code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import os
import numpy as np
import theano
import re
os.getcwd()
file='../data/AML/AML_miRNA_Seq3.csv'
patient = dict()
p = re.compile('TCGA-[^-]+-[0-9]+')
with open(file) as f:
record = f.readline().split(',')
while True:
record = f.readline().split(',')
if record[0] == '':
break;
m = p.match(record[0])
patient_id=m.group()
if not patient_id in patient:
patient[patient_id] = dict()
if record[1] != 'sample':
patient[patient_id][record[1]] = float(record[2])
# Number of patients
len(patient.keys())
# Number of genes
for k,v in patient.iteritems():
genes=v.keys()
print(len(genes))
break
patient_list = []
with open('../data/AML/pat_id.txt','r') as f:
f.readline() # skip the header
while True:
record = f.readline().strip()
if record == '':
break;
patient_list.append(record)
file2 = '../data/AML/AML_miRNA_Seq_table2.csv'
with open(file2,'w') as out:
out.write('ID')
for patient_id in patient_list:
out.write('\t')
out.write(patient_id)
out.write('\n')
for gene in genes:
out.write(gene)
for patient_id in patient_list:
out.write('\t')
out.write(str(patient[patient_id][gene]))
out.write('\n')
out.close()
| notebooks/preprocess_AML_miRNA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] hideCode=true hidePrompt=true
# # Chapter 8: Advanced Data Analysis
# In the last few chapters, we have become comfortable with the idea of building our own functions. These can become quite complex, as we have learned with the construction of the OLS regression in chapter 7. We will continue developing our tool set for working with and managing large sets of data by integrating data from different data sets. We will introduce the multi index to facilitate this process. Among other things, the multi index is useful for including identifiers across time and region or principality. We will also use the multi index to perform a panel regression that controls for level effects between different countries.
#
# ## Using a Double Index to Work with Panel Data
# In any project, it will not be uncommon for data to be attached to more than one indentifying category. Often, data will be labeled by polity and by date. In the next several examples, we will work with multiple data sets of this sort, working to combine different data sets, investigate the features of the double index, and use this data in a panel regression that can control for effects by polity and by time period.
#
# ### Plotting with Double Index
# We will be working with two datasets in the next example: the [Fraser Economic Freedom Index](https://www.fraserinstitute.org/economic-freedom/dataset?geozone=world&page=dataset&min-year=2&max-year=0&filter=0&year=2017) and GDP from the [Maddison Project](https://www.rug.nl/ggdc/historicaldevelopment/maddison/releases/maddison-project-database-2018).
#
# Due to the formatting of the Fraser Economic Freedom Index, the first column and the first three rows of data in the sheet titled “EFW Index 2018 Report” are blank. We will account for this when importing the data with pandas. Both should be saved in the same folder as the script below.
#
# First, import the GDP data from Maddison Project:
# + hideCode=true hidePrompt=true
#multiIndex.py
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# index_col = [0,2] will select countrycode as the primary index and year as
# the secondary index
data = pd.read_excel("mpd2018.xlsx",
sheet_name = "Full data",
index_col = [0,2])
# + [markdown] hideCode=true hidePrompt=true
# View the new dataframe by entering data in the console:
# + hideCode=true hidePrompt=true
data
# + [markdown] hideCode=true hidePrompt=true
# When working with a multi index, calling the values from a single index requires a few steps. If you were to call data.index in the console, both the countrycode and the year values would be returned:
# + hideCode=true hidePrompt=true
data.index
# + [markdown] hideCode=true hidePrompt=true
# To call only the year values from the multi index, we use the dataframe method, *.get_level_values(“year”)*. This returns the same list, but with only years:
# + hideCode=true hidePrompt=true
data.index.get_level_values("year")
# + [markdown] hideCode=true hidePrompt=true
# Since we don’t need to hold every repeated year value, only the full range years present in the data set. We will remove an repeated values from the list and then ensure that the list is in order:
# + hideCode=true hidePrompt=true
#multiIndex.py
# . . .
# to gather all of the years, we group observations by year,
# which will remove the other column of the multiindex
years = data.groupby("year").mean().index
# + [markdown] hideCode=true hidePrompt=true
# Once you understand the structure of the index, you may find it easier to call all of the commands in one line:
# + hideCode=true hidePrompt=true
years
# + [markdown] hideCode=true hidePrompt=true
# The data set has relatively few observations for data before the modern era. Before 1800, even for the years that have entries, data for most countries is not present.
#
# Next, we create a dictionary with tuples containing codes for each pair of countries whose real GDP per capita we will compare. The first pair is Canada and Finland, the second pair is France and Germany, and the third pair is Great Britain and the Netherlands. The first country in each pair will be represented by the linestyle *“-”* and the second pair by the linestyle *“--”*.
# + hideCode=true hidePrompt=true
#multiIndex.py
# . . .
# pairs of countries to compare in plots
pairs = [("CAN", "FIN"), ("FRA", "DEU"), ("USA","GBR", "ESP", "MEX")]
linestyles = ["-", ":","--","-."]
# + [markdown] hideCode=true hidePrompt=true
# Now that the dictionary has been prepared, cycle through each pair of countries in the dictionary. Using a for loop, we select one of these at a time with an accompanying linestyle. The code of the country calls the Real GDP data using the command *data.ix[(country),:]["cgdppc"]*. This selects data by index according to country and includes all years, as is indicated by the colon in the second part of the index entry. Last, the column *“cgdppc”* is selected. Conveniently, we can also use the code stored in country as the label.
#
# Script in lines 27-30 continue to adjust the plot. The command *plt.xlim(xmin, xmax)* selects the range of the x-axis. Only data from 1825 to the most recent observation, identified by *max(years)*, is included in the plot. The *plt.rcParams.update()* commands adjust fontsizes by accessing default parameter in matplotlib. Finally, the create a title that uses latex text by including “$text$”. This italicizes the title text and allows for the use latex commands such as subscripting, greek letters, etc....
# + hideCode=true hidePrompt=true
#multiIndex.py
# . . .
for pair in pairs:
fig, ax = plt.subplots(figsize=(16,8))
for i in range(len(pair)):
country = pair[i]
linestyle = linestyles[i]
data.loc[country,:]["cgdppc"].dropna(
).plot.line(ax = ax,
label = country, linestyle = linestyle)
plt.xlim([1825, max(years)])
plt.rcParams.update({"legend.fontsize": 25, "legend.handlelength": 2})
plt.rcParams.update({"font.size": 25})
plt.ylabel(
"$Real$ $GDP$ $Per$ $Capita$\n2011 U.S. Dollars",
fontsize=36)
plt.legend()
plt.show()
plt.close()
# + [markdown] hideCode=true hidePrompt=true
# ### Merge Data Sets with Double Index
# Next, we will import both sets of data as unique objects with the intention of combining them. First, we must successful import the Fraser Index. Unlike with other files we have imported, the column names are in the third row and the years are in the second column. Finally, we will call the *.dropna()* method twice to drop any row *(axis = 0)* and then any column *(axis=1)* with missing observations:
# + hideCode=true hidePrompt=true
#dataForPanel.py
import pandas as pd
#make sure dates are imported in the same format; to do this,
#we turned off parse_dates
fraser_data = pd.read_csv("cleanedEconFreedomData.csv",
index_col = [0, 1], parse_dates=False)
# drop any empty column and any empty row
fraser_data = fraser_data.dropna(
axis=0, thresh=1).dropna(axis=1, thresh=1)
maddison_data = pd.read_excel("mpd2018.xlsx", sheet_name = "Full data",
index_col = [0,2])#, parse_dates = True)
# + [markdown] hideCode=true hidePrompt=true
# The object fraserData is imported from the sheet #“EFW Index 2018 Report”#. The 1st and 0th columns are used as index columns in that order, respectively. The object maddisonData is imported from the sheet #“Full data”#. The 0th and 2nd columns columns are used as index columns. The double index of both dataframes refers to the #ISO_Code# (#countrycode#) and the year. Since the objects referred to by the indices match, we will be able to use the double index associated with a particular entry to refer to another double index.
#
# Since both dataframes employ the same double index format, we can copy any column from one dataframe to the other. We will copy for the Maddison GDP data to the Fraser Economic Freedom Index data.
# + hideCode=true hidePrompt=true
#dataForPanel.py
# . . .
fraser_data["RGDP Per Capita"] = maddison_data["cgdppc"]
fraser_data
# + [markdown] hideCode=true hidePrompt=true
# Now that the dataset has been transferred, we can save the result as a csv so that we can call it again in later examples. RGDP Per Capita is shown in the last column of the new csv file:
# + hideCode=true hidePrompt=true
fraser_data.to_csv("fraserDataWithRGDPPC.csv")
# + [markdown] hideCode=true hidePrompt=true
# ### Creating Indicator Variables
# Suppose that we wanted to test the idea that geography influences economic growth. We would need to clarify a hypothesis concerning this. We might believe, for example, that countries in North America tend to have a distinct real gross domestic product than in other continents i.e., real GDP tends to be higher or lower due to residing in North America. To represent this, we would create an indicator variable named *“North America”*. Countries residing in North America would be indicated with a 1 (i.e., True), and those outside of North America would receive a zero.
#
# To accomplish this task is straightforward if you know the appropriate commands to use. As usual, we import the data. Before creating an indicator variable, you will need to choose the name that will reference the indicator variable, indicator_name, and make a list of the index values, *target_index_list* that will be recorded as possessing the attribute referred to by the indicator variable. Finally, you will need to choose the name of the index column that includes the elements in the *target_index_list*. If you are not sure what this name is, you can check the names of the index columns using *df.index.names*.
# + hideCode=true hidePrompt=true
#indicatorVariable.py
import pandas as pd
def create_indicator_variable(data, indicator_name,
index_name, target_index_list):
# Prepare column with name of indicator variable
data[indicator_name] = 0
# for each index whose name matches an entry in target_index_list
# a value of 1 will be recorded
for index in target_index_list:
data.loc[data.index.get_level_values(\
index_name) == index, [indicator_name]] = 1
# Import data with "ISO_Code" and "Year" as index columns
data = pd.read_csv("fraserDataWithRGDPPC.csv",
index_col = ["ISO_Code", "Year"],
parse_dates = True)
# select "ISO_Code" from names of double index
index_name = data.index.names[0]
indicator_name = "North America"
# Cuba, Grenada, Saint Kitts, Saint Lucia, Saint Vincent are missing
# from Fraser Data
countries_in_north_america = [
"BHS", "BRB", "BLZ", "CAN", "CRI", "DOM", "SLV",
"GTM", "HTI", "HND", "JAM", "MEX", "NIC", "PAN",
"TTO", "USA"]
create_indicator_variable(data = data,
indicator_name = indicator_name,
index_name = index_name,
target_index_list = countries_in_north_america)
# + [markdown] hideCode=true hidePrompt=true
# Once the new column of data has been creative, it can be called by passing indicator¬¬_name to the dataframe. Be sure to use double brackets (i.e., *[[ , ]]*) so that the column names are included in the output.
# + hideCode=true hidePrompt=true
data[["North America"]]
# -
# It is useful to represent discrete variabls, like the indicator variable for "North America", by color in a scatter plot. If there is a difference means between two groups, it is often apparrent when data is visualized by color. In the case of the "North America" indicator variable, the difference in means between the groups does not appear to be causal (statistically significant), as we will show later in the chapter.
#
# We use "North America" as the variable indicating color in the scatter plot.
# +
import datetime
year = 2000
plot_data = data[data.index.get_level_values("Year")== datetime.datetime(year,1,1)]
fig, ax = plt.subplots(figsize = (24, 16))
plot_data.plot.scatter(x = "EFW", y = "RGDP Per Capita", c = "North America",
cmap = "coolwarm", ax = ax, s = 50)
ax.set_title(str(year), fontsize = 50)
# +
import datetime
from matplotlib import cm
year = 2000
# change colors, divide into 4 distinct colors
norm = cm.colors.Normalize()
cmap = cm.get_cmap('coolwarm', 2)
plt.cm.ScalarMappable(cmap=cmap, norm=norm)
plot_data = data[data.index.get_level_values("Year")== datetime.datetime(year,1,1)]
fig, ax = plt.subplots(figsize = (24, 16))
plot_data.plot.scatter(x = "EFW", y = "RGDP Per Capita", c = "North America",
cmap = cmap, ax = ax, norm = norm, s = 50)
# to remove numbers between 0 and 1, access the color axis through plt.gcf()
f = plt.gcf()
cax = f.get_axes()[1]
# access colorbar values
vals = cax.get_yticks()
print(vals)
# only include 0 or 1
cax.set_yticklabels([int(val) if val % 1 == 0 else "" for val in vals ])
# remove tick lines from color axis
cax.tick_params(length = 0)
ax.set_title(str(year), fontsize = 50)
# + [markdown] hideCode=true hidePrompt=true
# ### Create Quantile Ranking
# Similar to the pervious exercise, we may categorize data according ranked bins. This is accomplished by separating data into quantiles, often in the form of quartiles or quintiles, however this can be accomplished using any number of divisions. The following script allows you to create quantiles of the number of divisions of your choosing.
#
# First, we need to import the data that we have merged. Once the dataframe is created, we must prepare a place for quantile data to be registered. Out of convenience, we will refer to the n-tile, meaning that a quartile will be labeled a “4-tile”, a quintile will be label a “5-tile”, and so forth. Before recording the data, we will create blank entries using *np.nan*.
# + hideCode=true hidePrompt=true
#quantile.py
import pandas as pd
import numpy as np
# choose numbers of divisions
n = 5
# import data
data = pd.read_csv("fraserDataWithRGDPPC.csv",
index_col = ["ISO_Code", "Year"],
parse_dates = True)
#create column identifying n-tile rank
quantile_var = "RGDP Per Capita"
quantile_name = quantile_var + " " + str(n) + "-tile"
data[quantile_name] = np.nan
# + [markdown] hideCode=true hidePrompt=true
# Now that the dataframe has been prepped, we can create function that will record quantile values. Since the data frame includes data for nearly every country over many years, we will want to choose one of the index categories to identify quantiles. We prefer to compare countries, 8so unique quantile measures will compare countries in a given year. We will build a list of years and use a for-loop to cycle through values in the list. For each year, we will construct a quantile values comparing countries within the year.
#
# The *create_quantile()* function needs to be provided the number of divisions within a quantile (*n*), the dataframe (*data*), the year to which the quantile measure refers (*year*), the variable for which the quantile is constructed (*quantile_var*), and the key that will refer to the quantile data (*quantile_name*).
#
# First, we construct the year_index. This provides a slice of the original index that includes only the years of interest. Next, we identify the value that divides each quantile. The pandas dataframe has a convenient command, *df.quantile(percent)*, that will calculate the value for a particular percentile. For example, *df.quantile(.25)* will calculate the value representing the 25th percentile. A quantile is comprised of divisions whose size is a fraction of 1 and that sum to one. The value that divides each quantile is defined by *i/n* where *n* is the number of quartiles, and
# *i* includes all integers from 1 up to and including *n*.
#
# Now that the values dividing each quantile for a given year have been identified, we can check which in which quantile each country falls. Cycle through the index for the year with for index in *data[year_index].index*. This allows us to calls up each individual countries data for the given year, compare that data of the category of interest to the quantile values, and select the identify quantile that the nation falls in for the year.
# + hideCode=true hidePrompt=true
#quantile.py
import pandas as pd
import numpy as np
def create_quantile(n, data, year, quantile_var, quantile_name):
# index that indentifies countries for a given year
year_index = data.index.get_level_values("Year") == year
quantile_values_dict = {i:data[year_index][quantile_var]\
.quantile(i/n) for i in range(1, n + 1)}
# cycle through each country for a given year
for index in data[year_index].index:
# identtify value of the variable of interest
val = data.loc[index][quantile_var]
# compare that value to the values that divide each quantile
for i in range(1, n + 1):
# if the value is less than the highest in the quantile identified,
# save quantile as i
if val <= quantile_values_dict[i]:
data.loc[index,[quantile_name]]=int((n + 1) - i)
#exit loop
break
# otherwise check the higest value of the next quantile
else:
continue
# . . .
years = data.groupby("Year").mean().index
for year in years:
create_quantile(n, data, year, quantile_var, quantile_name)
# + [markdown] hideCode=true hidePrompt=true
# Now call "RGDP Per Capita" alongside the quantile rankings.
# + hideCode=true hidePrompt=true
data[["RGDP Per Capita", "RGDP Per Capita 5-tile"]]
# -
# Again, let's identify each quantile by color. We will use code slightly different from the last time. Since the values cover a greater range, all we need to do is shift the min and max values of the color axis.
# +
from matplotlib import cm
from mpl_toolkits.axes_grid1 import make_axes_locatable
year = 2016
plot_data = data[data.index.get_level_values("Year")== datetime.datetime(year,1,1)]
fig, ax = plt.subplots(figsize = (24, 16))
# change colors, divide into 4 distinct colors
norm = cm.colors.Normalize()
cmap = cm.get_cmap('jet', n)
plt.cm.ScalarMappable(cmap=cmap, norm=norm)
# set vmin to 0.5 to slight shift scale, otherwise quintile
# measures will be on border between colors and 1 and 2 will be
# indicated in lightblue
plot_data.plot.scatter(x = "EFW", y = "RGDP Per Capita",
c = "RGDP Per Capita " + str(n) + "-tile",
cmap = cmap, norm = norm, ax = ax, s = 50,
legend = False, vmin = 0.5, vmax = 5.5)
ax.set_title("Year: " + str(year), fontsize = 50)
# + [markdown] hideCode=true hidePrompt=true
# ### Lag Variables and Differenced Log Values
#
# With time series data, it is often useful to control for trends when data is autocorrelated. Consider, for example, that real GDP data is often highly correlated with values from the previous period. We might detect a false causal relationship between two variables that are actually unrelated but follow a similar trend. For example, we might regress your age against real GDP and find that there is a strong correlation between the two. To avaoid false positives like this, it is useful to account for the influence of lagged values and/or to detrend the data all together by using differenced logs.
#
# Creating lag variables is quite simple if the index is already recognized as containing dates and times. The method, *df.shift(n)* accomplishes this. Pass a negative value to create a lagged variable from n periods previous and a positive value to create a variable that refers to data n periods in the future.
#
# Because we are using a double index, we must specify to which index we must instruct Pandas as to which index the shift refers. We accomplish this by using *.groupby(level)* to target the index column that does not refer to a datetime data. This will group the data by entity, thus leaving only the date column to be referenced by *.shift(n)*. Since we only want a lag value from the period that immediately preceded the observation, *n=-1*.
# + hideCode=true hidePrompt=true
#logAndDifferenceData.py
import pandas as pd
import numpy as np
import datetime
# import data
data = pd.read_csv("fraserDataWithRGDPPC.csv", index_col = ["ISO_Code", "Year"],
parse_dates = True)
data["RGDP Per Capita Lag"] = data.groupby(level="ISO_Code")\
["RGDP Per Capita"].shift(-1)
# + [markdown] hideCode=true hidePrompt=true
# To understand what the command yielded, we will want to view data for an individual country. The following command will save *“RGDP Per Capita”* in the United States and the lag of that value to a csv file:
# + hideCode=true hidePrompt=true
data.loc["USA", :]
# + [markdown] hideCode=true hidePrompt=true
# Since not every year is included in the index, this actually results in false values for periods where observations are only available once every 5 years. If we use lagged values, we need to delimit the data to consecutive annual observations. Since consistent data is provided starting in the year 2000. To select data by year, we need to inicate that we wish to form selection criteria that refers to values form the year column in the index. The command *data.index.get_level_Values(“Year”)* calls these values. Once we execute the above script, we can call this command in the console.
# + hideCode=true hidePrompt=true
data.index.get_level_values("Year")
# + [markdown] hideCode=true hidePrompt=true
# If we compare these values to the critierion specified, a column of boolean values will be generated that identifies where values from the index meet the specified criterion.
# + hideCode=true hidePrompt=true
data.index.get_level_values("Year") >= datetime.datetime(1999, 1, 1)
# + [markdown] hideCode=true hidePrompt=true
# If we pass the command, *data.index.get_level_values("Year") > datetime.datetime(1999,1,1)*, to the dataframe, only observations that meet the criterion indicated will be included. In this case, any observation generated in years after 1999 will be included.
# + hideCode=true hidePrompt=true
#logAndDifferenceData.py
# . . .
data = data[data.index.get_level_values("Year") > datetime.datetime(1999,1,1)]
# + [markdown] hideCode=true hidePrompt=true
# Call *data* in the console to see that the range of dates included has narrowed, thereby removing the observations separated by 5 year intervals:
# + hideCode=true hidePrompt=true
data
# + [markdown] hideCode=true hidePrompt=true
# The earliest dates included nowonly reach as far back as the year 2001.
#
# Now that we’ve delimited the data, let’s perform some other transformation that will help to prepare the data for a regression. It will be useful to log GDP values as the rate of growth of GDP measured in raw units tends to increase over time. By logging data, beta estimates will approximate the cross elasticity of the endogenous variable y with respect to a change in an exogenous variable x.
#
# Since we have already created a lagged measure of real GDP, this is a good opportunitiy to use a for loop. We only want to log values referred to be a key that includes the string *“GDP”*. We check each key for inclusion of this term. If the term is included in the key, than we add a logged version of the variable.
#
# + hideCode=true hidePrompt=true
#logAndDifferenceData.py
# . . .
for key in data:
if "GDP" in key:
data["Log " + key] = np.log(data[key])
# + [markdown] hideCode=true hidePrompt=true
# The data is almost prepared. We have only left to create data that has been diferenced. Differenced log values approximate the rate of change of a variable. Likewise, we can difference the index values to test for the effect of an improvement in a nation’s economic freedom score on the rate of GDP growth.
#
# To difference data with a multi index, we must first organize the data so that observations are presented sequentially by entity. This uses the same command, .groupby(), that we used earlier to create a lag value. The result this time is that a new dataframe is created by taking the differenced values of all variables in the dataframe.
#
# We save the new results in a dictionary that holds both the original dataframe and the new dataframe with differenced data.
#
# Since it does not make sense to difference an indicator variable, we will reset the value of North America to 1.
# + hideCode=true hidePrompt=true
#logAndDifferenceData.py
# We do not want to difference the index values, only the Real GDP values
# so initialize the diff data as teh dataframe but only include index values
# from a differenced matrix (one year of observations will be missing)
diff_index = data.groupby(level=0).diff(-1).dropna().index
data_dict = {}
data_dict["Data"] = data
data_dict["Diff Data"] = data.copy().loc[diff_index]
data_dict["Diff Data"] = data.groupby(level=0).diff(-1)
data_dict["Diff Data"]["North America"] = 1
data_dict["Diff Data"] = data_dict["Diff Data"].dropna()
# + [markdown] hideCode=true hidePrompt=true
# Now let's check the data that we saved as data_dict["Diff Data"]
# + hideCode=true hidePrompt=true
data_dict["Data"]
# -
# Notice that there exist no data for the first year of observations. Without a previous year to draw from for the year 1999, the pandas dataframe is left blank for differenced values in this year.
# + [markdown] hideCode=true hidePrompt=true
# ### Using Indicator Variables in Regression
# Using the function created above, we can prepare indicator and quantile variables to be used in a regression. As before, the a regression follows the same form as a standard multiple regression with continuous exogenous variables, but also include a parameter, α_k, for each indicator or quantile variables.
#
# $y_i = \beta_0 + \sum_{j=1}^{n} \beta_jx_{j,i} + \sum_{k=1}^{m} \alpha_kx_{k,i} + e_i$
#
# For both cases, we can use the regression method that we created last class. We begin with use of indicator variables in regression. An indicator variable accounts for a level effect that is attributed to a predicted value due to an attribute of the observation. Suppose that we wanted to measure whether or not there is an effect of gender on wages in a particular field once all other relevant factors are accounted for. Observations where the wage earner is a woman would be indicated with a one. The beta value estimated by a regression would indicate the effect of gender on wage given the control variables included in the regression.
#
# We can perform a similar regression by identifying the effect of being a nation in North America on real GDP per capita of countries residing on the continent. In reality, this is actually a poor indicator variable, but we can expand our computational toolbelt by creating an indicator variable that identifies countries in North America and by including this variable in a regression. We will see that the creation of indicator variables is fundamental to the panel regression.
# + hideCode=true hidePrompt=true
#indicatorAndDiffRegression.py
import pandas as pd
import numpy as np
import datetime
def create_indicator_variable(data, indicator_name, index_name,
target_index_list):
# Prepare column with name of indicator variable
data[indicator_name] = 0
# for each index whose name matches an entry in target_index_list
# a value of 1 will be recorded
for index in target_index_list:
data.loc[data.index.get_level_values(\
index_name) == index, [indicator_name]] = 1
# import data
data = pd.read_csv("fraserDataWithRGDPPC.csv", index_col = ["ISO_Code", "Year"],
parse_dates = True)
data["RGDP Per Capita Lag"] = data.groupby(level="ISO_Code")\
["RGDP Per Capita"].shift(-1)
data = data[data.index.get_level_values("Year") > datetime.datetime(2000,1,1)]
for key in data:
if "GDP" in key:
data["Log " + key] = np.log(data[key])
# We do not want to difference the index values, only the Real GDP values
# so initialize the diff data as teh dataframe but only include index values
# from a differenced matrix (one year of observations will be missing)
diff_index = data.groupby(level=0).diff(-1).dropna().index
data_dict = {}
data_dict["Data"] = data
data_dict["Diff Data"] = data.copy().loc[diff_index]
for key in data:
if "GDP" in key:
data_dict["Diff Data"][key] = data[key].groupby(level=0).diff(-1)
data_dict["Diff Data"] = data_dict["Diff Data"].dropna()
# Create indicator variable for North Amierca in both data and diff_data
indicator_name = "North America"
index_name = "ISO_Code"
countries_in_north_america = ["BHS", "BRB", "BLZ", "CAN", "CRI", "DOM", "SLV",
"GTM", "HTI", "HND", "JAM", "MEX", "NIC", "PAN",
"TTO", "USA"]
for key in data_dict:
data = data_dict[key]
create_indicator_variable(data = data, indicator_name = indicator_name,
index_name = index_name,target_index_list = countries_in_north_america)
# + [markdown] hideCode=true hidePrompt=true
# As in the earlier example, we have created indicator variables. Notice that the created are not differenced even in the dataframe with differenced data. We will be using these values in regressions where the estimated value is either logged or log-differenced. In the first case, the indicator variable will effect the level estimated. In the second case, the indicator variable influences an estimated rate.
#
# We’ve left to estimate regression using data from each dataframe. We will estimate the impact of economic freedom (*“SUMMARY INDEX”*) on Real GDP Per Capita. To control for autocorrelation, we include the lag of Real GDP Per Capita. To run the regression once for each dataframe in data_dict, we use a for loop that cycles through the keys in data_dict. We print the statistics for the estimated beta values as well as statistics that describe the results of the regression overall. The key is also printed to identify each set of results.
# -
data_dict[key]
# +
#indicatorRegressoin.py
# . . .
from regression import Regression
# . . .
# prepare regression variables
X_names = ["EFW", "Log RGDP Per Capita Lag"]
y_name = ["Log RGDP Per Capita"]
# save instance of regression class
reg = Regression()
for key in data_dict:
# call OLS method
data = data_dict[key]
reg.OLS(reg_name = key, data = data.dropna(),
y_name = y_name, beta_names = X_names)
print(key, reg.estimates, sep = "\n")
print(reg.stats_DF)
print()
# + [markdown] hideCode=true hidePrompt=true
# We can check if the indicator variable, *“North America”*, adds any explanatory value by adding the variable to this regression. Only line 43 is changed by this addition.
# -
# . . .
X_names = ["EFW", "Log RGDP Per Capita Lag", "North America"]
# . . .
for key in data_dict:
# call OLS method
data = data_dict[key]
reg.OLS(reg_name = key, data = data.dropna(),
y_name = y_name, beta_names = X_names)
print(key, reg.estimates, sep = "\n")
print(reg.stats_DF)
print()
# Neither of regression yield an estimate for the indicator variable that is 1) statistically significant or 2) that significantly improves the goodness-of-fit (r\*\*2). In fact, the f-statistic and adjusted r-squared values have both fallen for the second set of regressions. It appears that inclusion of a control for North American countries does not improve the regression.
#
# ### Panel Regression
# The indicator variable plays a key role in a very popular regression within economics: the panel (or fixed effects) regression. A panel regression is an OLS regression that includes an indicator variable for certain fixed attributes. Conventiently, the panel regression is often used to control for effects between different political units – i.e., cities, states, nations, etc.... Using the data from the previous examples, we will run a panel regression with fixed effects for each nation. We regress the data over time, holding constant a level effect provided by the unique indicator variable associated with each nation. The indicator variables that drive results in the panel regression adjust the y-intercept indicated by the constant β0 with a unique adjustment for each state.
#
# We can accommodate a Panel Regression by making a few additions and edits to the *regress()* method that we have already built. The first step for creating a panel regression will be to add the *create_indicator_variable()* function that we created earlier. Be sure to add self to the terms that are passed to this method within the *Regression* class.
#
# For convenience, the entire Regression class is included with an additional
# +
#regression.py
# you may ignore import jdc, used to split class development
# other cells that edits a class will include the magic command %% add_to
import jdc
import pandas as pd
import numpy as np
import copy
# make sure to import sys, as we will use this in the next step
import sys
import stats
from scipy.stats import t, f
class Regression:
def __init__(self):
self.reg_history = {}
def OLS(self, reg_name, data, y_name, beta_names, min_val = 0,
max_val = None, constant = True):
self.min_val = min_val
if max_val != None:
self.max_val = max_val
else:
self.max_val = len(data)
self.reg_name = reg_name
self.y_name = y_name
self.beta_names = copy.copy(beta_names)
self.data = data.copy()
if constant:
self.add_constant()
self.build_matrices()
self.estimate_betas_and_yhat()
self.calculate_regression_stats()
self.save_output()
def calculate_regression_stats(self):
self.sum_square_stats()
self.calculate_degrees_of_freedom()
self.calculate_estimator_variance()
self.calculate_covariance_matrix()
self.calculate_t_p_error_stats()
self.calculate_root_MSE()
self.calculate_rsquared()
self.calculate_fstat()
self.build_stats_DF()
def add_constant(self):
self.data["Constant"] = 1
self.beta_names.append("Constant")
def build_matrices(self):
# Transform dataframews to matrices
self.y = np.matrix(self.data[self.y_name][self.min_val:self.max_val])
# create a k X n nested list containing vectors for each exogenous var
self.X = np.matrix(self.data[self.beta_names])
self.X_transpose = np.matrix(self.X).getT()
# (X'X)**-1
X_transp_X = np.matmul(self.X_transpose, self.X)
self.X_transp_X_inv = X_transp_X.getI()
# X'y
self.X_transp_y = np.matmul(self.X_transpose, self.y)
def estimate_betas_and_yhat(self):
# betas = (X'X)**-1 * X'y
self.betas = np.matmul(self.X_transp_X_inv, self.X_transp_y)
# y_hat = X * betas
self.y_hat = np.matmul(self.X, self.betas)
# Create a column that hold y-hat values
#.item(n) pulls nth value from matrix
self.data[self.y_name[0] + " estimator"] = \
[i.item(0) for i in self.y_hat]
# create a table that holds the estimated coefficient
# this will also be used to store SEs, t-stats, and p-values
self.estimates = pd.DataFrame(self.betas, index = self.beta_names,
columns = ["Coefficient"])
# identify y variable in index
self.estimates.index.name = "y = " + self.y_name[0]
def sum_square_stats(self):
ssr_list = []
sse_list = []
sst_list = []
mean_y = stats.mean(self.y).item(0)
for i in range(len(self.y)):
# ssr is sum of squared distances between the estimated y values
# (y-hat) and the average of y values (y-bar)
yhat_i = self.y_hat[i]
y_i = self.y[i]
r = yhat_i - mean_y
e = y_i - yhat_i
t = y_i - mean_y
ssr_list.append((r) ** 2)
sse_list.append((e) ** 2)
sst_list.append((t) ** 2)
# call item - call value instead of matrix
self.ssr = stats.total(ssr_list).item(0)
self.sse = stats.total(sse_list).item(0)
self.sst = stats.total(sst_list).item(0)
def calculate_degrees_of_freedom(self):
# Degrees of freedom compares the number of observations to the number
# of exogenous variables used to form the prediction
self.lost_degrees_of_freedom = len(self.estimates)
self.num_obs = self.max_val + 1 - self.min_val
self.degrees_of_freedom = self.num_obs - self.lost_degrees_of_freedom
def calculate_estimator_variance(self):
# estimator variance is the sse normalized by the degrees of freedom
# thus, estimator variance increases as the number of exogenous
# variables used in estimation increases(i.e., as degrees of freedom
# fall)
self.estimator_variance = self.sse / self.degrees_of_freedom
def calculate_covariance_matrix(self):
# Covariance matrix will be used to estimate standard errors for
# each coefficient.
# estimator variance * (X'X)**-1
self.cov_matrix = float(self.estimator_variance) * self.X_transp_X_inv
self.cov_matrix = pd.DataFrame(self.cov_matrix,
columns = self.beta_names,
index = self.beta_names)
def calculate_t_p_error_stats(self):
ratings = [.05, .01, .001]
results = self.estimates
stat_sig_names = ["SE", "t-stat", "p-value"]
# create space in data frame for SE, t, and p
for stat_name in stat_sig_names:
results[stat_name] = np.nan
# generate statistic for each variable
for var in self.beta_names:
# SE ** 2 of coefficient is found in the diagonal of cov_matrix
results.loc[var, "SE"] = self.cov_matrix[var][var] ** (1/2)
# t-stat = Coef / SE
results.loc[var,"t-stat"] = \
results.loc[var, "Coefficient"] / results.loc[var, "SE"]
# p-values is estimated using a table that transforms t-value in
# light of degrees of freedom
results.loc[var, "p-value"] = np.round(t.sf(np.abs(results.\
loc[var, "t-stat"]), self.degrees_of_freedom + 1) * 2, 5)
# values for significances will be blank unless p-values < .05
# pandas does not allow np.nan values or default blank strings to
# be replaced
significance = ["" for i in range(len(self.beta_names))]
for i in range(len(self.beta_names)):
var = self.beta_names[i]
for val in ratings:
if results.loc[var]["p-value"] < val:
significance[i] = significance[i] + "*"
results["signficance"] = significance
def calculate_root_MSE(self):
self.root_mse = self.estimator_variance ** (1/2)
def calculate_rsquared(self):
self.r_sq = self.ssr / self.sst
self.adj_r_sq = 1 - self.sse / self.degrees_of_freedom / (self.sst\
/ (self.num_obs - 1))
def calculate_fstat(self):
self.f_stat = (self.sst - self.sse) / (self.lost_degrees_of_freedom\
- 1) / self.estimator_variance
def build_stats_DF(self):
stats_dict = {"r**2":[self.r_sq],
"Adj. r**2":[self.adj_r_sq],
"f-stat":[self.f_stat],
"Est Var":[self.estimator_variance],
"rootMSE":[self.root_mse],
"SSE":[self.sse],
"SSR":[self.ssr],
"SST":[self.sst],
"Obs.":[int(self.num_obs)],
"DOF":[int(self.degrees_of_freedom)]}
self.stats_DF = pd.DataFrame(stats_dict)
self.stats_DF = self.stats_DF.rename(index={0:"Estimation Statistics"})
self.stats_DF = self.stats_DF.T
def save_output(self):
self.reg_history[self.reg_name] = {"Reg Stats": self.stats_DF.copy(),
"Estimates": self.estimates.copy(),
"Cov Matrix":self.cov_matrix.copy(),
"Data":self.data.copy()}
def joint_f_test(self, reg1_name, reg2_name):
# identify data for each regression
reg1 = self.reg_history[reg1_name]
reg2 = self.reg_history[reg2_name]
# identify beta estimates for each regression to draw variables
reg1_estimates = reg1["Estimates"]
reg2_estimates = reg2["Estimates"]
# name of y_var is saved as estimates index name
reg1_y_name = reg1_estimates.index.name
reg2_y_name = reg2_estimates.index.name
num_obs1 = reg1["Reg Stats"].loc["Obs."][0]
num_obs2 = reg2["Reg Stats"].loc["Obs."][0]
# check that the f-stat is measuring restriction, not for diff data sets
if num_obs1 != num_obs2:
self.joint_f_error()
if reg1_y_name == reg2_y_name:
restr_reg = reg1 if \
len(reg1_estimates.index) < len(reg2_estimates.index) else reg2
unrestr_reg = reg2 if restr_reg is reg1 else reg1
restr_var_names = restr_reg["Estimates"].index
unrestr_var_names = unrestr_reg["Estimates"].index
# identify statistics for each regression
restr_reg = restr_reg if False not in \
[key in unrestr_var_names for key in restr_var_names] else None
if restr_reg == None:
self.joint_f_error()
else:
sser = restr_reg["Reg Stats"].loc["SSE"][0]
sseu = unrestr_reg["Reg Stats"].loc["SSE"][0]
dofr = restr_reg["Reg Stats"].loc["DOF"][0]
dofu = unrestr_reg["Reg Stats"].loc["DOF"][0]
dfn = dofr - dofu
dfd = dofu - 1
f_stat = ((sser - sseu) / (dfn)) / (sseu / (dfd))
f_crit_val = 1 - f.cdf(f_stat,dfn = dfn, dfd = dfd)
#make dictionary?
f_test_label = "h_0:"
for key in unrestr_var_names:
if key not in restr_var_names:
f_test_label = f_test_label + str(key) + " == "
f_test_label = f_test_label + "0"
res_dict = {"f-stat":[f_stat],
"p-value":[f_crit_val],
"dfn":[dfn],
"dfd":[dfd]}
res_DF = pd.DataFrame(res_dict)
res_DF = res_DF.rename(index={0:""})
res_DF = res_DF.T
res_DF.index.name = f_test_label
return res_DF
def joint_f_error(self):
print("Regressions not comparable for joint F-test")
return None
def create_indicator_variable(self,data, indicator_name, index_name,
target_index_list):
# Prepare column with name of indicator variable
data[indicator_name] = 0
# for each index whose name matches an entry in target_index_list
# a value of 1 will be recorded
for index in target_index_list:
data.loc[data.index.get_level_values(\
index_name) == index, indicator_name] = 1
# -
# We will use the *create_indicator_variable()* method to create an indicator variable for every unique id in the index column labeled *“ISO_Codes”*. Each unique ISO Code represents a particular country, thus we will be creating one indicator variable for every country.
#
# Next, we include the script for the panel regression method in its entirety.
#
# +
# %%add_to Regression
def panel_regression(self, reg_name, data, y_name, X_names, min_val = 0,
max_val = None, entity = False, time = False,
constant = True):
self.indicator_lists = {}
#identify which index column holds dates, which holds entities
for i in range(len(data.index.levels)):
if isinstance(data.index.levels[i], pd.DatetimeIndex):
if time:
date_level = i
date_index_name = data.index.names[date_level]
self.indicator_lists[date_index_name] = list(data.groupby(date_index_name).mean().index)
self.indicator_lists[date_index_name].pop()
else:
if entity:
entity_level = i
entity_index_name = data.index.names[entity_level]
self.indicator_lists[entity_index_name] = list(data.groupby(entity_index_name).mean().index)
self.indicator_lists[entity_index_name].pop()
self.indicator_names = []
for index_name, lst in self.indicator_lists.items():
for indicator in lst:
self.create_indicator_variable(data, indicator, index_name,
[indicator])
self.indicator_names = self.indicator_names + lst
X_and_indicator_names = X_names + self.indicator_names
self.OLS(reg_name, data = data, y_name = y_name,
beta_names = X_and_indicator_names, min_val = min_val,
max_val = max_val, constant = constant)
self.X_names = X_names + ["Constant"]
self.data = self.data[self.X_names]
self.estimates = self.estimates.loc[self.X_names]
# -
# Since the index includes includes both *“Year”* and *“ISO_Code”*, we must select which type of fixed effect the regression will employ. We create the regression to allow for entity or time fixed effects in a single regression.
#
# Next, the panel_regression() method must determine the index to which it will refer for creating indicator variables. First, we identify which index column houses datetime data, and which houses entity_data by using isinstance(obj, type). Using a for loop, we can check the identity of both index columns. We save their locations as date_level and entity_level. We save the names of these columns as date_index_name and entity_index_name.
#
# Once the type of each column has been determined, save the name of the desired index name. We can create indicator variables for each state (or time period if that is preferred) using the create_indicator_variable() method. Before , extract the list of entities names, reduce them to the unique set, transform the set into a list, and sort the list. Remove the last indicator in indicators by using .pop(). The nation missing an indicator variable will implicitly depend upon $\beta_0$. This value will serve as the anchor for the underlying regression that the indicator variables marginally shift for each state. Next, cycle through the list of indicators to create an indicator variable for each entity. We save the names of the indicator variables with the variables in *X_names* as *X_and_indicator_names* so that they can be referenced in the regression.
#
# All that is left is to run the regression. The key here is to include the names of the indicator variables in the regression but not include them in the results. To do this, we set self.X_names to include the names passed to panel_regression() and "Constant". The dataframe owned by the Regression class, data, is saved as to only include the variables indicated in *self.X_names*. Likewise, the estimated beta values for the indicator variables are removed by passing *self.X_names* to the index of self.estimates.
#
# Finally, create a script modeled from indicatorRegression.py. We’ve removed any elements relating to the indicator variables from the previous script and saved the result as panelRegression.py. You may find copying this script manually to be easier than marginally editing the previous script. Note that we have removed “North America” from X_names and that OLS() has been replaced by panel_regression().
# +
#panelRegression.py
# . . . .
X_names = ["EFW", "Log RGDP Per Capita Lag"]
y_name = ["Log RGDP Per Capita"]
#save instance of regression class
reg = Regression()
for key, data in data_dict.items():
panel_name = key + " panel"
# call panel_regression method
reg.panel_regression(reg_name = panel_name, data = data.dropna(),
y_name = y_name, X_names = X_names, entity = True, time =True )
print(key, reg.estimates, sep = "\n")
print(reg.stats_DF)
# -
# Note also the exception that is included at the beginning of the panel_regression method. If you pass entity = True and time = True, the console will return the following:
reg = Regression()
for key, data in data_dict.items():
panel_name = key + " panel"
# call panel_regression method
reg.panel_regression(reg_name = panel_name, data = data.dropna(),
y_name = y_name, X_names = X_names, entity = True, time = True)
print(key, reg.estimates, sep = "\n")
print(reg.stats_DF)
# ### Checking Explanatory Power of Panel Regression
# It is possible that accounting for fixed effects for each political entity does not actually improve explanatory power of the regression. To check for this, we can compare the results from a standard OLS regression to the results from a panel regression.
# The inclusion of entity fixed effects has improved the overall fit, though it has also diminished both the estimate of statistical significance (indicated by the p-value) and economic significance (beta estimate) of a nation’s economic freedom on its real GDP per capita.
#
# We want to be sure that using a panel regression actually improved the estimates. To do this, we run a joint f-test, as in the previous chapter.
reg = Regression()
for key, data in data_dict.items():
# call OLS and Panel for comparison
data = data_dict[key]
reg.OLS(reg_name = key, data = data.dropna(),
y_name = y_name, beta_names = X_names)
print(key, reg.estimates, sep = "\n")
print(reg.stats_DF)
panel_name = key + " panel"
reg.panel_regression(reg_name = panel_name, data = data.dropna(),
y_name = y_name, X_names = X_names, entity = True, time = True)
print(key, reg.estimates, sep = "\n")
print(reg.stats_DF)
joint_f_test = reg.joint_f_test(key, key + " panel")
print(joint_f_test)
reg.indicator_lists
| Textbook/Chapter 8 - Advanced Data Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Clasificación con SVM Lineal con el dataset Digits de scikit-learn normalizando los datos
# http://scikit-learn.org/stable/modules/generated/sklearn.datasets.load_digits.html
# ## Información del dataset
# Casi 1800 imágenes de 8x8 con los dígitos 0..9. Cada píxel se representa con un número entre 0..15, que representa un color en la escala de grises.
# En este notebook se hará el training con los datos normalizados, igual que hacen en algunos ejemplo de scikit-learn (dividen cada fila por 16 y luego le restan la media). En este notebook el training se hará directamente con los datos normalizados, sin usar ningún tipo de sampler. Para ver el mismo código pero usando RBFSampler, estará en otro notebook.
from sklearn.datasets import load_digits
#from sklearn.tree import DecisionTreeClassifier
from sklearn.svm import LinearSVC
import math
import numpy as np
digits = load_digits()
data = digits.data
target = digits.target
N = data.shape[0]
prop_train = 2 / 3
N_train = math.ceil(N * prop_train)
N_test = N - N_train
# Normaliza los datos entre 0 y 1 y luego los centra en la media
data /= 16
data -= data.mean(axis = 0)
# Los primeros 2/3 de los datos son de train, y el resto son de test
# +
data_train = data[:N_train]
data_test = data[N_train:]
target_train = target[:N_train]
target_test = target[N_train:]
# -
# ## Hacer n_runs ejecuciones y devolver la media de puntuación de todas ellas
n_runs = 50
train_scores = []
test_scores = []
for i in range(n_runs):
#clf = DecisionTreeClassifier()
clf = LinearSVC()
# Samplear los datos
indices = np.arange(len(data))
indices = np.random.choice(indices, len(indices), replace = False) # Mezclar los indices
train_indices = indices[:N_train] # Los N_train primeros son de train
test_indices = indices[N_train:] # El resto son de test
dat_train = np.take(data, train_indices, axis = 0)
targ_train = np.take(target, train_indices, axis = 0)
dat_test = np.take(data, test_indices, axis = 0)
targ_test = np.take(target, test_indices, axis = 0)
clf.fit(dat_train, targ_train)
train_score = clf.score(dat_train, targ_train)
test_score = clf.score(dat_test, targ_test)
train_scores.append(train_score)
test_scores.append(test_score)
print("Mean of test scores:",np.mean(test_scores))
print("Mean of train scores:", np.mean(train_scores))
print("Standard deviation of test scores:",np.std(test_scores))
print("Standard deviation of train scores:",np.std(train_scores))
# ### Conclusiones
# El modelo es capaz de ajustar y generalizar correctamente. No parece que haya ninguna diferencia entre normalizar los datos y no hacerlo. Los resultados son bastante parecidos.
| code/notebooks/python/SVM/SVM Lineal con Digits normalizando.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# %pylab notebook
import glob
import matplotlib.pyplot as plt
from charistools.hypsometry import Hypsometry
list = glob.glob("/Users/brodzik/projects/CHARIS/derived_hypsometries/evapotranspiration/mod16/monthly/AM_*fullbasinmasks*txt")
list
fig, ax = plt.subplots(5, 2, figsize=(14,16))
basins = ['AM', 'BR', 'GA_v01', 'IN_v01', 'SY_v01']
vmax = [0.75, 5.5, 19.0, 5.5, 0.8]
for i, basin in enumerate(basins):
list = sort(glob.glob("/Users/brodzik/projects/CHARIS/derived_hypsometries/evapotranspiration/mod16/monthly/" + basin + "_*fullbasinmasks*txt"))
print(list)
snowhyps = Hypsometry(filename=list[1])
snowhyps.data = snowhyps.data['2001-01-01':'2001-12-31']
hyps = Hypsometry(filename=list[0])
hyps.data = hyps.data['2001-01-01':'2001-12-31']
snowhyps.imshow(ax=ax[i,0], title=basin + ' (snowy) ET', cmap='Greens_r',
xlabel='Date', dateFormat='%b', ylabel='Elevation ($m$)',
vmax=vmax[i])
hyps.imshow(ax=ax[i,1], title=basin + ' ET', cmap='Greens_r',
xlabel='Date', dateFormat='%b', ylabel='Elevation ($m$)',
vmax=vmax[i])
fig.tight_layout()
#ax[0,0].plot(hyps.data['2001-01-01':'2001-01-01'])
snowhyps.imshow(ax=ax[0,0], title='AM (snowy)', cmap='Greys_r',
xlabel='Date', dateFormat='%b',
ylabel='Elevation ($m$)')
hyps.imshow(ax=ax[0,1], title='AM', cmap='Greys_r',
xlabel='Date', dateFormat='%b',
ylabel='Elevation ($m$)')
hyps.data
fig.savefig("/Users/brodzik/ipython_notebooks/charis/ET.2001.png")
fig, ax = plt.subplots(1)
list = glob.glob("/Users/brodzik/projects/CHARIS/derived_hypsometries/evapotranspiration/mod16/monthly/AM_*fullbasinmasks*txt")
# snowhyps = Hypsometry(filename=list[0])
# snowhyps.data = snowhyps.data['2001-01-01':'2001-12-31']
# hyps = Hypsometry(filename=list[1])
# hyps.data = hyps.data['2001-01-01':'2001-12-31']
# snowhyps.imshow(ax=ax[i,0], title=basin + ' (snowy) ET', cmap='Greens_r',
# xlabel='Date', dateFormat='%b', ylabel='Elevation ($m$)')
# hyps.imshow(ax=ax[i,1], title=basin + ' ET', cmap='Greens_r',
# xlabel='Date', dateFormat='%b', ylabel='Elevation ($m$)')
fig.tight_layout()
| charis/Display ET by elevation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Developing Advanced User Interfaces
# *Using Jupyter Widgets, Pandas Dataframes and Matplotlib*
# While BPTK-Py offers a number of high-level functions to quickly plot equations (such as `bptk.plot_scenarios`) or create a dashboard (e.g. `bptk.dashboard`), it is sometimes necessary to create more sophisticated plots (e.g. plots with two axes) or a more sophisticated interface dashboard for your simulation.
#
# This is actually quite easy, because BPTK-Py's high-level functions utilize some very powerful open source libraries for data management, plotting, and dashboards: Pandas, Matplotlib, and Jupyter Widgets.
#
# In order to harness the full power of these libraries, it is only necessary to understand how to make the data generated by BPTK-Py available to them. This _How To_ illustrates this using a neat little simulation of customer acquisition strategies. It is not required to understand the simulation to follow this document, but if you are interested you can read more about it on our [blog](https://www.transentis.com/an-example-to-illustrate-the-business-prototyping-methodology/).
# ## Advanced Plotting
#
# We'll start with some advanced plotting of simulation results.
# +
## Load the BPTK Package
from BPTK_Py.bptk import bptk
bptk = bptk()
# -
# BPTK-Py's workhorse for creating plots is the `bptk.plot_scenarios` function. The function generates the data you want to plot using the simulation defined by the scenario manager and the settings defined by the scenarios. The data are stored in a Pandas dataframe. For plotting results, the framework uses Matplotlib. To illustrate this, we will recreate the plot below manually:
bptk.plot_scenarios(
scenario_managers=["smCustomerAcquisition"],
scenarios=["base"],
equations=['customers'],
title="Base",
freq="M",
x_label="Time",
y_label="No. of Customers"
)
# The data generated by a scenario is can be saved into a dataframe. This can be achieved by adding the `return_df` flag to `bptk.plot_scenario`:
df=bptk.plot_scenarios(
scenario_managers=["smCustomerAcquisition"],
scenarios=["base"],
equations=['customers'],
title="Base",
freq="M",
x_label="Time",
y_label="No. of Customers",
return_df=True
)
# The dataframe is indexed by timestep and stores the equations (in SD models) or agent properties (in Agent-based models) in the columns.
df[0:10] # just show the first ten items
# Internally, the `bptk.plot_scenarios` method first runs the simulation using the settings defined in the scenario and stores the data in a dataframe. It then plots the dataframe using Pandas `df.plot`method.
#
# We can do the same:
subplot=df.plot(None,"customers")
# Because we are missing some styling information, the plot above doesn't look quite as neat as the plots created by `bptk.plot_scenarios`. The styling information is stored in BPTK_Py.config, and you can access (and modify) it there.
#
# Now let's apply the config to `df.plot`:
# +
import BPTK_Py.config as config
subplot=df.plot(kind=config.configuration["kind"],
alpha=config.configuration["alpha"], stacked=config.configuration["stacked"],
figsize=config.configuration["figsize"],
title="Base",
color=config.configuration["colors"],
lw=config.configuration["linewidth"])
# -
# Yes! We've recreated the plot from the high level `btpk.plot_scenarios` method using basic plotting functions.
#
# Now let's do something that is currently not possible using the high-level BPTK-Py methods - let's create a graph that has two y-axes.
#
# This is useful when you want to show the results of two equations at the same time, but they need to have different scales on the y-axis. For instance, in the plot below, the number of customers is much smaller than the profit made, so the customer graph looks like a straight line.
bptk.plot_scenarios(
scenario_managers=["smCustomerAcquisition"],
scenarios=["base"],
equations=['customers','profit'],
title="Base",
freq="M",
x_label="Time",
y_label="No. of Customers"
)
# As before, we collect the data in a dataframe.
df=bptk.plot_scenarios(
scenario_managers=["smCustomerAcquisition"],
scenarios=["base"],
equations=['customers','profit'],
title="Base",
freq="M",
x_label="Time",
y_label="No. of Customers",
return_df = True
)
df[0:10]
# Plotting two axes is easy in Pandas (which itself uses the Matplotlib library):
# +
ax = df.plot(None,'customers', kind=config.configuration["kind"],
alpha=config.configuration["alpha"], stacked=config.configuration["stacked"],
figsize=config.configuration["figsize"],
title="Profit vs. Customers",
color=config.configuration["colors"],
lw=config.configuration["linewidth"])
# ax is a Matplotlib Axes object
ax1 = ax.twinx()
# Matplotlib.axes.Axes.twinx creates a twin y-axis.
plot =df.plot(None,'profit',ax=ax1)
# -
# Voilà!
#
# Once you have the data from BPTK you can do anything Pandas and Matplotlib allow you to do. It might be helpful to get a deeper look at how these libraries work when creating more complex dashboards and plots.
# ## Advanced interactive user interfaces
# Now let's try something a little more challenging: Let's build a dashboard for our simulation that let's you manipulate some of the scenrio settings interactively and plots results in tabs.
# > Note: You need to have widgets enabled in Jupyter for the following to work. Please check the [BPTK-Py installation instructions](https://bptk.transentis-labs.com/en/latest/docs/usage/installation.html) or refer to the [Jupyter Widgets](https://ipywidgets.readthedocs.io/en/latest/user_install.html) documentation
# First, we need to understand how to create tabs. For this we need to import the `ipywidget` Library and we also need to access Matplotlib's `pyplot`
# %matplotlib inline
import matplotlib.pyplot as plt
from ipywidgets import interact
import ipywidgets as widgets
# Then we can create some tabs that display scenario results as follows:
# +
out1 = widgets.Output()
out2 = widgets.Output()
tab = widgets.Tab(children = [out1, out2])
tab.set_title(0, 'Customers')
tab.set_title(1, 'Profit')
display(tab)
with out1:
# turn of pyplot's interactive mode to ensure the plot is not created directly
plt.ioff()
# create the plot, but don't show it yet
bptk.plot_scenarios(
scenario_managers=["smCustomerAcquisition"],
scenarios=["hereWeGo"],
equations=['customers'],
title="Here We Go",
freq="M",
x_label="Time",
y_label="No. of Customers"
)
# show the plot
plt.show()
# turn interactive mode on again
plt.ion()
with out2:
plt.ioff()
bptk.plot_scenarios(
scenario_managers=["smCustomerAcquisition"],
scenarios=["hereWeGo"],
equations=['profit'],
title="Here We Go",
freq="M",
x_label="Time",
y_label="Euro"
)
plt.show()
plt.ion()
# -
# That was easy! The only thing you really need to understand is to turn interactive plotting in `pyplot` off before creating the tabs and then turn it on again to create the plots. If you forget to do that, the plots appear above the tabs (try it and see!).
# In the next step, we need to add some sliders to manipulate the following scenario settings:
#
# * Referrals
# * Referral Free Months
# * Referral Program Adoption %
# * Advertising Success %
# Creating a slider for the referrals is easy using the integer slider from the `ipywidgets` widget library:
widgets.IntSlider(
value=7,
min=0,
max=15,
step=1,
description='Referrals:',
disabled=False,
continuous_update=False,
orientation='horizontal',
readout=True,
readout_format='d'
)
# When manipulating a simulation model, we mostly want to start with a particular scenario and then manipulate some of the scenario settings using interactive widgets. Let's set up a new scenario for this purpose and call it `interactiveScenario`:
bptk.register_scenarios(scenario_manager="smCustomerAcquisition", scenarios=
{
"interactiveScenario": {
"constants": {
"referrals":0,
"advertisingSuccessPct":0.1,
"referralFreeMonths":3,
"referralProgamAdoptionPct":10
}
}
}
)
# We can then access the scenario using `bptk.get_scenarios`:
scenario = bptk.get_scenario("smCustomerAcquisition","interactiveScenario")
scenario.constants
bptk.plot_scenarios(scenario_managers=["smCustomerAcquisition"],
scenarios=["interactiveScenario"],
equations=['profit'],
title="Interactive Scenario",
freq="M",
x_label="Time",
y_label="Euro"
)
# The scenario constants can be accessed in the constants variable:
# Now we have all the right pieces, we can put them together using the interact function.
@interact(referrals=widgets.FloatSlider(
value=1,
min=0,
max=15,
step=1,
continuous_update=False,
description='Advertising Success Pct'
))
def dashboard(referrals):
scenario= bptk.get_scenario("smCustomerAcquisition", "interactiveScenario")
scenario.constants["referrals"]=referrals
bptk.reset_scenario_cache(scenario_manager="smCustomerAcquisition", scenario="interactiveScenario")
bptk.plot_scenarios(scenario_managers=["smCustomerAcquisition"],
scenarios=["interactiveScenario"],
equations=['profit'],
title="Interactive Scenario",
freq="M",
x_label="Time",
y_label="Euro"
)
# Now let's combine this with the tabs from above.
# +
out1 = widgets.Output()
out2 = widgets.Output()
tab = widgets.Tab(children = [out1, out2])
tab.set_title(0, 'Customers')
tab.set_title(1, 'Profit')
display(tab)
@interact(referrals=widgets.FloatSlider(
value=0,
min=0,
max=15,
step=1,
continuous_update=False,
description='Advertising Success Pct'
))
def dashboardWithTabs(referrals):
scenario= bptk.get_scenario("smCustomerAcquisition","interactiveScenario")
scenario.constants["referrals"]=referrals
bptk.reset_scenario_cache(scenario_manager="smCustomerAcquisition", scenario="interactiveScenario")
with out1:
# turn of pyplot's interactive mode to ensure the plot is not created directly
plt.ioff()
# clear the widgets output ... otherwise we will end up with a long list of plots, one for each change of settings
# create the plot, but don't show it yet
bptk.plot_scenarios(
scenario_managers=["smCustomerAcquisition"],
scenarios=["interactiveScenario"],
equations=['customers'],
title="Interactive Scenario",
freq="M",
x_label="Time",
y_label="No. of Customers"
)
# show the plot
out1.clear_output()
plt.show()
# turn interactive mode on again
plt.ion()
with out2:
plt.ioff()
out2.clear_output()
bptk.plot_scenarios(
scenario_managers=["smCustomerAcquisition"],
scenarios=["interactiveScenario"],
equations=['profit'],
title="Interactive Scenario",
freq="M",
x_label="Time",
y_label="Euro"
)
plt.show()
plt.ion()
# -
| general/how-to/how_to_developing_advanced_user_interfaces/how_to_developing_advanced_user_interfaces.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#1
lista=[]
i=0
cantidad = int(input("Ingrese tamaño de la lista: "))
while i < cantidad:
print("Ingrese nombre:")
nombre=str(input())
lista.append(nombre)
i+= 1
for nombres in lista:
print("Hola, ", nombres)
# +
#2
nmax = int(input("Ingrese número máximo de invitados: "))
i=0
invitados = []
resp1=0
while i < nmax :
print("Desea agregar un invitado? -Si es si, oprima cualquier numero, sino, oprima 1-")
resp1 = int(input())
if resp1 == 1:
print("Saliendo")
break
else:
print("Ingrese invitado: ")
agregado=str(input())
invitados.append(agregado)
i +=1
print("\nLista completada")
print(invitados)
print("\nDesea cambiar un invitado? -Si es si, oprima cualquier numero, sino, oprima 1-")
resp2= int(input())
if resp2 ==1:
print("\nDesea eliminar un invitado? -Si es si, oprima cualquier numero, sino, oprima 1-")
resp33= int(input())
if resp33 ==1:
pass
else:
print("Ingrese numero del invitado a eliminar: -Partiendo desde la posicion 0-")
resp31 = int(input())
del invitados[resp31]
print("Lista completada nuevamente")
print(invitados)
else: print("Ingrese numero del invitado a cambiar: -Partiendo desde la posicion 0- ")
resp21= int(input())
print("Ingrese nuevo nombre a ingresar: ")
nuevo=str(input())
invitados[resp21] = nuevo
print("\nLista nuevamente completada")
print(invitados)
# +
#3
tamaño = int(input("Ingrese el tamaño de la lista:"))
valor = tamaño + 1
numeros = list(range(2, valor*2, 2))
print(numeros)
numeros = list(num**3 for num in numeros)
print(numeros)
print("\nValor inicial: ", numeros[0])
print("Valores del centro: ", numeros[1:-1])
print("Valor final: ", numeros[-1])
# +
#4
tamaño = int(input("Ingrese el tamaño de la lista: "))
i=0
admin = "admin"
lista = []
while i < tamaño:
print("Agregar usuario: ")
usuario=str(input()).lower()
if usuario==admin:
print("El nombre admin no está disponible")
elif usuario in lista:
print("Este nombre ya esta registrado, ingrese otro usuario")
else:
lista.append(usuario)
i+=1
print(lista)
# +
#5
i=0
dix = {"Nombre": "", "Apellido": ""}
r1 = str(input("Ingrese Nombre: "))
dix["Nombre"] = r1
r2 = str(input("Ingrese Apellido: "))
dix["Apellido"] = r2
print(dix)
r=int(input("Desea agregar algun valor a la lista? (Ejemplo: Lugar de residencia, email, telefono)"+
"\nSi desea agregar, oprima cualquier numero, sino, escriba 1)"))
while r!=1 :
c=str(input("Ingrese el campo a agregar: "))
res=str(input("Rellene el campo agregado: "))
dix[c]=res
r=int(input("Desea agegar otro campo?(INGRESE CUALQUIER NUMERO PARA SI-OPRIMA 1 PARA SALIR):\n"))
print(dix)
# +
#6
lista = []
r=""
basta = "basta"
print("Para acabar la encuesta, escriba 'basta'")
while r!=basta:
r=str(input("Ingrese su lugar favorito: "))
lista.append(r)
print(lista[:-1])
# +
#7(1)
lista=[]
i=0
cantidad = int(input("Ingrese tamaño de la lista: "))
while i < cantidad:
print("Ingrese nombre:")
nombre=str(input())
lista.append(nombre)
i+= 1
def saludo():
for nombres in lista:
print("Hola ", nombres)
saludo()
| TareaFINAL.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Startup Lending Model (Project 2)
#
# Insert a description of the model
#
# - [**Setup**](#Setup): Runs any imports and other setup
# - [**Inputs**](#Inputs): Defines the inputs for the model
# ## Setup
#
# Setup for the later calculations are here. The necessary packages are imported.
from dataclasses import dataclass
import pandas as pd
# %matplotlib inline
# ## Inputs
#
# All of the inputs for the model are defined here. A class is constructed to manage the data, and an instance of the class containing the default inputs is created.
# +
@dataclass
class ModelInputs:
price_machine: float = 1000000
loan_life: int = 5
initial_default: float = 0.3
default_decay: float = 0.9
final_default: float = 0.4
recovery_rate: float = 0.4
interest: float = 0.2
num_iterations: int = 1000
model_data = ModelInputs()
model_data
# +
# Insert your main model inbetween the inputs and output blocks
# -
# Main answer should be a DataFrame with columns of ['Interest Rate', 'Loan Life', 'Initial Default Probability', 'IRR']
# The IRR is the expected IRR across all the default cases for the interest rate,
# loan life, and initial default probability in that row
# Be sure to save this into irr_df
irr_df =
| docsrc/source/_static/Project Materials/Project 2/Project 2 Template.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Determinant
#
# Author: [<NAME>](https://patel-zeel.github.io/), [<NAME>](https://nipunbatra.github.io/)
# In this notebook, we will look at determinants. https://youtu.be/Ip3X9LOh2dk
# +
import numpy as np
import matplotlib.pyplot as plt
from matplotlib import rc
rc('font', size=16)
rc('text', usetex=True)
# -
# ## Transformation of N-D space
# Consider a N-D vector $\mathbf{v}_1$. We can get a transformed N-D vector $\mathbf{v}_1'$ by applying an appropriate linear transformation $A$ on $\mathbf{v}_1$.
#
# Note that $A$ will be a $N \times N$ matrix here. This transformarion is given as,
#
# $$
# \mathbf{v}_1' = A\mathbf{v}_1
# $$
# ### 1D case
# If $\mathbf{v}_1$ is a scaler, we get transformed scaler $\mathbf{v}_2$ by applying transformation $A$. $A$ would also be a scaler here.
#
# If $\mathbf{v}_1=3$, $A=5$ then $\mathbf{v}_1' = 5 \cdot 3 = 15$.
#
# Ultimately, $\mathbf{v}_1'$ is $\mathbf{v}_1$ scaled by factor of $A$.
# ### 2D case
# consider two vectors $\mathbf{v}_1$ and $\mathbf{v}_2$. We will focus on the area of the parallelogram spaned by these vectors.
#
# $$
# \mathbf{v}_1 =
# \begin{bmatrix}
# 0\\
# 1
# \end{bmatrix},
# \mathbf{v}_2 =
# \begin{bmatrix}
# 1\\
# 0
# \end{bmatrix}
# $$
def get_area(v1, v2):
return np.abs(np.sum(np.cross(v1.ravel(), v2.ravel())))
def plot_area(v1, v2, suffix='', annotate=False, c='b'):
vectors = [v1, v2]
for vi, v in enumerate(vectors, 1):
plt.arrow(x=0, y=0, dx=v[0, 0], dy=v[1, 0], shape='full',
head_width=0.2, head_length=0.2, length_includes_head=True, color=c)
if annotate:
plt.text(v[0, 0]-0.5, v[1, 0]+0.1, f'v{vi}{suffix}=({v[0, 0]}, {v[1, 0]})')
## Filling the area
plt.fill([0, v1[0,0], v1[0,0]+v2[0,0], v2[0,0]],
[0, v1[1,0], v1[1,0]+v2[1,0], v2[1,0]], alpha=0.4, label=f'area = {get_area(v1, v2)}')
plt.text(0-0.5, 0+0.1, f'({0}, {0})')
plt.grid()
plt.ylim(np.min([0, v1[1,0], v2[1,0]])-1, v1[1,0]+v2[1,0]+1)
plt.xlim(np.min([0, v1[0,0], v2[0,0]])-1, v1[0,0]+v2[0,0]+1)
plt.xlabel('x');plt.ylabel('y');
if annotate:
plt.legend(bbox_to_anchor=(1,1))
# +
vec1 = np.array([2, 0]).reshape(-1,1)
vec2 = np.array([2, 1]).reshape(-1,1)
plot_area(vec1, vec2, annotate=True)
# -
# Let us apply the following linear tranformation on $\mathbf{v_1}$ and $\mathbf{v_2}$.
#
# $$
# A =
# \begin{bmatrix}
# 2 & 1\\
# 2 & 3
# \end{bmatrix}
# $$
def plot_transform(v1, v2, A, annotate=False):
plot_area(v1, v2, annotate=annotate)
xlim, ylim = plt.xlim(), plt.ylim()
v1_, v2_ = A@v1, A@v2
plot_area(v1_, v2_, suffix='\'', annotate=annotate, c='r')
plt.xlim(min(plt.xlim()[0], xlim[0]), max(plt.xlim()[1], xlim[1]))
plt.ylim(min(plt.ylim()[0], ylim[0]), max(plt.ylim()[1], ylim[1]))
plt.title(f'Area should be scaled by $|A|={np.linalg.det(A)}$')
# +
A = np.array([[2, 1], [2, 3]])
plot_transform(vec1, vec2, A, annotate=True)
# -
# Note that the area spanned by $\mathbf{v}_1'$ and $\mathbf{v}_2'$ is scaled by factor factor of $|A|$ (determinant of $A$). More concretely,
#
# $$
# \text{Area}(\mathbf{v}_1', \mathbf{v}_2') = |A| \cdot \text{Area}(\mathbf{v}_1, \mathbf{v}_2)
# $$
#
# Let us choose one more example of $A$,
#
# $$
# A =
# \begin{bmatrix}
# 2 & 4\\
# 1 & 2
# \end{bmatrix}
# $$
# +
A = np.array([[2, 4], [1, 2]])
plot_transform(vec1, vec2, A, annotate=True)
# -
# Note that because $|A|=0$, area spanned by transformed vectors is $0$ and thus the plane spanned squeezes to a line.
#
# For a 3D case, $|A|$ will denote a factor by which volume spanned is scaled. This is true in general for N-D case.
| explain-ml-book/notebooks/2021-03-15-determinant.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# So far you mastered the notation of quantum mechanics and quantum computing, understood as much physics as needed to perform various operations on quantum states, and now you are ready to build quantum algorithms. In this notebook, we look at the basics of gate-model quantum computing, which is sometimes also referred to as universal quantum computing. Most academic and commercial efforts to build a quantum computer focus on this model: Alibaba, Baidu, Google, HP, IBM Q, Intel, IonQ, Microsoft, Rigetti Computing, and Tencent all aim at this, and the list keeps expanding. It remains unclear which implementation will prove scalable: superconducting chips, photonic systems, and ion traps are the most common types, each having its own advantages and disadvantages. We abstract away, we focus on the quantum algorithms irrespective of the physical implementation.
#
# To get there, first we have to familiarize ourselves with some gates and what happens to those gates on quantum computers. The following diagram shows the software stack that bridges a problem we want to solve with the actual computational back-end [[1](#1)]:
#
# <img src="../figures/universal_quantum_workflow.png" alt="Software stack on a gate-model quantum computer" style="width: 400px;"/>
#
# First, we define the problem at a high-level and a suitable quantum algorithm is chosen. Then, we express the quantum algorithm as a quantum circuit composed of gates. This in turn has to be compiled to a specific quantum gate set available. The last step is to execute the final circuit either on a quantum processor or on a simulator.
#
# The quantum algorithms we are interested in are about machine learning. In this notebook, we look at the levels below algorithms: the definition of circuits, their compilation, and the mapping to the hardware or a simulator.
#
#
# # Defining circuits
#
# Circuits are composed of qubit registers, gates acting on them, and measurements on the registers. To store the outcome of registers, many quantum computing libraries add classical registers to the circuits. Even by this language, you can tell that this is a very low level of programming a computer. It resembles the assembly language of digital computers, in which a program consists of machine code instructions.
#
# Qubit registers are indexed from 0. We often just say qubit 0, qubit 1, and so on, to refer to the register containing a qubit. This is not to be confused with the actual state of the qubit, which can be $|0\rangle$, $|1\rangle$, or any superposition thereof. For instance, qubit 0 can be in the state $|1\rangle$.
#
# Let's take a look at the gates. In digital computing, a processor transforms bit strings to bit strings with logical gates. Any bit string can be achieved with just two gates, which makes universal computations possible with simple operations composed only of these two types of gates. It is remarkable and surprising that the same is also true for quantum computers: any unitary operation can be decomposed into elementary gates, and three types of gates are sufficient. This is remarkable since we are talking about transforming continuous-valued probability amplitudes, not just discrete elements. Yet, this result is what provides the high-level theoretical foundation for being able to build a universal quantum computer at all.
#
# Let's look at some common gates, some of which we have already seen. Naturally, all of these are unitary.
#
# | Gate |Name | Matrix |
# |------|--------------------|---------------------------------------------------------------------|
# | X | Pauli-X or NOT gate|$\begin{bmatrix}0 & 1\\ 1& 0\end{bmatrix}$|
# | Z | Pauli-Z gate |$\begin{bmatrix}1 & 0\\ 0& -1\end{bmatrix}$|
# | H | Hadamard gate |$\frac{1}{\sqrt{2}}\begin{bmatrix}1 & 1\\ 1& -1\end{bmatrix}$|
# | Rx($\theta$)| Rotation around X|$\begin{bmatrix}\cos(\theta/2) & -i \sin(\theta/2)\\ -i \sin(\theta / 2) & \cos(\theta / 2)\end{bmatrix}$|
# | Ry($\theta$)| Rotation around Y|$\begin{bmatrix}\cos(\theta/2) & -\sin(\theta/2)\\ \sin(\theta / 2) & \cos(\theta / 2)\end{bmatrix}$|
# | CNOT, CX | Controlled-NOT | $\begin{bmatrix}1 & 0 & 0 &0\\ 0 & 1 & 0 &0\\ 0 & 0 & 0 &1\\ 0 & 0 & 1 &0\end{bmatrix}$|
#
# As we have seen before, the rotations correspond to axis defined in the Bloch sphere.
#
# There should be one thing immediately apparent from the table: there are many, in fact, infinitely many single-qubit operations. The rotations, for instance, are parametrized by a continuous value. This is in stark contrast with digital circuits, where the only non-trivial single-bit gate is the NOT gate.
#
# The CNOT gate is the only two-qubit gate in this list. It has a special role: we need two-qubit interactions to create entanglement. Let's repeat the circuit for creating the $|\phi^+\rangle = \frac{1}{\sqrt{2}}(|00\rangle+|11\rangle)$. We will have two qubit registers and two classical registers for measurement output. First, let's define the circuit and plot it:
# +
from qiskit import QuantumCircuit, ClassicalRegister, QuantumRegister
from qiskit import execute
from qiskit import Aer
from qiskit.tools.visualization import circuit_drawer, plot_histogram
q = QuantumRegister(2)
c = ClassicalRegister(2)
circuit = QuantumCircuit(q, c)
circuit.h(q[0])
circuit.cx(q[0], q[1])
circuit_drawer(circuit)
# -
# Note that we can't just initialize the qubit registers in a state we fancy. All registers are initialized in $|0\rangle$ and creating a desired state is **part** of the circuit. In a sense, arbitrary state preparation is the same as universal quantum computation: the end of the calculation is a state that we desired to prepare. Some states are easier to prepare than others. The above circuit has only two gates to prepare our target state, so it is considered very easy.
#
# Let us see what happens in this circuit. The Hadamard gate prepares an equal superposition $\frac{1}{\sqrt{2}}(|0\rangle+|1\rangle)$ in qubit 0. This qubit controls an X gate on qubit 1. Since qubit 0 is in the equal superposition after the Hadamard gate, it will not apply the X gate for the first part of the superposition ($|0\rangle$) and it will apply the X gate for the second part of the superposition ($|1\rangle$). Thus we create the final state $\frac{1}{\sqrt{2}}(|00\rangle+|11\rangle)$, and we entangle the two qubit registers.
#
# A digital computer's processing unit typically has 64-bit registers and it is able to perform universal calculations on bit strings. Any complex calculation is broken down into elementary 64-bit operations, either sequentially or in parallel execution. So you may wonder what is the deal with the thousands of qubits we expect from a quantum computer. Why can't a 64-qubit quantum computer be enough?
#
# Entanglement is the easiest way to understand why we need so many qubits. Entanglement is a key resource in quantum computing and we want to make use of it. If we have 64-qubits and we want to entangle another one outside these 64 registers, we would have get rid of the qubit in one of the registers, potentially destroying a superposition and definitely destroying entanglement between that register and any other qubit on the chip. The only way to make use of superpositions and the strong correlations provided by entanglement is if the entire problem is on the quantum processing unit for the duration of the calculation.
#
# This global nature of the calculation is also the reason why there is a focus on problems that are difficult to break down into elementary calculations. The travelling salesman problem is a great example: we need to consider all cities and all distances to minimize overall travel length.
#
# To finish off the circuit, we could add a measurement to each qubit:
circuit.measure(q, c)
circuit_drawer(circuit)
# Finally, we can plot the statistics:
backend = Aer.get_backend('qasm_simulator')
job = execute(circuit, backend, shots=100)
plot_histogram(job.result().get_counts(circuit))
# As we have seen before, 01 and 10 never appear.
# # Compilation
#
# The circuit is the way to describe a quantum algorithm. It may also contain some arbitrary single or two-qubit unitary and controlled versions thereof. A quantum compiler should be able to decompose these into elementary gates.
# For instance, in Qiskit, you can get access to the general unitary using the $u3$ gate
#
# $$
# u3(\theta, \phi, \lambda) = \begin{pmatrix}
# \cos(\theta/2) & -e^{i\lambda}\sin(\theta/2) \\
# e^{i\phi}\sin(\theta/2) & e^{i\lambda+i\phi}\cos(\theta/2)
# \end{pmatrix}.
# $$
#
# The compiler decomposes it into an actual gate sequence.
# This is one task of a quantum compiler. The next one is to translate the gates given in the circuit to the gates implemented in the hardware or the simulator. In the table above, we defined many gates, but a well-chosen set of three is sufficient for universality. For engineering constraints, typically one minimal set of universal gates is implemented in the hardware. It depends on the physical architecture which three.
#
# At this point, the number of gates applied is probably already increasing: the decomposition of unitary will create many gates and the translation of gates is also likely to add more gates. An additional problem is the topology of the qubits: in some implementations not all qubit registers are connected to each other. The most popular implementation is superconducting qubits, which are manufactured on silicon chips just like any digital device you have. Since this is a quintessentially two dimensional layout, most qubits on the chip will not be connected. Here is an example topology of eight qubits on a superconducting quantum computer:
#
# <img src="../figures/eight_qubits.svg" alt="8-qubit topology" style="width: 200px;"/>
#
# If we want to perform a two-qubit operations between two qubits that are not neighbouring, we have to perform SWAP operations to switch the qubit states between registers. A SWAP consists of three CNOT gates in a sequence.
#
# The total number of gates at the end of the compilation reflects the true requirement of the hardware. *Circuit depth* is the number of time steps required to execute the circuit, assuming that gates acting on distinct qubits can operate in parallel. On current and near-term quantum computers, we want circuits to be shallow, otherwise decoherence or other forms of noise destroy our calculations.
#
# We have to emphasize that the compilation depends on the backend. On the simulator, physical constraints do not apply. If we compile the circuit above, its depth will not increase:
from qiskit.compiler import assemble
compiled_circuit = assemble(circuit, backend)
compiled_circuit.experiments[0].instructions
# # References
#
# [1] <NAME>, <NAME>, <NAME>. (2018). [Open source software in quantum computing](https://doi.org/10.1371/journal.pone.0208561). *PLOS ONE* 13(12):e0208561. <a id='1'></a>
| qiskit_version/05_Gate-Model_Quantum_Computing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Codealong 06
# +
import os
import numpy as np
import pandas as pd
import csv
import matplotlib.pyplot as plt
from sklearn import neighbors, metrics, grid_search, cross_validation
pd.set_option('display.max_rows', 10)
pd.set_option('display.notebook_repr_html', True)
pd.set_option('display.max_columns', 10)
# %matplotlib inline
plt.style.use('ggplot')
# -
# ## `Iris` dataset
df = pd.read_csv(os.path.join('iris.csv'))
df
# ## Part A - Activity: Iris Dataset Exploratory Analysis
color = df.Species.map(pd.Series({'Setosa': 'red', 'Versicolor': 'green', 'Virginica': 'blue'}))
df.plot(kind = "scatter", x = "SepalLength", y = "SepalWidth", c = color, s = 60)
df.plot(kind = "scatter", x = "PetalLength", y = "PetalWidth", c = color, s = 60)
df.plot(kind = "scatter", x = "SepalLength", y = "PetalLength", c = color, s = 60)
df.plot(kind = "scatter", x = "PetalWidth", y = "SepalWidth", c = color, s = 60)
df.plot(kind = "scatter", x = "SepalLength", y = "PetalWidth", c = color, s = 60)
df.plot(kind = "scatter", x = "SepalWidth", y = "PetalLength", c = color, s = 60)
# ## Part B - First hand-coded classifier
def my_first_classifier(row):
if
# +
y_hat = df.apply(my_first_classifier, axis = 1)
y_hat
# +
species_df = pd.DataFrame({'Observed': df.Species, 'Predicted': y_hat})
species_df
# -
# ## Part C - Classification metrics
# ### Accuracy
1. * sum(species_df.Observed == species_df.Predicted) / len(df)
# ### Misclassification rate
# +
# TODO
# -
# ## Part D - Activity: Second hand-coded classifier
def my_second_classifier(row):
if row.PetalLength < 2:
return 'Setosa'
elif row.PetalLength < 5:
return 'Versicolor'
else:
return 'Virginica'
y_hat = df.apply(my_second_classifier, axis = 1)
sum(y_hat != df.Species)
1. * sum(y_hat == df.Species) / len(df)
# ## Part E
# ### Feature matrix and label vector
X = df[ ['SepalLength', 'SepalWidth', 'PetalLength', 'PetalWidth'] ]
y = df['Species']
X
y
# ### K-Nearest Neighbors (KNN)
model = neighbors.KNeighborsClassifier(n_neighbors = 5, weights = 'uniform')
model.fit(X, y)
y_hat = model.predict(X)
# +
species_df = pd.DataFrame({'Observed': y, 'Predicted': y_hat})
species_df
# -
species_df[species_df.Predicted != species_df.Observed]
sum(species_df.Predicted != species_df.Observed)
# You can measure the precision of your prediction either manually or with `.score()`
1. * sum(species_df.Predicted == species_df.Observed) / len(df)
model.score(X, y)
df.plot(kind = "scatter", x = "PetalWidth", y = "SepalWidth", c = color, s = 60)
model_color = species_df.Predicted.map(pd.Series({'Setosa': 'red', 'Versicolor': 'green', 'Virginica': 'blue'}))
df.plot(kind = "scatter", x = "PetalWidth", y = "SepalWidth", c = model_color, s = 60)
# # weights = 'uniform'
neighbors.KNeighborsClassifier(n_neighbors = 5, weights = 'uniform').fit(X, y).score(X, y)
# # weights = 'distance'
neighbors.KNeighborsClassifier(n_neighbors = 5, weights = 'distance').fit(X, y).score(X, y)
# +
model = neighbors.KNeighborsClassifier(n_neighbors = 5, weights = 'distance')
model.fit(X, y)
y_hat = model.predict(X)
species_df = pd.DataFrame({'Observed': y, 'Predicted': y_hat})
species_df
# -
species_df[species_df.Predicted != species_df.Observed]
model_color = species_df.Predicted.map(pd.Series({'Setosa': 'red', 'Versicolor': 'green', 'Virginica': 'blue'}))
df.plot(kind = "scatter", x = "PetalWidth", y = "SepalWidth", c = model_color, s = 60)
# ## Part F - What is the best value for `k`?
# +
k = range(1, len(df))
score_df = pd.DataFrame({'k': k})
models = score_df.\
apply(lambda row: neighbors.KNeighborsClassifier(n_neighbors = row['k']).fit(X, y), axis = 1)
models
score_df['Score'] = models.map(lambda model: model.score(X, y))
# -
score_df
plt.plot(score_df.k, score_df.Score)
# ## Part G - Validation
# 60% of the dataset to train the model; the rest to test the model
train_df = df.sample(frac = .6, random_state = 0).sort()
train_df
test_df = df.drop(train_df.index)
test_df
# The error in the training set is less than the error is the test set
# +
train_X = train_df[ ['SepalLength', 'SepalWidth', 'PetalLength', 'PetalWidth'] ]
train_y = train_df['Species']
model = neighbors.KNeighborsClassifier(n_neighbors = 5, weights = 'uniform')
model.fit(train_X, train_y)
print 'train = ', model.score(train_X, train_y)
test_X = test_df[ ['SepalLength', 'SepalWidth', 'PetalLength', 'PetalWidth'] ]
test_y = test_df['Species']
print 'test = ', model.score(test_X, test_y)
# +
k = range(1, len(train_df))
score_df = pd.DataFrame({'k': k})
models = score_df.\
apply(lambda row: neighbors.KNeighborsClassifier(n_neighbors = row['k']).fit(train_X, train_y), axis = 1)
score_df['TrainScore'] = models.apply(lambda model: model.score(train_X, train_y))
score_df['TestScore'] = models.apply(lambda model: model.score(test_X, test_y))
# -
score_df
score_df.set_index('k').plot()
# ## Part H - Cross-Validation
len(train_df)
# +
k_cv = 5 # 5-fold CV
k_nn = range(1, len(train_df) * (k_cv - 1) / k_cv) # k-NN
gs = grid_search.GridSearchCV(
estimator = neighbors.KNeighborsClassifier(),
param_grid = {'n_neighbors': k_nn},
cv = cross_validation.KFold(len(train_df), n_folds = k_cv)
)
gs.fit(train_X, train_y)
score_df = pd.DataFrame({'k': [score.parameters['n_neighbors'] for score in gs.grid_scores_],
'Score': [score.mean_validation_score for score in gs.grid_scores_]})
score_df
# -
plt.plot(score_df.k, score_df.Score)
score_df[score_df.Score == score_df.Score.max()]
score_df[score_df.Score == score_df.Score.min()]
gs.score(test_X, test_y)
| CSCI 183 - Data Science/codealong-06.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# # Kinesis Data Stream
# * https://github.com/aws-samples/aws-ml-data-lake-workshop
# * https://aws.amazon.com/blogs/big-data/snakes-in-the-stream-feeding-and-eating-amazon-kinesis-streams-with-python/
#
# 
# +
import boto3
import sagemaker
import pandas as pd
sess = sagemaker.Session()
bucket = sess.default_bucket()
role = sagemaker.get_execution_role()
region = boto3.Session().region_name
sm = boto3.Session().client(service_name='sagemaker', region_name=region)
kinesis = boto3.Session().client(service_name='kinesis', region_name=region)
# -
# %store -r stream_name
# +
# TODO: Adapt to any number of shards
shard_id_1 = kinesis.list_shards(StreamName=stream_name)['Shards'][0]['ShardId']
print(shard_id_1)
shard_id_2 = kinesis.list_shards(StreamName=stream_name)['Shards'][1]['ShardId']
print(shard_id_2)
# -
# # Download Dataset
# !aws s3 cp 's3://amazon-reviews-pds/tsv/amazon_reviews_us_Digital_Software_v1_00.tsv.gz' ./data/
# +
import csv
import pandas as pd
df = pd.read_csv('./data/amazon_reviews_us_Digital_Software_v1_00.tsv.gz',
delimiter='\t',
quoting=csv.QUOTE_NONE,
compression='gzip')
df.shape
# -
df.head(5)
df_star_rating_and_review_body = df[['star_rating', 'review_body']][:100]
df_star_rating_and_review_body.shape
df_star_rating_and_review_body.head()
reviews_tsv = df_star_rating_and_review_body.to_csv(sep='\t',
header=None,
index=False)
reviews_tsv
# # Simulate Application Writing Records to the Stream
# +
data_stream_response = kinesis.describe_stream(
StreamName=stream_name
)
print(data_stream_response)
# -
partition_key = 'CAFEPERSON'
# +
data_stream = boto3.Session().client(service_name='kinesis', region_name=region)
response = data_stream.put_records(
Records=[
{
'Data': reviews_tsv.encode('utf-8'),
'PartitionKey': partition_key
},
],
StreamName=stream_name
)
# -
# # Store Variables for the Next Notebooks
# %store partition_key
# # Get Records
# +
# TODO: Adapt to any number of shards
shard_id_1 = 'shardId-000000000000'
shard_id_2 = 'shardId-000000000001'
# +
# TODO: Adapt to any number of shards
shard_iter_1 = data_stream.get_shard_iterator(StreamName=stream_name,
ShardId=shard_id_1,
ShardIteratorType='TRIM_HORIZON')['ShardIterator']
shard_iter_2 = data_stream.get_shard_iterator(StreamName=stream_name,
ShardId=shard_id_2,
ShardIteratorType='TRIM_HORIZON')['ShardIterator']
# +
records_response_1 = data_stream.get_records(
ShardIterator=shard_iter_1,
Limit=100
)
if records_response_1['Records']:
print(records_response_1['Records'][0]['Data'].decode('utf-8'))
# +
records_response_2 = data_stream.get_records(
ShardIterator=shard_iter_2,
Limit=100
)
if records_response_2['Records']:
print(records_response_2['Records'][0]['Data'].decode('utf-8'))
# +
from IPython.core.display import display, HTML
display(HTML('<b>Review <a target="blank" href="https://console.aws.amazon.com/kinesis/home?region={}#/streams/details/{}/monitoring"> Stream</a></b>'.format(region, stream_name)))
# + language="javascript"
# Jupyter.notebook.save_checkpoint();
# Jupyter.notebook.session.delete();
# -
| 11_stream/archive/99_Put_Reviews_On_Kinesis_Data_Stream_TODO.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.0.3
# language: julia
# name: julia-1.0
# ---
# # Nuclear parameters & potential
# **Improve the appearence of this and other notebooks:** In order to modify the cell-output of the notebooks and to better print the wide tables, please, create or modify the file ~/.jupyter/custom/custom.css in your home directory and add the line: div.output_area pre { font-size: 7pt;}
using JAC
using Interact
# The charge and shape of the nucleus is essential to understand the electronic level structure and which particular processes may occur in an atom or ion. Usually, these and other nuclear parameters need to be defined prior to the generation of wave functions and the computation of atomic amplitudes, properties and processes.
#
# In JAC, all information about the underlying nuclear model are kept in the data structure `Nuclear.Model` which carries the following information:
? Nuclear.Model
# For quick and simple computations, it is often sufficient to just specify the nuclear charge $Z$ and to leave all other parameters to their default values. For example, we here define such a nucleus for xenon $(Z=54)$ and will immediatly see the details of this specification:
wa = Nuclear.Model(54.)
# Of course, we can also specify all details about the nucleus by using the standard constructor:
wb = Nuclear.Model(54., "uniform", 132., 5.75, AngularJ64(5//2), 1.0, 2.0)
# ... or we make use of the **graphical (GUI)** constructor by starting from scratch or by just refining a previous nuclear model. For this, all values can first be set and then *up-dated*. This constructor returns an observable whose value is obtained by obs[]:
wx = Nuclear.Model(Gui)
wd = wx[]
wx = Nuclear.Model(Gui; model=wb)
wd = wx[]
| tutorials/03-define-nuclear-model-parameters.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#example for the Millersville group
#first we do a bunch of imports
# +
import cartopy
import matplotlib.patheffects as mpatheffects
import matplotlib.pyplot as plt
import metpy.calc as mpcalc
from metpy.plots import simple_layout, StationPlot, StationPlotLayout
from metpy.plots.wx_symbols import sky_cover, current_weather
from metpy.units import units
import pytz
from siphon.catalog import TDSCatalog
from siphon.ncss import NCSS
from datetime import datetime, timedelta
import numpy as np
from scipy.spatial import cKDTree
# %matplotlib inline
# +
def _nearestDate(dates, pivot):
return min(dates, key=lambda x: abs(x - pivot))
# +
#WMO weather codes
wx_code_map = {'': 0, 'DU':7, 'BLDU':7, 'HZ': 5, 'BR': 10, '-UP':16, 'UP': 16, 'SQ': 18, 'FC': 19,
'DRSN': 36, 'FG': 45, 'FZFG': 49, '-DZ': 51, 'DZ': 53, '+DZ': 55,
'-RA': 61, 'RA': 63, '+RA': 65, '-FZRA': 66,
'-SN': 71, 'SN': 73, '+SN': 75, '-SHRA': 80, 'SHRA': 81, '-SHSN': 85,
'-TSRA' : 95, '+TSRA':97, 'TSRA':95, 'VCTSRA': 92 }
def to_code(text):
for s in text:
if ' ' in s:
yield max(to_code(s.split()))
else:
yield wx_code_map[s]
# +
def thin_points(xy, radius, sort_key=None):
# All points masked initially
mask = np.ones(xy.shape[0], dtype=np.bool)
if sort_key is not None:
# Need in decreasing priority
sorted_indices = np.argsort(sort_key)[::-1]
else:
sorted_indices = np.arange(len(xy))
# Make our tree
tree = cKDTree(xy)
# Loop over all the potential points
for sort_ind in sorted_indices:
val = mask[sort_ind]
# Only proceed if we haven't already excluded this point
if val:
# Loop over all the neighbors within the radius
for neighbor in tree.query_ball_point(xy[sort_ind], radius):
# Mask them out, but don't mask ourselves
if neighbor != sort_ind:
mask[neighbor] = False
return mask
# +
def filter_data(data_dict, projection, radius=100000, sort_key=None, bbox=None):
'Reduce station density and remove stations outside given bounding box.'
# Pull location information from our data dictionary
lats = data_dict['latitude'][:]
lons = data_dict['longitude'][:]
# Project the lons and lats
proj_pts = projection.transform_points(cartopy.crs.PlateCarree(), lons, lats)[..., :-1]
# Only pay attention to points in the passed-in bounding box (if given)
if bbox:
min_lon, max_lon, min_lat, max_lat = bbox
bounds_x, bounds_y = projection.transform_points(cartopy.crs.PlateCarree(),
np.array([min_lon, min_lon, max_lon, max_lon]),
np.array([min_lat, max_lat, min_lat, max_lat]))[..., :-1].T
min_x = bounds_x.min()
max_x = bounds_x.max()
min_y = bounds_y.min()
max_y = bounds_y.max()
proj_x, proj_y = proj_pts.T
box_mask = (proj_x > min_x) & (proj_x < max_x) & (proj_y > min_y) & (proj_y < max_y)
else:
box_mask = np.ones(lons.shape, dtype=np.bool)
# Reduce the stations on the map--order by the specified field
keep = thin_points(proj_pts, radius=radius, sort_key=data_dict.get(sort_key))
# Combine the masks
keep = keep & box_mask
# Use the mask on every field in the dictionary--use keys so we can modify the dict
# while iterating
for key in data_dict.keys():
data_dict[key] = data_dict[key][keep]
# +
def radar_plus_obs(bb, my_datetime,
station_radius=75000., station_layout=simple_layout,
figsize=[10,8], timezone = None):
min_lon = bb['west']
min_lat = bb['south']
max_lon = bb['east']
max_lat = bb['north']
if timezone is None:
timezone = pytz.timezone('US/Central')
local_time = timezone.fromutc(my_datetime)
fancy_date_string = local_time.strftime('%A %B %d at %I:%M %p %Z')
print(fancy_date_string)
metar_cat = TDSCatalog('http://thredds.ucar.edu/thredds/catalog/nws/metar/ncdecoded/catalog.xml?'
'dataset=nws/metar/ncdecoded/Metar_Station_Data_fc.cdmr')
dataset = list(metar_cat.datasets.values())[0]
ncss = NCSS(dataset.access_urls["NetcdfSubset"])
query = ncss.query().accept('csv').time(my_datetime)
query.lonlat_box(north=max_lat, south=min_lat, east=max_lon, west=min_lon)
query.variables('air_temperature', 'dew_point_temperature', 'inches_ALTIM',
'wind_speed', 'wind_from_direction', 'cloud_area_fraction', 'weather')
data = ncss.get_data(query)
lats = data['latitude'][:]
lons = data['longitude'][:]
tair = data['air_temperature'][:]
dewp = data['dew_point_temperature'][:]
slp = (data['inches_ALTIM'][:] * units('inHg')).to('mbar')
# Convert wind to components
u, v = mpcalc.get_wind_components(data['wind_speed'] * units.knot,
data['wind_from_direction'] * units.deg)
# Need to handle missing (NaN) and convert to proper code
cloud_cover = 8 * data['cloud_area_fraction']
cloud_cover[np.isnan(cloud_cover)] = 9
cloud_cover = cloud_cover.astype(np.int)
# For some reason these come back as bytes instead of strings
stid = [s.decode() for s in data['station']]
# Convert the text weather observations to WMO codes we can map to symbols
print(data['weather'])
print(data['air_temperature'])
try:
wx_text = [s.decode('ascii') for s in data['weather']]
wx_codes = np.array(list(to_code(wx_text)))
except: #all false
wx_codes = np.array(len(data['weather'])*[0])
print(wx_codes)
sfc_data = {'latitude': lats, 'longitude': lons,
'air_temperature': tair, 'dew_point_temperature': dewp, 'eastward_wind': u,
'northward_wind': v, 'cloud_coverage': cloud_cover,
'air_pressure_at_sea_level': slp, 'present_weather': wx_codes}
fig = plt.figure(figsize=figsize)
lat_0 = (min_lat + max_lat)/2.0
lon_0 = (min_lon + max_lon)/2.0
# Set our Projection
projection = cartopy.crs.Mercator(central_longitude=lon_0,
min_latitude=min_lat, max_latitude=max_lat)
ax = fig.add_subplot(1, 1, 1, projection=projection)
# Call our function to reduce data
filter_data(sfc_data, projection, radius=station_radius, sort_key='air_temperature')
# Make the station plot
stationplot = StationPlot(ax, sfc_data['longitude'], sfc_data['latitude'],
transform=cartopy.crs.PlateCarree(),
fontsize=20)
station_layout.plot(stationplot, sfc_data)
return ax
# +
plot_kwargs = dict(path_effects=[mpatheffects.withStroke(foreground='black', linewidth=1)],
clip_on=True)
layout = StationPlotLayout()
layout.add_barb('eastward_wind', 'northward_wind', 'knots')
layout.add_value('NW', 'air_temperature', color='red', **plot_kwargs)
layout.add_value('SW', 'dew_point_temperature', color='green', **plot_kwargs)
layout.add_value('NE', 'air_pressure_at_sea_level', units='mbar', fmt=lambda v: format(10 * v, '03.0f')[-3:], clip_on=True)
layout.add_symbol('C', 'cloud_coverage', sky_cover, clip_on=True)
layout.add_symbol('W', 'present_weather', current_weather, clip_on=True)
dt = datetime(2017,5,16,1,0)
bb={'west':-90.5, 'east':-86.0,'north':44, 'south':40}
figsize=[30,20]
# +
coast = cartopy.feature.NaturalEarthFeature(category='physical', scale='10m',
facecolor='none', name='lakes')
state_borders = cartopy.feature.NaturalEarthFeature(category='cultural',
name='admin_1_states_provinces_lakes', scale='50m', facecolor='none')
ax = radar_plus_obs(bb, datetime.utcnow(), station_radius=5000., station_layout=layout,
figsize=figsize, timezone = None)
ax.add_feature(coast, facecolor='none', edgecolor='black')
ax.add_feature(cartopy.feature.BORDERS)
ax.add_feature(state_borders, linestyle="--", edgecolor='blue')
gl = ax.gridlines( draw_labels=True,
linewidth=2, color='gray', alpha=0.5, linestyle='--')
# +
#boston 42.3601° N, 71.0589° W
dt = datetime.utcnow()
bb={'west':-75.5, 'east':-70.0,'north':44, 'south':40}
figsize=[30,20]
coast = cartopy.feature.NaturalEarthFeature(category='physical', scale='10m',
facecolor='none', name='lakes')
state_borders = cartopy.feature.NaturalEarthFeature(category='cultural',
name='admin_1_states_provinces_lakes', scale='50m', facecolor='none')
ax = radar_plus_obs(bb, dt, station_radius=1000., station_layout=layout,
figsize=figsize, timezone = None)
ax.add_feature(coast, facecolor='none', edgecolor='black')
ax.add_feature(cartopy.feature.BORDERS)
ax.add_feature(state_borders, linestyle="--", edgecolor='blue')
gl = ax.gridlines( draw_labels=True,
linewidth=2, color='gray',
alpha=0.5, linestyle='--')
gl.xlabel_style = {'size': 25, 'color': 'gray'}
gl.ylabel_style = {'size': 25, 'color': 'gray'}
# -
| notebooks/Using Siphon and MetPy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python3-venv
# language: python
# name: python3-venv
# ---
# +
import networkx as nx
import pickle
import itertools
from collections import Counter
from collections import defaultdict
# +
# import list of ligands to exclude (list pre-generated based on exclusion criteria)
ligs2excl = []
with open('ligands-to-exclude.txt','r') as file:
line_list = file.readlines()
for line in line_list:
ligs2excl.append(line.split()[0])
print(len(ligs2excl))
# +
# get total number of residues
total_res_dict = pickle.load(open('total_res_dict.p','rb'))
# +
# set filters on PDBspheres data
datecut = 'current' # to include all templates currently available: 'current'
resolutioncut = 'all' # to include all resolutions: 'all'
gdccut = '60'
Nccut = '15'
N4cut = '4'
ligsizecut = '8'
clcut = '0'
# +
# create ligand binding dictionary and calculate percentage of residues that each ligand binds
ligand_dict = {}
template_dict = {}
fracres_dict = {}
ligs_leaveout = {}
all_ligs_remove = []
bind_thresh = 0.333
for lig in ligs2excl:
all_ligs_remove.append(lig)
for protnow in ['E','S','ORF3a','nsp12','nsp13','nsp14','nsp15','nsp16','nsp3','nsp5','nsp7','nsp8','nsp9',\
'nsp1','nsp2','ORF7a','nsp4','nsp10','N','ORF8']:
rlist = []
llist = []
ligand_dict[protnow] = {}
template_dict[protnow] = {}
ligs_leaveout[protnow] = []
for lig in ligs2excl:
ligs_leaveout[protnow].append(lig)
file = open('./CCC.confidence_centroid_contacts.'+Nccut+'_10_'+gdccut+'_'+N4cut+'_'+clcut+'.ligs_'+ligsizecut+'.nCoV.'+datecut+'.res'+resolutioncut,'r')
line_list = file.readlines()
for line in line_list:
# viral protein
if line.split()[0].split('.')[0].split('_')[0]=='nCoV':
protein = line.split()[0].split('.')[0].split('_')[1]
if protein=='Spike':
protein = 'S'
if protein==protnow:
# ligand
ligand = line.split()[0].split('.')[6]
# residues
binding_residues = line.split()[-1].split(',')
del binding_residues[-1]
if len(binding_residues)>0:
if ligand not in llist:
llist.append(ligand)
for residue in binding_residues:
if residue not in rlist:
rlist.append(residue)
if ligand not in ligand_dict[protnow]:
ligand_dict[protnow][ligand] = [residue]
elif ligand in ligand_dict[protnow] and residue not in ligand_dict[protnow][ligand]:
ligand_dict[protnow][ligand].append(residue)
if ligand not in ligs_leaveout[protnow]:
if residue not in template_dict[protnow]:
template_dict[protnow][residue] = 1
elif residue in template_dict[protnow]:
template_dict[protnow][residue] = template_dict[protnow][residue] + 1
file.close()
rlist.sort()
llist.sort()
fracres_dict[protnow] = {}
for lig in llist:
fracres_dict[protnow][lig] = float(len(ligand_dict[protnow][lig]))/float(total_res_dict[protnow])
if fracres_dict[protnow][lig]>bind_thresh and lig not in ligs_leaveout[protnow]:
ligs_leaveout[protnow].append(lig)
rfd_sorted = sorted(fracres_dict[protnow].items(), key=lambda x: x[1], reverse=True)
for lig in ligs_leaveout[protnow]:
if lig not in all_ligs_remove:
all_ligs_remove.append(lig)
pickle.dump(ligs_leaveout,open('ligs_leaveout.p','wb'))
# +
# Load the contact ligand residues into data structures
# Filter ligands by SMILES strings and percentage of residues they bind
# Filter PDB templates by date available, resolution, GDC value
def findOccurrences(s, ch):
return [i for i, letter in enumerate(s) if letter == ch]
with open('./CCC.confidence_centroid_contacts.'+Nccut+'_10_'+gdccut+'_'+N4cut+'_'+clcut+'.ligs_'+ligsizecut+'.nCoV.'+datecut+'.res'+resolutioncut) as M:
wer=M.readlines()
ncovdict=defaultdict(lambda: ([], [])) #first is conta, second resid
ligdict=defaultdict(set)
filedict=defaultdict(set)
all_contacts = {}
for protnow in ['E','S','ORF3a','nsp12','nsp13','nsp14','nsp15','nsp16','nsp3','nsp5','nsp7','nsp8','nsp9',\
'nsp1','nsp2','ORF7a','nsp4','nsp10','N','ORF8']:
all_contacts[protnow] = 0
for lin in wer:
if lin.split()[0].split('.')[0].split('_')[0]=='nCoV':
ligand = lin.split()[0].split('.')[6]
ncovfind=lin.find('nCoV_')
underfind=findOccurrences(lin,'_')
virprot=lin[(underfind[min(k for k,x in enumerate(underfind) if x>ncovfind)]+1):underfind[min(k for k,x in enumerate(underfind) if x>ncovfind)+1]]
if virprot=='Spike':
virprot='S'
if ligand not in ligs_leaveout[virprot]:
fins=findOccurrences(lin, '.')
spherfind=lin.find('.Sphere.')
ligid=lin[(fins[min(k for k,x in enumerate(fins) if x>spherfind)]+1):fins[1+min(k for k,x in enumerate(fins) if x>spherfind)]]
contstr=lin.strip(',\n').split()[-1]
conts=contstr.split(',')
all_contacts[virprot] = all_contacts[virprot] + len(conts)
fileSrc=lin[:lin.find(':')]
nonodes=[1 if (not cont[-2]=='_') else 0 for cont in conts]
if any(nonodes):
continue
ncovdict[virprot][1].extend(conts)
for cont in conts:
ligdict[virprot+'.'+cont].add(ligid)
filedict[virprot+'.'+cont].add(fileSrc)
for pair in itertools.combinations(conts,2):
ncovdict[virprot][0].append(pair)
print(all_contacts)
# +
# Load the data into weighted networkx graphs, one for each viral protein
from networkx.algorithms import community
import numpy as np
H_all=dict()
shared_int_dict=dict()
all_int_dict=dict()
nnodes_dict=dict()
# Loop over the viral proteins
# Create a graph for each protein
# Loop over all residues that contact ligands
# Add the residues as nodes
# Store the ligands they contact in a list (as well as files to find them)
for protnow in ['E','S','ORF3a','nsp12','nsp13','nsp14','nsp15','nsp16','nsp3','nsp5','nsp7','nsp8','nsp9']:
shared_int_dict[protnow]=dict()
all_int_dict[protnow]=dict()
H=nx.Graph()
resa=Counter(ncovdict[protnow][1])
for ress in resa.most_common():
all_int_dict[protnow][ress[0]]=ress[1]
H.add_node(ress[0],contacts=ress[1],ligands=ligdict[protnow+'.'+ress[0]],files=filedict[protnow+'.'+ress[0]])
counta=Counter(ncovdict[protnow][0])
for conn in counta.most_common():
shared_int_dict[protnow][conn[0]]=conn[1]
H.add_edge(conn[0][0], conn[0][1], weight=conn[1], invweight=1.0/conn[1]) #np.exp(-float(conn[1])))
H_all[protnow]=H
nnodes_dict[protnow]=H.number_of_nodes()
# +
# distribution of contacts per residue
from matplotlib import pyplot as plt
from scipy.stats import poisson, gamma, expon, linregress
from scipy.optimize import curve_fit
from math import exp
import pickle
cntctsperres = {}
area = {}
histcounts = {}
num_contacts = {}
for protnow in ['nsp3','nsp5','nsp12','S','ORF3a','nsp13','nsp14','nsp15','nsp16','nsp9']:
num_contacts[protnow] = {}
cntctsperres[protnow] = []
contributes=nx.get_node_attributes(H_all[protnow],'contacts')
for nd in H_all[protnow]:
cntctsperres[protnow].append(contributes[nd])
if nd not in num_contacts[protnow]:
num_contacts[protnow][nd] = contributes[nd]
plt.figure()
histout = plt.hist(cntctsperres[protnow],bins=6500,range=(1,6500))
plt.title(protnow)
plt.xlabel('Contacts per residue')
plt.ylabel('Count')
plt.show()
bincounts = histout[0]
histcounts[protnow] = bincounts
area[protnow] = sum(bincounts)
binedges = histout[1]
bincenters = 0.5 * (binedges[1:] + binedges[:-1])
print('mean = '+str(np.mean(cntctsperres[protnow])))
print('std = '+str(np.std(cntctsperres[protnow])))
print('total = '+str(sum(cntctsperres[protnow])))
print('total area = '+str(area[protnow]))
# +
# distribution of contacts per residue (continued)
plt.figure()
colors = ['r','b','m','c','g']
labels = []
i = 0
sars2_normalized_counts_dict = {}
for protnow in ['S','nsp3','nsp5','nsp9','nsp12','nsp13','nsp14','nsp16','nsp15','ORF3a']:
normalized_counts = [float(num)/float(area[protnow]) for num in histcounts[protnow]]
sars2_normalized_counts_dict[protnow] = normalized_counts
labels.append(protnow)
plt.scatter(np.log(np.arange(1,len(normalized_counts)+1,1)),np.log(normalized_counts))
plt.xlabel('Log(Contacts per residue)',fontsize=16)
plt.ylabel('Log(Normalized count)',fontsize=16)
plt.xticks(fontsize=14)
plt.yticks(fontsize=14)
plt.legend(labels,fontsize=14,loc=(1.05,0.02))
#plt.savefig('figures/contacts_per_residue_plot_SARS2proteins.png')
#pickle.dump(sars2_normalized_counts_dict,open('sars2_normalized_counts_dict.p','wb'))
# +
# find min and max number of residues in contact across all ligands
def res_contacts(prtn,filename):
min_res_contact = 100
max_res_contact = 0
file = open(filename,'r')
line_list = file.readlines()
for line in line_list:
# viral protein
if line.split()[0].split('.')[0].split('_')[0]=='nCoV':
protein = line.split()[0].split('.')[0].split('_')[1]
if protein=='Spike':
protein = 'S'
if protein==prtn:
N4 = int(line.split()[11])
if N4 < min_res_contact and N4 > 0:
min_res_contact = N4
if N4 > max_res_contact:
max_res_contact = N4
file.close()
return min_res_contact, max_res_contact
# -
from networkx.algorithms import shortest_paths
from scipy.cluster.hierarchy import linkage,dendrogram
from scipy import cluster
from matplotlib import pyplot as plt
import pickle
# +
# Find max value across series of lists
def max_nested(list_of_lists):
return max([max(x) for x in list_of_lists])
# +
# Cut tree at specific height and find relevant clusters - threshold based on number of contacts
def cut_res_clust(protnow,comout,ordered_list_of_res,cut_height,H_all,cluster_ligand_dict,min_cp_size,max_cp_size,all_contacts):
cutree = cluster.hierarchy.cut_tree(comout,height=cut_height)
clusout=[(x,cutree[k][0]) for k,x in enumerate(ordered_list_of_res)]
clustall=[]
for k in range(max([x[1] for x in clusout])+1):
clustall.append([x[0] for x in clusout if x[1]==k])
contributes=nx.get_node_attributes(H_all[protnow],'contacts')
ligributes=nx.get_node_attributes(H_all[protnow],'ligands')
n_clusters=len(clustall)
cplist=[]
cluster_ligand_dict[cut_height]={}
for m,clust in enumerate(clustall):
totalContacts=(sum([contributes[res] for res in clust]))
contactsperres = totalContacts/float(len(clust))
cout=Counter(itertools.chain.from_iterable([list(ligributes[res]) for res in clust]))
commall=cout.most_common()
commadj=list()
for ite in commall:
commadj.append((ite[0],ite[1],len(clust),float(ite[1])/len(clust)))
commadj.sort(key=lambda tup: -tup[3])
cpnow=dict()
cpnow['index']=m
cluster_ligand_list=[]
if contactsperres>90 and len(clust)>(min_cp_size-1) and len(clust)<(max_cp_size+1):
cpnow['totalContacts']=totalContacts
cpnow['residuesList']=clust
cplist.append(clust)
mols=[]
proteincount=0
proteinset=set()
for r in range(len(commadj)):
if commadj[r][3]<0.75:
break
x=commadj[r]
cluster_ligand_list.append((x[0],x[3]))
if len(x[0])==3:
continue
else:
proteinset.add(x[0])
proteincount+=1
cluster_ligand_dict[cut_height][m]=(len(clust),cluster_ligand_list)
cpnow['proteinCount']=proteincount
ligroups=[]
if proteincount:
ligroups.append({'ligs':[x for x in proteinset]})
disty=[]
if not disty:
cpnow['ligroups']=ligroups
cpnow['ligroups'].append({'ligs':[x[0] for x in mols]})
continue
comout2=linkage(disty,method='complete',optimal_ordering=True)
cutree2 = cluster.hierarchy.cut_tree(comout2,height=0.1251)
for groupy in range(max_nested(cutree2)+1):
indsnow=[k for k,val in enumerate(cutree2) if val[0]==groupy]
molgroup=[]
for ind in indsnow:
molgroup.append(mols[ind][1])
print('-----')
return cplist, cluster_ligand_dict, n_clusters
# +
# identify unique clusters - bottom to top of dendrogram (largest unique within size range)
def unique_clusters(qvec,cpdict):
cpfinal = []
for q in qvec:
if len(cpdict[q])>0:
for check_cluster in cpdict[q]:
if len(cpfinal)>0:
unique=1
cpfinal_add = []
for final_cluster in cpfinal:
if set(final_cluster).issubset(set(check_cluster))==True or set(final_cluster)==set(check_cluster):
cpfinal.remove(final_cluster)
if check_cluster not in cpfinal_add and check_cluster not in cpfinal:
cpfinal_add.append(check_cluster)
unique=0
if len(cpfinal_add)>0:
cpfinal.extend(cpfinal_add)
if unique==1 and check_cluster not in cpfinal:
cpfinal.append(check_cluster)
elif len(cpfinal)==0:
cpfinal.append(check_cluster)
return cpfinal
# +
# Save final clusters to output file
def save_final_clusters(prtn,final_clusters,directory):
try:
os.system('mkdir '+directory)
except:
pass
f = open(directory+'/clusters_'+prtn+'.txt','w')
for i in range(1,len(final_clusters)+1):
f.write('%d:\t' % i)
for item in final_clusters[i-1]:
f.write(str(item)+',')
f.write('\n')
f.close()
f = open(directory+'/clusters_'+prtn+'_formatted.txt','w')
for i in range(1,len(final_clusters)+1):
f.write('%d:\t' % i)
for item in final_clusters[i-1]:
f.write(str(item[0:-2])+', ')
f.write('\n')
f.close()
return
# +
# Renumber clusters for current date for nsp12 and Spike
def renumber_final_clusters(prtn,final_clusters_temp):
if prtn=='nsp12':
final_clusters_renum = []
final_clusters_renum.append(final_clusters_temp[3])
final_clusters_renum.extend(final_clusters_temp[0:3])
elif prtn=='S':
final_clusters_renum = []
final_clusters_renum.extend(final_clusters_temp[0:2])
final_clusters_renum.append(final_clusters_temp[3])
final_clusters_renum.append(final_clusters_temp[2])
final_clusters_renum.extend(final_clusters_temp[4:])
return final_clusters_renum
# +
# make dictionary with keys = residues, values = ligands they bind
# ligands filtered by SMILES string and percentage of residues they bind
def reslig_dict(prtn,filename):
residue_list = []
ligand_list = []
res_lig_dict = {}
file = open(filename,'r')
line_list = file.readlines()
for line in line_list:
# viral protein
if line.split()[0].split('.')[0].split('_')[0]=='nCoV':
protein = line.split()[0].split('.')[0].split('_')[1]
if protein=='Spike':
protein = 'S'
if protein==prtn:
# ligand
ligand = line.split()[0].split('.')[6]
if ligand not in ligs_leaveout[prtn]:
#if ligand not in all_ligs_remove:
if ligand not in ligand_list:
ligand_list.append(ligand)
# residues
binding_residues = line.split()[-1].split(',')
del binding_residues[-1]
for residue in binding_residues:
if residue not in residue_list:
residue_list.append(residue)
if residue in res_lig_dict:
if ligand not in res_lig_dict[residue]:
res_lig_dict[residue].append(ligand)
elif residue not in res_lig_dict:
res_lig_dict[residue] = [ligand]
file.close()
residue_list.sort()
ligand_list.sort()
return res_lig_dict
# +
# make dictionary with ligands that bind to residues in each cluster
def cluster_dict(final_clusters,reslig_dict):
cluster_dict = {}
clind = 1
for clust in final_clusters:
cluster_dict[clind] = {}
cluster_dict[clind]['residues'] = clust
cluster_ligand_list = []
for res in clust:
n_res = 0
for lig in reslig_dict[str(res)]:
n_present = 0
liginlist = 0
# calculate fraction of residues in cluster to which ligand binds
for res1 in clust:
if lig in reslig_dict[str(res1)]:
n_present = n_present + 1
lig_present_frac = float(n_present)/float(len(clust))
if ((lig,lig_present_frac)) not in cluster_ligand_list:
cluster_ligand_list.append((lig,lig_present_frac))
cluster_ligand_list_sorted = sorted(cluster_ligand_list, key=lambda x: x[1], reverse=True)
cluster_dict[clind]['ligands'] = cluster_ligand_list_sorted
clind = clind+1
return cluster_dict
# +
# Save final ligands to output file
# Filter ligands by SMILES strings and percentage of residues they bind
def save_final_ligands(prtn,cluster_dict,directory):
try:
os.system('mkdir '+directory)
except:
pass
f = open(directory+'/ligands_'+prtn+'.txt','w')
for key,value in cluster_dict.items():
f.write('%d:\t' % key)
for item in value['ligands']:
if item[0] not in ligs_leaveout[prtn]:
f.write(str(item)+',')
f.write('\n')
f.close()
return
# +
# bar plot of consensus cluster info
# Filter ligands by SMILES strings and percentage of residues they bind
def pocket_info_plot(prot,clust_dict,lig_bound_frac):
clsize = []
nligs = []
labels = []
for key,value in clust_dict.items():
clsize.append(len(clust_dict[key]['residues']))
labels.append(str(key))
lig_count = []
for lig in clust_dict[key]['ligands']:
if lig[1]>=lig_bound_frac and lig[0] not in ligs_leaveout[prot]:
lig_count.append(lig[0])
nligs.append(len(lig_count))
if len(labels)<13:
x = 1.25*np.arange(1,13+1)
diff = 13-len(labels)
for i in range(0,diff):
clsize.append(0)
nligs.append(0)
elif len(labels)==13:
x = 1.25*np.arange(1,len(labels)+1)
width = 0.4 # width of bars
fig = plt.figure()
ax = fig.add_subplot(111) # Create matplotlib axes
ax2 = ax.twinx() # create another set of axes that shares the same x-axis as ax
rects1 = ax.bar(x - width/2, clsize, width, label='Residues', color='tab:blue')
rects2 = ax2.bar(x + width/2, nligs, width, label='Ligands', color='tab:orange')
ax.set_ylabel('Number of residues',fontsize=15)
ax.set_ylim([0,45])
ax2.set_ylabel('Number of ligands',fontsize=15)
ax2.set_ylim([0,250])
ax.yaxis.label.set_color('tab:blue')
ax2.yaxis.label.set_color('tab:orange')
ax.spines['left'].set_color('tab:blue')
ax2.spines['right'].set_color('tab:orange')
ax.tick_params(axis='y', colors='tab:blue')
ax2.tick_params(axis='y', colors='tab:orange')
ax.set_xlabel('Pocket',fontsize=15)
ax.set_title(prot,fontsize=15)
ax.set_xticks(x)
ax.set_xticklabels(labels,fontsize=13)
ax.set_yticks([0,5,10,15,20,25,30,35,40,45])
ax.set_yticklabels(['0','5','10','15','20','25','30','35','40','45'],fontsize=13)
ax2.set_yticks([0,25,50,75,100,125,150,175,200,225,250])
ax2.set_yticklabels(['0','25','50','75','100','125','150','175','200','225','250'],fontsize=13)
plt.show()
#plt.savefig('figures/residue_clusters_bar_chart_'+prot+'.png')
return
# +
# Run clustering on viral proteins
maindirectory = 'cluster-output-ncov-residues-shortestpath-CCC-'+Nccut+'-10-'+gdccut+'-'+N4cut+'-'+clcut+'.ligs_'+ligsizecut
try:
os.system('mkdir '+maindirectory)
except:
pass
directory = 'cluster-output-ncov-residues-shortestpath-CCC-'+Nccut+'-10-'+gdccut+'-'+N4cut+'-'+clcut+'.ligs_'+ligsizecut+'/date_'+datecut+'_res'+resolutioncut
filename = './CCC.confidence_centroid_contacts.'+Nccut+'_10_'+gdccut+'_'+N4cut+'_'+clcut+'.ligs_'+ligsizecut+'.nCoV.'+datecut+'.res'+resolutioncut
cpall=defaultdict(list)
cldict={}
for protnow in ['nsp12','S','nsp5','nsp3','ORF3a','nsp13','nsp14','nsp15','nsp16','nsp9']:
print(protnow)
Q=dict(shortest_paths.shortest_path_length(H_all[protnow],weight='invweight'))
contall=nx.get_node_attributes(H_all[protnow],'contacts')
cont_thresh=1
ordered_list_of_res=sorted([x for x in Q.keys() if contall[x]>cont_thresh],key=lambda qk: int(qk[1:-2]))
pdistmat=[]
for k,res in enumerate(ordered_list_of_res):
pdistmat.extend([Q[res][ordered_list_of_res[x]] if ordered_list_of_res[x] in Q[res] else 1 for x in range(k+1,len(ordered_list_of_res))])
try:
comout=linkage(pdistmat,method='complete',optimal_ordering=True)
#plt.figure(figsize=(100,25))
#dendrogram(comout,labels=ordered_list_of_res,leaf_font_size=10)
#plt.title(protnow)
mrc = res_contacts(protnow,filename)
min_cp_size=10
max_cp_size=mrc[1]
print('min',mrc[0],'max',mrc[1])
cluster_ligand_dict={}
cpdict={}
n_clusters=100
q=0.001
qvec = []
while n_clusters > 1:
crc=cut_res_clust(protnow,comout,ordered_list_of_res,q,H_all,cluster_ligand_dict,min_cp_size,max_cp_size,all_contacts)
cplist=crc[0]
cpdict[q]=crc[0]
cpall[protnow].append(cplist)
cluster_ligand_dict=crc[1]
n_clusters=crc[2]
qvec.append(q)
q=q+0.001
del qvec[-1]
final_clusters_temp=unique_clusters(qvec,cpdict)
if datecut=='current' and (protnow=='nsp12' or protnow=='S'):
final_clusters = renumber_final_clusters(protnow,final_clusters_temp)
else:
final_clusters = final_clusters_temp
save_final_clusters(protnow,final_clusters,directory)
rldict = reslig_dict(protnow,filename)
cldict[protnow] = cluster_dict(final_clusters,rldict)
save_final_ligands(protnow,cldict[protnow],directory)
lig_bound_frac = 0.5
pocket_info_plot(protnow,cldict[protnow],lig_bound_frac)
pickle.dump(cldict,open(directory+'/cldict.p', 'wb'))
except ValueError:
print('Empty distance matrix')
print('---:::::---:::::::---')
| residue-clustering.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="ok9eEc1E1iRS" colab_type="text"
# Notes:
#
# make flavors and effects into standard categories...?
#
# split string into up to four different categories
#
#
# maybe
# make a function that counts the commas,(or slashes?) and split strings the string into that many columns...
# or...
# make a function that counts and then adds commas so that they all have a uniform number of commas...
#
#
# maybe append to string...
#
# count commas in string and make a new collumn...
# https://www.geeksforgeeks.org/python-pandas-split-strings-into-two-list-columns-using-str-split/
#
# + [markdown] id="R7duO07q8VU6" colab_type="text"
# manually make a set of flavour columns
# and make a boolean ?function or line that adds a true to that column if
# flavor_text contains that string
#
# https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.Series.str.contains.html
#
# df['earthy_flavor'] = df.str.contains('earthy', regex=False)
#
#
# https://stackoverflow.com/questions/43055050/python-pandas-if-column-string-contains-word-flag
#
#
# + [markdown] id="tc7Y7GjxIomd" colab_type="text" endofcell="--"
# # Steps to create flavor categories:
#
# 1. Make a 'list' of unique flavor features/categories:
# -
# -
# -
# -
# -
# -
#
# 2. make a series of flavor text with:
#
# ### flavor_series = pd.Series(flavor_data)
#
# 3. create a new column of flavor_feature with:
#
# ### df['earthy_flavor'] = flavor_series.str.contains('earthy', regex=False)
#
#
#
# --
# + id="pUoQFjQR1h3n" colab_type="code" colab={}
# Import Libraries & Packages
import pandas as pd
import numpy as np
# + id="gJ176wWjyszp" colab_type="code" colab={}
#so more head displays
#override display option
pd.set_option('display.max_rows', 500)
pd.set_option('display.max_columns', 500)
pd.set_option('display.width', 1000)
# + id="jSwz_Q-A1h6I" colab_type="code" colab={}
# csv here for download
#https://drive.google.com/open?id=15-KMmSgxISrH8WtGGZPSB-C7DANSMLny
# + id="m0OXthCnWmT-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 217} outputId="c21ffd04-d53d-4ba6-f46f-88160bed800a"
# !wget https://raw.githubusercontent.com/MedCabinet/ML_Machine_Learning_Files/master/med1.csv
# + id="WyxvIA951h7x" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="73a8ed1c-3fbd-4da5-a0c5-3bddd65065a1"
# !ls
# + id="2jAu9xCe1h-I" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="d8243d7f-5aec-4992-d80b-3f62a47f524d"
# for colab
# upload file from local drives
import os
from google.colab import files
#uploaded = files.upload()
# !ls #check file is there
# + id="CIgTJzih1r6l" colab_type="code" colab={}
df = pd.read_csv('med1.csv')
# + id="8wsWPnG2TuYx" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="46b4cc6c-b623-484d-9092-d67002868c4d"
df.shape
# + id="JAr6jz3C1tGI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 168} outputId="d63020ca-8214-403c-e977-8cf737a90d57"
df.head(4)
# + id="nyfYRrnZ1tI8" colab_type="code" colab={}
# Making Flavour Features
# + id="Kyq6--TAICrt" colab_type="code" colab={}
#http://www.datasciencemadesimple.com/create-series-in-python-pandas/
flavor_data = df['Flavor']
# + id="i0QZ91DZIN1b" colab_type="code" colab={}
flavor_series = pd.Series(flavor_data)
# + id="9etSJP8827RU" colab_type="code" colab={}
# model: tested, works
# df['Earthy_flavor'] = flavor_series.str.contains('Earthy', regex=False)
# + id="3hzoCmJ2M-bJ" colab_type="code" colab={}
df['Ammonia_flavor'] = flavor_series.str.contains('Ammonia', regex=False)
df['Apple_flavor'] = flavor_series.str.contains('Apple', regex=False)
df['Apricot_flavor'] = flavor_series.str.contains('Apricot', regex=False)
df['Berry_flavor'] = flavor_series.str.contains('Berry', regex=False)
df['Blue_flavor'] = flavor_series.str.contains('Blue', regex=False)
df['Blueberry_flavor'] = flavor_series.str.contains('Blueberry', regex=False)
df['Citrus_flavor'] = flavor_series.str.contains('Citrus', regex=False)
df['Cheese_flavor'] = flavor_series.str.contains('Cheese', regex=False)
df['Chemical_flavor'] = flavor_series.str.contains('Chemical', regex=False)
df['Chestnut_flavor'] = flavor_series.str.contains('Chestnut', regex=False)
df['Diesel_flavor'] = flavor_series.str.contains('Diesel', regex=False)
df['Earthy_flavor'] = flavor_series.str.contains('Earthy', regex=False)
df['Flowery_flavor'] = flavor_series.str.contains('Flowery', regex=False)
df['Fruit_flavor'] = flavor_series.str.contains('Fruit', regex=False)
df['Grape_flavor'] = flavor_series.str.contains('Grape', regex=False)
df['Grapefruit_flavor'] = flavor_series.str.contains('Grapefruit', regex=False)
df['Honey_flavor'] = flavor_series.str.contains('Honey', regex=False)
df['Lavender_flavor'] = flavor_series.str.contains('Lavender', regex=False)
df['Lemon_flavor'] = flavor_series.str.contains('Lemon', regex=False)
df['Mango_flavor'] = flavor_series.str.contains('Mango', regex=False)
df['Menthol_flavor'] = flavor_series.str.contains('Menthol', regex=False)
df['Mint_flavor'] = flavor_series.str.contains('Mint', regex=False)
df['Minty_flavor'] = flavor_series.str.contains('Minty', regex=False)
df['Nutty_flavor'] = flavor_series.str.contains('Nutty', regex=False)
df['Orange_flavor'] = flavor_series.str.contains('Orange', regex=False)
df['Peach_flavor'] = flavor_series.str.contains('Peach', regex=False)
df['Pepper_flavor'] = flavor_series.str.contains('Pepper', regex=False)
df['Pine_flavor'] = flavor_series.str.contains('Pine', regex=False)
df['Pineapple_flavor'] = flavor_series.str.contains('Pineapple', regex=False)
df['Pungent_flavor'] = flavor_series.str.contains('Pungent', regex=False)
df['Sage_flavor'] = flavor_series.str.contains('Sage', regex=False)
df['Skunk_flavor'] = flavor_series.str.contains('Skunk', regex=False)
df['SpicyHerbal_flavor'] = flavor_series.str.contains('Spicy/Herbal', regex=False)
df['Strawberry'] = flavor_series.str.contains('Strawberry', regex=False)
df['Sweet_flavor'] = flavor_series.str.contains('Sweet', regex=False)
df['Tea_flavor'] = flavor_series.str.contains('Tea', regex=False)
df['Tobacco_flavor'] = flavor_series.str.contains('Tobacco', regex=False)
df['Tree_flavor'] = flavor_series.str.contains('Tree', regex=False)
df['Tropical_flavor'] = flavor_series.str.contains('Tropical', regex=False)
df['Vanilla_flavor'] = flavor_series.str.contains('Vanilla', regex=False)
df['Violet_flavor'] = flavor_series.str.contains('Violet', regex=False)
df['Woody_flavor'] = flavor_series.str.contains('Woody', regex=False)
# + id="oXoJXbIKM-lV" colab_type="code" colab={}
# + [markdown] id="zBcJE0nNVxD9" colab_type="text"
# ## List of Flavours
#
# Ammonia
# Apple
# Apricot
#
# Berry
# Blue
# Blueberry
#
# Citrus
# Cheese
# Chemical
# Chestnut
#
# Diesel
#
# Earthy
#
# Flowery
# Fruit
#
# Grape
# Grapefruit
#
# Honey
#
# Lavender
# Lemon
#
# Mango
# Menthol
# Mint
# Minty
#
# Nutty
#
# Orange
#
# Peach
# Pepper
# Pine
# Pineapple
# Pungent
#
# Sage
# Skunk
# Spicy/Herbal
# Strawberry
# Sweet
#
# Tea
# Tobacco
# Tree
# Tropical
#
# Vanilla
# Violet
#
# Woody
#
# + id="PQfaZ588TJ68" colab_type="code" colab={}
# + colab_type="code" id="KT_bqA6KT2jr" colab={}
# Making Effects Features
# + colab_type="code" id="qqfZdaKmT2jy" colab={}
#http://www.datasciencemadesimple.com/create-series-in-python-pandas/
effect_data = df['Effects']
# + colab_type="code" id="zV5xgAJ2T2j0" colab={}
effect_series = pd.Series(effect_data)
# + colab_type="code" id="OKa8ygx3T2j3" colab={}
# model, tested, works
# df['Aroused_effect'] = effect_series.str.contains('Aroused', regex=False)
# + id="BrR0u_IoTKPy" colab_type="code" colab={}
df['Aroused_effect'] = effect_series.str.contains('Aroused', regex=False)
df['Creative_effect'] = effect_series.str.contains('Creative', regex=False)
df['Energetic_effect'] = effect_series.str.contains('Energetic', regex=False)
df['Euphoric_effect'] = effect_series.str.contains('Euphoric', regex=False)
df['Focused_effect'] = effect_series.str.contains('Focused', regex=False)
df['Giggly_effect'] = effect_series.str.contains('Giggly', regex=False)
df['Happy_effect'] = effect_series.str.contains('Happy', regex=False)
df['Hungry_effect'] = effect_series.str.contains('Hungry', regex=False)
df['Relaxed_effect'] = effect_series.str.contains('Relaxed', regex=False)
df['Sleepy_effect'] = effect_series.str.contains('Sleepy', regex=False)
df['Talkative_effect'] = effect_series.str.contains('Talkative', regex=False)
df['Tingly_effect'] = effect_series.str.contains('Tingly', regex=False)
df['Uplifted_effect'] = effect_series.str.contains('Uplifted', regex=False)
df['None_effect'] = effect_series.str.contains('None', regex=False)
# + [markdown] id="oEcVwKjzTKlq" colab_type="text"
# ## List of Effects
#
# Aroused
#
# Creative
#
# Energetic
# Euphoric
#
# Focused
#
# Giggly
#
# Happy
# Hungry
#
# Relaxed
#
# Sleepy
#
# Talkative
# Tingly
#
# Uplifted
#
# None
# + id="eS_lybTmWuek" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 348} outputId="1e2a61d4-2cf3-4fc3-feec-61ed7e432cd7"
df.head(3)
# + id="1xqxsra8aIO0" colab_type="code" colab={}
# https://stackoverflow.com/questions/17383094/how-can-i-map-true-false-to-1-0-in-a-pandas-dataframe
df_int = df.replace([True, False],
[1.0, 0.0])
# + id="Wcuq0guMaItG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 348} outputId="f4c2394f-b30f-414a-a0bf-2727fc6ffce7"
df_int.head(3)
# + id="afRtnNbVboM0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="ea528282-7c65-45ed-d976-24895da53343"
df_int.dtypes
# + id="oJNg0Niodvpj" colab_type="code" colab={}
| Build_Data_inspection_Colab_v2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Métodos Úteis para Selenium
#
# 1. Apesar de não ser do selenium, a biblioteca time vai ajudar muito a gente, para esperar a página carregar ou esperar algo acontecer.
# - para descobrirmos o id, classe ou css selector, clicamos com o botão direito onde queremos achar, depois clicamos na setinha e clicamos no mesmo local
# - https://selenium-python.readthedocs.io/locating-elements.html
# + active=""
# import time
#
# time.sleep(2) -> espera 2 segundos
# -
# 2. Para selecionar elementos teremos os find_element e find_elements -> a biblioteca tem vários exemplos disso aqui: https://selenium-python.readthedocs.io/locating-elements.html
# + active=""
# elemento = driver.find_element_by_id('id_do_objeto') -> 1 único elemento com aquele id
#
# elementos = driver.find_elements_by_id('id_do_objeto') -> lista com vários elementos com aquele id
#
# Normalmente id é único, mas isso não é 100% garantido.
# -
# 3. Clicar em um elemento
# + active=""
# elemento.click()
# -
# 4. Para preencher um formulário, usaremos o send_keys. A biblioteca do selenium tem o KEYS que vamos importar porque isso pode ajudar
# + active=""
# from selenium.webdriver.common.keys import Keys
#
# elemento_input.send_keys('texto') -> preenche o campo com texto
# elemento_input.send_keys(Keys.RETURN) -> dá enter no campo
# -
# 5. Para selecionar um objeto pelo css dele, podemos usar o css_selector
# + active=""
# link_classe = driver.find_elements_by_css_selector('tag.classe')
# +
from selenium import webdriver
import time
driver = webdriver.Chrome()
driver.get('https:hashtagtreinamentos.com') # vai abrir o site
time.sleep(3) # vai demorar 3 segundos, para o site carregar por completo
# driver.find_element_by_class_name('btn-laranja').click() # primeiro tente pegar pelo id, class, css selector
# como tinha o mesmo id e classe que outros elementos, pegamos pelo 'name'
elementos = driver.find_elements_by_name('fullname')
# print(len(elementos)) - aqui viu que tem apenas 2 elementos com esse nome, então é de boa
# no name 0 vai escrever 'Fernanda'
elementos[0].send_keys('Fernanda')
# no email 0 vai colocar o e-mail
elementos = driver.find_elements_by_name('email')
elementos[0].send_keys('<EMAIL>')
# achou a classe do formulário inteiro, nunca pegue pelo id do formulário inteiro
driver.find_element_by_class_name('_form ').submit()
# ou faça desse jeito:
# driver.find_element_by_id('_form_173_submit').click()
| primeiros-passos.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import matplotlib.pyplot as plt
# Loads the aligned trajectory
import MDAnalysis as mda
ref = mda.Universe('protein.pdb')
sim = mda.Universe('protein.pdb', '0/1ubq_aligned_protein.dcd')
# # Principal components analysis
import MDAnalysis.analysis.pca as pca
sim_pca = pca.PCA(sim, select='protein and name CA')
sim_pca.run(start=40)
# ## The principal components themselves
plt.plot(sim_pca.p_components[:,0])
plt.plot(sim_pca.p_components[:,1])
# The first two principal components are focused on the floppy tail of the protein.
import numpy as np
print('The dot product of the first PC with itself is', np.dot(sim_pca.p_components[:,0],sim_pca.p_components[:,0]))
print('The dot product of the first and second PCs is', np.dot(sim_pca.p_components[:,0],sim_pca.p_components[:,1]))
# ## The variance and cumulated variance
plt.plot(sim_pca.variance,'.-')
plt.subplot(2,1,1)
plt.plot(sim_pca.cumulated_variance[:10],'.-')
plt.subplot(2,1,2)
plt.plot(sim_pca.cumulated_variance,'.-')
# The majority of the variance can be explained with two dimensions
# ## Projections onto the most important principal components
atomgroup = sim.select_atoms('protein and name CA')
pca_space = sim_pca.transform(atomgroup, n_components=10)
plt.plot(pca_space[:,0],'.-')
plt.plot(pca_space[:,1],'.-')
plt.plot(pca_space[:,2],'.-')
plt.scatter(pca_space[:,0], pca_space[:,1], c=range(pca_space.shape[0]), cmap="RdYlGn")
plt.colorbar()
| static_files/tutorials/ubq_wat-md/3-analysis/.ipynb_checkpoints/PCA-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from qiskit import(
QuantumCircuit,
QuantumRegister,
execute,
Aer)
from qiskit.visualization import plot_histogram
# # 1stepLBNL
from onestepSim_LBNL import runQuantum
circuit_LBNL1 = runQuantum(gLR=1,dophisplit=1)
# +
circuit=circuit_LBNL1
simulator = Aer.get_backend('qasm_simulator')
job = execute(circuit, simulator, shots=2000)
result = job.result()
counts = result.get_counts(circuit)
plot_histogram(counts,number_to_keep=None)
# -
circuit_LBNL1.draw(output='mpl', fold =100, filename='1step.pdf')
# # 2stepLBNL
from twostepSim_LBNL import runQuantum
circuit_LBNL2 = runQuantum(gLR=1,dophisplit=1)
# +
circuit=circuit_LBNL2
simulator = Aer.get_backend('qasm_simulator')
job = execute(circuit, simulator, shots=2000)
result = job.result()
counts = result.get_counts(circuit)
plot_histogram(counts,number_to_keep=None)
# -
circuit_LBNL2.draw(output='mpl', fold=100)
| sample/input_circuit.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Bernstein-Vazirani algorithm(Overview)
# We explain Bernstein-Vazirani algorithm.
#
# In this algorithm, we consider the following function that takes $x$ as input and computes the output using the internal bit sequence $a$.
#
# $$
# f_a(x) = (a\cdot x) \bmod2 = (\sum_i a_i x_i) \bmod2
# $$
#
# The goal is to find the bit sequence $a$ from the output of $f_a(x)$.
#
# $a, x$ are $n$-bit strings $a = a_0 a_1... a_{n-1}$, $x = x_0 x_1.... x_{n-1}$, respectively.
# Consider the case where $a$ is a 4-bit sequence $1001$.
# A foolproof way to find $x$ in classical calculations is to input $x$ in order, with only one bit set to 1 and the others to 0, as in $x=1000, 0100, 0010, 0001$.
# From $0 \cdot 0 = 0 \cdot 1 = 0$, the bit with $x_i=0$ do not affect the result, so we can determine the value of $a_i$ one bit at a time.
#
# Calculate $f_a(x)$ where $a=1001$ for four different $x$ as follows.
#
# $(1001 \cdot 1000) \bmod2 = 1$
# $(1001 \cdot 0100) \bmod2 = 0$
# $(1001 \cdot 0010) \bmod2 = 0$
# $(1001 \cdot 0001) \bmod2 = 1$
#
# From the above output, we can get $a=1001$.
# The Bernstein-Vazirani algorithm uses quantum superposition to find $a$ in a single calculation.
#
# The specific circuit for the case $a=1001$ is as follows.
#
# <img src="../tutorial-ja/img/102_img.png" width="30%">
#
# Let the $CX$ gate act on the qubits $i$, where $a_i=1$ are the control qubits, respectively.
# All target qubits are the fourth auxiliary qubit.
# Let's check the states.
#
# $$
# \begin{align}
# \lvert \psi_1\rangle &= \biggl(\otimes^4 H\lvert 0\rangle \biggr)\otimes H \lvert 1\rangle \\
# &= \frac{1}{\sqrt{2}}(\lvert 0\rangle + \lvert 1\rangle) \otimes \frac{1}{\sqrt{2}}(\lvert 0\rangle + \lvert 1\rangle) \otimes \frac{1}{\sqrt{2}}(\lvert 0\rangle + \lvert 1\rangle) \otimes \frac{1}{\sqrt{2}}(\lvert 0\rangle + \lvert 1\rangle) \otimes \frac{1}{\sqrt{2}}(\lvert 0\rangle - \lvert 1\rangle)
# \end{align}
# $$
# Next, check the state of $\lvert \psi_2\rangle$
# Let's focus only on the 0th and 4th qubits that the $CX$ gates act on.
#
# $$
# \begin{align}
# \lvert \psi_1\rangle_{04} &= \frac{1}{\sqrt{2}}(\lvert 0\rangle_0 + \lvert 1\rangle_0) \otimes \frac{1}{\sqrt{2}}(\lvert 0\rangle_4 - \lvert 1\rangle_4) \\
# &= \lvert +\rangle_0 \otimes \lvert -\rangle_4
# \end{align}
# $$
#
# $$
# \begin{align}
# \lvert \psi_2\rangle_{04} &= \frac{1}{\sqrt{2}}\lvert 0\rangle_0 \otimes \frac{1}{\sqrt{2}}(\lvert 0\rangle_4 - \lvert 1\rangle_4) + \frac{1}{\sqrt{2}}\lvert 1\rangle_0 \otimes \frac{1}{\sqrt{2}}(\lvert 1\rangle_4 - \lvert 0\rangle_4) \\
# &= \frac{1}{\sqrt{2}}\lvert 0\rangle_0 \otimes \frac{1}{\sqrt{2}}(\lvert 0\rangle_4 - \lvert 1\rangle_4) - \frac{1}{\sqrt{2}}\lvert 1\rangle_0 \otimes \frac{1}{\sqrt{2}}(\lvert 0\rangle_4 - \lvert 1\rangle_4) \\
# &= \frac{1}{\sqrt{2}}(\lvert 0\rangle_0 - \lvert 1\rangle_0 ) \otimes \frac{1}{\sqrt{2}}(\lvert 0\rangle_4 - \lvert 1\rangle_4) \\
# &= \lvert -\rangle_0 \otimes \lvert -\rangle_4
# \end{align}
# $$
#
#
# By acting on the 0th and 4th qubits with a $CX$ gate, we inverted the phase of the 0th qubit, changing it to $\lvert + \rangle \to \lvert - \rangle$.
# Similarly, with $CX$ gates acting on the third and fourth qubits, the state $\lvert\psi_3\rangle$ is as follows
#
# $$
# \lvert \psi_3\rangle = \frac{1}{\sqrt{2}}(\lvert 0\rangle - \lvert 1\rangle) \otimes \frac{1}{\sqrt{2}}(\lvert 0\rangle + \lvert 1\rangle) \otimes \frac{1}{\sqrt{2}}(\lvert 0\rangle + \lvert 1\rangle) \otimes \frac{1}{\sqrt{2}}(\lvert 0\rangle - \lvert 1\rangle) \otimes \frac{1}{\sqrt{2}}(\lvert 0\rangle - \lvert 1\rangle)
# $$
# Finally, let the H-gate act.
#
# $$
# \lvert \psi_4\rangle = \lvert 1\rangle \otimes \lvert 0\rangle \otimes \lvert 0\rangle \otimes \lvert 1\rangle \otimes \frac{1}{\sqrt{2}}(\lvert 0\rangle - \lvert 1\rangle)
# $$
#
# Measure the 0th to third qubits and we get $1001$ as the measurement result.
# Now we can find $a$.
# Let's implement this with blueqat.
from blueqat import Circuit
import numpy as np
# First, prepare a function to act oracle $U_f$.
def Uf(c, a):
N = len(a)
for i, val in enumerate(list(a)):
if val == '1':
c.cx[i, len(a)]
# The following is the main body of the algorithm.
# First, determine the $a$ you want to find by a random number.
#
# Find $a$ from the output result of the quantum circuit using the oracle, and check if the answer is correct.
# +
n = 4
a = ''
for i in range(n):
a += str(np.random.randint(2))
c = Circuit(n + 1)
c.x[n].h[:]
Uf(c, a)
c.h[:].m[:]
res = c.run(shots = 1000)
print(res)
if [arr[:n] for arr in res.keys()] == [a]:
print("OK")
else:
print("incorrect")
# -
# From the above, we were able to find the $a$ that the oracle has internally by using the Bernstein-Vazirani algorithm.
| tutorial/102_bernstein-vazirani.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import tarfile
from six.moves import urllib
DOWNLOAD_ROOT = "https://raw.githubusercontent.com/ageron/handson-ml/master/"
HOUSING_PATH = os.path.join("datasets", "housing")
HOUSING_URL = DOWNLOAD_ROOT + "datasets/housing/housing.tgz"
def fetch_housing_data(housing_url=HOUSING_URL, housing_path=HOUSING_PATH):
if not os.path.isdir(housing_path):
os.makedirs(housing_path)
tgz_path = os.path.join(housing_path, "housing.tgz")
urllib.request.urlretrieve(housing_url, tgz_path)
housing_tgz = tarfile.open(tgz_path)
housing_tgz.extractall(path=housing_path)
housing_tgz.close()
import pandas as pd
def load_housing_data(housing_path=HOUSING_PATH):
csv_path = os.path.join(housing_path, "housing.csv")
return pd.read_csv(csv_path)
# +
housing=load_housing_data()
housing.info()
# -
housing["ocean_proximity"].value_counts()
housing.describe()
# %matplotlib inline
# only in a Jupyter notebook
import matplotlib.pyplot as plt
housing.hist(bins=50, figsize=(20,15))
plt.show()
import numpy as np
def split_train_test(data, test_ratio):
shuffled_indices = np.random.permutation(len(data))
test_set_size = int(len(data) * test_ratio)
test_indices = shuffled_indices[:test_set_size]
train_indices = shuffled_indices[test_set_size:]
return data.iloc[train_indices], data.iloc[test_indices]b
train_set, test_set = split_train_test(housing, 0.2)
print(len(train_set), "train +", len(test_set), "test")
import hashlib
def test_set_check(identifier, test_ratio, hash):
return hash(np.int64(identifier)).digest()[-1] < 256 * test_ratio
def split_train_test_by_id(data, test_ratio, id_column, hash=hashlib.md5):
ids = data[id_column]
in_test_set = ids.apply(lambda id_: test_set_check(id_, test_ratio, hash))
return data.loc[~in_test_set], data.loc[in_test_set]
housing_with_id = housing.reset_index()
# adds an `index` column
train_set, test_set = split_train_test_by_id(housing_with_id, 0.2, "index")
housing["income_cat"] = np.ceil(housing["median_income"] / 1.5)
housing["income_cat"].where(housing["income_cat"] < 5, 5.0, inplace=True)
housing.hist(bins=50, figsize=(20,15))
plt.show()
# +
from sklearn.model_selection import StratifiedShuffleSplit
split = StratifiedShuffleSplit(n_splits=1, test_size=0.2, random_state=42)
for train_index, test_index in split.split(housing, housing["income_cat"]):
strat_train_set = housing.loc[train_index]
strat_test_set = housing.loc[test_index]
housing["income_cat"].value_counts() / len(housing)
# -
for set_ in (strat_train_set, strat_test_set):
set_.drop("income_cat", axis=1, inplace=True)
housing = strat_train_set.copy()
housing.plot(kind="scatter", x="longitude", y="latitude")
housing.plot(kind="scatter", x="longitude", y="latitude", alpha=0.1)
housing.plot(kind="scatter", x="longitude", y="latitude", alpha=0.4,
s=housing["population"]/100, label="population", figsize=(10,7),
c="median_house_value", cmap=plt.get_cmap("jet"), colorbar=True,
)
plt.legend()
corr_matrix = housing.corr()
corr_matrix["median_house_value"].sort_values(ascending=False)
# +
from pandas.plotting import scatter_matrix
attributes = ["median_house_value", "median_income", "total_rooms",
"housing_median_age"]
scatter_matrix(housing[attributes], figsize=(12, 8))
# -
housing.plot(kind="scatter", x="median_income", y="median_house_value",alpha=0.1)
# +
housing = strat_train_set.drop("median_house_value", axis=1)
housing_labels = strat_train_set["median_house_value"].copy()
housing
# -
housing.dropna(subset=["total_bedrooms"])
# option 1
housing.drop("total_bedrooms", axis=1)
# option 2
median = housing["total_bedrooms"].median() # option 3
median
housing["total_bedrooms"].fillna(median, inplace=True)
housing
from sklearn.preprocessing import Imputer
imputer = Imputer(strategy="median")
housing_num = housing.drop("ocean_proximity", axis=1)
imputer.fit(housing_num)
imputer.statistics_
housing_num.median().values
X = imputer.transform(housing_num)
X
housing_tr = pd.DataFrame(X, columns=housing_num.columns)
housing_tr
from sklearn.preprocessing import LabelEncoder
encoder = LabelEncoder()
housing_cat = housing["ocean_proximity"]
housing_cat_encoded = encoder.fit_transform(housing_cat)b
print(encoder.classes_)
from sklearn.preprocessing import OneHotEncoder
encoder = OneHotEncoder()
housing_cat_1hot = encoder.fit_transform(housing_cat_encoded.reshape(-1,1))
housing_cat_1hot
housing_cat_1hot.toarray()
from sklearn.preprocessing import LabelBinarizer
encoder = LabelBinarizer()
housing_cat_1hot = encoder.fit_transform(housing_cat)
housing_cat_1hot
# +
from sklearn.base import BaseEstimator, TransformerMixin
rooms_ix, bedrooms_ix, population_ix, household_ix = 3, 4, 5, 6
class CombinedAttributesAdder(BaseEstimator, TransformerMixin):
def __init__(self, add_bedrooms_per_room = True): # no *args or **kargs
self.add_bedrooms_per_room = add_bedrooms_per_room
def fit(self, X, y=None):
return self # nothing else to do
def transform(self, X, y=None):
rooms_per_household = X[:, rooms_ix] / X[:, household_ix]
population_per_household = X[:, population_ix] / X[:, household_ix]
if self.add_bedrooms_per_room:
bedrooms_per_room = X[:, bedrooms_ix] / X[:, rooms_ix]
return np.c_[X, rooms_per_household, population_per_household,
bedrooms_per_room]
else:
return np.c_[X, rooms_per_household, population_per_household]
attr_adder = CombinedAttributesAdder(add_bedrooms_per_room=False)
housing_extra_attribs = attr_adder.transform(housing.values)
# +
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler
num_pipeline = Pipeline([
('imputer', Imputer(strategy="median")),
('attribs_adder', CombinedAttributesAdder()),
('std_scaler', StandardScaler()),
])
housing_num_tr = num_pipeline.fit_transform(housing_num)
# -
from sklearn.base import BaseEstimator, TransformerMixin
class DataFrameSelector(BaseEstimator, TransformerMixin):
def __init__(self, attribute_names):
self.attribute_names = attribute_names
def fit(self, X, y=None):
return self
def transform(self, X):
return X[self.attribute_names].values
from sklearn.base import TransformerMixin #gives fit_transform method for free
class MyLabelBinarizer(TransformerMixin):
def __init__(self, *args, **kwargs):
self.encoder = LabelBinarizer(*args, **kwargs)
def fit(self, x, y=0):
self.encoder.fit(x)
return self
def transform(self, x, y=0):
return self.encoder.transform(x)
num_attribs = list(housing_num)
cat_attribs = ["ocean_proximity"]
num_pipeline = Pipeline([
('selector', DataFrameSelector(num_attribs)),
('imputer', Imputer(strategy="median")),
('attribs_adder', CombinedAttributesAdder()),
('std_scaler', StandardScaler()),
])
cat_pipeline = Pipeline([
('selector', DataFrameSelector(cat_attribs)),
('label_binarizer', MyLabelBinarizer()),
])
from sklearn.pipeline import FeatureUnion
full_pipeline = FeatureUnion(transformer_list=[
("num_pipeline", num_pipeline),
("cat_pipeline", cat_pipeline),
])
housing_prepared = full_pipeline.fit_transform(housing)
housing_prepared
housing_prepared.shape
# +
from sklearn.linear_model import LinearRegression
lin_reg = LinearRegression()
lin_reg.fit(housing_prepared, housing_labels)
# -
some_data = housing.iloc[:5]
some_data
some_labels = housing_labels.iloc[:5]
some_data_prepared = full_pipeline.transform(some_data)
some_data_prepared
print("Predictions:", lin_reg.predict(some_data_prepared))
print("Labels:", list(some_labels))
from sklearn.metrics import mean_squared_error
housing_predictions = lin_reg.predict(housing_prepared)
lin_mse = mean_squared_error(housing_labels, housing_predictions)
lin_rmse = np.sqrt(lin_mse)
lin_rmse
from sklearn.tree import DecisionTreeRegressor
tree_reg = DecisionTreeRegressor()
tree_reg.fit(housing_prepared, housing_labels)
housing_predictions = tree_reg.predict(housing_prepared)
tree_mse = mean_squared_error(housing_labels, housing_predictions)
tree_rmse = np.sqrt(tree_mse)
tree_rmse
from sklearn.model_selection import cross_val_score
scores = cross_val_score(tree_reg, housing_prepared, housing_labels,
scoring="neg_mean_squared_error", cv=10)
tree_rmse_scores = np.sqrt(-scores)
def display_scores(scores):
print("Scores:", scores)
print("Mean:", scores.mean())
print("Standard deviation:", scores.std())
display_scores(tree_rmse_scores)
lin_scores = cross_val_score(lin_reg, housing_prepared, housing_labels,
scoring="neg_mean_squared_error", cv=10)
lin_rmse_scores = np.sqrt(-lin_scores)
display_scores(lin_rmse_scores)
from sklearn.ensemble import RandomForestRegressor
forest_reg = RandomForestRegressor()
forest_reg.fit(housing_prepared, housing_labels)
scores = cross_val_score(forest_reg, housing_prepared, housing_labels,
scoring="neg_mean_squared_error", cv=10)
forest_rmse_scores = np.sqrt(-scores)
display_scores(forest_rmse_scores)
from sklearn.model_selection import GridSearchCV
param_grid = [
{'n_estimators': [3, 10, 30], 'max_features': [2, 4, 6, 8]},
{'bootstrap': [False], 'n_estimators': [3, 10], 'max_features': [2, 3, 4]},
]
forest_reg = RandomForestRegressor()
grid_search = GridSearchCV(forest_reg, param_grid, cv=5,
scoring='neg_mean_squared_error')
grid_search.fit(housing_prepared, housing_labels)
grid_search.best_params_
grid_search.best_estimator_
cvres = grid_search.cv_results_
num_attribs = list(housing_num)
cat_attribs = ["ocean_proximity"]
num_pipeline = Pipeline([
('selector', DataFrameSelector(num_attribs)),
('imputer', Imputer(strategy="median")),
('attribs_adder', CombinedAttributesAdder()),
('std_scaler', StandardScaler()),
])
cat_pipeline = Pipeline([
('selector', DataFrameSelector(cat_attribs)),
('label_binarizer', LabelBinarizer()),
])
for mean_score, params in zip(cvres["mean_test_score"], cvres["params"]):
print(np.sqrt(-mean_score), params)
feature_importances = grid_search.best_estimator_.feature_importances_
feature_importances
extra_attribs = ["rooms_per_hhold", "pop_per_hhold", "bedrooms_per_room"]
cat_one_hot_attribs = list(encoder.classes_)
attributes = num_attribs + extra_attribs + cat_one_hot_attribs
sorted(zip(feature_importances, attributes), reverse=True)
final_model = grid_search.best_estimator_
X_test = strat_test_set.drop("median_house_value", axis=1)
y_test = strat_test_set["median_house_value"].copy()
X_test_prepared = full_pipeline.transform(X_test)
final_predictions = final_model.predict(X_test_prepared)
final_mse = mean_squared_error(y_test, final_predictions)
final_rmse = np.sqrt(final_mse)
# => evaluates to 47,766.0
final_rmse
| chapter_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
# +
ipl18 = pd.DataFrame({'Team': ['SRH', 'CSK', 'KKR', 'RR', 'MI', 'RCB', 'KXIP', 'DD'],
'Matches': [14, 14, 14, 14, 14, 14, 14, 14],
'Won': [9, 9, 8, 7, 6, 6, 6, 5],
'Lost': [5, 5, 6, 7, 8, 8, 8, 9],
'Tied': [0, 0, 0, 0, 0, 0, 0, 0],
'N/R': [0, 0, 0, 0, 0, 0, 0, 0],
'Points': [18, 18, 16, 14, 12, 12, 12, 10],
'NRR': [0.284, 0.253, -0.070, -0.250, 0.317, 0.129, -0.502, -0.222],
'For': [2230, 2488, 2363, 2130, 2380, 2322, 2210, 2297],
'Against': [2193, 2433, 2425, 2141, 2282, 2383, 2259, 2304]},
index = range(1,9)
)
ipl18
# +
ipl17 = pd.DataFrame({'Team': ['MI', 'RPS', 'SRH', 'KKR', 'KXIP', 'DD', 'GL', 'RCB'],
'Matches': [14, 14, 14, 14, 14, 14, 14, 14],
'Won': [10, 9, 8, 8, 7, 6, 4, 3],
'Lost': [4, 5, 5, 6, 7, 8, 10, 10],
'Tied': [0, 0, 0, 0, 0, 0, 0, 0],
'N/R': [0, 0, 1, 0, 0, 0, 0, 1],
'Points': [20, 18, 17, 16, 14, 12, 8, 7],
'NRR': [0.784, 0.176, 0.469, 0.641, 0.123, -0.512, -0.412, -1.299],
'For': [2407, 2180, 2221, 2329, 2207, 2219, 2406, 1845],
'Against': [2242, 2165, 2118, 2300, 2229, 2255, 2472, 2033]},
index = range(1,9)
)
ipl17
# -
# Extracting the top 4 teams with their points using label based indexing.
# Since the rows are labelled from 1 to 8, you use '1:4' to extract the first four rows based on labels.
ipl18.loc[1:4, ['Team','Points']]
# Extracting the top 4 teams with their points using position based indexing. Position based indexing starts from 0 by default.
# The columns 'Team' and 'Points' have the index 0 and 6 respectively.
ipl18.iloc[0:4, [0, 6]]
ipl18.loc[(ipl18.NRR > 0) & (ipl18.For > ipl18.Against), ]
ipl18.set_index('Team', inplace = True)
ipl17.set_index('Team', inplace = True)
ipl = ipl18.add(ipl17, fill_value = 0)
ipl
ipl['Win Percentage'] = ipl['Won']/ipl['Matches']
ipl
ipl = ipl.loc[(ipl.Points > 25)].sort_values(by = ['Win Percentage'], ascending = False)
ipl
| python/Pandas+Graded+Questions+Solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1>Cybersecurity Breaches Effects on Stock Price</h1>
#
# By: <NAME>, <NAME>
# <h2>Introduction</h2>
# <h3>Some Background</h3>
# Most literature regarding financial forecasts of a company's future stock price does not include recent cybersecurity breaches as a key factor to take into consideration. Many studies looking at the economic effects of cybersecurity breaches on companies support this reasoning by showing that cybersecurity breaches have relatively <a href="https://meridian.allenpress.com/jis/article-abstract/33/3/227/10584/Much-Ado-about-Nothing-The-Lack-of-Economic-Impact">small economic consequences</a>. With the little effect cybersecurity breaches have on future performances, stock prices are able to rebound to their levels before the breach, with supposedly <a href= "http://web.csulb.edu/colleges/cba/intranet/vita/pdfsubmissions/26629-jis19-much-ado-about-nothing.pdf">no long term changes</a>. However, there is <a href= "https://www.researchgate.net/publication/336947778_The_Impact_of_Reported_Cybersecurity_Breaches_on_Firm_Innovation">research</a> suggesting that there are other long-term consequences like decline in firm productivity, research and development spending, patents, and investment efficiency.
#
# <h3>What are we doing and why?</h3>
# It seems that there is differing information on the long-term effects of cybersecurity breaches on companies. Through this project, we would like to see what the long-term effects of cybersecurity breaches on company stock truly are. Hopefully, we can show that cybersecurity breaches do have lasting consequences to show that it is important to consider breach disclosure in predictive financial models. If we were to prove this, we could bolster better cybersecurity efforts and funding.
#
# <h3>How will we do this?</h3>
# We first focus on the long-term by analyzing the differences between financial analysts’ predictions and actual stock prices a year after breach disclosure. In this long-term analysis, we hope to find a difference between financial analysts predictions and actual stock prices as to prove that predictive financial models are lacking, which hints that they should be considering cybersecurity breaches as important in long-term predictions for stock price. We will also quantify, through an event-study, the short-term impacts of cybersecurity breaches.
#
# <h2>Set Up</h2>
# This will all be done using <a href="https://www.wikiwand.com/en/Python_(programming_language)">Python</a>, leveraging <a href= "https://jupyter.org/">Jupyter Notebooks</a> to visualize various plots, graphs and tables. Here some useful information to install these tools: <a href="https://conda.io/projects/conda/en/latest/user-guide/install/index.html">1</a>, <a href="https://conda.io/projects/conda/en/latest/user-guide/getting-started.html">2</a>, <a href="https://jupyterlab.readthedocs.io/en/stable/getting_started/installation.html">3</a>, <a href="https://www.youtube.com/watch?v=HW29067qVWk">4</a>.
#
# Below are the packages we will use to collect and visualize our data.
# +
# Standard libraries
import pandas as pd
import numpy as np
from scipy import stats
# Visualization
import matplotlib.pyplot as plt
import datetime
import os
# os.system("pip install seaborn")
import seaborn as sns
pd.set_option('display.max_columns', None)
from sklearn.linear_model import LinearRegression
# Data Collection
# os.system("pip install wrds")
import wrds
# os.system("pip install pandas-datareader")
import pandas_datareader.data as web
# -
# <h2>Data Collection</h2>
# We will be using the <a href= "https://wrds-www.wharton.upenn.edu/">WRDS</a> (Wharton Research Data Services) database to find financial analyst predictions on stock price using their <a href= "https://www.investopedia.com/terms/i/ibes.asp">IBES</a> dataset. The WRDS database is provided to all UMD staff and students for free, you can sign up for an account <a href="https://wrds-www.wharton.upenn.edu/register/">here</a>. When running this following section of code, you must supply your account's credentials. We recommend setting up a <a href = "https://matteocourthoud.github.io/post/wrds/"> pgpass</a> to help automate the process.
db = wrds.Connection()
# We will be using the Audit Analytics February 2021 issue of cybersecurity databreaches. To get this dataset, we contacted the University of Maryland's Smith Business School, which provided this Excel file to us. <a href = "https://www.auditanalytics.com/">Audit Analytics</a> is an organization that tracks relevant business and financial data such as the information of a company cybersecurity breach.
#
# We will now load in the data by accessing the correct Excel sheet.
# Audit Analytics Dataset
xls = pd.ExcelFile('../data/audit_analytics.xlsx')
aa_records_df = pd.read_excel(xls, 'PublicCyber')
# Each row in this dataset represents a company's cybersecurity breach. The dataset contains useful information such as the company breached, date of public disclosure of the breach, and other varying information about the company and the type of cybersecurity breach performed.
#
# To clean the data, we need to drop all the rows that don't contain any company tickers. <a href ="https://www.wikiwand.com/en/Ticker_symbol">Tickers</a>, consisting of numbers and letters, are short abbreviations uniquely identifying a publicly traded company (Ex: Amazon is uniquely identified by their ticker, AMZN). If a row doesn't have a ticker symbol, then there may have been data corruption, human errors logging the data, or the company may not be publicly traded. Either way, we need to remove the rows. We also extract the relevant columns for our analysis (as shown in the table_columns array). Once this is all done, it leaves us with a dataset of 737 cybersecurity breaches on publicly traded companies.
# Data Cleaning
aa_records_df = aa_records_df[aa_records_df['Ticker'].isna() != True].reset_index(drop=True)
table_columns = ['Company name', 'Ticker', 'Date of Breach', 'Date Became Aware of Breach', 'Date of Disclosure',
'Number of Records', 'Type of Info', 'Information', 'Attack', 'Region', 'SIC Code']
aa_records_df = aa_records_df[aa_records_df.columns.intersection(table_columns)]
# Now, let's try to find the monthly stock price of each of these firms following the disclosure of the breach.
#
# Before we do that, we define a short helper function that will help us find the closest date in a set that corresponds to X months after the disclosure of a breach. We will make use of this utility in our main function for finding monthly stock prices.
def nearest(items, pivot):
"""
Gets closest day in a set (used to obtain stock price X months after disclosure)
"""
return min(items, key=lambda x: abs((x - pivot).days))
# With that out of the way, let's construct a function to obtain the monthly stock prices after the disclosure of the data breach. Let's break it down!
#
# Our function has two parameters: a row from our original breach dataframe and the number of months to get stock prices from. First, our function determines the range of dates to obtain monthly stock prices. Please note that our starting date is a day before the disclosure breach as to control for any fluctuations in stock price due to that disclosure. Following this, we leverage <a href="https://pandas-datareader.readthedocs.io/en/latest/index.html">pandas_datareader</a>, an <a href="https://rapidapi.com/blog/api-glossary/api-wrapper/#:~:text=In%20that%20line%2C%20an%20API,to%20automate%20API%2Dreliant%20processes.">API wrapper</a> for various API's. Specifically, we will be using its <a href = "https://finance.yahoo.com/">YahooFinance</a> API functionality, which will provide us with a dataframe of stock prices (df) beginning at our start date and ending at our end date. We then traverse through this dataframe, using our nearest helper function, to obtain the monthly stock prices and return them as an array.
#
# If the YahooFinance API cannot find a company's stock price for whatever reason, it returns an array of np.nan's. Likewise, if
# no stock price is available for a month because it's missing or because that month's date lies in the future and hasn't occurred yet, then the array will be populated by np.nan's for those months.
#
# Note: We record the <a href = "https://finance.zacks.com/significance-closing-price-stock-3007.html">closing</a> stock prices, meaning its the stock price at the end of a day. <br>
# Note: The YahooFinance API has a limit of 2,000 requests per hour. As we only have 737 breaches, we won't be hitting that limit, but keep that in mind when using API's.
# +
today = datetime.datetime.today().date()
def stock_after_disclosure(row, num_months):
"""
Returns an array containing the monthly stock price of a firm after date of disclosure (0 - num_months months after breach).
If firm exists in YahooFinance database, but no stock price available for a month (either b/c that date has yet to occur
or b/c simply N/A),
returns np.nan.
If firm does not exist in YahooFinance database, return array of np.nan's.
Parameters:
row : Dataframe row
Input dataframe's row (used along with df.apply)
num_months : int
Month limit
"""
start = pd.to_datetime(row['Date of Disclosure'])
end = start + pd.DateOffset(months = num_months)
start -= datetime.timedelta(days=1)
try:
df = web.DataReader(row['Ticker'], 'yahoo', start, end)
lst = []
for month in range(0, num_months + 1):
if today <= (start + pd.DateOffset(months = month)).date():
for x in range(month, num_months + 1):
lst.append(np.nan)
break
date = nearest(df.index, (start + pd.DateOffset(months = month)))
lst.append(df.loc[date]["Close"])
return lst
except Exception as e:
print("Error at %s" % row['Ticker'])
print(repr(e))
return [np.nan] * (num_months + 1)
# -
# Let's run our function on each row in our dataset. We'll be finding the monthly stock prices spanning a year after the disclosure of the breach.
#
# Note: This section of code takes a while to run (20-30 minutes) because we will be making API requests, loading in data from a server, and performing operations on said data.
# +
lst = []
months_after = 12 #Toggle this value
col = []
# Set the column names for new dataframe
for i in range(0, months_after + 1):
col.append("Stock Price (%s months DoD)" % i)
# Create array of arrays that contains stock prices after date of disclosure for each breach
for index, row in aa_records_df.iterrows():
x = stock_after_disclosure(row, months_after)
lst.append(x)
# -
# Following this, we can concatenate said data to our original dataframe.
# Merge stock price after breach with original dataframe
stock_prices = pd.DataFrame(lst, columns = col)
stock_price_aa_records = pd.concat([aa_records_df, stock_prices], axis=1, join='inner')
# We now have the actual stock prices. Let's move on to finding analyst predictions for these companies.
#
# We define the function below to find the analyst stock price predictions. It makes use of the <a href="https://www.investopedia.com/terms/i/ibes.asp">IBES</a> database in WRDS. The function takes all the financial analyst predictions within a month of the disclosure of the breach that forecast the company's stock price a year into the future. Since multiple financial analysts may make predictions, this function returns the median and mean of these predictions. If no predictions are found, the function returns np.nan's.
#
# Note: This function makes use of SQL, a programming language used to communicate with databases. Here are some helpful resources to get started learning about SQL: <a href = "https://www.codecademy.com/learn/learn-sql">CodeAcademy</a>, <a href = "https://www.khanacademy.org/computing/computer-programming/sql">KhanAcademy</a>
def analyst_stock_price(row):
"""
Returns the median and mean of analyst stock price forecasts for a firm, where the forecasts are within a month after the beach.
These forecasts predict the stock price 12 months into the future.
Parameters
row - Dataframe row
Input dataframe's row (used along with df.apply)
Returns
List of length 2. [median, mean]
"""
date = pd.to_datetime(row['Date of Disclosure'])
sql_query="""
SELECT VALUE as stock_price
FROM ibes.ptgdet
WHERE OFTIC ='{}' AND CAST(HORIZON as int) = 12 AND ANNDATS BETWEEN '{}' and '{}'
""".format(row['Ticker'], date, date + pd.DateOffset(months = 1))
df = db.raw_sql(sql_query)
if len(df.index) == 0:
return [np.nan] * 2
return [df['stock_price'].median(), df['stock_price'].mean()]
# We can now run the function on each company to get the financial analyst forecasts.
# +
# Create array of arrays that contains stock prices after date of disclosure for each breach
lst = []
for index, row in stock_price_aa_records.iterrows():
lst.append(analyst_stock_price(row))
# Merge stock price after breach with original dataframe
median_mean_df = pd.DataFrame(lst, columns = ['median stock forecast', 'mean stock forecast'])
stock_price_aa_records = pd.concat([stock_price_aa_records, median_mean_df], axis=1, join='inner')
# -
# Nice! We have now collected all the data to compare actual stock prices with financial analyst predicted stock prices. But before we do some Exploratory Data Analysis (EDA), we need to do...
# <h2>Data Transformation and Management</h2>
# As it turns out, we might want to transform some of our data relating to stock prices because of innate variation between companies.
#
# To better understand this problem, consider this hypothetical: Suppose company A's and company B's stock price both double after a year. However, company A's initial stock was much smaller, say it started at \\$10 and became \\$20 per share, while company B's stock went from \\$100 to \\$200 per share. Their numerical growth are rather different, but their percent growth are the same. When comparing growth between companies, it makes more sense to compare percent growth since it will better control for the already established stock of that company (whether that be high or low). In a way, finding the percent growth is like standardizing each company's current stock price by their initial stock price.
#
# For that reason, we need to be looking at percent stock price change for these companies, where the initial stock price corresponds to the stock price the day of the disclosure for the breach. The code below transforms the data to percent stock price change for the actual and predicted stock prices.
# +
stock_prices = pd.DataFrame()
n = 1
for x in col[1:]:
stock_prices[n] = stock_price_aa_records.apply(lambda row: (row[x] - row[col[0]])/row[col[0]], axis = 1)
n += 1
t1 = stock_price_aa_records.apply(lambda row: (row['median stock forecast']-row['Stock Price (0 months DoD)'])/row['Stock Price (0 months DoD)'], axis=1)
t2 = stock_price_aa_records.apply(lambda row: (row['mean stock forecast']-row['Stock Price (0 months DoD)'])/row['Stock Price (0 months DoD)'], axis=1)
analysts_percent = pd.concat([stock_prices[12], pd.DataFrame(t1), pd.DataFrame(t2)], axis=1, join='inner')
analysts_percent.columns = ['Actual', 'Analyst Median', 'Analyst Mean']
# -
# <h2>Exploratory Data Analysis</h2>
# To begin, let's make some boxplots and violin plots to get a better understanding of how actual stock prices change over time. We will be making use of the <a href="https://seaborn.pydata.org/index.html">seaborn</a> Python library. We also make use of <a href="https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.melt.html">melting</a> (more info <a href="http://www.jeannicholashould.com/tidy-data-in-python.html">here</a>).
# +
boxplot = sns.boxplot(x="variable", y = "value", data=pd.melt(stock_prices).dropna())
boxplot.set(xlabel="Months after Disclosure", ylabel='Percent Stock Price Change')
boxplot.set_title("Percent Change of Actual Stock Price (Box Plot)")
plt.show()
ax = sns.violinplot(x='variable', y='value', data=pd.melt(stock_prices).dropna())
plt.xlabel('Months after Disclosure')
plt.ylabel('Percent Stock Price Change')
ax.set_title("Percent Change of Actual Stock Price (Violin Plot)")
ax.plot()
plt.show()
# -
# Well...these plots don't really help but why? It seems that there are some major outliers that are making it hard to see how the percent change of actual stock price shifts over time. We have two options here: <a name="two_options"></a>
# 1. Remove the outliers and re-plot the data.
# 2. Find a better metric to represent these distributions over time.
#
# Let's opt to do the second option. There are other metrics to represent these distributions, namely seeing how the "middle" of these distributions change over time. We can define the "middle" of each of these distributions to be the mean or median stock price percent change over time.
#
# Let's take the naive approach of plotting the mean over time.
plt.plot(np.arange(1, 13), stock_prices.mean())
plt.xlabel('Months after Disclosure')
plt.ylabel('Percent Stock Price Change')
plt.title("Percent Change of Actual Stock Price (Mean)")
plt.show()
# It seems like the mean trends upwards over time. This is to say that over time, after public disclosure of a breach, the stock price of companies tend to still trend upwards. It seems that public disclosure of a breach might not have long-term consequences to a company's stock price.
#
# But wait! Let's not forget:
# > <i>"There are three kinds of lies: lies, damned lies, and <b>statistics</b>."</i>
# > \- <NAME>
#
# Means are only good representations of the "middle" of a distribution given that there are <a href = "https://statistics.laerd.com/statistical-guides/measures-central-tendency-mean-mode-median.php#:~:text=When%20not%20to%20use%20the%20mean">no influential outliers and no skewness</a>. A better way of representing the "middle" would be to use the median, which is less affected by outliers and skewness. Let's plot the medians.
plt.plot(np.arange(1, 13), stock_prices.median())
plt.xlabel('Months after Disclosure')
plt.ylabel('Percent Stock Price Change')
plt.title("Percent Change of Actual Stock Price (Median)")
plt.show()
# It seems like when we plot the median over time, the same trend occurs, <b>but</b> it's important to note that the percent stock price percent change values are not as large as the plot of means. Even more important is that the trend no longer seems exactly linear. It seems like the stock price grows slowly at the beginning and then gradually increases later on. This could be representative of a company recovering from the public disclosure of the breach at the start (which causes less stock gains) but after a while the company's growth goes back to normal. We'll look more into the details of this when we perform the event study.
#
# Another way of representing the "middle" is to take the <a href="https://www.investopedia.com/terms/t/trimmed_mean.asp">trimmed mean</a> to get rid of outliers. Below, we took the 5% trimmed means and plotted them. It looks to have the similar trends and observations as the plot of medians.
# Let's take 5% trimmed mean (get rid of some outliers)
trimmed_means = []
for x in range(1, months_after + 1):
trimmed_means.append(stats.trim_mean(stock_prices[x].dropna(), 0.05))
plt.plot(np.arange(1, 13), trimmed_means)
plt.xlabel('Months after Disclosure')
plt.ylabel('Percent Stock Price Change')
plt.title("Percent Change of Actual Stock Price (5% Trimmed Mean)")
plt.show()
# Let's now compare the actual versus financial analyst predictions of the stock price a year after the public disclosure of a cybersecurity breach. We will do this through a violin plot.
violinplot = sns.violinplot(x="variable", y = "value", data=pd.melt(analysts_percent[['Actual', 'Analyst Median', 'Analyst Mean']]).dropna())
violinplot.set(xlabel="Prediction Type", ylabel='Stock Price Percent Change')
violinplot.set_title("Violin Plot of Stock Price Percent Changes")
plt.show()
# Once again, there appears to be some outliers for these distributions, specifically for financial analyst predictions. As mentioned [before](#two_options), there are two options for handling these outliers. We don't really want to use a new metric like the "middle" because we want to compare the actual distributions, so we will instead opt for option one and remove the outliers.
#
# We will be removing the outliers for the financial analyst median and mean predictions. To remove the outliers, we need some rule to label something as an outlier. There are different methods for classifying outliers, but we opt to abide by the <a href="https://www.wikiwand.com/en/68%E2%80%9395%E2%80%9399.7_rule">three-sigma rule</a>, which states that nearly all values are taken to lie within three <a href= "https://www.wikiwand.com/en/Standard_deviation">standard deviations</a> of the mean. If a point were to be beyond three standard deviations from the mean, then we will classify that point as an outlier and remove it from the distribution. The following creates distributions without these outliers.
# +
z_scores = stats.zscore(analysts_percent['Analyst Median'].dropna())
abs_z_scores = np.abs(z_scores)
medians_no_outliers = analysts_percent['Analyst Median'].dropna()[abs_z_scores < 3]
z_scores = stats.zscore(analysts_percent['Analyst Mean'].dropna())
abs_z_scores = np.abs(z_scores)
means_no_outliers = analysts_percent['Analyst Mean'].dropna()[abs_z_scores < 3]
# -
# Let's now create a new violin plot without the outliers.
violinplot = sns.violinplot(x="variable", y = "value", data=pd.melt(pd.concat([analysts_percent['Actual'],medians_no_outliers, means_no_outliers], axis=1)))
violinplot.set(xlabel="Prediction Type", ylabel='Stock Price Percent Change')
violinplot.set_title("Violin Plot of Stock Price Percents (Analyst Outliers Removed)")
plt.show()
# This violin plot is way more legible than the previous. It seems from this violin plot that the financial analyst predictions tend to vary more than the actual stock prices. It also hints that financial analysts tend to overestimate the actual stock price of these companies. A better method of visualizing these differences is to look at the residuals of these stock prices, where the residual is the actual minus the predicted stock price percent change.
#
# The following code will compute the residuals for each prediction type and plot them. Note that for this part, the outlier financial analyst predictions have been kept.
# +
analysts_percent['Median Residual'] = analysts_percent.apply(lambda row: row['Actual'] - row["Analyst Median"], axis =1)
analysts_percent['Mean Residual'] = analysts_percent.apply(lambda row: row['Actual'] - row["Analyst Mean"], axis =1)
violinplot = sns.violinplot(x="variable", y = "value", data=pd.melt(analysts_percent[['Median Residual', 'Mean Residual']]))
violinplot.set(xlabel="Prediction Type", ylabel='Stock Price Percent Change')
violinplot.set_title("Violin Plot of Stock Price Percent Residuals")
plt.show()
# -
# It seems the distribution of these different residuals appear very similar. It also appears that these distributions center around 0 but are skewed towards the negative end. For this to happen, it means that financial analyst predictions are greater than the actual stock prices.
#
# For a better look, here's a display the summary statistics for each residual distribution. I would like to note that our initial sample of cybersecurity breaches was 737, but it has now shrunk down to a set of 474 breaches. This could either be due to the Yahoo Finance API not containing stock prices for certain companies, IBES not having predictions for smaller companies, or a year hasn't elapsed since the public disclosure of the cybersecurity breach.
print(analysts_percent['Median Residual'].describe())
print()
print(analysts_percent['Mean Residual'].describe())
# It does seem that financial analysts overestimate the actual stock prices for these firms that had recently issued public disclosures of cybersecurity breaches. We need a to perform a more scientific/mathematical study to conclude this. This takes us to our next section...
# <h2>Hypothesis Testing</h2>
# We want to perform a statistical test to confirm our findings that the financial analysts overestimate the actual stock price. Specifically, we want a test that proves that these residuals we've found skew towards negative. In other words, we want to prove that the mean ($\mu$) of the distribution of residuals is negative and that these results are statistically significant, meaning there is little to no doubt that the mean is 0.
#
# The best test for this situation would be a 1-sample t-test, specifically a <a href= "https://www.statisticssolutions.com/manova-analysis-paired-sample-t-test/#:~:text=The%20paired%20sample%20t%2Dtest,resulting%20in%20pairs%20of%20observations.">paired sample t-test</a>. To get an understanding of how this test works, we need to understand what a <a href = "https://support.minitab.com/en-us/minitab/18/help-and-how-to/statistics/basic-statistics/supporting-topics/basics/null-and-alternative-hypotheses">null and alternative hypothesis</a> are. The null hypothesis states that a population parameter is equal to a hypothesized value. The alternative hypothesis states that a population parameter is different from the hypothesized value posited in the null hypothesis. These two hypotheses are mutually exclusive (if one is true, the other is false). In this case, our hypotheses are:
#
# - Null Hypothesis: $\mu = 0$
# - Alternative Hypothesis: $\mu < 0$
#
# The null hypothesis roughly translates to the mean of the distribution of residuals is equal to 0, meaning that we expect no difference between the actual and predicted stock prices. The alternative hypothesis roughly translates to the mean of the distribution of residuals is less than 0, meaning that the predicted stock prices tend to be greater than the actual stock prices.
#
# In this statistical test, we begin with assuming the null hypothesis is true. We then study our sample dataset (through statistical methods) to see if given the null hypothesis is true, the sample dataset could reasonably occur. If our data could not reasonably happen under the premise of the null hypothesis, then we reject the null hypothesis and assume it's false. Since the null hypothesis is false in these circumstances, we would have to accept the alternative hypothesis as true. In the case that the data is not significantly opposed to the premise of the null hypothesis, we fail to reject the null hypothesis - meaning we have no evidence to believe its contrary. This is the essence of most statistical hypothesis testing. For a more through explanation, click <a href= "https://www.statisticshowto.com/probability-and-statistics/hypothesis-testing/">here</a>.
#
# Before we can perform this test, we need to meet certain prerequisites. We need to meet the three assumptions: independence of observations, approximately normal distributions, and no major outliers. We can reasonably assume independence of observations since one company’s residual does not affect another’s. From the violin plot, it seems that the distribution is approximately normal and to meet the assumption of no major outliers, we will remove outliers from the dataset when performing the test. Depicted below is a violin plot of the residuals with major outliers removed.
# +
z_scores = stats.zscore(analysts_percent['Median Residual'].dropna())
abs_z_scores = np.abs(z_scores)
no_outliers_median = analysts_percent['Median Residual'].dropna()[abs_z_scores < 3]
z_scores = stats.zscore(analysts_percent['Mean Residual'].dropna())
abs_z_scores = np.abs(z_scores)
no_outliers_mean = analysts_percent['Mean Residual'].dropna()[abs_z_scores < 3]
violinplot = sns.violinplot(x="variable", y = "value", data=pd.melt(pd.concat([no_outliers_mean, no_outliers_median], axis=1)))
violinplot.set(xlabel="Prediction Type", ylabel='Stock Price Percent Change')
violinplot.set_title("Violin Plot of Stock Price Percent Residuals (No Outliers)")
plt.show()
# -
# We also have to establish a <a href="https://blog.minitab.com/en/adventures-in-statistics-2/understanding-hypothesis-tests-significance-levels-alpha-and-p-values-in-statistics">significance level</a> before running the test. We choose a significance level of 5%, meaning if a sample has a less than 5% chance of occurring given the null hypothesis, then we will reject the null hypothesis and accept the alternative.
#
# The following code will output the p-values for the median and mean residual hypothesis tests respectively. Outliers have been removed for this test.
#
# Note: We are performing a <a href = "https://www.statisticssolutions.com/should-you-use-a-one-tailed-test-or-a-two-tailed-test-for-your-data-analysis/#:~:text=This%20is%20because%20a%20two,groups%20in%20a%20specific%20direction.">one-tailed test</a>, so we will halve the p-values.
print("Median residual p-value: %s" % (stats.ttest_1samp(a=no_outliers_median, popmean=0).pvalue/2))
print("Mean residual p-value: %s" % (stats.ttest_1samp(a=no_outliers_mean, popmean=0).pvalue/2))
# These p-values are extremely small, smaller than our significance level of 5%. Since these p-values are extremely small, the chance that these samples could have occurred given the null hypothesis are nearly impossible, so we reject the null hypothesis and accept that alternative hypothesis that $\mu$ < 0. We can now conclude that financial analysts tend to overestimate the stock price of a company after the public disclosure of a cybersecurity breach.
# <h3>Aside: What if we hadn't removed the outliers for the residuals?</h3>
# Suppose we hadn't removed the residuals and still ran the t-tests as is. We would have gotten these p-values:
print("Median residual p-value: %s" % (stats.ttest_1samp(a=analysts_percent['Median Residual'].dropna(), popmean=0).pvalue/2))
print("Mean residual p-value: %s" % (stats.ttest_1samp(analysts_percent['Mean Residual'].dropna(), popmean=0).pvalue/2))
# We would have still rejected the null hypothesis and accepted the alternative, but we would have to say that a major caveat was that we don't meet all the assumptions to run this test.
# <h1>Event-Study</h1>
# ## Introduction
#
# Our event study seeks to establish the effect of breaches on the stock price of affected companies. To measure this effect, we analyze the abnormal returns ($AR_{i,t}$), the actual returns ($r_{i,t}$) minus the normal returns ($NR_{i,t}$), in the aftermath of a breach announcement. Actual returns, $r_{i,t}$, are the real stock price changes, measured as $(p_{i,t} - p_{i,t-1}) / p_{i,t-1}$, where $p_{i,t}$ is the real adjusted stock price of firm $i$ on day $t$. The normal returns is what would have been the stock return of firm $i$ on day $t$, barring the occurrence of the breach event. The normal returns needs to be estimated using a model, because it is hypothetical.
#
# There are a variety of different models for normal returns, and they can generally be classified into two types: statistical and economic models. We chose to use statistical models because they offer good performance for their simplicity.
#
# Out of the statistical models, there are two major types: constant mean model and market model. A constant mean model takes the mean average of a firm's returns over the estimation period and uses this mean for all normal returns. A market model builds upon this concept, and creates a linear model that relates the market return (S\&P 500 returns) to the firm's return. The constant mean model can be viewed as market model with the market coefficient $\beta=0$.
#
# Market models make the following assumptions:
# * Stock returns are normally distributed
# * Stock returns are linearly correlated with market returns (S\&P500 returns)
# * The linear relationship between the stock returns and the market returns is the same throughout the estimation period.
# Although these assumptions may seem strong, in practice, these assumptions don’t present significant challenges. Further, market models are used widely in event studies, and are considered a powerful tool for estimating stock returns. For that reason, we opt to develop a market model.
# ### Data Loading
#
# We use the `stock_indicators.csv` file, containing the cybersecurity breaches and company stock prices that we prepared in the previous section.
df = pd.read_csv("../data/stock_indicators.csv")
date_cols= ["Date Became Aware of Breach", "Date of Disclosure"]
df[date_cols] = df[date_cols].apply(lambda x: pd.to_datetime(x, errors="coerce"))
df = df.dropna(subset=['Ticker'])
print("Date of Disclosure NAs: ", df["Date of Disclosure"].isna().sum())
print("Ticker NAs: ", df["Ticker"].isna().sum())
# Fix Ticker substocks
df["Ticker"] = df["Ticker"].apply(lambda x: x.replace(".", "-"))
df.head(50)
# Here, we define the estimation window and event windows for our analysis. The `max_normal_range` denotes the length of the maximum estimation window. We will retrieve `max_normal_range` days worth of stock prices before the event, but we won't neccessarily use. Similarly, the `max_event_range` denotes the days of stock prices we retrieve for the event window.
# +
max_normal_range = 1000
max_event_range = 10
earliest_breach_date = min(df["Date of Disclosure"])
latest_breach_date = max(df["Date of Disclosure"])
sp_start = earliest_breach_date - pd.to_timedelta(max_normal_range, unit="D")
sp_end = date.today()
# -
# We use the S&P 500 index as our market basket. Individual stock price's performance would be compared against this market index
sp500 = yf.download("^GSPC", start=sp_start, end=sp_end)
sp500.head()
sns.lineplot(data=sp500, x="Date", y="Adj Close")
# +
from contextlib import contextmanager
import sys, os
# This context is used to suppress output by redirecting it to null
# Some of the libraries used output verbose logs
@contextmanager
def suppress_stdout():
with open(os.devnull, "w") as devnull:
old_stdout = sys.stdout
sys.stdout = devnull
try:
yield
finally:
sys.stdout = old_stdout
# -
# ### Market Model
#
# Here, we implement the market model to predict the normal returns of the affect companies' stock price.
#
# The normal returns are the returns of a stock that would have happened without the breach. This market model consists of a regression with the following form:
# $$r_{i,t} = \alpha_i + \beta_i * r_{m,t} + \epsilon_{i,t}$$
#
# where $r_{i,t}$ is firm $i$'s normal return and $r_{m,t}$ is the actual market return. $\alpha_i$ and $\beta_i$ are the linear constant and coefficient, respectively, and $\epsilon_{i,t}$ is the regression error term. Our time increments are done by business days, so if the date of breach, $t=0$, is 4/23/2021 (Friday), then $t=1$ would be 4/26/21 (Monday). We then use this model to estimate $NR_{i,t}$, using $ \alpha_i + \beta_i * r_{m,t}$
#
# Our market model estimates, which describes the daily percentage changes in stock price. To measure the the full impact of a breach, we need to collect these percentage changes over a period. We track the Cumulative Average Return (CAR) within +/- 7 business days of the breach. The $CAR_i(a,b)$ is defined as the following:
#
# $$CAR_i(a,b) = \sum_{t=a}^{b}{AR_{i,t}}$$
#
# For small intervals, the CAR serves as a good aggregate of abnormal returns. For longer intervals, the abnormal returns would compound, causing the real stock price to differ from the cumulative returns.
# +
def window_date(date: datetime.datetime, window=(0, 0), business=True):
if business:
return (
date + BDay(window[0]),
date + BDay(window[1])
)
return (
date + pd.to_timedelta(window[0], unit="D"),
date + pd.to_timedelta(window[1], unit="D")
)
def date_range(df, window_dates: Tuple[datetime.datetime, datetime.datetime]):
return df.loc[(df.index >= window_dates[0]) & (df.index <= window_dates[1])]
def market_model(stock, sp500, window_date):
# Normal Model Estimation
window_sp500 = date_range(sp500, window_date)
window_stock = date_range(stock, window_date)
# pct_change - first entry is always nan
sp500_returns = window_sp500["Adj Close"].pct_change()[1:].values.reshape(-1, 1) # "X"
stock_returns = window_stock["Adj Close"].pct_change()[1:].values.reshape(-1, 1) # "Y"
joined = pd.DataFrame({
"sp500": window_sp500["Adj Close"],
"stock": window_stock["Adj Close"],
})
if joined.isnull().values.any():
print("Missing values for regression")
joined = joined.dropna()
if len(joined) == 0:
raise ValueError("No stock values available. Ticker not available?")
joined["sp500_returns"] = joined["sp500"].pct_change()
joined["stock_returns"] = joined["stock"].pct_change()
sp500_returns = joined["sp500_returns"].values[1:].reshape(-1, 1)
stock_returns = joined["stock_returns"].values[1:].reshape(-1, 1)
# Join the returns
reg = LinearRegression()
try:
reg.fit(
sp500_returns,
stock_returns
)
(alpha, beta) = reg.intercept_.item(), reg.coef_.item()
except Exception as e:
# print("<=========================== [Regression Error] ===========================>")
# print(sp500_returns.shape)
# print(stock_returns.shape)
# print(joined.head())
# print("<=========================== [Regression Error] ===========================>")
raise ValueError("Model failed to fit")
stock_returns_pred = reg.predict(sp500_returns)
mse = mean_squared_error(stock_returns, stock_returns_pred)
r2 = r2_score(stock_returns, stock_returns_pred)
# Reformatted
sp_ret = sp500_returns.ravel()
st_ret = stock_returns.ravel()
sp_ret = np.insert(sp_ret, 0, np.nan)
st_ret = np.insert(st_ret, 0, np.nan)
dat = pd.DataFrame({
"sp500": window_sp500["Adj Close"],
"window_stock": window_stock["Adj Close"],
"sp500_returns": sp_ret,
"stock_returns": st_ret,
}, index=window_sp500["Adj Close"].index)
return {
"alpha": alpha,
"beta": beta,
"mse": mse,
"r2": r2,
"data": dat
}
# -
# ### Stock price retrieval
#
# In this function, we use the specified estimation and event windows to retrieve the stock prices of interest for every company. We use `yfinance` library to retrieve stock prices by ticker and date from Yahoo Finance, a stock price database. After retrieving the relevant stock prices for a company, we execute. `market_model()` to find parameters alpha and beta. We cache the stock prices, markel parameters, normal returns and abnormal returns into a dictionary for every breach event.
# +
saves = {}
def normal_return_model(row, est_window=(-200, -7), event_window=(-7, 7), windows = [], coerce=True, debug=True):
if est_window[0] > est_window[1] or event_window[0] > event_window[1]:
raise ValueError("Invalid Window: Windows must be in form (a, b), a < b")
if est_window[1] > event_window[0]:
raise ValueError("Overlapping windows!")
ticker = row["Ticker"]
disclosure_date = row["Date of Disclosure"]
# Convert windows to date windows
est_window_date = window_date(disclosure_date, est_window)
event_window_date = window_date(disclosure_date, event_window)
start_date = est_window_date[0]
end_date = event_window_date[1]
# Ensure dates are valid
today_date = datetime.date.today()
if end_date > today_date:
if coerce:
end_date = today_date
else:
raise ValueError(
f"Ending interval of {end_date.strftime('%Y-%m-%d')} is later than today {today_date.strftime('%Y-%m-%d')}"
)
if debug:
print(f"Ticker: {row['Ticker']}")
print(f"Date of Disclosure: {row['Date of Disclosure'].strftime('%Y-%m-%d')}")
print(f"estimation window: {est_window_date}")
print(f"event window: {event_window_date}")
# Download
try:
with suppress_stdout():
raw_prices = yf.download(ticker, start=start_date, end=end_date)
raw_prices.dropna(axis = 0, how = 'all', inplace = True)
if len(raw_prices) == 0:
raise ValueError("Download Failed.")
except Exception as e:
raise ValueError('Download Failed.')
# Linear Regression
try:
results = market_model(raw_prices, sp500, est_window_date)
saves["results"] = results
alpha, beta = results["alpha"], results["beta"]
except Exception as e:
raise ValueError('Model fit failed.')
# Draw
if debug:
model_df = results["data"]
saves["model_df"] = model_df
# Event interval
event_df = pd.DataFrame()
event_df["actual"] = date_range(raw_prices, event_window_date)["Adj Close"].pct_change()[1:]
event_df["sp500"] = date_range(sp500, event_window_date)["Adj Close"].pct_change()[1:]
event_df["normal"] = alpha + beta * event_df["sp500"]
event_df["abnormal"] = event_df["actual"] - event_df["normal"]
# Compute CARs
CAR = {}
for w in windows:
dates = window_date(disclosure_date, w)
CAR[w] = date_range(event_df["abnormal"], dates).sum()
return {
"market_model": results,
"event_df": event_df,
"CAR": CAR,
}
normal_return_model(df.iloc[0], windows = [(0, 1), (0, 2), (0, 3), (0, 4), (0, 5)], debug=False)
# -
# We execute the `normal_return_model()`, which downloads and runs the market model regression, over all companies in the breach database. For some stocks, our downloader might fail because the ticker is malformed or because our stock database (Yahoo Finance) might not have the stock.
# +
import traceback
CARs = []
windows = [(0, i) for i in range(5)]
windows.extend([(-i, 0) for i in range(5)])
windows.extend([(-i, i) for i in range(5)])
print(windows)
arr = []
raw_results = []
for i, row in tqdm.tqdm(df.iterrows(), total=len(df)):
try:
result = normal_return_model(row, windows = windows, debug=False)
car = result['CAR']
obj = {
'alpha': result['market_model']['alpha'],
'beta': result['market_model']['beta']
}
obj.update({f'CAR{k}': v for k, v in car.items()})
arr.append(obj)
raw_results.append(result)
except Exception as e:
# print("<=============== Error ===============>")
# print(f"Row #: {i}")
# print(f"Ticker: {row['Ticker']}")
# print("<================ End ================>")
# traceback.print_exc()
arr.append({})
raw_results.append({})
assert len(arr) == len(df)
# -
# After running this regression, we save the Cumulative Abnormal Returns (CAR) into a seperate csv file and pickle our auxillary data into `raw_results.pkl`. This serves as a good checkpoint, as downloading and running the regressions takes a substantial amount of time.
data_df = pd.DataFrame(arr)
data_df.head(50)
data_df.to_csv("../data/car_window_multi.csv")
# +
import pickle
pkl_path = "../data/raw_results.pkl"
pickle.dump(raw_results, open(pkl_path , "wb"))
# -
# ### Reloading the data
#
import pickle
pkl_path = "../data/raw_results.pkl"
raw_results = pickle.load(open(pkl_path , "rb"))
data_df = pd.read_csv("../data/stock_indicators.csv")
df = pd.read_csv("../data/car_window_multi.csv")
# We join the results of the run back into our original dataframe (with company ticker and other related info)
df = pd.concat((df, data_df), axis=1)
df.head()
# ### Individual Company Analysis
#
# In this section, we analyze the financial impact of a particular breach, the SolarWinds breach, in detail.
#
# On December 14, 2020, SolarWinds publically announced that it experienced a major breach in its "Orion" system. Solarwinds is an information technology company that supplies software to major firms and government organizations. Through this breach, the actors were able to gain access to many organization's IT systems, allowing them to install further malware. Over 18,000 companies were affected - including Fortune 500 companies like Microsoft and government organizations like the Pentagon.
#
# We plot the market model and the Cumulative Abnormal Returns (CAR) for Solarwind during this breach.
# +
# Find indexes by company name (Equifax)
# df.index[df['Ticker'] == 'EFX']
np.where(df['Ticker'] == 'SWI')
# Equifax hack: 409
# Solarwinds: 43
# -
raw_results[0].keys()
result_df.head()
df.iloc[43].to_dict()
# #### General Company Data
# +
index = 43
# Need to match raw_results with table???
market_model_data = raw_results[index]["market_model"]
model_df = market_model_data["data"]
alpha = market_model_data["alpha"]
beta = market_model_data["beta"]
company_name = df.iloc[index]["Company name"]
company_ticker = df.iloc[index]["Ticker"]
breach_date = df.iloc[index]["Date of Disclosure"]
event_df = raw_results[index]["event_df"]
print(f"i:{index} > {company_name} ({company_ticker}), breach date: {breach_date.date()}")
print(f"Market Model: Y = {alpha:.3f} + {beta:.3f} * X")
model_df.head()
# -
# #### Market Model Return History (Estimation)
#
# Here, we compare the returns of SolarWinds alongside the S&P 500 index. The SolarWinds stock has higher variance than the S&P 500, as the S&P 500 is an aggregate over many different companies (and therefore diversified).
#
# +
estimation_return = pd.melt(model_df[["sp500_returns", "stock_returns"]], ignore_index=False)
melted.head()
# filtered.tail(100)
plt.figure(figsize=(8,5))
ax = sns.lineplot(data=estimation_return, x=estimation_return.index, y="value", hue="variable")
ax.set_title(f"{company_name} Adjusted Stock Return vs. S&P500 Returns (Estimation Period)")
ax.set_ylabel("Returns")
plt.xticks(rotation=45)
ax.legend(loc=1)
plt.show()
# +
sp500_returns = model_df["sp500_returns"]
stock_returns = model_df["stock_returns"]
plt.figure(figsize=(8,5))
# Display regression line
X = sp500_returns.values
Y = alpha + beta * X
ax = sns.lineplot(x=X, y=Y, label=f'Market Model: Y = {alpha:.3f} + {beta:.3f} * X')
# ----------------------------
sns.scatterplot(data=model_df, x="sp500_returns", y="stock_returns", color="black", alpha=0.5)
ax.legend()
ax.set_title(f"{company_name} Regression Model: Adjusted Stock Return vs. S&P500 Returns")
ax.set_ylabel("Stock Returns")
ax.set_xlabel("S&P 500 Market Returns")
# -
# In this regression plot, we pair each day's S&P 500 return with SolarWind's return. The data satisfies most of linear regression's assumptions:
#
# - The plot shows a clear linear relationship
# - The residuals have mostly constant variance across all values
# - The residuals are mostly normally distributed
#
# For the independence assumption, which suggests that data points must be independent, may be slightly violated as stock returns are correlated with one another temporally. However, over a large window, these violations do not affect the model by much, and research into market models suggest that linear regression remains an effective tool for estimating normal returns.
# #### Event Analysis
diff = (pd.Series(event_df.index - breach_date, index = event_df.index))
event_df["day_diff"] = diff
event_df.head()
plt.figure(figsize=(8,5))
event_return = pd.melt(event_df[['sp500', 'actual']], ignore_index=False)
ax = sns.lineplot(data=event_return, x=event_return.index, y="value", hue="variable")
ax.set_title(f"{company_name} Adjusted Stock Return vs. S&P500 Returns (Event Period)")
ax.set_ylabel("Returns")
plt.xticks(rotation=45)
ax.legend(loc=1)
plt.show()
# In this plot, we notice a substantial deviation in stock price proceeding the breach event. The company's stock price stopped over `15%` in two of the event window days. The S&P500 index remained stable across the week.
# +
f = pd.concat([estimation_return, event_return], axis=0)
plt.figure(figsize=(12, 8))
ax = sns.lineplot(data=f, x=f.index, y="value", hue="variable")
ax.set_title(f"{company_name} Adjusted Stock Return vs. S&P500 Returns (Full Period)")
ax.set_ylabel("Returns")
plt.xticks(rotation=45)
ax.legend(loc=3)
plt.show()
# -
# Here, we plot the S&P 500 and SolarWinds stock prices across the estimation **and** event window. Notice how the SolarWind's stock price deviated substantially from the S&P500 during this period. This serves as supporting evidence (but not sufficient) that it was the breach, and not market wide downturns, that caused this drop in stock price.
car_data = date_range(event_df, window_date(breach_date, CAR_range)) # Get the range of abnormal returns of interest
car_data.head()
# +
# Plotted with abnormal returns
plt.figure(figsize=(10,5))
ax = sns.lineplot(x=X, y=Y, label=f'Market Model: Y = {alpha:.3f} + {beta:.3f} * X')
sns.scatterplot(data=model_df, x="sp500_returns", y="stock_returns", color="black", alpha=0.2, label="Actual Returns [Estimation]")
sns.scatterplot(data=car_data, x="sp500", y="actual", color="red", alpha=0.7, label=f"Actual Returns [Breach ({breach_date.date()})]")
ax.legend(loc=3)
ax.set_title(f"{company_name}: Actual Returns vs. S&P500 Returns")
ax.set_ylabel("Stock Returns")
ax.set_xlabel("S&P 500 Market Returns")
for line in range(0,car_data.shape[0]):
ax.text(car_data["sp500"][line] + 0.0005, car_data["actual"][line],
f"+{car_data['day_diff'][line].days} days", horizontalalignment='left',
size='medium', color='black')
plt.show()
# -
# In this plot, we impose the event window returns on our market model. Once again, we observe substantial deviations of SolarWind's stock price relative to our market model, which is based on the S&P 500.
# ### CAR Plots
#
# Here, we analyze the Cumulative Abnormal Returns (CAR) over the entire database of breaches. We would like to detect whether there was a statistically signficant change in stock price after the breach.
data_df.describe()
# sns.histplot(data=data_df, x="CAR(0, 3)")
data_df.head()
data_df.columns
post_car_columns = ['CAR(0, 0)', 'CAR(0, 1)', 'CAR(0, 2)', 'CAR(0, 3)', 'CAR(0, 4)']
pre_car_columns = ['CAR(-1, 0)', 'CAR(-2, 0)', 'CAR(-3, 0)', 'CAR(-4, 0)']
wide_car_columns = ['CAR(-1, 1)', 'CAR(-2, 2)', 'CAR(-3, 3)', 'CAR(-4, 4)']
post_car = data_df[post_car_columns]
pre_car = data_df[pre_car_columns]
wide_car = data_df[wide_car_columns]
# +
plt.figure(figsize=(6,5))
A = 2
B = 2
fig, axs = plt.subplots(A, B, figsize=(6 * A, 5 * B))
fig.tight_layout()
plt.subplots_adjust(hspace=0.2)
for ax, col in zip(axs.flat, post_car_columns):
ax = sns.histplot(data_df[col], ax=ax)
ax.set_title(col)
ax.set_xlabel("Cumulative Average Return")
ax.set_ylabel("Frequency")
plt.show()
# -
# The cumulative returns over different date windows follows a normal distribution. Based on these histograms, there is a slight shift of the Cumulative Average Returns towards the negative.
import scipy
f = pd.DataFrame({name: scipy.stats.ttest_1samp(data.dropna().values, 0) for (name, data) in post_car.iteritems()})
f.insert(0, 'Description', ["t-statistic", "p-value"])
print(f.to_latex())
{name: scipy.stats.ttest_1samp(data.dropna().values, 0) for (name, data) in post_car.iteritems()}
# The t-tests of the CARs across the (0, 0) to (0, 4) windows suggests that the deviations are statistically significant. Four of the five windows show a p-value that is under an alpha of 5%. This supports our initial hypothesis that breaches would negatively affect a company's returns.
# <h1>Conclusion</h1>
# We have established that there are short-term and long-term consequences to stock price when cybersecurity breach disclosures occur. For short-term, we proved this through an event study, while for long-term, we used t-tests. We hope that these results will guide companies to be more wary of funding cybersecurity initiatives.
| notebooks/merge_final.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
import PIL
import torch, torchvision
import numpy as np
rng = np.random.default_rng()
import time
from tqdm.notebook import tqdm ### suitable for notebooks
# +
fname = 'EMNIST_torch'
### Download and initialize datasets
TrainDS_orig = torchvision.datasets.EMNIST(fname, train=True, download=True, split='letters', transform=torchvision.transforms.Compose([
lambda img: torchvision.transforms.functional.rotate(img, -90),
lambda img: torchvision.transforms.functional.hflip(img)
]))
TestDS_orig = torchvision.datasets.EMNIST(fname, train=False, split='letters', transform=torchvision.transforms.Compose([
lambda img: torchvision.transforms.functional.rotate(img, -90),
lambda img: torchvision.transforms.functional.hflip(img)
]))
# -
# A - 1
#
# B - 2
#
# .
# .
# .
# .
# +
# Size of dataset
print(len(TrainDS_orig)) #124800
# Details of dataset
print(TrainDS_orig)
## Show the pixel values of an image from training data
# print(np.asarray(TrainDS_orig[0][0].getdata()))
## Calculate mean alternative 1
tot = 0.0
for inputs,labels in TrainDS_orig:
tot = tot + np.asarray(inputs.getdata())/255
mean = tot.mean()/len(TrainDS_orig)
print(tot.mean()/len(TrainDS_orig))
## Calculate mean alternative 2
tot = 0.0
for inputs,labels in TrainDS_orig:
tot = tot + np.asarray(inputs.getdata()).mean()/255
print(tot/124800)
## Calculate standard deviation
tot = 0.0
for inputs,labels in TrainDS_orig:
tot = tot + (np.asarray(inputs.getdata()).mean()/255 - mean)**2
std = np.sqrt(tot/len(TrainDS_orig))
print(np.sqrt(tot/len(TrainDS_orig)))
# -
### Plot examples
fig, axes = plt.subplots(nrows=2, ncols=6, figsize=(15., 6.))
for axes_row in axes:
for ax in axes_row:
test_index = rng.integers(0, len(TestDS_orig))
image, orig_label = TestDS_orig[test_index]
ax.set_axis_off()
ax.imshow(image)
# ax.imshow(PIL.ImageOps.mirror(image.rotate(-90)))
ax.set_title('True: %i' % orig_label)
# +
### Define the label transform from an integer to a set of probabilities
def target_transform(inlabel):
newlabel = torch.zeros(27)
newlabel[inlabel] = 1.
return newlabel
def input_transform(image):
image = torchvision.transforms.functional.rotate(image,-90)
image = torchvision.transforms.functional.hflip(image)
tensor = torchvision.transforms.functional.to_tensor(image)
return tensor
### Reinitialize datasets with the transforms
TrainDS = torchvision.datasets.EMNIST(fname, train=True, download=True,
target_transform=target_transform, transform=torchvision.transforms.Compose([
lambda img: torchvision.transforms.functional.rotate(img, -90),
lambda img: torchvision.transforms.functional.hflip(img),
torchvision.transforms.ToTensor(),
torchvision.transforms.Normalize((0.1722273), (0.0521201))
]), split='letters')
TestDS = torchvision.datasets.EMNIST(fname, train=False,
target_transform=target_transform, transform=torchvision.transforms.Compose([
lambda img: torchvision.transforms.functional.rotate(img, -90),
lambda img: torchvision.transforms.functional.hflip(img),
torchvision.transforms.ToTensor(),
torchvision.transforms.Normalize((0.1722273), (0.0521201))
]), split='letters')
### Initialize DataLoaders as PyTorch convenience
TrainDL = torch.utils.data.DataLoader(TrainDS, shuffle=True, batch_size=200)
TestDL = torch.utils.data.DataLoader(TestDS, batch_size=200)
### Choose device: 'cuda' or 'cpu'
device = 'cpu'
# device = 'cuda'
### Define the dense neuron layer
# Network = torch.nn.Sequential(
# torch.nn.Flatten(), # 28x28 -> 784
# torch.nn.Linear(784, 10), # 784 -> 10
# torch.nn.Softmax(dim=1)
# )
Network = torch.nn.Sequential(
torch.nn.Flatten(),
torch.nn.Linear(784, 400),
torch.nn.ReLU(),
torch.nn.Linear(400, 27),
torch.nn.Softmax(dim=1)
)
Network.to(device=device)
### Get information about model
totpars = 0
for par in Network.parameters():
newpars = 1
for num in par.shape:
newpars *= num
totpars += newpars
print(Network)
print('%i trainable parameters' % totpars)
### Initialize loss function and optimizer
crit = torch.nn.BCELoss()
opt = torch.optim.SGD(Network.parameters(), lr=0.6)
# -
### Baseline: just say it's anything at probability 1/N, what's the loss?
N = 27
labels = torch.zeros(1, 27, dtype=torch.float32)
labels[0, 3] = 1.
output = torch.full_like(labels, 1./N)
print(crit(output, labels))
# +
### Set model in training mode and create the epochs axis
Network.train()
epochs = range(1, 40)
### Train the model
for e in tqdm(epochs):
tr_loss = 0.
samples = 0
### Loop over batches
for inputs, labels in tqdm(TrainDL, leave=False):
opt.zero_grad() # zero gradient values
inputs = inputs.to(device=device) # move input and label tensors to the device with the model
labels = labels.to(device=device)
outputs = Network(inputs) # compute model outputs
loss = crit(outputs, labels) # compute batch loss
loss.backward() # back-propagate the gradients
opt.step() # update the model weights
tr_loss += loss.clone().cpu().item()*len(inputs) # add the batch loss to the running loss
samples += len(inputs) # update the number of processed samples
tr_loss /= samples # compute training loss
print(e, tr_loss)
# +
### Set model in evaluation mode
Network.eval()
### Compute the test loss
with torch.no_grad():
te_loss = 0.
samples = 0
accuracy = 0
### Loop over batches
for inputs, labels in tqdm(TestDL):
inputs = inputs.to(device=device)
labels = labels.to(device=device)
outputs = Network(inputs)
loss = crit(outputs, labels)
te_loss += loss.clone().cpu().item()*len(inputs)
accuracy += torch.sum(torch.eq(torch.max(labels, 1)[1], torch.max(outputs, 1)[1]), dtype=int).clone().cpu().item()
samples += len(inputs)
te_loss /= samples
accuracy /= samples
print('Test loss: %f, accuracy: %f' % (te_loss, accuracy))
# -
torch.save(Network, 'model_torch_EMNIST_letters.chk')
Network = torch.load('model_torch_EMNIST_letters_0.chk')
device='cpu'
### Draw some random images from the test dataset and compare the true labels to the network outputs
fig, axes = plt.subplots(nrows=2, ncols=6, figsize=(15., 6.))
### Loop over subplots
for axes_row in axes:
for ax in axes_row:
### Draw the images
test_index = rng.integers(0, len(TestDS))
sample, label = TestDS[test_index]
image, orig_label = TestDS_orig[test_index]
### Compute the predictions
with torch.no_grad():
output = Network(torch.unsqueeze(sample, dim=0).to(device=device))
certainty, output = torch.max(output[0], 0)
certainty = certainty.clone().cpu().item()
output = output.clone().cpu().item()
### Show image
ax.set_axis_off()
ax.imshow(image)
ax.set_title('True: %i, predicted: %i\nat %f' % (orig_label, output, certainty))
# +
from PIL import ImageTk, Image, ImageDraw
import PIL
from tkinter import *
import cv2
width = 200 # canvas width
height = 200 # canvas height
center = height//2
white = (255, 255, 255) # canvas back
def save():
# save image to hard drive
filename = "user_input.png"
global output_image
output_image.save(filename)
###### Centering begin
# Load image as grayscale and obtain bounding box coordinates
image = cv2.imread('user_input.png', 0)
height, width = image.shape
x,y,w,h = cv2.boundingRect(image)
# Create new blank image and shift ROI to new coordinates
ROI = image[y:y+h, x:x+w]
mask = np.zeros([ROI.shape[0]+10,ROI.shape[1]+10])
width, height = mask.shape
print(ROI.shape)
print(mask.shape)
x = width//2 - ROI.shape[0]//2
y = height//2 - ROI.shape[1]//2
print(x,y)
mask[y:y+h, x:x+w] = ROI
# Check if centering/masking was successful
# plt.imshow(mask, cmap='viridis')
output_image = PIL.Image.fromarray(mask) # mask has values in [0-255] as expected
# Now we need to resize, but it causes problems with default arguments as it changes the range of pixel values to be negative or positive
# compressed_output_image = output_image.resize((22,22))
# Therefore, we use the following:
compressed_output_image = output_image.resize((22,22), PIL.Image.BILINEAR) # PIL.Image.NEAREST or PIL.Image.BILINEAR also performs good
# # Enhance Saturation
# converter = PIL.ImageEnhance.Color(compressed_output_image)
# compressed_output_image = converter.enhance(2.5)
# Enhance contrast
# converter = PIL.ImageEnhance.Contrast(compressed_output_image)
# compressed_output_image = converter.enhance(3.5)
convert_tensor = torchvision.transforms.ToTensor()
tensor_image = convert_tensor(compressed_output_image)
# Another problem we face is that in the above ToTensor() command, we should have gotten a normalized tensor with pixel values in [0,1]
# But somehow it doesn't happen. Therefore, we need to normalize manually
tensor_image = tensor_image/255.
# Padding
tensor_image = torch.nn.functional.pad(tensor_image, (3,3,3,3), "constant", 0)
# Normalization shoudl be done after padding i guess
convert_tensor = torchvision.transforms.Normalize((0.1722273), (0.0521201))
tensor_image = convert_tensor(tensor_image)
plt.imshow(tensor_image.detach().cpu().numpy().reshape(28,28), cmap='viridis')
# Debugging
# print(tensor_image)
# print(np.array(compressed_output_image.getdata())) # Get data values)
# print(np.array(image.getdata()))
### Compute the predictions
print()
with torch.no_grad():
# print(tensor_image)
output0 = Network(torch.unsqueeze(tensor_image, dim=0).to(device=device))
print(output0)
certainty, output = torch.max(output0[0], 0)
certainty = certainty.clone().cpu().item()
output = output.clone().cpu().item()
certainty1, output1 = torch.topk(output0[0],3)
certainty1 = certainty1.clone().cpu()#.item()
output1 = output1.clone().cpu()#.item()
# print(certainty)
print(output)
print(chr(ord('@')+output))
print(certainty1)
print(output1)
for i in output1:
print(chr(ord('@')+i))
def paint(event):
x1, y1 = (event.x - 1), (event.y - 1)
x2, y2 = (event.x + 1), (event.y + 1)
# canvas.create_oval(x1, y1, x2, y2, fill="white",width=24)
canvas.create_rectangle(x1, y1, x2, y2, fill="white",width=12)
draw.line([x1, y1, x2, y2],fill="white",width=12)
master = Tk()
# create a tkinter canvas to draw on
canvas = Canvas(master, width=width, height=height, bg='white')
canvas.pack()
# create an empty PIL image and draw object to draw on
output_image = PIL.Image.new("L", (width, height), 0)
draw = ImageDraw.Draw(output_image)
canvas.pack(expand=YES, fill=BOTH)
canvas.bind("<B1-Motion>", paint)
# add a button to save the image
button=Button(text="save",command=save)
button.pack()
master.mainloop()
# +
# Debugging
# plt.imshow(image, cmap='viridis')
# -
# # References:
# https://stackoverflow.com/questions/59535286/improve-real-life-results-of-neural-network-trained-with-mnist-dataset
| mnist_experiments/EMNIST_letters.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Métodos de las cadenas
# ### upper(): Devuelve la cadena con todos sus caracteres a mayúscula
"<NAME>".upper()
# ### lower(): Devuelve la cadena con todos sus caracteres a minúscula
"<NAME>".lower()
# ### capitalize(): Devuelve la cadena con su primer carácter en mayúscula
"<NAME>".capitalize()
# ### title(): Devuelve la cadena con el primer carácter de cada palabra en mayúscula
"<NAME>".title()
# ### count(): Devuelve una cuenta de las veces que aparece una subcadena en la cadena
"<NAME>".count('mundo')
# ### find(): Devuelve el índice en el que aparece la subcadena (-1 si no aparece)
"<NAME>".find('mundo')
"Hola mundo".find('mundoz')
# ### rfind(): Devuelve el índice en el que aparece la subcadena, empezando por el final
"Hola mundo mundo mundo".rfind('mundo')
# ### isdigit(): Devuelve True si la cadena es todo números (False en caso contrario)
c = "100"
c.isdigit()
# ### isalnum(): Devuelve True si la cadena es todo números o carácteres alfabéticos
c2 = "ABC10034po"
c2.isalnum()
# ### isalpha(): Devuelve True si la cadena es todo carácteres alfabéticos
c2.isalpha()
"Holamundo".isalpha()
# ### islower(): Devuelve True si la cadena es todo minúsculas
"<NAME>".islower()
# ### isupper(): Devuelve True si la cadena es todo mayúsculas
"<NAME>".isupper()
# ### istitle(): Devuelve True si la primera letra de cada palabra es mayúscula
"<NAME>".istitle()
# ### isspace(): Devuelve True si la cadena es todo espacios
" - ".isspace()
# ### startswith(): Devuelve True si la cadena empieza con una subcadena
"H<NAME>".startswith("Mola")
# ### endswith(): Devuelve True si la cadena acaba con una subcadena
"<NAME>".endswith('mundo')
# ### split(): Separa la cadena en subcadenas a partir de sus espacios y devuelve una lista
"Hola mundo mundo".split()[0]
# #### Podemos indicar el carácter a partir del que se separa:
"Hola,mundo,mundo,otra,palabra".split(',')
# ### join(): Une todos los caracteres de una cadena utilizando un caracter de unión
",".join("Hola mundo")
" ".join("Hola")
# ### strip(): Borra todos los espacios por delante y detrás de una cadena y la devuelve
" <NAME> ".strip()
# #### Podemos indicar el carácter a borrar:
"-----<NAME>---".strip('-')
# ### replace(): Reemplaza una subcadena de una cadena por otra y la devuelve
"Hola mundo".replace('o','0')
# #### Podemos indicar un límite de veces a reemplazar:
"Hola mundo mundo mundo mundo mundo".replace(' mundo','',4)
| MaterialCursoPython/Fase 3 - Programacion Orientada a Objetos/Tema 10 - Metodos de las colecciones/Apuntes/Leccion 1 (Apuntes) - Cadenas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Load Data from CSVs
# +
import unicodecsv
## Longer version of code (replaced with shorter, equivalent version below)
# enrollments = []
# f = open('enrollments.csv', 'rb')
# reader = unicodecsv.DictReader(f)
# for row in reader:
# enrollments.append(row)
# f.close()
with open('enrollments.csv', 'rb') as f:
reader = unicodecsv.DictReader(f)
enrollments = list(reader)
enrollments[0]
# +
#####################################
# 1 #
#####################################
## Read in the data from daily_engagement.csv and project_submissions.csv
## and store the results in the below variables.
## Then look at the first row of each table.
daily_engagement =
project_submissions =
# +
def read_csv(filename):
with open(filename,'rb') as f :
reader =unicodecsv.DictReader(f)
return list(reader)
enrollments = read_csv("enrollments.csv")
daily_engagement= read_csv("daily_engagement.csv")
project_submissions= read_csv("project_submissions.csv")
daily_engagement[0]
project_submissions[0]
# -
# ## Fixing Data Types
# +
from datetime import datetime as dt
# Takes a date as a string, and returns a Python datetime object.
# If there is no date given, returns None
def parse_date(date):
if date == '':
return None
else:
return dt.strptime(date, '%Y-%m-%d')
# Takes a string which is either an empty string or represents an integer,
# and returns an int or None.
def parse_maybe_int(i):
if i == '':
return None
else:
return int(i)
# Clean up the data types in the enrollments table
for enrollment in enrollments:
enrollment['cancel_date'] = parse_date(enrollment['cancel_date'])
enrollment['days_to_cancel'] = parse_maybe_int(enrollment['days_to_cancel'])
enrollment['is_canceled'] = enrollment['is_canceled'] == 'True'
enrollment['is_udacity'] = enrollment['is_udacity'] == 'True'
enrollment['join_date'] = parse_date(enrollment['join_date'])
enrollments[0]
# +
# Clean up the data types in the engagement table
for engagement_record in daily_engagement:
engagement_record['lessons_completed'] = int(float(engagement_record['lessons_completed']))
engagement_record['num_courses_visited'] = int(float(engagement_record['num_courses_visited']))
engagement_record['projects_completed'] = int(float(engagement_record['projects_completed']))
engagement_record['total_minutes_visited'] = float(engagement_record['total_minutes_visited'])
engagement_record['utc_date'] = parse_date(engagement_record['utc_date'])
daily_engagement[0]
# +
# Clean up the data types in the submissions table
for submission in project_submissions:
submission['completion_date'] = parse_date(submission['completion_date'])
submission['creation_date'] = parse_date(submission['creation_date'])
project_submissions[0]
# -
# ## Investigating the Data
# +
#####################################
# 2 #
#####################################
## Find the total number of rows and the number of unique students (account keys)
## in each table.
# -
for engagement_record in daily_engagement:
engagement_record['account_key'] = engagement_record['acct']
del [engagement_record['acct']]
def get_unique_students(data):
unique_students = set()
for data_point in data:
unique_students.add(data_point['account_key'])
return unique_students
len(enrollments)
unique_enrolled_students = get_unique_students(enrollments)
len(unique_enrolled_students)
len(daily_engagement)
unique_engagement_students = get_unique_students(daily_engagement)
len(unique_engagement_students)
len(project_submissions)
unique_project_submitters = get_unique_students(project_submissions)
len(unique_project_submitters)
len(enrollments)
unique_enrolled_students =set()
for enrollment in enrollments:
unique_enrolled_students.add(enrollment['account_key'])
len(unique_enrolled_students)
len(daily_engagement)
unique_daily_engagement_students =set()
for engagement in daily_engagement:
unique_daily_engagement_students.add(engagement['acct'])
len(unique_daily_engagement_students)
len(project_submissions)
# +
unique_project_submissions_students =set()
for submissions in project_submissions:
unique_project_submissions_students.add(submissions['account_key'])
len(unique_project_submissions_students)
# -
# ## Problems in the Data
# +
#####################################
# 3 #
#####################################
## Rename the "acct" column in the daily_engagement table to "account_key".
# -
"""
for engagement_record in daily_engagement:
engagement_record['account_key'] = engagement_record['acct']
del [engagement_record['acct']]
"""""
"""
def get_unique_students(data):
unique_students = set()
for data_point in data:
unique_students.add(data_point['account_key'])
return unique_students
len(enrollments)
unique_enrolled_students = get_unique_students(enrollments)
len(unique_enrolled_students)
len(daily_engagement)
unique_engagement_students = get_unique_students(daily_engagement)
len(unique_engagement_students)
len(project_submissions)
unique_project_submitters = get_unique_students(project_submissions)
len(unique_project_submitters)
"""
# ## Missing Engagement Records
# +
#####################################
# 4 #
#####################################
## Find any one student enrollments where the student is missing from the daily engagement table.
## Output that enrollment.
# -
for enrollment in enrollments:
student = enrollment['account_key']
if student not in unique_engagement_students:
print(enrollment)
break
# ## Checking for More Problem Records
# +
#####################################
# 5 #
#####################################
## Find the number of surprising data points (enrollments missing from
## the engagement table) that remain, if any.
# +
num_problem_students = 0
for enrollment in enrollments:
student = enrollment['account_key']
if student not in unique_engagement_students \
and enrollment['join_date'] != enrollment['cancel_date']:
num_problem_students += 1
num_problem_students
# -
# ## Tracking Down the Remaining Problems
# Create a set of the account keys for all Udacity test accounts
udacity_test_accounts = set()
for enrollment in enrollments:
if enrollment['is_udacity']:
udacity_test_accounts.add(enrollment['account_key'])
len(udacity_test_accounts)
# Given some data with an account_key field, removes any records corresponding to Udacity test accounts
def remove_udacity_accounts(data):
non_udacity_data = []
for data_point in data:
if data_point['account_key'] not in udacity_test_accounts:
non_udacity_data.append(data_point)
return non_udacity_data
# +
# Remove Udacity test accounts from all three tables
non_udacity_enrollments = remove_udacity_accounts(enrollments)
non_udacity_engagement = remove_udacity_accounts(daily_engagement)
non_udacity_submissions = remove_udacity_accounts(project_submissions)
print(len(non_udacity_enrollments))
print(len(non_udacity_engagement))
print(len(non_udacity_submissions))
# -
# ## Refining the Question
# +
#####################################
# 6 #
#####################################
## Create a dictionary named paid_students containing all students who either
## haven't canceled yet or who remained enrolled for more than 7 days. The keys
## should be account keys, and the values should be the date the student enrolled.
paid_students = {}
for enrollment in non_udacity_enrollments:
if not enrollment['is_canceled'] or enrollment['days_to_cancel'] > 7 :
account_key = enrollment['account_key']
enrollment_date = enrollment['join_date']
paid_students[account_key] = enrollment_date
if account_key not in paid_students or \
enrollment_date > paid_students[account_key]:
paid_students[account_key]= enrollment_date
len(paid_students)
# -
# ## Getting Data from First Week
# Takes a student's join date and the date of a specific engagement record,
# and returns True if that engagement record happened within one week
# of the student joining.
def within_one_week(join_date, engagement_date):
time_delta = engagement_date - join_date
return time_delta.days < 7
# +
#####################################
# 7 #
#####################################
## Create a list of rows from the engagement table including only rows where
## the student is one of the paid students you just found, and the date is within
## one week of the student's join date.
paid_engagement_in_first_week =
# -
# ## Exploring Student Engagement
# +
from collections import defaultdict
# Create a dictionary of engagement grouped by student.
# The keys are account keys, and the values are lists of engagement records.
engagement_by_account = defaultdict(list)
for engagement_record in paid_engagement_in_first_week:
account_key = engagement_record['account_key']
engagement_by_account[account_key].append(engagement_record)
# -
# Create a dictionary with the total minutes each student spent in the classroom during the first week.
# The keys are account keys, and the values are numbers (total minutes)
total_minutes_by_account = {}
for account_key, engagement_for_student in engagement_by_account.items():
total_minutes = 0
for engagement_record in engagement_for_student:
total_minutes += engagement_record['total_minutes_visited']
total_minutes_by_account[account_key] = total_minutes
# +
import numpy as np
# Summarize the data about minutes spent in the classroom
total_minutes = total_minutes_by_account.values()
print 'Mean:', np.mean(total_minutes)
print 'Standard deviation:', np.std(total_minutes)
print 'Minimum:', np.min(total_minutes)
print 'Maximum:', np.max(total_minutes)
# -
# ## Debugging Data Analysis Code
# +
#####################################
# 8 #
#####################################
## Go through a similar process as before to see if there is a problem.
## Locate at least one surprising piece of data, output it, and take a look at it.
# -
# ## Lessons Completed in First Week
# +
#####################################
# 9 #
#####################################
## Adapt the code above to find the mean, standard deviation, minimum, and maximum for
## the number of lessons completed by each student during the first week. Try creating
## one or more functions to re-use the code above.
# -
# ## Number of Visits in First Week
# +
######################################
# 10 #
######################################
## Find the mean, standard deviation, minimum, and maximum for the number of
## days each student visits the classroom during the first week.
# -
# ## Splitting out Passing Students
# +
######################################
# 11 #
######################################
## Create two lists of engagement data for paid students in the first week.
## The first list should contain data for students who eventually pass the
## subway project, and the second list should contain data for students
## who do not.
subway_project_lesson_keys = ['746169184', '3176718735']
passing_engagement =
non_passing_engagement =
# -
# ## Comparing the Two Student Groups
# +
######################################
# 12 #
######################################
## Compute some metrics you're interested in and see how they differ for
## students who pass the subway project vs. students who don't. A good
## starting point would be the metrics we looked at earlier (minutes spent
## in the classroom, lessons completed, and days visited).
# -
# ## Making Histograms
# +
######################################
# 13 #
######################################
## Make histograms of the three metrics we looked at earlier for both
## students who passed the subway project and students who didn't. You
## might also want to make histograms of any other metrics you examined.
# -
# ## Improving Plots and Sharing Findings
# +
######################################
# 14 #
######################################
## Make a more polished version of at least one of your visualizations
## from earlier. Try importing the seaborn library to make the visualization
## look better, adding axis labels and a title, and changing one or more
## arguments to the hist() function.
| code/L1_Starter_Code.Answers&Given.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # 9장 비지도학습
# + [markdown] slideshow={"slide_type": "slide"}
# #### 감사의 글
#
# 자료를 공개한 저자 오렐리앙 제롱과 강의자료를 지원한 한빛아카데미에게 진심어린 감사를 전합니다.
# + [markdown] slideshow={"slide_type": "slide"}
# * 레이블이 없는 데이터 학습
# * 예제: 사진에 포함된 사람들 분류하기
# + [markdown] slideshow={"slide_type": ""}
# * 용도
# * 군집(clustering)
# * 이상치 탐지
# * 밀도 추정
# + [markdown] slideshow={"slide_type": "slide"}
# * 군집(clustering): 비슷한 샘플끼리 군집 형성하기
# * 데이터 분석
# * 고객분류
# * 추천 시스템
# * 검색 엔진
# * 이미지 분할
# * 준지도 학습
# * 차원 축소
# + [markdown] slideshow={"slide_type": "slide"}
# * 이상치 탐지: 정상테이터 학습 후 이상치 탐지.
# * 제조라인에서 결함제품 탐지
# * 시계열데이터에서 새로운 트렌드 찾기
# + [markdown] slideshow={"slide_type": "slide"}
# * 밀도 추정: 데이터셋 생성확률과정의 확률밀도함수 추정 가능
# * 이상치 분류: 밀도가 낮은 지역에 위치한 샘플
# * 데이터분석
# * 시각화
# + [markdown] slideshow={"slide_type": "slide"}
# ## 주요 내용
# + [markdown] colab_type="text" id="LYbSTqz67D9H" slideshow={"slide_type": "slide"}
# * 군집
# * K-평균
# * DBSCAN
# * 가우시안혼합
# + [markdown] colab_type="text" id="l-Y7yrhc7cM6" slideshow={"slide_type": "slide"}
# ## 군집/군집화
# + [markdown] slideshow={"slide_type": "fragment"}
# * 군집(클러스터, cluster): 유사한 샘플들의 모음(집합, 그룹)
# * 군집화(clustering): 유사한 부류의 대상들로 이루어진 군집 만들기
# + [markdown] slideshow={"slide_type": "slide"}
# ### 분류 대 군집화
# + [markdown] slideshow={"slide_type": ""}
# * 유사점: 각 샘플에 하나의 그룹 할당
# * 차이점: 군집화는 군집이 미리 레이블(타깃)로 지정되지 않고 예측기 스스로 적절한 군집을 찾아내야 함.
# + [markdown] slideshow={"slide_type": "slide"}
# #### 예제
# + [markdown] slideshow={"slide_type": "slide"}
# * 왼편: 분류
# * 오른편: 군집화
# + [markdown] slideshow={"slide_type": ""}
# <img src="images/ch09/homl09-01.png" width="600"/>
# + [markdown] slideshow={"slide_type": "slide"}
# * 가우시안혼합 모델 적용하면 매우 정확한 군집화 가능
# * 4개의 특성 모두 사용할 경우
# * 꽃잎의 너비/길이, 꽃받침의 너비/길이
# + [markdown] slideshow={"slide_type": ""}
# <img src="images/ch09/homl09-02.png" width="400"/>
# + [markdown] slideshow={"slide_type": "slide"}
# ### 군집화 활용 예제
#
# * 고객 분류
# * 데이터 분석
# * 차원축소 기법
# * 이상치탐지
# * 준지도학습
# * 검색엔진
# * 이미지 분할
# + [markdown] slideshow={"slide_type": "slide"}
# ### 군집의 정의
# + [markdown] slideshow={"slide_type": "slide"}
# * 보편적 정의 없음. 사용되는 알고리즘에 따라 다른 형식으로 군집 형성
# + [markdown] slideshow={"slide_type": "fragment"}
# * K-평균: 센트로이드(중심)라는 특정 샘플을 중심으로 모인 샘플들의 그룹
# + [markdown] slideshow={"slide_type": "fragment"}
# * DBSCAN: 밀집된 샘플들의 연속으로 이루어진 그룹
# + [markdown] slideshow={"slide_type": "fragment"}
# * 가우시안혼합 모델: 특정 가우시안 분포를 따르는 샘플들의 그룹
# + [markdown] slideshow={"slide_type": "fragment"}
# * 경우에 따라 계층적 군집의 군집 형성 가능
# + [markdown] slideshow={"slide_type": "slide"}
# ## 비지도 학습 모델 1: K-평균
# + [markdown] slideshow={"slide_type": "slide"}
# ### 예제
# + [markdown] slideshow={"slide_type": ""}
# * 샘플 덩어리 다섯 개로 이루어진 데이터셋
# + [markdown] slideshow={"slide_type": ""}
# <img src="images/ch09/homl09-03.png" width="450"/>
# + [markdown] slideshow={"slide_type": "slide"}
# ### (사이킷런) K-평균 알고리즘 적용
# -
# * 각 군집의 중심을 찾고 가장 가까운 군집에 샘플 할당
# * 군집수(`n_clusters`) 지정해야 함.
# + [markdown] slideshow={"slide_type": "fragment"}
# ---
# ```python
# from sklearn.cluster import KMeans
#
# k = 5
# kmeans = KMeans(n_clusters=k, random_state=42)
# y_pred = kmeans.fit_predict(X)
# ```
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# ### 결정 경계
# -
# * 결과: 보로노이 다이어그램
# * 평면을 특정 점까지의 거리가 가장 가까운 점의 집합으로 분할한 그림
# * 경계 부분의 일부 샘플을 제외하고 기본적으로 군집이 잘 구성됨.
# + [markdown] slideshow={"slide_type": ""}
# <img src="images/ch09/homl09-04.png" width="450"/>
# + [markdown] slideshow={"slide_type": "slide"}
# ### K-평균 알고리즘의 단점 1
# + [markdown] slideshow={"slide_type": ""}
# * 군집의 크기가 서로 많이 다르면 잘 작동하지 않음.
# * 샘플과 센트로이드까지의 거리만 고려되기 때문.
# + [markdown] slideshow={"slide_type": "slide"}
# ### 하드 군집화 대 소프트 군집화
# + [markdown] slideshow={"slide_type": "fragment"}
# * 하드 군집화: 각 샘플에 대해 가장 가까운 군집 선택
# + [markdown] slideshow={"slide_type": "fragment"}
# * 소프트 군집화: 샘플별로 각 군집 센트로이드와의 거리 측정
# + [markdown] slideshow={"slide_type": "slide"}
# ### K-평균 알고리즘
# + [markdown] slideshow={"slide_type": "slide"}
# * 먼저 $k$ 개의 센트로이드 랜덤 선택
# * 수렴할 때까지 다음 과정 반복
# * 각 샘플을 가장 가까운 센트로이드에 할당
# * 군집별로 샘플의 평균을 계산하여 새로운 센트로이드 지정
# + [markdown] slideshow={"slide_type": "slide"}
# <img src="images/ch09/homl09-05.png" width="600"/>
# + [markdown] slideshow={"slide_type": "slide"}
# ### K-평균 알고리즘의 단점 2
# + [markdown] slideshow={"slide_type": ""}
# * 초기 센트로로이드에 따라 매우 다른 군집화 발생 가능
# + [markdown] slideshow={"slide_type": ""}
# <img src="images/ch09/homl09-06.png" width="600"/>
# + [markdown] slideshow={"slide_type": "slide"}
# ### 관성(inertia, 이너셔)
# -
# * k-mean 모델 평가 방법
# * 정의: 샘플과 가장 가까운 센트로이드와의 거리의 제곱의 합
# * 각 군집이 센트로이드에 얼마나 가까이 모여있는가를 측정
# * `score()` 메서드가 측정. (음수 기준)
# + [markdown] slideshow={"slide_type": "slide"}
# ### 좋은 모델 선택법
# -
# * 다양한 초기화 과정을 실험한 후에 가장 좋은 것 선택
# * `n_init = 10`이 기본값으로 사용됨. 즉, 10번 학습 후 가장 낮은 관성을 갖는 모델 선택.
# + [markdown] slideshow={"slide_type": "slide"}
# ### K-평균++
# -
# * 센트로이드를 무작위로 초기화하는 대신 특정 확률분포를 이용하여 선택
# * 센트로이드들 사이의 거리를 크게할 가능성이 높아짐.
# * `KMeans` 모델의 기본값으로 사용됨.
# + [markdown] slideshow={"slide_type": "slide"}
# ### elkan 알고리즘
# -
# * `algorithm=elkan`: 학습 속도 향상됨.
# + [markdown] slideshow={"slide_type": "fragment"}
# * 단, 밀집데이터(dense data)만 지원하며, 희소데이터는 지원하지 않음.
# + [markdown] slideshow={"slide_type": "fragment"}
# * 밀집데이터셋에 대한 기본값임.
# + [markdown] slideshow={"slide_type": "fragment"}
# * `algorithm=full`: 희소데이터에 대한 기본값.
# + [markdown] slideshow={"slide_type": "slide"}
# ### 미니배치 K-평균
# -
# * 미니배치를 지원하는 K-평균 알고리즘: `MiniBatchMeans`
# + [markdown] slideshow={"slide_type": "fragment"}
# * 사용법은 동일
# + [markdown] slideshow={"slide_type": ""}
# ```python
# from sklearn.cluster import MiniBatchKMeans
#
# minibatch_kmeans = MiniBatchKMeans(n_clusters=5, random_state=42)
# minibatch_kmeans.fit(X)
# ```
# + [markdown] slideshow={"slide_type": "slide"}
# #### `memmap` 활용
# -
# * 대용량 훈련 세트 활용하고자 할 경우
# * 8장 PCA에서 사용했던 기법과 동일
# + [markdown] slideshow={"slide_type": "slide"}
# #### memmap 활용이 불가능할 정도로 큰 데이터셋을 다뤄야 하는 경우
# -
# * 미니배치로 쪼개어 학습
# * `MiniBatchKMeans`의 `partial_fit()` 메서드 활용
# + [markdown] slideshow={"slide_type": "slide"}
# #### 미니배치 K-평균의 특징
# -
# * K-평균보다 훨씬 빠름.
# * 하지만 성능은 상대적으로 좀 떨어짐.
# * 군집수가 증가해도 마찬가지임.
# + [markdown] slideshow={"slide_type": "-"}
# <img src="images/ch09/homl09-07.png" width="600"/>
# + [markdown] slideshow={"slide_type": "slide"}
# ### 최적의 군집수 찾기
# -
# * 최적의 군집수를 사용하지 않으면 적절하지 못한 모델을 학습할 수 있음.
# + [markdown] slideshow={"slide_type": "fragment"}
# <img src="images/ch09/homl09-08.png" width="600"/>
# + [markdown] slideshow={"slide_type": "slide"}
# #### 관성과 군집수
# + [markdown] slideshow={"slide_type": ""}
# * 군집수 k가 증가할 수록 관성(inertia) 줄어듬.
# * 따라서 관성만으로 모델을 평가할 없음.
# + [markdown] slideshow={"slide_type": "slide"}
# * 관성이 더 이상 획기적으로 줄어들지 않는 지점의 군집수 선택 가능
# + [markdown] slideshow={"slide_type": ""}
# <img src="images/ch09/homl09-09.png" width="600"/>
# + [markdown] slideshow={"slide_type": "slide"}
# * 위 그래프에 의해 `k=4` 선택 가능.
# * 하지만 아래 그림에서 보듯이 좋은 성능이라 말하기 어려움.
# + [markdown] slideshow={"slide_type": ""}
# <img src="images/ch09/homl09-10.png" width="400"/>
# + [markdown] slideshow={"slide_type": "slide"}
# #### 실루엣 점수와 군집수
# -
# * 샘플별 실루엣 계수의 평균값
# + [markdown] slideshow={"slide_type": "fragment"}
# * 실루엣 계수: -1과 1사이의 값
# * 1에 가까운 값: 적절한 군집에 포함됨.
# * 0에 가까운 값: 군집 경계에 위치
# * -1에 가까운 값: 잘못된 군집에 포함됨
# + [markdown] slideshow={"slide_type": "slide"}
# * 아래 그림에 의하면 `k=5`도 좋은 선택이 될 수 있음.
# + [markdown] slideshow={"slide_type": ""}
# <img src="images/ch09/homl09-11.png" width="500"/>
# + [markdown] slideshow={"slide_type": "slide"}
# #### 실루엣 다이어그램과 군집수
# + [markdown] slideshow={"slide_type": ""}
# <img src="images/ch09/homl09-12.png" width="500"/>
# + [markdown] slideshow={"slide_type": "slide"}
# * 실루엣 다이어그램
# * 군집별 실루엣 계수 모음. 칼 모양.
# * 칼 두께: 군집에 포함된 샘플 수
# * 칼 길이: 군집에 포함된 각 샘플의 실루엣 계수
# + [markdown] slideshow={"slide_type": "fragment"}
# * 빨간 파선: 군집별 실루엣 점수
# * 대부분의 칼이 빨간 파선보다 길어야 함.
# * 칼의 두께가 서로 비슷해야, 즉, 군집별 크기가 비슷해야 좋은 모델임.
# + [markdown] slideshow={"slide_type": "fragment"}
# * 따라서 `k=5` 가 보다 좋은 모델임.
# + [markdown] slideshow={"slide_type": "slide"}
# ### K-평균의 한계
# + [markdown] slideshow={"slide_type": "slide"}
# * 최적의 모델을 구하기 위해 여러 번 학습해야 함.
# * 군집수를 미리 지정해야 함.
# * 군집의 크기나 밀집도가 다르거나, 원형이 아닐 경우 잘 작동하지 않음.
# + [markdown] slideshow={"slide_type": ""}
# <img src="images/ch09/homl09-13.png" width="600"/>
# + [markdown] slideshow={"slide_type": "slide"}
# ### 군집화 활용: 이미지 분할
# + [markdown] slideshow={"slide_type": "slide"}
# #### 이미지 분할
# -
# * 이미지를 여러 영역(segment)으로 분할하기
# + [markdown] slideshow={"slide_type": "fragment"}
# * 동일한 종류의 물체는 동일한 영역에 할당됨.
# * 자율주행: 보행자들을 모두 하나의 영역, 또는 각각의 영역으로 할당 가능
# + [markdown] slideshow={"slide_type": "fragment"}
# * 합성곱 신경망이 가장 좋은 성능 발휘
# + [markdown] slideshow={"slide_type": "slide"}
# * 여기서는 K-평균을 이용하여 색상분할 실행
# * 인공위성 사진 분석: 전체 산림 면적 측정
# * 군집수가 중요함.
# + [markdown] slideshow={"slide_type": "slide"}
# ---
# ```python
# segmented_imgs = []
# n_colors = (10, 8, 6, 4, 2)
# for n_clusters in n_colors:
# kmeans = KMeans(n_clusters=n_clusters, random_state=42).fit(X)
# segmented_img = kmeans.cluster_centers_[kmeans.labels_]
# segmented_imgs.append(segmented_img.reshape(image.shape))
# ```
# ---
# + [markdown] slideshow={"slide_type": "fragment"}
# <img src="images/ch09/homl09-14.png" width="600"/>
# + [markdown] slideshow={"slide_type": "slide"}
# ### 군집화 활용: 전처리
# + [markdown] slideshow={"slide_type": "slide"}
# #### 미니 MNIST 데이터셋 전처리
# -
# * MNIST와 비슷한 숫자 데이터셋
# + [markdown] slideshow={"slide_type": "fragment"}
# * 8x8 크기의 흑백 사진 1,797개.
# + [markdown] slideshow={"slide_type": "fragment"}
# * 전처리 없이 로지스틱회귀 학습시키면 96.89% 정확도 보임.
# + [markdown] slideshow={"slide_type": "slide"}
# #### K-평균 활용 전처리 후 로지스틱회귀 학습
# -
# * 구역수: 50
# + [markdown] slideshow={"slide_type": "fragment"}
# ---
# ```python
# pipeline = Pipeline([
# ("kmeans", KMeans(n_clusters=50, random_state=42)),
# ("log_reg", LogisticRegression(multi_class="ovr", solver="lbfgs", max_iter=5000, random_state=42)),
# ])
# pipeline.fit(X_train, y_train)
# ```
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# * 정확도: 97.78%로 증가
# + [markdown] slideshow={"slide_type": "fragment"}
# * 전처리 단계로 K-평균을 활용하기에 그리드 탐색 등을 이용하여 최적의 군집수 확인 가능.
# * 최적 군집수: 99
# * 모델 정확도: 98.22%
# -
# ---
# ```python
# param_grid = dict(kmeans__n_clusters=range(2, 100))
# grid_clf = GridSearchCV(pipeline, param_grid, cv=3, verbose=2)
# grid_clf.fit(X_train, y_train)
# ```
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# ### 군집화 활용: 준지도 학습
# + [markdown] slideshow={"slide_type": ""}
# * 레이블이 있는 데이터가 적고, 레이블이 없는 데이터가 많을 때 활용
# + [markdown] slideshow={"slide_type": "slide"}
# ### 예제: 미니 MNist (계속)
# + [markdown] slideshow={"slide_type": "fragment"}
# * 50개 샘플을 대상으로 학습한 모델의 성능: 83.33% 정도
# + [markdown] slideshow={"slide_type": "slide"}
# * 하지만 50개의 군집으로 나눈 후 군집별로 __대표 이미지__ 50개 선정.
# * 군집 센트로이드에 가장 가까운 샘플
# + [markdown] slideshow={"slide_type": "fragment"}
# <img src="images/ch09/homl09-15.png" width="600"/>
# + [markdown] slideshow={"slide_type": "slide"}
# * 50개 사진을 보고 수동으로 레이블 작성
# + [markdown] slideshow={"slide_type": "fragment"}
# * 위 50개 샘플을 이용하여 학습된 모델 성능: 92.22%로 향상
# + [markdown] slideshow={"slide_type": "slide"}
# #### 레이블 전파
# -
# * 위 50개의 그림과 동일한 군집에 속한 샘플에 동일한 레이블 전파하기.
# + [markdown] slideshow={"slide_type": "fragment"}
# * 군집에 속한 전체 샘플 보다 센트로이드에 가까운 20% 정도에게만 레이블 전파 후 학습
# * 센트로이드에 가깝기 떼문에 레이블의 정확도가 매우 높음.
# + [markdown] slideshow={"slide_type": "fragment"}
# * 정확도: 94%까지 향상.
# + [markdown] slideshow={"slide_type": "fragment"}
# * 참고
# * 전체 데이터셋으로 훈련된 로지스틱 회귀 모델 성능: 정확도 96.9%
# + [markdown] slideshow={"slide_type": "slide"}
# #### 준지도학습과 능동학습
# -
# * 분류기 모델이 가장 불확실하기 예측하는 샘플에 레이블 추가하기
# * 가능하면 서로 다른 군집에서 선택.
# * 새 모델 학습
# * 위 과정을 성능향상이 약해질 때까지 반복.
# + [markdown] slideshow={"slide_type": "slide"}
# ## 비지도학습 모델 2: DBSCAN
# + [markdown] slideshow={"slide_type": "slide"}
# * 연속적인 밀집 지역을 하나의 군집으로 설정.
# + [markdown] slideshow={"slide_type": "slide"}
# ### 사이킷런의 DBSCAN 모델
# -
# * 두 개의 하이퍼파라미터 사용
# * `eps`: $\varepsilon$-이웃 범위
# * 주어진 기준값 $\varepsilon$ 반경 내에 위치한 샘플
# * `min_samples`: $\varepsilon$ 반경 내에 위치하는 이웃의 수
# + [markdown] slideshow={"slide_type": "slide"}
# #### 핵심샘플과 군집
# -
# * 핵심샘플: $\varepsilon$ 반경 내에 자신을 포함해서 `min-samples`개의 이웃을 갖는 샘플
# + [markdown] slideshow={"slide_type": "fragment"}
# * 군집: 핵심샘플로 이루어진 이웃들로 구성된 그룹
# + [markdown] slideshow={"slide_type": "slide"}
# #### 이상치
# -
# * 핵심샘플이 아니면서 동시에 햄심샘플의 이웃도 아닌 샘플.
# + [markdown] slideshow={"slide_type": "slide"}
# #### 예제
# -
# * 반달모양 데이터 활용
# + [markdown] slideshow={"slide_type": "fragment"}
# ---
# ```python
# from sklearn.cluster import DBSCAN
#
# dbscan = DBSCAN(eps=0.05, min_samples=5)
# dbscan.fit(X)
# ```
# ---
# + [markdown] slideshow={"slide_type": "fragment"}
# <img src="images/ch09/homl09-16.png" width="600"/>
# + [markdown] slideshow={"slide_type": "slide"}
# ### DBSCAN과 예측
# -
# * `predict()` 메서드 지원하지 않음.
# + [markdown] slideshow={"slide_type": "fragment"}
# * 이유: `KNeighborsClassifier` 등 보다 좋은 성능의 분류 알고리즘 활용 가능.
# + [markdown] slideshow={"slide_type": "fragment"}
# * 아래 코드: 핵심샘플 대상 훈련.
# + [markdown] slideshow={"slide_type": ""}
# ---
# ```python
# from sklearn.neighbors import KNeighborsClassifier
#
# knn = KNeighborsClassifier(n_neighbors=50)
# knn.fit(dbscan.components_, dbscan.labels_[dbscan.core_sample_indices_])
# ```
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# * 이후 새로운 샘플에 대한 예측 가능
# * 아래 그림은 새로운 4개의 샘플에 대한 예측을 보여줌.
# + [markdown] slideshow={"slide_type": "fragment"}
# <img src="images/ch09/homl09-17.png" width="450"/>
# + [markdown] slideshow={"slide_type": "slide"}
# #### 이상치 판단
# -
# * 위 예제에서, 두 군집으로부터 일정거리 이상 떨어진 샘플을 이상치로 간주 가능.
# + [markdown] slideshow={"slide_type": "fragment"}
# * 예를 들어, 양편 끝쪽에 위치한 두 개의 샘플이 이상치로 간주될 수 있음.
# + [markdown] slideshow={"slide_type": ""}
# <img src="images/ch09/homl09-17a.png" width="450"/>
# + [markdown] slideshow={"slide_type": "slide"}
# ### DBSCAN의 장단점
# -
# * 매우 간단하면서 매우 강력한 알고리즘.
# * 하이퍼파라미터: 단 2개
# + [markdown] slideshow={"slide_type": "fragment"}
# * 군집의 모양과 개수에 상관없음.
# + [markdown] slideshow={"slide_type": "fragment"}
# * 이상치에 안정적임.
# + [markdown] slideshow={"slide_type": "fragment"}
# * 군깁 간의 밀집도가 크게 다르면 모든 군집 파악 불가능.
# + [markdown] slideshow={"slide_type": "slide"}
# #### 계산복잡도
# -
# * 시간복잡도: 약 $O(m\, \log m)$. 단, $m$은 샘플 수
# * 공간복잡도: 사이킷런의 DBSCAN 모델은 $O(m^2)$의 메모리 요구.
# * `eps`가 커질 경우.
# + [markdown] slideshow={"slide_type": "slide"}
# ### 기타 군집 알고리즘
# -
# * 응집 군집(병합 군집, agglomerative clustering)
# * BIRCH
# * 평균-이동
# * 유사도 전파
# * 스펙트럼 군집
# + [markdown] slideshow={"slide_type": "slide"}
# ## 비지도학습 모델 3: 가우시안혼합 모델
# + [markdown] slideshow={"slide_type": "slide"}
# * 데이터셋이 여러 개의 혼합된 가우시안 분포를 따르는 샘플들로 구성되었다고 가정.
# + [markdown] slideshow={"slide_type": ""}
# * 가우시안 분포 = 정규분포
# + [markdown] slideshow={"slide_type": "slide"}
# #### 정규분포 소개
# -
# * 종 모양의 확률밀도함수를 갖는 확률분포
# + [markdown] slideshow={"slide_type": ""}
# <img src="images/ch09/homl09-18.png" width="400"/>
# + [markdown] slideshow={"slide_type": "slide"}
# #### 군집
# -
# * 하나의 가우시안 분포에서 생생된 모든 샘플들의 그룹
# * 일반적으로 타원형 모양.
# + [markdown] slideshow={"slide_type": "slide"}
# ### 예제
# -
# * 아래 그림에서처럼 일반적으로 모양, 크기, 밀집도, 방향이 다름.
# * 따라서 각 샘플이 어떤 정규분포를 따르는지를 파악하는 게 핵심.
# + [markdown] slideshow={"slide_type": ""}
# <img src="images/ch09/homl09-13.png" width="600"/>
# + [markdown] slideshow={"slide_type": "slide"}
# ### GMM 활용
# + [markdown] slideshow={"slide_type": "slide"}
# * 위 데이터셋에 `GaussianMixture` 모델 적용
# + [markdown] slideshow={"slide_type": "fragment"}
# * `n_components`: 군집수 지정
# + [markdown] slideshow={"slide_type": "fragment"}
# * `n_init`: 모델 학습 반복 횟수.
# * 파라미터(평균값, 공분산 등)를 무작위로 추정한 후 수렴할 때까지 학습시킴.
# + [markdown] slideshow={"slide_type": "fragment"}
# ---
# ```python
# from sklearn.mixture import GaussianMixture
#
# gm = GaussianMixture(n_components=3, n_init=10, random_state=42)
# gm.fit(X)
# ```
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# * 아래 그림은 학습된 모델을 보여줌.
# * 군집 평균, 결정 경계, 밀도 등고선
# + [markdown] slideshow={"slide_type": ""}
# <img src="images/ch09/homl09-19.png" width="500"/>
# + [markdown] slideshow={"slide_type": "slide"}
# ### GMM 모델 규제
# -
# * 특성수가 크거나, 군집수가 많거나, 샘플이 적은 경우 최적 모델 학습 어려움.
# * 공분산(covariance)에 규제를 가해서 학습을 도와줄 수 있음.
# * `covariance_type` 설정.
# + [markdown] slideshow={"slide_type": "slide"}
# #### covariance_type 옵션값
# -
# * full
# * 아무런 제한 없음.
# * 기본값임.
# + [markdown] slideshow={"slide_type": "fragment"}
# * spherical
# * 군집이 원형이라 가정.
# * 지름(분산)은 다를 수 있음.
# + [markdown] slideshow={"slide_type": "slide"}
# * diag
# * 어떤 타원형도 가능.
# * 단. 타원의 축이 좌표축과 평행하다고 가정.
# + [markdown] slideshow={"slide_type": "fragment"}
# * tied
# * 모든 군집의 동일 모양, 동일 크기, 동일 방향을 갖는다고 가정.
# + [markdown] slideshow={"slide_type": "slide"}
# <img src="images/ch09/homl09-20.png" width="500"/>
# + [markdown] slideshow={"slide_type": "slide"}
# ### 가우시안혼합 모델 활용: 이상치 탐지
# -
# * 밀도가 임곗값보다 낮은 지역에 있는 샘플을 이상치로 간주 가능.
# + [markdown] slideshow={"slide_type": "slide"}
# <img src="images/ch09/homl09-21.png" width="500"/>
# + [markdown] slideshow={"slide_type": "slide"}
# ### 가우션 혼합모델 군집수 지정
# -
# * K-평균에서 사용했던 관성 또는 실루엣 점수 사용 불가.
# * 군집이 타원형일 때 값이 일정하지 않기 때문.
# + [markdown] slideshow={"slide_type": "fragment"}
# * 대신에 __이론적 정보 기준__ 을 최소화 하는 모델 선택 가능.
# + [markdown] slideshow={"slide_type": "slide"}
# #### 이론적 정보 기준
# -
# * BIC: Bayesian information criterion
#
# $$ \log(m)\, p - 2 \log (\hat L)$$
# + [markdown] slideshow={"slide_type": "fragment"}
# * AIC: Akaike information criterion
#
# $$ 2\, p - 2 \log (\hat L)$$
# + [markdown] slideshow={"slide_type": "slide"}
# * $m$: 샘플 수
# * $p$: 모델이 학습해야 할 파라미터 수
# * $\hat L$: 모델의 가능도 함수의 최댓값
# -
# * 학습해야 할 파라미터가 많을 수록 벌칙이 가해짐.
# * 데이터에 잘 학습하는 모델일 수록 보상을 더해줌.
# + [markdown] slideshow={"slide_type": "slide"}
# #### 군집수와 정보조건
# -
# * 아래 그림은 군집수 $k$와 AIC, BIC의 관계를 보여줌.
# * $k=3$이 최적으로 보임.
# + [markdown] slideshow={"slide_type": ""}
# <img src="images/ch09/homl09-22.png" width="500"/>
# + [markdown] slideshow={"slide_type": "slide"}
# ### 베이즈 가우시안혼합 모델
# -
# * 베이즈 확률통계론 활용
# + [markdown] slideshow={"slide_type": "slide"}
# #### BayesianGaussianMixture 모델
# -
# * 최적의 군집수를 자동으로 찾아줌.
# * 단, 최적의 군집수보다 큰 수를 `n_components`에 전달해야 함.
# * 즉, 군집에 대한 최소한의 정보를 알고 있다고 가정.
# * 자동으로 불필요한 군집 제거
# + [markdown] slideshow={"slide_type": "slide"}
# ---
# ```python
# from sklearn.mixture import BayesianGaussianMixture
#
# bgm = BayesianGaussianMixture(n_components=10, n_init=10, random_state=42)
# bgm.fit(X)
# ```
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# * 결과는 군집수 3개를 사용한 이전 결과와 거의 동일.
# * 군집수 확인 가능
# -
# ```python
# >>> np.round(bgm.weights_, 2)
# array([0.4 , 0.21, 0.4 , 0. , 0. , 0. , 0. , 0. , 0. , 0. ])
# ```
# + [markdown] slideshow={"slide_type": "slide"}
# #### 사전 믿음
# -
# * 군집수가 어느 정도일까를 나타내는 지수
# * `weight_concentration_prior` 하이퍼파라미터
# * `n_components`에 설정된 군집수에 대한 규제로 사용됨.
# * 작은 값이면 특정 군집의 가중치를 0에 가깝게 만들어 군집수를 줄이도록 함.
# * 즉, 큰 값일 수록 `n_components`에 설정된 군집수가 유지되도록 함.
# + [markdown] slideshow={"slide_type": "fragment"}
# <img src="images/ch09/homl09-24.png" width="500"/>
# + [markdown] slideshow={"slide_type": "slide"}
# ### 가우시안혼합 모델의 장단점
# -
# * 타원형 군집에 잘 작동.
# + [markdown] slideshow={"slide_type": "slide"}
# * 하지만 다른 모양을 가진 데이터셋에서는 성능 좋지 않음.
# + [markdown] slideshow={"slide_type": "fragment"}
# * 예제: 달모양 데이터에 적용하는 경우
# * 억지로 타원을 찾으려 시도함.
# + [markdown] slideshow={"slide_type": ""}
# <img src="images/ch09/homl09-23.png" width="500"/>
# + [markdown] slideshow={"slide_type": "slide"}
# ### 이상치 탐지와 특이치 탐지를 위한 다른 알고리즘
# -
# * PCA
# * Fast-MCD
# * 아이솔레이션 포레스트
# * LOF
# * one-class SVM
# + [markdown] slideshow={"slide_type": "slide"}
# __감사의 글__: 슬라이드에 사용할 이미지를 제공한 한빛아카데미에 감사드립니다.
| slides/handson-ml2-09.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/BaiganKing/lambdata_baiganking/blob/master/package_test.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="pddp1p3_ZAUB" colab_type="code" outputId="bdbf940f-281e-47c3-9970-edbfa698bb98" colab={"base_uri": "https://localhost:8080/", "height": 136}
# !pip install -i https://test.pypi.org/simple/ lambdata-baiganking==0.1.4
# + id="obFAiTjEZE8R" colab_type="code" colab={}
import lambdata_baiganKing as lb
from lambdata_baiganKing import df_utils
# + id="NMEsRY1PZMkw" colab_type="code" outputId="47980442-d550-403f-ed15-553f7f1131bb" colab={"base_uri": "https://localhost:8080/", "height": 221}
dir(df_utils)
# + id="3mxOJ-72c-xm" colab_type="code" colab={}
| package_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: percent
# format_version: '1.3'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:.conda-develop] *
# language: python
# name: conda-env-.conda-develop-py
# ---
# %% [markdown]
# ## Imports
# %%
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
import logging
import helpers.dbg as dbg
import helpers.env as henv
import helpers.printing as hprint
import im.kibot as vakibot
import pandas as pd
import os
import core.explore as exp
import numpy as np
import im.ib.data.extract.gateway.utils as ibutils
import datetime
import matplotlib.pyplot as plt
# %%
dbg.init_logger(verbosity=logging.INFO)
_LOG = logging.getLogger(__name__)
_LOG.info("%s", henv.get_system_signature()[0])
hprint.config_notebook()
# %%
def get_min_max_from_index(df):
min_dt = min(df.index)
max_dt = max(df.index)
if True:
min_dt = str(pd.Timestamp(min_dt).date())
max_dt = str(pd.Timestamp(max_dt).date())
print("min=", min_dt)
print("max=", max_dt)
return min_dt, max_dt
def print_df(df, n=3):
print("shape=", df.shape)
display(df.head(n))
display(df.tail(n))
# %% [markdown]
# # Kibot
# %%
df_kibot = vakibot.KibotS3DataLoader().read_data(
"Kibot",
"ES",
vakibot.AssetClass.Futures,
vakibot.Frequency.Minutely,
vakibot.ContractType.Continuous,
)
df_kibot.head()
df_kibot_orig = df_kibot.copy()
# %%
df_kibot = df_kibot_orig.copy()
df_kibot.rename({"vol": "volume"}, axis=1, inplace=True)
#df_kibot.index = pd.to_datetime(df_kibot.index, utc=True).tz_convert(tz='America/New_York')
df_kibot.index = pd.to_datetime(df_kibot.index).tz_localize(tz='America/New_York')
print_df(df_kibot, n=2)
# %%
fig, (ax1, ax2, ax3) = plt.subplots(nrows=3, ncols=1, sharex=True, figsize=(20, 10))
df_tmp = df_kibot.resample("1T").mean()
a = pd.Timestamp("2019-05-27 00:00:00")
#a = pd.Timestamp("2019-05-29 00:00:00")
#a = pd.Timestamp("2019-06-02 00:00:00")
print(a, a.day_name())
#b = a + pd.DateOffset(14)
#b = a + pd.DateOffset(7)
b = a + pd.DateOffset(3)
#b = a + pd.DateOffset(1)
print(b, b.day_name())
df_tmp = df_tmp[a:b][["close", "volume"]]
print("samples=%d [%s, %s]" % (df_tmp.shape[0], df_tmp.index[0], df_tmp.index[-1]))
dates = sorted(list(set(df_tmp.index.date)))#.unique()
for date in dates:
print(date, pd.Timestamp(date).day_name())
df_tmp = df_tmp.resample("1T").mean()
df_tmp["close"].plot(ax=ax1)
df_tmp["volume"].plot(ax=ax2)
print("samples=%d [%s, %s]" % (df_tmp.shape[0], df_tmp.index[0], df_tmp.index[-1]))
#
#df_tmp2 = df_tmp.resample("1T").mean()
#print("samples=%d [%s, %s]" % (df_tmp2.shape[0], df_tmp2.index[0], df_tmp2.index[-1]))
#support = ~pd.isnull(df_tmp["close"])
support = ~pd.isnull(df_tmp["volume"])
print(support[~support].head())
print("no support=", (~support[~support]).sum())
print("with support=", support[support].sum())
pd.DataFrame(1.0 * support).plot(ax=ax3)
for date in dates:
for h, m in ((9, 0), (16, 30)):
date_tmp = datetime.datetime.combine(date, datetime.time(h, m))
date_tmp = pd.Timestamp(date_tmp).tz_localize(tz="America/New_York")
plt.axvline(date_tmp, color="red", linestyle="--", linewidth=3)
# %% [markdown]
# # IB
# %%
import ib_insync
ib = ibutils.ib_connect(1)
# %%
contract = ib_insync.ContFuture("ES", "GLOBEX", "USD")
whatToShow = 'TRADES'
barSizeSetting = '1 min'
#barSizeSetting = '1 hour'
useRTH = False
start_ts = pd.Timestamp("2019-05-28 15:00").tz_localize(tz="America/New_York")
end_ts = pd.Timestamp("2019-05-29 15:00").tz_localize(tz="America/New_York")
#file_name = "ES.csv"
#if os.path.exists(file_name):
#df_ib = ibutils.get_data(ib, contract, start_ts, end_ts, barSizeSetting, whatToShow, useRTH)
#df_ib.to_csv("ES.csv")
durationStr = "1 D"
df_ib = ibutils.req_historical_data(ib, contract, end_ts, durationStr, barSizeSetting, whatToShow, useRTH)
# %%
display(df_ib.head(2))
print(df_ib.index[0], df_ib.index[-1])
display(df_kibot.head(2))
print(df_kibot.index[0], df_kibot.index[-1])
# %% [markdown]
# # Compare
# %%
target_col = "close"
#target_col = "open"
#target_col = "high"
#target_col = "volume"
# %%
if True:
print_df(df_ib, n=1)
print_df(df_kibot, n=1)
# %%
#min_dt = "2013-10-06"
#max_dt = "2013-10-09"
min_dt = start_ts
max_dt = end_ts
#
df_ib_tmp = df_ib.loc[min_dt:max_dt]
df_ib_tmp.columns = ["%s_ib" % c for c in df_ib_tmp.columns]
df_ib_tmp.head()
#
df_kibot_tmp = df_kibot.loc[min_dt:max_dt]
df_kibot_tmp.columns = ["%s_kibot" % c for c in df_kibot_tmp.columns]
df_kibot_tmp.head()
#df = pd.concat([df_ib_tmp, df_kibot_tmp], axis=1, join="outer")
df = pd.concat([df_ib_tmp, df_kibot_tmp], axis=1, join="inner")
display(df.head(1))
# Shift.
df["%s_ib" % target_col] = df["%s_ib" % target_col].shift(0)
# Filter columns.
display(df[cols].head(10))
cols = ["%s_%s" % (target_col, src) for src in "ib kibot".split()]
df[cols].plot()
# %%
df.iloc[:100][cols].plot()
# %%
ds1 = "ib"
ds2 = "kibot"
diff = df[target_col + "_" + ds1] - df[target_col + "_" + ds2]
diff.plot()
exp.drop_na(pd.DataFrame(diff), drop_infs=True).hist(bins=101)
# %%
intercept = False
exp.ols_regress(df,
target_col + "_" + ds1,
target_col + "_" + ds2,
intercept,
jointplot_=True,
max_nrows=None)
| im/ib/data/extract/gateway/notebooks/Task111_Compare_data_IB_vs_Kibot_and_understand_timing_semantic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:tiedecay_env]
# language: python
# name: conda-env-tiedecay_env-py
# ---
# # NHS Tie-Decay
# This notebook illustrates how to use the tie-decay code to load a temporal network dataset and analyze it.
import networkx as nx
import numpy as np
import operator
import pandas as pd
import pickle
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
from tiedecay.construct import TieDecayNetwork
from tiedecay.dataset import Dataset
# +
data_path = '../saved_data/NHSadjList.pkl'
users_path = '../saved_data/NHSUsersDict.pkl'
def open_file(file_path):
with open(file_path, 'rb') as f:
loaded = pickle.load(f)
return loaded
raw_data, user_mapping = [open_file(data_path), open_file(users_path)]
# +
dataset = Dataset(raw_data, user_mapping)
# half-life of one day
alpha = np.log(2)/24/3600
tdn = TieDecayNetwork(dataset, alpha=alpha)
# -
print(f"There are {dataset.num_interactions} interactions between {dataset.num_active_nodes} out of {dataset.num_nodes} total users.")
# ## Compute tie-decay graph at a given time
print(f'First time: {dataset.t_first}')
print(f'Last time: {dataset.t_last}')
t_select = '2012-06-01 00:00:00'
# %%time
B_t = tdn.compute_from_dataset(t_select)
# top 10 nodes by Tie-Decay PageRank at `t_select`
top_10_users = sorted(nx.pagerank(B_t).items(), key=operator.itemgetter(1), reverse=True)[:10]
top_10_users = [(dataset.node_mapping[x], y) for x,y in top_10_users]
top_10_users
# ## Compute tie-decay centrality trajectories
# Performant way of calculating centrality values sampled at equal intervals.
centrality_df = tdn.compute_centrality_trajectories_from_dataset(1000, 'pagerank')
centrality_df.head()
plot_df = centrality_df.sort_values(by=centrality_df.columns[-1], ascending=False)
sns.lineplot(x=plot_df.columns, y=plot_df.iloc[0,:])
plt.ylabel(dataset.node_mapping[plot_df.iloc[0].name])
| notebooks/nhs_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# # 9.13 dtypes
# %matplotlib inline
import pandas as pd
import numpy as np
dft = pd.DataFrame(dict(A = np.random.rand(3),
B = 1,
C = 'foo',
D = pd.Timestamp('20010102'),
E = pd.Series([1.0] * 3).astype('float32'),
F = False,
G = pd.Series([1] * 3,dtype='int8')))
dft
dft.dtypes
dft['A'].dtype
| CHAPTER 09 ESSENTIAL BASIC FUNCTIONALITY.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.10.4 64-bit
# language: python
# name: python3
# ---
import sys
import folium
import queue
import numpy as np
import osmnx as ox
import matplotlib.pyplot as plt
from geopy.geocoders import Nominatim
from geopy.distance import geodesic
ox.config(log_console=False, use_cache=False)
# Get the ROADS and NODES of Back bay, Boston
# NODES(white dots) and EDGES(grey lines) are stored in Digraph
place_name = "Back Bay, Massachusetts, USA"
mode = 'drive'
optimizer = 'length'
graph = ox.graph_from_place(place_name,network_type=mode)
fig, ax = ox.plot_graph(graph)
plt.tight_layout()
# Extract NODES and EDGES
nodes, edges = ox.graph_to_gdfs(graph, nodes=True, edges=True)
nodes
# +
# Feed nodes and lengths to algorithms, algo returns bunch of nodes, iterate through the nodes 2 by 2
NodesAndLength = list(graph.edges(data='length'))
EdgeAndLength = {}
for i in NodesAndLength:
EdgeAndLength[(i[0],i[1])] = i[2]
# +
# === Takes USER INPUTS and Find the nearest nodes on current map ===
print("Thank you for using our Navigator (Back Bay, Boston, USA)!\n\
Please enter your starting location and destination location (in Back Bay, Boston)\n\
You can either enter a place's name or the place's coordinate in this format: latitude, longitude \n\
*Note that only places that could be found on OpenStreetMap can be used*\n\
Recommendations: first church in boston --> Atlantic fish back bay\n\
boston architectural college --> gibson house back bay")
# startPoint = sys.argv[1]
# destPoint = sys.argv[2]
# startPoint = input("Your starting location: ")
# destPoint = input("Your destination: ")
startPoint = "first church in boston"
destPoint = "Atlantic fish back bay"
# ===================================================================
if startPoint[0].isdigit():
# Enter by HAND
(start_lat,start_long) = tuple(float(x) for x in startPoint.split(",")) # (latitude, longitude)
(end_lat,end_long) = tuple(float(x) for x in destPoint.split(",")) # (latitude, longitude)
orig_node = ox.distance.nearest_nodes(graph, start_long, start_lat)
dest_node = ox.distance.nearest_nodes(graph, end_long, end_lat)
else:
# Enter by GEOCODE
try:
locator = Nominatim(user_agent = "myapp")
startPoint = locator.geocode(startPoint)
print(startPoint.latitude, startPoint.longitude)
except:
raise ValueError("Cannot find the place entered for departure.. \nPlease enter a place that exists in OpenStreetMap\n")
try:
destPoint = locator.geocode(destPoint)
print(destPoint.latitude, destPoint.longitude)
except:
raise ValueError("Cannot find the place entered for destination.. \nPlease enter a place that exists in OpenStreetMap\n")
(start_long, start_lat) = (startPoint.longitude, startPoint.latitude)
(end_long, end_lat) = (destPoint.longitude, destPoint.latitude)
orig_node = ox.distance.nearest_nodes(graph, startPoint.longitude, startPoint.latitude)
dest_node = ox.distance.nearest_nodes(graph, destPoint.longitude, destPoint.latitude)
print(orig_node)
print(dest_node)
# -
# Starting computing shortest paths: Dijkstra, Bellman-Ford, and A* algorithm.
class Graph:
def __init__(self, num_of_vertices, node_list):
self.v = num_of_vertices
self.visited = []
self.graph = []
self.node_list = node_list # list of all the nodes
def add_edge(self, u, v, weight):
self.graph.append([u, v, weight])
def h(self, lat1, lon1, lat2, lon2):
'''
TO DO:
This is to calculate the heuristic distance
'''
distance = geodesic((lat1,lon1), (lat2,lon2)).m
return distance
def get_neighbors(self, n):
'''
TO DO: To get the neighbors and weights of node n
'''
neighbors_list = []
reshape_graph = np.array(self.graph).reshape(-1, 3)
# if n is at the first column
for array in reshape_graph:
if n in array[:-2]:
array = np.delete(array, np.where(array == n))
neighbors_list.append(array)
return neighbors_list
def dijkstra(self, start_vertex):
D = {v:float('inf') for v in self.node_list}
D[start_vertex] = 0
pre_nodes = {}
pq = queue.PriorityQueue()
pq.put((0, start_vertex))
while not pq.empty():
(dist, current_vertex) = pq.get()
self.visited.append(current_vertex)
for node in self.node_list:
for edge in self.graph:
if current_vertex == edge[0] and node == edge[1]:
distance = edge[2]
if node not in self.visited:
old_cost = D[node]
new_cost = D[current_vertex] + distance
if new_cost < old_cost:
pq.put((new_cost, node))
D[node] = new_cost
pre_nodes[node] = current_vertex
return D, pre_nodes
def bellman_ford(self, src):
q = queue.Queue()
inqueue = {v:False for v in self.node_list}
inqueue[src] = True
distance = {v:float('inf') for v in self.node_list}
distance[src] = 0
q.put(src)
pre_nodes = {}
while(q.empty() != True):
node = q.get()
for edge in self.graph:
if node == edge[0]:
end_node = edge[1]
weight = edge[2]
if distance[end_node] > distance[node] + weight:
distance[end_node] = distance[node] + weight
pre_nodes[end_node] = node
if inqueue[end_node] == False:
q.put(end_node)
inqueue[end_node] == True
return distance, pre_nodes
def a_star_algorithm(self, start, stop, graph):
# In this open_lst is a list of nodes which have been visited, but who's
# neighbours haven't all been always inspected, It starts off with the start node
# And closed_lst is a list of nodes which have been visited
# and who's neighbors have been always inspected
open_lst = set([start])
closed_lst = set([])
lat1 = graph.nodes[start]['y']
lon1 = graph.nodes[start]['x']
# poo has present distances from start to all other nodes
# the default value is +infinity
poo = {}
poo[start] = 0
# par contains an adjac mapping of all nodes
par = {}
par[start] = start
while len(open_lst) > 0:
n = None
# it will find a node with the lowest value of f() -
for v in open_lst:
lat2 = graph.nodes[v]['y']
lon2 = graph.nodes[v]['x']
if n == None or poo[v] + self.h(lat1, lon1, lat2, lon2) < poo[n] + self.h(lat1, lon1, lat2n, lon2n):
n = v
lat2n = graph.nodes[v]['y']
lon2n = graph.nodes[v]['x']
if n == None:
print('Path does not exist!')
return None
# if the current node is the stop
# then we start again from start
if n == stop:
reconst_path = []
while par[n] != n:
reconst_path.append(n)
n = par[n]
reconst_path.append(start)
print('Path found: {}'.format(reconst_path))
return reconst_path
# for all the neighbors of the current node do
neighbor_list = self.get_neighbors(n)
for array in neighbor_list:
m, weight = array[0], array[1]
# if the current node is not presentin both open_lst and closed_lst
# add it to open_lst and note n as it's par
if m not in open_lst and m not in closed_lst:
open_lst.add(m)
par[m] = n
poo[m] = poo[n] + weight
# otherwise, check if it's quicker to first visit n, then m
# and if it is, update par data and poo data
# and if the node was in the closed_lst, move it to open_lst
else:
if poo[m] > poo[n] + weight:
poo[m] = poo[n] + weight
par[m] = n
if m in closed_lst:
closed_lst.remove(m)
open_lst.add(m)
# remove n from the open_lst, and add it to closed_lst
# because all of his neighbors were inspected
open_lst.remove(n)
closed_lst.add(n)
print('Path does not exist!')
return None
def print_result(previous_nodes, shortest_path, start_node, target_node):
path = []
node = target_node
while node != start_node:
path.append(node)
node = previous_nodes[node]
# Add the start node manually
path.append(int(start_node))
print("We found the following best path with a value of {}.".format(shortest_path[target_node]))
print(path)
return path, "{:.2f}".format(shortest_path[target_node])
# +
# Dijkstra Algorithm
g = Graph(nodes.shape[0], list(nodes.index))
for t in NodesAndLength:
g.add_edge(t[0], t[1], t[2])
D, pre_nodes1 = g.dijkstra(orig_node)
dij_path, dij_length = print_result(pre_nodes1, D, start_node=orig_node, target_node=dest_node)
dij_path.reverse()
dij_length
# +
# Bellman-Ford Algorithm
g = Graph(nodes.shape[0], list(nodes.index))
for t in NodesAndLength:
g.add_edge(t[0], t[1], t[2])
D, pre_nodes2 = g.bellman_ford(orig_node)
bell_path, bell_length = print_result(pre_nodes2, D, start_node=orig_node, target_node=dest_node)
bell_path.reverse()
bell_length
# +
# Astar Algorithm
g = Graph(nodes.shape[0], list(nodes.index))
for t in NodesAndLength:
g.add_edge(t[0], t[1], t[2])
astar_path = g.a_star_algorithm(orig_node, dest_node, graph)
astar_path.reverse()
# Convert float to integer
for i in range(0,len(astar_path)):
astar_path[i] = int(astar_path[i])
astar_path
# +
# Computer length of Astar's path
astar_length = 0
for i in range(len(astar_path)-1):
subEdge = (astar_path[i],astar_path[i+1])
astar_length += EdgeAndLength[subEdge]
astar_length = "{:.2f}".format(astar_length)
# +
# Plot the shortest route on Openstreet Map
dijkstra_map = ox.plot_route_folium(graph, dij_path, popup_attribute="length", weight=10, color='lightblue')
bellman_map = ox.plot_route_folium(graph, bell_path, popup_attribute="length", weight=10, color='purple')
astar_map = ox.plot_route_folium(graph, astar_path, popup_attribute="length", weight=10, color='grey')
maps = [dijkstra_map,bellman_map,astar_map]
for i in maps:
folium.TileLayer('openstreetmap').add_to(i)
folium.TileLayer('Stamen Terrain').add_to(i)
folium.TileLayer('Stamen Toner').add_to(i)
folium.TileLayer('Stamen Water Color').add_to(i)
folium.TileLayer('cartodbpositron').add_to(i)
folium.TileLayer('cartodbdark_matter').add_to(i)
folium.LayerControl().add_to(i)
# Marker class only accepts coordinates in tuple form
start_marker = folium.Marker(
location = (start_lat,start_long),
popup = "Departure",
icon = folium.Icon(color='black'))
end_marker = folium.Marker(
location = (end_lat,end_long),
popup = "Destination",
icon = folium.Icon(color='green'))
# add the circle marker to the map
start_marker.add_to(i)
end_marker.add_to(i)
mapNames = ["Dijkstra","Bellman-Ford","A*"]
pathLengths = [dij_length,bell_length,astar_length]
for i in range(len(mapNames)):
# Add title
text = "Shortest path calculated with "+mapNames[i]+" algorithm - Total distance = "+str(pathLengths[i])+" meters"
title_html = '''
<h3 align="center" style="font-size:16px"><b>{}</b></h3>
'''.format(text)
maps[i].get_root().html.add_child(folium.Element(title_html))
dijkstra_map
# +
# Save the output map as html file
dijkstra_map.save("./dijkstra_map.html")
bellman_map.save("./bellman_map.html")
astar_map.save("./astar_map.html")
print("\n=========================\nMaps with plotted shortest path are stored in current folder, please open it with your browswer to view the path :)\n=== Program finished ===")
| Navigator.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.12 ('phys_fast')
# language: python
# name: python3
# ---
# +
# %load_ext autoreload
# %autoreload 2
import numpy as np
from koala import pointsets
from matplotlib import pyplot as plt
from koala import voronization
from koala import plotting
from koala.lattice import cut_boundaries
# +
from koala.example_graphs import *
points1 = pointsets.generate_random(50)
points2 = pointsets.generate_bluenoise(30,3,3)
graphs = [
tri_square_pent(),
two_tri(),
tutte_graph(),
n_ladder(6,True),
bridge_graph(),
voronization.generate_lattice(points2),
cut_boundaries(voronization.generate_lattice(points2), [False,True]),
cut_boundaries(voronization.generate_lattice(points2), [True,True]),
generate_honeycomb(8),
cut_boundaries(generate_honeycomb(8), [False,True]),
cut_boundaries(generate_honeycomb(8), [True,True]),
concave_plaquette()
]
# +
# now we plot the plaquettes
fig_width = 4
fig_depth = int(np.ceil(len(graphs)/fig_width))
width_size = 20
width_depth = (fig_depth*width_size)/fig_width
fig, axes = plt.subplots(fig_depth,4,figsize = (20,15))
for n,ax in enumerate(axes.flatten()[:len(graphs)]):
ax.set_title('n_plaquettes = '+ str(graphs[n].n_plaquettes))
plotting.plot_lattice(graphs[n], ax)
cmap = plt.get_cmap("tab10")
plaq_labels = np.arange(graphs[n].n_plaquettes)
color_scheme = cmap(plaq_labels % 10)
plotting.plot_plaquettes(graphs[n], plaq_labels, color_scheme = color_scheme, ax = ax, alpha = 0.3)
centers = np.array([p.center for p in graphs[n].plaquettes])
ax.scatter(centers[:,0], centers[:,1])
# -
| examples/pathological_graphs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + tags=["parameters"]
epochs = 10
# We don't use the whole dataset for efficiency purpose, but feel free to increase these numbers
n_train_items = 640
n_test_items = 640
# -
# # পার্ট এক্স - এমএনআইএসটিতে নিরাপদ প্রশিক্ষণ এবং মূল্যায়ন (Part X - Secure Training and Evaluation on MNIST)
#
# সার্ভিস সলিউশন (এমএলএএস/MLaaS) হিসাবে মেশিন লার্নিং তৈরি করার সময়, কোনও সংস্থাকে তার মডেলটি প্রশিক্ষণের জন্য অন্যান্য অংশীদারদের থেকে ডেটা অ্যাক্সেসের জন্য অনুরোধ করতে হবে। স্বাস্থ্য বা অর্থায়নে, মডেল এবং ডেটা উভয়ই অত্যন্ত সমালোচনামূলক: মডেল পরামিতিগুলি একটি ব্যবসায়িক সম্পদ হয় যখন ডেটা ব্যক্তিগত ডেটা থাকে যা শক্তভাবে নিয়ন্ত্রিত হয়।
#
# এই প্রসঙ্গে, একটি সম্ভাব্য সমাধান হ'ল মডেল এবং ডেটা উভয়ই এনক্রিপ্ট করা এবং মেশিন লার্নিং মডেলকে এনক্রিপ্ট করা মানগুলির উপরে প্রশিক্ষণ দেওয়া। এটি গ্যারান্টি দেয় যে সংস্থা উদাহরণস্বরূপ রোগীদের চিকিত্সার রেকর্ডগুলিতে অ্যাক্সেস করবে না এবং যে স্বাস্থ্য সুবিধাগুলি তারা যে মডেলটির অবদান রাখবে তা পর্যবেক্ষণ করতে সক্ষম হবে না। বেশ কয়েকটি এনক্রিপশন স্কিম বিদ্যমান যা এনক্রিপ্ট হওয়া ডেটাগুলিতে গণনার অনুমতি দেয়, যার মধ্যে সিকিউর মাল্টি-পার্টির গণনা (এসএমপিসি/SMPC), হোমোমর্ফিক এনক্রিপশন (FHE/SHE) এবং ফাংশনাল এনক্রিপশন (FE) রয়েছে। আমরা এখানে মাল্টি পার্টি পার্টি কম্পিউটেশনে ফোকাস করব (যা টিউটোরিয়াল 5 তে প্রবর্তন করা হয়েছে) যা ব্যক্তিগত যোগমূলক ভাগ করে নেওয়ার সাথে সাথে ক্রিপ্টো প্রোটোকল সিকিউরএনএন এবং এসপিডিজেডের উপর নির্ভর করে।
#
# এই টিউটোরিয়ালটির সঠিক সেটিংটি হ'ল: আপনি সার্ভার এবং আপনি model n $ শ্রমিকদের দ্বারা রাখা কিছু ডেটাতে আপনার মডেলটিকে প্রশিক্ষণ দিতে চান তা বিবেচনা করুন। সার্ভার সিক্রেট তার মডেল ভাগ করে এবং প্রতিটি ভাগ কর্মীর কাছে প্রেরণ করে। কর্মীরা গোপনীয়তার সাথে তাদের ডেটা ভাগ করে এবং তাদের মধ্যে এটি বিনিময় করে। আমরা যে কনফিগারেশনটি অধ্যয়ন করব সেগুলিতে 2 জন কর্মী আছেন: এলিস এবং বব। শেয়ার বিনিময় করার পরে, তাদের প্রত্যেকের এখন নিজস্ব একটি শেয়ার, অন্য শ্রমিকের একটি ভাগ এবং মডেলের একটি অংশ। গণনা এখন উপযুক্ত ক্রিপ্টো প্রোটোকল ব্যবহার করে মডেলটিকে ব্যক্তিগতভাবে প্রশিক্ষণ দেওয়া শুরু করতে পারে। মডেলটি প্রশিক্ষিত হয়ে গেলে, সমস্ত শেয়ার এটি ডিক্রিপ্ট করার জন্য সার্ভারে ফিরে পাঠানো যেতে পারে। এটি নিম্নলিখিত চিত্র সহ চিত্রিত:
# 
# এই প্রক্রিয়াটির উদাহরণ দেওয়ার জন্য, আসুন ধরে নেওয়া যাক এলিস এবং বব দুজনেই এমএনআইএসটি ডেটাসেটের একটি অংশ ধরে রাখি এবং আসুন ডিজিটের শ্রেণিবিন্যাস সম্পাদনের জন্য একটি মডেলকে প্রশিক্ষণ দিন!
#
# লেখক:
# - <NAME> - Twitter: [@theoryffel](https://twitter.com/theoryffel) · GitHub: [@LaRiffle](https://github.com/LaRiffle)
# অনুবাদক:
#
# - <NAME> - Twitter: [@adventuroussrv](https://twitter.com/adventuroussrv)
# # 1. এমএনআইএসটিতে এনক্রিপ্ট করা প্রশিক্ষণ ডেমো (Encrypted Training demo on MNIST)
# ## আমদানি এবং প্রশিক্ষণের কনফিগারেশন (Imports and training configuration)
# +
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from torchvision import datasets, transforms
import time
# -
# এই শ্রেণিটি প্রশিক্ষণের জন্য সমস্ত হাইপার-প্যারামিটারগুলি বর্ণনা করে। মনে রাখবেন যে এগুলি এখানে সর্বজনীন।
# +
class Arguments():
def __init__(self):
self.batch_size = 64
self.test_batch_size = 64
self.epochs = epochs
self.lr = 0.02
self.seed = 1
self.log_interval = 1 # Log info at each batch
self.precision_fractional = 3
args = Arguments()
_ = torch.manual_seed(args.seed)
# -
# পাইসাইফ্ট আমদানি এখানে। আমরা দুজন রিমোট কর্মীর সাথে যোগাযোগ করি যা কল হয়`alice` and `bob` এবং আরেকটি কর্মীকে অনুরোধ করুন `crypto_provider` আমাদের প্রয়োজন হতে পারে সমস্ত ক্রিপ্টো (crypto) প্রারম্ভিক(Primitive) কে দেয়।
# +
import syft as sy # import the Pysyft library
hook = sy.TorchHook(torch) # hook PyTorch to add extra functionalities like Federated and Encrypted Learning
# simulation functions
def connect_to_workers(n_workers):
return [
sy.VirtualWorker(hook, id=f"worker{i+1}")
for i in range(n_workers)
]
def connect_to_crypto_provider():
return sy.VirtualWorker(hook, id="crypto_provider")
workers = connect_to_workers(n_workers=2)
crypto_provider = connect_to_crypto_provider()
# -
# ## অনুমতি প্রাপ্তি এবং গোপন তথ্য ভাগ (Getting access and secret share data)
#
# এখানে আমরা একটি ইউটিলিটি ফাংশন (utility function) ব্যবহার করছি যা নিম্নলিখিত আচরণের অনুকরণ করে: আমরা ধরে নিই যে এমএনআইএসটি (MNIST) ডেটাসেটটি আমাদের প্রতিটি শ্রমিকের হাতে থাকা প্রতিটি অংশে বিতরণ করা হয়েছে। শ্রমিকরা তখন তাদের ডেটাগুলি ব্যাচে(batches) বিভক্ত করে এবং গোপনে তাদের ডেটা একে অপরের মধ্যে ভাগ করে দেয়। ফিরে আসা চূড়ান্ত বস্তুটি এই গোপন ভাগ করা ব্যাচগুলির একটি পুনরাবৃত্তিযোগ্য, যা আমরা **private data loader(ব্যক্তিগত ডেটা লোডার)** বলি। নোট করুন যে প্রক্রিয়া চলাকালীন স্থানীয় কর্মী (তাই আমাদের/so us) ডেটা অ্যাক্সেস ছিল না।
#
# আমরা যথারীতি একটি প্রশিক্ষণ এবং ব্যক্তিগত ডেটাসেট পরীক্ষা করি এবং ইনপুট এবং লেবেল উভয়ই গোপনে ভাগ করে নেওয়া হয়।
# +
def get_private_data_loaders(precision_fractional, workers, crypto_provider):
def one_hot_of(index_tensor):
"""
Transform to one hot tensor
Example:
[0, 3, 9]
=>
[[1., 0., 0., 0., 0., 0., 0., 0., 0., 0.],
[0., 0., 0., 1., 0., 0., 0., 0., 0., 0.],
[0., 0., 0., 0., 0., 0., 0., 0., 0., 1.]]
"""
onehot_tensor = torch.zeros(*index_tensor.shape, 10) # 10 classes for MNIST
onehot_tensor = onehot_tensor.scatter(1, index_tensor.view(-1, 1), 1)
return onehot_tensor
def secret_share(tensor):
"""
Transform to fixed precision and secret share a tensor
"""
return (
tensor
.fix_precision(precision_fractional=precision_fractional)
.share(*workers, crypto_provider=crypto_provider, requires_grad=True)
)
transformation = transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((0.1307,), (0.3081,))
])
train_loader = torch.utils.data.DataLoader(
datasets.MNIST('../data', train=True, download=True, transform=transformation),
batch_size=args.batch_size
)
private_train_loader = [
(secret_share(data), secret_share(one_hot_of(target)))
for i, (data, target) in enumerate(train_loader)
if i < n_train_items / args.batch_size
]
test_loader = torch.utils.data.DataLoader(
datasets.MNIST('../data', train=False, download=True, transform=transformation),
batch_size=args.test_batch_size
)
private_test_loader = [
(secret_share(data), secret_share(target.float()))
for i, (data, target) in enumerate(test_loader)
if i < n_test_items / args.test_batch_size
]
return private_train_loader, private_test_loader
private_train_loader, private_test_loader = get_private_data_loaders(
precision_fractional=args.precision_fractional,
workers=workers,
crypto_provider=crypto_provider
)
# -
# ## মডেল স্পেসিফিকেশন (Model specification)
#
# এখানে আমরা যে মডেলটি ব্যবহার করব তা এটি একটি সহজ তবে এটি [it has proved to perform reasonably well on MNIST](https://towardsdatascience.com/handwritten-digit-mnist-pytorch-977b5338e627)
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
self.fc1 = nn.Linear(28 * 28, 128)
self.fc2 = nn.Linear(128, 64)
self.fc3 = nn.Linear(64, 10)
def forward(self, x):
x = x.view(-1, 28 * 28)
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = self.fc3(x)
return x
# ## প্রশিক্ষণ এবং পরীক্ষার কার্যাদি (Training and testing functions)
#
# প্রশিক্ষণটি প্রায় যথারীতি করা হয়, আসল পার্থক্যটি হ'ল আমরা নেতিবাচক লগ-সম্ভাবনা negative log-likelihood (F.nll_loss in PyTorch) এর মতো ক্ষয়গুলি ব্যবহার করতে পারি না কারণ এসএমপিসি (SMPC) দিয়ে এই ফাংশনগুলি পুনরুত্পাদন করা বেশ জটিল। পরিবর্তে, আমরা একটি সরল গড়ের স্কোয়ার ত্রুটি হ্রাস(simpler Mean Square Error loss) ব্যবহার করি।
def train(args, model, private_train_loader, optimizer, epoch):
model.train()
for batch_idx, (data, target) in enumerate(private_train_loader): # <-- now it is a private dataset
start_time = time.time()
optimizer.zero_grad()
output = model(data)
# loss = F.nll_loss(output, target) <-- not possible here
batch_size = output.shape[0]
loss = ((output - target)**2).sum().refresh()/batch_size
loss.backward()
optimizer.step()
if batch_idx % args.log_interval == 0:
loss = loss.get().float_precision()
print('Train Epoch: {} [{}/{} ({:.0f}%)]\tLoss: {:.6f}\tTime: {:.3f}s'.format(
epoch, batch_idx * args.batch_size, len(private_train_loader) * args.batch_size,
100. * batch_idx / len(private_train_loader), loss.item(), time.time() - start_time))
# পরীক্ষার ফাংশন বদলায় না!
def test(args, model, private_test_loader):
model.eval()
test_loss = 0
correct = 0
with torch.no_grad():
for data, target in private_test_loader:
start_time = time.time()
output = model(data)
pred = output.argmax(dim=1)
correct += pred.eq(target.view_as(pred)).sum()
correct = correct.get().float_precision()
print('\nTest set: Accuracy: {}/{} ({:.0f}%)\n'.format(
correct.item(), len(private_test_loader)* args.test_batch_size,
100. * correct.item() / (len(private_test_loader) * args.test_batch_size)))
# ### প্রশিক্ষণ চালু করা যাক! (Let's launch the training!)
#
# এখানে কী ঘটছে সে সম্পর্কে কয়েকটি নোট। প্রথমত, আমরা গোপনে আমাদের কর্মীদের জুড়ে সমস্ত মডেল পরামিতি ভাগ করি। দ্বিতীয়ত, আমরা অপ্টিমাইজারের হাইপারপ্যারামিটারগুলি (optimizer's hyperparameters) স্থির নির্ভুলতায় রূপান্তর করি। নোট করুন যে আমাদের সেগুলি গোপনীয়ভাবে ভাগ করে নেওয়ার দরকার নেই কারণ এগুলি আমাদের প্রসঙ্গে প্রকাশ্য, তবে গোপনীয়ভাবে ভাগ করা মূল্যবোধগুলি সীমাবদ্ধ ক্ষেত্রগুলিতে বাস করে বলে আমাদের ধারাবাহিকভাবে ক্রিয়াকলাপ সম্পাদন করার জন্য তাদের এখনও ite `.fix_precision` ব্যবহার করে সীমাবদ্ধ ক্ষেত্রগুলিতে সরানো দরকার ওজন আপডেট $W \leftarrow W - \alpha * \Delta W$.
# +
model = Net()
model = model.fix_precision().share(*workers, crypto_provider=crypto_provider, requires_grad=True)
optimizer = optim.SGD(model.parameters(), lr=args.lr)
optimizer = optimizer.fix_precision()
for epoch in range(1, args.epochs + 1):
train(args, model, private_train_loader, optimizer, epoch)
test(args, model, private_test_loader)
# -
# তুমি এখানে! আপনি মাত্র 100% এনক্রিপ্টড প্রশিক্ষণ ব্যবহার করে এমএনআইএসটি ডেটাসেটের একটি ক্ষুদ্র ভগ্নাংশ ব্যবহার করে 75% নির্ভুলতা পাবেন!
# # 2. আলোচনা (Discussion)
#
# আসুন আমরা কী করেছি তা বিশ্লেষণ করে এনক্রিপ্ট করা প্রশিক্ষণের শক্তির আরও নিবিড় নজর দেওয়া যাক।
# ## 2.1 গণনার সময় (Computation time)
#
# প্রথম জিনিসটি স্পষ্টতই চলমান সময়! আপনি অবশ্যই লক্ষ্য করেছেন, এটি সাধারণ পাঠ্য প্রশিক্ষণের চেয়ে ধীর। বিশেষত, 64 টি আইটেমের 1 ব্যাচের ওপরে পুনরাবৃত্তিটি 3.2 সেকেন্ড লাগে যখন খাঁটি পাইটর্চে কেবল 13 মিমি থাকে। যদিও এটি কোনও ব্লকারের মতো মনে হতে পারে, কেবল মনে রাখবেন যে এখানে সবকিছু দূর থেকে এবং এনক্রিপ্ট করা বিশ্বে ঘটেছিল: কোনও একক ডেটা আইটেম প্রকাশ করা হয়নি। আরও সুনির্দিষ্টভাবে বলা যায় যে, একটি আইটেমটি প্রক্রিয়া করার সময় 50ms যা খুব খারাপ নয়। আসল প্রশ্নটি হ'ল এনক্রিপ্ট করা প্রশিক্ষণের প্রয়োজন হয় এবং যখন কেবল এনক্রিপ্ট করা পূর্বাভাসই যথেষ্ট analy পূর্বাভাসটি সম্পাদনের 50ms কোনও উত্পাদন-প্রস্তুত দৃশ্যে সম্পূর্ণ গ্রহণযোগ্য!
#
# একটি প্রধান বাধা ব্যয়বহুল অ্যাক্টিভেশন ফাংশন ব্যবহার: এসএমপিসির (SMPC) সাথে রিলু (ReLU) অ্যাক্টিভেশন খুব ব্যয়বহুল কারণ এটি ব্যক্তিগত তুলনা এবং সিকিউরএনএন (SecureNN) প্রোটোকল ব্যবহার করে। উদাহরণস্বরূপ, আমরা যদি ক্রিট্টোনেটসের (CryptoNets) মতো এনক্রিপ্ট করা গণনার ক্ষেত্রে বেশ কয়েকটি গবেষণাপত্রে করা হয় যেমন আমরা চতুষ্কোণীয় অ্যাক্টিভেশন (quadratic activation) দিয়ে রিলুকে প্রতিস্থাপন করি তবে আমরা 3.2 থেকে 1.2 তে নেমেছি।
#
# একটি সাধারণ নিয়ম হিসাবে, মূল ধারণাটি হ'ল প্রয়োজনীয় জিনিসগুলি কেবল এনক্রিপ্ট করা এবং এই টিউটোরিয়ালটি আপনাকে বোঝায় যে এটি কতটা সহজ হতে পারে।
# ## 2.2 এসএমপিসি সহ ব্যাকপ্রসারণ (Backpropagation with SMPC)
#
# আপনি বিস্মিত হতে পারেন যে আমরা সীমাবদ্ধ ক্ষেত্রগুলিতে পূর্ণসংখ্যার সাথে কাজ করলেও আমরা কীভাবে ব্যাকপ্রসারণ এবং গ্রেডিয়েন্ট আপডেটগুলি (backpropagation and gradient updates) করব। এটি করার জন্য, আমরা অটোগ্রাডটেনসর (AutogradTensor) নামে একটি নতুন সিফ্ট টেনসর তৈরি করেছি। আপনি যদি না ও দেখে থাকেন তবে এই টিউটোরিয়ালটি এটি নিবিড়ভাবে ব্যবহার করেছে! কোনও মডেলের ওজন মুদ্রণ (printing) করে এটি পরীক্ষা করা যাক:
model.fc3.bias
# এবং একটি তথ্য আইটেম (And a data item)
first_batch, input_data = 0, 0
private_train_loader[first_batch][input_data]
# আপনি পর্যবেক্ষণ করতে পারছেন অটোগ্রাডটেন্সার (AutogradTensor) আছে! এটি torch আবরণ এবং ফিক্সডপ্রেসিশনটেনসরের (FixedPrecisionTensor) মধ্যে বাস করে যা নির্দেশ করে যে মানগুলি এখন সীমাবদ্ধ ক্ষেত্রগুলিতে রয়েছে। এই অটোগ্র্যাডটেন্সারের (AutogradTensor) লক্ষ্য হ'ল যখন এনক্রিপ্ট করা মানগুলিতে অপারেশন করা হয় তখন গণনা গ্রাফ সংরক্ষণ করা। এটি কার্যকর কারণ কারণ ব্যাকপ্রোগেশনের (backpropagation) জন্য পিছনে কল করার সময়, এই অটোগ্রাডটেন্সার সমস্ত পশ্চাদপট ফাংশনগুলি ওভাররাইড (overrides) করে যা এনক্রিপ্ট করা গণনার সাথে সামঞ্জস্যপূর্ণ নয় এবং এই গ্রেডিয়েন্টগুলি কীভাবে গণনা করতে হবে তা নির্দেশ করে। উদাহরণস্বরূপ, গুণটি সম্পর্কে যা বিভার ট্রিপলস ট্রিক (Beaver triples trick) ব্যবহার করে করা হয়, আমরা সেই কৌশলটি আরও বেশি আলাদা করতে চাই না যে কোনও গুণকে আলাদা করা খুব সহজ হওয়া উচিত: $\partial_b (a \cdot b) = a \cdot \partial b$. এই গ্রেডিয়েন্টগুলি (gradients) কীভাবে গণনা করা যায় তা এখানে আমরা বর্ণনা করব:
#
# ```python
# class MulBackward(GradFunc):
# def __init__(self, self_, other):
# super().__init__(self, self_, other)
# self.self_ = self_
# self.other = other
#
# def gradient(self, grad):
# grad_self_ = grad * self.other
# grad_other = grad * self.self_ if type(self.self_) == type(self.other) else None
# return (grad_self_, grad_other)
# ```
#
# আপনি একবার তাকান করতে পারেন `tensors/interpreters/gradients.py` আমরা কীভাবে আরও গ্রেডিয়েন্ট প্রয়োগ করেছি তা যদি জানতে আগ্রহী হন।
#
# গণনা গ্রাফের ক্ষেত্রে, এর অর্থ গ্রাফের একটি অনুলিপি স্থানীয় রয়ে গেছে এবং যে সার্ভারটি ফরোয়ার্ড পাসের সাথে সমন্বয় সাধন করে তা পিছিয়ে পাস কীভাবে করতে হয় তার নির্দেশাবলীও সরবরাহ করে। এটি আমাদের সেটিংয়ে একটি সম্পূর্ণ বৈধ অনুমান।
# ## 2.3 সুরক্ষা গ্যারান্টি (Security guarantees)
#
#
# সর্বশেষে, আসুন আমরা এখানে যে সুরক্ষা পাচ্ছি সে সম্পর্কে কয়েকটি ইঙ্গিত দেওয়া যাক: আমরা এখানে যে বিরোধীরা বিবেচনা করছি তারা হলেন **সৎ কিন্তু কৌতূহলী (honest but curious)**: এর অর্থ হল যে কোনও শত্রুরা এই প্রোটোকলটি চালিয়ে ডেটা সম্পর্কে কিছুই জানতে পারে না, তবে একটি দূষিত শত্রু এখনও প্রোটোকল থেকে বিচ্যুত হতে পারে এবং উদাহরণস্বরূপ গণনাটি নাশকতার জন্য শেয়ারকে দূষিত করার চেষ্টা করে। বেসরকারী তুলনা সহ এসএমপিসি (SMPC) গণনাগুলিতে দূষিত বিরোধীদের বিরুদ্ধে সুরক্ষা এখনও একটি উন্মুক্ত সমস্যা।
#
# তদতিরিক্ত, সিকিওর মাল্টি-পার্টির (Secure Multi-Party) গণনাও যদি নিশ্চিত করে যে প্রশিক্ষণের ডেটা অ্যাক্সেস করা হয়নি তবে প্লেইন টেক্সট ওয়ার্ল্ড থেকে অনেক হুমকি এখনও এখানে উপস্থিত রয়েছে। উদাহরণস্বরূপ, আপনি যেমন মডেলটির কাছে অনুরোধ করতে পারেন (MLaaS প্রসঙ্গে), আপনি ভবিষ্যদ্বাণীগুলি পেতে পারেন যা প্রশিক্ষণ ডেটাসেট সম্পর্কে তথ্য প্রকাশ করতে পারে। বিশেষত সদস্যপদ আক্রমণগুলির বিরুদ্ধে আপনার কোনও সুরক্ষা নেই, মেশিন লার্নিং পরিষেবাগুলিতে একটি সাধারণ আক্রমণ যেখানে বিরোধীরা নির্ধারণ করতে চায় যে কোনও নির্দিষ্ট আইটেমটি ডেটাসেটে ব্যবহৃত হয়েছিল কিনা। এগুলি ছাড়াও অন্যান্য আক্রমণ যেমন অনিচ্ছাকৃত মুখস্থকরণ প্রক্রিয়াগুলি (কোনও ডেটা আইটেমের সুনির্দিষ্ট বৈশিষ্ট্য শেখা মডেল), মডেল বিপরীতকরণ বা নিষ্কাশন এখনও সম্ভব।
#
# উপরে উল্লিখিত হুমকির অনেকের জন্য কার্যকর একটি সাধারণ সমাধান হ'ল ডিফারেনশিয়াল প্রাইভেসি যুক্ত করা। এটি সিকিউর মাল্টি-পার্টির গণনার সাথে সুন্দরভাবে মিলিত হতে পারে এবং খুব আকর্ষণীয় সুরক্ষা গ্যারান্টি সরবরাহ করতে পারে। আমরা বর্তমানে বেশ কয়েকটি বাস্তবায়নের উপর কাজ করছি এবং আশা করি একটি উদাহরণ প্রস্তাব করুন যা উভয়ই সংক্ষেপে সংযুক্ত করা হবে!
# # উপসংহার (Conclusion)
#
# যেমনটি আপনি দেখেছেন, এসএমপিসি (SMPC) ব্যবহার করে কোনও মডেলকে প্রশিক্ষণ দেওয়া কোডের দৃষ্টিকোণ থেকে জটিল নয়, এমনকি আমরা হুডের (hood) নিচে জটিল বস্তুগুলিও ব্যবহার করি। এটি মাথায় রেখে, আপনার প্রশিক্ষণের জন্য বা মূল্যায়নের জন্য কখন এনক্রিপ্ট করা গণনা প্রয়োজন তা দেখার জন্য আপনার এখন ব্যবহারের ক্ষেত্রে বিশ্লেষণ করা উচিত। যদি এনক্রিপ্ট করা গণনা সাধারণভাবে খুব ধীর হয় তবে এটি সাবধানতার সাথে ব্যবহার করা যেতে পারে যাতে সামগ্রিক গণনা ওভারহেড হ্রাস পায়।
#
# আপনি যদি এটি উপভোগ করেন এবং গোপনীয়তা সংরক্ষণ, AI এবং AI সরবরাহ চেইনের (ডেটা) বিকেন্দ্রীভূত মালিকানার দিকে আন্দোলনে যোগ দিতে চান, আপনি নিম্নলিখিত উপায়ে এটি করতে পারেন!
#
# ### গিটহাবে পাইসিফ্ট কে স্টার দিন (Star PySyft on GitHub)
#
# আমাদের সম্প্রদায়কে সাহায্য করার সবচেয়ে সহজ উপায় হ'ল রিপোসিটোরি গুলোতে ষ্টার করা
# এটি আমরা যে অসাধারণ সরঞ্জামগুলি তৈরি করছি তার সচেতনতা বাড়াতে সহায়তা করে।
#
# - [Star PySyft](https://github.com/OpenMined/PySyft)
#
# ### গিটহাবে আমাদের টিউটোরিয়ালগুলি চয়ন করুন! (Pick our tutorials on GitHub!)
#
# ফেডারেটেড এবং প্রাইভেসি-প্রিজারভেভিং লার্নিংয়ের ( Federated and Privacy-Preserving Learning) দৃশ্য কেমন হওয়া উচিত এবং আমরা এটির জন্য bricks কীভাবে তৈরি করছি সে সম্পর্কে আরও ভাল ধারণা পেতে আমরা সত্যিই দুর্দান্ত টিউটোরিয়াল তৈরি করেছি।
#
# - [Checkout the PySyft tutorials](https://github.com/OpenMined/PySyft/tree/master/examples/tutorials)
#
#
# ### আমাদের স্ল্যাক যোগ দিন! (Join our Slack!)
#
# সর্বশেষতম অগ্রগতিতে আপ টু ডেট রাখার সর্বোত্তম উপায় হ'ল আমাদের সম্প্রদায়ে যোগদান করা!
#
# - [Join slack.openmined.org](http://slack.openmined.org)
#
# ### একটি কোড প্রকল্পে যোগদান করুন! (Join a Code Project!)
#
# আমাদের সম্প্রদায়ে অবদান রাখার সর্বোত্তম উপায় হ'ল কোড অবদানকারী হয়ে উঠুন! আপনি যদি মিনি-প্রকল্পগুলি "ওয়ান অফ" শুরু করতে চান তবে আপনি পাইসাইফ্ট গিটহাব ইস্যু পৃষ্ঠাতে গিয়ে চিহ্নিত বিষয়গুলির জন্য অনুসন্ধান করতে পারেন `Good First Issue`.
#
# - [Good First Issue Tickets](https://github.com/OpenMined/PySyft/issues?q=is%3Aopen+is%3Aissue+label%3A%22good+first+issue%22)
#
# ### দান করা (Donate)
#
# আপনার যদি আমাদের কোডবেসে অবদান রাখার সময় না থাকে তবে তবুও সমর্থন leণ দিতে চান, আপনি আমাদের ওপেন কালেক্টিভেরও ব্যাকের হয়ে উঠতে পারেন। সমস্ত অনুদান আমাদের ওয়েব হোস্টিং এবং অন্যান্য সম্প্রদায় ব্যয় যেমন হ্যাকাথনস এবং মেটআপগুলির দিকে যায়!
#
# - [Donate through OpenMined's Open Collective Page](https://opencollective.com/openmined)
| examples/tutorials/translations/bengali/Part 12 bis - Encrypted Training on MNIST.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # High-level introduction to <code>pandas</code>
# For a more complete introduction to <code>pandas</code>, see [https://pandas.pydata.org/](https://pandas.pydata.org/).
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# ## Series
# Series is a one-dimensional labeled array capable of holding any data type (integers, strings, floating point numbers, Python objects, etc.).
data = np.random.randn(20)
index = range(1990, 2010)
print (data)
print (index)
y = pd.Series(data, index=index)
print (y)
salaries = {
'gino': 1500, 'maria': 2560.34, 'luca': None, 'federico': 2451
}
s = pd.Series(salaries)
print (s)
k = pd.Series({
'a': 'v', 'b': None
})
print (k)
# ### Access series as arrays
print (s[:2], '\n')
print (s[s > s.median()], '\n')
print (np.log(s), '\n')
print (s + s, '\n')
print (s * 3, '\n')
print (y[4:8] + y[4:10])
# ### Access series as dictionaries
# # Data Frames
# From [http://pandas.pydata.org/pandas-docs/stable/dsintro.html#dataframe](http://pandas.pydata.org/pandas-docs/stable/dsintro.html#dataframe)
#
# DataFrame is a 2-dimensional labeled data structure with columns of potentially different types. You can think of it like a spreadsheet or SQL table, or a dict of Series objects. It is generally the most commonly used pandas object. Like Series, DataFrame accepts many different kinds of input:
#
# - Dict of 1D ndarrays, lists, dicts, or Series
# - 2-D numpy.ndarray
# - Structured or record ndarray
# - A Series
# - Another DataFrame
#
# Along with the data, you can optionally pass index (row labels) and columns (column labels) arguments. If you pass an index and / or columns, you are guaranteeing the index and / or columns of the resulting DataFrame. Thus, a dict of Series plus a specific index will discard all data not matching up to the passed index.
#
# If axis labels are not passed, they will be constructed from the input data based on common sense rules.
k = {'years': y, 'salaries': s}
df = pd.DataFrame(k)
print (df)
data = {}
for k, v in s.items():
data[k] = {}
for d, w in y.items():
data[k][d] = v + (v*w)
ydf = pd.DataFrame(data)
print (ydf)
ydf.head()
pd.DataFrame.from_dict(data, orient='index').head()
# ## Loading and manipulating data
football = '../data/fifa/fifa_2018.csv'
fifa = pd.read_csv(football)
fifa.head()
# +
selected_cols = [
'ID', 'name', 'club', 'league', 'height_cm', 'weight_kg', 'nationality', 'eur_value', 'overall',
'short_passing', 'long_passing', 'finishing', 'shot_power', 'marking', 'standing_tackle',
'sliding_tackle', 'gk_diving', 'gk_handling', 'gk_kicking', 'gk_positioning', 'gk_reflexes'
]
positions = ['rs', 'rw', 'rf', 'ram', 'rcm', 'rm', 'rdm', 'rcb', 'rb', 'rwb', 'st', 'lw', 'cf', 'cam',
'cm', 'lm', 'cdm', 'cb', 'lb', 'lwb', 'ls', 'lf', 'lam', 'lcm', 'ldm', 'lcb', 'gk']
selected_cols += positions
# -
F = fifa[selected_cols]
print (fifa.shape)
print (F.shape)
new_columns = {
'atk': ['finishing', 'shot_power'],
'dfs': ['marking', 'standing_tackle', 'sliding_tackle'],
'psg': ['short_passing', 'long_passing'],
'gkv': ['gk_diving', 'gk_handling', 'gk_kicking', 'gk_positioning', 'gk_reflexes']
}
# +
S = F.copy()
for k, v in new_columns.items():
S[k] = F[v].mean(axis=1) / 100
S = S.drop(v, axis=1)
S['role'] = S[positions].idxmax(axis=1)
S = S.drop(positions, axis=1)
# -
S.head()
M = np.array(S[list(new_columns.keys())])
M
S.role.unique()
S.plot(kind='scatter', x='atk', y='dfs', figsize=(10,7),
s=S['eur_value']/1000000.0, label='players',
cmap=plt.get_cmap('jet'),
alpha=0.2, c=S['overall']/100.0, colorbar=True)
plt.legend()
plt.show()
S[S.role == 'ram'].hist(bins=50, figsize=(20,15))
plt.show()
# ## Access dataframe by index and col
S.iloc[2:6] # gets rows (or columns) at particular positions in the index (so it only takes integers).
S.loc[2:6] # gets rows (or columns) with particular labels from the index
S[S['club'] == 'Paris Saint-Germain'].sort_values('overall', ascending=False)[:5]
S[S['league']=='French Ligue 1'][['club', 'overall', 'atk', 'dfs']].groupby('club').median()
| playground/game/pd_introduction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: ROS Python 2
# language: python
# name: ros_python
# ---
import rospy as rp
import jupyros as jr
from __future__ import print_function
from geometry_msgs.msg import Pose, Vector3
rp.init_node('runner')
from geometry_msgs.msg import Pose
import math
# %%thread_cell
from geometry_msgs.msg import Pose, Vector3
import rospy as rp
import math
i = 0
rate = rp.Rate(5)
pub = rp.Publisher('/poser', Vector3, queue_size=10)
while True:
msg = Vector3()
msg.x = math.sin(i * 0.1)
msg.y = math.cos(i * 0.1)
pub.publish(msg)
i += 1
rate.sleep()
jr.subscribe('/poser', Vector3, lambda msg: print(msg.x, msg.y))
plot_string = '/poser/:x:y'
topic = plot_string[:plot_string.find(':') - 1]
fields = plot_string.split(':')[1:]
jr.live_plot('/poser/:x:y', Vector3)
# !rqt_plot /poser
| notebooks/ROS Live Plotting.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # IST256 Lesson 07
# ## Files
#
# - Zybook Ch7
# - P4E Ch7
#
# ## Links
#
# - Participation: [https://poll.ist256.com](https://poll.ist256.com)
# - Zoom Chat!
# + [markdown] slideshow={"slide_type": "slide"}
# # Agenda
#
# ### Go Over Homework H06
#
#
# ### New Stuff
# - The importance of a persistence layer in programming.
# - How to read and write from files.
# - Techniques for reading a file a line at a time.
# - Using exception handling with files.
#
# + [markdown] slideshow={"slide_type": "slide"}
# # FEQT (Future Exam Questions Training) 1
#
# What is the output of the following code when `berry` is input on line `1`?
# + slideshow={"slide_type": "fragment"}
x = input()
if x.find("rr")!= -1:
y = x[1:]
else:
y = x[:-1]
print(y)
# + [markdown] slideshow={"slide_type": "fragment"}
# A. erry
# B. berr
# C. berry
# D. bey
#
# ## Vote Now: [https://poll.ist256.com](https://poll.ist256.com)
# + [markdown] slideshow={"slide_type": "slide"}
# # FEQT (Future Exam Questions Training) 2
#
# What is the output of the following code when `mike is cold` is input on line `1`?
# + slideshow={"slide_type": "fragment"}
x = input()
y = x.split()
w = ""
for z in y:
w = w + z[1]
print(w)
# + [markdown] slideshow={"slide_type": "fragment"}
# A. iic
# B. ike
# C. mic
# D. iso
#
# ## Vote Now: [https://poll.ist256.com](https://poll.ist256.com)
# + [markdown] slideshow={"slide_type": "slide"}
# # FEQT (Future Exam Questions Training) 3
#
# What is the output of the following code when `tony` is input on line `1`?
# + slideshow={"slide_type": "fragment"}
x = input()
x = x + x
x = x.replace("o","i")
x = x[:5]
print(x)
# + [markdown] slideshow={"slide_type": "fragment"}
# A. tony
# B. tiny
# C. tinyt
# D. tonyt
#
# ## Vote Now: [https://poll.ist256.com](https://poll.ist256.com)
# + [markdown] slideshow={"slide_type": "skip"}
# # Connect Activity
#
# Which of the following is not an example of secondary (persistent) memory?
# A. `Flash Memory`
# B. `Hard Disk Drive (HDD)`
# C. `Random-Access Memory (RAM)`
# D. `Solid State Disk (SSD)`
#
#
# ### Vote Now: [https://poll.ist256.com](https://poll.ist256.com)
#
# + [markdown] slideshow={"slide_type": "slide"}
# # Files == Persistence
#
# - Files add a **Persistence Layer** to our computing environment where we can store our data **after the program completes**.
# - **Think: Saving a game's progress or saving your work!**
# - When our program **Stores** data, we open the file for **writing**.
# - When our program **Reads** data, we open the file for **reading**.
# - To read or write a file we must first **open** it, which gives us a special variable called a **file handle**.
# - We then use the **file handle** to read or write from the file.
# - The **read()** function reads from the **write()** function writes to the file through the file handle.
# + [markdown] slideshow={"slide_type": "slide"}
# # Reading From a File
#
# Two approaches... that's it!
# + slideshow={"slide_type": "fragment"}
# all at once
with open(filename, 'r') as handle:
contents = handle.read()
# a line at a time
with open(filename, 'r') as handle:
for line in handle.readlines():
do_something_with_line
# + [markdown] slideshow={"slide_type": "slide"}
# # Writing a To File
#
# + slideshow={"slide_type": "fragment"}
# write mode
with open(filename, 'w') as handle:
handle.write(something)
# append mode
with open(filename, 'a') as handle:
handle.write(something)
# + [markdown] slideshow={"slide_type": "slide"}
# # Watch Me Code 1
#
# ### Let’s Write two programs.
# - Save a text message to a file.
# - Retrieve the text message from the file.
#
# + [markdown] slideshow={"slide_type": "slide"}
# # Check Yourself: Which line 1
#
# - Which line number creates the file handle?
#
# + slideshow={"slide_type": "fragment"}
a = "savename.txt"
with open(a,'w') as b:
c = input("Enter your name: ")
b.write(c)
# + [markdown] slideshow={"slide_type": "fragment"}
# A. `1`
# B. `2`
# C. `3`
# D. `4`
# ### Vote Now: [https://poll.ist256.com](https://poll.ist256.com)
# + [markdown] slideshow={"slide_type": "slide"}
# # Watch Me Code 2
#
# Common patterns for reading and writing more than one item to a file.
# - Input a series of grades, write them to a file one line at a time.
# - Read in that file one line at a time, print average.
#
# + [markdown] slideshow={"slide_type": "slide"}
# # Check Yourself: Which line 2
#
# - On which line number does the file handle no longer exist?
# + slideshow={"slide_type": "fragment"}
with open("sample.txt","r") as f:
for line in f.readlines():
print(line)
g = "done"
# + [markdown] slideshow={"slide_type": "fragment"}
# A. `1`
# B. `2`
# C. `3`
# D. `4`
# ### Vote Now: [https://poll.ist256.com](https://poll.ist256.com)
# + [markdown] slideshow={"slide_type": "slide"}
# # Your Operating System and You
#
# - Files are stored in your **secondary memory** in **folders**.
# - When the python program is in the **same** folder as the file, **no path is required**.
# - When the file is in a **different** folder, **a path is required**.
# - **Absolute paths** point to a file starting at the root of the hard disk.
# - **Relative paths** point to a file starting at the current place on the hard disk.
#
# + [markdown] slideshow={"slide_type": "slide"}
# # Python Path Examples
#
# <table style="font-size:1.0em;">
# <thead><tr>
# <th>What</th>
# <th>Windows</th>
# <th>Mac/Linux</th>
# </tr></thead>
# <tbody>
# <tr>
# <td><code> File in current folder </code></td>
# <td> "file.txt" </td>
# <td> "file.txt"</td>
# </tr>
# <tr>
# <td><code> File up one folder from the current folder </code></td>
# <td> "../file.txt"</td>
# <td> "../file.txt"</td>
# </tr>
# <tr>
# <td><code> File in a folder from the current folder </code></td>
# <td> "folder1/file.txt" </td>
# <td> "folder1/file.txt" </td>
# </tr>
# <tr>
# <td><code> Absolute path to file in a folder</code></td>
# <td> "C:/folder1/file.txt" </td>
# <td> "/folder1/file.txt"</td>
# </tr>
# </tbody>
# </table>
# + [markdown] slideshow={"slide_type": "slide"}
# # Check Yourself: Path
#
# ### - Is this path relative or absolute?
#
#
# "/path/to/folder/file.txt"
#
# A. `Relative`
# B. `Absolute`
# C. `Neither`
# D. `Not sure`
#
# ### Vote Now: [https://poll.ist256.com](https://poll.ist256.com)
#
# + [markdown] slideshow={"slide_type": "slide"}
# # Handling Errors with Try…Except
#
# - I/O is the ideal use case for exception handling.
# - Don't assume you can read a file!
# - Use try… except!
#
# + slideshow={"slide_type": "fragment"}
try:
file = 'data.txt'
with open(file,'r') as f:
print( f.read() )
except FileNotFoundError:
print(f"{file} was not found!")
# + [markdown] slideshow={"slide_type": "slide"}
# # End-To-End Example (Pre-Recorded)
#
# #### How Many Calories in that Beer?
#
# Let's write a program to search a data file of 254 popular beers. Given the name of the beer the program will return the number of calories.
#
# Watch this here:
# https://youtu.be/s-1ToO0dJIs
#
# + [markdown] slideshow={"slide_type": "slide"}
# # End-To-End Example
#
# #### A Better Spell Check
#
# In this example, we create a better spell checker than the one from small group.
#
# - read words from a file
# - read text to check from a file.
#
# + [markdown] slideshow={"slide_type": "slide"}
# # Conclusion Activity : One Question Challenge
#
# What is wrong with the following code:
# + slideshow={"slide_type": "fragment"}
file = "a.txt"
with open(file,'w'):
file.write("Hello")
# + [markdown] slideshow={"slide_type": "fragment"}
# A. `No file handle`
# B. `Cannot write - file opened for reading`
# C. `File a.txt does not exist`
# D. `Nothing is wrong!`
#
# ### Vote Now: [https://poll.ist256.com](https://poll.ist256.com)
| content/lessons/07-Files/Slides.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a href="https://colab.research.google.com/github/pykeen/pykeen/blob/master/notebooks/results_plots/results_plots.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# # Results Plotting Demo
#
# This notebook serves to make some simple plots of the 1) losses and 2) entities and relations following training with the PyKEEN pipeline.
# ! python -c "import pykeen" || pip install git+https://github.com/pykeen/pykeen.git
# +
import os
import numpy as np
import pykeen
from matplotlib import pyplot as plt
from pykeen.pipeline import pipeline
from pykeen.triples import TriplesFactory
# %matplotlib inline
# %config InlineBackend.figure_format = 'svg'
# -
pykeen.env()
# ## Toy Example
#
# Following the disussions proposed in https://github.com/pykeen/pykeen/issues/97, a very small set of triples are trained and visualized.
# +
os.makedirs('results', exist_ok=True)
triples = '''
Brussels locatedIn Belgium
Belgium partOf EU
EU hasCapital Brussels
'''.strip()
triples = np.array([triple.split('\t') for triple in triples.split('\n')])
tf = TriplesFactory.from_labeled_triples(triples=triples)
# -
# Training with default arguments
results = pipeline(
training=tf,
testing=tf,
model = 'TransE',
model_kwargs=dict(embedding_dim=2),
training_kwargs=dict(use_tqdm_batch=False),
evaluation_kwargs=dict(use_tqdm=False),
random_seed=1,
device='cpu',
)
results.plot(er_kwargs=dict(plot_relations=True))
plt.savefig('results/toy_1.png', dpi=300)
# Training with slower learning and more epochs
results = pipeline(
training=tf,
testing=tf,
model = 'TransE',
model_kwargs=dict(embedding_dim=2),
optimizer_kwargs=dict(lr=1.0e-1),
training_kwargs=dict(num_epochs=128, use_tqdm_batch=False),
evaluation_kwargs=dict(use_tqdm=False),
random_seed=1,
device='cpu',
)
results.plot(er_kwargs=dict(plot_relations=True))
plt.savefig('results/toy_2.png', dpi=300)
# Training with appropriate softplus
toy_results = pipeline(
training=tf,
testing=tf,
model='TransE',
loss='softplus',
model_kwargs=dict(embedding_dim=2),
optimizer_kwargs=dict(lr=1.0e-1),
training_kwargs=dict(num_epochs=128, use_tqdm_batch=False),
evaluation_kwargs=dict(use_tqdm=False),
random_seed=1,
device='cpu',
)
toy_results.plot(er_kwargs=dict(plot_relations=True))
plt.savefig('results/toy_3.png', dpi=300)
# ## Benchmark Dataset Example
nations_results = pipeline(
dataset='Nations',
model='TransE',
model_kwargs=dict(embedding_dim=8),
optimizer_kwargs=dict(lr=1.0e-1),
training_kwargs=dict(num_epochs=80, use_tqdm_batch=False),
evaluation_kwargs=dict(use_tqdm=False),
random_seed=1,
device='cpu',
)
nations_results.plot(er_kwargs=dict(plot_relations=True))
# Filter the ER plot down to a specific set of entities and relations
nations_results.plot_er(
relations={'treaties'},
apply_limits=False,
plot_relations=True,
);
| notebooks/results_plots/results_plots.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: pytorch1.6.0
# language: python
# name: pytorch1.6.0
# ---
# ## 导入必要的包
# +
import torch
from torch import nn
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from sklearn.utils import shuffle
from sklearn.preprocessing import scale
from torchsummary import summary
from matplotlib.font_manager import FontProperties
torch.__version__
# -
# ## 数据读取
# 本次数据读取将采用两种方式进行,一种是从原始的`housing.csv`文件进行读取,一种是利用`sklearn`集成的数据读取方式进行读取
# ### 从原始数据读取房价信息
data_frame = pd.read_csv('./boston/housing.csv', header=None)
# 原始的`housing.csv`将一行数据存储在一个单元格中,因此需要读取此单元格并将里面的内容从字符串转变为`float`类型
# +
all_data = []
for column in range(len(data_frame)):
column_data = []
for data in list(data_frame.iloc[column])[0].split(' '):
if data != '':
column_data.append(float(data))
all_data.append(column_data)
all_data = np.array(all_data)
# -
# 划分训练数据与标签数据`(GroundTruth)`
x_data = all_data[:, :13]
y_data = all_data[:, 13]
# 查看部分数据
print('x_data:\n', x_data, '\n x_data shape:', x_data.shape,
'\ny_data:\n', y_data, '\n y_data shape:', y_data.shape)
# ### sklearn自动读取
# 利用`sklearn`中的`load_boston`直接获取
# +
from sklearn.datasets import load_boston
boston = load_boston()
# -
# 获取数据键
boston.keys()
# 获取训练数据
boston['data'].shape
# 获取特征名
boston['feature_names']
# 获取一则训练数据
boston['data'][0]
# 获取标签数据`(GroundTruth)`
boston['target']
# ### 对比两种方式所获取数据
# 对比训练数据是否相同
(x_data == boston['data']).all()
# 对比标签数据`(GroundTruth)`是否相同
(y_data == boston['target']).all()
# ## 训练阶段
#
# 获取设备信息
# +
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
device
# -
# 构建模型
class Net(nn.Module):
def __init__(self, input_num, hidden_num, output_num):
super(Net, self).__init__()
self.net = nn.Sequential(
nn.Linear(input_num, hidden_num),
nn.ReLU(),
nn.Linear(hidden_num, output_num),
nn.ReLU()
)
def forward(self, input):
return self.net(input)
# +
net = Net(input_num=13, hidden_num=14, output_num=1).to(device)
summary(net, input_size=(13,))
# -
# 使用`scale`进行归一化,使用`unsqueeze`整理维度
x_data, y_data = torch.FloatTensor(scale(x_data)).to(device), torch.unsqueeze(torch.FloatTensor(y_data), dim=1).to(device)
# 划分训练集及验证集
train_x, train_y = x_data[0:400, :], y_data[0:400]
valid_x, valid_y = x_data[400:, :], y_data[400:]
# 设置超参数
epochs = 200
learning_rate = 0.001
batch_size = 10
total_step = int(train_x.shape[0] / batch_size)
# 定义优化器以及损失函数
# +
optimizer = torch.optim.Adam(net.parameters(), lr=learning_rate)
loss_func = torch.nn.MSELoss()
# -
# 定义参数重置函数,保证每次重新执行`for循环`时从`零`开始训练
def weight_reset(m):
if isinstance(m, nn.Conv2d) or isinstance(m, nn.Linear):
m.reset_parameters()
# +
# %%time
net.apply(weight_reset)
epoch_train_loss_value = []
step_train_loss_value = []
epoch_valid_loss_value = []
for i in range(epochs):
for step in range(total_step):
xs = train_x[step * batch_size:(step + 1) * batch_size, :]
ys = train_y[step * batch_size:(step + 1) * batch_size]
prediction = net(xs)
loss = loss_func(prediction, ys)
optimizer.zero_grad()
loss.backward()
optimizer.step()
step_train_loss_value.append(loss.cpu().detach().numpy())
valid_loss = loss_func(net(valid_x), valid_y)
epoch_valid_loss_value.append(valid_loss)
epoch_train_loss_value.append(np.mean(step_train_loss_value))
print('epoch={:3d}/{:3d}, train_loss={:.4f}, valid_loss={:.4f}'.format(i + 1,
epochs,
np.mean(step_train_loss_value),
valid_loss))
# -
# 损失值可视化
# +
fig = plt.gcf()
fig.set_size_inches(10, 5)
plt.xlabel('Epochs', fontsize=15)
plt.ylabel('Loss', fontsize=15)
plt.plot(epoch_train_loss_value, 'blue', label='Train loss')
plt.plot(epoch_valid_loss_value, 'red', label='Valid loss')
plt.legend(loc='best')
plt.title('Training and Validation loss', fontsize=15)
plt.show()
# -
# ## 模型预测
# 遍历valid数据,求出每一组的预测值
# +
prediction = []
for i in range(valid_x.shape[0]):
prediction.append(net(valid_x[i, :]).item())
# -
# 预测值可视化,可以发现经过200轮的训练,预测值与真实值交融性很好,表明模型效果较好
# +
fig = plt.gcf()
fig.set_size_inches(10, 5)
myfont = FontProperties(fname='./font/msyh.ttc')
plt.title('预测值与实际值', fontproperties=myfont, fontsize=15)
plt.scatter(np.arange(len(prediction)), prediction, label='Prediction', s=20)
plt.scatter(np.arange(len(prediction)), valid_y.cpu().numpy(), label='GroundTruth', s=20)
plt.xlabel('数据/组', fontproperties=myfont, fontsize=15)
plt.ylabel('房价', fontproperties=myfont, fontsize=15)
plt.legend()
plt.show()
| 多元线性回归.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/NeuromatchAcademy/course-content/blob/master/tutorials/W3D2_DynamicNetworks/student/W3D2_Tutorial1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text"
# # Neuromatch Academy: Week 2, Day 4, Tutorial 1
# # Neuronal Network Dynamics: Neural Rate Models
#
# __Content creators:__ <NAME>, <NAME>, <NAME>, <NAME>, <NAME>
#
# __Content reviewers:__ <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>
#
# + [markdown] colab_type="text"
# ---
# # Tutorial Objectives
#
# The brain is a complex system, not because it is composed of a large number of diverse types of neurons, but mainly because of how neurons are connected to each other. The brain is indeed a network of highly specialized neuronal networks.
#
# The activity of a neural network constantly evolves in time. For this reason, neurons can be modeled as dynamical systems. The dynamical system approach is only one of the many modeling approaches that computational neuroscientists have developed (other points of view include information processing, statistical models, etc.).
#
# How the dynamics of neuronal networks affect the representation and processing of information in the brain is an open question. However, signatures of altered brain dynamics present in many brain diseases (e.g., in epilepsy or Parkinson's disease) tell us that it is crucial to study network activity dynamics if we want to understand the brain.
#
# In this tutorial, we will simulate and study one of the simplest models of biological neuronal networks. Instead of modeling and simulating individual excitatory neurons (e.g., LIF models that you implemented yesterday), we will treat them as a single homogeneous population and approximate their dynamics using a single one-dimensional equation describing the evolution of their average spiking rate in time.
#
# In this tutorial, we will learn how to build a firing rate model of a single population of excitatory neurons.
#
# **Steps:**
# - Write the equation for the firing rate dynamics of a 1D excitatory population.
# - Visualize the response of the population as a function of parameters such as threshold level and gain, using the frequency-current (F-I) curve.
# - Numerically simulate the dynamics of the excitatory population and find the fixed points of the system.
# - Investigate the stability of the fixed points by linearizing the dynamics around them.
#
#
# + [markdown] colab_type="text"
# ---
# # Setup
# + cellView="both" colab={} colab_type="code"
# Imports
import matplotlib.pyplot as plt
import numpy as np
import scipy.optimize as opt # root-finding algorithm
# + cellView="form" colab={} colab_type="code"
# @title Figure Settings
import ipywidgets as widgets # interactive display
# %config InlineBackend.figure_format = 'retina'
plt.style.use("https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/nma.mplstyle")
# + cellView="form" colab={} colab_type="code"
# @title Helper functions
def plot_fI(x, f):
plt.figure(figsize=(6, 4)) # plot the figure
plt.plot(x, f, 'k')
plt.xlabel('x (a.u.)', fontsize=14)
plt.ylabel('F(x)', fontsize=14)
plt.show()
def plot_dr_r(r, drdt, x_fps=None):
plt.figure()
plt.plot(r, drdt, 'k')
plt.plot(r, 0. * r, 'k--')
if x_fps is not None:
plt.plot(x_fps, np.zeros_like(x_fps), "ko", ms=12)
plt.xlabel(r'$r$')
plt.ylabel(r'$\frac{dr}{dt}$', fontsize=20)
plt.ylim(-0.1, 0.1)
def plot_dFdt(x, dFdt):
plt.figure()
plt.plot(x, dFdt, 'r')
plt.xlabel('x (a.u.)', fontsize=14)
plt.ylabel('dF(x)', fontsize=14)
plt.show()
# + [markdown] colab_type="text"
# ---
# # Section 1: Neuronal network dynamics
# + cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 517} colab_type="code" outputId="83f11632-438e-4703-8853-a526b2e2fd35"
# @title Video 1: Dynamic networks
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="p848349hPyw", width=854, height=480, fs=1)
print("Video available at https://youtube.com/watch?v=" + video.id)
video
# + [markdown] colab_type="text"
# ## Section 1.1: Dynamics of a single excitatory population
#
# Individual neurons respond by spiking. When we average the spikes of neurons in a population, we can define the average firing activity of the population. In this model, we are interested in how the population-averaged firing varies as a function of time and network parameters. Mathematically, we can describe the firing rate dynamic as:
#
# \begin{align}
# \tau \frac{dr}{dt} &= -r + F(w\cdot r + I_{\text{ext}}) \quad\qquad (1)
# \end{align}
#
# $r(t)$ represents the average firing rate of the excitatory population at time $t$, $\tau$ controls the timescale of the evolution of the average firing rate, $w$ denotes the strength (synaptic weight) of the recurrent input to the population, $I_{\text{ext}}$ represents the external input, and the transfer function $F(\cdot)$ (which can be related to f-I curve of individual neurons described in the next sections) represents the population activation function in response to all received inputs.
#
# To start building the model, please execute the cell below to initialize the simulation parameters.
# + cellView="form" colab={} colab_type="code"
# @markdown *Execute this cell to set default parameters for a single excitatory population model*
def default_pars_single(**kwargs):
pars = {}
# Excitatory parameters
pars['tau'] = 1. # Timescale of the E population [ms]
pars['a'] = 1.2 # Gain of the E population
pars['theta'] = 2.8 # Threshold of the E population
# Connection strength
pars['w'] = 0. # E to E, we first set it to 0
# External input
pars['I_ext'] = 0.
# simulation parameters
pars['T'] = 20. # Total duration of simulation [ms]
pars['dt'] = .1 # Simulation time step [ms]
pars['r_init'] = 0.2 # Initial value of E
# External parameters if any
pars.update(kwargs)
# Vector of discretized time points [ms]
pars['range_t'] = np.arange(0, pars['T'], pars['dt'])
return pars
# + [markdown] colab_type="text"
# You can now use:
# - `pars = default_pars_single()` to get all the parameters, and then you can execute `print(pars)` to check these parameters.
# - `pars = default_pars_single(T=T_sim, dt=time_step)` to set new simulation time and time step
# - To update an existing parameter dictionary, use `pars['New_para'] = value`
#
# Because `pars` is a dictionary, it can be passed to a function that requires individual parameters as arguments using `my_func(**pars)` syntax.
# + [markdown] colab_type="text"
# ## Section 1.2: F-I curves
# In electrophysiology, a neuron is often characterized by its spike rate output in response to input currents. This is often called the **F-I** curve, denoting the output spike frequency (**F**) in response to different injected currents (**I**). We estimated this for an LIF neuron in yesterday's tutorial.
#
# The transfer function $F(\cdot)$ in Equation $1$ represents the gain of the population as a function of the total input. The gain is often modeled as a sigmoidal function, i.e., more input drive leads to a nonlinear increase in the population firing rate. The output firing rate will eventually saturate for high input values.
#
# A sigmoidal $F(\cdot)$ is parameterized by its gain $a$ and threshold $\theta$.
#
# $$ F(x;a,\theta) = \frac{1}{1+\text{e}^{-a(x-\theta)}} - \frac{1}{1+\text{e}^{a\theta}} \quad(2)$$
#
# The argument $x$ represents the input to the population. Note that the second term is chosen so that $F(0;a,\theta)=0$.
#
# Many other transfer functions (generally monotonic) can be also used. Examples are the rectified linear function $ReLU(x)$ or the hyperbolic tangent $tanh(x)$.
# + [markdown] colab_type="text"
# ### Exercise 1: Implement F-I curve
#
# Let's first investigate the activation functions before simulating the dynamics of the entire population.
#
# In this exercise, you will implement a sigmoidal **F-I** curve or transfer function $F(x)$, with gain $a$ and threshold level $\theta$ as parameters.
# + colab={} colab_type="code"
def F(x, a, theta):
"""
Population activation function.
Args:
x (float): the population input
a (float): the gain of the function
theta (float): the threshold of the function
Returns:
float: the population activation response F(x) for input x
"""
#################################################
## TODO for students: compute f = F(x) ##
# Fill out function and remove
raise NotImplementedError("Student excercise: implement the f-I function")
#################################################
# Define the sigmoidal transfer function f = F(x)
f = ...
return f
pars = default_pars_single() # get default parameters
x = np.arange(0, 10, .1) # set the range of input
# Uncomment below to test your function
# f = F(x, pars['a'], pars['theta'])
# plot_fI(x, f)
# + [markdown] cellView="both" colab={"base_uri": "https://localhost:8080/", "height": 288} colab_type="text" outputId="23aa765f-8d7d-481d-a204-985e5b7ec535"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W2D4_DynamicNetworks/solutions/W2D4_Tutorial1_Solution_45ddc05f.py)
#
# *Example output:*
#
# <img alt='Solution hint' align='left' width=416 height=272 src=https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/tutorials/W2D4_DynamicNetworks/static/W2D4_Tutorial1_Solution_45ddc05f_0.png>
#
#
# + [markdown] colab_type="text"
# ### Interactive Demo: Parameter exploration of F-I curve
# Here's an interactive demo that shows how the F-I curve changes for different values of the gain and threshold parameters. How do the gain and threshold parameters affect the F-I curve?
# + cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 494, "referenced_widgets": ["b61bd80d7d364ad7bf04d01c1cd74910", "245b4a9a358641198e846046ca13cefa", "144fe28586da4c43a999d02b60c7df79", "03250c609c324c2c891bf4a4ee465a26", "85a36101c38c4aa48deef2ab8946d246", "67d602777c7946e0abeb5955b326ce60", "95b37093de5d46c981097831e3419405", "ee845cd049f94bb19ae8222865495d38", "271c3bd6914c4808aa0aa39d6239a80b", "a53e1476f34a4e1aae19c69a62e60dc1"]} colab_type="code" outputId="47e4f04f-041d-4573-8114-ba644cb8e313"
# @title
# @markdown Make sure you execute this cell to enable the widget!
def interactive_plot_FI(a, theta):
"""
Population activation function.
Expecxts:
a : the gain of the function
theta : the threshold of the function
Returns:
plot the F-I curve with give parameters
"""
# set the range of input
x = np.arange(0, 10, .1)
plt.figure()
plt.plot(x, F(x, a, theta), 'k')
plt.xlabel('x (a.u.)', fontsize=14)
plt.ylabel('F(x)', fontsize=14)
plt.show()
_ = widgets.interact(interactive_plot_FI, a=(0.3, 3, 0.3), theta=(2, 4, 0.2))
# + [markdown] colab={} colab_type="text"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W2D4_DynamicNetworks/solutions/W2D4_Tutorial1_Solution_1c0165d7.py)
#
#
# + [markdown] colab_type="text"
# ## Section 1.3: Simulation scheme of E dynamics
#
# Because $F(\cdot)$ is a nonlinear function, the exact solution of Equation $1$ can not be determined via analytical methods. Therefore, numerical methods must be used to find the solution. In practice, the derivative on the left-hand side of Equation $1$ can be approximated using the Euler method on a time-grid of stepsize $\Delta t$:
#
# \begin{align}
# &\frac{dr}{dt} \approx \frac{r[k+1]-r[k]}{\Delta t}
# \end{align}
# where $r[k] = r(k\Delta t)$.
#
# Thus,
#
# $$\Delta r[k] = \frac{\Delta t}{\tau}[-r[k] + F(w\cdot r[k] + I_{\text{ext}}[k];a,\theta)]$$
#
#
# Hence, Equation (1) is updated at each time step by:
#
# $$r[k+1] = r[k] + \Delta r[k]$$
#
# + cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 283} colab_type="code" outputId="39ac5bc8-5083-44c<PASSWORD>"
# @markdown *Execute this cell to enable the single population rate model simulator: `simulate_single`*
def simulate_single(pars):
"""
Simulate an excitatory population of neurons
Args:
pars : Parameter dictionary
Returns:
rE : Activity of excitatory population (array)
Example:
pars = default_pars_single()
r = simulate_single(pars)
"""
# Set parameters
tau, a, theta = pars['tau'], pars['a'], pars['theta']
w = pars['w']
I_ext = pars['I_ext']
r_init = pars['r_init']
dt, range_t = pars['dt'], pars['range_t']
Lt = range_t.size
# Initialize activity
r = np.zeros(Lt)
r[0] = r_init
I_ext = I_ext * np.ones(Lt)
# Update the E activity
for k in range(Lt - 1):
dr = dt / tau * (-r[k] + F(w * r[k] + I_ext[k], a, theta))
r[k+1] = r[k] + dr
return r
help(simulate_single)
# + [markdown] colab_type="text"
# ### Interactive Demo: Parameter Exploration of single population dynamics
#
# Note that $w=0$, as in the default setting, means no recurrent input to the neuron population in Equation (1). Hence, the dynamics are entirely determined by the external input $I_{\text{ext}}$. Explore these dynamics in this interactive demo.
#
# How does $r_{\text{sim}}(t)$ change with different $I_{\text{ext}}$ values? How does it change with different $\tau$ values? Investigate the relationship between $F(I_{\text{ext}}; a, \theta)$ and the steady value of $r(t)$.
#
# Note that, $r_{\rm ana}(t)$ denotes the analytical solution - you will learn how this is computed in the next section.
# + cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 494, "referenced_widgets": ["4701ae22a90d47b49c0102c1d74dea93", "724ab274c39d4e00ac6ea1a63a3e5d29", "d3012a33cb774402874129a452dbc74e", "f276d852e7cc4313a129277277081c91", "<KEY>", "808a66d0557e4994aa6098643e9b1994", "fd5bc6df0676470bb62e6dadf3ef1aae", "c62ce0f30fc34364a816de5affce9108", "<KEY>", "85229a0353be4da285a280bff960af5f"]} colab_type="code" outputId="91dd1a42-b4ed-400d-e6e3-7a5186fec0b4"
# @title
# @markdown Make sure you execute this cell to enable the widget!
# get default parameters
pars = default_pars_single(T=20.)
def Myplot_E_diffI_difftau(I_ext, tau):
# set external input and time constant
pars['I_ext'] = I_ext
pars['tau'] = tau
# simulation
r = simulate_single(pars)
# Analytical Solution
r_ana = (pars['r_init']
+ (F(I_ext, pars['a'], pars['theta'])
- pars['r_init']) * (1. - np.exp(-pars['range_t'] / pars['tau'])))
# plot
plt.figure()
plt.plot(pars['range_t'], r, 'b', label=r'$r_{\mathrm{sim}}$(t)', alpha=0.5,
zorder=1)
plt.plot(pars['range_t'], r_ana, 'b--', lw=5, dashes=(2, 2),
label=r'$r_{\mathrm{ana}}$(t)', zorder=2)
plt.plot(pars['range_t'],
F(I_ext, pars['a'], pars['theta']) * np.ones(pars['range_t'].size),
'k--', label=r'$F(I_{\mathrm{ext}})$')
plt.xlabel('t (ms)', fontsize=16.)
plt.ylabel('Activity r(t)', fontsize=16.)
plt.legend(loc='best', fontsize=14.)
plt.show()
_ = widgets.interact(Myplot_E_diffI_difftau, I_ext=(0.0, 10., 1.),
tau=(1., 5., 0.2))
# + [markdown] colab={} colab_type="text"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W2D4_DynamicNetworks/solutions/W2D4_Tutorial1_Solution_65dee3e7.py)
#
#
# + [markdown] colab_type="text"
# ## Think!
# Above, we have numerically solved a system driven by a positive input. Yet, $r_E(t)$ either decays to zero or reaches a fixed non-zero value.
# - Why doesn't the solution of the system "explode" in a finite time? In other words, what guarantees that $r_E$(t) stays finite?
# - Which parameter would you change in order to increase the maximum value of the response?
# + [markdown] colab={} colab_type="text"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W2D4_DynamicNetworks/solutions/W2D4_Tutorial1_Solution_5a95a98e.py)
#
#
# + [markdown] colab_type="text"
# ---
# # Section 2: Fixed points of the single population system
#
# + cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 517} colab_type="code" outputId="f25d4b12-6499-414e-bee7-88ea1ea9c557"
# @title Video 2: Fixed point
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="Ox3ELd1UFyo", width=854, height=480, fs=1)
print("Video available at https://youtube.com/watch?v=" + video.id)
video
# + [markdown] colab_type="text"
# As you varied the two parameters in the last Interactive Demo, you noticed that, while at first the system output quickly changes, with time, it reaches its maximum/minimum value and does not change anymore. The value eventually reached by the system is called the **steady state** of the system, or the **fixed point**. Essentially, in the steady states the derivative with respect to time of the activity ($r$) is zero, i.e. $\displaystyle \frac{dr}{dt}=0$.
#
# We can find that the steady state of the Equation. (1) by setting $\displaystyle{\frac{dr}{dt}=0}$ and solve for $r$:
#
# $$-r_{\text{steady}} + F(w\cdot r_{\text{steady}} + I_{\text{ext}};a,\theta) = 0, \qquad (3)$$
#
# When it exists, the solution of Equation. (3) defines a **fixed point** of the dynamical system in Equation (1). Note that if $F(x)$ is nonlinear, it is not always possible to find an analytical solution, but the solution can be found via numerical simulations, as we will do later.
#
# From the Interactive Demo, one could also notice that the value of $\tau$ influences how quickly the activity will converge to the steady state from its initial value.
#
# In the specific case of $w=0$, we can also analytically compute the solution of Equation (1) (i.e., the thick blue dashed line) and deduce the role of $\tau$ in determining the convergence to the fixed point:
#
# $$\displaystyle{r(t) = \big{[}F(I_{\text{ext}};a,\theta) -r(t=0)\big{]} (1-\text{e}^{-\frac{t}{\tau}})} + r(t=0)$$ \\
#
# We can now numerically calculate the fixed point with a root finding algorithm.
# + [markdown] colab_type="text"
# ## Exercise 2: Visualization of the fixed points
#
# When it is not possible to find the solution for Equation (3) analytically, a graphical approach can be taken. To that end, it is useful to plot $\displaystyle{\frac{dr}{dt}}$ as a function of $r$. The values of $r$ for which the plotted function crosses zero on the y axis correspond to fixed points.
#
# Here, let us, for example, set $w=5.0$ and $I^{\text{ext}}=0.5$. From Equation (1), you can obtain
#
# $$\frac{dr}{dt} = [-r + F(w\cdot r + I^{\text{ext}})]\,/\,\tau $$
#
# Then, plot the $dr/dt$ as a function of $r$, and check for the presence of fixed points.
# + colab={} colab_type="code"
def compute_drdt(r, I_ext, w, a, theta, tau, **other_pars):
"""Given parameters, compute dr/dt as a function of r.
Args:
r (1D array) : Average firing rate of the excitatory population
I_ext, w, a, theta, tau (numbers): Simulation parameters to use
other_pars : Other simulation parameters are unused by this function
Returns
drdt function for each value of r
"""
#########################################################################
# TODO compute drdt and disable the error
raise NotImplementedError("Finish the compute_drdt function")
#########################################################################
# Calculate drdt
drdt = ...
return drdt
# Define a vector of r values and the simulation parameters
r = np.linspace(0, 1, 1000)
pars = default_pars_single(I_ext=0.5, w=5)
# Uncomment to test your function
# drdt = compute_drdt(r, **pars)
# plot_dr_r(r, drdt)
# + [markdown] colab={"base_uri": "https://localhost:8080/", "height": 485} colab_type="text" outputId="406680a9-25af-446f-df62-6b7ba39b4e12"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W2D4_DynamicNetworks/solutions/W2D4_Tutorial1_Solution_c5280901.py)
#
# *Example output:*
#
# <img alt='Solution hint' align='left' width=558 height=413 src=https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/tutorials/W2D4_DynamicNetworks/static/W2D4_Tutorial1_Solution_c5280901_0.png>
#
#
# + [markdown] colab_type="text"
# ## Exercise 3: Fixed point calculation
#
# We will now find the fixed points numerically. To do so, we need to specif initial values ($r_{\text{guess}}$) for the root-finding algorithm to start from. From the line $\displaystyle{\frac{dr}{dt}}$ plotted above in Exercise 2, initial values can be chosen as a set of values close to where the line crosses zero on the y axis (real fixed point).
#
# The next cell defines three helper functions that we will use:
#
# - `my_fp_single(r_guess, **pars)` uses a root-finding algorithm to locate a fixed point near a given initial value
# - `check_fp_single(x_fp, **pars)`, verifies that the values of $r_{\rm fp}$ for which $\displaystyle{\frac{dr}{dt}} = 0$ are the true fixed points
# - `my_fp_finder(r_guess_vector, **pars)` accepts an array of initial values and finds the same number of fixed points, using the above two functions
# + cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 247} colab_type="code" outputId="079fa0d8-885f-4f21-eca5-6904468e68ee"
# @markdown *Execute this cell to enable the fixed point functions*
def my_fp_single(r_guess, a, theta, w, I_ext, **other_pars):
"""
Calculate the fixed point through drE/dt=0
Args:
r_guess : Initial value used for scipy.optimize function
a, theta, w, I_ext : simulation parameters
Returns:
x_fp : value of fixed point
"""
# define the right hand of E dynamics
def my_WCr(x):
r = x
drdt = (-r + F(w * r + I_ext, a, theta))
y = np.array(drdt)
return y
x0 = np.array(r_guess)
x_fp = opt.root(my_WCr, x0).x.item()
return x_fp
def check_fp_single(x_fp, a, theta, w, I_ext, mytol=1e-4, **other_pars):
"""
Verify |dr/dt| < mytol
Args:
fp : value of fixed point
a, theta, w, I_ext: simulation parameters
mytol : tolerance, default as 10^{-4}
Returns :
Whether it is a correct fixed point: True/False
"""
# calculate Equation(3)
y = x_fp - F(w * x_fp + I_ext, a, theta)
# Here we set tolerance as 10^{-4}
return np.abs(y) < mytol
def my_fp_finder(pars, r_guess_vector, mytol=1e-4):
"""
Calculate the fixed point(s) through drE/dt=0
Args:
pars : Parameter dictionary
r_guess_vector : Initial values used for scipy.optimize function
mytol : tolerance for checking fixed point, default as 10^{-4}
Returns:
x_fps : values of fixed points
"""
x_fps = []
correct_fps = []
for r_guess in r_guess_vector:
x_fp = my_fp_single(r_guess, **pars)
if check_fp_single(x_fp, **pars, mytol=mytol):
x_fps.append(x_fp)
return x_fps
help(my_fp_finder)
# + colab={} colab_type="code"
r = np.linspace(0, 1, 1000)
pars = default_pars_single(I_ext=0.5, w=5)
drdt = compute_drdt(r, **pars)
#############################################################################
# TODO for students:
# Define initial values close to the intersections of drdt and y=0
# (How many initial values? Hint: How many times do the two lines intersect?)
# Calculate the fixed point with these initial values and plot them
#############################################################################
r_guess_vector = [...]
# Uncomment to test your values
# x_fps = my_fp_finder(pars, r_guess_vector)
# plot_dr_r(r, drdt, x_fps)
# + [markdown] colab={"base_uri": "https://localhost:8080/", "height": 485} colab_type="text" outputId="4cf2cddb-0cc0-45be-e577-48e69a6ee6f3"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W2D4_DynamicNetworks/solutions/W2D4_Tutorial1_Solution_0637b6bf.py)
#
# *Example output:*
#
# <img alt='Solution hint' align='left' width=558 height=413 src=https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/tutorials/W2D4_DynamicNetworks/static/W2D4_Tutorial1_Solution_0637b6bf_0.png>
#
#
# + [markdown] colab_type="text"
# ## Interactive Demo: fixed points as a function of recurrent and external inputs.
#
# You can now explore how the previous plot changes when the recurrent coupling $w$ and the external input $I_{\text{ext}}$ take different values. How does the number of fixed points change?
# + cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 494, "referenced_widgets": ["875329f08b434412a4e553ad777cf762", "abb9d829af484d84be1463074b52f5b4", "04f4018e9c2544618446287713fc8876", "b4066b09119c4faca10ee55f9438f63b", "b07bc0d64ad5444cb239d0ac25bc164f", "0e1d4030d1c44fdab228e94b40f5a66d", "1a161bda1e3e405289c0fc5d1f927aa9", "1e1c2f33a69842cb86d1a931e41182ad", "12a33e3c45bf43bf9e64366527bba3f6", "408c7f906c60459fb4375e995b8064c2"]} colab_type="code" outputId="b87c83c5-d95c-40a2-efa5-e657bbec0f2a"
# @title
# @markdown Make sure you execute this cell to enable the widget!
def plot_intersection_single(w, I_ext):
# set your parameters
pars = default_pars_single(w=w, I_ext=I_ext)
# find fixed points
r_init_vector = [0, .4, .9]
x_fps = my_fp_finder(pars, r_init_vector)
# plot
r = np.linspace(0, 1., 1000)
drdt = (-r + F(w * r + I_ext, pars['a'], pars['theta'])) / pars['tau']
plot_dr_r(r, drdt, x_fps)
_ = widgets.interact(plot_intersection_single, w=(1, 7, 0.2),
I_ext=(0, 3, 0.1))
# + [markdown] colab={} colab_type="text"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W2D4_DynamicNetworks/solutions/W2D4_Tutorial1_Solution_20486792.py)
#
#
# + [markdown] colab_type="text"
# ---
# # Summary
#
# In this tutorial, we have investigated the dynamics of a rate-based single population of neurons.
#
# We learned about:
# - The effect of the input parameters and the time constant of the network on the dynamics of the population.
# - How to find the fixed point(s) of the system.
#
# Next, we have two Bonus, but important concepts in dynamical system analysis and simulation. If you have time left, watch the next video and proceed to solve the exercises. You will learn:
#
# - How to determine the stability of a fixed point by linearizing the system.
# - How to add realistic inputs to our model.
# + [markdown] colab_type="text"
# ---
# # Bonus 1: Stability of a fixed point
# + cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 517} colab_type="code" outputId="11b2ff78-d268-404b-8b48-596a70315854"
# @title Video 3: Stability of fixed points
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="KKMlWWU83Jg", width=854, height=480, fs=1)
print("Video available at https://youtube.com/watch?v=" + video.id)
video
# + [markdown] colab_type="text"
# #### Initial values and trajectories
#
# Here, let us first set $w=5.0$ and $I_{\text{ext}}=0.5$, and investigate the dynamics of $r(t)$ starting with different initial values $r(0) \equiv r_{\text{init}}$. We will plot the trajectories of $r(t)$ with $r_{\text{init}} = 0.0, 0.1, 0.2,..., 0.9$.
# + cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 357} colab_type="code" outputId="e69ca3c9-9981-4fd4-e9bf-73d62cf764de"
# @markdown Execute this cell to see the trajectories!
pars = default_pars_single()
pars['w'] = 5.0
pars['I_ext'] = 0.5
plt.figure(figsize=(8, 5))
for ie in range(10):
pars['r_init'] = 0.1 * ie # set the initial value
r = simulate_single(pars) # run the simulation
# plot the activity with given initial
plt.plot(pars['range_t'], r, 'b', alpha=0.1 + 0.1 * ie,
label=r'r$_{\mathrm{init}}$=%.1f' % (0.1 * ie))
plt.xlabel('t (ms)')
plt.title('Two steady states?')
plt.ylabel(r'$r$(t)')
plt.legend(loc=[1.01, -0.06], fontsize=14)
plt.show()
# + [markdown] colab_type="text"
# ## Interactive Demo: dynamics as a function of the initial value
#
# Let's now set $r_{\rm init}$ to a value of your choice in this demo. How does the solution change? What do you observe?
# + cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 462, "referenced_widgets": ["a861ef888bd041ad8c47551e665cc9ba", "8d37cd431d964d96a50582482ff028d4", "ff23d44495b44d729861ba6cf52c0bca", "ba13f61549f24b14ba70e387aa1a9485", "f9e0df8ceaf647949c1c8975651dfda2", "cbec43cfc0064abf805c1d52bb395caa", "ed7948064a3b415d8429da73484125f7"]} colab_type="code" outputId="2962d6fb-cad4-45d2-8b39-910437af7b8b"
# @title
# @markdown Make sure you execute this cell to enable the widget!
pars = default_pars_single(w=5.0, I_ext=0.5)
def plot_single_diffEinit(r_init):
pars['r_init'] = r_init
r = simulate_single(pars)
plt.figure()
plt.plot(pars['range_t'], r, 'b', zorder=1)
plt.plot(0, r[0], 'bo', alpha=0.7, zorder=2)
plt.xlabel('t (ms)', fontsize=16)
plt.ylabel(r'$r(t)$', fontsize=16)
plt.ylim(0, 1.0)
plt.show()
_ = widgets.interact(plot_single_diffEinit, r_init=(0, 1, 0.02))
# + [markdown] colab={} colab_type="text"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W2D4_DynamicNetworks/solutions/W2D4_Tutorial1_Solution_4d2de6a0.py)
#
#
# + [markdown] colab_type="text"
# ### Stability analysis via linearization of the dynamics
#
# Just like Equation $1$ in the case ($w=0$) discussed above, a generic linear system
# $$\frac{dx}{dt} = \lambda (x - b),$$
# has a fixed point for $x=b$. The analytical solution of such a system can be found to be:
# $$x(t) = b + \big{(} x(0) - b \big{)} \text{e}^{\lambda t}.$$
# Now consider a small perturbation of the activity around the fixed point: $x(0) = b+ \epsilon$, where $|\epsilon| \ll 1$. Will the perturbation $\epsilon(t)$ grow with time or will it decay to the fixed point? The evolution of the perturbation with time can be written, using the analytical solution for $x(t)$, as:
# $$\epsilon (t) = x(t) - b = \epsilon \text{e}^{\lambda t}$$
#
# - if $\lambda < 0$, $\epsilon(t)$ decays to zero, $x(t)$ will still converge to $b$ and the fixed point is "**stable**".
#
# - if $\lambda > 0$, $\epsilon(t)$ grows with time, $x(t)$ will leave the fixed point $b$ exponentially, and the fixed point is, therefore, "**unstable**" .
# + [markdown] colab_type="text"
# ### Compute the stability of Equation $1$
#
# Similar to what we did in the linear system above, in order to determine the stability of a fixed point $r^{*}$ of the excitatory population dynamics, we perturb Equation (1) around $r^{*}$ by $\epsilon$, i.e. $r = r^{*} + \epsilon$. We can plug in Equation (1) and obtain the equation determining the time evolution of the perturbation $\epsilon(t)$:
#
# \begin{align}
# \tau \frac{d\epsilon}{dt} \approx -\epsilon + w F'(w\cdot r^{*} + I_{\text{ext}};a,\theta) \epsilon
# \end{align}
#
# where $F'(\cdot)$ is the derivative of the transfer function $F(\cdot)$. We can rewrite the above equation as:
#
# \begin{align}
# \frac{d\epsilon}{dt} \approx \frac{\epsilon}{\tau }[-1 + w F'(w\cdot r^* + I_{\text{ext}};a,\theta)]
# \end{align}
#
# That is, as in the linear system above, the value of
#
# $$\lambda = [-1+ wF'(w\cdot r^* + I_{\text{ext}};a,\theta)]/\tau \qquad (4)$$
#
# determines whether the perturbation will grow or decay to zero, i.e., $\lambda$ defines the stability of the fixed point. This value is called the **eigenvalue** of the dynamical system.
# + [markdown] colab_type="text"
# ## Exercise 4: Compute $dF$
#
# The derivative of the sigmoid transfer function is:
# \begin{align}
# \frac{dF}{dx} & = \frac{d}{dx} (1+\exp\{-a(x-\theta)\})^{-1} \\
# & = a\exp\{-a(x-\theta)\} (1+\exp\{-a(x-\theta)\})^{-2}. \qquad (5)
# \end{align}
#
# Let's now find the expression for the derivative $\displaystyle{\frac{dF}{dx}}$ in the following cell and plot it.
# + colab={} colab_type="code"
def dF(x, a, theta):
"""
Population activation function.
Args:
x : the population input
a : the gain of the function
theta : the threshold of the function
Returns:
dFdx : the population activation response F(x) for input x
"""
###########################################################################
# TODO for students: compute dFdx ##
raise NotImplementedError("Student excercise: compute the deravitive of F")
###########################################################################
# Calculate the population activation
dFdx = ...
return dFdx
pars = default_pars_single() # get default parameters
x = np.arange(0, 10, .1) # set the range of input
# Uncomment below to test your function
# df = dF(x, pars['a'], pars['theta'])
# plot_dFdt(x, df)
# + [markdown] colab={"base_uri": "https://localhost:8080/", "height": 433} colab_type="text" outputId="540be28c-ffc2-44d3-fbdb-6f27b3305405"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W2D4_DynamicNetworks/solutions/W2D4_Tutorial1_Solution_ce2e3bc5.py)
#
# *Example output:*
#
# <img alt='Solution hint' align='left' width=560 height=416 src=https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/tutorials/W2D4_DynamicNetworks/static/W2D4_Tutorial1_Solution_ce2e3bc5_0.png>
#
#
# + [markdown] colab_type="text"
# ## Exercise 5: Compute eigenvalues
#
# As discussed above, for the case with $w=5.0$ and $I_{\text{ext}}=0.5$, the system displays **three** fixed points. However, when we simulated the dynamics and varied the initial conditions $r_{\rm init}$, we could only obtain **two** steady states. In this exercise, we will now check the stability of each of the three fixed points by calculating the corresponding eigenvalues with the function `eig_single`. Check the sign of each eigenvalue (i.e., stability of each fixed point). How many of the fixed points are stable?
#
# Note that the expression of the eigenvalue at fixed point $r^*$
# $$\lambda = [-1+ wF'(w\cdot r^* + I_{\text{ext}};a,\theta)]/\tau$$
# + colab={} colab_type="code"
def eig_single(fp, tau, a, theta, w, I_ext, **other_pars):
"""
Args:
fp : fixed point r_fp
tau, a, theta, w, I_ext : Simulation parameters
Returns:
eig : eigevalue of the linearized system
"""
#####################################################################
## TODO for students: compute eigenvalue and disable the error
raise NotImplementedError("Student excercise: compute the eigenvalue")
######################################################################
# Compute the eigenvalue
eig = ...
return eig
# Find the eigenvalues for all fixed points of Exercise 2
pars = default_pars_single(w=5, I_ext=.5)
r_guess_vector = [0, .4, .9]
x_fp = my_fp_finder(pars, r_guess_vector)
# Uncomment below lines after completing the eig_single function.
# for fp in x_fp:
# eig_fp = eig_single(fp, **pars)
# print(f'Fixed point1 at {fp:.3f} with Eigenvalue={eig_fp:.3f}')
# + [markdown] colab_type="text"
# **SAMPLE OUTPUT**
#
# ```
# Fixed point1 at 0.042 with Eigenvalue=-0.583
# Fixed point2 at 0.447 with Eigenvalue=0.498
# Fixed point3 at 0.900 with Eigenvalue=-0.626
# ```
# + [markdown] colab={"base_uri": "https://localhost:8080/", "height": 70} colab_type="text" outputId="70e87372-6a77-4376-93d5-a2de111ec1ad"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W2D4_DynamicNetworks/solutions/W2D4_Tutorial1_Solution_e285f60d.py)
#
#
# + [markdown] colab_type="text"
# ## Think!
# Throughout the tutorial, we have assumed $w> 0 $, i.e., we considered a single population of **excitatory** neurons. What do you think will be the behavior of a population of inhibitory neurons, i.e., where $w> 0$ is replaced by $w< 0$?
# + [markdown] colab={} colab_type="text"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W2D4_DynamicNetworks/solutions/W2D4_Tutorial1_Solution_579bc9c9.py)
#
#
# + [markdown] colab_type="text"
# ---
# # Bonus 2: Noisy input drives the transition between two stable states
#
#
# + [markdown] colab_type="text"
# ## Ornstein-Uhlenbeck (OU) process
#
# As discussed in several previous tutorials, the OU process is usually used to generate a noisy input into the neuron. The OU input $\eta(t)$ follows:
#
# $$\tau_\eta \frac{d}{dt}\eta(t) = -\eta (t) + \sigma_\eta\sqrt{2\tau_\eta}\xi(t)$$
#
# Execute the following function `my_OU(pars, sig, myseed=False)` to generate an OU process.
# + cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 286} colab_type="code" outputId="5fcb932d-ba75-4b9f-8bb4-030363a53dd7"
# @title OU process `my_OU(pars, sig, myseed=False)`
# @markdown Make sure you execute this cell to visualize the noise!
def my_OU(pars, sig, myseed=False):
"""
A functions that generates Ornstein-Uhlenback process
Args:
pars : parameter dictionary
sig : noise amplitute
myseed : random seed. int or boolean
Returns:
I : Ornstein-Uhlenbeck input current
"""
# Retrieve simulation parameters
dt, range_t = pars['dt'], pars['range_t']
Lt = range_t.size
tau_ou = pars['tau_ou'] # [ms]
# set random seed
if myseed:
np.random.seed(seed=myseed)
else:
np.random.seed()
# Initialize
noise = np.random.randn(Lt)
I_ou = np.zeros(Lt)
I_ou[0] = noise[0] * sig
# generate OU
for it in range(Lt - 1):
I_ou[it + 1] = (I_ou[it]
+ dt / tau_ou * (0. - I_ou[it])
+ np.sqrt(2 * dt / tau_ou) * sig * noise[it + 1])
return I_ou
pars = default_pars_single(T=100)
pars['tau_ou'] = 1. # [ms]
sig_ou = 0.1
I_ou = my_OU(pars, sig=sig_ou, myseed=2020)
plt.figure(figsize=(10, 4))
plt.plot(pars['range_t'], I_ou, 'r')
plt.xlabel('t (ms)')
plt.ylabel(r'$I_{\mathrm{OU}}$')
plt.show()
# + [markdown] colab_type="text"
# ## Example: Up-Down transition
#
# In the presence of two or more fixed points, noisy inputs can drive a transition between the fixed points! Here, we stimulate an E population for 1,000 ms applying OU inputs.
# + cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 286} colab_type="code" outputId="e327a1fd-2d8a-41dc-a2d9-aef33a54d220"
# @title Simulation of an E population with OU inputs
# @markdown Make sure you execute this cell to spot the Up-Down states!
pars = default_pars_single(T=1000)
pars['w'] = 5.0
sig_ou = 0.7
pars['tau_ou'] = 1. # [ms]
pars['I_ext'] = 0.56 + my_OU(pars, sig=sig_ou, myseed=2020)
r = simulate_single(pars)
plt.figure(figsize=(10, 4))
plt.plot(pars['range_t'], r, 'b', alpha=0.8)
plt.xlabel('t (ms)')
plt.ylabel(r'$r(t)$')
plt.show()
| tutorials/W2D4_DynamicNetworks/student/W2D4_Tutorial1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import json
df_queue = pd.read_json("./single_request_without_queue.json", lines=True)
def filter_df(x):
return x['metric'] == 'completion_time' and 'tags' in x['data']
df_q_completion = df_queue[df_queue.apply(filter_df, axis=1)]
df_q_completion['start'] = df_q_completion.apply(lambda x: x['data']['tags']['start'], axis=1)
df_q_completion['start_success'] = df_q_completion.apply(lambda x: x['data']['tags']['startSuccess'], axis=1)
df_q_completion['end'] = df_q_completion.apply(lambda x: x['data']['tags']['end'], axis=1)
df_q_completion['vu'] = df_q_completion.apply(lambda x: x['data']['tags']['vu'], axis=1)
df_q_completion
min_start = df_q_completion['start'].min()
min_start
df_q_completion['start'] = df_q_completion['start'].apply(lambda x: int(x) - int(min_start))
df_q_completion['start_success'] = df_q_completion['start_success'].apply(lambda x: int(x) - int(min_start))
df_q_completion['end'] = df_q_completion['end'].apply(lambda x: int(x) - int(min_start))
df_q_completion
df_sampled = df_q_completion[:200] #.sample(n=100)
df_sampled_sorted = df_sampled.sort_values(by='start', ascending=False)
# +
import matplotlib.pyplot as plt
import numpy as np
target_bars_start = df_sampled_sorted['start']
target_bars_start_success = df_sampled_sorted['start_success']
target_bars_end = df_sampled_sorted['end']
N = len(target_bars_end)
ind = np.arange(N)
plt.subplots(figsize=(10, 20))
plt.barh(ind, target_bars_end, 0.5, color='orange')
plt.barh(ind, target_bars_start_success, 0.5)
plt.barh(ind, target_bars_start, 0.5, color='white')
# -
| analysis/Single Request Without Queue.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.3 32-bit
# language: python
# name: python38332bitabebdcd8e2a1488c90932336e3af1b49
# ---
# # Starbucks Store Information Scraping
from selenium import webdriver
from bs4 import BeautifulSoup as BS
import pandas as pd
# webdriver로 browser 준비
browser = webdriver.Chrome()
browser.get('https://www.starbucks.co.kr/store/store_map.do?disp=locale')
# ### CSS selector로 위치를 찾아서 클릭하기
# browser에서 모든 정보가 로딩된 뒤에 click해야 제대로 작동하기 때문에 sleep을 설정하는 것이 좋다. 그러나 Jupyter notebook에서는 cell이 하나씩 순차적으로 실행되니까 크게 상관이 없긴 하다. <br>
# 또는 전체를 run하는 경우, 너무 빨리 지나가버리기 때문에 sleep을 설정하면 진행을 천천히 확인할 수 있다.
# +
import time
time.sleep(5)
browser.find_element_by_css_selector('ul.sido_arae_box > li > a[data-sidocd="01"]').click() # '서울' 클릭
# -
time.sleep(5)
browser.find_element_by_css_selector('ul.gugun_arae_box > li > a[href*="0"][data-guguncd=""]').click() # '전체' 클릭
# 위 CSS selector 중 `[href*="0"]` 부분은 selector를 특정하는데 도움이 되지 않아서 필요 없음. 단지 tag attribute를 여러개 연이어 사용할 수 있다는 것을 보여주기 위해서 추가함
# ### BeautifulSoup으로 매장 리스트를 읽어오기
page = browser.page_source # 현재 webdriver에 열려있는 페이지의 내용을 불러옴
soup = BS(page, 'html.parser')
tags = soup.select('li.quickResultLstCon')
len(tags), tags[0] # 총 554개 매장
# +
# 매장이름, 주소, 위도, 경도, 매장타입을 스크래핑
starbucks = []
for tag in tags:
name = tag['data-name']
add = tag.select_one('p').text
lat = tag['data-lat']
long = tag['data-long']
store = tag.select_one('i').text
starbucks.append([name, add, lat, long, store])
starbucks[:5]
# -
# ### DataFrame으로 만들기
pd.DataFrame(starbucks, columns=['Store', 'Address', 'Latitude', 'Longitude', 'Type']).head()
# ---
# ### 번외) WebElement object의 html source 확인하는 법
# Selenium으로 element 찾기를 마친 뒤 BeautifulSoup으로 parsing 할 수도 있다!
# outerHTML: 지정한 태그(li) 포함해서 가져옴
# innerHTML: 지정한 태그 제외 안쪽만 가져옴
elmt = browser.find_element_by_css_selector('li.quickResultLstCon').get_attribute('outerHTML')
elmt, type(elmt) # str
print(BS(elmt, 'html.parser').prettify())
# ### Selenium만 사용해서 Starbucks 매장정보 가져오기
# 매장정보 모두 가져오기
tags = browser.find_elements_by_css_selector('li.quickResultLstCon')
tags[0].get_attribute('outerHTML') # 첫번째 WebElement의 html source 확인
# tags[0].find_element_by_css_selector('p').text # html은 제대로 가져오는데 왜 text는 안 불러와지지????
# +
# 속도가 너무 느리다...
stores = []
for tag in tags:
name = tag.get_attribute('data-name') # tag_name, css_selector로는 text가 안 불러와진다?????????????????????????
add = tag.find_element_by_xpath('//*[@id="mCSB_3_container"]/ul/li[5]/p').text.replace('\n', ' ')
lat = tag.get_attribute('data-lat')
long = tag.get_attribute('data-long')
store = tag.find_element_by_xpath('//*[@id="mCSB_3_container"]/ul/li[5]/i').text
stores.append([name, add, lat, long, store])
stores[:5]
# -
# Dataframe으로 만들기
pd.DataFrame(stores, columns=['Store', 'Address', 'Latitude', 'Longitude', 'Type']).head()
| Day28_03_starbucks_scraping_final.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import numpy as np
from RadSPy import simulator_1D
import matplotlib.pyplot as plt
# # 1). Create simulator object
testModel = simulator_1D()
# # 2). Set the layered model parameters
# +
# Layer thicknesses
thicknesses = np.array([1.5,1.5,25,5,50,150])
# dielectric constant values
eps = np.array([3.15,4.3,3.15,4.3,3.15,4.0,8.5])
# Loss tangent or conductivity values
lossOrCond = np.array([0.002, 0.02, 0.002, 0.02, 0.002, 0.02,0.02])
# 1 if using conductivity, 0 if using loss tangent
useConductivity = 0
# Spacecraft altitude in meters
H = 300*1000
testModel.setModel(H,thicknesses,eps,lossOrCond,useConductivity)
# -
# # 3). Set the radar pulse
# There are two ways to set the source pulse:
# (1) load the pulse from a csv file
# (2) set the pulse with an array.
# +
# Load pulse from csv file.
timeTest, pulseTest = testModel.loadPulse('Sharad_Ideal_sourcePulse.csv')
# Set pulse
testModel.setPulse(pulseTest,timeTest)
# -
# # 4). Set the radar matched filter
# There are two ways to set the matched filter for range compression:
# (1) load the filter from a csv file
# (2) set the filter with an array.
# +
tmfTest,fmfTest,mfTest = testModel.loadMatchFilter('Sharad_Ideal_matchedFilter.csv')
testModel.setMatchFilter(mfTest,fmfTest,tmfTest)
# -
# # 5). Set the windowing parameters
# +
# The SHARAD instument has a swept pulse from 15 MHz to 25 MHz. We will window over those frequencies
fmin = 15e6 # Low end of filter in Hz
fmax = 25e6 # High end of filter in Hz
fnum = 6 # Use zero for no windowing, 6 is a Hann window
testModel.setWindowParam(fnum,fmin,fmax)
# -
# # 6). Run simulation and plot result
# +
testModel.plotResult=True # This line sets the simulator to produce a plot when runSim() is called
testModel.plotTime=3.3 # in micro-seconds, this controls the time axis of the model plot
plt.rcParams.update({'font.size': 20}) # This changes the font size of the tick labels
testModel.runSim() # This runs the model.
# -
| demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Redes Convolucionales: Ejemplo clasificación de perros y gatos
#
# En este notebook utilizaremos datos de la [competición de Kaggle Dogs vs. Cats](https://www.kaggle.com/c/dogs-vs-cats/overview), a partir de los cuales intentaremos crearnos un sistema capaz de diferenciar entre perros y gatos mediante redes neuronales.
#
# Podrás acceder a la carpeta con las imágenes directamente desde el siguiente enlace a Google Drive: https://drive.google.com/drive/u/0/folders/1oVWZ2-2Whwiw9gFvR-aqQ4Y5sbDzYUtd
# + [markdown] _uuid="fe76d1d1ded592430e7548feacfa38dc42f085d9"
# # Importamos librerías
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from keras.preprocessing.image import ImageDataGenerator, load_img
from sklearn.model_selection import train_test_split
import matplotlib.pyplot as plt
import random
import os
# -
# # Definimos constantes
#
# Vamos a definir una serie de variables que se mantendrán constantes a lo largo de la ejecución del notebook, como tamaño de las imágenes que vamos a utilizar para analizarlas o el número de epochs que vamos a utilizar:
# +
IMAGE_WIDTH=64
IMAGE_HEIGHT=64
IMAGE_CHANNELS=3
IMAGE_SIZE=(IMAGE_WIDTH, IMAGE_HEIGHT, IMAGE_CHANNELS)
BATCH_SIZE = 64
EPOCHS = 5
# DATA_PATH = "dogs-vs-cats/mini_data/"
DATA_PATH = "dogs-vs-cats/train/"
# -
DATA_PATH
len(os.listdir(DATA_PATH))
# + [markdown] _uuid="7335a579cc0268fba5d34d6f7558f33c187eedb3"
# # Preparación de los datos
# 1. Descárgate el dataset de train de [la competición de Kaggle](https://www.kaggle.com/c/dogs-vs-cats/overview) o del [enlace](https://drive.google.com/drive/u/0/folders/1oVWZ2-2Whwiw9gFvR-aqQ4Y5sbDzYUtd) a Google Drive (si no te quieres registrar y todo eso).
# 2. Descomprime el dataset y guárdalo en la ruta actual, de modo que te queden las carpetas:
#
# ```
# 2-Redes Convolucionales:
# * dogs-vs-cats:
# * train
# * fotos
# * test
# * fotos
#
# ```
# -
DATA_PATH
# + _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a"
filenames = os.listdir(DATA_PATH)
categories = []
for filename in filenames:
category = filename.split('.')[0]
if category == 'dog':
categories.append(1)
else:
categories.append(0)
df = pd.DataFrame({
'filenames': filenames,
'category': categories
})
# + _uuid="915bb9ba7063ab4d5c07c542419ae119003a5f98"
df
# + [markdown] _uuid="a999484fc35b73373fafe2253ae9db7ff46fdb90"
# ## Representación de la distribución de las categorías
# + _uuid="fa26f0bc7a6d835a24989790b20f3c6f32946f45"
df['category'].value_counts().plot.bar();
# + [markdown] _uuid="400a293df3c8499059d9175f3915187074efd971"
# ## Representamos las imágenes
# +
# # !pip install opencv-python
# +
from skimage.io import imread
import cv2
sample = random.choice(filenames)
image = imread(DATA_PATH + sample)
print(image.shape)
print(np.max(image))
plt.imshow(image);
# -
# Una imagen no es mas que un array de HxWxC píxeles, siendo H(Height) y W(Width) las dimensiones de resolución de la imagen, y C el número de canales. Habrá tres valores por píxel.
# + jupyter={"outputs_hidden": true}
image
# -
# # Redimensionando imagen
#
# Cargar todas las imágenes a la vez es un problema, ya que son un total de 25000 (unos 500MB la carpeta de train). Este proceso require mucha memoria, por lo que tendremos que aplicarle un resize a cada imagen para bajarlas de resolución. Esto también nos sirve para solventar el problema de tener imágenes con distintas resoluciones.
# +
plt.figure(figsize=(12, 12))
sample = random.choice(filenames)
image = imread(DATA_PATH + sample)
imagesmall = cv2.resize(image, (IMAGE_WIDTH, IMAGE_HEIGHT))
print("Tamaño imagen original:", image.shape)
print("Tamaño imagen reshape:", imagesmall.shape)
print("Maximo valor por pixel:", np.max(imagesmall))
# Original image
plt.subplot(1, 2, 1)
plt.imshow(image)
# Resized image
plt.subplot(1, 2, 2)
plt.imshow(imagesmall);
# -
# # Color
# Podríamos cargar las imágenes como blanco y negro, de esta forma se reduciría el espacio de features considerablemente al contar con un único canal
sample = random.choice(filenames)
image = cv2.imread(filename=DATA_PATH + sample,
flags=cv2.IMREAD_GRAYSCALE)
print(image.shape)
plt.imshow(image, cmap='gray');
# # Cargamos los datos
#
# Llega el momento de cargar los datos. Ya no es tan sencillo como cuando teníamos datasets en CSVs puesto que ahora hay que cargar miles de archivos en memoria en este notebook. Para ello necesitaremos un programa iterativo que vaya recorriendo los archivos de la carpeta, cargarlos como array de numpy y almacenarlos en un objeto.
# +
from sklearn.model_selection import train_test_split
def read_data(path):
X = []
Y = []
for file in os.listdir(path):
image = imread(path + file)
smallimage = cv2.resize(image, (IMAGE_WIDTH, IMAGE_HEIGHT))
X.append(smallimage)
category = file.split('.')[0]
if category == 'dog':
Y.append(1)
else:
Y.append(0)
return np.array(X), np.array(Y)
X_train_full, y_train_full = read_data(DATA_PATH)
X_train, X_test, y_train, y_test = train_test_split(X_train_full, y_train_full, test_size=0.2, random_state=42)
print(X_train.shape)
print(X_test.shape)
# -
DATA_PATH
print(X_train[1].shape)
plt.imshow(X_train[1]);
# + jupyter={"outputs_hidden": true}
X_train[0]
# -
# # Normalizando los datos
#
# Normalizar los datos hará que entrene mucho mejor la red, al estar todos los pixeles en la misma escala.
print("Min:", np.min(X_train))
print("Max:", np.max(X_train))
# +
X_train = X_train / 255.0
X_test = X_test / 255.0
print("Min:", np.min(X_train))
print("Max:", np.max(X_train))
# -
# # Guardando los datos
#
# Podemos guardar los arrays de numpy en un archivo `.npz`, de tal manera que luego sea más rápido importarlo
np.savez('data.npz',
X_train=X_train, y_train=y_train, X_test=X_test, y_test=y_test)
# Para cargar:
# +
data = np.load('data.npz')
X_train = data['X_train']
X_test = data['X_test']
y_train = data['y_train']
y_test = data['y_test']
# + [markdown] _uuid="b244e6b7715a04fc6df92dd6dfa3d35c473ca600"
# # Modelo
#
# <img src="https://i.imgur.com/ebkMGGu.jpg" width="100%"/>
# -
# * **Conv Layer**: extraerá diferentes features de las imagenes
# * **Pooling Layer**: Reduce las dimensiones de las imágenes tras una capa convolucional
# * **Fully Connected Layer**: Tras las capas convolucionales, aplanamos las features y las introducimos como entrada de una red neuronal normal.
# * **Output Layer**: Las predicciones de la red
#
# Para el loss y la métrica, se puede usar un binary_crossentropy, al ser un target binario:
# +
from tensorflow import keras
layers = [
# 64 filtros
# (3,3) kernel de la convolucion
keras.layers.Conv2D(64,
(3,3),
activation='relu',
input_shape=IMAGE_SIZE),
keras.layers.MaxPooling2D(pool_size=(2,2)),
keras.layers.Conv2D(64,
(3,3),
activation='relu'),
keras.layers.MaxPooling2D(pool_size=(2,2)),
# Fully connected layer
keras.layers.Flatten(),
keras.layers.Dense(64, activation='relu'),
keras.layers.Dense(1, activation='sigmoid')
]
model = keras.Sequential(layers)
# loss para n clases (cambiar output) seria sparse_categorical_crossentropy
model.compile(optimizer='adam',
loss='binary_crossentropy',
metrics=['accuracy'])
# -
model.summary()
64*64*64
# + [markdown] _uuid="bd496f6c65888a969be3703135b0b03a8a1190c8"
# # Callbacks
# #### Early Stopping
# + _uuid="9aa032f0f6da539d23918890d2d131cc3aac8c7a"
from keras.callbacks import EarlyStopping
earlystop = EarlyStopping(patience=10)
# -
# # Entranando el modelo
model.fit(X_train,
y_train,
epochs=20,
batch_size=BATCH_SIZE,
#callbacks=[earlystop],
validation_split=0.2)
# Guardamos el histórico:
history = model.history.history
# # Evalúa los resultados
#
# Probemos los datos en el conjunto de test:
results = model.evaluate(X_test, y_test)
print("test loss, test acc", results)
plt.imshow(X_test[2000]);
y_test[2000]
X_test[2000].shape
predictions = model.predict(X_test[2000:2001]).round(2)
print("prections shape:", predictions.shape)
predictions
# + [markdown] _uuid="1b76c0a9040bc0babf0a453e567e41e22f8a1e0e"
# # Visualizando el rendimiento
#
# Utilizaremos los datos históricos guardados previamente:
# + jupyter={"outputs_hidden": true}
history
# + _uuid="79055f2dc3e2abb47bea758e0464c86ca42ab431"
fig, (ax1, ax2) = plt.subplots(2, 1, figsize=(12, 12))
ax1.plot(history['loss'], color='b', label="Training loss")
ax1.plot(history['val_loss'], color='r', label="validation loss")
ax1.legend(loc='best', shadow=True)
ax1.set_xticks(np.arange(1, EPOCHS, 1))
ax1.set_yticks(np.arange(0, 1, 0.1))
ax2.plot(history['accuracy'], color='b', label="Training accuracy")
ax2.plot(history['val_accuracy'], color='r',label="Validation accuracy")
ax2.set_xticks(np.arange(1, EPOCHS, 1))
legend = plt.legend(loc='best', shadow=True)
plt.tight_layout()
plt.show()
# -
| Bloque 3 - Machine Learning/05_Deep_Learning/2-Redes Convolucionales/01_Dogs&Cats.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Pygame
#
# Pygame is a wrapper for [SDL (Simple Directmedia Layer)](http://www.libsdl.org/), *a cross-platform development library designed to provide low level access to audio, keyboard, mouse, joystick, and graphics hardware via OpenGL and Direct3D*.
| multimedia/Pygame/basics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1 style="font-size:42px; text-align:center; margin-bottom:30px;"><span style="color:SteelBlue">Intro:</span> Python Crash Course</h1>
#
# <br><hr id="toc">
#
# ### Table of Contents
#
# * [Lesson 1: Jupyter Notebook Basics](#l0)
# * [Lesson 2: Python Basics](#l1)
# * [Lesson 3: Data Structures](#l2)
# * [Lesson 4: Flow and Functions](#l3)
# * [Lesson 5: Pandas](#l5)
#
# <br><hr>
# ### Jupyter Notebook is an open-source web application that allows you to create and share documents that contain:
# - live code
# - equations
# - visualizations
# - narrative text
#
# ### Uses include: data cleaning and transformation, numerical simulation, statistical modeling, data visualization, machine learning, and much more.
# <br id="l0">
#
# # Lesson 1: Jupyter Notebook Basics
# When running **multiple lines of code** in Jupyter Notebooks, only the last result is shown.
3 * 5 # Not shown
16 / 4 # Shown
# But you can display each line by explicitly printing them with the <code style="color:steelblue">print()</code> function.
print( 3 * 5 ) # Shown
print( 16 / 4 ) # Shown
# Hey, did you see that <code style="color:dimgray; font-weight:bold">Gray</code> text in the code cell?
#
# That's called a **comment**.
# * Comments add extra information, and they are not executed
# * In Python, comments start with the pound sign (a.k.a. hashtag):
#
# <pre style="color:dimgray"># This is a comment</pre>
# +
# print( 3 * 5 ) <-- This code does not get run
# -
# By the way, if you <code style="color:steelblue">print()</code> multiple objects, separated by commas, it will **concatenate** those objects into a single string (more on strings later).
# Print concatenation
print( 'Testing', 1, 2, 3 )
# Great, now let's start our tour of the basics of the Python programming language.
#
# By the way, you'll practice each of these topics throughout the course, so don't worry too much about remembering every little detail on your first pass.
# <hr style="border-color:royalblue;background-color:royalblue;height:1px;">
#
# <div style="text-align:center; margin: 40px 0 40px 0;">
# [**Back to Contents**](#toc)
# </div>
# <br id="l1">
#
# # Lesson 2: Python Basics
# A Python library is a collection of functions and methods that allows you to perform lots of actions without writing your own code.
#
# In general, it's good practice to keep all of your library imports at the top of your notebook or program.
#
# Lets import the math library to calculate the amount of water that can be carried in a container with a radius of 5cm and a height of 17cm.
#
# * The formula for the volume of a cylinder is $V = \pi r^2 h$
import math
total_volume = math.pi*math.pow(5, 2)*17
total_volume
# Do you have enough space to accomodate 2000 cm$^3$? Print <code style="color:steelblue">True</code> if you have enough space or <code style="color:steelblue">False</code> if you do not.**
# * Use the **greater-than-or-equal-to** operator.
# Do you have space for at least 2000 cm^3
if total_volume >= 2000:
print(True)
else:
print(False)
# Repeat the calculations from <span style="color:RoyalBlue">Above</span>, but this time use variables with descriptive names.
#
# * You have **3** empty cylinder-shaped bottles.
# * Each bottle has a height of **16** cm and a radius of **4** cm.
# * Each bottle can be completely filled (ignore the thickness of the bottle).
# * The formula for the volume of a cylinder is $V = \pi r^2 h$
#
# <br>
# **Start by setting variables with descriptive names for bottle dimensions, the number of bottles, and $\pi$.**
# Set variables
bottles = 3
bottle_height = 16
bottle_radius = 4
# <br>
# **Next, calculate the intermediary step of a single bottle's volume.**
# * Set it to a new variable.
# Volume of one bottle (in cm^3)
bottle_volume = math.pi*math.pow(bottle_radius, 2)*bottle_height
# Print volume of a single bottle
print(bottle_volume)
# <br>
# **Finally, calculate total volume.**
# * Set it to a new variable.
# Total volume
total_volume = bottle_volume * bottles
# Print total volume
print(total_volume)
# <hr style="border-color:royalblue;background-color:royalblue;height:1px;">
#
# <div style="text-align:center; margin: 40px 0 40px 0;">
# [**Back to Contents**](#toc)
# </div>
# <br id="l2">
# # Lesson 3: Data Structures
# ***In the previous lesson...***
#
# > *In the previous lesson, you learned about importing libraries, declaring variables and conditional statements.*
#
# > *You also learned how to use libraries by making calculations.*
#
#
# In this lesson we'll learn about importing files along with data structures while planning a trip to breweries in California.
#
# Before we start, let's import lists of breweries by locations.
# * These are stored in text files that we have provided for you.
# * Python has a variety of **input/output** methods. We won't cover them here, but you can learn more about them in the [documentation](https://docs.python.org/2/tutorial/inputoutput.html).
#
# <br>
# **First, run this code.**
# +
# Read lists of locations (simply run this code block)
with open('data/bay_area.txt', 'r') as f:
bay_area = f.read().splitlines()
with open('data/los_angeles_area.txt', 'r') as f:
los_angeles = f.read().splitlines()
with open('data/san_diego.txt', 'r') as f:
san_diego = f.read().splitlines()
# -
# Note that when text files are read using the <code style="color:steelblue">splitlines()</code> function, the resulting object is a list.
#
# So the three objects you just created from the files - <code style="color:steelblue">bay_area</code>, <code style="color:steelblue">los_angeles</code>, and<code style="color:steelblue">san_diego</code> - are all lists.
print( type(san_diego) )
# Let's start exploring this data.
#
# <br>
# **Print the first 5 locations in San Diego.**
# Print the first 5 locations in San Diego
san_diego[0:5]
# Next, we need to know how many breweries are in each location.
#
# <br>
# **Print the number of breweries in each list.**
# * Which city has the most locations?
# +
# Print length of each list
b_length = len(bay_area)
l_length = len(los_angeles)
s_length = len(san_diego)
print('Bay area has ' + str(b_length) + ' breweries.')
print('Los Angeles has ' + str(l_length) + ' breweries.')
print('San Diego has ' + str(s_length) + ' breweries.')
# -
# Next, your friend has a couple questions...
#
# They ask you to:
# * **Print <code style="color:steelblue">True</code> if <code>'Stone Brewing'</code> is in San Diego or <code style="color:steelblue">False</code> if it's not.**
# * **Print <code style="color:steelblue">True</code> if <code>'Area 51 Craft Brewing'</code> is in the Bay area or <code style="color:steelblue">False</code> if it's not.**
# +
# Is 'Stone Brewing' in San Diego?
if "Stone Brewing" in san_diego:
print(True)
else:
print(False)
# Is 'Area 51 Craft Brewing' in the Bay area?
if "Area 51 Craft Brewing" in bay_area:
print(True)
else:
print(False)
# -
# Print minimum value in san_diego
print('Minimum - ', sorted(san_diego)[0])
# Print maximum value in san_diego
print('Maximum - ', sorted(san_diego)[-1])
# Let's continue planning locations to visit. Before we continue, we need to remove duplicates from our lists because we don't have time to visit the same location twice.
#
# <br>
# **For each of the 3 lists of locations, print <code style="color:steelblue">True</code> if it has duplicate locations and <code style="color:steelblue">False</code> if it doesn't.**
# * Hint: A list with duplicates will have a greater length than a set of the same locations.
# +
# Bay area has duplicates?
b_len = len(bay_area)
print (b_len > len(set(bay_area)))
# Los Angeles has duplicates?
l_len = len(los_angeles)
print(l_len > len(set(los_angeles)))
# San Diego has duplicates?
s_len = len(san_diego)
print(s_len > len(set(san_diego)))
# -
# <br>
# **For the lists with duplicates, remove duplicates by converting them into sets. Then, convert them back into lists.**
# * Hint: <code style="color:steelblue">set()</code> and <code style="color:steelblue">list()</code> are your friends.
# +
# Convert lists to sets to remove duplicates, then convert them back to lists
bay_area = list(set(bay_area))
san_diego = list(set(san_diego))
# -
# Great, now lets double check to make sure the duplicates were removed.
# +
# Bay area has duplicates?
b_len = len(bay_area)
print (b_len > len(set(bay_area)))
# Los Angeles has duplicates?
l_len = len(los_angeles)
print(l_len > len(set(los_angeles)))
# San Diego has duplicates?
s_len = len(san_diego)
print(s_len > len(set(san_diego)))
# -
# Looks good! Now, let's look at a simple way to store the breweries in one place.
# We're almost ready to visit the breweries!
#
# However, it's too cumbersome to lug around the 3 different lists we created.
#
# <br>
# **Create a single dictionary named <code style="color:steelblue">brewery_dict</code> for the breweries in each location.**
# * Each key should be the name of the location.
# * Their values should be the lists of unique locations.
# Create location_dict
brewery_dict = {"Bay Area": bay_area, "Los Angeles": los_angeles, "San Diego": san_diego}
# Next, let's make sure the dictionary has the correct keys.
#
# <br>
# **Run the cell below and check the output.**
# * What do you think the code below is doing?
# * You'll learn more about <code style="color:steelblue">for</code> loops in the next lesson.
# Run this cell
for brewery in ['Bay Area', 'Los Angeles', 'San Diego']:
print( brewery in brewery_dict )
# <br>
# Did you get the expected output? If not, check the answer key before moving on.
#
# Suddenly, your friend walks over to you and says...
#
# > "Hmm... if you set up the dictionary correctly, you won't need the original lists anymore."
#
# > "Please get rid of them."
#
# <br>
# **Run this next code cell to overwrite the original borough lists with <code style="color:steelblue">None</code>.**
# Run this cell
bay_area, los_angeles, san_diego = None, None, None
# By the way, <code style="color:steelblue">None</code> is its own object type in Python.
#
# <br>
# > *<span style="color:tomato; font-weight:bold">None</span> is an object that denotes emptiness.*
#
# <br>
# For example:
print( type(None) )
# Now, we want to split our visit to California into two trips: one for Southern California and one for Northern California.
#
# <br>
# **Add two new items to your dictionary:**
# 1. **Key:** <code style="color:steelblue">'Southern California'</code>... **Value:** All locations in <code style="color:steelblue">'San Diego'</code> and <code style="color:steelblue">'Los Angeles'</code>.
# 2. **Key:** <code style="color:steelblue">'Northern California'</code>... **Value:** All locations in <code style="color:steelblue">'Bay Area'</code>.
#
# Since you got rid of your original lists, you'll have to use the values you've already stored in your dictionary.
# +
# Create a new key-value pair for 'Southern California'
brewery_dict["Southern California"] = brewery_dict["Los Angeles"] + brewery_dict["San Diego"]
# Create a new key-value pair for 'Northern California'
brewery_dict["Northern California"] = brewery_dict["Bay Area"]
# -
# ## Finally, let's just check that we have the right number of locations for each trip.
# * You should have 41 for Southern California
# * You should have 22 for Northern California
#
# <br>
# **Run the cell below and check that you get the expected output.**
print( len(brewery_dict['Southern California']) )
print( len(brewery_dict['Northern California']) )
# If you don't have the right number of locations, doublecheck that you removed duplicates and that you're concatenating the correct lists. You can also check the answer key for the solution.
#
# <br>
# **Once you have the right number of locations, let's save this object so we can use it in the next lesson. Run the cell below.**
# * We'll use a Python built-in package called <code style="color:steelblue">pickle</code> to do so.
# * Pickle saves an entire object in a file on your computer.
# +
# Import pickle library
import pickle
# Save object to disk
with open('./data/brewery_dict.pkl', 'wb') as f:
pickle.dump(brewery_dict, f)
# -
# <br>
#
# > *Now you have a dictionary of breweries in California.*
#
# > *In the next lesson, we'll look through the locations and pick one to start with.*
#
# <br>
#
# <hr style="border-color:royalblue;background-color:royalblue;height:1px;">
#
# <div style="text-align:center; margin: 40px 0 40px 0;">
# [**Back to Contents**](#toc)
# </div>
# <br id="l3">
# # Lesson 4: Flow and Functions
# ***In the previous lesson...***
#
# > *In the previous lesson, you created a brewery dictionary for the locations we're interested in visting.*
#
# Now we're ready to pick a location to start with.
#
# <br>
# **First, let's import <code style="color:steelblue">brewery_dict</code> again using <code style="color:steelblue">pickle</code>. Run this cell.**
# +
import pickle
# Read object from disk
with open('./data/brewery_dict.pkl', 'rb') as f:
brewery_dict = pickle.load(f)
# -
# Now we have the <code style="color:steelblue">brewery_dict</code> object again, but what if we forgot which keys are in the dictionary?
#
# <br>
# **Print the keys in <code style="color:steelblue">brewery_dict</code>.**
# Print the keys in brewery_dict
for keys in brewery_dict.keys():
print(keys)
# Ah, yes...
#
# Now, we need to choose between starting with <code style="color:steelblue">'Southern California'</code> or with <code style="color:steelblue">'Northern California'</code>. We should start with the list with more locations, so let's find which one it is.
#
# <br>
# **Write code, using <code style="color:steelblue">if</code> statements, that does the following:**
# * **If** our Southern California list has more locations than our Northern California list, print the message:
#
#
# <pre style="color:steelblue">I want to start in Southern California.</pre>
#
#
# * **Else if** our Northern California list has more locations than our Southern California list, print the message:
#
#
# <pre style="color:steelblue">I want to start in Northern California.</pre>
#
#
# * **Else** (i.e. they have the same number of locations), print the message:
#
#
# <pre style="color:steelblue">Either is fine. Flip a coin!</pre>
#
#
# Write code here
if len(brewery_dict["Southern California"]) > len(brewery_dict["Northern California"]):
print("I want to start in Southern California.")
elif len(location_dict["Northern California"]) > len(brewery_dict["Southern California"]):
print("I want to start in Northern California.")
else:
print ("Either is fine. Flip a coin!")
# <hr style="border-color:royalblue;background-color:royalblue;height:1px;">
#
# Remember we said that we wanted to start with the list with the most locations.
# * We already knew that it would be either Southern California or Northern California because those lists are combinations of 2 of the others.
# * However, what if we didn't know that?
#
# <br>
# **For each key in <code style="color:steelblue">brewery_dict</code>, print the number of locations in its list, like so:**
#
# <pre style="color:steelblue">
# Bay Area has 13 locations.
# San Diego has ...
# </pre>
#
# * **Tip:** You can iterate through keys and values of a dictionary at the same time using <code style="color:steelblue">.items()</code>, like so:
#
# <pre style="color:#bbb">
# for <strong style="color:steelblue">key, value</strong> in brewery_dict<strong style="color:steelblue">.items()</strong>:
# <span style="color:dimgray"># code block</span>
# </pre>
#
# * **Tip:** Remember, to insert multiple dynamic values into a string, you can just add more places to <code style="color:steelblue">.format()</code>, like so:
#
# <pre style="color:#bbb">
# '<strong style="color:steelblue">{}</strong> has <strong style="color:steelblue">{}</strong> locations.'.format(<strong style="color:steelblue">first_value, second_value</strong>)
# </pre>
# For each key in brewery_dict, print the number of breweries in its list
for key, value in brewery_dict.items():
print ("{} has {} breweries".format(key, len(value)))
# Now, let's give each brewery in Southern California a first impression based on its name.
#
# <br>
# **Combine <code style="color:steelblue">if</code> and <code style="color:steelblue">for</code> statements. For each brewery in Southern California...**
# * **If** its name has <code style="color:steelblue">'51'</code>, <code style="color:steelblue">'Coronado'</code>, <code style="color:steelblue">'Noble'</code> in it, print:
#
# <pre style="color:steelblue">{<strong>name</strong>} sounds good.</pre>
#
# * **Else If** its name has <code style="color:steelblue">'Stone'</code>, <code style="color:steelblue">'Ballast'</code> in it, print:
#
# <pre style="color:steelblue">{<strong>name</strong>} sounds awesome.</pre>
#
# * If its name doesn't sound pleasant or grand, just ignore it.
# * **Tip:** If you want to check if any word from a list is found in a string, you can use <code style="color:steelblue">any()</code>, like so:
#
# <pre style="color:steelblue">
# any( word in name for word in <strong>list_of_words</strong> )
# </pre>
# +
sounds_good = ['51', 'Coronado', 'Noble']
sounds_awesome = ['Stone', 'Ballast']
# Print first impression of each location in Southern California based on its names
for breweries in brewery_dict["Southern California"]:
if any(word in breweries for word in sounds_good):
print(breweries + " sounds good.")
elif any(word in breweries for word in sounds_awesome):
print (breweries + " sounds awesome.")
# -
# <hr style="border-color:royalblue;background-color:royalblue;height:1px;">
#
# **Using a list comprehension, create a new list called <code style="color:steelblue">good_locations</code>.**
# * It should contain locations in Southern California that <code style="color:steelblue">sound_good</code>.
# * Then print the list.
# * **Tip:** To check if any word from a list is found in a string, you can use <code style="color:steelblue">any()</code>.
# Create good_locations list using a list comprehension
good_breweries = [breweries for breweries in brewery_dict["Southern California"] if any(word in breweries for word in sounds_good)]
# Print the good-sounding locations
print(good_breweries)
# **Print the number pleasant-sounding locations we have.**
# Print number of good-sounding locations
print(str(len(good_breweries)))
# <hr style="border-color:royalblue;background-color:royalblue;height:1px;">
#
# **Write a function called <code style="color:steelblue">filter_locations</code> that takes two arguments:**
# 1. <code style="color:steelblue">location_list</code>
# 2. <code style="color:steelblue">words_list</code>
#
# The function should return the list of names in <code style="color:steelblue">location_list</code> that have any word in <code style="color:steelblue">words_list</code>.
#
# Code here
def filter_breweries(location_list, word_list):
names = [breweries for breweries in location_list if any(word in breweries for word in word_list)]
return names
# Next, let's test that function.
#
# <br>
# **Create a new <code style="color:steelblue">good_breweries</code> list using the function you just wrote.**
# * Pass in the list of Southern California breweries and the list of good-sounding words.
# * You should get the same breweries that you got just above.
# * Print the new list.
# +
# Create good_breweries using filter_breweries()
good_breweries = filter_breweries(brewery_dict["Southern California"], sounds_good)
# Print list of good-sounding breweries
print(good_breweries)
# -
# **Next, let's use this handy function to create a <code style="color:steelblue">awesome_breweries</code> list for breweries that sound awesome.**
# * Pass in the list of Southern California breweries and the list of awesome-sounding words.
# * Print the new list and confirm the expected output
# +
# Create awesome_locations using filter_locations()
awesome_locations = filter_breweries(brewery_dict["Southern California"], sounds_awesome)
# Print list of awesome-sounding breweries
print(awesome_locations)
# -
# Great, we'll start with these for our visit.
#
# <br>
#
# > *In this lesson, we filtered our lists of breweries.*
#
# <br>
# <hr style="border-color:royalblue;background-color:royalblue;height:1px;">
#
# <div style="text-align:center; margin: 40px 0 40px 0;">
# [**Back to Contents**](#toc)
# </div>
# <br id="l5">
# # Lesson 5: Pandas
# Then, let's import Pandas.
import pandas as pd
# Read Iris dataset:
# Read the iris dataset from a CSV file
df = pd.read_csv('./data/iris.csv')
# <br>
# **First, create a new DataFrame called <code style="color:steelblue">toy_df</code>. It should contain the first 5 rows plus the last 5 rows from our original Iris dataset.**
# * **Tip:** You already have a <code style="color:steelblue">.head()</code>, but what about a <code style="color:steelblue">.tail()</code>?
# * **Tip:** <code style="color:steelblue">pd.concat()</code> is your [friend](http://pandas.pydata.org/pandas-docs/stable/generated/pandas.concat.html).
# Create toy_df
df_h = df.head()
df_t = df.tail()
toy_df = pd.concat([df_h, df_t])
# **Next, display <code style="color:steelblue">toy_df</code>.**
# * After all, it will only be 10 rows.
# * In <code style="color:steelblue">toy_df</code>, you should have data from 2 different species of flower. Which are they?
# Display toy_df
toy_df
# You should have 'setosa' and 'virginica' flowers.
#
# **Next, display a summary table for <code style="color:steelblue">toy_df</code>.**
# * It should have the mean, standard deviation, and quartiles for each of the columns
# Describe toy_df
toy_df.describe()
# Since <code style="color:steelblue">toy_df</code> is only 10 rows, you can manually check the **mins** and **maxes**. Are they correct?
# Elementwise operations are very useful in machine learning, especially for feature engineering.
#
# <br>
#
# > *<span style="color:tomato; font-weight:bold">Feature engineering</span> is the process of creating new features (model input variables) from existing ones.*
#
# <br>
#
# We'll cover this topic in much more detail later, but let's first use our <code style="color:steelblue">toy_df</code> to illustrate the concept.
#
# In the Iris dataset, we have petal width and length, but what if we wanted to know petal area? Well, we can create a new <code style="color:steelblue">petal_area</code> feature (yes, the petals are not perfect rectangles, but that's fine).
#
# <br>
# **First, display the two columns of <code style="color:steelblue">petal_width</code> and <code style="color:steelblue">petal_length</code> in <code style="color:steelblue">toy_df</code>.**
# * **Tip:** You can index a DataFrame using a list of column names too, like so:
#
#
# <pre style="color:steelblue">df[['column_1', 'column_2']]</pre>
# Display petal_width and petal_length
petal_width = toy_df["petal_width"]
petal_length = toy_df["petal_length"]
toy_df[["petal_width", "petal_length"]]
# **Next, create a new <code style="color:steelblue">petal_area</code> feature in <code style="color:steelblue">toy_df</code>.**
# * Multiply the <code style="color:steelblue">petal_width</code> column by the <code style="color:steelblue">petal_length</code> column.
# * Display <code style="color:steelblue">toy_df</code> after creating the new feature.
# * Are the values for <code style="color:steelblue">petal_area</code> correct? Manually spot check a few of them just to make sure.
# +
# Create a new petal_area column
toy_df["petal_area"] = toy_df["petal_width"] * toy_df["petal_length"]
# Display toy_df
toy_df
# -
# **Finally, what do we now know about Iris flowers?**
#
# By creating a <code style="color:steelblue">petal_area</code> feature, it's now much easier to see that virginica flowers have significantly larger petals than setosa flowers do!
#
# Often, by creating new features, you can learn more about the data (and improve your machine learning models).
# <hr style="border-color:royalblue;background-color:royalblue;height:1px;">
#
#
# Let's say we wanted to display observations where <code style="color:steelblue">petal_area > 10</code> and <code style="color:steelblue">sepal_width > 3</code>. How could we do so?
#
# <br>
# **First, display <code style="color:steelblue">toy_df</code> again just to have it in front of you.**
# Display toy_df
toy_df
# **Take a look at the DataFrame and manually count the number that satisfy our conditions.**
# * How many observations have <code style="color:steelblue">petal_area > 10</code>?
# * How many observations have <code style="color:steelblue">sepal_width > 3</code>?
# * How many satisfy both conditions?
#
# Great. Now we'll see what's going on under the hood when we use our boolean masks.
#
# <br>
# **Create a boolean mask for <code style="color:steelblue">petal_area > 10</code>.**
# * Name it <code style="color:steelblue">petal_area_mask</code>.
# * Display the mask after you create it.
# * Does the result make sense?
# +
# Mask for petal_area > 10
petal_area_mask = toy_df["petal_area"] > 10
# Display petal_area_mask
petal_area_mask
# -
# **Next, create a boolean mask for <code style="color:steelblue">sepal_width > 3</code>.**
# * Name it <code style="color:steelblue">sepal_width_mask</code>.
# * Display the mask after you create it.
# * Does the result make sense?
# +
# Mask for sepal_width > 3
sepal_width_mask = toy_df["sepal_width"] > 3
# Display sepal_width_mask
sepal_width_mask
# -
# **Next, display the two masks combined using the <code style="color:steelblue">&</code> operator.**
# * Note how their combination results in another boolean mask!
# Display both masks, combined
sepal_width_mask & petal_area_mask
# **Finally, select the observations from <code style="color:steelblue">toy_df</code> where both conditions are met.**
# Index with both masks
toy_df[sepal_width_mask & petal_area_mask]
# <hr style="border-color:royalblue;background-color:royalblue;height:1px;">
#
# Now, armed with the power of **groupby**, let's just bring back our <code style="color:steelblue">toy_df</code> for one last hoorah, just to make sure we know what's going on under the hood.
#
# <br>
# Let's calculate the median <code style="color:steelblue">petal_area</code> for each species.
# * Since <code style="color:steelblue">toy_df</code> is small, we can do this manually as well and check to make sure the values are correct.
#
# <br>
# **First, let's manually calculate the median <code style="color:steelblue">petal_area</code> for the virginica flowers in our <code style="steelblue">toy_df</code>.**
# * Display all observations of the virginica species.
# * Sort them by <code style="color:steelblue">petal_area</code> in ascending order.
# * **Tip:** Check out the <code style="color:steelblue">.sort_values()</code> function ([documentation](http://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.sort_values.html)).
# Display all 'virginica' species, sorted by petal_area
toy_df[toy_df["species"] == "virginica"].sort_values("petal_area", axis=0, ascending="True")
# Based on the output above, what's median <code style="color:steelblue">petal_area</code> for the virginica species?
#
# <br>
# **Next, let's manually calculate the median <code style="color:steelblue">petal_area</code> for the setosa flowers in our <code style="steelblue">toy_df</code>.**
# * Display all observations of the setosa species.
# * Sort them by <code style="color:steelblue">petal_area</code> in ascending order.
# Display all 'setosa' species
toy_df[toy_df["species"] == "setosa"].sort_values("petal_area", axis=0, ascending="True")
# Based on the output above, what's median <code style="color:steelblue">petal_area</code> for the setosa species?
#
# <br>
# **Finally let's calculate the median values using a <code style="color:steelblue">.groupby()</code>.**
# * Do you get the same result?
# Median petal_area in toy_df
toy_df.groupby("species")["petal_area"].median()
# ** *Congratulations... You've completed the Python Crash Course!* **
#
# > *In this lesson, you explored the Iris dataset using Pandas.*
#
# <br>
# <hr style="border-color:royalblue;background-color:royalblue;height:1px;">
#
# <div style="text-align:center; margin: 40px 0 40px 0;">
# [**Back to Contents**](#toc)
# </div>
| Day_2/Python Crash Course.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + pycharm={"name": "#%%\n"}
from dataloader import get_loader
import numpy as np
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
import matplotlib.pyplot as plt
import numpy as np
import matplotlib.pyplot as plt
import mne
import pickle
import torch
import torch.nn as nn
from torch.utils.data import DataLoader, Subset, TensorDataset, RandomSampler
# -
class get_loader(nn.Module):
def __init__(self, root_dir, n_sess = 2, mode = 1):
'''
root_dir: the main folder with subject-wise subfolders
n_sess: number of sessions to consider. 1/2/3
mode: 0-pronounced
1-inner
2-visualized
'''
super(get_loader, self).__init__()
self.root_dir = root_dir
self.n_sess = n_sess
self.mode = mode #condition
def load_single_subject(self, sub_idx):
'''
sub_idx: 1, 2, ..... integers
'''
data = dict()
y = dict()
N_B_arr = np.arange(1, self.n_sess+1, 1)
N_S = sub_idx
print(N_S)
for N_B in N_B_arr:
# name correction if N_Subj is less than 10
if N_S<10:
Num_s='sub-0'+str(N_S)
else:
Num_s='sub-'+str(N_S)
file_name = root + Num_s + '/ses-0'+ str(N_B) + '/' +Num_s+'_ses-0'+str(N_B)+'_events.dat'
y[N_B] = np.load(file_name,allow_pickle=True)
# load data and events
file_name = root + Num_s + '/ses-0'+ str(N_B) + '/' +Num_s+'_ses-0'+str(N_B)+'_eeg-epo.fif'
X= mne.read_epochs(file_name,verbose='WARNING')
data[N_B]= X._data
#stack the sessions
X = data.get(1)
Y = y.get(1)
for i in range(2, self.n_sess+1, 1):
X = np.vstack((X, data.get(i)))
Y = np.vstack((Y, y.get(i)))
#select the recordings from required mode only
X_mode = X[Y[:,2] == self.mode]
Y_mode = Y[Y[:, 2] == self.mode]
return X_mode, Y_mode
def load_multiple_subjects(self, subjects):
'''
Load all subjects required and stack them into single array (n_rec*n_sub, 128, 1153)
'''
X_t, Y_t = [], []
for idx in subjects:
Xi, Yi = self.load_single_subject(idx)
X_t.append(Xi)
Y_t.append(Yi)
return np.vstack(X_t), np.vstack(Y_t)
def forward(self, subjects, batch_size = 1):
'''
subjects: list of subject indices to load data from
'''
X, Y = self.load_multiple_subjects(subjects)
X = torch.tensor(X)
Y = torch.tensor(Y)
train_data = TensorDataset(X, Y)
train_sampler = RandomSampler(X)
train_dataloader = DataLoader(X, sampler=train_sampler, batch_size=batch_size)
# can add test/validation loader too
return train_dataloader
# + pycharm={"name": "#%%\n"}
# Load in data
# root = '/Volumes/Datasets/inner_speech/derivatives/'
root = 'dataset/derivatives/' # -sil
creater = get_loader(root)
xn, yn = creater.load_multiple_subjects([1, 2, 3, 4, 5, 6, 7, 8])
# -
xt, yt = creater.load_single_subject(10)
# + [markdown] pycharm={"name": "#%% md\n"}
# # LDA
# + pycharm={"name": "#%%\n"}
# Vectorize data so that X is of size = (n_samples, n_features)
train_x = xn.reshape(xn.shape[0], xn.shape[1]*xn.shape[2])
test_x = xt.reshape(xt.shape[0], xt.shape[1]*xt.shape[2])
print("train_x: ", train_x.shape)
print("test_x: ", test_x.shape)
# Get Labels
train_y = yn[:,1]
test_y = yt[:,1]
print("train_y: ", train_y.shape)
print("test_y: ", test_y.shape)
# + pycharm={"name": "#%%\n"}
# Fit LDA
# solver='svd', shrinkage=None, priors=None, n_components=None, store_covariance=False, tol=0.0001, covariance_estimator=None
clf = LinearDiscriminantAnalysis()
train_x_proj = clf.fit_transform(train_x, train_y)
# + pycharm={"name": "#%%\n"}
# Predict on test data
predictions = clf.predict(test_x)
accuracy = clf.score(test_x, test_y)
print("Accuracy: ", accuracy*100)
# + pycharm={"name": "#%%\n"}
# Tranform data to LDA space
test_x_proj = clf.transform(test_x)
test_x_proj.shape
test_y
# +
# Visualize data in LDA Space
ax = plt.axes(projection ="3d")
for i in range(4):
data = test_x_proj[test_y == i, :]
ax.scatter3D(data[:,0], data[:,1], data[:,0], label=i)
plt.title("simple 3D scatter plot")
#ax.view_init(30,45)
ax.set_zlim([-2,3])
ax.set_ylim([-3,2])
ax.set_xlim([-2,2]) # there are some outliers which make it hard to see the giant clump unless you set axis limits
# But the projected data isnt very easily separable anyways
# -
# ## Classify with SVM
from sklearn import svm
import sklearn
# The learning is very fast
svm_clf = svm.SVC(C=.1, kernel="poly", degree=2) # best run
# svm_clf = svm.SVC(C=1, kernel="linear")
svm_clf.fit(train_x_proj, train_y)
acc = svm_clf.score(test_x_proj, test_y)
predictions = svm_clf.predict(test_x_proj)
print("Accuracy: ", acc)
# +
sklearn.metrics.ConfusionMatrixDisplay.from_predictions(test_y, predictions,display_labels=['up', 'down', 'right', 'left'])
# -
| lda_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Note: This example currently does not work in MyBinder.
# Install dependencies for this example
# Note: This does not include itkwidgets, itself
import sys
# !{sys.executable} -m pip install -U vedo
# +
from vedo import load, datadir, Sphere
from itkwidgets import view
embryo = load(datadir+'embryo.slc')
sph = Sphere(r=50).pos(10,20,30)
scals = sph.points()[:,2] # scalars are z coords
sph.pointColors(scals)
# -
embryo.printInfo()
view(actors=[embryo, sph])
| examples/vedo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# <div>
# <center><h1 style = "font-weight:lighter;">📍 Importing Libraries</h1></center>
# </div>
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LinearRegression
import reg2info as r2i
# <div>
# <center><h1 style = "font-weight:lighter;">📍 Loading dataset</h1></center>
# </div>
df = pd.read_csv("Salary_Data.csv")
df.head()
# <div>
# <center><h1 style = "font-weight:lighter;">📍 Splitting in to train and test set</h1></center>
# </div>
x= df.iloc[:, :-1].values
y= df.iloc[:, 1].values
x_train, x_test, y_train, y_test= train_test_split(x, y, test_size= 1/3, random_state=0)
# <div>
# <center><h1 style = "font-weight:lighter;">📍 Train your linear regression model</h1></center>
# </div>
lr= LinearRegression().fit(x_train, y_train)
# <div>
# <center><h1 style = "font-weight:lighter;">📍Getting the equation of lr</h1></center>
# </div>
eq = r2i.reg_equation(lr)
eq
# <div>
# <center><h1 style = "font-weight:lighter;">📍 Plotting the lr line and training data</h1></center>
# </div>
#
plot = r2i.reg_plot(lr, x, y)
| example/notebook_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Working with ML models for Time Series Analysis
# In this section, we will prepare the data for machine learning analysis by creating lagged variables. We will work with both stationary and non-stationary data. Three of the datasets that we are using are seasonally adjusted for stationarity through differencing. The two financial datasets are only resampled from daily to monthly data. We will create kernel density and autocorrelation plots as well as generate lag variables for a 12-time step period. We will then save these new dataframes. These dataframes will be used in the next 3 videos.
import pandas as pd
from pandas import read_csv
from matplotlib import pyplot
# ## Example 1: Vacation Dataset
# Read in data as a panda series
# https://trends.google.com/trends/?geo=US , google trends, search the word "vacation"
# Recall that data is from 2004 to 2019
vacation = pd.read_csv("~/Desktop/section_4/vacation_firstdiff.csv", index_col=0, parse_dates=True)
# display first few rows
print(vacation.head(5))
# line plot of dataset
vacation.plot(figsize=(8,5))
pyplot.show()
# data is monthly and is made stationary
# The plot shows that seasonality and trend were removed from the series through differening. Above is the plot of the differenced data.
vacation.describe()
# ### Kernel Density Plot
vacation.plot(kind='kde')
# ### Autocorrelation Plot
from statsmodels.graphics.tsaplots import plot_acf
# Check for autocorrelation of each lagged observation and whether it is statistically significant.
plot_acf(vacation)
pyplot.show()
# The blue shaded region is the margin of uncertainty. Candlesticks that extend out beyond the blue shaded region are considered statistically significant. Correlation values are between 1 and -1.
# ### Create Lagged Variables
from pandas import DataFrame
# reframe as supervised learning
# lag observation (t-1) is the input variable and t is the output variable.
df1 = DataFrame()
print(df1)
# Create 12 months of lag values to predict current observation
# Shift of 12 months
for i in range(12,0,-1):
df1[['t-'+str(i)]] = vacation.shift(i)
print(df1)
# Create column t
df1['t'] = vacation.values
print(df1.head(13))
# Create a new subsetted dataframe, removing Nans from first 12 rows
df1_vacat = df1[13:]
print(df1_vacat)
# save to new file
df1_vacat.to_csv('vacation_lags_12months_features.csv', index=False)
# ## Example 2: Furniture Dataset
# +
furniture = pd.read_csv("~/Desktop/section_4/furn_pctchange.csv", index_col=0, parse_dates=True)
furniture.head()
# data is monthly and made stationary
# -
# line plot of dataset
furniture.plot(figsize=(8,5))
pyplot.show()
furniture.describe()
# ## Kernel Density Plot
furniture.plot(kind='kde')
# ## Autocorrelation Plot
from statsmodels.graphics.tsaplots import plot_acf
# Check for autocorrelation of each lagged observation and whether it is statistically significant.
plot_acf(furniture)
pyplot.show()
# The blue shaded region is the margin of uncertainty. Candlesticks that extend out beyond the blue shaded region are considered statistically significant. Correlation values are between 1 and -1.
# ## Create Lagged Variables
from pandas import DataFrame
# reframe as supervised learning
# lab observation (t-1) is the input variable and t is the output variable.
df2 = DataFrame()
# print(df2)
# Create 12 months of lag values to predict current observation
# Shift of 12 months
for i in range(12,0,-1):
df2[['t-'+str(i)]] = furniture.shift(i)
print(df2)
# Create column t
df2['t'] = furniture.values
print(df2.head(13))
# Create a new subsetted dataframe, removing Nans from first 12 rows
df2_furn = df2[13:]
print(df2_furn)
# save to new file
df2_furn.to_csv('furniture_lags_12months_features.csv', index=False)
# ## Example 3: Bank of America Dataset
bac = pd.read_csv("~/Desktop/section_4/df_bankofamerica.csv", index_col=0, parse_dates=True)
# Convert the daily data to monthly data
bac= bac.resample(rule='M').last() # Don't use how='last', deprecated
# rule is monthly
# last means last day of the month
# +
# display first few rows
print(bac.head(5))
# line plot of dataset
bac.plot(figsize=(8,5))
pyplot.show()
# Notice that this data is monthly but not made stationary
# -
bac.describe()
bac.plot(kind='kde')
from statsmodels.graphics.tsaplots import plot_acf
# Check for autocorrelation of each lagged observation and whether it is statistically significant.
plot_acf(bac)
pyplot.show()
from pandas import DataFrame
# reframe as supervised learning
# lag observation (t-1) is the input variable and t is the output variable.
df3 = DataFrame()
# Create 12 months of lag values to predict current observation
# Shift of 12 months
for i in range(12,0,-1):
df3[['t-'+str(i)]] = bac.shift(i)
print(df3)
# Create column t
df3['t'] = bac.values
print(df3.head(13))
# Create a new subsetted dataframe, removing Nans from first 12 rows
df3_bac = df3[13:]
print(df3_bac)
# save to new file
df3_bac.to_csv('bac_lags_12months_features.csv', index=False)
# ## Example 4: J.P. Morgan Dataset
# +
jpm = pd.read_csv("~/Desktop/section_4/df_jpmorgan.csv", index_col=0, parse_dates=True)
# Convert the daily data to monthly data
jpm= jpm.resample(rule='M').last() # Don't use how='last', deprecated
# rule is monthly
# last means last day of the month
jpm.head()
# -
# display first few rows
print(jpm.head(5))
# line plot of dataset
jpm.plot(figsize=(8,5))
pyplot.show()
jpm.describe()
jpm.plot(kind='kde')
from statsmodels.graphics.tsaplots import plot_acf
# Check for autocorrelation of each lagged observation and whether it is statistically significant.
plot_acf(jpm)
pyplot.show()
from pandas import DataFrame
# reframe as supervised learning
# lag observation (t-1) is the input variable and t is the output variable.
df4 = DataFrame()
# print(df4)
# Create 12 months of lag values to predict current observation
# Shift of 12 months
for i in range(12,0,-1):
df4[['t-'+str(i)]] = jpm.shift(i)
print(df4)
# Create column t
df4['t'] = jpm.values
print(df4.head(13))
# Create a new subsetted dataframe, removing Nans from first 12 rows
df4_jpm = df4[13:]
print(df4_jpm)
# save to new file
df4_jpm.to_csv('jpm_lags_12months_features.csv', index=False)
# ## Example 5: Average Temperature of St. Louis dataset
temp = pd.read_csv("~/Desktop/section_4/temp_diffmean.csv", index_col=0, parse_dates=True)
# +
# display first few rows
print(temp.head(5))
# line plot of dataset
jpm.plot(figsize=(8,5))
pyplot.show()
# Notice that this is monthly and stationary data
# -
# summary statistics
temp.describe()
# kerndel density plot
temp.plot(kind='kde')
from statsmodels.graphics.tsaplots import plot_acf
# Check for autocorrelation of each lagged observation and whether it is statistically significant.
plot_acf(temp)
pyplot.show()
from pandas import DataFrame
# reframe as supervised learning
# lag observation (t-1) is the input variable and t is the output variable.
df5 = DataFrame()
# print(df5)
# Create 12 months of lag values to predict current observation
# Shift of 12 months
for i in range(12,0,-1):
df5[['t-'+str(i)]] = temp.shift(i)
print(df5)
# Create column t
df5['t'] = temp.values
print(df5.head(13))
# Create a new subsetted dataframe, removing Nans from first 12 rows
df5_temp = df5[13:]
print(df5_temp)
# save to new file
df5_temp.to_csv('temp_lags_12months_features.csv', index=False)
# +
# End
| Section 4/4.1_MachineLearningForTimeSeries.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8
# language: python
# name: python-3.8
# ---
# #### Node Embedding with Node2Vec in Stellargraph
#
# Requires Python 3.8
# +
import os
import psycopg2 as pg
import pandas.io.sql as psql
import pandas as pd
import networkx as nx
import matplotlib.pyplot as plt
from sklearn.manifold import TSNE
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LogisticRegressionCV
from sklearn.metrics import accuracy_score
from sklearn.metrics.pairwise import pairwise_distances
from sklearn import preprocessing
import numpy as np
from stellargraph.data import BiasedRandomWalk
from stellargraph import StellarGraph
from gensim.models import Word2Vec
import warnings
import collections
from stellargraph import datasets
from IPython.display import display, HTML
import matplotlib.pyplot as plt
# %matplotlib inline
print("finished")
# +
# connect to the database, download data
connection = pg.connect(host = 'postgis1', database = 'sdad',
user = os.environ.get('db_user'),
password = os.environ.get('db_pwd'))
edgelist = '''SELECT ctr1 AS source, ctr2 as target, repo_wts AS weight
FROM gh_sna.sna_intl_ctr_edgelist_dd_lchn_08
WHERE ctr1 != ctr2 AND ctr1 NOT IN (SELECT * FROM gh.bots_table) AND ctr2 NOT IN (SELECT * FROM gh.bots_table);'''
nodelist = '''SELECT login, country_code_vis FROM gh_sna.sna_ctr_ctry_codes'''
edgelist = pd.read_sql_query(edgelist, con=connection)
node_attrs = pd.read_sql_query(nodelist, con=connection)
edgelist.head()
# -
node_attrs.head()
# +
G = nx.from_pandas_edgelist(edgelist, source='source', target='target', edge_attr='weight')
def average_degree(G):
wtd_deg = list(G.degree(weight='weight'))
avg_wtd_degree = round(sum([row[1] for row in wtd_deg]) / len([row[1] for row in wtd_deg]), 3)
return avg_wtd_degree
num_edges = G.number_of_edges()
num_nodes = G.number_of_nodes()
avg_degree = average_degree(G)
print("Number of nodes: {}".format(num_edges))
print("Number of edges: {}".format(num_nodes))
print("Average degree: {}".format(avg_degree))
# -
plt.figure(figsize=(8,6))
nx.draw_networkx(G, arrows=False, with_labels=False, node_size=20)
nodelist = list(G.nodes())
nodelist = pd.DataFrame(nodelist, columns=['login'])
nodelist.count()
nodelist = pd.merge(nodelist,node_attrs,on='login',how='left')
nodelist = nodelist.rename(columns={"country_code_vis": "country"})
nodelist
attrs = nodelist.country.to_dict()
nx.set_node_attributes(G, attrs, "country") # left off here
nx.get_node_attributes(G, 'country')
# +
color_map = []
for node in G:
if node == "Pesa":
color_map.append('blue')
else:
color_map.append('orange')
plt.figure(figsize=(8,6))
nx.draw_networkx(G, node_color=color_map, arrows=False, with_labels=False, node_size=20)
# -
# #### Node Embedding
#
# First, we set the random walk global parameter and set a function for the jaccard_weights
# +
extract_weights = np.array(list(G.edges.data("weight")))
weights = extract_weights[:,2]
wt, cnt = np.unique(weights, return_counts=True)
plt.figure(figsize=(10, 8))
plt.bar(wt, cnt, width=0.005, color="b")
plt.title("Edge weights histogram")
plt.ylabel("Count")
plt.xlabel("edge weights")
plt.xticks(np.linspace(0, 1, 10))
plt.show()
# -
G = StellarGraph.from_networkx(G)
rw = BiasedRandomWalk(G)
# +
walk_length = 100
weighted_walks = rw.run(
nodes=G.nodes(), # root nodes
length=walk_length, # maximum length of a random walk
n=10, # number of random walks per root node
p=0.5, # Defines (unnormalised) probability, 1/p, of returning to source node
q=2.0, # Defines (unnormalised) probability, 1/q, for moving away from source node
weighted=True, # for weighted random walks
seed=42, # random seed fixed for reproducibility
)
print("Number of random walks: {}".format(len(weighted_walks)))
# -
weighted_model = Word2Vec(
weighted_walks, vector_size=128, window=5, min_count=0, sg=1, workers=1, epochs=1
)
emb = weighted_model.wv['rmehta']
emb.shape
# Retrieve node embeddings and corresponding subjects
node_ids = weighted_model.wv.index_to_key # list of node IDs
weighted_node_embeddings = (
weighted_model.wv.vectors
) # numpy.ndarray of size number of nodes times embeddings dimensionality
# the gensim ordering may not match the StellarGraph one, so rearrange
node_targets = nodelist.loc[node_ids].astype("category")
# Apply t-SNE transformation on node embeddings
tsne = TSNE(n_components=2, random_state=42)
weighted_node_embeddings_2d = tsne.fit_transform(weighted_node_embeddings)
# +
# draw the points
alpha = 0.7
plt.figure(figsize=(10, 8))
plt.scatter(
weighted_node_embeddings_2d[:, 0],
weighted_node_embeddings_2d[:, 1],
#c=node_targets.cat.codes,
cmap="jet",
alpha=0.7,
)
plt.show()
# -
# https://medium.com/stellargraph/can-graph-machine-learning-identify-hate-speech-in-online-social-networks-58e3b80c9f7e
dataset = datasets.Cora()
display(HTML(dataset.description))
G, subjects = dataset.load(
largest_connected_component_only=True,
edge_weights=jaccard_weights,
str_node_ids=True, # Word2Vec requires strings, not ints
)
subjects
nodes_list = np.array(list(G.nodes()))
node_name = nodes_list[4]
node_id = np.where(nodes_list)[0][0]
node_id
# References
#
# https://stellargraph.readthedocs.io/en/stable/demos/node-classification/node2vec-weighted-node-classification.html
# https://towardsdatascience.com/hands-on-graph-neural-networks-with-pytorch-pytorch-geometric-359487e221a8
# https://colab.research.google.com/drive/1b9rZIjD7MUEKwYbXZc3dchTBTpzdrvpd?usp=sharing#scrollTo=AUhES1VYo3tB
| src/05_github-networks/06_grl-testing/01_initial-code.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# https://www.hackerrank.com/challenges/py-set-mutations/problem
# +
_, the_set = int(input()), set(map(int,input().split()))
for _ in range(int(input())):
exec('the_set.{}({})'.format(input().split()[0], set(map(int, input().split()))))
print(sum(the_set))
| 04 - Sets/09 - Set Mutations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="ZFIqwYGbZ-df"
# # Timbre transfer FM
#
#
# + id="S_jXCnwZ2QYW" outputId="afaebd63-493d-4fb9-defa-c972b99457c6" colab={"base_uri": "https://localhost:8080/"}
USE_PRIVATE_DISTRO = True
DRIVE_BASE_DIR = '/content/drive/MyDrive/SMC 10/DDSP-10/'
DRIVE_DISTRO = DRIVE_BASE_DIR + 'dist/ddsp-1.2.0.tar.gz'
if USE_PRIVATE_DISTRO:
print("[INFO] Using private distro.")
from google.colab import drive
drive.mount('/content/drive')
# !pip install -qU "$DRIVE_DISTRO"
else:
# !pip install -qU ddsp
import warnings
import gin
import tensorflow as tf
# %reload_ext tensorboard
import tensorboard as tb
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
# %config InlineBackend.figure_format='retina'
from ddsp.colab.colab_utils import specplot
from ddsp.colab.colab_utils import play
from ddsp.training import data
from ddsp.training import models
from ddsp import core
# + [markdown] id="zpetvejYO0KQ"
# #### Configuration
# + id="mkFYv_DUZ7lW"
SAMPLE_RATE = 48000
DURATION = 4
FRAME_RATE = 250
TIME_STEPS = FRAME_RATE * DURATION
N_SAMPLES = SAMPLE_RATE * DURATION
MOD_FREQ = 100
INSTRUMENT = 'sr{}k_mf{}'.format(SAMPLE_RATE//1000, MOD_FREQ)
sns.set(style="whitegrid")
warnings.filterwarnings("ignore")
OUTPUT_FOLDER = 'fm_timbretrans_01_ar_fixed' #@param {type: "string"}
DRIVE_CHECKPOINTS_DIR = DRIVE_BASE_DIR + 'audio/' + OUTPUT_FOLDER + '/' + \
INSTRUMENT + '_checkpoints/'
DRIVE_TFRECORD_PATTERN = DRIVE_BASE_DIR + 'audio/' + OUTPUT_FOLDER + '/' + \
INSTRUMENT + '_dataset/train.synthrecord*'
# !mkdir -p "$DRIVE_CHECKPOINTS_DIR"
# + [markdown] id="Op0V8onI0VUK"
# #### Start Tensorboard
# + id="hBvbrMQvGzK9"
tb.notebook.start('--logdir "{}"'.format(DRIVE_CHECKPOINTS_DIR))
# + [markdown] id="Q9D9ozX6PAXB"
# #### Train the model
# + id="VDGtUMk3PGGy"
# !ddsp_run \
# --mode=train \
# --alsologtostderr \
# --stop_at_nan \
# --save_dir="$DRIVE_CHECKPOINTS_DIR" \
# --gin_file=models/fm_simple.gin \
# --gin_file=datasets/synthrecord.gin \
# --gin_param="SynthRecordProvider.file_pattern='$DRIVE_TFRECORD_PATTERN'" \
# --gin_param="SynthRecordProvider.sample_rate=$SAMPLE_RATE" \
# --gin_param="SynthRecordProvider.frame_rate=$FRAME_RATE" \
# --gin_param="SynthRecordProvider.example_secs=$DURATION" \
# --gin_param="RnnFcDecoder.rnn_channels=64" \
# --gin_param="RnnFcDecoder.ch=64" \
# --gin_param="F0MIDILoudnessPreprocessor.time_steps=$TIME_STEPS" \
# --gin_param="FrequencyModulation.n_samples=$N_SAMPLES" \
# --gin_param="FrequencyModulation.sample_rate=$SAMPLE_RATE" \
# --gin_param="FrequencyModulation.ar_scale=True" \
# --gin_param="FrequencyModulation.index_scale=True" \
# --gin_param="train_util.train.batch_size=16" \
# --gin_param="train_util.train.num_steps=6000" \
# --gin_param="train_util.train.steps_per_save=100" \
# --gin_param="train_util.train.steps_per_summary=50" \
# --gin_param="Trainer.checkpoints_to_keep=2" \
# --gin_param="Trainer.learning_rate=0.0005" \
# --gin_param="FilteredNoise.n_samples=$N_SAMPLES" \
# --early_stop_loss_value=4 \
# + [markdown] id="Ep_TMUitRz6y"
# #### Load pretrained model
# + id="qaS1PmlqR3JB"
data_provider_eval = data.SynthRecordProvider(DRIVE_TFRECORD_PATTERN,
sample_rate=SAMPLE_RATE,
frame_rate=FRAME_RATE,
example_secs=DURATION)
dataset_eval = data_provider_eval.get_batch(batch_size=1, shuffle=True, repeats=-1)
dataset_eval_iter = iter(dataset_eval)
gin_file = DRIVE_CHECKPOINTS_DIR + 'operative_config-0.gin'
gin.parse_config_file(gin_file)
model = models.Autoencoder()
model.restore(DRIVE_CHECKPOINTS_DIR)
for f in range(12):
frame = next(dataset_eval_iter)
# + id="jJNBloZ84hcW" outputId="951ac0f8-96a6-48d0-f590-849046623a0d" colab={"base_uri": "https://localhost:8080/", "height": 786}
# frame = next(dataset_eval_iter)
audio_baseline = frame['audio']
controls = model(frame, training=False)
audio_full = model.get_audio_from_outputs(controls)
audio_full /= tf.reduce_max(audio_full[0,:], axis=0).numpy()*1.5
print('Original Audio')
play(audio_baseline, sample_rate=SAMPLE_RATE)
print('Full reconstruction')
play(audio_full, sample_rate=SAMPLE_RATE)
for synth in ['harmonic', 'fm', 'filtered_noise']:
if synth in controls:
print('Only ' + synth)
play(controls[synth]['signal'], sample_rate=SAMPLE_RATE)
specplot(audio_baseline)
specplot(audio_full)
# get = lambda key: core.nested_lookup(key, controls)[0] #batch 0
# amps = get('fm/controls/amps')
# mod_amps = get('fm/controls/mod_amps')
# f0 = get('fm/controls/f0_hz')
# mod_f0 = get('fm/controls/mod_f0_hz')
# + id="i-IzBMyVST7d" outputId="11f87728-32af-4de5-80a0-d81178cea6e2" colab={"base_uri": "https://localhost:8080/", "height": 608}
c = controls['fm']['controls']
x = np.linspace(0,DURATION,TIME_STEPS)
f, ax = plt.subplots(4, 1, figsize=(15, 8), sharex=True, )
# f.suptitle('Inferred controls', fontsize=14)
plt.tight_layout()
ax[0].plot(x,c['op1'][0,:,0])
ax[0].plot(x,c['op2'][0,:,0])
ax[0].plot(x,c['op3'][0,:,0])
ax[0].plot(x,c['op4'][0,:,0])
ax[0].legend(['$A_1$','$A_2$','$A_3$','$A_4$'])
ax[0].set_title('Output levels')
ax[0].set_ylim(0)
# *c['f0'][0,:,0]
ax[1].plot(x,c['op1'][0,:,1])
ax[1].plot(x,c['op2'][0,:,1])
ax[1].plot(x,c['op3'][0,:,1])
ax[1].plot(x,c['op4'][0,:,1])
ax[1].legend(['$I_1$','$I_2$','$I_3$','$I_4$'])
ax[1].set_title('Freq. factor')
ax[2].plot(x,c['op1'][0,:,2])
ax[2].plot(x,c['op2'][0,:,2])
ax[2].plot(x,c['op3'][0,:,2])
ax[2].plot(x,c['op4'][0,:,2])
ax[2].legend(['$E_1$','$E_2$','$E_3$','$E_4$'])
ax[2].set_title('Envelopes')
ax[2].set_ylim(0)
for mod in range(6):
ax[3].plot(x,c['modulators'][0,:,mod])
ax[3].legend(['$M_{2,1}$','$M_{3,1}$','$M_{3,2}$','$M_{4,1}$','$M_{4,2}$','$M_{4,3}$'], loc='center right')
ax[3].set_ylim(0)
ax[3].set_title('Modulators')
ax[3].set_xlabel('Seconds')
f.show()
# + id="7q-rQHL1PAuS"
raise SystemExit("Stop right there!")
| ddsp/colab/fm/41_ttransfer_FM_simple.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
import scipy.io as sio
import math
import pandas as pd
import time
tic=time.time()
dataset = sio.loadmat('anomalyData.mat')
#server data
dataset
X=dataset['X']
#training dataset
Xval = dataset['Xval'] #cross-validation
yval = dataset['yval'] #the corresponding output
print(X.shape) #(rows,cols)
#the dataset contains throughput(mb/s) and latency(ms)
#plotting on the graph
plt.scatter(X[:, 0], X[:, 1], marker = "x")
plt.xlabel('Latency(ms)')
plt.ylabel('Throughput(mb/s)')
# +
#calculating the mean and variance
def estimateGaussian(X):
n = np.size(X, 1)
m = np.size(X, 0)
mu = np.zeros((n, 1))
sigma2 = np.zeros((n, 1))
mu = np.reshape((1/m)*np.sum(X, 0), (1, n))
sigma2 = np.reshape((1/m)*np.sum(np.power((X - mu),2), 0),(1, n))
return mu, sigma2
mu, sigma2 = estimateGaussian(X)
# -
mu, sigma2 = estimateGaussian(X)
print('mean: ',mu,' variance: ',sigma2)
#converting sigma2 into a covariance matrix
def multivariateGaussian(X, mu, sigma2):
n = np.size(sigma2, 1)
m = np.size(sigma2, 0)
#print(m,n)
if n == 1 or m == 1:
# print('Yes!')
sigma2 = np.diag(sigma2[0, :])
#print(sigma2)
X = X - mu
pi = math.pi
det = np.linalg.det(sigma2)
inv = np.linalg.inv(sigma2)
val = np.reshape((-0.5)*np.sum(np.multiply((X@inv),X), 1),(np.size(X, 0), 1))
#print(val.shape)
p = np.power(2*pi, -n/2)*np.power(det, -0.5)*np.exp(val)
return p
p = multivariateGaussian(X, mu, sigma2)
print(p.shape)
#calculating the threshold values
pval = multivariateGaussian(Xval, mu, sigma2)
def selectThreshHold(yval, pval):
F1 = 0
bestF1 = 0
bestEpsilon = 0
stepsize = (np.max(pval) - np.min(pval))/1000
epsVec = np.arange(np.min(pval), np.max(pval), stepsize)
noe = len(epsVec)
for eps in range(noe):
epsilon = epsVec[eps]
pred = (pval < epsilon)
prec, rec = 0,0
tp,fp,fn = 0,0,0
try:
for i in range(np.size(pval,0)):
if pred[i] == 1 and yval[i] == 1:
tp+=1
elif pred[i] == 1 and yval[i] == 0:
fp+=1
elif pred[i] == 0 and yval[i] == 1:
fn+=1
prec = tp/(tp + fp)
rec = tp/(tp + fn)
F1 = 2*prec*rec/(prec + rec)
if F1 > bestF1:
bestF1 = F1
bestEpsilon = epsilon
except ZeroDivisionError:
print('Warning dividing by zero!!')
return bestF1, bestEpsilon
#using F1 score method to determine the best parameter
F1, epsilon = selectThreshHold(yval, pval)
print('Epsilon and F1 are:',epsilon, F1)
#anomalies as outliers
outl = (p < epsilon)
# +
#returning the indices of the outliers to identify the faulty servers.
def findIndices(binVec):
l = []
for i in range(len(binVec)):
if binVec[i] == 1:
l.append(i)
return l
#This gives us a vector with binary entries where 1 means anomaly and 0 means normal.
# -
listOfOutliers = findIndices(outl)
count_outliers = len(listOfOutliers)
print('\n\nNumber of outliers:', count_outliers)
print('\n',listOfOutliers)
#got our faulty servers
plt.scatter(X[:, 0], X[:, 1], marker = "x")
plt.xlabel('Latency(ms)')
plt.ylabel('Throughput(mb/s)')
plt.scatter(X[listOfOutliers,0], X[listOfOutliers, 1], facecolors = 'none', edgecolors = 'r')
plt.show()
# +
#Creating the model
newDataset = sio.loadmat('anomalyDataTest.mat')
#this newDataset has 1000 examples each having 11 features.
#Xvaltest is the cross-validation set for the test samples
#yvaltest the corresponding labels.
Xtest = newDataset['X']
Xvaltest = newDataset['Xval']
yvaltest = newDataset['yval']
# +
mutest, sigma2test = estimateGaussian(Xtest)
ptest = multivariateGaussian(Xtest, mutest, sigma2test)
pvaltest = multivariateGaussian(Xvaltest, mutest, sigma2test)
F1test, epsilontest = selectThreshHold(yvaltest, pvaltest)
print('\nBest epsilon and F1 are\n',epsilontest, F1test)
# -
print(Xtest.shape)
print(Xvaltest.shape)
print(yvaltest.shape)
print(pvaltest.shape)
print(ptest.shape)
outliersTest = ptest < epsilontest
listOfOl = findIndices(outliersTest)
print('\n\n Outliers are:\n',listOfOl)
print('\n\nNumber of outliers are: ',len(listOfOl))
toc = time.time()
print('\n\nTotal time taken: ',str(toc - tic),'sec')
| Anomaly Detection on Server dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import scipy.sparse as sparse
import scipy.linalg as la
def grad(f, x):
'''
Input:
f: lambda function
x: function args
Output:
grad_f: function gradient at x
'''
n = len(x)
grad_f = np.zeros(n)
E = np.diag([pow(np.finfo(float).eps, 1/3) * (abs(a) + 1) for a in x])
for i in range(n):
grad_f[i] = (f(x + E[:, i]) - f(x - E[:, i])) * (0.5 / E[i, i])
return grad_f
def hess(f, x):
'''
Input:
f: lambda function
x: function args
Output:
hess_f: hessian of f at x
'''
n = len(x)
hess_f = np.zeros([n, n])
E = np.diag([pow(np.finfo(float).eps, 1/4) * (abs(a) + 1) for a in x])
for i in range(n):
for j in range(n):
hess_f[i, j] = ( f(x + E[:, i] + E[:, j])
- f(x - E[:, i] + E[:, j])
- f(x + E[:, i] - E[:, j])
+ f(x - E[:, i] - E[:, j]) ) * (0.25 / (E[i, i] * E[j, j]))
return hess_f
def cyclic_coordinate_descent(x0, A, b, tol=1e-5, maxiter=1000):
x = np.copy(x0)
n = A.shape[0]
m = 0
k = 0
r = A.dot(x) - b
while la.norm(r, np.inf) > tol and (m*n + k) < maxiter:
if k >= n:
k = 0
m = m + 1
alpha = -r[k]/A[k, k]
x[k] = x[k] + alpha
r = A.dot(x) - b
k += 1
return x, m*n + k
# -
A = sparse.diags([[2]*100, [1]*100, [1]*100], [0, -1, 1])
b = np.ones((100))
A
b
[x, i] = cyclic_coordinate_descent(np.zeros(100), A.toarray(), b, maxiter = np.inf)
A.dot(x)
i
B = np.array([[1, 0], [0, 3]])
d = np.array([1, 1])
x1 = np.array([-1, 3])
cyclic_coordinate_descent(x1, B, d, maxiter=10)
B.dot([3, 2])
B.dot(x1)
B[1, 1]
B[1][1]
| Labs/Lab 7.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] deletable=true editable=true
# # Introducing the Jupyter Notebook
# + [markdown] deletable=true editable=true
# ## What is it?
#
# From [jupyter.org](http://jupyter.org/):
#
# > An open-source web application that allows you to create and share documents that contain live code, equations, visualizations and explanatory text.
#
# The "document", aka "notebook", is composed of Code and Markdown cells, which you interact with using Command and Edit mode.
# + [markdown] deletable=true editable=true
# ## Command mode
#
# - Show keyboard shortcuts: `h`
# - Move up and down: `j`/`Down` and `k`/`Up`
# - Insert cell above or below: `a` or `b`
# - Run cell: `Shift`+`Enter`
# - Save: `S`
# - Switch to Edit mode: `Enter`
# + [markdown] deletable=true editable=true
# ## Markdown cells
#
# We're already using these!
# + [markdown] deletable=true editable=true
# ## Code cells
#
# Similar to running Python from the command line, but each one is a mini-editor with syntax highlighting, auto-indent, and more.
#
# * Autocomplete: `Tab`
# * Help: `Shift`+`Tab` (repeat for more)
# * Run cell: `Shift`+`Enter`
# * Switch to Command mode: `Esc`
# + deletable=true editable=true
hello = 'Hello, {}!'
who = input('Who? ')
hello.format(who)
# + deletable=true editable=true
# + [markdown] deletable=true editable=true
# ## IPython?
# + [markdown] deletable=true editable=true
# [IPython](http://ipython.org/) is a powerful replacement for the standard Python shell, which spawned the IPython Notebook. Eventually, the developers extracted the language-independent features of notebook web application to create the Jupyter Notebook. This "Big Split" occurred almost 2 years ago, but you'll still find lots of references to IPython in documentation and examples.
#
# - IPython still serves as the backend (aka "[kernel](http://jupyter.readthedocs.io/en/latest/projects/kernels.html)") for running Python code in the Notebook
# - "Jupyter" is an amalgamation of "Julia", "Python", and "R", which were the most popular languages used with the Notebook.
#
#
# + [markdown] deletable=true editable=true
# ## Getting started with Django
#
# [Django Extensions](https://django-extensions.readthedocs.io/en/latest/shell_plus.html) makes this easy. From within your virtual environment:
#
# ```
# (venv)$ pip install django-extensions jupyter
# (venv)$ cd /path/to/notebooks
# (venv)$ /path/to/manage.py shell_plus --notebook
# ```
#
# This will start the Notebook server, then open the Notebook Dashboard in your default web browser. The Dashboard lists the contents of the directory, and provides tools for starting and managing individual notebooks.
#
# 
#
# - Open an existing notebook by clicking on a `.ipynb` file
# - Create a new notebook by selecting "Django Shell-Plus" from the "New" menu in the upper-right corner.
#
# __NOTE__: There's a [known issue](https://github.com/django-extensions/django-extensions/issues/1026) with `manage.py shell_plus --notebook` for users who want to use the Notebook with multiple Django projects. Hopefully there will be a fix or a documented alternative soon.
# -
| 1 - Introducing the Jupyter Notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Magma Simulator
import magma as m
import mantle
# ## Simulating Combinational Circuits
#
# Create a 2-input multiplexor.
Mux2 = mantle.Mux2
print(Mux2)
# +
from magma.simulator import PythonSimulator
simulator = PythonSimulator(Mux2)
print('I0 I1 S O')
for i0 in [0, 1]:
for i1 in [0, 1]:
for s in [0, 1]:
simulator.set_value(Mux2.I0, i0)
simulator.set_value(Mux2.I1, i1)
simulator.set_value(Mux2.S, s)
simulator.evaluate()
print(i0, i1, s, int(simulator.get_value(Mux2.O)))
# -
# ## Simulating Sequential Logic Circuits with Clocks
#
# Create a 2-bit Counter.
# +
Counter = mantle.DefineCounter(2)
print(Counter)
# -
# Create a python simulator for the ice40.
# +
from magma.simulator import PythonSimulator
from hwtypes import BitVector
simulator = PythonSimulator(Counter, clock=Counter.CLK)
O0 = []
O1 = []
COUT = []
for i in range(16):
for j in range(2):
simulator.advance()
O = BitVector[2](simulator.get_value(Counter.O))
O0.append(int(O[0]))
O1.append(int(O[1]))
COUT.append(simulator.get_value(Counter.COUT))
# -
# Use `matplotlib` to plot the output of the simulator.
# +
import matplotlib.pyplot as plt
import numpy as np
n = len(O1) // 2
t = np.repeat(0.5 * np.arange(2*n + 1) ,2)[1:-1]
clock = np.tile(np.array([1, 1, 0, 0]), n)
O0 = np.repeat(np.array(O0), 2)
O1 = np.repeat(np.array(O1), 2)
COUT = np.repeat(np.array(COUT), 2)
plt.text(-1.5, 3.25, 'clock')
plt.plot(t, 0.5*clock + 3, 'r', linewidth=2)
plt.text(-1.5, 2.25, 'O[0]')
plt.plot(t, 0.5*O0 + 2, 'r', linewidth=2)
plt.text(-1.5, 1.25, 'O[1]')
plt.plot(t, 0.5*O1 + 1, 'r', linewidth=2)
plt.text(-1.5, 0.25, 'COUT')
plt.plot(t, 0.5*COUT, 'r', linewidth=2)
plt.xlim([0, n])
plt.ylim([0, 4])
plt.gca().axis('off')
plt.show()
# -
| notebooks/advanced/simulate.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import clima_anom as ca
import cartopy
import cartopy.feature as feature
import cartopy.crs as ccrs
import matplotlib.pyplot as plt
from netCDF4 import Dataset
Ano = '2018'
Mes = '03'
data_dir = '/mnt/Data/Data/GLM/Campinas/list/list_5min/'
data_out = '/mnt/Data/Data/GLM/Campinas/2018/point_5min/'
# +
# Region = 'America_sul'
Region = 'Sao_paulo'
print(f'Region: {Region}')
# -
for d in range(31):
Dia = ("{:02d}".format(d+1))
print(Dia)
for h in range(24):
Hora = ("{:02d}".format(h))
for m in range(12):
Minu = ("{:02d}".format(m))
Minu_s = 5*int(Minu)
Minu_s_s = ("{:02d}".format(Minu_s))
Minu_e = 5*int(Minu)+5
dir_lista = 'Lista_'+Ano+'_'+Mes+'_'+Dia+'_'+Hora+'_'+Minu_s_s+'.csv'
lista = pd.read_csv(data_dir+dir_lista)
# print(' ',Hora,':',Minu_s_s,dir_lista)
GLM_dia = []
GLM_hora = []
GLM_min = []
GLM_lon = []
GLM_lat = []
for t in range(len(lista)):
name_v1 = lista['File'][t]
data = Dataset(name_v1,mode = 'r')
lat = data['flash_lat']
lon = data['flash_lon']
area = data['flash_area']
energy = data['flash_energy']
GLM_lat = np.concatenate(([GLM_lat,lat]), axis=0)
GLM_lon = np.concatenate(([GLM_lon,lon]), axis=0)
GLM_flash = pd.DataFrame({'dia':Dia,'hora':Hora,'minuto':Minu_s_s,'Lat':GLM_lat,'Lon':GLM_lon})
if Region == 'America_sul':
out1 = GLM_flash[(GLM_flash['Lon'] > -85) & (GLM_flash['Lon'] < -30 )]
out2 = out1[(out1['Lat'] > -45) & (out1['Lat'] < 10 )]
elif Region == 'Sao_paulo':
out1 = GLM_flash[(GLM_flash['Lon'] > -50) & (GLM_flash['Lon'] < -44 )]
out2 = out1[(out1['Lat'] > -25) & (out1['Lat'] < -20 )]
else:
print('ERROR: this option dont exists')
break
salida = data_out + Ano + '_' + Mes + '_' + Dia + '_' + Hora + '_' + Minu_s_s + '.csv'
export_csv = out2.to_csv(salida, index = None, header=True)
| python/Accum_5min_GLM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from simtk.openmm.app import *
from simtk.openmm import *
from simtk.unit import *
f = open('ABCD.csv', 'w') #file to write chain data in
print('Loading...')
pdb = PDBFile('1kzy.clean.pdb')
forcefield = ForceField('amber99sb.xml', 'tip3p.xml')
modeller = Modeller(pdb.topology, pdb.positions) #initialize modeller
print('Adding hydrogens...')
modeller.addHydrogens(forcefield) #create forcefield
print('Adding solvent...')
modeller.addSolvent(forcefield, model='tip3p', padding=1*nanometer) #neutralize the system with ions
print('Minimizing...')
system = forcefield.createSystem(modeller.topology, nonbondedMethod=PME)
integrator = LangevinIntegrator(300*kelvin, 1/picosecond, 0.001*picoseconds)
##integrator = VerletIntegrator(2.0*femtoseconds)
platform = Platform.getPlatformByName('OpenCL')
simulation = Simulation(modeller.topology, system, integrator, platform)
simulation.context.setPositions(modeller.positions)
simulation.minimizeEnergy()
simulation.reporters.append(PDBReporter('ABCDtmp.pdb', 1000))
print('Saving...')
simulation.reporters.append(StateDataReporter(f, 100, step=True,kineticEnergy=True,
potentialEnergy=True,totalEnergy = True,density = True,temperature=True))
#for every 100 steps, write into f
simulation.step(30000) #run for 30000 steps
print('done')
# +
from simtk.openmm.app import *
from simtk.openmm import *
from simtk.unit import *
f = open('CD.csv', 'w')#file to write chain data in
print('Loading...')
pdb = PDBFile('CD.pdb')
forcefield = ForceField('amber99sb.xml', 'tip3p.xml')#create forcefield
modeller = Modeller(pdb.topology, pdb.positions)#initialize modeller
print('Adding hydrogens...')
modeller.addHydrogens(forcefield)
print('Adding solvent...')
modeller.addSolvent(forcefield, model='tip3p', padding=1*nanometer)#neutralize the system with ions
print('Minimizing...')
system = forcefield.createSystem(modeller.topology, nonbondedMethod=PME)
integrator = LangevinIntegrator(300*kelvin, 1/picosecond, 0.001*picoseconds)
platform = Platform.getPlatformByName('OpenCL')
simulation = Simulation(modeller.topology, system, integrator, platform)
simulation.context.setPositions(modeller.positions)
simulation.minimizeEnergy()
simulation.reporters.append(PDBReporter('CDtest.pdb', 1000))
print('Saving...')
simulation.reporters.append(StateDataReporter(f, 100, step=True,kineticEnergy=True,
potentialEnergy=True,totalEnergy = True,temperature=True,density = True))
#for every 100 steps, write into f
simulation.step(30000)#run for 30000 steps
print('done')
# +
from simtk.openmm.app import *
from simtk.openmm import *
from simtk.unit import *
f = open('AB.csv', 'w')#file to write chain data in
print('Loading...')
pdb = PDBFile('AB.pdb')
forcefield = ForceField('amber99sb.xml', 'tip3p.xml')#create forcefield
modeller = Modeller(pdb.topology, pdb.positions)#initialize modeller
print('Adding hydrogens...')
modeller.addHydrogens(forcefield)
print('Adding solvent...')
modeller.addSolvent(forcefield, model='tip3p', padding=1*nanometer)#neutralize the system with ions
print('Minimizing...')
system = forcefield.createSystem(modeller.topology, nonbondedMethod=PME)
integrator = LangevinIntegrator(300*kelvin, 1/picosecond, 0.001*picoseconds)
##integrator = VerletIntegrator(2.0*femtoseconds)
platform = Platform.getPlatformByName('OpenCL')
simulation = Simulation(modeller.topology, system, integrator, platform)
simulation.context.setPositions(modeller.positions)
simulation.minimizeEnergy()
simulation.reporters.append(PDBReporter('ABtmp.pdb', 1000))
print('Saving...')
simulation.reporters.append(StateDataReporter(f, 100, step=True,kineticEnergy=True,
potentialEnergy=True,totalEnergy = True,temperature=True,density = True))
#for every 100 steps, write into f
simulation.step(30000)#run for 30000 steps
print('done')
# -
# ## NVE
# #### constant number of particles
# #### constant volume
# #### constant energy or conserved energy
pdb.topology
import py3Dmol #visualize protein sample
view = py3Dmol.view(query='pdb:1kzy')
view.setStyle({'cartoon':{'color':'spectrum'}})
view
| bin/MD_Calculate_Energy_1KZY.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Analyze A/B Test Results
#
# You may either submit your notebook through the workspace here, or you may work from your local machine and submit through the next page. Either way assure that your code passes the project [RUBRIC](https://review.udacity.com/#!/projects/37e27304-ad47-4eb0-a1ab-8c12f60e43d0/rubric). **Please save regularly.**
#
# This project will assure you have mastered the subjects covered in the statistics lessons. The hope is to have this project be as comprehensive of these topics as possible. Good luck!
#
# ## Table of Contents
# - [Introduction](#intro)
# - [Part I - Probability](#probability)
# - [Part II - A/B Test](#ab_test)
# - [Part III - Regression](#regression)
#
#
# <a id='intro'></a>
# ### Introduction
#
# A/B tests are very commonly performed by data analysts and data scientists. It is important that you get some practice working with the difficulties of these
#
# For this project, you will be working to understand the results of an A/B test run by an e-commerce website. Your goal is to work through this notebook to help the company understand if they should implement the new page, keep the old page, or perhaps run the experiment longer to make their decision.
#
# **As you work through this notebook, follow along in the classroom and answer the corresponding quiz questions associated with each question.** The labels for each classroom concept are provided for each question. This will assure you are on the right track as you work through the project, and you can feel more confident in your final submission meeting the criteria. As a final check, assure you meet all the criteria on the [RUBRIC](https://review.udacity.com/#!/projects/37e27304-ad47-4eb0-a1ab-8c12f60e43d0/rubric).
#
# <a id='probability'></a>
# #### Part I - Probability
#
# To get started, let's import our libraries.
import pandas as pd
import numpy as np
import random
import matplotlib.pyplot as plt
# %matplotlib inline
#We are setting the seed to assure you get the same answers on quizzes as we set up
random.seed(42)
# `1.` Now, read in the `ab_data.csv` data. Store it in `df`. **Use your dataframe to answer the questions in Quiz 1 of the classroom.**
#
# a. Read in the dataset and take a look at the top few rows here:
df = pd.read_csv('ab_data.csv')
df.head()
# b. Use the cell below to find the number of rows in the dataset.
#Number of rows in the dataset.
df.shape[0]
# c. The number of unique users in the dataset.
#Number of unique users in the dataset.
df["user_id"].nunique()
# d. The proportion of users converted.
# + tags=[]
#Proportion of users converted.
obs_converted_rate = df.query("converted == 1")["user_id"].nunique()/ \
df["user_id"].nunique()
print('Proportion of users converted:', obs_converted_rate)
# -
# e. The number of times the `new_page` and `treatment` don't match.
df.loc[((df.group != 'treatment') & (df.landing_page == 'new_page')) | ((df.group == 'treatment') & (df.landing_page == 'old_page'))].shape[0]
# + tags=[]
#Number of times the new_page and treatment don't match
mismatch_count = df.query("group != 'treatment' \
and landing_page == 'new_page' \
or group == 'treatment' \
and landing_page != 'new_page'"
).shape[0]
print('Number of times the new_page and treatment don\'t match:', mismatch_count)
# -
# f. Do any of the rows have missing values?
#Do any rows have missing values
is_missing_values = df.isnull().values.any()
print('Do any rows have missing values:', is_missing_values)
# `2.` For the rows where **treatment** does not match with **new_page** or **control** does not match with **old_page**, we cannot be sure if this row truly received the new or old page. Use **Quiz 2** in the classroom to figure out how we should handle these rows.
#
# a. Now use the answer to the quiz to create a new dataset that meets the specifications from the quiz. Store your new dataframe in **df2**.
#Filter the dataframe where group and landing_page
#are treatment and language_page respectively.
df_tmt_newpage = df.query("group == 'treatment' \
and landing_page == 'new_page'"
)
#Filter the dataframe where group and landing_page
#are control and old_page respectively.
df_cntrl_oldpage = df.query("group == 'control' \
and landing_page == 'old_page'"
)
#Combine above treatment and control dataframes.
#This ensures we have clean data.
df2 = pd.concat([df_cntrl_oldpage, df_tmt_newpage], ignore_index=True)
df2.head()
# Double Check all of the correct rows were removed - this should be 0
invalid_row_count = df2[((df2['group'] == 'treatment') == \
(df2['landing_page'] == 'new_page')) == \
False].shape[0]
print('Invalid Row count:', invalid_row_count)
# `3.` Use **df2** and the cells below to answer questions for **Quiz3** in the classroom.
# a. How many unique **user_id**s are in **df2**?
#Unique users found in our cleaned up dataframe (df2).
unique_users = df2["user_id"].unique().shape[0]
print('Unique users in our cleaned dataframe df2:', unique_users)
# b. There is one **user_id** repeated in **df2**. What is it?
#Retrieve the user_id that was duplicated.
duplicated_user_id = df2[df2.duplicated("user_id")] \
.user_id.to_string(index=False)
print('Duplicate user id:', duplicated_user_id)
# c. What is the row information for the repeat **user_id**?
#Row information for the duplicated user.
df2[df2.duplicated("user_id")]
# d. Remove **one** of the rows with a duplicate **user_id**, but keep your dataframe as **df2**.
#Remove the duplicated user from the dataframe df2
df2.drop(df2[df2.duplicated("user_id")].index, inplace=True)
# `4.` Use **df2** in the cells below to answer the quiz questions related to **Quiz 4** in the classroom.
#
# a. What is the probability of an individual converting regardless of the page they receive?
# +
#Probability of individual converting regardless of the page they receive
prob_converted = df2.query("converted == 1")['user_id'].unique().shape[0] \
/df2["user_id"].unique().shape[0]
print('Probability of individual converted regardless of page:', prob_converted)
# -
# b. Given that an individual was in the `control` group, what is the probability they converted?
#Probability of control group individuals who converted.
cntrlgrp_converted_prob = df2.query("group == 'control' and converted == 1")['user_id'].unique().shape[0] \
/df2.query("group == 'control'")['user_id'].unique().shape[0]
print('Probability of control group individuals who converted:',cntrlgrp_converted_prob)
# c. Given that an individual was in the `treatment` group, what is the probability they converted?
#Probability of treatment group individuals who converted.
trmtgrp_converted_prob = df2.query("group == 'treatment' and converted == 1")['user_id'].unique().shape[0] \
/df2.query("group == 'treatment'")['user_id'].unique().shape[0]
print('Probability of treatment group individuals who converted:', trmtgrp_converted_prob)
# d. What is the probability that an individual received the new page?
#Probability that an individual received new page
newpage_prob = df2.query("landing_page == 'new_page'")['user_id'].unique().shape[0] \
/df["user_id"].unique().shape[0]
print('Probability that an individual received new page:', newpage_prob)
# e. Consider your results from parts (a) through (d) above, and explain below whether you think there is sufficient evidence to conclude that the new treatment page leads to more conversions.
# **Your answer goes here.**
# <br>
# <br>
#
# It is unlikely that the new page (treatment group) leads to more conversion, given that **12%** percent of individuals who received **old page (control group)** were converted, when compared to **11.8%** of individuals receiving **new page** were converted. With 0.2% percent difference in conversion, the older page is better then other.
# <br>
# <br>
# For us to support this observation, it requires a quantifying answers to questions about samples of data.
# <br>
# <br>
# Using statistical methods such as **Hypothesis testing** where we investigate a hypothesis. The result of this test allows us to interpert whether our assupmtion holds or whether the assumption has been violated.
# <br>
# <br>
#
#
# <a id='ab_test'></a>
# ### Part II - A/B Test
#
# Notice that because of the time stamp associated with each event, you could technically run a hypothesis test continuously as each observation was observed.
#
# However, then the hard question is do you stop as soon as one page is considered significantly better than another or does it need to happen consistently for a certain amount of time? How long do you run to render a decision that neither page is better than another?
#
# These questions are the difficult parts associated with A/B tests in general.
#
#
# `1.` For now, consider you need to make the decision just based on all the data provided. If you want to assume that the old page is better unless the new page proves to be definitely better at a Type I error rate of 5%, what should your null and alternative hypotheses be? You can state your hypothesis in terms of words or in terms of **$p_{old}$** and **$p_{new}$**, which are the converted rates for the old and new pages.
# **Put your answer here.**
# `2.` Assume under the null hypothesis, $p_{new}$ and $p_{old}$ both have "true" success rates equal to the **converted** success rate regardless of page - that is $p_{new}$ and $p_{old}$ are equal. Furthermore, assume they are equal to the **converted** rate in **ab_data.csv** regardless of the page. <br><br>
#
# Use a sample size for each page equal to the ones in **ab_data.csv**. <br><br>
#
# Perform the sampling distribution for the difference in **converted** between the two pages over 10,000 iterations of calculating an estimate from the null. <br><br>
#
# Use the cells below to provide the necessary parts of this simulation. If this doesn't make complete sense right now, don't worry - you are going to work through the problems below to complete this problem. You can use **Quiz 5** in the classroom to make sure you are on the right track.<br><br>
# a. What is the **conversion rate** for $p_{new}$ under the null?
#Conversion rate under the null Pnew
pnew = df2.query("converted == 1").user_id.nunique() /df2.user_id.nunique()
print('Conversion rate under the null hypothesis (Pnew):',pnew)
# b. What is the **conversion rate** for $p_{old}$ under the null? <br><br>
#We assume under the null hypothesis, both Pnew and Pold have equal success rate.
pold = pnew
print('Conversion rate under the null hypothesis (Pold):',pold)
# c. What is $n_{new}$, the number of individuals in the treatment group?
#Number of individuals in the treatment group.
nnew = df2.query("group == 'treatment'")['converted'].shape[0]
print('Number of individuals in the treatment group:', nnew)
# d. What is $n_{old}$, the number of individuals in the control group?
#Number of individuals in the control group
nold = df2.query("group == 'control'")['converted'].shape[0]
print('Number of individuals in the control group',nold)
#
# e. Simulate $n_{new}$ transactions with a conversion rate of $p_{new}$ under the null. Store these $n_{new}$ 1's and 0's in **new_page_converted**.
#Simulate nnew transactions with a conversion rate of pnew
new_page_converted = np.random.choice([0,1] , nnew, [1-pnew, pnew])
new_page_converted
# f. Simulate $n_{old}$ transactions with a conversion rate of $p_{old}$ under the null. Store these $n_{old}$ 1's and 0's in **old_page_converted**.
old_page_converted = np.random.choice([0,1] , nold, [1-pold, pold])
old_page_converted
# g. Find $p_{new}$ - $p_{old}$ for your simulated values from part (e) and (f).
# +
pnew_rate = np.mean(new_page_converted)
pold_rate = np.mean(old_page_converted)
base_line_converted = (pnew_rate - pold_rate)
print(base_line_converted)
# -
df2_newpage = df2.query("landing_page == 'new_page'")
df_oldpage = df2.query("landing_page == 'old_page'")
np.mean(df2_newpage[['converted']] == 0)['converted']
np.mean(df2_newpage[['converted']] == 1)['converted']
np.mean(df2_newpage.converted)
np.mean(df_oldpage[['converted']] == 1)['converted']
np.mean(df_oldpage[['converted']] == 0)['converted']
np.mean(df2[['converted']]==0)['converted']
# h. Create 10,000 $p_{new}$ - $p_{old}$ values using the same simulation process you used in parts (a) through (g) above. Store all 10,000 values in a NumPy array called **p_diffs**.
# +
# p_diffs = []
# for _ in range(10000):
# df_newsim = df2_newpage.sample(nnew, replace=True)
# df_oldsim = df_oldpage.sample(nold, replace=True)
# pnew_diff = np.mean(df_newsim.converted)
# pold_diff = np.mean(df_oldsim.converted)
# known_diff = pnew_diff - pold_diff
# p_diffs.append(known_diff)
# -
# i. Plot a histogram of the **p_diffs**. Does this plot look like what you expected? Use the matching problem in the classroom to assure you fully understand what was computed here.
# +
#plt.hist(p_diffs);
# -
# j. What proportion of the **p_diffs** are greater than the actual difference observed in **ab_data.csv**?
# +
# sim_diffs = np.array(p_diffs)
# null_values = np.random.normal(0, sim_diffs.std(), sim_diffs.size)
# +
#plt.hist(null_values)
#plt.axvline(x=base_line_converted, color='red');
#plt.axvline(x=np.mean(null_values), color='black');
# +
#(null_values > base_line_converted).mean()
# -
# k. Please explain using the vocabulary you've learned in this course what you just computed in part **j.** What is this value called in scientific studies? What does this value mean in terms of whether or not there is a difference between the new and old pages?
# **Put your answer here.**
# l. We could also use a built-in to achieve similar results. Though using the built-in might be easier to code, the above portions are a walkthrough of the ideas that are critical to correctly thinking about statistical significance. Fill in the below to calculate the number of conversions for each page, as well as the number of individuals who received each page. Let `n_old` and `n_new` refer the the number of rows associated with the old page and new pages, respectively.
# +
import statsmodels.api as sm
convert_old = df2.query("converted == 1 and landing_page == 'old_page'").shape[0]
convert_new = df2.query("converted == 1 and landing_page == 'new_page'").shape[0]
n_old = df2.query("landing_page == 'old_page'").shape[0]
n_new = df2.query("landing_page == 'new_page'").shape[0]
# -
# m. Now use `stats.proportions_ztest` to compute your test statistic and p-value. [Here](https://docs.w3cub.com/statsmodels/generated/statsmodels.stats.proportion.proportions_ztest/) is a helpful link on using the built in.
# +
n = n_old + n_new
pnull = obs_converted_rate
phat = pnew
significance = 0.05
#sm.stats.proportions_ztest(phat*n, n, pnull, alternative='larger')
#sm.stats.proportions_ztest([convert_old, convert_new], [n_old, n_new], alternative='larger')
stat, p_value = sm.stats.proportions_ztest([convert_old, convert_new], [n_old, n_new], pnull, alternative='larger')
print('z_stat: %0.3f, p_value: %0.3f' % (stat, p_value))
if p_value > significance:
print ("Fail to reject the null hypothesis - we have nothing else to say")
else:
print ("Reject the null hypothesis - suggest the alternative hypothesis is true")
# -
# n. What do the z-score and p-value you computed in the previous question mean for the conversion rates of the old and new pages? Do they agree with the findings in parts **j.** and **k.**?
# **Put your answer here.**
# <a id='regression'></a>
# ### Part III - A regression approach
#
# `1.` In this final part, you will see that the result you achieved in the A/B test in Part II above can also be achieved by performing regression.<br><br>
#
# a. Since each row is either a conversion or no conversion, what type of regression should you be performing in this case?
# **Put your answer here.**
# b. The goal is to use **statsmodels** to fit the regression model you specified in part **a.** to see if there is a significant difference in conversion based on which page a customer receives. However, you first need to create in df2 a column for the intercept, and create a dummy variable column for which page each user received. Add an **intercept** column, as well as an **ab_page** column, which is 1 when an individual receives the **treatment** and 0 if **control**.
df2.head()
df2['intercept'] = 1
#df2[['not_converted', 'converted']] = pd.get_dummies(df2['converted'])
df2[['not_converted', 'convert_dummy']] = pd.get_dummies(df2['converted'])
df2[['a_page', 'b_page']] = pd.get_dummies(df2['group'])
df2.head()
# +
#df2 = df2.drop('not_converted', axis=1)
# -
# c. Use **statsmodels** to instantiate your regression model on the two columns you created in part b., then fit the model using the two columns you created in part **b.** to predict whether or not an individual converts.
df2.head()
# +
#df2 = df2.drop('a_page', axis=1)
# +
from scipy import stats
stats.chisqprob = lambda chisq, df_new: stats.chi2.sf(chisq, df_new)
logit_mod = sm.Logit(df2['convert_dummy'], df2[['intercept', 'b_page']])
results = logit_mod.fit()
results.summary()
# -
#Control page - exponent
np.exp(0.0150)
#Treatment page - exponent
1/np.exp(-0.0150)
# d. Provide the summary of your model below, and use it as necessary to answer the following questions.
# e. What is the p-value associated with **ab_page**? Why does it differ from the value you found in **Part II**?<br><br> **Hint**: What are the null and alternative hypotheses associated with your regression model, and how do they compare to the null and alternative hypotheses in **Part II**?
# **Put your answer here.**
# f. Now, you are considering other things that might influence whether or not an individual converts. Discuss why it is a good idea to consider other factors to add into your regression model. Are there any disadvantages to adding additional terms into your regression model?
# **Put your answer here.**
# g. Now along with testing if the conversion rate changes for different pages, also add an effect based on which country a user lives in. You will need to read in the **countries.csv** dataset and merge together your datasets on the appropriate rows. [Here](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.join.html) are the docs for joining tables.
#
# Does it appear that country had an impact on conversion? Don't forget to create dummy variables for these country columns - **Hint: You will need two columns for the three dummy variables.** Provide the statistical output as well as a written response to answer this question.
countries_df = pd.read_csv('countries.csv')
countries_df.head()
countries_df.info()
df2.info()
df2.head()
countries_df.query("user_id == 851104")
df2.head()
df2 = df2.merge(countries_df, how='inner', on='user_id', validate='1:1')
df2.head()
df2['country'].unique()
df2[['CA', 'UK', 'USA']] = pd.get_dummies(df2['country'])
df2.head()
# +
from scipy import stats
stats.chisqprob = lambda chisq, df_new: stats.chi2.sf(chisq, df_new)
logit_mod = sm.Logit(df2['convert_dummy'], df2[['intercept', 'b_page', 'USA', 'UK']])
results = logit_mod.fit()
results.summary()
# -
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import precision_score, recall_score, accuracy_score, confusion_matrix,f1_score
# +
y = df2['convert_dummy']
X = df2[['a_page', 'b_page', 'CA', 'UK', 'USA']]
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.20, random_state=27)
# -
log_mod = LogisticRegression(solver='liblinear')
log_mod.fit(X_train, y_train)
y_pred = log_mod.predict(X_test)
print(precision_score(y_test, y_pred))
print(recall_score(y_test, y_pred))
print(accuracy_score(y_test, y_pred))
print(f1_score(y_test, y_pred))
confusion_matrix(y_test, y_pred)
# +
from sklearn.ensemble import RandomForestClassifier
rfc = RandomForestClassifier(n_estimators=20).fit(X_train, y_train)
rfc_pred = rfc.predict(X_test)
print(accuracy_score(y_test, rfc_pred))
print(f1_score(y_test, rfc_pred))
print(recall_score(y_test, rfc_pred))
print(precision_score(y_test, rfc_pred))
# -
(y_pred == 0).sum()
(y_pred == 1).sum()
(y_train == 0).sum()
(y_train == 1).sum()
(y_test == 0).sum()
(y_test == 1).sum()
y_pred.size
from sklearn.metrics import classification_report
target_names =['not_converted', 'converted']
print(classification_report(y_test, y_pred, target_names=target_names))
from sklearn.metrics import accuracy_score
print(accuracy_score(y_test, y_pred))
corr = df2.corr(method='pearson')
import seaborn as sns
sns.heatmap(corr)
# +
mask = np.zeros_like(corr, dtype=np.bool)
mask[np.triu_indices_from(mask)] = True
# Colors
cmap = sns.diverging_palette(240, 10, as_cmap=True)
# Plotting the heatmap
sns.heatmap(corr, mask=mask, linewidths=.5, cmap=cmap, center=0)
# -
# h. Though you have now looked at the individual factors of country and page on conversion, we would now like to look at an interaction between page and country to see if there significant effects on conversion. Create the necessary additional columns, and fit the new model.
#
# Provide the summary results, and your conclusions based on the results.
# <a id='conclusions'></a>
# ## Finishing Up
#
# > Congratulations! You have reached the end of the A/B Test Results project! You should be very proud of all you have accomplished!
#
# > **Tip**: Once you are satisfied with your work here, check over your report to make sure that it is satisfies all the areas of the rubric (found on the project submission page at the end of the lesson). You should also probably remove all of the "Tips" like this one so that the presentation is as polished as possible.
#
#
# ## Directions to Submit
#
# > Before you submit your project, you need to create a .html or .pdf version of this notebook in the workspace here. To do that, run the code cell below. If it worked correctly, you should get a return code of 0, and you should see the generated .html file in the workspace directory (click on the orange Jupyter icon in the upper left).
#
# > Alternatively, you can download this report as .html via the **File** > **Download as** submenu, and then manually upload it into the workspace directory by clicking on the orange Jupyter icon in the upper left, then using the Upload button.
#
# > Once you've done this, you can submit your project by clicking on the "Submit Project" button in the lower right here. This will create and submit a zip file with this .ipynb doc and the .html or .pdf version you created. Congratulations!
from subprocess import call
call(['python', '-m', 'nbconvert', 'Analyze_ab_test_results_notebook.ipynb'])
| old-Analyze_ab_test_results_notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
df = pd.read_csv(r"C:\Users\<NAME>\Desktop\ML EXAMPLE\Modular\ML_Live_Class\data\Advertising.csv")
df.head()
X = df.drop('sales', axis=1)
X.head()
y = df['sales']
y
from sklearn.preprocessing import PolynomialFeatures
from sklearn.linear_model import LinearRegression
poly_conv = PolynomialFeatures(degree=3, include_bias=False)
final_model = LinearRegression()
poly_features = poly_conv.fit_transform(X)
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(
poly_features, y, test_size=0.33, random_state=101)
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler() #it applies z score normalisation(standardisation)
scaler.fit(X_train)
scaled_X_train = scaler.transform(X_train)
scaled_X_test = scaler.transform(X_test)
from sklearn.linear_model import RidgeCV
ridge_cv_model = RidgeCV(alphas=(0.1, 1.0, 10.0), scoring='neg_mean_absolute_error')
ridge_cv_model.fit(scaled_X_train,y_train)
ridge_cv_model.alpha_ #best alpha hyperparameter
test_predictions = ridge_cv_model.predict(scaled_X_test)
from sklearn.metrics import mean_absolute_error, mean_squared_error
MAE = mean_absolute_error(y_test, test_predictions)
MAE
RMSE = np.sqrt(mean_squared_error(y_test, test_predictions))
RMSE
ridge_cv_model.coef_
ridge_cv_model.best_score_ #returns the highest negated mean absolute error
from sklearn.linear_model import LassoCV
help(LassoCV)
lasso_cv_model = LassoCV(eps=0.001, n_alphas=100, cv=10, max_iter=10000000)
lasso_cv_model.fit(scaled_X_train,y_train)
lasso_cv_model.alpha_
test_predictions = lasso_cv_model.predict(scaled_X_test)
MAE = mean_absolute_error(y_test, test_predictions)
MAE
RMSE = np.sqrt(mean_squared_error(y_test, test_predictions))
RMSE
from sklearn.linear_model import ElasticNetCV
help(ElasticNetCV)
elastic_model = ElasticNetCV(l1_ratio=[.1, .5, .7,.9, .95, .99, 1], eps=0.001, n_alphas=100, max_iter=10000000)
elastic_model.fit(scaled_X_train, y_train)
elastic_model.l1_ratio_
test_predictions = elastic_model.predict(scaled_X_test)
MAE = mean_absolute_error(y_test, test_predictions)
MAE
RMSE = np.sqrt(mean_squared_error(y_test, test_predictions))
RMSE
lasso_cv_model.coef_
| RidgeCV/Ridge_Lasso_CV.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Practica 1B
# <NAME>\
# <NAME>
# ### Ejercicio 1
#
# Un grupo de 5 personas quiere cruzar un viejo y estrecho puente. Es una noche cerrada y se necesita llevar una linterna para cruzar. El grupo solo dispone de una linterna, a la que le quedan 5 minutos de batería.
#
# 1. Cada persona tarda en cruzar 10, 30, 60, 80 y 120 segundos, respectivamente.
# 2. El puente solo resiste un máximo de 2 personas cruzando a la vez, y cuando cruzan dos personas juntas, caminan a la velocidad del más lento.
# 3. No se puede lanzar la linterna de un extremo a otro del puente, así que cada vez que crucen dos personas, alguien tiene que volver a cruzar hacia atrás con la linterna a buscar a los compañeros que falten, y así hasta que hayan cruzado todos
class Problem(object):
"""The abstract class for a formal problem. You should subclass
this and implement the methods actions and result, and possibly
__init__, goal_test, and path_cost. Then you will create instances
of your subclass and solve them with the various search functions."""
def __init__(self, initial, goal=None):
"""The constructor specifies the initial state, and possibly a goal
state, if there is a unique goal. Your subclass's constructor can add
other arguments."""
self.initial = initial
self.goal = goal
def actions(self, state):
"""Return the actions that can be executed in the given
state. The result would typically be a list, but if there are
many actions, consider yielding them one at a time in an
iterator, rather than building them all at once."""
raise NotImplementedError
def result(self, state, action):
"""Return the state that results from executing the given
action in the given state. The action must be one of
self.actions(state)."""
raise NotImplementedError
def goal_test(self, state):
"""Return True if the state is a goal. The default method compares the
state to self.goal or checks for state in self.goal if it is a
list, as specified in the constructor. Override this method if
checking against a single self.goal is not enough."""
if isinstance(self.goal, list):
return is_in(state, self.goal)
else:
return state == self.goal
def path_cost(self, c, state1, action, state2):
"""Return the cost of a solution path that arrives at state2 from
state1 via action, assuming cost c to get up to state1. If the problem
is such that the path doesn't matter, this function will only look at
state2. If the path does matter, it will consider c and maybe state1
and action. The default method costs 1 for every step in the path."""
return c + 1
def value(self, state):
"""For optimization problems, each state has a value. Hill-climbing
and related algorithms try to maximize this value."""
raise NotImplementedError
def coste_de_aplicar_accion(self, estado, accion):
"""Hemos incluido está función que devuelve el coste de un único operador (aplicar accion a estado). Por defecto, este
coste es 1. Reimplementar si el problema define otro coste """
return 1
class elPuente(Problem):
def __init__(self, initial, goal = None):
'''Inicializacion de nuestro problema.'''
Problem.__init__(self, (0, initial, (), 0), goal)
self._actions = [(10,"10"),(30,"30"),(60,"60"),(80,"80"),(120,"120"),(30,"10","30"),(60,"10","60"),(80,"10","80"),(120,"10","120"),(60,"30","60"),(80,"30","80"),(120,"30","120"),(80,"60","80"),(120,"60","120"),(120,"80","120")]
def actions(self, state):
'''Devuelve las acciones validas para un estado.'''
t = 0
for i in state[1]: t+= i
for i in state[2]: t+= i
ret = list()
#Recorre todas las acciones posibles
for act in self._actions:
i = 1
moves = list()
actTime = 0
#Controla que de tiempo a cruzar la barca
while(i in range(len(act))):
if(int(act[i]) > actTime): actTime = int(act[i])
moves.append(int(act[i]))
i+=1
#Si supera el tiempo no hay posibilidad de seguir
if((state[0] + actTime) < t):
insert = True
#Comprueba que las dos personas estén en el mismo lado del río
for j in range(len(moves)):
if ((int(act[j]) in state[1] and state[3] == 0) or (int(act[j]) in state[2] and state[3] == 1)):
insert = insert and True
else: insert = insert and False
j+=1
#Si cumple todas las condiciones se da como transición valida
if (insert): ret.append(act)
return ret
def result(self, state, act):
'''Devuelve el estado resultante de aplicar una accion a un estado determinado.'''
left = list(state[1])
right = list(state[2])
t = state[0] + act[0]
i = 1
#Para todo el estado actual ejecuta el camino de ida y el de vuelta
while(i in range(len(act))):
aux = int(act[i])
if state[3] == 0: #Ida
left.remove(aux)
right.append(aux)
else: #Vuelta
right.remove(aux)
left.append(aux)
i+=1
turno = (state[3] + 1) % 2
return (t, tuple(left), tuple(right), turno)
def goal_test(self,estado):
'''Devuelve si el estado actual es solución'''
return (estado[0] <= 300) and (len(estado[1]) == 0) and (estado[3] == 1)
puente.initial
puente = elPuente((10,30,60,80,120))
puente.actions(puente.initial)
puente.result(puente.initial, (120,"10","120"))
puente.goal_test(puente.initial)
puente.goal_test((300, (), (10,30,60,80,120), 1))
puente.actions(120,(30,60,80),(10.120),1)
from search import *
from search import breadth_first_tree_search, depth_first_tree_search, depth_first_graph_search, breadth_first_graph_search
# Para la ejecución se ha optado por la busqueda primero en anchura ya que la más eficiente dentro de las busquedas que no requieren de heurística. El tiempo de ejecución es ligeramente superior a otros algoritmos estudiados en clase y, aún así, son resultados muy buenos. Como podemos ver, se han adjuntado tanto los tiempo como las ejecuciones de todos los algorítmos comprobados.
breadth_first_tree_search(elPuente((10,30,60,80,120))).solution()
# 568 ms ± 2.42 ms per loop (mean ± std. dev. of 7 runs, 1 loop each)
depth_first_tree_search(elPuente((10,30,60,80,120))).solution()
# 998 ms ± 6.44 ms per loop (mean ± std. dev. of 7 runs, 1 loop each)
#
uniform_cost_search(elPuente((10,30,60,80,120))).solution()
# 2.92 s ± 26.3 ms per loop (mean ± std. dev. of 7 runs, 1 loop each)
# +
# Hacemos una definición ampliada de la clase Problem de AIMA que nos va a permitir experimentar con distintos
# estados iniciales, algoritmos y heurísticas, para resolver el 8-puzzle.
# The solvability of a configuration can be checked by calculating the Inversion Permutation. If the total Inversion Permutation is even then the initial configuration is solvable else the initial configuration is not solvable which means that only 9!/2 initial states lead to a solution.
# Añadimos en la clase ampliada la capacidad para contar el número de nodos analizados durante la
# búsqueda:
class Problema_con_Analizados(Problem):
"""Es un problema que se comporta exactamente igual que el que recibe al
inicializarse, y además incorpora unos atributos nuevos para almacenar el
número de nodos analizados durante la búsqueda. De esta manera, no
tenemos que modificar el código del algoritmo de búsqueda."""
def __init__(self, problem):
self.initial = problem.initial
self.problem = problem
self.analizados = 0
self.goal = problem.goal
def actions(self, estado):
return self.problem.actions(estado)
def result(self, estado, accion):
return self.problem.result(estado, accion)
def goal_test(self, estado):
self.analizados += 1
return self.problem.goal_test(estado)
def coste_de_aplicar_accion(self, estado, accion):
return self.problem.coste_de_aplicar_accion(estado,accion)
def check_solvability(self, state):
""" Checks if the given state is solvable """
inversion = 0
for i in range(len(state)):
for j in range(i+1, len(state)):
if (state[i] > state[j]) and state[i] != 0 and state[j]!= 0:
inversion += 1
return inversion % 2 == 0
# -
def resuelve_puente_puzzle(estado_inicial, algoritmo, h=None):
p = Problema_con_Analizados(elPuente(estado_inicial))
if p.check_solvability(estado_inicial):
if h:
sol = algoritmo(p,h).solution()
else:
sol = algoritmo(p).solution()
print("Solución: {0}".format(sol))
print("Algoritmo: {0}".format(algoritmo.__name__))
if h:
print("Heurística: {0}".format(h.__name__))
else:
pass
print("Longitud de la solución: {0}. Nodos analizados: {1}".format(len(sol),p.analizados))
else:
print("Este problema no tiene solucion. ")
E1 = (10,30,60,80,120)
resuelve_puente_puzzle(E1,breadth_first_tree_search)
| Practicas/Practica 1/P1_B_Ejercicio1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import bw2io as bi
import bw2data as bd
import sys
sys.path.append("/Users/akim/PycharmProjects/bigfood_db")
# Local files
from bigfood_db.import_databases import import_ecoinvent
from bigfood_db.importers import Agribalyse13Importer
from bigfood_db.utils import create_location_mapping_glo, create_location_mapping_rer, modify_exchanges
if __name__ == '__main__':
project = "Food ei38"
bd.projects.set_current(project)
ei_path = "/Users/akim/Documents/LCA_files/ecoinvent_38_cutoff/datasets"
ei_name = "ecoinvent 3.8 cutoff"
bi.bw2setup()
b3 = bd.Database('biosphere3')
import_ecoinvent(ei_path, ei_name)
ei = bd.Database(ei_name)
ag_path = "/Users/akim/Documents/LCA_files/agribalyse_13/Agribalyse CSV FINAL_no links_Nov2016v3.CSV"
ag_name = "Agribalyse 1.3"
ag = Agribalyse13Importer(ag_path, ag_name)
# ag.add_unlinked_activities()
# mapping = create_location_mapping(ag, ei_name)
# ag = modify_exchanges(ag, mapping, ei_name)
# -
ag.statistics()
# # Technosphere
# ## 1. Allocation for activities with RER locations
mapping_rer = create_location_mapping_rer(ag, ei_name)
# Add nitric acid manually
nitric_acid_rer_wo_ru = [
act for act in ei
if "nitric acid production" in act['name']
and "RER w/o" in act['location']
and "nitric acid, without water, in 50% solution state" in act['reference product']
]
assert len(nitric_acid_rer_wo_ru)==1
nitric_acid_ru = [
act for act in ei
if "nitric acid production" in act['name']
and "RU" == act['location']
and "nitric acid, without water, in 50% solution state" in act['reference product']
]
assert len(nitric_acid_ru)==1
mapping_rer.append(
{
("nitric acid production, product in 50% solution state", "RER"): [
(ei.name, nitric_acid_rer_wo_ru[0]['code']),
(ei.name, nitric_acid_ru[0]['code']),
]
}
)
mapping_rer
ag = modify_exchanges(ag, mapping_rer, ei_name)
ag.match_database(ei_name, fields=['name', 'location', 'unit', 'reference product'])
# ag.match_database(ei_name, fields=['name', 'location', 'unit'])
ag.statistics()
# ## 2. Allocation for activities with GLO locations
# %%time
mapping_glo = create_location_mapping_glo(ag, ei_name)
mapping_glo
# +
ag = modify_exchanges(ag, mapping_glo, ei_name)
ag.match_database(ei_name, fields=['name', 'location', 'unit'])
ag.statistics()
# -
# ## 3. Still unlinked
[(act['name'], act.get('location', None)) for act in ag.unlinked if "tech" in act['type']]
# # Biosphere
# +
# del bd.databases["Agribalyse biosphere"]
# bd.Database("Agribalyse biosphere").register()
# ag.add_unlinked_flows_to_biosphere_database("Agribalyse biosphere")
# -
# +
| dev/ei38_imports.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Compare $\nu_\textrm{max}$
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import sys
sys.path.insert(0, '../')
from shocksgo import generate_stellar_fluxes, generate_solar_fluxes, power_spectrum
# +
from glob import glob
paths = glob('validation_peaks/*')
print(len(paths))
observed = []
simulated = []
for path in paths:
o, s = np.load(path)
observed.append(o)
simulated.append(s)
plt.figure(figsize=(3, 3))
plt.plot(range(900, 3500, 100), range(900, 3500, 100), ls='--', color='gray')
plt.plot(observed, simulated, 'k.')
plt.xlabel('$\\nu_{\\rm max, obs}$', fontsize=15)
plt.ylabel('$\\nu_{\\rm max, sim}$', fontsize=15)
for s in 'right,top'.split(','):
plt.gca().spines[s].set_visible(False)
plt.savefig('paper_plots/nu_comparison.pdf', bbox_inches='tight')
# -
# # Generate simulated transits
# +
from astropy.constants import M_sun, R_sun, L_sun
import astropy.units as u
sample_times, sample_fluxes, kernel = generate_solar_fluxes(duration=1000*u.day)
sample_fluxes += np.random.normal(loc=0, scale=4e-5, size=len(sample_fluxes))
plt.plot(sample_times.to(u.day).value, sample_fluxes)
plt.xlabel('Time [days]')
plt.ylabel('Relative flux')
# +
freq, power = power_spectrum(sample_fluxes)
plt.loglog(freq * 1e6, power, ',')
plt.loglog(freq * 1e6, kernel.get_psd(2 * np.pi * freq) / 2 / np.pi, alpha=0.5)
plt.xlim([1e-1, 1e4])
plt.ylim([1e-11, 1e-4])
plt.xlabel('Freq [$\mu$Hz]')
plt.ylabel('Power [flux$^2$/Hz]')
# +
from astropy.constants import M_sun, R_sun, L_sun, R_earth
from batman import TransitModel, TransitParams
params = TransitParams()
params.per = 365.25
params.t0 = 1
params.inc = 90
params.w = 90
params.ecc = 0
params.u = [0.4, 0.2]
params.limb_dark = 'quadratic'
params.a = float(1*u.AU/R_sun)
params.rp = float(R_earth/R_sun)
fig, ax = plt.subplots(1, 4, figsize=(14, 4), sharey=True)
for i in range(4):
# times, fluxes, kernel = generate_stellar_fluxes(duration=2*u.day, M=1.0 * M_sun,
# T_eff=5777 * u.K,
# R=1.0 * R_sun, L=L_sun)
times, fluxes, kernel = generate_solar_fluxes(duration=2*u.day)
# add white noise
fluxes += np.random.normal(loc=0, scale=4e-5, size=len(fluxes))
print(fluxes.std())
fluxes += 1
model = TransitModel(params, times.to(u.day).value)
lc = model.light_curve(params)
# ax[i].plot(times.to(u.day).value, fluxes)
t = times.to(u.day).value - times.to(u.day).value.mean()
ax[i].scatter(t, lc*fluxes, marker='.', color='k', rasterized=True)
ax[i].set_ylim([0.9997, 1.0003])
ax[i].plot(t, lc, 'r')
ax[i].set_xlim([-1, 1])
ax[i].ticklabel_format(useOffset=False)
ax[i].set_xlabel('Time [days]')
np.save('simulated_transits/shocksgo_{0:02d}.npy'.format(i), np.vstack([t, lc*fluxes]))
for s in ['right', 'top']:
ax[i].spines[s].set_visible(False)
ax[0].set_ylabel('Relative flux')
fig.savefig('paper_plots/transits_shocksgo.pdf', bbox_inches='tight')
# +
from astropy.io import fits
data = fits.getdata('../data/VIRGO_1min_0083-7404.fits.gz', cache=False)
# +
virgo_fluxes = data[data != -99]/np.median(data[data != -99])
virgo_times = np.arange(len(data))[data != -99] / 60 / 24
plt.plot(virgo_times, virgo_fluxes)
# -
np.argmin(np.abs(virgo_times - 4500))
# +
fig, ax = plt.subplots(1, 4, figsize=(14, 4), sharey=True)
one_day = 24 * 60
offset = 0 # 5836000# 6249000 #1000
for i in range(0, 8, 2):
times = virgo_times[offset+one_day*(i+1) : offset+one_day*(i+3)].copy()
fluxes = virgo_fluxes[offset+one_day*(i+1) : offset+one_day*(i+3)].copy()
params.t0 = times.mean()
fluxes /= np.median(fluxes)
print(fluxes.std())
model = TransitModel(params, times)
lc = model.light_curve(params)
t = times - times.mean()
ax[i//2].scatter(t, lc*fluxes, marker='.', color='k', rasterized=True)
ax[i//2].set_ylim([0.9997, 1.0003])
ax[i//2].plot(t, lc, 'r')
ax[i//2].ticklabel_format(useOffset=False)
ax[i//2].set_xlabel('Time [days]')
for s in ['right', 'top']:
ax[i//2].spines[s].set_visible(False)
np.save('simulated_transits/virgo_{0:02d}.npy'.format(i//2), np.vstack([t, lc*fluxes]))
ax[0].set_ylabel('Relative flux')
fig.savefig('paper_plots/transits_virgo.pdf', bbox_inches='tight')
# -
# # Fit transits
# +
from glob import glob
import emcee
from copy import deepcopy
import batman
import celerite
from celerite import terms, modeling
virgo_transit_times = []
virgo_transit_fluxes = []
paths = glob('simulated_transits/virgo_??.npy')
for i, path in enumerate(paths):
t, f = np.load(path)
t += 365.25 * i
virgo_transit_times.append(t)
virgo_transit_fluxes.append(f)
virgo_transit_times = np.concatenate(virgo_transit_times)
virgo_transit_fluxes = np.concatenate(virgo_transit_fluxes)
# Set up batman+celerite model
def transit_model(p):
t0, rp = p[2:]
trial_params = deepcopy(params)
trial_params.t0 = t0
trial_params.rp = rp
m = batman.TransitModel(trial_params, virgo_transit_times)
return m.light_curve(trial_params)
class MeanModel(modeling.Model):
parameter_names = ("t0", "rp")
def get_value(self, t):
trial_params = deepcopy(params)
trial_params.t0 = self.t0
trial_params.rp = self.rp
m = batman.TransitModel(trial_params, t)
return m.light_curve(trial_params)
def log_probability(params):
gp.set_parameter_vector(params)
lp = gp.log_prior()
if not np.isfinite(lp):
return -np.inf
return gp.log_likelihood(virgo_transit_fluxes) + lp
kernel = terms.SHOTerm(log_omega0=0, log_S0=0, log_Q=np.log(1/np.sqrt(2)))
kernel.freeze_parameter('log_Q')
mean = MeanModel(t0=0, rp=float(R_earth/R_sun), bounds=dict(t0=(-0.5, 0.5), rp=(0, 1)))
gp = celerite.GP(kernel, mean=mean, fit_mean=True)
gp.compute(virgo_transit_times, 5e-5)
nwalkers, ndim = 20, 4
sampler = emcee.EnsembleSampler(nwalkers, ndim, log_probability, threads=8)
p0 = []
while len(p0) < nwalkers:
p = np.array([0, 0, 0, float(R_earth/R_sun)]) + 1e-5*np.random.randn(4)
# if np.isfinite(lnlike(p)):
p0.append(p)
p1 = sampler.run_mcmc(p0, 5000)[0]
sampler.reset()
sampler.run_mcmc(p1, 2000)
sampler.pool.close()
# -
from corner import corner
corner(sampler.flatchain, labels=gp.get_parameter_names(),
truths=[None, None, 0, float(R_earth/R_sun)]);
# +
# # Plot the data.
# plt.plot(virgo_transit_times, virgo_transit_fluxes, marker=".", color='k')
# # Plot 24 posterior samples.
# samples = sampler.flatchain
# for s in samples[np.random.randint(len(samples), size=24)]:
# gp.set_parameter_vector(s)
# mu = gp.predict(virgo_transit_fluxes, virgo_transit_times, return_cov=False)
# plt.plot(virgo_transit_times, mu, color='r', alpha=0.1)
# plt.ylabel(r"$y$")
# plt.xlabel(r"$t$")
# plt.xlim(-2, 2)
# plt.gca().yaxis.set_major_locator(plt.MaxNLocator(5))
# plt.title("posterior predictions");
# +
shocksgo_transit_times = []
shocksgo_transit_fluxes = []
paths = glob('simulated_transits/shocksgo_??.npy')
for i, path in enumerate(paths):
t, f = np.load(path)
t += 365.25 * i
shocksgo_transit_times.append(t)
shocksgo_transit_fluxes.append(f)
shocksgo_transit_times = np.concatenate(shocksgo_transit_times)
shocksgo_transit_fluxes = np.concatenate(shocksgo_transit_fluxes)
def log_probability_shocksgo(params):
gp.set_parameter_vector(params)
lp = gp.log_prior()
if not np.isfinite(lp):
return -np.inf
return gp.log_likelihood(shocksgo_transit_fluxes) + lp
kernel = terms.SHOTerm(log_omega0=0, log_S0=0, log_Q=np.log(1/np.sqrt(2)))
kernel.freeze_parameter('log_Q')
mean = MeanModel(t0=0, rp=float(R_earth/R_sun), bounds=dict(t0=(-0.01, 0.01), rp=(0, 1)))
gp = celerite.GP(kernel, mean=mean, fit_mean=True)
gp.compute(virgo_transit_times, 5e-5)
nwalkers, ndim = 20, 4
sampler_shocksgo = emcee.EnsembleSampler(nwalkers, ndim, log_probability_shocksgo, threads=8)
p0 = []
while len(p0) < nwalkers:
p = np.array([0, 0, 0, float(R_earth/R_sun)]) + 1e-5*np.random.randn(4)
p0.append(p)
p1 = sampler_shocksgo.run_mcmc(p0, 5000)[0]
sampler_shocksgo.reset()
sampler_shocksgo.run_mcmc(p1, 2000)
sampler_shocksgo.pool.close()
# -
corner(sampler_shocksgo.flatchain, labels=gp.get_parameter_names(),
truths=[None, None, 0, float(R_earth/R_sun)]);
# +
# Plot the data.
plt.plot(shocksgo_transit_times, shocksgo_transit_fluxes, marker=".", color='k')
# Plot 24 posterior samples.
samples = sampler.flatchain
for s in samples[np.random.randint(len(samples), size=24)]:
gp.set_parameter_vector(s)
mu = gp.predict(shocksgo_transit_fluxes, virgo_transit_times, return_cov=False)
plt.plot(shocksgo_transit_times, mu, color='r', alpha=0.1)
plt.ylabel(r"$y$")
plt.xlabel(r"$t$")
plt.xlim(-2 + 1*365.25, 2 + 1*365.25)
plt.gca().yaxis.set_major_locator(plt.MaxNLocator(5))
plt.title("posterior predictions");
# +
fig, ax = plt.subplots(2, 1, figsize=(3, 5))
ax[0].hist(sampler.flatchain[:, 2], bins=100, histtype='stepfilled', alpha=0.5, label='VIRGO', density=True)
ax[0].hist(sampler_shocksgo.flatchain[:, 2], bins=100, histtype='stepfilled', alpha=0.5, label='simulated', density=True)
ax[0].legend()
ax[0].set_xlabel("$t_0$")
ax[1].hist(sampler.flatchain[:, 3], bins=100, histtype='stepfilled', alpha=0.5, label='VIRGO', density=True)
ax[1].hist(sampler_shocksgo.flatchain[:, 3], bins=100, histtype='stepfilled', alpha=0.5, label='simulated', density=True)
ax[0].axvline(0, ls='--')
ax[1].axvline(float(R_earth/R_sun), ls='--')
# ax[1].legend()
ax[1].set_xlabel("$R_p/R_\star$")
for axis in ax:
for s in ['right', 'top', 'left']:
axis.set_yticks([])
axis.spines[s].set_visible(False)
fig.tight_layout()
fig.savefig('paper_plots/posterior_comparison.pdf', bbox_inches='tight')
# -
(1/(0.1*u.uHz)).to(u.day)
| notebooks/demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/ValentinoVizner/google_Colab/blob/master/Copy_of_lesson4_mnist_sgd.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="gOAL9bNMD6Xj" colab_type="text"
# **Important: This notebook will only work with fastai-0.7.x. Do not try to run any fastai-1.x code from this path in the repository because it will load fastai-0.7.x**
# + [markdown] id="3GYSUMQVD6Xr" colab_type="text"
# ## Using SGD on MNIST
# + [markdown] id="DpBNHi88D6Xw" colab_type="text"
# ## Background
# + [markdown] id="jLXEA33VD6X1" colab_type="text"
# ### ... about machine learning (a reminder from lesson 1)
# + [markdown] id="jZ0QyKDJD6X5" colab_type="text"
# The good news is that modern machine learning can be distilled down to a couple of key techniques that are of very wide applicability. Recent studies have shown that the vast majority of datasets can be best modeled with just two methods:
#
# 1. Ensembles of decision trees (i.e. Random Forests and Gradient Boosting Machines), mainly for structured data (such as you might find in a database table at most companies). We looked at random forests in depth as we analyzed the Blue Book for Bulldozers dataset.
#
# 2. Multi-layered neural networks learnt with SGD (i.e. shallow and/or deep learning), mainly for unstructured data (such as audio, vision, and natural language)
#
# In this lesson, we will start on the 2nd approach (a neural network with SGD) by analyzing the MNIST dataset. You may be surprised to learn that **logistic regression is actually an example of a simple neural net**!
# + [markdown] id="CHuCH19pD6X9" colab_type="text"
# ### About The Data
# + [markdown] id="K8d8S1KvD6YC" colab_type="text"
# In this lesson, we will be working with MNIST, a classic data set of hand-written digits. Solutions to this problem are used by banks to automatically recognize the amounts on checks, and by the postal service to automatically recognize zip codes on mail.
# + [markdown] id="SyV6yCDWD6YG" colab_type="text"
# <img src="https://github.com/fastai/fastai/blob/master/courses/ml1/images/mnist.png?raw=1" alt="" style="width: 60%"/>
# + [markdown] id="W9P8CrqcD6YK" colab_type="text"
# A matrix can represent an image, by creating a grid where each entry corresponds to a different pixel.
#
# <img src="https://github.com/fastai/fastai/blob/master/courses/ml1/images/digit.gif?raw=1" alt="digit" style="width: 55%"/>
# (Source: [<NAME>
# ](https://medium.com/@ageitgey/machine-learning-is-fun-part-3-deep-learning-and-convolutional-neural-networks-f40359318721))
#
# + [markdown] id="eLHa6IVuD6YR" colab_type="text"
# ## Imports and data
# + [markdown] id="Yyg1Ui3nD6YV" colab_type="text"
# We will be using the fastai library, which is still in pre-alpha. If you are accessing this course notebook, you probably already have it downloaded, as it is in the same Github repo as the course materials.
#
# We use [symbolic links](https://kb.iu.edu/d/abbe) (often called *symlinks*) to make it possible to import these from your current directory. For instance, I ran:
#
# ln -s ../../fastai
#
# in the terminal, within the directory I'm working in, `home/fastai/courses/ml1`.
# + id="34RUptuiJOyc" colab_type="code" outputId="c132182a-37e2-4313-8159-d3fa2b7750fa" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# !pip install torchvision==0.1.9
# !pip install fastai==0.7.0
# !pip install torchtext==0.2.3
# + id="whNq_5HAD6Yp" colab_type="code" colab={}
from fastai.imports import *
from fastai.torch_imports import *
from fastai.io import *
# + id="RxBA5eY3D6Y6" colab_type="code" colab={}
# !mkdir -p data/mnist
path = 'data/mnist/'
# + [markdown] id="_oahNjkQD6ZG" colab_type="text"
# Let's download, unzip, and format the data.
# + id="8gJoXZ1XD6ZJ" colab_type="code" colab={}
import os
os.makedirs(path, exist_ok=True)
# + id="HyE-hHv6D6ZP" colab_type="code" colab={}
URL='http://deeplearning.net/data/mnist/'
FILENAME='mnist.pkl.gz'
def load_mnist(filename):
return pickle.load(gzip.open(filename, 'rb'), encoding='latin-1')
# + id="2BNGd-uND6ZW" colab_type="code" colab={}
get_data(URL+FILENAME, path+FILENAME)
((x, y), (x_valid, y_valid), _) = load_mnist(path+FILENAME)
# + id="4P0e-lGtD6Zc" colab_type="code" outputId="b39fdaa2-302d-4397-dab4-274d690ee6ed" colab={}
type(x), x.shape, type(y), y.shape
# + [markdown] id="4p-sbOJ5D6Zj" colab_type="text"
# ### Normalize
# + [markdown] id="Uv_LzKelD6Zk" colab_type="text"
# Many machine learning algorithms behave better when the data is *normalized*, that is when the mean is 0 and the standard deviation is 1. We will subtract off the mean and standard deviation from our training set in order to normalize the data:
# + id="Jf6fadD_D6Zl" colab_type="code" colab={}
mean = x.mean()
std = x.std()
x=(x-mean)/std
mean, std, x.mean(), x.std()
# + [markdown] id="ZH8kOY99D6Zq" colab_type="text"
# Note that for consistency (with the parameters we learn when training), we subtract the mean and standard deviation of our training set from our validation set.
# + id="GYqIKmjkD6Zs" colab_type="code" outputId="483c9cc5-f314-41f4-e70c-f44fd6e1a000" colab={}
x_valid = (x_valid-mean)/std
x_valid.mean(), x_valid.std()
# + [markdown] id="IJc7VuIGD6Zx" colab_type="text"
# ### Look at the data
# + [markdown] id="Nu4GsotqD6Zy" colab_type="text"
# In any sort of data science work, it's important to look at your data, to make sure you understand the format, how it's stored, what type of values it holds, etc. To make it easier to work with, let's reshape it into 2d images from the flattened 1d format.
# + [markdown] id="4O057ZbUD6Zz" colab_type="text"
# #### Helper methods
# + id="Is3xYdGWD6Z0" colab_type="code" colab={}
def show(img, title=None):
plt.imshow(img, cmap="gray")
if title is not None: plt.title(title)
# + id="NwcKJGUtD6Z4" colab_type="code" colab={}
def plots(ims, figsize=(12,6), rows=2, titles=None):
f = plt.figure(figsize=figsize)
cols = len(ims)//rows
for i in range(len(ims)):
sp = f.add_subplot(rows, cols, i+1)
sp.axis('Off')
if titles is not None: sp.set_title(titles[i], fontsize=16)
plt.imshow(ims[i], cmap='gray')
# + [markdown] id="AtoZ3TkSD6Z6" colab_type="text"
# #### Plots
# + id="ekny-ln9D6Z7" colab_type="code" outputId="0a82aae8-4dbc-4a77-ae9b-621eed89715e" colab={}
x_valid.shape
# + id="6i93CFYFD6Z-" colab_type="code" outputId="ddab4f3b-dcde-448a-9d25-e2002c3df1d3" colab={}
x_imgs = np.reshape(x_valid, (-1,28,28)); x_imgs.shape
# + id="RsU8BzBiD6aC" colab_type="code" outputId="1be8c6b0-79ae-4b8f-840f-aceba98d2638" colab={}
show(x_imgs[0], y_valid[0])
# + id="mGurXCG7D6aG" colab_type="code" outputId="2f52c59c-e03b-4562-b75b-4ff23501f3aa" colab={}
y_valid.shape
# + [markdown] id="AdFokxBbD6aJ" colab_type="text"
# It's the digit 3! And that's stored in the y value:
# + id="Fmpo5C8OD6aK" colab_type="code" outputId="e93438ab-ebb6-4d16-f62e-322ab015e372" colab={}
y_valid[0]
# + [markdown] id="6DzmdUMGD6aN" colab_type="text"
# We can look at part of an image:
# + id="FJJZBwztD6aO" colab_type="code" outputId="0122f2bf-f1b0-4fc0-daec-e760668787e1" colab={}
x_imgs[0,10:15,10:15]
# + id="n_uXP41pD6aQ" colab_type="code" outputId="73905371-6e25-42ac-c407-f4abc7201cce" colab={}
show(x_imgs[0,10:15,10:15])
# + id="5K3kYDaWD6aV" colab_type="code" outputId="3d0028ca-5000-4fa9-a1b1-f6660983c9c7" colab={}
plots(x_imgs[:8], titles=y_valid[:8])
# + [markdown] id="1EYVosHaD6aa" colab_type="text"
# ## Neural Networks
# + [markdown] id="gBO60RMaD6ab" colab_type="text"
# We will take a deep look *logistic regression* and how we can program it ourselves. We are going to treat it as a specific example of a shallow neural net.
# + [markdown] id="ymasZOO0D6ac" colab_type="text"
# **What is a neural network?**
#
# A *neural network* is an *infinitely flexible function*, consisting of *layers*. A *layer* is a linear function such as matrix multiplication followed by a non-linear function (the *activation*).
#
# One of the tricky parts of neural networks is just keeping track of all the vocabulary!
# + [markdown] id="m2g1QGYqD6al" colab_type="text"
# ### Functions, parameters, and training
# + [markdown] id="01hfmiieD6an" colab_type="text"
# A **function** takes inputs and returns outputs. For instance, $f(x) = 3x + 5$ is an example of a function. If we input $2$, the output is $3\times 2 + 5 = 11$, or if we input $-1$, the output is $3\times -1 + 5 = 2$
#
# Functions have **parameters**. The above function $f$ is $ax + b$, with parameters a and b set to $a=3$ and $b=5$.
#
# Machine learning is often about learning the best values for those parameters. For instance, suppose we have the data points on the chart below. What values should we choose for $a$ and $b$?
# + [markdown] id="4E-9zuS7D6an" colab_type="text"
# <img src="https://github.com/fastai/fastai/blob/master/courses/ml1/images/sgd2.gif?raw=1" alt="" style="width: 70%"/>
# + [markdown] id="ZxyMdnquD6ao" colab_type="text"
# In the above gif from fast.ai's deep learning course, [intro to SGD notebook](https://github.com/fastai/courses/blob/master/deeplearning1/nbs/sgd-intro.ipynb)), an algorithm called stochastic gradient descent is being used to learn the best parameters to fit the line to the data (note: in the gif, the algorithm is stopping before the absolute best parameters are found). This process is called **training** or **fitting**.
#
# Most datasets will not be well-represented by a line. We could use a more complicated function, such as $g(x) = ax^2 + bx + c + \sin d$. Now we have 4 parameters to learn: $a$, $b$, $c$, and $d$. This function is more flexible than $f(x) = ax + b$ and will be able to accurately model more datasets.
#
# Neural networks take this to an extreme, and are infinitely flexible. They often have thousands, or even hundreds of thousands of parameters. However the core idea is the same as above. The neural network is a function, and we will learn the best parameters for modeling our data.
# + [markdown] id="ZyE7J-scD6ap" colab_type="text"
# ### PyTorch
# + [markdown] id="LBIWi0HQD6aq" colab_type="text"
# We will be using the open source [deep learning library, fastai](https://github.com/fastai/fastai), which provides high level abstractions and best practices on top of PyTorch. This is the highest level, simplest way to get started with deep learning. Please note that fastai requires Python 3 to function. It is currently in pre-alpha, so items may move around and more documentation will be added in the future.
#
# The fastai deep learning library uses [PyTorch](http://pytorch.org/), a Python framework for dynamic neural networks with GPU acceleration, which was released by Facebook's AI team.
#
# PyTorch has two overlapping, yet distinct, purposes. As described in the [PyTorch documentation](http://pytorch.org/tutorials/beginner/blitz/tensor_tutorial.html):
#
# <img src="https://github.com/fastai/fastai/blob/master/courses/ml1/images/what_is_pytorch.png?raw=1" alt="pytorch" style="width: 80%"/>
#
# The neural network functionality of PyTorch is built on top of the Numpy-like functionality for fast matrix computations on a GPU. Although the neural network purpose receives way more attention, both are very useful. We'll implement a neural net from scratch today using PyTorch.
#
# **Further learning**: If you are curious to learn what *dynamic* neural networks are, you may want to watch [this talk](https://www.youtube.com/watch?v=Z15cBAuY7Sc) by <NAME>, Facebook AI researcher and core PyTorch contributor.
#
# If you want to learn more PyTorch, you can try this [introductory tutorial](http://pytorch.org/tutorials/beginner/deep_learning_60min_blitz.html) or this [tutorial to learn by examples](http://pytorch.org/tutorials/beginner/pytorch_with_examples.html).
# + [markdown] id="UTEDJ8WBD6as" colab_type="text"
# ### About GPUs
# + [markdown] id="NU3ic2eoD6at" colab_type="text"
# Graphical processing units (GPUs) allow for matrix computations to be done with much greater speed, as long as you have a library such as PyTorch that takes advantage of them. Advances in GPU technology in the last 10-20 years have been a key part of why neural networks are proving so much more powerful now than they did a few decades ago.
#
# You may own a computer that has a GPU which can be used. For the many people that either don't have a GPU (or have a GPU which can't be easily accessed by Python), there are a few differnt options:
#
# - **Don't use a GPU**: For the sake of this tutorial, you don't have to use a GPU, although some computations will be slower.
# - **Use crestle, through your browser**: [Crestle](https://www.crestle.com/) is a service that gives you an already set up cloud service with all the popular scientific and deep learning frameworks already pre-installed and configured to run on a GPU in the cloud. It is easily accessed through your browser. New users get 10 hours and 1 GB of storage for free. After this, GPU usage is 34 cents per hour. I recommend this option to those who are new to AWS or new to using the console.
# - **Set up an AWS instance through your console**: You can create an AWS instance with a GPU by following the steps in this [fast.ai setup lesson](http://course.fast.ai/lessons/aws.html).] AWS charges 90 cents per hour for this.
# + [markdown] id="ZCT6Wgz1D6au" colab_type="text"
# ## Neural Net for Logistic Regression in PyTorch
# + id="3XuFGcOGD6av" colab_type="code" colab={}
from fastai.metrics import *
from fastai.model import *
from fastai.dataset import *
import torch.nn as nn
# + [markdown] id="_SENuT8xD6ay" colab_type="text"
# We will begin with the highest level abstraction: using a neural net defined by PyTorch's Sequential class.
# + id="_DUFbJE8D6az" colab_type="code" colab={}
net = nn.Sequential(
nn.Linear(28*28, 100),
nn.ReLU(),
nn.Linear(100, 100),
nn.ReLU(),
nn.Linear(100, 10),
nn.LogSoftmax()
).cuda()
# + [markdown] id="GDCnbXeyD6a3" colab_type="text"
# Each input is a vector of size `28*28` pixels and our output is of size `10` (since there are 10 digits: 0, 1, ..., 9).
#
# We use the output of the final layer to generate our predictions. Often for classification problems (like MNIST digit classification), the final layer has the same number of outputs as there are classes. In that case, this is 10: one for each digit from 0 to 9. These can be converted to comparative probabilities. For instance, it may be determined that a particular hand-written image is 80% likely to be a 4, 18% likely to be a 9, and 2% likely to be a 3.
# + id="OmkxRDTzD6a5" colab_type="code" colab={}
md = ImageClassifierData.from_arrays(path, (x,y), (x_valid, y_valid))
# + id="NWHu18TwD6a9" colab_type="code" colab={}
loss=nn.NLLLoss()
metrics=[accuracy]
# opt=optim.SGD(net.parameters(), 1e-1, momentum=0.9)
opt=optim.SGD(net.parameters(), 1e-1, momentum=0.9, weight_decay=1e-3)
# + [markdown] id="4P1VLKrSD6bI" colab_type="text"
# ### Loss functions and metrics
# + [markdown] id="WO6vvLZRD6bK" colab_type="text"
# In machine learning the **loss** function or cost function is representing the price paid for inaccuracy of predictions.
#
# The loss associated with one example in binary classification is given by:
# `-(y * log(p) + (1-y) * log (1-p))`
# where `y` is the true label of `x` and `p` is the probability predicted by our model that the label is 1.
# + id="atsE_h4VD6bL" colab_type="code" colab={}
def binary_loss(y, p):
return np.mean(-(y * np.log(p) + (1-y)*np.log(1-p)))
# + id="tEdenYmjD6bQ" colab_type="code" outputId="174f3761-171f-4979-9bf3-29ca2c983203" colab={}
acts = np.array([1, 0, 0, 1])
preds = np.array([0.9, 0.1, 0.2, 0.8])
binary_loss(acts, preds)
# + [markdown] id="WarT-znCD6bW" colab_type="text"
# Note that in our toy example above our accuracy is 100% and our loss is 0.16. Compare that to a loss of 0.03 that we are getting while predicting cats and dogs. Exercise: play with `preds` to get a lower loss for this example.
#
# **Example:** Here is an example on how to compute the loss for one example of binary classification problem. Suppose for an image x with label 1 and your model gives it a prediction of 0.9. For this case the loss should be small because our model is predicting a label $1$ with high probability.
#
# `loss = -log(0.9) = 0.10`
#
# Now suppose x has label 0 but our model is predicting 0.9. In this case our loss is should be much larger.
#
# `loss = -log(1-0.9) = 2.30`
#
# - Exercise: look at the other cases and convince yourself that this make sense.
# - Exercise: how would you rewrite `binary_loss` using `if` instead of `*` and `+`?
#
# Why not just maximize accuracy? The binary classification loss is an easier function to optimize.
#
# For multi-class classification, we use *negative log liklihood* (also known as *categorical cross entropy*) which is exactly the same thing, but summed up over all classes.
# + [markdown] id="4CaSn9DJD6bY" colab_type="text"
# ### Fitting the model
# + [markdown] id="Hr-LzRyKD6ba" colab_type="text"
# *Fitting* is the process by which the neural net learns the best parameters for the dataset.
# + id="rsno0GZgD6bb" colab_type="code" outputId="32925cb9-d9da-4294-85e6-714cc9dcf33e" colab={}
fit(net, md, n_epochs=5, crit=loss, opt=opt, metrics=metrics)
# + id="Ig4_ILk6D6bh" colab_type="code" colab={}
set_lrs(opt, 1e-2)
# + id="J5-EzOAnD6bn" colab_type="code" outputId="10574b44-3708-4302-a7e8-b61bfecef719" colab={}
fit(net, md, n_epochs=3, crit=loss, opt=opt, metrics=metrics)
# + id="vMI-O4lgD6bs" colab_type="code" colab={}
# + id="WiOfGv61D6bx" colab_type="code" outputId="d3fb56fc-5d72-41db-89b3-c7506cad7a62" colab={}
fit(net, md, n_epochs=5, crit=loss, opt=opt, metrics=metrics)
# + id="40UXnGBtD6b3" colab_type="code" colab={}
set_lrs(opt, 1e-2)
# + id="Uoztn7N2D6b6" colab_type="code" outputId="20335a1c-8c0f-4b14-982f-320f428a9722" colab={}
fit(net, md, n_epochs=3, crit=loss, opt=opt, metrics=metrics)
# + id="Kmdg-A-vD6cA" colab_type="code" outputId="7f16eb8f-c771-40a7-f597-ce745ac5eea8" colab={}
t = [o.numel() for o in net.parameters()]
t, sum(t)
# + [markdown] id="t1ak4v6DD6cF" colab_type="text"
# GPUs are great at handling lots of data at once (otherwise don't get performance benefit). We break the data up into **batches**, and that specifies how many samples from our dataset we want to send to the GPU at a time. The fastai library defaults to a batch size of 64. On each iteration of the training loop, the error on 1 batch of data will be calculated, and the optimizer will update the parameters based on that.
#
# An **epoch** is completed once each data sample has been used once in the training loop.
#
# Now that we have the parameters for our model, we can make predictions on our validation set.
# + id="vEyP3PQhD6cH" colab_type="code" colab={}
preds = predict(net, md.val_dl)
# + id="cc_BIy_vD6cK" colab_type="code" outputId="fba49c1a-2458-45fb-ecba-980888d70b6a" colab={}
preds.shape
# + [markdown] id="KoSEQDiCD6cM" colab_type="text"
# **Question**: Why does our output have length 10 (for each image)?
# + id="GPuuGMFkD6cN" colab_type="code" outputId="2fd252bf-6997-428e-92b1-0a88f005f2d2" colab={}
preds.argmax(axis=1)[:5]
# + id="ffKGWpbMD6cS" colab_type="code" colab={}
preds = preds.argmax(1)
# + [markdown] id="ydIqWkUGD6cU" colab_type="text"
# Let's check how accurate this approach is on our validation set. You may want to compare this against other implementations of logistic regression, such as the one in sklearn. In our testing, this simple pytorch version is faster and more accurate for this problem!
# + id="t5ecLN0mD6cV" colab_type="code" outputId="377ab77c-ac55-4833-e1ea-d9952659d6f8" colab={}
np.mean(preds == y_valid)
# + [markdown] id="WVc6iAcyD6ca" colab_type="text"
# Let's see how some of our predictions look!
# + id="mhMF6ON4D6cb" colab_type="code" outputId="e10d3b5f-8b3c-4105-ab32-e7cd77f6abc0" colab={}
plots(x_imgs[:8], titles=preds[:8])
# + [markdown] id="HNQtZlk-D6ce" colab_type="text"
# ## Defining Logistic Regression Ourselves
# + [markdown] id="W9q2RulID6cg" colab_type="text"
# Above, we used pytorch's `nn.Linear` to create a linear layer. This is defined by a matrix multiplication and then an addition (these are also called `affine transformations`). Let's try defining this ourselves.
#
# Just as Numpy has `np.matmul` for matrix multiplication (in Python 3, this is equivalent to the `@` operator), PyTorch has `torch.matmul`.
#
# Our PyTorch class needs two things: constructor (says what the parameters are) and a forward method (how to calculate a prediction using those parameters) The method `forward` describes how the neural net converts inputs to outputs.
#
# In PyTorch, the optimizer knows to try to optimize any attribute of type **Parameter**.
# + id="EZbOdMu-D6ch" colab_type="code" colab={}
def get_weights(*dims): return nn.Parameter(torch.randn(dims)/dims[0])
def softmax(x): return torch.exp(x)/(torch.exp(x).sum(dim=1)[:,None])
class LogReg(nn.Module):
def __init__(self):
super().__init__()
self.l1_w = get_weights(28*28, 10) # Layer 1 weights
self.l1_b = get_weights(10) # Layer 1 bias
def forward(self, x):
x = x.view(x.size(0), -1)
x = (x @ self.l1_w) + self.l1_b # Linear Layer
x = torch.log(softmax(x)) # Non-linear (LogSoftmax) Layer
return x
# + [markdown] id="FnCmaIP7D6cl" colab_type="text"
# We create our neural net and the optimizer. (We will use the same loss and metrics from above).
# + id="0YJEAi1sD6cl" colab_type="code" colab={}
net2 = LogReg().cuda()
opt=optim.Adam(net2.parameters())
# + id="XSQTerdGD6cn" colab_type="code" outputId="80e3ed96-f71a-49ba-9a6a-24512cc8a66c" colab={}
fit(net2, md, n_epochs=1, crit=loss, opt=opt, metrics=metrics)
# + id="UrA64OPvD6cr" colab_type="code" colab={}
dl = iter(md.trn_dl)
# + id="L2hqNQi3D6cu" colab_type="code" colab={}
xmb,ymb = next(dl)
# + id="QsbKCjabD6cy" colab_type="code" outputId="e285b4f9-1bb5-4de4-a7d4-3bc13e6058a2" colab={}
vxmb = Variable(xmb.cuda())
vxmb
# + id="SB26NMyFD6c2" colab_type="code" outputId="025589ff-447c-4033-e4fa-04e8c723ec5d" colab={}
preds = net2(vxmb).exp(); preds[:3]
# + id="8jVK6TDVD6c6" colab_type="code" outputId="bfedf594-ad90-402d-bc76-39cc5bfef585" colab={}
preds = preds.data.max(1)[1]; preds
# + [markdown] id="84UzLhGVD6dA" colab_type="text"
# Let's look at our predictions on the first eight images:
# + id="sNbPXZ2YD6dB" colab_type="code" outputId="2d0d13d8-569d-485f-c1af-f2911b80c55c" colab={}
preds = predict(net2, md.val_dl).argmax(1)
plots(x_imgs[:8], titles=preds[:8])
# + id="sd7sQts3D6dF" colab_type="code" outputId="4c306737-d458-4d3f-f19b-23a13dbe8342" colab={}
np.mean(preds == y_valid)
# + [markdown] id="7U1YQLpjD6dI" colab_type="text"
# ## Aside about Broadcasting and Matrix Multiplication
# + [markdown] id="XEhaSU2SD6dJ" colab_type="text"
# Now let's dig in to what we were doing with `torch.matmul`: matrix multiplication. First, let's start with a simpler building block: **broadcasting**.
# + [markdown] id="4dmR6fgkD6dK" colab_type="text"
# ### Element-wise operations
# + [markdown] id="W8cYRnfLD6dL" colab_type="text"
# Broadcasting and element-wise operations are supported in the same way by both numpy and pytorch.
#
# Operators (+,-,\*,/,>,<,==) are usually element-wise.
#
# Examples of element-wise operations:
# + id="4H7ltLxRD6dM" colab_type="code" outputId="bf661264-d443-4d38-90c6-ed2a155d739c" colab={}
a = np.array([10, 6, -4])
b = np.array([2, 8, 7])
a,b
# + id="Gws2iaJXD6dT" colab_type="code" outputId="7a549184-6765-4306-9072-8dcf50853037" colab={}
a + b
# + id="a9QeU0keD6dW" colab_type="code" outputId="7da315de-5b68-43e9-8f7b-e81c8db5c167" colab={}
(a < b).mean()
# + [markdown] id="5WdypDkaD6dZ" colab_type="text"
# ### Broadcasting
# + [markdown] id="IkoPQ7lED6da" colab_type="text"
# The term **broadcasting** describes how arrays with different shapes are treated during arithmetic operations. The term broadcasting was first used by Numpy, although is now used in other libraries such as [Tensorflow](https://www.tensorflow.org/performance/xla/broadcasting) and Matlab; the rules can vary by library.
#
# From the [Numpy Documentation](https://docs.scipy.org/doc/numpy-1.10.0/user/basics.broadcasting.html):
#
# The term broadcasting describes how numpy treats arrays with
# different shapes during arithmetic operations. Subject to certain
# constraints, the smaller array is “broadcast” across the larger
# array so that they have compatible shapes. Broadcasting provides a
# means of vectorizing array operations so that looping occurs in C
# instead of Python. It does this without making needless copies of
# data and usually leads to efficient algorithm implementations.
#
# In addition to the efficiency of broadcasting, it allows developers to write less code, which typically leads to fewer errors.
#
# *This section was adapted from [Chapter 4](http://nbviewer.jupyter.org/github/fastai/numerical-linear-algebra/blob/master/nbs/4.%20Compressed%20Sensing%20of%20CT%20Scans%20with%20Robust%20Regression.ipynb#4.-Compressed-Sensing-of-CT-Scans-with-Robust-Regression) of the fast.ai [Computational Linear Algebra](https://github.com/fastai/numerical-linear-algebra) course.*
# + [markdown] id="bj_cjV3LD6db" colab_type="text"
# #### Broadcasting with a scalar
# + id="RBRYrnmVD6db" colab_type="code" outputId="cde32ebd-373f-4868-a1e8-f968034fa360" colab={}
a
# + id="vBYpyHnPD6dd" colab_type="code" outputId="330e7a71-309c-4449-a9bb-608762305e65" colab={}
a > 0
# + [markdown] id="nPQg4lJCD6df" colab_type="text"
# How are we able to do a > 0? 0 is being **broadcast** to have the same dimensions as a.
#
# Remember above when we normalized our dataset by subtracting the mean (a scalar) from the entire data set (a matrix) and dividing by the standard deviation (another scalar)? We were using broadcasting!
#
# Other examples of broadcasting with a scalar:
# + id="MXY_HjzND6df" colab_type="code" outputId="b12b046f-1081-41b8-97d9-55d3810b01c7" colab={}
a + 1
# + id="NuBGyv5gD6dh" colab_type="code" outputId="1acda8e5-d821-44e7-f7c1-30ca1b253ea4" colab={}
m = np.array([[1, 2, 3], [4,5,6], [7,8,9]]); m
# + id="hBncUnoGD6dk" colab_type="code" outputId="cf60cdcc-d540-48f9-a265-ac53bcc36743" colab={}
2*m
# + [markdown] id="9kxhtJC6D6dm" colab_type="text"
# #### Broadcasting a vector to a matrix
# + [markdown] id="VK0XpASHD6dm" colab_type="text"
# We can also broadcast a vector to a matrix:
# + id="Lr9knCorD6dn" colab_type="code" outputId="cfeb9df4-8e46-4063-8f7d-45e697e75935" colab={}
c = np.array([10,20,30]); c
# + id="2x5GCFduD6dp" colab_type="code" outputId="029de891-2b65-44c9-ab5a-cdd63b969b7b" colab={}
m + c
# + id="8_YId0akD6dq" colab_type="code" outputId="f5cae2eb-169e-4180-81c7-46c8e00fc127" colab={}
c + m
# + [markdown] id="J4WFEg2KD6du" colab_type="text"
# Although numpy does this automatically, you can also use the `broadcast_to` method:
# + id="m1fzYNIdD6dv" colab_type="code" outputId="f8a8a76d-6eca-48e5-e2f8-15e473d36c99" colab={}
c.shape
# + id="WCUBCdE5D6dy" colab_type="code" outputId="a5a9ee6b-7f3e-4c39-cdba-bb0065e5ce01" colab={}
np.broadcast_to(c[:,None], m.shape)
# + id="ezKQhhBtD6d0" colab_type="code" outputId="e87c398a-7ca3-490a-9de7-8b5ee70cda02" colab={}
np.broadcast_to(np.expand_dims(c,0), (3,3))
# + id="xO_6eLLBD6d4" colab_type="code" outputId="819d7ea8-37a7-4570-e10e-4d79d8b54963" colab={}
c.shape
# + id="uIpFC5END6d6" colab_type="code" outputId="f159bb9e-1413-4417-d8d1-03de80e76c34" colab={}
np.expand_dims(c,0).shape
# + [markdown] id="b-ALeOHXD6d8" colab_type="text"
# The numpy `expand_dims` method lets us convert the 1-dimensional array `c` into a 2-dimensional array (although one of those dimensions has value 1).
# + id="0dsGji-PD6d8" colab_type="code" outputId="73d6758f-6959-4227-dea9-29a44cbfe29e" colab={}
np.expand_dims(c,0).shape
# + id="KQvJ8UDXD6d-" colab_type="code" outputId="6537ed0d-89c2-4bf4-9978-cf13f0d7c93d" colab={}
m + np.expand_dims(c,0)
# + id="4rbNpNbqD6eA" colab_type="code" outputId="2d5a3d99-baaa-409d-f63a-299dc9b2230d" colab={}
np.expand_dims(c,1)
# + id="py-y50phD6eB" colab_type="code" outputId="02eeb239-711e-4b1f-c8f1-1617dd5d42a9" colab={}
c[:, None].shape
# + id="nSZo-svoD6eD" colab_type="code" outputId="55f9f972-a2dc-4dd6-f72a-4dbf4ff55ee8" colab={}
m + np.expand_dims(c,1)
# + id="ZZCUwGHMD6eF" colab_type="code" outputId="04b46e43-5b18-48b3-dd2c-c4f779eb15f4" colab={}
np.broadcast_to(np.expand_dims(c,1), (3,3))
# + [markdown] id="ETRKbrK-D6eG" colab_type="text"
# #### Broadcasting Rules
# + id="cTd6RvklD6eH" colab_type="code" outputId="5de84a1b-cb1d-43c6-a683-31a48312e3d3" colab={}
c[None]
# + id="o-vk22GND6eL" colab_type="code" outputId="8e232c90-0d10-4d95-8f18-cb3e48a685fa" colab={}
c[:,None]
# + id="7rmJZCiND6eP" colab_type="code" outputId="e266180d-f6ee-4690-e122-99aa30607efb" colab={}
c[None] > c[:,None]
# + id="LGgopKEPD6eR" colab_type="code" outputId="1109ba63-c131-44aa-fefc-14dda8648d28" colab={}
xg,yg = np.ogrid[0:5, 0:5]; xg,yg
# + id="3K63-xopD6eW" colab_type="code" outputId="b80cb4c7-f308-4290-acab-6bdf6837265f" colab={}
xg+yg
# + [markdown] id="y2dYSsChD6eZ" colab_type="text"
# When operating on two arrays, Numpy/PyTorch compares their shapes element-wise. It starts with the **trailing dimensions**, and works its way forward. Two dimensions are **compatible** when
#
# - they are equal, or
# - one of them is 1
#
# Arrays do not need to have the same number of dimensions. For example, if you have a `256*256*3` array of RGB values, and you want to scale each color in the image by a different value, you can multiply the image by a one-dimensional array with 3 values. Lining up the sizes of the trailing axes of these arrays according to the broadcast rules, shows that they are compatible:
#
# Image (3d array): 256 x 256 x 3
# Scale (1d array): 3
# Result (3d array): 256 x 256 x 3
#
# The [numpy documentation](https://docs.scipy.org/doc/numpy-1.13.0/user/basics.broadcasting.html#general-broadcasting-rules) includes several examples of what dimensions can and can not be broadcast together.
# + [markdown] id="agtzn1fHD6eZ" colab_type="text"
# ### Matrix Multiplication
# + [markdown] id="-BGAP4m-D6ea" colab_type="text"
# We are going to use broadcasting to define matrix multiplication.
# + id="2MW-yt1rD6eb" colab_type="code" outputId="99ac05b6-c71d-4cef-cbd4-263fdda31e44" colab={}
m, c
# + id="j-9zNbKAD6ef" colab_type="code" outputId="313fa4ea-b293-4444-adfe-4122aaf6191c" colab={}
m @ c # np.matmul(m, c)
# + [markdown] id="nW4MRfjFD6eg" colab_type="text"
# We get the same answer using `torch.matmul`:
# + id="oyuICeIFD6eh" colab_type="code" outputId="ab806f6c-8b16-4471-d01c-b7f48e167fab" colab={}
T(m) @ T(c)
# + [markdown] id="s0UNVQMKD6ei" colab_type="text"
# The following is **NOT** matrix multiplication. What is it?
# + id="JRO7Jmg_D6ej" colab_type="code" outputId="c32b261f-322e-4536-a17e-9ac5a325957e" colab={}
m,c
# + id="KPmzJ_OED6el" colab_type="code" outputId="9be346cf-dc14-4db8-920d-62ac91f4eac8" colab={}
m * c
# + id="Z510WpBYD6ep" colab_type="code" outputId="4215fea5-f273-47af-aa65-1d1d4cb509bc" colab={}
(m * c).sum(axis=1)
# + id="CdDhw-p_D6er" colab_type="code" outputId="4b07d951-70e9-4f27-a169-408b75fd8673" colab={}
c
# + id="bULcRKP3D6es" colab_type="code" outputId="62c1928c-0d21-4825-f47a-41938efef145" colab={}
np.broadcast_to(c, (3,3))
# + [markdown] id="9e-m0WP4D6eu" colab_type="text"
# From a machine learning perspective, matrix multiplication is a way of creating features by saying how much we want to weight each input column. **Different features are different weighted averages of the input columns**.
#
# The website [matrixmultiplication.xyz](http://matrixmultiplication.xyz/) provides a nice visualization of matrix multiplcation
# + id="p2_5ZT96D6eu" colab_type="code" outputId="56f60c86-7bb9-47eb-9f98-194e40a22818" colab={}
n = np.array([[10,40],[20,0],[30,-5]]); n
# + id="MQb9PPiVD6ex" colab_type="code" outputId="02ae796d-e986-4e51-d1c9-e91dc6dc5596" colab={}
m
# + id="1Y826vR5D6ez" colab_type="code" outputId="d55ccb6d-ccb7-4501-cc00-9113ed85ab8b" colab={}
m @ n
# + id="lt-IBymhD6e1" colab_type="code" outputId="8582fd8c-b0b5-4873-cfc0-b2cca9e4eae9" colab={}
(m * n[:,0]).sum(axis=1)
# + id="9US1M_WCD6e3" colab_type="code" outputId="21012cf6-d415-4901-8fc3-ee8955a8c4a1" colab={}
(m * n[:,1]).sum(axis=1)
# + [markdown] id="X1KGDPlAD6e5" colab_type="text"
# ## Writing Our Own Training Loop
# + [markdown] id="BqhYVOzoD6e5" colab_type="text"
# As a reminder, this is what we did above to write our own logistic regression class (as a pytorch neural net):
# + id="yRPMnbpqD6e5" colab_type="code" outputId="2ed1929b-4376-4983-ac44-02f078330375" colab={}
# Our code from above
class LogReg(nn.Module):
def __init__(self):
super().__init__()
self.l1_w = get_weights(28*28, 10) # Layer 1 weights
self.l1_b = get_weights(10) # Layer 1 bias
def forward(self, x):
x = x.view(x.size(0), -1)
x = x @ self.l1_w + self.l1_b
return torch.log(softmax(x))
net2 = LogReg().cuda()
opt=optim.Adam(net2.parameters())
fit(net2, md, n_epochs=1, crit=loss, opt=opt, metrics=metrics)
# + [markdown] id="Evz5hfUvD6e8" colab_type="text"
# Above, we are using the fastai method `fit` to train our model. Now we will try writing the training loop ourselves.
#
# **Review question:** What does it mean to train a model?
# + [markdown] id="WWGRzvKiD6e8" colab_type="text"
# We will use the LogReg class we created, as well as the same loss function, learning rate, and optimizer as before:
# + id="AT3ny7MPD6e8" colab_type="code" colab={}
net2 = LogReg().cuda()
loss=nn.NLLLoss()
learning_rate = 1e-3
optimizer=optim.Adam(net2.parameters(), lr=learning_rate)
# + [markdown] id="vUlqVTM5D6e-" colab_type="text"
# md is the ImageClassifierData object we created above. We want an iterable version of our training data (**question**: what does it mean for something to be iterable?):
# + id="vNjGZlkSD6e_" colab_type="code" colab={}
dl = iter(md.trn_dl) # Data loader
# + [markdown] id="mH8GqD_oD6fA" colab_type="text"
# First, we will do a **forward pass**, which means computing the predicted y by passing x to the model.
# + id="ju6pZAYmD6fB" colab_type="code" colab={}
xt, yt = next(dl)
y_pred = net2(Variable(xt).cuda())
# + [markdown] id="JynnAzHFD6fD" colab_type="text"
# We can check the loss:
# + id="8G4Ihci3D6fD" colab_type="code" outputId="e7e759be-f427-40a8-b8d1-791b5ed9dcb3" colab={}
l = loss(y_pred, Variable(yt).cuda())
print(l)
# + [markdown] id="8CkOjoN-D6fF" colab_type="text"
# We may also be interested in the accuracy. We don't expect our first predictions to be very good, because the weights of our network were initialized to random values. Our goal is to see the loss decrease (and the accuracy increase) as we train the network:
# + id="DdoUs9rLD6fF" colab_type="code" outputId="19230735-4ef1-49fb-f518-d00197327fc4" colab={}
np.mean(to_np(y_pred).argmax(axis=1) == to_np(yt))
# + [markdown] id="vad8PXBTD6fG" colab_type="text"
# Now we will use the optimizer to calculate which direction to step in. That is, how should we update our weights to try to decrease the loss?
#
# Pytorch has an automatic differentiation package ([autograd](http://pytorch.org/docs/master/autograd.html)) that takes derivatives for us, so we don't have to calculate the derivative ourselves! We just call `.backward()` on our loss to calculate the direction of steepest descent (the direction to lower the loss the most).
# + id="RVYW5aidD6fH" colab_type="code" colab={}
# Before the backward pass, use the optimizer object to zero all of the
# gradients for the variables it will update (which are the learnable weights
# of the model)
optimizer.zero_grad()
# Backward pass: compute gradient of the loss with respect to model parameters
l.backward()
# Calling the step function on an Optimizer makes an update to its parameters
optimizer.step()
# + [markdown] id="xOTy53VvD6fJ" colab_type="text"
# Now, let's make another set of predictions and check if our loss is lower:
# + id="aUZdo0t_D6fJ" colab_type="code" colab={}
xt, yt = next(dl)
y_pred = net2(Variable(xt).cuda())
# + id="jHNbeqpWD6fK" colab_type="code" outputId="4b378991-f040-48a0-cb1b-47a6358b1f83" colab={}
l = loss(y_pred, Variable(yt).cuda())
print(l)
# + [markdown] id="NVG-ZHOID6fL" colab_type="text"
# Note that we are using **stochastic** gradient descent, so the loss is not guaranteed to be strictly better each time. The stochasticity comes from the fact that we are using **mini-batches**; we are just using 64 images to calculate our prediction and update the weights, not the whole dataset.
# + id="wv3-GewDD6fM" colab_type="code" outputId="400a382b-944b-478f-8f8e-76d90900d08c" colab={}
np.mean(to_np(y_pred).argmax(axis=1) == to_np(yt))
# + [markdown] id="HpC69k6KD6fO" colab_type="text"
# If we run several iterations in a loop, we should see the loss decrease and the accuracy increase with time.
# + id="vPaKHLzHD6fO" colab_type="code" outputId="52941853-a0ff-4d51-aa84-9e25a38c866d" colab={}
for t in range(100):
xt, yt = next(dl)
y_pred = net2(Variable(xt).cuda())
l = loss(y_pred, Variable(yt).cuda())
if t % 10 == 0:
accuracy = np.mean(to_np(y_pred).argmax(axis=1) == to_np(yt))
print("loss: ", l.data[0], "\t accuracy: ", accuracy)
optimizer.zero_grad()
l.backward()
optimizer.step()
# + [markdown] id="MkEGdQgtD6fQ" colab_type="text"
# ### Put it all together in a training loop
# + id="yrX23JBRD6fQ" colab_type="code" colab={}
def score(x, y):
y_pred = to_np(net2(V(x)))
return np.sum(y_pred.argmax(axis=1) == to_np(y))/len(y_pred)
# + id="SxnznGrTD6fS" colab_type="code" outputId="85d50606-472b-4d2e-cc67-95b831766277" colab={}
net2 = LogReg().cuda()
loss=nn.NLLLoss()
learning_rate = 1e-2
optimizer=optim.SGD(net2.parameters(), lr=learning_rate)
for epoch in range(1):
losses=[]
dl = iter(md.trn_dl)
for t in range(len(dl)):
# Forward pass: compute predicted y and loss by passing x to the model.
xt, yt = next(dl)
y_pred = net2(V(xt))
l = loss(y_pred, V(yt))
losses.append(l)
# Before the backward pass, use the optimizer object to zero all of the
# gradients for the variables it will update (which are the learnable weights of the model)
optimizer.zero_grad()
# Backward pass: compute gradient of the loss with respect to model parameters
l.backward()
# Calling the step function on an Optimizer makes an update to its parameters
optimizer.step()
val_dl = iter(md.val_dl)
val_scores = [score(*next(val_dl)) for i in range(len(val_dl))]
print(np.mean(val_scores))
# + [markdown] id="5LrnDvvqD6fV" colab_type="text"
# ## Stochastic Gradient Descent
# + [markdown] id="lf7gKiXYD6fV" colab_type="text"
# Nearly all of deep learning is powered by one very important algorithm: **stochastic gradient descent (SGD)**. SGD can be seeing as an approximation of **gradient descent (GD)**. In GD you have to run through all the samples in your training set to do a single itaration. In SGD you use only a subset of training samples to do the update for a parameter in a particular iteration. The subset used in each iteration is called a batch or minibatch.
#
# Now, instead of using the optimizer, we will do the optimization ourselves!
# + id="JgkXxyv_D6fW" colab_type="code" outputId="01ab654d-468b-436b-f2c2-7374f37144de" colab={}
net2 = LogReg().cuda()
loss_fn=nn.NLLLoss()
lr = 1e-2
w,b = net2.l1_w,net2.l1_b
for epoch in range(1):
losses=[]
dl = iter(md.trn_dl)
for t in range(len(dl)):
xt, yt = next(dl)
y_pred = net2(V(xt))
l = loss(y_pred, Variable(yt).cuda())
losses.append(loss)
# Backward pass: compute gradient of the loss with respect to model parameters
l.backward()
w.data -= w.grad.data * lr
b.data -= b.grad.data * lr
w.grad.data.zero_()
b.grad.data.zero_()
val_dl = iter(md.val_dl)
val_scores = [score(*next(val_dl)) for i in range(len(val_dl))]
print(np.mean(val_scores))
# + id="Sc_T2LPxD6fZ" colab_type="code" colab={}
| Copy_of_lesson4_mnist_sgd.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 1.1 Problema Lineal de Producción de Helados
# ### 1.2 Objetivos
# #### 1.2.1 Objetivo general.
# Determinar el plan semanal de producción de los diferentes tipos de paletas que conforman la “Gama Gourmet”, con el objetivo de maximizar beneficios.
#
# #### 1.2.2 Objetivos específicos
# * Cumplir con la demanda semanal de la empresa.
# * Maximizar la ganancia de la empresa.
# * Determinar la cantidad de paletas a producir de cada sabor.
# ### 1.3 Modelo que representa el problema
# Nuestra empresa de origen valenciano afincada en Sevilla desde la década de los años 70, se dedica a la elaboración de helados artesanos. Después de estos años de grandes progresos en su negocio, desea abrir mercado para poder enfrentarse a la situación actual.
# Esta ampliación tiene como objetivo introducir sus productos en el sector de la hostelería, mediante la propuesta de una gama de helados que podemos considerar “Gourmet”. A continuación detallaremos dicha gama.
# Creada por el gran Filippo Zampieron está compuesta por cinco tipos de paletas artesanales:
# 1. Paletas de menta
# 2. Paletas de chocolate
# 3. Paletas de yogurt y melocotón
# 4. Paletas de almendras
# 5. Paletas “Fiordilatte”.
#
# Aunque la elaboración de todas las paletas difieren en diversos aspectos, ya sea en la composición de la base, cobertura o en las proporciones de cada componente, hay un producto común en todas ellas; “*Jarabe Base*” ya que sin este no sería posible la fabricación de la base de las paletas.
# Este Jarabe, está compuesto por:
# * Agua: 655 gr
# * Azúcar de caña : 180 gr
# * Dextosa: 35 gr
# * Glucosa: 130 gr
#
# A continuación detallamos el proceso de elaboración y las cantidades utilizadas
# para la fabricación de un kilo de cada tipo de paletas.
#
# #### <font color="darkblue">Paletas de menta
# <img style="float: right; margin: 0px 0px 15px 15px;" src="https://likeavegan.com.au/wp-content/uploads/2014/10/chocmintpop.jpg" width="200px" height="75px" />
#
# La fabricación de este producto comienza con la elaboración de la base. Para ello se utilizan $550 gr$ del jarabe, seguido de unas gotas de esencia de menta ($10 gotas$) y posteriormente añadiendo unos $450 gr$ de leche fresca entera.
# Una vez que se ha mezclado la base y se ha dejado reposar para conseguir una textura idónea se procede a la elaboración de su cobertura.
# Está compuesta por unos $800 gr$ chocolate y $200 gr$ de manteca de cacao.
# #### <font color="darkblue">Paletas de Chocolate
# <img style="float: left; margin: 10px 10px 10px 10px;" src="https://s23991.pcdn.co/wp-content/uploads/2011/08/mexican-chocolate-pops-recipe.jpg" width="150px" height="50px" />
# La base de estas está compuesta por: $500 gr$ de jarabe, $440 gr$ de leche entera fresca unos $25 gr$ de azúcar invertido (una combinación de glucosa y fructosa) y por último, 35 gr de cacao.
# La cobertura al igual que el producto anterior está compuesta por: $800 gr$ de chocolate y $200 gr$ de manteca de cacao.
# #### <font color="darkblue">Paletas de Yogurt y Melocotón
# <img style="float: right; margin: 10px 10px 10px 10px;" src="https://cookieandkate.com/images/2015/08/peach-popsicles-recipe.jpg" width="150px" height="50px" />
# Con una base compuesta por: $430 gr$ de jarabe, $300 gr$ de yogurt desnatado, $20 gr$ de azúcar invertido y $250 gr$ de melocotón batido.
# Su cobertura es una dulce combinación de $500 gr$ de chocolate y $500 gr$ de nata.
# #### <font color="darkblue">Paletas de almendra
# <img style="float: left; margin: 10px 10px 10px 10px;" src="https://www.thelittleepicurean.com/wp-content/uploads/2015/01/banana-coconut-ice-pops.jpg" width="200px" height="50px" />
# Base elaborada por: $400 gr$ de jarabe, $495 gr$ de leche fresca entera, $25 gr$ de azúcar invertido.
# La cobertura está elaborada por $800 gr$ de chocolate, $200 gr$ de manteca de cacao y $80 gr$ de pasta de almendras.
# #### <font color="darkblue">Paletas “Fiordilatte”
# <img style="float: right; margin: 10px 10px 15px 15px;" src="https://justbakedbyme.files.wordpress.com/2012/07/vanilla-bean-custard-pops-7.jpg?w=768&h=559" width="200px" height="50px" />
# Su elaboración comienza con la base compuesta por $510 gr$ de jarabe, $510 gr$ de leche fresca entera, $250 gr$ de nata, $200 gr$ de azúcar invertido.
# Una vez que la base se haya mezclado y adoptado la textura deseada, se le inyecta un relleno compuesto por: $550 gr$ de nata y $500 gr$ de chocolate.
# Finalmente, esperado el tiempo necesario para que el relleno se adapte a la base, se le añade una cobertura de $800 gr$ de chocolate y $200 gr$ de manteca de cacao.
# **Se ha realizado un estudio de mercado y de producción, que nos proporcionan los siguientes datos:**
#
# #### <font color="blue">*Beneficio esperado por cada kilo de las diferentes paletas:*
#
# * Paletas de menta: $23$ pesos el kilo
# * Paletas de chocolate: $22.5$ pesos el kilo
# * Paletas de yogurt y melocotón: $21$ pesos el kilo
# * Paletas de almendras: $20.5$ pesos el kilo
# * Paletas “ Fiordilatte”: $21$ pesos el kilo
#
# #### <font color="blue">*Disponibilidad semanal de las siguientes materias primas :*
#
# * Jarabe Base: $20,5$ kg
# * Leche Fresca entera: $13$ kg
# * Yogurt desnatado: $5$ kg
# * Nata: $8,5$ kg
# * Azúcar invertido: $1,3$ kg
# * Chocolate: $27$ kg
# * Manteca de cacao: $5$ kg
# * Esencia de menta: $2$ frasco de $90 ml$, cada frasco proporciona $75$ gotas.
# * Cacao: $0,28$ kg ( $2$ bolsas de $140$ gr cada uno)
# * Melocotón batido: $4$ kg
# * Pasta de almendras: $0,8$ kg ( $2$ bolsas de $400$ gr cada una)
#
# #### <font color="blue">*Demanda esperada semanalmente para cada tipo de paleta:*
# * Demanda de paletas de menta y paletas de chocolate: $10$ kilos
# * Demanda de paletas de yogurt y paletas de almendras: $10$ kilos
# * No se ha estimado demanda alguna de paletas Fiordilatte.
#
# #### <font color="blue">*Variables de decisión:*
# * $x_1$ kilos a fabricar semanalmente de paletas de menta
# * $x_2$ kilos a fabricar semanalmente de paletas de chocolate
# * $x_3$ kilos a fabricar semanalmente de paletas de yogur y melocotón
# * $x_4$ kilos a fabricar semanalmente de paletas de almendras
# * $x_5$ kilos a fabricar semanalmente de paletas fiordilatte
# #### <font color="blue">*Restricciones*
#
# Limitación de Jarabe Base: $20,5$ kilos
#
# $550x_1+500x_2+430x_3+400x_4+510x_5+\leq20500$
#
# Limitación de Leche Fresca Entera: $13$ kilos
#
# $450x_1+440x_2+495x_4+510x_5\leq13000$
#
# Limitación de Yogurt desnatado: $5$ kilos
#
# $300x_3\leq5000$
#
# Limitación de Nata: $8,5$ kilos
#
# $500x_3+550x_5\leq8500$
#
# Limitación de Azúcar invertido: $1,3$ kilo
#
# $25x_2+20x_3+25x_4+200x_5\leq1300$
#
# Limitación de Chocolate: $27$ kilos
#
# $800x_1+800x_2+500x_3+800x_4+1300x_5\leq27000$
#
# Limitación de Manteca de cacao: 5 kilos
#
# $200x_1+200x_2+200x_4+200x_5\leq5000$
#
# Limitación de Esencia de menta: $150$ gotas
#
# $10x_1\leq150$
#
# Limitación de Cacao: 0,28 kg
#
# $35x_2\leq280$
#
# Limitación de Melocotón Batido: 4 kilos
#
# $250x_3\leq4000$
#
# Limitación de Pasta de Almendras: 0.8 kilo
#
# $80x_4\leq800$
#
# Restricciones con respecto a la demanda:
#
# $x_1+x_2\geq10$
#
# $x_3+x_4\geq10$
#
# Función Objetivo: Beneficio
#
# $23x_1+22.5x_2 21x_3 20.5x_4 21x_5 $
# ### 1.4 Solución del problema de optimización.
# Sintetizando las restricciones y nuestra función a optimizar obtenemos lo siguiente:
#
# Max $23x_1+22.5x_2+21x_3+20.5x_4+21x_5$
#
# s.a.
#
# $550x_1+500x_2+430x_3+400x_4+510x_5\leq20500$
#
# $450x_1+440x_2+495x_4+510x_5\leq13000$
#
# $300x_3\leq5000$
#
# $500x_3+550x5\leq8500$
#
# $25x_2+20x_3+25x_4+200x_5\leq1300$
#
# $800x_1+800x_2+500x_3+800x_4+1300x_5\leq27000$
#
# $200x_1+200x_2+200x_4+200x_5\leq5000$
#
# $10x_1\leq150$
#
# $35x_2\leq280$
#
# $250x_3\leq4000$
#
# $80x_4\leq800$
#
# $x_1+x_2\geq10$
#
# $x_3+x_4\geq10$
#
# $x_1,x_2,x_3,x_4,x_5\geq0$
import numpy as np
import scipy.optimize as opt
c = -np.array([23, 22.5, 21, 20.5, 21])
A = np.array([[550, 500, 430, 400, 510],
[450, 440, 0, 495, 510],
[0, 0, 300, 0, 0],
[0, 0, 500, 0, 550],
[0, 25, 20, 25, 200],
[800, 800, 500, 800, 1300],
[200, 200, 0, 200, 200],
[10, 0, 0, 0, 0],
[0, 35, 0, 0, 0],
[0, 0, 250, 0, 0],
[0, 0, 0, 80, 0],
[-1, -1, 0, 0, 0],
[0, 0, -1, -1, 0]])
b = np.array([20500, 13000, 5000, 8500, 1300, 27000, 5000, 150, 280, 4000, 800, -10, -10])
utilidad = opt.linprog(c, A_ub=A, b_ub=b)
utilidad
# ### 1.5 Visualización de la solución del problema
# Una vez que hemos descrito el proceso de producción de cada producto, hemos sintetizado toda la información en el siguiente cuadro para ver de una forma más clara, los requisitos de materias primas por tipo de paleta
#
# |Jarabe|Leche entera fresca|Yogurt desnatado|Nata|Azúcar inv.|Chocolate|Manteca cacao|Esencia menta|Cacao|Melocotón|Pasta de almendras
# :----|----
# <NAME>|550gr|450gr||||800gr|200gr|10gr
# <NAME>|500|440|||25|800|200||35
# <NAME> y Melocotón|430||300|500|20|500||||250
# <NAME>|400|495|||25|800|200||||80
# <NAME>|510|510||550|200|1300|200
#
# Teniendo una demanda:
#
#
# $P. Menta+P. Chocolate\geq10$
#
# $P. Yogurt y melocotón + P. Almendra\geq10$
resultado= utilidad.x
resultado
excedente= utilidad.slack
excedente
# ### 1.6 Conclusiones
#
#
# Nuestra empresa va a obtener un beneficio máximo de **876,375 pesos** cuando se producen $15 k$ de paletas de menta, $8 k$ de paletas de chocolate, $16 k $ de paletas de yogurt y melocotón junto con $0.75 k $ de paletas de almendra. En esta producción óptima no se contempla la fabricación de las paletas de "Fiordilatte" y como estas no estan contempladas en nuestra demanda no afecta nuestra proyección de ventas.
#
# Cuando se cumple esta producción óptima vamos a agotar las siguientes materias primas:
# * Chocolate
# * Esencia de menta
# * Cacao
# * Melocotón batido
#
# Mientras que quedamos con lo siguiente en el almacén:
# * $1070 gr$ de Jarabe Base
# * $2358.75 gr$ de leche entera fresca
# * $200 gr$ de yogurt desnatado
# * $500 gr$ de nata
# * $761.25 gr$ de azúcar invertido
# * $250 gr$ de manteca de cacao
# * $740 gr$ de pasta de almendras
#
# Con respecto a la demanda, encontramos un excedente de $13 k$ de paletas de menta y chocolate, mientras que son $6.75 k$ el excedente de paletas de almendra y yogurt con melocotón.
# ### 1.7 Referencias
# * https://idus.us.es/xmlui/bitstream/handle/11441/42359/conejero%20calvo%20de%20leon%20elena%20tr.pdf?sequence=1
| Proyecto_Modulo1_GalindoA_Pimentel.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import joblib
from os.path import join
# Define constants
eval_output_dir = "../output/word2vec_eval_analogies/word2vec_enwiki_jan_2021"
sswr_filepath = join(eval_output_dir, "sswr.joblib")
msr_filepath = join(eval_output_dir, "msr.joblib")
pad_filepath = join(eval_output_dir, "pad.joblib")
# Load evaluation results
sswr_analogy_accuracies = joblib.load(sswr_filepath)
msr_analogy_accuracies = joblib.load(msr_filepath)
pad_analogy_accuracies = joblib.load(pad_filepath)
# SSWR evaluation results
sswr_analogy_accuracies
# MSR evaluation results
msr_analogy_accuracies
# PAD evaluation results
pad_analogy_accuracies
| code/word_embeddings/evaluate_word2vec_sswr_msr_pad.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/komo135/tensorflow-rl/blob/master/ipynb/dqn.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab_type="code" id="vtJ6_QRbHucx" outputId="910c0cc0-fd8c-4333-9a6f-3534b4e475d1" colab={"base_uri": "https://localhost:8080/", "height": 51}
# Google ドライブをマウントするには、このセルを実行してください。
from google.colab import drive
drive.mount('/content/drive')
# %cd drive/My Drive
# + colab_type="code" id="nPAcy7EKniBL" outputId="ae1596e8-338c-4368-b807-b64c7d922b67" colab={"base_uri": "https://localhost:8080/", "height": 34}
# %tensorflow_version 2.x
import tensorflow as tf
# + colab_type="code" id="LBbFS0xIHubh" outputId="0398d08e-9dfe-42b3-bfc9-adec6a8afb93" colab={"base_uri": "https://localhost:8080/", "height": 34}
# %cd /content/drive/My Drive
try:
import imp
imp.reload(dqn)
except:
import dqn
agent = dqn.Agent(spread=10, pip_cost=1000, leverage=500, min_lots=0.01, assets=10000, available_assets_rate=0.4,
restore=not True, step_size=96, n=4, lr=1e-3)
# + colab_type="code" id="eG6AXzJYNXt5" outputId="c30cee89-4a9e-4d14-8c76-8da04627d388" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# %cd /content
agent.run()
# + id="Tqg6ocmoPfbz" colab_type="code" outputId="24814687-3f29-443b-feea-234ffeba81de" colab={"base_uri": "https://localhost:8080/", "height": 1000}
import numpy as np
def softmax(x):
"""Compute softmax values for each sets of scores in x."""
e_x = np.exp(x - np.max(x))
return e_x / e_x.sum()
# agent.test(spread=10, pip_cost=1000, los_cut=150,test_data=not True)
for _ in range(1):
tree_idx, replay = agent.memory.sample(128)
states = np.array([a[0][0] for a in replay], np.float32)
new_states = np.array([a[0][3] for a in replay], np.float32)
actions = np.array([a[0][1] for a in replay])
rewards = np.array([a[0][2] for a in replay], np.float32).reshape((-1, 1))
with tf.GradientTape() as tape:
q = agent.model(states)
target_q = agent.target_model(new_states).numpy()
arg_q = agent.model(new_states).numpy()
random = np.random.rand(actions.shape[0])
arg_q = np.argmax(arg_q, 1)
# arg_q = np.array([np.argmax(arg_q[i]) if random[i] > 0.1 else np.random.randint(arg_q.shape[1]) for i in
# range(arg_q.shape[0])])
q_backup = q.numpy()
for i in range(len(rewards)):
# q_backup[i, actions[i]] = rewards[i] if I < 1010 and not self.restore else rewards[i] + 0.2 * target_q[i, np.argmax(arg_q[i])]
q_backup[i, actions[i]] = rewards[i] + 0.1 * target_q[i, arg_q[i]]
mse = tf.reduce_mean(tf.reduce_sum(tf.abs(q_backup - q) ** 1.5, -1))
ae = np.array([sum(i) for i in np.abs(q_backup - q.numpy())])
agent.memory.batch_update(tree_idx, ae)
gradients = tape.gradient(mse, agent.model.trainable_variables)
# gradients = [(tf.clip_by_value(grad, -10.0, 10.0))
# for grad in gradients]
# agent.model.optimizer.apply_gradients(zip(gradients,agent.model.trainable_variables))
# print(np.mean(ae))
# print(q[0:5])
print(mse)
print(q)
# print(np.mean(rewards)))
# + id="uZdB0Wfu4R4I" colab_type="code" outputId="b06186c3-62af-4a38-b09c-405932f3bb4b" colab={"base_uri": "https://localhost:8080/", "height": 1000}
rewards
# + id="cPuyKpRvbijV" colab_type="code" colab={}
# !pip install ta
import numpy as np
import pandas as pd
import ta
from sklearn.preprocessing import MinMaxScaler
# + id="aGLMeGnLbA2b" colab_type="code" colab={}
def gen_data(file_path="gbpjpy15.csv"):
try:
print("load file")
df = pd.read_csv(file_path)
except:
print("Use 'python gen_data.py 'file_path''")
return
df["Close1"] = df["Close"] * 100
ma = np.array(ta.trend.ema(df["Close1"], 7) - ta.trend.ema(df["Close1"], 4)).reshape((-1,1))
ma2 = np.array(df["Close1"] - ta.trend.ema(df["Close1"], 7)).reshape((-1,1))
b1 = np.array(ta.volatility.bollinger_hband(df["Close1"]) - df["Close1"]).reshape((-1,1))
b2 = np.array(ta.volatility.bollinger_lband(df["Close1"]) - df["Close1"]).reshape((-1,1))
macd = np.array(ta.trend.macd_diff(df["Close1"])).reshape((-1,1))
rsi = np.array(ta.momentum.rsi(df["Close"]) - ta.momentum.rsi(df["Close"], 7)).reshape((-1,1))
stoch = np.array(ta.momentum.stoch_signal(df["High"], df["Low"], df["Close"]) - ta.momentum.stoch(df["High"], df["Low"], df["Close"])).reshape((-1,1))
x = np.concatenate([ma, macd, rsi, stoch], -1)
y = np.array(df[["Open"]])
atr = np.array(ta.volatility.average_true_range(df["High"], df["Low"], df["Close"]))
high = np.array(df[["High"]])
low = np.array(df[["Low"]])
print("gen time series data")
gen = tf.keras.preprocessing.sequence.TimeseriesGenerator(x, y, 30)
x = []
y = []
for i in gen:
x.extend(i[0].tolist())
y.extend(i[1].tolist())
x = np.asanyarray(x)[100:]
y = np.asanyarray(y)[100:]
atr = atr[-len(y):].reshape((-1, 1))
scale_atr = atr
high = high[-len(y):].reshape((-1, 1))
low = low[-len(y):].reshape((-1, 1))
np.save("x", x)
np.save("target", np.array([y, atr, scale_atr, high, low]))
print("done\n")
# + id="crWStywfcPY9" colab_type="code" colab={}
# %cd /content/drive/My Drive
gen_data()
# + id="gQ3ZAslQ38xE" colab_type="code" colab={}
def softmax(x):
"""Compute softmax values for each sets of scores in x."""
e_x = np.exp(x - np.max(x))
return e_x / e_x.sum()
# + id="_u_RsMfUl_9K" colab_type="code" colab={}
a1 = np.argmax(q,1)
# + id="6iwGD_4xl5U5" colab_type="code" colab={}
q1 = np.abs(q) / np.sum(np.abs(q),1).reshape((-1,1)) * (np.abs(q) / q)
q1 += .05 * np.random.randn(q1.shape[0], q1.shape[1])
a2 = np.argmax(q1,-1)
# + id="S_lJJT26mLMm" colab_type="code" colab={}
np.mean(a1 == a2)
# + id="bgv_CBM0kXWG" colab_type="code" colab={}
tree_idx, replay = agent.memory.sample(128)
# + id="iL75_g3EkZAg" colab_type="code" colab={}
3 in [1,2]
| ipynb/dqn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/yukinaga/ai_music/blob/main/section_1/02_exercise.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="z8g5FUr1BPri"
# # 演習
# NoteSequenceの扱いに慣れるために、「きらきら星」を様々な楽器で演奏しましょう。
# + [markdown] id="OHuehCAzjyk-"
# ## ライブラリのインストール
# + id="3iEXOSdljVZN"
# !apt-get update -qq && apt-get install -qq libfluidsynth1 fluid-soundfont-gm build-essential libasound2-dev libjack-dev
# !pip install -qU pyfluidsynth pretty_midi
# !pip install -qU magenta
# + [markdown] id="pqPT4STAlQIJ"
# 以下のコードの変数`pg`および`is_dm`の値を変更し、様々な楽器で「きらきら星」を演奏しましょう。
# + id="YYikkBYClqyc"
import magenta
import note_seq
from note_seq.protobuf import music_pb2
kira2 = music_pb2.NoteSequence() # NoteSequence
pg = 12 # 楽器の種類 -ここを変更-
is_dm = False # ドラムかどうか -ここを変更-
# notesにnoteを追加
kira2.notes.add(pitch=60, start_time=0.0, end_time=0.4, velocity=80, program=pg, is_drum=is_dm)
kira2.notes.add(pitch=60, start_time=0.4, end_time=0.8, velocity=80, program=pg, is_drum=is_dm)
kira2.notes.add(pitch=67, start_time=0.8, end_time=1.2, velocity=80, program=pg, is_drum=is_dm)
kira2.notes.add(pitch=67, start_time=1.2, end_time=1.6, velocity=80, program=pg, is_drum=is_dm)
kira2.notes.add(pitch=69, start_time=1.6, end_time=2.0, velocity=80, program=pg, is_drum=is_dm)
kira2.notes.add(pitch=69, start_time=2.0, end_time=2.4, velocity=80, program=pg, is_drum=is_dm)
kira2.notes.add(pitch=67, start_time=2.4, end_time=3.2, velocity=80, program=pg, is_drum=is_dm)
kira2.notes.add(pitch=65, start_time=3.2, end_time=3.6, velocity=80, program=pg, is_drum=is_dm)
kira2.notes.add(pitch=65, start_time=3.6, end_time=4.0, velocity=80, program=pg, is_drum=is_dm)
kira2.notes.add(pitch=64, start_time=4.0, end_time=4.4, velocity=80, program=pg, is_drum=is_dm)
kira2.notes.add(pitch=64, start_time=4.4, end_time=4.8, velocity=80, program=pg, is_drum=is_dm)
kira2.notes.add(pitch=62, start_time=4.8, end_time=5.2, velocity=80, program=pg, is_drum=is_dm)
kira2.notes.add(pitch=62, start_time=5.2, end_time=5.6, velocity=80, program=pg, is_drum=is_dm)
kira2.notes.add(pitch=60, start_time=5.6, end_time=6.4, velocity=80, program=pg, is_drum=is_dm)
note_seq.plot_sequence(kira2) # NoteSequenceの可視化
note_seq.play_sequence(kira2, synth=note_seq.fluidsynth) # NoteSequenceの再生
| section_1/02_exercise.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import matplotlib.pyplot as plt
# %matplotlib inline
from IPython import display
# -
def show_state(env, step=0, info=""):
plt.figure(3)
plt.clf()
plt.imshow(env.render(mode='rgb_array'))
plt.title("s"%(info['descriptions']))
#plt.title("%s | Step: %d %s" % (env._spec.id,step, info))
plt.axis('off')
display.clear_output(wait=True)
display.display(plt.gcf())
# +
import sys
import os
import cv2
from env import ClevrEnv
env = ClevrEnv()
# -
import sys
gym_loc = '/home/don/anaconda3/envs/clearn-pytorch/lib/python3.8/site-packages'
sys.path.append(gym_loc)
import cv2
count = 0
while count < 100:
action = env.sample_random_action()
obs, reward, _, info = env.step(action, update_des=True)
show_state(env, info="")
count += 1
| .ipynb_checkpoints/rendering-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="../images/QISKit-c.gif" alt="Note: In order for images to show up in this jupyter notebook you need to select File => Trusted Notebook" width="250 px" align="left">
# ## _*Single qubit states: amplitude and phase*_
#
# The latest version of this notebook is available on https://github.com/QISKit/qiskit-tutorial.
#
# ***
# ### Contributors
# <NAME>, <NAME>
# ## Introduction
# In [superposition and entanglement](superposition_and_entanglement.ipynb) we looked at the concept of computational states, and superpostion states. Here we continue the development of the understanding of a quantum state by introducing the concept of amplitude and phase.
# ## A qubit
#
# The fundamental unit of quantum information is a qubit. It is a two-dimensional vector space of the complex numbers $\mathbb{C}^2$. A qubit has some similarities to a classical bit, but is overall very different. Like a bit, a qubit can have two possible values - normally a 0 or a 1 which we denote $|0\rangle$ and $|1\rangle$. The difference is that whereas a bit must be either 0 or 1, a qubit can be 0, 1, or a superpostion of both. An arbitrary pure state of the qubit can be written as
#
# $$ |\psi\rangle = \alpha |0\rangle + \beta |1\rangle$$
#
# where $\alpha$ and $\beta$ are complex numbers. When a qubit in the state $|\psi\rangle$ is measured, the result is not deterministic, and one obtains the state $|0\rangle$ with probability $|\alpha|^2$ and the state $|1\rangle$ with probability $|\beta|^2$. This imposes the constrain that $|\alpha|^2 + |\beta|^2=1$ from conservation of probability. We can now define a pure state for a qubit as
#
# $$ |\psi\rangle = \cos(\theta/2)|0\rangle + e^{i\phi} \sin(\theta/2) |1\rangle$$
#
# where $0\leq \phi < 2\pi$ and $0 \leq \theta \leq \pi$.
#
#
# Quantum gates for a qubit are represented as a $2\times2$ unitary matrix $U$. The action on the quantum gate is found by
#
# $$ |\psi'\rangle = U |\psi \rangle $$ and the most general unitary must be able to take $|0\rangle$ to the above state.
# This gives
#
# $$U = \begin{pmatrix} \cos(\theta/2) & a \\ e^{i\phi}\sin(\theta/2) & b \end{pmatrix}$$
#
# where $a$ and $b$ are complex numbers constrained such that $U^\dagger U = I$ for all $0\leq \theta \leq \pi$ and $0\leq\phi < 2\pi$. This gives 3 constraints and as such $a\rightarrow -e^{i\lambda} \sin(\theta/2)$ and $b \rightarrow e^{i\lambda + i \phi} \cos(\theta/2)$ where $0\leq \lambda < 2\pi$
#
# $$U = \begin{pmatrix} \cos(\theta/2) & -e^{i\lambda} \sin(\theta/2) \\ e^{i\phi}\sin(\theta/2) & e^{i\lambda + i \phi} \cos(\theta/2) \end{pmatrix}$$
#
# This allows us to define a general single-qubit rotation as a function of the three parameters $\theta$, $\phi$ and $\lambda$, $U(\theta, \phi, \lambda)$
#
# It is generally useful to understand general single-qubit rotations in terms of a very special group of matrices called the Pauli matrices $X$, $Y$ and $Z$. We already saw in [superposition and entanglement](superposition_and_entanglement.ipynb) that the $X$ Pauli matrix could be defined as
#
# $$ X =\begin{pmatrix} 0 & 1 \\ 1 & 0 \end{pmatrix}$$
#
# Similarly, we can also define
#
# $$ Y =\begin{pmatrix} 0 & -i \\ i & 0 \end{pmatrix}$$
#
# and
#
# $$ Z =\begin{pmatrix} 1 & 0 \\ 0 & -1 \end{pmatrix}$$
#
# An important aspect of the Pauli matrices is that $X^2=Y^2=Z^2=I$. This allows us to simplify
#
# $$e^{i \theta P}= \cos(\theta)I + i\sin(\theta)P$$
#
# with $P$ a Pauli matrix. The Pauli matrices can be thus considered as the generators of rotations in the single-qubit space. We can then define the rotations
#
# $$r_x(\theta) = e^{-i \theta/2 X} = \cos(\theta/2)I - i \sin(\theta/2)X = \begin{pmatrix} \cos(\theta/2) & -i\sin(\theta/2) \\ -i\sin(\theta/2) & \cos(\theta/2) \end{pmatrix}$$
#
# $$r_y(\theta) = e^{-i \theta/2 Y} = \cos(\theta/2)I - i \sin(\theta/2)Y = \begin{pmatrix} \cos(\theta/2) & -\sin(\theta/2) \\ \sin(\theta/2) & \cos(\theta/2) \end{pmatrix}$$
#
# $$r_z(\theta) = e^{-i \theta/2 Z} = \cos(\theta/2)I - i \sin(\theta/2)Z = \begin{pmatrix} e^{-i\theta/2} & 0 \\ 0 & e^{i\theta/2} \end{pmatrix} := \begin{pmatrix} 1 & 0 \\ 0 & e^{i\theta} \end{pmatrix}$$
#
# And now, in terms of the previously defined $U(\theta, \phi, \lambda)$, we can define the parametrized rotations
#
# $$u_3(\theta, \phi, \lambda) = U(\theta, \phi, \lambda) = r_z(\phi)r_y(\theta)r_z(\lambda) = \begin{pmatrix} e^{-i(\phi + \lambda)/2}\cos(\theta/2) & -e^{-i(\phi - \lambda)/2} \sin(\theta/2) \\ e^{i(\phi-\lambda)/2}\sin(\theta/2) & e^{i(\phi + \lambda)/2} \cos(\theta/2) \end{pmatrix}$$
#
# where we have multiplied the above defined $U(\theta, \phi, \lambda)$ by a global phase $e^{-i(\phi + \lambda)/2}$
#
# Similarly, we can define the parametrized rotations
#
# $$u_2(\phi, \lambda) = U(\pi/2, \phi, \lambda)$$
#
# and
#
# $$u_1(\lambda) = U(0,0,\lambda) := r_z(\lambda)$$
# ## Phase gates
#
# Let's explore the action of some of the single-qubit gates we have just defined. We will start with the phase gate $u_1(\lambda)$. We can see from the gate matrix that this gate does not modify the state $|0\rangle$ and applies a phase of $\lambda$ to the state $|1\rangle$. In order for us to observe the effect of this gate on a qubit we need to make use of the Hadamard gate defined in [superposition and entanglement](superposition_and_entanglement.ipynb).
# +
# useful additional packages
import matplotlib.pyplot as plt
# %matplotlib inline
import numpy as np
# importing the QISKit
from qiskit import QuantumProgram
import Qconfig
# import basic plot tools
from qiskit.tools.visualization import plot_histogram
# -
# Following a Hadamard gate, we will perform a $u_1(\lambda)$ of varying angle $\lambda \in \{ - \pi, \pi \}$ and observe the qubit state oscillate between the states $|+\rangle$ ($X$ eigenvalue +1) and $|-\rangle$ ($X$ eigenvalue -1).
# +
backend = 'local_qasm_simulator' # the backend to run on
shots = 1024 # the number of shots in the experiment
Q_program = QuantumProgram()
Q_program.set_api(Qconfig.APItoken, Qconfig.config["url"]) # set the APIToken and API url
# Creating registers
qr = Q_program.create_quantum_register("qr", 1)
cr = Q_program.create_classical_register("cr", 1)
circuits = []
phase_vector = range(0,100)
for phase_index in phase_vector:
phase_shift = phase_index-50
phase = 2*np.pi*phase_shift/50
circuit_name = "phase_gate_%d"%phase_index
qc_phase_gate = Q_program.create_circuit(circuit_name, [qr], [cr])
qc_phase_gate.h(qr)
qc_phase_gate.u1(phase, qr)
qc_phase_gate.h(qr)
qc_phase_gate.measure(qr[0], cr[0])
circuits.append(circuit_name)
result = Q_program.execute(circuits, backend=backend, shots=shots, max_credits=3, wait=10, timeout=240)
# -
probz = []
phase_value = []
for phase_index in phase_vector:
phase_shift = phase_index - 50
phase_value.append(2*phase_shift/50)
if '0' in result.get_counts(circuits[phase_index]):
probz.append(2*result.get_counts(circuits[phase_index]).get('0')/shots-1)
else:
probz.append(-1)
# +
plt.plot(phase_value, probz, 'b',0.25,1/np.sqrt(2),'ro',0.5,0,'ko',1,-1,'go',-0.25,1/np.sqrt(2),'rx',-0.5,0,'kx',-1,-1,'gx')
plt.xlabel('Phase value (Pi)')
plt.ylabel('Eigenvalue of X')
plt.show()
# -
# We can see the eigenstate of $X$ oscillating between +1 and -1. The six symbols in the plot correspond to special gates that we use as part of our universal set of gates. These are $T$ ($T^{\dagger}$) for the red dot (cross), $S$ ($S^{\dagger}$) for the black dot (cross) and $Z$ ($Z^{\dagger}$) for the green dot (cross). Therefore, we can see that $T=u_1(\pi/4)$, $S=u_1(\pi/2)$ and $Z=u_1(\pi)$, with their adjoints corresponding to the negative arguments.
#
# Now let's have a look at how the unitaries $u_3$ operate on a qubit state.
# +
backend = 'local_qasm_simulator' # the backend to run on
shots = 1024 # the number of shots in the experiment
Q_program = QuantumProgram()
Q_program.set_api(Qconfig.APItoken, Qconfig.config["url"]) # set the APIToken and API url
# Creating registers
qr = Q_program.create_quantum_register("qr", 1)
cr = Q_program.create_classical_register("cr", 1)
circuits = []
phase_vector = range(0,100)
for phase_index in phase_vector:
phase_shift = phase_index-50
phase = 2*np.pi*phase_shift/50
circuit_name = "phase_gate_%d"%phase_index
qc_phase_gate = Q_program.create_circuit(circuit_name, [qr], [cr])
qc_phase_gate.u3(phase,0,np.pi, qr)
qc_phase_gate.measure(qr[0], cr[0])
circuits.append(circuit_name)
result = Q_program.execute(circuits, backend=backend, shots=shots, max_credits=3, wait=10, timeout=240)
# -
probz = []
phase_value = []
for phase_index in phase_vector:
phase_shift = phase_index - 50
phase_value.append(2*phase_shift/50)
if '0' in result.get_counts(circuits[phase_index]):
probz.append(2*result.get_counts(circuits[phase_index]).get('0')/shots-1)
else:
probz.append(-1)
# +
plt.plot(phase_value, probz, 'b',0.5,0,'ko',1,-1,'go',-0.5,0,'kx',-1,-1,'gx')
plt.xlabel('Phase value (Pi)')
plt.ylabel('Eigenvalue of Z')
plt.show()
# -
# In this case, we see that $u_3(\pi/2,0,\pi)$ corresponds to a $\pi/2$ rotation around $X$ (black dot) and $u_3(\pi,0,\pi)$ corresponds to a bit-flip around $X$ (green dot). The rotations in the negative directions are shown as crosses. We could also have implemented these rotations around the $Y$ axis by simply using $u_3(\lambda,\pi/2,\pi/2)$.
#
# As an exercise, prove that the Hadamard gate can be implemented as $H = u_2(0, \pi)$.
# %run "../version.ipynb"
| appendix/more_qis/single_qubit_states_amplitude_and_phase.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <EMAIL> © 2021
# ## Minimum of two random variables
# Question: Assume we have $X \sim Uniform(0, 1)$ and $Y \sim Uniform(0,1)$ and independent of each other. What is the expected value of the minimum of $X$ and $Y$?
#
# ### Intuition
# Before going to show the exact theoretical result, we can think heuristically,we have two uniform random variables that are independent of each other, we would expect two of these random variables to divide the interval (0,1) equally into 3 equal subintervals. Then the minimum would occur at 1/3 and the other one (which is maximum in this case) occurs at 2/3. We can even generalize our intuition for $n$ independent uniform random variables. In that case minimum would occur at $1/(n+1)$ and maximum would occur at $n/(n+1)$. Now let us verify intution theoretically.
# ### Theoritical result
# Let $Z$ be minimum of X and Y. We write this as $Z=\min(X,Y)$.
# \begin{equation}
# \mathbb{P}(Z\leq z)= \mathbb{P}(\min(X,Y)\leq z) =1-\mathbb{P}(\min(X,Y)\geq z)=1-\mathbb{P}(X> z, Y>z)
# \end{equation}
# Since our distribution is between 0 and 1 the following is true for uniform distribution
# \begin{equation}
# \mathbb{P}(X\leq z)= z \text{ and } \mathbb{P}(X> z)=1- z
# \end{equation}
# Also same goes for $Y$. Now since they are independent we have
# \begin{equation}
# \mathbb{P}(X> z, Y>z)=\mathbb{P}(X> z)\mathbb{P}(Y>z)=(1-z)^{2}
# \end{equation}
# Then equation (1) becomes
# \begin{equation}
# \mathbb{P}(Z\leq z)=1-\mathbb{P}(X> z, Y>z)=1-(1-z)^{2}
# \end{equation}
# We just calculated cumulative distribution function of $z$. Usually denoted as
# \begin{equation}
# F_{Z}(z)= \mathbb{P}(Z\leq z)
# \end{equation}
# If we take derivative of this $ F_{Z}(z)$, we will get density function of z. In this case it would be
# \begin{equation}
# F_{Z}'(z)=f_Z(z)=2(1-z)
# \end{equation}
# As last part we would take integral of $zf_Z(z)$ between 0 and 1 to find an expected value of minimum of two uniform random variables.
# \begin{equation}
# \mathbb{E}[Z]=\int_{0}^1 zf_Z(z)dz=\int_{0}^1 2z(1-z)dz=2\left(\frac{1}{2}-\frac{1}{3}\right)=\frac{1}{3}
# \end{equation}
# If you are allowed to use any programming language then you can simulate.
import numpy as np
x = np.random.uniform(0,1,100000)
y = np.random.uniform(0,1,100000)
z =np.minimum(x,y)
u =np.maximum(x,y)
np.mean(z), np.mean(u) # as you can see z is very close to 1/3
# ### To see the histogram of Z and U
import matplotlib.pyplot as plt
count, bins, ignored = plt.hist(z, 100, density=True)
plt.plot(bins, np.ones_like(bins), linewidth=2, color='r')
plt.show()
import matplotlib.pyplot as plt
count, bins, ignored = plt.hist(u, 100, density=True)
plt.plot(bins, np.ones_like(bins), linewidth=2, color='r')
plt.show()
| Minimim of two random variables March 3 2021.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/mrdbourke/tensorflow-deep-learning/blob/main/00_tensorflow_fundamentals.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="ielzxn52r2CB"
# ## Getting started with TensorFlow: A guide to the fundamentals
#
# ## What is TensorFlow?
#
# [TensorFlow](https://www.tensorflow.org/) is an open-source end-to-end machine learning library for preprocessing data, modelling data and serving models (getting them into the hands of others).
#
# ## Why use TensorFlow?
#
# Rather than building machine learning and deep learning models from scratch, it's more likely you'll use a library such as TensorFlow. This is because it contains many of the most common machine learning functions you'll want to use.
#
# ## What we're going to cover
#
# TensorFlow is vast. But the main premise is simple: turn data into numbers (tensors) and build machine learning algorithms to find patterns in them.
#
# In this notebook we cover some of the most fundamental TensorFlow operations, more specificially:
# * Introduction to tensors (creating tensors)
# * Getting information from tensors (tensor attributes)
# * Manipulating tensors (tensor operations)
# * Tensors and NumPy
# * Using @tf.function (a way to speed up your regular Python functions)
# * Using GPUs with TensorFlow
# * Exercises to try
#
# Things to note:
# * Many of the conventions here will happen automatically behind the scenes (when you build a model) but it's worth knowing so if you see any of these things, you know what's happening.
# * For any TensorFlow function you see, it's important to be able to check it out in the documentation, for example, going to the Python API docs for all functions and searching for what you need: https://www.tensorflow.org/api_docs/python/ (don't worry if this seems overwhelming at first, with enough practice, you'll get used to navigating the documentaiton).
#
#
# + [markdown] id="VArmDlu06sH0"
# ## Introduction to Tensors
#
# If you've ever used NumPy, [tensors](https://www.tensorflow.org/guide/tensor) are kind of like NumPy arrays (we'll see more on this later).
#
# For the sake of this notebook and going forward, you can think of a tensor as a multi-dimensional numerical representation (also referred to as n-dimensional, where n can be any number) of something. Where something can be almost anything you can imagine:
# * It could be numbers themselves (using tensors to represent the price of houses).
# * It could be an image (using tensors to represent the pixels of an image).
# * It could be text (using tensors to represent words).
# * Or it could be some other form of information (or data) you want to represent with numbers.
#
# The main difference between tensors and NumPy arrays (also an n-dimensional array of numbers) is that tensors can be used on [GPUs (graphical processing units)](https://blogs.nvidia.com/blog/2009/12/16/whats-the-difference-between-a-cpu-and-a-gpu/) and [TPUs (tensor processing units)](https://en.wikipedia.org/wiki/Tensor_processing_unit).
#
# The benefit of being able to run on GPUs and TPUs is faster computation, this means, if we wanted to find patterns in the numerical representations of our data, we can generally find them faster using GPUs and TPUs.
#
# Okay, we've been talking enough about tensors, let's see them.
#
# The first thing we'll do is import TensorFlow under the common alias `tf`.
# + colab={"base_uri": "https://localhost:8080/"} id="Z7ieIu8t9ijY" outputId="7548088b-91d5-412f-8544-3f6b3487e33a"
# Import TensorFlow
import tensorflow as tf
print(tf.__version__) # find the version number (should be 2.x+)
# + [markdown] id="JNtV5t1qz0VP"
# ### Creating Tensors with `tf.constant()`
#
# As mentioned before, in general, you usually won't create tensors yourself. This is because TensorFlow has modules built-in (such as [`tf.io`](https://www.tensorflow.org/api_docs/python/tf/io) and [`tf.data`](https://www.tensorflow.org/guide/data)) which are able to read your data sources and automatically convert them to tensors and then later on, neural network models will process these for us.
#
# But for now, because we're getting familar with tensors themselves and how to manipulate them, we'll see how we can create them ourselves.
#
# We'll begin by using [`tf.constant()`](https://www.tensorflow.org/api_docs/python/tf/constant).
# + colab={"base_uri": "https://localhost:8080/"} id="nC7aQgqi0M_Z" outputId="89250b3e-86a9-4d9b-b15f-8ae0812dde5b"
# Create a scalar (rank 0 tensor)
scalar = tf.constant(7)
scalar
# + [markdown] id="o6fXE5dXkO_3"
# A scalar is known as a rank 0 tensor. Because it has no dimensions (it's just a number).
#
# > 🔑 **Note:** For now, you don't need to know too much about the different ranks of tensors (but we will see more on this later). The important point is knowing tensors can have an unlimited range of dimensions (the exact amount will depend on what data you're representing).
# + colab={"base_uri": "https://localhost:8080/"} id="1sgUNKoFkJ21" outputId="5b244f42-05bc-4702-dc5c-a91716443c4c"
# Check the number of dimensions of a tensor (ndim stands for number of dimensions)
scalar.ndim
# + colab={"base_uri": "https://localhost:8080/"} id="irtCo2fs0V_o" outputId="e424c961-7a1a-4544-ed5a-becc1cc1d1bd"
# Create a vector (more than 0 dimensions)
vector = tf.constant([10, 10])
vector
# + colab={"base_uri": "https://localhost:8080/"} id="7DDc36pvmOse" outputId="f6fbc397-5058-4f2f-8f27-9d7c6adf4f95"
# Check the number of dimensions of our vector tensor
vector.ndim
# + colab={"base_uri": "https://localhost:8080/"} id="HXf5A5360V7A" outputId="3f773144-81c5-49d2-819b-192ff0153c03"
# Create a matrix (more than 1 dimension)
matrix = tf.constant([[10, 7],
[7, 10]])
matrix
# + colab={"base_uri": "https://localhost:8080/"} id="Asmn6YghlT6u" outputId="aa08a360-2917-4be3-e263-b4cea62ed569"
matrix.ndim
# + [markdown] id="qvQb7RT2s9Te"
# By default, TensorFlow creates tensors with either an `int32` or `float32` datatype.
#
# This is known as [32-bit precision](https://en.wikipedia.org/wiki/Precision_(computer_science) (the higher the number, the more precise the number, the more space it takes up on your computer).
# + colab={"base_uri": "https://localhost:8080/"} id="aEgthLq80V2u" outputId="f64d9ac7-1a83-4fc3-9511-a02e5e19dd6b"
# Create another matrix and define the datatype
another_matrix = tf.constant([[10., 7.],
[3., 2.],
[8., 9.]], dtype=tf.float16) # specify the datatype with 'dtype'
another_matrix
# + colab={"base_uri": "https://localhost:8080/"} id="v-Y-lXdOlXRg" outputId="7f7c851b-9487-4ce8-de4d-5a747b9912e3"
# Even though another_matrix contains more numbers, its dimensions stay the same
another_matrix.ndim
# + colab={"base_uri": "https://localhost:8080/"} id="fAy7J6fT0Vwz" outputId="77c91572-9b0a-4e0d-d39f-91455f41ee1e"
# How about a tensor? (more than 2 dimensions, although, all of the above items are also technically tensors)
tensor = tf.constant([[[1, 2, 3],
[4, 5, 6]],
[[7, 8, 9],
[10, 11, 12]],
[[13, 14, 15],
[16, 17, 18]]])
tensor
# + colab={"base_uri": "https://localhost:8080/"} id="FhIsj108mFOS" outputId="64bf5633-c17f-42b7-ce0a-14e3df8a2dec"
tensor.ndim
# + [markdown] id="w5MGwSpA100u"
# This is known as a rank 3 tensor (3-dimensions), however a tensor can have an arbitrary (unlimited) amount of dimensions.
#
# For example, you might turn a series of images into tensors with shape (224, 224, 3, 32), where:
# * 224, 224 (the first 2 dimensions) are the height and width of the images in pixels.
# * 3 is the number of colour channels of the image (red, green blue).
# * 32 is the batch size (the number of images a neural network sees at any one time).
#
# All of the above variables we've created are actually tensors. But you may also hear them referred to as their different names (the ones we gave them):
# * **scalar**: a single number.
# * **vector**: a number with direction (e.g. wind speed with direction).
# * **matrix**: a 2-dimensional array of numbers.
# * **tensor**: an n-dimensional arrary of numbers (where n can be any number, a 0-dimension tensor is a scalar, a 1-dimension tensor is a vector).
#
# To add to the confusion, the terms matrix and tensor are often used interchangably.
#
# Going forward since we're using TensorFlow, everything we refer to and use will be tensors.
#
# For more on the mathematical difference between scalars, vectors and matrices see the [visual algebra post by Math is Fun](https://www.mathsisfun.com/algebra/scalar-vector-matrix.html).
#
# 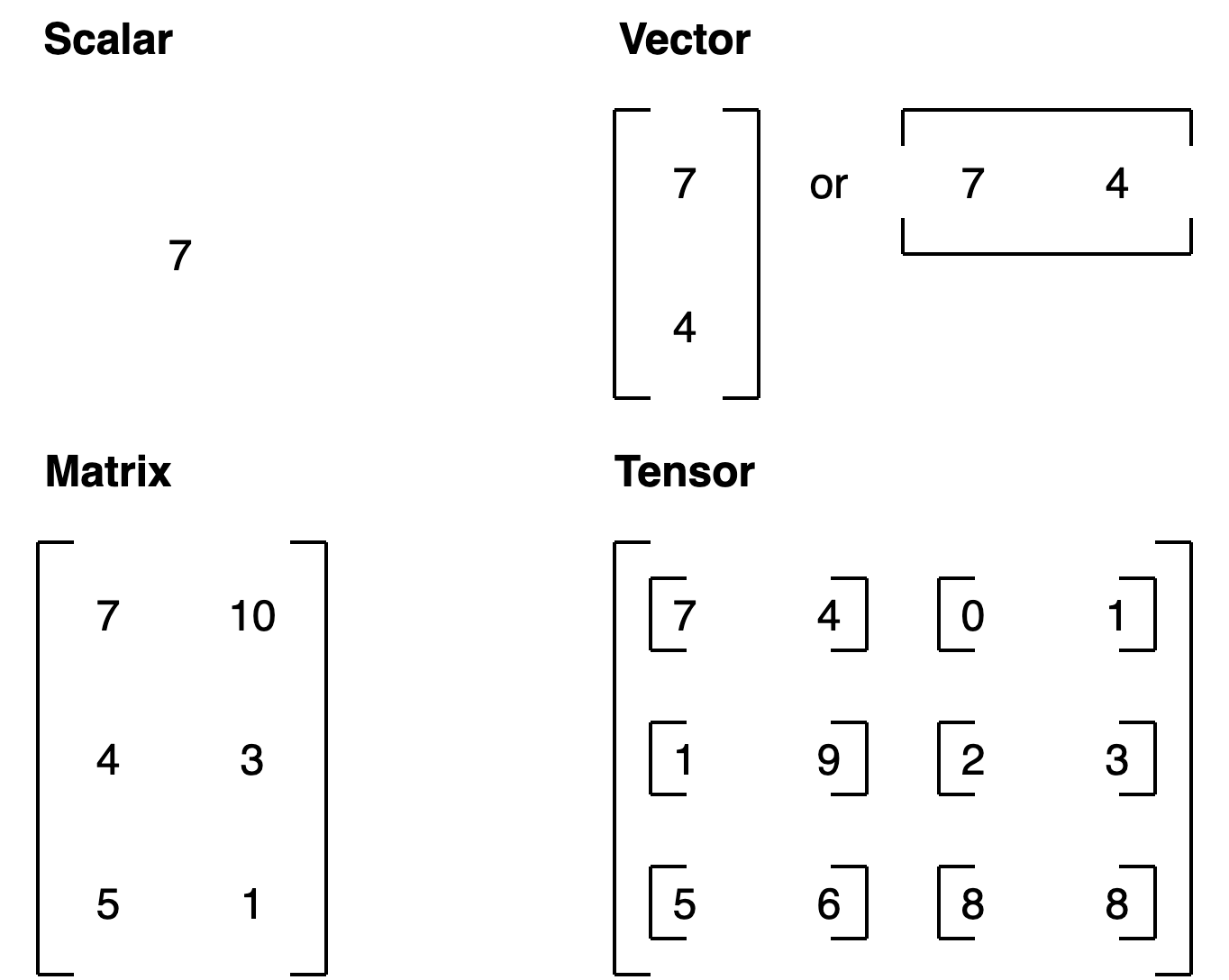
# + [markdown] id="TZMfDFKC0Cl7"
# ### Creating Tensors with `tf.Variable()`
#
# You can also (although you likely rarely will, because often, when working with data, tensors are created for you automatically) create tensors using [`tf.Variable()`](https://www.tensorflow.org/api_docs/python/tf/Variable).
#
# The difference between `tf.Variable()` and `tf.constant()` is tensors created with `tf.constant()` are immutable (can't be changed, can only be used to create a new tensor), where as, tensors created with `tf.Variable()` are mutable (can be changed).
# + colab={"base_uri": "https://localhost:8080/"} id="bv1SBbDe4TxN" outputId="2af43a4e-9d28-4788-f5f9-ab33c95a6551"
# Create the same tensor with tf.Variable() and tf.constant()
changeable_tensor = tf.Variable([10, 7])
unchangeable_tensor = tf.constant([10, 7])
changeable_tensor, unchangeable_tensor
# + [markdown] id="A4WrQ8c2ux15"
# Now let's try to change one of the elements of the changable tensor.
# + colab={"base_uri": "https://localhost:8080/", "height": 200} id="dfDwbF6i5Sy3" outputId="087b7aa3-d371-4741-d798-9aef73895635"
# Will error (requires the .assign() method)
changeable_tensor[0] = 7
changeable_tensor
# + [markdown] id="oWP-kZEVvGm8"
# To change an element of a `tf.Variable()` tensor requires the `assign()` method.
# + colab={"base_uri": "https://localhost:8080/"} id="FJV3iwvG4jg4" outputId="d110b678-03ff-49e8-dee6-9974ab75ba24"
# Won't error
changeable_tensor[0].assign(7)
changeable_tensor
# + [markdown] id="9UiV1Z0XvZ_B"
# Now let's try to change a value in a `tf.constant()` tensor.
# + colab={"base_uri": "https://localhost:8080/", "height": 200} id="5j_rOo8X5N9f" outputId="095cb0dc-dda0-41bc-e8dc-995934535957"
# Will error (can't change tf.constant())
unchangeable_tensor[0].assign(7)
unchangleable_tensor
# + [markdown] id="1t21IcYpverQ"
# Which one should you use? `tf.constant()` or `tf.Variable()`?
#
# It will depend on what your problem requires. However, most of the time, TensorFlow will automatically choose for you (when loading data or modelling data).
# + [markdown] id="pAAT59Ay0J0l"
# ### Creating random tensors
#
# Random tensors are tensors of some abitrary size which contain random numbers.
#
# Why would you want to create random tensors?
#
# This is what neural networks use to intialize their weights (patterns) that they're trying to learn in the data.
#
# For example, the process of a neural network learning often involves taking a random n-dimensional array of numbers and refining them until they represent some kind of pattern (a compressed way to represent the original data).
#
# **How a network learns**
# 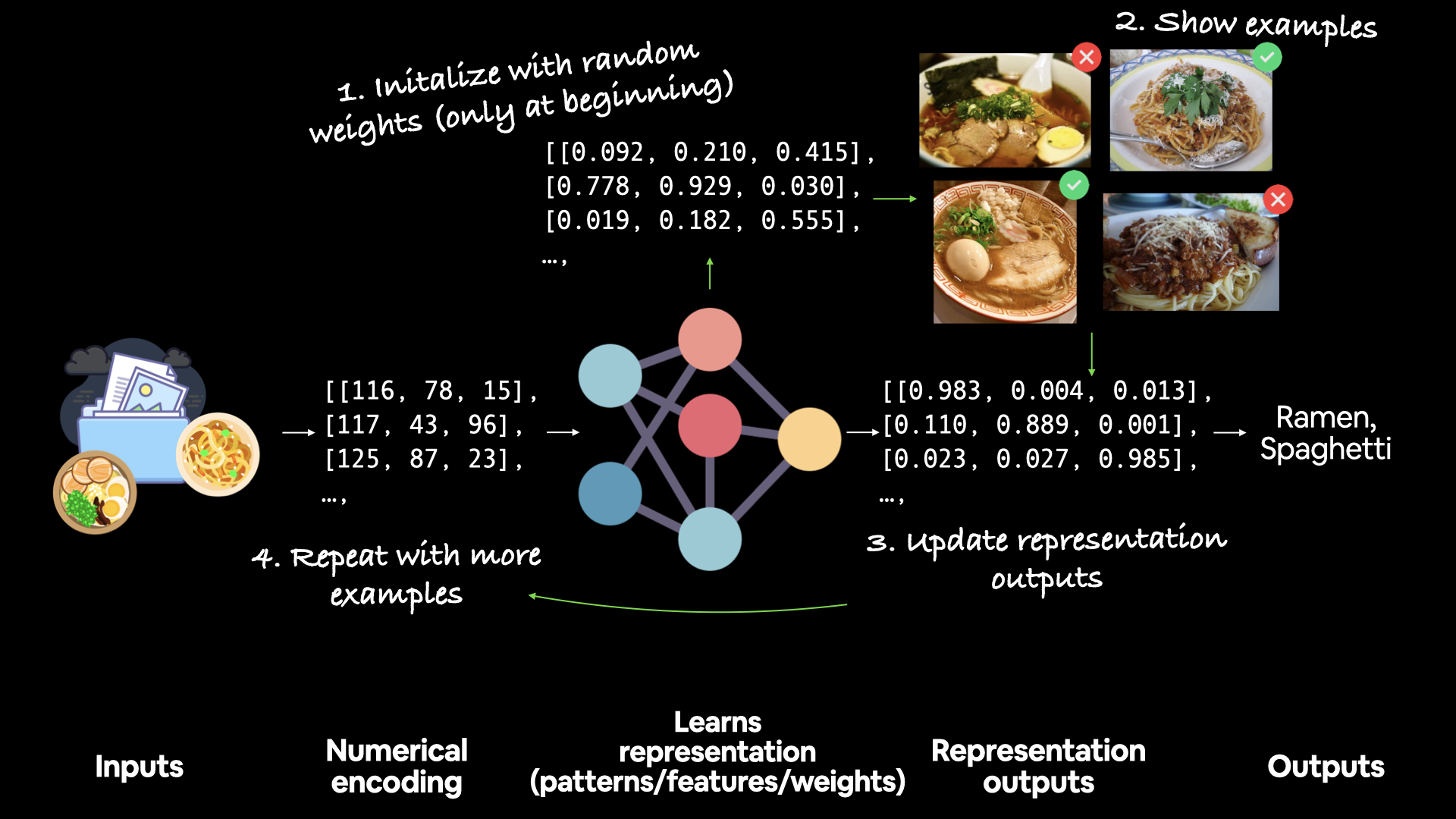
# *A network learns by starting with random patterns (1) then going through demonstrative examples of data (2) whilst trying to update its random patterns to represent the examples (3).*
#
# We can create random tensors by using the [`tf.random.Generator`](https://www.tensorflow.org/guide/random_numbers#the_tfrandomgenerator_class) class.
# + colab={"base_uri": "https://localhost:8080/"} id="yZ7Zu5Z178JL" outputId="79b4497d-0404-4171-f42f-050c854b9019"
# Create two random (but the same) tensors
random_1 = tf.random.Generator.from_seed(42) # set the seed for reproducibility
random_1 = random_1.normal(shape=(3, 2)) # create tensor from a normal distribution
random_2 = tf.random.Generator.from_seed(42)
random_2 = random_2.normal(shape=(3, 2))
# Are they equal?
random_1, random_2, random_1 == random_2
# + [markdown] id="k6Od5fpZ-S--"
# The random tensors we've made are actually [pseudorandom numbers](https://www.computerhope.com/jargon/p/pseudo-random.htm) (they appear as random, but really aren't).
#
# If we set a seed we'll get the same random numbers (if you've ever used NumPy, this is similar to `np.random.seed(42)`).
#
# Setting the seed says, "hey, create some random numbers, but flavour them with X" (X is the seed).
#
# What do you think will happen when we change the seed?
# + colab={"base_uri": "https://localhost:8080/"} id="9eStLqr1F4ZP" outputId="cf545015-ce96-46dd-9d43-fbb148a04782"
# Create two random (and different) tensors
random_3 = tf.random.Generator.from_seed(42)
random_3 = random_3.normal(shape=(3, 2))
random_4 = tf.random.Generator.from_seed(11)
random_4 = random_4.normal(shape=(3, 2))
# Check the tensors and see if they are equal
random_3, random_4, random_1 == random_3, random_3 == random_4
# + [markdown] id="Nji9AdFRIhBi"
# What if you wanted to shuffle the order of a tensor?
#
# Wait, why would you want to do that?
#
# Let's say you working with 15,000 images of cats and dogs and the first 10,000 images of were of cats and the next 5,000 were of dogs. This order could effect how a neural network learns (it may overfit by learning the order of the data), instead, it might be a good idea to move your data around.
# + colab={"base_uri": "https://localhost:8080/"} id="sl4HYEWMBI6x" outputId="195d45c3-9c2e-4e7d-ea71-f3382b4dc6b0"
# Shuffle a tensor (valuable for when you want to shuffle your data)
not_shuffled = tf.constant([[10, 7],
[3, 4],
[2, 5]])
# Gets different results each time
tf.random.shuffle(not_shuffled)
# + colab={"base_uri": "https://localhost:8080/"} id="-HYn0ME_H1SY" outputId="1a5c9383-9ca8-47c2-90a8-d2002dbf4ef8"
# Shuffle in the same order every time using the seed parameter (won't acutally be the same)
tf.random.shuffle(not_shuffled, seed=42)
# + [markdown] id="GmC3qGIHjAx6"
# Wait... why didn't the numbers come out the same?
#
# It's due to rule #4 of the [`tf.random.set_seed()`](https://www.tensorflow.org/api_docs/python/tf/random/set_seed) documentation.
#
# > "4. If both the global and the operation seed are set: Both seeds are used in conjunction to determine the random sequence."
#
# `tf.random.set_seed(42)` sets the global seed, and the `seed` parameter in `tf.random.shuffle(seed=42)` sets the operation seed.
#
# Because, "Operations that rely on a random seed actually derive it from two seeds: the global and operation-level seeds. This sets the global seed."
#
# + colab={"base_uri": "https://localhost:8080/"} id="cM6S8set-ixV" outputId="3ab2f553-5818-46eb-d276-a14853693bdc"
# Shuffle in the same order every time
# Set the global random seed
tf.random.set_seed(42)
# Set the operation random seed
tf.random.shuffle(not_shuffled, seed=42)
# + colab={"base_uri": "https://localhost:8080/"} id="xKJOsdE8yCn4" outputId="9cd2788b-95a7-4f31-997c-313a2297541d"
# Set the global random seed
tf.random.set_seed(42) # if you comment this out you'll get different results
# Set the operation random seed
tf.random.shuffle(not_shuffled)
# + [markdown] id="ouZ1fpJk0R3h"
# ### Other ways to make tensors
#
# Though you might rarely use these (remember, many tensor operations are done behind the scenes for you), you can use [`tf.ones()`](https://www.tensorflow.org/api_docs/python/tf/ones) to create a tensor of all ones and [`tf.zeros()`](https://www.tensorflow.org/api_docs/python/tf/zeros) to create a tensor of all zeros.
# + colab={"base_uri": "https://localhost:8080/"} id="aG8QNZP7kEe1" outputId="9ab2ec05-8b1d-45b0-a886-392e63d69149"
# Make a tensor of all ones
tf.ones(shape=(3, 2))
# + colab={"base_uri": "https://localhost:8080/"} id="GQKiWrB9kprj" outputId="8cd8ccee-2546-4473-ca89-0789100fbc1c"
# Make a tensor of all zeros
tf.zeros(shape=(3, 2))
# + [markdown] id="slcLTK5D7kc2"
# You can also turn NumPy arrays in into tensors.
#
# Remember, the main difference between tensors and NumPy arrays is that tensors can be run on GPUs.
#
# > 🔑 **Note:** A matrix or tensor is typically represented by a capital letter (e.g. `X` or `A`) where as a vector is typically represented by a lowercase letter (e.g. `y` or `b`).
# + colab={"base_uri": "https://localhost:8080/"} id="C0XP37xi7mn4" outputId="f59af147-58d8-4239-bbc1-1f140a858118"
import numpy as np
numpy_A = np.arange(1, 25, dtype=np.int32) # create a NumPy array between 1 and 25
A = tf.constant(numpy_A,
shape=[2, 4, 3]) # note: the shape total (2*4*3) has to match the number of elements in the array
numpy_A, A
# + [markdown] id="J1JLXa2P0wpx"
# ## Getting information from tensors (shape, rank, size)
#
# There will be times when you'll want to get different pieces of information from your tensors, in particuluar, you should know the following tensor vocabulary:
# * **Shape:** The length (number of elements) of each of the dimensions of a tensor.
# * **Rank:** The number of tensor dimensions. A scalar has rank 0, a vector has rank 1, a matrix is rank 2, a tensor has rank n.
# * **Axis** or **Dimension:** A particular dimension of a tensor.
# * **Size:** The total number of items in the tensor.
#
# You'll use these especially when you're trying to line up the shapes of your data to the shapes of your model. For example, making sure the shape of your image tensors are the same shape as your models input layer.
#
# We've already seen one of these before using the `ndim` attribute. Let's see the rest.
# + colab={"base_uri": "https://localhost:8080/"} id="qhckrmovCaAA" outputId="ac4307f8-d850-4c45-eaff-7cfa8a3919e9"
# Create a rank 4 tensor (4 dimensions)
rank_4_tensor = tf.zeros([2, 3, 4, 5])
rank_4_tensor
# + colab={"base_uri": "https://localhost:8080/"} id="ImJdhWnLtZ_2" outputId="92b9e814-51b1-4374-ae81-516dcc2e0c49"
rank_4_tensor.shape, rank_4_tensor.ndim, tf.size(rank_4_tensor)
# + colab={"base_uri": "https://localhost:8080/"} id="Vvb-4ZYdpI9f" outputId="0f1b35eb-b93f-422e-8979-17260c621ffb"
# Get various attributes of tensor
print("Datatype of every element:", rank_4_tensor.dtype)
print("Number of dimensions (rank):", rank_4_tensor.ndim)
print("Shape of tensor:", rank_4_tensor.shape)
print("Elements along axis 0 of tensor:", rank_4_tensor.shape[0])
print("Elements along last axis of tensor:", rank_4_tensor.shape[-1])
print("Total number of elements (2*3*4*5):", tf.size(rank_4_tensor).numpy()) # .numpy() converts to NumPy array
# + [markdown] id="S0SMO2ZOqL0G"
# You can also index tensors just like Python lists.
# + colab={"base_uri": "https://localhost:8080/"} id="CFzOo-7QqLJf" outputId="2629d691-a8de-4488-915a-b06868f5218d"
# Get the first 2 items of each dimension
rank_4_tensor[:2, :2, :2, :2]
# + colab={"base_uri": "https://localhost:8080/"} id="weQe2bBUqknd" outputId="16c61a8c-2ba3-42a3-d4b8-d19185553300"
# Get the dimension from each index except for the final one
rank_4_tensor[:1, :1, :1, :]
# + colab={"base_uri": "https://localhost:8080/"} id="YQKcZWz5rFXG" outputId="15a0bee6-6467-46b7-c315-e5ec963de8bd"
# Create a rank 2 tensor (2 dimensions)
rank_2_tensor = tf.constant([[10, 7],
[3, 4]])
# Get the last item of each row
rank_2_tensor[:, -1]
# + [markdown] id="cLZq3sHKrVdS"
# You can also add dimensions to your tensor whilst keeping the same information present using `tf.newaxis`.
# + colab={"base_uri": "https://localhost:8080/"} id="KuEEEQa4w1id" outputId="998dcc25-889f-4c5b-e417-f9e45dcaa350"
# Add an extra dimension (to the end)
rank_3_tensor = rank_2_tensor[..., tf.newaxis] # in Python "..." means "all dimensions prior to"
rank_2_tensor, rank_3_tensor # shape (2, 2), shape (2, 2, 1)
# + [markdown] id="a5_KyB-6r7z7"
# You can achieve the same using [`tf.expand_dims()`](https://www.tensorflow.org/api_docs/python/tf/expand_dims).
# + colab={"base_uri": "https://localhost:8080/"} id="HpPTBqt4rvr9" outputId="47a95be1-ba16-475f-b2cd-<KEY>"
tf.expand_dims(rank_2_tensor, axis=-1) # "-1" means last axis
# + [markdown] id="4EHae9iA04Ok"
# ## Manipulating tensors (tensor operations)
#
# Finding patterns in tensors (numberical representation of data) requires manipulating them.
#
# Again, when building models in TensorFlow, much of this pattern discovery is done for you.
# + [markdown] id="-NzdbYDqs1Ex"
# ### Basic operations
#
# You can perform many of the basic mathematical operations directly on tensors using Pyhton operators such as, `+`, `-`, `*`.
# + colab={"base_uri": "https://localhost:8080/"} id="tu3zJirLsMVw" outputId="7dce405b-292e-4d7a-efdd-eeda89807887"
# You can add values to a tensor using the addition operator
tensor = tf.constant([[10, 7], [3, 4]])
tensor + 10
# + [markdown] id="_smxbIkYwYY3"
# Since we used `tf.constant()`, the original tensor is unchanged (the addition gets done on a copy).
# + colab={"base_uri": "https://localhost:8080/"} id="BhJn3puhwOlM" outputId="92ce1cbe-e4c8-4f4e-c50b-c21d4ea57d43"
# Original tensor unchanged
tensor
# + [markdown] id="5N6tUM16xq9d"
# Other operators also work.
# + colab={"base_uri": "https://localhost:8080/"} id="6TW0_ZC_xoEC" outputId="f6aa4f99-a9e8-4e13-c06c-5c9450ef99c4"
# Multiplication (known as element-wise multiplication)
tensor * 10
# + colab={"base_uri": "https://localhost:8080/"} id="MN6XjwWfxu66" outputId="ca0a775c-e2e3-4a53-fbae-e4ced3f2c91c"
# Subtraction
tensor - 10
# + [markdown] id="1kMJe4GlyuZR"
# You can also use the equivalent TensorFlow function. Using the TensorFlow function (where possible) has the advantage of being sped up later down the line when running as part of a [TensorFlow graph](https://www.tensorflow.org/tensorboard/graphs).
# + colab={"base_uri": "https://localhost:8080/"} id="R2NDjqYIyyMc" outputId="b811b7c6-2c4a-4f89-fae6-89ceaf8dfa5e"
# Use the tensorflow function equivalent of the '*' (multiply) operator
tf.multiply(tensor, 10)
# + colab={"base_uri": "https://localhost:8080/"} id="lKEuDBFD49w7" outputId="99300b75-0f81-432b-f3d0-b32e4c651b5b"
# The original tensor is still unchanged
tensor
# + [markdown] id="8M-0dhjtzHoZ"
# ### Matrix mutliplication
#
# One of the most common operations in machine learning algorithms is [matrix multiplication](https://www.mathsisfun.com/algebra/matrix-multiplying.html).
#
# TensorFlow implements this matrix multiplication functionality in the [`tf.matmul()`](https://www.tensorflow.org/api_docs/python/tf/linalg/matmul) method.
#
# The main two rules for matrix multiplication to remember are:
# 1. The inner dimensions must match:
# * `(3, 5) @ (3, 5)` won't work
# * `(5, 3) @ (3, 5)` will work
# * `(3, 5) @ (5, 3)` will work
# 2. The resulting matrix has the shape of the outer dimensions:
# * `(5, 3) @ (3, 5)` -> `(5, 5)`
# * `(3, 5) @ (5, 3)` -> `(3, 3)`
#
# > 🔑 **Note:** '`@`' in Python is the symbol for matrix multiplication.
# + colab={"base_uri": "https://localhost:8080/"} id="pbpwVJrAsPpA" outputId="580b001a-f0c1-45ef-9918-cc5538b0fb1e"
# Matrix multiplication in TensorFlow
print(tensor)
tf.matmul(tensor, tensor)
# + colab={"base_uri": "https://localhost:8080/"} id="9vpDnpb10G7U" outputId="6693b6f2-e529-41f3-b08b-c19a90df1137"
# Matrix multiplication with Python operator '@'
tensor @ tensor
# + [markdown] id="TAV3S5YV0cDQ"
# Both of these examples work because our `tensor` variable is of shape (2, 2).
#
# What if we created some tensors which had mismatched shapes?
# + colab={"base_uri": "https://localhost:8080/"} id="UXSE6q1o0amm" outputId="77694ca8-b6c5-4997-8e0e-9cb43bd78145"
# Create (3, 2) tensor
X = tf.constant([[1, 2],
[3, 4],
[5, 6]])
# Create another (3, 2) tensor
Y = tf.constant([[7, 8],
[9, 10],
[11, 12]])
X, Y
# + colab={"base_uri": "https://localhost:8080/", "height": 241} id="3J4DGQa309Hc" outputId="d7a867bd-6c5f-4ef9-b9a9-b00c89b1cba1"
# Try to matrix multiply them (will error)
X @ Y
# + [markdown] id="v27FQ59v1N-H"
# Trying to matrix multiply two tensors with the shape `(3, 2)` errors because the inner dimensions don't match.
#
# We need to either:
# * Reshape X to `(2, 3)` so it's `(2, 3) @ (3, 2)`.
# * Reshape Y to `(3, 2)` so it's `(3, 2) @ (2, 3)`.
#
# We can do this with either:
# * [`tf.reshape()`](https://www.tensorflow.org/api_docs/python/tf/reshape) - allows us to reshape a tensor into a defined shape.
# * [`tf.transpose()`](https://www.tensorflow.org/api_docs/python/tf/transpose) - switches the dimensions of a given tensor.
#
# 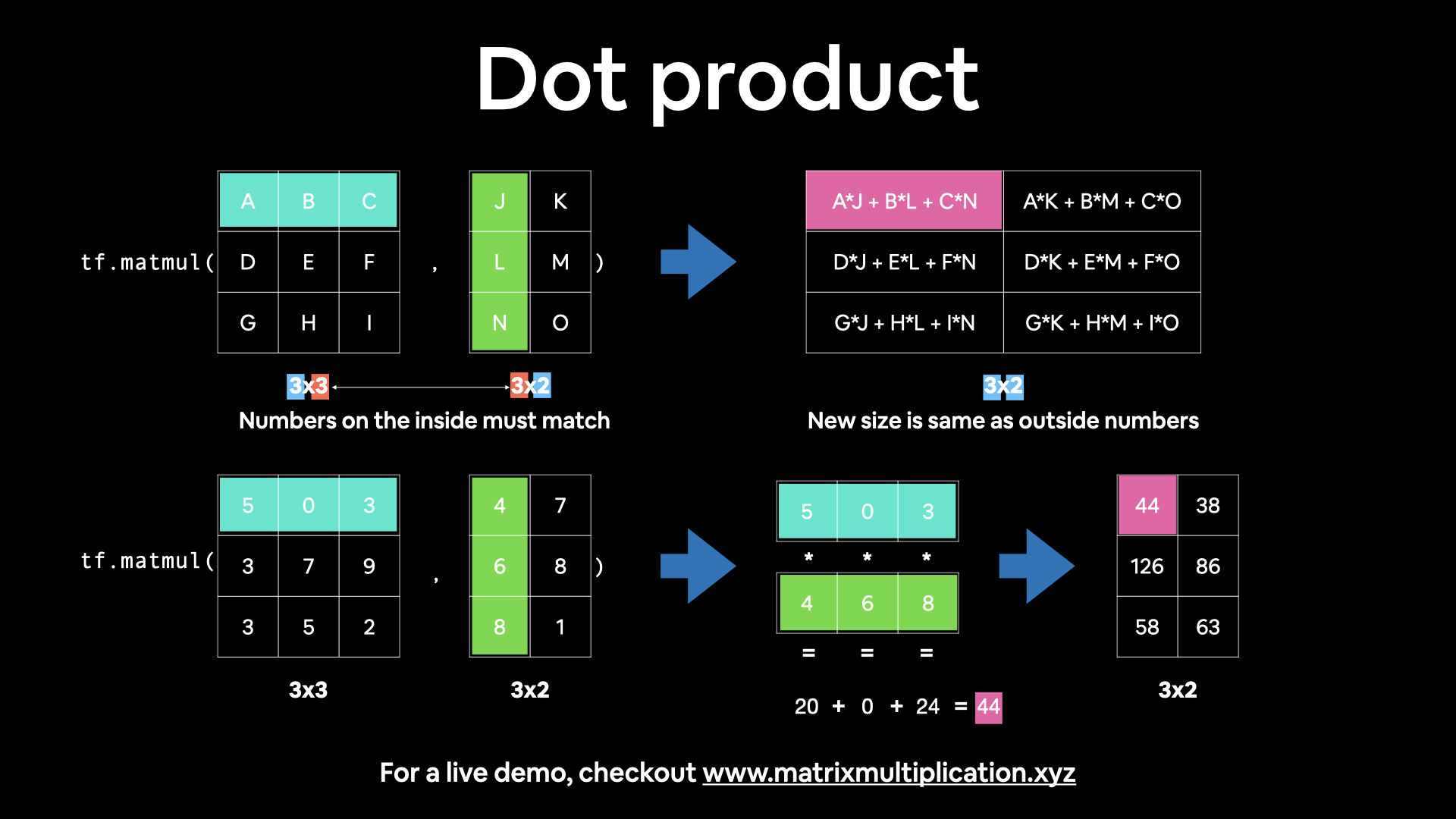
#
# Let's try `tf.reshape()` first.
# + colab={"base_uri": "https://localhost:8080/"} id="ZwvVl-k_2W9u" outputId="e408fd19-4705-476d-f1c5-302ff6859e54"
# Example of reshape (3, 2) -> (2, 3)
tf.reshape(Y, shape=(2, 3))
# + colab={"base_uri": "https://localhost:8080/"} id="1jBKPQVn1Nep" outputId="19906c7b-3e3b-4611-a639-7c8a3fff1d6a"
# Try matrix multiplication with reshaped Y
X @ tf.reshape(Y, shape=(2, 3))
# + [markdown] id="CWAgHKNk2_TO"
# It worked, let's try the same with a reshaped `X`, except this time we'll use [`tf.transpose()`](https://www.tensorflow.org/api_docs/python/tf/transpose) and `tf.matmul()`.
# + colab={"base_uri": "https://localhost:8080/"} id="qA2rCnik2OnQ" outputId="f23907d3-f052-4702-dcbb-371e0ccb8df3"
# Example of transpose (3, 2) -> (2, 3)
tf.transpose(X)
# + colab={"base_uri": "https://localhost:8080/"} id="zR8YdMfh3G0S" outputId="cd2fbe33-97c6-4e54-c27f-8febe63c8e4f"
# Try matrix multiplication
tf.matmul(tf.transpose(X), Y)
# + colab={"base_uri": "https://localhost:8080/"} id="SL45P1cC5tnJ" outputId="b2aa00a3-c1e2-44a5-a574-48a0a721f0ee"
# You can achieve the same result with parameters
tf.matmul(a=X, b=Y, transpose_a=True, transpose_b=False)
# + [markdown] id="AqE518TJ3GXG"
# Notice the difference in the resulting shapes when tranposing `X` or reshaping `Y`.
#
# This is because of the 2nd rule mentioned above:
# * `(3, 2) @ (2, 3)` -> `(3, 3)` done with `X @ tf.reshape(Y, shape=(2, 3))`
# * `(2, 3) @ (3, 2)` -> `(2, 2)` done with `tf.matmul(tf.transpose(X), Y)`
#
# This kind of data manipulation is a reminder: you'll spend a lot of your time in machine learning and working with neural networks reshaping data (in the form of tensors) to prepare it to be used with various operations (such as feeding it to a model).
#
# ### The dot product
#
# Multiplying matrices by eachother is also referred to as the dot product.
#
# You can perform the `tf.matmul()` operation using [`tf.tensordot()`](https://www.tensorflow.org/api_docs/python/tf/tensordot).
# + colab={"base_uri": "https://localhost:8080/"} id="qfSJHDpe2Oe9" outputId="67aab203-89c1-4b2f-afe5-b96416f043d9"
# Perform the dot product on X and Y (requires X to be transposed)
tf.tensordot(tf.transpose(X), Y, axes=1)
# + [markdown] id="waJcSOGf_Fg7"
# You might notice that although using both `reshape` and `tranpose` work, you get different results when using each.
#
# Let's see an example, first with `tf.transpose()` then with `tf.reshape()`.
# + colab={"base_uri": "https://localhost:8080/"} id="AAzB-F4l6Dc0" outputId="f2c1545c-e2e5-4e03-9319-65985cd299da"
# Perform matrix multiplication between X and Y (transposed)
tf.matmul(X, tf.transpose(Y))
# + colab={"base_uri": "https://localhost:8080/"} id="s-kQH7qh69PV" outputId="f8ff25a7-a7e6-4fbc-c1b7-91059ca8cfa7"
# Perform matrix multiplication between X and Y (reshaped)
tf.matmul(X, tf.reshape(Y, (2, 3)))
# + [markdown] id="2eCDnCX6AhbF"
# Hmm... they result in different values.
#
# Which is strange because when dealing with `Y` (a `(3x2)` matrix), reshaping to `(2, 3)` and tranposing it result in the same shape.
# + colab={"base_uri": "https://localhost:8080/"} id="P_RLV373ATAb" outputId="202e1c47-504a-4254-bcab-faac96ab9f86"
# Check shapes of Y, reshaped Y and tranposed Y
Y.shape, tf.reshape(Y, (2, 3)).shape, tf.transpose(Y).shape
# + [markdown] id="OilqUMBKAevX"
# But calling `tf.reshape()` and `tf.transpose()` on `Y` don't necessarily result in the same values.
# + colab={"base_uri": "https://localhost:8080/"} id="B5_aYjqeA_w_" outputId="39d8c934-c06e-4442-8a9d-fb91a7fb82e2"
# Check values of Y, reshape Y and tranposed Y
print("Normal Y:")
print(Y, "\n") # "\n" for newline
print("Y reshaped to (2, 3):")
print(tf.reshape(Y, (2, 3)), "\n")
print("Y transposed:")
print(tf.transpose(Y))
# + [markdown] id="g9g3l45LFO7K"
# As you can see, the outputs of `tf.reshape()` and `tf.transpose()` when called on `Y`, even though they have the same shape, are different.
#
# This can be explained by the default behaviour of each method:
# * [`tf.reshape()`](https://www.tensorflow.org/api_docs/python/tf/reshape) - change the shape of the given tensor (first) and then insert values in order they appear (in our case, 7, 8, 9, 10, 11, 12).
# * [`tf.transpose()`](https://www.tensorflow.org/api_docs/python/tf/transpose) - swap the order of the axes, by default the last axis becomes the first, however the order can be changed using the [`perm` parameter](https://www.tensorflow.org/api_docs/python/tf/transpose).
# + [markdown] id="fzjcZ4FHCOb5"
# So which should you use?
#
# Again, most of the time these operations (when they need to be run, such as during the training a neural network, will be implemented for you).
#
# But generally, whenever performing a matrix multiplication and the shapes of two matrices don't line up, you will transpose (not reshape) one of them in order to line them up.
#
# ### Matrix multiplication tidbits
# * If we transposed `Y`, it would be represented as $\mathbf{Y}^\mathsf{T}$ (note the capital T for tranpose).
# * Get an illustrative view of matrix multiplication [by Math is Fun](https://www.mathsisfun.com/algebra/matrix-multiplying.html).
# * Try a hands-on demo of matrix multiplcation: http://matrixmultiplication.xyz/ (shown below).
#
# 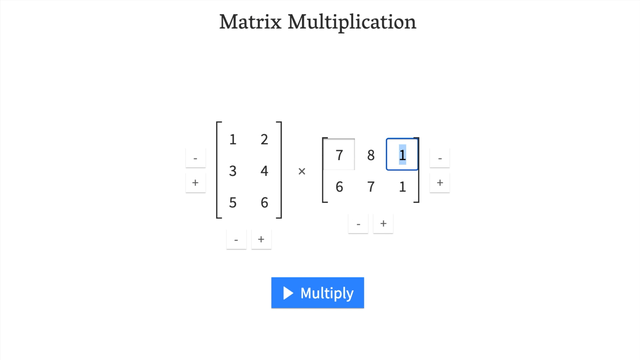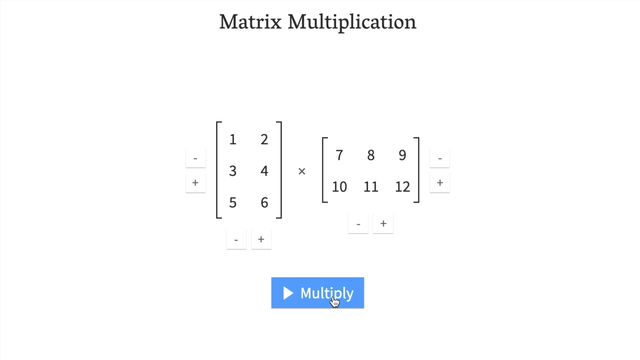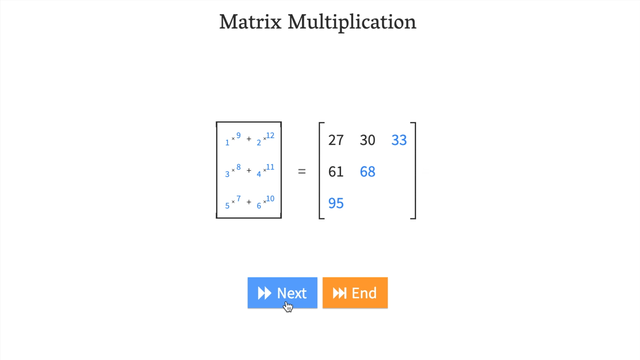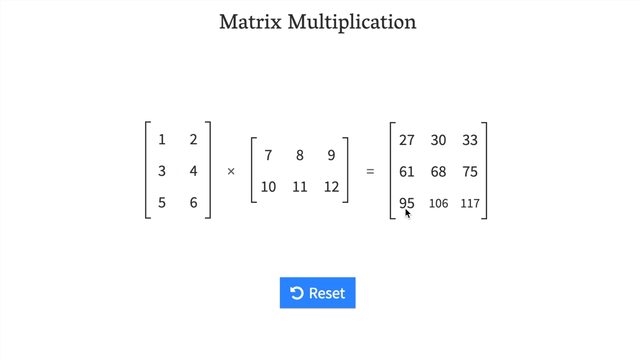
# + [markdown] id="bK8Kc94SL-JL"
# ### Changing the datatype of a tensor
#
# Sometimes you'll want to alter the default datatype of your tensor.
#
# This is common when you want to compute using less precision (e.g. 16-bit floating point numbers vs. 32-bit floating point numbers).
#
# Computing with less precision is useful on devices with less computing capacity such as mobile devices (because the less bits, the less space the computations require).
#
# You can change the datatype of a tensor using [`tf.cast()`](https://www.tensorflow.org/api_docs/python/tf/cast).
# + colab={"base_uri": "https://localhost:8080/"} id="jRkZhW35Lge_" outputId="a205ee88-9f2f-485c-e6e5-3be0b080fb6f"
# Create a new tensor with default datatype (float32)
B = tf.constant([1.7, 7.4])
# Create a new tensor with default datatype (int32)
C = tf.constant([1, 7])
B, C
# + colab={"base_uri": "https://localhost:8080/"} id="kdhV1rgcNpUP" outputId="83a85f1c-fdd3-44e9-f854-75c36b94af3b"
# Change from float32 to float16 (reduced precision)
B = tf.cast(B, dtype=tf.float16)
B
# + colab={"base_uri": "https://localhost:8080/"} id="Px2E1ANeNxRv" outputId="be744bf6-beca-4b27-e63d-ae1255be0593"
# Change from int32 to float32
C = tf.cast(C, dtype=tf.float32)
C
# + [markdown] id="1F3L1BDuQVJz"
# ### Getting the absolute value
# Sometimes you'll want the absolute values (all values are positive) of elements in your tensors.
#
# To do so, you can use [`tf.abs()`](https://www.tensorflow.org/api_docs/python/tf/math/abs).
# + colab={"base_uri": "https://localhost:8080/"} id="plNBFi51QOvW" outputId="79dbe7c8-e212-4b0d-9ee1-965c496afe28"
# Create tensor with negative values
D = tf.constant([-7, -10])
D
# + colab={"base_uri": "https://localhost:8080/"} id="bOSYmX37QFHS" outputId="8aa0a217-54e1-4edf-b812-4b0eb05f61a5"
# Get the absolute values
tf.abs(D)
# + [markdown] id="G4cnALehQ2CE"
# ### Finding the min, max, mean, sum (aggregation)
#
# You can quickly aggregate (perform a calculation on a whole tensor) tensors to find things like the minimum value, maximum value, mean and sum of all the elements.
#
# To do so, aggregation methods typically have the syntax `reduce()_[action]`, such as:
# * [`tf.reduce_min()`](https://www.tensorflow.org/api_docs/python/tf/math/reduce_min) - find the minimum value in a tensor.
# * [`tf.reduce_max()`](https://www.tensorflow.org/api_docs/python/tf/math/reduce_max) - find the maximum value in a tensor (helpful for when you want to find the highest prediction probability).
# * [`tf.reduce_mean()`](https://www.tensorflow.org/api_docs/python/tf/math/reduce_mean) - find the mean of all elements in a tensor.
# * [`tf.reduce_sum()`](https://www.tensorflow.org/api_docs/python/tf/math/reduce_sum) - find the sum of all elements in a tensor.
# * **Note:** typically, each of these is under the `math` module, e.g. `tf.math.reduce_min()` but you can use the alias `tf.reduce_min()`.
#
# Let's see them in action.
# + colab={"base_uri": "https://localhost:8080/"} id="kjW5_6i6Q7oo" outputId="7145a7b4-d9da-4120-a2cb-5ebe2bbc0f17"
# Create a tensor with 50 random values between 0 and 100
E = tf.constant(np.random.randint(low=0, high=100, size=50))
E
# + colab={"base_uri": "https://localhost:8080/"} id="slwMVgT-Rac0" outputId="34cf2c05-53b9-44eb-9df5-28df227794d4"
# Find the minimum
tf.reduce_min(E)
# + colab={"base_uri": "https://localhost:8080/"} id="voLqafGCRYqO" outputId="8bec8619-45e8-4b95-970b-4ee3b7c55d77"
# Find the maximum
tf.reduce_max(E)
# + colab={"base_uri": "https://localhost:8080/"} id="MPqwvy6TRYN3" outputId="905c4bf8-5d3f-443f-f92b-74752800987a"
# Find the mean
tf.reduce_mean(E)
# + colab={"base_uri": "https://localhost:8080/"} id="lqwCXeD6RhyE" outputId="4ce00693-124a-4875-a942-5a48672ace3c"
# Find the sum
tf.reduce_sum(E)
# + [markdown] id="zXGyiVPiRqgO"
# You can also find the standard deviation ([`tf.reduce_std()`](https://www.tensorflow.org/api_docs/python/tf/math/reduce_std)) and variance ([`tf.reduce_variance()`](https://www.tensorflow.org/api_docs/python/tf/math/reduce_variance)) of elements in a tensor using similar methods.
#
# ### Finding the positional maximum and minimum
#
# How about finding the position a tensor where the maximum value occurs?
#
# This is helpful when you want to line up your labels (say `['Green', 'Blue', 'Red']`) with your prediction probabilities tensor (e.g. `[0.98, 0.01, 0.01]`).
#
# In this case, the predicted label (the one with the highest prediction probability) would be `'Green'`.
#
# You can do the same for the minimum (if required) with the following:
# * [`tf.argmax()`](https://www.tensorflow.org/api_docs/python/tf/math/argmax) - find the position of the maximum element in a given tensor.
# * [`tf.argmin()`](https://www.tensorflow.org/api_docs/python/tf/math/argmin) - find the position of the minimum element in a given tensor.
# + colab={"base_uri": "https://localhost:8080/"} id="PspO0Vjp3Nm6" outputId="74c67bd9-a883-4b67-a7a3-55654d506e77"
# Create a tensor with 50 values between 0 and 1
F = tf.constant(np.random.random(50))
F
# + colab={"base_uri": "https://localhost:8080/"} id="ADbAMm9N3Zlb" outputId="6fc95755-3b77-495e-f382-0a8b7f2cdef2"
# Find the maximum element position of F
tf.argmax(F)
# + colab={"base_uri": "https://localhost:8080/"} id="aQrv1nVE3ckx" outputId="0a78d68f-6804-4125-da7c-d12a6811258e"
# Find the minimum element position of F
tf.argmin(F)
# + colab={"base_uri": "https://localhost:8080/"} id="yFHzARFwLmIf" outputId="2fbf3ccb-5ae2-4f20-c351-ed98401a135d"
# Find the maximum element position of F
print(f"The maximum value of F is at position: {tf.argmax(F).numpy()}")
print(f"The maximum value of F is: {tf.reduce_max(F).numpy()}")
print(f"Using tf.argmax() to index F, the maximum value of F is: {F[tf.argmax(F)].numpy()}")
print(f"Are the two max values the same (they should be)? {F[tf.argmax(F)].numpy() == tf.reduce_max(F).numpy()}")
# + [markdown] id="vSGIuNwm5QHM"
# ### Squeezing a tensor (removing all single dimensions)
#
# If you need to remove single-dimensions from a tensor (dimensions with size 1), you can use `tf.squeeze()`.
#
# * [`tf.squeeze()`](https://www.tensorflow.org/api_docs/python/tf/squeeze) - remove all dimensions of 1 from a tensor.
#
# + colab={"base_uri": "https://localhost:8080/"} id="6xDZLtNu5wUZ" outputId="b86ca328-b539-4701-e729-b7dc5c80dc49"
# Create a rank 5 (5 dimensions) tensor of 50 numbers between 0 and 100
G = tf.constant(np.random.randint(0, 100, 50), shape=(1, 1, 1, 1, 50))
G.shape, G.ndim
# + colab={"base_uri": "https://localhost:8080/"} id="oS91XOgO6lai" outputId="0359d710-ca66-40c1-ac48-cd60c164f5fe"
# Squeeze tensor G (remove all 1 dimensions)
G_squeezed = tf.squeeze(G)
G_squeezed.shape, G_squeezed.ndim
# + [markdown] id="46cKe32W65Ox"
# ### One-hot encoding
#
# If you have a tensor of indicies and would like to one-hot encode it, you can use [`tf.one_hot()`](https://www.tensorflow.org/api_docs/python/tf/one_hot).
#
# You should also specify the `depth` parameter (the level which you want to one-hot encode to).
# + colab={"base_uri": "https://localhost:8080/"} id="FlRkMjL-646U" outputId="f49a6bee-e62a-4175-b84c-e4f8b5500d4f"
# Create a list of indices
some_list = [0, 1, 2, 3]
# One hot encode them
tf.one_hot(some_list, depth=4)
# + [markdown] id="xYxYV0r08THk"
# You can also specify values for `on_value` and `off_value` instead of the default `0` and `1`.
# + colab={"base_uri": "https://localhost:8080/"} id="FZluadm88EcN" outputId="a129f06b-b23f-40d8-ed3a-ddbee35be070"
# Specify custom values for on and off encoding
tf.one_hot(some_list, depth=4, on_value="We're live!", off_value="Offline")
# + [markdown] id="-E-I1jFC84Qi"
# ### Squaring, log, square root
#
# Many other common mathematical operations you'd like to perform at some stage, probably exist.
#
# Let's take a look at:
# * [`tf.square()`](https://www.tensorflow.org/api_docs/python/tf/math/square) - get the square of every value in a tensor.
# * [`tf.sqrt()`](https://www.tensorflow.org/api_docs/python/tf/math/sqrt) - get the squareroot of every value in a tensor (**note:** the elements need to be floats or this will error).
# * [`tf.math.log()`](https://www.tensorflow.org/api_docs/python/tf/math/log) - get the natural log of every value in a tensor (elements need to floats).
# + colab={"base_uri": "https://localhost:8080/"} id="KTOvziCBLqhZ" outputId="9ef46ed1-6bd2-4e61-9068-11f5bb183150"
# Create a new tensor
H = tf.constant(np.arange(1, 10))
H
# + colab={"base_uri": "https://localhost:8080/"} id="qvPS3xbk9JBK" outputId="999851d2-5a61-4768-8a43-1b1862ffb0bd"
# Square it
tf.square(H)
# + colab={"base_uri": "https://localhost:8080/", "height": 275} id="SJibI0GO9uf4" outputId="0a50bff8-6654-470c-dad8-7177cdf29d7b"
# Find the squareroot (will error), needs to be non-integer
tf.sqrt(H)
# + colab={"base_uri": "https://localhost:8080/"} id="zlxRWy6Q-NHK" outputId="f61c7c31-34da-4c8f-8040-be761e939f95"
# Change H to float32
H = tf.cast(H, dtype=tf.float32)
H
# + colab={"base_uri": "https://localhost:8080/"} id="S73eO0p--TtN" outputId="6ae3c553-1df5-4f5a-a5a1-e3597300c583"
# Find the square root
tf.sqrt(H)
# + colab={"base_uri": "https://localhost:8080/"} id="RyCo55vz9u4f" outputId="3a8f9f11-4a00-482b-dde5-b5cebee8dcde"
# Find the log (input also needs to be float)
tf.math.log(H)
# + [markdown] id="urq3bx3l_Y4K"
# ### Manipulating `tf.Variable` tensors
#
# Tensors created with `tf.Variable()` can be changed in place using methods such as:
#
# * [`.assign()`](https://www.tensorflow.org/api_docs/python/tf/Variable#assign) - assign a different value to a particular index of a variable tensor.
# * [`.add_assign()`](https://www.tensorflow.org/api_docs/python/tf/Variable#assign_add) - add to an existing value and reassign it at a particular index of a variable tensor.
#
# + colab={"base_uri": "https://localhost:8080/"} id="tV7_uzdR_F4c" outputId="dbe2bd14-249b-4bf0-c2f1-1af02d98c59a"
# Create a variable tensor
I = tf.Variable(np.arange(0, 5))
I
# + colab={"base_uri": "https://localhost:8080/"} id="ukatOuCG_4CY" outputId="a6351712-d894-4e23-c57e-2ab45537d566"
# Assign the final value a new value of 50
I.assign([0, 1, 2, 3, 50])
# + colab={"base_uri": "https://localhost:8080/"} id="3r2Pya_vAnLz" outputId="a65b7db9-a0ec-47d7-f3cf-12cb60eb31ac"
# The change happens in place (the last value is now 50, not 4)
I
# + colab={"base_uri": "https://localhost:8080/"} id="gZie_FduAo5O" outputId="0b7d34bf-76e9-429e-ecec-7bb2beedb750"
# Add 10 to every element in I
I.assign_add([10, 10, 10, 10, 10])
# + colab={"base_uri": "https://localhost:8080/"} id="2qXODgVOBDzC" outputId="5fd97dbb-eb43-4e55-b74a-f4e94956b4c2"
# Again, the change happens in place
I
# + [markdown] id="2L2IyQhl-Z73"
# ## Tensors and NumPy
#
# We've seen some examples of tensors interact with NumPy arrays, such as, using NumPy arrays to create tensors.
#
# Tensors can also be converted to NumPy arrays using:
#
# * `np.array()` - pass a tensor to convert to an ndarray (NumPy's main datatype).
# * `tensor.numpy()` - call on a tensor to convert to an ndarray.
#
# Doing this is helpful as it makes tensors iterable as well as allows us to use any of NumPy's methods on them.
# + colab={"base_uri": "https://localhost:8080/"} id="HLHrij0vBywD" outputId="83109e2d-e3be-4e95-e6dd-d84713e9dc70"
# Create a tensor from a NumPy array
J = tf.constant(np.array([3., 7., 10.]))
J
# + colab={"base_uri": "https://localhost:8080/"} id="P0KBe_FqCKdU" outputId="c4de5d97-8095-4e0c-86de-67fad637d88d"
# Convert tensor J to NumPy with np.array()
np.array(J), type(np.array(J))
# + colab={"base_uri": "https://localhost:8080/"} id="xxKsJPvSCUPI" outputId="39cef6d1-2deb-4ec2-cbdf-b9e6f3b06488"
# Convert tensor J to NumPy with .numpy()
J.numpy(), type(J.numpy())
# + [markdown] id="SL4xHtNWDIT5"
# By default tensors have `dtype=float32`, where as NumPy arrays have `dtype=float64`.
#
# This is because neural networks (which are usually built with TensorFlow) can generally work very well with less precision (32-bit rather than 64-bit).
# + colab={"base_uri": "https://localhost:8080/"} id="HrQoUeyPCXU9" outputId="d4790fab-fc39-41d8-bf6b-c628d0f1ada0"
# Create a tensor from NumPy and from an array
numpy_J = tf.constant(np.array([3., 7., 10.])) # will be float64 (due to NumPy)
tensor_J = tf.constant([3., 7., 10.]) # will be float32 (due to being TensorFlow default)
numpy_J.dtype, tensor_J.dtype
# + [markdown] id="_ytLPBWF1wxt"
# ## Using `@tf.function`
#
# In your TensorFlow adventures, you might come across Python functions which have the decorator [`@tf.function`](https://www.tensorflow.org/api_docs/python/tf/function).
#
# If you aren't sure what Python decorators do, [read RealPython's guide on them](https://realpython.com/primer-on-python-decorators/).
#
# But in short, decorators modify a function in one way or another.
#
# In the `@tf.function` decorator case, it turns a Python function into a callable TensorFlow graph. Which is a fancy way of saying, if you've written your own Python function, and you decorate it with `@tf.function`, when you export your code (to potentially run on another device), TensorFlow will attempt to convert it into a fast(er) version of itself (by making it part of a computation graph).
#
# For more on this, read the [Better performnace with tf.function](https://www.tensorflow.org/guide/function) guide.
# + colab={"base_uri": "https://localhost:8080/"} id="URzFiwPoFDI7" outputId="a6c7bbe3-4bfb-499a-91e9-f8bd4313b6a0"
# Create a simple function
def function(x, y):
return x ** 2 + y
x = tf.constant(np.arange(0, 10))
y = tf.constant(np.arange(10, 20))
function(x, y)
# + colab={"base_uri": "https://localhost:8080/"} id="EHGbvlkXF7Gs" outputId="40f5d83c-f2fc-4c50-fdfe-f37b3fdfa75b"
# Create the same function and decorate it with tf.function
@tf.function
def tf_function(x, y):
return x ** 2 + y
tf_function(x, y)
# + [markdown] id="lsiapEnTGBbH"
# If you noticed no difference between the above two functions (the decorated one and the non-decorated one) you'd be right.
#
# Much of the difference happens behind the scenes. One of the main ones being potential code speed-ups where possible.
# + [markdown] id="y8AQmchkxKDn"
# ## Finding access to GPUs
#
# We've mentioned GPUs plenty of times throughout this notebook.
#
# So how do you check if you've got one available?
#
# You can check if you've got access to a GPU using [`tf.config.list_physical_devices()`](https://www.tensorflow.org/guide/gpu).
# + colab={"base_uri": "https://localhost:8080/"} id="3rFNSeODxi3L" outputId="7dd6e024-bed3-46ad-d96f-bea634317e24"
print(tf.config.list_physical_devices('GPU'))
# + [markdown] id="8165PtH8xyq5"
# If the above outputs an empty array (or nothing), it means you don't have access to a GPU (or at least TensorFlow can't find it).
#
# If you're running in Google Colab, you can access a GPU by going to *Runtime -> Change Runtime Type -> Select GPU* (**note:** after doing this your notebook will restart and any variables you've saved will be lost).
#
# Once you've changed your runtime type, run the cell below.
# + colab={"base_uri": "https://localhost:8080/"} id="iAvMwRbZyULE" outputId="eaab2fea-5960-43f1-899f-1eee870a76f9"
import tensorflow as tf
print(tf.config.list_physical_devices('GPU'))
# + [markdown] id="J3yu0LKiyZlJ"
# If you've got access to a GPU, the cell above should output something like:
#
# `[PhysicalDevice(name='/physical_device:GPU:0', device_type='GPU')]`
#
# You can also find information about your GPU using `!nvidia-smi`.
# + colab={"base_uri": "https://localhost:8080/"} id="yOuVvG5kyefS" outputId="fc734522-6270-447d-cefc-34e9fbe4a89b"
# !nvidia-smi
# + [markdown] id="pER7mLFezV9u"
# > 🔑 **Note:** If you have access to a GPU, TensorFlow will automatically use it whenever possible.
# + [markdown] id="HcCZxK61VIv0"
# ## 🛠 Exercises
#
# 1. Create a vector, scalar, matrix and tensor with values of your choosing using `tf.constant()`.
# 2. Find the shape, rank and size of the tensors you created in 1.
# 3. Create two tensors containing random values between 0 and 1 with shape `[5, 300]`.
# 4. Multiply the two tensors you created in 3 using matrix multiplication.
# 5. Multiply the two tensors you created in 3 using dot product.
# 6. Create a tensor with random values between 0 and 1 with shape `[224, 224, 3]`.
# 7. Find the min and max values of the tensor you created in 6.
# 8. Created a tensor with random values of shape `[1, 224, 224, 3]` then squeeze it to change the shape to `[224, 224, 3]`.
# 9. Create a tensor with shape `[10]` using your own choice of values, then find the index which has the maximum value.
# 10. One-hot encode the tensor you created in 9.
# + [markdown] id="EgHK5CjY3oVX"
# ## 📖 Extra-curriculum
#
# * Read through the [list of TensorFlow Python APIs](https://www.tensorflow.org/api_docs/python/), pick one we haven't gone through in this notebook, reverse engineer it (write out the documentation code for yourself) and figure out what it does.
# * Try to create a series of tensor functions to calculate your most recent grocery bill (it's okay if you don't use the names of the items, just the price in numerical form).
# * How would you calculate your grocery bill for the month and for the year using tensors?
# * Go through the [TensorFlow 2.x quick start for beginners](https://www.tensorflow.org/tutorials/quickstart/beginner) tutorial (be sure to type out all of the code yourself, even if you don't understand it).
# * Are there any functions we used in here that match what's used in there? Which are the same? Which haven't you seen before?
# * Watch the video ["What's a tensor?"](https://www.youtube.com/watch?v=f5liqUk0ZTw) - a great visual introduction to many of the concepts we've covered in this notebook.
| 1_tensorflow_fundamentals.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/lavanyashukla/CycleGAN/blob/master/SpaceInvaders_Baseline_Model_(3).ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="KfOuv88IHJTV"
# # Space Invaders
#
#
# + colab_type="code" id="o-uAdQfWkjIh" outputId="1972d90b-84bc-48ca-f929-88d21f61f5a5" colab={"base_uri": "https://localhost:8080/", "height": 51}
# !pip install wandb -qq
# !pip install gym pyvirtualdisplay -qq
# !apt-get install -y xvfb python-opengl ffmpeg -qq
# !pip install xdpyinfo -qq
# !apt-get update -qq
# !apt-get install cmake -qq
# !pip install --upgrade setuptools -qq
# !pip install ez_setup -qq
# + colab_type="code" id="ms0B7dnLHJTc" outputId="d28a30ed-eaed-4b42-edda-99ea89f5cfd1" colab={"base_uri": "https://localhost:8080/", "height": 63}
import gym
from gym import logger as gymlogger
from gym.wrappers import Monitor
gymlogger.set_level(30)
import numpy as np
import random
import math
import glob
import io
import base64
import tensorflow as tf
import matplotlib
import matplotlib.pyplot as plt
# %matplotlib inline
from datetime import datetime
from IPython.display import HTML
from IPython import display as ipythondisplay
from pyvirtualdisplay import Display
# import wandb
import wandb
# + [markdown] colab_type="text" id="wXDlF6cdHJTj"
# ## Preprocessing - crop images, convert them to 1D black and white image tensors
#
# - Image dimensions - (210, 160, 3)
# - Output dimensions - (88, 80, 1)
#
# + colab_type="code" id="29CSBYkPHJTl" colab={}
color = np.array([210, 164, 74]).mean()
def preprocess_frame(obs):
# Crop and resize
img = obs[25:201:2, ::2]
# Convert to greyscale
img = img.mean(axis=2)
# Improve contrast
img[img==color] = 0
# Normalzie image
img = (img - 128) / 128 - 1
# Reshape to 80*80*1
img = img.reshape(88,80)
return img
# + [markdown] colab_type="text" id="Y1qZntlnHJTs"
# ## Initialize gym environment and explore game screens
#
# + id="S4nYnj42iYY1" colab_type="code" outputId="0c23b727-0f07-4e0f-d26c-927b679aa94e" colab={"base_uri": "https://localhost:8080/", "height": 34}
env = gym.make("SpaceInvaders-v0")
print("Actions available(%d): %r"%(env.action_space.n, env.env.get_action_meanings()))
# + colab_type="code" id="jnbrjWstHJTu" outputId="1f4c425b-a482-4a89-a62a-67258767bae2" colab={"base_uri": "https://localhost:8080/", "height": 269}
observation = env.reset()
# Game Screen
for i in range(11):
if i > 9:
plt.imshow(observation)
plt.show()
observation, _, _, _ = env.step(1)
# + colab_type="code" id="_Ma31y4wFpst" outputId="427115b1-c72c-4f40-e453-1cd863783402" colab={"base_uri": "https://localhost:8080/", "height": 268}
# Preprocessed Game Screen
obs_preprocessed = preprocess_frame(observation).reshape(88,80)
plt.imshow(obs_preprocessed)
plt.show()
# + [markdown] id="2tXWpRTdirOQ" colab_type="text"
# ## Play Game
# + [markdown] id="wjcpwUfqjBrE" colab_type="text"
# ### Play a random game, log reward and gameplay video in wandb
# + id="I9Hwt7z_jB14" colab_type="code" colab={}
# initialize a new wandb run
wandb.init(project="qualcomm")
# define hyperparameters
wandb.config.episodes = 100
wandb.config.batch_size = 48
wandb.config.learning_rate = 0.001
input_shape = (None, 88, 80, 1)
# record gameplay video
display = Display(visible=0, size=(1400, 900))
display.start()
# reward across episodes
cumulative_reward = 0
# run for 100 episodes
for i in range(wandb.config.episodes):
# Set reward received in this episode = 0 at the start of the episode
episodic_reward = 0
# record a video of the game using wrapper
env = gym.wrappers.Monitor(env, './video', force=True)
episode = i+1
print("Episode: %d"%(episode))
# play a random game
state = env.reset()
done = False
while not done:
# take random action
# ****TODO: replace this with model prediction****
action = env.action_space.sample()
# perform the action and fetch next state, reward
state, reward, done, _ = env.step(action)
episodic_reward += reward
# log total reward received in this episode to wandb
wandb.log({'episodic_reward': episodic_reward})
# add reward from this episode to cumulative_reward
cumulative_reward += episodic_reward
# calculate the cumulative_avg_reward
# this is the metric your models will be evaluated on
cumulative_avg_reward = cumulative_reward/episode
# log cumulative_avg_reward over episodes to wandb
wandb.log({'cumulative_avg_reward': cumulative_avg_reward})
# your models will be evaluated on 100-episode average reward
# therefore, we stop logging after 100 episodes
if (episode > 100):
break
record_video = False
env.close()
# render gameplay video
mp4list = glob.glob('video/*.mp4')
if len(mp4list) > 0:
print(len(mp4list))
mp4 = mp4list[-1]
video = io.open(mp4, 'r+b').read()
encoded = base64.b64encode(video)
# log gameplay video in wandb
wandb.log({"gameplays": wandb.Video(mp4, fps=4, format="gif")})
# display gameplay video
ipythondisplay.display(HTML(data='''<video alt="test" autoplay
loop controls style="height: 400px;">
<source src="data:video/mp4;base64,{0}" type="video/mp4" />
</video>'''.format(encoded.decode('ascii'))))
# + colab_type="code" id="OKk22m-0CbEs" colab={}
| SpaceInvaders_Baseline_Model_(3).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # How to use this Framework
#
# ## Data
# - Store your raw, processed and interim data here
# - Make sure that you dont commit data to github. It cannot hold large datasets
# - Even if you get your data from weblinks, store your processed dataset
#
#
# ## Docs
# This holds the Sphinx documentation system for your project. You can add all the documentation that you need for the project.
# Make sure you keep filling in the documentations you go along. Sphinx also allows for storing docstrings as documentation
# **--more on sphinx here--**
# ## Reports
# This is where you will store the documentation for each of the Stress tests. You will need to ensure that you keep track of everything that you submitted for the documentation here
#
# ## Notebooks
# This is where you will store your jupyter notebooks. All the analysis that you will be doing will be in jupyter notebooks
#
# # Scripts
# There may be times where you may want to run individual python scripts to run checks. You can store those here
#
# ## src
#
# This folder contains the module src. This is a custom module that you will be adding to ask you finish doing your analysis in your notebooks. As you finish your analysis, you will package your code as functions then move them here so you can write a clean version for you to show.
# ## Models
# Store your model pickle files here
| notebooks/How_to_use_this_framework.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/kingpaglinawan/OOP-1-1/blob/main/Prelim_Exam.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="UBUO627o8qdV" outputId="29d8baf0-dea0-45d6-84bf-db6acc85ab93"
n = 20
total_numbers = n
sum = 0
while n >= 0:
sum += n
n -= 1
print("sum =", sum)
average = sum / total_numbers
print("Average = ", average)
# + [markdown] id="AzSSsAvpC5lr"
# Problem 2. (50 points)
#
# 1. Write a Python to display your full name, student number, age, and course
# 2. Create a class named Student with attributes: Name, Student_No, Age, School, and Course
# 3. Create an object name Myself and assign an instance for each attribute.
# 4. Create a method Info() using an instantiation of a class.
# 5. Insert your GitHub link "Prelim Exam" from your repository named "OOP 1-1"
#
# + colab={"base_uri": "https://localhost:8080/"} id="tH4v2rKCDIxT" outputId="3e9fe707-813e-447a-ec76-58dc690d9cbc"
class OOP_1_1:
def __init__(self,fullname,student_no,age,course,school):
self.fullname = fullname
self.student_no = student_no
self.age = age
self.course = course
self.school = school
def info(self):
#print(self.fullname,self.student_no,self.age,self.course,self.school)
print("Name: ", self.fullname)
print("Student No. ", self.student_no)
print("Age: ", self.age,"years old")
print("School: ", self.school)
print("Course: ", self.course)
Myself = OOP_1_1("<NAME>",202102061,19,"BSCPE/ BACHELOR OF SCIENCE IN COMPUTER ENGINEERING","Cavite State University (Main Campus)")
Myself.info()
| Prelim_Exam.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/KordingLab/ENGR344/blob/master/tutorials/W4D1_How_do_we_know_how_certain_we_should_be/TA/W4D1_Tutorial1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="wuSb692OG7s9"
# # Tutorial 1: Linear regression with MLE
# **Module 4: How do we know how certain we should be?**
#
# **Originally By Neuromatch Academy**
#
# **Content creators**: <NAME>, <NAME>, <NAME> with help from <NAME>
#
# **Content reviewers**: <NAME>, <NAME>, <NAME>, <NAME>, <NAME>
#
# **Content Modifiers**: <NAME>, <NAME>
# + [markdown] id="yUZ5o1SRG7tA"
# **Our 2021 Sponsors, including Presenting Sponsor Facebook Reality Labs**
#
# <p align='center'><img src='https://github.com/NeuromatchAcademy/widgets/blob/master/sponsors.png?raw=True'/></p>
# + colab={"base_uri": "https://localhost:8080/", "height": 563, "referenced_widgets": ["1cffd4ccdae54039b76ead26e65cbf21", "f7141cebf2f048f982eededdc3537861", "d73b81d96eae480b9ed9d2dfee7a60b4", "1514270a166d45299bc217f33faf7167"]} cellView="form" id="slvOzp2YOqfZ" outputId="adc22778-4d22-4afd-de13-0ded5fb79553"
# @title Due Dates Calendar
from ipywidgets import widgets
from IPython.display import display, IFrame, YouTubeVideo
out1 = widgets.Output()
with out1:
calendar = IFrame(src="https://calendar.google.com/calendar/embed?src=356b9d2nspjttvgbb3tvgk2f58%40group.calendar.google.com&ctz=America%2FNew_York", width=600, height=480)
display(calendar)
out = widgets.Tab([out1])
out.set_title(0, 'Calendar')
display(out)
# + [markdown] id="v9CTWLTsG7tB"
# ---
# # Tutorial Objectives
#
# *Estimated timing of tutorial: 30 minutes*
#
# This is Tutorial 2 of a series on fitting models to data. We start with simple linear regression, using least squares optimization (Tutorial 1) and Maximum Likelihood Estimation (Tutorial 2). We will use bootstrapping to build confidence intervals around the inferred linear model parameters (Tutorial 3). We'll finish our exploration of regression models by generalizing to multiple linear regression and polynomial regression (Tutorial 4). We end by learning how to choose between these various models. We discuss the bias-variance trade-off (Tutorial 5) and Cross Validation for model selection (Tutorial 6).
#
# In this tutorial, we will use a different approach to fit linear models that incorporates the random 'noise' in our data.
# - Learn about probability distributions and probabilistic models
# - Learn how to calculate the likelihood of our model parameters
# - Learn how to implement the maximum likelihood estimator, to find the model parameter with the maximum likelihood
#
#
# + cellView="form" id="zkAji4BkG7tC"
# @title Tutorial slides
# @markdown These are the slides for the videos in all tutorials today
from IPython.display import IFrame
IFrame(src=f"https://mfr.ca-1.osf.io/render?url=https://osf.io/2mkq4/?direct%26mode=render%26action=download%26mode=render", width=854, height=480)
# + [markdown] id="Vqg-pjSjG7tD"
# ---
# # Setup
# + id="fg-ZjpWuG7tE"
import numpy as np
import matplotlib.pyplot as plt
from scipy import stats
# + cellView="form" id="xUyQaEiUG7tE"
#@title Figure Settings
import ipywidgets as widgets # interactive display
# %config InlineBackend.figure_format = 'retina'
plt.style.use("https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/nma.mplstyle")
# + cellView="form" id="s-_f4Ok5G7tF"
# @title Plotting Functions
def plot_density_image(x, y, theta, sigma=1, ax=None):
""" Plots probability distribution of y given x, theta, and sigma
Args:
x (ndarray): An array of shape (samples,) that contains the input values.
y (ndarray): An array of shape (samples,) that contains the corresponding
measurement values to the inputs.
theta (float): Slope parameter
sigma (float): standard deviation of Gaussian noise
"""
# plot the probability density of p(y|x,theta)
if ax is None:
fig, ax = plt.subplots()
xmin, xmax = np.floor(np.min(x)), np.ceil(np.max(x))
ymin, ymax = np.floor(np.min(y)), np.ceil(np.max(y))
xx = np.linspace(xmin, xmax, 50)
yy = np.linspace(ymin, ymax, 50)
surface = np.zeros((len(yy), len(xx)))
for i, x_i in enumerate(xx):
surface[:, i] = stats.norm(theta * x_i, sigma).pdf(yy)
ax.set(xlabel='x', ylabel='y')
return ax.imshow(surface, origin='lower', aspect='auto', vmin=0, vmax=None,
cmap=plt.get_cmap('Wistia'),
extent=[xmin, xmax, ymin, ymax])
# + [markdown] id="rPT0sztBG7tG"
# ---
# # Section 1: Maximum Likelihood Estimation (MLE)
#
# + id="oJaU9BY-G7tG" colab={"base_uri": "https://localhost:8080/", "height": 581, "referenced_widgets": ["5a6bf15e553a472789d07bf400e19640", "f8fec86b1cc64c7d9a7ea4023d6c0bb9", "6b70f8f0505648d2af4ae1d4f67d6a14", "4af7ed809bcb4dbc8c4e48cb16663c63"]} outputId="705bbd53-b86b-45a8-8460-1a72efdcdfcc" cellView="form"
# @title Video 1: Maximum Likelihood Estimation
from ipywidgets import widgets
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="liTs_2PhiTU", width=854, height=480, fs=1, rel=0)
print('Video available at https://youtube.com/watch?v=' + video.id)
display(video)
out = widgets.Tab([out1])
out.set_title(0, 'Youtube')
display(out)
# + [markdown] id="4IkFiRTHG7tH"
# This video covers Maximum Likelihood Estimation (MLE) in the context of a 1D linear regression.
#
# + [markdown] id="386aghBnG7tH"
# ## Section 1.1: Gaussian noise
# + [markdown] id="Fjc70ZDzG7tH"
#
# <details>
# <summary> <font color='blue'>Click here for text recap of relevant part of video </font></summary>
#
# In the previous tutorial we made the assumption that the data was drawn from a linear relationship with noise added, and found an effective approach for estimating model parameters based on minimizing the mean squared error.
#
# In that case we treated the noise as simply a nuisance, but what if we factored it directly into our model?
#
# Recall our linear model:
#
# \begin{align}
# y = \theta x + \epsilon.
# \end{align}
#
# The noise component $\epsilon$ is often modeled as a random variable drawn from a Gaussian distribution (also called the normal distribution).
#
# The Gaussian distribution is described by its [probability density function](https://en.wikipedia.org/wiki/Probability_density_function) (pdf)
# \begin{align}
# \mathcal{N}(x; \mu, \sigma^2) = \frac{1}{\sqrt{2\pi\sigma^2}}e^{-\frac{1}{2\sigma^2}(x-\mu)^2}
# \end{align}
#
# and is dependent on two parameters: the mean $\mu$ and the variance $\sigma^2$. We often consider the noise signal to be Gaussian "white noise", with zero mean and unit variance:
#
# \begin{align}
# \epsilon \sim \mathcal{N}(0, 1).
# \end{align}
#
#
# + [markdown] id="bFUmLfIUG7tH"
# ### Interactive Demo 1.1: Gaussian Distribution Explorer
#
# Use the explorer widget below to see how varying the $\mu$ and $\sigma$ parameters change the location and shape of the samples.
#
#
# 1. What effect does varying $\mu$ have on the pdf?
# 2. What effect does varying $\sigma$ have on the pdf?
#
#
# + cellView="form" id="VDBqKEfJG7tI"
# @markdown Make sure you execute this cell to enable the widget!
@widgets.interact(mu=widgets.FloatSlider(0.0, min=-2.0, max=2.0),
sigma=widgets.FloatSlider(1.0, min=0.5, max=2.0))
def plot_normal_dist(mu=0, sigma=1):
# Generate pdf & samples from normal distribution with mu/sigma
rv = stats.norm(mu, sigma)
x = np.linspace(-5, 5, 100)
y = rv.pdf(x)
samples = rv.rvs(1000)
# Plot
fig, ax = plt.subplots()
ax.hist(samples, 20, density=True, color='g', histtype='stepfilled', alpha=0.8,
label='histogram')
ax.plot(x, y, color='orange', linewidth=3, label='pdf')
ax.vlines(mu, 0, rv.pdf(mu), color='y', linewidth=3, label='$\mu$')
ax.vlines([mu-sigma, mu+sigma], 0, rv.pdf([mu-sigma, mu+sigma]), colors='red',
color='b', linewidth=3, label='$\sigma$')
ax.set(xlabel='x', ylabel='probability density', xlim=[-5, 5], ylim=[0, 1.0])
ax.legend()
# + id="JjtuIgDIG7tI"
# to_remove explanation
"""
1. Mu controls the mean of the Gaussian distribution - it shifts it along the x-axis.
2. Sigma controls the width of the Gaussian distribution - small sigma results in a narrow
distribution, large sigma results in a very wide distribution
"""
# + [markdown] id="AgZLeLWkG7tI"
# ## Section 1.2: Probabilistic Models
#
# *Estimated timing to here from start of tutorial: 11 min*
#
# Now that we have a model of our noise component $\epsilon$ as random variable, how do we incorporate this back into our original linear model from before? Consider again our simplified model $y = \theta x + \epsilon$ where the noise has zero mean and unit variance $\epsilon \sim \mathcal{N}(0, 1)$. We can now also treat $y$ as a random variable drawn from a Gaussian distribution where $\mu = \theta x$ and $\sigma^2 = 1$:
#
# \begin{align}
# y \sim \mathcal{N}(\theta x, 1)
# \end{align}
#
# which is to say that the probability of observing $y$ given $x$ and parameter $\theta$ is
# \begin{align}
# p(y|x,\theta) = \frac{1}{\sqrt{2\pi}}e^{-\frac{1}{2}(y-\theta x)^2}
# \end{align}
#
#
# Note that in this and the following sections, we will focus on a single data point (a single pairing of $x$ and $y$). We have dropped the subscript $i$ just for simplicity (that is, we use $x$ for a single data point, instead of $x_i$).
#
# Now, let's revisit our original sample dataset where the true underlying model has $\theta = 1.2$.
# + cellView="form" id="PMA6b0F-G7tJ"
# @markdown Execute this cell to generate some simulated data
# setting a fixed seed to our random number generator ensures we will always
# get the same psuedorandom number sequence
np.random.seed(121)
theta = 1.2
n_samples = 30
x = 10 * np.random.rand(n_samples) # sample from a uniform distribution over [0,10)
noise = np.random.randn(n_samples) # sample from a standard normal distribution
y = theta * x + noise
# + [markdown] id="6TSqpPdDG7tJ"
# This time we can plot the density of $p(y|x,\theta=1.2)$ and see how $p(y)$ changes for different values of $x$.
# + cellView="form" id="AmnLdK4aG7tJ"
# @markdown Execute this cell to visualize p(y|x, theta=1.2)
fig, (ax1, ax2) = plt.subplots(ncols=2, figsize=(10, 4))
# Invokes helper function to generate density image plots from data and parameters
im = plot_density_image(x, y, 1.2, ax=ax1)
plt.colorbar(im, ax=ax1)
ax1.axvline(8, color='k')
ax1.set(title=r'p(y | x, $\theta$=1.2)')
# Plot pdf for given x
ylim = ax1.get_ylim()
yy = np.linspace(ylim[0], ylim[1], 50)
ax2.plot(yy, stats.norm(theta * 8, 1).pdf(yy), color='orange', linewidth=2)
ax2.set(
title=r'p(y|x=8, $\theta$=1.2)',
xlabel='y',
ylabel='probability density');
# + [markdown] id="GfPdKbY_G7tJ"
# ## Section 1.3: Likelihood Estimation
#
# *Estimated timing to here from start of tutorial: 15 min*
#
# Now that we have our probabilistic model, we turn back to our original challenge of finding a good estimate for $\theta$ that fits our data. Given the inherent uncertainty when dealing in probabilities, we talk about the [likelihood](https://en.wikipedia.org/wiki/Likelihood_function) that some estimate $\hat{\theta}$ fits our data. The likelihood function $\mathcal{L}(\theta)$ is equal to the probability density function parameterized by that $\theta$:
#
# \begin{align}
# \mathcal{L}(\theta|x,y) = p(y|x,\theta) = \frac{1}{\sqrt{2\pi\sigma^2}}e^{-\frac{1}{2\sigma^2}(y-\theta x)^2}
# \end{align}
# + [markdown] id="mzuNXo2GG7tJ"
# ### Coding Exercise 1.3: Likelihood Function
#
# In this exercise you will implement the likelihood function $\mathcal{L}(\theta|x, y)$ for our linear model where $\sigma = 1$.
#
# After implementing this function, we can produce probabilities that our estimate $\hat{\theta}$ generated the provided observations. We will try with one of the samples from our dataset.
#
# TIP: Use `np.exp` and `np.sqrt` for the exponential and square root functions, respectively.
# + id="-OG0S10EG7tJ"
def likelihood(theta_hat, x, y):
"""The likelihood function for a linear model with noise sampled from a
Gaussian distribution with zero mean and unit variance.
Args:
theta_hat (float): An estimate of the slope parameter.
x (ndarray): An array of shape (samples,) that contains the input values.
y (ndarray): An array of shape (samples,) that contains the corresponding
measurement values to the inputs.
Returns:
ndarray: the likelihood values for the theta_hat estimate
"""
sigma = 1
##############################################################################
## TODO for students: implement the likelihood function
# Fill out function and remove
raise NotImplementedError("Student exercise: implement the likelihood function")
##############################################################################
# Compute Gaussian likelihood
pdf = ...
return pdf
print(likelihood(1.0, x[1], y[1]))
# + cellView="both" id="WYKmqe7sG7tK"
# to_remove solution
def likelihood(theta_hat, x, y):
"""The likelihood function for a linear model with noise sampled from a
Gaussian distribution with zero mean and unit variance.
Args:
theta_hat (float): An estimate of the slope parameter.
x (ndarray): An array of shape (samples,) that contains the input values.
y (ndarray): An array of shape (samples,) that contains the corresponding
measurement values to the inputs.
Returns:
float: the likelihood value for the theta_hat estimate
"""
sigma = 1
# Compute Gaussian likelihood
pdf = 1 / np.sqrt(2 * np.pi * sigma**2) * np.exp(-(y - theta_hat * x)**2 / (2 * sigma**2))
return pdf
print(likelihood(1.0, x[1], y[1]))
# + [markdown] id="qZrmQ1PsG7tK"
# We should see that $\mathcal{L}(\theta=1.0|x=2.1,y=3.7) \approx 0.11$. So far so good, but how does this tell us how this estimate is better than any others?
#
# When dealing with a set of data points, as we are with our dataset, we are concerned with their joint probability -- the likelihood that all data points are explained by our parameterization. Since we have assumed that the noise affects each output independently, we can factorize the likelihood, and write:
#
# \begin{align}
# \mathcal{L}(\theta|\mathbf{x}, \mathbf{y}) = \prod_{i=1}^N \mathcal{L}(\theta|x_i,y_i),
# \end{align}
#
# where we have $N$ data points $\mathbf{x} = [x_1,...,x_N]$ and $\mathbf{y} = [y_1,...,y_N]$.
#
#
# In practice, such a product can be numerically unstable. Indeed multiplying small values together can lead to [underflow](https://en.wikipedia.org/wiki/Arithmetic_underflow), the situation in which the digital representation of floating point number reaches its limit. This problem can be circumvented by taking the logarithm of the likelihood because the logarithm transforms products into sums:
#
# \begin{align}
# \log\mathcal{L}(\theta|\mathbf{x}, \mathbf{y}) = \sum_{i=1}^N \log\mathcal{L}(\theta|x_i,y_i)
# \end{align}
#
# We can take the sum of the log of the output of our `likelihood` method applied to the full dataset to get a better idea of how different $\hat{\theta}$ compare. We can also plot the different distribution densities over our dataset and see how they line up qualitatively.
# + cellView="form" id="zGlIm4XnG7tK"
# @markdown Execute this cell to visualize different distribution densities
theta_hats = [0.5, 1.0, 2.2]
fig, axes = plt.subplots(ncols=3, figsize=(16, 4))
for theta_hat, ax in zip(theta_hats, axes):
ll = np.sum(np.log(likelihood(theta_hat, x, y))) # log likelihood
im = plot_density_image(x, y, theta_hat, ax=ax)
ax.scatter(x, y)
ax.set(title=fr'$\hat{{\theta}}$ = {theta_hat}, log likelihood: {ll:.2f}')
plt.colorbar(im, ax=ax);
# + [markdown] id="davoFIjxG7tK"
# Using the log likelihood calculation, we see that $\mathcal{L}(\theta=1.0) > \mathcal{L}(\theta=0.5) > \mathcal{L}(\theta=2.2)$.
#
# This is great: now we have a way to compare estimators based on likelihood. But like with the MSE approach, we want an analytic solution to find the best estimator. In this case, we want to find the estimator that maximizes the likelihood.
#
# + [markdown] id="PvVX4QpuG7tL"
# ## Section 1.4: Finding the Maximum Likelihood Estimator
#
# *Estimated timing to here from start of tutorial: 23 min*
#
# <details>
# <summary> <font color='blue'>Click here for text recap of relevant part of video </font></summary>
#
# We want to find the parameter value $\hat\theta$ that makes our data set most likely:
#
# \begin{align}
# \hat{\theta}_{\textrm{MLE}} = \underset{\theta}{\operatorname{argmax}} \mathcal{L}(\theta|X,Y)
# \end{align}
#
# We discussed how taking the logarithm of the likelihood helps with numerical stability, the good thing is that it does so without changing the parameter value that maximizes the likelihood. Indeed, the $\log()$ function is *monotonically increasing*, which means that it preserves the order of its inputs. So we have:
#
# \begin{align}
# \hat{\theta}_{\textrm{MLE}} = \underset{\theta}{\operatorname{argmax}} \sum_{i=1}^m \textrm{log} \mathcal{L}(\theta|x_i,y_i)
# \end{align}
#
# Now substituting our specific likelihood function and taking its logarithm, we get:
# \begin{align}
# \hat{\theta}_{\textrm{MLE}} = \underset{\theta}{\operatorname{argmax}} [-\frac{N}{2} \operatorname{log} 2\pi\sigma^2 - \frac{1}{2\sigma^2}\sum_{i=1}^N (y_i-\theta x_i)^2].
# \end{align}
#
# Note that maximizing the log likelihood is the same as minimizing the negative log likelihood (in practice optimization routines are developed to solve minimization not maximization problems). Because of the convexity of this objective function, we can take the derivative of our negative log likelihhood, set it to 0, and solve - just like our solution to minimizing MSE.
#
# \begin{align}
# \frac{\partial\operatorname{log}\mathcal{L}(\theta|x,y)}{\partial\theta}=\frac{1}{\sigma^2}\sum_{i=1}^N(y_i-\theta x_i)x_i = 0
# \end{align}
#
# This looks remarkably like the equation we had to solve for the optimal MSE estimator, and, in fact, we arrive to the exact same solution!
#
# \begin{align}
# \hat{\theta}_{\textrm{MLE}} = \hat{\theta}_{\textrm{MSE}} = \frac{\sum_{i=1}^N x_i y_i}{\sum_{i=1}^N x_i^2}
# \end{align}
# + id="SRVJL8iJG7tL"
# Compute theta_hat_MLE
theta_hat_mle = (x @ y) / (x @ x)
# + cellView="form" id="J2ku5KIVG7tL"
#@title
#@markdown Execute this cell to visualize density with theta_hat_mle
# Plot the resulting distribution density
fig, ax = plt.subplots()
ll = np.sum(np.log(likelihood(theta_hat_mle, x, y))) # log likelihood
im = plot_density_image(x, y, theta_hat_mle, ax=ax)
plt.colorbar(im, ax=ax);
ax.scatter(x, y)
ax.set(title=fr'$\hat{{\theta}}$ = {theta_hat_mle:.2f}, log likelihood: {ll:.2f}');
# + [markdown] id="WvJVY_TfG7tL"
# ---
# # Summary
#
# *Estimated timing of tutorial: 30 minutes*
#
#
# Likelihood vs probability:
#
# - $\mathcal{L}(\theta|x, y) = p(y|x, \theta)$
# - $p(y|x, \theta)$ -> "probability of observing the response $y$ given parameter $\theta$ and input $x$"
# - $\mathcal{L}(\theta|x, y)$ -> "likelihood model that parameters $\theta$ produced response $y$ from input $x$"
#
# Log-likelihood maximization:
#
# - We take the $\log$ of the likelihood function for computational convenience
# - The parameters $\theta$ that maximize $\log\mathcal{L}(\theta|x, y)$ are the model parameters that maximize the probability of observing the data.
#
# **Key point**: the log-likelihood is a flexible cost function, and is often used to find model parameters that best fit the data.
# + [markdown] id="6Ls_ehS3G7tL"
#
# ---
# # Notation
#
# \begin{align}
# x &\quad \text{input, independent variable}\\
# y &\quad \text{response measurement, dependent variable}\\
# \mathbf{x} &\quad \text{vector of input values}\\
# \mathbf{y} &\quad \text{vector of measurements}\\
# \epsilon &\quad \text{measurement error, noise contribution}\\
# \epsilon \sim \mathcal{N}(\mu, \sigma^2) &\quad \text{random variable } \epsilon \text{ is distributed according to a Gaussian distribution, with mean } \mu \text{ and variance } \sigma^2\\
# \mu &\quad \text{mean}\\
# \sigma^2 &\quad \text{variance}\\
# \sigma &\quad \text{standard deviation}\\
# \theta &\quad \text{parameter}\\
# \hat{\theta} &\quad \text{estimate of parameter}\\
# \mathcal{L}(\theta|x, y) &\quad \text{likelihood of that parameter } \theta \text{ producing response } y \text{ from input } x \\
# p(y|x, \theta) &\quad \text{probability of observing the response } y \text{ given input } x \text{ and parameter } \theta \\
# \end{align}
# + [markdown] id="QVmR-By3G7tM"
# ---
# # Bonus
# + [markdown] id="TFVz_KFyG7tM"
# We can also see $\mathrm{p}(\mathrm{y} | \mathrm{x}, \theta)$ as a function of $x$. This is the stimulus likelihood function, and it is useful in case we want to decode the input $x$ from observed responses $y$. This is what is relevant from the point of view of a neuron that does not have access to the outside world and tries to infer what's out there from the responses of other neurons!
#
#
#
#
| tutorials/W4D1_How_do_we_know_how_certain_we_should_be/TA/W4D1_Tutorial1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Flight seats downloader
#
# Data taken from [US Bureau of Transportation](https://www.transtats.bts.gov/DL_SelectFields.asp?Table_ID=236)
#
# Field descriptions provided [here](https://www.transtats.bts.gov/Fields.asp?Table_ID=311)
import requests
from selenium import webdriver
# +
chrome_options = webdriver.ChromeOptions()
prefs = {'download.default_directory' : '/Users/jaromeleslie/Documents/MDS/Personal_projects/Ohare_taxi_demand/data/seats'}
chrome_options.add_experimental_option('prefs', prefs)
driver = webdriver.Chrome(chrome_options=chrome_options)
driver.get('https://www.transtats.bts.gov/DL_SelectFields.asp?Table_ID=311')
# -
targ_years= list(range(2013,2020,1))
targ_years = list(map(str,targ_years))
# +
#STEP 1. LOCATE DOWNLOAD BUTTON
download_bt = driver.find_element_by_xpath('//*[@id="content"]/table[1]/tbody/tr/td[2]/table[3]/tbody/tr/td[2]/button[1]')
# download_bt.click()
# +
#STEP 2. SELECT FIELDS OF INTEREST (IGNORING DEFAULTS)
# DEPARTURES SCHEDULED
dep_sched_sel = driver.find_element_by_xpath('/html/body/div[3]/div[3]/table[1]/tbody/tr/td[2]/table[4]/tbody/tr[3]/td[1]/input')
dep_sched_sel.click()
# DEPARTURES PERFORMED
dep_perf_sel = driver.find_element_by_xpath('/html/body/div[3]/div[3]/table[1]/tbody/tr/td[2]/table[4]/tbody/tr[4]/td[1]/input')
dep_perf_sel.click()
# SEATS
seats_sel = driver.find_element_by_xpath('/html/body/div[3]/div[3]/table[1]/tbody/tr/td[2]/table[4]/tbody/tr[6]/td[1]/input')
seats_sel.click()
# PASSENGERS
pass_sel = driver.find_element_by_xpath('/html/body/div[3]/div[3]/table[1]/tbody/tr/td[2]/table[4]/tbody/tr[7]/td[1]/input')
pass_sel.click()
# +
#STEP 3. LOOP OVER YEARS OF INTEREST
#FIND DROPDOWN FOR SELECTABLE YEARS
year_sel = driver.find_element_by_id("XYEAR")
all_years = year_sel.find_elements_by_tag_name("option")
#OUTER LOOP FOR EACH YEAR
for year in all_years:
if year.get_attribute("value") in targ_years:
print("Value is: %s" % year.get_attribute("value"))
year.click()
#EXECUTE DOWNLOAD
download_bt.click()
# -
# # Merge downloads into single file
# +
#STARTING WITH 84 ZIPFILES, MAKE ORD_OTP.CSV
import pandas as pd
import numpy as np
import zipfile as zp
import requests
from selenium import webdriver
from bs4 import BeautifulSoup
from datetime import datetime
import pytest
# +
entries = []
for i in range(72,84,1):
if i == 0:
with zp.ZipFile('../data/seats/691436399_T_T100D_SEGMENT_ALL_CARRIER.zip') as myzip:
myzip.extract('691436399_T_T100D_SEGMENT_ALL_CARRIER.csv',path='../data/seats')
df = pd.read_csv('../data/seats/691436399_T_T100D_SEGMENT_ALL_CARRIER.csv')
entries.append(df.query('DEST == "ORD"'))
else:
with zp.ZipFile('../data/otp/1051953426_T_ONTIME_REPORTING '+'('+str(i)+').zip') as myzip:
myzip.extract('1051953426_T_ONTIME_REPORTING.csv', path ='../data/otp')
df = pd.read_csv('../data/otp/1051953426_T_ONTIME_REPORTING.csv')
entries.append(df.query('DEST == "ORD"'))
# -
combined_ord_seats = pd.concat(entries)
combined_ord_seats.head()
| src/auto_download_seats.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.5 64-bit (''maindev'': conda)'
# metadata:
# interpreter:
# hash: 8ef5afb6cf1f6c39027afcc5ffa2be2d0694f57907bf88ebd75e0075e6672cb9
# name: python3
# ---
# # JSMP: Model Development
# ## Return classification with LightGBM
# ***
#
# The goal here was to build a multi-category classifier for different return buckets using LightGBM. Instead of a more typical positive vs negative binary approach, I was hoping to also gauge the magnitude and the likelihood of having extreme gains vs extreme losses. The difficulty then lay into transforming these insights into a 0/1 *action* decision and I ended up abandoning this approach. Would be curious to see if anyone chose this approach and made it work!
#
# Below is just a sample quick training run to give an idea of how this approach worked.
import os
import jsmp
jsmp.env_config("config.json")
# ## Setup
#
# ### File paths
# set competition, training data paths
comp_dir = os.path.join(os.environ.get("DATA_DIR"), "jane-street-market-prediction")
pq_dir = os.path.join(comp_dir, "train") # training data in Parquet format!
# ### Training configuration
#
# Below I define `train_config`, which, in addition to typical LightGBM hyperparameters, also includes:
# * `'resp_splits'`: the split values to create discrete return bins for classificaiton. Bins the capture returns below -0.033 and above 0.033 are automatically added.
# * `'date_splits'`: implies that 100-380 is the training period, 380-440 the validation period and 440-499 the evaluation period.
train_config = {
"resp_splits": [-0.033, -0.005, 0, 0.005, 0.033],
"date_splits": [100, 380, 440],
"n_rounds": {'total': 100, 'early': 5},
"params": {
"objective": "multiclass",
"num_class": 6,
"boosting": "gbdt",
"eta": 0.05,
"num_leaves": 30,
"bagging_freq": 1,
"bagging_fraction": 0.6,
"feature_fraction": 0.6,
"seed": 42
}
}
# ## Model training
#
# Here I redeploy the above `train_config` in my custom-made training wrapper function:
lgb_model = jsmp.train_lgb_classifier(pq_dir, train_config, verbose=True)
# ## Predicting return bins
eval_df = jsmp.query_train_pq(pq_dir, date_range=[440, 499], return_cols=['resp', 'weight'])
pred_df = jsmp.gen_return_bins(eval_df, splits=train_config['resp_splits'])
pred_df = jsmp.predict_return_bin(pred_df, lgb_model)
pred_df
# ### Confusion matrix (percentages)
#
# We can see that the model can learn a systematic ability to predict the correct class more often than random chance. Also, the model appears to be quite strong in classifying **absolute** returns - in other words, it can identify volatility very accurately (even without using `weight` as a feature), but the direction of the return remains a key challenge!
n_bins = train_config['params']['num_class']
jsmp.confusion_matrix(pred_df, n_bins)
# ### Confusion matrix (counts)
jsmp.confusion_matrix(pred_df, n_bins, as_percent=False)
# ## Action evaluation
#
# ### Naive rule
#
# Given the model appears to perform better at the extreme ends, I ignored the median 2 bins and compared the sum of probabilities in the 2 lowest and 2 highest return bins to determine the optimal action:
good_bins = ["resp_bin_" + str(i) for i in range(n_bins) if i > (n_bins/2)]
bad_bins = ["resp_bin_" + str(i) for i in range(n_bins) if i < (n_bins/2 - 1)]
pred_df.loc[:, "action"] = (pred_df[good_bins].sum(axis=1) >
pred_df[bad_bins].sum(axis=1)).astype(int)
u, profit = jsmp.compute_utility(pred_df, verbose=True)
# ### Avoiding large weights
#
# I also found that large weights made a big impact on the final utility. My takeway was that if you can't predict well for trades with large weights, just ignore them and your score will increase! Also, larger weights are associated with lower volatility and that's where the model struggled to consistently predict the direction of returns.
pred_df.loc[(pred_df['weight'] > 40), "action"] = 0
u, profit = jsmp.compute_utility(pred_df, verbose=True)
# ## Conclusion
#
# * It was challenging to systematically come up with an optimal action function given a set of return class probabilities
# * Ultimately, I felt that a building a model that directly predicted the action would work best and that's when I started experimenting with Neural Networks
| lightGBM_experiment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.5 64-bit
# name: python395jvsc74a57bd0b62550ee9875c103988241e840a33346a8e052936219caf3064f7a25db6fe747
# ---
# +
import pandas
# wget https://raw.githubusercontent.com/MicrosoftDocs/mslearn-introduction-to-machine-learning/main/graphing.py
# wget https://raw.githubusercontent.com/MicrosoftDocs/mslearn-introduction-to-machine-learning/main/Data/doggy-boot-harness.csv
# Read the text file containing data using pandas
dataset = pandas.read_csv('doggy-boot-harness.csv')
# Print the data
# Because there are a lot of data, use head() to only print the first few rows
dataset.head()
# +
# Look at the harness sizes
print("Harness sizes")
print(dataset.harness_size)
# Remove the sex and age-in-years columns.
del dataset["sex"]
del dataset["age_years"]
# Print the column names
print("\nAvailable columns after deleting sex and age information:")
print(dataset.columns.values)
# +
# Print the data at the top of the table
print("TOP OF TABLE")
print(dataset.head())
# print the data at the bottom of the table
print("\nBOTTOM OF TABLE")
print(dataset.tail())
# +
# Print how many rows of data we have
print(f"We have {len(dataset)} rows of data")
# Determine whether each avalanche dog's harness size is < 55
# This creates a True or False value for each row where True means
# they are smaller than 55
is_small = dataset.harness_size < 55
print("\nWhether the dog's harness was smaller than size 55:")
print(is_small)
# Now apply this 'mask' to our data to keep the smaller dogs
data_from_small_dogs = dataset[is_small]
print("\nData for dogs with harness smaller than size 55:")
print(data_from_small_dogs)
# Print the number of small dogs
print(f"\nNumber of dogs with harness size less than 55: {len(data_from_small_dogs)}")
# +
# Make a copy of the dataset that only contains dogs with
# a boot size below size 40
# The call to copy() is optional but can help avoid unexpected
# behaviour in more complex scenarios
data_smaller_paws = dataset[dataset.boot_size < 40].copy()
# Print information about this
print(f"We now have {len(data_smaller_paws)} rows in our dataset. The last few rows are:")
data_smaller_paws.tail()
# +
# Load and prepare plotly to create our graphs
import plotly.express
import graphing # this is a custom file you can find in our code on github
# Show a graph of harness size by boot size:
plotly.express.scatter(data_smaller_paws, x="harness_size", y="boot_size")
# +
# Convert harness sizes from metric to imperial units
# and save the result to a new column
data_smaller_paws['harness_size_imperial'] = data_smaller_paws.harness_size / 2.54
# Show a graph but not in imperial units
plotly.express.scatter(data_smaller_paws, x="harness_size_imperial", y="boot_size")
| ML-For-Beginners/100-introduction-to-machine-learning/5-exercise-improve-models.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/mohanrajmit/Safety_Detection/blob/master/Helmet_Vest_Detection.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="jvtly9PG32X9" colab_type="code" outputId="a05c318a-262d-4ffe-ead2-d2863a3a753b" colab={"base_uri": "https://localhost:8080/", "height": 141}
# !git clone https://github.com/AlexeyAB/darknet.git
# + id="t6scTjLd3-Qf" colab_type="code" outputId="579c2a7e-e4c6-42a6-d8b3-859bbbc79421" colab={"base_uri": "https://localhost:8080/", "height": 35}
# cd /content/darknet
# + id="u4CGuKcJ4AZH" colab_type="code" outputId="5835bee6-6318-4294-c100-46b35dfc5d56" colab={"base_uri": "https://localhost:8080/", "height": 124}
# !ls
# + id="F_kYLRMN4XiI" colab_type="code" outputId="a38fb65c-97fb-43bd-a927-cb37f7146cc8" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# !make
# + id="iObW4HcV5Z5s" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 141} outputId="014c8d3b-2739-4a3f-fbcf-61679fc1e11a"
# !ls
# + id="mJWwbwuq4ZE1" colab_type="code" outputId="8d6fc627-025b-4f97-d24d-7d046c33dce9" colab={"base_uri": "https://localhost:8080/", "height": 35}
# ! ./darknet
# + id="IdJ6-6Jb4s5g" colab_type="code" outputId="794829bb-2350-496e-d63d-9b0abad507b8" colab={"base_uri": "https://localhost:8080/", "height": 212}
# !wget https://pjreddie.com/media/files/darknet19_448.conv.23
# + id="hmdEP12WjVSj" colab_type="code" colab={}
# !rm -rf /content/darknet/Safety_Detection
# + id="DNnU-E8U4zKv" colab_type="code" outputId="118eef90-3560-4bcf-ed33-5309bdc97c6c" colab={"base_uri": "https://localhost:8080/", "height": 141}
# !git clone https://github.com/mohanrajmit/Safety_Detection.git
# + id="0bDNJGzL44GB" colab_type="code" colab={}
# mv /content/darknet/Safety_Detection/dataset /content/darknet
# + id="wuCkJSp4486_" colab_type="code" outputId="40a9d1cc-59d6-4c36-b18c-d37f83677119" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# !./darknet detector train cfg/obj.data cfg/yolo-voc.2.0.cfg darknet19_448.conv.23 -dont_show 0
| Yolo_V4/Yolo_V2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import json
# +
experiments_path = "../"
def fc_to_string(k,cfg):
res = f"k_{k}bits_"
for layer in cfg:
res += str(layer["bits"]) + "."
res += "logsize_"
for layer in cfg:
res = res + str(layer["logsize"]) + "."
return res
def bits_str(cfg):
res = "b."
for layer in cfg:
res += str(layer["bits"]) + "."
return res
def bits_valid(cfg):
for layer in cfg:
if layer["bits"] > 128:
return False
return True
def logsize_str(cfg):
res = "sz."
for layer in cfg:
res = res + str(layer["logsize"]) + "."
return res
def load_configs(name):
with open(f'{experiments_path}config_{name}', 'r') as myfile:
data=myfile.read()
obj = json.loads(data)
return obj["config"]
def load_exp(fn):
try:
with open(fn, 'r') as myfile:
data=myfile.read()
# print(data)
obj = json.loads(data)
return obj
except Exception as err:
print(err)
return None
datasets =[
("Twitter 1 mio","twitter_1mio_coords.h5"),
("Twitter 10 mio","twitter_10mio_coords.h5"),
("Twitter 100 mio","twitter_100mio_coords.h5"),
("Twitter 200 mio","twitter_200mio_coords.h5"),
("Asia 500 mio","asia_500mio_coords.h5"),
("Asia 1 bil","asia_1bil_coords.h5")
]
# +
k = 8;
x = 0;
width = 1 * 8192
height = 1 * 8192;
exp_name = "test_datasets_new";
cfgs = load_configs(exp_name)
# +
import matplotlib.pyplot as plt
import matplotlib.ticker as ticker
import numpy as np
import matplotlib.backends.backend_pdf as pdf
import tqdm
import matplotlib
colors = ["#F72585","#7209B7", "#3A0CA3","#4361EE", "#4CC9F0"]
colors.reverse()
colormap = matplotlib.colors.LinearSegmentedColormap.from_list("neodym",[matplotlib.colors.to_rgb(x) for x in colors])
colors = ["#F72585", "#3A0CA3", "#4CC9F0"]
def save_figs(fn):
print(f"--> Saving all figures... {fn}")
out_pdf = pdf.PdfPages(fn)
for fig in tqdm.tqdm(range(1, plt.gcf().number+1)):
out_pdf.savefig( fig )
out_pdf.close()
plt.close("all")
def plot_matrix(cm, title='Experiment matrix', cmap=colormap, xlabels=None, ylabels=None, cblabel=None):
fig = plt.figure(figsize=(len(xlabels)*1.25, 2+ len(ylabels)))
ax = fig.add_subplot(111)
# ax.xaxis.set_major_locator(ticker.MultipleLocator(2))
# ax.yaxis.set_major_locator(ticker.MultipleLocator(1))
cax = ax.matshow(cm,cmap=cmap)
plt.title(title)
ax.set_xticks(range(len(xlabels)))
ax.set_yticks(range(len(ylabels)))
cbar = fig.colorbar(cax)
plt.xlabel('Sizes')
plt.ylabel('Bits')
if cblabel:
cbar.set_label(cblabel)
if xlabels:
ax.set_xticklabels(list(xlabels))
print(xlabels)
if ylabels:
ax.set_yticklabels(list(ylabels))
print(ylabels)
#plt.show()
def make_maps(size, cfgs):
bits_map = {}
logsize_map = {}
count_bits =0
count_sizes = 0
for c in cfgs :
if len(c) ==size and bits_valid(c):
if(not bits_str(c) in bits_map):
bits_map[bits_str(c)] = count_bits
count_bits+=1
if(not logsize_str(c) in logsize_map):
logsize_map[logsize_str(c)] = count_sizes
count_sizes+=1
return bits_map, logsize_map
def make_maps_smaller(size, cfgs):
bits_map = {}
logsize_map = {}
count_bits =0
count_sizes = 0
for c in cfgs :
if len(c) <size and bits_valid(c):
if(not bits_str(c) in bits_map):
bits_map[bits_str(c)] = count_bits
count_bits+=1
if(not logsize_str(c) in logsize_map):
logsize_map[logsize_str(c)] = count_sizes
count_sizes+=1
return bits_map, logsize_map
def make_maps_any( cfgs):
bits_map = {}
logsize_map = {}
count_bits =0
count_sizes = 0
for c in cfgs :
if bits_valid(c):
if(not bits_str(c) in bits_map):
bits_map[bits_str(c)] = count_bits
count_bits+=1
if(not logsize_str(c) in logsize_map):
logsize_map[logsize_str(c)] = count_sizes
count_sizes+=1
return bits_map, logsize_map
bits_map, logsize_map = make_maps(2,cfgs)
print(len(bits_map), len(logsize_map))
print(bits_map)
print(logsize_map)
a = np.random.rand(len(bits_map), len(logsize_map))
plot_matrix(a, ylabels=bits_map.keys(), xlabels=logsize_map.keys())
# +
figsize=(12,12)
folder_name = "test_datasets_new";
exp_name = "test_datasets_new";
def plot_some(cm, title='Experiment', xlabels=None, ylabels=None, cblabel=None, K=range(1,30)):
fig = plt.figure(figsize=figsize)
ax = fig.add_subplot(111)
# print(xlabels)
# print(ylabels)
sz = 1
lines = ['-', '--', '-.', ':']
i = 0
if xlabels:
ax.set_xticklabels(list(xlabels))
if ylabels:
ax.set_yticklabels(list(ylabels))
plt.plot(list(K), cm)
#plt.legend()
plt.title(title)
plt.xlabel('Size (kb)')
plt.ylabel('Time (Seconds)')
#plt.show()
def make_matrices(size, cfgs):
bits_map, logsize_map = make_maps(size,cfgs)
# print(bits_map)
matrices= {}
matrix_indices = ["insert_time", "query_time"]
for dn, ds in datasets :
matrices[ds] = {}
for mi in matrix_indices:
matrices[ds][mi] = np.zeros((len(bits_map),(len(logsize_map))))-1
x=0
for c in cfgs :
# print(fc_to_string(k,c))
y = 0;
for dn, ds in datasets :
fn = experiments_path + folder_name + "/" + exp_name + f'.w{width}h{height}.-' + fc_to_string(k,c) \
+ ds + ".json"
#print(fn)
data = load_exp(fn)
bits = bits_str(c)
sz = logsize_str(c)
if data and bits in bits_map and sz in logsize_map and bits_valid(c):
for mi in matrix_indices:
matrices[ds][mi][ bits_map[bits],logsize_map[sz]] = data["perf"][mi]
y+=1
x+=1
for dn, ds in datasets :
for mi in matrix_indices:
plot_matrix( matrices[ds][mi],title=f'{dn} {mi}', ylabels=bits_map.keys(), xlabels=logsize_map.keys(), cblabel=f"Time (Seconds)")
make_matrices(1, cfgs)
make_matrices(2, cfgs)
make_matrices(3, cfgs)
save_figs("test_datasets_new_matrices.pdf")
# +
figsize=(12,12)
folder_name = "test_datasets_new";
exp_name = "test_datasets_new";
def plot_some(cm, title='Experiment', xlabels=None, ylabels=None, cblabel=None, K=range(1,30)):
fig = plt.figure(figsize=figsize)
ax = fig.add_subplot(111)
# print(xlabels)
# print(ylabels)
sz = 1
lines = ['-', '--', '-.', ':']
i = 0
if xlabels:
ax.set_xticklabels(list(xlabels))
if ylabels:
ax.set_yticklabels(list(ylabels))
plt.plot(list(K), cm)
#plt.legend()
plt.title(title)
plt.xlabel('Size (kb)')
plt.ylabel('Time (Seconds)')
#plt.show()
def make_matrices(size, cfgs):
bits_map, logsize_map = make_maps(size,cfgs)
# print(bits_map)
matrices= {}
matrix_indices = ["insert_time", "query_time"]
for dn, ds in datasets :
matrices[ds] = {}
for mi in matrix_indices:
matrices[ds][mi] = {}
x=0
for c in cfgs :
# print(fc_to_string(k,c))
y = 0;
for dn, ds in datasets :
fn = experiments_path + folder_name + "/" + exp_name + f'.w{width}h{height}.-' + fc_to_string(k,c) \
+ ds + ".json"
#print(fn)
data = load_exp(fn)
bits = bits_str(c)
sz = logsize_str(c)
if data and bits in bits_map and sz in logsize_map and bits_valid(c):
for mi in matrix_indices:
kb_size = f'{data["summary"]["kb_size"]}kb ({bits}{sz})'
if len(kb_size) > 20:
kb_size = f'{data["summary"]["kb_size"]}'
kb_size = f'{data["summary"]["kb_size"]}'
#print(mi, kb_size, bits, sz)
#if kb_size in matrices[ds][mi]:
# print(mi,"double trouble", kb_size, bits, sz, (matrices[ds][mi][kb_size] == data["perf"][mi]) )
matrices[ds][mi][kb_size] = data["perf"][mi]
y+=1
x+=1
for dn, ds in datasets :
for mi in matrix_indices:
plot_some( list(matrices[ds][mi].values()), K=list(matrices[ds][mi].keys()),title=f'{dn} {mi}')
make_matrices(1, cfgs)
make_matrices(2, cfgs)
make_matrices(3, cfgs)
save_figs("test_datasets_new_plot_seperate.pdf")
# +
figsize=(6,6)
folder_name = "test_datasets_new";
exp_name = "test_datasets_new";
def plot_some(cm, title='Experiment', xlabels=None, ylabels=None, cblabel=None):
fig = plt.figure(figsize=figsize)
ax = fig.add_subplot(111)
# print(xlabels)
# print(ylabels)
sz = 1
lines = ['-', '--', '-.', ':']
i = 0
if xlabels:
ax.set_xticklabels(list(xlabels))
if ylabels:
ax.set_yticklabels(list(ylabels))
for k,v in cm.items():
plt.bar(list(v.keys()), list(v.values()), label=f'{k} layers')
plt.legend(loc='upper right')
plt.title(title)
plt.xlabel('Size (kb)')
plt.ylabel('Time (Seconds)')
plt.xticks(rotation=30, ha='right')
#plt.show()
def make_matrices(size, cfgs):
bits_map, logsize_map = make_maps(size,cfgs)
bits_map, logsize_map = make_maps_smaller(size, cfgs)
# print(bits_map)
matrices= {}
matrix_indices = ["query_time", "query_time_min", "query_time_max"]
for dn, ds in datasets :
matrices[ds] = {}
for mi in matrix_indices:
matrices[ds][mi] = {"global": {}}
for x in range(1,size):
matrices[ds][mi][str(x)] = {}
x=0
for c in cfgs :
# print(fc_to_string(k,c))
y = 0;
for dn, ds in datasets :
fn = experiments_path + folder_name + "/" + exp_name + f'.w{width}h{height}.-' + fc_to_string(k,c) \
+ ds + ".json"
#print(fn)
data = load_exp(fn)
bits = bits_str(c)
sz = logsize_str(c)
#print(c)
if data and bits in bits_map and sz in logsize_map and bits_valid(c):
for mi in matrix_indices:
kb_size = f'{data["summary"]["kb_size"]} {bits}{sz}'
#if len(kb_size) > 20:
# kb_size = f'{data["summary"]["kb_size"]}'
kb_size = f'{data["summary"]["kb_size"]}'
layer = f'{len(c)}'
#print(layer)
#print(mi, kb_size, bits, sz)
#if kb_size in matrices[ds][mi]:
# print(mi,"double trouble", kb_size, bits, sz, (matrices[ds][mi][kb_size] == data["perf"][mi]) )
matrices[ds][mi][layer][kb_size] = data["perf"][mi]
matrices[ds][mi]["global"][kb_size] = data["perf"][mi]
y+=1
x+=1
ds_min = {}
ds_max = {}
ds_mean = {}
ds_std = {}
print(matrices[ds]["query_time"])
for idx in matrices[ds]["query_time"]["global"].keys():
ds_min[idx] = []
ds_max[idx] = []
ds_mean[idx] = []
ds_std[idx] = []
for dn, ds in datasets :
ds_min[idx].append(matrices[ds]["query_time_min"]["global"][idx])
ds_max[idx].append(matrices[ds]["query_time_max"]["global"][idx])
ds_mean[idx].append(matrices[ds]["query_time"]["global"][idx])
ds_std[idx].append(matrices[ds]["query_time"]["global"][idx])
ds_min[idx] = np.min(np.array(ds_min[idx]))
ds_max[idx] = np.max(np.array(ds_max[idx]))
ds_mean[idx] = np.mean(np.array(ds_mean[idx]))
ds_std[idx] = np.std(ds_std[idx])
means = np.array(list(ds_mean.values()))
maxes = np.array(list(ds_max.values()))
mins = np.array(list(ds_min.values()))
std = np.array(list(ds_std.values()))
fig = plt.figure(figsize=figsize)
ax = fig.add_subplot(111)
markers = ["o", "s", "^"]
ds = datasets[0][1]
for i in range(1,4):
x = list(map(int, matrices[ds]["query_time"][str(i)].keys()))
glob_keys = list(matrices[ds]["query_time"]["global"].keys())
glob_keys = zip(glob_keys, range(len(glob_keys)))
gl = {key: value for key, value in glob_keys}
meanind = [gl[k] for k in matrices[ds]["query_time"][str(i)].keys()]
y = [means[j] for j in meanind]
plt.scatter(x , y, label=f"{i} Layers", marker=markers[i-1], s=40,zorder=2, c=colors[i-1])
# create stacked errorbars:
x = list(map(int, matrices[ds]["query_time"]["global"].keys()))
plt.errorbar(x, means, std, fmt='.k', lw=2,capsize=3, label="mean +- std",zorder=1)
plt.errorbar(x , means, [means - mins, maxes - means],
fmt='.k', ecolor='gray', capsize=6, lw=1, label="[min mean max]",zorder=3)
plt.legend(loc='upper left')
plt.title("Query Times")
plt.xlabel('Dataset')
plt.ylabel('Time (Seconds)')
#"""
for dn, ds in datasets :
fig = plt.figure(figsize=figsize)
ax = fig.add_subplot(111)
#plt.bar(list(ds_max.keys()), list(ds_max.values()), label=f'{k} max', color="red")
#plt.bar(list(ds_mean.keys()), list(ds_mean.values()), label=f'{k} mean', color="blue")
#plt.bar(list(ds_min.keys()), list(ds_min.values()), label=f'{k} min', color="green")
means = np.array(list( matrices[ds]["query_time"]["global"].values()))
maxes = np.array(list(matrices[ds]["query_time_max"]["global"].values()))
mins = np.array(list(matrices[ds]["query_time_min"]["global"].values()))
# create stacked errorbars:
x = list(map(int, matrices[ds]["query_time"]["global"].keys()))
plt.errorbar(x , means, [means - mins, maxes - means],
fmt='.k', ecolor='gray', capsize=6, lw=1, label="[min max]",zorder=2)
markers = ["o", "s", "^"]
for i in range(1,4):
x = list(map(int, matrices[ds]["query_time"][str(i)].keys()))
y = matrices[ds]["query_time"][str(i)].values()
plt.scatter(x , y, label=f"{i} Layers", marker=markers[i-1], s=40 ,zorder=1, c=colors[i-1])
plt.legend(loc='upper left')
plt.title(f"{dn}")
plt.xlabel('Dataset')
plt.ylabel('Time (Seconds)')
#make_matrices(1, cfgs)
#make_matrices(2, cfgs)
#make_matrices(3, cfgs)
make_matrices(4, cfgs)
save_figs("test_datasets_new_plot.pdf")
# +
figsize=(6,6)
folder_name = "test_datasets_new";
exp_name = "test_datasets_new";
def make_matrices_ds(size, cfgs):
bits_map, logsize_map = make_maps(size,cfgs)
bits_map, logsize_map = make_maps_smaller(size, cfgs)
# print(bits_map)
matrices= {}
matrix_indices = ["insert_time", "query_time"]
Title= {
"insert_time": "Insert Dataset", "query_time": "Query 10 mio points"
}
for dn, ds in datasets :
matrices[ds] = {}
for mi in matrix_indices:
matrices[ds][mi] = {}
x=0
for c in cfgs :
# print(fc_to_string(k,c))
y = 0;
for dn, ds in datasets :
fn = experiments_path + folder_name + "/" + exp_name + f'.w{width}h{height}.-' + fc_to_string(k,c) \
+ ds + ".json"
#print(fn)
data = load_exp(fn)
bits = bits_str(c)
sz = logsize_str(c)
#print(c)
if data and bits in bits_map and sz in logsize_map and bits_valid(c):
for mi in matrix_indices:
kb_size = f'{data["summary"]["kb_size"]} ({bits}{sz}'
if len(kb_size) > 20:
kb_size = f'{data["summary"]["kb_size"]}'
kb_size = f'{data["summary"]["kb_size"]}'
layer = f'{len(c)}'
#print(layer)
#print(mi, kb_size, bits, sz)
#if kb_size in matrices[ds][mi]:
# print(mi,"double trouble", kb_size, bits, sz, (matrices[ds][mi][kb_size] == data["perf"][mi]) )
matrices[ds][mi][kb_size] = data["perf"][mi]
y+=1
x+=1
for mi in matrix_indices:
ds_min = {}
ds_max = {}
ds_mean = {}
ds_std = {}
for dn, ds in datasets :
ar = np.array(list(matrices[ds][mi].values()) )
#print(ar)
ds_min[dn] = np.min(ar)
ds_max[dn] = np.max(ar)
ds_mean[dn] = np.mean(ar)
ds_std[dn] = np.std(ar)
fig = plt.figure(figsize=figsize)
ax = fig.add_subplot(111)
#plt.bar(list(ds_max.keys()), list(ds_max.values()), label=f'{k} max', color="red")
#plt.bar(list(ds_mean.keys()), list(ds_mean.values()), label=f'{k} mean', color="blue")
#plt.bar(list(ds_min.keys()), list(ds_min.values()), label=f'{k} min', color="green")
means = np.array(list(ds_mean.values()))
maxes = np.array(list(ds_max.values()))
mins = np.array(list(ds_min.values()))
std = np.array(list(ds_std.values()))
# create stacked errorbars:
plt.errorbar(ds_max.keys(), means, std, fmt='ok', lw=3,capsize=3, label="mean +- std")
plt.errorbar(ds_max.keys(), means, [means - mins, maxes - means],
fmt='.k', ecolor='gray', capsize=6, lw=1, label="[min max]")
plt.legend(loc='upper left')
plt.title(Title[mi])
plt.xlabel('Dataset')
plt.ylabel('Time (Seconds)')
plt.xticks(rotation=30, ha='right')
#make_matrices(1, cfgs)
#make_matrices(2, cfgs)
make_matrices_ds(3, cfgs)
save_figs("test_datasets_new_ds_size.pdf")
# -
| experiments/notebooks/.ipynb_checkpoints/test_datasets_full_time-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # What am I doing here?
#
# - GBT model
# - Setting up doing three runs, can change to three different models for a voting classifier
# - Every time I checkpoint a step to a file, it's in an 'if False' block. If you need to
# create a file, change that to True to make the file. Then change it back to False to
# get the faster way through the notebook.
import pyspark
import pyspark.sql.functions as F
import pyspark.sql.types as T
from pyspark.ml.feature import VectorAssembler
from pyspark.ml.tuning import TrainValidationSplit, ParamGridBuilder
from pyspark.ml.classification import GBTClassifier
from pyspark.ml.evaluation import BinaryClassificationEvaluator
# This is optional stuff - either pip install watermark
# or just comment it out (it just keeps track of what library
# versions I have)
# %load_ext watermark
# %watermark -iv
# Comment these out to run on a cluster. Also, adjust memory to size of your laptop
pyspark.sql.SparkSession.builder.config('spark.driver.memory', '8g')
pyspark.sql.SparkSession.builder.config('spark.sql.shuffle.paritions', 5)
spark = pyspark.sql.SparkSession.builder.getOrCreate()
# # Global Variables
unigrams = [ 'os', 'channel', 'app' ]
bigrams = [[ 'device', 'os'],
['device', 'channel'],
['device', 'app'],
['channel', 'app']]
# # Checkpoint 1
#
# Read the csv file, drop the attributed_time (because I didn't use it in the MVP),
# and downsample the 0 class to 25% because I'm still on my laptop
if False:
df = spark.read.csv('../data/train.csv',
header=True, inferSchema=True)
df = df.drop('attributed_time')
df = df.sampleBy('is_attributed', fractions={0:.25,1:1.})
test = spark.read.csv('../data/test.csv',
header= True, inferSchema=True)
df.write.parquet('../data/checkpoint1.parquet', mode='overwrite')
test.write.parquet('../data/test_checkpoint1.parquet', mode='overwrite')
else:
df = spark.read.parquet('../data/checkpoint1.parquet')
test = spark.read.parquet('../data/test_checkpoint1.parquet')
df.dtypes
test.dtypes
df.count()
# +
test.count()
# -
# # Daily IP prevalence
# Because IP addresses get reassigned, need to do these as feature engineering on train and test
# sets separately.
# (See the link Elyse posted on the slack.)
df = df.withColumn('doy',
F.dayofyear('click_time'))
test = test.withColumn('doy',
F.dayofyear('click_time'))
df_ip_counts = df[['doy', 'ip']].groupby(['doy','ip']).count()
test_ip_counts = test[['doy', 'ip']].groupby(['doy', 'ip']).count()
df_day_max = df_ip_counts[['doy','count']]\
.groupby(['doy'])\
.max()\
.withColumnRenamed('max(count)', 'day_max')\
.drop('max(doy)')
test_day_max = test_ip_counts[['doy','count']]\
.groupby(['doy'])\
.max()\
.withColumnRenamed('max(count)', 'day_max')\
.drop('max(doy)')
df_ip_counts = df_ip_counts.join(df_day_max,
['doy'],
how='left')
test_ip_counts = test_ip_counts.join(test_day_max,
['doy'],
how='left')
df_ip_counts.dtypes
# +
df_ip_counts = df_ip_counts.withColumn('ip_pct',
F.col('count').astype(T.FloatType())/
F.col('day_max').astype(T.FloatType()))
test_ip_counts = test_ip_counts.withColumn('ip_pct',
F.col('count').astype(T.FloatType())/
F.col('day_max').astype(T.FloatType()))
# -
df = df.join(
df_ip_counts[['doy','ip','ip_pct']],
on=['doy','ip'],
how='left'
)
test = test.join(
test_ip_counts[['doy','ip','ip_pct']],
on=['doy','ip'],
how='left'
)
# ## Same class balancing as MVP
# Still hacky - but I reordered it so that the join happens on a
# smaller table.
# And, now there are three versions to stack.
class1_a = df.filter(df.is_attributed == 1).sample(
withReplacement=True, fraction=4.0, seed=111)
class1_b = df.filter(df.is_attributed == 1).sample(
withReplacement=True, fraction=4.0, seed=222)
class1_c = df.filter(df.is_attributed == 1).sample(
withReplacement=True, fraction=4.0, seed=333)
df_a = df.sampleBy('is_attributed', {0:.11}, seed=111).unionAll(class1_a)
df_b = df.sampleBy('is_attributed', {0:.11}, seed=222).unionAll(class1_b)
df_c = df.sampleBy('is_attributed', {0:.11}, seed=333).unionAll(class1_c)
# ## Counting
#
# Built count tables except for IP with the full training set rather than the
# subset. Results here.
def get_count_table( group ):
if type(group) == str:
column_name = group + '_pct' # for example: ip_pct
else:
column_name = "_".join(group) # for example: device_os
table_name = 'table_' + column_name
counts_sdf = spark.read.parquet(f'../data/{table_name}.parquet')
return counts_sdf
def join_table( sdf, count_table, group ):
sdf = sdf.join(count_table, group, how='left')
return sdf
# create the count columns with the training data
# write everything out to disk so we don't have to redo
# feature engineering when all I want to do is tune hyperparameters
if False:
for c in unigrams:
ct = get_count_table( c )
df_a = join_table(df_a, ct, [c])
df_b = join_table(df_b, ct, [c])
df_c = join_table(df_c, ct, [c])
test = join_table(test, ct, [c])
for bigram in bigrams:
ct = get_count_table( bigram )
df_a = join_table(df_a, ct, bigram)
df_b = join_table(df_b, ct, bigram)
df_c = join_table(df_c, ct, bigram)
test = join_table(test, ct, bigram)
df_a.write.parquet('../data/dfa.parquet', mode='overwrite')
df_b.write.parquet('../data/dfb.parquet', mode='overwrite')
df_c.write.parquet('../data/dfc.parquet', mode='overwrite')
test.write.parquet('../data/test_stack.parquet', mode='overwrite')
else:
df_a = spark.read.parquet('../data/dfa.parquet')
df_b = spark.read.parquet('../data/dfb.parquet')
df_c = spark.read.parquet('../data/dfc.parquet')
test = spark.read.parquet('../data/test_stack.parquet')
df_a = df_a.fillna(0)
df_b = df_b.fillna(0)
df_c = df_c.fillna(0)
test = test.fillna(0)
for sdf in [ df_a, df_b, df_c ]:
sdf.groupby('is_attributed').count().show()
test.count()
# # Last minute model tweak - add hour column
# +
def add_hour(sdf):
return sdf.withColumn('hour',
(F.hour('click_time').astype(T.FloatType()) +
(F.minute('click_time').astype(T.FloatType()) / 60.)) / 24. )
test = add_hour(test)
df_a = add_hour(df_a)
df_b = add_hour(df_b)
df_c = add_hour(df_c)
# -
# # Create model data in format expected by Spark
input_cols = [ c + '_pct' for c in unigrams ]
input_cols += [ '_'.join(b) for b in bigrams ]
input_cols += ['ip_pct', 'hour']
input_cols
vec_assembler = VectorAssembler(inputCols=input_cols, outputCol = 'features')
evaluator = BinaryClassificationEvaluator(labelCol = 'is_attributed')
model_a = vec_assembler.transform(df_a).select('is_attributed', 'features')
model_b = vec_assembler.transform(df_b).select('is_attributed', 'features')
model_c = vec_assembler.transform(df_c).select('is_attributed', 'features')
# # GBT Classifier
# +
gbtc = GBTClassifier(
labelCol = 'is_attributed',
)
# Preparting for future hyperparameter tuning
pg = ParamGridBuilder(
).addGrid(
gbtc.maxDepth, [ 10 ]
).addGrid(
gbtc.subsamplingRate, [ .8 ]
).addGrid(
gbtc.featureSubsetStrategy, [ '6' ]
).addGrid(
gbtc.maxBins, [ 64 ]
).addGrid(
gbtc.stepSize, [ .2 ]
).addGrid(
gbtc.maxIter, [ 30 ]
).build(
)
tvs = TrainValidationSplit(
estimator = gbtc,
estimatorParamMaps = pg,
evaluator = evaluator,
trainRatio = .8
)
# -
tvs_a = tvs.fit(model_a)
results_a = tvs_a.transform(model_a)
evaluator.evaluate(results_a)
tvs_a.bestModel.extractParamMap()
tvs_b = tvs.fit(model_b)
results_b = tvs_b.transform(model_b)
evaluator.evaluate(results_b)
tvs_c = tvs.fit(model_c)
results_c = tvs_c.transform(model_c)
evaluator.evaluate(results_c)
# # Let's bring the test set in here
test_model = vec_assembler.transform(test)
results_a = tvs_a.transform(test_model)
results_b = tvs_b.transform(test_model)
results_c = tvs_c.transform(test_model)
def get_prediction(sdf):
sdf = sdf.select('click_id',
F.col('prediction').astype(T.ShortType()),
'probability')
sdf.groupby('prediction').count().show()
return sdf
results_a = get_prediction(results_a)
results_b = get_prediction(results_b)
results_c = get_prediction(results_c)
# # Extract probabilities
# +
mySchema = T.StructType([
T.StructField('click_id', T.IntegerType()),
T.StructField('prediction', T.ShortType()),
T.StructField('pclass1', T.FloatType())
])
def save_stuff(x):
return T.Row(click_id=x.click_id,
prediction=x.prediction,
pclass1=float(x.probability[1]))
vec_a = results_a.rdd.map(lambda x: save_stuff(x)).toDF(schema=mySchema)
vec_b = results_b.rdd.map(lambda x: save_stuff(x)).toDF(schema=mySchema)
vec_c = results_c.rdd.map(lambda x: save_stuff(x)).toDF(schema=mySchema)
# -
# # Take the median of the three models as my final answer
# +
vec_a = vec_a.select('click_id',
F.col('pclass1').alias('vec_a') )
vec_b = vec_b.select('click_id',
F.col('pclass1').alias('vec_b') )
vec_c = vec_c.select('click_id',
F.col('pclass1').alias('vec_c') )
joined = vec_a.join(vec_b, ['click_id']).join(vec_c, ['click_id'])
mySchema = T.StructType([
T.StructField('click_id', T.IntegerType()),
T.StructField('is_attributed', T.FloatType())
])
from statistics import median
def get_predict(x):
return T.Row(click_id=x.click_id,
is_attributed=median([x.vec_a, x.vec_b, x.vec_c]))
joined = joined.rdd.map(lambda x: get_predict(x)).toDF(schema=mySchema)
# -
joined.write.csv('../data/vote_results.csv', mode='overwrite')
spark.stop()
| notebook/Stack_GBT.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:loca]
# language: python
# name: conda-env-loca-py
# ---
# +
# %matplotlib inline
import geopandas
import regionmask
import pandas as pd
import xarray as xr
# -
# create GeoDataFrame with all HUC2 elements
# huc 2 to 4, should WBDHU2.shp
dfs = []
hucs = list(range(1, 19))
for i in hucs:
huc2_file = f'/glade/u/home/jhamman/projects/storylines/data/gis/WBD_{i:02d}_Shape/Shape/WBDHU2.shp'
gdf = geopandas.GeoDataFrame.from_file(huc2_file)
dfs.append(gdf)
gdf = pd.concat(dfs)
gdf.index = hucs
gdf.head()
# Create a Regions object for masking
huc_mask = regionmask.Regions_cls('USmask', range(len(gdf)), gdf.STATES.values, list(map(str, hucs)), gdf.geometry.values)
huc_mask.plot()
# load an xarray dataset
airtemps = xr.tutorial.load_dataset('air_temperature')
display(airtemps)
airtemps['air'].isel(time=0).plot()
# Create a mask object
mask = huc_mask.mask(airtemps, wrap_lon=True)
display(mask)
mask.plot()
# Group data by huck.
data_by_huc = airtemps.groupby(mask).mean('stacked_lat_lon')
display(data_by_huc)
# calculate the annual cycle and plot
data_by_huc.air.groupby('time.month').mean('time').plot()
| notebooks/prior_Aug2019/huc_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import torch
from torch.distributions import MultivariateNormal
import math
# ## Multivariate Bayesian Inferencing of Mean of Gaussian likelihood, known Precision
#
# Previously, we studied the univariate case. Now let us consider the scenario where the data instances are multi-dimensional i.e vectors. This leads us to the multivariate case.
#
# In particular the training dataset is
# $$
# X \equiv \left\lbrace
# \vec{x}^{ \left( 1 \right) }, \vec{x}^{ \left( 2 \right) }, \cdots, \vec{x}^{ \left( i \right) }, \cdots, \vec{x}^{ \left( n \right) }
# \right\rbrace
# $$
#
# Here, we assume the variance is known (a constant) but the mean of the data is unknown, modeled as a Gaussian random variable.
#
# We will express the Gaussian in terms of the precision matrix ${\Lambda}$, instead of the covariance matrix ${\Sigma}$ where ${\Lambda} = {\Sigma}^{-1}$.
#
# Since we assume that the data is Normally distributed:
# $$
# p\left( X \middle\vert \vec{\mu} \right) \propto e^{ -\frac{1}{2} \sum_{i=1}^{n} \left( \vec{x}^{ \left( i \right) } - \vec{ \mu } \right)^{T} {\Lambda} \left( \vec{x}^{ \left( i \right) } - \vec{ \mu } \right) }$$
#
# The variance is known - hence it is treated as a constant as opposed to a random variable.
#
# The mean $\vec{\mu}$ is unknown and is treated as a random variable. This too is assumed to be a Gaussian, with mean $\vec{\mu_{0}}$ and precision matrix $\Lambda_{0}$ (not to be confused with $\vec{\mu}$ and $\Lambda$ - the mean and precision matrix of the data itself ). Hence, the prior is
#
# $$p\left( \vec{\mu }\right) \propto e^{ -\frac{1}{2} \left( \vec{\mu }- \vec{ \mu_{0} }\right)^{T} {\Lambda}_{0} \left( \vec{\mu }- \vec{ \mu_{0} }\right) }
# $$
#
#
# Using Bayes theorem, the posterior probability is
#
# $$\overbrace{
# p\left(\vec{\mu} \middle\vert X \right)
# }^{posterior}
# =
# \overbrace{
# p\left( X \middle\vert \vec{\mu} \right)
# }^{likelihood}
# \overbrace{
# p\left(\vec{\mu} \right)
# }^{prior}$$
#
# The right hand side is the product of two Gaussians, which is a Gaussian itself. Let us denote its mean and precision matrix as $\vec{\mu_{n}}$ and $\Lambda_{n}$.
#
# where
# $$
# \begin{align*}
# &{\Lambda}_{n} = n {\Lambda} + {\Lambda}_{0} \\
# & \vec{\mu_{n}} = {\Lambda}_{n}^{-1} \left( n{\Lambda} \bar{\vec{x}} + {\Lambda}_{0} \vec{\mu}_{0} \right)
# \end{align*}$$
#
def inference_known_precision(X, prior_dist, precision_known):
mu_mle = X.mean(dim=0)
n = X.shape[0]
# Parameters of the prior
mu_0 = prior_dist.mean
precision_0 = prior_dist.precision_matrix
# Parameters of posterior
precision_n = n * precision_known + precision_0
mu_n = torch.matmul(n * torch.matmul(mu_mle.unsqueeze(0), precision_known) + torch.matmul(mu_0.unsqueeze(0), precision_0), torch.inverse(precision_n))
posterior_dist = MultivariateNormal(mu_n, precision_matrix=precision_n)
return posterior_dist
# Let us assume that the true distribution is a normal distribution. The true distribution corresponds
# to a single class.
precision_known = torch.tensor([[0.1, 0], [0, 0.1]], dtype=torch.float)
true_dist = MultivariateNormal(torch.tensor([20, 10], dtype=torch.float), precision_matrix=precision_known)
# +
# Case 1
# Let us assume our prior is a Normal distribution with a good estimate of the mean
prior_mu = torch.tensor([19, 9], dtype=torch.float)
prior_precision = torch.tensor([[0.33, 0], [0, 0.33]], dtype=torch.float)
prior_dist = MultivariateNormal(prior_mu, precision_matrix=prior_precision)
torch.manual_seed(0)
#Number of samples is low.
n = 3
X = true_dist.sample((n,))
posterior_dist_low_n = inference_known_precision(X, prior_dist, precision_known)
mu_mle = X.mean(dim=0)
mu_map = posterior_dist_low_n.mean
# When n is low, the posterior is dominated by the prior. Thus, a good prior can help offset the lack of data.
# We can see this in the following case.
# With a small sample (n=3), the MLE estimate of mean is worse compared to the MAP estimate of mean
print(f"True mean: {true_dist.mean}")
print(f"MAP mean: {mu_map}")
print(f"MLE mean: {mu_mle}")
# +
# Case 2
prior_mu = torch.tensor([19, 9], dtype=torch.float)
prior_precision = torch.tensor([[0.33, 0], [0, 0.33]], dtype=torch.float)
prior_dist = MultivariateNormal(prior_mu, precision_matrix=prior_precision)
torch.manual_seed(0)
#Number of samples is high.
n = 1000
X = true_dist.sample((n,))
posterior_dist_high_n = inference_known_precision(X, prior_dist, precision_known)
mu_mle = X.mean(dim=0)
mu_map = posterior_dist_high_n.mean
# When n is high, the MLE tends to converge to the true distribution. The MAP also tends to converge to the MLE,
# and in turn converges to the true distribution
print(f"True mean: {true_dist.mean}")
print(f"MAP mean: {mu_map}")
print(f"MLE mean: {mu_mle}")
# -
# ### How to use the estimated mean parameter?
#
# We typically find $\vec \mu_{∗}$, the value of $\vec\mu$ that maximizes this posterior probability. In this particular case, the maxima of a Gaussian probability density occurs at the mean, hence, $\vec\mu_{∗}$ = $\vec\mu_{n}$.
#
# Given an arbitrary new data instance $x$, its probability of belonging to the class from which the training data has been sampled is $\mathcal{N}\left( \vec x; \vec\mu_{n}, \Lambda_{n} \right)$.
map_dist = MultivariateNormal(posterior_dist_high_n.mean, precision_matrix=precision_known)
print(f"MAP distribution mu: {map_dist.mean} precision:{map_dist.precision_matrix}")
| python/ch13/13.8-bayesian-inference-unknown-mean-multivariate.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1><center> Facial Emotion Recognition - Preprocessing </center></h1>
# <center> A project for the French Employment Agency </center>
# <center> Telecom ParisTech 2018-2019 </center>
# # I. Context
# The aim of this notebook is to explore facial emotion recognition techniques from a live webcam video stream.
#
# The data set used for training is the Kaggle FER2013 emotion recognition data set : https://www.kaggle.com/c/challenges-in-representation-learning-facial-expression-recognition-challenge/data
#
# The models explored include :
# - Manual filters
# - Deep Learning Architectures
# - DenseNet Inspired Architectures
#
# This model will be combined with voice emotion recongition as well as psychological traits extracted from text inputs, and should provide a benchmark and a deep analysis of both verbal and non-verbal insights for candidates seeking for a job and their performance during an interview.
# # II. General imports
# Versions used :
# + active=""
# Python : 3.6.5
# Tensorflow : 1.10.1
# Keras : 2.2.2
# Numpy : 1.15.4
# OpenCV : 4.0.0
# +
### General imports ###
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from time import time
from time import sleep
import re
import os
import argparse
from collections import OrderedDict
import matplotlib.animation as animation
### Image processing ###
from scipy.ndimage import zoom
from scipy.spatial import distance
import imutils
from scipy import ndimage
import cv2
import dlib
from __future__ import division
from imutils import face_utils
### CNN models ###
import keras
from keras.preprocessing.image import ImageDataGenerator, array_to_img, img_to_array, load_img
from keras.callbacks import TensorBoard
from keras.models import Sequential
from keras.layers.core import Dense, Dropout, Activation, Flatten
from keras.layers.convolutional import Conv2D, MaxPooling2D, SeparableConv2D
from keras.utils import np_utils
from keras.regularizers import l2#, activity_l2
from keras.optimizers import SGD, RMSprop
from keras.utils import to_categorical
from keras.layers.normalization import BatchNormalization
from keras import models
from keras.utils.vis_utils import plot_model
from keras.layers import Input, GlobalAveragePooling2D
from keras.models import Model
from tensorflow.keras import layers
### Build SVM models ###
from sklearn.preprocessing import OneHotEncoder
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score
from sklearn import svm
### Same trained models ###
import h5py
from keras.models import model_from_json
import pickle
# -
# # III. Import datas
path = '/Users/maelfabien/filrouge_pole_emploi/Video/'
local_path = '/Users/maelfabien/Desktop/LocalDB/Videos/'
# +
pd.options.mode.chained_assignment = None # default='warn' #to suppress SettingWithCopyWarning
#Reading the dataset
dataset = pd.read_csv(local_path + 'fer2013.csv')
#Obtaining train data where usage is "Training"
train = dataset[dataset["Usage"] == "Training"]
#Obtaining test data where usage is "PublicTest"
test = dataset[dataset["Usage"] == "PublicTest"]
#Converting " " separated pixel values to list
train['pixels'] = train['pixels'].apply(lambda image_px : np.fromstring(image_px, sep = ' '))
test['pixels'] = test['pixels'].apply(lambda image_px : np.fromstring(image_px, sep = ' '))
dataset['pixels'] = dataset['pixels'].apply(lambda image_px : np.fromstring(image_px, sep = ' '))
# -
dataset.head()
plt.figure(figsize=(12,6))
plt.hist(dataset['emotion'], bins=30)
plt.title("Distribution of the number of images per emotion")
plt.show()
train.shape
test.shape
# # IV. Create the data set
shape_x = 48
shape_y = 48
# +
X_train = train.iloc[:, 1].values
y_train = train.iloc[:, 0].values
X_test = test.iloc[:, 1].values
y_test = test.iloc[:, 0].values
X = dataset.iloc[:,1].values
y = dataset.iloc[:,0].values
#np.vstack stack arrays in sequence vertically (picking element row wise)
X_train = np.vstack(X_train)
X_test = np.vstack(X_test)
X = np.vstack(X)
#Reshape X_train, y_train,X_test,y_test in desired formats
X_train = np.reshape(X_train, (X_train.shape[0],48,48,1))
y_train = np.reshape(y_train, (y_train.shape[0],1))
X_test = np.reshape(X_test, (X_test.shape[0],48,48,1))
y_test = np.reshape(y_test, (y_test.shape[0],1))
X = np.reshape(X, (X.shape[0],48,48,1))
y = np.reshape(y, (y.shape[0],1))
print("Shape of X_train and y_train is " + str(X_train.shape) +" and " + str(y_train.shape) +" respectively.")
print("Shape of X_test and y_test is " + str(X_test.shape) +" and " + str(y_test.shape) +" respectively.")
# +
# Change to float datatype
X_train = X_train.astype('float32')
X_test = X_test.astype('float32')
X = X.astype('float32')
# Scale the data to lie between 0 to 1
X_train /= 255
X_test /= 255
X /= 255
# Change the labels from integer to categorical data
y_train = to_categorical(y_train)
y_test = to_categorical(y_test)
y = to_categorical(y)
# -
# # V. Define the number of classes
# +
# Find the unique numbers from the train labels
classes = np.unique(y_train)
nClasses = len(classes)
print('Total number of outputs : ', nClasses)
print('Output classes : ', classes)
# Find the shape of input images and create the variable input_shape
nRows,nCols,nDims = X_train.shape[1:]
input_shape = (nRows, nCols, nDims)
# -
#Defining labels
def get_label(argument):
labels = {0:'Angry', 1:'Disgust', 2:'Fear', 3:'Happy', 4:'Sad' , 5:'Surprise', 6:'Neutral'}
return(labels.get(argument, "Invalid emotion"))
# +
plt.figure(figsize=[10,5])
# Display the first image in training data
plt.subplot(121)
plt.imshow(np.squeeze(X_train[25,:,:], axis = 2), cmap='gray')
plt.title("Ground Truth : {}".format(get_label(int(y_train[0]))))
# Display the first image in testing data
plt.subplot(122)
plt.imshow(np.squeeze(X_test[26,:,:], axis = 2), cmap='gray')
plt.title("Ground Truth : {}".format(get_label(int(y_test[1500]))))
# -
# # VI. Save the dataframes
np.save(local_path + 'X_train', X_train)
np.save(local_path + 'X_test', X_test)
np.save(local_path + 'X', X)
np.save(local_path + 'y_train', y_train)
np.save(local_path + 'y_test', y_test)
np.save(local_path + 'y', y)
# # VII. Feature Importance
# +
path = '/Users/maelfabien/filrouge_pole_emploi/Video/'
local_path = '/Users/maelfabien/Desktop/LocalDB/Videos/'
X_train = np.load(local_path + "X_train.npy")
X_test = np.load(local_path + "X_test.npy")
y_train = np.load(local_path + "y_train.npy")
y_test = np.load(local_path + "y_test.npy")
shape_x = 48
shape_y = 48
nRows,nCols,nDims = X_train.shape[1:]
input_shape = (nRows, nCols, nDims)
classes = np.unique(y_train)
nClasses = len(classes)
# -
model = OneVsRestClassifier(LGBMClassifier(learning_rate = 0.1, num_leaves = 50, n_estimators=100, verbose=1))
model.fit(X_train.reshape(-1,48*48*1), y_train)
model = XGBClassifier()
model.fit(X_train[:10000], y_train[:10000])
gray = cv2.cvtColor(model.feature_importances_.reshape(shape_x, shape_y,3), cv2.COLOR_BGR2GRAY)
plt.figure(figsize=(12,8))
sns.heatmap(gray)
plt.show()
# # VIII. Sources
# - Visualization : https://github.com/JostineHo/mememoji/blob/master/data_visualization.ipynb
# - State of the art Architecture : https://github.com/amineHorseman/facial-expression-recognition-using-cnn
# - Eyes Tracking : https://www.pyimagesearch.com/2017/04/24/eye-blink-detection-opencv-python-dlib/
# - Face Alignment : https://www.pyimagesearch.com/2017/05/22/face-alignment-with-opencv-and-python/
# - C.Pramerdorfer, and M.Kampel.Facial Expression Recognition using Con-volutional Neural Networks: State of the Art. Computer Vision Lab, TU Wien. https://arxiv.org/pdf/1612.02903.pdf
# - A Brief Review of Facial Emotion Recognition Based
# on Visual Information : https://www.mdpi.com/1424-8220/18/2/401/pdf
# - Going deeper in facial expression recognition using deep neural networks : https://ieeexplore.ieee.org/document/7477450
# - Emotional Deep Alignment Network paper : https://arxiv.org/abs/1810.10529
# - Emotional Deep Alignment Network github : https://github.com/IvonaTau/emotionaldan
| 03-Video/.ipynb_checkpoints/01-Pre-Processing-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # **`pycoco`** calling **`CoCo`** - Extra Functions
# ____
# +
try:
from importlib import reload
except:
pass
# # %matplotlib inline
# %matplotlib notebook
from matplotlib import pyplot as plt
import os
import numpy as np
from astropy.table import Table
import pycoco as pcc
# -
# # Run Light Curve Fits
#
# +
snname = "SN2006aj"
sn = pcc.classes.SNClass(snname)
sn.load_phot(verbose=False)
sn.plot_lc(multiplot=False)
pcc.coco.test_LCfit(snname)
# -
# ## Model Choice
# ----
# You can pass a model to **`CoCo lcfit`**, this is possible through **`pycoco`** using the `model` arg. This will only take models that are defined in CoCo/src/models and match the filenames. So, currently the valid models are:
models = np.unique([i.split(".")[0] for i in os.listdir(os.path.join(pcc.defaults._default_coco_dir_path, "src/models"))])
models
model_name = "Karpenka12"
# model_name = "Karpenka12Afterglow"
# model_name = "Kessler10"
pcc.coco.run_LCfit(os.path.join(pcc.defaults._default_data_dir_path, "lc/"+snname+".dat"), model=model_name, verbose=True)
# +
sn = pcc.classes.SNClass(snname)
sn.load_phot(verbose=False)
sn.get_lcfit(os.path.join(pcc.defaults._default_recon_dir_path, snname + ".dat"))
sn.plot_lc(multiplot=False)
# -
pcc.coco.run_LCfit_fileinput(os.path.join(pcc.defaults._default_coco_dir_path, "examples/lc.list"))
print("Done.")
# ## Call Specfit
# ---
#
# ### individual speclists
pcc.coco.get_all_spec_lists()
# +
sn.load_list(path=os.path.join(pcc.defaults._default_list_dir_path, sn.name+".list"))
sn.load_spec()
# sn.load_mangledspec()
sn.load_reconspec()
sn.load_simspec()
# +
path_to_filter = os.path.join(pcc.defaults._default_filter_dir_path,"BessellB.dat")
BessellB = pcc.classes.FilterClass()
BessellB.read_filter_file(path_to_filter)
BessellB.calculate_AB_zp()
sim_spec_flux = []
sim_mjd_obs = []
for spec in sn.sim_spec:
print(spec)
flux = pcc.kcorr.calc_spectrum_filter_flux(filter_object=BessellB, spectrum_object=sn.sim_spec[spec])
mjd_obs = float(spec.split("_")[-1])
sim_spec_flux.append(flux)
sim_mjd_obs.append(mjd_obs)
# -
fig = plt.figure()
ax = fig.add_subplot(111)
ax.scatter(sim_mjd_obs, sim_spec_flux)
# pcc.coco.run_specfit('/Users/berto/Code/CoCo/lists/SN2013ge.list')
pcc.coco.run_specfit(SNObject=sn, overwrite=True)
# ## All Speclists
# ___
pcc.coco.specfit_all()
# # Specphase
phase_path = os.path.join(pcc.defaults._default_coco_dir_path, "examples/phase.list")
pcc.coco.run_specphase("BessellV", phase_path)
pcc.coco.run_specphase("BessellV", phase_path, model="Foo")
pcc.coco.run_specphase("BessellV", phase_path, model="Bazin09")
| notebooks/Tutorials/1b - pycoco calling CoCo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Time Point Algebras
# <NAME>, Ph.D.
import os
import qualreas as qr
path = os.path.join(os.getenv('PYPROJ'), 'qualreas')
# ## References
# 1. <NAME>, <b>"Maintaining knowledge about temporal intervals"</b>, Communications of the ACM 26(11) pp.832-843, Nov. 1983
# 1. <NAME>, <b>“The Logic of Time”</b>, D. Reidel Publishing Co., 1983
# 1. <NAME>, <b>"Intervals, Points, and Branching Time"</b>, In: <NAME>., <NAME>. (eds.) Proceedings of the TIME-94 International Workshop on Temporal Reasoning, University of Regina, Regina, SK, Canada, pp. 121–133, 1994
# ## The Structures of Linear and Branching Time
# The point (and interval) algebras of time, supported by qualreas, consider the structure of time to be either linear or branching as shown in the figures below.
# <img src="Images/global_structures_of_time.png" width="400" align="center"/>
# ### Point Structure
# According to [van Benthem, 1983] a <b><i>Point Structure</i></b> is an ordered pair, $(\mathcal{T},\prec)$, where $\mathcal{T}$ is a non-empty set and $\prec$ is a transitive binary relation on $\mathcal{T}$. Equality is denoted by $=$, and the converse of $\prec$ is $\succ$.
# ### Linear Point Structure
# A <b><i>Linear Point Structure</i></b> is a Point Structure, $(\mathcal{T},\prec)$, such that for any two points, $x,y \in \mathcal{T}$, <b>one and only one</b> of the following three relationships holds:
#
# <p>$(x \prec y) \vee (x = y) \vee (x \succ y)$</p>
#
# <b>Example:</b> If $\mathbb{R}$ is the set of real numbers, then both $(\mathbb{R},<)$ and $(\mathbb{R},\le)$ are Linear Point Structures.
# ### Branching Point Structure
# A <b><i>Branching Point Structure</i></b> is an ordered triple, $(\mathcal{T},\prec,\sim)$, where $(\mathcal{T},\prec)$ is a Point Structure and $\sim$ is an irreflexive, symmetric binary relation on $\mathcal{T}$, called <b><i>incomparable</i></b>, such that for any $x,y \in \mathcal{T}$, <b>one and only one</b> of the following four relationships holds:
#
# <p>$(x \prec y) \vee (x = y) \vee (x \succ y) \vee (x \sim y)$</p>
#
# Basically, if $x$ and $y$ are on two different branches, then $x \sim y$.
# #### Binary-Branching vs. Poly-Branching
# There is a subtle difference in the composition of the incomparable relation with itself ($\sim;\sim$) depending on whether only two branches are allowed at a branch point (binary-branching) or more than two branches are allowed (poly-branching).
#
# * binary-branching: $(\sim ; \sim) = \{\prec, =, \succ\}$
# * poly-branching: $(\sim ; \sim) = \{\prec, =, \succ, \sim\}$
# ### Right-Branching Point Structure
# A <b>Right-Branching Point Structure</b> is a Branching Point Structure that has the property of <b><i>Left Linearity</i></b>:
#
# <p>$x,y,z \in \mathcal{T}$ and $(x < z) \wedge (y < z) \implies (x < y) \vee (x = y) \vee (x > y)$</p>
# <img src="Images/left_linearity_in_right_branching_time.png" width="300" align="center"/>
# ### Left-Branching Point Structure
# A <b>Left-Branching Point Structure</b> is a Branching Point Structure that has the property of <b><i>Right Linearity</i></b>:
#
# <p>$x,y,z \in \mathcal{T}$ and $(x > z) \wedge (y > z) \implies (x < y) \vee (x = y) \vee (x > y)$</p>
# <b>NOTE:</b> In the branching point algebras defined in qualreas, we distinguish between the right & left incomparable ($\sim$) relations by putting an "r" or an "l" in front of $\sim$ (i.e., "r\~", "l\~"). This is not really necessary, since right and left branching point structures cannot be mixed together, but this is how things got started in qualreas, so it remains that way, for now. In the discussion, below, the left and right branching <i>incomparable</i> relations are denoted by $\underset{L}{\sim}$ and $\underset{R}{\sim}$, respectively.
# ## Linear Point Algebra
# This algebra is based on the Linear Point Structure, $(\mathbb{R},<)$, and is used to derive Allen's algebra of proper time intervals [Allen, 1983]--known in qualreas as the "Linear Interval Algebra". (See the Jupyter Notebook, <i>"Notebooks/derive_allens_algebra.ipynb"</i>)
#
# An extension to Allen's algebra, the "Extended Linear Interval Algebra" [Reich, 1994], integrates proper time intervals with time points by using the Linear Point Structure, $(\mathbb{R},\le)$. (See the Jupyter Notebook, <i>"Notebooks/derive_extended_interval_algebra.ipynb"</i>)
pt_alg = qr.Algebra(os.path.join(path, "Algebras/Linear_Point_Algebra.json"))
pt_alg.summary()
qr.print_point_algebra_composition_table(pt_alg)
# ## Right-Branching Point Algebra
# An extension to Allen's algebra, the "Right-Branching Interval Algebra" [Reich, 1994], integrates proper time intervals with time points in a poly-branching, right-branching time structure, by using the Right-Branching Point Structure, $(\mathbb{R},\le, \underset{R}{\sim})$, below. (See the Jupyter Notebook, <i>"Notebooks/derive_right_branching_interval_algebra.ipynb"</i>)
rb_pt_alg = qr.Algebra(os.path.join(path, "Algebras/Right_Branching_Point_Algebra.json"))
rb_pt_alg.summary()
qr.print_point_algebra_composition_table(rb_pt_alg)
# ## Left-Branching Point Algebra
# An extension to Allen's algebra, the "Left-Branching Interval Algebra" [Reich, 1994], integrates proper time intervals with time points in a poly-branching, left-branching time structure, by using the Left-Branching Point Structure, $(\mathbb{R},\le, \underset{L}{\sim})$, below. (See the Jupyter Notebook, <i>"Notebooks/derive_right_branching_interval_algebra.ipynb"</i>)
lb_pt_alg = qr.Algebra(os.path.join(path, "Algebras/Left_Branching_Point_Algebra.json"))
lb_pt_alg.summary()
qr.print_point_algebra_composition_table(lb_pt_alg)
# ## Right-Binary-Branching Point Algebra
# The "Right-Binary-Branching Interval Algebra", is Allen's algebra of proper intervals, situated in a binary-branching, right-branching time structure, and is derived using the Right-Binary-Branching Point Structure, $(\mathbb{R},\le, \underset{L}{\sim})$, below. (See the Jupyter Notebook, <i>"Notebooks/derive_right_binary_branching_interval_algebra.ipynb"</i>)
rbb_pt_alg = qr.Algebra(os.path.join(path, "Algebras/Right_Binary_Branching_Point_Algebra.json"))
rbb_pt_alg.summary()
qr.print_point_algebra_composition_table(rbb_pt_alg)
# ## Left-Binary-Branching Point Algebra
# The "Left-Binary-Branching Interval Algebra", is Allen's algebra of proper intervals, situated in a binary-branching, left-branching time structure, and is derived using the Left-Binary-Branching Point Structure, $(\mathbb{R},\le, \underset{L}{\sim})$, below.
# (See the Jupyter Notebook, <i>"Notebooks/derive_left_binary_branching_interval_algebra.ipynb"</i>)
lbb_pt_alg = qr.Algebra(os.path.join(path, "Algebras/Left_Binary_Branching_Point_Algebra.json"))
lbb_pt_alg.summary()
qr.print_point_algebra_composition_table(lbb_pt_alg)
| Notebooks/time_point_algebras.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/kusumikakd/Data_Analysis-by-python/blob/master/Brain_Tumor_Classification_from_MRI.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="5DI_RmniQog3" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 72} outputId="50022db3-bc09-4405-c236-1c3bcebe0a7a"
import numpy as np
import seaborn as sns
import pandas as pd
import matplotlib.pyplot as plt
# + id="MpVMx80n2f4u" colab_type="code" colab={}
#For reading Image files
import cv2
from PIL import Image
# to read from directory
import os
from os import listdir
# + id="DayDyCWH2vQ_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="c653233d-93e1-4714-cc3a-83050b3ba66e"
from random import shuffle
from collections import Counter
import tensorflow as tf
from keras.models import Sequential
from keras.layers import Conv2D, MaxPooling2D
from keras.layers import Activation, Dropout, Flatten, Dense
from keras.utils.np_utils import to_categorical
# + id="82Rf5ppBSDXx" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="6b6d640d-1cb0-4d87-dc99-b4892ec29149"
os.listdir('/content/drive/My Drive/brain_tumor_dataset-20200604T045108Z-001')
# + id="BmnM4Wv8SP5p" colab_type="code" colab={}
no_image=Image.open('/content/drive/My Drive/brain_tumor_dataset-20200604T045108Z-001/no/1 no.jpeg')
yes_image = Image.open('/content/drive/My Drive/brain_tumor_dataset-20200604T045108Z-001/yes/Y1.jpg')
# + id="PXjqubGuTADZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 610} outputId="88404a81-5ab6-49da-efab-ecaaccf4884d"
# Let's plot these images
#Let's plot these x-ray images
fig = plt.figure(figsize=(20,10))
# 1,2,1 ==> 1 row, 2 columns, 1st Image
a1 = fig.add_subplot(1, 2, 1)
img_plot = plt.imshow(no_image , cmap= 'gray')
a1.set_title ('Normal MRI', fontsize = 15)
# 1,2,2 ==> 1 row, 2 columns, 2nd Image
a1 = fig.add_subplot(1, 2, 2)
img_plot = plt.imshow(yes_image , cmap= 'gray')
a1.set_title ('Brain Tumor MRI', fontsize = 15)
# + id="GL15nnVSTZYO" colab_type="code" colab={}
yes=os.listdir('/content/drive/My Drive/brain_tumor_dataset-20200604T045108Z-001/yes')
no= os.listdir('/content/drive/My Drive/brain_tumor_dataset-20200604T045108Z-001/no')
# + id="abNBJDhJUXrj" colab_type="code" colab={}
data=np.concatenate([yes, no])
# + id="d07K0JMFUgwC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 158} outputId="8f0a16d7-62fa-4e22-f3c8-28fe7d51eef6"
target_x=np.full(len(yes),1)
target_y = np.full(len(no), 0)
target_x
# + id="LzGshSZFU19c" colab_type="code" colab={}
data_target=np.concatenate([target_x, target_y])
# + id="ga7oqDr2WRVW" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 230} outputId="cd1d6816-dc02-4972-8c13-9442052c0834"
data_target
# + id="tzvJJzrRWTv4" colab_type="code" colab={}
x_data=[]
for file in yes:
img= cv2.imread('/content/drive/My Drive/brain_tumor_dataset-20200604T045108Z-001/yes/'+file)
mri= cv2.resize(img, (32,32))
(b, g, r)= cv2.split(mri)
img=cv2.merge([r,g,b])
x_data.append(img)
# + id="Wff8kiujYCBN" colab_type="code" colab={}
img= cv2.imread('/content/drive/My Drive/brain_tumor_dataset-20200604T045108Z-001/yes/Y1.jpg')
# + id="eTag-qvKYVPZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 887} outputId="8ca2da14-d2a6-42d2-d771-b5ccb12625e3"
img
# + id="F9KTotI5YZNg" colab_type="code" colab={}
for file in no:
img= cv2.imread('/content/drive/My Drive/brain_tumor_dataset-20200604T045108Z-001/no/'+file)
mri= cv2.resize(img, (32,32))
(b, g, r)= cv2.split(mri)
img=cv2.merge([r,g,b])
x_data.append(img)
# + id="g3gf6s9fY7RW" colab_type="code" colab={}
X= np.squeeze(x_data)
# + id="RvBjyu__bU3w" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="df3ee25f-36e7-4dbc-fa1d-b5e121fd9d2c"
X.shape
# + id="iKz621UubXE4" colab_type="code" colab={}
X=X.astype('float32')
X/=255
# + id="XE1inXIJbjsK" colab_type="code" colab={}
from sklearn.model_selection import train_test_split
x_train, x_test, y_train, y_test = train_test_split(X, data_target, test_size=0.1, random_state = 1)
# + id="MHoasTTOcNhP" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="c6693a80-b287-4983-b310-c42a93f8987c"
x_train.shape
# + id="a9MHksgocYQQ" colab_type="code" colab={}
x_train2, x_val, y_train2, y_val =train_test_split(X, data_target, test_size=0.25, random_state = 1)
# + id="KNsXV59FcmAH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="eecab57b-4b02-46dc-eb61-d0184b9daa83"
x_train2.shape
# + id="07aa62QvhAdG" colab_type="code" colab={}
def build_model():
# Input Layer
model = Sequential()
model.add(Conv2D(filters=16, kernel_size=9, activation='relu', padding='same', input_shape = (32, 32, 3)))
model.add(MaxPooling2D(pool_size=(2)))
model.add(Dropout(0.45))
# ConvNets - II
model.add(Conv2D(filters=16,kernel_size=9,padding='same', activation='relu'))
model.add(MaxPooling2D(pool_size=2))
model.add(Dropout(0.25))
# ConvNets - III
model.add(Conv2D(filters=36, kernel_size=9, padding='same', activation='relu'))
model.add(MaxPooling2D(pool_size=2))
model.add(Dropout(0.25))
# Faltten Matrix
model.add(Flatten())
# FC Layer
model.add(Dense(512, activation='relu'))
model.add(Dropout(0.15))
# Output Layer
model.add(Dense(1, activation='sigmoid'))
# Take a look at the model summary
model.compile(optimizer = 'adam', loss = 'binary_crossentropy', metrics = ['accuracy'])
return model
# + id="Qj4pmccmhBV-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 621} outputId="400c04b3-a872-4168-9769-e9242beffbb9"
model = build_model()
model.summary()
# + id="91XeC7jthGVk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="c729082e-d48f-4bd3-d8ae-3357d3f855b9"
history = model.fit(x_train2,
y_train2,
batch_size=128,
epochs=150,
validation_data=(x_val, y_val))
# + id="MxBoRcIvhMf3" colab_type="code" colab={}
pd.DataFrame(history.history).plot()
# + id="uUALyv19tIvE" colab_type="code" colab={}
score= model.evaluate(x_test, y_test)
# + id="uGiDNeOyt43F" colab_type="code" colab={}
score[1]
# + id="2ceBBp9kt_nU" colab_type="code" colab={}
prediction= model.predict(x_test)
# + id="05EB2PNxuGoh" colab_type="code" colab={}
from sklearn.metrics import confusion_matrix
predictions = np.round(prediction)
from sklearn.metrics import confusion_matrix
confusion_mtx = confusion_matrix(y_test, predictions)
confusion_mtx
# + id="6DZ10qNJvm9I" colab_type="code" colab={}
labels =["Yes","No"]
y_hat = model.predict(x_test)
# + id="VgIx6Pdav-n5" colab_type="code" colab={}
# Plot a random sample of 10 test images, their predicted labels and ground truth
figure = plt.figure(figsize=(20, 8))
for i, index in enumerate(np.random.choice(x_test.shape[0], size=15, replace=False)):
ax = figure.add_subplot(3, 5, i + 1, xticks=[], yticks=[])
# Display each image
ax.imshow(np.squeeze(x_test[index]))
predict_index = np.argmax(y_hat[index])
true_index = np.argmax(y_test[index])
# Set the title for each image
ax.set_title("{} ({})".format(labels[predict_index],
labels[true_index]),
color=("green" if predict_index == true_index else "red"))
plt.show()
# + id="UNFCfJAfwDDF" colab_type="code" colab={}
| Brain_Tumor_Classification_from_MRI.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Example: PCA applied to MNIST
#
# This notebook shows how PCA can be applied to a non-trivial data set to reduce its dimensionality.
# PCA (and its variations, like Incremental PCA) is a useful tool to reduce the dimensionality of your feature space to boost model training performance.
# In this example, we will examine how vanilla PCA affects the MNIST data set.
# +
# %matplotlib inline
import matplotlib
import matplotlib.pyplot as plt
matplotlib.rcParams.update({'font.size': 18})
import numpy as np
from sklearn.decomposition import PCA
from sklearn.model_selection import train_test_split
# Load my smaller MNIST data set...
X = np.load('data/mnist/mnist_data.pkl')
y = np.load('data/mnist/mnist_target.pkl')
# Create the training and test sets
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.25, random_state=42)
# -
# A single instance of the input has **784 features**!
sample_idx = 0
sample_digit = X_train[0,:]
print("Dimension = {0}".format(sample_digit.shape))
# Here is what that 28x28 = 784 dimension instance looks like:
digit_image = sample_digit.reshape(28,28)
plt.imshow(digit_image, cmap=matplotlib.cm.binary, interpolation='nearest')
plt.axis('on')
plt.show()
# Let us assume that this feature space is too large for our chose classification algorithm.
# We could map the data set into a 196 dimensional space...
pca_1 = PCA(n_components=196, random_state=42)
X_train_pca1 = pca_1.fit_transform(X_train)
sample_digit_pca1 = X_train_pca1[sample_idx]
print("Dimension = {0}".format(sample_digit_pca1.shape))
sample_digit_pca1_img = sample_digit_pca1.reshape(14,14)
plt.imshow(sample_digit_pca1_img, cmap=matplotlib.cm.binary, interpolation='nearest')
plt.axis('on')
plt.show()
# Yikes! I thought we were getting a 14x14 image!
# Hmmm...wait a tick. We **compressed** the image into some 192 dimensional space that does not directly map to the originally intended pixel space!
#
# We should transform them back to see the **loss** due to this compression.
X_train_recovered = pca_1.inverse_transform(X_train_pca1)
X_train_recovered.shape
sample_digit_pca1_restored = X_train_recovered[0,:]
print("Dimension = {0}".format(sample_digit_pca1_restored.shape))
# As humans, we see that reducing the input features by a factor of 4 still makes the number legible.
# To our training algorithm, this reduction will result in faster training as the features are mapped to 196 dimensions with **highest variance**.
# +
sample_digit_pca1_res_img = sample_digit_pca1_restored.reshape(28,28)
plt.imshow(sample_digit_pca1_res_img, cmap=matplotlib.cm.binary, interpolation='nearest')
plt.axis('off')
plt.title('Restored from 196D Space')
plt.show()
plt.imshow(digit_image, cmap=matplotlib.cm.binary, interpolation='nearest')
plt.axis('off')
plt.title('Original')
plt.show()
# -
# ## Pushing the limits
# Now let's get crazy. What if we could smash these numbers into a 3D space?
pca_2 = PCA(n_components=3, random_state=42)
X_train_pca2 = pca_2.fit_transform(X_train)
# As you will see, this compression has no intuition to us.
X_train_pca2[0,:]
X_train_recovered_2 = pca_2.inverse_transform(X_train_pca2)
sample_digit_pca2_restored = X_train_recovered_2[sample_idx,:]
sample_digit_pca2_res_img = sample_digit_pca2_restored.reshape(28,28)
plt.imshow(sample_digit_pca2_res_img, cmap=matplotlib.cm.binary, interpolation='nearest')
plt.axis('off')
plt.title('Restored from 3D Space')
plt.show()
# In this case, we see *some* visual cues that remain from the original instance; however, it is likely too lossy to be of any use!
| modules/14_stats_and_modeling/modeling/AI_and_ML_Intro_MITRE_with_python/Demo-PCA-MNIST.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Only for format the raw data.
import pandas as pd
# #### Load and format dataset Titanic survived dataset.
# laod original trainning dataset.
train_data = pd.read_csv(
"winequality-white.csv",
sep=r'\s*,\s*',
quotechar="'",
engine='python',
index_col=False,
na_values="?")
# Counting...
Target = 'label'
Labels = train_data[Target].unique()
counts = train_data[Target].value_counts()
print(counts)
# Fusion the datasets using only selected features.
final = train_data.dropna()
cols = final.columns.tolist()
temp = cols[1:]
temp.append(cols[0])
final = final[temp]
# Shuffling the data
final = final.sample(frac=1)
# Drop missing data
final = final.dropna()
# Convert to categorical features
final = final.astype('category')
# How much samples remaing.
Target = 'label'
Labels = final[Target].unique()
counts = final[Target].value_counts()
print(counts)
frac = 0.6
final.reset_index(inplace=True)
#train = final.loc[1:int(round(len(final.index)*frac)),:]
#test = final.loc[int(round(len(final.index)*frac)):,:]
train = final.loc[1:500,:]
test = final.loc[501:1001,:]
train.reset_index(inplace=True)
test.reset_index(inplace=True)
train = train.drop(train.columns[0:2], axis=1)
test = test.drop(test.columns[0:2], axis=1)
train = train.drop(train.columns[0:2], axis=1)
test = test.drop(test.columns[0:2], axis=1)
# +
# Counting...
Labels = train[Target].unique()
counts = train[Target].value_counts()
print(counts)
Labels = test[Target].unique()
counts = test[Target].value_counts()
print(counts)
# -
# Save the dataset in CSV.
train.to_csv('dataset.training.csv.mnist', sep=',', encoding='ascii', decimal='.', index=False, header=False)
test.to_csv('dataset.test.csv.mnist', sep=',', encoding='ascii', decimal='.', index=False, header=False)
train.head()
| RFCLP/.ipynb_checkpoints/Format_raw_data_wine-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#libraries for multi-variate encoder-decoder lstm
from pandas_datareader import data
import matplotlib.pyplot as plt
from math import sqrt
import pandas as pd
from pandas import read_csv
import datetime as dt
import urllib.request, json
import os
import numpy as np
from numpy import newaxis, split, array
import matplotlib.pyplot as plt
import tensorflow as tf
from sklearn.preprocessing import MinMaxScaler, StandardScaler, Normalizer
from sklearn.metrics import mean_squared_error
from sklearn import preprocessing
import time
import warnings
from keras.layers.core import Dense, Activation, Dropout
from keras.layers import Convolution1D, MaxPooling1D, Flatten, Embedding
from keras.layers import Conv1D, GlobalMaxPooling1D
from keras.layers.recurrent import LSTM
from keras.models import Sequential
from datetime import datetime
from keras.layers import Dense, Flatten, LSTM, RepeatVector, TimeDistributed
# +
#Load data
# +
#Asset data
def GetData(fileName):
return pd.read_csv(fileName, header=0,usecols=['Date','Adj Close'], parse_dates=True, index_col='Date')
# -
data_ivv = GetData('Asset_Dataset/IVV_daily.csv')
data_ivv.columns = ['IVV Adjusted Close']
# %store data_ivv
#normalize ivv
scaler = StandardScaler()
data_ivv_norm = scaler.fit_transform(data_ivv.values.reshape(-1, 1))
data_ivv_norm
#change to df for merging
data_ivv_norm_df = pd.DataFrame(data_ivv_norm)
#market and econ data
# %store -r scaled_macro_market
scaled_macro_market
scaled_macro_market_df = pd.DataFrame(scaled_macro_market)
merged_data = pd.concat([data_ivv_norm_df,scaled_macro_market_df],axis=1)
merged_data = merged_data.dropna()
merged_data.columns = ['IVV adj close','Real GDP','CPI All','Fed Rate','Unemployment rate','Personal Savings Rate','VIX']
merged_data
# +
##LSTM STARTS HERE##
# -
len(merged_data)
input_data = merged_data.values
# +
#try using plotly / seaborn / iplot
#to plot IVV & All parameters
#
# +
#The multiple input features make this an multivariate regression problem
lookback = 50
test_size = int(0.3*len(merged_data))
X=[]
y=[]
for i in range(len(merged_data)-lookback-1):
t=[]
for j in range(0,lookback):
t.append(input_data[[(i+j)],:])
X.append(t)
y.append(input_data[i+lookback,1])
# +
#transform input data into LSTM expected structure
#first convery X into array then reshape()
X,y = np.array(X), np.array(y)
X_test = X[:test_size+lookback]
X = X.reshape(X.shape[0]*350,lookback,2)
X_test = X_test.reshape(X_test.shape[0]*500, lookback , 2)
print(X.shape)
print(X_test.shape)
# -
model = Sequential()
model.add(LSTM(units=30, return_sequences= True, input_shape=(X.shape[1],2)))
model.add(LSTM(units=30, return_sequences=True))
model.add(LSTM(units=30))
model.add(Dense(units=1))
model.summary()
model.compile(optimizer='adam', loss='mean_squared_error')
model.fit(X, y, epochs=200, batch_size=32)
predicted_value= model.predict(X_test)
plt.plot(predicted_value, color= 'red')
plt.plot(input_data[lookback:test_size+(2*lookback),1], color='green')
plt.title("Opening price of stocks sold")
plt.xlabel("Time (latest-> oldest)")
plt.ylabel("Stock Opening Price")
plt.show()
| ! Dissertation/*LSTM/LSTM Prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Ensuring reproducibility and transparency can be so easy when using the right tooling from the start. This is a tutorial to showcase the use of [conda](https://conda.io) and [GitHub Actions](https://docs.github.com/en/actions). We use both to ensure easy reproducibility of our example notebook.
# ```console
# foo@bar:~$ git clone https://github.com/HumanCapitalAnalysis/student-project-template
# foo@bar:~$ cd student-project-template
# foo@bar:~$ conda env create -f environment.yml
# foo@bar:~$ conda activate student_project
# foo@bar:~$ jupyter nbconvert --execute student_project.ipynb
# ```
# ## conda - package management
# ! conda
# ! conda --version
# We can create a virtual environment for our student project and install some basic packages right from the beginning.
# ! conda env remove --name student_project_template
# ! conda create -y --name student_project_template numpy pandas
# Now we can have a look at all environments.
# ! conda env list
# We can now switch to the terminal window or the Anaconda prompt to activate the environment.
# ```console
# foo@bar:~$ conda acivate student_project_template
# foo@bar:~$ which python
# foo@bar:~$ conda list
# ```
# We are free to add / and remove packages form the environment.
# ```console
# foo@bar:~$ conda install scipy
# foo@bar:~$ conda remove pandas
# ```
# Only the installed packages will be available.
# ```console
# foo@bar:~$ python -c "import scipy"
# foo@bar:~$ python -c "import pandas"
# ```
# Returning to the notebook we can automate the process of environment generation using **environment.yml** files.
# ! cat environment_tutorial.yml
# ! conda env remove --name student_project_template
# ! conda env create -f environment_tutorial.yml
# ## GitHub Actions - continuous integration
#
# Now that we have automated the installation of all required software, we can move it to the cloud and execute our analysis there to ensure that there are no local dependencies that we are missing.
# I have linked your projects to **GitHub Actions** already, so all your commits are monitored now and since you have a **.github/workflows/ci.yml** in your repo a build will be triggered based on the instructions in there.
# ! cat .github/workflows/ci.yml
# Now we can run the notebooks on the CI server to ensure full reproducibility as (if passing) this means that all required files are available on **GitHub** and the whole software environment is also fully specified.
# Let's trigger a build and see the magic in action.
# 
# When all is working, don't forget to proudly add your [badge](https://docs.github.com/en/actions/guides/about-continuous-integration#status-badges-for-workflow-runs) to the **README.md** file.
| tutorial_conda_actions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] deletable=true editable=true
# # Examples of Data Distributions
# + [markdown] deletable=true editable=true
# ## Uniform Distribution
# + deletable=true editable=true
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
values = np.random.uniform(-10.0, 10.0, 100000)
plt.hist(values, 50)
plt.show()
# + [markdown] deletable=true editable=true
# ## Normal / Gaussian
# + [markdown] deletable=true editable=true
# Visualize the probability density function:
# + deletable=true editable=true
from scipy.stats import norm
import matplotlib.pyplot as plt
x = np.arange(-3, 3, 0.001)
plt.plot(x, norm.pdf(x))
# + [markdown] deletable=true editable=true
# Generate some random numbers with a normal distribution. "mu" is the desired mean, "sigma" is the standard deviation:
# + deletable=true editable=true
import numpy as np
import matplotlib.pyplot as plt
mu = 5.0
sigma = 2.0
values = np.random.normal(mu, sigma, 10000)
plt.hist(values, 50)
plt.show()
# + [markdown] deletable=true editable=true
# ## Exponential PDF / "Power Law"
# + deletable=true editable=true
from scipy.stats import expon
import matplotlib.pyplot as plt
x = np.arange(0, 10, 0.001)
plt.plot(x, expon.pdf(x))
# + [markdown] deletable=true editable=true
# ## Binomial Probability Mass Function
# + deletable=true editable=true
from scipy.stats import binom
import matplotlib.pyplot as plt
n, p = 10, 0.5
x = np.arange(0, 10, 0.001)
plt.plot(x, binom.pmf(x, n, p))
# + [markdown] deletable=true editable=true
# ## Poisson Probability Mass Function
# + [markdown] deletable=true editable=true
# Example: My website gets on average 500 visits per day. What's the odds of getting 550?
# + deletable=true editable=true
from scipy.stats import poisson
import matplotlib.pyplot as plt
mu = 500
x = np.arange(400, 600, 0.5)
plt.plot(x, poisson.pmf(x, mu))
# + [markdown] deletable=true editable=true
# ## Pop Quiz!
# + [markdown] deletable=true editable=true
# What's the equivalent of a probability distribution function when using discrete instead of continuous data?
# + deletable=true editable=true
| Distributions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Generalized linear models
# ## <NAME>, <NAME>, <NAME>, <NAME>
# > One of the central problems in systems neuroscience is that of characterizing the functional relationship between sensory stimuli and neural spike responses. Investigators call this the neural coding problem, because the spike trains of neurons can be considered a code by which the brain represents information about the state of the external world. One approach to understanding this code is to build mathematical models of the mapping between stimuli and spike responses; the code can then be interpreted by using the model to predict the neural response to a stimulus, or to decode the stimulus that gave rise to a particular response. [(Pillow, 2007)](http://pillowlab.princeton.edu/pubs/Pillow_BBchap07.pdf)
#
# Here, we will build probabilistic models for the response of a single neuron, starting from a simple model, that we will then extend. Conditional on a stimulus $x$ and model parameters $\theta$ we will model the probability of a neural reponse $y$, i.e., $p(y|x, \theta)$. Our central goal will be to find model parameters $\theta$ such that the $p(y|x,\theta)$ is a good fit to a dataset of stimulus-response pairs we observed, $\mathcal{D} = \{ (x_k,y_k) \}_{k=1}^K$.
#
#
# ### Goals of these exercises
#
# Central to inferring the best fitting parameters will be the likelihood function of the model. The simplest method goes by simply maximizing the likelihood using its gradient with respect to the model parameters (a technique called maximum likelihood estimation, MLE). You will learn to incorporate prior knowledge on the parameters, which leads to a method called maximum a posteriori (MAP). Finally, you will learn automatic differentiation (AD) which --- as the name suggests --- provides a automatic way to calcuate gradients of an objective function (here: the likelihood of parameters given the data). AD is a central ingredient to machine learning methods that are becoming increasingly popular.
#
# ### Assumptions and notation
#
# Throughout this tutorial, we will adopt the following conventions:
#
# - $T$ is the number of time bins within one trial; $t$ always specifies a time bin;
# - $K$ is the number of trials in the experiment; $k$ always identifies a trial;
# - to make the notation lighter, we will sometimes drop the subscript $k$;
# - $\hat{\pi}(\cdot)$ indicates an unnormalized probability, and $\pi(\cdot)$ the same probability normalize to integrate to 1;
# - $\mathcal{L}(\boldsymbol{\theta}) = p(\mathbf{y}\, |\, \boldsymbol{\theta})$ is the likelihood of the vector of parameters $\boldsymbol{\theta}$ for the (fixed) data $\mathbf{y}$.
#
# For all models we consider, we assume that time is discretized in bins of size $\Delta$. Given $z_t$, the instantaneous *input rate* of a neuron at time $\Delta \cdot t$, the spike counts $y_t$ are assumed to be independent, and distributed according to
#
# $\begin{equation}
# y_t \sim \mathrm{Poisson}\big(\eta(z_t)\big)
# \end{equation}$
#
# where $\eta(\cdot)$ is corresponding canonical link function (here, we will always use $\eta(\cdot) = \exp(\cdot)$ for Poisson). We further assume that there is a linear dependence between $z_i$ and a set of external covariates $\mathbf{x}_t$ at time $t$, i.e. $z_t = \boldsymbol{\theta}^\top \mathbf{x}$, and $\boldsymbol{\theta}$ is a vector of parameters which fully characterizes the neuron.
#
# Experiments are composed of $K$ trials, each subdivided into $T$ bins.
#
# Note, and in contrast to the lectures, we assume that the rate $\mu_t$ is already 'per bin size', i.e. the expected number of spikes is $\mu$ (and not $\mu\Delta$, as we had in lectures).
#
# For a Poisson neuron, the probability of producing $n$ spikes in an interval of size $\Delta$ is given by
#
# $\begin{align}
# P(y_t=n| \mu)= \frac{\mu^n e^{-\mu} }{n!}
# \end{align}$
#
#
# ## Exercise 1 (from lectures)
#
# Assume that you have spike counts $n_1$ to $n_K$ from $K$ trials, calculate the maximum likelihood estimate (MLE) of $\mu$.
#
#
# ## LNP model
#
# The stimulus $\mathbf{u}_t$ is a white noise sequence, and the input rate is:
#
# $\begin{equation}
# z_t = \mathbf{\beta}^\top \mathbf{u}_{t-\delta{}+1:t} + b = \boldsymbol{\theta}^\top \mathbf{x}
# \end{equation}$,
#
# i.e. $z_t$ is the result of a linear filter $\beta$ applied to the recent stimulus history $\mathbf{u}_{t-\delta{}+1:t}$, plus some offset $b$. This results in a vector of covariates at time $t$
# $\mathbf{x}_{kt} = \big[1, \mathbf{u}_{kt-\delta{}+1},\ldots, \mathbf{u}_{kt} \big]^\top$ for temporal filter length $\delta \in \mathbb{N}$. Note that we can deal with any form of input in the second and third column of $\mathbf{x}_{kt}$, not just white noise.
#
# The vector of parameters is $\boldsymbol{\theta} = \left[b, \beta^\top \right]^\top$.
#
#
#
# ### Simulating data from the model
#
# Next, we will want to generate data using this model. Execute the following code cell, which will load some functions you will need throughout the session.
# +
import numpy as np
# %run -i helpers.ipynb
# -
# The following cell generates a matrix $\mathbf{x}$ as specified above.
# +
binsize = 0.001 # seconds
T = 10000 # 1s trials
K = 10 # number of trials
nbins = T*K
delta = 10 # length of temporal filter
# stimulus
U = np.random.normal(size=nbins)
def toyDesignMatrix(U=U, T=T, K=K):
nbins = T*K
X = np.zeros((delta+1, nbins))
X[0,:] = 1. # bias
if delta > 0:
X[1, :] = U # instantaneous input
for i in range(1,delta):
X[i+1, i+1:] = U[:-(i+1)]
return X
X = toyDesignMatrix()
# -
# Next, we define $\mathbf{\theta}$.
# ground-truth vector of parameters
b = -6 # controls the offset and hence overall firing rate
beta = np.cos( np.linspace(0, PI, delta))
theta = np.hstack([b, beta])
# Given `X` and `theta`, we want to generate sample spike trains. In the following cell, we do so by just using
ys = []
for k in range(10):
y, fr = toyModel(X, theta) # spike train, firing rate
ys.append(y)
# ... plotting spike rasters and PSTH:
plt.subplot(311)
plt.plot(U[:200])
plt.subplot(312)
plt.imshow(np.asarray(ys)[:,:200], aspect=4, interpolation='None');
plt.subplot(313)
plt.plot(np.asarray(ys)[:, :200].mean(axis=0) / binsize); # PSTH
plt.plot(fr[:200], linewidth=2); # firing rate
#
# ### Optional: Try implementing the model yourself
#
# (Optional): Above, you used an implementation of the model we provided. You can try implementing the model as stated above yourself. To do so, complete the following function template
def toyModelExercise(X, theta):
# TODO: given stimulus and theta, return spikes and firing rate
return y, fr
# To check whether this model is correct, reproduce the PSTHs for both models and compare.
#
#
# ### MLE inference
#
# #### Likelihood ####
#
# The likelihood defines a model that connects model parameters to the observed data:
#
# $\begin{align}
# \log \mathcal{L}(\boldsymbol{\theta}) &= \log p(\mathbf{y} | \boldsymbol{\theta}) = \log \bigg[ \prod_{k=1}^K \prod_{t=1}^T p(y_{kt} | b, \beta_1, \beta_2) \bigg] \\ &= \sum_{k=1}^K \sum_{t=1}^T \log p(y_{tk} | b, \beta_1, \beta_2) = \sum_{k=1}^K \sum_{t=1}^T \big[ z_{tk} y_{tk} - \mathrm{e}^{z_{tk}} \big],
# \end{align}$
#
# where as above $z_{tk} = \theta^\top \mathbf{x}_{tk} = b + \beta^\top \mathbf{u}_{tk-\delta{}+1:tk}$.
#
# Large $\mathcal{L}$ for a given set of parameters $\theta$ indicates that the data is likely under that parameter set. We can iteratively find more likely parameters, starting from some initial guess $\theta_0$, by gradient ascent on $\mathcal{L}$.
#
# For this model, the likelihood function has a unique maximum.
#
#
# #### Gradients ####
#
# **Exercise 1:** Using pen and paper, derive the gradient of the $\log \mathcal{L}$ with respect to $\mathbf{\theta}$.
#
# ### MLE parameter inference
#
# We will now want to the use gradient you just derived to do parameter inference. For that, we will need to implement the functions `ll` and `dll` (the log-likelihood function and its derivative).
# **Exercise 2.1: ** Implement `ll` and `dll` in the cell below.
# +
# say, we got a single spike train
y, fr = toyModel(X, theta) # spike train, firing rate
def ll(theta):
# TODO: implement log-likelihood function
return NotImplemented
def dll(theta):
# TODO: implement derivative of log-likelihood function wrt theta
return NotImplemented
# -
# **Exercise 2.2**: Assume the true parameters that we used to generate the data, $\mathbf{\theta}^*$, were unknown. We want to recover $\mathbf{\theta}^*$ starting from an initial guess $\mathbf{\theta}_0$. Fill the gaps in the code block below. How good do you recover $\mathbf{\theta}^*$? What happens if you change `step_size`?
# +
theta_true = theta.copy()
theta_initial = np.random.randn(len(theta))
print('theta_star : {}'.format(theta_true))
print('theta_0 : {}'.format(theta_initial))
def gradientAscent(theta_initial, step_size=0.0001, num_iterations=1000):
theta_hat = theta_initial.copy()
for i in range(0, num_iterations):
# TODO: fix the next lines
log_likelihood = ...
gradient = ...
theta_hat = theta_hat + ...
return theta_hat
theta_hat = gradientAscent(theta_initial)
print('theta_hat : {}'.format(theta_hat))
# -
# ## Extending the model
#
# Our simple model assumed independent firing in each time-bin that only depends on the stimulus. In reality, we know that the activity of neurons depends also on their recent firing history.
#
# The GLM frameworks allows to flexibly extend our model simply by adding additional covariates and corresponding parameters, i.e. by adding columns to design matrix $\mathbf{X}$ and entries to parameter vector $\theta$.
#
# Let us try introducing the recent spiking history $\mathbf{y}_{kt-\tau}, \ldots, \mathbf{y}_{kt-1}$ as additional covariates.
#
# The vector of covariates at time $t$ becomes
# $\mathbf{x}_{kt} = \big[1, \mathbf{u}_{kt-\delta+1 \ : \ tk}, \mathbf{y}_{kt-\tau \ : \ tk-1}\big]^\top$,
#
# and we extend the vector of parameters as $\boldsymbol{\theta} = \left[b, \mathbf{\beta}^\top, \mathbf{\psi}^\top \right]^\top$, with history kernel $\mathbf{\psi} \in \mathbb{R}^\tau$ and history kernel length $\tau \in \mathbb{N}$.
#
# **Question:** What other covariates could help improve our model?
# ### MLE Inference
#
# **Exercise 3.1:** Write a function that implements the new design matrix $\mathbf{X}$ (now depends on data $\mathbf{y}$). Note that we provide a function `createDataset()` to generate data from the extended model with given parameter vector $\theta$.
# +
tau = 5 # length of history kernel (in bins)
psi = - 1.0 * np.arange(0, tau)[::-1]
theta_true = np.hstack((theta, psi))
y = createDataset(U, T, K, theta_true, delta)
def extendedDesignMatrix(y):
# TODO: implement design matrix X with
# X[kt,:] = [1, w*cos(t), w*sin(t), y_{kt-tau:kt-1}]
return NotImplemented
X = extendedDesignMatrix(y) # you might have to re-run the cell defining ll() and dll()
# to update the used design matrix X and data y
# -
# **Exercise 3.2:** Write down the gradients for the extended model. What changes from our earlier simpler model?
# ## MAP inference
#
# The solution $\hat{\theta}$ obtained by gradient ascent on the log-likelihood depends on the data $\mathcal{D} = \{ (x_{tk}, y_{tk}) \}_{(t,k)}$. In particular for very short traces and few trials, this data only weakly constrains the solution.
# We can often improve our obtained solutions by adding prior knowledge regarding what 'good' solutions should look like. In probabilistic modeling, this can be done by introducing prior distributions $p(\theta)$ on the model parameters, which together with the likelihood $\mathcal{L}$ define a posterior distribution over parameters given the data $p(\theta | \mathbf{y})$,
#
# $$ \log p(\theta | \mathbf{y}) = \log p(\mathbf{y}|\theta) + \log p(\theta) - \log p(\mathbf{y}) = \mathcal{L}(\theta) + \log p(\theta) + const.$$
#
# Maximum a-posterio (MAP) estimates parameters $\theta$ by gradient ascent on the (log-)posterior.
#
# We will assume zero-mean Gaussian priors on $\beta, \psi$, i.e.
# \begin{align}
# p(\beta) &= \mathcal{N}(0, \Sigma_\beta) \\
# p(\psi) &= \mathcal{N}(0, \Sigma_\psi).
# \end{align}
# We will not assume an explicit prior on $b$, which effectively assumes $b$ to be distributed 'uniformly' over $\mathbb{R}$.
#
# ### Gradients
#
# Compared to maximum likelihood, MAP only requires adding the prior gradient:
#
# \begin{align}
# \frac{\partial}{\partial \theta} p(\theta|\mathbf{y}) = \frac{\partial}{\partial \theta} \mathcal{L}(\theta) + \frac{\partial}{\partial \theta}p(\theta)
# \end{align}
#
# ### Exercises
# **Exercise 4: ** Derive the gradients for the prior. If you get stuck, or if you want to verify the solution, ask the tutors for help.
# **Exercise 5:** Fill gaps in codeblock below.
# +
## priors
# select prior covariance for input weights
Sig_beta = np.eye(delta)
# select prior covariance for history kernel
ir = np.atleast_2d(np.arange(tau, 0, -1))
Sig_psi = np.exp(- np.abs(ir.T - ir)/5) # assuming smoothness
# convenience
P_beta, P_psi = np.linalg.inv(Sig_beta), np.linalg.inv(Sig_psi)
## functions and gradients
def po(theta):
# TODO: implement log-posterior density function
return NotImplemented
def dpo(theta):
# TODO: implement derivative of log-posterior density function wrt theta
return NotImplemented
# Hint: it can be helpful to first derive the functions for the prior below:
def pr(theta):
# TODO: implement log-prior density function
return NotImplemented
def dpr(theta):
# TODO: implement derivative of log-prior density function wrt theta
return NotImplemented
# leave as is
def ll(theta):
z = np.dot(theta, X)
return np.sum( y * z - link(z) )
# leave as is
def dll(theta):
z = np.dot(theta, X)
r = y - link(z)
return np.dot(X, r)
# -
# **Exercise 6:** Numerical gradient checking -- use the code below to numerically ensure your gradients are correct
# +
from scipy import optimize
thrn = np.random.normal(size=theta_true.shape)
print(optimize.check_grad(ll, dll, thrn))
print(optimize.check_grad(pr, dpr, thrn))
print(optimize.check_grad(po, dpo, thrn))
# -
# **Exercise 7:** Do inference (WIP)
# +
data = createDataset(1000, 1, theta_true, omega)
# TODO: implement gradient ascent
# -
# ## Automatic differentiation
#
# Instead of calculating the gradients w.r.t. the model parameters by hand, we can calculate them automatically. Our objective function consists of many elementary functions, each of which is differentiable. [Automatic differentiation (AD)](https://en.wikipedia.org/wiki/Automatic_differentiation) applies the chain rule to the expression graph of our objective to find the gradient.
#
# Here, we will use a Python library called `autograd` to find the gradient of our objective. AD is a central ingredient in libraries used for training artifical neural networks, including theano, TensorFlow and PyTorch.
#
# ### Installation
#
# Install the [`autograd` package](https://github.com/HIPS/autograd) through your package manager.
#
# Depending on how things are set up on your machine, install `autograd` by `pip3 install autograd --user` or by `pip install autograd --user`.
#
# You might need to restart the notebook kernel in case the simple example which follows fails with an import error. If you restart the kernel, make sure to re-run the cells. You can do that by choosing `Kernel > Restart & Run All` from the menu.
# ### `autograd` by a simple example
# +
import autograd.numpy as np # thinly-wrapped numpy
from autograd import grad # the only autograd function you may ever need
def tanh(x): # Define a function
y = np.exp(-x)
return (1.0 - y) / (1.0 + y)
grad_tanh = grad(tanh) # Obtain its gradient function
print('Gradient at x=1.0 (autograd) : {}'.format(grad_tanh(1.0)))
print('Gradient at x=1.0 (finite diff): {}'.format((tanh(1.0001) - tanh(0.9999)) / 0.0002))
# -
ipython nbconvert exercises.ipynb --to pdf
# ### Exercises
# **Exercise 8**: Redo exercise 2 using `autograd`. To do so, go to the first code cell of the notebook. Instead of `import numpy as np` use `import autograd.numpy as np`. Restart the notebook kernel and run through the notebook from the beginning, till you get to the point where you implemented `dll`. Replace `dll` by a AD version that relies on the function `grad` (see simple example).
# **Exercise 9:** Similar to the previous exercise, adapt the extended version of the model to rely on `autograd` for differentiation.
# **Exercise 10 (open ended)** : Come up with your own extensions to one of the models.
| notebooks/block2/GLM_exercises.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.9 64-bit (''pyvizenv'': conda)'
# metadata:
# interpreter:
# hash: ff898a576814a6948258c6a280b2b4b3234985999fc4b3b5c5881f334a02846f
# name: 'Python 3.7.9 64-bit (''pyvizenv'': conda)'
# ---
# # San Francisco Rental Prices Dashboard
#
# In this notebook, you will compile the visualizations from the previous analysis into functions that can be used for a Panel dashboard.
# imports
import panel as pn
pn.extension('plotly')
import plotly.express as px
import pandas as pd
import hvplot.pandas
import matplotlib.pyplot as plt
import os
from pathlib import Path
from dotenv import load_dotenv
# +
# Read the Mapbox API key
load_dotenv()
mapbox_token = os.getenv("MAPBOX_API_KEY")
# Set the Mapbox API
px.set_mapbox_access_token(mapbox_token)
# -
# # Import Data
# +
# Import the CSVs to Pandas DataFrames
file_path = Path("data/sfo_neighborhoods_census_data.csv")
sfo_data = pd.read_csv(file_path, index_col="year")
file_path = Path("data/neighborhoods_coordinates.csv")
df_neighborhood_locations = pd.read_csv(file_path)
# -
# - - -
# ## Panel Visualizations
#
# In this section, you will copy the code for each plot type from your analysis notebook and place it into separate functions that Panel can use to create panes for the dashboard.
#
# These functions will convert the plot object to a Panel pane.
#
# Be sure to include any DataFrame transformation/manipulation code required along with the plotting code.
#
# Return a Panel pane object from each function that can be used to build the dashboard.
#
# Note: Remove any `.show()` lines from the code. We want to return the plots instead of showing them. The Panel dashboard will then display the plots.
# Custom functions for reusability
def group_sort_top_10(df):
"""Group, Sort and Retrieve the Top 10 Sale Prices"""
# Group the data
sfo_data_neighborhood_grouped = df.groupby(['neighborhood']).mean()
# Sort the data based on sale price per sqr foot
sfo_data_neighborhood_grouped_sorted = sfo_data_neighborhood_grouped.sort_values(by='sale_price_sqr_foot', ascending=False)
# Assign the top 10
sfo_data_grouped_sorted_top_ten = sfo_data_neighborhood_grouped_sorted[0:10].reset_index()
return sfo_data_grouped_sorted_top_ten
# +
# Define Panel Visualization Functions
def housing_units_per_year():
"""Housing Units Per Year."""
# Calculate the mean number of housing units per year
sfo_data_housing_by_year = sfo_data.groupby(['year']).mean()
sfo_data_housing_units = sfo_data_housing_by_year['housing_units']
# Use the Pandas plot function to plot the average housing units per year.
fig_housing_units = plt.figure(figsize=(8, 5))
plt.subplot()
plt.bar(sfo_data_housing_units.index, sfo_data_housing_units)
plt.title('Housing Units in San Francisco from 2010 to 2016', size=18)
plt.xlabel('Year', size=14)
plt.ylabel('Housing Units', size=14)
# Determine the y axis limits based on minimum and maximum values and standard deviation
low = sfo_data_housing_units.min() - sfo_data_housing_units.std()
high = sfo_data_housing_units.max() + sfo_data_housing_units.std()
plt.ylim(low , high)
plt.close(fig_housing_units)
return fig_housing_units
def average_gross_rent():
"""Average Gross Rent in San Francisco Per Year."""
# Calculate the mean number of housing units per year
sfo_data_rent_by_year = sfo_data.groupby(['year']).mean()
# Calculate the average gross rent and average sale price per square foot
sfo_data_sale_price_and_gross_rent = pd.concat([sfo_data_rent_by_year['sale_price_sqr_foot'], sfo_data_rent_by_year['gross_rent']], axis='columns', join='inner')
# Plot the Average Gross Rent per Year as a Line Chart
fig_gross_rent = plt.figure(figsize=(8, 5))
plt.subplot()
plt.plot(sfo_data_sale_price_and_gross_rent.index, sfo_data_sale_price_and_gross_rent['gross_rent'])
plt.title('Average Gross Rent in San Francisco', size=18)
plt.xlabel('Year', size=14)
plt.ylabel('Gross Rent', size=14)
plt.xlim(sfo_data_sale_price_and_gross_rent.index.min() , sfo_data_sale_price_and_gross_rent.index.max())
plt.close(fig_gross_rent)
return fig_gross_rent
def average_sales_price():
"""Average Sales Price Per Year."""
# Calculate the mean number of housing units per year
sfo_data_sales_by_year = sfo_data.groupby(['year']).mean()
# Calculate the average gross rent and average sale price per square foot
sfo_data_sale_price_and_gross_rent = pd.concat([sfo_data_sales_by_year['sale_price_sqr_foot'], sfo_data_sales_by_year['gross_rent']], axis='columns', join='inner')
# Plot the Average Sales Price per Year as a line chart
fig_sales_price = plt.figure(figsize=(8, 5))
plt.subplot()
plt.plot(sfo_data_sale_price_and_gross_rent.index, sfo_data_sale_price_and_gross_rent['sale_price_sqr_foot'])
plt.title('Average Sale Price per Square Foot in San Francisco', size=18)
plt.xlabel('Year', size=14)
plt.ylabel('Average Sale Price', size=14)
plt.xlim(sfo_data_sale_price_and_gross_rent.index.min() , sfo_data_sale_price_and_gross_rent.index.max())
plt.close(fig_sales_price)
# Return plot
return fig_sales_price
def average_price_by_neighborhood():
"""Average Prices by Neighborhood."""
# Group by year and neighborhood and then create a new dataframe of the mean values
sfo_data_neighborhood_mean_values = sfo_data.groupby(['year', 'neighborhood']).mean()
# Use hvplot to create an interactive line chart of the average price per sq ft.
# The plot should have a dropdown selector for the neighborhood
price_by_neighborhood = sfo_data_neighborhood_mean_values.hvplot(
kind='line',
x='year',
y='sale_price_sqr_foot',
groupby='neighborhood',
xlabel='Year',
ylabel='Average Price Per Square Foot',
title='Average Price Per Square Foot By Neighborhood'
)
# Return hvplot
return price_by_neighborhood
def top_most_expensive_neighborhoods():
"""Top 10 Most Expensive Neighborhoods."""
# Getting the data from the top 10 expensive neighborhoods
# Group, sort and pull top 10 rows of data for neighborhoods
sfo_data_top_ten = group_sort_top_10(sfo_data)
# Plotting the data from the top 10 expensive neighborhoods
top_ten = sfo_data_top_ten.hvplot.bar(
x='neighborhood',
y='sale_price_sqr_foot',
xlim=(0, 900),
xlabel='Neighborhood',
ylabel='Sale Price Per Square Foot',
title='Top 10 Highest Priced Neighborhoods',
rot=90,
height=500,
)
# Return hvplot
return top_ten
def parallel_coordinates():
"""Parallel Coordinates Plot."""
# Getting the data from the top 10 expensive neighborhoods
# Group, sort and pull top 10 rows of data for neighborhoods
sfo_data_top_ten = group_sort_top_10(sfo_data)
# Parallel Coordinates Plot
p_coordinates = px.parallel_coordinates(
sfo_data_top_ten,
dimensions=["sale_price_sqr_foot", "housing_units", "gross_rent"],
color="sale_price_sqr_foot",
color_continuous_scale=px.colors.sequential.Inferno,
labels={
"sale_price_sqr_foot": "Sale Price Per Sqr Ft",
"housing_units": "Housing Units",
"gross_rent": "Gross Rent"
}
)
# Return plotly express plot
return p_coordinates
def parallel_categories():
"""Parallel Categories Plot."""
# Getting the data from the top 10 expensive neighborhoods
sfo_data_neighborhood_cat = sfo_data.groupby(['neighborhood']).mean()
sfo_data_neighborhood_cat = sfo_data_neighborhood_cat.sort_values(by='sale_price_sqr_foot', ascending=False)
sfo_data_top_ten = sfo_data_neighborhood_cat[0:10].reset_index()
# Parallel Categories Plot
p_categor = px.parallel_categories(
sfo_data_top_ten,
dimensions=["neighborhood", "sale_price_sqr_foot", "housing_units", "gross_rent"],
color="sale_price_sqr_foot",
color_continuous_scale=px.colors.sequential.Inferno,
labels={
"neighborhood": "Neighborhood",
"sale_price_sqr_foot": "Sale Price Per Sqr Ft",
"housing_units": "Housing Units",
"gross_rent": "Gross Rent"
}
)
# Return plotly express plot
return p_categor
def neighborhood_map():
"""Neighborhood Map"""
# Calculate the mean values, group and sort for each neighborhood
sfo_data_neighborhood_map_grouped = sfo_data.groupby(['neighborhood']).mean()
sfo_data_neighborhood_map_grouped_sorted = sfo_data_neighborhood_map_grouped.sort_values(by='neighborhood', ascending=True)
sfo_data_neighborhood_map_grouped_sorted = sfo_data_neighborhood_map_grouped_sorted.reset_index()
# Join the average values with the neighborhood locations
sfo_data_neighborhood_map_coordinates = pd.concat([sfo_data_neighborhood_map_grouped_sorted, df_neighborhood_locations], axis='columns', join='inner')
sfo_data_neighborhood_map_coordinates = sfo_data_neighborhood_map_coordinates.drop('neighborhood', axis=1)
# Create a scatter mapbox to analyze neighborhood info
neighborhood_map = px.scatter_mapbox(
sfo_data_neighborhood_map_coordinates,
lat="Lat",
lon="Lon",
size="sale_price_sqr_foot",
color="gross_rent",
color_continuous_scale=px.colors.cyclical.IceFire,
labels={
"sale_price_sqr_foot": "Sales Price Sqr Foot",
"Lat": "Latitude",
"Lon": "Longitude",
"gross_rent": "Gross Rent"
},
zoom=11,
mapbox_style="streets",
title="Average Sale Price Per Square Foot and Gross Rent in San Francisco",
height=500
)
# Returns data map
return neighborhood_map
# -
# ## Panel Dashboard
#
# In this section, you will combine all of the plots into a single dashboard view using Panel. Be creative with your dashboard design!
# +
# Catch phrase for visitors of the dashboard
dashboard_eye_catcher = "# Rental Investing Has Never Been Easier!"
# Welcome message
welcome_message = "## Welcome to San Francisco Rentals"
# Thank you
thank_you_message = "#### Property investing in San Francisco has never been better and now is the time to get in! Let us help you start your journey using our friendly dashboard so you are ready when you meet with our team of investment professionals for next steps."
# Dashboard description
site_description = "#### We are excited you have chosen to use our tools to find your next property in San Francisco. Let's discuss a little of what you see on your screen to make yourself right at home. Across the top of your window you find various tabs to help narrow your search and decision making from providing a little bit of information to performing detail analysis. Most of the charts are interactive, allowing you to move the maps and information around in the frames with the ability to zoom in, select areas, and highlight specific neighborhoods you would like to consider. Enjoy!"
# Average price and gross rent plot descriptions
avg_price_and_gross_rent_description = "#### The following interactive map allows you to zoom in and out using the scroll wheel on your mouse. Hover over a colored dot for detailed information such as geographic location coordinates, average sale price per square foot and gross rent in the area. The scale on the right side of the map gives a color friendly perspective to quickly see higher grossing rental areas. To quickly compare neighborhood pricing models and the top 10 valued neighborhoods, click on the Neighborhood Analysis tab."
# Neighborhood plot description
neighborhood_plot_description = "#### Below you will find the 10 highest valued neighborhoods in San Francisco. Click the next tab, Yearly Market Analysis, at the top of the dashboard to view trends of San Francisco year to year."
# Put descriptions and eye catching neighborhood map in one column
welcome_layout = pn.Column(dashboard_eye_catcher, welcome_message, thank_you_message, site_description, avg_price_and_gross_rent_description, neighborhood_map(), neighborhood_plot_description, top_most_expensive_neighborhoods())
# Title for yearly market analysis tab
yearly_market_analysis_title = "## Yearly Market Analysis"
yearly_market_analysis_layout = pn.Column(yearly_market_analysis_title)
# Put Yearly Market Analysis description in a column
yearly_market_analysis_description = "#### The following information shows the average sale price per square foot and the average gross rent trend year over year in San Francisco."
yearly_market_analysis_layout.append(yearly_market_analysis_description)
# Put yearly markets plots in one column
yearly_market_analysis_layout.append(average_sales_price())
yearly_market_analysis_layout.append(average_gross_rent())
# Add description for housing units per year
housing_units_description = "#### Below is a snapshot of the steady growing number of housing units in San Francisco! Click the next tab, Neighborhood Analysis, to view individual neighborhoods."
yearly_market_analysis_layout.append(housing_units_description)
# Add plot of housing units to column
yearly_market_analysis_layout.append(housing_units_per_year())
# Title for neighborhood analysis tab
neighborhood_analysis_title = "## Neighborhood Analysis"
neighborhood_analysis_layout = pn.Column(neighborhood_analysis_title)
# Neighborhood analysis description created in a column
neighborhood_analysis_description = "#### The following figure is interactive! Click in the drop down menu on the right to select a neighborhood and view the average price per square foot trends year over year. If you would like to see the exact location for a specific neighborhood, click the Welcome tab at the top to find a specific location on the map."
# Assign neighborhood description to column
neighborhood_analysis_layout.append(neighborhood_analysis_description)
# Put neighborhood analaysis plots in a column
neighborhood_analysis_layout.append(average_price_by_neighborhood())
# Title for a detailed look tab
a_detailed_look_title = "## A Detailed Look"
a_detailed_look_layout = pn.Column(a_detailed_look_title)
# A detailed look layout description
a_detailed_look_description = "#### The following figures are coordinate (top figure) and category (bottom figure) plots to further dive into a detailed analysis of housing in San Francisco. The coordinate plot assists in finding groups of data and how they corrolate with other data points. The categories plot represents the connections of data as well with a little difference by using categories to visually see if groups of neighborhood connections may have a lower or higher number of housing units and possibly an increase or decrease of gross rent compared to other neighborhoods."
# Assign description to column
a_detailed_look_layout.append(a_detailed_look_description)
# Put parallel plots in column
a_detailed_look_layout.append(parallel_coordinates())
a_detailed_look_layout.append(parallel_categories())
# Create tabs
neighborhood_panel = pn.Tabs(
("Welcome", welcome_layout),
("Yearly Market Analysis", yearly_market_analysis_layout),
("Neighborhood Analysis", neighborhood_analysis_layout),
("A Detailed Look", a_detailed_look_layout),
)
# -
# ## Serve the Panel Dashboard
neighborhood_panel.servable()
| dashboard.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
from gensim.models import Word2Vec
# -
# # Load Data
df = pd.read_csv("../datasets/netflix_titles.csv")
df.head(5)
# # Data Cleaning
# +
import re
from nltk.corpus import stopwords
from nltk.tokenize import RegexpTokenizer
def data_cleaning(text):
# make lower case
text = text.lower()
# remove stopwords
text = text.split()
stops = set(stopwords.words("english"))
text = [w for w in text if not w in stops]
text = " ".join(text)
# remove punctuation
tokenizer = RegexpTokenizer(r'[a-zA-z]+')
text = tokenizer.tokenize(text)
text = " ".join(text)
return text
# -
df['cleaned'] = df['description'].apply(data_cleaning)
df['cleaned'][:5]
corpus = []
for words in df['cleaned']:
corpus.append(words.split())
# # Word Embedding (with pretrained embedded word)
import urllib.request
urllib.request.urlretrieve("https://s3.amazonaws.com/dl4j-distribution/GoogleNews-vectors-negative300.bin.gz", filename="GoogleNews-vectors-negative300.bin.gz")
word2vec_model = Word2Vec(size = 300, window=5, min_count = 2, workers = -1)
word2vec_model.build_vocab(corpus)
word2vec_model.intersect_word2vec_format('GoogleNews-vectors-negative300.bin.gz', lockf=1.0, binary=True)
word2vec_model.train(corpus, total_examples = word2vec_model.corpus_count, epochs = 15)
# # Average of word vectors
def vectors(document_list):
document_embedding_list = []
for line in document_list:
doc2vec = None
count = 0
for word in line.split():
if word in word2vec_model.wv.vocab:
count+=1
if doc2vec is None:
doc2vec = word2vec_model[word]
else:
doc2vec = doc2vec + word2vec_model[word]
if doc2vec is not None:
doc2vec = doc2vec / count
document_embedding_list.append(doc2vec)
return document_embedding_list
document_embedding_list = vectors(df['cleaned'])
print('number of vector',len(document_embedding_list))
from sklearn.metrics.pairwise import cosine_similarity
cosine_similarities = cosine_similarity(document_embedding_list, document_embedding_list)
print(cosine_similarities)
def recommendation(title):
show = df[['title','listed_in','description']]
indices = pd.Series(df.index, index = df['title']).drop_duplicates()
idx = indices[title]
sim_scores = list(enumerate(cosine_similarities[idx]))
sim_scores = sorted(sim_scores, key = lambda x: x[1], reverse=True)
sim_scores = sim_scores[1:6]
show_indices = [i[0] for i in sim_scores]
recommend = show.iloc[show_indices].reset_index(drop=True)
print('Recommended list')
recommend_df = pd.DataFrame(recommend)
recommend_df.head()
recommendation("3%")
| Netfilx/NetflixRecommeder.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import json
# open username and password from another file.
with open('C:\\Users\\xianx\\Documents\\xianw_secret.json') as f:
data = json.load(f)
username = data['username']
password = data['password']
#print (username+" :" +password)
# import pyodbc
import pyodbc
# setup connection to SQLserver, with specific credentials
connection=pyodbc.connect("DRIVER={SQL Server};"
"SERVER=LAPTOP-8VUQJOSL;"
"DATABASE=python_movies;"
"Trusted_Connection=Yes;"
"UID="+username+";"
"PWD="+password)
cursor=connection.cursor()
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
import numpy as np
#with open('C:\\Users\\xianx\\Documents\\bg.jpg') as pic:
# picture = json.load(pic)
#img=mpimg.imread(picture)
#imgplot = plt.imshow(img)
#plt.style.use(img)
# #%matplotlib inline
plt.rcParams.update({'figure.figsize':(8,5), 'figure.dpi':100})
sql1='select count(Score), year(ReleaseDate) from [dbo].[MetacriticMovies] where score>85 group by year(ReleaseDate)'
#sql2='select count(Score), year(ReleaseDate) from [dbo].[MetacriticMovies] where score<60 group by year(ReleaseDate)'
cursor.execute(sql1)
data1=cursor.fetchall()
#data2=cursor.fetchall()
#X = np.arange(3)
count1=[]
years=[]
for row in data1:
count1.append(row[0])
years.append(row[1])
#for row in data2:
# count2.append(row[0])
fig = plt.figure()
ax = fig.add_axes([0,0,1,1])
ax.bar(years,count1,color = 'b')
plt.gca().set(title='2010-2020 Top Movie Count Per Year',ylabel='Count');
# -
| backup.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Inference with GPs
# ## The Marginal Likelihood
#
# In the previous notebook, we learned how to construct and sample from a simple GP. This is useful for making predictions, i.e., interpolating or extrapolating based on the data you measured. But the true power of GPs comes from their application to *regression* and *inference*: given a dataset $D$ and a model $M(\theta)$, what are the values of the model parameters $\theta$ that are consistent with $D$? The parameters $\theta$ can be the hyperparameters of the GP (the amplitude and time scale), the parameters of some parametric model, or all of the above.
#
# A very common use of GPs is to model things you don't have an explicit physical model for, so quite often they are used to model "nuisances" in the dataset. But just because you don't care about these nuisances doesn't mean they don't affect your inference: in fact, unmodelled correlated noise can often lead to strong biases in the parameter values you infer. In this notebook, we'll learn how to compute likelihoods of Gaussian Processes so that we can *marginalize* over the nuisance parameters (given suitable priors) and obtain unbiased estimates for the physical parameters we care about.
# Given a set of measurements $y$ distributed according to
# $$
# \begin{align}
# y \sim \mathcal{N}(\mathbf{\mu}(\theta), \mathbf{\Sigma}(\alpha))
# \end{align}
# $$
# where $\theta$ are the parameters of the mean model $\mu$ and $\alpha$ are the hyperparameters of the covariance model $\mathbf{\Sigma}$, the *marginal likelihood* of $y$ is
# $$
# \begin{align}
# \ln P(y | \theta, \alpha) = -\frac{1}{2}(y-\mu)^\top \mathbf{\Sigma}^{-1} (y-\mu) - \frac{1}{2}\ln |\mathbf{\Sigma}| - \frac{N}{2} \ln 2\pi
# \end{align}
# $$
#
# where $||$ denotes the determinant and $N$ is the number of measurements. The term *marginal* refers to the fact that this expression implicitly integrates over all possible values of the Gaussian Process; this is not the likelihood of the data given one particular draw from the GP, but given the ensemble of all possible draws from $\mathbf{\Sigma}$.
# <div style="background-color: #D6EAF8; border-left: 15px solid #2E86C1;">
# <h1 style="line-height:2.5em; margin-left:1em;">Exercise 1</h1>
# </div>
#
# Define a function ``ln_gp_likelihood(t, y, sigma, **kwargs)`` that returns the log-likelihood defined above for a vector of measurements ``y`` at a set of times ``t`` with uncertainty ``sigma``. As before, ``**kwargs`` should get passed direcetly to the kernel function. Note that you're going to want to use [np.linalg.slogdet](https://docs.scipy.org/doc/numpy-1.14.0/reference/generated/numpy.linalg.slogdet.html) to compute the log-determinant of the covariance instead of ``np.log(np.linalg.det)``. (Why?)
# <div style="background-color: #D6EAF8; border-left: 15px solid #2E86C1;">
# <h1 style="line-height:2.5em; margin-left:1em;">Exercise 2</h1>
# </div>
#
# The following dataset was generated from a zero-mean Gaussian Process with a Squared Exponential Kernel of unity amplitude and unknown timescale. Compute the marginal log likelihood of the data over a range of reasonable values of $l$ and find the maximum. Plot the **likelihood** (not log likelihood) versus $l$; it should be pretty Gaussian. How well are you able to constrain the timescale of the GP?
import matplotlib.pyplot as plt
t, y, sigma = np.loadtxt("data/sample_data.txt", unpack=True)
plt.plot(t, y, "k.", alpha=0.5, ms=3)
plt.xlabel("time")
plt.ylabel("data");
# <div style="background-color: #D6EAF8; border-left: 15px solid #2E86C1;">
# <h1 style="line-height:2.5em; margin-left:1em;">Exercise 3a</h1>
# </div>
#
# The timeseries below was generated by a linear function of time, $y(t)= mt + b$. In addition to observational uncertainty $\sigma$ (white noise), there is a fair bit of correlated (red) noise, which we will assume is well described
# by the squared exponential covariance with a certain (unknown) amplitude $A$ and timescale $l$.
#
# Your task is to estimate the values of $m$ and $b$, the slope and intercept of the line, respectively. In this part of the exercise, **assume there is no correlated noise.** Your model for the $n^\mathrm{th}$ datapoint is thus
#
# $$
# \begin{align}
# y_n \sim \mathcal{N}(m t_n + b, \sigma_n\mathbf{I})
# \end{align}
# $$
#
# and the probability of the data given the model can be computed by calling your GP likelihood function:
#
# ```python
# def lnprob(params):
# m, b = params
# model = m * t + b
# return ln_gp_likelihood(t, y - model, sigma, A=0, l=1)
# ```
#
# Note, importantly, that we are passing the **residual vector**, $y - (mt + b)$, to the GP, since above we coded up a zero-mean Gaussian process. We are therefore using the GP to model the **residuals** of the data after applying our physical model (the equation of the line).
#
# To estimate the values of $m$ and $b$ we could generate a fine grid in those two parameters and compute the likelihood at every point. But since we'll soon be fitting for four parameters (in the next part), we might as well upgrade our inference scheme and use the ``emcee`` package to do Markov Chain Monte Carlo (MCMC). If you haven't used ``emcee`` before, check out the first few tutorials on the [documentation page](https://emcee.readthedocs.io/en/latest/). The basic setup for the problem is this:
#
# ```python
# import emcee
# sampler = emcee.EnsembleSampler(nwalkers, ndim, lnprob)
#
# print("Running burn-in...")
# p0, _, _ = sampler.run_mcmc(p0, nburn) # nburn = 500 should do
# sampler.reset()
#
# print("Running production...")
# sampler.run_mcmc(p0, nsteps); # nsteps = 1000 should do
# ```
#
# where ``nwalkers`` is the number of walkers (something like 20 or 30 is fine), ``ndim`` is the number of dimensions (2 in this case), and ``lnprob`` is the log-probability function for the data given the model. Finally, ``p0`` is a list of starting positions for each of the walkers. Pick some fiducial/eyeballed value for $m$ and $b$, then add a small random number to each to generate different initial positions for each walker. This will initialize all walkers in a ball centered on some point, and as the chain progresses they'll diffuse out and begin to explore the posterior.
#
# Once you have sampled the posterior, plot several draws from it on top of the data. Also plot the **true** line that generated the dataset (given by the variables ``m_true`` and ``b_true`` below). Do they agree, or is there bias in your inferred values? Use the ``corner`` package to plot the joint posterior. How many standard deviations away from the truth are your inferred values?
t, y, sigma = np.loadtxt("data/sample_data_line.txt", unpack=True)
m_true, b_true, A_true, l_true = np.loadtxt("data/sample_data_line_truths.txt", unpack=True)
plt.errorbar(t, y, yerr=sigma, fmt="k.", label="observed")
plt.plot(t, m_true * t + b_true, color="C0", label="truth")
plt.legend(fontsize=12)
plt.xlabel("time")
plt.ylabel("data");
# <div style="background-color: #D6EAF8; border-left: 15px solid #2E86C1;">
# <h1 style="line-height:2.5em; margin-left:1em;">Exercise 3b</h1>
# </div>
#
# This time, let's actually model the correlated noise. Re-define your ``lnprob`` function to accept four parameters (slope, intercept, amplitude, and timescale). If you didn't before, it's a good idea to enforce some priors to keep the parameters within reasonable (and physical) ranges. If any parameter falls outside this range, have ``lnprob`` return negative infinity (i.e., zero probability).
#
# You'll probably want to run your chains for a bit longer this time, too. As before, plot some posterior samples for the line, as well as the corner plot. How did you do this time? Is there any bias in your inferred values? How does the variance compare to the previous estimate?
# <div style="background-color: #D6EAF8; border-left: 15px solid #2E86C1;">
# <h1 style="line-height:2.5em; margin-left:1em;">Exercise 3c</h1>
# </div>
#
# If you didn't do this already, re-plot the posterior samples on top of the data, but this time draw them from the GP, *conditioned on the data*. How good is the fit?
| gps/02-Inference.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #GPU Accelerated Principal Component Analysis (PCA) using RAPIDS on a Sample Dataset with CPU vs GPU comparison
# #### Verifying GPUs
#
# RAPIDS requires GPUs with Pascal Architecture or better. That means any GPUs starting with K series (e.g. K80) or M series (e.g., M60) would not work. You can use the `nvidia-smi` command to verify the type of your GPU as well as the memory size which may be needed for some of the RAPIDS examples.
# %sh nvidia-smi
# ## Let's begin by importing RAPIDS and scikit learn libraries!
import numpy as np
import pandas as pd
from sklearn.decomposition import PCA as skPCA
from cuml import PCA as cumlPCA
import cudf
import os
# ## Downloading the data
# For this example we are downloading a sample dataset (Mortgage.csv) from Nvidia's repository
#
# __We have already completed Data prep (ETL) and feature engineering on this dataset and the dataset is ready for Machine Learning__
#
# __Note:__ If you already have a dataset, please don't run the next command
# +
# %sh
wget https://github.com/rapidsai/notebooks-extended/raw/master/data/mortgage/mortgage.csv.gz
gunzip mortgage.csv.gz
# mkdir -p /dbfs/RAPIDS/mortgage
# cp mortgage.csv /dbfs/RAPIDS/mortgage/
# -
# ## Loading the data with Spark
data = spark.read.csv("/RAPIDS/mortgage/mortgage.csv", header = "false", inferSchema = "true", sep = ",")
# __Let's check out the dataset in the spark dataframe and count the number of rows in the dataset__
display(data)
dataCount = data.count() # We're storing this value for later use
print(dataCount)
# __Helper Functions to compare CPU vs GPU results__
# +
from sklearn.metrics import mean_squared_error
def array_equal(a,b,threshold=2e-3,with_sign=True):
a = to_nparray(a)
b = to_nparray(b)
if with_sign == False:
a,b = np.abs(a),np.abs(b)
error = mean_squared_error(a,b)
res = error<threshold
return res
def to_nparray(x):
if isinstance(x,np.ndarray) or isinstance(x,pd.DataFrame):
return np.array(x)
elif isinstance(x,np.float64):
return np.array([x])
elif isinstance(x,cudf.DataFrame) or isinstance(x,cudf.Series):
return x.to_pandas().values
return x
# -
# __Helper Functions to remove any null values__
def null_workaround(df, **kwargs):
for column, data_type in df.dtypes.items():
if str(data_type) == "category":
df[column] = df[column].astype('int32').fillna(-1)
if str(data_type) in ['int8', 'int16', 'int32', 'int64', 'float32', 'float64']:
df[column] = df[column].fillna(-1)
return df
# ## Converting the Spark Dataframe into Pandas Dataframe
# __Load data function allows you to create a user defined sample of your data and converts the spark dataframe to pandas dataframe. Then, it removes any null values in the dataset. If you want to to experiment with a different dataset sizes, use the random array generator to load the random data.__
def load_data(nrows, ncols):
try:
frac = nrows/dataCount # as sample() takes an integer, we are creating a factor by which to get the approximate number of rows
print(frac) # just for checks :)
if (frac > 1):
frac = 1.0
print(frac) # just for checks++ :)
X = data.sample(True, frac)
print(X)
df = X.toPandas() # we then convert the Spark Dataframe to Pandas.
df = null_workaround(df)
print("everything worked")
except Exception as e:
print(e)
print('use random data')
X = np.random.rand(nrows,ncols)
df = pd.DataFrame({'fea%d'%i:X[:,i] for i in range(X.shape[1])})
print("only random data")
return df
# __Setting up data in Pandas Dataframe using Load data and null workaround function__
# +
# %%time
nrows = 2**20
nrows = int(nrows * 1.5)
ncols = 400
X = load_data(nrows,ncols)
X = null_workaround(X)
# -
# # Brief Intro to PCA parameters
# Let's take a look into all possible parameters that we can use when applying PCA:
# http://scikitlearn.org/stable/modules/generated/sklearn.decomposition.PCA.html
#
# We will start here with the following :
#
# __n_components__ : int, float, None or string
# Number of components to keep. if n_components is not set all components are kept
#
# __whiten__ : bool, optional (default False)
# When True (False by default) the components_ vectors are multiplied by the square root of n_samples and then divided by the singular values to ensure uncorrelated outputs with unit component-wise variances. Whitening will remove some information from the transformed signal (the relative variance scales of the components) but can sometime improve the predictive accuracy of the downstream estimators by making their data respect some hard-wired assumptions
#
# __random_state__ : int, RandomState instance or None, optional (default None)
# If int, random_state is the seed used by the random number generator
#
# __svd_solver__ : string {‘auto’, ‘full’, ‘arpack’, ‘randomized’}
# If "full" :run exact full SVD calling the standard LAPACK solver via scipy.linalg.svd and select the components by postprocessing
n_components = 10
whiten = False
random_state = 42
svd_solver="full"
# # Run PCA on CPU
# Let's check the time needed to execute PCA function using standard sklearn library.
# __Note: this algorithm runs on CPU only.__
import multiprocessing
print(multiprocessing.cpu_count()) # Return the number of CPUs in the system.
# %%time
pca_sk = skPCA(n_components=n_components,svd_solver=svd_solver,
whiten=whiten, random_state=random_state)
result_sk = pca_sk.fit_transform(X)
# # Run PCA on GPU
# Now, before we execute PCA function using RAPIDS cuml library we will first read the data in GPU data format using cudf.
#
# __cudf__ - GPU DataFrame manipulation library https://github.com/rapidsai/cudf
#
# __cuml__ - suite of libraries that implements a machine learning algorithms within the RAPIDS data science ecosystem https://github.com/rapidsai/cuml
# %%time
Xt = cudf.DataFrame.from_pandas(X) # Convert Pandas Dataframe to GPU Dataframe!
print(Xt)
# %%time
pca_cuml = cumlPCA(n_components=n_components,svd_solver=svd_solver,
whiten=whiten, random_state=random_state)
result_cuml = pca_cuml.fit_transform(Xt)
for attr in ['singular_values_','components_','explained_variance_',
'explained_variance_ratio_']:
passed = array_equal(getattr(pca_sk,attr),getattr(pca_cuml,attr))
message = 'compare pca: cuml vs sklearn {:>25} {}'.format(attr,'equal' if passed else 'NOT equal')
print(message)
# Spark ML accelerated with RAPIDS
passed = array_equal(result_sk,result_cuml)
message = 'compare pca: cuml vs sklearn transformed results %s'%('equal'if passed else 'NOT equal')
print(message)
| blog_notebooks/databricks/spark_rapids_pca_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Seasonality, Trend and Noise
# > You will go beyond summary statistics by learning about autocorrelation and partial autocorrelation plots. You will also learn how to automatically detect seasonality, trend and noise in your time series data. This is the Summary of lecture "Visualizing Time-Series data in Python", via datacamp.
#
# - toc: true
# - badges: true
# - comments: true
# - author: <NAME>
# - categories: [Python, Datacamp, Time_Series_Analysis, Visualization]
# - image: images/trend_seasonal.png
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
plt.rcParams['figure.figsize'] = (10, 5)
plt.style.use('fivethirtyeight')
# -
# ## Autocorrelation and Partial autocorrelation
# - Autocorrelation in time series data
# - Autocorrelation is measured as the correlation between a time series and a delayed copy of itself
# - For example, an autocorrelation of order 3 returns the correlation between a time series at points($t_1, t_2, t_3$) and its own values lagged by 3 time points. ($t_4, t_5, t_6$)
# - It is used to find repetitive paterns or periodic signal it time series
# - Partial autocorrelation in time series data
# - Contrary to autocorrelation, partial autocorrelation removes the effect of previous time points
# - For example, a partial autocorrelatio nfunction of order 3 returns the correlation between out time series ($t_1, t_2, t_3$) and lagged values of itself by 3 time points ($t_4, t_5, t_6$), but only after removing all effects attributable to lags 1 and 2
# ### Autocorrelation in time series data
# In the field of time series analysis, autocorrelation refers to the correlation of a time series with a lagged version of itself. For example, an autocorrelation of order 3 returns the correlation between a time series and its own values lagged by 3 time points.
#
# It is common to use the autocorrelation (ACF) plot, also known as self-autocorrelation, to visualize the autocorrelation of a time-series. The ```plot_acf()``` function in the statsmodels library can be used to measure and plot the autocorrelation of a time series.
co2_levels = pd.read_csv('./dataset/ch2_co2_levels.csv')
co2_levels.set_index('datestamp', inplace=True)
co2_levels = co2_levels.fillna(method='bfill')
# +
from statsmodels.graphics import tsaplots
# Display
fig = tsaplots.plot_acf(co2_levels['co2'], lags= 24);
# -
# ### Interpret autocorrelation plots
# If autocorrelation values are close to 0, then values between consecutive observations are not correlated with one another. Inversely, autocorrelations values close to 1 or -1 indicate that there exists strong positive or negative correlations between consecutive observations, respectively.
#
# In order to help you asses how trustworthy these autocorrelation values are, the ```plot_acf()``` function also returns confidence intervals (represented as blue shaded regions). If an autocorrelation value goes beyond the confidence interval region, you can assume that the observed autocorrelation value is statistically significant.
# ### Partial autocorrelation in time series data
# Like autocorrelation, the partial autocorrelation function (PACF) measures the correlation coefficient between a time-series and lagged versions of itself. However, it extends upon this idea by also removing the effect of previous time points. For example, a partial autocorrelation function of order 3 returns the correlation between our time series ($t_1, t_2, t_3, \dots$) and its own values lagged by 3 time points ($t_4, t_5, t_6, \dots$), but only after removing all effects attributable to lags 1 and 2.
#
# The ```plot_pacf()``` function in the statsmodels library can be used to measure and plot the partial autocorrelation of a time series.
# Display the partial autocorrelation plot of your time series
fig = tsaplots.plot_pacf(co2_levels['co2'], lags=24);
# ### Interpret partial autocorrelation plots
# If partial autocorrelation values are close to 0, then values between observations and lagged observations are not correlated with one another. Inversely, partial autocorrelations with values close to 1 or -1 indicate that there exists strong positive or negative correlations between the lagged observations of the time series.
#
# The ```.plot_pacf()``` function also returns confidence intervals, which are represented as blue shaded regions. If partial autocorrelation values are beyond this confidence interval regions, then you can assume that the observed partial autocorrelation values are statistically significant.
# ## Seasonality, trend and noise in time series data
# - The properties of time series
# - Seasonality: does the data display a clear periodic pattern?
# - Trend: does the data follow a consistent upwards or downwards slope?
# - Noise: are there any outlier points or missing values that are not consistent with the rest of the data?
# ### Time series decomposition
# You can rely on a method known as time-series decomposition to automatically extract and quantify the structure of time-series data. The statsmodels library provides the ```seasonal_decompose()``` function to perform time series decomposition out of the box.
# ```python
# decomposition = sm.tsa.seasonal_decompose(time_series)
# ```
# You can extract a specific component, for example seasonality, by accessing the seasonal attribute of the ```decomposition``` object.
co2_levels.index = pd.to_datetime(co2_levels.index)
# +
import statsmodels.api as sm
# Perform time series decomposition
decomposition = sm.tsa.seasonal_decompose(co2_levels)
# Print the seasonality component
print(decomposition.seasonal)
# -
# ### Plot individual components
# It is also possible to extract other inferred quantities from your time-series decomposition object. The following code shows you how to extract the observed, trend and noise (or residual, ```resid```) components.
# ```python
# observed = decomposition.observed
# trend = decomposition.trend
# residuals = decomposition.resid
# ```
# You can then use the extracted components and plot them individually.
# +
# Extract the trend component
trend = decomposition.trend
# Plot the values of the trend
ax = trend.plot(figsize=(12, 6), fontsize=10);
# Specify axis labels
ax.set_xlabel('Date', fontsize=10);
ax.set_title('Seasonal component the CO2 time-series', fontsize=10);
# -
# ## A quick review
#
# ### Visualize the airline dataset
# You will now review the contents of chapter 1. You will have the opportunity to work with a new dataset that contains the monthly number of passengers who took a commercial flight between January 1949 and December 1960.
airline = pd.read_csv('./dataset/ch3_airline_passengers.csv', parse_dates=['Month'], index_col='Month')
airline.info()
# +
# Plot the time series in your dataframe
ax = airline.plot(color='blue', fontsize=12);
# Add a red vertical line at the date 1955-12-01
ax.axvline('1955-12-01', color='red', linestyle='--');
# Specify the labels in your plot
ax.set_xlabel('Date', fontsize=12);
ax.set_title('Number of Monthly Airline Passengers', fontsize=12);
# -
# ### Analyze the airline dataset
# In Chapter 2 you learned:
#
# - How to check for the presence of missing values, and how to collect summary statistics of time series data contained in a pandas DataFrame.
# - To generate boxplots of your data to quickly gain insight in your data.
# - Display aggregate statistics of your data using groupby().
# +
# Print out the number of missing values
print(airline.isnull().sum())
# Print out summary statistics of the airline DataFrame
print(airline.describe())
# +
# Display boxplot of airline values
ax = airline.boxplot();
# Specify the title of your plot
ax.set_title('Boxplot of Monthly Airline\nPassengers Count', fontsize=20);
# +
# Get month for each dates from the index of airline
index_month = airline.index.month
# Compute the mean number of passengers for each month of the year
mean_airline_by_month = airline.groupby(index_month).mean()
# Plot the mean number of passengers for each month of the year
mean_airline_by_month.plot();
plt.legend(fontsize=20);
# -
# ### Time series decomposition of the airline dataset
# In this exercise, you will apply time series decomposition to the ```airline``` dataset, and visualize the ```trend``` and ```seasonal``` componenets.
# +
# Perform time series decomposition
decomposition = sm.tsa.seasonal_decompose(airline)
# Extract the trend and seasonal components
trend = decomposition.trend
seasonal = decomposition.seasonal
# -
airline_decomposed = pd.concat([trend, seasonal], axis=1)
# +
# Print the first 5 rows of airline_decomposed
print(airline_decomposed.head(5))
# Plot the values of the airline_decomposed DataFrame
ax = airline_decomposed.plot(figsize=(12, 6), fontsize=15);
# Specify axis labels
ax.set_xlabel('Date', fontsize=15);
plt.legend(fontsize=15);
plt.savefig('../images/trend_seasonal.png')
| _notebooks/2020-06-13-01-Seasonality-Trend-and-Noise.ipynb |