi*nx/2
print('traj shape', ktraj.shape)
k = hf['k_r'][()] + 1j*hf['k_i'][()]
[ncoil, nslice, nshot, ns] = k.shape
print('k shape', k.shape)
dcf = hf['dcf'][()]
print('dcf shape', dcf.shape)
ksp = k.reshape(ncoil,nslice,nshot*ns)
smap = np.transpose(hf['s_r'][()] + 1j*hf['s_i'][()], (3,0,1,2))
smap = np.transpose(smap, (0,1,3,2))
print('smap shape', smap.shape)
fmap = hf['b0'][()]
fmap = np.transpose(fmap, (0,2,1))
print('fmap shape', fmap.shape)
ktrajunder = ktraj.reshape(320,1280,2)
ktrajunder = ktrajunder[0:-1:5,:,:].reshape(81920,2)
kunder = k[:,:,0:-1:5,:]
im_size = (320,320)
iz = 6
k0 = torch.tensor(k[:,iz,:,:]).to(device = 'cuda:0').reshape(1,ncoil,nshot*ns)
s0 = torch.tensor(smap[iz,:,:,:]).to(device = 'cuda:0').unsqueeze(0)
ktraj = torch.tensor(ktraj).to(device = 'cuda:0')
# adjnufft_ob = tkbn.KbNufftAdjoint(im_size=im_size, grid_size=tuple(np.array(im_size)*2)).to(device = 'cuda:0')
Nop = NuSense(s0, ktraj)
# image_blurry = adjnufft_ob(k0, ktraj, smaps=s0)
image_blurry = Nop.H(k0*(torch.tensor(dcf).to(k0).unsqueeze(0)))
plt.figure(figsize=(20,10))
plt.imshow(torch.abs(image_blurry.squeeze(0).squeeze(0)).cpu().data.numpy())
plt.colorbar()
T = Diff2d(Nop.size_in, dim = (1,2))
CG_tik = CG(Nop.H*Nop+0.02*T.H*T, max_iter = 20)
I = Identity(Nop.size_in, Nop.size_in)
CG_tik = CG(Nop.H*Nop+0.2*I, max_iter = 20)
I0 = Nop.H*k0
I_tik = CG_tik.run(I0, I0)
plt.figure(figsize=(20,10))
plt.imshow(torch.abs(I_tik.squeeze(0)).cpu().data.numpy())
plt.colorbar()
| examples/.ipynb_checkpoints/test_linops-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h2>1. Two Sum () solved</h2>
# Easy
#
# Given an array of integers, return indices of the two numbers such that they add up to a specific target.
#
# You may assume that each input would have exactly one solution, and you may not use the same element twice.
#
# Example:
#
# Given nums = [2, 7, 11, 15], target = 9,
#
# Because nums[0] + nums[1] = 2 + 7 = 9,
# return [0, 1].
# +
def twoSum(nums, target):
a = -1
b = -1
indexi = 0
indexj = 0
flag = 0
for i in nums:
# print("here1")
if(a != -1 and b != -1):
break
for j in nums:
# print("here2")
if(i+j == target):
# print("here3")
# print(indexi)
# print(indexj)
if(indexi != indexj):
# print("here4")
a = indexi
b = indexj
break
indexj += 1
indexi += 1
indexj = 0
return [a, b]
twoSum([2, 7, 11, 15],9)
# -
# <h2>7. Reverse Integer (solved)</h2
# >
# Given a 32-bit signed integer, reverse digits of an integer.
#
# Example 1:
#
# Input: 123
# Output: 321
#
# Example 2:
#
# Input: -123
# Output: -321
#
# Example 3:
#
# Input: 120
# Output: 21
#
# Note:
# Assume we are dealing with an environment which could only store integers within the 32-bit signed integer range: [−231, 231 − 1]. For the purpose of this problem, assume that your function returns 0 when the reversed integer overflows.
# +
num = -11210344
def reverse(x):
isnegative = False
if(x<0):
isnegative = True
x = abs(x)
listnum = [int(y) for y in str(x)]
reversednumber=0
i=0
for y in listnum:
reversednumber += y*(10**i)
i+=1
if(isnegative):
reversednumber *= -1
if(reversednumber < -2147483648 or reversednumber > 2147483647):
return(0)
else:
return(reversednumber)
reverse(num)
# -
# <h2>9. Palindrome Number (Solved)</h2>
# Determine whether an integer is a palindrome. An integer is a palindrome when it reads the same backward as forward.
#
# Example 1:
#
# Input: 121
# Output: true
# Example 2:
#
# Input: -121
# Output: false
# Explanation: From left to right, it reads -121. From right to left, it becomes 121-. Therefore it is not a palindrome.
# Example 3:
#
# Input: 10
# Output: false
# Explanation: Reads 01 from right to left. Therefore it is not a palindrome.
# Follow up:
#
# Coud you solve it without converting the integer to a string?
# +
def isPalindrome(x):
string = str(x)
if(string == "".join(reversed(string)) ):
return True
else:
return False
def altisPalindrome(x):
listnum = [int(y) for y in str(x)]
if(listnum == listnum[::-1]):
return True
else:
return False
altisPalindrome(124421)
# -
# <h2>13. Roman to Integer (solved)</h2>
# Roman numerals are represented by seven different symbols: I, V, X, L, C, D and M.
#
# Symbol Value
# I 1
# V 5
# X 10
# L 50
# C 100
# D 500
# M 1000
# For example, two is written as II in Roman numeral, just two one's added together. Twelve is written as, XII, which is simply X + II. The number twenty seven is written as XXVII, which is XX + V + II.
#
# Roman numerals are usually written largest to smallest from left to right. However, the numeral for four is not IIII. Instead, the number four is written as IV. Because the one is before the five we subtract it making four. The same principle applies to the number nine, which is written as IX. There are six instances where subtraction is used:
#
# I can be placed before V (5) and X (10) to make 4 and 9.
# X can be placed before L (50) and C (100) to make 40 and 90.
# C can be placed before D (500) and M (1000) to make 400 and 900.
# Given a roman numeral, convert it to an integer. Input is guaranteed to be within the range from 1 to 3999.
#
# Example 1:
#
# Input: "III"
# Output: 3
# Example 2:
#
# Input: "IV"
# Output: 4
# Example 3:
#
# Input: "IX"
# Output: 9
# Example 4:
#
# Input: "LVIII"
# Output: 58
# Explanation: L = 50, V= 5, III = 3.
# Example 5:
#
# Input: "MCMXCIV"
# Output: 1994
# Explanation: M = 1000, CM = 900, XC = 90 and IV = 4.
# +
def romanToInt(s):
convlist = []
for x in s:
switcher = {
"I": 1,
"V": 5,
"X": 10,
"L": 50,
"C": 100,
"D": 500,
"M": 1000,
}
convlist.append(switcher.get(x, "nothing") )
i = 0
convint = 0
for x in convlist:
if(i < len(convlist) - 1 and x < convlist[i+1]):
convint -= x
else:
convint += x
i += 1
return(convint)
romanToInt('MCMXCIV')
# -
# <h2>2. Add Two Numbers (solved)</h2>
# You are given two non-empty linked lists representing two non-negative integers. The digits are stored in reverse order and each of their nodes contain a single digit. Add the two numbers and return it as a linked list.
#
# You may assume the two numbers do not contain any leading zero, except the number 0 itself.
#
# Example:
#
# Input: (2 -> 4 -> 3) + (5 -> 6 -> 4)
# Output: 7 -> 0 -> 8
# Explanation: 342 + 465 = 807.
# +
# Definition for singly-linked list.
class ListNode(object):
def __init__(self, x):
self.val = x
self.next = None
def addTwoNumbers(l1, l2):
"""
:type l1: ListNode
:type l2: ListNode
:rtype: ListNode
"""
r = (l1.val + l2.val)/10
l1temp = l1
l2temp = l2
result = ListNode((l1.val + l2.val)%10)
resulttemp = result
while(True):
if(l1temp.next == None and l2temp.next == None):
if(r==0):
break
else:
resulttemp.next = ListNode(r)
r = 0
break
elif(l1temp.next == None and l2temp.next != None):
l2temp = l2temp.next
resulttemp.next = ListNode(int((l2temp.val + r) % 10))
if(l2temp.val + r > 9):
r = int((l2temp.val + r) / 10 )
else:
r = 0
resulttemp = resulttemp.next
elif(l1temp.next != None and l2temp.next == None):
l1temp = l1temp.next
resulttemp.next = ListNode(int((l1temp.val + r) % 10))
if(l1temp.val + r > 9):
r = int((l1temp.val + r) / 10 )
else:
r = 0
resulttemp = resulttemp.next
else:
l1temp = l1temp.next
l2temp = l2temp.next
resulttemp.next = ListNode(int((l1temp.val + l2temp.val + r) % 10) )
if(l1temp.val + l2temp.val + r > 9):
r = int(int((l1temp.val + l2temp.val + r) / 10))
else:
r = 0
resulttemp = resulttemp.next
return (result)
# test :
l1 = ListNode(2)
l1head = l1
l1.next = ListNode(4)
l1 = l1.next
l1.next = ListNode(3)
l2 = ListNode(5)
l2head = l2
l2.next = ListNode(6)
l2 = l2.next
l2.next = ListNode(4)
result = addTwoNumbers(l1head,l2head)
print(result.next.next.val)
# -
# ## 23. Merge k Sorted Lists (solved)
#
# hard
#
# Merge k sorted linked lists and return it as one sorted list. Analyze and describe its complexity.
#
# Example:
#
# Input:
# [
# 1->4->5,
# 1->3->4,
# 2->6
# ]
# Output: 1->1->2->3->4->4->5->6
# +
# Definition for singly-linked list.
class ListNode(object):
def __init__(self, x):
self.val = x
self.next = None
def mergeKLists(lists):
smnumbers = []
while(True):
snum = float('inf')
sind = 0
i=0
comListCount = 0
for lis in lists:
try:
if(lis.val < snum):
snum = lis.val
sind = i
except:
comListCount += 1
continue
finally:
i+=1
if(comListCount == len(lists)):
break
smnumbers.append(snum)
lists[sind] = lists[sind].next
if(smnumbers != []) :
merglist = ListNode(smnumbers[0])
templist = merglist
for x in smnumbers[1:len(smnumbers)]:
templist.next = ListNode(x)
templist = templist.next
else:
return
return merglist
a = ListNode(1)
a.next = ListNode(4)
a.next.next = ListNode(5)
b = ListNode(1)
b.next = ListNode(3)
b.next.next = ListNode(4)
c = ListNode(2)
c.next = ListNode(6)
res = mergeKLists([a,b,c])
# -
ittorres = res
while(ittorres != None):
print(ittorres.val)
ittorres = ittorres.next
# ## 37. Sudoku Solver (unsolved)
#
# hard
#
# Write a program to solve a Sudoku puzzle by filling the empty cells.
#
# A sudoku solution must satisfy all of the following rules:
#
# Each of the digits 1-9 must occur exactly once in each row.
# Each of the digits 1-9 must occur exactly once in each column.
# Each of the the digits 1-9 must occur exactly once in each of the 9 3x3 sub-boxes of the grid.
# Empty cells are indicated by the character '.'.
#
#
# A sudoku puzzle...
#
# <img src = "https://upload.wikimedia.org/wikipedia/commons/thumb/f/ff/Sudoku-by-L2G-20050714.svg/250px-Sudoku-by-L2G-20050714.svg.png">
#
# ...and its solution numbers marked in red.
#
# <img src = "https://upload.wikimedia.org/wikipedia/commons/thumb/3/31/Sudoku-by-L2G-20050714_solution.svg/250px-Sudoku-by-L2G-20050714_solution.svg.png">
#
# Note:
#
# The given board contain only digits 1-9 and the character '.'.
# You may assume that the given Sudoku puzzle will have a single unique solution.
# The given board size is always 9x9.
# +
def solveSudoku(board):
# 1 - check the possible inputs in a box
# 2 - check the possible inputs in a row
# 3 - check the possible inputs in a column
i = 0
j = 0
elerow = []
for row in board:
nums = [1,2,3,4,5,6,7,8,9]
for i in row:
try:
i = int(i)
nums.remove(i)
except:
pass
elerow.append(nums)
print(elerow)
board1 = [["5","3",".",".","7",".",".",".","."],
["6",".",".","1","9","5",".",".","."],
[".","9","8",".",".",".",".","6","."],
["8",".",".",".","6",".",".",".","3"],
["4",".",".","8",".","3",".",".","1"],
["7",".",".",".","2",".",".",".","6"],
[".","6",".",".",".",".","2","8","."],
[".",".",".","4","1","9",".",".","5"],
[".",".",".",".","8",".",".","7","9"]]
solveSudoku(board1)
# -
# ## 224. Basic Calculator
#
# hard
#
# Implement a basic calculator to evaluate a simple expression string.
#
# The expression string may contain open ( and closing parentheses ), the plus + or minus sign -, non-negative integers and empty spaces .
#
# Example 1:
#
# Input: "1 + 1"
# Output: 2
# Example 2:
#
# Input: " 2-1 + 2 "
# Output: 3
# Example 3:
#
# Input: "(1+(4+5+2)-3)+(6+8)"
# Output: 23
# +
import re
def operation(a,b,opp):
if(opp == '+'):
return(a+b)
elif(opp == '-'):
return(a-b)
elif(opp == '*'):
return(a*b)
elif(opp == '/'):
return(a/b)
else:
return("error in operation")
def calculate(s):
arr=re.split('\W+',s)
arr =[int(i) for i in arr]
opp = re.findall("[-+*/]",s)
print(arr)
print(opp)
i = 0
while( len(opp)>0 ):
if('/' in opp):
ind = opp.index('/')
arr[ind] = operation(arr[ind],arr[ind+1],opp[ind])
del opp[ind]
del arr[ind+1]
elif('*' in opp):
ind = opp.index('*')
arr[ind] = operation(arr[ind],arr[ind+1],opp[ind])
del opp[ind]
del arr[ind+1]
elif('+' in opp):
ind = opp.index('+')
arr[ind] = operation(arr[ind],arr[ind+1],opp[ind])
del opp[ind]
del arr[ind+1]
elif('-' in opp):
ind = opp.index('-')
arr[ind] = operation(arr[ind],arr[ind+1],opp[ind])
del opp[ind]
del arr[ind+1]
print()
return(arr[-1])
str = "9 + 5 / 5 * 50"
print(calculate(str))
# -
# ## Google online coding interview questions:
# #### 1)
# One string is strictly smaller than another when the frequency of occurrence of the smallest character in the string is less than the frequency of occurrence of the smallest character in the comparison string.
# For example, string "abcd" is smaller than string "aaa" because the smallest character (in lexicographical order) in "abcd" is 'a', with a frequency of 1, and the smallest character in "aaa" is also 'a', but with a frequency of 3. In another example, string "a" is smaller than string "bb" because the smallest character in "a" is 'a' with a frequency of 1, and the smallest character in "bb" is 'b' with a frequency of 2.
# Write a function that, given string A (which contains M strings delimited by ',') and string B (which contains N strings delimited by ','), returns an array C of N integers. For 0 ≤ J < N, values of C[J] specify the number of strings in A which are strictly smaller than the comparison J-th string in B (starting from 0).
# For example, given strings A and B such that:
# A = "abcd,aabc,bd"
# B = "aaa,aa"
# the function should return [3, 2], because:
# All the strings in the array are strictly smaller than "aaa" on the basis of the given comparison criteria;
# Strings "abcd" and "bd" are strictly smaller than "aa".
# Assume that:
# 1 ≤ N, M ≤ 10000
# 1 ≤ length of any string contained by A or B ≤ 10
# All the input strings comprise only lowercase English alphabet letters (a-z)
# In your solution, focus on correctness. The performance of your solution will not be the primary focus of the assessment.
A = "aa,aaaa,abbca"
B = "aaa,bab"
# +
def get_freq(entry):
smallest = ord(entry[0])
freq = 0
for alpha in entry:
if(ord(alpha) == smallest):
freq += 1
elif(ord(alpha) < smallest):
smallest = ord(alpha)
freq = 1
return freq
dictA = {}
for entry in A.split(","):
freq = get_freq(entry)
if(freq in dictA):
dictA[freq] += 1
else:
dictA[freq] = 1
retlist = []
for entry in B.split(","):
freq = get_freq(entry)
cmpres = 0
for freqA in [*dictA]:
if(freqA<freq):
cmpres += dictA[freqA]
retlist.append(cmpres)
retlist
# -
# #### 2)
# Array X is greater than array Y if the first non-matching element in both arrays has a greater value in X than in Y.
# For example, for arrays X and Y such that:
# X = [1, 2, 4, 3, 5]
# Y = [1, 2, 3, 4, 5]
# X is greater than Y because the first element that does not match is larger in X (i.e. for X[2] and Y[2], X[2] > Y[2]).
# A contiguous subarray is defined by an interval of the indices. In other words, a contiguous subarray is a subarray which has consecutive indexes.
# Write a function that, given a zero-indexed array A consisting of N integers and an integer K, returns the largest contiguous subarray of length K from all the contiguous subarrays of length K.
# For example, given array A and K = 4 such that:
# A = [1, 4, 3, 2, 5]
# the function should return [4, 3, 2, 5], because there are two subarrays of size 4:
# [1, 4, 3, 2]
# [4, 3, 2, 5]
# and the largest subarray is [4, 3, 2, 5].
# Assume that:
# 1 ≤ K ≤ N ≤ 100;
# 1 ≤ A[J] ≤ 1000;
# given an array A contains N distinct integers.
# In your solution, focus on correctness. The performance of your solution will not be the primary focus of the assessment.
N = [1, 4, 3, 2, 5]
K = 3
# <i> Solution 1:
# +
def smallerList(a,b):
for i in range(0,len(a)):
if(a[0]<b[0]):
return a
if(b[0]<a[0]):
return b
return a
slices = []
i=0
while (i+K <= len(N)):
slices.append(N[i:i+K])
i+=1
print(slices)
while(len(slices) > 1):
slices.remove(smallerList(slices[0],slices[1]))
slices[0]
# -
# <i> solution 2:
maxp = max( N[0: len(N) - K + 1] )
N[N.index(maxp): N.index(maxp) + K]
| leetcode solved/leetcode questions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Generation of Sample Input Data for Scalability Analysis
# ---
#
# This notebook exemplifies the data assembly to include in the scalability analysis of the `respy` function `_full_solution`.
#
# The first step includes to generate **sample input data** from either of the Keane and Wolpin models. We have simulated both `kw_94_one` and `kw_97_basic` under a chosen `params` pd.Dataframe. Data was extracted for
# periods:
# - `"per1"`
# - `"per8"`
# - `"per18"`
# - `"per28"`
# - `"per38"`
# - `"per48"` (only for `kw_97_basic`)
#
# and saved under an appropriate file format (in our case `x` `@` [`.pickle`, `.npy`]). To minimize the effort during the timing analysis it is recommended to save each model as a separate file. For scalability analyses (timing analyses) of dynamic models it is indispensable to have data for various periods at hand. The reason: computational effort differs for the current period.
#
# To exclude any data impurity we generated sample input data for `kw_97_basic` twice. For each of the sample data we will create a separate file. The following data sets are gernerate as `x` `@` [`.pickle`, `.npy`].
#
# - `kw_94_one_input_params.x`
# - `kw_97_basic_one_input_params.x`
# - `kw_97_basic_two_input_params.x`
#
#
#
# +
from pathlib import Path
import numpy as np
import pickle
# %load_ext nb_black
# +
path_out_raw_data = Path("./resources/raw_input_data")
# Specific path of the sample data (available upon request)
path_in_raw_data = Path("../../development")
PERIODS = [1, 8, 18, 28, 38, 48]
# -
kw_94_one_input_params = {}
for period in PERIODS[:-1]:
filename = Path(f"{path_in_raw_data}/inputs_kw_94_one_per{period}.pickle")
infile = np.load(filename, allow_pickle=True)
label = "per" + str(period)
kw_94_one_input_params[label] = infile
pickle.dump(
kw_94_one_input_params,
open(f"{path_out_raw_data}/kw_94_one_input_params.pickle", "wb"),
)
np.save(
f"{path_out_raw_data}/kw_94_one_input_params",
kw_94_one_input_params,
allow_pickle=True,
)
for num in ["one", "two"]:
kw_97_basic_input_params = {}
for period in PERIODS[:-1]:
filename = Path(
f"{path_in_raw_data}/inputs_kw_97_basic_{num}_per{period}.pickle"
)
infile = np.load(filename, allow_pickle=True)
label = "per" + str(period)
kw_97_basic_input_params[label] = infile
pickle.dump(
kw_97_basic_input_params,
open(
f"{path_out_raw_data}/kw_97_basic_{num}_input_params.pickle",
"wb",
),
)
np.save(
f"{path_out_raw_data}/kw_97_basic_{num}_input_params",
kw_97_basic_input_params,
allow_pickle=True,
)
# +
input_params_pickle = pickle.load(
open(f"{path_out_raw_data}/kw_97_basic_two_input_params.pickle", "rb")
)
input_params_npy = np.load(
f"{path_out_raw_data}/kw_97_basic_two_input_params.npy", allow_pickle=True
).item()
# -
# ### Different data format: Adjustment in the scripts
#
# If, for some reasons, `.npy` is preferred, some lines in the script files have to be changed. Foremost, in `config.py` the `DATA_FORMAT` should be set to `"npy"`.
#
# In `caller_scalability_analysis.py` the following lines
#
# ```python
# input_params = pickle.load(open(INPUT_DATA, "rb"))[PERIOD]
# pickle.dump(input_params, open(PATH_AUXINPUT_PARAMS, "wb"))
# ```
#
# have to be replaced by:
#
# ```python
# input_params = np.load(INPUT_DATA, allow_pickle=True).item()[PERIOD]
# np.save(PATH_AUXINPUT_PARAMS, input_params, allow_pickle=True)
# ```
#
# In `exec_time_scalability.py` the following line
#
# ```python
# input_params = pickle.load(open(PATH_AUXINPUT_PARAMS, "rb"))
# ```
#
# has to be replaced by:
#
# ```python
# input_params = np.load(PATH_AUXINPUT_PARAMS, allow_pickle=True).item()
# ```
#
# The last change includes (if not already done) to change the imports in those modules. Instead `import pickle` we need to `import numpy as np`.
# ## Sample Input Data
# ---
#
# The resulting input data can be accessed via the period keys. In our case, the function `_full_solution` takes the arguments:
# - wages
# - nonpecs
# - continuation_values
# - period_draws_emax_risk
# - optim_paras
kw_94_one_input_params["per38"]
| scalability_analysis/DataTransformation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.5 64-bit (''base'': conda)'
# name: python3
# ---
# A sanity check for the implementation of MADE.
import optax
from jaxrl.networks.policies import NormalTanhMixturePolicy
from jaxrl.networks.autoregressive_policy import MADETanhMixturePolicy
import matplotlib.pyplot as plt
import jax
import numpy as np
import jax.numpy as jnp
import matplotlib
# %matplotlib inline
@jax.jit
def sample(rng, inputs, std=0.1):
num_points = len(inputs)
rng, key = jax.random.split(rng)
n = jnp.sqrt(jax.random.uniform(key, shape=(num_points // 2,))
) * 540 * (2 * np.pi) / 360
rng, key = jax.random.split(rng)
d1x = -jnp.cos(n) * n + jax.random.uniform(key,
shape=(num_points // 2,)) * 0.5
rng, key = jax.random.split(rng)
d1y = jnp.sin(n) * n + jax.random.uniform(key,
shape=(num_points // 2,)) * 0.5
x = jnp.concatenate(
[
jnp.stack([d1x, d1y], axis=-1),
jnp.stack([-d1x, -d1y], axis=-1)
]
)
rng, key = jax.random.split(rng)
x = x / 3 + jax.random.normal(key, x.shape) * std
return jnp.clip(x / 5 + inputs, -0.9999, 0.9999)
tmp = sample(jax.random.PRNGKey(1), jnp.zeros((10024, 2)))
x = plt.hist2d(tmp[:, 0], tmp[:, 1], bins=128)
# +
rng = jax.random.PRNGKey(1)
made = NormalTanhMixturePolicy((128, 128), 2) # Fails
made = MADETanhMixturePolicy((128, 128), 2) # Works
rng, key = jax.random.split(rng)
params = made.init(key, jnp.zeros(2))['params']
optim = optax.adamw(3e-4)
optim_state = optim.init(params)
# -
@jax.jit
def train_step(rng, params, optim_state):
rng, key1, key2 = jax.random.split(rng, 3)
xs = jax.random.normal(key1, shape=(1024, 2)) * 0.1
ys = sample(key2, xs)
def loss_fn(params):
dist = made.apply({'params': params}, xs)
log_probs = dist.log_prob(ys)
return -log_probs.mean()
loss_fn(params)
value, grads = jax.value_and_grad(loss_fn)(params)
updates, new_optim_state = optim.update(grads, optim_state, params)
new_params = optax.apply_updates(params, updates)
return value, rng, new_params, new_optim_state
for i in range(100000):
value, rng, params, optim_state = train_step(rng, params, optim_state)
if i % 10000 == 0:
print(value)
@jax.jit
def get_log_probs(xs, ys, params):
dist = made.apply({'params': params}, xs)
return dist.log_prob(ys)
# +
x = jnp.linspace(-0.9, 0.9, 256)
y = jnp.linspace(-0.9, 0.9, 256)
xv, yv = jnp.meshgrid(x, y)
ys = jnp.stack([xv, yv], -1)
xs = jnp.zeros_like(ys) - 0.2
log_probs = get_log_probs(xs, ys, params)
plt.imshow(jnp.exp(log_probs))
# -
| notebooks/test_made.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Homework 3: Semantic Retrieval, Text Clustering, and IR Evaluation
#
# In the following are my results for the 3rd Homework Assignement for Information Retrieval and Web Search.
# The notebook is divided into 4 parts:
# 0. Utility Functions
# 1. Latent Semantic Indexing
# 2. Text Clustering
# 3. IR Evaluation
# 4. Semantic Retrieval and Word Embeddings
from pprint import pprint
import numpy as np
from nltk.tokenize import RegexpTokenizer
import nltk
import math
from sklearn.cluster import KMeans
# # 0. Utilility Functions
# +
def preprocess_query(query):
"""Preprocessing of the corpus, filter for nouns and adjectives and lemmatize"""
# stop = set(stopwords.words('english'))
tags = {'NN', 'NNS', 'NNP', 'NNP', 'NNPS', 'JJ', 'JJR', 'JJS'}
wordnet_lemmatizer = WordNetLemmatizer()
# for i in range(len(query)):
query = [(word.lower(), convert(tag)) for (word, tag) in nltk.pos_tag(nltk.word_tokenize(query)) if tag in tags]
query = [wordnet_lemmatizer.lemmatize(w, t) for (w, t) in query ]
return query
def preprocess(docs):
"""Preprocessing of the corpus, filter for nouns and adjectives and lemmatize"""
# stop = set(stopwords.words('english'))
tags = {'NN', 'NNS', 'NNP', 'NNP', 'NNPS', 'JJ', 'JJR', 'JJS'}
for i in range(len(docs)):
docs[i] = [(word.lower(), convert(tag)) for (word, tag) in nltk.pos_tag(nltk.word_tokenize(docs[i])) if tag in tags]
return lemmatize_docs(docs)
def lemmatize_docs(docs):
"""Lemmatize the terms of the corpus"""
wordnet_lemmatizer = WordNetLemmatizer()
for i in range(len(docs)):
docs[i] = [wordnet_lemmatizer.lemmatize(w, t) for (w, t) in docs[i]]
return docs
def convert(tag):
"""Convert tag from treebank to wordnet format"""
if is_noun(tag):
return wn.NOUN
if is_adjective(tag):
return wn.ADJ
# -
# # 1. Latent Semantic Indexing
# +
d1 = """Frodo and Sam were trembling in the darkness, surrounded in darkness by hundreds of blood-
thirsty orcs. Sam was certain these beasts were about to taste the scent of their flesh."""
d2 = """The faceless black beast then stabbed Frodo. He felt like every nerve in his body was hurting.
Suddenly, he thought of Sam and his calming smile. Frodo had betrayed him."""
d3 = """Frodo’s sword was radiating blue, stronger and stronger every second. Orcs were getting
closer. And these weren’t just regular orcs either, Uruk-Hai were among them. Frodo had
killed regular orcs before, but he had never stabbed an Uruk-Hai, not with the blue stick."""
d4 = """Sam was carrying a small lamp, shedding some blue light. He was afraid that orcs might
spot him, but it was the only way to avoid deadly pitfalls of Mordor."""
docs = [d1, d2, d3, d4]
docs = preprocess(docs)
# -
def preprocess(docs):
tokenizer = RegexpTokenizer(r'\w+')
for i in range(len(docs)):
docs[i] = [word.lower() for word in tokenizer.tokenize(docs[i])]
return docs
# #### a)
# Your vocabulary consists of the following terms: Frodo, Sam, beast, orc, and blue.
# Compute the TF-IDF term-document occurrence matrix for given document collection and
# vocabulary terms.
terms = ["Frodo", "Sam", "beast", "blue"]
terms = [t.lower() for t in terms]
idf = calcIDF(terms, docs)
pprint(idf)
tf = calcTF(terms, docs)
pprint(tf)
tfidf = np.multiply(idf, tf)
pprint(tfidf)
def calcIDF(terms, docs):
doc_count = len(docs)
idf = np.zeros((len(terms), len(docs)))
for i in range(0, len(docs)):
for j in range(0, len(terms)):
term_count = 0
for t in docs[i]:
if t == terms[j]:
term_count += 1
idf[j][i] = term_count / doc_count
return idf
def calcTF(terms, docs):
tf = np.zeros((len(terms), len(docs)))
for i in range(0, len(docs)):
for j in range(0, len(terms)):
tf[j][i] = docs[i].count(terms[j])
return tf
# #### b)
# Perform the singular value decomposition of the above matrix and write down
# the obtained factor matrices U, Σ, and V. You can use some existing programming library
# to perform the SVD (e.g., numpy.linalg.svd in Python).
np.linalg.svd(tfidf)
# #### c)
# Reduce the rank of the factor matrices to K = 2, i.e., compute the 2-dimensional
# vectors for vocabulary terms and documents. Show terms and documents as points in a
# 2-dimensional graph.
# #### d)
# You are given the query “Sam blue orc”. Compute the latent vector for the query
# and rank the documents according to similarity of their latent vectors with the obtained
# latent vector of the query.
# # 2. Text Clustering
d1 = [0.17, 0.21, 0.35, 0.44, 0.49, 0.39, 0.09, 0.07, 0.37, 0.24]
d2 = [0.49, 0.48, 0.44, 0.09, 0.24, 0.2, 0.41, 0.16, 0.1, 0.15]
d3 = [0.41, 0.36, 0.27, 0.19, 0.15, 0.42, 0.23, 0.42, 0.02, 0.42]
d4 = [0.31, 0.41, 0.21, 0.19, 0.47, 0.28, 0.21, 0.39, 0.16, 0.38]
d5 = [0.46, 0.12, 0.21, 0.25, 0.38, 0.38, 0.46, 0.23, 0.31, 0.14]
d6 = [0.13, 0.33, 0.28, 0.42, 0.07, 0.13, 0.58, 0.15, 0.0, 0.49]
d7 = [0.21, 0.09, 0.07, 0.09, 0.3, 0.54, 0.24, 0.43, 0.51, 0.21]
d8 = [0.18, 0.39, 0.42, 0.05, 0.41, 0.1, 0.52, 0.12, 0.14, 0.38]
d9 = [0.4, 0.51, 0.01, 0.1, 0.12, 0.22, 0.26, 0.34, 0.42, 0.38]
docs = [d1, d2, d3, d4, d5, d6, d7, d8, d9]
# #### a)
# Assume that a news outlet is sequentially streaming these documents, from d 1
# to d 9 .
#
# Cluster the documents using the single pass clustering (SPC) algorithm based
# on cosine similarity between the given TF-IDF document vectors.
#
# Run the SPC using different values for similarity threshold:
#
# (i) λ = 0.6, (ii) λ = 0.8.
#
# What is the difference between the two clusterings, using different values for λ?
#
# Next, cluster the documents with SPC assuming the opposite order of streaming, from d 9 to d 1 (use λ = 0.8).
#
# Did you obtain the same clusters as before?
# +
lambda1 = 0.6
lambda2 = 0.8
lambda3 = 0.8
spc1 = singlepassclustering(docs, lambda1)
spc2 = singlepassclustering(docs, lambda2)
docs_reversed = docs[::-1]
spc3 = singlepassclustering(docs_reversed, lambda3)
print("(i)")
pprint(spc1)
print("(ii)")
pprint(spc2)
print("(iii)")
pprint(spc3)
# -
def singlepassclustering(docs, lambdax):
clusters = {}
result = {}
clusters[0] = []
result[0] = []
clusters[0].append(docs[0])
result[0].append(1)
cluster_count = 0
for i in range(1, len(docs)):
cluster_sim = {}
for c in range(0, len(clusters)):
cluster_sim[c] = simDocCluster(docs[i], clusters[c])
x = max(cluster_sim.keys(), key=(lambda key: cluster_sim[key]))
if(cluster_sim[x] > lambdax):
clusters[x].append(docs[i])
result[x].append(i+1)
else:
cluster_count +=1
clusters[cluster_count] = []
clusters[cluster_count].append(docs[i])
result[cluster_count] = []
result[cluster_count].append(i+1)
return result
def simDocCluster(doc, cluster):
similarities = 0.0
for c in cluster:
similarities += cosinesim(doc, c)
return similarities / len(cluster)
def cosinesim(doc1, doc2):
numerator = sum([doc1[x] * doc2[x] for x in range(0, len(doc1))])
sum1 = sum([doc1[x]**2 for x in range(0,len(doc1))])
sum2 = sum([doc2[x]**2 for x in range(0,len(doc2))])
denominator = math.sqrt(sum1) * math.sqrt(sum2)
if not denominator:
return 0.0
else:
return float(numerator) / denominator
# #### b)
# Cluster the above given documents using the k-means algorithm, with K = 3
# and using the following initial centroids:
#
# r1 = [0.33, 0.33, 0.42, 0.12, 0.2, 0.34, 0.58, 0.19, 0.07, 0.24]
#
# r2 = [0.29, 0.16, 0.38, 0.48, 0.43, 0.11, 0.12, 0.33, 0.03, 0.44]
#
# r3 = [0.01, 0.17, 0.11, 0.27, 0.23, 0.37, 0.35, 0.48, 0.54, 0.24].
#
# Use the cosine similarity between document vectors and centroids to guide the clustering.b
r1 = [0.33, 0.33, 0.42, 0.12, 0.2, 0.34, 0.58, 0.19, 0.07, 0.24]
r2 = [0.29, 0.16, 0.38, 0.48, 0.43, 0.11, 0.12, 0.33, 0.03, 0.44]
r3 = [0.01, 0.17, 0.11, 0.27, 0.23, 0.37, 0.35, 0.48, 0.54, 0.24]
centroids = np.array((r1,r2,r3))
kmeans = KMeans(n_clusters=3, random_state=0, init=centroids).fit(docs)
pprint(kmeans.labels_)
# #### Results
# Cluster 0 = d2, d4, d5, d7, d8
#
# Cluster 1 = d6, d9
#
# Cluster 2 = d1, d3,
# # 3. IR Evaluation
r1 = ['d1', 'd2', 'd5', 'd6', 'd13']
r2 = ['d1', 'd2', 'd4', 'd5', 'd6', 'd7', 'd8', 'd9', 'd10', 'd11', 'd12', 'd13', 'd19', 'd14', 'd17', 'd3', 'd15', 'd16', 'd18', 'd20']
r3 = ['d1', 'd2', 'd4', 'd5', 'd9', 'd10', 'd12', 'd13', 'd14', 'd15', 'd20']
# #### a)
# Compute the precision, recall and F 1 score for each of the three IR systems.
#
# What is the downside of using precision, recall, and F measure to evaluate IR systems?
#
# For some query q all odd documents are considered to be relevant and all documents with even identifiers are considered not relevant.
# precision = tp / (tp + fp)
#
# recall = tp / (tp + fn)
#
# f-measure = 2 * (precision * recall) / (precision + recall)
print("IR System 1")
tp_1 = 3
tn_1 = 0
fp_1 = 2
fn_1 = 0
precision_r1 = tp_1 / (tp_1 + fp_1)
print("precision: " + str(precision_r1))
recall_r1 = tp_1 / (tp_1 + fn_1)
print("recall: " + str(recall_r1))
F1_r1 = 2 * ((precision_r1 * recall_r1)/(precision_r1 + recall_r1))
print("f1: " + str(F1_r1))
print()
print("IR System 2")
tp_2 = 5
tn_2 = 5
fp_2 = 5
fn_2 = 5
precision_r2 = tp_2 / (tp_2 + fp_2)
print("precision: " + str(precision_r2))
recall_r2 = tp_2 / (tp_2 + fn_2)
print("recall: " + str(recall_r2))
F1_r2 = 2 * ((precision_r2 * recall_r2)/(precision_r2 + recall_r2))
print("f1: " + str(F1_r2))
print()
print("IR System 3")
tp_3 = 5
tn_3 = 1
fp_3 = 5
fn_3 = 0
precision_r3 = tp_3 / (tp_3 + fp_3)
print("precision: " + str(precision_r3))
recall_r3 = tp_3 / (tp_3 + fn_3)
print("recall: " + str(recall_r3))
F1_r3 = 2 * ((precision_r3 * recall_r3)/(precision_r3 + recall_r3))
print("f1: " + str(F1_r3))
# #### b)
# Compute the precision at rank 5 (P@5), R-precision, and average precision (AP)
# for each of the three IR systems.
# +
print("IR System 1")
pat5_1 = 3 / (3 + 2)
print("P@5: " + str(pat5_1))
r_precision_1 = 2 / (2 + 1)
print("R-precision [R@3]: " + str(r_precision_1))
ap_1 = (1 / 3) * ((1/1)+(2/3)+(3/5))
print("Average Precision: " + str(ap_1))
print()
print("IR System 2")
pat5_2 = 2 / (2 + 3)
print("P@5: " + str(pat5_2))
r_precision_2 = 4 / (4 + 6)
print("R-precision [R@10]: " + str(r_precision_2))
ap_2 = (1 / 10) * ((1/1)+(2/4)+(3/6)+(4/8)+(5/10)+(6/12)+(7/13)+(9/15)+(10/17))
print("Average Precision: " + str(ap_2))
print()
print("IR System 3")
pat5_3 = 3 / (3 + 2)
print("P@5: " + str(pat5_3))
r_precision_3 = 3 / (3 + 2)
print("R-precision [R@5]: " + str(r_precision_3))
ap_3 = (1/5)*((1/1)+(2/4)+(3/5)+(4/8)+(5/10))
print("Average Precision: " + str(ap_3))
# -
# #### c)
# You are given a toy IR system which is being evaluated on five queries.
#
# The following are the positions of relevant documents for each of these five queries, in the rankings returned by the toy IR system:
#
# • q 1 → [1, 6, 9, 17, 21]
#
# • q 2 → [1, 3, 4]
#
# • q 3 → [2, 5, 8, 9, 10]
#
# • q 4 → [4]
#
# • q 5 → [1, 2, 6]
#
# Evaluate the performance of this IR system in terms of mean average precision.
# +
precision_q1 = (1/4) * ((1/1)+(2/3)+(3/4)+(4/5))
precision_q2 = (1/2) * ((1/1)+(2/2))
precision_q3 = (1/2) * ((1/2)+(2/4))
precision_q4 = 0
precision_q5 = (1/1) * (1/1)
MAP = (1/5) * (precision_q1 + precision_q2 + precision_q3 + precision_q3 + precision_q4 + precision_q5)
print("MAP: " + str(MAP))
# -
# # 4. Semantic Retrieval with Word Embeddings
| Homework3/.ipynb_checkpoints/Homework3-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.widgets import Slider
from matplotlib.patches import Rectangle, Circle
import os
from PIL import Image
from scipy.special import erf
#import tifffile
fname = '09_2DWrinkAu_640kx_770mm_70um_0p64mrad_ss8_50x_50y_100z_216step_x256_y256.raw'
fnum = int(fname[0:2])
yrows = 130
xcols = 128
bright_disk_radius = 5
erf_sharpness = 5
hann2d = np.hanning(xcols)
hann2d = np.outer(hann2d, hann2d)
kx = np.arange(-xcols, xcols, 2)/2
kx,ky = np.meshgrid(kx, kx)
dist = np.hypot(kx, ky)
haadf_mask = np.array(dist >= 30, np.int)
x = np.arange(-xcols/2,xcols/2,1)
x,y = np.meshgrid(x,x)
dist = np.hypot(x,y)
bdisk_filter = erf((dist-bright_disk_radius)*erf_sharpness)/2 - erf((dist+bright_disk_radius)*erf_sharpness)/2 + 1
hann_filter = np.hanning(xcols)
hann_filter = np.outer(hann_filter, hann_filter)
#%%
#Pulls a yrows x xcols diffraction pattern at [first_index, second_index] from filename
#Will crop to xcols x xcols if crop == True
def dp_slice(filename, first_index = 0, second_index = 0, yrows = yrows,
xcols = xcols, crop = True, dtype = np.float32, min_count_to_zero = 20, min_offset = 1e-6):
dsize = np.dtype(dtype).itemsize
num_pixels = int((os.stat(filename).st_size/yrows/xcols/dsize)**0.5)
offset = int((num_pixels*first_index + second_index)*yrows*xcols*dsize)
dp_slice = np.memmap(filename, dtype = dtype, mode = 'r', shape = (yrows, xcols),
order = 'C', offset = offset)
dp_slice = np.array(dp_slice)
if crop:
dp_slice = dp_slice[:xcols, :xcols]
#Counts under min_count set to min_offset to be very close to zero but not exactly zero to avoid errors
#with taking logarithms
dp_slice[dp_slice <= min_count_to_zero] = min_offset
return dp_slice
#Transforms either dpslice or full 4D dp to cepstrum
def dp_to_cep(dp, window = hann2d):
# cep = dp*window
# cep[cep==0] = 0.0001
cep = np.log10(dp)
cep = np.fft.fft2(cep)
cep = np.fft.fftshift(cep, (-1, -2))
cep = np.abs(cep)**2.0
return cep
#Creates image from filename dp based on mask
def generate_image(filename, mask, yrows = yrows, xcols = xcols, dtype = np.float32):
dsize = np.dtype(dtype).itemsize
num_pixels = int((os.stat(filename).st_size/yrows/xcols/dsize)**0.5)
haadf = np.zeros((num_pixels, num_pixels))
for i in range(num_pixels):
for j in range(num_pixels):
haadf[i,j] = np.sum(dp_slice(filename, i, j)*haadf_mask)
return haadf
def browser(image, filename, cep_max):
#Cursor used to select which scanning point to show associated diffraction pattern and linescan
class Cursor(object):
def __init__(self, ax):
self.ax = ax
self.lock = False
self.lx = ax.axhline(color = 'k')
self.ly = ax.axvline(color = 'k')
self.x = 0
self.y = 0
def mouse_move(self, event):
if not event.inaxes == self.ax:
return
if self.lock:
return
x,y = event.xdata, event.ydata
x = int(round(x))
y = int(round(y))
self.x = x
self.y = y
self.lx.set_ydata(y)
self.ly.set_xdata(x)
update_dps(y, x)
plt.draw()
def click(self, event):
if not event.inaxes == self.ax:
return
if not event.dblclick:
return
self.lock = not self.lock
def update_dps(y, x):
dslice = dp_slice(filename, y, x)
cep = dp_to_cep(dslice, window = hann_filter)
dpdisp.set_data(dslice)
dpdisp.set_clim(dp_min_sl.val, dp_max_sl.val)
dpmin = np.min(dslice)
dpmax = np.max(dslice)
dp_min_sl.valmin = dpmin
dp_min_sl.valmax = dpmax
dp_min_ax.set_xlim(dpmin, dpmax)
dp_max_sl.valmin = dpmin
dp_max_sl.valmax = dpmax
dp_max_ax.set_xlim(dpmin, dpmax)
cepdisp.set_data(cep)
cepdisp.set_clim(cep_min_sl.val, cep_max_sl.val)
cepmin = np.min(cep)
cep_min_sl.valmin = cepmin
cep_min_ax.set_xlim(cepmin, cep_max)
cep_max_sl.valmin = cepmin
cep_max_ax.set_xlim(cepmin, cep_max)
plt.draw()
def update_clim(disp):
if disp == 'dp':
dpdisp.set_clim(dp_min_sl.val, dp_max_sl.val)
elif disp == 'cep':
cepdisp.set_clim(cep_min_sl.val, cep_max_sl.val)
plt.draw()
fig, ax = plt.subplots(1, 3)
fig.set_size_inches(15, 5)
plt.subplots_adjust(bottom = 0.20, left = 0.00, right = 0.95)
ax[0].imshow(image, origin = 'lower', aspect = 'equal')
ax[0].invert_xaxis()
ax[0].axis('off')
dslice = dp_slice(filename, 0, 0)
cep = dp_to_cep(dslice)
dpdisp = ax[1].imshow(dslice, origin = 'upper', aspect = 'equal')
c1 = Circle((64,64), 19.41858, fill = False, linestyle = '--', color = 'red', linewidth = 4)
c2 = Circle((64,64), 19.41858, fill = False, linestyle = '--', color = 'blue', linewidth = 4)
c1.set_radius(0)
c2.set_radius(0)
ax[1].add_artist(c1)
ax[1].add_artist(c2)
# ax[1].set_xlim(43, 85)
# ax[1].set_ylim(43, 85)
ax[1].axis('off')
plt.colorbar(dpdisp, ax = ax[1])
cepdisp = ax[2].imshow(cep, origin = 'upper', aspect = 'equal')
c1 = Circle((64,64), 11.02, fill = False, linestyle = '--', color = 'red', linewidth = 4)
c2 = Circle((64,64), 9.665, fill = False, linestyle = '--', color = 'blue', linewidth = 4)
ax[2].add_artist(c1)
ax[2].add_artist(c2)
c1.set_radius(0)
c2.set_radius(29.88/2*(3/11)**0.5)
ax[2].axis('off')
plt.colorbar(cepdisp, ax = ax[2])
cursor = Cursor(ax[0])
plt.connect('motion_notify_event', cursor.mouse_move)
plt.connect('button_press_event', cursor.click)
dp_min_ax = plt.axes([0.35, 0.15, 0.20, 0.03])
dp_max_ax = plt.axes([0.35, 0.10, 0.20, 0.03])
cep_min_ax = plt.axes([0.70, 0.15, 0.20, 0.03])
cep_max_ax = plt.axes([0.70, 0.10, 0.20, 0.03])
dp_min_sl = Slider(dp_min_ax, 'Min', np.min(dslice), np.max(dslice), valinit = 0)
dp_max_sl = Slider(dp_max_ax, 'Max', np.min(dslice), np.max(dslice), valinit = 0)
dp_max_sl.set_val(np.max(dslice))
dp_min_sl.slidermax = dp_max_sl
dp_max_sl.slidermin = dp_min_sl
cep_min_sl = Slider(cep_min_ax, 'Min', np.min(cep), cep_max, valinit = 0)
cep_max_sl = Slider(cep_max_ax, 'Max', np.min(cep), cep_max, valinit = 0)
cep_max_sl.set_val(cep_max)
cep_min_sl.slidermax = cep_max_sl
cep_max_sl.slidermin = cep_min_sl
dp_min_sl.on_changed(lambda x: update_clim('dp'))
dp_max_sl.on_changed(lambda x: update_clim('dp'))
cep_min_sl.on_changed(lambda x: update_clim('cep'))
cep_max_sl.on_changed(lambda x: update_clim('cep'))
return cursor, dp_min_sl, dp_max_sl, cep_min_sl, cep_max_sl
def pull_current_dp_cep(browser):
y = browser[0].y
x = browser[0].x
dp_fname = '%02d_dp_%d_%d.tif' % (fnum, y, x)
cep_fname = '%02d_cep_%d_%d.tif' % (fnum, y, x)
dp = dp_slice(fname, y, x)
cep = dp_to_cep(dp)
#tifffile.imwrite(dp_fname, dp)
#tifffile.imwrite(cep_fname, cep)
return dp_fname, cep_fname
#%%
im = generate_image(fname, haadf_mask)
plt.close('all')
brow = browser(im, fname, cep_max = 1e5)
# -
| Original.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # DEMO: Running FastCAM using SMOE Scale Saliency Maps
# ### NOTE: This notebook gives a detailed description of how FastCAM works. The other notebooks are shorter and aimed at just running the code.
#
# ### See also:
# demo_simple_fast-cam.ipynb
# demo_very-simple_fast-cam.ipynb
# demo-captum.ipynb
# ## Initial code setup
import os
import cv2
import numpy as np
import torch
from torchvision import models
from IPython.display import Image
# Lets load things we need for **Grad-CAM**
# +
from torchvision.utils import make_grid, save_image
import torch.nn.functional as F
from gradcam.utils import visualize_cam
from gradcam import GradCAMpp, GradCAM
# The GradCAM kit throws a warning we don't need to see for this demo.
import warnings
warnings.filterwarnings('ignore')
# -
# Now we import the code for **this package**.
import maps
import mask
import draw
import norm
import misc
# This is where we can set some parameters like the image name and the layer weights.
# +
input_image_name = "ILSVRC2012_val_00049934.224x224.png" # Our input image to process
output_dir = 'outputs' # Where to save our output images
input_dir = 'images' # Where to load our inputs from
weights = [1.0, 1.0, 1.0, 1.0, 1.0] # Equal Weights work best
# when using with GradCAM
#weights = [0.18, 0.15, 0.37, 0.4, 0.72] # Our saliency layer weights
# From paper:
# https://arxiv.org/abs/1911.11293
save_prefix = input_image_name[:-4].split('/')[-1] # Chop the file extension and path
load_image_name = os.path.join(input_dir, input_image_name)
os.makedirs(output_dir, exist_ok=True)
# -
# Good Doggy!
Image(filename=load_image_name)
# Now we create a model in PyTorch and send it to our device.
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
model = models.resnet50(pretrained=True)
model = model.to(device)
# ## Setting up Hooks
# Now we will set up our layer hooks as callback methods. This will keep a copy of a layers output data from a forward pass. This is how we will get the data out of the network's layers.
#
# So, lets look at the second spatial scale of ResNet-50 which has three bottleneck layers. We want that very last ReLU at the very end.
print("{}".format(model.layer1))
# <br>
#
# **So...** we would select
#
# model.layer1[2].relu
#
# with a command like
#
# model.layer1[2].relu.register_forward_hook(hooks[1])
#
# This attaches a callback which will store this layers output, when the network is run.
#
# Below we attach a hook at the end of each spatial scale.
# +
hooks = [misc.CaptureLayerOutput() for i in range(5)] # Create 5 callback hooks in a liszt
map_num = len(hooks) # Store size for later use
# Chopin off the "_ =" is fine. We technically don't
# need anything bach but the handel to each hook.
_ = model.relu.register_forward_hook(hooks[0])
_ = model.layer1[2].relu.register_forward_hook(hooks[1])
_ = model.layer2[3].relu.register_forward_hook(hooks[2])
_ = model.layer3[5].relu.register_forward_hook(hooks[3])
_ = model.layer4[2].relu.register_forward_hook(hooks[4])
# -
# Notice that the layers we access are the same as in the resnet50 python model layers specificiation which is **[3, 4, 6, 3]**, we just subtract 1 from each.
#
# OK, now let's load in our image and set it to the standard *ImageNet* size: 224x224.
# +
in_tensor = misc.LoadImageToTensor(load_image_name, device)
in_tensor = F.interpolate(in_tensor, size=(224, 224), mode='bilinear', align_corners=False)
in_height = in_tensor.size()[2]
in_width = in_tensor.size()[3]
# -
# OK, let's run our ResNet network *forward only*. The callbacks will then contain the layer data afterwards. Since we are only doing a forward computation, this sails on a CPU.
model.eval()
with torch.set_grad_enabled(False):
_ = model(in_tensor)
# ## Running saliency on our network data
#
# Now that the network has run, we can finally do some work. First we create objects for our saliency layers. These are *PyTorch nn layers*, but we can talk to them even if they are not inside a network.
# All three objects here are technically layers. So, they can be used inside your network as well.
#
# If you want to use these **inside your network**:
#
# example, in your __init__ something like:
#
# self.salmap_layer = maps.SMOEScaleMap()
#
# then in forward(x) something like:
#
# x = self.relu(x)
# x = self.salmap_layer(x)
# First, we create an object to get each saliency map given the data stored in the hooks. This will run SMOE Scale on each of the layer hook outputs and produce raw a saliency map. This package also includes code for using *Standard Deviation* or *Truncated Normal Entropy*.
getSmap = maps.SMOEScaleMap()
# This next object will normalize the saliency maps from 0 to 1 using a Gaussian CDF squashing function.
#getNorm = norm.GaussNorm2D()
getNorm = norm.GammaNorm2D() # A little more accurate, but much slower
# Now we will create an object to combine the five saliency maps from each scale into one.
getCsmap = maps.CombineSaliencyMaps(output_size=[in_height,in_width],
map_num=map_num, weights=weights, resize_mode='bilinear')
# Once we have our objects, we will run SMOE Scale on each of the output hooks and then normalize the output.
smaps = [ getNorm(getSmap(x.data)) for x in hooks ]
# Now, we combine the different saliency maps into a single combined saliency map. Notice that we also get back each saliency map in *smaps* rescaled by the method.
csmap,smaps = getCsmap(smaps)
# Then we save and view it.
# +
output_name = "{}.MAP_COMBINED.jpg".format(save_prefix)
output_path = os.path.join(output_dir, output_name)
misc.SaveGrayTensorToImage(csmap, output_path)
np_smaps = misc.TensorToNumpyImages(smaps) # For later use, keep the Numpy version
# of the saliency maps.
Image(filename=output_path)
# -
# Now lets get our individual saliency maps for each of the five layers and look at them.
# +
il = [smaps[0,i,:,:] for i in range(map_num)] # Put each saliency map into the figure
il.append(csmap[0,:,:]) # add in the combined map at the end of the figure
images = [torch.stack(il, 0)]
images = make_grid(torch.cat(images, 0), nrow=5)
output_name = "{}.SAL_MAPS.jpg".format(save_prefix)
sal_img_file = os.path.join(output_dir, output_name)
save_image(images.unsqueeze(1), sal_img_file)
Image(filename=sal_img_file)
# -
# ## LOVI and Heat Maps
# Now we will take our saliency maps and create the LOVI and Heat Map versions.
# +
SHM = draw.HeatMap(shape=np_smaps.shape, weights=weights ) # Create our heat map drawer
LOVI = draw.LOVI(shape=np_smaps.shape, weights=None) # Create out LOVI drawer
shm_im = SHM.make(np_smaps) # Combine the saliency maps
# into one heat map
lovi_im = LOVI.make(np_smaps) # Combine the saliency maps
# into one LOVI image
# -
# Next, save and display our images.
# +
output_name = "{}.HEAT.jpg".format(save_prefix)
output_path = os.path.join(output_dir, output_name)
cv2.imwrite(output_path, (shm_im*255.0).astype(np.uint8))
Image(filename=output_path)
# -
# Let's create on overlay with our original image by alpha blending.
# +
cv_im = cv2.imread(load_image_name).astype(np.float32) / 255.0
cv_im = cv2.resize(cv_im, (224,224))
ab_shm = misc.AlphaBlend(shm_im, cv_im) # Blend the heat map and the original image
output_name = "{}.ALPHA_HEAT.jpg".format(save_prefix)
output_path = os.path.join(output_dir, output_name)
cv2.imwrite(output_path, (ab_shm*255.0).astype(np.uint8))
Image(filename=output_path)
# -
# Now we view our LOVI map with and without alpha blending of the original image. The LOVI image tells us which parts of the network are most active by layer. We range like a rainbow with violet/blue representing early layers and yellow/red representing later layers. White areas are active over all layers.
# +
output_name = "{}.LOVI.jpg".format(save_prefix)
output_path = os.path.join(output_dir, output_name)
cv2.imwrite(output_path, (lovi_im*255.0).astype(np.uint8))
Image(filename=output_path)
# -
# You can see how this image is composed by looking again at all the individual saliency maps...
Image(filename=sal_img_file)
# +
ab_lovi = misc.AlphaBlend(lovi_im, cv_im) # Blend original image and LOVI image
output_name = "{}.ALPHA_LOVI.jpg".format(save_prefix)
output_path = os.path.join(output_dir, output_name)
cv2.imwrite(output_path, (ab_lovi*255.0).astype(np.uint8))
Image(filename=output_path)
# -
# ## The Masked Image
# From the combined saliency map, we can extract the masked out input image. This illustrates What parts of the image are the top xx% most salient.
#
# First we set up objects to create the mask from the input tensor version of the image and the combined saliency map.
getMask = mask.SaliencyMaskDropout(keep_percent = 0.1, scale_map=False)
# We define a denormalization object to get things back to normal image pixel values.
denorm = misc.DeNormalize([0.485, 0.456, 0.406], [0.229, 0.224, 0.225])
# Next we process the tensor and saliency map to create the masked image.
masked_tensor,drop_map = getMask(in_tensor, csmap)
# Finally, we denormalize the tensor image, save and display it. This shows us the **top 10% most salient parts** of the images.
# +
masked_tensor = denorm(masked_tensor)
output_name = "{}.MASK.jpg".format(save_prefix)
output_path = os.path.join(output_dir, output_name)
misc.SaveColorTensorToImage(masked_tensor, output_path)
Image(filename=output_path)
# -
# ## Run With Grad-CAM++
# Let's go ahead and push our network model into the Grad-CAM library.
#
# **NOTE** much of this code is borrowed from the Pytorch GradCAM package.
resnet_gradcampp4 = GradCAMpp.from_config(model_type='resnet', arch=model, layer_name='layer4')
# Let's get our original input image back.
raw_tensor = misc.LoadImageToTensor(load_image_name, device, norm=False)
raw_tensor = F.interpolate(raw_tensor, size=(224, 224), mode='bilinear', align_corners=False)
# Now we will create illustrations of the combined saliency map.
# +
masked_tensor_raw,drop_map = getMask(raw_tensor, csmap)
cs_heatmap, cs_result = visualize_cam(csmap, raw_tensor)
cs_masked = misc.AlphaMask(raw_tensor, csmap).squeeze(0)
cs_masked = misc.RangeNormalize(cs_masked)
images = []
images.append(torch.stack([raw_tensor.squeeze().cpu(), cs_heatmap.cpu(),
cs_result.cpu(), cs_masked.cpu(), masked_tensor_raw[0,].cpu()], 0))
# -
# Now, lets get the Grad-CAM++ saliency map only.
mask_pp1, logit = resnet_gradcampp4(in_tensor)
# Let's double check and make sure it's picking the correct class
too_logit = logit.max(1)
print("Network Class Output: {} : Value {} ".format(too_logit[1][0],too_logit[0][0]))
# Now visualize the results
# +
heatmap_pp1, result_pp1 = visualize_cam(mask_pp1, raw_tensor)
hard_masked_pp1,_ = getMask(raw_tensor, mask_pp1.squeeze(0))
hard_masked_pp1 = hard_masked_pp1.squeeze(0)
masked_pp1 = misc.AlphaMask(raw_tensor, mask_pp1.squeeze(0)).squeeze(0)
masked_pp1 = misc.RangeNormalize(masked_pp1)
images.append(torch.stack([raw_tensor.squeeze().cpu(), heatmap_pp1.cpu(),
result_pp1.cpu(), masked_pp1.cpu(), hard_masked_pp1.cpu()], 0))
# -
# **Now we combine the Grad-CAM map and the SMOE Scale saliency maps** in the same way we would combine Grad-CAM with Guided Backprop.
mask_pp2 = csmap*mask_pp1
# Now let's visualize the combined saliency map from SMOE Scale and GradCAM++.
# +
heatmap_pp2, result_pp2 = visualize_cam(mask_pp2, raw_tensor)
hard_masked_pp2,_ = getMask(raw_tensor,mask_pp2.squeeze(0))
hard_masked_pp2 = hard_masked_pp2.squeeze(0)
masked_pp2 = misc.AlphaMask(raw_tensor, mask_pp2.squeeze(0)).squeeze(0)
masked_pp2 = misc.RangeNormalize(masked_pp2)
images.append(torch.stack([raw_tensor.squeeze().cpu(), heatmap_pp2.cpu(),
result_pp2.cpu(), masked_pp2.cpu(), hard_masked_pp2.cpu()], 0))
# -
# Now we combine the Grad-CAM map and the SMOE Scale saliency maps but create a map of the **non-class** objects. These are salient locations that the network found interesting, but are not part of the object class.
mask_pp3 = csmap*(1.0 - mask_pp1)
# Now let's visualize the combined non-class saliency map from SMOE Scale and GradCAM++.
# +
heatmap_pp3, result_pp3 = visualize_cam(mask_pp3, raw_tensor)
hard_masked_pp3,_ = getMask(raw_tensor,mask_pp3.squeeze(0))
hard_masked_pp3 = hard_masked_pp3.squeeze(0)
masked_pp3 = misc.AlphaMask(raw_tensor, mask_pp3.squeeze(0)).squeeze(0)
masked_pp3 = misc.RangeNormalize(masked_pp3)
images.append(torch.stack([raw_tensor.squeeze().cpu(), heatmap_pp3.cpu(),
result_pp3.cpu(), masked_pp3.cpu(), hard_masked_pp3.cpu()], 0))
# -
# We now put all the images into a nice grid for display.
images = make_grid(torch.cat(images,0), nrow=5)
# ... save and look at it.
# +
output_name = "{}.CAM_PP.jpg".format(save_prefix)
output_path = os.path.join(output_dir, output_name)
save_image(images, output_path)
Image(filename=output_path)
# -
# The top row is the SMOE Scale based saliency map. The second row is GradCAM++ only. Next we have the FastCAM output from combining the two. The last row is the non-class map showing salient regions that are not associated with the output class.
#
# This image should look **exactly** like the one on the README.md on Github minus the text.
| demo_fast-cam.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Isinstance() before Numpy arraying:
#
# This is to test if it is more efficient to np.array() things all the time or to first check if they are already np.ndarrays.
#
# <NAME>
# 31/5/2016
# +
import numpy as np
# Make up some random data
large_test_numpy = np.array(np.random.randn(100000), dtype="float64")
small_test_numpy = np.array(np.random.randn(10), dtype="float64")
large_test_list = list(large_test_numpy)
small_test_list = list(small_test_numpy)
# -
# ### Timing calls
# From the following cells an isinstance call is around an order of magnitude faster than a np.array call on a small np.ndarray and around 3 orders of magnitude for a large numpy array. A np.array call on a np.ndarray is order of magnitudes faster then a np.array call on a large list.
#
# Strangely a negative isinstace response is twice as long as a positive response.
#
# +
# Time Small lists and arrays
print("Positive isinstance call:")
% timeit isinstance(small_test_numpy, np.ndarray)
print("\nNegative isinstance call:")
% timeit isinstance(small_test_list, np.ndarray)
print("\nSmall numpy array -> numpy array:")
% timeit np.array(small_test_numpy, dtype="float64")
print("\nSmall list -> numpy array:")
% timeit np.array(small_test_list, dtype="float64")
# +
# Time larger lsits and arrays
print("Positive isinstance call:")
% timeit isinstance(large_test_numpy, np.ndarray)
print("\nNegative isinstance call:")
% timeit isinstance(large_test_list, np.ndarray)
print("\nLarge numpy array -> numpy array:")
% timeit np.array(large_test_numpy, dtype="float64")
print("\nLarge list -> numpy array:")
% timeit np.array(large_test_list, dtype="float64")
# -
# %%timeit
if not isinstance(small_test_numpy, np.ndarray):
np.array(small_test_numpy, dtype="float64")
# %%timeit
if not isinstance(large_test_numpy, np.ndarray):
np.array(large_test_numpy, dtype="float64")
# %%timeit
if type(small_test_list) is not np.ndarray:
np.array(small_test_list, dtype="float64")
# %%timeit
if type(large_test_list) is not np.ndarray:
np.array(large_test_list, dtype="float64")
# ## Conclusion:
#
# If want to make sure that you value is a numpy array instead of just calling np.array on the object first do an isinstance call to check that it is not an numpy array already for a slight efficiency gain 10-100 microseconds per instance.
#
# #### Note:
# Not suitable if you are wanting to change the type of numpy array. I.e float32, float64, int. This should probably be done in the begining.
| Numpy - Arraying an Array.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <table> <tr>
# <td style="background-color:#ffffff;">
# <a href="http://qworld.lu.lv" target="_blank"><img src="..\images\qworld.jpg" width="25%" align="left"> </a></td>
# <td style="background-color:#ffffff;vertical-align:bottom;text-align:right;">
# prepared by <a href="http://abu.lu.lv" target="_blank"><NAME></a> (<a href="http://qworld.lu.lv/index.php/qlatvia/" target="_blank">QLatvia</a>)
# </td>
# </tr></table>
# <table width="100%"><tr><td style="color:#bbbbbb;background-color:#ffffff;font-size:11px;font-style:italic;text-align:right;">This cell contains some macros. If there is a problem with displaying mathematical formulas, please run this cell to load these macros. </td></tr></table>
# $ \newcommand{\bra}[1]{\langle #1|} $
# $ \newcommand{\ket}[1]{|#1\rangle} $
# $ \newcommand{\braket}[2]{\langle #1|#2\rangle} $
# $ \newcommand{\dot}[2]{ #1 \cdot #2} $
# $ \newcommand{\biginner}[2]{\left\langle #1,#2\right\rangle} $
# $ \newcommand{\mymatrix}[2]{\left( \begin{array}{#1} #2\end{array} \right)} $
# $ \newcommand{\myvector}[1]{\mymatrix{c}{#1}} $
# $ \newcommand{\myrvector}[1]{\mymatrix{r}{#1}} $
# $ \newcommand{\mypar}[1]{\left( #1 \right)} $
# $ \newcommand{\mybigpar}[1]{ \Big( #1 \Big)} $
# $ \newcommand{\sqrttwo}{\frac{1}{\sqrt{2}}} $
# $ \newcommand{\dsqrttwo}{\dfrac{1}{\sqrt{2}}} $
# $ \newcommand{\onehalf}{\frac{1}{2}} $
# $ \newcommand{\donehalf}{\dfrac{1}{2}} $
# $ \newcommand{\hadamard}{ \mymatrix{rr}{ \sqrttwo & \sqrttwo \\ \sqrttwo & -\sqrttwo }} $
# $ \newcommand{\vzero}{\myvector{1\\0}} $
# $ \newcommand{\vone}{\myvector{0\\1}} $
# $ \newcommand{\stateplus}{\myvector{ \sqrttwo \\ \sqrttwo } } $
# $ \newcommand{\stateminus}{ \myrvector{ \sqrttwo \\ -\sqrttwo } } $
# $ \newcommand{\myarray}[2]{ \begin{array}{#1}#2\end{array}} $
# $ \newcommand{\X}{ \mymatrix{cc}{0 & 1 \\ 1 & 0} } $
# $ \newcommand{\Z}{ \mymatrix{rr}{1 & 0 \\ 0 & -1} } $
# $ \newcommand{\Htwo}{ \mymatrix{rrrr}{ \frac{1}{2} & \frac{1}{2} & \frac{1}{2} & \frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & \frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} & \frac{1}{2} } } $
# $ \newcommand{\CNOT}{ \mymatrix{cccc}{1 & 0 & 0 & 0 \\ 0 & 1 & 0 & 0 \\ 0 & 0 & 0 & 1 \\ 0 & 0 & 1 & 0} } $
# $ \newcommand{\norm}[1]{ \left\lVert #1 \right\rVert } $
# $ \newcommand{\pstate}[1]{ \lceil \mspace{-1mu} #1 \mspace{-1.5mu} \rfloor } $
# <h2> Python: Quick Reference </h2>
# <hr>
# <ul>
# <li><a href="#Variables">Variables</a></li>
# <li><a href="#Arithmetic_operators">Arithmetic operators</a></li>
# <li><a href="#Objects">Objects</a></li>
# <li><a href="#Size_of_an_object">Size of an object</a></li>
# <li><a href="#Loops">Loops</a></li>
# <li><a href="#Conditionals">Conditionals</a></li>
# <li><a href="#Logical_and_Boolean_operators">Logical and Boolean operators</a></li>
# <li><a href="#Double_list">Double list</a></li>
# <li><a href="#List_operations">List operations</a></li>
# <li><a href="#Functions">Functions</a></li>
# <li><a href="#Random_number">Random number</a></li>
# </ul>
# <hr>
# <a id="Variables"></a>
# <h3> Variables </h3>
# +
number = 5 # integer
real = -3.4 # float
name = 'Asja' # string
surname = "Sarkana" # string
boolean1 = True # Boolean
boolean2 = False # Boolean
# -
# <hr>
# <a id="Arithmetic_operators"></a>
# <h3> Arithmetic operators </h3>
# <h4> Basic operators </h4>
# +
a = 13
b = 5
print("a =",a)
print("b =",b)
print()
# basics operators
print("a + b =",a+b)
print("a - b =",a-b)
print("a * b =",a*b)
print("a / b =",a/b)
# -
# <h4> Integer division and modulus operators </h4>
# +
a = 13
b = 5
print("a =",a)
print("b =",b)
print()
print("integer division:")
print("a//b =",a//b)
print("modulus operator:")
print("a mod b =",a % b)
# -
# <h4> Exponent operator</h4>
#
# number\*\*exponent
# +
b = 5
print("b =",b)
print()
print("b*b =",b**2)
print("b*b*b =",b**3)
print("sqrt(b)=",b**0.5)
# -
# <hr>
# <a id="Objects"></a>
# <h3> Objects </h3>
# <h4> Lists </h4>
# +
# list
mylist = [10,8,6,4,2]
print(mylist)
# -
# <h4> Tuple </h4>
# +
# tuple
mytuple=(1,4,5,'Asja')
print(mytuple)
# -
# <h4> Dictionary </h4>
# +
# dictionary
mydictionary = {
'name' : "Asja",
'surname':'Sarkane',
'age': 23
}
print(mydictionary)
print(mydictionary['surname'])
# -
# <h4>List of the other objects or variables </h4>
# +
# list of the other objects or variables
list_of_other_objects =[
mylist,
mytuple,
3,
"Asja",
mydictionary
]
print(list_of_other_objects)
# -
# <hr>
# <a id="Size_of_an_object"></a>
# <h3> Size of an object </h3>
#
# We use the method "len()" that takes an object as the input.
# +
# length of a string
print(len("<NAME>"))
# size of a list
print(len([1,2,3,4]))
# size of a dictionary
mydictionary = { 'name' : "Asja", 'surname':'Sarkane', 'age': 23}
print(len(mydictionary))
# -
# <hr>
# <a id="Loops"></a>
# <h3> Loops </h3>
# <h4> While-loop </h4>
i = 10
while i>0: # while condition(s):
print(i)
i = i - 1
# <h4> For-loop </h4>
for i in range(10): # i is in [0,1,...,9]
print(i)
for i in range(-5,6): # i is in [-5,-4,...,0,...,4,5]
print(i)
for i in range(0,23,4): # i is in [0,4,8,12,16,20]
print(i)
for i in [3,8,-5,11]:
print(i)
for i in "Sarkane":
print(i)
# +
# dictionary
mydictionary = {
'name' : "Asja",
'surname':'Sarkane',
'age': 23,
}
for key in mydictionary:
print("key is",key,"and its value is",mydictionary[key])
# -
# <hr>
# <a id="Conditionals"></a>
# <h3>Conditionals </h3>
for a in range(4,7):
# if condition(s)
if a<5:
print(a,"is less than 5")
# elif conditions(s)
elif a==5:
print(a,"is equal to 5")
# else
else:
print(a,"is greater than 5")
# <hr>
# <a id="Logical_and_Boolean_operators"></a>
# <h3>Logical and Boolean operators</h3>
# <h4> Logical operator "and"</h4>
# Logical operator "and"
i = -3
j = 4
if i<0 and j > 0:
print(i,"is negative AND",j,"is positive")
# <h4>Logical operator "or"</h4>
# Logical operator "or"
i = -2
j = 2
if i==2 or j == 2:
print("i OR j is 2: (",i,",",j,")")
# <h4>Logical operator "not" </h4>
# Logical operator "not"
i = 3
if not (i==2):
print(i,"is NOT equal to 2")
# <h4> Operator "equal to" </h4>
# Operator "equal to"
i = -1
if i == -1:
print(i,"is EQUAL TO -1")
# <h4> Operator "not equal to" </h4>
# Operator "not equal to"
i = 4
if i != 3:
print(i,"is NOT EQUAL TO 3")
# <h4> Operator "less than or equal to" </h4>
# Operator "not equal to"
i = 2
if i <= 5:
print(i,"is LESS THAN OR EQUAL TO 5")
# <h4> Operator "greater than or equal to" </h4>
# Operator "not equal to"
i = 5
if i >= 1:
print(i,"is GREATER THAN OR EQUAL TO 3")
# <hr>
# <a id="Double_list"></a>
# <h3> Double list </h3>
# +
A =[
[1,2,3],
[-2,-4,-6],
[3,6,9]
]
# print all
print(A)
print()
# print each list in a new line
for list in A:
print(list)
# -
# <hr>
# <a id="List_operations"></a>
# <h3>List operations</h3>
# <h4> Concatenation of two lists </h4>
# +
list1 = [1,2,3]
list2 = [4,5,6]
#concatenation of two lists
list3 = list1 + list2
print(list3)
list4 = list2 + list1
print(list4)
# -
# <h4> Appending a new element </h4>
# +
list = [0,1,2]
list.append(3)
print(list)
list = list + [4]
print(list)
# -
# <hr>
# <a id="Functions"></a>
# <h3>Functions</h3>
# +
def summation_of_integers(n):
summation = 0
for integer in range(n+1):
summation = summation + integer
return summation
print(summation_of_integers(10))
print(summation_of_integers(20))
# -
# <hr>
# <a id="Random_number"></a>
# <h3>Random number</h3>
#
# We can use method "randrange()".
# +
from random import randrange
print(randrange(10),"is picked randomly between 0 and 9")
print(randrange(-9,10),"is picked randomly between -9 and 9")
print(randrange(0,20,3),"is picked randomly from the list [0,3,6,9,12,15,18]")
# -
| python/Python04_Quick_Reference.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Determine distance between line segments in 2D.
#
# Aims to handle many fringe cases not usually handled by generic linear algebra equations.
#
# Convention: Line segments defined by X,Y values of endpoints.
#
# Exposes intermediary support calculations.
import numpy as np
# +
# First function calculates shortest distance between a point and an infinite line
def point_to_inf_line(x_pt, y_pt, x_line_1, y_line_1, x_line_2, y_line_2):
dist = (np.abs((y_line_2 - y_line_1) * x_pt - (x_line_2 - x_line_1) *
y_pt + x_line_2 * y_line_1 - y_line_2 * x_line_1) /
np.sqrt((y_line_2 - y_line_1)**2 + (x_line_2 - x_line_1)**2))
return(dist)
point_to_inf_line(1,0,2,1,3,1) # returns distance 1.0
# +
# Calculate minimum distance between point and a line segment
def point_to_line_segment(x_pt, y_pt, x_line_1, y_line_1, x_line_2, y_line_2):
dist_inf_line = point_to_inf_line(x_pt, y_pt, x_line_1, y_line_1, x_line_2, y_line_2)
# define line from points, in ax + by + c = 0 form
# check for vertical line case
if x_line_1 == x_line_2:
line_a = 1
line_b = 0
line_c = -1 * x_line_1
else:
line_m = (y_line_2 - y_line_1) / (x_line_2 - x_line_1) # slope
line_int = y_line_2 - line_m * x_line_2 # y-intercept
line_a = -1 * line_m # put in ax + by + c = 0 form
line_b = 1 # assume equal to 1
line_c = -1 * line_int
# find point on line for shortest distance
x_int = (line_b*(line_b * x_pt - line_a * y_pt) - line_a * line_c) / (line_a ** 2 + line_b ** 2)
y_int = (line_a * (-1 * line_b * x_pt + line_a * y_pt) - line_b * line_c) / (line_a ** 2 + line_b ** 2)
# if point falls in bounds of line segment, treat as infinite line for calculating distance
if (x_int >= np.min([x_line_1, x_line_2]) and x_int <= np.max([x_line_1, x_line_2]) and
y_int >= np.min([y_line_1, y_line_2]) and y_int <= np.max([y_line_1, y_line_2])):
return(dist_inf_line)
else:
# if nearest point not on segment, use segment endpoints as potential nearest points
# return shrtest distance from pt to segment endpoints
dist_end_pt_1 = np.sqrt((y_line_1 - y_pt)**2 + (x_line_1 - x_pt)**2)
dist_end_pt_2 = np.sqrt((y_line_2 - y_pt)**2 + (x_line_2 - x_pt)**2)
return(np.min([dist_end_pt_1, dist_end_pt_2]))
point_to_line_segment(1,0,1,1,3,1) # should equal 1
point_to_line_segment(0,0,0,-1,0,1) # equal 0, point falls on line
point_to_line_segment(2, 1, 0, 0, 0, 1) # vertical line
# -
def segment_to_segment(x1, y1, x2, y2, x3, y3, x4, y4):
# center data to origin to avoid precision problems
# this can be encountered with survey data where points are millions of feet from datum
min_x = np.min([x1, x2, x3, x4])
min_y = np.min([y1, y2, y3, y4])
x1 = x1 - min_x
x2 = x2 - min_x
x3 = x3 - min_x
x4 = x4 - min_x
y1 = y1 - min_y
y2 = y2 - min_y
y3 = y3 - min_y
y4 = y4 - min_y
# check vertical lines
if x1 == x2 and x3 == x4: # if both vertical
# then flip x and y axes, so that slope is not Inf
min_dist = np.min([point_to_line_segment(y1, x1, y3, x3, y4, x4),
point_to_line_segment(y2, x2, y3, x3, y4, x4),
point_to_line_segment(y3, x3, y1, x1, y2, x2),
point_to_line_segment(y4, x4, y1, x1, y2, x2)])
return(min_dist)
# find intersection between the 2 as infinite lines
elif x1 == x2: # if first vertical
x_int = x1
y_int = (y4 - y3)/(x4 - x3)*(x_int - x3) + y3 # point-slope intercept form
elif x3 == x4: # if second vertical
x_int = x3
y_int = (y2 - y1)/(x2 - x1)*(x_int - x1) + y1 # point-slope intercept form
elif (y2 - y1)/(x2 - x1) == (y4 - y3)/(x4 - x3): # if parallel lines
min_dist = np.min([point_to_line_segment(x1, y1, x3, y3, x4, y4),
point_to_line_segment(x2, y2, x3, y3, x4, y4),
point_to_line_segment(x3, y3, x1, y1, x2, y2),
point_to_line_segment(x4, y4, x1, y1, x2, y2)])
return(min_dist)
else:
x_int = (((x1 * y2 - y1 * x2) * (x3 - x4) - (x1 - x2) * (x3*y4 - y3 * x4)) /
(x1 - x2) * (y3 - y4) - (y1 - y2) * (x3 - x4))
y_int = (((x1 * y2 - y1 * x2) * (y3 - y4) - (y1 - y2) * (x3 * y4 - y3 * x4))/
((x1 - x2) * (y3 - y4) - (y1 - y2) * (x3 - x4)))
# check intersection
if (x_int >= np.min([x1, x2]) and x_int <= np.max([x1, x2]) and
x_int >= np.min([x3, x4]) and x_int <= np.max([x3, x4]) and
y_int >= np.min([y1, y2]) and y_int <= np.max([y1, y2]) and
y_int >= np.min([y3, y4]) and y_int <= np.max([y3, y4])):
return(0.0)
else:
min_dist = np.min([point_to_line_segment(x1, y1, x3, y3, x4, y4),
point_to_line_segment(x2, y2, x3, y3, x4, y4),
point_to_line_segment(x3, y3, x1, y1, x2, y2),
point_to_line_segment(x4, y4, x1, y1, x2, y2)])
return(min_dist)
segment_to_segment(0, 0, 0, 1, 1, 0, 1, 1) #vertical, adjacent lines
segment_to_segment(0, 0, 0, 1, 1, 2, 1, 3) #vertical, offset lines
segment_to_segment(-1, 0, 1, 0, 0, -1, 0, 1) # intersection
segment_to_segment(3e6, 3e6, 3e6+1, 3e6+1, 3e6, 3e6+1, 3e6+1, 3e6) # far from axis
segment_to_segment(-3e6, -3e6, -3e6+1, -3e6+1, -3e6, -3e6+1, -3e6+1, -3e6) # far from axis, negative
segment_to_segment(0,0, 1,1, 1,0, 2,1) # diagonal parallel
segment_to_segment(0,0, 1,1, 2,1, 3,2) # diagonal offset parallel
segment_to_segment(0,0, 1,1, 1,0, 2,0.9) # diagonal off-parallel
segment_to_segment(0,0, 1,3, 2,3, 3,0) # / \ shape
segment_to_segment(0,0, 0,3, 2,3, 3,0) # | \ shape
segment_to_segment(0,0, 0,2, 2,3, 3,0) # | \ shape
| Min distance between segments.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h3 style='color:blue' align='center'>Handling imbalanced data in customer churn prediction</h3>
# Customer churn prediction is to measure why customers are leaving a business. In this tutorial we will be looking at customer churn in telecom business. We will build a deep learning model to predict the churn and use precision,recall, f1-score to measure performance of our model.
# We will then handle imbalance in data using various techniques and improve f1-score
import pandas as pd
from matplotlib import pyplot as plt
import numpy as np
# %matplotlib inline
import warnings
warnings.filterwarnings('ignore')
# **Load the data**
df = pd.read_csv("customer_churn.csv")
df.sample(5)
df.Churn.value_counts()
517400/df.shape[0]
# **First of all, drop customerID column as it is of no use**
df.drop('customerID',axis='columns',inplace=True)
df.dtypes
# **Quick glance at above makes me realize that TotalCharges should be float but it is an object. Let's check what's going on with this column**
df.TotalCharges.values
# **Ahh... it is string. Lets convert it to numbers**
pd.to_numeric(df.TotalCharges,errors='coerce').isnull()
df[pd.to_numeric(df.TotalCharges,errors='coerce').isnull()]
df.shape
df.iloc[488].TotalCharges
df[df.TotalCharges!=' '].shape
# **Remove rows with space in TotalCharges**
df1 = df[df.TotalCharges!=' ']
df1.shape
df1.dtypes
df1.TotalCharges = pd.to_numeric(df1.TotalCharges)
df1.TotalCharges.values
df1[df1.Churn=='No']
# **Data Visualization**
# +
tenure_churn_no = df1[df1.Churn=='No'].tenure
tenure_churn_yes = df1[df1.Churn=='Yes'].tenure
plt.xlabel("tenure")
plt.ylabel("Number Of Customers")
plt.title("Customer Churn Prediction Visualiztion")
blood_sugar_men = [113, 85, 90, 150, 149, 88, 93, 115, 135, 80, 77, 82, 129]
blood_sugar_women = [67, 98, 89, 120, 133, 150, 84, 69, 89, 79, 120, 112, 100]
plt.hist([tenure_churn_yes, tenure_churn_no], rwidth=0.95, color=['green','red'],label=['Churn=Yes','Churn=No'])
plt.legend()
# +
mc_churn_no = df1[df1.Churn=='No'].MonthlyCharges
mc_churn_yes = df1[df1.Churn=='Yes'].MonthlyCharges
plt.xlabel("Monthly Charges")
plt.ylabel("Number Of Customers")
plt.title("Customer Churn Prediction Visualiztion")
blood_sugar_men = [113, 85, 90, 150, 149, 88, 93, 115, 135, 80, 77, 82, 129]
blood_sugar_women = [67, 98, 89, 120, 133, 150, 84, 69, 89, 79, 120, 112, 100]
plt.hist([mc_churn_yes, mc_churn_no], rwidth=0.95, color=['green','red'],label=['Churn=Yes','Churn=No'])
plt.legend()
# -
# **Many of the columns are yes, no etc. Let's print unique values in object columns to see data values**
def print_unique_col_values(df):
for column in df:
if df[column].dtypes=='object':
print(f'{column}: {df[column].unique()}')
print_unique_col_values(df1)
# **Some of the columns have no internet service or no phone service, that can be replaced with a simple No**
df1.replace('No internet service','No',inplace=True)
df1.replace('No phone service','No',inplace=True)
print_unique_col_values(df1)
# **Convert Yes and No to 1 or 0**
yes_no_columns = ['Partner','Dependents','PhoneService','MultipleLines','OnlineSecurity','OnlineBackup',
'DeviceProtection','TechSupport','StreamingTV','StreamingMovies','PaperlessBilling','Churn']
for col in yes_no_columns:
df1[col].replace({'Yes': 1,'No': 0},inplace=True)
for col in df1:
print(f'{col}: {df1[col].unique()}')
df1['gender'].replace({'Female':1,'Male':0},inplace=True)
df1.gender.unique()
# **One hot encoding for categorical columns**
df2 = pd.get_dummies(data=df1, columns=['InternetService','Contract','PaymentMethod'])
df2.columns
df2.sample(5)
df2.dtypes
# +
cols_to_scale = ['tenure','MonthlyCharges','TotalCharges']
from sklearn.preprocessing import MinMaxScaler
scaler = MinMaxScaler()
df2[cols_to_scale] = scaler.fit_transform(df2[cols_to_scale])
# -
for col in df2:
print(f'{col}: {df2[col].unique()}')
# **Train test split**
# +
X = df2.drop('Churn',axis='columns')
y = testLabels = df2.Churn.astype(np.float32)
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=15, stratify=y)
# -
y_train.value_counts()
y.value_counts()
5163/1869
y_test.value_counts()
X_train.shape
X_test.shape
X_train[:10]
len(X_train.columns)
# **Build a model (ANN) in tensorflow/keras**
from tensorflow_addons import losses
import tensorflow as tf
from tensorflow import keras
from sklearn.metrics import confusion_matrix , classification_report
def ANN(X_train, y_train, X_test, y_test, loss, weights):
model = keras.Sequential([
keras.layers.Dense(26, input_dim=26, activation='relu'),
keras.layers.Dense(15, activation='relu'),
keras.layers.Dense(1, activation='sigmoid')
])
model.compile(optimizer='adam', loss=loss, metrics=['accuracy'])
if weights == -1:
model.fit(X_train, y_train, epochs=100)
else:
model.fit(X_train, y_train, epochs=100, class_weight = weights)
print(model.evaluate(X_test, y_test))
y_preds = model.predict(X_test)
y_preds = np.round(y_preds)
print("Classification Report: \n", classification_report(y_test, y_preds))
return y_preds
y_preds = ANN(X_train, y_train, X_test, y_test, 'binary_crossentropy', -1)
# ## Mitigating Skewdness of Data
# ### Method 1: Undersampling
# reference: https://www.kaggle.com/rafjaa/resampling-strategies-for-imbalanced-datasets
# +
# Class count
count_class_0, count_class_1 = df1.Churn.value_counts()
# Divide by class
df_class_0 = df2[df2['Churn'] == 0]
df_class_1 = df2[df2['Churn'] == 1]
# +
# Undersample 0-class and concat the DataFrames of both class
df_class_0_under = df_class_0.sample(count_class_1)
df_test_under = pd.concat([df_class_0_under, df_class_1], axis=0)
print('Random under-sampling:')
print(df_test_under.Churn.value_counts())
# +
X = df_test_under.drop('Churn',axis='columns')
y = df_test_under['Churn']
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=15, stratify=y)
# -
# Number of classes in training Data
y_train.value_counts()
# **Printing Classification in the last, Scroll down till the last epoch to watch the classification report**
y_preds = ANN(X_train, y_train, X_test, y_test, 'binary_crossentropy', -1)
# Check classification report above. f1-score for minority class 1 improved from **0.57 to 0.76**. Score for class 0 reduced to 0.75 from 0.85 but that's ok. We have more generalized classifier which classifies both classes with similar prediction score
# ### Method2: Oversampling
# +
# Oversample 1-class and concat the DataFrames of both classes
df_class_1_over = df_class_1.sample(count_class_0, replace=True)
df_test_over = pd.concat([df_class_0, df_class_1_over], axis=0)
print('Random over-sampling:')
print(df_test_over.Churn.value_counts())
# +
X = df_test_over.drop('Churn',axis='columns')
y = df_test_over['Churn']
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=15, stratify=y)
# -
# Number of classes in training Data
y_train.value_counts()
loss = keras.losses.BinaryCrossentropy()
weights = -1
y_preds = ANN(X_train, y_train, X_test, y_test, 'binary_crossentropy', -1)
# Check classification report above. f1-score for minority class 1 improved from **0.57 to 0.76**. Score for class 0 reduced to 0.75 from 0.85 but that's ok. We have more generalized classifier which classifies both classes with similar prediction score
# ### Method3: SMOTE
# To install imbalanced-learn library use **pip install imbalanced-learn** command
X = df2.drop('Churn',axis='columns')
y = df2['Churn']
# +
from imblearn.over_sampling import SMOTE
smote = SMOTE(sampling_strategy='minority')
X_sm, y_sm = smote.fit_sample(X, y)
y_sm.value_counts()
# -
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X_sm, y_sm, test_size=0.2, random_state=15, stratify=y_sm)
# Number of classes in training Data
y_train.value_counts()
y_preds = ANN(X_train, y_train, X_test, y_test, 'binary_crossentropy', -1)
# SMOT Oversampling increases f1 score of minority class 1 from **0.57 to 0.81 (huge improvement)** Also over all accuracy improves from 0.78 to 0.80
# ### Method4: Use of Ensemble with undersampling
df2.Churn.value_counts()
# Regain Original features and labels
X = df2.drop('Churn',axis='columns')
y = df2['Churn']
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=15, stratify=y)
y_train.value_counts()
# model1 --> class1(1495) + class0(0, 1495)
#
# model2 --> class1(1495) + class0(1496, 2990)
#
# model3 --> class1(1495) + class0(2990, 4130)
df3 = X_train.copy()
df3['Churn'] = y_train
df3.head()
df3_class0 = df3[df3.Churn==0]
df3_class1 = df3[df3.Churn==1]
def get_train_batch(df_majority, df_minority, start, end):
df_train = pd.concat([df_majority[start:end], df_minority], axis=0)
X_train = df_train.drop('Churn', axis='columns')
y_train = df_train.Churn
return X_train, y_train
# +
X_train, y_train = get_train_batch(df3_class0, df3_class1, 0, 1495)
y_pred1 = ANN(X_train, y_train, X_test, y_test, 'binary_crossentropy', -1)
# +
X_train, y_train = get_train_batch(df3_class0, df3_class1, 1495, 2990)
y_pred2 = ANN(X_train, y_train, X_test, y_test, 'binary_crossentropy', -1)
# +
X_train, y_train = get_train_batch(df3_class0, df3_class1, 2990, 4130)
y_pred3 = ANN(X_train, y_train, X_test, y_test, 'binary_crossentropy', -1)
# -
len(y_pred1)
y_pred_final = y_pred1.copy()
for i in range(len(y_pred1)):
n_ones = y_pred1[i] + y_pred2[i] + y_pred3[i]
if n_ones>1:
y_pred_final[i] = 1
else:
y_pred_final[i] = 0
cl_rep = classification_report(y_test, y_pred_final)
print(cl_rep)
# f1-score for minority class 1 improved to 0.62 from 0.57. The score for majority class 0 is suffering and reduced to 0.80 from 0.85 but at least there is some balance in terms of prediction accuracy across two classes
| Program's_Contributed_By_Contributors/AI-Summer-Course/py-master/DeepLearningML/14_imbalanced/handling_imbalanced_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ## VAE-Experiment-3:
#
#
# #### Variational Autoencoder
# Variational Autoencoders (VAE) are generative models which use deep feed-forward neural networks to model complicated probability density functions $p(x)$. As discussed in the first notebook, a VAE models a complicated probability distribution as a complex deterministic transformation of a simple probability distribution. A $VAE$ model can be trained to fit a data distribution by maximizing the log-likelihood of the samples from the distribution. Specifically, as discussed in the second notebook, VAE actually maximizes a variational lower-bound to the log-likelihood. It simultaneously learns an encoder network in order to estimate the variational lower-bound during training.
#
# #### Let's learn a simple fully-connected Variational Autoencoder that generates digits similar to the ones in the MNIST dataset.
# +
##########################
# Import necessary modules
##########################
import pickle
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
from matplotlib import gridspec
import warnings
warnings.filterwarnings('ignore')
# %matplotlib inline
import torch
from torch.autograd import Variable
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
# +
################
# Set parameters
################
cuda = torch.cuda.is_available()
kwargs = {'num_workers': 1, 'pin_memory': True} if cuda else {}
seed = 1
n_classes = 10
z_dim = 2
X_dim = 784
train_batch_size = 100
valid_batch_size = train_batch_size
N = 1000
epochs = 5
params = {}
params['cuda'] = cuda
params['n_classes'] = n_classes
params['z_dim'] = z_dim
params['X_dim'] = X_dim
params['train_batch_size'] = train_batch_size
params['valid_batch_size'] = valid_batch_size
params['N'] = N
params['epochs'] = epochs
# +
###################################
# Load data and create Data loaders
###################################
def load_data(data_path='./data/VAE/processed_data/'):
print('loading data!')
trainset_labeled = pickle.load(open(data_path + "train_labeled.p", "rb"))
trainset_unlabeled = pickle.load(open(data_path + "train_unlabeled.p", "rb"))
# Set -1 as labels for unlabeled data
trainset_unlabeled.train_labels = torch.from_numpy(np.array([-1] * 47000))
validset = pickle.load(open(data_path + "validation.p", "rb"))
train_labeled_loader = torch.utils.data.DataLoader(trainset_labeled,
batch_size=train_batch_size,
shuffle=True, **kwargs)
train_unlabeled_loader = torch.utils.data.DataLoader(trainset_unlabeled,
batch_size=train_batch_size,
shuffle=True, **kwargs)
valid_loader = torch.utils.data.DataLoader(validset, batch_size=valid_batch_size, shuffle=True)
return train_labeled_loader, train_unlabeled_loader, valid_loader
# +
#################
# Define Networks
#################
# Encoder
class Q_net(nn.Module):
def __init__(self):
super(Q_net, self).__init__()
self.lin1 = nn.Linear(X_dim, N)
self.lin2 = nn.Linear(N, N)
# Gaussian code (z)
self.lin3gauss_mean = nn.Linear(N, z_dim)
self.lin3gauss_logvar = nn.Linear(N, z_dim)
def forward(self, x):
x = F.dropout(self.lin1(x), p=0.2, training=self.training)
x = F.relu(x)
x = F.dropout(self.lin2(x), p=0.2, training=self.training)
x = F.relu(x)
xgauss_mean = self.lin3gauss_mean(x)
xgauss_logvar = self.lin3gauss_logvar(x)
return xgauss_mean, xgauss_logvar
# Decoder
class P_net(nn.Module):
def __init__(self):
super(P_net, self).__init__()
self.lin1 = nn.Linear(z_dim, N)
self.lin2 = nn.Linear(N, N)
self.lin3 = nn.Linear(N, X_dim)
def forward(self, x):
x = self.lin1(x)
x = F.dropout(x, p=0.2, training=self.training)
x = F.relu(x)
x = self.lin2(x)
x = F.dropout(x, p=0.2, training=self.training)
x = self.lin3(x)
return F.sigmoid(x)
# +
###################
# Utility functions
###################
def save_model(model, filename):
print('Best model so far, saving it...')
torch.save(model.state_dict(), filename)
def report_loss(epoch, recon_loss):
'''
Print loss
'''
print('Epoch-{}; recon_loss: {:.4}'.format(epoch,recon_loss.data[0]))
def create_latent(Q, loader):
'''
Creates the latent representation for the samples in loader
return:
z_values: numpy array with the latent representations
labels: the labels corresponding to the latent representations
'''
Q.eval()
labels = []
for batch_idx, (X, target) in enumerate(loader):
X = X * 0.3081 + 0.1307
# X.resize_(loader.batch_size, X_dim)
X, target = Variable(X), Variable(target)
labels.extend(target.data.tolist())
if cuda:
X, target = X.cuda(), target.cuda()
# Reconstruction phase
z_sample = Q(X)
if batch_idx > 0:
z_values = np.concatenate((z_values, np.array(z_sample.data.tolist())))
else:
z_values = np.array(z_sample.data.tolist())
labels = np.array(labels)
return z_values, labels
def get_X_batch(data_loader, params, size=None):
if size is None:
size = data_loader.batch_size
data_loader.batch_size = size
for X, target in data_loader:
break
train_batch_size = params['train_batch_size']
X_dim = params['X_dim']
cuda = params['cuda']
X = X * 0.3081 + 0.1307
X = X[:size]
target = target[:size]
X.resize_(size, X_dim)
X, target = Variable(X), Variable(target)
if cuda:
X, target = X.cuda(), target.cuda()
return X, target
# -
# The VAE objective function and the reparameterization trick that allows us to back-propagate gradient through the sampling process in the latent layer.
# +
####################
# Training procedure
####################
def train(P, Q, P_decoder, Q_encoder, data_loader):
'''
Train procedure for one epoch.
'''
TINY = 1e-15
# Set the networks in train mode (apply dropout when needed)
Q.train()
P.train()
# Loop through the labeled and unlabeled dataset getting one batch of samples from each
# The batch size has to be a divisor of the size of the dataset or it will return
# invalid samples
for X, target in data_loader:
# Load batch and normalize samples to be between 0 and 1
X = X * 0.3081 + 0.1307
X.resize_(train_batch_size, X_dim)
X, target = Variable(X), Variable(target)
if cuda:
X, target = X.cuda(), target.cuda()
# Init gradients
P.zero_grad()
Q.zero_grad()
# Reconstruction phase
z_mean, z_logvar = Q(X)
std = z_logvar.mul(0.5).exp_()
if cuda:
eps = torch.cuda.FloatTensor(std.size()).normal_()
else:
eps = torch.FloatTensor(std.size()).normal_()
eps = Variable(eps)
z_sample = eps.mul(std).add_(z_mean)
X_sample = P(z_sample)
criterion = nn.BCELoss()
criterion.size_average = False
recon_loss = criterion(X_sample, X.resize(train_batch_size, X_dim))
# -0.5 * sum(1 + log(sigma^2) - mu^2 - sigma^2)
KLD_element = z_mean.pow(2).add_(z_logvar.exp()).mul_(-1).add_(1).add_(z_logvar)
KLD_loss = torch.sum(KLD_element).mul_(-0.5)
loss = recon_loss + KLD_loss
loss.backward()
P_decoder.step()
Q_encoder.step()
P.zero_grad()
Q.zero_grad()
return recon_loss
def generate_model(train_labeled_loader, train_unlabeled_loader, valid_loader):
torch.manual_seed(10)
if cuda:
Q = Q_net().cuda()
P = P_net().cuda()
else:
Q = Q_net()
P = P_net()
# Set learning rates
gen_lr = 0.0005
# Set optimizators
P_decoder = optim.Adam(P.parameters(), lr=gen_lr)
Q_encoder = optim.Adam(Q.parameters(), lr=gen_lr)
for epoch in range(epochs):
recon_loss = train(P, Q, P_decoder, Q_encoder,
train_unlabeled_loader)
if epoch % 1 == 0:
report_loss(epoch, recon_loss)
return Q, P
# +
##########################
# Train a generative model
##########################
train_labeled_loader, train_unlabeled_loader, valid_loader = load_data()
Q, P = generate_model(train_labeled_loader, train_unlabeled_loader, valid_loader)
# +
####################
# Save trained model
####################
# Save trained model
torch.save(Q,'./data/VAE/TrainedModels/VAE_mytraining_Q.pt')
torch.save(P,'./data/VAE/TrainedModels/VAE_mytraining_P.pt')
# +
####################
# Load trained model
####################
# Load model trained for 200 epochs
Q_pt = torch.load('./data/VAE/TrainedModels/VAE_preTrained_Q.pt')
P_pt = torch.load('./data/VAE/TrainedModels/VAE_preTrained_P.pt')
# +
##########################
# Visualize reconstruction
##########################
def create_reconstruction(Q, P, data_loader, params):
Q.eval()
P.eval()
X, label = get_X_batch(data_loader, params, size=1)
## Sampling from latent distribution
z_mean, z_logvar = Q(X)
std = z_logvar.mul(0.5).exp_()
if cuda:
eps = torch.cuda.FloatTensor(std.size()).normal_()
else:
eps = torch.FloatTensor(std.size()).normal_()
eps = Variable(eps)
z_sample = eps.mul(std).add_(z_mean)
## Forwarding the mean of the latent distribution
#z_mean, z_logvar = Q(X)
#z_sample = z_mean
x = P(z_sample)
img_orig = np.array(X[0].data.tolist()).reshape(28, 28)
img_rec = np.array(x[0].data.tolist()).reshape(28, 28)
plt.subplot(1, 2, 1)
plt.imshow(img_orig)
plt.subplot(1, 2, 2)
plt.imshow(img_rec)
data_loader = valid_loader # Training data: train_unlabeled_loader | Validation data: valid_loader
create_reconstruction(Q_pt, P_pt, data_loader, params)
plt.show()
# +
######################
# Visualize generation
######################
def grid_plot2d(Q, P, params):
Q.eval()
P.eval()
cuda = params['cuda']
z1 = Variable(torch.from_numpy(np.arange(-1, 1, 0.15).astype('float32')))
z2 = Variable(torch.from_numpy(np.arange(-1, 1, 0.15).astype('float32')))
if cuda:
z1, z2 = z1.cuda(), z2.cuda()
nx, ny = len(z1), len(z2)
plt.subplot()
gs = gridspec.GridSpec(nx, ny, hspace=0.05, wspace=0.05)
for i, g in enumerate(gs):
z = torch.cat((z1[i / ny], z2[i % nx])).resize(1, 2)
x = P(z)
ax = plt.subplot(g)
img = np.array(x.data.tolist()).reshape(28, 28)
ax.imshow(img, )
ax.set_xticks([])
ax.set_yticks([])
ax.set_aspect('auto')
grid_plot2d(Q_pt, P_pt, params)
plt.show()
# -
# ### C-shaped distribution
#
# Now, remember the C-shaped distribution from the first notebook? Here it is again, Can you learn a Variational Autoencoder that models this distribution.
#
# +
#####################################
# Reset workspace for next experiment
#####################################
# %reset -f
# +
##########################
# Import necessary modules
##########################
import pickle
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
from matplotlib import gridspec
import warnings
warnings.filterwarnings('ignore')
import torch
from torch.autograd import Variable
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
# +
################
# Set parameters
################
cuda = True
kwargs = {'num_workers': 1, 'pin_memory': True} if cuda else {}
seed = 1
z_dim = 2
X_dim = 2
train_batch_size = 100
valid_batch_size = train_batch_size
N = 100
epochs = 100
params = {}
params['cuda'] = cuda
params['z_dim'] = z_dim
params['X_dim'] = X_dim
params['train_batch_size'] = train_batch_size
params['valid_batch_size'] = valid_batch_size
params['N'] = N
params['epochs'] = epochs
# +
####################
# Define Data-loader
####################
def data_loader_Cdist(numSamples):
z = np.random.randn(numSamples,2).astype(np.float32) # Sample from Gaussian distribution
z1 = z[:,0]
z2 = z[:,1]
os = 10
ost = np.pi/2
x1 = -(1.5*os+z1)*(np.sin(z2+ost))
x2 = (os+z1)*np.cos(z2+ost)
x = np.concatenate([np.expand_dims(x1,1),np.expand_dims(x2,1)],1)
dummy = np.zeros(len(x))
return torch.from_numpy(x), torch.from_numpy(dummy)
# +
#################
# Define Networks
#################
# Encoder
class Q_net(nn.Module):
def __init__(self):
super(Q_net, self).__init__()
self.lin1 = nn.Linear(X_dim, N)
self.lin2 = nn.Linear(N, N)
# Gaussian code (z)
self.lin3gauss_mean = nn.Linear(N, z_dim)
self.lin3gauss_logvar = nn.Linear(N, z_dim)
def forward(self, x):
x = self.lin1(x)
x = F.relu(x)
x = self.lin2(x)
x = F.relu(x)
xgauss_mean = self.lin3gauss_mean(x)
xgauss_logvar = self.lin3gauss_logvar(x)
return xgauss_mean, xgauss_logvar
# Decoder
class P_net(nn.Module):
def __init__(self):
super(P_net, self).__init__()
self.lin1 = nn.Linear(z_dim, N)
self.lin2 = nn.Linear(N, N)
self.lin3 = nn.Linear(N, X_dim)
def forward(self, x):
x = self.lin1(x)
x = F.relu(x)
x = self.lin2(x)
x = F.relu(x)
x = self.lin3(x)
return x
# +
###################
# Utility functions
###################
def save_model(model, filename):
print('Best model so far, saving it...')
torch.save(model.state_dict(), filename)
def report_loss(epoch, recon_loss):
'''
Print loss
'''
print('Epoch-{}; recon_loss: {:.4}'.format(epoch,recon_loss.data[0]))
# +
####################
# Training procedure
####################
def train(P, Q, P_decoder, Q_encoder, data_loader):
'''
Train procedure for one epoch.
'''
TINY = 1e-15
# Set the networks in train mode (apply dropout when needed)
Q.train()
P.train()
# Loop through the labeled and unlabeled dataset getting one batch of samples from each
# The batch size has to be a divisor of the size of the dataset or it will return
# invalid samples
for it in range(100):
X, target = data_loader(train_batch_size)
# Load batch and normalize samples to be 0-mean and unit-std
X = (X + 4.55)/ 8.08
X.resize_(train_batch_size, X_dim)
X, target = Variable(X), Variable(target)
if cuda:
X, target = X.cuda(), target.cuda()
# Init gradients
P.zero_grad()
Q.zero_grad()
# Reconstruction phase
z_mean, z_logvar = Q(X)
std = z_logvar.mul(0.5).exp_()
if cuda:
eps = torch.cuda.FloatTensor(std.size()).normal_()
else:
eps = torch.FloatTensor(std.size()).normal_()
eps = Variable(eps)
z_sample = eps.mul(std).add_(z_mean)
X_sample = P(z_sample)
criterion = nn.MSELoss()
criterion.size_average = False
recon_loss = criterion(X_sample, X.resize(train_batch_size, X_dim))
# -0.5 * sum(1 + log(sigma^2) - mu^2 - sigma^2)
KLD_element = z_mean.pow(2).add_(z_logvar.exp()).mul_(-1).add_(1).add_(z_logvar)
KLD_loss = torch.sum(KLD_element).mul_(-0.5)
loss = recon_loss + KLD_loss
loss.backward()
P_decoder.step()
Q_encoder.step()
P.zero_grad()
Q.zero_grad()
return recon_loss
def generate_model(train_labeled_loader, train_unlabeled_loader, valid_loader):
torch.manual_seed(10)
if cuda:
Q = Q_net().cuda()
P = P_net().cuda()
else:
Q = Q_net()
P = P_net()
# Set learning rates
gen_lr = 0.0001
# Set optimizators
P_decoder = optim.Adam(P.parameters(), lr=gen_lr)
Q_encoder = optim.Adam(Q.parameters(), lr=gen_lr)
for epoch in range(epochs):
#x,dummy = train_unlabeled_loader(train_batch_size)
recon_loss = train(P, Q, P_decoder, Q_encoder,
train_unlabeled_loader)
if epoch % 10 == 0:
report_loss(epoch, recon_loss)
return Q, P
# +
##########################
# Train a generative model
##########################
Q, P = generate_model(data_loader_Cdist, data_loader_Cdist, data_loader_Cdist)
# +
##################################
# Visualize generated distribution
##################################
def generate_Cdist(numSamples):
z = np.random.randn(numSamples,2).astype(np.float32) # Sample from Gaussian distribution
z = Variable(torch.from_numpy(z))
if cuda:
z = z.cuda()
x = P(z)
return x
x = generate_Cdist(5000)
x = np.array(x.data.tolist())
x = x*8.08 - 4.55
x1 = x[:,0]
x2 = x[:,1]
plt.scatter(x1,x2)
plt.hold()
plt.grid(True)
plt.show()
# -
# The VAE seems to have done a near-decent job at modelling the C-shaped distribution.
# ### Exercise
#
# 1. In function 'grid_plot2d()', we visualize the ability of the decoder to generate MNIST digits when as we change samples from the latent distribution. Play with it by varying its range and visualizing the outputs. You can use the pretrained decoder model for this. What do you observe?
#
# 2. In the MNIST example, plot the reconstruction loss and the KL Divergence, as training progresses. How many epochs are required to get reasonable results?
#
# 3. In the C-shaped distribution, tune the VAE so that it models the distribution even better.
# 4. VAEs have a problem that the generations are not sharp enough, although meaningful. Can you come up with an idea to make them sharper? Implement it!
#
| VAE-Lab3-VAE.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: all
# language: python
# name: all
# ---
# +
# %pylab inline
import time
import pandas as pd
import pysumma as ps
import xarray as xr
from matplotlib import cm
import itertools
import seaborn as sns
from pathlib import Path
import matplotlib.dates as mdates
from matplotlib.patches import Patch
from matplotlib.lines import Line2D
from matplotlib.collections import LineCollection
from matplotlib.colors import ListedColormap, BoundaryNorm
from pysumma.plotting.utils import justify
sns.set_context('poster')
mpl.style.use('seaborn-bright')
mpl.rcParams['figure.figsize'] = (18, 12)
color_1l = 'lightseagreen'
#colors_2l = ['wheat', 'orange', 'peru']
colors_2l = ['orange', 'chocolate', 'saddlebrown']
colors_3l = ['dodgerblue', 'royalblue', 'navy']
colors_4l = ['violet', 'deeppink', 'crimson']
dt_list = ['+0.0K', '+2.0K', '+4.0K']
model_list = ['JRDN', 'CLM', '2L_thin', '2L_mid', '2L_thick', '3L_thin', '3L_mid', '3L_thick', '4L_thin', '4L_mid', '4L_thick']
site_list = ['Dana', 'Reynolds_Creek', 'Col-de-Port']
# +
dana_obs = pd.read_csv('../obs_data/dana_obs.csv',
index_col=0, skipinitialspace=True, parse_dates=True)
cdp_obs = xr.open_dataset('../obs_data/coldeport_obs.nc')
reynolds_obs = xr.open_dataset('../obs_data/reynolds_obs.nc')
dana = xr.open_dataset('../processed/dana_perturbations.nc')
dana['time'] = dana.time.dt.round('H')
coldeport = xr.open_dataset('../processed/coldeport_perturbations.nc')
coldeport['time'] = coldeport.time.dt.round('H')
reynolds = xr.open_dataset('../processed/reynolds_perturbations.nc')
reynolds['time'] = reynolds.time.dt.round('H')
# +
dana_ts = slice('2009-10-01', '2010-08-01')
dana_swe = dana['scalarSWE'].sel(dt='+0.0K', time=dana_ts).resample(time='D').mean()
reynolds_ts = slice('2003-11-01', '2004-06-01')
reynolds_swe = reynolds['scalarSWE'].sel(dt='+0.0K', time=reynolds_ts).resample(time='D').mean()
coldeport_ts = slice('2000-10-01', '2001-07-01')
coldeport_swe = coldeport['scalarSWE'].sel(dt='+0.0K', time=coldeport_ts).resample(time='D').mean()
dana_mintemp = dana['airtemp'].sel(dt='+0.0K', time=dana_ts).resample(time='D').min() - 273.16
dana_maxtemp = dana['airtemp'].sel(dt='+0.0K', time=dana_ts).resample(time='D').max() - 273.16
dana_depth = dana['scalarSnowDepth'].sel(dt='+0.0K', time=dana_ts).resample(time='D').mean()
dana_snotemp = dana['scalarSnowTemp'].sel(dt='+0.0K', time=dana_ts).resample(time='D').mean() - 273.16
coldeport_mintemp = coldeport['airtemp'].sel(dt='+0.0K', time=coldeport_ts).resample(time='D').min() - 273.16
coldeport_maxtemp = coldeport['airtemp'].sel(dt='+0.0K', time=coldeport_ts).resample(time='D').max() - 273.16
coldeport_depth = coldeport['scalarSnowDepth'].sel(dt='+0.0K', time=coldeport_ts).resample(time='D').mean()
coldeport_snotemp = coldeport['scalarSnowTemp'].sel(dt='+0.0K', time=coldeport_ts).resample(time='D').mean() - 273.16
reynolds_mintemp = reynolds['airtemp'].sel(dt='+0.0K', time=reynolds_ts).resample(time='D').min() - 273.16
reynolds_maxtemp = reynolds['airtemp'].sel(dt='+0.0K', time=reynolds_ts).resample(time='D').max() - 273.16
reynolds_depth = reynolds['scalarSnowDepth'].sel(dt='+0.0K', time=reynolds_ts).resample(time='D').mean()
reynolds_snotemp = reynolds['scalarSnowTemp'].sel(dt='+0.0K', time=reynolds_ts).resample(time='D').mean()- 273.16
# +
mpl.rcParams['figure.figsize'] = (24, 15)
sizes = {'height_ratios': [3, 3, 3 ,1]}
fig, axes = plt.subplots(4, 3, gridspec_kw=sizes, sharey='row', sharex='col')
# Plotting for Dana Meadows
axes[3, 0].fill_between(dana_mintemp.time.values, dana_mintemp, dana_maxtemp, alpha=0.5)
axes[3, 0].axhline(0, color='black')
axes[3, 0].set_ylabel('Daily air \n temperature \n range (C)')
dana_snotemp.sel(model='JRDN' ).plot(color='black', label='JRDN', ax=axes[2, 0], linewidth=4);
dana_snotemp.sel(model='CLM' ).plot(color='lime', label='CLM', ax=axes[2, 0], linewidth=4);
dana_snotemp.sel(model='2L_thin' ).plot(color=colors_2l[0], label='2l_thin', ax=axes[2, 0], linewidth=1.5);
dana_snotemp.sel(model='2L_mid' ).plot(color=colors_2l[1], label='2l_mid', ax=axes[2, 0], linewidth=1.5);
dana_snotemp.sel(model='2L_thick').plot(color=colors_2l[2], label='2l_thick', ax=axes[2, 0], linewidth=1.5);
dana_snotemp.sel(model='3L_thin' ).plot(color=colors_3l[0], label='3l_thin', ax=axes[2, 0], linewidth=1.5);
dana_snotemp.sel(model='3L_mid' ).plot(color=colors_3l[1], label='3l_mid', ax=axes[2, 0], linewidth=1.5);
dana_snotemp.sel(model='3L_thick').plot(color=colors_3l[2], label='3l_thick', ax=axes[2, 0], linewidth=1.5);
dana_snotemp.sel(model='4L_thin' ).plot(color=colors_4l[0], label='4l_thin', ax=axes[2, 0], linewidth=1.5);
dana_snotemp.sel(model='4L_mid' ).plot(color=colors_4l[1], label='4l_mid', ax=axes[2, 0], linewidth=1.5);
dana_snotemp.sel(model='4L_thick').plot(color=colors_4l[2], label='4l_thick', ax=axes[2, 0], linewidth=1.5);
axes[2, 0].set_ylabel('Mean snowpack \n temperature (C)')
dana_depth.sel(model='JRDN' ).plot(color='black', label='JRDN', ax=axes[1, 0], linewidth=4);
dana_depth.sel(model='CLM' ).plot(color='lime', label='CLM', ax=axes[1, 0], linewidth=4);
dana_depth.sel(model='2L_thin' ).plot(color=colors_2l[0], label='2l_thin', ax=axes[1, 0], linewidth=1.5);
dana_depth.sel(model='2L_mid' ).plot(color=colors_2l[1], label='2l_mid', ax=axes[1, 0], linewidth=1.5);
dana_depth.sel(model='2L_thick').plot(color=colors_2l[2], label='2l_thick', ax=axes[1, 0], linewidth=1.5);
dana_depth.sel(model='3L_thin' ).plot(color=colors_3l[0], label='3l_thin', ax=axes[1, 0], linewidth=1.5);
dana_depth.sel(model='3L_mid' ).plot(color=colors_3l[1], label='3l_mid', ax=axes[1, 0], linewidth=1.5);
dana_depth.sel(model='3L_thick').plot(color=colors_3l[2], label='3l_thick', ax=axes[1, 0], linewidth=1.5);
dana_depth.sel(model='4L_thin' ).plot(color=colors_4l[0], label='4l_thin', ax=axes[1, 0], linewidth=1.5);
dana_depth.sel(model='4L_mid' ).plot(color=colors_4l[1], label='4l_mid', ax=axes[1, 0], linewidth=1.5);
dana_depth.sel(model='4L_thick').plot(color=colors_4l[2], label='4l_thick', ax=axes[1, 0], linewidth=1.5);
axes[1, 0].set_ylabel('Snow depth (m)')
dana_swe.sel(model='JRDN' ).plot(color='black', label='JRDN', ax=axes[0, 0], linewidth=4);
dana_swe.sel(model='CLM' ).plot(color='lime', label='CLM', ax=axes[0, 0], linewidth=4);
dana_swe.sel(model='2L_thin' ).plot(color=colors_2l[0], label='2l_thin', ax=axes[0, 0], linewidth=1.5);
dana_swe.sel(model='2L_mid' ).plot(color=colors_2l[1], label='2l_mid', ax=axes[0, 0], linewidth=1.5);
dana_swe.sel(model='2L_thick').plot(color=colors_2l[2], label='2l_thick', ax=axes[0, 0], linewidth=1.5);
dana_swe.sel(model='3L_thin' ).plot(color=colors_3l[0], label='3l_thin', ax=axes[0, 0], linewidth=1.5);
dana_swe.sel(model='3L_mid' ).plot(color=colors_3l[1], label='3l_mid', ax=axes[0, 0], linewidth=1.5);
dana_swe.sel(model='3L_thick').plot(color=colors_3l[2], label='3l_thick', ax=axes[0, 0], linewidth=1.5);
dana_swe.sel(model='4L_thin' ).plot(color=colors_4l[0], label='4l_thin', ax=axes[0, 0], linewidth=1.5);
dana_swe.sel(model='4L_mid' ).plot(color=colors_4l[1], label='4l_mid', ax=axes[0, 0], linewidth=1.5);
dana_swe.sel(model='4L_thick').plot(color=colors_4l[2], label='4l_thick', ax=axes[0, 0], linewidth=1.5);
axes[0, 0].set_ylabel('SWE (mm)')
axes[0, 0].set_xlabel('')
axes[0, 0].set_title('Dana Meadows')
# Plotting for Col-de-Port
axes[3, 1].fill_between(coldeport_mintemp.time.values, coldeport_mintemp, coldeport_maxtemp, alpha=0.5)
axes[3, 1].axhline(0, color='black')
coldeport_snotemp.sel(model='JRDN' ).plot(color='black', label='JRDN', ax=axes[2, 1], linewidth=4);
coldeport_snotemp.sel(model='CLM' ).plot(color='lime', label='CLM', ax=axes[2, 1], linewidth=4);
coldeport_snotemp.sel(model='2L_thin' ).plot(color=colors_2l[0], label='2l_thin', ax=axes[2, 1], linewidth=1.5);
coldeport_snotemp.sel(model='2L_mid' ).plot(color=colors_2l[1], label='2l_mid', ax=axes[2, 1], linewidth=1.5);
coldeport_snotemp.sel(model='2L_thick').plot(color=colors_2l[2], label='2l_thick', ax=axes[2, 1], linewidth=1.5);
coldeport_snotemp.sel(model='3L_thin' ).plot(color=colors_3l[0], label='3l_thin', ax=axes[2, 1], linewidth=1.5);
coldeport_snotemp.sel(model='3L_mid' ).plot(color=colors_3l[1], label='3l_mid', ax=axes[2, 1], linewidth=1.5);
coldeport_snotemp.sel(model='3L_thick').plot(color=colors_3l[2], label='3l_thick', ax=axes[2, 1], linewidth=1.5);
coldeport_snotemp.sel(model='4L_thin' ).plot(color=colors_4l[0], label='4l_thin', ax=axes[2, 1], linewidth=1.5);
coldeport_snotemp.sel(model='4L_mid' ).plot(color=colors_4l[1], label='4l_mid', ax=axes[2, 1], linewidth=1.5);
coldeport_snotemp.sel(model='4L_thick').plot(color=colors_4l[2], label='4l_thick', ax=axes[2, 1], linewidth=1.5);
coldeport_depth.sel(model='JRDN' ).plot(color='black', label='JRDN', ax=axes[1, 1], linewidth=4);
coldeport_depth.sel(model='CLM' ).plot(color='lime', label='CLM', ax=axes[1, 1], linewidth=4);
coldeport_depth.sel(model='2L_thin' ).plot(color=colors_2l[0], label='2l_thin', ax=axes[1, 1], linewidth=1.5);
coldeport_depth.sel(model='2L_mid' ).plot(color=colors_2l[1], label='2l_mid', ax=axes[1, 1], linewidth=1.5);
coldeport_depth.sel(model='2L_thick').plot(color=colors_2l[2], label='2l_thick', ax=axes[1, 1], linewidth=1.5);
coldeport_depth.sel(model='3L_thin' ).plot(color=colors_3l[0], label='3l_thin', ax=axes[1, 1], linewidth=1.5);
coldeport_depth.sel(model='3L_mid' ).plot(color=colors_3l[1], label='3l_mid', ax=axes[1, 1], linewidth=1.5);
coldeport_depth.sel(model='3L_thick').plot(color=colors_3l[2], label='3l_thick', ax=axes[1, 1], linewidth=1.5);
coldeport_depth.sel(model='4L_thin' ).plot(color=colors_4l[0], label='4l_thin', ax=axes[1, 1], linewidth=1.5);
coldeport_depth.sel(model='4L_mid' ).plot(color=colors_4l[1], label='4l_mid', ax=axes[1, 1], linewidth=1.5);
coldeport_depth.sel(model='4L_thick').plot(color=colors_4l[2], label='4l_thick', ax=axes[1, 1], linewidth=1.5);
coldeport_swe.sel(model='JRDN' ).plot(color='black', label='JRDN', ax=axes[0, 1], linewidth=4);
coldeport_swe.sel(model='CLM' ).plot(color='lime', label='CLM', ax=axes[0, 1], linewidth=4);
coldeport_swe.sel(model='2L_thin' ).plot(color=colors_2l[0], label='2l_thin', ax=axes[0, 1], linewidth=1.5);
coldeport_swe.sel(model='2L_mid' ).plot(color=colors_2l[1], label='2l_mid', ax=axes[0, 1], linewidth=1.5);
coldeport_swe.sel(model='2L_thick').plot(color=colors_2l[2], label='2l_thick', ax=axes[0, 1], linewidth=1.5);
coldeport_swe.sel(model='3L_thin' ).plot(color=colors_3l[0], label='3l_thin', ax=axes[0, 1], linewidth=1.5);
coldeport_swe.sel(model='3L_mid' ).plot(color=colors_3l[1], label='3l_mid', ax=axes[0, 1], linewidth=1.5);
coldeport_swe.sel(model='3L_thick').plot(color=colors_3l[2], label='3l_thick', ax=axes[0, 1], linewidth=1.5);
coldeport_swe.sel(model='4L_thin' ).plot(color=colors_4l[0], label='4l_thin', ax=axes[0, 1], linewidth=1.5);
coldeport_swe.sel(model='4L_mid' ).plot(color=colors_4l[1], label='4l_mid', ax=axes[0, 1], linewidth=1.5);
coldeport_swe.sel(model='4L_thick').plot(color=colors_4l[2], label='4l_thick', ax=axes[0, 1], linewidth=1.5);
axes[0, 1].set_ylabel('')
axes[1, 1].set_ylabel('')
axes[0, 1].set_xlabel('')
axes[0, 1].set_title('Col de Porte')
# Plotting for Reynolds Creek
axes[3, 2].fill_between(reynolds_mintemp.time.values, reynolds_mintemp, reynolds_maxtemp, alpha=0.5)
axes[3, 2].axhline(0, color='black')
reynolds_snotemp.sel(model='JRDN' ).plot(color='black', label='SNTHERM-like', ax=axes[2, 2], linewidth=4);
reynolds_snotemp.sel(model='CLM' ).plot(color='lime', label='CLM-like', ax=axes[2, 2], linewidth=4);
reynolds_snotemp.sel(model='2L_thin' ).plot(color=colors_2l[0], label='2l_thin', ax=axes[2, 2], linewidth=1.5);
reynolds_snotemp.sel(model='2L_mid' ).plot(color=colors_2l[1], label='2l_mid', ax=axes[2, 2], linewidth=1.5);
reynolds_snotemp.sel(model='2L_thick').plot(color=colors_2l[2], label='2l_thick', ax=axes[2, 2], linewidth=1.5);
reynolds_snotemp.sel(model='3L_thin' ).plot(color=colors_3l[0], label='3l_thin', ax=axes[2, 2], linewidth=1.5);
reynolds_snotemp.sel(model='3L_mid' ).plot(color=colors_3l[1], label='3l_mid', ax=axes[2, 2], linewidth=1.5);
reynolds_snotemp.sel(model='3L_thick').plot(color=colors_3l[2], label='3l_thick', ax=axes[2, 2], linewidth=1.5);
reynolds_snotemp.sel(model='4L_thin' ).plot(color=colors_4l[0], label='4l_thin', ax=axes[2, 2], linewidth=1.5);
reynolds_snotemp.sel(model='4L_mid' ).plot(color=colors_4l[1], label='4l_mid', ax=axes[2, 2], linewidth=1.5);
reynolds_snotemp.sel(model='4L_thick').plot(color=colors_4l[2], label='4l_thick', ax=axes[2, 2], linewidth=1.5);
reynolds_depth.sel(model='JRDN' ).plot(color='black', label='SNTHERM-like', ax=axes[1, 2], linewidth=4);
reynolds_depth.sel(model='CLM' ).plot(color='lime', label='CLM-like', ax=axes[1, 2], linewidth=4);
reynolds_depth.sel(model='2L_thin' ).plot(color=colors_2l[0], label='2l_thin', ax=axes[1, 2], linewidth=1.5);
reynolds_depth.sel(model='2L_mid' ).plot(color=colors_2l[1], label='2l_mid', ax=axes[1, 2], linewidth=1.5);
reynolds_depth.sel(model='2L_thick').plot(color=colors_2l[2], label='2l_thick', ax=axes[1, 2], linewidth=1.5);
reynolds_depth.sel(model='3L_thin' ).plot(color=colors_3l[0], label='3l_thin', ax=axes[1, 2], linewidth=1.5);
reynolds_depth.sel(model='3L_mid' ).plot(color=colors_3l[1], label='3l_mid', ax=axes[1, 2], linewidth=1.5);
reynolds_depth.sel(model='3L_thick').plot(color=colors_3l[2], label='3l_thick', ax=axes[1, 2], linewidth=1.5);
reynolds_depth.sel(model='4L_thin' ).plot(color=colors_4l[0], label='4l_thin', ax=axes[1, 2], linewidth=1.5);
reynolds_depth.sel(model='4L_mid' ).plot(color=colors_4l[1], label='4l_mid', ax=axes[1, 2], linewidth=1.5);
reynolds_depth.sel(model='4L_thick').plot(color=colors_4l[2], label='4l_thick', ax=axes[1, 2], linewidth=1.5);
reynolds_swe.sel(model='JRDN' ).plot(color='black', label='SNTHERM-like', ax=axes[0, 2], linewidth=4);
reynolds_swe.sel(model='CLM' ).plot(color='lime', label='CLM-like', ax=axes[0, 2], linewidth=4);
reynolds_swe.sel(model='2L_thin' ).plot(color=colors_2l[0], label='2l_thin', ax=axes[0, 2], linewidth=1.5);
reynolds_swe.sel(model='2L_mid' ).plot(color=colors_2l[1], label='2l_mid', ax=axes[0, 2], linewidth=1.5);
reynolds_swe.sel(model='2L_thick').plot(color=colors_2l[2], label='2l_thick', ax=axes[0, 2], linewidth=1.5);
reynolds_swe.sel(model='3L_thin' ).plot(color=colors_3l[0], label='3l_thin', ax=axes[0, 2], linewidth=1.5);
reynolds_swe.sel(model='3L_mid' ).plot(color=colors_3l[1], label='3l_mid', ax=axes[0, 2], linewidth=1.5);
reynolds_swe.sel(model='3L_thick').plot(color=colors_3l[2], label='3l_thick', ax=axes[0, 2], linewidth=1.5);
reynolds_swe.sel(model='4L_thin' ).plot(color=colors_4l[0], label='4l_thin', ax=axes[0, 2], linewidth=1.5);
reynolds_swe.sel(model='4L_mid' ).plot(color=colors_4l[1], label='4l_mid', ax=axes[0, 2], linewidth=1.5);
reynolds_swe.sel(model='4L_thick').plot(color=colors_4l[2], label='4l_thick', ax=axes[0, 2], linewidth=1.5);
axes[0, 2].set_ylabel('')
axes[1, 2].set_ylabel('')
axes[0, 2].set_xlabel('')
axes[2, 1].set_ylabel('')
axes[2, 2].set_ylabel('')
axes[0, 2].set_title('Reynolds Creek')
axes[1, 0].set_title('')
axes[1, 1].set_title('')
axes[1, 2].set_title('')
axes[2, 0].set_title('')
axes[2, 1].set_title('')
axes[2, 2].set_title('')
plt.tight_layout(pad=-0.1)
fig.autofmt_xdate()
#axes[1, 2].legend(bbox_to_anchor=(.1, -.4))
axes[2, 2].legend(bbox_to_anchor=(0.8, 1.5), framealpha=1.0)
# +
mpl.rcParams['figure.figsize'] = (20, 6)
fig, axes = plt.subplots(1, 3, sharey=True)
dana_ts = slice('2009-10-28 06', '2010-06-15 18')
coldeport_ts = slice('2000-11-15 06', '2001-06-01 18')
reynolds_ts = slice('2003-12-14 06', '2004-04-01 18')
dana_snotemp = 1000 * dana['scalarSnowDensity'].sel(dt='+0.0K', time=dana_ts)
coldeport_snotemp = 1000 * coldeport['scalarSnowDensity'].sel(dt='+0.0K', time=coldeport_ts)
reynolds_snotemp = 1000 * reynolds['scalarSnowDensity'].sel(dt='+0.0K', time=reynolds_ts)
ms = 7
# Plotting for dana meadows
#dana_airtemp.plot(color='black', label='Air temp', ax=axes[0], linewidth=6);
(dana_snotemp.sel(model='JRDN' ) ).plot(color='black', label='SNTHERM-like', ax=axes[0], linewidth=4);
(dana_snotemp.sel(model='CLM' ) ).plot(color='lime', label='CLM-like', ax=axes[0], linewidth=4);
(dana_snotemp.sel(model='2L_thin' ) ).plot(color=colors_2l[0], label='2l_thin', ax=axes[0], linewidth=1.5);
(dana_snotemp.sel(model='2L_mid' ) ).plot(color=colors_2l[1], label='2l_mid', ax=axes[0], linewidth=1.5);
(dana_snotemp.sel(model='2L_thick') ).plot(color=colors_2l[2], label='2l_thick', ax=axes[0], linewidth=1.5);
(dana_snotemp.sel(model='3L_thin' ) ).plot(color=colors_3l[0], label='3l_thin', ax=axes[0], linewidth=1.5);
(dana_snotemp.sel(model='3L_mid' ) ).plot(color=colors_3l[1], label='3l_mid', ax=axes[0], linewidth=1.5);
(dana_snotemp.sel(model='3L_thick') ).plot(color=colors_3l[2], label='3l_thick', ax=axes[0], linewidth=1.5);
(dana_snotemp.sel(model='4L_thin' ) ).plot(color=colors_4l[0], label='4l_thin', ax=axes[0], linewidth=1.5);
(dana_snotemp.sel(model='4L_mid' ) ).plot(color=colors_4l[1], label='4l_mid', ax=axes[0], linewidth=1.5);
(dana_snotemp.sel(model='4L_thick') ).plot(color=colors_4l[2], label='4l_thick', ax=axes[0], linewidth=1.5);
# Plotting for Col-de-Port
#coldeport_airtemp.plot(color='black', label='Air temp', ax=axes[1], linewidth=6);
(coldeport_snotemp.sel(model='JRDN' ) ).plot(color='black', label='SNTHERM-like', ax=axes[1], linewidth=4);
(coldeport_snotemp.sel(model='CLM' ) ).plot(color='lime', label='CLM-like', ax=axes[1], linewidth=4);
(coldeport_snotemp.sel(model='2L_thin' ) ).plot(color=colors_2l[0], label='2l_thin', ax=axes[1], linewidth=1.5);
(coldeport_snotemp.sel(model='2L_mid' ) ).plot(color=colors_2l[1], label='2l_mid', ax=axes[1], linewidth=1.5);
(coldeport_snotemp.sel(model='2L_thick') ).plot(color=colors_2l[2], label='2l_thick', ax=axes[1], linewidth=1.5);
(coldeport_snotemp.sel(model='3L_thin' ) ).plot(color=colors_3l[0], label='3l_thin', ax=axes[1], linewidth=1.5);
(coldeport_snotemp.sel(model='3L_mid' ) ).plot(color=colors_3l[1], label='3l_mid', ax=axes[1], linewidth=1.5);
(coldeport_snotemp.sel(model='3L_thick') ).plot(color=colors_3l[2], label='3l_thick', ax=axes[1], linewidth=1.5);
(coldeport_snotemp.sel(model='4L_thin' ) ).plot(color=colors_4l[0], label='4l_thin', ax=axes[1], linewidth=1.5);
(coldeport_snotemp.sel(model='4L_mid' ) ).plot(color=colors_4l[1], label='4l_mid', ax=axes[1], linewidth=1.5);
(coldeport_snotemp.sel(model='4L_thick') ).plot(color=colors_4l[2], label='4l_thick', ax=axes[1], linewidth=1.5);
# Plotting for reynolds creek
(reynolds_snotemp.sel(model='JRDN' )).plot(color='black', label='SNTHERM-like', ax=axes[2], linewidth=4);
(reynolds_snotemp.sel(model='CLM' )).plot(color='lime', label='CLM-like', ax=axes[2], linewidth=4);
(reynolds_snotemp.sel(model='2L_thin' )).plot(color=colors_2l[0], label='2l_thin', ax=axes[2], linewidth=1.5);
(reynolds_snotemp.sel(model='2L_mid' )).plot(color=colors_2l[1], label='2l_mid', ax=axes[2], linewidth=1.5);
(reynolds_snotemp.sel(model='2L_thick')).plot(color=colors_2l[2], label='2l_thick', ax=axes[2], linewidth=1.5);
(reynolds_snotemp.sel(model='3L_thin' )).plot(color=colors_3l[0], label='3l_thin', ax=axes[2], linewidth=1.5);
(reynolds_snotemp.sel(model='3L_mid' )).plot(color=colors_3l[1], label='3l_mid', ax=axes[2], linewidth=1.5);
(reynolds_snotemp.sel(model='3L_thick')).plot(color=colors_3l[2], label='3l_thick', ax=axes[2], linewidth=1.5);
(reynolds_snotemp.sel(model='4L_thin' )).plot(color=colors_4l[0], label='4l_thin', ax=axes[2], linewidth=1.5);
(reynolds_snotemp.sel(model='4L_mid' )).plot(color=colors_4l[1], label='4l_mid', ax=axes[2], linewidth=1.5);
(reynolds_snotemp.sel(model='4L_thick')).plot(color=colors_4l[2], label='4l_thick', ax=axes[2], linewidth=1.5);
axes[0].set_title('Dana Meadows')
axes[1].set_title('Col de Porte')
axes[2].set_title('Reynolds Creek')
axes[0].set_ylabel(r'Snow density ($kg / m^3$)')
axes[1].set_ylabel('')
axes[2].set_ylabel('')
axes[0].set_xlabel('')
axes[1].set_xlabel('')
axes[2].set_xlabel('')
print(xticks()[1])
#plt.tight_layout(pad=-1.1)
axes[2].fmt_xdata = mdates.DateFormatter('%Y-%m')
fig.autofmt_xdate()
axes[2].legend(bbox_to_anchor=(1., 1.))
# +
mpl.rcParams['figure.figsize'] = (20, 12)
fig, axes = plt.subplots(2, 3, sharey='row')
axes = axes.flatten()
dana_ts = slice('2010-01-15 06', '2010-01-16 18')
coldeport_ts = slice('2001-01-15 06', '2001-01-16 18')
reynolds_ts = slice('2004-01-15 06', '2004-01-16 18')
dana_airtemp = dana['airtemp'].sel(time=dana_ts) - 273.16
dana_snotemp = dana['scalarSurfaceTemp'].sel(dt='+0.0K', time=dana_ts) - 273.16
coldeport_airtemp = coldeport['airtemp'].sel(time=coldeport_ts) - 273.16
coldeport_snotemp = coldeport['scalarSurfaceTemp'].sel(dt='+0.0K', time=coldeport_ts) - 273.16
reynolds_airtemp = reynolds['airtemp'].sel(time=reynolds_ts) - 273.16
reynolds_snotemp = reynolds['scalarSurfaceTemp'].sel(dt='+0.0K', time=reynolds_ts) - 273.16
ms = 7
# Plotting for dana meadows
(dana_snotemp.sel(model='JRDN')).plot(color='black', label='CLM', ax=axes[0], linewidth=4);
#dana_airtemp.plot(color='black', label='Air temp', ax=axes[0], linewidth=6);
(dana_snotemp.sel(model='CLM' ) - dana_snotemp.sel(model='JRDN')).plot(color='lime', label='CLM-like', ax=axes[3], linewidth=4);
(dana_snotemp.sel(model='2L_thin' ) - dana_snotemp.sel(model='JRDN')).plot(color=colors_2l[0], label='2l_thin', ax=axes[3],marker='o', markersize=ms, linewidth=1.5);
(dana_snotemp.sel(model='2L_mid' ) - dana_snotemp.sel(model='JRDN')).plot(color=colors_2l[1], label='2l_mid', ax=axes[3],marker='s', markersize=ms, linewidth=1.5);
(dana_snotemp.sel(model='2L_thick') - dana_snotemp.sel(model='JRDN')).plot(color=colors_2l[2], label='2l_thick', ax=axes[3],marker='d', markersize=ms, linewidth=1.5);
(dana_snotemp.sel(model='3L_thin' ) - dana_snotemp.sel(model='JRDN')).plot(color=colors_3l[0], label='3l_thin', ax=axes[3],marker='o', markersize=ms, linewidth=1.5);
(dana_snotemp.sel(model='3L_mid' ) - dana_snotemp.sel(model='JRDN')).plot(color=colors_3l[1], label='3l_mid', ax=axes[3],marker='s', markersize=ms, linewidth=1.5);
(dana_snotemp.sel(model='3L_thick') - dana_snotemp.sel(model='JRDN')).plot(color=colors_3l[2], label='3l_thick', ax=axes[3],marker='d', markersize=ms, linewidth=1.5);
(dana_snotemp.sel(model='4L_thin' ) - dana_snotemp.sel(model='JRDN')).plot(color=colors_4l[0], label='4l_thin', ax=axes[3],marker='o', markersize=ms, linewidth=1.5);
(dana_snotemp.sel(model='4L_mid' ) - dana_snotemp.sel(model='JRDN')).plot(color=colors_4l[1], label='4l_mid', ax=axes[3],marker='s', markersize=ms, linewidth=1.5);
(dana_snotemp.sel(model='4L_thick') - dana_snotemp.sel(model='JRDN')).plot(color=colors_4l[2], label='4l_thick', ax=axes[3],marker='d', markersize=ms, linewidth=1.5);
# Plotting for Col-de-Port
(coldeport_snotemp.sel(model='JRDN')).plot(color='black', label='', ax=axes[1], linewidth=4);
#coldeport_airtemp.plot(color='black', label='Air temp', ax=axes[1], linewidth=6);
(coldeport_snotemp.sel(model='CLM' ) - coldeport_snotemp.sel(model='JRDN')).plot(color='lime', label='CLM-like', ax=axes[4], linewidth=4);
(coldeport_snotemp.sel(model='2L_thin' ) - coldeport_snotemp.sel(model='JRDN')).plot(color=colors_2l[0], label='2l_thin', ax=axes[4],marker='o', markersize=ms, linewidth=1.5);
(coldeport_snotemp.sel(model='2L_mid' ) - coldeport_snotemp.sel(model='JRDN')).plot(color=colors_2l[1], label='2l_mid', ax=axes[4],marker='s', markersize=ms, linewidth=1.5);
(coldeport_snotemp.sel(model='2L_thick') - coldeport_snotemp.sel(model='JRDN')).plot(color=colors_2l[2], label='2l_thick', ax=axes[4],marker='d', markersize=ms, linewidth=1.5);
(coldeport_snotemp.sel(model='3L_thin' ) - coldeport_snotemp.sel(model='JRDN')).plot(color=colors_3l[0], label='3l_thin', ax=axes[4],marker='o', markersize=ms, linewidth=1.5);
(coldeport_snotemp.sel(model='3L_mid' ) - coldeport_snotemp.sel(model='JRDN')).plot(color=colors_3l[1], label='3l_mid', ax=axes[4],marker='s', markersize=ms, linewidth=1.5);
(coldeport_snotemp.sel(model='3L_thick') - coldeport_snotemp.sel(model='JRDN')).plot(color=colors_3l[2], label='3l_thick', ax=axes[4],marker='d', markersize=ms, linewidth=1.5);
(coldeport_snotemp.sel(model='4L_thin' ) - coldeport_snotemp.sel(model='JRDN')).plot(color=colors_4l[0], label='4l_thin', ax=axes[4],marker='o', markersize=ms, linewidth=1.5);
(coldeport_snotemp.sel(model='4L_mid' ) - coldeport_snotemp.sel(model='JRDN')).plot(color=colors_4l[1], label='4l_mid', ax=axes[4],marker='s', markersize=ms, linewidth=1.5);
(coldeport_snotemp.sel(model='4L_thick') - coldeport_snotemp.sel(model='JRDN')).plot(color=colors_4l[2], label='4l_thick', ax=axes[4],marker='d', markersize=ms, linewidth=1.5);
# Plotting for reynolds creek
(reynolds_snotemp.sel(model='JRDN')).plot(color='black', label='', ax=axes[2], linewidth=4);
(reynolds_snotemp.sel(model='CLM' ) - reynolds_snotemp.sel(model='JRDN')).plot(color='lime', label='CLM-like', ax=axes[5], linewidth=4);
(reynolds_snotemp.sel(model='2L_thin' ) - reynolds_snotemp.sel(model='JRDN')).plot(color=colors_2l[0], label='2l_thin', ax=axes[5], marker='o', markersize=ms, linewidth=1.5);
(reynolds_snotemp.sel(model='2L_mid' ) - reynolds_snotemp.sel(model='JRDN')).plot(color=colors_2l[1], label='2l_mid', ax=axes[5], marker='s', markersize=ms, linewidth=1.5);
(reynolds_snotemp.sel(model='2L_thick') - reynolds_snotemp.sel(model='JRDN')).plot(color=colors_2l[2], label='2l_thick', ax=axes[5], marker='d', markersize=ms, linewidth=1.5);
(reynolds_snotemp.sel(model='3L_thin' ) - reynolds_snotemp.sel(model='JRDN')).plot(color=colors_3l[0], label='3l_thin', ax=axes[5], marker='o', markersize=ms, linewidth=1.5);
(reynolds_snotemp.sel(model='3L_mid' ) - reynolds_snotemp.sel(model='JRDN')).plot(color=colors_3l[1], label='3l_mid', ax=axes[5], marker='s', markersize=ms, linewidth=1.5);
(reynolds_snotemp.sel(model='3L_thick') - reynolds_snotemp.sel(model='JRDN')).plot(color=colors_3l[2], label='3l_thick', ax=axes[5], marker='d', markersize=ms, linewidth=1.5);
(reynolds_snotemp.sel(model='4L_thin' ) - reynolds_snotemp.sel(model='JRDN')).plot(color=colors_4l[0], label='4l_thin', ax=axes[5], marker='o', markersize=ms, linewidth=1.5);
(reynolds_snotemp.sel(model='4L_mid' ) - reynolds_snotemp.sel(model='JRDN')).plot(color=colors_4l[1], label='4l_mid', ax=axes[5], marker='s', markersize=ms, linewidth=1.5);
(reynolds_snotemp.sel(model='4L_thick') - reynolds_snotemp.sel(model='JRDN')).plot(color=colors_4l[2], label='4l_thick', ax=axes[5], marker='d', markersize=ms, linewidth=1.5);
axes[0].set_title('Dana Meadows')
axes[1].set_title('Col de Porte')
axes[2].set_title('Reynolds Creek')
axes[3].set_title('')
axes[4].set_title('')
axes[5].set_title('')
axes[0].set_ylabel('SNTHERM-like snow \n surface temperature [C]')
axes[1].set_ylabel('')
axes[2].set_ylabel('')
axes[3].set_ylabel('Difference from SNTHERM-like \n snow surface temperature [C]')
axes[4].set_ylabel('')
axes[5].set_ylabel('')
axes[0].set_xlabel('')
axes[1].set_xlabel('')
axes[2].set_xlabel('')
axes[3].set_xlabel('')
axes[4].set_xlabel('')
axes[5].set_xlabel('')
axes[0].axhline(0, color='grey')
axes[1].axhline(0, color='grey')
axes[2].axhline(0, color='grey')
axes[3].axhline(0, color='grey')
axes[4].axhline(0, color='grey')
axes[5].axhline(0, color='grey')
plt.tight_layout(pad=-1.3)
fig.autofmt_xdate()
axes[-1].legend(bbox_to_anchor=(1., 1.6))
# +
mpl.rcParams['figure.figsize'] = (20, 12)
fig, axes = plt.subplots(2, 3, sharey='row', sharex=False)
colors = ['powderblue', 'azure', '#ffe119', '#f58231']
dana_ts = slice('2009-10-01', '2010-08-01')
dana_swe = dana['scalarSWE'].sel(time=dana_ts).resample(time='D').mean()
dana_swe.sel(dt='-2.0K').plot.line(x='time', label='-2.0K', add_legend=False, color='skyblue', ax=axes[0,0])
dana_swe.sel(dt='+0.0K').plot.line(x='time', label='-2.0K', add_legend=False, color='paleturquoise', ax=axes[0,0])
dana_swe.sel(dt='+2.0K').plot.line(x='time', label='-2.0K', add_legend=False, color='#ffe119', ax=axes[0,0])
dana_swe.sel(dt='+4.0K').plot.line(x='time', label='-2.0K', add_legend=False, color='#f58231', ax=axes[0,0])
dana_ts = slice('2016-10-01', '2017-09-01')
dana_swe = dana['scalarSWE'].sel(time=dana_ts).resample(time='D').mean()
dana_swe.sel(dt='-2.0K').plot.line(x='time', label='-2.0K', add_legend=False, color='skyblue', ax=axes[1,0])
dana_swe.sel(dt='+0.0K').plot.line(x='time', label='-2.0K', add_legend=False, color='paleturquoise', ax=axes[1,0])
dana_swe.sel(dt='+2.0K').plot.line(x='time', label='-2.0K', add_legend=False, color='#ffe119', ax=axes[1,0])
dana_swe.sel(dt='+4.0K').plot.line(x='time', label='-2.0K', add_legend=False, color='#f58231', ax=axes[1,0])
coldeport_ts = slice('2002-10-01', '2003-09-01')
coldeport_swe = coldeport['scalarSWE'].sel(time=coldeport_ts).resample(time='D').mean()
coldeport_swe.sel(dt='-2.0K').plot.line(x='time', label='-2.0K', add_legend=False, color='skyblue', ax=axes[0,1])
coldeport_swe.sel(dt='+0.0K').plot.line(x='time', label='-2.0K', add_legend=False, color='paleturquoise', ax=axes[0,1])
coldeport_swe.sel(dt='+2.0K').plot.line(x='time', label='-2.0K', add_legend=False, color='#ffe119', ax=axes[0,1])
coldeport_swe.sel(dt='+4.0K').plot.line(x='time', label='-2.0K', add_legend=False, color='#f58231', ax=axes[0,1])
coldeport_ts = slice('1998-09-01', '1999-09-01')
coldeport_swe = coldeport['scalarSWE'].sel(time=coldeport_ts).resample(time='D').mean()
coldeport_swe.sel(dt='-2.0K').plot.line(x='time', label='-2.0K', add_legend=False, color='skyblue', ax=axes[1,1])
coldeport_swe.sel(dt='+0.0K').plot.line(x='time', label='-2.0K', add_legend=False, color='paleturquoise', ax=axes[1,1])
coldeport_swe.sel(dt='+2.0K').plot.line(x='time', label='-2.0K', add_legend=False, color='#ffe119', ax=axes[1,1])
coldeport_swe.sel(dt='+4.0K').plot.line(x='time', label='-2.0K', add_legend=False, color='#f58231', ax=axes[1,1])
reynolds_ts = slice('2002-10-01', '2003-06-01')
reynolds_swe = reynolds['scalarSWE'].sel(time=reynolds_ts).resample(time='D').mean()
reynolds_swe.sel(dt='-2.0K').plot.line(x='time', label='-2.0K', add_legend=False, color='skyblue', ax=axes[0,2])
reynolds_swe.sel(dt='+0.0K').plot.line(x='time', label='-2.0K', add_legend=False, color='paleturquoise', ax=axes[0,2])
reynolds_swe.sel(dt='+2.0K').plot.line(x='time', label='-2.0K', add_legend=False, color='#ffe119', ax=axes[0,2])
reynolds_swe.sel(dt='+4.0K').plot.line(x='time', label='-2.0K', add_legend=False, color='#f58231', ax=axes[0,2])
reynolds_ts = slice('2003-10-01', '2004-06-01')
reynolds_swe = reynolds['scalarSWE'].sel(time=reynolds_ts).resample(time='D').mean()
reynolds_swe.sel(dt='-2.0K').plot.line(x='time', label='-2.0K', add_legend=False, color='skyblue', ax=axes[1,2])
reynolds_swe.sel(dt='+0.0K').plot.line(x='time', label='-2.0K', add_legend=False, color='paleturquoise', ax=axes[1,2])
reynolds_swe.sel(dt='+2.0K').plot.line(x='time', label='-2.0K', add_legend=False, color='#ffe119', ax=axes[1,2])
reynolds_swe.sel(dt='+4.0K').plot.line(x='time', label='-2.0K', add_legend=False, color='#f58231', ax=axes[1,2])
axes[0, 0].set_xlabel('')
axes[0, 1].set_xlabel('')
axes[0, 2].set_xlabel('')
axes[1, 0].set_xlabel('')
axes[1, 1].set_xlabel('')
axes[1, 2].set_xlabel('')
axes[0, 0].set_ylabel('')
axes[0, 1].set_ylabel('')
axes[0, 2].set_ylabel('')
axes[1, 0].set_ylabel('')
axes[1, 1].set_ylabel('')
axes[1, 2].set_ylabel('')
axes[0, 0].set_title('Dana Meadows (low snow year)', fontsize=22)
axes[0, 1].set_title('Col de Porte (low snow year)', fontsize=22)
axes[0, 2].set_title('Reynolds Creek (low snow year)', fontsize=22)
axes[1, 0].set_title('Dana Meadows (high snow year)', fontsize=22)
axes[1, 1].set_title('Col de Porte (high snow year)', fontsize=22)
axes[1, 2].set_title('Reynolds Creek (high snow year)', fontsize=22)
#axes[0, 2].legend(bbox_to_anchor=(1., 1.))
plt.setp( axes[0, 0].xaxis.get_majorticklabels(), rotation=45 );
plt.setp( axes[0, 1].xaxis.get_majorticklabels(), rotation=45 );
plt.setp( axes[0, 2].xaxis.get_majorticklabels(), rotation=45 );
plt.setp( axes[1, 0].xaxis.get_majorticklabels(), rotation=45 );
plt.setp( axes[1, 1].xaxis.get_majorticklabels(), rotation=45 );
plt.setp( axes[1, 2].xaxis.get_majorticklabels(), rotation=45 );
fig.text(-0.03, 0.5, r'SWE (mm)', va='center', rotation='vertical', fontsize=22)
colors_all = ['skyblue', 'paleturquoise', '#ffe119', '#f58231']
layers = ['-2.0K', '+0.0K', '+2.0K', '+4.0K']
legend_elements = [Line2D([0], [0], color=c, label=m, markersize=15) for m, c in zip(layers, colors_all)]
plt.legend(handles=legend_elements, bbox_to_anchor=(1., 1.))
plt.tight_layout()
# +
mpl.rcParams['figure.figsize'] = (24, 8)
fig, axes = plt.subplots(1, 3, sharex=False, sharey=False)
axes = axes.flatten()
date1 = '2004-10-01'
date2 = '2007-08-01'
coldeport['scalarSWE'].sel(dt='+0.0K', time=slice(date1, date2), model='JRDN').plot(color='black', label='SNTHERM-like', linewidth=4, ax=axes[0]);
cdp_obs['SWE_auto'].sel(time=slice(date1, date2)).plot(color='red', label='Observed', ax=axes[0])
plt.xlabel('')
plt.title('')
plt.legend()
date1 = '2009-10-01'
date2 = '2012-08-01'
dana['scalarSWE'].sel(dt='+0.0K', time=slice(date1, date2), model='JRDN').plot(color='black', label='SNTHERM-like', linewidth=4, ax=axes[1]);
dana_obs['Dana Meadows Pillow SWE [mm]'][date1:date2].plot(color='red',linewidth=3, label='Observed', ax=axes[1])
plt.xlabel('')
plt.title('')
plt.legend()
date1 = '1999-10-01'
date2 = '2002-08-01'
reynolds['scalarSWE'].sel(dt='+0.0K', time=slice(date1, date2), model='JRDN').plot(color='black', label='SNTHERM-like', linewidth=4, ax=axes[2]);
reynolds_obs['SWE'].sel(time=slice(date1, date2)).plot(color='red',linewidth=3, label='Observed', ax=axes[2])
plt.xlabel('')
plt.ylabel(r'Snow water equivalent $(kg / m^3)$')
plt.title('')
axes[0].set_ylabel(r'Snow Water Equivalent $(kg / m^3)$')
axes[1].set_ylabel(r'')
axes[2].set_ylabel(r'')
axes[0].set_title(r'Col de Porte')
axes[1].set_title(r'Dana Meadows')
axes[2].set_title(r'Reynolds Creek')
axes[0].set_xlabel(r'')
axes[1].set_xlabel(r'')
axes[2].set_xlabel(r'')
plt.legend()
# -
| gen_plots/timeseries_plots.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # July 17 - Latent vector analysis 2
# +
# Imports
import math
import os
import sys
import pandas as pd
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
from matplotlib.colors import LogNorm
from mpl_toolkits.mplot3d import Axes3D
# Add the path to the parent directory to augment search for module
par_dir = os.path.abspath(os.path.join(os.getcwd(), os.pardir))
if par_dir not in sys.path:
sys.path.append(par_dir)
# Import the custom plotting module
from plot_utils import plot_utils
import random
import torch
# +
# Label dict - Dictionary mapping integer labels to str
label_dict = {0:"gamma", 1:"e", 2:"mu"}
np.set_printoptions(threshold=np.inf)
def plot_event(run_id, iteration, mode):
dump_dir = "/home/akajal/WatChMaL/VAE/dumps/" + run_id + "/"
if mode is "validation":
np_arr_path = dump_dir + "val_iteration_" + str(iteration) + ".npz"
else:
np_arr_path = dump_dir + "iteration_" + str(iteration) + ".npz"
# Load the numpy array
np_arr = np.load(np_arr_path)
np_event, np_recon, np_labels, np_energies = np_arr["events"], np_arr["prediction"], np_arr["labels"], np_arr["energies"]
i = random.randint(0, np_labels.shape[0]-1)
plot_utils.plot_actual_vs_recon(np_event[i], np_recon[i],
label_dict[np_labels[i]], np_energies[i].item(),
show_plot=True)
plot_utils.plot_charge_hist(torch.tensor(np_event).permute(0,2,3,1).numpy(),
np_recon, iteration, num_bins=200)
def plot_log(run_id, model_name, iteration, variant, mode):
dump_dir = "/home/akajal/WatChMaL/VAE/dumps/" + run_id + "/"
# Setup the path to the training log file
if mode is "training":
log = dump_dir + "log_train.csv"
elif mode is "training_validation":
log = dump_dir + "val_test.csv"
elif mode is "validation":
log = dump_dir + "validation_log.csv"
else:
print("mode has to be one of training, training_validation, validation")
return None
downsample_interval = 32 if mode is "training" else None
if variant is "AE":
plot_utils.plot_ae_training([log], [model_name], {model_name:["red"]},
downsample_interval=downsample_interval, show_plot=True, legend_loc=(0.8,0.8))
elif variant is "VAE":
plot_utils.plot_vae_training([log], [model_name], {model_name:["red", "blue"]},
downsample_interval=downsample_interval, show_plot=True, legend_loc=(0.8,0.8))
if iteration is not None:
plot_event(run_id, iteration, mode=mode)
# -
# ## Latent vector analysis to find the difference b/w the latent vectors generated by the buggy ( model that works ) v.s. debugged ( model that does not work )
# ## 1. Debugged model ( Regular VAE which does not work )
# +
run_id = "20190717_181756"
dump_dir = "/home/akajal/WatChMaL/VAE/dumps/" + run_id + "/"
# Load the numpy array for the various iterations
np_arr_0 = np.load(dump_dir + "iteration_" + str(3) + ".npz")
np_arr_1 = np.load(dump_dir + "iteration_" + str(1500) + ".npz")
np_arr_2 = np.load(dump_dir + "iteration_" + str(3000) + ".npz")
# Extract the latent vector np arrays from the loaded archive
np_arr_0_z, np_arr_1_z, np_arr_2_z = np_arr_0["z"], np_arr_1["z"], np_arr_2["z"]
# -
plot_log("20190717_181756", "ENet(VAE)", None, "VAE", "training")
# ## Plot the histogram for only one of the dimensions (dim = 0) from each iteration
np_arr_0_z_0 = np_arr_0_z[:,0]
np_arr_1_z_0 = np_arr_1_z[:,0]
np_arr_2_z_0 = np_arr_2_z[:,0]
# +
num_bins = 100
# Initialize the plot and corresponding parameters
fig, ax = plt.subplots(figsize=(16,9),facecolor="w")
ax.tick_params(axis="both", labelsize=20)
# Setup the bins beforehand
bins = np.linspace(min(np.amin(np_arr_0_z_0), np.amin(np_arr_1_z_0), np.amin(np_arr_2_z_0)),
max(np.amax(np_arr_0_z_0), np.amax(np_arr_1_z_0), np.amax(np_arr_2_z_0)),
num_bins)
plt.hist(np_arr_0_z_0, bins, density=False,
label="iteration 3", color="red",
alpha=0.5, stacked=True, histtype="step")
plt.hist(np_arr_1_z_0, bins, density=False,
label="iteration 1500", color="blue",
alpha=0.5, stacked=True, histtype="step")
plt.hist(np_arr_2_z_0, bins, density=False,
label="iteration 3000", color="green",
alpha=0.5, stacked=True, histtype="step")
# Setup the axes
ax.set_xlabel(r"$z_0$", fontsize=20)
ax.set_ylabel("Frequency", fontsize=20)
plt.legend(loc="upper right", prop={"size":20})
plt.grid(True)
plt.title(r"Distribution of $z_0$ at intervals 3, 1500, 3000", fontsize=20)
plt.show()
# -
# ## Plot the histogram for all 64 dimensions flattened
# +
run_id = "20190717_181756"
dump_dir = "/home/akajal/WatChMaL/VAE/dumps/" + run_id + "/"
# Load the numpy array for the various iterations
np_arr_0 = np.load(dump_dir + "iteration_" + str(3) + ".npz")
np_arr_1 = np.load(dump_dir + "iteration_" + str(1500) + ".npz")
np_arr_2 = np.load(dump_dir + "iteration_" + str(3000) + ".npz")
# Extract the latent vector np arrays from the loaded archive
np_arr_0_z, np_arr_1_z, np_arr_2_z = np_arr_0["z"], np_arr_1["z"], np_arr_2["z"]
np_arr_0_z_0 = np_arr_0_z.reshape(-1, 1)
np_arr_1_z_0 = np_arr_1_z.reshape(-1, 1)
np_arr_2_z_0 = np_arr_2_z.reshape(-1, 1)
num_bins = 100
# Initialize the plot and corresponding parameters
fig, ax = plt.subplots(figsize=(16,9),facecolor="w")
ax.tick_params(axis="both", labelsize=20)
# Setup the bins beforehand
bins = np.linspace(min(np.amin(np_arr_0_z_0), np.amin(np_arr_1_z_0), np.amin(np_arr_2_z_0)),
max(np.amax(np_arr_0_z_0), np.amax(np_arr_1_z_0), np.amax(np_arr_2_z_0)),
num_bins)
plt.hist(np_arr_0_z_0, bins, density=False,
label="iteration 3", color="red",
alpha=0.5, stacked=True, histtype="step")
plt.hist(np_arr_1_z_0, bins, density=False,
label="iteration 1500", color="blue",
alpha=0.5, stacked=True, histtype="step")
plt.hist(np_arr_2_z_0, bins, density=False,
label="iteration 3000", color="green",
alpha=0.5, stacked=True, histtype="step")
# Setup the axes
ax.set_xlabel(r"$z$", fontsize=20)
ax.set_ylabel("Frequency", fontsize=20)
plt.legend(loc="upper right", prop={"size":20})
plt.yscale("log")
plt.grid(True)
plt.title(r"Distribution of $z$ at intervals 3, 1500, 3000", fontsize=20)
plt.show()
# -
# ## Collect and compute the scatter plot values of $\mu$ and $\Sigma$ for each of the dimensions
# +
run_id = "20190718_015626"
dump_dir = "/home/akajal/WatChMaL/VAE/dumps/" + run_id + "/"
#iterations = [3,1500,3000]
iterations = [1500]
iteration_colors = ["red", "blue", "green"]
iteration_color_dict = {}
for i,iteration in enumerate(iterations):
iteration_color_dict[iteration] = iteration_colors[i]
# Load the numpy array for the various iterations
np_arr_dict = {}
for iteration in iterations:
np_arr_dict[iteration] = np.load(dump_dir + "iteration_" + str(iteration) + ".npz")["z_prime"]
num_dims = np_arr_dict[list(np_arr_dict.keys())[0]].shape[1]
scatter_values = {}
for iteration in iterations:
iter_scatter_values_mu = []
iter_scatter_values_var = []
for i in range(num_dims):
iter_scatter_values_mu.append(np.mean(np_arr_dict[iteration][:,i]))
iter_scatter_values_var.append(np.var(np_arr_dict[iteration][:,i]))
scatter_values[iteration] = [iter_scatter_values_mu, iter_scatter_values_var]
# -
# ## Plot the scatter plot for 𝜇
# +
# Initialize the figure
fig, ax = plt.subplots(figsize=(16,9),facecolor="w")
ax.tick_params(axis="both", labelsize=20)
# Use the dimensions as the values for the x-axis
x = np.linspace(1, num_dims, num_dims)
# Plot the scatter for mu values for each of the iteration
for iteration in iterations:
plt.scatter(x, scatter_values[iteration][0], color=iteration_color_dict[iteration],
label="iteration {0}".format(iteration), s=100)
# Setup the axes
ax.set_xlabel(r"$z'$ dimension", fontsize=20)
ax.set_ylabel(r"$\mu$", fontsize=20)
plt.legend(loc="upper right", prop={"size":20})
plt.grid(True)
plt.title(r"Mean of different $z'$ dimensions at iterations {0}".format(iterations), fontsize=20)
plt.show()
# -
# ## Plot the scatter plot for $\Sigma$
# +
# Initialize the figure
fig, ax = plt.subplots(figsize=(16,9),facecolor="w")
ax.tick_params(axis="both", labelsize=20)
# Use the dimensions as the values for the x-axis
x = np.linspace(1, num_dims, num_dims)
# Plot the scatter for mu values for each of the iteration
for iteration in iterations:
plt.scatter(x, scatter_values[iteration][1], s=100, color=iteration_color_dict[iteration],
label="iteration {0}".format(iteration))
# Setup the axes
ax.set_xlabel(r"$z'$ dimension", fontsize=20)
ax.set_ylabel(r"$\sigma$", fontsize=20)
plt.legend(loc="upper right", prop={"size":20})
plt.grid(True)
plt.title(r"Variance of different $z'$ dimensions at iterations {0}".format(iterations), fontsize=20)
plt.show()
# -
# ## 2. Buggy model which does work
# +
run_id = "20190718_015626"
dump_dir = "/home/akajal/WatChMaL/VAE/dumps/" + run_id + "/"
# Load the numpy array for the various iterations
np_arr_0 = np.load(dump_dir + "iteration_" + str(3) + ".npz")
np_arr_1 = np.load(dump_dir + "iteration_" + str(1500) + ".npz")
np_arr_2 = np.load(dump_dir + "iteration_" + str(3000) + ".npz")
# Extract the latent vector np arrays from the loaded archive
np_arr_0_z, np_arr_1_z, np_arr_2_z = np_arr_0["z_prime"], np_arr_1["z_prime"], np_arr_2["z_prime"]
np_arr_0_z_0 = np_arr_0_z[:,0]
np_arr_1_z_0 = np_arr_1_z[:,0]
np_arr_2_z_0 = np_arr_2_z[:,0]
#--------------------------------------------------------------------------------
# Plot the single dimension histogram
#--------------------------------------------------------------------------------
num_bins = 100
# Initialize the plot and corresponding parameters
fig, ax = plt.subplots(figsize=(16,9),facecolor="w")
ax.tick_params(axis="both", labelsize=20)
# Setup the bins beforehand
bins = np.linspace(min(np.amin(np_arr_0_z_0), np.amin(np_arr_1_z_0), np.amin(np_arr_2_z_0)),
max(np.amax(np_arr_0_z_0), np.amax(np_arr_1_z_0), np.amax(np_arr_2_z_0)),
num_bins)
plt.hist(np_arr_0_z_0, bins, density=False,
label="iteration 3", color="red",
alpha=0.5, stacked=True)
plt.hist(np_arr_1_z_0, bins, density=False,
label="iteration 1500", color="blue",
alpha=0.5, stacked=True)
plt.hist(np_arr_2_z_0, bins, density=False,
label="iteration 3000", color="green",
alpha=0.5, stacked=True)
# Setup the axes
ax.set_xlabel(r"$z'_0$", fontsize=20)
ax.set_ylabel("Frequency", fontsize=20)
plt.yscale("log")
plt.xlim(-0.001, 0.001)
plt.legend(loc="upper right", prop={"size":20})
plt.grid(True)
plt.title(r"Distribution of $z'_0$ at intervals 3, 1500, 3000", fontsize=20)
plt.show()
#--------------------------------------------------------------------------------
# Plot the all dimension histogram
#--------------------------------------------------------------------------------
np_arr_0_z_0 = np_arr_0_z.reshape(-1, 1)
np_arr_1_z_0 = np_arr_1_z.reshape(-1, 1)
np_arr_2_z_0 = np_arr_2_z.reshape(-1, 1)
num_bins = 100
# Initialize the plot and corresponding parameters
fig, ax = plt.subplots(figsize=(16,9),facecolor="w")
ax.tick_params(axis="both", labelsize=20)
# Setup the bins beforehand
bins = np.linspace(min(np.amin(np_arr_0_z_0), np.amin(np_arr_1_z_0), np.amin(np_arr_2_z_0)),
max(np.amax(np_arr_0_z_0), np.amax(np_arr_1_z_0), np.amax(np_arr_2_z_0)),
num_bins)
plt.hist(np_arr_0_z_0, bins, density=False,
label="iteration 3", color="red",
alpha=0.5, stacked=True,histtype="step")
plt.hist(np_arr_1_z_0, bins, density=False,
label="iteration 1500", color="blue",
alpha=0.5, stacked=True, histtype="step")
plt.hist(np_arr_2_z_0, bins, density=False,
label="iteration 3000", color="green",
alpha=0.5, stacked=True, histtype="step")
# Setup the axes
ax.set_xlabel(r"$z$", fontsize=20)
ax.set_ylabel("Frequency", fontsize=20)
plt.legend(loc="upper right", prop={"size":20})
plt.yscale("log")
plt.xlim(-0.001, 0.001)
plt.grid(True)
plt.title(r"Distribution of $z$ at intervals 3, 1500, 3000", fontsize=20)
plt.show()
# -
# ## Analysis of the latent vectors being generated from the encoder
# +
run_id = "20190718_015626"
dump_dir = "/home/akajal/WatChMaL/VAE/dumps/" + run_id + "/"
# Load the numpy array for the various iterations
np_arr_0 = np.load(dump_dir + "iteration_" + str(3) + ".npz")
np_arr_1 = np.load(dump_dir + "iteration_" + str(1500) + ".npz")
np_arr_2 = np.load(dump_dir + "iteration_" + str(3000) + ".npz")
# Extract the latent vector np arrays from the loaded archive
np_arr_0_z, np_arr_1_z, np_arr_2_z = np_arr_0["z_prime"], np_arr_1["z_prime"], np_arr_2["z_prime"]
# -
| notebooks/notebooks_archive/July 17 - Latent vector analysis 2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 1. In the following code, print 98.6. What is "98.6"?
# - A constant
# 2. In the following code, x = 42. What is "x"?
# - A variable
# 3. Which of the following variables is the "most mnemonic"?
# - hours
# 4. Which of the following is not a Python reserved word?
# - speed
# 5. Assume the variable x has been initialized to an integer value (e.g., x = 3). What does the following statement do? x = x + 2
# - Retrieve the current value for x, add two to it and put the sum back into x
# 6. Which of the following elements of a mathematical expression in Python is evaluated first?
# -Parenthesis()
# 7. What is the value of the following expression
# - 2
# 8. What will be the value of x after the following statement executes: x = 1 + 2 * 3 - 8 / 4
# - 5
# 9. What will be the value of x when the following statement is executed: x = int(98.6)
# - 98
# 10. What does the Python raw_input() function do?
# - Pause the program and read data from the user
| Course-1-Programming-for-Everybody-Getting-Started-with-Python/Quizzes/Quiz Chapter 2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
from rnn_utils import *
def rnn_cell_forward(xt, a_prev, parameters):
Wax = parameters["Wax"]
Waa = parameters["Waa"]
Wya = parameters["Wya"]
ba = parameters["ba"]
by = parameters["by"]
a_next = np.tanh(np.dot(Waa, a_prev) + np.dot(Wax, xt) + ba)
yt_pred = softmax(np.dot(Wya, a_next) + by)
cache = (a_next, a_prev, xt, parameters)
return a_next, yt_pred, cache
def rnn_forward(x, a0, parameters):
caches = []
n_x, m, T_x = x.shape
n_y, n_a = parameters["Wya"].shape
a = np.zeros((n_a, m, T_x))
y_pred = np.zeros((n_y, m, T_x))
a_next = a0
for t in range(T_x):
xt = x[:, :, t]
a_next, yt_pred, cache = rnn_cell_forward(xt, a_next, parameters)
a[:,:,t] = a_next
y_pred[:,:,t] = yt_pred
caches.append(cache)
caches = (caches, x)
return a, y_pred, caches
def lstm_cell_forward(xt, a_prev, c_prev, parameters):
Wf = parameters["Wf"]
bf = parameters["bf"]
Wi = parameters["Wi"]
bi = parameters["bi"]
Wc = parameters["Wc"]
bc = parameters["bc"]
Wo = parameters["Wo"]
bo = parameters["bo"]
Wy = parameters["Wy"]
by = parameters["by"]
n_x, m = xt.shape
n_y, n_a = Wy.shape
concat = np.concatenate((a_prev, xt), axis = 0)
ft = sigmoid(np.dot(Wf, concat) + bf) # forget gate
it = sigmoid(np.dot(Wi, concat) + bi) # update gate
cct = np.tanh(np.dot(Wc, concat) + bc) # candidate value
c_next = ft * c_prev + it * cct # cell state
ot = sigmoid(np.dot(Wo, concat) + bo) # output gate
a_next = ot * np.tanh(c_next) # hidden state
yt_pred = softmax(np.dot(Wy, a_next) + by)
cache = (a_next, c_next, a_prev, c_prev, ft, it, cct, ot, xt, parameters)
return a_next, c_next, yt_pred, cache
def lstm_forward(x, a0, parameters):
caches = []
Wy = parameters['Wy']
n_x, m, T_x = x.shape
n_y, n_a = Wy.shape
a = np.zeros((n_a, m, T_x))
c = np.zeros_like(a)
y = np.zeros((n_y, m, T_x))
a_next = a0
c_next = np.zeros_like(a_next)
for t in range(T_x):
xt = x[:, :, t]
a_next, c_next, yt, cache = lstm_cell_forward(xt, a_next, c_next, parameters)
a[:,:,t] = a_next
c[:,:,t] = c_next
y[:,:,t] = yt
caches.append(cache)
caches = (caches, x)
return a, y, c, caches
def rnn_cell_backward(da_next, cache):
(a_next, a_prev, xt, parameters) = cache
Wax = parameters["Wax"]
Waa = parameters["Waa"]
Wya = parameters["Wya"]
ba = parameters["ba"]
by = parameters["by"]
dtanh = (1 - a_next ** 2) * da_next
dxt = np.dot(Wax.T, dtanh)
dWax = np.dot(dtanh, xt.T)
da_prev = np.dot(Waa.T, dtanh)
dWaa = np.dot(dtanh, a_prev.T)
dba = np.sum(dtanh, axis = 1,keepdims=1)
gradients = {"dxt": dxt, "da_prev": da_prev, "dWax": dWax, "dWaa": dWaa, "dba": dba}
return gradients
def rnn_backward(da, caches):
(caches, x) = caches
(a1, a0, x1, parameters) = caches[0]
n_a, m, T_x = da.shape
n_x, m = x1.shape
dx = np.zeros((n_x, m, T_x))
dWax = np.zeros((n_a, n_x))
dWaa = np.zeros((n_a, n_a))
dba = np.zeros((n_a, 1))
da0 = np.zeros((n_a, m))
da_prevt = np.zeros((n_a, m))
for t in reversed(range(T_x)):
gradients = rnn_cell_backward(da[:,:,t] + da_prevt, caches[t])
dxt, da_prevt, dWaxt, dWaat, dbat = gradients["dxt"], gradients["da_prev"], gradients["dWax"], gradients["dWaa"], gradients["dba"]
dx[:, :, t] = dxt
dWax += dWaxt
dWaa += dWaat
dba += dbat
da0 = da_prevt
gradients = {"dx": dx, "da0": da0, "dWax": dWax, "dWaa": dWaa,"dba": dba}
return gradients
def lstm_cell_backward(da_next, dc_next, cache):
(a_next, c_next, a_prev, c_prev, ft, it, cct, ot, xt, parameters) = cache
n_x, m = xt.shape
n_a, m = a_next.shape
dot = da_next * np.tanh(c_next) * ot * (1 - ot)
dcct = (da_next * ot * (1 - np.tanh(c_next) ** 2) + dc_next) * it * (1 - cct ** 2)
dit = (da_next * ot * (1 - np.tanh(c_next) ** 2) + dc_next) * cct * (1 - it) * it
dft = (da_next * ot * (1 - np.tanh(c_next) ** 2) + dc_next) * c_prev * ft * (1 - ft)
dWf = np.dot(dft, np.hstack([a_prev.T, xt.T]))
dWi = np.dot(dit, np.hstack([a_prev.T, xt.T]))
dWc = np.dot(dcct, np.hstack([a_prev.T, xt.T]))
dWo = np.dot(dot, np.hstack([a_prev.T, xt.T]))
dbf = np.sum(dft, axis=1, keepdims=True)
dbi = np.sum(dit, axis=1, keepdims=True)
dbc = np.sum(dcct, axis=1, keepdims=True)
dbo = np.sum(dot, axis=1, keepdims=True)
da_prev = np.dot(Wf[:, :n_a].T, dft) + np.dot(Wc[:, :n_a].T, dcct) + np.dot(Wi[:, :n_a].T, dit) + np.dot(Wo[:, :n_a].T, dot)
dc_prev = (da_next * ot * (1 - np.tanh(c_next) ** 2) + dc_next) * ft
dxt = np.dot(Wf[:, n_a:].T, dft) + np.dot(Wc[:, n_a:].T, dcct) + np.dot(Wi[:, n_a:].T, dit) + np.dot(Wo[:, n_a:].T, dot)
gradients = {"dxt": dxt, "da_prev": da_prev, "dc_prev": dc_prev, "dWf": dWf,"dbf": dbf, "dWi": dWi,"dbi": dbi,
"dWc": dWc,"dbc": dbc, "dWo": dWo,"dbo": dbo}
return gradients
def lstm_backward(da, caches):
(caches, x) = caches
(a1, c1, a0, c0, f1, i1, cc1, o1, x1, parameters) = caches[0]
n_a, m, T_x = da.shape
n_x, m = x1.shape
dx = np.zeros((n_x, m, T_x))
da0 = np.zeros((n_a, m))
da_prevt = np.zeros((n_a, m))
dc_prevt = np.zeros((n_a, m))
dWf = np.zeros((n_a, n_a + n_x))
dWi = np.zeros((n_a, n_a + n_x))
dWc = np.zeros((n_a, n_a + n_x))
dWo = np.zeros((n_a, n_a + n_x))
dbf = np.zeros((n_a, 1))
dbi = np.zeros((n_a, 1))
dbc = np.zeros((n_a, 1))
dbo = np.zeros((n_a, 1))
for t in reversed(range(T_x)):
gradients = lstm_cell_backward(da[:,:,t] + da_prevt, dc_prevt, caches[t])
dx[:,:,t] = gradients["dxt"]
dWf += gradients["dWf"]
dWi += gradients["dWi"]
dWc += gradients["dWc"]
dWo += gradients["dWo"]
dbf += gradients["dbf"]
dbi += gradients["dbi"]
dbc += gradients["dbc"]
dbo += gradients["dbo"]
da0 = gradients["da_prev"]
gradients = {"dx": dx, "da0": da0, "dWf": dWf,"dbf": dbf, "dWi": dWi,"dbi": dbi,
"dWc": dWc,"dbc": dbc, "dWo": dWo,"dbo": dbo}
return gradients
| RNN with LSTM/Basic RNN with LSTM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Step B
import warnings
warnings.filterwarnings('ignore')
import numpy as np
import cv2
from matplotlib import pyplot as plt
import skimage.morphology as sk
from operator import itemgetter
# The ```compute_all_keypoints``` function calculates all keypoints of all query and train images and stores them in a dictionary, in order to easily access them later.
def compute_all_keypoints(query_imgs, train_imgs, sift):
img_dict = {}
for img in query_imgs:
file = 'models/' + img + '.jpg'
query = cv2.imread(file, 0)
kp, des = sift.detectAndCompute(query, None)
img_dict[img] = {'kp': kp, 'des': des, 'shape': query.shape}
for img in train_imgs:
file = 'scenes/' + img + '.png'
train = cv2.imread(file, 0)
kp, des = sift.detectAndCompute(train, None)
img_dict[img] = {'kp': kp, 'des': des, 'shape': train.shape}
return img_dict
# The ```apply_ratio_test``` function takes all the matches found between the query and the train image, it chooses the good ones with the usual ratio test and it stores them in a dictionary using the indexes of the query keypoints as keys and the indexes of the train keypoints as values.
def apply_ratio_test(all_matches):
# map of matches kp_train_idx -> kp_query_idx
good_matches = {}
for m, n in all_matches:
if m.distance < LOWE_COEFF * n.distance:
good_matches[m.queryIdx] = m.trainIdx
return good_matches
# The ```compute_entry_hough_space``` function maps a point into the 4-dimensional Hough space, the ```create_hough_space``` function computes the Hough space entries for all keypoints.
def compute_entry_hough_space(kp_q, kp_t, q_xc, q_yc):
entry = {}
v = ((q_xc - kp_q.pt[0]), (q_yc - kp_q.pt[1]))
scale_ratio = kp_t.size / kp_q.size
delta_angle = kp_t.angle - kp_q.angle
x_c = kp_t.pt[0] + scale_ratio * (np.cos(delta_angle) * v[0] - np.sin(delta_angle) * v[1])
y_c = kp_t.pt[1] + scale_ratio * (np.sin(delta_angle) * v[0] + np.cos(delta_angle) * v[1])
entry['x_c'] = x_c
entry['y_c'] = y_c
entry['scale_ratio'] = scale_ratio
entry['delta_angle'] = delta_angle
return entry
def create_hough_space(good_matches, kp_query, kp_train, query_xc, query_yc):
# map of hough space kp_train_idx -> map name-values
hough_space = {}
for t_idx, q_idx in good_matches.items():
hough_space[t_idx] = compute_entry_hough_space(kp_query[q_idx], kp_train[t_idx], query_xc, query_yc)
return hough_space
# The ```compute_bins``` function partitions the 4-dimensional Hough space into discrete bins.
def compute_bins(hough_space,query_shape,train_shape):
values = {}
data_scale = [entry['scale_ratio'] for entry in hough_space.values()]
counts_scale, bins_scale, patches_size = plt.hist(data_scale, bins='auto')
img_scale = np.mean([bins_scale[np.argmax(counts_scale)], bins_scale[np.argmax(counts_scale) + 1]])
plt.close();
data_angle = [entry['delta_angle'] for entry in hough_space.values()]
counts_angle, bins_angle, patches_angle = plt.hist(data_angle, bins='auto')
plt.close();
x_bin_size = img_scale * query_shape[1] * BIN_PRECISION_FACTOR
y_bin_size = img_scale * query_shape[0] * BIN_PRECISION_FACTOR
x_bins = int(np.ceil(train_shape[1] / x_bin_size) + 2)
y_bins = int(np.ceil(train_shape[0] / y_bin_size) + 2)
x_min = train_shape[1] / 2 - x_bins / 2 * x_bin_size
y_min = train_shape[0] / 2 - y_bins / 2 * y_bin_size
angle_bin_size = np.std(data_angle) * ANGLE_BIN_SIZE_COEFF
angle_bin_center = np.mean(data_angle)
angle_min = angle_bin_center - ANGLE_BINS / 2 * angle_bin_size
angle_max = angle_bin_center + ANGLE_BINS / 2 * angle_bin_size
scale_bin_size = np.std(data_scale) * SCALE_BIN_SIZE_COEFF
scale_bin_center = np.mean(data_scale)
scale_min = 0
scale_max = scale_bin_center * 2
scale_bins = int((scale_max - scale_min) / scale_bin_size)
values['x_bins'] = x_bins
values['y_bins'] = y_bins
values['x_min'] = x_min
values['y_min'] = y_min
values['x_bin_size'] = x_bin_size
values['y_bin_size'] = y_bin_size
values['scale_bins'] = scale_bins
values['scale_min'] = scale_min
values['scale_bin_size'] = scale_bin_size
values['angle_min'] = angle_min
values['angle_bin_size'] = angle_bin_size
return values
# The ```voting``` function attributes 16 bins to each point in the 4-dimensional Hough space.
def voting(b,h_s):
accumulator = np.zeros((b['x_bins'], b['y_bins'], ANGLE_BINS, b['scale_bins']))
votes = {}
for idx, v in h_s.items():
try:
for x in range(0, 2):
for y in range(0, 2):
for z in range(0, 2):
for w in range(0, 2):
i = int(np.floor((v['x_c'] - b['x_min'] + (x - 1 / 2) * b['x_bin_size']) / b['x_bin_size']))
j = int(np.floor((v['y_c'] - b['y_min'] + (y - 1 / 2) * b['y_bin_size']) / b['y_bin_size']))
k = int(np.floor((v['delta_angle'] - b['angle_min'] + (z - 1 / 2) * b['angle_bin_size']) / b['angle_bin_size']))
l = int(np.floor((v['scale_ratio'] - b['scale_min'] + (w - 1 / 2) * b['scale_bin_size']) / b['scale_bin_size']))
if i >= 0 and j >= 0 and k >= 0 and l >= 0:
accumulator[i, j, k, l] += 1
votes[(i, j, k, l)] = votes.get((i, j, k, l), [])
votes[(i, j, k, l)].append(idx)
except: pass
return accumulator, votes
# The ```find_all_correspondeces``` function computes all the correspondeces between query points and train points that voted for a local maxima.
def find_all_correspondeces(query_imgs, train_img, img_dict, bf):
# list of lists of all correspondent point of every image
global_correspondences = []
for query_img in query_imgs:
# compute keypoints and desctiptors for query and train
kp_query, des_query = img_dict[query_img]['kp'], img_dict[query_img]['des']
kp_train, des_train = img_dict[train_img]['kp'], img_dict[train_img]['des']
# match descriptors between the two images
all_matches = bf.knnMatch(des_train, des_query, k=2)
# create map of matching keypoint indexes surviving the lowe ratio test
good_matches = apply_ratio_test(all_matches)
# barycenter of found query keypoint
query_xc = np.mean(list(kp_query[p].pt[0] for _, p in good_matches.items()))
query_yc = np.mean(list(kp_query[p].pt[1] for _, p in good_matches.items()))
# create hough space
hough_space = create_hough_space(good_matches, kp_query, kp_train, query_xc, query_yc)
# do not go on with this query image if the number of entries in the hough space are below a certain threshold
if len(hough_space) < HOUGH_T: continue
# compute all the values related to the size
bins_values = compute_bins(hough_space, img_dict[query_img]['shape'], img_dict[train_img]['shape'])
# create and populate accumulator with voting by each entry of the hough space
accumulator, votes= voting(bins_values, hough_space)
# compute local maxima of the 4-dimensional accumulator
mask = sk.local_maxima(accumulator)
accumulator[mask != 1] = 0
# store in a list all the correspondeces between query points and train points that voted for a local maxima
# the list contains: number of votes that a local maxima bin has received, name of query image, list of query and train keypoints which voted for that bin
for b in list(np.argwhere(accumulator >= T_Q)): # thresholding the accumulator to come up with few maxima
keypoint_index_list = votes[tuple(b)] # all query keypoint who voted for a local maxima bin
correspondence_list = [(kp_train[k], kp_query[good_matches[k]]) for k in keypoint_index_list]
global_correspondences.append([accumulator[tuple(b)], query_img, correspondence_list])
# sorted correspondences based on number of votes found in local maxima bins
g_c = sorted(global_correspondences, key=itemgetter(0), reverse=True)
return g_c
# The ```check_matches``` function orders the good matches in decreasing number of keypoints and it runs a series of tests on them, checking the geometric arrangement and the color consistency.
def check_matches(correspondences, train_img):
train_file = 'scenes/' + train_img + '.png'
train_bgr = cv2.imread(train_file)
#dict query_name -> list of projected query vertex into train image
recognised = {}
for entry in correspondences:
try:
query_file = 'models/' + entry[1] + '.jpg'
query_bgr = cv2.imread(query_file)
# compute homography through correspondent keypoints
src_pts = np.float32([e[1].pt for e in entry[2]]).reshape(-1, 1, 2)
dst_pts = np.float32([e[0].pt for e in entry[2]]).reshape(-1, 1, 2)
M, _ = cv2.findHomography(src_pts, dst_pts, cv2.RANSAC, 5.0)
h, w, d = query_bgr.shape
pts = np.float32([[0, 0], [0, h - 1], [w - 1, h - 1], [w - 1, 0]]).reshape(-1,1,2)
dst = cv2.perspectiveTransform(pts, M)
# determine center of the train ROI matching with the image query
center = tuple((dst[0, 0, i] + dst[1, 0, i] + dst[2, 0, i] + dst[3, 0, i]) / 4 for i in (0, 1))
# determine extreme points of the quadrilateral shape of query image projected into train scene
x_min = int(max((dst[0,0,0] + dst[1,0,0]) / 2, 0))
y_min = int(max((dst[0,0,1] + dst[3,0,1]) / 2, 0))
x_max = int(min((dst[2,0,0] + dst[3,0,0]) / 2, train_bgr.shape[1]))
y_max = int(min((dst[1,0,1] + dst[2,0,1]) / 2, train_bgr.shape[0]))
# compute main color of both query and train ROI to tell similar boxes apart
query_color = query_bgr.mean(axis=0).mean(axis=0)
train_crop = train_bgr[y_min:y_max, x_min:x_max]
train_color = train_crop.mean(axis=0).mean(axis=0)
color_diff = np.sqrt(np.sum([value ** 2 for value in abs(query_color - train_color)]))
# recognise a query in the scene only if its ROI color is similar to the query color and it does not overlap with another already placed box
temp = True
if color_diff < COLOR_T:
for k, v in recognised.items():
for corners in v:
r_center = tuple((corners[0, 0, i] + corners[1, 0, i] + corners[2, 0, i] + corners[3, 0, i]) / 4 for i in (0, 1))
if (center[0] > min(corners[0, 0, 0], corners[1, 0, 0]) and center[0] < max(corners[2, 0, 0], corners[3, 0, 0])\
and center[1] > min(corners[0, 0, 1], corners[3, 0, 1]) and center[1] < max(corners[1, 0, 1], corners[2, 0, 1]))\
or (r_center[0] > x_min and r_center[0] < x_max\
and r_center[1] > y_min and r_center[1] < y_max):
temp = False
break
if temp:
recognised[entry[1]] = recognised.get(entry[1], [])
recognised[entry[1]].append(dst)
except: pass
return recognised
# The ```print_matches``` function takes all the recognised images and prints their details, i.e. their position, width, and height.
def print_matches(train_img, query_imgs, recognised, true_imgs, verbose):
print('Scene: ' + train_img + '\n')
for query_img in query_imgs:
total = len(recognised.get(query_img, []))
true_total = true_imgs[train_img][query_img]
if total != true_total:
print('\033[1m' + 'Product ' + query_img + ' – ' + str(total) + '/' + str(true_total) + ' instances found' + '\033[0m')
elif total > 0 or verbose == True:
print('Product ' + query_img + ' – ' + str(total) + '/' + str(true_total) + ' instances found')
for j in range(total):
dst = recognised[query_img][j]
center = tuple(int((dst[0, 0, i] + dst[1, 0, i] + dst[2, 0, i] + dst[3, 0, i]) / 4) for i in (0, 1))
w = int(((dst[3, 0, 0] - dst[0, 0, 0]) + (dst[2, 0, 0] - dst[1, 0, 0])) /2)
h = int(((dst[1, 0, 1] - dst[0, 0, 1]) + (dst[2, 0, 1] - dst[3, 0, 1])) /2)
print('\t' + 'Position: ' + str(center)\
+ '\t' + 'Width: ' + str(w)\
+ '\t' + 'Height: ' + str(h))
# The ```draw_matches``` function draws on the train image the boxes' homographies and the numbers corresponding to the query images.
def draw_matches(recognised, train_img, color):
train_file = 'scenes/' + train_img + '.png'
# if color option is enabled all the results are printed on colored images
if color == True:
train_bgr = cv2.imread(train_file)
train_temp = cv2.cvtColor(train_bgr, cv2.COLOR_BGR2RGB)
train_rgb = np.zeros(train_bgr.shape, train_bgr.dtype)
for y in range(train_temp.shape[0]):
for x in range(train_temp.shape[1]):
for c in range(train_temp.shape[2]):
train_rgb[y, x, c] = np.clip(0.5 * train_temp[y, x, c], 0, 255)
else:
train_gray = cv2.imread(train_file, 0)
train_rgb = cv2.cvtColor(train_gray // 2, cv2.COLOR_GRAY2RGB)
# for each recognised box in the scene draw the bounding box with its number in it
for k, v in recognised.items():
for dst in v:
train_rgb = cv2.polylines(train_rgb, [np.int32(dst)], True, (0, 255, 0), 3, cv2.LINE_AA)
font = cv2.FONT_HERSHEY_SIMPLEX
cv2.putText(train_rgb, k,\
(int((dst[3, 0, 0] - dst[0, 0, 0]) * 0.25 + dst[0, 0, 0]), int((dst[1, 0, 1] - dst[0, 0, 1]) * 0.67 + dst[0, 0, 1])),\
font, 5, (0, 255, 0), 10, cv2.LINE_AA)
plt.imshow(train_rgb),plt.show();
if color == True:
cv2.imwrite('output/step_B/' + train_img + '.png', cv2.cvtColor(train_rgb, cv2.COLOR_RGB2BGR))
# The ```step_B``` function takes the lists of query and train images and performs the product recognition.
# + tags=[]
def step_B(query_imgs, train_imgs, true_imgs, verbose, color):
sift = cv2.xfeatures2d.SIFT_create()
# descriptor matcher
bf = cv2.BFMatcher()
img_dict = compute_all_keypoints(query_imgs, train_imgs, sift) #compute all keypoints for all images once for all
for train_img in train_imgs:
g_c = find_all_correspondeces(query_imgs, train_img, img_dict, bf)
recognised = check_matches(g_c, train_img)
print_matches(train_img, query_imgs, recognised, true_imgs, verbose)
draw_matches(recognised, train_img, color)
print('\n')
# -
# Parameters:
LOWE_COEFF = 0.8
BIN_PRECISION_FACTOR = 0.25
ANGLE_BINS = 7
ANGLE_BIN_SIZE_COEFF = 0.1
SCALE_BIN_SIZE_COEFF = 0.1
T_Q = 5
COLOR_T = 50
HOUGH_T = 100
# + tags=[]
query_imgs = ['0', '1', '11', '19', '24', '25', '26']
train_imgs = ['e1', 'e2', 'e3', 'e4', 'e5',\
'm1', 'm2', 'm3', 'm4', 'm5']
true_imgs = {
'e1': {'0': 1, '1': 0, '11': 1, '19': 0, '24': 0, '25': 0, '26': 0},
'e2': {'0': 0, '1': 0, '11': 0, '19': 0, '24': 1, '25': 1, '26': 1},
'e3': {'0': 1, '1': 1, '11': 1, '19': 0, '24': 0, '25': 0, '26': 0},
'e4': {'0': 1, '1': 0, '11': 1, '19': 0, '24': 0, '25': 1, '26': 1},
'e5': {'0': 0, '1': 0, '11': 0, '19': 1, '24': 0, '25': 1, '26': 0},
'm1': {'0': 0, '1': 0, '11': 0, '19': 0, '24': 2, '25': 1, '26': 1},
'm2': {'0': 1, '1': 2, '11': 1, '19': 0, '24': 0, '25': 0, '26': 0},
'm3': {'0': 0, '1': 0, '11': 0, '19': 1, '24': 0, '25': 2, '26': 1},
'm4': {'0': 0, '1': 0, '11': 0, '19': 0, '24': 2, '25': 2, '26': 1},
'm5': {'0': 0, '1': 2, '11': 1, '19': 1, '24': 0, '25': 2, '26': 0},
}
# -
# verbose=False does not print the true negative instances
# color=True outputs all the scenes in color insted of grayscale, but the process is quite slow, therefore it is False by default
step_B(query_imgs, train_imgs, true_imgs, verbose=False, color=False)
| step_B.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # SQL
# Before you start, download the SQLite version of the [Chinook database](https://github.com/lerocha/chinook-database) from [GitHub](https://github.com/lerocha/chinook-database/raw/master/ChinookDatabase/DataSources/Chinook_Sqlite.sqlite).
# +
import numpy as np
import pandas as pd
import sqlite3
# Open connection to database
db_connection = sqlite3.connect('Chinook_Sqlite.sqlite')
# -
# ## Example
#
# Select the first 10 customers.
pd.read_sql(
'''SELECT *
FROM customer
LIMIT 10''', con=db_connection)
# ## Exercise 1
#
# Select the first name of all customers from the UK.
pd.read_sql(
'''SELECT FirstName
FROM customer
WHERE Country == \'United Kingdom\'''', con=db_connection)
# ## Exercise 2
#
# Select the city and country of all customers from the UK or Portugal.
pd.read_sql(
'''SELECT City, Country
FROM customer
WHERE Country == \'United Kingdom\'
OR Country == \'Portugal\'''', con=db_connection)
# ## Exercise 3
#
# Select the first 10 invoices.
pd.read_sql(
'''SELECT *
FROM invoice
LIMIT 10''', con=db_connection)
# ## Exercise 4
#
# Join the tables `customer` and `invoice`, and retrieve customer ID and invoice amount.
pd.read_sql(
'''SELECT c.CustomerId, i.Total
FROM customer AS c
JOIN invoice AS i
ON c.CustomerId == i.CustomerId''', con=db_connection)
# Now compute the total of all invoices by customer.
pd.read_sql(
'''SELECT c.CustomerId, SUM(i.Total)
FROM customer AS c
JOIN invoice AS i
ON c.CustomerId == i.CustomerId
GROUP BY c.CustomerId''', con=db_connection)
# Now aggregate only invoices from 2013.
#
# Hint: use the SQLite function `STRFTIME` on `InvoiceDate`.
pd.read_sql(
'''SELECT c.CustomerId, SUM(i.Total)
FROM customer AS c
JOIN invoice AS i
ON c.CustomerId == i.CustomerId
WHERE STRFTIME(\'%Y\', i.InvoiceDate) == \'2013\'
GROUP BY c.CustomerId''', con=db_connection)
# Now order by total amount in descending order.
pd.read_sql(
'''SELECT c.CustomerId, SUM(i.Total) AS total
FROM customer AS c
JOIN invoice AS i
ON c.CustomerId == i.CustomerId
WHERE STRFTIME(\'%Y\', i.InvoiceDate) == \'2013\'
GROUP BY c.CustomerId
ORDER BY total DESC''', con=db_connection)
# Finally, add the first name of the support rep from table `employee`.
pd.read_sql(
'''SELECT c.CustomerId, e.FirstName, SUM(i.Total) AS total
FROM customer AS c
JOIN invoice AS i
ON c.CustomerId == i.CustomerId
JOIN employee AS e
ON c.SupportRepId == e.EmployeeId
WHERE STRFTIME(\'%Y\', i.InvoiceDate) == \'2013\'
GROUP BY c.CustomerId
ORDER BY total DESC''', con=db_connection)
| 02_managing_data_and_analyses/notebooks/01_solutions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" papermill={"duration": 0.040845, "end_time": "2021-08-10T00:16:02.727422", "exception": false, "start_time": "2021-08-10T00:16:02.686577", "status": "completed"} tags=[]
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python Docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
# Input data files are available in the read-only "../input/" directory
# For example, running this (by clicking run or pressing Shift+Enter) will list all files under the input directory
import os
for dirname, _, filenames in os.walk('/kaggle/input'):
for filename in filenames:
print(os.path.join(dirname, filename))
# You can write up to 20GB to the current directory (/kaggle/working/) that gets preserved as output when you create a version using "Save & Run All"
# You can also write temporary files to /kaggle/temp/, but they won't be saved outside of the current session
# + papermill={"duration": 1.28814, "end_time": "2021-08-10T00:16:04.026834", "exception": false, "start_time": "2021-08-10T00:16:02.738694", "status": "completed"} tags=[]
import numpy as np
import pandas as pd
df1=pd.read_csv('../input/tmdb-movie-metadata/tmdb_5000_credits.csv')
df2=pd.read_csv('../input/tmdb-movie-metadata/tmdb_5000_movies.csv')
# + papermill={"duration": 0.084023, "end_time": "2021-08-10T00:16:04.126312", "exception": false, "start_time": "2021-08-10T00:16:04.042289", "status": "completed"} tags=[]
df2.rename(columns={'id':'movie_id'}, inplace=True)
print(df2.columns)
df2.head()
# + papermill={"duration": 0.029724, "end_time": "2021-08-10T00:16:04.168177", "exception": false, "start_time": "2021-08-10T00:16:04.138453", "status": "completed"} tags=[]
print(df1.columns)
df1.head()
# + papermill={"duration": 0.076808, "end_time": "2021-08-10T00:16:04.257365", "exception": false, "start_time": "2021-08-10T00:16:04.180557", "status": "completed"} tags=[]
movies=df2.merge(df1,on='movie_id')
movies.head()
# + [markdown] papermill={"duration": 0.013236, "end_time": "2021-08-10T00:16:04.284740", "exception": false, "start_time": "2021-08-10T00:16:04.271504", "status": "completed"} tags=[]
# # Let's try the simplest recommendation algorithm first; **Demographic Filtering**
#
# * Demographic Filtering (DF) technique uses the demographic data of a user to determine which items may be appropriate for recommendation.
# * This require a metric based on which movies can be rated.
# * After sorting based on the metric, we recommend the top movies to the user.
# * Generally used matric for this purpose is: weighted rating (wr)
#
# .
#
# Here,
# * v is the number of votes for the movie;
# * m is the minimum votes required to be listed in the chart;
# * R is the average rating of the movie; And
# * C is the mean vote across the whole report
#
# v(**vote_count**) and R (**vote_average**) are listed in our dataset.
#
# C is calculated by finding the mean of the average rating of all ovies
#
# We also pick an **m** value. This can be set as a quantile of the **vote_count** data. We pick 0.8 quantile.
#
#
# + papermill={"duration": 0.028928, "end_time": "2021-08-10T00:16:04.327225", "exception": false, "start_time": "2021-08-10T00:16:04.298297", "status": "completed"} tags=[]
C=movies['vote_average'].mean()
print('C = ', C)
m=movies['vote_count'].quantile(0.8)
print('m = ', m)
# + papermill={"duration": 0.040915, "end_time": "2021-08-10T00:16:04.382005", "exception": false, "start_time": "2021-08-10T00:16:04.341090", "status": "completed"} tags=[]
data=movies[movies['vote_count']>m]
# + [markdown] papermill={"duration": 0.013128, "end_time": "2021-08-10T00:16:04.408850", "exception": false, "start_time": "2021-08-10T00:16:04.395722", "status": "completed"} tags=[]
# ### We create a **score** to evaluate the movies using the WR, then sort the dataframe based on the score. Then pick the top 10 movies to feature.
# + papermill={"duration": 0.050836, "end_time": "2021-08-10T00:16:04.473155", "exception": false, "start_time": "2021-08-10T00:16:04.422319", "status": "completed"} tags=[]
def weighted_rating(x, m=m, C=C):
v = x['vote_count']
R = x['vote_average']
# Calculation based on the IMDB formula
return (v/(v+m) * R) + (m/(m+v) * C)
data['score']=data.apply(weighted_rating, axis=1)
data=data.sort_values('score', ascending=False)
# + papermill={"duration": 0.030632, "end_time": "2021-08-10T00:16:04.517823", "exception": false, "start_time": "2021-08-10T00:16:04.487191", "status": "completed"} tags=[]
data[['original_title', 'vote_count', 'vote_average', 'score']].head(10)
# + papermill={"duration": 0.014295, "end_time": "2021-08-10T00:16:04.546628", "exception": false, "start_time": "2021-08-10T00:16:04.532333", "status": "completed"} tags=[]
| demographic-filtering-recommendation.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.1.0
# language: julia
# name: julia-1.1
# ---
# # MLP model on MNIST
using Knet, Plots, Statistics, LinearAlgebra, Base.Iterators, Random, StatsBase, JLD
ENV["COLUMNS"] = 80
ARRAY = KnetArray{Float64}
XSIZE=784 # input dimension
HIDDENSIZE=32 # hidden layer dimension
YSIZE=10 # output dimension
WSIZE=HIDDENSIZE*(XSIZE+1)+YSIZE*(HIDDENSIZE+1) # DY: total size of W
WZERO=0.01 # DY: to be used by w initialization, do not use LAMBDA for this
BATCHSIZE=10 # minibatch size
LAMBDA=1e-2 # regularization parameter
#LAMBDA=1e-10
#LR=1e-1 # learning rate
LR=1e-2 # learning rate
MITER=10^4 # iterations for finding minimum
#MITER=10^5 # iterations for finding minimum # DY: do we really need 100K? Takes more than 2 hours!
DITER=10^5 # iterations for diffusion tensor
CITER=10^7 # iterations for covariance trajectory
CFREQ=10^2 # keep every CFREQ points on trajectory
@show gpu();
# # Define regularized linear model with softmax loss
# +
loss(w,x,y;λ=LAMBDA) = nll(pred(w,x), y) + (λ/2) * sum(abs2,w)
function pred(w,x)
w1 = reshape(w[1:HIDDENSIZE*XSIZE],HIDDENSIZE,XSIZE)
b1 = w[HIDDENSIZE*XSIZE+1:HIDDENSIZE*(XSIZE+1)]
w2 = reshape(w[1+HIDDENSIZE*(XSIZE+1):HIDDENSIZE*(XSIZE+1+YSIZE)],YSIZE,HIDDENSIZE)
b2 = w[1+HIDDENSIZE*(XSIZE+1+YSIZE):end] # DY: you forgot the final bias vector
x = reshape(x,XSIZE,:)
z = relu.(w1*x .+ b1) # DY: relu may be faster than max.(0,x)
return w2*z .+ b2
end
# -
# # Load MNIST data
include(Knet.dir("data/mnist.jl"))
xtrn,ytrn,xtst,ytst = mnist()
atrn,atst = ARRAY(xtrn), ARRAY(xtst) # GPU copies for batch training
println.(summary.((xtrn,ytrn,xtst,ytst,atrn,atst)));
# # Minibatch data
# Minibatching for SGD-I, i.e. with replacement. Knet.minibatch can't do this, we define new struct
struct MB; x; y; n; end
Base.Iterators.IteratorSize(::Type{MB}) = Base.IsInfinite() # need this for collect to work
Base.iterate(d::MB, s...)=(r = rand(1:length(d.y),d.n); ((ARRAY(mat(d.x)[:,r]), d.y[r]), true))
dtrn = MB(xtrn, ytrn, BATCHSIZE)
println.(summary.(first(dtrn)));
# # Find minimum
WSIZE,LAMBDA,WZERO,MITER # we are not using BATCHSIZE here
# Find minimum without minibatching
# ~11 iters/sec, converges in 15 mins 10K iters to
# loss=0.4994713306015728, nll=0.2576288962706217, reg=0.24184243433095112 (LAMBDA=0.01)
wminfile = "MLP-wmin-$WSIZE-$LAMBDA-$WZERO-$MITER.jld2"
if !isfile(wminfile)
wmin = Param(ARRAY(WZERO*randn(WSIZE))) # DY: why use LAMBDA here? why rand and not randn()? LAMBDA*rand((XSIZE+1)*HIDDENSIZE + HIDDENSIZE*YSIZE)))
args = repeat([(wmin,atrn,ytrn)],MITER)
Knet.gc()
losses = collect(progress(adam(loss,args)))
Knet.save(wminfile, "wmin", wmin, "losses", losses)
else
wmin, losses = Knet.load(wminfile, "wmin", "losses");
end
@show summary(wmin)
losses[end-4:end]'
println.((
(loss(wmin,atrn,ytrn),nll(pred(wmin,atrn),ytrn),(LAMBDA/2)*sum(abs2,wmin)),
(loss(wmin,atst,ytst),nll(pred(wmin,atst),ytst),(LAMBDA/2)*sum(abs2,wmin)),
(accuracy(pred(wmin,atrn),ytrn),accuracy(pred(wmin,atst),ytst))));
∇loss = grad(loss)
∇lossi(w,x,y,i) = ∇loss(w,x,y)[i]
∇∇lossi = grad(∇lossi)
w = value(wmin)
@show typeof.((w,atrn,ytrn))
@show ∇lossi(w,atrn,ytrn,1)
∇∇lossi(w,atrn,ytrn,1)'
#wcpu = Array(w)
#xcpu = convert(Array{Float64},xtrn)
#@show typeof.((wcpu,xcpu,ytrn))
#@show ∇lossi(wcpu,xcpu,ytrn,1)
#∇∇lossi(wcpu,xcpu,ytrn,1)'
# # Hessian of loss around minimum
function hessian(loss,w,x,y)
∇loss = grad(loss)
∇lossi(w,x,y,i) = ∇loss(w,x,y)[i]
∇∇lossi = grad(∇lossi)
w = value(w)
n = length(w)
# h = similar(w,n,n)
h = zeros(n,n)
for i in progress(1:n)
h[:,i] .= Array(vec(∇∇lossi(w,x,y,i)))
end
return h
end
# Compute hessian: ~80 mins
hessfile = "MLP-hess-$WSIZE-$LAMBDA.jld"
if !isfile(hessfile)
hmin = hessian(loss,wmin,atrn,ytrn)
save(hessfile,"h",hmin)
else
hmin = load(hessfile,"h")
end
println.((summary(hmin),extrema(Array(hmin)),norm(hmin),norm(hmin-hmin')));
# # Eigenvalues of the Hessian
heigfile = "MLP-nosym_heig-$LAMBDA.jld2"
H = Array(hmin)
if !isfile(heigfile)
# @time eigenH = eigen(Symmetric(H)) # ~53s
@time eigenH = eigen(H) # ~53s
Knet.save(heigfile,"eigenH",eigenH)
else
eigenH = Knet.load(heigfile,"eigenH")
end
eigenH.values'
summarystats(real.(eigenH.values)) |> dump
scatter(real.(eigenH.values),xaxis=:log10,yaxis=:log10)
# # Diffusion Tensor
function diffusiontensor(loss,w,x,y;iters=DITER,lr=LR,batchsize=BATCHSIZE)
∇loss = grad(loss)
grad0 = Array(∇loss(w, ARRAY(x), y))
data = MB(x,y,batchsize)
grads = zeros(length(w), iters)
for (i,d) in progress(enumerate(take(data,iters)))
grads[:,i] .= Array(∇loss(w,d...))
end
prefac = (lr^2)/(2iters)
grads = grad0 .- grads
@time v = prefac * (grads * grads')
return v
end
LAMBDA,LR,BATCHSIZE,DITER
dtfile = "MLP-dt-$LAMBDA-$LR-$BATCHSIZE-$DITER.jld2"
if !isfile(dtfile)
Knet.gc()
D = diffusiontensor(loss,wmin,xtrn,ytrn) # ~700 iters/sec
Knet.save(dtfile,"D",D)
else
D = Knet.load(dtfile,"D")
end
summarystats(vec(D)) |> dump
# # Record trajectory with SGD starting at minimum
LAMBDA,LR,BATCHSIZE,CITER,CFREQ
# Trajectory of w starting from wmin recorded after each update:
# ~1000 updates/sec, ~16 secs total
trajfile = "MLP-traj-$LAMBDA-$LR-$BATCHSIZE-$CITER-$CFREQ.jld2"
if !isfile(trajfile)
w = Param(ARRAY(value(wmin)))
data = MB(xtrn,ytrn,BATCHSIZE)
d = take(data,CITER)
W = zeros(eltype(w),length(w),div(CITER,CFREQ))
f(x,y) = loss(w,x,y)
Knet.gc()
i = 0
for t in progress(sgd(f,d; lr=LR))
i += 1; (div,rem)=divrem(i,CFREQ)
if rem == 0
W[:,div] = Array(vec(w))
end
end
Knet.save(trajfile,"W",W)
else
W = Knet.load(trajfile,"W")
end
summary(W)
# Plot losses on whole dataset, first steps seem transient, ~10 secs
r = 1:100:size(W,2)
@time plot(r, [loss(ARRAY(W[:,i]),atrn,ytrn) for i in r])
rr1,rr2 = rand(1:size(W,1)),rand(1:size(W,1))
# Plot trajectory of two random dimensions
@show rr1,rr2 = rand(1:size(W,1)),rand(1:size(W,1))
if norm(W[rr1,:]) > 0 && norm(W[rr2,:]) > 0
histogram2d(W[rr1,:],W[rr2,:],background_color="black")
end
# Minibatch training seems to converge to a slightly worse spot
w0 = Array(value(wmin))
μ = mean(W[:,2500:end],dims=2)
w1 = W[:,end]
@show norm(w0), norm(μ), norm(w0 - μ)
@show extrema(w0), extrema(μ), extrema(w0 - μ)
@show mean(abs.(w0 - μ) .> 0.01)
@show loss(w0,xtrn,ytrn)
@show loss(μ,xtrn,ytrn)
@show loss(w1,xtrn,ytrn)
# # Covariance of SGD trajectory around minimum
#Wstable = W[:,2500:end]; @show summary(Wstable)
Wstable = W
μ = mean(Wstable,dims=2); @show summary(μ)
Wzero = Wstable .- μ; @show summary(Wzero)
Σ = (Wzero * Wzero') / size(Wzero,2)
@show summary(Σ)
@show norm(Σ)
@show extrema(Σ)
@show norm(diag(Σ));
# check for convergence
n2 = div(size(W,2),2)
w1 = W[:,1:n2]
w2 = W[:,1+n2:end]
w1 = w1 .- mean(w1,dims=2)
w2 = w2 .- mean(w2,dims=2)
Σ1 = (w1 * w1') / size(w1,2)
Σ2 = (w2 * w2') / size(w2,2);
# The variances (diagonal elements) converge
norm(diag(Σ1)),norm(diag(Σ2)),norm(diag(Σ1)-diag(Σ2))
# The off diagonal elements are still not there
norm(Σ1),norm(Σ2),norm(Σ1-Σ2)
# # Check Einstein relation
summary.((H,D,Σ))
a = H*Σ + Σ*H
b = (2/LR)*D
norm(a),norm(b),norm(a-b)
# ## Covariance eigenspace
# 46 sec
@time eigenΣ = eigen(Σ);
ΛΣ = eigenΣ.values; O = eigenΣ.vectors;
# +
# check that the eigenvectors/values are OK
# norm(ΛΣ[end]*O[:,end]),norm(Σ*O[:,end]-ΛΣ[end]*O[:,end])
# -
# transform the trajectory to the eigenbasis
Weig = O'*W;
## check Einstein relation in top Neig eigen-directions of Σ
Neig=100
Or = O[:,end-Neig+1:end];
aa = Or'*a*Or
bb = Or'*b*Or
norm(aa),norm(bb)/norm(aa),norm(aa-bb)/norm(aa)
aa[1:10,1:10]
bb[1:10,1:10]
# +
# pick two eigen directions
Nweights = size(W,1)
Xid = Nweights
Yid = Nweights-1
#O = eigvecs(Σ);
#W_ss = W*O; # sample weights are row vectors
Wx = Weig[Xid,:]
Wy = Weig[Yid,:]
#COV_ss = O'*Σ*O
#COV_xy_inv = inv(COV_ss[[Xid,Yid],[Xid,Yid]])
COV_xy_inv = Diagonal([1/ΛΣ[Xid],1/ΛΣ[Yid]]) + zeros(2,2)
μeig = O'*μ
W0eig = O'*w0;
# -
myhplot = histogram2d(Wx,Wy
,bins=100
,aspect_ratio=1
,background_color="black"
);
display(myhplot)
# +
# Construct a grid enclosing the steady-state trajectory
minmaxdiff(t) = maximum(t)-minimum(t)
function makegrid(xvec,yvec,mean,xindex,yindex;Nx=50,Ny=50,zoom=0.35)
Lx,Ly = minmaxdiff(xvec),minmaxdiff(yvec)
xrange,yrange = zoom*Lx,zoom*Ly
dx,dy = xrange/Nx,yrange/Ny
x = collect(-xrange:dx:xrange) .+ mean[xindex]
y = collect(-yrange:dy:yrange) .+ mean[yindex]
# some mumbo-jumbo for calculating weights corresponding to grid points
Identity = Diagonal(ones(Nweights,Nweights)); # unit matrix
xmask = Identity[:,xindex];
ymask = Identity[:,yindex];
Imask = Identity - xmask*xmask' - ymask*ymask' # set two diagonal elements to zero
return (x,y,Imask,xmask,ymask)
end
# -
(x,y,Imask,xmask,ymask) = makegrid(Wx,Wy,W0eig,Xid,Yid;Nx=12,Ny=12,zoom=0.5)
#(x,y,Imask,xmask,ymask) = makegrid(Wx,Wy,μeig,Xid,Yid;Nx=4,Ny=4);
meanXY = W0eig[[Xid Yid]]
# Contours of the fit mv-Gaussian
ffit(s,t) = -(([s t]-meanXY)*COV_xy_inv*([s t]-meanXY)')[1]
Ffit(s,t) = 250*ffit(s,t)/ffit(x[end],y[end])
contour!(x,y,Ffit
,linestyle=:dash
,levels=10
,linewidth=2
)
# contours of loss
midx = Int((length(x)-1)/2)
midy = Int((length(y)-1)/2)
#fexp(s,t) = 1e5*(loss(O*(Imask*W0eig + s*xmask + t*ymask),xtrn,ytrn) - loss(w0,xtrn,ytrn))
fexp(s,t) = 1e5*(loss(O*(Imask*μeig + s*xmask + t*ymask),xtrn,ytrn) - loss(w0,xtrn,ytrn))
logfexpmidp1 = log(1+fexp(x[midx],y[midy]))
Flossxy(s,t) = 1e2*(log((1+fexp(s,t))) - logfexpmidp1)
Flossxy(0,0)
# ## N,N-1
@time contour!(x,y,Flossxy
#contour!(x,y,fexp
,levels=10
,linewidth=2
)
| archive/20190502-mnist-mlp-production.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Additional Resources
# Notebooks in science
#
# - [Will Jupyter notebook eventually replace scientific papers](https://www.quora.com/Will-Jupyter-notebooks-eventually-replace-scientific-papers)
#
#
# Examples of Notebooks
# - [Gallery of interesting notebooks](https://github.com/jupyter/jupyter/wiki/A-gallery-of-interesting-Jupyter-Notebooks)
# - [nbviewer - render a notebook](https://nbviewer.jupyter.org)
# - [28 Jupyter Notebook tips, tricks, and shortcuts](https://www.dataquest.io/blog/jupyter-notebook-tips-tricks-shortcuts/)
# - [Estimate of Public Jupyter Notebooks on GitHub](https://github.com/parente/nbestimate)
#
#
# Enabling slideshows in Jupyter notebook
#
# - [RISE - slideshows](http://www.damian.oquanta.info/index.html)
#
#
# Thoughts on version control in Jupyter notebooks
# - https://medium.freecodecamp.org/how-to-handle-version-control-and-reproducibility-with-jupyter-notebook-e1fbc0b8f922
#
#
# Tools using Jupyter notebooks:
#
# - [jupyter-repo2docker](https://github.com/jupyter/repo2docker) - jupyter-repo2docker takes as input a repository source, such as a GitHub repo. It then builds, runs, and/or pushes Docker images built from that source.
# - [nodebook](https://github.com/stitchfix/nodebook) - Repeatable analysis plugin for Jupyter notebook; designed to enforce an ordered flow of cell execution
# - [Building textbooks with Jupyter](https://github.com/choldgraf/textbooks-with-jupyter)
# - [Nikola - static site generate](https://getnikola.com)
#
#
| notebooks/4-Last-things/4-2-Additional-resources.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import arviz as az
import matplotlib.pyplot as plt
import numpy as np
import pymc as pm
az.style.use("arviz-white")
# # Using PyMC for MCMC sampling
#
# ### Chapter 3.3: Introduction to PyMC
#
# In this example, we use PyMC to conduct simple linear regression.
#
# The response is the mass of a T. Rex and the covariate is the age. The model is:
#
# $$ mass_i\sim\mbox{Normal}(\beta_1+\beta_2age_i,\sigma^2).$$
#
# The priors are:
# * $\beta_1,\beta_2\sim\mbox{Normal}(0,1000)$
# * $\sigma^2\sim\mbox{InvGamma}(0.1,0.1)$.
## Load T-Rex data
mass = np.array([29.9, 1761, 1807, 2984, 3230, 5040, 5654])
age = np.array([2, 15, 14, 16, 18, 22, 28])
# ## Define the model
with pm.Model() as model:
# Priors
τ = pm.Gamma("τ", 0.1, 10)
σ = pm.Deterministic("σ", 1 / (τ**0.5))
# σ = pm.HalfNormal('σ', np.std(mass))
β1 = pm.Normal("β1", 0, 1000)
β2 = pm.Normal("β2", 0, 1000)
# likelihood
mass_y = pm.Normal("mass_y", β1 + β2 * age, σ, observed=mass)
trace = pm.sample(2000)
# ## Summarize the output
az.summary(trace, var_names=["~τ"])
az.plot_trace(trace, var_names=["~τ"]);
| BSM/Chapter_03_09_Simple_linear_regression_in_PyMC3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Лабораторная работа 2. Введение в машинное обучение.
#
# Результат лабораторной работы − отчет. Мы принимаем отчеты в формате ноутбуков IPython (ipynb-файл). Постарайтесь сделать ваш отчет интересным рассказом, последовательно отвечающим на вопросы из заданий. Помимо ответов на вопросы, в отчете также должен быть код, однако чем меньше кода, тем лучше всем: нам − меньше проверять, вам — проще найти ошибку или дополнить эксперимент. При проверке оценивается четкость ответов на вопросы, аккуратность отчета и кода.
#
# ### Оценивание
# Каждая из задач имеет определенную «стоимость» (указана в скобках около задачи). Максимально допустимая оценка за работу — 10 баллов. Сдавать задание после указанного срока сдачи нельзя.
# ## Данные
#
#
# В этой лабораторной работе мы научимся обучать модели машинного обучения, корректно ставить эксперименты, подбирать гиперпараметры, сравнивать и смешивать модели. Вам предлагается решить задачу бинарной классификации, а именно построить алгоритм, определяющий превысит ли средний заработок человека порог $50k. Каждый объект выборки — человек, для которого известны следующие признаки:
# - age
# - workclass
# - fnlwgt
# - education
# - education-num
# - marital-status
# - occupation
# - relationship
# - race
# - sex
# - capital-gain
# - capital-loss
# - hours-per-week
#
# Более подробно про признаки можно почитать [здесь](http://archive.ics.uci.edu/ml/machine-learning-databases/adult/adult.names). Целевой признак записан в переменной *>50K,<=50K*.
# ## Метрика качества
#
# Обучение и оценка качества модели производится на независимых множествах примеров. Как правило, имеющиеся примеры разбивают на два подмножества: обучение (train) и контроль (test). Выбор пропорции разбиения — компромисс. Действительно, большой размер обучения ведет к более качественным алгоритмам, но бОльшему шуму в оценке модели на контроле. И наоборот, большой размер контрольной выборки ведет к менее шумной оценке качества, однако модели получаются менее точными.
#
# Многие модели классификации получают оценку принадлежности положительному классу $\tilde{y}(x) \in R$ (например, вероятность принадлежности классу 1). Затем принимают решение о классе объекта путем сравнения оценки с некоторым порогом $\theta$:
#
# $y(x) = +1$, если $\tilde{y}(x) \geq \theta$, $y(x) = -1$, если $\tilde{y}(x) < \theta$
#
# В этом случае можно рассматривать метрики, которые умеют работать с исходным ответом классификатора. В задании мы будем работать с метрикой AUC-ROC, которую в данном случае можно считать как долю неправильно упорядоченных пар объектов, отсортированных по возрастанию предсказанной оценки принадлежности классу 1 (более подробно можно узнать [здесь](https://github.com/esokolov/ml-course-msu/blob/master/ML15/lecture-notes/Sem05_metrics.pdf)). Детального понимания принципов работы метрики AUC-ROC для выполнения лабораторной не требуется.
# ## Подбор гиперпараметров модели
#
# В задачах машинного обучения следует различать параметры модели и гиперпараметры (структурные параметры). Обычно параметры модели настраиваются в ходе обучения (например, веса в линейной модели или структура решающего дерева), в то время как гиперпараметры задаются заранее (например, регуляризация в линейной модели или максимальная глубина решающего дерева). Каждая модель обычно имеет множество гиперпараметров, и нет универсальных наборов гиперпараметров, оптимально работающих во всех задачах, для каждой задачи нужно подбирать свой набор.
#
# Для оптимизации гиперпараметров модели часто используют _перебор по сетке (grid search)_: для каждого гиперпараметра выбирается несколько значений, перебираются все комбинации значений и выбирается комбинация, на которой модель показывает лучшее качество (с точки зрения метрики, которая оптимизируется). Однако в этом случае нужно грамотно оценивать построенную модель, а именно делать разбиение на обучающую и тестовую выборку. Есть несколько схем, как это можно реализовать:
#
# - Разбить имеющуюся выборку на обучающую и тестовую. В этом случае сравнение большого числа моделей при переборе параметров приводит к ситуации, когда лучшая на тестовой подвыборке модель не сохраняет свои качества на новых данных. Можно сказать, что происходит _переобучение_ на тестовую выборку.
# - Для устранения описанной выше проблемы, можно разбить данные на 3 непересекающихся подвыборки: обучение (`train`), валидация (`validation`) и контроль (`test`). Валидационную подвыборку используют для сравнения моделей, а `test` — для окончательной оценки качества и сравнения семейств моделей с подобранными параметрами.
# - Другой способ сравнения моделей — [кросс-валидация](http://en.wikipedia.org/wiki/Cross-validation_(statistics). Существуют различные схемы кросс-валидации:
# - Leave-One-Out
# - K-Fold
# - Многократное случайное разбиение выборки
#
# Кросс-валидация вычислительно затратна, особенно если вы делаете перебор по сетке с очень большим числом комбинаций. С учетом конечности времени на выполнение задания возникает ряд компромиссов:
# - сетку можно делать более разреженной, перебирая меньше значений каждого параметра; однако, надо не забывать, что в таком случае можно пропустить хорошую комбинацию параметров;
# - кросс-валидацию можно делать с меньшим числом разбиений или фолдов, но в таком случае оценка качества кросс-валидации становится более шумной и увеличивается риск выбрать неоптимальный набор параметров из-за случайности разбиения;
# - параметры можно оптимизировать последовательно (жадно) — один за другим, а не перебирать все комбинации; такая стратегия не всегда приводит к оптимальному набору;
# - перебирать не все комбинации параметров, а небольшое число случайно выбранных.
# ## Задание
#
# Прежде чем приступать к заданию, рекомендуем прочитать его до конца.
# Загрузите набор данных *data.adult.csv*. Чтобы лучше понимать, с чем вы работаете/корректно ли вы загрузили данные, можно вывести несколько первых строк на экран.
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
url = 'https://archive.ics.uci.edu/ml/machine-learning-databases/adult/adult.data'
data = pd.read_csv(url, sep=',')
data.head()
data=pd.read_csv("data.adult.csv")
data.head()
# Иногда в данных встречаются пропуски. Как задаются пропуски, обычно либо прописывается в описании к данным, либо просто на месте пропуска после чтения данных оказывается значение numpy.nan. Более подробно о работе с пропусками в Pandas можно прочитать например [здесь](http://pandas.pydata.org/pandas-docs/stable/missing_data.html).
#
# В данном датасете пропущенные значения обозначены как "?".
#
# **(0.5 балла)** Найдите все признаки, имеющие пропущенные значения. Удалите из выборки все объекты с пропусками.
data.columns
s = 0
for i in data.columns:
#print(data[i])
s+=1
print(s)
data.shape
for i in data.columns:
data[i]
data[' Never-married'].isin(['?']).unique()
data.head()
for i in data.columns:
if len(data[i].isin(['?']).unique())==2: #isin выводит список правды-неправды, unique их посчитает, и если есть и то, и то, то длина ответа два (если только неправды - 1)
data[i].replace('?',np.nan,inplace=True)
data[i].dropna(inplace=True)
print(i)
# Обычно после загрузки датасета необходима его некоторая предобработка. В данном случае она будет заключаться в следующем:
#
# - Выделите целевую переменную (ту, которую мы хотим предсказывать) в отдельную переменную, удалите ее из датасета и преобразуйте к бинарному формату.
# - Обратите внимание, что не все признаки являются вещественными (числовыми). В начале мы будем работать только с вещественными признаками. Выделите их отдельно.
label=pd.factorize(data[' <=50K'])[0]
data.drop([' <=50K'],axis = 1, inplace = True)
data.head()
data.dtypes
intcolumns=[]
for i in data.columns:
if data[i].dtype.name=='int64':
intcolumns.append(i)
intcolumns
# ## Обучение классификаторов на вещественных признаках
#
# В данном разделе необходимо будет работать только с вещественными признаками и целевой переменной.
#
# В начале посмотрим, как работает подбор параметров по сетке и как влияет на качество разбиение выборки. Сейчас и далее будем рассматривать 4 алгоритма:
# - [kNN](http://scikit-learn.org/stable/modules/generated/sklearn.neighbors.KNeighborsClassifier.html)
# - [DecisonTree](http://scikit-learn.org/stable/modules/generated/sklearn.tree.DecisionTreeClassifier.html#sklearn.tree.DecisionTreeClassifier)
# - [RandomForest](http://scikit-learn.org/stable/modules/generated/sklearn.ensemble.RandomForestClassifier.html)
# - [SGD Linear Classifier](http://scikit-learn.org/stable/modules/generated/sklearn.linear_model.SGDClassifier.html)
#
# Для начала у первых трёх алгоритмов выберем один гиперпараметр, который будем оптимизировать:
# - kNN — число соседей (*n_neighbors*)
# - DecisonTree — глубина дерева (*max_depth*)
# - SGD Linear Classifier — оптимизируемая функция (*loss*)
#
# Остальные параметры оставляйте в значениях по умолчанию. Для подбора гиперпараметров воспользуйтесь перебором по сетке, который реализован в классе [GridSearchCV](http://scikit-learn.org/stable/modules/generated/sklearn.grid_search.GridSearchCV.html#sklearn.grid_search.GridSearchCV). В качестве схемы кросс-валидации используйте 5-fold cv, которую можно задать с помощью класса [KFoldCV](http://scikit-learn.org/stable/modules/generated/sklearn.cross_validation.KFold.html).
#
# **(1.5 балла)** Для каждого алгоритма подберите оптимальные значения указанных гиперпараметров. Постройте график среднего качества по кросс-валидации алгоритма при заданном значении гиперпараметра, на котором также отобразите доверительный интервал.
from sklearn.neighbors import KNeighborsClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn import linear_model
from sklearn.model_selection import GridSearchCV
knn = KNeighborsClassifiereighborsClassifier()
params = {'n_neighbors':np.arange(GridSearchCV50)}
clf_knn = GridSearchCV(knn, params, cv = 5)
clf_knn.fit(data[intcolumns], label)
knn = KNeighborsClassifier()
params={'n_neighbors':np.arange(1,40)}
clf_knn = GridSearchCV(knn, params,cv=5)
clf_knn.fit(data[intcolumns], label)
# +
#clf_knn.cv_results_
# -
#чтоб не томить, лучшее значение ищется вот так
print(clf_knn.cv_results_['param_n_neighbors'][clf_knn.cv_results_['mean_test_score'].argmax()])
print(clf_knn.cv_results_['mean_test_score'].argmax())
#clf_knn.cv_results_['mean_test_score'].max()
x = params['n_neighbors']
y = clf_knn.cv_results_['mean_test_score']
err = clf_knn.cv_results_['std_test_score']
plt.plot(x, y)
plt.fill_between(x, y-err, y+err, facecolor = 'red')
x,y,err=params['n_neighbors'],clf_knn.cv_results_['mean_test_score'],clf_knn.cv_results_['std_test_score']
plt.plot(x,y,'b')
plt.xlabel('n neighbors')
plt.ylabel('mean test score')
plt.fill_between(x,y-err,y+err,facecolor='green')
plt.show()
# Лучшее значение параметра по кроссвалидации:
clf_knn.cv_results_['param_n_neighbors'][clf_knn.cv_results_['mean_test_score'].argmax()],clf_knn.cv_results_['mean_test_score'].max()
# Качество на кросвалидации стабилизируется и падает после оптимального значения параметра.
tree = DecisionTreeClassifier(random_state=0)
params={'max_depth':np.arange(1,30)}
clf_tree = GridSearchCV(tree, params,cv=5)
clf_tree.fit(data[intcolumns], label)
x,y,err=params['max_depth'],clf_tree.cv_results_['mean_test_score'],clf_tree.cv_results_['std_test_score']
plt.plot(x,y,'b')
plt.xlabel('max depth')
plt.ylabel('mean test score')
plt.fill_between(x,y-err,y+err,facecolor='green')
plt.show()
x = params['max_depth']
y = clf_tree.cv_results_['mean_test_score']
err = clf_tree.cv_results_['std_test_score']
plt.plot(x, y)
plt.fill_between(x, y-err, y+err, facecolor = 'red')
clf_tree.cv_results_['param_max_depth'][clf_tree.cv_results_['mean_test_score'].argmax()]
clf_tree.cv_results_['mean_test_score'].max()
# Лучшее значение параметра по кроссвалидации:
clf_tree.cv_results_['param_max_depth'][clf_tree.cv_results_['mean_test_score'].argmax()],clf_tree.cv_results_['mean_test_score'].max()
# Сравним random forest c decision tree
randomcoffee = RandomForestClassifier()
params = {'max_depth': np.arange(1, 51)}
clf_rf = GridSearchCV(randomcoffee, params, cv=5)
clf_rf.fit(data[intcolumns], label)
rf = RandomForestClassifier()
params={'max_depth':np.arange(1,50)}
clf_rf = GridSearchCV(rf, params,cv=5)
clf_rf.fit(data[intcolumns], label)
[clf_rf.cv_results_['mean_test_score'].argmax()]
clf_rf.cv_results_['param_max_depth'][clf_rf.cv_results_['mean_test_score'].argmax()]
clf_rf.cv_results_['mean_test_score'].max()
x = params['max_depth']
y = clf_rf.cv_results_['mean_test_score']
err = clf_rf.cv_results_['std_test_score']
plt.plot(x, y)
plt.fill_between(x, y-err, y+err)
x,y,err=params['max_depth'],clf_rf.cv_results_['mean_test_score'],clf_rf.cv_results_['std_test_score']
plt.plot(x,y,'b')
plt.xlabel('max depth')
plt.ylabel('mean test score')
plt.fill_between(x,y-err,y+err,facecolor='green')
plt.show()
# Лучшее значение параметра по кроссвалидации:
clf_rf.cv_results_['param_max_depth'][clf_rf.cv_results_['mean_test_score'].argmax()],clf_rf.cv_results_['mean_test_score'].max()
# Desion tree и random forest на тесте ухудшают качество после оптимального параметра.
# Наверное, глубокие деревья переобучаются, а на трейне результаты лучше именно у них.
clf_tree.cv_results_['param_max_depth'][clf_tree.cv_results_['mean_train_score'].argmax()]
clf_tree.cv_results_['mean_train_score'].max()
clf_rf.cv_results_['param_max_depth'][clf_rf.cv_results_['mean_train_score'].argmax()]
clf_rf.cv_results_['mean_train_score'].max()
# +
sgd = linear_model.SGDClassifier()
params={'loss': ('hinge', 'log', 'modified_huber')}
clf_sgd = GridSearchCV(sgd, params,cv=5)
clf_sgd.fit(data[intcolumns], label)
# -
x,y,err=range(len(params['loss'])),clf_sgd.cv_results_['mean_test_score'],clf_sgd.cv_results_['std_test_score']
plt.xticks(x, params['loss'])
plt.plot(x,y,'b')
plt.xlabel('loss')
plt.ylabel('mean test score')
plt.fill_between(x,y-err,y+err,facecolor='green')
plt.show()
x = params['loss']
y = clf_sgd.cv_results_['mean_test_score']
err = clf_sgd.cv_results_['std_test_score']
plt.plot(x, y)
plt.fill_between(x,y-err, y+err)
# Лучшее значение параметра по кроссвалидации:
clf_sgd.cv_results_['param_loss'][clf_sgd.cv_results_['mean_test_score'].argmax()],clf_sgd.cv_results_['mean_test_score'].max()
# Оптимальный параметр обладает наименьшей дисперсией на тесте.
# # Что вы можете сказать о получившихся графиках?
#
# **(1 балл)** Также подберём число деревьев (*n_estimators*) в алгоритме RandomForest. Как известно, в общем случае Random Forest не переобучается с увеличением количества деревьев, так что при увеличении этого гиперпараметра его качество не будет становиться хуже. Поэтому подберите такое количество деревьев, при котором качество на кросс-валидации стабилизируется. Обратите внимание, что для проведения этого эксперимента не нужно с нуля обучать много случайных лесов с различными количествами деревьев. Обучите один случайный лес с максимальным интересным количеством деревьев, а затем рассмотрите подмножества разных размеров, состоящие из деревьев построенного леса (поле [*estimators_*](http://scikit-learn.org/stable/modules/generated/sklearn.ensemble.RandomForestClassifier.html)). В дальнейших экспериментах используйте это количество деревьев.
from sklearn.model_selection import KFold
from sklearn.metrics import accuracy_score
import random
# +
rf = RandomForestClassifier(n_estimators=1000,max_depth=12)
kf = KFold(n_splits=5)
h=10
score_kf=np.zeros(rf.n_estimators//h)
score_kf_max=np.zeros(rf.n_estimators//h)
score_kf_min=np.ones(rf.n_estimators//h)
for train_index, test_index in kf.split(data[intcolumns]):
print("TRAIN:", train_index, "TEST:", test_index)
X_train, X_test = data[intcolumns].loc[train_index], data[intcolumns].loc[test_index]
y_train, y_test = label[train_index], label[test_index]
rf.fit(X_train,y_train)
score=[]
for j in range(1,rf.n_estimators//h):
proba = sum(list(map(lambda x: x.predict_proba(X_test),random.sample(rf.estimators_,j*h))))/(j*h)
pred=rf.classes_.take(np.argmax(proba, axis=1), axis=0)
score=np.append(score,accuracy_score(y_test,pred))
score=np.append(score,rf.score(X_test,y_test))
score_kf+=score
score_kf_max=np.maximum(score,score_kf_max)
score_kf_min=np.minimum(score,score_kf_min)
score_kf/=kf.n_splits
# -
x,y=range(1,rf.n_estimators//h+1),score_kf
plt.plot(x,y,'b')
plt.xticks(x[0::10], tuple(map(str,list(np.array(x)*h)))[0::10])
plt.xlabel('n estimators')
plt.ylabel('mean test score')
plt.fill_between(x,score_kf_min,score_kf_max,facecolor='green')
plt.show()
# Качество на кроссвалидации стабилизируется после примерно 300 деревьев.
# При обучении алгоритмов стоит обращать внимание не только на качество, но и каким образом они работают с данными. В этой задаче получилось так, что некоторые из используемых алгоритмов чувствительны к масштабу признаков. Чтобы убедиться, что это как-то могло повлиять на качество, давайте посмотрим на сами признаки.
#
# **(0.5 балла)** Постройте гистограммы для признаков *age*, *fnlwgt*, *capital-gain*.
# +
features=['age', 'fnlwgt', 'capital-gain']
fig, axes = plt.subplots(3,1,figsize=(15,15))
params = dict(bins=150)
for i in range(3):
axes[i].hist(data[features[i]],**params)
axes[i].set_title(features[i])
plt.show()
# -
# **(0.5 балла)** Глядя на получившиеся графики, объясните, в чем заключается особенность данных? На какие алгоритмы это может повлиять? Может ли масшитабирование повлиять на работу этих алгоритмов?
# Данные различны по масштабу. Максимальное значение age 90, а значения fnlwgt и capital-gain могут быть сотнями тысяч. Значения capital-gain сконцентрированны в пределах 20000.
# Различия масштабов признаков не влияют на построение решающих деревьев (на decision tree и random forest), так как в вершине дерева происходит разбиение по конкретному признаку независимо от остальных.
# Но на алгоритм knn эти различия влияют, так как расстояние до объектов сильно зависит от признаков, принимающих большие значения.
# Так же различие масштабов влияет на линейные классификаторы, оптимизируемые с помощью sgd, (такие как логистическая регрессия, svm), так как масштаб признаков влияет на веса w линеной модели (веса соответствующие большим признакам могут принимать маленькие значения) и на градиент функции ошибки.
#
# Масштабирование признаков можно выполнить, например, одним из следующих способов:
# - $x_{new} = \dfrac{x - \mu}{\sigma}$, где $\mu, \sigma$ — среднее и стандартное отклонение значения признака по всей выборке (см. функцию [scale](http://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.scale.html))
# - $x_{new} = \dfrac{x - x_{min}}{x_{max} - x_{min}}$, где $[x_{min}, x_{max}]$ — минимальный интервал значений признака
#
# **(1 балл)** Масштабируйте все вещественные признаки одним из указанных способов и подберите оптимальные значения гиперпараметров аналогичо пункту выше.
data_norm=(data[intcolumns]-data[intcolumns].mean())/data[intcolumns].std()
knn = KNeighborsClassifier()
params={'n_neighbors':np.arange(1,40)}
clf_knn = GridSearchCV(knn, params,cv=5)
clf_knn.fit(data_norm, label)
x,y,err=params['n_neighbors'],clf_knn.cv_results_['mean_test_score'],clf_knn.cv_results_['std_test_score']
plt.plot(x,y,'b')
plt.xlabel('n neighbors')
plt.ylabel('mean test score')
plt.fill_between(x,y-err,y+err,facecolor='green')
plt.show()
# Лучшее значение параметра по кроссвалидации:
clf_knn.cv_results_['param_n_neighbors'][clf_knn.cv_results_['mean_test_score'].argmax()],clf_knn.cv_results_['mean_test_score'].max()
tree = DecisionTreeClassifier(random_state=0)
params={'max_depth':np.arange(1,30)}
clf_tree = GridSearchCV(tree, params,cv=5)
clf_tree.fit(data_norm, label)
x,y,err=params['max_depth'],clf_tree.cv_results_['mean_test_score'],clf_tree.cv_results_['std_test_score']
plt.plot(x,y,'b')
plt.xlabel('max depth')
plt.ylabel('mean test score')
plt.fill_between(x,y-err,y+err,facecolor='green')
plt.show()
# Лучшее значение параметра по кроссвалидации:
clf_tree.cv_results_['param_max_depth'][clf_tree.cv_results_['mean_test_score'].argmax()],clf_tree.cv_results_['mean_test_score'].max()
rf = RandomForestClassifier()
params={'max_depth':np.arange(1,50)}
clf_rf = GridSearchCV(rf, params,cv=5)
clf_rf.fit(data_norm, label)
x,y,err=params['max_depth'],clf_rf.cv_results_['mean_test_score'],clf_rf.cv_results_['std_test_score']
plt.plot(x,y,'b')
plt.xlabel('max depth')
plt.ylabel('mean test score')
plt.fill_between(x,y-err,y+err,facecolor='green')
plt.show()
# Лучшее значение параметра по кроссвалидации:
clf_rf.cv_results_['param_max_depth'][clf_rf.cv_results_['mean_test_score'].argmax()],clf_rf.cv_results_['mean_test_score'].max()
sgd = linear_model.SGDClassifier()
params={'loss': ('hinge', 'log', 'modified_huber')}
clf_sgd = GridSearchCV(sgd, params,cv=5)
clf_sgd.fit(data_norm, label)
x,y,err=range(len(params['loss'])),clf_sgd.cv_results_['mean_test_score'],clf_sgd.cv_results_['std_test_score']
plt.xticks(x, params['loss'])
plt.plot(x,y,'b')
plt.xlabel('loss')
plt.ylabel('mean test score')
plt.fill_between(x,y-err,y+err,facecolor='green')
plt.show()
# Лучшее значение параметра по кроссвалидации:
clf_sgd.cv_results_['param_loss'][clf_sgd.cv_results_['mean_test_score'].argmax()],clf_sgd.cv_results_['mean_test_score'].max()
# Изменилось ли качество у некоторых алгоритмов?
# Результаты классификаторов knn и sgd изменились в лучшую сторону. На решающие деревья нормализация не повлияла.
# **(1,5 балла)** Теперь сделайте перебор нескольких гиперпараметров по сетке и найдите оптимальные комбинации (лучшее среднее значение качества) для каждого алгоритма в данном случае:
# - KNN — число соседей (*n_neighbors*) и метрика (*metric*)
# - DecisonTree — глубина дерева (*max_depth*) и критерий разбиения (*criterion*)
# - RandomForest — критерий разбиения в деревьях (*criterion*) и *max_features* (при фиксированном количестве деревьев, найденном ранее)
# - SGDClassifier — оптимизируемая функция (*loss*) и *penalty*
#
# Обратите внимание, что эта операция может быть ресурсо- и трудоемкой. Как оптимизировать подбор параметров по сетке, сказано в разделе "Подбор гиперпараметров модели"
knn = KNeighborsClassifier()
params=[{'n_neighbors':np.arange(1,20),'metric':['minkowski','manhattan','chebyshev']}]
clf_knn = GridSearchCV(knn, params,cv=5)
clf_knn.fit(data_norm, label)
params_knn=clf_knn.cv_results_['params'][clf_knn.cv_results_['mean_test_score'].argmax()]
params_knn,clf_knn.cv_results_['mean_test_score'].max()
tree = DecisionTreeClassifier(random_state=0)
params={'max_depth':np.arange(1,15),'criterion':['gini','entropy']}
clf_tree = GridSearchCV(tree, params,cv=5)
clf_tree.fit(data_norm, label)
params_tree=clf_tree.cv_results_['params'][clf_tree.cv_results_['mean_test_score'].argmax()]
params_tree,clf_tree.cv_results_['mean_test_score'].max()
rf = RandomForestClassifier(n_estimators=300,max_depth=11)
params={'criterion':['gini','entropy'],'max_features':np.arange(1,len(data_norm.columns))}
clf_rf = GridSearchCV(rf, params,cv=5)
clf_rf.fit(data_norm, label)
params_rf=clf_rf.cv_results_['params'][clf_rf.cv_results_['mean_test_score'].argmax()]
params_rf,clf_rf.cv_results_['mean_test_score'].max()
sgd = linear_model.SGDClassifier()
params={'loss': ['hinge', 'log', 'modified_huber'],'penalty':['none', 'l2', 'l1', 'elasticnet']}
clf_sgd = GridSearchCV(sgd, params,cv=5)
clf_sgd.fit(data_norm, label)
params_sgd=clf_sgd.cv_results_['params'][clf_sgd.cv_results_['mean_test_score'].argmax()]
params_sgd,clf_sgd.cv_results_['mean_test_score'].max()
# Какой из алгоритмов имеет наилучшее качество?
#
# **(0.5 балла)** Сравните алгоритмы с точки зрения времени обучения. Обучение какого из алгоритмов работает дольше всего и, как вы думаете, почему?
clf_knn.cv_results_['mean_fit_time'].mean(),clf_tree.cv_results_['mean_fit_time'].mean(),clf_rf.cv_results_['mean_fit_time'].mean(),clf_sgd.cv_results_['mean_fit_time'].mean()
# Лучшее качество имеют решающие деревья (random forest и decision tree).
# Дольше всего работает random forest, так как строится 300 решающих деревьев. Второй по длительности работы knn, так как для каждого объекта считаются расстояния до других объектов для нахождения ближайших соседей. Линейная модель и decision tree работают быстро.
# ## Добавление категориальных признаков в модели
#
# Пока мы не использовали нечисловые признаки, которые есть в датасете. Давайте посмотрим, правильно ли мы сделали и увеличится ли качество моделей после добавления этих признаков.
#
# **(0.5 балла)** Преобразуйте все категориальные признаки с помощью метода one-hot-encoding (например, это можно сделать с помощью функции [pandas.get_dummies](http://pandas.pydata.org/pandas-docs/stable/generated/pandas.get_dummies.html) или [DictVectorizer](http://scikit-learn.org/stable/modules/generated/sklearn.feature_extraction.DictVectorizer.html) из sklearn).
categories=pd.get_dummies(data[list(set(data.columns)-set(intcolumns))])
categories.head()
# Так как после кодирования признаков получилось достаточно много, в этой работе мы не будем добавлять их и подбирать заново оптимальные гиперпараметры.
#
# **(0.5 балла)** Добавьте к нормированным вещественным признакам закодированные категориальные и обучите алгоритмы с наилучшими гиперпараметрами из предыдущего пункта. Дало ли добавление новых признаков прирост качества? Измеряйте качество как и раньше используя 5-Fold CV. Для этого удобно воспользоваться функцией [cross_val_score](http://scikit-learn.org/stable/modules/generated/sklearn.cross_validation.cross_val_score.html#sklearn.cross_validation.cross_val_score).
from sklearn.cross_validation import cross_val_score
data=pd.concat([data_norm,categories],axis=1)
data.head()
knn = KNeighborsClassifier(metric=params_knn['metric'],n_neighbors=params_knn['n_neighbors'])
(cross_val_score(knn, data, label,cv=5)).mean()
tree = DecisionTreeClassifier(max_depth=params_tree['max_depth'],criterion=params_tree['criterion'])
(cross_val_score(tree, data, label,cv=5)).mean()
rf = RandomForestClassifier(n_estimators=300,max_depth=11,criterion=params_rf['criterion'],max_features=params_rf['max_features'])
(cross_val_score(rf, data, label,cv=5)).mean()
sgd = linear_model.SGDClassifier(loss=params_sgd['loss'],penalty=params_sgd['penalty'])
(cross_val_score(sgd, data, label,cv=5)).mean()
# Отличается ли теперь наилучший классификатор от наилучшего в предыдущем пункте?
# Да, есть прирост качества у всех классификаторов, в том числе и у деревьев.
# ## Смешивание моделей
#
# Во всех предыдущих пунктах мы получили много сильных моделей, которые могут быть достаточно разными по своей природе (например, метод ближайших соседей и случайный лес). Часто на практике оказывается возможным увеличить качество предсказани путем смешивания подобных разных моделей. Давайте посмотрим, действительно ли это дает прирост в качестве.
#
# Выберете из построенных моделей двух предыдущих пунктов две, которые дали наибольшее начество на кросс-валидации (обозначим их $clf_1$ и $clf_2$). Далее постройте новый классификатор, ответ которого на некотором объекте $x$ будет выглядеть следующим образом:
#
# $$result(x) = clf_1(x) * \alpha + clf_2(x) * (1 - \alpha)$$
#
# где $\alpha$ — гиперпараметр нового классификатора.
#
# **(1 балл)** Подберите по сетке от 0 до 1 $\alpha$ для этого классификатора с помощью 5-Fold CV и постройте график качества в зависимости от $\alpha$ (аналогичный графику в разделе "Обучение классификаторов и оценка качества"). Дал ли этот подход прирост к качеству по сравнению с моделями ранее?
# Лучшее качество показали random forest и decision tree. Берем комбинации предсказанных вероятностей и по полученной вероятности строим ответ.
# +
m1 = rf
m2 = tree
k=5
score_kf=np.zeros(k+1)
score_kf_max=np.zeros(k+1)
score_kf_min=np.ones(k+1)
kf = KFold(n_splits=5)
for train_index, test_index in kf.split(data[intcolumns]):
print("TRAIN:", train_index, "TEST:", test_index)
X_train, X_test = data.loc[train_index], data.loc[test_index]
y_train, y_test = label[train_index], label[test_index]
score=[]
for alpha in range(k+1):
m1.fit(X_train,y_train)
m2.fit(X_train,y_train)
pred=(m1.predict(X_test) * alpha/k + m2.predict(X_test) * (1 - alpha/k))>=0.5
score=np.append(score,accuracy_score(y_test,pred))
score_kf+=score
score_kf_max=np.maximum(score,score_kf_max)
score_kf_min=np.minimum(score,score_kf_min)
score_kf/=kf.n_splits
# -
x,y=range(k+1),score_kf
plt.plot(x,y,'b')
plt.xticks(x, tuple(map(str,list(np.array(x)/k))))
plt.xlabel('alpha')
plt.ylabel('mean test score')
plt.fill_between(x,score_kf_min,score_kf_max,facecolor='green')
plt.show()
# Оптимальное значение alpha:
(np.array(x)/k)[score_kf.argmax()],score_kf.max()
# Теперь попробуем смешать непохожие классификаторы: decision tree и knn.
# +
m1 = knn
m2 = tree
k=5
score_kf=np.zeros(k+1)
score_kf_max=np.zeros(k+1)
score_kf_min=np.ones(k+1)
kf = KFold(n_splits=5)
for train_index, test_index in kf.split(data[intcolumns]):
print("TRAIN:", train_index, "TEST:", test_index)
X_train, X_test = data.loc[train_index], data.loc[test_index]
y_train, y_test = label[train_index], label[test_index]
score=[]
for alpha in range(k+1):
m1.fit(X_train,y_train)
m2.fit(X_train,y_train)
pred=(m1.predict(X_test) * alpha/k + m2.predict(X_test) * (1 - alpha/k))>=0.5
score=np.append(score,accuracy_score(y_test,pred))
score_kf+=score
score_kf_max=np.maximum(score,score_kf_max)
score_kf_min=np.minimum(score,score_kf_min)
score_kf/=kf.n_splits
# -
x,y=range(k+1),score_kf
plt.plot(x,y,'b')
plt.xticks(x, tuple(map(str,list(np.array(x)/k))))
plt.xlabel('alpha')
plt.ylabel('mean test score')
plt.fill_between(x,score_kf_min,score_kf_max,facecolor='green')
plt.show()
# Оптимальное значение alpha:
(np.array(x)/k)[score_kf.argmax()],score_kf.max()
# Смешивание random forest и decision tree увеличило качество. При смешивании knn и decision tree, knn не получил веса.
# ## Сравнение построенных моделей
#
#
# **(1 балл)** Сделайте общие итоговые выводы о классификаторах с точки зрения их работы с признаками и сложности самой модели (какие гиперпараметры есть у модели, сильно ли изменение значения гиперпараметра влияет на качество модели).
# KNN
# Количество ближайших соседей - важный параметр. При небольших значениях (1,2) классификатор неустойчив к шуму, при большом классификатор уделяет внимание далеким объектам и предсказывает для всех объектов похожие значения. Как видно из графиков этот параметр влияет на качество модели. Очень важен и выбор метрики. Для большинства метрик нужна нормированность признаков. Манхэттенская метрика дала хороший прирост качества. Но подбор метрики не поддается каким-то конкретным правилам, лучше выбирать ее эмпирически среди частоиспользуемых. Кроме нормированности признаков стоит следить за их количеством, так как при большом числе сильно вырастает размерность, и все объекты становятся одинаково далеки, выбор ближайших соседей трудно осуществим.
# Decision Tree Для построения дерева важный параметр - его максимальная глубина, на графиках было видно как она влияет на тестовый результат. Не очень глубоким деревом сложно классифицировать объекты, а слишком глубокое дерево переобучится. Так же оптимизировался параметр criterion - критерий неоднородности, который используется при построении дерева (характеризует долю класса в вершине, максимален при равенстве долей классов, то есть описательную способность вершины). Изменение этого параметра не сыграло большой роли, так что можно или оставлять значение по кмолчанию, или подбирать эмпирически. В случае деревьев признаки можно не нормировать. При очень большом количестве признаков не все они могут быть задействованы, так как глубина дерева ограничена. Зато решающие деревья позволяют оценить значимость признаков.
# Random Forest О нормированности, глубине и критерии неоднородности можно сказать то же самое. Параметр n estimators (количество голосующих деревьев) важен: чем больше деревьев, тем надежнее предсказанный ответ. Но этот параметр влияет на время работы классификатора, так что нужно найти его значение, при котором ошибка на тесте стабилизируется. Так же оптимизировался параметр max features (количество признаков в дереве). Он сильно влияет на качество ответа, так как при его большом значении деревья могут оказаться "похожими", возможно переобучение, и увеличивается время построения дерева.
# SGD Для линейных классификаторов оптимизировалась функция ошибки, я выбирала между svm, логистической регрессией и персептроном ('hinge', 'log', 'perceptron'). Этот параметр, естественно, очень влияет на результат, так как это разные подходы к оценке качества. Так же оптимизировался параметр, отвечающий за регуляризацию. Этот параметр тоже лучше оптимизировать, так как, например, l1 регуляризация может занулить веса перед признаками, а l2 - нет, это может оказаться важным. В случае линейных классификаторов данные нужно нормировать.
| L_02.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import os
import sys
import random
import math
import re
import time
import numpy as np
import cv2
import matplotlib
import matplotlib.pyplot as plt
import tensorflow as tf
from mrcnn.config import Config
#import utils
from mrcnn import model as modellib,utils
from mrcnn import visualize
import yaml
from mrcnn.model import log
from PIL import Image
# +
ROOT_DIR = os.path.abspath("")
# Directory to save logs and trained model
MODEL_DIR = os.path.join(ROOT_DIR, "logs")
print(MODEL_DIR)
iter_num=0
# Local path to trained weights file
COCO_MODEL_PATH = os.path.join(ROOT_DIR, "mask_rcnn_coco.h5")
# Download COCO trained weights from Releases if needed
if not os.path.exists(COCO_MODEL_PATH):
utils.download_trained_weights(COCO_MODEL_PATH)
class ShapesConfig(Config):
"""Configuration for training on the toy shapes dataset.
Derives from the base Config class and overrides values specific
to the toy shapes dataset.
"""
# Give the configuration a recognizable name
NAME = "shape"
# Train on 1 GPU and 8 images per GPU. We can put multiple images on each
# GPU because the images are small. Batch size is 8 (GPUs * images/GPU).
GPU_COUNT = 1
IMAGES_PER_GPU = 2
# Number of classes (including background)
NUM_CLASSES = 1 + 2 # background + 6 shapes
# Use small images for faster training. Set the limits of the small side
# the large side, and that determines the image shape.
IMAGE_MIN_DIM = 320
IMAGE_MAX_DIM = 384
# Use smaller anchors because our image and objects are small
RPN_ANCHOR_SCALES = (8 * 6, 16 * 6, 32 * 6, 64 * 6, 128 * 6) # anchor side in pixels
# Reduce training ROIs per image because the images are small and have
# few objects. Aim to allow ROI sampling to pick 33% positive ROIs.
TRAIN_ROIS_PER_IMAGE = 100
# Use a small epoch since the data is simple
STEPS_PER_EPOCH = 100
# use small validation steps since the epoch is small
VALIDATION_STEPS = 50
config = ShapesConfig()
# config.display()
# -
class DrugDataset(utils.Dataset):
# 得到该图中有多少个实例(物体)
def get_obj_index(self, image):
n = np.max(image)
return n
# 解析labelme中得到的yaml文件,从而得到mask每一层对应的实例标签
def from_yaml_get_class(self, image_id):
info = self.image_info[image_id]
with open(info['yaml_path']) as f:
temp = yaml.load(f.read())
labels = temp['label_names']
del labels[0]
return labels
# 重新写draw_mask
def draw_mask(self, num_obj, mask, image,image_id):
#print("draw_mask-->",image_id)
#print("self.image_info",self.image_info)
info = self.image_info[image_id]
#print("info-->",info)
#print("info[width]----->",info['width'],"-info[height]--->",info['height'])
for index in range(num_obj):
for i in range(info['width']):
for j in range(info['height']):
#print("image_id-->",image_id,"-i--->",i,"-j--->",j)
#print("info[width]----->",info['width'],"-info[height]--->",info['height'])
at_pixel = image.getpixel((i, j))
if at_pixel == index + 1:
mask[j, i, index] = 1
return mask
# 重新写load_shapes,里面包含自己的类别,可以任意添加
# 并在self.image_info信息中添加了path、mask_path 、yaml_path
# yaml_pathdataset_root_path = "/tongue_dateset/"
# img_floder = dataset_root_path + "rgb"
# mask_floder = dataset_root_path + "mask"
# dataset_root_path = "/tongue_dateset/"
def load_shapes(self, count, img_floder, mask_floder, imglist, dataset_root_path):
"""Generate the requested number of synthetic images.
count: number of images to generate.
height, width: the size of the generated images.
"""
# Add classes,可通过这种方式扩展多个物体
self.add_class("shapes", 1, "baby") # 黑色素瘤
self.add_class("shapes", 2, "quilt")
for i in range(count):
# 获取图片宽和高
filestr = imglist[i].split(".")[0]
#print(imglist[i],"-->",cv_img.shape[1],"--->",cv_img.shape[0])
#print("id-->", i, " imglist[", i, "]-->", imglist[i],"filestr-->",filestr)
#filestr = filestr.split("_")[1]
mask_path = mask_floder + "/" + filestr + ".png"
yaml_path = dataset_root_path + "labelme_json/rgb_" + filestr + "_json/info.yaml"
# print(dataset_root_path + "labelme_json/" + filestr + "_json/img.png")
cv_img = cv2.imread(dataset_root_path + "pic/" + filestr + ".png")
self.add_image("shapes", image_id=i, path=img_floder + "/" + imglist[i],
width=cv_img.shape[1], height=cv_img.shape[0], mask_path=mask_path, yaml_path=yaml_path)
# 重写load_mask
def load_mask(self, image_id):
"""Generate instance masks for shapes of the given image ID.
"""
global iter_num
print("image_id",image_id)
info = self.image_info[image_id]
count = 1 # number of object
img = Image.open(info['mask_path'])
num_obj = self.get_obj_index(img)
mask = np.zeros([info['height'], info['width'], num_obj], dtype=np.uint8)
mask = self.draw_mask(num_obj, mask, img,image_id)
occlusion = np.logical_not(mask[:, :, -1]).astype(np.uint8)
for i in range(count - 2, -1, -1):
mask[:, :, i] = mask[:, :, i] * occlusion
occlusion = np.logical_and(occlusion, np.logical_not(mask[:, :, i]))
labels = []
labels = self.from_yaml_get_class(image_id)
labels_form = []
for i in range(len(labels)):
if labels[i].find("baby") != -1:
labels_form.append("baby")
elif labels[i].find("quilt") != -1:
labels_form.append("quilt")
class_ids = np.array([self.class_names.index(s) for s in labels_form])
return mask, class_ids.astype(np.int32)
# +
dataset_root_path=ROOT_DIR+"/quilt_data/"
img_floder = dataset_root_path + "pic"
# print(img_floder)
mask_floder = dataset_root_path + "cv2_mask"
#yaml_floder = dataset_root_path
imglist = os.listdir(img_floder)
count = len(imglist)
print(imglist)
print(count)
#train与val数据集准备
dataset_train = DrugDataset()
dataset_train.load_shapes(count, img_floder, mask_floder, imglist, dataset_root_path)
dataset_train.prepare()
dataset_val = DrugDataset()
dataset_val.load_shapes(10, img_floder, mask_floder, imglist, dataset_root_path)
dataset_val.prepare()
# -
image_ids = np.random.choice(dataset_train.image_ids, 4)
for image_id in image_ids:
image = dataset_train.load_image(image_id)
mask, class_ids = dataset_train.load_mask(image_id)
# visualize.display_top_masks(image, mask,
# class_ids, dataset_train.class_names)
print("dataset_val-->",dataset_val._image_ids)
image_ids = np.random.choice(dataset_val.image_ids, 4)
for image_id in image_ids:
image = dataset_train.load_image(image_id)
mask, class_ids = dataset_train.load_mask(image_id)
# visualize.display_top_masks(image, mask, class_ids, dataset_train.class_names)
# +
model = modellib.MaskRCNN(mode="training", config=config,
model_dir=MODEL_DIR)
init_with = "coco" # imagenet, coco, or last
if init_with == "imagenet":
model.load_weights(model.get_imagenet_weights(), by_name=True)
elif init_with == "coco":
# Load weights trained on MS COCO, but skip layers that
# are different due to the different number of classes
# See README for instructions to download the COCO weights
model.load_weights(COCO_MODEL_PATH, by_name=True,
exclude=["mrcnn_class_logits", "mrcnn_bbox_fc",
"mrcnn_bbox", "mrcnn_mask"])
elif init_with == "last":
# Load the last model you trained and continue training
model.load_weights(model.find_last()[1], by_name=True)
# -
model.train(dataset_train, dataset_val,
learning_rate=config.LEARNING_RATE/10,
epochs=120,
layers='heads')
# +
# Save weights
# Typically not needed because callbacks save after every epoch
# Uncomment to save manually
import os
MODEL_DIR = os.path.join(ROOT_DIR, "logs")
model_2_path = os.path.join(MODEL_DIR, "shape20181022T1535")
model_path = os.path.join(model_2_path, "mask_rcnn_shape_0027.h5")
print(model_path)
model.keras_model.save_weights(model_path)
| samples/shapes/train_quilt.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a href="https://colab.research.google.com/github/adasegroup/ML2021_seminars/blob/master/seminar8/Multiclass_Imbalanced_solutions.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# # Advanced classification: Imbalanced and Multi-class cases
# In this seminar we will learn how to perform classification in case of multiple balanced or imbalanced classes.
#
# The dataset, which we will use for this tutorial, is the smaller version [Stanford Dogs Dataset](http://vision.stanford.edu/aditya86/ImageNetDogs/). The initial dataset consists from images of 120 breeds of dogs. In our case we are going to use just 4 classes out of those 120.
# 
# #### The plan of the seminar:
# * a small introduction to Stanford Dogs Dataset
# * Producing the features of the images using the pretrained neural network (we will consider it as a black box)
# * Multi-class classification methods: One-vs-One and One-vs-Rest
# * Imbalanced dataset - why is it a problem?
# * Imbalanced classification methods: Over and Under-Sampling, SMOTE
# Let us start with some library imports.
# ##### NOTES:
# * Class description
# * dataframe creation in class or in seminar
# !pip install -U imbalanced-learn
# !wget https://github.com/adasegroup/ML2021_seminars/raw/main/seminar8/data/dog_breeds.zip
# !unzip -oqd "./" "dog_breeds.zip"
# !ls .
# !rm -rf ./__MACOSX ./sample_data .config ./dog_breeds.zip
import torch
import pandas as pd
import matplotlib.pyplot as plt
import urllib
# %matplotlib inline
from PIL import Image
from torchvision import transforms
import os
import sklearn
import os.path
from tqdm.autonotebook import tqdm
# +
#if you load your data from the local directory
#################################
#path_doggies ="dog_breeds/small"
#paths_doggies = [path_doggies +'/'+ i for i in os.listdir(path_doggies) if '.DS_' not in i]
#################################
# -
paths_doggies = [i for i in os.listdir('./') if '.DS_' not in i]
# <br>
# Now let us have a look at the data
def img_show(img, ax, title = None):
"""
Plots the image on the particular axis
Parameters
----------
img: Image,image to plot.
axis: matplotlib axis to plot on.
title: string, the title of the image
"""
ax.imshow(img)
ax.axis('off')
if title:
ax.set_title(title)
#images for plotting
img_names = {}
for num, i in enumerate(paths_doggies[:4]):
img_names.update({i.split('-')[-1]:paths_doggies[num]+'/'+os.listdir(i)[0]})
#plot the images from img_names
fig, ax = plt.subplots(1,4, figsize=(20,10))
k = 0
for i, key in enumerate(img_names.keys()):
img_show(Image.open(img_names[key]), ax[i], title = key)
plt.show()
# In order to make working with the data much easier, we are going to create a class, that will store the ```image_to_features``` model, the ```data_list```, containing all the vectors of features of the image samples and the ```data_path```.
class DogBreedDataset:
def __init__(self, data_path, feature_generator, num_samples=None):
"""
A wrapper class for Stanford Dog Breeds dataset.
Parameters
----------
data_path: string, the path to the dataset.
feature_generator: torch.nn.Module, the model, that receives the torch.tensor of the preprocessed image
as the input and produces the tensor of features as the output.
num_samples: integer, the number of samples in each class to load, default: None.
"""
self.data_path = data_path
self.model = feature_generator
self.num_samples = num_samples
self.data_list = []
def preprocess_image(self, image):
"""
Opens and preprocesses an Image according to the requirements mentioned at https://pytorch.org/hub/pytorch_vision_vgg/
Parameters
----------
path: the path to the image.
img_name: the name of the image file.
Returns
-------
input_tensor: the tensor of the preprocessed image.
input_batch: input_tensor with an extra dim, representing a batch
"""
preprocess = transforms.Compose([
transforms.Resize(256),
transforms.CenterCrop(224),
transforms.ToTensor(),
transforms.Normalize(mean=[0.485, 0.456, 0.406],
std=[0.229, 0.224, 0.225]),
])
input_tensor = preprocess(image)
input_batch = input_tensor.unsqueeze(0)
return input_batch
def load_dataset(self):
"""
Loads and preprocesses the images from the dataset
Parameters
----------
path: the path to the image.
img_name: the name of the image file.
Returns
-------
data_list: the list of vectors of features of dogs' images
"""
data_list = []
for path in tqdm(self.data_path):
counter = 0
for filename in tqdm(os.listdir(path)):
counter += 1
# input
with open(os.path.join(path, filename), 'rb') as file:
batch = self.preprocess_image(Image.open(file))
with torch.no_grad():
features = self.model(batch).flatten().cpu().numpy()
# label
_, label = path.split('-', 1)
data_list.append((features, label))
if counter >= self.num_samples:
break
return data_list
# The model that we are going to use to get our features from this raw images is the Neural Network called **VGG-11** (you are going to learn about these types of NN models later in this course).
# Lucky for us, [```PyTorch```](https://pytorch.org) library stores some of the most popular [pretrained Neural Networks](https://pytorch.org/hub/), so we don't have to design and train the VGG-11 NN from sctratch.
# +
import ssl
ssl._create_default_https_context = ssl._create_unverified_context
# -
#dowbload the VGG11 model from pytorch hub
model = torch.hub.load('pytorch/vision:v0.4.0', 'vgg11', pretrained=True)
# 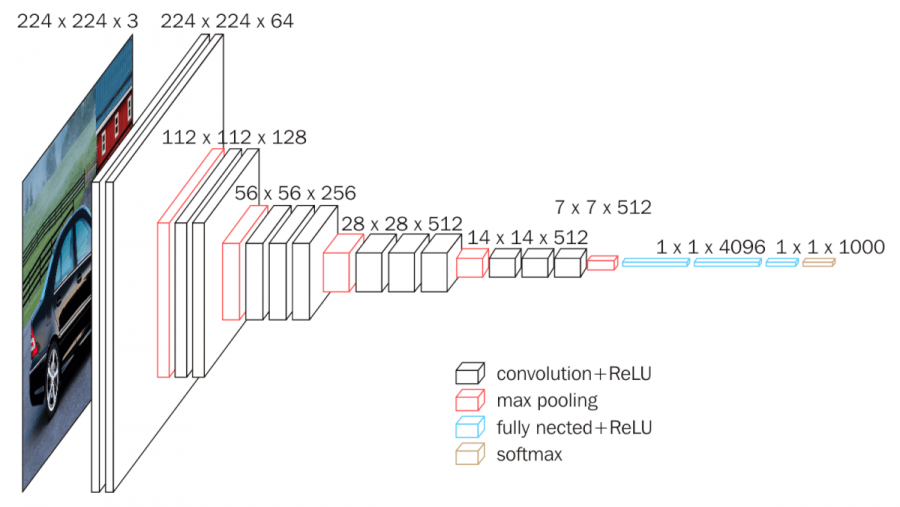
# However, we do not need the whole network for producing the images' features - we will take only the part of it, just before the first __fully connected__ layer.
#take only the "head" that outputs the images' features
image_to_feats = model.features
image_to_feats.eval()
# Let's have a look at our data:
# Let us download, preprocess and store the features of the images in a pandas dataframe
dataset_class = DogBreedDataset(paths_doggies, image_to_feats, num_samples = 150)
datalist = dataset_class.load_dataset()
# Let's create a pandas dataframe with all the features and labels.
# +
#derive the list of labels from the paths_doggies list
#take the data_list and after each list of features add the data label
#create a list of column names
#create a df_doggies dataframe
# -
features, label = datalist[0]
# +
columns = [f"feat_{i+1}" for i in range(len(features))]
df_doggies = pd.DataFrame(
[feat for feat, lab in datalist],
columns=columns)
df_doggies["y"] = [lab for feat, lab in datalist]
# -
df_doggies.shape
df_doggies.head()
# Turn the labels to Categorical type and create the dictionary, in case we would like to recover the original labels
df_doggies.y = pd.Categorical(df_doggies.y)
label_map = dict(enumerate(df_doggies.y.cat.categories) )
label_map
df_doggies.y = df_doggies.y.cat.codes
# ## Plotting the data using dimensinonality reduction techniques
# DataPlotter is another blackbox that we are going to use for representing our features in a more convenient way for plotting (later in the course you will learn about PCA and TSNE).
#
# Let's plot our data!
# +
from sklearn.manifold import TSNE
import seaborn as sns
from sklearn.decomposition import PCA
class DataPlotter:
def __init__(self, data, dim_red = 'pca', X=None, y=None):
"""
A wrapper class for dimensionality reduction and plotting.
Parameters
----------
data_path: dataframe, the dataset.
dim_red: string, the dimensionality reduction technique to use, either 'tsne' or 'pca'.
"""
self.data = data
self.dim_red = dim_red
self.X = X
self.y = y
if X is None:
self.X = self.data.loc[:, self.data.columns!='y']
if y is None:
self.y = self.data.y.astype(int)
def shuffle_data(self):
"""
Randomly shuffling the data.
"""
self.X = self.X.sample(frac=1).reset_index(drop=True)
self.y = self.y.sample(frac=1).reset_index(drop=True)
def reduce_dimension(self):
"""
Reduce the current dimension of the feature data to 2 dimensions using either pca or tsne.
"""
if self.dim_red =='tsne':
self.X_embedded = TSNE(n_components=2, perplexity=30.0).fit_transform(self.X)
elif self.dim_red == 'pca':
self.X_embedded = PCA(n_components=2).fit_transform(self.X)
def plot_data(self):
plt.figure(figsize=(20,10))
sns.scatterplot(self.X_embedded[:,0], self.X_embedded[:,1], hue = self.y, palette="rainbow", s=100,
legend = "full")
# -
data_pltr = DataPlotter(df_doggies, dim_red = 'pca')
data_pltr.reduce_dimension()
data_pltr.plot_data()
# ## Multi-class classification
# Finally, let's try some multi-clall classification methods.
from sklearn.model_selection import train_test_split
from sklearn.base import clone
# Train-test split
# +
y = df_doggies.y.astype(int)
X = df_doggies.loc[:, df_doggies.columns!='y']
split = train_test_split(X, y, test_size=0.5,
random_state=42, stratify=y)
train_X, test_X, train_y, test_y = split
# -
# Most of the binary classification methods that you have already discussed in the previous seminars, unfortuntelly, only allow to distinguish one class from the other. However, in our case, we want to classify several dog breeds, so how can we do that?
# One way to this problem is using **One-vs-All** approach:
# 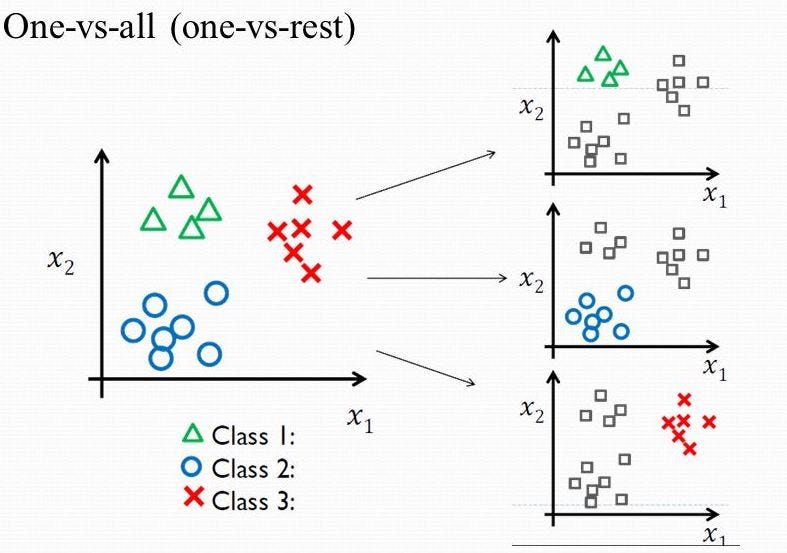
# +
from sklearn.svm import LinearSVC
# from sklearn.linear_model import LogisticRegression
model_SVC = LinearSVC(random_state=0)
# model_LogReg = LogisticRegression(random_state=0)
# +
from sklearn.multiclass import OneVsRestClassifier
ovr_classifier = OneVsRestClassifier(clone(model_SVC), n_jobs=-1)
ovr_classifier.fit(train_X, train_y)
# -
from sklearn.metrics import confusion_matrix
# +
predict_y = ovr_classifier.predict(test_X)
cmatrix = confusion_matrix(test_y, predict_y)
pd.DataFrame(cmatrix)
# -
# rows -- fact
# columns -- predict
print("Accuracy %.3f%%" % (100 * ovr_classifier.score(test_X, test_y)))
# ### One-vs-One approach to multi-class classification
# 
# +
from sklearn.multiclass import OneVsOneClassifier
ovo_classifier = OneVsOneClassifier(clone(model_SVC))
ovo_classifier.fit(train_X, train_y)
# -
predict_y = ovo_classifier.predict(test_X)
print("Accuracy %.3f%%" % (100 * ovo_classifier.score(test_X, test_y)))
pd.DataFrame(confusion_matrix(test_y, predict_y))
# ## Imbalanced data
# Data imbalance is a very common problem for many machine learning problems. Consider volcano erruption, or plane crush prediction - there is an abundance of negative examples, when the event does not happen and very little recorded cases of the events, the occurence of which we want to predict.
# This is where various methods of class balancing is going to help.
# +
# df_doggies_subset = df_doggies1.loc[df_doggies1.y.isin([0,1,2,3])]
# -
X_sub, y_sub = df_doggies.loc[:, df_doggies.columns!='y'], df_doggies.y.astype(int)
data_pltr = DataPlotter(df_doggies, dim_red = 'pca')
data_pltr.reduce_dimension()
data_pltr.plot_data()
from collections import Counter
print('Distribution before imbalancing: {}'.format(Counter(y_sub)))
from imblearn.datasets import make_imbalance
X_res, y_res = make_imbalance(
X_sub, y_sub, sampling_strategy={0: 150, 1: 150, 2: 30, 3: 150},
random_state=1)
print('Distribution after imbalancing: {}'.format(Counter(y_res)))
data_pltr = DataPlotter(df_doggies, dim_red = 'pca', X = X_res, y = y_res)
data_pltr.reduce_dimension()
data_pltr.plot_data()
split = train_test_split(X_res, y_res, test_size=0.3,
random_state=42, stratify=y_res)
train_X, test_X, train_y, test_y = split
from sklearn.linear_model import RidgeClassifier
model_SVC = LinearSVC(random_state=50)
#model_SVC = RidgeClassifier(random_state=0)
ovr_classifier = OneVsRestClassifier(clone(model_SVC), n_jobs=-1)
ovr_classifier.fit(train_X, train_y)
predictions = ovr_classifier.predict(test_X)
#predictions = model_SVC.predict(test_X[test_y==0])
from imblearn.metrics import classification_report_imbalanced
print("Accuracy %.3f%%" % (100 * ovr_classifier.score(test_X, test_y)))
print(classification_report_imbalanced(test_y, predictions))
# ## Techniques to try, when dealing with the imabalanced dataset:
# * Under/Over Sampling
# * Synthetic minority over-sampling technique and its variants (ADASYN, BorderlineSMOTE, etc)
from imblearn.over_sampling import RandomOverSampler, SMOTE
from imblearn.under_sampling import RandomUnderSampler
balancer = RandomUnderSampler()
balanced_train_x, balanced_train_y = balancer.fit_resample(train_X, train_y)
print('Distribution before balancing: {}'.format(Counter(train_y)))
print('Distribution after balancing: {}'.format(Counter(balanced_train_y)))
model = LinearSVC(random_state=50, )
ovr_classifier = OneVsRestClassifier(clone(model), n_jobs=-1)
ovr_classifier.fit(balanced_train_x, balanced_train_y)
print("Accuracy %.3f%%" % (100 * ovr_classifier.score(test_X, test_y)))
predictions = ovr_classifier.predict(test_X)
#predictions = model.predict(test_X[test_y==0])
print(classification_report_imbalanced(test_y, predictions))
pd.DataFrame(confusion_matrix(test_y, predictions))
balancer = RandomOverSampler()
balanced_train_x, balanced_train_y = balancer.fit_resample(train_X, train_y)
print('Distribution after balancing: {}'.format(Counter(balanced_train_y)))
model = LinearSVC(random_state=50)
ovr_classifier = OneVsRestClassifier(clone(model), n_jobs=-1)
ovr_classifier.fit(balanced_train_x, balanced_train_y)
print("Accuracy %.3f%%" % (100 * ovr_classifier.score(test_X, test_y)))
predictions = ovr_classifier.predict(test_X)
pd.DataFrame(confusion_matrix(test_y, predictions))
print(classification_report_imbalanced(test_y, predictions))
# ## SMOTE
# <img src="https://ars.els-cdn.com/content/image/1-s2.0-S0950705119302898-gr1.jpg" alt="smote" width="600"/>
rebalancer = SMOTE(sampling_strategy='not majority', k_neighbors=5, random_state = 1)
under_balancer = RandomUnderSampler(sampling_strategy={0:20, 1: 30, 3:50}, random_state = 1)
balanced_train_x, balanced_train_y = under_balancer.fit_resample(train_X, train_y)
print('Distribution before balancing: {}'.format(Counter(balanced_train_y)))
# +
model = LinearSVC(random_state=0)
ovr_classifier = OneVsRestClassifier(clone(model))
X_SMOTE, y_SMOTE = rebalancer.fit_resample(balanced_train_x, balanced_train_y)
print('Distribution after balancing: {}'.format(Counter(y_SMOTE)))
ovr_classifier = ovr_classifier.fit(X_SMOTE, y_SMOTE)
# -
predict_y_balanced = ovr_classifier.predict(test_X)
pd.DataFrame(confusion_matrix(test_y, predict_y_balanced))
print("Accuracy %.3f%%" % (100 * ovr_classifier.score(test_X, test_y)))
print(classification_report_imbalanced(test_y, predict_y_balanced))
data_pltr = DataPlotter(df_doggies, dim_red = 'pca', X = X_SMOTE, y = y_SMOTE)
data_pltr.reduce_dimension()
data_pltr.plot_data()
# There are different variations of SMOTE method, such as ADASYN, BalancedSMOTE etc. Many of them are avaliable in [```imblearn```](https://imbalanced-learn.readthedocs.io/en/stable/api.html) library.
#
# **Try out those methods yourself, using the mentioned methods, plot and analyze the results.**
# +
from imblearn.over_sampling import BorderlineSMOTE, ADASYN, SVMSMOTE
rebalancer_list = [BorderlineSMOTE(),ADASYN(n_neighbors = 3), SVMSMOTE()]
model = LinearSVC(random_state=0)
for rebalancer in rebalancer_list:
ovr_classifier = OneVsRestClassifier(clone(model))
X_SMOTE, y_SMOTE = rebalancer.fit_resample(train_X, train_y)
print('Distribution after balancing: {}'.format(Counter(y_SMOTE)))
ovr_classifier = ovr_classifier.fit(X_SMOTE, y_SMOTE)
predict_y_balanced = ovr_classifier.predict(test_X)
print(classification_report_imbalanced(test_y, predict_y_balanced))
data_pltr = DataPlotter(df_doggies, dim_red = 'pca', X = X_SMOTE, y = y_SMOTE)
data_pltr.reduce_dimension()
data_pltr.plot_data()
# -
| seminar8/Multiclass_Imbalanced_solutions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# # "Getting Started with Audio data in Python"
# > "In this article, I am will be explaining some of the modules which can be used to load and manipulate some audio files in python."
#
# - toc: true
# - branch: master
# - badges: true
# - comments: true
# - categories: [data, audio, file, sound, functions, wave, dockship, article, speech, recognition]
# - hide: false
# Nowadays Audio data is also becoming more and more common in the field of Data Science.
#
# In this article, I am going to explain how to load, manipulate and store audio data in python.
#
# There are different kinds of audio formats:
#
# 1. mp3
# 2. wav
# 3. flac
# 4. m4a
#
# Audio data is stored and measured in frequency(Hz)
#
# 1KHz means 1000 bytes/units of information stored per second
#
# In a small sample of the audio clip, there are some thousands of byte information.
#
# Let's dive into some code
# +
# Required modules
import wave
import numpy as np
from matplotlib import pyplot as plt
# -
# filename is the name of the audio file which needs to be loaded
# +
# Loading audio file as wave object
gm_wave = wave.open(filename, 'r')
# +
# Converting wave object into bytes
gm_bytes = gm_wave.readframes(-1)
# +
# Output of the variable
gm_bytes
# Output
# b'\x04\xbb\x05\x86\t\x10\x06\x82\r\xe4\x06\xda\x0e...'
# -
# The output of the sound variable is in bytes which is now not in human readable form. So let's convert this into human readable form.
#
# For that we can use numpy module's frombuffer function.
# +
# Changing into bytes
signal_gm = np.frombuffer(gm_bytes, dtype='int16')
signal_gm[:10]
# Output
# array([ -3, -5, -8, -8, -9, -13, -8, -10, -9, -11], dtype=int16)
# -
# The wave object also has so many functions which can be used to get the characteristics of the audio file, they are getframerate(), getnchannels(), getsamplewidth(), getnframes() etc.
#
# Frame rate is the number of frequency bytes in one second.
# +
# Get the frame rate
framerate_gm = gm_wave.getframerate()
# Show the frame rate
framerate_gm
# Output
# 8000
# -
# **Now it's time to visualize the sound wave.**
#
# Let's get the timestamp values of the audio file, for that we are using np.linspace() function to create a evenly spaced numpy array
#
# Syntax: np.linspace(start, stop, step) which creates step number of floating point values in between start and stop.
# +
# Creating timestamp values
time_gm = np.linspace(start=0, stop=len(signal_gm)/framerate_gm, step=len(signal_gm))
time_gm[:10]
# Output
# array([0.00000000e+00, 1.25000117e-04, 2.50000233e-04, ...,
1.34118250e+02, 1.34118375e+02, 1.34118500e+02])
# -
# The above code creates a time stamp array values. last value of the array is the duration of the audio clip.
#
# Plotting the audio file
# +
plt.title("Audio Clip")
plt.plot(time_gm, signal_gm)
# x and y axis labels
plt.xlabel("Time(s)")
plt.ylabel("Amplitude")
# show our plot
plt.show()
# -
# Apart from wave there are also some other third party libraries for processing the audio data.
#
# 1. CMU Sphinx
# 2. Speech Recognition
# 3. Kaldi
# 4. Wav2letter++
# +
# Import the module
import speech_recognition as sr
# Create the instance of recognizer
recog = sr.Recognizer()
# Set the limit of the energy
recog.energy_threshold = 350
# -
# Recognizer class contains many of the built-in functions to convert audio into text data
#
# 1. recognize_bing()
# 2. recognize_google()
# 3. recognize_google_cloud()
# 4. recognize_ibm()
# 5. recognize_wit()
# 6. recognize_houndify()
# Input: Audio file
#
# Output: Transcribed text
#
# **Note:** Some of the api calls require credentials
# Translate using google api
text = recog.recognize_google(audio_data=audio_file, language='en-US')
# if the audio file which you are passing to the recognize function is of different language, then specify the corresponding language to the second argument.
#
# Else it will print the text in the English language.
#
# Creating a AudioFile class using sr module
# +
# Read in audio file
audio = sr.AudioFile()
# Check type of audio
type(audio)
# Output
#
# -
# If we try to pass the audio variable to any one of the recognize function, it will throw and error. As the recognize functions accept only the audio_data input.
#
# In this case, we need to convert the AudioFile to AudioData
# +
# Convert from AudioFile to AudioData
with audio as src:
audio_data = recognizer.record(src)
# Check the type
type(audio_data)
# Output
#
# -
# In the above code snippet, the record function records the audio file in the form of audio_data.
# +
# Leave duration and offset as default
with clean_support_call as source:
clean_support_call_audio = recognizer.record(source, duration=None, offset=None)
# -
# The record function also takes two other arguments which is duration and offset.
#
# Duration specifies the time for which the audio data should be recorded and the offset specifies the beginning byte from which the function start capturing the audio data.
#
# Audio file can also be of non-speech data i.e. a roar of the lion, barking of the dog etc.
#
# In this case, if you pass the audio file, you will get a **UnknownValueError**
# +
# Import the leopard roar audio file
leopard_roar = sr.AudioFile("leopard_roar.wav")
# Convert the AudioFile to AudioData
with leopard_roar as source:
leopard_roar_audio = recognizer.record(source)
# Recognize the AudioData
recognizer.recognize_google(leopard_roar_audio)
# -
# If you have trouble in hearing the audio file, the api will also have trouble.
# +
# Import audio file with background nosie
noisy_support_call = sr.AudioFile(noisy_support_call.wav)
with noisy_support_call as source:
# Adjust for ambient noise and record
recognizer.adjust_for_ambient_noise(source,duration=0.5)
noisy_support_call_audio = recognizer.record(source)
# Recognize the audio
recognizer.recognize_google(noisy_support_call_audio)
# -
# The above code removes the noise from the audio clip by hearing the audio clip by listening to it for 0.5 duration.
| _notebooks/2021-02-27-Getting-Started-with-Audio-data-in-Python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# hvPlot is written to work well inside a Jupyter notebook, from the interactive Python command prompt, or inside a Python batch script. In this user guide we will discover how to use hvPlot to view plots in each of these cases and how to save the plots to a separate file.
#
# ## Notebook
#
# In a Jupyter notebook, hvPlot will return HoloViews objects that display themselves using Bokeh, as long as the extension has been loaded. The easiest way of loading the extension is to import one of the plotting extensions such as `hvplot.pandas`:
import hvplot
import hvplot.pandas
# Here we will load some of the sample data and then compose the HoloViews objects into a layout:
# +
from hvplot.sample_data import airline_flights, us_crime
violent_crime = us_crime.hvplot(x='Year', y='Violent Crime rate', width=400)
burglaries = us_crime.hvplot(x='Year', y='Burglary rate', width=400)
violent_crime + burglaries
# -
# To display the object from inside a function we can instead call the ``display`` function. The ``display`` function also supports an ``display_id`` in the notebook (for JupyterLab and classic Jupyter Notebook versions >5.5), which allows us to obtain a handle for the plot:
handle = display(violent_crime, display_id='example')
# This display handle may then be updated with another HoloViews object replacing the plot:
#
# ```python
# handle.update(burglaries)
# ```
# ## Converting to bokeh
#
# In some cases it can be convenient to construct a plot with ``hvPlot`` and then convert it to a bokeh model to embed in a more complex plot. This can be achieved using HoloViews' ``render`` function, e.g. we can convert the `violent_crime` plot from above into a bokeh `Figure`, which can be composed into a more complex figure as needed:
# +
import holoviews as hv
hv.render(violent_crime)
# -
# ## Python Command Prompt & Scripts
#
# When working outside the notebook we can instead use the ``hvplot.show`` function, which will open the plot in a new browser window:
#
# <img src="../assets/console.png" style="display: table; margin: 0 auto;"></img>
#
# For static plots this will simply save a temporary file and open it, however for dynamic and [datashaded](http://datashader.org) plots it will automatically launch a Bokeh server, enabling all the dynamic features.
#
# <img src="../assets/console_server.gif" style="display: table; margin: 0 auto;"></img>
# ## Saving plots
#
# When looking at any Bokeh plot in a web browser, you can use the toolbar's "Save" tool to export the plot as a PNG (try it on one of the plots above!).
#
# hvPlot also provides a convenient ``save`` function to export HoloViews objects to a file. By default it will save the plot as HTML:
# +
hex_bins = airline_flights.hvplot.hexbin(x='airtime', y='arrdelay', colorbar=True, width=600, height=500)
hvplot.save(hex_bins, 'test.html')
# -
# By default, the HTML file generated will depend on loading JavaScript code for BokehJS from the online CDN repository, to reduce the file size. If you need to work in an airgapped or no-network environment, you can declare that `INLINE` resources should be used instead of `CDN`:
from bokeh.resources import INLINE
hvplot.save(hex_bins, 'test.html', resources=INLINE)
# Finally, if a 'png' file extension is specified, the exported plot will be rendered as a PNG, which currently requires Selenium and PhantomJS to be installed:
hvplot.save(hex_bins, 'test.png')
| examples/user_guide/Viewing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Progress Reporting and Command Observers
#
# SimpleITK `Filter`s and other classes derived from `ProcessObject`s have the ability for user code to be executed when certain events occur. This is known as the Command and Observer design patters to implement user callbacks. This allows for the monitoring and abortion of processes as they are being executed.
#
# Consider the following image source which takes a few seconds to execute. It would be nice to quickly know how long your going to need to wait, to know if you can go get a cup of coffee.
# +
from __future__ import print_function
# %matplotlib inline
import matplotlib.pyplot as plt
import SimpleITK as sitk
print(sitk.Version())
import sys
import os
import threading
from myshow import myshow
from myshow import myshow3d
# -
size=256 # if this is too fast increase the size
img = sitk.GaborSource(sitk.sitkFloat32, size=[size]*3, sigma=[size*.2]*3, mean=[size*0.5]*3, frequency=.1)
myshow3d(img,zslices=[int(size/2)],dpi=40);
myshow(img);
# We need to add a command to display the progress reported by the `ProcessObject::GetProgress` method during the `sitkProgressEvent`. This involves three components:
#
# 1. Events
# 2. ProcessObject's methods
# 3. Commands
#
# We'll look at some examples after a brief explanation of these components.
#
# ## Events
#
# The avaiable events to observed are defined in a namespace enumeration.
#
# <table>
# <tr><td>sitkAnyEvent</td><td>Occurs for all event types.</td></tr>
# <tr><td>sitkAbortEvent</td><td>Occurs after the process has been aborted, but before exiting the Execute method.</td></tr>
# <tr><td>sitkDeleteEvent</td><td>Occurs when the underlying itk::ProcessObject is deleted.</td></tr>
# <tr><td>sitkEndEvent</td><td>Occurs at then end of normal processing.</td></tr>
# <tr><td>sitkIterationEvent</td><td>Occurs with some algorithms that run for a fixed or undetermined number of iterations.</td></tr>
# <tr><td>sitkProgressEvent</td><td>Occurs when the progress changes in most process objects.</td></tr>
# <tr><td>sitkStartEvent</td><td>Occurs when then itk::ProcessObject is starting.</td></tr>
# <tr><td>sitkUserEvent</td><td>Other events may fall into this enumeration.</td></tr>
# </table>
#
# The convention of pre-fixing enums with "sitk" is continued, although it's getting a little crowded.
#
# C++ is more strongly typed than Python it allows for implicit conversion from an enum type to an int, but not from an int to an enum type. Care needs to be made to ensure the correct enum value is passed in Python.
#
# ## ProcessObject's methods
#
# To be able to interface with the `ProcessObject` during execution, the object-oriented interface must be used to access the method of the ProcessObject. While any constant member function can be called during a command call-back there are two common methods:
#
# 1. `ProcessObject::GetProgress()`
# 2. `ProcessObject::Abort()`
#
# The methods are only valid during the `Command` while a process is being executed, or when the process is not in the `Execute` method.
#
# Additionally it should be noted that follow methods can *not* be called during a command or from another thread during execution `Execute` and `RemoveAllCommands`. In general the `ProcessObject` should not be modified during execution.
#
#
# ## Commands
#
# The command design pattern is used to allow user code to be executed when an event occurs. It is implemented in the `Command` class. The `Command` class provides an `Execute` method to be overridden in derived classes.
#
# There are three ways to define a command with SimpleITK in Python.
#
# 1. Derive from the `Command` class.
# 2. Use the `PyCommand` class' `SetCallbackPyCallable` method.
# 3. Use an inline `lambda` function in `ProcessOject::AddCommand`.
help(sitk.Command)
# +
class MyCommand(sitk.Command):
def __init__(self):
# required
super(MyCommand,self).__init__()
def Execute(self):
print("MyCommand::Execute Called")
cmd = MyCommand()
cmd.Execute()
# + slideshow={"slide_type": "-"}
help(sitk.PyCommand)
# -
cmd = sitk.PyCommand()
cmd.SetCallbackPyCallable( lambda: print("PyCommand Called") )
cmd.Execute()
# Back to watching the progress of out Gabor image source. First lets create the filter as an object
size=256
filter = sitk.GaborImageSource()
filter.SetOutputPixelType(sitk.sitkFloat32)
filter.SetSize([size]*3)
filter.SetSigma([size*.2]*3)
filter.SetMean([size*0.5]*3)
filter.SetFrequency(.1)
img = filter.Execute()
myshow3d(img,zslices=[int(size/2)],dpi=40);
# ## The ProcessObject interface for the Invoker or Subject
#
# SimpleITK doesn't have a large heirachy of inheritance. It has been kept to a minimal, so there is no common `Object` or `LightObject` base class as ITK has. As most of the goals for the events have to do with observing processes, the "Subject" interface of the Observer patter or the "Invoker" part of the Command design pattern, has been added to a `ProcessObject` base class for filters.
#
# The `ProcessObject` base class has the following methods of handling commands: `AddCommand`, `RemoveAllCommands`, and `HasCommand`.
#
# Adding these functionalities are not available in the procedural interface available for SimpleITK. They are only available through the Object Oriented interface, and break the method chaining interface.
help(sitk.ProcessObject)
# ### Deriving from the `Command` class
# The traditional way of using Commands in ITK involves deriving from the `Command` class and adding to the `ProcessObject`.
class MyCommand(sitk.Command):
def __init__(self, msg):
# required
super(MyCommand,self).__init__()
self.msg = msg
def __del__(self):
print("MyCommand begin deleted: \"{0}\"".format(self.msg))
def Execute(self):
print(self.msg)
cmd1 = MyCommand("Start")
cmd2 = MyCommand("End")
filter.RemoveAllCommands() # this line is here so we can easily re-execute this code block
filter.AddCommand(sitk.sitkStartEvent, cmd1)
filter.AddCommand(sitk.sitkEndEvent, cmd2)
filter.Execute()
# A reference to the `Command` object must be maintained, or else it will be removed from the `ProcessObject`.
filter.AddCommand(sitk.sitkStartEvent, MyCommand("stack scope"))
print("Before Execution")
filter.Execute()
# ### Using a `labmda` function as the `Command`
# In Python the `AddCommand` has been extended to accept `PyCommand` objects and implicitly creates a `PyCommand` from a callable python argument. This is **really** useful.
filter.RemoveAllCommands() # this line is here so we can easily re-execute this code block
filter.AddCommand(sitk.sitkStartEvent, lambda: print("Starting...",end=''))
filter.AddCommand(sitk.sitkStartEvent, lambda: sys.stdout.flush())
filter.AddCommand(sitk.sitkEndEvent, lambda: print("Done"))
filter.Execute()
# ## Access to ITK data during command execution
# The commands are not too useful unless you can query the filter through the SimpleITK interface. A couple status variables and methods are exposed in the SimpleITK `ProcessObject` through the polymorphic interface of the same ITK class.
filter.RemoveAllCommands()
filter.AddCommand(sitk.sitkProgressEvent, lambda: print("\rProgress: {0:03.1f}%...".format(100*filter.GetProgress()),end=''))
filter.AddCommand(sitk.sitkProgressEvent, lambda: sys.stdout.flush())
filter.AddCommand(sitk.sitkEndEvent, lambda: print("Done"))
filter.Execute()
# # Utilizing Jupyter Notebooks and Commands
#
# Utilization of commands and events frequently occurs with advanced integration into graphical user interfaces. Let us now export this advanced integration into Jupyter Notebooks.
#
# Jupyter notebooks support displaying output as HTML, and execution of javascript on demand. Together this can produce animation.
# +
import uuid
from IPython.display import HTML, Javascript, display
divid = str(uuid.uuid4())
html_progress="""
<p style="margin:5px">FilterName:</p>
<div style="border: 1px solid black;padding:1px;margin:5px">
<div id="{0}" style="background-color:blue; width:0%%"> </div>
</div>
""".format(divid)
def command_js_progress(processObject):
p = processObject.GetProgress()
display(Javascript("$('div#%s').width('%i%%')" % (divid, int(p*100))))
# +
filter.RemoveAllCommands()
filter.AddCommand(sitk.sitkStartEvent, lambda: display(HTML(html_progress)))
filter.AddCommand(sitk.sitkProgressEvent, lambda: command_js_progress(filter))
filter.Execute()
# -
# ### Support for Bi-direction JavaScript
#
# It's possible to get button in HTML to execute python code...
# +
import uuid
from IPython.display import HTML, Javascript, display
g_Abort = False
divid = str(uuid.uuid4())
html_progress_abort="""
<div style="background-color:gainsboro; border:2px solid black;padding:15px">
<p style="margin:5px">FilterName:</p>
<div style="border: 1px solid black;padding:1px;margin:5px">
<div id="{0}" style="background-color:blue; width:0%%"> </div>
</div>
<button onclick="set_value()" style="margin:5px" >Abort</button>
</div>
""".format(divid)
javascript_abort = """
<script type="text/Javascript">
function set_value(){
var command = "g_Abort=True"
console.log("Executing Command: " + command);
var kernel = IPython.notebook.kernel;
kernel.execute(command);
}
</script>
"""
def command_js_progress_abort(processObject):
p = processObject.GetProgress()
display(Javascript("$('div#%s').width('%i%%')" % (divid, int(p*100))))
if g_Abort:
processObject.Abort()
def command_js_start_abort():
g_Abort=False
# -
g_Abort=False
filter.RemoveAllCommands()
filter.AddCommand(sitk.sitkStartEvent, command_js_start_abort )
filter.AddCommand(sitk.sitkStartEvent, lambda: display(HTML(html_progress_abort+javascript_abort)))
filter.AddCommand(sitk.sitkProgressEvent, lambda: command_js_progress_abort(filter))
# A caveat with this approach is that the IPython kernel must continue to execute while the filter is running. So we must place the filter in a thread.
import threading
threading.Thread( target=lambda:filter.Execute() ).start()
# While the `lambda` command are convenient, the lack for having an object to hold data can still be problematic. For example in the above code the uuid, is used to uniquely identify the HTML element. So if the filter is executed multiple times then the JavaScript update will be confused on what to update.
#### The following shows a failure that you will want to avoid.
threading.Thread( target=lambda:filter.Execute() ).start()
# ### A Reusable class for IPython Progress
#
# There currently are too many caveats without support for Abort. Let us create a reusable class which will automatically generate the UUID and just display the progress.
# +
import uuid
from IPython.display import HTML, Javascript, display
class HTMLProgressWatcher:
def __init__(self, po):
self.processObject = po
self.abort = False
po.AddCommand(sitk.sitkStartEvent, lambda: self.cmdStartEvent() )
po.AddCommand(sitk.sitkProgressEvent, lambda: self.cmdProgressEvent() )
po.AddCommand(sitk.sitkEndEvent, lambda: self.cmdEndEvent() )
def cmdStartEvent(self):
global sitkIPythonProgress_UUID
self.abort=False
self.divid = str(uuid.uuid4())
try:
sitkIPythonProgress_UUID[self.divid] = self
except NameError:
sitkIPythonProgress_UUID = {self.divid: self}
html_progress_abort="""
<p style="margin:5px">{0}:</p>
<div style="border: 1px solid black;padding:1px;margin:5px">
<div id="{1}" style="background-color:blue; width:0%%"> </div>
</div>
""".format(self.processObject.GetName(), self.divid)
display(HTML(html_progress_abort+javascript_abort))
def cmdProgressEvent(self):
p = self.processObject.GetProgress()
display(Javascript("$('div#%s').width('%i%%')" % (self.divid, int(p*100))))
if self.abort:
self.processObject.Abort()
def cmdEndEvent(self):
global sitkIPythonProgress_UUID
del sitkIPythonProgress_UUID[self.divid]
del self.divid
# -
filter.RemoveAllCommands()
watcher = HTMLProgressWatcher(filter)
filter.Execute()
# +
# threading.Thread.start?
# -
| Python/41_Progress.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# Let's use the trap method, simpsons rule, and romberg integration
#
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
# # Change function
# +
#define a funtion
def func(x):
return np.exp( -2*x)*np.cos(10*x)
# -
# # define core of trap method
def func_integral(x):
return (np.exp(-2*x)*np.sin(10*x)*5)/52.0 - (np.exp(-2*x)*np.cos(10*x))/52.0
def trapezoid_core(f,x,h):
return 0.5*h*(f(x+h) + f(x))
# +
def trapezoid_method(f,a,b,N):
#f == tunction to integrate
#a == lower limit if integration
#b == upper limit of integration
#N == number of intervals to use
#define x valuesto perform the trapezoid rule
x = np.linspace(a,b,N)
h = x[1]-x[0]
# define the value of the integral
Fint = 0.0
#perform the integral using the trapezoid method
for i in range(0,len(x)-1,1):
Fint += trapezoid_core(f,x[i],h)
#return the answer
return Fint
# -
# ## define core of simpson;s method
def simpsons_core(f,x,h):
return h*(f(x) + 4*f(x+h) +f(x+2*h))/3.
# +
def simpsons_method(f,a,b,N):
#f == tunction to integrate
#a == lower limit if integration
#b == upper limit of integration
#N == number of intervals to use
#note the number of chunks will be N-1
#define x valuesto perform the trapezoid rule
x = np.linspace(a,b,N)
h = x[1]-x[0]
# define the value of the integral
Fint = 0.0
#perform the integral using the
#simpsons method
for i in range(0,len(x)-2,2):
Fint += simpsons_core(f,x[i],h)
#apply rule over the last interval if N is even
if((N%2)==2):
Fint += simpsons_core(f,x[-2],0.5*h)
#return the answer
return Fint
# -
# # define romberg core
def romberg_core(f,a,b,i):
#we need the difference between a and b
h = b-a
#interval between functions evaluations at refine level i
dh = h/2.**(i)
#we need the cofactor
K= h/2. **(i+1)
# and the function evaluations
M = 0.0
for j in range(2**i):
M += f(a + 0.5*dh + j*dh)
#return the answer
return K*M
# ## define a wrapper function to perform Romberg integration
def romberg_integration(f,a,b,tol):
#define an iteration variable
i=0
#define a maximum number of iterations
imax = 10000
# define an error estimate, set to a large value
delta = 100.0*np.fabs(tol)
#set an array of integral answers
I = np.zeros(imax,dtype=float)
#get the zeroth romberg iteration
I[0] = 0.5*(b-a)*(f(a) + f(b))
#iterate by 1
i += 1
while(delta>tol):
#find this romberg iteration
I[i] = 0.5*I[i-1] + romberg_core(f,a,b,i)
#compute the new functional error estimate
delta = np.fabs( (I[i] - I[i-1])/I[i] )
print(i,I[i],I[i-1],delta)
if(delta>tol):
#iterate
i += 1
#if we've reached the maximum iterations
if(i>imax):
print("Max iterations reached.")
raise StopIteration('Stopping iterations after' ,i)
#return the answer
return I[i]
# +
integral_value = func_integral(np.pi)-func_integral(0)
print("integral value using actual integration formulae for e^(-2x)cos(10x) = ", integral_value)
# Trapezoid method
#set initial interval size to 50
N = 50
tolerance = 1.0e-6
delta = 1 # some high number
while (delta > tolerance):
N = N + 50
num_int_value = trapezoid_method(func,0,np.pi,N);
delta = np.fabs(num_int_value - integral_value)
print()
print("numeric integral value by Trapezoid method", num_int_value)
print( "interval size = ",N,"delta = ", delta)
#Simpson method
N = 50
delta = 1 # some high number
while ((delta) > tolerance):
N = N + 50
num_int_value = simpsons_method(func,0,np.pi,N);
delta = np.fabs(num_int_value - integral_value)
print()
print("numeric integral value by simpson method", num_int_value)
print( "interval size = ",N,"delta = ", delta, )
print()
print("Integral value by Romberg method:")
tolerance = 1.0e-6
RI = romberg_integration(func,0,np.pi,tolerance)
print(RI, (RI-integral_value)/integral_value, tolerance)
# -
# ## Number of iterations for Romburg = 26
# ## Interval size for Trapezoid = 1300
# ## Interval size for Simpsons method = 5900
| astr-119-hw-5/HW-5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Building a multiclass classification model
# > Data cleaning, adding structures to PubChem data, building a multiclass model, dealing with imbalanced data
#
# - toc: True
# - branch: master
# - badges: true
# - comments: true
# - categories: [jupyter, multiclass, PubChem, imbalanced]
# - image: images/blog/multiclass.png
# - hide: false
# - search_exclude: false
# # Introduction
#
# At the Fall 2021 ACS Meeting, the group from [NCATS](https://opendata.ncats.nih.gov/adme/home) described a number of ADME models they had developed. Even better, the NCATS team also released some of the data used to build these models. In this notebook we'll use the data from the [NCATS CYP3A4](https://pubchem.ncbi.nlm.nih.gov/bioassay/1645841) assay to classify molecules as CYP3A4 activators, inhibitors, or inactive.
# In order to run this notebook, the following Python libraries should be installed
# - [pandas](https://pandas.pydata.org/) - handling data tables
# - [pubchempy](https://pubchempy.readthedocs.io/en/latest/) - grabbing chemical structures from PubChem
# - [tqdm](https://github.com/tqdm/tqdm) - progress bars
# - [numpy](https://numpy.org/) - linear algebra and matrices
# - [itertools](https://docs.python.org/3/library/itertools.html) - advanced list handling
# - [sklearn](https://scikit-learn.org/stable/) - machine learning
# - [lightgbm](https://lightgbm.readthedocs.io/en/latest/) - gradient boosted trees for machine learning
# - [matplotlib](https://matplotlib.org/) - plotting
# - [seaborn](https://seaborn.pydata.org/) - even better plotting
# - [pingouin](https://pingouin-stats.org/api.html) - stats
# - [imbalanced-learning](https://imbalanced-learn.org/stable/) - machine learning with imbalanced datasets
import pandas as pd
import pubchempy as pcp
from tqdm.auto import tqdm
import numpy as np
import itertools
from lib.descriptor_gen import DescriptorGen
from sklearn.preprocessing import LabelEncoder
from sklearn.model_selection import train_test_split
from lightgbm import LGBMClassifier
from sklearn.metrics import plot_confusion_matrix, matthews_corrcoef, roc_auc_score
import matplotlib.pyplot as plt
import seaborn as sns
# Enable Pandas progress_apply so that we can get progress bars for Pandas operations
tqdm.pandas()
# # Read and clean the PubChem data
df = pd.read_csv("data/AID_1645841_datatable.csv",skiprows=[1,2,3,4,5],low_memory=False)
df
# Note that the data file doesn't have chemical structures as SMILES strings. We're going to add those using the pubchempy library, which can look up the chemical structure based on the **PUBCHEM_CID** field in our dataframe. This is great, but the pubchempy service will have problems if we pass it a null value. Let's check and see if we have any null values in our **PUBCHEM_CID** column.
sum(df.PUBCHEM_CID.isna())
# We have four null values. Let's look at those rows.
df[df.PUBCHEM_CID.isna()]
# We can drop the four rows where **PUBCHEM_CID** is null.
df.dropna(subset=["PUBCHEM_CID"],inplace=True)
df
# In order to lookup a structure based on **PUBCHEM_CID** the **PUBCHEM_CID** field must be an integer. Let's take a look at the datatypes for our dataframe.
df.dtypes
# The **PUBCHEM_CID** field is a float64, which is not what we want. Let's convert that column to an integer.
df.PUBCHEM_CID = df.PUBCHEM_CID.astype(int)
# The field we want to model is **Phenotype-Replicate_1** which takes one of three values. Let's look at the possible values and their distribution. We can see that the class **Activator** is somewhat underrepresented. We'll start by building a model with the data as provided. Once we've done this, we'll also take a look at whether we can improve our model by employing strategies to compensate for the data imbalance.
df['Phenotype-Replicate_1'].value_counts(normalize=True)
# In order to build our model, we need the column were predicting to be represented as numeric values. We can convert the text labels to numbers using the [LabelEncoder](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.LabelEncoder.html) method from skikit-learn.
labels = df['Phenotype-Replicate_1'].unique().tolist()
le = LabelEncoder()
le.fit(labels)
df['label'] = le.transform(df['Phenotype-Replicate_1'])
# Our dataframe has a bunch of extra fields that are not necessary for this analysis. Let's simplify and create a new dataframe with only fields we care about.
data_df = df[['PUBCHEM_CID','Phenotype-Replicate_1','label']].copy()
data_df
# # Add chemical structures to the PubChem data
# Now we'll use pubchempy to look up chemical structures based on **PUBCHEM_CID** and add them to our dataframe. In order to keep the PubChem server happy, we'll break the **PUBCHEM_CID** list into chunks of 100.
cmpd_list = []
num_chunks = len(df)/100
for chunk in tqdm(np.array_split(data_df.PUBCHEM_CID,num_chunks)):
cmpd_list.append(pcp.get_compounds(chunk.tolist()))
# We collected the chemical structures in a list of lists. We need to flatten this into a single list. The operation works something like this
#
# <pre>[[1,2,3],[4,5,6],[7,8,9]] -> [1,2,3,4,5,6,7,8,9]</pre>
data_df['Compound'] = list(itertools.chain(*cmpd_list))
# Extract the **SMILES** from the **Compound** objects in the **Compound** column.
data_df['SMILES'] = [x.canonical_smiles for x in data_df.Compound]
# # Calculate molecular descriptors
# Create a **DescriptorGen** object for generating molecular descriptors.
desc_gen = DescriptorGen()
# Add the descriptors to the dataframe.
data_df['desc'] = data_df.SMILES.progress_apply(desc_gen.from_smiles)
# # Split the data into training and test sets
train, test = train_test_split(data_df)
train_X = np.stack(train.desc)
train_y = train.label
test_X = np.stack(test.desc)
test_y = test.label
# # Create and evaluate a machine learning model
# - Intstantiate a [LighGBM classifier](https://lightgbm.readthedocs.io/en/latest/pythonapi/lightgbm.LGBMClassifier.html#lightgbm.LGBMClassifier)
# - Train the model
# - Predict on the test set
lgbm = LGBMClassifier()
lgbm.fit(train_X, train_y)
pred = lgbm.predict_proba(test_X)
# Evaluate model performance using the ROC AUC score. Note that this is a little different with a multiclass classifer. We specify **class='ovo'** which means that we are evaluating "one vs one". We evaluate the AUC for all pairs of classes. The argument **average='macro'** indicates that the reported AUC is the average of all of the **one vs one** comparisons.
roc_auc_score(test_y,pred,multi_class='ovo',average='macro')
# We can also plot a confusion matrix to examine the model's performance on each of the three classes.
sns.set_style("white")
sns.set_context('talk')
plt.rcParams["figure.figsize"] = (8,8)
plot_confusion_matrix(lgbm,test_X,test_y,display_labels=sorted(labels),cmap=plt.cm.Blues)
# # Use oversampling to compensate for imbalanced data
# Load the imbalanced-learn library to perform oversampling. In a simple oversampling approach we repeated sample the imbalanced class(es) to create a balanced dataset.
from imblearn.over_sampling import RandomOverSampler
# We will create an oversampling object and use it to resample our training set.
ros = RandomOverSampler()
resample_X, resample_y = ros.fit_resample(train_X,train_y)
# Recall that our original training set is somewhat imbalanced. The minority class (0 or Activator) only accounts for ~7% of the data.
pd.Series(train_y).value_counts()
# After oversampling the dataset is balanced.
pd.Series(resample_y).value_counts()
# Build a model with the balanced, oversampled data
resample_lgbm = LGBMClassifier()
resample_lgbm.fit(resample_X, resample_y)
# Make a prediction with the new model, built with the resampled data.
resample_pred = resample_lgbm.predict_proba(test_X)
roc_auc_score(test_y,resample_pred,multi_class='ovr',average='macro')
# As above, we can plot a confusion matrix to examine the performance of the classifier trained on the oversampled data. Let's put the two confusion matrices side by side to compare.
sns.set_style("white")
sns.set_context('talk')
classifiers = [lgbm,resample_lgbm]
titles = ["Standard","Oversampled"]
fig, axes = plt.subplots(nrows=1, ncols=2, figsize=(15,5))
for cls,ax,title in zip(classifiers, axes, titles):
plot_confusion_matrix(cls,test_X,test_y,display_labels=sorted(labels),cmap=plt.cm.Blues,ax=ax)
ax.title.set_text(title)
plt.tight_layout()
# # Comparing the standard and oversampled models
# Let's see if there is a difference in AUC between the Standard and Resampled models. In order to compare we'll perform ten rounds of cross validation.
res = []
for i in tqdm(range(0,10)):
# split the data into training and test sets
train, test = train_test_split(data_df)
train_X = np.stack(train.desc)
train_y = train.label
test_X = np.stack(test.desc)
test_y = test.label
# Create the standard model
lgbm = LGBMClassifier()
lgbm.fit(train_X,train_y)
pred = lgbm.predict_proba(test_X)
auc = roc_auc_score(test_y,pred,multi_class='ovo',average='macro')
# Create the resampled model
resample_lgbm = LGBMClassifier()
resample_X, resample_y = ros.fit_resample(train_X,train_y)
resample_lgbm.fit(resample_X, resample_y)
resample_pred = resample_lgbm.predict_proba(test_X)
resample_auc = roc_auc_score(test_y,resample_pred,multi_class='ovr',average='macro')
res.append([auc, resample_auc])
# Create a dataframe with the AUC values using the Standard and Oversampled models
res = np.array(res)
res_df = pd.DataFrame(res,columns=["Standard","Oversampled"])
res_df.head()
# Reformat the dataframe to combine the two columns in **res_df**
melt_df = res_df.melt()
melt_df.columns = ["Method","AUC"]
melt_df
# Plot the AUC distributions for the Standard and Oversampled models as a kernel density estimate (KDE)
sns.set(rc={'figure.figsize': (10, 10)})
sns.set_context('talk')
sns.kdeplot(x="AUC",hue="Method",data=melt_df);
# Another way of comparing the distributions is to use the plot_paired method available in the pingouin library. Note that the AUC for the Resampled method is always greater than that for the Standard method.
from pingouin import wilcoxon, plot_paired
melt_df['cycle'] = list(range(0,10))+list(range(0,10))
plot_paired(data=melt_df,dv="AUC",within="Method",subject="cycle");
# In order to compare distributions, we sometime perform a t-test. However, a t-test assumes that the data is normally distributed. Since we can't make this assumption, we can use the Wilcoxon ranked sum test, which is the non-parametric equivalent to the t-test. The pingouin library provides a convenient implementation in the **wilcoxon** function. As we can see from the p-value in the table below, the difference in the means of the distributions is statistically significant.
wilcoxon(res_df.Standard,res_df.Oversampled)
| _notebooks/2021-08-28-multiclass-classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python36
# ---
# Copyright (c) Microsoft Corporation. All rights reserved.
#
# Licensed under the MIT License.
# # Training with the Azure Machine Learning Accelerated Models Service
# This notebook will introduce how to apply common machine learning techniques, like transfer learning, custom weights, and unquantized vs. quantized models, when working with our Azure Machine Learning Accelerated Models Service (Azure ML Accel Models).
#
# We will use Tensorflow for the preprocessing steps, ResNet50 for the featurizer, and the Keras API (built on Tensorflow backend) to build the classifier layers instead of the default ImageNet classifier used in Quickstart. Then we will train the model, evaluate it, and deploy it to run on an FPGA.
#
# #### Transfer Learning and Custom weights
# We will walk you through two ways to build and train a ResNet50 model on the Kaggle Cats and Dogs dataset: transfer learning only and then transfer learning with custom weights.
#
# In using transfer learning, our goal is to re-purpose the ResNet50 model already trained on the [ImageNet image dataset](http://www.image-net.org/) as a basis for our training of the Kaggle Cats and Dogs dataset. The ResNet50 featurizer will be imported as frozen, so only the Keras classifier will be trained.
#
# With the addition of custom weights, we will build the model so that the ResNet50 featurizer weights as not frozen. This will let us retrain starting with custom weights trained with ImageNet on ResNet50 and then use the Kaggle Cats and Dogs dataset to retrain and fine-tune the quantized version of the model.
#
# #### Unquantized vs. Quantized models
# The unquantized version of our models (ie. Resnet50, Resnet152, Densenet121, Vgg16, SsdVgg) uses native float precision (32-bit floats), which will be faster at training. We will use this for our first run through, then fine-tune the weights with the quantized version. The quantized version of our models (i.e. QuantizedResnet50, QuantizedResnet152, QuantizedDensenet121, QuantizedVgg16, QuantizedSsdVgg) will have the same node names as the unquantized version, but use quantized operations and will match the performance of the model when running on an FPGA.
#
# #### Contents
# 1. [Setup Environment](#setup)
# * [Prepare Data](#prepare-data)
# * [Construct Model](#construct-model)
# * Preprocessor
# * Classifier
# * Model construction
# * [Train Model](#train-model)
# * [Test Model](#test-model)
# * [Execution](#execution)
# * [Transfer Learning](#transfer-learning)
# * [Transfer Learning with Custom Weights](#custom-weights)
# * [Create Image](#create-image)
# * [Deploy Image](#deploy-image)
# * [Test the service](#test-service)
# * [Clean-up](#cleanup)
# * [Appendix](#appendix)
# <a id="setup"></a>
# ## 1. Setup Environment
# #### 1.a. Please set up your environment as described in the [Quickstart](./accelerated-models-quickstart.ipynb), meaning:
# * Make sure your Workspace config.json exists and has the correct info
# * Install Tensorflow
#
# #### 1.b. Download dataset into ~/catsanddogs
# The dataset we will be using for training can be downloaded [here](https://www.microsoft.com/en-us/download/details.aspx?id=54765). Download the zip and extract to a directory named 'catsanddogs' under your user directory ("~/catsanddogs").
#
#
# #### 1.c. Import packages
import os
import sys
import tensorflow as tf
import numpy as np
from keras import backend as K
import sklearn
import tqdm
# #### 1.d. Create directories for later use
# After you train your model in float32, you'll write the weights to a place on disk. We also need a location to store the models that get downloaded.
custom_weights_dir = os.path.expanduser("~/custom-weights")
saved_model_dir = os.path.expanduser("~/models")
# <a id="prepare-data"></a>
# ## 2. Prepare Data
# Load the files we are going to use for training and testing. By default this notebook uses only a very small subset of the Cats and Dogs dataset. That makes it run relatively quickly.
# +
import glob
import imghdr
datadir = os.path.expanduser("~/catsanddogs")
cat_files = glob.glob(os.path.join(datadir, 'PetImages', 'Cat', '*.jpg'))
dog_files = glob.glob(os.path.join(datadir, 'PetImages', 'Dog', '*.jpg'))
# Limit the data set to make the notebook execute quickly.
cat_files = cat_files[:64]
dog_files = dog_files[:64]
# The data set has a few images that are not jpeg. Remove them.
cat_files = [f for f in cat_files if imghdr.what(f) == 'jpeg']
dog_files = [f for f in dog_files if imghdr.what(f) == 'jpeg']
if(not len(cat_files) or not len(dog_files)):
print("Please download the Kaggle Cats and Dogs dataset form https://www.microsoft.com/en-us/download/details.aspx?id=54765 and extract the zip to " + datadir)
raise ValueError("Data not found")
else:
print(cat_files[0])
print(dog_files[0])
# -
# Construct a numpy array as labels
image_paths = cat_files + dog_files
total_files = len(cat_files) + len(dog_files)
labels = np.zeros(total_files)
labels[len(cat_files):] = 1
# +
# Split images data as training data and test data
from sklearn.model_selection import train_test_split
onehot_labels = np.array([[0,1] if i else [1,0] for i in labels])
img_train, img_test, label_train, label_test = train_test_split(image_paths, onehot_labels, random_state=42, shuffle=True)
print(len(img_train), len(img_test), label_train.shape, label_test.shape)
# -
# <a id="construct-model"></a>
# ## 3. Construct Model
# We will define the functions to handle creating the preprocessor and the classifier first, and then run them together to actually construct the model with the Resnet50 featurizer in a single Tensorflow session in a separate cell.
#
# We use ResNet50 for the featurizer and build our own classifier using Keras layers. We train the featurizer and the classifier as one model. We will provide parameters to determine whether we are using the quantized version and whether we are using custom weights in training or not.
# ### 3.a. Define image preprocessing step
# Same as in the Quickstart, before passing image dataset to the ResNet50 featurizer, we need to preprocess the input file to get it into the form expected by ResNet50. ResNet50 expects float tensors representing the images in BGR, channel last order. We've provided a default implementation of the preprocessing that you can use.
#
# **Note:** Expect to see TF deprecation warnings until we port our SDK over to use Tensorflow 2.0.
# +
import azureml.accel.models.utils as utils
def preprocess_images(scaling_factor=1.0):
# Convert images to 3D tensors [width,height,channel] - channels are in BGR order.
in_images = tf.placeholder(tf.string)
image_tensors = utils.preprocess_array(in_images, 'RGB', scaling_factor)
return in_images, image_tensors
# -
# ### 3.b. Define classifier
# We use Keras layer APIs to construct the classifier. Because we're using the tensorflow backend, we can train this classifier in one session with our Resnet50 model.
def construct_classifier(in_tensor, seed=None):
from keras.layers import Dropout, Dense, Flatten
from keras.initializers import glorot_uniform
K.set_session(tf.get_default_session())
FC_SIZE = 1024
NUM_CLASSES = 2
x = Dropout(0.2, input_shape=(1, 1, int(in_tensor.shape[3]),), seed=seed)(in_tensor)
x = Dense(FC_SIZE, activation='relu', input_dim=(1, 1, int(in_tensor.shape[3]),),
kernel_initializer=glorot_uniform(seed=seed), bias_initializer='zeros')(x)
x = Flatten()(x)
preds = Dense(NUM_CLASSES, activation='softmax', input_dim=FC_SIZE, name='classifier_output',
kernel_initializer=glorot_uniform(seed=seed), bias_initializer='zeros')(x)
return preds
# ### 3.c. Define model construction
# Now that the preprocessor and classifier for the model are defined, we can define how we want to construct the model.
#
# Constructing the model has these steps:
# 1. Get preprocessing steps
# * Get featurizer using the Azure ML Accel Models SDK:
# * import the graph definition
# * restore the weights of the model into a Tensorflow session
# * Get classifier
#
def construct_model(quantized, starting_weights_directory = None):
from azureml.accel.models import Resnet50, QuantizedResnet50
# Convert images to 3D tensors [width,height,channel]
in_images, image_tensors = preprocess_images(1.0)
# Construct featurizer using quantized or unquantized ResNet50 model
if not quantized:
featurizer = Resnet50(saved_model_dir)
else:
featurizer = QuantizedResnet50(saved_model_dir, custom_weights_directory = starting_weights_directory)
features = featurizer.import_graph_def(input_tensor=image_tensors)
# Construct classifier
preds = construct_classifier(features)
# Initialize weights
sess = tf.get_default_session()
tf.global_variables_initializer().run()
featurizer.restore_weights(sess)
return in_images, image_tensors, features, preds, featurizer
# <a id="train-model"></a>
# ## 4. Train Model
def read_files(files):
""" Read files to array"""
contents = []
for path in files:
with open(path, 'rb') as f:
contents.append(f.read())
return contents
def train_model(preds, in_images, img_train, label_train, is_retrain = False, train_epoch = 10, learning_rate=None):
""" training model """
from keras.objectives import binary_crossentropy
from tqdm import tqdm
learning_rate = learning_rate if learning_rate else 0.001 if is_retrain else 0.01
# Specify the loss function
in_labels = tf.placeholder(tf.float32, shape=(None, 2))
cross_entropy = tf.reduce_mean(binary_crossentropy(in_labels, preds))
optimizer = tf.train.GradientDescentOptimizer(learning_rate).minimize(cross_entropy)
def chunks(a, b, n):
"""Yield successive n-sized chunks from a and b."""
if (len(a) != len(b)):
print("a and b are not equal in chunks(a,b,n)")
raise ValueError("Parameter error")
for i in range(0, len(a), n):
yield a[i:i + n], b[i:i + n]
chunk_size = 16
chunk_num = len(label_train) / chunk_size
sess = tf.get_default_session()
for epoch in range(train_epoch):
avg_loss = 0
for img_chunk, label_chunk in tqdm(chunks(img_train, label_train, chunk_size)):
contents = read_files(img_chunk)
_, loss = sess.run([optimizer, cross_entropy],
feed_dict={in_images: contents,
in_labels: label_chunk,
K.learning_phase(): 1})
avg_loss += loss / chunk_num
print("Epoch:", (epoch + 1), "loss = ", "{:.3f}".format(avg_loss))
# Reach desired performance
if (avg_loss < 0.001):
break
# <a id="test-model"></a>
# <a id="test-model"></a>
# ## 5. Test Model
def test_model(preds, in_images, img_test, label_test):
"""Test the model"""
from keras.metrics import categorical_accuracy
in_labels = tf.placeholder(tf.float32, shape=(None, 2))
accuracy = tf.reduce_mean(categorical_accuracy(in_labels, preds))
contents = read_files(img_test)
accuracy = accuracy.eval(feed_dict={in_images: contents,
in_labels: label_test,
K.learning_phase(): 0})
return accuracy
# <a id="execution"></a>
# ## 6. Execute steps
# You can run through the Transfer Learning section, then skip to Create AccelContainerImage. By default, because the custom weights section takes much longer for training twice, it is not saved as executable cells. You can copy the code or change cell type to 'Code'.
#
# <a id="transfer-learning"></a>
# ### 6.a. Training using Transfer Learning
# +
# Launch the training
tf.reset_default_graph()
sess = tf.Session(graph=tf.get_default_graph())
with sess.as_default():
in_images, image_tensors, features, preds, featurizer = construct_model(quantized=True)
train_model(preds, in_images, img_train, label_train, is_retrain=False, train_epoch=10, learning_rate=0.01)
accuracy = test_model(preds, in_images, img_test, label_test)
print("Accuracy:", accuracy)
# -
# #### Save Model
# +
model_name = 'resnet50-catsanddogs-tl'
model_save_path = os.path.join(saved_model_dir, model_name)
tf.saved_model.simple_save(sess, model_save_path,
inputs={'images': in_images},
outputs={'output_alias': preds})
input_tensors = in_images.name
output_tensors = preds.name
print(input_tensors)
print(output_tensors)
# -
# <a id="custom-weights"></a>
# ### 6.b. Traning using Custom Weights
#
# Because the quantized graph defintion and the float32 graph defintion share the same node names in the graph definitions, we can initally train the weights in float32, and then reload them with the quantized operations (which take longer) to fine-tune the model.
#
# First we train the model with custom weights but without quantization. Training is done with native float precision (32-bit floats). We load the training data set and batch the training with 10 epochs. When the performance reaches desired level or starts decredation, we stop the training iteration and save the weights as tensorflow checkpoint files.
# #### Launch the training
# ```
# tf.reset_default_graph()
# sess = tf.Session(graph=tf.get_default_graph())
#
# with sess.as_default():
# in_images, image_tensors, features, preds, featurizer = construct_model(quantized=False)
# train_model(preds, in_images, img_train, label_train, is_retrain=False, train_epoch=10)
# accuracy = test_model(preds, in_images, img_test, label_test)
# print("Accuracy:", accuracy)
# featurizer.save_weights(custom_weights_dir + "/rn50", tf.get_default_session())
# ```
# #### Test Model
# After training, we evaluate the trained model's accuracy on test dataset with quantization. So that we know the model's performance if it is deployed on the FPGA.
# ```
# tf.reset_default_graph()
# sess = tf.Session(graph=tf.get_default_graph())
#
# with sess.as_default():
# print("Testing trained model with quantization")
# in_images, image_tensors, features, preds, quantized_featurizer = construct_model(quantized=True, starting_weights_directory=custom_weights_dir)
# accuracy = test_model(preds, in_images, img_test, label_test)
# print("Accuracy:", accuracy)
# ```
# #### Fine-Tune Model
# Sometimes, the model's accuracy can drop significantly after quantization. In those cases, we need to retrain the model enabled with quantization to get better model accuracy.
# ```
# if (accuracy < 0.93):
# with sess.as_default():
# print("Fine-tuning model with quantization")
# train_model(preds, in_images, img_train, label_train, is_retrain=True, train_epoch=10)
# accuracy = test_model(preds, in_images, img_test, label_test)
# print("Accuracy:", accuracy)
# ```
# #### Save Model
# ```
# model_name = 'resnet50-catsanddogs-cw'
# model_save_path = os.path.join(saved_model_dir, model_name)
#
# tf.saved_model.simple_save(sess, model_save_path,
# inputs={'images': in_images},
# outputs={'output_alias': preds})
#
# input_tensors = in_images.name
# output_tensors = preds.name
# ```
# <a id="create-image"></a>
# ## 7. Create AccelContainerImage
#
# Below we will execute all the same steps as in the [Quickstart](./accelerated-models-quickstart.ipynb#create-image) to package the model we have saved locally into an accelerated Docker image saved in our workspace. To complete all the steps, it may take a few minutes. For more details on each step, check out the [Quickstart section on model registration](./accelerated-models-quickstart.ipynb#register-model).
# +
from azureml.core import Workspace
from azureml.core.model import Model
from azureml.core.image import Image
from azureml.accel import AccelOnnxConverter
from azureml.accel import AccelContainerImage
# Retrieve workspace
ws = Workspace.from_config()
print("Successfully retrieved workspace:", ws.name, ws.resource_group, ws.location, ws.subscription_id, '\n')
# Register model
registered_model = Model.register(workspace = ws,
model_path = model_save_path,
model_name = model_name)
print("Successfully registered: ", registered_model.name, registered_model.description, registered_model.version, '\n', sep = '\t')
# Convert model
convert_request = AccelOnnxConverter.convert_tf_model(ws, registered_model, input_tensors, output_tensors)
# If it fails, you can run wait_for_completion again with show_output=True.
convert_request.wait_for_completion(show_output=False)
converted_model = convert_request.result
print("\nSuccessfully converted: ", converted_model.name, converted_model.url, converted_model.version,
converted_model.id, converted_model.created_time, '\n')
# Package into AccelContainerImage
image_config = AccelContainerImage.image_configuration()
# Image name must be lowercase
image_name = "{}-image".format(model_name)
image = Image.create(name = image_name,
models = [converted_model],
image_config = image_config,
workspace = ws)
image.wait_for_creation()
print("Created AccelContainerImage: {} {} {}\n".format(image.name, image.creation_state, image.image_location))
# -
# <a id="deploy-image"></a>
# ## 8. Deploy image
# Once you have an Azure ML Accelerated Image in your Workspace, you can deploy it to two destinations, to a Databox Edge machine or to an AKS cluster.
#
# ### 8.a. Deploy to Databox Edge Machine using IoT Hub
# See the sample [here](https://github.com/Azure-Samples/aml-real-time-ai/) for using the Azure IoT CLI extension for deploying your Docker image to your Databox Edge Machine.
#
# ### 8.b. Deploy to AKS Cluster
# #### Create AKS ComputeTarget
# +
from azureml.core.compute import AksCompute, ComputeTarget
# Uses the specific FPGA enabled VM (sku: Standard_PB6s)
# Standard_PB6s are available in: eastus, westus2, westeurope, southeastasia
prov_config = AksCompute.provisioning_configuration(vm_size = "Standard_PB6s",
agent_count = 1,
location = "eastus")
aks_name = 'aks-pb6-tl'
# Create the cluster
aks_target = ComputeTarget.create(workspace = ws,
name = aks_name,
provisioning_configuration = prov_config)
# -
# Provisioning an AKS cluster might take awhile (15 or so minutes), and we want to wait until it's successfully provisioned before we can deploy a service to it. If you interrupt this cell, provisioning of the cluster will continue. You can re-run it or check the status in your Workspace under Compute.
aks_target.wait_for_completion(show_output = True)
print(aks_target.provisioning_state)
print(aks_target.provisioning_errors)
# #### Deploy AccelContainerImage to AKS ComputeTarget
# +
from azureml.core.webservice import Webservice, AksWebservice
# Set the web service configuration (for creating a test service, we don't want autoscale enabled)
# Authentication is enabled by default, but for testing we specify False
aks_config = AksWebservice.deploy_configuration(autoscale_enabled=False,
num_replicas=1,
auth_enabled = False)
aks_service_name ='my-aks-service'
aks_service = Webservice.deploy_from_image(workspace = ws,
name = aks_service_name,
image = image,
deployment_config = aks_config,
deployment_target = aks_target)
aks_service.wait_for_deployment(show_output = True)
# -
# <a id="test-service"></a>
# ## 9. Test the service
#
# <a id="create-client"></a>
# ### 9.a. Create Client
# The image supports gRPC and the TensorFlow Serving "predict" API. We have a client that can call into the docker image to get predictions.
#
# **Note:** If you chose to use auth_enabled=True when creating your AksWebservice.deploy_configuration(), see documentation [here](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.core.webservice(class)?view=azure-ml-py#get-keys--) on how to retrieve your keys and use either key as an argument to PredictionClient(...,access_token=key).
# **WARNING:** If you are running on Azure Notebooks free compute, you will not be able to make outgoing calls to your service. Try locating your client on a different machine to consume it.
# +
# Using the grpc client in AzureML Accelerated Models SDK
from azureml.accel.client import PredictionClient
address = aks_service.scoring_uri
ssl_enabled = address.startswith("https")
address = address[address.find('/')+2:].strip('/')
port = 443 if ssl_enabled else 80
# Initialize AzureML Accelerated Models client
client = PredictionClient(address=address,
port=port,
use_ssl=ssl_enabled,
service_name=aks_service.name)
# -
# <a id="serve-model"></a>
# ### 9.b. Serve the model
# Let's see how our service does on a few images. It may get a few wrong.
# Specify an image to classify
print('CATS')
for image_file in cat_files[:8]:
results = client.score_file(path=image_file,
input_name=input_tensors,
outputs=output_tensors)
result = 'CORRECT ' if results[0] > results[1] else 'WRONG '
print(result + str(results))
print('DOGS')
for image_file in dog_files[:8]:
results = client.score_file(path=image_file,
input_name=input_tensors,
outputs=output_tensors)
result = 'CORRECT ' if results[1] > results[0] else 'WRONG '
print(result + str(results))
# <a id="cleanup"></a>
# ## 10. Cleanup
# It's important to clean up your resources, so that you won't incur unnecessary costs.
aks_service.delete()
aks_target.delete()
image.delete()
registered_model.delete()
converted_model.delete()
# <a id="appendix"></a>
# ## 11. Appendix
# License for plot_confusion_matrix:
#
# New BSD License
#
# Copyright (c) 2007-2018 The scikit-learn developers.
# All rights reserved.
#
#
# Redistribution and use in source and binary forms, with or without
# modification, are permitted provided that the following conditions are met:
#
# a. Redistributions of source code must retain the above copyright notice,
# this list of conditions and the following disclaimer.
# b. Redistributions in binary form must reproduce the above copyright
# notice, this list of conditions and the following disclaimer in the
# documentation and/or other materials provided with the distribution.
# c. Neither the name of the Scikit-learn Developers nor the names of
# its contributors may be used to endorse or promote products
# derived from this software without specific prior written
# permission.
#
#
# THIS SOFTWARE IS PROVIDED BY THE COPYRIGHT HOLDERS AND CONTRIBUTORS "AS IS"
# AND ANY EXPRESS OR IMPLIED WARRANTIES, INCLUDING, BUT NOT LIMITED TO, THE
# IMPLIED WARRANTIES OF MERCHANTABILITY AND FITNESS FOR A PARTICULAR PURPOSE
# ARE DISCLAIMED. IN NO EVENT SHALL THE REGENTS OR CONTRIBUTORS BE LIABLE FOR
# ANY DIRECT, INDIRECT, INCIDENTAL, SPECIAL, EXEMPLARY, OR CONSEQUENTIAL
# DAMAGES (INCLUDING, BUT NOT LIMITED TO, PROCUREMENT OF SUBSTITUTE GOODS OR
# SERVICES; LOSS OF USE, DATA, OR PROFITS; OR BUSINESS INTERRUPTION) HOWEVER
# CAUSED AND ON ANY THEORY OF LIABILITY, WHETHER IN CONTRACT, STRICT
# LIABILITY, OR TORT (INCLUDING NEGLIGENCE OR OTHERWISE) ARISING IN ANY WAY
# OUT OF THE USE OF THIS SOFTWARE, EVEN IF ADVISED OF THE POSSIBILITY OF SUCH
# DAMAGE.
#
| how-to-use-azureml/deployment/accelerated-models/accelerated-models-training.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# [idiomatic-python-coding-the-smart-way](https://medium.com/the-andela-way/idiomatic-python-coding-the-smart-way-cc560fa5f1d6)
# # 1. Chained comparison operators
x = 4
# Bad
if x >= 3 and x <= 5:
print("Hello")
# Good
if 3 <= x <= 5:
print("Hello")
# # 2. Indentation
# Bad
name = 'John'; address = 'Kampala'
if name: print(name)
print(address)
# Good
name = 'John'
address = 'Kampala'
if name:
print(name)
print(address)
# # 3. Use the Falsy & Truthy Concepts
# Bad
x = True
y = 0
if x == True:
print("Hello1")
elif x == False:
print("Hello2")
if y == 0:
print("Hello3")
ls = [2, 5]
if len(ls) > 0:
print("Hello4")
# Good
(x, y) = (True, 0)
# x is truthy
if x:
print("Hello1")
else:
print("Hello2")
# y is falsy
if not y:
print("Hello3")
ls = [2, 5]
if ls:
print("Hello4")
# +
# 4. Ternary Operator replacement
# -
# Bad
a = True
value = 0
if a:
value = 1
print(value)
# Good
a = True
value = 1 if a else 0
print(value)
# # 5. Use the 'in' keyword
# Bad
city = 'Nairobi'
found = False
if city == 'Nairobi' or city == 'Kampala' or city == 'Lagos':
found = True
# Good
city = 'Nairobi'
found = city in {'Nairobi', 'Kampala', 'Lagos'}
# Bad
cities = ['Nairobi', 'Kampala', 'Lagos']
index = 0
while index < len(cities):
print(cities[index])
index += 1
# Good
cities = ['Nairobi', 'Kampala', 'Lagos']
for city in cities:
print(city)
# # 6. Use 'return' to evaluate expressions, in addition to return values
# Bad
def check_equal(x, y):
result = False
if x == Y:
result = True
return result
# Good
def check_equal(x, y):
return x == y
# # 7. Multiple assignment
# Bad
x = 'foo'
y = 'foo'
z = 'foo'
# Good
x = y = z = 'foo'
# # 8. Formatting Strings
# Bad
def user_info(user):
return 'Name: ' + user.name + ' Age: '+ user.age
# Good
def user_info(user):
return 'Name: {user.name} Age: {user.age}'.format(user=user)
# # 9. List comprehension
# Bad
ls = list()
for element in range(10):
if not(element % 2):
ls.append(element)
# We may also employ a lambda function
ls = list(filter(lambda element: not(element % 2), range(10)))
# Good
ls = [element for element in range(10) if not(element % 2)]
# # 10. enumerate(list)
# Bad
ls = list(range(10))
index = 0
while index < len(ls):
print(ls[index], index)
index += 1
# Good
ls = list(range(10))
for index, value in enumerate(ls):
print(value, index)
# # 11. Dictionary Comprehension
# Bad
emails = {}
users = [{'name': 'a', 'email':'<EMAIL>'}, {'name': 'b', 'email': '<EMAIL>'}]
for user in users:
if user['email']:
emails[user['name']] = user['email']
print(emails)
# Good
users = [{'name': 'a', 'email':'<EMAIL>'}, {'name': 'b', 'email': '<EMAIL>'}]
emails = {user['name']: user['email'] for user in users if user['email']}
print(emails)
# # 12. Sets
# Bad
ls1 = [1, 2, 3, 4, 5]
ls2 = [4, 5, 6, 7, 8]
elements_in_both = []
for element in ls1:
if element in ls2:
elements_in_both.append(element)
print(elements_in_both)
# Good
ls1 = [1, 2, 3, 4, 5]
ls2 = [4, 5, 6, 7, 8]
elements_in_both = list( set(ls1) & set(ls2) )
print(elements_in_both)
# # 13. SetComprehension
# Bad
elements = [1, 3, 5, 2, 3, 7, 9, 2, 7]
unique_elements = set()
for element in elements:
unique_elements.add(element)
print(unique_elements)
# Good
elements = [1, 3, 5, 2, 3, 7, 9, 2, 7]
unique_elements = {element for element in elements}
print(unique_elements)
# # 14. Use the default parameter of ‘dict.get’ to provide default values
# Bad
auth = None
payload = {}
if 'auth_token' in payload:
auth = payload['auth_token']
else:
auth = 'Unauthorized'
print(auth)
# Good
payload = {}
auth = payload.get('auth_token', 'Unauthorized')
print(auth)
# # 15. Don’t Repeat Yourself (DRY)
# Bad
user = {'age': 10}
if user:
print('------------------------------')
print(user)
print('------------------------------')
# Good
user = {'age': 10}
if user:
print('{0}\n{1}\n{0}'.format('-'*30, user))
| idiomatic_python_coding_the_smart_way.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import networkx as nx
import pandas as pd
import itertools
import matplotlib.pyplot as plt
import seaborn as sns
import os
import linecache
import random
import sys
root_dir = "\\".join(sys.path[0].split("\\")[:-2])
KG = nx.read_gpickle(root_dir+"\\data\\graph\\training_KG_concepts.gpickle")
G = nx.convert_node_labels_to_integers(KG)
all_nodes = list(KG.nodes)
# 10000 pivots
# +
# betweenness_dict_10000 = nx.betweenness_centrality(KG, k=10000)
# np.save(root_dir+"\\data\\betweenness\\betweenness_dict_10000.npy", betweenness_dict_10000)
# -
betweenness_dict_10000 = np.load(
root_dir+"\\data\\betweenness\\betweenness_dict_10000.npy",
allow_pickle=True
).item()
edges_cc = pd.read_csv(root_dir+"\\data\\og\\edges_cc.csv")
all_edges = [
(edges_cc.src[i], edges_cc.dst[i]) \
for i \
in range(len(edges_cc))
]
training_edges = list(KG.edges())
validation_edges = list(set(all_edges)-set(training_edges))
training_betweenness = [
max(betweenness_dict_10000[edge[0]], betweenness_dict_10000[edge[1]]) \
for edge in training_edges
]
validation_betweenness = [
max(betweenness_dict_10000[edge[0]], betweenness_dict_10000[edge[1]]) \
for edge in validation_edges \
if edge[0] in betweenness_dict_1000 \
and edge[1] in betweenness_dict_1000
]
#
n = 1000
all_pairs = []
no_edge = []
validation = validation_betweenness
while len(all_pairs)<n or len(no_edge)<n or len(validation)<n:
pair = sample_node_pair(G)
all_pairs.append((all_nodes[pair[0]], all_nodes[pair[1]]))
if pair not in G.edges and pair[::-1] not in G.edges:
KG_pair = (all_nodes[pair[0]], all_nodes[pair[1]])
if KG_pair not in validation_edges and KG_pair[::-1] not in validation_edges:
no_edge.append(KG_pair)
# Betweenness Centrality Measurements
all_pairs_betweenness = []
for i in set(itertools.chain(*all_pairs)):
try:
all_pairs_betweenness.append(betweenness_dict_10000[i])
except:
continue
no_edge_betweenness = []
for i in set(itertools.chain(*no_edge)):
try:
no_edge_betweenness.append(betweenness_dict_10000[i])
except:
continue
training_betweenness = []
for i in set(itertools.chain(*training_edges)):
try:
training_betweenness.append(betweenness_dict_10000[i])
except:
continue
validation_betweenness = []
for i in set(itertools.chain(*validation_edges)):
try:
validation_betweenness.append(betweenness_dict_10000[i])
except:
continue
print(
np.mean(all_pairs_betweenness),
np.mean(no_edge_betweenness),
np.mean(training_betweenness),
np.mean(validation_betweenness)
)
print(
np.median(all_pairs_betweenness),
np.median(no_edge_betweenness),
np.median(training_betweenness),
np.median(validation_betweenness)
)
# +
df = pd.DataFrame(
data = {
"Normalized Betweenness (Unique Nodes)" : no_edge_betweenness+training_betweenness+validation_betweenness,
"" : ["No Edge"]*len(no_edge_betweenness)+["Training"]*len(training_betweenness)+["Validation"]*len(validation_betweenness)
}
)
#palette = {"Training": "#ABC9EA", "Validation": "#EFB792"}
palette = sns.color_palette("pastel")
g = sns.boxplot(x="", y="Normalized Betweenness (Unique Nodes)",
data=df, palette=palette)
g.set_yscale("log")
# -
all_pairs_betweenness = []
for i in all_pairs:
try:
b = max(betweenness_dict_10000[i[0]], betweenness_dict_10000[i[1]])
if b != b:
continue
all_pairs_betweenness.append(b)
except:
continue
no_edge_betweenness = []
for i in no_edge:
try:
b = max(betweenness_dict_10000[i[0]], betweenness_dict_10000[i[1]])
if b != b:
continue
no_edge_betweenness.append(b)
except:
continue
training_betweenness = []
for i in training_edges:
try:
b = max(betweenness_dict_10000[i[0]], betweenness_dict_10000[i[1]])
if b != b:
continue
training_betweenness.append(b)
except:
continue
validation_betweenness = []
for i in validation_edges:
try:
b = max(betweenness_dict_10000[i[0]], betweenness_dict_10000[i[1]])
if b != b:
continue
validation_betweenness.append(b)
except:
continue
print(
np.mean(all_pairs_betweenness),
np.mean(no_edge_betweenness),
np.mean(training_betweenness),
np.mean(validation_betweenness)
)
print(
np.median(all_pairs_betweenness),
np.median(no_edge_betweenness),
np.median(training_betweenness),
np.median(validation_betweenness)
)
# +
df = pd.DataFrame(
data = {
"Normalized Betweenness (Pairwise Max)" : no_edge_betweenness+training_betweenness+validation_betweenness,
"" : ["No Edge"]*len(no_edge_betweenness)+["Training"]*len(training_betweenness)+["Validation"]*len(validation_betweenness)
}
)
#palette = {"All Pairs": "black", "No Edge": "black", "Training": "#ABC9EA", "Validation": "#EFB792"}
palette = sns.color_palette("pastel")
g = sns.boxplot(x="", y="Normalized Betweenness (Pairwise Max)",
data=df, palette=palette)
g.set_yscale("log")
# -
| dev code/betweenness/Betweenness Centrality.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/aimancreator/phytonaiman/blob/main/03_0_Feature_Scaling.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + deletable=false editable=false id="0beRirb_nVr_"
# Copyright (c) 2019 Skymind AI Bhd.
# Copyright (c) 2020 <EMAIL>ifAI Sdn. Bhd.
#
# This program and the accompanying materials are made available under the
# terms of the Apache License, Version 2.0 which is available at
# https://www.apache.org/licenses/LICENSE-2.0.
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS, WITHOUT
# WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the
# License for the specific language governing permissions and limitations
# under the License.
#
# SPDX-License-Identifier: Apache-2.0
# + [markdown] id="3WzqzHY_nVsL"
# # Feature Scaling
# + [markdown] id="VA_kWZEmnVsM"
# While many algorithms (such as SVM, K-nearest neighbors, and logistic regression) require features to be normalized, intuitively we can think of **Principle Component Analysis (PCA)** as being a prime example of when normalization is important. In PCA we are interested in the components that maximize the variance. **If one component (e.g. human height) varies less than another (e.g. weight) because of their respective scales (meters vs. kilos)**, PCA might determine that the direction of maximal variance more closely corresponds with the ‘weight’ axis, if those features are **not scaled**. As a change in height of one meter can be considered much more important than the change in weight of one kilogram, this is clearly **incorrect**.
# + id="A7kOgrO1nVsN"
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn import preprocessing
from sklearn import datasets
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split
from sklearn import metrics
# %matplotlib inline
# + [markdown] id="SBwsa5j4nVsO"
# We will use a small dataset that contains (Physics,Biology and Maths) marks of a classroom of students.
# + id="_0oYFQHpnVsO"
df = pd.read_csv("https://archive.org/download/ml-fundamentals-data/machine-learning-fundamentals-data/grades.csv", index_col=0)
# + [markdown] id="qkahNBkenVsP"
# Show the first 5 rows of data.
# + id="rd8s0F8NnVsP" outputId="14bb5b1d-4f9a-43f1-a8f3-339c94b00c63" colab={"base_uri": "https://localhost:8080/", "height": 235}
df.head()
# + [markdown] id="_O-uunZQnVsR"
# <br>We can use boxplot to visualize the data
# + id="HdzM6Mw6nVsR" outputId="2911977a-5c2e-4ee0-fe81-9c2bd0ba188a" colab={"base_uri": "https://localhost:8080/", "height": 282}
df.boxplot()
# + [markdown] id="EisAOn-XnVsS"
# We could notice that the data is spread around the range of 1 to 100
# + [markdown] id="bZiaTHIenVsT"
# <br>We will use scaling functions from scikit learn to perform some preprocessing techniques to scale our data.
# <br>**Min-Max normalization** involves scaling features to lie between a given minimum and maximum value, often between zero and one.
# + id="ybm5po1bnVsU"
scaler = preprocessing.MinMaxScaler()
# + [markdown] id="DHLUu8YFnVsU"
# We will use Min-Max Scaling to scale all the columns of data.
# + id="gv9D3QhWnVsV"
data_scaled = scaler.fit_transform(df)
# + [markdown] id="7NlMe7GznVsV"
# Transform the numpy array containing our scaled data into a pandas data frame.
# + id="Kk0o4zhEnVsV" outputId="c529f044-bad6-4d72-9e5f-a6b4f467b23c" colab={"base_uri": "https://localhost:8080/"}
df_new = pd.DataFrame(data_scaled, index=df.index)
df.index
# + [markdown] id="aj3UwK3rnVsW"
# <br>Shows the first 5 rows of the scaled data
# + id="jJAcLIy6nVsZ" outputId="20e461e0-705e-41d5-d9b9-3b98fbc867b0"
df_new.head()
# + [markdown] id="iUYbBLhDnVsa"
# As you can see, our values are scaled into a *range from 0 to 1*.
# + id="cbKX52j1nVsa" outputId="4905079e-15a3-445e-9c24-2014570afcb8" colab={"base_uri": "https://localhost:8080/", "height": 282}
df_new.boxplot()
# + [markdown] id="zfgxwHKSnVsb"
# Another common scaling technique is the **Standard Scaler**.
# <br>Standard Scaler Standardize features by removing the mean and scaling to unit variance
#
# The standard score of a sample x is calculated as:
#
# $$z = \frac{x - u}{s}$$
#
# where $u$ is the mean of the training samples or zero if `with_mean = False`, and $s$ is the standard deviation of the training samples or one if `with_std = False`.
#
# **Standardization** of a dataset is a common requirement for many machine learning estimators: they might behave badly if the individual features do not more or less look like standard normally distributed data (e.g. Gaussian with 0 mean and unit variance).
# + [markdown] id="7zO-_hjOnVsb"
# <br>Now use the standard scaler to scale the data
# + id="30wb2rownVsc"
scaler = preprocessing.StandardScaler()
# + [markdown] id="2K4G931xnVsc"
# <br>Apply the scaler on the dataset and use df.head to visual the data
# + id="Ua4kPfbsnVsc" outputId="a7c77e42-ca0a-47d6-dd04-8d8eb09e757c" colab={"base_uri": "https://localhost:8080/", "height": 235}
data_scaled = scaler.fit_transform(df)
df_new = pd.DataFrame(data_scaled, index=df.index)
df_new.head()
# + [markdown] id="0iFkbb9SnVsd"
# <br>
# By plotting boxplot we can see that the range of the data changed where their average mean will be 0
# + id="B8R9va9DnVsd" outputId="0f3bf94b-2aff-4e04-d5f3-50f0724071c8" colab={"base_uri": "https://localhost:8080/", "height": 282}
df_new.boxplot()
# + [markdown] id="Dr2mo-PvnVsd"
# # Example of improved performance using Standard Scaling
# + [markdown] id="JE7iD3-enVse"
# In this example, we will use a social network advertisment dataset which the result will be whether the targeted person purchased the items in the advertisement
# + id="LTsWEMcDnVse"
df = pd.read_csv("https://storage.googleapis.com/kagglesdsdata/datasets/20757/26793/Social_Network_Ads.csv?X-Goog-Algorithm=GOOG4-RSA-SHA256&X-Goog-Credential=<PASSWORD>-kaggle-com%40k<PASSWORD>-161<PASSWORD>.iam.gserviceaccount.com%2F<PASSWORD>%2Fauto%2Fstorage%2Fgoog4_request&X-Goog-Date=20210409T032645Z&X-Goog-Expires=259199&X-Goog-SignedHeaders=host&X-Goog-Signature=31f455e07f1bf184f0e61cb7b25691a276321989e5600bab749db340aa1893f5d33da6d9308dd1a2fa29806bf79a5b4e2a010d972314895069898c053cc5607503484e7ed99f2e266546a77c29cce2fb87dc5702a8bd99473e2e0ac32cffeefe0b12ac7dfb1b734255dea35c2532709bb6dd2042fa9204492cbbf52822b15fa4e5c6b1cd89e6139722b85e35065c14eb74b84a2a4e57df46a1fea60f94e1f486c04a4f232a5f4705c32006c6e988fd0e3b6a19cd2771acbab07860aaceb451d7c46fa7f346b519ac2bc6740757d60218ee4a8e6b53782c360b9c297c3df03662fbde2baf5ef3d067d61058cd5d1338cc6515ba970869468561cb26af20f37b98")
# + [markdown] id="qyf1dkIpnVse"
# Visualizing the first 5 example of the dataset
# + id="eDGWEFCmnVse" outputId="07162ffc-e7e2-4aaf-ccb2-735cdef7fad8" colab={"base_uri": "https://localhost:8080/", "height": 204}
df.head()
# + [markdown] id="bZ9etSoPnVsf"
# Dropping column User ID which are not needed in the training
# + id="WbLG10YwnVsf" outputId="0ea53f6c-4a16-401f-a43a-d17b8d1756ea" colab={"base_uri": "https://localhost:8080/", "height": 204}
df.drop(columns=['Age'], axis=1, inplace=True)
df.head()
# + [markdown] id="zAuoFFAGnVsf"
# Use One Hot Encoding to split the Gender attribute into binary values
# + id="UdTVqvsanVsg" outputId="8cfc4b3f-b526-4382-dda5-3b3ce95c028d" colab={"base_uri": "https://localhost:8080/", "height": 323}
#df = pd.get_dummies(df, columns=['Gender'])
#df.head()
# + [markdown] id="iZr2OnNynVsg"
# Assigning the attributes that are needed for training
# + id="TQY5SL97nVsh" outputId="7b51d6fe-21d6-411f-9b82-dca923b501ad" colab={"base_uri": "https://localhost:8080/", "height": 561}
df.columns
y = df['Purchased']
df.drop(columns=['Purchased'], axis=1, inplace=True)
X = df
# + [markdown] id="wW_e9ye6nVsh"
# Splitting the data into training set and test set
# + id="IRKJJlIwnVsh"
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=123)
# + [markdown] id="kwdC3-EgnVsh"
# Training the model with Logistic Regression
# + id="7jZpGKBdnVsi" outputId="197afb91-d71f-41b9-b134-a31ed97f59dd" colab={"base_uri": "https://localhost:8080/"}
model = LogisticRegression()
model.fit(X_train, y_train)
# + [markdown] id="JpH5Mf-anVsi"
# Get the prediction using the test set
# + id="6dIlIA59nVsi"
predict = model.predict(X_test)
# + [markdown] id="4euigyY-nVsj"
# Calculating the accuracy score for the trained model
# + id="wFitY7P3nVsj" outputId="105ad8fc-6649-4dbc-b4d4-729d6cf3b7ba" colab={"base_uri": "https://localhost:8080/"}
accuracy = metrics.accuracy_score(predict, y_test)
print(accuracy)
# + [markdown] id="zTfxk9honVsj"
# As from here we can see the the performance of the model is not that good, if we use feature scaling, the performance will be better.
# + [markdown] id="6PCpiOaRnVsk"
# Scale the training set data with Standard Scaler
# <br> We will only fit the training set only as we learn the means and standard deviation of the training set, and then:
#
# - Standardize the training set using the training set means and standard deviations.
# - Standardize the test set using the training set means and standard deviations.
#
# Anything that we want to learn, must be learned from the model's training data, this is because we want to keep the test set data to be "unknown" by the model. If we scale the training set and test set together, we are actually using the knowledge of the whole data which result in "leaking information"
#
# Besides that, the test set data should be using the scaler from the training set data . If we scale the training set data and test set data separately, the value of 1.0 in training set data and 1.0 in test set data would actually be different as they learned from each respective mean and standard deviation. In the end it would greatly impact the model performance.
# + id="hGapL_TPnVsk"
scaler = preprocessing.StandardScaler()
X_train = scaler.fit_transform(X_train)
X_test = scaler.transform(X_test)
# + [markdown] id="YcNr4ce1nVsl"
# Now let us train the model again with the scaled data and look at the output of the accuracy
# + id="IWlX8UnhnVsl" outputId="db731499-6bf9-4080-ddbf-7f379b5b5407" colab={"base_uri": "https://localhost:8080/"}
model.fit(X_train, y_train)
predict = model.predict(X_test)
accuracy = metrics.accuracy_score(predict, y_test)
print(accuracy)
# + [markdown] id="XX8UHRCMnVsm"
# We can see that there is a great improvement in the accuracy after we scaled the data
| 03_0_Feature_Scaling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="VYNA79KmgvbY" colab_type="text"
# Copyright 2018 The Dopamine Authors.
#
# Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
# + [markdown] id="Ctd9k0h6wnqT" colab_type="text"
# # Visualize Dopamine baselines with Tensorboard
# This colab allows you to easily view the trained baselines with Tensorboard (even if you don't have Tensorboard on your local machine!).
#
# Simply specify the game you would like to visualize and then run the cells in order.
#
# _The instructions for setting up Tensorboard were obtained from https://www.dlology.com/blog/quick-guide-to-run-tensorboard-in-google-colab/_
# + id="s8r_45_0qpmb" colab_type="code" colab={} cellView="form"
# @title Prepare all necessary files and binaries.
# !wget https://bin.equinox.io/c/4VmDzA7iaHb/ngrok-stable-linux-amd64.zip
# !unzip ngrok-stable-linux-amd64.zip
# !gsutil -q -m cp -R gs://download-dopamine-rl/compiled_tb_event_files.tar.gz /content/
# !tar -xvzf /content/compiled_tb_event_files.tar.gz
# + id="D-oZRzeWwHZN" colab_type="code" colab={} cellView="form"
# @title Select which game to visualize.
game = 'Asterix' # @param['AirRaid', 'Alien', 'Amidar', 'Assault', 'Asterix', 'Asteroids', 'Atlantis', 'BankHeist', 'BattleZone', 'BeamRider', 'Berzerk', 'Bowling', 'Boxing', 'Breakout', 'Carnival', 'Centipede', 'ChopperCommand', 'CrazyClimber', 'DemonAttack', 'DoubleDunk', 'ElevatorAction', 'Enduro', 'FishingDerby', 'Freeway', 'Frostbite', 'Gopher', 'Gravitar', 'Hero', 'IceHockey', 'Jamesbond', 'JourneyEscape', 'Kangaroo', 'Krull', 'KungFuMaster', 'MontezumaRevenge', 'MsPacman', 'NameThisGame', 'Phoenix', 'Pitfall', 'Pong', 'Pooyan', 'PrivateEye', 'Qbert', 'Riverraid', 'RoadRunner', 'Robotank', 'Seaquest', 'Skiing', 'Solaris', 'SpaceInvaders', 'StarGunner', 'Tennis', 'TimePilot', 'Tutankham', 'UpNDown', 'Venture', 'VideoPinball', 'WizardOfWor', 'YarsRevenge', 'Zaxxon']
agents = ['dqn', 'c51', 'rainbow', 'iqn']
for agent in agents:
for run in range(1, 6):
# !mkdir -p "/content/$game/$agent/$run"
# !cp -r "/content/$agent/$game/$run" "/content/$game/$agent/$run"
LOG_DIR = '/content/{}'.format(game)
get_ipython().system_raw(
'tensorboard --logdir {} --host 0.0.0.0 --port 6006 &'
.format(LOG_DIR)
)
# + id="zlKKnaP4y9FA" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} cellView="form" outputId="3abff714-c484-436e-dc5f-88b15511f4f2"
# @title Start the tensorboard
get_ipython().system_raw('./ngrok http 6006 &')
# ! curl -s http://localhost:4040/api/tunnels | python3 -c \
# "import sys, json; print(json.load(sys.stdin)['tunnels'][0]['public_url'])"
| dopamine/colab/tensorboard.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="SC-JCbJfjcbg"
# # Reconstructed autoencoder
# + id="Q0aw5yLOy041" colab={"base_uri": "https://localhost:8080/", "height": 36} outputId="30568199-8558-4918-ef3e-39d55e838e93"
## load the libraries
import tensorflow as tf
from tensorflow import keras
from keras import Model, models, layers, optimizers, regularizers
from keras.layers import Dense, Input, LSTM, Dropout, Activation
from keras.models import Sequential
from tensorflow.keras.optimizers import Adam
from tensorflow.keras import layers
from tensorflow.keras.metrics import *
from tensorflow.python.client import device_lib
from tensorflow.keras.callbacks import ModelCheckpoint, EarlyStopping
from sklearn.preprocessing import LabelEncoder, MinMaxScaler, RobustScaler
from sklearn.model_selection import train_test_split
from sklearn.metrics import *
from sklearn import metrics
from pylab import rcParams
from collections import Counter
from itertools import repeat
from time import perf_counter
from imblearn.over_sampling import SMOTE
import matplotlib.pyplot as plt
import seaborn as sns
import pandas as pd
import numpy as np
import warnings
import os
warnings.filterwarnings('ignore')
warnings.filterwarnings("ignore", category=np.VisibleDeprecationWarning)
pd.set_option('display.max_columns', None)
tf.keras.backend.set_floatx('float32')
os.getcwd()
# + id="GYLK0-I3kK9h"
cols = {'obj': [],
'cat': [],
'cont': []
}
def data_split():
file_name = 'final_data_v2.csv'
file_path = os.getcwd()+'/drive/MyDrive/Colab Notebooks/'
df = pd.read_csv(file_path+file_name, encoding='utf-8')
df.rename(columns={'category_id_1':'category1'}, inplace=True)
# 데이터 유형별 분류하기
for dt_idx, dt_val in zip(df.dtypes.index, df.dtypes.values):
if dt_val == 'object':
if ('id' in dt_idx) | ('time' in dt_idx) | ('name' in dt_idx) | ('keyword' in dt_idx) |('url' in dt_idx):
df.drop(columns = dt_idx, axis=1, inplace=True)
else:
cols['obj'].append(dt_idx)
else:
if ('id' in dt_idx) | ('time' in dt_idx):
df.drop(columns = dt_idx, axis=1, inplace=True)
else:
if len(df[dt_idx].value_counts()) <= 30: #연속형 데이터 중 30개 내의 범주로 나눌 수 있는 데이터 = category로 구분.
cols['cat'].append(dt_idx)
else:
if ('hour' in dt_idx) | ('group' in dt_idx):
pass
else:
cols['cont'].append(dt_idx)
return cols
# + id="S4FL1ImtkK9h"
def reorganization(df):
data = pd.DataFrame()
cols = data_split()
for k, v in cols.items():
if k == 'obj':
data = pd.concat([data, df[v]], axis=1)
elif k == 'cont':
data = pd.concat([data, df[v]], axis=1)
else:
data = pd.concat([data, df[v]], axis=1)
return data
# + id="wYtsspxpkK9h"
def preprocessing():
file_name = 'final_data_v2.csv'
file_path = os.getcwd()+'/drive/MyDrive/Colab Notebooks/'
df = pd.read_csv(file_path+file_name, encoding='utf-8')
df.rename(columns={'category_id_1':'category1'}, inplace=True)
# 데이터 유형별 분류하기
data = reorganization(df)
modified_df = pd.DataFrame()
for i, c in enumerate(data.columns):
if c in cols['obj']:
obj_data = pd.get_dummies(data[c], prefix=c, prefix_sep = "/")
modified_df = pd.concat([modified_df, obj_data], axis=1)
elif c in cols['cat']: # click_label 컬럼 = y 변수로 사용
if 'click' in c:
pass
else:
cat_data = pd.get_dummies(data[c], prefix=c, prefix_sep = "/")
modified_df = pd.concat([modified_df, cat_data], axis=1)
else:
scaled_num_data = MinMaxScaler().fit_transform(df[[c]])
scaled_num_data = pd.DataFrame(scaled_num_data, columns = [c])
modified_df = pd.concat([modified_df,scaled_num_data], axis=1)
print('---- Data info ----')
print(cols)
print('Data Frame shape: {}'.format(modified_df.shape))
return modified_df
# + colab={"base_uri": "https://localhost:8080/"} outputId="553bd97b-9ad5-48b8-ca1d-52fa73ca1610" id="PkerczPKkK9h"
# 데이터 7:3으로 나누기 (검증 데이터는 8:2)
def split_data():
### read dataset
file_name = 'final_data_v2.csv'
file_path = os.getcwd()+'/drive/MyDrive/Colab Notebooks/'
df = pd.read_csv(file_path+file_name, encoding='utf-8')
df.rename(columns={'category_id_1':'category1'}, inplace=True)
modified_df = preprocessing()
X = modified_df
y = df['click_label']
# split the train/test data (7:3 ratio)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.3, random_state = 2022, stratify = y) #y 비율에 따른 층화추출 및 데이터를 7:3의 비율로 나누기
print(f"# of train_data's rows: {X_train.shape[0]} \n# of test_data's rows: {X_test.shape[0]}")
print(f'train:test ratio = {round(X_train.shape[0]/(X_train.shape[0]+ X_test.shape[0]),2)}:{round(X_test.shape[0]/(X_train.shape[0]+ X_test.shape[0]), 2)}')
oversample = SMOTE(random_state=2022) # 불균형 데이터 셋인 번개장터 데이터 셋 불균형 문제 완화
X_train, y_train = oversample.fit_resample(X_train, y_train)
## create train and validation datasets (8:2 ratio)
x_train, x_val, y_train, y_val = train_test_split(X_train, y_train, test_size=0.2, random_state = 2022, stratify = y_train)
print(f"# of train_data's rows: {x_train.shape[0]} \n# of validation_data's rows: {x_val.shape[0]}")
print(f'train:test ratio = {round(x_train.shape[0]/(x_train.shape[0]+ x_val.shape[0]),2)}:{round(x_val.shape[0]/(x_train.shape[0]+ x_val.shape[0]), 2)}')
## reshape the inputs
x_train = x_train.values.reshape(-1, x_train.shape[1])
x_val = x_val.values.reshape(-1, x_val.shape[1])
return x_train, X_test, x_val, y_train, y_test, y_val
x_train, X_test, x_val, y_train, y_test, y_val = split_data()
# + colab={"base_uri": "https://localhost:8080/"} outputId="8027f795-065a-403d-96b6-975f8f02497b" id="jfKPSwrIkK9i"
# Stacked autoencoder code
## input layer
input_layer = Input(shape=(x_train.shape[1],))
print(f"input layer's shape: {input_layer.shape}")
## encoding architecture
encode_layer = Dense(150, activation='relu', activity_regularizer = 'l2')(input_layer)
## latent view
latent_view = Dense(10, activation='sigmoid')(encode_layer)
## decoding architecture
decode_layer = Dense(150, activation='relu', activity_regularizer = 'l2')(latent_view)
## output layer
output_layer = Dense(x_train.shape[1],)(decode_layer)
print(f"output layer's shape: {output_layer.shape}")
# layer_loss = tf.losses.binary_crossentropy(input_layer, output_layer)
# print(f"binary_crossentropy of Autoencoder model: {layer_loss}")
model = tf.keras.Model(input_layer, output_layer)
print(model.summary())
# + colab={"base_uri": "https://localhost:8080/"} outputId="31ee6c20-cbd9-4933-ff79-ae791048182b" id="k4bOrliykK9i"
# patience: patience 는 성능이 증가하지 않는 epoch 을 몇 번이나 허용할 것인가를 정의
learning_rate = 0.01
optimizer = keras.optimizers.Adam(learning_rate=learning_rate)
model.compile(optimizer=optimizer, loss = 'binary_crossentropy', metrics = ['binary_accuracy'])
early_stopping = EarlyStopping(monitor='val_loss', patience=5, verbose=1, mode='auto')
start = perf_counter()
history = model.fit(x_train, y_train, epochs=200, batch_size=512, validation_data=(x_val, y_val), callbacks=[early_stopping])
print("End of Training")
print("걸린 시간: {:g}분 {:.2f}초".format((perf_counter() - start)//60, round((perf_counter() - start)%60)))
# + colab={"base_uri": "https://localhost:8080/", "height": 520} id="N2WjzAvSwUio" outputId="644728ea-bd55-4082-ccc0-bb87233f4098"
plt.figure(figsize=(10, 8))
sns.lineplot(data= pd.DataFrame(history.history)[['loss', 'val_loss']])
plt.grid(True)
plt.xlabel('Epoch', fontsize=15)
plt.title('Training with validation data about loss value (Batch size = 512)', fontsize=15)
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 520} id="mGSjREK212qu" outputId="9499053e-80ab-42d2-da0d-5b5e95484b25"
plt.figure(figsize=(10, 8))
sns.lineplot(data= pd.DataFrame(history.history)[['binary_accuracy', 'val_binary_accuracy']])
plt.grid(True)
plt.xlabel('Epoch', fontsize=15)
plt.title('Training with validation data about loss value (Batch size = 512)', fontsize=15)
plt.show()
# + id="mY20x_Liw6VN"
def flatten(X):
flattened_X = np.empty((X.shape[0], X.shape[1])) # sample x features array.
for i in range(X.shape[0]):
flattened_X[i] = X[i, :]
return flattened_X
# + colab={"base_uri": "https://localhost:8080/", "height": 350} id="3MIwnzG8w1xJ" outputId="c9ea724b-8d94-451e-8f08-b76753c97b64"
# %matplotlib inline
val_preds = model.predict(x_val)
bin_acc = tf.metrics.binary_accuracy(flatten(x_val), flatten(val_preds))
result_df = pd.DataFrame({'Reconstruction_error':bin_acc,
'True_class':list(y_val)})
precision_rt, recall_rt, threshold_rt = metrics.precision_recall_curve(result_df['True_class'], result_df['Reconstruction_error'])
plt.figure(figsize=(8,5))
plt.plot(threshold_rt, precision_rt[1:], label='Precision')
plt.plot(threshold_rt, recall_rt[1:], label='Recall')
plt.xlabel('Threshold'); plt.ylabel('Precision/Recall/ROC-AUC value')
plt.title('Precision/Recall Curve & ROC-AUC score per threshold about Validation data')
auc = []
for thr in threshold_rt:
pred_y = [1 if e >= thr else 0 for e in result_df.Reconstruction_error.values]
result_df['pred'] = pred_y
auc.append(roc_auc_score(result_df.True_class, result_df['pred']))
plt.plot(threshold_rt, auc,label='roc-auc')
plt.legend(); plt.grid()
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="dT0GfPse-7Mt" outputId="de650068-86e7-4748-c70e-10709cab8757"
# best position of threshold
index_cnt = [cnt for cnt, (p, r) in enumerate(zip(precision_rt, recall_rt)) if p==r][0]
print('precision: ',precision_rt[index_cnt],', recall: ',recall_rt[index_cnt], end = ', ')
# fixed Threshold
threshold_fixed = threshold_rt[index_cnt]
print('threshold: ',threshold_fixed)
# + colab={"base_uri": "https://localhost:8080/", "height": 550} id="hCs6Ny_19qBk" outputId="8dba5c97-a598-42a7-cb21-d33f56e6d07c"
# for i in threshold_rt:
pred_y_val = [1 if e > threshold_fixed else 0 for e in result_df.Reconstruction_error.values]
cm = confusion_matrix(result_df.True_class, pred_y_val)
print(f'threshold: {threshold_fixed}')
print(cm)
print(classification_report(result_df.True_class, pred_y_val))
print("f1 score: ",f1_score(result_df['True_class'], pred_y_val))
print('ROC-AUC:', roc_auc_score(result_df.True_class, pred_y_val))
# confusion matrix heatmap
LABELS = ['No', 'Yes']
plt.figure(figsize=(4, 4))
sns.heatmap(cm, xticklabels=LABELS, yticklabels=LABELS, annot=True, fmt="g", cmap='Pastel1'); plt.title("Confusion matrix")
plt.ylabel('True class')
plt.xlabel('Predicted class')
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} outputId="b2e2c91b-796c-41a6-c525-daa1c1ffe6cb" id="xYl5PqNiNVVo"
loss, accuracy = model.evaluate(x_val, y_val)
print("테스트 데이터 정확도", accuracy)
# + colab={"base_uri": "https://localhost:8080/"} outputId="fc3e9fbc-6138-48d0-a3e2-a3f0b1743475" id="Kw9o2VJyNVVp"
print("Validation data - Accuracy value: ",accuracy_score(result_df['True_class'], pred_y_val))
# + colab={"base_uri": "https://localhost:8080/", "height": 350} id="CnsM1NbwBUas" outputId="50d8393e-069c-4967-de9b-5dbbc450b48d"
# %matplotlib inline
preds = model.predict(X_test)
bin_acc = tf.metrics.binary_accuracy(flatten(X_test.values.reshape(-1, x_val.shape[1])), flatten(preds))
result_df = pd.DataFrame({'Reconstruction_error': bin_acc,
'True_class': y_test.tolist()})
precision_rt, recall_rt, threshold_rt = metrics.precision_recall_curve(result_df['True_class'], result_df['Reconstruction_error'])
plt.figure(figsize=(8,5))
plt.plot(threshold_rt, precision_rt[1:], label='Precision')
plt.plot(threshold_rt, recall_rt[1:], label='Recall')
plt.xlabel('Threshold'); plt.ylabel('Precision/Recall/ROC-AUC value')
plt.title('Precision/Recall Curve & ROC-AUC score per threshold about Test data')
auc= []
for thr in threshold_rt:
pred_y = [1 if e >= thr else 0 for e in result_df.Reconstruction_error.values]
result_df['pred'] = pred_y
auc.append(roc_auc_score(result_df.True_class, result_df['pred']))
plt.plot(threshold_rt, auc,label='roc-auc')
plt.legend(); plt.grid()
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="HYCcRDP_AqgX" outputId="8a6214ad-9b14-4a59-c55a-5b688cb0cc9e"
# best position of threshold
# index_cnt = [cnt for cnt, (p, r) in enumerate(zip(precision_rt, recall_rt)) if p==r]
print('precision: ',precision_rt[auc.index(np.max(auc))],', recall: ',recall_rt[auc.index(np.max(auc))], end = ', ')
# fixed Threshold
threshold_fixed = threshold_rt[auc.index(np.max(auc))]#threshold_rt[index_cnt]
print('threshold: ',threshold_fixed)
# + colab={"base_uri": "https://localhost:8080/"} id="_PXMtahPCFzj" outputId="84454977-d0e9-468c-9946-73122dae44bd"
loss, accuracy = model.evaluate(X_test, y_test)
print("테스트 데이터 정확도", accuracy)
# + colab={"base_uri": "https://localhost:8080/"} id="SF2XnVtFLyap" outputId="e815d50d-412b-4b3f-f8b3-7d9ff7dd7322"
pred_y = [1 if e >= threshold_fixed else 0 for e in result_df.Reconstruction_error.values]
result_df['pred'] = pred_y
print("Test data - Accuracy value: ",accuracy_score(result_df['True_class'], result_df['pred']))
# + colab={"base_uri": "https://localhost:8080/", "height": 550} id="oM8yRc6OB8qu" outputId="3672fb00-30ec-4414-ea9b-8aa56caaa1e9"
# for i in threshold_rt:
pred_y_test = [1 if e > threshold_fixed else 0 for e in result_df.Reconstruction_error.values]
cm = confusion_matrix(result_df.True_class, pred_y_test)
print(f'threshold: {threshold_fixed}')
print(cm)
print(classification_report(result_df.True_class, pred_y_test))
print("f1 score: ",f1_score(result_df['True_class'], pred_y_test))
print('ROC-AUC:', roc_auc_score(result_df.True_class, pred_y_test))
# confusion matrix heatmap
LABELS = ['No', 'Yes']
plt.figure(figsize=(4, 4))
sns.heatmap(cm, xticklabels=LABELS, yticklabels=LABELS, annot=True, fmt="g", cmap='Pastel1'); plt.title("Confusion matrix")
plt.ylabel('True class')
plt.xlabel('Predicted class')
plt.show()
| revised_autoencoder.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from matplotlib import rcParams
rcParams['legend.fontsize'] = 14
rcParams['axes.labelsize'] = 14
rcParams['axes.titlesize'] = 14
# %matplotlib inline
# -
# # INS-GPS Integration
# This notebook shows an idealized example of loose INS-GPS integration.
# # Creating a trajectory and generating inertial readings
# First we need to generate a trajectory. To keep things simple we generate sort of a random walk trajectory by summing random displacements.
from pyins import sim
from pyins.coord import perturb_ll
def generate_trajectory(n_points, min_step, max_step, angle_spread, random_state=0):
rng = np.random.RandomState(random_state)
xy = [np.zeros(2)]
angle = rng.uniform(2 * np.pi)
heading = [90 - angle]
angle_spread = np.deg2rad(angle_spread)
for i in range(n_points - 1):
step = rng.uniform(min_step, max_step)
xy.append(xy[-1] + step * np.array([np.cos(angle), np.sin(angle)]))
angle += rng.uniform(-angle_spread, angle_spread)
heading.append(90 - angle)
return np.asarray(xy), np.asarray(heading)
xy, h = generate_trajectory(1000, 70, 100, 20, random_state=1)
# Assume that each step is done in 10 seconds and define time stamps:
t = np.arange(1000) * 10
# Add displacements to initial latitude and longitude:
lat0 = 58
lon0 = 56
lat, lon = perturb_ll(lat0, lon0, xy[:, 1], xy[:, 0])
# We also append 20 minutes of INS being at rest:
t = np.hstack((-1200, t))
lat = np.hstack((lat[0], lat))
lon = np.hstack((lon[0], lon))
h = np.hstack((h[0], h))
# Set pitch and roll angles to zeros:
p = np.zeros_like(h)
r = np.zeros_like(h)
# Sensor sampling period is set to 0.1:
dt = 0.1
# Run the simulation routine which will interpolate the trajectory and generate inertial readings:
traj_ref, gyro, accel = sim.from_position(dt, lat, lon, t, h=h, p=p, r=r)
# The final trajectory is drawn below, the initial point is marked with a cross.
plt.plot(traj_ref.lon, traj_ref.lat)
plt.plot(traj_ref.lon[0], traj_ref.lat[0], 'kx', markersize=12)
plt.xlabel("lon, deg")
plt.ylabel("lat, deg")
# ## Integrating ideal data
# Just to check that everything is correct we want to integrate the generated gyro and accel readings.
from pyins.integrate import coning_sculling, integrate
from pyins.filt import traj_diff
# First we apply coning and sculling corrections:
theta, dv = coning_sculling(gyro, accel)
# And the run the integration.
traj_ideal = integrate(dt, *traj_ref.iloc[0], theta, dv)
# Compute integration error using a convenience function:
err_ideal = traj_diff(traj_ideal, traj_ref)
def plot_errors(dt, err, step=1000):
plt.figure(figsize=(15, 10))
plt.subplot(331)
err = err.iloc[::step]
t = err.index * dt / 3600
plt.plot(t, err.lat, label='lat')
plt.xlabel("time, h")
plt.ylabel("m")
plt.legend(loc='best')
plt.subplot(334)
plt.plot(t, err.lon, label='lon')
plt.xlabel("time, h")
plt.ylabel("m")
plt.legend(loc='best')
plt.subplot(332)
plt.plot(t, err.VE, label='VE')
plt.xlabel("time, h")
plt.ylabel("m/s")
plt.legend(loc='best')
plt.subplot(335)
plt.plot(t, err.VN, label='VN')
plt.xlabel("time, h")
plt.ylabel("m/s")
plt.legend(loc='best')
plt.subplot(333)
plt.plot(t, err.h, label='heading')
plt.xlabel("time, h")
plt.ylabel("deg")
plt.legend(loc='best')
plt.subplot(336)
plt.plot(t, err.p, label='pitch')
plt.xlabel("time, h")
plt.ylabel("deg")
plt.legend(loc='best')
plt.subplot(339)
plt.plot(t, err.r, label='roll')
plt.xlabel("time, h")
plt.ylabel("deg")
plt.legend(loc='best')
plt.tight_layout()
# We see that attitude and velocity errors are vanishingly small. The position errors are less than 3 meters during 3 hours of operations, which is completely negligible compared to errors of even the most accurate INS.
plot_errors(dt, err_ideal)
# ## Integrating "real" data
# Now we will run the navigation using inertial sensors with errors.
#
# The error will be a sum of a random bias and additive white noise. We define magnitudes typical for moderately accurate navigation grade sensors.
gyro_bias_sd = np.deg2rad(0.05) / 3600 # 0.05 d/h
accel_bias_sd = 5e-3
gyro_bias_sd
gyro_noise = 1e-6 # rad / s^0.5
accel_noise = 3e-4 # m / s^1.5
# Compute biases as a random constants. To avoid a "bad case" in this example we generated biases uniformly within $[-2 \sigma, 2 \sigma]$.
np.random.seed(1)
gyro_bias = gyro_bias_sd * np.random.uniform(-2, 2, 3)
accel_bias = accel_bias_sd * np.random.uniform(-2, 2, 3)
gyro_bias, accel_bias
from pyins import earth
# Now we apply errors to inertial readings:
gyro_e = gyro + gyro_bias * dt + gyro_noise * np.random.randn(*gyro.shape) * dt**0.5
accel_e = accel + accel_bias * dt + accel_noise * np.random.randn(*accel.shape) * dt**0.5
# Compute coning and sculling corrections:
theta, dv = coning_sculling(gyro_e, accel_e)
# An INS operation have to start with the self alignment. We devote 15 minutes of the initial rest for it:
t_align = 15 * 60
align_samples = int(t_align / dt)
# Split the readings into alignment and navigation parts:
theta_align = theta[:align_samples]
theta_nav = theta[align_samples:]
dv_align = dv[:align_samples]
dv_nav = dv[align_samples:]
from pyins.align import align_wahba
(h0, p0, r0), P_align = align_wahba(dt, theta_align, dv_align, 58)
# Compare estimated attitude angles with the true angles.
h0 - traj_ref.h.loc[align_samples], p0 - traj_ref.p.loc[align_samples], r0 - traj_ref.r.loc[align_samples]
# Assume that the initial position is known with the accuracy typical to GPS receivers:
lat0, lon0 = perturb_ll(traj_ref.lat.loc[align_samples], traj_ref.lon.loc[align_samples],
10 * np.random.randn(1), 10 * np.random.randn(1))
# Assume that it is known that the navigation starts at rest ans set initial velocities to 0:
VE0 = 0
VN0 = 0
# Now we can run the integration:
traj_real = integrate(dt, lat0, lon0, VE0, VN0, h0, p0, r0, theta_nav, dv_nav, stamp=align_samples)
traj_error = traj_diff(traj_real, traj_ref)
# We see that even with very accurate gyros pure INS performance is not that good.
plot_errors(dt, traj_error)
# ## Aiding from GPS
# Now we will use idealize GPS position observations for INS errors estimation and correction.
# We assume that GPS is available every second and use known exact values of latitude and longitude:
gps_data = pd.DataFrame(index=traj_ref.index[::10])
gps_data['lat'] = traj_ref.lat[::10]
gps_data['lon'] = traj_ref.lon[::10]
# We will use an idealized model that GPS observations contain only additive normal errors with a standard deviation of 10 meters (note that in reality errors in outputs from GPS receivers behave much worse).
gps_pos_sd = 10
gps_data['lat'], gps_data['lon'] = perturb_ll(gps_data.lat, gps_data.lon,
gps_pos_sd * np.random.randn(*gps_data.lat.shape),
gps_pos_sd * np.random.randn(*gps_data.lon.shape))
# To use GPS measurements in a navigation Kalman filter we wrap this data into a special object:
from pyins.filt import LatLonObs
gps_obs = LatLonObs(gps_data, gps_pos_sd)
# Also define gyro and accelerometer models using parameters defined above:
from pyins.filt import InertialSensor
gyro_model = InertialSensor(bias=gyro_bias_sd, noise=gyro_noise)
accel_model = InertialSensor(bias=accel_bias_sd, noise=accel_noise)
# Now we can run a navigation Kalman filter which will blend INS and GPS data. In this example INS errors didn't grow very large, thus we can use a feedforward filter.
from pyins.filt import FeedforwardFilter
# We create a filter by passing sampling period and computed trajectory. To initialize the covariance matrix we pass standard deviations of the initial errors.
#
# Currently the covariance matrix is initialized as diagonal, even though it can be done more rigorously, i.e consider correlations between sensor biases and attitude errors. But my view is that a reliable filter should not depend on such fine details, otherwise it is likely to fail in real conditions. So for the sake of simplicity it is implemented like this for now (can be changed later).
# Theoretical attitude accuracy (sd values) from static gyrocompassing in our case is: 0.35 deg for heading (`azimuth_sd`) and 0.03 deg for pitch and roll (`level_sd`). Here we set values slightly higher to account for a non-perfect alignment:
ff_filt = FeedforwardFilter(dt, traj_real,
pos_sd=10, vel_sd=0.1, azimuth_sd=0.5, level_sd=0.05,
gyro_model=gyro_model, accel_model=accel_model)
# We run the filter and pass available measurements to it. The return value is the INS trajectory corrected by estimated errors.
ff_res = ff_filt.run(observations=[gps_obs])
# Now we want to investigate errors in the filtered trajectory.
filt_error = traj_diff(ff_res.traj, traj_ref)
# Obviously performance in terms of position and velocity accuracy is very good, but this is sort of expected because GPS provides coordinates directly.
#
# Attitude angle errors are generally decreasing as well, but the picture is less clear. We want to plot their standard deviation bounds estimated by the filter as well.
plot_errors(dt, filt_error, step=10)
# The return value of `FeedforwardFilter` contains attributes `err`, `sd`, `gyro_err`, `gyro_sd`, `accel_err`, `accel_sd` for estimated trajectory errors and inertial sensor states and their standard deviations. Below we plot true errors for heading, pitch and roll with their 1-sigma bounds provided by the filter.
# Generally we see that the filter's performance is adequate. It can be measured more precisely by Monte-Carlo simulation, but this will not be included in this example.
# +
plt.figure(figsize=(15, 5))
t_plot = filt_error.index * dt / 3600
plt.subplot(131)
plt.plot(t_plot, filt_error.h, 'b')
plt.plot(t_plot, ff_res.sd.h, 'b--')
plt.plot(t_plot, -ff_res.sd.h, 'b--')
plt.xlabel("time, h")
plt.ylabel("deg")
plt.title("heading error")
plt.subplot(132)
plt.plot(t_plot, filt_error.p, 'b')
plt.plot(t_plot, ff_res.sd.p, 'b--')
plt.plot(t_plot, -ff_res.sd.p, 'b--')
plt.xlabel("time, h")
plt.ylabel("deg")
plt.title("pitch error")
plt.subplot(133)
plt.plot(t_plot, filt_error.r, 'b')
plt.plot(t_plot, ff_res.sd.r, 'b--')
plt.plot(t_plot, -ff_res.sd.r, 'b--')
plt.xlabel("time, h")
plt.ylabel("deg")
plt.title("roll error")
plt.tight_layout()
# -
# Also it is interesting to assess the filter's sensor bias estimation. Plots below show $\pm \sigma$ bands of gyro bias estimates, the straight line depicts the true value. We see that estimation of gyro biases is quite successful.
# +
plt.figure(figsize=(15, 5))
t_plot = filt_error.index[::10] * dt / 3600
gyro_err = ff_res.gyro_err.iloc[::10]
gyro_sd = ff_res.gyro_sd.iloc[::10]
plt.subplot(131)
plt.plot(t_plot, gyro_err.BIAS_1 + gyro_sd.BIAS_1, 'b')
plt.plot(t_plot, gyro_err.BIAS_1 - gyro_sd.BIAS_1, 'b')
plt.hlines(gyro_bias[0], *plt.xlim())
plt.xlabel("time, h")
plt.ylabel("rad/s")
plt.title("Gyro 1 bias")
plt.subplot(132)
plt.plot(t_plot, gyro_err.BIAS_2 + gyro_sd.BIAS_2, 'b')
plt.plot(t_plot, gyro_err.BIAS_2 - gyro_sd.BIAS_2, 'b')
plt.hlines(gyro_bias[1], *plt.xlim())
plt.xlabel("time, h")
plt.ylabel("rad/s")
plt.title("Gyro 2 bias")
plt.subplot(133)
plt.plot(t_plot, gyro_err.BIAS_3 + gyro_sd.BIAS_3, 'b')
plt.plot(t_plot, gyro_err.BIAS_3 - gyro_sd.BIAS_3, 'b')
plt.hlines(gyro_bias[2], *plt.xlim())
plt.xlabel("time, h")
plt.ylabel("rad/s")
plt.title("Gyro 3 bias")
plt.tight_layout()
# -
# Below the same done for accelerometer biases. Horizontal accelerometer biases are less observable on the given trajectory than gyro biases, and the vertical bias is not observable at all because pitch and roll are held zero.
# +
plt.figure(figsize=(15, 5))
t_plot = filt_error.index[::10] * dt / 3600
accel_err = ff_res.accel_err.iloc[::10]
accel_sd = ff_res.accel_sd.iloc[::10]
plt.subplot(131)
plt.plot(t_plot, accel_err.BIAS_1 + accel_sd.BIAS_1, 'b')
plt.plot(t_plot, accel_err.BIAS_1 - accel_sd.BIAS_1, 'b')
plt.hlines(accel_bias[0], *plt.xlim())
plt.xlabel("time, h")
plt.ylabel("rad/s")
plt.title("Accel 1 bias")
plt.subplot(132)
plt.plot(t_plot, accel_err.BIAS_2 + accel_sd.BIAS_2, 'b')
plt.plot(t_plot, accel_err.BIAS_2 - accel_sd.BIAS_2, 'b')
plt.hlines(accel_bias[1], *plt.xlim())
plt.xlabel("time, h")
plt.ylabel("rad/s")
plt.title("Accel 2 bias")
plt.subplot(133)
plt.plot(t_plot, accel_err.BIAS_3 + accel_sd.BIAS_3, 'b')
plt.plot(t_plot, accel_err.BIAS_3 - accel_sd.BIAS_3, 'b')
plt.hlines(accel_bias[2], *plt.xlim())
plt.xlabel("time, h")
plt.ylabel("rad/s")
plt.title("Accel 3 bias")
plt.tight_layout()
# -
| examples/ins_gps.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.1.0
# language: julia
# name: julia-1.1
# ---
# Join properties to DataFrames of the oc
# +
using DataFrames , Statistics
using Query, Glob
using PyPlot
import CSV
## directory
rootdir = ENV["GAIA_ROOT"]
wdir = "$rootdir/products"
votdir = "$rootdir/products/votable"
cd(wdir)
# +
# import final sample and original one with ages, etc..
sample= rootdir*"/master/notebooks/data/BrowseTargets.18292.1530479692.fake"
df1= CSV.read(sample, delim= "|")
df1.name= strip.(df1.name)
finalsample=wdir*"/votlist.finalSample_metric0.01.csv"
df2= CSV.read(finalsample, delim= ";")
ndat= size(df2)
## UGLY fix for name with a "-" in it
name=[]
for vot in df2.votname
t= split(vot,"-")
if length(t) == 3
t[1]= t[1]*"-"*t[2]
end
push!(name,t[1])
end
df2[:name]= name
df3= join(df1, df2, on = :name, makeunique= true)
i= 0
for na in df2.name
i += 1
if !(na in df1.name)
println(na)
println(i)
end
end
dfselect= sort(df3[:, [:votname, :name,:log_age]], order(:votname))
CSV.write("votlist.finalSample_metric0.01-age.csv", dfselect, delim= ";")
# +
## Joint age with ALL votname
cd(votdir)
files= glob("*vot")
cd(wdir)
dfsample = DataFrames.DataFrame(votname= String[],name= String[])
for f in files
t= split(f,"-")
if length(t) == 3
t[1]= t[1]*"-"*t[2]
end
nameoc= t[1]
push!(dfsample,[f, nameoc])
end
dfage= join(df1, dfsample, on = :name, makeunique= true)
dfselect= sort(dfage[:, [:votname,:log_age]], order(:votname))
println(dfsample)
CSV.write("votlist-age.csv", dfselect, delim= ";")
# -
PyPlot.hist(dfselect[:log_age], 50, density=true)
println("## Age distribution")
| notebooks/analysis/joinProperties.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="qHtYXGO7lr75"
# # Create CIFAR10 shards
# + id="LgebGLa8levP"
# !pip install webdataset
# + id="W0pgJukBlwat"
import torchvision
import webdataset as wds
from sklearn.model_selection import train_test_split
from pathlib import Path
import sys
# + [markdown] id="xdtIJvJSlx6G"
# ### Download train and test set
# * Loop Train set 26 times to get 1,300,000 train samples
# * Loop Test set 10 times to get 100,000 test samples
# + id="G2IU9ADrlzv8"
train_list = []
for x in range(26):
trainset = torchvision.datasets.CIFAR10(root="./", train=True, download=True)
train_list.extend(trainset)
print("Size of train_list:",sys.getsizeof(train_list))
print("Length of train_list:",len(train_list))
# + id="SxjhB0_Bl2H_"
test_list = []
for x in range(10):
testset = torchvision.datasets.CIFAR10(root="./", train=False, download=True)
test_list.extend(testset)
print("Size of test_list:",sys.getsizeof(test_list))
print("Length of test_list:",len(test_list))
# + [markdown] id="g4hue4Qsl5Q5"
# ## Create Tar Shards
# + [markdown] id="EHDh9C5SmlA2"
# Create local path for storing shards
# + id="sVWg5zy_l2Ao"
output_pth = "cifar-shards"
Path(output_pth).mkdir(parents=True, exist_ok=True)
Path(output_pth + "/train").mkdir(parents=True, exist_ok=True)
Path(output_pth + "/val").mkdir(parents=True, exist_ok=True)
# + [markdown] id="_DUTvpemmpt7"
# Write sharded tar files; 2,000 samples per shard
# + id="em__dalwl12o"
output_pth = "cifar-shards"
for name in [(train_list, "train"), (test_list, "val")]:
with wds.ShardWriter(
output_pth + "/" + str(name[1]) + "/" + "cifar-" + str(name[1]) + "-%06d.tar",
maxcount=2000,
) as sink:
for index, (image, cls) in enumerate(name[0]):
sink.write(
{"__key__": "%07d" % index, "ppm": image, "cls": cls}
)
# + [markdown] id="oPyQ_4rlm3kU"
# Copy shards to your GCS bucket
# + id="gRVzI2Mal-Xg"
# !gsutil -m cp -r cifar-shards/val gs:// # TODO: Add your GCS bucket location
# + id="tQOrAaAxjCER"
# !gsutil -m cp -r cifar-shards/train gs:// # TODO: Add your GCS bucket location
| pytorch_on_gcp/Create_CIFAR_shards.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from Bio import Seq
from Bio.Blast import NCBIWWW
from Bio.Blast import NCBIXML
seq_string='MIGRILGGLLEKNPPHLIVDCHGVGYEVDVPQSTFYNLPQTGEKVVLLTQQIVREDAHLLYGFGTVEERSTFRELLKISGIGARQALAVLSGMSVPELAQAVTLQDAGRLTRVPGIGKKTAERLLLELKGKLGADLGDLAGAASYSDHAIDILNALLALGYNEKEALAAIKNVPAGTGVSEGIKLALKALSKG'
seq_string = seq_string.replace('0', '')
testSeq = Seq.Seq(seq_string)
result_handle = NCBIWWW.qblast(program="blastp", database="nr", alignments = 3, descriptions=3, hitlist_size=3, sequence=seq_string)
all_titles = ""
blast_records = NCBIXML.parse(result_handle)
for blast_record in blast_records:
for alignment in blast_record.alignments:
for hsp in alignment.hsps:
print('****Alignment****')
print('Sequence:', alignment.title)
print('Length:', alignment.length)
print('Score:', hsp.score)
print('e value:', hsp.expect)
print(hsp.query)
print(hsp.match)
print(hsp.sbjct)
all_titles += alignment.title
blast_record = NCBIXML.read(result_handle)
all_titles = ""
for alignment in blast_record.alignments:
for hsp in alignment.hsps:
print('****Alignment****')
print('Sequence:', alignment.title)
print('Length:', alignment.length)
print('Score:', hsp.score)
print('e value:', hsp.expect)
print(hsp.query)
print(hsp.match)
print(hsp.sbjct)
all_titles += alignment.title
import re
m = re.findall('gi\|(.+?)\|', all_titles)
ids_to_search = ",".join(m)
a.keys()
from Bio import Entrez
handle = Entrez.efetch(db="Protein", id=ids_to_search, retmode="xml")
records = Entrez.read(handle)
handle.close()
for record in records:
for e in record["GBSeq_feature-table"]:
if e['GBFeature_key'] =="Protein":
for ee in e["GBFeature_quals"]:
if ee['GBQualifier_name'] =="EC_number":
print(ee["GBQualifier_value"])
ids_to_search
import tensorflow as tf
path = "C:\\Users\\Donatas\\Workspace\\Machine Learning\\SynBioAi\\PREnzyme\\weights\\protein\\wgan\\mini_sample\\sngan\\protein_tcn_batch_2_lr_0.0004_b1_0.0_b2_0.999_dim_16_image_3x192\\events.out.tfevents.1533809858.DONATAS-PC"
for e in tf.train.summary_iterator(path):
print(e)
# for v in e.summary.value:
# if v.tag == 'model/fake':
# print(v)
| notebooks/data_analysis/protein_blast.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
def splitDataFrameList(df,target_column,separator):
''' df = dataframe to split,
target_column = the column containing the values to split
separator = the symbol used to perform the split
returns: a dataframe with each entry for the target column separated, with each element moved into a new row.
The values in the other columns are duplicated across the newly divided rows.
'''
def splitListToRows(row,row_accumulator,target_column,separator):
split_row = row[target_column].split(separator)
for s in split_row:
new_row = row.to_dict()
new_row[target_column] = s
row_accumulator.append(new_row)
new_rows = []
df.apply(splitListToRows,axis=1,args = (new_rows,target_column,separator))
new_df = pd.DataFrame(new_rows)
return new_df
# Load csv file
df = pd.read_csv('data/outfit.csv')
#df.transpose()
# ### Pieces distribution in outfits
df2 = splitDataFrameList(df,"pieces",";")
#df2.transpose()
df2["pieces"].fillna("Nan").value_counts().plot.pie()
plt.show()
# ### Add hierarchial index
# +
df2_hier = df2.reset_index()
df2_hier.columns = 'id body outfit_id pieces style weather'.split()
df2_hier.transpose()
#df2_hier.pivot_table(values='body',index=['id','pieces'],columns=['weather'])
pivot_table = df2_hier.pivot_table(index=['outfit_id','id'], values=['pieces'], aggfunc=lambda x: "aaa"+x)
#pivot_table.transpose()
# -
# ### Weather distribution in outfits
df3 = splitDataFrameList(df,"weather",";")
df4 = df3[["outfit_id","weather"]]
df4.columns = 'outfit_id weather_category_id'.split()
#df4.transpose()
# Weather dictionary
weather = {
# -1: "Unknown",
1: "Freezing",
2: "Cold",
3: "Chilly",
4: "Fresh",
5: "Warm",
6: "Hot",
7: "Wind",
8: "Rain",
9: "Snow"
}
# Weather category dictionary
df_weathercat = pd.DataFrame(weather, index=['weather_category_name']).transpose()
# Add index column
df_weathercat.reset_index(inplace=True)
df_weathercat.columns = 'weather_category_id weather_category_name'.split()
# Change weather_id values to string - there is no "inplace" version of this!?
df_weathercat["weather_category_id"] = df_weathercat["weather_category_id"].apply(lambda x: str(x))
df_weathercat
# Left outer join.
pd.merge(df4,df_weathercat,how='left',on='weather_category_id')["weather_category_name"].fillna("Nan").value_counts().plot.pie()
plt.show()
| panda/outfit-visualization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Scalable GP Regression (w/ KISS-GP)
#
# ## Introduction
#
# If the function you are modeling has additive structure across its dimensions, then SKI can be one of the most efficient methods for your problem.
#
# Here, we model the kernel as a sum of kernels that each act on one dimension. Additive SKI (or KISS-GP) can work very well out-of-the-box on larger datasets (100,000+ data points) with many dimensions. This is a strong assumption though, and may not apply to your problem.
#
# This is the same as [the KISSGP Kronecker notebook](./KISSGP_Kronecker_Regression.ipynb), but applied to more dimensions.
# +
# Imports
import math
import torch
import gpytorch
import matplotlib.pyplot as plt
# Inline plotting
# %matplotlib inline
# -
# ### Set up train data
#
# Here we're learning a simple sin function - but in 2 dimensions
# +
# We store the data as a 10k 1D vector
# It actually represents [0,1]x[0,1] in cartesian coordinates
n = 30
train_x = torch.zeros(pow(n, 2), 2)
for i in range(n):
for j in range(n):
# Each coordinate varies from 0 to 1 in n=100 steps
train_x[i * n + j][0] = float(i) / (n-1)
train_x[i * n + j][1] = float(j) / (n-1)
train_x = train_x.cuda()
train_y = torch.sin(train_x[:, 0]) + torch.cos(train_x[:, 1]) * (2 * math.pi) + torch.randn_like(train_x[:, 0]).mul(0.01)
train_y = train_y / 4
# -
# ## The model
#
# As with the Kronecker example case, applying SKI to a multidimensional kernel is as simple as wrapping that kernel with a `GridInterpolationKernel`. You'll want to be sure to set `num_dims` though!
#
# To use an additive decomposition of the kernel, wrap it in an `AdditiveStructureKernel`.
#
# SKI has only one hyperparameter that you need to worry about: the grid size. For 1D functions, a good starting place is to use as many grid points as training points. (Don't worry - the grid points are really cheap to use, especially with an additive function!).
#
# If you want, you can also explicitly determine the grid bounds of the SKI approximation using the `grid_bounds` argument. However, it's easier if you don't use this argument - then GPyTorch automatically chooses the best bounds for you.
# +
class GPRegressionModel(gpytorch.models.ExactGP):
def __init__(self, train_x, train_y, likelihood):
super(GPRegressionModel, self).__init__(train_x, train_y, likelihood)
# SKI requires a grid size hyperparameter. This util can help with that
# We're setting Kronecker structure to False because we're using an additive structure decomposition
grid_size = gpytorch.utils.grid.choose_grid_size(train_x, kronecker_structure=False)
self.mean_module = gpytorch.means.ConstantMean()
self.covar_module = gpytorch.kernels.AdditiveStructureKernel(
gpytorch.kernels.GridInterpolationKernel(
gpytorch.kernels.ScaleKernel(
gpytorch.kernels.RBFKernel(),
), grid_size=128, num_dims=1
), num_dims=2
)
def forward(self, x):
mean_x = self.mean_module(x)
covar_x = self.covar_module(x)
return gpytorch.distributions.MultivariateNormal(mean_x, covar_x)
likelihood = gpytorch.likelihoods.GaussianLikelihood()
model = GPRegressionModel(train_x, train_y, likelihood).cuda()
# -
# ## Train the model hyperparameters
#
# +
# Optimize the model
model.train()
likelihood.train()
# Use the adam optimizer
optimizer = torch.optim.Adam([
{'params': model.parameters()}, # Includes GaussianLikelihood parameters
], lr=0.1)
# "Loss" for GPs - the marginal log likelihood
mll = gpytorch.mlls.ExactMarginalLogLikelihood(likelihood, model)
# Sometimes we get better performance on the GPU when we don't use Toeplitz math
# for SKI. This flag controls that
def train(num_iter):
with gpytorch.settings.use_toeplitz(False):
for i in range(num_iter):
optimizer.zero_grad()
output = model(train_x)
loss = -mll(output, train_y)
loss.backward()
print('Iter %d/%d - Loss: %.3f' % (i + 1, num_iter, loss.item()))
optimizer.step()
# %time train(num_iter=20)
# +
# Set into eval mode
model.eval()
likelihood.eval()
# Create 100 test data points
# Over the square [0,1]x[0,1]
n = 30
test_x = torch.zeros(int(pow(n, 2)), 2).cuda()
for i in range(n):
for j in range(n):
test_x[i * n + j][0] = float(i) / (n-1)
test_x[i * n + j][1] = float(j) / (n-1)
with torch.no_grad(), gpytorch.settings.fast_pred_var():
observed_pred = likelihood(model(test_x))
pred_labels = observed_pred.mean.view(n, n)
# Calculate the true test values
test_y_actual = torch.sin(test_x.data[:, 0]) + (torch.cos(test_x.data[:, 1]) * (2 * math.pi))
test_y_actual = test_y_actual.view(n, n) / 8
delta_y = torch.abs(pred_labels - test_y_actual)
# Define a plotting function
def ax_plot(f, ax, y_labels, title):
im = ax.imshow(y_labels)
ax.set_title(title)
f.colorbar(im)
# Make a plot of the predicted values
f, observed_ax = plt.subplots(1, 1, figsize=(4, 3))
ax_plot(f, observed_ax, pred_labels.cpu(), 'Predicted Values (Likelihood)')
# Make a plot of the actual values
f, observed_ax2 = plt.subplots(1, 1, figsize=(4, 3))
ax_plot(f, observed_ax2, test_y_actual.cpu(), 'Actual Values (Likelihood)')
# Make a plot of the errors
f, observed_ax3 = plt.subplots(1, 1, figsize=(4, 3))
ax_plot(f, observed_ax3, delta_y.cpu(), 'Absolute Error Surface')
# -
| examples/05_Scalable_GP_Regression_Multidimensional/KISSGP_Additive_Regression_CUDA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Cold Start Problem
#
# In the previous notebook, you learned about the **Cold Start Problem** first hand. In cases where you are introduced to a new user or new movie, collaborative flitering is not helpful as a technique to make predictions.
#
# Instead, you will need to use one of the techniques from the previous lesson like content based recommendations for new items or rank based recommendations for new users.
#
# As a final step to completing out our recommendation system, we will build in these edge cases. Run the cell below to get started.
#
# ### Matrix Factorization - Collaborative Filtering Where Possible
#
# Notice the following information is available by running the below cell:
#
# `1.` **reviews** - a dataframe of reviews
#
# `2.` **movies** - a dataframe of movies
#
# `3.` **create_train_test** - a function for creating the training and validation datasets
#
# `4.` **predict_rating** - a function that takes a user and movie and gives a prediction using FunkSVD
#
# `5.` **train_df** and **val_df** - the training and test datasets used in the previous notebook
#
# `6.` **user_mat** and **movie_mat** - the u and v matrices from FunkSVD
#
# `7.` **train_data_df** - a user-movie matrix with ratings where available. FunkSVD was performed on this matrix
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
import pickle
# Read in the datasets
movies = pd.read_csv('data/movies_clean.csv')
reviews = pd.read_csv('data/reviews_clean.csv')
del movies['Unnamed: 0']
del reviews['Unnamed: 0']
def create_train_test(reviews, order_by, training_size, testing_size):
'''
INPUT:
reviews - (pandas df) dataframe to split into train and test
order_by - (string) column name to sort by
training_size - (int) number of rows in training set
testing_size - (int) number of columns in the test set
OUTPUT:
training_df - (pandas df) dataframe of the training set
validation_df - (pandas df) dataframe of the test set
'''
reviews_new = reviews.sort_values(order_by)
training_df = reviews_new.head(training_size)
validation_df = reviews_new.iloc[training_size:training_size+testing_size]
return training_df, validation_df
def predict_rating(user_matrix, movie_matrix, user_id, movie_id):
'''
INPUT:
user_matrix - user by latent factor matrix
movie_matrix - latent factor by movie matrix
user_id - the user_id from the reviews df
movie_id - the movie_id according the movies df
OUTPUT:
pred - the predicted rating for user_id-movie_id according to FunkSVD
'''
# Create series of users and movies in the right order
user_ids_series = np.array(train_data_df.index)
movie_ids_series = np.array(train_data_df.columns)
# User row and Movie Column
user_row = np.where(user_ids_series == user_id)[0][0]
movie_col = np.where(movie_ids_series == movie_id)[0][0]
# Take dot product of that row and column in U and V to make prediction
pred = np.dot(user_matrix[user_row, :], movie_matrix[:, movie_col])
return pred
# Use our function to create training and test datasets
train_df, val_df = create_train_test(reviews, 'date', 8000, 2000)
# Create user-by-item matrix - this will keep track of order of users and movies in u and v
train_user_item = train_df[['user_id', 'movie_id', 'rating', 'timestamp']]
train_data_df = train_user_item.groupby(['user_id', 'movie_id'])['rating'].max().unstack()
train_data_np = np.array(train_data_df)
# Read in user and movie matrices
user_file = open("user_matrix", 'rb')
user_mat = pickle.load(user_file)
user_file.close()
movie_file = open("movie_matrix", 'rb')
movie_mat = pickle.load(movie_file)
movie_file.close()
# -
# ### Validating Predictions
#
# Unfortunately, you weren't able to make predictions on every user-movie combination in the test set, as some of these users or movies were new.
#
# However, you can validate your predictions for the user-movie pairs that do exist in the user_mat and movie_mat matrices.
#
# `1.` Complete the function below to see how far off we were on average across all of the predicted ratings.
def validation_comparison(val_df, user_mat=user_mat, movie_mat=movie_mat):
'''
INPUT:
val_df - the validation dataset created in the third cell above
user_mat - U matrix in FunkSVD
movie_mat - V matrix in FunkSVD
OUTPUT:
rmse - RMSE of how far off each value is from it's predicted value
perc_rated - percent of predictions out of all possible that could be rated
actual_v_pred - a 10 x 10 grid with counts for actual vs predicted values
'''
val_users = np.array(val_df['user_id'])
val_movies = np.array(val_df['movie_id'])
val_ratings = np.array(val_df['rating'])
sse = 0
num_rated = 0
preds, acts = [], []
actual_v_pred = np.zeros((10,10))
for idx in range(len(val_users)):
try:
pred = predict_rating(user_mat, movie_mat, val_users[idx], val_movies[idx])
sse += (val_ratings[idx] - pred)**2
num_rated+=1
preds.append(pred)
acts.append(val_ratings[idx])
actual_v_pred[11-int(val_ratings[idx]-1), int(round(pred)-1)]+=1
except:
continue
rmse = np.sqrt(sse/num_rated)
perc_rated = num_rated/len(val_users)
return rmse, perc_rated, actual_v_pred, preds, acts
# How well did we do?
rmse, perc_rated, actual_v_pred, preds, acts = validation_comparison(val_df)
print(rmse, perc_rated)
sns.heatmap(actual_v_pred);
plt.xticks(np.arange(10), np.arange(1,11));
plt.yticks(np.arange(10), np.arange(1,11));
plt.xlabel("Predicted Values");
plt.ylabel("Actual Values");
plt.title("Actual vs. Predicted Values");
plt.figure(figsize=(8,8))
plt.hist(acts, normed=True, alpha=.5, label='actual');
plt.hist(preds, normed=True, alpha=.5, label='predicted');
plt.legend(loc=2, prop={'size': 15});
plt.xlabel('Rating');
plt.title('Predicted vs. Actual Rating');
# `2.` We didn't do so bad on making those predictions! But, how many user-movie pairs were we unable to make predictions for? Use the cell below to answer this question.
# From the above, this can be calculated as follows:
print("Number not rated {}".format(int(len(val_df['rating'])*(1-perc_rated))))
print("Number rated {}.".format(int(len(val_df['rating'])*perc_rated)))
# ### Content Based For New Movies
#
# If all of the above went well, you will notice we still have work to do! We need to bring in a few things we picked up from the last lesson to use for those new users and movies. Below is the code used to make the content based recommendations, which found movies that were similar to one another. This was from **5_Content_Based_Recommendations** in the previous lesson.
#
# The below function **find_similar_movies** will provide similar movies to any movie based only on content.
#
# Run the cell below to gain access to the content based similarity functions.
# +
# Subset so movie_content is only using the dummy variables for each genre and the 3 century based year dummy columns
movie_content = np.array(movies.iloc[:,4:])
# Take the dot product to obtain a movie x movie matrix of similarities
dot_prod_movies = movie_content.dot(np.transpose(movie_content))
def find_similar_movies(movie_id):
'''
INPUT
movie_id - a movie_id
OUTPUT
similar_movies - an array of the most similar movies by title
'''
# find the row of each movie id
movie_idx = np.where(movies['movie_id'] == movie_id)[0][0]
# find the most similar movie indices - to start I said they need to be the same for all content
similar_idxs = np.where(dot_prod_movies[movie_idx] == np.max(dot_prod_movies[movie_idx]))[0]
# pull the movie titles based on the indices
similar_movies = np.array(movies.iloc[similar_idxs, ]['movie'])
return similar_movies
def get_movie_names(movie_ids):
'''
INPUT
movie_ids - a list of movie_ids
OUTPUT
movies - a list of movie names associated with the movie_ids
'''
movie_lst = list(movies[movies['movie_id'].isin(movie_ids)]['movie'])
return movie_lst
# -
# ### Rank Based For New Users
#
# From the above two code cells, we have a way to make recommendations for movie-user pairs that have ratings in any part of our user-movie matrix. We also have a way to make ratings for movies that have never received a rating using movie similarities.
#
# In this last part here, we need a way to make recommendations to new users. For this, our functions from **2_Most_Popular_Recommendations** in Lesson 1 will come in handy. Run the cell below to have these functions available.
#
# Run the cell below to gain access to the rank based functions.
# +
def create_ranked_df(movies, reviews):
'''
INPUT
movies - the movies dataframe
reviews - the reviews dataframe
OUTPUT
ranked_movies - a dataframe with movies that are sorted by highest avg rating, more reviews,
then time, and must have more than 4 ratings
'''
# Pull the average ratings and number of ratings for each movie
movie_ratings = reviews.groupby('movie_id')['rating']
avg_ratings = movie_ratings.mean()
num_ratings = movie_ratings.count()
last_rating = pd.DataFrame(reviews.groupby('movie_id').max()['date'])
last_rating.columns = ['last_rating']
# Add Dates
rating_count_df = pd.DataFrame({'avg_rating': avg_ratings, 'num_ratings': num_ratings})
rating_count_df = rating_count_df.join(last_rating)
# merge with the movies dataset
movie_recs = movies.set_index('movie_id').join(rating_count_df)
# sort by top avg rating and number of ratings
ranked_movies = movie_recs.sort_values(['avg_rating', 'num_ratings', 'last_rating'], ascending=False)
# for edge cases - subset the movie list to those with only 5 or more reviews
ranked_movies = ranked_movies[ranked_movies['num_ratings'] > 4]
return ranked_movies
def popular_recommendations(user_id, n_top, ranked_movies):
'''
INPUT:
user_id - the user_id (str) of the individual you are making recommendations for
n_top - an integer of the number recommendations you want back
ranked_movies - a pandas dataframe of the already ranked movies based on avg rating, count, and time
OUTPUT:
top_movies - a list of the n_top recommended movies by movie title in order best to worst
'''
top_movies = list(ranked_movies['movie'][:n_top])
return top_movies
# -
# ### Now For Your Task
#
# The above cells set up everything we need to use to make predictions. Your task is to write a function, which uses the above information as necessary to provide recommendations for every user in the **val_df** dataframe. There isn't one right way to do this, but using a blend between the three could be your best bet.
#
# You can see the blended approach I used in the video on the next page, but feel free to be creative with your solution!
#
# `3.` Use the function below along with the document strings to assist with completing the task for this notebook.
def make_recommendations(_id, _id_type='movie', train_data=train_data_df,
train_df=train_df, movies=movies, rec_num=5, user_mat=user_mat):
'''
INPUT:
_id - either a user or movie id (int)
_id_type - "movie" or "user" (str)
train_data - dataframe of data as user-movie matrix
train_df - dataframe of training data reviews
movies - movies df
rec_num - number of recommendations to return (int)
user_mat - the U matrix of matrix factorization
movie_mat - the V matrix of matrix factorization
OUTPUT:
recs - (array) a list or numpy array of recommended movies like the
given movie, or recs for a user_id given
'''
# if the user is available from the matrix factorization data,
# I will use this and rank movies based on the predicted values
# For use with user indexing
val_users = train_data_df.index
rec_ids = create_ranked_df(movies, train_df)
if _id_type == 'user':
if _id in train_data.index:
# Get the index of which row the user is in for use in U matrix
idx = np.where(val_users == _id)[0][0]
# take the dot product of that row and the V matrix
preds = np.dot(user_mat[idx,:],movie_mat)
# pull the top movies according to the prediction
indices = preds.argsort()[-rec_num:][::-1] #indices
rec_ids = train_data_df.columns[indices]
rec_names = get_movie_names(rec_ids)
else:
# if we don't have this user, give just top ratings back
rec_names = popular_recommendations(_id, rec_num, ranked_movies)
# Find similar movies if it is a movie that is passed
else:
rec_ids = find_similar_movies(_id)
rec_names = get_movie_names(rec_ids)
return rec_ids, rec_names
make_recommendations(48, 'user')
len(set(val_df['user_id']))
# Make recommendations
user_recs_dict_with_top = dict()
for user_id in set(val_df['user_id']):
user_recs_dict_with_top[user_id] = make_recommendations(user_id, 'user')[1]
cnter = 0
for user, rec in user_recs_dict_with_top.items():
if cnter < 12:
print("For user {}, our recommendations are: \n {}".format(user, rec))
cnter+=1
else:
break
# **This recommendation style looks like it may do okay with accuracy, but it seems like a lot of the same movies are showing up. When we think back to serendipity, novelty, and diversity as means of a good recommendation system, this set of recommendations still isn't great. We might consider providing some content based recommendations from movies an individual has watched along with these recommendations to meet those categories of a good recommender.**
| lessons/Recommendations/2_Matrix_Factorization_for_Recommendations/4_Cold_Start_Problem_Solution.ipynb |
# # Training a new tokenizer from an old one
# Install the Transformers and Datasets libraries to run this notebook.
# !pip install datasets transformers[sentencepiece]
# +
from datasets import load_dataset
# This can take a few minutes to load, so grab a coffee or tea while you wait!
raw_datasets = load_dataset("code_search_net", "python")
# -
raw_datasets["train"]
print(raw_datasets["train"][123456]["whole_func_string"])
# +
# Don't uncomment the following line unless your dataset is small.
# training_corpus = [raw_datasets["train"][i: i + 1000]["whole_func_string"] for i in range(0, len(raw_datasets["train"]), 1000)]
# -
training_corpus = (raw_datasets["train"][i: i + 1000]["whole_func_string"] for i in range(0, len(raw_datasets["train"]), 1000))
gen = (i for i in range(10))
print(list(gen))
print(list(gen))
def get_training_corpus():
return (raw_datasets["train"][i: i + 1000]["whole_func_string"] for i in range(0, len(raw_datasets["train"]), 1000))
def get_training_corpus():
dataset = raw_datasets["train"]
for start_idx in range(0, len(dataset), 1000):
samples = dataset[start_idx: start_idx + 1000]
yield samples["whole_func_string"]
# +
from transformers import AutoTokenizer
old_tokenizer = AutoTokenizer.from_pretrained("gpt2")
# +
example = '''def add_numbers(a, b):
"""Add the two numbers `a` and `b`."""
return a + b'''
tokens = old_tokenizer.tokenize(example)
tokens
# -
tokenizer = old_tokenizer.train_new_from_iterator(training_corpus, 52000)
tokens = tokenizer.tokenize(example)
tokens
print(len(tokens))
print(len(old_tokenizer.tokenize(example)))
example = """class LinearLayer():
def __init__(self, input_size, output_size):
self.weight = torch.randn(input_size, output_size)
self.bias = torch.zeros(output_size)
def __call__(self, x):
return x @ self.weights + self.bias
"""
tokenizer.tokenize(example)
tokenizer.save_pretrained("code-search-net-tokenizer")
tokenizer.push_to_hub("code-search-net-tokenizer")
# Replace sgugger below by your actual namespace to use your own tokenizer.
tokenizer = AutoTokenizer.from_pretrained("sgugger/code-search-net-tokenizer")
| course/chapter6/section2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Question 1:
# + active=""
# Create a function that takes a list and string. The function should remove the letters
# in the string from the list, and return the list.
#
# Examples
#
# remove_letters(["s", "t", "r", "i", "n", "g", "w"], "string") ["w"]
# remove_letters(["b", "b", "l", "l", "g", "n", "o", "a", "w"], "balloon") ["b", "g","w"]
# remove_letters(["d", "b", "t", "e", "a", "i"], "edabit") []
# -
# Answer :
# +
def remove_letters(lst,str_):
for i in str_:
if i in lst:
lst.remove(i)
return lst
print(remove_letters(["s", "t", "r", "i", "n", "g", "w"], "string"))
print(remove_letters(["b", "b", "l", "l", "g", "n", "o", "a", "w"], "balloon"))
print(remove_letters(["d", "b", "t", "e", "a", "i"], "edabit"))
# -
# Question 2:
# + active=""
# A block sequence in three dimensions. We can write a formula for this one:
# Create a function that takes a number (step) as an argument and returns the amount of blocks in that step.
#
# Examples
#
# blocks(1) 5
# blocks(5) 39
# blocks(2) 12
# -
# Answer :
# +
def blocks(num):
blocks_num = 5
for i in range(0,num-1):
blocks_num += 7 + i
return blocks_num
print(blocks(1))
print(blocks(5))
print(blocks(2))
# -
# Question 3:
# + active=""
# Create a function that subtracts one positive integer from another,
# without using any arithmetic operators such as -, %, /, +, etc.
#
# Examples
#
# my_sub(5, 9) 4
# my_sub(10, 30) 20
# my_sub(0, 0) 0
# -
# Answer :
# +
def my_sub(a,b):
return len(range(a,b))
print(my_sub(5,9))
print(my_sub(10,30))
print(my_sub(0,0))
# -
# Question 4:
# + active=""
# Create a function that takes a string containing money in dollars and
# pounds sterling (seperated by comma) and returns the sum of dollar bills only, as an integer.
# For the input string:
# - Each amount is prefixed by the currency symbol: $ for dollars and £ for pounds.
# - Thousands are represented by the suffix k. i.e. $4k = $4,000 and £40k = £40,000
#
# Examples
#
# add_bill("d20,p40,p60,d50") 20 + 50 = 70
# add_bill("p30,d20,p60,d150,p360") 20 + 150 = 170
# add_bill("p30,d2k,p60,d200,p360") 2 * 1000 + 200 = 2200
# -
# Answer :
# +
import re
def add_bill(str_):
str1 = re.findall("d[0-9k]+",str_)
sum_ = sum([int(i[1::]) if "k" not in i else int(i[1:~0])*1000 for i in str1])
return sum_
print(add_bill("d20,p40,p60,d50"))
print(add_bill("p30,d20,p60,d150,p360"))
print(add_bill("p30,d2k,p60,d200,p360"))
# -
# Question 5:
# + active=""
# Create a function that flips a horizontal list into a vertical list, and a vertical list
# into a horizontal list.In other words, take an 1 x n list (1 row + n columns) and
# flip it into a n x 1 list (n rows and 1 column), and vice versa.
#
# Examples
#
# flip_list([1, 2, 3, 4]) [[1], [2], [3], [4]]
# # Take a horizontal list and flip it vertical.
# flip_list([[5], [6], [9]]) [5, 6, 9]
# # Take a vertical list and flip it horizontal.
# flip_list([]) []
# -
# Answer :
# +
def flip_list(lst):
flip = []
for i in lst:
if type(lst[0]) == list:
flip.append(i[0])
else:
flip.append([i])
return flip
print(flip_list([1, 2, 3, 4]))
print(flip_list([[5], [6], [9]]))
print(flip_list([]))
# -
| Python Advance Programming Assignment/Assignment_13.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # GOOGLE PLAYSTORE ANALYSIS
# The dataset used in this analysis is taken from [kaggle datasets](https://www.kaggle.com/datasets)
# In this analysis we took a raw data which is in csv format and then converted it into a dataframe.Performed some operations, cleaning of the data and finally visualizing some necessary conclusions obtained from it.
# Let's import necessary libraries required for the analysis
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# Convert the csv file into dataframe using pandas
df=pd.read_csv('googleplaystore.csv')
df.head(5)
# This is the data we obtained from the csv file.Let's see some info about this dataframe
df.info()
# This dataframe consists of 10841 entries ie information about 10841 apps.
# It tells about the category to which the app belongs,rating given by the users,size of the app,number of reviews given,count of number of installs and some other information
# # DATA CLEANING
# Some columns have in-appropriate data,data types.This columns needed to be cleaned to perform the analysis.
# ##### SIZE :
# This column has in-appropriate data type.This needed to be converted into numeric type after converting every value into MB's
#
# For example, the size of the app is in “string” format. We need to convert it into a numeric value. If the size is “10M”, then ‘M’ was removed to get the numeric value of ‘10’. If the size is “512k”, which depicts app size in kilobytes, the first ‘k’ should be removed and the size should be converted to an equivalent of ‘megabytes’.
df['Size'] = df['Size'].map(lambda x: x.rstrip('M'))
df['Size'] = df['Size'].map(lambda x: str(round((float(x.rstrip('k'))/1024), 1)) if x[-1]=='k' else x)
df['Size'] = df['Size'].map(lambda x: np.nan if x.startswith('Varies') else x)
# 10472 has in-appropriate data in every column, may due to entry mistake.So we are removing that entry from the table
df.drop(10472,inplace=True)
# By using pd.to_numeric command we are converting into numeric type
df['Size']=df['Size'].apply(pd.to_numeric)
# ##### Installs :
# The value of installs is in “string” format. It contains numeric values with commas. It should be removed. And also, the ‘+’ sign should be removed from the end of each string.
df['Installs'] = df['Installs'].map(lambda x: x.rstrip('+'))
df['Installs'] = df['Installs'].map(lambda x: ''.join(x.split(',')))
# By using pd.to_numeric command we are converting it into numeric data type
df['Installs']=df['Installs'].apply(pd.to_numeric)
# ##### Reviews :
# The reviews column is in string format and we need to convert it into numeric type
df['Reviews']=df['Reviews'].apply(pd.to_numeric)
# After cleaning some columns and rows we obtained the required format to perform the analysis
df.head(5)
# # DATA VISUALIZATION
# In this we are taking a parameter as reference and checking the trend of another parameter like whether there is a rise or fall,which category are more,what kinds are of more intrest and so on.
# ###### Basic pie chart to view distribution of apps across various categories
fig, ax = plt.subplots(figsize=(10, 10), subplot_kw=dict(aspect="equal"))
number_of_apps = df["Category"].value_counts()
labels = number_of_apps.index
sizes = number_of_apps.values
ax.pie(sizes,labeldistance=2,autopct='%1.1f%%')
ax.legend(labels=labels,loc="right",bbox_to_anchor=(0.9, 0, 0.5, 1))
ax.axis("equal")
plt.show()
# ## App count for certain range of Ratings
# In this we are finding the count of apps for each range from 0 to 5 ie how many apps have more rating,how many are less rated.
bins=pd.cut(df['Rating'],[0.0,1.0,2.0,3.0,4.0,5.0])
rating_df=pd.DataFrame(df.groupby(bins)['App'].count())
rating_df.reset_index(inplace=True)
rating_df
plt.figure(figsize=(12, 6))
axis=sns.barplot('Rating','App',data=rating_df);
axis.set(ylabel= "App count",title='APP COUNT STATISTICS ACCORDING TO RATING');
# We can see that most of the apps are with rating 4 and above and very less apps have rating below 2.
# ## Top5 Apps with highest review count
# In this we are retrieving the top5 apps with more number of reviews and seeing it visually how their review count is changing.
reviews_df=df.sort_values('Reviews').tail(15).drop_duplicates(subset='App')[['App','Reviews','Rating']]
reviews_df
plt.figure(figsize=(12, 6))
axis=sns.lineplot(x="App",y="Reviews",data=reviews_df)
axis.set(title="Top 5 most Reviewed Apps");
sns.set_style('darkgrid')
# Facebook has more reviews compared to other apps in the playstore
# ## Which content type Apps are more in playstore
# In this we are grouping the apps according to their content type and visually observing the result
content_df=pd.DataFrame(df.groupby('Content Rating')['App'].count())
content_df.reset_index(inplace=True)
content_df
plt.figure(figsize=(12, 6))
plt.bar(content_df['Content Rating'],content_df['App']);
plt.xlabel('Content Rating')
plt.ylabel('App count')
plt.title('App count for different Contents');
# Most of the apps in playstore can be used by everyone irrespective of the age.Only 3 apps are A rated
# ##### ---------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------
# ## Free vs Paid Apps
# Let's see variations considering type of App ie paid and free apps
Type_df=df.groupby('Type')[['App']].count()
Type_df['Rating']=df.groupby('Type')['Rating'].mean()
Type_df.reset_index(inplace=True)
Type_df
# We found the number of apps that are freely available and their average rating and also number of paid apps and their average rating.
fig, axes = plt.subplots(1, 2, figsize=(18, 6))
axes[0].bar(Type_df.Type,Type_df.App)
axes[0].set_title("Number of free and paid apps")
axes[0].set_ylabel('App count')
axes[1].bar(Type_df.Type,Type_df.Rating)
axes[1].set_title('Average Rating of free and paid apps')
axes[1].set_ylabel('Average Rating');
# #### Conclusion
# Average rating of Paid Apps is more than Free apps.So,we can say that paid apps are trust worthy and we can invest in them
# ##### -----------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------
# ## Max Installs
# In this we are finding the apps with more number of installs and as we dont have exact count of installs we got around 20 apps with 1B+ downloads
# From the 20 apps we will see some analysis of what types are more installed
max_installs=df.loc[df['Installs']==df.Installs.max()][['App','Category','Reviews','Rating','Installs','Content Rating']]
max_installs=max_installs.drop_duplicates(subset='App')
max_installs
# These are the 20 apps which are with 1B+ downloads
# ### Which App has more rating and trend of 20 apps rating
plt.figure(figsize=(12, 6))
sns.barplot('Rating','App',data=max_installs);
# We can see that Google photos,Instagram and Subway Surfers are the most rated Apps which have 1B+ downloads.
# Though the Apps are used by 1B+ users they have a good rating too
# ### Which content Apps are most Installed
# We will group the most installed apps according to their content and see which content apps are most installed
content_max_df=pd.DataFrame(max_installs.groupby('Content Rating')['App'].count())
content_max_df.reset_index(inplace=True)
content_max_df
plt.figure(figsize=(12, 6))
axis=sns.barplot('Content Rating','App',data=content_max_df);
axis.set(ylabel= "App count",title='Max Installed APP COUNT STATISTICS ACCORDING TO Content RATING');
# More than 10 apps are of type which can be used by any age group and about 8 apps are teen aged apps.Only 1 app is to used by person with age 10+
# ### Which category Apps are more Installed
# In this we will group the most installed apps according to their category and see which category are on high demand
category_max_df=pd.DataFrame(max_installs.groupby('Category')['App'].count())
category_max_df.reset_index(inplace=True)
category_max_df
plt.figure(figsize=(12, 6))
axis=sns.barplot('App','Category',data=category_max_df);
plt.plot(category_max_df.App,category_max_df.Category,'o--r')
axis.set(ylabel= "App count",title='Max Installed APP COUNT STATISTICS ACCORDING TO Category');
# Communication Apps are mostly installed by people like facebook,whatsapp,instagram..and then social apps are in demand.
# #### Conclusion
# The most installed apps ie apps with downloads more than 1 Billion are mostly Communication related apps and can be used by any age group without any restriction and also have high user rating.
# ###### ----------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------
# ## Final Conclusion
# This analysis is mostly based on the existing data in the dataset , how one parameter is changing with respect to another parameter,whether paid apps are trust worthy and intrests of people towards some particular categories.
#
# This analysis can further be extended to predict the number of installs and ratings would be if a new app is launched by some Machine Learning algortihms and models.
#
# ##### -------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------
# ### THANK YOU :)
| playstore analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Bibliotecas:
# +
#importanto bibliotecas
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
from sklearn import datasets, linear_model, preprocessing
import statsmodels.api as sm
from sklearn.metrics import mean_squared_error, r2_score
from sklearn.preprocessing import StandardScaler
from datetime import datetime
from sklearn.linear_model import Lasso, LassoCV
from sklearn.impute import SimpleImputer
from sklearn.tree import DecisionTreeRegressor
from sklearn.metrics import mean_squared_error as MSE
from sklearn.model_selection import train_test_split
# -
# #### Lendo csv e removendo algumas colunas redundantes:
Y_train=pd.read_csv('shared/bases_budokai_ufpr/produtividade_soja_modelagem.csv')
df=pd.read_csv('shared/bases_budokai_ufpr/agroclimatology_budokai.csv')
Y_train.head()
Y_train['nivel'].unique()
# +
#Y_codigo=Y_train['codigo_ibge']
Y_train=Y_train.drop(columns=['nivel', 'name'])
# -
Y_train.head()
# ### De acordo com as respostas esperadas temos:
codigo_saida=[4102000,4104303,4104428,4104808,4104907,4109401,4113205,4113700,4113734,4114005,4114401,4117701,4117909,4119608,4119905,4127007,4127403,4127502,4127700,4128005
]
# +
dataset=pd.DataFrame(columns=df.keys())
#dataset.loc[dataset['codigo_ibge']==codigo_saida[j]]
for k in codigo_saida:
dataset=pd.concat([df.loc[df['codigo_ibge']==k],dataset])
dataset=dataset.reset_index(drop=True)
# -
dataset.info()
dataset['codigo_ibge']=list(map(int,dataset['codigo_ibge']))
#transformar o codigo em inteiro novamente
#df.loc[df['codigo_ibge']==codigo_saida[0]]
# ### Vamos começar transformando os dados de data em 'datetime' para melhor agrupar em semanas, meses ou anos.
# +
def t_data_day(data):
data=str(data)
data=datetime.strptime(data,"%Y%m%d")
return data.day
def t_data_month(data):
data=str(data)
data=datetime.strptime(data,"%Y%m%d")
return data.month
def t_data_year(data):
data=str(data)
data=datetime.strptime(data,"%Y%m%d")
return data.year
#return data.day, data.month, data.year
# -
dataset['day']=list(map(t_data_day, dataset['data']))
dataset['month']=list(map(t_data_month, dataset['data']))
dataset['year']=list(map(t_data_year, dataset['data']))
dataset.head()
# ## *Uma feature importante é o valor da produção do ano passado, inserir depois.
dataset=dataset.drop(dataset.loc[dataset['year']==2003].index).reset_index(drop=True)
# #### A ideia posteriormente aqui é fazer um algoritmo de pesos para cada mês (media ponderada)
# ### A titulo de teste do sistema podemos implementar um algoritmo rápido para previsões baseado na média dos valores anuais.
dataset=dataset.drop(columns=['data','latitude','longitude','day','month'])
# #### Para cada codigo postal agrupamos os dados anuais pela média.
# +
codigos=dataset['codigo_ibge'].unique()
datanew=pd.DataFrame(columns=dataset.keys())
for i in codigos:
aux = dataset.loc[dataset['codigo_ibge']==i].groupby(by='year').agg('mean').reset_index()
datanew=pd.concat([datanew,aux])
datanew=datanew.reset_index(drop=True)
# -
datanew.head()
X_train=datanew.loc[datanew['year']<2018]
X_test=datanew.loc[datanew['year']>2017]
# #### Scalando os dados:
# +
colunas_=list(X_train.keys())
colunas_.remove('year')
colunas_.remove('codigo_ibge')
#removendo os dados que não serão escalados
# +
scaler=StandardScaler()
X_train_scaled=scaler.fit_transform(X_train[colunas_])
X_train_scaled=pd.DataFrame(X_train_scaled, columns=colunas_)
X_test_scaled=scaler.transform(X_test[colunas_])
X_test_scaled=pd.DataFrame(X_test_scaled, columns=colunas_)
# -
# ### Precisamos reinserir o codigo e ano e resetar o indice.
X_train_scaled[['codigo_ibge','year']]=X_train[['codigo_ibge','year']].reset_index(drop=True)
X_test_scaled[['codigo_ibge','year']]=X_test[['codigo_ibge','year']].reset_index(drop=True)
X_train_scaled
#k-fold para achar melhor valor de alpha
model = LassoCV(cv=5, random_state=0, max_iter=10000)
# +
df_out=pd.DataFrame(columns=['codigo_ibge','2018','2019','2020'])
for i in codigo_saida:
X_train_menor=X_train_scaled.loc[X_train_scaled['codigo_ibge']==i].drop(columns=['codigo_ibge','year'])
Y_train_menor=Y_train[Y_train['codigo_ibge']==i].drop(columns='codigo_ibge').T
model.fit(X_train_menor, Y_train_menor)
##########################
# Teste com árvore de decisão
dt = DecisionTreeRegressor(max_depth=4, min_samples_leaf=0.1, random_state=3)
dt.fit(X_train_menor,Y_train_menor)
##############################################3
lasso_t= Lasso(alpha=model.alpha_, max_iter= 10000).fit(X_train_menor,Y_train_menor)
print(f'\n Alpha usado: {model.alpha_}')
print('Features non-zero para {}: {}'.format(i, np.sum(lasso_t.coef_ != 0)))
print('Features com valores non-zero (ordenados pela magnitude absoluta):')
for e in sorted (list(zip(list(X_train), lasso_t.coef_)), key = lambda e: -abs(e[1])):
if e[1] != 0:
print('\t{}, {:.3f}'.format(e[0], e[1]))
X_test_menor=X_test_scaled.loc[X_test_scaled['codigo_ibge']==i].drop(columns=['codigo_ibge','year'])
##################
Y_prd = dt.predict(X_test_menor)
predicoes = Y_prd
#Analisando a acuracia
##################
#predicoes=lasso_t.predict(X_test_menor)
data_saida={'codigo_ibge':i, '2018':predicoes[0], '2019':predicoes[1], '2020':predicoes[2]}
data_saida=pd.DataFrame([data_saida])
df_out=pd.concat([df_out,data_saida])
# -
df_out=df_out.reset_index(drop=True)
df_out
df_out.to_csv('submission.csv',index=False)
| notebook_v1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
"""
Author: <NAME>
Modifier: <NAME>, <NAME>
"""
import numpy as np
import pandas as pd
import datetime as dt
from datetime import datetime
# %matplotlib inline
import matplotlib.pyplot as plt
import plotly as py
import plotly.graph_objs as go
from plotly.offline import download_plotlyjs, init_notebook_mode, plot, iplot
from sklearn.decomposition import PCA
from sklearn.preprocessing import StandardScaler
from mpl_toolkits.mplot3d import Axes3D
from sklearn.model_selection import KFold
from sklearn.neighbors import KNeighborsRegressor
from sklearn.tree import DecisionTreeRegressor
from sklearn.metrics import mean_squared_error
from sklearn.cluster import KMeans
from kneed import KneeLocator
from sklearn.metrics import silhouette_score
from sklearn.manifold import TSNE
from sklearn import preprocessing
# +
"""
TASK 1-2
"""
# Read csv from current path.
def read_csv():
df1 = pd.read_csv('2020_US_weekly_symptoms_dataset.csv')
df2 = pd.read_csv('aggregated_cc_by.csv')
return df1, df2
df1, df2 = read_csv()
print(df1.shape)
print(df2.shape)
# +
"""
TASK 1-3
"""
# Since we are going to handle regions in USA from 2020_US_weekly_symptoms_dataset.csv
# To extract the records of USA only from dataframe
def extract_from_aggregated(df1, df2):
# Get all region codes
region_codes = df1.open_covid_region_code.unique()
result = df2[df2['open_covid_region_code'].isin(region_codes)]
result = result.reset_index(drop = True)
result = result[['open_covid_region_code', 'region_name', 'date', 'hospitalized_new']]
return result
df2 = extract_from_aggregated(df1,df2)
print(df2.shape)
# +
# Clean data in threshold num_rows * 0.05 and num_cols * 0.05
def clean_dataframe(df):
num_rows,num_cols = df1.shape
thresh_rows = int(num_rows * 0.05);
thresh_cols = int(num_cols * 0.05);
df = df.dropna(axis = 1, thresh = thresh_rows)
df = df.dropna(axis = 0, thresh = thresh_cols)
df = df.reset_index(drop = True)
return df;
df1 = clean_dataframe(df1)
print('after clean: ', df1.shape)
print(df1.head())
# +
def convert_to_datetime(df1, df2):
df1['date'] = pd.to_datetime(df1.date)
df2['date'] = pd.to_datetime(df2.date)
return df1,df2
df1, df2 = convert_to_datetime(df1,df2)
# +
def daily_to_weekly(df2):
df2['date'] = df2['date'] - pd.to_timedelta(7, unit='d')
df2 = df2.groupby(['open_covid_region_code', pd.Grouper(key='date', freq='W-MON')])['hospitalized_new'].sum().reset_index().sort_values(['open_covid_region_code', 'date'])
return df2
print(df2['date'])
df2 = daily_to_weekly(df2)
print(df2['date'])
# +
"""
TASK 1-4
"""
# Merge two data, delete unnecessary columns
def merge_two_dfs(df1, df2):
result = df1.merge(df2, how='inner', on=['open_covid_region_code', 'date'])
result = result.reset_index(drop = True)
result.drop('country_region', axis='columns', inplace=True)
result = result.fillna(0)
return result
final = merge_two_dfs(df1,df2)
print(final.head())
# -
#### TASK 2!!!!!!!!!!
# +
"""
TASK 3
"""
# df: mereged data
# Case1) To keep all data from some regions in the val set and train on the rest
def split_data_regions(df):
regions = df.open_covid_region_code.unique()
col_name = 'open_covid_region_code'
kf = KFold()
for train_index, val_index in kf.split(regions):
train, validation = regions[train_index], regions[val_index]
train_df = df[df[col_name].isin(train)]
val_df = df[df[col_name].isin(validation)]
X_train = train_df.iloc[:, 5:-1]
Y_train = train_df['hospitalized_new']
# print("X_train: \n", X_train.head(3))
# print("Y_train: \n", Y_train.head(3))
X_val = val_df.iloc[:, 5:-1]
Y_val = val_df['hospitalized_new']
# print("X_validation: \n", X_val.head(3))
# print("Y_validation: \n", Y_val.head(3))
# X_train = preprocessing.StandardScaler().fit(X_train).transform(X_train.astype(float))
# X_val = preprocessing.StandardScaler().fit(X_train).transform(X_train.astype(float))
# Y_train = preprocessing.StandardScaler().fit(X_train).transform(X_train.astype(float))
# Y_val = preprocessing.StandardScaler().fit(X_train).transform(X_train.astype(float))
yield X_train, Y_train, X_val, Y_val, train
# Case2) To keep data for the last couple of timepoints (keep data after 2020-08-10) from all regions in the va
# set and train on the rest
# df: merged data // date: date as string
def split_data_time(df, d):
# Convert to datetime from String
date = datetime.strptime(d, '%Y-%m-%d')
train_df = df[df["date"] <= date]
val_df = df[df["date"] > date]
X_train = train_df.iloc[:, 5:-1]
Y_train = train_df['hospitalized_new']
X_val = val_df.iloc[:, 5:-1]
Y_val = val_df['hospitalized_new']
return X_train, Y_train, X_val, Y_val, train_df['date']
# +
"""
TASK 3: KNeighborsRegressor
@Params: k:int, X_train, Y_train, X_val: dataframe
"""
def KNN_regression(k, X_train, Y_train, X_val):
neigh = KNeighborsRegressor(n_neighbors=k)
neigh.fit(X_train, Y_train)
pred = neigh.predict(X_val)
return pred
"""
TASK 3: DecisionTreeRegressor
@Params: l:int, X_train, Y_train, X_val: dataframe
"""
def DecisionTree_regression(l, X_train, Y_train, X_val):
model = DecisionTreeRegressor(min_samples_leaf=l)
model.fit(X_train, Y_train)
pred = model.predict(X_val)
return pred
# +
"""
TASK 3: KNeighborsRegressor based on regions
"""
errors_knn = []
errors_tmp = []
fold_regions = []
for X_train, Y_train, X_val, Y_val, train in split_data_regions(final):
for n in range(100):
pred = KNN_regression(n+1, X_train, Y_train, X_val)
mse = mean_squared_error(Y_val, pred, squared=False)
errors_knn.append(mse)
errors_tmp.append(mse)
fold_regions.append(train)
plt.figure(figsize=(12,9))
plt.plot(range(100), errors_tmp, color='blue', markerfacecolor='red', linestyle='dashed', marker='o', markersize=8)
plt.title('Regions')
plt.ylabel('Error')
plt.xlabel('K')
errors_tmp = []
err = np.array(errors_knn)
min_index = np.argmin(err)
k = (min_index // 5) + 1
min_err = err[min_index]
min_reg = fold_regions[min_index]
print('[BASED ON REGION] K: ', k)
print('[BASED ON REGION] Minimum MSE: ', min_err)
print('[BASED ON REGION] Regions used in Train: \n', min_reg)
# +
"""
TASK 3: KNeighborsRegressor based on times
"""
errors_knn = []
errors_tmp = []
fold_times = []
d = '2020-08-01'
X_train, Y_train, X_val, Y_val, train = split_data_time(final, d)
for n in range(100):
pred = KNN_regression(n+1, X_train, Y_train, X_val)
mse = mean_squared_error(Y_val, pred, squared=False)
errors_knn.append(mse)
errors_tmp.append(mse)
fold_times.append(train)
plt.figure(figsize=(12,9))
plt.plot(range(100), errors_tmp, color='blue', markerfacecolor='red', linestyle='dashed', marker='o', markersize=8)
plt.title('Times')
plt.ylabel('Error')
plt.xlabel('K')
# for X_train, Y_train, X_val, Y_val, train in split_data_time(final, d):
# for n in range(100):
# pred = KNN_regression(n+1, X_train, Y_train, X_val)
# mse = mean_squared_error(Y_val, pred, squared=False)
# errors_knn.append(mse)
# errors_tmp.append(mse)
# fold_times.append(train)
# plt.figure(figsize=(12,9))
# plt.plot(range(100), errors_tmp, color='blue', markerfacecolor='red', linestyle='dashed', marker='o', markersize=8)
# plt.title('Times')
# plt.ylabel('Error')
# plt.xlabel('K')
# errors_tmp = []
err = np.array(errors_knn)
min_index = np.argmin(err)
k = (min_index // 5) + 1
min_err = err[min_index]
min_reg = fold_regions[min_index]
print('[BASED ON TIME] K: ', k)
print('[BASED ON TIME] Minimum MSE: ', min_err)
print('[BASED ON TIME] Dates used in Train: \n', min_reg, '\n[BASED ON TIME] Time: ', d)
# +
"""
TASK 3: DecisionTreeRegressor based on regions
"""
errors_tmp = []
errors_dt = []
fold_regions = []
for X_train, Y_train, X_val, Y_val, train in split_data_regions(final):
for l in range(100):
pred = DecisionTree_regression(l+1, X_train, Y_train, X_val)
mse = mean_squared_error(Y_val, pred, squared=False)
errors_dt.append(mse)
errors_tmp.append(mse)
fold_regions.append(train)
plt.figure(figsize=(12,9))
plt.plot(range(100), errors_tmp, color='blue', markerfacecolor='red', linestyle='dashed', marker='o', markersize=8)
plt.title('Regions')
plt.ylabel('Error')
plt.xlabel('Leaf')
errors_tmp = []
err = np.array(errors_dt)
min_index = np.argmin(err)
l = (min_index // 5) + 1
min_err = err[min_index]
min_reg = fold_regions[min_index]
print('[BASED ON REGION] Leaf: ', l)
print('[BASED ON REGION] Minimum MSE: ', min_err)
print('[BASED ON REGION] Regions used in Train: \n', min_reg)
# +
"""
TASK 3: DecisionTreeRegressor based on times
"""
errors_tmp = []
errors_dt = []
fold_times = []
d = '2020-08-01'
X_train, Y_train, X_val, Y_val, train = split_data_time(final, d)
for l in range(100):
pred = DecisionTree_regression(l+1, X_train, Y_train, X_val)
mse = mean_squared_error(Y_val, pred)
errors_dt.append(mse)
errors_tmp.append(mse)
fold_times.append(train)
plt.figure(figsize=(12,9))
plt.plot(range(100), errors_tmp, color='blue', markerfacecolor='red', linestyle='dashed', marker='o', markersize=8)
plt.title('Regions')
plt.ylabel('Error')
plt.xlabel('Leaf')
errors_tmp = []
err = np.array(errors_dt)
min_index = np.argmin(err)
l = (min_index // 5) + 1
min_err = err[min_index]
min_reg = fold_times[min_index]
print('\n')
print('[BASED ON TIME] L: ', l)
print('[BASED ON TIME] Minimum MSE: ', min_err)
print('[BASED ON TIME] Times used in Train: \n', min_reg, '\n[BASED ON TIME] Time: ', d)
# -
| project1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <div class="alert alert-success">
# <b>PROGRAMMING ASSIGNMENT : image compression using k-means</b>:
#
#
# use the cluster labels as the color
#
#
#
# </div>
# %matplotlib inline
import matplotlib.pyplot as plt
from matplotlib.pyplot import imread
from sklearn import cluster
image = imread("p.png")
plt.figure(figsize = (15,8))
plt.imshow(image)
image.shape
x, y, z = image.shape
image_2d = image.reshape(x*y, z)
image_2d.shape
kmeans_cluster = cluster.KMeans(n_clusters=16)
kmeans_cluster.fit(image_2d)
cluster_centers = kmeans_cluster.cluster_centers_
cluster_labels = kmeans_cluster.labels_
plt.figure(figsize = (15,8))
plt.imshow(cluster_centers[cluster_labels].reshape(x, y, z))
| image_class.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Representing Data with EntitySets
#
# An ``EntitySet`` is a collection of dataframes and the relationships between them. They are useful for preparing raw, structured datasets for feature engineering. While many functions in Featuretools take ``dataframes`` and ``relationships`` as separate arguments, it is recommended to create an ``EntitySet``, so you can more easily manipulate your data as needed.
#
# ## The Raw Data
#
# Below we have two tables of data (represented as Pandas DataFrames) related to customer transactions. The first is a merge of transactions, sessions, and customers so that the result looks like something you might see in a log file:
# +
import featuretools as ft
data = ft.demo.load_mock_customer()
transactions_df = data["transactions"].merge(data["sessions"]).merge(data["customers"])
transactions_df.sample(10)
# -
# And the second dataframe is a list of products involved in those transactions.
products_df = data["products"]
products_df
# ## Creating an EntitySet
#
# First, we initialize an ``EntitySet``. If you'd like to give it a name, you can optionally provide an ``id`` to the constructor.
es = ft.EntitySet(id="customer_data")
# ## Adding dataframes
#
# To get started, we add the transactions dataframe to the `EntitySet`. In the call to ``add_dataframe``, we specify three important parameters:
#
# * The ``index`` parameter specifies the column that uniquely identifies rows in the dataframe.
# * The ``time_index`` parameter tells Featuretools when the data was created.
# * The ``logical_types`` parameter indicates that "product_id" should be interpreted as a Categorical column, even though it is just an integer in the underlying data.
# +
from woodwork.logical_types import Categorical, PostalCode
es = es.add_dataframe(
dataframe_name="transactions",
dataframe=transactions_df,
index="transaction_id",
time_index="transaction_time",
logical_types={
"product_id": Categorical,
"zip_code": PostalCode,
},
)
es
# -
# You can also use a setter on the ``EntitySet`` object to add dataframes
# + raw_mimetype="text/restructuredtext" active=""
# .. currentmodule:: featuretools
#
#
# .. note ::
#
# You can also use a setter on the ``EntitySet`` object to add dataframes
#
# ``es["transactions"] = transactions_df``
#
# that this will use the default implementation of `add_dataframe`, notably the following:
#
# * if the DataFrame does not have `Woodwork <https://woodwork.alteryx.com/>`_ initialized, the first column will be the index column
# * if the DataFrame does not have Woodwork initialized, all columns will be inferred by Woodwork.
# * if control over the time index column and logical types is needed, Woodwork should be initialized before adding the dataframe.
#
# .. note ::
#
# You can also display your `EntitySet` structure graphically by calling :meth:`.EntitySet.plot`.
# -
# This method associates each column in the dataframe to a [Woodwork](https://woodwork.alteryx.com/) logical type. Each logical type can have an associated standard semantic tag that helps define the column data type. If you don't specify the logical type for a column, it gets inferred based on the underlying data. The logical types and semantic tags are listed in the schema of the dataframe. For more information on working with logical types and semantic tags, take a look at the [Woodwork documention](https://woodwork.alteryx.com/).
es["transactions"].ww.schema
# Now, we can do that same thing with our products dataframe.
# +
es = es.add_dataframe(
dataframe_name="products",
dataframe=products_df,
index="product_id")
es
# -
# With two dataframes in our `EntitySet`, we can add a relationship between them.
#
# ## Adding a Relationship
#
# We want to relate these two dataframes by the columns called "product_id" in each dataframe. Each product has multiple transactions associated with it, so it is called the **parent dataframe**, while the transactions dataframe is known as the **child dataframe**. When specifying relationships, we need four parameters: the parent dataframe name, the parent column name, the child dataframe name, and the child column name. Note that each relationship must denote a one-to-many relationship rather than a relationship which is one-to-one or many-to-many.
es = es.add_relationship("products", "product_id", "transactions", "product_id")
es
# Now, we see the relationship has been added to our `EntitySet`.
#
# ## Creating a dataframe from an existing table
#
# When working with raw data, it is common to have sufficient information to justify the creation of new dataframes. In order to create a new dataframe and relationship for sessions, we "normalize" the transaction dataframe.
es = es.normalize_dataframe(
base_dataframe_name="transactions",
new_dataframe_name="sessions",
index="session_id",
make_time_index="session_start",
additional_columns=[
"device",
"customer_id",
"zip_code",
"session_start",
"join_date",
],
)
es
# Looking at the output above, we see this method did two operations:
#
# 1. It created a new dataframe called "sessions" based on the "session_id" and "session_start" columns in "transactions"
# 2. It added a relationship connecting "transactions" and "sessions"
#
# If we look at the schema from the transactions dataframe and the new sessions dataframe, we see two more operations that were performed automatically:
es["transactions"].ww.schema
es["sessions"].ww.schema
# 1. It removed "device", "customer_id", "zip_code" and "join_date" from "transactions" and created a new columns in the sessions dataframe. This reduces redundant information as the those properties of a session don't change between transactions.
# 2. It copied and marked "session_start" as a time index column into the new sessions dataframe to indicate the beginning of a session. If the base dataframe has a time index and ``make_time_index`` is not set, ``normalize_dataframe`` will create a time index for the new dataframe. In this case it would create a new time index called "first_transactions_time" using the time of the first transaction of each session. If we don't want this time index to be created, we can set ``make_time_index=False``.
#
# If we look at the dataframes, we can see what ``normalize_dataframe`` did to the actual data.
es["sessions"].head(5)
es["transactions"].head(5)
# To finish preparing this dataset, create a "customers" dataframe using the same method call.
# +
es = es.normalize_dataframe(
base_dataframe_name="sessions",
new_dataframe_name="customers",
index="customer_id",
make_time_index="join_date",
additional_columns=["zip_code", "join_date"],
)
es
# -
# ## Using the EntitySet
#
# Finally, we are ready to use this EntitySet with any functionality within Featuretools. For example, let's build a feature matrix for each product in our dataset.
# +
feature_matrix, feature_defs = ft.dfs(entityset=es, target_dataframe_name="products")
feature_matrix
# + raw_mimetype="text/restructuredtext" active=""
# As we can see, the features from DFS use the relational structure of our `EntitySet`. Therefore it is important to think carefully about the dataframes that we create.
#
# Dask and Koalas EntitySets
# ~~~~~~~~~~~~~~~~~~~~~~~~~~
#
# EntitySets can also be created using Dask dataframes or Koalas dataframes. For more information refer to :doc:`../guides/using_dask_entitysets` and :doc:`../guides/using_koalas_entitysets`.
| docs/source/getting_started/using_entitysets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
from __future__ import print_function
import cobra
import cobra.test
import mackinac
import numpy as np
import csv
import glob
import pickle
import pandas as pd
import time
import sys
from collections import defaultdict
from cobra.flux_analysis import gapfill
# -
# Read in list of all genomes on PATRIC and list of existing models in folder
with open('../Data/20_species_1023_genomes.csv') as csvfile:
genome_ids_list = []
for line in csvfile:
genome_ids_list.append(line.strip())
len(genome_ids_list)
# models = glob.glob('../gap_models/*.xml')
# models = [x.replace("../gap_models/","").replace(".xml","") for x in models]
# +
# Do any ungapfilled models produce lactate?
# Lactate = cpd01022
# D-lactate = cpd00221
# L-lactate = cpd00159
lactate = []
dlactate = []
llactate = []
missing_lactate = []
missing_dlactate = []
missing_llactate = []
for genome_id in genome_ids_list:
model_file_name = "../gap_models/%s.xml" % (genome_id)
model = cobra.io.read_sbml_model(model_file_name)
# Lactate = cpd01022
try:
metabolite = model.metabolites.get_by_id('cpd01022_c')
demand = model.add_boundary(metabolite, type='demand')
model.objective = demand
obj_val = model.slim_optimize(error_value=0.)
if obj_val > 1e-3:
lactate.append([genome_id, obj_val])
model.remove_reactions([demand])
except:
missing_lactate.append(genome_id)
pass
# D-lactate = cpd00221
try:
metabolite = model.metabolites.get_by_id('cpd00221_c')
demand = model.add_boundary(metabolite, type='demand')
model.objective = demand
obj_val = model.slim_optimize(error_value=0.)
if obj_val > 1e-3:
dlactate.append([genome_id, obj_val])
model.remove_reactions([demand])
except:
missing_dlactate.append(genome_id)
pass
# L-lactate = cpd00159
try:
metabolite = model.metabolites.get_by_id('cpd00159_c')
demand = model.add_boundary(metabolite, type='demand')
model.objective = demand
obj_val = model.slim_optimize(error_value=0.)
if obj_val > 1e-3:
llactate.append([genome_id, obj_val])
model.remove_reactions([demand])
except:
missing_llactate.append(genome_id)
pass
print(lactate)
print(dlactate)
print(llactate)
print(missing_lactate)
print(missing_dlactate)
print(missing_llactate)
# +
# Do any gapfilled models produce lactate?
# Lactate = cpd01022
# D-lactate = cpd00221
# L-lactate = cpd00159
lactate2 = []
dlactate2 = []
llactate2 = []
missing_lactate2 = []
missing_dlactate2 = []
missing_llactate2 = []
for genome_id in genome_ids_list:
model_file_name = "../models/%s.xml" % (genome_id)
model = cobra.io.read_sbml_model(model_file_name)
# Lactate = cpd01022
try:
metabolite = model.metabolites.get_by_id('cpd01022_c')
demand = model.add_boundary(metabolite, type='demand')
model.objective = demand
obj_val = model.slim_optimize(error_value=0.)
if obj_val > 1e-3:
lactate2.append([genome_id, obj_val])
model.remove_reactions([demand])
except:
missing_lactate2.append(genome_id)
pass
# D-lactate = cpd00221
try:
metabolite = model.metabolites.get_by_id('cpd00221_c')
demand = model.add_boundary(metabolite, type='demand')
model.objective = demand
obj_val = model.slim_optimize(error_value=0.)
if obj_val > 1e-3:
dlactate2.append([genome_id, obj_val])
model.remove_reactions([demand])
except:
missing_dlactate2.append(genome_id)
pass
# L-lactate = cpd00159
try:
metabolite = model.metabolites.get_by_id('cpd00159_c')
demand = model.add_boundary(metabolite, type='demand')
model.objective = demand
obj_val = model.slim_optimize(error_value=0.)
if obj_val > 1e-3:
llactate2.append([genome_id, obj_val])
model.remove_reactions([demand])
except:
missing_llactate2.append(genome_id)
pass
print(lactate2)
print(dlactate2)
print(llactate2)
print(missing_lactate2)
print(missing_dlactate2)
print(missing_llactate2)
# +
# Do any gap models produce pyruvate?
# pyruvate = cpd00020
pyruvate = []
for genome_id in genome_ids_list[10:20]:
model_file_name = "../gap_models/%s.xml" % (genome_id)
model = cobra.io.read_sbml_model(model_file_name)
metabolite = model.metabolites.get_by_id('cpd00020_c')
demand = model.add_boundary(metabolite, type='demand')
model.objective = demand
obj_val = model.slim_optimize(error_value=0.)
# print(obj_val)
if obj_val > 1e-3:
pyruvate.append([genome_id, obj_val])
model.remove_reactions([demand])
print(pyruvate)
# -
bio1_mets = model.reactions.get_by_id('bio1').metabolites.keys()
bio1_met_names_ids = []
for bio1_met in bio1_mets:
bio1_met_names_ids.append([bio1_met.name, bio1_met.id])
bio1_met_names_ids
len(bio1_met_names_ids)
# +
t = time.time()
counter = 0
media_set = set()
for genome_id in genome_ids_list:
sys.stdout.write('\r'+ str(counter) + ': ' + str(len(media_set)))
sys.stdout.flush()
model_file_name = "../gap_models/%s.xml" % (genome_id)
model = cobra.io.read_sbml_model(model_file_name)
media_EXs = model.medium.keys()
media_set |= set(media_EXs)
counter += 1
# media_set
# +
media_list = list(media_set)
media_list = [x.replace("EX_","").replace("_e","_c") for x in media_list]
ex_dict = defaultdict(list)
for cpd in media_list:
cpd_obj = universal.metabolites.get_by_id(cpd)
ex_dict[cpd_obj.name].append(cpd_obj.id)
ex_dict
# -
universal = cobra.io.load_json_model("../Data/GramPosUni.json")
comp_media = model.medium.keys()
media_mets = []
for exchange_id in comp_media:
exchange = model.reactions.get_by_id(exchange_id).reactants
met = exchange[0].id.replace("_e","_c")
name = universal.metabolites.get_by_id(met).name
media_mets.append([name,met])
media_mets
universal = cobra.io.load_json_model("../Data/GramPosUni.json")
comp_media = model.medium.keys()
media_mets = []
for exchange_id in comp_media:
exchange = model.reactions.get_by_id(exchange_id).reactants
met = exchange[0].id.replace("_e","_c")
name = universal.metabolites.get_by_id(met).name
media_mets.append([name,met])
media_mets
aa_ex_remove = ['EX_cpd11590_e',
'EX_cpd00065_e', 'EX_cpd11592_e',
'EX_cpd11588_e', 'EX_cpd11580_e',
'EX_cpd00117_e', 'EX_cpd11584_e',
'EX_cpd11582_e', 'EX_cpd11586_e',
'EX_cpd00156_e', 'EX_cpd00033_e',
'EX_cpd11591_e', 'EX_cpd00550_e',
'EX_cpd11583_e', 'EX_cpd11581_e',
'EX_cpd11587_e', 'EX_cpd11585_e',
'EX_cpd01914_e', 'EX_cpd00637_e',
'EX_cpd01017_e', 'EX_cpd00039_e',
'EX_cpd00122_e', 'EX_cpd15603_e',
'EX_cpd11593_e', 'EX_cpd15605_e',
'EX_cpd00051_e', 'EX_cpd00064_e',
'EX_cpd00322_e', 'EX_cpd11576_e',
'EX_cpd00060_e', 'EX_cpd11589_e',
'EX_cpd00054_e', 'EX_cpd00053_e',
'EX_cpd15606_e', 'EX_cpd15604_e']
aas =[
['D-lactate','cpd00221_c'],
['L-lactate','cpd00159_c'],
['D-Alanine', 'cpd00117_c'],
['L-Arginine', 'cpd00051_c'],
['D-Aspartate', 'cpd00320_c'],
['L-Aspartate', 'cpd00041_c'],
['D-Cysteine', 'cpd00587_c'],
['L-Cysteine', 'cpd00084_c'],
['D-Glutamine', 'cpd00610_c'],
['L-Glutamine', 'cpd00053_c'],
['Glycine', 'cpd00033_c'],
['Histidine','cpd00572_c'],
['D-Histidine','cpd03842_c'],
['L-Histidine','cpd00119_c'],
['L-Hydroxyproline','cpd00747_c'],
['D-Isoleucine', 'cpd03841_c'],
['L-Isoleucine', 'cpd00322_c'],
['D-Leucine','cpd01106_c'],
['L-Leucine','cpd00107_c'],
['D-Lysine', 'cpd00549_c'],
['L-Lysine', 'cpd00039_c'],
['D-Methionine', 'cpd00637_c'],
['L-Methionine', 'cpd00060_c'],
['Phenylalanine','cpd01400_c'],
['D-Phenylalanine','cpd01526_c'],
['L-Phenylalanine','cpd00066_c'],
['D-Proline', 'cpd00567_c'],
['L-Proline', 'cpd00129_c'],
['D-Serine', 'cpd00550_c'],
['L-Serine', 'cpd00054_c'],
['D-Threonine', 'cpd00611_c'],
['L-Threonine', 'cpd00161_c'],
['D-Tryptophan', 'cpd00411_c'],
['L-Tryptophan', 'cpd00065_c'],
['D-Tyrosine', 'cpd03843_c'],
['L-Tyrosine', 'cpd00069_c'],
['Valine', 'cpd15141_c'],
['D-Valine', 'cpd03840_c'],
['L-Valine', 'cpd00156_c'],
['Ornithine', 'cpd00064_c'],
['D-Ornithine', 'cpd00404_c']
]
# +
t = time.time()
total_genomes = len(genome_ids_list)
aa_prod = defaultdict(list)
counter = 0
failure_report = []
for genome_id in genome_ids_list:
sys.stdout.write('\r'+ str(counter))
sys.stdout.flush()
model_file_name = "../gap_models/%s.xml" % (genome_id)
model = cobra.io.read_sbml_model(model_file_name)
model_rxns = []
for rxn in model.reactions:
model_rxns.append(rxn.id)
model_mets = []
for met in model.metabolites:
model_mets.append(met.id)
for ex_rxn_id in aa_ex_remove:
if ex_rxn_id in model_rxns:
model.reactions.get_by_id(ex_rxn_id).lower_bound = 0
model.reactions.get_by_id(ex_rxn_id).upper_bound = 0
for met_id in aas:
if met_id[1] in model_mets:
try:
metabolite = model.metabolites.get_by_id(met_id[1])
demand = model.add_boundary(metabolite, type='demand')
model.objective = demand
obj_val = model.slim_optimize(error_value=0.)
if obj_val > 1e-3:
model.remove_reactions([demand])
aa_prod[met_id[0]].append(genome_id)
else:
model.remove_reactions([demand])
except:
failure_report.append([genome_id,met_id[1]])
pass
counter += 1
percent_dict = {}
for aa in aas:
percent_dict[aa[0]] = 100*len(aa_prod[aa[0]])/total_genomes
elapsed = time.time() - t
print("Time to complete:", elapsed/60, "mins")
# -
percent_dict
# +
# 'L-Aspartate', 'cpd00041_c'
float(len(aa_prod['L-Aspartate']))/1023
missing_asp = set(genome_ids_list).difference(set(aa_prod['L-Aspartate']))
for genome_id in missing_asp:
print(genome_id)
# +
# t = time.time()
total_genomes = len(genome_ids_list)
# aa_prod = defaultdict(list)
asp_obj_val = []
counter = 0
# failure_report = []
# for genome_id in genome_ids_list:
# sys.stdout.write('\r'+ str(counter))
# sys.stdout.flush()
for genome_id in missing_asp:
model_file_name = "../gap_models/%s.xml" % (genome_id)
model = cobra.io.read_sbml_model(model_file_name)
model_rxns = []
for rxn in model.reactions:
model_rxns.append(rxn.id)
model_mets = []
for met in model.metabolites:
model_mets.append(met.id)
for ex_rxn_id in aa_ex_remove:
if ex_rxn_id in model_rxns:
model.reactions.get_by_id(ex_rxn_id).lower_bound = 0
model.reactions.get_by_id(ex_rxn_id).upper_bound = 0
metabolite = model.metabolites.get_by_id('cpd00041_c')
demand = model.add_boundary(metabolite, type='demand')
model.objective = demand
obj_val = model.slim_optimize(error_value=0.)
if obj_val > 1e-3:
model.remove_reactions([demand])
asp_obj_val.append(genome_id)
else:
model.remove_reactions([demand])
# counter += 1
# percent_dict = {}
# for aa in aas:
# percent_dict[aa[0]] = 100*len(aa_prod[aa[0]])/total_genomes
# elapsed = time.time() - t
# print("Time to complete:", elapsed/60, "mins")
# +
# model.solver = 'gurobi'
# +
t = time.time()
universal = cobra.io.load_json_model("../Data/GramPosUni.json")
genome_id = '1050107.3'
model_file_name = "../gap_models/%s.xml" % (genome_id)
model = cobra.io.read_sbml_model(model_file_name)
model_rxns = []
for rxn in model.reactions:
model_rxns.append(rxn.id)
model_mets = []
for met in model.metabolites:
model_mets.append(met.id)
for ex_rxn_id in aa_ex_remove:
if ex_rxn_id in model_rxns:
model.reactions.get_by_id(ex_rxn_id).lower_bound = 0
model.reactions.get_by_id(ex_rxn_id).upper_bound = 0
metabolite = model.metabolites.get_by_id('cpd00041_c')
demand = model.add_boundary(metabolite, type='demand')
model.objective = demand
obj_val = model.slim_optimize(error_value=0.)
if obj_val > 1e-3:
model.remove_reactions([demand])
print('model_has_pathway')
else:
solution = gapfill(model, universal, demand_reactions=False)
model.remove_reactions([demand])
elapsed = time.time() - t
print("Time to complete:", elapsed/60, "mins")
solution
# +
genome_id = '1050107.3'
model_file_name = "../gap_models/%s.xml" % (genome_id)
model = cobra.io.read_sbml_model(model_file_name)
rxn_to_add = universal.reactions.get_by_id('rxn00249_c')
model.add_reactions([rxn_to_add])
# model.reactions.get_by_id('rxn00249_c')
model_rxns = []
for rxn in model.reactions:
model_rxns.append(rxn.id)
model_mets = []
for met in model.metabolites:
model_mets.append(met.id)
for ex_rxn_id in aa_ex_remove:
if ex_rxn_id in model_rxns:
model.reactions.get_by_id(ex_rxn_id).lower_bound = 0
model.reactions.get_by_id(ex_rxn_id).upper_bound = 0
metabolite = model.metabolites.get_by_id('cpd00041_c')
demand = model.add_boundary(metabolite, type='demand')
model.objective = demand
solution = model.optimize()
# -
df = solution.fluxes.to_frame()
df_active = df.loc[(abs(df['fluxes'])) > 0.1]
df_active
active_fva_solution = fva_solution.loc[(abs(fva_solution['minimum'] - fva_solution['maximum']) < 200)
& (abs(fva_solution['minimum'] * fva_solution['maximum']) > 1.5)]
active_fva_solution
# +
# solution.shadow_prices
# -
model.summary()
from cobra.flux_analysis.summary import metabolite_summary
metabolite_summary(model.metabolites.get_by_id('cpd00041_c'))
metabolite_summary(model.metabolites.get_by_id('cpd00106_c'))
metabolite_summary(model.metabolites.get_by_id('cpd00130_c'))
metabolite_summary(model.metabolites.get_by_id('cpd00032_c'))
metabolite_summary(model.metabolites.get_by_id('cpd00061_c'))
metabolite_summary(model.metabolites.get_by_id('cpd00482_c'))
metabolite_summary(model.metabolites.get_by_id('cpd00223_c'))
metabolite_summary(model.metabolites.get_by_id('cpd00482_c'))
metabolite_summary(model.metabolites.get_by_id('cpd00020_c'))
metabolite_summary(model.metabolites.get_by_id('cpd00276_e'))
model.reactions.get_by_id('rxn00347_c')
from cobra.flux_analysis import gapfill
universal.reactions.get_by_id('rxn00109_c').check_mass_balance()
# +
# sink = model.add_boundary(metabolite, type='sink')
# sink
# -
for rxn in model.metabolites.get_by_id('cpd00032_c').reactions:
print(rxn.id)
print('REACTANTS')
for react in rxn.reactants:
print(react.name)
print('PRODUCTS')
for prod in rxn.products:
print(prod.name)
model.reactions.get_by_id('rxn00260')
# +
# Flux sampling
from cobra.flux_analysis.sampling import OptGPSampler
optgp = OptGPSampler(cd630_PATRIC, 6)
print('\nHomogeneous: ' + str(optgp.problem.homogeneous))
s = optgp.sample(10000)
s.std().describe()
# -
active_fva_solution = fva_solution.loc[(abs(fva_solution['minimum'] - fva_solution['maximum']) < 200)
& (abs(fva_solution['minimum'] * fva_solution['maximum']) > 1.5)]
active_fva_solution
fva_solution.loc['EX_cpd00013_e']
model.reactions.get_by_id('DM_cpd00041_c')
# +
t = time.time()
from cobra.flux_analysis import flux_variability_analysis
fva_solution = flux_variability_analysis(model, model.reactions, loopless=True)
# list(fva_solution.index)
# type(fva_solution['minimum'])
active_fva_solution = fva_solution.loc[(fva_solution['minimum'] > -9999.9)
& (fva_solution['maximum'] < 9999.9)
& (fva_solution['minimum'] < -0.1)
& (fva_solution['maximum'] > 0.1)]
elapsed = time.time() - t
print("Time to complete:", elapsed/60, "mins")
active_fva_solution
# -
active_fva_solution = fva_solution.loc[(fva_solution['minimum'] > -999)
& (fva_solution['maximum'] < 999)
& (fva_solution['minimum'] < -1)
& (fva_solution['maximum'] > 1)]
active_fva_solution
active_fva_solution = fva_solution.loc[(((fva_solution['minimum'] > -999) & (fva_solution['minimum'] < -1))|
((fva_solution['minimum'] < 999) & (fva_solution['minimum'] > 1)))
& (((fva_solution['maximum'] > -999) & (fva_solution['maximum'] < -1))|
((fva_solution['maximum'] < 999) & (fva_solution['maximum'] > 1)))]
active_fva_solution
# +
likelihoods = pickle.load(open('../likelihoods/1050107.3.probs'))
likelihoods['rxn00260_c']
# need to test with probannopy gapfill function on cluster...
# +
t = time.time()
model = cobra.io.read_sbml_model('../gap_models/1051650.8.xml')
with model as model_x:
gaps = findGapFilled(model_x)
for rxn in gaps:
model_x.remove_reactions([rxn])
# print(model_x.optimize().objective_value)
solution = gapfill(model, universal, demand_reactions=False)
for reaction in solution[0]:
print(reaction.id)
elapsed = time.time() - t
print "Time to complete:", elapsed/60, "mins"
# +
# Basal Synthetic Media
bsm = [
['H+','cpd00067'],
['H2O','cpd00001'],
['CO2','cpd00011'],
['O2','cpd00007'],
['N2','cpd00528'],
# ['H2','cpd11640'], # Only with no O2
['K+','cpd00205'],
['Na+','cpd00971'],
['Mg','cpd00254'],
['Mn2+','cpd00030'],
['Fe2+','cpd10515'], # Iron ion in heme
['Ca2+','cpd00063'], # Calcium pantothenate;cpd19112
['Vitamin B12r','cpd00423'], # C62H91CoN13O14P : cobalamin;cpd03424;cpd00730 : not present in any exchange reactions
['Cobinamide','cpd03422'], #EXs : related to cobalamin (B12) Added to ensure cells have access to B12
['BIOT','cpd00104'], # C10H15N2O3S : biotin B7
['PAN','cpd00644'], # C9H16NO5 : Pantothenate B5
['Folate','cpd00393'], # C19H17N7O6 : B9
['Niacin','cpd00218'], # C6H4NO2 : B3
['Pyridoxal','cpd00215'], # C8H9NO3 : B6
['Riboflavin','cpd00220'], # C17H19N4O6 : B2
['thiamin','cpd00305'] # C12H17N4OS : B1
['Phosphate','cpd00009'], # HO4P :
['Thioglycolate','cpd01415'], # C2H3O2S : not present in any exchange reactions
['Sulfate','cpd00048'], # O4S :
['Acetate','cpd00029'], # C2H3O2 : not present in any exchange reactions
['Citrate','cpd00137'], # C6H5O7 : Consider removing.
['Polysorbate 60','cpd24450'], # C35H68O10 : Almost tween 80 : not present in any exchange reactions
['Ethyl acetate','cpd00633'], # C4H8O2 : not present in any exchange reactions
['ABEE','cpd00443'], # C7H6NO2 : aminobenzoate : not present in any exchange reactions
]
# Potentially add to BSM (from M9 media)
M9_ions = [
['Cl-','cpd00099'],
['Co2+','cpd00149'],
['Cu2+','cpd00058'],
['Fe3','cpd10516'],
# ['Sodium molybdate','cpd11145'], # This doesn't connect to anything
['Ni2+','cpd00244'],
['Selenate','cpd03396'],
['Selenite','cpd03387'],
['Zn2+','cpd00034']
]
# Enviromental Metabolites with Exchange reactions
[
# ['CO2','cpd00011'], #EXs :
# ['Ca2+','cpd00063'], #EXs :
# ['Cd2+','cpd01012'], #EXs : Removed because toxic
# ['chromate','cpd11595'], #EXs : Removed because toxic
# ['Cl-','cpd00099'], #EXs :
# ['Co2+','cpd00149'], #EXs : In M9
# ['Cu2+','cpd00058'], #EXs : In M9
# ['Fe2+','cpd10515'], #EXs :
# ['H+','cpd00067'], #EXs :
# ['H2','cpd11640'], #EXs :
# ['H2O','cpd00001'], #EXs :
# ['Hg2+','cpd00531'], #EXs : Removed because toxic
# ['K+','cpd00205'], #EXs :
# ['Mg','cpd00254'], #EXs :
# ['Mn2+','cpd00030'], #EXs :
# ['Na+','cpd00971'], #EXs :
# ['Ni2+','cpd00244'], #EXs : In M9
# ['O2','cpd00007'], #EXs :
# ['Pb','cpd04097'], #EXs : Removed because toxic
# ['Zn2+','cpd00034'], #EXs : In M9
# ['fe3','cpd10516'] #EXs : In M9
]
# M9 Base : https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4932939/
# [
# ['Ca2+','cpd00063'],
# ['Cl-','cpd00099'],
# ['CO2','cpd00011'],
# ['Co2+','cpd00149'],
# ['Cu2+','cpd00058'],
# ['Fe2+','cpd10515'],
# ['Fe3','cpd10516'],
# ['H+','cpd00067'],
# ['H2O','cpd00001'],
# ['K+','cpd00205'],
# ['Mg','cpd00254'],
# ['Mn2+','cpd00030'],
# ['Sodium molybdate','cpd11145'],
# ['Na+','cpd00971'],
# ['Ni2+','cpd00244'],
# ['Selenate','cpd03396'],
# ['Selenite','cpd03387'],
# ['Zn2+','cpd00034']
# ]
# M9 default carbon, nitrogen, phosphorous, and sulfur sources
M9_sources = [
['D-Glucose','cpd00027'],
['NH3','cpd00013'], # this is actually NH4 : ammonium
['Phosphate','cpd00009'],
['Sulfate','cpd00048']
]
# Vitamins
vit_k = [
# ['BIOT','cpd00104'], #EXs : Biotin
# ['Cobinamide','cpd03422'], #EXs : related to cobalamin (B12)
# ['Folate','cpd00393'], #EXs :
['Menaquinone 7','cpd11606'], #EXs : Vitamine K2 : Add when there is no O2
# ['Niacin','cpd00218'], #EXs :
# ['PAN','cpd00644'], #EXs : Pantothenate
# ['Pyridoxal','cpd00215'], #EXs :
# ['Riboflavin','cpd00220'], #EXs :
# ['Thiamin','cpd00305'] #EXs :
]
# For aerobic simulations, O2 was added with a lower bound of −20 and to 0 for anaerobic simulations.
# +
# Carbon Sources from all Exchanges and additional interesting sources
[
['4-Hydroxybenzoate','cpd00136'], #EXs : found in coconuts
['2-keto-3-deoxygluconate','cpd00176'], #EXs : degraded pectin product
['Amylotriose','cpd01262'], #EXs :
['CELB','cpd00158'], #EXs : Cellobiose
['D-Fructose','cpd00082'], #EXs :
['D-Glucose','cpd00027'], #EXs :
['D-Mannitol','cpd00314'], #EXs : sweetener the is poorly absorbed in the gut
['D-Mannose','cpd00138'], #EXs : related to mucin
['Ribose','cpd00105'], #EXs :
['Dextrin','cpd11594'], #EXs :
['Dulcose','cpd01171'], #EXs : Galactitol
['GLCN','cpd00222'], #EXs : Gluconate
['GLUM','cpd00276'], #EXs : Glucosamine
['Galactose','cpd00108'], #EXs :
['L-Arabinose','cpd00224'], #EXs :
['L-Inositol','cpd00121'], #EXs :
['L-Lactate','cpd00159'], #EXs :
['L-Malate','cpd00130'], #EXs :
['Glycerol','cpd00100'], #EXs :
['LACT','cpd00208'], #EXs : lactose
['Maltohexaose','cpd01329'], #EXs :
['Maltose','cpd00179'], #EXs :
['Melibiose','cpd03198'], #EXs :
['Palmitate','cpd00214'], #EXs :
['Propionate','cpd00141'], #EXs :
['Salicin','cpd01030'], #EXs :
['Sorbitol','cpd00588'], #EXs :
['Stachyose','cpd01133'], #EXs :
['Succinate','cpd00036'], #EXs :
['Sucrose','cpd00076'], #EXs :
['TRHL','cpd00794'], #EXs : Trehalose
['Ursin','cpd03696'], #EXs : Arbutin
['Xylose','cpd00154'], #EXs :
['hexadecenoate','cpd15237'] #EXs :
]
# Nitrogen Sources
[
# ['NH3','cpd00013'], #EXs :
['Allantoin','cpd01092'], #EXs : degradation product of purines
['BET','cpd00540'], #EXs : Betaine
['Choline','cpd00098'], #EXs : Found in milk
['GABA','cpd00281'], #EXs : Could also be a carbon source
['Nitrate','cpd00209'], #EXs :
['Nitrite','cpd00075'], #EXs :
['Spermidine','cpd00264'], #EXs :
['Urea','cpd00073'], #EXs :
['crotonobetaine','cpd08305'] #EXs :
]
# Sulfur Sources
[
['H2S2O3','cpd00268'], #EXs : Thiosulfate
['Isethionate','cpd03048'], #EXs : C2H5O4S
# ['Sulfate','cpd00048'], #EXs : O4S
['Sulfite','cpd00081'], #EXs : HO3S
['Sulfoacetate','cpd09878'], #EXs : C2H2O5S
['ethanesulfonate','cpd11579'], #EXs : C2H5O3S
['methanesulfonate','cpd08023'] #EXs : CH3O3S
]
# Phosphorus Sources
[
['Phosphate','cpd00009'] #EX :
]
# +
# Amino Acid related metabolites
aas = [
['D-Alanine','cpd00117'], #EXs :
['D-Glutamate','cpd00186'], #EXs :
['D-Methionine','cpd00637'], #EXs :
['D-Serine','cpd00550'], #EXs :
['Glycine','cpd00033'], #EXs : 1
['L-Alanine','cpd00035'], #EXs : 2
['L-Arginine','cpd00051'], #EXs : 3
['L-Asparagine','cpd00132'], #EXs : 4
['L-Aspartate','cpd00041'], #EXs : 5
['L-Cysteate','cpd00395'], #EXs : 6
['L-Cysteine','cpd00084'], #EXs : 7
['L-Glutamate','cpd00023'], #EXs : 8
['L-Glutamine','cpd00053'], #EXs : 9
['L-Histidine','cpd00119'], #EXs : 10
['L-Isoleucine','cpd00322'], #EXs : 11
['L-Leucine','cpd00107'], #EXs : 12
['L-Lysine','cpd00039'], #EXs : 13
['L-Methionine','cpd00060'], #EXs : 14
['L-Phenylalanine','cpd00066'], #EXs : 15
['L-Proline','cpd00129'], #EXs : 16
['L-Serine','cpd00054'], #EXs : 17
['L-Threonine','cpd00161'], #EXs : 18
['L-Tryptophan','cpd00065'], #EXs : 19
['L-Tyrosine','cpd00069'], #EXs : 20
['L-Valine','cpd00156'], #EXs : 21
]
# Explore leave one out with core amino acids.
# Dimers, and other amino acid related mets
aa_related = [
['2-Oxoglutarate','cpd00024'], #EXs :
['Ala-Gln','cpd11587'], #EXs :
['Ala-His','cpd11584'], #EXs :
['Ala-Leu','cpd11583'], #EXs :
['ala-L-asp-L','cpd11593'], #EXs :
['ala-L-glu-L','cpd11586'], #EXs :
['ala-L-Thr-L','cpd11582'], #EXs :
['Aminoethanol','cpd00162'], #EXs : Ethanolamine
['Carnitine','cpd00266'], #EXs :
['Chorismate','cpd00216'], #EXs :
['Cys-Gly','cpd01017'], #EXs :
['Gly-Cys','cpd15603'], #EXs :
['Gly-Gln','cpd11580'], #EXs :
['Gly-Leu','cpd15604'], #EXs :
['Gly-Met','cpd11591'], #EXs :
['Gly-Phe','cpd15605'], #EXs :
['Gly-Tyr','cpd15606'], #EXs :
['gly-asn-L','cpd11581'], #EXs :
['gly-asp-L','cpd11589'], #EXs :
['gly-glu-L','cpd11592'], #EXs :
['gly-pro-L','cpd11588'], #EXs :
['L-Methionine S-oxide','cpd01914'], #EXs :
['L-alanylglycine','cpd11585'], #EXs :
['L-methionine R-oxide','cpd11576'], #EXs :
['met-L-ala-L','cpd11590'], #EXs :
['S-Adenosyl-L-methionine','cpd00017'], #EXs :
['S-Methyl-L-methionine','cpd02027'], #EXs :
['S-Ribosylhomocysteine','cpd02227'], #EXs :
['N-Acetyl-D-glucosamine','cpd00122'], #EXs :
['N-Acetyl-D-mannosamine','cpd00492'], #EXs :
['Ornithine','cpd00064'], #EXs :
['Putrescine','cpd00118'], #EXs :
['Taurine','cpd00210'], #EXs :
['meso-2,6-Diaminopimelate','cpd00516'] #EXs : related to lysine
]
# +
# DNA/RNA related metabolites
[
['35ccmp','cpd00696'], #EXs :
['AMP','cpd00018'], #EXs :
['Adenosine','cpd00182'], #EXs :
['Adenosine 3-5-bisphosphate','cpd00045'], #EXs :
['Cytosine','cpd00307'], #EXs :
['Deoxyadenosine','cpd00438'], #EXs :
['Deoxycytidine','cpd00654'], #EXs :
['Deoxyguanosine','cpd00277'], #EXs :
['Deoxyinosine','cpd03279'], #EXs :
['Deoxyuridine','cpd00412'], #EXs :
['GMP','cpd00126'], #EXs :
['GTP','cpd00038'], #EXs :
['Guanosine','cpd00311'], #EXs :
['Inosine','cpd00246'], #EXs :
['HYXN','cpd00226'], #EXs : Hypoxanthine
['Nicotinamide ribonucleotide','cpd00355'], #EXs :
['TTP','cpd00357'], #EXs : Deoxythymidine triphosphate
['Thymidine','cpd00184'], #EXs :
['Thyminose','cpd01242'], #EXs : deoxyribose
['Uracil','cpd00092'], #EXs :
['Uridine','cpd00249'], #EXs :
['XAN','cpd00309'], #EXs : Xanthine
['Xanthosine','cpd01217'], #EXs :
['dATP','cpd00115'], #EXs :
['dGTP','cpd00241'], #EXs :
['dTMP','cpd00298'] #EXs :
]
# Check to see if these metabolites are used in pathways? Should I add some of these to media?
# +
# Energy metabolite precursors
[
['NADP','cpd00006'], #EXs :
['PPi','cpd00012'] #EXs :
]
# Enviromental Metabolites (Checked and added above)
[
['CO2','cpd00011'], #EXs :
['Ca2+','cpd00063'], #EXs :
['Cd2+','cpd01012'], #EXs :
['chromate','cpd11595'], #EXs :
['Cl-','cpd00099'], #EXs :
['Co2+','cpd00149'], #EXs :
['Cu2+','cpd00058'], #EXs :
['Fe2+','cpd10515'], #EXs :
['H+','cpd00067'], #EXs :
['H2','cpd11640'], #EXs :
['H2O','cpd00001'], #EXs :
['Hg2+','cpd00531'], #EXs :
['K+','cpd00205'], #EXs :
['Mg','cpd00254'], #EXs :
['Mn2+','cpd00030'], #EXs :
['N2','cpd00528'], #EXs :
['Na+','cpd00971'], #EXs :
['Ni2+','cpd00244'], #EXs :
['O2','cpd00007'], #EXs :
['Pb','cpd04097'], #EXs :
['Zn2+','cpd00034'], #EXs :
['fe3','cpd10516'] #EXs :
]
# 'Don't include' and Misc. metabolites
[
['4-Hydroxybenzoate','cpd00136'], #EXs :
['ARSENOBETAINE','cpd11597'], #EXs :
['Butyro-betaine','cpd08306'], #EXs :
['Choline sulfate','cpd00681'], #EXs :
['Citrate','cpd00137'], #EXs : Already in BSM
['CoA','cpd00010'], #EXs : Organisms should be required to produce this
['Dephospho-CoA','cpd00655'], #EXs :
['Ferrichrome','cpd03724'], #EXs :
['Glycerol-3-phosphate','cpd00080'], #EXs :
['Heme','cpd00028'], #EXs : Already have Fe2+
['MOPS','cpd11575'], #EXs :
['Lanosterol','cpd01188'], #EXs :
['Oxidized glutathione','cpd00111'], #EXs : Important antioxident : allicin (http://www.jbc.org/content/291/22/11477.full)
['butanesulfonate','cpd11596'], #EXs :
['hexanesulfonate','cpd11578'], #EXs :
['ocdca','cpd01080'] #EXs :
]
# +
# Production
# H2O2 -- cpd00025
# Acetate -- cpd00029
# Butyrate -- cpd00211
# isobutyrate -- cpd01711
# GABA -- cpd00281
# ethanol -- cpd00363
# Propionate -- cpd00141
# formate -- cpd00047
# Valerate -- cpd00597
# Isovaleric acid -- cpd05178 (wrong eqn)
# sulforaphane --
# https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5541232/
# thiamin -- cpd00305
# Pyridoxal phosphate (B6) -- cpd00016
# BIOT (biotin, B7) -- cpd00104
# (CH3)3NO (TMAO) -- cpd00811
# Indole-3-(carb)aldehyde -- cpd05401
# https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4337795/
# Acetaldehyde -- cpd00071
# Deoxycholate -- cpd02733
# Chorismate -- cpd00216
# Hexanoate -- cpd01113
#
# Consumption
# Galactose -- cpd00108; cpd01112
# L-galactose -- cpd01257
# lactose -- cpd00208
# beta-lactose -- cpd01354
# sucrose -- cpd00076
# trehalose (TRHL) -- cpd00794
# maltose -- cpd00179
# D-Mannose -- cpd00138
# D-Fructose -- cpd00082
# Inulin -- cpd27312
# ethanol -- cpd00363
# Carnitine -- cpd00266
# Citrate -- cpd00137
# GLUM (D-glucosamine) -- cpd00276
#
# -
model = cobra.io.read_sbml_model('../gap_models/1051650.8.xml')
model.reactions.get_by_id('bio1')
biomass = model.reactions.get_by_id('bio1')
# +
biomass_rcts = defaultdict(list)
for rct in biomass.reactants:
biomass_rcts[rct.name].append(rct.id)
biomass_rcts
# +
biomass_prdt = defaultdict(list)
for prdt in biomass.products:
biomass_prdt[prdt.name].append(prdt.id)
biomass_prdt
# +
| Code/OldCode/Test_lactate_production.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import h5py
import MFParse
import matplotlib.pyplot as plt
test_egg = '/Users/ziegler/Documents/egg_files/locust_mc_Seed603_Angle90.00_Pos0.000.egg'
#test_egg = '/Users/ziegler/Documents/egg_files/locust_mc_Seed400_LO25.8781G_Radius0.100_Pos0.000.egg'
parsed_egg = MFParse.parse_egg(test_egg)
# +
summed_egg = np.sum(parsed_egg, axis=0)
time_data = summed_egg[2*8192: 3*8192]
plt.figure()
plt.plot(time_data)
plt.xlim(0,100)
fft_data = np.fft.fftshift(np.fft.fft(time_data))
plt.figure()
plt.plot(freqs, (abs(fft_data)))
# -
freqs = np.fft.fftshift(np.fft.fftfreq(8192, 5e-9))
freqs
| analysis/202141_compress_egg_file.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1>Launch Websocket OpenGrid Node on Heroku</h1>
# <h2>Step 1: Import required dependencies</h2>
import syft as sy
import torch as th
import grid as gr
hook = sy.TorchHook(th)
# <h2>Step 2: Deploy websocket application </h2>
address = gr.launch_on_heroku("websocketgridtest",
dev_user="IonesioJunior",
app_type="websocket",
branch="add_websocket_approach",
verbose=True,
check_deps=True)
# <h2>Step 3 : Create / connect client to grid application</h2>
worker = gr.WebsocketGridClient(hook, "https://websocketgridtest.herokuapp.com/",id="grid")
worker.connect()
# <h2>Step 4: Use PySyft Like Normal</h2>
x = th.tensor([1,2,3,4,5]).send(worker)
x
x.get()
y = th.tensor([1,2,3,4,5,6]).send(worker)
z = y + y
z
z.get()
| examples/experimental/Launch_Websocket_Grid_worker.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={} colab_type="code" id="BI0iVW92iSJd"
# !git clone https://github.com/IBM/dl-interpretability-compbio.git
# !pip3 install dl-interpretability-compbio/
# + colab={} colab_type="code" id="aSAvcwv1iVQO"
# !pip3 install -I scikit-image>=0.14.2
# + [markdown] colab_type="text" id="UAqIrqFGiSJj"
# TODO NOW: Restart the runtime
# + colab={} colab_type="code" id="Fq9eHyvuibYp"
import skimage
print(skimage.__version__)
# + colab={} colab_type="code" id="3MUp7-fOiSJl"
# !wget https://repo.anaconda.com/archive/Anaconda3-5.2.0-Linux-x86_64.sh
# !bash Anaconda3-5.2.0-Linux-x86_64.sh -b -f -p /usr/local
# !conda install -y rdkit=2019.03.1=py36hc20afe1_1 -c https://conda.anaconda.org/rdkit
# + colab={} colab_type="code" id="21wgjLuJiSJo"
import sys
sys.path.insert(0, '/usr/local/lib/python3.6/site-packages')
# %cd /content/dl-interpretability-compbio/notebooks
# + [markdown] colab_type="text" id="REAVyd2-iSJr"
# # Training of a super simple model for celltype classification
# + colab={} colab_type="code" id="55vponomiSJs"
import tensorflow as tf
# !which python
# !python --version
print(tf.VERSION)
print(tf.keras.__version__)
# !pwd # start jupyter under notebooks/ for correct relative paths
# + colab={} colab_type="code" id="Jawnc9gBiSJu"
import datetime
import inspect
import pandas as pd
import numpy as np
import seaborn as sns
from tensorflow.keras import layers
from tensorflow.keras.utils import to_categorical
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import MinMaxScaler
from depiction.models.examples.celltype.celltype import one_hot_encoding, one_hot_decoding
# + [markdown] colab_type="text" id="eV99rgAwiSJx"
# ### a look at the data
# labels are categories 1-20, here's the associated celltype:
# + colab={} colab_type="code" id="q8ZrSw9CiSJy"
meta_series = pd.read_csv('../data/single-cell/metadata.csv', index_col=0)
meta_series
# + [markdown] colab_type="text" id="mPDAVEQmiSJ0"
# There are 13 unbalanced classes, and over 80k samples
# + colab={} colab_type="code" id="GVakpKb1iSJ1"
data_df = pd.read_csv('../data/single-cell/data.csv')
data_df.groupby('category').count()['CD45']
# + colab={} colab_type="code" id="xb-1Jj1kiSJ3"
data_df.sample(n=10)
# + colab={} colab_type="code" id="NsLSuRHUiSJ5"
print(inspect.getsource(one_hot_encoding)) # from keras, but taking care of 1 indexed classes
print(inspect.getsource(one_hot_decoding))
# + colab={} colab_type="code" id="BxWRaWAeiSJ7"
classes = data_df['category'].values
labels = one_hot_encoding(classes)
#scale the data from 0 to 1
min_max_scaler = MinMaxScaler(feature_range=(0, 1), copy=True)
data = min_max_scaler.fit_transform(data_df.drop('category', axis=1).values)
data.shape
# + colab={} colab_type="code" id="IQ8HhcwKiSJ-"
one_hot_decoding(labels)
# + colab={} colab_type="code" id="l6eON_pCiSKA"
data_train, data_test, labels_train, labels_test = train_test_split(
data, labels, test_size=0.33, random_state=42, stratify=data_df.category)
# + colab={} colab_type="code" id="S64rOJlKiSKD"
labels
# + colab={} colab_type="code" id="aEjhisuLiSKF"
batchsize = 32
# + colab={} colab_type="code" id="InmbMsjtiSKH"
dataset = tf.data.Dataset.from_tensor_slices((data_train, labels_train))
dataset = dataset.shuffle(2 * batchsize).batch(batchsize)
dataset = dataset.repeat()
testset = tf.data.Dataset.from_tensor_slices((data_test, labels_test))
testset = testset.batch(batchsize)
# + [markdown] colab_type="text" id="8rESJF4JiSKK"
# ### I don't know how a simpler network would look like
# + colab={} colab_type="code" id="cQtHkUufiSKK"
model = tf.keras.Sequential()
# Add a softmax layer with output units per celltype:
model.add(layers.Dense(
len(meta_series), activation='softmax',
batch_input_shape=tf.data.get_output_shapes(dataset)[0]
))
# + colab={} colab_type="code" id="eAeszUQliSKN"
model.summary()
# + colab={} colab_type="code" id="X7CgpdruiSKP"
model.compile(optimizer=tf.keras.optimizers.Adam(0.001),
loss='categorical_crossentropy',
metrics=[tf.keras.metrics.categorical_accuracy])
# + colab={} colab_type="code" id="5eFeD1IIiSKR"
# evaluation on testset on every epoch
# log_dir="logs/fit/" + datetime.datetime.now().strftime("%Y%m%d-%H%M%S")
# tensorboard_callback = tf.keras.callbacks.TensorBoard(log_dir=log_dir, histogram_freq=1)
model.fit(
dataset,
epochs=20, steps_per_epoch=np.ceil(data_train.shape[0]/batchsize),
validation_data=testset, # callbacks=[tensorboard_callback]
)
# + [markdown] colab_type="text" id="eGPzA56qiSKV"
# ### Is such a simple model interpretable?
# + colab={} colab_type="code" id="TmjmEdh6iSKX"
# Save entire model to a HDF5 file
model.save('./celltype_model.h5')
# + colab={} colab_type="code" id="h0frkNhEiSKa"
# tensorboard --logdir logs/fit
# + colab={} colab_type="code" id="0NNW9sjdiSKb"
# To recreate the exact same model, including weights and optimizer.
# model = tf.keras.models.load_model('../data/models/celltype_dnn_model.h5')
# + [markdown] colab_type="text" id="2EMPgmwMiSKd"
# ## What is the effect of increasing model complexity?
# Play around by adding some layers, train and save the model under some name to use with the other notebook.
# + [markdown] colab_type="text" id="VQeWMkhxiSKe"
# 
# + colab={} colab_type="code" id="GpcD6psdiSKe"
model = tf.keras.Sequential()
# Adds a densely-connected layers with 64 units to the model:
model.add(layers.Dense(64, activation='relu', batch_input_shape=tf.data.get_output_shapes(dataset)[0])) #
# ...
# do whatever you want
# model.add(layers.Dense(64, activation='relu'))
# model.add(layers.Dropout(0.5))
# ...
# Add a softmax layer with output units per celltype:
model.add(layers.Dense(len(meta_series), activation='softmax'))
# + colab={} colab_type="code" id="ev4jSLwDinZm"
# %reset
# + [markdown] colab_type="text" id="y9eFrzKRiSKh"
# # Interpreting Cell Typer
# + colab={} colab_type="code" id="6DENxkXAiSKi"
import pandas as pd
import numpy as np
import seaborn as sns
import ipywidgets as widgets
from ipywidgets import interact, interact_manual
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import MinMaxScaler
from depiction.models.base.base_model import BaseModel
from depiction.models.examples.celltype.celltype import CellTyper
from depiction.interpreters.u_wash.u_washer import UWasher
from depiction.interpreters.alibi import Counterfactual
from depiction.interpreters.aix360.rule_based_model import RuleAIX360
from depiction.models.base import BinarizedClassifier
from depiction.core import Task, DataType
from tensorflow import keras
# + [markdown] colab_type="text" id="ngLakJpmiSKk"
# ### Data
# + colab={} colab_type="code" id="EYzJqtxAiSKk"
# Load data
datapath = '../data/single-cell/data.csv'
data_df = pd.read_csv(datapath)
#scale the data from 0 to 1
min_max_scaler = MinMaxScaler(feature_range=(0, 1), copy=True)
data = min_max_scaler.fit_transform(data_df.drop('category', axis=1).values)
data_df = pd.DataFrame(
np.append(data, data_df['category'].values[:, None], axis=1), index=data_df.index, columns=data_df.columns
)
# split as in traing of the model
train_df, test_df = train_test_split(data_df, test_size=0.33, random_state=42, stratify=data_df.category)
test_df, valid_df = train_test_split(test_df, test_size=0.67, random_state=42, stratify=test_df.category)
train_df.head()
# + colab={} colab_type="code" id="MRzHOTWqiSKm"
markers = train_df.columns[:-1]
X_train = train_df[markers].values
X_test = test_df[markers].values
X_valid = valid_df[markers].values
y_train = train_df['category'].values.astype(np.int)
y_test = test_df['category'].values.astype(np.int)
y_valid = valid_df['category'].values.astype(np.int)
# + colab={} colab_type="code" id="P8T5vhFGiSKo"
sns.countplot(data_df.category)
CellTyper.celltype_names
# + [markdown] colab_type="text" id="Kjs3N2JyiSKq"
# ### Loading a pretrained model
# is actually done under the hood by a child implementation of `depiction.models.uri.HTTPModel`.
# Change `filename`, `cache_dir` (with fixed subdir `models/`) and/or `origin` to load/download a different model.
# Or have a look at other uri models, e.g `FileSystemModel` or `RESTAPIModel`.
# + colab={} colab_type="code" id="OYAIrv76iSKr"
# Import trained classifier
classifier = CellTyper(filename='celltype_model.h5', cache_dir='/Users/dow/Box/Molecular_SysBio/data/dl-interpretability-compbio') # FIX
# + colab={} colab_type="code" id="aH3BX_a6iSKu"
classifier.model.summary()
# + [markdown] colab_type="text" id="XBrX2_iMiSKx"
# ### Layer weights
# + colab={} colab_type="code" id="rdqdqiGKiSKx"
weights = classifier.model.layers[0].get_weights()[0]
sns.heatmap(pd.DataFrame(
weights,
index=markers,
# columns=CellTyper.celltype_names.values()
).T)
# + [markdown] colab_type="text" id="M15PQCQxiSKz"
# Compare qualitatively to __B__ and **C** (thought the image is not depicting this exact dataset)
# 
# from https://science.sciencemag.org/content/332/6030/687/tab-figures-data
# + [markdown] colab_type="text" id="MNpC4qYIiSK0"
# helper/widget functions
# + colab={} colab_type="code" id="eE0UnzXjiSK0"
def random_from_class(label):
id_sample_to_explain = test_df.reset_index().query('category==@label').sample(n=1).index[0]
print('Interpreting sample with index {} in test_df'.format(id_sample_to_explain))
return id_sample_to_explain
# + colab={} colab_type="code" id="MlDToOWuiSK2"
def visualize_logits(id_sample_to_explain):
sample = X_test[id_sample_to_explain]
logits = pd.DataFrame(classifier.predict([[sample]]), columns=CellTyper.celltype_names.values()).T
sns.heatmap(logits)
def visualize(id_sample_to_explain, layer):
sample = X_test[id_sample_to_explain]
if layer is None:
visualize_logits(id_sample_to_explain)
return
elif layer==0:
# output of last "layer" is the sample
layer_output = sample.transpose()
else:
# for vizualization of output of a layer we access the model
activation_model = keras.models.Model(
inputs=classifier.model.input,
outputs=classifier.model.layers[layer-1].output
)
layer_output = activation_model.predict([[sample]])[0]
weights = classifier.model.layers[layer].get_weights()[0]
weighted_output = (weights.transpose() * layer_output)
sns.heatmap(weighted_output)
def visualize_random_from_class(label, layer):
visualize(random_from_class(label), layer)
# + colab={} colab_type="code" id="fSnQhwUNiSK3"
interact_manual(
visualize_random_from_class,
label=[(v, k) for k, v in classifier.celltype_names.items()],
layer=dict(
**{layer.name: i for i, layer in enumerate(classifier.model.layers)}, logits=None
)
);
# + colab={} colab_type="code" id="C-miVx5niSK5"
visualize_logits(4368)
# + [markdown] colab_type="text" id="udEv_avZiSK7"
# ## Interpretability methods
# starting with "local" methods, explaining a given sample.
# + [markdown] colab_type="text" id="iC6x1bWYiSK7"
# ### Lime
# + colab={} colab_type="code" id="0LRUbxytiSK8"
# Create a LIME tabular interpreter
lime_params = {
'training_data': X_train,
'training_labels': y_train,
'feature_names': markers,
'verbose': True,
'class_names': classifier.celltype_names.values(),
'discretize_continuous': False,
'sample_around_instance': True
}
lime = UWasher("lime", classifier, **lime_params)
# + [markdown] colab_type="text" id="-6qrEnSbiSK-"
# ### Anchor
# + colab={} colab_type="code" id="ZumEHjyyiSK-"
anchors_params = {
'feature_names': markers,
'class_names': classifier.celltype_names.values(),
'categorical_names': {}
}
fit_params = { # depiction fits the anchor (tabular) on contruction.
'train_data': X_train,
'train_labels': y_train,
'validation_data': X_valid,
'validation_labels': y_valid
}
anchors = UWasher('anchors', classifier, **fit_params, **anchors_params)
# + [markdown] colab_type="text" id="puFPXmFMiSLA"
# ### Counterfactual
# + colab={} colab_type="code" id="ux8jtGm4iSLA"
counterfactual_params = {
# setting some parameters
'shape': (1, 13), # with batch size
'target_proba': 1.0,
'tol': 0.1, # tolerance for counterfactuals
'max_iter': 10,
'lam_init': 1e-1,
'max_lam_steps': 10,
'learning_rate_init': 0.1,
'feature_range': (X_train.min(),X_train.max())
}
counterfactual = Counterfactual(
classifier,
target_class='other', # any other class
**counterfactual_params,
)
# + [markdown] colab_type="text" id="2TZuV4qZiSLC"
# helper/widget functions
# + colab={} colab_type="code" id="5RX1nHYeiSLD"
def interpret_with_lime(id_sample_to_explain):
"""Explain the chosen instance wrt the chosen label."""
lime.interpret(X_test[id_sample_to_explain], explanation_configs={'top_labels': 1})
def anchor_callback(sample, **kwargs):
"""Explain the chosen instance wrt the chosen labels."""
return np.argmax(classifier.predict(sample,**kwargs), axis=1)
def interpret_with_anchor(id_sample_to_explain):
anchors.interpret(X_test[id_sample_to_explain], explanation_configs={},callback=anchor_callback)
def interpret_with_counterfactual(id_sample_to_explain):
"""Explain the chosen instance wrt the chosen label."""
explanation = counterfactual.interpret(np.expand_dims(X_test[id_sample_to_explain], axis=0)) # with batch size
predicted_class = explanation['cf']['class']
probability = explanation['cf']['proba'][0][predicted_class]
print(f'Counterfactual prediction: {predicted_class} with probability {probability}')
print(explanation['cf']['X'])
# + colab={} colab_type="code" id="_QOXqYlUjRwQ"
interpret_with_lime(4368)
# + colab={} colab_type="code" id="ok_mixlIiSLI"
interpret_with_anchor(4368)
# + colab={} colab_type="code" id="Qhl8HCDVjRIJ"
interpret_with_counterfactual(4368)
# + [markdown] colab_type="text" id="g_CH6WLEiSLK"
# ## Global interpretation with rule-based models
# + colab={} colab_type="code" id="49qpykyBiSLL"
LABEL2ID = {CellTyper.celltype_names[i]: i for i in CellTyper.celltype_names.keys()}
# + colab={} colab_type="code" id="lZjxJ4hFiSLM"
LABEL_TO_EXPLAIN = 'Mature CD4+ T'
LABEL_ID = LABEL2ID[LABEL_TO_EXPLAIN]
# + [markdown] colab_type="text" id="kE7hBlIGiSLP"
# ### Data preparation and auxiliary functions
# + colab={} colab_type="code" id="_tV-CwjwiSLP"
# Binarize the task to use this method
model = BinarizedClassifier(classifier, data_type=DataType.TABULAR, label_index=LABEL_ID)
# + [markdown] colab_type="text" id="KhFhOYehiSLQ"
# ### Post-Hoc explanation
# + [markdown] colab_type="text" id="B3GzOI4niSLR"
# #### BRCG
# + colab={} colab_type="code" id="6oq_RSfxiSLR"
interpreter = RuleAIX360('brcg', X=X_train, model=model)
interpreter.interpret()
# + [markdown] colab_type="text" id="a_J86oImiSLT"
# #### GLRM - Linear
# + colab={} colab_type="code" id="0Tz1yzsjiSLU"
interpreter = RuleAIX360('glrm_linear', X=X_train, model=model)
interpreter.interpret()
# + [markdown] colab_type="text" id="i8X06lUMiSLV"
# #### GLRM - Logistic
# + colab={} colab_type="code" id="KPZiAcWAiSLW"
interpreter = RuleAIX360('glrm_logistic', X=X_train, model=model)
interpreter.interpret()
# + [markdown] colab_type="text" id="xtK1dmS7iSLX"
# ### Ante-Hoc explanation
# + colab={} colab_type="code" id="BvgKj6WRiSLY"
y_train_binary = y_train == LABEL_ID
# + [markdown] colab_type="text" id="xdysL5mtiSLa"
# #### BRCG
# + colab={} colab_type="code" id="NLI3f8OOiSLa"
interpreter = RuleAIX360('brcg', X=X_train, y=y_train_binary)
interpreter.interpret()
# + [markdown] colab_type="text" id="wQbK3FbBiSLc"
# #### GLRM - Linear
# + colab={} colab_type="code" id="toLPn-R8iSLc"
interpreter = RuleAIX360('glrm_linear', X=X_train, y=y_train_binary)
interpreter.interpret()
# + [markdown] colab_type="text" id="37c9s7hdiSLd"
# #### GLRM - Logistic
# + colab={} colab_type="code" id="0D1TeBo4iSLe"
interpreter = RuleAIX360('glrm_logistic', X=X_train, y=y_train_binary)
interpreter.interpret()
# + colab={} colab_type="code" id="pXaAyU_VjmwD"
# %reset
# + [markdown] colab_type="text" id="kMbwiDk_iSLf"
# # Having fun with DeepBind
# + colab={} colab_type="code" id="0C6L0pJYiSLf"
from depiction.models.examples.deepbind.deepbind_cli import DeepBind
from depiction.core import Task, DataType
from depiction.interpreters.u_wash.u_washer import UWasher
from depiction.models.examples.deepbind.deepbind_cli import create_DNA_language
from ipywidgets import interact
# + [markdown] colab_type="text" id="WGelUbnEiSLh"
# ### Setup task
# + colab={} colab_type="code" id="-_TDAJpniSLh"
class_names = ['NOT BINDING', 'BINDING']
tf_factor_id = 'D00328.003' # CTCF
classifier = DeepBind(tf_factor_id = tf_factor_id)
# + [markdown] colab_type="text" id="MhycDJg1iSLl"
# ## Instantiate the interpreters
# + colab={} colab_type="code" id="tBVe1f1fiSLl"
# Create a LIME text interpreter
interpreter = 'lime'
lime_explanation_configs = {
'labels': (1,),
}
interpreter_params = {
'class_names': class_names,
'split_expression': list,
'bow': False,
'char_level': True
}
lime_explainer = UWasher(interpreter, classifier, **interpreter_params)
# Create Anchor text intepreter
interpreter = 'anchors'
anchors_explanation_configs = {
'use_proba': False,
'batch_size': 100
}
interpreter_params = {
'class_names': class_names,
'nlp': create_DNA_language(),
'unk_token': 'N',
'sep_token': '',
'use_unk_distribution': True
}
anchor_explainer = UWasher(interpreter, classifier, **interpreter_params)
# + [markdown] colab_type="text" id="TiJgyKaJiSLq"
# ## Let's use LIME and Anchors
# + colab={} colab_type="code" id="j7y4z_6miSLr"
# sequences = [
# 'AGGCTAGCTAGGGGCGCCC', 'AGGCTAGCTAGGGGCGCTT', 'AGGGTAGCTAGGGGCGCTT',
# 'AGGGTAGCTGGGGGCGCTT', 'AGGCTAGGTGGGGGCGCTT', 'AGGCTCGGTGGGGGCGCTT',
# 'AGGCTCGGTAGGGGGCGATT'
# ]
sequence = 'AGGCTCGGTAGGGGGCGATT'
classifier.use_labels = False
lime_explainer.interpret(sequence, explanation_configs=lime_explanation_configs)
classifier.use_labels = True
anchor_explainer.interpret(sequence, explanation_configs=anchors_explanation_configs)
# + [markdown] colab_type="text" id="ZDZmQgtpiSLt"
# CTCF binding motif
# 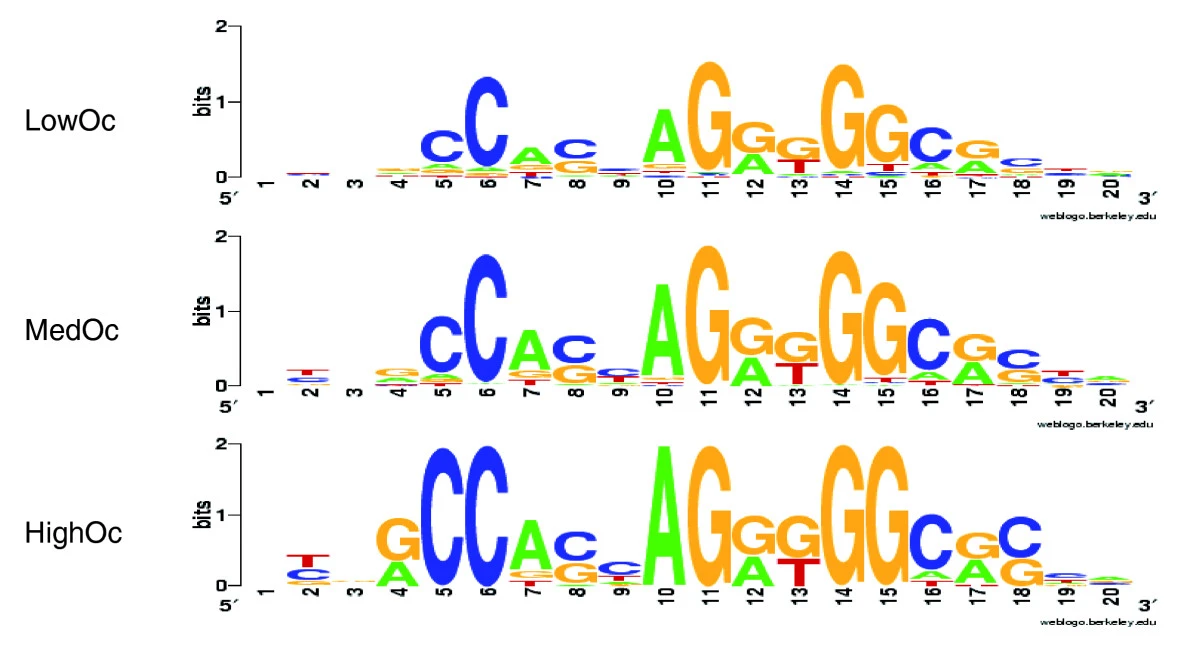
# from Essien, Kobby, et al. "CTCF binding site classes exhibit distinct evolutionary, genomic, epigenomic and transcriptomic features." Genome biology 10.11 (2009): R131.
# + colab={} colab_type="code" id="3cP4GKyXiSLt"
tf_factor_id = 'D00761.001' # FOXA1
classifier.tf_factor_id = tf_factor_id
# + colab={} colab_type="code" id="xSwBJ-gakCMb"
sequence = 'TGTGTGTGTG'
classifier.use_labels = False
lime_explainer.interpret(sequence, explanation_configs=lime_explanation_configs)
classifier.use_labels = True
anchor_explainer.interpret(sequence, explanation_configs=anchors_explanation_configs)
# + [markdown] colab_type="text" id="8teB12aXiSLu"
# FOXA1 binding motif
# 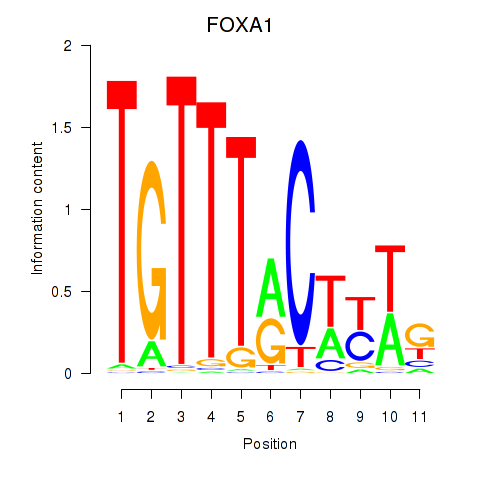
# from https://ismara.unibas.ch/supp/dataset1_IBM_v2/ismara_report/pages/FOXA1.html
# + colab={} colab_type="code" id="Er_2zKiNkFm8"
# %reset
# + [markdown] colab_type="text" id="zKE7J7g-iSLv"
# # Understanding PaccMann
# + colab={} colab_type="code" id="2jD1S2g5iSLv"
# %%capture
# import all the needed libraries
import numpy as np
import pandas as pd
import tempfile
from rdkit import Chem
from sklearn.model_selection import train_test_split
import ipywidgets as widgets
from ipywidgets import interact, interact_manual
from IPython.display import SVG, display
from depiction.models.examples.paccmann import PaccMannSmiles, PaccMannCellLine
from depiction.models.examples.paccmann.smiles import (
get_smiles_language, smiles_attention_to_svg,
process_smiles, get_atoms
)
from depiction.core import Task, DataType
from depiction.interpreters.u_wash.u_washer import UWasher
cache_dir = tempfile.mkdtemp()
# + [markdown] colab_type="text" id="T0zz9L-CiSLz"
# ### Data
# + colab={} colab_type="code" id="lLpPOiwiiSL0"
# Parse data from GDSC
# drugs
drugs = pd.read_csv(
'../data/paccmann/gdsc.smi', sep='\t',
index_col=1, header=None,
names=['smiles']
)
# cell lines
cell_lines = pd.read_csv('../data/paccmann/gdsc.csv.gz', index_col=1)
genes = cell_lines.columns[3:].tolist()
# sensitivity data
drug_sensitivity = pd.read_csv('../data/paccmann/gdsc_sensitivity.csv.gz', index_col=0)
# labels
class_names = ['Not Effective', 'Effective']
# + [markdown] colab_type="text" id="__OHTJMjiSL2"
# ### Interpretability on the drug level for a cell line of interest
# + [markdown] colab_type="text" id="VPXGSQ8PiSL2"
# #### LIME and Anchor
# + colab={} colab_type="code" id="goRqf0l8iSL3"
# pick a cell line
selected_cell_line = 'NCI-H1648'
# filter and prepare data
selected_drug_sensitivity = drug_sensitivity[
drug_sensitivity['cell_line'] == selected_cell_line
]
selected_drugs = drugs.reindex(selected_drug_sensitivity['drug']).dropna()
selected_drug_sensitivity = selected_drug_sensitivity.set_index('drug').reindex(
selected_drugs.index
).dropna()
# setup a classifier for the specific cell line
classifier = PaccMannSmiles(cell_lines.loc[selected_cell_line][genes].values, cache_dir=cache_dir)
# interpretablity methods
def interpret_smiles_with_lime(example):
explanation_configs = {
'labels': (1,),
}
interpreter_params = {
'class_names': class_names,
'split_expression': list,
'bow': False,
'char_level': True
}
explainer = UWasher('lime', classifier, **interpreter_params)
explainer.interpret(example, explanation_configs=explanation_configs)
def interpret_smiles_with_anchor(example):
explanation_configs = {
'use_proba': False,
'batch_size': 32,
}
interpreter_params = {
'class_names': class_names,
'nlp': get_smiles_language(),
'unk_token': '*',
'sep_token': '',
'use_unk_distribution': True
}
explainer = UWasher('anchors', classifier, **interpreter_params)
def predict_wrapper(samples):
return np.argmax(classifier.predict(samples), axis=1)
explainer.interpret(example, explanation_configs=explanation_configs, callback=predict_wrapper)
def interpret_smiles(interpreter, drug):
if interpreter == 'lime':
interpret_smiles_with_lime(drugs.loc[drug].item())
else:
interpret_smiles_with_anchor(drugs.loc[drug].item())
# + colab={} colab_type="code" id="YKOVmjtIiSL5"
# Dummy just to visualize the drugs neatly in Colab
interact_manual(
lambda drug: print(drug),
drug=drugs.index
);
# + colab={} colab_type="code" id="Zu-CDGwgkRuh"
interpret_smiles_with_lime(drugs.loc['PHA-793887'].item())
# + colab={} colab_type="code" id="BImYLoe3kUZa"
interpret_smiles_with_anchor(drugs.loc['PHA-793887'].item())
# + [markdown] colab_type="text" id="uAkABHTeiSL7"
# #### What about PaccMann's attention?
# + colab={} colab_type="code" id="sUbhTly4iSL7"
# pick a cell line
selected_cell_line = 'NCI-H1648'
# setup a classifier for the specific cell line
classifier = PaccMannSmiles(cell_lines.loc[selected_cell_line][genes].values, cache_dir=cache_dir)
# + colab={} colab_type="code" id="qaTxmpA3iSL8"
def attention_smiles(drug):
try:
smiles = drugs.loc[drug].item()
molecule = Chem.MolFromSmiles(smiles)
atoms = get_atoms(smiles)
_ = classifier.predict([smiles])
smiles_attention = next(classifier.predictor.predictions)['smiles_attention'][0]
display(SVG(smiles_attention_to_svg(smiles_attention, atoms, molecule)))
except:
print('Structure visualization not supported')
# + colab={} colab_type="code" id="362fNGJOiSL-"
interact(
attention_smiles,
drug=drugs.index
);
# + [markdown] colab_type="text" id="mRCCc8ONiSL_"
# ### Interpretability on the cell line level for a drug of interest
# + [markdown] colab_type="text" id="b1Mh0ApkiSL_"
# #### LIME and Anchor
# + colab={} colab_type="code" id="fjcSoA67iSMA"
# pick a drug
selected_drug = 'Imatinib'
# filter and prepare data
selected_drug_sensitivity = drug_sensitivity[
drug_sensitivity['drug'] == selected_drug
]
selected_cell_lines = cell_lines.reindex(selected_drug_sensitivity['cell_line']).dropna()
selected_drug_sensitivity = selected_drug_sensitivity.set_index('cell_line').reindex(
selected_cell_lines.index
).dropna()
X_train, X_test, y_train, y_test = train_test_split(
selected_cell_lines[genes].values, selected_drug_sensitivity['effective'].values
)
X_test, X_valid, y_test, y_valid = train_test_split(
X_test, y_test
)
# setup a classifier for the specific drug
classifier = PaccMannCellLine(drugs.loc[selected_drug].item(), cache_dir=cache_dir)
# interpretablity methods
def interpret_cell_line_with_lime(example):
explanation_configs = {
'labels': (1,),
}
interpreter_params = {
'training_data': X_train,
'training_labels': y_train,
'feature_names': genes,
'class_names': class_names,
'discretize_continuous': False,
'sample_around_instance': True
}
explainer = UWasher('lime', classifier, **interpreter_params)
explainer.interpret(example, explanation_configs=explanation_configs)
def interpret_cell_line_with_anchor(example):
explanation_configs = {}
interpreter_params = {
'feature_names': genes,
'class_names': class_names,
'categorical_names': {}
}
explainer = UWasher('anchors', classifier, **interpreter_params)
explainer.explainer.fit(
X_train, y_train, X_valid, y_valid
)
def predict_wrapper(samples):
return np.argmax(classifier.predict(samples), axis=1)
explainer.interpret(example, explanation_configs=explanation_configs, callback=predict_wrapper)
def interpret_cell_line(interpreter, cell_line):
if interpreter == 'lime':
interpret_cell_line_with_lime(
cell_lines.loc[cell_line][genes].values
)
else:
interpret_cell_line_with_anchor(
cell_lines.loc[cell_line][genes].values
)
# + colab={} colab_type="code" id="uoXjLbSRiSMB"
# Dummy just to visualize the cell lines neatly
interact_manual(
lambda cell_line: print(cell_line),
cell_line=cell_lines.index
);
# + colab={} colab_type="code" id="FMxvtFZJkikp"
interpret_cell_line_with_lime(
cell_lines.loc['JiyoyeP-2003'][genes].values
)
# + colab={} colab_type="code" id="YjMI87FekkXI"
interpret_cell_line_with_anchor(
cell_lines.loc['JiyoyeP-2003'][genes].values
)
# + [markdown] colab_type="text" id="mL5Z8YjviSMC"
# #### What about PaccMann's attention?
# + colab={} colab_type="code" id="FJ1wJFYoiSMC"
# pick a drug
selected_drug = 'Imatinib'
classifier = PaccMannCellLine(drugs.loc[selected_drug].item(), cache_dir=cache_dir)
# + colab={} colab_type="code" id="gDM6Sy8WiSMD"
def attention_cell_line(cell_line, top_k=10):
try:
_ = classifier.predict([cell_lines.loc[cell_line][genes].values])
gene_attention = next(classifier.predictor.predictions)['gene_attention'][0]
pd.Series(dict(zip(genes, gene_attention))).sort_values(ascending=False)[:top_k].plot.bar()
except:
print('Cell line visualization not supported')
# + colab={} colab_type="code" id="29phf-POiSMG"
interact(
attention_cell_line, cell_line=cell_lines.index,
top_k=(1, 30, 1)
);
# + colab={} colab_type="code" id="jMWjBB62iSMI"
| workshops/20191121_PyPharma19/tutorial_colab.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
# default_exp utils
# -
# # Utils
#
# > Some utility functions.
#hide
from nbdev.showdoc import *
# +
#export
import json # 2.0.9
import numpy as np
import glob
import pandas as pd
import os
def merge_data(folder, drop=[], limit = None):
all_files = glob.glob(os.path.join(folder, "*.pkl"))
li = []
for filename in all_files[:limit]:
df = pd.read_pickle(filename)
df = df.drop(columns = drop)
li.append(df)
data = pd.concat(li, axis=0, ignore_index=True)
return data
def loadJson(path, default = {}):
"""Return dictionary from json file."""
if os.path.isfile(path):
try:
with open(path, 'r', encoding = 'utf-8') as f:
return json.loads(f.read(), strict = False)
except:
return default
else:
return default
| notebooks/04_utils.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#Authors: <NAME> - C16315253, <NAME> - C16497656
#import libraries
import pandas as pd
import sklearn
from sklearn.model_selection import train_test_split
from sklearn.naive_bayes import GaussianNB
from sklearn import preprocessing
from sklearn.metrics import accuracy_score
#gethering the columns names
columnsName = open("headers.txt")
text_reader = columnsName.read()
#reading the csv file on which the predictions will be made
dataFrame = pd.read_csv('queries.csv',header = None, na_values = ["?", " ?", "? "])
dataFrame.columns = text_reader.split("\n")
#reading the file which we will train the model on
dataFrame2 = pd.read_csv('trainingset.csv',header = None, na_values = ["?", " ?", "? "])
dataFrame2.columns = text_reader.split("\n")
#gethering the converter to translate the catergorically features to continous
le = preprocessing.LabelEncoder()
#creating the collection which we do not want to train on
DictCollection = ["id"]
#Formatting the subscibed feature
dictFrameTarget = dataFrame2['subscribed'].to_dict()
for r in dictFrameTarget:
if (dictFrameTarget[r] == "yes"):
dictFrameTarget[r] = 1
elif(dictFrameTarget[r] == "no"):
dictFrameTarget[r] = 0
#converting the dataframe coloumns to continius features
ProcessedData = {}
le = preprocessing.LabelEncoder()
DictCollection = ["id","subscribed"]
for key in dataFrame2.columns:
if key not in DictCollection:
ProcessedData[key] = le.fit_transform(dataFrame2[key])
ProcessedPrediction = {}
for key in dataFrame.columns:
if key not in DictCollection:
ProcessedPrediction[key] = le.fit_transform(dataFrame[key])
#converting the dict to dataframe
df = pd.DataFrame(ProcessedData)
predDf = pd.DataFrame(ProcessedPrediction)
#training the dataset with an appropriate amount of training data.
train,test,train_labels,test_labels = train_test_split(df,
dictFrameTarget,
test_size = 0.45,
random_state=42)
#Creating the model and fitting it to the data
gnb = GaussianNB()
model = gnb.fit(train, train_labels)
#Make a prediction and test accuracy
preds = gnb.predict(predDf)
#changing the numeric representation of the predictions to string representation
predsAns = []
for i in preds:
if i == 0:
predsAns.append("no")
elif i == 1:
predsAns.append("yes")
dataPreds = pd.DataFrame(predsAns)
#Old code for testing accuracy against the left over training set
#preds = gnb.predict(test)
#print(preds)
#print(accuracy_score(test_labels, preds))
#Result: 80%
#concat the two dataframes and write to csv file
compPredictions = pd.concat([dataFrame["id"], dataPreds],axis=1)
compPredictions.to_csv('PredictionFile.csv')
| ML_CA2_Classifier_2019_20.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Quantum Fourier Transform
#
# Takes an input state $|x\rangle$ on the computational basis and takes it to the Fourier basis, which can be done by applying the following circuit $U$.
#
# Let $|x\rangle$ be a all zeros state on $n$ qubits,
#
# $$
# \begin{align*}
# U|x\rangle = \frac{1}{\sqrt{N}}\sum_{y = 0}^{N-1}\exp\bigg(\frac{2\pi i xy}{N}\bigg)|y\rangle
# \end{align*}
# $$
#
# You can find a detailed explanation about the QFT right [here](https://github.com/matheusmtta/Quantum-Computing/blob/master/Algorithms/QFT.ipynb).
# +
import matplotlib.pyplot as plt
# %matplotlib inline
import numpy as np
import math
from qiskit.visualization import plot_state_qsphere
from qiskit import *
from qiskit.tools.visualization import plot_histogram
from IPython.display import display, Math, Latex
# +
def reflect(U, n):
for i in range(int(n/2)):
U.swap(i, n-i-1)
def myQFT(n):
U = QuantumCircuit(n)
for x_k in range(n):
U.h(x_k)
for x_j in range(x_k+1, n):
angle = math.pi/2**(x_j-x_k)
U.cu1(angle, x_j, x_k)
reflect(U, n)
U = U.to_gate()
U.name = 'Quantum Fourier Tranform'
#ctl_U = U.control() make it a controlled gate
return U
# +
n = 5
mc = QuantumCircuit(n, n)
#state |x> = |10101>
mc.x(0)
mc.x(2)
mc.x(4)
#Computational basis state qsphere
backend = BasicAer.get_backend('statevector_simulator')
job0 = execute(mc, backend).result()
U = myQFT(n)
mc.append(U, range(n))
#Fourier basis state qsphere
job1 = execute(mc, backend).result()
mc.measure(range(n), range(n))
backend = BasicAer.get_backend('qasm_simulator')
atp = 1024
res = execute(mc, backend=backend, shots=atp).result()
ans = res.get_counts()
mc.draw('mpl')
# -
plot_histogram(ans)
plot_state_qsphere(job0.get_statevector(mc))
plot_state_qsphere(job1.get_statevector(mc))
| Miscellaneous Quantum Computing Notebooks/QFT Implementation.ipynb |