code
stringlengths 38
801k
| repo_path
stringlengths 6
263
|
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Environment (conda_tensorflow_p36)
# language: python
# name: conda_tensorflow_p36
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/amilkh/cs230-fer/blob/transfer-learning/Final-SeNet50.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab_type="code" id="gwdg7Sv3XBaP" outputId="898c8cb4-3e8b-4f63-a2da-fc392f6d08cc" colab={"base_uri": "https://localhost:8080/", "height": 527}
# %tensorflow_version 1.x
# !pip install keras-vggface
# !pip install scikit-image
# !pip install pydot
# + colab_type="code" id="2nz38mJZXN_P" outputId="20e85bee-7622-43fa-c448-c31cb819fbd0" colab={"base_uri": "https://localhost:8080/", "height": 34}
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import tensorflow as tf
from tensorflow.keras.layers import *
from tensorflow.python.lib.io import file_io
# %matplotlib inline
import keras
from keras import backend as K
from keras.callbacks import ModelCheckpoint, EarlyStopping
from keras.models import load_model
from keras.preprocessing.image import ImageDataGenerator
from keras_vggface.vggface import VGGFace
from keras.utils import plot_model
from sklearn.metrics import *
from keras.engine import Model
from keras.layers import Input, Flatten, Dense, Activation, Conv2D, MaxPool2D, BatchNormalization, Dropout, MaxPooling2D
import skimage
from skimage.transform import rescale, resize
import pydot
# + id="1fZczU8lGkX-" colab_type="code" outputId="d9e5dcea-c0be-4f4c-99b4-4ce966c3ea93" colab={"base_uri": "https://localhost:8080/", "height": 122}
from google.colab import drive
drive.mount('/content/drive')
# + colab_type="code" id="nUcd6yIGduUW" outputId="1bd89817-9058-4050-fb7f-5b7071f4c9b8" colab={"base_uri": "https://localhost:8080/", "height": 51}
print(tf.__version__)
print(keras.__version__)
# + id="v60q28mDHnN9" colab_type="code" colab={}
EPOCHS = 100
BS = 128
DROPOUT_RATE = 0.5
FROZEN_LAYER_NUM = 201
ADAM_LEARNING_RATE = 0.001
SGD_LEARNING_RATE = 0.01
SGD_DECAY = 0.0001
Resize_pixelsize = 197
# + id="itKZtFV0F7b1" colab_type="code" outputId="e03b20a2-6738-4c26-b216-074a9f8df5fa" colab={"base_uri": "https://localhost:8080/", "height": 632}
vgg_notop = VGGFace(model='senet50', include_top=False, input_shape=(Resize_pixelsize, Resize_pixelsize, 3), pooling='avg')
last_layer = vgg_notop.get_layer('avg_pool').output
x = Flatten(name='flatten')(last_layer)
x = Dropout(DROPOUT_RATE)(x)
x = Dense(4096, activation='relu', name='fc6')(x)
x = Dropout(DROPOUT_RATE)(x)
x = Dense(1024, activation='relu', name='fc7')(x)
x = Dropout(DROPOUT_RATE)(x)
# l=0
# for layer in vgg_notop.layers:
# print(layer,"["+str(l)+"]")
# l=l+1
batch_norm_indices = [2, 6, 9, 12, 21, 25, 28, 31, 42, 45, 48, 59, 62, 65, 74, 78, 81, 84, 95, 98, 101, 112, 115, 118, 129, 132, 135, 144, 148, 151, 154, 165, 168, 171, 182, 185, 188, 199, 202, 205, 216, 219, 222, 233, 236, 239, 248, 252, 255, 258, 269, 272, 275]
for i in range(FROZEN_LAYER_NUM):
if i not in batch_norm_indices:
vgg_notop.layers[i].trainable = False
# print('vgg layer 2 is trainable: ' + str(vgg_notop.layers[2].trainable))
# print('vgg layer 3 is trainable: ' + str(vgg_notop.layers[3].trainable))
out = Dense(7, activation='softmax', name='classifier')(x)
model = Model(vgg_notop.input, out)
optim = keras.optimizers.Adam(lr=ADAM_LEARNING_RATE, beta_1=0.9, beta_2=0.999, epsilon=1e-08, decay=0.0)
#optim = keras.optimizers.Adam(lr=0.0005, beta_1=0.9, beta_2=0.999, epsilon=1e-08, decay=0.0)
sgd = keras.optimizers.SGD(lr=SGD_LEARNING_RATE, momentum=0.9, decay=SGD_DECAY, nesterov=True)
rlrop = keras.callbacks.ReduceLROnPlateau(monitor='val_acc',mode='max',factor=0.5, patience=10, min_lr=0.00001, verbose=1)
model.compile(optimizer=sgd, loss='categorical_crossentropy', metrics=['accuracy'])
# plot_model(model, to_file='model2.png', show_shapes=True)
# + id="v0mXUNZB-yI5" colab_type="code" colab={}
# ! rm -rf train; mkdir train
# ! unzip -q '/content/drive/My Drive/cs230 project/dataset/emotion.zip' -d train
# ! unzip -q '/content/drive/My Drive/cs230 project/dataset/facesdb.zip' -d train
# ! unzip -q '/content/drive/My Drive/cs230 project/dataset/fer2013/train.zip' -d train
# ! unzip -q '/content/drive/My Drive/cs230 project/dataset/googlesearch.zip' -d train
# ! unzip -q '/content/drive/My Drive/cs230 project/dataset/googleset.zip' -d train
# ! unzip -q '/content/drive/My Drive/cs230 project/dataset/jaffe.zip' -d train
# ! unzip -q '/content/drive/My Drive/cs230 project/dataset/umea.zip' -d train
# + id="TUuN9SLX_Qvl" colab_type="code" colab={}
# ! rm -rf dev; mkdir dev
# ! unzip -q '/content/drive/My Drive/cs230 project/dataset/fer2013/test-public.zip' -d dev
# ! rm -rf test; mkdir test
# ! unzip -q '/content/drive/My Drive/cs230 project/dataset/fer2013/test-private.zip' -d test
# + id="egSWDlktpwwQ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 136} outputId="6e23e651-d3d0-4800-d044-bb4310698f13" language="bash"
# root='/content/test/'
# IFS=$(echo -en "\n\b")
# (for dir in $(ls -1 "$root")
# do printf "$dir: " && ls -i "$root$dir" | wc -l
# done)
# + id="560jefnZ_Cyq" colab_type="code" colab={}
from tensorflow.keras.preprocessing.image import ImageDataGenerator
def get_datagen(dataset, aug=False):
if aug:
datagen = ImageDataGenerator(
rescale=1./255,
featurewise_center=False,
featurewise_std_normalization=False,
rotation_range=10,
width_shift_range=0.1,
height_shift_range=0.1,
zoom_range=0.1,
horizontal_flip=True)
else:
datagen = ImageDataGenerator(rescale=1./255)
return datagen.flow_from_directory(
dataset,
target_size=(197, 197),
color_mode='rgb',
shuffle = True,
class_mode='categorical',
batch_size=BS)
# + id="4aQAGQP5_Gpl" colab_type="code" outputId="39926d21-b2a1-4280-ba59-0e3afe7ef03e" colab={"base_uri": "https://localhost:8080/", "height": 102}
train_generator = get_datagen('/content/train', True)
dev_generator = get_datagen('/content/dev')
test_generator = get_datagen('/content/test')
# + colab_type="code" id="pLISdlaStbUn" outputId="c4f6da09-20e4-4b04-d2cb-efccdda0d88e" colab={"base_uri": "https://localhost:8080/", "height": 1000}
history = model.fit_generator(
generator = train_generator,
validation_data=dev_generator,
#steps_per_epoch=28709// BS,
#validation_steps=3509 // BS,
shuffle=True,
epochs=100,
callbacks=[rlrop],
use_multiprocessing=True,
)
# + id="JSSv08SHF0bC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="a7179cab-5f0b-4414-e049-ec5a6ce50967"
print('\n# Evaluate on dev data')
results_dev = model.evaluate_generator(dev_generator, 3509 // BS)
print('dev loss, dev acc:', results_dev)
# + id="Ev4sDYDlOsqk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="aa5ea7f8-b78d-4459-fd5f-5013490b7ecf"
print('\n# Evaluate on test data')
results_test = model.evaluate_generator(test_generator, 3509 // BS)
print('test loss, test acc:', results_test)
# + id="m9f7smhHUQus" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 590} outputId="53028197-9731-4dc2-a13e-2cfaeec79e96"
# list all data in history
print(history.history.keys())
# summarize history for accuracy
plt.plot(history.history['acc'])
plt.plot(history.history['val_acc'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'dev'], loc='upper left')
plt.show()
# summarize history for loss
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'dev'], loc='upper left')
plt.show()
# + id="F-QKHTmJQhi1" colab_type="code" colab={}
epoch_str = '-EPOCHS_' + str(EPOCHS)
test_acc = 'test_acc_%.3f' % results_test[1]
model.save('/content/drive/My Drive/cs230 project/models/final/' + 'SENET50' + epoch_str + test_acc + '.h5')
# + id="yJceV2fuFEfQ" colab_type="code" colab={}
|
models/tf-SeNet50.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:env-TM2020] *
# language: python
# name: conda-env-env-TM2020-py
# ---
# + pycharm={"name": "#%%\n"}
import os
import numpy as np
import pandas as pd
import scipy.optimize as spo
import matplotlib.pyplot as plt
# + pycharm={"name": "#%%\n"}
# baseline stock to compare our portfolio performance with
BASE_LINE = 'SPY'
# companies stocks in our portfolio
SYMBOLS = ['AAPL', 'XOM', 'IBM', 'PG']
# initial allocations
allocations = np.array([0.3, 0.2, 0.1, 0.4])
# risk free rate, percent return when amount invested in secure asset
risk_free_rate = 0.0
# sampling frequency, currently configured for daily.
# weekly: 52, monthly: 12
sampling_freq = 252
# date range for which we wish to optimize our portfolio
start_date = '2020-09-01'
end_date = '2021-05-31'
# initial investment amount
initial_investment = 100000
# + pycharm={"name": "#%%\n"}
def symbol_to_path(symbol, base_dir="data"):
return os.path.join(base_dir, f"{str(symbol)}.csv")
def get_df(data_frame, symbol, columns, jhow="left"):
path = symbol_to_path(symbol)
df_temp = pd.read_csv(path,
index_col="Date",
parse_dates=True,
usecols=columns,
na_values=["nan"])
df_temp = df_temp.rename(columns={columns[1]: symbol})
data_frame = data_frame.join(df_temp, how=jhow)
return data_frame
def get_data(symbols, dates):
data_frame = pd.DataFrame(index=dates)
if "SPY" in symbols:
symbols.pop(symbols.index("SPY"))
data_frame = get_df(data_frame, "SPY", ["Date", "Adj Close"], jhow="inner")
for s in symbols:
data_frame = get_df(data_frame, s, ["Date", "Adj Close"])
return data_frame
def plot_data(df, title="Stock prices"):
df.plot(figsize=(20, 15), fontsize=15)
plt.title(title, fontsize=30)
plt.ylabel("Price [$]", fontsize=20)
plt.xlabel("Dates", fontsize=20)
plt.legend(fontsize=20)
plt.show()
def plot_selected(df, columns, start_date, end_date):
plt_df = normalize_data(df.loc[start_date:end_date][columns])
plot_data(plt_df)
def normalize_data(df):
return df / df.iloc[0, :]
# + pycharm={"name": "#%%\n"}
# plotting cumulative performance of stocks
dates = pd.date_range(start_date, end_date)
df = get_data(SYMBOLS, dates)
plot_selected(df, SYMBOLS, start_date, end_date)
# + pycharm={"name": "#%%\n"}
# computing portfolio value based on initial allocation and invetment
price_stocks = df[SYMBOLS]
price_SPY = df[BASE_LINE]
normed_price: pd.DataFrame = price_stocks/price_stocks.values[0]
allocated = normed_price.multiply(allocations)
position_value = allocated.multiply(initial_investment)
portfolio_value = position_value.sum(axis=1)
# + pycharm={"name": "#%%\n"}
# plotting portfolio's performance before optimum allocation
port_val = portfolio_value / portfolio_value[0]
prices_SPY = price_SPY / price_SPY[0]
df_temp = pd.concat([port_val, prices_SPY], keys=['Portfolio', 'SPY'], axis=1)
plot_data(df_temp, title="Daily portfolio value and SPY (Before optimization)")
# + pycharm={"name": "#%%\n"}
def compute_daily_returns(df: pd.DataFrame) -> pd.DataFrame:
daily_returns = df.copy()
# daily_returns[1:] = (daily_returns[1:] / daily_returns[:-1].values) - 1
daily_returns = daily_returns / daily_returns.shift(1) - 1
daily_returns.iloc[0] = 0
return daily_returns
def compute_sharpe_ratio(sampling_freq: int, risk_free_rate: float, daily_return: pd.DataFrame) -> pd.DataFrame:
daily_return_std = daily_return.std()
return np.sqrt(sampling_freq) * ((daily_return.subtract(risk_free_rate)).mean()) / daily_return_std
# + pycharm={"name": "#%%\n"}
daily_return = compute_daily_returns(portfolio_value)
sharpe_ratio = compute_sharpe_ratio(sampling_freq, risk_free_rate, daily_return)
print('Sharpe Ratio (Before Optimization): ', sharpe_ratio)
# + pycharm={"name": "#%%\n"}
# function used by minimizer to find optimum allocation. Minimizes negative sharpe ratio
def f(allocations: np.array, starting_investment: float, normed_prices):
allocated = normed_prices.multiply(allocations)
position_values = allocated.multiply(starting_investment)
portfolio_value = position_values.sum(axis=1)
daily_return = (portfolio_value/portfolio_value.shift(1)) - 1
return compute_sharpe_ratio(252, 0.0, daily_return) * -1
# + pycharm={"name": "#%%\n"}
# finding optimum allocation with bounds(each stock can take value between 0 and 1) and constraints (sum of allocation must be 1)
bounds = [(0.0, 1.0) for _ in normed_price.columns]
constraints = ({'type': 'eq', 'fun': lambda inputs: 1.0 - np.sum(inputs)})
result = spo.minimize(f, allocations, args=(initial_investment, normed_price, ), method='SLSQP',
constraints=constraints, bounds=bounds, options={'disp': True})
# + pycharm={"name": "#%%\n"}
# plotting portfolio's performance after optimum allocation
opt_allocation = result.x
opt_allocated = normed_price.multiply(opt_allocation)
opt_position_value = opt_allocated.multiply(initial_investment)
opt_port_value = opt_position_value.sum(axis=1)
normed_opt_port_value = opt_port_value / opt_port_value.values[0]
plot_data(pd.concat([normed_opt_port_value, prices_SPY], keys=['Portfolio', 'SPY'], axis=1), 'Daily Portfolio and SPY values (After Optimization)')
# + pycharm={"name": "#%%\n"}
print('Optimum Allocation: ', opt_allocation)
# + pycharm={"name": "#%%\n"}
daily_return_opt = compute_daily_returns(opt_port_value)
sharpe_ratio_opt = compute_sharpe_ratio(sampling_freq, risk_free_rate, daily_return_opt)
print('Sharpe Ratio (After Optimization): ', sharpe_ratio_opt)
# + pycharm={"name": "#%%\n"}
|
optimizer.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.5 64-bit (''ai_tutorial'': venv)'
# name: python37564bitaitutorialvenvbfa9976514ab457184b1b6f4ee41b3e6
# ---
# +
import cv2
import matplotlib.pyplot as plt
img = cv2.imread('./flower.jpg')
img = cv2.resize(img, dsize=None, fx=0.5, fy=0.5)
gray = cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)
gray = cv2.GaussianBlur(gray, (7, 7), 0)
img2 = cv2.threshold(gray, 140, 240, cv2.THRESH_BINARY_INV)[1]
plt.subplot(1, 2, 1)
plt.imshow(img2, cmap='gray')
cnts = cv2.findContours(img2, cv2.RETR_LIST, cv2.CHAIN_APPROX_SIMPLE)[0]
for pt in cnts:
x, y, w, h = cv2.boundingRect(pt)
if w < 30 or w > 200:
continue
cv2.rectangle(img, (x, y), (x+w, y+h), (0, 255, 0), thickness=2)
plt.subplot(1, 2, 2)
plt.imshow(cv2.cvtColor(img, cv2.COLOR_BGR2RGB))
plt.show()
# -
|
flower.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Lab 1: Quantum Computing Operations and Algorithms
#
# <div class="youtube-wrapper">
# <iframe src="https://www.youtube.com/embed/uI02dn7PsHI" title="YouTube video player" frameborder="0" allow="accelerometer; autoplay; clipboard-write; encrypted-media; gyroscope; picture-in-picture" allowfullscreen></iframe>
# </div>
#
# - Download the notebook: <a href="/content/summer-school/2021/resources/lab-notebooks/lab-1.ipynb">[en]</a> <a href="/content/summer-school/2021/resources/lab-notebooks/lab-1-ja.ipynb">[ja]</a>
#
# In this lab, you will learn how to construct quantum states and circuits, and run a simple quantum algorithm.
#
# Quantum States and Circuits:
# * Graded Exercise 1-1: Bit Flip
# * Graded Exercise 1-2: Plus State
# * Graded Exercise 1-3: Minus State
# * Graded Exercise 1-4: Cmplex State
# * Graded Exercise 1-5: Bell State
# * Graded Exercise 1-6: GHZ-like State
#
# The Deutsch-Jozsa Algorithm:
# * Graded Exercise 1-7: Classical Deutsch-Jozsa
# * Graded Exercise 1-8: Quantum Deutsch-Jozsa
#
# ### Lab Tips / Hints
#
# * For all the exercises with gates just add the gate(s) and only gates.
# * For Ex6, look at the example with GHZ state and how it will look if you add X gate.
# * For the Ex7, the balanced aspect is related to fact that half of the output are '0' and the other one is ‘1'. So in the worst case you have to test half of possibilities plus 1.
# * For the Ex8, think of the algorithm by parts. A good help is the [Qiskit textbook chapter on D-J algorithm](https://learn.qiskit.org/course/ch-algorithms/deutsch-jozsa-algorithm#full_alg).
#
# * To test your gate, please go to IBM Composer [here](https://quantum-computing.ibm.com/composer/files/new). As per the docs, "IBM Quantum Composer is a graphical quantum programming tool that lets you drag and drop operations to build quantum circuits and run them on real quantum hardware or simulators."
# * For Ex7, the formula is mentioned in the lecture 2.1 notes, under the Deutsch-Jozsa Algorithm>Classical Solution
# * For an alternate solution to Ex6, one can use “Y” gate it performs a bit flip and phase flip on the second qubit for e.g, $|000\rangle + |111\rangle$ changes to $|010\rangle - |101\rangle$
#
# <!-- ::: q-block.reminder -->
#
# ### FAQ
#
# <details>
# <summary>Is it needed to watch the two firsts lectures to complete the lab?</summary>
# It’s recommended but not mandatory.
# </details>
#
# <!-- ::: -->
#
# ### Suggested resources
# - Watch daytonellwanger on [Introduction to Quantum Computing (14) - Quantum Circuits and Gates](https://www.youtube.com/watch?v=wLv20RHqlgw)
# - Watch sentdex on [Deutsch Jozsa Algorithm - Quantum Computer Programming w/ Qiskit p.3](https://www.youtube.com/watch?v=_BHvE_pwF6E)
# - Read Qiskit on [Single Qubit Gates](/course/ch-states/single-qubit-gates)
# - Read Qiskit on [Deutsch-Jozsa Algorithm](https://learn.qiskit.org/course/ch-algorithms/deutsch-jozsa-algorithm#full_alg )
# - Read StackExchange on [Implementing Four Bell States on IBMQ](https://quantumcomputing.stackexchange.com/questions/2258/how-to-implement-the-4-bell-states-on-the-ibm-q-composer)
# - Watch Dr. <NAME> on [Quantum Computing with Qiskit Fundamentals](https://www.youtube.com/watch?v=A2ViFq0yhLI)
# - Watch Dr. <NAME> on [Quantum Computing with Qiskit Introduction to Algorithms](https://www.youtube.com/watch?v=uuCUUW6Bu2g)
# - Read Analysis of Deutsch-Jozsa Quantum Algorithm on [Suggested Resources: Materials ](https://eprint.iacr.org/2018/249.pdf)
# - Read <NAME> al. on [Quantum Engineer’s Guide to Superconducting Qubits](https://arxiv.org/pdf/1904.06560.pdf)
# - Read <NAME>. on [Classical and Quantum Logic Gates](http://www2.optics.rochester.edu/~stroud/presentations/muthukrishnan991/LogicGates.pdf)
|
notebooks/summer-school/2021/lab1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Imports
#
# We use *io* for byte IO streaming, *cairosvg* for SVG rasterization, *pdf2image* for PDF rasterization, and *PIL* for image processing:
# +
import io
from cairosvg import svg2png
from pdf2image import convert_from_path
from PIL import Image
# -
# # Synthetic data generation
#
# First we load an example document. We use a low DPI (dots per inch), since the added detail is not expected to increase detection accuracy:
# +
pdf_file = '../data/documents/ak10900_selostus.pdf'
pdf_page = convert_from_path(pdf_file, dpi=50)
# -
# We can visualize one of the pages by simply indexing into the list:
# + tags=[]
print('Image size:', pdf_page[0].size, '(px)')
pdf_page[0]
# -
# And here's one of the pages that originally may have contained a signature:
# + tags=[]
print('Image size:', pdf_page[36].size, '(px)')
pdf_page[36]
# -
# Next we'll load the signature of a former US head of state:
# + tags=[]
signature = Image.open(io.BytesIO(svg2png(url='https://upload.wikimedia.org/wikipedia/commons/4/46/George_HW_Bush_Signature.svg')))
# Reduce the width to be more in line with document
signature = signature.resize([signature.width//4, signature.height//4])
print('Image size:', signature.size, '(px)')
signature
# -
# Finally we simply paste the signature onto the target document page at the correct location:
page = pdf_page[36].copy()
page.paste(signature, [130,360], signature)
page
|
notebooks/Synthetic data generation demo.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
from flask import Flask
from flask_sqlalchemy import SQLAlchemy
from sqlalchemy import BigInteger, Boolean, CheckConstraint, Column, DateTime, Float, ForeignKey, Integer, SmallInteger, String, Text, UniqueConstraint
from sqlalchemy.orm import relationship
app = Flask(__name__)
app.config['SQLALCHEMY_DATABASE_URI'] = 'postgresql+psycopg2://postgres@localhost/dev_logware3'
app.config['SQLALCHEMY_TRACK_MODIFICATIONS'] = False
db = SQLAlchemy(app)
conn = db.engine.connect().connection
# +
class Annotation(db.Model):
__tablename__ = 'annotations'
annotation_guid = db.Column(db.String(32), primary_key=True)
reading_guid = db.Column(db.ForeignKey('readings.reading_guid'), nullable=False)
annotation = db.Column(db.Text)
reading = db.relationship('Reading', primaryjoin='Annotation.reading_guid == Reading.reading_guid', backref='annotations')
class Asset(db.Model):
__tablename__ = 'assets'
__table_args__ = (
db.CheckConstraint("(model)::text <> ''::text"),
db.CheckConstraint("(serial)::text <> ''::text"),
db.UniqueConstraint('model', 'serial')
)
asset_guid = db.Column(db.String(32), primary_key=True)
asset_type = db.Column(db.SmallInteger, nullable=False)
model = db.Column(db.String(32), nullable=False)
serial = db.Column(db.String(32), nullable=False)
active = db.Column(db.Boolean)
deleted = db.Column(db.Boolean)
asset_password = db.Column(db.String(20))
notes = db.Column(db.Text)
class LicenseInUse(db.Model):
__tablename__ = 'license_in_use'
license_in_use_guid = db.Column(db.String(32), primary_key=True)
computer_name = db.Column(db.Text, nullable=False)
user_guid = db.Column(db.ForeignKey('users.user_guid'), nullable=False)
license_guid = db.Column(db.ForeignKey('licenses.license_guid'), nullable=False)
time_stamp = db.Column(db.DateTime, nullable=False)
license = db.relationship('License', primaryjoin='LicenseInUse.license_guid == License.license_guid', backref='license_in_uses')
user = db.relationship('User', primaryjoin='LicenseInUse.user_guid == User.user_guid', backref='license_in_uses')
class License(db.Model):
__tablename__ = 'licenses'
license_guid = db.Column(db.String(32), primary_key=True)
license_type = db.Column(db.SmallInteger, nullable=False)
license_serial = db.Column(db.String(20))
version = db.Column(db.String(20))
date_applied = db.Column(db.DateTime, nullable=False)
logins_remaining = db.Column(db.Integer)
license_id = db.Column(db.Text, nullable=False)
deleted = db.Column(db.Boolean)
class Location(db.Model):
__tablename__ = 'locations'
__table_args__ = (
db.CheckConstraint("(location_name)::text <> ''::text"),
)
location_guid = db.Column(db.String(32), primary_key=True)
location_name = db.Column(db.String(20), nullable=False, unique=True)
active = db.Column(db.Boolean)
deleted = db.Column(db.Boolean)
notes = db.Column(db.Text)
class LogSession(db.Model):
__tablename__ = 'log_sessions'
log_session_guid = db.Column(db.String(32), primary_key=True)
session_start = db.Column(db.DateTime, nullable=False)
session_end = db.Column(db.DateTime)
logging_interval = db.Column(db.Integer, nullable=False)
logger_guid = db.Column(db.ForeignKey('assets.asset_guid'), nullable=False)
user_guid = db.Column(db.ForeignKey('users.user_guid'), nullable=False)
session_type = db.Column(db.SmallInteger, nullable=False)
computer_name = db.Column(db.Text, nullable=False)
asset = db.relationship('Asset', primaryjoin='LogSession.logger_guid == Asset.asset_guid', backref='log_sessions')
user = db.relationship('User', primaryjoin='LogSession.user_guid == User.user_guid', backref='log_sessions')
class Reading(db.Model):
__tablename__ = 'readings'
reading_guid = db.Column(db.String(32), primary_key=True)
reading = db.Column(db.Float(53), nullable=False)
reading_type = db.Column(db.SmallInteger, nullable=False)
time_stamp = db.Column(db.DateTime, nullable=False)
log_session_guid = db.Column(db.ForeignKey('log_sessions.log_session_guid'), nullable=False)
sensor_guid = db.Column(db.ForeignKey('assets.asset_guid'), nullable=False)
location_guid = db.Column(db.ForeignKey('locations.location_guid'), nullable=False)
channel = db.Column(db.SmallInteger, nullable=False)
max_alarm = db.Column(db.Boolean)
max_alarm_value = db.Column(db.Float(53))
min_alarm = db.Column(db.Boolean)
min_alarm_value = db.Column(db.Float(53))
compromised = db.Column(db.Boolean)
location = db.relationship('Location', primaryjoin='Reading.location_guid == Location.location_guid', backref='readings')
log_session = db.relationship('LogSession', primaryjoin='Reading.log_session_guid == LogSession.log_session_guid', backref='readings')
asset = db.relationship('Asset', primaryjoin='Reading.sensor_guid == Asset.asset_guid', backref='readings')
class SensorParameter(db.Model):
__tablename__ = 'sensor_parameters'
log_session_guid = db.Column(db.ForeignKey('log_sessions.log_session_guid'), primary_key=True, nullable=False)
channel = db.Column(db.SmallInteger, primary_key=True, nullable=False)
parameter_name = db.Column(db.String(128), primary_key=True, nullable=False)
parameter_value = db.Column(db.String(128), nullable=False)
log_session = db.relationship('LogSession', primaryjoin='SensorParameter.log_session_guid == LogSession.log_session_guid', backref='sensor_parameters')
class User(db.Model):
__tablename__ = 'users'
__table_args__ = (
db.CheckConstraint("(login_name)::text <> ''::text"),
)
user_guid = db.Column(db.String(32), primary_key=True)
login_name = db.Column(db.String(32), nullable=False, unique=True)
first_name = db.Column(db.String(64))
last_name = db.Column(db.String(64))
user_password = db.Column(db.String(64))
user_group = db.Column(db.SmallInteger)
permissions = db.Column(db.BigInteger)
active = db.Column(db.Boolean)
deleted = db.Column(db.Boolean)
change = db.Column(db.Boolean)
notes = db.Column(db.Text)
class Version(db.Model):
__tablename__ = 'versions'
db_version = db.Column(db.String(20), primary_key=True)
client_version = db.Column(db.String(20))
# -
records = d SELECT readings.reading_guid, readings.reading , readings.reading_type , readings.time_stamp , locations.location_name , readings.sensor_guid , readings.location_guid AS readings_location_guid, readings.channel AS readings_channel, readings.max_alarm AS readings_max_alarm, readings.max_alarm_value AS readings_max_alarm_value, readings.min_alarm AS readings_min_alarm, readings.min_alarm_value AS readings_min_alarm_value, readings.compromised AS readings_compromised
b.session.query(Reading).join(Reading.location).filter_by(location_name='ONSITE1').filter(Reading.time_stamp.between('2017-03-26', '2017-03-28'))
criteria = {"location_name_1": 'ONSITE1', "time_stamp_1":'2017-01-26', "time_stamp_2":'2017-03-28'}
df = pd.read_sql(('''
SELECT readings.reading_guid, readings.reading , readings.reading_type , readings.time_stamp , locations.location_name , readings.sensor_guid , readings.location_guid AS readings_location_guid, readings.channel AS readings_channel, readings.max_alarm AS readings_max_alarm, readings.max_alarm_value AS readings_max_alarm_value, readings.min_alarm AS readings_min_alarm, readings.min_alarm_value AS readings_min_alarm_value, readings.compromised AS readings_compromised
FROM readings JOIN locations ON readings.location_guid = locations.location_guid
WHERE locations.location_name = %(location_name_1)s AND readings.time_stamp BETWEEN %(time_stamp_1)s AND %(time_stamp_2)s'''),
conn, params=criteria)
df.info()
df
def celsius_to_fahr(temp_celsius):
"""Convert Fahrenheit to Celsius
Return Celsius conversion of input"""
temp_fahr = (temp_celsius * 1.8) + 32
return temp_fahr
# convert temps to fahrenheit
df.loc[df['reading_type'] == 0, 'reading'] = df.reading.apply(celsius_to_fahr)
df
locs = [loc for loc, in db.session.query(Location.location_name)]
summary = pd.DataFrame(index=None, columns=['LOCATION', 'SPECIFICATION', 'START_DATE', 'END_DATE', 'FIRST_POINT_RECORDED', 'LAST_POINT_RECORDED', 'TOTAL_HOURS_EVALUATED', 'TOTAL_HOURS_RECORDED', 'TOTAL_HOURS_OUT', 'PERCENT_OUT', 'HOURS_TEMP_HIGH', 'HOURS_TEMP_LOW', 'HOURS_RH_HIGH', 'HOURS_RH_LOW', 'HOURS_OVERLAP', 'HOURS_NO_DATA', 'INT_GREATER_THAN_15', 'HRS_DOWN_FOR_MAINT', 'DUPE_RECORDS'])
summary
df.location_name.unique()
summary.LOCATION = df.location_name.unique()
summary
temps = df[df['reading_type']==0]
temps.dtypes
temps = temps.set_index('time_stamp')
temps['duration'] = temps.index.to_series().diff().dt.seconds.div(60, fill_value=0)
temps.describe()
temp_hi = temps[temps.reading > 72]
temp_low = temps[temps.reading < 69]
temp_low.describe()
t_hr_hi = temp_hi.duration.sum(axis=0) / 60
t_hr_hi
t_hr_low = temp_low.duration.sum(axis=0) / 60
t_hr_low
t_gap = temps[temps.duration > 15]
t_gap_hrs = t_gap.duration.sum(axis=0) / 60
t_gap
t_gap_hrs
# df.loc[df['line_race'] == 0, 'rating'] = 0
summary.loc[summary.LOCATION == criteria.get('location_name_1'), 'START_DATE'] = pd.to_datetime(criteria.get('time_stamp_1'))
summary.loc[summary.LOCATION == criteria.get('location_name_1'), 'END_DATE'] = pd.to_datetime(criteria.get('time_stamp_2'))
summary.loc[summary.LOCATION == criteria.get('location_name_1'), 'FIRST_POINT_RECORDED'] = df.time_stamp.min()
summary.loc[summary.LOCATION == criteria.get('location_name_1'), 'LAST_POINT_RECORDED'] = df.time_stamp.max()
summary.loc[summary.LOCATION == criteria.get('location_name_1'), 'TOTAL_HOURS_EVALUATED'] = (summary.END_DATE - summary.START_DATE).astype('timedelta64[s]') / 3600.0
summary.loc[summary.LOCATION == criteria.get('location_name_1'), 'TOTAL_HOURS_RECORDED'] = (summary.LAST_POINT_RECORDED - summary.FIRST_POINT_RECORDED).astype('timedelta64[s]') / 3600.0
summary.loc[summary.LOCATION == criteria.get('location_name_1'), 'HOURS_TEMP_HIGH'] = temp_hi.duration.sum(axis=0) / 60
summary.loc[summary.LOCATION == criteria.get('location_name_1'), 'HOURS_TEMP_LOW'] = temp_low.duration.sum(axis=0) / 60
summary.loc[summary.LOCATION == criteria.get('location_name_1'), 'HOURS_RH_HIGH'] = rh_hi.duration.sum(axis=0) / 60
summary.loc[summary.LOCATION == criteria.get('location_name_1'), 'HOURS_RH_LOW'] = rh_low.duration.sum(axis=0) / 60
summary.loc[summary.LOCATION == criteria.get('location_name_1'), 'TOTAL_HOURS_RECORDED'] = (((summary.LAST_POINT_RECORDED - summary.FIRST_POINT_RECORDED).astype('timedelta64[s]') / 3600.0) -(t_gap_hrs / 60))
summary
rh = df[df['reading_type']==1]
rh.dtypes
rh = rh.set_index('time_stamp')
rh['duration'] = rh.index.to_series().diff().dt.seconds.div(60, fill_value=0)
rh.describe()
rh_hi = rh[rh.reading > 29.5]
rh_low = rh[rh.reading < 25.5]
rh_low.describe()
rh_hr_hi = rh_hi.duration.sum(axis=0) / 60
rh_hr_hi
rh_hr_low = rh_low.duration.sum(axis=0) / 60
rh_hr_low
rh_gap = rh[rh.duration > 15]
rh_gap_hrs = rh_gap.duration.sum(axis=0)
rh_gap
rh_gap_hrs
a = ((summary.LAST_POINT_RECORDED - summary.FIRST_POINT_RECORDED).astype('timedelta64[s]') / 3600.0)
a - t_gap_hrs
pd.concat([t_gap, rh_gap], keys=['reading_type', 'time_stamp'])
len(pd.merge(t_gap, rh_gap, left_index=True, right_index=True))
# +
# df.loc[df['line_race'] == 0, 'rating'] = 0
summary.loc[summary.LOCATION == criteria.get('location_name_1'), 'START_DATE'] = pd.to_datetime(criteria.get('time_stamp_1'))
summary.loc[summary.LOCATION == criteria.get('location_name_1'), 'END_DATE'] = pd.to_datetime(criteria.get('time_stamp_2'))
summary.loc[summary.LOCATION == criteria.get('location_name_1'), 'FIRST_POINT_RECORDED'] = df.time_stamp.min()
summary.loc[summary.LOCATION == criteria.get('location_name_1'), 'LAST_POINT_RECORDED'] = df.time_stamp.max()
summary.loc[summary.LOCATION == criteria.get('location_name_1'), 'TOTAL_HOURS_EVALUATED'] = (summary.END_DATE - summary.START_DATE).astype('timedelta64[s]') / 3600.0
summary.loc[summary.LOCATION == criteria.get('location_name_1'), 'HOURS_TEMP_HIGH'] = temp_hi.duration.sum(axis=0) / 60
summary.loc[summary.LOCATION == criteria.get('location_name_1'), 'HOURS_TEMP_LOW'] = temp_low.duration.sum(axis=0) / 60
summary.loc[summary.LOCATION == criteria.get('location_name_1'), 'TOTAL_HOURS_RECORDED'] = ((summary.LAST_POINT_RECORDED - summary.FIRST_POINT_RECORDED).astype('timedelta64[s]') / 3600.0) - t_gap_hrs
summary.loc[summary.LOCATION == criteria.get('location_name_1'), 'HOURS_NO_DATA'] = t_gap_hrs
summary.loc[summary.LOCATION == criteria.get('location_name_1'), 'INT_GREATER_THAN_15'] = len(pd.merge(t_gap, rh_gap, left_index=True, right_index=True))
summary.loc[summary.LOCATION == criteria.get('location_name_1'), 'TOTAL_HOURS_OUT'] = summary[['HOURS_TEMP_HIGH', 'HOURS_TEMP_LOW', 'HOURS_RH_HIGH', 'HOURS_RH_LOW']].sum(axis=1)
summary
# -
criteria.get('location_name_1')
|
notebooks/exploratory/0.4-jlawton-logware3-exploration-with-flask-sqlalchemy.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import keras
import tensorflow as tf
print(keras.__version__)
print(tf.__version__)
# +
import numpy as np
import pandas as pd
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.model_selection import train_test_split
from sklearn.metrics import classification_report,confusion_matrix
NGRAMS = 2
EPOCHS = 25
# Wikilabels
df = pd.read_csv('../data/wiki/wiki_name_race.csv')
df.dropna(subset=['name_first', 'name_last'], inplace=True)
sdf = df
# Additional features
sdf['name_first'] = sdf.name_first.str.title()
sdf.groupby('race').agg({'name_first': 'count'})
# -
# ## Preprocessing the input data
# +
# only last name will be use to train the model
sdf['name_last_name_first'] = sdf['name_last']
# build n-gram list
vect = CountVectorizer(analyzer='char', max_df=0.3, min_df=3, ngram_range=(NGRAMS, NGRAMS), lowercase=False)
a = vect.fit_transform(sdf.name_last_name_first)
vocab = vect.vocabulary_
# sort n-gram by freq (highest -> lowest)
words = []
for b in vocab:
c = vocab[b]
#print(b, c, a[:, c].sum())
words.append((a[:, c].sum(), b))
#break
words = sorted(words, reverse=True)
words_list = ['UNK']
words_list.extend([w[1] for w in words])
num_words = len(words_list)
print("num_words = %d" % num_words)
def find_ngrams(text, n):
a = zip(*[text[i:] for i in range(n)])
wi = []
for i in a:
w = ''.join(i)
try:
idx = words_list.index(w)
except:
idx = 0
wi.append(idx)
return wi
# build X from index of n-gram sequence
X = np.array(sdf.name_last_name_first.apply(lambda c: find_ngrams(c, NGRAMS)))
# check max/avg feature
X_len = []
for x in X:
X_len.append(len(x))
max_feature_len = max(X_len)
avg_feature_len = int(np.mean(X_len))
print("Max feature len = %d, Avg. feature len = %d" % (max_feature_len, avg_feature_len))
y = np.array(sdf.race.astype('category').cat.codes)
# Split train and test dataset
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=21, stratify=y)
# -
# ## Train a LSTM model
#
# ref: http://machinelearningmastery.com/sequence-classification-lstm-recurrent-neural-networks-python-keras/
# +
'''The dataset is actually too small for LSTM to be of any advantage
compared to simpler, much faster methods such as TF-IDF + LogReg.
Notes:
- RNNs are tricky. Choice of batch size is important,
choice of loss and optimizer is critical, etc.
Some configurations won't converge.
- LSTM loss decrease patterns during training can be quite different
from what you see with CNNs/MLPs/etc.
'''
from keras.preprocessing import sequence
from keras.models import Sequential
from keras.layers import Dense, Embedding, Dropout, Activation
from keras.layers import LSTM
from keras.layers.convolutional import Conv1D
from keras.layers.convolutional import MaxPooling1D
from keras.models import load_model
max_features = num_words # 20000
feature_len = 20 # avg_feature_len # cut texts after this number of words (among top max_features most common words)
batch_size = 32
print(len(X_train), 'train sequences')
print(len(X_test), 'test sequences')
print('Pad sequences (samples x time)')
X_train = sequence.pad_sequences(X_train, maxlen=feature_len)
X_test = sequence.pad_sequences(X_test, maxlen=feature_len)
print('X_train shape:', X_train.shape)
print('X_test shape:', X_test.shape)
num_classes = np.max(y_train) + 1
print(num_classes, 'classes')
print('Convert class vector to binary class matrix '
'(for use with categorical_crossentropy)')
y_train = tf.keras.utils.to_categorical(y_train, num_classes)
y_test = tf.keras.utils.to_categorical(y_test, num_classes)
print('y_train shape:', y_train.shape)
print('y_test shape:', y_test.shape)
# +
print('Build model...')
model = Sequential()
model.add(Embedding(num_words, 32, input_length=feature_len))
model.add(LSTM(128, dropout=0.2, recurrent_dropout=0.2))
model.add(Dense(num_classes, activation='softmax'))
# try using different optimizers and different optimizer configs
model.compile(loss='categorical_crossentropy',
optimizer='adam',
metrics=['accuracy'])
print(model.summary())
# -
print('Train...')
model.fit(X_train, y_train, batch_size=batch_size, epochs=EPOCHS,
validation_split=0.1, verbose=1)
score, acc = model.evaluate(X_test, y_test,
batch_size=batch_size, verbose=1)
print('Test score:', score)
print('Test accuracy:', acc)
# ## Confusion Matrix
p = model.predict(X_test, verbose=2) # to predict probability
y_pred = np.argmax(p, axis=-1)
target_names = list(sdf.race.astype('category').cat.categories)
print(classification_report(np.argmax(y_test, axis=1), y_pred, target_names=target_names))
print(confusion_matrix(np.argmax(y_test, axis=1), y_pred))
# ## Save model
model.save('./wiki/lstm/wiki_ln_lstm.h5')
words_df = pd.DataFrame(words_list, columns=['vocab'])
words_df.to_csv('./wiki/lstm/wiki_ln_vocab.csv', index=False, encoding='utf-8')
sdf.groupby('race').agg({'name_first': 'count'}).to_csv('./wiki/lstm/wiki_race.csv', columns=[])
|
ethnicolr/models/ethnicolr_keras_lstm_wiki_ln.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="w_6GmnEWZntb"
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
import matplotlib.pyplot as plt #Plotting
# %matplotlib inline
import warnings #What to do with warnings
warnings.filterwarnings("ignore") #Ignore the warnings
plt.rcParams["figure.figsize"] = (10,10) #Make the plots bigger by default
plt.rcParams["lines.linewidth"] = 2 #Setting the default line width
plt.style.use("ggplot") #Define the style of the plot
from statsmodels.tsa.seasonal import seasonal_decompose #Describes the time data
from statsmodels.tsa.stattools import adfuller #Check if data is stationary
from statsmodels.graphics.tsaplots import plot_acf #Compute lag for ARIMA
from statsmodels.graphics.tsaplots import plot_pacf #Compute partial lag for ARIMA
from statsmodels.tsa.arima_model import ARIMA #Predictions and Forecasting
# + id="Gr5EhDRKZt3a" outputId="4ccd6c6f-0fe8-49f2-e1b1-2d9abb63be78"
amazon = pd.read_csv("../input/amazonstocks/AMZN (1).csv") #Get our stock data from the CSV
amazon.head(10) #Take a peek at the Amazon data
# + id="zBBHnd46Zt6u" outputId="2bc19821-82e5-4709-f3c5-68cc205d425f"
print(amazon.isnull().any()) #Check for null values
# + id="Km0Bm2RiZt9s" outputId="8f0f22a6-3709-49ae-afd5-9d44522be758"
amazonOpen = amazon[["Date", "Open"]].copy() #Get the date and open columns
amazonOpen["Date"] = pd.to_datetime(amazonOpen["Date"]) #Ensure the date data is in datetime format
amazonOpen.set_index("Date", inplace = True) #Set the date to the index
amazonOpen = amazonOpen.asfreq("b") #Set the frequency
amazonOpen = amazonOpen.fillna(method = "bfill") #Fill null values with future values
#amazonOpen.index #Make sure the frequency remains intact
amazonOpen.head(12) #Take a peek at the open data
# + id="pPrdpiNfZuAu" outputId="e75000a9-3842-44c2-9403-c80bae49dd85"
y = amazonOpen.plot(title = "Amazon Stocks (Open)") #Get an idea of the data
y.set(ylabel = "Price at Open") #Set the y label to open
plt.show() #Show the plot
# + id="2gwW04PrZuDt" outputId="ccf71d28-5fcd-404a-f933-ffcf6479399b"
decomp = seasonal_decompose(amazonOpen, model = "multiplicative") #Decompose the data
x = decomp.plot() #Plot the decomposed data
# + id="7KpDogq6ZuG-" outputId="d4bbeeef-0718-42bb-b683-47b8246df2c8"
print("ADFuller Test; Significance: 0.05") #Print the significance level
adf = adfuller(amazonOpen["Open"]) #Call adfuller to test
print("ADF test static is {}".format(adf[1])) #Print the adfuller results
# + id="5Z1Anpc_ZuKu" outputId="ec27294e-298f-4759-aeff-3d01a88d8e23"
openLog = np.log(amazonOpen) #Take the log of the set for normalization
openStationary = openLog - openLog.shift() #Get a stationary set by subtracting the shifted set
openStationary = openStationary.dropna() #Drop generated null values from the set
openStationary.plot(title = "Stationary Amazon Stocks") #Plot the stationary set
# + id="g-EEcTxzZuNt" outputId="88d1c3ea-24f7-4897-ac31-acc7812d565d"
print("ADFuller Test; Significance: 0.05") #Print the significance level
adf = adfuller(openStationary["Open"]) #Call adfuller to test
print("ADF test static is {}".format(adf[1])) #Print the adfuller results
# + id="y5oUqCq2ZuQx" outputId="a55405b1-7956-4223-89ae-38f64adda97a"
decomp = seasonal_decompose(openStationary) #Decompose the stationary data
x = decomp.plot() #Plot the decomposition
# + id="601me8pDZuUC" outputId="66145dc1-564b-4544-87f9-d7beca4deace"
fig,axes = plt.subplots(2,2) #Set a subset for the data visualizations
a = axes[0,0].plot(amazonOpen["Open"]) #Plot the original data
a = axes[0,0].set_title("Original Data") #Give the original data a name
b = plot_acf(amazonOpen["Open"],ax=axes[0,1]) #Plot the ACF of the original data
x = axes[1,0].plot(openStationary["Open"]) #Plot the stationary data
x = axes[1,0].set_title("Stationary Data") #Give the stationary data a name
y = plot_acf(openStationary["Open"],ax=axes[1,1]) #Plot the ACF of the stationary data
# + id="KpcEJo2AZuW9" outputId="8a84b085-f71f-4771-ffe0-f65b60b52d25"
fig,axes = plt.subplots(1,2) #Create a subplot for the Partial ACF
a = axes[0].plot(openStationary["Open"]) #Plot the stationary data
a = axes[0].set_title("Stationary") #Ensure the stationary data is named
b = plot_pacf(openStationary["Open"], ax = axes[1], method = "ols") #Plot the partial ACF
# + id="s2QPQw11dEgU" outputId="fcf8ca2e-b7d4-4e76-dc74-e863e2ee78a4"
model = ARIMA(openStationary, order = (5, 1, 5)) #Build the ARIMA model
fitModel = model.fit(disp = 1) #Fit the ARIMA model
# + id="eD0btK0adEjo" outputId="1901e11a-91b4-4797-9213-1e40e7ae1650"
plt.rcParams.update({"figure.figsize" : (12,6), "lines.linewidth" : 0.05, "figure.dpi" : 100}) #Fix the look of the graph, dimming it to show the red
x = fitModel.plot_predict(dynamic = False) #Fit the ARIMA model
x = plt.title("Forecast Fitting") #Add a stock title
plt.show() #Show the ARIMA plot
# + id="2iBlXULxdEtD" outputId="19cf55f6-d7da-42e5-ef44-c4065b863aa4"
plt.rcParams.update({"figure.figsize" : (12,5), "lines.linewidth": 2}) #Fix the line width
length = int((len(amazonOpen)*975)/1000) #Get 975/1000 of the length of the data
print(length) #Print the length to make sure it actually is an int
# + id="t2BA-WGPdEwC"
train = amazonOpen[:length] #Use 975/1000 of the data for the train set
test = amazonOpen[length:] #Use the rest for testing
modelValid = ARIMA(train,order=(5,1,5)) #Create a model for the train set
fitModelValid = modelValid.fit(disp= -1) #Fit the model
# + id="6W7-nNH-dEzC"
fc,se,conf = fitModelValid.forecast(len(amazonOpen) - length) #Forcast over the test area
forecast = pd.Series(fc, index = test.index) #Get the forecast for the area
# + id="Kjm9SfzJdncG" outputId="e41f8db4-7f5c-45fe-aab8-006d348deef1"
#Add labels for the train, test, and forecast
plt.plot(train,label = "Training Data")
plt.plot(test,label = "Actual Continuation")
plt.plot(forecast,label = "Forecasted Continuation", color = "g")
plt.title("ARIMA Forecast") #Add the Forecast title
plt.legend(loc = "upper left") #Put the legend in the top left
plt.xlabel("Year") #Add the year label to the bottom
plt.ylabel("Open Price") #Add the open price to the y axis
# + id="QTOTgC3LdnfN" outputId="53fcfc62-9eec-4708-d59b-914fea97c658"
modelPred = ARIMA(amazonOpen,order=(5,1,5)) #Create a model for the whole data
fitModelPred = modelPred.fit(disp= -1) #Fit the model
# + id="wIXOY-WCdnjA" outputId="7c90f9c9-366d-4fbc-dc95-0de6d78cc4e9"
fitModelPred.plot_predict(1,len(amazonOpen) + 500) #Plot predictions for the next 500 days
x = fitModelPred.forecast(500) #Forecast the prediction for the next 500 days.
x = plt.title("Amazon Stock Forecast") #Add a stock title
x = plt.xlabel("Year") #Add the year label to the bottom
x = plt.ylabel("Open Price") #Add the open price to the y axis
|
Amazon_stock_prediction_model.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Logistic Regression
#
# Logistic regression is a statistical method for predicting binary outcomes from data.
#
# Examples of this are "yes" vs. "no" or "young" vs. "old".
#
# These are categories that translate to a probability of being a 0 or a 1.
#
# Source: [Logistic Regression](https://towardsdatascience.com/real-world-implementation-of-logistic-regression-5136cefb8125)
# + [markdown] slideshow={"slide_type": "subslide"}
# We can calculate the logistic regression by applying an activation function as the final step to our linear model.
#
# This converts the linear regression output to a probability.
# + slideshow={"slide_type": "subslide"}
# %matplotlib inline
import matplotlib.pyplot as plt
import pandas as pd
# + [markdown] slideshow={"slide_type": "subslide"}
# Generate some data
# + slideshow={"slide_type": "fragment"}
from sklearn.datasets import make_blobs
X, y = make_blobs(centers=2, random_state=42)
print(f"Labels: {y[:10]}")
print(f"Data: {X[:10]}")
# + slideshow={"slide_type": "subslide"}
# Visualizing both classes
plt.scatter(X[:, 0], X[:, 1], c=y)
# + [markdown] slideshow={"slide_type": "subslide"}
# Split our data into training and testing data
# + slideshow={"slide_type": "fragment"}
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=1)
# + [markdown] slideshow={"slide_type": "subslide"}
# Create a logistic regression model
# + slideshow={"slide_type": "fragment"}
from sklearn.linear_model import LogisticRegression
classifier = LogisticRegression()
classifier
# + [markdown] slideshow={"slide_type": "subslide"}
# Fit (train) our model by using the training data
# + slideshow={"slide_type": "fragment"}
classifier.fit(X_train, y_train)
# + [markdown] slideshow={"slide_type": "subslide"}
# Validate the model by using the test data
# + slideshow={"slide_type": "fragment"}
print(f"Training Data Score: {classifier.score(X_train, y_train)}")
print(f"Testing Data Score: {classifier.score(X_test, y_test)}")
# + [markdown] slideshow={"slide_type": "subslide"}
# Make predictions
# -
# Generate a new data point (the red circle)
import numpy as np
new_data = np.array([[-2, 6]])
plt.scatter(X[:, 0], X[:, 1], c=y)
plt.scatter(new_data[0, 0], new_data[0, 1], c="r", marker="o", s=100)
# + slideshow={"slide_type": "fragment"}
# Predict the class (purple or yellow) of the new data point
predictions = classifier.predict(new_data)
print("Classes are either 0 (purple) or 1 (yellow)")
print(f"The new point was classified as: {predictions}")
# + slideshow={"slide_type": "subslide"}
predictions = classifier.predict(X_test)
pd.DataFrame({"Prediction": predictions, "Actual": y_test})
|
01-Lesson-Plans/19-Supervised-Machine-Learning/1/Activities/05-Ins_Logistic_Regression/Solved/Ins_Logistic_Regression.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Analysis
# language: python
# name: ana
# ---
# # The Seats-Votes Curve in a Single Election
#
# *<NAME><br>
# University of Bristol<br>
# <EMAIL><br>
# (Supported by NSF #1657689)*
# In the last notebook, we talked about the concept underlying *historical* seats-votes relationships. At its core, though, estimating seats-votes relationships in this manner is
#
# 1. **wasteful**: in each election, we observe a ton of district-level information. However, we disregard nearly all of this information about district-level *vote* results in favor of exclusive examination of the *district* winners. While winning a district or not *is* information about district-level vote (it's a censored variable with a threshold of $.5$), this is not the same as the original information in the vote shares.
# 2. **dubious**: over many elections, the seats-votes model does *not* account for structural changes to the electorate, the party system, or the electoral process. This means that each election is considered comparable to every other election, even when they're separated by *large* expanses of time or structural changes to the electoral system.
#
# These kinds of objections are noted by Browning & King (1987) but are most forcefully discussed by Gelman & King (1994). Incorporating older arguments (e.g. Mackerras (1962), among others), this line of argument seeks to define seats votes curves *in a single election* using a basic counterfactual argument: when shifts in party popular vote occur, these votes tend to be distributed uniformly at random among districts.
#
# In this notebook, we'll talk about how this kind of seats-votes curve is built within a single election, how the *uniform partisan swing* assumption works, and how it might not quite capture the true reality of how votes change from election to election. But, first, though, we'll have to read the data again.
import pandas
import pysal
import numpy
import seaborn
import seatsvotes
import statsmodels.api as sm
import matplotlib.pyplot as plt
# %matplotlib inline
house = seatsvotes.data.congress(geo=True)
house.crs = {"init":"epsg:4269"}
house = house.to_crs(epsg=5070)
house.head()
# # Seats & Votes in a single election
# At its core, analyzing a seats-votes relationship in a single election is somewhat suspect. This is because the relationship between votes and seats is a *strict hinge-point* relationship in most two-party elections. Once the reference party gets beyond 50% in the two-party vote (i.e. once Dems get more votes than Republicans), the district is "won," and represented by a $1$. In the other direction, the district is lost. This makes for a pretty boring scatterplot:
plt.scatter(house.vote_share, house.vote_share > .5, marker='.', color='r')
plt.vlines(.5, 0,1, linestyle=':', color='k', )
# Indeed, there's no real mechanism to be learned here; the basic relationship is simply one of a censored variable; when Dems win a majority of the two-party vote, they win the seat; when Dems win a minority, Republicans win.
# ### Uniform Partisan Swing
# So, how do we model the seats-votes curve in a single election? We can't do it *directly* based on the split of districts that are won (or lost). Instead, a classic method to construct the seats-votes curve relies on the assumption of *uniform partisan swing*, that changes in a party's popular vote are reasonably modelled by assigning those changes to *districts* uniformly. So, if Dems change in their popular vote by 10%, then we simply subtract 10% of each districts' vote shares.
#
# Unfortunately, this returns us to the same issue noted before about the difference between *average district voteshare* and *party popular vote*. This distinction leads to some rather finnicky issues with assuming uniform partisan swing. For example, a change of 10% in a district with *many* voters will result in a larger bump to the popular vote than a 10% change to a small district. Thus, it becomes more common to examine a party's average voteshare across districts, rather than the party's popular vote. Further, it again becomes common to assume that, when vote shares are expected to go below $0$ or above $100%$, the vote shares stop changing for that district. Complexities remain about *uncontested* districts, and we'll treat those later.
#
# Thinking about a *seats-votes* curve estimated using uniform partisan swing, we can build one in a pretty direct fashion. We'll focus first on building the seats-votes curve under uniform swing around a point we *did observe:* a single election. Let's pick out an election in particular, the 2006 election:
house06 = house.query('year == 2006')
# Since this is only a single election, we only get one single observation of the average Democrat vote share, and one observation of how many districts the Dems won in that election. Below, we'll plot this as a single point, marked as a red `x`:
f = plt.figure()
ax = plt.gca()
ax.axis((0,1,0,1))
ax.plot(house06.vote_share.mean(), (house06.vote_share > .5).mean(), marker='x', color='r')
ax.set_xlabel("Average District Pct. Dem Vote")
ax.set_ylabel("Pct. of Congress Won by Dems")
plt.show()
# Now, uniform partisan swing basically works by asking:
#
# > if a party were to win $\delta$% more in each district, how many districts that party win?
#
# By asking that question for a few different values of $\delta$, we can build up a sequence of hypothetical district average vote shares which show the underlying seat-vote relationship latent in our data, *assuming (of course)* that the uniform swing hypothesis holds as a reasonable model of how votes in districts change as the average district vote changes.
#
# So, let's see what happens when Democrats win 1% more in each district. Below, we'll add `.01` to each observation, and record the number of seats Dems would have won if each district increases in Democrat vote share by exactly one percentage point:
f = plt.figure()
ax = plt.gca()
ax.axis((0,1,0,1))
ax.plot(house06.vote_share.mean(),
(house06.vote_share > .5).mean(), marker='x', color='r')
ax.plot((house06.vote_share + .01).mean(),
((house06.vote_share + .01) > .5).mean(), marker='x', color='k')
ax.set_xlabel("Average District Pct. Dem Vote")
ax.set_ylabel("Pct. of Congress Won by Dems")
plt.show()
# Wow! there's a very small increase in the share of seats Dems win when every district's voteshare increases by 1%. This suggests that one or more districts $i$ have a vote share $v_i$ such that $v_i < .5$ but $v_i + .01 > .5$. Thus, when all districts increase in their Democrat vote share by a single percentage point, the districts flip from being won by Republicans to being won by Democrats.
#
# Given that we can do this with an increase towards *Democrats*, we can also *decrease* all Democrat vote shares by a single percentage point and model a uniform partisan swing towards Republicans:
f = plt.figure()
ax = plt.gca()
ax.axis((0,1,0,1))
ax.plot(house06.vote_share.mean(),
(house06.vote_share > .5).mean(), marker='x', color='r')
ax.plot((house06.vote_share + .01).mean(),
((house06.vote_share + .01) > .5).mean(), marker='x', color='grey')
ax.plot((house06.vote_share - .01).mean(),
((house06.vote_share - .01) > .5).mean(), marker='x', color='k')
ax.set_xlabel("Average District Pct. Dem Vote")
ax.set_ylabel("Pct. of Congress Won by Dems")
plt.show()
# More districts flip! This suggests that some district is won by Democrats with a margin smaller than 1%.
#
# Again, contingent on the uniform partisan swing assumption being realistic, we can do this over and over again for different values of $\delta$ and obtain the "seats-votes curve" for that election:
f = plt.figure()
ax = plt.gca()
ax.axis((0,1,0,1))
ax.plot((house06.vote_share.mean() + numpy.linspace(-.25, .25, num=20)),
[((house06.vote_share + delta) > .5).mean() for delta
in numpy.linspace(-.25,.25,num=20)], marker='x', color='grey')
ax.plot(house06.vote_share.mean(),
(house06.vote_share > .5).mean(), marker='x', color='r')
ax.set_xlabel("Average District Pct. Dem Vote")
ax.set_ylabel("Pct. of Congress Won by Dems")
plt.show()
# The curve above is the "empirical" seats votes curve, like that implemented by <NAME> in the `pscl` library in R. This "empirical" seats votes curve requires the uniform partisan swing assumption to construct; each point on the curve is given by some swing $\delta$ away from the observed average party vote share $\bar{v}$ and the fraction of districts at swing $\delta$ that are won by Democrats $(v_i + \delta > .5)$. Letting $\mathcal{I}$ stand for the indicator function which is 1 when the statement inside of it is true and zero otherwise, this means every point on the empirical seats-votes curve is a coordinate:
#
# $$ \left(\bar{v}, \sum_i^N \mathcal{I}\left( (v_i + \delta) > .5\right)n^{-1}\right)$$
#
# And the functional form of the "empirical" seats-votes curve is:
#
# $$f(\delta) = \sum_i^N \mathcal{I}\left((v_i + \delta) > .5\right)n^{-1} $$
#
# Looking at this definition, a few things become apparent. First, this method practically ignores districts that go way above $1$ or below $0$; while these are impossible (no district can have more Republican votes than there are voters in general), the seats-votes curve only considers districts relative to the threshold of victory, $.5$. This means that some vote shares from $v_i + \delta$ may actually be invalid vote shares. Second, while many doubt whether or not this model is a *realistic* model of how the electorate behaves, it's still suggested as a good first approximation for how the electoral system actually works, especially in the small region around the observed voteshare (e.g. Jackman 2014). There is absolutely *no* uncertainty in this model of the seats-votes curve, however. Further, assuming that the average swing affects all districts equally can be patently unrealistic when thinking about elections as social or geographical processes. Finally, we can see that the empirical seats-votes curve is actually *much* bumpier than the plot above suggests: there is a finite number ($N$) of observed $v_i$, but $\delta$ changes continuously. This means that the $f(\delta)$ function changes in a stepwise fashion: as $\delta$ increases, some $v_i + \delta$ switches from being below $.5$ to being above $.5$; when this happens, the function increases by a single step $1/n$. In general, if districts are allowed to have *idential* $v_i$, the changes may only occur in multiples of $1/n$, but these remain *integral* changes.
#
# Indeed, this step change property belies a much more important fact: this "model" of the seats-votes curve built from this restrictive "uniform partisan swing" assumption is actually a much more basic expression of the empirical structure of the data. Nagle (2018) provides a very good discussion of this; the seats-votes curve (when constructed this way) contains the same information as the empirical cumulative distribution function for $v_i$.
#
# This means that no theory of partisan swing is necessary to construct this curve at all. We'll discuss this now.
# # Seats-Votes Curves as an Empirical CDF
# Going back to our first observed point from the 2006 election, we'll zoom in really tight on the observed result.
f = plt.figure()
ax = plt.gca()
ax.axis((0,1,0,1))
ax.plot(house06.vote_share.mean(), (house06.vote_share > .5).mean(), marker='x', color='r')
ax.axis((.55,.58, .52,.55))
ax.set_xlabel("Average District Pct. Dem Vote")
ax.set_ylabel("Pct. of Congress Won by Dems")
plt.show()
# Now, we've discussed that *really small* changes in $\delta$ should reveal the integrality property. So, instead of changing $\delta$ by $.01$, we'll change $\delta$ **really finely** (like, $.00004$) near our observed value. This reveals the stepwise structure:
f = plt.figure()
ax = plt.gca()
ax.axis((0,1,0,1))
ax.plot((house06.vote_share.mean() + numpy.linspace(-.01, .01, num=500)),
[((house06.vote_share + delta) > .5).mean()
for delta in numpy.linspace(-.01,.01,num=500)],
marker='x', color='grey')
ax.plot(house06.vote_share.mean(),
(house06.vote_share > .5).mean(), marker='x', color='r')
ax.axis((.55,.58, .52,.55))
ax.set_xlabel("Average District Pct. Dem Vote")
ax.set_ylabel("Pct. of Congress Won by Dems")
plt.show()
# Now we can see the stepwise structure. Again, this happens because seats are either *won* or they're *lost* by a party; no fractional seats are possible, so the seats-votes curve always assigns a discrete number of "wins" to Democrats. If there are 435 seats, the "estimated" seats-votes curve can only change $1/435$ at any step. Sometimes, the seats-votes curve can change more than $1/435$ at a time, like if two districts tie exactly in their vote shares. But, it can never change less than $1/435$ due to integrality. If we visualize this as light small lines in the plot we can see this effect very clearly:
f = plt.figure()
ax = plt.gca()
ax.axis((0,1,0,1))
ax.plot((house06.vote_share.mean() + numpy.linspace(-.01, .01, num=500)),
[((house06.vote_share + delta) > .5).mean() for delta in numpy.linspace(-.01,.01,num=500)], marker='x', color='grey')
ax.plot(house06.vote_share.mean(),
(house06.vote_share > .5).mean(), marker='x', color='r')
ax.hlines((numpy.arange(1,435)/435), 0,1, color='k', linewidth=.15)
ax.set_xlabel("Average District Pct. Dem Vote")
ax.set_ylabel("Pct. of Congress Won by Dems")
ax.axis((.55,.58, .52,.55))
plt.show()
# Each of the horizontal lines is a change in exactly one seat. Note that two seats tie for the voteshares right around $.565$; we can see this by noting that the two district vote share percentages are so close to one another, they fall through the grid of values together. But, unless something ties *exactly*, the seats-votes curve only ever changes by a single seat at each percent vote value, and if the grid of $\delta$ became even finer, the curve would only increase exactly $1/435$ at a time.
#
# To complete the last bit of relationship between the seats-votes curve and a cumulative density function, we need to obtain an expression that relates the two kinds of curves.
#
# So, can see that when a party popular vote share is exactly $\bar{v}$, then the seats-votes curve will be the count where $v_i>.5$. If we're *instead* at some swung vote share $\bar{v} + \delta$, the height of the seats-votes curve will be the percent of seats won at that *swung* vote share ($v_i + \delta > .5$). Moving to the *empirical cumulative density function* (ECDF), the height of the ECDF at an arbitrary value $x$ is the percent of all observations less than or equal to that value ($v_i < x$). Arbitrarily, let $x = \bar{v} - \delta$, we can see that the criteria for the ECDF is *identical to* the within-election seats-votes curve, shifted by a factor, $.5 - \bar{v}$:
#
# $$
# \begin{align}
# v_i &< x \\
# v_i &< \bar{v} - \delta \\
# v_i &< \bar{v} - \delta + (.5 - .5) \\
# v_i + \delta + .5- \bar{v} &< .5 \\
# v_i + \delta + (.5- \bar{v}) &< .5
# \end{align}
# $$
#
# This holds for any $x$ in the domain of the cumulative density function, since $\delta$ is a free parameter and $\bar{v}$ is observed.
#
# The $.5 - \bar{v}$ factor isn't totally arbitrary: it's the *aggregate margin*, or the difference between the between the party's average voteshare in districts and $.5$, which is the threshold the party would need to control the legislature. Second, note that we've defined $x = \bar{v} - \delta$. This "flips" the direction of the seats-votes curve, since swings of size $\delta$ in the seats-votes curve are swings of size $-\delta$ in the ECDF. For a more rigorous discussion, again see my dissertation (Wolf, 2017) or Nagle (2018).
# In practice, this means that the ECDF *for Republicans* and the seats-votes curve *for Democrats* are simply shifted & scaled versions of the same curve. Since we can construct the ECDF without reference to a theory of partisan swing, this suggests the "empirical" seats-votes curve is sufficiently more general than it might appear at first glance.
#
# As far as our ability to compute seats-votes curves is concerned, this means we can use all of the standard methods for computing histograms to "estimate" empirical seats-votes curves under the uniform partisan swing assumption. Below, I'll show the empirical CDF and swing-built seats-votes curve from above, zoomed in:
f = plt.figure()
ax = plt.gca()
ax.axis((0,1,0,1))
shift = .5 - house06.vote_share.mean()
ax.plot((house06.vote_share.mean() + numpy.linspace(-.10, .10, num=1000)),
[((house06.vote_share + delta) > .5).mean()
for delta in numpy.linspace(-.10,.10,num=1000)],
marker='x', color='grey', label='Uniform Swing')
ax.plot(house06.vote_share.mean(),
(house06.vote_share > .5).mean(), marker='x', color='r')
ax.hist(1-house06.vote_share - shift, cumulative=True, histtype='step',
density=True, bins=9000, color='red', zorder=100,
label='Republican Shifted ECDF')
ax.hist(1-house06.vote_share, cumulative=True, histtype='step',
density=True, bins=9000, color='purple', zorder=100,
label='Republican ECDF')
ax.hlines((numpy.arange(1,436)/435), 0,1, color='k', linewidth=.15)
ax.arrow(.475,.501,-shift, 0, color='b',
length_includes_head=True, zorder=100, label='Shift')
ax.hlines(1000,10001,1000, color='b', label='Shift') # because the shift label is disappearing
ax.legend(loc='center right', bbox_to_anchor=(2,.5), fontsize=20, frameon=False)
ax.axis((.46, .59, .48,.58))
ax.set_title("Seats-Votes & ECDF Equivalence")
ax.set_xlabel("Average District Pct. Dem Vote")
ax.set_ylabel("Pct. of Congress Won by Dems")
plt.show()
# We can do this for the nation for each year to get a collection of seats-votes curves that apply to each year, or use *all* seats across the elections to build one collected seats-votes curve. In practice, when we have multiple seats-votes curves, it's usually best to visualize each separately, and consider the *consensus* curve to to be the median among the many different replications. We'll see this strategy in Gelman & King (1994), as well as McGann et al. (2015) and Wolf (2018).
#
# Below, though, I'll plot seats-votes curves using the shifted Republican ECDF for each election since 1994.
for year,chunk in house.groupby('year'):
shift = .5 - chunk.vote_share.mean()
plt.hist(numpy.clip(1-chunk.vote_share.dropna().values - shift, 0,1),
cumulative=True, density=True, histtype='step',
color='k', bins=100, alpha=.2, zorder=1000,
label='Single Year' if year==2006 else None)
plt.scatter(house.groupby('year').vote_share.mean(),
house.groupby('year').vote_share.apply(lambda x: (x > .5).mean()),
label='Observed District\nAverages', marker='x', color='b')
plt.hist(numpy.clip((1-house.dropna(subset=['vote_share']).vote_share.values)
- (.5 - house.vote_share.mean()), 0,1),
cumulative=True, density=True, histtype='step',
color='orangered', bins=100, linewidth=3, label='Pooled')
plt.legend(loc='upper left', fontsize=14)
plt.xlabel("Average District %Dem")
plt.ylabel("Percent of Legislature Won")
plt.show()
# # Is the seats-votes curve realistic?
#
# The question of whether or not a seats-votes curve is *realistic* is quite distinct from *whether or not we can estimate it*. Again, following Nagle (2018), we can clearly "estimate" seats-votes curves for any number of replications from a model of elections. We can do this quickly, easily, and in a pretty straight-forward fashion. I'll talk a lot later about how there may be many different ways of constructing *stochastic* estimates of seats-votes curves, those that take into account the various random and co-varying factors that are involved in electoral processes. But, at its core, the question of whether or not we can estimate seats-votes curves easily is solved using Nagle (2018)'s relization about empirical CDFs and seats-votes curves.
#
# Their *realism* depends, though, on their interpretation. Interpreting seats-votes curves as empirical CDFs is not problematic at all. However, interpreting them *as a model* of how congressional control might change as district average vote shares change... that is problematic. In general, this interpretation hinges on how realistic the assumption of uniform partisan swing is.
#
# While we may take the uniform partisan swing curves as a good first-approximation for how the electoral system will work, there is a common perception that extremes of the seats-votes curves, the points that are far away from an observed $(\bar{v},\bar{s})$ point, are not "realible" representations of reality. In my own dissertation research, I found that policymakers, public officials, and stakeholders in Washington and and Arizona did believe that the whole range of $\bar{v}$ to $1-\bar{v}$ were possible, but that it was probably more likely that the structure of partisanship in the state would not change radically or flip in the control of the congressional delegation. This suggests that, for practitioners I interviewed, seats-votes curves may indeed have a narrow band of validity around the observed election results, and this band may encompass values where a party both wins or loses. This doesn't mean that the *estimation* of the curve is unrealible, but it *does* mean the model becomes unrealistic at large swings.
#
# However, another component of the assumption of uniform partisan swing that I examined in my dissertation considered whether the assumption of *uniformity* makes sense *geographically*. That is, do nearby districts tend to swing the same direction (or, more-so than they may otherwise if swing were uniformly random)? It may be the case that small but *highly correlated* swings will behave differently from small uncorrelated swings. If *this* is true, it suggests that assuming that *all districts accrue the same changes* might be unrealistic, and this unrealism may affect the validity of the seats-votes curve as a conceptual model.
#
# To examine this more in depth, we can see the votes over the 2002-2010 elections to Congress.
aughties = house.query('year > 2000 & year < 2012')
# Now, admitting that there are a few inter-censal redistrictings during this period, I'll assume that the redistricting map remains *mainly* unchanged, and that it is reasonable to compute swing between two districts in subsequent elections so long as they have the same district number.
aughties['district_id'] = aughties.state_fips.astype(str).str.rjust(2,'0') \
+ aughties.lewis_dist.astype(str).str.rjust(2, '0')
# Now, I'm going to pivot the dataframe so that each row is a congressional district and each column is a year in which a vote share is recorded.
pivot_table = aughties.pivot(index='district_id',
columns='year',
values='vote_share')
pivot_table.head()
# Then, we'll compute the `diff` along columns and drop the first (since it has nothing to diff against):
swings = pivot_table.diff(axis=1).drop(2002, axis=1)
swings.head()
# We can see the swing distributions in each year:
swings.hist(bins=40)
# Now, swing is *quite* noisy when considering both contested and uncontested elections together. We see that, in general, there appear to be three modes in these distributions. These tend to be where elections change between contested and uncontested across two elections. If we focus in on the cases where elections are contested, we'll have a bit of a cleaner time when we move to the map. Thus, let's focus on the middle 80% first, and return later to the problem of uncontested elections.
midswing = aughties.query('vote_share < .9 & vote_share > .1')\
.pivot(index='district_id',
columns='year',
values='vote_share')\
.diff(axis=1).drop(2002,axis=1)
midswing.hist(bins=40)
plt.tight_layout()
plt.show()
# Given this drop of uncontested elections, contests where *either* year is uncontested result in null swing. But, looking generally at a map of the swings (here, we'll focus on 2004), we can see that it does indeed appear to cluster in space:
spswing = aughties.merge(midswing[2004].to_frame('swing'),
left_on='district_id',
right_index=True,how='left')
spswing = spswing.query('state_name not in ("alaska", "hawaii")')
ax = spswing[~spswing.swing.isnull()].plot('swing', cmap='RdYlBu')
spswing[spswing.swing.isnull()].plot(color='lightgrey', edgecolor='k',
linewidth=.1, zorder=-1,ax=ax)
# Given that we've focused only on contested elections in both cycles, it appears that the changes in vote share between years *does* tend to be spatially dependent. To verify this, we can start by using Moran's $I$, which is kind of like a spatial correlation coefficient. This will tell us how correlated a given congressional district's swing is with other nearby districts. Over all districts, this will tell us whether the *swing* at any site tends to be related to swing in nearby sites. If districts that are near each other tend to experience the same swing, then it is not really the case that swing exists uniformly at random across contested elections. Instead, there are *correlated swings*: areas that are near each other tend to swing together.
Is = []
for y in numpy.arange(2004,2012,2):
spswing = aughties.merge(midswing[y].to_frame('swing'),
left_on='district_id',
right_index=True,how='left')
spswing = spswing.query('state_name not in ("alaska", "hawaii")')
spswing = spswing.dropna(subset=['swing'])
W = pysal.weights.Queen.from_dataframe(spswing)
Is.append(pysal.Moran(spswing.swing.values, W))
# We can visualize this in on of two ways. First, we can simply look at the estimate of Moran's $I$ and its standard error in each year:
plt.errorbar(numpy.arange(2004,2012,2),
[mi.I for mi in Is],
[2*(mi.VI_rand**.5) for mi in Is],
fmt='none', color='k')
plt.scatter(numpy.arange(2004,2012,2),
[mi.I for mi in Is], marker='x', color='r', zorder=100)
plt.xticks(numpy.arange(2004,2012,2))
seaborn.despine()
plt.xlabel("Moran's I for queen congressional district swings")
# Or, we can look at the scatterplot relating the swing in a contested election to the swing in neighboring districts in each year:
f,ax = plt.subplots(2,2,figsize=(8,8))
for i,year in enumerate(numpy.arange(2004,2012,2)):
ax_ = ax.flatten()[i]
seaborn.regplot(Is[i].z,
pysal.weights.lag_spatial(Is[i].w, Is[i].z),
ax=ax_, scatter_kws=dict(color='k', marker='.', s=4),
line_kws=dict(color='r'))
ax_.hlines(0,*plt.xlim(), linestyle='--', color='grey')
ax_.vlines(0,*plt.ylim(), linestyle='--', color='grey')
ax_.plot((-.1,.1), (-.1,.1), color='skyblue', linestyle='--')
ax_.set_title("Moran Plot for {}".format(year))
ax_.set_xlabel("Swing")
f.tight_layout()
plt.show()
# Thus, we see clearly that swing is *not* uniformly random over space; areas that are near each other tend to break towards (or away from) candidates in a simlar fashion.
|
notebooks/01 - Seat-Vote, Rank-Vote, & Single Elections.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19"
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python Docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
# Input data files are available in the read-only "../input/" directory
# For example, running this (by clicking run or pressing Shift+Enter) will list all files under the input directory
import os
for dirname, _, filenames in os.walk('/kaggle/input'):
for filename in filenames:
print(os.path.join(dirname, filename))
from tensorflow import keras
from tensorflow.keras import layers
from tensorflow.keras.datasets import mnist
from tensorflow.keras.utils import to_categorical
import matplotlib.pyplot as plt
# You can write up to 20GB to the current directory (/kaggle/working/) that gets preserved as output when you create a version using "Save & Run All"
# You can also write temporary files to /kaggle/temp/, but they won't be saved outside of the current session
# -
(train_images, train_labels), (test_images, test_labels) = mnist.load_data()
print("The shape of train_image, train_labels, test_images and test_labels are {}, {},{} & {}".format(
train_images.shape, train_labels.shape, test_images.shape, test_labels.shape))
#Visualizating loaded data
digit = train_images[1]
plt.imshow(digit, cmap=plt.cm.binary)
plt.show()
#Manipulating tensor images
my_slice = train_images[10:100]
my_slice.shape
#Creating a batch
batch = train_images[:128]
#Tensor reshaping
train_images_reshaped = train_images.reshape((60000, 28 * 28))
train_images_reshaped.shape
# ***Creating convnet layers***
train_x = train_images.reshape((60000, 28,28,1))
train_y = train_labels.astype('float32')/255
test_x = test_images.reshape((10000, 28,28,1))
test_y = test_labels.astype('float32')/255
model = keras.Sequential(([
layers.Conv2D(32, (3, 3), activation='relu', input_shape=(28, 28, 1)),
layers.MaxPooling2D((2, 2)),
layers.Conv2D(64, (3, 3), activation='relu'),
#layers.MaxPooling2D((2, 2)),
#layers.Conv2D(64, (3, 3), activation='relu'),
layers.Flatten(),
layers.Dense(64, activation = 'relu'),
layers.Dense(10, activation = 'softmax')
]))
# ***Training the convent layers***
train_x = train_images.reshape((60000, 28, 28, 1))
train_y = train_images.astype('float32') / 255
test_x = test_images.reshape((10000, 28, 28, 1))
test_y = test_images.astype('float32') / 255
train_y = to_categorical(train_y)
test_x = to_categorical(train_x)
model.compile(optimizer='rmsprop',
loss='categorical_crossentropy',
metrics=['accuracy'])
model.fit(train_x, train_y, epochs=5, batch_size=64)
|
.ipynb_checkpoints/mnist-digit-classification-by-tensorflow-b2055443-14b1-4923-b369-f9d038824a44-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# -*- coding: utf-8 -*-
import numpy as np
import cv2
import matplotlib.pyplot as plt
import os
# plot vermell
ruta=os.path.dirname(os.path.abspath(__file__))
img=cv2.imread(ruta+'/Users/Dani_Retamosa/Downloads/PROJECTE/castell cartoixa Terrassa/castell_101.jpeg',1)
surf=cv2.SURF(4000)
kp,des=surf.detectAndCompute(img, None)
img2=cv2.drawKeypoints(img, kp, None, (255,0,0), 4)
plt.imshow(img2),plt.show()
print len(kp)
|
Tasca_1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
# +
# More Data Types
# -
# Casting
np.array([1,2,3]) + 1.5
# assignment never changes the type, though
a = np.array([4,2,3])
print(a.dtype)
a[0] = 1.9 # <-- Float gets truncated (not rounded) to Int to match the array type
a
# But you CAN force a cast to a new type
a = np.array([1,2, 1.5, 1.6, 2.5, 3.5, 4.5])
b = np.around(a)
print(b)
c = np.around(a).astype(int)
c
# +
# Different Data Type Sizes
# -
np.array([1], dtype=int).dtype
np.iinfo(np.int32).max, 2**31 - 1
np.iinfo(np.uint32).max, 2**32 - 1
np.finfo(np.float32).eps
np.finfo(np.float64).eps
np.float32(1e-8) + np.float32(1) == 1
np.float64(1e-8) + np.float64(1) == 1
# +
# Don't use special datatypes if you don't know you need them.
a = np.zeros((1e6,), dtype=np.float64)
b = np.zeros((1e6,), dtype=np.float32)
# %timeit a*a
# %timeit b*b
# +
# Structured Data Types
# +
samples = np.zeros((6,), dtype=[('sensor_code', 'S4'),
('position', float),
('value', float)])
print(samples.ndim)
print(samples.shape)
print(samples.dtype.names)
print(samples)
samples[:] = [('ALFA', 1, 0.37), ('BETA', 1, 0.11), ('TAU', 1, 0.13),
('ALFA', 1.5, 0.37), ('BETA', 3, 0.11), ('TAU', 1.2, 0.13)]
samples
# -
# Field access works by indexing by the field names. For example...
samples['sensor_code']
samples['value']
samples['position']
samples[0]
samples[0]['sensor_code']
samples[0]['sensor_code'] = 'TAU'
samples[0]
# Multiple simultaneous field access
samples[['sensor_code', 'value']]
# And the fancy indexing still works
samples[samples['sensor_code'] == 'ALFA']
# +
# maskedarray: dealing with (propogation of) missing data
# -
# For floats, one could use NaN, but masks work for all types
x = np.ma.array([1,2,3,4], mask=[0,1,0,1])
x
y = np.ma.array([1,2,3,4], mask=[0,1,1,1])
y
x + y
# Masking versions of common functions
np.ma.sqrt([1,-1,2,-2])
|
Ch1/SciPy_NumPy/Ch_1_3_3_More_Elaborate_Arrays.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Plot pre-projected data defined in PDEF
# **12 April 2020 by MiniUFO**
#
# ---
# [TOC]
#
# ---
# ### 1. Introduction
# The current [**xgrads**](https://github.com/miniufo/xgrads) package is capable of parsing `PDEF` info defined in `ctl` ([**see the doc here**](http://cola.gmu.edu/grads/gadoc/pdef.html)). With a `PDEF` defined, the `ctl` contains two sets of grids. One is the **native** grid with grid points given in `PDEF` and the data are known as **pre-projected** data. Another is the rectilinear lat/lon grid commonly defined by `XDEF` and `YDEF`. As a result, the number of grid points defined in `PDEF` is used for reading and converting to [**xarray.Dataset**](http://xarray.pydata.org/en/stable/). The common lat/lon grid is used internally by [**GrADS**](http://cola.gmu.edu/grads/) for interpolation and displaying. So the lat/lon grid and its resolution can be modified at will.
#
# As a light-weighted package, [**xgrads**](https://github.com/miniufo/xgrads) does not intend to do the interpolation and display the interpolated fields as [**GrADS**](http://cola.gmu.edu/grads/) does. For those who may want to do a similar job, this notebook is a quick guide to read the `ctl` file with `PDEF` and plot the pre-projected data with [**cartopy**](https://scitools.org.uk/cartopy/docs/latest/)
#
#
# ---
# ### 2. A case of Lambert Conformal Conic Projection (LCC)
# The example `ctl` file is:
#
# ```python
# dset ^EMI_2019_monthly.grd
# options big_endian
# title CUACE_emi_index data
# undef -9999.
# pdef 360 320 nps 100 170 130 15
# xdef 720 linear 0.00 0.5
# ydef 682 linear 41.18 0.0676
# zdef 1 levels 1
# tdef 10 linear JAN2019 1mo
# vars 12
# emi_index 0 99 pm u2/m3
# demi_index 0 99 pm u2/m3
# emisdep_index 0 99 pm u2/m3
# diff_index 0 99 pm u2/m3
# trans_index 0 99 pm u2/m3
# trans_in 0 99 pm u2/m3
# trans_out 0 99 pm u2/m3
# surf_index 0 99 pm 1/1
# semi_index 0 99 pm u2/m3
# sdemi_index 0 99 pm u2/m3
# emitest_index 0 99 pm u2/m3
# deptest_index 0 99 pm u2/m3
# endvars
# ```
#
# First parse the `ctl` file as:
# +
import sys
sys.path.append('../')
from xgrads import CtlDescriptor, open_CtlDataset
ctl = CtlDescriptor(file='d:/EMI_2019_monthly.ctl')
print(ctl.pdef)
# -
# `PDEF` is parsed OK. So load the data into `xarray.Dataset`:
# +
dset = open_CtlDataset('d:/EMI_2019_monthly.ctl')
print(dset)
# -
# Plot the pre-projected data without map projection using [**xarray**](http://xarray.pydata.org/en/stable/)'s wrapper of [**matplotlib**](https://matplotlib.org/):
data = dset.emi_index[0]
data.where(data!=ctl.undef).plot(figsize=(9,5), cmap='jet')
# Now we are going to display the pre-projected data on a map. This is easy under the help of [**cartopy**](https://scitools.org.uk/cartopy/docs/latest/). Notice that the `ctl` object has a `get_data_projection` function to provide a map projection in accordance with [**cartopy**](https://scitools.org.uk/cartopy/docs/latest/).
# +
import cartopy.crs as ccrs
import cartopy.feature as cf
import matplotlib.pyplot as plt
# this data projection is defined by PDEF, and will
# be used by cartopy for plotting.
data_proj = ctl.get_data_projection()
# Note that data projection is uniquely defined by PDEF.
# But we can plot the data in different map projections.
# Here choose three for demonstration.
map_proj_pcr = ccrs.PlateCarree(central_longitude=105)
map_proj_lcc = ccrs.LambertConformal(central_longitude=105)
map_proj_orth = ccrs.Orthographic(central_longitude=105)
plt.figure(figsize=(15,10))
ax = plt.subplot(131, projection = map_proj_pcr)
ax.contourf(data.x, data.y, data, transform=data_proj, cmap='jet')
ax.coastlines('50m')
ax.add_feature(cf.BORDERS)
ax.set_title('PlateCarree projection (similar to GrADS)')
ax = plt.subplot(132, projection = map_proj_lcc)
ax.contourf(data.x, data.y, data, transform=data_proj, cmap='jet')
ax.coastlines('50m')
ax.add_feature(cf.BORDERS)
ax.set_title('Lambert conformal projection')
ax = plt.subplot(133, projection = map_proj_orth)
ax.contourf(data.x, data.y, data, transform=data_proj, cmap='jet')
ax.coastlines('50m')
ax.add_feature(cf.BORDERS)
ax.set_title('Orthographic projection')
ax.set_global()
# -
# ---
# ### 3. A case of North Polar Stereo projection (NPS)
# The ctl file has been modified slightly as:
# ```python
# dset ^EMI_2019_monthly.grd
# options big_endian
# title CUACE_emi_index data
# undef -9999.
# pdef 360 320 nps 100 170 130 15
# xdef 720 linear 0.00 0.5
# ydef 682 linear 41.18 0.0676
# zdef 1 levels 1
# tdef 10 linear JAN2019 1mo
# vars 12
# emi_index 0 99 pm u2/m3
# demi_index 0 99 pm u2/m3
# emisdep_index 0 99 pm u2/m3
# diff_index 0 99 pm u2/m3
# trans_index 0 99 pm u2/m3
# trans_in 0 99 pm u2/m3
# trans_out 0 99 pm u2/m3
# surf_index 0 99 pm 1/1
# semi_index 0 99 pm u2/m3
# sdemi_index 0 99 pm u2/m3
# emitest_index 0 99 pm u2/m3
# deptest_index 0 99 pm u2/m3
# endvars
# ```
# Note the projection in `PDEF` becomes `nps`. The loading and plotting is similar:
# +
dset, ctl = open_CtlDataset('d:/EMI_2019_monthly2.ctl', returnctl=True)
data = dset.emi_index[0]
# this time it is North Polar Stereo (NPS) projection
data_proj = ctl.get_data_projection()
# Note that data projection is uniquely defined by PDEF.
# But we can plot the data in different map projections.
# Here choose three for demonstration.
map_proj_pcr = ccrs.PlateCarree(central_longitude=180)
map_proj_nps = ccrs.NorthPolarStereo(central_longitude=180)
map_proj_orth = ccrs.Orthographic(central_longitude=105, central_latitude=70)
plt.figure(figsize=(15,10))
ax = plt.subplot(131, projection = map_proj_pcr)
ax.contourf(data.x, data.y, data, transform=data_proj, cmap='jet')
ax.coastlines('50m')
ax.add_feature(cf.BORDERS)
ax.set_title('PlateCarree projection (similar to GrADS)')
ax = plt.subplot(132, projection = map_proj_nps)
ax.contourf(data.x, data.y, data, transform=data_proj, cmap='jet')
ax.coastlines('50m')
ax.add_feature(cf.BORDERS)
ax.set_title('North Polar Stereo projection')
ax = plt.subplot(133, projection = map_proj_orth)
ax.contourf(data.x, data.y, data, transform=data_proj, cmap='jet')
ax.coastlines('50m')
ax.add_feature(cf.BORDERS)
ax.set_title('Orthographic projection')
ax.set_global()
# -
# ---
# ### 4. Summary
#
# [GrADS](http://cola.gmu.edu/grads/) has two jobs with a `PDEF` ctl file. One is loading the binary data and the other is plotting with proper map projection. Here [xgrads](https://github.com/miniufo/xgrads) package will only do the loading task and let [cartopy](https://scitools.org.uk/cartopy/docs/latest/) do the plotting job.
#
# ---
# ### References
# [http://cola.gmu.edu/grads/gadoc/pdef.html](http://cola.gmu.edu/grads/gadoc/pdef.html)
#
# [https://scitools.org.uk/cartopy/docs/latest/](https://scitools.org.uk/cartopy/docs/latest/)
#
# [https://matplotlib.org/](https://matplotlib.org/)
#
# [http://xarray.pydata.org/en/stable/](http://xarray.pydata.org/en/stable/)
|
notebooks/Plot preprojected data defined in PDEF.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Simple RNN
#
# In this notebook, we're going to train a simple RNN to do **time-series prediction**. Given some set of input data, it should be able to generate a prediction for the next time step!
# <img src='assets/time_prediction.png' width=40% />
#
# > * First, we'll create our data
# * Then, define an RNN in PyTorch
# * Finally, we'll train our network and see how it performs
# ### Import resources and create data
import torch
from torch import nn
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# +
plt.figure(figsize=(8,5))
# how many time steps/data pts are in one batch of data
seq_length = 20
# generate evenly spaced data pts
time_steps = np.linspace(0, np.pi, seq_length + 1)
data = np.sin(time_steps)
data.resize((seq_length + 1, 1)) # size becomes (seq_length+1, 1), adds an input_size dimension
x = data[:-1] # all but the last piece of data
y = data[1:] # all but the first
# display the data
plt.plot(time_steps[1:], x, 'r.', label='input, x') # x
plt.plot(time_steps[1:], y, 'b.', label='target, y') # y
plt.legend(loc='best')
plt.show()
# -
# ---
# ## Define the RNN
#
# Next, we define an RNN in PyTorch. We'll use `nn.RNN` to create an RNN layer, then we'll add a last, fully-connected layer to get the output size that we want. An RNN takes in a number of parameters:
# * **input_size** - the size of the input
# * **hidden_dim** - the number of features in the RNN output and in the hidden state
# * **n_layers** - the number of layers that make up the RNN, typically 1-3; greater than 1 means that you'll create a stacked RNN
# * **batch_first** - whether or not the input/output of the RNN will have the batch_size as the first dimension (batch_size, seq_length, hidden_dim)
#
# Take a look at the [RNN documentation](https://pytorch.org/docs/stable/nn.html#rnn) to read more about recurrent layers.
class RNN(nn.Module):
def __init__(self, input_size, output_size, hidden_dim, n_layers):
super(RNN, self).__init__()
self.hidden_dim=hidden_dim
# define an RNN with specified parameters
# batch_first means that the first dim of the input and output will be the batch_size
self.rnn = nn.RNN(input_size, hidden_dim, n_layers, batch_first=True)
# last, fully-connected layer
self.fc = nn.Linear(hidden_dim, output_size)
def forward(self, x, hidden):
# x (batch_size, seq_length, input_size)
# hidden (n_layers, batch_size, hidden_dim)
# r_out (batch_size, time_step, hidden_size)
batch_size = x.size(0)
# get RNN outputs
r_out, hidden = self.rnn(x, hidden)
# shape output to be (batch_size*seq_length, hidden_dim)
r_out = r_out.view(-1, self.hidden_dim)
# get final output
output = self.fc(r_out)
return output, hidden
# ### Check the input and output dimensions
#
# As a check that your model is working as expected, test out how it responds to input data.
# +
# test that dimensions are as expected
test_rnn = RNN(input_size=1, output_size=1, hidden_dim=10, n_layers=2)
# generate evenly spaced, test data pts
time_steps = np.linspace(0, np.pi, seq_length)
data = np.sin(time_steps)
data.resize((seq_length, 1))
test_input = torch.Tensor(data).unsqueeze(0) # give it a batch_size of 1 as first dimension
print('Input size: ', test_input.size())
# test out rnn sizes
test_out, test_h = test_rnn(test_input, None)
print('Output size: ', test_out.size())
print('Hidden state size: ', test_h.size())
# -
# ---
# ## Training the RNN
#
# Next, we'll instantiate an RNN with some specified hyperparameters. Then train it over a series of steps, and see how it performs.
# +
# decide on hyperparameters
input_size=1
output_size=1
hidden_dim=32
n_layers=1
# instantiate an RNN
rnn = RNN(input_size, output_size, hidden_dim, n_layers)
print(rnn)
# -
# ### Loss and Optimization
#
# This is a regression problem: can we train an RNN to accurately predict the next data point, given a current data point?
#
# >* The data points are coordinate values, so to compare a predicted and ground_truth point, we'll use a regression loss: the mean squared error.
# * It's typical to use an Adam optimizer for recurrent models.
# MSE loss and Adam optimizer with a learning rate of 0.01
criterion = nn.MSELoss()
optimizer = torch.optim.Adam(rnn.parameters(), lr=0.01)
# ### Defining the training function
#
# This function takes in an rnn, a number of steps to train for, and returns a trained rnn. This function is also responsible for displaying the loss and the predictions, every so often.
#
# #### Hidden State
#
# Pay close attention to the hidden state, here:
# * Before looping over a batch of training data, the hidden state is initialized
# * After a new hidden state is generated by the rnn, we get the latest hidden state, and use that as input to the rnn for the following steps
# train the RNN
def train(rnn, n_steps, print_every):
# initialize the hidden state
hidden = None
for batch_i, step in enumerate(range(n_steps)):
# defining the training data
time_steps = np.linspace(step * np.pi, (step+1)*np.pi, seq_length + 1)
data = np.sin(time_steps)
data.resize((seq_length + 1, 1)) # input_size=1
x = data[:-1]
y = data[1:]
# convert data into Tensors
x_tensor = torch.Tensor(x).unsqueeze(0) # unsqueeze gives a 1, batch_size dimension
y_tensor = torch.Tensor(y)
# outputs from the rnn
prediction, hidden = rnn(x_tensor, hidden)
## Representing Memory ##
# make a new variable for hidden and detach the hidden state from its history
# this way, we don't backpropagate through the entire history
hidden = hidden.data
# calculate the loss
loss = criterion(prediction, y_tensor)
# zero gradients
optimizer.zero_grad()
# perform backprop and update weights
loss.backward()
optimizer.step()
# display loss and predictions
if batch_i%print_every == 0:
print('Loss: ', loss.item())
plt.plot(time_steps[1:], x, 'r.') # input
plt.plot(time_steps[1:], prediction.data.numpy().flatten(), 'b.') # predictions
plt.show()
return rnn
# +
# train the rnn and monitor results
n_steps = 75
print_every = 15
trained_rnn = train(rnn, n_steps, print_every)
# -
# ### Time-Series Prediction
#
# Time-series prediction can be applied to many tasks. Think about weather forecasting or predicting the ebb and flow of stock market prices. You can even try to generate predictions much further in the future than just one time step!
|
recurrent-neural-networks/time-series/Simple_RNN.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
messages=pd.read_csv("SMSSpamCollection",sep="\t",names=['label','message'])
messages
messages.info()
import re
import nltk
from nltk.corpus import stopwords
from nltk.stem import PorterStemmer
ps=PorterStemmer()
corpus=[]
for i in range(0,len(messages)):
review=re.sub('[^a-zA-Z]','',messages['message'][i])
review=review.lower()
review=review.split()
review=[ps.stem(word) for word in review if not word in set(stopwords.words('english'))]
review=' '.join(review)
corpus.append(review)
corpus[:5]
from sklearn.feature_extraction.text import CountVectorizer
cv=CountVectorizer(max_features=1000)
X=cv.fit_transform(corpus).toarray()
X.shape
y=pd.get_dummies(messages['label'],drop_first=True)
y
from sklearn.model_selection import train_test_split
X_train,X_test,y_train,y_test=train_test_split(X,y,test_size=0.4,random_state=0)
from sklearn.naive_bayes import MultinomialNB
spam_detect_model=MultinomialNB().fit(X_train,y_train)
y_pred=spam_detect_model.predict(X_test)
from sklearn.metrics import confusion_matrix,accuracy_score
confusion_matrix(y_pred,y_test)
accuracy_score(y_pred,y_test)
|
Spamclassifier.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from scipy.stats import rankdata
# +
df = pd.read_csv("data.csv")
df.index = pd.to_datetime(df['date'], format='%Y-%m-%d')
df = df.drop('date', axis=1)
close_columns = []
high_columns = []
low_columns = []
open_columns = []
volume_columns = []
open_int_columns = []
for i in df.columns:
if "close" in i:
close_columns.append(i)
elif "high" in i:
high_columns.append(i)
elif "low" in i:
low_columns.append(i)
elif "open_int" in i:
open_int_columns.append(i)
elif "open" in i:
open_columns.append(i)
elif "volume" in i:
volume_columns.append(i)
close_df = df[close_columns]
high_df = df[high_columns]
low_df = df[low_columns]
open_df = df[open_columns]
volume_df = df[volume_columns]
open_int_df = df[open_int_columns]
# -
df.tail()
# ## Signals
# +
# daily data
returns_daily = close_df.pct_change().dropna()
vol_daily = returns_daily.ewm(adjust=True, com=60, min_periods=0).std().dropna()
#monthly data
returns_monthly = close_df.pct_change(20).dropna().resample('BM').last().ffill()
vol_monthly = (np.sqrt(261)*vol_daily).resample('BM').last().ffill()
# -
vol_monthly.head()
# # TSMOM
# +
def signal2(df, date, passive, method):
num_assets = len(df.iloc[-1])
signal = []
if method == "momentum":
returns = df.pct_change(20 * 12).resample('BM').last().ffill()[:date]
if passive:
signal = np.ones(num_assets)
else:
signal = np.where(returns.iloc[-1] > 0, 1, -1)
elif method == "momentum_lagged":
returns_12 = df.pct_change(21 * 12).resample('BM').last().ffill()[:date]
returns_6 = df.pct_change(21 * 6).resample('BM').last().ffill()[:date]
returns_3 = df.pct_change(21 * 3).resample('BM').last().ffill()[:date]
momentum_mean = (returns_12.iloc[-1] + returns_6.iloc[-1] + returns_3.iloc[-1]) / 3
if passive:
signal = np.ones(num_assets)
else:
signal = np.where(momentum_mean > 0, 1, -1)
return signal
def tsmom (df,returns_monthly, vol_monthly,date, method = 'momentum', risk=0.4, passive=False, momentum_window=12):
position = signal2(df, date, passive, method)
weights = (risk / vol_monthly.iloc[date-1])
weights /= len(weights)
portfolio = position * weights
return (1+np.dot(portfolio, returns_monthly.iloc[date]))
# -
# # CSMOM
# +
def signal(df, date):
num_assets = len(df.iloc[-1])
signal = []
returns = df.pct_change(20 * 12).resample('BM').last().ffill()[:date]
returns_rank = rankdata(returns.iloc[-1])
signal = np.where(returns_rank > int(num_assets * 0.7), 1, np.where(returns_rank < int(num_assets * 0.3), -1, 0))
return signal
def csmom (df,returns_monthly, vol_monthly, date):
position = signal(df, date)
num_assets = len(df.iloc[-1])
weights = 1 / (int(num_assets - num_assets * 0.8) + int(num_assets * 0.2))
print(weights, int(num_assets * 0.8), int(num_assets * 0.2))
portfolio = position * weights
return (1+np.dot(portfolio, returns_monthly.iloc[date]))
# -
# ## Classic CSMOM
r = [] # retorno do TSMOM
rp = [] # retorno passivo
start = 12 -1
years = 19
end = 12*(int(start/12) + years)
for i in range(start, end):
r.append(csmom (close_df,returns_monthly, vol_monthly, date=i))
rp.append(tsmom (close_df,returns_monthly, vol_monthly, date=i, risk=0.4))
# +
from pandas.plotting import register_matplotlib_converters
register_matplotlib_converters()
r = pd.DataFrame(r)
rp = pd.DataFrame(rp)
r.index = returns_monthly.iloc[start:end].index
rp.index = returns_monthly.iloc[start:end].index
plt.figure(figsize=(16,9))
plt.plot(100*r.cumprod(), label = 'CSMOM', color='blue')
plt.plot(100*rp.cumprod(), label = 'TSMOM', color='red')
plt.yscale('log')
plt.legend()
plt.title('Cumulative Excess Return of CSMOM and Diversified Passive Long')
plt.show()
tsmom_return = float((rp.cumprod().iloc[-1] - 1)*100)
csmom_return = float((r.cumprod().iloc[-1] - 1)*100)
print('Passive return: ', round(tsmom_return,2), '%,',
" Annualized: ", round((((100+tsmom_return)/100) ** (1/years) - 1)*100 , 2), "%", sep='')
print('CSMOM return: ', round(csmom_return,2), "%,",
" Annualized: ", round((((100+csmom_return)/100) ** (1/years) - 1)*100 , 2), "%", sep='')
|
CSMOM.ipynb
|
# ---
# jupyter:
# jupytext:
# cell_metadata_filter: collapsed
# formats: ipynb,py:percent
# text_representation:
# extension: .py
# format_name: percent
# format_version: '1.3'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %% [markdown]
# # Micro- and Macroeconomic Implications of Very Impatient Households
# %% [markdown]
# ## Introduction
#
# Buffer stock saving models of the kind implemented in $\texttt{ConsIndShockType}$ say that, if a standard ['Growth Impatience Condition'](https://econ.jhu.edu/people/ccarroll/papers/BufferStockTheory/#Growth-Modified-Conditions), holds:
#
# \begin{eqnarray}
# \newcommand{\Rfree}{\mathsf{R}}\newcommand{\DiscFac}{\beta}\newcommand{\PermGroFac}{\Gamma}\newcommand{\PermShk}{\psi}\newcommand{\CRRA}{\rho}
# \left(\frac{(\Rfree\DiscFac)^{1/\CRRA}\mathbb{E}[\PermShk^{-1}]}{\PermGroFac}\right) & < & 1
# \end{eqnarray}
#
# then the _ratio_ of asets $\newcommand{\aLev}{\mathbf{a}}\aLev$ to permanent income $\newcommand{\pLev}{\mathbf{p}}\pLev$, $a=\aLev/\pLev$, has a target value $\newcommand{\aTarg}{\check{a}}\aTarg$ that depends on the consumer's preferences (relative risk aversion $\CRRA$, time preference $\DiscFac$) and circumstances (interest factor $\Rfree$, growth factor $\PermGroFac$, uncertainty about permanent income shocks $\sigma^{2}_{\PermShk}$).
#
# If everyone had identical preferences and everyone were at their target $\check{a}$, then inequality in the level of $\aLev$ would be exactly the same as inequality in $\pLev$.
#
# ["The Distribution of Wealth and the Marginal Propensity to Consume"](http://econ.jhu.edu/people/ccarroll/papers/cstwMPC) (Carroll, Slacalek, Tokuoka, and White 2017; hereafter: "cstwMPC") shows that, when such a model is simulated and agents draw their idiosyncratic shocks (so, agents are _ex post_ heterogeneous -- see the definition in [Intro-To-HARK](http://github.com/econ-ark/PARK/tree/master/Intro-To-HARK.pdf)) -- asset inequality is indeed close to $\pLev$ inequality even though everyone is not always at exactly their target $a$.
#
# But a large body of evidence shows that _actual_ inequality in assets is much greater than _actual_ inequality in permanent income. Thus, to make a model that qualifies as what cstwMPC call a 'serious' microfounded macro model of consumption (one that matches the key facts _theory says_ should be first-order important), the model must be modified to incorporate some form of _ex ante_ heterogeneity: That is, there must be differences across people in $\DiscFac$ or $\Rfree$ or $\CRRA$ or $\PermGroFac$ or $\sigma^{2}_{\PermShk}$.
#
# The most transparent and simplest of these to change is the time preference factor $\beta$. So that is what the paper does. The main results are:
#
# 1. The distribution of $\beta$ need not be particularly wide to match the extreme concentration of wealth: roughly 0.91 to 0.98 (annual); that is, the most impatient person discounts the future about 6 percentage points more per year than the most patient agent agent
# 2. With such a distribution of $\beta$, simulated agents' (annual) marginal propensity to consume (MPC) from transitory income shocks to income matches large body of microeconomic evidence that typically finds evidence of MPC's in the range of 0.2 to 0.6. This is much better than RA macro models that typically yield MPC's in the range of 0.01 to 0.05.
#
# While the most impatient agents in the cstwMPC model have fairly high MPCs (~0.6 annual), there is microeconomic evidence that a significant fraction of households have *even higher* MPCs than the model predicts, especially at the quarterly frequency. This group of households is commonly referred to as "hand-to-mouth" -- they consume most of their transitory shocks to income not too long after they receive them (mostly within a quarter). There are several reasons why a household could be hand-to-mouth, but one plausible explanation is that these households are *even more impatient* than estimated by cstwMPC for the most impatient agent.
#
# %% [markdown]
# ### PROBLEM
# In this exercise, you will explore the micro- and macroeconomic implications of some households being *very impatient*. Specifically, you will address the following questions:
#
# 1. How does the distribution of the MPC change (relative to cstwMPC's baseline) if some simulated households are extremely impatient? Do we observe a significant portion of hand-to-mouth households?
# 2. How does the distribution (and aggregate level) of wealth change if some households are extremely impatient? Does this distribution of $\beta$ still generate a wealth distribution like the one seen in U.S. data?
# %% code_folding=[25]
# This cell does some setup and imports generic tools used to produce the figures
import sys
import os
sys.path.insert(0, os.path.abspath('../lib'))
from util import log_progress
import numpy as np
from copy import deepcopy
import HARK # Prevents import error from Demos repo
from HARK.utilities import plotFuncs
Generator=False # Is this notebook the master or is it generated?
# Import related generic python packages
# Set how many digits past the decimal point should be printed?
from time import clock
mystr = lambda number : "{:.4f}".format(number)
decfmt4 = lambda number : "{:.4f}".format(number)
decfmt3 = lambda number : "{:.3f}".format(number)
decfmt2 = lambda number : "{:.2f}".format(number)
decfmt1 = lambda number : "{:.1f}".format(number)
# This is a jupytext paired notebook that autogenerates BufferStockTheory.py
# which can be executed from a terminal command line via "ipython BufferStockTheory.py"
# But a terminal does not permit inline figures, so we need to test jupyter vs terminal
# Google "how can I check if code is executed in the ipython notebook"
from IPython import get_ipython # In case it was run from python instead of ipython
def in_ipynb():
try:
if str(type(get_ipython())) == "<class 'ipykernel.zmqshell.ZMQInteractiveShell'>":
return True
else:
return False
except NameError:
return False
# Determine whether to make the figures inline (for spyder or jupyter)
# vs whatever is the automatic setting that will apply if run from the terminal
if in_ipynb():
# # %matplotlib inline generates a syntax error when run from the shell
# so do this instead
get_ipython().run_line_magic('matplotlib', 'inline')
else:
get_ipython().run_line_magic('matplotlib', 'auto')
# Import the plot-figure library matplotlib
import matplotlib.pyplot as plt
# In order to use LaTeX to manage all text layout in our figures, we import rc settings from matplotlib.
from matplotlib import rc
plt.rc('font', family='serif')
# LaTeX is huge and takes forever to install on mybinder
# so if it is not installed then do not use it
from distutils.spawn import find_executable
iflatexExists=False
if find_executable('latex'):
iflatexExists=True
plt.rc('text', usetex= iflatexExists)
# The warnings package allows us to ignore some harmless but alarming warning messages
import warnings
warnings.filterwarnings("ignore")
from copy import copy, deepcopy
# %% [markdown]
# ## Calibrating a Basic Version of cstwMPC
#
# To get started, let's reproduce a simplified version of the main results from cstwMPC.
#
# In cstwMPC, the authors calibrated nearly all of the model parameters-- risk aversion, income shock process, etc-- to commonly used or previously estimated values. The only parameter to be estimated is the distribution of $\beta$. cstwMPC assumed that $\beta$ is uniformly distributed on $[\grave{\beta}-\nabla,\grave{\beta}+\nabla]$, approximated by a seven point distribution.
#
# Their estimation procedure seeks the values of $\grave{\beta}$ and $\nabla$ that generate a simulated distribution of wealth that best matches empirical U.S. data. Their definition of "best match" has two aspects:
#
# 1. The simulated aggregate capital-to-income ratio matches the true U.S. value.
# 2. The sum of squared distances between the simulated and empirical Lorenz curves (at the 20th, 40th, 60th, and 80th percentiles) is minimized (conditional on item 1).
#
# cstwMPC's target empirical moments are a capital-to-income ratio of 10.26 and cumulative wealth shares as given in the table below. Yes, you are reading the table correctly: The "poorest" 80 percent of households own 17.5 percent of wealth.
#
# | Net worth percentile | Cumulative wealth share |
# |:---:|:---:|
# | 20th | -0.2% |
# | 40th | 1.0% |
# | 60th | 5.5% |
# | 80th | 17.5% |
#
# To reproduce their basic results, we must import an $\texttt{AgentType}$ subclass and define a dictionary with calibrated parameters identical to those in the paper.
# %% code_folding=[0, 4]
# Import IndShockConsumerType
from HARK.ConsumptionSaving.ConsIndShockModel import IndShockConsumerType
# Define a dictionary with calibrated parameters
cstwMPC_calibrated_parameters = {
"CRRA":1.0, # Coefficient of relative risk aversion
"Rfree":1.01/(1.0 - 1.0/160.0), # Survival probability,
"PermGroFac":[1.000**0.25], # Permanent income growth factor (no perm growth),
"PermGroFacAgg":1.0,
"BoroCnstArt":0.0,
"CubicBool":False,
"vFuncBool":False,
"PermShkStd":[(0.01*4/11)**0.5], # Standard deviation of permanent shocks to income
"PermShkCount":5, # Number of points in permanent income shock grid
"TranShkStd":[(0.01*4)**0.5], # Standard deviation of transitory shocks to income,
"TranShkCount":5, # Number of points in transitory income shock grid
"UnempPrb":0.07, # Probability of unemployment while working
"IncUnemp":0.15, # Unemployment benefit replacement rate
"UnempPrbRet":None,
"IncUnempRet":None,
"aXtraMin":0.00001, # Minimum end-of-period assets in grid
"aXtraMax":40, # Maximum end-of-period assets in grid
"aXtraCount":32, # Number of points in assets grid
"aXtraExtra":[None],
"aXtraNestFac":3, # Number of times to 'exponentially nest' when constructing assets grid
"LivPrb":[1.0 - 1.0/160.0], # Survival probability
"DiscFac":0.97, # Default intertemporal discount factor; dummy value, will be overwritten
"cycles":0,
"T_cycle":1,
"T_retire":0,
'T_sim':1200, # Number of periods to simulate (idiosyncratic shocks model, perpetual youth)
'T_age': 400,
'IndL': 10.0/9.0, # Labor supply per individual (constant),
'aNrmInitMean':np.log(0.00001),
'aNrmInitStd':0.0,
'pLvlInitMean':0.0,
'pLvlInitStd':0.0,
'AgentCount':10000,
}
# %% [markdown]
# Now let's make several instances of our class of agents and give them different values of $\beta$, following cstwMPC's estimated distribution. In our specification of interest, we will use $\grave{\beta}=0.9855583$ and $\nabla = 0.0085$.
#
# NB: Reported parameter estimates in cstwMPC use a model with aggregate shocks and wage and interest rates determined dynamically (a heterogeneous agents DSGE model); this is the $\texttt{AggShockConsumerType}$ in HARK. The estimated parameters are slightly different in this exercise, as we are ignoring general equilibrium aspects and only using the $\texttt{IndShockConsumerType}$
# %%
# This cell constructs seven instances of IndShockConsumerType with different discount factors
from HARK.utilities import approxUniform
BaselineType = IndShockConsumerType(**cstwMPC_calibrated_parameters)
# Specify the distribution of the discount factor
num_types = 7 # number of types we want
DiscFac_mean = 0.9855583 # center of beta distribution
DiscFac_spread = 0.0085 # spread of beta distribution
DiscFac_dstn = approxUniform(num_types, DiscFac_mean-DiscFac_spread, DiscFac_mean+DiscFac_spread)[1]
MyTypes = [] # initialize an empty list to hold our consumer types
for nn in range(num_types):
# Now create the types, and append them to the list MyTypes
NewType = deepcopy(BaselineType)
NewType.DiscFac = DiscFac_dstn[nn]
NewType.seed = nn # give each consumer type a different RNG seed
MyTypes.append(NewType)
# %% [markdown]
# ## Solving and Simulating the Baseline Agents
#
# Now let's solve and simulate each of our types of agents. If you look in the parameter dictionary (or at any of the agent objects themselves), you will see that each one has an $\texttt{AgentCount}$ attribute of 10000. That is, these seven ex ante heterogeneous types each represent ten thousand individual agents that will experience ex post heterogeneity when they draw different income (and mortality) shocks over time.
#
# In the code block below, fill in the contents of the loop to solve and simulate each agent type for many periods. To do this, you should invoke the methods $\texttt{solve}$, $\texttt{initializeSim}$, and $\texttt{simulate}$ in that order. Simulating for 1200 quarters (300 years) will approximate the long run distribution of wealth in the population.
# %%
# Progress bar keeps track interactively of how many have been made
for ThisType in log_progress(MyTypes, every=1):
ThisType.solve()
ThisType.initializeSim()
ThisType.simulate()
# %% [markdown]
# To verify that you wrote that code correctly, let's check that the aggregate level of capital (total assets held by all households) to income ratio equals what we expected it would be. To do that, let's combine the asset holdings of all types, take the mean, and see if we get the desired capital to income ratio of 10.26.
#
# NB: Because there is no permanent income growth in this model, all shocks are mean one and idiosyncratic, and we have many agents, aggregate or average income is 1.0.
# %%
aLvl_all = np.concatenate([ThisType.aLvlNow for ThisType in MyTypes])
print('The ratio of aggregate capital to permanent income is ' + decfmt2(np.mean(aLvl_all)))
# %% [markdown]
# ## Plotting the Lorenz Curve
# %%
# Plot Lorenz curves for model with uniform distribution of time preference
from HARK.cstwMPC.SetupParamsCSTW import SCF_wealth, SCF_weights
from HARK.utilities import getLorenzShares, getPercentiles
pctiles = np.linspace(0.001,0.999,200)
sim_wealth = np.concatenate([ThisType.aLvlNow for ThisType in MyTypes])
SCF_Lorenz_points = getLorenzShares(SCF_wealth,weights=SCF_weights,percentiles=pctiles)
sim_Lorenz_points = getLorenzShares(sim_wealth,percentiles=pctiles)
plt.plot(pctiles,SCF_Lorenz_points,'--k')
plt.plot(pctiles,sim_Lorenz_points,'-b')
plt.xlabel('Percentile of net worth')
plt.ylabel('Cumulative share of wealth')
plt.show(block=False)
# %% [markdown]
# ## Calculating the Lorenz Distance at Targets
#
# Now we want to construct a function that calculates the Euclidean distance between simulated and actual Lorenz curves at the four percentiles of interest: 20, 40, 60, and 80.
# %% [markdown]
# ### PROBLEM - Create a Function to Calculate Lorenz Distance
# Now let's write a function that calculates the Euclidean distance between simulated and actual Lorenz curves at the four percentiles of interest: 20, 40, 60, and 80. Fill in the skeleton of the function below, and then test your function using the input $\texttt{MyTypes}$. If you did it correctly, the Lorenz distance should be 0.03.
#
# You may find it useful to check out some documentation for $\texttt{HARK.utilities}$ [at this link](https://econ-ark.github.io/HARK/generated/HARKutilities.html).
# %% [markdown]
# ## The Distribution Of the Marginal Propensity to Consume
#
# For many macroeconomic purposes, the distribution of the MPC $\kappa$ is more important than the distribution of wealth. Ours is a quarterly model, and MPC's are typically reported on an annual basis; we can compute an approximate MPC from the quraterly ones as $\kappa_{Y} \approx 1.0 - (1.0 - \kappa_{Q})^4$
#
# In the cell below, we retrieve the MPCs from our simulated consumers and show that the 10th percentile in the MPC distribution is only about 6 percent, while at the 90th percentile it is almost 0.5
# %%
# Retrieve the MPC's
percentiles=np.linspace(0.1,0.9,9)
MPC_sim = np.concatenate([ThisType.MPCnow for ThisType in MyTypes])
MPCpercentiles_quarterly = getPercentiles(MPC_sim,percentiles=percentiles)
MPCpercentiles_annual = 1.0 - (1.0 - MPCpercentiles_quarterly)**4
print('The MPC at the 10th percentile of the distribution is '+str(decfmt2(MPCpercentiles_annual[0])))
print('The MPC at the 50th percentile of the distribution is '+str(decfmt2(MPCpercentiles_annual[4])))
print('The MPC at the 90th percentile of the distribution is '+str(decfmt2(MPCpercentiles_annual[-1])))
# %% [markdown]
# ### PROBLEM
#
# Now let's look in more detail at the distribution of the MPC. In the code block below, write a function that produces text output of the following form:
#
# $\texttt{The 35th percentile of the MPC is 0.15623}$
#
# Your function should take two inputs: a list of types of consumers and an array of percentiles (numbers between 0 and 1). It should return no outputs, merely print to screen one line of text for each requested percentile. The model is calibrated at a quarterly frequency, but Carroll et al report MPCs at an annual frequency. To convert, use the formula:
#
# $\kappa_{Y} \approx 1.0 - (1.0 - \kappa_{Q})^4$
# %% [markdown]
# ## Adding Very Impatient Households
#
# Now that we have some tools for examining both microeconomic (the MPC across the population) and macroeconomic (the distribution and overall level of wealth) outcomes from our model, we are all set to conduct our experiment.
#
# In this exercise, we are going to add very impatient households to the economy in a very direct way: by replacing the *most impatient consumer type* with an *even more impatient type*. Specifically, we will have these agents have a discount factor of $\beta = 0.80$ at a quarterly frequency, which corresponds to $\beta \approx 0.41$ annual.
#
# In the code block below, we:
#
# 1. Replicate the list of agents using $\texttt{deepcopy}$.
# 2. Set the $\beta$ of the most impatient type to $0.80$ (for the copied set of agents).
# 3. Solve and simulate the most impatient type (for the copied set of agents).
# %%
# Follow the instructions above to make another list of agents that includes *very* impatient households.
NewTypes = deepcopy(MyTypes)
NewTypes[0].DiscFac = 0.8
NewTypes[0].solve()
NewTypes[0].initializeSim()
NewTypes[0].simulate()
# Retrieve the MPC's
percentiles=np.linspace(0.1,0.9,9)
MPC_sim = np.concatenate([ThisType.MPCnow for ThisType in NewTypes])
MPCpercentiles_quarterly = getPercentiles(MPC_sim,percentiles=percentiles)
MPCpercentiles_annual = 1.0 - (1.0 - MPCpercentiles_quarterly)**4
print('The MPC at the 10th percentile of the distribution is '+str(decfmt2(MPCpercentiles_annual[0])))
print('The MPC at the 50th percentile of the distribution is '+str(decfmt2(MPCpercentiles_annual[4])))
print('The MPC at the 90th percentile of the distribution is '+str(decfmt2(MPCpercentiles_annual[-1])))
# %% [markdown]
# ### PROBLEM
# ## Testing the Implications of Very Impatient Households
#
# Now that we have the baseline set of simulated agents (in $\texttt{MyTypes}$) and the altered set of simulated agents (in whatever you named your copied version), let's investigate what this means for micro- and macroeconomic outcomes. In the code block below, use both lists of agents and the data tools you wrote above to investigate the following questions:
#
# 1. Did introducing very impatient households generate a substantial proportion of hand-to-mouth households?
# - Define 'hand to mouth' as households whose quarterly MPC is greater than 0.7
# 2. Did introducing very impatient households affect the simulated model's ability to match the empirical distribution of wealth and its aggregate level?
# 3. Much of the "behavioral" consumption literature concludes, when consumers are found to have very high MPC's, that the standard optimal consumption model "doesn't work"
# * Given what you have found, can you reject the hypothesis that hand-to-mouth households arise in the data because they are very impatient?
#
# Use the markdown block below the code block to briefly answer those questions.
# %% [markdown]
# ### PROBLEM -- Plot the new distribution of wealth
#
# The $\texttt{matplotlib}$ library provides plotting functionality that replicates Matlab's plot features (more or less). As an example of how to use it, we have written a few lines of code that plot the empirical vs simulated Lorenz curves. Write some code that plots the CDF of the MPC before and after adding very impatient households, and plots the DIFFERENCES between the Lorenz curves across the two populations. Interpret the two graphs.
|
notebooks/Micro-and-Macro-Implications-of-Very-Impatient-HHs-Problems.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/misqualzarabi/DS-Unit-1-Sprint-2-Data-Wrangling-and-Storytelling/blob/master/Join_and_Reshape_Data.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="UQqT5ie4vdDS" colab_type="text"
# Lambda School Data Science
#
# *Unit 1, Sprint 1, Module 3*
#
# ---
#
#
# + [markdown] id="kOI3YimSvopw" colab_type="text"
# # Join and Reshape Data
#
# - Student can concatenate data with pandas
# - Student can merge data with pandas
# - Student can understand and describe tidy data formatting
# - Student can use the `.melt()` and `.pivot()` functions to translate between wide and tidy data format.
#
# Helpful Links:
# - [Pandas Cheat Sheet](https://github.com/pandas-dev/pandas/blob/master/doc/cheatsheet/Pandas_Cheat_Sheet.pdf)
# - [Tidy Data](https://en.wikipedia.org/wiki/Tidy_data)
# - Combine Data Sets: Standard Joins
# - Tidy Data
# - Reshaping Data
# - Python Data Science Handbook
# - [Chapter 3.6](https://jakevdp.github.io/PythonDataScienceHandbook/03.06-concat-and-append.html), Combining Datasets: Concat and Append
# - [Chapter 3.7](https://jakevdp.github.io/PythonDataScienceHandbook/03.07-merge-and-join.html), Combining Datasets: Merge and Join
# - [Chapter 3.8](https://jakevdp.github.io/PythonDataScienceHandbook/03.08-aggregation-and-grouping.html), Aggregation and Grouping
# - [Chapter 3.9](https://jakevdp.github.io/PythonDataScienceHandbook/03.09-pivot-tables.html), Pivot Tables
# + [markdown] id="b_gXHprXvqVx" colab_type="text"
# # [Objective](#concat) Concatenate dataframes with pandas
#
#
# + [markdown] id="mPVHZevR04pV" colab_type="text"
# ## Overview
#
# "Concatenate" is a fancy word for joining two things together. For example, we can concatenate two strings together using the `+` operator.
# + id="NeAeYKwN08q3" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="e59085c2-1317-4d5b-bc0b-03e6399135b0"
'We can join/concatenate two strings together ' + 'using the "+" operator.'
# + [markdown] id="QIgxXomn7iHC" colab_type="text"
# When we "concatenate" two dataframes we will "stick them together" either by rows or columns. Lets look at some simple examples:
# + id="O6MbummV9kgH" colab_type="code" colab={}
import pandas as pd
# + id="q1aKHYuH8BTX" colab_type="code" colab={}
df1 = pd.DataFrame({'a': [1,2,3,4], 'b': [4,5,6,7], 'c': [7,8,9,10]})
df2 = pd.DataFrame({'a': [6,4,8,7], 'b': [9,4,3,2], 'c': [1,6,2,9]})
# + id="blLFOpK-8Zwq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 166} outputId="e84ef389-4b24-45aa-ea4a-ddb6a12ec53c"
df1.head()
# + id="olRWT5VK8bl2" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 166} outputId="f411cf92-ad28-4758-a86d-1a4b92b5d2a9"
df2.head()
# + [markdown] id="FBh-mGzI8k3l" colab_type="text"
# ### Concatenate by Rows
#
# concatenating by rows is the default behavior of `pd.concat()` This is often the most common form of concatenation.
# + id="QCw6DJxR8m6T" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 284} outputId="088eeaff-2db5-489f-ce4f-3ff08b238e56"
# Pass in the dataframes that we want to concatenate as as list.
concatenated_by_rows = pd.concat([df1, df2])
# Reset the index so that we don't have repeated row identifiers
concatenated_by_rows = concatenated_by_rows.reset_index()
concatenated_by_rows.head(8)
# + [markdown] id="EvZH9k-e8ohe" colab_type="text"
# ### Concatenate by Columns
# + id="-fCzFQxx9D7b" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 166} outputId="6d7eeda1-6152-4e17-de07-f9253f062ef6"
concatenated_by_columns = pd.concat([df1,df2], axis=1)
concatenated_by_columns.head()
# + [markdown] id="2-NxpMCr9WOS" colab_type="text"
# When concatenating dataframes, it is done using the column headers and row index values to match rows up. If these don't match up, then `NaN` values will be added where matches can't be found.
# + id="luR-nvD99tBa" colab_type="code" colab={}
df3 = pd.DataFrame({'a': [4,3,2,1], 'b': [4,5,6,7], 'c': [7,8,9,10]})
df4 = pd.DataFrame({'a': [6,4,8,7,8], 'b': [9,4,3,2,1], 'd': [1,6,2,9,5]})
# + id="Bj-FdzVf97mn" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 166} outputId="157363e0-bac5-4cd9-b392-5614d2e733fb"
df3.head()
# + id="vfZ_wekl99-e" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 195} outputId="bf7cb2bf-d4c9-41d4-b50c-3ad9676b5b02"
df4.head()
# + [markdown] id="pOuoIdey-kCD" colab_type="text"
# ### Concatenate by rows when not all column headers match
# + id="5FpZdgat-EQD" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 468} outputId="415d7ea1-7feb-4f95-96af-f3f309037408"
concatenated_by_rows = pd.concat([df3, df4])
concatenated_by_rows.head(9)
# + [markdown] id="Al203GNp-qVS" colab_type="text"
# ### Concatenate by columns when not all row indexes match
# + id="lc2ngk3O-YCv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 195} outputId="ecbb8055-68cf-433e-a923-98f244541dae"
concatenated_by_columns = pd.concat([df3, df4], axis=1)
concatenated_by_columns.head()
# + [markdown] id="jUpWwpdSBJGd" colab_type="text"
# Whenever we are combining dataframes, if appropriate values cannot be found based on the rules of the method we are using, then missing values will be filled with `NaNs`.
# + [markdown] id="k8YGJ8Wm1AG3" colab_type="text"
# ## Follow Along
#
#
# + [markdown] id="hbH8CcozBcyI" colab_type="text"
# We’ll work with a dataset of [3 Million Instacart Orders, Open Sourced](https://tech.instacart.com/3-million-instacart-orders-open-sourced-d40d29ead6f2)!
#
# The files that we will be working with are in a folder of CSVs, we need to load that folder of CSVs, explore the CSVs to make sure that we understand what we're working with, and where the important data lies, and then work to combine the dataframes together as necessary.
#
#
#
# Our goal is to reproduce this table which holds the first two orders for user id 1.
#
# + id="3xugHGV5C60D" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 312} outputId="e573b368-2bb8-4a5f-a456-d4df7cf6a196"
from IPython.display import display, Image
url = 'https://cdn-images-1.medium.com/max/1600/1*vYGFQCafJtGBBX5mbl0xyw.png'
example = Image(url=url, width=600)
display(example)
# + id="4E3wKrdTChuC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 202} outputId="0607a570-6045-4ab4-8905-5fbf0abd8f3e"
# !wget https://s3.amazonaws.com/instacart-datasets/instacart_online_grocery_shopping_2017_05_01.tar.gz
# + id="dkxqMNGrDJrM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 235} outputId="2f6b1758-e0bf-4cf6-c916-b6c2fd949e03"
# !tar --gunzip --extract --verbose --file=instacart_online_grocery_shopping_2017_05_01.tar.gz
# + id="HI5E_tNjCjsV" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="cf769d16-43d2-41fd-b261-af1b64df9c58"
# %cd instacart_2017_05_01
# + id="qaDdXbQqCnGc" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 118} outputId="1b1cb7ab-cc3e-4752-cf1c-537b5b41c22a"
# !ls -lh *.csv
# + id="Z-e2iYLUSDY7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 312} outputId="434fa20e-e4bb-4a96-ddf6-0935153594ab"
display(example)
# + [markdown] id="MxwByNLoEG9p" colab_type="text"
# ### aisles
#
# We don't need anything from aisles.csv
# + id="pLmGMr_rCoi-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 212} outputId="35beee96-7177-490e-d99f-911f9bfed6b5"
aisles = pd.read_csv('aisles.csv')
print(aisles.shape)
aisles.head()
# + [markdown] id="oTPRZsLvENgJ" colab_type="text"
# ### departments
#
# We don't need anything from departments.csv
# + id="fRVjREe8D6yj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 212} outputId="0ab5c816-2e79-4a42-dab5-b08407142db3"
departments = pd.read_csv('departments.csv')
print(departments.shape)
departments.head()
# + [markdown] id="U38aOM6nEWOe" colab_type="text"
# ### order_products__prior
#
# We need:
# - order id
# - proudct id
# - add to cart order
#
# Everything except for 'reordered'
# + id="-CMWcWSiD8aW" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 212} outputId="0a9df715-f7f5-42ef-deb8-38d6ebcde8f1"
order_products__prior = pd.read_csv('order_products__prior.csv')
print(order_products__prior.shape)
order_products__prior.head()
# + [markdown] id="_KRCdSl5E63N" colab_type="text"
# ### order_products__train
#
# We need:
# - order id
# - proudct id
# - add to cart order
#
# Everything except for 'reordered'
#
# Do you see anything similar between order_products__train and order_products__prior?
#
#
# + id="2Pq7lgVUD-a-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 212} outputId="ed47e00d-9381-4bcf-fcf1-ca55f6ef7729"
order_products__train = pd.read_csv('order_products__train.csv')
print(order_products__train.shape)
order_products__train.head()
# + [markdown] id="V2cdftjYFj1k" colab_type="text"
# ### orders
#
# We need:
# - order id
# - user id
# - order number
# - order dow
# - order hour of day
# + id="t9PyqoneEBPd" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 212} outputId="805fa275-1f3e-4a7f-ff6d-4adafcca6015"
orders = pd.read_csv('orders.csv')
print(orders.shape)
orders.head()
# + [markdown] id="yXyRiuIdFmGU" colab_type="text"
# ### products
#
# We need:
# - product id
# - product name
# + id="3J917C0NEDhG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 212} outputId="efd663dc-8e82-4eb7-ca10-b8a0b88933c2"
products = pd.read_csv('products.csv')
print(products.shape)
products.head()
# + [markdown] id="ALhA76X1GkgY" colab_type="text"
# ## Concatenate order_products__prior and order_products__train
#
#
#
# + id="QU4nkwnPGz4A" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="7749efee-d2a4-47fc-9323-fb4228f47f75"
order_products__prior.shape
# + id="XrBZ0y8TG09_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="f3c6b2d6-b678-42ed-c2b3-d495fbca7f2a"
order_products__train.shape
# + id="gZk8V7yxG2Qg" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 212} outputId="beb0f397-d438-4b5a-d2fc-a06848769b35"
order_products = pd.concat([order_products__prior, order_products__train])
print(order_products.shape)
order_products.head()
# + [markdown] id="XSiHrHuj1ME-" colab_type="text"
# ## Challenge
#
# Concatenating dataframes means to stick two dataframes together either by rows or by columns. The default behavior of `pd.concat()` is to take the rows of one dataframe and add them to the rows of another dataframe. If we pass the argument `axis=1` then we will be adding the columns of one dataframe to the columns of another dataframe.
#
# Concatenating dataframes is most useful when the columns are the same between two dataframes or when we have matching row indices between two dataframes.
#
# Be ready to use this method to combine dataframes together during your assignment.
# + [markdown] id="17PV3bEtz449" colab_type="text"
# # [Objective](#merge) Merge dataframes with pandas
#
#
# + [markdown] id="DAiMlm5Q05LW" colab_type="text"
# ## Overview
# + id="oH4J87G4LZjd" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 313} outputId="85b50959-7a7f-4ce0-abc0-1f479a318111"
display(example)
# + [markdown] id="p1o6R29VLwwu" colab_type="text"
# Before we can continue we need to understand where the data in the above table is coming from and what why specific pieces of data are held in the specific dataframes.
#
# Each of these CSVs has a specific unit of observation (row). The columns that we see included in each CSV were selected purposefully. For example, everything each row of the `orders` dataframe is a specific and unique order -telling us who made the order, and when they made it. Every row in the `products` dataframe tells us about a specific and unique product that thestore offers. And everything in the `order_products` dataframe tells us about how products are associated with specific orders -including when the product was added to the shopping cart.
#
# ### The Orders Dataframe
#
# Holds information about specific orders, things like who placed the order, what
#
# - user_id
# - order_id
# - order_number
# - order_dow
# - order_hour_of_day
#
# ### The Products Dataframe
#
# Holds information about individual products.
#
# - product_id
# - product_name
#
# ### The Order_Products Dataframe
#
# Tells us how products are associated with specific orders since an order is a group of products.
#
# - order_id
# - product_id
# - add_to_cart_order
#
# As we look at the table that we're trying to recreate, we notice that we're not looking at specific orders or products, but at a specific **USER**. We're looking at the first two orders for a specific user and the products associated with those orders, so we'll need to combine dataframes to get all of this data together into a single table.
#
# **The key to combining all of this information is that we need values that exist in both datasets that we can use to match up rows and combine dataframes.**
# + [markdown] id="g38DqtNj1BnI" colab_type="text"
# ## Follow Along
#
# We have two dataframes, so we're going to need to merge our data twice. As we approach merging datasets together we will take the following approach.
#
# 1) Identify which to dataframes we would like to combine.
#
# 2) Find columns that are common between both dataframes that we can use to match up information.
#
# 3) Slim down both of our dataframes so that they only relevant data before we merge.
#
# 4) Merge the dataframes.
# + id="yvtGLx2XWzTz" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 312} outputId="57ae9662-5e49-4496-b1ca-50b5fd97a063"
display(example)
# + [markdown] id="259BJMJ43Ka5" colab_type="text"
# ### First Merge
#
# 1) Combine `orders` and `order_products`
#
# 2) We will use the `order_id` column to match information between the two datasets
#
# 3) Lets slim down our dataframes to only the information that we need. We do this because the merge process is complex. Why would we merge millions of rows together if we know that we're only going to need 11 rows when we're done
#
# What specific conditions could we use to slim down the `orders` dataframe?
#
# `user_id == 1` and `order_id <=2`
#
# or
#
# `order_id == 2539329` or `order_id == 2398795`
# + id="q2mZmxF8XEwI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 166} outputId="cdffd1cd-3659-4e2c-ce95-d0aab1d77094"
df1.head()
# + id="geCgwtX4XIEt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 106} outputId="4453fd01-22fd-448a-e566-266a21420955"
# What if I only wanted the rows where column C is > 8
condition = (df1['c'] > 8)
df1[condition]
# + id="HU5MvyI2YSTN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 218} outputId="9794a353-9d89-49eb-8c92-abf5859b9158"
orders['user_id'] == 1
# + id="I6r8i8tN1H1S" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 123} outputId="0ed60b80-dcc3-49a1-fd94-d17721bb7d0f"
# An example of dataframe filtering
# Create a condition
condition = ((orders['user_id'] == 1) & (orders['order_number'] <=2))
# Pass that condition into the square brackets
# that we use to access portions of a dataframe
# only the rows where that condition evaluates to *TRUE*
# will be retained in the dataframe
orders_subset = orders[condition]
# Look at the subsetted dataframe
print(orders_subset.shape)
orders_subset.head()
# + id="cWUCmgx66Td3" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 123} outputId="7a91e818-c94b-4f37-8ff0-821d37a60c1b"
# We don't necessarily have to save our condition to the variable "condition"
# we can pass the condition into the square brackest directly
# I just wanted to be clear what was happening inside of the square brackets
orders_subset = orders[(orders['order_id'] == 2539329) | (orders['order_id'] == 2398795)]
print(orders_subset.shape)
orders_subset.head()
# + [markdown] id="1kQG4sxP6lod" colab_type="text"
# Remember there are multiple ways that we could have filtered this dataframe. We also could have done it by specific `order_id`s
#
# + id="8IA4Kwyw6vk6" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 123} outputId="5fa2db1b-bd1c-4c27-aed9-6764b28c96b0"
condition = (orders['order_id'] == 2539329) | (orders['order_id'] == 2398795)
orders_subset = orders[condition]
print(orders_subset.shape)
orders_subset.head()
# + [markdown] id="Nt8qiPCl7Lh8" colab_type="text"
# Now we'll filter down the order_products dataframe
#
# What conditions could we use for subsetting that table?
#
# We can use order_id again.
# + id="DHE_-PKs7e4s" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 390} outputId="2dcd2407-841c-4fd8-8343-af58d51b4fcb"
condition = (order_products['order_id'] == 2539329) | (order_products['order_id'] == 2398795)
order_products_subset = order_products[condition]
print(order_products_subset.shape)
order_products_subset.head(11)
# + id="rkX2DdgNZ52d" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 312} outputId="e6022e52-a0ff-47e9-d39f-94c78cbba8f7"
display(example)
# + [markdown] id="yfi9zxpR7ugQ" colab_type="text"
# 4) Now we're ready to merge these two tables together.
# + id="gnhC5A2m7yQz" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 373} outputId="89b7deae-a7ee-446e-d956-854db666b1c9"
# on = The column header for the unique identifier column that I'm using to
# match the two dataframes' information
# how = The way that I want any non-matching rows to be retained or not retained
# Which NaNs should be kept vs dropped
orders_and_products = pd.merge(orders_subset,
order_products_subset,
on='order_id',
how='inner')
# inner = do the merge, but drop any rows with NaNs
# outer = do the merge, but keep all the rows even the ones with NaNs
# right = do the merge, but only keep the rows with Nans that came from the
# right-hand dataframe
# left = do the merge, but only keep the rows with Nans that came from the
# left-hand dataframe
orders_and_products.head(11)
# + id="RHGqFlPJ80-k" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 312} outputId="a8ff16aa-791d-479c-fbe0-e780ed43547d"
display(example)
# + id="DVdIEQsQ8D6x" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 373} outputId="66f85ea8-ca16-4570-e4b8-00e310b62b92"
# Remove columns that we don't need
orders_and_products = orders_and_products.drop(['eval_set',
'reordered',
'days_since_prior_order'],
axis=1)
orders_and_products.head(11)
# + [markdown] id="gLDN1ueb8_pY" colab_type="text"
# Okay, we're looking pretty good, we're missing one more column `product_name` so we're going to need to merge one more time
#
# 1) merge `orders_and_products` with `products`
#
# 2) Use `product_id` as our identifier in both tables
#
# 3) We need to slim down the `products` dataframe
# + id="Hy0fJFKn8--C" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 218} outputId="6850a80f-f627-48eb-b057-52d43d1ac560"
orders_and_products['product_id']
# + id="7yz6FwtG9bhd" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 218} outputId="64858b7d-763d-4879-9c42-e5f173979cf6"
orders_and_products['product_id'].isin([196, 26088])
# + id="xd_hwRyC9s1a" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 284} outputId="ceae8ebc-f979-41c3-ce90-7437d4531f85"
condition = products['product_id'].isin(orders_and_products['product_id'])
products_subset = products[condition]
products_subset
# + id="LjdgAOtb96LF" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 440} outputId="83af746c-5220-4b7b-b770-0e1798c37a1a"
final = pd.merge(orders_and_products, products_subset, on='product_id', how='inner')
final
# + id="vfOdijKidc4_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 312} outputId="29b54bca-7571-4d04-bf31-25e21280723a"
display(example)
# + [markdown] id="75lbOTf3-csq" colab_type="text"
# ### Some nitpicky cleanup:
# + id="2NODYWri_CYp" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 373} outputId="8d92e4e7-fc13-4d7d-c3e5-e3a97bdbe83f"
# reorder columns
final = final[['user_id', 'order_id', 'order_number', 'order_dow', 'order_hour_of_day', 'add_to_cart_order', 'product_id', 'product_name']]
final
# + id="_dL8nGeN-eiH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 373} outputId="69664338-6597-4960-f89d-1d6e115bacf2"
# sort rows
final = final.sort_values(by=['order_number', 'add_to_cart_order'])
final
# + id="RSdatcjM-r2B" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 373} outputId="270e6527-911d-46c2-a59a-3c4575a5f43d"
# remove underscores from column headers
final.columns = [column.replace('_', ' ') for column in final]
final
# + id="a0blpL6O-99U" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 313} outputId="87a1f297-a0f1-44c3-cb60-ca9579e6b494"
display(example)
# + [markdown] id="sNS2FhVW1NxV" colab_type="text"
# ## Challenge
#
# Review this Chis Albon documentation about [concatenating dataframes by row and by column](https://chrisalbon.com/python/data_wrangling/pandas_join_merge_dataframe/) and then be ready to master this function and practice using different `how` parameters on your assignment.
# + [markdown] id="xTz5WPngz5BA" colab_type="text"
# # [Objective](#tidy) Learn Tidy Data Format
# + [markdown] id="rUL61OCE06Dd" colab_type="text"
# ## Overview
#
# ### Why reshape data?
#
# #### Some libraries prefer data in different formats
#
# For example, the Seaborn data visualization library prefers data in "Tidy" format often (but not always).
#
# > "[Seaborn will be most powerful when your datasets have a particular organization.](https://seaborn.pydata.org/introduction.html#organizing-datasets) This format ia alternately called “long-form” or “tidy” data and is described in detail by <NAME>. The rules can be simply stated:
#
# > - Each variable is a column
# - Each observation is a row
#
# > A helpful mindset for determining whether your data are tidy is to think backwards from the plot you want to draw. From this perspective, a “variable” is something that will be assigned a role in the plot."
#
# #### Data science is often about putting square pegs in round holes
#
# Here's an inspiring [video clip from _Apollo 13_](https://www.youtube.com/watch?v=ry55--J4_VQ): “Invent a way to put a square peg in a round hole.” It's a good metaphor for data wrangling!
# + [markdown] id="NoUBeGKlAcCh" colab_type="text"
# ### Hadley Wickham's Examples
#
# From his paper, [Tidy Data](http://vita.had.co.nz/papers/tidy-data.html)
# + id="S_b6SOZz091T" colab_type="code" colab={}
# %matplotlib inline
import pandas as pd
import numpy as np
import seaborn as sns
table1 = pd.DataFrame(
[[np.nan, 2],
[16, 11],
[3, 1]],
index=['<NAME>', '<NAME>', '<NAME>'],
columns=['treatmenta', 'treatmentb'])
# + [markdown] id="gjEInTFNA54j" colab_type="text"
# "Table 1 provides some data about an imaginary experiment in a format commonly seen in the wild.
#
# The table has two columns and three rows, and both rows and columns are labelled."
# + id="fzoZDHtAA30k" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 136} outputId="77039484-7f7e-47fa-b4e6-04b072e447ee"
table1
# + [markdown] id="Q7lDPNIwA9t5" colab_type="text"
# "There are many ways to structure the same underlying data.
#
# Table 2 shows the same data as Table 1, but the rows and columns have been transposed. The data is the same, but the layout is different."
# + id="kQTFbHJWA_X2" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 106} outputId="d4866f00-369b-419d-f3c4-abd8d5e704e0"
table2 = table1.T
table2
# + [markdown] id="OfCVyRL3BI5h" colab_type="text"
# "Table 3 reorganises Table 1 to make the values, variables and obserations more clear.
#
# Table 3 is the tidy version of Table 1. Each row represents an observation, the result of one treatment on one person, and each column is a variable."
#
# | name | trt | result |
# |--------------|-----|--------|
# | <NAME> | a | - |
# | <NAME> | a | 16 |
# | <NAME> | a | 3 |
# | <NAME> | b | 2 |
# | <NAME> | b | 11 |
# | <NAME> | b | 1 |
# + [markdown] id="_bZaQTSfBp4s" colab_type="text"
# ## Follow Along
# + [markdown] id="9s42XuCqBN86" colab_type="text"
# ### Table 1 --> Tidy
#
# We can use the pandas `melt` function to reshape Table 1 into Tidy format.
# + id="aT8z1LRKhpiy" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 136} outputId="a04ded8b-c12f-4aa1-d72a-78d57aa3bfce"
table1
# + id="91Pw2zgkBHlg" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 136} outputId="a704fa0a-985c-4693-ebbb-2e3db92dffef"
# Take the row index, and add it as a new column
table1 = table1.reset_index()
table1.head()
# + id="UOlvjeBwBTNo" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 225} outputId="a564f966-1030-4e7e-ca97-74de7b9c2ed0"
# What is the unique identifier for each row
# Where is the data at that I want to be in my single "tidy" column
# MELT FUNCTION - Go from WIDE -> TIDY
tidy1 = table1.melt(id_vars='index', value_vars=['treatmenta', 'treatmentb'])
tidy1
# + id="hkXdIKDjBZC4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 225} outputId="ea821e51-b5b0-48dd-9d8a-2466c7b4028b"
# rename columns
tidy1 = tidy1.rename(columns={
'index': 'name',
'variable': 'trt',
'value': 'result'
})
tidy1
# + id="-BnEq18qBbBK" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 225} outputId="2b800cbf-2bae-4ad0-996b-d30e0b1fc0d8"
tidy1.trt = tidy1.trt.str.replace('treatment', '')
tidy1
# + [markdown] id="UuFviZYbBebc" colab_type="text"
# ### Tidy --> Table 1
#
# The `pivot_table` function is the inverse of `melt`.
# + id="LdfbFRI5Bgnh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 166} outputId="1ffe9ac6-e228-418c-f626-573a8d7913ad"
# index: unique identifier
# columns: What do you want to differentiate the columns in wide format
# values: Where are the numbers at - go in the middle of the wide dataframe
wide = tidy1.pivot_table(index='name', columns='trt', values='result')
wide
# + [markdown] id="_sfVVJg91Ri9" colab_type="text"
# ## Challenge
#
# On your assignment, be prepared to take table2 (the transpose of table1) and reshape it to be in tidy data format using `.melt()` and then put it back in "wide format" using `.pivot_table()`
# + [markdown] id="9H9XAmbTz5D6" colab_type="text"
# # [Objective](#melt-pivot) Transition between tidy and wide data formats with `.melt()` and `.pivot()`.
# + [markdown] id="346ZYlh7vsbx" colab_type="text"
# ## Overview
#
# Tidy data format can be particularly useful with certain plotting libraries like Seaborn for example. Lets practice reshaping our data and show how this can be extremely useful in preparing our data for plotting.
#
# Remember that tidy data format means:
#
# - Each variable is a column
# - Each observation is a row
#
# A helpful mindset for determining whether your data are tidy is to think backwards from the plot you want to draw. From this perspective, a “variable” is something that will be assigned a role in the plot." When plotting, this typically means that the values that we're most interested in and that represent the same thing will all be in a single column. You'll see that in the different examples that we show. The important data will be in a single column.
#
#
# + id="oc3h1LK6ulP-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 153} outputId="2b1d3fc4-46cc-4f43-904e-14b16bdb8878"
# Look at some of the awesome out-of-the-box seaborn functionality:
import seaborn as sns
sns.catplot(x='trt', y='result', col='name',
kind='bar', data=tidy1, height=2);
# + [markdown] id="RDs7HSDtvwp7" colab_type="text"
# ## Follow Along
#
# Now with Instacart Data. We're going to try and reproduce a small part of this visualization:
# + id="UYaAh9i2Cth8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 383} outputId="3af56ed3-9445-4186-a340-f016983228bf"
from IPython.display import display, Image
url = 'https://cdn-images-1.medium.com/max/1600/1*wKfV6OV-_1Ipwrl7AjjSuw.png'
example = Image(url=url, width=600)
display(example)
# + [markdown] id="cWA17E72Csb8" colab_type="text"
# Instead of a plot with 50 products, we'll just do two — the first products from each list
# - Half And Half Ultra Pasteurized
# - Half Baked Frozen Yogurt
#
# So, given a `product_name` we need to calculate its `order_hour_of_day` pattern.
# + id="Afch5TnbvzYH" colab_type="code" colab={}
products = pd.read_csv('products.csv')
order_products = pd.concat([pd.read_csv('order_products__prior.csv'),
pd.read_csv('order_products__train.csv')])
orders = pd.read_csv('orders.csv')
# + [markdown] id="a-wZZuvJC9XR" colab_type="text"
# ### Subset and Merge
#
# One challenge of performing a merge on this data is that the `products` and `orders` datasets do not have any common columns that we can merge on. Due to this we will have to use the `order_products` dataset to provide the columns that we will use to perform the merge.
#
# Here's the two products that we want to work with.
# + id="8kiwQevWC-ky" colab_type="code" colab={}
product_names = ['Half Baked Frozen Yogurt', 'Half And Half Ultra Pasteurized']
# + [markdown] id="qQS97tQ4DMcU" colab_type="text"
# Lets remind ourselves of what columns we have to work with:
# + id="ObLpUAJdDDKs" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="18d21169-e5a9-4527-b3fb-eb0668a19db6"
products.columns.to_list()
# + id="3bMVJbGMDGeY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 134} outputId="2c4f07ab-30ab-45b7-bf53-03a9a808aa14"
orders.columns.to_list()
# + [markdown] id="NWeDbbc4DSQ9" colab_type="text"
# This might blow your mind, but we're going to subset the dataframes to select specific columns **and** merge them all in one go. Ready?
# + id="jYG576viDYTY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="c1f1f1a4-3846-43c8-f568-000490faee4e"
order_products.columns.to_list()
# + [markdown] id="nyCeGkhyDlR0" colab_type="text"
# Ok, so we were a little bit lazy and probably should have subsetted our the rows of our dataframes before we merged them. We are going to filter after the fact. This is something that you can try out for practice. Can you figure out how to filter these dataframes **before** merging rather than after?
# + id="GRaUjNYQDqlv" colab_type="code" colab={}
merged = (products[['product_id', 'product_name']]
.merge(order_products[['order_id', 'product_id']])
.merge(orders[['order_id', 'order_hour_of_day']]))
# + id="HkvxpkkdlRSR" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 195} outputId="969b2d32-00db-45ae-f671-1f5ec6ba2a2b"
merged.head()
# + [markdown] id="PsCbAF1aD6mI" colab_type="text"
# Again, there are multiple effective ways to write conditions.
# + id="3Q3-vSjFD5r2" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 212} outputId="dc591de7-1847-4af9-d0fa-fb6022a358d0"
condition = (merged['product_name'] == 'Half Baked Frozen Yogurt') | (merged['product_name'] == 'Half And Half Ultra Pasteurized')
merged = merged[condition]
print(merged.shape)
merged.head()
# + id="J4IxDetIELzM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 212} outputId="c292dbe5-49a3-43a0-fc04-2ebd99206560"
product_names = ['Half Baked Frozen Yogurt', 'Half And Half Ultra Pasteurized']
condition = merged['product_name'].isin(product_names)
subset = merged[condition]
print(subset.shape)
subset.head()
# + [markdown] id="cuGmrZ52ECOS" colab_type="text"
# ### 4 ways to reshape and plot
#
#
# + id="0FKFamTREFiw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 384} outputId="bc5c8b56-7196-42b0-a004-23a5e072d957"
display(example)
# + id="nUSwmXAtmVxm" colab_type="code" colab={}
froyo = subset[subset['product_name']=='Half Baked Frozen Yogurt']
cream = subset[subset['product_name']=='Half And Half Ultra Pasteurized']
# + [markdown] id="-5udAYCYEQK5" colab_type="text"
# 1) The `.value_counts()` approach.
#
# Remember, that we're trying to get the key variables (values) listed as a single column.
# + id="mZih8j2QEIb9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 420} outputId="f32183ef-5a82-4583-b7c7-ba70148638a9"
cream['order_hour_of_day'].value_counts(normalize=True).sort_index()
# + id="RVvLVPgeEZXK" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 437} outputId="3d69bd9f-c295-49eb-ae1f-0e78c0f37fd2"
froyo['order_hour_of_day'].value_counts(normalize=True).sort_index()
# + id="xdiov3Vdmou8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 265} outputId="b5ee572d-7cc6-4317-f4a1-162bfd3e22e3"
import matplotlib.pyplot as plt
(cream['order_hour_of_day']
.value_counts(normalize=True)
.sort_index()
.plot())
# plt.show()
(froyo['order_hour_of_day']
.value_counts(normalize=True)
.sort_index()
.plot());
# + [markdown] id="35cWYAYJEdNU" colab_type="text"
# 2) Crosstab
# + id="q5LgBIUSEjCt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 787} outputId="f156f30b-1f44-4943-8694-18d7f4284036"
pd.crosstab(subset['order_hour_of_day'],
subset['product_name'],
normalize='columns').plot()
# + id="hxH0cGLjnHn1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 280} outputId="d6cc480f-f2a6-49e3-fb1c-adfcb6615ffe"
pd.crosstab(subset['order_hour_of_day'],
subset['product_name'],
normalize='columns').plot();
# + [markdown] id="AtzievA2El7W" colab_type="text"
# 3) Pivot Table
# + id="t3YVUT9znR2y" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 195} outputId="4c23354c-bfad-4be8-9f40-0e1827bbec51"
subset.head()
# + id="u1zTNxRWEndk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 283} outputId="95708730-30ce-4c2c-9f5f-495e72b09c67"
subset.pivot_table(index='order_hour_of_day',
columns='product_name',
values='order_id',
aggfunc=len).plot();
# + [markdown] id="Q-uexEZoErje" colab_type="text"
# 4) Melt
#
# We've got to get it into wide format first. We'll use a crosstab which is a specific type of pivot_table.
# + id="l2YFirZPE2DL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 787} outputId="6d38f5e5-9d8d-45cf-abf3-bde096364964"
# My crosstab is WIDE format
wide = pd.crosstab(subset['order_hour_of_day'],
subset['product_name'],
normalize=True)
wide
# + id="tGsUDMq8E6TP" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="bc537f43-9842-40c0-fcce-e92bdbd1c285"
melted = wide.reset_index().melt(id_vars='order_hour_of_day').rename(columns={
'order_hour_of_day': 'Hour of Day Ordered',
'product_name': 'Product',
'value': 'Percent of Orders by Product'
})
melted
# + [markdown] id="blu1emcSE95R" colab_type="text"
# Now, with Seaborn:
# + id="gTusUe1WE-57" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 369} outputId="fb178b6d-29f6-4df3-c8a2-2ed16e97ae70"
sns.relplot(x='Hour of Day Ordered',
y='Percent of Orders by Product',
hue='Product',
data=melted,
kind='line');
# + id="-87wbxbunwK3" colab_type="code" colab={}
|
Join_and_Reshape_Data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
num = int(input("Enter a number to generate its pattern = "))
for i in range(1,num + 1):
for j in range(1,i + 1):
print (j, end = " ")
print()
num = 2
for i in range(2,50):
j=2
while (j <= (1/2)):
if (i %j ==0):
break
j+= 1
if ( j> 1/j):
print (i, "is a prime number")
print ("Bye Bye!!")
num = int(input("Enter a number: "))
fact = 1
if num < 0:
print("Sorry, factorial does not exist for negative number")
elif num == 0:
print("The factorial of 0 is 1")
else:
for i in range(1, num + 1):
fact = fact * i
print("factorial of ",num , "is", fact)
num = int(input("Enter the number to be checked: "))
flag = 0
if num > 1 :
for i in range(2, int(num / 2)):
if (num % i == 0):
flag = 1
if flag == 1:
print(num, "is not a prime number")
else:
print(num, "is a prime number")
else:
print ("Entered number is <= 1, think again!")
num = int(input("Enter a number to find its factor: "))
print (1, end= ' ')
factor = 2
while factor<= num/2 :
if num % factor == 0:
print(factor, end=' ')
factor += 1
print (num, end=' ')
|
Class-Exercise-I- PeseAlo.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.0 64-bit ('ai-env')
# metadata:
# interpreter:
# hash: ee89ffdf677b068b3969c9c92fc557abd8fe9bdccee3c1b3432324df722c1402
# name: python3
# ---
# ### Generating human faces with Adversarial Networks (5 points)
# <img src="https://www.strangerdimensions.com/wp-content/uploads/2013/11/reception-robot.jpg" width=320>
# This time we'll train a neural net to generate plausible human faces in all their subtlty: appearance, expression, accessories, etc. 'Cuz when us machines gonna take over Earth, there won't be any more faces left. We want to preserve this data for future iterations. Yikes...
#
# Based on https://github.com/Lasagne/Recipes/pull/94 .
#
# +
import matplotlib.pyplot as plt
# %matplotlib inline
import numpy as np
plt.rcParams.update({'axes.titlesize': 'small'})
from sklearn.datasets import load_digits
#The following line fetches you two datasets: images, usable for autoencoder training and attributes.
#Those attributes will be required for the final part of the assignment (applying smiles), so please keep them in mind
from lfw_dataset import fetch_lfw_dataset
data,attrs = fetch_lfw_dataset(dimx=36, dimy=36)
# # !wget https://raw.github.com/yandexdataschool/Practical_DL/fall20/week08_generative/lfw_dataset.py
#preprocess faces
data = np.float32(data).transpose([0,3,1,2]) / 255.
IMG_SHAPE = data.shape[1:]
# -
#print random image
plt.imshow(data[np.random.randint(data.shape[0])].transpose([1,2,0]),
cmap="gray", interpolation="none")
# # Generative adversarial nets 101
#
# <img src="https://raw.githubusercontent.com/torch/torch.github.io/master/blog/_posts/images/model.png" width=320px height=240px>
#
# Deep learning is simple, isn't it?
# * build some network that generates the face (small image)
# * make up a __measure__ of __how good that face is__
# * optimize with gradient descent :)
#
#
# The only problem is: how can we engineers tell well-generated faces from bad? And i bet you we won't ask a designer for help.
#
# __If we can't tell good faces from bad, we delegate it to yet another neural network!__
#
# That makes the two of them:
# * __G__enerator - takes random noize for inspiration and tries to generate a face sample.
# * Let's call him __G__(z), where z is a gaussian noize.
# * __D__iscriminator - takes a face sample and tries to tell if it's great or fake.
# * Predicts the probability of input image being a __real face__
# * Let's call him __D__(x), x being an image.
# * __D(x)__ is a predition for real image and __D(G(z))__ is prediction for the face made by generator.
#
# Before we dive into training them, let's construct the two networks.
# +
import torch, torch.nn as nn
import torch.nn.functional as F
use_cuda = torch.cuda.is_available()
print("Torch version:", torch.__version__)
if use_cuda:
print("Using GPU")
else:
print("Not using GPU")
# +
def sample_noise_batch(batch_size):
noise = torch.randn(batch_size, CODE_SIZE)
return noise.cuda() if use_cuda else noise.cpu()
class Reshape(nn.Module):
def __init__(self, shape):
nn.Module.__init__(self)
self.shape=shape
def forward(self,input):
return input.view(self.shape)
# +
CODE_SIZE = 256
from itertools import count
# automatic layer name maker. Don't do this in production :)
# лол а зачем так вообще писать в учебных материалах м?
ix = ('layer_%i'%i for i in count())
generator = nn.Sequential()
generator.add_module(next(ix), nn.Linear(CODE_SIZE, 10*8*8))
generator.add_module(next(ix), nn.ELU())
generator.add_module(next(ix), Reshape([-1, 10, 8, 8]))
generator.add_module(next(ix), nn.ConvTranspose2d(10, 64, kernel_size=(5,5)))
generator.add_module(next(ix), nn.ELU())
generator.add_module(next(ix), nn.ConvTranspose2d(64, 64, kernel_size=(5,5)))
generator.add_module(next(ix), nn.ELU())
generator.add_module(next(ix), nn.Upsample(scale_factor=2))
generator.add_module(next(ix), nn.ConvTranspose2d(64, 32, kernel_size=(5,5)))
generator.add_module(next(ix), nn.ELU())
generator.add_module(next(ix), nn.ConvTranspose2d(32, 32, kernel_size=(5,5)))
generator.add_module(next(ix), nn.ELU())
generator.add_module(next(ix), nn.Conv2d(32, 3, kernel_size=(5,5)))
if use_cuda: generator.cuda()
# -
generated_data = generator(sample_noise_batch(5))
assert tuple(generated_data.shape)[1:] == IMG_SHAPE, "generator must output an image of shape %s, but instead it produces %s"%(IMG_SHAPE,generated_data.shape)
# ### Discriminator
# * Discriminator is your usual convolutional network with interlooping convolution and pooling layers
# * The network does not include dropout/batchnorm to avoid learning complications.
# * We also regularize the pre-output layer to prevent discriminator from being too certain.
# +
def sample_data_batch(batch_size):
idxs = np.random.choice(np.arange(data.shape[0]), size=batch_size)
batch = torch.tensor(data[idxs], dtype=torch.float32)
return batch.cuda() if use_cuda else batch.cpu()
# a special module that converts [batch, channel, w, h] to [batch, units]
class Flatten(nn.Module):
def forward(self, input):
return input.view(input.shape[0], -1)
# +
# discriminator = nn.Sequential()
# (W−F+2P)/S+1 => output image size
# W - image size, F - kernel size, P - padding size, S - stride size
### YOUR CODE - create convolutional architecture for discriminator
### Note: please start simple. A few convolutions & poolings would do, inception/resnet is an overkill
# discriminator.add_module("disc_logit", nn.Linear(50, 1))
# (36 - 3 + 2)/1 + 1 =
# 36 -> 18 -> 9
class Net(nn.Module):
def __init__(self, batch_size = 5):
super(Net, self).__init__()
self.batch_size = batch_size
self.conv1 = nn.Conv2d(3, 16, 3, padding=1)
self.conv2 = nn.Conv2d(16, 32, 3, padding=1)
self.flat = nn.Flatten()
self.pool = nn.MaxPool2d(2, 2)
self.fc1 = nn.Linear(32 * 9 * 9, 500)
self.fc2 = nn.Linear(500, 1)
self.dropout = nn.Dropout(0.25)
def forward(self, x):
x = self.pool(F.relu(self.conv1(x)))
x = self.pool(F.relu(self.conv2(x)))
x = self.flat(x)
x = self.dropout(x)
x = F.relu(self.fc1(x))
x = self.dropout(x)
x = self.fc2(x)
return x
discriminator = Net()
if use_cuda: discriminator.cuda()
discriminator(sample_data_batch(5))
# -
sample_data_batch(5).shape
# # Training
#
# We train the two networks concurrently:
# * Train __discriminator__ to better distinguish real data from __current__ generator
# * Train __generator__ to make discriminator think generator is real
# * Since discriminator is a differentiable neural network, we train both with gradient descent.
#
# 
#
# Training is done iteratively until discriminator is no longer able to find the difference (or until you run out of patience).
#
#
# ### Tricks:
# * Regularize discriminator output weights to prevent explosion
# * Train generator with __adam__ to speed up training. Discriminator trains with SGD to avoid problems with momentum.
# * More: https://github.com/soumith/ganhacks
#
def generator_loss(noise):
"""
1. generate data given noise
2. compute log P(real | gen noise)
3. return generator loss (should be scalar)
"""
generated_data = generator(noise)
disc_on_generated_data = discriminator(generated_data)
logp_gen_is_real = F.logsigmoid(disc_on_generated_data)
loss = -torch.mean(-logp_gen_is_real)
return loss
# +
loss = generator_loss(sample_noise_batch(32))
print(loss)
assert len(loss.shape) == 0, "loss must be scalar"
# -
def discriminator_loss(real_data, generated_data):
"""
1. compute discriminator's output on real & generated data
2. compute log-probabilities of real data being real, generated data being fake
3. return discriminator loss (scalar)
"""
disc_on_real_data = discriminator(real_data)
disc_on_fake_data = discriminator(generated_data)
logp_real_is_real = F.logsigmoid(disc_on_real_data)
logp_gen_is_fake = F.logsigmoid(disc_on_fake_data)
loss = torch.mean(-logp_real_is_real - logp_gen_is_fake)
return loss
# +
loss = discriminator_loss(sample_data_batch(32),
generator(sample_noise_batch(32)))
print(loss)
assert len(loss.shape) == 0, "loss must be scalar"
# -
# ### Auxilary functions
# Here we define a few helper functions that draw current data distributions and sample training batches.
# +
def sample_images(nrow, ncol, sharp=False):
images = generator(sample_noise_batch(batch_size=nrow*ncol))
images = images.data.cpu().numpy().transpose([0, 2, 3, 1])
if np.var(images)!=0:
images = images.clip(np.min(data),np.max(data))
for i in range(nrow*ncol):
plt.subplot(nrow,ncol,i+1)
if sharp:
plt.imshow(images[i], cmap="gray", interpolation="none")
else:
plt.imshow(images[i], cmap="gray")
plt.show()
def sample_probas(batch_size):
plt.title('Generated vs real data')
D_real = F.sigmoid(discriminator(sample_data_batch(batch_size)))
generated_data_batch = generator(sample_noise_batch(batch_size))
D_fake = F.sigmoid(discriminator(generated_data_batch))
plt.hist(D_real.data.cpu().numpy(),
label='D(x)', alpha=0.5, range=[0,1])
plt.hist(D_fake.data.cpu().numpy(),
label='D(G(z))', alpha=0.5, range=[0,1])
plt.legend(loc='best')
plt.show()
# -
# ### Training
# Main loop.
# We just train generator and discriminator in a loop and draw results once every N iterations.
#optimizers
disc_opt = torch.optim.SGD(discriminator.parameters(), lr=5e-3)
gen_opt = torch.optim.Adam(generator.parameters(), lr=1e-4)
# +
from IPython import display
from tqdm import tnrange
batch_size = 100
for epoch in tnrange(50000):
# Train discriminator
for i in range(5):
real_data = sample_data_batch(batch_size)
fake_data = generator(sample_noise_batch(batch_size))
loss = discriminator_loss(real_data, fake_data)
disc_opt.zero_grad()
loss.backward()
disc_opt.step()
# Train generator
noise = sample_noise_batch(batch_size)
loss = generator_loss(noise)
gen_opt.zero_grad()
loss.backward()
gen_opt.step()
if epoch %100==0:
display.clear_output(wait=True)
sample_images(2,3,True)
sample_probas(1000)
# +
plt.figure(figsize=[16, 24])
sample_images(16, 8)
# Note: a no-nonsense neural network should be able to produce reasonably good images after 15k iterations
# By "reasonably good" we mean "resembling a car crash victim" or better
# -
# ### Evaluation
# __The code below__ dumps a batch of images so that you could use them for precision/recall evaluation.
#
# Please, if you want to compare the result with the autoencoder result, generate the same number of images as for autoencoders for a fair comparison.
# +
num_images = <YOUR CODE>
batch_size = 100
all_images = []
for batch_i in range((num_images - 1) / batch_size + 1):
with torch.no_grad():
images = generator(sample_noise_batch(batch_size=batch_size))
images = images.data.cpu().numpy().transpose([0, 2, 3, 1])
if np.var(images)!=0:
images = images.clip(np.min(data), np.max(data))
all_images.append(images)
all_images = np.concatenate(all_images, axis=0)[:num_images]
np.savez("./gan.npz", Pictures=all_images)
|
week08_generative/adversarial_pytorch.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Importing pandas and setting the option to see all the rows
import numpy as np
from sklearn.impute import KNNImputer
import pandas as pd
pd.set_option('display.max_rows', None)
df = pd.read_csv('data_scientist_jobs.csv')
df.head()
# ## Removing the rows that don't have a salary estimate
df = df[df['Salary Estimate'] != '-1']
df.head()
# ## Removing the term "Glassdoor estimate" from the salary estimate
salary = df['Salary Estimate'].apply(lambda x: x.split('(')[0])
salary.head()
# ## Removing the dollar signs and K
minus_kd = salary.apply(lambda x: x.replace('K', '').replace('$',''))
# ## Adding one hot encoding for hourly rates and employer provided salary. Also getting the salary ranges by removing all kinds of text
df['hourly'] = df['Salary Estimate'].apply(lambda x: 1 if 'per hour' in x.lower() else 0)
df['employer_provided'] = df['Salary Estimate'].apply(lambda x: 1 if 'employer provided salary:' in x.lower() else 0)
range_ = minus_kd.apply(lambda x: x.lower().replace('per hour', '').replace('employer provided salary:', ''))
print(range_)
# Adding min, max and average salary columns from the salary ranges extracted above
df['min_salary'] = range_.apply(lambda x: int(x.split('-')[0]))
df['max_salary'] = range_.apply(lambda x: int(x.split('-')[1]))
df['avg_salary'] = (df.min_salary + df.max_salary)/2
df.head()
# Adding a column just for company name while separating the name from the rating
df['company_txt'] = df.apply(lambda x: x['Company Name'] if x['Rating'] <0 else x['Company Name'][:-4], axis = 1)
df.head()
# Adding a seprate column just for the state of the location of the job
df['job_state'] = df['Location'].apply(lambda x: x.split(',')[1])
#print(df.job_state)
# ## Adding a column with the age of the company
df['age'] = df.Founded.apply(lambda x: x if x <1 else 2020 - x)
df.age.value_counts()
# ## Extracting skills from the job descriptions and adding one hot encoding for each type of skill. 1st one is for python
df['python_'] = df['Job Description'].apply(lambda x: 1 if 'python' in x.lower() else 0)
df.python_.value_counts()
# OHE for R
df['R_'] = df['Job Description'].apply(lambda x: 1 if 'r studio' in x.lower() or 'r-studio' in x.lower() else 0)
df.R_.value_counts()
# OHE for spark
df['spark_'] = df['Job Description'].apply(lambda x: 1 if 'spark' in x.lower() else 0)
df.spark_.value_counts()
# OHE for aws
df['aws_'] = df['Job Description'].apply(lambda x: 1 if 'aws' in x.lower() else 0)
df.aws_.value_counts()
# OHE for excel
df['excel_'] = df['Job Description'].apply(lambda x: 1 if 'excel' in x.lower() else 0)
df.excel_.value_counts()
# ## OHE for the type of ownership. Private is coded as 1 while rest is 0
df['private'] = df['Type of ownership'].apply(lambda x: 1 if 'private' in x.lower() else 0)
df.private.value_counts()
# ## Filling in data for Rating by np.nan for knn-imputation
df['Rating'] = df['Rating'].apply(lambda x: np.nan if x==-1 else x)
# ## Doing the same for age
df['age'] = df['age'].apply(lambda x: np.nan if x==-1 else x)
# ## Converting the size column to numreic data by getting the average size of the companies
df.Size.value_counts()
df['Size'] = df['Size'].apply(lambda x: -1 if x == "Unknown" else x)
df['Size'] = df['Size'].apply(lambda x: -1 if x== "-1" else x)
df.Size.value_counts()
# +
size = df['Size']
size_max = []
size_min = []
for s in size:
if s == -1:
size_max.append(s)
size_min.append(s)
else:
s = str(s)
if s == "10000+ Employees":
size_max.append(10000)
size_min.append(10000)
else:
#print(s)
size_min.append(float(s.split(" ")[0]))
size_max.append(float(s.split(" ")[2]))
avg_size = []
for i in range(len(size_min)):
avg_size.append((size_min[i] + size_max[i])/2)
#print(avg_size)
df['avg_size'] = avg_size
# -
df['avg_size'] = df['avg_size'].apply(lambda x: np.nan if x==-1 else x)
df['Revenue'] = df['Revenue'].apply(lambda x: -1 if x[:7] == "Unknown" else x)
#print((df['Size'][0].split('E')[0]).split(' ')[2])
# ## Doing the same thing with revenues
df['Revenue'] = df['Revenue'].apply(lambda x: -1 if x == "-1" else x)
df.Revenue.value_counts()
# +
revenue = df['Revenue']
revenue_min = []
revenue_max = []
#revenue_max = df['Revenue'].apply(lambda x: x if x ==(-1 or "$10+ billion (USD)") else x.split(" ")[2][1:])
for r in revenue:
if r == -1:
revenue_min.append(r)
revenue_max.append(r)
elif r == "$10+ billion (USD)":
revenue_min.append(10000000000)
revenue_max.append(10000000000)
elif r == "$500 million to $1 billion (USD)":
revenue_min.append(500000000)
revenue_max.append(1000000000)
elif r == "Less than $1 million (USD)":
revenue_min.append(900000)
revenue_max.append(900000)
else:
fact = r.split(" ")[3][0]
if fact == "m":
factor = 1000000
else:
factor = 1000000000
revenue_min.append(float(r.split(" ")[0][1:])*factor)
revenue_max.append(float(r.split(" ")[2][1:])*factor)
avg_revenue = []
for i in range(len(revenue_min)):
avg_revenue.append((revenue_min[i] + revenue_max[i])/2)
#df['avg_revenue'] = avg_revenue
print(len(avg_revenue))
# -
df['avg_revenue'] = avg_revenue
df['avg_revenue'] = df['avg_revenue'].apply(lambda x: np.nan if x==-1 else x)
imputer = KNNImputer(n_neighbors=2)
continous_df = df[['avg_revenue', 'avg_size', 'age', 'Rating']]
continous_df = pd.DataFrame(imputer.fit_transform(continous_df), columns = continous_df.columns)
continous_df.shape
# # Modifying the data frame with all the imputed columns
df.columns
df = df.drop(['avg_revenue', 'avg_size', 'age', 'Rating'], axis = 1)
df.shape
df.columns
df.index = range(770)
df.head()
final_df = pd.concat([df, continous_df], axis=1)
final_df.shape
final_df.head()
# ## Exporting the data frame as a csv for further exploratory data analysis
final_df.to_csv('salary_data_cleaned.csv',index = False)
|
data_cleaning.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:LDCT]
# language: python
# name: conda-env-LDCT-py
# ---
# # 一、构建数据集和图像预处理
# +
import os
import numpy as np
import pydicom.filereader
from tqdm import trange
import torch
import torchvision
import torchvision.transforms as transforms
# -
# ## 1、继承torchvision.transforms.Compose类
# +
class My_Compose(transforms.Compose):
def __init__(self, transforms):
super().__init__(self)
self.transforms = transforms
def __call__(self, image):
for t in self.transforms:
image = t(image)
return image
class My_ToTensor(transforms.ToTensor):
def __init__(self):
super().__init__()
def __call__(self, image):
return self.to_tensor(image)
@staticmethod
def to_tensor(pic):
pic = pic[:, :, None]
img = torch.from_numpy(pic.transpose((2, 0, 1))).contiguous()
return img.float().div(4096)
class My_Normalize(transforms.Normalize):
def __init__(self, mean, std):
self.mean = mean
self.std = std
def __call__(self, image):
image = transforms.functional.normalize(image, mean=self.mean, std=self.std)
return image
# -
# ## 2、继承torch.utils.data.Dataset类
# + slideshow={"slide_type": "-"}
class Mydataset(torch.utils.data.Dataset):
'''读取LDCT和NDCT图像,进行归一化和标准化处理,返回(LDCT, NDCT) if "train = True"或(LDCT, NDCT, LD_ds) if "train = False"元组列表'''
def __init__(self, LDCT_root, NDCT_root, transform, normalize, train = True):
'''请指定LDCT和NDCT图像路径,以及图像预处理transform'''
super().__init__()
self.LDCT_root = LDCT_root
self.NDCT_root = NDCT_root
self.transform = transform
self.normalize = normalize
self.train = train
LDCT_list = os.listdir(LDCT_root)
NDCT_list = os.listdir(NDCT_root)
self.data_path = list(zip(LDCT_list, NDCT_list))
if len(LDCT_list) == len(NDCT_list):
self.len = len(LDCT_list)
else:
print('LDCT和NDCT图像数量不一致,请检查!')
def __getitem__(self, index):
'''根据索引获取image和label'''
LD, ND = self.data_path[index]
LD_path = self.LDCT_root + '\\' + LD
ND_path = self.NDCT_root + '\\' + ND
preprocessed = self.get_preprocess(LD_path, ND_path)
if self.train:
return preprocessed[0], preprocessed[1]
else:
return preprocessed[0], preprocessed[1], LD_path
def __len__(self):
'''返回数据集长度'''
return self.len
def get_preprocess(self, LD_path, ND_path):
'''读取图像并预处理'''
# 读取
LD_ds, LD_image = self.get_dcm_array(LD_path)
ND_ds, ND_image = self.get_dcm_array(ND_path)
# 归一化和标准化
LD_image = self.normalize(self.transform(LD_image))
ND_image = self.normalize(self.transform(ND_image))
return [LD_image, ND_image, LD_ds]
@staticmethod
def get_dcm_array(path):
'''读取dcm,并转换为像素为CT值'''
ds = pydicom.filereader.dcmread(path)
return ds, (ds.pixel_array).astype(np.int16)
# -
# ## 3、计算图像经过my_totensor处理后的mean和std
def cal_mean_and_std():
root = r'E:\NBIA\Sampling\LDCT_ALL\\'
dcm_list = os.listdir(root)
my_totensor = My_ToTensor()
transform = My_Compose([my_totensor])
mean_list = []
std_list = []
# print(len(dcm_list))
for idx in trange(len(dcm_list)):
dcm = dcm_list[idx]
ds = pydicom.filereader.dcmread(root+dcm)
img = (ds.pixel_array).astype(np.int16)
img = transform(img)
mean_list.append(img.mean())
std_list.append(img.std())
# mean
print('mean_len:', len(mean_list))
print('mean_sum:', np.array(mean_list).sum())
print('mean_mean:', np.array(mean_list).sum()/len(mean_list))
# std
print('\nstd_len:', len(std_list))
print('std_sum:', np.array(std_list).sum())
print('std_mean:', np.array(std_list).sum()/len(std_list))
print('\n' + '*'*50)
# ## 4、测试函数
def test():
LDCT_path = r'E:\Jupyter notebook\LDCT\LDCT'
NDCT_path = r'E:\Jupyter notebook\LDCT\NDCT'
my_totensor = My_ToTensor()
my_normalize = My_Normalize(0.131, 0.121)
transform = My_Compose([my_totensor])
normalize = My_Compose([my_normalize])
train_set = Mydataset(LDCT_root = LDCT_path, NDCT_root = NDCT_path,
matrix = 256,
transform = transform,
normalize = normalize)
train_loader = torch.utils.data.DataLoader(train_set,
batch_size = 2,
num_workers = 0,
shuffle = False,)
dataiter = iter(train_loader)
data = list(dataiter.next())
print(len(data))
for i in range(len(data[0])):
LD_img = data[0][i]
Res_img = data[1][i]
print('图像数量:', len(data[0]))
print('size:', LD_img.size() , Res_img.size())
print('type:', LD_img.type() , Res_img.type())
print('max:' , LD_img.max().item() , Res_img.max().item())
print('min:' , LD_img.min().item() , Res_img.min().item())
print('mean:', LD_img.mean().item(), Res_img.mean().item())
print('std:' , LD_img.std().item() , Res_img.std().item())
print('LD_img:', LD_img)
print('Res_img:', Res_img)
# # Test
if __name__ == '__main__':
# 计算图像经过my_totensor处理后的mean和std
cal_mean_and_std()
# 测试函数
# test()
|
Deep-Learning-and-Dicom-Viewer/LDCT/Dataset.ipynb
|
# # Sample Code
#
# Run this script. Edit it and make it your own.
# +
# measurements.py
"""
Functions to calculate distance between points using different metrics.
"""
import numpy as np
def crow(pointA, pointB):
"""
Distance between points A and B "as the crow flies."
pointA = (x1, y1)
pointB = (x2, y2)
returns sqrt( (x2-x1)**2 + (y2-y1)**2 )
"""
interval = np.sqrt( (pointA[0] - pointB[0])**2 + \
(pointA[1] - pointB[1])**2 )
return interval
def taxicab(pointA, pointB):
"""
Distance between points A and B "as the cab drives."
pointA = (x1, y1)
pointB = (x2, y2)
returns |x2-x1| + |y2-y1|
"""
interval = abs(pointB[0] - pointA[0]) + \
abs(pointB[1] - pointA[1])
return interval
def distance(pointA, pointB=(0,0), metric='taxi'):
"""
Return distance between points A and B. If metric is 'taxi', use taxicab
metric. Otherwise, use Euclidean distance.
pointA = (x1, y1)
pointB = (x2, y2)
"""
if metric == 'taxi':
return taxicab(pointA, pointB)
else:
return crow(pointA, pointB)
|
measurements.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#importing some useful packages
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
import numpy as np
import cv2
# %matplotlib inline
i_plot = False
# -
import os
cal_img_list = os.listdir("camera_cal/")
test_img_list = os.listdir("test_images/")
# +
def calibrate_camera():
nx = 9
ny = 6
# prepare object points, like (0,0,0), (1,0,0), (2,0,0) ....,(6,5,0)
objp = np.zeros((nx*ny,3), np.float32)
objp[:,:2] = np.mgrid[0:nx, 0:ny].T.reshape(-1,2)
# Arrays to store object points and image points from all the images.
objpoints = [] # 3d points in real world space
imgpoints = [] # 2d points in image plane.
# Make a list of calibration images
for image_file in cal_img_list:
img = mpimg.imread('camera_cal/' + image_file)
gray = cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)
# Find the chessboard corners
ret, corners = cv2.findChessboardCorners(gray, (nx,ny), None)
# If found, add object points, image points
if ret == True:
objpoints.append(objp)
imgpoints.append(corners)
# Do camera calibration given object points and image points
img_size = (img.shape[1], img.shape[0])
ret, mtx, dist, rvecs, tvecs = cv2.calibrateCamera(objpoints, imgpoints, img_size,None,None)
if 1:
dst = cv2.undistort(img, mtx, dist, None, mtx)
print(mtx)
print(dist)
for image_file in cal_img_list:
img = mpimg.imread('camera_cal/' + image_file)
dst = cv2.undistort(img, mtx, dist, None, mtx)
plt.figure()
plt.imshow(img)
plt.draw()
fig = plt.figure()
plt.imshow(dst)
plt.draw()
fig.savefig('output_images/calibrate.png')
break
return [mtx, dist]
[mtx, dist] = calibrate_camera()
print(mtx, dist)
if 0:
for img_file in test_img_list:
img = mpimg.imread('test_images/' + img_file)
dst = cv2.undistort(img, mtx, dist, None, mtx)
plt.figure()
plt.imshow(img)
plt.draw()
plt.figure()
plt.imshow(dst)
plt.draw()
break
# +
def region_of_interest(img, vertices):
"""
Applies an image mask.
Only keeps the region of the image defined by the polygon
formed from `vertices`. The rest of the image is set to black.
`vertices` should be a numpy array of integer points.
"""
#defining a blank mask to start with
mask = np.zeros_like(img)
#defining a 3 channel or 1 channel color to fill the mask with depending on the input image
if len(img.shape) > 2:
channel_count = img.shape[2] # i.e. 3 or 4 depending on your image
ignore_mask_color = (255,) * channel_count
else:
ignore_mask_color = 255
#filling pixels inside the polygon defined by "vertices" with the fill color
cv2.fillPoly(mask, vertices, ignore_mask_color)
# plt.figure()
# plt.imshow(mask)
# plt.draw()
#returning the image only where mask pixels are nonzero
masked_image = cv2.bitwise_and(img, mask)
return masked_image
# +
# Use color transforms, gradients, etc., to create a thresholded binary image.
# def create_binary_image(img, s_thresh=(170, 255), sx_thresh=(20, 100)):
def create_binary_image(img, s_thresh=(90, 255), sx_thresh=(20, 100), h_thresh=(5, 55)):
img = np.copy(img)
# Convert to HLS color space and select the channels
hls = cv2.cvtColor(img, cv2.COLOR_RGB2HLS)
h_channel = hls[:,:,0]
l_channel = hls[:,:,1]
s_channel = hls[:,:,2]
# Restrict relevant hues for saturation thresholding to filter out shadows
h_binary = np.zeros_like(h_channel)
h_binary[(h_channel >= h_thresh[0]) & (h_channel <= h_thresh[1])] = 1
# Compute the X gradient on the l channel
sobelx = cv2.Sobel(l_channel, cv2.CV_64F, 1, 0)
abs_sobelx = np.absolute(sobelx)
scaled_sobel = np.uint8(255*abs_sobelx/np.max(abs_sobelx))
# Threshold x gradient
sxbinary = np.zeros_like(scaled_sobel)
sxbinary[(scaled_sobel >= sx_thresh[0]) & (scaled_sobel <= sx_thresh[1])] = 1
# Threshold saturation channel
s_binary = np.zeros_like(s_channel)
s_binary[(s_channel >= s_thresh[0]) & (s_channel <= s_thresh[1])] = 1
# Stack each channel using the hue binary to mask shadows from the saturation binary
color_binary = np.dstack((np.zeros_like(sxbinary), sxbinary, s_binary & h_binary)) * 255
# Mask region of interest
ysize = color_binary.shape[0]
xsize = color_binary.shape[1]
vertices = np.array([[[0, ysize], [0.438*xsize, 0.627*ysize], [(1-0.438)*xsize, 0.627*ysize], [xsize, ysize]]], dtype=np.int32)
#returning the image only where mask pixels are nonzero
masked_color_binary = region_of_interest(color_binary, vertices)
return masked_color_binary
# for img_file in test_img_list:
# img = mpimg.imread('test_images/' + img_file)
# break
# binary_img = create_binary_image(img)
# plt.figure()
# plt.imshow(img)
# plt.draw()
# plt.figure()
# plt.imshow(binary_img)
# plt.draw()
# plt.figure()
# plt.imshow(masked_binary_img)
# plt.draw()
# +
# * Apply a perspective transform to rectify binary image ("birds-eye view").
def compute_perspective_transform_matrix(img):
width = img.shape[1]
height = img.shape[0]
xfactor1 = 0
xfactor2 = 0.447
xfactor3 = 0.3
yfactor = 0.627
# Source pointe hand-picked from lane image
src = np.float32([[xfactor2*width, yfactor*height],
[(1-xfactor2)*width, yfactor*height],
[(1-xfactor1)*width, height],
[xfactor1*width, height]])
# Destination pointe hand-picked from assumed bird's eye view
dst = np.float32([[xfactor3*width, 0],
[(1-xfactor3)*width, 0],
[(1-xfactor3)*width, height],
[xfactor3*width, height]])
# Compute the tranform matrix
return cv2.getPerspectiveTransform(src, dst)
for img_file in test_img_list:
img = mpimg.imread('test_images/' + img_file)
perspective_matrix = compute_perspective_transform_matrix(img)
break
# +
def create_birds_eye_view(img, perspective_matrix):
img_size = (img.shape[1], img.shape[0])
return cv2.warpPerspective(img, perspective_matrix, img_size)
if 1:
for img_file in test_img_list:
# img = mpimg.imread('test_images/' + img_file)
img = mpimg.imread('test_images/' + 'straight_lines1.jpg')
birds_eye_view = create_birds_eye_view(img, perspective_matrix)
fig = plt.figure()
plt.imshow(birds_eye_view)
plt.draw()
fig.savefig('output_images/birds_eye.png')
break
# +
# Bootstrap lane finding method when no prior polynomial fits exist
def find_lane_pixels(binary_warped):
# Take a histogram of the bottom half of the image
histogram = np.sum(binary_warped[binary_warped.shape[0]//2:,:], axis=0)
# Create an output image to draw on and visualize the result
out_img = np.dstack((binary_warped, binary_warped, binary_warped))
# Find the peak of the left and right halves of the histogram
# These will be the starting point for the left and right lines
midpoint = np.int(histogram.shape[0]//2)
leftx_base = np.argmax(histogram[:midpoint])
rightx_base = np.argmax(histogram[midpoint:]) + midpoint
# HYPERPARAMETERS
# Choose the number of sliding windows
nwindows = 9
# Set the width of the windows +/- margin
margin = 100
# Set minimum number of pixels found to recenter window
minpix = 50
# Set height of windows - based on nwindows above and image shape
window_height = np.int(binary_warped.shape[0]//nwindows)
# Identify the x and y positions of all nonzero pixels in the image
nonzero = binary_warped.nonzero()
nonzeroy = np.array(nonzero[0])
nonzerox = np.array(nonzero[1])
# Current positions to be updated later for each window in nwindows
leftx_current = leftx_base
rightx_current = rightx_base
# Create empty lists to receive left and right lane pixel indices
left_lane_inds = []
right_lane_inds = []
# Step through the windows one by one
for window in range(nwindows):
# Identify window boundaries in x and y (and right and left)
win_y_low = binary_warped.shape[0] - (window+1)*window_height
win_y_high = binary_warped.shape[0] - window*window_height
win_xleft_low = leftx_current - margin
win_xleft_high = leftx_current + margin
win_xright_low = rightx_current - margin
win_xright_high = rightx_current + margin
# # Draw the windows on the visualization image
# cv2.rectangle(out_img,(win_xleft_low,win_y_low),
# (win_xleft_high,win_y_high),(0,255,0), 2)
# cv2.rectangle(out_img,(win_xright_low,win_y_low),
# (win_xright_high,win_y_high),(0,255,0), 2)
# Identify the nonzero pixels in x and y within the window #
good_left_inds = ((nonzeroy >= win_y_low) & (nonzeroy < win_y_high) &
(nonzerox >= win_xleft_low) & (nonzerox < win_xleft_high)).nonzero()[0]
good_right_inds = ((nonzeroy >= win_y_low) & (nonzeroy < win_y_high) &
(nonzerox >= win_xright_low) & (nonzerox < win_xright_high)).nonzero()[0]
# Append these indices to the lists
left_lane_inds.append(good_left_inds)
right_lane_inds.append(good_right_inds)
# If you found > minpix pixels, recenter next window on their mean position
if len(good_left_inds) > minpix:
leftx_current = np.int(np.mean(nonzerox[good_left_inds]))
if len(good_right_inds) > minpix:
rightx_current = np.int(np.mean(nonzerox[good_right_inds]))
# Concatenate the arrays of indices (previously was a list of lists of pixels)
try:
left_lane_inds = np.concatenate(left_lane_inds)
right_lane_inds = np.concatenate(right_lane_inds)
except ValueError:
# Avoids an error if the above is not implemented fully
pass
# Extract left and right line pixel positions
leftx = nonzerox[left_lane_inds]
lefty = nonzeroy[left_lane_inds]
rightx = nonzerox[right_lane_inds]
righty = nonzeroy[right_lane_inds]
return leftx, lefty, rightx, righty, out_img
# Lane finding method when previous polynomial exists
def search_around_poly(binary_warped, left_fit, right_fit):
# HYPERPARAMETER
# Choose the width of the margin around the previous polynomial to search
# The quiz grader expects 100 here, but feel free to tune on your own!
margin = 100
# Grab activated pixels
nonzero = binary_warped.nonzero()
nonzeroy = np.array(nonzero[0])
nonzerox = np.array(nonzero[1])
### Set the area of search based on activated x-values ###
### within the +/- margin of our polynomial function ###
left_lane_inds = ((nonzerox > (left_fit[0]*(nonzeroy**2) + left_fit[1]*nonzeroy +
left_fit[2] - margin)) & (nonzerox < (left_fit[0]*(nonzeroy**2) +
left_fit[1]*nonzeroy + left_fit[2] + margin)))
right_lane_inds = ((nonzerox > (right_fit[0]*(nonzeroy**2) + right_fit[1]*nonzeroy +
right_fit[2] - margin)) & (nonzerox < (right_fit[0]*(nonzeroy**2) +
right_fit[1]*nonzeroy + right_fit[2] + margin)))
# Again, extract left and right line pixel positions
leftx = nonzerox[left_lane_inds]
lefty = nonzeroy[left_lane_inds]
rightx = nonzerox[right_lane_inds]
righty = nonzeroy[right_lane_inds]
out_img = np.dstack((binary_warped, binary_warped, binary_warped))
return leftx, lefty, rightx, righty, out_img
def fit_polynomial(binary_warped, ym_per_pix, xm_per_pix, left_fit_prev, right_fit_prev):
global i_plot
# Find our lane pixels first
if left_fit_prev == [] or right_fit_prev == []:
leftx, lefty, rightx, righty, out_img = find_lane_pixels(binary_warped)
else:
leftx, lefty, rightx, righty, out_img = search_around_poly(binary_warped, left_fit_prev, right_fit_prev)
# Fit a second order polynomial to each using `np.polyfit`
left_fit = np.polyfit(lefty, leftx, 2)
right_fit = np.polyfit(righty, rightx, 2)
left_fit_cr = np.polyfit(lefty*ym_per_pix, leftx*xm_per_pix, 2)
right_fit_cr = np.polyfit(righty*ym_per_pix, rightx*xm_per_pix, 2)
# Generate x and y values for plotting
ploty = np.linspace(0, binary_warped.shape[0]-1, binary_warped.shape[0] )
try:
left_fitx = left_fit[0]*ploty**2 + left_fit[1]*ploty + left_fit[2]
right_fitx = right_fit[0]*ploty**2 + right_fit[1]*ploty + right_fit[2]
except TypeError:
# Avoids an error if `left` and `right_fit` are still none or incorrect
print('The function failed to fit a line!')
left_fitx = 1*ploty**2 + 1*ploty
right_fitx = 1*ploty**2 + 1*ploty
if i_plot:
## Visualization ##
# Colors in the left and right lane regions
out_img[lefty, leftx] = [255, 0, 0]
out_img[righty, rightx] = [0, 0, 255]
fig = plt.figure()
plt.imshow(out_img)
# Plots the left and right polynomials on the lane lines
plt.plot(left_fitx, ploty, color='yellow')
plt.plot(right_fitx, ploty, color='yellow')
plt.draw()
fig.savefig('output_images/polyfit.png')
return left_fit_cr, right_fit_cr, left_fit, right_fit, left_fitx, right_fitx, ploty
# -
def measure_curvature_real(binary_warped, left_fit_cr, right_fit_cr, ym_per_pix, xm_per_pix):
'''
Calculates the curvature of polynomial functions in meters.
'''
# Define y-value where we want radius of curvature
# We'll choose the maximum y-value, corresponding to the bottom of the image
y_eval = binary_warped.shape[0]*ym_per_pix
# Calculation of R_curve (radius of curvature)
left_curverad = ((1 + (2*left_fit_cr[0]*y_eval + left_fit_cr[1])**2)**1.5) / np.absolute(2*left_fit_cr[0])
right_curverad = ((1 + (2*right_fit_cr[0]*y_eval + right_fit_cr[1])**2)**1.5) / np.absolute(2*right_fit_cr[0])
# Calculate lateral position
left_pos = left_fit_cr[0]*y_eval**2 + left_fit_cr[1]*y_eval + left_fit_cr[2]
right_pos = right_fit_cr[0]*y_eval**2 + right_fit_cr[1]*y_eval + right_fit_cr[2]
image_width_in_pixels = binary_warped.shape[1]
lateral_pos = 0.5 * (left_pos + right_pos) - 0.5*image_width_in_pixels*xm_per_pix
# Noise tends to reduce curvature so assume that lower curve radius estimate is more accurate
if (abs(left_curverad) < abs(right_curverad)):
curverad = left_curverad
else:
curverad = right_curverad
return curverad, lateral_pos
def low_pass_filter(x, x_prev, factor):
if x_prev == []:
x_filt = x
else:
x_filt = factor*x + (1-factor)*x_prev
return x_filt
def display_result(undistorted_image, birds_eye, left_fitx, right_fitx, ploty, perspective_matrix, curverad, lateral_pos):
# Create an image to draw the lines on
warp_zero = np.zeros_like(birds_eye).astype(np.uint8)
color_warp = np.dstack((warp_zero, warp_zero, warp_zero))
# Recast the x and y points into usable format for cv2.fillPoly()
pts_left = np.array([np.transpose(np.vstack([left_fitx, ploty]))])
pts_right = np.array([np.flipud(np.transpose(np.vstack([right_fitx, ploty])))])
pts = np.hstack((pts_left, pts_right))
# Draw the lane onto the warped blank image
cv2.fillPoly(color_warp, np.int_([pts]), (0,255, 0))
# Warp the blank back to original image space using inverse perspective matrix (Minv)
Minv = np.linalg.inv(perspective_matrix)
newwarp = cv2.warpPerspective(color_warp, Minv, (undistorted_image.shape[1], undistorted_image.shape[0]))
# Combine the result with the original image
result = cv2.addWeighted(undistorted_image, 1, newwarp, 0.3, 0)
font = cv2.FONT_HERSHEY_SIMPLEX
txt1 = "Radius of curvature = %0.1f m" % curverad
if lateral_pos > 0:
txt2 = "Vehicle is %0.3f m left of road center" % abs(lateral_pos)
else:
txt2 = "Vehicle is %0.3f m right of road center" % abs(lateral_pos)
cv2.putText(result, txt1, (50, 50), font, 1.2, (255, 255, 255), 2, cv2.LINE_AA)
cv2.putText(result, txt2, (50, 100), font, 1.2, (255, 255, 255),2 , cv2.LINE_AA)
return result
# +
left_fit_prev = []
right_fit_prev = []
curverad_prev = []
lateral_pos_prev = []
def process_image(img):
global mtx, dist, perspective_matrix, left_fit_prev, right_fit_prev, curverad_prev, lateral_pos_prev, i_plot
# Define conversions in x and y from pixels space to meters
ym_per_pix = 30/720 # meters per pixel in y dimension
xm_per_pix = 3.7/700 # meters per pixel in x dimension
# Apply distortion correction
undistorted_image = cv2.undistort(img, mtx, dist, None, mtx)
# Create binary image that is color coded to show detection method
color_binary_image = create_binary_image(undistorted_image)
# Convert to monochrome binary image
gray = cv2.cvtColor(color_binary_image, cv2.COLOR_RGB2GRAY)
binary_image = np.zeros_like(gray)
binary_image[gray > 5] = 1
# Create birds-eye view
birds_eye = create_birds_eye_view(binary_image, perspective_matrix)
# Compute lane boundaries
left_fit_cr, right_fit_cr, left_fit, right_fit, left_fitx, right_fitx, ploty = fit_polynomial(birds_eye, ym_per_pix, xm_per_pix, left_fit_prev, right_fit_prev)
# Record previous fit for faster find
left_fit_prev = left_fit
right_fit_prev = right_fit
# Compute lane curvature and vehicle position
curverad, lateral_pos = measure_curvature_real(birds_eye, left_fit_cr, right_fit_cr, ym_per_pix, xm_per_pix)
# Low-pass filter curvature and position
curverad_filt = low_pass_filter(curverad, curverad_prev, 0.05)
lateral_pos_filt = low_pass_filter(lateral_pos, lateral_pos_prev, 0.1)
curverad_prev = curverad_filt
lateral_pos_prev = lateral_pos_filt
# Display result
result = display_result(undistorted_image, birds_eye, left_fitx, right_fitx, ploty, perspective_matrix, curverad_filt, lateral_pos_filt)
if i_plot:
fig = plt.figure()
plt.imshow(undistorted_image)
plt.draw()
fig.savefig('output_images/undistort.png')
fig = plt.figure()
plt.imshow(binary_image)
plt.draw()
fig.savefig('output_images/binary_image.png')
plt.figure()
plt.imshow(birds_eye)
plt.draw()
fig = plt.figure()
plt.imshow(result)
plt.draw()
fig.savefig('output_images/result.png')
return result
if i_plot:
for img_file in test_img_list:
# img = mpimg.imread('test_images/' + img_file)
img = mpimg.imread('test_images/' + 'test2.jpg')
process_image(img)
break
# -
# Import everything needed to edit/save/watch video clips
from moviepy.editor import VideoFileClip
from IPython.display import HTML
project_output = 'project_video_output.mp4'
## To speed up the testing process you may want to try your pipeline on a shorter subclip of the video
## To do so add .subclip(start_second,end_second) to the end of the line below
## Where start_second and end_second are integer values representing the start and end of the subclip
## You may also uncomment the following line for a subclip of the first 5 seconds
# clip1 = VideoFileClip("project_video.mp4").subclip(0,5)
clip1 = VideoFileClip("project_video.mp4")
project_clip = clip1.fl_image(process_image) #NOTE: this function expects color images!!
# %time project_clip.write_videofile(project_output, audio=False)
HTML("""
<video width="960" height="540" controls>
<source src="{0}">
</video>
""".format(project_output))
challenge_output = 'challenge_video_output.mp4'
## To speed up the testing process you may want to try your pipeline on a shorter subclip of the video
## To do so add .subclip(start_second,end_second) to the end of the line below
## Where start_second and end_second are integer values representing the start and end of the subclip
## You may also uncomment the following line for a subclip of the first 5 seconds
#clip2 = VideoFileClip("challenge_video.mp4").subclip(0,5)
clip2 = VideoFileClip("challenge_video.mp4")
challenge_clip = clip2.fl_image(process_image) #NOTE: this function expects color images!!
# %time challenge_clip.write_videofile(challenge_output, audio=False)
HTML("""
<video width="960" height="540" controls>
<source src="{0}">
</video>
""".format(challenge_output))
|
AdvancedLaneFinding.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # carnd_t3_p1_path_planning_project
#
# [//]: # (Image References)
# [image1]: ./output/FSM.png
# [image2]: ./output/Right_turn.png
# [image3]: ./output/Left_turn_at_curve.png
# [image4]: ./output/Wall_of_cars.png
# [image7]: ./output/Vehicle_cut_in_front.png
# [image5]: ./output/Best_lap_46miles.png
# [image6]: ./output/Cost_function_design.png
# [video6]: ./output/IMG_4661.TRIM.MOV
#
#
# ## Carnd - term 3 - project 1 - path planning project
#
# ### Overview
# The goal of this project is to build a path planner model to drive vehicle around a 3-lane track and change lane as it sees opportunity to go ahead of other vehicles in front. The lane change action has to be performed safely without bumping into other vehicle as well as causing crash. The vehicle acceleration and turning has to be performed with minimal jerk to ensure comfort level for the passengers. The model processes a series of sensor data (x,y,s,d,yaw,speed,previous-path_x,previous_path_y,end_path_s,end_path_d,sensor_fusion,etc) to track the reference path and to predict the vehicle drive path, including lane changing. Spline library is used to optimize the waypoint fitting of the predicted reference path. Similar to other pervious projects in Term2, the path planning model is connected to a simulator via uWebSocketIO with waypoints of the reference path.
#
#
# ### Project Repository
# All resource are located in Udacity's project repository
# [CarND-Path-Planning-Project](https://github.com/udacity/CarND-Path-Planning-Project)
#
#
# ### Project Submission
# All modified code including results are committed to my personal github page
# [carnd_t3_p1_path_planning_project](https://github.com/chriskcheung/carnd_t3_p1_path_planning_project)
#
#
# ### Key Files
# ##### main.cpp
# establishes communication between simulator and path_planning model using uWebSocketIO, and reads in sensor data during a set time intervals. Data is passed into path planning class for initialization, processing sensor data, updating finite state machine (FSM) of path planner, and generating trajectory of the new predicted path.
#
# ##### path_planner.h and path_planner.cpp
# contains 1 main class: pathPlanning.
#
# pathPlanning class receives sensor data periodically. Base on the new sensor data, pathPlanning loops through all other vehicles data, for their locations and speed, then calculates the latest closest distance between other vehicles with respect to the subjected car, the cost functions of each lane with respect to their closest vehicle in both front and back.
#
# There are 5 major functions involved: initPathPlanning(), processSensorData(), isChangeLaneSafe(), updateFSM(), generateTrajectory().
#
# initPathPlanning() initializes all costs and predicted waypoints back to their default state or zero at every new data receives to start fresh.
#
# processSensorData() loops through all other vehicles data from sensor\_fusion to find the closest vehicle to the subjected car in each lane, considering both front and back. This maps out the landscape of the surround and ignores those that are outside of laneVisibleDist\_ to avoid unnecessary tracking. Base on their s distance to the subjected car, and the velocity difference, cost\_[] of changing lane into each lane is established for FSM to make decision.
#
# isChangeLaneSafe() determines if it is safe to proceed from the existing lane to the targeting lane base on other vehicles position. The min\_safe\_dist\_front\_ and min\_safe\_dist\_back\_ ensure that there is enough room to the front and back vehicle on the left and the right before signaling safe to proceed.
#
# updateFSM() tracks its current state of the subjected car. With the lowest cost\_[\*] and isChangeLaneSafe(), it determine which direction the car should turn, (target\_lane\_), vs staying in the current lane. The FSM also controls the speed of the car to either catch up or slow down to match the speed of vehicle in front to avoid collision, or the max legal speed. maxTravelSpeed\_.
#
# generateTrajectory() reuses up to 10 previous\_path\_x and previous\_path\_y waypoints to ensure smooth lane change transition. New way points are added from the 11th previous\_path\_\* alone with new points from the spline fitting line to ensure a smooth fitting line for lane change.
#
#
# ## Project Setup
# I create an AWS instance to execute pathPlanning executible and run term3_sim simulator in local PC. Data is sent through AWS server to local PC client, which create latency. The latency is obvious during implementation when switching between wired and WiFi connections. I end up tuning my model with wired connect to keep things consistent.
#
#
# ## Implementation Challenge
#
# ### Cost function
# Determine the right cost function is important to for the FSM to make decision. I start dividing the distance between subjected car and the closest vechicle in front of each lane (closest\_front\_s\_diff\_[\*]) by the delta of velocity between two cars (ref\_v\_ - closest\_front\_v\_[\*]) to determine the time the two vehicle apart. The larger the time is the bigger the buffer and the less cost it should be. With exponential function on the result of the division, it doesn't reveal such relationships. Reciprocal of exponential results seems to provides that relationship, but the spread is too heavy on one side rather than evenly distributed. After a few try, I settle with 100-exp(x) where x is (closest\_front\_s\_diff\_[\*])/(ref\_v\_ - closest\_front\_v\_[\*]). It provides the gradual increase of weight as x decrease which fits the bill of what I am looking for.
# ![alt text][image6]
#
# I filter out any vehicles that are fall into the following conditions to simply the cost calculation:
# 1. for vehicle outside of the visible distance, cost will be 0 to represent an open lane for advancing
# 2. for vehicle that is 2 lanes away from current lane, cost set cost to 100 as it is not impossible to jump 2 lanes without crossing the adjacent lane
#
# ```c++
# // use the end of previous path info to determine the cost of changing lane
# switch(int(ref_lane_)){
# // left most lane
# case 0:
# // cost of staying in same lane base on the distance of vehicle in the front
# cost_[0] += closest_front_s_diff_[0] > laneVisibleDist_ ? 0 :
# (ref_v_ - closest_front_v_[0])==0 ? 100 :
# max(0.0, min(100.0 - exp(closest_front_s_diff_[0]/(ref_v_ - closest_front_v_[0])), 100.0));
# // cost of turn right base on any room on the right front
# cost_[1] += closest_front_s_diff_[1] > laneVisibleDist_ ? 0 :
# (ref_v_ - closest_front_v_[1])==0 ? 100 :
# max(0.0, min(100.0 - exp(closest_front_s_diff_[1]/(ref_v_ - closest_front_v_[1])), 100.0));
# // not supporting to jump 2 lanes, so max cost for right most lane
# cost_[2] += 100;
# break;
# ...
# }
# }
#
# ```
#
# ### isChangeLaneSafe()
# Checking whether it is safe to change lane depends on the subject vehicle position with respect the other vehicle in the front or in the back on current lane and the adjacent lane that is targeting for lane change. We have to look ahead of both vehicle in future (reference point from the previous 10th waypoint) to make sure they won't collide to each other when lane change actually happens. closest\_front\_s\_diff\_ subtracts previous\_path\_x[10] with future distance of other vehicle in adjacent lane at time of previous\_path\_x[10]. Same goes to vehicle from the back for each lane.
#
# ```c++
# // cap the previous point to be reused at the 10th point for smoothing
# prv_size_ = min(int(previous_path_x.size()), 10);
# ...
# double prev_x1 = prv_x_[prv_size_-1];
# ...
# ref_x_ = prev_x1;
# ...
#
# double time_2_prev_path_end = dt_ * prv_size_;
# // vehicle velocity
# double vehicle_v = sqrt(vehicle_vx*vehicle_vx + vehicle_vy*vehicle_vy);
# // vehicle s distance with respect to end of subjected car's previous path trajectory
# double vehicle_s_at_prev_path_end = vehicle_s + vehicle_v*time_2_prev_path_end;
# // s diff between subjected car and vehicle from sensor fusion
# double s_diff = fabs(ref_s_ - vehicle_s_at_prev_path_end);
# ...
# closest_front_s_diff_[vehicle_lane] = s_diff;
# ...
# if (closest_front_s_diff_[1] > min_safe_dist_front_ && closest_back_s_diff_[1] > min_safe_dist_back_)
# return true;
# else
# return false;
# ```
#
#
# ### FSM
# This implementation is based on a 3 state finite state machine: keepLane, laneChangeLeft, and laneChangeRight states.
# ![alt text][image1]
#
# It starts at keepLane state. The state transition is based on 3 conditions:
# 1) if gap between the front vehicle is at or shorter than the minimum distance to consider lane change,
# 2) if cost of changing lane lower than cost of staying in current lane,
# 3) if it is safe to change lane to the targeting lane.
# keepLane state determines the target\_lane\_, and then pass it to the next state. laneChangeLeft and laneChangeRight both monitors the current lane from sensor data. Once the current lane matches with target\_lane\_, which signals that the lane change is completed successfully, it will transition back to keepLane state. Each state is responsible to maintain the subject vehicle speed to accelerate or decelerate its speed to match the front vehicle speed of the target\_lane\_ if presence, or catch up to legal speed limit. Speed should not exceed legal limit at any given time, thus speed will be decelerate when it gets too close to limit.
#
# ```c++
# case fsmStateType::keepLane:
# // consider change lane if gap between car in front is closer than minium lane change distance
# if(closest_front_s_diff_[ref_lane_] < min_lane_change_dist_){
#
# // reduce speed first to match the speed of car in front
# ref_v_ = max(ref_v_-0.6, closest_front_v_[ref_lane_]-0.6);
#
# // skip changing lane if it just finished changing lane within 10th of sec
# if(timer_ == 0){
# //cost of changing lane
# if(ref_lane_==0 && isChangeLaneSafe(0, 1) && (cost_[0] > cost_[1])){
# f_ = fsmStateType::laneChangeRight;
# target_lane_ = 1;
# break;
# }
# else if(ref_lane_==2 && isChangeLaneSafe(2, -1) && (cost_[2] > cost_[1])){
# f_ = fsmStateType::laneChangeLeft;
# target_lane_ = 1;
# break;
# }
# else if(ref_lane_==1){
# // change to left if cost of left lane is the lowest and if it is feasible
# if(cost_[0] <= cost_[2] && cost_[0] < cost_[1] && isChangeLaneSafe(1, -1)){
# // change lane left
# f_ = fsmStateType::laneChangeLeft;
# target_lane_ = 0;
# break;
# }
# // change to right if cost of right lane is the lowest and if it is feasible
# else if(cost_[2] <= cost_[0] && cost_[2] < cost_[1] && isChangeLaneSafe(1, 1)){
# // change lane right
# f_ = fsmStateType::laneChangeRight;
# target_lane_ = 2;
# cout << "SSS: change lane from 1 to RIGHT to " << target_lane_<< endl;
# break;
# }
# // else stay in its lane if cost of its lane is the lowest
# }
# }
# }
# else if(ref_v_ < maxTravelSpeed_*MILE_PER_HOUR_2_METER_PER_SEC)
# ref_v_ = min(ref_v_+0.8, maxTravelSpeed_*MILE_PER_HOUR_2_METER_PER_SEC);
# else if (ref_v_ > maxTravelSpeed_*MILE_PER_HOUR_2_METER_PER_SEC)
# ref_v_ = max(ref_v_-0.6, maxTravelSpeed_*MILE_PER_HOUR_2_METER_PER_SEC);
#
# if(timer_ < 0)
# timer_ = 0;
# else if(timer_ > 0)
# timer_ -= dt_*(nextPathSize_-prv_size_);
# target_lane_ = ref_lane_;
# break;
#
# case fsmStateType::laneChangeLeft:
# // lane change completed
# if(car_lane_ == target_lane_){
# f_ = fsmStateType::keepLane;
# // start timer to settle in to new lane before another lane change
# timer_ = 2;
# }
#
# // set targeting speed to either match the speed limit or the vehicle in front of target lane
# if(closest_front_v_[target_lane_] > 0 && closest_front_v_[target_lane_] < target_v)
# target_v = closest_front_v_[target_lane_];
#
# // adjust speed
# if(ref_v_ < target_v)
# // purposely lower acceleration to +0.6 from +0.8 to reduce jerk when changing lane at the curve
# ref_v_ = min(ref_v_+0.6, target_v);
# else
# // purposely increase decceleration to -0.6 from -0.8 to reduce jerk when changing lane at the curve
# ref_v_ = max(ref_v_-0.8, target_v);
# break;
#
# case fsmStateType::laneChangeRight:
# // complete lane change
# if(car_lane_ == target_lane_){
# f_ = fsmStateType::keepLane;
# // start timer to settle in to new lane before another lane change
# timer_ = 2;
# }
# // set targeting speed to either match the speed limit or the vehicle in front of target lane
# if(closest_front_v_[target_lane_] > 0 && closest_front_v_[target_lane_] < target_v)
# target_v = closest_front_v_[target_lane_];
# if(ref_v_ < target_v)
# // purposely lower acceleration to +0.6 from +0.8 to reduce jerk when changing lane at the curve
# ref_v_ = min(ref_v_+0.6, target_v);
# else
# // purposely increase decceleration to -0.6 from -0.8 to reduce jerk when changing lane at the curve
# ref_v_ = max(ref_v_-0.8, target_v);
# break;
# ```
#
# #### State Machine Improvement
# Notice that both laneChangeLeft and laneChangeRight states are identical. Since the target\_lane\_ is determined in keepLane and stays the same throughout laneChangeLeft and laneChangeRight, this FSM works just fine with 2 state implementation: keepLane and laneChange states. This will be for future improvement.
#
# #### Jerk prevention
# During lane change at straight lanes, jerk will not be an issue. However, when changing lane at the curve, with the extra turning on top of the curvature of the lane, it may apply additional force to create jerk. Simple tricks can be done to prevent this. It is by purpose to lower acceleration from 0.8 to 0.6 and increase deceleration from 0.6 to 0.8 to reduce speed as well as jerk create during lane change.
#
# ```c++
# if(ref_v_ < target_v)
# // purposely lower acceleration to +0.6 from +0.8 to reduce jerk when changing lane at the curve
# ref_v_ = min(ref_v_+0.6, target_v);
# else
# // purposely increase decceleration to -0.6 from -0.8 to reduce jerk when changing lane at the curve
# ref_v_ = max(ref_v_-0.8, target_v);
# ```
#
# ### Trajectory
# Trajectory generation consists of two parts: reuse of preivous waypoints as the base points, and newly generated points from the last of the previous.
#
# The reuse of previous waypoints is for the purpose of continuation from previous prediction to avoid a suddent change in direction and avoid jerk.
# ```c++
# // Since the spline fitting line starts from the last 2 points from the previous path points
# // as a continuation and the rest of the points are newly plotted/picked to predict the new path
# // for the next 30 meters. With the spline fitting line ready, break the spline evenly to N points
# // with the distance and time interval base on the the speed that we want to travel
# // start with all of previous path points from last time
# for(int i=0; i<prv_size_; i++) {
# next_x.push_back(previous_path_x[i]);
# next_y.push_back(previous_path_y[i]);
# }
# ```
# The new waypoint generation is based on the last two points from the reuse waypoints, plus three more points that are 30 meters apart to create a nice even sketch of the line. These five points are fitted into Spline equation to fit a line with nice and smooth curvature for our prediction.
#
# ##### Note
# Previously, I use 30 meters as suggested by <NAME> in his walkthrough video. I find that not enough to create a smooth fitting line that would avoid jerk. I could have blamed it to my vehicle speed being too fast. However, increasing the distance fo these three waypoints from 30 meters to 50 meters apart makes the spline fitting line even my smooth and the subject vehicle changes lane as smooth as lane splitting. Therefore, I leave it at 50 meters as my final submission version.
#
#
# ```c++
# // in Frenet add evenly 30m spaced points ahead of the starting reference so the points are not only
# // cover the even distance points. original 30m apart, curently increase to 50m apart to further
# // reduce jerk during lane change by sketching out the spline fit line
# vector<double> next_wp0 = getXY(car_s_+50, (2+laneWidth_*target_lane_), map_s_, map_x_, map_y_);
# vector<double> next_wp1 = getXY(car_s_+100, (2+laneWidth_*target_lane_), map_s_, map_x_, map_y_);
# vector<double> next_wp2 = getXY(car_s_+150, (2+laneWidth_*target_lane_), map_s_, map_x_, map_y_);
# ...
# ptsx.push_back(prv_x2);
# ptsy.push_back(prv_y2);
# ...
# // push the ref_x_ and ref_y_ as 2nd point, see processSensorData() for their origin
# ptsx.push_back(ref_x_);
# ptsy.push_back(ref_y_);
#
# ptsx.push_back(next_wp0[0]);
# ptsx.push_back(next_wp1[0]);
# ptsx.push_back(next_wp2[0]);
#
# ptsy.push_back(next_wp0[1]);
# ptsy.push_back(next_wp1[1]);
# ptsy.push_back(next_wp2[1]);
# ```
#
# These coordinates are based in global coordinates. It is easier to convert them to car's coordinates (or Frenet coordinates) before fitting happens.
#
# ```c++
# for(int i=0; i<ptsx.size(); i++){
# //shift car reference angle to 0 degrees to align with local coordinates or in car's prespective
# double shift_x = ptsx[i]-ref_x_;
# double shift_y = ptsy[i]-ref_y_;
#
# ptsx[i] = (shift_x*cos(0-ref_yaw_)) - (shift_y*sin(0-ref_yaw_));
# ptsy[i] = (shift_x*sin(0-ref_yaw_)) + (shift_y*cos(0-ref_yaw_));
# }
#
# // create a spline for fitting
# tk::spline s;
#
# // set (x,y) points to spline
# s.set_points(ptsx, ptsy);
# ```
#
# Once the spline fitting line is done, spread the points evenly over a distance of 30 meters, using the target speed, ref\_v\_, that we obtain from FSM. It is important to use ref\_v\_ as it how fast the subject vehicle should go to avoid colliding into the vehicle in front. Also, each waypoint has to be converted back to global coordinates before feeding back to simulator to drive the subject vehicle.
#
# ```c++
# // fill up the rest of out path planner after filling it with previous points, here we will always output 50 points
# double N = (target_dist/(0.02*ref_v_)); // ref_v base on meter/sec
#
# for(int i=1; i<=nextPathSize_-prv_size_; i++){
# double x_point = x_add_on+(target_x/N);
# double y_point = s(x_point);
#
# x_add_on = x_point;
#
# double x_ref = x_point;
# double y_ref = y_point;
#
# // rotate back to normal after rotating it eariler (back to global coordinates)
# x_point = (x_ref*cos(ref_yaw_)) - (y_ref*sin(ref_yaw_));
# y_point = (x_ref*sin(ref_yaw_)) + (y_ref*cos(ref_yaw_));
#
# x_point += ref_x_;
# y_point += ref_y_;
#
# next_x.push_back(x_point);
# next_y.push_back(y_point);
# }
#
# ```
#
# ### Results
# Unlike all previous project, this project is quite challenging as it is not given a lot of hints or guidance to begin with. However, all materials are covered by lecture one way or the other. It is up to us the student to put them together. There are many ways to approach the problem set and I found this very interesting when working out all the piece to finalize my solutions with 3 state FSM or even possibly to improve to a 2 state FSM. The result is quite satisfactory as one of my best lap reaches 46 miles without instance. The challenging part is to tuning the safety distance, speeds, and even the newly generated waypoints to avoid jerk when changing lanes. Below, I capture a few of the screenshot to illustrate the lane change moments and the best lap result.
# [image3]: ./output/Left_turn_at_curve.png
# [image4]: ./output/Wall_of_cars.png
# [image7]: ./output/Vehicle_cut_in_front.png
# [image5]: ./output/Best_lap_46miles.png
# ![alt text][image2]
# Making right turn
# ![alt text][image3]
# Marking left turn at the curve
# ![alt text][image4]
# Two cars in parallel in front, car turns to middle lane, then the right most lane
# ![alt text][image7]
# Black vehicle cut in front, car makes left turn right away
# ![alt text][image5]
# Best lap of 46 miles with incidence
|
README.md.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.10 64-bit (''fcc_tf'': conda)'
# name: python3
# ---
# +
from __future__ import absolute_import, division, print_function, unicode_literals
from path import Path
import pandas as pd
from IPython.display import clear_output
import tensorflow_probability as tfp
import tensorflow as tf
tfd = tfp.distributions
# -
initial_distribution = tfd.Categorical(probs=[0.8, 0.2])
transition_distribution = tfd.Categorical(probs=[[0.7, 0.3],
[0.2, 0.8]])
observation_distribution = tfd.Normal(loc=[0., 15.], scale=[5., 10.])
model = tfd.HiddenMarkovModel(
initial_distribution=initial_distribution,
transition_distribution=transition_distribution,
observation_distribution=observation_distribution,
num_steps=7
)
# +
mean = model.mean()
with tf.compat.v1.Session() as sess:
print(mean.numpy())
|
tensorflow/Hidden Markov Model.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/yoraoui/PingPong/blob/main/point_cloud.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="3LWqyH8NU4Rp" outputId="c4cf0ce7-b739-4d8b-9c7d-1783486b778f"
# ! python -V
# ! pip3 install trimesh
# ! pip3 install pyglet
# + id="YJzmxRkvP5ea"
import pdb
import os
import glob
import trimesh
import trimesh.sample
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
import random as random
# + id="FMh2XfozY16v"
# If using a GPU keep these lines to avoid CUDNN errors
gpus = tf.config.experimental.list_physical_devices('GPU')
for gpu in gpus:
tf.config.experimental.set_memory_growth(gpu, True)
# + colab={"base_uri": "https://localhost:8080/"} id="c8g0BARGaMxs" outputId="76cbd8ca-50af-4d3f-8263-dd3c85e665cd"
# Testing Stuff in this cell, Not Relevant for the problem
from typing import NewType
PointCloud = NewType("PointCloud", np.ndarray)
print(PointCloud)
def get_point_cloud():
return np.random.randint(1, 10, size = (1024, 3))
pcs = []
for i in range(10):
pcs.append(get_point_cloud())
for pc in pcs:
# print(pc.shape)
# print(PointCloud(pc))
pass
# + [markdown] id="xRYB748BM-va"
# ## Getting the data to local system
# + id="WKvZGQ97KtK_" colab={"base_uri": "https://localhost:8080/"} outputId="19aeb2db-da03-4ffa-99e8-36d6d5c6b166"
import os
if not os.path.exists("ModelNet10.zip"):
# ! curl http://3dvision.princeton.edu/projects/2014/3DShapeNets/ModelNet10.zip -L -o ModelNet10.zip
# + id="Qhm1UutaMbaz"
import os
if not os.path.exists("data/ModelNet10"):
os.mkdir("data")
# !unzip ModelNet10.zip -d ./data/
# + colab={"base_uri": "https://localhost:8080/"} id="B1XjOZ7AQPJV" outputId="69560324-0510-4b70-c4a2-19f1ae42bbec"
# !ls data/ModelNet10
# + id="ma8jqZ-eNwhv" colab={"base_uri": "https://localhost:8080/"} outputId="25ee36fb-5d2a-4a2c-c723-aabc1d049f18"
data_folder = os.path.join(os.curdir, "data", "ModelNet10")
with open(os.path.join(data_folder, "README.txt")) as f:
print(f.read())
f.close()
# + [markdown] id="EH0EiSzxQXPL"
# ## Reading and Visualising the data
# + id="Nu3J3yHUQU1i"
# ! export data_folder="./data/ModelNet10"
# + id="9w0pe0iEUl6J" colab={"base_uri": "https://localhost:8080/"} outputId="76980128-a493-4437-f8c8-98d3dd67d6a6"
DATA_DIR = data_folder
print(DATA_DIR)
# + colab={"base_uri": "https://localhost:8080/"} id="SQwb8-fTUB7_" outputId="a4cba183-6f74-4f76-830b-d6153d8be401"
# !ls $data_folder/bathtub/test
# + id="5WtnVlkn548I"
num_points_per_cloud = 1024 # <- you can modify this number as needed
# + id="QI9N00PJUv5f"
cad_mesh = trimesh.load(os.path.join(DATA_DIR, "bed/train/bed_0010.off")) # <- Set path to a .off file
#cad_mesh.show()
# + [markdown] id="DlytQV9jVynz"
# ## Utils functions
# + colab={"base_uri": "https://localhost:8080/"} id="parHwyOMkwte" outputId="e8ca2c28-8688-420c-f8c8-364eb06be416"
# Test Code
class_ids = {}
folders = glob.glob(os.path.join(DATA_DIR, "[!README]*"))
for class_id, folder in enumerate(folders):
print(os.path.basename(folder))
class_ids[class_id] = os.path.basename(folder)
sub_folder = "train"
for f in folders:
# train_folder = os.path.join(f, "train")
# print(train_folder)
# # print(os.listdir(train_folder))
print(glob.glob(os.path.join(f, f"{sub_folder}/*.off")))
print(class_ids)
# + id="6YExEW8g04Fs"
# function for getting point_cloud by passing cad mesh filepath
def mesh_file_2_pc(file_path, n_points):
"function takes path to a cad file and returns point cloud (generated with n_points)"
cad_mesh = trimesh.load(file_path)
point_cloud_array = trimesh.sample.sample_surface(cad_mesh, n_points)[0]
return point_cloud_array
# + colab={"base_uri": "https://localhost:8080/"} id="9ByVcZcTihu2" outputId="761d0fd6-471f-40fe-f11d-9fda5ad0b4f0"
def test_dry_run():
" to test extraction of all point clouds to list"
def process_folders(folders, subFolder_type = "train"):
"collecting all the point clouds in a folder into array"
pcs = [] # array to hold all point clouds
for f in folders:
mesh_files = glob.glob(os.path.join(f, f"{subFolder_type}/*.off"))
print(f"Getting point cloud for: \n {mesh_files}")
for file in mesh_files[:2]:
# reading 2 files from each to keep the run short
point_cloud = mesh_file_2_pc(file, 1024)
pcs.append(point_cloud)
return pcs
folders = glob.glob(os.path.join(DATA_DIR, "[!README]*"))
pcs = process_folders(folders)
for pc in pcs:
print(pc.shape)
return pcs
pcs = test_dry_run()
# + id="MIzOgbPj26vi"
assert np.array(pcs).shape == (20, 1024, 3)
# 20 point cloud files with each of shape (1024, 3)
# + id="B5kZqYRNVxqv"
def create_point_cloud_dataset(data_dir, num_points_per_cloud=1024):
"""
Given the path to the ModelNet10 dataset, samples the models and creates point clouds
:param data_dir: path to the ModelNet10 dataset
:type data_dir: str
:param num_points_per_cloud: number of points to sample per cloud. 1024, 2048....
:type num_points_per_cloud: int
:return: tuple of numpy array containing training and test point clouds, their corresponding labels and a list of
class IDs
:rtype: tuple
"""
train_pc = [] # array of training point clouds
test_pc = [] # array of test point clouds
train_labels = [] # array of corresponding training labels
test_labels = [] # array of corresponding test labels
class_ids = {} # list of class names
# get all the folders except the readme file
folders = glob.glob(os.path.join(data_dir, "[!README]*"))
print(folders)
for class_id, folder in enumerate(folders):
print("processing class: {}".format(os.path.basename(folder)))
# Custom code here, get the name of the folder (class) and save it
##############################
class_ids[class_id] = os.path.basename(folder)
##############################
# get the files in the train folder
train_files = glob.glob(os.path.join(folder, "train/*"))
for f in train_files:
# Custom Code Here
##################################
pc = mesh_file_2_pc(f, num_points_per_cloud)
train_pc.append(pc)
train_labels.append(class_id)
###############################3
# get the files in the test folder
test_files = glob.glob(os.path.join(folder, "test/*"))
for f in test_files:
# Custom Code Here
##########################
pc = mesh_file_2_pc(f, num_points_per_cloud)
test_pc.append(pc)
test_labels.append(class_id)
########################
return (np.array(train_pc), np.array(test_pc),
np.array(train_labels), np.array(test_labels), class_ids)
# + id="xm9ioj_i0Gua"
def visualize_cloud(point_cloud):
"""
Utility function to visualize a point cloud
:param point_cloud: input point cloud
:type point_cloud: numpy array
"""
fig = plt.figure()
ax = fig.add_subplot(111, projection="3d")
ax.scatter(point_cloud[:, 0], point_cloud[:, 1], point_cloud[:, 2])
plt.show()
def add_noise_and_shuffle(point_cloud, label):
"""
Adds noise to a point cloud and shuffles it
:param point_cloud: input point cloud
:type point_cloud: tensor
:param label: corresponding label
:type label: tensor
:return: the processed point cloud and the label
:rtype: tensors
"""
dev_in_metres = 0.002 # <- change this value to change amount of noise
# add noise to the points
point_cloud += tf.random.uniform(point_cloud.shape, -dev_in_metres, dev_in_metres, dtype=tf.float64)
# shuffle points
point_cloud = tf.random.shuffle(point_cloud)
return point_cloud, label
# + [markdown] id="vXCGvJQhVjNN"
# ### Generating the point cloud for a geometry
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="yI7Ulo3LU3ed" outputId="9f59149b-0b74-451b-ce2b-e34e551d60b8"
points = trimesh.sample.sample_surface(cad_mesh, 1024)[0]
# visualize the point cloud using matplotlib
visualize_cloud(points)
print(points.dtype, points.shape)
# + colab={"base_uri": "https://localhost:8080/", "height": 248} id="bUKde8-R32kB" outputId="ac6fb908-7691-428a-c413-d3d4dfbd346a"
def object_pc(category: str):
n = np.random.randint(1, 10)
cad_mesh = trimesh.load(os.path.join(DATA_DIR, f"{category}/train/{category}_000{n}.off"))
points = trimesh.sample.sample_surface(cad_mesh, 1024)[0]
# visualize the point cloud using matplotlib
visualize_cloud(points)
object_pc("chair")
# + [markdown] id="4HBAk2QwWgJp"
# ### Generating point clouds from complete dataset
# + id="INAjEP7EWOgZ" colab={"base_uri": "https://localhost:8080/"} outputId="d8bcabfd-0c85-41dd-e779-6129052e7b32"
num_points_per_cloud = 1024 # <- you can modify this number as needed
train_pc, test_pc, train_labels, test_labels, class_ids = create_point_cloud_dataset(DATA_DIR, num_points_per_cloud)
# + [markdown] id="06fbbEDLXLo6"
# ### Saving Point Cloud in pickle files
# + id="shUcTEweXDGw"
import pickle
# once loaded save the numpy arrays to pickle files to use later
pickle.dump(train_pc, open("trainpc.pkl", "wb"))
pickle.dump(test_pc, open("testpc.pkl", "wb"))
pickle.dump(train_labels, open("trainlabels.pkl", "wb"))
pickle.dump(test_labels, open("testlabels.pkl", "wb"))
pickle.dump(class_ids, open("class_ids.pkl", "wb"))
# + id="hz3iApxNgVgc" colab={"base_uri": "https://localhost:8080/"} outputId="f857c814-f17f-47b5-9274-68fc03ba823c"
# ls
# + id="4RBlxfJc8wK9"
# from google.colab import files
def download_files():
"download files to avoid preprocessing again"
for file in ["trainpc.pkl", "trainlabels.pkl", "testpc.pkl", "testlabels.pkl", "class_ids.pkl"]:
files.download(file)
# download_files()
# + id="PMxNBMml-B6s"
# RUN THIS ONLY IF UPLOADING FILE FROM LOCAL SYSTEM
from google.colab import files
def upload_files():
uploaded = files.upload()
for fn in uploaded.keys():
print('User uploaded file "{name}" with length {length} bytes'.format(
name=fn, length=len(uploaded[fn])))
# upload_files()
# + [markdown] id="oI4r_BRoYPCx"
# # CAN START HERE IF POINT CLOUD IS ALREADY PROCESSED AND SAVED AS PICKLE FILES
# + [markdown] id="G4Sxyd8tYYtl"
# ### Getting the already saved point cloud arrays and labels
# + colab={"base_uri": "https://localhost:8080/"} id="ga0DBlz3bSi0" outputId="6b1d63e7-49c9-4fd8-a1b1-277e697506e4"
# ls
# + id="jue-0DcyYM21"
# load the data from pickle files if already present
import pickle
def load_from_files():
train_pc = pickle.load(open("trainpc.pkl", "rb"))
train_labels = pickle.load(open("trainlabels.pkl", "rb"))
test_pc = pickle.load(open("testpc.pkl", "rb"))
test_labels = pickle.load(open("testlabels.pkl", "rb"))
class_ids = pickle.load(open("class_ids.pkl", "rb"))
# load_from_files()
# + id="wpR9Zp0h7sY7" colab={"base_uri": "https://localhost:8080/"} outputId="e461a22d-723c-49bb-d3c8-898d719f99b3"
# Glipse of the data that was generated in preprocessing
print(train_pc.shape)
print(train_pc[0])
print(train_labels.shape)
print(train_labels[0:3900: 100])
print(test_pc.shape)
print(test_labels.shape)
print(test_labels[0:900: 100])
print(class_ids)
# + [markdown] id="YLLYtBb-W5MQ"
# ### Generating scene with multiple objects for inputs to segmentation model
# + id="NYGVnK-qWsB4"
# import pdb
import numpy as np
import pdb
import random
import pandas as pd
from tqdm import tqdm
def as_list(x):
if type(x) is list:
return x
else:
return [x]
def one_hot(scene_labels):
# pdb.set_trace()
" return [0, 1, 0, 1, 0, 0, 0, 0, 1, 0] for scene_labels = [8, 1, 3]"
one_hot_array = np.zeros(10)
for i in scene_labels:
one_hot_array[i] = 1
return one_hot_array
def get_tiled(label_onehot):
tiled_labels = np.tile(label_onehot, (1024, 1))
return tiled_labels
def pc_transform(pc):
"takes a point cloud and transform"
# [4, 4, 4] * [2.5, 2.5, 2.5].T = [10, 10, 10]
# scaling and objects by say 2.6 times
scaled_pcs = []
factor = np.random.randint(1, 2) + np.random.randn()
# factor = 1.
scaling_factor = [factor, factor, factor]
# print(f"scaling factor: {scaling_factor}")
scaling_diag_array = np.diag(scaling_factor)
scaled_pc = np.dot(pc, scaling_diag_array.T)
# print(f"scaled point cloud shape: {scaled_pc.shape}")
a = [[1, 3, 5], [2, 4, 6]]
sf = np.diag([2, 2, 2])
assert (np.dot(a, sf.T) == [[2, 6, 10], [4, 8, 12]]).all()
assert scaled_pc.shape == (1024, 3), "scales_pcs dims mismatch issue"
# for row in point_cloud_array:
# scaled_row = np.dot(row, scaling_diag_array.T) # do np.dot(A, B)
# scaled_pcs.append(scaled_row)
# translation
translated_length = np.random.randint(-40, 40, size= (1, 3))
translation_mat = np.tile(translated_length, (1024, 1))
# print(f"translated_length : {translated_length }")
translated_pc = np.add(scaled_pc, translation_mat)
assert translated_pc.shape == (1024, 3), "translated_pcs dims mismatch"
# for row in scaled_pcs:
# print(f"point cloud row for")
# new_point_loc = row + translated_length
# print(f"new point loc: {new_point_loc}")
# translated_pcs.append(new_point_loc)
return translated_pc
def get_random_index(grouped_idx_df):
random.seed(123)
df1 = grouped_idx_df.apply(np.random.choice)
random_indexes = df1.sample(n=3, replace = False).tolist()
return random_indexes
def generate_scenes(pc_array, label_array, n_scenes, n_objects):
"this generates the scene from the point cloud given"
scene_pc_array = []
scene_label_array = []
# n_records_in_pc = len(pc_array)
# print("n_records_in_pc", n_records_in_pc)
# l = list(range(n_records_in_pc))
# random.seed(123)
# random.shuffle(l)
# generating groups of indexes with same label
df = pd.DataFrame(label_array, columns = ["labels"])
df["index"] = df.index
grouped_idx_df = df.groupby("labels")["index"].apply(list)
for i in tqdm(range(n_scenes)):
"this look will run to create a scene"
index_values = get_random_index(grouped_idx_df)
# print("index_values", index_values)
scene_pc = np.empty((0, 3))
pointwise_labels = np.empty((0, 10))
for j, idx in enumerate(index_values):
"this loop will run to add objects to a single scene"
# print("idx", idx)
# print(pc_array[idx], "pc_array[idx]")
#scene_pc.append(pc_array[idx]) # before adding an object in scece, transform it (scale it and translate)
scene_pc = np.append(scene_pc, pc_transform(pc_array[idx]), axis = 0)
pointwise_labels = np.append(pointwise_labels, get_tiled(one_hot(as_list(label_array[idx]))), axis= 0)
# print(f"scene labels for scene {i}: {scene_label}")
scene_pc_array.append(np.array(scene_pc))
scene_label_array.append(np.array(pointwise_labels))
return np.array(scene_pc_array), np.array(scene_label_array)
# CTRL + M + L
def segmentation_pc_gen(train_pc, train_labels, test_pc, test_labels):
"""
Generates point cloud from segmentation problem
Input: list of point clouds [total_objects, 1024, 3]
Output: list of point clouds where each point cloud is [2048, 3] => anything which has more thatn 2 objects will need to resample so the size is amaintained
2 objects in a scene => (2048, 3)
4 objects in a scene => (4096, 3) => we need to resample from this so that overall point cloud size of (2048, 3) is maintained
"""
N_TRAIN = 2000
N_TEST = 600
N_OBJECTS_IN_SCENE = 3
# the size of each point cloud is [3072, 3]
# Overall 2000 point clouds generated for training and 600 point clouds generated for test
scene_train_pc, scene_train_labels = generate_scenes(train_pc, train_labels, N_TRAIN, n_objects = N_OBJECTS_IN_SCENE)
# print(scene_train_pc)
scene_test_pc, scene_test_labels = generate_scenes(test_pc, test_labels, N_TEST, n_objects = N_OBJECTS_IN_SCENE)
assert scene_train_pc.shape == (N_TRAIN, 3072, 3)
assert scene_train_labels.shape == (N_TRAIN, 3072, 10)
assert scene_test_pc.shape == (N_TEST, 3072, 3)
assert scene_test_labels.shape == (N_TEST, 3072, 10)
return scene_train_pc, scene_test_pc, scene_train_labels, scene_test_labels
# + colab={"base_uri": "https://localhost:8080/"} id="YtHY0_TgbgAf" outputId="53da7f2d-925b-4d1d-a373-04190f3e1b71"
scene_train_pc, scene_test_pc, scene_train_labels, scene_test_labels = segmentation_pc_gen(train_pc, train_labels, test_pc, test_labels)
# + id="0Gw8Q_TGce8_"
def visualize_scene(point_cloud, label):
"""
Utility function to visualize a point cloud
:param point_cloud: input point cloud
:type point_cloud: numpy array
"""
fig = plt.figure()
ax = fig.add_subplot(111, projection="3d")
ax.scatter(point_cloud[:, 0], point_cloud[:, 1], point_cloud[:, 2], c= label)
plt.show()
def get_point_class(scene_labels):
c = [np.where(r==1)[0][0] for r in scene_labels]
return c
# + colab={"base_uri": "https://localhost:8080/", "height": 248} id="BeTElRvCp_BL" outputId="685e35e1-1d94-49c2-d09b-fd8544fd81fc"
IDX = 38
visualize_scene(scene_train_pc[IDX], get_point_class(scene_train_labels[IDX]))
# + id="lnEhNJ-TfZnS"
import pickle
# once loaded save the numpy arrays to pickle files to use later
pickle.dump(scene_train_pc, open("scene_trainpc.pkl", "wb"))
pickle.dump(scene_test_pc, open("scene_testpc.pkl", "wb"))
pickle.dump(scene_train_labels, open("scene_trainlabels.pkl", "wb"))
pickle.dump(scene_test_labels, open("scene_testlabels.pkl", "wb"))
# pickle.dump(class_ids, open("class_ids.pkl", "wb"))
# + id="njAONFd9KQUG"
# labels = np.random.randint(0, 9, size = (10, ))
# get_tiled(one_hot(as_list(labels[3])))
# + id="g50TUO1nV8tN"
# print(one_hot([2]))
# some_pc = np.random.random(size= (1024, 3))
# print(some_pc)
# print(pc_transform(some_pc))
# pcs = np.random.random(size = (10, 1024, 3))
# labels = np.random.randint(0, 9, size = (10, ))
# arr1, arr2 = generate_scenes(pcs, labels, n_scenes = 5, n_objects= 3 )
# print(np.random.random(size = (10, 1024, 3)))
# print(np.random.randint(0, 9, size = 10))
# + id="UFSlzgbxsYaQ"
def test_assert():
a = np.array([[1, 3, 5], [2, 4, 6]])
sf = np.diag([2, 2, 2])
print(sf)
b = np.dot(a, sf.T)
print(b.dtype)
assert (b == [[2, 6, 10], [4, 8, 12]]).all()
assert True == True
# assert 1//2 == 1
print([[1, 2], [3,4]] == [[1, 2], [3, 4]])
print(b == [[2, 6, 10], [4, 8, 12]])
# + colab={"base_uri": "https://localhost:8080/"} id="Bf30HVmufWhh" outputId="e67280ce-2994-42cd-9d58-d142566c8bf2"
# arr1.shape, arr2.shape
# + id="BdGzB_i0BEPK"
# arr2
# + id="VtmagB62qWR8"
# arr1[0], arr1[3]
# + colab={"base_uri": "https://localhost:8080/"} id="Mx8HYN67nF38" outputId="c8d82d89-dae8-4edb-9786-752526670288"
# arr1[0].shape
# + colab={"base_uri": "https://localhost:8080/"} id="jL3kSsvZo6Yq" outputId="cca65471-d7a0-4576-c6b1-763999edef88"
# arr1.min(), arr1.max()
# + colab={"base_uri": "https://localhost:8080/", "height": 248} id="SRudMQKQnE1W" outputId="f2619d8b-33e1-4fe7-c970-e4c3ef7fcad4"
# visualize_cloud(arr1[0])
# + colab={"base_uri": "https://localhost:8080/"} id="WxciT5SdkwuK" outputId="f134dabf-4ed4-426c-f470-6bdf6aa58d93"
# arr1.shape
# + colab={"base_uri": "https://localhost:8080/"} id="gw0GhVQmJsXh" outputId="9142240f-1fd9-4973-c7c7-15cd8714f9bb"
# trying ideas on hw to pick unique type of objects in a scene
"""import pandas as pd
labels = np.random.randint(0, 10 , size = (100, ))
def get_random_index(labels):
df = pd.DataFrame(labels, columns = ["labels"])
df["index"] = df.index
df.head()
df1 = df.groupby("labels")["index"].apply(list).apply(np.random.choice)
random_indexes = df1.sample(n=3, replace = False).tolist()
return random_indexes
get_random_index(labels)"""
# + id="LqpBbPma55nV"
"""import random
def generate_scene(pc_with_labels, n_objects):
"takes the number of objects and return a combined point cloud"
# Do we duplicate an object in a scene? If not, random indexes have to be generated such that
# all objects are represented once only in the scene.
combined_pc = []
label_group = []
N_RECORDS = len(pc_with_labels)
l = range(N_RECORDS)
random.seed(123)
random.shuffle(l)
index_values = random.sample(list(enumerate(l)), n_objects)
# will return the index and the value of an iterable as a tuple
return combined_pc, label_group
def combine_object_pcs(pc_array, labels):
composed_pcs_array = []
composed_labels_array = []
# As the number of objects in each scene will be variable, the array shape will vary between records?
# Please ask the supervisor if we need to keep same number of objects in each scene or do we need to resample the scene
pc_with_labels = np.vstack(pc_array, labels)
for i in range(2000):
with num_objects in random.randint(2, 8):
new_pc, label_group = generate_scene(pc_with_labels, num_obejcts)
composed_pcs_array.append(new_pc)
composed_labels_array.append(label_group)
return composed_pcs_array, composed_labels_array"""
# + id="O_bez8QyMWXS"
# some_dataset = tf.data.Dataset.from_tensor_slices((arr1, arr2))
# len(some_dataset),
# some_dataset.take(2)
# + id="u_Ba9BAHYiFS"
# Create tensorflow data loaders from the numpy arrays
train_dataset = tf.data.Dataset.from_tensor_slices((train_pc, train_labels))
test_dataset = tf.data.Dataset.from_tensor_slices((test_pc, test_labels))
# + id="FmvZl5X0hNXr"
# Create tensorflow data loaders from the numpy arrays
scene_train_dataset = tf.data.Dataset.from_tensor_slices((scene_train_pc, scene_train_labels))
scene_test_dataset = tf.data.Dataset.from_tensor_slices((scene_test_pc, scene_test_labels))
# + colab={"base_uri": "https://localhost:8080/"} id="Y6UzJdQZapys" outputId="b878cabf-9d53-4814-f43e-323f38461a91"
len(train_pc)
# + [markdown] id="nwymY0pYYpkH"
# ### Data Augmentation
# + id="7LrAVLDVYoke"
# 3. Perform data augmentation by adding noise and shuffling the dataset.
# In this step you need to fill in the utils.add_noise_and_shuffle function
# to add noise to the sampled points and shuffle the points around
batch_size = 16 # <- You can modify this value as needed
train_dataset = train_dataset.shuffle(len(train_pc)).map(add_noise_and_shuffle).batch(batch_size)
test_dataset = test_dataset.shuffle(len(test_pc)).batch(batch_size)
# + [markdown] id="bEbDlmoQgxFi"
# ## Building the model from here
# + [markdown] id="hD8nmcseg5oi"
# ### Model Building Utilities - Taken from network.py
# + id="P_hENQLphyBk"
import os
import numpy as np
import tensorflow as tf
import trimesh.sample
from tensorflow import keras
from tensorflow.keras import layers
from matplotlib import pyplot as plt
import pickle
import keras.backend as K
# import network
# import utils
def tnet(inputs, num_features):
"""
This is the core t-net of the pointnet paper
:param inputs: the input tensor
:type inputs: tensor
:param num_features: number of features in the tensor (3 for point cloud, N if N features)
:type num_features: int
:return: output tensor
:rtype: tensor
"""
# Initalise bias as the indentity matrix
bias = keras.initializers.Constant(np.eye(num_features).flatten())
# TODO: Build the tnet with the following layers
# Some convolutional layers (1D) - with batch normalization, RELU activation
x = inputs
# x = layers.Input(shape= [None, 3072, 3])(x)
x = layers.Conv1D(filters=64, kernel_size=7, strides=2, padding="same")(x) # produces 1D feature map, 7 kernels => 7 1D sequences
x = layers.BatchNormalization()(x)
x = layers.Activation(tf.nn.relu)(x)
x = layers.Conv1D(filters=128, kernel_size=3, strides=2, padding="same")(x)
x = layers.BatchNormalization()(x)
x = layers.Activation(tf.nn.relu)(x)
x = layers.Conv1D(filters= 512, kernel_size=3, strides=2, padding="same")(x)
x = layers.BatchNormalization()(x)
x = layers.Activation(tf.nn.relu)(x)
# Global max pooling
x = layers.GlobalMaxPool1D()(x)
# Some dense fully connected layers - with batch normalization, RELU activation
# x = layers.Flatten()(x)
x = layers.Dense(units=512)(x)
x = layers.BatchNormalization()(x)
x = layers.Activation("relu")(x)
x = layers.Dense(units=192)(x)
x = layers.BatchNormalization()(x)
x = layers.Activation("relu")(x)
# final layer with custom regularizer on the output
# TODO: this custom regularizer needs to be defined
x = layers.Dense(
num_features * num_features,
kernel_initializer="zeros",
bias_initializer=bias,
activity_regularizer=CustomRegularizer(num_features))(x)
feat_t = layers.Reshape((num_features, num_features))(x)
# Apply affine transformation to input features
return layers.Dot(axes=(2, 1))([inputs, feat_t])
class CustomRegularizer(keras.regularizers.Regularizer):
"""
This class implements a regularizer that makes the output to be orthogonal.
In other words, it adds a loss |I-AA^T|^2 on the output A. Equation 2 of the paper.
"""
def __init__(self, dim, weight=0.001):
"""
Initializes the class
:param dim: dimensions of the input tensor
:type dim: int
:param weight: weight to apply on the regularizer
:type weight: float
"""
self.dim = dim
self.weight = weight
def __call__(self, x):
# TODO: define the custom regularizer here
x = tf.reshape(x, (-1, self.dim, self.dim))
# print(f"reshaped input shape: {K.shape(x)}")
# compute the outer product and reshape it to batch size x num_features x num_features
# outerproduct = tf.tensordot(x, tf.transpose(x), axes = [1, 2])
outerproduct = tf.linalg.matmul(a =x, b = x, transpose_b = True)
# print(f"outerproduct shape: {K.shape(outerproduct)}")
outerproduct = tf.reshape(outerproduct, (K.shape(x)[0], K.shape(x)[1], K.shape(x)[1]))
# Compute (I-outerproduct)^2 element wise. use tf.square()
z = tf.square( tf.eye(K.shape(x)[1]) - outerproduct)
# Apply weight
z = self.weight*z
# Compute reduce sum using tf.reduce_sum()
output = tf.reduce_sum(z)
return output
# + colab={"base_uri": "https://localhost:8080/"} id="3ZIBAEu_fjkJ" outputId="f39cc498-e8ab-438a-9aeb-e483adf64007"
# print(arr1[:4].shape)
# x = np.random.random((4, 1800, 3))
# dim = 3
# x.shape, tf.reshape(x, (-1, 3, 3)).shape
# + colab={"base_uri": "https://localhost:8080/"} id="LSX8uStJkC5Z" outputId="0d1b405d-db52-496c-c6b2-0af759d4c722"
x1 = tnet(train_pc, 3)
x1
# + id="N1LyJSUw0KER"
# x = np.random.random((4, 1024, 3))
# x1 = tnet(x, 3)
# x1
# + id="zSrLT6CCgKZc"
def pointnet_classifier(num_points_per_cloud, num_classes):
"""
This is the object classifier version of PointNet
:param inputs: input point clouds tensor
:type inputs: tensor
:param num_classes: number of classes
:type num_classes: int
:return: the predicted labels
:rtype: tensor
"""
# TODO: build the network using the following layers
# apply tnet to the input data
inputs = keras.Input(shape=(num_points_per_cloud, 3))
f_3 = tnet(inputs, 3)
# extract features using some Convolutional Layers - with batch normalization and RELU activation
x = layers.Conv1D(filters= 32, kernel_size= 3, strides =1, activation= "relu")(f_3)
x = layers.BatchNormalization()(x)
x = layers.Conv1D(filters= 64, kernel_size= 2, strides =1, activation= "relu")(x)
x = layers.BatchNormalization()(x)
# apply tnet on the feature vector
f_64 = tnet(x, 64) # output of this layer to be used in segmentation model
# extract features using some Convolutional Layers - with batch normalization and RELU activation
x = layers.Conv1D(filters= 128, kernel_size= 2, strides =1, activation= "relu")(f_64)
x = layers.BatchNormalization()(x)
x = layers.Conv1D(filters= 256, kernel_size= 2, strides =1, activation="relu")(x)
x = layers.BatchNormalization()(x)
x = layers.Conv1D(filters= 512, kernel_size= 2, strides =1, activation="relu")(x)
x = layers.BatchNormalization()(x)
# apply 1D global max pooling
global_features = layers.GlobalMaxPool1D()(x) # output of this layer to be used in segmentation model
# Add a few dense layers with dropout between the layers
x = layers.Dense(units = 128, activation = "relu")(global_features)
x = layers.Dense(units = 64, activation = "relu")(x)
# Finally predict classes using a dense layer with a softmax activation
outputs = layers.Dense(num_classes, activation="softmax")(x)
model = keras.Model(inputs=inputs, outputs=outputs, name="pointnet")
return model
def pointnet_segmenter(num_points_per_cloud, labels):
"""
This is the semantic segmentation version of Pointnet
:param inputs: input point cloud
:type inputs: tensor
:param labels: labels for each point of the point cloud
:type labels: tensor
:return: predicted labels for each point of the point cloud
:rtype: tensor
"""
# TODO: build the network using the following layers
# apply tnet to the input data
inputs = keras.Input(shape=(num_points_per_cloud, 3))
f_3 = tnet(inputs, 3)
# extract features using some Convolutional Layers - with batch normalization and RELU activation
x = layers.Conv1D(filters= 32, kernel_size= 3, strides =1, activation= "relu")(f_3)
x = layers.BatchNormalization()(x)
x = layers.Conv1D(filters= 64, kernel_size= 3, strides =1, activation= "relu")(x)
x = layers.BatchNormalization()(x)
# apply tnet on the feature vector
f_64 = tnet(x, 64)
# extract features using some Convolutional Layers - with batch normalization and RELU activation
x = layers.Conv1D(filters= 128, kernel_size= 2, strides =1, activation= "relu")(f_64)
x = layers.BatchNormalization()(x)
x = layers.Conv1D(filters= 256, kernel_size= 2, strides =1, activation="relu")(x)
x = layers.BatchNormalization()(x)
x = layers.Conv1D(filters= 1024, kernel_size= 2, strides =1, activation="relu")(x)
x = layers.BatchNormalization()(x)
# apply 1D global max pooling
global_features = layers.GlobalMaxPool1D()(x)
# concatenate these features with the earlier features (f)
# you can also use skip connections if you like
z = layers.concatenate([f_64, global_features])
# extract features using some Convolutional Layers - with batch normalization and RELU activation
x = layers.Conv1D(filters= 512, kernel_size= 3, strides =1, activation= "relu")(z)
x = layers.BatchNormalization()(x)
x = layers.Conv1D(filters= 256, kernel_size= 3, strides =1, activation="relu")(x)
x = layers.BatchNormalization()(x)
x = layers.Conv1D(filters= 128, kernel_size= 3, strides =1, activation="relu")(x)
x = layers.BatchNormalization()(x)
x = layers.Dense(128, activation = "relu")(x)
outputs = layers.Dense(128, activation = "softmax")(x)
# return the output
model = keras.Model(inputs=inputs, outputs=outputs, name="segpointnet")
return model
# + [markdown] id="XYFsv_Atg2VT"
# ### Classification Model
# + id="cUqZZGZ_lJPe" colab={"base_uri": "https://localhost:8080/"} outputId="b53abf03-683c-4111-f6e1-46475b94af68"
# 1. Fill in the skeleton code given in the network.py file
num_points_per_cloud = 1024
model = pointnet_classifier(num_points_per_cloud, num_classes=10)
model.summary()
# outputs = network.pointnet_segmenter(inputs, train_labels)
model.compile(
loss="mse", # <- choose a suitable loss function
optimizer=keras.optimizers.Adam(learning_rate=0.001), # <- you may modify this if you like
metrics=["accuracy"] # <- choose a suitable metric, https://www.tensorflow.org/api_docs/python/tf/keras/metrics
)
# mirrored_strategy = tf.distribute.MirroredStrategy()
# with mirrored_strategy.scope():
# # build the network and visualize its architecture
# # model = keras.Model(inputs=inputs, outputs=outputs, name="pointnet")
# # 2. Set the loss function, optimizer and metrics to print
# model.compile(
# loss="categorical_crossentropy", # <- choose a suitable loss function
# optimizer=keras.optimizers.Adam(learning_rate=0.001), # <- you may modify this if you like
# metrics=["accuracy"] # <- choose a suitable metric, https://www.tensorflow.org/api_docs/python/tf/keras/metrics
# )
# + colab={"base_uri": "https://localhost:8080/"} id="sYSCDt2W8Q19" outputId="0f9cd64f-ded8-4d4f-9a4e-8f70f8dd68d9"
# train the network
num_epochs = 25 # <- change this value as needed
model.fit(train_dataset, epochs=num_epochs, validation_data=test_dataset)
# + id="PqjJ9or38qjD"
# + id="PoO_wyWDlXUV"
# visualize results
data = test_dataset.take(1)
point_clouds, labels = list(data)[0] # this is one batch of data
# predict labels using the model
preds = model.predict(point_clouds)
preds = tf.math.argmax(preds, -1)
# + id="esHSATpxsT36"
# 3. Display some clouds using matplotlib scatter plot along with true and predicted labels
# 4. Display a confusion matrix
# + [markdown] id="YWqUmW6Jg-rd"
# ### Segmentation Model
# + id="mJiU2y4n2goi" colab={"base_uri": "https://localhost:8080/", "height": 460} outputId="2704d724-9a1e-46c0-cfa9-e605445e4610"
num_points_per_cloud = 3072
# inputs = keras.Input(shape=(num_points_per_cloud, 3))
# outputs = pointnet_classifier(inputs, num_classes=10)
outputs = pointnet_segmenter(num_points_per_cloud, scene_train_labels)
model = keras.Model(inputs=inputs, outputs=outputs, name="segmentnet")
model.summary()
# + id="cpFrrft_rghc"
model.compile(loss = "categorical_crossentropy", optimizer = tf.optimizers.Adam, metrics = ["accuracy"])
# + id="XHjyob9Krz8B"
N_EPOCHS = 5
model.fit(scene_train_dataset
epochs = N_EPOCHS,
verbose = 1,
validation_data = scene_test_dataset)
|
point_cloud.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Part 2: Checkouts, Branching, & Merging
#
# This section deals with navigating repository history, creating & merging
# branches, and understanding conflicts.
# ### The Hangar Workflow
#
# The hangar workflow is intended to mimic common ``git`` workflows in which small
# incremental changes are made and committed on dedicated ``topic`` branches.
# After the ``topic`` has been adequatly set, ``topic`` branch is merged into
# a separate branch (commonly referred to as ``master``, though it need not to be the
# actual branch named ``"master"``), where well vetted and more permanent changes
# are kept.
#
# Create Branch -> Checkout Branch -> Make Changes -> Commit
#
# #### Making the Initial Commit
#
# Let's initialize a new repository and see how branching works in Hangar:
#
# <!-- However, unlike GIT, remember that it is not possible to make changes in a DETACHED HEAD state. Hangar enforces the requirement that all work is performed at the tip of a branch. -->
from hangar import Repository
import numpy as np
repo = Repository(path='foo/pth')
repo_pth = repo.init(user_name='Test User', user_email='<EMAIL>')
# When a repository is first initialized, it has no history, no commits.
repo.log() # -> returns None
# Though the repository is essentially empty at this point in time, there is one
# thing which is present: a branch with the name: ``"master"``.
repo.list_branches()
# This ``"master"`` is the branch we make our first commit on; until we do, the
# repository is in a semi-unstable state; with no history or contents, most of the
# functionality of a repository (to store, retrieve, and work with versions of
# data across time) just isn't possible. A significant portion of otherwise
# standard operations will generally flat out refuse to execute (ie. read-only
# checkouts, log, push, etc.) until the first commit is made.
#
# One of the only options available at this point is to create a
# write-enabled checkout on the ``"master"`` branch and to begin to add data so we
# can make a commit. Let’s do that now:
co = repo.checkout(write=True)
# As expected, there are no arraysets nor metadata samples recorded in the checkout.
print(f'number of metadata keys: {len(co.metadata)}')
print(f'number of arraysets: {len(co.arraysets)}')
# Let’s add a dummy array just to put something in the repository history to
# commit. We'll then close the checkout so we can explore some useful tools which
# depend on having at least one historical record (commit) in the repo.
dummy = np.arange(10, dtype=np.uint16)
aset = co.arraysets.init_arrayset(name='dummy_arrayset', prototype=dummy)
aset['0'] = dummy
initialCommitHash = co.commit('first commit with a single sample added to a dummy arrayset')
co.close()
# If we check the history now, we can see our first commit hash, and that it is labeled with the branch name `"master"`
repo.log()
# So now our repository contains:
# - [A commit](api.rst#hangar.checkout.WriterCheckout.commit_hash): a fully
# independent description of the entire repository state as
# it existed at some point in time. A commit is identified by a `commit_hash`.
# - [A branch](api.rst#hangar.checkout.WriterCheckout.branch_name): a label
# pointing to a particular `commit` / `commit_hash`.
#
# Once committed, it is not possible to remove, modify, or otherwise tamper with
# the contents of a commit in any way. It is a permanent record, which Hangar has
# no method to change once written to disk.
#
# In addition, as a `commit_hash` is not only calculated from the `commit` ’s
# contents, but from the `commit_hash` of its parents (more on this to follow),
# knowing a single top-level `commit_hash` allows us to verify the integrity of
# the entire repository history. This fundamental behavior holds even in cases of
# disk-corruption or malicious use.
#
# ### Working with Checkouts & Branches
#
# As mentioned in the first tutorial, we work with the data in a repository through
# a [checkout](api.rst#hangar.repository.Repository.checkout). There are two types
# of checkouts (each of which have different uses and abilities):
#
# **[Checking out a branch / commit for reading:](api.rst#read-only-checkout)** is
# the process of retrieving records describing repository state at some point in
# time, and setting up access to the referenced data.
#
# - Any number of read checkout processes can operate on a repository (on
# any number of commits) at the same time.
#
# **[Checking out a branch for writing:](api.rst#write-enabled-checkout)** is the
# process of setting up a (mutable) ``staging area`` to temporarily gather
# record references / data before all changes have been made and staging area
# contents are committed in a new permanent record of history (a `commit`).
#
# - Only one write-enabled checkout can ever be operating in a repository
# at a time.
# - When initially creating the checkout, the `staging area` is not
# actually “empty”. Instead, it has the full contents of the last `commit`
# referenced by a branch’s `HEAD`. These records can be removed / mutated / added
# to in any way to form the next `commit`. The new `commit` retains a
# permanent reference identifying the previous ``HEAD`` ``commit`` was used as
# its base `staging area`.
# - On commit, the branch which was checked out has its ``HEAD`` pointer
# value updated to the new `commit`’s `commit_hash`. A write-enabled
# checkout starting from the same branch will now use that `commit`’s
# record content as the base for its `staging area`.
#
# #### Creating a branch
#
# A branch is an individual series of changes / commits which diverge from the main
# history of the repository at some point in time. All changes made along a branch
# are completely isolated from those on other branches. After some point in time,
# changes made in a disparate branches can be unified through an automatic
# `merge` process (described in detail later in this tutorial). In general, the
# `Hangar` branching model is semantically identical to the `Git` one; The one exception
# is that in Hangar, a branch must always have a `name` and a `base_commit`. (No
# "Detached HEAD state" is possible for a `write-enabled` checkout). If No `base_commit` is
# specified, the current writer branch `HEAD` `commit` is used as the `base_commit`
# hash for the branch automatically.
#
# Hangar branches have the same lightweight and performant properties which
# make working with `Git` branches so appealing - they are cheap and easy to use,
# create, and discard (if necessary).
#
# To create a branch, use the [create_branch()](api.rst#hangar.repository.Repository.create_branch)
# method.
branch_1 = repo.create_branch(name='testbranch')
branch_1
# We use the [list_branches()](api.rst#hangar.repository.Repository.list_branches) and [log()](api.rst#hangar.repository.Repository.log) methods to see that a new branch named `testbranch` has been created and is indeed pointing to our initial commit.
print(f'branch names: {repo.list_branches()} \n')
repo.log()
# If instead, we actually specify the base commit (with a different branch
# name) we see we do actually get a third branch. pointing to the same commit as
# `master` and `testbranch`
branch_2 = repo.create_branch(name='new', base_commit=initialCommitHash)
branch_2
repo.log()
# #### Making changes on a branch
#
# Let’s make some changes on the `new` branch to see how things work.
#
# We can see that the data we added previously is still here (`dummy` arrayset containing
# one sample labeled `0`).
co = repo.checkout(write=True, branch='new')
co.arraysets
co.arraysets['dummy_arrayset']
co.arraysets['dummy_arrayset']['0']
# Let's add another sample to the `dummy_arrayset` called `1`
# +
arr = np.arange(10, dtype=np.uint16)
# let's increment values so that `0` and `1` aren't set to the same thing
arr += 1
co['dummy_arrayset', '1'] = arr
# -
# We can see that in this checkout, there are indeed two samples in the `dummy_arrayset`:
len(co.arraysets['dummy_arrayset'])
# That's all, let's commit this and be done with this branch.
co.commit('commit on `new` branch adding a sample to dummy_arrayset')
co.close()
# #### How do changes appear when made on a branch?
#
# If we look at the log, we see that the branch we were on (`new`) is a commit ahead of `master` and `testbranch`
repo.log()
# The meaning is exactly what one would intuit. We made some changes, they were
# reflected on the `new` branch, but the `master` and `testbranch` branches
# were not impacted at all, nor were any of the commits!
# ### Merging (Part 1) Fast-Forward Merges
#
# Say we like the changes we made on the ``new`` branch so much that we want them
# to be included into our ``master`` branch! How do we make this happen for this
# scenario??
#
# Well, the history between the ``HEAD`` of the ``new`` and the ``HEAD`` of the
# ``master`` branch is perfectly linear. In fact, when we began making changes
# on ``new``, our staging area was *identical* to what the ``master`` ``HEAD``
# commit references are right now!
#
# If you’ll remember that a branch is just a pointer which assigns some ``name``
# to a ``commit_hash``, it becomes apparent that a merge in this case really
# doesn’t involve any work at all. With a linear history between ``master`` and
# ``new``, any ``commits`` exsting along the path between the ``HEAD`` of
# ``new`` and ``master`` are the only changes which are introduced, and we can
# be sure that this is the only view of the data records which can exist!
#
# What this means in practice is that for this type of merge, we can just update
# the ``HEAD`` of ``master`` to point to the ``HEAD`` of ``"new"``, and the
# merge is complete.
#
# This situation is referred to as a **Fast Forward (FF) Merge**. A FF merge is
# safe to perform any time a linear history lies between the ``HEAD`` of some
# ``topic`` and ``base`` branch, regardless of how many commits or changes which
# were introduced.
#
# For other situations, a more complicated **Three Way Merge** is required. This
# merge method will be explained a bit more later in this tutorial.
co = repo.checkout(write=True, branch='master')
# #### Performing the Merge
#
# In practice, you’ll never need to know the details of the merge theory explained
# above (or even remember it exists). Hangar automatically figures out which merge
# algorithms should be used and then performed whatever calculations are needed to
# compute the results.
#
# As a user, merging in Hangar is a one-liner! just use the [merge()](api.rst#hangar.checkout.WriterCheckout.merge)
# method from a `write-enabled` checkout (shown below), or the analogous methods method
# from the Repository Object [repo.merge()](api.rst#hangar.repository.Repository.merge)
# (if not already working with a `write-enabled` checkout object).
co.merge(message='message for commit (not used for FF merge)', dev_branch='new')
# Let's check the log!
repo.log()
co.branch_name
co.commit_hash
co.arraysets['dummy_arrayset']
# As you can see, everything is as it should be!
co.close()
# #### Making changes to introduce diverged histories
#
# Let’s now go back to our `testbranch` branch and make some changes there so
# we can see what happens when changes don’t follow a linear history.
co = repo.checkout(write=True, branch='testbranch')
co.arraysets
co.arraysets['dummy_arrayset']
# We will start by mutating sample `0` in `dummy_arrayset` to a different value
old_arr = co['dummy_arrayset', '0']
new_arr = old_arr + 50
new_arr
co['dummy_arrayset', '0'] = new_arr
# Let’s make a commit here, then add some metadata and make a new commit (all on
# the `testbranch` branch).
co.commit('mutated sample `0` of `dummy_arrayset` to new value')
repo.log()
co.metadata['hello'] = 'world'
co.commit('added hellow world metadata')
co.close()
# Looking at our history how, we see that none of the original branches reference
# our first commit anymore.
repo.log()
# We can check the history of the `master` branch by specifying it as an argument to the `log()` method.
repo.log('master')
# ### Merging (Part 2) Three Way Merge
#
# If we now want to merge the changes on `testbranch` into `master`, we can't just follow a simple linear history; **the branches have diverged**.
#
# For this case, Hangar implements a **Three Way Merge** algorithm which does the following:
# - Find the most recent common ancestor `commit` present in both the `testbranch` and `master` branches
# - Compute what changed between the common ancestor and each branch's `HEAD` commit
# - Check if any of the changes conflict with each other (more on this in a later tutorial)
# - If no conflicts are present, compute the results of the merge between the two sets of changes
# - Create a new `commit` containing the merge results reference both branch `HEAD`s as parents of the new `commit`, and update the `base` branch `HEAD` to that new `commit`'s `commit_hash`
co = repo.checkout(write=True, branch='master')
# Once again, as a user, the details are completely irrelevant, and the operation
# occurs from the same one-liner call we used before for the FF Merge.
co.merge(message='merge of testbranch into master', dev_branch='testbranch')
# If we now look at the log, we see that this has a much different look than
# before. The three way merge results in a history which references changes made
# in both diverged branches, and unifies them in a single ``commit``
repo.log()
# #### Manually inspecting the merge result to verify it matches our expectations
#
# `dummy_arrayset` should contain two arrays, key `1` was set in the previous
# commit originally made in `new` and merged into `master`. Key `0` was
# mutated in `testbranch` and unchanged in `master`, so the update from
# `testbranch` is kept.
#
# There should be one metadata sample with they key `hello` and the value
# ``"world"``.
co.arraysets
co.arraysets['dummy_arrayset']
co['dummy_arrayset', ['0', '1']]
co.metadata
co.metadata['hello']
# **The Merge was a success!**
co.close()
# ### Conflicts
#
# Now that we've seen merging in action, the next step is to talk about conflicts.
#
# #### How Are Conflicts Detected?
#
# Any merge conflicts can be identified and addressed ahead of running a `merge`
# command by using the built in [diff](api.rst#hangar.diff.WriterUserDiff) tools.
# When diffing commits, Hangar will provide a list of conflicts which it identifies.
# In general these fall into 4 categories:
#
# 1. **Additions** in both branches which created new keys (samples /
# arraysets / metadata) with non-compatible values. For samples &
# metadata, the hash of the data is compared, for arraysets, the schema
# specification is checked for compatibility in a method custom to the
# internal workings of Hangar.
# 2. **Removal** in `Master Commit/Branch` **& Mutation** in `Dev Commit / Branch`. Applies for samples, arraysets, and metadata identically.
# 3. **Mutation** in `Dev Commit/Branch` **& Removal** in `Master Commit / Branch`. Applies for samples, arraysets, and metadata identically.
# 4. **Mutations** on keys of both branches to non-compatible values. For
# samples & metadata, the hash of the data is compared; for arraysets, the
# schema specification is checked for compatibility in a method custom to the
# internal workings of Hangar.
#
# #### Let's make a merge conflict
#
# To force a conflict, we are going to checkout the `new` branch and set the
# metadata key `hello` to the value `foo conflict... BOO!`. Then if we try
# to merge this into the `testbranch` branch (which set `hello` to a value
# of `world`) we see how hangar will identify the conflict and halt without
# making any changes.
#
# Automated conflict resolution will be introduced in a future version of Hangar,
# for now it is up to the user to manually resolve conflicts by making any
# necessary changes in each branch before reattempting a merge operation.
co = repo.checkout(write=True, branch='new')
co.metadata['hello'] = 'foo conflict... BOO!'
co.commit ('commit on new branch to hello metadata key so we can demonstrate a conflict')
repo.log()
# **When we attempt the merge, an exception is thrown telling us there is a conflict!**
co.merge(message='this merge should not happen', dev_branch='testbranch')
# #### Checking for Conflicts
#
# Alternatively, use the diff methods on a checkout to test for conflicts before attempting a merge.
#
# It is possible to diff between a checkout object and:
#
# 1. Another branch ([diff.branch()](api.rst#hangar.diff.WriterUserDiff.branch))
# 2. A specified commit ([diff.commit()](api.rst#hangar.diff.WriterUserDiff.commit))
# 3. Changes made in the staging area before a commit is made
# ([diff.staged()](api.rst#hangar.diff.WriterUserDiff.staged))
# (for `write-enabled` checkouts only.)
#
# Or via the [CLI status tool](cli.rst#hangar-status) between the staging area and any branch/commit
# (only a human readable summary is produced).
merge_results, conflicts_found = co.diff.branch('testbranch')
conflicts_found
conflicts_found.t1.metadata
# The type codes for a `Conflicts` `namedtuple` such as the one we saw:
#
# Conflicts(t1=('hello',), t21=(), t22=(), t3=(), conflict=True)
#
# are as follow:
#
# - ``t1``: Addition of key in master AND dev with different values.
# - ``t21``: Removed key in master, mutated value in dev.
# - ``t22``: Removed key in dev, mutated value in master.
# - ``t3``: Mutated key in both master AND dev to different values.
# - ``conflict``: Bool indicating if any type of conflict is present.
# #### To resolve, remove the conflict
del co.metadata['hello']
co.metadata['resolved'] = 'conflict by removing hello key'
co.commit('commit which removes conflicting metadata key')
co.merge(message='this merge succeeds as it no longer has a conflict', dev_branch='testbranch')
# We can verify that history looks as we would expect via the log!
repo.log()
|
docs/Tutorial-002.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Table of Content
# 1. Introduction of Tool
# 2. Features of Tool
# 3. Requirements of Tool
# 4. How to Work
# 5. Folders Description
# 6. Toy examples
# ### 1. Introduction of Tool
# Dynamic Analyzer tool has leverage the capabilities of LibClang to analyze the dynamic
# aspects of C language programs. LibClang is considered as a stable high level C interface to clang and used for high level abstractions by traversing an AST through a cursor.
#
# ### 2. Features of Tool
# Besides dynamic aspects of a program, the tool would help to obtain the following features as follow.
#
# • To extract the name, mangled name and qualified name of functions
# • To extract the function definitions with its line number of source code.
# • To extract the type of a function such as Template function, a method or normal function
# • To extract the calling points location of a function along with parameters
# • To extract the cyclomatic complexity of each function
# • To extract the longest functions call complexity
# • To accumulate the total cyclomatic complexity
# ### 3. Requirments of Tool
# • llvm (Preferable llvm-11.0.0)
# • Clang
# • Python ( Supports both Python 2 ad 3)
# • Conda
# • Ubuntu/macOS
# ### 3. How to work
# Firstly open the Terminal and clone the repository as follow.
# > git clone https://github.com/HPCL/code-analysis.git
# Code-analysis folder will be created on system current root path. It will be open by applying the following command.
# > cd code-analysis
# In code-analysis folder execute the environments.yml file as follow.
# > conda env create –f environments.yml
# The content of environments.yml is covering the dependencies which are used to execute the Dynamic analyzer tool smoothly. If you are executing environments.yml first time and initially dependency requirements are not available on your system then it will take few seconds.
#
# To check the status of all tests, apply the following commands
# > cd /Users/…/code-analysis/Dynamic/tests
# > pytest
# To execute a specific test, apply the following command and observe the output.
# > pytest –q –s –name example3.c test_Function_List.py
# This command is passing a file example3.c as an argument with reference –name to test_Function_List.py. In case of passing more than one arguments with references, you can update conftest.py file in tests folder.
# ### 4. Folder Description
# The hierarchy of clone folder is as follows
#
# Code-analysis
# ------Dynamic
# ---------- examples
# ---------- src
# ---------- tests
# ---------- README.md
# ---------- requirments.txt
# ------Static
# ------__init__.py
# -------environments.yml
#
# src Folder:
#
# This folder contains the python scripts which are used to cover the different aspects of dynamic analysis.
#
# tests Folder:
#
# This folder contain three types of files as follows
# 1. Tests script written in python and can be executed through pytest.
# 2. Toy examples of *.c to understand the functionality of scripts of src folder and are passed as an argument when a test is checked through pytest.
# 3. Conftest.py file is used to declare the arguments reference which are passed through pytest. Initially, --name reference is given for passing *.c file as argument of pytest command.
#
# ### 5. Toy Examples
# ### Example-1: (Example2 in tests folder)
#
# void Large(int a, int b)
# {
# if(a>=b)
# printf("large number is %d", a);
# else
# printf("Large number is %d", b);
# }
#
# void main() {
# int A=2, B=3, Larg;
# if(A == B)
# printf("Both number are same");
# else
# Large(A, B);
# Large(5,4);
# printf("Bye");
# }
# ### Script-1 (Function_List.py) : Retreive information about all functions of example2.c
# +
import csv, os, glob
import sys
import clang.cindex
from clang.cindex import Config
os.environ['DYLD_LIBRARY_PATH']= '/usr/local/Cellar/llvm/11.0.0/lib/'
#Config.set_library_path('/usr/local/Cellar/llvm/11.0.0/lib')
#Config.set_library_path('/usr/local/anaconda3/lib/python3.8/site-packages/')
def Extract_Function_Names(file_name, tu):
List_of_Functions=[]
col=[]
col.append("File Name")
col.append("Function Name")
col.append("Mangled Name")
col.append("Function Type")
List_of_Functions.append(col)
filename = tu.cursor.spelling
for c in tu.cursor.walk_preorder():
if c.location.file is None:
pass
elif c.location.file.name != filename:
pass
elif c.kind == clang.cindex.CursorKind.FUNCTION_DECL or c.kind==clang.cindex.CursorKind.CXX_METHOD or c.kind==clang.cindex.CursorKind.FUNCTION_TEMPLATE:
col=[]
col.append(file_name)
col.append(c.spelling)
col.append(c.mangled_name)
if c.kind==clang.cindex.CursorKind.FUNCTION_TEMPLATE:
col.append('Template Function')
elif c.kind==clang.cindex.CursorKind.CXX_METHOD:
col.append("Class Method")
elif c.kind==clang.cindex.CursorKind.FUNCTION_DECL:
col.append("Function")
List_of_Functions.append(col)
return List_of_Functions
idx = clang.cindex.Index.create()
os.chdir("/Users/shussain/code-analysis/dynamic/tests")
tu = idx.parse("example2.c", args='-xc++ --std=c++11'.split())
List_of_Functions = Extract_Function_Names("example2.c", tu)
for f in List_of_Functions:
print (f)
# -
# ### Script-2 (Function_Definition_Location.py) : Retreive information functions definitions and their location of example2.c
# +
import csv, os, glob
import sys
import clang.cindex
from clang.cindex import Config
os.environ['DYLD_LIBRARY_PATH']= '/usr/local/Cellar/llvm/11.0.0/lib/'
File_Content_Array=[]
def File_to_Array(file_name):
with open(file_name) as file:
for line in file:
File_Content_Array.append(line)
#return File_Content_Array
def Extract_Line_Column(cursor):
x=str(cursor.location)
y=x.split(',')
line = y[1]
column=y[2]
line = line.split()
line=line[1]
column=column.split()
column = column[1]
return line, column
def Extract_Function_Qualified_Name(cursor):
if cursor is None:
return ''
elif cursor.kind== clang.cindex.CursorKind.TRANSLATION_UNIT:
return ''
else:
res = Extract_Function_Qualified_Name(cursor.semantic_parent)
if res != '':
return res + '::' + cursor.spelling
return cursor.spelling
def Extract_Function_Definition_Location(file_name, tu):
List_of_Functions=[]
File_to_Array(file_name)
# Heading row for the list of functions
col=[]
col.append("File Name")
col.append("Function Name")
col.append("Function Qualified Name")
col.append("Function Definition")
col.append("Line Number")
col.append("Column Number")
List_of_Functions.append(col)
filename = tu.cursor.spelling
for c in tu.cursor.walk_preorder():
if c.location.file is None:
pass
elif c.location.file.name != filename:
pass
elif c.kind == clang.cindex.CursorKind.FUNCTION_DECL or c.kind==clang.cindex.CursorKind.CXX_METHOD or c.kind==clang.cindex.CursorKind.FUNCTION_TEMPLATE:
col=[]
col.append(file_name)
col.append(c.spelling)
col.append(Extract_Function_Qualified_Name(c))
lin1, col1 = Extract_Line_Column(c)
col1=col1[:-1]
st=File_Content_Array[int(lin1)-1]
col.append(st.strip())
col.append(lin1)
col.append(col1)
List_of_Functions.append(col)
return List_of_Functions
idx = clang.cindex.Index.create()
os.chdir("/Users/shussain/code-analysis/dynamic/tests")
tu = idx.parse("example2.c", args='-xc++ --std=c++11'.split())
List_of_Function = Extract_Function_Definition_Location("example2.c", tu)
for f in List_of_Function:
print (f)
# -
# ### Script-3 (Function_Definition_Calls.py) : Retreive the calling points of functions including calling point definitions of example2.c
import csv, os, glob
import sys
import clang.cindex
from clang.cindex import Config
os.environ['DYLD_LIBRARY_PATH']= '/usr/local/Cellar/llvm/11.0.0/lib/'
File_Content_Array=[]
Function_Complexity=[]
List_of_Functions=[]
Function_List=[]
def File_to_Array(file_name):
with open(file_name) as file:
for line in file:
File_Content_Array.append(line)
#return File_Content_Array
def Extract_Line_Column(cursor):
x=str(cursor.location)
y=x.split(',')
line = y[1]
column=y[2]
line = line.split()
line=line[1]
column=column.split()
column = column[1]
return line, column
def Extract_Function_List(tu):
filename = tu.cursor.spelling
for c in tu.cursor.walk_preorder():
if c.location.file is None:
pass
elif c.location.file.name != filename:
pass
elif c.kind == clang.cindex.CursorKind.FUNCTION_DECL or c.kind==clang.cindex.CursorKind.CXX_METHOD or c.kind==clang.cindex.CursorKind.FUNCTION_TEMPLATE:
Function_List.append(c.spelling)
def Extract_Function_Definition_Calls(file_name, tu):
Extract_Function_List(tu)
Final_List=[]
col=[]
col.append("Function Name")
col.append("Name Space")
col.append("Calling Point")
col.append("Line#")
col.append("Column#")
Final_List.append(col)
Name_Space="Anonymous NameSpace"
File_to_Array(file_name)
filename = tu.cursor.spelling
for c in tu.cursor.get_tokens():
if c.location.file is None:
pass
elif c.location.file.name != filename:
pass
elif c.cursor.kind == clang.cindex.CursorKind.NAMESPACE:
Name_Space=c.spelling
elif (c.cursor.kind == clang.cindex.CursorKind.DECL_REF_EXPR or c.cursor.kind == clang.cindex.CursorKind.CALL_EXPR or c.cursor.kind == clang.cindex.CursorKind.OVERLOADED_DECL_REF) and c.spelling in Function_List: # or c.kind==clang.cindex.TokenKind.IDENTIFIER) and c.spelling in Function_List:
lin1, col1 = Extract_Line_Column(c)
st1=File_Content_Array[int(lin1)-1]
col=[]
col.append(c.spelling)
col.append(Name_Space)
col.append(st1)
col.append(lin1)
col.append(col1)
if col not in Final_List:
Final_List.append(col)
else:
col=[]
return Final_List
idx = clang.cindex.Index.create()
os.chdir("/Users/shussain/code-analysis/dynamic/tests")
tu = idx.parse("example2.c", args='-xc++ --std=c++11'.split())
List_of_Function_Calls = Extract_Function_Definition_Calls("example2.c", tu)
for f in List_of_Function_Calls:
print (f)
# ### Script-4 (Function_Complexity.py) : Retreive the information about functions and their complexities of example2.c
# +
import csv, os, glob
import sys
import clang.cindex
from clang.cindex import Config
os.environ['DYLD_LIBRARY_PATH']= '/usr/local/Cellar/llvm/11.0.0/lib/'
File_Content_Array=[]
Function_Complexity=[]
List_of_Functions=[]
Function_List=[]
Cursor_Types = {clang.cindex.CursorKind.IF_STMT, clang.cindex.CursorKind.WHILE_STMT, clang.cindex.CursorKind.FOR_STMT,clang.cindex.CursorKind.DEFAULT_STMT,clang.cindex.CursorKind.CASE_STMT, clang.cindex.CursorKind.COMPOUND_STMT}
Keywords= {"if", "case", "default", "for", "while", "else"}
# A function to copy the content of file into an array
def File_to_Array(file_name):
with open(file_name) as file:
for line in file:
File_Content_Array.append(line)
#return File_Content_Array
def Extract_Function_Complexity(tu):
cnt=0
flag=False
filename = tu.cursor.spelling
fn=""
for c in tu.cursor.get_tokens():
if c.location.file is None:
pass
elif c.location.file.name != filename:
pass
else:
cond1=c.cursor.kind in Cursor_Types
cond2=c.spelling in Keywords
if (c.cursor.kind==clang.cindex.CursorKind.FUNCTION_DECL or c.cursor.kind==clang.cindex.CursorKind.FUNCTION_TEMPLATE) and c.spelling in Function_List:
flag=True
fn=c.spelling
cnt=cnt+1
if cond1==True and cond2==True and (cnt<=len(Function_Complexity) and Function_List[cnt-1] ==fn):# and len(Function_List)==len(Function_Complexity)):
temp=Function_Complexity[cnt-1]
temp=temp+1
Function_Complexity[cnt-1]=temp
def Extract_Line_Column(cursor):
x=str(cursor.location)
y=x.split(',')
line = y[1]
column=y[2]
line = line.split()
line=line[1]
column=column.split()
column = column[1]
return line, column
def Extract_Function_Qualified_Name(cursor):
if cursor is None:
return ''
elif cursor.kind== clang.cindex.CursorKind.TRANSLATION_UNIT:
return ''
else:
res = Extract_Function_Qualified_Name(cursor.semantic_parent)
if res != '':
return res + '::' + cursor.spelling
return cursor.spelling
def Merge_Function_Complexity(Functions, Complexity):
Final_List=[]
# Heading row for the list of functions
col=[]
col.append("File Name")
col.append("Function Name")
col.append("Function Qualified Name")
col.append("Mangled Name")
col.append("Name Space")
col.append("Function Definition")
col.append("Line Number")
col.append("Column Number")
col.append("Cyclomatic Complexity")
Final_List.append(col)
cnt=0
for func in Functions:
col=[]
col.append(func[0])
col.append(func[1])
col.append(func[2])
col.append(func[3])
col.append(func[4])
col.append(func[5])
col.append(func[6])
col.append(func[7])
col.append(Complexity[cnt])
Final_List.append(col)
cnt=cnt+1
return Final_List
def Extract_Function_Definition_Location(file_name, tu):
Name_Space="Anonymous NameSpace"
File_to_Array(file_name)
filename = tu.cursor.spelling
for c in tu.cursor.walk_preorder():
if c.location.file is None:
pass
elif c.location.file.name != filename:
pass
elif c.kind == clang.cindex.CursorKind.NAMESPACE:
Name_Space=c.spelling
elif c.kind == clang.cindex.CursorKind.FUNCTION_DECL or c.kind==clang.cindex.CursorKind.CXX_METHOD or c.kind==clang.cindex.CursorKind.FUNCTION_TEMPLATE:
col=[]
col.append(file_name)
col.append(c.spelling)
col.append(Extract_Function_Qualified_Name(c))
col.append(c.mangled_name)
col.append(Name_Space)
lin1, col1 = Extract_Line_Column(c)
col1=col1[:-1]
st=File_Content_Array[int(lin1)-1]
col.append(st.strip())
col.append(lin1)
col.append(col1)
List_of_Functions.append(col)
Function_List.append(c.spelling)
Function_Complexity.append(0)
Extract_Function_Complexity(tu)
Final_List = Merge_Function_Complexity(List_of_Functions, Function_Complexity)
return Final_List
idx = clang.cindex.Index.create()
os.chdir("/Users/shussain/code-analysis/dynamic/tests")
tu = idx.parse("example2.c", args='-xc++ --std=c++11'.split())
List_of_Function_Complexity = Extract_Function_Definition_Location("example2.c", tu)
for f in List_of_Function_Complexity:
print (f)
# -
# ### Script-5 (Called_Calling_Function.py) : Retreive the information about called and calling points of all functions of example2.c
# +
import csv, os, glob
import sys
import clang.cindex
from clang.cindex import Config
os.environ['DYLD_LIBRARY_PATH']= '/usr/local/Cellar/llvm/11.0.0/lib/'
File_Content_Array=[]
Function_Complexity=[]
List_of_Functions=[]
Function_List=[]
with open(file_name) as file:
for line in file:
File_Content_Array.append(line)
#return File_Content_Array
def Extract_Line_Column(cursor):
x=str(cursor.location)
y=x.split(',')
line = y[1]
column=y[2]
line = line.split()
line=line[1]
column=column.split()
column = column[1]
return line, column
def Extract_Function_Qualified_Name(cursor):
if cursor is None:
return ''
elif cursor.kind== clang.cindex.CursorKind.TRANSLATION_UNIT:
return ''
else:
res = Extract_Function_Qualified_Name(cursor.semantic_parent)
if res != '':
return res + '::' + cursor.spelling
return cursor.spelling
def Extract_Function_List(tu):
filename = tu.cursor.spelling
for c in tu.cursor.walk_preorder():
if c.location.file is None:
pass
elif c.location.file.name != filename:
pass
elif c.kind == clang.cindex.CursorKind.FUNCTION_DECL or c.kind==clang.cindex.CursorKind.CXX_METHOD or c.kind==clang.cindex.CursorKind.FUNCTION_TEMPLATE:
Function_List.append(c.spelling)
def Extract_Function_Definition_Calls(file_name, tu):
Extract_Function_List(tu)
Final_List=[]
col=[]
col.append("Calling Function Name")
col.append("Function Definition")
col.append("Line#")
col.append("Column#")
col.append("Name Space")
col.append("Called Function")
col.append("Called Function Definition")
col.append("Line#")
col.append("Column#")
Final_List.append(col)
Name_Space="Anonymous NameSpace"
File_to_Array(file_name)
filename = tu.cursor.spelling
for c in tu.cursor.walk_preorder():
if c.location.file is None:
pass
elif c.location.file.name != filename:
pass
elif c.kind == clang.cindex.CursorKind.NAMESPACE:
Name_Space=c.spelling
elif c.kind == clang.cindex.CursorKind.FUNCTION_DECL or c.kind==clang.cindex.CursorKind.CXX_METHOD or c.kind==clang.cindex.CursorKind.FUNCTION_TEMPLATE:
Calling_Func= c.spelling
lin1, col1 = Extract_Line_Column(c)
st1=File_Content_Array[int(lin1)-1]
col1=col1[:-1]
elif (c.kind == clang.cindex.CursorKind.DECL_REF_EXPR or c.kind == clang.cindex.CursorKind.CALL_EXPR or c.kind == clang.cindex.CursorKind.OVERLOADED_DECL_REF or c.kind==clang.cindex.TokenKind.IDENTIFIER) and c.spelling in Function_List:
Called_Func= c.spelling
lin2, col2 = Extract_Line_Column(c)
st2=File_Content_Array[int(lin2)-1]
col2=col2[:-1]
col=[]
col.append(Calling_Func)
col.append(Name_Space)
col.append(st1)
col.append(lin1)
col.append(col1)
col.append(Called_Func)
col.append(st2)
col.append(lin2)
col.append(col2)
if col not in Final_List:
Final_List.append(col)
else:
col=[]
return Final_List
idx = clang.cindex.Index.create()
os.chdir("/Users/shussain/code-analysis/dynamic/tests")
tu = idx.parse("example2.c", args='-xc++ --std=c++11'.split())
List_of_Function_Calling = Extract_Function_Definition_Calls("example2.c", tu)
for f in List_of_Function_Calling:
print (f)
# -
# ### Script-6 (Function_Metrics.py) : Retreive the set of metrics related to functions of example2.c
# +
import csv, os, glob
import sys
import clang.cindex
from clang.cindex import Config
os.environ['DYLD_LIBRARY_PATH']= '/usr/local/Cellar/llvm/11.0.0/lib/'
File_Content_Array=[]
Function_Complexity=[]
List_of_Functions=[]
Function_List=[]
Function_Identifiers=[]
Function_Literals=[]
Cursor_Types = {clang.cindex.CursorKind.IF_STMT, clang.cindex.CursorKind.WHILE_STMT, clang.cindex.CursorKind.FOR_STMT,clang.cindex.CursorKind.DEFAULT_STMT,clang.cindex.CursorKind.CASE_STMT, clang.cindex.CursorKind.COMPOUND_STMT}
Keywords= {"if", "case", "default", "for", "while", "else"}
def File_to_Array(file_name):
with open(file_name) as file:
for line in file:
File_Content_Array.append(line)
#return File_Content_Array
def Extract_Identifiers_Literals(tu):
cnt=0
filename = tu.cursor.spelling
for c in tu.cursor.get_tokens():
#filename = tu.cursor.spelling
if c.location.file is None:
pass
elif c.location.file.name != filename:
pass
# elif c.kind==clang.cindex.TokenKind.IDENTIFIER and c.spelling in Function_List:
elif (c.cursor.kind==clang.cindex.CursorKind.FUNCTION_DECL or c.cursor.kind==clang.cindex.CursorKind.FUNCTION_TEMPLATE) and c.spelling in Function_List:
# flag=True
# fn=c.spelling
cnt=cnt+1
elif c.kind==clang.cindex.TokenKind.IDENTIFIER and c.spelling not in Function_List:
temp=Function_Identifiers[cnt-1]
temp=temp+1
Function_Identifiers[cnt-1]=temp
elif c.kind==clang.cindex.TokenKind.LITERAL:
temp=Function_Literals[cnt-1]
temp=temp+1
Function_Literals[cnt-1]=temp
def Extract_Function_Complexity(tu):
cnt=0
flag=False
filename = tu.cursor.spelling
fn=""
for c in tu.cursor.get_tokens():
if c.location.file is None:
pass
elif c.location.file.name != filename:
pass
else:
cond1=c.cursor.kind in Cursor_Types
cond2=c.spelling in Keywords
if (c.cursor.kind==clang.cindex.CursorKind.FUNCTION_DECL or c.cursor.kind==clang.cindex.CursorKind.FUNCTION_TEMPLATE) and c.spelling in Function_List:
flag=True
fn=c.spelling
cnt=cnt+1
if cond1==True and cond2==True and (cnt<=len(Function_Complexity) and Function_List[cnt-1] ==fn):# and len(Function_List)==len(Function_Complexity)):
temp=Function_Complexity[cnt-1]
temp=temp+1
Function_Complexity[cnt-1]=temp
def Extract_Line_Column(cursor):
x=str(cursor.location)
y=x.split(',')
line = y[1]
column=y[2]
line = line.split()
line=line[1]
column=column.split()
column = column[1]
return line, column
def Extract_Function_Qualified_Name(cursor):
if cursor is None:
return ''
elif cursor.kind== clang.cindex.CursorKind.TRANSLATION_UNIT:
return ''
else:
res = Extract_Function_Qualified_Name(cursor.semantic_parent)
if res != '':
return res + '::' + cursor.spelling
return cursor.spelling
def Merge_Function_Complexity(Functions, Complexity, Identifiers, literals):
Final_List=[]
# Heading row for the list of functions
col=[]
col.append("File Name")
col.append("Function Name")
col.append("Function Qualified Name")
col.append("Mangled Name")
col.append("Name Space")
col.append("Function Definition")
col.append("Line Number")
col.append("Column Number")
col.append("Cyclomatic Complexity")
col.append("Total_Identifiers")
col.append("Total_Literals")
Final_List.append(col)
cnt=0
for func in Functions:
col=[]
col.append(func[0])
col.append(func[1])
col.append(func[2])
col.append(func[3])
col.append(func[4])
col.append(func[5])
col.append(func[6])
col.append(func[7])
col.append(Complexity[cnt])
col.append(Identifiers[cnt])
col.append(literals[cnt])
Final_List.append(col)
cnt=cnt+1
return Final_List
def Extract_Function_Definition_Location(file_name, tu):
Name_Space="Anonymous NameSpace"
File_to_Array(file_name)
filename = tu.cursor.spelling
for c in tu.cursor.walk_preorder():
if c.location.file is None:
pass
elif c.location.file.name != filename:
pass
elif c.kind == clang.cindex.CursorKind.NAMESPACE:
Name_Space=c.spelling
elif c.kind == clang.cindex.CursorKind.FUNCTION_DECL or c.kind==clang.cindex.CursorKind.CXX_METHOD or c.kind==clang.cindex.CursorKind.FUNCTION_TEMPLATE:
col=[]
col.append(file_name)
col.append(c.spelling)
col.append(Extract_Function_Qualified_Name(c))
col.append(c.mangled_name)
col.append(Name_Space)
lin1, col1 = Extract_Line_Column(c)
col1=col1[:-1]
st=File_Content_Array[int(lin1)-1]
col.append(st.strip())
col.append(lin1)
col.append(col1)
List_of_Functions.append(col)
Function_List.append(c.spelling)
Function_Complexity.append(0)
Function_Identifiers.append(0)
Function_Literals.append(0)
Extract_Function_Complexity(tu)
Extract_Identifiers_Literals(tu)
Final_List = Merge_Function_Complexity(List_of_Functions, Function_Complexity, Function_Identifiers, Function_Literals)
return Final_List
idx = clang.cindex.Index.create()
os.chdir("/Users/shussain/code-analysis/dynamic/tests")
tu = idx.parse("example2.c", args='-xc++ --std=c++11'.split())
List_of_Function_Metrics = Extract_Function_Definition_Location("example2.c", tu)
for f in List_of_Function_Metrics:
print (f)
# -
|
sandbox/dynamic/Dynamic_analysis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Learning Objectives:
#
#
# 1. Selection via indexing
# 2. Row indexing
# 3. Column Selection
# 4. .loc[ ] method in dataframes
# 5. Slicing in loc method
# 6. iloc[ ] method for indexing
import numpy as np
import pandas as pd
# +
##creating a dataframe using dictionary
scores_data = {"Student": ["Tom", "Katey", "Mak", "Bill", "Rahul"],
"Age": np.random.randint(20, 25, size=5),
"Sub1_Score": np.random.randint(40, 100, size=5),
"Sub2_Score": np.random.randint(40, 100, size=5),
"Sub3_Score": np.random.randint(40, 100, size=5)
}
scores_df = pd.DataFrame(scores_data)
scores_df
# -
scores_df[2]
scores_df['Student']
scores_df[:]
scores_df[1:4]
scores_df[2:3]
# ## Column selection
scores_df[['Sub1_Score', 'Sub2_Score']]
# ## `.loc[ ]` method in dataframes
scores_df = scores_df.set_index('Student')
scores_df
scores_df.loc['Tom']
scores_df.loc[['Tom', 'Katey']]
scores_df.loc[['Tom', 'Katey'], ['Sub1_Score', 'Sub3_Score']]
scores_df.loc['Katey': 'Bill']
# ## `iloc[ ]` method in dataframe
scores_df = scores_df.reset_index(drop=True)
scores_df
scores_df[1:4:2]
scores_df[::-1]
scores_df.iloc[2]
scores_df.iloc[1:3]
scores_df.iloc[:, 1:]
scores_df.iloc[::2, 2:]
|
module_4_pandas/indexing_&_subsetting_dataframe.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
from scipy import stats, optimize, interpolate
from sklearn.linear_model import LinearRegression
import netCDF4 # module that reads in .nc files (built on top of HDF5 format)
import pandas as pd
import geopandas as gpd
from geopandas.tools import sjoin
import xarray
import rioxarray
from shapely.geometry import Point, mapping
from shapely.geometry.polygon import Polygon
from pyproj import CRS, Transformer # for transforming projected coordinates to elliptical coordinates
import cartopy.crs as ccrs # for defining and transforming coordinate systems
import cartopy.feature as cfeature # to add features to a cartopy map
import cartopy.io.shapereader as shpreader
from fire_utils import ncdump, coord_transform, tindx_func, bailey_ecoprovince_shp, bailey_ecoprovince_mask, update_reg_indx, \
mon_fire_freq, mon_burned_area, seas_burnarea, clim_pred_var
from stats_utils import uni_lsq_regression_model, reg_uni_climate_fire_corr, multi_regression_model, reg_multi_climate_fire_corr
#self-library
from datetime import datetime, timedelta
from cftime import num2date, date2num, DatetimeGregorian
from tqdm import tqdm
import matplotlib.pyplot as plt
import matplotlib.colors as colors
from matplotlib.patches import Rectangle
import matplotlib.patches as patches
import matplotlib.path as mpltPath
# %matplotlib inline
# %config IPython.matplotlib.backend = 'retina'
# %config InlineBackend.figure_format = 'retina'
# -
data_dir= "../data"
pred_input_path= "/12km/"
resp_input_path= "/firelist/"
outfilepath= "../plots/"
# ## Data pre-processing
# +
wildfire_df= pd.read_csv(data_dir + resp_input_path + "west_US_fires_1984_2020.txt",
usecols= (0, 1, 9, 18, 19, 20, 21, 22, 23, 24), delimiter= ',') #west_US_fires_1984_2020.txt
wildfire_x, wildfire_y= coord_transform(wildfire_df['final_lat'], wildfire_df['final_lon'])
wildfire_df['final_x']= wildfire_x
wildfire_df['final_y']= wildfire_y
wildfire_gdf= gpd.GeoDataFrame(wildfire_df, crs= 'EPSG:5070', geometry=gpd.points_from_xy(wildfire_df['final_x'], wildfire_df['final_y']))
reg_indx_arr= update_reg_indx(wildfire_gdf) #sorts all fires into the respective regions using polygon matching
wildfire_df['reg_indx']= reg_indx_arr #adding regional index as a column in the dataframe
grouped= wildfire_df.groupby(['reg_indx'])
# -
fire_file= data_dir + pred_input_path + "wildfire/burnarea_combined.nc"
burnarea_data= netCDF4.Dataset(fire_file, 'r')
lat_long_fire_grid= coord_transform(burnarea_data['X'][:].data, burnarea_data['Y'][:].data, input_crs= 'EPSG:5070')
tot_months= 36*12
month_arr= np.linspace(0, tot_months - 1, tot_months, dtype= int)
year_arr= np.asarray([1984, 1991, 1998, 2005, 2012, 2019])
# ## Statistical analyses
# +
freq_sierra= mon_fire_freq(wildfiredf= wildfire_df, regindx= 1, threshold= True).flatten()
freq_imdesert= mon_fire_freq(wildfiredf= wildfire_df, regindx= 13, threshold= True).flatten()
sum_indx_1, sum_indx_2= tindx_func(startmon= 3, duration= 8, tim_size= 432)
sum_freq_sierra= np.asarray([np.sum(freq_sierra[sum_indx_1[i]:sum_indx_2[i]]) for i in range(len(sum_indx_1))])
sum_freq_imdesert= np.asarray([np.sum(freq_imdesert[sum_indx_1[i]:sum_indx_2[i]]) for i in range(len(sum_indx_1))])
# -
pred_var_sierra, pred_freq_sierra, r_sierra= uni_lsq_regression_model(sum_freq_sierra, pred_file_indx= 2, pred_seas_indx= 1, regindx= 1, freq_flag= True)
pred_var_imdesert, pred_freq_imdesert, r_imdesert= uni_lsq_regression_model(sum_freq_imdesert, pred_file_indx= 2, pred_seas_indx= 1, regindx= 13, freq_flag= True)
sierra_sum_burnarea= seas_burnarea(firefile= fire_file, season= "summer", regindx= 1)
imdesert_sum_burnarea= seas_burnarea(firefile= fire_file, season= "summer", regindx= 13)
coeff_sierra, r2_sierra, _ = multi_regression_model(sierra_sum_burnarea, regression= "enetCV", regindx= 1, freq_flag= False)
coeff_imdesert, r2_imdesert, _ = multi_regression_model(imdesert_sum_burnarea, regression= "enetCV", regindx= 13, freq_flag= False)
# ## Plotting
# +
fig2= plt.figure(figsize=(20, 20))
gs = fig2.add_gridspec(4, 4)
fig2.subplots_adjust(hspace= 0.4, wspace= 0.2)
pred_var_arr= ["Tmax", "VPD", "Prec", "Antprc", "PET", "Forest"]
ypos= np.arange(len(pred_var_arr))
f2_ax1 = fig2.add_subplot(gs[0, 0:2])
f2_ax1.set_title(r'Sierra Nevada', fontsize= 14);
ax2= f2_ax1.twinx()
f2_ax1.plot(month_arr, mon_fire_freq(wildfiredf= wildfire_df, regindx= 1).flatten(), color= 'turquoise', label= 'Large (> 405 ha) fire frequency');
ax2.plot(month_arr, mon_burned_area(firefile= fire_file, regindx= 1, final_year= 2019), color= 'forestgreen', label= 'Summer burned area');
f2_ax1.set_xticks((year_arr - 1984 + 1)*12 - 1);
f2_ax1.set_xticklabels(year_arr)
f2_ax1.set_ylim(0, 100);
f2_ax1.set_ylabel(r'Frequency', fontsize= 12);
ax2.set_ylim(0, 4500);
#ax2.set_ylabel(r'Burned area [in ${\rm km}^2$]', fontsize= 12, labelpad= 10, rotation= 270);
f2_ax1.tick_params(labeltop=False, top=True, labelright=False, right=False, which='both', labelsize= 12);
ax2.tick_params(labeltop=False, top=True, labelright=False, right=True, which='both', labelsize= 12);
f2_ax1.grid(b=True, which='major', color='black', alpha=0.05, linestyle='-');
f2_ax1.grid(b=True, which='minor', color='black', alpha=0.01, linestyle='-');
f2_ax1.legend(loc= (0.28, 0.90), frameon=False, fontsize= 12);
ax2.legend(loc= (0.28, 0.82), frameon=False, fontsize= 12);
f2_ax2 = fig2.add_subplot(gs[0, 2:4])
f2_ax2.set_title(r'IM Desert', fontsize= 14);
ax3= f2_ax2.twinx()
f2_ax2.plot(month_arr, mon_fire_freq(wildfiredf= wildfire_df, regindx= 13).flatten(), color= 'coral', label= 'Large (> 405 ha) fire frequency');
ax3.plot(month_arr, mon_burned_area(firefile= fire_file, regindx= 13, final_year= 2019), color= 'gold', label= 'Summer burned area');
f2_ax2.set_xticks((year_arr - 1984 + 1)*12 - 1);
f2_ax2.set_xticklabels(year_arr)
f2_ax2.set_ylim(0, 100);
#f2_ax2.set_ylabel(r'Frequency', fontsize= 12);
ax3.set_ylim(0, 4500);
ax3.set_ylabel(r'Burned area [${\rm km}^2$]', fontsize= 12, labelpad= 15, rotation= 270);
f2_ax2.tick_params(labeltop=False, top=True, labelleft= False, labelright=False, right=False, which='both', labelsize= 12);
ax3.tick_params(labeltop=False, top=True, labelright=True, right=True, which='both', labelsize= 12);
f2_ax2.grid(b=True, which='major', color='black', alpha=0.05, linestyle='-');
f2_ax2.grid(b=True, which='minor', color='black', alpha=0.01, linestyle='-');
f2_ax2.legend(loc= (0.45, 0.90), frameon=False, fontsize= 12);
ax3.legend(loc= (0.45, 0.82), frameon=False, fontsize= 12);
f2_ax3 = fig2.add_subplot(gs[1, 0])
f2_ax3.plot(pred_var_sierra, sum_freq_sierra, 'o', markersize= 10,
markerfacecolor= 'turquoise',
markeredgecolor= 'turquoise',
linestyle= 'None');
f2_ax3.plot(pred_var_sierra, pred_freq_sierra, color= 'black', lw= 2, label=r'$r = %.2f$'%np.sqrt(r_sierra));
f2_ax3.set_xlabel(r"Mar-Oct VPD $\ [{\rm hPa}]$", fontsize= 12);
f2_ax3.set_title(r'Frequency', fontsize= 12)
f2_ax3.legend(loc='best', frameon=True, fontsize=12);
f2_ax3.tick_params(labeltop=False, top=True, labelright=False, right=False, which='both', labelsize= 12);
f2_ax3.grid(b=True, which='major', color='black', alpha=0.05, linestyle='-');
f2_ax3.grid(b=True, which='minor', color='black', alpha=0.01, linestyle='-');
f2_ax4 = fig2.add_subplot(gs[1, 1])
f2_ax4.barh(ypos, coeff_sierra, align= "center", color= 'forestgreen');
f2_ax4.set_xlim(-1.2, 1.2);
f2_ax4.set_xlabel(r"Normalized coefficients", fontsize= 12);
f2_ax4.set_yticks(ypos);
f2_ax4.set_yticklabels(pred_var_arr, fontsize= 12);
f2_ax4.tick_params(labeltop=False, top=True, labelright=False, right=False, which='both', labelsize= 12);
f2_ax4.grid(b=True, which='major', color='black', alpha=0.05, linestyle='-');
f2_ax4.grid(b=True, which='minor', color='black', alpha=0.01, linestyle='-');
f2_ax4.text(0.55, 4.7, r"${\rm R}^2 = %.2f$"%r2_sierra, fontsize= 12, bbox=dict(facecolor='none', edgecolor='grey', boxstyle='round', pad=0.3));
f2_ax4.set_title(r'Burned Area', fontsize= 12);
f2_ax5 = fig2.add_subplot(gs[1, 2])
f2_ax5.plot(pred_var_imdesert, sum_freq_imdesert, 'o', markersize= 10,
markerfacecolor= 'coral',
markeredgecolor= 'coral',
linestyle= 'None');
f2_ax5.plot(pred_var_imdesert, pred_freq_imdesert, color= 'black', lw= 2, label=r'$r = %.2f$'%np.sqrt(r_imdesert));
f2_ax5.set_xlabel(r"Mar-Oct VPD $\ [{\rm hPa}]$", fontsize= 12);
f2_ax5.set_title(r'Frequency', fontsize= 12)
f2_ax5.legend(loc='best', frameon=True, fontsize=12);
f2_ax5.tick_params(labeltop=False, top=True, labelright=False, right=False, which='both', labelsize= 12);
f2_ax5.grid(b=True, which='major', color='black', alpha=0.05, linestyle='-');
f2_ax5.grid(b=True, which='minor', color='black', alpha=0.01, linestyle='-');
f2_ax6= fig2.add_subplot(gs[1, 3])
f2_ax6.barh(ypos, coeff_imdesert, align= "center", color= 'gold');
f2_ax6.set_xlim(-1.2, 1.2);
f2_ax6.set_xlabel(r"Normalized coefficients", fontsize= 12);
f2_ax6.set_yticks(ypos);
f2_ax6.set_yticklabels(pred_var_arr, fontsize= 12);
f2_ax6.tick_params(labeltop=False, top=True, labelright=False, right=False, which='both', labelsize= 12);
f2_ax6.grid(b=True, which='major', color='black', alpha=0.05, linestyle='-');
f2_ax6.grid(b=True, which='minor', color='black', alpha=0.01, linestyle='-');
f2_ax6.text(0.55, 4.7, r"${\rm R}^2 = %.2f$"%r2_imdesert, fontsize= 12, bbox=dict(facecolor='none', edgecolor='grey', boxstyle='round', pad=0.3));
f2_ax6.set_title(r'Burned Area', fontsize= 12);
#plt.savefig(outfilepath + 'clim_fire_freq_area.pdf', bbox_inches='tight');
# -
|
notebooks/.ipynb_checkpoints/grant_plot_nb-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # **Traffic Sign Recognition**
#
# ## Writeup
#
#
#
# ---
#
# **Build a Traffic Sign Recognition Project**
#
# The goals / steps of this project are the following:
# * Load the data set (see below for links to the project data set)
# * Explore, summarize and visualize the data set
# * Design, train and test a model architecture
# * Use the model to make predictions on new images
# * Analyze the softmax probabilities of the new images
# * Summarize the results with a written report
#
#
# [//]: # (Image References)
#
# [image1]: ./output_figures/train_set_class.png
# [image2]: ./output_figures/test_set_class.png
# [image3]: ./output_figures/validation_set_class.png
# [image4]: ./output_figures/before_process.png
# [image5]: ./output_figures/after_process.png
# [image6]: ./output_figures/learning_curve.png
# [image7]: ./germain_traffic_signs/online_images.png
# [image8]: ./output_figures/online_test.png
#
#
#
# ## Rubric Points
# ### Here I will consider the [rubric points](https://review.udacity.com/#!/rubrics/481/view) individually and describe how I addressed each point in my implementation.
#
# ---
#
#
# You're reading it! and here is a link to my [project code](https://github.com/zhou-wenbin/UdacityND-Traffic-Sign-Classifier-Project/blob/master/Traffic%20Sign%20Classifier%20for%20submission.ipynb)
#
# ### Data Set Summary & Exploration
#
# #### 2. A basic summary of the data set.
#
#
# | Data description | number |
# |:---------------------:|:---------------------------------------------:|
# | size of training set | 34799 |
# | size of the validation set | 4410 |
# | size of test set | 12630 |
# | shape of a traffic sign image | (32, 32, 3) |
# | number of unique labels | 43 |
#
#
# #### 1. The distribution of labels in train/test/validation data set.
#
# train data | test data | validation data
# :-------------------------:|:-------------------------:|:-------------------------:
# ![alt text][image1] | ![alt text][image2]| ![alt text][image3]
#
#
#
#
# ### Design and Test a Model Architecture
#
# I have tried different ways to process the images, for example, equalize the histogram of the Y channel of images, add weight to the gussian blurred version of the image and so on. But that did not increased the accuracy. For the record, I will post how I did the image process even though in the training process I did not use them.
#
# * After I checked the training set, I saw that there are some dark images in the set, as follows,
# ![alt text][image4]
#
# * After I preprocess the training set, I made the signs more standing out, as follows,
# ![alt text][image5]
#
#
# Before I train my model, the only thing I did to my data set is to shuffle the training data because after shuffling the data we cag get rid of the correlation between images.
#
#
# #### 1. My final model consisted of the following layers:
#
# | Layer | Description |
# |:---------------------:|:---------------------------------------------:|
# | Input | 32x32x3 RGB image |
# | Layer 1:conv1 | filter=5x5x6, strides=[1,1,1,1], outputs=28x28x6 |
# | ReLU | activation function |
# | Layer 2 :conv2 | filter=5x5x10, strides=[1,1,1,1], outputs=24x24x10 |
# | ReLU | activation function |
# |Layer 3: conv3 | filter=5x5x16, strides=[1,1,1,1], outputs=20x20x16 |
# | ReLU | activation function |
# | Max pooling | strides=[1,2,2,1], Input = 20x20x16, Output = 10x10x16. |
# | Layer 4:fully connected | Input = 10x10x16, Output =120 |
# | Dropout | |
# | Layer 5:fully connected | Input = 120, Output =100 |
# | ReLU | activation function |
# | Layer 6:fully connected | Input = 100, Output =84 |
# | ReLU | activation function |
# | Layer 7:fully connected | Input = 84, Output =43 |
#
#
#
# #### 2. Hyper parameters
#
#
# |Hyper parameters | Description |
# |:---------------------:|:---------------------------------------------:|
# | optimizer | Adam |
# | batch size | 128 |
# | number of epochs | 50 |
# |learning rate| 0.001 |
#
# #### 3. My approach to get the above model.
#
# In the beginning, I implemeted the simple LeNet network with only 5 layers that was explained in the lecture and no matter how I tried to process the image I was not able to achieve test accuracy above 0.9. Then I started to increased the layer a bit, because I think the deeper the network, the better the performance will be. In order to prevent overfitting I also added a dropout process in between layer 4 and layer 5 because layer 4 has many nurons so it might cause overfitting. Surprisingly, after I increased 2 more layers the accuracy increased immediately. However, it will not increase the performance even if I did a preprocess of training set. I will leave this as following research to see why the preprocess of the image did not help.
#
# ### 4. My final model results were:
#
# * training set accuracy of 0.998
# * validation set accuracy of 0.958
# * test set accuracy of 0.950
#
# #### The learning curve is shown as follows,
# ![alt text][image6]
#
#
# ### 5. Test a Model on New Images
#
# * The online images are chosen as follows :
#
# ![alt text][image7]
#
# I manully screenshot traffic signs from google images.
#
# * Test result is (with top five guessing),
# ![alt text][image8]
# with test accuray =0.800
# ### 6. Reflections
# * w.r.t the testing on online images:
#
# The only image that was not recongnized is due to the reason of resize, which makes the sign diffictult to recongnize either for out human eyes. You can that the input image of the unrecongnized one is very blur.
#
# * w.r.t the model constructions:
#
# I experimented that if I increased the NN layer from 5 to 7, the accuracy increased surprisely. However, there is no theory that can explain this. And also those parameters are tuned by trials and errors and I had no clue why should we decrease the learning rate or increase the EPOCH size or why should we do the image process. These will be left as future reseach.
|
writeup report.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.5 64-bit (''voicenet_venv'': venv)'
# language: python
# name: python37564bitvoicenetvenvvenv3211909c721248e184b0b91a67b5318b
# ---
# # Gender Recognition Based on Voice msms
# By <NAME>
# ---
# A lot can be achieved by analyzing Voice in Speech Analytics. And one of the most foundational tasks can be: Identifying the Gender with the help of Voice. In this project, I'll analyze and cover the workflow of how to detect the gender of the speaker using **MFCC** (Mel Frequency and Cepstral Coefficients) and **GMM** (Gaussian Mixture Models). I'll make use of the mentioned techniques to achieve noteworthy performance.
# ---
# # Outline
# 1. Introduction to Project
# - 1.1 Project Objective
# - 1.2 Historical Context
# - 1.3 Project Workflow
#
#
# 2. Project Setup
# - 2.1 Importing the Libraries
# - 2.2 Importing the Data
# - 2.3 Managing the Data
#
#
# 3. What is MFCC?
# - 3.1 Building the Features Extractor
#
#
# 4. What are GMMs?
# - 3.2 Training the Models
#
#
# 5. Identifying the Gender
#
#
# 6. Conclusions and Analysis
#
#
# 7. Acknowledgements and References
# ---
# # 1. Introduction of Project
# ## 1.1 Project Objective
# To predict the gender of the speaker based on his/her voice samples.
# ---
# ## 1.2 Historical Context
# Large amounts of computing power available alongwith Artificial Intelligent systems has resulted inflection into capability of machines to recognize the voices. Faster Processing and large amount of Speech Data available makes the performance of these sytems roughly on par with humans. From **Audrey** a speech recognizing system which could recognize a single voice speaking digits aloud at Bell Labs in 1952; we've reached to having day to day conversations with voice assistants like Google Assistant and Siri in our smartphones.
#
# But most of these systems are usually neutral to the gender of the speaker and results being given. Having systems which can respond as per the user's gender is indeed an amazing capability. A large amount of tasks which are based on gender preferences can be handled by them. It results into better customer service and enhances user experience.
# 
# ---
# ## 1.3 Project Workflow
# 
# ---
# # 2. Project Setup
# ## 2.1 Importing the Libraries
# Here I'm using one of the standard machine learning libraries which are available in scikit-learn along with numpy and pandas for data manipulation.
# +
# Importing Libraries and Modules
# For Importing Files
import os
import sys
import math
import tarfile
# For Data Manipulation
import numpy as np
import pandas as pd
# For Audio Files Processing
from scipy.io.wavfile import read
from sklearn.mixture import GaussianMixture as GMM
from python_speech_features import mfcc
from python_speech_features import delta
from sklearn import preprocessing
# To Ignore Warnings
import warnings
warnings.filterwarnings('ignore')
# To Save Models
import pickle
# -
# ---
# ## 2.2 Importing the Data
# Data about voice samples of males and females is **The Free ST American English Corpus dataset** which can be downloaded from [here](http://www.openslr.org/45)!. It contains utterances from 10 speakers, 5 from each gender. Each speaker has about 350 utterances.
# Once you download your dataset, you need to split it into two parts:Training Set and Testing Set.
#
# - **Training Set** : It's used to train the gender models.
#
#
# - **Testing Set** : It's used for testing the accuracy of the gender recognition.
#
# The spilliting criterion depends totally on you. I'll prefer going with 2/3 for Training Set and rest for Testing Set. I'll create a class which will help us managing and formatting our data. We may need functions for following tasks:
#
# 1. A function for getting the path where our compressed dataset resides.
#
#
# 2. A function to extract files out of our compressed dataset.
#
#
# 3. A function to create separate folders for our training and testing files.
#
#
# 4. A function which can fill filenames into an empty dictionary.
#
#
# 5. A function which can move files into their respective folders.
#
#
# 6. And ofcourse a driver function for all of the above functions.
#
# ---
# ## 2.3 Managing the Data
# +
class Data_Manager:
# Function #1
def __init__(self, dataset_path):
self.dataset_path = dataset_path
#-------------------------------------------------------------------------------------------------------------------------------
# Function #2
def extract_dataset(self, compressed_dataset_file_name, dataset_directory):
try:
tar = tarfile.open(compressed_dataset_file_name, "r:gz")
tar.extractall(dataset_directory)
tar.close()
print("Files extraction was successful!")
except:
print("No extraction was performed !")
#-------------------------------------------------------------------------------------------------------------------------------
# Function #3
def make_folder(self, folder_path):
try:
os.mkdir(folder_path)
print(folder_path, "was created !")
except:
print("Exception raised: ", folder_path, "could not be created !")
#-------------------------------------------------------------------------------------------------------------------------------
# Function #4
def get_fnames_from_dict(self, dataset_dict, f_or_m):
training_data, testing_data = [], []
for i in range(1,5):
length_data = len(dataset_dict[f_or_m +"000" + str(i)])
length_separator = math.trunc(length_data*2/3)
training_data += dataset_dict[f_or_m + "000" + str(i)][:length_separator]
testing_data += dataset_dict[f_or_m + "000" + str(i)][length_separator:]
return training_data, testing_data
#------------------------------------------------------------------------------------------------------------------------------
# Function #5
def move_files(self, src, dst, group):
for fname in group:
os.rename(src + '/' + fname, dst + '/' + fname)
#------------------------------------------------------------------------------------------------------------------------------
# Function #6
def manage(self):
compressed_dataset_file_name = self.dataset_path
dataset_directory = compressed_dataset_file_name.split(".")[0]
try:
os.mkdir(dataset_directory)
except:
pass
self.extract_dataset(compressed_dataset_file_name, dataset_directory)
file_names = [fname for fname in os.listdir(dataset_directory) if ("f0" in fname or "m0" in fname)]
dataset_dict = {"f0001": [], "f0002": [], "f0003": [], "f0004": [], "f0005": [],
"m0001": [], "m0002": [], "m0003": [], "m0004": [], "m0005": [], }
for fname in file_names:
dataset_dict[fname.split('_')[0]].append(fname)
training_set, testing_set = {},{}
training_set["females"], testing_set["females"] = self.get_fnames_from_dict(dataset_dict, "f")
training_set["males" ], testing_set["males" ] = self.get_fnames_from_dict(dataset_dict, "m")
self.make_folder("TrainingData")
self.make_folder("TestingData")
self.make_folder("TrainingData/females")
self.make_folder("TrainingData/males")
self.make_folder("TestingData/females")
self.make_folder("TestingData/males")
self.move_files(dataset_directory, "TrainingData/females", training_set["females"])
self.move_files(dataset_directory, "TrainingData/males", training_set["males"])
self.move_files(dataset_directory, "TestingData/females", testing_set["females"])
self.move_files(dataset_directory, "TestingData/males", testing_set["males"])
#-------------------------------------------------------------------------------------------------------------------------------
if __name__== "__main__":
data_manager = Data_Manager("SLR45.tgz")
data_manager.manage()
# -
# Let me expain briefly what I've done here:
#
# 1. **Function #1** : It gets path where our dataset resides!
#
#
# 2. **Function #2** : It extracts tar format file to a directory.
#
#
# 3. **Function #3** : It creates a folder for the Data.
#
#
# 4. **Function #4** : Create dictionaries from Training Set and Testing Set.
#
#
# 5. **Function #5** : Move files to their respective folders.
#
#
# 6. **Function #6** : It reads file & creates folder for the data where it'll decompress our dataset. Later it'll select our files and fill them in our dictionary, divide and group our file names. And finally when we're done creating folders for our files, it'll move them into their respective folders.
# ---
# # 3. What is MFCC?
# It's time to build a feature extractor now. There can be many acoustic features which can help distinguishing males from females, but I'm here gonna use **MFCC** or **Mel Frequency Cepstral Co-efficients**, since it's one of the best acoustic feature in terms of results. Generally here's how they're derived:
#
# 1. Take the Fourier transform of (a windowed excerpt of) a signal. It transforms the time domain signal into spectral domain signal where source and filter part are now in multiplication.
#
#
# 2. Map the powers of the spectrum obtained above onto the mel scale, using triangular overlapping windows.
#
#
# 3. Take the logs of the powers at each of the mel frequencies. It helps in separating source and filter.
#
#
# 4. Take the discrete cosine transform of the list of mel log powers, as if it were a signal.
#
#
# 5. The MFCCs are the amplitudes of the resulting spectrum.
# 
# ---
# ## 3.1 Building Features Extractor
# To extract MFCC features, I'm gonna make use of a python module named:python_speech_features. It's simple to apply, and has a good documentation for support.
#
# It's best suitable to build a class and encaspulate a function which does features extraction for us:
class Features_Extractor:
def __init__(self):
pass
def extract_features(self, audio_path):
rate, audio = read(audio_path)
mfcc_feature = mfcc(audio, rate, winlen = 0.05, winstep = 0.01, numcep = 5, nfilt = 30, nfft = 800, appendEnergy = True)
mfcc_feature = preprocessing.scale(mfcc_feature)
deltas = delta(mfcc_feature, 2)
double_deltas = delta(deltas, 2)
combined = np.hstack((mfcc_feature, deltas, double_deltas))
return combined
# Let's see what I've just done here! I've built a function which extracts MFCC from audio files and performs the CMS normalization and later I've combined it with MFCC deltas and double_deltas. It takes audio_path i.e path to the audio wave and returns an array or extracted features matrix.
#
# MFCC function has several arguments, they signify:
#
# - **audio**: Audio signal from which we've to compute features
#
#
# - **rate** : Sample rate of the audio signal we're working with
#
#
# - **winlen**: Length of the analysis window in seconds; default is 0.025s (25 milliseconds)
#
#
# - **winstep**: Default step between successive windows in seconds; default is 0.01s (10 milliseconds)
#
#
# - **numcep**: Number of Cepstrum to return; default is 13
#
#
# - **nfilt**: Number of filters in the filterbank; default is 26
#
#
# - **nfft**: Size of the fft; default is 512
#
#
# - **appendEnergy**: If it's set True, the zeroth cepstral coefficient is replaced with log of total frame energy
# ---
# # 4. What are GMMs?
# > A Gaussian Mixture Model (GMM) is a parametric probability density function represented as a weighted sum of Gaussian component densities. GMMs are commonly used as a parametric model of the probability distribution of continuous measurements or features in a biometric system, such as vocal-tract related spectral features in a speaker recognition system. GMM parameters are estimated from training data using the iterative Expectation-Maximization (EM) algorithm or Maximum A Posteriori(MAP) estimation from a well-trained prior model.
# >
# > [<NAME>](https://www.semanticscholar.org/paper/Gaussian-Mixture-Models-Reynolds/734b07b53c23f74a3b004d7fe341ae4fce462fc6)
# 
# A Gaussian Mixture Model popularly known as GMM is a probabilistic clustering model for reprenting a certain data distribution as a sum of Gaussian Density Functions. These densities forming a GMM are also known as components of GMM. The likelihood of a data point is given by the following equation:
#
#
# $P(X|\lambda) = \sum_{k=1}^{K} w_k P_k(X|\mu_k, \Sigma_k) $
#
#
#
#
# where $P_k(X|\mu_k, \Sigma_k) $ is the Gaussian Distribution:
#
#
#
#
# $P_k(X|\mu_k,\Sigma_k) = \frac{1}{{\sqrt{2\pi|\Sigma_k|}}} \thinspace e^{\frac{1}{2}(X-\mu_k)^T \Sigma^{-1}(X-\mu_k)}$
#
# where:
#
# $\lambda$ : It represents Training Data.
#
# $\mu$ : It represents the mean.
#
# $\Sigma$ : It represents the co-variance matrices.
#
# $w_k$ : It represents the weights.
#
# $k$ : It represents the index of the components.
# ---
# ## 4.1 Training the Models
# I'm going to build a class where I'll train my audio samples. It'll be a tedious task if I'll write it in separate cells, so I'll stick with the same cell. Let's see what I aim to achieve here:
#
# 1. A function which can assign paths to where our voice samples resides
#
#
# 2. A function which collects voice features from the files
#
#
# 3. A function where I will generate GMM Models and later would fit our features
#
#
# 4. A function where I will save our newly constructed GMM Models
# +
class Models_Trainer:
# Function #1
def __init__(self, females_files_path, males_files_path):
self.females_training_path = females_files_path
self.males_training_path = males_files_path
self.features_extractor = Features_Extractor()
#-----------------------------------------------------------------------------------------------------------------------------
# Function #2
def get_file_paths(self, females_training_path, males_training_path):
females = [ os.path.join(females_training_path, f) for f in os.listdir(females_training_path) ]
males = [ os.path.join(males_training_path, f) for f in os.listdir(males_training_path) ]
return females, males
#-----------------------------------------------------------------------------------------------------------------------------
# Function #3
def collect_features(self, files):
features = np.asarray(())
for file in files:
print("%5s %10s" % ("Processing ", file))
vector = self.features_extractor.extract_features(file)
if features.size == 0:
features = vector
else:
features = np.vstack((features, vector))
return features
#------------------------------------------------------------------------------------------------------------------------------
# Function #4
def process(self):
females, males = self.get_file_paths(self.females_training_path,self.males_training_path)
female_voice_features = self.collect_features(females)
male_voice_features = self.collect_features(males)
females_gmm = GMM(n_components = 16, max_iter = 200, covariance_type='diag', n_init = 3)
males_gmm = GMM(n_components = 16, max_iter = 200, covariance_type='diag', n_init = 3)
females_gmm.fit(female_voice_features)
males_gmm.fit(male_voice_features)
self.save_gmm(females_gmm, "females")
self.save_gmm(males_gmm, "males")
#-----------------------------------------------------------------------------------------------------------------------------
# Function #5
def save_gmm(self, gmm, name):
filename = name + ".gmm"
with open(filename, 'wb') as gmm_file:
pickle.dump(gmm, gmm_file)
print ("%5s %10s" % ("Saving", filename,))
#-----------------------------------------------------------------------------------------------------------------------------
if __name__== "__main__":
models_trainer = Models_Trainer("TrainingData/females", "TrainingData/males")
models_trainer.process()
# -
# Okay, I'll explain what I've done here. Lemme go through each function and succinctly tell you what's happening:
#
# 1. **Function #1** : It assigns the paths of the female and male audio samples to their respective variables; signifying that these are training samples.
#
#
# 2. **Function #2** : It gets the file paths and stores them in their respective appropriate variable names.
#
#
# 3. **Function #3** : It collects various features from the people of the same gender. It takes up audio samples, and returns extracted features matrix. It extracts MFCC and delta features and stacks them.
#
#
# 4. **Function #4** : This function gathers features from Function #3, generates GMM Models and later fits features collected to them. There are 2 separate models for males and females. Finally, generated models are saved.
#
#
# 5. **Function #5** : It's always to better to save your models so you don't have to iterate the whole process again.It takes the GMM models and the filename. Pickle Module is used to dump the models just generated.
# ---
# # 5. Identifying the Gender
# Finally, all the pieces are about to chip in the right place. We've already collected features, fitted them to our generated GMM models. It's time to see how it works on samples it hasn't seen yet!
# I'm going to create a class once again, which encapsulates several functions. Let's see what I wish to achieve here:
#
# 1. A function for necessary variables and to load our previously saved GMM models.
#
#
# 2. A function which can return where our voice samples to be tested resides.
#
#
# 3. A function to identify the gender by computing the likelihood of male and female voice samples.
#
#
# 4. A function which can read the samples and can declare the better likelihood out of two and to predict results.
# +
class Gender_Identifier:
# Function #1
def __init__(self, females_files_path, males_files_path, females_model_path, males_model_path):
self.females_training_path = females_files_path
self.males_training_path = males_files_path
self.error = 0
self.total_sample = 0
self.features_extractor = Features_Extractor()
self.females_gmm = pickle.load(open(females_model_path, 'rb'))
self.males_gmm = pickle.load(open(males_model_path, 'rb'))
#------------------------------------------------------------------------------------------------------------------------------
# Function #2
def get_file_paths(self, females_training_path, males_training_path):
females = [ os.path.join(females_training_path, f) for f in os.listdir(females_training_path) ]
males = [ os.path.join(males_training_path, f) for f in os.listdir(males_training_path) ]
files = females + males
return files
#------------------------------------------------------------------------------------------------------------------------------
# Function #3
def identify_gender(self, vector):
female_scores = np.array(self.females_gmm.score(vector))
female_log_likelihood = female_scores.sum()
male_scores = np.array(self.males_gmm.score(vector))
male_log_likelihood = male_scores.sum()
print("%10s %5s %1s" % ("+ Female Score",":", str(round(female_log_likelihood, 3))))
print("%10s %7s %1s" % ("+ Male Score", ":", str(round(male_log_likelihood,3))))
if male_log_likelihood > female_log_likelihood:
winner = "male"
else:
winner = "female"
return winner
#---------------------------------------------------------------------------------------------------------------------------
# Function #4
def process(self):
files = self.get_file_paths(self.females_training_path, self.males_training_path)
for file in files:
self.total_sample += 1
print("%10s %8s %1s" % ("--> Testing", ":", os.path.basename(file)))
vector = self.features_extractor.extract_features(file)
winner = self.identify_gender(vector)
expected_gender = file.split("/")[1][:-26]
print("%10s %6s %1s" % ("+ Expectation",":", expected_gender))
print("%10s %3s %1s" % ("+ Identification", ":", winner))
if winner != expected_gender:
self.error += 1
print("----------------------------------------------------")
accuracy = ( float(self.total_sample - self.error) / float(self.total_sample) ) * 100
accuracy_msg = "*** Accuracy = " + str(round(accuracy, 3)) + "% ***"
print(accuracy_msg)
#------------------------------------------------------------------------------------------------------------------------------
if __name__== "__main__":
gender_identifier = Gender_Identifier("TestingData/females", "TestingData/males", "females.gmm", "males.gmm")
gender_identifier.process()
# -
# ---
# # 6. Conclusions & Analysis
# Looking at the predictions, it's pretty evident that it resulted into 95.749% accuracy. It maybe different for other voice samples. The accuracy can be further improved using GMM Normalization also known as UBM-GMM system.
#
# It was fun contributing my time towards this project!
# ---
# # 7. Acknowledgements & References
# - Machine Learning in Action: Voice Gender Detection
#
# - Reynolds et al. : Using Adapted Gaussian Mixture Models, Digital signal processing 10.1 (2000)
#
# - <NAME>'s Blog
|
other-files/Gender Recognition using Voice.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # PS Orthotile and Landsat 8 Crossovers
#
# Have you ever wanted to compare PS images to Landsat 8 images? Both image collections are made available via the Planet API. However, it takes a bit of work to identify crossovers - that is, images of the same area that were collected within a reasonable time difference of each other. Also, you may be interested in filtering out some imagery, e.g. cloudy images.
#
# This notebook walks you through the process of finding crossovers between PS Orthotiles and Landsat 8 scenes. In this notebook, we specify 'crossovers' as images that have been taken within 1hr of eachother. This time gap is sufficiently small that we expect the atmospheric conditions won't change much (this assumption doesn't always hold, but is the best we can do for now). We also filter out cloudy images and constrain our search to images collected in 2017, January 1 through August 23.
# +
# Notebook dependencies
from __future__ import print_function
import datetime
import json
import os
import ipyleaflet as ipyl
import ipywidgets as ipyw
from IPython.core.display import HTML
from IPython.display import display
import pandas as pd
from planet import api
from planet.api import filters
from shapely import geometry as sgeom
# -
# ## Define AOI
#
# Define the AOI as a geojson polygon. This can be done at [geojson.io](http://geojson.io). If you use geojson.io, only copy the single aoi feature, not the entire feature collection.
aoi = {u'geometry': {u'type': u'Polygon', u'coordinates': [[[-121.3113248348236, 38.28911976564886], [-121.3113248348236, 38.34622533958], [-121.2344205379486, 38.34622533958], [-121.2344205379486, 38.28911976564886], [-121.3113248348236, 38.28911976564886]]]}, u'type': u'Feature', u'properties': {u'style': {u'opacity': 0.5, u'fillOpacity': 0.2, u'noClip': False, u'weight': 4, u'color': u'blue', u'lineCap': None, u'dashArray': None, u'smoothFactor': 1, u'stroke': True, u'fillColor': None, u'clickable': True, u'lineJoin': None, u'fill': True}}}
json.dumps(aoi)
# ## Build Request
#
# Build the Planet API Filter request for the Landsat 8 and PS Orthotile imagery taken in 2017 through August 23.
# define the date range for imagery
start_date = datetime.datetime(year=2017,month=1,day=1)
stop_date = datetime.datetime(year=2017,month=8,day=23)
# +
# filters.build_search_request() item types:
# Landsat 8 - 'Landsat8L1G'
# Sentinel - 'Sentinel2L1C'
# PS Orthotile = 'PSOrthoTile'
def build_landsat_request(aoi_geom, start_date, stop_date):
query = filters.and_filter(
filters.geom_filter(aoi_geom),
filters.range_filter('cloud_cover', lt=5),
# ensure has all assets, unfortunately also filters 'L1TP'
# filters.string_filter('quality_category', 'standard'),
filters.range_filter('sun_elevation', gt=0), # filter out Landsat night scenes
filters.date_range('acquired', gt=start_date),
filters.date_range('acquired', lt=stop_date)
)
return filters.build_search_request(query, ['Landsat8L1G'])
def build_ps_request(aoi_geom, start_date, stop_date):
query = filters.and_filter(
filters.geom_filter(aoi_geom),
filters.range_filter('cloud_cover', lt=0.05),
filters.date_range('acquired', gt=start_date),
filters.date_range('acquired', lt=stop_date)
)
return filters.build_search_request(query, ['PSOrthoTile'])
print(build_landsat_request(aoi['geometry'], start_date, stop_date))
print(build_ps_request(aoi['geometry'], start_date, stop_date))
# -
# ## Search Planet API
#
# The client is how we interact with the planet api. It is created with the user-specific api key, which is pulled from $PL_API_KEY environment variable. Create the client then use it to search for PS Orthotile and Landsat 8 scenes. Save a subset of the metadata provided by Planet API as our 'scene'.
# +
def get_api_key():
return os.environ['PL_API_KEY']
# quick check that key is defined
assert get_api_key(), "PL_API_KEY not defined."
# +
def create_client():
return api.ClientV1(api_key=get_api_key())
def search_pl_api(request, limit=500):
client = create_client()
result = client.quick_search(request)
# note that this returns a generator
return result.items_iter(limit=limit)
items = list(search_pl_api(build_ps_request(aoi['geometry'], start_date, stop_date)))
print(len(items))
# uncomment below to see entire metadata for a PS orthotile
# print(json.dumps(items[0], indent=4))
del items
items = list(search_pl_api(build_landsat_request(aoi['geometry'], start_date, stop_date)))
print(len(items))
# uncomment below to see entire metadata for a landsat scene
# print(json.dumps(items[0], indent=4))
del items
# -
# In processing the items to scenes, we are only using a small subset of the [product metadata](https://www.planet.com/docs/spec-sheets/sat-imagery/#product-metadata).
# +
def items_to_scenes(items):
item_types = []
def _get_props(item):
props = item['properties']
props.update({
'thumbnail': item['_links']['thumbnail'],
'item_type': item['properties']['item_type'],
'id': item['id'],
'acquired': item['properties']['acquired'],
'footprint': item['geometry']
})
return props
scenes = pd.DataFrame(data=[_get_props(i) for i in items])
# acquired column to index, it is unique and will be used a lot for processing
scenes.index = pd.to_datetime(scenes['acquired'])
del scenes['acquired']
scenes.sort_index(inplace=True)
return scenes
scenes = items_to_scenes(search_pl_api(build_landsat_request(aoi['geometry'],
start_date, stop_date)))
# display(scenes[:1])
print(scenes.thumbnail.tolist()[0])
del scenes
# -
# ## Investigate Landsat Scenes
#
# There are quite a few Landsat 8 scenes that are returned by our query. What do the footprints look like relative to our AOI and what is the collection time of the scenes?
# +
landsat_scenes = items_to_scenes(search_pl_api(build_landsat_request(aoi['geometry'],
start_date, stop_date)))
# How many Landsat 8 scenes match the query?
print(len(landsat_scenes))
# -
# ### Show Landsat 8 Footprints on Map
# +
def landsat_scenes_to_features_layer(scenes):
features_style = {
'color': 'grey',
'weight': 1,
'fillColor': 'grey',
'fillOpacity': 0.15}
features = [{"geometry": r.footprint,
"type": "Feature",
"properties": {"style": features_style,
"wrs_path": r.wrs_path,
"wrs_row": r.wrs_row}}
for r in scenes.itertuples()]
return features
def create_landsat_hover_handler(scenes, label):
def hover_handler(event=None, id=None, properties=None):
wrs_path = properties['wrs_path']
wrs_row = properties['wrs_row']
path_row_query = 'wrs_path=={} and wrs_row=={}'.format(wrs_path, wrs_row)
count = len(scenes.query(path_row_query))
label.value = 'path: {}, row: {}, count: {}'.format(wrs_path, wrs_row, count)
return hover_handler
def create_landsat_feature_layer(scenes, label):
features = landsat_scenes_to_features_layer(scenes)
# Footprint feature layer
feature_collection = {
"type": "FeatureCollection",
"features": features
}
feature_layer = ipyl.GeoJSON(data=feature_collection)
feature_layer.on_hover(create_landsat_hover_handler(scenes, label))
return feature_layer
# +
# Initialize map using parameters from above map
# and deleting map instance if it exists
try:
del fp_map
except NameError:
pass
zoom = 6
center = [38.28993659801203, -120.14648437499999] # lat/lon
# +
# Create map, adding box drawing controls
# Reuse parameters if map already exists
try:
center = fp_map.center
zoom = fp_map.zoom
print(zoom)
print(center)
except NameError:
pass
# Change tile layer to one that makes it easier to see crop features
# Layer selected using https://leaflet-extras.github.io/leaflet-providers/preview/
map_tiles = ipyl.TileLayer(url='http://{s}.basemaps.cartocdn.com/light_all/{z}/{x}/{y}.png')
fp_map = ipyl.Map(
center=center,
zoom=zoom,
default_tiles = map_tiles
)
label = ipyw.Label(layout=ipyw.Layout(width='100%'))
fp_map.add_layer(create_landsat_feature_layer(landsat_scenes, label)) # landsat layer
fp_map.add_layer(ipyl.GeoJSON(data=aoi)) # aoi layer
# Display map and label
ipyw.VBox([fp_map, label])
# -
# This AOI is located in a region covered by 3 different path/row tiles. This means there is 3x the coverage than in regions only covered by one path/row tile. This is particularly lucky!
#
# What about the within each path/row tile. How long and how consistent is the Landsat 8 collect period for each path/row?
# +
def time_diff_stats(group):
time_diff = group.index.to_series().diff() # time difference between rows in group
stats = {'median': time_diff.median(),
'mean': time_diff.mean(),
'std': time_diff.std(),
'count': time_diff.count(),
'min': time_diff.min(),
'max': time_diff.max()}
return pd.Series(stats)
landsat_scenes.groupby(['wrs_path', 'wrs_row']).apply(time_diff_stats)
# -
# It looks like the collection period is 16 days, which lines up with the [Landsat 8 mission description](https://landsat.usgs.gov/landsat-8).
#
# path/row 43/33 is missing one image which causes an unusually long collect period.
#
# What this means is that we don't need to look at every Landsat 8 scene collect time to find crossovers with Planet scenes. We could look at the first scene for each path/row, then look at every 16 day increment. However, we will need to account for dropped Landsat 8 scenes in some way.
#
# What is the time difference between the tiles?
# +
def find_closest(date_time, data_frame):
# inspired by:
# https://stackoverflow.com/questions/36933725/pandas-time-series-join-by-closest-time
time_deltas = (data_frame.index - date_time).to_series().reset_index(drop=True).abs()
idx_min = time_deltas.idxmin()
min_delta = time_deltas[idx_min]
return (idx_min, min_delta)
def closest_time(group):
'''group: data frame with acquisition time as index'''
inquiry_date = datetime.datetime(year=2017,month=3,day=7)
idx, _ = find_closest(inquiry_date, group)
return group.index.to_series().iloc[idx]
# for accurate results, we look at the closest time for each path/row tile to a given time
# using just the first entry could result in a longer time gap between collects due to
# the timing of the first entries
landsat_scenes.groupby(['wrs_path', 'wrs_row']).apply(closest_time)
# -
# So the tiles that are in the same path are very close (24sec) together from the same day. Therefore, we would want to only use one tile and pick the best image.
#
# Tiles that are in different paths are 7 days apart. Therefore, we want to keep tiles from different paths, as they represent unique crossovers.
# ## Investigate PS Orthotiles
#
# There are also quite a few PS Orthotiles that match our query. Some of those scenes may not have much overlap with our AOI. We will want to filter those out. Also, we are interested in knowing how many unique days of coverage we have, so we will group PS Orthotiles by collect day, since we may have days with more than one collect (due multiple PS satellites collecting imagery).
# +
all_ps_scenes = items_to_scenes(search_pl_api(build_ps_request(aoi['geometry'], start_date, stop_date)))
# How many PS scenes match query?
print(len(all_ps_scenes))
all_ps_scenes[:1]
# -
# What about overlap? We really only want images that overlap over 20% of the AOI.
#
# Note: we do this calculation in WGS84, the geographic coordinate system supported by geojson. The calculation of coverage expects that the geometries entered are 2D, which WGS84 is not. This will cause a small inaccuracy in the coverage area calculation, but not enough to bother us here.
# +
def aoi_overlap_percent(footprint, aoi):
aoi_shape = sgeom.shape(aoi['geometry'])
footprint_shape = sgeom.shape(footprint)
overlap = aoi_shape.intersection(footprint_shape)
return overlap.area / aoi_shape.area
overlap_percent = all_ps_scenes.footprint.apply(aoi_overlap_percent, args=(aoi,))
all_ps_scenes = all_ps_scenes.assign(overlap_percent = overlap_percent)
all_ps_scenes.head()
# -
print(len(all_ps_scenes))
ps_scenes = all_ps_scenes[all_ps_scenes.overlap_percent > 0.20]
print(len(ps_scenes))
# Ideally, PS scenes have daily coverage over all regions. How many days have PS coverage and how many PS scenes were taken on the same day?
# +
# ps_scenes.index.to_series().head()
# ps_scenes.filter(items=['id']).groupby(pd.Grouper(freq='D')).agg('count')
# -
# Use PS acquisition year, month, and day as index and group by those indices
# https://stackoverflow.com/questions/14646336/pandas-grouping-intra-day-timeseries-by-date
daily_ps_scenes = ps_scenes.index.to_series().groupby([ps_scenes.index.year,
ps_scenes.index.month,
ps_scenes.index.day])
# +
daily_count = daily_ps_scenes.agg('count')
daily_count.index.names = ['y', 'm', 'd']
# How many days is the count greater than 1?
daily_multiple_count = daily_count[daily_count > 1]
print('Out of {} days of coverage, {} days have multiple collects.'.format( \
len(daily_count), len(daily_multiple_count)))
daily_multiple_count.head()
# +
def scenes_and_count(group):
entry = {'count': len(group),
'acquisition_time': group.index.tolist()}
return pd.DataFrame(entry)
daily_count_and_scenes = daily_ps_scenes.apply(scenes_and_count)
# need to rename indices because right now multiple are called 'acquired', which
# causes a bug when we try to run the query
daily_count_and_scenes.index.names = ['y', 'm', 'd', 'num']
multiplecoverage = daily_count_and_scenes.query('count > 1')
multiplecoverage.query('m == 7') # look at just occurrence in July
# -
# Looks like the multiple collects on the same day are just a few minutes apart. They are likely crossovers between different PS satellites. Cool! Since we only want to us one PS image for a crossover, we will chose the best collect for days with multiple collects.
# ## Find Crossovers
#
# Now that we have the PS Orthotiles filtered to what we want and have investigated the Landsat 8 scenes, let's look for crossovers between the two.
#
# First we find concurrent crossovers, PS and Landsat collects that occur within 1hour of each other.
# +
def find_crossovers(acquired_time, landsat_scenes):
'''landsat_scenes: pandas dataframe with acquisition time as index'''
closest_idx, closest_delta = find_closest(acquired_time, landsat_scenes)
closest_landsat = landsat_scenes.iloc[closest_idx]
crossover = {'landsat_acquisition': closest_landsat.name,
'delta': closest_delta}
return pd.Series(crossover)
# fetch PS scenes
ps_scenes = items_to_scenes(search_pl_api(build_ps_request(aoi['geometry'],
start_date, stop_date)))
# for each PS scene, find the closest Landsat scene
crossovers = ps_scenes.index.to_series().apply(find_crossovers, args=(landsat_scenes,))
# filter to crossovers within 1hr
concurrent_crossovers = crossovers[crossovers['delta'] < pd.Timedelta('1 hours')]
print(len(concurrent_crossovers))
concurrent_crossovers
# -
# Now that we have the crossovers, what we are really interested in is the IDs of the landsat and PS scenes, as well as how much they overlap the AOI.
# +
def get_crossover_info(crossovers, aoi):
def get_scene_info(acquisition_time, scenes):
scene = scenes.loc[acquisition_time]
scene_info = {'id': scene.id,
'thumbnail': scene.thumbnail,
# we are going to use the footprints as shapes so convert to shapes now
'footprint': sgeom.shape(scene.footprint)}
return pd.Series(scene_info)
landsat_info = crossovers.landsat_acquisition.apply(get_scene_info, args=(landsat_scenes,))
ps_info = crossovers.index.to_series().apply(get_scene_info, args=(ps_scenes,))
footprint_info = pd.DataFrame({'landsat': landsat_info.footprint,
'ps': ps_info.footprint})
overlaps = footprint_info.apply(lambda x: x.landsat.intersection(x.ps),
axis=1)
aoi_shape = sgeom.shape(aoi['geometry'])
overlap_percent = overlaps.apply(lambda x: x.intersection(aoi_shape).area / aoi_shape.area)
crossover_info = pd.DataFrame({'overlap': overlaps,
'overlap_percent': overlap_percent,
'ps_id': ps_info.id,
'ps_thumbnail': ps_info.thumbnail,
'landsat_id': landsat_info.id,
'landsat_thumbnail': landsat_info.thumbnail})
return crossover_info
crossover_info = get_crossover_info(concurrent_crossovers, aoi)
print(len(crossover_info))
# -
# Next, we filter to overlaps that cover a significant portion of the AOI.
significant_crossovers_info = crossover_info[crossover_info.overlap_percent > 0.9]
print(len(significant_crossovers_info))
significant_crossovers_info
# Browsing through the crossovers, we see that in some instances, multiple crossovers take place on the same day. Really, we are interested in 'unique crossovers', that is, crossovers that take place on unique days. Therefore, we will look at the concurrent crossovers by day.
# +
def group_by_day(data_frame):
return data_frame.groupby([data_frame.index.year,
data_frame.index.month,
data_frame.index.day])
unique_crossover_days = group_by_day(significant_crossovers_info.index.to_series()).count()
print(len(unique_crossover_days))
print(unique_crossover_days)
# -
# There are 6 unique crossovers between Landsat 8 and PS that cover over 90% of our AOI between January and August in 2017. Not bad! That is definitely enough to perform comparison.
# ### Display Crossovers
#
# Let's take a quick look at the crossovers we found to make sure that they don't look cloudy, hazy, or have any other quality issues that would affect the comparison.
# +
# https://stackoverflow.com/questions/36006136/how-to-display-images-in-a-row-with-ipython-display
def make_html(image):
return '<img src="{0}" alt="{0}"style="display:inline;margin:1px"/>' \
.format(image)
def display_thumbnails(row):
print(row.name)
display(HTML(''.join(make_html(t)
for t in (row.ps_thumbnail, row.landsat_thumbnail))))
_ = significant_crossovers_info.apply(display_thumbnails, axis=1)
# -
# They all look pretty good although the last crossover (2017-08-10) could be a little hazy.
|
jupyter-notebooks/crossovers/ps_l8_crossovers.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
# ### <font color = "darkblue">Updates to Assignment</font>
#
# #### If you were working on the older version:
# * Please click on the "Coursera" icon in the top right to open up the folder directory.
# * Navigate to the folder: Week 3/ Planar data classification with one hidden layer. You can see your prior work in version 6b: "Planar data classification with one hidden layer v6b.ipynb"
#
# #### List of bug fixes and enhancements
# * Clarifies that the classifier will learn to classify regions as either red or blue.
# * compute_cost function fixes np.squeeze by casting it as a float.
# * compute_cost instructions clarify the purpose of np.squeeze.
# * compute_cost clarifies that "parameters" parameter is not needed, but is kept in the function definition until the auto-grader is also updated.
# * nn_model removes extraction of parameter values, as the entire parameter dictionary is passed to the invoked functions.
# # Planar data classification with one hidden layer
#
# Welcome to your week 3 programming assignment. It's time to build your first neural network, which will have a hidden layer. You will see a big difference between this model and the one you implemented using logistic regression.
#
# **You will learn how to:**
# - Implement a 2-class classification neural network with a single hidden layer
# - Use units with a non-linear activation function, such as tanh
# - Compute the cross entropy loss
# - Implement forward and backward propagation
#
# ## 1 - Packages ##
#
# Let's first import all the packages that you will need during this assignment.
# - [numpy](https://www.numpy.org/) is the fundamental package for scientific computing with Python.
# - [sklearn](http://scikit-learn.org/stable/) provides simple and efficient tools for data mining and data analysis.
# - [matplotlib](http://matplotlib.org) is a library for plotting graphs in Python.
# - testCases provides some test examples to assess the correctness of your functions
# - planar_utils provide various useful functions used in this assignment
# +
# Package imports
import numpy as np
import matplotlib.pyplot as plt
from testCases_v2 import *
import sklearn
import sklearn.datasets
import sklearn.linear_model
from planar_utils import plot_decision_boundary, sigmoid, load_planar_dataset, load_extra_datasets
# %matplotlib inline
np.random.seed(1) # set a seed so that the results are consistent
# -
# ## 2 - Dataset ##
#
# First, let's get the dataset you will work on. The following code will load a "flower" 2-class dataset into variables `X` and `Y`.
X, Y = load_planar_dataset()
# Visualize the dataset using matplotlib. The data looks like a "flower" with some red (label y=0) and some blue (y=1) points. Your goal is to build a model to fit this data. In other words, we want the classifier to define regions as either red or blue.
# Visualize the data:
plt.scatter(X[0, :], X[1, :], c=Y, s=40, cmap=plt.cm.Spectral);
# You have:
# - a numpy-array (matrix) X that contains your features (x1, x2)
# - a numpy-array (vector) Y that contains your labels (red:0, blue:1).
#
# Lets first get a better sense of what our data is like.
#
# **Exercise**: How many training examples do you have? In addition, what is the `shape` of the variables `X` and `Y`?
#
# **Hint**: How do you get the shape of a numpy array? [(help)](https://docs.scipy.org/doc/numpy/reference/generated/numpy.ndarray.shape.html)
# +
### START CODE HERE ### (≈ 3 lines of code)
shape_X = X.shape
shape_Y = Y.shape
m = len(X[0]) # training set size
### END CODE HERE ###
print ('The shape of X is: ' + str(shape_X))
print ('The shape of Y is: ' + str(shape_Y))
print ('I have m = %d training examples!' % (m))
# -
# **Expected Output**:
#
# <table style="width:20%">
#
# <tr>
# <td>**shape of X**</td>
# <td> (2, 400) </td>
# </tr>
#
# <tr>
# <td>**shape of Y**</td>
# <td>(1, 400) </td>
# </tr>
#
# <tr>
# <td>**m**</td>
# <td> 400 </td>
# </tr>
#
# </table>
# ## 3 - Simple Logistic Regression
#
# Before building a full neural network, lets first see how logistic regression performs on this problem. You can use sklearn's built-in functions to do that. Run the code below to train a logistic regression classifier on the dataset.
# Train the logistic regression classifier
clf = sklearn.linear_model.LogisticRegressionCV();
clf.fit(X.T, Y.T);
# You can now plot the decision boundary of these models. Run the code below.
# +
# Plot the decision boundary for logistic regression
plot_decision_boundary(lambda x: clf.predict(x), X, Y)
plt.title("Logistic Regression")
# Print accuracy
LR_predictions = clf.predict(X.T)
print ('Accuracy of logistic regression: %d ' % float((np.dot(Y,LR_predictions) + np.dot(1-Y,1-LR_predictions))/float(Y.size)*100) +
'% ' + "(percentage of correctly labelled datapoints)")
# -
# **Expected Output**:
#
# <table style="width:20%">
# <tr>
# <td>**Accuracy**</td>
# <td> 47% </td>
# </tr>
#
# </table>
#
# **Interpretation**: The dataset is not linearly separable, so logistic regression doesn't perform well. Hopefully a neural network will do better. Let's try this now!
# ## 4 - Neural Network model
#
# Logistic regression did not work well on the "flower dataset". You are going to train a Neural Network with a single hidden layer.
#
# **Here is our model**:
# <img src="images/classification_kiank.png" style="width:600px;height:300px;">
#
# **Mathematically**:
#
# For one example $x^{(i)}$:
# $$z^{[1] (i)} = W^{[1]} x^{(i)} + b^{[1]}\tag{1}$$
# $$a^{[1] (i)} = \tanh(z^{[1] (i)})\tag{2}$$
# $$z^{[2] (i)} = W^{[2]} a^{[1] (i)} + b^{[2]}\tag{3}$$
# $$\hat{y}^{(i)} = a^{[2] (i)} = \sigma(z^{ [2] (i)})\tag{4}$$
# $$y^{(i)}_{prediction} = \begin{cases} 1 & \mbox{if } a^{[2](i)} > 0.5 \\ 0 & \mbox{otherwise } \end{cases}\tag{5}$$
#
# Given the predictions on all the examples, you can also compute the cost $J$ as follows:
# $$J = - \frac{1}{m} \sum\limits_{i = 0}^{m} \large\left(\small y^{(i)}\log\left(a^{[2] (i)}\right) + (1-y^{(i)})\log\left(1- a^{[2] (i)}\right) \large \right) \small \tag{6}$$
#
# **Reminder**: The general methodology to build a Neural Network is to:
# 1. Define the neural network structure ( # of input units, # of hidden units, etc).
# 2. Initialize the model's parameters
# 3. Loop:
# - Implement forward propagation
# - Compute loss
# - Implement backward propagation to get the gradients
# - Update parameters (gradient descent)
#
# You often build helper functions to compute steps 1-3 and then merge them into one function we call `nn_model()`. Once you've built `nn_model()` and learnt the right parameters, you can make predictions on new data.
# ### 4.1 - Defining the neural network structure ####
#
# **Exercise**: Define three variables:
# - n_x: the size of the input layer
# - n_h: the size of the hidden layer (set this to 4)
# - n_y: the size of the output layer
#
# **Hint**: Use shapes of X and Y to find n_x and n_y. Also, hard code the hidden layer size to be 4.
# +
# GRADED FUNCTION: layer_sizes
def layer_sizes(X, Y):
"""
Arguments:
X -- input dataset of shape (input size, number of examples)
Y -- labels of shape (output size, number of examples)
Returns:
n_x -- the size of the input layer
n_h -- the size of the hidden layer
n_y -- the size of the output layer
"""
### START CODE HERE ### (≈ 3 lines of code)
n_x = len(X) # size of input layer
n_h = 4
n_y = len(Y) # size of output layer
### END CODE HERE ###
return (n_x, n_h, n_y)
# -
X_assess, Y_assess = layer_sizes_test_case()
(n_x, n_h, n_y) = layer_sizes(X_assess, Y_assess)
print("The size of the input layer is: n_x = " + str(n_x))
print("The size of the hidden layer is: n_h = " + str(n_h))
print("The size of the output layer is: n_y = " + str(n_y))
# **Expected Output** (these are not the sizes you will use for your network, they are just used to assess the function you've just coded).
#
# <table style="width:20%">
# <tr>
# <td>**n_x**</td>
# <td> 5 </td>
# </tr>
#
# <tr>
# <td>**n_h**</td>
# <td> 4 </td>
# </tr>
#
# <tr>
# <td>**n_y**</td>
# <td> 2 </td>
# </tr>
#
# </table>
# ### 4.2 - Initialize the model's parameters ####
#
# **Exercise**: Implement the function `initialize_parameters()`.
#
# **Instructions**:
# - Make sure your parameters' sizes are right. Refer to the neural network figure above if needed.
# - You will initialize the weights matrices with random values.
# - Use: `np.random.randn(a,b) * 0.01` to randomly initialize a matrix of shape (a,b).
# - You will initialize the bias vectors as zeros.
# - Use: `np.zeros((a,b))` to initialize a matrix of shape (a,b) with zeros.
# +
# GRADED FUNCTION: initialize_parameters
def initialize_parameters(n_x, n_h, n_y):
"""
Argument:
n_x -- size of the input layer
n_h -- size of the hidden layer
n_y -- size of the output layer
Returns:
params -- python dictionary containing your parameters:
W1 -- weight matrix of shape (n_h, n_x)
b1 -- bias vector of shape (n_h, 1)
W2 -- weight matrix of shape (n_y, n_h)
b2 -- bias vector of shape (n_y, 1)
"""
np.random.seed(2) # we set up a seed so that your output matches ours although the initialization is random.
### START CODE HERE ### (≈ 4 lines of code)
W1 = np.random.randn(n_h,n_x) * 0.01
b1 = np.zeros([n_h, 1])
W2 = np.random.randn(n_y, n_h) * 0.01
b2 = np.zeros([n_y, 1])
### END CODE HERE ###
assert (W1.shape == (n_h, n_x))
assert (b1.shape == (n_h, 1))
assert (W2.shape == (n_y, n_h))
assert (b2.shape == (n_y, 1))
parameters = {"W1": W1,
"b1": b1,
"W2": W2,
"b2": b2}
return parameters
# +
n_x, n_h, n_y = initialize_parameters_test_case()
parameters = initialize_parameters(n_x, n_h, n_y)
print("W1 = " + str(parameters["W1"]))
print("b1 = " + str(parameters["b1"]))
print("W2 = " + str(parameters["W2"]))
print("b2 = " + str(parameters["b2"]))
# -
# **Expected Output**:
#
# <table style="width:90%">
# <tr>
# <td>**W1**</td>
# <td> [[-0.00416758 -0.00056267]
# [-0.02136196 0.01640271]
# [-0.01793436 -0.00841747]
# [ 0.00502881 -0.01245288]] </td>
# </tr>
#
# <tr>
# <td>**b1**</td>
# <td> [[ 0.]
# [ 0.]
# [ 0.]
# [ 0.]] </td>
# </tr>
#
# <tr>
# <td>**W2**</td>
# <td> [[-0.01057952 -0.00909008 0.00551454 0.02292208]]</td>
# </tr>
#
#
# <tr>
# <td>**b2**</td>
# <td> [[ 0.]] </td>
# </tr>
#
# </table>
#
#
# ### 4.3 - The Loop ####
#
# **Question**: Implement `forward_propagation()`.
#
# **Instructions**:
# - Look above at the mathematical representation of your classifier.
# - You can use the function `sigmoid()`. It is built-in (imported) in the notebook.
# - You can use the function `np.tanh()`. It is part of the numpy library.
# - The steps you have to implement are:
# 1. Retrieve each parameter from the dictionary "parameters" (which is the output of `initialize_parameters()`) by using `parameters[".."]`.
# 2. Implement Forward Propagation. Compute $Z^{[1]}, A^{[1]}, Z^{[2]}$ and $A^{[2]}$ (the vector of all your predictions on all the examples in the training set).
# - Values needed in the backpropagation are stored in "`cache`". The `cache` will be given as an input to the backpropagation function.
# +
# GRADED FUNCTION: forward_propagation
def forward_propagation(X, parameters):
"""
Argument:
X -- input data of size (n_x, m)
parameters -- python dictionary containing your parameters (output of initialization function)
Returns:
A2 -- The sigmoid output of the second activation
cache -- a dictionary containing "Z1", "A1", "Z2" and "A2"
"""
# Retrieve each parameter from the dictionary "parameters"
### START CODE HERE ### (≈ 4 lines of code)
W1 = parameters['W1']
b1 = parameters['b1']
W2 = parameters['W2']
b2 = parameters['b2']
### END CODE HERE ###
# Implement Forward Propagation to calculate A2 (probabilities)
### START CODE HERE ### (≈ 4 lines of code)
Z1 = np.dot(W1, X) + b1
A1 = np.tanh(Z1)
Z2 = np.dot(W2, A1) + b2
A2 = sigmoid(Z2)
### END CODE HERE ###
assert(A2.shape == (1, X.shape[1]))
cache = {"Z1": Z1,
"A1": A1,
"Z2": Z2,
"A2": A2}
return A2, cache
# +
X_assess, parameters = forward_propagation_test_case()
A2, cache = forward_propagation(X_assess, parameters)
# Note: we use the mean here just to make sure that your output matches ours.
print(np.mean(cache['Z1']) ,np.mean(cache['A1']),np.mean(cache['Z2']),np.mean(cache['A2']))
# -
# **Expected Output**:
# <table style="width:50%">
# <tr>
# <td> 0.262818640198 0.091999045227 -1.30766601287 0.212877681719 </td>
# </tr>
# </table>
# Now that you have computed $A^{[2]}$ (in the Python variable "`A2`"), which contains $a^{[2](i)}$ for every example, you can compute the cost function as follows:
#
# $$J = - \frac{1}{m} \sum\limits_{i = 1}^{m} \large{(} \small y^{(i)}\log\left(a^{[2] (i)}\right) + (1-y^{(i)})\log\left(1- a^{[2] (i)}\right) \large{)} \small\tag{13}$$
#
# **Exercise**: Implement `compute_cost()` to compute the value of the cost $J$.
#
# **Instructions**:
# - There are many ways to implement the cross-entropy loss. To help you, we give you how we would have implemented
# $- \sum\limits_{i=0}^{m} y^{(i)}\log(a^{[2](i)})$:
# ```python
# logprobs = np.multiply(np.log(A2),Y)
# cost = - np.sum(logprobs) # no need to use a for loop!
# ```
#
# (you can use either `np.multiply()` and then `np.sum()` or directly `np.dot()`).
# Note that if you use `np.multiply` followed by `np.sum` the end result will be a type `float`, whereas if you use `np.dot`, the result will be a 2D numpy array. We can use `np.squeeze()` to remove redundant dimensions (in the case of single float, this will be reduced to a zero-dimension array). We can cast the array as a type `float` using `float()`.
# +
# GRADED FUNCTION: compute_cost
def compute_cost(A2, Y, parameters):
"""
Computes the cross-entropy cost given in equation (13)
Arguments:
A2 -- The sigmoid output of the second activation, of shape (1, number of examples)
Y -- "true" labels vector of shape (1, number of examples)
parameters -- python dictionary containing your parameters W1, b1, W2 and b2
Returns:
cost -- cross-entropy cost given equation (13)
"""
m = Y.shape[1] # number of example
# Compute the cross-entropy cost
### START CODE HERE ### (≈ 2 lines of code)
logprobs = np.multiply(np.log(A2), Y) + np.multiply((1 - Y), np.log(1 - A2))
cost = -np.sum(logprobs) / m
### END CODE HERE ###
cost = np.squeeze(cost) # makes sure cost is the dimension we expect.
# E.g., turns [[17]] into 17
assert(isinstance(cost, float))
return cost
# +
A2, Y_assess, parameters = compute_cost_test_case()
print("cost = " + str(compute_cost(A2, Y_assess, parameters)))
# -
# **Expected Output**:
# <table style="width:20%">
# <tr>
# <td>**cost**</td>
# <td> 0.693058761... </td>
# </tr>
#
# </table>
# Using the cache computed during forward propagation, you can now implement backward propagation.
#
# **Question**: Implement the function `backward_propagation()`.
#
# **Instructions**:
# Backpropagation is usually the hardest (most mathematical) part in deep learning. To help you, here again is the slide from the lecture on backpropagation. You'll want to use the six equations on the right of this slide, since you are building a vectorized implementation.
#
# <img src="images/grad_summary.png" style="width:600px;height:300px;">
#
# <!--
# $\frac{\partial \mathcal{J} }{ \partial z_{2}^{(i)} } = \frac{1}{m} (a^{[2](i)} - y^{(i)})$
#
# $\frac{\partial \mathcal{J} }{ \partial W_2 } = \frac{\partial \mathcal{J} }{ \partial z_{2}^{(i)} } a^{[1] (i) T} $
#
# $\frac{\partial \mathcal{J} }{ \partial b_2 } = \sum_i{\frac{\partial \mathcal{J} }{ \partial z_{2}^{(i)}}}$
#
# $\frac{\partial \mathcal{J} }{ \partial z_{1}^{(i)} } = W_2^T \frac{\partial \mathcal{J} }{ \partial z_{2}^{(i)} } * ( 1 - a^{[1] (i) 2}) $
#
# $\frac{\partial \mathcal{J} }{ \partial W_1 } = \frac{\partial \mathcal{J} }{ \partial z_{1}^{(i)} } X^T $
#
# $\frac{\partial \mathcal{J} _i }{ \partial b_1 } = \sum_i{\frac{\partial \mathcal{J} }{ \partial z_{1}^{(i)}}}$
#
# - Note that $*$ denotes elementwise multiplication.
# - The notation you will use is common in deep learning coding:
# - dW1 = $\frac{\partial \mathcal{J} }{ \partial W_1 }$
# - db1 = $\frac{\partial \mathcal{J} }{ \partial b_1 }$
# - dW2 = $\frac{\partial \mathcal{J} }{ \partial W_2 }$
# - db2 = $\frac{\partial \mathcal{J} }{ \partial b_2 }$
#
# !-->
#
# - Tips:
# - To compute dZ1 you'll need to compute $g^{[1]'}(Z^{[1]})$. Since $g^{[1]}(.)$ is the tanh activation function, if $a = g^{[1]}(z)$ then $g^{[1]'}(z) = 1-a^2$. So you can compute
# $g^{[1]'}(Z^{[1]})$ using `(1 - np.power(A1, 2))`.
# +
# GRADED FUNCTION: backward_propagation
def backward_propagation(parameters, cache, X, Y):
"""
Implement the backward propagation using the instructions above.
Arguments:
parameters -- python dictionary containing our parameters
cache -- a dictionary containing "Z1", "A1", "Z2" and "A2".
X -- input data of shape (2, number of examples)
Y -- "true" labels vector of shape (1, number of examples)
Returns:
grads -- python dictionary containing your gradients with respect to different parameters
"""
m = X.shape[1]
# First, retrieve W1 and W2 from the dictionary "parameters".
### START CODE HERE ### (≈ 2 lines of code)
W1 = parameters["W1"]
W2 = parameters["W2"]
### END CODE HERE ###
# Retrieve also A1 and A2 from dictionary "cache".
### START CODE HERE ### (≈ 2 lines of code)
A1 = cache['A1']
A2 = cache['A2']
### END CODE HERE ###
# Backward propagation: calculate dW1, db1, dW2, db2.
### START CODE HERE ### (≈ 6 lines of code, corresponding to 6 equations on slide above)
dZ2= A2 - Y
dW2 = (1 / m) * np.dot(dZ2, A1.T)
db2 = (1 / m) * np.sum(dZ2, axis=1, keepdims=True)
dZ1 = np.multiply(np.dot(W2.T, dZ2), 1 - np.power(A1, 2))
dW1 = (1 / m) * np.dot(dZ1, X.T)
db1 = (1 / m) * np.sum(dZ1, axis=1, keepdims=True)
### END CODE HERE ###
grads = {"dW1": dW1,
"db1": db1,
"dW2": dW2,
"db2": db2}
return grads
# +
parameters, cache, X_assess, Y_assess = backward_propagation_test_case()
grads = backward_propagation(parameters, cache, X_assess, Y_assess)
print ("dW1 = "+ str(grads["dW1"]))
print ("db1 = "+ str(grads["db1"]))
print ("dW2 = "+ str(grads["dW2"]))
print ("db2 = "+ str(grads["db2"]))
# -
# **Expected output**:
#
#
#
# <table style="width:80%">
# <tr>
# <td>**dW1**</td>
# <td> [[ 0.00301023 -0.00747267]
# [ 0.00257968 -0.00641288]
# [-0.00156892 0.003893 ]
# [-0.00652037 0.01618243]] </td>
# </tr>
#
# <tr>
# <td>**db1**</td>
# <td> [[ 0.00176201]
# [ 0.00150995]
# [-0.00091736]
# [-0.00381422]] </td>
# </tr>
#
# <tr>
# <td>**dW2**</td>
# <td> [[ 0.00078841 0.01765429 -0.00084166 -0.01022527]] </td>
# </tr>
#
#
# <tr>
# <td>**db2**</td>
# <td> [[-0.16655712]] </td>
# </tr>
#
# </table>
# **Question**: Implement the update rule. Use gradient descent. You have to use (dW1, db1, dW2, db2) in order to update (W1, b1, W2, b2).
#
# **General gradient descent rule**: $ \theta = \theta - \alpha \frac{\partial J }{ \partial \theta }$ where $\alpha$ is the learning rate and $\theta$ represents a parameter.
#
# **Illustration**: The gradient descent algorithm with a good learning rate (converging) and a bad learning rate (diverging). Images courtesy of <NAME>.
#
# <img src="images/sgd.gif" style="width:400;height:400;"> <img src="images/sgd_bad.gif" style="width:400;height:400;">
#
#
# +
# GRADED FUNCTION: update_parameters
def update_parameters(parameters, grads, learning_rate = 1.2):
"""
Updates parameters using the gradient descent update rule given above
Arguments:
parameters -- python dictionary containing your parameters
grads -- python dictionary containing your gradients
Returns:
parameters -- python dictionary containing your updated parameters
"""
# Retrieve each parameter from the dictionary "parameters"
### START CODE HERE ### (≈ 4 lines of code)
W1 = parameters["W1"]
b1 = parameters["b1"]
W2 = parameters["W2"]
b2 = parameters["b2"]
### END CODE HERE ###
# Retrieve each gradient from the dictionary "grads"
### START CODE HERE ### (≈ 4 lines of code)
dW1 = grads["dW1"]
db1 = grads["db1"]
dW2 = grads["dW2"]
db2 = grads["db2"]
## END CODE HERE ###
# Update rule for each parameter
### START CODE HERE ### (≈ 4 lines of code)
W1 = W1 - learning_rate * dW1
b1 = b1 - learning_rate * db1
W2 = W2 - learning_rate * dW2
b2 = b2 - learning_rate * db2
### END CODE HERE ###
parameters = {"W1": W1,
"b1": b1,
"W2": W2,
"b2": b2}
return parameters
# +
parameters, grads = update_parameters_test_case()
parameters = update_parameters(parameters, grads)
print("W1 = " + str(parameters["W1"]))
print("b1 = " + str(parameters["b1"]))
print("W2 = " + str(parameters["W2"]))
print("b2 = " + str(parameters["b2"]))
# -
# **Expected Output**:
#
#
# <table style="width:80%">
# <tr>
# <td>**W1**</td>
# <td> [[-0.00643025 0.01936718]
# [-0.02410458 0.03978052]
# [-0.01653973 -0.02096177]
# [ 0.01046864 -0.05990141]]</td>
# </tr>
#
# <tr>
# <td>**b1**</td>
# <td> [[ -1.02420756e-06]
# [ 1.27373948e-05]
# [ 8.32996807e-07]
# [ -3.20136836e-06]]</td>
# </tr>
#
# <tr>
# <td>**W2**</td>
# <td> [[-0.01041081 -0.04463285 0.01758031 0.04747113]] </td>
# </tr>
#
#
# <tr>
# <td>**b2**</td>
# <td> [[ 0.00010457]] </td>
# </tr>
#
# </table>
# ### 4.4 - Integrate parts 4.1, 4.2 and 4.3 in nn_model() ####
#
# **Question**: Build your neural network model in `nn_model()`.
#
# **Instructions**: The neural network model has to use the previous functions in the right order.
# +
# GRADED FUNCTION: nn_model
def nn_model(X, Y, n_h, num_iterations = 10000, print_cost=False):
"""
Arguments:
X -- dataset of shape (2, number of examples)
Y -- labels of shape (1, number of examples)
n_h -- size of the hidden layer
num_iterations -- Number of iterations in gradient descent loop
print_cost -- if True, print the cost every 1000 iterations
Returns:
parameters -- parameters learnt by the model. They can then be used to predict.
"""
np.random.seed(3)
n_x = layer_sizes(X, Y)[0]
n_y = layer_sizes(X, Y)[2]
# Initialize parameters, then retrieve W1, b1, W2, b2. Inputs: "n_x, n_h, n_y". Outputs = "W1, b1, W2, b2, parameters".
### START CODE HERE ### (≈ 5 lines of code)
parameters = initialize_parameters(n_x,n_h,n_y)
W1 = parameters["W1"]
b1 = parameters["b1"]
W2 = parameters["W2"]
b2 = parameters["b2"]
### END CODE HERE ###
# Loop (gradient descent)
for i in range(0, num_iterations):
### START CODE HERE ### (≈ 4 lines of code)
# Forward propagation. Inputs: "X, parameters". Outputs: "A2, cache".
A2, cache = forward_propagation(X,parameters)
# Cost function. Inputs: "A2, Y, parameters". Outputs: "cost".
cost = compute_cost(A2,Y,parameters)
# Backpropagation. Inputs: "parameters, cache, X, Y". Outputs: "grads".
grads = backward_propagation(parameters,cache,X,Y)
# Gradient descent parameter update. Inputs: "parameters, grads". Outputs: "parameters".
parameters = update_parameters(parameters,grads)
### END CODE HERE ###
# Print the cost every 1000 iterations
if print_cost and i % 1000 == 0:
print ("Cost after iteration %i: %f" %(i, cost))
return parameters
# -
X_assess, Y_assess = nn_model_test_case()
parameters = nn_model(X_assess, Y_assess, 4, num_iterations=10000, print_cost=True)
print("W1 = " + str(parameters["W1"]))
print("b1 = " + str(parameters["b1"]))
print("W2 = " + str(parameters["W2"]))
print("b2 = " + str(parameters["b2"]))
# **Expected Output**:
#
# <table style="width:90%">
#
# <tr>
# <td>
# **cost after iteration 0**
# </td>
# <td>
# 0.692739
# </td>
# </tr>
#
# <tr>
# <td>
# <center> $\vdots$ </center>
# </td>
# <td>
# <center> $\vdots$ </center>
# </td>
# </tr>
#
# <tr>
# <td>**W1**</td>
# <td> [[-0.65848169 1.21866811]
# [-0.76204273 1.39377573]
# [ 0.5792005 -1.10397703]
# [ 0.76773391 -1.41477129]]</td>
# </tr>
#
# <tr>
# <td>**b1**</td>
# <td> [[ 0.287592 ]
# [ 0.3511264 ]
# [-0.2431246 ]
# [-0.35772805]] </td>
# </tr>
#
# <tr>
# <td>**W2**</td>
# <td> [[-2.45566237 -3.27042274 2.00784958 3.36773273]] </td>
# </tr>
#
#
# <tr>
# <td>**b2**</td>
# <td> [[ 0.20459656]] </td>
# </tr>
#
# </table>
# ### 4.5 Predictions
#
# **Question**: Use your model to predict by building predict().
# Use forward propagation to predict results.
#
# **Reminder**: predictions = $y_{prediction} = \mathbb 1 \text{{activation > 0.5}} = \begin{cases}
# 1 & \text{if}\ activation > 0.5 \\
# 0 & \text{otherwise}
# \end{cases}$
#
# As an example, if you would like to set the entries of a matrix X to 0 and 1 based on a threshold you would do: ```X_new = (X > threshold)```
# +
# GRADED FUNCTION: predict
def predict(parameters, X):
"""
Using the learned parameters, predicts a class for each example in X
Arguments:
parameters -- python dictionary containing your parameters
X -- input data of size (n_x, m)
Returns
predictions -- vector of predictions of our model (red: 0 / blue: 1)
"""
# Computes probabilities using forward propagation, and classifies to 0/1 using 0.5 as the threshold.
### START CODE HERE ### (≈ 2 lines of code)
A2, cache = forward_propagation(X,parameters)
predictions = 1*(A2>0.5)
### END CODE HERE ###
return predictions
# +
parameters, X_assess = predict_test_case()
predictions = predict(parameters, X_assess)
print("predictions mean = " + str(np.mean(predictions)))
# -
# **Expected Output**:
#
#
# <table style="width:40%">
# <tr>
# <td>**predictions mean**</td>
# <td> 0.666666666667 </td>
# </tr>
#
# </table>
# It is time to run the model and see how it performs on a planar dataset. Run the following code to test your model with a single hidden layer of $n_h$ hidden units.
# +
# Build a model with a n_h-dimensional hidden layer
parameters = nn_model(X, Y, n_h = 4, num_iterations = 10000, print_cost=True)
# Plot the decision boundary
plot_decision_boundary(lambda x: predict(parameters, x.T), X, Y)
plt.title("Decision Boundary for hidden layer size " + str(4))
# -
# **Expected Output**:
#
# <table style="width:40%">
# <tr>
# <td>**Cost after iteration 9000**</td>
# <td> 0.218607 </td>
# </tr>
#
# </table>
#
# Print accuracy
predictions = predict(parameters, X)
print ('Accuracy: %d' % float((np.dot(Y,predictions.T) + np.dot(1-Y,1-predictions.T))/float(Y.size)*100) + '%')
# **Expected Output**:
#
# <table style="width:15%">
# <tr>
# <td>**Accuracy**</td>
# <td> 90% </td>
# </tr>
# </table>
# Accuracy is really high compared to Logistic Regression. The model has learnt the leaf patterns of the flower! Neural networks are able to learn even highly non-linear decision boundaries, unlike logistic regression.
#
# Now, let's try out several hidden layer sizes.
# ### 4.6 - Tuning hidden layer size (optional/ungraded exercise) ###
#
# Run the following code. It may take 1-2 minutes. You will observe different behaviors of the model for various hidden layer sizes.
# +
# This may take about 2 minutes to run
plt.figure(figsize=(16, 32))
hidden_layer_sizes = [1, 2, 3, 4, 5, 20, 50]
for i, n_h in enumerate(hidden_layer_sizes):
plt.subplot(5, 2, i+1)
plt.title('Hidden Layer of size %d' % n_h)
parameters = nn_model(X, Y, n_h, num_iterations = 5000)
plot_decision_boundary(lambda x: predict(parameters, x.T), X, Y)
predictions = predict(parameters, X)
accuracy = float((np.dot(Y,predictions.T) + np.dot(1-Y,1-predictions.T))/float(Y.size)*100)
print ("Accuracy for {} hidden units: {} %".format(n_h, accuracy))
# -
# **Interpretation**:
# - The larger models (with more hidden units) are able to fit the training set better, until eventually the largest models overfit the data.
# - The best hidden layer size seems to be around n_h = 5. Indeed, a value around here seems to fits the data well without also incurring noticeable overfitting.
# - You will also learn later about regularization, which lets you use very large models (such as n_h = 50) without much overfitting.
# **Optional questions**:
#
# **Note**: Remember to submit the assignment by clicking the blue "Submit Assignment" button at the upper-right.
#
# Some optional/ungraded questions that you can explore if you wish:
# - What happens when you change the tanh activation for a sigmoid activation or a ReLU activation?
# - Play with the learning_rate. What happens?
# - What if we change the dataset? (See part 5 below!)
# <font color='blue'>
# **You've learnt to:**
# - Build a complete neural network with a hidden layer
# - Make a good use of a non-linear unit
# - Implemented forward propagation and backpropagation, and trained a neural network
# - See the impact of varying the hidden layer size, including overfitting.
# Nice work!
# ## 5) Performance on other datasets
# If you want, you can rerun the whole notebook (minus the dataset part) for each of the following datasets.
# +
# Datasets
noisy_circles, noisy_moons, blobs, gaussian_quantiles, no_structure = load_extra_datasets()
datasets = {"noisy_circles": noisy_circles,
"noisy_moons": noisy_moons,
"blobs": blobs,
"gaussian_quantiles": gaussian_quantiles}
### START CODE HERE ### (choose your dataset)
dataset = "noisy_moons"
### END CODE HERE ###
X, Y = datasets[dataset]
X, Y = X.T, Y.reshape(1, Y.shape[0])
# make blobs binary
if dataset == "blobs":
Y = Y%2
# Visualize the data
plt.scatter(X[0, :], X[1, :], c=Y, s=40, cmap=plt.cm.Spectral);
# -
# Congrats on finishing this Programming Assignment!
#
# Reference:
# - http://scs.ryerson.ca/~aharley/neural-networks/
# - http://cs231n.github.io/neural-networks-case-study/
|
Neural networks and deeplearning/Planar_data_classification_with_onehidden_layer_v6c.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Feature engineering on NCAA data
#
# Domain knowledge is critical to getting the best out of data analysis and machine learning.
# In the case of basketball, <NAME> identified four factors that are critical to success:
# * Shooting
# * Turnovers
# * Rebounding
# * Free Throws
#
# Of course, it is not enough to identify factors, you need a way to measure them.
#
# Read [this article](https://www.basketball-reference.com/about/factors.html) about the four factors and how they are measured. In this notebook, we will compute them from the box score data. The numbers are slightly different from that of the article because the article is about the NBA, but these numbers are Dean Oliver's variants for NCAA games.
# ## Shooting efficiency
#
# Shooting is measured as the fraction of field goal attempts made, weighting 3 points higher:
#
# $(FG + 0.5 * 3P) / FGA$
#
# Let's compute the offensive and defensive shooting efficiency and see how correlated they are to winning teams.
#
# See [%%bigquery documentation](https://googleapis.github.io/google-cloud-python/latest/bigquery/magics.html) for how to use it.
# %%bigquery df1
SELECT
team_code,
AVG(SAFE_DIVIDE(fgm + 0.5 * fgm3,fga)) AS offensive_shooting_efficiency,
AVG(SAFE_DIVIDE(opp_fgm + 0.5 * opp_fgm3,opp_fga)) AS opponents_shooting_efficiency,
AVG(win) AS win_rate,
COUNT(win) AS num_games
FROM lab_dev.team_box
WHERE fga IS NOT NULL
GROUP BY team_code
# Let's remove the entries corresponding to teams that played fewer than 100 games, and then plot it.
df1 = df1[df1['num_games'] > 100]
df1.plot(x='offensive_shooting_efficiency', y='win_rate', style='o');
df1.plot(x='opponents_shooting_efficiency', y='win_rate', style='o');
# Does the relationship make sense? Do you think offensive and defensive efficiency are good predictors of a team's performance?
# ## Turnover Percentage
#
# Turnover percentage is measured as:
#
# $TOV / (FGA + 0.475 * FTA + TOV - OREB)$
#
# As before, let's compute this, and see whether it is a good predictor. For simplicity, we will compute only offensive turnover percentage, although we should really compute both sides as we did for scoring efficiency.
# %%bigquery df2
SELECT
team_code,
AVG(SAFE_DIVIDE(tov,fga+0.475*fta+tov-oreb)) AS turnover_percent,
AVG(win) AS win_rate,
COUNT(win) AS num_games
FROM lab_dev.team_box
WHERE fga IS NOT NULL
GROUP BY team_code
HAVING num_games > 100
df2.plot(x='turnover_percent', y='win_rate', style='o');
# ## Rebounding
#
# Again, we'd have to measure both sides, but for simplicity, we'll do only the offensive rebounds.
#
# $ORB / (ORB + Opp DRB)$
# %%bigquery df3
SELECT
team_code,
AVG(SAFE_DIVIDE(oreb,oreb + opp_dreb)) AS rebounding,
AVG(win) AS win_rate,
COUNT(win) AS num_games
FROM lab_dev.team_box
WHERE fga IS NOT NULL
GROUP BY team_code
HAVING num_games > 100
df3.plot(x='rebounding', y='win_rate', style='o');
# The relationship doesn't seem all that strong here. One way to measure the strength of the relationship is through the correlation. Numbers near 0 mean not correlated and numbers near +/- 1 indicate high correlation:
df3.corr()['win_rate']
# The correlation between rebounding and win_rate is 0.38. Compare that to the first data frame:
df1.corr()['win_rate']
# Notice that the offensive and opponents efficiency have correlation of 0.67 and -0.66, which are higher.
df2.corr()['win_rate']
# ## Free throw factor
#
# This is a measure of both how often a team gets to the line and how often they make them:
#
# $FT / FGA$
#
# %%bigquery df3
SELECT
team_code,
AVG(SAFE_DIVIDE(ftm,fga)) AS freethrows,
AVG(win) AS win_rate,
COUNT(win) AS num_games
FROM lab_dev.team_box
WHERE fga IS NOT NULL
GROUP BY team_code
HAVING num_games > 100
df3.plot(x='freethrows', y='win_rate', style='o');
df3.corr()['win_rate']
# ## Machine Learning
#
# Let's use these factors to create a simple ML model
# %%bigquery
SELECT
team_code,
is_home,
SAFE_DIVIDE(fgm + 0.5 * fgm3,fga) AS offensive_shooting_efficiency,
SAFE_DIVIDE(opp_fgm + 0.5 * opp_fgm3,opp_fga) AS opponents_shooting_efficiency,
SAFE_DIVIDE(tov,fga+0.475*fta+tov-oreb) AS turnover_percent,
SAFE_DIVIDE(opp_tov,opp_fga+0.475*opp_fta+opp_tov-opp_oreb) AS opponents_turnover_percent,
SAFE_DIVIDE(oreb,oreb + opp_dreb) AS rebounding,
SAFE_DIVIDE(opp_oreb,opp_oreb + dreb) AS opponents_rebounding,
SAFE_DIVIDE(ftm,fga) AS freethrows,
SAFE_DIVIDE(opp_ftm,opp_fga) AS opponents_freethrows,
win
FROM lab_dev.team_box
WHERE fga IS NOT NULL and win IS NOT NULL
LIMIT 10
# +
# %%bigquery
CREATE OR REPLACE MODEL lab_dev.four_factors_model
OPTIONS(model_type='logistic_reg', input_label_cols=['win'])
AS
SELECT
team_code,
is_home,
SAFE_DIVIDE(fgm + 0.5 * fgm3,fga) AS offensive_shooting_efficiency,
SAFE_DIVIDE(opp_fgm + 0.5 * opp_fgm3,opp_fga) AS opponents_shooting_efficiency,
SAFE_DIVIDE(tov,fga+0.475*fta+tov-oreb) AS turnover_percent,
SAFE_DIVIDE(opp_tov,opp_fga+0.475*opp_fta+opp_tov-opp_oreb) AS opponents_turnover_percent,
SAFE_DIVIDE(oreb,oreb + opp_dreb) AS rebounding,
SAFE_DIVIDE(opp_oreb,opp_oreb + dreb) AS opponents_rebounding,
SAFE_DIVIDE(ftm,fga) AS freethrows,
SAFE_DIVIDE(opp_ftm,opp_fga) AS opponents_freethrows,
win
FROM lab_dev.team_box
WHERE fga IS NOT NULL and win IS NOT NULL
# -
# %%bigquery
SELECT * FROM ML.EVALUATE(MODEL lab_dev.four_factors_model)
# 87% isn't bad, but ... there is a *huge* problem with the above approach.
# How are we supposed to know Team A's free throw shooting percentage against Team B before the game is played?
#
# What we could do is to get the free throw shooting percentage of Team A in the 3 games prior to this one and use that. This requires analytic functions in SQL. If you are not familar with these, make a copy of the select statement and modify it in stages until you grasp what is happening.
# +
# %%bigquery
CREATE OR REPLACE MODEL lab_dev.four_factors_model
OPTIONS(model_type='logistic_reg', input_label_cols=['win'])
AS
WITH all_games AS (
SELECT
game_date,
team_code,
is_home,
SAFE_DIVIDE(fgm + 0.5 * fgm3,fga) AS offensive_shooting_efficiency,
SAFE_DIVIDE(opp_fgm + 0.5 * opp_fgm3,opp_fga) AS opponents_shooting_efficiency,
SAFE_DIVIDE(tov,fga+0.475*fta+tov-oreb) AS turnover_percent,
SAFE_DIVIDE(opp_tov,opp_fga+0.475*opp_fta+opp_tov-opp_oreb) AS opponents_turnover_percent,
SAFE_DIVIDE(oreb,oreb + opp_dreb) AS rebounding,
SAFE_DIVIDE(opp_oreb,opp_oreb + dreb) AS opponents_rebounding,
SAFE_DIVIDE(ftm,fga) AS freethrows,
SAFE_DIVIDE(opp_ftm,opp_fga) AS opponents_freethrows,
win
FROM lab_dev.team_box
WHERE fga IS NOT NULL and win IS NOT NULL
)
, prevgames AS (
SELECT
is_home,
AVG(offensive_shooting_efficiency)
OVER(PARTITION BY team_code ORDER BY game_date ASC ROWS BETWEEN 4 PRECEDING AND 1 PRECEDING) AS offensive_shooting_efficiency,
AVG(opponents_shooting_efficiency)
OVER(PARTITION BY team_code ORDER BY game_date ASC ROWS BETWEEN 4 PRECEDING AND 1 PRECEDING)AS opponents_shooting_efficiency,
AVG(turnover_percent)
OVER(PARTITION BY team_code ORDER BY game_date ASC ROWS BETWEEN 4 PRECEDING AND 1 PRECEDING) AS turnover_percent,
AVG(opponents_turnover_percent)
OVER(PARTITION BY team_code ORDER BY game_date ASC ROWS BETWEEN 4 PRECEDING AND 1 PRECEDING) AS opponents_turnover_percent,
AVG(rebounding)
OVER(PARTITION BY team_code ORDER BY game_date ASC ROWS BETWEEN 4 PRECEDING AND 1 PRECEDING) AS rebounding,
AVG(opponents_rebounding)
OVER(PARTITION BY team_code ORDER BY game_date ASC ROWS BETWEEN 4 PRECEDING AND 1 PRECEDING) AS opponents_rebounding,
AVG(freethrows)
OVER(PARTITION BY team_code ORDER BY game_date ASC ROWS BETWEEN 4 PRECEDING AND 1 PRECEDING) AS freethrows,
AVG(opponents_freethrows)
OVER(PARTITION BY team_code ORDER BY game_date ASC ROWS BETWEEN 4 PRECEDING AND 1 PRECEDING) AS oppponents_freethrows,
win
FROM all_games
)
SELECT * FROM prevgames
WHERE offensive_shooting_efficiency IS NOT NULL
# -
# %%bigquery
SELECT * FROM ML.EVALUATE(MODEL lab_dev.four_factors_model)
# Based on just the teams' performance coming in, we can predict the outcome of games with a 69.4% accuracy.
# ## More complex ML model
#
# We can write a more complex ML model using Keras and a deep neural network.
# The code is not that hard but you'll have to do a lot more work (scaling, hyperparameter tuning)
# to get better performance than you did with the BigQuery ML model.
# +
# %%bigquery games
WITH all_games AS (
SELECT
game_date,
team_code,
is_home,
SAFE_DIVIDE(fgm + 0.5 * fgm3,fga) AS offensive_shooting_efficiency,
SAFE_DIVIDE(opp_fgm + 0.5 * opp_fgm3,opp_fga) AS opponents_shooting_efficiency,
SAFE_DIVIDE(tov,fga+0.475*fta+tov-oreb) AS turnover_percent,
SAFE_DIVIDE(opp_tov,opp_fga+0.475*opp_fta+opp_tov-opp_oreb) AS opponents_turnover_percent,
SAFE_DIVIDE(oreb,oreb + opp_dreb) AS rebounding,
SAFE_DIVIDE(opp_oreb,opp_oreb + dreb) AS opponents_rebounding,
SAFE_DIVIDE(ftm,fga) AS freethrows,
SAFE_DIVIDE(opp_ftm,opp_fga) AS opponents_freethrows,
win
FROM lab_dev.team_box
WHERE fga IS NOT NULL and win IS NOT NULL
)
, prevgames AS (
SELECT
is_home,
AVG(offensive_shooting_efficiency)
OVER(PARTITION BY team_code ORDER BY game_date ASC ROWS BETWEEN 4 PRECEDING AND 1 PRECEDING) AS offensive_shooting_efficiency,
AVG(opponents_shooting_efficiency)
OVER(PARTITION BY team_code ORDER BY game_date ASC ROWS BETWEEN 4 PRECEDING AND 1 PRECEDING)AS opponents_shooting_efficiency,
AVG(turnover_percent)
OVER(PARTITION BY team_code ORDER BY game_date ASC ROWS BETWEEN 4 PRECEDING AND 1 PRECEDING) AS turnover_percent,
AVG(opponents_turnover_percent)
OVER(PARTITION BY team_code ORDER BY game_date ASC ROWS BETWEEN 4 PRECEDING AND 1 PRECEDING) AS opponents_turnover_percent,
AVG(rebounding)
OVER(PARTITION BY team_code ORDER BY game_date ASC ROWS BETWEEN 4 PRECEDING AND 1 PRECEDING) AS rebounding,
AVG(opponents_rebounding)
OVER(PARTITION BY team_code ORDER BY game_date ASC ROWS BETWEEN 4 PRECEDING AND 1 PRECEDING) AS opponents_rebounding,
AVG(freethrows)
OVER(PARTITION BY team_code ORDER BY game_date ASC ROWS BETWEEN 4 PRECEDING AND 1 PRECEDING) AS freethrows,
AVG(opponents_freethrows)
OVER(PARTITION BY team_code ORDER BY game_date ASC ROWS BETWEEN 4 PRECEDING AND 1 PRECEDING) AS oppponents_freethrows,
win
FROM all_games
)
SELECT * FROM prevgames
WHERE offensive_shooting_efficiency IS NOT NULL
# -
import tensorflow as tf
import tensorflow.keras as keras
nrows = len(games)
ncols = len(games.iloc[0])
ntrain = (nrows * 7) // 10
print(nrows, ncols, ntrain)
# 0:ntrain are the training data; remaining rows are testing
# last col is the label
train_x = games.iloc[:ntrain, 0:(ncols-1)]
train_y = games.iloc[:ntrain, ncols-1]
test_x = games.iloc[ntrain:, 0:(ncols-1)]
test_y = games.iloc[ntrain:, ncols-1]
model = keras.models.Sequential()
model.add(keras.layers.Dense(5, input_dim=ncols-1, activation='relu'))
model.add(keras.layers.Dense(1, activation='sigmoid'))
model.compile(loss='binary_crossentropy', optimizer='adam', metrics=['accuracy'])
history = model.fit(train_x, train_y, epochs=5, batch_size=32)
score = model.evaluate(test_x, test_y, batch_size=512)
print(score)
# With a deep neural network, we are able to get 71.5% accuracy using the four factors model.
# +
# Copyright 2019 Google Inc. All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
|
blogs/ncaa/ncaa_feateng_solution.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] nbsphinx="hidden"
# This notebook is part of the `nbsphinx` documentation: https://nbsphinx.readthedocs.io/.
# -
# # Usage
# ## Sphinx Setup
#
# In the directory with your notebook files, run this command (assuming you have [Sphinx](https://www.sphinx-doc.org/) installed already):
#
# python3 -m sphinx.cmd.quickstart
#
# Answer the questions that appear on the screen. In case of doubt, just press the `<Return>` key repeatedly to take the default values.
#
# After that, there will be a few brand-new files in the current directory.
# You'll have to make a few changes to the file named `conf.py`. You should at least check if this variable contains the right things:
#
# ```python
# extensions = [
# 'nbsphinx',
# 'sphinx.ext.mathjax',
# ]
# ```
#
# For an example, see this project's [conf.py](conf.py) file.
#
# Once your `conf.py` is in place,
# edit the file named `index.rst` and add the file names of your notebooks
# (without the `.ipynb` extension)
# to the
# [toctree](https://www.sphinx-doc.org/en/master/usage/restructuredtext/directives.html#directive-toctree)
# directive.
# For an example, see this project's `doc/index.rst` file.
#
# Alternatively, you can delete the file `index.rst`
# and replace it with your own notebook called `index.ipynb`
# which will serve as main page.
# In this case you can create the main [toctree](subdir/toctree.ipynb)
# in `index.ipynb`.
# ### Sphinx Configuration Values
#
# All configuration values are described in the
# [Sphinx documentation](http://www.sphinx-doc.org/en/master/usage/configuration.html),
# here we mention only the ones which may be relevant
# in combination with `nbsphinx`.
# #### `exclude_patterns`
#
# Sphinx builds all potential source files (reST files, Jupyter notebooks, ...)
# that are in the source directory (including any sub-directories),
# whether you want to use them or not.
# If you want certain source files not to be built,
# specify them in
# [exclude_patterns](http://www.sphinx-doc.org/en/master/usage/configuration.html#confval-exclude_patterns).
# For example, you might want to ignore source files in your build directory:
#
# ```python
# exclude_patterns = ['_build']
# ```
#
# Note that the directory `.ipynb_checkpoints`
# is automatically added
# to `exclude_patterns`
# by `nbsphinx`.
# #### `extensions`
#
# This is the only required value.
# You have to add `'nbsphinx'` to the list of
# [extensions](http://www.sphinx-doc.org/en/master/usage/configuration.html#confval-extensions),
# otherwise it won't work.
#
# Other interesting extensions are:
#
# * `'sphinx.ext.mathjax'`
# for [math formulas](markdown-cells.ipynb#Equations)
# * `'sphinxcontrib.bibtex'`
# for [bibliographic references](a-normal-rst-file.rst#references)
# * `'sphinxcontrib.rsvgconverter'`
# for [SVG->PDF conversion in LaTeX output](markdown-cells.ipynb#SVG-support-for-LaTeX)
# * `'sphinx_copybutton'`
# for [adding "copy to clipboard" buttons](https://sphinx-copybutton.readthedocs.io/)
# to all text/code boxes
# * `'sphinx_gallery.load_style'` to load CSS styles for [thumbnail galleries](subdir/gallery.ipynb)
# #### `highlight_language`
#
# Default language for syntax highlighting in reST and Markdown cells,
# when no language is specified explicitly.
#
# By default, this is `'python3'`,
# while Jupyter doesn't have a default language.
# Set
# [highlight_language](http://www.sphinx-doc.org/en/master/usage/configuration.html#confval-highlight_language)
# to `'none'` to get the same behavior as in Jupyter:
#
# ```python
# highlight_language = 'none'
# ```
#
# See also [nbsphinx_codecell_lexer](#nbsphinx_codecell_lexer).
# #### `html_css_files`
#
# See [Custom CSS](custom-css.ipynb) and
# [html_css_files](https://www.sphinx-doc.org/en/master/usage/configuration.html#confval-html_css_files).
# #### `html_sourcelink_suffix`
#
#
# By default, a `.txt` suffix is added to source files.
# This is only relevant if the chosen HTML theme supports source links and if
# [html_show_sourcelink](http://www.sphinx-doc.org/en/master/usage/configuration.html#confval-html_show_sourcelink)
# is `True`.
#
# Jupyter notebooks with the suffix `.ipynb.txt` are normally not very useful,
# so if you want to avoid the additional suffix, set
# [html_sourcelink_suffix](http://www.sphinx-doc.org/en/master/usage/configuration.html#confval-html_sourcelink_suffix) to the empty string:
#
# ```python
# html_sourcelink_suffix = ''
# ```
# #### `latex_additional_files`
#
# [latex_additional_files](http://www.sphinx-doc.org/en/master/usage/configuration.html#confval-latex_additional_files)
# can be useful if you are using BibTeX files, see
# [References](a-normal-rst-file.rst#references).
# #### `mathjax_config`
#
# The configuration value
# [mathjax_config](https://www.sphinx-doc.org/en/master/usage/extensions/math.html#confval-mathjax_config)
# can be useful to enable
# [Automatic Equation Numbering](markdown-cells.ipynb#Automatic-Equation-Numbering).
# #### `suppress_warnings`
#
# Warnings can be really helpful to detect small mistakes,
# and you should consider invoking Sphinx with the
# [-W](https://www.sphinx-doc.org/en/master/man/sphinx-build.html#cmdoption-sphinx-build-W)
# option,
# which turns warnings into errors.
# However, warnings can also be annoying,
# especially if you are fully aware of the "problem",
# but you simply don't care about it for some reason.
# In this case, you can use
# [suppress_warnings](https://www.sphinx-doc.org/en/master/usage/configuration.html#confval-suppress_warnings)
# to silence specific types of warnings.
#
# If you want to suppress all warnings from `nbsphinx`, use this:
#
# ```python
# suppress_warnings = [
# 'nbsphinx',
# ]
# ```
#
# You can also be more specific:
#
# ```python
# suppress_warnings = [
# 'nbsphinx.localfile',
# 'nbsphinx.gallery',
# 'nbsphinx.thumbnail',
# 'nbsphinx.notebooktitle',
# 'nbsphinx.ipywidgets',
# ]
# ```
# ### `nbsphinx` Configuration Values
# #### `nbsphinx_allow_errors`
#
# If `True`, the build process is continued even if an exception occurs.
#
# See [Ignoring Errors](allow-errors.ipynb).
# #### `nbsphinx_codecell_lexer`
#
# Default Pygments lexer for syntax highlighting in code cells.
# If available,
# this information is taken from the notebook metadata instead.
#
# Please note that this is not the same as
# [highlight_language](#highlight_language),
# which is used for formatting code in Markdown cells!
# #### `nbsphinx_custom_formats`
#
# See [Custom Notebook Formats](custom-formats.ipynb).
# #### `nbsphinx_epilog`
#
# See [Prolog and Epilog](prolog-and-epilog.ipynb).
# #### `nbsphinx_execute`
#
# Whether to execute notebooks before conversion or not.
# Possible values: `'always'`, `'never'`, `'auto'` (default).
#
# See [Explicitly Dis-/Enabling Notebook Execution](never-execute.ipynb).
# #### `nbsphinx_execute_arguments`
#
# Kernel arguments used when executing notebooks.
#
# If you [use Matplotlib for plots](code-cells.ipynb#Plots),
# this setting is recommended:
#
# ```python
# nbsphinx_execute_arguments = [
# "--InlineBackend.figure_formats={'svg', 'pdf'}",
# "--InlineBackend.rc={'figure.dpi': 96}",
# ]
# ```
#
# If you don't use LaTeX/PDF output,
# you can drop the `'pdf'` figure format.
#
# See [Configuring the Kernels](configuring-kernels.ipynb#Kernel-Arguments).
# #### `nbsphinx_input_prompt`
#
# Input prompt for code cells. `%s` is replaced by the execution count.
#
# To get a prompt similar to the Classic Notebook, use
#
# ```python
# nbsphinx_input_prompt = 'In [%s]:'
# ```
# #### `nbsphinx_kernel_name`
#
# Use a different kernel than stored in the notebook metadata, e.g.:
#
# ```python
# nbsphinx_kernel_name = 'python3'
# ```
#
# See [Configuring the Kernels](configuring-kernels.ipynb#Kernel-Name).
# #### `nbsphinx_output_prompt`
#
# Output prompt for code cells. `%s` is replaced by the execution count.
#
# To get a prompt similar to the Classic Notebook, use
#
# ```python
# nbsphinx_output_prompt = 'Out[%s]:'
# ```
# #### `nbsphinx_prolog`
#
# See [Prolog and Epilog](prolog-and-epilog.ipynb).
# #### `nbsphinx_prompt_width`
#
# Width of input/output prompts (HTML only).
#
# If a prompt is wider than that, it protrudes into the left margin.
#
# Any CSS length can be specified.
# #### `nbsphinx_requirejs_options`
#
# Options for loading RequireJS.
# See [nbsphinx_requirejs_path](#nbsphinx_requirejs_path).
# #### `nbsphinx_requirejs_path`
#
# URL or local path to override the default URL
# for [RequireJS](https://requirejs.org/).
#
# If you use a local file,
# it should be located in a directory listed in
# [html_static_path](http://www.sphinx-doc.org/en/master/usage/configuration.html#confval-html_static_path).
#
# Set to empty string to disable loading RequireJS.
# #### `nbsphinx_responsive_width`
#
# If the browser window is narrower than this,
# input/output prompts are on separate lines
# (HTML only).
#
# Any CSS length can be specified.
# #### `nbsphinx_thumbnails`
#
# A dictionary mapping from a document name
# (i.e. source file without suffix but with subdirectories)
# -- optionally containing wildcards --
# to a thumbnail path to be used in a
# [thumbnail gallery](subdir/gallery.ipynb).
#
# See [Specifying Thumbnails](gallery/thumbnail-from-conf-py.ipynb).
# #### `nbsphinx_timeout`
#
# Controls when a cell will time out.
# The timeout is given in seconds.
# Given `-1`, cells will never time out,
# which is also the default.
#
# See [Cell Execution Timeout](timeout.ipynb).
# #### `nbsphinx_widgets_options`
#
# Options for loading Jupyter widgets resources.
# See [nbsphinx_widgets_path](#nbsphinx_widgets_path).
# #### `nbsphinx_widgets_path`
#
# URL or local path to override the default URL
# for Jupyter widgets resources.
# See [Interactive Widgets (HTML only)](code-cells.ipynb#Interactive-Widgets-(HTML-only)).
#
# If you use a local file,
# it should be located in a directory listed in
# [html_static_path](http://www.sphinx-doc.org/en/master/usage/configuration.html#confval-html_static_path).
#
# For loading the widgets resources,
# RequireJS is needed,
# see [nbsphinx_requirejs_path](#nbsphinx_requirejs_path).
#
# If `nbsphinx_widgets_path` is not specified,
# widgets resources are only loaded if at least one notebook
# actually uses widgets.
# If you are loading the relevant JavaScript code by some other means already,
# you can set this option to the empty string to avoid loading it a second time.
# ## Running Sphinx
#
# To create the HTML pages, use this command:
#
# python3 -m sphinx <source-dir> <build-dir>
#
# If you have many notebooks, you can do a parallel build by using the `-j` option:
#
# python3 -m sphinx <source-dir> <build-dir> -j<number-of-processes>
#
# For example, if your source files are in the current directory and you have 4 CPU cores, you can run this:
#
# python3 -m sphinx . _build -j4
#
# Afterwards, you can find the main HTML file in `_build/index.html`.
#
# Subsequent builds will be faster, because only those source files which have changed will be re-built.
# To force re-building all source files, use the `-E` option.
#
# <div class="alert alert-info">
#
# Note
#
# By default, notebooks will be executed during the Sphinx build process only if they do not have any output cells stored.
# See [Controlling Notebook Execution](executing-notebooks.ipynb).
#
# </div>
#
# To create LaTeX output, use:
#
# python3 -m sphinx <source-dir> <build-dir> -b latex
#
# If you don't know how to create a PDF file from the LaTeX output, you should have a look at [Latexmk](http://personal.psu.edu/jcc8//software/latexmk-jcc/) (see also [this tutorial](https://mg.readthedocs.io/latexmk.html)).
#
# Sphinx can automatically check if the links you are using are still valid.
# Just invoke it like this:
#
# python3 -m sphinx <source-dir> <build-dir> -b linkcheck
# ## Watching for Changes with `sphinx-autobuild`
#
# If you think it's tedious to run the Sphinx build command again and again while you make changes to your notebooks, you'll be happy to hear that there is a way to avoid that: [sphinx-autobuild](https://pypi.org/project/sphinx-autobuild)!
#
# It can be installed with
#
# python3 -m pip install sphinx-autobuild --user
#
# You can start auto-building your files with
#
# python3 -m sphinx_autobuild <source-dir> <build-dir>
#
# This will start a local webserver which will serve the generated HTML pages at http://localhost:8000/.
# Whenever you save changes in one of your notebooks, the appropriate HTML page(s) will be re-built and when finished, your browser view will be refreshed automagically.
# Neat!
#
# You can also abuse this to auto-build the LaTeX output:
#
# python3 -m sphinx_autobuild <source-dir> <build-dir> -b latex
#
# However, to auto-build the final PDF file as well, you'll need an additional tool.
# Again, you can use `latexmk` for this (see [above](#Running-Sphinx)).
# Change to the build directory and run
#
# latexmk -pdf -pvc
#
# If your PDF viewer isn't opened because of LaTeX build errors, you can use the command line flag `-f` to *force* creating a PDF file.
# ## Automatic Creation of HTML and PDF output on readthedocs.org
#
# There are two different methods, both of which are described below.
#
# In both cases, you'll first have to create an account on https://readthedocs.org/
# and connect your GitLab/Github/Bitbucket/... account.
# Instead of connecting, you can also manually add
# any publicly available Git/Subversion/Mercurial/Bazaar/... repository.
#
# After doing the steps described below,
# you only have to "push" to your repository,
# and the HTML pages and the PDF file of your stuff
# are automagically created on readthedocs.org.
# Awesome!
#
# You can even have different versions of your stuff,
# just use Git tags and branches and select in the
# [readthedocs.org settings](https://readthedocs.org/dashboard/)
# which of those should be created.
#
# <div class="alert alert-info">
#
# Note
#
# If you want to execute notebooks
# (see [Controlling Notebook Execution](executing-notebooks.ipynb)),
# you'll need to install the appropriate Jupyter kernel.
# In the examples below,
# the IPython kernel is installed from the packet `ipykernel`.
#
# </div>
# ### Using `requirements.txt`
#
# 1. Create a file named `.readthedocs.yml`
# in the main directory of your repository
# with the following contents:
#
# ```yaml
# version: 2
# formats: all
# python:
# version: 3
# install:
# - requirements: doc/requirements.txt
# system_packages: true
# ```
#
# For further options see https://docs.readthedocs.io/en/latest/config-file/.
#
# 1. Create a file named `doc/requirements.txt`
# (or whatever you chose in the previous step)
# containing the required `pip` packages:
#
# ```
# ipykernel
# nbsphinx
# ```
#
# You can also install directly from Github et al., using a specific branch/tag/commit, e.g.
#
# ```
# git+https://github.com/spatialaudio/nbsphinx.git@master
# ```
# ### Using `conda`
#
# 1. Create a file named `.readthedocs.yml`
# in the main directory of your repository
# with the following contents:
#
# ```yaml
# version: 2
# formats: all
# conda:
# file: doc/environment.yml
# ```
#
# For further options see https://docs.readthedocs.io/en/latest/config-file/.
#
# 1. Create a file named `doc/environment.yml`
# (or whatever you chose in the previous step)
# describing a
# [conda environment](https://docs.conda.io/projects/conda/en/latest/user-guide/tasks/manage-environments.html)
# like this:
#
# ```yaml
# channels:
# - conda-forge
# dependencies:
# - python>=3
# - pandoc
# - ipykernel
# - pip
# - pip:
# - nbsphinx
# ```
#
# It is up to you if you want to install `nbsphinx` with `conda` or with `pip`
# (but note that the `conda` package might be outdated).
# And you can of course add further `conda` and `pip` packages.
# You can also install packages directly from Github et al.,
# using a specific branch/tag/commit, e.g.
#
# ```yaml
# - pip:
# - git+https://github.com/spatialaudio/nbsphinx.git@master
# ```
#
# <div class="alert alert-info">
#
# Note
#
# The specification of the `conda-forge` channel is recommended because it tends to have more recent package versions than the default channel.
#
# </div>
# ## HTML Themes
#
# The `nbsphinx` extension does *not* provide its own theme, you can use any of the available themes or [create a custom one](https://www.sphinx-doc.org/en/master/theming.html#creating-themes), if you feel like it.
#
# The following (incomplete) list of themes contains up to three links for each theme:
#
# 1. The documentation (or the official sample page) of this theme (if available; see also the [documentation of the built-in Sphinx themes](https://www.sphinx-doc.org/en/master/usage/theming.html#builtin-themes))
# 1. How the `nbsphinx` documentation looks when using this theme
# 1. How to enable this theme using either `requirements.txt` or `readthedocs.yml` and theme-specific settings (in some cases)
#
# ### Sphinx's Built-In Themes
#
# * `agogo`:
# [example](https://nbsphinx.readthedocs.io/en/agogo-theme/),
# [usage](https://github.com/spatialaudio/nbsphinx/compare/agogo-theme^...agogo-theme)
#
# * [alabaster](https://alabaster.readthedocs.io/):
# [example](https://nbsphinx.readthedocs.io/en/alabaster-theme/),
# [usage](https://github.com/spatialaudio/nbsphinx/compare/alabaster-theme^...alabaster-theme)
#
# * `bizstyle`:
# [example](https://nbsphinx.readthedocs.io/en/bizstyle-theme/),
# [usage](https://github.com/spatialaudio/nbsphinx/compare/bizstyle-theme^...bizstyle-theme)
#
# * `classic`:
# [example](https://nbsphinx.readthedocs.io/en/classic-theme/),
# [usage](https://github.com/spatialaudio/nbsphinx/compare/classic-theme^...classic-theme)
#
# * `haiku`:
# [example](https://nbsphinx.readthedocs.io/en/haiku-theme/),
# [usage](https://github.com/spatialaudio/nbsphinx/compare/haiku-theme^...haiku-theme)
#
# * `nature`:
# [example](https://nbsphinx.readthedocs.io/en/nature-theme/),
# [usage](https://github.com/spatialaudio/nbsphinx/compare/nature-theme^...nature-theme)
#
# * `pyramid`:
# [example](https://nbsphinx.readthedocs.io/en/pyramid-theme/),
# [usage](https://github.com/spatialaudio/nbsphinx/compare/pyramid-theme^...pyramid-theme)
#
# * `scrolls`:
# [example](https://nbsphinx.readthedocs.io/en/scrolls-theme/),
# [usage](https://github.com/spatialaudio/nbsphinx/compare/scrolls-theme^...scrolls-theme)
#
# * `sphinxdoc`:
# [example](https://nbsphinx.readthedocs.io/en/sphinxdoc-theme/),
# [usage](https://github.com/spatialaudio/nbsphinx/compare/sphinxdoc-theme^...sphinxdoc-theme)
#
# * `traditional`:
# [example](https://nbsphinx.readthedocs.io/en/traditional-theme/),
# [usage](https://github.com/spatialaudio/nbsphinx/compare/traditional-theme^...traditional-theme)
#
# ### 3rd-Party Themes
#
# * [alabaster_jupyterhub](https://github.com/jupyterhub/alabaster-jupyterhub):
# [example](https://nbsphinx.readthedocs.io/en/alabaster-jupyterhub-theme/),
# [usage](https://github.com/spatialaudio/nbsphinx/compare/alabaster-jupyterhub-theme^...alabaster-jupyterhub-theme)
#
# * [basicstrap](https://pythonhosted.org/sphinxjp.themes.basicstrap/):
# [example](https://nbsphinx.readthedocs.io/en/basicstrap-theme/),
# [usage](https://github.com/spatialaudio/nbsphinx/compare/basicstrap-theme^...basicstrap-theme)
#
# * [better](https://sphinx-better-theme.readthedocs.io/):
# [example](https://nbsphinx.readthedocs.io/en/better-theme/),
# [usage](https://github.com/spatialaudio/nbsphinx/compare/better-theme^...better-theme)
#
# * [bootstrap](https://sphinx-bootstrap-theme.readthedocs.io/):
# [example](https://nbsphinx.readthedocs.io/en/bootstrap-theme/),
# [usage](https://github.com/spatialaudio/nbsphinx/compare/bootstrap-theme^...bootstrap-theme)
#
# * [bootstrap-astropy](https://github.com/astropy/astropy-sphinx-theme):
# [example](https://nbsphinx.readthedocs.io/en/astropy-theme/),
# [usage](https://github.com/spatialaudio/nbsphinx/compare/astropy-theme^...astropy-theme)
#
# * [cloud/redcloud/greencloud](https://cloud-sptheme.readthedocs.io/):
# [example](https://nbsphinx.readthedocs.io/en/cloud-theme/),
# [usage](https://github.com/spatialaudio/nbsphinx/compare/cloud-theme^...cloud-theme)
#
# * [dask_sphinx_theme](https://github.com/dask/dask-sphinx-theme):
# [example](https://nbsphinx.readthedocs.io/en/dask-theme/),
# [usage](https://github.com/spatialaudio/nbsphinx/compare/dask-theme^...dask-theme)
#
# * [guzzle_sphinx_theme](https://github.com/guzzle/guzzle_sphinx_theme):
# [example](https://nbsphinx.readthedocs.io/en/guzzle-theme/),
# [usage](https://github.com/spatialaudio/nbsphinx/compare/guzzle-theme^...guzzle-theme)
#
# * [julia](https://github.com/JuliaLang/JuliaDoc):
# [example](https://nbsphinx.readthedocs.io/en/julia-theme/),
# [usage](https://github.com/spatialaudio/nbsphinx/compare/julia-theme^...julia-theme)
#
# * [jupyter](https://github.com/jupyter/jupyter-sphinx-theme/):
# [example](https://nbsphinx.readthedocs.io/en/jupyter-theme/),
# [usage](https://github.com/spatialaudio/nbsphinx/compare/jupyter-theme^...jupyter-theme)
#
# * [maisie_sphinx_theme](https://github.com/maisie-dev/maisie-sphinx-theme):
# [example](https://nbsphinx.readthedocs.io/en/maisie-theme/),
# [usage](https://github.com/spatialaudio/nbsphinx/compare/maisie-theme^...maisie-theme)
#
# * [pangeo](https://github.com/pangeo-data/sphinx_pangeo_theme/):
# [example](https://nbsphinx.readthedocs.io/en/pangeo-theme/),
# [usage](https://github.com/spatialaudio/nbsphinx/compare/pangeo-theme^...pangeo-theme)
#
# * [pydata_sphinx_theme](https://pydata-sphinx-theme.readthedocs.io/):
# [example](https://nbsphinx.readthedocs.io/en/pydata-theme/),
# [usage](https://github.com/spatialaudio/nbsphinx/compare/pydata-theme^...pydata-theme)
#
# * [pytorch_sphinx_theme](https://github.com/shiftlab/pytorch_sphinx_theme):
# [example](https://nbsphinx.readthedocs.io/en/pytorch-theme/),
# [usage](https://github.com/spatialaudio/nbsphinx/compare/pytorch-theme^...pytorch-theme)
#
# * [sizzle](https://docs.red-dove.com/sphinx_sizzle_theme/):
# [example](https://nbsphinx.readthedocs.io/en/sizzle-theme/),
# [usage](https://github.com/spatialaudio/nbsphinx/compare/sizzle-theme^...sizzle-theme)
#
# * [sphinx_material](https://github.com/bashtage/sphinx-material):
# [example](https://nbsphinx.readthedocs.io/en/material-theme/),
# [usage](https://github.com/spatialaudio/nbsphinx/compare/material-theme^...material-theme)
#
# * [sphinx_py3doc_enhanced_theme](https://github.com/ionelmc/sphinx-py3doc-enhanced-theme):
# [example](https://nbsphinx.readthedocs.io/en/py3doc-enhanced-theme/),
# [usage](https://github.com/spatialaudio/nbsphinx/compare/py3doc-enhanced-theme^...py3doc-enhanced-theme)
#
# * [sphinx_pyviz_theme](https://github.com/pyviz-dev/sphinx_pyviz_theme):
# [example](https://nbsphinx.readthedocs.io/en/pyviz-theme/),
# [usage](https://github.com/spatialaudio/nbsphinx/compare/pyviz-theme^...pyviz-theme)
#
# * [sphinx_rtd_theme](https://github.com/readthedocs/sphinx_rtd_theme):
# [example](https://nbsphinx.readthedocs.io/en/rtd-theme/),
# [usage](https://github.com/spatialaudio/nbsphinx/compare/rtd-theme^...rtd-theme)
#
# * [typlog](https://github.com/typlog/sphinx-typlog-theme):
# [example](https://nbsphinx.readthedocs.io/en/typlog-theme/),
# [usage](https://github.com/spatialaudio/nbsphinx/compare/typlog-theme^...typlog-theme)
#
# If you know of another Sphinx theme that should be included here, please open an [issue on Github](https://github.com/spatialaudio/nbsphinx/issues).
# An overview of many more themes can be found at https://sphinx-themes.org/.
# ## Using Notebooks with Git
#
# [Git](https://git-scm.com/) is extremely useful for managing source code and it can and should also be used for managing Jupyter notebooks.
# There is one caveat, however:
# Notebooks can contain output cells with rich media like images, plots, sounds, HTML, JavaScript and many other types of bulky machine-created content.
# This can make it hard to work with Git efficiently, because changes in those bulky contents can completely obscure the more interesting human-made changes in text and source code.
# Working with multiple collaborators on a notebook can become very tedious because of this.
#
# It is therefore highly recommended that you remove all outputs from your notebooks before committing changes to a Git repository (except for the reasons mentioned in [Pre-Executing Notebooks](pre-executed.ipynb)).
#
# If there are no output cells in a notebook, `nbsphinx` will by default execute the notebook, and the pages generated by Sphinx will therefore contain all the output cells.
# See [Controlling Notebook Execution](executing-notebooks.ipynb) for how this behavior can be customized.
#
# In the Jupyter Notebook application, you can manually clear all outputs by selecting
# "Cell" $\to$ "All Output" $\to$ "Clear" from the menu.
# In JupyterLab, the menu items are "Edit" $\to$ "Clear All Outputs".
#
# There are several tools available to remove outputs from multiple files at once without having to open them separately.
# You can even include such a tool as "clean/smudge filters" into your Git workflow, which will strip the output cells automatically whenever a Git command is executed.
# For details, have a look at those links:
#
# * https://github.com/kynan/nbstripout
# * https://github.com/toobaz/ipynb_output_filter
# * https://tillahoffmann.github.io/2017/04/17/versioning-jupyter-notebooks-with-git.html
# * http://timstaley.co.uk/posts/making-git-and-jupyter-notebooks-play-nice/
# * https://pascalbugnion.net/blog/ipython-notebooks-and-git.html
# * https://github.com/choldgraf/nbclean
# * https://jamesfolberth.org/articles/2017/08/07/git-commit-hook-for-jupyter-notebooks/
|
doc/usage.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: KERNEL
# language: python
# name: kernel
# ---
# + [markdown] id="8a7EldJxII_3"
# # Abstract
#
# This notebook provides in-depth look at DCGANs, the motivation behind them, and a highly detailed overview of the optimization techniques and tricks necessary to stabilize training between a Generator/Discriminator pair. This is my implementation of [Radford et al, 2015](https://arxiv.org/abs/1511.06434) with the addition of techniques presented in [Improved GAN, Salimans et al 2016](https://arxiv.org/abs/1606.03498), [Spectral Normalization, Miyato et al 2018](https://arxiv.org/abs/1802.05957), and others. I stray from the common path and try to generate sort-of-HD CelebA images in their original, rectangular, 5:4 aspect ratio at 157x128.
#
# The GAN was first introduced in 2014 by [Goodfellow et al](https://arxiv.org/abs/1406.2661): A pair of two feed-forward networks - a Generator and Discriminator - play a min-max game with each other in an attempt to learn the distribution of image training data and reconstruct it from low-dimensional Gaussian latent vectors.
#
# The GAN was quickly followed by the DCGAN by [Radford et al](https://arxiv.org/pdf/1511.06434v2), a GAN implemented with two separate CNN models. The DCGAN has since taken the mainstage for image generation tasks and has seen many improved iterations dealing with the stability of the min-max game played by the two opposing networks.
#
# The difficulty of training the GAN speaks for itself - once you give it a shot. Achieving stability in training and ensuring convergence to a nash equillibrium is not nearly as easy as it is made out to be by recent DCGAN papers implementing weight and image regularization techniques such as SELU and Spectral Norm. The plain DCGAN, even with contemporary regularization techniques, doesn't quite cut it in the state-of-the-art - but comes surprisingly close for its simplicity. Progressively Growing GANs and the MSG-GAN are some better, more recent candidates which build on the DCGAN to produce truly impressive results at higher resolutions.
# + [markdown] id="ZKfq5PRc9-Pj"
# <!--TABLE OF CONTENTS-->
# # Table of Contents:
# - [Intro to GANs](#Intro-to-GANs)
# - [Dataset Setup & Inspection](#Dataset-Setup-&-Inspection)
# - [Preprocess Images: Resize, Normalize](#Preprocess-Images:-Resize,-Normalize)
# - [Training a GAN](#Training-a-GAN)
# - [The DCGAN Architecture](#The-DCGAN-Architecture)
# - [Stabilizing a DCGAN: One Weird Trick(s)](#Stabilizing-a-DCGAN:-One-Weird-Trick)
# - [Activation functions](#Activation-functions)
# - [SELU and Self-Normalizing Neural Networks](#SELU-and-Self-Normalizing-Neural-Networks)
# - [Spectral Normalization](#Spectral-Normalization)
# - [Minibatch Standard Deviation](#Minibatch-Standard-Deviation)
# - [Pixelwise Normalization](#Pixelwise-Normalization)
# - [WGAN / Wasserstein Loss](#WGAN-/-Wasserstein-Loss)
# - [Gaussian Instance Noise](#Gaussian-Instance-Noise)
# - [Batch Size](#Batch-Size)
# - [Latent Vector Dimension](#Latent-Vector-Dimension)
# - [Building a DCGAN](#Building-a-DCGAN)
# - [Building the Generator Network](#Build-Generator-Network)
# - [Building the Discriminator Network](#Build-Discriminator-Network)
# - [Verifying Model Architecture](#Verify-Model-Architecture)
# - [Choice of Optimizer and Loss Function](#Choice-of-Optimizer-and-Loss-Function)
# - [Training a DCGAN](#Training-a-DCGAN)
# - [Making Training Utilities](#Make-Training-Utilities)
# - [Building Training Loop](#Build-Training-Loop)
# - [Tracking Progress with TensorBoard](#Track-Progress-with-TensorBoard)
# - [Configuring Training Params & Training Model](#Configure-Training-Params-&-Train-Model)
# - [Loading Pre-Trained Checkpoints to Resume Training](#Load-Pre-Trained-Checkpoints-to-Resume-Training)
# - [Testing Model](#Test-Model:-Generate-Images)
# - [Image Generating Utilities](#Image-Generating-Utilities)
# - [Generated Images and Checkpoint Progression](#Generated-Images-and-Checkpoint-Progression)
# - [Lessons Learned: How to Stabilize and Optimize a GAN](#Lessons-Learned:-How-to-Stabilize-and-Optimize-a-GAN)
# - [References](#References)
#
# <br>
#
# - [Appendix A](#Appendix-A)
# - [Animating GAN Training Progress](#Animating-GAN-Training-Progress)
# - [Spherical Interpolation](#Spherical-Interpolation)
# - [CelebA Attributes](#CelebA-Attributes)
# - [Wasserstein Loss: The WGAN](#Wasserstein-Loss:-The-WGAN)
# + [markdown] id="Y585MOPUhanI"
# # Intro to GANs
#
# In this notebook I'm going to give a brief intro to Generative Adversial Networks (GANs) and an in-depth analysis of Deep Convolutional GANs (DCGANs) and their tuning.
#
# [GANs were first introduced by Goodfellow et al in 2014.](https://papers.nips.cc/paper/5423-generative-adversarial-nets.pdf)
#
# We can imagine any training data to lie in an n-dimensional space - for example, 28x28 images from MNIST
# can be sampled from a 28x28 = 784-dimensional space; each of 784 pixels in each image has a probability
# associated with all its possible values, i.e. pixel values in range [0,255]). **Hence, MNIST images can be represented by a probability distribution - one image is a point in a 784-dimensional space**
#
# <img src="img/bivariate gaussian distro.jpg">
#
# <sub><center> A bivariate gaussian distribution of a 2 dimensional space. The height of the plot is the probability of a certain data point appearing. In reality, this plot would have many peaks and can only be visualized on a 784-dimensional MNIST image by first reducing its dimensionality (e.g. with PCA). Image by [<NAME>](https://www.researchgate.net/project/Bayesian-Tracking-of-Multiple-Point-Targets)</center></sub>
#
# The idea is simple: Two networks act as adversaries - the generator (G) has the objective to fool the discriminator (D), while D has the objective to not be fooled.
# G samples data from a random distribution. D is fed either samples from G or samples from real training data and attempts to classify
# G's output as having come from the real training data distribution or not.
#
#
# **G is penalized by outputs which D accurately classifies as fakes; it thus learns to generate data as close
# as possible to the real training data distribution.**
#
# <img src="img/generator.png">
#
# <sub><center>Generator training process; loss is based on discriminator's prediction on generated images. Figure adapted from [Google GAN overview](https://developers.google.com/machine-learning/gan/gan_structure)</center></sub>
#
# **D is penalized by incorrectly classifying fake inputs as real and vice versa; it thus learns to identify fake
# data; D's role is to provide feedback to and facilitate G's learning.**
#
# <img src="img/discriminator.png">
#
# <sub><center> Discriminator training process; loss is based on discriminator's predictions on both fake and real images. Figure adapted from [Google GAN overview](https://developers.google.com/machine-learning/gan/gan_structure)</center></sub>
#
#
#
# For game theory enthusiasts, the generator-discriminator network pair are playing a [minimax game](https://en.wikipedia.org/wiki/Minimax) against each other; their objective function is the predictability of G's output. G tries to minimize predictability while D tries to maximize it.
#
# **When G reaches its optimal strategy, loss converges at a Nash equillibrium for both networks at a saddle point in the objective function representing their game.**
#
# According to Goodfellow et al, the networks' strategy converges when G recovers the training data as
# truthfully as possible and D can't distinguish between fake and real data, being 50% sure that anything G makes is real - and at this point neither G nor D can do their task any better.
#
# **Once G can generate such data, it will theoretically be able to generate images closely resembling real training data by sampling from a low-dimensional space.**
#
# Both networks can be trained by backpropogation; common implementations of generator-discriminator network pairs are
# multilayer perceptrons and CNNs. In this notebook, we'll implement the CNN-based GAN: the DCGAN
#
#
# + id="K52ZH9MUhanO"
import os
import re
import time
import enum
import cv2 as cv
import numpy as np
from scipy.special import expit
import matplotlib.pyplot as plt
import torch
import torchvision.utils
from torch import nn
from torch.optim import Adam, AdamW, SGD
from torchvision import transforms, datasets
from torch.utils.data import DataLoader
from torch.utils.tensorboard import SummaryWriter
# + colab={"base_uri": "https://localhost:8080/"} id="QRvcogx4h1Mw" outputId="4bd3cf47-d079-45ab-c408-737862a19a6f"
# need to import files from google drive into colab
from google.colab import drive
drive.mount("/content/gdrive")
# + [markdown] id="DIR2M0wZhanU"
# We'll try applying the GAN paradigm on the MNIST dataset, our objective being to generate real handwritten digits.
#
#
# Just kidding. [MNIST is no good for computer vision](https://twitter.com/fchollet/status/852594987527045120): Ian Goodfellow, the original author of GANs, and <NAME>, Keras author, give a few details in the above link to their twitter conversation: the gist is that **when you use MNIST, often good CV optimizations don't work, and bad ones do. So MNIST is a poor representation of CV tasks.**
#
# Instead, we're going to use CelebA: Celebrity Face Attributes by [Liu et al, 2015](http://mmlab.ie.cuhk.edu.hk/projects/CelebA.html)
# + colab={"base_uri": "https://localhost:8080/"} id="Ftw1SpJhkOVB" outputId="93c948c4-c5d8-41d8-ed3b-f087c95230b9"
# # copy CelebA dataset from (my) gdrive and unzip
# !cp '/content/gdrive/My Drive/DL/DCGAN/datasets1.zip' .
# !unzip -q datasets1.zip
print('CelebA dataset loaded from gdrive')
# + [markdown] id="QivMboAQvM9Q"
# # Dataset Setup & Inspection
#
# We can download the CelebA dataset using PyTorch directly, as below - but it usually errors due to a limitation with the gdrive where it is hosted. Instead, download it from the [CelebA homepage by Liu et al](http://mmlab.ie.cuhk.edu.hk/projects/CelebA.html). More details on obtaining CelebA provided in code comments.
#
# We'll use torch.datasets.CelebA to make use of its built-in train set split, in place of torch.datasets.ImageFolder, which we would need for a custom dataset. We don't really need train and test sets here - but the **CelebA dataset does specify train/test/validation partition in a text file, so we'll use the training partition to make our results comparable to other DCGAN implementations trained on CelebA.**
# + id="WYOU9kouhanV"
# CelebA (Celebrity Face Attributes), included with PyTorch
# if you try to download fom PyTorch you may encounter an error
# in that case download from official (Liu et al) baidu drive: https://pan.baidu.com/s/1CRxxhoQ97A5qbsKO7iaAJg
# password "<PASSWORD>" for baidu drive
# torch tries to download from (Liu et al) gdrive which has a limit on how many downloads it can provide
# but try it out; you might get lucky. Otherwise, this function just loads from 'root' on disk
class CelebA():
def __init__(self, split, transform=transforms.ToTensor()):
self.data = datasets.CelebA(root='./datasets/CelebA', split=split, download=True, transform=transform)
def __getitem__(self, index):
x = self.data[index]
return x
def __len__(self):
return len(self.data)
# + colab={"base_uri": "https://localhost:8080/"} id="7FFTcOuhL-DE" outputId="b43ea504-945d-42b5-d31a-fa306d460b91"
# instantiate the dataset as an object
dataset = CelebA(split='train', transform=transforms.ToTensor())
# choose a batch size - 128 used in original DCGAN by Radford et al, but lower is better. More on this later.
# I ultimately used a batch size of 32 which has been used by more recent DCGAN papers such as HDCGAN by Curto et al.
# Create a PyTorch DataLoader object which returns CelebA images in shuffled batches of 32,
# discarding the last batch of <32 images with drop_last=True
dataloader = DataLoader(
dataset,
batch_size=32,
num_workers=1,
shuffle=True,
drop_last=True
)
# check dataset size
print(f'CelebA: {dataset.__len__()} celebrity face training images.')
# generate a batch of images from our dataloader
batch = next(iter(dataloader))
# images in (C,H,W) format
print(f'Images are size {batch[0].size()[1:]} (C,H,W)')
# check pixel values
print(f'Pixel values are in range [{torch.min((batch[0][0]))}, {torch.max((batch[0][0]))}]')
# Check labels
print(f'\nLabels are size {batch[1].size()[1:]}')
# should have 40 boolean labels
print(f'CelebA attribute labels: {batch[1][0]}')
# + [markdown] id="NkstdhRHPO13"
# We get ~160k images in the CelebA training set and our dataloader returns two tensors per batch - one with 32 RGB images, one with 32 labels. According to [Liu et al, 2015](http://mmlab.ie.cuhk.edu.hk/projects/CelebA.html), images each have 40 boolean labels representing various visual attributes such as:
# - wearing a hat
# - pointy nose
# - wavy hair
# - sunglasses
# - and so forth.
# + [markdown] id="Q5WJzUV-OpKp"
# # Preprocess Images: Resize, Normalize
#
# The dataset is loaded by TorchVision into a PIL Image with pixel values in range [0,255], while torch.transforms.ToTensor converts the PIL Image to a Float Tensor of shape (C,H,W) in range [0,1]. **We're going to want to normalize these images to [-1,1] before training the discriminator on them because the DCGAN generator uses tanh activation, producing images also in range [-1,1]. From [Radford et al, 2015](https://arxiv.org/pdf/1511.06434v2):** _"We observed that using a bounded [tanh] activation [in the generator's final layer] allowed the model to learn more quickly to saturate and cover the color space of the training distribution."_
#
# We normalize pixels with _pixels = (pixels - mean) / stdev_. **I've seen some DCGAN implementations use a mean of 0.5 and stdev of 0.5 for all pixel values and image batch dimensions, which is incorrect**. The PyTorch image normalization documentation uses values of mean = (0.485, 0.456, 0.406), stdev = (0.229, 0.224, 0.225), which are the values from the [ImageNet dataset](http://www.image-net.org/) - which is the correct way to do it. We'll calculate the mean and stdev of each pixel dimension in (C,H,W) and use those to properly normalize the images to [-1,1].
#
# We use the aligned & cropped CelebA dataset, provided at 218x178. **We'll keep the 5:4 rectangular aspect ratio and resize the images to 157x128 to slightly lower the complexity of the required network, else we would require extra layers to upsample/downsample larger images.**
# + colab={"base_uri": "https://localhost:8080/"} id="VDrRs1zzFdaD" outputId="670844db-2b84-442c-b9b1-da60c18dee9c"
# Get real mean and stdev of each pixel dimension in (C,H,W) in the batch of 10k images
# For reference, ImageNet uses mean=[0.485, 0.456, 0.406], std=[0.229, 0.224, 0.225]
def get_stats(dataloader):
tot_pixels = 0
num_samples = 0
mean = torch.empty(3)
stdev = torch.empty(3)
for data in dataloader:
data = data[0]
b, c, h, w = data.shape
num_pixels = b * h * w
sum_ = torch.sum(data, dim=[0, 2, 3])
sum_of_square = torch.sum(data ** 2, dim=[0, 2, 3])
mean = (tot_pixels * mean + sum_) / (tot_pixels + num_pixels)
stdev = (tot_pixels * stdev + sum_of_square) / (tot_pixels + num_pixels)
num_samples += 1
tot_pixels += num_pixels
print('\r'+f'{(num_samples / len(dataloader)*100):.3f}% processed',end='')
return mean, torch.sqrt(stdev - mean ** 2)
mean, stdev = get_stats(dataloader)
print(f'\nPixel Mean: {mean}, Pixel Stdev: {stdev}')
# + [markdown] id="EXfxkhdWAoFJ"
# Close, but not exactly the same as ImageNet. **Certainly not (0.5,0.5,0.5) for both mean and stdev.**
#
# Now reload the dataset resized, and normalized to [-1,1] according to the values from the previous step. We'll create a composite transform using TorchVision for this:
# + colab={"base_uri": "https://localhost:8080/"} id="02iX0v3AAmf5" outputId="2015d7fe-a00c-47c3-c59b-c4c6649b1a36"
# images are often represented by pixel brightness values in a range [0,255]
# Normalizing pixel values to the [-1,1] range helps train GANs: see https://github.com/soumith/ganhacks
# make a PyTorch function which moves PIL images to a tensor and normalizes them to [-1,1]:
resize_normalize = transforms.Compose([
transforms.Resize(size=(157,128)),
transforms.ToTensor(),
#For reference, ImageNet uses mean=[0.485, 0.456, 0.406], std=[0.229, 0.224, 0.225]
transforms.Normalize(mean=[0.5063,0.4258, 0.3831], std=[0.3107,0.2904,0.2897], inplace=True)
])
# instantiate the resized and normalized dataset
dataset = CelebA(split='train', transform=resize_normalize)
# create a new dataloader with the processed dataset images
dataloader = DataLoader(
dataset,
batch_size=32,
num_workers=1,
shuffle=True,
drop_last=True
)
# get a batch from the dataloader
batch = next(iter(dataloader))
# check that we've normalized to [-1,1] correctly by inspecting pixel dimensions
# batch[0] --> (N,C,H,W); batch[0][0] --> (C,H,W) containing image pixel values
# we want to make sure all pixels in all 3 RGB channels are between [-1,1], i.e. in (C,H,W)
print(f'{batch[0][0][0].numel()} pixels in each {batch[0].size()[2]}x{batch[0].size()[3]} RGB image')
print(f'Pixel values are in range [{torch.min((batch[0][0]))}, {torch.max((batch[0][0]))}]')
# + [markdown] id="wry2wM6lhani"
# Looks good - 157x128 RGB images with pixels normalized to [-1,1].
#
# Let's look at the images:
# + colab={"base_uri": "https://localhost:8080/", "height": 390} id="Pa0C5bPuhani" outputId="6ee005a5-33ae-4ee9-cd3c-85f6b406d900"
# get the images from the batch
img = batch[0]
# we'll visualize a 4x4 grid of images
img_grid_size = 5
img_grid = img[:img_grid_size**2]
# make a grid of images from the tensor; normalize to [0,1] so all pixels are visible
# make_grid pads images with 2 black pixels by default; set pad_value==white (1) so we can see a grid
grid = torchvision.utils.make_grid(tensor=img_grid,nrow=img_grid_size, normalize=True,pad_value=1)
# swap H,W dimensions for matplotlib's imshow() to work properly
grid = np.moveaxis(grid.numpy(), 0, 2)
# plot images
plt.figure(figsize=(6, 6))
plt.title("CelebA Samples")
# imshow plots axes with # of pixels by defaul|t
plt.imshow(grid)
plt.show()
# + [markdown] id="qGQJf5Khhanm"
# Those are faces alright, presumably of celebrities.
#
# And our dataset is ready to use.
# + [markdown] id="0dBm8ujchanu"
# # Training a GAN
# Common implementations of generator-discriminator network pairs use
# multilayer perceptrons and CNNs.
# Both networks of a GAN can be trained by backpropogation of the error from the adversarial loss function, as introduced by [Goodfellow et al, 2014](https://arxiv.org/pdf/1406.2661.pdf):
#
# <img src="img/adversarial loss.GIF"><sub><center>Image from Goodfellow et al, 2014.</center></sub>
#
# - _x_ is an image in (C,H,W) format.
# - _z_ is a vector constructed by sampling from a latent space learned by the generator
# - _D(x)_ is the discriminator function which outputs the probability that _x_ comes from the distribution of the real training data. The function _D_ is simply a binary classifier - real data, or fake data.
# - _G(z)_ is the generator function, transforms the latent space vector _z_ to the same space as the training data (i.e. 3x32x32 for MNIST)
# - _p<sub>data</sub>_ is the probability distribution of the real training data
# - _p<sub>z</sub>_ is the probability distribution of the fake training data output by _G(z)_
# - _D(G(z))_ is the probability that G(z) is classfied same as real data, G(x)
# - D learns to output lim_D(x)_ -> 1, and lim_D(G(z))_ -> 0
# - D's objective is to maximize log(D(x)) - log probability of classifying real data as real data
# - G's objective is to minimize log(1-D(G(z))) - log inverted probability of classifying fake data as real data (i.e. prob of classifying fake data as fake data
#
# We use log probability so probabilities are additive (otherwise the product of probabilities would always approach 0).
# The GAN's loss function is then just binary cross-entropy over outputs from two batches of data; real and fake.
# In Goodfellow et al, the authors say that log(1-D(G(z)) saturates when G is poorly trained and D is well trained.
# According to [Arjovsky and Bottou, 2017](arxiv.org/pdf/1701.04862.pdf), if the discriminator is too good, then its loss gradients will vanish and so the generator will not learn: An optimized disciminator precludes the generator from making progress.
#
# **As such, in practice G is trained by maximizing log(D(G(z))** which provides higher gradients, faster training, and a lesser likelihood of a vanishing gradient. In sum: Instead of training the GAN to minimize D being correct, we train the GAN to maximize that D is incorrect.
#
# And that's it - that's the meat of the GAN, the adversarial loss function: just a doubled application of binary cross entropy.
#
#
# + [markdown] id="p1WiqYbmhanv"
# # The DCGAN Architecture
# It is identical to the original GAN architecture, but uses CNNs for both G and D (instead of dense MLPs).
#
# We should first understand transpose 2D convolutional layers, used for upsampling latent Gaussian vectors into 2D generated images.
#
# <img src="img/conv2dtranspose.gif" width=500 height=500>
# <sub><center> Conv2D Fractionally Strided, i.e. Transpose Kernels; stacked across input channels, they create 3D Conv2D Transpose Filters</center></sub>
#
# Conv transpse is sort of like the opposite of maxpool; conv transpose layers increase dimensionality of their inputs.
# However, conv transpose is unlike maxpool in that it uses kernels with weights, as in regular Conv2D layers. **In transpose convolutional layers, we achieve upsampling by taking the product of _each_ (single) value in the input volume
# with each (of many) weights of the filter.**
#
# From [The original DCGAN by Radford et al](https://arxiv.org/abs/1511.06434), we build the discriminator CNN with:
# - Strided convolutions (CNN filters)
# - No Pooling layers
# - Batch Norm
# - LeakyReLU activation (ReLU that doesn't saturate below 0) for all layers
#
# We build the generator CNN with:
# - Fractional Strided Convolutions (Conv-Transpose Filters)
# - No Pooling layer
# - BatchNorm
# - ReLU activation for all layers except
# - Tanh only on output layer
#
# And they also suggest to avoid fully connected layers in deeper architectures. I've followed these guidelines and then made improvements on them, drawing from papers that followed the DCGAN such as [The Improved GAN, Salimans et al, 2016](https://arxiv.org/abs/1606.03498) and [Progressive Growing GANs, Karras et al, 2017](https://arxiv.org/abs/1710.10196).
#
#
# The input to the generator is a latent space vector _z_ sampled from a standard normal distribution. Its output is an image in standard (C,H,W) format. The Conv2D Transpose filters transform the 1D vector into a 3D volume,
# in direct and opposite analogy to the way regular Conv2D layers transform 3D inputs into 1D vectors, as will our discriminator.
#
# <img src="img/conv2d transpose.png">
#
# <sub><center>Generator from Radford et al. A "backwards" CNN, where a 100-dimensional Gaussian vector is projected to 3D feature map representation by reshaping. Then transpose Conv2D layers transform the input vector into intermediate feature maps and eventually, an image.</center></sub>
#
# Notably, fully connected hidden layers are avoided. **I've used an FC linear layer to upsample the 100-dim input vector to the discriminator's input convolutional filter, while the discriminator 'flattens' its output by a final convolutional layer which produces a scalar.**
#
# Some DCGAN implementations use 1024 filters in the first discriminator and last generator layer - I found no improvement over 512 filters in that layer, with about half the model parameters required. [HDCGAN](https://arxiv.org/abs/1711.06491) used 64 filters in each of its layers - so **it seems that number of filters is less significant than other aspects of the network architecture, particularly normalization of output feature maps.**
#
# Finally, the original DCGAN uses sigmoid activation on the final discriminator layer along with regular binary cross entropy (BCE) loss. **However, I use BCE with logits and no sigmoid activation on the final discriminator layer, which is more numerically stable because we use the log-sum-exp (LSE) trick by combining a sigmoid layer and log-loss in one class.**
#
# ----------
# + [markdown] id="7hqDKZF_hanv"
# # Stabilizing a DCGAN: One Weird Trick
# Now we can define the GAN with some caveats. GANs in general, and especially DCGANs are notoriously unstable due to the competitive interplay between the Discriminator/Generator pair, which encourages weights to spiral out of control for one network to beat the other - and that is the case unless both networks are very carefully tuned. Some recent advances in GAN architecture will help:
#
# ### Activation functions
#
# I found LeakyReLU to work best, as suggested in the original DCGAN paper. I tried ReLU - which lead to less stable loss dynamics and lower gradients - and SELU (Scaled Exponential Linear Unit) - which had its advantages, see below.
#
# Otherwise, I'm going to implement some tricks from recent years that were developed after DCGAN came out. These are mostly normalization techniques - this is necessary so that the discriminator loss is stable and doesn't go to 0 - training collapse - when the discriminator learns too quickly and and the generator is left with zero information gain.
#
# ### SELU and Self-Normalizing Neural Networks
# Self-Normalizing Neural Networks (SNNs) were introduced by [Klambauer et al, 2017](https://arxiv.org/abs/1706.02515) with the SELU activation. They show that activations with close to zero mean and unit variance converge towards zero mean and unit variance as they are propogated through network layers. They show that there is an upper bound on activations that aren't close to zero mean and unit variance as well - so gradient issues become "impossible". SELU worked to normalize regular machine learning models and dense feedforward networks with many layers.
#
# <img src="img/SNN.GIF"><sub><center>Loss on CIFAR-10 classificaiton task using FC feedforward networks, comparing BatchNorm vs SNN at different numbers of hidden layers. Image from [Klambauer et al, 2017](https://arxiv.org/pdf/1706.02515.pdf)</center></sub>
#
# In the [HDCGAN](https://arxiv.org/pdf/1711.06491.pdf), they found that using SELU in conjunction with [BatchNorm](https://arxiv.org/abs/1502.03167) results in model activations with mean close to 0 and variance close to 1. I took inspiration from this paper to try SELU here.
#
# "[Using SELU+BatchNorm] ... given a sufficiently large sample size by the Central Limit Theorem, we can attain NASH equilibrium." -[Curto et al, 2020](https://arxiv.org/pdf/1711.06491.pdf)
#
# However, they found SELU not to work as well in practice, supposedly due to numerical errors introduced by GPU calculation - so they used BatchNorm to overcome this problem. They say this paradigm greatly increased convergence speed.
#
# **I found SELU activation + BatchNorm to work well, but not better than properly tuned LeakyReLU, and at a higher computational cost. I ultimately stuck with LeakyReLU. I attribute this result to the normalizing properties of Spectral Normalization + Batch Normalization - so SELU had little to work on. The idea is sound though - I suspect it will be much more important in larger (>512x512) DCGAN tasks.**
#
# ### Spectral Normalization
# Proposed in [Miyato et al, 2018, Spectral Normalization for Generative Adversarial Networks](https://arxiv.org/abs/1802.05957). They find that Spectral Norm for model weights outperforms the use of gradient penalty for weight normalization in the WGAN-GP, and at lower computational cost.
#
# **Spectral Normalization constrains the [Lipschitz constant](https://en.wikipedia.org/wiki/Lipschitz_continuity) (uniform continuity) of the convolutional filters. In the discriminator, for example, discriminator(x) and discriminator(y) will be sufficiently close granted that distribution of (x) and (y) are sufficiently close, which stabilizes training.**
#
# <img src="img/specnorm.GIF"><sub><center> Imapct of Spectral Normalization (SN) on CIFAR-10 image generation task (Inception score ~= generated image quality) at different Adam hyperparameters (lr, b1, b2) represented by A,B,C,D,E,F. SN makes GANs highly robust to hyperparameter choice.</center></sub>
#
# **With the global weight regularization of the discriminator using Spectral Normalization, we obtain more diverse generated images. I found this to be true, and to also help when added on the Generator.**
#
# ### Minibatch Standard Deviation
# From [Salimans et al 2016, Improved Techniques for Training GANs](https://arxiv.org/abs/1606.03498), they introduce
# "Minibatch discrimination". The idea is to compute feature stats across entire minibatches (not just for each image) and append it to the feature map of each image within that minibatch, the aim being to have generated and real training images match more closely in their statistics. They implement it with a layer at the end of the discriminator which learns a tensor of weights to project the activation to an array of statistics which is used as a feature by the discriminator. [Karras et al, 2018 - Progressive Growing of GANs For Improved Quality, Stability, and Variation](https://arxiv.org/pdf/1710.10196.pdf) suggest a simpler approach they call "Minibatch Standard Deviation", without the weighted tensor. In their approach, **we take the standard deviations of each feature over the entire minibatch and average them, concatenating that to the minibatch as a single additional feature map. Goes on the last layer of the discriminator, before activation.**
#
# ### Pixelwise Normalization
# Pixelwise Normalization is another technique suggested by [Karras et al, 2018](https://arxiv.org/pdf/1710.10196.pdf) in their progressively growing GANs paper. While we usually use BatchNorm in the generator after each conv2d layer, the progressive GAN instead uses pixelwise normalization to transform the feature vector of each pixel in each image to unit length after the last conv2d layer. I've also seen some implementations stack BatchNorm and pixelwise normalization. Karras et al say that pixelwise normalization does not harm the generator in any way, but does prevent activation magnitudes from spiralling out of control due to the competition between discriminator and generator. **In the implementation of pixelwise norm, each pixel in the feature map is simply divided by the L2 norm over that pixel's RGB channels; goes after BatchNorm in generator layers.**
#
#
# ### WGAN / Wasserstein Loss
# I've discussed the WGAN above - and since the idea is to minimize the distance betwene the distribution of the training data and generated data, this should work great. Unfortunately,
# WGAN doesn't work as well as intended. For example, in [Mescheder et al, 2018, Which Training Methods for GANs do actually Converge?](https://arxiv.org/pdf/1801.04406.pdf), WGAN seems to fail to reach convergence and is outperformed in this regard by all other recent implementations - even the regular unregularized GAN authored by Goodfellow et al in 2014.
#
# WGAN loss is simple to implement:
#
# Discriminator (Critic) Loss = [average critic score on real images] – [average critic score on fake images]
# Generator Loss = -[average critic score on fake images]
#
# And we would then clip the weights of the discriminator to [-C, C] (using torch.clamp on model params) at every iteration simplying using to enforce a [Lipschitz constraint](https://en.wikipedia.org/wiki/Lipschitz_continuity) on the discriminator. C=0.05 worked OK for me but not at all great.
#
# From Mescehder et al, 2018: _"Our
# findings show that neither Wasserstein GANs (WGANs) (Arjovsky et al., 2017) nor Wasserstein GANs with Gradient
# Penalty (WGAN-GP) (Gulrajani et al., 2017) converge on this simple example ... On the other hand, we show that instance noise (Sønderby
# et al., 2016; Arjovsky & Bottou, 2017) ... lead to local convergence_"
#
# **WGANs aren't ideal. So I used instance noise instead as suggested by [Sonderby at al, 2016](https://arxiv.org/abs/1610.04490) and verified by [Mescheder et al, 2018](https://arxiv.org/pdf/1801.04406.pdf).** I implemented WGAN loss in the training loop to test it out - it didn't do me any favours.
#
# ### Gaussian Instance Noise
# From [Sonderby et al, 2016](https://arxiv.org/abs/1610.04490), they propose (in the appendix) using instance noise in GANs to stabilize training. It's motivated by the lack of congruence between the supports of the generated and real training data.
#
# **I implemented instance noise by using 10% of the input tensor's pixel values as the standard deviation of noise added to that tensor from a normal (Gaussian) distribution.**
#
# ### Batch Size
# Recent papers use smaller batch sizes, such as [HDCGAN](https://arxiv.org/pdf/1711.06491.pdf) with a batch size of 32. There are many more implementations using smaller batch sizes, and I found this to indeed work better than batch sizes of 128 or larger, especially when dealing with image resolutions larger than 64x64. **Smaller batch sizes should avoid the discriminator learning too quickly and overpowering the generator, especially early in training.**
#
#
# ### Latent Vector Dimension
# The dimension of the latent space vector we give to the generator should determine the quality of its output. However, 100-dimensional latent vectors (_z_) are used in the original DCGAN by Radford et al and in a ton of DCGAN papers to this day. **This number can be reduced with little loss in quality, while higher latent vector dimension leads to higher computational cost with impereceptible improvements, if any.**
# + [markdown] id="V1kGQrdv4OPP"
# **We'll implement and try out Gaussian instance noise, Spectral Normalization, Pixelwise Norm, and Minibatch Standard Deviation. We'll also try SELU activation.**
# + [markdown] id="qWZ-PdSRa6hZ"
# # Building a DCGAN
# + id="6Lunf8aE4NJI"
# simple addition layer of gaussian instance noise
# suggested in https://arxiv.org/abs/1906.04612
class GaussianNoise(nn.Module):
# sigma: sigma*pixel value = stdev of added noise from normal distribution
def __init__(self, sigma=0.1):
super().__init__()
self.sigma = sigma
self.register_buffer('noise', torch.tensor(0))
def forward(self, x):
if self.training and self.sigma != 0:
# scale of noise = stdev of gaussian noise = sigma * pixel value
scale = self.sigma * x.detach()
sampled_noise = self.noise.expand(*x.size()).float().normal_() * scale
x = x + sampled_noise
return x
# spectralnorm constrains the Lipschitz constant (strong form of uniform continuity) of the convolutional filters
# i.e. discriminator(x), discriminator(y) will be sufficiently close granted that distribution of (x) and (y) are sufficiently close
# stabilizes the training of the| discriminator network. works well in practice, but slows down discriminator
# may need to adust discriminator learning rate (2-4x greater than that of generator)
class SpectralNorm(nn.Module):
def __init__(self, module):
super().__init__()
self.module = nn.utils.spectral_norm(module)
def forward(self, x):
return self.module(x)
# From Progressively Growing GANs https://arxiv.org/abs/1710.10196
# For every pixel in a feature map, divide that pixel
# by the L2 norm over that pixel's channels
# theoretically goes after batchnorm only in generator layers
# didn't help my training
class PixelwiseNorm(nn.Module):
def __init__(self, alpha=1e-8):
super().__init__()
self.alpha = alpha
def forward(self, x):
y = x.pow(2.).mean(dim=1, keepdim=True).add(self.alpha).sqrt()
y = x / y
return y
# From Progressively Growing GANs https://arxiv.org/abs/1710.10196
# Standard deviation of each feature in the activation map is calculated
# and then averaged over the minibatch.
# goes on the final layer of discriminator, just before activation
# didn't find it to help my network and I wanted to reduce complexity for easier debugging
class MinibatchStdDev(nn.Module):
def __init__(self, alpha=1e-8):
super().__init__()
self.alpha = alpha
def forward(self, x):
batch_size, _, height, width = x.shape
y = x - x.mean(dim=0, keepdim=True)
y = y.pow(2.).mean(dim=0, keepdim=False).add(self.alpha).sqrt()
y = y.mean().view(1, 1, 1, 1)
y = y.repeat(batch_size, 0, height, width)
y = torch.cat([x, y], 1)
return y
# + [markdown] id="cCihlB7Bhanw"
# ## Build Generator Network
#
# Using all the tricks defined above. I'll make a base transpose convolutional layer and build off of that. The goal of the generator is to upsample a latent vecotr of 100 features up to a 3x157x128 image. **We transform the latent vector with a dense layer to 512x4x4 so that a 512-channel 4x4 kernel filter can take it as input. We also take care to not BatchNorm the last layer of the generator, and use tanh activation instead of SELU/ReLU.** We then proceed with transpose convolutional layers more or less as in a standard CNN. Some DCGAN implementations use dropout in the generator - I found this to actually hurt performance, but to give a moderate boost to diversity of generated images when used in the discriminator.
#
# + id="8TUBZ9yLhanw"
# dimension of the latent space vector we give to the generator
# dim (100,) 1D array
# 100 is used in a ton of GAN papers up to this day; this number can be reduced with little loss in quality
# higher latent vector dim leads to higher computational cost
latent_vector_dim = 100
####################### GENERATOR CNN ############################################
# kernel to (5,4) for rectangular original CelebA aspect ratio, else (4,4)
def conv2d_transpose_block(in_channels, out_channels, kernel=(5,4), stride=2, padding=1, noise=True, normalize=True, dropout=True, activation=False):
# build the base conv2d layer for downsampling
conv2d_upsample = nn.ConvTranspose2d(
in_channels=in_channels,
out_channels=out_channels,
kernel_size=kernel,
stride=stride,
padding=padding,
bias=False)
# make a list of layers which we'll unpack when building the sequential model for the generator
layers = [SpectralNorm(conv2d_upsample)]
# add a BatchNorm layer before activation (batchnorm not present on last layer of generator)
# Goodfellow has a talk where he says placement of activation does not matter w.r.t. BatchNorm
# Others have said that activation is better after batchnorm (Szegedy)
if normalize:
layers.append(nn.BatchNorm2d(out_channels))
# we only use gaussian noise in the discriminator; didn't help training to have it on generator
#if noise:
# layers.append(GaussianNoise(out_channels))
# use only in D, didn't help here
#if dropout:
# layers.append(nn.Dropout(0.3))
if activation:
layers.append(activation)
else:
# modify tensors in place to save VRAM when processing high res images
#nn.SELU(inplace=True)) works very well; however ReLU trained faster
layers.append(nn.ReLU(inplace=True))
return layers
class ConvolutionalGenerativeNet(nn.Module):
def __init__(self):
super().__init__()
# linear layer of generator takes 1D latent vector dim to generator image input volume dim
# generator input volume dim == 512x4x4: 512 input filters and 4x4 kernel size
self.linear = nn.Linear(latent_vector_dim, 512 * (4*4))
# unpack layers from conv2d_transpose_block for the transpose (upsampling) conv layers
self.generator = nn.Sequential(
*conv2d_transpose_block(512,512),
*conv2d_transpose_block(512,256),
*conv2d_transpose_block(256,128),
*conv2d_transpose_block(128,64),
# padding to (2,1) on last layer for rectangular 218x178 (resized to 157x128) original CelebA size
*conv2d_transpose_block(64,3, padding=(2,1), normalize=False, dropout=False, noise=False, activation=nn.Tanh())
)
def forward(self, latent_vector_batch):
# pass 1D latent vectors through linaer layer to transform into input image volume dim 1024x4x4
latent_vector_batch = self.linear(latent_vector_batch)
# reshape into 3D volume 512x4x4 (input filter depth on 1st generator layer * kernel size)
latent_vector_batch = latent_vector_batch.view(latent_vector_batch.shape[0], 512, 4, 4)
return self.generator(latent_vector_batch)
# + [markdown] id="FkSQ13nrhan3"
# ## Build Discriminator Network
#
# A standard CNN, for all intents and purposes - plus some GAN tricks.
#
# The DCGAN paper by Radford et al doesn't use a final FC layer to flatten the feature map; instead, we flatten with a final conv2d layer. **We structure the network such that the final Conv2D layer receives a 512x4x4 feature map and passes a single 512x4x4 filter over it to output a tensor of size (1,1,1), i.e. a scalar value.**
#
# **Take care not to normalize the first or last layer. Spectral Normalization on all layers except the last one, which outputs a scalar. We use dropout on all layers except the last one, and I found that to improve the diversity of images created by the generator.**
# + id="V8XLODabhan3"
############################# DISCRIMINATOR CNN ##########################################
# make a function to create a conv2d block since first DCGAN layer doesn't use batchnorm
# and last layer doesn't use ReLU (tanh in Radford et al, 2014)
def conv2d_block(in_channels, out_channels, kernel=4, stride=2, padding=1, specnorm=True, normalize=True, noise=True, activation=False, dropout=True):
# build the base conv2d layer for downsampling
conv2d_downsample = nn.Conv2d(
in_channels=in_channels,
out_channels=out_channels,
kernel_size=kernel,
stride=stride,
padding=padding,
bias=False)
# make a list of layers to unpack when building the sequential model for discriminator
# don't use specnorm in final layer which outputs scalar and simulates an FC layer
if specnorm:
layers = [SpectralNorm(conv2d_downsample)]
else:
layers = [(conv2d_downsample)]
# add a BatchNorm layer if desired
# but spectral norm replaces batch norm completely in discriminator; present in all layers
#if normalize:
# layers.append(nn.BatchNorm2d(out_channels))
# Goodfellow has a talk where he says placement of activation does not matter w.r.t. BatchNorm
# Others have said that activation is better after batchnorm (such as Francois Chollet quoting Szegedy directly - easily google-able)
if activation:
layers.append(activation)
else:
# modify tensors in place to save VRAM when processing high res images
# SELU() worked well but is outperformed in speed by nn.LeakyReLU(0.2)
layers.append(nn.LeakyReLU(negative_slope=0.2, inplace=True))
# works well in discriminator with dropout p value in range [0.15,0.5]
if dropout:
layers.append(nn.Dropout(0.3))
return layers
class ConvolutionalDiscriminativeNet(nn.Module):
def __init__(self):
super().__init__()
# 2nd last layer outputs (512), and the last layer applies a single 512 filter
# The DCGAN paper doesn't use a final FC layer; instead of FC dense layer to flatten, we use a final conv2d layer
# the final conv2d layer replaces the FC layer because a single 512 filter will output (Kx1x1), i.e. a scalar value K
# we then pass K into sigmoid activation and the "FC conv layer"/"flattening" is complete
# unpack layers from conv2d_block for the regular (downsampling) conv layers
self.discriminator = nn.Sequential(
# Gaussian noise with stdev 0.1 relative to input tensor
# Salimans et al used it in the output layer of D while OpenAI has it on the input
GaussianNoise(),
# no batch norm layer on layer 1 helps performance,
*conv2d_block(3, 64, normalize=False),
*conv2d_block(64, 128),
*conv2d_block(128, 256),
*conv2d_block(256, 512),
*conv2d_block(512, 512),
# Minibatch standard deviation as in Karras et al, on final layer
# MinibatchStdDev(),
# "fake FC layer": no batchnorm, no padding
# no Gaussian noise, batchnorm, dropout on last layer
# use BCELossWithLogits which implements sigmoid activation; so no activation on last layer
*conv2d_block(512, 1, noise=False, specnorm=False, normalize=False, padding=0, dropout=False),
)
def forward(self, batch):
return self.discriminator(batch)
# + [markdown] id="6Iw8XQy-Eg_H"
# # Verify Model Architecture
# Let's make sure we correctly downsample our 3x157x128 images to a binary cross entropy-classifiable scalar, and upsample our gaussian latent vector (with 100 features total per image) to the original 3x157x128 image size.
# + colab={"base_uri": "https://localhost:8080/"} id="dFoF64bRtIGu" outputId="59937384-02bb-4dce-ba69-48d4252fb91b"
from torchsummary import summary
# need device to instantiate model
device = 'cuda'
# Instantiate Discriminator
model = ConvolutionalDiscriminativeNet().to(device)
# USE INPUT IMAGE SIZE. Should downsample to scalar (1x1x1) CxHxW
print('DISCRIMINATOR ARCHITECTURE: Downsample Image dim==(3x157x128) to Scalar (1x1x1)')
summary(model, input_size=(3,157,128))
# Instantiate Generator
model = ConvolutionalGenerativeNet().to(device)
# USE LATENT VECTOR (z) as input. Should upsample to (3x157x128) CxHxW
print('\n\nGENERATOR ARCHITECTURE: Upsample Latent Space Vector dim==(100,) to Image (3x157x128)')
summary(model, input_size=((latent_vector_dim,)))
# + [markdown] id="tpQa0tzpFybV"
# Looks good. We should be able to compute a prediction on a generated image by passing a latent vector through G, and the result through D to get D(G(z)) to plug into the adversarial loss function.
# + [markdown] id="vh2CMoEEhaoH"
# # Choice of Optimizer and Loss Function
# [Radford et al 2015](https://arxiv.org/abs/1511.06434) used Adam for both discriminator and generator with lr=0.0002. I extensively tried to optimize on this and could find no better - although SGD is commonly used to slow down the discriminator in absence of the normalization techniques we've implemented. SGD was much too slow for the normalized discriminator to learn, and Adam worked decently with the parameters lr = 0.0002, b1=0.5, b2 = 0.999 from the original DCGAN by Radford et al 2015.
#
# However, I have swapped Adam for AdamW, [Decoupled Weight Decay Regularization by Loshchilov and Hutter, 2017](https://arxiv.org/abs/1711.05101). Adam incorrectly implements weight decay regularization by equating it with L2 regularization, which is not correct. AdamW overcomes this issue by decoupling the weight decay from the learning rate, with the paper showing improved performance particularly on image classification tasks - precisely as we have here in the discriminator.
#
# As for learning rates - lr=0.0002 as proposed in the original DCGAN paper by Radford et al worked great for me. [Heusel et al, 2017 - GANs Trained by a Two Time-Scale Update Rule [TTUR] Converge to a Local Nash Equilibrium](https://arxiv.org/abs/1706.08500) suggest that different learning rates for the discriminator and generator can also work well, but I didn't find "TTUR" necesary or helpful.
#
# Instead of using the standard torch binary cross entropy class BCELoss(), we'll use BCELossWithLogits. This is because it is more numerically stable since it uses the [log-sum-exp (LSE) trick](https://en.wikipedia.org/wiki/LogSumExp#log-sum-exp_trick_for_log-domain_calculations) by combining a sigmoid layer and log-loss in one class. Because of this, we won't use a sigmoid activation as the final layer of the discriminator as in Radford et al.
# + id="g_hRgUqZhaoI"
####### Note that these are here for a proper python implementation #############################
####### But they have to be specified explicitly in the training loop for google colab - a bug #####
def get_optimizers(G, D):
# optimizer for discrminator
D_optimizer = Adam(D.parameters(), lr=0.0002,betas=(0.5, 0.999))
# optimizer for generator
# currently using same optimizer for both networks; can (and should) play with lr
G_optimizer = Adam(G.parameters(), lr=0.0002, betas=(0.5, 0.999))
return D_optimizer, G_optimizer
# using standard binary cross entropy works great
def get_criterion():
GAN_loss = nn.BCEWithLogitsLoss()
return GAN_loss
# + [markdown] id="iSaQ0qwta_a5"
# # Training a DCGAN
# + [markdown] id="ZNq4_lTXhaoS"
# ## Make Training Utilities
#
# Generating the latent vectors for each batch.
#
# Weight initialization to a standard normal distribution as in Radford et al, 2015, with mean=0 and stdev = 0.02.
#
# And we plot the mean and max of the gradients in each layer of the discriminator (whose gradients are ultimately responsible for the entire network's learning) to make sure we don't have anything vanish or explode. Will show us whether gradients are behaving properly, and at which layer. We call this function specifically for the discriminator network, since we're most concerned with creating stable dynamics for the discriminator gradients - they define the learning of the generator as well (and so the entire DCGAN).
# + id="JNpIMHvPhaoZ"
# generates a latent vector (z) sampled from a standard normal (Gaussian) distribution
# adapted from https://discuss.pytorch.org/t/check-gradient-flow-in-network/15063/10
def get_gaussian_latent_vectors(batch_size, device):
return torch.randn((batch_size, latent_vector_dim), device=device)
# this explicit weight init from standard normal distribution didn't improve performance
# possibly because batchnorm is enough to mitigate the impact of poorly initialized weights
def init_weights(model):
classname = model.__class__.__name__
#print(classname)
if classname.find("Conv2d") != -1:
torch.nn.init.normal_(model.weight.data, 0.0, 0.02)
elif classname.find("BatchNorm2d") != -1:
# 1-centered normal distribution, stdev==0.02 as specified in Radford et al, 2015
torch.nn.init.normal_(model.weight.data, 1.0, 0.02)
torch.nn.init.constant_(model.bias.data, 0.0)
print(f'Weights initialized for {model.__class__.__name__}')
# function to plot gradients after training loop is over
# will make it obvious if any gradients are exploding/vanishing
# goes into training loop after loss.backwards()
from matplotlib.lines import Line2D
def plot_grad_flow(named_parameters):
ave_grads = []
max_grads= []
layers = []
for n, p in named_parameters:
if(p.requires_grad) and ("bias" not in n):
layers.append(n)
ave_grads.append(p.grad.abs().mean())
max_grads.append(p.grad.abs().max())
plt.bar(np.arange(len(max_grads)), max_grads, alpha=0.1, lw=1, color="c")
plt.bar(np.arange(len(max_grads)), ave_grads, alpha=0.1, lw=1, color="b")
plt.hlines(0, 0, len(ave_grads)+1, lw=2, color="k" )
plt.xticks(range(0,len(ave_grads), 1), layers, rotation="vertical")
plt.xlim(left=0, right=len(ave_grads))
plt.ylim(bottom = -0.001, top=0.02) # zoom in on the lower gradient regions
plt.xlabel("Layers")
plt.ylabel("average gradient")
plt.title("Gradient flow")
plt.grid(True)
plt.legend([Line2D([0], [0], color="c", lw=4),
Line2D([0], [0], color="b", lw=4),
Line2D([0], [0], color="k", lw=4)], ['max-gradient', 'mean-gradient', 'zero-gradient'])
# + [markdown] id="QoYR8anphaow"
# ## Build Training Loop
#
# Based on the [original DCGAN paper by Radford et al](https://arxiv.org/abs/1511.06434), here's what we're going for:
#
# **Update Discriminator:**
#
# - Forward real images minibatch through the Discriminator;
#
# - Compute Discriminator loss and calculate gradients for the backward pass;
#
# - Generate fake images minibatch via forward pass of latent vector z to Generator
#
# - Forward generated fake minibatch (G(z)) through the Discriminator;
#
# - Compute the Discriminator loss and compute gradients for the backward pass;
#
# - Add: real minibatch gradients + fake minibatch gradients
#
# - Update Discriminator using Adam or SGD.
#
# **Update Generator:**
#
# - **Log Trick:** Flip the target labels: fake images from the generator G(z) get labeled as real. This step configures binary cross-entropy to maximize log(D(G(z))) for the Discriminator loss passed to the Generator and helps overcome the problem of Generator's vanishing gradients. This is the log trick where instead of minizming log(1 - D(G(z))) we instead maximize log(D(G(z))).
#
# - Forward fake images minibatch through the updated Discriminator: D(G(z)), where z is labeled as real images for the log trick;
#
# - Compute Generator loss based on the updated Discriminator output;
#
# - Update the Generator using Adam (SGD results in a too-slow generator that is outpaced by the Discriminator, whose loss goes to 0 signalling a failure mode)
#
#
# In the above, we specifically train the discriminator first; this is not by chance. This is because if the generator produces an image that looks entirely real to the discriminator, the generator will just get stuck producing that image. This is called [mode collapse](https://developers.google.com/machine-learning/gan/problems). **Training the discriminator first is one way to avoid mode collapse.**
# + id="pvfHxO9ohaox"
def train(training_config, G=None, D=None, epoch=0):
# instantiate tensorboard
writer = SummaryWriter() # (tensorboard) writer will output to ./runs/ directory by default
# need GPU/TPU for training purposes on such a heavy model and dataset
#for TPU: device = device = xm.xla_device()
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
########## instantiate discriminator ##############
# instantiate, set to training mode and move to GPU
discriminator = ConvolutionalDiscriminativeNet().train().to(device)
if not D:
# init weights from normal distribution with mean=0, stdev=0.2 as in Radford et al 2015
# didn't help training much in my experiments
# intiiate discriminator weights for fresh models
discriminator.apply(init_weights)
# instantiate optimizer
D_optimizer = AdamW(discriminator.parameters(), lr=0.002, betas=(0.5,0.999))
else:
# load discriminator weights from checkpoint
discriminator.load_state_dict(D_checkpoint['model_state_dict'])
# instantiate optimizer and load state from checkpoint
D_optimizer = AdamW(discriminator.parameters())
D_optimizer.load_state_dict(D_checkpoint['optimizer_state_dict'])
print(f'Epoch {epoch} checkpoint D model and optimizer state loaded')
########## instantiate generator ###############
# instantiate, set to training mode and move to GPU
generator = ConvolutionalGenerativeNet().train().to(device)
if not G:
# initiate generator weights for fresh models
generator.apply(init_weights)
G_optimizer = AdamW(generator.parameters(), lr=0.0002, betas=(0.5, 0.999))
else:
# load generator weights from discriminator
generator.load_state_dict(G_checkpoint['model_state_dict'])
# instantiate optimizer and load state from checkpoint
G_optimizer = AdamW(generator.parameters())
G_optimizer.load_state_dict(G_checkpoint['optimizer_state_dict'])
print(f'Epoch {epoch} checkpoint G model and optimizer state loaded\n')
# We use binary cross-entropy since we are effectively training the discriminator binary classifier
# loss from the discriminator will be used to train both the discriminator and the generator
# so BCELoss covers loss for both discriminator and generator model training
GAN_loss = nn.BCEWithLogitsLoss()
# We use a single BCELoss function to train the entire GAN using the adversarial loss as in from Radford et al 2015:
# passing real image labels (torch.ones) into BCEloss will return -log(x)
# Using one-sided label smoothing (only on real labels) - do not use on fake images
# purpose is to slow down the discriminator training on real samples, so we don't smooth generator labels
real_image_label = torch.autograd.Variable(torch.Tensor(training_config['batch_size'], 1,1,1).fill_(0.9).type(torch.cuda.HalfTensor))
# unsmoothed labels:
# real_image_label = torch.ones((training_config['batch_size'], 1, 1, 1), device=device)
# passing fake image labels (torch.zeros) into BCEloss will return -log(1-x)
fake_image_label = torch.zeros((training_config['batch_size'], 1, 1, 1), device=device)
# record progress for logging to TensorBoard
# number of debug images generated every so often during training
# keeps track of gradients and training progress/direction
ref_batch_size = 25
# this is just a latent vector (z) of dim (100,) of random Gaussian noise
# used to keep track of progress to generate intermediate/debug imagery
reference_latent_vec = get_gaussian_latent_vectors(num_reference_debug_images, device)
# Store losses for logging to TensorBoard
D_loss_values = []
G_loss_values = []
# Number of generated ebug images
image_count = 0
# measure training time (to get an idea of differences w.r.t. hyperparameters)
start_time = time.time()
# training loop, encompasses training of entire GAN
for epoch in range(epoch, training_config['num_epochs']):
for batch_idx, (real_images, _) in enumerate(dataloader):
# move images to GPU
real_images = real_images.to(device)
######## Train discriminator by maximizing adversarial loss log(D(x)) + log(1-D(G(z))) #######
# Notation: x = real images, z = latent Gaussian vectors
# G(z) = fake images,
# D(G(z)) = probability that fake image is real,
# D(G(x)) = probability that real image is real
# zero out the gradient for each new batch so it doesn't accumulate
D_optimizer.zero_grad()
# get D's predictions on real images D(x)
D_output_real = discriminator(real_images)#.view(-1)
# Store D(x) to track progress
# remember D outputs logits, so take sigmoid activation (scipy.expit)
D_x = expit(D_output_real.mean().item())
# -log(D(x)) is minimized when D(x) = discriminator(real_images) == 1
real_D_loss = GAN_loss(D_output_real, real_image_label)
# get latent vector
z = get_gaussian_latent_vectors(training_config['batch_size'], device)
# get G(z) = fake image generated on latent vector
fake_images = generator(z)
# D(G(z)) = prob that fake image is real
# add call to .detach() to discard gradidents on generator's backward() pass
fake_images_predictions = discriminator(fake_images.detach())
# store prediction on fake images to track progress
# remember D outputs logits, so take sigmoid activation (scipy.expit)
D_G_z = expit(fake_images_predictions.mean().item())
# -log(1 - D(G(z))) is minimized when D(G(z)) = 0
# BCELoss returns -log(1 - D(G(z))) when we pass fake image labels (0) as ground truth
fake_D_loss = GAN_loss(fake_images_predictions, fake_image_label)
# train the discriminator first, before the generator
# using BCELoss
D_loss = real_D_loss + fake_D_loss
## IF USING WGAN ##
# D_loss = -(D_out_real.mean() - D_out_fake.mean())
# compute gradients after discriminator's forward pass
D_loss.backward()
####### this function shows discriminator gradients at the end of the training run for debdugging
# a little costly to run so only enable when debugging
# DO NOT ENABLE FOR FINAL TRAINING LOOP
# plot_grad_flow(discriminator.named_parameters())
# update discriminator weights
D_optimizer.step()
## IF USING WGAN: CLIP WEIGHTS TO [-0.05, 0.05] ##
#for p in discriminator.parameters():
# p.data.clamp(-0.05, 0.05)
############# Train generator by maximizing log(D(G(z))) ###################
# zero out the gradient for each new forward pass so it doesn't accumulate
G_optimizer.zero_grad()
# z = latent vector
z = get_gaussian_latent_vectors(training_config['batch_size'], device)
# pass z through G to get generated (fake) images
generated_images = generator(z)
# get D(G(z)) = prob that fake image is real
generated_images_predictions = discriminator(generated_images)#.view(-1)
# Store fake image prediction after updating D in previous step
# remember D outputs logits, so take sigmoid activation (scipy.expit)
D2_G_z = expit(generated_images_predictions.mean().item())
# -log(D(G(z))) is minimized when D(G(z)) = 1
# BCELoss returns -log(D(G(z))) when we pass real image labels (1) as ground truth
# by passing real image labels for fake (generated) images we configure
# BCELoss to the "log trick"
G_loss = GAN_loss(generated_images_predictions, real_image_label)
## IF USING WGAN ##
# G_loss = -D_out_fake.mean()
# compute gradients after generator's forward pass
G_loss.backward()
# update discriminator weights
G_optimizer.step()
############################ TensorBoard Logging ######################
G_loss_values.append(G_loss.item())
D_loss_values.append(D_loss.item())
# log to TensorBoard
if training_config['enable_tensorboard']:
# write losses for G and D to tensorboard
writer.add_scalars('LOSS: ', {'GEN': G_loss.item(), 'DISC': D_loss.item(), 'DISC_REAL': real_D_loss.item(), 'DISC_FAKE': fake_D_loss.item(), 'D_x': D_x, 'D_G_z': D_G_z, 'D2_G_z':D2_G_z}, len(dataloader) * epoch + batch_idx + 1)
# show generated images in TensorBoard for easy tracking of training progress
if training_config['debug_imagery_log_freq'] is not None and batch_idx % training_config['debug_imagery_log_freq'] == 0:
# don't compute gradients on debug imagery
with torch.no_grad():
# generate the debug imagery on current generator model
log_generated_images = (generator(reference_latent_vec))
# resize generated debug imagery for easier viewing if desired
log_generated_images_resized = nn.Upsample(scale_factor=2, mode='nearest')(log_generated_images)
# make grid of 5x5 generated images to get an idea of G's performance
intermediate_imagery_grid = torchvision.utils.make_grid(log_generated_images, nrow=int(np.sqrt(ref_batch_size)), normalize=True)
writer.add_image('intermediate generated imagery', intermediate_imagery_grid, len(dataloader) * epoch + batch_idx + 1)
# also save images to disk
torchvision.utils.save_image(log_generated_images, os.path.join(training_config['debug_path'], f'{str(image_count).zfill(6)}.jpg'), nrow=int(np.sqrt(ref_batch_size)), normalize=True)
# count of how many image grids have been generated (once for every 'debug_imagery_log_freq' size batch)
image_count += 1
# log to stdodut once for every 'console_log_freq' size batch
if training_config['console_log_freq'] is not None and batch_idx % training_config['console_log_freq'] == 0:
print(f'GAN training: time elapsed= {(time.time() - start_time):.2f} [s] | epoch={epoch + 1} | batch= [{batch_idx + 1}/{len(dataloader)}]')
print(f'G loss: { G_loss.item():.3f}, D loss: {D_loss.item():.3f}, D_REAL: {real_D_loss.item():.3f}, D_FAKE: {fake_D_loss.item():.3f}, \nD_X: {D_x:.3f}, D_G_z: {D_G_z:.3f}, D2_G_z:{D2_G_z:.3f} Debug Image: {str(image_count - 1).zfill(6)}.jpg')
print('\n')
# save a checkpoint of the DCGAN every epoch
# performance often degrades past a certain epoch and may not recover
# save generator
G_checkpoint_name = f'G-DCGAN-{epoch}.pt'
torch.save({
'epoch': epoch,
'model_state_dict': generator.state_dict(),
'optimizer_state_dict': G_optimizer.state_dict(),
'loss': G_loss,
}, './binaries/'+G_checkpoint_name)
# save discriminator
D_checkpoint_name = f'D-DCGAN-{epoch}.pt'
torch.save({
'epoch': epoch,
'model_state_dict': discriminator.state_dict(),
'optimizer_state_dict': D_optimizer.state_dict(),
'loss': D_loss,
}, './binaries/'+D_checkpoint_name)
# + [markdown] id="ABVLzhlP5SBO"
# ## Track Progress with TensorBoard
# Google Colab has magic functions for this to run TensorBoard inline, as long as we've logged to SummaryWriter() correctly. Nice to have so we don't have to scroll through stdout, and we'll also configure it to display the latest generated images so we know when (if ever) we're happy with the DCGAN's results.
# + id="54hwpKMp5Rm5"
# %load_ext tensorboard
# %tensorboard --logdir runs
# + [markdown] id="nJ2aSRRkhao2"
# ## Configure Training Params & Train Model
#
# Training config is mostly for logging purposes. The batch_size was set by the dataloader in the first couple of cells, so make sure to change it there if you want to change it here.
#
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="RJwMnJdve_eR" outputId="7874427a-1ec8-4686-958b-4c5ccdb152cf"
# Track generator output on a reference latent Gaussian vector throughout training
# also allows us to build a nice animation of training progress
# since the intermediate/dedbug generated faces will always be the 'same'
# will make a 5x5 grid of debug images
num_reference_debug_images = 25
# configuration for training; mostly for logging
training_config = {
'num_epochs' : 200, # more than necessary to enable manual stopping
'batch_size' : 32, # lower batch size is better for GAN; 128 used in Radford et al, lower works even better.
'enable_tensorboard' : True,
# save images each time this many batches are processed
'debug_imagery_log_freq' : 200,
# log training progress to stdout each time this many batches are processed
'console_log_freq' : 200,
'debug_path' : './debug_path/'
}
# use pretrained G/D loaded from checkpoint if desired
train(training_config) #G=generator, D=discriminator)
# -
# You get the idea.
# + [markdown] id="BsJ1kJWn_bFR"
# ## Load Pre-Trained Checkpoints to Resume Training
#
# We'll pass the loaded checkpoint to the training loop function and load the model and optimizer state dicts in there (to make sure Colab plays nicely).
# + colab={"base_uri": "https://localhost:8080/", "height": 646} id="CBGCJxBzXT9e" outputId="fe8d3f81-36bc-41b6-a3bf-c7beaa6c849e"
# path to saved generator checkpoint
G = './binaries1/G-DCGAN-11.pt'
# load generator checkpoint
G_checkpoint = torch.load(G)
epoch = G_checkpoint['epoch']
print('G loaded from ',G)
# path to save discriminator checkpoint
D = './binaries1/D-DCGAN-11.pt'
# load discriminator checkpoint
D_checkpoint = torch.load(D)
print('D loaded from ',D)
print('\n')
# note you can change optimizer learning rate before restarting a checkpoint
# using G/D_optimizer.param_groups[0]['lr'] = 0.0001
# but this always leads to mode collapse. I'm not exactly sure why - let me know if you do.
train(training_config, G=G_checkpoint, D=D_checkpoint, epoch=epoch)
# -
# Again, you get the idea.
# + [markdown] id="Yku61L3phao9"
# # Test Model: Generate Images
#
# ## Image Generating Utilities
# We need to process generated images: we rescale back to [0,1] after tanh outputs pixels in [-1,1] and resize to OpenCV's (H,W,C), BGR format.
#
# We'll also make a function to generate and save images from a random latent vector so we can play with the trained generator.
#
# + id="pxo5xBTFhao-"
def process_generated_image(generated_image_tensor):
# Move the tensor from GPU to CPU, convert to numpy array, extract first batch
generated_image = generated_image_tensor.detach().to('cpu').numpy()[0]
# move colour channel from dim 0 to dim 2 (C,H,W) -> (H,W,C)
generated_image = np.moveaxis(generated_image,0, 2)
# Since CelebA images are RGB we don't use this
# if generating grayscale/1 channel images use this to repeat channel 3 times to get RGB image for OpenCV to display
# generated_image = np.repeat(generated_image, 3, axis=2)
# Generator outputs pixel valeus in [-1,1] due to tanh activation on last layer
# transform to [0,1] range for display: add (-1), divide by 2
generated_image -= np.min(generated_image)
generated_image /= np.max(generated_image)
return generated_image
# wrapper to make latent vector z, generate image G(z), and process it for display
def generate_from_random_latent_vector(generator):
# don't compute gradients when just generating images for dedbugging/tracking (saves VRAM)
with torch.no_grad():
# Generate latent vector (z)
latent_vector = get_gaussian_latent_vectors(1, next(generator.parameters()).device)
# Generate image G(z)
image = generator(latent_vector)
# Process the generated image for display (i.e. convert pixel values from [-1,1] to [0,1])
generated_image = process_generated_image(image)
return generated_image
# find next available file name (XXXXXXX.jpg) for generating image
def get_image_ID(input_dir):
# search for present images in xxxxxxx.jpg format and dget next available image name
def valid_frame_name(str):
pattern = re.compile(r'[0-9]{6}\.jpg') # regex, examples it covers: 000000.jpg or 923492.jpg, etc.
return re.fullmatch(pattern, str) is not None
# Filter out only images with xxxxxx.jpg format from the input_dir
valid_frames = list(filter(valid_frame_name, os.listdir(input_dir)))
if len(valid_frames) > 0:
# Images in xxxxxx.jpg format: find the biggest such xxxxxx number and increment by 1
last_img_name = sorted(valid_frames)[-1]
new_prefix = int(last_img_name.split('.')[0]) + 1 # increment by 1
# fill with 0's
return f'{str(new_prefix).zfill(6)}.jpg'
else:
# return first image if dir is empty
return '000000.jpg'
# save generated image to directory
# resolution is 2* 157x128 for easier veiwing by default
def save_image(image_dir, image, resolution=(256, 314), display=False):
# should get numpy array after postprocessing function
assert isinstance(image, np.ndarray), f'Expected numpy array got {type(image)}.'
# get name for image that isn't taken yet
image_name = get_image_ID(image_dir)
# convert to uint8 for OpenCV to display
if image.dtype != np.uint8:
image = (image*255).astype(np.uint8)
# OpenCV expects BGR format, so reverse image's channel dim (from RGB) using [::-1]
cv.imwrite(os.path.join(image_dir, image_name), cv.resize(image[:, :, ::-1], resolution, interpolation=cv.INTER_NEAREST))
image = cv.resize(image, resolution, interpolation=cv.INTER_NEAREST)
# print image to stdout if desired
if display:
plt.imshow(image)
plt.show()
# + [markdown] id="3df-vuJ-hapj"
# # Generated Images and Checkpoint Progression
#
# Now we can check how the generator has progressed through the training epochs.
#
# ### 5 Epochs
# + colab={"base_uri": "https://localhost:8080/", "height": 286} id="dVnt_nhEhapk" outputId="02677edb-4b26-4168-c2a7-3f0b15fddfc1"
# load the desired Generator checkpoint
# select dedsired epoch
epoch = 5
model_path = f'./final_binaries/G-DCGAN-{epoch}.pt'
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
# instantiate model
generator = ConvolutionalGenerativeNet().to(device)
# load model weights from checkpoint
generator.load_state_dict(torch.load(model_path)['model_state_dict'])
# set to validation phase: don't update gradients, no batchnorm in validation phase
generator.eval()
# store generated images here
generated_image_path = './images'
# Generate faces (using DCGAN Generator trained on CelebA)
print('Generating celebrity face')
generated_image = generate_from_random_latent_vector(generator)
save_image(generated_image_path, generated_image, display=True)
# + [markdown] id="vaz4Y_YKtUWv"
# ### 10 Epochs
# + colab={"base_uri": "https://localhost:8080/", "height": 286} id="NCNvdoDDhaqI" outputId="1022fe1a-8b5f-4374-b549-b3ab814527a9"
# load the desired Generator checkpoint
# select dedsired epoch
epoch = 10
model_path = f'./final_binaries/G-DCGAN-{epoch}.pt'
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
# instantiate model
generator = ConvolutionalGenerativeNet().to(device)
# load model weights from checkpoint
generator.load_state_dict(torch.load(model_path)['model_state_dict'])
# set to validation phase: don't update gradients, no batchnorm in validation phase
generator.eval()
# store generated images here
generated_image_path = './images'
# Generate faces (using DCGAN Generator trained on CelebA)
print('Generating celebrity face')
generated_image = generate_from_random_latent_vector(generator)
save_image(generated_image_path, generated_image, display=True)
# + [markdown] id="WXifjtLatWIO"
# ### 25 Epochs
# + colab={"base_uri": "https://localhost:8080/", "height": 286} id="kS2yuV-ZhaqO" outputId="f60dd763-ad90-44fb-a96f-f173580ec4fb"
# load the desired Generator checkpoint
# select dedsired epoch
epoch = 25
model_path = f'./final_binaries/G-DCGAN-{epoch}.pt'
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
# instantiate model
generator = ConvolutionalGenerativeNet().to(device)
# load model weights from checkpoint
generator.load_state_dict(torch.load(model_path)['model_state_dict'])
# set to validation phase: don't update gradients, no batchnorm in validation phase
generator.eval()
# store generated images here
generated_image_path = './images'
# Generate faces (using DCGAN Generator trained on CelebA)
print('Generating celebrity face')
generated_image = generate_from_random_latent_vector(generator)
save_image(generated_image_path, generated_image, display=True)
# + [markdown] id="JbMCxEoUtZjF"
# ### 37 Epochs
# + colab={"base_uri": "https://localhost:8080/", "height": 286} id="ChJM5ZYMtJRy" outputId="673aed60-5b96-48ba-b4b1-bdcd85fc7c16"
# load the desired Generator checkpoint
# select dedsired epoch
epoch = 37
model_path = f'./binaries/G-DCGAN-{epoch}.pt'
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
# instantiate model
generator = ConvolutionalGenerativeNet().to(device)
# load model weights from checkpoint
generator.load_state_dict(torch.load(model_path)['model_state_dict'])
# set to validation phase: don't update gradients, no batchnorm in validation phase
generator.eval()
# store generated images here
generated_image_path = './images'
# Generate faces (using DCGAN Generator trained on CelebA)
print('Generating celebrity face')
generated_image = generate_from_random_latent_vector(generator)
save_image(generated_image_path, generated_image, display=True)
# + [markdown] id="comwPEDs-QZy"
# Now that might fool me even on a good day - it's certainly a palatable human face, perhaps even a little celebrity-like.
# + [markdown] id="Plq-vNyiuHUV"
# # Lessons Learned: How to Stabilize and Optimize a GAN
#
# The difficulty of training the GAN speaks for itself - once you give it a shot. Achieving stability in training and ensuring convergence (of the min-max game between the Generator and Discriminator) to a nash equillibrium is not nearly as easy as it is made out to be by recent papers implementing weight and image regularization techniques such as SELU and Spectral Norm. I've concluded that the plain DCGAN, even with contemporary regularization, doesn't quite cut it. [Progressively Growing GANs](https://arxiv.org/abs/1710.10196) and the [MSG-GAN](https://arxiv.org/abs/1903.06048) are some better candidates for truly impressive results at higher resolutions.
#
# **Here's what I've gathered from trying to get something impressive out of the plain DCGAN architecture:**
# + [markdown] id="IGXhEQTFzOhQ"
# **Training and Failure Modes:**
# - Best results are rarely at the last epoch of training - check intermediate epochs, because the generator's output sometimes degrades before picking back up
# - When the DCGAN is properly tuned, D loss decreases with time and has low variance.
# - When G loss is low or consistently decreases, it's likely fooling D with garbage images.
# - Slowing down D with different learning rates as in the TTUR paper sometimes works, but isn't necessary with proper normalization (SpectralNorm + Noise).
# - We can actually get away with a 5-10x faster learning rate on D when using all the regularization techniques in this notebook: Spectral Norm + Gaussian Instance Noise + Dropout.
# - Slowing down D with altered learning schedule (training D/G in a for loop for each batch) won't avoid training collapse - it'll only delay it.
#
# **Architecture/Hyperparameters:**
# - More filters is not better. 512 is a good maximum. 64 filters in each layer of both G and D works decently as well. Other hyperparams appear more important.
# - Use Dropout ONLY ON D, and not on its final layer. Using dropout on G leads to poor generated images.
# - Use Gaussian Instance Noise ONLY ON D - works well on input or in multiple layers (but not the last layer). Use in G causes it to be unable to generate decent images early on and it never recovers.
# - Adding Gaussian Instance Noise generally made discriminator less sensitive to hyperparameters.
# - Gaussian Noise didn't seem to make a big difference to stability of D when using SpectralNorm + BatchNorm.
# - SpectralNorm is a powerful normalization technique and can be used together with BatchNorm in G.
# - SpectralNorm performed better alone than SpectralNorm + BatchNorm in D. Both improved performance in G.
# - SpectralNorm can replace the normalization power of SELU, working well with leakyReLU.
# - Large batch is much faster to train (1024), but smaller batches train MUCH better GANs.
# - Batch in radford et al (128) works well; smaller batches (64, 32) are more stable but slower to train.
#
# **Activation & Optimizers:**
# - Activation function: In terms of performance for this model, I found LeakyReLU+ReLU to lead to faster equillibrium than SELU, though SELU also produced good generated images.
# - Use BCELossWithLogits and no sigmoid activation on final discriminator layer: More stable than BCELoss() because of log-sum-exp trick.
# - Optimizer: I found AdamW to work better than Adam. AdamW is better than SGD for discriminator, although SGD has been used to slow down the discriminator - it seems SGD slows down D too much with proper discriminator normalization.
# - SELU from HDCGAN paper (SNN): Performed as well as LeakyReLU for this model. SpecNorm+BatchNorm probably means weights are already regularized such that SELU behaves similarly to LeakyReLU.
# - SELU is indeed self normalizing, as it produced decent images even without SpectralNorm and BatchNorm - however, I found it better to have explicit normalization such as SpecNorm and BatchNorm and just use LeakyReLU + ReLU.
#
# **Other tricks:**
# - One-sided label smoothing: set discriminator labels for real images from 1 -> 0.9 to make the only the discriminator's job harder.
# - Popular GANHacks GitHub page incorrectly says to smooth both real and fake labels (1->0.9, 0->0.1).
# - Pixelwise Normalization didn't really change performance, just added complexity. Likely due to Spectral Normalization.
# - Minibatch Standard Deviation didn't help much either. Again, likely has to do with Spectral Normalization working so well.
#
#
# <br>
#
# **Thanks for reading - I sincerely appreciate it. Any comments, errors, or questions, please drop me a line at <EMAIL>.**
#
#
#
# + [markdown] id="4yQzIvdEx2pt"
# # References
# - Arjovsky and Bottou, 2017. Towards Principled Methods for Training Generative Adversarial Networks. https://arxiv.org/abs/1701.04862
# - Arjovsky et al, 2017. Wasserstein GAN. https://arxiv.org/abs/1701.07875
# - Curto et al, 2020. High-resolution Deep Convolutional Generative Adversarial Networks. https://arxiv.org/pdf/1711.06491.pdf
# - Dumoulin and Visin, 2018. A guide to convolution arithmetic for deep learning. https://arxiv.org/pdf/1603.07285.pdf
# - Goodfellow et al, 2014. Generative Adversarial Networks. https://arxiv.org/abs/1406.2661
# - Heusel et al, 2017. GANs Trained by a Two Time-Scale Update Rule Converge to a Local Nash Equilibrium. https://arxiv.org/abs/1706.08500
# - Jenni and Favaro, 2019. On Stabilizing Generative Adversarial Training with Noise. https://arxiv.org/abs/1906.04612
# - Karras et al, 2017. Progressive Growing of GANs for Improved Quality, Stability, and Variation. https://arxiv.org/abs/1710.10196
# - Klambauer et al, 2017. Self-Normalizing Neural Networks. https://arxiv.org/abs/1706.02515
# - Liu et al, 2016. Large-scale CelebFaces Attributes (CelebA) Dataset. http://mmlab.ie.cuhk.edu.hk/projects/CelebA.html
# - Mescheder et al, 2018. Which Training Methods for GANs do actually Converge? https://arxiv.org/pdf/1801.04406.pdf
# - Miyato et al, 2018. Spectral Normalization for Generative Adversarial Networks. https://arxiv.org/abs/1802.05957
# - Radford et al, 2015. Unsupervised Representation Learning with Deep Convolutional Generative Adversarial Networks. https://arxiv.org/abs/1511.06434
# - Salimans et al, 2016. Improved Techniques for Training GANs. https://arxiv.org/pdf/1606.03498.pdf
# - Sonderby et al, 2016. Amortised MAP Inference for Image Super-resolution. https://arxiv.org/abs/1610.04490
#
#
# Notable mention to GanHacks GitHub: https://github.com/soumith/ganhacks/blob/master/README.md
#
# + [markdown] id="e1Y_ZpI88p_N"
# # Appendix A
#
# ## Animating GAN Training Progress
#
# We can use imageio to take all the intermediate debug images dumped during training and write it to a GIF. We'll need to load, resize, and rescale the images out of OpenCV format.
#
# We'll wrap imageio.mimwrite to write a series of images from our debug image path at a chosen framerate, and implement downsampling so we don't end up with a 2GB GIF.
#
#
#
# + id="Z2pskxgv6WRo"
import imageio
# load a single face image from disk, resize if desired
def load_image(img_path, target_shape=None):
# convert openCV BGR images back into RGB
# [:, :, ::-1] converts BGR (images saved in opencv format) into RGB
img = cv.imread(img_path)[:, :, ::-1]
# resize with OpenCV
if target_shape is not None: # resize section
if isinstance(target_shape, int) and target_shape != -1:
current_height, current_width = img.shape[:2]
new_width = target_shape
new_height = int(current_height * (new_width / current_width))
img = cv.resize(img, (new_width, new_height), interpolation=cv.INTER_CUBIC)
else: # set both dimensions to target shape
img = cv.resize(img, (target_shape[1], target_shape[0]), interpolation=cv.INTER_CUBIC)
# normalize to [0,1] range after cv.resize pushes it out of that range
img = img.astype(np.float32) # convert from uint8 to float32
img /= 255.0 # get to [0, 1] range
return img
def create_gif(frames_dir, out_path, downsample=1, img_width=None, fps=60, start_frame = 0, stop_frame=None):
# get images saved as .jpg
frame_paths = [os.path.join(frames_dir, frame_name) for count, frame_name in enumerate(os.listdir(frames_dir)) if count % downsample == 0]
# for resizing image for GIF
if img_width is not None:
for frame_path in frame_paths:
img = load_image(frame_path, target_shape=img_width)
cv.imwrite(frame_path, np.uint8(img[:, :, ::-1] * 255))
# make sure we build the GIF from first to last image to visualize G's learning
frame_paths.sort()
# store list of images from frame_path
images = [imageio.imread(frame_path) for frame_path in frame_paths]
# truncate images at stop frame if provided
if stop_frame:
images = images[start_frame:stop_frame]
# make it into a gif with mimwrite, which writes a series of images to the specified URI (out_path)
imageio.mimwrite(out_path, images, fps=fps)
print(f'\nGIF saved to {out_path} at {fps} FPS.')
# + colab={"base_uri": "https://localhost:8080/"} id="o9aFpsCs8XOW" outputId="f4c18d74-b015-412b-e588-4d9a128fed45"
# debug_path/xxxxxx.jpg contains all intermediate generated training images
create_gif('debug_path_final', 'debug_path_final.gif', downsample=100, fps = 5, stop_frame=400)
#create_gif('debug_path_relu_nospecD', 'debug_path_relu_nospecD.gif', downsample=2, fps = 20)
#create_gif('debug_pathSELU', 'debug_pathSELU.gif', downsample=2, fps = 20)
# + id="1oY6B_WIe91i"
# !unzip -q debug_path_final.zip
# + [markdown] id="4LM-A_mj6oEI"
# And here's a sample of what we get:
#
# <img src="generated_images/training_progress 2.gif">
#
# And here's when things go wrong - this is an example of mode collapse:
#
# <img src="generated_images/mode_collapse.gif">
# + [markdown] id="xRqP4gFh6c9f"
#
# ## Spherical Interpolation
#
# We can save the various latent vectors (_z_) we use to generate images, and then use these vectors to interpolate between them, generating new images. This can help us deduce which features are present in a certain latent vector - but it's mostly for fun.
#
# Spherical interpolation is suggested over linear interpolation for GANs by [GAN Hacks](https://github.com/soumith/ganhacks).
# + id="MqYZe-2l6Di3"
# Suggested to use slerp instead of linear interp for GANs by https://github.com/soumith/ganhacks
# Spherical interpolation formula from https://en.wikipedia.org/wiki/Slerp
def spherical_interpolation(t, p0, p1):
# t is the interpolation parameter in range [0,1]
# p0 and p1 are the two latent vectors
if t <= 0:
return p0
elif t >= 1:
return p1
elif np.allclose(p0, p1):
return p0
# Convert p0 and p1 to unit vectors and find the angle between them (omega)
omega = np.arccos(np.dot(p0 / np.linalg.norm(p0), p1 / np.linalg.norm(p1)))
return np.sin((1.0 - t) * omega) / np.sin(omega) * p0 + np.sin(t * omega) / np.sin(omega) * p1
# generates intermediate interpolated imagery
# between two latent vectors a and b using
def interpolation_gif(generator, a, b, gif_dir='interpolation_result', image_dir='images', fps=5):
# number of images between the vectors a and b, including a and b
interpolation_resolution = 48
# make intermediate image directory and gif directory
os.mkdir(image_dir)
# store intermediate images
generated_imgs = []
for i in range(interpolation_resolution):
# t in range [0,1] i.e. fraction of total interpolation
t = i / (interpolation_resolution - 1)
# generate intermediate interpolated vector
current_latent_vector = spherical_interpolation(t, a, b)
# convert to tensor for compatibility with image processing functions previously defined
current_latent_vector = torch.unsqueeze(torch.tensor(current_latent_vector, device='cuda'), dim=0)
# generate image from latent vector and process for saving
generated_img = process_generated_image(generator(current_latent_vector))#.detach())
# track progress for sanity
print('\r'+f'Image {i+1}/{interpolation_resolution} processed',end='')
# make directory to save intermediate images
# save intermediate interpolated image
save_image(image_dir, generated_img)
# convert from openCV (H,W,C) to (C,H,W) for torchvision.utils.save_image
# make list of interpolated images
generated_imgs.append(torch.tensor(np.moveaxis(generated_img, 2, 0)))
# make a GIF of the interpolation
create_gif(image_dir, gif_dir+'.gif', downsample=1, fps = fps)
# + colab={"base_uri": "https://localhost:8080/"} id="tHCcWb04CT_z" outputId="863ff2ff-c6c7-4c86-fa58-7760c4863e0d"
# load the desired Generator checkpoint
# select dedsired epoch
epoch = 37
model_path = f'./final_binaries/G-DCGAN-{epoch}.pt'
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
# instantiate model
generator = ConvolutionalGenerativeNet().to(device)
# load model weights from checkpoint
generator.load_state_dict(torch.load(model_path)['model_state_dict'])
# set to validation phase: don't compute gradients/update weights, no batchnorm in validation phase
generator.eval()
# generate two random Gaussian latent vectors
z1 = np.random.normal(size=100).astype('Float32')
z2 = np.random.normal(size=100).astype('Float32')
# generate and save interpolated images between z1 and z2
# make a couple of gifs so we can pick through them later
for i in range(5):
# generate two random Gaussian latent vectors
z1 = np.random.normal(size=100).astype('Float32')
z2 = np.random.normal(size=100).astype('Float32')
# create interpolations between z1 and z2, and save to a gif
interpolation_gif(generator, z1, z2, fps=10, image_dir=f'____interpolation-{i}', gif_dir=f'____interpolated_gifs/interp-{i}')
# + [markdown] id="navDUhj4Uyli"
# <img src="generated_images/interpolation/interp-2.gif">
# + [markdown] id="D5VaOc256D15"
# ## CelebA Attributes
# We can make a dictionary to hold the attributes so we can play with the trained DCGAN later on. Attributes from Liu et al's baidu drive 'list_attr_celeba.txt'.
#
# **This will mostly be useful to figure out which feature is which using linear interpolation once our generator is trained; for labelling samples we would use an Auxilliary GAN (AC-GAN) where we embed image labels into an extra tensor dimension.**
# + id="Dj_yzuCSLRPa"
# CelebA dataset attributes
attributes_dict = {
'5_o_Clock_Shadow': torch.bool,
'Arched_Eyebrows': torch.bool,
'Attractive': torch.bool,
'Bags_Under_Eyes': torch.bool,
'Bald': torch.bool,
'Bangs': torch.bool,
'Big_Lips': torch.bool,
'Big_Nose': torch.bool,
'Black_Hair': torch.bool,
'Blond_Hair': torch.bool,
'Blurry': torch.bool,
'Brown_Hair': torch.bool,
'Bushy_Eyebrows': torch.bool,
'Chubby': torch.bool,
'Double_Chin': torch.bool,
'Eyeglasses': torch.bool,
'Goatee': torch.bool,
'Gray_Hair': torch.bool,
'Heavy_Makeup': torch.bool,
'High_Cheekbones': torch.bool,
'Male': torch.bool,
'Mouth_Slightly_Open': torch.bool,
'Mustache': torch.bool,
'Narrow_Eyes': torch.bool,
'No_Beard': torch.bool,
'Oval_Face': torch.bool,
'Pale_Skin': torch.bool,
'Pointy_Nose': torch.bool,
'Receding_Hairline': torch.bool,
'Rosy_Cheeks': torch.bool,
'Sideburns': torch.bool,
'Smiling': torch.bool,
'Straight_Hair': torch.bool,
'Wavy_Hair': torch.bool,
'Wearing_Earrings': torch.bool,
'Wearing_Hat': torch.bool,
'Wearing_Lipstick': torch.bool,
'Wearing_Necklace': torch.bool,
'Wearing_Necktie': torch.bool,
'Young': torch.bool,
}
# + [markdown] id="-2q1ue_CLSj6"
# ## Wasserstein Loss: The WGAN
# In addition to the many normalization techniques above, one way to avoid mode collapse is by using [Wasserstein loss, as first described by Arjovsky et al, 2017](https://arxiv.org/abs/1701.07875) instead of adversarial loss as described by Radford et al. Wasserstein loss measures the distance between two probability distributions (similar to KL divergence, used in sparse autoencoders to ensure training weights are close to 0 in each hidden layer of a network). Roughly speaking, Wasserstein loss computes the quantity (percentage) of the difference between probability distributions, and the distance between the portion of the probability disributions which are different. [Earth mover's distance](https://en.wikipedia.org/wiki/Earth_mover%27s_distance) is another name for Wasserstein loss, and here's why: We imagine two different probability disributions as two differently-shaped dirt piles: the Wasserstein distance (loss) is then the product of the amount of dirt moved and the distance over which it is moved so that the two piles take on the exacft same shape. Wasserstein distance quantifies the minimum energy cost in transforming one probability distribution into the shape of another. **Wasserstein loss ultimately penalizes the generator w.r.t. the distance between the probability distributions of generated training images _P<sub>z</sub>_ and real training images _P<sub>data</sub>_**
|
notebooks/DCGAN_rectangular.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.2 64-bit (''gymenv'': venv)'
# name: python3
# ---
# +
import warnings # python运行代码的时候,经常会碰到代码可以正常运行但是会提出警告,不想看到这些不重要的警告,所以使用控制警告输出
warnings.filterwarnings("ignore") # 使用警告过滤器来控制忽略发出的警告
import pandas as pd
import numpy as np
import matplotlib # python中类似于MATLAB的绘图工具,是一个2D绘图库
import matplotlib.pyplot as plt
import datetime # datetime模块提供了各种类,用于操作日期和时间
# %matplotlib inline
# #%matplotlib inline 表示内嵌绘图,有了这个命令就可以省略掉plt.show()命令了
from finrl.config import config # 引入finrl包的配置
from finrl.marketdata.yahoodownloader import YahooDownloader
from finrl.preprocessing.preprocessors import FeatureEngineer
from finrl.preprocessing.data import data_split
from finrl.env.env_stocktrading import StockTradingEnv
from finrl.model.models import DRLAgent, DRLEnsembleAgent
from finrl.trade.backtest import (
backtest_stats,
get_daily_return,
get_baseline,
backtest_plot,
)
from pprint import pprint # 用于打印 Python 数据结构. 使输出数据格式整齐, 便于阅读
import sys # 该语句告诉Python,我们想要使用sys,此模块包含了与Python解释器和它的环境有关的函数
sys.path.append("../FinRL-Library")
# 在Python执行import sys语句的时候,python会根据sys.path的路径来寻找sys.py模块。
# 添加自己的模块路径, Sys.path.append(“mine module path”)
import itertools # itertools模块中的函数可以用来对数据进行循环操作
# +
import os
if not os.path.exists("./" + config.DATA_SAVE_DIR): # "./"代表当前目录
os.makedirs("./" + config.DATA_SAVE_DIR)
if not os.path.exists("./" + config.TRAINED_MODEL_DIR):
os.makedirs("./" + config.TRAINED_MODEL_DIR)
if not os.path.exists("./" + config.TENSORBOARD_LOG_DIR):
os.makedirs("./" + config.TENSORBOARD_LOG_DIR)
if not os.path.exists("./" + config.RESULTS_DIR):
os.makedirs("./" + config.RESULTS_DIR)
# +
config.START_DATE
config.END_DATE
print(config.SSE_choose30_TICKER)
# 缓存数据,如果日期或者股票列表发生变化,需要删除该缓存文件重新下载
SAVE_PATH = "./datasets/China_SSE_20210101-12h19.csv"
if os.path.exists(SAVE_PATH):
df = pd.read_csv(SAVE_PATH)
else:
df = YahooDownloader(
config.START_DATE, #'2000-01-01',
config.END_DATE, # 2021-01-01,预计将改日期改为'2021-06-20'(今日日期)
ticker_list=config.SSE_choose30_TICKER,
).fetch_data() # SSE_choose30_TICKER上证选择30只股票
df.to_csv(SAVE_PATH)
# +
df.head() # 最开始5条
df.tail() # tail仅展示了最后五条数据
df.shape
df.sort_values(["date", "tic"]).head() # ticker表示股票代码,e.g.AAPL是苹果的股票
tech_indicators = ["macd", "rsi_30", "cci_30", "dx_30"]
fe = FeatureEngineer(
use_technical_indicator=True,
tech_indicator_list=tech_indicators,
use_turbulence=True,
user_defined_feature=False,
)
SAVE_PATH = "./datasets/_China_SSE_20210101-12h19.preprocess.csv"
if os.path.exists(SAVE_PATH):
processed = pd.read_csv(SAVE_PATH)
else:
processed = fe.preprocess_data(df)
processed.to_csv(SAVE_PATH)
list_ticker = processed["tic"].unique().tolist() # 按照processed的"tic"列去重
list_date = list(
pd.date_range(processed["date"].min(), processed["date"].max()).astype(str)
) # 成一个固定频率的时间索引
combination = list(itertools.product(list_date, list_ticker))
processed_full = pd.DataFrame(combination, columns=["date", "tic"]).merge(
processed, on=["date", "tic"], how="left"
)
processed_full = processed_full[processed_full["date"].isin(processed["date"])]
# isin函数,清洗数据,删选过滤掉processed_full中一些行,processed_full新加一列['date']若和processed_full中的['date']不相符合,则被剔除
processed_full = processed_full.sort_values(["date", "tic"])
processed_full = processed_full.fillna(0)
# 对于processed_full数据集中的缺失值使用 0 来填充.
processed_full.sample(5)
# +
stock_dimension = len(processed_full.tic.unique())
state_space = 1 + 2*stock_dimension + len(tech_indicators)*stock_dimension
print(f"Stock Dimension: {stock_dimension}, State Space: {state_space}")
env_kwargs = {
"hmax": 100,
"initial_amount": 1000000,
"buy_cost_pct": 0.001,
"sell_cost_pct": 0.001,
"state_space": state_space,
"stock_dim": stock_dimension,
"tech_indicator_list": tech_indicators,
"action_space": stock_dimension,
"reward_scaling": 1e-4,
"print_verbosity":5
}
rebalance_window = 63 # rebalance_window is the number of days to retrain the model
validation_window = 63 # validation_window is the number of days to do validation and trading (e.g. if validation_window=63, then both validation and trading period will be 63 days)
train_start = '2009-01-01'
train_end = '2015-10-01'
val_test_start = '2015-10-01'
val_test_end = '2020-07-20'
ensemble_agent = DRLEnsembleAgent(df=processed_full,
train_period=(train_start,train_end),
val_test_period=(val_test_start,val_test_end),
rebalance_window=rebalance_window,
validation_window=validation_window,
**env_kwargs)
A2C_model_kwargs = {
'n_steps': 5,
'ent_coef': 0.01,
'learning_rate': 0.0005
}
PPO_model_kwargs = {
"ent_coef":0.01,
"n_steps": 2048,
"learning_rate": 0.00025,
"batch_size": 128
}
DDPG_model_kwargs = {
"action_noise":"ornstein_uhlenbeck",
"buffer_size": 50_000,
"learning_rate": 0.000005,
"batch_size": 128
}
timesteps_dict = {'a2c' : 30_000,
'ppo' : 100_000,
'ddpg' : 10_000
}
timesteps_dict = {'a2c' : 1_000,
'ppo' : 1_000,
'ddpg' : 1_000
}
df_summary = ensemble_agent.run_ensemble_strategy(A2C_model_kwargs,
PPO_model_kwargs,
DDPG_model_kwargs,
timesteps_dict)
# -
df_summary
unique_trade_date = processed_full[(processed_full.date > val_test_start)&(processed_full.date <= val_test_end)].date.unique()
# +
df_trade_date = pd.DataFrame({'datadate':unique_trade_date})
df_account_value=pd.DataFrame()
for i in range(rebalance_window+validation_window, len(unique_trade_date)+1,rebalance_window):
temp = pd.read_csv('results/account_value_trade_{}_{}.csv'.format('ensemble',i))
df_account_value = df_account_value.append(temp,ignore_index=True)
sharpe=(252**0.5)*df_account_value.account_value.pct_change(1).mean()/df_account_value.account_value.pct_change(1).std()
print('Sharpe Ratio: ',sharpe)
df_account_value=df_account_value.join(df_trade_date[validation_window:].reset_index(drop=True))
# -
df_account_value.head()
# %matplotlib inline
df_account_value.account_value.plot()
# +
print("==============Get Backtest Results===========")
now = datetime.datetime.now().strftime('%Y%m%d-%Hh%M')
perf_stats_all = backtest_stats(account_value=df_account_value)
perf_stats_all = pd.DataFrame(perf_stats_all)
# +
#baseline stats
print("==============Get Baseline Stats===========")
baseline_df = get_baseline(
ticker="^DJI",
start = df_account_value.loc[0,'date'],
end = df_account_value.loc[len(df_account_value)-1,'date'])
stats = backtest_stats(baseline_df, value_col_name = 'close')
# +
print("==============Compare to DJIA===========")
# %matplotlib inline
# S&P 500: ^GSPC
# Dow Jones Index: ^DJI
# NASDAQ 100: ^NDX
backtest_plot(df_account_value,
baseline_ticker = '^DJI',
baseline_start = '2019-01-01',
baseline_end = '2021-01-01')
print("==============Compare to DJIA===========")
# %matplotlib inline
# S&P 500: ^GSPC
# Dow Jones Index: ^DJI
# NASDAQ 100: ^NDX
backtest_plot(df_account_value,
baseline_ticker = '^DJI',
baseline_start = df_account_value.loc[0,'date'],
baseline_end = df_account_value.loc[len(df_account_value)-1,'date'])
|
ensemble_Chinadata.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # PS 88 Lab 4: A Warm Fuzzy Theory of Cooperation, with Real Data
# This is the first lab where we will develop some theoretical ideas *and* see how they play out in some data; in this case, the survey that we had you fill out.
#
# As always, let's first import some libraries we will use in the lab
import numpy as np
import matplotlib.pyplot as plt
from datascience import Table
# %matplotlib inline
# ## Part 1: Theory
# One reason that people may not always defect in a prisoners' dilemma situation (whether in the lab or real life) is that they might put weight on considerations beyond just material payoffs.
#
# Here is a simple way to get at this notion. Suppose that our players play a game with the same *material* payoffs as the prisoners' dilemma from lecture:
#
# *Material Payoffs*
#
# | | Cooperate | Defect |
# |----------|----------|--------------|
# | Cooperate | 2,2 | 0,3 |
# | Defect | 3,0 | 1,1 |
#
# Let the players' utility be equal to these material payoffs, except they get an additional $w > 0$ if they both pick cooperate. (Think of this as a "warm fuzzy" feeling for getting the best collective outcome.
#
# **Question 1.1. Write a table which represents this version of the prisoners' dilemma. (Hint: you can double click on this cell and to copy the markdown code for the table above and paste it as a starting point.)**
# *Answer to 1.1 here*
# **Question 1.2. If $w=1/2$, what are the Nash Equilibria to this game?**
# *Answer to 1.2 here*
# **Question 1.3 . If $w=3/2$, what are the Nash Equilibria to this game**?
# *Answer to 1.3 here*
# If one player is uncertain about what strategy the other will use, then we can use the concept of Expected Utility to figure out the best response. For example, the plot below gives the expectued utility to cooperating and defecting as a function of the probability that the other player cooperates.
w = 1/2
p = np.arange(0,1, step=.01)
euc = p*(2 + w) + (1-p)*0
eud = p*3 + (1-p)*1
plt.ticklabel_format(style='plain')
plt.plot(p,euc, label='Cooperating Expected Utility')
plt.plot(p,eud, label='Defecting Expected Utility')
plt.xlabel('Probability of Cooperation')
plt.ylabel('Expected Utility')
plt.legend()
# Note that the EU for defecting (gold) is always above the EU for cooperating (blue). So, for any probability of the other cooperating, it is always better to defect. This should line up with what you found in question 1.1.
# **Question 1.4 Create a similar plot for $w=3/2$**
# Code for 1.4
w = 3/2
p = np.arange(0,1, step=.01)
euc = p*(2 + w) + (1-p)*0
eud = p*3 + (1-p)*1
plt.ticklabel_format(style='plain')
plt.plot(p,euc, label='Cooperating Expected Utility')
plt.plot(p,eud, label='Defecting Expected Utility')
plt.xlabel('Probability of Cooperation')
plt.ylabel('Expected Utility')
plt.legend()
# **Question 1.6. Using the last two graphs, explain why the NE to this game are different when $w=1/2$ and $w=3/2$.**
# *Answer to 1.6*
# **Question 1.7. Now make the same graph where w=3. Compared to the $w=3/2$ case, how does this change the range of $p$ where Cooperating maximizing the player's EU?**
w = 3
p = np.arange(0,1, step=.01)
euc = p*(2 + w) + (1-p)*0
eud = p*3 + (1-p)*1
plt.ticklabel_format(style='plain')
plt.plot(p,euc, label='Cooperating Expected Utility')
plt.plot(p,eud, label='Defecting Expected Utility')
plt.xlabel('Probability of Cooperation')
plt.ylabel('Expected Utility')
plt.legend()
# *Answer to 1.7*
# **Question 1.8. Given the analysis above, suppose this is a reasonable model of how people make decisions in hypothetical prisoners' dilemmas, but individuals vary in both their "warm fuzzy" level $w$ and their optimism about their partner cooperating $p$. Will they be more or less likely to cooperate when $w$ is high? When $p$ is high?**
# *Answer to 1.8*
# ## Part 2: An Empirical Test of the Warm Fuzzy Theory
# Now let's explore how you all played this game in the survey. First we can load up the data with the `read_table` function
classdata = Table.read_table("PS88GTSurvey.csv")
classdata
# Let's say we want to see how often you said you would cooperate in when matched with a random adult in the United States. (Note: there are nicer ways to do this, but let's stick with things that have already been covered in our class and Data 8.)
#
# The following line of code asks whether each of these answers is "Cooperate". The answer to this question is in the `PD_US` column.
classdata.column("PD_US") == "Cooperate"
# We can think of this as the Yes/No answer to whether each person cooperated. We can then count how many said they would cooperate by summing these up, since Python treats 'True' as a 1 and 'False' as a 0.
sum(classdata.column("PD_US") == "Cooperate")
# We might be more interested in the proportion of cooperators. To get this we want to divide by the number of respondents, which we can do with the `num_rows` function applied to the table.
classdata.num_rows
sum(classdata.column("PD_US") == "Cooperate")/classdata.num_rows
# It might be interesting to contrast this with how you al said you would behave with a randomly picked class member, which is storted in the `PD_Class` column.
#
# **Question 2.1. Write a line of code to compute the proportion of respondents who said they would cooperate when matched with a random member of the class.**
sum(classdata.column("PD_Class") == "Cooperate")/classdata.num_rows
# **Question 2.2. Is this more or less than the level of cooperation with a random adult in the United States? Does this result speak to any of the theories of when people cooperate that we discussed in class?**
# *Answer to 2.2*
# Now let's take a look at how you all expect others to behave. The `PD_prC_US` column has your answers to the question about the probability that a random US adult would cooperate. A first cut is to take the average of that.
np.mean(classdata.column("PD_prC_US"))
# Hmm that's weird: probabilities should be between 0 and 1! However, I didn't force this in the survey, I just asked you to put a number between 0 and 1. If you go back to the table you can see that someone didn't follow this instruction. But this is actually a nice teachable moment: we frequently need to "clean" our data before we can analyze it. In this case let's assume anyone who put an answer above 1 entered their answer as a percent. So, we want to take any answer above 1 and divide it by 100.
#
# There are a few ways to do this, here is one that only uses some basic comments. First, we want to identify who answered above 1:
classdata.column("PD_prC_Class") > 1
# What we want to do is take the original answer for any row where this is 'False", and divide by 100 for any row where it is true. The following line of code does this by adding together two terms:
# - `(classdata.column("PD_prC_US") <= 1)*classdata.column("PD_prC_US")`: the first part will be equal to 1 if the original variable is less than or equal to 1 and equal to 0 otherwise. So if the original variable is less than 1, this will return the original variable, and if not we get 0.
# - `(classdata.column("PD_prC_US") > 1)*classdata.column("PD_prC_US")/100`: by a similar logic, if the original variable is 1 this returns the original variable divided by 100, and if it is less than 1 it returns 0.
#
# Combining, one of these is equal to 0, and the other is equal to what we want, and so by adding them we get what we want.
prC_US_cleaned = (classdata.column("PD_prC_US") <= 1)*classdata.column("PD_prC_US") + (classdata.column("PD_prC_US") > 1)*classdata.column("PD_prC_US")/100
prC_US_cleaned
# Now we can compute the average after this cleaning.
np.mean(prC_US_cleaned)
# **Question 2.3. Compare this to the real probability of cooperation with a random US adult. What does this say about the correctness of your expectations, on average?**
# *Answer to 2.3*
# **Question 2.4. Write code to clean the variable for the expected probability of cooperation with a random member of the class, and compare this to the real probability of cooperation.**
# Code for 2.4
prC_Class_cleaned = (classdata.column("PD_prC_Class") <= 1)*classdata.column("PD_prC_Class") + (classdata.column("PD_prC_Class") > 1)*classdata.column("PD_prC_Class")/100
np.mean(prC_Class_cleaned)
# *Words for 2.4*
# We may also want to add the cleaned variable to our Table.
classdata = classdata.with_column("prC_US_cleaned", prC_US_cleaned)
classdata = classdata.with_column("prC_Class_cleaned", prC_Class_cleaned)
classdata
# Now let's start to look at the relationship between the cooperation choice and the expectation that one's partner will cooperate. There are many ways to do this, but a simple one is to compare the average expectation about cooperation among those who cooperated vs defected.
#
# We can do this by first using the `where` function to selection people who cooperated, and then take the average of their expectation about the probability of the partner cooperating. Let's first do this for the question about a random US adult.
np.mean(classdata.where("PD_US", "Cooperate").column("prC_US_cleaned"))
np.mean(classdata.where("PD_US", "Defect").column("prC_US_cleaned"))
# We can also compare the histograms of the two groups using the `hist` function and using a `group="PD_US"` option.
classdata.hist("prC_US_cleaned", group="PD_US")
# **Question 2.5. Are people who choose to cooperate more or less pessimistic about their partner cooperating than those who choose to defect? How does this compare to what the "warm fuzzy" theory of cooperation predicted? (Note: even if you know how to do it, there is no need to do any formal hypothesis testing here, just compare the averages)**
# *Answer to 2.5*
# **Question 2.6. Write code to compare the expected probability of cooperation with a random class member for those who chose to cooperate in this scenario vs those who chose to defect. How does this compare to the "warm fuzzy" theory prediction?**
# Code for 2.6
np.mean(classdata.where("PD_Class", "Cooperate").column("prC_Class_cleaned"))
np.mean(classdata.where("PD_Class", "Defect").column("prC_Class_cleaned"))
classdata.hist("prC_Class_cleaned", group="PD_Class")
# *Words for 2.6*
# The `WarmFuzzy` column contains your answers to the question "On a scale from 0 to 10, how much do you agree with the following statement: "It makes me feel good when a group I am working with sets aside their differences to achieve a common goal"". We might think that this is a reasonable measure of the $w$ variable discussed in the theory section.
#
# **Question 2.7. Write some code to see if people who gave higher answers to this question are more likely to cooperate with members of the class. How does this compare to the predictions of the Warm Fuzzy Theory?**
# Code for 2.7
np.mean(classdata.where("PD_US", "Cooperate").column("WarmFuzzy"))
np.mean(classdata.where("PD_US", "Defect").column("WarmFuzzy"))
np.mean(classdata.where("PD_Class", "Cooperate").column("WarmFuzzy"))
np.mean(classdata.where("PD_Class", "Defect").column("WarmFuzzy"))
# *Words for 2.7*
# **Question 2.8. What do you think might be an important factor determining how people decide whether to cooperate or defect in PD like scenarios? What question(s) could you ask in a survey like this to try and test the importance of this factor?**
# *Words for 2.8*
# ## Section 3. Ultimatums with classmates
# Now lets look at how your behavior in the ultimatum game compares with the theoretical predictions.
#
# **Question 3.1. Make histograms of the offer one would make as a proposer (`Proposer`) or responder (`Responder`)**
# Code for 3.1
classdata.hist("Proposer")
classdata.hist("Responder")
# **Quetion 3.2 How does this compare to the theoretical predictions discussed in lecture?**
# *Words for 3.2*
# A related question we can ask is "Taking the responder behavior as fixed, do proposers make optimal offers?" To figure this out, we can write the expected utility for making offer $x$ as:
# $$
# EU(x) = Pr(x \text{ accepted}) (10-x) + Pr(x \text{ rejected})*0 = Pr(x \text{ accepted}) (10-x)
# $$
#
# Further, we have the information required to compute the probability that an offer is accepted. For each member of the class we know the minimal acceptable offer, which we can use to compute the proportion that would accept a given offer. Let's walk through one way to do this.
#
# First, let's make an array with the possible offers:
offers = np.arange(0, 11)
offers
# If we want to know how many people would accept an offer of, say, 3, we can ask how many set their minimal acceptable offer to 0,1,2, or 3. We can do that with the following code:
sum(classdata.column("Responder") <= 3)
# And we can convert this into a probability by dividing by the number of respondents:
sum(classdata.column("Responder") <= 3)/classdata.num_rows
# **Question 3.3. Write code to compute the probability that an offer of 2 would be *rejected***
# Answer to 3.3
sum(classdata.column("Responder") > 2)/classdata.num_rows
# We would like to know the probability of all possible offers being accepted. We can create an array with these probabilities with a for loop:
paccept = []
for i in offers:
paccept = np.append(paccept, sum(classdata.column("Responder")<= i)/classdata.num_rows)
paccept
# Now let's combine the offer and probability of acceptance into a table.
offertable = Table().with_columns("Offer", offers, "PrAccept", paccept)
offertable
# One interesting thing to lok at is the probability of acceptance as a function of the offer:
offertable.plot("Offer", "PrAccept")
# **Question 3.4. Write code to compute the expected utility to making each offer, add this as a column to the `offertable`, and then plot the expected utility as a function of the offer**
offertable=offertable.with_column("EU", (10-offertable.column("Offer"))*offertable.column("PrAccept"))
offertable
offertable.plot("Offer", "EU")
# **Question 3.5. Compare this to the offers made. Did you all typically choose to make offers that give close to the maximum EU (assuming all you care about is money!)**
# *Words for 3.5*
# There are some other variables from the survey that we haven't looked at yet:
# - `GT` is how much game theory one knew before starting the class
# - `LibCon` is the 1-7 point scale of political views, where 1 is most conservative and 7 is most liberal.
# - `LibConGuess` is the belief about the average `LibCon`.
#
# **Question 3.6. Come up with a quick theory (2-3 sentences) for why one of these variables might be related with the offers made and/or accepted in the ultimatum game. Then write some code to look at this relationship, and describe what you find.**
# *Theory for 3.6*
# +
# Code for 3.6
# -
# *What you found in 3.6*
|
lab/lab4/PS88_lab4_cooperation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 0.3.11
# language: julia
# name: julia-0.3
# ---
using yinsGraph
include("sampler.jl")
include("randTrees.jl")
include("solvers.jl")
a = grid2(30,30,isotropy=10);
@time t = randishKruskal(a)
@time st = comp_stretches(t,a);
sum(st)/nnz(a)
t
@time at = augmentTree(t,a,convert(Int,4*round(sqrt(size(a)[1]))))
la = lap(a)
la[1,1] = la[1,1] + 1
lat = lap(at)
lat[1,1] = lat[1,1] + 1;
@time F = cholfact(lat)
@show nnz(lat)
n = size(la)[1]
b = randn(n);
using IterativeSolvers
@time y = cg(la,b)
norm(b - la*y[1])
y[2]
@time z = cg(la, b, F)
norm(b-la*z[1])
b = randn(n)
b = b / norm(b)
for i in 1:10
b = F \ (la * b)
@show norm(b)
b = b / norm(b)
end
E = eigs(x -> (F \ la * x))
@time fla = cholfact(la);
@time w = fla \ b
norm(b - la*w)
@profile at = augmentTree(t,a,convert(Int,round(sqrt(size(a)[1]))))
# +
function augTreeSolver{Tv,Ti}(ddmat::SparseMatrixCSC{Tv,Ti})
adjmat = -triu(ddmat,1)
adjmat = adjmat + adjmat'
tree = randishKruskal(adjmat)
augtree = augmentTree(tree,adjmat,convert(Int,round(sqrt(size(a)[1]))))
n = size(ddmat)[1]
Dx = spdiagm(ddmat*ones(n))
augDD = Dx + spdiagm(augtree*ones(n)) - augtree
F = cholfact(augDD)
f(b) = cg(ddmat, b, F)
return f
end
# -
f = augTreeSolver(la)
n = size(la)[1]
b = randn(n)
x = f(b)
norm(la*x[1]-b)
y[1]
(x,y) = grid2coords(30);
plotGraph(at,x,y,"red",dots=false)
plotGraph(t,x,y,dots=false)
x, y = grid2coords(50)
plotGraph(t2, x, y, dots = false)
|
notebooks/RandomTrees.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## TensorFlow Intro
# ### Dependencies
#
# ```
# conda create -n tensorflow python=3.5
# source activate tensorflow
# conda install pandas matplotlib jupyter notebook scipy scikit-learn
# pip install tensorflow
# ```
#
# ### Hello, world!
#
# Try running the following code in your Python console to make sure you have TensorFlow properly installed. The console will print "Hello, world!" if TensorFlow is installed. Don’t worry about understanding what it does. You’ll learn about it in the next section.
# +
import tensorflow as tf
# Create TensorFlow object called tensor
hello_constant = tf.constant('Hello World!')
with tf.Session() as sess:
# Run the tf.constant operation in the session
output = sess.run(hello_constant)
print(output)
# -
# ### Tensor
#
# In TensorFlow, data isn’t stored as integers, floats, or strings. These values are encapsulated in an object called a tensor. In the case of hello_constant = tf.constant('Hello World!'), hello_constant is a 0-dimensional string tensor, but tensors come in a variety of sizes as shown below:
# A is a 0-dimensional int32 tensor
A = tf.constant(1234)
# B is a 1-dimensional int32 tensor
B = tf.constant([123,456,789])
# C is a 2-dimensional int32 tensor
C = tf.constant([ [123,456,789], [222,333,444] ])
# tf.constant() is one of many TensorFlow operations you will use in this lesson. The tensor returned by tf.constant() is called a constant tensor, because the value of the tensor never changes.
# ### Session
# TensorFlow’s api is built around the idea of a computational graph, a way of visualizing a mathematical process which you learned about in the MiniFlow lesson. Let’s take the TensorFlow code you ran and turn that into a graph.
# A "TensorFlow Session" is an environment for running a graph. The session is in charge of allocating the operations to GPU(s) and/or CPU(s), including remote machines. Let’s see how you use it.
with tf.Session() as sess:
output = sess.run(hello_constant)
# The code has already created the tensor, hello_constant, from the previous lines. The next step is to evaluate the tensor in a session.
# The code creates a session instance, sess, using tf.Session. The sess.run() function then evaluates the tensor and returns the results.
# ### Input
#
# In the last section, you passed a tensor into a session and it returned the result. What if you want to use a non-constant? This is where tf.placeholder() and feed_dict come into place. In this section, you'll go over the basics of feeding data into TensorFlow.
# ### tf.placeholder()
#
# Sadly you can’t just set x to your dataset and put it in TensorFlow, because over time you'll want your TensorFlow model to take in different datasets with different parameters. You need tf.placeholder()!
#
# tf.placeholder() returns a tensor that gets its value from data passed to the tf.session.run() function, allowing you to set the input right before the session runs.
# ### Session’s feed_dict
# +
x = tf.placeholder(tf.string)
with tf.Session() as sess:
output = sess.run(x, feed_dict={x: 'Hello World'})
# -
# Use the feed_dict parameter in tf.session.run() to set the placeholder tensor. The above example shows the tensor x being set to the string "Hello, world". It's also possible to set more than one tensor using feed_dict as shown below.
# +
x = tf.placeholder(tf.string)
y = tf.placeholder(tf.int32)
z = tf.placeholder(tf.float32)
with tf.Session() as sess:
output = sess.run(x, feed_dict={x: 'Test String', y: 123, z: 45.67})
# -
# Note: If the data passed to the feed_dict doesn’t match the tensor type and can’t be cast into the tensor type, you’ll get the error “ValueError: invalid literal for...”.
# ### Quiz
#
# Let's see how well you understand tf.placeholder() and feed_dict. The code below throws an error, but I want you to make it return the number 123. Change line 11, so that the code returns the number 123.
#
# Note: The quizzes are running TensorFlow version 0.12.1. However, all the code used in this course is compatible with version 1.0. We'll be upgrading our in class quizzes to the newest version in the near future.
# +
import tensorflow as tf
def run():
output = None
x = tf.placeholder(tf.int32)
with tf.Session() as sess:
# TODO: Feed the x tensor 123
output = sess.run(x, feed_dict = {x: 123})
return output
# -
print(run())
# ### TensorFlow Math
# Getting the input is great, but now you need to use it. You're going to use basic math functions that everyone knows and loves - add, subtract, multiply, and divide - with tensors.
# #### Addition
x = tf.add(5, 2) # 7
# You’ll start with the add function. The tf.add() function does exactly what you expect it to do. It takes in two numbers, two tensors, or one of each, and returns their sum as a tensor.
# #### Subtraction and Multiplication
#
# Here’s an example with subtraction and multiplication.
x = tf.subtract(10, 4) # 6
y = tf.multiply(2, 5) # 10
# The x tensor will evaluate to 6, because 10 - 4 = 6. The y tensor will evaluate to 10, because 2 * 5 = 10. That was easy!
# #### Converting types
#
# It may be necessary to convert between types to make certain operators work together. For example, if you tried the following, it would fail with an exception:
tf.subtract(tf.constant(2.0),tf.constant(1))
# Fails with ValueError: Tensor conversion requested dtype float32 for Tensor with dtype int32:
# That's because the constant 1 is an integer but the constant 2.0 is a floating point value and subtract expects them to match.
#
# In cases like these, you can either make sure your data is all of the same type, or you can cast a value to another type. In this case, converting the 2.0 to an integer before subtracting, like so, will give the correct result:
tf.subtract(tf.cast(tf.constant(2.0), tf.int32), tf.constant(1)) # 1
# #### Quiz
#
# Let's apply what you learned to convert an algorithm to TensorFlow. The code below is a simple algorithm using division and subtraction. Convert the following algorithm in regular Python to TensorFlow and print the results of the session. You can use tf.constant() for the values 10, 2, and 1.
# +
# Note: You can't run code in this tab
import tensorflow as tf
# TODO: Convert the following to TensorFlow:
x = 10
y = 2
z = x/y - 1
x = tf.constant(10)
y = tf.constant(2)
z = tf.subtract(tf.divide(x, y), tf.cast(tf.constant(1), tf.float64))
# TODO: Print z from a session
with tf.Session() as sess:
output = sess.run(z)
print(output)
# -
# ### Linear functions in TensorFlow
#
# The most common operation in neural networks is calculating the linear combination of inputs, weights, and biases. As a reminder, we can write the output of the linear operation as:
#
# y = xW + b
#
# Here, W is a matrix of the weights connecting two layers. The output y, the input x, and the biases b are all vectors.
#
# ### Weights and Bias in TensorFlow
#
# The goal of training a neural network is to modify weights and biases to best predict the labels. In order to use weights and bias, you'll need a Tensor that can be modified. This leaves out tf.placeholder() and tf.constant(), since those Tensors can't be modified. This is where tf.Variable class comes in.
# tf.Variable()
x = tf.Variable(5)
# The tf.Variable class creates a tensor with an initial value that can be modified, much like a normal Python variable. This tensor stores its state in the session, so you must initialize the state of the tensor manually. You'll use the tf.global_variables_initializer() function to initialize the state of all the Variable tensors.
init = tf.global_variables_initializer()
with tf.Session() as sess:
sess.run(init)
# The tf.global_variables_initializer() call returns an operation that will initialize all TensorFlow variables from the graph. You call the operation using a session to initialize all the variables as shown above. Using the tf.Variable class allows us to change the weights and bias, but an initial value needs to be chosen.
#
# Initializing the weights with random numbers from a normal distribution is good practice. Randomizing the weights helps the model from becoming stuck in the same place every time you train it. You'll learn more about this in the next lesson, when you study gradient descent.
#
# Similarly, choosing weights from a normal distribution prevents any one weight from overwhelming other weights. You'll use the tf.truncated_normal() function to generate random numbers from a normal distribution.
# #### tf.truncated_normal()
n_features = 120
n_labels = 5
weights = tf.Variable(tf.truncated_normal((n_features, n_labels)))
# The tf.truncated_normal() function returns a tensor with random values from a normal distribution whose magnitude is no more than 2 standard deviations from the mean.
#
# Since the weights are already helping prevent the model from getting stuck, you don't need to randomize the bias. Let's use the simplest solution, setting the bias to 0.
# #### tf.zeros()
n_labels = 5
bias = tf.Variable(tf.zeros(n_labels))
# The tf.zeros() function returns a tensor with all zeros.
# ### Linear Classifier Quiz
# You'll be classifying the handwritten numbers 0, 1, and 2 from the MNIST dataset using TensorFlow. The above is a small sample of the data you'll be training on. Notice how some of the 1s are written with a serif at the top and at different angles. The similarities and differences will play a part in shaping the weights of the model.
# +
# Solution is available in the other "quiz_solution.py" tab
import tensorflow as tf
def get_weights(n_features, n_labels):
"""
Return TensorFlow weights
:param n_features: Number of features
:param n_labels: Number of labels
:return: TensorFlow weights
"""
# TODO: Return weights
weights = tf.Variable(tf.truncated_normal((n_features, n_labels)))
return weights
def get_biases(n_labels):
"""
Return TensorFlow bias
:param n_labels: Number of labels
:return: TensorFlow bias
"""
# TODO: Return biases
bias = tf.Variable(tf.zeros(n_labels))
return bias
def linear(input, w, b):
"""
Return linear function in TensorFlow
:param input: TensorFlow input
:param w: TensorFlow weights
:param b: TensorFlow biases
:return: TensorFlow linear function
"""
# TODO: Linear Function (xW + b)
return tf.add(tf.matmul(input,w), b)
# -
# ### TensorFlow Softmax
#
# You might remember in the Intro to TFLearn lesson we used the softmax function to calculate class probabilities as output from the network. The softmax function squashes it's inputs, typically called logits or logit scores, to be between 0 and 1 and also normalizes the outputs such that they all sum to 1. This means the output of the softmax function is equivalent to a categorical probability distribution. It's the perfect function to use as the output activation for a network predicting multiple classes.
#
# We're using TensorFlow to build neural networks and, appropriately, there's a function for calculating softmax.
x = tf.nn.softmax([2.0, 1.0, 0.2])
# Easy as that! tf.nn.softmax() implements the softmax function for you. It takes in logits and returns softmax activations.
# #### Quiz
#
# Use the softmax function in the quiz below to return the softmax of the logits.
# +
import tensorflow as tf
def run():
output = None
logit_data = [2.0, 1.0, 0.1]
logits = tf.placeholder(tf.float32)
softmax = tf.nn.softmax(logits)
with tf.Session() as sess:
output = sess.run(softmax, feed_dict={logits: logit_data})
return output
# -
# ### Cross Entropy in TensorFlow
#
# In the Intro to TFLearn lesson we discussed using cross entropy as the cost function for classification with one-hot encoded labels. Again, TensorFlow has a function to do the cross entropy calculations for us.
#
# Let's take what you learned from the video and create a cross entropy function in TensorFlow. To create a cross entropy function in TensorFlow, you'll need to use two new functions:
#
# tf.reduce_sum()
# tf.log()
#
# <img src="images/cross-entropy-diagram.png">
# #### Reduce Sum
x = tf.reduce_sum([1, 2, 3, 4, 5]) # 15
# The tf.reduce_sum() function takes an array of numbers and sums them together.
# #### Natural Log
x = tf.log(100.0) # 4.60517
# This function does exactly what you would expect it to do. tf.log() takes the natural log of a number.
# #### Quiz
# Print the cross entropy using softmax_data and one_hot_encod_label.
# +
import tensorflow as tf
softmax_data = [0.7, 0.2, 0.1]
one_hot_data = [1.0, 0.0, 0.0]
softmax = tf.placeholder(tf.float32)
one_hot = tf.placeholder(tf.float32)
# ToDo: Print cross entropy from session
cross_entropy = -tf.reduce_sum(tf.multiply(one_hot, tf.log(softmax)))
with tf.Session() as sess:
print(sess.run(cross_entropy, feed_dict={softmax: softmax_data, one_hot: one_hot_data}))
# -
# ### Mini-batching
#
# In this section, you'll go over what mini-batching is and how to apply it in TensorFlow.
#
# Mini-batching is a technique for training on subsets of the dataset instead of all the data at one time. This provides the ability to train a model, even if a computer lacks the memory to store the entire dataset.
#
# Mini-batching is computationally inefficient, since you can't calculate the loss simultaneously across all samples. However, this is a small price to pay in order to be able to run the model at all.
#
# It's also quite useful combined with SGD. The idea is to randomly shuffle the data at the start of each epoch, then create the mini-batches. For each mini-batch, you train the network weights with gradient descent. Since these batches are random, you're performing SGD with each batch.
#
# Let's look at the MNIST dataset with weights and a bias to see if your machine can handle it#.
# +
from tensorflow.examples.tutorials.mnist import input_data
import tensorflow as tf
n_input = 784 # MNIST data input (img shape: 28*28)
n_classes = 10 # MNIST total classes (0-9 digits)
# Import MNIST data
mnist = input_data.read_data_sets('/datasets/ud730/mnist', one_hot=True)
# The features are already scaled and the data is shuffled
train_features = mnist.train.images
test_features = mnist.test.images
train_labels = mnist.train.labels.astype(np.float32)
test_labels = mnist.test.labels.astype(np.float32)
# Weights & bias
weights = tf.Variable(tf.random_normal([n_input, n_classes]))
bias = tf.Variable(tf.random_normal([n_classes]))
# -
# #### Question 1
#
# Calculate the memory size of train_features, train_labels, weights, and bias in bytes. Ignore memory for overhead, just calculate the memory required for the stored data.
#
# * Single-precision floating-point format is a computer number format that occupies 4 bytes (32 bits) in computer memory
#
# train_features Shape: (55000, 784) Type: float32
#
# train_labels Shape: (55000, 10) Type: float32
#
# weights Shape: (784, 10) Type: float32
#
# bias Shape: (10,) Type: float32
# a) How many bytes of memory does train_features need?
#
# 172480000
#
# b) How many bytes of memory does train_labels need?
#
# 2200000
#
# c) How many bytes of memory does weights need?
#
# 31360
#
# d) How many bytes of memory does bias need?
#
# 40
# The total memory space required for the inputs, weights and bias is around 174 megabytes, which isn't that much memory. You could train this whole dataset on most CPUs and GPUs.
#
# But larger datasets that you'll use in the future measured in gigabytes or more. It's possible to purchase more memory, but it's expensive. A Titan X GPU with 12 GB of memory costs over $1,000.
#
# Instead, in order to run large models on your machine, you'll learn how to use mini-batching.
#
# Let's look at how you implement mini-batching in TensorFlow.
# ### TensorFlow Mini-batching
# In order to use mini-batching, you must first divide your data into batches.
#
# Unfortunately, it's sometimes impossible to divide the data into batches of exactly equal size. For example, imagine you'd like to create batches of 128 samples each from a dataset of 1000 samples. Since 128 does not evenly divide into 1000, you'd wind up with 7 batches of 128 samples, and 1 batch of 104 samples. (7*128 + 1*104 = 1000)
#
# In that case, the size of the batches would vary, so you need to take advantage of TensorFlow's tf.placeholder() function to receive the varying batch sizes.
#
# Continuing the example, if each sample had n_input = 784 features and n_classes = 10 possible labels, the dimensions for features would be [None, n_input] and labels would be [None, n_classes].
# Features and Labels
features = tf.placeholder(tf.float32, [None, n_input])
labels = tf.placeholder(tf.float32, [None, n_classes])
# What does None do here?
#
# The None dimension is a placeholder for the batch size. At runtime, TensorFlow will accept any batch size greater than 0.
#
# Going back to our earlier example, this setup allows you to feed features and labels into the model as either the batches of 128 samples or the single batch of 104 samples.
# #### Question 2
#
# Use the parameters below, how many batches are there, and what is the last batch size?
#
# features is (50000, 400)
#
# labels is (50000, 10)
#
# batch_size is 128
# a) How many batches are there?
#
# b) What is the last batch size?
#
# #### Question 3
#
# Implement the batches function to batch features and labels. The function should return each batch with a maximum size of batch_size. To help you with the quiz, look at the following example output of a working batches function.
# 4 Samples of features
example_features = [
['F11','F12','F13','F14'],
['F21','F22','F23','F24'],
['F31','F32','F33','F34'],
['F41','F42','F43','F44']]
# 4 Samples of labels
example_labels = [
['L11','L12'],
['L21','L22'],
['L31','L32'],
['L41','L42']]
# example_batches = batches(3, example_features, example_labels)
# The example_batches variable would be the following:
[
# 2 batches:
# First is a batch of size 3.
# Second is a batch of size 1
[
# First Batch is size 3
[
# 3 samples of features.
# There are 4 features per sample.
['F11', 'F12', 'F13', 'F14'],
['F21', 'F22', 'F23', 'F24'],
['F31', 'F32', 'F33', 'F34']
], [
# 3 samples of labels.
# There are 2 labels per sample.
['L11', 'L12'],
['L21', 'L22'],
['L31', 'L32']
]
], [
# Second Batch is size 1.
# Since batch size is 3, there is only one sample left from the 4 samples.
[
# 1 sample of features.
['F41', 'F42', 'F43', 'F44']
], [
# 1 sample of labels.
['L41', 'L42']
]
]
]
# Implement the batches function in the "quiz.py" file below.
import math
def batches(batch_size, features, labels):
"""
Create batches of features and labels
:param batch_size: The batch size
:param features: List of features
:param labels: List of labels
:return: Batches of (Features, Labels)
"""
assert len(features) == len(labels)
# TODO: Implement batching
output_batches = []
sample_size = len(features)
for start_i in range(0, sample_size, batch_size):
end_i = start_i + batch_size
batch = [features[start_i:end_i], labels[start_i:end_i]]
output_batches.append(batch)
return output_batches
# Let's use mini-batching to feed batches of MNIST features and labels into a linear model.
#
# Set the batch size and run the optimizer over all the batches with the batches function. The recommended batch size is 128. If you have memory restrictions, feel free to make it smaller.
#
# +
from tensorflow.examples.tutorials.mnist import input_data
import tensorflow as tf
import numpy as np
from helper import batches
learning_rate = 0.001
n_input = 784 # MNIST data input (img shape: 28*28)
n_classes = 10 # MNIST total classes (0-9 digits)
# Import MNIST data
mnist = input_data.read_data_sets('/datasets/ud730/mnist', one_hot=True)
# The features are already scaled and the data is shuffled
train_features = mnist.train.images
test_features = mnist.test.images
train_labels = mnist.train.labels.astype(np.float32)
test_labels = mnist.test.labels.astype(np.float32)
# Features and Labels
features = tf.placeholder(tf.float32, [None, n_input])
labels = tf.placeholder(tf.float32, [None, n_classes])
# Weights & bias
weights = tf.Variable(tf.random_normal([n_input, n_classes]))
bias = tf.Variable(tf.random_normal([n_classes]))
# Logits - xW + b
logits = tf.add(tf.matmul(features, weights), bias)
# Define loss and optimizer
cost = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(logits=logits, labels=labels))
optimizer = tf.train.GradientDescentOptimizer(learning_rate=learning_rate).minimize(cost)
# Calculate accuracy
correct_prediction = tf.equal(tf.argmax(logits, 1), tf.argmax(labels, 1))
accuracy = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
# TODO: Set batch size
batch_size = 128
assert batch_size is not None, 'You must set the batch size'
init = tf.global_variables_initializer()
with tf.Session() as sess:
sess.run(init)
# TODO: Train optimizer on all batches
for batch_features, batch_labels in batches(batch_size, train_features, train_labels):
sess.run(optimizer, feed_dict={features: batch_features, labels: batch_labels})
# Calculate accuracy for test dataset
test_accuracy = sess.run(
accuracy,
feed_dict={features: test_features, labels: test_labels})
print('Test Accuracy: {}'.format(test_accuracy))
# -
# Extracting /datasets/ud730/mnist/train-images-idx3-ubyte.gz Extracting /datasets/ud730/mnist/train-labels-idx1-ubyte.gz Extracting /datasets/ud730/mnist/t10k-images-idx3-ubyte.gz Extracting /datasets/ud730/mnist/t10k-labels-idx1-ubyte.gz
#
# Test Accuracy: 0.11649999767541885
# The accuracy is low, but you probably know that you could train on the dataset more than once. You can train a model using the dataset multiple times. You'll go over this subject in the next section where we talk about "epochs".
# ### Epochs
#
# An epoch is a single forward and backward pass of the whole dataset. This is used to increase the accuracy of the model without requiring more data. This section will cover epochs in TensorFlow and how to choose the right number of epochs.
#
# The following TensorFlow code trains a model using 10 epochs.
# +
from tensorflow.examples.tutorials.mnist import input_data
import tensorflow as tf
import numpy as np
from helper import batches # Helper function created in Mini-batching section
def print_epoch_stats(epoch_i, sess, last_features, last_labels):
"""
Print cost and validation accuracy of an epoch
"""
current_cost = sess.run(
cost,
feed_dict={features: last_features, labels: last_labels})
valid_accuracy = sess.run(
accuracy,
feed_dict={features: valid_features, labels: valid_labels})
print('Epoch: {:<4} - Cost: {:<8.3} Valid Accuracy: {:<5.3}'.format(
epoch_i,
current_cost,
valid_accuracy))
n_input = 784 # MNIST data input (img shape: 28*28)
n_classes = 10 # MNIST total classes (0-9 digits)
# Import MNIST data
mnist = input_data.read_data_sets('/datasets/ud730/mnist', one_hot=True)
# The features are already scaled and the data is shuffled
train_features = mnist.train.images
valid_features = mnist.validation.images
test_features = mnist.test.images
train_labels = mnist.train.labels.astype(np.float32)
valid_labels = mnist.validation.labels.astype(np.float32)
test_labels = mnist.test.labels.astype(np.float32)
# Features and Labels
features = tf.placeholder(tf.float32, [None, n_input])
labels = tf.placeholder(tf.float32, [None, n_classes])
# Weights & bias
weights = tf.Variable(tf.random_normal([n_input, n_classes]))
bias = tf.Variable(tf.random_normal([n_classes]))
# Logits - xW + b
logits = tf.add(tf.matmul(features, weights), bias)
# Define loss and optimizer
learning_rate = tf.placeholder(tf.float32)
cost = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(logits=logits, labels=labels))
optimizer = tf.train.GradientDescentOptimizer(learning_rate=learning_rate).minimize(cost)
# Calculate accuracy
correct_prediction = tf.equal(tf.argmax(logits, 1), tf.argmax(labels, 1))
accuracy = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
init = tf.global_variables_initializer()
batch_size = 128
epochs = 10
learn_rate = 0.001
train_batches = batches(batch_size, train_features, train_labels)
with tf.Session() as sess:
sess.run(init)
# Training cycle
for epoch_i in range(epochs):
# Loop over all batches
for batch_features, batch_labels in train_batches:
train_feed_dict = {
features: batch_features,
labels: batch_labels,
learning_rate: learn_rate}
sess.run(optimizer, feed_dict=train_feed_dict)
# Print cost and validation accuracy of an epoch
print_epoch_stats(epoch_i, sess, batch_features, batch_labels)
# Calculate accuracy for test dataset
test_accuracy = sess.run(
accuracy,
feed_dict={features: test_features, labels: test_labels})
print('Test Accuracy: {}'.format(test_accuracy))
# -
# Running the code will output the following:
# Epoch: 0 - Cost: 11.0 Valid Accuracy: 0.204
#
# Epoch: 1 - Cost: 9.95 Valid Accuracy: 0.229
#
# Epoch: 2 - Cost: 9.18 Valid Accuracy: 0.246
#
# Epoch: 3 - Cost: 8.59 Valid Accuracy: 0.264
#
# Epoch: 4 - Cost: 8.13 Valid Accuracy: 0.283
#
# Epoch: 5 - Cost: 7.77 Valid Accuracy: 0.301
#
# Epoch: 6 - Cost: 7.47 Valid Accuracy: 0.316
#
# Epoch: 7 - Cost: 7.2 Valid Accuracy: 0.328
#
# Epoch: 8 - Cost: 6.96 Valid Accuracy: 0.342
#
# Epoch: 9 - Cost: 6.73 Valid Accuracy: 0.36
#
# Test Accuracy: 0.3801000118255615
# * Each epoch attempts to move to a lower cost, leading to better accuracy.
#
# * This model continues to improve accuracy up to Epoch 9. Let's increase the number of epochs to 100.
# ...
# Epoch: 79 - Cost: 0.111 Valid Accuracy: 0.86
#
# Epoch: 80 - Cost: 0.11 Valid Accuracy: 0.869
#
# Epoch: 81 - Cost: 0.109 Valid Accuracy: 0.869
#
# Epoch: 85 - Cost: 0.107 Valid Accuracy: 0.869
#
# Epoch: 86 - Cost: 0.107 Valid Accuracy: 0.869
#
# Epoch: 87 - Cost: 0.106 Valid Accuracy: 0.869
#
# Epoch: 88 - Cost: 0.106 Valid Accuracy: 0.869
#
# Epoch: 89 - Cost: 0.105 Valid Accuracy: 0.869
#
# Epoch: 90 - Cost: 0.105 Valid Accuracy: 0.869
#
# Epoch: 91 - Cost: 0.104 Valid Accuracy: 0.869
#
# Epoch: 92 - Cost: 0.103 Valid Accuracy: 0.869
#
# Epoch: 93 - Cost: 0.103 Valid Accuracy: 0.869
#
# Epoch: 94 - Cost: 0.102 Valid Accuracy: 0.869
#
# Epoch: 95 - Cost: 0.102 Valid Accuracy: 0.869
#
# Epoch: 96 - Cost: 0.101 Valid Accuracy: 0.869
#
# Epoch: 97 - Cost: 0.101 Valid Accuracy: 0.869
#
# Epoch: 98 - Cost: 0.1 Valid Accuracy: 0.869
#
# Epoch: 99 - Cost: 0.1 Valid Accuracy: 0.869
#
# Test Accuracy: 0.8696000006198883
#
# * From looking at the output above, you can see the model doesn't increase the validation accuracy after epoch 80.
# * Let's see what happens when we increase the learning rate.
#
# learn_rate = 0.1
# Epoch: 76 - Cost: 0.214 Valid Accuracy: 0.752
#
# Epoch: 77 - Cost: 0.21 Valid Accuracy: 0.756
#
# Epoch: 78 - Cost: 0.21 Valid Accuracy: 0.756
#
#
# Epoch: 85 - Cost: 0.207 Valid Accuracy: 0.756
#
# Epoch: 86 - Cost: 0.209 Valid Accuracy: 0.756
#
# Epoch: 87 - Cost: 0.205 Valid Accuracy: 0.756
#
# Epoch: 88 - Cost: 0.208 Valid Accuracy: 0.756
#
# Epoch: 89 - Cost: 0.205 Valid Accuracy: 0.756
#
# Epoch: 90 - Cost: 0.202 Valid Accuracy: 0.756
#
# Epoch: 91 - Cost: 0.207 Valid Accuracy: 0.756
#
# Epoch: 92 - Cost: 0.204 Valid Accuracy: 0.756
#
# Epoch: 93 - Cost: 0.206 Valid Accuracy: 0.756
#
# Epoch: 94 - Cost: 0.202 Valid Accuracy: 0.756
#
# Epoch: 95 - Cost: 0.2974 Valid Accuracy: 0.756
#
# Epoch: 96 - Cost: 0.202 Valid Accuracy: 0.756
#
# Epoch: 97 - Cost: 0.2996 Valid Accuracy: 0.756
#
# Epoch: 98 - Cost: 0.203 Valid Accuracy: 0.756
#
# Epoch: 99 - Cost: 0.2987 Valid Accuracy: 0.756
#
# Test Accuracy: 0.7556000053882599
# * Looks like the learning rate was increased too much. The final accuracy was lower, and it stopped improving earlier.
# * Let's stick with the previous learning rate, but change the number of epochs to 80.
# Epoch: 65 - Cost: 0.122 Valid Accuracy: 0.868
#
# Epoch: 66 - Cost: 0.121 Valid Accuracy: 0.868
#
# Epoch: 67 - Cost: 0.12 Valid Accuracy: 0.868
#
# Epoch: 68 - Cost: 0.119 Valid Accuracy: 0.868
#
# Epoch: 69 - Cost: 0.118 Valid Accuracy: 0.868
#
# Epoch: 70 - Cost: 0.118 Valid Accuracy: 0.868
#
# Epoch: 71 - Cost: 0.117 Valid Accuracy: 0.868
#
# Epoch: 72 - Cost: 0.116 Valid Accuracy: 0.868
#
# Epoch: 73 - Cost: 0.115 Valid Accuracy: 0.868
#
# Epoch: 74 - Cost: 0.115 Valid Accuracy: 0.868
#
# Epoch: 75 - Cost: 0.114 Valid Accuracy: 0.868
#
# Epoch: 76 - Cost: 0.113 Valid Accuracy: 0.868
#
# Epoch: 77 - Cost: 0.113 Valid Accuracy: 0.868
#
# Epoch: 78 - Cost: 0.112 Valid Accuracy: 0.868
#
# Epoch: 79 - Cost: 0.111 Valid Accuracy: 0.868
#
# Epoch: 80 - Cost: 0.111 Valid Accuracy: 0.869
#
# Test Accuracy: 0.86909999418258667
# The accuracy only reached 0.86, but that could be because the learning rate was too high. Lowering the learning rate would require more epochs, but could ultimately achieve better accuracy.
#
# In the upcoming TensorFLow Lab, you'll get the opportunity to choose your own learning rate, epoch count, and batch size to improve the model's accuracy.
|
tensorflow/intro_to_tensorflow.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ! wget http://www.hobieco.com/linked_images/H18-Magnum.jpg
from IPython.display import Image
Image(filename='H18-Magnum.jpg')
# +
from matplotlib.pyplot import imshow
import numpy as np
from PIL import Image
# %matplotlib inline
pil_im = Image.open('H18-Magnum.jpg', 'r')
imshow(np.asarray(pil_im))
# -
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
from ipywidgets import interactive
from IPython.display import Audio, display
import numpy as np
def beat_freq(f1=220.0, f2=224.0):
max_time = 3
rate = 8000
times = np.linspace(0,max_time,rate*max_time)
signal = np.sin(2*np.pi*f1*times) + np.sin(2*np.pi*f2*times)
print(f1, f2, abs(f1-f2))
display(Audio(data=signal, rate=rate))
return signal
v = interactive(beat_freq, f1=(200.0,300.0), f2=(200.0,300.0))
display(v)
v.kwargs
f1, f2 = v.children
f1.value = 255
f2.value = 260
plt.plot(v.result[0:6000])
# +
'''
Make a colorbar as a separate figure.
'''
from matplotlib import pyplot
import matplotlib as mpl
# Make a figure and axes with dimensions as desired.
fig = pyplot.figure(figsize=(8,3))
ax1 = fig.add_axes([0.05, 0.80, 0.9, 0.15])
ax2 = fig.add_axes([0.05, 0.475, 0.9, 0.15])
ax3 = fig.add_axes([0.05, 0.15, 0.9, 0.15])
# Set the colormap and norm to correspond to the data for which
# the colorbar will be used.
cmap = mpl.cm.cool
norm = mpl.colors.Normalize(vmin=5, vmax=10)
# ColorbarBase derives from ScalarMappable and puts a colorbar
# in a specified axes, so it has everything needed for a
# standalone colorbar. There are many more kwargs, but the
# following gives a basic continuous colorbar with ticks
# and labels.
cb1 = mpl.colorbar.ColorbarBase(ax1, cmap=cmap,
norm=norm,
orientation='horizontal')
cb1.set_label('Some Units')
# The second example illustrates the use of a ListedColormap, a
# BoundaryNorm, and extended ends to show the "over" and "under"
# value colors.
cmap = mpl.colors.ListedColormap(['r', 'g', 'b', 'c'])
cmap.set_over('0.25')
cmap.set_under('0.75')
# If a ListedColormap is used, the length of the bounds array must be
# one greater than the length of the color list. The bounds must be
# monotonically increasing.
bounds = [1, 2, 4, 7, 8]
norm = mpl.colors.BoundaryNorm(bounds, cmap.N)
cb2 = mpl.colorbar.ColorbarBase(ax2, cmap=cmap,
norm=norm,
# to use 'extend', you must
# specify two extra boundaries:
boundaries=[0]+bounds+[13],
extend='both',
ticks=bounds, # optional
spacing='proportional',
orientation='horizontal')
cb2.set_label('Discrete intervals, some other units')
# The third example illustrates the use of custom length colorbar
# extensions, used on a colorbar with discrete intervals.
cmap = mpl.colors.ListedColormap([[0., .4, 1.], [0., .8, 1.],
[1., .8, 0.], [1., .4, 0.]])
cmap.set_over((1., 0., 0.))
cmap.set_under((0., 0., 1.))
bounds = [-1., -.5, 0., .5, 1.]
norm = mpl.colors.BoundaryNorm(bounds, cmap.N)
cb3 = mpl.colorbar.ColorbarBase(ax3, cmap=cmap,
norm=norm,
boundaries=[-10]+bounds+[10],
extend='both',
# Make the length of each extension
# the same as the length of the
# interior colors:
extendfrac='auto',
ticks=bounds,
spacing='uniform',
orientation='horizontal')
cb3.set_label('Custom extension lengths, some other units')
pyplot.show()
# +
"""
Show how to make date plots in matplotlib using date tick locators and
formatters. See major_minor_demo1.py for more information on
controlling major and minor ticks
All matplotlib date plotting is done by converting date instances into
days since the 0001-01-01 UTC. The conversion, tick locating and
formatting is done behind the scenes so this is most transparent to
you. The dates module provides several converter functions date2num
and num2date
"""
import datetime
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.dates as mdates
import matplotlib.cbook as cbook
# %matplotlib inline
years = mdates.YearLocator() # every year
months = mdates.MonthLocator() # every month
yearsFmt = mdates.DateFormatter('%Y')
# load a numpy record array from yahoo csv data with fields date,
# open, close, volume, adj_close from the mpl-data/example directory.
# The record array stores python datetime.date as an object array in
# the date column
datafile = cbook.get_sample_data('goog.npy')
r = np.load(datafile).view(np.recarray)
fig, ax = plt.subplots()
ax.plot(r.date, r.adj_close)
# format the ticks
ax.xaxis.set_major_locator(years)
ax.xaxis.set_major_formatter(yearsFmt)
ax.xaxis.set_minor_locator(months)
datemin = datetime.date(r.date.min().year, 1, 1)
datemax = datetime.date(r.date.max().year+1, 1, 1)
ax.set_xlim(datemin, datemax)
# format the coords message box
def price(x): return '$%1.2f'%x
ax.format_xdata = mdates.DateFormatter('%Y-%m-%d')
ax.format_ydata = price
ax.grid(True)
# rotates and right aligns the x labels, and moves the bottom of the
# axes up to make room for them
fig.autofmt_xdate()
plt.show()
# +
'''
Demo to show use of the engineering Formatter.
'''
import matplotlib.pyplot as plt
import numpy as np
# %matplotlib inline
from matplotlib.ticker import EngFormatter
fig, ax = plt.subplots()
ax.set_xscale('log')
formatter = EngFormatter(unit='Hz', places=1)
ax.xaxis.set_major_formatter(formatter)
xs = np.logspace(1, 9, 100)
ys = (0.8 + 0.4 * np.random.uniform(size=100)) * np.log10(xs)**2
ax.plot(xs, ys)
plt.show()
# +
"""
Show how to modify the coordinate formatter to report the image "z"
value of the nearest pixel given x and y
"""
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.cm as cm
X = 10*np.random.rand(5,3)
fig, ax = plt.subplots()
ax.imshow(X, cmap=cm.jet, interpolation='nearest')
numrows, numcols = X.shape
def format_coord(x, y):
col = int(x+0.5)
row = int(y+0.5)
if col>=0 and col<numcols and row>=0 and row<numrows:
z = X[row,col]
return 'x=%1.4f, y=%1.4f, z=%1.4f'%(x, y, z)
else:
return 'x=%1.4f, y=%1.4f'%(x, y)
ax.format_coord = format_coord
plt.show()
# +
"""
Thanks to <NAME> <<EMAIL>> for the logo design
"""
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
import matplotlib.cm as cm
mpl.rcParams['xtick.labelsize'] = 10
mpl.rcParams['ytick.labelsize'] = 12
mpl.rcParams['axes.edgecolor'] = 'gray'
axalpha = 0.05
#figcolor = '#EFEFEF'
figcolor = 'white'
dpi = 80
fig = plt.figure(figsize=(6, 1.1),dpi=dpi)
fig.figurePatch.set_edgecolor(figcolor)
fig.figurePatch.set_facecolor(figcolor)
def add_math_background():
ax = fig.add_axes([0., 0., 1., 1.])
text = []
text.append((r"$W^{3\beta}_{\delta_1 \rho_1 \sigma_2} = U^{3\beta}_{\delta_1 \rho_1} + \frac{1}{8 \pi 2} \int^{\alpha_2}_{\alpha_2} d \alpha^\prime_2 \left[\frac{ U^{2\beta}_{\delta_1 \rho_1} - \alpha^\prime_2U^{1\beta}_{\rho_1 \sigma_2} }{U^{0\beta}_{\rho_1 \sigma_2}}\right]$", (0.7, 0.2), 20))
text.append((r"$\frac{d\rho}{d t} + \rho \vec{v}\cdot\nabla\vec{v} = -\nabla p + \mu\nabla^2 \vec{v} + \rho \vec{g}$",
(0.35, 0.9), 20))
text.append((r"$\int_{-\infty}^\infty e^{-x^2}dx=\sqrt{\pi}$",
(0.15, 0.3), 25))
#text.append((r"$E = mc^2 = \sqrt{{m_0}^2c^4 + p^2c^2}$",
# (0.7, 0.42), 30))
text.append((r"$F_G = G\frac{m_1m_2}{r^2}$",
(0.85, 0.7), 30))
for eq, (x, y), size in text:
ax.text(x, y, eq, ha='center', va='center', color="#11557c", alpha=0.25,
transform=ax.transAxes, fontsize=size)
ax.set_axis_off()
return ax
def add_matplotlib_text(ax):
ax.text(0.95, 0.5, 'matplotlib', color='#11557c', fontsize=65,
ha='right', va='center', alpha=1.0, transform=ax.transAxes)
def add_polar_bar():
ax = fig.add_axes([0.025, 0.075, 0.2, 0.85], polar=True)
ax.axesPatch.set_alpha(axalpha)
ax.set_axisbelow(True)
N = 7
arc = 2. * np.pi
theta = np.arange(0.0, arc, arc/N)
radii = 10 * np.array([0.2, 0.6, 0.8, 0.7, 0.4, 0.5, 0.8])
width = np.pi / 4 * np.array([0.4, 0.4, 0.6, 0.8, 0.2, 0.5, 0.3])
bars = ax.bar(theta, radii, width=width, bottom=0.0)
for r, bar in zip(radii, bars):
bar.set_facecolor(cm.jet(r/10.))
bar.set_alpha(0.6)
for label in ax.get_xticklabels() + ax.get_yticklabels():
label.set_visible(False)
for line in ax.get_ygridlines() + ax.get_xgridlines():
line.set_lw(0.8)
line.set_alpha(0.9)
line.set_ls('-')
line.set_color('0.5')
ax.set_yticks(np.arange(1, 9, 2))
ax.set_rmax(9)
if __name__ == '__main__':
main_axes = add_math_background()
add_polar_bar()
add_matplotlib_text(main_axes)
plt.show()
# +
import matplotlib.path as mpath
import matplotlib.patches as mpatches
import matplotlib.pyplot as plt
Path = mpath.Path
fig, ax = plt.subplots()
pp1 = mpatches.PathPatch(
Path([(0, 0), (1, 0), (1, 1), (0, 0)],
[Path.MOVETO, Path.CURVE3, Path.CURVE3, Path.CLOSEPOLY]),
fc="none", transform=ax.transData)
ax.add_patch(pp1)
ax.plot([0.75], [0.25], "ro")
ax.set_title('The red point should be on the path')
plt.show()
# +
"""
Illustrate some helper functions for shading regions where a logical
mask is True
See :meth:`matplotlib.collections.BrokenBarHCollection.span_where`
"""
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.collections as collections
t = np.arange(0.0, 2, 0.01)
s1 = np.sin(2*np.pi*t)
s2 = 1.2*np.sin(4*np.pi*t)
fig, ax = plt.subplots()
ax.set_title('using span_where')
ax.plot(t, s1, color='black')
ax.axhline(0, color='black', lw=2)
collection = collections.BrokenBarHCollection.span_where(
t, ymin=0, ymax=1, where=s1>0, facecolor='green', alpha=0.5)
ax.add_collection(collection)
collection = collections.BrokenBarHCollection.span_where(
t, ymin=-1, ymax=0, where=s1<0, facecolor='red', alpha=0.5)
ax.add_collection(collection)
plt.show()
# +
"""
Use a Text as a watermark
"""
import numpy as np
#import matplotlib
#matplotlib.use('Agg')
import matplotlib.pyplot as plt
fig, ax = plt.subplots()
ax.plot(np.random.rand(20), '-o', ms=20, lw=2, alpha=0.7, mfc='orange')
ax.grid()
# position bottom right
fig.text(0.95, 0.05, 'Property of MPL',
fontsize=50, color='gray',
ha='right', va='bottom', alpha=0.5)
plt.show()
# +
# a bar plot with errorbars
import numpy as np
import matplotlib.pyplot as plt
N = 5
menMeans = (20, 35, 30, 35, 27)
menStd = (2, 3, 4, 1, 2)
ind = np.arange(N) # the x locations for the groups
width = 0.35 # the width of the bars
fig, ax = plt.subplots()
rects1 = ax.bar(ind, menMeans, width, color='r', yerr=menStd)
womenMeans = (25, 32, 34, 20, 25)
womenStd = (3, 5, 2, 3, 3)
rects2 = ax.bar(ind+width, womenMeans, width, color='y', yerr=womenStd)
# add some text for labels, title and axes ticks
ax.set_ylabel('Scores')
ax.set_title('Scores by group and gender')
ax.set_xticks(ind+width)
ax.set_xticklabels( ('G1', 'G2', 'G3', 'G4', 'G5') )
ax.legend( (rects1[0], rects2[0]), ('Men', 'Women') )
def autolabel(rects):
# attach some text labels
for rect in rects:
height = rect.get_height()
ax.text(rect.get_x()+rect.get_width()/2., 1.05*height, '%d'%int(height),
ha='center', va='bottom')
autolabel(rects1)
autolabel(rects2)
plt.show()
# -
|
PhotoTest.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# IMPORTANT: must add the "%matplotlib notebook" for notebook's matplotlib graphs.
# %matplotlib notebook
# Import numpy, matplotlib, and pandas
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# +
# Load the csv from Resources folder as school_ranking_df
school_ranking_df = pd.read_csv("Resources/cwurDataTop10.csv")
# Show the first 10 rows of school_ranking_df
school_ranking_df.head(10)
# +
# Declare a variable called institution_avg_score_df and set it with the group of institution and the average of score.
# Hint: When grouping, use as_index=False to retain column names.
institution_avg_score_df = school_ranking_df.groupby("institution", as_index=False)["score"].mean()
# Show the first 5 rows of school_ranking_df
institution_avg_score_df.head()
# -
# Set x axis and tick locations
x_axis = np.arange(len(institution_avg_score_df))
tick_locations = [value+0.4 for value in x_axis]
# Create a list indicating where to write x labels and set figure size to adjust for space
# Bars should indicate score
# Horizontal axis should show the institutions
plt.figure(figsize=(10,4))
plt.bar(x_axis, institution_avg_score_df["score"], color='r', alpha=0.5, align="edge")
plt.xticks(tick_locations, institution_avg_score_df["institution"], rotation="vertical")
# Set x and y limits
plt.xlim(-0.25, len(x_axis))
plt.ylim(0, 100)
|
Matplotlib_Code-Drills/day-01/02/.ipynb_checkpoints/Day-01_02_solved-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Preamble
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import warnings
with warnings.catch_warnings():
warnings.simplefilter("ignore")
import scanpy as sc
# -
## local paths etc. You'll want to change these
DATASET_DIR = "/scratch1/rsingh/work/schema/data/tasic-nature"
import sys; sys.path.extend(['/scratch1/rsingh/tools','/afs/csail.mit.edu/u/r/rsingh/work/schema/'])
# #### Import Schema and tSNE
# We use fast-tsne here, but use whatever you like
from fast_tsne import fast_tsne
from schema import SchemaQP
# ### Get example data
# * This data is from Tasic et al. (Nature 2018, DOI: 10.1038/s41586-018-0654-5 )
# * Shell commands to get our copy of the data:
# * wget http://schema.csail.mit.edu/datasets/Schema_demo_Tasic2018.h5ad.gz
# * gunzip Schema_demo_Tasic2018.h5ad.gz
# * The processing of raw data here broadly followed the steps in Kobak & Berens, https://www.biorxiv.org/content/10.1101/453449v1
# * The gene expression data has been count-normalized and log-transformed.
#
adata = sc.read(DATASET_DIR + "/" + "Schema_demo_Tasic2018.h5ad")
# ### Schema examples
# * In all of what follows, the primary dataset is gene expression. The secondary datasets are 1) cluster IDs; and 2) cell-type "class" variables which correspond to superclusters (i.e. higher-level clusters) in the Tasic et al. paper.
# #### Recommendations for parameter settings
# * min_desired_corr and w_max_to_avg are the names for the hyperparameters $s_1$ and $\bar{w}$ from our paper
# * *min_desired_corr*: at first, you should try a range of values for min_desired_corr (e.g., 0.99, 0.90, 0.50). This will give you a sense of what might work well for your data; after this, you can progressively narrow down your range. In typical use-cases, high min_desired_corr values (> 0.80) work best.
# * *w_max_to_avg*: start by keeping this constraint very loose. This ensures that min_desired_corr remains the binding constraint. Later, as you get a better sense for min_desired_corr values, you can experiment with this too. A value of 100 is pretty high and should work well in the beginning.
#
# #### With PCA as change-of-basis, min_desired_corr=0.75, positive correlation with secondary datasets
# +
afx = SchemaQP(0.75) # min_desired_corr is the only required argument.
dx_pca = afx.fit_transform(adata.X, # primary dataset
[adata.obs["class"].values], # one secondary dataset
['categorical'] #it has labels, i.e., is a categorical datatype
)
# -
# #### Similar to above, with NMF as change-of-basis and a different min_desired_corr
# +
afx = SchemaQP(0.6, params= {"decomposition_model": "nmf", "num_top_components": 50})
dx_nmf = afx.fit_transform(adata.X,
[adata.obs["class"].values, adata.obs.cluster_id.values], # two secondary datasets
['categorical', 'categorical'], # both are labels
[10, 1] # relative wts
)
# -
# #### Now let's do something unusual. Perturb the data so it *disagrees* with cluster ids
# +
afx = SchemaQP(0.97, # Notice that we bumped up the min_desired_corr so the perturbation is limited
params = {"decomposition_model": "nmf", "num_top_components": 50})
dx_perturb = afx.fit_transform(adata.X,
[adata.obs.cluster_id.values], # could have used both secondary datasets, but one's fine here
['categorical'],
[-1] # This is key: we are putting a negative wt on the correlation
)
# -
# ### tSNE plots of the baseline and Schema transforms
fig = plt.figure(constrained_layout=True, figsize=(8,2), dpi=300)
tmps = {}
for i,p in enumerate([("Original", adata.X),
("PCA1 (pos corr)", dx_pca),
("NMF (pos corr)", dx_nmf),
("Perturb (neg corr)", dx_perturb)
]):
titlestr, dx1 = p
ax = fig.add_subplot(1,4,i+1, frameon=False)
tmps[titlestr] = dy = fast_tsne(dx1, seed=42)
ax = plt.gca()
ax.set_aspect('equal', adjustable='datalim')
ax.scatter(dy[:,0], dy[:,1], s=1, color=adata.obs['cluster_color'])
ax.set_title(titlestr)
ax.axis("off")
|
examples/Schema_demo.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/mlvlab/COSE474/blob/master/3_Object_Detection_and_MOT_tutorial.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="QX1esx9JMpFR"
# # **Multiple Object Tracking with PyTorch**
#
#
#
# ---
# **Reference**
#
#
# * Object Detection with Faster R-CNN: https://www.learnopencv.com/faster-r-cnn-object-detection-with-pytorch/
# * Simple Online and Realtime Tracking (SORT) algorithm for object ID tracking: https://arxiv.org/abs/1602.00763
#
#
# **Question**
# : What is Multiple Object Tracking?
# - Object tracking is one of the tasks in computer vision, which is detecting an object and searching for that object in a video or a series of images (actually both meaning the same thing).
# - Surveillance cameras in public places for spotting suspicious activities or crimes, and a computer system called 'Hawk-eye' for tracking the trajectory of the ball in various sports are typical examples of applying object tracking in a real life.
#
#
# **Goals**
#
#
# 1. We will use MOT17Det Dataset
# 2. First part: Object Detection with **Faster R-CNN**
# 3. Second part: Multiple Object(ID) Tracking with **Simple Online and Realtime Tracking (SORT)** algorithm
#
#
#
# ---
#
#
#
# + [markdown] colab_type="text" id="hzmQhgb43MMg"
# **0. Preparation**
#
#
# * For your convenience, it is recommended to mount your Google Drive first.
# * Then create extra space for this tutorial in there.
#
#
# ---
#
#
#
#
#
# + id="cnS6Q7M7C2VP" colab_type="code" colab={}
from google.colab import drive
root = '/content/drive/'
drive.mount(root)
# + id="gf8jnmft4v7Q" colab_type="code" colab={}
# Making Directory
import os
from os.path import join
mot = "My Drive/Colab Notebooks/MOT/" # a custom path. you can change if you want to
MOT_PATH = join(root,mot)
# !mkdir "{MOT_PATH}"
# + [markdown] id="1zHl8FW6p1DI" colab_type="text"
#
#
# ---
#
#
# **1. Dataset**
#
#
# * https://motchallenge.net/ : MOT17Det Dataset for Pedestrian Detection Challenge
# * We will only use MOT17-09 dataset for our task.
#
#
# ---
#
#
#
#
#
# + id="O9lkwFuo2CK3" colab_type="code" colab={}
# Download MOT17Det Dataset
# !wget -P "{MOT_PATH}" https://motchallenge.net/data/MOT17Det.zip
# !cd "{MOT_PATH}";unzip MOT17Det.zip
# + id="9oHlCPGp6xZx" colab_type="code" colab={}
# Remove unwanted data for drive volume issue (optional)
# !cd "{MOT_PATH}";rm -rf test
# !cd "{MOT_PATH}";rm -rf train/MOT17-02;rm -rf train/MOT17-04;rm -rf train/MOT17-05
# !cd "{MOT_PATH}";rm -rf train/MOT17-10;rm -rf train/MOT17-11;rm -rf train/MOT17-13
# + id="Ct_3R8Nf-GlT" colab_type="code" colab={}
import sys
motdata = join(MOT_PATH,'train/MOT17-09/img1/')
sys.path.append(motdata)
# + id="_evT8k1S9GQU" colab_type="code" colab={}
# Example: Original picture before detection
import matplotlib.pylab as plt
import cv2
list_motdata = os.listdir(motdata)
list_motdata.sort()
img_ex_path = motdata + list_motdata[0]
img_ex_origin = cv2.imread(img_ex_path)
img_ex = cv2.cvtColor(img_ex_origin, cv2.COLOR_BGR2RGB)
plt.imshow(img_ex)
plt.axis('off')
plt.show()
# + [markdown] id="nvuU_xZjqrJd" colab_type="text"
#
#
# ---
#
#
# **2. Object Detection with Faster R-CNN**
#
# * We will use a pretrained Faster R-CNN model using ResNet50 as a backbone with FPN.
#
#
#
#
#
# ---
#
#
#
# + id="7IEdaeDV6xfZ" colab_type="code" colab={}
# Import required packages/modules first
from PIL import Image
import numpy as np
import torch
import torchvision
from torchvision import transforms as T
# + id="vaFru4rA6xrr" colab_type="code" colab={}
# Download the pretrained Faster R-CNN model from torchvision
model = torchvision.models.detection.fasterrcnn_resnet50_fpn(pretrained=True)
model.eval()
# + id="KBcUWr2W9Tfe" colab_type="code" colab={}
# Define the class names given by PyTorch's official Docs
COCO_INSTANCE_CATEGORY_NAMES = [
'__background__', 'person', 'bicycle', 'car', 'motorcycle', 'airplane', 'bus',
'train', 'truck', 'boat', 'traffic light', 'fire hydrant', 'N/A', 'stop sign',
'parking meter', 'bench', 'bird', 'cat', 'dog', 'horse', 'sheep', 'cow',
'elephant', 'bear', 'zebra', 'giraffe', 'N/A', 'backpack', 'umbrella', 'N/A', 'N/A',
'handbag', 'tie', 'suitcase', 'frisbee', 'skis', 'snowboard', 'sports ball',
'kite', 'baseball bat', 'baseball glove', 'skateboard', 'surfboard', 'tennis racket',
'bottle', 'N/A', 'wine glass', 'cup', 'fork', 'knife', 'spoon', 'bowl',
'banana', 'apple', 'sandwich', 'orange', 'broccoli', 'carrot', 'hot dog', 'pizza',
'donut', 'cake', 'chair', 'couch', 'potted plant', 'bed', 'N/A', 'dining table',
'N/A', 'N/A', 'toilet', 'N/A', 'tv', 'laptop', 'mouse', 'remote', 'keyboard', 'cell phone',
'microwave', 'oven', 'toaster', 'sink', 'refrigerator', 'N/A', 'book',
'clock', 'vase', 'scissors', 'teddy bear', 'hair drier', 'toothbrush'
]
# + id="ZAbS5uwE9vd0" colab_type="code" colab={}
# Defining a function for get a prediction result from the model
def get_prediction(img_path, threshold):
img = Image.open(img_path) # Load the image
transform = T.Compose([T.ToTensor()]) # Defing PyTorch Transform
img = transform(img) # Apply the transform to the image
pred = model([img]) # Pass the image to the model
pred_class = [COCO_INSTANCE_CATEGORY_NAMES[i] for i in list(pred[0]['labels'].numpy())] # Get the Prediction Score
pred_boxes = [[(i[0], i[1]), (i[2], i[3])] for i in list(pred[0]['boxes'].detach().numpy())] # Bounding boxes
pred_score = list(pred[0]['scores'].detach().numpy())
pred_t = [pred_score.index(x) for x in pred_score if x > threshold][-1] # Get list of index with score greater than threshold.
pred_boxes = pred_boxes[:pred_t+1]
pred_class = pred_class[:pred_t+1]
return pred_boxes, pred_class
# + id="Sa4XIuLH6xoa" colab_type="code" colab={}
# Defining a api function for object detection
def object_detection_api(img_path, threshold=0.5, rect_th=3, text_size=1.5, text_th=3):
boxes, pred_cls = get_prediction(img_path, threshold) # Get predictions
img = cv2.imread(img_path) # Read image with cv2
img = cv2.cvtColor(img, cv2.COLOR_BGR2RGB) # Convert to RGB
for i in range(len(boxes)):
cv2.rectangle(img, boxes[i][0], boxes[i][1],color=(0, 255, 0), thickness=rect_th) # Draw Rectangle with the coordinates
cv2.putText(img,pred_cls[i], boxes[i][0], cv2.FONT_HERSHEY_SIMPLEX, text_size, (0,255,0),thickness=text_th) # Write the prediction class
plt.figure(figsize=(15,20)) # display the output image
plt.imshow(img)
plt.xticks([])
plt.yticks([])
plt.show()
# + id="cDlyeXSXHMLo" colab_type="code" colab={}
# Example: After detection
object_detection_api(img_ex_path,threshold=0.8)
# + [markdown] id="6X8gXp0eInzV" colab_type="text"
#
#
#
#
# * The picture above is an example of applying Detection Network (in our case, Faster R-CNN).
# * Since the purpose of dataset we are using is 'tracking', you can see that most of the detected classes are 'person'.
# * We need a prediction result (bbs offset, class label, pred scores) for all the images.
#
#
#
# + [markdown] id="iCelLE4jq8ye" colab_type="text"
#
#
# ---
#
#
# **3. Object ID Tracking with SORT**
#
#
# * Simple Online and Realtime Tracking (SORT) algorithm for object ID tracking
#
# ---
#
#
# + id="j1WuiRXsHMPG" colab_type="code" colab={}
# Git clone: SORT Algorithm
# !cd "{MOT_PATH}";git clone https://github.com/abewley/sort.git
sort = join(MOT_PATH,'sort/')
sys.path.append(sort)
# + id="byjLd9LkKTux" colab_type="code" colab={}
# requirement for sort
# !cd "{sort}";pip install -r requirements.txt
# + id="Rk2pt2SyuH9s" colab_type="code" colab={}
# Optional: if error occurs, you might need to re-install scikit-image and imgaug
# !pip uninstall scikit-image
# !pip uninstall imgaug
# !pip install imgaug
# !pip install -U scikit-image
import skimage
print(skimage.__version__)
# + id="9nfV0kTkMmsd" colab_type="code" colab={}
# Detection information on all the images is well-refined as a json file, which is available at our course git repo
# !cd "{MOT_PATH}";git clone https://github.com/mlvlab/COSE474.git
# + id="zdErbmxk96w1" colab_type="code" colab={}
import json
import collections
from pprint import pprint
from sort import *
jsonpath = join(MOT_PATH,'COSE474/3_MOT_detinfo.json')
with open(jsonpath) as data_file:
data = json.load(data_file)
odata = collections.OrderedDict(sorted(data.items()))
# + id="eONi2h58yKjD" colab_type="code" colab={}
# Let's check out downloaded json file
pprint(odata)
# + [markdown] id="WnsyoUpmz42L" colab_type="text"
#
#
# ---
#
#
#
# * For each image, bbs offset, class label, pred scores are all annotated.
# * Labels are annotated as a number - not a word, and for further information, go to the website below.
# * https://tech.amikelive.com/node-718/what-object-categories-labels-are-in-coco-dataset/
#
#
#
# ---
#
#
# + id="vUB25un4yJsX" colab_type="code" colab={}
img_path = motdata # img root path
# Making new directory for saving results
save_path = join(MOT_PATH,'save/')
# !mkdir "{save_path}"
# + id="_k9vAOQQ960r" colab_type="code" colab={}
mot_tracker = Sort() # Tracker using SORT Algorithm
# + id="MLvfa1Ls964o" colab_type="code" colab={}
for key in odata.keys():
arrlist = []
det_img = cv2.imread(os.path.join(img_path, key))
overlay = det_img.copy()
det_result = data[key]
for info in det_result:
bbox = info['bbox']
labels = info['labels']
scores = info['scores']
templist = bbox+[scores]
if labels == 1: # label 1 is a person in MS COCO Dataset
arrlist.append(templist)
track_bbs_ids = mot_tracker.update(np.array(arrlist))
mot_imgid = key.replace('.jpg','')
newname = save_path + mot_imgid + '_mot.jpg'
print(mot_imgid)
for j in range(track_bbs_ids.shape[0]):
ele = track_bbs_ids[j, :]
x = int(ele[0])
y = int(ele[1])
x2 = int(ele[2])
y2 = int(ele[3])
track_label = str(int(ele[4]))
cv2.rectangle(det_img, (x, y), (x2, y2), (0, 255, 255), 4)
cv2.putText(det_img, '#'+track_label, (x+5, y-10), 0,0.6,(0,255,255),thickness=2)
cv2.imwrite(newname,det_img)
# + [markdown] colab_type="text" id="kyVRJJLMnNGo"
#
# ---
# It's all done!
#
#
# * Finally, you can get a sequence of image with each Tracking ID for every detected person.
# * Check '3_MOT_result.gif' for whole demo experience.
#
#
#
# ---
#
#
#
|
3_Object_Detection_and_MOT_tutorial.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# Chapter 9 of [A guided tour of mathematical methods for the physical sciences](http://www.cambridge.org/nz/academic/subjects/physics/mathematical-methods/guided-tour-mathematical-methods-physical-sciences-3rd-edition#KUoGXYx5FTwytcUg.97) introduces the theorem of Stokes. In Chapter 7, you saw a special vector field, for which the divergence and curl are zero. This gave rise to a model for Hurricane winds and the magnetic field around a wire, captured in one of Maxwell's equations representing the Biot-Savart Law.
# ### Biot-Savart Law
# The current in an long wire creates a magnetic field around it. If we align the wire with the z-axis, the field cannot vary with $z$, or $\phi$, so that ${\bf B}(r,\phi,z) = {\bf B}(r)$. The divergence of magnetic fields, and of incompressible flows, is zero, we found in Chapter 7 that
#
# $$ {\bf B} = \frac{A}{r} \hat{\mathbf \phi}$$
#
# But how do we find the value of $A$? This is where we apply the theorem of Stokes to the Biot-Savart Law:
# $$ \nabla\times{\bf B} = \mu_0{\bf J}.$$
# Integrating this
# over a disk $d{\bf S}$ centered on the wire of radius $r$ in the plane perpendicular to the wire, we have:
# $$ \iint_S \nabla\times{\bf B} \cdot d{\bf S} = \iint_S\mu_0{\bf J}\cdot d{\bf S}.$$
#
# 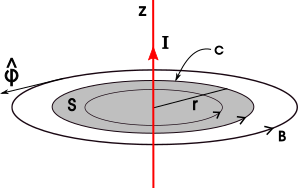
#
# For the left hand side, we apply Stokes Theorem:
# $$ \iint_S \nabla\times{\bf B} \cdot d{\bf S} = \oint_C {\bf B}\cdot d{\bf r} = 2\pi r{\bf B},$$
# because $\bf B$ is constant for each circular path $C$ around the wire.
#
# For the right hand side, we recognize that as long as the radius of the disk $r$ is larger than the thickness of our wire:
# $$\iint_S\mu_0{\bf J}\cdot d{\bf S}= \mu_0 I,$$
# where $I$ is the current. Putting these two results together, we find that the magnetic field is
# $$ {\bf B}(r) = \frac{\mu_0 I}{2\pi r} \hat{\mathbf \phi}.$$ Breaking the radius $r$ and the unit vector $\hat{\phi}$ down in an x- and y-component (see Problem c of Section 7.2 of the book), we define a python function for the three Cartesian components of the magnetic field:
def B(I,x,y):
mu0 = 1.26 * 10**(-6)
r = np.sqrt((x)**2+(y)**2)
c = mu0*I/(2*np.pi)
Bx = -y*c/r**2
By = x*c/r**2
Bz = z*0
return Bx,By,Bz
# Then, we set up a grid of points:
# +
from mpl_toolkits.mplot3d import axes3d
import matplotlib.pyplot as plt
import numpy as np
# %matplotlib notebook
X = np.linspace(-1,1,12)
Y = np.linspace(-1,1,12)
Z = np.linspace(-1,1,12)
x,y,z = np.meshgrid(X,Y,Z)
# -
# The magnetic field on the grid points around the wire with current $I$ is:
I = 200000
Bx,By,Bz = B(I,x,y)
# And finally, we plot of the magnetic field (our first 3D plot with matplotlib!):
fig = plt.figure()
ax = fig.add_subplot(111, projection='3d')
ax.quiver(x,y,z,Bx,By,Bz)
ax.plot([0, 0],[0, 0],[-1,1],linewidth=3,color='r')
plt.xlabel('x')
plt.ylabel('y')
plt.show()
# You can now rotate the graph to look at this field from all angles, and/or change the current. From the top, it should look just like the flow in the python notebook for Chapter 7. What happens when we run the current in the other direction? Chapter 9 concludes with applications of Stokes' Theorem to introduce Lenz' Law, the mysterious quantum mechanical Aharanov-Bohm effect, and the impact of vortices at the tips of sails and wings. Let us now return to the [overview of jupyter notebooks](https://pal.blogs.auckland.ac.nz/2017/12/02/jupyter-notebooks-for-mathematical-methods-in-the-physical-sciences/)!
|
09_Stokes.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 0.3.11
# language: julia
# name: julia-0.3
# ---
addprocs(7)
@everywhere begin
using NetworkDiscovery
using POMDPs
using POMDPToolbox
end
rng=MersenneTwister(1)
nodes = 1000
comms = 5
probes = 30
p_inter = 0.01
p_intra = 0.3
N = 1000
function est_rew(policy, nodes, comms, probes, p_intra, p_inter, N)
sum = @parallel (+) for i in 1:N
prob_rng = MersenneTwister(i)
sim_rng = MersenneTwister(i)
nw = generate_network(rng, nodes, comms, p_intra, p_inter)
pomdp = generate_problem(rng, nw, probes, 1, 100.0, 10, 10, p_intra, p_inter)
revealed = initial_belief(pomdp)
sim = RolloutSimulator(rng=sim_rng, initial_state=nw, initial_belief=revealed)
simulate(sim, pomdp, policy)
end
end
policy = DiscoveryHeuristic(ProbeHighestDegree(true), GuessBasedOnNeighbors(rng))
@time est_rew(policy, nodes, comms, probes, p_intra, p_inter, N)
|
notebooks/Evaluate Heuristic.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # The purpose of this notebook is to test out different data sets in order to decide which suits best to work on
# ### Problem statement
#
# For this project you must create a data set by `simulating a real-world phenomenon of
# your choosing.` You may pick any phenomenon you wish – you might pick one that is
# of interest to you in your personal or professional life. Then, rather than collect data
# related to the phenomenon, you should `model and synthesise such data using Python.`
# We suggest you use the numpy.random package for this purpose.
# Specifically, in this project you should:
#
# * Choose a real-world phenomenon that can be measured and for which you could collect at least one-hundred data points across at least four different variables.
# * Investigate the types of variables involved, their likely distributions, and their relationships with each other.
# * Synthesise/simulate a data set as closely matching their properties as possible.
# * Detail your research and implement the simulation in a Jupyter notebook – the data set itself can simply be displayed in an output cell within the notebook.
#
#
# Note that `this project is about simulation` – you must synthesise a data set. Some
# students may already have some real-world data sets in their own files. It is okay to
# base your synthesised data set on these should you wish (please reference it if you do),
# but the main task in this project is to create a synthesised data set. The next section
# gives an example project idea
#
# -------------------
# # Ideas for project
#
# Corona virus
# * galway specefic
# * compare with national / international.
# * simulate exponential growth if no lockdown?
#
# # Corona Virus
import pandas as pd
data = pd.read_csv("https://opendata-geohive.hub.arcgis.com/datasets/d9be85b30d7748b5b7c09450b8aede63_0.csv?outSR=%7B%22latestWkid%22%3A3857%2C%22wkid%22%3A102100%7D")
data
data.head()
data["CountyName"].value_counts()
galData = data.loc[data["CountyName"] == "Galway"]
galData
df = galData[["TimeStamp", "CountyName", "ConfirmedCovidCases", "ConfirmedCovidDeaths" ]]
df
import matplotlib.pyplot as plt
df.plot(x="TimeStamp", y="ConfirmedCovidCases")
# # Marathon Results
# Marathon results
#
# https://www.kaggle.com/rojour/boston-results
#
# https://towardsdatascience.com/half-marathon-finish-time-prediction-part-1-5807760033eb
#
# https://www.kaggle.com/daniboy370/boston-marathon-2019
#
#
# https://raw.githubusercontent.com/adrian3/Boston-Marathon-Data-Project/master/results2019.csv
# https://github.com/adrian3/Boston-Marathon-Data-Project
marathonData = pd.read_csv("./results2019.csv")
marathonData
marathonData.head()
mdf = marathonData[["gender", "age", "official_time", "place_overall" ]]
mdf
# # Strava Data
#
strava = pd.read_csv("./Strava/activities.csv")
strava
dfStrava = strava[["Activity Date", "Activity Type", "Elapsed Time", "Distance", "Moving Time", "Max Speed","Average Speed","Calories" ]]
dfStrava
StravaData = dfStrava.loc[dfStrava["Activity Type"] == "Run"]
StravaData
x = StravaData["Distance"]
y = StravaData["Activity Date"]
plt.hist(x)
# ## Research
#
# https://data.gov.ie/blog/coronavirus-covid-19
#
# https://data.gov.ie/
#
# https://covid19ireland-geohive.hub.arcgis.com/pages/helpfaqs
# # References
#
# [[1]](www.address.com)
#
# [1] Title; Website; www.address.com
|
old files/Test Notebook.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Orbit Homework
#
# <NAME>
#
# +
# Configure Jupyter so figures appear in the notebook
# %matplotlib inline
# Configure Jupyter to display the assigned value after an assignment
# %config InteractiveShell.ast_node_interactivity='last_expr_or_assign'
# import functions from the modsim.py module
from modsim import *
import numpy as np
print('done')
# -
# ### Earth falling into the sun
#
# Here's a question from the web site [Ask an Astronomer](http://curious.astro.cornell.edu/about-us/39-our-solar-system/the-earth/other-catastrophes/57-how-long-would-it-take-the-earth-to-fall-into-the-sun-intermediate):
#
# "If the Earth suddenly stopped orbiting the Sun, I know eventually it would be pulled in by the Sun's gravity and hit it. How long would it take the Earth to hit the Sun? I imagine it would go slowly at first and then pick up speed."
#
# Here's a solution.
# +
# Here are the units we'll need
s = UNITS.second
N = UNITS.newton
kg = UNITS.kilogram
m = UNITS.meter
# +
# And an inition condition (with everything in SI units)
#r_0 = 147e9 * m
G = 6.674e-11 * N / kg**2 * m**2
m1 = 1.989e30 * kg #mass of sun
m2 = 5.972e24 * kg #mass of earth
t_0 = 0 * s
t_end = 60*60*24*7*52 *s
x_0 = 0 * m
y_0 = 147e9 * m
vx_0 = 30330 * m/s
vy_0 = 0 * m/s
init = State(x = x_0, y = y_0 , vx = vx_0, vy = vy_0)
# +
# Making a system object
r_earth = 6.371e6 * m
r_sun = 695.508e6 * m
system = System(init=init,
G=G,
m1=m1,
r_final=r_sun + r_earth,
m2=m2,
t_0=t_0,
t_end=t_end)
# +
# Here's a function that computes the force of gravity
def universal_gravitation(state, system):
"""Computes gravitational force.
state: State object with distance r
system: System object with m1, m2, and G
"""
x, y, vx, vy = state
unpack(system)
r = Vector(x,y)
angle = r.angle
force = G * m1 * m2 / r.mag**2
fx, fy = pol2cart(angle, force)
Force = Vector(fx,fy)
return Force
# -
test = Vector(1,1)
test.mag
test.angle
universal_gravitation(init, system)
# +
# The slope function
def slope_func(state, t, system):
"""Compute derivatives of the state.
state: position, velocity
t: time
system: System object containing `g`
returns: derivatives of y and v
"""
x, y, vx, vy = state
unpack(system)
v = Vector(vx,vy)
force = universal_gravitation(state, system)
dxdt = vx
dydt = vy
print(v.x)
dvxdt = -force.x / m2
dvydt = -force.y / m2
print(force.x)
return dxdt, dydt, dvxdt, dvydt
# -
def universal_gravitation_2(state,system):
x, y, vx, vy = state
unpack(system)
r = Vector(x,y)
force = (G*m1*m2/r.mag**2)
direction = -r.hat()
forcevec = direction * force
return forcevec
# +
# Always test the slope function!
slope_func(init, 0, system)
# +
# Here's an event function that stops the simulation
# before the collision
def event_func(state, t, system):
x, y, vx, vy = state
r = Vector(x,y)
return r.y - system.r_final
# +
# Always test the event function!
event_func(init, 0, system)
# -
timestep = linspace(t_0, t_end, 200)
# +
# Finally we can run the simulation
results, details = run_ode_solver(system, slope_func, t_eval = timestep )
details
# +
# Here's how long it takes...
t_final = get_last_label(results) * s
# +
# ... expressed in units we understand
t_final.to(UNITS.day)
# +
# Before plotting, we run the simulation again with `t_eval`
'''ts = linspace(t_0, t_final, 201)
results, details = run_ode_solver(system, slope_func, events=event_func, t_eval=ts)
details'''
# +
# Scaling the time steps to days
results.index /= 60 * 60 * 24
# +
# Scaling the distance to million km
x = results.x / 1e9;
y = results.y / 1e9;
# +
# And plotting
plot(x, label='x')
plot(y, label='y')
decorate(xlabel='Time (day)',
ylabel='Distance from sun (million km)')
# +
def plot_trajectory(results):
plot(results.x, results.y, label='trajectory')
decorate(xlabel='x position (m)',
ylabel='y position (m)')
plot_trajectory(results)
savefig('figs/chap10-fig02.pdf')
# -
|
code/orbit_homework.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: model_notebooks
# language: python
# name: model_notebooks
# ---
# +
import pickle
import pandas as pd
import numpy as np
with open('./DF_genres_hashed.pkl', 'rb') as f:
df = pickle.load(f)
# -
df["genres"].value_counts()
df.head()
# # DBSCAN Attempt - Takes too long on local machine - Will try Sagemaker
from sklearn.cluster import DBSCAN
# un-pickling model
with open('DBSCAN_Model.pkl', 'rb') as f:
dbscan = pickle.load(f)
len(dbscan.labels_)
len(set(dbscan.labels_))
dbscan.labels_[1]
np.where(dbscan.labels_ == 2950)[0]
print(len(np.where(dbscan.labels_ == 11)[0]))
for result in np.where(dbscan.labels_ == 11)[0]:
display(pd.DataFrame(df.loc[result][1:2]),pd.DataFrame(df.loc[result][10:11]),pd.DataFrame(df.loc[result][17:18]))
|
model_notebooks/_archive/DBSCAN_test.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys
sys.path.append('/user/ms5941/NLP')
from config import *
from utilities import *
import gensim
import glob
import os
import re
import numpy as np
import pandas as pd
import json
# import logging
# logging.basicConfig(format='%(asctime)s : %(levelname)s : %(message)s', level=logging.INFO)
from gensim.models import ldamodel
# -
THEME = 'Inflation'
with open(TEMP_PATH + '/%s/%s_yearly_slices.txt' % (THEME, THEME), 'r') as f:
yearly_slices = json.load(f)
print(yearly_slices)
dictionary_all = gensim.corpora.Dictionary.load(TEMP_PATH + '/%s/%s_less_restricted.dict' % (THEME, THEME))
corpus_all = gensim.corpora.MmCorpus(TEMP_PATH + '/%s/%s_less_restricted.mm' % (THEME, THEME))
# +
cum_yearly_slices = np.cumsum(yearly_slices)
corpus_by_year = dict()
corpus_by_year[START_YEAR] = corpus_all[:cum_yearly_slices[0]]
for i in range(1, 25):
corpus_by_year[START_YEAR + i] = corpus_all[cum_yearly_slices[i-1]:cum_yearly_slices[i]]
# -
len(corpus_by_year[2009])
avg_topics_all = get_topics(THEME, corpus_all, dictionary_all, corpus_by_year, num_topics=15)
# +
lda = get_model(THEME, corpus_all, dictionary_all, num_topics=15)
topic_word_distribution = lda.get_topics()
df = []
for ind in range(15):
topic = lda.show_topic(ind, topn=15)
topicframe = pd.DataFrame(topic).T
topicframe.index = [ind + 1, ind + 1]
df.append(topicframe)
df = pd.concat(df)
df.columns = ['Word ' + str(i + 1) for i in df.columns]
df.to_csv('LDA_Outputs/%s_Topics_15.csv' % THEME)
yearly_topic_avg_probabilities = pd.DataFrame(avg_topics_all).T
yearly_topic_avg_probabilities.index += 1
yearly_topic_avg_probabilities.columns = [START_YEAR + i for i in range(25)]
yearly_topic_avg_probabilities.to_csv('LDA_Outputs/%s_Average_Topic_Probabilities_Per_Year.csv' % THEME)
# -
|
Regular LDA Runs/Regular LDA - Inflation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import xarray as xr
import cartopy
import cartopy.crs as ccrs
import salem
import pandas as pd
import geopandas as gpd
from oggm import utils
import os, glob
# +
# Get the RGI
rgi_dir = utils.get_rgi_dir(version='62')
fs = list(sorted(glob.glob(rgi_dir + "/*/*_rgi6*_*.shp")))[2:]
out = []
for f in fs:
sh = gpd.read_file(f).set_index('RGIId')
del sh['geometry']
out.append(pd.DataFrame(sh))
mdf = pd.concat(out)
mdf['O1Region'] = ['{:02d}'.format(int(i)) for i in mdf['O1Region']]
mdf['O2Region'] = ['{:02d}'.format(int(i)) for i in mdf['O2Region']]
# Read glacier attrs
gtkeys = {0: 'Glacier',
1: 'Ice cap',
2: 'Perennial snowfield',
3: 'Seasonal snowfield',
9: 'Not assigned',
}
ttkeys = {0: 'Land-terminating',
1: 'Marine-terminating',
2: 'Lake-terminating',
3: 'Dry calving',
4: 'Regenerated',
5: 'Shelf-terminating',
9: 'Not assigned',
}
stkeys = {0: 'Glacier or ice cap',
1: 'Glacier complex',
2: 'Nominal glacier',
9: 'Not assigned',
}
mdf['GlacierType'] = [gtkeys[g] for g in mdf.Form]
mdf['TerminusType'] = [ttkeys[g] for g in mdf.TermType]
mdf['GlacierStatus'] = [stkeys[g] for g in mdf.Status]
mdf['IsTidewater'] = [ttype in ['Marine-terminating', 'Lake-terminating'] for ttype in mdf.TerminusType]
mdf['IsNominal'] = [stype == 'Nominal glacier' for stype in mdf.GlacierStatus]
# -
mdf = mdf.drop(['check_geom'], axis=1)
for i, d in mdf.iterrows():
assert i[6:8] == d['O1Region']
mdf.to_hdf('rgi62_stats.h5', key='df', mode='w', complevel=5)
df = pd.read_hdf('rgi62_stats.h5')
df.columns
df = df.loc[df['Connect'] != 2]
df['rgi_year'] = [int(s[0:4]) for s in df.BgnDate]
len(df.loc[df['rgi_year'] < 0])
df = df.loc[df['rgi_year'] > 0]
df['rgi_year'].min()
df.groupby('O1Region').median()['rgi_year']
|
prepare_rgi_summary_table.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ## Decision Tree
# ---
#
# ### Purity
# - **Target:** get pure subset
# - Can tell us not only prediction but also confidence on prediction
#
# ### Algorithm
#
# - **Split(node,{examples}):**
# 1. A <- the best attribute for splitting the {examples}
# 2. Decision attribute for this node <- A
# 3. For each value of A, create new child node
# 4. Spliting training {examples} to child nodes
# 5. For each child node/subset:<br>
# if subset is pure: STOP<br>
# else:Split(child_node,{subset})
# - **<NAME>(ID3:1986),(C4.5:1993)**
# - **Breimanetal(CaRT:1984) from statistics**
# <div class="alert alert-block alert-warning">
# <b>Which attribute to split on?</b>
# </div>
#
# - Want to measure "purity" of the split
# - more certain after about Yes/No after the split
# - pure set(__<font color='red'>4 yes</font>__ / __<font color='blue'>0 no</font>__)=>completely certain(100%)
# - inpure(__<font color='red'>3 yes</font> / <font color='red'>3 no</font>__)=>completely uncertain(50%)
# - can't use __P("yes"|set)__:
# - must be symmetric: 4 yes / 0 no as pure as 0 yes / 4 no
#
# <div class="alert alert-block alert-info">
# <b>Entropy</b>
# </div>
#
# > A way to measure uncertainty of the class in a subset of examples
#
# $$H(s) = -p_{(+)}log_2{p_{(+)}} - p_{(-)}log_2{p_{(-)}}$$
# - Interpretation: assume item X belongs to S
# - how many bits need to tell if X positive or negtive
# - impure(3 yes / 3 no)
#
# $H(s) = - \sub{3,5}$
#
# <div class="alert alert-block alert-danger">
# <b>Information Gain</b>
# </div>
#
# > ID3
#
# - Want many iterms in pure sets
# - Expected drop in entropy after split (**<font color='red'>Expected Entropy,EH</font>**)
# $$Gain(S,A)=H(S) - \sum_{V}$$
# - Mutual Information
# - between attribute A and class labels of S
#
# <div class="alert alert-block alert-danger">
# <b>Information gain ratio</b>
# </div>
#
# > ID4.5
#
#
# <div class="alert alert-block alert-danger">
# <b>Gini Index</b>
# </div>
#
# > CART, Classification and Regression Trees
#
# $$Gini(A)=1-\sum_{i=1}^{C}p_i^2$$
# $$Gini_{split} = \sum{{N_i \over N} Gini(T_i)}$$
#
# ### Gini
#
# #### Defination of Gini Index
# $$Gini(p)=\sum_{k=1}^{K}{p_k(1-p_k)}=1-\sum_{k=1}^{K}p_k^2$$
#
# #### Gini Index of Sample Set D
# $$Gini(D)=1-\sum_{k=1}^{K}{\big({|C_k|\over|D|}\big)^2}$$
#
# #### Gain Gini
# $$Gain\_Gini(D,A)={|D_1|\over|D|}Gini(D_1)+{|D_2|\over|D|}Gini(D_2)$$
#
# #### Split Attribute
# $$\min_{i \in A}(Gain_Gini(D,A))$$
#
# #### Split Point
# $$\min_{A \in Attribute}(\min_{i \in A}(Gain\_Gini(D,A_i))$$
# ### <font color='red'>How to deal with continuous attributes? </font>
# ### CART
# #### CART methodology
# - Binary Split
# - Split Based on One Variable
# - Estimation the misclassification rate
# - Pruning procedure
# #### Tree growing procedure
# - Splitting strategy
# - Continuous or numerial variable
#
# #### Pipline
#
# - 选择最优切分变量j与切分点s
# - 用选用的(j,s)对,划分区域并决定相应的输出值
# - 继续对两个子区域调用上述步骤,将输入空间分为M个区域R1,R2,...,Rm,生成决策树。
# - 当输入空间划分确定时,用平方误差来表示回归树对训练数据的预测方法,用平方误差最小的准则求解每个单元的最优输出值。
#
# #### Pruning procedure
# > <font color='red'><b>分为两部分,分别是生成子树序列和交叉验证</b></font>
#
# $$C_\alpha(T)=C(T)+\alpha(T)$$
#
# - T 为任意树,|T|为树T的叶节点个数
# - $\alpha$是参数,权衡拟合程度与树的复杂度
# - C|T|为预测误差,可以是平方误差也可以是基尼指数,C|T|衡量训练数据的拟合程度
# ### Guassian information gain to decide splits
# ### Overfitting
# ### Pros and Cons
#
# - Cons
# - only axis-aligned splits of data
# - greedy(may not find best tree
# ### Summary
# ### Realize with Sklearn
# +
import sklearn
from sklearn.datasets import load_iris
import pandas as pd
import graphviz
from sklearn import tree
# -
iris = load_iris()
# +
X = iris.data
y = iris.target
clf = tree.DecisionTreeClassifier()
# -
clf = clf.fit(X,y)
dot_data = tree.export_graphviz(clf, out_file=None, feature_names=iris.feature_names, class_names=iris.target_names,
filled=True, rounded=True, special_characters=True)
graph = graphviz.Source(dot_data)
graph.view()
|
Decision Tree & Random Forest.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.5 64-bit
# metadata:
# interpreter:
# hash: aee8b7b246df8f9039afb4144a1f6fd8d2ca17a180786b69acc140d282b71a49
# name: Python 3.8.5 64-bit
# ---
# +
# Imports
import torch
import torchvision
from torchvision.datasets import MNIST
import torchvision.transforms as transforms
from torch.utils.data import random_split
from torch.utils.data import DataLoader
import torch.nn as nn
import torch.nn.functional as F
import matplotlib.pyplot as plt
import jovian
# + tags=[]
dataset = MNIST(root='data/', download=True, train=True, transform=transforms.ToTensor())
# + tags=[]
img_tensor, label = dataset[0]
print(img_tensor.shape, label)
# -
plt.imshow(img_tensor[0, 0:24, 13:25], cmap='gray') #needs 1st part as 0, channel not expected by imshow or is the last dimension
# +
train_ds, val_ds = random_split(dataset, [50000, 10000])
batch_size = 128
train_loader = DataLoader(train_ds, batch_size, shuffle=True)
val_loader = DataLoader(val_ds, batch_size, shuffle=True)
# -
def accuracy(outputs, labels):
_, preds = torch.max(outputs, dim=1)
return torch.tensor(torch.sum(preds == labels).item() / len(preds))
# +
input_size = 28*28
num_classes = 10
class MnistModel(nn.Module):
def __init__(self, input_size, hidden_size, output_size):
super().__init__()
self.linear1 = nn.Linear(input_size, hidden_size)
#output layer
self.linear2 = nn.Linear(hidden_size, output_size)
def forward(self, xb):
xb = xb.reshape(-1, input_size)
out = self.linear1(xb)
out = F.relu(out)
out = self.linear2(out)
return out
def training_step(self, batch):
images, labels = batch
out = self(images)
loss = F.cross_entropy(out, labels)
return loss
def validation_step(self, batch):
images, labels = batch
out = self(images)
loss = F.cross_entropy(out, labels)
acc = accuracy(out, labels)
return {"val_loss": loss, "val_acc": acc}
def validation_epoch_end(self, outputs):
batch_losses = [x['val_loss'] for x in outputs]
epoch_loss = torch.stack(batch_losses).mean()
batch_accuracies = [x['val_acc'] for x in outputs]
epoch_accuracy = torch.stack(batch_accuracies).mean()
return {'val_loss': epoch_loss.item(), 'val_acc': epoch_accuracy.item()}
def epoch_end(self, epoch, result):
print('Epoch {0}: Validation Loss: {1}, Validation Accuracy: {2}'.format(epoch, result['val_loss'], result['val_acc']))
# -
def evaluate(model, val_loader):
val_outs = [model.validation_step(val_batch) for val_batch in val_loader]
val_result = model.validation_epoch_end(val_outs)
return val_result
# + tags=[]
def fit(epochs, lr, model, train_loader, val_loader, opt_func=torch.optim.SGD):
optimizer = opt_func(model.parameters(), lr)
history = []
for epoch in range(epochs):
for batch in train_loader:
loss = model.training_step(batch)
loss.backward()
optimizer.step()
optimizer.zero_grad()
val_result = evaluate(model, val_loader)
model.epoch_end(epoch, val_result)
history.append(val_result)
return history
# -
def get_default_device():
if torch.cuda.is_available():
return torch.device('cuda')
else:
return torch.device('cpu')
device = get_default_device()
device
def to_device(data, device):
"""Move tensor(s) to chosen device"""
if isinstance(data, (list,tuple)):
return [to_device(x, device) for x in data]
return data.to(device, non_blocking=True)
class DeviceDataLoader():
"""Wrap a dataloader to move data to a device"""
def __init__(self, dl, device):
self.dl = dl
self.device = device
def __iter__(self):
"""Yield a batch of data after moving it to device"""
for b in self.dl:
yield to_device(b, self.device)
def __len__(self):
"""Number of batches"""
return len(self.dl)
# +
input_size = 784
hidden_size = 32 # you can change this
num_classes = 10
model = MnistModel(input_size=input_size, hidden_size=hidden_size, output_size=num_classes)
to_device(model, device)
# -
train_loader = DeviceDataLoader(train_loader, device)
val_loader = DeviceDataLoader(val_loader, device)
# + tags=[]
history1 = fit(5, 0.5, model, train_loader, val_loader)
# + tags=[]
history2 = fit(5, 0.1, model, train_loader, val_loader)
# -
history = history1+history2
# + tags=[]
losses = [x['val_loss'] for x in history]
plt.plot(losses, '-x')
plt.xlabel('epoch')
plt.ylabel('loss')
plt.title('Loss vs. No. of epochs')
# -
def predict(image, model=model):
xb = image.unsqueeze(0)
yb = model(xb)
_, preds = torch.max(yb, dim=1)
return preds[0].item()
test_dataset = MNIST(root='data/',
train=False,
transform=transforms.ToTensor())
# + tags=[]
img, label = test_dataset[0]
plt.imshow(img[0], cmap='gray')
print('Label:', label, ', Predicted:', predict(img, model))
# + tags=[]
jovian.commit(filename='logistic_regression.ipynb', project='pytorch_learn', git_commit=True, git_message='2 layered model')
|
mnist.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Advanced Feature Engineering
# ## Statistics and Distance based Features
#
# ### Statistic based feature is `groupby` feature.
#
# Dataset example:
#
# 
#
#
# Approaches to solve the task:
#
# 1. The most strightforward approach is apply `LabelEncoding` on `Ad_position` feature. In this case, model captures all hiden relationships between variables, but no matter how good it is it still treats all the data points independently.
#
#
# 2. Perform feature engineering to derive intersting relationships of `user-page` pair.
#
# 
#
# 
#
#
#
# On the same way we can create similar usefull features.
# - How many pages user visited
# - Standard deviation of prices
# - Most visited page
# - Many many more
#
#
# ### Distance based feature is `kNN` feature.
#
# - Explicit group is not needed
# - More flexible
# - Much harder to implement
#
# On house price prediction competition, distance based features can be usefull. For example:
#
# 1. Number of houses in 500m, 1000m,..
# 2. Average price per square meter in 500m, 1000m,..
# 3. Number of schools/supermarkets/parking lots in 500m, 1000m,..
# 4. Distance to closest subway station
#
#
# Not only geographical positions, but can apply to others too.
# ## Matrix Factorization for Feature Engineering
#
# Not clear how to use it for feature engineering.
#
# - Matrix Factorization is a very general approach fordimensionality reduction and feature extraction.
# - It can be applied for transforming categorical features into real-valued.
# - Many of tricks trick suitable for linear models can be useful for MF.
#
# Reduces dimension using latent facors. Look at sklearn SVD and TruncatedSVD.
#
# ```python
# # Right way:
# X_all= np.concatenate([X_train,X_test])
# pca.fit(X_all)
# X_train_pca= pca.transform(X_train)
# X_test_pca= pca.transform(X_test)
# ```
# ## Feature Interactions
#
# Very usefull for tree-based models.
#
# 
#
#
# `ad_site` is combination of `category_ad` and `category_site`.
#
# Why we need it? Because model treat each feature independently, by combining them we add some knowledge to model.
#
#
# Implementation:
#
# 1) First variant:
#
# 
#
#
# 2) Second variant:
#
# 
#
#
# Both implementation results are the same.
#
#
# Interaction can be applied for numerical features too.
#
# Combine both numerical features by:
#
# - Multiplication
# - Sum
# - Diff
# - Division
#
# Example:
#
# 
#
#
# Practical Notes:
#
# - We have a lot of possible interactions −N*N for Nfeatures.
# - Need to reduce its’ number
# - a.Dimensionality reduction (Inpractical, don't use)
# - b.Feature selection. (Practically usefull, because only several combinations are important. To find important combinations, run RandomForest or XGBoost and select most import feature combination.)
#
#
# - 2nd order intearctions. (only 2 features combination), easy to construct as above example. For higher order, use:
#
# ```python
# # in sklearn
# tree_model.apply()
#
# # in xgboost
# booster.predict(pred_leaf=True)
# ```
#
# ## t-SNE
#
# - tSNE is a great tool for visualization
# - It can be used as feature as well
# - Be careful with interpretation of results
# - Try different perplexities
|
advanced_feature_engineering/advanced_feature_engineering.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Sweeping Parameters
# + [markdown] tags=[]
# *Modeling and Simulation in Python*
#
# Copyright 2021 <NAME>
#
# License: [Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International](https://creativecommons.org/licenses/by-nc-sa/4.0/)
# + tags=[]
# install Pint if necessary
try:
import pint
except ImportError:
# !pip install pint
# + tags=[]
# download modsim.py if necessary
from os.path import basename, exists
def download(url):
filename = basename(url)
if not exists(filename):
from urllib.request import urlretrieve
local, _ = urlretrieve(url, filename)
print('Downloaded ' + local)
download('https://raw.githubusercontent.com/AllenDowney/' +
'ModSimPy/master/modsim.py')
# + tags=[]
# import functions from modsim
from modsim import *
# + [markdown] tags=[]
# The following cells download the code from Chapter 3 and import the `step` function we defined.
# + tags=[]
download('https://raw.githubusercontent.com/AllenDowney/' +
'ModSimPy/master/chap03.py')
# + tags=[]
from chap03 import step
# -
# In the previous chapter we defined metrics that quantify the performance of a bike sharing system.
# In this chapter we'll see how those metrics depend on the parameters of the system, like the arrival rate of customers at the stations.
#
# And I will present a program development strategy, called incremental
# development, that might help you write programs faster and spend less
# time debugging.
# This chapter is available as a Jupyter notebook where you can read the text, run the code, and work on the exercises.
# Click here to access the notebooks: <https://allendowney.github.io/ModSimPy/>.
# ## Functions That Return Values
#
# We have used several functions that return values.
# For example, when you run `sqrt`, it returns a number you can assign to a variable.
# +
from numpy import sqrt
root_2 = sqrt(2)
root_2
# -
# And when you run `State`, it returns a new `State` object:
bikeshare = State(olin=10, wellesley=2)
bikeshare
# Not all functions have return values. For example, when you run `step`,
# it updates a `State` object, but it doesn't return a value.
#
# To write functions that return values, we can use a `return` statement, like this:
def add_five(x):
return x + 5
# `add_five` takes a parameter, `x`, which could be any number. It
# computes `x + 5` and returns the result. So if we run it like this, the
# result is `8`:
add_five(3)
# As a more useful example, here's a version of `run_simulation` that
# creates a `State` object, runs a simulation, and then returns the
# `State` object:
def run_simulation(p1, p2, num_steps):
state = State(olin=10, wellesley=2,
olin_empty=0, wellesley_empty=0)
for i in range(num_steps):
step(state, p1, p2)
return state
# We can call `run_simulation` like this:
final_state = run_simulation(0.3, 0.2, 60)
# The result is a `State` object that represents the final state of the system, including the metrics we'll use to evaluate the performance of the system:
print(final_state.olin_empty,
final_state.wellesley_empty)
# The simulation we just ran starts with `olin=10` and `wellesley=2`, and uses the values `p1=0.3`, `p2=0.2`, and `num_steps=60`.
# These five values are *parameters of the model*, which are quantities that determine the behavior of the system.
#
# It is easy to get the parameters of a model confused with the parameters of a function.
# It is especially easy because the parameters of a model often appear as parameters of a function.
#
# For example, the previous version of `run_simulation` takes `p1`, `p2`, and `num_steps` as parameters.
# So we can call `run_simulation` with different parameters and see how
# the metrics, like the number of unhappy customers, depend on the
# parameters. But before we do that, we need a new version of a `for` loop.
# ## Loops and Arrays
#
# In `run_simulation`, we use this `for` loop:
#
# ```
# for i in range(num_steps):
# step(state, p1, p2)
# ```
#
# In this example, `range` creates a sequence of numbers from `0` to `num_steps` (including `0` but not `num_steps`).
# Each time through the loop, the next number in the sequence gets assigned to the loop variable, `i`.
#
# But `range` only works with integers; to get a sequence of non-integer
# values, we can use `linspace`, which is defined NumPy:
# +
from numpy import linspace
p1_array = linspace(0, 1, 5)
p1_array
# -
# The arguments indicate where the sequence should start and stop, and how
# many elements it should contain. In this example, the sequence contains
# `5` equally-spaced numbers, starting at `0` and ending at `1`.
#
# The result is a NumPy *array*, which is a new kind of object we have
# not seen before. An array is a container for a sequence of numbers.
#
# We can use an array in a `for` loop like this:
for p1 in p1_array:
print(p1)
# When this loop runs, it
#
# 1. Gets the first value from the array and assigns it to `p1`.
#
# 2. Runs the body of the loop, which prints `p1`.
#
# 3. Gets the next value from the array and assigns it to `p1`.
#
# 4. Runs the body of the loop, which prints `p1`.
#
# 5. ...
#
# And so on, until it gets to the end of the array. This will come in handy in the next section.
# ## Sweeping Parameters
#
# If we know the actual values of parameters like `p1` and `p2`, we can
# use them to make specific predictions, like how many bikes will be at
# Olin after one hour.
#
# But prediction is not the only goal; models like this are also used to
# explain why systems behave as they do and to evaluate alternative
# designs. For example, if we observe the system and notice that we often run out of bikes at a particular time, we could use the model to figure out why that happens. And if we are considering adding more bikes, or another station, we could evaluate the effect of various "what if" scenarios.
#
# As an example, suppose we have enough data to estimate that `p2` is
# about `0.2`, but we don't have any information about `p1`. We could run simulations with a range of values for `p1` and see how the results vary. This process is called *sweeping* a parameter, in the sense that the value of the parameter "sweeps" through a range of possible values.
#
# Now that we know about loops and arrays, we can use them like this:
# +
p1_array = linspace(0, 0.6, 6)
p2 = 0.2
num_steps = 60
for p1 in p1_array:
final_state = run_simulation(p1, p2, num_steps)
print(p1, final_state.olin_empty)
# -
# Each time through the loop, we run a simulation with a different value
# of `p1` and the same value of `p2`, `0.2`. Then we print `p1` and the
# number of unhappy customers at Olin.
#
# To save and plot the results, we can use a `SweepSeries` object, which
# is similar to a `TimeSeries`; the difference is that the labels in a
# `SweepSeries` are parameter values rather than time values.
#
# We can create an empty `SweepSeries` like this:
sweep = SweepSeries()
# And add values like this:
# +
p1_array = linspace(0, 0.6, 31)
for p1 in p1_array:
final_state = run_simulation(p1, p2, num_steps)
sweep[p1] = final_state.olin_empty
# -
# The result is a `SweepSeries` that maps from each value of `p1` to the
# resulting number of unhappy customers.
# + [markdown] tags=[]
# We can display the results like this:
# + tags=[]
show(sweep)
# -
# We can plot the results like this:
# +
sweep.plot(label='Olin', color='C1')
decorate(title='Olin-<NAME>',
xlabel='Customer rate at Olin (p1 in customers/min)',
ylabel='Number of unhappy customers at Olin')
# -
# The keyword argument `color='C1'` specifies the color of the line.
# The `TimeSeries` we have plotted so far use the default color, `C0`, which is blue.
# I use a different color for `SweepSeries` to remind us that it is not a `TimeSeries`.
#
# When the arrival rate at Olin is low, there are plenty of bikes and no unhappy customers.
# As the arrival rate increases, we are more likely to run out of bikes and the number of unhappy customers increases. The line is jagged because the simulation is based on random numbers. Sometime we get lucky and there are relatively few unhappy customers; other times are are unlucky and there are more.
# ## Incremental Development
#
# When you start writing programs that are more than a few lines, you
# might find yourself spending more time debugging. The more code you write before you start debugging, the harder it is to find the problem.
#
# *Incremental development* is a way of programming that tries to
# minimize the pain of debugging. The fundamental steps are:
#
# 1. Always start with a working program. If you have an example from a
# book, or a program you wrote that is similar to what you are working
# on, start with that. Otherwise, start with something you *know* is
# correct, like `x=5`. Run the program and confirm that it does what
# you expect.
#
# 2. Make one small, testable change at a time. A "testable" change is
# one that displays something or has some other effect you can check.
# Ideally, you should know what the correct answer is, or be able to
# check it by performing another computation.
#
# 3. Run the program and see if the change worked. If so, go back to
# Step 2. If not, you have to do some debugging, but if the
# change you made was small, it shouldn't take long to find the
# problem.
#
# When this process works, your changes usually work the first time, or if they don't, the problem is obvious. In practice, there are two problems with incremental development:
#
# - Sometimes you have to write extra code to generate visible output
# that you can check. This extra code is called *scaffolding*
# because you use it to build the program and then remove it when you
# are done. That might seem like a waste, but time you spend on
# scaffolding is almost always time you save on debugging.
#
# - When you are getting started, it might not be obvious how to choose
# the steps that get from `x=5` to the program you are trying to
# write. You will see more examples of this process as we go along,
# and you will get better with experience.
#
# If you find yourself writing more than a few lines of code before you
# start testing, and you are spending a lot of time debugging, try
# incremental development.
# ## Summary
#
# This chapter introduces functions that return values, which we use to write a version of `run_simulation` that returns a `State` object with the final state of the system.
#
# It also introduces `linspace`, which we use to create a NumPy array, and `SweepSeries`, which we use to store the results of a parameter sweep.
#
# We used a parameter sweep to explore the relationship between one of the parameters, `p1`, and the number of unhappy customers, which is a metric that quantifies how well (or badly) the system works.
#
# In the exercises, you'll have a chance to sweep other parameters and compute other metrics.
#
# In the next chapter, we'll move on to a new problem, modeling and predicting world population growth.
# ## Exercises
# ### Exercise 1
#
# Write a function called `make_state` that creates a `State` object with the state variables `olin=10` and `wellesley=2`, and then returns the new `State` object.
#
# Write a line of code that calls `make_state` and assigns the result to a variable named `init`.
# +
# Solution goes here
# +
# Solution goes here
# -
# ### Exercise 2
#
# Read the documentation of `linspace` at <https://numpy.org/doc/stable/reference/generated/numpy.linspace.html>. Then use it to make an array of 101 equally spaced points between 0 and 1 (including both).
# +
# Solution goes here
# -
# ### Exercise 3
#
# Wrap the code from this chapter in a function named `sweep_p1` that takes an array called `p1_array` as a parameter. It should create a new `SweepSeries` and run a simulation for each value of `p1` in `p1_array`, with `p2=0.2` and `num_steps=60`.
# It should store the results in the `SweepSeries` and return it.
#
# Use your function to plot the number of unhappy customers at Olin as a function of `p1`. Label the axes.
# +
# Solution goes here
# +
# Solution goes here
# -
# ### Exercise 4
#
# Write a function called `sweep_p2` that runs simulations with `p1=0.5` and a range of values for `p2`. It should store the results in a `SweepSeries` and return the `SweepSeries`.
#
# +
# Solution goes here
# +
# Solution goes here
# -
# ## Challenge Exercises
#
# The following two exercises are a little more challenging. If you are comfortable with what you have learned so far, you should give them a try. If you feel like you have your hands full, you might want to skip them for now.
# ### Exercise 5
#
# Because our simulations are random, the results vary from one run to another, and the results of a parameter sweep tend to be noisy. We can get a clearer picture of the relationship between a parameter and a metric by running multiple simulations with the same parameter and taking the average of the results.
#
# Write a function called `run_multiple_simulations` that takes as parameters `p1`, `p2`, `num_steps`, and `num_runs`.
# `num_runs` specifies how many times it should call `run_simulation`.
#
# After each run, it should store the total number of unhappy customers (at Olin or Wellesley) in a `TimeSeries`.
# At the end, it should return the `TimeSeries`.
#
# Test your function with parameters
#
# ```
# p1 = 0.3
# p2 = 0.3
# num_steps = 60
# num_runs = 10
# ```
#
# Display the resulting `TimeSeries` and use the `mean` function from NumPy to compute the average number of unhappy customers.
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# -
# ### Exercise 6
#
# Continuing the previous exercise, use `run_multiple_simulations` to run simulations with a range of values for `p1` and
#
# ```
# p2 = 0.3
# num_steps = 60
# num_runs = 20
# ```
#
# Store the results in a `SweepSeries`, then plot the average number of unhappy customers as a function of `p1`. Label the axes.
#
# What value of `p1` minimizes the average number of unhappy customers?
# +
# Solution goes here
# +
# Solution goes here
# -
# ## Under the Hood
#
# The object you get when you call `SweepSeries` is actually a Pandas `Series`, the same as the object you get from `TimeSeries`.
# I give them different names to help us remember that they play different roles.
#
# `Series` provides a number of functions, which you can read about at <https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.Series.html>.
#
# They include `mean`, which computes the average of the values in the `Series`, so if you have a `Series` named `totals`, for example, you can compute the mean like this:
#
# ```
# totals.mean()
# ```
#
# `Series` provides other statistical functions, like `std`, which computes the standard deviation of the values in the series.
#
# In this chapter I use the keyword argument `color` to specify the color of a line plot.
# You can read about the other available colors at <https://matplotlib.org/3.3.2/tutorials/colors/colors.html>.
|
chapters/chap04.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="PQFFW6WMUi8-" colab_type="code" colab={}
import sys
import types
import cv2
import numpy as np
from io import StringIO ## for Python 3
from urllib.request import urlopen
def load_code_from_url(url_path):
code_str = urlopen(url_path).read()
code_str = code_str.decode('utf-8')
return code_str
code_str = load_code_from_url("https://raw.githubusercontent.com/hoat23/VisionArtificialAndImageProcessing/master/bin/utils_imgprocessing.py")
exec(code_str)
# + id="rXVck7jGU9Uk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 264} outputId="2343e218-c0f8-4e1e-db9d-45a1d2d98015"
#url_img = "https://raw.githubusercontent.com/hoat23/VisionArtificialAndImageProcessing/master/img/img_02.jpg"
url_img = "https://raw.githubusercontent.com/hoat23/VisionArtificialAndImageProcessing/master/img/img_04_20x20.jpg"
img_orig = load_image_from_url(url_img)
plt.imshow(img_orig)
plt.show()
# + id="gIAYihnaVchZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 350} outputId="34e29b7c-950c-486d-c6cf-ea9e49a68bf7"
from skimage import data, segmentation
from skimage.segmentation import mark_boundaries
from skimage.future import graph
from google.colab.patches import cv2_imshow
img_tmp = img_orig.copy()
# Aplying SLIC algorithm to get the matrix of labels with 4 segments
n_segments = 4
labels = segmentation.slic(img_tmp, compactness=30, n_segments=n_segments)
img_boundaries_slic = mark_boundaries(img_tmp, labels,color=(255,0,0),background_label=3)
print(labels)
# + id="ymKBwkXeV6Ba" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 597} outputId="36762b3b-56b3-40fb-fa4b-71bd6953cf4c"
img_copy = img_orig.copy()
# Countours Detection
filter_value = 2; threshold_level = 0; mode = cv2.RETR_EXTERNAL # _LIST _EXTERNAL _CCOMP _TREE
mask_8bit = np.uint8( np.where(labels == filter_value, 1 , 0) )
print(mask_8bit)
_, binarized = cv2.threshold(mask_8bit, threshold_level, 255, cv2.THRESH_BINARY)
contours, hierarchy = cv2.findContours(binarized, mode, cv2.CHAIN_APPROX_SIMPLE)
# Drawing contours
countourIdx=255; color = (0,255,0); thickness = 3
img_show = cv2.drawContours(img_copy, contours, -1, (0, 255, 0), 1)
plt.imshow(img_show)
plt.show()
# + id="APPnQjzcVwMJ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 331} outputId="bc988aab-22d5-47fa-d6ad-11ea453b9fbe"
cnt = contours[0]
area = cv2.contourArea(cnt)
x,y,w,h = cv2.boundingRect(cnt)
print(" Area :", area)
print(" Po : (", x,",",y,")")
print(" Width :", w)
print(" Higth :", h)
img_copy = img_orig.copy()
img_rectangle_contour = cv2.rectangle(img_copy,(x,y),(x+w,y+h),(0,255,0),1)
plt.imshow(img_rectangle_contour)
plt.show()
# + id="0c0waDGDM7F5" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 264} outputId="10e0fb33-1835-4c49-b372-ec19241d2b6b"
# Building a image with 4 channel (transparency)
img_copy = img_orig.copy()
img_4ch = np.dstack([img_copy, binarized])
# Setting the figure size
plt.figure(figsize=(10,4))
ax = plt.axes()
# Setting the background color grey
ax.set_facecolor("gray")
plt.imshow(img_4ch)
plt.show()
# + id="I3ad_zpCWNTV" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 597} outputId="d838c84e-8804-4caf-a4b7-ed42357cd066"
# Cutting image using a mask
img_copy = img_orig.copy()
#mask = mask_8bit
print(mask_8bit)
mask_uint8 = mask_8bit.astype('uint8')
img_mask = cv2.merge((mask_uint8*255,mask_uint8*255,mask_uint8*255))
img_bitwise = cv2.bitwise_or(img_copy, img_mask)
bw_r, bw_g, bw_b = cv2.split(img_bitwise)
plt.imshow(img_bitwise)
plt.show()
# + id="dA7ISkFaMCJU" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 264} outputId="5a2835b9-f03e-405c-a982-7f1bcc8ba248"
# Cutting a image
# imagen[y : y + h , x : x + w]
img_cut = img_copy[ y : y+h , x : x+w ]
plt.imshow(img_cut)
plt.show()
|
notebook/08_SLIC_and_Contours.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Modelling of the impact of social tracing apps: a contact network model
#
# 23th April 2020
#
# by <NAME>, <NAME>, <NAME>, <NAME> and <NAME>.
#
# contact : [<EMAIL>]
#
# We wish to thank the _Soyez Prevenus_ team for proposing and organising this modelling challenge !
#
# **An online interactive version of this document is available [here](https://mybinder.org/v2/gh/GuilhemN/covid-19-tracing-app-modelling/master?filepath=model.ipynb)**
# ### Context of creation
#
# This model was created during the "Hack COVID19" Hackathon, which took place between the 10th and the 12th of April 2020 and was organized by HEC and l'École Polytechnique.
# It was created to model the impact of the adoption of a social tracing app, like STOP COVID in France, on the spread of the pandemic.
# Its primary goal is to show the influence of parameters such as the proportion of people using the app in the global population or the effect of individual behaviour after receiving an exposure notification from the app on the epidemic.
#
#
# ### Motivation
#
# We are students in computer science at ENS de Lyon, so we are not experts in the field of epidemiology. This model doesn't aim to provide precise predictions for the future of the pandemic.
# However, it is easily understandable and still reasonnably effective to get an overall sense of what we could expect from a social tracing app.
# More specifically, we believe it gives a good intuition about the impact a social tracing app may have on the epidemic depending on the adoption rate of the app.
# You may try changing the parameters to convince yourself that targeted quarantining of suspected cases can lead to a great limitation of the spread - if it is well adopted by the population.
# So feel free to play with our model!
#
# ### Bibliography
#
# For this model we used the contact network epidemiology approach, as it has been widely studied and it can account for the discrepency of transmission between individuals.
# It has been discussed that this approach is suited to model an epidemic as SARS-CoV-1 in [1].
# Moreover, the structure of the chosen graph has a great impact on the final extent of the outbreak, as discussed in [2].
# We chose two models for the graph:
# - a model using an exponential law for the degrees distribution - which seems suited for a disease such as SARS in urban areas [3]
# - a method from [3] for building the network by stages: first we build highly connected households, then the connections between them.
#
# As for the epidemiological parameters of the disease, we adapted parameters from [4] to suit our model.
# Also, other models for the impact of a social tracing app have been proposed such as in [4] and [5].
#
# ### The model
#
# Each vertex of the graph is an individual that can be : Healthy, Asymptomatic, Presymptomatic, Symptomatic, Dead or Recovered.
# At each step we go trough all the edges and with the probability given by these edges, there is a contact between the individuals.
# There is a parameterized proportion of close contact which are likely to propagate the virus and of being detected by the app, whereas further contact are less likely to propagate the virus but are not detected by the app. The last type of contact models evironnemental contamination by surfaces for example.
#
# The model implements several policies based on screening when an individual gets symptoms:
# - Warning after symptoms:
# * If this policy is applied, the app sends notifications to the persons the infected individual met in the past 14 days as soon as the symptoms are detected. To be more realistic, in this case, non infected individuals may also send notifications at random to model other diseases with similar symptoms.
# * If not, the individual passes a screening test and notifications are only sent if the test is positive (the result is only available after a predefined number of days, which is approximately 5 days for now in France).
#
# - Quarantine after notification:
# * If this policy is applied, a proportion of individuals receiving a notification goes in quarantine directly. They pass a screening test and stop the quarantine if it turns out to be negative.
# * If not, individuals receiving a notification wait until they get the result of their test before going in quarantine.
#
# ### Limits of the model
#
# * Some parameters like the proportion of asymptomatic persons and their infectiosity are not well known. While our work is based on their latest evaluation, they may be reevaluated quite differently in a few days or weeks.
#
# * We don't take into account the eventual false positives of nasal tests. We consider here that there are none.
#
# * The variability of infectiousness seems to be really high: an infected person that will develop symptoms is really contagious during the 2 days before the symptoms appear and is less contagious the 4 days after the symptoms appear, according to [6]. This could moderate our conclusions about the validity window in the "results" part as it gives us 2 to 3 free days during the incubation period.
#
# * The geographical heterogeneity of a national territory is not taken into account, this is a simulation just for one small isolated community (the simulations presented below in the result part are done with 4000 individuals)
#
# * The community is also homogeneous : differences in sex, age or prior conditions that would affect the infection are not taken into account.
#
# * In the initial state, a proportion of the population (5%) is contaminated at the same time. In reality, at the end of the lockdown, the infected will have been contaminated at different instant, smoothing the pressure on healthcare systems.
#
# * The distribution of the time an individual spends as asymptomatic or symptomatic follows a geometrical law. In reality it seems that they are more close to a normal or lognormal distribution. However, beacause symptomatic individuals are quarantined and asymptomatic ones are not the most infectious, this bias should not impact the dynamic of the epidemic too much.
#
# ### Key takeaways
#
#
# * Contaminations from presymptomatic individuals are critical and social distancing can be effective to limit their effect. However, individuals must be warned early (since they do not yet display symptoms). The duration of the signaling chain must be less than the 5 days incubation period.
# * Even with quicker and more precise screening, the app seems unlikely to be effective because of the validity window of nasal tests.
# * The advice of quarantining upon recieving a notification, especially for recent contact, can be a response to this issue but at a high social and economic cost.
# * The "warning after symptoms" policy can be useful to get around the testing delays of symptomatic people if those are longer than 2/3 days. For this, though, we need to precisely identify COVID-19 symptoms and distinguish them from similar symptoms of other diseases.
# * The proportion of users of the app in the population is a central parameter. Under 20% of users, the effects of the app are insignificant, even in a best case scenario. In reality, depending on the policies adopted this threshold could be much higher.
#
# ### References
#
# * [1] Contact network epidemiology: Bond percolation applied to infectious disease prediction and control, <NAME> [(link)](https://www.ams.org/journals/bull/2007-44-01/S0273-0979-06-01148-7/S0273-0979-06-01148-7.pdf)
# * [2] Contact Network Epidemiology: Mathematical Methods of Modeling a Mutating Pathogen on a Two-type Network by <NAME> [(link)](https://repositories.lib.utexas.edu/bitstream/handle/2152/13376/Seilheimer_-_M_08.pdf)
# * [3] Network theory and SARS: predicting outbreak diversity by <NAME> & al. [(link)](https://doi.org/10.1016/j.jtbi.2004.07.026)
# * [4] Quantifying SARS-CoV-2 transmission suggests epidemic control with digital contact tracing. - PubMed - NCBI [(link)](https://science.sciencemag.org/content/sci/early/2020/04/09/science.abb6936.full.pdf)
# * [5] The Impact of Contact Tracing in Clustered Populations by <NAME> & <NAME> [(link)](https://journals.plos.org/ploscompbiol/article/file?id=10.1371/journal.pcbi.1000721&type=printable)
# * [6] Temporal dynamics in viral shedding and transmissibility of COVID-19 by <NAME> & al [(link)](https://www.nature.com/articles/s41591-020-0869-5)
# * [7] Clinical characteristics of 24 asymptomatic infections with COVID-19 screened among close contacts in Nanjing, China by <NAME> & al [(link)](https://link.springer.com/article/10.1007/s11427-020-1661-4)
# * [8] Report of the WHO-China Joint Mission on Coronavirus Disease 2019 (COVID-19) [(link)](https://www.who.int/docs/default-source/coronaviruse/who-china-joint-mission-on-covid-19-final-report.pdf)
#
#
# ### Graph visualisation
#
# Our model is based on a graph simulation: we show a visualisation of the spread of the epidemic in the contact network.
# The contact network is organised in households (little groups of 2 to 6 nodes packed together). The members of a household have a greater probability of contact between them than with members of different households.
#
# Here is the initial state:
# <img src="https://raw.githubusercontent.com/GuilhemN/covid-19-tracing-app-modelling/master/images/init_state.png" alt="init state" style="width: 500px;"/>
#
# Color correspondence:
#
#
# | Color | Meaning |
# | :-------: | :------------------------: |
# | Green | healthy |
# | Red | infected without symptoms |
# | Pink | infected with symptoms |
# | Blue | cured |
# | Black | dead |
#
# Let's see what happens when nobody installs the app.
#
# Day 20:
#
# <img src="https://raw.githubusercontent.com/GuilhemN/covid-19-tracing-app-modelling/master/images/state_20_noapp.png" alt="no app, day 20" style="width: 500px;"/>
#
# After 50 days, almost everyone has been infected (because almost everyone is cured):
#
# <img src="https://raw.githubusercontent.com/GuilhemN/covid-19-tracing-app-modelling/master/images/state_50_no_app.png" alt="no app, day 50" style="width: 500px;"/>
#
# Note that these visualisations are just meant to illustrate the model. In the "Results" section, we used another inital state and experimented different policies.
#
# The graph visualisations were made with Gephi and the Force Atlas 2 algorithm.
#
# ### Detailed results
#
# #### Baseline case: no contact tracing
# <img src="https://raw.githubusercontent.com/GuilhemN/covid-19-tracing-app-modelling/master/images/0%25%2C-%2C-%2C[3%2C10]%2C5j%2C0.3.png" alt="no contact tracing" style="width: 500px;"/>
#
# The epidemic spreads freely from the 5% initial infected rate until almost all the population gets infected.
# The initial Rt (R0) (red curve) is a bit lower than 2 because in the initial population, 10% of the individuals are cured and because people who suffer of symptoms directly quarantine themselves.
# This was not the case at the beginning of the outbreak when R0 was evaluated around 2.
#
# We will take this graph as reference for further analysis.
#
# #### Influence of the waiting time for tests results
#
# We use these test parameters to model the current test abilities in France:
# * 5 days to wait for test results
# * a 3 to 10 days validity window
# * 30% of false positives
#
# Even with a great adoption of the app by the population (80%), no change in the way the epidemic spreads is visible because of the test delay.
# <img src="https://raw.githubusercontent.com/GuilhemN/covid-19-tracing-app-modelling/master/images/80%25%2C-%2C-%2C[3%2C10]%2C5j%2C0.3.png" alt="current testing params in France, 80% adoption" style="width: 500px;"/>
#
# In fact, the critical time period to keep in mind to curb the spread of the disease is the 5 days incubation period.
# According to the last epidemiological statistics, it seems that asymptomatic people make up to about 40% of the infected people - yet they are the cause of only 6% of infections[4].
# On the contrary, the infectiousness of presymptomatic people appears to be really high [4] [6] - so much so that most contaminations originate from them.
# Moreover, when symptoms arise the affected person goes into quarantine. This means that symptomatic transmission is contained if this measure is well respected.
# Thus, presymptomatic contaminations are the most significant, and efforts should focus on them.
# To avoid them, it is necessary to identify and quarantine infected people at the beginning of the incubation period. Therefore, after 5-day delay it is already too late.
#
# *Note: The spikes every 5 days in test demand are due to the fact that an individual cannot ask for a new test if he is already waiting for results.*
#
# In the case of the current testing abilities in France, the characteristic time is of `2*5=10` days between the suspicions of the first infected person and the quarantining of the person notified and infected by the latter. The quarantine arrives far too late to be effective.
#
# Furthermore, the validity window of nasal tests is a key factor that limits their effectiveness. Even if we succeed in having almost perfect tests (1% of false positives) and results available with no delay, we will not be able to identify and isolate presymptomatic people that just got infected because of the 3-10 days validity window.
# Here is a comparison with and without the validity window:
#
# * with the validity window:
# <img src="https://raw.githubusercontent.com/GuilhemN/covid-19-tracing-app-modelling/master/images/80%25%2C-%2C-%2C[3%2C10]%2C0j%2C0.01.png" alt="with validity window" style="width: 500px;"/>
# * without the validity window:
# <img src="https://raw.githubusercontent.com/GuilhemN/covid-19-tracing-app-modelling/master/images/80%25%2C-%2C-%2C[-1%2C100]%2C0j%2C0.01.png" alt="without validity window" style="width: 500px;"/>
#
# With the validity window, we can only identify infected people 3 days after the infection, which is already a significant part of the incubation period.
#
# * with more realistic testing parameters (2 days of delay and 15% of false negative) (still better than current tests) :
# <img src="https://raw.githubusercontent.com/GuilhemN/covid-19-tracing-app-modelling/master/images/80%25%2C-%2C-%2C[3%2C10]%2C2j%2C0.15.png" alt="with better tests" style="width: 500px;"/>
#
# Even with better screening, we can see that the curve of the test requests appears earlier, but the
# quarantine curve doesn't change much. This can be explained by the fact that a close contact has a probability of only about 2% of causing a contamination (according to a WHO report in China [8]). This leads to a high test demand from persons that only have a low chance of being infected. Moreover, in the best case, the decision to quarantine is taken 4 days after the infection, also too late.
#
# In these simulations the demand for tests is unrealistically high because we don't add any constraints on test capacity.
# We may imagine a more nuanced system to determine if we need a test: the number of notifications received, the presumed infectiousness of the infected person at the time of the contact, the comorbidity factors, etc. This could allow a more precise evaluation of the risk an individual has of getting infected.
#
# #### 'Quarantine after notification' policy
#
# * application of the "quarantine after notification" policy and with current tests
# <img src="https://raw.githubusercontent.com/GuilhemN/covid-19-tracing-app-modelling/master/images/80%25%2CN%2C-%2C[3%2C10]%2C5j%2C0.3.png" alt="quarantine after notification with current tests" style="width: 500px;"/>
#
# It provides a more aggressive response against the epidemic and is proven to be effective even with the current test abilities. However, a great part of its effectiveness is likely to be due to the massive quarantine it causes (80% of the population).
# Notice the oscillations of the number of people in quarantine: when the pandemic first spreads, a great part of the population quarantines itself because of the many notifications received. After testing, a lot of them get negative results and end their quarantine, causing the epidemic to spread again.
#
# * application of "quarantine after notification" and with better tests (2 days of delay and 15% of false negative):
# <img src="https://raw.githubusercontent.com/GuilhemN/covid-19-tracing-app-modelling/master/images/80%25%2CN%2C-%2C[3%2C10]%2C2j%2C0.15.png" alt="quarantine after notification with better tests" style="width: 500px;"/>
#
# The effectiveness of this policy is greater with quicker tests because presymptomatic people get notified earlier.
#
# This is illustrated by the rapid collapse of Rt during the first days (red curve).
# In this case, this policy uses quick testing to get earlier and more efficient quarantines.
# With such tests parameters, the average number of quarantine days per person is also reduced as the waiting time for test results is shorter.
#
# However, it should be expected that such a policy comes with a high social cost because of its strict quarantining.
# Indeed, because of the great number of close contacts in a day (13 on average), almost all the users of the app would quarantine themselves after the first wave of contaminations.
# This high social cost could hamper the adoption of the app.
#
# A response to this problem can be to send a notification only to latest contacts. Instead of warning all the close contacts that happended in the last 14 days when testing positive as we did in all our other simulations, we can warn only the contacts of the 7 last days.
# * application of "quarantine after notification" and with better tests (2 days of delay and 15% of false negative) and notificaiton of contacts in the last 7 days:
# <img src="https://raw.githubusercontent.com/GuilhemN/covid-19-tracing-app-modelling/master/images/80%2CN%2C-%2C%5B3%2C10%5D%2C2j%2C0.15_avec_daysNotif_a_7.png" alt="quarantine after notification with better tests" style="width: 500px;"/>
#
# This way, we still warn presymptomatics in the begining of their incubation period without useless qurantine of older contacts that, even if they led to an infection, are now symptomatic or asymptomatic.
# This lowers the timer spent by healthy person in quarantine by 2 days without speeding the spread of the virus too much.
#
# In short, if test results can be available quickly, both the effectiveness of this policy and the social cost are improved.
#
# #### 'Warning after symptoms' policy
#
# * application of "quarantine after notification" and "warning after symptoms" with current tests (5 days of delay and 30% of false negatives):
# <img src="https://raw.githubusercontent.com/GuilhemN/covid-19-tracing-app-modelling/master/images/80%25%2CN%2CW%2C[3%2C10]%2C5j%2C0.3.png" alt="warning after symptoms with current tests" style="width: 500px;"/>
#
# The proportion of notifications sent because of symptoms unrelated to COVID-19 is difficult to estimate. However if we manage to keep it fairly low ( < 0.5% chance of having COVID-looking symptoms per person per day) this policy can be an effective workaround for test delays and test reliability if applied with the "quarantine after notification" policy. The spread of the disease is contained at the same level as with better tests but at the cost of 50% more days of quarantine.
#
# * application of "quarantine after notification" and "warning after symptoms" with better tests (1 days of delay and 1% of false negatives):
# <img src="https://raw.githubusercontent.com/GuilhemN/covid-19-tracing-app-modelling/master/images/80%25%2CN%2CW%2C[3%2C10]%2C1j%2C0.01.png" alt="warning after symptoms with better tests" style="width: 500px;"/>
#
# In both cases it leads to the lockdown of a high proportion of the population.
#
# #### Influence of the proportion of users
#
# In the case of better tests (2 days of delay and 15% of false negatives) and the use of the "quarantine after notification" policy, we plot the influence of the proportion of users on several key values.
#
# <img src="https://raw.githubusercontent.com/GuilhemN/covid-19-tracing-app-modelling/master/images/day_in_quarantine.png" alt="days in quarantine" style="width: 500px;"/>
# <img src="https://raw.githubusercontent.com/GuilhemN/covid-19-tracing-app-modelling/master/images/max_assymp.png" alt="maximum number of asymptomatic cases" style="width: 500px;"/>
# <img src="https://raw.githubusercontent.com/GuilhemN/covid-19-tracing-app-modelling/master/images/proportion_of_healthy.png" alt="proportion of healthy people after 60 days" style="width: 500px;"/>
#
# We see that the use of the app will not reduce the maximal pressure on healthcare system (max. symptomatic) by much but will significantly curb the total spread of the disease.
# By doubling the average number of days in quarantine per person, it triples the final proportion of healthy people.
#
#
# # Parameters
# +
####################
# GRAPH GENERATION #
####################
nbIndividuals = 1000 # number of people in the graph | nombre d'individus dans le graphe
initHealthy = 0.85 # proportion of healthy people at start | la proportion de personnes saines à l'intant initial
initCured = 0.1 # proportion of cured people at start | proportion de personnes guéries à l'instant initial
# The other people are 60% presymptomatic and 40% asymptomatic at start | Les autres personnes sont 40% d'asymptomatiques et 60% de présymptomatiques au départ
# graph generation for exponential degrees distribution
#------------------------------------------------------
deg_avg = 100 # average number of connexions per person | le nombre moyen de connexions par personne
av_household_size = 6 # average size of household | la taille moyenne d'un foyer
household_proba = 1 # probability of meeting a person of the same household | la probabilité de contact par jour entre membres d'un même foyer
extern_contact_proba = 0.3 # probabilty of meeting a person of a different household | la probabilité de contact par jour entre personne de foyers différents
# average contacts per day = 0.3*(100-6) + 1*6 = 34.2
# graph generation with organization in households
#-------------------------------------------------
household_size = (3, 5) # min and max size of an household (uniform distribution) | extremums de la taille d'un foyer
household_link = 1 # probability of contact between members of a household | proba de contact entre membres d'un foyer
number_of_households = 300 # 2500 is good but a bit slow | number of households in the community | nombre de foyers dans une communauté
community_link = 0.3 # probability of contact across households | proba de contact entre foyers
av_deg_by_household = 400 # number of link from a household | nombre moyen de liens depuis un foyer
# average external degree of an individual : 400/4 (4 is the average size of an household)
# average contacts per day = (400/4)*0.3 + 4 = 34
# This paramter was estimated, this is a limit of the model
##############
# APP PARAMS #
##############
daysNotif = 14 # number of days the app checks back for contact notification | nombre de jours vérifiés par l'appli pour notifier un contact
utilApp = 0.8 # percentage of people having the app | la proportion d'utilisateurs de l'application dans la population générale
pDetection = 0.9 # prob. that the app detects a contact | proba que l'appli détecte un contact
pReport = 0.9 # prob. that a user reports his symptoms | proba qu'un utilisateur alerte de ses symptômes
pReadNotif = 0.8 # probablity of taking a notification into account (ask for a test, quarantine) | proba de prendre en compte une notification (demande de test, quarantaine)
pSymptomsNotCovid = 0.005 # every day, everyone sends a notification with prob. pSymptomsNotCovid | chaque jour, tout le monde envoie une notif avec proba PSymptomsNotCovid
############
# POLICIES #
############
# people warn the app immediately after having symptoms | on prévient l'application directement après avoir développé les symptômes
warningAfterSymptoms = False
# upon notification, an individual asks for a test (with some prob.)
# if true, user waits for test results in quarantine, else he goes in quarantine only upon reception of positive test results
# |
# à la reception d'une notif, l'utilisateur demande un test (avec une certaine proba)
# si vrai, il attend les résultats en quarantaine, sinon il ne se met en quarantaine qu'aux résultats d'un test positif
quarantineAfterNotification = True
###############
# TEST PARAMS #
###############
testWindow = (3, 10) # tests are only effective in a given window (time since infection) | les tests ne sont efficaces que dans une fenêtre de temps après infection
daysUntilResult = 2 # Time to wait before test results | attente pour l'obtention des résultats des tests
pFalseNegative = 0.15 # prob. of false negative | proba d'avoir un faux négatif
daysBetweenTests = 0 # the time to wait before being allowed to ask for un new test | temps avant de pouvoir demander un nouveau test
##############
# QUARANTINE #
##############
pQSymptoms = 0.9 # probability of going into quarantine when one has symptoms | proba de confinement lors de détection des symptômes
quarantineFactor = 100 # reduction factor applied to the probabilities when one is in quarantine | réduction des probas de rencontre lors du confinement
daysQuarantine = 14 # duration of the quarantine | durée de la quarantaine
#################
# PROBABILITIES #
#################
pCloseContact = 0.375 # prob. that a contact is a close contact (those detected by the app) | proba qu'un contact soit rapproché (ceux détectés par l'appli)
pContaminationCloseContact = 0.02 # prob. of contamination after close contact with an infected person | proba de contamination après contact rapproché avec qqn d'infecté
#according to [8] around 1 to 5% of close contact lead to virus transmission
pContaminationCloseContactAsymp = 0.006
# infectiousness of asymptomatic people appears to be very low according to [4] and [6]
pContaminationFar = 0.001 # prob. of contamination upon non close contact (environmental or short contact) | proba de contamination par contact environnemental ou bref
pContaminationFarAsymp = 0.0003
# we took R0=2 estimate from [4] and : 34 contacts/day, an average time of infectiousness of 10 days (pre symptomatic + begining of symptoms period)
#average number of infected by symptomatic : (0.375*0.02+0.625*0.001)*34*10 = 2.76
#average number of infected by asymptomatic : (0.375*0.006+0.625*0.0003)*34*10 = 0.83
# this gives 0.6*2.76 + 0.4*0.83 = 1.99 persons infected in average by an infected
# this is plausible given the estimate of R0 and the fact that asymptomatic contamination appears to be minor
# [4] and [6]
# and (0.6*0.625*0.001 + 0.4*0.625*0.0003)*34*10 / R0 = 0.0765 -> the proportion of contaminations which are not due to close contact (environmental / short contact) (contaminations by asymptomatic people are neglected) estimated according to environmental contamination estimate in [4]
# thus most infections (92%) are susceptible to be noticed by the app
# -> the proportion of contaminations by asympt. people is : 0.4*0.83/(0.6*2.76 + 0.4*0.0.83) = 0.17 plausible according to the presumed low infectiosity shown in [4], but this is a conservative estimate (not the 0.06 given by this paper) given the high uncertainty around the results
pAsympt = 0.4 # probability of being asymptomatic when infected | proba qu'une personne infectée soit asymptomatique
# according to [4] and Diamond Princess estimates
# parameters for the lognormal law of the incubation period | paramètres pour la loi lognormale de la période d'incubation
incubMeanlog = 1.644 # -> ~5.5 days
incubSdlog = 0.363 # -> ~2.1 days
# according to [4]
# The next probabilities are given for 1 step of the process, thus overall time spent in these states follows a geometric law for which expected values have been calculated
pAtoG = 0.1 # probability of going from asymptomatic state to cured | proba de passer de asymptomatique à guéri
# according to [7]
pIStoC = 0.07 # probability of going from symptomatic state to cured | proba de passer de avec symptômes à gueri
pIStoD = 0.003 # probability of dying when symptomatic | proba de décès d'une personne présentant des symptômes
# average time with symptoms : 1/(0.07+0.003) = 13.7 days : plausible according to [4]
# death rate when symptoms : 0.003/0.07 = 4.3% : plausible in France according to estimate of 1.6M cases with symptoms and 6 000 deaths the 3 April
# https://www.mgfrance.org/publication/communiquepresse/2525-enquete-mg-france-plus-d-un-million-et-demi-de-personnes-prises-en-charge-par-leur-medecin-generaliste-pour-le-covid-19-entre-le-17-mars-et-le-3-avril
# -
# # Libs and defs
# +
# Librairies
import random
import numpy as np
import sys
# -> sliders
from ipywidgets import interact, interactive, fixed, interact_manual
import ipywidgets as widgets
# -
HEALTHY = 0
ASYMP = 1
PRESYMP = 2
SYMP = 3
CURED = 4
DEAD = 5
# +
class Graph:
""" Object holding the representation of the graph and some metrics """
def __init__(self):
self.individuals = []
self.adj = []
self.encounters = [[[] for day in range(daysNotif)] for individual in range(nbIndividuals)]
self.nbHealthy = 0 # number of healthy people
self.nbAS = 0 # number of asymptomatic people
self.nbPS = 0 # number of premptomatic people
self.nbS = 0 # number of symptomatic people
self.nbCured = 0 # number of cured persons
self.nbDead = 0 # number of deceased people
self.nbQuarantineI = 0 # number of infected people in quarantine
self.nbQuarantineNonI = 0 # number of non infected people in quarantine
self.nbTest = 0 # number of tests made
# cumulative counters :
self.nbQuarantineTotal = 0 # number of people in quarantine
self.nbInfectedByASPS = 0 # number of people infected by asymp. + presymp. people
#to compute Rt
self.stepNb = 0
self.contaminations = [] # number of people contaminated at a given time
self.numInfectedByNewInfected = [] # total number of people who will get infected by people contaminated at a given time
class Individual:
""" Object holding the representation of an individual """
def __init__(self, state, daysQuarantine, app, sentNotification, daysIncubation, timeSinceInfection, timeLeftForTestResult):
self.state = state
self.daysQuarantine = daysQuarantine
self.app = app
self.sentNotification = sentNotification
self.daysIncubation = daysIncubation
self.timeSinceInfection = timeSinceInfection
self.timeSinceLastTest = np.inf # we don't want to test people too often
self.timeLeftForTestResult = timeLeftForTestResult
self.nbInfected = 0
def in_state(self, state):
return self.state == state
def is_infected(self):
return self.state in [PRESYMP, ASYMP, SYMP]
def has_no_covid(self):
return self.state in [HEALTHY, CURED]
def in_quarantine(self):
return self.daysQuarantine > 0
def go_quarantine(self):
if self.daysQuarantine <= 0:
self.daysQuarantine = daysQuarantine # goes into quarantine if isn't already
# -
# # Graph generation
# +
def create_individuals(graph):
graph.contaminations.append(0)
for i in range(nbIndividuals):
app = False
if random.uniform(0,1) < utilApp:
app = True
s = PRESYMP
time_since_infection = -1
incub = 0
r = random.random()
if r < initHealthy:
s = HEALTHY
graph.nbHealthy += 1
elif r < initHealthy + initCured:
s = CURED
graph.nbCured += 1
else:
graph.contaminations[0] += 1 # we start as if a proportion of the population just got infected
time_since_infection = 0
if random.random() < pAsympt:
s = ASYMP
graph.nbAS += 1
else:
s = PRESYMP
incub = round(np.random.lognormal(incubMeanlog, incubSdlog))
graph.nbPS += 1
# state, quarantine, app, notif, incubation, timeSinceInfection, timeLeftForTestResult
graph.individuals.append(Individual(s, 0, app, False, incub, time_since_infection, -1))
def init_graph_exp(graph):
""" Graph initialisation based on exponential ditribution of degrees """
create_individuals(graph)
# affecting degrees to vertices
degrees = np.around(np.random.exponential(deg_avg, nbIndividuals))
# to get an even number of total degrees
S = sum(degrees)
if S%2 == 1:
degrees[0] += 1
S += 1
graph.adj = [[] for i in range(nbIndividuals)]
while S > 0:
# creating an edge
[p1, p2] = np.random.choice(len(degrees), 2, replace=False, p=degrees/S)
if degrees[p1] <= av_household_size or degrees[p2] <= av_household_size:
# the last edges created are edges within households
graph.adj[p1].append({"node" : p2, "proba" : household_proba})
graph.adj[p2].append({"node" : p1, "proba" : household_proba})
else:
graph.adj[p1].append({"node" : p2, "proba" : extern_contact_proba})
graph.adj[p2].append({"node" : p1, "proba" : extern_contact_proba})
degrees[p1] -= 1
degrees[p2] -= 1
S -= 2
def init_graph_household(graph):
""" Graph generation based on households organisation """
global nbIndividuals
# creation of the households
graph.adj = []
for i in range(number_of_households):
size = random.randint(household_size[0], household_size[1])
nb = len(graph.adj)
for i in range(nb, nb+size):
household = []
for j in range(nb, nb+size):
if (i != j):
household.append({"node": j, "proba": household_link})
graph.adj.append(household)
# linkage of the households
for i in range(av_deg_by_household*number_of_households):
[p1, p2] = np.random.choice(len(graph.adj), 2, replace=False)
graph.adj[p1].append({"node": p2, "proba": community_link})
graph.adj[p2].append({"node": p1, "proba": community_link})
nbIndividuals = len(graph.adj)
create_individuals(graph)
graph.encounters = [[[] for day in range(daysNotif)] for individual in range(nbIndividuals)]
# -
# # Updating the graph
def contamination(graph, i, j, closeContact):
""" Individuals i and j have come into contact, leading to a possible contamination | Les individus i et j sont entrés en contact, une contamination est possible """
if graph.individuals[i].state == graph.individuals[j].state:
return
if graph.individuals[i].in_state(HEALTHY):
contamination(graph, j, i, closeContact)
return
# i is the infected individual
if graph.individuals[i].is_infected():
if graph.individuals[j].in_state(HEALTHY):
if closeContact:
pContamination = pContaminationCloseContact
pContaminationAsymp = pContaminationCloseContactAsymp
else:
pContamination = pContaminationFar
pContaminationAsymp = pContaminationFarAsymp
if (random.random() < pContamination and (not graph.individuals[i].in_state(ASYMP))) or \
(random.random() < pContaminationAsymp and graph.individuals[i].in_state(ASYMP)):
# j becomes infected
# for Rt computation
graph.contaminations[graph.stepNb] += 1
graph.numInfectedByNewInfected[graph.stepNb - graph.individuals[i].timeSinceInfection] += 1 # parent infection took place timeSinceInfection ago
if graph.individuals[i].in_state(ASYMP) or graph.individuals[i].in_state(PRESYMP):
graph.nbInfectedByASPS += 1
graph.individuals[j].timeSinceInfection = 0
graph.individuals[i].nbInfected += 1 # i has infected one more person
graph.nbHealthy -= 1
if random.random() < pAsympt:
graph.individuals[j].state = ASYMP
graph.nbAS += 1
else:
graph.individuals[j].state = PRESYMP
graph.individuals[j].daysIncubation = round(np.random.lognormal(incubMeanlog, incubSdlog))
graph.nbPS += 1
# +
def test_individual(individual, graph):
# if there is a test incoming, the person is not tested again
if individual.timeLeftForTestResult >= 0 or individual.in_state(DEAD):
return
# the person was tested not long ago
if individual.timeSinceLastTest < daysBetweenTests:
return
# the person is tested
individual.timeSinceLastTest = 0
graph.nbTest += 1
individual.timeLeftForTestResult = daysUntilResult
if individual.has_no_covid():
individual.latestTestResult = False # we assume that there are no false positives
return
if individual.timeSinceInfection < testWindow[0] or individual.timeSinceInfection > testWindow[1]:
individual.latestTestResult = False # not in the detection window, the test fails
return
# otherwise the person is ill
# the test result depends whether we have a false negative or not
individual.latestTestResult = not (random.random() < pFalseNegative)
# -
def send_notification(graph, i):
""" Send notification to people who have been in touch with i | Envoi d'une notif aux personnes ayant été en contact avec i """
if graph.individuals[i].sentNotification:
return # notifications already sent
graph.individuals[i].sentNotification = True
for daysEncounter in graph.encounters[i]:
# note: graph.encounter[i] is empty if i does not have the app so there is no need to have an additional condition
for contact in daysEncounter:
if random.random() < pReadNotif: # if the person takes the notification into account
# the person is always tested
test_individual(graph.individuals[contact], graph) # asks for a test
if quarantineAfterNotification: # in this case, the person waits for test results in quarantine
graph.individuals[contact].go_quarantine()
# +
def make_encounters(graph, i):
""" Assess all encounters made by i in one day | Détermine toutes les rencontres faites par i en un jour """
for edge in graph.adj[i]:
j = edge['node']
if j < i:
continue # only check one way of the edge | on ne regarde qu'un sens de chaque arête
# if i and/or j are in quarantine, reduce the probability that they meet | si i et/ou j sont confinés, réduction de leur proba de rencontre
factor = 1
if graph.individuals[i].in_quarantine():
factor *= quarantineFactor
if graph.individuals[j].in_quarantine():
factor *= quarantineFactor
if random.random() < edge['proba'] / factor:
if random.random() < pCloseContact: # if this is a close contact
# if i and j have the app, we save their encounter | si i et j ont l'appli, on note la rencontre
if graph.individuals[i].app and graph.individuals[j].app and random.random() < pDetection: # contact detections are symmetric in our model
graph.encounters[i][-1].append(j)
graph.encounters[j][-1].append(i)
contamination(graph, i, j, True)
else:
contamination(graph, i, j, False)
# +
def step(graph):
""" Step from a day to the next day | Passage au jour suivant du graphe """
graph.nbTest = 0
for encounter in graph.encounters:
encounter.append([]) # will contain every encounter of the day | contiendra les nouvelles rencontres du jour
graph.contaminations.append(0)
graph.numInfectedByNewInfected.append(0)
## go through each possible encounter | on constate toutes les rencontres entre individus
for i in range(nbIndividuals):
make_encounters(graph, i)
## update the states | on met à jour les états des individus
for i, individual in enumerate(graph.individuals):
if individual.in_state(ASYMP):
if random.random() < pAtoG:
graph.nbAS -= 1
graph.nbCured += 1
individual.state = CURED
elif individual.in_state(PRESYMP):
if individual.daysIncubation == 0: # the person develops symptoms
graph.nbPS -= 1
graph.nbS += 1
individual.state = SYMP
# send the notifications (encounters[i] is empty if i doesn't have the app) | envoi des notifs (encounters[i] vide si i n'a pas l'appli)
if random.random() < pReport and warningAfterSymptoms:
send_notification(graph, i)
if random.random() < pQSymptoms: # go into quarantine if symptoms appear | mise en confinement à la détection des symptômes
individual.go_quarantine()
test_individual(individual, graph) # all individuals developing symptoms are tested
elif individual.in_state(SYMP):
action = random.random()
if action < pIStoC:
graph.nbS -= 1
graph.nbCured += 1
individual.state = CURED
elif action > 1 - pIStoD:
graph.nbS -= 1
graph.nbDead += 1
individual.state = DEAD
# if warningAfterSymptoms is True, each individual has a probability of sending a false notification due to symptoms that are misinterpreted as from COVID-19
# | si warningAfterSymptoms est vrai, chaque individu a une probabilité d'envoyer une notification en raison de symptômes faussement perçus comme relevant du COVID-19
if warningAfterSymptoms and random.random() < pSymptomsNotCovid:
send_notification(graph, i)
# reception of test results | réception des résultats de test
if individual.timeLeftForTestResult == 0:
if individual.in_quarantine() and individual.latestTestResult == False: # is in quarantine and gets a negative test
individual.daysQuarantine = 0 # end of quarantine
if individual.latestTestResult == True:
individual.go_quarantine()
individual.timeLeftForTestResult = np.inf # people tested positive are not tested again
if random.random() < pReport: # not everyone reports a positive test to the app
send_notification(graph, i)
individual.app = False # unsubscribe from the app in order to not consider new notifications
individual.timeLeftForTestResult -= 1
## results of the day | bilan du jour
graph.nbQuarantineNonI = 0
graph.nbQuarantineI = 0
for individual in graph.individuals:
if individual.in_state(DEAD):
continue
individual.daysQuarantine -= 1
individual.daysIncubation -= 1
individual.timeSinceLastTest += 1
# if there are still symptoms we don't end the quarantine
if (not individual.in_quarantine()) and individual.in_state(SYMP):
individual.daysQuarantine = 1
if individual.in_quarantine():
graph.nbQuarantineTotal += 1/nbIndividuals
if not individual.is_infected():
graph.nbQuarantineNonI += 1
else:
graph.nbQuarantineI += 1
if individual.timeSinceInfection >= 0:
individual.timeSinceInfection += 1
## deleting oldest recorded day | suppression du plus vieux jour de l'historique
for encounter in graph.encounters:
encounter.pop(0)
graph.stepNb += 1
# -
# # Display
# Interactive model below (it takes about 10-15 sec to appear and to run a simulation)
# +
# %matplotlib notebook
import matplotlib.pyplot as plt
fig, ((ax, ax2), (ax3, ax4)) = plt.subplots(2, 2, figsize=[9,8])
axRt = ax3.twinx()
xs = []
y_D = []
y_MS = []
y_MPS = []
y_MAS = []
y_S = []
y_G = []
y_Q = []
y_InfectByASPS = []
y_QuarantineNonI = []
y_QuarantineI = []
y_QuarantineNonITotal = []
y_Test = []
y_TestTotal = []
y_Rt = []
ax.set_ylim([0, nbIndividuals])
def update_viz(graph):
if y_QuarantineNonITotal != []:
y_QuarantineNonITotal.append((graph.nbQuarantineNonI + nbIndividuals*y_QuarantineNonITotal[-1])/nbIndividuals)
y_TestTotal.append((graph.nbTest + nbIndividuals*y_TestTotal[-1])/nbIndividuals)
else:
y_QuarantineNonITotal.append(graph.nbQuarantineNonI/nbIndividuals)
y_TestTotal.append(graph.nbTest/nbIndividuals)
xs.append(len(xs))
y_D.append(graph.nbDead/nbIndividuals*100)
y_MS.append(graph.nbS/nbIndividuals*100)
y_MPS.append(graph.nbPS/nbIndividuals*100)
y_MAS.append(graph.nbAS/nbIndividuals*100)
y_S.append(graph.nbHealthy/nbIndividuals*100)
y_G.append(graph.nbCured/nbIndividuals*100)
y_Q.append(graph.nbQuarantineTotal)
y_InfectByASPS.append(graph.nbInfectedByASPS)
y_QuarantineNonI.append(graph.nbQuarantineNonI/nbIndividuals*100)
y_QuarantineI.append(graph.nbQuarantineI/nbIndividuals*100)
y_Test.append(graph.nbTest/nbIndividuals*100)
def draw_viz(graph):
ax.clear()
ax2.clear()
ax3.clear()
ax4.clear()
axRt.clear()
ax.set_xlabel("Days")
ax2.set_xlabel("Days")
ax3.set_xlabel("Days")
ax4.set_xlabel("Days")
# computing Rt | calcul de Rt
for i in range(graph.stepNb):
if graph.contaminations[i] != 0 and graph.contaminations[i] > 5: # we just take into account days where there were more than 5 contaminations to reduce random fluctuations
y_Rt.append(graph.numInfectedByNewInfected[i]/graph.contaminations[i])
else:
y_Rt.append(0)
for i in range(1, graph.stepNb-1): # smoothing Rt curve
if y_Rt[i] == 0:
y_Rt[i] = (y_Rt[i-1] + y_Rt[i+1])/2
labels = [ "Symptomatic", "Deceased", "Asymptomatic","Presymptomatic", "Cured", "Healthy"]
ax.stackplot(xs, y_MS, y_D, y_MAS,y_MPS, y_G, y_S, labels=labels, edgecolor="black", colors=["red", "darkred", "orange","yellow", "dodgerblue", "mediumseagreen"])
ax.set_ylabel("Proportion of the population")
labels2 = ["In quarantine and non infected (percentage)", "In quarantine and infected (percentage)"]
ax2.stackplot(xs, y_QuarantineNonI, y_QuarantineI, labels=labels2)
ax2.set_ylabel("Proportion of the population")
#line, = ax3.plot(xs, y_InfectByASPS)
#line.set_label("Total infections by asympt.")
ax3.set_ylabel("Quarantine days / Tests")
line, = ax3.plot(xs, y_Q)
line.set_label("Cumulative quarantine days per person")
line, = ax3.plot(xs, y_QuarantineNonITotal)
line.set_label("Cumulative quarantine days of healthy people per person")
line, = ax3.plot(xs, y_TestTotal)
line.set_label("Cumulative number of tests per person")
axRt.set_ylabel("Rt", color = 'red')
line, = axRt.plot(xs, y_Rt, color = 'red')
line.set_label("Rt (average number of infections caused by one infected)")
line, = ax4.plot(xs, y_Test)
line.set_label("Number of tests (in percentage of population)")
ax4.set_ylabel("Tests")
ax.legend(loc='upper center', bbox_to_anchor=(0.5, -0.15), shadow=True, ncol=3)
ax2.legend(loc='upper center', bbox_to_anchor=(0.5, -0.15), shadow=True, ncol=1)
#ax3.legend(loc='upper center', bbox_to_anchor=(0.5, -0.15), shadow=True, ncol=1)
ax3.legend(loc='upper center', bbox_to_anchor=(0.5, -0.15), shadow=True, ncol=1)
#axRt.legend(loc='upper center', bbox_to_anchor=(0.5, -0.15), shadow=True, ncol=1) #to avoid legend on top of the other
ax4.legend(loc='upper center', bbox_to_anchor=(0.5, -0.15), shadow=True, ncol=2)
plt.tight_layout()
def update_prob(app_use_rate, report_to_app, read_notif, false_negative, time_for_test, max_notif_days, warning_after_symptoms, quarantine_after_notification):
global nbIndividuals
global utilApp, pReport, pReadNotif
global pFalseNegative, daysUntilResult, daysNotif
global quarantineAfterNotification, warningAfterSymptoms
global xs, y_D, y_MS, y_MPS, y_MAS, y_S, y_G, y_Q, y_InfectByASPS, y_Rt
global y_QuarantineNonI, y_QuarantineNonITotal, y_QuarantineI, y_Test, y_TestTotal
utilApp = app_use_rate
pReport = report_to_app
pReadNotif = read_notif
pFalseNegative = false_negative
daysUntilResult = time_for_test
daysNotif = max_notif_days
warningAfterSymptoms = warning_after_symptoms
quarantineAfterNotification = quarantine_after_notification
nbSteps = 60
nbIndividuals = 4000 # you may change the number of individuals for the exponential distribution graph here
sys.stdout.write('\r' + "Creation of the graph ...")
sys.stdout.flush()
graph = Graph()
init_graph_household(graph) # default graph generation using households structure, as shown in the Results section
# uncomment this to get a graph with degrees following an exponential distribution
#init_graph_exp(graph)
sys.stdout.write('\r' + " ")
sys.stdout.flush()
xs.clear()
y_D.clear()
y_MS.clear()
y_MPS.clear()
y_MAS.clear()
y_S.clear()
y_G.clear()
y_Q.clear()
y_InfectByASPS.clear()
y_QuarantineNonI.clear()
y_QuarantineNonITotal.clear()
y_QuarantineI.clear()
y_Test.clear()
y_TestTotal.clear()
y_Rt.clear()
maxSymp = 0
for step_ind in range(nbSteps):
# update matplotlib
update_viz(graph)
# update simulation
step(graph)
sys.stdout.write('\r'+'Progress: '+(str((100*step_ind/nbSteps))[:4]) + "%")
sys.stdout.flush()
maxSymp = max(maxSymp, graph.nbS)
sys.stdout.write('\r' + " ")
sys.stdout.flush()
#usefull information to decomment if a more precise evaluation is needed
# print("Total individuals:", nbIndividuals)
# print("Number of deceased:", graph.nbDead)
# print("Max. nb of symptomatic people:", maxSymp)
# print("Test per people:", y_TestTotal[-1])
# print("Final healthy:", y_S[-1])
#print(maxSymp/nbIndividuals,",", y_S[-1],",", y_Q[-1], ",", y_TestTotal[-1])
draw_viz(graph)
plt.show()
update_prob(utilApp, pReport, pReadNotif, pFalseNegative, daysUntilResult, daysNotif, warningAfterSymptoms, quarantineAfterNotification)
interact_manual(update_prob, \
app_use_rate = widgets.FloatSlider(min=0.0, max=1.0, step=0.01, value=utilApp), \
report_to_app = widgets.FloatSlider(min=0.0, max=1.0, step=0.01, value=pReport), \
read_notif = widgets.FloatSlider(min=0.0, max=1.0, step=0.01, value=pReadNotif), \
false_negative = widgets.FloatSlider(min=0.0, max=1.0, step=0.01, value=pFalseNegative), \
time_for_test = widgets.IntSlider(min=0, max=10, step=1, value=daysUntilResult), \
max_notif_days = widgets.IntSlider(min=0, max=30, step=1, value=daysNotif), \
warning_after_symptoms = widgets.Checkbox(value=warningAfterSymptoms), \
quarantine_after_notification = widgets.Checkbox(value=quarantineAfterNotification))
# -
|
model.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="9FACFsyE9iFT"
# # Manejando datos con PANDAS
# + [markdown] id="8-i2lHtH9iFU"
# En el capítulo anterior, hemos dado un cordial saludo a NumPy y sus objetos ``` ndarray ```, que proporcionan un almacenamiento y manipulación eficientes de matrices en Python.
#
# Ahora vamos a conocer a uno de los que van a ser nuestros mejores amigos de aquí al final del bootcamp: los ```DataFrames```, las estructuras de datos proporcionadas por la biblioteca Pandas, que nos permitirán manejar datos con formato tabular.
#
#
# Pandas es un paquete más construido sobre NumPy que nos proporciona una implementación eficiente de un ```DataFrame```.
# Los ```DataFrames``` son esencialmente matrices multidimensionales con etiquetas de fila y columna adjuntas que, a menudo, presentan tipos heterogéneos y / o datos faltantes.
#
#
# Además de ofrecer una interfaz de almacenamiento conveniente para datos etiquetados, Pandas implementa una serie de poderosas operaciones de datos familiares para los usuarios de base de datos y hojas de cálculo.
#
#
# Como hemos visto, la estructura de datos ```ndarray``` de NumPy proporciona características esenciales para el tratamiento de datos adecuados a la computación numérica tan eficiente que comentábamos.
#
#
# Si bien sirve muy bien para este propósito, sus limitaciones se vuelven claras cuando necesitamos más flexibilidad (por ejemplo, adjuntar etiquetas a los datos, trabajar con datos faltantes, etc.) o cuando intentamos realizar operaciones que no se adaptan a estas estructuras, como podrían ser operaciones que vemos en Excel como agrupaciones o _pivot tables_. Estas operaciones no son fáciles de realizar con los arrays de numpy debido precisamente a su estructura, ya que está pensada para ser más eficiente a cabio de perder organización de los datos, cosa que Pandas intenta combinar, es decir, Pandas busca poder realizar un montón de operaciones con datos estructurados sin verse demasiado penalizado desde el punto de vista de la eficiencia.
#
#
# Pandas, y en particular sus objetos ```Series``` y ```DataFrame```, se basan en la estructura array de NumPy proporcionando un acceso eficiente a este tipo de tareas de manipulación de datos que ocupan gran parte del tiempo de un científico de datos.
#
#
# En este capítulo, nos centraremos en la mecánica de usar ```Series```, ```DataFrames``` y estructuras relacionadas de manera efectiva.
#
# -
# ## Primeros pasos
#
# Como ya sabéis, lo primero que tenemos que hacer para poder aprovechar todas las bondades que nos brinda una librería es importarla:
# + colab={"base_uri": "https://localhost:8080/", "height": 34} id="udeK7bx29iFW" jupyter={"outputs_hidden": false} outputId="e8606582-4041-4f6e-ab54-85254c4818d0"
import pandas
pandas.__version__
# + [markdown] id="jJdV68419iFa"
# Sin embargo, así como ```numpy``` lo importábamos con el alias ```np``` para no tener que escribir sieeeempre tanto, ```pandas``` lo importaremos con el alias ```pd```. Qué originales somos, ¿verdad?
# + id="KCyuhicf9iFb"
import pandas as pd
# + [markdown] id="hXN7L_-g9iFe"
# Otro de los pasos que solemos (deberíamos) hacer a la hora de utilizar una librería es leer la documentación, la cual puede ser consultada en su web oficial:
#
# http://pandas.pydata.org/
#
# O mediante código:
# + jupyter={"outputs_hidden": true}
# pd?
# + jupyter={"outputs_hidden": true}
help(pd.DataFrame)
# + [markdown] id="_nFPKUvZ9iFf"
# ## Los objetos de Pandas
# + [markdown] id="bKrZM0jp9iFf"
# Al nivel más básico, los objetos de PAndas pueden ser vistos como una mejora de los arrays de NumPy, donde los datos están organizados mediante el uso de identificadores de las filas y columnas, que serán índices u eitquetas respectivamente.
#
# Como veremos, Pandas nos proporciona una cantidad enorme de herramientas, métodos y funcionalidades para las estructuras básicas más utilizadas. Para aprovecharlo, deberemos dominar 3 estructuras propias de Pandas:
# 1. [Series](#1.-Series)
# 2. [DataFrame](#2.-DataFrame)
# 3. [Index](#3.-Index)
#
# En vista de que Pandas se apoya mucho en NumPy, utilizaremos ambas librerías a lo largo de este notebook:
# + id="SJHNwhKO9iFg"
import numpy as np
import pandas as pd
# + [markdown] id="C5gXfgpy9iFj"
# ## 1. Series
#
# El tipo ``Series`` no es nada más que un array unidimensional de datos indexados. Al igual que los arrays de NumPy, puede ser creado a partir de una lista:
#
# + colab={"base_uri": "https://localhost:8080/", "height": 101} id="s5wj_-V_9iFk" jupyter={"outputs_hidden": false} outputId="d90ab060-7659-42ab-87be-3bec9b7f63df"
data = pd.Series([0.25, 0.5, 0.75, 1.0])
data
# + [markdown] id="W4OxIJJ49iFn"
# Como podemos comprobar, el objeto ``Series`` nos indica unos valores asociados a unos índices, a los cuales podemos acceder mediante los atributos ``values`` e ``index``, respectivamente.
#
# Estos ``values`` son un array de NumPy:
# + colab={"base_uri": "https://localhost:8080/", "height": 34} id="_JF2MpLk9iFn" jupyter={"outputs_hidden": false} outputId="ce71db91-a104-40ea-de26-4315ce2b68dd"
data.values
# + [markdown] id="6gCGfG8s9iFs"
# El atributo ``index`` nos devolverá un objeto de tipo ``pd.Index``, que se parece al tipo ```range()``` que hemos visto en las clases de Python y que detallaremos más adelante.
# + colab={"base_uri": "https://localhost:8080/", "height": 34} id="osrZH0TT9iFs" jupyter={"outputs_hidden": false} outputId="d3cd9c2b-a47f-4ea0-8d6b-e6550d5dc8b8"
data.index
# + [markdown] id="CHMO4eAe9iFw"
# Al igual que hemos visto para las listas y los arrays, podemos acceder a los elementos de un ```Series``` mediante indexación y slicing:
# + colab={"base_uri": "https://localhost:8080/", "height": 34} id="fX4sw1sC9iFw" jupyter={"outputs_hidden": false} outputId="bca47732-3937-4bb9-ccde-f62fa3efb8d7"
data[2]
# + colab={"base_uri": "https://localhost:8080/", "height": 67} id="5M7PR4fi9iFz" jupyter={"outputs_hidden": false} outputId="38504cb6-670c-4e5c-df02-06e56317bb3a"
data[1:3]
# + [markdown] id="_MyxcleC9iF3"
# ### ``Series`` como un array de NumPy generalizado
# + [markdown] id="0bI7Jupb9iF3"
# Lo que hemos visto hasta ahora es que el ```Series``` de Pandas es como un array de NumPy con un índice. Sin embargo, este íncide puede ser determinado de forma explícita, lo que le confiere capacidades adicionales a este objeto. Este índice pordría consistir en valores de cualquier tipo como, por ejemplo, strings:
# + colab={"base_uri": "https://localhost:8080/", "height": 101} id="xgM85EUU9iF3" jupyter={"outputs_hidden": false} outputId="123c50e4-71f5-4dca-9939-26a9038d2d17"
data = pd.Series([0.25, 0.5, 0.75, 1.0],
index=['a', 'b', 'c', 'd'])
data
# + [markdown] id="Wwc6Kv2s9iF7"
# De este modo, podemos acceder a los valores de nuestro ```Series``` como si de un diccionario se tratase:
# + colab={"base_uri": "https://localhost:8080/", "height": 34} id="muXmfPQz9iF7" jupyter={"outputs_hidden": false} outputId="d9c7a49a-1d56-4a50-fc42-c4aecdb3aabc"
data['b']
# + [markdown] id="meVD3j5j9iF9"
# También podríamos utilizar números salteados:
# + colab={"base_uri": "https://localhost:8080/", "height": 101} id="HbdrKBw99iF-" jupyter={"outputs_hidden": false} outputId="10a947a6-c19c-4ade-dfac-9279e768b832"
data = pd.Series([0.25, 0.5, 0.75, 1.0],
index=[2, 5, 3, 7])
data
# + colab={"base_uri": "https://localhost:8080/", "height": 34} id="jnvpvIdv9iGA" jupyter={"outputs_hidden": false} outputId="b048d8ce-e0af-4e25-9d96-24b97ed5704f"
data[5]
# + [markdown] id="8GE1ObxL9iGC"
# ### ``Series`` como un diccionario evolucionado
#
# En vista de lo expuesto anteriormente, también podemos entender un ```Series``` como una especie de diccionario de Python, pues es una estructura que mapea una serie de claves con sus correspondientes valores.
#
# Así como el tipo array de NumPy es más generalmente más eficiente que un Series de Pandas debido a que está especialmente diseñado para optimizar ciertas operaciones, un Series de Pandas es más eficiente que un diccionario para ciertas operaciones.
#
# Tal es la similitud entre el Series de Pandas y el diccionario de Python, que podemos construir un Series a partir de un diccionario de una forma directa:
# + colab={"base_uri": "https://localhost:8080/", "height": 118} id="a-zIOdt49iGD" jupyter={"outputs_hidden": false} outputId="f9d35371-caf1-41f8-fbc2-66ad2b8b567a"
population_dict = {'California': 38332521,
'Texas': 26448193,
'New York': 19651127,
'Florida': 19552860,
'Illinois': 12882135}
population = pd.Series(population_dict)
population
# + [markdown] id="I4zDml449iGF"
# Lo que se hace al crear un series de esta forma es tratar las claves del diccionario como índices. De este modo, si queremos acceder al valor asociado a un índice concreto, utilizaremos la misma notación que para los diccionarios:
# + colab={"base_uri": "https://localhost:8080/", "height": 34} id="ApgWunV79iGG" jupyter={"outputs_hidden": false} outputId="4810be25-7932-4181-b862-b3af7dafa633"
population['California']
# + [markdown] id="q_n6VEbW9iGI"
# Vale. Hasta aquí todo correcto: un ``Series`` es un diccionario. ¿Y eso qué me aporta? ¿No decías que era como un diccionario evolucionado?
#
# Pues sí, y una de las ventajas que tenemos con este objeto de PAndas es que soporta operaciones del estilo array, como slicing, el cual detallaremos más adelante:
# + colab={"base_uri": "https://localhost:8080/", "height": 118} id="fV1J23Ww9iGJ" jupyter={"outputs_hidden": false} outputId="fc0a4b39-a3ce-4ab7-da23-b7d46894fc14"
population['California':'Florida']
# + [markdown] id="5lvgTva79iGM"
# ### Construyendo ~~series de objetos~~ objetos Series
#
# Ya hemos visto cómo crear ```Series``` desde listas, arrays o diccionarios, para lo que hemos seguido la misma sintaxis (aunque en algunos casos omitiéramos el parámetro index):
#
# ```python
# >>> pd.Series(data, index=index)
# ```
#
# donde ``index`` es un argumento opcional que identificará en índice que se asignará a los valores, representados por el argumento ``data``, que podría ser una gran variedad de entidades (ints, floats, strings...).
#
# Por ejemplo, ``data`` podría ser un array de Numpy, como hemos visto anteriormente, mmientras que ```index```, en caso de no ser definido, tomará por defecto lso valores numéricos correspondientes a la posición de cada valor:
# + colab={"base_uri": "https://localhost:8080/", "height": 84} id="I_7_QbuH9iGM" jupyter={"outputs_hidden": false} outputId="c81b3e46-66e1-44ef-a319-a4502ea919d5"
pd.Series([2, 4, 6])
# + [markdown] id="QSiWVch49iGO"
# ``data`` también podría ser un escalar, el cual se repetirá en caso de que especifiquemos más de un índice, al igual que ocurría con los arrays de NumPy cuando especificábamos una forma concreta y le decíamos que lo llenara con un escalar:
# + id="9YJxJG8c9iGP" jupyter={"outputs_hidden": false}
prueba = pd.Series(5, index=['WH', 'LV', 'LV', 'BW'])
prueba
# + colab={"base_uri": "https://localhost:8080/", "height": 67} id="drSKlD5Oo5lr" outputId="b2521dd1-9f8f-4047-a5e0-0f53d160b8ec"
prueba['LV']
# + [markdown] id="F8dEVn_j9iGR"
# En el caso de crearnos un Series a partir de un diccionario, el índice se tomará de las claves:
# + colab={"base_uri": "https://localhost:8080/", "height": 84} id="M5HL00ku9iGR" jupyter={"outputs_hidden": false} outputId="420e9bd5-14dc-42cd-8589-8729abde70d0"
pd.Series({2:'a', 1:'b', 3:'c'})
# + [markdown] id="ycA8sPLg9iGT"
# Y, en caso de ser especificado explícitamente (mediante el parámetro), funcionará como si filtrásemos el diccionario original:
# + colab={"base_uri": "https://localhost:8080/", "height": 67} id="D-8r-ynw9iGU" jupyter={"outputs_hidden": false} outputId="d42b8f2c-ad78-48a5-e01a-561275184277"
pd.Series({2:'a', 1:'b', 3:'c'}, index=[3, 2])
# + [markdown] id="xddzdAVO9iGW"
# ## 2. DataFrame
#
# Una vez vista la estructura ```Series```, ya podemos pasar a los ```DataFrame```, que serán nuestros aliados de aquí hasta el final de los tiempos.
#
# Un ```DataFrame``` es un conjunto de datos estructurados en forma de tabla. Por poner un ejemplo, es como si tuviéramos una tabla de Excel a la que pudiéramos aplicar código Python. En realidad, un DataFrame está compuesto por ```Series``` organizados en forma de tabla, donde podremos acceder a los datos mediante el índice (que seleccionará la/s fila/s) y las columnas (etiquetas que utilizaremos para referirnos a las Series que componen nuestro DataFrame).
#
# Haciendo un símil con el apartado anterior, podemos ver el DataFrame como un array de NumPy de dos dimensiones generalizado, o como un diccionario especializado.
# + [markdown] id="gjzFp2ri9iGY"
# ### ```DataFrame``` como array bidimensional de NumPy generalizado
#
# Así como deciamos que un ```Series``` puede ser visto como un array 1D con las ventajas de los índices, un ```DataFrame``` puede ser interpretado como un array bidimensional con ejes evolucionados, que también nos permitirán esa flexibilidad de los ```Series```.
#
# Como hemos dicho anteriormente, un DataFrame se compone de Series. Para demostrarlo, nos crearemos un Series con los datos del área de cada estado, y lo usaremos, junto con el Series de la población que teníamos creado, para construir nuestro DataFrame:
# + colab={"base_uri": "https://localhost:8080/", "height": 118} id="oe0NAbyg9iGY" jupyter={"outputs_hidden": false} outputId="15f7a80f-3aab-4dc8-d52b-c5f09d97edb5"
area_dict = {'California': 423967, 'Texas': 695662, 'New York': 141297,
'Florida': 170312, 'Illinois': 149995, 'Madrid': 24000}
area = pd.Series(area_dict)
area
# + colab={"base_uri": "https://localhost:8080/", "height": 118} id="kILpFBcFqnQi" outputId="2cc99b30-f7ca-4d8a-ce5c-1dd4fc090c77"
population
# + [markdown] id="naK1oJjw9iGd"
# Ahora que tenemos los 2 Series, nos creamos nuestro DataFrame:
# + colab={"base_uri": "https://localhost:8080/", "height": 202} id="vMsaHkES9iGe" jupyter={"outputs_hidden": false} outputId="b5509f45-0f51-4c50-9843-47324e80fe04"
states = pd.DataFrame({'population': population,
'area': area})
states
# + [markdown] id="15kNfT_z9iGg"
# Al igual que los objetos ``Series``, los ``DataFrame`` tienen un atributo ``index`` que nos da acceso a las filas:
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="F3KCMGrI9iGg" jupyter={"outputs_hidden": false} outputId="b0ad00cc-5bb9-460a-b2c1-75b08811c289"
states.index
# + [markdown] id="LfaoYe2u9iGi"
# Además, los ``DataFrame`` tienen un atributo ``columns``, que es como un índice para las columnas:
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="p0Nhztz99iGi" jupyter={"outputs_hidden": false} outputId="ba9ebd27-f6a8-47da-efb8-8bf7985612a9"
states.columns
# -
# En el fondo, los ```DataFrames``` son arrays con índices que nos permiten trabajar con ellos como si de una tabla se tratase. De hecho, podemos acceder a los valores mediante el atributo ```values```, que es de tipo array:
states.values
# + [markdown] id="goLHEwdM9iGk"
# ### ```DataFrame``` como diccionario especializado
#
# Del mismo modo que hemos visto para las Series, los DataFrames pueden verse como diccionarios especiales, que en lugar de mapear (asociar) una clave a un valor, mapea una clave a una columnade datos, que será de tipo Series.
#
# Cabe destacar que, a diferencia de los arrays, estas estructuras dan más importancia a las columnas, por lo que si hacemos el indexado sobre el DataFrame, estaremos pidiendo la columna en lugar de la fila:
# + colab={"base_uri": "https://localhost:8080/", "height": 121} id="bC-CyZ5L9iGl" jupyter={"outputs_hidden": false} outputId="9d02e1cc-876c-4ad4-c4d9-b26ba9109412"
states['population']
# -
# Si quisiéramos pedir una fila, tendríamos que hacerlo del siguiente modo:
# + colab={"base_uri": "https://localhost:8080/", "height": 69} id="pbgwH5OFs1G7" outputId="3a657da0-9361-4a27-c3eb-6538f521d84f"
states.loc['California']
# + [markdown] id="FCCmuoRN9iGn"
# Y si quisiéramos pedir unas columnas y filas concretas, deberíamos hacer una combinación de ambos:
# + colab={"base_uri": "https://localhost:8080/", "height": 69} id="pbgwH5OFs1G7" outputId="3a657da0-9361-4a27-c3eb-6538f521d84f"
print(states['population'].loc['California':"Madrid"])
# Son intecambiables:
print(states.loc['California']['population'])
# -
# Nos centraremos en la indexación más adelante.
# + [markdown] id="dIQePttK9iGn"
# ### Construyendo DataFrames
#
# Podemos construir nuestros queridos DataFrames de varias formas. A continuación, se mostrarán las más utilizadas:
# + [markdown] id="401tE-HB9iGn"
# #### Series --> DataFrame
#
# Como ya hemos mencionado, los DataFrame se componen de Series, que serán sus columnas, por lo que podemos crear un DataFrame de una sola columna mediante un Series de la siguiente manera:
# -
population
# + colab={"base_uri": "https://localhost:8080/", "height": 202} id="Z6uTbDsk9iGo" jupyter={"outputs_hidden": false} outputId="aaaddad4-7626-462b-9d7e-dd798af864d0"
pd.DataFrame(population, columns=['population'])
# + [markdown] id="NkwJRc6j9iGp"
# #### Lista de diccionarios --> DataFrame
#
# También hemos visto que un diccionario puede ser convertido a un Series. De forma análoga, de una lista de diccionarios podremos crear un DataFrame, donde cada uno de los diccionarios de esa lista será una columna del DataFrame:
# + colab={"base_uri": "https://localhost:8080/", "height": 141} id="_iAsTqN19iGq" jupyter={"outputs_hidden": false} outputId="bb449950-1891-4891-9d74-078221916c92"
# Por cierto, esto que hacemos ahora se llama list comprehension, ¿recordáis para qué servía?
data = [{'a': i, 'b': 2 * i, 'c': 9, 'd': i**2} for i in range(3)]
pd.DataFrame(data)
# -
# OJO: No es necesario que los diccionarios sean del mismo tamaño ni que tengan los mismos índices. En estos casos, los huecos que queden sin valores se rellenarán con ```Nan```, que significa que son valores que no existen, que no es un número (Not a Number):
# + colab={"base_uri": "https://localhost:8080/", "height": 106} id="nu9L1r709iGs" jupyter={"outputs_hidden": false} outputId="8e286546-acc8-4b96-a8d2-14f5cc73d7c9"
pd.DataFrame([{'a': 1, 'b': 2}, {'a': 5, 'v': 89, 'z': 190}, {'x': 10}])
# + [markdown] id="SL21CUSe9iGu"
# #### Diccionario de Series --> DataFrame
#
# Tal como hemos hecho en uno de los apartados anteriores, si juntamos en un diccionario los series con un índice (claves del diccionario), podemos crear un DataFrame de una forma entendible:
# + colab={"base_uri": "https://localhost:8080/", "height": 195} id="Jkhht2nf9iGu" jupyter={"outputs_hidden": false} outputId="7ec48794-56ac-413d-89e8-ff366be8a38c"
pd.DataFrame({'population': population,
'area': area})
# + [markdown] id="9LiO0xPe9iGw"
# #### 2D NumPy array --> DataFrame
#
# Otra forma de crear un DataFrame es hacerlo a partir de un array bidimensional de Numpy, donde las filas y columnas se adaptan perfectamente a la estructura del DataFrame. Además, podremos especificar el índice y las columnas que queremos para esos datos:
# + colab={"base_uri": "https://localhost:8080/", "height": 136} id="auUiOVY-9iGw" jupyter={"outputs_hidden": false} outputId="a0eb7e47-2255-47bf-b21f-1e4ecb0a3271"
pd.DataFrame(np.random.rand(3, 2),
columns=['foo', 'bar'],
index=['a', 'b', 'c'])
# + [markdown] id="laPeuZHb9iG3"
# ## 3. Index
#
# Hemos visto que tanto los ```Series``` como los ```DataFrame``` tienen un índice explícito que les permite acceder a sus datos de manera ordenada. Este índice es un objeto en sí mismo, que puede ser interpretado como un array inmutable o como un set ordenado (que admita valores repetidos, porque sí, los índices pueden tener valores repetidos aunque acabe desembocando en un buen lío).
#
# Para mostrar cómo es un objeto ```Index```, vamos a hacer un ejemplo sencillo:
# + colab={"base_uri": "https://localhost:8080/", "height": 34} id="fDOpTeAa9iG3" jupyter={"outputs_hidden": false} outputId="2c41a344-c8f1-44dd-bebb-b7ae1fa1f61a"
ind = pd.Index([1, 3, 5, 7, 11])
ind
# + [markdown] id="NpdQglf_9iG6"
# ### ```Index``` como array inmutable
#
# El obejto ``Index`` funciona como un array en la mayoría de casos- Por ejemplo, podemos utilizar la misma notación que en el indexing de las listas o los arrays, para devolver valores o trozos (slices):
# + colab={"base_uri": "https://localhost:8080/", "height": 34} id="MCMs2MDU9iG6" jupyter={"outputs_hidden": false} outputId="047ca76a-4100-402b-8729-18b0a21ae58e"
ind[1]
# + colab={"base_uri": "https://localhost:8080/", "height": 34} id="64pT9ua59iG8" jupyter={"outputs_hidden": false} outputId="798dcdcc-6965-4d28-ed7d-4769df583122"
ind[::2]
# + [markdown] id="EasjVw719iG_"
# Dada su naturaleza, los ``Index`` tienen muhos atributos compartidos con los arrays de NumPy:
# + colab={"base_uri": "https://localhost:8080/", "height": 34} id="ssX7_kGQ9iG_" jupyter={"outputs_hidden": false} outputId="7f65429b-efc0-4371-e4aa-b403389bcfb2"
print(ind.size, ind.shape, ind.ndim, ind.dtype)
# + [markdown] id="BQNIkzXm9iHB"
# Aunque, a diferencia de las listas o los arrays, los ``Index`` son inmutables, es decir, no se pueden cambiar de una forma normal. Esto hace que sea más seguro compartir los índices entre múltiples DataFrames y arrays, sin los efectos colaterales de los cambios de los cambios de índice de forma inadvertida.
# + colab={"base_uri": "https://localhost:8080/", "height": 280} id="OQHyFi9f9iHB" jupyter={"outputs_hidden": false} outputId="c579daad-eb64-4af6-cace-7b5953ac1ab3"
ind[1] = 0
# + [markdown] id="0noLLgZI9iHE"
# ### ```Index``` como set ordenado
#
# Los objetos de Pandas están diseñados para facilitar operaciones como los cruces de datasets, lo cual depende de varios aspectos de la aritmética de conjuntos que ya hemos visto mediante el uso de los ```set```.
#
# Los objetos ``Index`` siguen muchas de las convenciones usadas por estos, tales como uniones, intersecciones, diferencias, y otras combinaciones que pueden ser realizadas de un modo similar.
# + id="0QNG-iLd9iHE" jupyter={"outputs_hidden": false}
indA = pd.Index([1, 3, 5, 7, 9])
indB = pd.Index([2, 3, 5, 7, 11])
# + colab={"base_uri": "https://localhost:8080/", "height": 34} id="I6cSkULc9iHF" jupyter={"outputs_hidden": false} outputId="bd7e23ea-f53e-485d-ec99-41c49f08ea04"
indA & indB # intersection
# + colab={"base_uri": "https://localhost:8080/", "height": 34} id="9ZGmkHWt9iHH" jupyter={"outputs_hidden": false} outputId="76c8c90a-dd37-4b85-e2b1-0b443bf707d8"
indA | indB # union
# + colab={"base_uri": "https://localhost:8080/", "height": 34} id="wpSwvHbl9iHJ" jupyter={"outputs_hidden": false} outputId="2e49ca26-4e8e-4746-ace0-69e4b0eef75d"
indA ^ indB # symmetric difference
# + [markdown] id="zlFFlsEi9iHK"
# Al igual que con los sets, estas operaciones también podrían ser realizadas mediante los métodos correspondientes, como:
# + id="YMkLiFqi9iHK"
indA.intersection(indB)
# -
df = pd.DataFrame({'population': population,
'area': area})
df.index = [1, 2, 3, 4, 5, 6]
df
# +
series1 = pd.Series({'a': 1, 'b': 2, 'c':3, 'd': 14})
series2 = pd.Series({'a': 1, 'e': 2, 'c':4, 'd': 14})
series1.index = ['a', 'b', 'b', 'c']
# series1
# pd.DataFrame(series1)
pd.DataFrame({'population': series1,
'population2': series2})
# + jupyter={"outputs_hidden": true}
prueba = pd.Series(5, index=['WH', 'LV', 'LV', 'BW'])
pd.DataFrame({'a': prueba}, index=['WH', 'LV', 'BW'])
# + [markdown] id="laPeuZHb9iG3"
# ## Practica lo aprendido
#
# A continuación, vamos a hacer unos ejercicios relacionados con lo que acabamos de ver:
#
# ### Ejercicio 1: el Creador
#
# 1. Crea una lista de 10 elementos del tipo que quieras y llámala ``lista1``
# 2. Crea un array con los 10 primeros números pares (el 0 no es par) y llámalo ``array1``
# 3. Crea 2 objetos tipo Series a partir de ``lista1`` y ``array1``, y llámalos ``series1`` y ``series2``, respectivamente. Los índices de ``series1`` deberán ser las 10 primeras letras del abecedario, y los de ``series2`` deberán ser aleatorios entre la 'a' y la 'z' (puedes usar la función np.random.choice() para seleccionar aleatoriamente de una lista que le pases).
# PISTA: la función ``chr(n)`` nos devolverá un caracter en función de un número, siendo ``chr(97) = 'a', chr(98) = 'b'...``
# 4. Crea un DataFrame a partir de los 2 Series que te has creado en el punto anterior, llámalo ``df1``
# 5. Crea un DataFrame (``df2``) con 5 columnas, cuyos nombres y valores serán:
# 1. 'Columna1' (``series1``)
# 1. 'Columna1_otra_vez' (``series1``)
# 1. 'Columna_diferente' (``series2``)
# 1. 'Col_dif' (``series2``)
# 1. 'Coolumna' (``series1``)
# +
import pandas as pd
import numpy as np
# 1
lista1 = ['Messi', 'Son', 'Sancho', '<NAME>', 'Mbappe', 'Kimmich', 'Haaland', 'Pedri', 'Trincao', 'Dest']
# 2
array1 = np.arange(10) * 2 + 2
# 3
indices = []
n = 97
for i in range(26):
indices.append(chr(n + i))
ind1 = indices[0:10]
series1 = pd.Series(lista1, ind1)
print(series1)
print("\n")
ind2 = np.random.choice(indices, 10, replace = False)
series2 = pd.Series(array1, ind2)
print(series2)
print("\n")
# 4
df1 = pd.DataFrame({'Serie 1': series1, 'Serie 2': series2})
print(df1)
# 5
df2 = pd.DataFrame({'Columna1': series1,
'Columna1_otra_vez': series1,
'Columna_diferente': series2,
'Col_dif': series2,
'Columna': series1})
df2
# +
l = []
for i in range(10):
l.append(np.random.choice(indices, replace = False))
l
# + [markdown] id="laPeuZHb9iG3"
# ### Ejercicio 2: el Creador 2
#
# 1. Crea un par de diccionarios, ``dic1`` y ``dic2``, con los valores de ``lista1`` y ``array1`` y con las claves los índices ``ind1`` e ``ind2``. Llámalos dic1 y dic 2, respectivamente.
# 2. Crea un par de objetos índice ``ind3`` y ``ind4`` a partir de los índices anteriores
# 3. Saca la intersección de los índices
# 4. Créate ahora los objetos series ``series3`` y ``series4`` a partir de los diccionarios anteriores y con el índice común
# 5. Crea un DataFrame a partir de estas series
# -
|
semana_6/dia_1/1_IntroPandas.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.5 64-bit (''ml'': conda)'
# metadata:
# interpreter:
# hash: aa12e81bb39beeca05898a0aa92eeb9ac673e8b720e9a11bb6f6303f215907fc
# name: python3
# ---
import pandas as pd
df = pd.read_csv('winequality-red.csv')
df.head()
df.describe()
# +
from sklearn.model_selection import train_test_split
X = df.drop('quality', axis=1).to_numpy()
y = df['quality'].to_numpy()
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=123, shuffle=True)
print(f'Shapes:\nX_train: {X_train.shape}, y_train: {y_train.shape}\nX_test: {X_test.shape}, y_test: {y_test.shape}')
# +
from tensorflow.keras.layers import Input, Dense, BatchNormalization
from tensorflow.keras.models import Model
input = Input(shape=X_train.shape[1:])
norm = BatchNormalization()(input)
l1 = Dense(units=50, activation='relu')(norm)
l2 = Dense(units=1)(l1)
model = Model(inputs=[input], outputs=[l2])
model.compile(
loss='mse',
optimizer='SGD'
)
from tensorflow.keras.callbacks import EarlyStopping
EScb = EarlyStopping(patience=3, restore_best_weights=True)
history = model.fit(
X_train,
y_train,
epochs=100,
validation_data=(X_test, y_test),
callbacks=[EScb]
)
# +
import matplotlib.pyplot as plt
fig, ax = plt.subplots(ncols=1, nrows=1, figsize=(20, 10))
ax.plot(history.history['loss'], label='loss')
ax.plot(history.history['val_loss'], label = 'val_loss')
ax.set_xlabel('Epoch')
ax.set_ylabel('Loss')
ax.legend(loc='upper right');
#plt.savefig('loss_plot.png', dpi=300)
# -
import numpy as np
test = np.array([[8.319637,0.527821,0.270976,2.538806,0.087467,15.874922,46.467792,0.996747,3.311113,0.658149,10.422983],])
model.predict(test)
|
Regression/WineQualityPrediction/winequality.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="_ikqR4WcZTtg"
# # Cavity Detection Tool (CADET)
# + [markdown] id="oMMhLTf0FRGN"
# ## Download files
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 4587, "status": "ok", "timestamp": 1626286820330, "user": {"displayName": "Tom\u00e1\u0161 Pl\u0161ek", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgeoFcGyfJEsJXX2kJpPud49PzRQCXp2EYggF61aA=s64", "userId": "12667160157784747384"}, "user_tz": -120} id="hs8JG11EGomr" outputId="4f719ff7-b497-4a58-d756-041b2db88c51"
from google_drive_downloader import GoogleDriveDownloader as gdd
gdd.download_file_from_google_drive(file_id='1XpUkeadqowZATYCN2YwLXGjXcJRT33dJ',dest_path='./CADET_size.h5')
gdd.download_file_from_google_drive(file_id='1eneLmdvzq_onbVxTIRgZJmVu3vTiTSbU',dest_path='./NGC4696.fits')
gdd.download_file_from_google_drive(file_id='11P66DF_M3L2V12tV3MzBHqYzQdlFRj7T',dest_path='./NGC4778.fits')
gdd.download_file_from_google_drive(file_id='1QE1gZShYhXrfyuEk2eUFLAoAk1G2F8U9',dest_path='./NGC5813.fits')
# + [markdown] id="H-W7kwe6FUVT"
# ## Import libraries
# + executionInfo={"elapsed": 231, "status": "ok", "timestamp": 1626287609024, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgeoFcGyfJEsJXX2kJpPud49PzRQCXp2EYggF61aA=s64", "userId": "12667160157784747384"}, "user_tz": -120} id="6_sR5sacFiZz"
import os, sys, glob
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from astropy.io import fits
from astropy.wcs import WCS
from astropy.nddata import CCDData
from keras.models import load_model
from keras.layers import LeakyReLU
from sklearn.cluster import DBSCAN
from scipy.ndimage import center_of_mass, rotate
import warnings
warnings.filterwarnings("ignore")
import plotly.express as px
import plotly.graph_objects as go
# + [markdown] id="qTGMIZG5Zd3d"
# ## Function for producing 3D cavity models
# + executionInfo={"elapsed": 222, "status": "ok", "timestamp": 1626288232658, "user": {"displayName": "Tom\u<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgeoFcGyfJEsJXX2kJpPud49PzRQCXp2EYggF61aA=s64", "userId": "12667160157784747384"}, "user_tz": -120} id="Ov-mo-FSEUyL"
def produce_3D_cavity(img, name="", plot=False):
'''
Function produces a 3D cavity model from 2D cavity prediction and calculates it's volume. The 3D model can be also alternatively plotted or stored in the .npy format.
Input
-----
- two dimensional decomposed cavity prediction produced by the CADET network
Parameters
----------
img : 2D numpy array of size 128x128
Two dimensional cavity prediction.
name : string, optional
Name of the
Returns
-------
volume : float
Volume of the predicted cavity calculated under the assumption of rotational symmetry in every point along the direction from the galactic centre.
Output (optional)
------
- 3D cavity model stored in the .npy format
- 3D cavity plot
'''
# find the center of mass of the cavity and de-rotate it
cen = center_of_mass(img)
phi = np.arctan2(cen[0]-63.5, cen[1]-63.5)
img = rotate(img, phi*180/np.pi, reshape=False, prefilter=False)
# iterate over image slices and determine cavity width in each slice
means, widths, indices = [], [], []
for n in range(128):
rang = np.where(img[:,n] > 0, np.arange(0,128), 0)
if not (rang > 0).any(): continue
x = 0
for i,r in enumerate(rang):
if r > 0 and x == 0: x = i
elif x != 0 and r == 0:
widths.append((i-x)/2)
means.append((x+i)/2)
indices.append(n)
x = 0
# produce 3D cavity model
cube = np.zeros((128,128,128))
for m, w, i in zip(means, widths, indices):
x, y = np.indices((128, 128))
r = np.sqrt((x-abs(m))**2 + (y-63.5)**2)
sliced = np.where(r <= w, 1, 0)
cube[:,:,i] += sliced
# (optional) plot 3D cavity model
if plot:
Zi,Xi,Yi = cube.nonzero()
# fig = plt.figure()
# ax = fig.add_subplot(111, projection='3d')
# ax.plot(Xi,Yi,Zi, "sy", markersize=3, mec="k")
# ax.plot(63.5, 63.5, 63.5, "or", ms=2)
# ax.set_xlim(0,128)
# ax.set_ylim(0,128)
# ax.set_zlim(0,128)
# ax.autoscale_view('tight')
# fig.tight_layout()
# plt.show()
df = pd.DataFrame(list(zip(Xi,Yi,Zi)), columns=["x", "y", "z"])
fig = px.scatter_3d(df, x='x', y='y', z='z')
fig.update_layout(scene = dict(
xaxis = dict(nticks=4, range=[0,128],),
yaxis = dict(nticks=4, range=[0,128],),
zaxis = dict(nticks=4, range=[0,128],),),
width=700,
margin=dict(r=20, l=10, b=10, t=10))
fig.update_traces(marker=dict(size=1.5, color="yellow", symbol="square",
line=dict(width=1,
color='DarkSlateGrey')),
selector=dict(mode='markers'))
fig.show()
# (optional) store the 3D cavity model in the .npy format
if name: np.save("{0}.npy".format(name), cube)
# return the cavity volume
volume = np.sum(cube)
return volume
# + [markdown] id="il9oN84DcQ5I"
# ## Usage
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} executionInfo={"elapsed": 10728, "status": "ok", "timestamp": 1626287544053, "user": {"displayName": "Tom\u00e1\u0161 Pl\u0161ek", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgeoFcGyfJEsJXX2kJpPud49PzRQCXp2EYggF61aA=s64", "userId": "12667160157784747384"}, "user_tz": -120} id="4VXYW_ewFp5X" outputId="98502b91-7e1d-4a55-a3a6-465047693fc9"
threshold = 0.55
# load the model
path_to_model = "CADET_size.h5"
if not os.path.exists(path_to_model):
print("\nThere is no {0} file in the current directory".format(path_to_model))
model = load_model(path_to_model, custom_objects = {"LeakyReLU": LeakyReLU})
# find FITS files
files = glob.glob("*.fits")
if len(files) == 0:
print("\nThe current directory does not seem to contain any FITS files.")
os.system("mkdir -p predictions decomposed txts")
# iterate over all FITS files
for filename in files:
name = filename.split("/")[-1].split(".")[0]
# load FITS file
with fits.open(filename) as file:
data = file[0].data
wcs = WCS(file[0].header)
# check the input shape
shape = data.shape
if shape != (128, 128):
print("The image of size {0}x{1} is not supported. Please, transform your image into the 128x128 format.".format(*shape))
continue
# produce prediction
data = np.log10(data+1) / np.amax(np.log10(data+1))
pred = model.predict(data.reshape(1,128,128,1))
pred = pred.reshape(128,128)
ccd = CCDData(pred, unit="adu", wcs=wcs)
ccd.write("predictions/{0}_CADET_size.fits".format(name), overwrite=True)
# cut-off by the threshold and select nonzero points
pred_cut = np.where(pred > threshold, pred, 0)
x, y = pred_cut.nonzero()
points = np.array([x,y]).reshape(2, -1)
# decompose the prediction
clusters = DBSCAN(eps=3).fit(points.T)
clusters = clusters.labels_
N = len(set(clusters))
cavs, areas, volumes, xn, yn, clustersn = [], [], [], [], [], []
for i in range(N):
img = np.zeros((128,128))
b = clusters == i
xi, yi = x[b], y[b]
img[xi,yi] = 1
# ignore too small cavities
if np.sum(img) < 30: continue
xn = np.concatenate((xn, xi))
yn = np.concatenate((yn, yi))
clustersn = np.concatenate((clustersn, clusters[b]))
cavs.append(img)
areas.append(np.sum(img))
volumes.append(produce_3D_cavity(img))
#volumes.append(produce_3D_cavity(img, plot=True))#,name="{0}/{1}".format(dir,name))
# save areas and volumes into txt file
with open("txts/{0}_size.txt".format(name), "w") as file:
print("\n{0}\n# AREA (px^2) VOLUME (px^3)".format(name))
file.write("# AREA (px^2) VOLUME (px^3)")
for i, area, volume in zip(range(len(areas)), areas, volumes):
line = "{0} {1:.0f} {2:.0f}".format(i+1, area, volume)
print(line)
file.write("\n"+line)
# plot decomposed predictions
plt.figure(figsize=(9,3))
plt.subplot(131)
plt.text(35, 113, name, color="white", size=18, va="center", ha="center")
plt.imshow(data, origin="lower")
plt.xticks([]); plt.yticks([])
plt.subplot(132)
plt.imshow(pred, origin="lower")
plt.xticks([]); plt.yticks([])
plt.subplot(133)
plt.scatter(yn, xn, c=clustersn, marker="o")
for j,cav in enumerate(cavs):
center = center_of_mass(cav)
if not np.isnan(center[0]):
plt.text(center[1], center[0], str(j+1), fontsize=20,
c="black", ha="center", va="center")
plt.xticks([]); plt.yticks([])
plt.xlim(0,128); plt.ylim(0,128)
plt.tight_layout()
plt.savefig("decomposed/{0}_CADET_size.png".format(name))
# + [markdown] id="UoehdNkBS6U7"
# ## 3D cavity model
# + colab={"base_uri": "https://localhost:8080/", "height": 542} executionInfo={"elapsed": 548, "status": "ok", "timestamp": 1626288236637, "user": {"displayName": "Tom\u00e1\u0161 Pl\u0161ek", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgeoFcGyfJEsJXX2kJpPud49PzRQCXp2EYggF61aA=s64", "userId": "12667160157784747384"}, "user_tz": -120} id="mkNCz9k8RB9F" outputId="78c1eb15-153a-4a5e-8229-25c7bca36a5b"
produce_3D_cavity(img, plot=True);
# + id="Z8t1w0fNSgVo"
|
CADET_example_colab.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
data = pd.read_csv("50_Startups-Copy1.csv")
data.head(10)
X = data.iloc[:, :3].values
Y = data.iloc[:, 4].values
data.describe()
data.shape
data.isnull().sum()
# +
from sklearn.preprocessing import LabelEncoder, OneHotEncoder
labelencoder = LabelEncoder()
data["State"] = labelencoder.fit_transform(data["State"])
# -
X = data.iloc[:, :3].values
Y = data.iloc[:, 4:].values
# +
from sklearn.cross_validation import train_test_split
X_train, X_test,Y_train, Y_test = train_test_split(X, Y, test_size = 0.4, random_state =0)
print(X_train.shape, Y_train.shape)
print(X_test.shape, Y_test.shape)
# +
from sklearn.linear_model import LinearRegression
regressor = LinearRegression().fit(X_train, Y_train)
# -
predict = regressor.predict(X_test)
|
Code/Day3/Multiple_Linear_Regression.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.6.13 64-bit (''nlp-tutorial'': conda)'
# language: python
# name: python3
# ---
from nltk.tokenize import word_tokenize, WordPunctTokenizer
from tensorflow.keras.preprocessing.text import text_to_word_sequence
print('단어 토큰화1 :',word_tokenize("Don't be fooled by the dark sounding name, Mr. Jone's Orphanage is as cheery as cheery goes for a pastry shop."))
print('단어 토큰화2 :',WordPunctTokenizer().tokenize("Don't be fooled by the dark sounding name, Mr. Jone's Orphanage is as cheery as cheery goes for a pastry shop."))
print('단어 토큰화3 :',text_to_word_sequence("Don't be fooled by the dark sounding name, Mr. Jone's Orphanage is as cheery as cheery goes for a pastry shop."))
# +
from nltk.tokenize import TreebankWordTokenizer
tokenizer = TreebankWordTokenizer()
text = "Starting a home-based restaurant may be an ideal. it doesn't have a food chain or restaurant of their own."
print('트리뱅크 워드토크나이저 :',tokenizer.tokenize(text))
# +
from nltk.tokenize import sent_tokenize
text = "His barber kept his word. But keeping such a huge secret to himself was driving him crazy. Finally, the barber went up a mountain and almost to the edge of a cliff. He dug a hole in the midst of some reeds. He looked about, to make sure no one was near."
print('문장 토큰화1 :',sent_tokenize(text))
# -
text = "I am actively looking for Ph.D. students. and you are a Ph.D student."
print('문장 토큰화2 :',sent_tokenize(text))
# +
import kss
text = '<NAME> 자연어 처리가 재미있기는 합니다. 그런데 문제는 영어보다 한국어로 할 때 너무 어렵습니다. 이제 해보면 알걸요?'
print('한국어 문장 토큰화 :',kss.split_sentences(text))
# +
import nltk
nltk.download('averaged_perceptron_tagger')
from nltk.tokenize import word_tokenize
from nltk.tag import pos_tag
text = "I am actively looking for Ph.D. students. and you are a Ph.D. student."
tokenized_sentence = word_tokenize(text)
print('단어 토큰화 :',tokenized_sentence)
print('품사 태깅 :',pos_tag(tokenized_sentence))
# +
from konlpy.tag import Mecab
from konlpy.tag import Kkma
mecab = Mecab()
kkma = Kkma()
print('Mecab 형태소 분석 :',mecab.morphs("열심히 코딩한 당신, 연휴에는 여행을 가봐요"))
print('Mecab 품사 태깅 :',mecab.pos("열심히 코딩한 당신, 연휴에는 여행을 가봐요"))
print('Mecab 명사 추출 :',mecab.nouns("열심히 코딩한 당신, 연휴에는 여행을 가봐요"))
print('꼬꼬마 형태소 분석 :',kkma.morphs("열심히 코딩한 당신, 연휴에는 여행을 가봐요"))
print('꼬꼬마 품사 태깅 :',kkma.pos("열심히 코딩한 당신, 연휴에는 여행을 가봐요"))
print('꼬꼬마 명사 추출 :',kkma.nouns("열심히 코딩한 당신, 연휴에는 여행을 가봐요"))
|
CH2/2-1)tokenization.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
import h5py
from lr_utils import load_dataset
# + pycharm={"name": "#%%\n"}
train_set_x_orig , train_set_y , test_set_x_orig , test_set_y , classes = load_dataset()
# + pycharm={"name": "#%%\n"}
index = 25
plt.imshow(train_set_x_orig[index])
# + pycharm={"name": "#%%\n"}
#打印出当前的训练标签值
#使用np.squeeze的目的是压缩维度,【未压缩】train_set_y[:,index]的值为[1] , 【压缩后】np.squeeze(train_set_y[:,index])的值为1
#print("【使用np.squeeze:" + str(np.squeeze(train_set_y[:,index])) + ",不使用np.squeeze: " + str(train_set_y[:,index]) + "】")
#只有压缩后的值才能进行解码操作
print("y=" + str(train_set_y[:,index]) + ", it's a " + classes[np.squeeze(train_set_y[:,index])].decode("utf-8") + "' picture")
# + pycharm={"name": "#%%\n"}
m_train = train_set_y.shape[1]
m_test = test_set_y.shape[1]
num_px = train_set_x_orig.shape[1]
#现在看一看我们加载的东西的具体情况
print ("训练集的数量: m_train = " + str(m_train))
print ("测试集的数量 : m_test = " + str(m_test))
print ("每张图片的宽/高 : num_px = " + str(num_px))
print ("每张图片的大小 : (" + str(num_px) + ", " + str(num_px) + ", 3)")
print ("训练集_图片的维数 : " + str(train_set_x_orig.shape))
print ("训练集_标签的维数 : " + str(train_set_y.shape))
print ("测试集_图片的维数: " + str(test_set_x_orig.shape))
print ("测试集_标签的维数: " + str(test_set_y.shape))
# + pycharm={"name": "#%%\n"}
#X_flatten = X.reshape(X.shape[0],-1).T #X.T是X的转置
#将训练集的维度降低并转置。
train_set_x_flatten = train_set_x_orig.reshape(train_set_x_orig.shape[0], -1).T
#将测试集的维度降低并转置。
test_set_x_flatten = test_set_x_orig.reshape(test_set_x_orig.shape[0], -1).T
# + pycharm={"name": "#%%\n"}
print ("训练集降维最后的维度: " + str(train_set_x_flatten.shape))
print ("训练集_标签的维数 : " + str(train_set_y.shape))
print ("测试集降维之后的维度: " + str(test_set_x_flatten.shape))
print ("测试集_标签的维数 : " + str(test_set_y.shape))
# + pycharm={"name": "#%%\n"}
train_set_x = train_set_x_flatten / 255
test_set_x = test_set_x_flatten / 255
# + [markdown] pycharm={"name": "#%% md\n"}
# 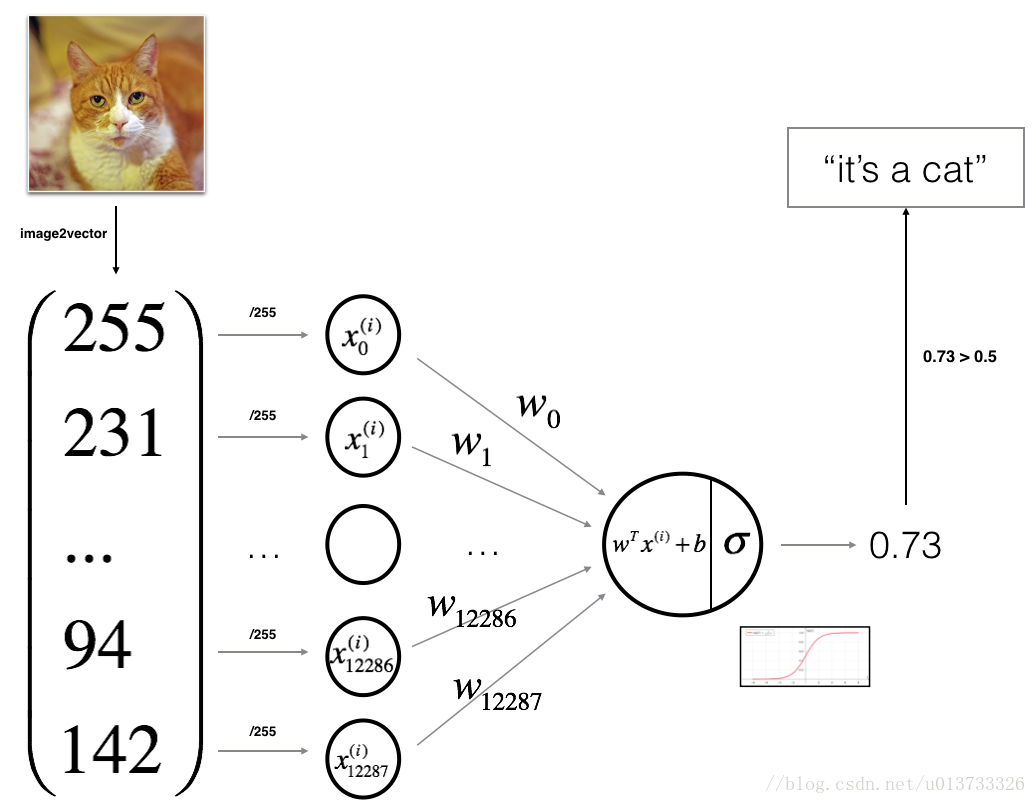
# + pycharm={"name": "#%%\n"}
def sigmoid(z):
"""
参数:
z - 任何大小的标量或numpy数组。
返回:
s - sigmoid(z)
"""
s = 1 / (1 + np.exp(-z))
return s
# + pycharm={"name": "#%%\n"}
# 测试sigmoid()
print("====================测试sigmoid====================")
print ("sigmoid(0) = " + str(sigmoid(0)))
print ("sigmoid(9.2) = " + str(sigmoid(9.2)))
# + pycharm={"name": "#%%\n"}
def initialize_with_zeros(dim):
"""
此函数为w创建一个维度为(dim,1)的0向量,并将b初始化为0。
参数:
dim - 我们想要的w矢量的大小(或者这种情况下的参数数量)
返回:
w - 维度为(dim,1)的初始化向量。
b - 初始化的标量(对应于偏差)
"""
w = np.zeros(shape=(dim, 1))
b = 0
# 使用断言来确保我要的数据是正确的
assert w.shape == (dim, 1)
assert isinstance(b, float) or isinstance(b, int)
return w, b
# + pycharm={"name": "#%%\n"}
def propagate(w, b, X, Y):
"""
实现前向和后向传播的成本函数及其梯度。
参数:
w - 权重,大小不等的数组(num_px * num_px * 3,1)
b - 偏差,一个标量
X - 矩阵类型为(num_px * num_px * 3,训练数量)
Y - 真正的“标签”矢量(如果非猫则为0,如果是猫则为1),矩阵维度为(1,训练数据数量)
返回:
cost- 逻辑回归的负对数似然成本
dw - 相对于w的损失梯度,因此与w相同的形状
db - 相对于b的损失梯度,因此与b的形状相同
"""
m = X.shape[1]
# 正向传播
A = sigmoid(np.dot(w.T, X) + b) # 计算激活值
cost = (-1 /m) * np.sum(Y * np.log(A) + (1 - Y) * (np.log(1 - A))) # 计算成本
# 反向传播
dw = (1 / m) * np.dot(X, (A - Y).T)
db = (1 / m) * np.sum(A - Y)
# 使用断言确保我的数据是正确的
assert dw.shape == w.shape
assert db.dtype == float
cost = np.squeeze(cost)
assert cost.shape == ()
# 创建一个字典,把dw和db保存起来
grads = {"dw": dw,
"db": db}
return grads, cost
# + pycharm={"name": "#%%\n"}
def optimize(w, b, X, Y, num_iterations, learning_rate, print_cost=False):
"""
此函数通过运行梯度下降算法来优化w和b
参数:
w - 权重,大小不等的数组(num_px * num_px * 3,1)
b - 偏差,一个标量
X - 维度为(num_px * num_px * 3,训练数据的数量)的数组。
Y - 真正的“标签”矢量(如果非猫则为0,如果是猫则为1),矩阵维度为(1,训练数据的数量)
num_iterations - 优化循环的迭代次数
learning_rate - 梯度下降更新规则的学习率
print_cost - 每100步打印一次损失值
返回:
params - 包含权重w和偏差b的字典
grads - 包含权重和偏差相对于成本函数的梯度的字典
成本 - 优化期间计算的所有成本列表,将用于绘制学习曲线。
提示:
我们需要写下两个步骤并遍历它们:
1)计算当前参数的成本和梯度,使用propagate()。
2)使用w和b的梯度下降法则更新参数。
"""
costs = []
for i in range(num_iterations):
grads, cost = propagate(w, b, X, Y)
dw = grads["dw"]
db = grads["db"]
w = w - learning_rate * dw
b = b - learning_rate * db
# 记录成本
if i % 100 == 0:
costs.append(cost)
if print_cost:
print("迭代的次数: %i , 误差值: %f" % (i, cost))
params = {"w": w,
"b": b}
grads = {"dw": dw,
"db": db}
return params, grads, costs
# + pycharm={"name": "#%%\n"}
def predict(w, b, X):
"""
使用学习逻辑回归参数logistic (w,b)预测标签是0还是1,
参数:
w - 权重,大小不等的数组(num_px * num_px * 3,1)
b - 偏差,一个标量
X - 维度为(num_px * num_px * 3,训练数据的数量)的数据
返回:
Y_prediction - 包含X中所有图片的所有预测【0 | 1】的一个numpy数组(向量)
"""
m = X.shape[1]
Y_prediction = np.zeros((1, m))
w = w.reshape(X.shape[0], 1)
# 预测猫在图片中出现的概率
A = sigmoid(np.dot(w.T, X) + b)
for i in range(A.shape[1]):
Y_prediction[0, i] = 1 if A[0, i] > 0.5 else 0
assert Y_prediction.shape == (1, m)
return Y_prediction
# + [markdown] pycharm={"name": "#%% md\n"}
# 就目前而言,我们基本上把所有的东西都做完了,现在我们要把这些函数统统整合到一个model()函数中,届时只需要调用一个model()就基本上完成所有的事了。
# + pycharm={"name": "#%%\n"}
def model(X_train, Y_train, X_test, Y_test,
num_iterations=2000, learning_rate=0.5, print_cost=False):
"""
通过调用之前实现的函数来构建逻辑回归模型
参数:
X_train - numpy的数组,维度为(num_px * num_px * 3,m_train)的训练集
Y_train - numpy的数组,维度为(1,m_train)(矢量)的训练标签集
X_test - numpy的数组,维度为(num_px * num_px * 3,m_test)的测试集
Y_test - numpy的数组,维度为(1,m_test)的(向量)的测试标签集
num_iterations - 表示用于优化参数的迭代次数的超参数
learning_rate - 表示optimize()更新规则中使用的学习速率的超参数
print_cost - 设置为true以每100次迭代打印成本
返回:
d - 包含有关模型信息的字典。
"""
w, b = initialize_with_zeros(X_train.shape[0])
parameters, grads, costs = optimize(w, b, X_train, Y_train, num_iterations, learning_rate, print_cost)
# 从字典“参数”中检索参数w和b
w, b = parameters['w'], parameters['b']
# 预测测试/训练集的例子
Y_prediction_test = predict(w , b, X_test)
Y_prediction_train = predict(w , b, X_train)
# 打印训练后的准确性
print("训练集准确性:" , format(100 - np.mean(np.abs(Y_prediction_train - Y_train)) * 100) ,"%")
print("测试集准确性:" , format(100 - np.mean(np.abs(Y_prediction_test - Y_test)) * 100) ,"%")
d = {"costs" : costs,
"Y_prediction_test" : Y_prediction_test,
"Y_prediciton_train" : Y_prediction_train,
"w" : w,
"b" : b,
"learning_rate" : learning_rate,
"num_iterations" : num_iterations }
return d
# + pycharm={"name": "#%%\n"}
print("====================测试model====================")
d = model(train_set_x, train_set_y, test_set_x, test_set_y, num_iterations=2000, learning_rate=0.005, print_cost=True)
# + pycharm={"name": "#%%\n"}
costs = np.squeeze(d['costs'])
print(costs)
plt.plot(costs)
plt.ylabel('cost')
plt.xlabel('iterations (per hundreds)')
plt.title('Learning rate =' + str(d['learning_rate']))
plt.show()
# + pycharm={"name": "#%%\n"}
learning_rates = [0.01, 0.001, 0.0001]
models = {}
for lr in learning_rates:
print('learning rate is: {}'.format(lr))
models[str(lr)] = model(train_set_x, train_set_y, test_set_x, test_set_y, num_iterations=2000, learning_rate=lr, print_cost=False)
print ('\n' + "-------------------------------------------------------" + '\n')
for lr in learning_rates:
plt.plot(np.squeeze(models[str(lr)]['costs']), label=str(models[str(lr)]['learning_rate']))
plt.ylabel('cost')
plt.xlabel('iterations')
legend = plt.legend(loc='upper center', shadow=True)
frame = legend.get_frame()
frame.set_facecolor('0.90')
plt.show()
# + pycharm={"name": "#%%\n"}
|
assignments/c1-w2/c1-week2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import numpy as np
from copy import copy
import math
import random
import matplotlib.pyplot as plt
from matplotlib.patches import Ellipse
# ランドマークのクラス
class Landmark:
def __init__(self, x, y):
self.pos = np.array([[x],[y]])
def relative_pos(self, pose):
x, y, theta = pose
lx, ly = self.pos[0][0], self.pos[1][0]
distance = math.sqrt((x-lx)**2 + (y-ly)**2)
direction = math.atan2(ly-y, lx-x) - theta
return (distance, direction, lx, ly)
# マップのクラス
class Map:
def __init__(self):
self.landmarks = []
def append_landmark(self, x, y):
self.landmarks.append(Landmark(x, y))
def draw(self):
xs = [e.pos[0][0] for e in self.landmarks]
ys = [e.pos[1][0] for e in self.landmarks]
plt.scatter(xs,ys,s=300,marker="*",label="landmarks",color="orange")
def relative_landmark_positions(self, pose):
positions = []
for i, ln in enumerate(self.landmarks):
distance, direction, lx, ly = ln.relative_pos(pose)
positions.append([distance, direction, lx, ly, i])
return positions
# +
m = Map()
m.append_landmark(-0.5,0.0)
m.append_landmark(0.5,0.0)
m.append_landmark(0.0,0.5)
m.draw()
# -
# ロボットのクラス
class Robot:
def __init__(self, pose):
self.pose = pose
def draw(self):
x,y,theta = self.pose
plt.quiver([x], [y], [math.cos(theta)], [math.sin(theta)],
color="red", label="actual robot motion")
def observation(self, m):
measurements = m.relative_landmark_positions(self.pose)
observations = []
for m in measurements:
distance, direction, lx, ly, i = m
# 視界
if (math.cos(direction) < 0.0): continue
# 観測値にノイズを混ぜる
measured_distance = random.gauss(distance, distance*0.1) # 距離 10%
measured_direction = random.gauss(direction, 5.0/180.0*math.pi) # 方向 5[deg]
observations.append([measured_distance, measured_direction, lx, ly, i])
return observations
def motion_model(self, pos, fw, rot):
# 移動にノイズを混ぜる
actual_fw = random.gauss(fw, fw/10) # 移動距離 10%
dir_error = random.gauss(0.0, math.pi / 180.0 * 3.0) # 前進時方向ズレ 3[deg]
px, py, pt = pos
#前進
x = px + actual_fw * math.cos(pt + dir_error)
y = py + actual_fw * math.sin(pt + dir_error)
# 回転 10%
t = pt + dir_error + random.gauss(rot,rot/10)
# あとで使うので位置の変数を変えずに移動量を返す
return np.array([x,y,t])
robot = Robot(np.array([0.1, 0.2, math.pi*20.0/180]) )
robot.draw()
m.draw()
# +
observations = robot.observation(m)
print(observations)
fig = plt.figure(0,figsize=(8, 8))
sp = fig.add_subplot(111, aspect='equal')
sp.set_xlim(-1.0,1.0)
sp.set_ylim(-0.5,1.5)
for observation in observations:
x,y,theta = robot.pose
distance, direction,lx,ly, i = observation
lx = distance*math.cos(theta + direction) + x
ly = distance*math.sin(theta + direction) + y
plt.plot([robot.pose[0], lx],[robot.pose[1], ly],color="pink")
c = math.cos(theta + direction)
s = math.sin(theta + direction)
rot = np.array([[ c, -s],
[ s, c]])
err_robot = np.array([[(distance*0.1)**2,0.0],
[0.0,(distance*math.sin(5.0/180.0*math.pi))**2]])
err_world = (rot).dot(err_robot).dot((rot).T)
eig_vals,eig_vec = np.linalg.eig(err_world)
v1 = eig_vals[0] * eig_vec[:, 0]
v2 = eig_vals[1] * eig_vec[:, 1]
v1_direction = math.atan2(v1[1], v1[0])
elli = Ellipse([lx, ly], width=3*math.sqrt(np.linalg.norm(v1)), height=3*math.sqrt(np.linalg.norm(v2)), angle=v1_direction/3.14*180)
elli.set_alpha(0.2)
sp.add_artist(elli)
robot.draw()
m.draw()
# -
class LandmarkEstimation:
def __init__(self):
self.pos = np.array([[0.0], [0.0]])
# 最初は大きな共分散行列にしておく
self.cov = np.array([[1000000.0**2, 0.0],
[0.0, 1000000**2]])
class Particle:
def __init__(self, pose, w):
self.w = w
self.pose = pose
self.map = [LandmarkEstimation(), LandmarkEstimation(), LandmarkEstimation()] #数は3で既知とする
def motion_update(self, fw, rot, robot):
self.pose = robot.motion_model(self.pose, fw, rot)
def measurement_update(self, measurement):
x,y,theta = self.pose
distance, direction, lx, ly, i = measurement
ln = self.map[i]
lx = distance * math.cos(theta + direction) + x
ly = distance * math.sin(theta + direction) + y
## 重みの更新
delta = np.array([[x], [y]]) - np.array([[lx], [ly]])
coef = 2*math.pi * math.sqrt(np.linalg.det(ln.cov))
inexp = -0.5 * (delta.T.dot(np.linalg.inv(ln.cov))).dot(delta)
self.w *= 1.0/coef * math.exp(inexp)
## 地図の書き換え
z = np.array([[lx], [ly]])
c = math.cos(theta + direction)
s = math.sin(theta + direction)
rot = np.array([[ c, -s],
[ s, c]])
err_robot = np.array([[(distance*0.1)**2, 0.0],
[0.0,(distance*math.sin(5.0/180.0*math.pi))**2]])
err_world = (rot).dot(err_robot).dot((rot).T)
ln.cov = np.linalg.inv( np.linalg.inv(ln.cov) + np.linalg.inv(err_world) )
K = (ln.cov).dot(np.linalg.inv(err_world))
ln.pos += K.dot( z - ln.pos )
def draw(self, i):
fig = plt.figure(i, figsize=(4, 4))
sp = fig.add_subplot(111, aspect='equal')
sp.set_xlim(-1.0, 1.0)
sp.set_ylim(-0.5, 1.5)
m.draw()
x, y, theta = self.pose
plt.quiver([x], [y], [math.cos(theta)], [math.sin(theta)], color="red", label="actual robot motion")
for e in self.map:
eig_vals, eig_vec = np.linalg.eig(e.cov)
v1 = eig_vals[0] * eig_vec[:, 0]
v2 = eig_vals[1] * eig_vec[:, 1]
v1_direction = math.atan2(v1[1], v1[0])
x, y = e.pos
elli = Ellipse([x,y], width=3*math.sqrt(np.linalg.norm(v1)), height=3*math.sqrt(np.linalg.norm(v2)), angle=v1_direction/3.14*180)
elli.set_alpha(0.5)
sp.add_artist(elli)
# +
import copy
class FastSLAM():
def __init__(self, pose):
self.particles = [Particle(pose,1.0/100) for i in range(100)]
def draw(self):
for (i, p) in enumerate(self.particles):
if i == 3: return
p.draw(i)
def motion_update(self, fw, rot, robot):
for p in self.particles:
p.motion_update(fw, rot, robot)
def measurement_update(self, measurement):
for p in self.particles:
p.measurement_update(measurement)
self.resampling()
def resampling(self):
num = len(self.particles) # numはパーティクルの個数
ws = [e.w for e in self.particles] # 重みのリストを作る
#重みの和がゼロに丸め込まれるとサンプリングできなくなるので小さな数を足しておく
if sum(ws) < 1e-100:
ws = [e + 1e-100 for e in ws]
# パーティクルのリストから、weightsのリストの重みに比例した確率で、num個選ぶ
ps = random.choices(self.particles, weights=ws, k=num)
# 選んだリストからパーティクルを取り出し、パーティクルの姿勢から重み1/numの新しいパーティクルを作成
self.particles = [copy.deepcopy(e) for e in ps]
# +
robot.pose = np.array([0.0, 0.0, 0.0])
slam = FastSLAM(robot.pose)
def one_step(m):
slam.motion_update(0.2, math.pi / 180.0 * 20, robot)
robot.pose = robot.motion_model(robot.pose, 0.2, math.pi / 180.0 * 20)
measurements = robot.observation(m)
for m in measurements:
slam.measurement_update(m)
n = 30
for i in range(n):
one_step(m)
print(str(n) + "step後の地図")
slam.draw()
# -
|
fast_slam.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PyCharm (btcpaper)
# language: python
# name: pycharm-8016b31a
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn.decomposition import PCA
from sklearn.preprocessing import Normalizer, StandardScaler, MinMaxScaler, RobustScaler
from sklearn.pipeline import Pipeline
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_absolute_error, r2_score, mean_squared_error
# %matplotlib inline
data=pd.read_csv('BTC_Data_736_features_raw.csv',sep=',')
data.interpolate(axis=0,inplace=True)
interval3=(data['Date'] >= '2013/04/01') & (data['Date'] <= '2019/12/31')
df=data.loc[interval3]
df.head()
X=df.iloc[:,2:]
X.head()
estimators=[]
estimators.append(['minmax',MinMaxScaler(feature_range=(-1,1))])
scale=Pipeline(estimators)
X=scale.fit_transform(X)
pca = PCA(n_components=6,random_state=7)
pca.fit(X)
pca.explained_variance_ratio_
np.cumsum(pca.explained_variance_ratio_)
#Plotting the Cumulative Summation of the Explained Variance
plt.figure()
plt.plot(np.cumsum(pca.explained_variance_ratio_))
plt.xlabel('Number of Components')
plt.ylabel('Variance (%)') #for each component
plt.title('Explained Variance')
plt.show()
df_pca=pd.DataFrame(pca.components_).transpose()
df_pca.columns=['comp_1','comp_2','comp_3','comp_4','comp_5','comp_6']
y=df.iloc[:,1:2]
y.reset_index(drop=True,inplace=True)
df_pca['priceUSD']=y
df_pca.head()
df_pca.to_csv('pca_75_reg.csv',index=False)
# # Classification
one=data['priceUSD'].shift(-1,fill_value=1)
df['one']=one.loc[interval3]
df['difference']=((df['one']-df['priceUSD'])/df['priceUSD'])*100
df.reset_index(drop=True,inplace=True)
category=[]
for x in range(len(df['difference'])):
if df['difference'][x]>=0:
category.append(1)
else:
category.append(0)
sum(category)
df['category']=pd.DataFrame(category)
df.tail()
df['priceUSD']=df['category']
df.drop(columns=['category','one','difference'],inplace=True)
df.head(3)
X=df.iloc[:,2:]
X.head()
estimators=[]
estimators.append(['minmax',MinMaxScaler(feature_range=(-1,1))])
scale=Pipeline(estimators)
X=scale.fit_transform(X)
pca = PCA(n_components=50,random_state=7)
pca.fit(X)
pca.explained_variance_ratio_
np.cumsum(pca.explained_variance_ratio_)
#Plotting the Cumulative Summation of the Explained Variance
plt.figure()
plt.plot(np.cumsum(pca.explained_variance_ratio_))
plt.xlabel('Number of Components')
plt.ylabel('Variance (%)') #for each component
plt.title('Explained Variance')
plt.show()
df_pca=pd.DataFrame(pca.components_).transpose()
# + active=""
# df_pca.columns=['comp_1','comp_2','comp_3','comp_4','comp_5','comp_6']
# -
y=df.iloc[:,1:2]
y.reset_index(drop=True,inplace=True)
df_pca['priceUSD']=y
df_pca.head()
df_pca.to_csv('pca_75_clas.csv',index=False)
|
PCA.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python2
# ---
import numpy as np
import pandas as pd
from pandas import DataFrame, Series
import pandas.util.testing as tm
tm.N = 3
def unpivot(frame):
N, K = frame.shape
data = {'value': frame.values.ravel('F'),
'variable': np.asarray(frame.columns).repeat(N),
'date': np.tile(np.asarray(frame.index), K)}
return DataFrame(data, columns=['date', 'variable', 'value'])
dframe = unpivot(tm.makeTimeDataFrame())
dframe
dframe_piv = dframe.pivot('date', 'variable', 'value')
dframe_piv
|
l6/Lecture 34 - Pivoting.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Time
# ## Time structure and local time string
import time
now_time = time.localtime()
print(now_time)
print(time.asctime(now_time))
print(time.tzname)
# ## Create time
# +
print('time.time(): {}, time.process_time():{}'.format(time.time(), time.process_time()))
# mktime
ti = time.localtime(time.mktime((2020,8,8,0,0,0,0,0,0)))
print(ti)
print(time.asctime(ti))
# strptime
ti2 = time.strptime('2019年11月29日', '%Y年%m月%d日')
print(ti2)
# -
# ## Format time
print(time.strftime('%Y年%m月%d日', (2020,8,8,0,0,0,0,0,0)))
# # Calendar
import calendar
calendar.setfirstweekday(6)
this_month = calendar.month(now_time.tm_year, now_time.tm_mon, 5, 2)
print(this_month)
weekdaynames = ['星期一','星期二','星期三','星期四','星期五','星期六','星期天']
print(weekdaynames[calendar.weekday(2019,12,5)])
# # Datetime
# +
import datetime
d1 = datetime.date(2019, 2, 15)
d2 = d1 + datetime.timedelta(50)
print('今天是:', datetime.date.today())
print('此刻本地时间是:', datetime.datetime.now())
print('{} ---> {}'.format(d1, d2))
print(d1.strftime('%Y/%m/%d'))
print('{}年{}月{}日'.format(d2.year, d2.month, d2.day))
print('--------------------------------------------------------------------------------')
d3 = d1.replace(month=d1.month+3)
print(d3)
print(d3.timetuple())
dd = d3 - d1
print('>>> The duration from {0} to {1} is {2} day(s).'.format(
d1, d3, dd.days))
|
test/py_time_test.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Using gradient hidden states on regression
# ## Purpose
# * Kalman filtered states have previously been used for the regressions unsuccessfully.
# * Numerical gradient will now instead be used.
# ## Methodology
# * Load simulated data generated by: [12.05_regression_simulated_data_simple_nonlinear.ipynb](12.05_regression_simulated_data_simple_nonlinear.ipynb)
# * Determine velocity and acceleration and compared...
# ## Setup
# +
# # %load imports.py
## Local packages:
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
# %config Completer.use_jedi = False ## (To fix autocomplete)
## External packages:
import pandas as pd
pd.options.display.max_rows = 999
pd.options.display.max_columns = 999
pd.set_option("display.max_columns", None)
import numpy as np
import os
import matplotlib.pyplot as plt
#if os.name == 'nt':
# plt.style.use('presentation.mplstyle') # Windows
import plotly.express as px
import plotly.graph_objects as go
import seaborn as sns
import sympy as sp
from sympy.physics.mechanics import (dynamicsymbols, ReferenceFrame,
Particle, Point)
from sympy.physics.vector.printing import vpprint, vlatex
from IPython.display import display, Math, Latex
from src.substitute_dynamic_symbols import run, lambdify
import pyro
import sklearn
import pykalman
from statsmodels.sandbox.regression.predstd import wls_prediction_std
import statsmodels.api as sm
from scipy.integrate import solve_ivp
## Local packages:
from src.data import mdl
from src.symbols import *
from src.parameters import *
import src.symbols as symbols
from src import prime_system
from src.models import regression
from src.visualization.regression import show_pred
from src.visualization.plot import track_plot
## Load models:
# (Uncomment these for faster loading):
import src.models.vmm_simple_nonlinear as vmm
from src.data.transform import transform_to_ship
from src.data.wpcc import ship_parameters, df_parameters, ps, ship_parameters_prime, meta_data
from src.data.lowpass_filter import lowpass_filter
# -
# ## Load test
# +
#id=22773
#id=22616
id=22774
#id=22770
df = mdl.load_run(id=id, dir_path='../data/raw/')
df.index = df.index.total_seconds()
df = df.iloc[0:-100].copy()
df.index-=df.index[0]
df['t'] = df.index
df['thrust'] = df['Prop/PS/Thrust'] + df['Prop/SB/Thrust']
df.sort_index(inplace=True)
# -
# # Ship parameters
# +
T_ = (meta_data.TA + meta_data.TF)/2
L_ = meta_data.lpp
m_ = meta_data.mass
rho_ = meta_data.rho
B_ = meta_data.beam
CB_ = m_/(T_*B_*L_*rho_)
I_z_ = m_*meta_data.KZZ**2
#I_z_ = 900
ship_parameters = {
'T' : T_,
'L' : L_,
'CB' :CB_,
'B' : B_,
'rho' : rho_,
'x_G' : 0, # motions are expressed at CG
'm' : m_,
'I_z': I_z_,
'volume':meta_data.Volume,
}
ps = prime_system.PrimeSystem(**ship_parameters) # model
scale_factor = meta_data.scale_factor
ps_ship = prime_system.PrimeSystem(L=ship_parameters['L']*scale_factor, rho=meta_data['rho']) # ship
# -
# ## Brix parameters
mask = df_parameters['prime'].notnull()
index = df_parameters.loc[mask,'prime'].index
coefficients=vmm.simulator.get_all_coefficients(sympy_symbols=False)
missing_coefficients = set(coefficients) - set(index)
missing_coefficients
mask = df_parameters['prime'].notnull()
df_parameters.loc[mask,'prime']
ship_parameters['x_G']
# ### Estimate hidden states with gradients
# +
df_measurement = df.copy()
t_ = df_measurement.index
ts = np.mean(np.diff(t_))
fs=1/ts
suffix = ['','1d','2d']
for i in range(2):
for key in ['x0','y0','z0','psi']:
y = df_measurement[f'{key}{suffix[i]}'].copy()
y_filt = lowpass_filter(data=y, cutoff=0.5, fs=fs, order=4)
df_measurement[f'{key}{suffix[i+1]}'] = np.gradient(y_filt, t_)
df_measurement = transform_to_ship(df=df_measurement)
df_measurement=df_measurement.iloc[3000:-1000].copy()
df_measurement['V'] = np.sqrt(df_measurement['u']**2 + df_measurement['v']**2)
df_measurement['U'] = df_measurement['V']
df_measurement['beta'] = -np.arctan2(df_measurement['v'],df_measurement['u'])
interesting = ['x0','y0','z0','psi','u','v','r','u1d','v1d','r1d','delta','thrust','U']
df_measurement = df_measurement[interesting].copy()
# -
for key in df_measurement:
fig,ax=plt.subplots()
fig.set_size_inches(10,2)
df_measurement.plot(y=key, ax=ax)
# ## Regression
# +
df = ps.prime(df_measurement, U=df_measurement['U']) # Note primed!
for key in df_measurement:
fig,ax=plt.subplots()
fig.set_size_inches(10,2)
df.plot(y=key, ax=ax)
# -
# ## N
# +
N_ = sp.symbols('N_')
diff_eq_N = regression.DiffEqToMatrix(ode=vmm.N_qs_eq.subs(N_qs,N_),
label=N_, base_features=[delta,u,v,r])
# +
X = diff_eq_N.calculate_features(data=df)
y = diff_eq_N.calculate_label(y=df['r1d'])
model_N = sm.OLS(y,X)
results_N = model_N.fit()
show_pred(X=X,y=y,results=results_N, label=r'$\dot{r}$')
# -
# ## Y
Y_ = sp.symbols('Y_')
diff_eq_Y = regression.DiffEqToMatrix(ode=vmm.Y_qs_eq.subs(Y_qs,Y_),
label=Y_, base_features=[delta,u,v,r])
# +
X = diff_eq_Y.calculate_features(data=df)
y = diff_eq_Y.calculate_label(y=df['v1d'])
model_Y = sm.OLS(y,X)
results_Y = model_Y.fit()
show_pred(X=X,y=y,results=results_Y, label=r'$\dot{v}$')
# -
# ## X
X_ = sp.symbols('X_')
diff_eq_X = regression.DiffEqToMatrix(ode=vmm.X_qs_eq.subs(X_qs,X_),
label=X_, base_features=[delta,u,v,r,thrust])
# +
X = diff_eq_X.calculate_features(data=df)
y = diff_eq_X.calculate_label(y=df['u1d'])
model_X = sm.OLS(y,X)
results_X = model_X.fit()
show_pred(X=X,y=y,results=results_X, label=r'$\dot{u}}$')
# -
results_summary_X = regression.results_summary_to_dataframe(results_X)
results_summary_Y = regression.results_summary_to_dataframe(results_Y)
results_summary_N = regression.results_summary_to_dataframe(results_N)
# ### Decoupling
# +
u1d,v1d,r1d = sp.symbols('u1d, v1d, r1d')
u_,v_,r_ = sp.symbols('u, v, r')
X_qs_, Y_qs_, N_qs_ = sp.symbols('X_qs, Y_qs, N_qs')
X_eq = vmm.X_eom.subs([
(X_force,sp.solve(vmm.fx_eq,X_force)[0]),
])
Y_eq = vmm.Y_eom.subs([
(Y_force,sp.solve(vmm.fy_eq,Y_force)[0]),
])
N_eq = vmm.N_eom.subs([
(N_force,sp.solve(vmm.mz_eq,N_force)[0]),
])
subs = [
#(X_qs,X_qs_),
#(Y_qs,Y_qs_),
#(N_qs,N_qs_),
(u1d,u1d),
(v1d,v1d),
(r1d,r1d),
(u,u_),
(v,v_),
(r,r_),
]
eq_X_ = X_eq.subs(subs)
eq_Y_ = Y_eq.subs(subs)
eq_N_ = N_eq.subs(subs)
A,b = sp.linear_eq_to_matrix([eq_X_,eq_Y_,eq_N_],[u1d,v1d,r1d])
# -
subs = {value:key for key,value in p.items()}
A_ = A*sp.matrices.MutableDenseMatrix([A_coeff,B_coeff,C_coeff])
A_lambda=lambdify(A_.subs(subs))
# +
A_coeff_ = results_summary_X['coeff']
B_coeff_ = results_summary_Y['coeff']
C_coeff_ = results_summary_N['coeff']
coeffs = run(A_lambda,A_coeff=A_coeff_.values, B_coeff=B_coeff_.values, C_coeff=C_coeff_.values,
**df_parameters['prime'], **ship_parameters_prime)
# -
# The way that the regression is formulated, inertial forces, such as centrifugal force will be included into the derivatives (I think) which means that centrifugal force : $-m \cdot r \cdot u$ will be included into $Y_{ur}$ coefficient. This coefficient is therefore not pure hydrodynamic, and can potentially be counted twice..?
# The coefficients are recalculated below to avooid this:
# +
results_summary_X['decoupled'] = coeffs[0][0]
results_summary_Y['decoupled'] = coeffs[1][0]
results_summary_N['decoupled'] = coeffs[2][0]
x_G_ = ship_parameters_prime['x_G']
m_ = ship_parameters_prime['m']
results_summary_X.loc['Xrr','decoupled']+=(-m_*x_G_)
results_summary_X.loc['Xvr','decoupled']+=(-m_)
results_summary_Y.loc['Yur','decoupled']+=m_
results_summary_N.loc['Nur','decoupled']+=m_*x_G_
#results_summary_X.loc['Xr','decoupled']+=(-m_*x_G_)
#results_summary_Y.loc['Yr','decoupled']+=m_
#results_summary_N.loc['Nr','decoupled']+=m_*x_G_
# -
results_summary_N
# ## Add the regressed parameters
# Hydrodynamic derivatives that depend on acceleration cannot be obtained from the VCT regression. They are however essential if a time simulation should be conducted. These values have then been taken from Brix semi empirical formulas for the simulations below.
# +
df_parameters_all = df_parameters.copy()
for other in [results_summary_X, results_summary_Y, results_summary_N]:
df_parameters_all = df_parameters_all.combine_first(other)
df_parameters_all.rename(columns={'decoupled':'regressed'}, inplace=True)
df_parameters_all.drop(columns=['brix_lambda'], inplace=True)
df_parameters_all['regressed'] = df_parameters_all['regressed'].combine_first(df_parameters_all['prime']) # prefer regressed
#df_parameters_all['regressed'].fillna(0,inplace=True)
# +
df_plot_parameters = df_parameters_all[['regressed','prime']].copy()
mask = ((df_plot_parameters['prime'].abs() > 0) &
(df_plot_parameters['prime'].abs() < 0.8)
)
df_plot_parameters.loc[mask].plot(kind='bar')
mask = (
(df_plot_parameters['prime'].abs() > 0.8)
)
if mask.any():
df_plot_parameters.loc[mask].plot(kind='bar')
# -
# # Simulation
df_measurement.head()
# +
parameters=df_parameters_all['regressed'].copy()
result_regression = vmm.simulator.simulate(df_=df_measurement, parameters=parameters, ship_parameters=ship_parameters,
control_keys=['delta','thrust'],
primed_parameters=True,
prime_system=ps,
name='regressed')
# -
result_regression.track_plot()
result_regression.plot();
|
notebooks/14.02_hidden_states_gradient_regression.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Color Threshold, Blue Screen
# ### Import resources
# +
import matplotlib.pyplot as plt
import numpy as np
import cv2
# %matplotlib inline
# -
# ### Read in and display the image
# +
# Read in the image
image = cv2.imread('images/pizza_bluescreen.jpg')
# Print out the type of image data and its dimensions (height, width, and color)
print('This image is:', type(image),
' with dimensions:', image.shape)
# +
# Make a copy of the image
image_copy = np.copy(image)
# Change color to RGB (from BGR)
image_copy = cv2.cvtColor(image_copy, cv2.COLOR_BGR2RGB)
# Display the image copy
plt.imshow(image_copy)
# -
# ### Define the color threshold
## TODO: Define the color selection boundaries in RGB values
# play around with these values until you isolate the blue background
lower_blue = np.array([0,0,200])
upper_blue = np.array([250,250,255])
# ### Create a mask
# +
# Define the masked area
mask = cv2.inRange(image_copy, lower_blue, upper_blue)
# Vizualize the mask
plt.imshow(mask, cmap='gray')
# +
# Mask the image to let the pizza show through
masked_image = np.copy(image_copy)
masked_image[mask != 0] = [0, 0, 0]
# Display it!
plt.imshow(masked_image)
# -
# ### Mask and add a background image
# +
# Load in a background image, and convert it to RGB
background_image = cv2.imread('images/space_background.jpg')
background_image = cv2.cvtColor(background_image, cv2.COLOR_BGR2RGB)
# Crop it to the right size (514x816)
crop_background = background_image[0:514, 0:816]
# Mask the cropped background so that the pizza area is blocked
crop_background[mask == 0] = [0, 0, 0]
# Display the background
plt.imshow(crop_background)
# -
# ### Create a complete image
# +
# Add the two images together to create a complete image!
complete_image = masked_image + crop_background
# Display the result
plt.imshow(complete_image)
# -
|
1_1_Image_Representation/3. Blue Screen.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # **1. Handout Part3**
# ## **1.1. Introduction**
# > 지난 Chapter에서는 간단한 Machine-learning Method를 활용하여 실제로 레일 온도를 예측하는 모델을 생성해 봤지만 성능이 좋지 못했습니다.
# ><br><br> 이번 시간에는 Tree 기반의 모델을 사용하여 레일의 온도를 예측해 보고 Model의 성능을 높이기 위한 방법들을 공부해 봅시다.
#
# ***
# <br>
#
# ## **1.2. Objective**
# > 본 과제의 목적은 다음과 같습니다. <br>
# > 1. Random Forest 모델 생성
# > 2. 데이터 전처리를 통한 모델 성능 향상
# > 3. 새로운 Feature 생성을 통한 모델 성능 향상
# <br>
# ***
# <br>
# # **2. EXERCISE**
# ## **2.1. Loading Library**
# > 이번시간에도 똑같이 library를 로딩합니다.<br>
# > 필요한 library들은 pandas, matplolib, sklearn의 svm, randomforest 입니다.<br>
# > svm 과 RandomForest는 회귀를 사용합니다.<br>
# >문제 : **이전 시간의 답과 Kaggle과 google의 힘을 빌려 라이브러리를 import 해보세요(svm, randomforest)**
# + pycharm={"name": "#%%\n"}
import pandas as pd
import matplotlib.pyplot as plt
# -
# ## **2.2. Loading Data File**
# > library들을 불러왔으면, 파일을 입력해보자!<br>
# > 이번에도 북쪽 데이터만을 사용할겁니다!<br><br>
# > 데이터를 불러오는 것은 항상 해야하는 일이기 때문에 다시 한 번 복습해봅시다<br>
#
# > **북쪽 레일의 온도를 대표하는 지점은 I-7 지점입니다.(이전 과제의 레일 데이터를 참고하세요)**<br>
# >
# >**본 예측 모델은 I-7의 온도를 기후데이터만으로 예측하는 것을 목표로 합니다.**<br><br>
# > 머신러닝 모델을 만들기 위해서는 feature와 target으로 데이터를 나누어야 합니다.<br>
# > 이를 위해서 다음 문제를 해결해 봅시다.<br>
# > 문제 : 데이터를 로드하고 feature와 target으로 데이터를 나누자<br>
# > - rd에 전체 데이터를 읽어오자
# > - loc또는 iloc을 활용하여, x에는 기후데이터를, y에는 target인 i-7값을 입력하자
# ***
#
# + pycharm={"name": "#%%\n"}
rd = pd.read_excel()
x = rd.loc[]
y = rd.loc[]
# -
# ## **2.3. Data Split and fitting**
# > 이전 시간에 했던 datasplit를 다시 복습해봅시다.<br>
# > 문제 : **trainset의 비율을 70%로 잡고 X_train X_val y_train y_val로 데이터를 나누시오!**
#
# + pycharm={"name": "#%%\n"}
# -
# ## **2.4. RandomForest**
# > 이제 RandomForest의 회귀모델을 만들어볼 차례입니다 <br>
# > 문제 : **X_train과 y_train을 통해 모델을 RandomForest을 생성하시오**<br>
# > 문제 : **X_val과 y_val을 통해 모델을 평가하시오 이때의 평가지표는 (MAE, R2, RMSE)를 사용!**<br>
# > - MAE, R2, RMSE에 대해서는 공부해볼 것!<br>
# + pycharm={"name": "#%%\n"}
# -
# ## **2.5. Prediction**
# > 만들어진 RandomForest 모델을 통해 실제로 본 모델이 얼마나 잘 예측하는지까지 평가를 해보았습니다.<br>
# > 그렇다면, 모델의 예측값이 어떻게 나타나는지 한번 visualization해보는게 좋겠죠?<br><br>
# > 위에서 원 데이터를 로드한 rd에 대해서 <br>
# > 2018.08.09~2018.08.12 까지의 기후데이터와 i-7의 온도데이터를 로드합시다.<br>
# > 변수명은 원하는대로 진행해주시면 됩니다.<br>
# > 기후데이터와 앞서 만든 모델을 통해 레일 온도를 예측하여, predict_rail_temp에 저장합시다.<br>
# > 여기까지 따라하셨다면 아래의 문제를 해결하시오!<br><br>
# > 문제 : **matplotlib를 통해 실제 i-7값과 예측된 predict_rail_temp를 동시에 나타나도록 코딩하라.**<br>
#
#
# + pycharm={"name": "#%%\n"}
# -
# ## **2.6. Comparison to SVM**
# > 이전에 만들었던 SVM 회귀 모델과의 성능을 비교해봅시다.<br>
# > SVM 회귀 모델의 작성하여서 MAE, RMSE, R2_Score를 비교해봅시다.<br>
# > 문제 : **평가지표들을 이용해서 SVM과 RandomForest 성능을 비교하고 성능의 차이가 나는 이유를 google 등을 이용해 서술하시오.**<br>
# >(주의 : 이유를 알게 되면 여러분은 나무와 사랑에 빠질 가능성이 있다.)
# >
# + pycharm={"name": "#%%\n"}
# -
# ## **2.7. Data Normalization**
# > Data Normalization은 일부 모델에서는 필수적입니다.<br>
# > Sklearn에서는 다양한 Normalization 함수를 제공합니다.<br>
# > google에 이에 대한 설명을 해주기 때문에 data normalization 또는 데이터 정규화로 검색해서 자세한 내용을 공부해보세요<br>
# > 문제 : **standardization 과 minmaxscaler를 각각 해보세요**
# > 문제 : **정규화를 하지 않은 경우와 성능을 비교하시오(성능지표 이용)**
# + pycharm={"name": "#%%\n"}
# -
# ## **2.8. Feature generation**
# > 일부 응용에서는 모델의 성능을 올리기 위해 새로운 Feature를 생성해야만 합니다.<br>
# > 특히, 현재 Feature만으로 현상을 명확히 설명할 수 없을 경우 필수적입니다.<br>
# > 기존의 논문 스터디를 통해 우리는 태양의 방사량과 위치가 레일의 온도에 영향을 미치는 것을 알았습니다.<br>
# > 하지만 현재 Feature에는 이를 반영하기 위한 Feature가 존재하지 않습니다.<br>
# > 따라서 성능이 높은 모델을 만들기 위해서 새로운 Feature를 생성해야합니다.<br>
#
#
#
# ### **2.8.1 Feature generation**
# > TSI는 Total solar irradiation를 의미합니다. <br>
# > 문제 : **google 검색, 코드 분석 및 논문 스터디를 통해 'TSI','azimuth','altitude'에 대해 정리해오세요(한글파일로 개념, 계산식 정리)**<br>
# > (azimuth, altitude, TSI에 대한 코드는 feature_preprocessor.py에 있습니다. (azimuth = az, altitude = al)<br>
# > 문제 : **작성되어 있는 feature_preprocessor.py를 load하여 TSI, azimuth, altitude를 데이터에 추가해보세요.**<br>
# >(단순 복붙 금지, .py 파일 import 방법 찾아볼것)
# + pycharm={"name": "#%%\n"}
# -
# ## **2.9. Improve model performance**
# > 위에서 생성한 Feature 3개를 추가하여 모델을 학습시키고 성능을 비교해봅시다.<br>
# > 문제 : **TSI, azimuth, altitude를 기존에 데이터에 추가하여 SVM, RandomForest 모델을 생성해봅시다.**<br>
# > 문제 : **TSI, azimuth, altitude가 포함된 모델과 그렇지 않은 모델의 성능을 비교해보세요.(총 4가지 모델 비교)**
#
# + pycharm={"name": "#%%\n"}
|
handout3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # U-Net for multi-class semantic segmentation
#
# First load all the libraries
#
#
# tensorboard --logdir=~/roaddetection/notebooks/networks/logs/
# +
# %load_ext autoreload
# %autoreload 2
import numpy as np
np.random.seed(42)
#from keras.layers import merge
from src.models.catdata import *
from src.models.catmodel import *
from src.models.catsegnet import *
from src.models.catloss import *
from src.data.utils import get_tile_prefix
from src.models.metrics_img import auc_roc, auc_pr, auc_pr_multiclass, dummy_metric
#import rasterio.plot as rioplot
import matplotlib
import matplotlib.pyplot as plt
#import matplotlib.image as mpimg
from pathlib import Path
import os, shutil
import sys
# %matplotlib inline
import pandas as pd
import math
from sklearn.utils import shuffle
from sklearn.model_selection import train_test_split
from keras.callbacks import RemoteMonitor, TensorBoard, ReduceLROnPlateau, EarlyStopping, History
from keras import backend as keras
sys.path.append("/home/ubuntu/roaddetection/")
# -
# # Create the train, validation and test data sets
#
# To increase the U-Nets learning capacity, it is helpful to **first** train on high road-pixel density data. This is achived by creating the corresponding image-sets according to the requirements.
#
# In a **second** training step all the images, e.g. also low and even zero road-pixel density data is considered.
#
# ### Define the paths where the data is stored
# +
original_dataset_dir = "../../data/train_raw"
raw_images_path = "../../data/raw/images"
dirs = []
base_dir = "../../data"
train_dir = os.path.join(base_dir, "train")
dirs.append(train_dir)
validation_dir = os.path.join(base_dir, "validate")
dirs.append(validation_dir)
test_dir = os.path.join(base_dir, "test")
dirs.append(test_dir)
print(train_dir)
# -
# ### Create the directories if they don't exist
for directory in dirs:
if not os.path.exists(directory):
os.mkdir(directory)
# ### Define a function which takes care of the assignement of image names to one of the three sets accoring to the desired pixel density of each class
# +
r = 5
def selectData(tNoRoad = 0.95, tPRoad = 0, tURoad = 0, tEmpty = 0):
df = pd.read_csv('../visualize_imagery/numOfPixPerClassPerTile_256.csv').drop(['Unnamed: 0'], axis = 1)
#df = df[(df['region'] == 'Borneo')]
any_RP = df[((df['relative_noRoad']) < tNoRoad)
& ((df['relative_pavedRoad']) > 0)
& ((df['relative_unpavedRoad']) > 0)]
unpaved_RP = df[((df['relative_noRoad']) < tNoRoad)
& ((df['relative_pavedRoad']) == 0)
& ((df['relative_unpavedRoad']) > tURoad)]
paved_RP = df[((df['relative_noRoad']) < tNoRoad)
& ((df['relative_pavedRoad']) > tPRoad)
& ((df['relative_unpavedRoad']) == 0)]
no_RP = df[(df['relative_noRoad'] == 1)].sample(frac=tEmpty, random_state=r)
assert len(any_RP.merge(unpaved_RP)) == 0
assert len(unpaved_RP.merge(paved_RP)) == 0
assert len(paved_RP.merge(no_RP)) == 0
assert len(no_RP.merge(any_RP)) == 0
sdf = any_RP
if tEmpty > 0:
sdf = pd.concat([sdf, no_RP])
if tPRoad > 0:
sdf = pd.concat([sdf, paved_RP])
if tURoad > 0:
sdf = pd.concat([sdf, unpaved_RP])
sdf = shuffle(sdf, random_state=r).sample(frac=.4, random_state=r).reset_index(drop=True)
train_tmp, test = train_test_split(sdf, test_size=0.2, random_state=r)
train, valid = train_test_split(train_tmp, test_size=0.2, random_state=r)
return train, valid, test
# -
# ### Create the dataframes assigning the image names to the corresponding classes
train, valid, test = selectData(tNoRoad = .95)#tURoad = 0.06)
print("Train set")
print(train.region.value_counts())
print("")
print("Validation set")
print(valid.region.value_counts())
print("")
print("Test set")
print(test.region.value_counts())
# ### Define the function which copies the data according to the three dataframes
# +
def should_make_tiles_from(r_analytic_name):
is_analytic_tif = r_analytic_name.endswith(
('AnalyticMS.tif', 'AnalyticMS_SR.tif', 'AnalyticMS.tiff', 'AnalyticMS_SR.tiff')
)
return is_analytic_tif
def copy(fnames, src_dir):
for name in fnames:
for file_type in ["sat", "map", "sat_rgb"]:
src = os.path.join(original_dataset_dir, file_type, name)
dest = os.path.join(src_dir, file_type, name)
if(os.path.exists(src)):
shutil.copy(src, dest)
def make_datasets(show_progress = False):
file_prefixes = [ get_tile_prefix(r_analytic.name)
for r_analytic in Path(raw_images_path).iterdir()
if should_make_tiles_from(r_analytic.name)
]
# copy files to train dir
train_fnames = train.name.values
copy(train_fnames, train_dir)
print("Create train data.")
# copy files to validation dir
validation_fnames = valid.name.values
copy(validation_fnames, validation_dir)
print("Create validation data.")
# copy files to test dir
test_fnames = test.name.values
copy(test_fnames, test_dir)
print("Create test data.")
# print overview
if show_progress == True:
for directory in dirs:
for file_type in ["sat", "map", "sat_rgb"]:
target = os.path.join(directory, file_type)
print(target, ":", len(os.listdir(target)))
print("Done.")
# -
# ### Copy the data
# +
#make_datasets()
# -
# # Create class weights accoriding to their train statistics
#
# This is recommended as the classes are imbalanced (more no-road pixels than road pixels in each image)
labeldict = {
0: train.numPixel_noRoad.mean(),
1: train.numPixel_pavedRoad.mean(),
2: train.numPixel_unpavedRoad.mean()
}
def create_class_weight(labels_dict,mu=0.25):
total = sum(labels_dict.values())
keys = labels_dict.keys()
class_weight = dict()
weights = []
for key in keys:
score = math.log(mu*total/float(labels_dict[key]))
class_weight[key] = score if score > 1.0 else 1.0
weights.append(score if score > 1.0 else 1.0)
return weights
# # Define the data generator to flow the images from the directory while training
def data_generator(batch_size, data_gen_args, data_dir, imgdatagen_dir, target_size):
return trainGenerator(
batch_size,
data_dir,
'sat',
'map',
data_gen_args,
save_to_dir = imgdatagen_dir,
image_color_mode="rgba",
target_size=target_size,
flag_multi_class=True,
num_class=3)
# # Create the names of the models to save them on the hard drive
def model_name(model, th0, th1, th2, th3, batch_size, epochs):
MODELDIR = '../../models/'
mname = 'neuneu_multicat_' + model + '_NL_th0-' + str(th0) \
+ '_th1-' + str(th1) \
+ '_th2-' + str(th2) \
+ '_th3-' + str(th3) + '_bs-' + str(batch_size) \
+ '_ep-' + str(epochs)
versions = []
for file in Path(MODELDIR).iterdir():
if file.name.startswith((mname)):
versions.append(int(file.name.rsplit(mname+'_r-')[1].split('.')[0]))
latest = 1
if len(versions) > 0:
latest = np.max(versions) + 1
model_name = MODELDIR + mname + '_r-' + str(latest) + '.hdf5'
pretrained = ''
if(latest > 1):
pretrained = MODELDIR + mname + '_r-' + str(latest-1) + '.hdf5'
return model_name, pretrained
def plot_history(history):
plt.plot(history["acc"], label="acc")
plt.plot(history["val_acc"], label="val_acc")
plt.legend()
plt.show()
plt.close()
plt.plot(history["loss"], label="loss")
plt.plot(history["val_loss"], label="val_loss")
plt.legend()
plt.show()
plt.close()#
plt.plot(history["auc_pr_multiclass"], label="auc_pr_multiclass")
plt.plot(history["val_auc_pr_multiclass"], label="val_auc_pr_multiclass")
plt.legend()
plt.show()
plt.close()
def prediction(model):
testGene = testGenerator("../../data/test/sat",target_size=(256, 256),as_gray=False)
n = 0
for img, name in testGene:
results = model.predict(img, batch_size=1)
saveResult("../../data/test/predict", results, name, True, 3)
n += 1
if(n>300):
break
# # Define the training schedule
def go(model = 'unet', th0 = 0.01, th1 = 0.08, th2=0.03, th3 = 0.04, target_size = (256, 256), batch_size = 3, epochs = 5, data_aug = True, pretrained = False):
valid_gen_args = dict(data_format="channels_last")
if (data_aug == True):
train_gen_args = dict(
data_format="channels_last",
horizontal_flip=True,
vertical_flip=True
)
else:
train_gen_args = valid_gen_args
trained_model_fn, pretrained_model_fn = model_name(model, th0, th1, th2, th3, batch_size, epochs)
steps_per_epoch = len(train) // batch_size
validation_steps = len(valid) // batch_size
optimizer = Adam(lr=1e-4)
loss_weights = None
metrics = ['accuracy', auc_pr_multiclass]
imgdatagen_dir = None
train_gen = data_generator(batch_size, train_gen_args, train_dir, imgdatagen_dir, target_size)
validation_gen = data_generator(batch_size, valid_gen_args, validation_dir, imgdatagen_dir,target_size)
width, height = target_size
if model == 'unet':
model = munet((width, height, 4), 3)
if model == 'segnet':
model = segnet((width, height, 4), 3)
model.compile(optimizer=optimizer,
loss=noisy_loss(th0, th1, th2, th3),
loss_weights=loss_weights,
metrics=metrics)
#model.summary()
if (pretrained == True):
if (len(pretrained_model_fn) > 0):
print('... loading the pretrained model', pretrained_model_fn)
model.load_weights(pretrained_model_fn)
else:
print('load the weights from the pretrained binary model...')
binary_model = load_model('../../models/unet_borneo_and_harz_05_09_16_22.hdf5', custom_objects={'auc_pr': auc_pr})
for layer, pretrained_layer in zip(model.layers[1:38], binary_model.layers[1:38]):
layer.set_weights(pretrained_layer.get_weights())
#else:
# print('load default model')
# # model.load_weights('../../models/0905_1500_NL_multiCat_unet.hdf5')#multicat_unet_NL_th0-0.01_th1-0.01_bs-3_ep-18_r-3.hdf5')#multicat_unet_NL_th0-0.08_th1-0.08_bs-3_ep-10_r-2.hdf5')
# model.load_weights('../../models/multicat_unet_NL_th0-0.01_th1-0.08_bs-3_ep-30_r-2.hdf5')#multicat_unet_NL_th0-0.08_th1-0.04_bs-3_ep-10_r-1.hdf5')#../../models/multicat_unet_NL_th0-0.01_th1-0.08_bs-3_ep-5_r-6.hdf5')
model_checkpoint = ModelCheckpoint(trained_model_fn, monitor='loss',verbose=1, save_best_only=True)
leaning_rate = ReduceLROnPlateau(monitor='val_loss', factor=0.5, patience=5, verbose=0, mode='auto', min_delta=0.0001, cooldown=0, min_lr=0)
early_stop = EarlyStopping(monitor='loss', min_delta=0, patience=10, verbose=1, mode='auto', baseline=None)
hist = History()
tesorboard = TrainValTensorBoard(log_dir='./logs', batch_size=batch_size,
write_graph=True,
write_grads=True,
write_images=True)
class_weights = create_class_weight(labeldict)
history = model.fit_generator(
train_gen,
steps_per_epoch=steps_per_epoch,
epochs=epochs,
class_weight=class_weights,
callbacks=[model_checkpoint, leaning_rate, early_stop, hist, tesorboard],
validation_data=validation_gen,
validation_steps=validation_steps
)
return history, model
# # Modify the TensorBoard output such that training and validation metrics and losses are shown in one graph
class TrainValTensorBoard(TensorBoard):
def __init__(self, log_dir='./logs', **kwargs):
# Make the original `TensorBoard` log to a subdirectory 'training'
training_log_dir = os.path.join(log_dir, 'training')
super(TrainValTensorBoard, self).__init__(training_log_dir, **kwargs)
# Log the validation metrics to a separate subdirectory
self.val_log_dir = os.path.join(log_dir, 'validation')
def set_model(self, model):
# Setup writer for validation metrics
self.val_writer = tf.summary.FileWriter(self.val_log_dir)
super(TrainValTensorBoard, self).set_model(model)
def on_epoch_end(self, epoch, logs=None):
# Pop the validation logs and handle them separately with
# `self.val_writer`. Also rename the keys so that they can
# be plotted on the same figure with the training metrics
logs = logs or {}
val_logs = {k.replace('val_', ''): v for k, v in logs.items() if k.startswith('val_')}
for name, value in val_logs.items():
summary = tf.Summary()
summary_value = summary.value.add()
summary_value.simple_value = value.item()
summary_value.tag = name
self.val_writer.add_summary(summary, epoch)
self.val_writer.flush()
# Pass the remaining logs to `TensorBoard.on_epoch_end`
logs = {k: v for k, v in logs.items() if not k.startswith('val_')}
super(TrainValTensorBoard, self).on_epoch_end(epoch, logs)
def on_train_end(self, logs=None):
super(TrainValTensorBoard, self).on_train_end(logs)
self.val_writer.close()
# # Train the model and create the prediction afterwards
glob_history = {}
for t0 in [0.0]:
for t1 in [0.0]:
for t2 in [0.0]:
for t3 in [0.0]:
for i in range(3):
history, model = go(batch_size = 3, epochs = 30, pretrained = True, th0=t0, th1=t1, th2=t2, th3=t3)
glob_history.update(history.history)
plot_history(glob_history)
prediction(model)
del history
del model
keras.clear_session()
# !rm -rf logs/*
# !rm -rf ../../models/neuneu*
|
notebooks/networks/run_multiCatNetwork_V2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
# %load_ext autoreload
# %reload_ext autoreload
# %autoreload 2
# %matplotlib inline
import os
# TO USE A DATABASE OTHER THAN SQLITE, USE THIS LINE
# Note that this is necessary for parallel execution amongst other things...
# os.environ['SNORKELDB'] = 'postgres:///snorkel-intro'
from snorkel import SnorkelSession
session = SnorkelSession()
# Here, we just set how many documents we'll process for automatic testing- you can safely ignore this!
n_docs = 500 if 'CI' in os.environ else 2591
from snorkel.models import candidate_subclass
Spouse = candidate_subclass('Spouse', ['person1', 'person2'])
train_cands = session.query(Spouse).filter(Spouse.split == 0).order_by(Spouse.id).all()
dev_cands = session.query(Spouse).filter(Spouse.split == 1).order_by(Spouse.id).all()
test_cands = session.query(Spouse).filter(Spouse.split == 2).order_by(Spouse.id).all()
# +
from util import load_external_labels
# %time load_external_labels(session, Spouse, annotator_name='gold')
from snorkel.annotations import load_gold_labels
#L_gold_dev = load_gold_labels(session, annotator_name='gold', split=1, zero_one=True)
#L_gold_test = load_gold_labels(session, annotator_name='gold', split=2, zero_one=True)
L_gold_dev = load_gold_labels(session, annotator_name='gold', split=1)
L_gold_test = load_gold_labels(session, annotator_name='gold', split=2)
# +
#gold_labels_dev = [x[0,0] for x in L_gold_dev.todense()]
#for i,L in enumerate(gold_labels_dev):
# print(i,gold_labels_dev[i])
gold_labels_dev = []
for i,L in enumerate(L_gold_dev):
gold_labels_dev.append(L[0,0])
gold_labels_test = []
for i,L in enumerate(L_gold_test):
gold_labels_test.append(L[0,0])
print(len(gold_labels_dev),len(gold_labels_test))
# -
import gensim
gensim.scripts.glove2word2vec.glove2word2vec('../glove.6B.300d.txt', '../glove_w2v.txt')
# +
from gensim.parsing.preprocessing import STOPWORDS
import gensim.matutils as gm
from gensim.models.keyedvectors import KeyedVectors
# Load pretrained model (since intermediate data is not included, the model cannot be refined with additional data)
model = KeyedVectors.load_word2vec_format('../glove_w2v.txt', binary=False) # C binary format
wordvec_unavailable= set()
def write_to_file(wordvec_unavailable):
with open("wordvec_unavailable.txt","w") as f:
for word in wordvec_unavailable:
f.write(word+"\n")
def preprocess(tokens):
btw_words = [word for word in tokens if word not in STOPWORDS]
btw_words = [word for word in btw_words if word.isalpha()]
return btw_words
def get_word_vectors(btw_words): # returns vector of embeddings of words
word_vectors= []
for word in btw_words:
try:
word_v = np.array(model[word])
word_v = word_v.reshape(len(word_v),1)
#print(word_v.shape)
word_vectors.append(model[word])
except:
wordvec_unavailable.add(word)
return word_vectors
def get_similarity(word_vectors,target_word): # sent(list of word vecs) to word similarity
similarity = 0
target_word_vector = 0
try:
target_word_vector = model[target_word]
except:
wordvec_unavailable.add(target_word+" t")
return similarity
target_word_sparse = gm.any2sparse(target_word_vector,eps=1e-09)
for wv in word_vectors:
wv_sparse = gm.any2sparse(wv, eps=1e-09)
similarity = max(similarity,gm.cossim(wv_sparse,target_word_sparse))
return similarity
# +
##### Continuous ################
import re
from snorkel.lf_helpers import (
get_left_tokens, get_right_tokens, get_between_tokens,
get_text_between, get_tagged_text,
)
spouses = {'spouse', 'wife', 'husband', 'ex-wife', 'ex-husband'}
family = {'father', 'mother', 'sister', 'brother', 'son', 'daughter',
'grandfather', 'grandmother', 'uncle', 'aunt', 'cousin'}
family = family | {f + '-in-law' for f in family}
other = {'boyfriend', 'girlfriend' 'boss', 'employee', 'secretary', 'co-worker'}
# Helper function to get last name
def last_name(s):
name_parts = s.split(' ')
return name_parts[-1] if len(name_parts) > 1 else None
def LF_husband_wife(c):
sc = 0
word_vectors = get_word_vectors(preprocess(get_between_tokens(c)))
for sw in spouses:
sc=max(sc,get_similarity(word_vectors,sw))
if sc<0.8:
return (0,0)
return (1,sc)
def LF_husband_wife_left_window(c):
sc_1 = 0
word_vectors = get_word_vectors(preprocess(get_left_tokens(c[0])))
for sw in spouses:
sc_1=max(sc_1,get_similarity(word_vectors,sw))
sc_2 = 0
word_vectors = get_word_vectors(preprocess(get_left_tokens(c[1])))
for sw in spouses:
sc_2=max(sc_2,get_similarity(word_vectors,sw))
return(1,max(sc_1,sc_2))
def LF_same_last_name(c):
p1_last_name = last_name(c.person1.get_span())
p2_last_name = last_name(c.person2.get_span())
if p1_last_name and p2_last_name and p1_last_name == p2_last_name:
if c.person1.get_span() != c.person2.get_span():
return (1,1)
return (0,0)
def LF_no_spouse_in_sentence(c):
return (-1,0.75) if np.random.rand() < 0.75 and len(spouses.intersection(c.get_parent().words)) == 0 else (0,0)
def LF_and_married(c):
word_vectors = get_word_vectors(preprocess(get_right_tokens(c)))
sc = get_similarity(word_vectors,'married')
if sc<0.7:
return (0,0)
if 'and' in get_between_tokens(c):
return (1,sc)
else:
return (0,0)
def LF_familial_relationship(c):
sc = 0
word_vectors = get_word_vectors(preprocess(get_between_tokens(c)))
for fw in family:
sc=max(sc,get_similarity(word_vectors,fw))
return (-1,sc)
def LF_family_left_window(c):
sc_1 = 0
word_vectors = get_word_vectors(preprocess(get_left_tokens(c[0])))
for fw in family:
sc_1=max(sc_1,get_similarity(word_vectors,fw))
sc_2 = 0
word_vectors = get_word_vectors(preprocess(get_left_tokens(c[1])))
for fw in family:
sc_2=max(sc_2,get_similarity(word_vectors,fw))
return (-1,max(sc_1,sc_2))
def LF_other_relationship(c):
sc = 0
word_vectors = get_word_vectors(preprocess(get_between_tokens(c)))
for ow in other:
sc=max(sc,get_similarity(word_vectors,ow))
return (-1,sc)
def LF_other_relationship_left_window(c):
sc = 0
word_vectors = get_word_vectors(preprocess(get_left_tokens(c)))
for ow in other:
sc=max(sc,get_similarity(word_vectors,ow))
return (-1,sc)
import bz2
# Function to remove special characters from text
def strip_special(s):
return ''.join(c for c in s if ord(c) < 128)
# Read in known spouse pairs and save as set of tuples
with bz2.BZ2File('data/spouses_dbpedia.csv.bz2', 'rb') as f:
known_spouses = set(
tuple(strip_special(x).strip().split(',')) for x in f.readlines()
)
# Last name pairs for known spouses
last_names = set([(last_name(x), last_name(y)) for x, y in known_spouses if last_name(x) and last_name(y)])
def LF_distant_supervision(c):
p1, p2 = c.person1.get_span(), c.person2.get_span()
return (1,1) if (p1, p2) in known_spouses or (p2, p1) in known_spouses else (0,0)
def LF_distant_supervision_last_names(c):
p1, p2 = c.person1.get_span(), c.person2.get_span()
p1n, p2n = last_name(p1), last_name(p2)
return (1,1) if (p1 != p2) and ((p1n, p2n) in last_names or (p2n, p1n) in last_names) else (0,1)
import numpy as np
def LF_Three_Lists_Left_Window(c):
c1,s1 = LF_husband_wife_left_window(c)
c2,s2 = LF_family_left_window(c)
c3,s3 = LF_other_relationship_left_window(c)
sc = np.array([s1,s2,s3])
c = [c1,c2,c3]
sharp_param = 1.5
prob_sc = np.exp(sc * sharp_param - np.max(sc))
prob_sc = prob_sc / np.sum(prob_sc)
#print 'Left:',s1,s2,s3,prob_sc
if s1==s2 or s3==s1 or np.max(sc)<0.5:
return (0,0)
return c[np.argmax(prob_sc)],1
def LF_Three_Lists_Between_Words(c):
c1,s1 = LF_husband_wife(c)
c2,s2 = LF_familial_relationship(c)
c3,s3 = LF_other_relationship(c)
sc = np.array([s1,s2,s3])
c = [c1,c2,c3]
sharp_param = 1.5
prob_sc = np.exp(sc * sharp_param - np.max(sc))
prob_sc = prob_sc / np.sum(prob_sc)
#print 'BW:',s1,s2,s3,prob_sc
#if s1==s2 or s3==s1 or np.max(sc)<0.5:
# return (0,0)
return c[np.argmax(prob_sc)],1
LFs = [LF_distant_supervision, LF_distant_supervision_last_names,LF_same_last_name,
LF_and_married, LF_Three_Lists_Between_Words,LF_Three_Lists_Left_Window
]
# +
import re
from snorkel.lf_helpers import (
get_left_tokens, get_right_tokens, get_between_tokens,
get_text_between, get_tagged_text,
)
spouses = {'spouse', 'wife', 'husband', 'ex-wife', 'ex-husband'}
family = {'father', 'mother', 'sister', 'brother', 'son', 'daughter',
'grandfather', 'grandmother', 'uncle', 'aunt', 'cousin'}
family = family | {f + '-in-law' for f in family}
other = {'boyfriend', 'girlfriend' 'boss', 'employee', 'secretary', 'co-worker'}
# Helper function to get last name
def last_name(s):
name_parts = s.split(' ')
return name_parts[-1] if len(name_parts) > 1 else None
def LF_husband_wife(c):
return (1,1) if len(spouses.intersection(get_between_tokens(c))) > 0 else (0,1)
def LF_husband_wife_left_window(c):
if len(spouses.intersection(get_left_tokens(c[0], window=2))) > 0:
return (1,1)
elif len(spouses.intersection(get_left_tokens(c[1], window=2))) > 0:
return (1,1)
else:
return (0,1)
def LF_same_last_name(c):
p1_last_name = last_name(c.person1.get_span())
p2_last_name = last_name(c.person2.get_span())
if p1_last_name and p2_last_name and p1_last_name == p2_last_name:
if c.person1.get_span() != c.person2.get_span():
return (1,1)
return (0,1)
def LF_no_spouse_in_sentence(c):
return (-1,1) if np.random.rand() < 0.75 and len(spouses.intersection(c.get_parent().words)) == 0 else (0,1)
def LF_and_married(c):
return (1,1) if 'and' in get_between_tokens(c) and 'married' in get_right_tokens(c) else (0,1)
def LF_familial_relationship(c):
return (-1,1) if len(family.intersection(get_between_tokens(c))) > 0 else (0,1)
def LF_family_left_window(c):
if len(family.intersection(get_left_tokens(c[0], window=2))) > 0:
return (-1,1)
elif len(family.intersection(get_left_tokens(c[1], window=2))) > 0:
return (-1,1)
else:
return (0,1)
def LF_other_relationship(c):
return (-1,1) if len(other.intersection(get_between_tokens(c))) > 0 else (0,1)
import bz2
# Function to remove special characters from text
def strip_special(s):
return ''.join(c for c in s if ord(c) < 128)
# Read in known spouse pairs and save as set of tuples
with bz2.BZ2File('data/spouses_dbpedia.csv.bz2', 'rb') as f:
known_spouses = set(
tuple(strip_special(x).strip().split(',')) for x in f.readlines()
)
# Last name pairs for known spouses
last_names = set([(last_name(x), last_name(y)) for x, y in known_spouses if last_name(x) and last_name(y)])
def LF_distant_supervision(c):
p1, p2 = c.person1.get_span(), c.person2.get_span()
return (1,1) if (p1, p2) in known_spouses or (p2, p1) in known_spouses else (0,1)
def LF_distant_supervision_last_names(c):
p1, p2 = c.person1.get_span(), c.person2.get_span()
p1n, p2n = last_name(p1), last_name(p2)
return (1,1) if (p1 != p2) and ((p1n, p2n) in last_names or (p2n, p1n) in last_names) else (0,1)
# -
def LF_Three_Lists_Between_Words(c):
c1,s1 = LF_husband_wife(c)
c2,s2 = LF_familial_relationship(c)
c3,s3 = LF_other_relationship(c)
sc = np.array([s1,s2,s3])
c = [c1,c2,c3]
sharp_param = 1.5
prob_sc = np.exp(sc * sharp_param - np.max(sc))
prob_sc = prob_sc / np.sum(prob_sc)
#print 'BW:',s1,s2,s3,prob_sc
#if s1==s2 or s3==s1 or np.max(sc)<0.5:
# return (0,0)
return c[np.argmax(prob_sc)],1
LFs = [
LF_distant_supervision, LF_distant_supervision_last_names,
LF_husband_wife, LF_husband_wife_left_window, LF_same_last_name,
LF_no_spouse_in_sentence, LF_and_married, LF_familial_relationship,
LF_family_left_window, LF_other_relationship, LF_Three_Lists_Between_Words
]
LFs = [
LF_distant_supervision, LF_distant_supervision_last_names,
LF_husband_wife_left_window, LF_same_last_name,
LF_no_spouse_in_sentence, LF_and_married,
LF_family_left_window, LF_Three_Lists_Between_Words
]
# +
import numpy as np
import math
def PHI(K,LAMDAi,SCOREi):
return [K*l*s for (l,s) in zip(LAMDAi,SCOREi)]
def softmax(THETA,LAMDAi,SCOREi):
x = []
for k in [1,-1]:
product = np.dot(PHI(k,LAMDAi,SCOREi),THETA)
x.append(product)
return np.exp(x) / np.sum(np.exp(x), axis=0)
def function_conf(THETA,LAMDA,P_cap,Confidence):
s = 0.0
i = 0
for LAMDAi in LAMDA:
s = s + Confidence[i]*np.dot(np.log(softmax(THETA,LAMDAi)),P_cap[i])
i = i+1
return -s
def function(THETA,LAMDA,SCORE,P_cap):
s = 0.0
i = 0
for i in range(len(LAMDA)):
s = s + np.dot(np.log(softmax(THETA,LAMDA[i],SCORE[i])),P_cap[i])
i = i+1
return -s
def P_K_Given_LAMDAi_THETA(K,THETA,LAMDAi,SCOREi):
x = softmax(THETA,LAMDAi,SCOREi)
if(K==1):
return x[0]
else:
return x[1]
np.random.seed(78)
THETA = np.random.rand(len(LFs),1)
def PHIj(j,K,LAMDAi,SCOREi):
return LAMDAi[j]*K*SCOREi[j]
def RIGHT(j,LAMDAi,SCOREi,THETA):
phi = []
for k in [1,-1]:
phi.append(PHIj(j,k,LAMDAi,SCOREi))
x = softmax(THETA,LAMDAi,SCOREi)
return np.dot(phi,x)
def function_conf_der(THETA,LAMDA,P_cap,Confidence):
der = []
for j in range(len(THETA)):
i = 0
s = 0.0
for LAMDAi in LAMDA:
p = 0
for K in [1,-1]:
s = s + Confidence[i]*(PHIj(j,K,LAMDAi)-RIGHT(j,LAMDAi,THETA))*P_cap[i][p]
p = p+1
i = i+1
der.append(-s)
return np.array(der)
def function_der(THETA,LAMDA,SCORE,P_cap):
der = []
for j in range(len(THETA)):
i = 0
s = 0.0
for index in range(len(LAMDA)):
p = 0
for K in [1,-1]:
s = s + (PHIj(j,K,LAMDA[index],SCORE[index])-RIGHT(j,LAMDA[index],SCORE[index],THETA))*P_cap[i][p]
p = p+1
i = i+1
der.append(-s)
return np.array(der)
import numpy as np
def get_LAMDA(cands):
LAMDA = []
SCORE = []
for ci in cands:
L=[]
S=[]
P_ik = []
for LF in LFs:
#print LF.__name__
l,s = LF(ci)
L.append(l)
S.append((s+1)/2) #to scale scores in [0,1]
LAMDA.append(L)
SCORE.append(S)
return LAMDA,SCORE
def get_Confidence(LAMDA):
confidence = []
for L in LAMDA:
Total_L = float(len(L))
No_zeros = L.count(0)
No_Non_Zeros = Total_L - No_zeros
confidence.append(No_Non_Zeros/Total_L)
return confidence
def get_Initial_P_cap(LAMDA):
P_cap = []
for L in LAMDA:
P_ik = []
denominator=float(L.count(1)+L.count(-1))
if(denominator==0):
denominator=1
P_ik.append(L.count(1)/denominator)
P_ik.append(L.count(-1)/denominator)
P_cap.append(P_ik)
return P_cap
#print(np.array(LAMDA))
#print(np.array(P_cap))append(L)
#LAMDA=np.array(LAMDA).astype(int)
#P_cap=np.array(P_cap)
#print(np.array(LAMDA).shape)
#print(np.array(P_cap).shape)
#print(L)
#print(ci.chemical.get_span(),ci.disease.get_span(),"No.Os",L.count(0),"No.1s",L.count(1),"No.-1s",L.count(-1))
#print(ci.chemical.get_span(),ci.disease.get_span(),"P(0):",L.count(0)/len(L)," P(1)",L.count(1)/len(L),"P(-1)",L.count(-1)/len(L))
def get_P_cap(LAMDA,SCORE,THETA):
P_cap = []
for i in range(len(LAMDA)):
P_capi = softmax(THETA,LAMDA[i],SCORE[i])
P_cap.append(P_capi)
return P_cap
def score(predicted_labels,gold_labels):
tp =0.0
tn =0.0
fp =0.0
fn =0.0
for i in range(len(gold_labels)):
if(predicted_labels[i]==gold_labels[i]):
if(predicted_labels[i]==1):
tp=tp+1
else:
tn=tn+1
else:
if(predicted_labels[i]==1):
fp=fp+1
else:
fn=fn+1
print("tp",tp,"tn",tn,"fp",fp,"fn",fn)
precision = tp/(tp+fp)
recall = tp/(tp+fn)
f1score = (2*precision*recall)/(precision+recall)
print("precision:",precision)
print("recall:",recall)
print("F1 score:",f1score)
from scipy.optimize import minimize
import cPickle as pickle
def get_marginals(P_cap):
marginals = []
for P_capi in P_cap:
marginals.append(P_capi[0])
return marginals
def predict_labels(marginals):
predicted_labels=[]
for i in marginals:
if(i<0.5):
predicted_labels.append(-1)
else:
predicted_labels.append(1)
return predicted_labels
def print_details(label,THETA,LAMDA,SCORE):
print(label)
P_cap = get_P_cap(LAMDA,SCORE,THETA)
marginals=get_marginals(P_cap)
plt.hist(marginals, bins=20)
plt.show()
plt.bar(range(0,2796),marginals)
plt.show()
predicted_labels=predict_labels(marginals)
print(len(marginals),len(predicted_labels),len(gold_labels_dev))
#score(predicted_labels,gold_labels_dev)
print(precision_recall_fscore_support(np.array(gold_labels_dev),np.array(predicted_labels),average='binary'))
def train(No_Iter,Use_Confidence=True,theta_file_name="THETA"):
global THETA
global dev_LAMDA,dev_SCORE
LAMDA,SCORE = get_LAMDA(train_cands)
P_cap = get_Initial_P_cap(LAMDA)
Confidence = get_Confidence(LAMDA)
for iteration in range(No_Iter):
if(Use_Confidence==True):
res = minimize(function_conf,THETA,args=(LAMDA,P_cap,Confidence), method='BFGS',jac=function_conf_der,options={'disp': True, 'maxiter':20}) #nelder-mead
else:
res = minimize(function,THETA,args=(LAMDA,SCORE,P_cap), method='BFGS',jac=function_der,options={'disp': True, 'maxiter':20}) #nelder-mead
THETA = res.x # new THETA
print(THETA)
P_cap = get_P_cap(LAMDA,SCORE,THETA) #new p_cap
print_details("train iteration: "+str(iteration),THETA,dev_LAMDA,dev_SCORE)
#score(predicted_labels,gold_labels)
NP_P_cap = np.array(P_cap)
np.savetxt('Train_P_cap.txt', NP_P_cap, fmt='%f')
pickle.dump(NP_P_cap,open("Train_P_cap.p","wb"))
NP_THETA = np.array(THETA)
np.savetxt(theta_file_name+'.txt', NP_THETA, fmt='%f')
pickle.dump( NP_THETA, open( theta_file_name+'.p', "wb" )) # save the file as "outfile_name.npy"
def test(THETA):
global dev_LAMDA,dev_SCORE
P_cap = get_P_cap(dev_LAMDA,dev_SCORE,THETA)
print_details("test:",THETA,dev_LAMDA,dev_SCORE)
NP_P_cap = np.array(P_cap)
np.savetxt('Dev_P_cap.txt', NP_P_cap, fmt='%f')
pickle.dump(NP_P_cap,open("Dev_P_cap.p","wb"))
def load_marginals(s):
marginals = []
if(s=="train"):
train_P_cap = np.load("Train_P_cap.npy")
marginals = train_P_cap[:,0]
return marginals
# +
from sklearn.metrics import precision_recall_fscore_support
import matplotlib.pyplot as plt
dev_LAMDA,dev_SCORE = get_LAMDA(dev_cands)
write_to_file(wordvec_unavailable)
# +
# with Three lists between nothing removed
train(3,Use_Confidence=False,theta_file_name="Three_lists_between_THETA")
test(THETA)
# +
# with Three lists between and removed husband_wife,familial_relationship,other_relationship
# of only between words
train(3,Use_Confidence=False,theta_file_name="Three_lists_between_THETA")
test(THETA)
# +
# except LF_husband_wife
train(3,Use_Confidence=False,theta_file_name="THETA")
test(THETA)
# +
#All are descrete
train(3,Use_Confidence=False,"THETA")
test(THETA)
# +
def print_details(label,THETA,LAMDA,SCORE):
print(label)
P_cap = get_P_cap(LAMDA,SCORE,THETA)
marginals=get_marginals(P_cap)
plt.hist(marginals, bins=20)
plt.show()
#plt.bar(range(0,2796),marginals)
#plt.show()
predicted_labels=predict_labels(marginals)
print(len(marginals),len(predicted_labels),len(gold_labels_dev))
#score(predicted_labels,gold_labels_dev)
print(precision_recall_fscore_support(np.array(gold_labels_dev),np.array(predicted_labels),average='binary'))
def predict_labels(marginals):
predicted_labels=[]
for i in marginals:
if(i<0.5):
predicted_labels.append(-1)
else:
predicted_labels.append(1)
return predicted_labels
#import cPickle as pickle
#THETA = pickle.load( open( "THETA.p", "rb" ) )
#test(THETA)
#LAMDA,SCORE = get_LAMDA(dev_cands)
#Confidence = get_Confidence(LAMDA)
#P_cap = get_P_cap(LAMDA,SCORE,THETA)
#marginals=get_marginals(P_cap)
#plt.hist(marginals, bins=20)
#plt.show()
#plt.bar(range(0,888),train_marginals)
#plt.show()
print_details("dev set",THETA,dev_LAMDA,dev_SCORE)
predicted_labels=predict_labels(marginals)
sorted_predicted_labels=[x for (y,x) in sorted(zip(Confidence,predicted_labels))] #sort Labels as per Confidence
sorted_predicted_labels=list(reversed(sorted_predicted_labels))
for i,j in enumerate(reversed(sorted(zip(Confidence,predicted_labels,gold_labels_dev)))):
if i>20:
break
print i,j
#print(len(marginals),len(predicted_labels),len(gold_labels_dev))
#no_of_labels=186#int(len(predicted_labels)*0.1) #54 - >0.2 , 108>= 0.15 , 186>= 0.12
#print(len(sorted_predicted_labels[0:no_of_labels]))
no_of_labels=2796
score(predicted_labels[0:no_of_labels],gold_labels_dev[0:no_of_labels])
|
intro-glove/LFS_ContinousSpace_one_by_one.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.5.1
# language: julia
# name: julia-1.5
# ---
# ---
# title: Facility Location
# ---
# **Originally Contributed by**: <NAME> and <NAME>
# +
using Random
using LinearAlgebra
using JuMP
import GLPK
using Plots
# -
# ## Uncapacitated facility location
# ### Problem description
#
# We are given
# * $M=\{1, \dots, m\}$ clients
# * $N=\{ 1, \dots, n\}$ sites where a facility can be built
#
# **Decision variables**
# Decision variables are split into two categories:
# * Binary variable $y_{j}$ indicates whether facility $j$ is built or not
# * Binary variable $x_{i, j}$ indicates whether client $i$ is assigned to facility $j$
#
# **Objective**
# The objective is to minimize the total cost of serving all clients.
# This costs breaks down into two components:
# * Fixed cost of building a facility.
# In this example, this cost is $f_{j} = 1, \ \forall j$.
# * Cost of serving clients from the assigned facility.
# In this example, the cost $c_{i, j}$
# of serving client $i$ from facility $j$
# is the Euclidean distance between the two.
#
# **Constraints**
# * Each customer must be served by exactly one facility
# * A facility cannot serve any client unless it is open
# ### MILP formulation
#
# The problem can be formulated as the following MILP:
#
# \begin{align}
# \min_{x, y} \ \ \ &
# \sum_{i, j} c_{i, j} x_{i, j} +
# \sum_{j} f_{j} y_{j}\\
# s.t. &
# \sum_{j} x_{i, j} = 1, && \forall i \in M\\
# & x_{i, j} \leq y_{j}, && \forall i \in M, j \in N\\
# & x_{i, j}, y_{j} \in \{0, 1\}, && \forall i \in M, j \in N
# \end{align}
#
# where the first set of constraints ensures
# that each client is served exactly once,
# and the second set of constraints ensures
# that no client is served from an unopened facility.
# ### Problem data
# +
Random.seed!(314)
m = 12 # number of clients
n = 5 # number of facility locations
# Clients' locations
Xc = rand(m)
Yc = rand(m)
# Facilities' potential locations
Xf = rand(n)
Yf = rand(n)
# Fixed costs
f = ones(n);
# Distance
c = zeros(m, n)
for i in 1:m
for j in 1:n
c[i, j] = norm([Xc[i] - Xf[j], Yc[i] - Yf[j]], 2)
end
end
# -
# Display the data
scatter(Xc, Yc, label = "Clients", markershape=:circle, markercolor=:blue)
scatter!(Xf, Yf, label="Facility",
markershape=:square, markercolor=:white, markersize=6,
markerstrokecolor=:red, markerstrokewidth=2
)
# ### JuMP implementation
# +
# Create a JuMP model
ufl = Model(GLPK.Optimizer)
# Variables
@variable(ufl, y[1:n], Bin);
@variable(ufl, x[1:m, 1:n], Bin);
# Each client is served exactly once
@constraint(ufl, client_service[i in 1:m],
sum(x[i, j] for j in 1:n) == 1
);
# A facility must be open to serve a client
@constraint(ufl, open_facility[i in 1:m, j in 1:n],
x[i, j] <= y[j]
)
# Objective
@objective(ufl, Min, f'y + sum(c .* x));
# -
# Solve the uncapacitated facility location problem with GLPK
optimize!(ufl)
println("Optimal value: ", objective_value(ufl))
# ### Visualizing the solution
# +
# The threshold 1e-5 ensure that edges between clients and facilities are drawn when x[i, j] ≈ 1.
x_ = value.(x) .> 1 - 1e-5
y_ = value.(y) .> 1 - 1e-5
# Display clients
p = scatter(Xc, Yc, markershape=:circle, markercolor=:blue, label=nothing)
# Show open facility
mc = [(y_[j] ? :red : :white) for j in 1:n]
scatter!(Xf, Yf,
markershape=:square, markercolor=mc, markersize=6,
markerstrokecolor=:red, markerstrokewidth=2,
label=nothing
)
# Show client-facility assignment
for i in 1:m
for j in 1:n
if x_[i, j] == 1
plot!([Xc[i], Xf[j]], [Yc[i], Yf[j]], color=:black, label=nothing)
end
end
end
display(p)
# -
# ## Capacitated Facility location
# ### Problem formulation
#
# The capacitated variant introduces a capacity constraint on each facility, i.e., clients have a certain level of demand to be served, while each facility only has finite capacity which cannot be exceeded.
#
# Specifically, let
# * $a_{i} \geq 0$ denote the demand of client $i$
# * $q_{j} \geq 0$ denote the capacity of facility $j$
#
# The capacity constraints then write
# \begin{align}
# \sum_{i} a_{i} x_{i, j} &\leq q_{j} y_{j} && \forall j \in N
# \end{align}
#
# Note that, if $y_{j}$ is set to $0$, the capacity constraint above automatically forces $x_{i, j}$ to $0$.
# Thus, the capacitated facility location can be formulated as follows
#
# \begin{align}
# \min_{x, y} \ \ \ &
# \sum_{i, j} c_{i, j} x_{i, j} +
# \sum_{j} f_{j} y_{j}\\
# s.t. &
# \sum_{j} x_{i, j} = 1, && \forall i \in M\\
# & \sum_{i} a_{i} x_{i, j} \leq q_{j} y_{j}, && \forall j \in N\\
# & x_{i, j}, y_{j} \in \{0, 1\}, && \forall i \in M, j \in N
# \end{align}
#
# For simplicity, we will assume that there is enough capacity to serve the demand,
# i.e., there exists at least one feasible solution.
# +
# Demands
a = rand(1:3, m);
# Capacities
q = rand(5:10, n);
# +
# Display the data
scatter(Xc, Yc, label=nothing,
markershape=:circle, markercolor=:blue, markersize= 2 .*(2 .+ a)
)
scatter!(Xf, Yf, label=nothing,
markershape=:rect, markercolor=:white, markersize= q,
markerstrokecolor=:red, markerstrokewidth=2
)
# -
# ### JuMP implementation
# +
# Create a JuMP model
cfl = Model(GLPK.Optimizer)
# Variables
@variable(cfl, y[1:n], Bin);
@variable(cfl, x[1:m, 1:n], Bin);
# Each client is served exactly once
@constraint(cfl, client_service[i in 1:m], sum(x[i, :]) == 1)
# Capacity constraint
@constraint(cfl, capacity, x'a .<= (q .* y))
# Objective
@objective(cfl, Min, f'y + sum(c .* x));
# -
# Solve the problem
optimize!(cfl)
println("Optimal value: ", objective_value(cfl))
# ### Visualizing the solution
# +
# The threshold 1e-5 ensure that edges between clients and facilities are drawn when x[i, j] ≈ 1.
x_ = value.(x) .> 1 - 1e-5
y_ = value.(y) .> 1 - 1e-5
# Display the solution
p = scatter(Xc, Yc, label=nothing,
markershape=:circle, markercolor=:blue, markersize= 2 .*(2 .+ a)
)
mc = [(y_[j] ? :red : :white) for j in 1:n]
scatter!(Xf, Yf, label=nothing,
markershape=:rect, markercolor=mc, markersize=q,
markerstrokecolor=:red, markerstrokewidth=2
)
# Show client-facility assignment
for i in 1:m
for j in 1:n
if x_[i, j] == 1
plot!([Xc[i], Xf[j]], [Yc[i], Yf[j]], color=:black, label=nothing)
break
end
end
end
display(p)
# -
# ## Further
# * [Benders decomposition](https://github.com/JuliaOpt/JuMPTutorials.jl/blob/master/script/optimization_concepts/benders_decomposition.jl)
# is a method of choice for solving facility location problems.
# * Benchmark instances can be found
# [here](https://resources.mpi-inf.mpg.de/departments/d1/projects/benchmarks/UflLib/).
|
notebook/modelling/facility_location.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# ## Deploy and perform inference on ML Model packages from AWS Marketplace.
#
# There are two simple ways to try/deploy [ML model packages from AWS Marketplace](https://aws.amazon.com/marketplace/search/results?page=1&filters=FulfillmentOptionType%2CSageMaker::ResourceType&FulfillmentOptionType=SageMaker&SageMaker::ResourceType=ModelPackage), either using AWS console to deploy an ML model package (see [this blog](https://aws.amazon.com/blogs/machine-learning/adding-ai-to-your-applications-with-ready-to-use-models-from-aws-marketplace/)) or via code written typically in a Jupyter notebook. Many listings have a high-quality sample Jupyter notebooks provided by the seller itself, usually, these sample notebooks are linked to the AWS Marketplace listing (E.g. [Source Separation](https://aws.amazon.com/marketplace/pp/prodview-23n4vi2zw67we?qid=1579739476471&sr=0-1&ref_=srh_res_product_title)), If a sample notebook exists, try it out.
#
# If such a sample notebook does not exist and you want to deploy and try an ML model package via code written in python language, this generic notebook can guide you on how to deploy and perform inference on an ML model package from AWS Marketplace.
#
#
# > **Note**:If you are facing technical issues while trying an ML model package from AWS Marketplace and need help, please open a support ticket or write to the team on <EMAIL> for additional assistance.
#
# #### Pre-requisites:
# 1. Open this notebook from an Amazon SageMaker Notebook instance.
# 1. Ensure that Amazon SageMaker notebook instance used has IAMExecutionRole with **AmazonSageMakerFullAccess**
# 1. Your IAM role has these three permisions - **aws-marketplace:ViewSubscriptions**, **aws-marketplace:Unsubscribe**, **aws-marketplace:Subscribe** and you have authority to make AWS Marketplace subscriptions in the AWS account used.
#
# > **Note**: If you are viewing this notebook from a GitHub repository, then to try this notebook successfully, [create an Amazon SageMaker Notebook Instance](https://docs.aws.amazon.com/sagemaker/latest/dg/howitworks-create-ws.html) and then [access Notebook Instance](https://docs.aws.amazon.com/sagemaker/latest/dg/howitworks-access-ws.html) you just created. Next, upload this Jupyter notebook to your notebook instance.
#
#
#
# #### Additional Resources:
# **Background on Model Packages**:
# 1. An ML model can be created from a Model Package, to know how, see [Use a Model Package to Create a Model](https://docs.aws.amazon.com/sagemaker/latest/dg/sagemaker-mkt-model-pkg-model.html).
# 2. An ML Model accepts data and generates predictions.
# 3. To perform inference, you first need to deploy the ML Model. An ML model typically supports two types of predictions:
# 1. [Use Batch Transform](https://docs.aws.amazon.com/sagemaker/latest/dg/batch-transform.html) to asynchronously generate predictions for multiple input data observations.
# 2. Send input data to Amazon SageMaker endpoint to synchronously generate predictions for individual data observations. For information, see [Deploy a Model on Amazon SageMaker Hosting Services](https://docs.aws.amazon.com/sagemaker/latest/dg/how-it-works-hosting.html)
#
# **Background on AWS Marketplace Model packages**:
# If you are new to Model packages from AWS Marketplace, here are some additional resources.
# * For a high level overview of how AWS Marketplace for Machine Learning see the [Using AWS Marketplace for machine learning workloads](https://aws.amazon.com/blogs/awsmarketplace/using-aws-marketplace-for-machine-learning-workloads/) blog post.
# * For a high level overview on Model packages from AWS Marketplace, see [this blog post](https://aws.amazon.com/blogs/aws/new-machine-learning-algorithms-and-model-packages-now-available-in-aws-marketplace/).
# * For an overview on how to deploy a Model package using AWS Console and using AWS CLI for performing inference, see the [Adding AI to your applications with ready-to-use models from AWS Marketplace](https://aws.amazon.com/blogs/machine-learning/adding-ai-to-your-applications-with-ready-to-use-models-from-aws-marketplace/) blog post.
# * For a Jupyter notebook of the sample solution for **Automating auto insurance claim processing workflow** outlined in [this re:Mars session](https://www.youtube.com/watch?v=GkKZt0s_ku0), see [amazon-sagemaker-examples/aws-marketplace](https://github.com/awslabs/amazon-sagemaker-examples/tree/master/aws_marketplace/using_model_packages/auto_insurance) GitHub repository.
# * For a Jupyter notebook of the sample solution for **Improving workplace safety solution** outlined in [this re:Invent session](https://www.youtube.com/watch?v=iLOXaWpK6ag), see [amazon-sagemaker-examples/aws-marketplace](https://github.com/awslabs/amazon-sagemaker-examples/tree/master/aws_marketplace/using_model_packages/improving_industrial_workplace_safety) GitHub repository.
#
# #### Contents:
# 1. [Subscribe to the model package](#Subscribe-to-the-model-package)
# 1. [Identify compatible instance-type](#A.-Identify-compatible-instance-type)
# 2. [Identify content-type](#B.-Identify-content_type)
# 3. [Specify model-package-arn](#C.-Specify-model-package-arn)
# 2. [Create an Endpoint and perform real-time inference](#2.-Create-an-Endpoint-and-perform-real-time-inference)
# 1. [Create an Endpoint](#A.-Create-an-Endpoint)
# 2. [Create input payload](#B.-Create-input-payload)
# 3. [Perform Real-time inference](#C.-Perform-Real-time-inference)
# 4. [Visualize output](#D.-Visualize-output)
# 5. [Delete the endpoint](#E.-Delete-the-endpoint)
# 3. [Perform Batch inference](#3.-Perform-Batch-inference)
# 1. [Prepare input payload](#A.-Prepare-input-payload)
# 2. [Run a batch-transform job](#B.-Run-a-batch-transform-job)
# 3. [Visualize output](#C.-Visualize-output)
# 4. [Delete the model](#4.-Delete-the-model)
# 5. [Unsubscribe to the model package](#Unsubscribe-to-the-model-package)
#
# #### Usage instructions
# You can run this notebook one cell at a time (By using Shift+Enter for running a cell).
# +
#Following boilerplate code includes all major libraries that you might need.
import base64
import json
import uuid
from sagemaker import ModelPackage
import sagemaker as sage
from sagemaker import get_execution_role
from sagemaker import ModelPackage
from urllib.parse import urlparse
import boto3
from IPython.display import Image
from PIL import Image as ImageEdit
import urllib.request
import numpy as np
role = get_execution_role()
sagemaker_session = sage.Session()
# -
bucket=sagemaker_session.default_bucket()
bucket
# ### 1. Subscribe to the model package
# Before you can deploy the model, your account needs to be subscribed to it. This section covers instructions for populating necessary parameters and for subscribing to the Model package, if the subscription does not already exist.
# 1. Open the Model Package listing page (E.g. [GluonCV YOLOv3 Object Detector](https://aws.amazon.com/marketplace/pp/prodview-5jlvp43tsn3ny?qid=1578429923058&ref_=srh_res_product_title&sr=0-1)) from AWS Marketplace that you wish to try/use.
# 2. Read the **product overview** section and **Highlights** section of the listing to understand the value proposition of the model package.
# 3. View **usage information** and then **additional resources** sections. These sections will contain following things:
# 1. Input content-type
# 2. Sample input file (optional)
# 3. Sample Jupyter notebook
# 4. Output format
# 5. Any additional information.
# #### A. Identify compatible instance-type
#
# 1. On the listing, Under **Pricing Information**, you will see **software pricing** for **real-time inference** as well as **batch-transform usage** for specific instance-types.
#
# > **Note**: Software pricing is in addition to regular SageMaker infrastructure charges.
#
# 2. In the pricing chart, you will also notice the **vendor recommended instance-type** . E.g [GluonCV YOLOv3 Object Detector](https://aws.amazon.com/marketplace/pp/prodview-5jlvp43tsn3ny?qid=1578429923058&ref_=srh_res_product_title&sr=0-1) has recommended real-time inference instance-type as
# **ml.m4.xlarge** and recommended batch transform inference as **ml.m4.xlarge**
#
# 3. Specify the recommended instance-types in the following cell and then run the cell.
real_time_inference_instance_type=''
batch_transform_inference_instance_type=''
#real_time_inference_instance_type='ml.m4.xlarge'
#batch_transform_inference_instance_type='ml.m4.xlarge'
# #### B. Identify content_type
# You need to specify input content-type and payload while performing inference on the model. In this sub-section you will identify input content type that is accepted by the model you wish to try.
# Sellers has provided content_type information via:
# 1. a sample invoke_endpoint api/CLI call in the **usage instructions** section, of the listing. E.g [GluonCV YOLOv3 Object Detector](https://aws.amazon.com/marketplace/pp/prodview-5jlvp43tsn3ny?qid=1578429923058&sr=0-1&ref_=srh_res_product_title) has following AWS CLI snippet, with --content-type specified as **image/jpeg**.
# >```Bash
# aws sagemaker-runtime invoke-endpoint --endpoint-name your_endpoint_name --body fileb://img.jpg --content-type image/jpeg --custom-attributes '{"threshold": 0.2}' --accept json >(cat) 1>/dev/null
# ```
#
# 2. plain-text information in the **usage instructions** section, of the listing. E.g. [Lyrics Generator (CPU)](https://aws.amazon.com/marketplace/pp/prodview-qqzh5iao6si4c?qid=1578429518061&sr=0-2&ref_=srh_res_product_title) has following snippet which indicates that **application/json** is the content-type.
#
# >```Javascript
# Input (application/json): Artist name and seed lyrics (start of song).
# Payload: {"instances": [{"artist":"<singer>", "seed": "<seed_word>"}]}
# ```
#
# 3. Sample notebook, linked under **usage instructions**/**additional information**/**support information** and the sample notebook might use AWS CLI/Boto3 or SDK to perform inference.
#
# 1. E.g., [Vehicle Damage Inspection](https://aws.amazon.com/marketplace/pp/prodview-xhj66rbazm6oe?qid=1579723100840&sr=0-1&ref_=srh_res_product_title) has a link to a file under **Additional resources** section that containing **Vehicle-Damage-Inspection.ipynb**, a jupyter notebook that has following snippet with ContentType specified as **image/jpeg**.
# > ```Python
# invocation_api_handle.invoke_endpoint(EndpointName=ENDPOINT_NAME, ContentType='image/jpeg', ...
# ```
# 2. [A Sample notebook from sagemaker-examples repo](https://github.com/awslabs/amazon-sagemaker-examples/blob/master/aws_marketplace/using_model_packages/auto_insurance/automating_auto_insurance_claim_processing.ipynb) uses Python SDK to perform inference, and the predictor class defined shows that content type is **image/jpeg**
# > ```Python
# def damage_detection_predict_wrapper(endpoint, session):
# return sage.RealTimePredictor(endpoint, session,content_type='image/jpeg')
# ```
#
# Once you have identified the input content type, specify the same in following cell.
content_type=''
#content_type='image/jpeg'
# #### C. Specify model-package-arn
# A model-package-arn is a unique identifier for each ML model package from AWS Marketplace within a chosen region.
# 1. On the AWS Marketplace listing, click on **Continue to subscribe** button.
# 2. On the **Subscribe to this software** page (E.g. [here](https://aws.amazon.com/marketplace/ai/procurement?productId=d9949c88-fe3b-4a2d-923e-9458fe7e9f2c)), Review the **End user license agreement**, **support terms**, as well as **pricing information**.
# 3. **"Accept Offer"** button needs to be clicked if your organization agrees with EULA, pricing information as well as support terms.
# 4. Next, **Continue to configuration** button becomes activated and when you click on the button, you will see that a **Product Arn** will appear. In the **Region** dropdown, Choose the region in which you have opened this notebook from, Copy the product ARN and replace it in the next cell.
model_package_arn=''
#model_package_arn='arn:aws:sagemaker:us-east-1:865070037744:model-package/gluoncv-yolo3-darknet531547760-bdf604d6d9c12bf6194b6ae534a638b2'
# Congratulations, you have identified necessary information to be able to create an endpoint for performing real-time inference.
# ### 2. Create an Endpoint and perform real-time inference.
# In this section, you will stand up an Amazon SageMaker endpoint. Each endpoint must have a unique name which you can use for performing inference.
#
# Specify a short name you wish to use for naming endpoint.
model_name=''
#model_name='gluoncv-object-detector'
# #### A. Create an Endpoint
# +
def predict_wrapper(endpoint, session):
return sage.RealTimePredictor(endpoint, session,content_type)
#create a deployable model from the model package.
model = ModelPackage(role=role,
model_package_arn=model_package_arn,
sagemaker_session=sagemaker_session,
predictor_cls=predict_wrapper)
#Deploy the model
predictor = model.deploy(1, real_time_inference_instance_type, endpoint_name=model_name)
# -
# Once endpoint has been created, you would be able to perform real-time inference.
# #### B. Create input payload
# **Background**: A Machine Learning model accepts a payload and returns an inference. E.g. [Deep Vision vehicle recognition](https://aws.amazon.com/marketplace/pp/prodview-a7wgrolhu54ts?qid=1579728052169&sr=0-1&ref_=srh_res_product_title) accepts an image as a payload and returns an inference containing make,model, and year of the car.
# In this step, you will prepare a payload to perform a prediction. This step varies from model to model.
# ##### Identify a sample Input file you can use:
# 1. Sometimes file is available in **additional resources** section of the listing. E.g. [Mphasis DeepInsights Document Classifier](https://aws.amazon.com/marketplace/pp/prodview-u5jlb2ba6xmaa?qid=1579793398686&sr=0-1&ref_=srh_res_product_title) has multiple sample files in an archieve.
#
#
# 2. Sometimes file is available in a Github Repo link associated with the listing. E.g. [Source Separation](https://aws.amazon.com/marketplace/pp/prodview-23n4vi2zw67we?qid=1579739476471&sr=0-1&ref_=srh_res_product_title) has a sample file in the GitHUB repo. In which case, please copy the link to the raw data file.
#
#
# 3. Sometimes a sample file is not available, however, clear instructions on how to prepare the payload are available. E.g. [Face Anonymizer (CPU)](https://aws.amazon.com/marketplace/pp/prodview-3olpixsfcqfq6?qid=1560287886810&sr=0-3&ref_=srh_res_product_title)), then you would need to manually identify an input file you can use. I identified that I can use an image shown on [this blog](https://aws-preview.aka.amazon.com/blogs/machine-learning/adding-ai-to-your-applications-with-ready-to-use-models-from-aws-marketplace/), and then manually prepare a payload for performing inference
#
# 4. For models for which there is no sample file (E.g. [Demisto Phishing Email Classifier](https://aws.amazon.com/marketplace/pp/prodview-k5354ho27eyps)) but it accepts a simple input, jump to [Step B.2](#Step-B.2-Manually-prepare-data-(applicable-only-if-your-payload-is-not-ready-yet))
# Specify the URL of the sample file you identified in the following cell to download the file for creating payload.
url=''
#url='https://upload.wikimedia.org/wikipedia/commons/thumb/b/b2/Vincent_van_Gogh_-_Self-Portrait_-_Google_Art_Project.jpg/512px-Vincent_van_Gogh_-_Self-Portrait_-_Google_Art_Project.jpg'
#url='https://d1.awsstatic.com/webteam/architecture-icons/AWS-Architecture-Icons-Deck_For-Light-BG_20191031.pptx.6fcecd0cf65442a1ada0ce1674bc8bfc8de0cb1d.zip'
# Next, specify a file_name that you would like to save the file to.
file_name=''
#file_name='input.json'
#file_name='input.jpg'
#file_name='file.zip'
#Download the file
urllib.request.urlretrieve(url,file_name)
# #### View the file you just downloaded
#
# Based on the type of file used, uncomment, modify, and run appropriate code snippets.
# ###### ZIP/Tar file
# If your input file was inside a zip file, uncomment appropriate line from following two lines.
# +
# #!unzip $file_name
# #!tar -xvf $file_name
# +
#Update the file_name variable with an appropriate file-path from the folder created by unzipping the archieve
#file_name=''
#file_name='images/AWS-Architecture-Icons-Deck_For-Light-BG_20191031.pptx'
# -
# ###### Image File
# +
#Uncomment and run the following line to view the image
#Image(url= file_name, width=400, height=800)
# -
# ###### Text/Json/CSV File
# If your input file is a text/json/csv file, view the file by un-commenting following line. If your file contains multiple payloads, consider keeping just one.
# +
# #!head $file_name
# -
# ##### Video File
# +
#View and play the video by uncommenting following two lines
#from IPython.display import HTML
#HTML('<iframe width="560" height="315" src="'+file_name+'?rel=0&controls=0&showinfo=0" frameborder="0" allowfullscreen></iframe>')
# -
# ##### Audio File
# +
#Uncomment following two lines to view and play the audio
#import IPython.display as ipd
#ipd.Audio(file_name)
# -
# If your model's input **content-type** is one of the following and file_name variable is pointing to a file that can directly be sent to ML model, congratulations, you have prepared the payload, you can jump to Step [C. Perform Real-time inference](#C.-Perform-Real-time-inference):
# * **wav/mp3**
# * **application/pdf**
# * **image/png**
# * **image/jpeg**:
# * **text/plain**
# * **text/csv**
# * **application/json** (Only if file_name variable is pointing to a JSON file that can directly be sent to ML model)
#
# If your content-type is any other, your model might need additional pre-processing, proceed to [Step B.1](#Step-B.1-Pre-process-the-data-(Optional-for-some-models))
# #### Step B.1 Pre-process the data (Optional for some models)
# Some models require preprocessing, others dont. If model you want to try/use requires additional pre-processing, Please refer to sample notebook or usage instructions for pre-processing required. This section contains some re-usable code that you might need to tweak as per your requirement. Uncomment, tweak and use following as required. Ensure that final payload is written to a variable with name 'payload'.
# ##### Some models require Base64 encoded data
# > [Background Noise Classifier (CPU)](https://aws.amazon.com/marketplace/pp/prodview-vpd6qdjm4d7u4?qid=1579792115621&sr=0-2&ref_=srh_res_product_title) requires payload to be in following format
#
# ```javascript
# Payload: {"instances": [{"audio": {"b64": "BASE_64_ENCODED_WAV_FILE_CONTENTS"}}]}
# ```
#
# > [Neural Style Transfer](https://aws.amazon.com/marketplace/pp/prodview-g5i35lg4qmplu) requires payload to be in following format. You would need to tweak code appropriately to convert two images into base64 format for this model.
#
# ```javascript
# {
# "content": "base64 characters of your content image",
# "style": "base64 characters of your style image",
# "iterations": 2
# }
# ```
#
# +
#Here is a sample code that does Base64 encoding
#file_read = open(file_name, 'rb').read()
#base64_encoded_value = base64.b64encode(file_read).decode('utf-8')
#payload="{\"style\":\""+str(style_image_base64_encoded_value)+"\", \"iterations\": 2,\"content\":\""+str(base64_encoded_value)+"\"}"
# -
# ##### Some models require images in serialized format
# > E.g. [Mphasis DeepInsights Damage Prediction](https://aws.amazon.com/marketplace/pp/prodview-2f5br37zmuk2y?qid=1576781776298&sr=0-1&ref_=srh_res_product_title) requires the image to be re-sized to (300 x 300) and then JSON serialised before it can be fed to the model. To make it easy to do so, they also have provided snippet identical to following one in the sample jupyter notebook.
# +
#from PIL import Image
#image = Image.open(file_name).convert(mode = 'RGB')
#resized_image = image.resize((300,300))
#image_array = np.array(resized_image).tolist()
#payload = json.dumps({'instances': [{'input_image': image_array}]})
# -
# Next, jump to [Step B.3](#Step-B.3-Write-payload-you-created-to-a-file)
# ##### Step B.2 Manually prepare data (applicable only if your payload is not ready yet)
# If sample notebook is not available but input format is simple, write code required for creating the input file. E.g. [Demisto Phishing Email Classifier](https://aws.amazon.com/marketplace/pp/prodview-k5354ho27eyps) does not have a sample file but sample notebook has some code that can be used for prepared payload.
#
# ```Javascript
# email1 = "<Email content shortened for brevity>"
# email2 = "<Email content shortened for brevity>"
# emails = [email1, email2]
# json.dumps(emails)
# ```
#
# Prepare appropriate payload and store the same in a variable called 'payload'.
# +
#Write your code here.
# -
# Jump to [Step B.3](#Step-B.3-Write-payload-you-created-to-a-file) to write your payload to a file.
# ##### Step B.3 Write payload you created to a file
# Assuming that you have populated payload json/csv in a variable called 'payload', here is a sample generic code that writes the payload to a file you can un-comment and reuse.
# +
#file_name='output_file'
#file = open(file_name, "w") #Change w to wb if you intend to write bytes isntead of text.
#file.write(payload)
#file.close()
# -
# Once your payload is ready and is written to a file referenced by the file_name variable, you are ready to perform an inference. Proceed to Step [C. Perform Real-time inference](#C.-Perform-Real-time-inference).
#
# #### C. Perform Real-time inference
# Specify name and extension of the file you would like to store the inference output to. The output type varies from model to model and this information is usually available in the **Usage instructions**/**sample notebook** associated with the listing.
#
# For Example:
# * [Neural Style Transfer](https://aws.amazon.com/marketplace/pp/prodview-g5i35lg4qmplu) model's output type is image, so specify **.png** as file extension.
# * [Source Separation](https://aws.amazon.com/marketplace/pp/prodview-23n4vi2zw67we?qid=1579807024600&sr=0-1&ref_=srh_res_product_title) model's output type is a zip file, so specify **.zip** as file extension.
# * [Mphasis DeepInsights Address Extraction](https://aws.amazon.com/marketplace/pp/prodview-z4wgslad4b27g?qid=1579802907920&sr=0-2&ref_=srh_res_product_title) model's output type is text/plain, so specify **.txt** as file extension.
# * Sample notebook provided by seller usually makes it evident what the output type is. If one doesnt exist, and instructions are unclear, try a few options, start with text - Many ML models return response in a simple textual format.
real_time_inference_output_file_name=''
#real_time_inference_output_file_name='output.json'
#real_time_inference_output_file_name='output.zip'
#real_time_inference_output_file_name='output.txt'
#real_time_inference_output_file_name='output.png'
# The following AWS CLI command sends the **payload** and the **content-type** to the model hosted on the endpoint.
# > **Note on Custom Attributes**: Some models accept additional attributes such as [GluonCV YOLOv3 Object Detector](https://aws.amazon.com/marketplace/pp/prodview-5jlvp43tsn3ny?qid=1578429923058&ref_=srh_res_product_title&sr=0-1)
# accepts a custom attribute called threshold as specified in following sample code snippet.
# > ```Bash
# aws sagemaker-runtime invoke-endpoint --endpoint-name your_endpoint_name --body fileb://img.jpg --content-type image/jpeg --custom-attributes '{"threshold": 0.2}' --accept json >(cat) 1>/dev/null
# ```
# Please modify the following AWS-CLI command appropriately if the model you wish to perform inference on requires any custom attribute, if not, execute following command to perform inference.
#
# Once inference has been performed, the output gets written to the output file.
# !aws sagemaker-runtime invoke-endpoint \
# --endpoint-name $model_name \
# --body fileb://$file_name \
# --content-type $content_type \
# --region $sagemaker_session.boto_region_name \
# $real_time_inference_output_file_name
# If the above invocation shows a snippet such as following, it means the command executed successfully. Otherwise, check whether input payload is in correct format.
#
# ```Javascript
# {
# "ContentType": "<content_type>; charset=utf-8",
# "InvokedProductionVariant": "<Variant>"
# }
# ```
# View the output available in file referenced by **real_time_inference_output_file_name** variable.
# #### D. Visualize output
# If the output is in **text**/**CSV**/**JSON** format, view the output file by uncommenting and running following command. Otherwise use an appropriate command (Please see reference commands from step [View the file you just downloaded](#View-the-file-you-just-downloaded)) for viewing the output OR open the output file directly from Jupyter console.
# +
#f=open(real_time_inference_output_file_name, "r")
#data=f.read()
#print(data)
#Sometimes output is a json, load it into a variable with json.loads(data) and then print the variable to see formatted output.
# -
# #### E. Delete the endpoint
# Now that you have successfully performed a real-time inference, you do not need the endpoint any more. you can terminate the same to avoid being charged.
predictor=sage.RealTimePredictor(model_name, sagemaker_session,content_type)
predictor.delete_endpoint(delete_endpoint_config=True)
# ### 3. Perform Batch inference
# To run a batch transform job, we will use the same payload we used for performing real-time inference. file_name variable points to the payload.
#
# > **Note**: If you followed instructions closely, your input file contains a single payload. However, batch-transform can be used to perform a batch inference on multiple records at a time. To know more, see documentation.
#upload the file to S3
transform_input = sagemaker_session.upload_data(file_name, key_prefix=model_name)
print("Transform input uploaded to " + transform_input)
#Run a batch-transform job
transformer = model.transformer(1, batch_transform_inference_instance_type)
transformer.transform(transform_input, content_type=content_type)
transformer.wait()
#output is available on following path
transformer.output_path
# #### C. Visualize output
# +
from urllib.parse import urlparse
parsed_url = urlparse(transformer.output_path)
bucket_name = parsed_url.netloc
file_key = '{}/{}.out'.format(parsed_url.path[1:], file_name.split("/")[-1])
print(file_key)
s3_client = sagemaker_session.boto_session.client('s3')
response = s3_client.get_object(Bucket = sagemaker_session.default_bucket(), Key = file_key)
# -
# If the output is in **text**/**CSV**/**JSON** format, view the output file by uncommenting and running following command. Otherwise go to S3, download the file and open it using appropriate editor.
response_bytes = response['Body'].read().decode('utf-8')
print(response_bytes)
# ### 4. Delete the model
model.delete_model()
# **Note** - You need to write appropriate code here to clean-up any files you may have uploaded/created while trying out this notebook.
# ### 5. Cleanup
# Finally, if the AWS Marketplace subscription was created just for the experiment and you would like to unsubscribe to the product, here are the steps that can be followed.
# Before you cancel the subscription, ensure that you do not have any [deployable model](https://console.aws.amazon.com/sagemaker/home#/models) created from the model package or using the algorithm. Note - You can find this information by looking at the container name associated with the model.
#
# **Steps to un-subscribe to product from AWS Marketplace**:
# 1. Navigate to __Machine Learning__ tab on [__Your Software subscriptions page__](https://aws.amazon.com/marketplace/ai/library?productType=ml&ref_=lbr_tab_ml)
# 2. Locate the listing that you would need to cancel subscription for, and then __Cancel Subscription__ can be clicked to cancel the subscription.
#
#
|
aws_marketplace/using_model_packages/generic_sample_notebook/A_generic_sample_notebook_to_perform_inference_on_ML_model_packages_from_AWS_Marketplace.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# These are the libraries that we use.
# + tags=["outputPrepend"]
import os
import re
import operator
import warnings
import joblib
import pandas as pd
import matplotlib as mpl
import matplotlib.pyplot as plt
from matplotlib.legend_handler import HandlerLine2D
import numpy as np
import math
import nltk
nltk.download('all')
from nltk.stem import PorterStemmer
from nltk.stem import LancasterStemmer
from nltk.stem import WordNetLemmatizer
from nltk.stem import PorterStemmer
from nltk.stem import LancasterStemmer
from nltk.corpus import stopwords
from nltk.tokenize import sent_tokenize, word_tokenize
from sklearn import tree
from sklearn.tree import DecisionTreeClassifier
from sklearn.model_selection import train_test_split
from sklearn.feature_extraction.text import TfidfVectorizer, CountVectorizer
from sklearn.metrics import confusion_matrix, accuracy_score, recall_score, f1_score, roc_auc_score, roc_curve, auc
from sklearn.metrics import classification_report
from sklearn.model_selection import cross_val_score, cross_validate
from sklearn.pipeline import Pipeline
from sklearn.utils.extmath import cartesian
from IPython.display import SVG, clear_output
from graphviz import Source
from IPython.display import display
# -
# Here we create the dataset we're going to use to train our Binary Decision Tree model. We make sure that both accepted and rejected tites are not duplicated. We get all the accepted titles (which are way fewer than the rejected ones), and then we get a sample of the rejected ones big enough as to make the union between accepted and rejected contain 1000 samples.
# + tags=[]
df = pd.read_csv('scopus_search_results.csv', encoding='UTF-8')
acceptedDataSet = df[['Title','Accepted']][df['Accepted'] == True]
acceptedDataSet = acceptedDataSet.drop_duplicates()
# since we have many rejected titles, get a random sample among them
# so that the union between accepted and rejected is 1000 titles
rejectedDataSet = df[['Title','Accepted']][df['Accepted'] == False].drop_duplicates().sample(n=1000-len(acceptedDataSet), random_state=1)
joinDataset = pd.concat([acceptedDataSet,rejectedDataSet],ignore_index=True)
print(len(acceptedDataSet))
print(len(rejectedDataSet))
print(len(joinDataset))
# print(joinDataset)
# -
# Titles preprocessing:
# - remove special and single characters
# - trim whitespaces
# - convert to lowercase
# - lemmatize the words: return the root of the words E.G. studying -> study
# + tags=[]
titles = joinDataset.Title
classifications = joinDataset.Accepted
lemmatizer = WordNetLemmatizer()
def titlePreprocessing(titles):
preProcessedTitles = []
for title in titles:
# Remove all the special characters
title = re.sub(r'\W', ' ', title)
# remove all single characters
title = re.sub(r'\s+[a-zA-Z]\s+', ' ', title)
# Remove single characters from the start
title = re.sub(r'\^[a-zA-Z]\s+', ' ', title)
# Substituting multiple spaces with single space
title = re.sub(r'\s+', ' ', title, flags=re.I)
# Removing prefixed 'b'
title = re.sub(r'^b\s+', '', title)
# Converting to Lowercase
title = title.lower()
# Lemmatization
title = title.split()
title = [lemmatizer.lemmatize(word) for word in title]
title = ' '.join(title)
preProcessedTitles.append(title)
return preProcessedTitles
preProcessedTitles = titlePreprocessing(titles)
for i in range(0,10):
print(preProcessedTitles[i])
# -
# Split the dataset into train and test and convert the classes into binary values because that's a requirement to fit a Binary Decision Tree.
# + tags=[]
train_titles, test_titles, train_classes, test_classes = train_test_split(preProcessedTitles, classifications, test_size=0.2, random_state=1)
# Classes must be binary to train a Binary Decision Tree so False -> 0 and True -> 1
train_classes = train_classes.astype(int)
test_classes = test_classes.astype(int)
print(train_classes.value_counts())
print(test_classes.value_counts())
# -
# Train the Binary Decision Tree on the train set.
# +
vectorizer = CountVectorizer(ngram_range=(1, 2), token_pattern = r'\b[\w]{4,}(?<![\d])\b', stop_words=stopwords.words('english'), min_df = 2)
# Create Decision Tree classifer object
max_depth = 11
min_samples_split = 0.015789474
min_samples_leaf = 0.00526315789473684
criterion="gini"
class_weight = "balanced"
splitter = "best"
dtc = DecisionTreeClassifier(max_depth = max_depth, min_samples_split = min_samples_split, min_samples_leaf = min_samples_leaf, criterion=criterion, class_weight = class_weight, splitter = splitter)
# dtc = DecisionTreeClassifier(criterion="entropy", splitter = "best", max_depth = 5, min_samples_leaf = 5, class_weight = "balanced")
dtc_pipe = Pipeline([('vectorizer', vectorizer), ('dtc', dtc)])
# Train Decision Tree Classifer
dtc_pipe.fit(train_titles,train_classes)
# -
# Test the accuracy of the prediction on the test set.
# + tags=[]
#Predict the response for test dataset
predicted_classes = dtc_pipe.predict(test_titles)
print(classification_report(test_classes, predicted_classes))
print(pd.crosstab(test_classes, predicted_classes, rownames=['True'], colnames=['Predicted'], margins=True))
print("Accuracy:",accuracy_score(test_classes, predicted_classes))
test_recall_score = recall_score(test_classes, predicted_classes, average='binary',pos_label = 1)
print("test_recall_score: %s"%test_recall_score)
# -
# Test the accuracy of the cross validation.
# + tags=[]
classifications = classifications.astype(int)
scores = cross_validate(dtc_pipe, titles, classifications, cv=10, scoring = ('recall', 'precision', 'roc_auc'))
print("recall cross validation: %0.2f (+/- %0.2f)" % (scores['test_recall'].mean(), scores['test_recall'].std() * 2))
print("test_precision cross validation: %0.2f (+/- %0.2f)" % (scores['test_precision'].mean(), scores['test_precision'].std() * 2))
print("test_roc_auc cross validation: %0.2f (+/- %0.2f)" % (scores['test_roc_auc'].mean(), scores['test_roc_auc'].std() * 2))
# -
# This is to visualize the resulting Binary Decision Tree Model. It requires "graphviz" (pip install graphviz)
# +
os.environ["PATH"] += os.pathsep + 'C:\\Program Files (x86)\\Graphviz2.38\\bin\\'
graph = Source(
tree.export_graphviz(
dtc,
out_file=None,
feature_names=vectorizer.get_feature_names(),
class_names=['1', '0'],
filled = True)
)
savePath = ".\models\\renders\\"
graph.format = 'pdf'
graph.render(str('%sdtc_maxdepth_%s_minsampleleaf_%s'%(savePath, max_depth,min_samples_leaf)),view=True)
# -
# Parameter Fitting: we try to get the best possible parameters for our Binary Decision Tree Classifier by bruteforcing and plotting the results.
# +
max_depths = np.linspace(1, 32, 32, endpoint=True)
train_results = []
test_results = []
for max_depth in max_depths:
dtc = DecisionTreeClassifier(max_depth=max_depth)
dtc_pipe = Pipeline([('vectorizer', vectorizer), ('dtc', dtc)])
dtc_pipe.fit(train_titles,train_classes)
train_pred = dtc_pipe.predict(train_titles)
false_positive_rate, true_positive_rate, thresholds = roc_curve(train_classes, train_pred)
roc_auc = auc(false_positive_rate, true_positive_rate)
# Add auc score to previous train results
train_results.append(roc_auc)
y_pred = dtc_pipe.predict(test_titles)
false_positive_rate, true_positive_rate, thresholds = roc_curve(test_classes, y_pred)
roc_auc = auc(false_positive_rate, true_positive_rate)
# Add auc score to previous test results
test_results.append(roc_auc)
line1, = plt.plot(max_depths, train_results, 'b', label="Train AUC")
line2, = plt.plot(max_depths, test_results, 'r', label="Test AUC")
plt.legend(handler_map={line1: HandlerLine2D(numpoints=2)})
plt.ylabel("AUC score")
plt.xlabel("Tree depth")
# plt.show()
plt.savefig("tree_depth_optimization.pdf", format="pdf")
# +
min_samples_splits = np.linspace(0.1, 1.0, 10, endpoint=True)
train_results = []
test_results = []
for min_samples_split in min_samples_splits:
dtc = DecisionTreeClassifier(min_samples_split=min_samples_split)
dtc_pipe = Pipeline([('vectorizer', vectorizer), ('dtc', dtc)])
dtc_pipe.fit(train_titles,train_classes)
train_pred = dtc_pipe.predict(train_titles)
false_positive_rate, true_positive_rate, thresholds = roc_curve(train_classes, train_pred)
roc_auc = auc(false_positive_rate, true_positive_rate)
# Add auc score to previous train results
train_results.append(roc_auc)
y_pred = dtc_pipe.predict(test_titles)
false_positive_rate, true_positive_rate, thresholds = roc_curve(test_classes, y_pred)
roc_auc = auc(false_positive_rate, true_positive_rate)
# Add auc score to previous test results
test_results.append(roc_auc)
line1, = plt.plot(min_samples_splits, train_results, 'b', label="Train AUC")
line2, = plt.plot(min_samples_splits, test_results, 'r', label="Test AUC")
plt.legend(handler_map={line1: HandlerLine2D(numpoints=2)})
plt.ylabel("AUC score")
plt.xlabel("Min Samples Split")
# plt.show()
plt.savefig("min_samples_splits_optimization.pdf", format="pdf")
# +
min_samples_leafs = np.linspace(0.1, 0.5, 5, endpoint=True)
train_results = []
test_results = []
for min_samples_leaf in min_samples_leafs:
dtc = DecisionTreeClassifier(min_samples_leaf=min_samples_leaf)
dtc_pipe = Pipeline([('vectorizer', vectorizer), ('dtc', dtc)])
dtc_pipe.fit(train_titles,train_classes)
train_pred = dtc_pipe.predict(train_titles)
false_positive_rate, true_positive_rate, thresholds = roc_curve(train_classes, train_pred)
roc_auc = auc(false_positive_rate, true_positive_rate)
# Add auc score to previous train results
train_results.append(roc_auc)
y_pred = dtc_pipe.predict(test_titles)
false_positive_rate, true_positive_rate, thresholds = roc_curve(test_classes, y_pred)
roc_auc = auc(false_positive_rate, true_positive_rate)
# Add auc score to previous test results
test_results.append(roc_auc)
line1, = plt.plot(min_samples_leafs, train_results, 'b', label="Train AUC")
line2, = plt.plot(min_samples_leafs, test_results, 'r', label="Test AUC")
plt.legend(handler_map={line1: HandlerLine2D(numpoints=2)})
plt.ylabel("AUC score")
plt.xlabel("min samples leaf")
# plt.show()
plt.savefig("min_samples_leafs_splits_optimization.pdf", format="pdf")
# -
# Progress Bar Function
# Print iterations progress
def printProgressBar (iteration, total, prefix = '', suffix = '', decimals = 1, length = 100, fill = '█', printEnd = "\r"):
"""
Call in a loop to create terminal progress bar
@params:
iteration - Required : current iteration (Int)
total - Required : total iterations (Int)
prefix - Optional : prefix string (Str)
suffix - Optional : suffix string (Str)
decimals - Optional : positive number of decimals in percent complete (Int)
length - Optional : character length of bar (Int)
fill - Optional : bar fill character (Str)
printEnd - Optional : end character (e.g. "\r", "\r\n") (Str)
"""
percent = ("{0:." + str(decimals) + "f}").format(100 * (iteration / float(total)))
filledLength = int(length * iteration // total)
bar = fill * filledLength + '-' * (length - filledLength)
print('\r%s |%s| %s%% %s' % (prefix, bar, percent, suffix), end = printEnd)
# Print New Line on Complete
if iteration == total:
print()
# Find the best combination of parameters.
# +
warnings.filterwarnings('ignore')
max_depths = np.linspace(7, 12, 5, endpoint=True)
min_samples_splits = np.linspace(np.nextafter(0,1), 0.1, 20, endpoint=True)
min_samples_leafs = np.linspace(np.nextafter(0,1), 0.1, 20, endpoint=True)
criterion="gini"
class_weight = "balanced"
splitter = "best"
paramCombinations = cartesian([max_depths, min_samples_splits, min_samples_leafs])
data = []
columns = ['max_depth', 'min_samples_split', 'min_samples_leaf', 'train_accuracy_score', 'train_f1_score', 'train_recall_score', 'train_roc_auc_score', 'test_accuracy_score','test_f1_score','test_recall_score','test_roc_auc_score']
count = 0
for pComb in paramCombinations:
printProgressBar(count, len(paramCombinations), prefix = 'Progress:', suffix = 'Complete', length = 50)
# print advancement percentage
# if(((count/len(paramCombinations)*100) % 1) == 0):
# clear_output()
# print(str("%d%%"%(count/len(paramCombinations)*100)))
dtc = DecisionTreeClassifier(max_depth = pComb[0], min_samples_split = pComb[1], min_samples_leaf = pComb[2],
criterion=criterion,splitter=splitter,class_weight=class_weight)
dtc_pipe = Pipeline([('vectorizer', vectorizer), ('dtc', dtc)])
dtc_pipe.fit(train_titles,train_classes)
train_pred = dtc_pipe.predict(train_titles)
train_accuracy_score = accuracy_score(train_classes, train_pred)
train_f1_score = f1_score(train_classes, train_pred, average='weighted')
# train_recall_score = recall_score(train_classes, train_pred, average='weighted')
# train_f1_score = f1_score(train_classes, train_pred, average='binary',pos_label = 1)
train_recall_score = recall_score(train_classes, train_pred, average='binary',pos_label = 1)
train_roc_auc_score = roc_auc_score(train_classes, train_pred)
test_pred = dtc_pipe.predict(test_titles)
test_accuracy_score = accuracy_score(test_classes, test_pred)
test_f1_score = f1_score(test_classes, test_pred, average='weighted')
# test_recall_score = recall_score(test_classes, test_pred, average='weighted')
# test_f1_score = f1_score(test_classes, test_pred, average='binary',pos_label = 1)
test_recall_score = recall_score(test_classes, test_pred, average='binary',pos_label = 1)
test_roc_auc_score = roc_auc_score(test_classes, test_pred)
data.append((pComb[0],pComb[1],pComb[2],train_accuracy_score, train_f1_score, train_recall_score,train_roc_auc_score,
test_accuracy_score, test_f1_score, test_recall_score,test_roc_auc_score))
count += 1
# create DataFrame using data
paramCombinationsDataFrame = pd.DataFrame(data, columns = columns)
paramCombinationsDataFrame.to_csv('models/parameters/bruteforce_param_combinations.csv', index=False, header=True)
# +
print("#Max Depts: %s"%len(max_depths))
print("Max Depts: %s"%max_depths)
print("#Min Samples Splits: %s"%len(min_samples_splits))
print("Min Samples Splits: %s"%min_samples_splits)
print("#Min Samples Leaves: %s"%len(min_samples_leafs))
print("Min Samples Leaves: %s"%min_samples_leafs)
bestTrainIndex, bestTrainRes = max(enumerate(train_results), key=operator.itemgetter(1))
# bestTrainRes = max(train_results)
# bestTrainIndex = train_results.index(max(train_results))
bestTrainParamCombination = paramCombinations[bestTrainIndex]
print(str("Training Set Best Combination: %s Index: %d Score: %f"%(bestTrainParamCombination,bestTrainIndex,bestTrainRes)))
bestTestIndex, bestTestRes = max(enumerate(test_results), key=operator.itemgetter(1))
# bestTestRes = max(test_results)
# bestTestIndex = test_results.index(max(test_results))
bestTestParamCombination = paramCombinations[bestTestIndex]
print(str("Test Set Best Combination: %s Index: %d Score: %f"%(bestTestParamCombination,bestTestIndex,bestTestRes)))
# -
# Save the model so that it can later be re-imported.
savePath = ".\models\\"
extension = ".sav"
filename = str('%sdtc_maxdepth_%s_minsampleleaf_%s%s'%(savePath, max_depth,min_samples_leaf,extension))
joblib.dump(dtc_pipe, filename)
# Use the model to predict stuff.
# +
savePath = ".\models\\"
extension = ".sav"
filename = str('%sdtc_maxdepth_%s_minsampleleaf_%s%s'%(savePath, max_depth,min_samples_leaf,extension))
model = joblib.load(filename)
unreadTitles = df['Title'][df['ReadTimeStamp'].isnull()].drop_duplicates()
preprocessedTitles = titlePreprocessing(unreadTitles)
predictedClasses = model.predict(preprocessedTitles)
class0Titles = set()
class1Titles = set()
for i in range(0,len(predictedClasses),1):
if(predictedClasses.item(i) == 1):
class1Titles.add(unreadTitles.iloc[i])
else:
class0Titles.add(unreadTitles.iloc[i])
# -
# Print the interesting titles.
for class1Title in class1Titles:
print(class1Title)
print("\n--- CLASSIFIER: %s ---" %filename)
print("--- %d interesting titles found out of %d ---" %(len(class1Titles), len(unreadTitles)))
# Export the interesting articles into a separate CSV file.
# +
unreadArticles = df[df['ReadTimeStamp'].isnull()]
indexes = set(unreadArticles.index)
class0Rows = set()
class1Rows = set()
for index, article in unreadArticles.iterrows():
if((article['Title'] in class1Titles) == False):
class0Rows.add(index)
class1Rows = indexes.difference(class0Rows)
class0Articles = unreadArticles.drop(class1Rows)
class1Articles = unreadArticles.drop(class0Rows)
print(unreadArticles.shape[0])
print("%d + %d = %d"%(class0Articles.shape[0],class1Articles.shape[0], class0Articles.shape[0]+class1Articles.shape[0]))
class0Articles.to_csv('class_0_articles.csv', index=False, header=True)
class1Articles.to_csv('class_1_articles.csv', index=False, header=True)
# -
|
binary_decision_tree.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Connecting content with JupyterHub and Binder
#
# Because Jupyter Books are built with Jupyter Notebooks, you can connect your online
# book with a Jupyter kernel running in the cloud. This lets readers quickly interact
# with your content in a traditional coding interface using either JupyterHub or BinderHub.
# This page describes a few ways to accomplish this.
#
# ## Creating interact buttons for BinderHub
#
# BinderHub can be used to build the environment needed to run a repository, and provides
# a link that lets others interact with that repository. If your Jupyter Book is hosted online
# on GitHub, you can automatically insert buttons that link to the Jupyter Notebook running in a BinderHub.
# When a user clicks the button, they'll be taken to a live version of the page. If your code
# doesn't require a significant amount of CPU or RAM, you can use the free, public BinderHub running
# at https://mybinder.org.
#
# To automatically include Binder link buttons in each page of your Jupyter Book, use the following
# configuration:
#
# ```yaml
# # Binder link settings
# use_binder_button : true # If 'true', add a binder button for interactive links
# ```
#
# In addition, you can configure the components of your Binder links, which control things like
# where your BinderHub exists, which repository is used to define the environment, etc. Here's
# an example configuration with some explanation of each field.
#
# ```yaml
# binderhub_url : "https://mybinder.org" # The URL for your BinderHub.
# binder_repo_base : "https://github.com/" # The site on which the textbook repository is hosted
# binder_repo_org : "jupyter" # The username or organization that owns this repository
# binder_repo_name : "jupyter-book" # The name of the repository on the web
# binder_repo_branch : "master" # The branch on which your textbook is hosted.
# binderhub_interact_text : "Interact" # The text that interact buttons will contain.
# ```
#
# ## Creating interact buttons for JupyterHub
#
# JupyterHub lets you host an online service that gives users their own Jupyter servers
# with an environment that you specify for them. It allows you to give users access to
# resources and hardware that you provision in the cloud, and allows you to authenticate users
# in order to control who has access to your hardware.
#
# Similar to Binder link buttons, you can also automatically include interact links that send
# your readers to a JupyterHub that is running a live, interactive version of your page. This
# is accomplished using the [nbgitpuller](https://github.com/jupyterhub/nbgitpuller) server
# extension.
#
# ```yaml
# use_jupyterhub_button : false # If 'true', display a button that will direct users to a JupyterHub (that you provide)
# ```
#
# You can configure the location of the JupyterHub (which you may set up on your own using a guide
# such as [zero to jupyterhub for kubernetes](https://z2jh.jupyter.org) or [the littlest jupyterhub](https://tljh.jupyter.org)) with the following configuration.
#
# ```yaml
# jupyterhub_url : "" # The URL for your JupyterHub.
# ```
# ## Letting users define their own JupyterHub location
#
# ✨**Experimental**✨
#
# If you use interact links with your Jupyter Book, you can also allow users to update
# these links to their own JupyterHub location by using parameters specified in the URL.
# If an interact button is present on a page, append the following to a page's URL in order
# to update where the link points:
#
# ```
# mybook.com/mypage?jupyterhub=myhuburl.com
# ```
#
# You should see a message displayed next to the interact link that lets the user know
# where the link now points. This can be useful if you'd like to share content but allow
# users to run this content wherever they like.
|
docs/old_docs/features/interact.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Multi-generation climate model inter-comparison
#
# This notebook uses [`intake-esm`](https://intake-esm.readthedocs.io/en/latest/) to ingest and organize climate model output from various model generations from 1990-2019 and [`xskillscore`](https://github.com/raybellwaves/xskillscore) to calculate model skill with respect to NCEP Reanalysis
# +
from matplotlib import pyplot as plt
import numpy as np
import pandas as pd
import xarray as xr
import xskillscore as xs
import xesmf as xe
from tqdm.autonotebook import tqdm # Fancy progress bars for our loops!
import intake
# util.py is in the local directory
# it contains code that is common across project notebooks
# or routines that are too extensive and might otherwise clutter
# the notebook design
import util
# %matplotlib inline
plt.rcParams['figure.figsize'] = 12, 6
# %config InlineBackend.figure_format = 'retina'
# -
# ## Data catalogs
col_dict = {}
if util.is_ncar_host():
col = intake.open_esm_datastore("../catalogs/glade-cmip6.json")
else:
col = intake.open_esm_datastore("../catalogs/pangeo-cmip6.json")
col_dict["CMIP6"] = col
col
col = intake.open_esm_datastore("../catalogs/adhoc-ipcc-ar.json")
col_dict["pre-CMIP6"] = col
col
mip_catalog_dict = {"FAR": "pre-CMIP6",
"SAR": "pre-CMIP6",
"TAR": "pre-CMIP6",
"CMIP6": "CMIP6"}
mip_ids = mip_catalog_dict.keys()
mip_ids
# ## The task at hand
#
# - Loop through the `source_id` for `experiment_id == 'historical'`
# - For each model:
# - select only a single ensemble member (at least for now)
# - Take a slice `time=slice('1981', '2010')` and compute time average over that slice (date range consistent with the long-term climatology of the NCEP reanalysis, to which we are comparing
# - Regrid the `tas` data to a common grid
# - drop redundant variables (like "height: 2m")
# - concatenate to ensemble dataset along `ensemble` dimension
# ### Defining a regridding function
# +
# Define the common target grid axes
dx, dy = 1., 1.
ds_out = xr.Dataset({'lat': (['lat'], np.arange(-90.+dy/2., 90., dy)),
'lon': (['lon'], np.arange(0.+dx/2., 360., dx)),})
# Regridding function
def regrid_to_common(ds, ds_out):
"""
Regrid from rectilinear grid to common grid
"""
regridder = xe.Regridder(ds, ds_out, 'bilinear',periodic=True, reuse_weights=True)
return regridder(ds)
# -
# ### Loading data
#
# `intake-esm` enables loading data directly into an [xarray.Dataset](http://xarray.pydata.org/en/stable/api.html#dataset).
#
# Note that data on the cloud are in
# [zarr](https://zarr.readthedocs.io/en/stable/) format and data on
# [glade](https://www2.cisl.ucar.edu/resources/storage-and-file-systems/glade-file-spaces) are stored as
# [netCDF](https://www.unidata.ucar.edu/software/netcdf/) files. This is opaque to the user.
#
# `intake-esm` has rules for aggegating datasets; these rules are defined in the collection-specification file.
#varnames = ['tas','psl','pr','uas','vas']
varnames = ['tas', 'pr']
time_slice = slice('1981', '2010') # date range consistent with NCEP reanalysis long-term-mean
# For converting units for precip output
cm_to_m = 1.e-2
rho_water = 1.e3
day_in_s = (24.*60.*60.)
# +
ds_dict = {}
for mip_id in tqdm(mip_ids):
ds_dict[mip_id] = {}
for varname in varnames:
col = col_dict[mip_catalog_dict[mip_id]]
cat = col.search(experiment_id='historical',
table_id='Amon',
variable_id=varname,
member_id='r1i1p1f1' # choose first ensemble member only (for now)
)
dset_dict = cat.to_dataset_dict(zarr_kwargs={'consolidated': True, 'decode_times': False})
ds_dict[mip_id][varname] = {}
for key, ds in dset_dict.items():
if (mip_catalog_dict[mip_id] == 'pre-CMIP6') and (mip_id != key.split(".")[-1]): continue
# rename spatial dimensions if necessary
if ('longitude' in ds.dims) and ('latitude' in ds.dims):
ds = ds.rename({'longitude':'lon', 'latitude': 'lat'})
ds = xr.decode_cf(ds) # Need this temporarily because setting 'decode_times': True appears broken
ds = ds.squeeze() # get rid of member_id (for now)
# take long-term mean
timeave = ds.sel(time=time_slice).mean(dim='time')
# modify pre-CMIP6 chunks
if mip_catalog_dict[mip_id] == 'pre-CMIP6':
timeave = timeave.chunk({'lat':timeave['lat'].size, 'lon':timeave['lon'].size})
# regrid to common grid
ds_new = regrid_to_common(timeave[varname], ds_out)
# Add metadata and apply various corrections
if mip_catalog_dict[mip_id] == 'CMIP6':
# Correct MCM-UA precipitation due to broken units (<NAME>, personal communication)
if ('MCM-UA' in ds.attrs['parent_source_id']) and (varname == 'pr'):
# convert from cm/day to kg/m^2/s
ds_new *= (cm_to_m * rho_water / day_in_s)
# TEMPORARY FIX: Correct BCC-ESM1 and CanESM5 which inexplicably have latitude flipped
if ("BCC-ESM1" in key) or ("CanESM5" in key):
ds_new['lat'].values = ds_new['lat'].values[::-1]
ds_new.attrs['name'] = ds.attrs['source_id']
else:
# Maybe chance this at pre-processing stage?
ds_new.attrs['name'] = ds.attrs['institution']
# drop redundant variables (like "height: 2m")
for coord in ds_new.coords:
if coord not in ['lat','lon']:
ds_new = ds_new.drop(coord)
# Add ensemble as new dimension
ds_new = ds_new.expand_dims({'ensemble': np.array([ds_new.attrs['name']])}, 0)
# Add var as new dimension
ds_new = ds_new.expand_dims({'var': np.array([varname])}, 0)
# We should keep the metadata!!!
ds_new.attrs['mip_id'] = mip_id
ds_dict[mip_id][varname][key] = ds_new # add this to the dictionary
# +
ens_dict = {}
for mip_id in mip_ids:
ens = (
xr.concat([
xr.concat([ds for name, ds in ds_dict[mip_id][varname].items()], dim='ensemble')
for varname in varnames
],dim='var')
)
ens[mip_id] = xr.concat(
[ens, ens.mean(dim='ensemble', skipna=True).expand_dims({'ensemble': np.array(['ensemble-mean'])}, 0)],
dim='ensemble'
)
ens.name = mip_id
ens_dict[mip_id] = ens
ens_dict
# -
# ## Some 'observational' reference data to compare models against
#
# We will use long-term mean climatology from NCEP reanalysis (because it's easy)
# +
# Paths gotten from:
# https://www.esrl.noaa.gov/psd/data/gridded/data.ncep.reanalysis.derived.surface.html
# https://www.esrl.noaa.gov/psd/data/gridded/data.ncep.reanalysis.derived.surfaceflux.html
ncep_url = "http://www.esrl.noaa.gov/psd/thredds/dodsC/Datasets/ncep.reanalysis.derived/"
ncep_paths_dict = {"tas": "surface/air.mon.ltm.nc",
"psl": "surface/slp.mon.ltm.nc",
"pr": "surface_gauss/prate.sfc.mon.ltm.nc",
"uas": "surface/uwnd.mon.ltm.nc",
"vas": "surface/vwnd.mon.ltm.nc"}
def convert_ncep_units_to_cf(da):
if da.attrs['units'] == "degC":
da += 273.15
da.attrs['units'] = "K"
elif da.attrs['units'] == "millibars":
da *= 100.
da.attrs['units'] = "Pa"
return da
ncep_list = []
for varname, path in ncep_paths_dict.items():
# Check that we read in same variables as models
if varname not in varnames: continue
ncep_ds = xr.open_dataset(ncep_url + path)
ncep_var_name = path.split("/")[1].split(".")[0]
# Long-term monthly climatology 1981-2010
ncep_var_native = ncep_ds[ncep_var_name]
# Convert NCEP data to CF units
ncep_var_native = convert_ncep_units_to_cf(ncep_var_native)
# Long-term mean 1981-2010
ncep_var_native = ncep_var_native.mean(dim='time')
# Regrid to common grid
ncep_var_regridded = regrid_to_common(ncep_var_native, ds_out)
# keep all the original attributes
ncep_var_regridded.attrs.update(ncep_ds.attrs)
# Expand dimensions
ncep_var_regridded = ncep_var_regridded.expand_dims({'var': np.array([varname])}, 0)
ncep_list.append(ncep_var_regridded)
# -
obs = xr.concat([ds for ds in ncep_list], dim='var')
obs.name = "NCEP"
big_plots = True
if big_plots:
for ens in ens_dict.values():
for varname in varnames:
# Plot mean state
plt.figure()
q = ens.sel(var=varname).plot(x='lon', y='lat', col='ensemble', col_wrap=6)
plt.title(ens.name+" "+varname+" long-term mean")
# Plot anomalies with respect to observations (temporarily, actually multi-model mean)
plt.figure()
q = (ens-obs).sel(var=varname).plot(x='lon', y='lat', col='ensemble', col_wrap=6)
plt.title(ens.name+" "+varname+" long-term mean bias")
# How to change color bar limits?
break
# ## Regionmask
### create mask for 'obs' and 'ens'
import regionmask
mask = regionmask.defined_regions.giorgi.mask(obs, wrap_lon=True)
ssa_mask = regionmask.defined_regions.giorgi.map_keys('SSA')
wna_mask = regionmask.defined_regions.giorgi.map_keys('WNA')
obs_ssa_mask = obs.where(mask == ssa_mask)
obs_wna_mask = obs.where(mask == wna_mask)
obs_ssa_mask[0].plot()
obs_wna_mask[0].plot()
# +
masked_dict = {}
for key, ens in ens_dict.items():
masked_dict[key] = ens.copy().where(mask == wna_mask)
# +
rmse_masked_dict = {}
for key, ens in tqdm(masked_dict.items()):
rmse_masked_dict[key] = xs.rmse(obs_wna_mask, ens, ['lat', 'lon']).compute()
# +
rmse_dict = {}
for key, ens in ens_dict.items():
rmse_dict[key] = xs.rmse(obs, ens, ['lat', 'lon']).compute()
# -
rmse_dict
# +
rmse_dict = {}
for key, ens in ens_dict.items():
rmse_dict[key] = xs.rmse(obs, ens, ['lat', 'lon']).compute()
# -
rmse_dict['FAR']
rmse_med = rmse_dict['FAR'].sel(ensemble=[key for key in rmse_dict['FAR'].ensemble.values if key!='ens-mean']).median(dim='ensemble').compute()
rmse_med
# +
skillscore_dict = {}
for key, ens in tqdm(rmse_dict.items()):
skillscore_dict[key] = ens / rmse_med
# -
skillscore_dict
# ## Pick up here with scatterplots
tas = skillscore_dict['FAR'].sel(var='tas')
pr = skillscore_dict['FAR'].sel(var='pr')
plt.scatter(skillscore_dict['FAR'].sel(var='tas'), skillscore_dict['FAR'].sel(var='pr'))
for key, ens in skillscore_dict.items():
plt.figure(figsize=(10,3))
q = plt.scatter(skillscore_dict[key].sel(var='tas'), skillscore_dict[key].sel(var='pr'))
# q = carpet_plot(skillscore_dict[key])
plt.title("Climate model skill metrics in "+key)
# q.colorbar.set_label("normalized mean absolute error")
plt.scatter(skillscore_dict['FAR'].sel(var='tas'), skillscore_dict['FAR'].sel(var='pr'), label = 'FAR', alpha = 0.5)
plt.scatter(skillscore_dict['SAR'].sel(var='tas'), skillscore_dict['SAR'].sel(var='pr'), label = 'SAR', alpha = 0.5)
plt.scatter(skillscore_dict['TAR'].sel(var='tas'), skillscore_dict['TAR'].sel(var='pr'), label = 'TAR', alpha = 0.5)
plt.scatter(CMIP6_dict.sel(var='tas'), CMIP6_dict.sel(var='pr'), label = 'CMIP6', alpha = 0.5)
plt.legend()
plt.ylabel('precipitation rate normalized error')
plt.xlabel('near-surface air temperature normalized error')
plt.title('Climate model skill metrics');
plt.savefig('model_performance_scatter.png', dpi=100)
skillscore_dict['FAR'].sel(var='tas')
# calculate correlations
FAR_pearson_corr = xs.pearson_r(skillscore_dict['FAR'].sel(var='tas'), skillscore_dict['FAR'].sel(var='pr'), dim = ['ensemble'])
FAR_pearson_corr
SAR_pearson_corr = xs.pearson_r(skillscore_dict['SAR'].sel(var='tas'), skillscore_dict['SAR'].sel(var='pr'), dim = ['ensemble'])
SAR_pearson_corr
TAR_pearson_corr = xs.pearson_r(skillscore_dict['TAR'].sel(var='tas'), skillscore_dict['TAR'].sel(var='pr'), dim = ['ensemble'])
TAR_pearson_corr
CMIP6_pearson_corr = xs.pearson_r(CMIP6_dict.sel(var='tas'), CMIP6_dict.sel(var='pr'), dim = ['ensemble'])
CMIP6_pearson_corr
CMIP6_dict.sel(var='tas')
CMIP6_dict = skillscore_dict['CMIP6'].sel(ensemble = [ensemble for ensemble in rmse_dict['CMIP6'].ensemble.values if np.logical_not(np.isnan(rmse_dict['CMIP6'].sel(ensemble = ensemble)).any())])
CMIP6_dict
[key for key in rmse_dict['CMIP6'].sel(var='tas') if key!= np.isnan(rmse_dict['CMIP6'].sel(var='tas'))]
CMIP6_dict = rmse_dict['CMIP6'].sel(ensemble = [key for key in rmse_dict['CMIP6'].sel(var='tas') if (rmse_dict['CMIP6'].sel(var='tas')).any() != np.nan])
CMIP6_dict
# to deal with NaNs in CMIP6
CMIP6_dict = rmse_dict['CMIP6'].sel(ensemble = [key for key in rmse_dict['CMIP6'].sel(var='tas') if not np.isnan(rmse_dict['CMIP6'].sel(var='tas', ensemble=key))])
np.isnan(rmse_dict['CMIP6'].sel(var='tas'))
# to deal with NaNs in CMIP6
CMIP6_dict = rmse_dict['CMIP6'].sel(ensemble = [key for key in rmse_dict['CMIP6'].ensemble])
CMIP6_dict = rmse_dict['CMIP6'].sel(ensemble=[key for key in rmse_dict['CMIP6'].var if key != 'nan'])
#rmse_med = rmse_dict['FAR'].sel(ensemble=[key for key in rmse_dict['FAR'].ensemble.values if key!='ens-mean']).median(dim='ensemble').compute()
for ens in ens_dict.values():
for varname in varnames:
# Plot mean state
plt.figure()
q = ens.sel(var=varname).plot(x='lon', y='lat', col='ensemble', col_wrap=6)
plt.title(ens.name+" "+varname+" long-term mean")
# ## Computing skill metrics
rmse = xs.rmse(obs, ens, ['lat', 'lon']).compute()
rmse_med = rmse.median(dim='ensemble').compute()
skill_score = rmse/rmse_med
def carpet_plot(da):
coords = list(da.coords)
q = plt.pcolormesh(da.values, cmap='RdYlBu_r')
plt.yticks(np.arange(da.coords[coords[0]].size)+0.5, list(da.coords[coords[0]].values))
plt.xticks(np.arange(da.coords[coords[1]].size)+0.5, list(da.coords[coords[1]].values), rotation=90.)
plt.colorbar()
plt.clim([0.5,1.5])
return q
for key, ens in ens_dict.items():
plt.figure(figsize=(10,3))
q = carpet_plot(skillscore_dict[key])
plt.title("Climate model skill metrics in "+key)
q.colorbar.set_label("normalized mean absolute error")
plt.figure(figsize=(12,6))
q = carpet_plot(skill_score)
q.colorbar.set_label("normalized model error")
|
notebooks/var_skill_scatter_plot_avc.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="xx5CxeVubU3W" colab_type="text"
# # Hypothesis Testing
# + [markdown] id="6OR23eWKbU3X" colab_type="text"
# The purpose of the test is to tell if there is any significant difference between two data sets.
#
#
# + [markdown] id="604OoWnTbU3Z" colab_type="text"
# ## Overview
# + [markdown] id="o-il1iBSbU3b" colab_type="text"
# This module covers,
#
# 1) One sample and Two sample t-tests
#
# 2) ANOVA
#
# 3) Type I and Type II errors
#
# 4) Chi-Squared Tests
# + [markdown] id="G2yYl5cvbU3c" colab_type="text"
# ## Question 1
#
# *A student is trying to decide between two GPUs. He want to use the GPU for his research to run Deep learning algorithms, so the only thing he is concerned with is speed.*
#
# *He picks a Deep Learning algorithm on a large data set and runs it on both GPUs 15 times, timing each run in hours. Results are given in the below lists GPU1 and GPU2.*
# + id="ohh5XJ4ZbU3d" colab_type="code" colab={}
from scipy import stats
import numpy as np
# + id="6yGkYI6EbU3i" colab_type="code" colab={}
GPU1 = np.array([11,9,10,11,10,12,9,11,12,9,11,12,9,10,9])
GPU2 = np.array([11,13,10,13,12,9,11,12,12,11,12,12,10,11,13])
#Assumption: Both the datasets (GPU1 & GPU 2) are random, independent, parametric & normally distributed
# + [markdown] id="L55jqmIXbU3m" colab_type="text"
# Hint: You can import ttest function from scipy to perform t tests
# + [markdown] id="v8fsIhPFbU3n" colab_type="text"
# **First T test**
#
# *One sample t-test*
#
# Check if the mean of the GPU1 is equal to zero.
# - Null Hypothesis is that mean is equal to zero.
# - Alternate hypothesis is that it is not equal to zero.
# + id="wgMSWwApbU3o" colab_type="code" colab={}
# + [markdown] id="byu8iw46bU3v" colab_type="text"
# ## Question 2
# + [markdown] id="E65pzWcJbU3w" colab_type="text"
# Given,
#
# Null Hypothesis : There is no significant difference between data sets
#
# Alternate Hypothesis : There is a significant difference
#
# *Do two-sample testing and check whether to reject Null Hypothesis or not.*
# + [markdown] id="N-zpWvyXbU32" colab_type="text"
# https://docs.scipy.org/doc/scipy/reference/generated/scipy.stats.ttest_ind.html
# + id="UDDkuOtObU3x" colab_type="code" colab={}
# + [markdown] id="MbXv5aZvbU33" colab_type="text"
# ## Question 3
#
# He is trying a third GPU - GPU3.
# + id="kkh_sQl4bU34" colab_type="code" colab={}
GPU3 = np.array([9,10,9,11,10,13,12,9,12,12,13,12,13,10,11])
#Assumption: Both the datasets (GPU1 & GPU 3) are random, independent, parametric & normally distributed
# + [markdown] id="WoYNz3g7bU37" colab_type="text"
# *Do two-sample testing and check whether there is significant differene between speeds of two GPUs GPU1 and GPU3.*
#
# #### Answer:
# + id="X4N11XArbU38" colab_type="code" colab={}
# + [markdown] id="oyDFS4WZbU4A" colab_type="text"
# ## ANOVA
# + [markdown] id="wknPzstFbU4B" colab_type="text"
# ## Question 4
#
# If you need to compare more than two data sets at a time, an ANOVA is your best bet.
#
# *The results from three experiments with overlapping 95% confidence intervals are given below, and we want to confirm that the results for all three experiments are not significantly different.*
#
# But before conducting ANOVA, test equality of variances (using Levene's test) is satisfied or not. If not, then mention that we cannot depend on the result of ANOVA
# + id="kGb0GeK8bU4C" colab_type="code" colab={}
import numpy as np
e1 = np.array([1.595440,1.419730,0.000000,0.000000])
e2 = np.array([1.433800,2.079700,0.892139,2.384740])
e3 = np.array([0.036930,0.938018,0.995956,1.006970])
#Assumption: All the 3 datasets (e1,e2 & e3) are random, independent, parametric & normally distributed
# + [markdown] id="_xiYN6gVbU4G" colab_type="text"
# Perform levene test on the data
#
# The Levene test tests the null hypothesis that all input samples are from populations with equal variances. Levene’s test is an alternative to Bartlett’s test bartlett in the case where there are significant deviations from normality.
#
# source: scipy.org
# + [markdown] id="f2MlJTXgbU4H" colab_type="text"
# #### Answer:
# + id="VUJP_GGQbU4R" colab_type="code" colab={}
# + [markdown] id="FigxGCQtbU4Y" colab_type="text"
# ## Question 5
#
# The one-way ANOVA tests the null hypothesis that two or more groups have the same population mean. The test is applied to samples from two or more groups, possibly with differing sizes.
#
# use stats.f_oneway() module to perform one-way ANOVA test
# + id="P3bPYPCbbU4Z" colab_type="code" colab={}
# + [markdown] id="yrMK1qb7bU4j" colab_type="text"
# ## Question 6
#
# *In one or two sentences explain about **TypeI** and **TypeII** errors.*
#
# #### Answer:
# + [markdown] id="H08OGwdIbU4k" colab_type="text"
# answer here
# + [markdown] id="hm7v3pcIbU4m" colab_type="text"
# ## Question 7
#
# You are a manager of a chinese restaurant. You want to determine whether the waiting time to place an order has changed in the past month from its previous population mean value of 4.5 minutes.
# State the null and alternative hypothesis.
#
# #### Answer:
#
# + [markdown] id="pbbaU2I4bU4n" colab_type="text"
# answer here
# + [markdown] id="1SGpq-dKbU4r" colab_type="text"
# ## Chi square test
# + [markdown] id="WlxmIu_rdgpc" colab_type="text"
# ## Question 8
# + [markdown] id="8J3V015PbU4s" colab_type="text"
# Let's create a small dataset for dice rolls of four players
# + id="xrO5BbIEbU4t" colab_type="code" colab={}
import numpy as np
d1 = [5, 8, 3, 8]
d2 = [9, 6, 8, 5]
d3 = [8, 12, 7, 2]
d4 = [4, 16, 7, 3]
d5 = [3, 9, 6, 5]
d6 = [7, 2, 5, 7]
dice = np.array([d1, d2, d3, d4, d5, d6])
# + [markdown] id="uF7GRMChbU4x" colab_type="text"
# run the test using SciPy Stats library
#
# Depending on the test, we are generally looking for a threshold at either 0.05 or 0.01. Our test is significant (i.e. we reject the null hypothesis) if we get a p-value below our threshold.
#
# For our purposes, we’ll use 0.01 as the threshold.
#
# use stats.chi2_contingency() module
#
# This function computes the chi-square statistic and p-value for the hypothesis test of independence of the observed frequencies in the contingency table
#
# Print the following:
#
# - chi2 stat
# - p-value
# - degree of freedom
# - contingency
#
#
# + id="vqaTIKmgbU4y" colab_type="code" colab={}
# + [markdown] id="VzyfaNoabU42" colab_type="text"
# ## Question 9
#
# ### Z-test
#
# Get zscore on the above dice data using stats.zscore module from scipy. Convert zscore values to p-value and take mean of the array.
# + id="Sd5xPCuRbU43" colab_type="code" colab={}
# + [markdown] id="OyoFUxf5bU47" colab_type="text"
# ## Question 10
#
# A Paired sample t-test compares means from the same group at different times.
#
# The basic two sample t-test is designed for testing differences between independent groups.
# In some cases, you might be interested in testing differences between samples of the same group at different points in time.
# We can conduct a paired t-test using the scipy function stats.ttest_rel().
# + id="SwVmQ1gRbU48" colab_type="code" colab={}
before= stats.norm.rvs(scale=30, loc=100, size=500) ## Creates a normal distribution with a mean value of 100 and std of 30
after = before + stats.norm.rvs(scale=5, loc=-1.25, size=500)
# + [markdown] id="rB_os5FjbU4_" colab_type="text"
# Test whether a weight-loss drug works by checking the weights of the same group patients before and after treatment using above data.
# + id="Fq2wyEqlbU5A" colab_type="code" colab={}
|
Hypothesis_testing.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.1.0
# language: julia
# name: julia-1.1
# ---
# # Testing Navier-Stokes in package `ViscousFlow`
using ViscousFlow
using Plots
pyplot()
clibrary(:colorbrewer)
default(grid = false)
# ### Non-linear term basic calculation
using Random
Random.seed!(1);
nx = 129; ny = 129;
w = Nodes(Dual,(nx,ny))
w .= rand(Float64,size(w));
Qq = Edges(Dual,w);
Ww = deepcopy(Qq);
ψ = deepcopy(w);
L = plan_laplacian(w,with_inverse=true)
@time nl = divergence(cellshift!(Qq,curl(L\w))∘cellshift!(Ww,w));
# ### Solve the Lamb-Oseen vortex
# First, construct the exact solution
woseen(x::Tuple{Real,Real},t;Re=1.0,x0::Tuple{Real,Real}=(0,0),t0=1) =
exp(-((x[1]-x0[1])^2+(x[2]-x0[2])^2)/(4(t+t0)/Re))/(1+t/t0)
Re = 200 + 50rand()
U = 1.0 + 0.2randn()
U∞ = (U,0.0)
Δx = 0.015;
Δt = min(0.5*Δx,0.5*Δx^2*Re);
xlim = (0.0,3.0);
ylim = (0.0,2.0);
# Construct exact solution in shape of grid data
sys = Systems.NavierStokes(Re,Δx,xlim,ylim,Δt,U∞ = U∞)
w₀ = Nodes(Dual,size(sys));
xg,yg = coordinates(w₀,dx=Δx,I0=Systems.origin(sys))
x0 = (1.0,1.0); t0 = 1;
wexact(t) = [woseen((x,y),t;Re=Re,x0=x0.+U∞.*t,t0=t0) for x in xg, y in yg]
ifrk = IFRK(w₀,sys.Δt,
(t,w) -> Systems.plan_intfact(t,w,sys),
(w,t) -> Systems.r₁(w,t,sys) ,rk=TimeMarching.RK31)
t = 0.0
w₀ .= wexact(t)
w = deepcopy(w₀)
tf = 1.0
T = 0:Δt:tf;
t = 0.0;
for ti in T
global t, w = ifrk(t,w)
end
using LinearAlgebra
LinearAlgebra.norm(w-wexact(t),Inf)
plot(xg,yg,w)
plot(xg,w[:,65],label="numerical",ylim=(0,1))
plot!(xg,wexact(t)[:,65],label="exact")
|
examples/Basic Navier-Stokes.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# =============================================
# Example: detecting defaults on retail credits
# =============================================
#
#
# SkopeRules finds logical rules with high precision and fuse them. Finding
# good rules is done by fitting classification and regression trees
# to sub-samples.
# A fitted tree defines a set of rules (each tree node defines a rule); rules
# are then tested out of the bag, and the ones with higher precision are kept.
#
# This example aims at finding logical rules to predict credit defaults. The
# analysis shows that setting.
#
#
#
# Data import and preparation
# ...........................
#
# There are 3 categorical variables (SEX, EDUCATION and MARRIAGE) and 20
# numerical variables.
# The target (credit defaults) is transformed in a binary variable with
# integers 0 (no default) and 1 (default).
# From the 30000 credits, 50% are used for training and 50% are used
# for testing. The target is unbalanced with a 22%/78% ratio.
#
#
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.metrics import roc_curve, precision_recall_curve
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import GridSearchCV
from sklearn.utils import shuffle
from skrules import SkopeRules
from skrules.datasets import load_credit_data
print(__doc__)
rng = np.random.RandomState(1)
# Importing data
dataset = load_credit_data()
X = dataset.data
y = dataset.target
# Shuffling data, preparing target and variables
data, y = shuffle(np.array(X), y, random_state=rng)
data = pd.DataFrame(data, columns=X.columns)
for col in ['ID']:
del data[col]
# Quick feature engineering
data = data.rename(columns={"PAY_0": "PAY_1"})
old_PAY = ['PAY_3', 'PAY_4', 'PAY_5', 'PAY_6']
data['PAY_old_mean'] = data[old_PAY].apply(lambda x: np.mean(x), axis=1)
old_BILL_AMT = ['BILL_AMT3', 'BILL_AMT4', 'BILL_AMT5', 'BILL_AMT6']
data['BILL_AMT_old_mean'] = data[old_BILL_AMT].apply(
lambda x: np.mean(x), axis=1)
data['BILL_AMT_old_std'] = data[old_BILL_AMT].apply(
lambda x: np.std(x),
axis=1)
old_PAY_AMT = ['PAY_AMT3', 'PAY_AMT4', 'PAY_AMT5', 'PAY_AMT6']
data['PAY_AMT_old_mean'] = data[old_PAY_AMT].apply(
lambda x: np.mean(x), axis=1)
data['PAY_AMT_old_std'] = data[old_PAY_AMT].apply(
lambda x: np.std(x), axis=1)
data.drop(old_PAY_AMT + old_BILL_AMT + old_PAY, axis=1, inplace=True)
# Creating the train/test split
feature_names = list(data.columns)
print("List of variables used to train models : " + str(feature_names))
data = data.values
n_samples = data.shape[0]
n_samples_train = int(n_samples / 2)
y_train = y[:n_samples_train]
y_test = y[n_samples_train:]
X_train = data[:n_samples_train]
X_test = data[n_samples_train:]
# -
# Benchmark with a Random Forest classifier
# .........................................
#
# This part shows the training and performance evaluation of a random forest
# model. The objective remains to extract rules which targets credit defaults.
#
#
# +
rf = GridSearchCV(
RandomForestClassifier(
random_state=rng,
n_estimators=50,
class_weight='balanced'),
param_grid={'max_depth': range(3, 8, 1),
'max_features': np.linspace(0.1, 1., 5)},
scoring={'AUC': 'roc_auc'}, cv=5,
refit='AUC', n_jobs=-1)
rf.fit(X_train, y_train)
scoring_rf = rf.predict_proba(X_test)[:, 1]
print("Random Forest selected parameters : %s" % rf.best_params_)
# Plot ROC and PR curves
fig, axes = plt.subplots(1, 2, figsize=(12, 5),
sharex=True, sharey=True)
ax = axes[0]
fpr_RF, tpr_RF, _ = roc_curve(y_test, scoring_rf)
ax.step(fpr_RF, tpr_RF, linestyle='-.', c='g', lw=1, where='post')
ax.set_title("ROC", fontsize=20)
ax.legend(loc='upper center', fontsize=8)
ax.set_xlabel('False Positive Rate', fontsize=18)
ax.set_ylabel('True Positive Rate (Recall)', fontsize=18)
ax = axes[1]
precision_RF, recall_RF, _ = precision_recall_curve(y_test, scoring_rf)
ax.step(recall_RF, precision_RF, linestyle='-.', c='g', lw=1, where='post')
ax.set_title("Precision-Recall", fontsize=20)
ax.set_xlabel('Recall (True Positive Rate)', fontsize=18)
ax.set_ylabel('Precision', fontsize=18)
plt.show()
# -
# The ROC and Precision-Recall curves illustrate the performance of Random
# Forests in this classification task.
# Suppose now that we add an interpretability contraint to this setting:
# Typically, we want to express our model in terms of logical rules detecting
# defaults. A random forest could be expressed in term of weighted sum of
# rules, but 1) such a large weighted sum, is hardly interpretable and 2)
# simplifying it by removing rules/weights is not easy, as optimality is
# targeted by the ensemble of weighted rules, not by each rule.
# In the following section, we show how SkopeRules can be used to produce
# a number of rules, each seeking for high precision on a potentially small
# area of detection (low recall).
#
#
# Getting rules with skrules
# ..........................
#
# This part shows how SkopeRules can be fitted to detect credit defaults.
# Performances are compared with the random forest model previously trained.
#
#
# +
# fit the model
clf = SkopeRules(
similarity_thres=.8, max_depth=3, max_features=0.5,
max_samples_features=0.5, random_state=rng, n_estimators=20,
feature_names=feature_names, recall_min=0.04, precision_min=0.6)
clf.fit(X_train, y_train)
# in the score_top_rules method, a score of k means that rule number k
# vote positively, but not rules 1, ..., k-1. It will allow us to plot
# performance of each rule separately on the ROC and PR plots.
scoring = clf.score_top_rules(X_test)
print(str(len(clf.rules_)) + ' rules have been built.')
print('The 5 most precise rules are the following:')
for rule in clf.rules_[:5]:
print(rule[0])
curves = [roc_curve, precision_recall_curve]
xlabels = ['False Positive Rate', 'Recall (True Positive Rate)']
ylabels = ['True Positive Rate (Recall)', 'Precision']
fig, axes = plt.subplots(1, 2, figsize=(12, 5),
sharex=True, sharey=True)
ax = axes[0]
fpr, tpr, _ = roc_curve(y_test, scoring)
fpr_rf, tpr_rf, _ = roc_curve(y_test, scoring_rf)
ax.scatter(fpr[:-1], tpr[:-1], c='b', s=10, label="rules of SkopeRules")
ax.step(fpr_RF, tpr_RF, linestyle='-.', c='g', lw=1, where='post',
label="Random Forest")
ax.set_title("ROC", fontsize=20)
ax.legend(loc='upper center', fontsize=8)
ax.set_xlabel('False Positive Rate', fontsize=18)
ax.set_ylabel('True Positive Rate (Recall)', fontsize=18)
ax = axes[1]
precision, recall, _ = precision_recall_curve(y_test, scoring)
precision_rf, recall_rf, _ = precision_recall_curve(y_test, scoring_rf)
ax.scatter(recall[1:-1], precision[1:-1], c='b', s=10,
label="rules of SkopeRules")
ax.step(recall_RF, precision_RF, linestyle='-.', c='g', lw=1, where='post',
label="Random Forest")
ax.set_title("Precision-Recall", fontsize=20)
ax.set_xlabel('Recall (True Positive Rate)', fontsize=18)
ax.set_ylabel('Precision', fontsize=18)
plt.show()
# -
# The ROC and Precision-Recall curves show the performance of the rules
# generated by SkopeRules the (the blue points) and the performance of the
# Random Forest classifier fitted above.
# Each blue point represents the performance of a set of rules: Starting from
# the left on the precision-recall cruve, the kth point
# represents the score associated to the concatenation (union) of the k first
# rules, etc. Thus, each blue point is associated with an interpretable
# classifier, which is a combination of a few rules.
# In terms of performance, each of these interpretable classifiers compare well
# with Random Forest, while offering complete interpretation.
# The range of recall and precision can be controlled by the precision_min and
# recall_min parameters. Here, setting precision_min to 0.6 force the rules to
# have a limited recall.
#
#
|
doc/_build/html/_downloads/plot_credit_default.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
dataset = pd.read_csv('../0_datasets/credit_data.csv')
dataset.shape
dataset.dropna(inplace=True)
dataset.shape
dataset.head()
# ### Sem pradronização
X = dataset.iloc[:, 1:4].values
X
y = dataset.iloc[:, 4].values
y
from sklearn.neighbors import KNeighborsClassifier
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2,
stratify=y)
# Salários da base de treinamento
np.mean(X_train[0]), np.median(X_train[0]), np.std(X_train[0])
# Salários da base de testes
np.mean(X_test[0]), np.median(X_test[0]), np.std(X_test[0])
knn = KNeighborsClassifier()
knn.fit(X_train, y_train)
previsoes = knn.predict(X_test)
accuracy_score(y_test, previsoes)
# ### Com padronização
from sklearn.preprocessing import StandardScaler
# +
z_score_train = StandardScaler()
z_score_test = StandardScaler()
X_train_p = z_score_train.fit_transform(X_train)
X_test_p = z_score_test.fit_transform(X_test)
# -
X_train_p, X_test_p
min(X_train_p[0]), max(X_train_p[0]), min(X_test_p[0]), max(X_test_p[0])
# Salários da base de treinamento
np.mean(X_train_p), np.median(X_train_p), np.std(X_train_p)
# Salários da base de testes
np.mean(X_test_p), np.median(X_test_p), np.std(X_test_p)
knn = KNeighborsClassifier()
knn.fit(X_train_p, y_train)
previsoes = knn.predict(X_test_p)
accuracy_score(y_test, previsoes)
|
6_distribuicoes_estatisticas/z-score_knn.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="8-oXiSGwTFGJ"
# ##Task 2 Apply algorithm on breast cancer wisconsin dataset - One Hot Encoding of features: and Train test Division 60%-40%
# + id="19-VWLo5dRM_" executionInfo={"status": "ok", "timestamp": 1629714859946, "user_tz": -330, "elapsed": 1202, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0Ix_OxEFfMvg474o7FEouUO3wMGLVoXqHui807w=s64", "userId": "01938360022046049253"}}
#Import scikit-learn dataset library
from sklearn import datasets
from sklearn.tree import DecisionTreeClassifier
import pandas as pd
import numpy as np
#Load dataset
data = datasets.load_breast_cancer()
# + colab={"base_uri": "https://localhost:8080/"} id="STpVNWyrdRNb" executionInfo={"status": "ok", "timestamp": 1629714867528, "user_tz": -330, "elapsed": 631, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0Ix_OxEFfMvg474o7FEouUO3wMGLVoXqHui807w=s64", "userId": "01938360022046049253"}} outputId="73f80824-59d2-449d-d907-4e7393331281"
# print the names of the features
print("Features: ", data.feature_names)
# print the label type of breast cancer
print("\n class: \n",data.target_names)
# print data(feature)shape
print( "\n",data.data.shape)
# + id="SNoEzna3dRN2" executionInfo={"status": "ok", "timestamp": 1629714903985, "user_tz": -330, "elapsed": 824, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0Ix_OxEFfMvg474o7FEouUO3wMGLVoXqHui807w=s64", "userId": "01938360022046049253"}}
#import the necessary module
from sklearn.model_selection import train_test_split
#split data set into train and test sets
data_train, data_test, target_train, target_test = train_test_split(data.data,data.target, test_size = 0.40, random_state = 69)
# + colab={"base_uri": "https://localhost:8080/"} id="CRct6OrjdROI" executionInfo={"status": "ok", "timestamp": 1629714906568, "user_tz": -330, "elapsed": 548, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0Ix_OxEFfMvg474o7FEouUO3wMGLVoXqHui807w=s64", "userId": "01938360022046049253"}} outputId="ea6c759d-1f2b-459b-f7b7-4bd93dd8c866"
#Create a Decision Tree Classifier (using Gini)
cli=DecisionTreeClassifier(criterion='gini',max_leaf_nodes=100)
cli.fit(data_train,target_train)
#Train the model using the training sets
# + colab={"base_uri": "https://localhost:8080/"} id="EUmpqOTcdROY" executionInfo={"status": "ok", "timestamp": 1629714910638, "user_tz": -330, "elapsed": 759, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0Ix_OxEFfMvg474o7FEouUO3wMGLVoXqHui807w=s64", "userId": "01938360022046049253"}} outputId="da966e71-b576-467d-e504-a2b929319c22"
# Predict the classes of test data
prediction=cli.predict(data_test)
#print(test_pred.dtype)
prediction.dtype
# + colab={"base_uri": "https://localhost:8080/"} id="ygJOfDiAdROs" executionInfo={"status": "ok", "timestamp": 1629714914128, "user_tz": -330, "elapsed": 433, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0Ix_OxEFfMvg474o7FEouUO3wMGLVoXqHui807w=s64", "userId": "01938360022046049253"}} outputId="559ad0c1-41c9-4f34-9b6a-04bf35f42cf4"
from sklearn import metrics
# Model Accuracy, how often is the classifier correct?
print("Accuracy :",metrics.accuracy_score(target_test,prediction))
# + colab={"base_uri": "https://localhost:8080/"} id="n2qEF6RvTFGi" executionInfo={"status": "ok", "timestamp": 1629714917063, "user_tz": -330, "elapsed": 593, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0Ix_OxEFfMvg474o7FEouUO3wMGLVoXqHui807w=s64", "userId": "01938360022046049253"}} outputId="c80c60d0-c7d7-4dea-f451-8c58f6a866b7"
from sklearn.metrics import precision_score
from sklearn.metrics import recall_score
precision = precision_score(target_test, prediction,average=None)
recall = recall_score(target_test, prediction,average=None)
print('precision: \n {}'.format(precision))
print('\n')
print('recall: \n {}'.format(recall))
|
LAB-4/069_04_02.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: isa-api-py38
# language: python
# name: isa-api-py38
# ---
# #
# ## Abstract:
#
# In this notebook, we'll show how to generate an ISA-Tab and an ISA JSON representation of a metabolomics study.
# The study uses GC-MS and 13C NMR on 3 distinct sample types (liver, blood and heart) collected from study subjects assigned to 3 distinct study arms.
#
# GC-MS acquisition were carried out in duplicate, extracts were derivatized using BSA and acquired on an Agilent QTOF in both positive and negative modes.
# 13C NMR free induction decays were acquired on a Bruker Avance, using CPMG and PSEQ pulse sequences in duplicates.
#
#
# ### 1. Loading ISA-API model and relevant library
# +
from isatools.model import *
from isatools.create.model import *
from isatools import isatab
from isatools import isajson
from isatools.isajson import ISAJSONEncoder
from isatools.model import (Investigation, Study, Assay, Person, Material,
DataFile, plink,
OntologySource, OntologyAnnotation, Sample,
Source, Characteristic, Protocol, Process)
from isatools.isatab import dump_tables_to_dataframes as dumpdf
import networkx as nx
import numpy as np
import os
# -
# ### 2. Setting variables:
# +
NAME = 'name'
FACTORS_0_VALUE = OntologyAnnotation(term='nitroglycerin')
FACTORS_0_VALUE_ALT = OntologyAnnotation(term='alcohol')
FACTORS_0_VALUE_THIRD = OntologyAnnotation(term='water')
FACTORS_1_VALUE = 5
FACTORS_1_UNIT = OntologyAnnotation(term='kg/m^3')
FACTORS_2_VALUE = 100.0
FACTORS_2_VALUE_ALT = 50.0
FACTORS_2_UNIT = OntologyAnnotation(term='s')
TEST_EPOCH_0_NAME = 'test epoch 0'
TEST_EPOCH_1_NAME = 'test epoch 1'
TEST_EPOCH_2_NAME = 'test epoch 2'
TEST_STUDY_ARM_NAME_00 = 'test arm'
TEST_STUDY_ARM_NAME_01 = 'another arm'
TEST_STUDY_ARM_NAME_02 = 'yet another arm'
TEST_STUDY_DESIGN_NAME = 'test study design'
TEST_EPOCH_0_RANK = 0
SCREEN_DURATION_VALUE = 100
FOLLOW_UP_DURATION_VALUE = 5*366
WASHOUT_DURATION_VALUE = 30
DURATION_UNIT = OntologyAnnotation(term='day')
# -
# ### 3. Declaration of ISA Sample / Biomaterial templates for liver, blood and heart
sample_list = [
{
'node_type': SAMPLE,
'characteristics_category': OntologyAnnotation(term='organism part'),
'characteristics_value': OntologyAnnotation(term='liver'),
'size': 1,
'technical_replicates': None,
'is_input_to_next_protocols': True
},
{
'node_type': SAMPLE,
'characteristics_category': OntologyAnnotation(term='organism part'),
'characteristics_value': OntologyAnnotation(term='blood'),
'size': 1,
'technical_replicates': None,
'is_input_to_next_protocols': True
},
{
'node_type': SAMPLE,
'characteristics_category': OntologyAnnotation(term='organism part'),
'characteristics_value': OntologyAnnotation(term='heart'),
'size': 1,
'technical_replicates': None,
'is_input_to_next_protocols': True
}
]
# ### 4. Declaration of ISA Assay templates as Python `OrderedDict`
# +
# A Mass Spectrometry based metabolite profiling assay
ms_assay_dict = OrderedDict([
('measurement_type', OntologyAnnotation(term='metabolite profiling')),
('technology_type', OntologyAnnotation(term='mass spectrometry')),
('extraction', {}),
('extract', [
{
'node_type': EXTRACT,
'characteristics_category': OntologyAnnotation(term='extract type'),
'characteristics_value': OntologyAnnotation(term='polar fraction'),
'size': 1,
'is_input_to_next_protocols': True
},
{
'node_type': EXTRACT,
'characteristics_category': OntologyAnnotation(term='extract type'),
'characteristics_value': OntologyAnnotation(term='lipids'),
'size': 1,
'is_input_to_next_protocols': True
}
]),
('derivatization', {
'#replicates': 1,
OntologyAnnotation(term='derivatization'): ['sylalation'],
OntologyAnnotation(term='derivatization'): ['bis(trimethylsilyl)acetamide'],
}),
('labeled extract', [
{
'node_type': LABELED_EXTRACT,
'characteristics_category': OntologyAnnotation(term='labeled extract type'),
'characteristics_value': '',
'size': 1,
'is_input_to_next_protocols': True
}
]),
('mass spectrometry', {
'#replicates': 2,
OntologyAnnotation(term='instrument'): ['Agilent QTOF'],
OntologyAnnotation(term='injection_mode'): ['GC'],
OntologyAnnotation(term='acquisition_mode'): ['positive mode','negative mode']
}),
('raw spectral data file', [
{
'node_type': DATA_FILE,
'size': 1,
'is_input_to_next_protocols': False
}
])
])
# A high-throughput phenotyping imaging based phenotyping assay
phti_assay_dict = OrderedDict([
('measurement_type', OntologyAnnotation(term='phenotyping')),
('technology_type', OntologyAnnotation(term='high-throughput imaging')),
('extraction', {}),
('extract', [
{
'node_type': EXTRACT,
'characteristics_category': OntologyAnnotation(term='extract type'),
'characteristics_value': OntologyAnnotation(term='supernatant'),
'size': 1,
'technical_replicates': None,
'is_input_to_next_protocols': True
},
{
'node_type': EXTRACT,
'characteristics_category': OntologyAnnotation(term='extract type'),
'characteristics_value': OntologyAnnotation(term='pellet'),
'size': 1,
'technical_replicates': None,
'is_input_to_next_protocols': True
}
]),
('phenotyping by high throughput imaging', {
'OntologyAnnotation(term=instrument)': ['lemnatech gigant'],
'OntologyAnnotation(term=acquisition_mode)': ['UV light','near-IR light','far-IR light','visible light'],
'OntologyAnnotation(term=camera position)': ['top','120 degree','240 degree','360 degree'],
'OntologyAnnotation(term=imaging daily schedule)': ['06.00','19.00']
}),
('raw_spectral_data_file', [
{
'node_type': DATA_FILE,
'size': 1,
'technical_replicates': 2,
'is_input_to_next_protocols': False
}
])
])
# A liquid chromatography diode-array based metabolite profiling assay
lcdad_assay_dict = OrderedDict([
('measurement_type', OntologyAnnotation(term='metabolite identification')),
('technology_type', OntologyAnnotation(term='liquid chromatography diode-array detector')),
('extraction', {}),
('extract', [
{
'node_type': EXTRACT,
'characteristics_category': OntologyAnnotation(term='extract type'),
'characteristics_value': OntologyAnnotation(term='supernatant'),
'size': 1,
'technical_replicates': None,
'is_input_to_next_protocols': True
},
{
'node_type': EXTRACT,
'characteristics_category': OntologyAnnotation(term='extract type'),
'characteristics_value': OntologyAnnotation(term='pellet'),
'size': 1,
'technical_replicates': None,
'is_input_to_next_protocols': True
}
]),
('lcdad_spectroscopy', {
'OntologyAnnotation(term=instrument)': ['Shimadzu DAD 400'],
}),
('raw_spectral_data_file', [
{
'node_type': DATA_FILE,
'size': 1,
'technical_replicates': 2,
'is_input_to_next_protocols': False
}
])
])
# A NMR spectroscopy based metabolite profiling assay:
nmr_assay_dict = OrderedDict([
('measurement_type', OntologyAnnotation(term='metabolite profiling')),
('technology_type', OntologyAnnotation(term='nmr spectroscopy')),
('extraction', {}),
('extract', [
{
'node_type': EXTRACT,
'characteristics_category': OntologyAnnotation(term='extract type'),
'characteristics_value': OntologyAnnotation(term='supernatant'),
'size': 1,
'technical_replicates': None,
'is_input_to_next_protocols': True
},
{
'node_type': EXTRACT,
'characteristics_category': OntologyAnnotation(term='extract type'),
'characteristics_value': OntologyAnnotation(term='pellet'),
'size': 1,
'technical_replicates': None,
'is_input_to_next_protocols': True
}
]),
('nmr spectroscopy', {
OntologyAnnotation(term='instrument'): [OntologyAnnotation(term='Bruker AvanceII 1 GHz')],
OntologyAnnotation(term='acquisition_mode'): [OntologyAnnotation(term='1D 13C NMR')],
OntologyAnnotation(term='pulse_sequence'): [OntologyAnnotation(term='CPMG')]
}),
('raw_spectral_data_file', [
{
'node_type': DATA_FILE,
'size': 1,
'technical_replicates': 1,
'is_input_to_next_protocols': False
}
])
])
# +
first_treatment = Treatment(factor_values=(
FactorValue(factor_name=BASE_FACTORS[0], value=FACTORS_0_VALUE),
FactorValue(factor_name=BASE_FACTORS[1], value=FACTORS_1_VALUE, unit=FACTORS_1_UNIT),
FactorValue(factor_name=BASE_FACTORS[2], value=FACTORS_2_VALUE, unit=FACTORS_2_UNIT)
))
second_treatment = Treatment(factor_values=(
FactorValue(factor_name=BASE_FACTORS[0], value=FACTORS_0_VALUE_ALT),
FactorValue(factor_name=BASE_FACTORS[1], value=FACTORS_1_VALUE, unit=FACTORS_1_UNIT),
FactorValue(factor_name=BASE_FACTORS[2], value=FACTORS_2_VALUE, unit=FACTORS_2_UNIT)
))
third_treatment = Treatment(factor_values=(
FactorValue(factor_name=BASE_FACTORS[0], value=FACTORS_0_VALUE_ALT),
FactorValue(factor_name=BASE_FACTORS[1], value=FACTORS_1_VALUE, unit=FACTORS_1_UNIT),
FactorValue(factor_name=BASE_FACTORS[2], value=FACTORS_2_VALUE_ALT, unit=FACTORS_2_UNIT)
))
fourth_treatment = Treatment(factor_values=(
FactorValue(factor_name=BASE_FACTORS[0], value=FACTORS_0_VALUE_THIRD),
FactorValue(factor_name=BASE_FACTORS[1], value=FACTORS_1_VALUE, unit=FACTORS_1_UNIT),
FactorValue(factor_name=BASE_FACTORS[2], value=FACTORS_2_VALUE, unit=FACTORS_2_UNIT)
))
screen = NonTreatment(element_type=SCREEN, duration_value=SCREEN_DURATION_VALUE, duration_unit=DURATION_UNIT)
run_in = NonTreatment(element_type=RUN_IN, duration_value=WASHOUT_DURATION_VALUE, duration_unit=DURATION_UNIT)
washout = NonTreatment(element_type=WASHOUT, duration_value=WASHOUT_DURATION_VALUE, duration_unit=DURATION_UNIT)
follow_up = NonTreatment(element_type=FOLLOW_UP, duration_value=FOLLOW_UP_DURATION_VALUE, duration_unit=DURATION_UNIT)
potential_concomitant_washout = NonTreatment(element_type=WASHOUT, duration_value=FACTORS_2_VALUE,
duration_unit=FACTORS_2_UNIT)
cell_screen = StudyCell(SCREEN, elements=(screen,))
cell_run_in = StudyCell(RUN_IN, elements=(run_in,))
cell_other_run_in = StudyCell('OTHER RUN-IN', elements=(run_in,))
cell_screen_and_run_in = StudyCell('SCREEN AND RUN-IN', elements=[screen, run_in])
cell_concomitant_treatments = StudyCell('CONCOMITANT TREATMENTS',
elements=([{second_treatment, fourth_treatment}]))
cell_washout_00 = StudyCell(WASHOUT, elements=(washout,))
cell_washout_01 = StudyCell('ANOTHER WASHOUT', elements=(washout,))
cell_single_treatment_00 = StudyCell('SINGLE TREATMENT FIRST', elements=[first_treatment])
cell_single_treatment_01 = StudyCell('SINGLE TREATMENT SECOND', elements=[second_treatment])
cell_single_treatment_02 = StudyCell('SINGLE TREATMENT THIRD', elements=[third_treatment])
cell_multi_elements = StudyCell('MULTI ELEMENTS',
elements=[{first_treatment, second_treatment,
fourth_treatment}, washout, second_treatment])
cell_multi_elements_padded = StudyCell('MULTI ELEMENTS PADDED',
elements=[first_treatment, washout, {
second_treatment,
fourth_treatment
}, washout, third_treatment, washout])
cell_follow_up = StudyCell(FOLLOW_UP, elements=(follow_up,))
cell_follow_up_01 = StudyCell('ANOTHER FOLLOW_UP', elements=(follow_up,))
qc = QualityControl()
ms_sample_assay_plan = SampleAndAssayPlan.from_sample_and_assay_plan_dict("ms_sap", sample_list, ms_assay_dict)
nmr_sample_assay_plan = SampleAndAssayPlan.from_sample_and_assay_plan_dict("nmr_sap", sample_list, nmr_assay_dict)
first_arm = StudyArm(name=TEST_STUDY_ARM_NAME_00, group_size=3, arm_map=OrderedDict([
(cell_screen, None), (cell_run_in, None),
(cell_single_treatment_00, ms_sample_assay_plan),
(cell_follow_up, ms_sample_assay_plan)
]))
second_arm = StudyArm(name=TEST_STUDY_ARM_NAME_01, group_size=5, arm_map=OrderedDict([
(cell_screen, None), (cell_run_in, None),
(cell_multi_elements, ms_sample_assay_plan),
(cell_follow_up, ms_sample_assay_plan)
]))
third_arm = StudyArm(name=TEST_STUDY_ARM_NAME_02, group_size=3, arm_map=OrderedDict([
(cell_screen, None), (cell_run_in, None),
(cell_multi_elements_padded, ms_sample_assay_plan),
(cell_follow_up, ms_sample_assay_plan)
]))
third_arm_no_run_in = StudyArm(name=TEST_STUDY_ARM_NAME_02, group_size=3, arm_map=OrderedDict([
(cell_screen, None),
(cell_multi_elements_padded, ms_sample_assay_plan),
(cell_follow_up, ms_sample_assay_plan)
]))
arm_same_name_as_third = StudyArm(name=TEST_STUDY_ARM_NAME_02, group_size=5, arm_map=OrderedDict([
(cell_screen, None), (cell_run_in, None),
(cell_single_treatment_01, ms_sample_assay_plan),
(cell_follow_up, ms_sample_assay_plan)
]))
# Sample QC (for mass spectroscopy and other)
pre_run_sample_type = ProductNode(
id_='pre/00', node_type=SAMPLE, name='water', size=2, characteristics=(
Characteristic(category='dilution', value=10, unit='mg/L'),
)
)
post_run_sample_type = ProductNode(
id_='post/00', node_type=SAMPLE, name='ethanol', size=2, characteristics=(
Characteristic(category='dilution', value=1000, unit='mg/L'),
Characteristic(category='dilution', value=100, unit='mg/L'),
Characteristic(category='dilution', value=10, unit='mg/L'),
Characteristic(category='dilution', value=1, unit='mg/L'),
Characteristic(category='dilution', value=0.1, unit='mg/L')
))
dummy_sample_type = ProductNode(id_='dummy/01', node_type=SAMPLE, name='dummy')
more_dummy_sample_type = ProductNode(id_='dummy/02', node_type=SAMPLE, name='more dummy')
interspersed_sample_types = [(dummy_sample_type, 20)]
qc = QualityControl(
interspersed_sample_type=interspersed_sample_types,
pre_run_sample_type=pre_run_sample_type,
post_run_sample_type=post_run_sample_type
)
# -
single_arm = StudyArm(name=TEST_STUDY_ARM_NAME_00, group_size=10, arm_map=OrderedDict([
(cell_screen, ms_sample_assay_plan), (cell_run_in,ms_sample_assay_plan),
(cell_single_treatment_00, nmr_sample_assay_plan),
(cell_follow_up, nmr_sample_assay_plan)
]))
study_design = StudyDesign(study_arms=(single_arm,))
study = study_design.generate_isa_study()
study
treatment_assay = next(iter(study.assays))
treatment_assay.graph
[(process.name, getattr(process.prev_process, 'name', None), getattr(process.next_process, 'name', None)) for process in treatment_assay.process_sequence]
a_graph = treatment_assay.graph
len(a_graph.nodes)
isa_investigation = Investigation(studies=[study])
isa_tables = dumpdf(isa_investigation)
[type(x) for x in study.assays[0].graph.nodes()]
[(getattr(el, 'name', None), type(el))for el in treatment_assay.graph.nodes()]
from isatools.model import _build_assay_graph
gph = _build_assay_graph(treatment_assay.process_sequence)
[key for key in isa_tables.keys()]
isa_tables['s_study_01.txt']
isa_tables['a_AT0_metabolite-profiling_nmr-spectroscopy.txt']
isa_tables['a_AT0_metabolite-profiling_mass-spectrometry.txt']
final_dir = os.path.abspath(os.path.join('notebook-output', 'sd-test'))
isa_j = json.dumps(isa_investigation, cls=ISAJSONEncoder, sort_keys=True, indent=4, separators=(',', ': '))
open(os.path.join(final_dir,"isa_as_json_from_dumps2.json"),"w").write(isa_j) # this call write the string 'isa_j' to the file called 'isa_as_json_from_dumps.json'
isatab.dump(isa_obj=isa_investigation, output_path=final_dir)
print(final_dir)
with open(os.path.join(final_dir,'i_investigation.txt')) as isa:
validation_report=isatab.validate(isa)
validation_report["errors"]
# ## Conclusion:
#
#
# ## About this notebook
#
# - authors: <EMAIL>, <EMAIL>
# - license: CC-BY 4.0
# - support: <EMAIL>
# - issue tracker: https://github.com/ISA-tools/isa-api/issues
|
_build/jupyter_execute/content/notebooks/isa-metabolomics-study-design-create.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Format GEOJSON
# Dernier format de données que nous aborderons, le **geoJSON**.
#
# Le geoJSON est un format (<http://geojson.org/>) qui permet d'encoder des
# données à "**caractère géographique**". Voici ce que dit Wikipédia à propos
# de ce format :
#
# > GeoJSON (de l'anglais Geographic JSON, signifiant littéralement JSON
# > géographique) est un format ouvert d'encodage d'ensemble de données
# > géospatiales simples utilisant la norme JSON (JavaScript Object
# > Notation). Il permet de décrire des données de type point, ligne,
# > chaîne de caractères, polygone, ainsi que des ensembles et
# > sous-ensembles de ces types de données et d'y ajouter des attributs
# > d'information qui ne sont pas spatiale. Le format GeoJSON,
# > contrairement à la majorité des standards de systèmes d'informations
# > géographiques, n'est pas écrit par l'Open Geospatial Consortium, mais
# > par un groupe de travail de développeurs au travers d'internet.
#
# Comme indiqué dans Wikipédia, le geoJSON est avant tout du JSON, nous
# retrouverons donc les mêmes caractéristiques que le JSON (système de
# clé/valeur).
# ### À faire vous-même
#
# Dans la barre d'adresse de votre navigateur, tapez l'adresse suivante
# :
#
# ```
# https://earthquake.usgs.gov/fdsnws/event/1/query?format=geojson&starttime=2019-07-31&endtime=2019-08-01
#
# ```
#
# Vous devriez obtenir quelque chose ressemblant à ceci
# :
# ```json
# {
# "type": "FeatureCollection",
# "metadata": {
# "generated": 1567103163000,
# "url": "https://earthquake.usgs.gov/fdsnws/event/1/query?format=geojson&starttime=2019-07-31&endtime=2019-08-01",
# "title": "USGS Earthquakes",
# "status": 200,
# "api": "1.8.1",
# "count": 635
# },
# "features": [
# {
# "type": "Feature",
# "properties": {
# "mag": 0.33,
# "place": "6km NW of The Geysers, CA",
# "time": 1564617341320,
# "updated": 1564619343353,
# "tz": -480,
# "url": "https://earthquake.usgs.gov/earthquakes/eventpage/nc73239641",
# "detail": "https://earthquake.usgs.gov/fdsnws/event/1/query?eventid=nc73239641&format=geojson",
# "felt": null,
# "cdi": null,
# "mmi": null,
# "alert": null,
# "status": "automatic",
# "tsunami": 0,
# "sig": 2,
# "net": "nc",
# "code": "73239641",
# "ids": ",nc73239641,",
# "sources": ",nc,",
# "types": ",geoserve,nearby-cities,origin,phase-data,scitech-link,",
# "nst": 7,
# "dmin": 0.01258,
# "rms": 0.03,
# "gap": 78,
# "magType": "md",
# "type": "earthquake",
# "title": "M 0.3 - 6km NW of The Geysers, CA"
# },
# "geometry": {
# "type": "Point",
# "coordinates": [
# -122.8026657,
# 38.8191681,
# 3.29
# ]
# },
# "id": "nc73239641"
# },
# {
# "type": "Feature",
# ...
# ```
#
# -----
# Vous avez obtenu des informations au format geoJSON, sur les
# tremblements de Terre ayant eu lieu entre le 31 juillet 2019 et le 01
# août 2019, partout dans le monde.
#
# Le site "earthquake.usgs.gov", comme le site
# "http://openweathermap.org/", propose une API qui renvoie des données à
# partir d'une simple url. Le site vous propose différentes options pour
# la requête, vous trouverez une description complète de ces options
# [ici](http://earthquake.usgs.gov/fdsnws/event/1/)
#
# Attention, vous aurez un message d'erreur si votre requête renvoie plus
# de 20000 événements
# ### À faire vous-même
#
# En vous aidant de la documentation présente sur le site
# [http://earthquake.usgs.gov](http://earthquake.usgs.gov/fdsnws/event/1/),
# écrivez une requête sous forme d'url qui permettra d'obtenir des données
# (au format geoJSON) sur les tremblements de terre, d'une magnitude
# supérieure à 5, ayant eu lieu ces 30 derniers jours partout dans le
# monde.
#
# Testez votre requête en la copiant dans la barre d'adresse de votre
# navigateur. Une fois les données obtenues, étudiez-les afin de
# comprendre la structure de ces données.
# **ATTENTION** : les dates et les heures sont fournies au format "timestamp".
# Le "timestamp" désigne le nombre de secondes écoulé depuis le 1er
# janvier 1970 à minuit UTC précise. Au lieu de donner une date et une
# heure pour un événement donné, il est possible de donner son
# "timestamp". Par exemple, au lieu de dire l'événement A à eu lieu le 24
# octobre 2018 à 13h 11 minutes et 10 secondes, on pourra dire que
# l'événement A à pour "timestamp" 1540379470 (durée qui s'est écoulé en
# seconde entre le 1er janvier 1970 à minuit UTC et le 24 octobre 2018 à
# 13h 11 minutes et 10 secondes). Vous trouverez un convertisseur de
# timestamp sur ce [site](http://www.timestamp.fr/?). Attention, dans le
# JSON renvoyé par le site "earthquake.usgs.gov" le timestamp est donné en
# milliseconde, il est donc nécessaire de diviser par 1000 la valeur que
# vous allez trouver dans le JSON (et garder uniquement la partie entière
# du résultat de votre division).
#
# -----
|
1_les_donnees_structurees_et_leur_traitement/activites/02_Le_format_GEOJSON.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Region Based Data Analysis
# The following notebook will go through prediction analysis for region based Multiple Particle Tracking (MPT) using OGD severity datasets for non-treated (NT) hippocampus, ganglia, thalamus, cortex, and striatum.
#
# ## Table of Contents
#
#
# [1. Load Data](#1.-load-data)<br />
# [2. Analysis](#2.-analysis)<br />
# [3. Modelling](#modelling)<br />
# [4. Evaluate Results](#evaluate-results)<br />
# ---
# ## 1. Load Data
# Loading feature dataset from OGD folders:
#
# There are 15 total videos from each age group.
# +
# libraries used
import boto3
import diff_classifier.aws as aws
import pandas as pd
import seaborn as sn
import numpy as np
import matplotlib.pyplot as pl
from os import listdir, getcwd, chdir
from os.path import isfile, join
import os
from matplotlib import colors as plt_colors
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
from sklearn import metrics
from sklearn.metrics import classification_report
from sklearn.metrics import confusion_matrix
from sklearn import preprocessing
import xgboost as xgb
# from xgboost import cv
import shap
# -
workbookDir = getcwd()
print('Current Notebook Dir: ' + workbookDir)
chdir(workbookDir) # Go to current workbook Dir
chdir('..') # Go up one
print(f'Using current directory for loading data: {getcwd()}')
workbookDir = getcwd()
# !pwd
dataset_path = workbookDir + '/region_feature_folder/'
filelist = [f for f in listdir(dataset_path) if isfile(join(dataset_path, f)) and 'feat' in f and 'ganglia' not in f and 'hippocampus' not in f and 'thalamus' not in f]
filelist
fstats_tot = None
video_num = 0
for filename in filelist:
# try:
fstats = pd.read_csv(dataset_path + filename, encoding = "ISO-8859-1", index_col='Unnamed: 0')
print('{} size: {}'.format(filename, fstats.shape))
if 'cortex' in filename:
fstats['region'] = pd.Series(fstats.shape[0]*['cortex'], index=fstats.index)
elif 'striatum' in filename:
fstats['region'] = pd.Series(fstats.shape[0]*['striatum'], index=fstats.index)
elif 'ganglia' in filename:
fstats['region'] = pd.Series(fstats.shape[0]*['ganglia'], index=fstats.index)
elif 'thalamus' in filename:
fstats['region'] = pd.Series(fstats.shape[0]*['thalamus'], index=fstats.index)
elif 'hippocampus' in filename:
fstats['region'] = pd.Series(fstats.shape[0]*['hippocampus'], index=fstats.index)
else:
print('Error, no target')
fstats['Video Number'] = pd.Series(fstats.shape[0]*[video_num], index=fstats.index)
if fstats_tot is None:
fstats_tot = fstats
else:
fstats_tot = fstats_tot.append(fstats, ignore_index=True)
video_num += 1
# except Exception:
# print('Skipped!: {}'.format(filename))
# ## 2. Analysis
# The following columns are present within the downloaded datasets:
fstats_tot.columns
# Many of these features are not useful for prediction or have data which may negatively impact classification. The following features and the target feature are defined in the following cell. We also remove any datapoints that are empty or infinite:
# +
fstats_tot
features = [
'alpha', # Fitted anomalous diffusion alpha exponenet
'D_fit', # Fitted anomalous diffusion coefficient
'kurtosis', # Kurtosis of track
'asymmetry1', # Asymmetry of trajecory (0 for circular symmetric, 1 for linear)
'asymmetry2', # Ratio of the smaller to larger principal radius of gyration
'asymmetry3', # An asymmetric feature that accnts for non-cylindrically symmetric pt distributions
'AR', # Aspect ratio of long and short side of trajectory's minimum bounding rectangle
'elongation', # Est. of amount of extension of trajectory from centroid
'boundedness', # How much a particle with Deff is restricted by a circular confinement of radius r
'fractal_dim', # Measure of how complicated a self similar figure is
'trappedness', # Probability that a particle with Deff is trapped in a region
'efficiency', # Ratio of squared net displacement to the sum of squared step lengths
'straightness', # Ratio of net displacement to the sum of squared step lengths
'MSD_ratio', # MSD ratio of the track
'frames', # Number of frames the track spans
'Deff1', # Effective diffusion coefficient at 0.33 s
'Deff2', # Effective diffusion coefficient at 3.3 s
# 'angle_mean', # Mean turning angle which is counterclockwise angle from one frame point to another
# 'angle_mag_mean', # Magnitude of the turning angle mean
# 'angle_var', # Variance of the turning angle
# 'dist_tot', # Total distance of the trajectory
# 'dist_net', # Net distance from first point to last point
# 'progression', # Ratio of the net distance traveled and the total distance
'Mean alpha',
'Mean D_fit',
'Mean kurtosis',
'Mean asymmetry1',
'Mean asymmetry2',
'Mean asymmetry3',
'Mean AR',
'Mean elongation',
'Mean boundedness',
'Mean fractal_dim',
'Mean trappedness',
'Mean efficiency',
'Mean straightness',
'Mean MSD_ratio',
'Mean Deff1',
'Mean Deff2',
]
target = 'region' # prediction target (y)
ecm = fstats_tot[features + [target] + ['X'] + ['Y']]
ecm = ecm[~ecm.isin([np.nan, np.inf, -np.inf]).any(1)] # Removing nan and inf data points
# -
# Showing a piece of our data:
ecm[target].unique()
# Before prediction, it is required to balance data. As shown, The current dataset is highly imbalance with most datapoints belonging to P21 and P35 categories. The dataset is reduced using random sampling of each target category.
#--------------NOT-ADDED-----------------------------
def balance_data(df, target, **kwargs):
if 'random_state' not in kwargs:
random_state = 1
else:
random_state = kwargs['random_state']
if isinstance(target, list):
target = target[0]
df_target = []
bal_df = []
for name in df[target].unique():
df_target.append((name, df[df[target] == name]))
print(f"Ratio before data balance ({':'.join([str(i[0]) for i in df_target])}) = {':'.join([str(len(i[1])) for i in df_target])}")
for i in range(len(df_target)):
ratio = min([len(i[1]) for i in df_target])/len(df_target[i][1])
bal_df.append(df_target[i][1].sample(frac=ratio, random_state=random_state))
print(f"Ratio after balance ({':'.join([str(i[0]) for i in df_target])}) = {':'.join([str(len(i)) for i in bal_df])}")
return pd.concat(bal_df)
bal_ecm = balance_data(ecm, target, random_state=1)
# +
# ecm_14 = ecm[ecm[target] == 14]
# ecm_21 = ecm[ecm[target] == 21]
# ecm_28 = ecm[ecm[target] == 28]
# ecm_35 = ecm[ecm[target] == 35]
# print(f"Ratio before data balance (P14:P21:P28:P35) = {len(ecm_14)}:{len(ecm_21)}:{len(ecm_28)}:{len(ecm_35)}")
# ecm_list = [ecm_14, ecm_21, ecm_28, ecm_35]
# for i in range(len(ecm_list)):
# ratio = min([len(i) for i in ecm_list])/len(ecm_list[i])
# ecm_list[i] = ecm_list[i].sample(frac=ratio, random_state=1)
# print(f"Ratio after balance (P14:P21:P28:P35) = {len(ecm_list[0])}:{len(ecm_list[1])}:{len(ecm_list[2])}:{len(ecm_list[3])}")
# bal_ecm = pd.concat(ecm_list)
# -
# ## 3. Modelling
# The model used for this study is an extreme gradient boosting (XGBoost) decision tree which is a boosted decision tree. This model was used due to its past results within competitions and research.
# Due to the use of statistical surroundings in our feature analysis, binning is required in order to avoid data leakage between training/testing. The followingcode will implement binning and a checkerboard implementation to select certain bins for the training dataset.
# Using checkerboard binning for data split:
def checkerboard(size):
rows = int(size/2)
checks = list(range(0, size*size, size+1))
for i in range(1, rows):
ssize = size - 2*i
for j in range(0, ssize):
checks.append(2*i + (size+1)*j)
for i in range(1, rows):
ssize = size - 2*i
for j in range(0, ssize):
checks.append(size*size - 1 - (2*i + (size+1)*j))
checks.sort()
return checks
# +
# Old method
# bins = list(range(0, 2048+1, 256))
# bal_ecm['binx'] = pd.cut(bal_ecm.X, bins, labels=[0, 1, 2, 3, 4, 5, 6, 7], include_lowest=True)
# bal_ecm['biny'] = pd.cut(bal_ecm.Y, bins, labels=[0, 1, 2, 3, 4, 5, 6, 7], include_lowest=True)
# bal_ecm['bins'] = 8*bal_ecm['binx'].astype(np.int8) + bal_ecm['biny'].astype(np.int8)
# bal_ecm = bal_ecm[np.isfinite(bal_ecm['bins'])]
# bal_ecm['bins'] = bal_ecm['bins'].astype(int)
# cols = bal_ecm.columns.tolist()
# cols = cols[-3:] + cols[:-3]
# bal_ecm = bal_ecm[cols]
# +
# def bin_data(data, ):
# pass
resolution = 128
assert not 2048%resolution and resolution >= 128, "resolution needs to be a factor of 2048 and > 128"
bins = list(range(0, 2048+1, resolution))
bin_labels = [int(i/resolution) for i in bins][:-1]
bal_ecm['binx'] = pd.cut(bal_ecm.X, bins, labels=bin_labels, include_lowest=True)
bal_ecm['biny'] = pd.cut(bal_ecm.Y, bins, labels=bin_labels, include_lowest=True)
bal_ecm['bins'] = (len(bins)-1)*bal_ecm['binx'].astype(np.int32) + bal_ecm['biny'].astype(np.int32)
bal_ecm = bal_ecm[np.isfinite(bal_ecm['bins'])]
bal_ecm['bins'] = bal_ecm['bins'].astype(int)
# cols = bal_ecm.columns.tolist()
# cols = cols[-3:] + cols[:-3]
# bal_ecm = bal_ecm[cols]
# +
# Checkerboard method
# seed = 1234
# np.random.seed(seed)
# test_val_split = 0.5
# le = preprocessing.LabelEncoder()
# bal_ecm['encoded_target'] = le.fit_transform(bal_ecm[target])
# X_train = bal_ecm[~bal_ecm.bins.isin(checkerboard((len(bins)-1)))].reset_index()
# X_test_val = bal_ecm[bal_ecm.bins.isin(checkerboard((len(bins)-1)))].reset_index()
# y_train = X_train['encoded_target']
# X_val, X_test = train_test_split(X_test_val, test_size=test_val_split, random_state=seed)
# y_test = X_test['encoded_target']
# y_val = X_val['encoded_target']
# dtrain = xgb.DMatrix(X_train[features], label=y_train)
# dtest = xgb.DMatrix(X_test[features], label=y_test)
# dval = xgb.DMatrix(X_val[features], label=y_val)
# +
# Regular split
seed = 1234
np.random.seed(seed)
train_split = 0.8
test_split = 0.5
le = preprocessing.LabelEncoder()
bal_ecm['encoded_target'] = le.fit_transform(bal_ecm[target])
training_bins = np.random.choice(bal_ecm.bins.unique(), int(len(bal_ecm.bins.unique())*train_split), replace=False)
X_train = bal_ecm[bal_ecm.bins.isin(training_bins)]
X_test_val = bal_ecm[~bal_ecm.bins.isin(training_bins)]
X_val, X_test = train_test_split(X_test_val, test_size=test_split, random_state=seed)
y_train = X_train['encoded_target']
y_test = X_test['encoded_target']
y_val = X_val['encoded_target']
dtrain = xgb.DMatrix(X_train[features], label=y_train)
dtest = xgb.DMatrix(X_test[features], label=y_test)
dval = xgb.DMatrix(X_val[features], label=y_val)
# -
#Check lengths of datasets:
def get_lengths(df, X_train, X_test, X_val=None):
print(f'Tot before split: {len(df)}')
print(f'Training: {len(X_train)} ({len(X_train)/len(bal_ecm):.3f}%)')
print(f'Testing: {len(X_test)} ({len(X_test)/len(bal_ecm):.3f}%)')
try:
print(f'Evaluation: {len(X_val)} ({len(X_val)/len(bal_ecm):.3f}%)')
except:
pass
get_lengths(bal_ecm, X_train, X_test, X_val)
from xgboost.libpath import find_lib_path
import ctypes
lib_path = find_lib_path()
lib = ctypes.cdll.LoadLibrary(lib_path[0])
# Model parameters are based on the best possible XGBoost parameters to minimize logloss error.
# Init_params for binary logistic classification
init_param = {'max_depth': 3,
'eta': 0.005,
'min_child_weight': 0,
'verbosity': 0,
'objective': 'binary:logistic',
'silent': 'True',
'gamma': 5,
'subsample': 0.15,
'colsample_bytree': 0.8,
'eval_metric': 'logloss'}
# from xgboost import XGBClassifier
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score
# model = XGBClassifier()
# +
# model.predict(X_test[features])
# -
features
# +
from xgboost.training import CVPack
from xgboost import callback
from xgboost.core import CallbackEnv
from xgboost.core import EarlyStopException
def cv(params, X_train, y_train, features=None, num_boost_round=20, nfold=3, stratified=False, folds=None,
metrics=(), obj=None, feval=None, maximize=False, early_stopping_rounds=None,
fpreproc=None, as_pandas=True, verbose_eval=None, show_stdv=True,
seed=0, callbacks=None, shuffle=True):
# pylint: disable = invalid-name
"""Cross-validation with given parameters.
Parameters
----------
params : dict
Booster params.
dtrain : DMatrix
Data to be trained.
num_boost_round : int
Number of boosting iterations.
nfold : int
Number of folds in CV.
stratified : bool
Perform stratified sampling.
folds : a KFold or StratifiedKFold instance or list of fold indices
Sklearn KFolds or StratifiedKFolds object.
Alternatively may explicitly pass sample indices for each fold.
For ``n`` folds, **folds** should be a length ``n`` list of tuples.
Each tuple is ``(in,out)`` where ``in`` is a list of indices to be used
as the training samples for the ``n`` th fold and ``out`` is a list of
indices to be used as the testing samples for the ``n`` th fold.
metrics : string or list of strings
Evaluation metrics to be watched in CV.
obj : function
Custom objective function.
feval : function
Custom evaluation function.
maximize : bool
Whether to maximize feval.
early_stopping_rounds: int
Activates early stopping. Cross-Validation metric (average of validation
metric computed over CV folds) needs to improve at least once in
every **early_stopping_rounds** round(s) to continue training.
The last entry in the evaluation history will represent the best iteration.
If there's more than one metric in the **eval_metric** parameter given in
**params**, the last metric will be used for early stopping.
fpreproc : function
Preprocessing function that takes (dtrain, dtest, param) and returns
transformed versions of those.
as_pandas : bool, default True
Return pd.DataFrame when pandas is installed.
If False or pandas is not installed, return np.ndarray
verbose_eval : bool, int, or None, default None
Whether to display the progress. If None, progress will be displayed
when np.ndarray is returned. If True, progress will be displayed at
boosting stage. If an integer is given, progress will be displayed
at every given `verbose_eval` boosting stage.
show_stdv : bool, default True
Whether to display the standard deviation in progress.
Results are not affected, and always contains std.
seed : int
Seed used to generate the folds (passed to numpy.random.seed).
callbacks : list of callback functions
List of callback functions that are applied at end of each iteration.
It is possible to use predefined callbacks by using
:ref:`Callback API <callback_api>`.
Example:
.. code-block:: python
[xgb.callback.reset_learning_rate(custom_rates)]
shuffle : bool
Shuffle data before creating folds.
Returns
-------
evaluation history : list(string)
"""
if stratified is True and not SKLEARN_INSTALLED:
raise XGBoostError('sklearn needs to be installed in order to use stratified cv')
if isinstance(metrics, str):
metrics = [metrics]
if not features:
features = X_train.columns
if isinstance(params, list):
_metrics = [x[1] for x in params if x[0] == 'eval_metric']
params = dict(params)
if 'eval_metric' in params:
params['eval_metric'] = _metrics
else:
params = dict((k, v) for k, v in params.items())
if (not metrics) and 'eval_metric' in params:
if isinstance(params['eval_metric'], list):
metrics = params['eval_metric']
else:
metrics = [params['eval_metric']]
params.pop("eval_metric", None)
results = {}
# create folds in data
cvfolds, wt_list = mknfold(X_train, y_train, nfold, params, metrics, features)
# setup callbacks
callbacks = [] if callbacks is None else callbacks
if early_stopping_rounds is not None:
callbacks.append(callback.early_stop(early_stopping_rounds,
maximize=maximize,
verbose=False))
if isinstance(verbose_eval, bool) and verbose_eval:
callbacks.append(callback.print_evaluation(show_stdv=show_stdv))
elif isinstance(verbose_eval, int):
callbacks.append(callback.print_evaluation(verbose_eval, show_stdv=show_stdv))
callbacks_before_iter = [
cb for cb in callbacks if
cb.__dict__.get('before_iteration', False)]
callbacks_after_iter = [
cb for cb in callbacks if
not cb.__dict__.get('before_iteration', False)]
for i in range(num_boost_round):
for cb in callbacks_before_iter:
cb(CallbackEnv(model=None,
cvfolds=cvfolds,
iteration=i,
begin_iteration=0,
end_iteration=num_boost_round,
rank=0,
evaluation_result_list=None))
for fold in cvfolds:
fold.update(i, obj)
res = aggcv([f.eval(i, feval) for f in cvfolds], wt_list)
for key, mean, std in res:
if key + '-mean' not in results:
results[key + '-mean'] = []
if key + '-std' not in results:
results[key + '-std'] = []
results[key + '-mean'].append(mean)
results[key + '-std'].append(std)
try:
for cb in callbacks_after_iter:
cb(CallbackEnv(model=None,
cvfolds=cvfolds,
iteration=i,
begin_iteration=0,
end_iteration=num_boost_round,
rank=0,
evaluation_result_list=res))
except EarlyStopException as e:
for k in results:
results[k] = results[k][:(e.best_iteration + 1)]
break
if as_pandas:
try:
import pandas as pd
results = pd.DataFrame.from_dict(results)
except ImportError:
pass
return results
# +
def bin_fold(X_train, nfold):
bin_list = [X_train[X_train['bins'] == i_bin].index.to_numpy() for i_bin in X_train.bins.unique()]
bin_list = sorted(bin_list, key=len)
i = 0
while(len(bin_list) > nfold):
if (i >= len(bin_list)-1):
i = 0
bin_list[i] = np.concatenate([bin_list[i], bin_list.pop()])
i += 1
wt_list = [len(i)/sum(len(s) for s in bin_list) for i in bin_list]
return bin_list, wt_list
def mknfold(X_train, y_train, nfold, param, evals=(), features=None):
if not features:
features = X_train.columns
dall = xgb.DMatrix(X_train[features], label=y_train)
out_idset, wt_list = bin_fold(X_train, nfold)
in_idset = [np.concatenate([out_idset[i] for i in range(nfold) if k != i]) for k in range(nfold)]
evals = list(evals)
ret = []
for k in range(nfold):
# perform the slicing using the indexes determined by the above methods
x_train_snip = X_train.loc[in_idset[k]][features]
y_train_snip = X_train.loc[in_idset[k]]['encoded_target']
x_test_snip = X_train.loc[out_idset[k]][features]
y_test_snip = X_train.loc[out_idset[k]]['encoded_target']
dtrain = xgb.DMatrix(x_train_snip, label=y_train_snip)
dtest = xgb.DMatrix(x_test_snip, label=y_test_snip)
tparam = param
plst = list(tparam.items()) + [('eval_metric', itm) for itm in evals]
ret.append(CVPack(dtrain, dtest, plst))
return ret, wt_list
# +
from xgboost.core import STRING_TYPES
def aggcv(rlist, wt_list):
# pylint: disable=invalid-name
"""
Aggregate cross-validation results.
If verbose_eval is true, progress is displayed in every call. If
verbose_eval is an integer, progress will only be displayed every
`verbose_eval` trees, tracked via trial.
"""
cvmap = {}
idx = rlist[0].split()[0]
for line in rlist:
arr = line.split()
assert idx == arr[0]
for metric_idx, it in enumerate(arr[1:]):
if not isinstance(it, STRING_TYPES):
it = it.decode()
k, v = it.split(':')
if (metric_idx, k) not in cvmap:
cvmap[(metric_idx, k)] = []
cvmap[(metric_idx, k)].append(float(v))
msg = idx
results = []
for (metric_idx, k), v in sorted(cvmap.items(), key=lambda x: x[0][0]):
v = np.array(v)
if not isinstance(msg, STRING_TYPES):
msg = msg.decode()
mean = np.average(v, weights=wt_list)
std = np.average((v-mean)**2, weights=wt_list)
results.extend([(k, mean, std)])
return results
# -
cv(init_param, X_train, y_train, features, num_boost_round=10, nfold=5, early_stopping_rounds=3, metrics={'logloss', 'error'})
from scipy.stats import skewnorm
a=10
data = [round(i, 3) for i in skewnorm.rvs(a, size=10, random_state=seed)*0.3]
data
seed = 1234
np.random.seed(seed)
# +
import operator
import numpy as np
def xgb_paramsearch(X_train, y_train, features, init_params, nfold=5, num_boost_round=2000, early_stopping_rounds=3, metrics=None, **kwargs):
params = {**init_params}
if 'use_gpu' in kwargs and kwargs['use_gpu']:
# GPU integration will cut cv time in ~half:
params.update({'gpu_id' : 0,
'tree_method': 'gpu_hist',
'predictor': 'gpu_predictor'})
if 'metrics' not in kwargs:
metrics = {params['eval_metric']}
else:
metrics.add(params['eval_metric'])
if params['eval_metric'] in ['map', 'auc', 'aucpr']:
eval_f = operator.gt
else:
eval_f = operator.lt
if 'early_break' not in kwargs:
early_break = 5
else:
early_break = kwargs['early_break']
if 'thresh' not in kwargs:
thresh = 0.01
else:
thresh = kwargs['thresh']
if 'seed' not in kwargs:
seed = 1111
else:
seed = kwargs['seed']
best_param = params
best_model = cv(params,
X_train,
y_train,
features,
nfold=nfold,
num_boost_round=num_boost_round,
early_stopping_rounds=early_stopping_rounds,
metrics=metrics)
best_eval = best_model[f"test-{params['eval_metric']}-mean"].min()
best_boost_rounds = best_model[f"test-{params['eval_metric']}-mean"].idxmin()
def _gs_helper(var1n, var2n, best_model, best_param, best_eval, best_boost_rounds):
local_param = {**best_param}
for var1, var2 in gs_params:
print(f"Using CV with {var1n}={{{var1}}}, {var2n}={{{var2}}}")
local_param[var1n] = var1
local_param[var2n] = var2
cv_model = cv(local_param,
X_train,
y_train,
features,
nfold=nfold,
num_boost_round= num_boost_round,
early_stopping_rounds=early_stopping_rounds,
metrics=metrics)
cv_eval = cv_model[f"test-{local_param['eval_metric']}-mean"].min()
boost_rounds = cv_model[f"test-{local_param['eval_metric']}-mean"].idxmin()
if(eval_f(cv_eval, best_eval)):
best_model = cv_model
best_param[var1n] = var1
best_param[var2n] = var2
best_eval = cv_eval
best_boost_rounds = boost_rounds
print(f"New best param found: "
f"{local_param['eval_metric']} = {{{best_eval}}}, "
f"boost_rounds = {{{best_boost_rounds}}}")
return best_model, best_param, best_eval, best_boost_rounds
while(early_break >= 0):
np.random.seed(seed)
best_eval_init = best_eval
gs_params = {
(subsample, colsample)
for subsample in np.random.choice([i/10. for i in range(5,11)], 3)
for colsample in np.random.choice([i/10. for i in range(5,11)], 3)
}
best_model, best_param, best_eval, best_boost_rounds = _gs_helper('subsample',
'colsample_bytree',
best_model,
best_param,
best_eval,
best_boost_rounds)
gs_params = {
(max_depth, min_child_weight)
for max_depth in [10] + list(np.random.randint(1, 10, 3))
for min_child_weight in [0, 10] + list(np.random.randint(0, 10, 3))
}
best_model, best_param, best_eval, best_boost_rounds = _gs_helper('max_depth',
'min_child_weight',
best_model,
best_param,
best_eval,
best_boost_rounds)
gs_params = {
(eta, gamma)
for eta in np.random.choice([.005, .01, .05, .1, .2, .3], 3)
for gamma in [0] + list(np.random.choice([0.01, 0.001, 0.2, 0.5, 1.0, 2.0, 3.0, 5.0, 10.0], 3))
}
best_model, best_param, best_eval, best_boost_rounds = _gs_helper('eta',
'gamma',
best_model,
best_param,
best_eval,
best_boost_rounds)
if (abs(best_eval_init - best_eval) < thresh):
early_break-=1
seed+=1
return best_model, best_param, best_eval, best_boost_rounds
# -
best_model, best_param, best_eval, best_boost_rounds = xgb_paramsearch(X_train, y_train, features, init_params=init_param, nfold=5, num_boost_round=2000, early_stopping_rounds=3, metrics={'logloss', 'error'}, use_gpu='True')
param['alpha'] = 50
cv_model[f"test-merror-mean"].min()
best_param
*** only use PEG (try to find 100nm)
*** maybe look at different features (poor distributions)
heterogenious in different ways
different features are responsible to accuracies
*** think about to present code/results!
# +
evals = [(dtrain, 'train'), (dval, 'eval')]
num_round = best_boost_rounds
bst = xgb.train(best_param, dtrain, num_round, evals, early_stopping_rounds=3, )
######
label = dtest.get_label()
ypred1 = bst.predict(dtest)
# by default, we predict using all the trees
alpha = 0.62
pred = [0 if i < alpha else 1 for i in ypred1]
print("Accuracy:",metrics.accuracy_score(y_test, pred))
# -
from datetime import date
import json
bst.save_model(f'model_xgboost_region_based_cortex_striatum_80_20_split_{str(date.today())}')
with open(f'config_xgboost_region_based_cortex_striatum_80_20_split_{str(date.today())}', 'w', encoding='utf-8') as f:
json.dump(bst.save_config(), f, ensure_ascii=False, indent=4)
from datetime import date
import json
bst.load_model(f'model_xgboost_P14_P21_P28_P32_50-50-split_2020-07-18')
with open(f'config_xgboost_P14_P21_P28_P32_50-50-split_2020-07-18', 'r', encoding='utf-8') as f:
config = f.read()
config = json.loads(config)
setting = bst.load_config(config)
# +
ypred1 = bst.predict(dtest)
# by default, we predict using all the trees
pred = [0 if i < alpha else 1 for i in ypred1]
print("Accuracy:",metrics.accuracy_score(y_test, pred))
# -
model_bytearray = bst.save_raw()[4:]
def myfun(self=None):
return model_bytearray
bst.save_raw = myfun
# +
# import ctypes
# def c_array(ctype, values):
# """Convert a python string to c array."""
# if (isinstance(values, np.ndarray)
# and values.dtype.itemsize == ctypes.sizeof(ctype)):
# return (ctype * len(values)).from_buffer_copy(values)
# return (ctype * len(values))(*values)
# mats = c_array(ctypes.c_void_p, [dtrain.handle])
# +
# tst = X_test[features + [target]]
# tst['tst'] = y_test
# -
results = X_test[features]
results['predicted'] = pred
results['actual'] = y_test
# ## 4. Evaluate Results
# +
print('0 == {}'.format(le.inverse_transform([0])))
print('1 == {}'.format(le.inverse_transform([1])))
class_names = ['cortex', 'striatum']
class_results = classification_report(y_test, pred, digits=4, target_names = class_names)
print(str(class_results))
# +
confusion_matrix(y_test, pred)
pl.figure(figsize=(12,10))
cm_array = confusion_matrix(y_test, pred)
df_cm = pd.DataFrame(cm_array, index = class_names, columns = class_names)
sn.set(font_scale=1.4) # for label size
ax = sn.heatmap(df_cm, annot=True, annot_kws={"size": 16}, cmap="YlGnBu")
ax.set(xlabel='Actual', ylabel='Predicted')
pl.show()
# -
explainer = shap.TreeExplainer(bst)
shap_values = explainer.shap_values(X_test[features])
# %matplotlib inline
colors = ['#999999', '#7995e9']
class_inds = np.argsort([-np.abs(shap_values[i]).mean() for i in range(len(shap_values))])
cmap = plt_colors.ListedColormap(np.array(colors)[class_inds])
# +
# sn.reset_orig() # Reset matplot lib to no longer use seaborn
# -
shap.summary_plot(shap_values, X_test[features], class_names=np.array(class_names), title='Total SHAP Values', plot_type='bar', color='#999999')
# +
pl.ioff()
# %matplotlib inline
#------SHAP-FILE--------------
import random
def get_cmap(shap_values):
class_inds = np.argsort([-np.abs(shap_values[i]).mean() for i in range(len(shap_values))])
cmap = plt_colors.ListedColormap(np.array(colors)[class_inds])
return cmap
def plot_dependency(feature_name, shap_values, X_df, fig_dim, color, figsize=None, y_range=None, alpha=None):
if len(list(color)) is not 1:
color = get_cmap(shap_values)
colors = enumerate(color)
fig, axs = pl.subplots(*fig_dim, figsize=figsize)
# ax = axs.ravel()
cnt = 0
if (fig_dim == (1, 1)):
if figsize is not None:
axs[x][y].set_ylim(*figsize)
shap.dependence_plot(feature_name, shap_values, X_df, interaction_index=None, color=next(colors)[1], ax=axs)
else:
for x in range(fig_dim[0]):
for y in range(fig_dim[1]):
if figsize is not None:
axs[x][y].set_ylim(*figsize)
shap.dependence_plot(feature_name, shap_values, X_df, interaction_index=None, color=next(colors)[1], ax=axs[x][y])
cnt+=1
plot_dependency("Mean Deff1", shap_values, X_test[features], (1,1), ['#999999'])
# -
plot_dependency("Mean fractal_dim", shap_values, X_test[features], (1,1), ['#999999'])
plot_dependency("Mean kurtosis", shap_values, X_test[features], (1,1), ['#999999'])
plot_dependency("straightness", shap_values, X_test[features], (1,1), ['#999999'])
plot_dependency("Mean alpha", shap_values, X_test[features], (1,1), ['#999999'])
shap.summary_plot(shap_values, X_test[features], max_display=5, class_names = class_names, title = 'SHAP Value cortex')
# +
from modules import anim_plot_changed
from importlib import reload
reload(anim_plot_changed)
_ = anim_plot_changed.rotate_3d(results, [top_feat[0], top_feat[1], top_feat[2]])
_ = anim_plot_changed.rotate_3d(results, [top_feat[0], top_feat[2], top_feat[3]])
_ = anim_plot_changed.rotate_3d(results, [top_feat[1], top_feat[2], top_feat[3]])
# +
from modules import anim_plot_changed
from importlib import reload
reload(anim_plot_changed)
_ = anim_plot_changed.rotate_3d(results, [top_feat[0], top_feat[1], top_feat[2]], anim_param={'frames':np.arange(0,720,1)}, save_param={'filename':'This_is_a_test.gif','fps':50})
# -
from matplotlib import animation
from matplotlib.animation import PillowWriter
from sklearn import model
print(model.feature_importances_)
# +
# Feature search (new) -------not in file--------:
import operator
from sklearn.metrics import accuracy_score
def feature_thresholding_helper(X_train, X_test, X_val, new_feat):
dtrain = xgb.DMatrix(X_train[new_feat], label=y_train)
dtest = xgb.DMatrix(X_test[new_feat], label=y_test)
dval = xgb.DMatrix(X_val[new_feat], label=y_val)
return dtrain, dtest, dval
def feature_thresholding(X_train, y_train, X_test, y_test, X_val, y_val, params, features, nfold=5, num_boost_round=2000, early_stopping_rounds=3, metrics={'mlogloss', 'merror'}, thresh=np.arange(0,.1,.002)):
best_thresh = -1
if params['eval_metric'] in ['map', 'auc', 'aucpr']:
best_eval = -np.inf
eval_f = operator.gt
else:
best_eval = np.inf
eval_f = operator.lt
best_eval = -np.inf
eval_f = operator.gt
for t in thresh:
print(f"Using thresh = {t} ",end = '| ')
new_feat = list(np.array(features)[np.array(model.feature_importances_ > t)])
# cv_model = cv(params,
# X_train,
# y_train,
# features=new_feat,
# nfold=nfold,
# num_boost_round=num_boost_round,
# early_stopping_rounds=early_stopping_rounds,
# metrics=metrics)
# cv_eval = cv_model[f"test-{'merror'}-mean"].min()
# print(f"Eval = {cv_eval} ", end = '| ')
# if eval_f(cv_eval, best_eval):
# best_thresh = t
# best_eval = cv_eval
dtrain, dtest, dval = feature_thresholding_helper(X_train, X_test, X_val, new_feat)
evals = [(dtrain, 'train'), (dval, 'eval')]
bst2 = xgb.train(best_param, dtrain, 1500, evals, early_stopping_rounds=3, verbose_eval=False)
######
label = dtest.get_label()
ypred1 = bst2.predict(dtest)
# by default, we predict using all the trees
pred2 = [np.where(x == np.max(x))[0][0] for x in ypred1]
cv_eval = accuracy_score(y_test, pred2)
if eval_f(cv_eval, best_eval):
best_thresh = t
best_eval = cv_eval
print(f"Best eval = {best_eval}, Best threshold = {best_thresh}")
print(f"Features used:\n{np.array(features)[np.array(model.feature_importances_ > best_thresh)]}")
return list(np.array(features)[np.array(model.feature_importances_ > best_thresh)])
new_feat = feature_thresholding(X_train, y_train, X_test, y_test, X_val, y_val, best_param, features)
# -
new_feat = list(np.array(features)[np.array(model.feature_importances_ > best_thresh)])
cv_model = cv(best_param,
X_train,
y_train,
features=new_feat,
nfold=5,
num_boost_round=best_boost_rounds,
early_stopping_rounds=3,
metrics={'mlogloss', 'merror'})
cv_model
dtrain = xgb.DMatrix(X_train[new_feat], label=y_train)
dtest = xgb.DMatrix(X_test[new_feat], label=y_test)
dval = xgb.DMatrix(X_val[new_feat], label=y_val)
# +
evals = [(dtrain, 'train'), (dval, 'eval')]
num_round = best_boost_rounds
bst = xgb.train(best_param, dtrain, num_round, evals, early_stopping_rounds=3, )
######
label = dtest.get_label()
ypred1 = bst.predict(dtest)
# by default, we predict using all the trees
pred = [np.where(x == np.max(x))[0][0] for x in ypred1]
# -
# print('0 == {}'.format(le.inverse_transform([0])))
# print('1 == {}'.format(le.inverse_transform([1])))
# print('2 == {}'.format(le.inverse_transform([2])))
# print('3 == {}'.format(le.inverse_transform([3])))
class_names = ['P14', 'P21', 'P28', 'P35']
class_results = classification_report(y_test, pred, digits=4, target_names = ['P14', 'P21', 'P28', 'P35'])
print(str(class_results))
# Running CV with newly thresholded features; using new seed of 123 to get different unique GS hyperparams
best_model2, best_param2, best_eval2, best_boost_rounds2 = xgb_paramsearch(X_train, y_train, new_feat, init_params=best_param, nfold=5, num_boost_round=2000, early_stopping_rounds=3, metrics={'mlogloss', 'merror'}, use_gpu='True', seed=123)
# +
seed = 1234
np.random.seed(seed)
train_split = 0.7
test_split = 0.5
le = preprocessing.LabelEncoder()
bal_ecm['encoded_target'] = le.fit_transform(bal_ecm[target])
training_bins = np.random.choice(bal_ecm.bins.unique(), int(len(bal_ecm.bins.unique())*train_split), replace=False)
X_train = bal_ecm[bal_ecm.bins.isin(training_bins)]
X_test_val = bal_ecm[~bal_ecm.bins.isin(training_bins)]
X_val, X_test = train_test_split(X_test_val, test_size=test_split, random_state=seed)
y_train = X_train['encoded_target']
y_test = X_test['encoded_target']
y_val = X_val['encoded_target']
dtrain = xgb.DMatrix(X_train[new_feat], label=y_train)
dtest = xgb.DMatrix(X_test[new_feat], label=y_test)
dval = xgb.DMatrix(X_val[new_feat], label=y_val)
# -
best_param2={'max_depth': 5,
'eta': 0.005,
'min_child_weight': 10,
'verbosity': 0,
'objective': 'multi:softprob',
'num_class': 4,
'silent': 'True',
'gamma': 5,
'subsample': 0.6,
'colsample_bytree': 0.5,
'eval_metric': 'mlogloss',
'gpu_id': 0,
'tree_method': 'gpu_hist',
'predictor': 'gpu_predictor'}
# +
evals = [(dtrain, 'train'), (dval, 'eval')]
num_round = best_boost_rounds
bst2 = xgb.train(best_param, dtrain, num_round, evals, early_stopping_rounds=3, )
######
label = dtest.get_label()
ypred1 = bst2.predict(dtest)
# by default, we predict using all the trees
pred2 = [np.where(x == np.max(x))[0][0] for x in ypred1]
print("Accuracy:",metrics.accuracy_score(y_test, pred2))
# -
class_names = ['P14', 'P21', 'P28', 'P35']
class_results = classification_report(y_test, pred2, digits=4, target_names = ['P14', 'P21', 'P28', 'P35'])
print(str(class_results))
# +
# param2 = {'max_depth': 2,
# 'eta': 0.005,
# 'min_child_weight': 0,
# 'verbosity': 0,
# 'objective': 'multi:softprob',
# 'num_class': 4,
# 'silent': 'True',
# 'gamma': 5,
# 'subsample': 0.25,
# 'colsample_bytree': 0.3,
# 'colsample_bynode':.5,
# 'reg_alpha': 0}
# +
from sklearn.metrics import accuracy_score
model_final = XGBClassifier(**param2)
new_feat = np.array(features)[np.array(model.feature_importances_ > t)]
eval_set = [(X_train[new_feat], y_train), (X_test[new_feat], y_test)]
model_final.fit(X_train[new_feat], y_train, verbose=False, eval_set=eval_set, eval_metric=["merror", 'mlogloss'])
y_pred_f = model_final.predict(X_test[new_feat])
accuracy = accuracy_score(y_test, y_pred_f)
print("Accuracy: %.2f%%" % (accuracy * 100.0))
results = model_final.evals_result()
epochs = len(results['validation_0']['merror'])
x_axis = range(0, epochs)
fig, ax = pl.subplots(figsize=(12,12))
ax.plot(x_axis, results['validation_0']['mlogloss'], label='Train')
ax.plot(x_axis, results['validation_1']['mlogloss'], label='Test')
ax.legend()
pl.ylabel('Log Loss')
pl.title('XGBoost Log Loss')
pl.show()
# -
sorted(dict_importance, key=dict_importance.get, reverse=True)[:5]
new_feat = np.array(features)[np.array(model.feature_importances_ > best_thresh)]
model2.fit(X_train[new_feat], y_train, verbose=False, eval_set=[(X_val[new_feat],y_val)], eval_metric='mlogloss')
pred3 = model2.predict(X_test[new_feat])
acc = metrics.accuracy_score(y_test, pred3)
print("Accuracy:",metrics.accuracy_score(y_test, pred3))
|
notebooks/.ipynb_checkpoints/XGBoost_region_analysis_cortex_striatum-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "slide"}
# # Tours through the Book
#
# This book is _massive_. With 17,000 lines of code and 125,000 words of text, a printed version would cover more than 1,000 pages of text. Obviously, we do not assume that everybody wants to read everything.
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "slide"}
# While the chapters of this book can be read one after the other, there are many possible paths through the book. In this graph, an arrow $A \rightarrow B$ means that chapter $A$ is a prerequisite for chapter $B$. You can pick arbitrary paths in this graph to get to the topics that interest you most:
# -
from IPython.display import SVG
SVG(filename='PICS/Sitemap.svg')
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "slide"}
# But since even this map can be overwhelming, here are a few _tours_ to get you started. Each of these tours allows you to focus on a particular view, depending on whether you are a programmer, student, or researcher.
# -
# ## The Pragmatic Programmer Tour
#
# You have a program to test. You want to generate tests as quickly as possible and as thorough as possible. You don't care so much how something is implemented, but it should get the job done. You want to get to the point.
#
# 1. __Start with [Introduction to Testing](Intro_Testing.ipynb) to get the basic concepts.__ (You would know most of these anyway, but it can't hurt to get quick reminders).
#
# 2. __Use the simple fuzzers from [the chapter on Fuzzers](Fuzzer.ipynb)__ to test your program against the first random inputs.
#
# 3. __Get [coverage](Coverage.ipynb) from your program__ and use coverage information to [guide test generation towards code coverage](GreyboxFuzzer.ipynb).
#
# 4. __Define an [input grammar](Grammars.ipynb) for your program__ and use this grammar to thoroughly fuzz your program with syntactically correct inputs. As fuzzer, we would recommend a [grammar coverage fuzzer](GrammarCoverageFuzzer), as this ensures coverage of input elements.
#
# 5. If you want __more control over the generated inputs,__ consider [probabilistic fuzzing](ProbabilisticGrammarFuzzer.ipynb) and [fuzzing with generator functions](GeneratorGrammarFuzzer.ipynb).
#
# 6. If you want to __deploy a large set of fuzzers__, learn how to [manage a large set of fuzzers](FuzzingInTheLarge.ipynb).
#
# In each of these chapters, start with the "Synopsis" parts; these will give you quick introductions on how to use things, as well as point you to relevant usage examples. With this, enough said. Get back to work and enjoy!
# ## The Page-by-Page Tours
#
# These tours are how the book is organized. Having gone through the [Introduction to Testing](Intro_Testing.ipynb) for the basic concepts, you can read your way through these parts:
#
# 1. __The [lexical tour](02_Lexical_Fuzzing.ipynb)__ focuses on _lexical_ test generation techniques, i.e. techniques that compose an input character by character and byte by byte. Very fast and robust techniques with a minimum of bias.
#
# 1. __The [syntactical tour](03_Syntactical_Fuzzing.ipynb)__ focuses on _grammars_ as a means to specify the syntax of inputs. The resulting test generators produce syntactically correct inputs, making tests much faster, and provide lots of control mechanisms for the tester.
#
# 1. __The [semantical tour](04_Semantical_Fuzzing.ipynb)__ makes use of _code semantics_ to shape and guide test generation. Advanced techniques include extracting input grammars, mining function specifications, and symbolic constraint solving to cover as many code paths as possible.
#
# 1. __The [application tour](05_Domain-Specific_Fuzzing.ipynb)__ applies the techniques defined in the earlier parts on domains such as Web servers, user interfaces, APIs, or configurations.
#
# 1. __The [management tour](06_Managing_Fuzzing.ipynb)__ finally focuses on how to handle and organize large sets of test generators, and when to stop fuzzing.
# Most of these chapters start with a "Synopsis" section that explains how to use the most important concepts. You can choose whether you want a "usage" perspective (then just read the synopsis) or a "understanding" perspective (then read on).
# ## The Undergraduate Tour
#
# You are a student of computer science and/or software engineering. You want to know basics of testing and related fields. You not only want to use techniques, but also dig deeper into algorithms and implementations. We have the following recommendation for you:
#
# 1. Start with [Introduction to Testing](Intro_Testing.ipynb) and [Coverage](Coverage.ipynb) to get the __basic concepts.__ (You may know some of these already, but hey, you're a student, right?)
#
# 2. __Learn how simple fuzzers work__ from [the chapter on Fuzzers](Fuzzer.ipynb). This already gives you tools that took down 30% of UNIX utilities in the 90s. What happens if you test some tool that never has been fuzzed before?
#
# 3. __[Mutation-based fuzzing](MutationFuzzer.ipynb)__ is pretty much the standard in fuzzing today: Take a set of seeds, and mutate them until we find a bug.
#
# 4. __Learn how [grammars](Grammars.ipynb) can be used to generate syntactically correct inputs.__ This makes test generation much more efficient, but you have to write (or [mine](GrammarMiner.ipynb) a grammar in the first place.
#
# 5. __Learn how to [fuzz APIs](APIFuzzer.ipynb) and [graphical user interfaces](GUIFuzzer.ipynb)__. Both of these are important domains for software test generation.
#
# 6. __Learn how to [reduce failure-inducing inputs](Reducer.ipynb) to a minimum automatically__. This is a great time saver for debugging, especially in conjunction with automated testing.
#
# For all these chapters, experiment with the implementations to understand their concepts. Feel free to experiment as you wish.
# If you are a teacher, the above chapters can be useful in programming and/or software engineering courses. Make use of slides and/or live programming, and have students work on exercises.
# ## The Graduate Tour
#
# On top of the "Undergraduate" tour, you want to get deeper into test generation techniques, including techniques that are more demanding.
#
# 1. __[Search-based testing](SearchBasedFuzzer.ipynb)__ allows you to guide test generation towards specific goals, such as code coverage. Robust and efficient.
#
# 1. Get an introduction to __[configuration testing](ConfigurationFuzzer.ipynb)__. How does one test and cover a system that comes with multiple configuration options?
#
# 1. __[Mutation analysis](MutationAnalysis.ipynb)__ seeds synthetic defects (mutations) into program code to check whether the tests find them. If the tests do not find mutations, they likely won't find real bugs either.
#
# 1. __Learn how to [parse](Parser.ipynb) inputs__ using grammars. If you want to analyze, decompose, mutate existing inputs, you need a parser for that.
#
# 1. __[Concolic](ConcolicFuzzer.ipynb) and [symbolic](SymbolicFuzzer.ipynb) fuzzing__ solve constraints along program paths to reach code that is hard to test. Used wherever reliability is paramount; also a hot research topic.
#
# 1. __Learn how to [estimate when to stop fuzzing](WhenToStopFuzzing.ipynb)__. There has to be a stop at some point, right?
#
# For all these chapters, experiment with the code; feel free to create your own variations and extensions. This is how we get to research!
# If you are a teacher, the above chapters can be useful in advanced courses on software engineering and testing. Again, you can make use of slides and/or live programming, and have students work on exercises.
# ## The Researcher Tour
#
# On top of the "Graduate" tour, you are looking for techniques that are somewhere between lab stage and widespread usage – in particular, techniques where there is still room for lots of improvement. If you look for research ideas, go for these topics.
#
# 1. __[Mining function specifications](DynamicInvariants.ipynb)__ is a hot topic in research: Given a function, how can we infer an abstract model that describes its behavior? The conjunction with test generation offers several opportunities here, in particular for dynamic specification mining.
#
# 2. __[Mining input grammars](GrammarMiner.ipynb)__ promises to join the robustness and ease of use of lexical fuzzing with the efficiency and speed of syntactical fuzzing. The idea is to mine an input grammar from a program automatically, which then serves as base for syntactical fuzzing. Still in an early stage, but lots of potential.
#
# 3. __[Probabilistic grammar fuzzing](ProbabilisticGrammarFuzzer.ipynb)__ gives programmers lots of control over which elements should be generated. Plenty of research possibilities at the intersection of probabilistic fuzzing and mining data from given tests, as sketched in this chapter.
#
# 4. __[Fuzzing with generators](GeneratorGrammarFuzzer.ipynb)__ gives programmers the ultimate control over input generation, namely by allowing them to define their own generator functions. The big challenge is: How can one best exploit the power of syntactic descriptions with a minimum of contextual constraints?
#
# 5. __[Carving unit tests](Carver.ipynb)__ brings the promise of speeding up test execution (and generation) dramatically, by extracting unit tests from program executions that replay only individual function calls (possibly with new, generated arguments). In Python, carving is simple to realize; here's plenty of potential to toy with.
#
# 6. __Testing [web servers](WebFuzzer.ipynb) and [GUIs](GUIFuzzer.ipynb)__ is a hot research field, fueled by the need of practitioners to test and secure their interfaces (and the need of other practitioners to break through these interfaces). Again, there's still lots of unexplored potential here.
#
# 7. __[Greybox fuzzing](GreyboxFuzzer.ipynb) and [greybox fuzzing with grammars](GreyboxGrammarFuzzer.ipynb)__ bring in _statistical estimators_ to guide test generation towards inputs and input properties that are most likely to discover new bugs. The intersection of testing, program analysis, and statistics offers lots of possibilities for future research.
# For all these topics, having Python source available that implements and demonstrates the concepts is a major asset. You can easily extend the implementations with your own ideas and run evaluations right in a notebook. Once your approach is stable, consider porting it to a language with a wider range of available subjects (such as C, for example).
# ## The Author Tour
#
# This is the ultimate tour – you have learned everything there is and want to contribute to the book. Then, you should read two more chapters:
#
# 1. The __[guide for authors](Guide_for_Authors.ipynb)__ gives an introduction on how to contribute to this book (coding styles, writing styles, conventions, and more).
#
# 2. The __[template chapter](Template.ipynb)__ serves as a blueprint for your chapter.
#
# If you want to contribute, feel free to contact us – preferably before writing, but after writing is fine just as well. We will be happy to incorporate your material.
# + [markdown] button=false new_sheet=true run_control={"read_only": false}
# ## Lessons Learned
#
# * You can go through the book from beginning to end...
# * ...but it may be preferable to follow a specific tour, based on your needs and resources.
# * Now [go and explore generating software tests](index.ipynb)!
|
notebooks/Tours.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Bolsa de valores
# Prevendo o volume de ações utilizando o modelo Random Forest Regressor com inclusão de indicadores financeiros como CMO, DX e MFI
# * Utilizaremos dados do Yahoo Finance da PETR4 num intervalo de tempo específico
# * Dados serão obtido através da biblioteca Pandas DataReader
# Importando bibliotecas
import datetime as dt
import pandas_datareader.data as web
import matplotlib.pyplot as plt
import numpy as np
# Definindo janela de tempo
start = dt.datetime(2018,1,1)
end = dt.datetime(2020,9,30)
# Obtendo dados da PETR4
PETR4 = web.DataReader('PETR4.SA',"yahoo",start,end)
# Exibindo as cinco primeiras linhas
PETR4.head()
# Adicionando indicadores
High = PETR4['High'].values
Low = PETR4['Low'].values
Open = PETR4['Open'].values
Close = PETR4['Close'].values
Volume = PETR4['Volume'].values
from talib._ta_lib import ADX, APO, CCI, CMO, DX, RSI, DX, MACD, MFI, ROC, RSI, ULTOSC
# Indicador ADX
PETR4['ADX'] = ADX(High, Low, Close, timeperiod=14)
# Indicador APO
PETR4['APO'] = APO(Close, fastperiod=12, slowperiod=26, matype=0)
# Indicador CCI
PETR4['CCI'] = CCI(High, Low, Close, timeperiod=14)
# Indicador CMO
PETR4['CMO'] = CMO(Close, timeperiod=14)
# Indicador DX
PETR4['DX'] = DX(High, Low, Close, timeperiod=14)
# Indicador MACD
macd, macdsignal, macdhist = MACD(Close, fastperiod=12, slowperiod=26, signalperiod=9)
PETR4['MACD'] = macd
# Indicador MFI
PETR4['MFI'] = MFI(High, Low, Close, Volume, timeperiod=14)
# Indicador ROC
PETR4['ROC'] = ROC(Close, timeperiod=14)
# Indicador RSI
PETR4['RSI'] = RSI(Close, timeperiod=14)
# Indicador ULTOSC
PETR4['ULTOSC'] = ULTOSC(High, Low, Close, timeperiod1=7, timeperiod2=14, timeperiod3=28)
# Verificando amostra
PETR4.head()
# Removendo NaNs
PETR4 = PETR4.dropna()
PETR4.head()
# * Removendo High, Low, Open, Close, Adj Close
PETR4 = PETR4.drop(['High','Low','Open','Close', 'Adj Close'],axis=1)
# * Normalizando dados do Volume
from sklearn.preprocessing import RobustScaler
scaler = RobustScaler()
PETR4['Volume'] = scaler.fit_transform(PETR4['Volume'].values.reshape(-1, 1))
# Definindo variáveis X e Y
X = PETR4.drop(['Volume'],axis=1)
Y = PETR4['Volume']
# * Criando amostra de treino e teste
# +
X_treino = X[X.index<'2020-08-01']
X_teste = X[X.index>='2020-08-01']
Y_treino = Y[X.index<'2020-08-01']
Y_teste = Y[X.index>='2020-08-01']
# -
# * Prevendo volume utilizando modelo Random Forest Regressor
from sklearn.ensemble import RandomForestRegressor
rfr = RandomForestRegressor()
rfr.fit(X_treino,Y_treino)
Y_previsto = rfr.predict(X_teste)
# * Desnormalizando dados
Y_previsto = scaler.inverse_transform(Y_previsto.reshape(-1, 1))
Y_teste = scaler.inverse_transform(Y_teste.values.reshape(-1, 1))
# * Graficando Y_previsto em função de Y_teste
plt.scatter(Y_teste,Y_previsto)
plt.xlabel('Y_teste')
plt.ylabel('Y_previsto')
plt.tight_layout()
# Calculando métricas de erro
from sklearn.metrics import mean_absolute_error, mean_squared_error
def mean_absolute_percentage_error(y_true, y_pred):
y_true, y_pred = np.array(y_true), np.array(y_pred)
return np.mean(np.abs((y_true - y_pred) / y_true)) * 100
MAE = mean_absolute_error(Y_teste,Y_previsto)
MAPE = mean_absolute_percentage_error(Y_teste,Y_previsto)
MSE = mean_squared_error(Y_teste,Y_previsto)
RMSE = np.sqrt(MSE)
print("MAE = {:0.2e}".format(MAE))
print("MAPE = {:0.2f}%".format(MAPE))
print("MSE = {:0.2e}".format(MSE))
print("RMSE = {:0.2e}".format(RMSE))
# Podemos prever o valor com uma incerteza de 24.26%. Um pouco melhor do que o caso sem indicadores que foi de 28.78%.
|
Data_Science/Economia/PETR4-Indicadores.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### 数据解析
# Import required libraries
from tpot import TPOTRegressor
from sklearn.model_selection import train_test_split
import pandas as pd
import numpy as np
# Load the data
medical = pd.read_csv('train_fenlie1.csv', low_memory=False)
medical.head(5)
# ### 数据整理
medical.dtypes
for cat in ['Molecule_max_phase', 'RO5_violations']:
print("Number of levels in category '{0}': \b {1:2.2f} ".format(cat, medical[cat].unique().size))
medical = medical.fillna(-999)
pd.isnull(medical).any()
# #### 去掉不相关特征
medical_new = medical.drop(['ID','Label'], axis=1)
medical_new = medical_new.fillna(-999)
pd.isnull(medical_new).any()
medical_new = np.array(medical_new.values,dtype=float)
medical_new.shape
# medical_new.astype(float)
medical_new.dtype
np.isnan(medical_new)
medical_new[np.isnan(medical_new)] = -999
np.isnan(medical_new)
medical_new.shape
medical_label = medical['Label'].values
# medical_label = medical_label[:,np.newaxis]
medical_label
# ## 使用TPOT进行数据分析
X_train, X_test, y_train, y_test = train_test_split(medical_new[:1000],medical_label[:1000],
train_size=0.75, test_size=0.25, random_state=42)
X_train.shape, X_test.shape, y_train.size, y_test.size
tpot = TPOTRegressor(generations=5, population_size=50, verbosity=2, random_state=42)
tpot.fit(X_train, y_train)
print(tpot.score(X_test, y_test))
tpot.export('tpot_medical_pipeline_fenlie1.py')
# medical_new = medical.drop(['ID','Label'], axis=1)
|
Baseline_train_fenlie1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 0. Import Packages
import sympy
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# ## 1. Gradient Vector Field and Contour
# +
#define symbolic vars, function
x, y = sympy.symbols('x y')
fun = (x-1)**2+(y-1)**2
#take the gradient symbolically
gradfun = [sympy.diff(fun,var) for var in (x,y)]
#turn into a bivariate lambda for numpy
numgradfun = sympy.lambdify([x,y],gradfun)
x = np.linspace(-4, 6, 15)
y = np.linspace(-4, 6, 15)
x1 = np.linspace(-4, 6, 100)
y1 = np.linspace(-4, 6, 100)
X,Y = np.meshgrid(x, y)
X1, Y1 = np.meshgrid(x1, y1)
graddat = numgradfun(X,Y)
Z = (X1-1)**2 + (Y1-1)**2
plt.figure()
plt.axes().set_aspect('equal', 'datalim')
plt.quiver(X,Y,graddat[0],graddat[1])
plt.contour(x1, y1, Z, cmap='seismic')
plt.show()
# -
# ## 2. Linear Regression
# - x = np.array([1, 2, 3])
# - y = np.array([4.1, 6.9, 9.9])
# - Hypothesis H = wx+b
#
# +
x = np.array([1, 2, 3])
y = np.array([4.1, 6.9, 9.9])
w = 0
b = 0
a = 0.05 # learning rate
H = w*x + b
for i in range(20):
dw = 2*sum(x*(H-y))/len(y)
db = 2*sum(H-y)/len(y)
w = w - a*dw
b = b - a*db
H = w*x+b
c = sum((H-y)*(H-y))/len(y)
plt.plot(x, y, "ro")
plt.plot(x, w*x+b, "b-")
print("w: ", w, "b: ", b, "Cost: ", c)
print("Prediction:", w*4+b)
# -
# - x = np.array([1, 2, 3])
# - y = np.array([4.1, 6.9, 9.9])
# - Hypothesis H = wx
# +
x = np.array([1, 2, 3])
y = np.array([4.1, 6.9, 9.9])
w = 0
a = 0.05 # learning rate
H = w*x
for i in range(20):
g = 2*sum(x*(H-y))/len(y)
w = w - a*g
H = w*x
c = sum((H-y)*(H-y))/len(y)
plt.plot(x, y, "ro")
plt.plot(x, w*x, "b-")
print("w: ", w, "Cost: ", c)
print("Prediction:", w*4)
# -
# - x = np.array([30, 20, 40])
# - y = np.array([6.1, 3.1, 6.8])
# - Hypothesis H = w*x + b
# +
x = np.array([30, 20, 40])
y = np.array([6.1, 3.1, 6.8])
w = 0
b = 0
a = 0.001 # learning rate
H = w*x + b
for i in range(1000):
dw = 2*sum(x*(H-y))/len(y)
db = 2*sum(H-y)/len(y)
w = w - a*dw
b = b - a*db
H = w*x+b
c = sum((H-y)*(H-y))/len(y)
if i % 100 == 0:
print("w: ", w, "b: ", b, "Cost: ", c)
print("Prediction:", w*25+b)
# -
# ## 2. Multi-Variable Linear regression
# +
X = np.matrix([[30, 20, 40], [1, 2, 3], [5, 10, 15]])
Y = np.matrix([6.1, 3.1, 6.8])
W = np.matrix([0, 0, 0])
b = 0
a = 1e-4 # learning rate
H = W*X + b
for t in range(3000):
dW = 2*((H-Y)*X.T)/Y.shape[1] # gradient
db = 2*np.mean(H-Y)
W = W - a*dW
b = b - a*db
c = (H-Y)*(H-Y).T/Y.shape[1]
H = W*X+b
if t % 100 == 0:
print('Iteration:', t, 'Cost:', c[0,0])
print('')
X_test = np.matrix([[25], [2], [10]])
print("Price Prediction:", (W*X_test+b)[0,0])
|
4_Regression/Gradient_Vector_and_Linear_Regression.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="QWUdwHrBkctr"
# # CIFAR10_Test
#
# 使用CIFAR10数据集对CNN进行训练及测试
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="Ykfy4xGQkct0"
import tensorflow as tf
import numpy as np
import math
#import matplotlib.pyplot as plt
# #%matplotlib inline
import time
import os
import tensornets as nets
import cv2
import scipy.io as sio
# for auto-reloading external modules
# see http://stackoverflow.com/questions/1907993/autoreload-of-modules-in-ipython
# %load_ext autoreload
# %autoreload 2
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 125, "output_extras": [{"item_id": 1}]} colab_type="code" executionInfo={"elapsed": 31366, "status": "ok", "timestamp": 1522747101694, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "115487754759868122157"}, "user_tz": -480} id="zG95i-smkcuA" outputId="e588e385-dee3-40c0-cb6f-e792f146bc08"
from cs231n.data_utils import load_CIFAR10
import sys
if sys.platform == "linux" :
cifar10_dir = "/home/z_tomcato/cs231n/assignment2/assignment2/cs231n/datasets/cifar-10-batches-py"
else:
cifar10_dir = 'cs231n/datasets'
def get_CIFAR10_data(num_training=10000, num_validation=1000, num_test=10000):
"""
Load the CIFAR-10 dataset from disk and perform preprocessing to prepare
it for the two-layer neural net classifier. These are the same steps as
we used for the SVM, but condensed to a single function.
"""
# Load the raw CIFAR-10 data
# cifar10_dir = '../assignment2/cs231n/datasets'
X_train, y_train, X_test, y_test = load_CIFAR10(cifar10_dir)
# Subsample the data
mask = range(num_training, num_training + num_validation)
X_val = X_train[mask]
y_val = y_train[mask]
mask = range(num_training)
X_train = X_train[mask]
y_train = y_train[mask]
mask = range(num_test)
X_test = X_test[mask]
y_test = y_test[mask]
# Normalize the data: subtract the mean image
mean_image = np.mean(X_train, axis=0)
X_train -= mean_image
X_val -= mean_image
X_test -= mean_image
return X_train, y_train, X_val, y_val, X_test, y_test
# Invoke the above function to get our data.
X_train, y_train, X_val, y_val, X_test, y_test = get_CIFAR10_data()
print('Train data shape: ', X_train.shape)
print('Train labels shape: ', y_train.shape)
print('Validation data shape: ', X_val.shape)
print('Validation labels shape: ', y_val.shape)
print('Test data shape: ', X_test.shape)
print('Test labels shape: ', y_test.shape)
# -
# ## Tensornets测试
"""
inputs = tf.placeholder(tf.float32, [None, 224, 224, 3])
#model = nets.InceptionResNet2
model = nets.MobileNet100
newModel = model(inputs)
img = nets.utils.load_img('cat.png', target_size=256, crop_size=224)
assert img.shape == (1, 224, 224, 3)
with tf.Session() as sess:
img = newModel.preprocess(img) # equivalent to img = nets.preprocess(model, img)
sess.run(newModel.pretrained()) # equivalent to nets.pretrained(model)
scores = sess.run(newModel, {inputs: img})
#rint(preds)
print(nets.utils.decode_predictions(scores, top=1)[0])
#print(tf.argmax(tf.squeeze(preds, [0], axis = 1)))
predictions = tf.argmax(scores, axis = 1)
print(predictions)
[(u'n02124075', u'Egyptian_cat', 0.28067636), (u'n02127052', u'lynx', 0.16826575)]"""
# +
def load_img_from_tensor(x, target_size=None, crop_size=None, interp=cv2.INTER_CUBIC):
minSize = min(x.shape[1:3])
imgs = None
if target_size:
if isinstance(target_size, int):
hw_tuple = (x.shape[1] * target_size // minSize, x.shape[2] * target_size // minSize)
else:
hw_tuple = (target_size[1], target_size[0])
imgs = np.zeros((x.shape[0],hw_tuple[0],hw_tuple[1], 3), dtype=np.uint8)
if x.shape[1:3] != hw_tuple:
for i in range(x.shape[0]):
imgs[i,:, :, :] = cv2.resize(x[i, :, :, :], hw_tuple, interpolation=interp)
if crop_size is not None:
imgs = nets.utils.crop(imgs, crop_size)
return imgs
def img_preprocess(x):
# Copied from keras (equivalent to the same as in TF Slim)
x = x.copy()
x = x / 255.
x = x - 0.5
x = x * 2.
return x
# -
# ## Run Module
# +
#model = nets.MobileNet100
def run_model(session, Xd, yd, Xv, yv, num_class = 10, epochs=3, batch_size=100,print_every=10, learning_rate = 1e-5, dropout = 0.5):
print("Batch dataset initialized.\n# of training data: {}\n# of test data: {}\n# of class: {}"
.format(Xd.shape[0], Xv.shape[0], 10))
# shuffle indicies
train_indicies = np.arange(Xd.shape[0])
np.random.shuffle(train_indicies)
with tf.Session() as sess:
inputs = tf.placeholder(tf.float32, [None, 224, 224, 3])
outputs = tf.placeholder(tf.int32, [None])
cnn_net = nets.MobileNet100(inputs, is_training = True, classes = num_class)
cnn_loss = tf.losses.softmax_cross_entropy(tf.one_hot(outputs,num_class, dtype=tf.int32), cnn_net)
cnn_train = tf.train.AdamOptimizer(learning_rate=learning_rate).minimize(cnn_loss)
sess.run(tf.global_variables_initializer())
nets.pretrained(cnn_net)
# tensorboard setting
fileName = time.strftime("%Y%m%d_%H%M%S", time.localtime())
fileName = os.path.normcase("./result/" + fileName)
summary_writer = tf.summary.FileWriter(fileName, sess.graph)
global_step = 0
for current_epoch in range(epochs):
# training step
###for x_batch, y_batch in batch_set.batches():
print("#############################Epoch Start##############################")
for i in range(int(math.ceil(Xd.shape[0]/batch_size))):
start = time.time()
start_idx = (i*batch_size)%Xd.shape[0]
idx = np.int32(train_indicies[start_idx:start_idx+batch_size])
batch_Xd = load_img_from_tensor(Xd[idx,:, :, :], target_size=256, crop_size=224)
batch_Xd = cnn_net.preprocess(batch_Xd)
batch_yd = yd[idx]
feed = {inputs : batch_Xd, outputs : batch_yd}
global_step = global_step + 1
cnn_predictions = tf.argmax(cnn_net, axis = 1)
cnn_correct_prediction = tf.equal(tf.cast(cnn_predictions, dtype=tf.int32), batch_yd)
cnn_accuracy = tf.reduce_mean(tf.cast(cnn_correct_prediction, tf.float32))
train_summary = tf.summary.merge([tf.summary.scalar("train_loss", cnn_loss),
tf.summary.scalar("train_accuracy", cnn_accuracy)])
_, loss, scores,accuracy, summary = sess.run([cnn_train, cnn_loss,
cnn_net, cnn_accuracy, train_summary], feed_dict=feed)
summary_writer.add_summary(summary, global_step)
if global_step % print_every == 0:
print("{}/{} ({} epochs) step, loss : {:.6f}, accuracy : {:.3f}, time/batch : {:.3f}sec"
.format(global_step, int(round(Xd.shape[0]/batch_size)) * epochs, current_epoch,
loss, accuracy, time.time() - start))
# test step
start, avg_loss, avg_accuracy = time.time(), 0, 0
test_summary = tf.summary.merge([tf.summary.scalar("val_loss", cnn_loss),
tf.summary.scalar("val_accuracy", cnn_net)])
Xv = cnn_net.preprocess(Xv)
feed = {inputs : Xv, outputs : yv}
loss, accuracy, summary = sess.run([cnn_loss, cnn_accuracy, test_summary], feed_dict=feed)
summary_writer.add_summary(summary, current_epoch)
print("{} epochs test result. loss : {:.6f}, accuracy : {:.3f}, time/batch : {:.3f}sec"
.format(current_epoch, loss , accuracy , time.time() - start))
print("\n")
return
# -
def load_wiki(wiki_path, num_training=49000, num_validation=1000, num_test=1000,):
wiki_path = ""
if sys.platform == "linux" :
wiki_path = "/devdata/wiki/"
else:
wiki_path = "G:\\MachineLearning\\wiki\\wiki\\"
mat_path = wiki_path + 'wiki_with_age.mat'
data = sio.loadmat(mat_path)
img_paths = []
for i in range(len(wiki_data[6][0])):
wiki_data = data['wiki'][0][0]
face_score =wiki_data[6][0][i]
if face_score != float("-inf"):
full_path = wiki_path + wiki_data[0][0][2][0][i][0]
img = cv2.imread(full_path)
cv2.imshow("test", img)
face_loc = wiki_data[5][0][i][0]
print(face_loc)
face_loc = face_loc.astype("int32")
print(face_loc)
roi_img = img[face_loc[1]:face_loc[3], face_loc[0]:face_loc[2]]
temp_img = cv2.resize(roi_img, ((face_loc[3]-face_loc[1]) * 3, (face_loc[2]-face_loc[0]) * 3))
cv2.imshow("temp_img", temp_img)
cv2.imshow("roi_img", roi_img)
print("age: ", wiki_data[8][0][i])
gender = wiki_data[3][0][i]
if gender == 0:
print("女")
else:
print("男")
else:
print("没找到人脸")
tf.reset_default_graph()
with tf.Session() as sess:
#with tf.device("/cpu:0"): #"/cpu:0" or "/gpu:0"
#sess.run(tf.global_variables_initializer())
#print('Training')
run_model(sess,X_train,y_train,X_val,y_val, epochs=4, batch_size=500,print_every=100, learning_rate = 1e-5)
|
Wiki_test.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Plots y visualización de los datos
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
data = pd.read_csv("../datasets/customer-churn-model/Customer Churn Model.txt")
data
% matplotlib inline
# +
#savefig("path_donde_guardar_im.jpeg")
# -
# ### Scatter Plot
data.plot(kind="scatter", x="Day Mins", y="Day Charge")
data.plot(kind="scatter", x="Night Mins", y="Night Charge")
figure, axs = plt.subplots(2,2, sharey=True, sharex=True)
data.plot(kind="scatter", x="Day Mins", y ="Day Charge", ax=axs[0][0])
data.plot(kind="scatter", x="Night Mins", y="Night Charge", ax=axs[0][1])
data.plot(kind="scatter", x="Day Calls", y ="Day Charge", ax=axs[1][0])
data.plot(kind="scatter", x="Night Calls", y="Night Charge", ax=axs[1][1])
# ### Histogramas de frecuencias
k = int(np.ceil(1+np.log2(3333)))
plt.hist(data["Day Calls"], bins = k) #bins = [0,30,60,...,200]
plt.xlabel("Número de llamadas al día")
plt.ylabel("Frecuencia")
plt.title("Histograma del número de llamadas al día")
# ### Boxplot, diagrama de caja y bigotes
plt.boxplot(data["Day Calls"])
plt.ylabel("Número de llamadas diarias")
plt.title("Boxplot de las llamadas diarias")
data["Day Calls"].describe()
IQR=data["Day Calls"].quantile(0.75)-data["Day Calls"].quantile(0.25)
IQR
data["Day Calls"].quantile(0.25) - 1.5*IQR
data["Day Calls"].quantile(0.75) + 1.5*IQR
|
notebooks/T1 - 3 - Data Cleaning - Plots.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="d1S1dk6ie07E" executionInfo={"status": "ok", "timestamp": 1616643844403, "user_tz": 300, "elapsed": 740, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}}
# #!/usr/bin/env python3.7
# + colab={"base_uri": "https://localhost:8080/"} id="TXh511SHe2Zb" executionInfo={"status": "ok", "timestamp": 1616643856153, "user_tz": 300, "elapsed": 4389, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="c47558b8-7f1a-480b-ba75-89d5c86bf78c"
# !pip install -U fortran-magic
# + ein.tags="worksheet-0" colab={"base_uri": "https://localhost:8080/", "height": 50} id="7DR24b3hewJ2" executionInfo={"status": "ok", "timestamp": 1616643873900, "user_tz": 300, "elapsed": 1725, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="1e91c29f-8988-4741-dae5-c390584d6c35"
# %matplotlib inline
# %load_ext fortranmagic
import sys; sys.path.append('..')
import pandas as pd
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
import seaborn as sns
mpl.rc('figure', figsize=(12, 7))
ran_the_first_cell = True
jan2017 = pd.to_datetime(['2017-01-03 00:00:00+00:00',
'2017-01-04 00:00:00+00:00',
'2017-01-05 00:00:00+00:00',
'2017-01-06 00:00:00+00:00',
'2017-01-09 00:00:00+00:00',
'2017-01-10 00:00:00+00:00',
'2017-01-11 00:00:00+00:00',
'2017-01-12 00:00:00+00:00',
'2017-01-13 00:00:00+00:00',
'2017-01-17 00:00:00+00:00',
'2017-01-18 00:00:00+00:00',
'2017-01-19 00:00:00+00:00',
'2017-01-20 00:00:00+00:00',
'2017-01-23 00:00:00+00:00',
'2017-01-24 00:00:00+00:00',
'2017-01-25 00:00:00+00:00',
'2017-01-26 00:00:00+00:00',
'2017-01-27 00:00:00+00:00',
'2017-01-30 00:00:00+00:00',
'2017-01-31 00:00:00+00:00',
'2017-02-01 00:00:00+00:00'])
calendar = jan2017.values.astype('datetime64[D]')
event_dates = pd.to_datetime(['2017-01-06 00:00:00+00:00',
'2017-01-07 00:00:00+00:00',
'2017-01-08 00:00:00+00:00']).values.astype('datetime64[D]')
event_values = np.array([10, 15, 20])
# + [markdown] ein.tags="worksheet-0" id="uPT0lbFsewJ-"
# <center>
# <h1>The PyData Toolbox</h1>
# <h3><NAME> (Twitter: @scottbsanderson, GitHub: ssanderson)</h3>
# <h3><a href="https://github.com/ssanderson/pydata-toolbox">https://github.com/ssanderson/pydata-toolbox</a></h3>
# </center>
# + [markdown] id="L0L2fCDuewJ_"
# ## Outline
#
# - Built-in Data Structures
# - Numpy `array`
# - Pandas `Series`/`DataFrame`
# - Plotting and "Real-World" Analyses
# + [markdown] id="hOM4UMypewKA"
# # Data Structures
# + [markdown] id="mV52uEAXewKA"
# > Rule 5. Data dominates. If you've chosen the right data structures and organized things well, the algorithms
# will almost always be self-evident. Data structures, not algorithms, are central to programming.
#
# - *Notes on Programming in C*, by <NAME>.
# + [markdown] id="LxpCFTcnewKA"
# # Lists
# + id="9TvitxHDewKA" executionInfo={"status": "ok", "timestamp": 1616644018856, "user_tz": 300, "elapsed": 450, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}}
assert ran_the_first_cell, "Oh noes!"
# + id="baG-f87zfxVi"
a = 10
b = 5
c = a + b
d = a + b - 1
assert c==15, "Algo salió mal c=15"
# + colab={"base_uri": "https://localhost:8080/"} id="1hsRxszKewKB" executionInfo={"status": "ok", "timestamp": 1616644140918, "user_tz": 300, "elapsed": 463, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="a2072644-3f6b-4164-d9dd-fb1ae805ba58"
l = [1, 'dos', 3.0, 'a', 5.0, "seis"]
l
# + colab={"base_uri": "https://localhost:8080/"} id="da5z_pctewKB" executionInfo={"status": "ok", "timestamp": 1616644162858, "user_tz": 300, "elapsed": 459, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="f672c9c8-3184-4649-ab80-ed73c6322682"
first = l[0]
second = l[1]
print("first:", first)
print("second:", second)
# + colab={"base_uri": "https://localhost:8080/"} id="mTmKER0fewKC" executionInfo={"status": "ok", "timestamp": 1616644164117, "user_tz": 300, "elapsed": 551, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="d1e4bc42-f341-4e19-b25a-33b2872feb16"
last = l[-1]
penultimate = l[-2]
print("last:", last)
print("second to last:", penultimate)
# + colab={"base_uri": "https://localhost:8080/"} id="znN79DrEewKC" executionInfo={"status": "ok", "timestamp": 1616644184935, "user_tz": 300, "elapsed": 1326, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="3f18926f-eb29-4774-ffcd-7cb5c440cb04"
sublist = l[2:5]
sublist
# + colab={"base_uri": "https://localhost:8080/"} id="S52qQT25ewKC" executionInfo={"status": "ok", "timestamp": 1616644193081, "user_tz": 300, "elapsed": 420, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="6f38dbe4-8200-4f77-ae77-b6465f26c45b"
first_three = l[:2]
first_three
# + colab={"base_uri": "https://localhost:8080/"} id="VA1ebFRLewKD" executionInfo={"status": "ok", "timestamp": 1616644204603, "user_tz": 300, "elapsed": 412, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="13b6a288-9fb1-4b17-fd6c-ff9eec1f7608"
after_three = l[4:]
after_three
# + colab={"base_uri": "https://localhost:8080/"} id="yN7B3LYEewKD" executionInfo={"status": "ok", "timestamp": 1616644306024, "user_tz": 300, "elapsed": 419, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="58e48a4e-1bab-4b8c-ddab-515f3f0ae7c8"
l = ['h', 'g', 'f', 'e', 'd', 'c', 'b','a']
l[1:7:2]
# + colab={"base_uri": "https://localhost:8080/"} id="9VA_EXk7ewKD" executionInfo={"status": "ok", "timestamp": 1616644331205, "user_tz": 300, "elapsed": 587, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="9e9779c0-d7ec-4898-d0d9-dbcd0e59d1bf"
l[::-1]
# + colab={"base_uri": "https://localhost:8080/"} id="5WfLk3SaewKE" executionInfo={"status": "ok", "timestamp": 1616644355736, "user_tz": 300, "elapsed": 2023, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="21dc0d40-c380-4345-8932-d21adabc43bb"
l = [1, 2, 3, 4, 5]
print("Before:", l)
l.append('siete')
print("After:", l)
# + colab={"base_uri": "https://localhost:8080/"} id="CZ_lMxspewKE" executionInfo={"status": "ok", "timestamp": 1616644390539, "user_tz": 300, "elapsed": 2098, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="d4049fac-61d8-460d-9077-a499f0c468b1"
l = [1, 2, 3, 4, 5]
[x**2 for x in l]
# + [markdown] ein.tags="worksheet-0" id="VsqTBEd3ewKE"
# ## Review: Python Lists
#
# - Zero-indexed sequence of arbitrary Python values.
# - Slicing syntax: `l[start:stop:step]` copies elements at regular intervals from `start` to `stop`.
# - Efficient (`O(1)`) appends and removes from end.
# - Comprehension syntax: `[f(x) for x in l if cond(x)]`.
# + [markdown] id="QYHiITX8ewKE"
# # Dictionaries
# + colab={"base_uri": "https://localhost:8080/"} id="n84fZDzIewKF" executionInfo={"status": "ok", "timestamp": 1616644435966, "user_tz": 300, "elapsed": 457, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="d656124c-8b2e-4b2f-a04a-aa2b8c2702d9"
philosophers = {'<NAME>': 'Humberto', '<NAME>': 'Carlos', 'Berna': 'Rodrigo'}
philosophers
# + colab={"base_uri": "https://localhost:8080/"} id="Sov-nwZXewKF" executionInfo={"status": "ok", "timestamp": 1616644478805, "user_tz": 300, "elapsed": 503, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="cefc8f08-dbbf-4ceb-c4ea-96e24970654b"
philosophers['Lucho'] = 'Wilson'
philosophers
# + colab={"base_uri": "https://localhost:8080/"} id="zmCJbJv9ewKF" executionInfo={"status": "ok", "timestamp": 1616644506660, "user_tz": 300, "elapsed": 498, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="42680580-df8f-4832-c099-92265598b356"
del philosophers['Ludwig']
philosophers
# + colab={"base_uri": "https://localhost:8080/", "height": 162} id="00k-R9NkewKF" executionInfo={"status": "error", "timestamp": 1616644512207, "user_tz": 300, "elapsed": 458, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="7d75ad65-c175-4c22-966e-e1dbb3678e04"
philosophers['Bertrand':'Immanuel'] #Upps no funciona :c
# + [markdown] ein.tags="worksheet-0" id="pf4esEhtewKG"
# ## Review: Python Dictionaries
#
# - Unordered key-value mapping from (almost) arbitrary keys to arbitrary values.
# - Efficient (`O(1)`) lookup, insertion, and deletion.
# - No slicing (would require a notion of order).
# + id="RmtpC96yewKG" executionInfo={"status": "ok", "timestamp": 1616644594959, "user_tz": 300, "elapsed": 1688, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}}
a = [[10, 20, 30],
[20, 30, 40],
[50, 60, 70],
[10, 10, 10]]
b = [[10, 20, 30, 40],
[20, 30, 40, 50]]
# + id="d33TyqaqewKG" executionInfo={"status": "ok", "timestamp": 1616644598388, "user_tz": 300, "elapsed": 1994, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}}
def matmul(A, B):
"""Multiply matrix A by matrix B."""
rows_out = len(A)
cols_out = len(B[0])
out = [[0 for col in range(cols_out)] for row in range(rows_out)]
for i in range(rows_out):
for j in range(cols_out):
for k in range(len(B)):
out[i][j] += A[i][k] * B[k][j]
return out
# + colab={"base_uri": "https://localhost:8080/"} id="_2Z0lwJAewKH" executionInfo={"status": "ok", "timestamp": 1616644599703, "user_tz": 300, "elapsed": 366, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="e762f85a-23dc-4dc4-e9c1-8a25538967a8"
# %%time
matmul(a, b)
# + colab={"base_uri": "https://localhost:8080/"} id="yCmGWA0bewKH" executionInfo={"status": "ok", "timestamp": 1616644607008, "user_tz": 300, "elapsed": 439, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="10dddb0b-c605-4b74-eb95-78c6a1596d14"
import random
def random_matrix(m, n):
out = []
for row in range(m):
out.append([random.random() for _ in range(n)])
return out
randm = random_matrix(2, 3)
randm
# + colab={"base_uri": "https://localhost:8080/"} id="D0c3WH3IewKH" executionInfo={"status": "ok", "timestamp": 1616644632006, "user_tz": 300, "elapsed": 11348, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="e6363ee8-2217-48e5-f2ff-b2dd1a20b50d"
# %%time
randa = random_matrix(600, 100)
randb = random_matrix(100, 600)
x = matmul(randa, randb)
# + id="9sZjK9mgewKI" executionInfo={"status": "ok", "timestamp": 1616644679502, "user_tz": 300, "elapsed": 457, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}}
def python_dot_product(xs, ys):
return sum(x * y for x, y in zip(xs, ys))
# + id="A6jbz1o8ewKI" executionInfo={"status": "ok", "timestamp": 1616644684582, "user_tz": 300, "elapsed": 2772, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}}
# %%fortran
subroutine fortran_dot_product(xs, ys, result)
double precision, intent(in) :: xs(:)
double precision, intent(in) :: ys(:)
double precision, intent(out) :: result
result = sum(xs * ys)
end
# + id="oNkLKOZEewKI" executionInfo={"status": "ok", "timestamp": 1616644690743, "user_tz": 300, "elapsed": 490, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}}
list_data = [float(i) for i in range(100000)]
array_data = np.array(list_data)
# + colab={"base_uri": "https://localhost:8080/"} id="IrApTzH0ewKI" executionInfo={"status": "ok", "timestamp": 1616644692926, "user_tz": 300, "elapsed": 496, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="3bf3a415-0cd1-4a11-e583-7c87bd79844f"
# %%time
python_dot_product(list_data, list_data)
# + colab={"base_uri": "https://localhost:8080/"} id="OyulAO2AewKM" executionInfo={"status": "ok", "timestamp": 1616644718475, "user_tz": 300, "elapsed": 453, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="62bc0466-7ec8-40c2-901c-9a963683b946"
# %%time
fortran_dot_product(array_data, array_data)
# + [markdown] id="y9NeflnJewKM"
# <center><img src="images/sloth.gif" alt="Drawing" style="width: 1080px;"/></center>
#
# + [markdown] id="G0uE1bKEewKN"
# ## Why is the Python Version so Much Slower?
# + colab={"base_uri": "https://localhost:8080/"} id="hj4BF6O6ewKN" executionInfo={"status": "ok", "timestamp": 1616644724145, "user_tz": 300, "elapsed": 443, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="74025aaa-0a9e-4e36-e77b-2812e1106025"
def mul_elemwise(xs, ys):
return [x * y for x, y in zip(xs, ys)]
mul_elemwise([1, 2, 3, 4], [1, 2 + 0j, 3.0, 'four'])
# + colab={"base_uri": "https://localhost:8080/"} id="sY6_9Qg2ewKN" executionInfo={"status": "ok", "timestamp": 1616644726329, "user_tz": 300, "elapsed": 453, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="6451f8dc-9825-4f7c-e463-7e86ad87abea"
source_code = 'a + b * c'
bytecode = compile(source_code, '', 'eval')
import dis; dis.dis(bytecode)
# + [markdown] id="o2YxTY0KewKN"
# ## Why is the Python Version so Slow?
# - Dynamic typing means that every single operation requires dispatching on the input type.
# - Having an interpreter means that every instruction is fetched and dispatched at runtime.
# - Other overheads:
# - Arbitrary-size integers.
# - Reference-counted garbage collection.
# + [markdown] id="wMZDbYeEewKO"
# > This is the paradox that we have to work with when we're doing scientific or numerically-intensive Python. What makes Python fast for development -- this high-level, interpreted, and dynamically-typed aspect of the language -- is exactly what makes it slow for code execution.
#
# - <NAME>, [*Losing Your Loops: Fast Numerical Computing with NumPy*](https://www.youtube.com/watch?v=EEUXKG97YRw)
# + [markdown] id="3ds45C8yewKO"
# # What Do We Do?
# + [markdown] id="pyZcSdKRewKP"
# - Python is slow for numerical computation because it performs dynamic dispatch on every operation we perform...
# + [markdown] id="I9wpcdU8ewKP"
# - ...but often, we just want to do the same thing over and over in a loop!
# + [markdown] id="cm3NhRTNewKP"
# - If we don't need Python's dynamicism, we don't want to pay (much) for it.
# + [markdown] id="g21J9MsrewKP"
# - **Idea:** Dispatch **once per operation** instead of **once per element**.
# + colab={"base_uri": "https://localhost:8080/"} id="DOUAFMxwewKP" executionInfo={"status": "ok", "timestamp": 1616644756312, "user_tz": 300, "elapsed": 469, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="5f0e2984-92d9-4a5b-af8b-85275b633b00"
import numpy as np
data = np.array([1, 2, 3, 4])
data
# + colab={"base_uri": "https://localhost:8080/"} id="icQ0LlA4ewKQ" executionInfo={"status": "ok", "timestamp": 1616644757327, "user_tz": 300, "elapsed": 490, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="901ff846-2c1d-43c5-c36f-7de00003d0db"
data + data
# + colab={"base_uri": "https://localhost:8080/"} id="FOHhQoG8ewKQ" executionInfo={"status": "ok", "timestamp": 1616644757809, "user_tz": 300, "elapsed": 382, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="a3db1fad-a543-40c7-c990-1c8b26934c50"
# %%time
(array_data * array_data).sum()
# + colab={"base_uri": "https://localhost:8080/"} id="VWiPzZZ4ewKQ" executionInfo={"status": "ok", "timestamp": 1616644765259, "user_tz": 300, "elapsed": 414, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="c83e99c2-4502-4e88-d371-5557308d1d73"
# %%time
array_data.dot(array_data)
# + colab={"base_uri": "https://localhost:8080/"} id="X7XstGWbewKR" executionInfo={"status": "ok", "timestamp": 1616644769670, "user_tz": 300, "elapsed": 713, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="1fddb2cc-582d-4f46-9690-5d4eeae63d6b"
# %%time
fortran_dot_product(array_data, array_data)
# + colab={"base_uri": "https://localhost:8080/"} id="6f3jCjmkewKS" executionInfo={"status": "ok", "timestamp": 1616644795993, "user_tz": 300, "elapsed": 432, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="70b5d91e-e0c6-4c31-ef8b-5ed3fa2de670"
two_by_two = data.reshape(2, 2)
two_by_two
# + [markdown] id="JAa81oOqewKS"
# Numpy arrays are:
#
# - Fixed-type
# + [markdown] id="f7QTnb1qewKS"
# - Size-immutable
# + [markdown] id="Cd9c0zNNewKS"
# - Multi-dimensional
# + [markdown] id="Opmwj7puewKS"
# - Fast\*
# + [markdown] id="-SLCqXTTewKS"
# \* If you use them correctly.
# + [markdown] id="lxJiIgrfewKT"
# # What's in an Array?
# + colab={"base_uri": "https://localhost:8080/"} id="5sMQLRVJewKT" executionInfo={"status": "ok", "timestamp": 1616644802480, "user_tz": 300, "elapsed": 1456, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="0d78dc16-9b45-4b30-cf61-f638b97a8b7d"
arr = np.array([1, 2, 3, 4, 5, 6], dtype='int16').reshape(2, 3)
print("Array:\n", arr, sep='')
print("===========")
print("DType:", arr.dtype)
print("Shape:", arr.shape)
print("Strides:", arr.strides)
print("Data:", arr.data.tobytes())
# + [markdown] id="oM8ziW6-ewKT"
# # Core Operations
#
# - Vectorized **ufuncs** for elementwise operations.
# - Fancy indexing and masking for selection and filtering.
# - Aggregations across axes.
# - Broadcasting
# + [markdown] id="gBe5a57mewKT"
# # UFuncs
#
# UFuncs (universal functions) are functions that operate elementwise on one or more arrays.
# + colab={"base_uri": "https://localhost:8080/"} id="GeJvA-CXewKT" executionInfo={"status": "ok", "timestamp": 1616644808846, "user_tz": 300, "elapsed": 600, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="551d0cec-ccd6-4d60-b9e9-d215ed1361c2"
data = np.arange(15).reshape(3, 5)
data
# + colab={"base_uri": "https://localhost:8080/"} id="yZGOonV3ewKU" executionInfo={"status": "ok", "timestamp": 1616644809731, "user_tz": 300, "elapsed": 430, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="35d49d8f-c0be-4087-a2f0-20994223e78f"
data * data
# + colab={"base_uri": "https://localhost:8080/"} id="cdxdTrEQewKU" executionInfo={"status": "ok", "timestamp": 1616644816073, "user_tz": 300, "elapsed": 609, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="4085af8a-b661-4432-f5c0-5df6c706bde3"
np.sqrt(data)
# + colab={"base_uri": "https://localhost:8080/"} id="tSgjwQjDewKU" executionInfo={"status": "ok", "timestamp": 1616644819258, "user_tz": 300, "elapsed": 713, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="ebb17da8-3a7b-40dd-a228-1df7c9851e34"
(data % 3) == 0
# + colab={"base_uri": "https://localhost:8080/"} id="CvDf2bpJewKU" executionInfo={"status": "ok", "timestamp": 1616644821038, "user_tz": 300, "elapsed": 455, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="3bc5eb0d-007e-4ab6-a924-bec6dd6020c0"
((data % 2) == 0) & ((data % 3) == 0)
# + colab={"base_uri": "https://localhost:8080/"} id="pCcuPV5LewKV" executionInfo={"status": "ok", "timestamp": 1616644824545, "user_tz": 300, "elapsed": 838, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="e247d7d1-1664-453b-842d-7a32421e9d79"
data @ data.T
# + [markdown] id="AOXNMWRfewKV"
# # UFuncs Review
#
# - UFuncs provide efficient elementwise operations applied across one or more arrays.
# - Arithmetic Operators (`+`, `*`, `/`)
# - Comparisons (`==`, `>`, `!=`)
# - Boolean Operators (`&`, `|`, `^`)
# - Trigonometric Functions (`sin`, `cos`)
# - Transcendental Functions (`exp`, `log`)
# + [markdown] id="zNSLFNVMewKV"
# # Selections
# + [markdown] id="YTAO28-QewKW"
# We often want to perform an operation on just a subset of our data.
# + colab={"base_uri": "https://localhost:8080/"} id="pt6tya9AewKW" executionInfo={"status": "ok", "timestamp": 1616644830205, "user_tz": 300, "elapsed": 479, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="22286e74-162f-4ee6-ca27-7578066d45e7"
sines = np.sin(np.linspace(0, 3.14, 10))
cosines = np.cos(np.linspace(0, 3.14, 10))
sines
# + colab={"base_uri": "https://localhost:8080/"} id="t09KpHX4ewKW" executionInfo={"status": "ok", "timestamp": 1616644840284, "user_tz": 300, "elapsed": 504, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="d99e3b3f-a4a6-47a4-c653-a0000a9cf67b"
sines[2]
# + colab={"base_uri": "https://localhost:8080/"} id="MGTNYZmZewKW" executionInfo={"status": "ok", "timestamp": 1616644846335, "user_tz": 300, "elapsed": 614, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="aa575585-7558-47f2-b32a-e55ff390e700"
sines[:6]
# + colab={"base_uri": "https://localhost:8080/"} id="Zq86Kp_yewKX" executionInfo={"status": "ok", "timestamp": 1616644850065, "user_tz": 300, "elapsed": 440, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="d17a4c86-fb0d-470c-9cfe-93f93d8b1e54"
sines[2:]
# + colab={"base_uri": "https://localhost:8080/"} id="lL0JLCHCewKX" executionInfo={"status": "ok", "timestamp": 1616644854280, "user_tz": 300, "elapsed": 787, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="48b51800-29ea-4171-802f-a9da1846be75"
sines[::8]
# + id="AO2nni8dewKX" outputId="ed700597-d975-45c0-d9a0-f68bc3a26499"
print("sines:\n", sines)
print("sines > 0.5:\n", sines > 0.5)
print("sines[sines > 0.5]:\n", sines[sines > 0.5])
# + colab={"base_uri": "https://localhost:8080/"} id="7iKpUBsdewKX" executionInfo={"status": "ok", "timestamp": 1616644871133, "user_tz": 300, "elapsed": 448, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="84cfb894-d2af-42a8-df37-d0b9573becfd"
print(sines)
sines[[0, 4, 7]]
# + id="PyqOkpegewKY" executionInfo={"status": "ok", "timestamp": 1616644874092, "user_tz": 300, "elapsed": 296, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}}
unsorted_data = np.array([1, 3, 2, 12, -1, 5, 2])
# + colab={"base_uri": "https://localhost:8080/"} id="dK7Jcs7JewKY" executionInfo={"status": "ok", "timestamp": 1616644882169, "user_tz": 300, "elapsed": 433, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="7d3f82d9-4581-4622-d896-33bc2ab43473"
sort_indices = np.argsort(unsorted_data)
sort_indices
# + colab={"base_uri": "https://localhost:8080/"} id="RIE4AFjaewKY" executionInfo={"status": "ok", "timestamp": 1616644883211, "user_tz": 300, "elapsed": 438, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="27382622-bb8d-470a-eaf0-4204c6699a49"
unsorted_data[sort_indices]
# + id="A2Bq6QfJewKY" executionInfo={"status": "ok", "timestamp": 1616644909026, "user_tz": 300, "elapsed": 444, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}}
market_caps = np.array([2, 7, 12, 15, 14])
assets = np.array(['E', 'D', 'C', 'B', 'A'])
# + colab={"base_uri": "https://localhost:8080/"} id="hqHQ1VUOewKZ" executionInfo={"status": "ok", "timestamp": 1616644913264, "user_tz": 300, "elapsed": 402, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="8b4ad854-1e46-431a-f61b-9dec407b50d8"
sort_by_mcap = np.argsort(market_caps)
assets[sort_by_mcap]
# + colab={"base_uri": "https://localhost:8080/"} id="vuYxSmWYewKZ" executionInfo={"status": "ok", "timestamp": 1616644916231, "user_tz": 300, "elapsed": 439, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="74f4c5b9-7e5e-4a49-f2cc-18bee01d27fe"
print("Dates:\n", repr(event_dates))
print("Values:\n", repr(event_values))
print("Calendar:\n", repr(calendar))
# + colab={"base_uri": "https://localhost:8080/"} id="-O0Sw72xewKZ" executionInfo={"status": "ok", "timestamp": 1616644919793, "user_tz": 300, "elapsed": 581, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="9fb19f96-f736-4f12-e4cd-7b5f0bf6a9f7"
print("Raw Dates:", event_dates)
print("Indices:", calendar.searchsorted(event_dates))
print("Forward-Filled Dates:", calendar[calendar.searchsorted(event_dates)])
# + [markdown] id="CsCuaP6xewKZ"
# On multi-dimensional arrays, we can slice along each axis independently.
# + colab={"base_uri": "https://localhost:8080/"} id="olwP1smBewKZ" executionInfo={"status": "ok", "timestamp": 1616644921179, "user_tz": 300, "elapsed": 591, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="12331800-0536-4611-f96e-ff479c5961d5"
data = np.arange(25).reshape(5, 5)
data
# + colab={"base_uri": "https://localhost:8080/"} id="FMvp5WQWewKa" executionInfo={"status": "ok", "timestamp": 1616644928461, "user_tz": 300, "elapsed": 1719, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="8477a6a6-c0d2-466b-8875-b98081bc0e6a"
data[:3, :3]
# + colab={"base_uri": "https://localhost:8080/"} id="Vhibso3bewKa" executionInfo={"status": "ok", "timestamp": 1616644938151, "user_tz": 300, "elapsed": 639, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="81aefa1d-7dfd-4265-9213-fbc560313b92"
data[:4, [1, -1]]
# + colab={"base_uri": "https://localhost:8080/"} id="jcr50RLPewKa" executionInfo={"status": "ok", "timestamp": 1616644950172, "user_tz": 300, "elapsed": 505, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="4620bbde-1614-46ae-a5e9-c4bf56c58240"
data[(data[:, 1] % 3) == 0]
# + [markdown] id="VIudSOfAewKb"
# # Selections Review
#
# - Indexing with an integer removes a dimension.
# - Slicing operations work on Numpy arrays the same way they do on lists.
# - Indexing with a boolean array filters to True locations.
# - Indexing with an integer array selects indices along an axis.
# - Multidimensional arrays can apply selections independently along different axes.
# + [markdown] id="jsc5jtv4ewKb"
# ## Reductions
#
# Functions that reduce an array to a scalar.
# + [markdown] id="12FAOT1AewKb"
# $$Var(X) = \frac{1}{N}\sqrt{\sum_{i=1}^N (x_i - \bar{x})^2}$$
# + id="nmsuCodRewKb" executionInfo={"status": "ok", "timestamp": 1616645018297, "user_tz": 300, "elapsed": 450, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}}
def variance(x):
return ((x - x.mean()) ** 2).sum() / len(x)
# + colab={"base_uri": "https://localhost:8080/"} id="vBE0kgvlewKb" executionInfo={"status": "ok", "timestamp": 1616645022911, "user_tz": 300, "elapsed": 485, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="58f5dc4e-89d5-4d16-e8d2-587e660f25dd"
variance(np.random.standard_normal(100))
# + [markdown] id="v9oQPcFQewKc"
# - `sum()` and `mean()` are both **reductions**.
# + [markdown] id="at2SFtlYewKc"
# - In the simplest case, we use these to reduce an entire array into a single value...
# + colab={"base_uri": "https://localhost:8080/"} id="New-Kkq3ewKc" executionInfo={"status": "ok", "timestamp": 1616645040533, "user_tz": 300, "elapsed": 499, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="dc1cf744-8b11-4da1-8ef5-d383b775304d"
data = np.arange(25)
data.mean()
# + [markdown] id="jBnz0p2VewKc"
# - ...but we can do more interesting things with multi-dimensional arrays.
# + colab={"base_uri": "https://localhost:8080/"} id="13ENc1c-ewKc" executionInfo={"status": "ok", "timestamp": 1616645042536, "user_tz": 300, "elapsed": 397, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="6b2cf81f-b1f7-4964-af56-3960ba3febc4"
data = np.arange(30).reshape(3, 10)
data
# + colab={"base_uri": "https://localhost:8080/"} id="lHL5XAb4ewKd" executionInfo={"status": "ok", "timestamp": 1616645043599, "user_tz": 300, "elapsed": 362, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="f13140f9-b22e-4a1a-db07-eaba77b68dc7"
data.mean()
# + colab={"base_uri": "https://localhost:8080/"} id="w8QqwhDDewKd" executionInfo={"status": "ok", "timestamp": 1616645048396, "user_tz": 300, "elapsed": 645, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="4c8f98ea-fa80-4776-9836-8a7039ab6ccb"
data.mean(axis=0)
# + colab={"base_uri": "https://localhost:8080/"} id="joll3yZJewKd" executionInfo={"status": "ok", "timestamp": 1616645049257, "user_tz": 300, "elapsed": 464, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="eeb89dd8-9ca5-405e-8de8-c11aac00ca3c"
data.mean(axis=1)
# + [markdown] id="LBsjwj0NewKd"
# ## Reductions Review
#
# - Reductions allow us to perform efficient aggregations over arrays.
# - We can do aggregations over a single axis to collapse a single dimension.
# - Many built-in reductions (`mean`, `sum`, `min`, `max`, `median`, ...).
# + [markdown] id="ftCc2eZdewKe"
# # Broadcasting
# + colab={"base_uri": "https://localhost:8080/"} id="pVxe2VpyewKe" executionInfo={"status": "ok", "timestamp": 1616645054532, "user_tz": 300, "elapsed": 683, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="e0676781-1a59-454d-c94e-ba59927a712b"
row = np.array([1, 2, 3, 4])
column = np.array([[1], [2], [3]])
print("Row:\n", row, sep='')
print("Column:\n", column, sep='')
# + colab={"base_uri": "https://localhost:8080/"} id="pB6S48A8ewKe" executionInfo={"status": "ok", "timestamp": 1616645060427, "user_tz": 300, "elapsed": 1462, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="8c70b49c-d642-4216-a427-0dfd5ced52f2"
row + column
# + colab={"base_uri": "https://localhost:8080/"} id="tm8YLcGPewKf" executionInfo={"status": "ok", "timestamp": 1616645082666, "user_tz": 300, "elapsed": 475, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="5e8ceefa-8287-42f6-c223-69bac6f7ff37"
print("Data:\n", data, sep='')
print("Mean:\n", data.mean(axis=0), sep='')
print("Data - Mean:\n", data - data.mean(axis=0), sep='')
# + [markdown] id="p2yd5CpPewKf"
# # Broadcasting Review
#
# - Numpy operations can work on arrays of different dimensions as long as the arrays' shapes are still "compatible".
# - Broadcasting works by "tiling" the smaller array along the missing dimension.
# - The result of a broadcasted operation is always at least as large in each dimension as the largest array in that dimension.
# + [markdown] id="mNyzLdsnewKf"
# # Numpy Review
# + [markdown] id="2aZOsvr-ewKf"
# - Numerical algorithms are slow in pure Python because the overhead dynamic dispatch dominates our runtime.
# + [markdown] id="Al8v22bmewKf"
# - Numpy solves this problem by:
# 1. Imposing additional restrictions on the contents of arrays.
# 2. Moving the inner loops of our algorithms into compiled C code.
# + [markdown] id="Ctq-qD7qewKg"
# - Using Numpy effectively often requires reworking an algorithms to use vectorized operations instead of for-loops, but the resulting operations are usually simpler, clearer, and faster than the pure Python equivalent.
# + [markdown] id="z75FvRYrewKg"
# Numpy is great for many things, but...
# + [markdown] id="-lJ3pSmXewKg"
# - Sometimes our data is equipped with a natural set of **labels**:
# - Dates/Times
# - Stock Tickers
# - Field Names (e.g. Open/High/Low/Close)
# + [markdown] id="qmFmtBpbewKg"
# - Sometimes we have **more than one type of data** that we want to keep grouped together.
# - Tables with a mix of real-valued and categorical data.
# + [markdown] id="cwfXzivfewKg"
# - Sometimes we have **missing** data, which we need to ignore, fill, or otherwise work around.
# + [markdown] id="WeHHqcFSewKh"
# Pandas extends Numpy with more complex data structures:
#
# - `Series`: 1-dimensional, homogenously-typed, labelled array.
# - `DataFrame`: 2-dimensional, semi-homogenous, labelled table.
# + [markdown] id="rHVUAwjrewKh"
# Pandas also provides many utilities for:
# - Input/Output
# - Data Cleaning
# - Rolling Algorithms
# - Plotting
# + [markdown] id="IrENp8qOewKh"
# # Selection in Pandas
# + colab={"base_uri": "https://localhost:8080/"} id="ntt6fl9vewKi" executionInfo={"status": "ok", "timestamp": 1616645101399, "user_tz": 300, "elapsed": 583, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="b4dbf32c-6bfa-4d52-c03b-05fa310c650b"
s = pd.Series(index=['a', 'b', 'c', 'd', 'e'], data=[1, 2, 3, 4, 5])
s
# + colab={"base_uri": "https://localhost:8080/"} id="JibdzjFMewKi" executionInfo={"status": "ok", "timestamp": 1616645129322, "user_tz": 300, "elapsed": 2085, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="ed25cb79-6005-44cb-bbc2-112f5d08df09"
print("Los Indices son:", s.index)
print("Los Valores son:", s.values)
# + colab={"base_uri": "https://localhost:8080/"} id="-hndCqUHewKi" executionInfo={"status": "ok", "timestamp": 1616645133364, "user_tz": 300, "elapsed": 1165, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="a02038ec-9010-452d-ffdf-3da32460e863"
s.iloc[0]
# + colab={"base_uri": "https://localhost:8080/"} id="0Msp_CpWewKi" executionInfo={"status": "ok", "timestamp": 1616645138137, "user_tz": 300, "elapsed": 465, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="660aac38-b545-48e5-eed5-4cf2772b7e16"
s.loc['a']
# + colab={"base_uri": "https://localhost:8080/"} id="yrbHhfRaewKj" executionInfo={"status": "ok", "timestamp": 1616645151648, "user_tz": 300, "elapsed": 473, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="a75b42d2-c7b9-4184-ccd1-39f41d90fa05"
s.iloc[:2]
# + colab={"base_uri": "https://localhost:8080/"} id="_rqP8nsFewKj" executionInfo={"status": "ok", "timestamp": 1616645160418, "user_tz": 300, "elapsed": 2610, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="64516809-71aa-4f48-80b2-03cf4e10b4af"
s.loc[:'c']
# + colab={"base_uri": "https://localhost:8080/"} id="Cd4XI64VewKj" executionInfo={"status": "ok", "timestamp": 1616645163433, "user_tz": 300, "elapsed": 561, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="4ce0000f-4715-40bc-a1b6-a07349e73f90"
s.iloc[[0, -1]]
# + colab={"base_uri": "https://localhost:8080/"} id="jUW-3PUFewKj" executionInfo={"status": "ok", "timestamp": 1616645167874, "user_tz": 300, "elapsed": 449, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="89ae8002-1b5c-4da5-b2a2-d1975d81cb34"
s.loc[s > 2]
# + colab={"base_uri": "https://localhost:8080/"} id="45kGBrTiewKk" executionInfo={"status": "ok", "timestamp": 1616645186542, "user_tz": 300, "elapsed": 489, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="425ca730-d714-4823-f76e-dd7661f72f85"
other_s = pd.Series({'A': 10.0, 'C': 20.0, 'D': 30.0, 'Z': 40.0})
other_s
# + colab={"base_uri": "https://localhost:8080/"} id="Hv-Sri7bewKk" executionInfo={"status": "ok", "timestamp": 1616645190760, "user_tz": 300, "elapsed": 1272, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="2b0c461f-b88d-4463-d170-2f5e2a312b34"
s + other_s
# + colab={"base_uri": "https://localhost:8080/"} id="IssM5D3wewKk" executionInfo={"status": "ok", "timestamp": 1616645198221, "user_tz": 300, "elapsed": 401, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="71d6175c-2d42-4d71-f4c6-64115e6c7fb5"
(s + other_s).fillna(0.0)
# + colab={"base_uri": "https://localhost:8080/"} id="G2NuULKskQfc" executionInfo={"status": "ok", "timestamp": 1616645472241, "user_tz": 300, "elapsed": 28653, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="965a10fe-2f0f-4749-cf6c-27ea6bc84d2f"
from google.colab import drive
drive.mount('/content/drive/')
# + colab={"base_uri": "https://localhost:8080/", "height": 225} id="ZKpYA8BLewKk" executionInfo={"status": "ok", "timestamp": 1616645516188, "user_tz": 300, "elapsed": 772, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="5655f6e4-bac4-4625-c958-df43b5d89582"
aapl = pd.read_csv('/content/drive/MyDrive/Notebooks de Métodos Numéricos/Lab 2/AAPL.csv', parse_dates=['Date'], index_col='Date')
aapl.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 136} id="q3Qt-WpIewKl" executionInfo={"status": "ok", "timestamp": 1616645530124, "user_tz": 300, "elapsed": 1139, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="27b98a76-cc48-464a-d852-26a87638b981"
aapl.iloc[:2, :2]
# + colab={"base_uri": "https://localhost:8080/", "height": 195} id="HDIE4L-VewKl" executionInfo={"status": "ok", "timestamp": 1616645536340, "user_tz": 300, "elapsed": 451, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="8aabad16-dd51-4949-8922-a5b5e6ffee14"
aapl.loc[pd.Timestamp('2010-02-01'):pd.Timestamp('2010-02-04'), ['Close', 'Volume']]
# + [markdown] id="SiRhDiSOewKl"
# # Rolling Operations
# + colab={"base_uri": "https://localhost:8080/", "height": 417} id="vQ2wMNoKewKm" executionInfo={"status": "ok", "timestamp": 1616645544304, "user_tz": 300, "elapsed": 948, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="52d559c0-dc32-4f46-b198-b974e24d0034"
aapl.rolling(5)[['Close', 'Adj Close']].mean().plot();
# + colab={"base_uri": "https://localhost:8080/", "height": 442} id="UXqAH9aZewKm" executionInfo={"status": "ok", "timestamp": 1616645549513, "user_tz": 300, "elapsed": 927, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="99405fe7-a741-4fea-ce2f-8a67473567a2"
aapl.drop('Volume', axis=1).resample('2W').max().plot();
# + colab={"base_uri": "https://localhost:8080/", "height": 416} id="pVQp7HoaewKm" executionInfo={"status": "ok", "timestamp": 1616645557396, "user_tz": 300, "elapsed": 3068, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="895f41af-1228-4491-d912-2178f1c4580b"
aapl['Close'].pct_change().ewm(span=30).std().plot();
# + [markdown] id="gqQLe1R3ewKm"
# # "Real World" Data
# + colab={"base_uri": "https://localhost:8080/", "height": 195} id="el03o8A3ewKn" executionInfo={"status": "ok", "timestamp": 1616645891737, "user_tz": 300, "elapsed": 4393, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="f24e26a5-463e-4110-cec0-ad0a39baf7a7"
import sys
sys.path.append('/content/drive/MyDrive/Notebooks de Métodos Numéricos/Lab 2/demos')
from avocados import read_avocadata
avocados = read_avocadata('2014', '2016')
avocados.head()
# + colab={"base_uri": "https://localhost:8080/"} id="alRABilfewKn" executionInfo={"status": "ok", "timestamp": 1616645897986, "user_tz": 300, "elapsed": 476, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="57372bc5-521a-443f-f593-dcd66f11219f"
avocados.dtypes
# + colab={"base_uri": "https://localhost:8080/", "height": 424} id="PPIjIujjewKn" executionInfo={"status": "ok", "timestamp": 1616645904788, "user_tz": 300, "elapsed": 1966, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="4501989c-c275-4f97-caed-678d68dcf24e"
hass = avocados[avocados.Variety == 'HASS']
hass.groupby(['Date', 'Region'])['Weighted Avg Price'].mean().unstack().ffill().plot();
# + id="zGzou3IeewKn" executionInfo={"status": "ok", "timestamp": 1616645911120, "user_tz": 300, "elapsed": 478, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}}
def _organic_spread(group):
if len(group.columns) != 2:
return pd.Series(index=group.index, data=0.0)
is_organic = group.columns.get_level_values('Organic').values.astype(bool)
organics = group.loc[:, is_organic].squeeze()
non_organics = group.loc[:, ~is_organic].squeeze()
diff = organics - non_organics
return diff
def organic_spread_by_region(df):
"""What's the difference between the price of an organic
and non-organic avocado within each region?
"""
return (
df
.set_index(['Date', 'Region', 'Organic'])
['Weighted Avg Price']
.unstack(level=['Region', 'Organic'])
.ffill()
.groupby(level='Region', axis=1)
.apply(_organic_spread)
)
# + colab={"base_uri": "https://localhost:8080/", "height": 440} id="0IE07Mh7ewKo" executionInfo={"status": "ok", "timestamp": 1616645915256, "user_tz": 300, "elapsed": 1962, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="4194b328-4159-4779-b6eb-4b86274fd6cd"
organic_spread_by_region(hass).plot();
plt.gca().set_title("Daily Regional Organic Spread");
plt.legend(bbox_to_anchor=(1, 1));
# + colab={"base_uri": "https://localhost:8080/", "height": 343} id="VFYHViXMewKo" executionInfo={"status": "ok", "timestamp": 1616645918341, "user_tz": 300, "elapsed": 460, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="fa5abcf0-fb11-4553-86f3-55f52e163b11"
spread_correlation = organic_spread_by_region(hass).corr()
spread_correlation
# + colab={"base_uri": "https://localhost:8080/", "height": 712} id="LJKE6A-AewKo" executionInfo={"status": "ok", "timestamp": 1616645923497, "user_tz": 300, "elapsed": 1568, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiwFNcAkUDPfjz-UOdCxnj_SMO03iKNKbuGMh1l2Q=s64", "userId": "07981164818721997713"}} outputId="a675d495-1dc3-4fa7-b550-72df254df12c"
import seaborn as sns
grid = sns.clustermap(spread_correlation, annot=True)
fig = grid.fig
axes = fig.axes
ax = axes[2]
ax.set_xticklabels(ax.get_xticklabels(), rotation=45);
# + [markdown] id="KcamdQDdewKo"
# # Pandas Review
#
# - Pandas extends numpy with more complex datastructures and algorithms.
# - If you understand numpy, you understand 90% of pandas.
# - `groupby`, `set_index`, and `unstack` are powerful tools for working with categorical data.
# - Avocado prices are surprisingly interesting :)
# + [markdown] id="B94JypMlewKp"
# # Thanks!
|
Lab2/crvargasmscottsanderson.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Bite Size Bayes
#
# Copyright 2020 <NAME>
#
# License: [Attribution-NonCommercial-ShareAlike 4.0 International (CC BY-NC-SA 4.0)](https://creativecommons.org/licenses/by-nc-sa/4.0/)
# ## 101 Bowls
#
# In [Notebook 4](https://colab.research.google.com/github/AllenDowney/BiteSizeBayes/blob/master/05_dice.ipynb), we saw that the Bayes table works with more than two hypotheses. As an example, we solved a cookie problem with five bowls.
#
# Now we'll take it even farther and solve a cookie problem with 101 bowls:
#
# * Bowl 0 contains no vanilla cookies,
#
# * Bowl 1 contains 1% vanilla cookies,
#
# * Bowl 2 contains 2% vanilla cookies,
#
# and so on, up to
#
# * Bowl 99 contains 99% vanilla cookies, and
#
# * Bowl 100 contains all vanilla cookies.
#
# As in the previous problems, there are only two kinds of cookies, vanilla and chocolate. So Bowl 0 is all chocolate cookies, Bowl 1 is 99% chocolate, and so on.
#
# Suppose we choose a bowl at random, choose a cookie at random, and it turns out to be vanilla. What is the probability that the cookie came from Bowl $x$, for each value of $x$?
#
# To solve this problem, I'll use `np.arange` to represent 101 hypotheses, numbered from 0 to 100.
# ## Review
#
# [In the previous notebook](https://colab.research.google.com/github/AllenDowney/BiteSizeBayes/blob/master/05_test.ipynb) we used a Bayes table to interpret medical tests.
#
# In this notebook we'll solve an expanded version of the cookie problem with 101 Bowls. It might seem like a silly problem, but it's not: the solution demonstrates a Bayesian way to estimate a proportion, and it applies to lots of real problems that don't involve cookies.
#
# Then I'll introduce an alternative to the Bayes table, a probability mass function (PMF), which is a useful way to represent and do computations with distributions.
#
# Here's the function, from the previous notebook, we'll use to make Bayes tables:
# +
import pandas as pd
def make_bayes_table(hypos, prior, likelihood):
"""Make a Bayes table.
hypos: sequence of hypotheses
prior: prior probabilities
likelihood: sequence of likelihoods
returns: DataFrame
"""
table = pd.DataFrame(index=hypos)
table['prior'] = prior
table['likelihood'] = likelihood
table['unnorm'] = table['prior'] * table['likelihood']
prob_data = table['unnorm'].sum()
table['posterior'] = table['unnorm'] / prob_data
return table
# +
import numpy as np
xs = np.arange(101)
# -
# The prior probability for each bowl is $1/101$. I could create a sequence with 101 identical values, but if all of the priors are equal, we only have to probide one value:
prior = 1/101
# Because of the way I numbered the bowls, the probability of a vanilla cookie from Bowl $x$ is $x/100$. So we can compute the likelihoods like this:
likelihood = xs/100
# And that's all we need; the Bayes table does the rest:
table = make_bayes_table(xs, prior, likelihood)
# Here's a feature we have not seen before: we can give the index of the Bayes table a name, which will appear when we display the table.
table.index.name = 'Bowl'
# Here are the first few rows:
table.head()
# Because Bowl 0 contains no vanilla cookies, its likelihood is 0, so its posterior probability is 0. That is, the cookie cannot have come from Bowl 0.
#
# Here are the last few rows of the table.
table.tail()
# The posterior probabilities are substantially higher for the high-numbered bowls.
#
# There is a pattern here that will be clearer if we plot the results.
# +
import matplotlib.pyplot as plt
def plot_table(table):
"""Plot results from the 101 Bowls problem.
table: DataFrame representing a Bayes table
"""
table['prior'].plot()
table['posterior'].plot()
plt.xlabel('Bowl #')
plt.ylabel('Probability')
plt.legend()
# -
plot_table(table)
plt.title('One cookie');
# The prior probabilities are uniform; that is, they are the same for every bowl.
#
# The posterior probabilities increase linearly; Bowl 0 is the least likely (actually impossible), and Bowl 100 is the most likely.
# ## Two cookies
#
# Suppose we put the first cookie back, stir the bowl thoroughly, and draw another cookie from the same bowl. and suppose it turns out to be another vanilla cookie.
#
# Now what is the probability that we are drawing from Bowl $x$?
#
# To answer this question, we can use the posterior probabilities from the previous problem as prior probabilities for a new Bayes table, and then update with the new data.
# +
prior2 = table['posterior']
likelihood2 = likelihood
table2 = make_bayes_table(xs, prior2, likelihood2)
plot_table(table2)
plt.title('Two cookies');
# -
# The blue line shows the posterior after one cookie, which is the prior before the second cookie.
#
# The orange line shows the posterior after two cookies, which curves upward. Having see two vanilla cookies, the high-numbered bowls are more likely; the low-numbered bowls are less likely.
#
# I bet you can guess what's coming next.
# ## Three cookies
#
# Suppose we put the cookie back, stir, draw another cookie from the same bowl, and get a chocolate cookie.
#
# What do you think the posterior distribution looks like after these three cookies?
#
# Hint: what's the probability that the chocolate cookie came from Bowl 100?
#
# We'll use the posterior after two cookies as the prior for the third cookie:
prior3 = table2['posterior']
# Now, what about the likelihoods? Remember that the probability of a vanilla cookie from Bowl $x$ is $x/100$. So the probability of a chocolate cookie is $(1 - x/100)$, which we can compute like this.
likelihood3 = 1 - xs/100
# That's it. Everything else is the same.
table3 = make_bayes_table(xs, prior3, likelihood3)
# And here are the results
plot_table(table3)
plt.title('Three cookies');
# The blue line is the posterior after two cookies; the orange line is the posterior after three cookies.
#
# Because Bowl 100 contains no chocolate cookies, the posterior probability for Bowl 100 is 0.
#
# The posterior distribution has a peak near 60%. We can use `idxmax` to find it:
table3['posterior'].idxmax()
# The peak in the posterior distribution is at 67%.
#
# This value has a name; it is the **MAP**, which stands for "Maximum Aposteori Probability" ("aposteori" is Latin for posterior).
#
# In this example, the MAP is close to the proportion of vanilla cookies in the dataset: 2/3.
# **Exercise:** Let's do a version of the dice problem where we roll the die more than once. Here's the statement of the problem again:
#
# > Suppose you have a 4-sided, 6-sided, 8-sided, and 12-sided die. You choose one at random, roll it and get a 1. What is the probability that the die you rolled is 4-sided? What are the posterior probabilities for the other dice?
#
# And here's a solution using a Bayes table:
# +
hypos = ['H4', 'H6', 'H8', 'H12']
prior = 1/4
likelihood = 1/4, 1/6, 1/8, 1/12
table = make_bayes_table(hypos, prior, likelihood)
table
# -
# Now suppose you roll the same die again and get a 6. What are the posterior probabilities after the second roll?
#
# Use `idxmax` to find the MAP.
prior2 = table['posterior']
likelihood2 = 0, 1/6, 1/8, 1/12
table2 = make_bayes_table(hypos, prior2, likelihood2)
table2
plot_table(table2)
plt.title('One and then Six');
table2['posterior'].idxmax()
# ## Probability Mass Functions
#
# When we do more than one update, we don't always want to keep the whole Bayes table. In this section we'll replace the Bayes table with a more compact representation, a probability mass function, or PMF.
#
# A PMF is a set of possible outcomes and their corresponding probabilities. There are many ways to represent a PMF; in this notebook I'll use a Pandas Series.
#
# Here's a function that takes a sequence of outcomes, `xs`, and a sequence of probabilities, `ps`, and returns a Pandas Series that represents a PMF.
def make_pmf(xs, ps, **options):
"""Make a Series that represents a PMF.
xs: sequence of values
ps: sequence of probabilities
options: keyword arguments passed to Series constructor
returns: Pandas Series
"""
pmf = pd.Series(ps, index=xs, **options)
return pmf
# And here's a PMF that represents the prior from the 101 Bowls problem.
# +
xs = np.arange(101)
prior = 1/101
pmf = make_pmf(xs, prior)
pmf.head()
# -
# Now that we have a priod, we need to compute likelihoods.
#
# Here are the likelihoods for a vanilla cookie:
likelihood_vanilla = xs / 100
# And for a chocolate cookie.
likelihood_chocolate = 1 - xs / 100
# To compute posterior probabilities, I'll use the following function, which takes a PMF and a sequence of likelihoods, and updates the PMF:
def bayes_update(pmf, likelihood):
"""Do a Bayesian update.
pmf: Series that represents the prior
likelihood: sequence of likelihoods
"""
pmf *= likelihood
pmf /= pmf.sum()
# The steps here are the same as in the Bayes table:
#
# 1. Multiply the prior by the likelihoods.
#
# 2. Add up the products to get the total probability of the data.
#
# 3. Divide through to normalize the posteriors.
# Now we can do the update for a vanilla cookie.
bayes_update(pmf, likelihood_vanilla)
# Here's what the PMF looks like after the update.
# +
pmf.plot()
plt.xlabel('Bowl #')
plt.ylabel('Probability')
plt.title('One cookie');
# -
# That's consistent with what we got with the Bayes table.
#
# The advantage of using a PMF is that it is easier to do multiple updates. The following cell starts again with the uniform prior and does updates with two vanilla cookies and one chocolate cookie:
# +
data = 'VVC'
pmf = make_pmf(xs, prior)
for cookie in data:
if cookie == 'V':
bayes_update(pmf, likelihood_vanilla)
else:
bayes_update(pmf, likelihood_chocolate)
# -
# Here's what the results look like:
# +
pmf.plot()
plt.xlabel('Bowl #')
plt.ylabel('Probability')
plt.title('Three cookies');
# -
# Again, that's consistent with what we got with the Bayes table.
#
# In the next section, I'll use a PMF and `bayes_update` to solve a dice problem.
# ## The dice problem
#
# As an exercise, let's do one more version of the dice problem:
#
# > Suppose you have a 4-sided, 6-sided, 8-sided, 12-sided, and a **20-sided die**. You choose one at random, roll it and **get a 7**. What is the probability that the die you rolled is 4-sided? What are the posterior probabilities for the other dice?
#
# Notice that in this version I've added a 20-sided die and the outcome is 7, not 1.
#
# Here's a PMF that represents the prior:
# +
sides = np.array([4, 6, 8, 12, 20])
prior = 1/5
pmf = make_pmf(sides, prior)
pmf
# -
# In this version, the hypotheses are integers rather than strings, so we can compute the likelihoods like this:
likelihood = 1 / sides
# But the outcome is 7, so any die with fewer than 7 sides has likelihood 0.
#
# We can adjust `likelihood` by making a Boolean Series:
too_low = (sides < 7)
# And using it to set the corresponding elements of `likelihood` to 0.
likelihood[too_low] = 0
likelihood
# Now we can do the update and display the results.
bayes_update(pmf, likelihood)
pmf
# The 4-sided and 6-sided dice have been eliminated. Of the remaining dice, the 8-sided die is the most likely.
# **Exercise:** Suppose you have the same set of 5 die. You choose a die, roll it six times, and get 6, 7, 2, 5, 1, and 2 again. Use `idxmax` to find the MAP. What is the posterior probability of the MAP?
# +
sides = np.array([4, 6, 8, 12, 20])
rolls = np.array([6, 7, 2, 5, 1])
prior = 1/5
pmf = make_pmf(sides, prior)
for roll in rolls:
likelihood = 1 / sides
too_low = (sides < roll)
likelihood[too_low] = 0
bayes_update(pmf, likelihood)
print('Rolled: ', roll)
print(pmf, '\n')
# -
pmf.idxmax()
# +
# Solution goes here
# +
# Solution goes here
# -
# ## Summary
#
# In this notebook, we extended the cookie problem with more bowls and the dice problem with more dice.
#
# I defined the MAP, which is the quantity in a posterior distribution with the highest probability.
#
# Although the cookie problem is not particularly realistic or useful, the method we used to solve it applies to many problems in the real world where we want to estimate a proportion.
#
# [In the next notebook](https://colab.research.google.com/github/AllenDowney/BiteSizeBayes/blob/master/07_euro.ipynb) we'll use the same method to take another step toward doing Bayesian statistics.
|
06_pmf_my_soln.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# This example shows how to use a `GridInducingVariationalGP` module. This classification module is designed for when the function you're modeling has 2-3 dimensional inputs and you don't believe that the output can be additively decomposed.
# # Need clarification on additive classification again
#
#
# In this example, the function is checkerboard of 1/3x1/3 squares with labels of -1 or 1
#
#
# Here we use KISS-GP (https://arxiv.org/pdf/1503.01057.pdf) to classify
# +
import math
import torch
import gpytorch
from matplotlib import pyplot as plt
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
# +
from torch.autograd import Variable
# We make an nxn grid of training points
# In [0,1]x[0,1] spaced every 1/(n-1)
n = 30
train_x = torch.zeros(int(pow(n, 2)), 2)
train_y = torch.zeros(int(pow(n, 2)))
for i in range(n):
for j in range(n):
train_x[i * n + j][0] = float(i) / (n - 1)
train_x[i * n + j][1] = float(j) / (n - 1)
# True function is checkerboard of 1/3x1/3 squares with labels of -1 or 1
train_y[i * n + j] = pow(-1, int(3 * i / n + int(3 * j / n)))
train_x = Variable(train_x)
train_y = Variable(train_y)
# -
from torch import nn, optim
from gpytorch.kernels import RBFKernel, GridInterpolationKernel
from gpytorch.means import ConstantMean
from gpytorch.likelihoods import GaussianLikelihood, BernoulliLikelihood
from gpytorch.random_variables import GaussianRandomVariable
# +
# Our classification model is just KISS-GP run through a Bernoulli likelihood
class GPClassificationModel(gpytorch.models.GridInducingVariationalGP):
def __init__(self):
super(GPClassificationModel, self).__init__(grid_size=10, grid_bounds=[(0, 1), (0, 1)])
# Near-zero mean
self.mean_module = ConstantMean(constant_bounds=[-1e-5, 1e-5])
# RBF as universal approximator
self.covar_module = RBFKernel(log_lengthscale_bounds=(-5, 6))
self.register_parameter('log_outputscale', nn.Parameter(torch.Tensor([0])), bounds=(-5,6))
def forward(self,x):
# Learned mean is near-zero
mean_x = self.mean_module(x)
# Get predictive and scale
covar_x = self.covar_module(x)
covar_x = covar_x.mul(self.log_outputscale.exp())
# Store as Gaussian
latent_pred = GaussianRandomVariable(mean_x, covar_x)
return latent_pred
# Initialize classification model
model = GPClassificationModel()
# Likelihood is Bernoulli, warm predictive mean
likelihood = BernoulliLikelihood()
# +
# Find optimal model hyperparameters
model.train()
likelihood.train()
# Use the adam optimizer
optimizer = torch.optim.Adam([
{'params': model.parameters()},
# BernoulliLikelihood has no parameters
], lr=0.1)
# "Loss" for GPs - the marginal log likelihood
# n_data refers to the amount of training data
mll = gpytorch.mlls.VariationalMarginalLogLikelihood(likelihood, model, n_data=len(train_y))
def train():
num_training_iterations = 200
for i in range(num_training_iterations):
# zero back propped gradients
optimizer.zero_grad()
# Make prediction
output = model(train_x)
# Calc loss and use to compute derivatives
loss = -mll(output, train_y)
loss.backward()
print('Iter %d/%d - Loss: %.3f log_lengthscale: %.3f' % (
i + 1, num_training_iterations, loss.data[0],
model.covar_module.base_kernel_module.log_lengthscale.data.squeeze()[0],
))
optimizer.step()
# %time train()
# +
# Set model and likelihood into eval mode
model.eval()
likelihood.eval()
# Initialize figiure an axis
f, observed_ax = plt.subplots(1, 1, figsize=(4, 3))
# Test points are 100x100 grid of [0,1]x[0,1] with spacing of 1/99
n = 100
test_x = Variable(torch.zeros(int(pow(n, 2)), 2))
for i in range(n):
for j in range(n):
test_x.data[i * n + j][0] = float(i) / (n-1)
test_x.data[i * n + j][1] = float(j) / (n-1)
# Make binary predictions by warmping the model output through a Bernoulli likelihood
with gpytorch.beta_features.fast_pred_var():
predictions = likelihood(model(test_x))
# Define plotting function
def ax_plot(ax, rand_var, title):
# prob<0.5 --> label -1 // prob>0.5 --> label 1
pred_labels = rand_var.mean().ge(0.5).float().mul(2).sub(1).data.numpy()
# Colors = yellow for 1, red for -1
color = []
for i in range(len(pred_labels)):
if pred_labels[i] == 1:
color.append('y')
else:
color.append('r')
# Plot data a scatter plot
ax.scatter(test_x.data[:, 0].numpy(), test_x.data[:, 1].numpy(), color=color, s=1)
ax.set_ylim([-0.5, 1.5])
ax.set_title(title)
# Plot predictions
ax_plot(observed_ax, predictions, 'Predicted Values')
# -
|
examples/kissgp_kronecker_product_classification.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# x_new = x - alpha * gradient(x)
import numpy as np
def gradient_descent(f, gradient, x0, alpha, eps, max_iter):
x = x0
for i in range(max_iter):
x_new = x - alpha * gradient(x)
if np.abs(f(x_new) - f(x)) < eps:
break
x = x_new
converged = i != max_iter
result = {}
result['converged'] = converged
result['num_iter'] = i
result['x'] = x_new
return result
def f(x):
return 0.5 * (x[0] ** 2 + 10*x[1]**2)
def gradient(x):
return np.array([x[0], 10 * x[1]])
# +
x0 = np.array([3, 5])
alpha = 0.1
eps = 0.001
max_iter = 1000
gradient_descent(f, gradient, x0, alpha, eps, max_iter)
# -
def momentum(f, gradient, x0, alpha, eps, max_iter, beta):
x = x0
d = 0
for i in range(max_iter):
d = beta * d + alpha * gradient(x)
x_new = x - d
if np.abs(f(x_new) - f(x)) < eps:
break
x = x_new
converged = i != max_iter
result = {}
result['converged'] = converged
result['num_iter'] = i
result['x'] = x_new
return result
momentum(f, gradient, x0, alpha, eps, max_iter, beta=0.5)
def nesterov(f, gradient, x0, alpha, eps, max_iter, beta):
x = x0
d = 0
for i in range(max_iter):
d = beta * d + alpha * gradient(x - beta * d)
x_new = x - d
if np.abs(f(x_new) - f(x)) < eps:
break
x = x_new
converged = i != max_iter
result = {}
result['converged'] = converged
result['num_iter'] = i
result['x'] = x_new
return result
nesterov(f, gradient, x0, alpha, eps, max_iter, beta=0.9)
# n-ti momenat: E(X^n)
# E(X)
# E(X^2)
#
# $$ m_0 = 0 $$
# $$ m_1 = \beta_1 m_0 + (1 - \beta_1)g_1 = (1 - \beta_1)g_1 $$
# $$ m_2 = \beta_1 m_1 + (1 - \beta_1)g_2 = \beta_1 (1 - \beta_1)g_1 + (1 - \beta_1)g_2 $$
# $$ m_3 = \beta_1 m_2 + (1 - \beta_1)g_3 = \beta_1 ^ 2(1 - \beta_1)g_1 + \beta_1(1 - \beta_1)g_2 + (1 - \beta_1)g_3 $$
#
# $$ m_t = (1 - \beta_1) \sum_{i=0}^t \beta_1^{t - i}g_i $$
#
# $$ E[m_t] = E[(1 - \beta_1) \sum_{i=0}^t \beta_1^{t - i}g_i]
# = E[g_t] [(1 - \beta_1) \sum_{i=0}^t \beta_1^{t - i} + greska$$
#
# $$ (1 - \beta_1) \sum_{i=0}^t \beta_1^{t - i} = 1 - \beta_1 ^t $$
def adam(f, gradient, x0, alpha, eps, max_iter, beta1, beta2, delta):
m = 0
v = 0
x = x0
for i in range(1, max_iter):
g = gradient(x)
m = beta1 * m + (1 - beta1) * g
v = beta2 * v + (1 - beta2) * g ** 2
m_hat = m / (1 - beta1 ** i)
v_hat = v / (1 - beta2 ** i)
x_new = x - alpha * (m_hat / (np.sqrt(v_hat) + delta))
if np.abs(f(x_new) - f(x)) < eps:
break
x = x_new
converged = i != max_iter
result = {}
result['converged'] = converged
result['num_iter'] = i
result['x'] = x_new
return result
adam(f, gradient, x0, alpha, eps, max_iter, beta1=0.9, beta2=0.999, delta=1e-7)
# +
# Branch and Bound
# +
# Q---
# ---
# -Q--
# ----
# -
def print_solution(board):
n = board.shape[0]
for i in range(n):
for j in range(n):
print(board[i][j], end=' ')
print()
def solve(n):
board = np.full((n, n), '.', dtype=str)
row_check = np.full(n, False)
d1_check = np.full(2 * n - 1, False)
d2_check = np.full(2 * n - 1, False)
if not bnb(board, 0, row_check, d1_check, d2_check):
print('Nema resenja')
return False
print_solution(board)
def bnb(board, c, row_check, d1_check, d2_check):
n = board.shape[0]
if c >= n:
return True
for r in range(n):
if is_free(r, c, row_check, d1_check, d2_check):
board[r][c] = 'Q'
row_check[r] = True
d1_check[r + c] = True
d2_check[r - c + n - 1] = True
if bnb(board, c + 1, row_check, d1_check, d2_check):
return True
board[r][c] = '.'
row_check[r] = False
d1_check[r + c] = False
d2_check[r - c + n - 1] = False
return False
def is_free(r, c, row_check, d1_check, d2_check):
n = row_check.shape[0]
if row_check[r] or d1_check[r + c] or d2_check[r - c + n - 1]:
return False
return True
solve(8)
|
2020_2021/live/05_gradient_descent.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Draft Bot Version 1 Exploration
# ### J&T
#
import pandas as pd
import numpy as np
# ### Load in dataset
draft_csv = 'data/draft_vow_100000.csv'
df = pd.read_csv(draft_csv)
df
df.columns
pd.set_option('display.max_columns', None)
# pd.set_option('display.max_rows', None)
# columns = ['draft_id', 'user_rank', 'event_match_wins', 'event_match_losses', 'pack_number', 'pick_number']
# columns = []
max_cols = 14
nocard_df = df.iloc[:, :max_cols]
nocard_df
nocard_df.describe()
df.describe()
# ## Question 1
# What cards correlate to the highest draft win rate?
# (Uncommmons)
|
Draft Bot Exploration.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="rVKzWm1PRmaD"
# Importing all the required libraries
import numpy as np
import pandas as pd
import matplotlib as mpl
import matplotlib.pyplot as plt
from matplotlib import style
from sklearn import preprocessing
from sklearn.linear_model import LinearRegression, Ridge, Lasso
from sklearn.model_selection import train_test_split
from datetime import datetime, timedelta
# %matplotlib inline
# + colab={"base_uri": "https://localhost:8080/", "height": 238} id="vkuIvKLiRsQ5" outputId="86b76565-ecb8-404d-d24b-95364f6d1b28"
# The historical data of stock prices of Apple was downloaded from Yahoo! Finance Website in the .csv format
# Website - https://finance.yahoo.com/quote/AAPL/history?p=AAPL
# Reading the CSV file
df = pd.read_csv('AAPL.csv')
df.set_index('Date', inplace=True)
df.tail()
# + colab={"base_uri": "https://localhost:8080/", "height": 584} id="XYZVf1NLSW6P" outputId="9ead5bfd-faf2-4a40-8373-7a93cfee87ef"
# Visualizing the stock prices
df['Adj Close'].plot(label='AAPL', figsize=(15, 9), title='Adjusted Closing Price', color='red', linewidth=1.0, grid=True)
plt.legend()
# + id="ySTPqKLeTP9y"
# Rolling Mean / Moving Average to remove the noise in the graph and smoothen it
close_col = df['Adj Close']
mvag = close_col.rolling(window=100).mean() # Taking an average over the window size of 100.
# Increasing the window size can make it more smoother, but less informative and vice-versa.
# + colab={"base_uri": "https://localhost:8080/", "height": 638} id="5mD0edxCTboB" outputId="eb01d713-e026-42e0-c364-5357ab7c357e"
# Visualizing Rolling Mean and Adjusted Closing Price together
df['Adj Close'].plot(label='AAPL', figsize=(15,10), title='Adjusted Closing Price vs Moving Average', color='red', linewidth=1.0, grid=True)
mvag.plot(label='MVAG', color='blue')
plt.legend()
# + colab={"base_uri": "https://localhost:8080/", "height": 638} id="NhGoIScjTf94" outputId="b24be804-eafe-407a-85b6-340f2e48adac"
# Return Deviation measures the Mean of the Probability Distribution of Investment Returns if it has a positive/negative Average Net Outcome
rd = close_col / close_col.shift(1) - 1
rd.plot(label='Return', figsize=(15, 10), title='Return Deviation', color='red', linewidth=1.0, grid=True)
plt.legend()
# + id="bquFgRT7TkRc"
# Number of days for which to predict the stock prices
predict_days = 30
# + id="SH36wayeTokU"
# Shifting by the Number of Predict days for Prediction array
df['Prediction'] = df['Adj Close'].shift(-predict_days)
# print(df['Prediction'])
# print(df['Adj Close'])
# + colab={"base_uri": "https://localhost:8080/"} id="i7u-XK2vTr3V" outputId="a7a42c93-908e-4bed-ab40-d736a75aaa2c"
# Dropping the Prediction Row
X = np.array(df.drop(['Prediction'], axis = 1))
X = X[:-predict_days] # Size upto predict days
# print(X)
print(X.shape)
# + colab={"base_uri": "https://localhost:8080/"} id="ZY-Irh8DTu87" outputId="3108280a-3743-46fd-dce6-82406f4bec3e"
# Creating the Prediction Row
y = np.array(df['Prediction'])
y = y[:-predict_days] # Size upto predict_days
# print(y)
print(y.shape)
# + colab={"base_uri": "https://localhost:8080/"} id="y64AyU3ITyFb" outputId="51dbfc3e-4e49-401d-e183-90771801a5eb"
# Splitting the data into Training data & Testing data
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2) #Splitting the data into 80% for training & 20% for testing
print(X_train.shape)
print(y_train.shape)
print(X_test.shape)
print(y_test.shape)
# + [markdown] id="7_HT-Vx0T_fu"
# **1. Linear Regression**
# + colab={"base_uri": "https://localhost:8080/"} id="AZ-B0DYXUJ0b" outputId="5a485142-e471-455d-f7f3-6cb6e84674f5"
# Defining the Linear Regression Model
linear_model = LinearRegression()
linear_model.fit(X_train, y_train) # Training the algorithm
# + colab={"base_uri": "https://localhost:8080/"} id="OS9LhKyXUUPf" outputId="5c737bd0-189d-4670-a5f5-d9b33c33d8b1"
# Score of the Linear Regression Model (Using the Test Data)
linear_model_score = linear_model.score(X_test, y_test)
print('Linear Model score:', linear_model_score)
# + colab={"base_uri": "https://localhost:8080/"} id="_8aUwKkCUWYX" outputId="c2e1be4e-a88d-48cb-bb5c-610adb9e9b1d"
# Define the Real & Prediction Values
X_predict = np.array(df.drop(['Prediction'], 1))[-predict_days:]
linear_model_predict_prediction = linear_model.predict(X_predict)
linear_model_real_prediction = linear_model.predict(np.array(df.drop(['Prediction'], 1)))
# + id="pn5Ub_2SUcBb"
# Defining some Parameters
predicted_dates = []
recent_date = df.index.max()
display_at = 1000
alpha = 0.5
for i in range(predict_days):
recent_date += str(timedelta(days=1))
predicted_dates.append(recent_date)
# + colab={"base_uri": "https://localhost:8080/", "height": 384} id="GoV4o0YMUdMs" outputId="1a1d7d24-74ed-46d3-9450-0ce1f8e4b0bd"
# Plotting the Actual and Prediction Prices
plt.figure(figsize=(15, 9))
plt.plot(df.index[display_at:], linear_model_real_prediction[display_at:], label='Linear Prediction', color='blue', alpha=alpha)
plt.plot(predicted_dates, linear_model_predict_prediction, label='Forecast', color='green', alpha=alpha)
plt.plot(df.index[display_at:], df['Close'][display_at:], label='Actual', color='red')
plt.legend()
# + [markdown] id="9G5vEWNWUr3x"
# **2. Lasso Regression**
# + colab={"base_uri": "https://localhost:8080/"} id="Y9Ws7-J2U8Da" outputId="7ec4c6f2-fad4-458d-e48b-89e708d041b2"
# Defining the Lasso Regression Model
lasso_model = Lasso()
lasso_model.fit(X_train, y_train) # Training the algorithm
# + colab={"base_uri": "https://localhost:8080/"} id="Naf2uxglU9AC" outputId="9ee95f37-0a23-47a6-daed-f7dfba7e9881"
# Score of the Lasso Regression Model (Using the Test Data)
lasso_model_score = lasso_model.score(X_test, y_test)
print('Lasso Model score:', lasso_model_score)
# + colab={"base_uri": "https://localhost:8080/"} id="cDIruJcEU_a4" outputId="1042def7-fd62-4292-f82c-89947ecc9cb3"
# Define the Real & Prediction Values
lasso_model_predict_prediction = lasso_model.predict(X_predict)
lasso_model_real_prediction = lasso_model.predict(np.array(df.drop(['Prediction'], 1)))
# + colab={"base_uri": "https://localhost:8080/", "height": 384} id="JLJ8cl2JVCEn" outputId="cf528112-2013-46cf-99de-4f257d027ef7"
# Plotting the Actual and Prediction Prices
plt.figure(figsize=(15, 9))
plt.plot(df.index[display_at:], lasso_model_real_prediction[display_at:], label='Lasso Prediction', c='blue', alpha=alpha)
plt.plot(predicted_dates, lasso_model_predict_prediction, label='Forecast', color='green', alpha=alpha)
plt.plot(df.index[display_at:], df['Close'][display_at:], label='Actual', color='red')
plt.legend()
# + [markdown] id="W8XZXRBPYBws"
# **3. Ridge Regression**
# + colab={"base_uri": "https://localhost:8080/"} id="gu8W-w_PYNfC" outputId="5a64f3c1-cc1f-4551-b8f9-6dc9a516af89"
# Defining the Ridge Regression Model
ridge_model = Ridge()
ridge_model.fit(X_train, y_train) # Training the algorithm
# + colab={"base_uri": "https://localhost:8080/"} id="ikDonCQxYRJL" outputId="f1cdbede-35ff-4dbd-b116-46dbcd582a05"
# Score of the Ridge Regression Model (Using the Test Data)
ridge_model_score = ridge_model.score(X_test, y_test)
print('Ridge Model score:', ridge_model_score)
# + colab={"base_uri": "https://localhost:8080/"} id="MxSpuD8ZYU_S" outputId="961f6728-97a9-40d3-b95d-b0a2b6103165"
# Define the Real & Prediction Values
ridge_model_predict_prediction = ridge_model.predict(X_predict)
ridge_model_real_prediction = ridge_model.predict(np.array(df.drop(['Prediction'], 1)))
# + colab={"base_uri": "https://localhost:8080/", "height": 384} id="inyiq90WYYZc" outputId="7ebe2478-c5ac-4a7d-9c85-38831542f45d"
# Plotting the Actual and Prediction Prices
plt.figure(figsize=(15, 9))
plt.plot(df.index[display_at:], ridge_model_real_prediction[display_at:], label='Ridge Prediction', color='blue', alpha=alpha)
plt.plot(predicted_dates, ridge_model_predict_prediction, label='Forecast', color='green', alpha=alpha)
plt.plot(df.index[display_at:], df['Close'][display_at:], label='Actual', color='red')
plt.legend()
# + colab={"base_uri": "https://localhost:8080/"} id="1iMIcAHNVE4M" outputId="82253e76-1254-41ec-e669-cbf8d3c44651"
# Best Performance of the Regressor Models
best_score = max(linear_model_score, lasso_model_score, ridge_model_score)
index = np.argmax([linear_model_score, lasso_model_score, ridge_model_score])
best_regressor = {0:'Linear Regression Model',
1:'Lasso Model',
2:'Ridge Model'}
print("The Best Performing Model is {0} with the score of {1}%.".format(best_regressor[index], best_score*100))
|
stock-price-prediction.ipynb
|