code
stringlengths 38
801k
| repo_path
stringlengths 6
263
|
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
'''
import libraries and set print options
'''
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import csv
pd.set_option('display.max_rows', 1000)
# # Exploratory Data Analysis - DISREGARD
data = pd.read_csv('../data/receipt_data_manual.csv') ##read csv as a dataframe
full_data = data.copy()
data.head() ## show the top 5
for i in range(len(data.database)):
if (pd.isnull(data.database[i])):
data.database[i]=''
else:
data.database[i] = data.database[i].strip()
titles = data.database.tolist() ## get the long_name column into a list
len(titles) ## print the length
possible_brands = [x.split(",")[0] for x in titles if len(x.split(",")) > 1] ## what is the 0th index for items with commas
possible_brands[:10]
possible_brands = [x for x in titles if len(x.split(",")) == 4]
print (len(possible_brands))
possible_brands
# ## RULES:
# ### - The zeroth index seems to be brand for the most part (few exceptions)
# ### - When there are 3 items, the first index is the main product with the 2nd index being a descriptor/flavor/type
# ### - When there are 4 items (only ~4k examples), the 2nd index is the product, the 0th index is still brand, the 3rd index is the descriptor
# # Data Cleaning - Process Is Commented
if 'no_brand_descriptor_title' in data.columns:
DROP_COLS = [
'receipt_name',
'full_product_title',
'general_product_title',
'store',
'no_brand_descriptor_title',
'no_descriptor_title'
]
else:
DROP_COLS = [
'receipt_name',
'full_product_title',
'general_product_title',
'store'
]
data = data.drop(DROP_COLS, axis=1) ## drop unnecessary columns for now
data.head()
# +
'''
Find the number of commas in each title and only use those with less than 5
'''
data['num_commas'] = data.apply(lambda x: len(x.database.split(","))-1, axis=1)
# data = data[data['num_commas'] < 4]
for index in range(len(data.database)):
if (data.num_commas[index] >= 4):
data.database[index] = ''
# -
len(data)
'''
This will keep brands and separate it from the rest of the titles with a ",". EX: "FRESH & EASY, CANOLA OIL"
'''
def remove_descriptor(x):
x_ = x.split(",")
if len(x_) == 2 or len(x_) == 3:
if 'fl oz' in x_[1]:
return x_[0].strip()
else:
return x_[0].strip() + ", " + x_[1].strip()
elif len(x_) == 4:
return x_[0].strip() + ", " + x_[2].strip()
else:
return x_[0].strip()
'''
This will keep brands and separate it from the rest of the titles with a ",". EX: "FRESH & EASY, CANOLA OIL"
'''
def remove_brand_descriptor(x):
x_ = x.split(",")
if len(x_) > 1:
if 'fl oz' in x_[1]:
return x_[0].strip()
else:
return x_[1].strip()
elif len(x_) == 4:
return x_[2].strip()
else:
return x_[0].strip()
'''
Apply the 2 methods to obtain two different columns with the cleaning process applied
'''
data['no_brand_descriptor_title'] = data.apply(lambda x: remove_brand_descriptor(x.database), axis=1)
data['no_descriptor_title'] = data.apply(lambda x: remove_descriptor(x.database), axis=1)
data.tail() ##print dataframe
# +
'''
Get rid of leading and ending spaces, make all one case, in csv file
'''
for i in range(len(full_data.database)):
full_data.receipt_name[i] = '' if pd.isnull(full_data.receipt_name[i]) else (full_data.receipt_name[i].upper()).strip()
full_data.full_product_title[i] = '' if pd.isnull(full_data.full_product_title[i]) else (full_data.full_product_title[i].lower()).strip()
full_data.general_product_title[i] = '' if pd.isnull(full_data.general_product_title[i]) else (full_data.general_product_title[i].lower()).strip()
full_data.store[i] = '' if pd.isnull(full_data.store[i]) else (full_data.store[i].lower()).strip()
# -
for i in range(len(full_data.database)):
if (data.database[i] == ''):
print([full_data.receipt_name[i], full_data.full_product_title[i], full_data.general_product_title[i], full_data.store[i], data.database[i], data.no_brand_descriptor_title[i], data.no_descriptor_title[i]])
'''
Create csv rows where there is a product match. Put all unmatched products in a separate file
'''
csvRows = []
no_match_row = []
csvRows.append(['receipt_name', 'full_product_title', 'general_product_title', 'store', 'database', 'no_brand_descriptor_title', 'no_descriptor_title'])
for i in range(len(full_data.database)):
if (data.database[i] == ''):
no_match_row.append([full_data.receipt_name[i], full_data.full_product_title[i], full_data.general_product_title[i], full_data.store[i]])
else:
csvRows.append([full_data.receipt_name[i], full_data.full_product_title[i], full_data.general_product_title[i], full_data.store[i], data.database[i], data.no_brand_descriptor_title[i], data.no_descriptor_title[i]])
'''
Create new, cleaned csv of manual data
'''
csvfile = '../data/cleaned_receipt_data_manual.csv'
with open(csvfile, "w") as fp:
wr = csv.writer(fp, dialect='excel')
wr.writerows(csvRows)
'''
Create csv of manual data with no matches
'''
csvfile = '../data/no_matches_manual_data.csv'
with open(csvfile, "w") as fp:
wr = csv.writer(fp, dialect='excel')
wr.writerows(no_match_row)
|
synthetic_data/scripts/manual_data_cleaning.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/JeffreyAsuncion/DS-Unit-2-Linear-Models/blob/master/DSPT6_U2S1M2_JeffreyAsuncion_assignment.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="X2ZGQqwRoxI6" colab_type="text"
# Lambda School Data Science
#
# *Unit 2, Sprint 1, Module 2*
#
# ---
# + [markdown] colab_type="text" id="7IXUfiQ2UKj6"
# # Regression 2
#
# ## Assignment
#
# You'll continue to **predict how much it costs to rent an apartment in NYC,** using the dataset from renthop.com.
#
# - [x] Do train/test split. Use data from April & May 2016 to train. Use data from June 2016 to test.
# - [x] Engineer at least two new features. (See below for explanation & ideas.)
# - [x] Fit a linear regression model with at least two features.
# - [x] Get the model's coefficients and intercept.
# - [x] Get regression metrics RMSE, MAE, and $R^2$, for both the train and test data.
# - [ ] What's the best test MAE you can get? Share your score and features used with your cohort on Slack!
# - [ ] As always, commit your notebook to your fork of the GitHub repo.
#
#
# #### [Feature Engineering](https://en.wikipedia.org/wiki/Feature_engineering)
#
# > "Some machine learning projects succeed and some fail. What makes the difference? Easily the most important factor is the features used." — <NAME>, ["A Few Useful Things to Know about Machine Learning"](https://homes.cs.washington.edu/~pedrod/papers/cacm12.pdf)
#
# > "Coming up with features is difficult, time-consuming, requires expert knowledge. 'Applied machine learning' is basically feature engineering." — <NAME>, [Machine Learning and AI via Brain simulations](https://forum.stanford.edu/events/2011/2011slides/plenary/2011plenaryNg.pdf)
#
# > Feature engineering is the process of using domain knowledge of the data to create features that make machine learning algorithms work.
#
# #### Feature Ideas
# - Does the apartment have a description?
# - How long is the description?
# - How many total perks does each apartment have?
# - Are cats _or_ dogs allowed?
# - Are cats _and_ dogs allowed?
# - Total number of rooms (beds + baths)
# - Ratio of beds to baths
# - What's the neighborhood, based on address or latitude & longitude?
#
# ## Stretch Goals
# - [ ] If you want more math, skim [_An Introduction to Statistical Learning_](http://faculty.marshall.usc.edu/gareth-james/ISL/ISLR%20Seventh%20Printing.pdf), Chapter 3.1, Simple Linear Regression, & Chapter 3.2, Multiple Linear Regression
# - [ ] If you want more introduction, watch [<NAME>, Statistics 101: Simple Linear Regression](https://www.youtube.com/watch?v=ZkjP5RJLQF4)
# (20 minutes, over 1 million views)
# - [ ] Add your own stretch goal(s) !
# + colab_type="code" id="o9eSnDYhUGD7" colab={}
# %%capture
import sys
# If you're on Colab:
if 'google.colab' in sys.modules:
DATA_PATH = 'https://raw.githubusercontent.com/LambdaSchool/DS-Unit-2-Applied-Modeling/master/data/'
# !pip install category_encoders==2.*
# If you're working locally:
else:
DATA_PATH = '../data/'
# Ignore this Numpy warning when using Plotly Express:
# FutureWarning: Method .ptp is deprecated and will be removed in a future version. Use numpy.ptp instead.
import warnings
warnings.filterwarnings(action='ignore', category=FutureWarning, module='numpy')
# + colab_type="code" id="cvrw-T3bZOuW" colab={}
import numpy as np
import pandas as pd
# Read New York City apartment rental listing data
df = pd.read_csv(DATA_PATH+'apartments/renthop-nyc.csv')
assert df.shape == (49352, 34)
# Remove the most extreme 1% prices,
# the most extreme .1% latitudes, &
# the most extreme .1% longitudes
df = df[(df['price'] >= np.percentile(df['price'], 0.5)) &
(df['price'] <= np.percentile(df['price'], 99.5)) &
(df['latitude'] >= np.percentile(df['latitude'], 0.05)) &
(df['latitude'] < np.percentile(df['latitude'], 99.95)) &
(df['longitude'] >= np.percentile(df['longitude'], 0.05)) &
(df['longitude'] <= np.percentile(df['longitude'], 99.95))]
# + id="OKYGKuKPgjL7" colab_type="code" outputId="66dc66d0-7bdb-4640-bc41-ce61937570b7" colab={"base_uri": "https://localhost:8080/", "height": 35}
df.shape
# + id="_I0ewQDHgnHx" colab_type="code" outputId="8b5f506a-ca47-4ca0-a5d7-c6b92ebf7e6c" colab={"base_uri": "https://localhost:8080/", "height": 181}
df.columns
# + id="N4xrhp1tzeSZ" colab_type="code" outputId="7ebbf496-fa19-485d-d989-6dd1d94e33c6" colab={"base_uri": "https://localhost:8080/", "height": 249}
df.head(2)
# + id="8lbKPb3bgpjW" colab_type="code" outputId="032c4535-e69f-4e3a-fb3d-ac59ba3488a1" colab={"base_uri": "https://localhost:8080/", "height": 108}
# date df['created']
df['created'].describe()
# + [markdown] id="vUbWKTe3ppwn" colab_type="text"
# Split the data into two sets
# ====================
#
# - Train data from April & May 2016
# - Test data from June 2016
# + id="yiBcZyoblcez" colab_type="code" outputId="e0a23301-ba7f-449d-c227-c57528cc8c3c" colab={"base_uri": "https://localhost:8080/", "height": 35}
# Split the data into two sets
train = df[df['created'] < '2016-06-01']
test = df[df['created'] >= '2016-06-01']
train.shape, test.shape
# + [markdown] id="6a9h5d3uuMKU" colab_type="text"
# Create 2 New Features
# ==========
#
# - Does the apartment have a description?
# - How long is the description?
# - How many total perks does each apartment have?
# - Are cats _or_ dogs allowed?
# - Are cats _and_ dogs allowed?
# - Total number of rooms (beds + baths)
# - Ratio of beds to baths
# - What's the neighborhood, based on address or latitude & longitude?
# + id="mZEXyqVL-b6Z" colab_type="code" colab={}
# to Avoid SettingWithCopyWarning
train = train.copy()
test = test.copy()
# + id="MMNI3PzlxYsu" colab_type="code" outputId="900e40ef-21f9-4aeb-818b-b23621d440f2" colab={"base_uri": "https://localhost:8080/", "height": 181}
train.columns
# + [markdown] id="Qml3R23VzUba" colab_type="text"
# Total number of rooms (beds + baths)
# + id="vh9YNqMhqdiA" colab_type="code" colab={}
# Total number of rooms (beds + baths)
train['total_num_of_rooms'] = train['bedrooms'] + train['bathrooms']
test['total_num_of_rooms'] = test['bedrooms'] + test['bathrooms']
# + id="q8BKYG8KtYfb" colab_type="code" outputId="a9697530-f9ec-42a0-80df-fa1d2c44fec5" colab={"base_uri": "https://localhost:8080/", "height": 108}
train[['total_num_of_rooms', 'bedrooms', 'bathrooms']].head(2)
# + id="j_2lH21L7vbC" colab_type="code" outputId="19279ba8-3f1d-437c-a3f3-f54edaeada5b" colab={"base_uri": "https://localhost:8080/", "height": 108}
test[['total_num_of_rooms', 'bedrooms', 'bathrooms']].head(2)
# + [markdown] id="XqlN5SP2zZ4Q" colab_type="text"
# Ratio of beds to baths
# + id="V717ZDuTu5ct" colab_type="code" colab={}
# Ratio of beds to baths
train['ratio_bed_to_baths'] = train['bedrooms'] / train['bathrooms']
test['ratio_bed_to_baths'] = test['bedrooms'] / test['bathrooms']
# + id="1BRtcuM6vp4Q" colab_type="code" outputId="0aec43a0-630b-4b8c-a832-a228b9286f60" colab={"base_uri": "https://localhost:8080/", "height": 108}
# check the 'ratio_bed_to_baths
train[['ratio_bed_to_baths', 'bedrooms', 'bathrooms']].head(2)
# + id="Ko_6Ph5wNSgV" colab_type="code" outputId="4859cffc-7bc9-423c-f6cc-8a1c2676c01a" colab={"base_uri": "https://localhost:8080/", "height": 581}
train['ratio_bed_to_baths'].value_counts()
#### inf == infinity
#### Really that's no good
# + [markdown] id="FPutMz8EzP64" colab_type="text"
# How many total perks does each apartment have?
# + id="iHlbaFrhzMkx" colab_type="code" outputId="e192fdb1-aa50-471b-8dd6-7a641e1b4137" colab={"base_uri": "https://localhost:8080/", "height": 199}
train.columns
# + id="2G-tpJtw6S7w" colab_type="code" colab={}
perks =['elevator', 'cats_allowed', 'hardwood_floors', 'dogs_allowed',
'doorman', 'dishwasher', 'no_fee', 'laundry_in_building',
'fitness_center', 'pre-war', 'laundry_in_unit', 'roof_deck',
'outdoor_space', 'dining_room', 'high_speed_internet', 'balcony',
'swimming_pool', 'new_construction', 'terrace', 'exclusive', 'loft',
'garden_patio', 'wheelchair_access', 'common_outdoor_space']
# + id="tg2wQkyxzMbw" colab_type="code" outputId="77f74fbd-1484-40a4-d2a2-a0496a425aac" colab={"base_uri": "https://localhost:8080/", "height": 235}
train[perks].sample(5)
# + id="BE2m5p6y1ETu" colab_type="code" outputId="2d582429-e0a8-4f8e-edc5-81bedd4a1602" colab={"base_uri": "https://localhost:8080/", "height": 198}
# create new feature 'total_perks'
train['total_perks'] = train[perks].sum(axis=1)
test['total_perks'] = test[perks].sum(axis=1)
train[['total_perks']].sample(5)
# + [markdown] id="7zTzNoZnw9xk" colab_type="text"
# ##What's the neighborhood, based on address or latitude & longitude?
# + id="t2v2YVd6xB0B" colab_type="code" outputId="32135b06-8241-42e8-e794-52fd3d90e729" colab={"base_uri": "https://localhost:8080/", "height": 199}
## how to you do this????
## let's look at the columns given
train.columns # hello again
# + id="GDSuFcBwxBw5" colab_type="code" outputId="080f9968-3c63-4a2c-b007-161a4aff4d51" colab={"base_uri": "https://localhost:8080/", "height": 198}
train[['display_address','latitude', 'longitude','street_address']].sample(5)
# + id="fzvV5UxUxBtU" colab_type="code" outputId="ea3b8f31-7f18-42fa-b15c-0370213937a8" colab={"base_uri": "https://localhost:8080/", "height": 35}
train['latitude'].value_counts().sum()
# what if we just include the long and lat into the linear regression without feature engineering it
# that's ludicrous/ludicris ;-o
## ok let's try
# + [markdown] id="WZ7tTPCZxMkz" colab_type="text"
# Fit a linear regression model with at least two features.
# ========
#
# + id="gbFnhrdhHt5z" colab_type="code" outputId="2bb4e3fb-ac22-4477-9147-7e3f9d733ba6" colab={"base_uri": "https://localhost:8080/", "height": 35}
# 1. Begin with baseline (0 features)
train['price'].mean()
# + id="wRUZXGHBJV_l" colab_type="code" colab={}
# 2. Arrange y target vectors
target = 'price'
y_train = train[target]
y_test = test[target]
# + id="LE1kO7Y3J0eA" colab_type="code" outputId="5bfb51f7-acde-4f64-f340-75e6fa090f3a" colab={"base_uri": "https://localhost:8080/", "height": 35}
# 3. Get mean baseline
print('Mean Baseline (using 0 Features)')
guess = y_train.mean()
# make sure that, this mean is only from the train data
# do not contaminate with the test data
# + id="1O-686k3KYjB" colab_type="code" outputId="b9a2aa17-b11a-48b8-ecad-68504b09152f" colab={"base_uri": "https://localhost:8080/", "height": 35}
# Train Error
from sklearn.metrics import mean_absolute_error
y_pred = [guess] * len(y_train)
mae = mean_absolute_error(y_train, y_pred)
print(f'Train Error (April and May 2016) : ${mae:.2f}')
# + id="dj7PYobVKYck" colab_type="code" outputId="55e5b82f-ac93-41db-c6f6-f7587f683d18" colab={"base_uri": "https://localhost:8080/", "height": 35}
# Test Error
y_pred = [guess] * len(y_test)
mae = mean_absolute_error(y_test, y_pred)
print(f'Train Error (June 2016) : ${mae:.2f}')
# + id="eYDSMobyLeqo" colab_type="code" colab={}
# + id="TPZEZz4Mvwde" colab_type="code" colab={}
# 1. Import the appropriate estimator class from SciKit-Learn
from sklearn.linear_model import LinearRegression
# + id="9BfmZB9dx1tR" colab_type="code" colab={}
# 2. Instantiate this class
model = LinearRegression()
# + id="obBoHqLiyE6q" colab_type="code" outputId="1f4cadbf-7c3e-4da0-d00a-fad7c2e91a80" colab={"base_uri": "https://localhost:8080/", "height": 35}
# 3. Arrange X features matrices (alread did y targets vectors)
# Let's use the 2 new features
features = ['bedrooms', 'bathrooms', 'total_perks','latitude', 'longitude' ] #added lat and long
X_train = train[features]
X_test = test[features]
print(f'Linear Regression, dependent on: {features}')
# + id="KKtjZvaJy41V" colab_type="code" outputId="f3e9bceb-6dbd-4175-aa63-60db871f0d51" colab={"base_uri": "https://localhost:8080/", "height": 35}
# 4. Fit the model
model.fit(X_train, y_train)
y_pred_train = model.predict(X_train)
mae = mean_absolute_error(y_train, y_pred_train)
print(f'Train Error: ${mae:.2f}')
# + id="89rvtmXqL1Qf" colab_type="code" outputId="67e8bd42-dc3b-44e7-e31f-4599d8c493eb" colab={"base_uri": "https://localhost:8080/", "height": 35}
# 5. Apply the model to new data
y_pred_test = model.predict(X_test)
mae = mean_absolute_error(y_test, y_pred_test)
print(f'Test Error: ${mae:.2f}')
# + [markdown] id="FxxABT-TPv0k" colab_type="text"
# Get the model's coefficients and intercept.
# ========
#
# + id="DwF8XcKOOvXa" colab_type="code" outputId="0749dca5-00e3-4865-96f2-d3fc256003a3" colab={"base_uri": "https://localhost:8080/", "height": 72}
model.intercept_, model.coef_
# + id="VN0tYe5GQFXa" colab_type="code" outputId="819c68f4-26b9-412a-d6f9-16eda4214c30" colab={"base_uri": "https://localhost:8080/", "height": 235}
beta0 = model.intercept_
beta1, beta2, beta3, beta4, beta5 = model.coef_
print(f'intercept : {beta0:.2f}')
print(f'beta1 : {beta1:.2f}')
print(f'beta2 : {beta2:.2f}')
print(f'beta3 : {beta3:.2f}\n')
print(f'beta4 : {beta4:.2f}\n')
print(f'beta5 : {beta5:.2f}\n')
print(f'y = {beta0:.2f} + {beta1:.2f}*x1 + {beta2:.2f}*x2 + {beta3:.2f}*x3+ {beta4:.2f}*x4 + {beta5:.2f}*x5\n')
print(f'price = {beta0:.2f} + {beta1:.2f}*num_of_bedrooms + {beta2:.2f}*num_of_bathrooms + {beta3:.2f}*total_perks + {beta4:.2f}*lat + {beta5:.2f}*long')
# + [markdown] id="lzey2v6BS0Om" colab_type="text"
# Get regression metrics RMSE, MAE, and $R^2$, for both the train and test data.
# ======
#
# + id="57vqZF06S81j" colab_type="code" colab={}
from sklearn.metrics import mean_absolute_error, mean_squared_error, r2_score
# + id="pWHJ1zaiS8v2" colab_type="code" outputId="dac51d91-de39-4be1-899a-d1f01450a77b" colab={"base_uri": "https://localhost:8080/", "height": 90}
# Print Regression Metrics
mse_train = mean_squared_error(y_train, y_pred_train)
rmse_train = np.sqrt(mse_train)
mae_train = mean_absolute_error(y_train, y_pred_train)
r2_train = r2_score(y_train, y_pred_train)
mse_test = mean_squared_error(y_test, y_pred_test)
rmse_test = np.sqrt(mse_test)
mae_test = mean_absolute_error(y_test, y_pred_test)
r2_test = r2_score(y_test, y_pred_test)
print(f'Mean Squared Error \ttrain : {mse_train}\ttest : {mse_test}')
print(f'Root Mean Squared Error\ttrain : {rmse_train}\ttest : {rmse_test}')
print(f'Mean Absolute Error \ttrain : {mae_train}\ttest : {mae_test}')
print(f'R^2 \t\t\ttrain : {r2_train}\ttest : {r2_test}')
# + [markdown] id="RaE_HndY5BU2" colab_type="text"
# ## Before adding the latitude and longitude
#
# error test | train | test
# --------------------|-----------------------------|-------------
# Mean Squared Error | 1449490.55 | 1421073.10
# Root Mean Squared Error | 1203.94 | 1192.08
# Mean Absolute Error | 795.40 | 799.73
# R^2 | 0.5331 | 0.5427
# + [markdown] id="SOZjnQ_t9f_N" colab_type="text"
# ## After adding Latitude and Longitude with Feature Engineering
#
# error test | train | test
# --------------------|-----------------------------|-------------
# Mean Squared Error | 1291105.58| 1258411.98
# Root Mean Squared Error| 1136.26 | 1121.78
# Mean Absolute Error | 729.17 | 733.70
# R^2 | 0.5841 | 0.5951
# + [markdown] id="P4TYjACe-cAf" colab_type="text"
# <div align="center">
#
# There is a 60%(test) Reduction invariance
#
#
# when we take into account the following features
# (bedrooms, bathrooms, total_perks, latitude, longitude) into account.
#
# Alternatively, we can say that the following features
#
#
# (bedrooms, bathrooms, total_perks, latitude, longitude)
# explain 60%(test) of the variance in price.
#
# + id="-onY32vDo-ir" colab_type="code" colab={}
# There is a 53%(train) & 54%(test) Reduction invariance
# when we take into account the following features
# (bedrooms, bathrooms, total_perks) into account
# Alternatively, we can say that the following features
# (bedrooms, bathrooms, total_perks)
# explain 53%(train) & 54%(test) of the variance in price.
# thanks StatQuest
# + [markdown] id="RDItWRgKoUNO" colab_type="text"
# This is a great way to explain the R^2
# https://www.youtube.com/watch?v=nk2CQITm_eo&feature=youtu.be
# + [markdown] id="kcQajphAS9MA" colab_type="text"
# What's the best test MAE you can get? Share your score and features used with your cohort on Slack!
# ========
#
# + id="f65MxOnrTHIZ" colab_type="code" colab={}
# + id="0I4uwrrhTMrm" colab_type="code" colab={}
# + [markdown] id="DTU2UjudTIbQ" colab_type="text"
#
# As always, commit your notebook to your fork of the GitHub repo.
# =========
#
# + id="Jla8sBwyTNjm" colab_type="code" colab={}
# + id="L21aapLgTIJ3" colab_type="code" colab={}
|
DSPT6_U2S1M2_JeffreyAsuncion_assignment.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <div class="alert alert-block alert-info" style="margin-top: 20px">
# <a href="https://cocl.us/topNotebooksPython101Coursera">
# <img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/PY0101EN/Ad/TopAd.png" width="750" align="center">
# </a>
# </div>
# <a href="https://cognitiveclass.ai/">
# <img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/PY0101EN/Ad/CCLog.png" width="200" align="center">
# </a>
# <h1>Functions in Python</h1>
# <p><strong>Welcome!</strong> This notebook will teach you about the functions in the Python Programming Language. By the end of this lab, you'll know the basic concepts about function, variables, and how to use functions.</p>
# <h2>Table of Contents</h2>
# <div class="alert alert-block alert-info" style="margin-top: 20px">
# <ul>
# <li>
# <a href="#func">Functions</a>
# <ul>
# <li><a href="content">What is a function?</a></li>
# <li><a href="var">Variables</a></li>
# <li><a href="simple">Functions Make Things Simple</a></li>
# </ul>
# </li>
# <li><a href="pre">Pre-defined functions</a></li>
# <li><a href="if">Using <code>if</code>/<code>else</code> Statements and Loops in Functions</a></li>
# <li><a href="default">Setting default argument values in your custom functions</a></li>
# <li><a href="global">Global variables</a></li>
# <li><a href="scope">Scope of a Variable</a></li>
# <li>
# <a href="#quiz">Quiz on Loops</a>
# </li>
# </ul>
# <p>
# Estimated time needed: <strong>40 min</strong>
# </p>
# </div>
#
# <hr>
# <h2 id="func">Functions</h2>
# A function is a reusable block of code which performs operations specified in the function. They let you break down tasks and allow you to reuse your code in different programs.
#
# There are two types of functions :
#
# - <b>Pre-defined functions</b>
# - <b>User defined functions</b>
# <h3 id="content">What is a Function?</h3>
# You can define functions to provide the required functionality. Here are simple rules to define a function in Python:
# - Functions blocks begin <code>def</code> followed by the function <code>name</code> and parentheses <code>()</code>.
# - There are input parameters or arguments that should be placed within these parentheses.
# - You can also define parameters inside these parentheses.
# - There is a body within every function that starts with a colon (<code>:</code>) and is indented.
# - You can also place documentation before the body
# - The statement <code>return</code> exits a function, optionally passing back a value
#
# An example of a function that adds on to the parameter <code>a</code> prints and returns the output as <code>b</code>:
# +
# First function example: Add 1 to a and store as b
def add(a):
b = a + 1
print(a, "if you add one", b)
return(b)
# -
# The figure below illustrates the terminology:
# <img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/PY0101EN/Chapter%203/Images/FuncsDefinition.png" width="500" />
# We can obtain help about a function :
# +
# Get a help on add function
help(add)
# -
# We can call the function:
# +
# Call the function add()
add(1)
# -
# If we call the function with a new input we get a new result:
# +
# Call the function add()
add(2)
# -
# We can create different functions. For example, we can create a function that multiplies two numbers. The numbers will be represented by the variables <code>a</code> and <code>b</code>:
# +
# Define a function for multiple two numbers
def Mult(a, b):
c = a * b
return(c)
# -
# The same function can be used for different data types. For example, we can multiply two integers:
#
# +
# Use mult() multiply two integers
Mult(2, 3)
# -
# Two Floats:
# +
# Use mult() multiply two floats
Mult(10.0, 3.14)
# -
# We can even replicate a string by multiplying with an integer:
# +
# Use mult() multiply two different type values together
Mult(2, "<NAME> ")
# -
# <h3 id="var">Variables</h3>
# The input to a function is called a formal parameter.
#
# A variable that is declared inside a function is called a local variable. The parameter only exists within the function (i.e. the point where the function starts and stops).
#
# A variable that is declared outside a function definition is a global variable, and its value is accessible and modifiable throughout the program. We will discuss more about global variables at the end of the lab.
#
# +
# Function Definition
def square(a):
# Local variable b
b = 1
c = a * a + b
print(a, "if you square + 1", c)
return(c)
# -
# The labels are displayed in the figure:
# <img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/PY0101EN/Chapter%203/Images/FuncsVar.png" width="500" />
# We can call the function with an input of <b>3</b>:
# +
# Initializes Global variable
x = 3
# Makes function call and return function a y
y = square(x)
y
# -
# We can call the function with an input of <b>2</b> in a different manner:
# +
# Directly enter a number as parameter
square(2)
# -
# If there is no <code>return</code> statement, the function returns <code>None</code>. The following two functions are equivalent:
# +
# Define functions, one with return value None and other without return value
def MJ():
print('<NAME>')
def MJ1():
print('<NAME>')
return(None)
# +
# See the output
MJ()
# +
# See the output
MJ1()
# -
# Printing the function after a call reveals a **None** is the default return statement:
# +
# See what functions returns are
print(MJ())
print(MJ1())
# -
# Create a function <code>con</code> that concatenates two strings using the addition operation:
# +
# Define the function for combining strings
def con(a, b):
return(a + b)
# +
# Test on the con() function
con("This ", "is")
# -
# <hr/>
# <div class="alert alert-success alertsuccess" style="margin-top: 20px">
# <h4> [Tip] How do I learn more about the pre-defined functions in Python? </h4>
# <p>We will be introducing a variety of pre-defined functions to you as you learn more about Python. There are just too many functions, so there's no way we can teach them all in one sitting. But if you'd like to take a quick peek, here's a short reference card for some of the commonly-used pre-defined functions: <a href="http://www.astro.up.pt/~sousasag/Python_For_Astronomers/Python_qr.pdf">Reference</a></p>
# </div>
# <hr/>
# <h3 id="simple">Functions Make Things Simple</h3>
# Consider the two lines of code in <b>Block 1</b> and <b>Block 2</b>: the procedure for each block is identical. The only thing that is different is the variable names and values.
# <h4>Block 1:</h4>
# +
# a and b calculation block1
a1 = 4
b1 = 5
c1 = a1 + b1 + 2 * a1 * b1 - 1
if(c1 < 0):
c1 = 0
else:
c1 = 5
c1
# -
# <h4>Block 2:</h4>
# +
# a and b calculation block2
a2 = 0
b2 = 0
c2 = a2 + b2 + 2 * a2 * b2 - 1
if(c2 < 0):
c2 = 0
else:
c2 = 5
c2
# -
# We can replace the lines of code with a function. A function combines many instructions into a single line of code. Once a function is defined, it can be used repeatedly. You can invoke the same function many times in your program. You can save your function and use it in another program or use someone else’s function. The lines of code in code <b>Block 1</b> and code <b>Block 2</b> can be replaced by the following function:
# +
# Make a Function for the calculation above
def Equation(a,b):
c = a + b + 2 * a * b - 1
if(c < 0):
c = 0
else:
c = 5
return(c)
# -
# This function takes two inputs, a and b, then applies several operations to return c.
# We simply define the function, replace the instructions with the function, and input the new values of <code>a1</code>, <code>b1</code> and <code>a2</code>, <code>b2</code> as inputs. The entire process is demonstrated in the figure:
# <img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/PY0101EN/Chapter%203/Images/FuncsPros.gif" width="850" />
# Code **Blocks 1** and **Block 2** can now be replaced with code **Block 3** and code **Block 4**.
# <h4>Block 3:</h4>
a1 = 4
b1 = 5
c1 = Equation(a1, b1)
c1
# <h4>Block 4:</h4>
a2 = 0
b2 = 0
c2 = Equation(a2, b2)
c2
# <hr>
# <h2 id="pre">Pre-defined functions</h2>
# There are many pre-defined functions in Python, so let's start with the simple ones.
# The <code>print()</code> function:
# +
# Build-in function print()
album_ratings = [10.0, 8.5, 9.5, 7.0, 7.0, 9.5, 9.0, 9.5]
print(album_ratings)
# -
# The <code>sum()</code> function adds all the elements in a list or tuple:
# +
# Use sum() to add every element in a list or tuple together
sum(album_ratings)
# -
# The <code>len()</code> function returns the length of a list or tuple:
# +
# Show the length of the list or tuple
len(album_ratings)
# -
# <h2 id="if">Using <code>if</code>/<code>else</code> Statements and Loops in Functions</h2>
# The <code>return()</code> function is particularly useful if you have any IF statements in the function, when you want your output to be dependent on some condition:
# +
# Function example
def type_of_album(artist, album, year_released):
print(artist, album, year_released)
if year_released > 1980:
return "Modern"
else:
return "Oldie"
x = type_of_album("<NAME>", "Thriller", 1980)
print(x)
# -
# We can use a loop in a function. For example, we can <code>print</code> out each element in a list:
# +
# Print the list using for loop
def PrintList(the_list):
for element in the_list:
print(element)
# +
# Implement the printlist function
PrintList(['1', 1, 'the man', "abc"])
# -
# <hr>
# <h2 id="default">Setting default argument values in your custom functions</h2>
# You can set a default value for arguments in your function. For example, in the <code>isGoodRating()</code> function, what if we wanted to create a threshold for what we consider to be a good rating? Perhaps by default, we should have a default rating of 4:
# +
# Example for setting param with default value
def isGoodRating(rating=4):
if(rating < 7):
print("this album sucks it's rating is",rating)
else:
print("this album is good its rating is",rating)
# +
# Test the value with default value and with input
isGoodRating()
isGoodRating(10)
# -
# <hr>
# <h2 id="global">Global variables</h2>
# So far, we've been creating variables within functions, but we have not discussed variables outside the function. These are called global variables.
# <br>
# Let's try to see what <code>printer1</code> returns:
# +
# Example of global variable
artist = "<NAME>"
def printer1(artist):
internal_var = artist
print(artist, "is an artist")
printer1(artist)
# -
# If we print <code>internal_var</code> we get an error.
# <b>We got a Name Error: <code>name 'internal_var' is not defined</code>. Why?</b>
#
# It's because all the variables we create in the function is a <b>local variable</b>, meaning that the variable assignment does not persist outside the function.
#
# But there is a way to create <b>global variables</b> from within a function as follows:
# +
artist = "<NAME>"
def printer(artist):
global internal_var
internal_var= "<NAME>"
print(artist,"is an artist")
printer(artist)
printer(internal_var)
# -
# <h2 id="scope">Scope of a Variable</h2>
# The scope of a variable is the part of that program where that variable is accessible. Variables that are declared outside of all function definitions, such as the <code>myFavouriteBand</code> variable in the code shown here, are accessible from anywhere within the program. As a result, such variables are said to have global scope, and are known as global variables.
# <code>myFavouriteBand</code> is a global variable, so it is accessible from within the <code>getBandRating</code> function, and we can use it to determine a band's rating. We can also use it outside of the function, such as when we pass it to the print function to display it:
# +
# Example of global variable
myFavouriteBand = "AC/DC"
def getBandRating(bandname):
if bandname == myFavouriteBand:
return 10.0
else:
return 0.0
print("AC/DC's rating is:", getBandRating("AC/DC"))
print("Deep Purple's rating is:",getBandRating("Deep Purple"))
print("My favourite band is:", myFavouriteBand)
# -
# Take a look at this modified version of our code. Now the <code>myFavouriteBand</code> variable is defined within the <code>getBandRating</code> function. A variable that is defined within a function is said to be a local variable of that function. That means that it is only accessible from within the function in which it is defined. Our <code>getBandRating</code> function will still work, because <code>myFavouriteBand</code> is still defined within the function. However, we can no longer print <code>myFavouriteBand</code> outside our function, because it is a local variable of our <code>getBandRating</code> function; it is only defined within the <code>getBandRating</code> function:
# +
# Example of local variable
def getBandRating(bandname):
myFavouriteBand = "AC/DC"
if bandname == myFavouriteBand:
return 10.0
else:
return 0.0
print("AC/DC's rating is: ", getBandRating("AC/DC"))
print("Deep Purple's rating is: ", getBandRating("Deep Purple"))
print("My favourite band is", myFavouriteBand)
# -
# Finally, take a look at this example. We now have two <code>myFavouriteBand</code> variable definitions. The first one of these has a global scope, and the second of them is a local variable within the <code>getBandRating</code> function. Within the <code>getBandRating</code> function, the local variable takes precedence. **Deep Purple** will receive a rating of 10.0 when passed to the <code>getBandRating</code> function. However, outside of the <code>getBandRating</code> function, the <code>getBandRating</code> s local variable is not defined, so the <code>myFavouriteBand</code> variable we print is the global variable, which has a value of **AC/DC**:
# +
# Example of global variable and local variable with the same name
myFavouriteBand = "AC/DC"
def getBandRating(bandname):
myFavouriteBand = "Deep Purple"
if bandname == myFavouriteBand:
return 10.0
else:
return 0.0
print("AC/DC's rating is:",getBandRating("AC/DC"))
print("Deep Purple's rating is: ",getBandRating("Deep Purple"))
print("My favourite band is:",myFavouriteBand)
# -
# <h2>Quiz on Functions</h2>
# Come up with a function that divides the first input by the second input:
# Write your code below and press Shift+Enter to execute
def div(a, b):
return(a/b)
# Double-click __here__ for the solution.
#
# <!--
# def div(a, b):
# return(a/b)
# -->
# <hr>
# Use the function <code>con</code> for the following question.
# +
# Use the con function for the following question
def con(a, b):
return(a + b)
# -
# Can the <code>con</code> function we defined before be used to add to integers or strings?
# Write your code below and press Shift+Enter to execute
#yes, for example:
con(2, 2)
# Double-click __here__ for the solution.
#
# <!--
# yes, for example:
# con(2, 2)
# -->
# <hr>
# Can the <code>con</code> function we defined before be used to concentrate a list or tuple?
# Write your code below and press Shift+Enter to execute
#yes,for example:
con(['a', 1], ['b', 1])
# Double-click __here__ for the solution.
#
# <!--
# yes,for example:
# con(['a', 1], ['b', 1])
# -->
# <hr>
# <h2>The last exercise!</h2>
# <p>Congratulations, you have completed your first lesson and hands-on lab in Python. However, there is one more thing you need to do. The Data Science community encourages sharing work. The best way to share and showcase your work is to share it on GitHub. By sharing your notebook on GitHub you are not only building your reputation with fellow data scientists, but you can also show it off when applying for a job. Even though this was your first piece of work, it is never too early to start building good habits. So, please read and follow <a href="https://cognitiveclass.ai/blog/data-scientists-stand-out-by-sharing-your-notebooks/" target="_blank">this article</a> to learn how to share your work.
# <hr>
# <div class="alert alert-block alert-info" style="margin-top: 20px">
# <h2>Get IBM Watson Studio free of charge!</h2>
# <p><a href="https://cocl.us/bottemNotebooksPython101Coursera"><img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/PY0101EN/Ad/BottomAd.png" width="750" align="center"></a></p>
# </div>
# <h3>About the Authors:</h3>
# <p><a href="https://www.linkedin.com/in/joseph-s-50398b136/" target="_blank"><NAME></a> is a Data Scientist at IBM, and holds a PhD in Electrical Engineering. His research focused on using Machine Learning, Signal Processing, and Computer Vision to determine how videos impact human cognition. Joseph has been working for IBM since he completed his PhD.</p>
# Other contributors: <a href="www.linkedin.com/in/jiahui-mavis-zhou-a4537814a"><NAME></a>, <a href="https://www.linkedin.com/in/reevejamesd/"><NAME></a>
# <hr>
# <p>Copyright © 2018 IBM Developer Skills Network. This notebook and its source code are released under the terms of the <a href="https://cognitiveclass.ai/mit-license/">MIT License</a>.</p>
|
PY0101EN-3-3-Functions.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
from bs4 import BeautifulSoup
import requests
import pandas as pd
import re
import csv
# TODO: Auto-parse of NCAA brackets. Not necessary now since already grabbed manually.
base_url = 'https://www.sports-reference.com'
def getTeamsUrlsForYear(year):
postseason_url = '/cbb/postseason/' + year + '-ncaa.html'
web = base_url + postseason_url
html = requests.get(web).text
soup = BeautifulSoup(html, 'html5lib')
brackets_div = soup('div', {'id': 'bracket'})
team_urls = {}
for bracket in brackets_div:
# First round of each bracket
rounds = bracket.find('div', 'round')
refs = rounds.findAll('a')
for ref in refs:
if 'schools' in ref['href']:
team_urls[ref.text.strip()] = ref['href']
return team_urls
# +
def data_mapper(stats):
# 'team_g', 'team_fg', 'team_fga', 'team_fg_pct', 'team_fg2', 'team_fg2a', 'team_fg2_pct',
# 'team_fg3', 'team_fg3a', 'team_fg3_pct', 'team_ft', 'team_fta', 'team_ft_pct', 'team_orb', 'team_drb',
# 'team_trb', 'team_ast', 'team_stl', 'team_blk', 'team_tov', 'team_pf', 'team_pts', 'opp_opp_fg',
# 'opp_opp_fga', 'opp_opp_fg_pct', 'opp_opp_fg2', 'opp_opp_fg2a', 'opp_opp_fg2_pct', 'opp_opp_fg3',
# 'opp_opp_fg3a', 'opp_opp_fg3_pct', 'opp_opp_ft', 'opp_opp_fta', 'opp_opp_ft_pct', 'opp_opp_orb',
# 'opp_opp_drb', 'opp_opp_trb', 'opp_opp_ast', 'opp_opp_stl', 'opp_opp_blk', 'opp_opp_tov',
# 'opp_opp_pf', 'opp_opp_pts'
delim = ','
data_string = stats['team_g'] + delim + stats['team_fg'] + delim + stats['team_fga'] + delim + \
stats['team_fg_pct'] + delim + stats['team_fg2'] + delim + stats['team_fg2a'] + delim + \
stats['team_fg2_pct'] + delim + stats['team_fg3'] + delim + stats['team_fg3a'] + delim + \
stats['team_fg3_pct'] + delim + stats['team_ft'] + delim + stats['team_fta'] + delim + \
stats['team_ft_pct'] + delim + stats['team_orb'] + delim + stats['team_drb'] + delim + \
stats['team_trb'] + delim + stats['team_ast'] + delim + stats['team_stl'] + delim + \
stats['team_blk'] + delim + stats['team_tov'] + delim + stats['team_pf'] + delim + \
stats['team_pts'] + delim + stats['opp_opp_fg'] + delim + stats['opp_opp_fga'] + delim + \
stats['opp_opp_fg_pct'] + delim + stats['opp_opp_fg2'] + delim + stats['opp_opp_fg2a'] + delim + \
stats['opp_opp_fg2_pct'] + delim + stats['opp_opp_fg3'] + delim + stats['opp_opp_fg3a'] + delim + \
stats['opp_opp_fg3_pct'] + delim + stats['opp_opp_ft'] + delim + stats['opp_opp_fta'] + delim + \
stats['opp_opp_ft_pct'] + delim + stats['opp_opp_orb'] + delim + stats['opp_opp_drb'] + delim + \
stats['opp_opp_trb'] + delim + stats['opp_opp_ast'] + delim + stats['opp_opp_stl'] + delim +\
stats['opp_opp_blk'] + delim + stats['opp_opp_tov'] + delim + stats['opp_opp_pf'] + delim +\
stats['opp_opp_pts'] + '\n'
return data_string
base_url = 'https://www.sports-reference.com'
def getTeamsData(teams_urls):
teams_data = {}
for team_name, team_url in teams_urls.items():
web = base_url + team_url
html = requests.get(web).text
soup = BeautifulSoup(html, 'html5lib')
table = soup.find('div', {'id': 'div_schools_totals', 'class': 'table_container'})
thead = table.find('thead')
tbody = table.find('tbody')
trows = tbody.findAll('tr')
stats = {}
for trow in trows:
if trow.find('th').text == "Team":
tds = trow.findAll('td')
prefix = 'team_'
elif trow.find('th').text == "Opponent":
tds = trow.findAll('td')
prefix = 'opp_'
else:
continue
for td in tds:
stats[prefix+td['data-stat']] = td.text
stat_string = data_mapper(stats)
teams_data[team_name] = stat_string
return teams_data
# +
delim = ','
def createNewFileWithTeamData(in_filename, out_filename, teams_data):
in_file = open(in_filename, 'r')
out_file = open(out_filename, 'w')
header = in_file.readline()
out_file.write(header)
for line in in_file:
try:
line = line.split(',')
newline = line[0] + delim + line[1] + delim + line[2] + delim + line[3] + delim + teams_data[line[1]]
out_file.write(newline)
except:
print('KEYS:', teams_data.keys())
print(line[1])
break
in_file.close()
out_file.close()
# -
years = ['1997', '1998', '1999', '2000', '2001', '2002', '2003', '2004', '2005', '2006', '2007', '2008', '2009', '2010', '2011', '2012', '2013', '2014', '2015', '2016', '2017', '2018', '2019', '2021']
for year in years:
old_filename = './NCAA-Tournaments/' + year + '-NCAA-Tournament.xlsx'
read_file = pd.read_excel(old_filename)
new_filename = './NCAA-Tournaments/' + year + '-NCAA-Tournament.csv'
read_file.to_csv(new_filename, index = None, header=True)
years = ['1997', '1998', '1999', '2000', '2001', '2002', '2003', '2004', '2005', '2006', '2007', '2008', '2009', '2010', '2011', '2012', '2013', '2014', '2015', '2016', '2017', '2018', '2019', '2021']
for year in years:
print('NOW ON YEAR: ', year)
in_filename = './NCAA-Tournaments/' + year + '-NCAA-Tournament.csv'
out_filename = './NCAA-Tournaments/' + year + '-NCAA-Tournament-updated.csv'
teams_urls = getTeamsUrlsForYear(year)
teams_data = getTeamsData(teams_urls)
createNewFileWithTeamData(in_filename, out_filename, teams_data)
print('----------------------------------------')
years = ['1997', '1998', '1999', '2000', '2001', '2002', '2003', '2004', '2005', '2006', '2007', '2008', '2009', '2010', '2011', '2012', '2013', '2014', '2015', '2016', '2017', '2018', '2019', '2021']
out_filename = './Consolidated-NCAA-Tournament-data.csv'
out_file = open(out_filename, 'w')
delim = ','
for year in years:
print('NOW ON YEAR: ', year)
in_filename = './NCAA-Tournaments/' + year + '-NCAA-Tournament-updated.csv'
in_file = open(in_filename, 'r')
if year == '1997':
line = 'YEAR' + delim + in_file.readline()
out_file.write(line)
else:
in_file.readline()
for line in in_file:
out_file.write(year + delim + line)
in_file.close()
print('----------------------------------------')
out_file.close()
df = pd.read_csv('./Consolidated-NCAA-Tournament-data.csv')
df
df['YEAR']
df_keys_for_per_game = ['TEAM_FG', 'TEAM_FGA', 'TEAM_2P', 'TEAM_2PA', 'TEAM_3P',
'TEAM_3PA', 'TEAM_FT', 'TEAM_FTA', 'TEAM_ORB', 'TEAM_DRB', 'TEAM_TRB', 'TEAM_AST', 'TEAM_STL', 'TEAM_BLK',
'TEAM_TOV','TEAM_PF', 'TEAM_PTS', 'OPP_FG', 'OPP_FGA', 'OPP_2P', 'OPP_2PA', 'OPP_3P', 'OPP_3PA', 'OPP_FT',
'OPP_FTA', 'OPP_FT%', 'OPP_ORB', 'OPP_DRB', 'OPP_TRB', 'OPP_AST', 'OPP_STL', 'OPP_BLK', 'OPP_TOV',
'OPP_PF', 'OPP_PTS']
for key in df_keys_for_per_game:
df[key] = df[key] / df['GAMES']
df
df.to_csv('./consolidated-ncaa-tournament-per-game-data.csv', index = None, header=True)
|
ncaa-data-processing.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Imports
# +
import pandas as pd
import time
import numpy as np
import datetime as dt
import seaborn as sns
import matplotlib.pyplot as plt
import utilities.densmore_v3 as dns
# -
df = pd.read_csv('../data/api_football_data/05_merged_data/elo_merged_data_names_short.csv')
df.head()
# # SETUP
# ### ELO Model Equation
# **Where:**
#
# | Ratings | Stats | Coefficients|
# |:-----------------------------------|:------------------------------|:------------|
# | $R_{old}$ = Team's previous rating | $GF - GA$ = Goal differential | $K$ = Impact of all terms on team's new rating |
# | $R_{new}$ = Team's new rating | $R_{opp} - R_{old}$ = Matchup difficulty | $C$ = Spread of all ratings |
# | $R_{opp}$ = Opponent's rating | | |
# | | | |
# **FIRST GAME OF SEASON**
# $$\large R_{new} = R_{start} + K \begin{pmatrix} GF - GA + \frac{R_{opp} - R_{old}}{C} \end{pmatrix} $$
#
# **SUBSEQUENT GAMES**
#
# $$\large R_{new} = R_{old} + K \begin{pmatrix} GF - GA + \frac{R_{opp} - R_{old}}{C} \end{pmatrix} $$
# ```python
# new_h_rating = h_rating + K * (h_goals - a_goals + ((a_rating - h_rating) / C))
# new_a_rating = a_rating + K * (a_goals - h_goals + ((h_rating - a_rating) / C))
# ````
# ### Reference Variables
# +
teams = ['atl', 'chi', 'cin', 'cor', 'crw', 'dcu',
'fcd', 'hou', 'lafc', 'lag', 'min', 'mtl',
'ner', 'nyc', 'nyrb', 'orl', 'phi', 'por',
'rsl', 'sea', 'sje', 'skc', 'tfc', 'van']
years = [2016, 2017, 2018, 2019]
# -
# ### Add Game Numbers by Season
# #### Splitt Data by Season
df_2016 = df[df['season'] == 2016].copy()
df_2017 = df[df['season'] == 2017].copy()
df_2018 = df[df['season'] == 2018].copy()
df_2019 = df[df['season'] == 2019].copy()
# +
# df_2016
# df_2017
# df_2018
# df_2019
# -
# #### Add Game Numbers
def add_game_nums(season_df):
game_counts = {team: 1 for team in teams}
game_nums_by_id = []
for row, data in season_df.iterrows():
for team, game_num in game_counts.items():
if team == data['h_name']:
game_counts[team] += 1
data['h_game_num'] = game_num
elif team == data['a_name']:
game_counts[team] += 1
data['a_game_num'] = game_num
else:
continue
game_nums_by_id.append({'game_id': data['game_id'],
'h_game_num': data['h_game_num'],
'a_game_num': data['a_game_num']})
# print(f"{data['h_name']}: {data['h_game_num']}", f"{data['a_name']}: {data['a_game_num']}")
game_nums_df = pd.DataFrame(game_nums_by_id, index=range(len(game_nums_by_id)))
new_df = pd.merge(left=season_df, right=game_nums_df, on='game_id')
return new_df
df_2016 = add_game_nums(df_2016)
df_2017 = add_game_nums(df_2017)
df_2018 = add_game_nums(df_2018)
df_2019 = add_game_nums(df_2019)
df_2016.index = df_2016.index + 1
df_2017.index = df_2017.index + 1
df_2018.index = df_2018.index + 1
df_2019.index = df_2019.index + 1
# #### Re-join Tables
df = pd.concat([df_2016, df_2017, df_2018, df_2019], axis=0)
df.drop(columns=['h_possession', 'h_pass_acc', 'a_possession', 'a_pass_acc'], inplace=True)
df.columns
df = df[['result', 'game_id', 'season', 'game_date',
'h_name', 'h_id', 'h_goals', 'h_game_num',
'a_name', 'a_id', 'a_goals', 'a_game_num']]
df.to_csv('../data/api_football_data/05_merged_data/elo_merged_data_ready.csv')
# ### Setting Up ELO Table
# #### Method 1 - As dictionary first, then convert to df
# +
# elo_dict = {'game_num' : 0,
# 'season' : 0,
# 'new_false_preds': 0,
# 'total_false_preds': 0}
# for team in teams:
# elo_dict[team] = [0]
# elo_table = pd.DataFrame(elo_dict)
# for i in range(1,136):
# elo_table = elo_table.append(pd.Series(name=i, dtype=int)).fillna(0).astype(int)
# elo_table.index = elo_table.index + 1
# +
# elo_table.head()
# -
# #### Method 2 (better) - As Dataframe
elo_table = pd.DataFrame(columns=['new_false_preds', 'total_false_preds'] + [team for team in teams])
for i in range(1,137):
elo_table = elo_table.append(pd.Series(name=i, dtype=int)).fillna(0).astype(int)
indeces = list(zip([2016]*34 + [2017]*34 + [2018]*34 + [2019]*34, list(range(1,35)) * 4))
elo_table.index = pd.MultiIndex.from_tuples(indeces, names=['year', 'game'])
elo_table.head()
# ### Rating Update Function
# +
def update_ratings(h_rating, h_goals, a_rating, a_goals, K=20, C=200):
if h_rating > a_rating:
expected_result = 'home'
elif h_rating < a_rating:
expected_result = 'away'
else:
expected_result = np.random.Generator.choice(['home', 'away'])
if h_goals > a_goals:
actual_result = 'home'
elif h_goals < a_goals:
actual_result = 'away'
else:
actual_result = 'draw'
new_h_rating = h_rating + K * (h_goals - a_goals + ((a_rating - h_rating) / C))
new_a_rating = a_rating + K * (a_goals - h_goals + ((h_rating - a_rating) / C))
# new_h_rating = h_rating + K * (h_goals - a_goals + ((a_rating - h_rating) / C) + D * (h_ball_poss + h_pass_acc))
# new_a_rating = a_rating + K * (a_goals - h_goals + ((h_rating - a_rating) / C) + D * (a_ball_poss + a_pass_acc))
if expected_result != actual_result:
false_pred = 1
else:
false_pred = 0
return new_h_rating, new_a_rating, false_pred, expected_result, actual_result
# -
def create_elo_table(games=df, K=30, C=300):
log = []
# establish list of team name abbreviations
teams = ['atl', 'chi', 'cin', 'cor', 'crw', 'dcu',
'fcd', 'hou', 'lafc', 'lag', 'min', 'mtl',
'ner', 'nyc', 'nyrb', 'orl', 'phi', 'por',
'rsl', 'sea', 'sje', 'skc', 'tfc', 'van']
# create elo table with necessary columns
elo_table = pd.DataFrame(columns=['new_false_preds', 'total_false_preds'] + [team for team in teams])
# fill 136 rows with 0s
for i in range(1,137):
elo_table = elo_table.append(pd.Series(name=i, dtype=int)).fillna(0).astype(int)
# set up MultiIndex - will allow me to update specific values precisely
indeces = list(zip([2016]*34 + [2017]*34 + [2018]*34 + [2019]*34, list(range(1,35)) * 4))
elo_table.index = pd.MultiIndex.from_tuples(indeces, names=['year', 'game'])
# start a running total of false predictions
false_preds_count = 0
# loop through match data and updating
for row, data in games.iterrows():
# set variables for each game - for easier referencing in code below
h_team, a_team = data['h_name'], data['a_name']
h_game_num, a_game_num = data['h_game_num'], data['a_game_num']
h_goals, a_goals = data['h_goals'], data['a_goals']
season = data['season']
# collect each team's previous rating
try:
h_prev_rating = elo_table.at[(season, h_game_num-1), h_team]
except:
h_prev_rating = 500
try:
a_prev_rating = elo_table.at[(season, a_game_num-1), a_team]
except:
a_prev_rating = 500
# running 'update_rating' function and saving results
new_h_rating, new_a_rating, false_pred, expected_result, actual_result = \
update_rating(h_prev_rating, h_goals, a_prev_rating, a_goals, K, C)
# update running count of total false predictions
false_preds_count += false_pred
# adding results to elo_table if game number is the same for both teams
if h_game_num == a_game_num:
game_num = h_game_num
elo_table.at[(season, game_num), h_team] = new_h_rating
elo_table.at[(season, game_num), a_team] = new_a_rating
elo_table.at[(season, game_num), 'new_false_preds'] += false_pred
elo_table.at[(season, game_num), 'total_false_preds'] = false_preds_count
# adding results to elo_table if game number is different for each team
else:
# home
elo_table.at[(season, h_game_num), h_team] = new_h_rating
elo_table.at[(season, h_game_num), 'new_false_preds'] += false_pred
elo_table.at[(season, h_game_num), 'total_false_preds'] = false_preds_count
# away
elo_table.at[(season, a_game_num), a_team] = new_a_rating
elo_table.at[(season, a_game_num), 'new_false_preds'] += false_pred
elo_table.at[(season, a_game_num), 'total_false_preds'] = false_preds_count
log.append({'match_id' : data['game_id'],
'season' : season,
'home_team' : h_team,
'home_game_num': h_game_num,
'away_team' : a_team,
'away_game_num' : a_game_num,
'result_expected' : expected_result,
'result_actual' : actual_result,
'false_pred' : false_pred})
return elo_table, pd.DataFrame(log)
elo_table_filled, elo_table_log = create_elo_table()
elo_table_filled.to_csv('../data/api_football_data/05_merged_data/elo_table_filled.csv')
elo_table_log.to_csv('../data/api_football_data/05_merged_data/elo_table_log.csv')
elo_table_log['false_pred'].value_counts(normalize=True)
# #### Code Testing
update_rating(723, 3, 438, 0, K=20, C=200)
update_rating(723, 0, 438, 3, K=20, C=200)
723-754.5, 723-634.5, 438-406.5, 438-526.5
# ## Saved Code
# +
# Realized an easier way to handle first games of the season. Using try/except to set previous ratings.
# When doing it for game 1, 1-1 will be 0, which will raise a 'key' error. If an error occurs for one
# or both of the teams their previous_rating variable will be set to 500. There shouldnt be errors
# otherwise.
# saving this code for safety
# # first game - update each team's game 1 using predetermined ratings at the start of each season
# if h_game_num == 1 or a_game_num == 1:
# # running 'update_rating' function and saving results
# new_h_rating, new_a_rating, false_pred, expected_result, actual_result = \
# update_rating(500, h_goals, 500, a_goals, K, C)
# # add results to elo_table if game number is the same for both teams
# if h_game_num == a_game_num:
# game_num = h_game_num
# elo_table.at[(season, game_num), h_team] = new_h_rating
# elo_table.at[(season, game_num), a_team] = new_a_rating
# elo_table.at[(season, game_num), 'new_false_preds'] += false_pred
# elo_table.at[(season, game_num), 'total_false_preds'] = false_preds_count
# # add results to elo_table if game numbers aren't the same (when number of teams is odd)
# else:
# # home
# elo_table.at[(season, h_game_num), h_team] = new_h_rating
# elo_table.at[(season, h_game_num), 'new_false_preds'] += false_pred
# elo_table.at[(season, h_game_num), 'total_false_preds'] = false_preds_count
# # away
# elo_table.at[(season, a_game_num), a_team] = new_a_rating
# elo_table.at[(season, a_game_num), 'new_false_preds'] += false_pred
# elo_table.at[(season, a_game_num), 'total_false_preds'] = false_preds_count
# -
# ### ELO Model Equation
# **Where:**
#
# | Ratings | Stats | Coefficients|
# |:---------|:------|:------------|
# | $R_{start}$ = 500 | $Pass\%$ = Pass accuracy percentage | $K$ = Impact of all terms on team's new rating |
# | $R_{old}$ = Team's previous rating | $Pos.\%$ = Possession percentage | $C$ = Spread of all ratings |
# | $R_{new}$ = Team's new rating | $GF - GA$ = Goal differential | $D$ = Impact of misc stats on rating ($Pass\%$ and $Pos.\%$) |
# | $R_{opp}$ = Opponent's rating | $R_{opp} - R_{old}$ = Matchup difficulty | |
# **FIRST GAME OF SEASON**
# $$\large R_{new} = R_{start} + K \begin{pmatrix} GF - GA + \frac{R_{opp} - R_{old}}{C} + D( Pass\% + Pos.\%) \end{pmatrix} $$
#
# **SUBSEQUENT GAMES**
#
# $$\large R_{new} = R_{old} + K \begin{pmatrix} GF - GA + \frac{R_{opp} - R_{old}}{C} + D( Pass\% + Pos.\%) \end{pmatrix} $$
|
code/04_ELO_Model.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Euler Method
# The Euler method works by assuming the gradient of a solution, $y'=f(t_n,y_n)$ can be approximated by a linear gradient between $y_n$ and $y_{n+1}$, i.e.,
#
# $$ \frac{y_{n+1}-y_n}{\Delta t} = f(t_n,y_n)$$
#
# so $y_{n+1}$ can be expressed as,
#
# $$ y_{n+1} = y_n+\Delta t \; f(t_n,y_n)$$
#
# The example below shows the Euler method solution for $y'=y+t$, change the slider to increase the number of solution points (thereby decreasing $\Delta t$) and watch what happens to the root-mean-square (RMS) error of the numerical solution compared to the analytic solution.
# +
# %matplotlib inline
# import packages
from matplotlib import pyplot as plt
import numpy as np
import time
from scipy import integrate
from ipywidgets import interact
# define ODE
def ode(y, t):
return y+t
# define plot function
def plotter(n):
# dt, time
dt = 1/(n-1);
t = np.linspace(0, 1, n)
# solution loop
y = np.zeros([n])
y_a = y;
y[0] = 0;
err = 0;
for i in range(1,n):
f_e = ode(y[i-1],t[i-1])
y_e = y[i-1]+dt*f_e
#y[i] = y[i-1]+dt*(ode(y_e,t[i])+f_e)/2
y[i] = y[i-1]+dt*ode(y[i-1],t[i-1])
err = err+(abs(y[i]-(np.exp(t[i])-t[i]-1)))**2
err = (err/n)**0.5;
#analytic solution
t_a = np.linspace(0,1,101)
y_a = np.zeros(len(t_a))
for j in range(1,len(t_a)):
y_a[j] = np.exp(t_a[j])-t_a[j]-1
f,ax = plt.subplots(1,1, figsize=(14,8))
ax.plot(t_a,y_a, color='b', label='Analytic')
ax.plot(t,y, 'o',color='r', label='Euler')
plt.title('RMS error %.4f' %err)
legend = ax.legend(loc='upper left', shadow=False)
#plt.yticks(np.arange(-1.2, 1.3, 0.4))
plt.xlabel('t')
plt.ylabel('y')
#ax.plot([t_i],y[1,0],marker='o', markersize=3, color="red")
plt.show()
interact(plotter, n =(2,21,1))
|
engsci211_ode3_euler.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Model Tuning
# > A Summary of lecture "Machine Learning with Tree-Based Models in Python", via datacamp
#
# - toc: true
# - badges: true
# - comments: true
# - author: <NAME>
# - categories: [Python, Datacamp, Machine_Learning]
# - image:
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# ## Tuning a CART's Hyperparameters
# - Hyperparameters
# - Machine learning model:
# - parameters: learned from data
# - CART example: split-point of a node, split-feature of a node, ...
# - hyperparameters: not learned from data, set prior to training
# - CART example: ```max_depth```, ```min_samples_leaf```, splitting criterion, ...
# - What is hyperparameter tuning?
# - Problem: search for a set of optimal hyperparameters for a learning algorithm.
# - Solution: find a set of optimal hyperparameters that results in an optimal model.
# - Optimal model: yields an optimal score
# - Score : defaults to accuracy (classification) and $R^2$ (regression)
# - Cross-validation is used to estimate the generalization performance.
# - Approaches to hyperparameter tuning
# - Grid Search
# - Random Search
# - Bayesian Optimization
# - Genetic Algorithm
# - ...
# - Grid search cross validation
# - Manually set a grid of discrete hyperparameter values.
# - Set a metric for scoring model performance.
# - Search exhaustively through the grid.
# - For each set of hyperparameters, evaluate each model's CV score
# - The optimal hyperparameters are those of the model achieving the best CV score.
# ### Tree hyperparameters
# In the following exercises you'll revisit the [Indian Liver Patient](https://www.kaggle.com/uciml/indian-liver-patient-records) dataset which was introduced in a previous chapter.
#
# Your task is to tune the hyperparameters of a classification tree. Given that this dataset is imbalanced, you'll be using the ROC AUC score as a metric instead of accuracy.
# - Preprocess
indian = pd.read_csv('./datasets/indian_liver_patient_preprocessed.csv', index_col=0)
indian.head()
X = indian.drop('Liver_disease', axis='columns')
y = indian['Liver_disease']
# +
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=2)
# +
from sklearn.tree import DecisionTreeClassifier
# Instantiate dt
dt = DecisionTreeClassifier()
# Check default hyperparameter
dt.get_params()
# -
# ### Set the tree's hyperparameter grid
# In this exercise, you'll manually set the grid of hyperparameters that will be used to tune the classification tree ```dt``` and find the optimal classifier in the next exercise.
#
#
# Define params_dt
params_dt = {
'max_depth': [2, 3, 4],
'min_samples_leaf': [0.12, 0.14, 0.16, 0.18],
}
# ### Search for the optimal tree
# In this exercise, you'll perform grid search using 5-fold cross validation to find ```dt```'s optimal hyperparameters. Note that because grid search is an exhaustive process, it may take a lot time to train the model. Here you'll only be instantiating the ```GridSearchCV``` object without fitting it to the training set. As discussed in the video, you can train such an object similar to any scikit-learn estimator by using the ```.fit()``` method:
# ```python
# grid_object.fit(X_train, y_train)
# ```
#
# +
from sklearn.model_selection import GridSearchCV
# Instantiate grid_dt
grid_dt = GridSearchCV(estimator=dt, param_grid=params_dt, scoring='roc_auc', cv=5, n_jobs=-1)
grid_dt.fit(X_train, y_train)
# -
# ### Evaluate the optimal tree
# In this exercise, you'll evaluate the test set ROC AUC score of grid_dt's optimal model.
#
# In order to do so, you will first determine the probability of obtaining the positive label for each test set observation. You can use the method ```predict_proba()``` of an sklearn classifier to compute a 2D array containing the probabilities of the negative and positive class-labels respectively along columns.
# +
from sklearn.metrics import roc_auc_score
# Extract the best estimator
best_model = grid_dt.best_estimator_
# Predict the test set probabilities of the positive class
y_pred_proba = best_model.predict_proba(X_test)[:, 1]
# Compute test_roc_auc
test_roc_auc = roc_auc_score(y_test, y_pred_proba)
# Print test_roc_auc
print("Test set ROC AUC score: {:.3f}".format(test_roc_auc))
# -
# ## Tuning a RF's Hyperparameters
# - Random Forest Hyperparameters
# - CART hyperparameters
# - number of estimators
# - Whether it uses bootstrapping or not
# - ...
# - Tuning is expensive
# - Hyperparameter tuning:
# - Computationally expensive,
# - sometimes leads to very slight improvement
# - Weight the impact of tuning on the whole project
# ### Random forests hyperparameters
# In the following exercises, you'll be revisiting the [Bike Sharing Demand](https://www.kaggle.com/c/bike-sharing-demand) dataset that was introduced in a previous chapter. Recall that your task is to predict the bike rental demand using historical weather data from the Capital Bikeshare program in Washington, D.C.. For this purpose, you'll be tuning the hyperparameters of a Random Forests regressor.
# - Preprocess
bike = pd.read_csv('./datasets/bikes.csv')
bike.head()
X = bike.drop('cnt', axis='columns')
y = bike['cnt']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=2)
# +
from sklearn.ensemble import RandomForestRegressor
# Instantiate rf
rf = RandomForestRegressor()
# Get hyperparameters
rf.get_params()
# -
# ### Set the hyperparameter grid of RF
# In this exercise, you'll manually set the grid of hyperparameters that will be used to tune ```rf```'s hyperparameters and find the optimal regressor. For this purpose, you will be constructing a grid of hyperparameters and tune the number of estimators, the maximum number of features used when splitting each node and the minimum number of samples (or fraction) per leaf.
# Define the dicrionary 'params_rf'
params_rf = {
'n_estimators': [100, 350, 500],
'max_features': ['log2', 'auto', 'sqrt'],
'min_samples_leaf': [2, 10, 30],
}
# ### Search for the optimal forest
# In this exercise, you'll perform grid search using 3-fold cross validation to find ```rf```'s optimal hyperparameters. To evaluate each model in the grid, you'll be using the negative mean squared error metric.
#
# Note that because grid search is an exhaustive search process, it may take a lot time to train the model. Here you'll only be instantiating the ```GridSearchCV``` object without fitting it to the training set. As discussed in the video, you can train such an object similar to any scikit-learn estimator by using the ```.fit()``` method:
# ```python
# grid_object.fit(X_train, y_train)
# ```
# +
from sklearn.model_selection import GridSearchCV
# Instantiate grid_rf
grid_rf = GridSearchCV(estimator=rf, param_grid=params_rf, scoring='neg_mean_squared_error', cv=3,
verbose=1, n_jobs=-1)
# fit model
grid_rf.fit(X_train, y_train)
# -
# ### Evaluate the optimal forest
# In this last exercise of the course, you'll evaluate the test set RMSE of ```grid_rf```'s optimal model.
# +
from sklearn.metrics import mean_squared_error as MSE
# Extract the best estimator
best_model = grid_rf.best_estimator_
# Predict test set labels
y_pred = best_model.predict(X_test)
# Compute rmse_test
rmse_test = MSE(y_test, y_pred) ** 0.5
# Print rmse_test
print('Test RMSE of best model: {:.3f}'.format(rmse_test))
|
_notebooks/2020-06-04-03-Model-Tuning.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Deviations from Normality
#
# _(plus python functions with default parameters plus a quick example of recursive functions)_
#
# Today, we'll develop the code for skewness and kurtosis even though these are already available in the `scipy.stats` module, and then we'll apply them to hedge fund index returns.
#
# We'll also look at using `scipy.stats` module to apply the _Jarque-Bera_ test for normality, and apply them to different return series.
#
# First, add the following code to our `edhec_risk_kit.py`
#
# ```python
# def get_hfi_returns():
# """
# Load and format the EDHEC Hedge Fund Index Returns
# """
# hfi = pd.read_csv("data/edhec-hedgefundindices.csv",
# header=0, index_col=0, parse_dates=True)
# hfi = hfi/100
# hfi.index = hfi.index.to_period('M')
# return hfi
# ```
#
# +
# %load_ext autoreload
# %autoreload 2
import pandas as pd
import edhec_risk_kit_105 as erk
hfi = erk.get_hfi_returns()
hfi.head()
# -
# ## Skewness
#
# Intuitively, a negative skew means that you get more negative returns than you would have expected if the returns were distributed like the normal distribution.
#
# Another way of thinking about it is if that returns are normally distributed, the mean and the median would be very close.
#
# However, if they are negatively skewed, the expected value i.e. the mean is less than the median. If they are positively skewed, the expected value (again, the mean) is greater than the median.
pd.concat([hfi.mean(), hfi.median(), hfi.mean()>hfi.median()], axis=1)
# Now, let's develop the code to compute the skewness of a series of numbers.
#
# Recall that the skewness is given by:
#
# $$ S(R) = \frac{E[ (R-E(R))^3 ]}{\sigma_R^3} $$
#
def skewness(r):
"""
Alternative to scipy.stats.skew()
Computes the skewness of the supplied Series or DataFrame
Returns a float or a Series
"""
demeaned_r = r - r.mean()
# use the population standard deviation, so set dof=0
sigma_r = r.std(ddof=0)
exp = (demeaned_r**3).mean()
return exp/sigma_r**3
skewness(hfi).sort_values()
# Just to see if we get the same answer, let's use the skewness function that is built into `scipy.stats`
import scipy.stats
scipy.stats.skew(hfi)
# So, let's add that to our `edhec_risk_kit.py`.
#
# Finally, let's look at the skewness that you would expect from a truly random sequence of returns. Let's use the random normal generator from numpy and generate the same number of returns as we have for the hedge fund data.
hfi.shape
import numpy as np
normal_rets = np.random.normal(0, 0.15, (263, 1))
normal_rets.mean(), normal_rets.std()
erk.skewness(normal_rets)
# # Kurtosis
#
# Intuitively, the kurtosis measures the "fatness" of the tails of the distribution. The normal distribution has a kurtosis of 3 and so if the kurtosis of your returns is less than 3 then it tends to have thinner tails, and if the kurtosis is greater than 3 then the distribution has fatter tails.
#
# Kurtosis is given by:
#
# $$ K(R) = \frac{E[ (R-E(R))^4 ]}{\sigma_R^4} $$
#
# This is very similar to the skewness, so we can just copy and paste it and then edit it to compute the 4th rather than the 3rd power (as was the case for skewness).
#
erk.kurtosis(hfi)
# Let's compare it with `scipy.stats` ...
scipy.stats.kurtosis(hfi)
# Note that these numbers are all lower by 3 from the number we have computed. That's because, as we said above, the expected kurtosis of a normally distributed series of numbers is 3, and `scipy.stats` is returning the _Excess Kurtosis_. We can see this by applying it on the random normal numbers we generated:
scipy.stats.kurtosis(normal_rets)
erk.kurtosis(normal_rets)
# ## Running the Jarque-Bera Test for Normality
#
# The `scipy.stats` module contains a function that runs the _Jarque-Bera_ test on a sequence of numbers. Let's apply that to the normally generated returns:
scipy.stats.jarque_bera(normal_rets)
# The first number is the test statistic and the second number is the one we want. It represents the p-value for the hypothesis test. If you want to run the test at a 1% level of significance, you want this number to be greater than 0.01 to accept the hypothesis that the data is normally distributed, and if that number is less than 0.01 then you must reject the hypothesis of normality.
#
# In this case, since we got a number higher than 0.01 we can accept the hypothesis that the numbers are random. Now, let's try this on our different hedge fund indices.
scipy.stats.jarque_bera(hfi)
# Why didn't we get the results for the individual indices? Because the implementation of the test isn't smart enough to realize that we want to treat each column as a separate set of returns. We can write out own wrapper for it to fix that, so let's start by writing a simple wrapper, and adding this code to our python file:
#
# ```python
# import scipy.stats
# def is_normal(r, level=0.01):
# """
# Applies the Jarque-Bera test to determine if a Series is normal or not
# Test is applied at the 1% level by default
# Returns True if the hypothesis of normality is accepted, False otherwise
# """
# statistic, p_value = scipy.stats.jarque_bera(r)
# return p_value > level
# ```
erk.is_normal(normal_rets)
# There are a few different ways to handle the problem. The first is to use the `.aggregate` method on a dataframe, that takes a function as an argument and applies that function to each column:
hfi.aggregate(erk.is_normal)
# However, we can fix this in our wrapper so that we have a uniform interface to test normality:
#
# ```python
# import scipy.stats
# def is_normal(r, level=0.01):
# """
# Applies the Jarque-Bera test to determine if a Series is normal or not
# Test is applied at the 1% level by default
# Returns True if the hypothesis of normality is accepted, False otherwise
# """
# if isinstance(r, pd.DataFrame):
# return r.aggregate(is_normal)
# else:
# statistic, p_value = scipy.stats.jarque_bera(r)
# return p_value > level
# ```
#
import pandas as pd
isinstance(hfi, pd.DataFrame)
erk.is_normal(normal_rets)
# ## Testing CRSP SmallCap and Large Cap returns for Normality
#
# Let's see whether any of the returns we've been studying so far pass the normality hypothesis.
ffme = erk.get_ffme_returns()
erk.skewness(ffme)
erk.kurtosis(ffme)
erk.is_normal(ffme)
|
Investment Management/Course1/lab_105.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# General:
import tweepy # To consume Twitter's API
import pandas as pd # To handle data
import numpy as np # For number computing
# For plotting and visualization:
from IPython.display import display
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
# -
CONSUMER_KEY = 'g98ma1alivKBO1F8mgxqsmhwW'
CONSUMER_SECRET = '<KEY>'
ACCESS_TOKEN = '<KEY>'
ACCESS_SECRET = '<KEY>'
# API's setup:
def twitter_setup():
"""
Utility function to setup the Twitter's API
with our access keys provided.
"""
# Authentication and access using keys:
auth = tweepy.OAuthHandler(CONSUMER_KEY, CONSUMER_SECRET)
auth.set_access_token(ACCESS_TOKEN, ACCESS_SECRET)
# Return API with authentication:
api = tweepy.API(auth)
return api
# +
# We create an extractor object:
extractor = twitter_setup()
# We create a tweet list as follows:
tweets = extractor.user_timeline(screen_name="NBA", count=200)
print("Number of tweets extracted: {}.\n".format(len(tweets)))
# We print the most recent 5 tweets:
print("20 recent tweets:\n")
for tweet in tweets[:20]:
print(tweet.text)
print()
# +
# We create a pandas dataframe as follows:
data = pd.DataFrame(data=[tweet.text for tweet in tweets], columns=['Tweets'])
# We display the first 10 elements of the dataframe:
display(data.head(20))
# -
# Internal methods of a single tweet object:
print(dir(tweets[0]))
# We print info from the first tweet:
print(tweets[0].id)
print(tweets[0].created_at)
print(tweets[0].source)
print(tweets[0].favorite_count)
print(tweets[0].retweet_count)
print(tweets[0].geo)
print(tweets[0].coordinates)
print(tweets[0].entities)
print(tweets[0].user.followers_count)
# +
# We add relevant data:
data['len'] = np.array([len(tweet.text) for tweet in tweets])
data['ID'] = np.array([tweet.id for tweet in tweets])
data['Date'] = np.array([tweet.created_at for tweet in tweets])
data['Source'] = np.array([tweet.source for tweet in tweets])
data['Likes'] = np.array([tweet.favorite_count for tweet in tweets])
data['RTs'] = np.array([tweet.retweet_count for tweet in tweets])
data['text'] = np.array([tweet.text for tweet in tweets])
data['Followers'] = np.array([tweet.user.followers_count for tweet in tweets])
# -
# Display of first 20 elements from dataframe:
display(data.head(20))
|
6210_project_YufanYang_JiahaoZhao_YoumingZheng/code_collecting_data/homepage_tweets.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.7.2
# language: julia
# name: julia-1.7
# ---
# # Strings
#
# Topics:
# 1. How to get a string
# 2. String interpolation
# 3. String concatenation
# ## How to get a string
#
# Enclose your characters in " " or """ """!
s1 = "I am a string."
s2 = """I am also a string. """
# There are a couple functional differences between strings enclosed in single and triple quotes. <br>
# One difference is that, in the latter case, you can use quotation marks within your string.
"Here, we get an "error" because it's ambiguous where this string ends "
"""Look, Mom, no "errors"!!! """
# Note that ' ' define a character, but NOT a string!
typeof('a')
'We will get an error here'
# ## String interpolation
#
# We can use the $ sign to insert existing variables into a string and to evaluate expressions within a string. <br>
# Below is an example that contains some highly sensitive personal information.
name = "Jane"
num_fingers = 10
num_toes = 10
println("Hello, my name is $name.")
println("I have $num_fingers fingers and $num_toes toes.")
println("That is $(num_fingers + num_toes) digits in all!!")
# ## String concatenation
#
# Below are three ways we can concatenate strings! <br><br>
# The first way is to use the `string()` function. <br>
# `string()` converts non-string inputs to strings.
s3 = "How many cats ";
s4 = "is too many cats?";
😺 = 10
string(s3, s4)
string("I don't know, but ", 😺, " is too few.")
# We can also use `*` for concatenation!
s3*s4
# ### Exercises
#
# #### 2.1
# Create a string that says "hi" 1000 times, first with `repeat` and then with the exponentiation operator, which can call `*` under the hood. Assign it the variable `hi` below.
# + deletable=false nbgrader={"checksum": "2046f6f9c3d290e2d63b4c261629d27d", "grade": false, "grade_id": "cell-5b38bdb1115161b4", "locked": false, "schema_version": 1, "solution": true}
# + deletable=false editable=false hide_input=true nbgrader={"checksum": "8b3cc1d8a062b67271109b980793144e", "grade": true, "grade_id": "cell-20b1027a968fbb48", "locked": true, "points": 1, "schema_version": 1, "solution": false}
@assert hi == "hihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihihi"
# -
# #### 2.2
# Declare two variables
#
# ```julia
# a = 3
# b = 4
# ```
# and use them to create two strings:
# ```julia
# "3 + 4"
# "7"
# ```
# and store the results in `c` and `d` respectively
# + deletable=false nbgrader={"checksum": "fad4e601d321aac4b6a1ea9b4e859309", "grade": false, "grade_id": "cell-36beb524c8ecd33b", "locked": false, "schema_version": 1, "solution": true}
# + deletable=false editable=false hide_input=true nbgrader={"checksum": "191b413e8c273e1d2c6423d090ce6f0f", "grade": true, "grade_id": "cell-14c3c5b015a97892", "locked": true, "points": 1, "schema_version": 1, "solution": false}
@assert c == "3 + 4"
@assert d == "7"
# -
|
Julia/Notebooks/1. Introduction-to-Julia-main/1 - Strings.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
list0 = [1,2,"3",4]
if "3" in list0:
print("있음")
else:
print("없음")
listyou = ['you','You','YOu','YoU']
my_school______ = listyou
for stupidperson in my_school______:
print(stupidperson*4)
단 = int(input("몃단?"))
for i in [1,2,3,4,5,6,7,8,9]:
print(i*단)
단= int(input("??"))
for i in range (1,101):
print(i * 단)
answer = range(1,101)[0:11]
print(answer)
numbers = range(1,101)
for bla in numbers:
print(bla)
numbers = range(1,101)
for bla in numbers:
if bla % 2 == 0:
print (bla)
numbers = range(0,101)
for bla in numbers:
if bla % 5 == 0:
print (bla)
# +
for e in range(1,10):
for r in range(1,10):
for t in range(1,10):
if (e * r * t)%2==0:
print (e * r * t,end=" ")
print( )
print(" ")
# -
|
nest_loop.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <script async src="https://www.googletagmanager.com/gtag/js?id=UA-59152712-8"></script>
# <script>
# window.dataLayer = window.dataLayer || [];
# function gtag(){dataLayer.push(arguments);}
# gtag('js', new Date());
#
# gtag('config', 'UA-59152712-8');
# </script>
#
# # Tutorial-IllinoisGRMHD: MoL_registration.C
#
# ## Authors: <NAME> & <NAME>
#
# <font color='red'>**This module is currently under development**</font>
#
# ## In this tutorial module we explain the Method of Lines (MoL) options used by the `IllinoisGRMHD` codes. This module will likely be absorbed by another one once we finish documenting the code.
#
# ### Required and recommended citations:
#
# * **(Required)** <NAME>., <NAME>., <NAME>., <NAME>., and <NAME>. IllinoisGRMHD: an open-source, user-friendly GRMHD code for dynamical spacetimes. Class. Quantum Grav. 32 (2015) 175009. ([arxiv:1501.07276](http://arxiv.org/abs/1501.07276)).
# * **(Required)** <NAME>., <NAME>., <NAME>., <NAME>. Primitive Variable Solvers for Conservative General Relativistic Magnetohydrodynamics. Astrophysical Journal, 641, 626 (2006) ([astro-ph/0512420](https://arxiv.org/abs/astro-ph/0512420)).
# * **(Recommended)** <NAME>., <NAME>., <NAME>. An efficient shock-capturing central-type scheme for multidimensional relativistic flows - II. Magnetohydrodynamics. A&A 400 (2) 397-413 (2003). DOI: 10.1051/0004-6361:20021641 ([astro-ph/0210618](https://arxiv.org/abs/astro-ph/0210618)).
# <a id='toc'></a>
#
# # Table of Contents
# $$\label{toc}$$
#
# This module is organized as follows
#
# 0. [Step 0](#src_dir): **Source directory creation**
# 1. [Step 1](#introduction): **Introduction**
# 1. [Step 2](#mol_registration__c): **`MoL_registration.C`**
# 1. [Step n-1](#code_validation): **Code validation**
# 1. [Step n](#latex_pdf_output): **Output this notebook to $\LaTeX$-formatted PDF file**
# <a id='src_dir'></a>
#
# # Step 0: Source directory creation \[Back to [top](#toc)\]
# $$\label{src_dir}$$
#
# We will now use the [cmdline_helper.py NRPy+ module](Tutorial-Tutorial-cmdline_helper.ipynb) to create the source directory within the `IllinoisGRMHD` NRPy+ directory, if it does not exist yet.
# +
# Step 0: Creation of the IllinoisGRMHD source directory
# Step 0a: Add NRPy's directory to the path
# https://stackoverflow.com/questions/16780014/import-file-from-parent-directory
import os,sys
nrpy_dir_path = os.path.join("..","..")
if nrpy_dir_path not in sys.path:
sys.path.append(nrpy_dir_path)
# Step 0b: Load up cmdline_helper and create the directory
import cmdline_helper as cmd
IGM_src_dir_path = os.path.join("..","src")
cmd.mkdir(IGM_src_dir_path)
# Step 0c: Create the output file path
outfile_path__MoL_registration__C = os.path.join(IGM_src_dir_path,"MoL_registration.C")
# -
# <a id='introduction'></a>
#
# # Step 1: Introduction \[Back to [top](#toc)\]
# $$\label{introduction}$$
# <a id='mol_registration__c'></a>
#
# # Step 2: `MoL_registration.C` \[Back to [top](#toc)\]
# $$\label{mol_registration__c}$$
# +
# %%writefile $outfile_path__MoL_registration__C
//--------------------------------------------------------------------------
// Register with the time stepper
// (MoL thorn, found in arrangements/CactusBase/MoL)
// To understand this, read documentation in arrangements/CactusBase/MoL/doc
//--------------------------------------------------------------------------
#include "cctk.h"
#include <cstdio>
#include <cmath>
#include <cstddef>
#include "cctk_Parameters.h"
#include "cctk_Arguments.h"
#include "Symmetry.h"
extern "C" void IllinoisGRMHD_RegisterVars(CCTK_ARGUMENTS)
{
DECLARE_CCTK_ARGUMENTS;
DECLARE_CCTK_PARAMETERS;
CCTK_INT ierr = 0, group, rhs;
// Register evolution & RHS gridfunction groups
/* Ax and Ax_rhs */
group = CCTK_GroupIndex("IllinoisGRMHD::em_Ax");
rhs = CCTK_GroupIndex("IllinoisGRMHD::em_Ax_rhs");
ierr += MoLRegisterEvolvedGroup(group, rhs);
/* Ay and Ay_rhs */
group = CCTK_GroupIndex("IllinoisGRMHD::em_Ay");
rhs = CCTK_GroupIndex("IllinoisGRMHD::em_Ay_rhs");
ierr += MoLRegisterEvolvedGroup(group, rhs);
/* Az and Az_rhs */
group = CCTK_GroupIndex("IllinoisGRMHD::em_Az");
rhs = CCTK_GroupIndex("IllinoisGRMHD::em_Az_rhs");
ierr += MoLRegisterEvolvedGroup(group, rhs);
/* psi6phi and psi6phi_rhs */
group = CCTK_GroupIndex("IllinoisGRMHD::em_psi6phi");
rhs = CCTK_GroupIndex("IllinoisGRMHD::em_psi6phi_rhs");
ierr += MoLRegisterEvolvedGroup(group, rhs);
/* ALL OTHER EVOLVED VARIABLES (rho_star,tau,mhd_st_x,mhd_st_y,mhd_st_z) */
group = CCTK_GroupIndex("IllinoisGRMHD::grmhd_conservatives");
rhs = CCTK_GroupIndex("IllinoisGRMHD::grmhd_conservatives_rhs");
ierr += MoLRegisterEvolvedGroup(group, rhs);
if (ierr) CCTK_ERROR("Problems registering with MoL");
}
# -
# <a id='code_validation'></a>
#
# # Step n-1: Code validation \[Back to [top](#toc)\]
# $$\label{code_validation}$$
#
# First we download the original `IllinoisGRMHD` source code and then compare it to the source code generated by this tutorial notebook.
# +
# Verify if the code generated by this tutorial module
# matches the original IllinoisGRMHD source code
# First download the original IllinoisGRMHD source code
import urllib
from os import path
original_IGM_file_url = "https://bitbucket.org/zach_etienne/wvuthorns/raw/5611b2f0b17135538c9d9d17c7da062abe0401b6/IllinoisGRMHD/src/MoL_registration.C"
original_IGM_file_name = "MoL_registration-original.C"
original_IGM_file_path = os.path.join(IGM_src_dir_path,original_IGM_file_name)
# Then download the original IllinoisGRMHD source code
# We try it here in a couple of ways in an attempt to keep
# the code more portable
try:
original_IGM_file_code = urllib.request.urlopen(original_IGM_file_url).read().decode("utf-8")
# Write down the file the original IllinoisGRMHD source code
with open(original_IGM_file_path,"w") as file:
file.write(original_IGM_file_code)
except:
try:
original_IGM_file_code = urllib.urlopen(original_IGM_file_url).read().decode("utf-8")
# Write down the file the original IllinoisGRMHD source code
with open(original_IGM_file_path,"w") as file:
file.write(original_IGM_file_code)
except:
# If all else fails, hope wget does the job
# !wget -O $original_IGM_file_path $original_IGM_file_url
# Perform validation
# Validation__MoL_registration__C = !diff $original_IGM_file_path $outfile_path__MoL_registration__C
if Validation__MoL_registration__C == []:
# If the validation passes, we do not need to store the original IGM source code file
# !rm $original_IGM_file_path
print("Validation test for MoL_registration.C: PASSED!")
else:
# If the validation fails, we keep the original IGM source code file
print("Validation test for MoL_registration.C: FAILED!")
# We also print out the difference between the code generated
# in this tutorial module and the original IGM source code
print("Diff:")
for diff_line in Validation__MoL_registration__C:
print(diff_line)
# -
# <a id='latex_pdf_output'></a>
#
# # Step n: Output this notebook to $\LaTeX$-formatted PDF file \[Back to [top](#toc)\]
# $$\label{latex_pdf_output}$$
#
# The following code cell converts this Jupyter notebook into a proper, clickable $\LaTeX$-formatted PDF file. After the cell is successfully run, the generated PDF may be found in the root NRPy+ tutorial directory, with filename
# [Tutorial-IllinoisGRMHD__MoL_registration.pdf](Tutorial-IllinoisGRMHD__MoL_registration.pdf) (Note that clicking on this link may not work; you may need to open the PDF file through another means).
latex_nrpy_style_path = os.path.join(nrpy_dir_path,"latex_nrpy_style.tplx")
# #!jupyter nbconvert --to latex --template $latex_nrpy_style_path --log-level='WARN' Tutorial-IllinoisGRMHD__MoL_registration.ipynb
# #!pdflatex -interaction=batchmode Tutorial-IllinoisGRMHD__MoL_registration.tex
# #!pdflatex -interaction=batchmode Tutorial-IllinoisGRMHD__MoL_registration.tex
# #!pdflatex -interaction=batchmode Tutorial-IllinoisGRMHD__MoL_registration.tex
# !rm -f Tut*.out Tut*.aux Tut*.log
|
IllinoisGRMHD/doc/Tutorial-IllinoisGRMHD__MoL_registration.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # About: LDAP認証の設定
#
# ---
#
# MoodleのLDAP認証プラグインの設定を行います。
# ## 概要
# ### 前提条件
#
# LDAP認証を設定するために必要となる準備事項を以下に示します。
# * LDAPサーバにアクセスするための認証情報など
# * CA証明書(TLS接続を行う場合かつ、追加が必要な場合)
# ## LDAPサーバに接続するための設定
# LDAPサーバにTLS接続を行う場合、以下の設定が必要となることがあります。
#
# * CA証明書の配置
# * LDAPサーバの名前解決に関する設定
#
# 追加の設定が不要な場合はこの章の実行をスキップしてください。
# ### 設定対象となるMoodle環境の指定
#
# 設定対象となるMoodle環境にアクセスするためのAnsibleのグループ名を指定してください。
# +
# (例)
# target_group = 'Moodle'
target_group =
# -
# Ansibleでアクセスできることを確認します。
# !ansible {target_group} -m ping
# ### CA証明書の追加
#
# Moodle環境にCA証明書を追加します。
# 次のセルにCA証明書の内容を指定してください。
# +
# (例)
# ca_certificate = '''-----BEGIN CERTIFICATE-----
# MIIC0zCCAlmgAwIBAgIUCfQ+m0pgZ/BjYAJvxrn/bdGNZokwCgYIKoZIzj0EAwMw
# (中略)
# 8FdfWPypvA==
# -----END CERTIFICATE-----'''
ca_certificate = '''-----BEGIN CERTIFICATE-----
-----END CERTIFICATE-----'''
# -
# MoodleコンテナにCA証明書を配置します。
# +
from tempfile import TemporaryDirectory
from pathlib import Path
with TemporaryDirectory() as workdir:
ca_pem = Path(workdir) / 'ldap-ca.pem'
with ca_pem.open(mode='w') as f:
f.write(ca_certificate)
# !ansible {target_group} -b -m copy -a \
# 'src={ca_pem} dest=/srv/moodle/moodle/conf/ca-trust/'
# -
# CA証明書を追加します。
# !ansible {target_group} -a 'chdir=/srv/moodle \
# docker-compose exec moodle \
# update-ca-trust extract'
# ### LDAPサーバの名前解決に関する設定
#
# MoodleコンテナからLDAPサーバのホスト名の名前解決を行えるように設定を追加します。
# MoodleコンテナからLDAPサーバの名前解決を行うための設定を`docker-compose.yml`に追加するには、以下に示す2つの設定方法があります。
#
# * [DNSサーバの設定を追加する](https://docs.docker.com/compose/compose-file/#dns)
# * [コンテナ内の/etc/hostsにエントリを追加する](https://docs.docker.com/compose/compose-file/#extra_hosts)
# DNSサーバの設定を`docker-compose.yml`に追加する場合の例を以下に示します。この例ではIPアドレス `8.8.8.8`のDNSサーバをMoodleコンテナに追加しています。
#
# ```
# moodle:
# dns:
# - 8.8.8.8 # DNSサーバのIPアドレスを指定する
# ```
# Moodleコンテナの`/etc/hosts` にエントリを追加する場合の例を以下に示します。この例ではホスト名 `ldap.example.org`、IPアドレス`192.168.10.100`のエントリをMoodleコンテナの`/etc/hosts`に追加しています。
#
# ```
# moodle:
# extra_hosts:
# - "ldap.example.org:192.168.10.100"
# ```
# 実際に`docker-compose.yml`を編集するために、Moodle環境からローカル環境に `docker-compose.yml` ファイルの取得を行います。実行結果に表示されているリンクをクリックすることで、ローカル環境に取得した `docker-compose.yml` を編集することができます。
#
# > ファイルの編集後は**必ず**、メニューの[File]-[Save]を選択してファイルの保存を行ってください。
# %run scripts/edit_conf.py
fetch_docker_compose(target_group)
# `docker-compose.yml` を編集した後に、次のセルを実行すると編集の前後での差分を確認することができます。
show_local_docker_compose_diff(target_group)
# DNSサーバの設定を追加した場合の差分の表示例を以下に示します。
#
# ```
# --- docker-compose.yml.orig
# +++ docker-compose.yml
# @@ -25,6 +25,8 @@
# - --explicit_defaults_for_timestamp=true
# - --log-error=/var/log/mysql/error.log
# moodle:
# + dns:
# + - 8.8.8.8
# image: harbor.vcloud.nii.ac.jp/vcp/moodle-simple:3.9
# container_name: moodle
# restart: always
# ```
# `/etc/hosts`のエントリを追加する設定を行った場合の差分の表示例を以下に示します。
#
# ```
# --- docker-compose.yml.orig
# +++ docker-compose.yml
# @@ -25,6 +25,8 @@
# - --explicit_defaults_for_timestamp=true
# - --log-error=/var/log/mysql/error.log
# moodle:
# + extra_hosts:
# + - "ldap.example.org:192.168.10.100"
# image: harbor.vcloud.nii.ac.jp/vcp/moodle-simple:3.9
# container_name: moodle
# restart: always
# ```
# ローカル環境の`docker-compose.yml`をMoodle環境に配置して、さらに変更内容を変更するためにMoodleコンテナの再起動を行います。
apply_docker_compose(target_group)
# ## LDAP認証プラグインの設定
#
# MoodleでLDAP認証行うための設定を行います。
# ### LDAP認証プラグインの有効化
# まずLDAP認証プラグインを有効にします。管理者ユーザでログインして[サイト管理]-[プラグイン]-[認証管理]を選択し認証プラグインの一覧を表示してください。以下のような画面が表示されます。
#
# 
#
# 図の「LDAPサーバ」の行の `Yes` の列にあるアイコンを選択してLDAP認証プラグインを有効化してください。
# ### LDAPサーバの設定
# 「LDAPサーバ」の「設定」のリンクを選択すると以下のような設定画面が表示されます。
#
# 
#
# 「ホストURL」などにLDAPサーバのアドレスなどを設定してください。
#
# また、同じ画面の「バインド設定」でLDAPサーバに接続するための識別名、パスワードも必要に応じて設定してください。
#
# 
#
# 最後に設定画面の最下部にある「変更を保存する」ボタンを選択してください。
# ### LDAPサーバへの接続確認
# 認証プラグインの一覧画面にある「設定テスト」リンクを選択すると、現在の設定でLDAPサーバに接続できるかをテストすることができます。
#
# 
#
# LDAPサーバへの接続が成功すると以下のような表示となります。
#
# 
|
Moodle-Simple/notebooks/211-LDAP認証の設定.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import easyib
api = easyib.REST()
# default parameters: url="https://localhost:5000", ssl=False
# SSL warnings can be suppressed by setting up a SSL certificate, or type "export PYTHONWARNINGS="ignore:Unverified HTTPS request" in a shell
api.get_bars("TSLA")["data"]
list_of_orders = [
{
"conid": api.get_conid("TSLA"),
"orderType": "MKT",
"side": "BUY",
"quantity": 6,
"tif": "GTC",
}
]
api.submit_orders(list_of_orders)
order = {
"conid": api.get_conid("TSLA"),
"orderType": "MKT",
"side": "BUY",
"quantity": 7,
"tif": "GTC",
}
api.modify_order(1258176647,order)
api.get_order(1258176647)
api.get_live_orders()
|
examples/examples.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Python Lambda Functions
# Anonymous, single-use, or throw-away functions.
# **lambda arguments : expression**
# Here are some single-argument lambdas:
add5 = lambda x: x + 5
print(add5(7))
square = lambda x: x * x
print(square(8))
get_tens = lambda p: int(p/10)%10
print(get_tens(749))
print(get_tens(836.21))
# **Lambdas as an argument in other functions**
# One of the most popular uses for lambda functions is as an argument inside sort, or filter functions.
# ### Sorting a List of Tuples using Lambda
list1 = [('eggs', 5.25), ('honey', 9.70), ('carrots', 1.10), ('peaches', 2.45)]
list1.sort(key = lambda x: x[1])
print(list1)
# ### Sorting a List of Dictionaries using Lambda
import pprint as pp
list1 = [{'make':'Ford', 'model':'Focus', 'year':2013}, {'make':'Tesla', 'model':'X', 'year':1999}, {'make':'Mercedes', 'model':'C350E', 'year':2008}]
list2 = sorted(list1, key = lambda x: x['year'])
pp.pprint(list2)
# ### Filtering a List of Integers using Lambda
list1 = [1, 2, 3, 4, 5, 6]
list2 = list(filter(lambda x: x%2 == 0, list1))
print(list2)
odds = lambda x: x%2 == 1
list1 = [1, 2, 3, 4, 5, 6]
list2 = list(filter(odds, list1))
print(list2)
# ### Lambda Function on a List using Map
# Python's map function applies the lambda to every element in the list.
list1 = [1, 2, 3, 4, 5, 6]
list2 = list(map(lambda x: x ** 2, list1))
print(list2)
# ### Lambda Conditionals
# **lambda args: a if boolean_expression else b**
starts_with_J = lambda x: True if x.startswith('J') else False
print(starts_with_J('Joey'))
wordb4 = lambda s, w: s.split()[s.split().index(w)-1] if w in s else None
sentence = 'Four score and seven years ago'
print(wordb4(sentence, 'seven'))
# ### Lambdas on DataTime Objects
# You sometimes want to get just the year, month, date or time for comparision.
# This would typically be most useful as a parameter in sort or filter functions.
# +
import datetime
now = datetime.datetime.now()
print(now)
year = lambda x: x.year
print(year(now))
# +
def do_something(f, val):
return f(val)
func = lambda x: x**3
print(func(16))
print(do_something(func, 5))
# -
# ### Extreme Lambdas
# This is probably a stretch -- you shouldn't be trying to do this much with Lambdas.
# Some things are better done in a regular function. But this shows what's possible with Lambdas.
# +
isnum = lambda q: q.replace('.','',1).isdigit()
print(isnum('25983'))
print(isnum('3.1415'))
print(isnum('T57'))
print(isnum('-16'))
is_num = lambda r: isnum(r[1:]) if r[0]=='-' else isnum(r)
print(is_num('-16.4'))
tonum = lambda s: float(s) if is_num(s) else -1
print(tonum('30y'))
print(tonum('-21.67'), type(tonum('-21.67')))
|
Python Lambda Functions.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from matplotlib import pyplot as plt
import numpy as np
def tax(income):
THRESH1 = 18200
THRESH2 = 37000
THRESH3 = 80000
THRESH4 = 180000
RATE1 = 0.190
RATE2 = 0.325
RATE3 = 0.37
RATE4 = 0.45
tax = 0
if income > THRESH4:
tax += (income - THRESH4) * RATE4
income = THRESH4
if income > THRESH3:
tax += (income - THRESH3) * RATE3
income = THRESH3
if income > THRESH2:
tax += (income - THRESH2) * RATE2
income = THRESH2
if income > THRESH1:
tax += (income - THRESH1) * RATE1
income = THRESH1
return tax
# +
brackets = [0, 18200, 37000, 80000, 180000, 250000]
for i in range(len(brackets) - 1):
income_data = np.arange(brackets[i], brackets[i+1], 10)
tax_data = np.array([tax(j) for j in income_data])
plt.plot(income_data, tax_data)
plt.xlabel('Gross Income (A$)')
plt.ylabel('Tax Due (A$)')
plt.title('Income Tax - Piecewise Defintion')
plt.legend(['< $' + str(threshold) for threshold in brackets[1:]])
# -
print(tax(150000))
# +
def E_of_ring(x, R=1):
return x/(x**2 + R**2)**(3/2)
x_list = np.linspace(0, 10, 100)
E_list = np.array([E_of_ring(x) for x in x_list])
plt.plot(x_list, E_list)
plt.plot(x_list, np.array([E_of_ring(.25*x) for x in x_list]) )
|
jupyter/tax.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="s8XtcgbGj5xN" colab_type="text"
# #Linear Regression for MNIST Images
# + id="j4BNOnZe46CQ" colab_type="code" outputId="af1bf2ab-4a25-474b-8805-f7c1bec8fc44" executionInfo={"status": "ok", "timestamp": 1568778283409, "user_tz": 360, "elapsed": 660, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13266317426275772701"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
import torch
import torchvision
n_epochs = 10
batch_size_train = 128
batch_size_test = 1000
learning_rate = 1e-2
momentum = 0.5
log_interval = 100
random_seed = 1
torch.backends.cudnn.enabled = False
torch.manual_seed(random_seed)
# Checking GPU availability
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
print(device)
# + [markdown] id="FWd5IGhbkCnz" colab_type="text"
# # MNIST dataset is part of torchvision
# ## Divide MNIST into training, validation and test sets
# ## Use DataLoader iterator for loading data in batches
# + id="Y2uiYpfC4_aW" colab_type="code" colab={}
from torch.utils.data import random_split
MNIST_training = torchvision.datasets.MNIST('/MNIST_dataset/', train=True, download=True,
transform=torchvision.transforms.Compose([
torchvision.transforms.ToTensor(),
torchvision.transforms.Normalize((0.1307,), (0.3081,))]))
MNIST_test_set = torchvision.datasets.MNIST('/MNIST_dataset/', train=False, download=True,
transform=torchvision.transforms.Compose([
torchvision.transforms.ToTensor(),
torchvision.transforms.Normalize((0.1307,), (0.3081,))]))
# create a training and a validation set
MNIST_training_set, MNIST_validation_set = random_split(MNIST_training, [55000, 5000])
train_loader = torch.utils.data.DataLoader(MNIST_training_set,batch_size=batch_size_train, shuffle=True)
validation_loader = torch.utils.data.DataLoader(MNIST_validation_set,batch_size=batch_size_train, shuffle=True)
test_loader = torch.utils.data.DataLoader(MNIST_test_set,batch_size=batch_size_test, shuffle=True)
# + [markdown] id="vzUWismvsZeC" colab_type="text"
# ## Check DataLoader
# + id="PSq8BOG85GyN" colab_type="code" outputId="33002280-b4fe-48a0-978c-ca465778a49d" executionInfo={"status": "ok", "timestamp": 1568778284505, "user_tz": 360, "elapsed": 1734, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13266317426275772701"}} colab={"base_uri": "https://localhost:8080/", "height": 50}
examples = enumerate(test_loader)
batch_idx, (example_data, example_targets) = next(examples)
print(example_data.shape)
print(example_targets.shape)
# + [markdown] id="W4Bhye8YspZM" colab_type="text"
# ## Also, make sure to display some images
# + id="g1hgLmOT5KsW" colab_type="code" outputId="0581c6b4-76ed-4433-c7ee-ff4b05b355e6" executionInfo={"status": "ok", "timestamp": 1568778284823, "user_tz": 360, "elapsed": 2044, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13266317426275772701"}} colab={"base_uri": "https://localhost:8080/", "height": 551}
import matplotlib.pyplot as plt
fig = plt.figure()
for i in range(6):
plt.subplot(2,3,i+1)
plt.tight_layout()
plt.imshow(example_data[i][0], cmap='gray', interpolation='none')
plt.title("Ground Truth: {}".format(example_targets[i]))
plt.xticks([])
plt.yticks([])
fig
# + id="aM32G_zp52dY" colab_type="code" colab={}
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
# + id="cd1MU6Yh56HF" colab_type="code" colab={}
# Linear regression
class LinearRegression(nn.Module):
def __init__(self):
super(LinearRegression, self).__init__()
self.fc = nn.Linear(28*28, 1)
def forward(self, x):
x = x.view(x.size(0), -1)
x = self.fc(x)
return x
# + id="_hjjozQl6ARg" colab_type="code" colab={}
linear_model = LinearRegression().to(device)
optimizer = optim.Adam(linear_model.parameters(), lr=learning_rate,amsgrad=True)
classes = torch.tensor([0,1,2,3,4,5,6,7,8,9]).view(1,10).float().to(device)
# + id="Cy8SKSrG6KxV" colab_type="code" colab={}
def train(epoch):
linear_model.train()
for batch_idx, (data, target) in enumerate(train_loader):
data = data.to(device)
target = target.to(device)
optimizer.zero_grad()
output = linear_model(data)
loss = F.mse_loss(output, target.float().view_as(output)) # notice the use of view_as
loss.backward()
optimizer.step()
if batch_idx % log_interval == 0:
print('Train Epoch: {} [{}/{} ({:.0f}%)]\tLoss: {:.6f}'.format(
epoch, batch_idx * len(data), len(train_loader.dataset),
100. * batch_idx / len(train_loader), loss.item()))
# + id="Tr4Bd0BI6P7D" colab_type="code" colab={}
def validation():
linear_model.eval()
validation_loss = 0
correct = 0
with torch.no_grad(): # notice the use of no_grad
for data, target in validation_loader:
data = data.to(device)
target = target.to(device)
output = linear_model(data)
validation_loss += F.mse_loss(output, target.float().view_as(output), size_average=False).item()
_, pred = torch.min(torch.abs(output - classes),dim=1) # notice the type of broadcasting the subtraction uses here
correct += pred.eq(target.data.view_as(pred)).sum()
validation_loss /= len(validation_loader.dataset)
print('\nValidation set: Avg. loss: {:.4f}, Accuracy: {}/{} ({:.0f}%)\n'.format(validation_loss, correct, len(validation_loader.dataset),100. * correct / len(validation_loader.dataset)))
# + id="DGcIxeFW0rR2" colab_type="code" colab={}
def test():
linear_model.eval()
test_loss = 0
correct = 0
with torch.no_grad():
for data, target in test_loader:
data = data.to(device)
target = target.to(device)
output = linear_model(data)
test_loss += F.mse_loss(output, target.float().view_as(output), size_average=False).item()
_, pred = torch.min(torch.abs(output - classes),dim=1) # notice the type of broadcasting the subtraction uses here
correct += pred.eq(target.data.view_as(pred)).sum()
test_loss /= len(test_loader.dataset)
print('\nTest set: Avg. loss: {:.4f}, Accuracy: {}/{} ({:.0f}%)\n'.format(test_loss, correct, len(test_loader.dataset),100. * correct / len(test_loader.dataset)))
# + id="qRG7c7Sy6XOs" colab_type="code" outputId="2d23ee7c-162e-4283-ca77-5ac6a3f75670" executionInfo={"status": "ok", "timestamp": 1568778367947, "user_tz": 360, "elapsed": 85111, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13266317426275772701"}} colab={"base_uri": "https://localhost:8080/", "height": 1000}
validation()
for epoch in range(1, n_epochs + 1):
train(epoch)
validation()
test()
|
Week 3/MNIST_Linear_Regression.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# default_exp ai_platform_constants
# +
#export
from enum import Enum
# https://cloud.google.com/sdk/gcloud/reference/ai-platform/jobs/submit/training#--master-accelerator
class AcceleratorType(Enum):
NVIDIA_TESLA_K80 = 'nvidia-tesla-k80'
NVIDIA_TESLA_P100 = 'nvidia-tesla-p100'
NVIDIA_TESLA_V100 = 'nvidia-tesla-v100'
NVIDIA_TESLA_P4 = 'nvidia-tesla-p4'
NVIDIA_TESLA_T4 = 'nvidia-tesla-t4'
TPU_V2 = 'tpu-v2'
TPU_V2_POD = 'tpu-v2-pod'
TPU_V3 = 'tpu-v3'
TPU_V3_POD = 'tpu-v3-pod'
# +
#export
# https://cloud.google.com/sdk/gcloud/reference/ai-platform/jobs/submit/training#--master-machine-type
class ScaleTier(Enum):
"""Single worker instance.
This tier is suitable for learning how to use AI Platform, and for experimenting
with new models using small datasets."""
BASIC = 'basic'
"""Single worker instance with a GPU."""
BASIC_GPU = 'basic-gpu'
"""Single worker instance with a Cloud TPU."""
BASIC_TPU = 'basic-tpu'
"""The CUSTOM tier is not a set tier, but rather enables you to use your
own cluster specification. When you use this tier, set values to
configure your processing cluster according to these guidelines"""
CUSTOM = 'custom'
"""Many workers and a few parameter servers."""
STANDARD_1 = 'standard-1';
"""A large number of workers with many parameter servers."""
PREMIUM_1 = 'premium-1'
# +
#export
# https://cloud.google.com/compute/docs/machine-types
class MachineType(Enum):
N1_STANDARD_4 = 'n1-standard-4'
N1_STANDARD_8 = 'n1-standard-8'
N1_STANDARD_16 = 'n1-standard-16'
N1_STANDARD_32 = 'n1-standard-32'
N1_STANDARD_64 = 'n1-standard-64'
N1_STANDARD_96 = 'n1-standard-96'
N1_HIGHMEM_2 = 'n1-highmem-2'
N1_HIGHMEM_4 = 'n1-highmem-4'
N1_HIGHMEM_8 = 'n1-highmem-8'
N1_HIGHMEM_16 = 'n1-highmem-16'
N1_HIGHMEM_32 = 'n1-highmem-32'
N1_HIGHMEM_64 = 'n1-highmem-64'
N1_HIGHMEM_96 = 'n1-highmem-96'
N1_HIGHCPU_16 = 'n1-highcpu-16'
N1_HIGHCPU_32 = 'n1-highcpu-32'
N1_HIGHCPU_64 = 'n1-highcpu-64'
N1_HIGHCPU_96 = 'n1-highcpu-96'
# -
#export
class DistributionStrategyType(Enum):
def __str__(self):
return str(self.value)
MIRRORED_STRATEGY = "tf.distribute.MirroredStrategy"
ONE_DEVICE_STRATEGY = "tf.distribute.OneDeviceStrategy"
CENTRAL_STORAGE_STRATEGY = "tf.distribute.experimental.CentralStorageStrategy"
PARAMETER_SERVERSTRATEGY = "tf.distribute.experimental.ParameterServerStrategy"
MULTI_WORKER_MIRRORED_STRATEGY = "tf.distribute.experimental.MultiWorkerMirroredStrategy"
TPU_STRATEGY = "tf.distribute.experimental.TPUStrategy"
|
ai_platform_constants.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# <h1><center> Exogeneous closure for the PKF applied to the Burgers dynamics </center></h1>
# <center>
# <NAME> <br> 2020
# </center>
# + [markdown] slideshow={"slide_type": "notes"}
# The aim is to train a closure to predict the uncertainty dynamics for the Burgers dynamics.
#
# The notebook present the situation where uknown physical processes are represented by an exogeneous NN to train.
# + [markdown] slideshow={"slide_type": "fragment"}
# ---
# <center> <b>Table of contents</b> </center>
#
# 1. [Introduction](#introduction)
# 1. [The Burgers dynamics](#the-burgers-dynamics)
# 1. [PKF for the Burgers dynamics](#pkf-forthe-burgers-dynamics)
# 1. [Numerical application](#numerical-application)
# - [Generation of a database](#generation-of-a-database)
# - [Training of the closure](#training-of-the-closure)
# - [Comparison with the theoretically designed closure](#comparison-with-the-theoretically-designed-closure)
# 1. [Conclusion](#conclusion)
# ---
# + slideshow={"slide_type": "skip"}
import numpy as np
import matplotlib.pyplot as plt
# #%matplotlib inline
# #%matplotlib notebook
# -
# ## Introduction
# The aim is to design a NN which merges known and unknown physics.
import sympy
from sympy import (Function, symbols, init_printing, Derivative,
latex, Add, Mul, Pow,
Integer, Rational, Float, Symbol, symbol,
srepr, Tuple
)
init_printing()
# ## The Burgers dynamics
from pdenetgen import NNModelBuilder, Eq
import tensorflow.keras as keras
def display_system(system):
print(50*'*')
for equation in system:
display(equation)
print(50*'*')
# #### Set of the function and symbols <a id='burgers-pkf-sympy-definition'>
# +
t, x = symbols('t x')
u = Function('u')(t,x)
closure = sympy.Function('closure')(t,x)
V = Function('{V_{u}}')(t,x)
nu = Function('{\\nu_{u,xx}}')(t,x)
Kappa = symbols('\\kappa')
# -
# #### Set constants for numerical experiments <a id='burgers-pkf-num-definition'>
# Constant setting following Pannekoucke et al. (2018)
n = 241
kappa = 0.0025
dt = 0.002
# #### Set of the Burgers equation <a id='burgers-pkf-dyn-burgers'>
burgers_dynamics = [
Eq(
Derivative(u,t),
Kappa*Derivative(u,x,2)-u*Derivative(u,x)
),
]
display_system(burgers_dynamics)
burgers_NN_builder = NNModelBuilder(burgers_dynamics, "Burgers")
print(burgers_NN_builder.code)
exec(burgers_NN_builder.code)
burgers = Burgers(shape=(n,), kappa=kappa)
# ##### Example of forecast from a given initial condition
# +
import matplotlib.pyplot as plt
def plot_results(data, label=None, labelx=True, title=None, save_file=None, normalisation=None,
selected_times=None,style=None, name=None, alpha=1., bolds=[0., 1.]):
normalisation = 1. if normalisation is None else normalisation
selected_times = [time for time in data] if selected_times is None else selected_times
style = 'k' if style is None else style
for time in selected_times:
lalpha = alpha if time in bolds else 0.2
lname = name if time==selected_times[-1] else None
plt.plot(domain.x[0],data[time]/normalisation, style, alpha = lalpha, label=lname)
if labelx:
plt.xlabel('$x/D$', fontsize=15)
if label:
plt.ylabel(label, fontsize=15)
if title:
plt.title(title)
if save_file:
plt.savefig(save_file)
# -
domain = burgers
# Set initial condition for 'u'
U0=0.25*( 1+np.cos(2*np.pi/ domain.lengths[0] *(domain.x[0]-0.25)) )
Umax = U0.max()
burgers.set_dt(dt)
end_time_forecast = 1.
times = burgers.window(end_time_forecast)
saved_times = times[::50]
print('saved_times :' ,saved_times)
forecast = burgers.forecast(times, np.array([U0.reshape((1,)+U0.shape+(1,)) ]))
for time in times:
plt.plot(domain.x[0], forecast[time][0,0,:,0])
# ## PKF for the Burgers dynamics
# #### Set of the PKF equations for the Burgers equation <a id='burgers-pkf-dyn-pkf'>
# +
# From Pannekoucke et al. (2018)
pkf_dynamics = [
# Trend of the expectation of 'u'
Eq(
Derivative(u,t),
Kappa*Derivative(u,x,2)-u*Derivative(u,x)-Derivative(V,x)/Integer(2)
),
# Trend of the variance
Eq(
Derivative(V,t),
-Kappa*V/nu + Kappa*Derivative(V,x,2)-Kappa*Derivative(V,x)**Integer(2)/(Integer(2)*V)
-u*Derivative(V,x)-Integer(2)*V*Derivative(u,x)
),
# Trend of the diffusion
Eq(
Derivative(nu,t),
Integer(4)*Kappa*nu**Integer(2)*closure
-Integer(3)*Kappa*Derivative(nu,x,2)
-Kappa
+Integer(6)*Kappa*Derivative(nu,x)**Integer(2)/nu
-Integer(2)*Kappa*nu*Derivative(V,x,2)/V
+Kappa*Derivative(V,x)*Derivative(nu,x)/V
+Integer(2)*Kappa*nu*Derivative(V,x)**Integer(2)/V**Integer(2)
-u*Derivative(nu,x)
+Integer(2)*nu*Derivative(u,x)
)
]
display_system(pkf_dynamics)
# -
pkf_NN_builder = NNModelBuilder(pkf_dynamics,'NN_Unclosed_PKF_Burgers')
print(pkf_NN_builder.code)
exec(pkf_NN_builder.code)
# #### Construction of a closure as a NN from parameterized form
# The aim is to compute the closure proposal
# $$a\frac{\frac{\partial^{2}}{\partial x^{2}} \operatorname{{\nu_{u,xx}}}{\left(t,x \right)}}{\operatorname{{\nu_{u,xx}}}^{2}{\left(t,x \right)}} +b \frac{1}{ \operatorname{{\nu_{u,xx}}}^{2}{\left(t,x \right)}} +c\frac{ \left(\frac{\partial}{\partial x} \operatorname{{\nu_{u,xx}}}{\left(t,x \right)}\right)^{2}}{\operatorname{{\nu_{u,xx}}}^{3}{\left(t,x \right)}},$$
# as an exogeneous neural network, where $(a,b,c)$ are trainable.
class ClosedPKFBurgers(NN_Unclosed_PKF_Burgers):
def _make_exogenous_model(self):
u = keras.layers.Input(shape=(self.input_shape_x,1))
V = keras.layers.Input(shape=(self.input_shape_x,1))
nu_u_xx = keras.layers.Input(shape=(self.input_shape_x,1))
#
# Computation of the spatial derivatives
#
kernel_Dnu_u_xx_x_o2 = np.asarray([self.dx[self.coordinates.index('x')]**(-2),
-2/self.dx[self.coordinates.index('x')]**2,
self.dx[self.coordinates.index('x')]**(-2)]).reshape((3,)+(1,1))
Dnu_u_xx_x_o2 = DerivativeFactory((3,),kernel=kernel_Dnu_u_xx_x_o2,name='Dnu_u_xx_x_o2')(nu_u_xx)
kernel_Dnu_u_xx_x_o1 = np.asarray([-1/(2*self.dx[self.coordinates.index('x')]),0.0,
1/(2*self.dx[self.coordinates.index('x')])]).reshape((3,)+(1,1))
Dnu_u_xx_x_o1 = DerivativeFactory((3,),kernel=kernel_Dnu_u_xx_x_o1,name='Dnu_u_xx_x_o1')(nu_u_xx)
#
# Design of the unknown closure to train
#
# Terme 1
div_14 = keras.layers.Lambda(lambda x: 1/x,name='DivLayer_14')(nu_u_xx)
pow_12 = keras.layers.multiply([div_14,div_14,] ,name='PowLayer_12')
term1 = keras.layers.multiply([pow_12,Dnu_u_xx_x_o2],name='MulLayer_25')
# Terme 2
div_13 = keras.layers.Lambda(lambda x: 1/x,name='DivLayer_13')(nu_u_xx)
term2 = keras.layers.multiply([div_13,div_13,] ,name='PowLayer_11')
# Terme 3
pow_13 = keras.layers.multiply([Dnu_u_xx_x_o1,Dnu_u_xx_x_o1,] ,name='PowLayer_13')
div_15 = keras.layers.Lambda(lambda x: 1/x,name='DivLayer_15')(nu_u_xx)
pow_14 = keras.layers.multiply([div_15,div_15,div_15,] ,name='PowLayer_14')
term3 = keras.layers.multiply([pow_13,pow_14],name='MulLayer_26')
# Product by (a,b,c), implemented as Conv1D
term1 = keras.layers.Conv1D(1,1,name='times_a',padding='same',use_bias=False,activation='linear')(term1)
term2 = keras.layers.Conv1D(1,1,name='times_b',padding='same',use_bias=False,activation='linear')(term2)
term3 = keras.layers.Conv1D(1,1,name='times_c',padding='same',use_bias=False,activation='linear')(term3)
closure = keras.layers.add([term1, term2, term3],name='Closure')
self._exogenous_model = keras.models.Model(inputs=[u,V,nu_u_xx], outputs=[closure])
def compute_exogenous(self, t, state):
if self._exogenous_model is None:
self._make_exogenous_model()
u,V,nu = state
closure = self._exogenous_model.predict([u,V,nu])
if not isinstance(closure, list):
closure = [closure]
return closure
def _make_full_trend(self):
if self._trend_model is None:
self._make_trend_model()
if self._exogenous_model is None:
self._make_exogenous_model()
state = keras.layers.Input(shape=(3,self.input_shape_x,1))
u = keras.layers.Lambda(lambda x : x[:,0,:,:])(state)
V = keras.layers.Lambda(lambda x : x[:,1,:,:])(state)
nu_u_xx = keras.layers.Lambda(lambda x : x[:,2,:,:])(state)
closure = self._exogenous_model([u,V,nu_u_xx])
trend_u, trend_V, trend_nu = self._trend_model([u,V,nu_u_xx,closure])
trend_u = keras.layers.Reshape((1,self.input_shape_x,1))(trend_u)
trend_V = keras.layers.Reshape((1,self.input_shape_x,1))(trend_V)
trend_nu = keras.layers.Reshape((1,self.input_shape_x,1))(trend_nu)
trend = keras.layers.Concatenate(axis=1)([trend_u,trend_V,trend_nu])
self._full_trend = keras.models.Model(inputs=state,outputs=trend)
closed_burgers = ClosedPKFBurgers(shape=(241,),kappa=kappa)
closed_burgers._make_full_trend()
# **Set initial PKF fields**
# +
# Set initial condition for the variance parameter 'V_u'
V0 = (0.01*Umax)**2 + 0*U0
# Set the initial condition for the diffusion
# L**2 = 2nu t => nu = 0.5*L**2
lh = 0.02*domain.lengths[0]
nu0 = 0.5*lh**2 + 0*U0
state0 = np.asarray([U0, V0,nu0])
normalization = {
'Velocity':U0.max(),
'Variance':V0.max(),
'Length-scale':lh
}
# -
length_scale = lambda nu: np.sqrt(2*nu)
plt.figure(figsize=(12,12))
for k,field in enumerate(normalization):
plt.subplot(221+k)
if field=='Length-scale':
data = {0:length_scale(state0[k])}
else:
data = {0:state0[k]}
plot_results(data, label=field)
# ## Numerical application
def plot_pkf_traj_ensemble(traj):
plt.figure(figsize=(15,5))
for k,field in enumerate(normalization):
if field=='Length-scale':
data = {time:length_scale(traj[time][k]) for time in traj}
else:
data = {time:traj[time][k] for time in traj}
plt.subplot(131+k)
plot_results(data,label=field,normalisation=normalization[field])
def plot_pkf_traj_NN(traj):
plt.figure(figsize=(15,5))
for k,field in enumerate(normalization):
if field=='Length-scale':
data = {time:length_scale(traj[time][k][0,:,0]) for time in traj}
else:
data = {time:traj[time][k][0,:,0] for time in traj}
plt.subplot(131+k)
plot_results(data,label=field,normalisation=normalization[field])
state0 = np.asarray([U0.reshape((1,)+U0.shape+(1,)),
V0.reshape((1,)+V0.shape+(1,)),
nu0.reshape((1,)+nu0.shape+(1,))])
# ### Generation of a database
# **Gaussian random vector of Gaussian correlation function**
# +
# Création d'une matrice de covariance d'erreur de prévision initiale: $P_0$
# Cette matrice est construite comme une matrice homogène de corrélation Gaussienne et de longueur de portée l_h
# 1) Définition de la fonction de corrélation homogène
gauss = lambda x : np.exp(-0.5*x**2/lh**2) # lh has been previously specified
correlation = gauss(domain.x[0]-domain.x[0][domain.shape[0]//2])
spectrum = np.abs(np.fft.fft(correlation))
# 2) Construction de B^(1/2)
std_spectrum = np.sqrt(spectrum)
def make_sample():
zeta = np.random.normal(size=domain.shape)
zeta = np.fft.fft(zeta)
ef = np.fft.ifft(std_spectrum * zeta)
ef = np.real(ef)
return ef
# -
plt.figure(figsize=(12,5))
plt.subplot(121)
plt.plot(domain.x[0], correlation)
plt.title('Homogenous correlation function');
plt.subplot(122)
for k in range(10):
plt.plot(domain.x[0], make_sample())
plt.title("Example of sample errors");
# ##### **Diagnosis tool for ensemble estimation of expectation/variance/diffusion tensor**
# +
def make_init_ensemble(Ne):
return np.array([make_sample() for k in range(Ne)])
def estimate_covariance(ensemble):
mean = ensemble.mean(0)
error = (ensemble - mean)/np.sqrt(len(ensemble))
return error.T @ error
class EnsembleDiagnosis(object):
def __init__(self, ensemble, base_space):
self.base_space = base_space
if isinstance(ensemble, list):
ensemble = np.array(ensemble)
if len(ensemble.shape)==3:
ensemble = np.array([elm[0] for elm in ensemble])
# 1) Computation of the mean
self.mean = ensemble.mean(axis=0)
# 2) Computation of the variance
self.std = ensemble.std(axis=0)
self.variance = self.std*self.std
# 3) Computation of the metric terms
# we use the formula g_ij = E[(D_i eps)(D_j eps)]
# a) Computation of the normalized error
epsilon = (ensemble-self.mean)/self.std
# b) Computation of derivatives
n = self.base_space.shape[0]
K = np.arange(n)
kp = (K+1)%n
km = (K-1)%n
dx = self.base_space.dx[0]
Depsilon = np.array([(eps[kp]-eps[km])/(2*dx) for eps in epsilon])
self.metric = (Depsilon*Depsilon).mean(axis=0) # see Pannekoucke et al. (2018) for details
# Computation of the diffusion tensor
self.diffusion = 0.5*1/self.metric
self.length_scale = np.sqrt(2*self.diffusion)
# -
# ##### **Ensemble validation for the covariance setting**
# +
Ne = 1600
ensemble = make_init_ensemble(Ne)
mean = ensemble.mean(axis=0)
std = ensemble.std(axis=0)
print(f"Validation of the mean (=0): {mean.mean()} +/- {mean.std()}" )
print(f"Validation of the standard-deviation (=1): {std.mean()} +/- {std.std()}" )
ens_diagnosis = EnsembleDiagnosis(ensemble, domain)
nu_h = 0.5*lh**2
plt.figure(figsize=(15,5))
plt.subplot(131)
plt.plot(ens_diagnosis.mean)
plt.title('Moyenne')
plt.subplot(132)
plt.plot(ens_diagnosis.variance)
plt.title('Variance')
plt.subplot(133)
plt.plot(ens_diagnosis.diffusion/nu_h)
plt.title('diffusion (normalisée par $nu_h$)')
# -
# **Computation of a large ensemble (1600 members) to build a reference**
# Standard deviation for the initial perturbation
sigma_f = 0.01*U0.max()
# +
# Set parameters for ensemble estimation
large_Ne = 1600
# 1. Set the initial background state
random_U0 = U0 + sigma_f*make_init_ensemble(1)[0]
# 2. Build an ensemble of initial perturbed state
ensemble = make_init_ensemble(large_Ne)
ensemble_ea = np.array([random_U0+sigma_f*ea for ea in ensemble])
ensemble_ea = ensemble_ea.reshape((1,)+ensemble_ea.shape+(1,))
print(f"shape of ensemble_ea: {ensemble_ea.shape}")
# 3. Build the ensemble of forecast using the NN architecture
ensemble_forecast = burgers.forecast(times,ensemble_ea)
# -
# 4. Compute diagnosis from ensemble
ensemble_traj = {}
for time in times[::50]:
diagnosis = EnsembleDiagnosis(ensemble_forecast[time][0,:,:,0], domain)
ensemble_traj[time] = [diagnosis.mean, diagnosis.variance, diagnosis.diffusion]
plot_pkf_traj_ensemble(ensemble_traj)
# #### **Generation of the training data set**
def generate_data(k, Ne=400):
# 1. Set the initial background state
random_U0 = U0 + sigma_f*make_init_ensemble(1)[0]
# 2. Build an ensemble of initial perturbed state
ensemble = make_init_ensemble(Ne)
ensemble_ea = np.array([random_U0+sigma_f*ea for ea in ensemble])
ensemble_ea = ensemble_ea.reshape((1,)+ensemble_ea.shape+(1,))
# 3. Compute the ensemble of forecasts
ensemble_forecast = burgers.forecast(times,ensemble_ea)
# 4. Compute the diagnosis
diagnosis_list = []
for time in times:
diagnosis = EnsembleDiagnosis(ensemble_forecast[time][0,:,:,0], domain)
diagnosis_list.append( np.array([diagnosis.mean, diagnosis.variance, diagnosis.diffusion]))
return diagnosis_list
# +
data_size = 400 # for Ne=400, this takes 1h09'01'' so take care with this..
save_file = "pkf-dataset.npy"
generate_data_set = False
parallel_diagnosis = False
try:
# load data
data = np.load(save_file)
data = data.reshape(data.shape+(1,))
except:
# 1. Generate data
#data = [generate_data(k) for k in range(data_size)]
data = []
for k in range(data_size):
if k%5==0:
print(k)
data.append(generate_data(k))
# 2. Save data
data = np.array(data)
np.save(save_file,data)
# -
data.shape
# ### Training of the closure
# **Make a RK4 time scheme**
# Schéma temporelle de type RK4
def make_time_scheme(dt, trend):
""" Implémentation d'un schéma de RK4 sous forme de réseau de neurones """
state = keras.layers.Input(shape = trend.input_shape[1:])
# k1
k1 = trend(state)
# k2
_tmp_1 = keras.layers.Lambda(lambda x : 0.5*dt*x)(k1)
input_k2 = keras.layers.add([state,_tmp_1])
k2 = trend(input_k2)
# k3
_tmp_2 = keras.layers.Lambda(lambda x : 0.5*dt*x)(k2)
input_k3 = keras.layers.add([state,_tmp_2])
k3 = trend(input_k3)
# k4
_tmp_3 = keras.layers.Lambda(lambda x : dt*x)(k3)
input_k4 = keras.layers.add([state,_tmp_3])
k4 = trend(input_k4)
# output
# k2+k3
add_k2_k3 = keras.layers.add([k2,k3])
add_k2_k3_mul2 = keras.layers.Lambda(lambda x:2.*x)(add_k2_k3)
# Add k1,k4
_sum = keras.layers.add([k1,add_k2_k3_mul2,k4])
# *dt
_sc_mul = keras.layers.Lambda(lambda x:dt/6.*x)(_sum)
output = keras.layers.add([state, _sc_mul])
time_scheme = keras.models.Model(inputs =[state], outputs=[output])
return time_scheme
# +
closed_pkf_burgers = ClosedPKFBurgers(shape=(241,),kappa=kappa)
closed_pkf_burgers._make_full_trend()
trained = closed_pkf_burgers._full_trend.get_weights()
trained = np.array((trained[0], trained[1], trained[2])).flatten()
trained
# -
closed_pkf_burgers._make_full_trend()
closed_pkf_burgers._full_trend.summary()
closed_pkf_burgers._exogenous_model.summary()
time_scheme = make_time_scheme(dt, closed_pkf_burgers._full_trend)
#time_scheme.summary()
# **Extract data for the training**
data[0].shape
select_from = 400 # 200
X = np.array([elm[select_from:-1] for elm in data])
Y = np.array([elm[select_from+1:] for elm in data])
X = X.reshape((np.prod(X.shape[:2]),3,241,1))
Y = Y.reshape((np.prod(Y.shape[:2]),3,241,1))
X.shape
# **Training of the NN**
trained = closed_pkf_burgers._full_trend.get_weights()
trained = np.array((trained[0], trained[1], trained[2])).flatten()
trained
# +
# Expérience d'apprentissage:
# 2. Adam
lr = 0.1
epochs = 30
for iteration in range(3):
# 1. Set the learning
time_scheme.compile(optimizer=keras.optimizers.Adam(lr=lr),
loss='mean_squared_error')
# 2. Train
history = time_scheme.fit(X,Y,epochs=epochs, batch_size=32,verbose=0)
print(f"iteration {iteration} is complet")
# 3. Plot history
plt.figure()
plt.plot(history.history['loss'])
# 4. Update the learning rate for next iteration
lr = lr/10
# -
# ### Comparison with the theoretically designed closure
# **Compare the weights with the previous theoretical closure**
# The weights of the theoretical closure are : 1, 3/4, -2
trained = closed_pkf_burgers._full_trend.get_weights()
trained = np.array((trained[0], trained[1], trained[2])).flatten()
trained
theoretical = np.array([1,3/4,-2])
relative_error = (trained - theoretical)/theoretical
relative_error*100
# **Exemple de prévision réalisée avec le modèle calibré**
# default
closed_pkf_burgers.set_dt(dt)
times = closed_pkf_burgers.window(1)
saved_times = times[::50]
print('saved_times :' ,saved_times)
state0.shape
trained_unclosed_traj = closed_pkf_burgers.forecast(times, state0, saved_times)
normalization
# PKF using trained closure
plot_pkf_traj_NN(trained_unclosed_traj)
plt.savefig('./figures/burgers-exogeneous-b.pdf')
# ensemble of forecast statistics
plot_pkf_traj_ensemble(ensemble_traj)
plt.savefig('./figures/burgers-exogeneous-a.pdf')
# ## Conclusion <a id='conclusion'/>
# In this notebook, we have considered the uncertainty prediction for the Burgers dynamics where an unclosed term is present.
#
# A closure has been proposed and implemented as an exogeneous neural network with three unknowns.
#
# A synthetic dataset has been used to train the NN. The resulting closure has shown to be relevant to predict the uncertainty.
|
example/pdenetgen-NN-PKF_burgers_learn-exogenous-closure.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="Y_WGyPRajc2P" colab_type="text"
# # MNIST Dynamic Filter Classification - Result
# + [markdown] id="sSeMh3LCjpwa" colab_type="text"
# Note: This notebook is desinged to run with Python3 and GPU runtime.
#
# 
# + [markdown] id="-xkr13nCB0Il" colab_type="text"
# This notebook uses TensorFlow2.x.
# + id="kop8_9RihcJX" colab_type="code" outputId="332ff586-5747-47aa-e0c3-b3cc5186bb21" colab={"base_uri": "https://localhost:8080/", "height": 34}
# %tensorflow_version 2.x
# + [markdown] id="VJO3PPzqsq8d" colab_type="text"
# ####[MDR-01]
# Import modules.
# + id="gB5UUoAXIVmC" colab_type="code" colab={}
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
from tensorflow.keras import layers, models
from tensorflow.keras.datasets import mnist
# + [markdown] id="yz2h7_8St1wi" colab_type="text"
# ####[MDR-02]
# Download the MNIST dataset and store into NumPy arrays.
# + id="ASgzWK5AjWvn" colab_type="code" colab={}
(train_images, train_labels), (test_images, test_labels) = mnist.load_data()
train_images = train_images.reshape(
(len(train_images), 784)).astype('float32') / 255
test_images = test_images.reshape(
(len(test_images), 784)).astype('float32') / 255
train_labels = tf.keras.utils.to_categorical(train_labels, 10)
test_labels = tf.keras.utils.to_categorical(test_labels, 10)
# + [markdown] id="qdQ0Tp2IvFy8" colab_type="text"
# ####[MDR-03]
# Mount your Google Drive on `/content/gdrive`.
# + id="tpL_niBTXggS" colab_type="code" outputId="350e0c55-6796-4d65-d027-208014ae201b" colab={"base_uri": "https://localhost:8080/", "height": 34}
from google.colab import drive
drive.mount('/content/gdrive')
# + [markdown] id="UmuSINelkooA" colab_type="text"
# ####[MDR-04]
# Confirm that the exported model file `MNIST_single.hd5` is on your Google Drive.
# + id="zq-uJjvNgO6A" colab_type="code" outputId="8ea0089b-d6cc-48a8-b033-52dbd549aaa3" colab={"base_uri": "https://localhost:8080/", "height": 34}
# !ls -l '/content/gdrive/My Drive/MNIST_single.hd5'
# + [markdown] id="fBltXsSRvZn0" colab_type="text"
# ####[MDR-05]
# Restore the model from the exported model file.
# + id="BakcuKxdQoSL" colab_type="code" outputId="4b6101a2-2990-4b07-82ec-7a4a127d1955" colab={"base_uri": "https://localhost:8080/", "height": 445}
model = models.load_model('/content/gdrive/My Drive/MNIST_single.hd5')
model.summary()
# + [markdown] id="YCp1qNMQluZj" colab_type="text"
# ####[MDR-06]
# Define a model to extract outputs from intermediate layers.
# + id="gCrvgrephgYN" colab_type="code" colab={}
layer_outputs = [model.get_layer('conv_filter').output,
model.get_layer('max_pooling').output]
model2 = models.Model(inputs=model.input, outputs=layer_outputs)
# + [markdown] id="RNkIEPULmGJl" colab_type="text"
# ####[MDR-07]
# Apply the trained filters to the MNIST dataset.
# + id="oD-7fgQjibJb" colab_type="code" colab={}
conv_output, pool_output = model2.predict(test_images[:9])
filter_vals = model.get_layer('conv_filter').get_weights()[0]
# + [markdown] id="whBG8JpVmozt" colab_type="text"
# ####[MDR-08]
# Show images after applying the convolutional filters.
# + id="Ix937xqIiJCi" colab_type="code" outputId="fe7e0d69-1e2e-4f1d-c450-e36b02c0c597" colab={"base_uri": "https://localhost:8080/", "height": 975}
num_filters = 16
fig = plt.figure(figsize=(10, num_filters+1))
v_max = np.max(conv_output)
for i in range(num_filters):
subplot = fig.add_subplot(num_filters+1, 10, 10*(i+1)+1)
subplot.set_xticks([])
subplot.set_yticks([])
subplot.imshow(filter_vals[:,:,0,i], cmap=plt.cm.gray_r)
for i in range(9):
subplot = fig.add_subplot(num_filters+1, 10, i+2)
subplot.set_xticks([])
subplot.set_yticks([])
subplot.set_title('%d' % np.argmax(test_labels[i]))
subplot.imshow(test_images[i].reshape((28,28)),
vmin=0, vmax=1, cmap=plt.cm.gray_r)
for f in range(num_filters):
subplot = fig.add_subplot(num_filters+1, 10, 10*(f+1)+i+2)
subplot.set_xticks([])
subplot.set_yticks([])
subplot.imshow(conv_output[i,:,:,f],
vmin=0, vmax=v_max, cmap=plt.cm.gray_r)
# + [markdown] id="ay0hH6cmmu8O" colab_type="text"
# ####[MDR-09]
# Show images after applying the pooling layer.
# + id="6zxP1eJtjINb" colab_type="code" outputId="3677525c-cda4-4eb7-d4ef-ee05ded4ffb4" colab={"base_uri": "https://localhost:8080/", "height": 975}
num_filters = 16
fig = plt.figure(figsize=(10, num_filters+1))
v_max = np.max(pool_output)
for i in range(num_filters):
subplot = fig.add_subplot(num_filters+1, 10, 10*(i+1)+1)
subplot.set_xticks([])
subplot.set_yticks([])
subplot.imshow(filter_vals[:,:,0,i],
cmap=plt.cm.gray_r, interpolation='nearest')
for i in range(9):
subplot = fig.add_subplot(num_filters+1, 10, i+2)
subplot.set_xticks([])
subplot.set_yticks([])
subplot.set_title('%d' % np.argmax(test_labels[i]))
subplot.imshow(test_images[i].reshape((28,28)),
vmin=0, vmax=1, cmap=plt.cm.gray_r)
for f in range(num_filters):
subplot = fig.add_subplot(num_filters+1, 10, 10*(f+1)+i+2)
subplot.set_xticks([])
subplot.set_yticks([])
subplot.imshow(pool_output[i,:,:,f],
vmin=0, vmax=v_max, cmap=plt.cm.gray_r)
# + [markdown] id="5FR5LlCSm4f1" colab_type="text"
# ####[MDR-10]
# Show prodiction results (probability for each digit) for incorrect predictions.
# + id="inzg7CcljP1z" colab_type="code" outputId="e70adde0-3af0-4656-ea51-6f0535116ab2" colab={"base_uri": "https://localhost:8080/", "height": 608}
fig = plt.figure(figsize=(12, 10))
c = 0
for (image, label) in zip(test_images, test_labels):
image
p_val = model.predict(np.array([image]))
pred = p_val[0]
prediction, actual = np.argmax(pred), np.argmax(label)
if prediction == actual:
continue
subplot = fig.add_subplot(5, 4, c*2+1)
subplot.set_xticks([])
subplot.set_yticks([])
subplot.set_title('%d / %d' % (prediction, actual))
subplot.imshow(image.reshape((28, 28)),
vmin=0, vmax=1, cmap=plt.cm.gray_r)
subplot = fig.add_subplot(5, 4, c*2+2)
subplot.set_xticks(range(10))
subplot.set_xlim(-0.5, 9.5)
subplot.set_ylim(0,1)
subplot.bar(range(10), pred, align='center', edgecolor='b')
c += 1
if c == 10:
break
|
Chapter04/5. MNIST dynamic filter classification result.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # T4 Time series
# ## Time series with elapsed time relative to start time
# Experimental data has been recorded and saved as numpy array in a file. We will load the file and extract information about the data by displaying it and calculating various statistics.
# In order to complete the below excercies, you need to download the files `time-series1N.npy` and `time-series2.xlsx` from the github repository. Both are located in the `tutorials` folder. Note that both files need to be saved in the same folder as this notebook.
# #### Load data
# First, we will load the data using the numpy `load` function which can be used to open files with the `.npy` ending.
# +
# first we load the numpy module
import numpy as np
# and we also load the matplotlib module for later plotting
import matplotlib.pyplot as plt
# %matplotlib inline
# data will be loaded from the file named 'time-series1N.npy' and saved in the data variable
data = np.load('time-series1N.npy')
# -
# #### Get a first idea about the data
#
# We don't know at all what the data is about. Let's try to get a first idea about what is contained in the varialbe `data`.
#
# - What is the type of the imported data?
# - What are the dimensions of the data (you can use the `np.shape()` function)?
# - Which data is contained in each dimension? What do you think?
# - Try to plot the data in a meaningful way.
# your code goes here
# #### Extract simple statistics from the data
#
# We move on to get more detailed information about the data. Use `numpy` functions for the below stats.
#
# - Cacluate the ***maximum*** (`np.max()`), ***minimum*** (`np.min()`), ***mean*** (`np.mean()`) and ***standard deviation*** (`np.std()`) of the data.
# your code goes here
# #### Get information about the time axis
#
# The first row of the data contains the relative time elapsed during the experiment. Let's extract more information about the time axis.
#
# - What is the interval between data points? Try the `np.diff()` function and see what it returns.
# - Are all intervals equal? What is the sampling frequency of the data?
# +
# your code goes here
# -
# #### Plot histogram of the data
#
# Let's learn about the distribution of the recorded data by using the histogram function (`hist`) from `matplotlib.pyplot`.
#
# - Plot the histogram of the data using different bin sizes. Use the matplotlib hist function `plt.hist([data],bins=[number of bins])`. Start using `bins=10` and increase that value to see what happens.
# - What can you say about the distribution of the data (e.g. unimodal, bimodal, skewed, ...)?
# - Calculate the ***median*** as well as the following percentils: ***5th, 25th, 50th, 75th and 95th***. You can use the `numpy` median and percential functions `np.median()` and `np.percentile([data],[percentile])`.
# +
# your code goes here
# -
# ## Time series with time stamps - using pandas
# `pandas` provides operations for maniuplating time series in a very efficient way. Let's explore a time series using some of the functions provided by `pandas`.
# Note that you need the `time-series2.xlsx` file from the github repository to complete the below exercises. The file has to be located in the same directory as this notebook.
# #### Load data
# First we will load the data saved in an excel file using the pandas `read_excel` function.
# +
import pandas as pd
from pandas import ExcelWriter
from pandas import ExcelFile
data2 = pd.read_excel('time-series2.xlsx',sheet_name='NZRainfall',index_col='DATE') # load excel spreadsheet
data2.index = pd.to_datetime(data2.index) # convert the index column to the date format
# -
# The data format in which pandas keeps the data is call `DataFrame`.
# #### Get an idea about the data
#
# We already have an idea from the sheetname what the data is about. Let's find out more about the data.
#
# - How does that data look like? What information are contained in the data?
# - What is the interval/sampling frequency of the data?
# - What are the dimensions of the `DataFrame`?
# - Get the statistics of the data by using the pandas `[name of the DataFrame].describe()` function.
#
# **Hint :** You can simply display the `DataFrame` to the screen, or see the first lines with `data2.head()`.
# your code goes here
# The table is 2D array (3 column and 154 rows/entries) of time stamps and rainfall numbers of three towns - Auckland, Christchurch, Wellington - in New Zealand. The rainfall is measured monthly (interval is 1 month) and given for the period from Jan 2000 through Dec 2012.
# ### Plotting and slicing the data
#
# `pandas` has a build-in plot function which is called by `[name of the DataFrame].plot()`.
#
# - Plot the entire data.
# - Plot the data for the year 2004.
# - Plot the data in from the year 2006 through 2011.
# your code goes here
# your code goes here
# your code goes here
# ### Resample the data
#
# Let's leverage the build-in function to resample the time series data in pandas. Pandas allows to reample data with the `[name of DataFrame].resample()` function.
#
# - Get information on the input arguments of the `resample` function online.
# - Downsample the data to 6 month periods and 1 year periods (using the mean of the original data) and plot the data in each case.
# your code goes here
# your code goes here
# ## The end
|
tutorials/T04_Time-series-Empty.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.10.0 64-bit (''adventofcode-mOkh6lsX'': pipenv)'
# language: python
# name: python3
# ---
# # Matching 3D coordinates, with transformations
#
# - <https://adventofcode.com/2021/day/19>
#
# We are asked to figure out how many different beacons the scanners can 'see', by matching overlapping beacon coordinates. The trick is to know, without spending too much computing resources on this, when two sets of scanners are likely to be referencing the same set of beacons.
#
# To do this, I pre-calculate the distances between each pair of beacons in a cloud; this is easily achieved by using the [`scipy.spacial.distance.pdist()` function](https://docs.scipy.org/doc/scipy/reference/generated/scipy.spatial.distance.pdist.html). A cloud of $n$ beacons can be paired up into $n(n-1)/2$ pairs, and the distances between these pairs remains constant no matter what rotation you apply. You can then _intersect_ the sets of distances for two scanners, and see if _enough_ of those distances match. For two scanners to have 12 beacons in common, you need at least $12(12 - 1)/2 = 66$ distances matching.
#
# Once you have determined that two scanners do overlap (have at least 66 distances in common), we need to figure out the correct rotation for the second scanner. I used numpy vectorisation here; a single matrix of rotation transformations is used to produce all 24 possible rotations of the beacon cloud, with a dot product operation (a single `@` matrix multiplication). You can then try each of the beacons involved with one of the matching distances (we don't know which of these will match with which beacon in the other scanner, but their number is limited to 2 or 4 or so), and see if shifting the rotated beacon positions leads to a match.
#
# Determining which of the 24 orientations has a match means we need to figure out how many beacon positions are the same; the orientation with the most matches (provided there are at least 12), is the correct rotation. I used [`numpy.unique()`](https://numpy.org/doc/stable/reference/generated/numpy.unique.html) to generate an array of all unique vectors in both the beacon matrix of the scanner we are trying to match against and all rotations of the beacon cloud, together with the _inverse index_, an array where each position is the index into the unique values array for each input vector. You can then create two [sparse boolean matrices](https://docs.scipy.org/doc/scipy/reference/sparse.html) where rows and columns represent vectors in the input and the indices of the unique values, one for the fixed scanner, the other for all orientations of the scanner we are matching. A `True` value in any given cell connects a vector in one of the beacon clouds to one of the unique values, so representing the _set memberships_ of each cloud. If you then produce the _dot product_ of these two sparse matrices, you essentially create their intersection, and you can then sum this, per rotation, to get a count of intersections. Using [`numpy.argmax()`](https://numpy.org/doc/stable/reference/generated/numpy.argmax.html) on this gives us the index of the rotation that matches. With the right orientation, and a known distance, you can then use the updated scanner object to help locate the other scanners.
#
# To position all scanners, you start with a list with a single positioned scanner (it doesn't actually matter which one). I then use a [dequeu](https://docs.python.org/3/library/collections.html#collections.deque) (double-ended queue) for all scanners without a definite positon, and as long as there are scanners in the queue, take the next one and test that one against all positioned scanners, one by one, until I find a match. If matched, it is added to the positioned scanners list, otherwise, if there was no match against any of the positioned scanners, the tested scanner is added to the back of the queue again. Once they all have a position, we only have to count the unique beacon vectors.
#
# And because everything is achieved with vectorised array operations, the whole thing takes milliseconds to run.
#
# +
from __future__ import annotations
from collections import deque
from dataclasses import dataclass
from functools import cached_property
from itertools import combinations, permutations
from typing import Final
import math
import numpy as np
from scipy.spatial.distance import pdist
from scipy.sparse import lil_matrix
MIN_BEACONS_IN_COMMON: Final[int] = 12
MIN_COMMON_DISTANCES: Final[int] = math.comb(MIN_BEACONS_IN_COMMON, 2)
# Generate the 24 unique 3D rotation matrices
def _rotations() -> np.array:
# the identity transformation matrix, [1 0 0 0] [0 1 0 0] [0 0 1 0] [0 0 0 1]
eye = np.identity(4, dtype=np.int8)
# all permutations of 0, 1 and 2, padded with 3 to keep the bottom eye row in place.
# used to re-arrange the rows of the eye matrix
rows = np.pad(
np.array(list(permutations(range(3)))), ((0, 0), (0, 1)), constant_values=3
)
# the product of (-1, 1), times 3, with a 1 added to the end; these are the
# signs for each row of the rotation matrix.
signs = np.pad(
np.array([-1, 1])[
np.stack(np.meshgrid(*([np.arange(2)] * 3)), axis=-1).reshape(-1, 3)
],
((0, 0), (0, 1)),
constant_values=1,
)
# produce the product of signs and rows.
signs, rows = np.repeat(signs, rows.shape[0], axis=0), np.tile(
rows, (signs.shape[0], 1)
)
# lower half of the transformation matrix, used to calculate permutation parity
# see https://en.wikipedia.org/wiki/Parity_of_a_permutation
tx, ty = np.tril_indices(3, -1)
rowsparity = np.prod(np.sign(rows[:, tx] - rows[:, ty]), axis=-1)
signsparity = np.prod(signs[:, :3], axis=1)
# all signs and all permutations with the same parity
signs, rows = signs[rowsparity == signsparity], rows[rowsparity == signsparity]
count = signs.shape[0]
# alter eye with the signs combo (cols 0-3), then permute the rows (cols 4-7)
return (signs[:, :, None] * np.tile(eye, (count, 1)).reshape(-1, 4, 4))[
np.arange(count)[:, None], rows
]
ROTATIONS: Final[np.array] = _rotations()
NOR: int = ROTATIONS.shape[0]
@dataclass
class Scanner:
beacons: np.array
position: np.array = np.zeros(3, dtype=np.int16)
@classmethod
def from_lines(cls, lines: list[str]) -> Scanner:
return Scanner(np.genfromtxt(lines, delimiter=",", dtype=np.int16))
@cached_property
def distances(self) -> dict[np.float64, set[int]]:
"""Map from distance to pair of indices of beacons"""
map, combos = {}, combinations(range(self.beacons.shape[0]), 2)
for dist, pair in zip(pdist(self.beacons), combos):
try:
map[dist].update(pair)
except KeyError:
map[dist] = set(pair)
return map
@cached_property
def orientations(self) -> np.array:
# pad with 1s, apply the rotations transformation, then un-pad.
b = self.beacons
aug = np.concatenate((b, np.ones((*b.shape[:-1], 1), dtype=b.dtype)), axis=-1)
return (aug @ ROTATIONS)[..., :-1]
def __and__(self, other: Scanner) -> Scanner | None:
"""Check for scanner overlap
Returns new Scanner at correct rotation with position updated, relative to
other.
"""
# how many distances are the same? If enough match, there is overlap
shared = self.distances.keys() & other.distances.keys()
if sum(len(self.distances[d]) // 2 for d in shared) < MIN_COMMON_DISTANCES:
return None
# track some of the sizes involved, number of other and self beacons and
# orientations (reposititioned)
nob = other.beacons.shape[0]
nsb = self.beacons.shape[0]
nsr = nsb * NOR
# pick one of the beacons from other that we know has distances in common
distance = next(iter(shared))
reference = other.beacons[next(iter(other.distances[distance]))]
own_pairs = self.distances[distance]
# try all ends of the matching pairs in self; we don't know what side
# matches with the reference beacon.
for i in own_pairs:
offsets = reference - self.orientations[:, i]
repositioned = self.orientations + offsets[:, None, :]
# find unique vectors, and their inverse index, used to quantify
# how many vectors in a repositioned oriention fit.
values, ix = np.unique(
np.vstack((other.beacons, repositioned.reshape(-1, 3))),
axis=0,
return_inverse=True,
)
if values.shape[0] > nsr + nob - 12:
continue # not enough overlap between target beacons and repositioned
# find the matching orientation intersecting the beacons and repositioned
# matrix as sets; we count the unique values and create boolean matrices
# mapping vector values to their index in the beacons and repositioned
# matrices, then taking the dot product of these two mappings.
ix_beacons, ix_repos = ix[:nob], ix[nob:]
obvmembers = lil_matrix((nob, values.shape[0]), dtype=bool)
obvmembers[np.arange(nob), ix_beacons] = True
rvmembers = lil_matrix((nsr, values.shape[0]), dtype=bool)
rvmembers[np.arange(nsr), ix_repos] = True
matches = (obvmembers.tocsr() @ rvmembers.tocsr().T).T.sum(axis=-1)
counts = matches.reshape(-1, nsb).sum(axis=-1)
if not np.any(counts >= MIN_BEACONS_IN_COMMON):
continue
# orientation determined, get the corrected beacon positions
orientation = np.argmax(counts)
new_beacons = repositioned[orientation]
new_pos = reference - self.orientations[orientation][i]
return Scanner(new_beacons, new_pos)
@dataclass
class BeaconMap:
scanners: list[Scanner]
@classmethod
def from_text(cls, text: str) -> BeaconMap:
scanners = [
Scanner.from_lines(sc.splitlines()[1:]) for sc in text.split("\n\n")
]
return cls(scanners)
@cached_property
def positioned_scanners(self) -> list[Scanner]:
to_position = deque(self.scanners)
positioned = [to_position.popleft()]
while to_position:
scanner = to_position.popleft()
for other in positioned:
if placed := scanner & other:
positioned.append(placed)
break
else:
to_position.append(scanner)
return positioned
@cached_property
def positioned_beacons(self) -> set[np.array]:
return {pos for s in self.positioned_scanners for pos in zip(*s.beacons.T)}
test_map = BeaconMap.from_text(
"""\
--- scanner 0 ---\n404,-588,-901\n528,-643,409\n-838,591,734\n390,-675,-793
-537,-823,-458\n-485,-357,347\n-345,-311,381\n-661,-816,-575\n-876,649,763
-618,-824,-621\n553,345,-567\n474,580,667\n-447,-329,318\n-584,868,-557
544,-627,-890\n564,392,-477\n455,729,728\n-892,524,684\n-689,845,-530
423,-701,434\n7,-33,-71\n630,319,-379\n443,580,662\n-789,900,-551\n459,-707,401
--- scanner 1 ---\n686,422,578\n605,423,415\n515,917,-361\n-336,658,858
95,138,22\n-476,619,847\n-340,-569,-846\n567,-361,727\n-460,603,-452
669,-402,600\n729,430,532\n-500,-761,534\n-322,571,750\n-466,-666,-811
-429,-592,574\n-355,545,-477\n703,-491,-529\n-328,-685,520\n413,935,-424
-391,539,-444\n586,-435,557\n-364,-763,-893\n807,-499,-711\n755,-354,-619
553,889,-390
--- scanner 2 ---\n649,640,665\n682,-795,504\n-784,533,-524\n-644,584,-595
-588,-843,648\n-30,6,44\n-674,560,763\n500,723,-460\n609,671,-379\n-555,-800,653
-675,-892,-343\n697,-426,-610\n578,704,681\n493,664,-388\n-671,-858,530
-667,343,800\n571,-461,-707\n-138,-166,112\n-889,563,-600\n646,-828,498
640,759,510\n-630,509,768\n-681,-892,-333\n673,-379,-804\n-742,-814,-386
577,-820,562
--- scanner 3 ---\n-589,542,597\n605,-692,669\n-500,565,-823\n-660,373,557
-458,-679,-417\n-488,449,543\n-626,468,-788\n338,-750,-386\n528,-832,-391
562,-778,733\n-938,-730,414\n543,643,-506\n-524,371,-870\n407,773,750
-104,29,83\n378,-903,-323\n-778,-728,485\n426,699,580\n-438,-605,-362
-469,-447,-387\n509,732,623\n647,635,-688\n-868,-804,481\n614,-800,639
595,780,-596
--- scanner 4 ---\n727,592,562\n-293,-554,779\n441,611,-461\n-714,465,-776
-743,427,-804\n-660,-479,-426\n832,-632,460\n927,-485,-438\n408,393,-506
466,436,-512\n110,16,151\n-258,-428,682\n-393,719,612\n-211,-452,876
808,-476,-593\n-575,615,604\n-485,667,467\n-680,325,-822\n-627,-443,-432
872,-547,-609\n833,512,582\n807,604,487\n839,-516,451\n891,-625,532
-652,-548,-490\n30,-46,-14
"""
)
assert len(test_map.positioned_beacons) == 79
# +
import aocd
beacon_map = BeaconMap.from_text(aocd.get_data(day=19, year=2021))
print("Part 1:", len(beacon_map.positioned_beacons))
# -
# # Part 2
#
# For part two, we only need to know the maximum Manhattan distance between the scanner positions. That's trivial to produce, all we need to do is use the `scipy.distance.pdist()` function again, this time on all scanner positions and with the `"cityblock"` metric instead of the default `euclidian` metric, and take the maximum value.
#
# +
def max_distance(scanners: list[Scanner]) -> int:
return int(pdist(np.array([s.position for s in scanners]), "cityblock").max())
assert max_distance(test_map.positioned_scanners) == 3621
# -
print("Part 2:", max_distance(beacon_map.positioned_scanners))
|
2021/Day 19.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/duke-sunshine/NYU_Econometrics_Summer2021/blob/main/Economics_Milestone5_Causal_ipynb.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="baoVO4VazP9Q"
# # Data Preparation
# + [markdown] id="HMG1iPdFzTHr"
# Story Background: https://www.gemini.com/cryptopedia/the-dao-hack-makerdao
#
# Event: July 20, 2016, at block 192,000
# + [markdown] id="M1AUq193zmQg"
# [Data Metrics](https://github.com/coinmetrics-io/data/blob/master/csv/metrics.csv)
# + id="7tHY6FVuxaSN"
import pandas as pd
import numpy as np
# + [markdown] id="qt5hVJ3Mzz6g"
# ###import the data for Ethereum
# + colab={"base_uri": "https://localhost:8080/", "height": 342} id="eznPu_z5zvBc" outputId="d9d33b47-c5ad-4f81-b7cd-cec45c0ff22e"
df_eth=pd.read_csv("https://raw.githubusercontent.com/coinmetrics-io/data/master/csv/eth.csv")
df_eth.head()
# + colab={"base_uri": "https://localhost:8080/"} id="JwqB5hAk7row" outputId="e3aa900d-0fee-4505-c579-d1df00908544"
df_eth['time']=pd.to_datetime(df_eth['time'])
df_eth.dtypes
# + colab={"base_uri": "https://localhost:8080/", "height": 342} id="s5c7PnODz46B" outputId="633f7cae-2fa7-4bbf-9308-7e6a8569fcf3"
df_eth['Type'] = 'eth'
df_eth.head()
# + [markdown] id="P7GMGNoSz8FM"
# ### Import the Data for Ethereum Classic
# + colab={"base_uri": "https://localhost:8080/", "height": 342} id="Bp0Y-sPez91E" outputId="0d311c1f-b869-4d19-c50a-a198365304ac"
df_etc=pd.read_csv("https://raw.githubusercontent.com/coinmetrics-io/data/master/csv/etc.csv")
df_etc.head()
# + colab={"base_uri": "https://localhost:8080/"} id="qYVvsug1GWHU" outputId="2cd20dfa-91cc-4fcf-cc06-dd293c657d53"
### change to dateime
df_etc['time']=pd.to_datetime(df_etc['time'])
df_etc.dtypes
# + colab={"base_uri": "https://localhost:8080/", "height": 342} id="8aaYGs2sGhk6" outputId="8b9ffc3f-ec6b-4ea0-d656-031b8cc0ac1f"
df_etc['Type'] = 'etc'
df_etc.head()
# + [markdown] id="qgHxty-pxejg"
# # Regression Discontinguity
# + [markdown] id="3_Q5kfEJ7YWl"
# https://youtu.be/TfKwgGT2fSM
# + [markdown] id="w6zkeOJZ7ZY2"
# ### create the identifier varaible for Dao Hack
# + id="1-x1Zdvs8DRm"
from datetime import date
# + colab={"base_uri": "https://localhost:8080/", "height": 342} id="f4B5JEcG7PWC" outputId="debd2217-ccdf-4088-d9a9-eff612fd6d39"
df_eth ['After_DaoHack'] =df_eth['time'].apply(lambda x: 1 if x>= date(2016,7,20) else 0)
df_eth[df_eth.time.dt.date>=date(2016,7,18)].head()
# + colab={"base_uri": "https://localhost:8080/", "height": 342} id="vh_XxOpoD7Zz" outputId="4b4654ef-5230-4dae-c11c-2192dd6327b1"
df_eth['days']=df_eth.index
df_eth.head()
# + [markdown] id="J58BVTE1EBZM"
# ### normalize the data
# + colab={"base_uri": "https://localhost:8080/"} id="IIKWCP4mEgd8" outputId="e80924fb-bcc3-4917-9544-567dca6d151f"
df_eth['days'][df_eth.time.dt.date==date(2016,7,20)]
# + colab={"base_uri": "https://localhost:8080/"} id="W4EBX56wEEZI" outputId="8df5ccc5-9465-48ab-9571-0d0cf847632c"
df_eth['days']=df_eth['days']-356
df_eth[df_eth.time.dt.date>=date(2016,7,18)].head()
# + [markdown] id="9EnjgUYS8hyE"
# ## create the regression function
# + [markdown] id="I75Dz7Lk8mNU"
# https://www.statsmodels.org/stable/index.html
# + id="stLbYqt18hg7"
import statsmodels.api as sm
import statsmodels.formula.api as smf
# + id="rmA9-dnY8qgU"
results = smf.ols('PriceUSD ~ 1+ After_DaoHack +days+TxTfrValAdjUSD ', data=df_eth).fit()
# + colab={"base_uri": "https://localhost:8080/"} id="L_j8WQIm9F2w" outputId="4478d7af-8738-4644-c0e9-8dccdc5cb8ed"
print(results.summary())
# + id="AOCpbBdb9UmW"
results = smf.ols('PriceUSD ~ 1+ After_DaoHack + days +After_DaoHack*days +TxTfrValAdjUSD', data=df_eth).fit()
# + colab={"base_uri": "https://localhost:8080/"} id="pn4MDJj19ZHB" outputId="a3bb7063-a5fd-4a53-9ca5-42fa8c4891cb"
print(results.summary())
# + [markdown] id="ETLFygpxxjt5"
# # Difference in Difference
# + [markdown] id="4qdGg7YKxmuk"
# https://towardsdatascience.com/causal-inference-101-difference-in-differences-1fbbb0f55e85
# + [markdown] id="WL9RDDGyJhVq"
# ### prepare the datasets
# + id="5uOwr1ndITdh"
Columns =["time","PriceUSD","TxTfrValAdjUSD","FeeMedUSD","RevUSD","GasLmtTxMean","Type"]
df_eth =df_eth[Columns]
df_etc =df_etc[Columns]
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="OWbaXLXKIX0b" outputId="c2d07780-680d-4396-9fe6-d36e1e98358f"
df_etc =df_etc.dropna()
df_etc.head()
# + id="vkHvoa1wIAhh"
df_eth =df_eth[df_eth.time.dt.date>=date(2016,7,25)]
df_etc =df_etc[df_etc.time.dt.date>=date(2016,7,25)]
# + id="fd60jQVmI20J"
df_eth=df_eth.reset_index()
df_etc=df_etc.reset_index()
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="t8401h32IsgK" outputId="7b83b69e-80d6-4819-8692-590681d9316e"
df_eth["days"]=df_eth.index
df_eth.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="FZ-uORuVI8pS" outputId="2498bc36-e190-405f-bbbb-938bf2711dab"
df_etc["days"]=df_etc.index
df_etc.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="7n8h9KRFG1CF" outputId="e8d2a954-67dd-4307-928d-76afdeae19f4"
df = pd.concat([df_eth, df_etc], axis=0)
df.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="UfrlCbnhHXRb" outputId="8967d0c5-24da-4320-ea70-9f001866c7b7"
df["etc"]=df["Type"].apply(lambda x: 1 if x=="etc" else 0)
df.head()
# + [markdown] id="mUFwPq7KJkQw"
# ### run the regression
#
# Treatment: ETC hard-fork
# + id="oQ2RfZhUJQ0I"
import statsmodels.api as sm
import statsmodels.formula.api as smf
# + colab={"base_uri": "https://localhost:8080/"} id="Ns-3OEd-J1Zy" outputId="8519b045-79bb-4903-b2ec-afd806fa7c9e"
results = smf.ols('PriceUSD ~ 1+ etc +days+TxTfrValAdjUSD+FeeMedUSD+RevUSD+GasLmtTxMean', data=df).fit()
print(results.summary())
# + colab={"base_uri": "https://localhost:8080/"} id="gN-_gCC7K8p4" outputId="8101fb7b-cf5d-48e0-b4e7-bb51ef6e52c2"
results = smf.ols('PriceUSD ~ 1+ etc +days +etc*days+TxTfrValAdjUSD+FeeMedUSD+RevUSD+GasLmtTxMean', data=df).fit()
print(results.summary())
|
Economics_Milestone5_Causal_ipynb.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Web-Scraping Project
#
# ### Preparing for Web-Scraping on the following web sites.
#
# https://admin.nber.org/xsearch?q=early+childhood+development+OR+Education&whichsearch=ftpub&restrict_papers=yes&fullresults=1&datefilter=&b=search+again
#
# https://openknowledge.worldbank.org/discover
#
# https://www.unicef.org/search/search.php?querystring_en=%28%22early+childhood+development%22+OR+%22early+development%22+OR+%22early+child+care%22+OR+%22ecd%22%29+AND+%28%22literacy%22+OR+%22cognition%22+OR+%22education%22+OR+%22school*%22%29&hits=&type=&navigation=&Go.x=0&Go.y=0
#
# +
# Import Dependencies
# # !pip install BeautifulSoup4 as bs
# # !pip install pandas
# # !pip install splinter
from splinter import Browser
from bs4 import BeautifulSoup
import requests
import pandas as pd
import time
import random
# -
# # Mac Users
# https://splinter.readthedocs.io/en/latest/drivers/chrome.html
# !which chromedriver
executable_path = {'executable_path': '/usr/local/bin/chromedriver'}
browser = Browser('chrome', **executable_path, headless=False)
# # Windows Users
executable_path = {'executable_path': './chromedriver.exe'}
browser = Browser('chrome', **executable_path, headless=False)
# Using googlechrom drive open test explorer
url = 'https://openknowledge.worldbank.org/discover'
browser.visit(url)
# Retrieve page from the request module
response = requests.get(url)
# Create BeautifulSoup Object; parse with 'html.parser'
soup = BeautifulSoup(response.text, 'html.parser')
# Print the main page html
print(soup.prettify())
# results are returned as an iterable list
headers = soup.find_all('h4')
# Check the list of the headers
print(headers[0])
print(headers[1])
print(headers[3])
print(headers[9])
# +
# Loop through the headers on the main page
t = []
titles = []
for i in range(0, 10):
try:
#Look for the title on the main page under 'h4' tag
title = soup.find_all('h4')[i].text
# append the title name into an empty list, 't'
if (title):
t.append(title)
# print(title)
# Remove the new line split '\n' from the title name
name = t[i].splitlines()
titles.append(name[1])
# print(titles)
except AttributeError as e:
print(e)
# -
# Creating a click action on the book title
browser.click_link_by_text(titles[2])
# Create loop to click on each report link
for i in range(0, 10):
browser.click_link_by_text(titles[i])
html = browser.html
reports_soup = BeautifulSoup(html, 'html.parser')
slide_element = reports_soup.select_one('div.main-content')
publish_date = slide_element.find('div', class_='simple-item-view-other word-break').get_text()
print(publish_date)
browser.back()
print('page' + str(i+1))
browser.is_element_present_by_css('a[class="no-decor"]')
report_link = browser.find_by_css('a[class="no-decor"]')
report_link[0].click()
# Go back to the previous page
browser.back()
# +
# Find the next page button and click the link to the next page
# page = soup.find('a', class_='next-page-link')
# print(page)
browser.is_element_present_by_css('a[class="next-page-link"]')
next_page = browser.find_by_css('a[class="next-page-link"]')
print(next_page[1])
next_page[1].click()
# -
# Find the first title and click the link
browser.is_element_present_by_css('a[class="no-decor"]')
report_link = browser.find_by_css('a[class="no-decor"]')
report_link.click()
print(report_link)
# Create report soup for extracting data from the report page
html = browser.html
reports_soup = BeautifulSoup(html, 'html.parser')
# Create slide element to find the information to store
slide_element = reports_soup.select_one('div.main-content')
print(slide_element)
# Target the title header text
slide_element.find('h2', class_='ds-div-head')
# +
# Store the journal title, summary, citation, link, publish date, and author
title = [slide_element.find('h2', class_='ds-div-head').get_text()]
summary = [slide_element.find('div', class_='okr-item-page-field-wrapper abstract').get_text().splitlines()]
citation = [slide_element.find('div', class_='citation').get_text().splitlines()]
link_path = slide_element.find('div', class_='okr-item-page-field-wrapper uri').get_text()
link = [link_path[4:].splitlines()]
publish_date = [slide_element.find('div', class_='simple-item-view-other word-break').get_text()]
author = [slide_element.find('div', class_='authorprofile-item-view-link').get_text().splitlines()]
print(title)
print(summary)
print(citation)
print(link)
print(publish_date)
print(author)
# -
# Go back to the previous page
browser.back()
# +
# Loop through the headers on the main page
titles = []
report_title = []
report_summary = []
report_citation = []
report_link = []
report_publish_date = []
report_author = []
for i in range(0, 10):
try:
#Look for the title on the main page under 'h4' tag
name = soup.find_all('h4')[i].text
# Remove the new line split '\n' from the title name
if (name):
t = name.splitlines()
titles.append(t[1])
# print(titles)
# Click on the book title link and nevigate to the report page
browser.click_link_by_text(titles[i])
# Creating the BeautifulSoup parser to extract the information from the report page
html = browser.html
reports_soup = BeautifulSoup(html, 'html.parser')
report = reports_soup.select_one('div.main-content')
# Extracting the information from the report page
title = report.find('h2', class_='ds-div-head').get_text()
summary = report.find('div', class_='okr-item-page-field-wrapper abstract').get_text().splitlines()
citation = report.find('div', class_='citation').get_text().splitlines()
link_path = report.find('div', class_='okr-item-page-field-wrapper uri').get_text()
link = link_path[4:].splitlines()
publish_date = report.find('div', class_='simple-item-view-other word-break').get_text()
author = report.find('div', class_='authorprofile-item-view-link').get_text().splitlines()
# Append the information into the empty lists
report_title.append(title)
report_summary.append(summary[1])
report_citation.append(citation[1])
report_link.append(link[1])
report_publish_date.append(publish_date)
report_author.append(author[1])
# Go back to the main page after extracting the information
browser.back()
except AttributeError as e:
print(e)
print(report_title)
print(report_link)
# -
# print the popup windown html
html = browser.html
page_soup = BeautifulSoup(html, 'html.parser')
print(page_soup.prettify())
# Click the "No, thanks." button on the popup window
browser.click_link_by_text('No, thanks.')
# +
# Create a loop that can navigate to the next page
report_title = []
report_summary = []
report_citation = []
report_link = []
report_publish_date = []
report_author = []
executable_path = {'executable_path': './chromedriver.exe'}
browser = Browser('chrome', **executable_path, headless=False)
url = 'https://openknowledge.worldbank.org/discover'
browser.visit(url)
response = requests.get(url)
soup = BeautifulSoup(response.text, 'html.parser')
# Loop through the first 5 pages
for j in range(0, 5):
titles = []
for i in range(0, 10):
try:
html = browser.html
page_soup = BeautifulSoup(html, 'html.parser')
if browser.is_text_present('No, thanks.') == True:
browser.click_link_by_text('No, thanks.')
#Look for the title on the main page under 'h4' tag
browser.is_element_present_by_css('h4')
name = page_soup.find_all('h4')[i].text
# Remove the new line split '\n' from the title name
if (name):
t = name.splitlines()
titles.append(t[1])
# Click on the book title link and nevigate to the report page
browser.is_element_present_by_text(titles[i], wait_time=2)
browser.click_link_by_text(titles[i])
# Creating the BeautifulSoup parser to extract the information from the report page
html = browser.html
reports_soup = BeautifulSoup(html, 'html.parser')
report = reports_soup.select_one('div.main-content')
# Extracting the information from the report page
title = report.find('h2', class_='ds-div-head').get_text()
summary = report.find('div', class_='okr-item-page-field-wrapper abstract').get_text().splitlines()
citation = report.find('div', class_='citation').get_text().splitlines()
link_path = report.find('div', class_='okr-item-page-field-wrapper uri').get_text()
link = link_path[4:].splitlines()
publish_date = report.find('div', class_='simple-item-view-other word-break').get_text()
author = report.find('div', class_='authorprofile-item-view-link').get_text().splitlines()
# Append the information into the empty lists
report_title.append(title)
report_summary.append(summary[1])
report_citation.append(citation[1])
report_link.append(link[1])
report_publish_date.append(publish_date)
report_author.append(author[1])
# Go back to the main page after extracting the information
browser.back()
# Stop the loop for 2 seconds to make sure catching the popup window
time.sleep(2)
except AttributeError as e:
print(e)
# Create a click action to navigate to the next page
browser.is_element_present_by_css('a[class="next-page-link"]', wait_time=1)
next_page = browser.find_by_css('a[class="next-page-link"]')
next_page[1].click()
time.sleep(2)
print(report_link)
print(len(report_summary))
# -
print(len(report_link))
print(len(report_publish_date))
print(report_summary[0])
# Test to extract title from the second main page
# Create page_soup when the browser is at the second main page
html = browser.html
page_soup = BeautifulSoup(html, 'html.parser')
name = page_soup.find_all('h4')[1].text
print(name)
# +
report_abstract = []
for i in report_summary:
abstract = i[8:]
# print(abstract)
report_abstract.append(abstract)
print(report_abstract)
# -
# Storing the variables into a dataframe
data_df = pd.DataFrame(list(zip(report_title, report_abstract, report_link, report_publish_date, report_author)),
columns=["title", "summary", "link", "publish_date", "author"])
# Replace the "\n" in the dataframe
data_df = data_df.replace('\n',' ', regex=True)
data_df.head()
# Saving the dataframe into CSV
data_df.to_csv("data.csv", index=False, encoding='utf-8')
# Scraping NEBR
url = "https://admin.nber.org/xsearch?q=early+childhood+development+OR+Education&whichsearch=ftpub&restrict_papers=yes&fullresults=1&datefilter=&b=search+again"
browser.visit(url)
# Retrieve page from the request module
response = requests.get(url)
# Create BeautifulSoup Object; parse with 'html.parser'
soup = BeautifulSoup(response.text, 'html.parser')
# Print the main page html
print(soup.prettify())
# header results are returned as an iterable list
headers = soup.find_all('a', class_="resultTitle")
# Check the list of the headers
print(len(headers))
print(headers[0].get_text())
print(headers[1].get_text())
print(headers[3].get_text())
print(headers[9].get_text())
# publish date
publish_date = soup.find_all('span', class_='searchResultNiceDate')
print(len(publish_date))
print(publish_date[0].get_text())
# Authors
authors = soup.find_all('span', class_='searchResultAuthor')
print(len(authors))
print(authors[0].get_text())
# Summary
summary = soup.find_all('div', class_='searchResultAbstract')
print(len(summary))
print(summary[0].get_text())
# URL
link = soup.find_all('p', class_='url')
print(len(link))
print(link[0].get_text())
# Find the next page button and click on it
browser.is_text_present('More results.')
browser.click_link_by_text('More results.')
# Create a new soup for the new page
html = browser.html
soup = BeautifulSoup(html, 'html.parser')
# +
# Creating Loop for browsing on NBER web site
executable_path = {'executable_path': './chromedriver.exe'}
browser = Browser('chrome', **executable_path, headless=False)
url = "https://admin.nber.org/xsearch?q=early+childhood+development+OR+Education&whichsearch=ftpub&restrict_papers=yes&fullresults=1&datefilter=&b=search+again"
browser.visit(url)
for i in range(0, 10):
try:
html = browser.html
page_soup = BeautifulSoup(html, 'html.parser')
# results are returned as an iterable list
headers = soup.find_all('a', class_="resultTitle")
publish_date = soup.find_all('span', class_='searchResultNiceDate')
authors = soup.find_all('span', class_='searchResultAuthor')
summary = soup.find_all('div', class_='searchResultAbstract')
link = soup.find_all('p', class_='url')
print(len(headers))
t = random.randint(5, 15)
browser.is_element_present_by_text('More results', wait_time=t)
browser.click_link_by_text('More results.')
except AttributeError as e:
print(e)
# -
random.randint(5, 15)
|
USF_Web_Scraping_Project.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Classwork 4
#
# ### _<NAME>, <NAME>, <NAME>_:
# # The Sieve of Eratosthenes
#
# #### **The goal of the algorithm and how it works**
# It is an ancient algorithm for finding all prime numbers up to any given limit. We start by producing a list of all the numbers lower than the given value starting at 2 in a list, then we traverse through the list one at a time, removing any number if it is divisible by another number within the list. We are then left with only prime numbers remaining in our list, lower than our given value.
# #### **Describe your design decision:**
# We have two different functions in our module which finds the prime list in two different ways.
# eratosthenes(n) - uses the sieve of eratosthenes algorithm with nested while loops
# eratosthenes2(n) - wraps around a generator function to calculate the prime numbers
#
# The first function calls the for loop and cycles through as many times as need to. This is easy for readers to see as everything is done in one function.
# The second function uese a generator function. The generator can run infinitely and we only need to call it as many times as we need. It is also easy to edit and find errors as the work is split in two different functions, and is also easy for readers as each function has a specific task to perform.
#
#
# #### **Which data structure(s) did you use and why?:**
# We chose to use a list to store our numbers because we can add or delete to a list easiliy. Whereas a tuple, for example, would be more permanent and we would be unable to remove numbers from it. We also did not used a dictonary as we felt the list would store exacly what we needed and a dictionary also has keys for it's variables, which we decided was unnecessary for this particular module.
#
# ### INSTRUCTOR COMMENT:
# Why not a set?
# ### Code:
# +
import primes
print(primes.eratosthenes(5))
print(primes.eratosthenes2(5))
# -
# This shows that both of our functions: eratosthenes(n) and eratosthenes2(n) are both able to produce the correct output.
# ### INSTRUCTOR COMMENTS
# Generally, if you want to show that a sequence is correct, you need more than 2 elements! It is more convincing to show something nontrivial.
|
cw04-primes.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:miniconda3]
# language: python
# name: conda-env-miniconda3-py
# ---
# +
# %matplotlib inline
import mne
import matplotlib.pyplot as plt
fname = "oddball-epo.fif"
epochs = mne.read_epochs(fname)
# -
# ## Evokeds
# Finally, if we average an epoched dataset over trials, we can use the `mne.Evoked` object.
target = epochs["target"].average()
target
standard = epochs["standard"].average()
# To quickly investigate evoked activity, the `Evoked` object has a number of plotting functions available.
target.plot_joint();
# For condition contrasts, you can use `mne.combine.evoked`:
diff = mne.combine_evoked((target, -standard), weights='equal')
diff.plot_joint(times=.35);
# Or as an image:
diff.plot_image();
# + [markdown] hideCode=true hidePrompt=false
# Because we have a 10/20 electrode layout, we can easily use a somewhat nicer layout:
# -
rois = mne.channels.make_1020_channel_selections(diff.info, midline="z12")
diff.plot_image(group_by=rois, show=False, show_names="all");
# To contrast multiple conditions, `mne.viz.plot_compare_evokeds` is available:
mne.viz.plot_compare_evokeds({"standard": standard,
"target": target}, picks=[13]);
# ## Time-Frequency stuff
# For an overview over the spectral shape of the data, we can use a plotting method of `raw`, `raw.plot_psd`:
epochs_for_tfr = mne.read_epochs("oddball-long-epo.fif")
epochs_for_tfr.plot_psd(fmin=2, fmax=20);
# But what about the time/frequency correlates of the Oddball effect?
#
# We will extract power per time and frequency with Morlet wavelets.
from mne.time_frequency import tfr_morlet
freqs = list(range(3, 30))
tfr_target = tfr_morlet(epochs_for_tfr["target"], freqs, 3, return_itc=False)
tfr_standard = tfr_morlet(epochs_for_tfr["standard"], freqs, 3, return_itc=False)
# Time-frequency data (single trial or averaged) is stored in TFR objects. These objects behave in many ways like Evoked objects ...
tfr_contrast = mne.combine_evoked((tfr_standard, tfr_target), (-.5, .5))
tfr_contrast.apply_baseline((None, 0))
# Plotting time-frequencyy activity (event-related spectral perturbations): observe the alpha-band ERD and the time-frequency correlates of the P3 effect.
tfr_contrast.plot_joint();
tfr_contrast.plot(picks=[27]);
del epochs_for_tfr
# ## Statistics
# Remember what the data look like:
diff.plot_image(group_by=rois, show=False, show_names="all");
# Can we statistically threshold this image to see which effects are reliable?
# ### Cluster-based permutation stats
# Exploratory analysis with nonparametric control of the error rate is commonly done with
# cluster-based permutation tests (i.e., Maris 2012). To cluster across space, we first need a
# channel adjacency matrix.
from mne.channels import find_ch_connectivity
connectivity, ch_names = find_ch_connectivity(epochs.info, ch_type='eeg')
plt.imshow(connectivity.toarray(), cmap="Greys")
# Now we need the data in the right shape. Sadly, because the space dimension needs
# to be last, we need to manually swap the time and space axes.
epochs.pick_types(eeg=True)
target_epochs, standard_epochs = epochs["target"].get_data(), epochs["standard"].get_data()
target_epochs.shape, standard_epochs.shape
target_epochs = target_epochs.swapaxes(1, 2)
standard_epochs = standard_epochs.swapaxes(1, 2)
target_epochs.shape, standard_epochs.shape
# MNE has various cluster-based permutation test options. Here, we test for single-trial
# differences between conditions with `mne.stats.spatio_temporal_cluster_test`.
#
# We use threshold-free cluster enhancement to reduce the number of parameters.
#
# Warning: the next cell takes a lot of time and computational power.
# +
from mne.stats import spatio_temporal_cluster_test
mne.set_log_level(True)
tfce = dict(start=.2, step=.5) # decrease both for real analyses
cluster_stats = spatio_temporal_cluster_test([target_epochs, standard_epochs],
threshold=tfce,
n_permutations=200, # way too low, increase for real analyses
n_jobs=1, # increase for decent CPUs
connectivity=connectivity)
T_obs, clusters, p_values, _ = cluster_stats
# -
# Now we can visualise the *t* values over time and space ...
extent = (*epochs.times[[0, -1]], 0, len(epochs.ch_names))
im = plt.imshow(T_obs.T, aspect="auto", cmap="RdBu_r",
vmin=-100, vmax=100, extent=extent
)
plt.colorbar(im)
# ... and the p-values.
# +
plt.hist(p_values)
alpha = .01
print(sum(p_values < alpha))
# -
# We can use the resulting mask to mask the image:
# +
pvals = p_values.reshape(T_obs.shape).T < alpha
diff.plot_image(group_by=rois, show=False, show_names="all", mask=pvals);
# -
# ### Parametric stats
# Sometimes, e.g. because we wish to test a specific hypothesis, cluster-based permutation tests are too much.
# We can also simply access the data in array form and test with parametric (or nonparametric) tests.
# For this, we first need to identify the spatial and temporal coordinates of an effect we want to test -
# for example, the N2 at Cz.
time_mask = (.2 < epochs.times) & (epochs.times < .25)
electrode_pz = epochs.ch_names.index("Cz")
plt.plot(time_mask)
# Now we extract the target data. Reminder: the shape of epochs data is (trial, channel, time)
epochs["target"].get_data().shape
cond_a = epochs["target"].get_data()[:, electrode_pz, time_mask].mean(-1)
cond_b = epochs["standard"].get_data()[:, electrode_pz, time_mask].mean(-1)
cond_a.shape
# Now we can simply use ordinary tests on these statistics.
from scipy.stats import ttest_ind, wilcoxon
ttest_ind(cond_a, cond_b)
wilcoxon(cond_a, cond_b)
# It is also straight-forward to convert the data into a (pandas) dataframe.
df = epochs.to_data_frame()
df.head(20)
df_cz = df.query("200 < time < 250")["Cz"].groupby(["epoch", "condition"]).mean().reset_index()
df_cz.head()
import seaborn as sns
sns.factorplot(y="Cz", data=df_cz, x="condition")
|
2018_06_Amsterdam/mne_notebook_2_evoked_data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#Importing libraries
from __future__ import print_function
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina'
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import seaborn as sns
import warnings
warnings.filterwarnings('ignore')
pd.options.display.float_format = '{:,.2f}'.format
pd.set_option('display.max_rows', 100)
pd.set_option('display.max_columns', 200)
from datetime import datetime
from matplotlib.colors import ListedColormap
from sklearn.datasets import make_classification, make_moons, make_circles
from sklearn.metrics import confusion_matrix, classification_report, mean_squared_error, mean_absolute_error, r2_score
from sklearn.linear_model import LogisticRegression
from sklearn.utils import shuffle
from keras.models import Sequential
from keras.layers import Dense, Dropout, BatchNormalization, Activation
from keras.optimizers import Adam
from keras.callbacks import EarlyStopping
from keras.utils.np_utils import to_categorical
from sklearn.preprocessing import StandardScaler, LabelEncoder, OneHotEncoder, MinMaxScaler
from sklearn.model_selection import train_test_split, cross_val_score, StratifiedKFold, KFold
import keras.backend as K
from keras.wrappers.scikit_learn import KerasClassifier
# +
def plot_decision_boundary(func, X, y, figsize=(9, 6)):
amin, bmin = X.min(axis=0) - 0.1
amax, bmax = X.max(axis=0) + 0.1
hticks = np.linspace(amin, amax, 101)
vticks = np.linspace(bmin, bmax, 101)
aa, bb = np.meshgrid(hticks, vticks)
ab = np.c_[aa.ravel(), bb.ravel()]
c = func(ab)
cc = c.reshape(aa.shape)
cm = plt.cm.RdBu
cm_bright = ListedColormap(['#FF0000', '#0000FF'])
fig, ax = plt.subplots(figsize=figsize)
contour = plt.contourf(aa, bb, cc, cmap=cm, alpha=0.8)
ax_c = fig.colorbar(contour)
ax_c.set_label("$P(y = 1)$")
ax_c.set_ticks([0, 0.25, 0.5, 0.75, 1])
plt.scatter(X[:, 0], X[:, 1], c=y, cmap=cm_bright)
plt.xlim(amin, amax)
plt.ylim(bmin, bmax)
def plot_multiclass_decision_boundary(model, X, y):
x_min, x_max = X[:, 0].min() - 0.1, X[:, 0].max() + 0.1
y_min, y_max = X[:, 1].min() - 0.1, X[:, 1].max() + 0.1
xx, yy = np.meshgrid(np.linspace(x_min, x_max, 101), np.linspace(y_min, y_max, 101))
cmap = ListedColormap(['#FF0000', '#00FF00', '#0000FF'])
Z = model.predict_classes(np.c_[xx.ravel(), yy.ravel()], verbose=0)
Z = Z.reshape(xx.shape)
fig = plt.figure(figsize=(8, 8))
plt.contourf(xx, yy, Z, cmap=plt.cm.Spectral, alpha=0.8)
plt.scatter(X[:, 0], X[:, 1], c=y, s=40, cmap=plt.cm.RdYlBu)
plt.xlim(xx.min(), xx.max())
plt.ylim(yy.min(), yy.max())
def plot_data(X, y, figsize=None):
if not figsize:
figsize = (8, 6)
plt.figure(figsize=figsize)
plt.plot(X[y==0, 0], X[y==0, 1], 'or', alpha=0.5, label=0)
plt.plot(X[y==1, 0], X[y==1, 1], 'ob', alpha=0.5, label=1)
plt.xlim((min(X[:, 0])-0.1, max(X[:, 0])+0.1))
plt.ylim((min(X[:, 1])-0.1, max(X[:, 1])+0.1))
plt.legend()
def plot_loss_accuracy(history):
historydf = pd.DataFrame(history.history, index=history.epoch)
plt.figure(figsize=(8, 6))
historydf.plot(ylim=(0, max(1, historydf.values.max())))
loss = history.history['loss'][-1]
acc = history.history['acc'][-1]
plt.title('Loss: %.3f, Accuracy: %.3f' % (loss, acc))
def plot_loss(history):
historydf = pd.DataFrame(history.history, index=history.epoch)
plt.figure(figsize=(8, 6))
historydf.plot(ylim=(0, historydf.values.max()))
plt.title('Loss: %.3f' % history.history['loss'][-1])
def plot_confusion_matrix(model, X, y):
y_pred = model.predict_classes(X, verbose=0)
plt.figure(figsize=(8, 6))
sns.heatmap(pd.DataFrame(confusion_matrix(y, y_pred)), annot=True, fmt='d', cmap='YlGnBu', alpha=0.8, vmin=0)
def plot_compare_histories(history_list, name_list, plot_accuracy=True):
dflist = []
min_epoch = len(history_list[0].epoch)
losses = []
for history in history_list:
h = {key: val for key, val in history.history.items() if not key.startswith('val_')}
dflist.append(pd.DataFrame(h, index=history.epoch))
min_epoch = min(min_epoch, len(history.epoch))
losses.append(h['loss'][-1])
historydf = pd.concat(dflist, axis=1)
metrics = dflist[0].columns
idx = pd.MultiIndex.from_product([name_list, metrics], names=['model', 'metric'])
historydf.columns = idx
plt.figure(figsize=(6, 8))
ax = plt.subplot(211)
historydf.xs('loss', axis=1, level='metric').plot(ylim=(0,1), ax=ax)
plt.title("Training Loss: " + ' vs '.join([str(round(x, 3)) for x in losses]))
if plot_accuracy:
ax = plt.subplot(212)
historydf.xs('acc', axis=1, level='metric').plot(ylim=(0,1), ax=ax)
plt.title("Accuracy")
plt.xlabel("Epochs")
plt.xlim(0, min_epoch-1)
plt.tight_layout()
def make_sine_wave():
c = 3
num = 2400
step = num/(c*4)
np.random.seed(0)
x0 = np.linspace(-c*np.pi, c*np.pi, num)
x1 = np.sin(x0)
noise = np.random.normal(0, 0.1, num) + 0.1
noise = np.sign(x1) * np.abs(noise)
x1 = x1 + noise
x0 = x0 + (np.asarray(range(num)) / step) * 0.3
X = np.column_stack((x0, x1))
y = np.asarray([int((i/step)%2==1) for i in range(len(x0))])
return X, y
def make_multiclass(N=500, D=2, K=3):
"""
N: number of points per class
D: dimensionality
K: number of classes
"""
np.random.seed(0)
X = np.zeros((N*K, D))
y = np.zeros(N*K)
for j in range(K):
ix = range(N*j, N*(j+1))
# radius
r = np.linspace(0.0,1,N)
# theta
t = np.linspace(j*4,(j+1)*4,N) + np.random.randn(N)*0.2
X[ix] = np.c_[r*np.sin(t), r*np.cos(t)]
y[ix] = j
fig = plt.figure(figsize=(6, 6))
plt.scatter(X[:, 0], X[:, 1], c=y, s=40, cmap=plt.cm.RdYlBu, alpha=0.8)
plt.xlim([-1,1])
plt.ylim([-1,1])
return X, y
# -
#Load data
dataset=pd.read_csv('C:\\Users\\sagi\\Desktop\\Learning\\ML\\Datasets\\petrol_consumption.csv')
#Exploring data
sns.heatmap(dataset.corr(), annot=True, vmin=-1, vmax=1)
dataset.hist(figsize=(10, 8))
plt.tight_layout()
plt.figure(figsize=(6, 8))
sns.heatmap(dataset.corr()[['Petrol_Consumption']], annot=True, vmin=-1, vmax=1)
#Normalizing
ss = StandardScaler()
scale_features = ['Average_income','Paved_Highways','Petrol_tax','Average_income','Petrol_Consumption']
dataset[scale_features] = ss.fit_transform(dataset[scale_features])
# +
#Split the data
dataset.shape
X=dataset.iloc[:,:4].values
y=dataset.iloc[:,4].values
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=0)
print(X_train.shape, y_train.shape, X_test.shape, y_test.shape)
# -
#Check some more correlations
plt.figure(figsize=(4, 8))
tempdf = dataset.corr()[['Petrol_Consumption']].sort_values('Petrol_Consumption', ascending=False).iloc[:20, :]
sns.heatmap(tempdf, annot=True, vmin=-1, vmax=1)
# +
#Linear Regression Model
linr_model = Sequential()
linr_model.add(Dense(1, input_shape=(X.shape[1],)))
linr_model.compile('adam', 'mean_squared_error')
linr_history = linr_model.fit(X_train, y_train, epochs=1500, verbose=0, validation_split=0.2)
plot_loss(linr_history)
linr_model.evaluate(X_test, y_test, verbose=0)
# +
#linr_model.evaluate(X_test, y_test, verbose=0)
# -
#weights data frame
linr_wdf = pd.DataFrame(linr_model.get_weights()[0].T,
columns=dataset.drop(['Petrol_Consumption'], axis=1).columns).T.sort_values(0, ascending=False)
linr_wdf.columns = ['feature_weight']
linr_wdf.iloc[:20,:]
|
Regression-Predicting-a-real-valued-output-for-Petrol_Consumption.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Heap Class and Heap Operations
#
# The Heap_Class.ipynb contains following:
#
# **`Private Variables`**:<br>
# - **self.heap:** It is Python list containing the array elements organized in Heap Structure
# - **self.heap_type:** Tracks if the Heap is MinHeap or MaxHeap
# - **self.heap_size** Tracks Heap Size
#
# **`Class Methods`**:<br>
# **build_heap**: This method is used to build heap structure from a given array
#
# **insert**: This method is used to add an element in the heap struture and calls heapify_UP method to ensure the heap properties are preserved
#
# **delete**: This method is used to delete an element from the heap structure and calls heapify_DOWN method to ensure the heap properties are preserved
#
# **heapify_UP**: This method is called from the **insert** method when a new element is **inserted** to the heap structure. It scans the heap with a bottom-up approach starting from the newly added element(which is the last node in the structure and going up until it the heap properties are as desired.
#
#
# **heapify_DOWN**: This method is called from the **delete** method when a element is **deleted** from the heap structure. It scans the heap with Top-Down approach starting from the element to be deleted. In this method
# 1. Find the index of the element to be deleted
# 2. Swap the last node element with the element to be deleted
# 2. Pop out the last node element (which is the element we wanted to delete)
# 3. Starting from the original index of the element to be deleted, Go down the heap structure to ensure Heap properties are met
#
#
# **get_array():**: This method returns the heap array (self.heap)
#
#
# **get_heap_size():**: This method returns the size of the heap
#
#
|
DataStructures/Tree_DataStructures/HEAPS/About_Heap_Class_And_Operations.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.2 64-bit (''base'': conda)'
# name: python3
# ---
# +
import re
import folium
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from geopy.geocoders import Nominatim
from wordcloud import WordCloud, STOPWORDS
from geopy.extra.rate_limiter import RateLimiter
import ipywidgets as widgets
from ipywidgets import Layout
from IPython.display import display, HTML
# -
data = pd.read_csv("train.csv")
duplicated_data = data['text'].duplicated().sum()
data = data.drop_duplicates(subset=['text'], keep='first')
def clean_data(text):
text = text.lower()
text = re.sub(r'http[s]?:\/\/.*[\r\n]*', '', text)
text = re.sub(r'[^A-Za-z0-9 ]+', '', text)
text = " ".join(text.split())
return text
def remove_emojis(text):
emoji_pattern = re.compile("["
u"\U0001F600-\U0001F64F" # emoticons
u"\U0001F300-\U0001F5FF" # symbols & pictographs
u"\U0001F680-\U0001F6FF" # transport & map symbols
u"\U0001F1E0-\U0001F1FF" # flags (iOS)
u"\U00002702-\U000027B0"
u"\U000024C2-\U0001F251"
"]+", flags=re.UNICODE
)
return emoji_pattern.sub(r'', text)
data['text'] = data['text'].apply(clean_data)
data['text'] = data['text'].apply(remove_emojis)
data['location'] = data['location'].replace({
"United States": "USA",
"London, UK": "UK",
"Atlanta, GA": "Atlanta",
"New York City": "New York",
"NYC": "New York",
"San Francisco, CA": "San Francisco",
"California, USA": "California",
"Chicago, IL": "Chicago",
"Los Angeles, CA": "Los Angeles",
"New York, NY": "New York",
"United Kingdom": "UK",
"London, England": "London",
"Everywhere": "Worldwide",
"Earth": "Worldwide",
"Washington, DC": "Washington",
"Washington, D.C.": "Washington"
})
# <h1 align="center"> Disaster Tweets </h1>
# <br>
# +
target_values = ['Non-Disaster', 'Disaster']
target = data['target'].value_counts()
fig, ax = plt.subplots(1,2, figsize=(25,10))
colors = sns.color_palette('prism')
sns.barplot(x=target_values, y=target, palette=['#55B56A', '#ED6A5A'], ax=ax[0])
data.groupby('target').count()['id'].plot(kind='pie', ax=ax[1], labels=target_values, autopct="%.1f%%", colors = ['#55B56A', '#ED6A5A'], fontsize=18)
ax[0].set_xlabel('Target', fontsize=12)
ax[0].set_ylabel('Cantidad', fontsize=12)
ax[1].set_ylabel('')
plt.show()
# -
# <br>
# <br>
# +
target1 = data[data['target']==1]['keyword'].value_counts()
target0 = data[data['target']==0]['keyword'].value_counts()
target_data = [value for value in target1.index.tolist() if value in target0.index.tolist()]
unique_keyword = data.keyword.unique()
u = data['keyword'].value_counts().nlargest(10)
keywords = widgets.SelectMultiple(
options = target_data,
value = u.index.tolist(),
description='Keyword',
disabled = False,
layout = Layout(width='50%', height='80px', display='flex')
)
# +
location1 = data[data['target']==1]['location'].value_counts()
location0 = data[data['target']==0]['location'].value_counts()
target_data = [value for value in location1.index.tolist() if value in location0.index.tolist()]
unique_location = data.location.unique()
h = data[data['target']==1]['location'].value_counts().nlargest(10)
locations = widgets.SelectMultiple(
options = target_data,
value = h.index.tolist(),
description='Location',
disabled=False,
layout = Layout(width='50%', height='80px', display='flex')
)
# -
def disaster_keywords_plot(keywords):
temp_data = data.loc[data['keyword'].isin(keywords)]
disaster_keywords = temp_data[temp_data['target']==1]['keyword'].value_counts().nlargest(10)
non_disaster_keywords = temp_data[temp_data['target']==0]['keyword'].value_counts().nlargest(10)
fig, ax = plt.subplots(1,2, figsize=(25,10))
sns.barplot(x=disaster_keywords.tolist(), y=disaster_keywords.index, orient='h', palette=['#ED6A5A'], ax=ax[0])
sns.barplot(x=non_disaster_keywords.tolist(), y=non_disaster_keywords.index, orient='h', palette=['#55B56A'], ax=ax[1])
ax[0].set_title('Disaster Keywords\n', fontsize=20)
ax[0].set_xlabel('Cantidad', fontsize=12)
ax[0].set_ylabel('Palabra', fontsize=12)
ax[1].set_title('Non-Disaster Keywords\n', fontsize=20)
ax[1].set_xlabel('Cantidad', fontsize=12)
ax[1].set_ylabel('Palabra', fontsize=12)
plt.show()
widgets.interactive(disaster_keywords_plot, keywords=keywords)
# <br>
# <br>
def disaster_locations_plot(locations):
temp_data = data.loc[data['location'].isin(locations)]
disaster_locations = temp_data[temp_data['target']==1]['location'].value_counts().nlargest(10)
non_disaster_locations = temp_data[temp_data['target']==0]['location'].value_counts().nlargest(10)
fig, ax = plt.subplots(1,2, figsize=(25,10))
sns.barplot(x=disaster_locations.tolist(), y=disaster_locations.index, orient='h', palette=['#ED6A5A'], ax=ax[0])
sns.barplot(x=non_disaster_locations.tolist(), y=non_disaster_locations.index, orient='h', palette=['#55B56A'], ax=ax[1])
ax[0].set_title('Disaster Locations\n', fontsize=20)
ax[0].set_xlabel('Cantidad', fontsize=12)
ax[0].set_ylabel('Palabra', fontsize=12)
ax[1].set_title('Non-Disaster Locations\n', fontsize=20)
ax[1].set_xlabel('Cantidad', fontsize=12)
ax[1].set_ylabel('Palabra', fontsize=12)
plt.show()
widgets.interactive(disaster_locations_plot, locations=locations)
# <br>
# <br>
def disaster_map_plot(locations):
df = data['location'].value_counts()[:20,]
df = pd.DataFrame(df)
df = df.reset_index()
df.columns = ['location', 'counts']
geolocator = Nominatim(user_agent="main")
geocode = RateLimiter(geolocator.geocode, min_delay_seconds=1)
dictt_latitude = {}
dictt_longitude = {}
for i in df['location'].values:
location = geocode(i)
dictt_latitude[i] = location.latitude
dictt_longitude[i] = location.longitude
df['latitude']= df['location'].map(dictt_latitude)
df['longitude'] = df['location'].map(dictt_longitude)
map1 = folium.Map(location=[10.0, 10.0], tiles='CartoDB dark_matter', zoom_start=2.3)
markers = []
for i, row in df.iterrows():
loss = row['counts']
if row['counts'] > 0:
count = row['counts']*0.4
folium.CircleMarker([float(row['latitude']), float(row['longitude'])], radius=float(count), color='#ef4f61', fill=True).add_to(map1)
display(map1)
widgets.interactive(disaster_map_plot, locations=locations)
|
main.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="pfbg_NxOEZ-k"
# # Exercise Sheet 9
# + colab={} colab_type="code" id="3HYjFWp2EZ-o"
from __future__ import print_function, division
import numpy as np
import matplotlib.pyplot as plt
import sys
import tensorflow as tf
from tensorflow.keras.layers import Input, Dense, Reshape, Flatten, Dropout
from tensorflow.keras.layers import BatchNormalization, Activation, ZeroPadding2D
from tensorflow.keras.layers import LeakyReLU
from tensorflow.keras.layers import UpSampling2D, Conv2D
from tensorflow.keras.models import Sequential, Model
from tensorflow.keras.optimizers import Adam, RMSprop
# + [markdown] colab_type="text" id="dgzAZA0EEZ_A"
# ## GAN Optimisation
# + [markdown] colab_type="text" id="RDNkoCxqEZ_C"
# For the example $V(x, y) = x y$ perform alternating gradient descent updates with respect to $x$ and $y$ as we would perform when training a GAN, i.e. one player is performing updates to optimise $V$ and one player to optimise $−V$. Identify the trajectory of this update. Which ‘late-time’ behaviour do you find?
# + [markdown] colab_type="text" id="mDcKpLVOEZ_D"
# ### Solution
# + [markdown] colab_type="text" id="TvOD0QpxVuKP"
# First we see what happens when both players update with respect to the full gradient
# + colab={} colab_type="code" id="fKqDnXCsfE3T"
# X = [x, y]
V = lambda X: X[0] * X[1]
Vneg = lambda X: -V(X)
dV = lambda X: np.array([X[1], X[0]])
dVneg = lambda X: -dV(X)
# + colab={} colab_type="code" id="oNe47-FDeGD7"
def alt_grad_descent1(epochs, lrate):
X = np.random.normal(0,10,2)
hist = np.zeros([epochs ,2])
for j in range(epochs):
# minimize V
X = X - lrate*dV(X)
# minimize -V
X = X - lrate*dVneg(X)
hist[j] = X
return hist
hist = alt_grad_descent1(50000, 10**-2)
Vhist = [V(hist[i]) for i in range(len(hist))]
# + colab={"base_uri": "https://localhost:8080/", "height": 286} colab_type="code" id="BQCgGZXzPbj9" outputId="e380c8ba-25d7-4f48-b5a4-98b4a6a5985b"
plt.plot(Vhist)
# + colab={"base_uri": "https://localhost:8080/", "height": 286} colab_type="code" id="mGpx-aryqvbb" outputId="63e522b2-9ed7-4a28-f340-07e532d286ef"
plt.plot(hist[:,0], hist[:,1])
# + [markdown] colab_type="text" id="xS0YXqAiV3Bt"
# Now we check what happens when player 1 updates only the x-direction, while player 2 updates only the y-direction. See [here](https://arxiv.org/pdf/1801.04406.pdf) for the relation to GAN training
# + colab={} colab_type="code" id="os6jB8-GWBKw"
def alt_grad_descent2(epochs, lrate):
X = np.random.normal(0,10,2)
hist = np.zeros([epochs ,2])
for j in range(epochs):
# minimize V
X[0] = X[0] - lrate*dV(X)[0]
# minimize -V
X[1] = X[1] - lrate*dVneg(X)[1]
hist[j] = X
return hist
hist = alt_grad_descent2(50000, 10**-3)
Vhist = [V(hist[i]) for i in range(len(hist))]
# + colab={"base_uri": "https://localhost:8080/", "height": 286} colab_type="code" id="qCNM_aGDWO8j" outputId="c4057f03-6e81-4ce1-d2bb-19c64e1d0437"
plt.plot(Vhist)
# + colab={"base_uri": "https://localhost:8080/", "height": 286} colab_type="code" id="Fk4ZcO1aWVSb" outputId="d6c116f7-f2b7-467b-9128-e92c7f8391f1"
plt.plot(hist[:,0], hist[:,1])
# + [markdown] colab_type="text" id="JnawiBWUs_5V"
# ## GAN
# + [markdown] colab_type="text" id="YuwPFkkNs_5Y"
# The aim of this exercise is to implement a GAN architecture and to test it on the polynomial dataset from the previous exercise sheets. You can adapt the architecture from the MNIST example in the lectures.
# $\star$ Update your loss function to implement a Wasserstein GAN.
# Note that in case you do not have a local environment where you can calculate on a GPU, you can calculate online on collab on a GPU.
# + [markdown] colab_type="text" id="dsTG_su8XDoj"
# The references are: [GAN](https://arxiv.org/abs/1406.2661), [WGAN](https://arxiv.org/abs/1701.07875)
# + [markdown] colab_type="text" id="fPWdYWQls_5Z"
# ### Solution
# + colab={} colab_type="code" id="r8AkzCKKVpZd"
# !mkdir images
# + colab={} colab_type="code" id="HJv2sUfHpPWv"
# Polynomial data
size = 40
# evaluate polynomial over grid of size 40x40
def polynomial(degree):
coeff = np.random.normal(0,1,(degree+1, degree+1))
#coeff = np.random.uniform(-1,1,(degree+1, degree+1))
return [[sum([coeff[i,j]*((x/size)**i)*((y/size)**j)
for i in range(degree+1) for j in range(degree+1) if (i+j)<=degree])
for x in range(size)] for y in range(size)]
# training set of polynomial images of degree <=5
maxdegree = 5
size = 40
num_polys = 20000
polydata = np.array([polynomial(np.random.randint(0,maxdegree)) for i in range(num_polys)])
polydata = tf.keras.utils.normalize(polydata)
np.save('polydata.npy', polydata)
# + colab={} colab_type="code" id="dROAjd5Tpo0O"
polydata = np.load('polydata.npy')
# + colab={"base_uri": "https://localhost:8080/", "height": 286} colab_type="code" id="B2HGmBMMc-Q-" outputId="b74646aa-26ec-4e31-d549-0762cd38caa9"
plt.imshow(polydata[1])
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="7nUTkLKCj0_M" outputId="be7d7bfe-f51a-4181-b74d-0a38f7098336"
class DCGAN():
def __init__(self):
# Input shape
self.img_rows = 40
self.img_cols = 40
self.channels = 1
self.img_shape = (self.img_rows, self.img_cols, self.channels)
self.latent_dim = 2
optimizer = Adam(0.0002, 0.5)
# Build and compile the discriminator
self.discriminator = self.build_discriminator()
self.discriminator.compile(loss='binary_crossentropy',
optimizer=optimizer,
metrics=['accuracy'])
# Build the generator
self.generator = self.build_generator()
# The generator takes noise as input and generates imgs
z = Input(shape=(self.latent_dim,))
img = self.generator(z)
# For the combined model we will only train the generator
self.discriminator.trainable = False
# The discriminator takes generated images as input and determines validity
valid = self.discriminator(img)
# The combined model (stacked generator and discriminator)
# Trains the generator to fool the discriminator
self.combined = Model(z, valid)
self.combined.compile(loss='binary_crossentropy', optimizer=optimizer)
def build_generator(self):
model = Sequential()
model.add(Dense(128 * 10 * 10, activation="relu", input_dim=self.latent_dim))
model.add(Reshape((10, 10, 128)))
model.add(UpSampling2D())
model.add(Conv2D(128, kernel_size=3, padding="same"))
model.add(BatchNormalization(momentum=0.8))
model.add(Activation("relu"))
model.add(UpSampling2D())
model.add(Conv2D(64, kernel_size=3, padding="same"))
model.add(BatchNormalization(momentum=0.8))
model.add(Activation("relu"))
model.add(Conv2D(self.channels, kernel_size=3, padding="same"))
model.add(Activation("tanh"))
model.summary()
noise = Input(shape=(self.latent_dim,))
img = model(noise)
return Model(noise, img)
def build_discriminator(self):
model = Sequential()
model.add(Conv2D(16, kernel_size=3, strides=2, input_shape=self.img_shape, padding="same"))
model.add(LeakyReLU(alpha=0.2))
model.add(Dropout(0.25))
model.add(Conv2D(32, kernel_size=3, strides=2, padding="same"))
model.add(ZeroPadding2D(padding=((0,1),(0,1))))
model.add(BatchNormalization(momentum=0.8))
model.add(LeakyReLU(alpha=0.2))
model.add(Dropout(0.25))
model.add(Conv2D(64, kernel_size=3, strides=2, padding="same"))
model.add(BatchNormalization(momentum=0.8))
model.add(LeakyReLU(alpha=0.2))
model.add(Dropout(0.25))
model.add(Conv2D(128, kernel_size=3, strides=1, padding="same"))
model.add(BatchNormalization(momentum=0.8))
model.add(LeakyReLU(alpha=0.2))
model.add(Dropout(0.25))
model.add(Flatten())
model.add(Dense(1, activation='sigmoid'))
model.summary()
img = Input(shape=self.img_shape)
validity = model(img)
return Model(img, validity)
def train(self, epochs, batch_size=128, save_interval=50):
# Load the dataset
# (X_train, _), (_, _) = mnist.load_data()
X_train = polydata
# Rescale -1 to 1
# X_train = X_train / 127.5 - 1.
X_train = np.expand_dims(X_train, axis=3)
# Adversarial ground truths
valid = np.ones((batch_size, 1))
fake = np.zeros((batch_size, 1))
for epoch in range(epochs):
# ---------------------
# Train Discriminator
# ---------------------
# Select a random half of images
idx = np.random.randint(0, X_train.shape[0], batch_size)
imgs = X_train[idx]
# Sample noise and generate a batch of new images
noise = np.random.normal(0, 1, (batch_size, self.latent_dim))
gen_imgs = self.generator.predict(noise)
# Train the discriminator (real classified as ones and generated as zeros)
d_loss_real = self.discriminator.train_on_batch(imgs, valid)
d_loss_fake = self.discriminator.train_on_batch(gen_imgs, fake)
d_loss = 0.5 * np.add(d_loss_real, d_loss_fake)
# ---------------------
# Train Generator
# ---------------------
# Train the generator (wants discriminator to mistake images as real)
g_loss = self.combined.train_on_batch(noise, valid)
# Plot the progress
print ("%d [D loss: %f, acc.: %.2f%%] [G loss: %f]" % (epoch, d_loss[0], 100*d_loss[1], g_loss))
# If at save interval => save generated image samples
if epoch % save_interval == 0:
self.save_imgs(epoch)
def save_imgs(self, epoch):
r, c = 5, 5
noise = np.random.normal(0, 1, (r * c, self.latent_dim))
gen_imgs = self.generator.predict(noise)
# Rescale images 0 - 1
gen_imgs = 0.5 * gen_imgs + 0.5
fig, axs = plt.subplots(r, c)
cnt = 0
for i in range(r):
for j in range(c):
axs[i,j].imshow(gen_imgs[cnt, :,:,0], cmap='gray')
axs[i,j].axis('off')
cnt += 1
fig.savefig("images/poly_%d.png" % epoch)
plt.close()
if __name__ == '__main__':
dcgan = DCGAN()
dcgan.train(epochs=4000, batch_size=32, save_interval=50)
# + colab={"base_uri": "https://localhost:8080/", "height": 607} colab_type="code" id="guZ22DU3rbRo" outputId="8985c31d-d627-4d98-c443-28c85682c131"
fig, axes = plt.subplots(2, 8, figsize=(19,5))
for i in range(2):
for j in range(8):
out = dcgan.generator.predict(np.reshape(np.random.normal(0,1,2),(1,2)))
axes[i,j].imshow(out[0,:,:,0])
fig, axes = plt.subplots(2, 8, figsize=(19,5))
for i in range(2):
for j in range(8):
axes[i,j].imshow(polydata[i*j+j])
# + [markdown] colab_type="text" id="luLhXLGlhElx"
# #### Wasserstein GAN
# [Paper](https://arxiv.org/pdf/1701.07875.pdf)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="s6bNbndrG9jR" outputId="207e843f-f261-478f-b944-fee92eb7612c"
from tensorflow.keras import backend as K
class WGAN():
def __init__(self):
self.img_rows = 40
self.img_cols = 40
self.channels = 1
self.img_shape = (self.img_rows, self.img_cols, self.channels)
self.latent_dim = 10
# Following parameter and optimizer set as recommended in paper
self.n_critic = 10
self.clip_value = 0.03
optimizer = RMSprop(lr=0.00003)
# Build and compile the critic
self.critic = self.build_critic()
self.critic.compile(loss=self.wasserstein_loss,
optimizer=optimizer,
metrics=['accuracy'])
# Build the generator
self.generator = self.build_generator()
# The generator takes noise as input and generated imgs
z = Input(shape=(self.latent_dim,))
img = self.generator(z)
# For the combined model we will only train the generator
self.critic.trainable = False
# The critic takes generated images as input and determines validity
valid = self.critic(img)
# The combined model (stacked generator and critic)
self.combined = Model(z, valid)
self.combined.compile(loss=self.wasserstein_loss,
optimizer=optimizer)
def wasserstein_loss(self, y_true, y_pred):
return K.mean(y_true * y_pred)
def build_generator(self):
model = Sequential()
model.add(Dense(128 * 10 * 10, activation="relu", input_dim=self.latent_dim))
model.add(Reshape((10, 10, 128)))
model.add(UpSampling2D())
model.add(Conv2D(128, kernel_size=4, padding="same"))
# model.add(BatchNormalization(momentum=0.8))
model.add(Activation("relu"))
model.add(UpSampling2D())
model.add(Conv2D(64, kernel_size=4, padding="same"))
# model.add(BatchNormalization(momentum=0.8))
model.add(Activation("relu"))
model.add(Conv2D(self.channels, kernel_size=4, padding="same"))
model.add(Activation("tanh"))
model.summary()
noise = Input(shape=(self.latent_dim,))
img = model(noise)
return Model(noise, img)
def build_critic(self):
model = Sequential()
x=8
model.add(Conv2D(x*16, kernel_size=3, strides=2, input_shape=self.img_shape, padding="same"))
model.add(LeakyReLU(alpha=0.2))
model.add(Dropout(0.25))
model.add(Conv2D(x*32, kernel_size=3, strides=2, padding="same"))
model.add(ZeroPadding2D(padding=((0,1),(0,1))))
model.add(BatchNormalization(momentum=0.8))
model.add(LeakyReLU(alpha=0.2))
model.add(Dropout(0.25))
model.add(Conv2D(x*64, kernel_size=3, strides=2, padding="same"))
model.add(BatchNormalization(momentum=0.8))
model.add(LeakyReLU(alpha=0.2))
model.add(Dropout(0.25))
model.add(Conv2D(x*128, kernel_size=3, strides=1, padding="same"))
model.add(BatchNormalization(momentum=0.8))
model.add(LeakyReLU(alpha=0.2))
model.add(Dropout(0.25))
model.add(Flatten())
model.add(Dense(1))
model.summary()
img = Input(shape=self.img_shape)
validity = model(img)
return Model(img, validity)
def train(self, epochs, batch_size=128, sample_interval=50):
# Load the dataset
# (X_train, _), (_, _) = mnist.load_data()
X_train = polydata
# Rescale -1 to 1
# X_train = (X_train.astype(np.float32) - 127.5) / 127.5
X_train = np.expand_dims(X_train, axis=3)
# Adversarial ground truths
valid = np.ones((batch_size, 1))
fake = -np.ones((batch_size, 1))
for epoch in range(epochs):
self.critic.trainable = True
for _ in range(self.n_critic):
# ---------------------
# Train Discriminator
# ---------------------
# Select a random batch of images
idx = np.random.randint(0, X_train.shape[0], batch_size)
imgs = X_train[idx]
# Sample noise as generator input
noise = np.random.normal(0, 1, (batch_size, self.latent_dim))
# Generate a batch of new images
gen_imgs = self.generator.predict(noise)
# Train the critic
d_loss_real = self.critic.train_on_batch(imgs, valid)
d_loss_fake = self.critic.train_on_batch(gen_imgs, fake)
d_loss = 0.5 * np.add(d_loss_fake, d_loss_real)
# Clip critic weights
for l in self.critic.layers:
weights = l.get_weights()
weights = [np.clip(w, -self.clip_value, self.clip_value) for w in weights]
l.set_weights(weights)
# ---------------------
# Train Generator
# ---------------------
#self.critic.trainable = False
g_loss = self.combined.train_on_batch(noise, valid)
# Plot the progress
#print(epoch)
print ("%d [D loss: %f] [G loss: %f]" % (epoch, 1 - d_loss[0], 1 - g_loss))
# If at save interval => save generated image samples
if epoch % sample_interval == 0:
self.sample_images(epoch)
def sample_images(self, epoch):
r, c = 5, 5
noise = np.random.normal(0, 1, (r * c, self.latent_dim))
gen_imgs = self.generator.predict(noise)
# Rescale images 0 - 1
gen_imgs = 0.5 * gen_imgs + 0.5
fig, axs = plt.subplots(r, c)
cnt = 0
for i in range(r):
for j in range(c):
axs[i,j].imshow(gen_imgs[cnt, :,:,0], cmap='gray')
axs[i,j].axis('off')
cnt += 1
fig.savefig("images/wgan_poly_%d.png" % epoch)
plt.close()
if __name__ == '__main__':
wgan = WGAN()
wgan.train(epochs=4000, batch_size=32, sample_interval=50)
# + colab={"base_uri": "https://localhost:8080/", "height": 607} colab_type="code" id="1fF8hLgfiuV9" outputId="2e256503-e43d-43fc-f6e8-521bafc32731"
fig, axes = plt.subplots(2, 8, figsize=(19,5))
for i in range(2):
for j in range(8):
out = wgan.generator.predict(np.reshape(np.random.normal(0,1,10),(1,10)))
axes[i,j].imshow(out[0,:,:,0])
fig, axes = plt.subplots(2, 8, figsize=(19,5))
for i in range(2):
for j in range(8):
axes[i,j].imshow(polydata[i*j+j])
# + colab={} colab_type="code" id="3l2L3-YX5U28"
|
09/.ipynb_checkpoints/SolutionExercise9-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Project 3
#
#
# # Movie Genre Classification
#
# Classify a movie genre based on its plot.
#
# <img src="moviegenre.png"
# style="float: left; margin-right: 10px;" />
#
#
#
#
# https://www.kaggle.com/c/miia4201-202019-p3-moviegenreclassification/overview
#
# ### Data
#
# Input:
# - movie plot
#
# Output:
# Probability of the movie belong to each genre
#
#
# ### Evaluation
#
# - 20% API
# - 30% Create a solution using with a Machine Learning algorithm - Presentation (5 slides)
# - 50% Performance in the Kaggle competition (Normalized acording to class performance in the private leaderboard)
#
#
# ### Acknowledgements
#
# We thank Professor <NAME>, Ph.D. and his student <NAME> for providing this dataset.
#
# See https://arxiv.org/abs/1702.01992
# ## Sample Submission
import pandas as pd
import os
import numpy as np
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.preprocessing import MultiLabelBinarizer
from sklearn.multiclass import OneVsRestClassifier
from sklearn.ensemble import RandomForestRegressor, RandomForestClassifier
from sklearn.metrics import r2_score, roc_auc_score
from sklearn.model_selection import train_test_split
dataTraining = pd.read_csv('https://github.com/albahnsen/AdvancedMethodsDataAnalysisClass/raw/master/datasets/dataTraining.zip', encoding='UTF-8', index_col=0)
dataTesting = pd.read_csv('https://github.com/albahnsen/AdvancedMethodsDataAnalysisClass/raw/master/datasets/dataTesting.zip', encoding='UTF-8', index_col=0)
dataTraining.head()
dataTesting.head()
# ### Create count vectorizer
#
vect = CountVectorizer(max_features=1000)
X_dtm = vect.fit_transform(dataTraining['plot'])
X_dtm.shape
print(vect.get_feature_names()[:50])
# ### Create y
# +
dataTraining['genres'] = dataTraining['genres'].map(lambda x: eval(x))
le = MultiLabelBinarizer()
y_genres = le.fit_transform(dataTraining['genres'])
# -
y_genres
X_train, X_test, y_train_genres, y_test_genres = train_test_split(X_dtm, y_genres, test_size=0.33, random_state=42)
# ### Train multi-class multi-label model
clf = OneVsRestClassifier(RandomForestClassifier(n_jobs=-1, n_estimators=100, max_depth=10, random_state=42))
clf.fit(X_train, y_train_genres)
y_pred_genres = clf.predict_proba(X_test)
roc_auc_score(y_test_genres, y_pred_genres, average='macro')
# ### Predict the testing dataset
# +
X_test_dtm = vect.transform(dataTesting['plot'])
cols = ['p_Action', 'p_Adventure', 'p_Animation', 'p_Biography', 'p_Comedy', 'p_Crime', 'p_Documentary', 'p_Drama', 'p_Family',
'p_Fantasy', 'p_Film-Noir', 'p_History', 'p_Horror', 'p_Music', 'p_Musical', 'p_Mystery', 'p_News', 'p_Romance',
'p_Sci-Fi', 'p_Short', 'p_Sport', 'p_Thriller', 'p_War', 'p_Western']
y_pred_test_genres = clf.predict_proba(X_test_dtm)
# -
res = pd.DataFrame(y_pred_test_genres, index=dataTesting.index, columns=cols)
res.head()
res.to_csv('pred_genres_text_RF.csv', index_label='ID')
|
Exercises/P3-MovieGenrePrediction.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] heading_collapsed=true
# # EDA of processed openFDA drug event data
# + hidden=true
import os
import numpy as np
import pandas as pd
import pickle
import matplotlib.pyplot as plt
import seaborn as sns
sns.set_style('white')
# %matplotlib inline
data_dir = "../../data/openFDA_drug_event/"
# + [markdown] heading_collapsed=true hidden=true
# ## Standard mappings
# + [markdown] heading_collapsed=true hidden=true
# ### drugs
# + hidden=true
drugs = (pd.
read_csv('../../data/openFDA_drug_event/er_tables/drugs.csv.gz',
compression='gzip',index_col=0))
# + hidden=true
print(drugs.shape)
drugs.head()
# + hidden=true
drugs_reports = drugs.safetyreportid.astype(str).unique()
# + hidden=true
print(len(drugs_reports))
drugs_reports[:5]
# + hidden=true
drugs_rxcuis = drugs.rxcui.astype(int).unique()
# + hidden=true
print(len(drugs_rxcuis))
drugs_rxcuis[:5]
# + hidden=true
del drugs
# + hidden=true
standard_drugs = (pd.
read_csv('../../data/openFDA_drug_event/er_tables/standard_drugs.csv.gz',
compression='gzip',
index_col=0,
dtype={
'safetyreportid' : 'str'
}))
standard_drugs.head()
# + hidden=true
print(standard_drugs.shape)
# + hidden=true
standard_drugs_reports = standard_drugs.safetyreportid.astype(str).unique()
# + hidden=true
print(len(standard_drugs_reports))
# + hidden=true
print(standard_drugs.RxNorm_concept_id.nunique())
print(standard_drugs.RxNorm_concept_code.nunique())
# + hidden=true
standard_drugs_rxids = standard_drugs.RxNorm_concept_id.dropna().astype(int).unique()
standard_drugs_rxcuis = standard_drugs.RxNorm_concept_code.dropna().astype(int).unique()
# + hidden=true
del standard_drugs
# + hidden=true
print(len(standard_drugs_reports)/len(drugs_reports))
# + hidden=true
print(len(np.intersect1d(standard_drugs_rxcuis,drugs_rxcuis))/len(drugs_rxcuis))
print(len(np.intersect1d(standard_drugs_rxcuis,drugs_rxcuis))/len(standard_drugs_rxcuis))
# + hidden=true
standard_drugs_atc = (pd.
read_csv('../../data/openFDA_drug_event/er_tables/standard_drugs_atc.csv.gzip',
compression='gzip',index_col=0))
print(standard_drugs_atc.shape)
print(standard_drugs_atc.head())
tmp = standard_drugs_atc[['RxNorm_concept_id','ATC_concept_id']].drop_duplicates()
print(tmp.shape)
tmp = tmp[tmp.RxNorm_concept_id.notnull()]
print(tmp.shape)
print(tmp.shape)
print(tmp.dropna().shape)
# + hidden=true
standard_drugs_atc_reports = standard_drugs_atc.safetyreportid.astype(str).unique()
# + hidden=true
del standard_drugs_atc
# + hidden=true
standard_drugs_rxnorm_ingredients = (pd.
read_csv('../../data/openFDA_drug_event/er_tables/standard_drug_rxnorm_ingredients.csv.gzip',
compression='gzip',index_col=0))
# + hidden=true
standard_drugs_rxnorm_ingredients.head()
# + hidden=true
standard_drugs_rxnorm_ingredients_reports = (standard_drugs_rxnorm_ingredients.
safetyreportid.astype(str).unique()
)
# + hidden=true
print(len(standard_drugs_rxnorm_ingredients_reports))
print(len(drugs_reports))
print(len(np.intersect1d(standard_drugs_rxnorm_ingredients_reports,drugs_reports))/ \
len(drugs_reports))
# + hidden=true
del standard_drugs_rxnorm_ingredients
# + [markdown] heading_collapsed=true hidden=true
# ### reactions
# + hidden=true
reactions = (pd.
read_csv('../../data/openFDA_drug_event/er_tables/reactions.csv.gz',
compression='gzip',
index_col=0,
dtype={
'safetyreportid' : 'str',
}))
reactions.head()
# + hidden=true
reactions_meddraptnames = reactions.reaction_meddrapt.dropna().astype(str).str.title().unique()
print(len(reactions_meddraptnames))
reactions_meddraptnames[:5]
# + hidden=true
reactions_reports = reactions.safetyreportid.unique()
# + hidden=true
print(len(reactions_reports))
# + hidden=true
del reactions
# + hidden=true
standard_reactions = (pd.
read_csv('../../data/openFDA_drug_event/er_tables/standard_reactions.csv.gz',
compression='gzip',
index_col=0,
dtype={
'safetyreportid' : 'str',
'MedDRA_concept_id' : 'int',
'MedDRA_concept_code' : 'int'
}))
standard_reactions.head()
# + hidden=true
standard_reactions.MedDRA_concept_class_id.value_counts()
# + hidden=true
standard_reactions_reports = standard_reactions.safetyreportid.unique()
# + hidden=true
len(standard_reactions_reports)
# + hidden=true
len(standard_reactions_reports)/len(reactions_reports)
# + hidden=true
standard_reactions_meddraptids = standard_reactions.MedDRA_concept_id.unique()
standard_reactions_meddraptnames = standard_reactions.MedDRA_concept_name.unique()
print(len(standard_reactions_meddraptnames))
standard_reactions_meddraptnames[:5]
# + hidden=true
del standard_reactions
# + hidden=true
reactions_meddraptnames
# + hidden=true
print(len(np.intersect1d(standard_reactions_meddraptnames,reactions_meddraptnames))/ \
len(reactions_meddraptnames))
print(len(np.intersect1d(standard_reactions_meddraptnames,reactions_meddraptnames))/ \
len(standard_reactions_meddraptnames))
# + hidden=true
standard_reactions_meddra_hlt = (pd.
read_csv('../../data/openFDA_drug_event/er_tables/standard_reactions_meddra_hlt.csv.gz',
compression='gzip',index_col=0,
dtype={
'safetyreportid' : 'str',
'MedDRA_concept_id' : 'int',
'MedDRA_concept_code' : 'int'
}))
standard_reactions_meddra_hlt.head()
# + hidden=true
standard_reactions_meddra_hlt_reports = standard_reactions_meddra_hlt.safetyreportid.unique()
# + hidden=true
print(len(standard_reactions_meddra_hlt_reports))
# + hidden=true
len(standard_reactions_meddra_hlt_reports)/len(reactions_reports)
# + hidden=true
del standard_reactions_meddra_hlt
# + hidden=true
standard_reactions_meddra_hlgt = (pd.
read_csv('../../data/openFDA_drug_event/er_tables/standard_reactions_meddra_hlgt.csv.gz',
compression='gzip',index_col=0,
dtype={
'safetyreportid' : 'str',
'MedDRA_concept_id' : 'int',
'MedDRA_concept_code' : 'int'
}))
standard_reactions_meddra_hlgt.head()
# + hidden=true
standard_reactions_meddra_hlgt_reports = standard_reactions_meddra_hlgt.safetyreportid.unique()
# + hidden=true
print(len(standard_reactions_meddra_hlgt_reports))
# + hidden=true
len(standard_reactions_meddra_hlgt_reports)/len(reactions_reports)
# + hidden=true
del standard_reactions_meddra_hlgt
# + hidden=true
standard_reactions_meddra_soc = (pd.
read_csv('../../data/openFDA_drug_event/er_tables/standard_reactions_meddra_soc.csv.gz',
compression='gzip',index_col=0,
dtype={
'safetyreportid' : 'str',
'MedDRA_concept_id' : 'int',
'MedDRA_concept_code' : 'int'
}
))
standard_reactions_meddra_soc.head()
# + hidden=true
standard_reactions_meddra_soc_reports = standard_reactions_meddra_soc.safetyreportid.unique()
# + hidden=true
print(len(standard_reactions_meddra_soc_reports))
# + hidden=true
len(standard_reactions_meddra_soc_reports)/len(reactions_reports)
# + hidden=true
del standard_reactions_meddra_soc
# + hidden=true
standard_reactions_snomed = (pd.
read_csv('../../data/openFDA_drug_event/er_tables/'+
'standard_reactions_snomed.csv.gz',
compression='gzip',index_col=0,
dtype={
'safetyreportid' : 'str',
'SNOMED_concept_id' : 'int',
'SNOMED_concept_code' : 'int'
}
))
standard_reactions_snomed.head()
# + hidden=true
standard_reactions_snomed_reports = (standard_reactions_snomed.safetyreportid.unique())
len(standard_reactions_snomed_reports)
# + hidden=true
len(np.intersect1d(standard_reactions_snomed_reports,reactions_reports))/ \
len(reactions_reports)
# + hidden=true
del standard_reactions_meddrapt_to_snomed
# + [markdown] heading_collapsed=true hidden=true
# ## Number of reports over the years
# + hidden=true
report = (pd.
read_csv('../../data/openFDA_drug_event/er_tables/report.csv.gzip',
compression='gzip',index_col=0))
report.head()
# + hidden=true
report.receive_date = pd.to_datetime(report.receive_date,format='%Y%m%d')
report['Year'] = report.receive_date.apply(lambda x : x.year)
# + hidden=true
import matplotlib.dates as mdates
# + hidden=true
nreportoverdates = (report.
groupby('Year').
safetyreportid.
count()
)
nreportoverdates.head()
fig,ax = plt.subplots(dpi=200)
nreportoverdates.plot(kind='bar',ax=ax)
ax.set_yscale('log')
ax.set_xlabel('')
# + hidden=true
patient = (pd.
read_csv('../../data/openFDA_drug_event/er_tables/patient.csv.gzip',
compression='gzip',index_col=0))
patient.head()
# + hidden=true
ped_reports = (patient.
query('patient_custom_master_age>0 & patient_custom_master_age<18').
safetyreportid.unique()
)
# + hidden=true
npedreportoverdates = (report.
query('safetyreportid in @ped_reports').
groupby('Year').
safetyreportid.
count()
)
fig,ax = plt.subplots(dpi=200)
npedreportoverdates.plot(kind='bar',ax=ax)
ax.set_yscale('log')
ax.set_xlabel('')
# + [markdown] heading_collapsed=true hidden=true
# ## Age distribution
# + hidden=true
patient_df = (pd.
read_csv('../../data/openFDA_drug_event/er_tables/patient.csv.gzip',
compression="gzip",index_col=0))
patient_df.head()
# + hidden=true
col = 'patient_custom_master_age'
display(patient_df[col].astype(float).dropna().shape[0])
# + hidden=true
values = patient_df.groupby('safetyreportid')[col].agg('mean').dropna().values
# + hidden=true
print(len(values))
print(len(values)/patient_df.shape[0])
# + hidden=true
gt_100 = values>100
sum(gt_100)
# + hidden=true
gt_115 = values>115
sum(gt_115)
# + hidden=true
lt_0 = values<0
sum(lt_0)
# + hidden=true
eq_0 = values==0
sum(eq_0)
# + hidden=true
eq_100 = values==100
sum(eq_100)
# + hidden=true
lt_100 = values<100
gt_0 = values>0
num = sum(np.logical_and(gt_0,lt_100))
print(num)
num/len(values)
# + hidden=true
lt_18 = values<18
num = sum(np.logical_and(gt_0,lt_18))
print(num)
num/len(values)
# + hidden=true
gtoeq_18 = values>=18
num = sum(np.logical_and(gtoeq_18,lt_100))
print(num)
num/len(values)
# + hidden=true
sensical_values = values[np.logical_and(gt_0,lt_100)]
# + hidden=true
integer_year_counts = np.unique(np.floor(sensical_values),return_counts=True)
integer_year_counts
# + hidden=true
plt.figure(dpi=200)
plt.plot(integer_year_counts[0],integer_year_counts[1])
# + hidden=true
plt.figure(dpi=200)
plt.bar(integer_year_counts[0][:19],integer_year_counts[1][:19])
plt.xticks(integer_year_counts[0][:19])
# + [markdown] heading_collapsed=true hidden=true
# ## Age units used across childhood
# + hidden=true
pediatric_patient_df = (patient_df.
query('patient_custom_master_age>0 & patient_custom_master_age<18'))
df = pediatric_patient_df[['patient_onsetageunit','patient_custom_master_age']].query('patient_custom_master_age<18').copy()
df['patient_custom_master_age'] = np.floor(df['patient_custom_master_age'])
df['mem'] = 1.
df_pivot = df.pivot_table(index='patient_custom_master_age',columns='patient_onsetageunit',
values='mem',aggfunc=sum,fill_value=0)
order=['Decade','Year','Month','Week','Day','Hour']
df_pivot = df_pivot[order].apply(lambda x : x / sum(x),axis=1).round(2)
display(df_pivot.T)
fig,ax=plt.subplots(dpi=200)
g = df_pivot.plot(kind='bar',stacked=True,ax=ax)
g.legend(bbox_to_anchor=(1,1))
g = g.set_xticklabels(g.get_xticklabels(),rotation=30)
# + [markdown] heading_collapsed=true hidden=true
# ## NDrugindications across childhood
# + hidden=true
drugcharacteristics = (pd.
read_csv('../../data/openFDA_drug_event/er_tables/drugcharacteristics.csv.gzip',
compression='gzip',
index_col=0)
)
drugcharacteristics.head()
# + hidden=true
patients = (pd.
read_csv('../../data/openFDA_drug_event/er_tables/patient.csv.gzip',
compression='gzip',
index_col=0)
)
patients.head()
# + hidden=true
patient_drugcharacteristics = (patients.
set_index('safetyreportid').
join(drugcharacteristics.
set_index('safetyreportid')
)
)
patient_drugcharacteristics.head()
# + hidden=true
pediatric_patient_df = (patient_drugcharacteristics.
loc[:,['patient_custom_master_age','drug_indication']].
dropna().
query('patient_custom_master_age>0 & patient_custom_master_age<25').
reset_index()
)
pediatric_patient_df['year'] = np.floor(pediatric_patient_df.patient_custom_master_age).astype(int)
# + hidden=true
fig,ax = plt.subplots(dpi=200)
(pediatric_patient_df.
loc[:,['year','drug_indication']].
drop_duplicates().
groupby('year')['drug_indication'].
count()
).plot.barh(ax=ax)
ax.set_xlabel('Number of Drug Indications')
ax.set_ylabel('Age')
# + [markdown] heading_collapsed=true hidden=true
# ## NDrugs across childhood
# + hidden=true
pediatric_patients = (pd.
read_csv('../../data/openFDA_drug_event/er_tables/patient.csv.gzip',
compression='gzip',
index_col=0)
).query('patient_custom_master_age<25')
pediatric_patients.head()
# + hidden=true
ped_reports = pediatric_patients.safetyreportid.astype(str).unique()
len(ped_reports)
# + hidden=true
pediatric_standard_drugs = (pd.
read_csv('../../data/openFDA_drug_event/er_tables/standard_drugs.csv.gzip',
compression='gzip',
index_col=0)
).query('safetyreportid in @ped_reports')
pediatric_standard_drugs.safetyreportid = pediatric_standard_drugs.safetyreportid.astype(str)
pediatric_standard_drugs.RxNorm_concept_id = pediatric_standard_drugs.RxNorm_concept_id.astype(int)
pediatric_standard_drugs.head()
# + hidden=true
pediatric_patient_drugs = (pediatric_patients.
loc[:,['patient_custom_master_age','safetyreportid']].
drop_duplicates().
set_index('safetyreportid').
join(pediatric_standard_drugs.
set_index('safetyreportid')
).
dropna()
)
pediatric_patient_drugs.RxNorm_concept_id = pediatric_patient_drugs.RxNorm_concept_id.astype(int)
pediatric_patient_drugs.RxNorm_concept_code = pediatric_patient_drugs.RxNorm_concept_code.astype(int)
pediatric_patient_drugs['year'] = np.floor(pediatric_patient_drugs.patient_custom_master_age).astype(int)
pediatric_patient_drugs.head()
# + hidden=true
fig,ax = plt.subplots(dpi=200)
(pediatric_patient_drugs.
loc[:,['year','RxNorm_concept_id']].
drop_duplicates().
groupby('year')['RxNorm_concept_id'].
count()
).plot.barh(ax=ax)
ax.set_xlabel('Number of Drugs')
ax.set_ylabel('Age')
# + hidden=true
del pediatric_patient_drugs
del pediatric_standard_drugs
del pediatric_patients
# + [markdown] heading_collapsed=true hidden=true
# ## NReactions across Childhood
# + hidden=true
pediatric_patients = (pd.
read_csv('../../data/openFDA_drug_event/er_tables/patient.csv.gzip',
compression='gzip',
index_col=0)
).query('patient_custom_master_age<25')
pediatric_patients.head()
# + hidden=true
ped_reports = pediatric_patients.safetyreportid.astype(str).unique()
len(ped_reports)
# + hidden=true
pediatric_standard_reactions = (pd.
read_csv('../../data/openFDA_drug_event/er_tables/standard_reactions.csv.gzip',
compression='gzip',
index_col=0)
).query('safetyreportid in @ped_reports')
pediatric_standard_reactions.safetyreportid = pediatric_standard_reactions.safetyreportid.astype(str)
pediatric_standard_reactions.MedDRA_concept_id = pediatric_standard_reactions.MedDRA_concept_id.astype(int)
pediatric_standard_reactions.head()
# + hidden=true
pediatric_patient_reactions = (pediatric_patients.
loc[:,['patient_custom_master_age','safetyreportid']].
drop_duplicates().
set_index('safetyreportid').
join(pediatric_standard_reactions.
set_index('safetyreportid')
).
dropna()
)
pediatric_patient_reactions.MedDRA_concept_id = pediatric_patient_reactions.MedDRA_concept_id.astype(int)
pediatric_patient_reactions.MedDRA_concept_code = pediatric_patient_reactions.MedDRA_concept_code.astype(int)
pediatric_patient_reactions['year'] = np.floor(pediatric_patient_reactions.patient_custom_master_age).astype(int)
pediatric_patient_reactions.head()
# + hidden=true
fig,ax = plt.subplots(dpi=200)
(pediatric_patient_reactions.
loc[:,['year','MedDRA_concept_id']].
drop_duplicates().
groupby('year')['MedDRA_concept_id'].
count()
).plot.barh(ax=ax)
ax.set_xlabel('Number of Reactions')
ax.set_ylabel('Age')
# + hidden=true
del pediatric_patient_drugs
del pediatric_standard_drugs
del pediatric_patients
# + hidden=true
# + [markdown] heading_collapsed=true hidden=true
# ## Medicinal product frequency across childhood
# + hidden=true
df = (pediatric_patient_df.
set_index('safetyreportid').
join(
pediatric_patient_drug_df.
set_index('safetyreportid')
).
loc[:,['master_age','medicinalproduct']].
query('master_age<18')
)
df['master_age'] = np.floor(df['master_age'])
df['mem'] = 1.
medproduct_freq_in_childhood = df.pivot_table(index='master_age',columns='medicinalproduct',
values='mem',aggfunc=sum,fill_value=0)
cols = medproduct_freq_in_childhood.columns[medproduct_freq_in_childhood.sum(axis=0)>medproduct_freq_in_childhood.shape[0]]
display(medproduct_freq_in_childhood[cols])
# + [markdown] heading_collapsed=true hidden=true
# ## Drug administration route distribution
# + hidden=true
patient_drug_df = (pd.read_csv('../../data/openFDA_drug_event/er_tables/drugcharacteristics.csv.gzip',
compression='gzip',index_col=0))
# + hidden=true
patient_drug_df.head()
# + hidden=true
drug_route_counts = np.unique(patient_drug_df['drug_administration'].dropna().values,return_counts=True)
# + hidden=true
print(drug_route_counts)
len(drug_route_counts[0])
# + hidden=true
plt.figure(figsize=(5,25),dpi=200)
order = drug_route_counts[1].argsort()
plt.barh(drug_route_counts[0][order],drug_route_counts[1][order])
plt.xscale('log')
# + hidden=true
|
ntbks/openFDA_drug_event_parsing/Exploring.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
"""
1. Standard deviation.
A number that describes how spread out the values are.
- A low standard deviation means that most of the numbers are close to the mean (average) value.
- A high standard deviation means that the values are spread out over a wider range.
"""
import numpy
# +
# LOW std deviation
speed = [86,87,88,86,87,85,86]
low_std = numpy.std(speed)
print(low_std)
# +
# HIGH std deviation
speed = [32,111,138,28,59,77,97]
high_std = numpy.std(speed)
print(high_std)
# -
"""
2. Variance.
Variance is another number that indicates how spread out the values are.
FORMULA: standard deviation = 'square root' of the variance.
Or
Variance = (standard deviation x standard deviation)
EXAMPLE.
To calculate the variance you have to do as follows:
1. Find the mean:
(32+111+138+28+59+77+97) / 7 = 77.4
2. For each value: find the difference from the mean:
32 - 77.4 = -45.4
111 - 77.4 = 33.6
138 - 77.4 = 60.6
28 - 77.4 = -49.4
59 - 77.4 = -18.4
77 - 77.4 = - 0.4
97 - 77.4 = 19.6
3. For each difference: find the square value:
(-45.4)2 = 2061.16
(33.6)2 = 1128.96
(60.6)2 = 3672.36
(-49.4)2 = 2440.36
(-18.4)2 = 338.56
(- 0.4)2 = 0.16
(19.6)2 = 384.16
4. The variance is the average number of these squared differences:
(2061.16+1128.96+3672.36+2440.36+338.56+0.16+384.16) / 7 = 1432.2
"""
# +
import numpy
speed = [32,111,138,28,59,77,97]
variance = numpy.var(speed)
print(variance)
# +
# std dev
import numpy
speed = [32,111,138,28,59,77,97]
std_dev = numpy.std(speed)
print(std_dev)
|
0.ML-Basics-to-pro-Python-Lab/.ipynb_checkpoints/2.StandardDeviation-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/samirlenka/pandas_exercises/blob/master/04_Apply/US_Crime_Rates/Exercises.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="k8KTkBjfUije"
# # United States - Crime Rates - 1960 - 2014
# + [markdown] id="y7XzbR66Uijn"
# ### Introduction:
#
# This time you will create a data
#
# Special thanks to: https://github.com/justmarkham for sharing the dataset and materials.
#
# ### Step 1. Import the necessary libraries
# + id="USXE4OW9Uijo"
import pandas as pd
import numpy as np
# + [markdown] id="Pmk7PbEuUijp"
# ### Step 2. Import the dataset from this [address](https://raw.githubusercontent.com/guipsamora/pandas_exercises/master/04_Apply/US_Crime_Rates/US_Crime_Rates_1960_2014.csv).
# + [markdown] id="AvPlw-CnUijp"
# ### Step 3. Assign it to a variable called crime.
# + id="UAxnY4k0Uijq" outputId="8366718f-289c-46c6-f290-9c9a3ed7b0ea" colab={"base_uri": "https://localhost:8080/", "height": 204}
crime = pd.read_csv('https://raw.githubusercontent.com/guipsamora/pandas_exercises/master/04_Apply/US_Crime_Rates/US_Crime_Rates_1960_2014.csv')
crime.head(5)
# + [markdown] id="HgcDUGY0Uijq"
# ### Step 4. What is the type of the columns?
# + id="gzLINEybUijq" outputId="f2630f28-bdec-47b2-9877-330b85ec7551" colab={"base_uri": "https://localhost:8080/"}
crime.info()
# + [markdown] id="PKwdCiQjUijr"
# ##### Have you noticed that the type of Year is int64. But pandas has a different type to work with Time Series. Let's see it now.
#
# ### Step 5. Convert the type of the column Year to datetime64
# + id="AyNpEeQlUijr" outputId="f90b393e-049c-4351-81c1-615d696595fa" colab={"base_uri": "https://localhost:8080/"}
crime.Year = pd.to_datetime(crime.Year, format='%Y')
#crime.head(10)
crime.info()
# + [markdown] id="BFbmpyyhUijr"
# ### Step 6. Set the Year column as the index of the dataframe
# + id="Oscz82o_Uijs" outputId="414377f4-2bda-419d-f228-33e3022c790e" colab={"base_uri": "https://localhost:8080/", "height": 235}
crime = crime.set_index('Year', drop = True)
crime.head(5)
# + [markdown] id="bmlsleFfUijs"
# ### Step 7. Delete the Total column
# + id="qF8PQPviUijs" outputId="47019080-3667-43d1-f7ca-ee5259e52e8e" colab={"base_uri": "https://localhost:8080/", "height": 235}
del crime['Total']
crime.head(5)
# + [markdown] id="t9zI-Db-Uijt"
# ### Step 8. Group the year by decades and sum the values
#
# #### Pay attention to the Population column number, summing this column is a mistake
# + id="8RyGR8eKUijt" outputId="f63181c5-9eb7-48fe-fe53-e22d67843107" colab={"base_uri": "https://localhost:8080/", "height": 266}
crimes = crime.resample('10AS').sum()
crimes
# + id="JoLH2wNOTahS" outputId="17bdeadd-0ba7-4e2b-dfbb-becaa42a5db4" colab={"base_uri": "https://localhost:8080/", "height": 266}
population = crime['Population'].resample('10AS').max()
# Updating the "Population" column
crimes['Population'] = population
crimes.head(10)
# + [markdown] id="9DlmfX1gUijt"
# ### Step 9. What is the most dangerous decade to live in the US?
# + id="4QMM9tPcUiju" outputId="d9c57791-d2c7-4f19-da0f-05e1c39861da" colab={"base_uri": "https://localhost:8080/"}
# apparently the 90s was a pretty dangerous time in the US
crime.idxmax(axis=0)
crime.idxmax(0)
|
04_Apply/US_Crime_Rates/Exercises.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # WACV -- CityScapes
#
# ## 19 semantic classes
#
# ### NAS architectures based on MobileNet-v2 with 3 initial layers
# %matplotlib inline
import glob
import sys
sys.path.append('../../src/')
from functools import partial
# +
from PIL import Image
import cv2
import matplotlib.pyplot as plt
import numpy as np
import torch
import torch.nn as nn
import torch.nn.functional as F
# -
from nn.encoders import mbv2
from nn.micro_decoders import TemplateDecoder as Decoder
from utils.helpers import prepare_img
from utils.model_zoo import load_url
class Segmenter(nn.Module):
"""Create Segmenter"""
def __init__(self, encoder, decoder):
super(Segmenter, self).__init__()
self.encoder = encoder
self.decoder = decoder
def _reset_clf(self, num_classes):
self.decoder._reset_clf(num_classes)
def forward(self, x):
return self.decoder(self.encoder(x))
models_info = {
'arch0': (
[[[3, 0, 1], [4, 1, 1], [3, 1, 1]],
[[0, 1, 0, 0, 1],
[2, 1, 2, 1, 0],
[3, 1, 1, 1, 0],
[1, 1, 2, 0, 0],
[3, 0, 2, 0, 0],
[5, 3, 2, 1, 0],
[0, 5, 0, 1, 0]]],
'wacv_cs-2dcef44e.pth',
'https://cloudstor.aarnet.edu.au/plus/s/EhSSQUPeyKvL5Zk/download'
),
'arch1': (
[[[1, 1, 0], [1, 3, 0], [3, 4, 0]],
[[1, 1, 0, 0, 0],
[0, 1, 1, 1, 1],
[3, 1, 2, 3, 0],
[3, 0, 2, 2, 0],
[0, 1, 2, 0, 0],
[2, 1, 1, 3, 0],
[4, 0, 2, 2, 0]]],
'wacv_cs-2bcc2420.pth',
'https://cloudstor.aarnet.edu.au/plus/s/pnGC9DB03uLIXhW/download'
),
}
# Configuration
AGG_SIZE = 64
REPEATS = 2
NUM_CLASSES = 19
cmap = np.load('../../src/utils/cs_cmap.npy')
device = 'cuda' if torch.cuda.is_available() else 'cpu'
dec_fn = partial(Decoder,
num_classes=NUM_CLASSES,
agg_size=AGG_SIZE,
repeats=REPEATS)
img_dir = '../imgs/CityScapes/'
imgs = glob.glob('{}*leftImg8bit.png'.format(img_dir))
# +
# Initialise models
models = dict()
for arch, (config, filename, url) in models_info.items():
enc = mbv2(pretrained=False, return_layers=[1, 2])
dec = dec_fn(config=config, inp_sizes=enc.out_sizes)
segm = Segmenter(enc, dec).to(device).eval()
segm.load_state_dict(load_url((arch + '_' + filename, url), map_location=device), strict=False)
models[arch] = segm
del enc
# +
# Figure 10 from the paper
n_cols = len(models) + 2 # 1 - for image, 1 - for GT
n_rows = len(imgs)
plt.figure(figsize=(24, 12))
idx = 1
with torch.no_grad():
for img_path in imgs:
img = np.array(Image.open(img_path))
msk = np.array(Image.open(img_path.replace('leftImg8bit', 'gtFine_labelIds')))
orig_size = img.shape[:2][::-1]
img_inp = torch.tensor(
prepare_img(img).transpose(2, 0, 1)[None]).float().to(device)
plt.subplot(n_rows, n_cols, idx)
plt.imshow(img)
plt.title('img')
plt.axis('off')
idx += 1
plt.subplot(n_rows, n_cols, idx)
plt.imshow(msk)
plt.title('gt')
plt.axis('off')
idx += 1
for mname, mnet in models.items():
segm = mnet(img_inp)[0].squeeze().data.cpu().numpy().transpose((1, 2, 0))
segm = cv2.resize(segm, orig_size, interpolation=cv2.INTER_CUBIC)
segm = cmap[segm.argmax(axis=2).astype(np.uint8)]
plt.subplot(n_rows, n_cols, idx)
plt.imshow(segm)
plt.title(mname)
plt.axis('off')
idx += 1
# -
|
examples/inference/WACV-CS-segm.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from keras.models import Sequential
from keras.layers import Dense
from keras.layers import LSTM
from sklearn.preprocessing import MinMaxScaler
from sklearn.metrics import mean_squared_error
from sklearn.model_selection import train_test_split
# %config InlineBackend.figure_format='svg'
# -
df=pd.read_csv('train_Data.csv')
df.columns
df_min=df[df['8']==1]
df_min.to_csv('pima_minority_train.csv',index=False)
df_min=pd.read_csv('pima_minority_train.csv')
df_majority=df[df['8']==0]
df_majority.to_csv('pima_majority_train.csv',index=False)
df_majority=pd.read_csv('pima_majority_train.csv')
# %matplotlib inline
sns.countplot(x='8',data=df)
def create_dataset(dataset,look_back=1):
datax,datay=[],[]
for i in range(len(dataset)-look_back-1):
a=dataset[i:(i+look_back),:]
datax.append(a)
datay.append(dataset[i+look_back,:])
return np.array(datax),np.array(datay)
# +
df_minor=np.array(df_min)
scaler=MinMaxScaler(feature_range=(0,1))
df_minor=scaler.fit_transform(df_min)
x,y=create_dataset(df_minor,3)#5
print(x.shape)
print(y.shape)
# -
Xtrain,xtest,Ytrain,ytest=train_test_split(x,y,test_size=0.40,random_state=60)
# +
model=Sequential()
model.add(LSTM(20,input_shape=(Xtrain.shape[1],Xtrain.shape[2])))#5
model.add(Dense(9))
print(model.summary())
# -
model.compile(loss='mse',optimizer='adam')
history=model.fit(Xtrain,Ytrain,epochs=500,verbose=1)
model.save('7-24-2019-pima.h5')
Xtrain.shape
plt.plot(history.history['loss'],label='train')
#plt.plot(history.history['val_loss'],label='test')
plt.xlabel('number of epochs')
plt.ylabel('val_loss')
plt.legend()
#pyplot.savefig('LSTM training.png',dpi=300)
plt.show()
prediction=model.predict(xtest)
def draw_prediction(ytest,d,columns):
_,axes=plt.subplots(len(columns),1,figsize=(10,20))
for i,cols in enumerate(columns):
axes[i].plot(ytest[:,i],label='real',color='blue')
axes[i].plot(d[:,i],label='prediction',color='orange')
#axes[i].set_xlabel='index'
#axes[i].set_ylabel=cols
axes[i].xlabel='index'
axes[i].ylabel=cols
clmns=df.columns
draw_prediction(ytest,prediction,clmns)
prediction
prediction2=scaler.inverse_transform(prediction)
ytest2=scaler.inverse_transform(ytest)
draw_prediction(ytest2,prediction2,clmns)
prediction
new_data=pd.DataFrame(prediction2)
new_data.to_csv('new_corrected_data-v1-7-24-2019.csv',index=False)
|
pima- seed-19/sample generation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # `allcools mcds`
# + tags=["remove-cell"]
import subprocess
from IPython.display import display, Markdown
def execute_command_and_return_markdown(command):
output = subprocess.run(command,
shell=True,
check=True,
stdout=subprocess.PIPE,
encoding='utf8').stdout
markdown = f"""
```shell
$ {command}
{output}
```
"""
display(Markdown(markdown))
return
# + tags=["hide-input"]
execute_command_and_return_markdown('allcools generate-mcds -h')
# -
|
docs/allcools/command_line/allcools_mcds.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
matplotlib.style.use('seaborn')
# -
# Load data
df = pd.read_csv('../kryptos/performance_results/all_strategies/backtest_summary.csv', sep=',')
df.head()
df.columns
def show_bar_histogram(values, label):
labels = df.namespace.tolist()
ind = np.arange(len(labels))
width = 0.9
fig, ax = plt.subplots(figsize=(5,10))
rects = ax.barh(ind, np.array(values), color='y')
ax.set_yticks(ind+((width)/2.))
ax.set_yticklabels(labels, rotation='horizontal')
ax.set_xlabel(label)
plt.show()
print('Number of simulations: ' + str(df.head(1).number_of_simulations[0]))
print('Data Frequency: ' + str(df.head(1).data_freq[0]))
print('Asset: ' + str(df.head(1).asset[0]))
print('Exchange: ' + str(df.head(1).exchange[0]))
print('History Frequency: ' + str(df.head(1).history_freq[0]))
values = []
for row in df.number_of_trades:
values.append(row)
label_x = 'Number of Trades'
show_bar_histogram(values, label_x)
values = []
for row in df.net_profit_pct:
values.append(row)
label_x = 'Net Profit %'
show_bar_histogram(values, label_x)
values = []
for row in df.max_daily_drawdown_pct:
values.append(row)
label_x = 'Max Daily Drawdown %'
show_bar_histogram(values, label_x)
values = []
for row in df.average_exposure_pct:
values.append(row)
label_x = 'Average Exposure %'
show_bar_histogram(values, label_x)
|
edas/.ipynb_checkpoints/Explore Kryptos Dataset-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# split_at_heading: true
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#default_exp transform
# -
#export
from fastcore.imports import *
from fastcore.foundation import *
from fastcore.utils import *
from fastcore.dispatch import *
from PIL import Image
import torch
from nbdev.showdoc import *
from fastcore.test import *
# # Transforms
#
# > Definition of `Transform` and `Pipeline`
# The classes here provide functionality for creating a composition of *partially reversible functions*. By "partially reversible" we mean that a transform can be `decode`d, creating a form suitable for display. This is not necessarily identical to the original form (e.g. a transform that changes a byte tensor to a float tensor does not recreate a byte tensor when decoded, since that may lose precision, and a float tensor can be displayed already).
#
# Classes are also provided and for composing transforms, and mapping them over collections. `Pipeline` is a transform which composes several `Transform`, knowing how to decode them or show an encoded item.
# ## Transform -
# +
#export
_tfm_methods = 'encodes','decodes','setups'
class _TfmDict(dict):
def __setitem__(self,k,v):
if k not in _tfm_methods or not callable(v): return super().__setitem__(k,v)
if k not in self: super().__setitem__(k,TypeDispatch())
self[k].add(v)
# -
#export
class _TfmMeta(type):
def __new__(cls, name, bases, dict):
res = super().__new__(cls, name, bases, dict)
for nm in _tfm_methods:
base_td = [getattr(b,nm,None) for b in bases]
if nm in res.__dict__: getattr(res,nm).bases = base_td
else: setattr(res, nm, TypeDispatch(bases=base_td))
res.__signature__ = inspect.signature(res.__init__)
return res
def __call__(cls, *args, **kwargs):
f = args[0] if args else None
n = getattr(f,'__name__',None)
if callable(f) and n in _tfm_methods:
getattr(cls,n).add(f)
return f
return super().__call__(*args, **kwargs)
@classmethod
def __prepare__(cls, name, bases): return _TfmDict()
#export
def _get_name(o):
if hasattr(o,'__qualname__'): return o.__qualname__
if hasattr(o,'__name__'): return o.__name__
return o.__class__.__name__
# +
#export
class Transform(metaclass=_TfmMeta):
"Delegates (`__call__`,`decode`,`setup`) to (`encodes`,`decodes`,`setups`) if `split_idx` matches"
split_idx,init_enc,as_item_force,as_item,order,train_setup = None,False,None,True,0,None
def __init__(self, enc=None, dec=None, split_idx=None, as_item=False, order=None):
self.split_idx,self.as_item = ifnone(split_idx, self.split_idx),as_item
if order is not None: self.order=order
self.init_enc = enc or dec
if not self.init_enc: return
self.encodes,self.decodes,self.setups = TypeDispatch(),TypeDispatch(),TypeDispatch()
if enc:
self.encodes.add(enc)
self.order = getattr(enc,'order',self.order)
if len(type_hints(enc)) > 0: self.input_types = first(type_hints(enc).values())
self._name = _get_name(enc)
if dec: self.decodes.add(dec)
@property
def use_as_item(self): return ifnone(self.as_item_force, self.as_item)
@property
def name(self): return getattr(self, '_name', _get_name(self))
def __call__(self, x, **kwargs): return self._call('encodes', x, **kwargs)
def decode (self, x, **kwargs): return self._call('decodes', x, **kwargs)
def __repr__(self): return f'{self.name}: {self.use_as_item} {self.encodes} {self.decodes}'
def setup(self, items=None, train_setup=False):
train_setup = train_setup if self.train_setup is None else self.train_setup
return self.setups(getattr(items, 'train', items) if train_setup else items)
def _call(self, fn, x, split_idx=None, **kwargs):
if split_idx!=self.split_idx and self.split_idx is not None: return x
f = getattr(self, fn)
if self.use_as_item or not is_listy(x): return self._do_call(f, x, **kwargs)
res = tuple(self._do_call(f, x_, **kwargs) for x_ in x)
return retain_type(res, x)
def _do_call(self, f, x, **kwargs):
return x if f is None else retain_type(f(x, **kwargs), x, f.returns_none(x))
add_docs(Transform, decode="Delegate to `decodes` to undo transform", setup="Delegate to `setups` to set up transform")
# -
show_doc(Transform)
# A `Transform` is the main building block of the fastai data pipelines. In the most general terms a transform can be any function you want to apply to your data, however the `Transform` class provides several mechanisms that make the process of building them easy and flexible.
#
# ### The main `Transform` features:
#
# - **Type dispatch** - Type annotations are used to determine if a transform should be applied to the given argument. It also gives an option to provide several implementations and it choses the one to run based on the type. This is useful for example when running both independent and dependent variables through the pipeline where some transforms only make sense for one and not the other. Another usecase is designing a transform that handles different data formats. Note that if a transform takes multiple arguments only the type of the first one is used for dispatch.
# - **Handling of tuples** - When a tuple (or another collection satisfying `is_listy`) of data is passed to a transform it will get applied to each element separately. Most comonly it will be a *(x,y)* tuple, but it can be anything for example a list of images. You can opt out of this behavior by setting the flag `as_item=True`. For transforms that must always operate on the tuple level you can set `as_item_force=True` which takes precedence over `as_item`, an example of that is `PointScaler`.
# - **Reversability** - A transform can be made reversible by implementing the `decodes` method. This is mainly used to turn something like a category which is encoded as a number back into a label understandable by humans for showing purposes.
# - **Type propagation** - Whenever possible a transform tries to return data of the same type it received. Mainly used to maintain semantics of things like `ArrayImage` which is a thin wrapper of pytorches `Tensor`. You can opt out of this behavior by adding `->None` return type annotation.
# - **Preprocessing** - The `setup` method can be used to perform any one-time calculations to be later used by the transform, for example generating a vocabulary to encode categorical data.
# - **Filtering based on the dataset type** - By setting the `split_idx` flag you can make the transform be used only in a specific `DataSource` subset like in training, but not validation.
# - **Ordering** - You can set the `order` attribute which the `Pipeline` uses when it needs to merge two lists of transforms.
# - **Appending new behavior with decorators** - You can easily extend an existing `Transform` by creating `encodes` or `decodes` methods for new data types. You can put those new methods outside the original transform definition and decorate them with the class you wish them patched into. This can be used by the fastai library users to add their own behavior, or multiple modules contributing to the same transform.
#
# ### Defining a `Transform`
# There are a few ways to create a transform with different ratios of simplicity to flexibility.
# - **Extending the `Transform` class** - Use inheritence to implement the methods you want.
# - **Passing methods to the constructor** - Instantiate the `Transform` class and pass your functions as `enc` and `dec` arguments.
# - **@Transform decorator** - Turn any function into a `Transform` by just adding a decorator - very straightforward if all you need is a single `encodes` implementation.
# - **Passing a function to fastai APIs** - Same as above, but when passing a function to other transform aware classes like `Pipeline` or `TfmdDS` you don't even need a decorator. Your function will get converted to a `Transform` automatically.
# +
class A(Transform): pass
@A
def encodes(self, x): return x+1
f1 = A()
test_eq(f1(1), 2)
class B(A): pass
@B
def decodes(self, x): return x-1
f2 = B()
test_eq(f2(1), 2)
test_eq(f2.decode(2), 1)
test_eq(f1.decode(2), 2)
class A(Transform): pass
f3 = A()
test_eq_type(f3(2), 2)
test_eq_type(f3.decode(2.0), 2.0)
# -
# `Transform` can be used as a decorator, to turn a function into a `Transform`.
f = Transform(lambda o:o//2)
test_eq_type(f(2), 1)
test_eq_type(f.decode(2.0), 2.0)
# +
@Transform
def f(x): return x//2
test_eq_type(f(2), 1)
test_eq_type(f.decode(2.0), 2.0)
@Transform
def f(x): return x*2
test_eq_type(f(2), 4)
test_eq_type(f.decode(2.0), 2.0)
# -
# You can derive from `Transform` and use `encodes` for your encoding function.
# +
class ArrayImage(ndarray):
_show_args = {'cmap':'viridis'}
def __new__(cls, x, *args, **kwargs):
if isinstance(x,tuple): super().__new__(cls, x, *args, **kwargs)
if args or kwargs: raise RuntimeError('Unknown array init args')
if not isinstance(x,ndarray): x = array(x)
return x.view(cls)
def show(self, ctx=None, figsize=None, **kwargs):
if ctx is None: _,ctx = plt.subplots(figsize=figsize)
ctx.imshow(im, **{**self._show_args, **kwargs})
ctx.axis('off')
return ctx
im = Image.open(TEST_IMAGE)
im_t = ArrayImage(im)
# -
class A(Transform):
def encodes(self, x:ArrayImage): return -x
def decodes(self, x:ArrayImage): return x+1
def setups (self, x:ArrayImage): x.foo = 'a'
f = A()
t = f(im_t)
test_eq(t, -im_t)
test_eq(f(1), 1)
test_eq(type(t), ArrayImage)
test_eq(f.decode(t), -im_t+1)
test_eq(f.decode(1), 1)
f.setup(im_t)
test_eq(im_t.foo, 'a')
t2 = array(1)
f.setup(t2)
assert not hasattr(f2,'foo')
f
# Without return annotation we get an `Int` back since that's what was passed.
# +
class A(Transform): pass
@A
def encodes(self, x:Int): return x//2
@A
def encodes(self, x:float): return x+1
f = A()
test_eq_type(f(Int(2)), Int(1))
test_eq_type(f(2), 2)
test_eq_type(f(2.), 3.)
# -
# Without return annotation we don't cast if we're not a subclass of the input type. If the annotation is a tuple, then any type in the tuple will match.
# +
class A(Transform):
def encodes(self, x:(Int,float)): return x/2
def encodes(self, x:(str,list)): return str(x)+'1'
f = A()
test_eq_type(f(Int(2)), 1.)
test_eq_type(f(2), 2)
test_eq_type(f(Float(2.)), Float(1.))
test_eq_type(f('a'), 'a1')
# -
# With return annotation `None` we get back whatever Python creates usually.
def func(x)->None: return x/2
f = Transform(func)
test_eq_type(f(2), 1.)
test_eq_type(f(2.), 1.)
# Since `decodes` has no return annotation, but `encodes` created an `Int` and we pass that result here to `decode`, we end up with an `Int`.
def func(x): return Int(x+1)
def dec (x): return x-1
f = Transform(func,dec)
t = f(1)
test_eq_type(t, Int(2))
test_eq_type(f.decode(t), Int(1))
# If the transform has `split_idx` then it's only applied if `split_idx` param matches.
f.split_idx = 1
test_eq(f(1, split_idx=1),2)
test_eq_type(f(1, split_idx=0), 1)
# If `as_item=True` the transform takes tuples as a whole and is applied to them.
# +
class A(Transform):
def encodes(self, xy): x,y=xy; return (x+y,y)
def decodes(self, xy): x,y=xy; return (x-y,y)
f = A(as_item=True)
t = f((1,2))
test_eq(t, (3,2))
test_eq(f.decode(t), (1,2))
f.split_idx = 1
test_eq(f((1,2), split_idx=1), (3,2))
test_eq(f((1,2), split_idx=0), (1,2))
# +
class AL(Transform): pass
@AL
def encodes(self, x): return L(x_+1 for x_ in x)
@AL
def decodes(self, x): return L(x_-1 for x_ in x)
f = AL(as_item=True)
t = f([1,2])
test_eq(t, [2,3])
test_eq(f.decode(t), [1,2])
# -
# If `as_item=False` the transform is applied to each element of a listy input.
# +
def neg_int(x:numbers.Integral): return -x
f = Transform(neg_int, as_item=False)
test_eq(f([1]), (-1,))
test_eq(f([1.]), (1.,))
test_eq(f([1.,2,3.]), (1.,-2,3.))
test_eq(f.decode([1,2]), (1,2))
# -
#hide
test_eq(f.input_types, numbers.Integral)
#export
class InplaceTransform(Transform):
"A `Transform` that modifies in-place and just returns whatever it's passed"
def _call(self, fn, x, split_idx=None, **kwargs):
super()._call(fn,x,split_idx,**kwargs)
return x
#hide
import pandas as pd
class A(InplaceTransform): pass
@A
def encodes(self, x:pd.Series): x.fillna(10, inplace=True)
f = A()
test_eq_type(f(pd.Series([1,2,None])),pd.Series([1,2,10]))
# ### TupleTransform
#export
class TupleTransform(Transform):
"`Transform` that always treats `as_item` as `False`"
as_item_force=False
#export
class ItemTransform (Transform):
"`Transform` that always treats `as_item` as `True`"
as_item_force=True
# +
def float_to_int(x:(float,int)): return Int(x)
f = TupleTransform(float_to_int)
test_eq_type(f([1.]), (Int(1),))
test_eq_type(f([1]), (Int(1),))
test_eq_type(f(['1']), ('1',))
test_eq_type(f([1,'1']), (Int(1),'1'))
test_eq(f.decode([1]), [1])
test_eq_type(f(Tuple(1.)), Tuple(Int(1)))
# -
class B(TupleTransform): pass
class C(TupleTransform): pass
f = B()
test_eq(f([1]), [1])
# +
@B
def encodes(self, x:int): return x+1
@B
def encodes(self, x:str): return x+'1'
@B
def encodes(self, x)->None: return str(x)+'!'
b,c = B(),C()
test_eq(b([1]), [2])
test_eq(b(['1']), ('11',))
test_eq(b([1.0]), ('1.0!',))
test_eq(c([1]), [1])
test_eq(b([1,2]), (2,3))
test_eq(b.decode([2]), [2])
assert pickle.loads(pickle.dumps(b))
# -
@B
def decodes(self, x:int): return x-1
test_eq(b.decode([2]), [1])
test_eq(b.decode(('2',)), ('2',))
# Non-type-constrained functions are applied to all elements of a tuple.
# +
class A(TupleTransform): pass
@A
def encodes(self, x): return x+1
@A
def decodes(self, x): return x-1
f = A()
t = f((1,2.0))
test_eq_type(t, (2,3.0))
test_eq_type(f.decode(t), (1,2.0))
# -
# Type-constrained functions are applied to only matching elements of a tuple, and return annotations are only applied where matching.
# +
class B(TupleTransform):
def encodes(self, x:int): return Int(x+1)
def encodes(self, x:str): return x+'1'
def decodes(self, x:Int): return x//2
f = B()
start = (1.,2,'3')
t = f(start)
test_eq_type(t, (1.,Int(3),'31'))
test_eq(f.decode(t), (1.,Int(1),'31'))
# -
# The same behavior also works with `typing` module type classes.
# +
class A(Transform): pass
@A
def encodes(self, x:numbers.Integral): return x+1
@A
def encodes(self, x:float): return x*3
@A
def decodes(self, x:int): return x-1
f = A()
start = 1.0
t = f(start)
test_eq(t, 3.)
test_eq(f.decode(t), 3)
f = A(as_item=False)
start = (1.,2,3.)
t = f(start)
test_eq(t, (3.,3,9.))
test_eq(f.decode(t), (3.,2,9.))
# -
# Transform accepts lists
# +
def a(x): return L(x_+1 for x_ in x)
def b(x): return L(x_-1 for x_ in x)
f = TupleTransform(a,b)
t = f((L(1,2),))
test_eq(t, (L(2,3),))
test_eq(f.decode(t), (L(1,2),))
# -
# ### Func -
#export
def get_func(t, name, *args, **kwargs):
"Get the `t.name` (potentially partial-ized with `args` and `kwargs`) or `noop` if not defined"
f = getattr(t, name, noop)
return f if not (args or kwargs) else partial(f, *args, **kwargs)
# This works for any kind of `t` supporting `getattr`, so a class or a module.
test_eq(get_func(operator, 'neg', 2)(), -2)
test_eq(get_func(operator.neg, '__call__')(2), -2)
test_eq(get_func(list, 'foobar')([2]), [2])
t = get_func(torch, 'zeros', dtype=torch.int64)(5)
test_eq(t.dtype, torch.int64)
a = [2,1]
get_func(list, 'sort')(a)
test_eq(a, [1,2])
# Transforms are built with multiple-dispatch: a given function can have several methods depending on the type of the object received. This is done directly with the `TypeDispatch` module and type-annotation in `Transform`, but you can also use the following class.
#export
class Func():
"Basic wrapper around a `name` with `args` and `kwargs` to call on a given type"
def __init__(self, name, *args, **kwargs): self.name,self.args,self.kwargs = name,args,kwargs
def __repr__(self): return f'sig: {self.name}({self.args}, {self.kwargs})'
def _get(self, t): return get_func(t, self.name, *self.args, **self.kwargs)
def __call__(self,t): return mapped(self._get, t)
# You can call the `Func` object on any module name or type, even a list of types. It will return the corresponding function (with a default to `noop` if nothing is found) or list of functions.
# +
test_eq(Func('sqrt')(math), math.sqrt)
test_eq(Func('sqrt')(torch), torch.sqrt)
@patch
def powx(x:math, a): return math.pow(x,a)
@patch
def powx(x:torch, a): return torch.pow(x,a)
tst = Func('powx',a=2)([math, torch])
test_eq([f.func for f in tst], [math.powx, torch.powx])
for t in tst: test_eq(t.keywords, {'a': 2})
# +
#export
class _Sig():
def __getattr__(self,k):
def _inner(*args, **kwargs): return Func(k, *args, **kwargs)
return _inner
Sig = _Sig()
# -
show_doc(Sig, name="Sig")
# `Sig` is just sugar-syntax to create a `Func` object more easily with the syntax `Sig.name(*args, **kwargs)`.
f = Sig.sqrt()
test_eq(f(math), math.sqrt)
test_eq(f(torch), torch.sqrt)
# ## Pipeline -
#export
def compose_tfms(x, tfms, is_enc=True, reverse=False, **kwargs):
"Apply all `func_nm` attribute of `tfms` on `x`, maybe in `reverse` order"
if reverse: tfms = reversed(tfms)
for f in tfms:
if not is_enc: f = f.decode
x = f(x, **kwargs)
return x
def to_int (x): return Int(x)
def to_float(x): return Float(x)
def double (x): return x*2
def half(x)->None: return x/2
# +
def test_compose(a, b, *fs): test_eq_type(compose_tfms(a, tfms=map(Transform,fs)), b)
test_compose(1, Int(1), to_int)
test_compose(1, Float(1), to_int,to_float)
test_compose(1, Float(2), to_int,to_float,double)
test_compose(2.0, 2.0, to_int,double,half)
# +
class A(Transform):
def encodes(self, x:float): return Float(x+1)
def decodes(self, x): return x-1
tfms = [A(), Transform(math.sqrt)]
t = compose_tfms(3., tfms=tfms)
test_eq_type(t, Float(2.))
test_eq(compose_tfms(t, tfms=tfms, is_enc=False), 1.)
test_eq(compose_tfms(4., tfms=tfms, reverse=True), 3.)
# -
tfms = [A(as_item=False), Transform(math.sqrt, as_item=False)]
test_eq(compose_tfms((9,3.), tfms=tfms), (3,2.))
#export
def mk_transform(f, as_item=True):
"Convert function `f` to `Transform` if it isn't already one"
f = instantiate(f)
return f if isinstance(f,Transform) else Transform(f, as_item=as_item)
def neg(x): return -x
test_eq(type(mk_transform(neg)), Transform)
test_eq(type(mk_transform(math.sqrt)), Transform)
test_eq(type(mk_transform(lambda a:a*2)), Transform)
#export
def gather_attrs(o, k, nm):
"Used in __getattr__ to collect all attrs `k` from `self.{nm}`"
if k.startswith('_') or k==nm: raise AttributeError(k)
att = getattr(o,nm)
res = [t for t in att.attrgot(k) if t is not None]
if not res: raise AttributeError(k)
return res[0] if len(res)==1 else L(res)
#export
def gather_attr_names(o, nm):
"Used in __dir__ to collect all attrs `k` from `self.{nm}`"
return L(getattr(o,nm)).map(dir).concat().unique()
#export
class Pipeline:
"A pipeline of composed (for encode/decode) transforms, setup with types"
def __init__(self, funcs=None, as_item=False, split_idx=None):
self.split_idx,self.default = split_idx,None
if isinstance(funcs, Pipeline): self.fs = funcs.fs
else:
if isinstance(funcs, Transform): funcs = [funcs]
self.fs = L(ifnone(funcs,[noop])).map(mk_transform).sorted(key='order')
for f in self.fs:
name = camel2snake(type(f).__name__)
a = getattr(self,name,None)
if a is not None: f = L(a)+f
setattr(self, name, f)
self.set_as_item(as_item)
def set_as_item(self, as_item):
self.as_item = as_item
for f in self.fs: f.as_item = as_item
def setup(self, items=None, train_setup=False):
tfms = self.fs[:]
self.fs.clear()
for t in tfms: self.add(t,items, train_setup)
def add(self,t, items=None, train_setup=False):
t.setup(items, train_setup)
self.fs.append(t)
def __call__(self, o): return compose_tfms(o, tfms=self.fs, split_idx=self.split_idx)
def __repr__(self): return f"Pipeline: {' -> '.join([f.name for f in self.fs])}"
def __getitem__(self,i): return self.fs[i]
def __setstate__(self,data): self.__dict__.update(data)
def __getattr__(self,k): return gather_attrs(self, k, 'fs')
def __dir__(self): return super().__dir__() + gather_attr_names(self, 'fs')
def decode (self, o, full=True):
if full: return compose_tfms(o, tfms=self.fs, is_enc=False, reverse=True, split_idx=self.split_idx)
#Not full means we decode up to the point the item knows how to show itself.
for f in reversed(self.fs):
if self._is_showable(o): return o
o = f.decode(o, split_idx=self.split_idx)
return o
def show(self, o, ctx=None, **kwargs):
o = self.decode(o, full=False)
o1 = [o] if self.as_item or not is_listy(o) else o
for o_ in o1:
if hasattr(o_, 'show'): ctx = o_.show(ctx=ctx, **kwargs)
return ctx
def _is_showable(self, o):
return all(hasattr(o_, 'show') for o_ in o) if is_listy(o) else hasattr(o, 'show')
add_docs(Pipeline,
__call__="Compose `__call__` of all `fs` on `o`",
decode="Compose `decode` of all `fs` on `o`",
show="Show `o`, a single item from a tuple, decoding as needed",
add="Add transform `t`",
set_as_item="Set value of `as_item` for all transforms",
setup="Call each tfm's `setup` in order")
# `Pipeline` is a wrapper for `compose_tfm`. You can pass instances of `Transform` or regular functions in `funcs`, the `Pipeline` will wrap them all in `Transform` (and instantiate them if needed) during the initialization. It handles the transform `setup` by adding them one at a time and calling setup on each, goes through them in order in `__call__` or `decode` and can `show` an object by applying decoding the transforms up until the point it gets an object that knows how to show itself.
# Empty pipeline is noop
pipe = Pipeline()
test_eq(pipe(1), 1)
pipe.set_as_item(False)
test_eq(pipe((1,)), (1,))
# Check pickle works
assert pickle.loads(pickle.dumps(pipe))
# +
class IntFloatTfm(Transform):
def encodes(self, x): return Int(x)
def decodes(self, x): return Float(x)
foo=1
int_tfm=IntFloatTfm()
def neg(x): return -x
neg_tfm = Transform(neg, neg)
# +
pipe = Pipeline([neg_tfm, int_tfm])
start = 2.0
t = pipe(start)
test_eq_type(t, Int(-2))
test_eq_type(pipe.decode(t), Float(start))
test_stdout(lambda:pipe.show(t), '-2')
pipe.set_as_item(False)
test_stdout(lambda:pipe.show(pipe((1.,2.))), '-1\n-2')
test_eq(pipe.foo, 1)
assert 'foo' in dir(pipe)
assert 'int_float_tfm' in dir(pipe)
# -
# Transforms are available as attributes named with the snake_case version of the names of their types. Attributes in transforms can be directly accessed as attributes of the pipeline.
# +
test_eq(pipe.int_float_tfm, int_tfm)
test_eq(pipe.foo, 1)
pipe = Pipeline([int_tfm, int_tfm])
pipe.int_float_tfm
test_eq(pipe.int_float_tfm[0], int_tfm)
test_eq(pipe.foo, [1,1])
# -
# Check opposite order
pipe = Pipeline([int_tfm,neg_tfm])
t = pipe(start)
test_eq(t, -2)
test_stdout(lambda:pipe.show(t), '-2')
# +
class A(Transform):
def encodes(self, x): return int(x)
def decodes(self, x): return Float(x)
pipe = Pipeline([neg_tfm, A])
t = pipe(start)
test_eq_type(t, -2)
test_eq_type(pipe.decode(t), Float(start))
test_stdout(lambda:pipe.show(t), '-2.0')
# -
s2 = (1,2)
pipe.set_as_item(False)
t = pipe(s2)
test_eq_type(t, (-1,-2))
test_eq_type(pipe.decode(t), (Float(1.),Float(2.)))
test_stdout(lambda:pipe.show(t), '-1.0\n-2.0')
class B(Transform):
def encodes(self, x): return x+1
def decodes(self, x): return x-1
# +
from PIL import Image
def f1(x:ArrayImage): return -x
def f2(x): return Image.open(x).resize((128,128))
def f3(x:Image.Image): return(ArrayImage(array(x)))
# -
pipe = Pipeline([f2,f3,f1])
t = pipe(TEST_IMAGE)
test_eq(type(t), ArrayImage)
test_eq(t, -array(f3(f2(TEST_IMAGE))))
pipe = Pipeline([f2,f3])
t = pipe(TEST_IMAGE)
ax = pipe.show(t)
test_fig_exists(ax)
#Check filtering is properly applied
add1 = B()
add1.split_idx = 1
pipe = Pipeline([neg_tfm, A(), add1])
test_eq(pipe(start), -2)
pipe.split_idx=1
test_eq(pipe(start), -1)
pipe.split_idx=0
test_eq(pipe(start), -2)
for t in [None, 0, 1]:
pipe.split_idx=t
test_eq(pipe.decode(pipe(start)), start)
test_stdout(lambda: pipe.show(pipe(start)), "-2.0")
# ### Methods
# +
#TODO: method examples
# -
show_doc(Pipeline.__call__)
show_doc(Pipeline.decode)
show_doc(Pipeline.setup)
# During the setup, the `Pipeline` starts with no transform and adds them one at a time, so that during its setup, each transform gets the items processed up to its point and not after.
# +
#hide
#Test is with TfmdList
# -
# ## Export -
#hide
from nbdev.export import notebook2script
notebook2script()
|
nbs/04_transform.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os,sys,inspect
currentdir = os.path.dirname(os.path.abspath(inspect.getfile(inspect.currentframe())))
parentdir = os.path.dirname(currentdir)
# Add parent dir to path, so that python finds the lenskit package
sys.path.insert(0,parentdir)
from lenskit.metrics import dataGenerator
from lenskit import batch, topn, util, topnFair
from lenskit import crossfold as xf
from lenskit.algorithms import Recommender, als, funksvd, user_knn as uknn, item_knn as iknn
from lenskit import topn, topnFair
import numpy as np
import pandas as pd
import math
import pickle
# %matplotlib inline
test_data = pd.read_pickle("test_data_Lenskit2")
all_recs_joined = pd.read_pickle("all_recs_joined_Lenskit2")
def evaluate_genre(genre):
providers = list(all_recs_joined.iloc[:,8:])
proItems_genre_n = all_recs_joined.loc[ all_recs_joined[genre] == 1].item.nunique()
print (genre)
rla = topnFair.FairRecListAnalysis(['user', "Algorithm"])
rla.add_metric("rND")
rla.add_metric("rKL")
rla.add_metric("rRD")
rla.add_metric("APCR")
rla.add_metric("nd_APCR")
rla.add_metric("equal_ex")
rla.add_metric("ndcg")
results= rla.compute(all_recs_joined, test_data, genre, providers)
print("done" , genre)
return results.groupby('Algorithm').mean()
testproviders = ["Drama"]
#testproviders = list(all_recs_joined.iloc[:,8:])
allResults = []
for genre in testproviders:
rep = evaluate_genre(genre)
allResults.append(rep)
output = pd.concat(allResults, keys=testproviders)
output
output.to_csv('out_1.csv')
|
notebooks/Evaluation-of-measures-LensKit-100k.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Lab 2: Data Types
# Welcome to Lab 2!
#
# Last time, we had our first look at Python and Jupyter notebooks. So far, we've only used Python to manipulate numbers. There's a lot more to life than numbers, so Python lets us represent many other types of data in programs.
#
# In this lab, you'll first see how to represent and manipulate another fundamental type of data: text. A piece of text is called a *string* in Python.
#
# You'll also see how to invoke *methods*. A method is very similar to a function. It just looks a little different because it's tied to a particular piece of data (like a piece of text or a number).
#
# Last, you'll see how to work with datasets in Python -- *collections* of data, like the numbers 2 through 5 or the words "welcome", "to", and "lab".
# Initialize the OK tests to get started.
# +
# Don't change this cell; just run it.
# The result will give you directions about how to log in to the submission system, called OK.
# Once you're logged in, you can run this cell again, but it won't ask you who you are because
# it remembers you. However, you will need to log in once per assignment.
from client.api.notebook import Notebook
ok = Notebook('lab02.ok')
import os
if not os.path.exists(os.path.join(os.environ.get("HOME"), ".config/ok/auth_refresh")):
ok.auth(force=True)
else:
ok.auth(inline=True)
_ = ok.submit()
# -
# **Deadline**: TODO: Submit this assignment by October 16, 2018 09:00 am
#
# **Submission**: Once you're finished, select "Save and Checkpoint" in the File menu and then execute the submit cell below (or at the end). The result will contain a link that you can use to check that your assignment has been submitted successfully.
_ = ok.submit()
# # 1. Review: The building blocks of Python code
#
# The two building blocks of Python code are *expressions* and *statements*. An **expression** is a piece of code that
#
# * is self-contained, meaning it would make sense to write it on a line by itself, and
# * usually has a value.
#
#
# Here are two expressions that both evaluate to 3
#
# 3
# 5 - 2
#
# One important form of an expression is the **call expression**, which first names a function and then describes its arguments. The function returns some value, based on its arguments. Some important mathematical functions are
#
# | Function | Description |
# |----------|---------------------------------------------------------------|
# | `abs` | Returns the absolute value of its argument |
# | `max` | Returns the maximum of all its arguments |
# | `min` | Returns the minimum of all its arguments |
# | `pow` | Raises its first argument to the power of its second argument |
# | `round` | Round its argument to the nearest integer |
#
# Here are two call expressions that both evaluate to 3
#
# abs(2 - 5)
# max(round(2.8), min(pow(2, 10), -1 * pow(2, 10)))
#
# All these expressions but the first are **compound expressions**, meaning that they are actually combinations of several smaller expressions. `2 + 3` combines the expressions `2` and `3` by addition. In this case, `2` and `3` are called **subexpressions** because they're expressions that are part of a larger expression.
#
# A **statement** is a whole line of code. Some statements are just expressions. The expressions listed above are examples.
#
# Other statements *make something happen* rather than *having a value*. After they are run, something in the world has changed. For example, an **assignment statement** assigns a value to a name.
#
# A good way to think about this is that we're **evaluating the right-hand side** of the equals sign and **assigning it to the left-hand side**. Here are some assignment statements:
#
# height = 1.3
# the_number_five = abs(-5)
# absolute_height_difference = abs(height - 1.688)
#
# A key idea in programming is that large, interesting things can be built by combining many simple, uninteresting things. The key to understanding a complicated piece of code is breaking it down into its simple components.
#
# For example, a lot is going on in the last statement above, but it's really just a combination of a few things. This picture describes what's going on.
#
# <img src="statement.jpg">
# **Question 1.1.** In the next cell, assign the name `new_year` to the larger number among the following two numbers:
#
# 1. the absolute value of $2^{5}-2^{11}-2^1$, and
# 2. $5 \times 13 \times 31 + 2$.
#
# Try to use just one statement (one line of code).
new_year = ...
new_year
# Check your work by executing the next cell.
_ = ok.grade('q11')
# # 2. Text
# Programming doesn't just concern numbers; text is one of the most common types of values used in programs.
#
# A snippet of text is represented by a **string value** in Python. The word "*string*" is a programming term for a sequence of characters. A string might contain a single character, a word, a sentence, or a whole book.
#
# To distinguish text data from actual code, we demarcate strings by putting quotation marks around them. Single quotes (`'`) and double quotes (`"`) are both valid, but the types of opening and closing quotation marks **must** match. The contents can be any sequence of characters, including numbers and symbols.
#
# We've seen strings before in `print` statements. Below, two different strings are passed as arguments to the `print` function.
print("I <3", 'Data Science')
# Just like how numbers can be assigned to variables, strings can be assigned to variables as well. Any variable name can be assigned to a string, but you should use a variable name that describes what the variable is used for.
number = 'two'
operator = '*'
print(number, operator, number)
# **Question 2.1.** In Avengers: Endgame, <NAME> said his famous last words in a battle with Thanos. Here was his final conversation:
#
# Thanos said: "I am inevitable"
# <NAME> replied: "I am Iron Man"
#
# The cell below contains unfinished code. Fill in the `...`s so that it prints out this conversation *exactly* as it appears above.
# +
thanos_speaking = ...
thanos_quote = '"I am inevitable"'
tony_reply = '<NAME> replied:'
tony_quote = ...
print(thanos_speaking, thanos_quote)
print(tony_reply, tony_quote)
# -
_ = ok.grade('q21')
# ## 2.1. String Methods
# Strings can be transformed using **methods**, which are functions that involve an existing string and some other arguments. One example is the `replace` method, which replaces all instances of some part of a string with some alternative.
#
# A method is invoked on a string by placing a `.` after the string value, then the name of the method, and finally parentheses containing the arguments. Here's a sketch, where the `<` and `>` symbols aren't part of the syntax; they just mark the boundaries of sub-expressions.
#
# <expression that evaluates to a string>.<method name>(<argument>, <argument>, ...)
#
# You can also "chain" them by calling methods one after another:
#
# <expression>.<method name>(<argument>, <argument>, ...).<method name>(<argument>, <argument>, ...)
#
# Try to predict the output of these examples, then execute them.
#
# +
greet = 'hello'
first_person = 'hitchhiker'
# Replace one letter and assign to a new variable
slang = greet.replace('o', 'y').replace('e', 'o')
# Capitalize the word
slang = slang.title()
# Replace a sequence of letters, which appears twice, and assign to a new variable
second_person = first_person.replace('hi', 'ma')
# Print the results
print(greet, 'to', slang)
print(first_person, 'to', second_person)
# -
# You can also invoke methods on the results of other method (or function) calls. Here's an example:
#
# Calling replace on the output of another call to
# replace
'train'.replace('t', 'ing').replace('in', 'de')
# Here's a picture of how Python evaluates a "chained" method call like that:
#
# <img src="chaining_method_calls.jpg"/>
# **Question 2.1.1.** Assign strings to the names `you` and `this` so that the final expression evaluates to a 10-letter English word with three double letters in a row.
#
# *Hint:* After you guess at some values for `you` and `this`, it's helpful to see the value of the variable `the`. Try printing the value of `the` by adding a line like this:
#
# print(the)
#
# *Hint 2:* Run the tests if you're stuck. They sometimes will offer help.
you = ...
this = ...
a = 'beeper'
the = a.replace('p', you)
the.replace('bee', this)
_ = ok.grade('q211')
# Other string methods do not take any arguments at all, because the original string is all that's needed to compute the result. In this case, parentheses are still needed, but there's nothing in between the parentheses. Here are some methods that work that way:
#
# |Method name|Value|
# |-|-|
# |`lower`|a lowercased version of the string|
# |`upper`|an uppercased version of the string|
# |`capitalize`|a version with the first letter capitalized|
# |`title`|a version with the first letter of every word capitalized||
#
'unIverSITy of caliFORnia'.title()
# All these string methods are useful, but most programmers don't memorize their names or how to use them. In the "real world," people usually just search the internet for documentation and examples. A complete [list of string methods](https://docs.python.org/3/library/stdtypes.html#string-methods) appears in the Python language documentation. [Stack Overflow](http://stackoverflow.com) has a huge database of answered questions that often demonstrate how to use these methods to achieve various ends.
# ## 2.2. Converting to and from Strings
# Strings and numbers are different *types* of values, even when a string contains the digits of a number. For example, evaluating the following cell causes an error because an integer cannot be added to a string. **After you see the error, comment out the line using `#`, so that you don't get an error.**
8 + "8"
# However, there are built-in functions to convert numbers to strings and strings to numbers.
#
# int(): Converts a string of digits to an integer ("int") value
# float(): Converts a string of digits, perhaps with a decimal point, to a decimal ("float") value
# str(): Converts any value to a string
# Try to predict what the following cell will evaluate to, then evaluate it.
8 + int("8")
# Suppose you're writing a program that looks for dates in a text, and you want your program to find the amount of time that elapsed between two years it has identified. It doesn't make sense to subtract two texts, but you can first convert the text containing the years into numbers.
#
# **Question 2.2.1.** Finish the code below to compute the number of years that elapsed between `one_year` and `another_year`. Don't just write the numbers `1989` and `2019` (or `30`); use a conversion function to turn the given text data into numbers. We should be able to change the values in `one_year` and `another_year` and use your `difference` expression to get the correct result.
# +
# Some text data:
one_year = "1989"
another_year = "2019"
# Complete the next line. Note that we can't just write:
# another_year - one_year
# If you don't see why, try seeing what happens when you
# write that here.
difference = ...
difference
# -
_ = ok.grade('q221')
# ## 2.3. Strings as function arguments
#
# String values, like numbers, can be arguments to functions and can be returned by functions. The function `len` takes a single string as its argument and returns the number of characters in the string: its **len**-gth.
#
# Note that it doesn't count *words*; it counts characters. `len("one small step for man")` is 22, not 5.
#
# **Question 2.3.1.** Use `len` to find out the number of characters in the very long string in the next cell. (It's the first sentence of the English translation of the French [Declaration of the Rights of Man](http://avalon.law.yale.edu/18th_century/rightsof.asp).) The length of a string is the total number of characters in it, including things like spaces and punctuation. Assign `sentence_length` to that number.
a_very_long_sentence = "The representatives of the French people, organized as a National Assembly, believing that the ignorance, neglect, or contempt of the rights of man are the sole cause of public calamities and of the corruption of governments, have determined to set forth in a solemn declaration the natural, unalienable, and sacred rights of man, in order that this declaration, being constantly before all the members of the Social body, shall remind them continually of their rights and duties; in order that the acts of the legislative power, as well as those of the executive power, may be compared at any moment with the objects and purposes of all political institutions and may thus be more respected, and, lastly, in order that the grievances of the citizens, based hereafter upon simple and incontestable principles, shall tend to the maintenance of the constitution and redound to the happiness of all."
sentence_length = ...
sentence_length
_ = ok.grade('q231')
# # 3. Importing code
#
# > What has been will be again,
# > what has been done will be done again;
# > there is nothing new under the sun.
#
# Most programming involves work that is very similar to work that has been done before. Since writing code is time-consuming, it's good to rely on others' published code when you can. Rather than copy-pasting, Python allows us to **import** other code, creating a **module** that contains all of the names created by that code.
#
# Python includes many useful modules that are just an `import` away. We'll look at the `math` module as a first example. The `math` module is extremely useful in computing mathematical expressions in Python.
#
# Suppose we want to very accurately compute the area of a circle with radius 5 meters. For that, we need the constant $\pi$, which is roughly 3.14. Conveniently, the `math` module has `pi` defined for us:
import math
radius = 5
area_of_circle = radius**2 * math.pi
area_of_circle
# `pi` is defined inside `math`, and the way that we access names that are inside modules is by writing the module's name, then a dot, then the name of the thing we want:
#
# <module name>.<name>
#
# In order to use a module at all, we must first write the statement `import <module name>`. That statement creates a module object with things like `pi` in it and then assigns the name `math` to that module. Above we have done that for `math`.
# **Question 3.1.** `math` also provides the name `e` for the base of the natural logarithm, which is roughly 2.71. Compute $e^{\pi}-\pi$, giving it the name `near_twenty`. (_Hint_: the comic below shows you what the answer is supposed to be. :-))
near_twenty = ...
near_twenty
_ = ok.grade('q31')
# (Source: [XKCD Comics](https://www.xkcd.com/217))
# ## 3.1. Importing functions
#
# **Modules** can provide other named things, including **functions**. For example, `math` provides the name `sin` for the sine function. Having imported `math` already, we can write `math.sin(3)` to compute the sine of 3. (Note that this sine function considers its argument to be in [radians](https://en.wikipedia.org/wiki/Radian), not degrees. 180 degrees are equivalent to $\pi$ radians.)
#
# **Question 3.1.1.** A $\frac{\pi}{4}$-radian (45-degree) angle forms a right triangle with equal base and height, pictured below. If the hypotenuse (the radius of the circle in the picture) is 1, then the height is $\sin(\frac{\pi}{4})$. Compute that using `sin` and `pi` from the `math` module. Give the result the name `sine_of_pi_over_four`.
#
# 
# (Source: [Wolfram MathWorld](http://mathworld.wolfram.com/images/eps-gif/TrigonometryAnglesPi4_1000.gif))
sine_of_pi_over_four = ...
sine_of_pi_over_four
_ = ok.grade('q311')
# For your reference, here are some more examples of functions from the `math` module.
#
# Note how different methods take in different number of arguments. Often, the documentation of the module will provide information on how many arguments is required for each method. You can check documentation by adding a question mark `?` at the end of the function name, e.g., `math.log?`.
# Calculating factorials.
math.factorial(5)
# Calculating logarithms (the logarithm of 8 in base 2).
# The result is 3 because 2 to the power of 3 is 8.
math.log(8, 2)
# Calculating square roots.
math.sqrt(5)
# Calculating cosines.
math.cos(math.pi)
# There's many variations of how we can import methods from outside sources. For example, we can import just a specific method from an outside source, we can rename a library we import, and we can import every single method from a whole library.
#Importing just cos and pi from math.
#Notice that we now don't have to use `math.` before calling cos and pi
from math import cos, pi
print(cos(pi))
#We do have to use it infront of other methods from math, though
math.log(pi)
#We can nickname math as something else, if we don't want to type math
import math as m
m.log(m.pi)
#Lastly, we can import everything from math
from math import *
log(pi)
# ##### Applications : Continuously Compounded Interest
#
# **Question 3.1.2.** Suppose we decide to invest some money in an account that continously compounds interest. In other words, the amount that we initially invested is constantly earning interest and any interest that we earned also keeps acquiring interest.
#
# The formula for _continously compounded interest_ is modeled by $A = Pe^{Rt}$, where $A$ is the amount earned, $P$ is the _principal_, which is our initial investment, $e$ is the Euler's constant, $R$ is the rate of interest per year, and $t$ is time in years.
#
# Suppose we decide to invest 50,000 dollars at the bank of INT 5. The bank offers an interest rate of 8.0%. Assuming that our investment continously compounds interest, compute the amount of money earned after 10 years (using `math.e`).
# +
# Import math module
import math as m
# Using the parameters from above, Calculate the total amount earned
# Principal
initial_investment = ...
# Time in years
time = ...
# Rate of interest
interest_rate = ...
# Amount earned
total = ...
total
# -
_ = ok.grade('q312')
# ##### A function that displays a picture
# People have written Python functions that do very cool and complicated things, like crawling web pages for data, transforming videos, or doing machine learning with lots of data. Now that you can import things, when you want to do something with code, first check to see if someone else has done it for you.
#
# Let's see an example of a function that's used for downloading and displaying pictures.
#
# The module `IPython.display` provides a function called `Image`. The `Image` function takes a single argument, **a string** that is the URL of the image on the web. It returns an *image* value that this Jupyter notebook understands how to display. To display an image, make it the value of the last expression in a cell, just like you'd display a number or a string.
#
# **Question 3.1.3.** In the next cell, import the module `IPython.display` and use its `Image` function to display the image at this URL:
#
# https://upload.wikimedia.org/wikipedia/commons/thumb/8/8c/David_-_The_Death_of_Socrates.jpg/1024px-David_-_The_Death_of_Socrates.jpg
#
# Since we are importing `IPython.display`, instead of all the functions from it, the process is similar to what we did when we wrote `import math`: we need to **use the name of the module** (in this case, `IPython.display`) **before we call its function**: e.g., `math.pi`, instead of just `pi`.
#
# Give the name `art` to the output of the call to `Image`. (It might take a few seconds to load the image. It's a painting called *The Death of Socrates* by <NAME>, depicting events from a philosophical text by Plato.)
#
# *Hint*: A link isn't any special type of data type in Python. You can't just write a link into Python and expect it to work; you need to type the link in as a **specific data type**. Which one makes the most sense? (See the answer in the explanation above.)
# Import the module IPython.display. Watch out for capitalization.
import IPython.display
# Replace the ... with a call to the Image function
# in the IPython.display module, which should produce
# a picture.
art = ...
art
_ = ok.grade('q313')
# # 4. Arrays
#
# Up to now, we haven't done much that you couldn't do yourself by hand, without going through the trouble of learning Python. Computers are most useful when you can use a small amount of code to *do the same action* to *many different things*.
#
# For example, in the time it takes you to calculate the 18% tip on a restaurant bill, a laptop can calculate 18% tips for every restaurant bill paid by every human on Earth that day. (That's if you're pretty fast at doing arithmetic in your head!)
#
# **Arrays** are how we put many values in one place so that we can operate on them as a group. For example, if `billions_of_numbers` is an array of numbers, the expression
#
# 0.18 * billions_of_numbers
#
# gives a new array of numbers that's the result of multiplying each number in `billions_of_numbers` by 0.18 (18%). Arrays are not limited to numbers; we can also put all the words in a book into an array of strings.
#
# Concretely, an array is a **collection of values of the same type**, like a column in an Excel spreadsheet.
#
# <img src="excel_array.jpg">
# ## 4.1. Making arrays
# You can type in the data that goes in an array yourself, but that's not typically how programs work. Normally, we create arrays by loading them from an external source, like a data file.
#
# First, though, let's learn how to do it the hard way. Execute the following cell so that all the methods from the `datascience` module are available to you.
from datascience import *
# Now, to create an array, call the function `make_array`. Each argument you pass to `make_array` will be in the array it returns. Run this cell to see an example:
make_array(0.125, 4.75, -1.3)
# Each value in an array (in the above case, the numbers 0.125, 4.75, and -1.3) is called an *element* of that array.
#
# Arrays themselves are also values, just like numbers and strings. That means you can assign them names or use them as arguments to functions.
#
# **Question 4.1.1.** Make an array containing the numbers 1, 2, and 3, in that order. Name it `small_numbers`.
small_numbers = ...
small_numbers
_ = ok.grade('q411')
# **Question 4.1.2.** Make an array containing the numbers 0, 1, -1, $\pi$, and $e$, in that order. Name it `interesting_numbers`. *Hint:* How did you get the values $\pi$ and $e$ earlier? You can refer to them in exactly the same way here.
import math
interesting_numbers = [0, 1, -1, math.pi, math.e]
interesting_numbers
_ = ok.grade('q412')
# **Question 4.1.3.** Make an array containing the five strings `"Hello"`, `","`, `" "`, `"world"`, and `"!"`. (The third one is a single space inside quotes.) Name it `hello_world_components`.
#
# *Note:* If you print `hello_world_components`, you'll notice some extra information in addition to its contents: `dtype='<U5'`. That's just NumPy's extremely cryptic way of saying that the things in the array are strings.
hello_world_components = ...
hello_world_components
_ = ok.grade('q413')
# The `join` method of a string takes an array of strings as its argument and puts all of the elements together into one string. Try it:
'.'.join(make_array('U','C','S','B'))
# **Question 4.1.4.** Assign `separator` to a string so that the name `hello` is bound to (i.e., assigned to) the string `'Hello, world!'` in the cell below.
separator = ...
hello = separator.join(hello_world_components)
hello
_ = ok.grade('q414')
# ### 4.1.1. `np.arange`
# Arrays are provided by a package called [NumPy](http://www.numpy.org/) (pronounced "NUM-pie" or, if you prefer to pronounce things incorrectly, "NUM-pee"). The package is called `numpy`, but it's standard to rename it `np` for brevity. You can do that with:
#
# import numpy as np
#
# Very often in data science, we want to work with many numbers that are evenly spaced within some range. NumPy provides a special function for this called `arange`. `np.arange(start, stop, space)` produces an array with all the numbers starting at `start` and counting up by `space`, stopping **before** `stop` is reached.
#
# For example, the value of `np.arange(1, 6, 2)` is an array with elements 1, 3, and 5 -- it starts at 1 and counts up by 2, then stops before 6. In other words, it's equivalent to `make_array(1, 3, 5)`.
#
# `np.arange(4, 9, 1)` is an array with elements 4, 5, 6, 7, and 8. (It doesn't contain 9 because `np.arange` stops *before* the stop value is reached.)
#
# **Question 4.1.1.1.** Import `numpy` as `np` and then use `np.arange` to create an array with the multiples of 99 from 0 up to (**and including**) 9999. (So its elements are 0, 99, 198, 297, etc.)
...
multiples_of_99 = ...
multiples_of_99
_ = ok.grade('q4111')
# ##### Temperature readings
# NOAA (the US National Oceanic and Atmospheric Administration) operates weather stations that measure surface temperatures at different sites around the United States. The hourly readings are [publicly available](http://www.ncdc.noaa.gov/qclcd/QCLCD?prior=N).
#
# Suppose we download all the hourly data from the Oakland, California site for the month of December 2015. To analyze the data, we want to know when each reading was taken, but we find that the data don't include the timestamps of the readings (the time at which each one was taken).
#
# However, we know the first reading was taken at the first instant of December 2015 (midnight on December 1st) and each subsequent reading was taken exactly 1 hour after the last.
#
# **Question 4.1.1.2.** Create an array of the *time, in seconds, since the start of the month* at which each hourly reading was taken. Name it `collection_times`.
#
# *Hint 1:* There were 31 days in December, which is equivalent to ($31 \times 24$) hours or ($31 \times 24 \times 60 \times 60$) seconds. So your array should have $31 \times 24$ elements in it.
#
# *Hint 2:* The `len` function works on arrays, too. If your `collection_times` isn't passing the tests, check its length and make sure it has $31 \times 24$ elements.
collection_times = ...
collection_times
_ = ok.grade('q4112')
# ## 4.2. Working with single elements of arrays ("indexing")
# Let's work with a more interesting dataset. The next cell creates an array called `population` that includes estimated world populations in every year from **1950** to roughly the present. (The estimates come from the [US Census Bureau website](http://www.census.gov/population/international/data/worldpop/table_population.php).)
#
# Rather than type in the data manually, we've loaded them from a file on your computer called `world_population.csv`. You'll learn how to do that next week.
# Don't worry too much about what goes on in this cell.
from datascience import *
population = Table.read_table("world_population.csv").column("Population")
population
# Here's how we get the first element of `population`, which is the world population in the first year in the dataset, 1950.
population.item(0)
# The value of that expression is the number 2557628654 (around 2.5 billion), because that's the first thing in the array `population`.
#
# Notice that we wrote `.item(0)`, not `.item(1)`, to get the first element. This is a weird convention in computer science. 0 is called the *index* of the first item. It's the number of elements that appear *before* that item. So 3 is the index of the 4th item.
#
# Here are some more examples. In the examples, we've given names to the things we get out of `population`. Read and run each cell.
# The third element in the array is the population
# in 1952.
population_1952 = population.item(2)
population_1952
# The thirteenth element in the array is the population
# in 1962 (which is 1950 + 12).
population_1962 = population.item(12)
population_1962
# The 66th element is the population in 2015.
population_2015 = population.item(65)
population_2015
# The array has only 66 elements, so this doesn't work.
# (There's no element with 66 other elements before it.)
population_2016 = population.item(66)
population_2016
# Since make_array returns an array, we can call .item(3)
# on its output to get its 4th element, just like we
# "chained" together calls to the method "replace" earlier.
make_array(-1, -3, 4, -2).item(3)
# **Question 4.2.1.** Set `population_1973` to the world population in 1973, by getting the appropriate element from `population` using `item`.
population_1973 = ...
population_1973
_ = ok.grade('q421')
# ## 4.3. Doing something to every element of an array
# Arrays are primarily useful for doing the same operation many times, so we don't often have to use `.item` and work with single elements.
#
# ##### Logarithms
# Here is one simple question we might ask about world population:
#
# > How big was the population in *orders of magnitude* in each year?
#
# The logarithm function is one way of measuring how big a number is. The logarithm (base 10) of a number increases by 1 every time we multiply the number by 10. It's like a measure of how many decimal digits the number has, or how big it is in orders of magnitude.
#
# We could try to answer our question like this, using the `log10` function from the `math` module and the `item` method you just saw:
# +
import math
population_1950_magnitude = math.log10(population.item(0))
population_1951_magnitude = math.log10(population.item(1))
population_1952_magnitude = math.log10(population.item(2))
population_1953_magnitude = math.log10(population.item(3))
...
# -
# But this is tedious and doesn't really take advantage of the fact that we are using a computer.
#
# Instead, NumPy provides its own version of `log10` that takes the logarithm of each element of an array. It takes a single array of numbers as its argument. It returns an array of the same length, where the first element of the result is the logarithm of the first element of the argument, and so on.
#
# **Question 4.3.1.** Use it to compute the logarithms of the world population in every year. Give the result (an array of 66 numbers) the name `population_magnitudes`. Your code should be very short.
population_magnitudes = ...
population_magnitudes
_ = ok.grade('q431')
# <img src="array_logarithm.jpg">
#
# This is called *elementwise* application of the function, since it operates separately on each element of the array it's called on. The textbook's section on arrays has a useful list of NumPy functions that are designed to work elementwise, like `np.log10`.
#
# ##### Arithmetic
# Arithmetic also works elementwise on arrays. For example, you can divide all the population numbers by 1 billion to get numbers in billions:
population_in_billions = population / 1000000000
population_in_billions
# You can do the same with addition, subtraction, multiplication, and exponentiation (`**`). For example, you can calculate a tip on several restaurant bills at once (in this case just 3):
restaurant_bills = make_array(20.12, 39.90, 31.01)
print("Restaurant bills:\t", restaurant_bills)
tips = .2 * restaurant_bills
print("Tips:\t\t\t", tips)
# <img src="array_multiplication.jpg">
#
# **Question 4.3.2.** Suppose the total charge at a restaurant is the original bill plus the tip. That means we can multiply the original bill by 1.2 to get the total charge. Compute the total charge for each bill in `restaurant_bills`.
total_charges = ...
total_charges
_ = ok.grade('q432')
# **Question 4.3.3.** `more_restaurant_bills.csv` contains 100,000 bills! Compute the total charge for each one. How is your code different?
more_restaurant_bills = Table.read_table("more_restaurant_bills.csv").column("Bill")
more_total_charges = ...
more_total_charges
_ = ok.grade('q433')
# The function `sum` takes a single array of numbers as its argument. It returns the sum of all the numbers in that array (so it returns a single number, not an array).
#
# **Question 4.3.4.** What was the sum of all the bills in `more_restaurant_bills`, *including tips*?
sum_of_bills = ...
sum_of_bills
_ = ok.grade('q434')
# **Question 4.3.5.** The powers of 2 ($2^0 = 1$, $2^1 = 2$, $2^2 = 4$, etc) arise frequently in computer science. (For example, you may have noticed that storage on smartphones or USBs come in powers of 2, like 16 GB, 32 GB, or 64 GB.) Use `np.arange` and the exponentiation operator `**` to compute the first 30 powers of 2, starting from `2^0`.
powers_of_2 = ...
powers_of_2
_ = ok.grade('q435')
# ## 4.4 Example: Growth Rates
# A natural example of how we can use arrays to reduce large amounts of computation is growth rates.
#
# **Question 4.4.1** Let's say we are investing in stocks, and we initially invest 10.23 dollars into the market. We check back in one year later, and we see that our total money in the market is now 14.32 dollars. What was our anual growth rate?
annual_growth_rate = ...
annual_growth_rate
_ = ok.grade('q441')
# **Question 4.4.2** If we wanted to see multiple people's annual stock growth rates, we could continue the above process per person. However, this can become tedious.
#
# Let's use the power of arrays! Assume that `initials` contains the initial amount of money for 5 different people, and `changed` contains the amount of money after one year for the same corresponding people. Assign `annual_growth_rates` to an array of all of the different growth rates for the 5 people.
initials = make_array(10.21, 11.32, 15.21, 13.22, 19.10)
changed = make_array(14.20, 35.44, 10.43, 9.62, 20.10)
annual_growth_rates = ...
annual_growth_rates
_ = ok.grade('q442')
# **Question 4.4.3** Now, let's use an array arithmetic to deduce the annual growth rate on peoples stocks given the amount of money in their market 10 years from now, found in the variable `ten_years`. Assuming everyone initially started with 10 dollars in their market, calculate the annual growth rate per person over these 10 years and assign this array of values to `annual_rates_over_ten_years`.
#
# *Hint*: If you don't remember this formula, check out the textbook!
ten_years = make_array(50.32, 1.04, 0.40, 14.50, 11.12)
annual_rates_over_ten_years = ...
annual_rates_over_ten_years
_ = ok.grade('q443')
# **Question 4.4.4** Lastly, let's use array arithmetic to figure the final amount of money in people's market 10 years from now, assuming they all invested different amounts of money (`invested`) in the same stock, INT5. The annual growth rate for INT5 was .045. Assign `money_in_ten_years` to an array of the money people ended with in the INT5 stock based on how much they initially invested.
#
# *Hint*: If you don't remember this formula, check out the textbook!
invested = make_array(10,11,15,20,25)
money_in_ten_years = ...
money_in_ten_years
_ = ok.grade('q444')
# Congratulations, you're done with lab 2!
#
# Before you submit the notebook, make sure that you
# * select from the top menu `Save and Checkpoint` from the `File` menu
# * `Kernel -> Restart & Clear Output`
# * `Cell -> Run All`
#
# Verify that all computations execute correctly. You should see **no errors**; if there are any errors, make sure to correct them before you submit the notebook.
# For your convenience, you can run this cell to run all the tests at once!
import os
_ = [ok.grade(q[:-3]) for q in os.listdir("tests") if q.startswith('q')]
_ = ok.submit()
|
lab02/lab02.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 計算 sin(x)
import numpy as np
import matplotlib.pyplot as plt
x = float(input('x (0.0 <= x <= 2PI) ='))
list1 = []
for k in range(1,15):
sum = 0
for i in range(k):
factorial = 1.0
for j in range(2*i + 1):
factorial *= j + 1
sum += ( ( (-1)**i) * (x**(2*i+1) ) ) / factorial
list1.append(sum)
len(list1)
b = [i for i in range(1,15)]
len(b)
plt.plot(b,list1)
plt.show()
# ## 相比numpy裡的sin(x)套件
print("sin(x) = " , np.sin(x))
|
NCCU Applications of mathematics softwares/lesson_6_4.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <center><i><img src="https://www.python.org/static/apple-touch-icon-72x72-precomposed.png"></i></center>
# ## <center>Script dove l'utente inserisce i minuti come un intero e lo script riavvia il Pc o il Server dopo gli x minuti<i><br><br><font color="blue">AlanTurista</font></i></center>
# ## Main
# +
import countdown_def
import restart_def
print("\n\n\t~ Script per riavvio Pc o Server ~\n\n@Author: <NAME>, <EMAIL>\n")
print('*'*60)
#Give time to restart in minutes
x = int(input("\n\tInserire minuti: "))
t = x*60
print("\n\n\tIl calcolatore verrà riavviato in:\n")
countdown_def.countdown(t)
restart_def.restart()
# -
# ## Countdown
# +
import time
def countdown(t):
while t > 0:
m, s = divmod(t, 60)
h, m = divmod(m, 60)
time_left = str(h).zfill(2) + ":" + str(m).zfill(2) + ":" + str(s).zfill(2)
print(time_left + "\r", end="")
time.sleep(1)
t -= 1
# -
# ## Restart
# +
import os
def restart():
os.system("shutdown -g -t 0")
|
IT/restart_server.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + code_folding=[0]
# Standard Imports and Extensions
# %matplotlib inline
# %config InlineBackend.figure_format='retina' # Comment out if not on Mac
import matplotlib as mpl
mpl.rcParams['figure.figsize'] = (16.0, 10.0)
# from IPython.display import set_matplotlib_formats
# set_matplotlib_formats('retina')
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
# In my python 2.7 environment, this is not set up by default - fix this.
if np.datetime64 not in mpl.units.registry:
from pandas.tseries import converter as pdtc
pdtc.register()
# + code_folding=[0]
# Simple STL plotting utility
def stl_plot(stl):
"""
Plot components of STL decomposition using matplotlib
:param stl_decomposition: pandas.DataFrame with data, trend, seasonal and residual columns
"""
fig, axes = plt.subplots(4, 1, sharex=True)
axes[0].set_ylabel("observations")
axes[1].set_ylabel("trend")
axes[2].set_ylabel("seasonal")
axes[3].set_ylabel("residual")
_ = axes[0].plot(stl.data)
_ = axes[1].plot(stl.trend)
_ = axes[2].plot(stl.seasonal)
_ = axes[3].plot(stl.residual)
# +
columns = ["data", "seasonal", "trend", "residual"]
time = pd.date_range('3/1/1958 00:00:00', periods=708, freq='M')
stl_java = pd.read_csv("output.csv", header=None)
stl_java.columns = columns
stl_java.index = time
stl_fortran = pd.read_csv("fortran_benchmark/output.csv", header=None)
stl_fortran.columns = columns
stl_fortran.index = time
# -
stl_plot(stl_java)
stl_plot(stl_fortran)
_ = plt.semilogy(np.abs(stl_java.trend - stl_fortran.trend), "bx")
_ = plt.semilogy(np.abs(stl_java.seasonal - stl_fortran.seasonal), "g+")
|
examples/StlPerfTest/StlJavaFortranComparison.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="wd-PuFkHXHAW" colab_type="text"
# Code from https://www.inferentialthinking.com/chapters/17/Classification.html was not running.
#
# Code updated by <NAME>
#
# Date: 12/17/2019
# + id="jEo4WNhrXGRg" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 459} outputId="f53756fa-5d0d-43c3-ed8a-37933991baeb"
import numpy as np
from datascience import *
path_data = 'https://github.com/data-8/textbook/raw/gh-pages/data/'
data = Table.read_table(path_data + 'ckd.csv')
ckd = data.relabeled('Blood Glucose Random', 'Glucose')
ckd
# + id="pLnv7MyAQ-mJ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 388} outputId="26922de1-0d95-425e-d9fe-2435335b12ac"
def standard_units(x):
return (x - np.mean(x))/np.std(x)
ckd = Table().with_columns(
'Hemoglobin', standard_units(ckd.column('Hemoglobin')),
'Glucose', standard_units(ckd.column('Glucose')),
'White Blood Cell Count',
standard_units(ckd.column('White Blood Cell Count')),
'Class', ckd.column('Class'))
ckd
# + id="GsjtHVw8Q-o4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 388} outputId="d3ec7331-38d4-422a-e6a7-e4e59195009a"
color_table = Table().with_columns(
'Class', make_array(1, 0),
'Color', make_array('darkblue', 'gold'))
ckd = ckd.join('Class', color_table)
ckd
# + id="fKCa7Ps9Q-rp" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 359} outputId="10b9e2d4-2446-4f9c-edbb-335bd3722e13"
# %matplotlib inline
import matplotlib.pyplot as plt
plt.style.use('fivethirtyeight')
ckd.scatter('Hemoglobin', 'Glucose', colors='Color')
# + id="smZhIefYQ-uZ" colab_type="code" colab={}
def distance(point1, point2):
"""The distance between two arrays of numbers."""
return np.sqrt(np.sum((point1 - point2)**2))
def all_distances(training, point):
"""The distance between p (an array of numbers)
and the numbers in row i of attribute_table."""
attributes = training.drop('Class')
def distance_from_point(row):
return distance(point, np.array(row))
return attributes.apply(distance_from_point)
def table_with_distances(training, point):
"""A copy of the training table with
the distance from each row to array p."""
return training.with_column('Distance',
all_distances(training, point))
def closest(training, point, k):
"""A table containing the k closest
rows in the training table to array p."""
with_dists = table_with_distances(training, point)
sorted_by_distance = with_dists.sort('Distance')
topk = sorted_by_distance.take(np.arange(k))
return topk
# + id="Vj6xVOKPQ-w5" colab_type="code" colab={}
def show_closest(point):
"""point = array([x,y])
gives the coordinates of a new point
shown in red"""
HemoGl = ckd.drop('White Blood Cell Count', 'Color')
t = closest(HemoGl, point, 1)
x_closest = t.row(0).item(1)
y_closest = t.row(0).item(2)
ckd.scatter('Hemoglobin', 'Glucose', colors='Color')
plt.scatter(point.item(0), point.item(1), color='red', s=30)
plt.plot(make_array(point.item(0), x_closest),
make_array(point.item(1), y_closest), color='k', lw=2);
# + id="M3V9ac7PQ-zh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 359} outputId="b36f4de7-b3fa-4d95-8808-616592fa2f8e"
# In this example, Alice's Hemoglobin
# attribute is 0 and her Glucose is 1.5.
alice = make_array(0, 1.5)
show_closest(alice)
# + id="NZI2aePIQ-2K" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 359} outputId="2a2d7824-ae2d-4cc0-88a4-f3364b3d00db"
alice = make_array(0, 0.97)
show_closest(alice)
# + id="EWoQiP6gXxEo" colab_type="code" colab={}
x_array = make_array()
y_array = make_array()
for x in np.arange(-2, 2.1, 0.1):
for y in np.arange(-2, 2.1, 0.1):
x_array = np.append(x_array, x)
y_array = np.append(y_array, y)
test_grid = Table().with_columns(
'Hemoglobin', x_array,
'Glucose', y_array)
# + id="xgN3clqGXxHh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 365} outputId="645bb5cd-89f5-4d19-d5a5-6b4768309130"
test_grid.scatter('Hemoglobin', 'Glucose',
color='red', alpha=0.4, s=30)
plt.scatter(ckd.column('Hemoglobin'), ckd.column('Glucose'),
c=ckd.column('Color'), edgecolor='k')
plt.xlim(-2, 2)
plt.ylim(-2, 2);
# + id="pmzKUPocXxKB" colab_type="code" colab={}
def majority(topkclasses):
"""1 if the majority of the "Class" column is 1s, and 0 otherwise."""
ones = topkclasses.where('Class', are.equal_to(1)).num_rows
zeros = topkclasses.where('Class', are.equal_to(0)).num_rows
if ones > zeros:
return 1
else:
return 0
def classify(training, p, k):
"""Classify an example with attributes p using k-nearest
neighbor classification with the given training table."""
closestk = closest(training, p, k)
topkclasses = closestk.select('Class')
return majority(topkclasses)
def classify_grid(training, test, k):
c = make_array()
for i in range(test.num_rows):
c = np.append(c, classify(training,
make_array(test.row(i)), k))
return c
# + id="pXhgy8K3XxO_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 365} outputId="62e8e4bb-2625-405b-af00-67e469b77cb4"
c = classify_grid(ckd.drop('White Blood Cell Count',
'Color'), test_grid, 1)
test_grid = test_grid.with_column('Class',
c).join('Class', color_table)
test_grid.scatter('Hemoglobin', 'Glucose',
colors='Color', alpha=0.4, s=30)
plt.scatter(ckd.column('Hemoglobin'), ckd.column('Glucose'),
c=ckd.column('Color'), edgecolor='k')
plt.xlim(-2, 2)
plt.ylim(-2, 2);
# + id="76TdMXBwXxRn" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 359} outputId="0900da67-0472-4c34-fba4-0b4babf14ec1"
ckd.scatter('White Blood Cell Count',
'Glucose', colors='Color')
# + id="VlluK4xZXxUN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 365} outputId="4ab04061-3bdb-47b5-c879-03e80d1ba608"
shuffled_ckd = ckd.sample(with_replacement=False)
training = shuffled_ckd.take(np.arange(79))
testing = shuffled_ckd.take(np.arange(79, 158))
training.scatter('White Blood Cell Count',
'Glucose', colors='Color')
plt.xlim(-2, 6)
plt.ylim(-2, 6);
# + id="V5ExsRIkZTpY" colab_type="code" colab={}
x_array = make_array()
y_array = make_array()
for x in np.arange(-2, 6.1, 0.25):
for y in np.arange(-2, 6.1, 0.25):
x_array = np.append(x_array, x)
y_array = np.append(y_array, y)
test_grid = Table().with_columns(
'Glucose', x_array,
'White Blood Cell Count', y_array)
c = classify_grid(training.drop('Hemoglobin',
'Color'), test_grid, 1)
# + id="4yLTy870ZTsI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 365} outputId="f72076ee-c702-4209-87f6-ef78014d698c"
test_grid = test_grid.with_column('Class',
c).join('Class', color_table)
test_grid.scatter('White Blood Cell Count',
'Glucose', colors='Color', alpha=0.4, s=30)
plt.xlim(-2, 6)
plt.ylim(-2, 6);
# + id="Ys1sCOVnZTvI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 365} outputId="d2c4a3af-c05d-4f3a-e1dc-8cd22a86ad80"
test_grid = test_grid.with_column('Class',
c).join('Class', color_table)
test_grid.scatter('White Blood Cell Count',
'Glucose', colors='Color', alpha=0.4, s=30)
plt.scatter(testing.column('White Blood Cell Count'),
testing.column('Glucose'), c=testing.column('Color'),
edgecolor='k')
plt.xlim(-2, 6)
plt.ylim(-2, 6);
# + id="6CnX-4qTZTx2" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="ac2127cb-b0c9-48b8-daf5-00709fb73a93"
ckd.row(0)
# + id="La2rrBQbZT0e" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="96450330-fb5d-4ab2-9840-4b93d12079ee"
ckd.row(0).item(1)
# + id="ItOCzqF8agG4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="04fd5b65-0dae-45e0-fee6-e9e8914705af"
ckd_attributes = ckd.select('Hemoglobin', 'Glucose')
ckd_attributes.row(3)
# + id="yMufUnK-agJ0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="fdf19af3-9724-4723-ef75-738008297488"
patient3 = np.array(ckd_attributes.row(3))
alice = make_array(0, 1.1)
alice, patient3
# + id="Ti3HqgnIagMr" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="f068cfa9-7cfd-493b-de2d-506160476deb"
distance = np.sqrt(np.sum((alice - patient3)**2))
distance
# + id="-9rDygOqb5dG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="949bb905-4811-4020-9f5b-02de8517c199"
t = ckd_attributes.take(np.arange(5))
t
# + id="VPqrmtAcb5k-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="8bb1e4e2-379b-4636-c2c2-551ed3166bf4"
def max_abs(row):
return np.max(np.abs(np.array(row)))
max_abs(t.row(4))
# + id="TApRb7qmb5oP" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="fb6ee957-1436-44ca-8042-f30b399fcf6b"
t.apply(max_abs)
# + id="89-UEAm1jc7l" colab_type="code" colab={}
def distance(point1, point2):
return np.sqrt(np.sum((point1 - point2)**2))
# + id="rD-sGU6TagPW" colab_type="code" colab={}
def distance_from_alice(row):
"""Returns distance between Alice and a row of the attributes table"""
return distance(alice, np.array(row))
# + id="ptOI6Lrmbpvx" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="52d5f8cf-8b80-4d38-a6a1-86bc7b8f2da5"
distance_from_alice(ckd_attributes.row(3))
# + id="8PBTK0k3bp1f" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 388} outputId="a3b602e6-3ef9-4f01-c124-9a735208ef27"
distances = ckd_attributes.apply(distance_from_alice)
ckd_with_distances = ckd.with_column('Distance from Alice', distances)
ckd_with_distances
# + id="x8BFGI_5bp4A" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 388} outputId="3f218a54-727c-4299-8f99-21db8dbffba9"
sorted_by_distance = ckd_with_distances.sort('Distance from Alice')
sorted_by_distance
# + id="3O6KUOnaj7uU" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="305eeb19-89e1-4ee2-d0b9-617d77ab4397"
alice_5_nearest_neighbors = sorted_by_distance.take(np.arange(5))
alice_5_nearest_neighbors
# + id="5mOGlJ7sj7xJ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 520} outputId="41790753-019a-4c5d-8446-0a6b391ad05f"
import matplotlib.pyplot as plots
plots.figure(figsize=(8,8))
plots.scatter(ckd.column('Hemoglobin'),
ckd.column('Glucose'), c=ckd.column('Color'), s=40)
plots.scatter(alice.item(0), alice.item(1), color='red', s=40)
radius = sorted_by_distance.column('Distance from Alice').item(4)+0.014
theta = np.arange(0, 2*np.pi+1, 2*np.pi/200)
plots.plot(radius*np.cos(theta)+alice.item(0),
radius*np.sin(theta)+alice.item(1), color='g', lw=1.5);
plots.xlim(-2, 2.5)
plots.ylim(-2, 2.5);
|
Slides/Winter 2020/16) K-Nearest Neighbors/17_1_Nearest_Neighbors.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <small><small><i>
# All the IPython Notebooks in **Python Introduction** lecture series by Dr. <NAME> are available @ **[GitHub](https://github.com/milaan9/01_Python_Introduction)**
# </i></small></small>
# # Python Type Conversion and Type Casting
#
# In this class, you will learn about the Type conversion and uses of type conversion.
#
# Before learning Type Conversion in Python, you should have knowledge about **[Python Data Types](https://github.com/milaan9/01_Python_Introduction/blob/main/009_Python_Data_Types.ipynb)**.
# ## Type Conversion
#
# The process of converting the value of one data type (integer, string, float, etc.) to another data type is called type conversion. Python has two types of type conversion.
#
# 1. **Implicit Type Conversion**
# 2. **Explicit Type Conversion**
# ### 1. Implicit Type Conversion
#
# In Implicit type conversion, Python automatically converts one data type to another data type. This process doesn't need any user involvement.
#
# Let's see an example where Python promotes the conversion of the lower data type (integer) to the higher data type (float) to avoid data loss.
# +
# Example 1: Converting integer to float
num_int = 123
num_flo = 1.23
num_new = num_int + num_flo
print("datatype of num_int:",type(num_int))
print("datatype of num_flo:",type(num_flo))
print("Value of num_new:",num_new)
print("datatype of num_new:",type(num_new))
# -
# **Explanation**:
#
# In the above program,
#
# 1. We add two variables **`num_int`** and **`num_flo`**, storing the value in **`num_new`**.
# 2. We will look at the data type of all three objects respectively.
# 3. In the output, we can see the data type of **`num_int`** is an **`integer`** while the data type of **`num_flo`** is a **`float`**.
# 4. Also, we can see the **`num_new`** has a **`float`** data type because Python always converts smaller data types to larger data types to avoid the loss of data.
# Now, let's try adding a string and an integer, and see how Python deals with it.
# +
# Example 2: Addition of string(higher) data type and integer(lower) datatype
num_int = 123
num_str = "456"
print("Data type of num_int:",type(num_int))
print("Data type of num_str:",type(num_str))
print(num_int+num_str) # ERROR! cannot add two different data types (int and string)
# -
# **Explanation**:
#
# In the above program,
#
# 1. We add two variables **`num_int`** and **`num_str`**.
# 2. As we can see from the output, we got **`TypeError`**. Python is not able to use Implicit Conversion in such conditions.
# 3. However, Python has a solution for these types of situations which is known as Explicit Conversion.
# ### 2. Explicit Type Conversion
#
# In Explicit Type Conversion, users convert the data type of an object to required data type. We use the predefined functions like **`int()`**, **`float()`**, **`str()`**, etc to perform explicit type conversion.
#
# This type of conversion is also called typecasting because the user casts (changes) the data type of the objects.
#
# **Syntax:**
#
# ```python
# <required_datatype>(expression)
# ```
#
# Typecasting can be done by assigning the required data type function to the expression.
# int to float
num_int = 10
print('num_int',num_int) # 10
num_float = float(num_int)
print('num_float:', num_float) # 10.0
# float to int
gravity = 9.81
print(int(gravity)) # 9
# +
# Example 3: Addition of string and integer using explicit conversion
num_int = 123
num_str = "456" # there is a number inside " " so I CAN change it to integer
#num_str = "jason" # there is a name inside " " so I cannot change to integer
print("Data type of num_int:",type(num_int))
print("Data type of num_str before Type Casting:",type(num_str))
num_str = int(num_str) # change from "string" data type to integer datatype
print("Data type of num_str after Type Casting:",type(num_str))
num_sum = num_int + num_str # Now we can add two same data types (int and int)
print("Sum of num_int and num_str:",num_sum)
print("Data type of the sum:",type(num_sum))
# -
# **Explanation**:
#
# In the above program,
#
# 1. We add **`num_str`** and **`num_int`** variable.
# 2. We converted **`num_str`** from string(higher) to integer(lower) type using **`int()`** function to perform the addition.
# 3. After converting **`num_str`** to an integer value, Python is able to add these two variables.
# 4. We got the **`num_sum`** value and data type to be an integer.
float(6) # means converting interger 6 to a float value
# Conversion from **`float`** to **`int`** will truncate the value (make it closer to zero).
# +
int(33.6) # means converting float 33.6 to a integer value
# When you change from FLOAT to INTEGER it will round-up the number
# -
int(-33.6)
# Conversion to and from **`string`** must contain compatible values.
# int to str
num_int = 10
print(num_int) # 10
num_str = str(num_int)
print(num_str) # '10'
float('7.5') # means converting string '7.5' to a float value
# str to int or float
num_str = '10.6' #or '10'
# print('num_int', int(num_str)) # 10
print('num_float', float(num_str)) # 10.6
str(65) # means converting integer 65 to a string value
int('1p') # means converting string '1p' to a integer value
str("1p")
# We can even convert one sequence to another.
set([1,2,3]) # [1,2,3] is tuple and now converting to a set {}
tuple({5,6,7}) # {1,2,3} is set and now converting to a tuple ()
list('hello') # ("hello") is string and now converting to a list []
# str to list
first_name = 'Milaan'
print(first_name) # 'Milaan'
first_name_to_list = list(first_name)
print(first_name_to_list) # ['M', 'i', 'l', 'a', 'a', 'n']
# To convert to dictionary, each element must be a pair:
dict([[1,2],[3,4]]) # [[1,2],[3,4]] is tuple and now converting to a dictionary
dict([(3,63),(7,91)]) # [(3,63),(7,91)] is tuple and now converting to a dictionary
# ## Key Points to Remember
#
# 1. Type Conversion is the conversion of object from one data type to another data type.
# 2. Implicit Type Conversion is automatically performed by the Python interpreter.
# 3. Python avoids the loss of data in Implicit Type Conversion.
# 4. Explicit Type Conversion is also called Type Casting, the data types of objects are converted using predefined functions by the user.
# 5. In Type Casting, loss of data may occur as we enforce the object to a specific data type.
|
010_Python_Type_Conversion.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Collect data
# 1. Find yourself an data set that you want to analyse, this dataset has to be an csv file (excel), we are going to use this dataset in the coming weeks to give you real working experience.
#
# - The dataset can be from your own work of you can explore https://www.kaggle.com/datasets
# - Look for dataset with not to much columns
#
#
# 2. Write at least 3 questions down, you want to know from the data set
# - Do not go to hard on yourself
#
# # Explain concepts
#
# Because we all know how hard it is to explain concepts, that's why we need to practise. You do not have to type things done, just try to explain the concept to someone you know. Of course you can google first
# 1. Explain why we use the print statement
# 2. Explain why for loops are handy
# 3. What is the difference between a set, a tuple and a list
# 4. Why do we need to use comments and how do you use them best
#
|
homework/Week1-Assignment.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: furniture
# language: python
# name: furniture
# ---
# #### imports
1
# +
from annoy import AnnoyIndex
from fastai.vision.all import *
from mytools.tools import *
from fur.paths import *
from fur.learn import *
from IPython.display import Image,display
import requests
# -
model_path = models_path/'model_sofas_styles__color_singlelabel_stable'
# +
# model_path =models_path/path_info(models_path).sort_values(['time'])['name'].iloc[-1];model_path
# -
model_path.ls()
len(model_path.ls())
# +
learn = load_learner(model_path/'model.pkl')
sf = SaveFeatures(learn.model[1][7])
t = AnnoyIndex(512, 'euclidean')
t.load(str(model_path/'test.ann'))
feature_dict = from_pickle(model_path/'feature_dict.pkl')
# -
t.get_n_items()
models_path
len(feature_dict)
# #### predict
pred_item_url = 'https://bekolmebel.ru/sites/default/files/cb947cbfd550a4d60f19.jpg'
save_file_from_url(pred_item_url, 'pred.jpg')
Image(filename ='pred.jpg', width=500, height=500)
pred = learn.predict('pred.jpg');pred[0]
len(sf.features)
similar = t.get_nns_by_vector(sf.features[-1], 5, include_distances=False)
similar
for i in similar:
display(Image(filename =ssd_pictures_path/list(feature_dict.keys())[i], width=500, height=500))
|
.ipynb_checkpoints/presentation-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:gsoc_ensembl]
# language: python
# name: conda-env-gsoc_ensembl-py
# ---
# +
####################################################################################################
# Copyright 2019 <NAME> and EMBL-European Bioinformatics Institute
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
# http://www.apache.org/licenses/LICENSE-2.0
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
####################################################################################################
# -
# # Approach:-
# 1. Get all lncRNA transcript IDs.
# 2. Use _rna fasta file.
# 3. Get sequence of all those whose transcript IDs lies in fasta file.
from tqdm import tqdm
import csv
import pandas as pd
import numpy as np
path = '/all_type=lncRNA_data/all_lncrna_transcript_data_corrected.csv'
df = pd.read_csv(path)
transcript_ids = df['RefSeq Transcript ID'].tolist()
trans_len = df['Transcript Length'].tolist()
parent = df['Parent'].tolist()
print(len(transcript_ids))
print(len(trans_len))
ncrna_ids = []
for i in range(len(transcript_ids)):
ncrna_ids.append(transcript_ids[i][4:])
len(ncrna_ids)
ncrna_ids[0:10]
# +
# %%time
f12 = '/fasta/GRCh38_p12/GCF_000001405.38_GRCh38.p12_rna.fna'
from Bio import SeqIO
records12 = list(SeqIO.parse(f12, "fasta"))
# -
records12_ids = []
records12_sequences = []
for i in range(len(records12)):
records12_ids.append(records12[i].id)
records12_sequences.append(records12[i].seq)
len(records12_ids)
len(records12)
len(records12_sequences)
# +
# str(records12_sequences[10000])
# -
ncrna_sequences = []
for i in range(len(ncrna_ids)):
ncrna_sequences.append('NA')
print(len(ncrna_sequences))
# ## Below - Extracting IDs and their sequences, from fasta file, in a list (for simplicity)
lncrna_records12_index = []
s = set(ncrna_ids)
# records12_lncrna_ids = []
c = 0
for i in tqdm(range(len(records12_ids))):
if records12_ids[i] in s:
lncrna_records12_index.append(i)
c = c + 1
# records12_lncrna_ids.append
c
len(lncrna_records12_index)
index_ncrna_records12 = []
index_ncrna_ids = []
# +
from tqdm import tqdm
def get_lncrna_seq():
for i in tqdm(lncrna_records12_index):
# for i in tqdm(range(len(records12_ids))):
for j in range(len(ncrna_ids)):
if records12_ids[i] == ncrna_ids[j]:
ncrna_sequences[j] = str(records12_sequences[i])
index_ncrna_ids.append(j)
index_ncrna_records12.append(i)
# ncrna_sequences.append(str(records12_sequences[i]))
else:
continue
# -
# %%time
get_lncrna_seq()
(len(ncrna_sequences))
# +
# ncrna_sequences[0:5]
# -
c = 0
for i in range(len(ncrna_sequences)):
if ncrna_sequences[i] != 'NA':
c = c + 1
print(c)
print(len(index_ncrna_ids))
seq_len = []
for i in range(len(ncrna_sequences)):
if ncrna_sequences[i] != 'NA':
seq_len.append(len(ncrna_sequences[i]))
else:
seq_len.append('NA')
s = set(trans_len)
c = 0
for i in range(len(seq_len)):
if seq_len[i] == 'NA':
continue
elif seq_len[i] == trans_len[i]:
c = c + 1
print(c) #sequence length which match with trans_len
len(len_match)
# %%time
len_match = []
c = 0
na = 0
difference = []
for i in tqdm(range(len(seq_len))):
if seq_len[i] == 'NA':
len_match.append('NA')
na = na + 1
difference.append('NA')
elif seq_len[i] == trans_len[i]:
len_match.append('YES')
difference.append(abs(trans_len[i]-seq_len[i]))
elif seq_len[i] != trans_len[i]:
len_match.append('NO')
difference.append(abs(trans_len[i]-seq_len[i]))
c = c + 1
print(len(difference))
# +
import pandas as pd
df = pd.read_csv('/all_type=lncRNA_data/all_lncrna_transcript_data_corrected.csv', index_col=0)
df['Length of Sequences'] = seq_len
df['| Seq_len - Trans_len |'] = difference
df['Match ?'] = len_match
df['Sequences'] = ncrna_sequences
# -
df.head(5)
df.to_csv('/all_type=lncRNA_data/all_lncrna_transcript_data_with_sequences_corrected.csv')
c = 0
no_index = []
for i in range(len(len_match)):
if len_match[i] == 'NO':
c = c + 1
no_index.append(i)
c
no_difference = []
for i in (no_index):
no_difference.append(difference[i])
len(no_difference)
max(no_difference)
max_difference_index = []
for i in no_index:
if difference[i] == 159:
max_difference_index.append(i)
break
max_difference_index
difference[19823]
df['RefSeq Transcript ID'][19823]
df['Index'][19823]
df.iloc[[19823]]
c = 0
na = 0
for i in range(len(len_match)):
if difference[i] == 0:
c = c + 1
elif difference[i] == 'NA':
na = na + 1
|
RefSeq-analysis/data_acquisition/genome_sequence_data/python_scripts/get_refseq_lncrna_seq.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys
import warnings
if not sys.warnoptions:
warnings.simplefilter('ignore')
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import pandas as pd
from sklearn.preprocessing import MinMaxScaler
from datetime import datetime
from datetime import timedelta
from tqdm import tqdm
sns.set()
tf.compat.v1.random.set_random_seed(1234)
df = pd.read_csv('../dataset/GOOG-year.csv')
df.head()
minmax = MinMaxScaler().fit(df.iloc[:, 4:5].astype('float32')) # Close index
df_log = minmax.transform(df.iloc[:, 4:5].astype('float32')) # Close index
df_log = pd.DataFrame(df_log)
df_log.head()
test_size = 30
simulation_size = 5
df_train = df_log.iloc[:-test_size]
df_test = df_log.iloc[-test_size:]
df.shape, df_train.shape, df_test.shape
class Model:
def __init__(
self,
learning_rate,
num_layers,
size,
size_layer,
output_size,
forget_bias = 0.1,
):
def lstm_cell(size_layer):
return tf.nn.rnn_cell.LSTMCell(size_layer, state_is_tuple = False)
backward_rnn_cells = tf.nn.rnn_cell.MultiRNNCell(
[lstm_cell(size_layer) for _ in range(num_layers)],
state_is_tuple = False,
)
forward_rnn_cells = tf.nn.rnn_cell.MultiRNNCell(
[lstm_cell(size_layer) for _ in range(num_layers)],
state_is_tuple = False,
)
self.X = tf.placeholder(tf.float32, (None, None, size))
self.Y = tf.placeholder(tf.float32, (None, output_size))
drop_backward = tf.contrib.rnn.DropoutWrapper(
backward_rnn_cells, output_keep_prob = forget_bias
)
forward_backward = tf.contrib.rnn.DropoutWrapper(
forward_rnn_cells, output_keep_prob = forget_bias
)
self.backward_hidden_layer = tf.placeholder(
tf.float32, shape = (None, num_layers * 2 * size_layer)
)
self.forward_hidden_layer = tf.placeholder(
tf.float32, shape = (None, num_layers * 2 * size_layer)
)
_, last_state = tf.nn.bidirectional_dynamic_rnn(
forward_backward,
drop_backward,
self.X,
initial_state_fw = self.forward_hidden_layer,
initial_state_bw = self.backward_hidden_layer,
dtype = tf.float32,
)
with tf.variable_scope('decoder', reuse = False):
backward_rnn_cells_decoder = tf.nn.rnn_cell.MultiRNNCell(
[lstm_cell(size_layer) for _ in range(num_layers)],
state_is_tuple = False,
)
forward_rnn_cells_decoder = tf.nn.rnn_cell.MultiRNNCell(
[lstm_cell(size_layer) for _ in range(num_layers)],
state_is_tuple = False,
)
drop_backward_decoder = tf.contrib.rnn.DropoutWrapper(
backward_rnn_cells_decoder, output_keep_prob = forget_bias
)
forward_backward_decoder = tf.contrib.rnn.DropoutWrapper(
forward_rnn_cells_decoder, output_keep_prob = forget_bias
)
self.outputs, self.last_state = tf.nn.bidirectional_dynamic_rnn(
forward_backward_decoder, drop_backward_decoder, self.X,
initial_state_fw = last_state[0],
initial_state_bw = last_state[1],
dtype = tf.float32
)
self.outputs = tf.concat(self.outputs, 2)
self.logits = tf.layers.dense(self.outputs[-1], output_size)
self.cost = tf.reduce_mean(tf.square(self.Y - self.logits))
self.optimizer = tf.train.AdamOptimizer(learning_rate).minimize(
self.cost
)
def calculate_accuracy(real, predict):
real = np.array(real) + 1
predict = np.array(predict) + 1
percentage = 1 - np.sqrt(np.mean(np.square((real - predict) / real)))
return percentage * 100
def anchor(signal, weight):
buffer = []
last = signal[0]
for i in signal:
smoothed_val = last * weight + (1 - weight) * i
buffer.append(smoothed_val)
last = smoothed_val
return buffer
num_layers = 1
size_layer = 128
timestamp = 5
epoch = 300
dropout_rate = 0.8
future_day = test_size
learning_rate = 0.01
def forecast():
tf.reset_default_graph()
modelnn = Model(
learning_rate, num_layers, df_log.shape[1], size_layer, df_log.shape[1], dropout_rate
)
sess = tf.InteractiveSession()
sess.run(tf.global_variables_initializer())
date_ori = pd.to_datetime(df.iloc[:, 0]).tolist()
pbar = tqdm(range(epoch), desc = 'train loop')
for i in pbar:
init_value_forward = np.zeros((1, num_layers * 2 * size_layer))
init_value_backward = np.zeros((1, num_layers * 2 * size_layer))
total_loss, total_acc = [], []
for k in range(0, df_train.shape[0] - 1, timestamp):
index = min(k + timestamp, df_train.shape[0] - 1)
batch_x = np.expand_dims(
df_train.iloc[k : index, :].values, axis = 0
)
batch_y = df_train.iloc[k + 1 : index + 1, :].values
logits, last_state, _, loss = sess.run(
[modelnn.logits, modelnn.last_state, modelnn.optimizer, modelnn.cost],
feed_dict = {
modelnn.X: batch_x,
modelnn.Y: batch_y,
modelnn.backward_hidden_layer: init_value_backward,
modelnn.forward_hidden_layer: init_value_forward,
},
)
init_value_forward = last_state[0]
init_value_backward = last_state[1]
total_loss.append(loss)
total_acc.append(calculate_accuracy(batch_y[:, 0], logits[:, 0]))
pbar.set_postfix(cost = np.mean(total_loss), acc = np.mean(total_acc))
future_day = test_size
output_predict = np.zeros((df_train.shape[0] + future_day, df_train.shape[1]))
output_predict[0] = df_train.iloc[0]
upper_b = (df_train.shape[0] // timestamp) * timestamp
init_value_forward = np.zeros((1, num_layers * 2 * size_layer))
init_value_backward = np.zeros((1, num_layers * 2 * size_layer))
for k in range(0, (df_train.shape[0] // timestamp) * timestamp, timestamp):
out_logits, last_state = sess.run(
[modelnn.logits, modelnn.last_state],
feed_dict = {
modelnn.X: np.expand_dims(
df_train.iloc[k : k + timestamp], axis = 0
),
modelnn.backward_hidden_layer: init_value_backward,
modelnn.forward_hidden_layer: init_value_forward,
},
)
init_value_forward = last_state[0]
init_value_backward = last_state[1]
output_predict[k + 1 : k + timestamp + 1] = out_logits
if upper_b != df_train.shape[0]:
out_logits, last_state = sess.run(
[modelnn.logits, modelnn.last_state],
feed_dict = {
modelnn.X: np.expand_dims(df_train.iloc[upper_b:], axis = 0),
modelnn.backward_hidden_layer: init_value_backward,
modelnn.forward_hidden_layer: init_value_forward,
},
)
output_predict[upper_b + 1 : df_train.shape[0] + 1] = out_logits
future_day -= 1
date_ori.append(date_ori[-1] + timedelta(days = 1))
init_value_forward = last_state[0]
init_value_backward = last_state[1]
for i in range(future_day):
o = output_predict[-future_day - timestamp + i:-future_day + i]
out_logits, last_state = sess.run(
[modelnn.logits, modelnn.last_state],
feed_dict = {
modelnn.X: np.expand_dims(o, axis = 0),
modelnn.backward_hidden_layer: init_value_backward,
modelnn.forward_hidden_layer: init_value_forward,
},
)
init_value_forward = last_state[0]
init_value_backward = last_state[1]
output_predict[-future_day + i] = out_logits[-1]
date_ori.append(date_ori[-1] + timedelta(days = 1))
output_predict = minmax.inverse_transform(output_predict)
deep_future = anchor(output_predict[:, 0], 0.3)
return deep_future[-test_size:]
results = []
for i in range(simulation_size):
print('simulation %d'%(i + 1))
results.append(forecast())
accuracies = [calculate_accuracy(df['Close'].iloc[-test_size:].values, r) for r in results]
# %matplotlib inline
plt.figure(figsize = (15, 5))
for no, r in enumerate(results):
plt.plot(r, label = 'forecast %d'%(no + 1))
plt.plot(df['Close'].iloc[-test_size:].values, label = 'true trend', c = 'black')
plt.legend()
plt.title('Bidirectional-Lstm-Seq2seq average accuracy: %.4f'%(np.mean(accuracies)))
plt.show()
# -
|
deep-learning/011bidirectional-lstm-seq2seq.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Density estimation ussing GMM
#
# This is an example of techniques for density estimation ussing mixtures of Gaussians. The example uses Python classes from Scikit-learn.
# +
# %matplotlib inline
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
import sklearn
from sklearn.neighbors import KernelDensity
from sklearn.mixture import GMM
from sklearn.mixture import DPGMM
from sklearn.mixture import VBGMM
import warnings
warnings.filterwarnings("ignore")
plt.rcParams['figure.figsize'] = (10,4)
colors = ['blue', 'red']
np.random.seed(42)
print(("Using the following versions\n"
"Numpy : {}\n"
"Scikit-learn : {}\n"
"Matplotlib : {}\n"
).format(np.__version__, sklearn.__version__,
matplotlib.__version__))
# -
# ## Dataset: toy example
#
# Given a dataset of independent and identically distributed observations $X = \{x_1, \dots, x_N\}$ comming from a mixture of $K$ Gaussians with unknown set of parameter values $\Theta = \{\theta_1, \dots, \theta_K\}$ where $\theta_i = \{\mu_i, \sigma_i\}$ and the latent variable $Z = \{z_1, \dots, z_N\}$ where $z_n \in \{1,\dots,K\}$ denoting which Gaussian has generated the observation $x_n$.
# +
# Hidden or latent variables
Z = {1: (-10.0, 2.0, 100),
2: (-5.0, 3.0, 90),
3: (0.0, 1.0, 100),
4: (5.0, 2.0, 150),
5: (9.0, 2.0, 200),
6: (15.0, 1.0, 100)}
Y = np.concatenate([np.ones(size)*y for y, (loc, scale, size) in Z.iteritems()]).reshape(-1,1)
# Observed variables
X = np.concatenate([np.random.normal(loc,scale,size) for loc, scale, size in Z.values()]).reshape(-1,1)
# A grid for the following density estimations
x_grid = np.linspace(X.min(), X.max(), 100).reshape(-1,1)
fig = plt.figure(figsize=(16,4))
ax1 = fig.add_subplot(121)
ax1.set_title('Sample distribution of X per component')
ax1.scatter(X, Y, c=Y)
ax1.scatter(X, np.zeros_like(X), marker='x', c='black', s=100, label='All (y=0)', alpha=0.1)
ax1.set_ylim([0,len(Z)+1])
ax1.set_ylabel('y')
ax1.set_xlabel('x')
ax1.legend(loc='upper left')
ax2 = fig.add_subplot(122)
ax2.set_title('Histogram of X')
hist = ax2.hist(X, bins=30)
ax2.set_ylabel('frequency')
ax2.set_xlabel('x')
# -
# ## Modeling the data
#
# We want to model the previous distribution with a mixture of Gaussians. In this example we will assume that we know the original number of Gaussians $K$.
#
# We want to find the set of parameter values $\Theta$ and the mixing coefficients $\pi = \{\pi_1, \dots, \pi_K\}$ that maximize the likelihood function
#
# $$
# p(X|\mu, \Sigma, \pi) = \prod_{n=1}^N \sum_{k=1}^K \pi_k \mathcal{N}(x_n|\mu_k, \Sigma_k)
# $$
# +
models = [(KernelDensity, {'kernel':'gaussian'},"Gaussian Kernel Density"),
(GMM, {'n_components':7},"Gaussian Mixture Model"),
(DPGMM, {'n_components':7},"Dirichlet Process GMM"),
(VBGMM, {'n_components':7},"Variational Bayes GMM")]
fig = plt.figure()
ax = fig.add_subplot(111)
for i, (model, args, name) in enumerate(models):
print("Computing {}".format(name))
kde = model(**args)
kde.fit(X)
if model in [KernelDensity]:
q_x = np.exp(kde.score_samples(x_grid))
else:
q_x = np.exp(kde.score(x_grid))
ax.plot(x_grid, q_x, label=name)
ax.legend(loc='upper left')
# -
|
jupyter/EM_Gaussian_mixture.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.8 64-bit (''base'': conda)'
# language: python
# name: python3
# ---
# +
import torch
import numpy as np
import matplotlib.pyplot as plt
plt.rcParams['figure.figsize'] = [10, 10]
import torch.nn as nn
from torch import tensor
# -
# ## Creating random data for linear model
# +
### y = 2*X + 1 + ε, where e is a noise
# -
# column vector X
X = torch.linspace(1, 50, 50).reshape(-1, 1)
X.shape
torch.manual_seed(71)
ε = torch.randint(-8, 9, (50,1), dtype=torch.float) # noise (bias)
ε.shape
y = 2*X + 1 + ε
y.shape
# matplotlib doesn't work with pytorch tensors so need to be converted to numpy
plt.figure(figsize=(10,10))
plt.scatter(X.numpy(), y.numpy());
# ---
# ## Build a simple linear model
# +
torch.manual_seed(59)
model = nn.Linear(in_features=1, out_features=1)
print(model.weight)
print(model.bias)
# -
# so from above it is y = 0.1060x + 0.9638
#
# nn.Linear is a neural network linear layer (just a linear function), not linear regression
# The same but other way - as pytorch convention:
class Model(nn.Module):
def __init__(self, in_features, out_features):
super().__init__()
self.linear = nn.Linear(in_features, out_features)
def forward(self, x):
y_pred = self.linear(x)
return y_pred
# +
# setting up a linear model for the problem of 1 input / 1 output with random weight and bias values
# here with manual seed for reproductivity
torch.manual_seed(59)
model = Model(1, 1)
print(model.linear.weight.item())
print(model.linear.bias.item())
# -
for name, param in model.named_parameters():
print(name, '\t', param.item())
X_try = torch.tensor([2.0])
print('y for X_try =', model.forward(X_try).item())
# Generating random values of x
x_1 = np.linspace(0.0, 50.0, 50)
x_1
# +
# by hand calculating y for random values from x_1 using initially generated weight and bias
weight_1 = 0.1059
bias_1 = 0.9637
y_1 = weight_1 * x_1 + bias_1
y_1
# -
plt.scatter(X.numpy(), y.numpy())
plt.plot(x_1, y_1, 'r');
# ## Loss
# Mean squared error to be used. 'criterion' is used as convention for loss funtion
criterion = nn.MSELoss()
# ## Optimizer
# Stochastic gradient descent
optimizer = torch.optim.SGD(model.parameters(), lr=1e-3)
# ## Train
# +
epochs = 50
losses = []
for i in range(epochs): # Loop over epochs
i += 1 # count epoch
y_pred = model.forward(X) # forward propagation
loss = criterion(y_pred, y) # calculate loss with chosen criterion loss
losses.append(loss.detach().numpy()) # append calculated loss value to tracking list
optimizer.zero_grad() # Resets all gradients (derivatives of losses) optimised previously by back propagation
loss.backward() # Does backpropagation
optimizer.step() # Updates hyperparameters of the model
print('loss:', losses[-1])
print('weight:', model.linear.weight.item(), '\nbias:', model.linear.bias.item())
# -
# Trained weights and bias give optimal loss for the problem. Both may be used for predictions now.
plt.plot(range(epochs), losses)
plt.ylabel('MSE Loss value')
plt.xlabel('Epoch');
# ## Check
# Predictions on random generated numbers
# +
x = np.linspace(0.0, 50.0, 50)
current_weight = model.linear.weight.item()
current_bias = model.linear.bias.item()
predicted_y = current_weight * x + current_bias
# -
predicted_y
# Ploting generated data and linear model with trained weight and bias
plt.scatter(X.numpy(), y.numpy())
plt.plot(x, predicted_y, 'r');
|
Courses/PyTorch for Deep Learning with Python/07 ANN/040_Linear_regression.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Creating the action client
# This time, we'll make the action client node for the action server.
# <t>
# Create a file called **demo_action_client.py** and copy the following into it:
# +
# #! /usr/bin/env python
import actionlib
import rospy
from action_tutorials.msg import DemoFeedback, DemoResult, DemoAction, DemoGoal
number = 1
def feedback_callback(feedback):
global number
print '[Feedback] number %d received' %number
number += 1
client = actionlib.SimpleActionClient('/demo_as', DemoAction)
rospy.init_node('demo_action_client')
client.wait_for_server()
while not rospy.is_shutdown():
goal = DemoGoal()
goal.count = input('Enter a number: ')
client.send_goal(goal, feedback_cb = feedback_callback)
client.wait_for_result()
print '[Result] State: %d'%(client.get_state())
print '---'
number = 1
# -
# Compile and run it after running **demo_action_server.py**
|
Modules/Module 6 - ROS Actions/4. Creating the ROS action client.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Task 5: Group Data Analysis
# ---
# In this notebook, we will be answering our research questions as a group with an in depth analysis.
#
# Some of our research questions were already approached within our individual EDA's, for example, if smokers typically have higher charges on average (they did), or if certain regions paid more on average (southeast paid marginally more than the other groups) Some other questions remain un answered, and that's what we are going to focus on for this group data analysis.
#
# The first two research questions we would like to answer are:
# - Is there any correlation between age and charges? Our group predict
#
|
analysis/Germaine/Group Data Analysis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ModSim project 1
# +
# Configure Jupyter so figures appear in the notebook
# %matplotlib inline
# Configure Jupyter to display the assigned value after an assignment
# %config InteractiveShell.ast_node_interactivity='last_expr_or_assign'
# import functions from the modsim.py module
from modsim import *
from pandas import read_csv
from math import exp
# -
table = read_csv('data.csv',index_col=0);
# +
plot(table.Population,':', label='Bee Population')
decorate(xlabel='Year',
ylabel='Bee population (Million)',
title='Bee population over time')
# -
population = table.Population
t_0 = 1947;
t_end = get_last_label(population);
p_0 = population[1947];
system = System(t_0=t_0, t_end=t_end, p_0=p_0, alpha = -0.01);
def update_func2(pop, t, system):
"""Compute the population next year.
pop: current population
t: current year
system: system object containing parameters of the model
returns: population next year
"""
net_growth = system.alpha * pop
return pop + net_growth
def run_simulation(system, update_func):
"""Simulate the system using any update function.
system: System object
update_func: function that computes the population next year
returns: TimeSeries
"""
results = TimeSeries()
results[system.t_0] = system.p_0
for t in linrange(system.t_0, system.t_end):
results[t+1] = update_func(results[t], t, system)
return results
def plot_results(population, timeseries, title):
"""Plot the estimates and the model.
census: TimeSeries of population estimates
un: TimeSeries of population estimates
timeseries: TimeSeries of simulation results
title: string
"""
plot(timeseries, color='gray', label='model')
plot(population,':', label='Bee Population')
decorate(xlabel='Year',
ylabel='Bee population (Million)',
title='Bee population over time')
results = run_simulation(system, update_func2)
plot_results(population, results, 'Quadratic model')
# Will bees go exticnt in the US?
#
# Since 1947 the number of Honey Bees colonies have been decreasing. One of the posible reasons for this sudden decline could be the increased use of insecticides in agrigulture. Honey Bees are a main natural pollinator that farmers acknowledge as good, but are inadvertly the subjects of the toxic insecticides.
# To explore this question we will develop a quadratic model that predicts the number of Honey Bee colonies in the United States.
# Specifically, we will add one new feature to the model
# 1. Insecticide Usage: The current model doesn't account for the change in insectide usage. We will use this feature to more accurately predict the number of bee colonies.
# We will begin our model with the data from 1947, which is when the number of colonies began to decrease.
|
code/Project1.2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
#
# ---
# # NumPy Data X - BKHW
#
# **Author:** <NAME>, <NAME>, <NAME> 1/22/2017, midified June 2017
#
# **License Agreement:** Feel free to do whatever you want with this code
#
# ___
# # Introduction to NumPy
# # What is NumPy:
#
# NumPy stands for Numerical Python and it is the fundamental package for scientific computing with Python. It is a package that lets you efficiently store and manipulate numerical arrays. It contains among other things:
#
# * a powerful N-dimensional array object
# * sophisticated (broadcasting) functions
# * tools for integrating C/C++ and Fortran code
# * useful linear algebra, Fourier transform, and random number capabilities
#
# # NumPy contains an array object that is "fast"
#
#
# <img src="https://github.com/ikhlaqsidhu/data-x/raw/master/imgsource/threefundamental.png">
#
#
# It stores:
# * location of a memory block (allocated all at one time)
# * a shape (3 x 3 or 1 x 9, etc)
# * data type / size of each element
#
# The core feauture that NumPy supports is its multi-dimensional arrays. In NumPy, dimensions are called axes and the number of axes is called a rank.
# written for Python 3.6
import numpy as np
#
# ## Creating a NumPy Array: -
# ### 1. Simplest possible: We use a list as an argument input in making a NumPy Array
#
# Create array from Python list
list1 = [1, 2, 3, 4]
data = np.array(list1)
data
# Find out object type
type(data)
# See data type that is stored in the array
data.dtype
# The data types are specified for the full array, if we store
# a float in an int array, the float will be up-casted to an int
data[0] = 3.14159
print(data)
data[0] = 0
# NumPy converts to most logical data type
list2 = [1.2, 2, 3, 4]
data2 = np.array(list2)
print(data2)
print(data2.dtype) # all values will be converted to floats if we have one
# We can manually specify the datatype
list3 = [1, 2, 3]
data3 = np.array(list3, dtype=str) #manually specify data type
print(data3)
print(data3.dtype)
# lists can also be much longer
list4 = range(100000)
data = np.array(list4)
data
len(data) # to see the length of the full array
# data = np.array(1,2,3,4, 5,6,7,8,9) # wrong
data = np.array([1,2,3,4,5,6,7,8,9]) # right
data
# +
# see documentation, the first keyword is the object to be passed in
# np.array?
# -
# More info on data types can be found here:
# https://docs.scipy.org/doc/numpy-1.13.0/user/basics.types.html
# # Accessing elements: Slicing and indexing
# Similar to indexing adn slicing Python lists:
print(data[:])
print (data[0:3])
print (data[3:])
print (data[::-2])
# more slicing
x = np.array(range(25))
print ('x:',x)
print()
print (x[5:15:2])
print (x[15:5:-1])
# ## Arrays are like lists, but different
# NumPy utilizes efficient pointers to a location in memory and it will store the full array in data. Lists on the other hand are pointers to many different objects in memory.
# +
# Slicing returns a view in Numpy,
# and not a copy as is the case with Python lists
data = np.array(range(10))
view = data[0:3]
view
# -
l = list(range(10))
copy = l[0:3]
# copy
copy[0] = 99
view[0] = 99
print(copy)
print(view)
print('Python list:',l) # has not changed
print('NumPy array:',data) # has changed
# Creating copies of the array instead of views
arr_copy = data[:3].copy()
print('Array copy',arr_copy)
arr_copy[0] = 555
print('New array copy',arr_copy)
print('Original array',data) # now it is not a view any more
# same thing with assignment, its not a copy, its the same data
x = np.array(range(25))
print (x)
y = x
y[:] = 0
print (x)
x is y
# # Arrays are also a lot faster than lists
# +
# Arrays are faster and more efficient than lists
x = list(range(100000))
y = [i**2 for i in x]
print (y[0:5])
# -
# Time the operation with some IPython magic command
print('Time for Python lists:')
# list_time = %timeit -o -n 20 [i**2 for i in x]
z = np.array(x)
w = z**2
print(w[:5])
print('Time for NumPy arrays:')
# np_time = %timeit -o -n 20 z**2
print('NumPy is ' + str(list_time.all_runs[0]//np_time.all_runs[0]) + ' times faster than lists at squaring 100 000 elements.')
# # Universal functions
# A function that is applied on an `ndarray` in an element-by-element fashion. Several universal functions can be found the NumPy documentation here:
# https://docs.scipy.org/doc/numpy-1.13.0/reference/ufuncs.html
# +
# Arrays are different than lists in another way:
# x and y are lists
x = list(range(5))
y = list(range(5,10))
print ("list x = ", x)
print ("list y = ", y)
print ("x + y = ", x+y)
# -
# now lets try with NumPy arrays:
xn = np.array(x)
yn = np.array(y)
print ('np.array xn =', xn)
print ('np.array xn =', yn)
print ("xn + yn = ", xn + yn)
# # + for np.arrays is a wrapper around the function np.add
np.add(xn,yn)
# +
# An array is a sequence that can be manipulated easily
# An arithmatic operation is applied to each element individually
# When two arrays are added, they must have the same size
# (otherwise they might be broadcasted)
print (3* x)
print (3 * xn)
# -
# # Join, add, concatenate
print(xn)
print(yn)
# if you need to join numpy arrays, try hstack, vstack, column_stack, or concatenate
print (np.hstack((xn,yn)))
print (np.vstack((xn,yn)))
print (np.column_stack((xn,yn)))
print (np.concatenate((xn, yn), axis = 0))
# +
# the elements of an array must be of a type that is valid to perform
# a specific mathematical operation on
data = np.array([1,2,'cat', 4])
print(data)
print(data.dtype)
print (data+1) # results in error
# -
# ### Creating arrays with 2 axis:
#
# This list has two dimensions
list3 = [[1, 2, 3],
[4, 5, 6]]
list3 # nested list
# data = np.array([[1, 2, 3], [4, 5, 6]])
data = np.array(list3)
data
# # Attributes of a multidim array
print('Dimensions:',data.ndim)
print ('Shape:',data.shape)
print('Size:', data.size)
# +
# You can also transpose an array Matrix with either np.transpose(arr)
# or arr.T
print ('Transpose:')
data.T
# print (list3.T) # note, this would not work
# -
# # Other ways to create NumPy arrays
# +
# np.arange() is similar to built in range()
# Creates array with a range of consecutive numbers
# starts at 0 and step=1 if not specified. Exclusive of stop.
np.arange(12)
# -
#Array increasing from start to end: np.arange(start, end)
np.arange(10, 20)
# Array increasing from start to end by step: np.arange(start, end, step)
# The range always includes start but excludes end
np.arange(1, 10, 2)
# Returns a new array of specified size, filled with zeros.
array=np.zeros((2,5), dtype=np.int8)
array
#Returns a new array of specified size, filled with ones.
array=np.ones((2,5), dtype=np.float128)
array
# Returns the identity matrix of specific squared size
array = np.eye(5)
array
# ## Some useful indexing strategies
# ### There are two main types of indexing: Integer and Boolean
x = np.array([[1, 2], [3, 4], [5, 6]])
x
# #### Integer indexing
# first element is the row, 2nd element is the column
print(x[1,0])
print(x[1:,:]) # all rows after first, all columns
# first list contains row indices, 2nd element contains column indices
idx = x[[0,1,2], [0,1,1]] # create index object
print (idx)
# ### Boolean indexing
print('Comparison operator, find all values greater than 3:\n')
print(x>3)
print('Boolean indexing, only extract elements greater than 3:\n')
print(x[x>3])
# ### Masks
arr = np.arange(10)
mask = arr>5
print(mask)
arr[mask]
# Functions any / all
print( np.any( arr==9 ) )
print( np.all( arr>-1 ) )
# ## Extra NumPy array methods
# +
# Reshape is used to change the shape
a = np.arange(0, 15)
print('Original:',a)
a = a.reshape(3, 5)
# a = np.arange(0, 15).reshape(3, 5) # same thing
print ('Reshaped:')
print(a)
# -
# We can also easily find the sum, min, max, .. are easy
print (a)
print ('Sum:',a.sum())
print('Min:', a.min())
print('Max:', a.max())
# +
print ('Sum along rows:',a.sum(axis=0))
print ('Sum along rows:',a.sum(axis=1))
# Note here axis specifies what dimension to "collapse"
# -
# ## Arrray Axis
# <img src= "https://github.com/ikhlaqsidhu/data-x/raw/master/imgsource/anatomyarray.png">
#
#
# To get the cumulative product:
print (np.arange(1, 10))
print (np.cumprod(np.arange(1, 10)))
# To get the cumulative sum:
print (np.arange(1, 10))
np.cumsum((np.arange(1, 10)))
# Creating a 3D array:
a = np.arange(0, 96).reshape(2, 6, 8)
print(a)
# The same methods typically apply in multiple dimensions
print (a.sum(axis = 0))
print ('---')
print (a.sum(axis = 1))
# # More ufuncs and Basic Operations
# One of the coolest parts of NumPy is the ability for you to run mathematical operations on top of arrays. Here are some basic operations:
a = np.arange(11, 21)
b = np.arange(0, 10)
print ("a = ",a)
print ("b = ",b)
print (a + b)
a * b
a ** 2
# You can even do things like matrix operations
a.dot(b)
# Matrix multiplication
c = np.arange(1,5).reshape(2,2)
print ("c = \n", c)
print()
d = np.arange(5,9).reshape(2,2)
print ("d = \n", d)
print (d.dot(c))
np.matmul(d,c)
# # Random numbers
# Radom numbers
np.random.seed(0) # set the seed to zero for reproducibility
print(np.random.uniform(1,5,10)) # 10 random uniform numbers from 1 to 5
print()
print (np.random.exponential(1,5)) # 5 random exp numbers with rate 1
print (np.random.random(8).reshape(2,4)) #8 random 0-1 in a 2 x 4 array
# If you want to learn more about "random" numbers in NumPy go to: https://docs.scipy.org/doc/numpy-1.12.0/reference/routines.random.html
# # Trignometric functions
# +
# linspace: Create an array with numbers from a to b
# with n equally spaced numbers (inclusive)
data = np.linspace(0,10,5)
print (data)
# -
from numpy import pi
x = np.linspace(0,pi, 3)
print('x = ', x)
print()
print ("sin(x) = ", np.sin(x))
# flatten matrices using ravel()
x = np.array(range(24))
x = x.reshape(4,6)
print('Original:\n',x)
print()
x = x.ravel() # make it flat
print ('Flattened:\n',x)
|
02a-tools-numpy/archive/numpy-predict-intro-v6_1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # **Basic problem solution**
# imports
import numpy as np
import pandas as pd
import networkx as nx
import matplotlib.pyplot as plt
# global parameters
rng = np.random.default_rng(12345)
# ### **Generate random graph**
# local parameters
nnodes = 10
nedges = 12
# Checks
assert nedges <= (nnodes - 1) * nnodes / 2 # nedges > max possible nedges in the full graph
G = nx.gnm_random_graph(nnodes, nedges)
nx.draw(
G,
with_labels=True,
node_color="indigo",
font_color="w",
)
# Betweenness centrality
nx.betweenness_centrality(G)
# Adjacency matrix
nx.to_pandas_adjacency(G, dtype=int)
# ### **Read graph from file**
df = pd.read_csv("../Data/example_adjacency_matrix.csv", header=None)
H = nx.from_pandas_adjacency(df)
nx.draw(
H,
with_labels=True,
node_color="cyan",
font_color="k",
)
# Betweenness centrality
nx.betweenness_centrality(H)
# Adjacency matrix
nx.to_pandas_adjacency(H, dtype=int)
# + [markdown] pycharm={"name": "#%% md\n"}
# ### **Apply mutations to the graph**
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
prob = 0.1 # mutation probability
n = len(H.nodes)
ad_mat = nx.to_numpy_array(H, dtype=int)
mask = np.tril(rng.choice([0, 1], (n, n), p=[1 - prob, prob]), k=-1)
mask += mask.T
new_ad_mat = ad_mat ^ mask
Q = nx.from_numpy_array(new_ad_mat)
nx.is_isomorphic(H, Q)
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
nx.draw(
Q,
with_labels=True,
node_color="lightgreen",
font_color="k",
)
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
|
Notebooks/basic_graph_example.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={} colab_type="code" id="wPY4it_wHzvW"
import torch
# + colab={} colab_type="code" id="HY0eFW_8H-3r"
def activation(x):
"""Create a sigmoid activation function.
Good for outputs that fall between 0 and 1. (probability)
args x: a torch tensor.
"""
return 1/(1 + torch.exp(-x))
# + colab={} colab_type="code" id="y9qIUKojIYcj"
# generate some data
# set some random seed so that the result is predictatble
data = torch.manual_seed(7)
# set some features to 5 random variables
# 2-dimensional matrix/tensor of 1 row and 5 columns
features = torch.randn((1,5))
# set weights
weights = torch.randn_like(features)
# set true bias term
bias = torch.randn((1,1))
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="7vdqvgK3c29m" outputId="aff8fffd-2c90-467c-9e74-c4285523bfc9"
# calculate y output
# y = (weights.features + bias)
x = torch.sum(weights * features) + bias
y = activation(x)
print(y)
# + colab={} colab_type="code" id="aJ09sXxNd4gP"
# better to do matrix multiplication because it's optimized
torch.mm(weights, features)
# + colab={} colab_type="code" id="F_AApoJiWvAJ"
# + [markdown] colab_type="text" id="68IgplvGXA5J"
# Since we are doing matrix multiplication, we need the matrices shapes to match.
# We'll change the shape of weights for the mm to work.
# + colab={} colab_type="code" id="eMgeaC0-W_ZA"
# weight.reshape(a, b) reshapes the data into a tensor of size (a, b)
# weight.resize_(a, b) returns the same tensor with a different shape.
# if
# weight.view(a, b) returns a new tensor
print(weights.shape)
reshaped_weights = weights.view(5, 1)
y = activation(torch.mm(reshaped_weights, features) + bias)
print(y)
# + colab={} colab_type="code" id="Y-t9h0_RXAAB"
|
intro-notebooks/single_layer_network.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from tensorflow import keras
from tensorflow.keras import *
from tensorflow.keras.models import *
from tensorflow.keras.layers import *
from tensorflow.keras.regularizers import l2#正则化L2
import tensorflow as tf
import numpy as np
import pandas as pd
# 12-0.2
# 13-2.4
# 18-12.14
import pandas as pd
import numpy as np
normal = np.loadtxt(r'F:\张老师课题学习内容\code\数据集\试验数据(包括压力脉动和振动)\2013.9.12-未发生缠绕前\2013-9.12振动\2013-9-12振动-1250rmin-mat\1250rnormalviby.txt', delimiter=',')
chanrao = np.loadtxt(r'F:\张老师课题学习内容\code\数据集\试验数据(包括压力脉动和振动)\2013.9.17-发生缠绕后\振动\9-18上午振动1250rmin-mat\1250r_chanraoviby.txt', delimiter=',')
print(normal.shape,chanrao.shape,"***************************************************")
data_normal=normal[6:8] #提取前两行
data_chanrao=chanrao[6:8] #提取前两行
print(data_normal.shape,data_chanrao.shape)
print(data_normal,"\r\n",data_chanrao,"***************************************************")
data_normal=data_normal.reshape(1,-1)
data_chanrao=data_chanrao.reshape(1,-1)
print(data_normal.shape,data_chanrao.shape)
print(data_normal,"\r\n",data_chanrao,"***************************************************")
#水泵的两种故障类型信号normal正常,chanrao故障
data_normal=data_normal.reshape(-1, 512)#(65536,1)-(128, 515)
data_chanrao=data_chanrao.reshape(-1,512)
print(data_normal.shape,data_chanrao.shape)
# +
import numpy as np
def yuchuli(data,label):#(4:1)(51:13)
#打乱数据顺序
np.random.shuffle(data)
train = data[0:102,:]
test = data[102:128,:]
label_train = np.array([label for i in range(0,102)])
label_test =np.array([label for i in range(0,26)])
return train,test ,label_train ,label_test
def stackkk(a,b,c,d,e,f,g,h):
aa = np.vstack((a, e))
bb = np.vstack((b, f))
cc = np.hstack((c, g))
dd = np.hstack((d, h))
return aa,bb,cc,dd
x_tra0,x_tes0,y_tra0,y_tes0 = yuchuli(data_normal,0)
x_tra1,x_tes1,y_tra1,y_tes1 = yuchuli(data_chanrao,1)
tr1,te1,yr1,ye1=stackkk(x_tra0,x_tes0,y_tra0,y_tes0 ,x_tra1,x_tes1,y_tra1,y_tes1)
x_train=tr1
x_test=te1
y_train = yr1
y_test = ye1
#打乱数据
state = np.random.get_state()
np.random.shuffle(x_train)
np.random.set_state(state)
np.random.shuffle(y_train)
state = np.random.get_state()
np.random.shuffle(x_test)
np.random.set_state(state)
np.random.shuffle(y_test)
#对训练集和测试集标准化
def ZscoreNormalization(x):
"""Z-score normaliaztion"""
x = (x - np.mean(x)) / np.std(x)
return x
x_train=ZscoreNormalization(x_train)
x_test=ZscoreNormalization(x_test)
# print(x_test[0])
#转化为一维序列
x_train = x_train.reshape(-1,512,1)
x_test = x_test.reshape(-1,512,1)
print(x_train.shape,x_test.shape)
def to_one_hot(labels,dimension=2):
results = np.zeros((len(labels),dimension))
for i,label in enumerate(labels):
results[i,label] = 1
return results
one_hot_train_labels = to_one_hot(y_train)
one_hot_test_labels = to_one_hot(y_test)
# -
x = layers.Input(shape=[512,1,1])
Flatten=layers.Flatten()(x)
Dense1=layers.Dense(12, activation='relu')(Flatten)
Dense2=layers.Dense(6, activation='relu')(Dense1)
Dense3=layers.Dense(2, activation='softmax')(Dense2)
model = keras.Model(x, Dense3)
model.summary()
# +
#定义优化
model.compile(loss='categorical_crossentropy',
optimizer='adam',metrics=['accuracy'])
# -
import time
time_begin = time.time()
history = model.fit(x_train,one_hot_train_labels,
validation_split=0.1,
epochs=50,batch_size=10,
shuffle=True)
time_end = time.time()
time = time_end - time_begin
print('time:', time)
# +
import time
time_begin = time.time()
score = model.evaluate(x_test,one_hot_test_labels, verbose=0)
print('Test loss:', score[0])
print('Test accuracy:', score[1])
time_end = time.time()
time = time_end - time_begin
print('time:', time)
# +
#绘制acc-loss曲线
import matplotlib.pyplot as plt
plt.plot(history.history['loss'],color='r')
plt.plot(history.history['val_loss'],color='g')
plt.plot(history.history['accuracy'],color='b')
plt.plot(history.history['val_accuracy'],color='k')
plt.title('model loss and acc')
plt.ylabel('Accuracy')
plt.xlabel('epoch')
plt.legend(['train_loss', 'test_loss','train_acc', 'test_acc'], loc='center right')
# plt.legend(['train_loss','train_acc'], loc='upper left')
#plt.savefig('1.png')
plt.show()
# +
import matplotlib.pyplot as plt
plt.plot(history.history['loss'],color='r')
plt.plot(history.history['accuracy'],color='b')
plt.title('model loss and sccuracy ')
plt.ylabel('loss/sccuracy')
plt.xlabel('epoch')
plt.legend(['train_loss', 'train_sccuracy'], loc='center right')
plt.show()
|
BCNcode/0_vibratioon_signal/1250/DNN/CNN_1250-015-512-y.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from os import path
from enmspring.graphs import hosts
from enmspring.graphs import Stack
rootfolder = '/home/yizaochen/codes/dna_rna/fluctmatch_sequence'
enmroot = '/home/yizaochen/codes/dna_rna/enmspring'
# ### Part 1: Initialize
host = 'a_tract_21mer'
g_agent = Stack(host, rootfolder)
g_agent.build_node_list()
g_agent.initialize_three_mat()
# ### Part 2: Show crd in VMD
g_agent.vmd_show_crd()
# ### Part 3: Show nodes in the graph
tcl_out = path.join(enmroot, 'tclscripts', f'{host}_base_nodes.tcl')
colorid = 0
vdw_radius = 0.2
g_agent.write_show_nodes_tcl(tcl_out, colorid=colorid, vdw_radius=vdw_radius)
# ### Part 4: Show edges in the graph
tcl_out = path.join(enmroot, 'tclscripts', f'{host}_base_edges.tcl')
radius = 0.15
g_agent.write_show_base_edges_tcl(tcl_out, radius=radius)
# ### Additional Part 1: Copy back
for host in hosts:
g_agent = GraphAgent(host, rootfolder)
g_agent.copy_nohydrogen_crd()
# ### Additional Part 2: Find 5' of Guide
print('serial 6 7 8 9')
|
notebooks/draw_graph_basestack_vmd.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ## Final Project: Linear Regression
#
# - We want to present the relationship between (two) variables linearly
#
# - For example, recall the running distance and drinking water
#
# - We are interested to obtain the best line describing by `y_pred[i] = w_1 x[i] +w_0` that maps running distance to drinking water
#
# - Below, list `x` represents running distance in miles and list `y` represents the drinking water in litres
# +
import numpy as np
import matplotlib.pyplot as plt
# Running Distance in Mile
x = np.array([3.3,4.4,5.5,6.71,6.93,4.168,9.779,6.182,7.59,2.167,
7.042,10.791,5.313,7.997,5.654,9.27,3.1])
# Water Drinks in Litre
y = np.array([1.7,2.76,2.09,3.19,1.694,1.573,3.366,2.596,2.53,1.221,
2.827,3.465,1.65,2.904,2.42,2.94,1.3])
plt.scatter(x, y)
plt.xlabel('Running Distance (Mile)')
plt.ylabel('Water Drinks (Litre)')
# -
# ## In order to obtain the best line, we should define error first
#
# - For linear relationship, mean-square-error (MSE) represents is it a good line (good model) or not
#
# - $y[i]$ and $y_{pred}[i]$ are i-th element of list `y` and list `y_pred` respectively where `y_pred[i] = w_1 x[i] +w_0`
#
# - Define error as: $E[i] = y_{pred}[i] - y[i]$
#
# - Define mean-square-error as: $MSE = \frac{1}{N} \sum_{i=0}^{N-1} E[i]^2$
#
# - Also mean-square-error is equal to: $MSE = \frac{1}{N} \sum_{i=0}^{N-1} (y_{pred}[i] - y[i])^2$
#
# - The parameter $N$ is: $N = len(y)$
# ### Question 1: Obtain the MSE for the following two lines:
#
# 1- `y_pred[i] = 0.7*x[i] + 0.3`
#
# 2- `y_pred[i] = 0.25163494*x[i] + 0.79880123`
#
# Hint: Your function take four input arguments: 1- y, 2- x, 3- slope, 4-intercept
# +
def min_sq_error(Y, X, w1, w0):
y_pred = [i*w1 + w0 for i in X]
sum_squared_error = sum((y_p-y)**2 for y, y_p in zip(Y, y_pred))
N = len(y)
mse = sum_squared_error/N
return mse
print(min_sq_error(y, x, 0.7, 0.3))
print(min_sq_error(y, x, 0.25163494, 0.79880123))
# -
# ## Question 2: Obtain the best line (Coding is not required)
#
# - In order the best line, we want to obtain which slope ($w_1$) and intercept ($w_0$) minimize the mean-square-error (MSE)
#
# - Mathematically:
#
# - $MSE = \frac{1}{N} \sum_{i=0}^{N-1} (y_{pred}[i] - y[i])^2$
#
# - $MSE = f(w_1, w_0)= \frac{1}{N} \sum_{i=0}^{N-1} (w_1x[i] + w_0 - y[i])^2$
#
# The steps in order to obtain the best line:
#
# 1- Compute:
#
# $\frac{\partial MSE}{\partial w_1}$
#
# and
#
# $\frac{\partial MSE}{\partial w_0}$
#
# 2- then obtain $w_1$ and $w_0$ such that:
#
#
# $\frac{\partial MSE}{\partial w_1} = 0$
#
# and
#
# $\frac{\partial MSE}{\partial w_0} = 0$
#
# - For this part, you need to use partial derivative and you use derivative table
#
# - For this part, write down the steps and the solution on a paper
# ## Question 3: Write a code to return the slope and intercept with given list x and y
#
# - After taking partial derivative of mean-squared-error and setting to zero for both $w_1$ and $w_0$ ($\frac{\partial MSE}{\partial w_1} = 0$ $\frac{\partial MSE}{\partial w_0} = 0$)
#
# - $w_1$ = is obtained from list x and list y
# - $w_0 $ = is obtained from $w_1$, list x and list y
# - Write a Python function that return $w_1$ and $w_0$ from your calculation on the paper
# +
def slope_intercept_LR(x, y):
x_bar = np.mean(x)
y_bar = np.mean(y)
xy_bar = np.mean(x*y)
x_squared_bar = np.mean(np.square(x))
x_bar_squared = x_bar **2
w1 = ((xy_bar - x_bar * y_bar) / (x_squared_bar - x_bar_squared))
w0 = (y_bar - w1*x_bar)
return w1, w0
print(slope_intercept_LR(x, y))
# -
# ## Question 4: After obtain the best line, obtain the variance and mean of error
#
# - In Question 3, we have obtained the best line
# - So, we can error list which its element is: $E[i] = y_{pred}[i] - y[i]$
# - Write a function that calculate variance and mean of list $E$
# - Plot the distribution of the error for optimal line
# +
def variance_mean(x, y, w1, w0):
y_pred = [w1*x[i] + w0 for i in range(len(x))]
error = [y_pred[i] - y[i] for i in range(len(y))]
mean_e = np.mean(error)
var_e = np.var(error)
plt.scatter(x, y)
plt.xlabel("Running Distance (Miles)")
plt.ylabel("Water Drinks (Liters)")
plt.plot(x, y_pred, 'ro-') # line of fit
return str(var_e), str(mean_e)
print(variance_mean(x, y, 0.25163494428355315, 0.79880123 ))
#print(variance_mean(y, x, 0.7, 0.3))
# +
import seaborn as sns
y_pred = [0.25163494428355315*x[i] + 0.79880123 for i in range(len(x))]
a = y_pred - y
sns.distplot(a, hist=True, kde=True, bins=5)
# -
# ## Question 5: (Optional but Bonus point) In almost all applications, we update the slope and intercept through iteration
#
# - Instead of putting the $\frac{\partial MSE}{\partial w_1} = 0$ $\frac{\partial MSE}{\partial w_0} = 0$
# - Initialize the $w_1$ and $w_0$ with arbitrary value, then update them by follwing Gradient Updating Rule:
# - $w_1 = w_1 - step*\frac{\partial MSE}{\partial w_1}$
# - $w_0 = w_0 - step*\frac{\partial MSE}{\partial w_0}$
# +
import numpy as np
w_0 = np.random.randn()
w_1 = np.random.randn()
x_bar = np.mean(x)
y_bar = np.mean(y)
xy_bar = np.mean(x*y)
x_squared_bar = np.mean(np.square(x))
step = 0.01
epoch = 5000
for _ in range(epoch):
w_1 = w_1 - step*((w_1*x_squared_bar) + (w_0*x_bar) - (xy_bar))
w_0 = w_0 - step*((w_1*x_bar) + (w_0) - (y_bar))
print(w_1)
print(w_0)
# -
|
Final_Project/.ipynb_checkpoints/Final_Project-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Populate MOF with given molecule and initialize LAMMPS
# - Read MOF xyz file
# - Get size of the MOF
# - Center MOF to size/2
# - Add molecule
# - Run packmol
# - Convert xyz to cif
import os, shutil
from angstrom import Molecule
from packmol import Packmol
# +
cells = {'BEDYEQ': [17.1430, 41.9680, 41.9680, 120.0, 90.0, 90.0],
'IRMOF-1': [25.8320, 25.8320, 25.8320, 90.0, 90.0, 90.0],
'UIO-66': [20.7004, 20.7004, 20.7004, 90.0, 90.0, 90.0]}
molpos = {'BEDYEQ': [10, 10, 10, 30, 30, 30],
'IRMOF-1': [5, 5, 5, 20, 20, 20],
'UIO-66': [5, 5, 5, 20, 20, 20]}
n_molecules = 1
molecule_padding = 5
packmol_dir = '/home/kutay/Documents/Research/DMMP/packmol'
# -
for mof_name in cells:
mof_file = os.path.join(packmol_dir, '%s-111.xyz' % mof_name)
mof = Molecule(read=mof_file)
# Replicate and get box size
mof.set_cell(cells[mof_name])
mof.cell.calculate_vertices()
box_size = cells[mof_name]
pmol = Packmol()
pmol.options['tolerance'] = 3.0
# Add mof
mof_position = {'fixed': '%.1f %.1f %.1f 0. 0. 0.' % (box_size[0] / 2, box_size[1] / 2, box_size[2] / 2), 'centerofmass': ''}
pmol.add_structure({'structure': '%s-111.xyz' % mof_name, 'number': '1', 'position': mof_position})
# Add molecule
mol_file = os.path.join(packmol_dir, 'dmma.xyz')
mol_position = {'inside box': '%s' % ' '.join([str(round(i, 1)) for i in molpos[mof_name]])}
pmol.add_structure({'structure': 'dmmp.xyz', 'number': n_molecules, 'position': mol_position})
run_dir = os.path.join(packmol_dir, '%s' % mof_name)
pmol.run(run_dir, packmol_dir)
# Create cif file
packed_xyz = Molecule(read=os.path.join(run_dir, pmol.options['output']))
cell_center = (mof.cell.vertices[0] + mof.cell.vertices[-1]) / 2
packed_xyz.center(cell_center)
packed_xyz.write(os.path.join(packmol_dir, '%s-dmmp.cif' % mof_name), cell=mof.cell.to_list())
pmol.clear()
# ### LAMMPS
from thermof import Simulation
from thermof import Parameters
lammps_dir = '/home/kutay/Documents/Research/DMMP/lammps'
simpar = Parameters()
simpar.lammps
for mof_name in cells:
sim_dir = os.path.join(lammps_dir, mof_name)
cif_file = os.path.join(packmol_dir, '%s-dmmp.cif' % mof_name)
simpar = Parameters()
sim = Simulation(mof=os.path.abspath(cif_file), parameters=simpar)
sim.set_dir(sim_dir)
simpar.lammps['replication'] = '2 2 2'
simpar.lammps['force_field'] = 'UFF4MOF'
simpar.lammps['dont_replicate_molecules'] = True
simpar.thermof['fix'] = ['MIN', 'NVT', 'NVE']
simpar.thermof['min']['edif'] = 1e-3
simpar.thermof['nvt']['steps'] = 500000
simpar.thermof['nve']['steps'] = 1000000
simpar.thermof['thermo_style'] = ['step', 'temp', 'press', 'pe', 'etotal', 'emol', 'epair', 'vol', 'lx', 'ly', 'lz']
simpar.job['nodes'] = 1
simpar.job['ppn'] = 28
simpar.job['walltime'] = '36:00:00'
simpar.job['name'] = '%s-dmmp' % mof_name
sim.initialize()
|
scripts/lammps/Diffusion-111.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="0zFs256qPZi6"
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
import tensorflow_hub as hub
from tensorflow.keras.layers import Conv1D
from tensorflow.keras.layers import LSTM, Bidirectional
from tensorflow.keras.layers import Dense, Flatten, MaxPooling1D, Input, Activation, concatenate
from tensorflow.keras.models import Sequential, Model
from sklearn.preprocessing import OneHotEncoder
from tensorflow.keras.layers import Dropout, BatchNormalization, GaussianNoise
from sklearn.utils import resample
from sklearn.utils import shuffle
from tensorflow.keras.models import load_model
# + id="Uif3patwPiIM"
data = pd.read_csv("test.csv", sep=",")
print(data.shape, data.columns)
data = data.dropna()
features1 = data["title1_en"]
features2 = data["title2_en"]
# + id="aI8vfUhEPtd4"
embed = hub.load("https://tfhub.dev/google/universal-sentence-encoder/4")
# + id="PxpZ03j3P37q"
embedded_features1 = embed(features1)
embedded_features1 = np.array(embedded_features1)
result = pd.DataFrame(embedded_features1)
result.to_csv("features1.csv", index=False)
# + id="mbgtVV-mQL2f"
embedded_features2 = embed(features2)
embedded_features2 = np.array(embedded_features2)
result = pd.DataFrame(embedded_features2)
result.to_csv("features2.csv", index=False)
# + id="QDKd8tm9Q0g3"
features1 = pd.read_csv("features1.csv")
features2 = pd.read_csv("features2.csv")
# + id="J9vbU172Q-LC"
processed = pd.concat([features1, features2], axis=1)
# + id="9e6baVhiW3oZ"
processed = pd.read_csv("test_merged.csv")
# + id="JKOyt4WeW-dr"
processed.shape
# + id="vnNz4bfHRi4y"
from sklearn.preprocessing import LabelEncoder
from tensorflow.keras.utils import to_categorical
# + [markdown] id="q7LSpcJhjD4V"
# #### Testing 1 score 76
# + id="sBykOsX7RrI3"
a = Sequential()
a.add(Dense(units=64, input_shape=(1024,),activation='relu'))
a.add(Dropout(0.2))
a.add(Dense(units=128, input_shape=(1024,),activation='relu'))
a.add(Dense(units=32, input_shape=(1024,),activation='relu'))
a.add(Dense(3, activation='softm.7790918e-01 6.1159653e-08 8.220908sentropy', optimizer="adam", metrics=['accuracy'])
# + [markdown] id="rR5ozPLhjHIs"
# #### Testing 2
# + id="RlFN_gcHi_8d"
model = Sequential()
model.add(LSTM(units=128, return_sequences=True, kernel_initializer='he_normal'))
model.add(Dropout(0.5))
model.add(BatchNormalization())
model.add(LSTM(units=128, return_sequences=True, go_backwards=True, kernel_initializer='he_normal'))
model.add(Dropout(0.5))
model.add(BatchNormalization())
model.add(LSTM(units=128, kernel_initializer='he_normal'))
model.add(Dense(3, activation='softmax'))
model.compile(loss='categorical_crossentropy', optimizer="adam", metrics=['accuracy'])
# + id="RPfe6UqeG-F1"
input1 = Input(shape=(None, 512))
inner = Conv1D(256, 1, padding='same', name='conv1', kernel_initializer='he_normal')(input1)
inner = BatchNormalization()(inner)
inner = Dropout(0.5)(inner)
inner = Activation('relu')(inner)
inner1 = MaxPooling1D(pool_size=1, name='max1')(inner)
# inner = Conv1D(256, 1, padding='same', name='conv1', kernel_initializer='he_normal')(inner1)
# inner = BatchNormalization()(inner)
# inner = Dropout(0.5)(inner)
# inner = Activation('relu')(inner)
# inner1 = MaxPooling1D(pool_size=1, name='max1')(inner)
# inner = Conv1D(512, 1, padding='same', name='conv1', kernel_initializer='he_normal')(inner1)
# inner = BatchNormalization()(inner)
# inner = Dropout(0.5)(inner)
# inner = Activation('relu')(inner)
# inner1 = MaxPooling1D(pool_size=1, name='max1')(inner)
# inner = Conv1D(512, 1, padding='same', name='conv1', kernel_initializer='he_normal')(inner1)
# inner = BatchNormalization()(inner)
# inner = Dropout(0.5)(inner)
# inner = Activation('relu')(inner)
# inner1 = MaxPooling1D(pool_size=1, name='max1')(inner)
# inner = Conv1D(768, 1, padding='same', name='conv1', kernel_initializer='he_normal')(inner1)
# inner = BatchNormalization()(inner)
# inner = Dropout(0.5)(inner)
# inner = Activation('relu')(inner)
# inner1 = MaxPooling1D(pool_size=1, name='max1')(inner)
input2 = Input(shape=(None, 512))
inner2 = Conv1D(256, 1, padding='same', name='conv1', kernel_initializer='he_normal')(input2)
inner2 = BatchNormalization()(inner2)
inner2 = Dropout(0.5)(inner2)
inner2 = Activation('relu')(inner2)
inner2 = MaxPooling1D(pool_size=1, name='max1')(inner2)
# inner2 = Conv1D(256, 1, padding='same', name='conv1', kernel_initializer='he_normal')(inner2)
# inner2 = BatchNormalization()(inner2)
# inner2 = Dropout(0.5)(inner2)
# inner2 = Activation('relu')(inner2)
# inner2 = MaxPooling1D(pool_size=1, name='max1')(inner2)
# inner2 = Conv1D(512, 1, padding='same', name='conv1', kernel_initializer='he_normal')(inner2)
# inner2 = BatchNormalization()(inner2)
# inner2 = Dropout(0.5)(inner2)
# inner2 = Activation('relu')(inner2)
# inner2 = MaxPooling1D(pool_size=1, name='max1')(inner2)
# inner2 = Conv1D(512, 1, padding='same', name='conv1', kernel_initializer='he_normal')(inner2)
# inner2 = BatchNormalization()(inner2)
# inner2 = Dropout(0.5)(inner2)
# inner2 = Activation('relu')(inner2)
# inner2 = MaxPooling1D(pool_size=1, name='max1')(inner2)
# inner2 = Conv1D(768, 1, padding='same', name='conv1', kernel_initializer='he_normal')(inner2)
# inner2 = BatchNormalization()(inner2)
# inner2 = Dropout(0.5)(inner2)
# inner2 = Activation('relu')(inner2)
# inner2 = MaxPooling1D(pool_size=1, name='max1')(inner2)
x = concatenate([input1, input2])
x = Dense(1024, activation='relu')(x)
x = BatchNormalization()(x)
x = Dropout(0.5)(x)
x = Dense(1024, activation='relu')(x)
x = BatchNormalization()(x)
x = Dropout(0.5)(x)
x = Dense(1024, activation='relu')(x)
x = Dense(3, activation='softmax')(x)
model = Model(inputs=[input1, input2], outputs = x)
model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
# + id="HKM5tORjR__w"
model = load_model("cnn+concat")
# + id="lZpvfhgKSp_p"
mapping = {0:"agreed", 1:"disagreed", 2:"unrelated"}
# + id="1YnsF7Yw59YX"
processed = np.array(processed)
# + id="e7CjgRT550tL"
processed = processed.reshape((processed.shape[0],1, processed.shape[1]))
# + id="Ndx1YHfU727w"
feature1 = np.array(features1).reshape((features1.shape[0],1, features1.shape[1]))
feature2 = np.array(features2).reshape((features2.shape[0],1, features2.shape[1]))
# + colab={"base_uri": "https://localhost:8080/"} id="jiH5U-Sw7mUJ" outputId="f6404bcc-1e0d-4c17-ce78-e9b648c68d35"
processed.shape
# + id="kzqyVEMZUTrf"
result = model.predict((feature1, feature2))
# + id="ubvKkeBCKJdz"
result = model.predict(processed)
# + id="g-_HKldWaua6"
predicted_labels = list()
# + id="B7yYOQaXaKR4"
for i in range(result.shape[0]):
print(i)
predicted_labels.append(mapping[np.argmax(result[i])])
# + id="5sZa0L04ceso"
data = pd.read_csv("test.csv", sep=",")
# + id="EJk_X_erSHM7"
data["label"] = pd.DataFrame(predicted_labels)
# + id="bP72lX9mTS4P"
data.to_csv("cnn+concat.csv", index=False)
|
code/Notebooks/fake_news_classification_inference.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # IR-Reproducibility with Transferred Relevance Judgments
# +
import pandas as pd
import json
import numpy as np
import seaborn as sns
import random
import sys
sys.path.append('../../../simulate-label-transfer/transfer_simulation')
from repro_measures import effect_ratio, delta_relative_improvement, classify_repro_pairs
def regression_by_measure(df, measure: str):
g = sns.FacetGrid(
df.loc[:,["corpus","tag",measure]].sort_values(measure).replace({k: v for v,k in enumerate(df.sort_values(measure).tag.unique())}),
hue="corpus",
height=6,
aspect=1.2
)
g.map(sns.regplot, "tag", measure, scatter_kws={'alpha':0.4})
g.add_legend()
return g
def query_regression_by_measure(df, topic: int, measure: str):
regression_by_measure(df[df['topic'] == topic], measure)
def extract_metadata_from_Tag(tag):
tag = tag.replace('clueweb12-and-wayback12', 'clueweb12_and_wayback12')
return dict(list(map(lambda x: x.split("^"), ("collection^"+tag).split("-"))))
def sorted_df(df):
return df.sort_values("tag",ascending=False)
def import_df(file_name):
df = pd.read_json(file_name, lines=True)
df = pd.merge(
pd.DataFrame(df.tag.apply(lambda t: extract_metadata_from_Tag(t)).tolist()).reset_index(),
df.reset_index(),
on="index"
).drop(["index","tag"],axis=1)
df["tag"] = df['body_lang.en'].astype(str)+df['title_lang.en'].astype(str)+df['meta_desc_lang.en'].astype(str)
return sorted_df(df)
def build_topic_df(df, topics: list=None, topic_start: int=None, topic_end: int=None):
if topics and topic_start and topic_end:
raise
elif not topics:
topics = [i for i in range(topic_start, topic_end+1)]
nqueries = len(topics)
df = df[(df['topic'].astype(int).isin(topics))]
df = df.groupby(['collection', 'body_lang.en', 'title_lang.en', 'meta_desc_lang.en', 'corpus', 'tag']).agg({
'bpref': np.mean,
'pseudoNDCG@10': np.mean,
'pseudoNDCG': np.mean
}).reset_index()
columns = ['collection', 'body_lang.en', 'title_lang.en', 'meta_desc_lang.en', 'corpus', 'bpref', 'pseudoNDCG@10', 'pseudoNDCG', 'tag']
if topic_start and topic_end:
columns = ['collection', 'body_lang.en', 'title_lang.en', 'meta_desc_lang.en', 'corpus', 'topics', 'bpref', 'pseudoNDCG@10', 'pseudoNDCG', 'tag']
df['topics'] = 'topics-' + str(topic_start) + '-' + str(topic_end)
df = df[columns]
return sorted_df(df)
def is_source_collection(r, target_collection):
if target_collection in ['cw12', 'cw12wb12']:
return r['topic'] <= 200 and r['corpus'] == 'cw09'
elif target_collection == 'cc15':
return (r['corpus'] == 'cw09' and r['topic'] <= 200) or (r['corpus'] == 'cw12' and r['topic'] > 200)
raise
def label_transfer_df(df, target_collection):
df = df.copy()
df['is_source_collection'] = df.apply(lambda i: is_source_collection(i, target_collection), axis=1)
df = df[(df['is_source_collection'] == True) | (df['corpus'] == target_collection)]
df['corpus'] = df.apply(lambda i: 'source' if i['is_source_collection'] else target_collection, axis=1)
df['collection'] = df.apply(lambda i: 'source' if i['is_source_collection'] else i['collection'], axis=1)
df['keep'] = df.apply(lambda i: is_shallow_topic(target_collection, i['topic']), axis=1)
df = df[df['keep'] == True]
return df
# -
base_dir = '/mnt/ceph/storage/data-in-progress/kibi9872/sigir2021/'
topic_df = import_df(base_dir + 'data-26-10-2020/rankings/reproducibility-evaluation-zero-scores-removed.jsonl')
query_df = import_df(base_dir + 'data-26-10-2020/rankings/reproducibility-evaluation-per-query-zero-scores-removed.jsonl')
# # Helpers from transferred-topics
# +
import pandas as pd
def is_shallow_topic(topic):
return topic['relevant'] > 0 and topic['irrelevant'] > 0 and (topic['relevant'] + topic['irrelevant']) >= 10
transferred_topics = pd.read_json('../../test/resources/overview-of-transferred-topics.jsonl', lines=True)
transferred_topics['shallowTopic'] = transferred_topics.apply(lambda i: is_shallow_topic(i), axis=1)
def is_shallow_topic(corpus, topic):
tmp_df = transferred_topics[(transferred_topics['targetCorpus'] == corpus) & (transferred_topics['topic'] == topic)]
return len(tmp_df) > 0 and tmp_df.iloc[0]['shallowTopic']
# -
# # Precalculations
query_dfs = {
'cc15': label_transfer_df(query_df, 'cc15'),
'cw12': label_transfer_df(query_df, 'cw12'),
'cw12wb12': label_transfer_df(query_df, 'cw12wb12'),
}
topic_df
[i for i in query_dfs['cc15'][(query_dfs['cc15']['is_source_collection'] == True) & (query_dfs['cc15']['collection'] == 'source')].tag.unique()]
# # Sample Pairs
#
# - Load from file if exists, or calculate new pairs
# +
def position_pair(corpus, topics, measure, firstPos=None, secondPos=None):
df = performance_ranking_on_source(corpus, topics, measure)
tags = [i for i in df.keys()]
tags = sorted(tags, key=lambda i: df[i]['pos'], reverse=False)
if firstPos == None:
firstPos = random.randrange(0, len(tags))
if secondPos == None:
secondPos = random.randrange(0, len(tags))
firstTag = tags[firstPos]
secondTag = tags[secondPos]
if firstTag == secondTag:
return position_pair(corpus, topics, measure)
else:
first_topic_level_df = topics_to_performance_on_source_and_on_target_for_tag(corpus, firstTag, topics, measure)
first_topic_level_df['tag'] = firstTag
second_topic_level_df = topics_to_performance_on_source_and_on_target_for_tag(corpus, secondTag, topics, measure)
second_topic_level_df['tag'] = secondTag
topic_level_df = pd.concat([first_topic_level_df, second_topic_level_df])
if df[firstTag]['pos'] > df[secondTag]['pos']:
return calculate_position_pair(df, secondTag, firstTag, topic_level_df)
else:
return calculate_position_pair(df, firstTag, secondTag, topic_level_df)
def calculate_position_pair(df, first, second, topic_level_df):
return {
'firstTag': first,
'sourceFirstPos': df[first]['pos'],
'sourceFirstScore': df[first]['measure'],
'sourceSecondPost': df[second]['pos'],
'sourceSecondTag': second,
'sourceSecondScore': df[second]['measure'],
'effectRatio': effect_ratio(df=topic_level_df, baseline=second, advanced=first),
'delta_relative_improvement': delta_relative_improvement(df=topic_level_df, baseline=second, advanced=first)
}
def topics_to_performance_on_source_and_on_target_for_tag(corpus, tag, topics, measure):
df = query_dfs[corpus]
df = df[df['tag'] == tag]
df = df[df['topic'].astype(int).isin(topics)].copy()
df['measure'] = df[measure]
return df[['collection', 'topic', 'measure']].reset_index()
def query_df_for_topic(corpus, topics):
df = query_dfs[corpus]
df = df[(df['topic'].astype(int).isin(topics))]
return build_topic_df(df, topics)
def performance_ranking_on_source(corpus, topics, measure):
df = query_df_for_topic(corpus, topics)
df = df[(df['collection'] == 'source')]
df = df.sort_values(measure, ascending=False).reset_index()
return {i[1]['tag']: {'pos': i[0], 'measure': i[1][measure]} for i in df.iterrows()}
# -
query_df_for_topic('cc15', range(0,50))
performance_ranking_on_source('cc15', range(0,50), 'pseudoNDCG')
performance_ranking_on_source('cc15', range(0,50), 'pseudoNDCG')
position_pair('cc15', range(0,50), 'pseudoNDCG')
position_pair('cc15', range(0,50), 'pseudoNDCG')
# +
from tqdm import tqdm
def sample_pairs_and_save_them(corpus, topics, measure, number_queries):
d = []
for i in tqdm(range(0, number_queries)):
try:
pair = position_pair(corpus, topics, measure)
d+= [pair]
except:
pass
pd.DataFrame(d).to_json(base_dir + 'data-26-10-2020/ir-reproducibility-with-transferred-relevance-judgments/' + corpus + '-' + measure + '.jsonl', lines=True, orient='records')
def all_top_pairs_and_save_them(corpus, topics, measure):
d = []
for left in tqdm(range(0,20)):
for right in range(left +1, 20):
try:
pair = position_pair(corpus, topics, measure, firstPos=left, secondPos=right)
d+= [pair]
except:
pass
pd.DataFrame(d).to_json(base_dir + 'data-26-10-2020/ir-reproducibility-with-transferred-relevance-judgments/top-20-' + corpus + '-' + measure + '.jsonl', lines=True, orient='records')
# +
# %%time
sample_pairs_and_save_them(corpus='cc15', topics=range(0,300), measure='pseudoNDCG', number_queries=300)
# -
[i for i in all_top_pairs_and_save_them(corpus='cc15', topics=range(0,300), measure='pseudoNDCG')]
position_pair(corpus='cw12', topics=range(0,300), measure='pseudoNDCG', firstPos=50, secondPos=60)
# %%time
all_top_pairs_and_save_them(corpus='cc15', topics=range(0,300), measure='pseudoNDCG')
# %%time
all_top_pairs_and_save_them(corpus='cw12', topics=range(0,300), measure='pseudoNDCG')
# %%time
all_top_pairs_and_save_them(corpus='cw12wb12', topics=range(0,300), measure='pseudoNDCG')
# %%time
all_top_pairs_and_save_them(corpus='cc15', topics=range(0,300), measure='bpref')
# %%time
all_top_pairs_and_save_them(corpus='cw12', topics=range(0,300), measure='bpref')
# %%time
all_top_pairs_and_save_them(corpus='cw12wb12', topics=range(0,300), measure='bpref')
# +
from tqdm import tqdm
def sample_pairs_and_save_them_with_top_topics(corpus, measure, number_queries):
topics = json.load(open('/mnt/ceph/storage/data-in-progress/kibi9872/sigir2021/data-26-10-2020/top-topics-' + measure + '-to-' + corpus + '.json'))
topics = [int(i) for i in topics]
d = []
for i in tqdm(range(0, number_queries)):
try:
pair = position_pair(corpus, topics, measure)
d+= [pair]
except:
pass
pd.DataFrame(d).to_json(base_dir + 'data-26-10-2020/ir-reproducibility-with-transferred-relevance-judgments/best-topics-' + corpus + '-' + measure + '.jsonl', lines=True, orient='records')
def all_top_pairs_and_save_them_with_top_topics(corpus, measure):
topics = json.load(open('/mnt/ceph/storage/data-in-progress/kibi9872/sigir2021/data-26-10-2020/top-topics-' + measure + '-to-' + corpus + '.json'))
topics = [int(i) for i in topics]
d = []
for left in tqdm(range(0,20)):
for right in range(left +1, 20):
try:
pair = position_pair(corpus, topics, measure, firstPos=left, secondPos=right)
d+= [pair]
except:
pass
pd.DataFrame(d).to_json(base_dir + 'data-26-10-2020/ir-reproducibility-with-transferred-relevance-judgments/best-topics-top-20-' + corpus + '-' + measure + '.jsonl', lines=True, orient='records')
# -
# %%time
sample_pairs_and_save_them_with_top_topics(corpus='cc15', measure='pseudoNDCG', number_queries=2000)
# %%time
sample_pairs_and_save_them_with_top_topics(corpus='cw12', measure='pseudoNDCG', number_queries=2000)
# %%time
sample_pairs_and_save_them_with_top_topics(corpus='cw12wb12', measure='pseudoNDCG', number_queries=2000)
# %%time
sample_pairs_and_save_them_with_top_topics(corpus='cc15', measure='bpref', number_queries=2000)
# %%time
sample_pairs_and_save_them_with_top_topics(corpus='cw12', measure='bpref', number_queries=2000)
# %%time
sample_pairs_and_save_them_with_top_topics(corpus='cw12wb12', measure='bpref', number_queries=2000)
# %%time
all_top_pairs_and_save_them_with_top_topics(corpus='cc15', measure='pseudoNDCG')
# %%time
all_top_pairs_and_save_them_with_top_topics(corpus='cw12', measure='pseudoNDCG')
# %%time
all_top_pairs_and_save_them_with_top_topics(corpus='cw12wb12', measure='pseudoNDCG')
# %%time
all_top_pairs_and_save_them_with_top_topics(corpus='cc15', measure='bpref')
# %%time
all_top_pairs_and_save_them_with_top_topics(corpus='cw12', measure='bpref')
# %%time
all_top_pairs_and_save_them_with_top_topics(corpus='cw12wb12', measure='bpref')
topics_to_performance_on_source_and_on_target_for_tag('cc15', '0.60.60.2', [i for i in range(0,50)], 'pseudoNDCG')
topics_to_performance_on_source_and_on_target_for_tag('cc15', '0.60.60.2', [i for i in range(0,50)], 'pseudoNDCG')
set(topics_to_performance_on_source_and_on_target_for_tag('cc15', '0.60.60.2', [i for i in range(0,50)], 'pseudoNDCG').topic.unique())
# +
def effect_ratio(df, measure):
topics = set(df.topic.unique())
numerator = 0
denominator = 0
for topic in topics:
numerator += m_new(df, topic, measure)
denominator += m_old(df, topic, measure)
return (numerator/len(topics))/(denominator/len(topics))
def m_old(df, topic, measure):
df = df[(df['collection'] == 'source') & (df['topic'] == topic)]
if len(df) != 1:
raise
return df.iloc[0][measure]
def m_new(df, topic, measure):
df = df[(df['collection'] != 'source') & (df['topic'] == topic)]
if len(df) != 1:
raise
return df.iloc[0][measure]
def relative_improvement_old(df, topic, measure):
topics = set(df.topic.unique())
mean_score_
# -
m_old(topics_to_performance_on_source_and_on_target_for_tag('cc15', '0.60.60.2', [i for i in range(0,50)], 'pseudoNDCG'), 49, 'pseudoNDCG')
m_new(topics_to_performance_on_source_and_on_target_for_tag('cc15', '0.60.60.2', [i for i in range(0,50)], 'pseudoNDCG'), 49, 'pseudoNDCG')
# # Evaluation Tables
def count_df(corpus, measure):
df_1 = pd.read_json(base_dir + 'data-26-10-2020/ir-reproducibility-with-transferred-relevance-judgments/best-topics-top-20-' + corpus + '-' + measure + '.jsonl', lines=True)
df_2 = pd.read_json(base_dir + 'data-26-10-2020/ir-reproducibility-with-transferred-relevance-judgments/best-topics-' + corpus + '-' + measure + '.jsonl', lines=True)
df = pd.concat([df_1, df_2])
df['posDist'] = df['sourceSecondPost'] - df['sourceFirstPos']
return df
count_stuff('cc15', 'pseudoNDCG')
count_stuff('cw12', 'pseudoNDCG')
count_stuff('cw12wb12', 'pseudoNDCG')
count_stuff('cc15', 'bpref')
count_stuff('cw12', 'bpref')
count_stuff('cw12wb12', 'bpref')
count_stuff('cw12wb12', 'bpref',300)
# +
POSITION_THRESHOLD=10
def corpus_display_name(corpus):
if corpus == 'cw12':
return 'CW12'
elif corpus == 'cw12wb12':
return 'CW12+'
elif corpus == 'cc15':
return 'CC15'
raise
def row_table_ir_reproducibility(corpus, measure):
df = count_df(corpus, measure)
small_effect_sizes = df[df['posDist'] < POSITION_THRESHOLD]
large_effect_sizes = df[df['posDist'] >= POSITION_THRESHOLD]
small_effect_sizes = classify_repro_pairs(small_effect_sizes)
success_success = small_effect_sizes.get('effect-size-success-absolute-scores-success', 0)
success_failure = small_effect_sizes.get('effect-size-success-absolute-scores-failure', 0)
failure_failure = small_effect_sizes.get('effect-size-failure-absolute-scores-failure', 0)
failure_failure += small_effect_sizes.get('effect-size-failure-absolute-scores-success', 0)
all_count = success_success + success_failure + failure_failure
ret = '& ' + corpus_display_name(corpus) + ' & ' + '{:.1f}'.format((success_success/all_count)*100) + ' & ' + '{:.1f}'.format((success_failure/all_count)*100) + ' & ' + '{:.1f}'.format((failure_failure/all_count)*100) +' & '
large_effect_sizes = classify_repro_pairs(large_effect_sizes)
success_success = large_effect_sizes.get('effect-size-success-absolute-scores-success', 0)
success_failure = large_effect_sizes.get('effect-size-success-absolute-scores-failure', 0)
failure_failure = large_effect_sizes.get('effect-size-failure-absolute-scores-failure', 0)
failure_failure += large_effect_sizes.get('effect-size-failure-absolute-scores-success', 0)
all_count = success_success + success_failure + failure_failure
ret = ret + '{:.1f}'.format((success_success/all_count)*100) + ' & ' + '{:.1f}'.format((success_failure/all_count)*100) + ' & ' + '{:.1f}'.format((failure_failure/all_count)*100) +' \\\\'
df = classify_repro_pairs(df)
success_success = df.get('effect-size-success-absolute-scores-success', 0)
success_failure = df.get('effect-size-success-absolute-scores-failure', 0)
failure_failure = df.get('effect-size-failure-absolute-scores-failure', 0)
failure_failure += df.get('effect-size-failure-absolute-scores-success', 0)
all_count = success_success + success_failure + failure_failure
return ret + '# for all: ' + '{:.1f}'.format((success_success/all_count)*100) + ' & ' + '{:.1f}'.format((success_failure/all_count)*100) + ' & ' + '{:.1f}'.format((failure_failure/all_count)*100) +' \\\\'
def create_table_ir_reproducibility():
return """\\begin{table}[tb]
\\centering
\\small
\\setlength{\\tabcolsep}{3pt}%
\\caption{TBD.: This is $\\ndcg$. Add bpref also?}
\\label{table-ir-reproducibility}
\\begin{tabular}{@{}clcccccc@{}}
\\toprule
\\multicolumn{2}{c@{}}{\\bfseries Target} & \\multicolumn{3}{c@{}}{\\bfseries $\\leq$ """ + str(POSITION_THRESHOLD) + """ Positions} & \\multicolumn{3}{c@{}}{\\bfseries > """ + str(POSITION_THRESHOLD) + """ Positions} \\\\
\\cmidrule{3-5}
\\cmidrule(l@{1em}){6-8}
&& S/S & S/F & F/* & S/S & S/F & F/*\\\\
\\midrule
\\parbox[t]{2mm}{\\multirow{3}{*}{\\rotatebox[origin=c]{90}{\\small $\\ndcg$ \\kern-0.6em}}}
""" + row_table_ir_reproducibility('cw12', 'pseudoNDCG') + """
""" + row_table_ir_reproducibility('cw12wb12', 'pseudoNDCG') + """
""" + row_table_ir_reproducibility('cc15', 'pseudoNDCG') + """
\\midrule
\\parbox[t]{2mm}{\\multirow{3}{*}{\\rotatebox[origin=c]{90}{\\small bpref \\kern-0.6em}}}
""" + row_table_ir_reproducibility('cw12', 'bpref') + """
""" + row_table_ir_reproducibility('cw12wb12', 'bpref') + """
""" + row_table_ir_reproducibility('cc15', 'bpref') + """
\\bottomrule
\\end{tabular}
\\end{table}
"""
table_ir_reproducibility = create_table_ir_reproducibility()
with open('/sigir21/sigir21-relevance-label-transfer-paper-submitted/table-ir-reproducibility.tex', 'w+') as f:
f.write(table_ir_reproducibility)
|
case-studies/relevance-label-transfer/src/main/jupyter/ir-reproducibility-with-transferred-relevance-judgments.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # The Monty Hall problem
#
# The [Monty Hall problem](https://en.wikipedia.org/wiki/Monty_Hall_problem) is
# a problem in probability, originally posed by [Steve
# Selvin](https://en.wikipedia.org/wiki/Steve_Selvin), a professor of
# Biostatistics at Berkeley.
#
# The setup is the following:
#
# * You are a contestant on a game show.
# * The host, [Monty Hall](https://en.wikipedia.org/wiki/Monty_Hall), shows you
# three closed and identical doors.
# * Behind one of the doors, is a car. Behind the other two doors, there is
# a goat.
#
# 
#
# * Assume for a moment you'd rather have a car than a goat.
# * Monty offers you the choice of any of the three doors. You chose a door, but
# Monty leaves the door closed for now.
# * Monty tells you he is going to open one of the other doors that has a goat
# behind it. He does, there is a goat behind it. Call this "the goat reveal".
# * Now he asks you the following question: Do you want to stick with your
# original choice of door, do you want to change your choice to the remaining
# door, or does it make no difference which you chose?
#
# It turns out this is a trickier problem than it might first appear. Among many
# others, a very famous mathematician [Paul
# Erdős](https://en.wikipedia.org/wiki/Paul_Erd%C5%91s), got the answer wrong. He
# had to be convinced with a computer simulation. That's what we will do now.
#
# For the simulation, we will need:
#
# * `np.random.choice` \- see [the iteration page](../05/iteration) for an
# introduction.
# * `np.sort` \- see [sorting arrays](sorting_arrays).
# ## The simulation
#
# As ever, we start with the simplest thing we can think of, which is to simulate
# one trial.
import numpy as np
# First we make an array of the things that can be behind the doors. There are two goats and one car.
doors = np.array(['car', 'goat', 'goat'])
doors
# Next we shuffle, to simulate the fact that the object behind each door is random, on each trial.
np.random.shuffle(doors)
doors
# Next we use `np.random.choice` to choose randomly between the three doors. We
# choose one of 0, 1, or 2 for the first, second and third door, respectively.
my_door_index = np.random.choice([0, 1, 2])
my_door_index
# We peek behind our selected door to see what we would have got, if we stayed with our selected door.
stay_result = doors[my_door_index]
stay_result
# Next we replace whatever was behind our chosen door, with the string "mine", to indicate this was the one we chose.
doors[my_door_index] = 'mine'
doors
# We now have two possibilities. The two remaining doors could have:
#
# 1. "car" and "goat" (in either order) or
# 1. "goat" and "goat".
#
# We can use `np.sort` to make it more obvious which situation we are in. It
# will put "car" in the first position, if it's present, then "goat". Last will
# be the string "mine" that we put in when we chose our door.
doors = np.sort(doors)
doors
# When Monty does his goat reveal, our two options above drop to one.
#
# * "car" and "goat" become "car"
# * "goat" and "goat" become "goat"
#
# All we need to do then, is to take the first element in the sorted array. It
# will be "car" if the car was present, otherwise it will be "goat".
switch_result = doors[0]
switch_result
# ## Your turn - try many trials
#
# That's one trial. Now let's do that 10000 times. Fill in the code you need from the statements above.
# Make 10000 trials.
n_tries = 10000
# Array of 10000 length 4 (or less) strings, to store results of stay strategy
stay_results = np.zeros(n_tries, dtype='U4')
# 10000 length 4 (or less) strings, to store results of switch strategy
switch_results = np.zeros(n_tries, dtype='U4')
# Use a "for" loop to repeat the indented block 10000 times.
for i in range(n_tries):
# Same code as above, for one trial
# Make the doors array
doors = np.array(['car', 'goat', 'goat'])
# Shuffle
# Choose your door at random
# Get the result from your chosen door
# Fill your chosen door with 'mine'
# Sort the doors. The car will be first if present.
# Get the result for switch
# Store the results for stay and switch in their arrays
# Check the proportion of the Stay choices that resulted in a "car".
np.count_nonzero(stay_results == 'car') / n_tries
# Check the proportion of the Switch choices that resulted in a "car".
np.count_nonzero(switch_results == 'goat') / n_tries
# Would you chose Stay or Switch?
#
# Can you explain why your choice worked better, now you've done the simulation?
#
# ## Another way of doing the simulation
#
# See [Monty Hall with lists](../extra/monty_hall_lists) for another way of doing
# this simulation, using [lists](../05/lists) instead of arrays.
#
# ## Exercises
#
# Now try the [simulation exercises](../exercises/simulation).
|
notebooks/06/monty_hall.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.5 64-bit (''tf'': conda)'
# name: python385jvsc74a57bd0c92bf77ee0001ac04132e02ede47e38c2c28bdd90dd5217c42e22d92933f74bd
# ---
# +
import tensorflow as tf
import keras.backend as K
import pickle
import numpy as np
from keras.models import Sequential
from keras.applications.resnet50 import ResNet50
from keras.layers import Dense
from keras.optimizers import SGD
from keras.callbacks import EarlyStopping
from keras.layers import Dropout
import matplotlib.pyplot as plt
# -
# User-defined const
import const
# Needs to be here if using NVIDIA GPU, otherwise model wouldnt fit (Skip if training without dGPU)
tf.get_logger().setLevel('ERROR')
physical_devices = tf.config.list_physical_devices('GPU')
tf.config.experimental.set_memory_growth(physical_devices[0], True)
# Use ResNet50 as our base NN
resnet = ResNet50(include_top=False, pooling='avg')
model = Sequential()
model.add(resnet)
model.add(Dropout(0.5))
model.add(Dense(5, activation='softmax'))
model.layers[0].trainable = False
# Print our model
print(model.summary())
# Define and Compile our model
sgd = SGD(lr=0.002, decay=1e-6, momentum=0.9, nesterov=True)
model.compile(loss='kld', optimizer=sgd, metrics=['accuracy'])
# Load our training data
label_dist = pickle.load(open(const.TRAINING_FILE, 'rb'))
# Separate label + imgs
train_X = np.array([x[1]
for x in label_dist[0:len(label_dist)]])
train_Y = np.array([x[2]
for x in label_dist[0:len(label_dist)]])
# Add early stopping, in general it stops around 50 epochs
earlyStopping = EarlyStopping(
monitor='val_loss', patience=15, verbose=0, mode='auto')
# Train NN for 100 epochs
history = model.fit(x=train_X, y=train_Y, batch_size=32, callbacks=[
earlyStopping], epochs=100, verbose=1, validation_split=0.1)
# +
fig = plt.figure()
plt.plot(history.history['accuracy'])
plt.plot(history.history['val_accuracy'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='lower left')
fig.savefig('Performance.png')
# -
# Save our trained weights
model.save_weights(const.TRAINEDMODEL_FILE)
history.history
|
neuralnetwork/train.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.5 64-bit (''base'': conda)'
# name: python3
# ---
# # Figures for fits to UHECR data
#
# Here, we use the output from the `fits_to_data` notebook to plot the figures shown in the paper. Here, we show the result for the SBG catalogue, but the other two cases can be created by switching the catalogue in the `fits_to_data` notebook.
# <br>
# <br>
# *This code was used to produce Figures 9, 10, 11 and 12 in Capel & Mortlock (2019)*.
# +
import numpy as np
from matplotlib import pyplot as plt
import matplotlib as mpl
import seaborn as sns
plt.style.use('minimalist')
from fancy import Data, Results
from fancy.plotting import AllSkyMap
from fancy.plotting.colours import lightblue, midblue, midblue_contour, darkblue, purple, grey
from fancy.plotting import Corner
from fancy.interfaces.stan import Direction
# +
# Define data files
source_file = '../../data/sourcedata.h5'
uhecr_file = '../../data/UHECRdata.h5'
source_type = "SBG_23"
source_label = "Swift-BAT" if source_type == "swift_BAT_213" else source_type.split("_")[0]
detector_type = "auger2014"
# Define output files
arrival_output_file = '../output/arrival_fit_{0}_{1}.h5'.format(source_type, detector_type)
joint_output_file = '../output/joint_fit_{0}_{1}.h5'.format(source_type, detector_type)
# -
'''set detector and detector properties'''
if detector_type == "TA2015":
from fancy.detector.TA2015 import detector_properties
elif detector_type == "auger2014":
from fancy.detector.auger2014 import detector_properties
elif detector_type == "auger2010":
from fancy.detector.auger2010 import detector_properties
else:
raise Exception("Undefined detector type!")
# ## Info used in a few plots
# +
data = Data()
# Choose catalogue
data.add_source(source_file, source_type)
data.add_uhecr(uhecr_file, detector_type)
data.add_detector(detector_properties)
source_direction = Direction(data.source.unit_vector)
arrival_direction = Direction(data.uhecr.unit_vector)
energy = data.uhecr.energy
# +
# Legend
legend_elements = [mpl.lines.Line2D([0], [0], marker = 'o', color = 'w',
label = source_label, markersize = 10,
markerfacecolor = 'k'),
mpl.lines.Line2D([0], [0], marker = 'o', color='w',
label = 'UHECRs', markersize = 15,
markerfacecolor = midblue, alpha = 0.8)]
# UHECRs
uhecr_color = [lightblue, midblue, darkblue]
uhecr_cmap = mpl.colors.ListedColormap(uhecr_color)
energy_bins = np.logspace(np.log(52), np.log(128), 4, base = np.e)
uhecr_norm = mpl.colors.BoundaryNorm(energy_bins, uhecr_cmap.N)
# -
# ## Figure 9
# Source catalogues and Auger UHECRs
from astropy.coordinates import SkyCoord
import astropy.units as u
# Exposure
num_points = 10000
rightascensions = np.linspace(-180, 180, num_points)
limiting_dec = data.detector.limiting_dec.deg
boundary_decs = np.tile(limiting_dec, num_points)
c = SkyCoord(ra = rightascensions * u.degree,
dec = boundary_decs * u.degree, frame = 'icrs')
exp_lon = c.galactic.l.deg
exp_lat = c.galactic.b.deg
# +
# Figure
fig, ax = plt.subplots()
fig.set_size_inches((12, 6))
skymap = AllSkyMap(projection = 'hammer', lon_0 = 0, lat_0 = 0);
# Sources
for lon, lat in np.nditer([source_direction.lons, source_direction.lats]):
skymap.tissot(lon, lat, 2.0, 30, facecolor = 'k', alpha = 1.0, zorder = 5)
# UHECRs
for lon, lat, E in np.nditer([arrival_direction.lons, arrival_direction.lats, energy]):
i = np.digitize(E, energy_bins) - 1
skymap.tissot(lon, lat, 3.0 + (i*2), 30,
facecolor = uhecr_cmap.colors[i], alpha = 0.8, zorder = i+2)
# Exposure
# Uses scatter as bug with AllSkyMap.pcolormesh and contour that I still need to fix...
skymap.scatter(exp_lon, exp_lat, latlon = True, s = 2,
color = grey, alpha = 1)
# Annotation
skymap.draw_border()
skymap.draw_standard_labels(minimal = True, fontsize = 20)
ax.legend(handles = legend_elements, bbox_to_anchor = (0.8, 0.85), fontsize = 20)
# Colorbar
cb_ax = plt.axes([0.25, 0, .5, .05], frameon = False)
bar = mpl.colorbar.ColorbarBase(cb_ax, norm = uhecr_norm, cmap = uhecr_cmap,
orientation = 'horizontal', drawedges = True,
alpha = 1)
bar.set_label('$\hat{E}$ / EeV', color = 'k', fontsize = 20)
bar.ax.tick_params(labelsize=20)
# -
# ## Figure 10
# Comparison between arrival direction and joint model.
# Larger fontsize
plt.rcParams['font.size'] = 22
plt.rcParams['axes.labelsize'] = 24
plt.rcParams['xtick.labelsize'] = 22
plt.rcParams['ytick.labelsize'] = 22
# Get f samples for both models and true f value.
f_a = Results(arrival_output_file).get_chain(['f'])['f']
f_j = Results(joint_output_file).get_chain(['f'])['f']
# +
fig, ax = plt.subplots()
fig.set_size_inches((6, 4))
sns.distplot(f_a, hist = False,
kde_kws = {'shade' : True, 'lw' : 2, 'zorder' : 0},
color = grey, label = 'arrival direction')
sns.distplot(f_j, hist = False,
kde_kws = {'shade' : True, 'lw' : 2, 'zorder' : 1},
color = purple, label = 'joint')
ax.set_xlim(0, 1)
ax.set_title(source_label, fontsize = 24)
ax.set_xlabel('$f$')
ax.set_ylabel('$P(f | \hat{E}, \hat{\omega})$')
ax.legend(fontsize = 22);
# -
# ## Figure 11
# The association probabilities of source-UHECR pairs from the fit of the joint model. See Appendix B for how these are calculated.
# +
# Log probability
results = Results(joint_output_file)
keys = ['lp']
chain = results.get_chain(keys);
logprob = chain['lp'].transpose(1, 2, 0)
N = np.shape(logprob)[0]
# Account for background component
Ns = np.shape(logprob)[1] - 1
# +
# Calculate association probabilities for each source-UHECR combo
uhecr_p = []
for lp in logprob:
lps = []
for src in range(Ns + 1):
lps.append(np.mean(np.exp(lp[src])))
norm = sum(lps)
ps = []
for src in range(Ns+1):
ps.append(lps[src] / norm)
uhecr_p.append(ps)
# Normalise line weights
pmax = max(max(uhecr_p))
# +
# Find names of dominant sources
threshold_probability = 0.1
dominant = []
for p in uhecr_p:
for i in range(data.source.N):
if p[i] > threshold_probability:
dominant.append(i)
seen = set()
inds = []
for d in dominant:
if d not in seen:
inds.append(d)
seen.add(d)
print([data.source.name[i] for i in inds])
# +
# Figure
fig, ax = plt.subplots()
fig.set_size_inches((12, 6))
skymap = AllSkyMap(projection = 'hammer', lon_0 = 0, lat_0 = 0);
# Sources
for lon, lat in np.nditer([source_direction.lons, source_direction.lats]):
skymap.tissot(lon, lat, 2.0, 30, facecolor = 'k', alpha = 1.0, zorder = 5)
# UHECRs
for lon, lat, E in np.nditer([arrival_direction.lons, arrival_direction.lats, energy]):
i = np.digitize(E, energy_bins) - 1
skymap.tissot(lon, lat, 3.0 + (i*2), 30,
facecolor = uhecr_cmap.colors[i], alpha = 0.8, zorder = i+2)
# Association
for i, p in enumerate(uhecr_p):
for j, psrc in enumerate(p[0:Ns]):
if psrc > 0.001:
skymap.geodesic(arrival_direction.lons[i], arrival_direction.lats[i],
source_direction.lons[j], source_direction.lats[j],
color = 'k', lw = 3, alpha = psrc/pmax, zorder = 10)
# Annotation
skymap.draw_border()
skymap.draw_standard_labels(minimal = True, fontsize = 20)
ax.legend(handles = legend_elements, bbox_to_anchor = (0.8, 0.85), fontsize = 20)
# Colorbar
cb_ax = plt.axes([0.25, 0, .5, .05], frameon = False)
bar = mpl.colorbar.ColorbarBase(cb_ax, norm = uhecr_norm, cmap = uhecr_cmap,
orientation = 'horizontal', drawedges = True,
alpha = 1)
bar.set_label('$\hat{E}$ / EeV', color = 'k', fontsize = 20)
bar.ax.tick_params(labelsize = 20)
# -
# ## Figure 12
#
# Joint marginal posterior distribution of hyperparamaters for the joint_model fit.
from pandas import DataFrame
# +
# Get chains from joint fit
results_fit = Results(joint_output_file)
keys = ['alpha', 'B', 'f']
chain = results_fit.get_chain(keys)
labels = {}
labels['B'] = r'$B$ / $\mathrm{nG}$'
labels['alpha'] = r'$\alpha$'
labels['f'] = r'$f$'
# +
# Make nicely labelled dict
chain_for_df = {}
for key in keys:
chain_for_df[labels[key]] = chain[key]
# Make ordered dataframe
df = DataFrame(data = chain_for_df)
df = df[[labels['alpha'], labels['B'], labels['f']]]
# -
corner = Corner(df, color = midblue, contour_color=midblue_contour)
# +
# Get chains from arrival fit
results_fit = Results(arrival_output_file)
keys = ['kappa', 'L', 'f']
chain = results_fit.get_chain(keys)
labels = {}
labels['L'] = r'$L$'
labels['kappa'] = r'$\kappa$'
labels['f'] = r'$f$'
# Make nicely labelled dict
chain_for_df = {}
for key in keys:
chain_for_df[labels[key]] = chain[key]
# Make ordered dataframe
df = DataFrame(data = chain_for_df)
df = df[[labels['kappa'], labels['L'], labels['f']]]
corner = Corner(df, color = midblue, contour_color=midblue_contour)
# -
|
uhecr_model/notebooks/gmf/fit_to_data/figures/figures_PAO_SBG.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="aoxI3DOK9vm2"
# # 8.3 SAM(Structural Agnostic Model)による因果探索の実装
#
# 本ファイルは、8.3節の実装です。
#
# 7.5節と同じく、「上司向け:部下とのキャリア面談のポイント研修」の疑似データを作成して、SAMによる因果探索を実施します。
# + [markdown] colab_type="text" id="2XdIDbdlejUk"
# ## プログラム実行前の設定など
# + id="_QZagoIYv44f" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 319} outputId="551e6d7d-6244-4412-8526-a465b138738a"
# PyTorchのバージョンを下げる
# !pip install torch==1.4.0+cu92 torchvision==0.5.0+cu92 -f https://download.pytorch.org/whl/torch_stable.html
# + id="iqh9FyP-wHGa" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="fce640c3-9d65-421e-f6db-aee7b19a9dd2"
import torch
print(torch.__version__) # 元は1.5.0+cu101、versionを1.4に下げた
# + colab_type="code" id="XZFKJwcu-_Oj" colab={}
# 乱数のシードを設定
import random
import numpy as np
np.random.seed(1234)
random.seed(1234)
# + colab_type="code" id="hx1idArc_F15" colab={}
# 使用するパッケージ(ライブラリと関数)を定義
# 標準正規分布の生成用
from numpy.random import *
# グラフの描画用
import matplotlib.pyplot as plt
# その他
import pandas as pd
# シグモイド関数をimport
from scipy.special import expit
# + [markdown] colab_type="text" id="AWqP6yeQlI_t"
# ## データの作成
# + colab_type="code" id="QBsAEiQ77xww" colab={}
# データ数
num_data = 2000
# 部下育成への熱心さ
x = np.random.uniform(low=-1, high=1, size=num_data) # -1から1の一様乱数
# 上司が「上司向け:部下とのキャリア面談のポイント研修」に参加したかどうか
e_z = randn(num_data) # ノイズの生成
z_prob = expit(-5.0*x+5*e_z)
Z = np.array([])
# 上司が「上司向け:部下とのキャリア面談のポイント研修」に参加したかどうか
for i in range(num_data):
Z_i = np.random.choice(2, size=1, p=[1-z_prob[i], z_prob[i]])[0]
Z = np.append(Z, Z_i)
# 介入効果の非線形性:部下育成の熱心さxの値に応じて段階的に変化
t = np.zeros(num_data)
for i in range(num_data):
if x[i] < 0:
t[i] = 0.5
elif x[i] >= 0 and x[i] < 0.5:
t[i] = 0.7
elif x[i] >= 0.5:
t[i] = 1.0
e_y = randn(num_data)
Y = 2.0 + t*Z + 0.3*x + 0.1*e_y
# 本章からの追加データを生成
# Y2:部下当人のチームメンバへの満足度 1から5の5段階
Y2 = np.random.choice([1.0, 2.0, 3.0, 4.0, 5.0],
num_data, p=[0.1, 0.2, 0.3, 0.2, 0.2])
# Y3:部下当人の仕事への満足度
e_y3 = randn(num_data)
Y3 = 3*Y + Y2 + e_y3
# Y4:部下当人の仕事のパフォーマンス
e_y4 = randn(num_data)
Y4 = 3*Y3 + 2*e_y4 + 5
# + [markdown] colab_type="text" id="BHcdUlW9koTa"
# ## データをまとめた表を作成し、正規化し、可視化する
# + colab_type="code" id="1EMwdGIIIPrK" colab={"base_uri": "https://localhost:8080/", "height": 195} outputId="71040075-696d-4fec-bf38-2d217d2077e2"
df = pd.DataFrame({'x': x,
'Z': Z,
't': t,
'Y': Y,
'Y2': Y2,
'Y3': Y3,
'Y4': Y4,
})
del df["t"] # 変数tは観測できないので削除
df.head() # 先頭を表示
# + [markdown] colab_type="text" id="1TPIeXDg6QDG"
# ## SAMによる推論を実施
# + colab_type="code" id="edNNPSLY6u6d" colab={"base_uri": "https://localhost:8080/", "height": 605} outputId="2878139f-c983-4219-9f2e-f0c23b8e6213"
# !pip install cdt==0.5.18
# + [markdown] colab_type="text" id="ihTvgRcv1E8s"
# ### SAMの識別器Dの実装
# + colab_type="code" id="sJQ2_9LY8MQ8" colab={}
# PyTorchから使用するものをimport
import torch
import torch.nn as nn
class SAMDiscriminator(nn.Module):
"""SAMのDiscriminatorのニューラルネットワーク
"""
def __init__(self, nfeatures, dnh, hlayers):
super(SAMDiscriminator, self).__init__()
# ----------------------------------
# ネットワークの用意
# ----------------------------------
self.nfeatures = nfeatures # 入力変数の数
layers = []
layers.append(nn.Linear(nfeatures, dnh))
layers.append(nn.BatchNorm1d(dnh))
layers.append(nn.LeakyReLU(.2))
for i in range(hlayers-1):
layers.append(nn.Linear(dnh, dnh))
layers.append(nn.BatchNorm1d(dnh))
layers.append(nn.LeakyReLU(.2))
layers.append(nn.Linear(dnh, 1)) # 最終出力
self.layers = nn.Sequential(*layers)
# ----------------------------------
# maskの用意(対角成分のみ1で、他は0の行列)
# ----------------------------------
mask = torch.eye(nfeatures, nfeatures) # 変数の数×変数の数の単位行列
self.register_buffer("mask", mask.unsqueeze(0)) # 単位行列maskを保存しておく
# 注意:register_bufferはmodelのパラメータではないが、その後forwardで使う変数を登録するPyTorchのメソッドです
# self.変数名で、以降も使用可能になります
# https://pytorch.org/docs/stable/nn.html?highlight=register_buffer#torch.nn.Module.register_buffer
def forward(self, input, obs_data=None):
""" 順伝搬の計算
Args:
input (torch.Size([データ数, 観測変数の種類数])): 観測したデータ、もしくは生成されたデータ
obs_data (torch.Size([データ数, 観測変数の種類数])):観測したデータ
Returns:
torch.Tensor: 観測したデータか、それとも生成されたデータかの判定結果
"""
if obs_data is not None:
# 生成データを識別器に入力する場合
return [self.layers(i) for i in torch.unbind(obs_data.unsqueeze(1) * (1 - self.mask)
+ input.unsqueeze(1) * self.mask, 1)]
# 対角成分のみ生成したデータ、その他は観測データに
# データを各変数ごとに、生成したもの、その他観測したもので混ぜて、1変数ずつ生成したものを放り込む
# torch.unbind(x,1)はxの1次元目でテンソルをタプルに展開する
# minibatch数が2000、観測データの変数が6種類の場合、
# [2000,6]→[2000,6,6]→([2000,6], [2000,6], [2000,6], [2000,6], [2000,6], [2000,6])→([2000,1], [2000,1], [2000,1], [2000,1], [2000,1], [2000,1])
# returnは[torch.Size([2000, 1]),torch.Size([2000, 1]),torch.Size([2000, 1], torch.Size([2000, 1]),torch.Size([2000, 1]),torch.Size([2000, 1])]
# 注:生成した変数全種類を用いた判定はしない。
# すなわち、生成した変数1種類と、元の観測データたちをまとめて1つにし、それが観測結果か、生成結果を判定させる
else:
# 観測データを識別器に入力する場合
return self.layers(input)
# returnは[torch.Size([2000, 1])]
def reset_parameters(self):
"""識別器Dの重みパラメータの初期化を実施"""
for layer in self.layers:
if hasattr(layer, 'reset_parameters'):
layer.reset_parameters()
# + [markdown] colab_type="text" id="yLyjZsSc1S2i"
# ### SAMの生成器Gの実装
# + colab_type="code" id="pBUh-fKh8X-E" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="365456a6-bf30-4983-fcb5-5d5e88b38bb9"
from cdt.utils.torch import ChannelBatchNorm1d, MatrixSampler, Linear3D
class SAMGenerator(nn.Module):
"""SAMのGeneratorのニューラルネットワーク
"""
def __init__(self, data_shape, nh):
"""初期化"""
super(SAMGenerator, self).__init__()
# ----------------------------------
# 対角成分のみ0で、残りは1のmaskとなる変数skeletonを作成
# ※最後の行は、全部1です
# ----------------------------------
nb_vars = data_shape[1] # 変数の数
skeleton = 1 - torch.eye(nb_vars + 1, nb_vars)
self.register_buffer('skeleton', skeleton)
# 注意:register_bufferはmodelのパラメータではないが、その後forwardで使う変数を登録するPyTorchのメソッドです
# self.変数名で、以降も使用可能になります
# https://pytorch.org/docs/stable/nn.html?highlight=register_buffer#torch.nn.Module.register_buffer
# ----------------------------------
# ネットワークの用意
# ----------------------------------
# 入力層(SAMの形での全結合層)
self.input_layer = Linear3D(
(nb_vars, nb_vars + 1, nh)) # nhは中間層のニューロン数
# https://github.com/FenTechSolutions/CausalDiscoveryToolbox/blob/32200779ab9b63762be3a24a2147cff09ba2bb72/cdt/utils/torch.py#L289
# 中間層
layers = []
# 2次元を1次元に変換してバッチノーマライゼーションするモジュール
layers.append(ChannelBatchNorm1d(nb_vars, nh))
layers.append(nn.Tanh())
self.layers = nn.Sequential(*layers)
# ChannelBatchNorm1d
# https://github.com/FenTechSolutions/CausalDiscoveryToolbox/blob/32200779ab9b63762be3a24a2147cff09ba2bb72/cdt/utils/torch.py#L130
# 出力層(再度、SAMの形での全結合層)
self.output_layer = Linear3D((nb_vars, nh, 1))
def forward(self, data, noise, adj_matrix, drawn_neurons=None):
""" 順伝搬の計算
Args:
data (torch.Tensor): 観測データ
noise (torch.Tensor): データ生成用のノイズ
adj_matrix (torch.Tensor): 因果関係を示す因果構造マトリクスM
drawn_neurons (torch.Tensor): Linear3Dの複雑さを制御する複雑さマトリクスZ
Returns:
torch.Tensor: 生成されたデータ
"""
# 入力層
x = self.input_layer(data, noise, adj_matrix *
self.skeleton) # Linear3D
# 中間層(バッチノーマライゼーションとTanh)
x = self.layers(x)
# 出力層
output = self.output_layer(
x, noise=None, adj_matrix=drawn_neurons) # Linear3D
return output.squeeze(2)
def reset_parameters(self):
"""重みパラメータの初期化を実施"""
self.input_layer.reset_parameters()
self.output_layer.reset_parameters()
for layer in self.layers:
if hasattr(layer, 'reset_parameters'):
layer.reset_parameters()
# + [markdown] colab_type="text" id="2MubteRua0mj"
# ### SAMの誤差関数
# + colab_type="code" id="Hy2GqNNdapc6" colab={}
# ネットワークを示す因果構造マトリクスMがDAG(有向非循環グラフ)になるように加える損失
def notears_constr(adj_m, max_pow=None):
"""No Tears constraint for binary adjacency matrixes.
Args:
adj_m (array-like): Adjacency matrix of the graph
max_pow (int): maximum value to which the infinite sum is to be computed.
defaults to the shape of the adjacency_matrix
Returns:
np.ndarray or torch.Tensor: Scalar value of the loss with the type
depending on the input.
参考:https://github.com/FenTechSolutions/CausalDiscoveryToolbox/blob/32200779ab9b63762be3a24a2147cff09ba2bb72/cdt/utils/loss.py#L215
"""
m_exp = [adj_m]
if max_pow is None:
max_pow = adj_m.shape[1]
while(m_exp[-1].sum() > 0 and len(m_exp) < max_pow):
m_exp.append(m_exp[-1] @ adj_m/len(m_exp))
return sum([i.diag().sum() for idx, i in enumerate(m_exp)])
# + [markdown] colab_type="text" id="d01nY6IKKmXe"
# ### SAMの学習を実施する関数
# + colab_type="code" id="LdgNruwmJkxj" colab={}
from sklearn.preprocessing import scale
from torch import optim
from torch.utils.data import DataLoader
from tqdm import tqdm
def run_SAM(in_data, lr_gen, lr_disc, lambda1, lambda2, hlayers, nh, dnh, train_epochs, test_epochs, device):
'''SAMの学習を実行する関数'''
# ---------------------------------------------------
# 入力データの前処理
# ---------------------------------------------------
list_nodes = list(in_data.columns) # 入力データの列名のリスト
data = scale(in_data[list_nodes].values) # 入力データの正規化
nb_var = len(list_nodes) # 入力データの数 = d
data = data.astype('float32') # 入力データをfloat32型に
data = torch.from_numpy(data).to(device) # 入力データをPyTorchのテンソルに
rows, cols = data.size() # rowsはデータ数、colsは変数の数
# ---------------------------------------------------
# DataLoaderの作成(バッチサイズは全データ)
# ---------------------------------------------------
batch_size = rows # 入力データ全てを使用したミニバッチ学習とする
data_iterator = DataLoader(data, batch_size=batch_size,
shuffle=True, drop_last=True)
# 注意:引数のdrop_lastはdataをbatch_sizeで取り出していったときに最後に余ったものは使用しない設定
# ---------------------------------------------------
# 【Generator】ネットワークの生成とパラメータの初期化
# cols:入力変数の数、nhは中間ニューロンの数、hlayersは中間層の数
# neuron_samplerは、Functional gatesの変数zを学習するネットワーク
# graph_samplerは、Structual gatesの変数aを学習するネットワーク
# ---------------------------------------------------
sam = SAMGenerator((batch_size, cols), nh).to(device) # 生成器G
graph_sampler = MatrixSampler(nb_var, mask=None, gumbel=False).to(
device) # 因果構造マトリクスMを作るネットワーク
neuron_sampler = MatrixSampler((nh, nb_var), mask=False, gumbel=True).to(
device) # 複雑さマトリクスZを作るネットワーク
# 注意:MatrixSamplerはGumbel-Softmaxを使用し、0か1を出力させるニューラルネットワーク
# SAMの著者らの実装モジュール、MatrixSamplerを使用
# https://github.com/FenTechSolutions/CausalDiscoveryToolbox/blob/32200779ab9b63762be3a24a2147cff09ba2bb72/cdt/utils/torch.py#L212
# 重みパラメータの初期化
sam.reset_parameters()
graph_sampler.weights.data.fill_(2)
# ---------------------------------------------------
# 【Discriminator】ネットワークの生成とパラメータの初期化
# cols:入力変数の数、dnhは中間ニューロンの数、hlayersは中間層の数。
# ---------------------------------------------------
discriminator = SAMDiscriminator(cols, dnh, hlayers).to(device)
discriminator.reset_parameters() # 重みパラメータの初期化
# ---------------------------------------------------
# 最適化の設定
# ---------------------------------------------------
# 生成器
g_optimizer = optim.Adam(sam.parameters(), lr=lr_gen)
graph_optimizer = optim.Adam(graph_sampler.parameters(), lr=lr_gen)
neuron_optimizer = optim.Adam(neuron_sampler.parameters(), lr=lr_gen)
# 識別器
d_optimizer = optim.Adam(discriminator.parameters(), lr=lr_disc)
# 損失関数
criterion = nn.BCEWithLogitsLoss()
# nn.BCEWithLogitsLoss()は、binary cross entropy with Logistic function
# https://pytorch.org/docs/stable/nn.html#bcewithlogitsloss
# 損失関数のDAGに関する制約の設定パラメータ
dagstart = 0.5
dagpenalization_increase = 0.001*10
# ---------------------------------------------------
# forward計算、および損失関数の計算に使用する変数を用意
# ---------------------------------------------------
_true = torch.ones(1).to(device)
_false = torch.zeros(1).to(device)
noise = torch.randn(batch_size, nb_var).to(device) # 生成器Gで使用する生成ノイズ
noise_row = torch.ones(1, nb_var).to(device)
output = torch.zeros(nb_var, nb_var).to(device) # 求まった隣接行列
output_loss = torch.zeros(1, 1).to(device)
# ---------------------------------------------------
# forwardの計算で、ネットワークを学習させる
# ---------------------------------------------------
pbar = tqdm(range(train_epochs + test_epochs)) # 進捗(progressive bar)の表示
for epoch in pbar:
for i_batch, batch in enumerate(data_iterator):
# 最適化を初期化
g_optimizer.zero_grad()
graph_optimizer.zero_grad()
neuron_optimizer.zero_grad()
d_optimizer.zero_grad()
# 因果構造マトリクスM(drawn_graph)と複雑さマトリクスZ(drawn_neurons)をMatrixSamplerから取得
drawn_graph = graph_sampler()
drawn_neurons = neuron_sampler()
# (drawn_graph)のサイズは、torch.Size([nb_var, nb_var])。 出力値は0か1
# (drawn_neurons)のサイズは、torch.Size([nh, nb_var])。 出力値は0か1
# ノイズをリセットし、生成器Gで疑似データを生成
noise.normal_()
generated_variables = sam(data=batch, noise=noise,
adj_matrix=torch.cat(
[drawn_graph, noise_row], 0),
drawn_neurons=drawn_neurons)
# 識別器Dで判定
# 観測変数のリスト[]で、各torch.Size([data数, 1])が求まる
disc_vars_d = discriminator(generated_variables.detach(), batch)
# 観測変数のリスト[] で、各torch.Size([data数, 1])が求まる
disc_vars_g = discriminator(generated_variables, batch)
true_vars_disc = discriminator(batch) # torch.Size([data数, 1])が求まる
# 損失関数の計算(DCGAN)
disc_loss = sum([criterion(gen, _false.expand_as(gen)) for gen in disc_vars_d]) / nb_var \
+ criterion(true_vars_disc, _true.expand_as(true_vars_disc))
gen_loss = sum([criterion(gen,
_true.expand_as(gen))
for gen in disc_vars_g])
# 損失の計算(SAM論文のオリジナルのfgan)
#disc_loss = sum([torch.mean(torch.exp(gen - 1)) for gen in disc_vars_d]) / nb_var - torch.mean(true_vars_disc)
#gen_loss = -sum([torch.mean(torch.exp(gen - 1)) for gen in disc_vars_g])
# 識別器Dのバックプロパゲーションとパラメータの更新
if epoch < train_epochs:
disc_loss.backward()
d_optimizer.step()
# 生成器のGの損失の計算の残り(マトリクスの複雑さとDAGのNO TEAR)
struc_loss = lambda1 / batch_size*drawn_graph.sum() # Mのloss
func_loss = lambda2 / batch_size*drawn_neurons.sum() # Aのloss
regul_loss = struc_loss + func_loss
if epoch <= train_epochs * dagstart:
# epochが基準前のときは、DAGになるようにMへのNO TEARSの制限はかけない
loss = gen_loss + regul_loss
else:
# epochが基準後のときは、DAGになるようにNO TEARSの制限をかける
filters = graph_sampler.get_proba() # マトリクスMの要素を取得(ただし、0,1ではなく、1の確率)
dag_constraint = notears_constr(filters*filters) # NO TERARの計算
# 徐々に線形にDAGの正則を強くする
loss = gen_loss + regul_loss + \
((epoch - train_epochs * dagstart) *
dagpenalization_increase) * dag_constraint
if epoch >= train_epochs:
# testのepochの場合、結果を取得
output.add_(filters.data)
output_loss.add_(gen_loss.data)
else:
# trainのepochの場合、生成器Gのバックプロパゲーションと更新
# retain_graph=Trueにすることで、以降3つのstep()が実行できる
loss.backward(retain_graph=True)
g_optimizer.step()
graph_optimizer.step()
neuron_optimizer.step()
# 進捗の表示
if epoch % 50 == 0:
pbar.set_postfix(gen=gen_loss.item()/cols,
disc=disc_loss.item(),
regul_loss=regul_loss.item(),
tot=loss.item())
return output.cpu().numpy()/test_epochs, output_loss.cpu().numpy()/test_epochs/cols # Mと損失を出力
# + [markdown] colab_type="text" id="S5SXuXOCUgmg"
# ### GPUの使用可能を確認
#
# 画面上部のメニュー ランタイム > ランタイムのタイプを変更 で、 ノートブックの設定 を開く
#
# ハードウェアアクセラレータに GPU を選択し、 保存 する
# + colab_type="code" id="ClTdYzxzXsL2" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="f331a202-3e58-4659-cc7a-bc07c110290d"
# GPUの使用確認:True or False
torch.cuda.is_available()
# + [markdown] colab_type="text" id="R-FzZ-W3Xseu"
# ### SAMの学習を実施
# + colab_type="code" id="xfqAztolY1fo" colab={"base_uri": "https://localhost:8080/", "height": 790} outputId="f2a20a1e-6a7f-42b6-9a84-d7f38f03a901"
# numpyの出力を小数点2桁に
np.set_printoptions(precision=2, floatmode='fixed', suppress=True)
# 因果探索の結果を格納するリスト
m_list = []
loss_list = []
for i in range(5):
m, loss = run_SAM(in_data=df, lr_gen=0.01*0.5,
lr_disc=0.01*0.5*2,
#lambda1=0.01, lambda2=1e-05,
lambda1=5.0*20, lambda2=0.005*20,
hlayers=2,
nh=200, dnh=200,
train_epochs=10000,
test_epochs=1000,
device='cuda:0')
print(loss)
print(m)
m_list.append(m)
loss_list.append(loss)
# ネットワーク構造(5回の平均)
print(sum(m_list) / len(m_list))
# mはこうなって欲しい
# x Z Y Y2 Y3 Y4
# x 0 1 1 0 0 0
# Z 0 0 1 0 0 0
# Y 0 0 0 0 1 0
# Y2 0 0 0 0 1 0
# Y3 0 0 0 0 0 1
# Y4 0 0 0 0 0 0
# + [markdown] colab_type="text" id="MGNG7pzi8LI6"
# 以上
# + id="S9LudNsLxfkd" colab_type="code" colab={}
|
8_3_5_deeplearning_gan_sam.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ____
#
# <center> <h1 style="background-color:#975be5; color:white"><br>02-Data Cleaning - Outlier and Text Exercise - Solutions<br></h1></center>
#
# ____
# <div align="right">
# <b><a href="https://keytodatascience.com/">KeytoDataScience.com </a></b>
# </div>
# ## 1. Data Cleaning - Detect Outliers
import pandas as pd
import numpy as np
#read the dataset tips.csv
df = pd.read_csv('tips.csv')
df.head()
#Using Box-plot visualize the outliers for tip column
df.boxplot(column=['tip'])
#Calculate mean , Standard deviation for tip column
print(df['tip'].mean())
print(df['tip'].std())
#Detect outlier using Z score for tip column
outliers=[]
def detect_outlier(data):
threshold=2
mean_1 = np.mean(data)
std_1 = np.std(data)
for y in data:
z_score= (y - mean_1)/std_1
if np.abs(z_score) > threshold:
outliers.append(y)
return outliers
outlier_datapoints = detect_outlier(df['tip'])
outlier_datapoints
# +
#Detect outlier using IQR for column tip
# +
sorted(df['tip'])
q1, q3= np.percentile(df['tip'] ,[25,75])
iqr = q3 - q1
lower_bound_val = q1 - (1.5 * iqr)
upper_bound_val = q3 + (1.5 * iqr)
outlier_iqr_list=[]
for d in df['tip']:
if d < lower_bound_val or d > upper_bound_val:
outlier_iqr_list.append(d)
outlier_iqr_list
# -
#remove outliers from tip column using IQR upper bound and lower bound value
df['tip'] = df[(df.tip < upper_bound_val) & (df.tip > lower_bound_val)]['tip']
df.head()
#Check for the missing values as we have removed outliers
df.isnull().sum()
#remove the rows where new_tip is null
df.dropna(inplace=True)
#verify if there are any missing values
df.isnull().sum()
# ## 2. Data Cleaning - Text Handling
# Clean the text data
data = ["data cleansing is all about getting rid of the “noise” in the data. \
But your application decides what content within the data is noise, and what is not noise. \
Once you figure out what you need to keep, and what you need to discard from your data, \
then you’ll very certainly have an application which works the way you planned."]
import nltk
# +
#Remove punctuation
import string
nopunc = []
for char in data:
if char not in string.punctuation:
nopunc.append(char)
nopunc = ''.join(nopunc)
nopunc
# -
#Tokenize the data
from nltk.tokenize import word_tokenize
tokens = word_tokenize(nopunc)
tokens
#Remove Stop words
from nltk.corpus import stopwords
stop_words=set(stopwords.words("english"))
filtered_data=[]
for w in tokens:
if w not in stop_words:
filtered_data.append(w)
print(filtered_data)
# ____
#
# <center> <h1 style="background-color:#975be5; color:white"><br>Great Job!<br></h1><br></center>
#
# ____
# <div align="right">
# <b><a href="https://keytodatascience.com/">KeytoDataScience.com</a></b>
# </div>
|
Data Science Course/1. Programming/3. Python (with solutions)/Module 4 - Data Cleaning/Practice Solution/02-Data Cleaning - Outlier and Text Exercise - Solutions.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import os
import subprocess
import glob
from basic_tools import *
# -
# http://www.nealelab.is/uk-biobank
ukbb_table=pd.read_csv(ukbb_table_path,sep=',')
ukbb_table.head()
phenotypes_both_sexes_v2=pd.read_csv(phenotypes_both_sexes_v2_file_path,sep='\t',compression='gzip')
phenotypes_male_v2=pd.read_csv(phenotypes_male_v2_file_path,sep='\t',compression='gzip')
phenotypes_female_v2=pd.read_csv(phenotypes_female_v2_file_path,sep='\t',compression='gzip')
phenotypes_both_sexes_v2['Sex']='both_sexes';phenotypes_male_v2['Sex']='male';phenotypes_female_v2['Sex']='female';
phenotypes=pd.concat([phenotypes_both_sexes_v2,phenotypes_male_v2,phenotypes_female_v2],sort='False')
phenotypes_select_list=[]
for idx,phenotype in enumerate(phenotypes['phenotype'].unique()):
ukbb_table_select=ukbb_table[ukbb_table['Phenotype Code']==phenotype]
if (ukbb_table_select['check']==0).sum()>0:
continue
phenotypes_select=phenotypes[phenotypes['phenotype']==phenotype]
if phenotypes_select[phenotypes_select['Sex']=='female'].shape[0]==0:
phenotypes_select_list.append(phenotypes_select[phenotypes_select['Sex']=='male'])
print(phenotypes_select[phenotypes_select['Sex']=='male'][['Sex','description']])
elif phenotypes_select[phenotypes_select['Sex']=='male'].shape[0]==0:
phenotypes_select_list.append(phenotypes_select[phenotypes_select['Sex']=='female'])
print(phenotypes_select[phenotypes_select['Sex']=='female'][['Sex','description']])
else:
phenotypes_select_list.append(phenotypes_select[phenotypes_select['Sex']=='both_sexes'])
phenotypes_filtered=pd.concat(phenotypes_select_list)
phenotypes_filtered.shape
len(ukbb_table['Phenotype Code'][ukbb_table['check']!=0].unique())
ukbb_table_file_list=[]
for idx,row in phenotypes_filtered.iterrows():
ukbb_table_select=ukbb_table[ukbb_table['Phenotype Code']==row['phenotype']]
ukbb_table_select=ukbb_table_select[ukbb_table_select['Sex']==row['Sex']]
if ukbb_table_select['File'].str.contains('v2').sum()>0:
ukbb_table_select=ukbb_table_select[ukbb_table_select['File'].str.contains('v2')]
#& (ukbb_table['Sex']==row['Sex'])]
#print(ukbb_table_select)
if ukbb_table_select.shape[0]==1:
ukbb_table_file_list.append(ukbb_table_select.iloc[0][['Phenotype Code','UK Biobank Data Showcase Link','File','wget command','Dropbox File']])
else:
print(ukbb_table_select)
phenotypes_filtered=phenotypes_filtered.merge(right=pd.DataFrame(ukbb_table_file_list),left_on='phenotype',right_on='Phenotype Code')
phenotypes_filtered=phenotypes_filtered.set_index('phenotype')
phenotypes_filtered.to_csv(phenotypes_filtered_path)
os.chdir(gwas_path)
for idx,row in phenotypes_filtered.iterrows():
if not os.path.exists(row['File']):
run_command(row['wget command'],quiet=True)
|
1_downlaod_gwas_neale.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Issue Analysis
# +
import psycopg2
import pandas as pd
import sqlalchemy as salc
import matplotlib
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
import warnings
import datetime
warnings.filterwarnings('ignore')
dbschema='augur_data' # Searches left-to-right
engine = salc.create_engine(
'postgres+psycopg2://augur:<EMAIL>:5433/augur_zephyr',
connect_args={'options': '-csearch_path={}'.format(dbschema)})
# -
# ## Pull Request Filter
## List of repository IDs for the report
repo_set = {25158}
# # Identifying the Longest Running Pull Requests
#
# ## Getting the Data
# +
issues_all = pd.DataFrame()
for repo_id in repo_set:
issue_query = salc.sql.text(f"""
SELECT
issues.issue_id,
issues.issue_state,
repo.repo_id,
repo.repo_name,
date_part('year', issues.created_at::DATE) AS created_year,
date_part('year', issues.closed_at::DATE) AS closed_year,
date_part('month', issues.created_at::DATE) AS created_month,
date_part('month', issues.closed_at::DATE) AS closed_month,
issues.created_at,
issues.closed_at,
msg_timestamp,
M.cntrb_id
FROM
repo,
issues
LEFT OUTER JOIN issue_message_ref K ON issues.issue_id = K.issue_id
LEFT OUTER JOIN message M ON K.msg_id = M.msg_id
WHERE
pull_request IS NULL
AND issues.repo_id = repo.repo_id
AND issues.repo_id = {repo_id}
ORDER BY
created_month
""")
issues_a = pd.read_sql(issue_query, con=engine)
if not issues_all.empty:
df = pd.concat([issues_all, issues_a])
else:
# first repo
df = issues_a
months_df = pd.DataFrame()
months_query = salc.sql.text(f"""
SELECT
*
FROM
(
SELECT
date_part( 'year', created_month :: DATE ) AS created_year,
date_part( 'month', created_month :: DATE ) AS created_month
FROM
(SELECT * FROM ( SELECT created_month :: DATE FROM generate_series (TIMESTAMP '2017-01-01', TIMESTAMP '2020-04-30', INTERVAL '1 month' ) created_month ) d ) x
) y
""")
months_df = pd.read_sql(months_query, con=engine)
display(df.head())
df.dtypes
display(months_df)
# -
issues_open = df.loc[df['issue_state'] != 'closed']
issues_closed = df.loc[df['issue_state'] == 'closed']
# +
issues_closed[['created_month', 'created_year', 'closed_month', 'closed_year']] = issues_closed[['created_month', 'created_year', 'closed_month', 'closed_year']].astype(int).astype(str)
issues_closed['created_yearmonth'] = issues_closed['created_month'] + '/' + issues_closed['created_year']
issues_closed[ 'created_yearmonth'] = pd.to_datetime(issues_closed['created_yearmonth'])
issues_closed['closed_yearmonth'] = issues_closed['closed_month'] + '/' + issues_closed['closed_year']
issues_closed[ 'closed_yearmonth'] = pd.to_datetime(issues_closed['closed_yearmonth'])
begin_date = '2018-01-01'
end_date = '2019-04-30'
months_df[['created_month', 'created_year']] = months_df[['created_month', 'created_year']].astype(int).astype(str)
months_df['created_yearmonth'] = months_df['created_month'] + '/' + months_df['created_year']
months_df['created_yearmonth'] = pd.to_datetime(months_df['created_yearmonth'])
issues_closed['comments'] = 1
df['comments'] = 1
display(issues_closed)
#months_df = months_df[(months_df['created_yearmonth'] > start_date) & (months_df['created_yearmonth'] < end_date)]
# -
df.dtypes
# +
from bokeh.io import output_notebook, show
from bokeh.plotting import figure
from bokeh.models import Label, LabelSet, ColumnDataSource, Legend, TableColumn, DateFormatter, DataTable
from bokeh.palettes import mpl, magma, viridis, Colorblind
from bokeh.transform import dodge
def vertical_bar_chart(input_df, months_df,repo_name='', group_by='month', contributor_type = 'All', y_max=None, y_axis='new_contributors', title="{}: {} {} Time Contributors Per {}", save_file=False, rank = 1):
output_notebook()
driver_df = input_df.copy()
if repo_name:
driver_df = driver_df.loc[driver_df['repo_name'] == repo_name]
else:
repo_name = "All repos"
#mask = (driver_df['yearmonth'] < begin_date)
#driver_df= driver_df[~driver_df['cntrb_id'].isin(driver_df.loc[mask]['cntrb_id'])]
#mask = (driver_df['yearmonth'] < end_date)
#driver_df = driver_df.loc[mask]
#driver_df = driver_df.loc[driver_df['rank'] == rank]
#adds all months to driver_df so the lists of dates will include all months and years
driver_df = pd.concat([driver_df, months_df])
data = pd.DataFrame()
if group_by == 'year':
#x-axis dates
data['dates'] = driver_df['created_year'].unique()
data['dates'] = data['dates'].dropna()
display(data['dates'])
#average issues each month
data['average_comments_per_issue'] = driver_df.groupby(['created_year']).mean().reset_index()['comments'] / driver_df.groupby(['created_year', 'issue_id']).count().reset_index()['comments']
#display(driver_df.groupby(['created_year']).mean().reset_index()['comments'])
display(driver_df.groupby(['issue_id']).count().reset_index()['comments'])
#used to format x-axis and title
group_by_format_string = "Year"
elif group_by == 'month':
#x-axis dates
dates = np.datetime_as_string(driver_df['yearmonth'], unit='M')
dates = np.unique(dates)
data['dates'] = dates
display(data['dates'])
#new contributor counts for y-axis
data['average_comments_per_issue'] = driver_df.groupby(['yearmonth']).sum().reset_index()[y_axis]
#used to format x-axis and title
group_by_format_string = "Month"
if len(data['average_comments_per_issue']) >= 15:
plot_width = 46 * len(data['average_comments_per_issue'])
else:
plot_width = 670
p = figure(x_range=data['dates'], plot_height=400, plot_width = plot_width, title='Title',
toolbar_location=None, y_range=(0, max(data['average_comments_per_issue'])* 1.15), margin = (0, 0, 200, 0))
p.vbar(x=data['dates'], top=data['average_comments_per_issue'], width=0.8)
source = ColumnDataSource(data=dict(dates=data['dates'], average_comments_per_issue=data['average_comments_per_issue']))
labels = LabelSet(x='dates', y='average_comments_per_issue', text='average_comments_per_issue', y_offset=4,
text_font_size="13pt", text_color="black",
source=source, text_align='center')
p.add_layout(labels)
caption = 'caption'
caption = Label(x=-10, y=-120, x_units='screen', y_units='screen',
text='{}'.format(caption), render_mode='css',
background_fill_color='white', text_font_size = '15pt')
p.add_layout(caption)
p.xgrid.grid_line_color = None
p.y_range.start = 0
p.axis.minor_tick_line_color = None
p.outline_line_color = None
p.title.align = "center"
p.title.text_font_size = "18px"
p.yaxis.axis_label = 'Average Comments per Issues per Year'
p.xaxis.axis_label = group_by_format_string
p.xaxis.axis_label_text_font_size = "18px"
p.yaxis.axis_label_text_font_size = "16px"
p.xaxis.major_label_text_font_size = "16px"
p.xaxis.major_label_orientation = 45.0
p.yaxis.major_label_text_font_size = "16px"
show(p)
# -
vertical_bar_chart(issues_closed, months_df, group_by = 'year')
|
templates/.ipynb_checkpoints/issues_template-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Deep Equals
#
# Write a function that determines if two lists of numbers are equal
a = [1, 2, 3]
b = [1, 2, 3]
c = [1, 2, 3, 4]
d = [3, 2, 1]
e = [1, 1, 2, 3]
# +
def listAreEqual(list1, list2):
if len(list1) != len(list2):
return False
for i, item in enumerate(list1):
if list1[i] != list2[i]:
return False
return True
listAreEqual(a, b)
# -
|
JavaScript/Python-Drills/06-Deep_Equals/Unsolved/Deep_Equals.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="y00b5TQZnqs_"
# # Your First AI application
#
# Going forward, AI algorithms will be incorporated into more and more everyday applications. For example, you might want to include an image classifier in a smart phone app. To do this, you'd use a deep learning model trained on hundreds of thousands of images as part of the overall application architecture. A large part of software development in the future will be using these types of models as common parts of applications.
#
# In this project, you'll train an image classifier to recognize different species of flowers. You can imagine using something like this in a phone app that tells you the name of the flower your camera is looking at. In practice you'd train this classifier, then export it for use in your application. We'll be using [this dataset](http://www.robots.ox.ac.uk/~vgg/data/flowers/102/index.html) from Oxford of 102 flower categories, you can see a few examples below.
#
# <img src='assets/Flowers.png' width=500px>
#
# The project is broken down into multiple steps:
#
# * Load the image dataset and create a pipeline.
# * Build and Train an image classifier on this dataset.
# * Use your trained model to perform inference on flower images.
#
# We'll lead you through each part which you'll implement in Python.
#
# When you've completed this project, you'll have an application that can be trained on any set of labeled images. Here your network will be learning about flowers and end up as a command line application. But, what you do with your new skills depends on your imagination and effort in building a dataset. For example, imagine an app where you take a picture of a car, it tells you what the make and model is, then looks up information about it. Go build your own dataset and make something new.
# + [markdown] colab_type="text" id="kKnPjnLAftRV"
# ## Import Resources
# -
import warnings
warnings.filterwarnings('ignore')
# + colab={} colab_type="code" id="2dCk6873paNW"
# TODO: Make all necessary imports.
# %matplotlib inline
import json
import matplotlib.pyplot as plt
import numpy as np
import tensorflow as tf
import tensorflow_datasets as tfds
import tensorflow_hub as hub
tfds.disable_progress_bar()
# -
import logging
logger = tf.get_logger()
logger.setLevel(logging.ERROR)
print('GPU available' if tf.test.is_gpu_available() else 'GPU no available')
# + [markdown] colab_type="text" id="tWKF0YOarpCx"
# ## Load the Dataset
#
# Here you'll use `tensorflow_datasets` to load the [Oxford Flowers 102 dataset](https://www.tensorflow.org/datasets/catalog/oxford_flowers102). This dataset has 3 splits: `'train'`, `'test'`, and `'validation'`. You'll also need to make sure the training data is normalized and resized to 224x224 pixels as required by the pre-trained networks.
#
# The validation and testing sets are used to measure the model's performance on data it hasn't seen yet, but you'll still need to normalize and resize the images to the appropriate size.
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="vXISRjfdrrQ6" outputId="6edf59b2-b468-4c4a-cff4-7cc7cfcc3c2d"
# TODO: Load the dataset with TensorFlow Datasets.
train_split = 60
validation_split = 20
test_split = 20
split = tfds.Split.ALL.subsplit([train_split, validation_split, test_split])
dataset, dataset_info = tfds.load('oxford_flowers102', split=split, as_supervised=True, with_info=True)
# TODO: Create a training set, a validation set and a test set.
training_set, validation_set, testing_set = dataset
# + [markdown] colab_type="text" id="S5pdQnDbf0-j"
# ## Explore the Dataset
# + colab={"base_uri": "https://localhost:8080/", "height": 102} colab_type="code" id="XikJ4X7FUv8v" outputId="10240009-1148-41ae-8ce0-4025c2f2fa87"
# TODO: Get the number of examples in each set from the dataset info.
total_examples = dataset_info.splits['train'].num_examples + dataset_info.splits['test'].num_examples + dataset_info.splits['validation'].num_examples
n_training = (total_examples*train_split)//100
n_testing = (total_examples*test_split)//100
n_validation = (total_examples*validation_split)//100
# TODO: Get the number of classes in the dataset from the dataset info.
n_classes = dataset_info.features['label'].num_classes
print(f'Samples for train: {n_training}')
print(f'Samples for test: {n_testing}')
print(f'Samples for validation: {n_validation}')
print(f'\nThere are {n_classes} species of flowers')
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" id="CWR9ScCbPI_D" outputId="fdf01c8d-2db9-4d7c-a566-4db2599fd1ab"
# TODO: Print the shape and corresponding label of 3 images in the training set.
for _, (image, label) in zip(range(3), training_set):
print(f'Shape: {image.shape} - Label: {label}')
# + colab={"base_uri": "https://localhost:8080/", "height": 280} colab_type="code" id="DQbnq8htRTnl" outputId="32a0e1af-2b04-440e-ddb4-835732be3e83"
# TODO: Plot 1 image from the training set. Set the title
# of the plot to the corresponding image label.
first_image = next(iter(training_set))[0]
plt.imshow(first_image.numpy())
plt.show()
# + [markdown] colab_type="text" id="zuh1841cs-j1"
# ### Label Mapping
#
# You'll also need to load in a mapping from label to category name. You can find this in the file `label_map.json`. It's a JSON object which you can read in with the [`json` module](https://docs.python.org/3.7/library/json.html). This will give you a dictionary mapping the integer coded labels to the actual names of the flowers.
# + colab={} colab_type="code" id="JoVzdO3KsdSk"
with open('label_map.json', 'r') as f:
class_names = json.load(f)
# + colab={"base_uri": "https://localhost:8080/", "height": 280} colab_type="code" id="fc6pMUZgEvUo" outputId="4274fd43-5cee-4523-885f-a18f6f277dd6"
# TODO: Plot 1 image from the training set. Set the title
# of the plot to the corresponding class name.
first_image, first_label = next(iter(training_set))
plt.imshow(first_image.numpy())
plt.title(class_names[str(first_label.numpy())])
plt.axis('off')
plt.show()
# + [markdown] colab_type="text" id="0gL7AaqNf-NC"
# ## Create Pipeline
# + colab={"base_uri": "https://localhost:8080/", "height": 88} colab_type="code" id="5hNznLbPNZxS" outputId="7c114910-b75f-4220-cda9-f84426ec2728"
# TODO: Create a pipeline for each set.
IMG_SIZE = 224
BATCH_SIZE = 64
def normalize(image, label):
image = tf.cast(image, tf.float32)
image = tf.image.resize(image, (IMG_SIZE, IMG_SIZE))
image /= 255.0
return image, label
training_batches = training_set.cache().shuffle(n_training//4).map(normalize).batch(BATCH_SIZE).prefetch(1)
validation_batches = validation_set.cache().map(normalize).batch(BATCH_SIZE).prefetch(1)
testing_batches = testing_set.map(normalize).batch(BATCH_SIZE).prefetch(1)
# + [markdown] colab_type="text" id="gR9gtRbeXPYx"
# # Build and Train the Classifier
#
# Now that the data is ready, it's time to build and train the classifier. You should use the MobileNet pre-trained model from TensorFlow Hub to get the image features. Build and train a new feed-forward classifier using those features.
#
# We're going to leave this part up to you. If you want to talk through it with someone, chat with your fellow students!
#
# Refer to the rubric for guidance on successfully completing this section. Things you'll need to do:
#
# * Load the MobileNet pre-trained network from TensorFlow Hub.
# * Define a new, untrained feed-forward network as a classifier.
# * Train the classifier.
# * Plot the loss and accuracy values achieved during training for the training and validation set.
# * Save your trained model as a Keras model.
#
# We've left a cell open for you below, but use as many as you need. Our advice is to break the problem up into smaller parts you can run separately. Check that each part is doing what you expect, then move on to the next. You'll likely find that as you work through each part, you'll need to go back and modify your previous code. This is totally normal!
#
# When training make sure you're updating only the weights of the feed-forward network. You should be able to get the validation accuracy above 70% if you build everything right.
#
# **Note for Workspace users:** One important tip if you're using the workspace to run your code: To avoid having your workspace disconnect during the long-running tasks in this notebook, please read in the earlier page in this lesson called Intro to GPU Workspaces about Keeping Your Session Active. You'll want to include code from the workspace_utils.py module. Also, If your model is over 1 GB when saved as a checkpoint, there might be issues with saving backups in your workspace. If your saved checkpoint is larger than 1 GB (you can open a terminal and check with `ls -lh`), you should reduce the size of your hidden layers and train again.
# + colab={} colab_type="code" id="4zElEHViXLni"
# TODO: Build and train your network.
URL = 'https://tfhub.dev/google/imagenet/mobilenet_v2_140_224/feature_vector/4'
model = tf.keras.Sequential([
hub.KerasLayer(URL, input_shape=(IMG_SIZE, IMG_SIZE, 3), trainable=False),
tf.keras.layers.Dense(n_classes, activation='softmax')
])
model.summary()
# -
model.compile(optimizer='adam', loss='sparse_categorical_crossentropy', metrics=['accuracy'])
history = model.fit(training_batches, epochs=8, validation_data=validation_batches)
# + colab={"base_uri": "https://localhost:8080/", "height": 498} colab_type="code" id="VU6sWzx4e7Yb" outputId="f7b5c7c5-683a-463c-9228-68c4918bdd5b"
# TODO: Plot the loss and accuracy values achieved during training for the training and validation set.
_, (ax1, ax2) = plt.subplots(1, 2, figsize=(10, 5))
ax1.set_title('Accuracy')
ax1.plot(np.arange(1, 9), history.history['accuracy'], label='Training accuracy')
ax1.plot(np.arange(1, 9), history.history['val_accuracy'], label='Validation accuracy')
ax1.legend()
ax2.set_title('Loss')
ax2.plot(np.arange(1, 9), history.history['loss'], label='Training loss')
ax2.plot(np.arange(1, 9), history.history['val_loss'], label='Validation loss')
ax2.legend()
# + [markdown] colab_type="text" id="qcTDnyvop3ky"
# ## Testing your Network
#
# It's good practice to test your trained network on test data, images the network has never seen either in training or validation. This will give you a good estimate for the model's performance on completely new images. You should be able to reach around 70% accuracy on the test set if the model has been trained well.
# + colab={"base_uri": "https://localhost:8080/", "height": 85} colab_type="code" id="79l7-HM1cafO" outputId="6cf468a4-1e27-4f20-d63a-a8bdd78bcdbe"
# TODO: Print the loss and accuracy values achieved on the entire test set.
loss, accuracy = model.evaluate(testing_batches)
print(f'Loss for testing set: {loss}')
print(f'Accuracy for testing set: {accuracy}')
# + [markdown] colab_type="text" id="pLsIDWnuqfkl"
# ## Save the Model
#
# Now that your network is trained, save the model so you can load it later for making inference. In the cell below save your model as a Keras model (*i.e.* save it as an HDF5 file).
# + colab={} colab_type="code" id="7XOwdOjSptp-"
# TODO: Save your trained model as a Keras model.
path_to_model = './saved_model.h5'
model.save(path_to_model, save_format='h5')
# + [markdown] colab_type="text" id="rbeLSRC1rxuj"
# ## Load the Keras Model
#
# Load the Keras model you saved above.
# + colab={"base_uri": "https://localhost:8080/", "height": 394} colab_type="code" id="3T6Dgc7Nrzds" outputId="f5d356dc-183f-4cd3-f15d-88ebb4966082"
# TODO: Load the Keras model
reloaded_model = tf.keras.models.load_model(path_to_model, custom_objects={'KerasLayer': hub.KerasLayer})
reloaded_model.summary()
# + [markdown] colab_type="text" id="ZjucwuFrsyhJ"
# # Inference for Classification
#
# Now you'll write a function that uses your trained network for inference. Write a function called `predict` that takes an image, a model, and then returns the top $K$ most likely class labels along with the probabilities. The function call should look like:
#
# ```python
# probs, classes = predict(image_path, model, top_k)
# ```
#
# If `top_k=5` the output of the `predict` function should be something like this:
#
# ```python
# probs, classes = predict(image_path, model, 5)
# print(probs)
# print(classes)
# > [ 0.01558163 0.01541934 0.01452626 0.01443549 0.01407339]
# > ['70', '3', '45', '62', '55']
# ```
#
# Your `predict` function should use `PIL` to load the image from the given `image_path`. You can use the [Image.open](https://pillow.readthedocs.io/en/latest/reference/Image.html#PIL.Image.open) function to load the images. The `Image.open()` function returns an `Image` object. You can convert this `Image` object to a NumPy array by using the `np.asarray()` function.
#
# The `predict` function will also need to handle pre-processing the input image such that it can be used by your model. We recommend you write a separate function called `process_image` that performs the pre-processing. You can then call the `process_image` function from the `predict` function.
#
# ### Image Pre-processing
#
# The `process_image` function should take in an image (in the form of a NumPy array) and return an image in the form of a NumPy array with shape `(224, 224, 3)`.
#
# First, you should convert your image into a TensorFlow Tensor and then resize it to the appropriate size using `tf.image.resize`.
#
# Second, the pixel values of the input images are typically encoded as integers in the range 0-255, but the model expects the pixel values to be floats in the range 0-1. Therefore, you'll also need to normalize the pixel values.
#
# Finally, convert your image back to a NumPy array using the `.numpy()` method.
# + colab={} colab_type="code" id="oG7mJ1-5s1qe"
# TODO: Create the process_image function
def process_image(np_image):
image = tf.convert_to_tensor(np_image)
image = tf.image.resize(image, (224, 224))
image = tf.cast(image, tf.float32)
image /= 255.0
return image.numpy()
# -
# To check your `process_image` function we have provided 4 images in the `./test_images/` folder:
#
# * cautleya_spicata.jpg
# * hard-leaved_pocket_orchid.jpg
# * orange_dahlia.jpg
# * wild_pansy.jpg
#
# The code below loads one of the above images using `PIL` and plots the original image alongside the image produced by your `process_image` function. If your `process_image` function works, the plotted image should be the correct size.
# +
from PIL import Image
image_path = './test_images/hard-leaved_pocket_orchid.jpg'
im = Image.open(image_path)
test_image = np.asarray(im)
processed_test_image = process_image(test_image)
fig, (ax1, ax2) = plt.subplots(figsize=(10,10), ncols=2)
ax1.imshow(test_image)
ax1.set_title('Original Image')
ax2.imshow(processed_test_image)
ax2.set_title('Processed Image')
plt.tight_layout()
plt.show()
# -
# Once you can get images in the correct format, it's time to write the `predict` function for making inference with your model.
#
# ### Inference
#
# Remember, the `predict` function should take an image, a model, and then returns the top $K$ most likely class labels along with the probabilities. The function call should look like:
#
# ```python
# probs, classes = predict(image_path, model, top_k)
# ```
#
# If `top_k=5` the output of the `predict` function should be something like this:
#
# ```python
# probs, classes = predict(image_path, model, 5)
# print(probs)
# print(classes)
# > [ 0.01558163 0.01541934 0.01452626 0.01443549 0.01407339]
# > ['70', '3', '45', '62', '55']
# ```
#
# Your `predict` function should use `PIL` to load the image from the given `image_path`. You can use the [Image.open](https://pillow.readthedocs.io/en/latest/reference/Image.html#PIL.Image.open) function to load the images. The `Image.open()` function returns an `Image` object. You can convert this `Image` object to a NumPy array by using the `np.asarray()` function.
#
# **Note:** The image returned by the `process_image` function is a NumPy array with shape `(224, 224, 3)` but the model expects the input images to be of shape `(1, 224, 224, 3)`. This extra dimension represents the batch size. We suggest you use the `np.expand_dims()` function to add the extra dimension.
# + colab={} colab_type="code" id="SBnPKFJuGB32"
# TODO: Create the predict function
def predict(image_path, model, top_k):
image = Image.open(image_path)
np_image = np.asarray(image)
processed_image = process_image(np_image)
input_image = np.expand_dims(processed_image, axis=0)
probs = model.predict(input_image)[0]
indexes = np.argsort(probs)[-top_k:][::-1]
probs = probs[indexes]
classes = list(map(lambda n: str(n + 1), indexes))
return probs, classes
# + [markdown] colab_type="text" id="aft8f_n5C7Co"
# # Sanity Check
#
# It's always good to check the predictions made by your model to make sure they are correct. To check your predictions we have provided 4 images in the `./test_images/` folder:
#
# * cautleya_spicata.jpg
# * hard-leaved_pocket_orchid.jpg
# * orange_dahlia.jpg
# * wild_pansy.jpg
#
# In the cell below use `matplotlib` to plot the input image alongside the probabilities for the top 5 classes predicted by your model. Plot the probabilities as a bar graph. The plot should look like this:
#
# <img src='assets/inference_example.png' width=600px>
#
# You can convert from the class integer labels to actual flower names using `class_names`.
# + colab={"base_uri": "https://localhost:8080/", "height": 336} colab_type="code" id="I_tBH8xGGVxQ" outputId="ef0fe795-65f3-49c5-fab0-086fac7d409d"
# TODO: Plot the input image along with the top 5 classes
image_path = './test_images/cautleya_spicata.jpg'
image = Image.open(image_path)
top_k = 5
probs, classes = predict(image_path, reloaded_model, top_k)
plt.subplot(1, 2, 1)
plt.imshow(np.asarray(image))
plt.axis('off')
plt.subplot(1, 2, 2)
plt.title('Class Probability')
plt.barh(np.arange(top_k), probs)
plt.yticks(np.arange(top_k), [class_names[key] for key in classes])
plt.tight_layout()
plt.show()
|
image-classifier/Project_Image_Classifier_Project.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .sh
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Bash
# language: bash
# name: bash
# ---
# Exam by <NAME>
# ## Problem 2
#
# The command 'ls' lists the files in a directory. The option -l displays the list with details of the files, and the option -h makes the size of the files human-readable.
ls -lh
# The owner and the group have permissions to read and write the file 'Locus_Germplasm_Phenotype_20130122.txt', while "anyone" has permissions to only read the file.
# ## Problem 3
#
# The command 'head' prints the first lines of a file. Adding a number as option limits the number of lines displayed.
head -1 Locus_Germplasm_Phenotype_20130122.txt
# ## Problem 4
#
# The command 'wc' (that stands for 'word count') has the function of counting the number of elements in a file. By adding the option -l the command counts the number of lines in the specified file.
wc -l Locus_Germplasm_Phenotype_20130122.txt
# ## Problem 5
#
# Using the previous problems as hints, I found the following solution: knowing the total number of lines in the file, we could get the last lines (using the command 'tail') and specify to retrieve all of them minus one (the first). After that, we use the operator '>' to redirect the output of the command to a file called "Data_Only.csv".
tail -7215 Locus_Germplasm_Phenotype_20130122.txt > Data_Only.csv
# To find out if the result is correct, we just count the number of lines in that file.
wc -l Data_Only.csv
# In order to check even further, we can see if the header has been removed correctly
echo "Locus file: "
head -3 Locus_Germplasm_Phenotype_20130122.txt
echo ""
echo "Data only file: "
head -2 Data_Only.csv
# ## Problem 6
#
# The command 'grep' can find strings and regular expressions inside a file or a set of files. Useful command options for this problem are -i, that makes the search insensitive to case, and -n, that displays the number of line of the resulting matches.
#
# In addition, I will be using the options -E to be able to use extended regular expressions and -a to treat the file as text in case it is binary, as I don't know its source.
#
# The problem specifies that the word 'root' is contained in the phenotype part of the line. The locus_name and pubmed_id cannot contain the pattern 'root' as they can only contain numbers and certain letters. If no germplasm name contains the substring 'root', then we can search for the substring in any part of the line and the results will be from the phenotype part.
#
# We can find if any germplasm name contains 'root' with the next command:
grep -E -ina '^\w+\s\S*root\S*\s.*' Locus_Germplasm_Phenotype_20130122.txt
# There is no output, so there is no germplasm name containing 'root'. Then, the lines that contain the word root in the phenotype are found by the next command:
grep -ina 'root' Locus_Germplasm_Phenotype_20130122.txt
# If you want to see only the line number, the command would be:
grep -ina 'root' Locus_Germplasm_Phenotype_20130122.txt | grep -E -o '^[0-9]+'
# ## Problem 7
#
# To retrieve the locus name of the lines that contain 'root' we follow a similar process to the one I used to get the line numbers: using a pipeline to obtain the first word characters of the lines that contain 'root'.
#
# Just like in Problem 5, to redirect the result we use the operator '>'.
grep -ia 'root' Locus_Germplasm_Phenotype_20130122.txt | grep -E -o '^\w+' > Root-associated-Loci.txt
# ## Problem 8
#
# The command is exactly the same as the one used in Problem 7, just changing the condition of the second 'grep' that the pattern must match at the beggining of the line for the condition of matching at the end of the line, and changing the match from 'any word character' to 'digits'.
#
# Of course, the file name also changes.
grep -ia 'root' Locus_Germplasm_Phenotype_20130122.txt | grep -E -o '[0-9]+$' > Root-associated-Publications.txt
# ## Problem 9
#
# Using the background information that one of the characters in the AGI Locus Code correspond to the number of chromosome (specifically the third character) we can find out if there is a chromosome with no genes related to roots.
#
# We have already a file with all the AGI Locus Codes associated to roots. Using 5 slightly different commands on it, we can find if any of the 5 chromosomes does not have genes related to roots.
#
# There is no need to specify in the pattern that we are looking for that the first two letters can be capital or lower-case, as we are using the option -i. The next character in the pattern is the number of the chromosome, different in each command. After searching for matches, we count the number of them to find out if any of the chromosomes has 0 genes related to roots.
grep -i 'at1' Root-associated-Loci.txt | wc -l
grep -i 'at2' Root-associated-Loci.txt | wc -l
grep -i 'at3' Root-associated-Loci.txt | wc -l
grep -i 'at4' Root-associated-Loci.txt | wc -l
grep -i 'at5' Root-associated-Loci.txt | wc -l
# In this case, it seems that all chromosomes have genes associated to roots!
# ## Problem 10
#
# I think I can skip this question.
|
Exam_1_Answers.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Convolutional Layer
# Visualize four filtered outputs (a.k.a. feature maps) of a convolutional layer.
# ### Import the image
# Import the Necessary Packages
import torch
import torch.nn as nn
import torch.nn.functional as F
import cv2
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# +
# image path
img_path = 'images/AA.JPG'
# load color image (cv2: bgr)
bgr_img = cv2.imread(img_path)
# convert to gray
gray_img = cv2.cvtColor(bgr_img, cv2.COLOR_BGR2GRAY)
# normalize scale to [0,1]
gray_img = gray_img.astype('float32')/255
# plot image
plt.imshow(gray_img, cmap='gray')
plt.show()
# -
# ## Create 4-Filters (vertical and horizontal)
# +
# used 4x4 filter
filter_vals = np.ones((4,4))
filter_vals[:,0:2] *= -1
# print(filter_vals)
print('Filter shape: ', filter_vals.shape)
# make four different filters
filter_1 = filter_vals
filter_2 = -filter_1
filter_3 = filter_1.T
filter_4 = -filter_3
filters = np.array([filter_1, filter_2, filter_3, filter_4])
# visualize four filters
fig = plt.figure(figsize=(10,5))
for i in range(4):
ax = fig.add_subplot(1, 4, i+1, xticks=[], yticks=[])
ax.imshow(filters[i], cmap='gray')
ax.set_title('Filter {}'.format(str(i+1)))
width, height = filters[i].shape
for x in range(width):
for y in range(height):
ax.annotate(str(filters[i][x][y]),
xy=(y,x),
horizontalalignment='center',
verticalalignment='center',
color='white' if filters[i][x][y] < 0 else 'black')
# -
# ## Define a Network Architecture
# For CNN
# * Convolutional layers
#
# +
# CNN with a single convolutional layer with four filters
class Net(nn.Module):
def __init__(self, weight):
super(Net, self).__init__()
# initializes the weights of the convolutional layer to be the weights of the 4 defined filters
k_height, k_width = weight.shape[2:]
# 1 input image channel(grayscale), 4 output channels/feature maps (filter)
# 4x4 square convolution kernel
self.conv = nn.Conv2d(1, 4, kernel_size=(k_height, k_width), bias=False)
self.conv.weight = torch.nn.Parameter(weight)
# define the feedforward behavior
def forward(self, x):
conv_x = self.conv(x)
activated_x = F.relu(conv_x)
# return both layers
return conv_x, activated_x
# +
# instantiate the model ans set the weights
weight = torch.from_numpy(filters).unsqueeze(1).type(torch.FloatTensor)
print(weight.shape)
model = Net(weight)
print(model)
# -
# ## Visualize the output of each filter
# helper function for visualizing the output of a given layer
# default number of filters is 4
def viz_layer(layer, n_filters= 4):
fig = plt.figure(figsize=(20, 20))
for i in range(n_filters):
ax = fig.add_subplot(1, n_filters, i+1, xticks=[], yticks=[])
# grab layer outputs
ax.imshow(np.squeeze(layer[0,i].data.numpy()), cmap='gray')
ax.set_title('Output %s' % str(i+1))
# +
# plot original image
plt.imshow(gray_img, cmap='gray')
# visualize all filters
fig = plt.figure(figsize=(12, 6))
fig.subplots_adjust(left=0, right=1.5, bottom=0.8, top=1, hspace=0.05, wspace=0.05)
for i in range(4):
ax = fig.add_subplot(1, 4, i+1, xticks=[], yticks=[])
ax.imshow(filters[i], cmap='gray')
ax.set_title('Filter %s' % str(i+1))
# convert the image into an input Tensor
gray_img_tensor = torch.from_numpy(gray_img).unsqueeze(0).unsqueeze(1)
# get the convolutional layer (pre and post activation)
conv_layer, activated_layer = model(gray_img_tensor)
# visualize the output of a conv layer
viz_layer(conv_layer)
# -
# visualize the output of an activated conv layer
viz_layer(activated_layer)
|
01_Convolution_Layer/Convolutional Layer.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Using NumPy arrays enables you to express many kinds of data processing tasks as concise array expressions that might otherwise require writing loops. This practice of replacing explicit loops with array expressions is commonly referred to as vectorization.
# vectorize is a class to convert an ordinary Python function which accepts scalars and returns scalars into a “vectorized-function” with the same broadcasting rules as other Numpy functions
import numpy as np
def addsubtract(a,b):
"Return a-b if a > b, otherwise return a + b"
if a > b:
return a - b
else:
return a + b
addsubtract(5,7)
vec_addsubtract = np.vectorize(addsubtract)
vec_addsubtract([0,3,6,9],[1,3,5,7])
# As always, the shapes need to match
# We get an error if they don't
vec_addsubtract([0,3,6,9],[1,3,5,7,8])
|
Vectorization_m03_demo03.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import math
import collections
import numpy as np
import pandas as pd
import matplotlib.pyplot as pp
# %matplotlib inline
# -
squares = []
for i in range(1, 11):
squares.append(i ** 2)
squares
square = [i**2 for i in range(1,11)]
#square brackets around it since it is a list
#put the computation first then the for loop
square
pow_to_pow = [i**i for i in range(1,11)]
pow_to_pow
sum = [1,2,3,4,5,6,7,8,9,10]
add = 0
for i in range(0,10):
add = add+ sum[i]
print(add)
|
Comprehensions.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: learning_nlp
# language: python
# name: learning_nlp
# ---
import pandas as pd
combined = pd.read_csv('data/Combined_News_DJIA.csv')
news = pd.read_csv('data/RedditNews.csv')
market = pd.read_csv('data/upload_DJIA_table.csv')
print(combined.shape)
combined.head(1)
# Remove 'b'-prefixes
# Applymap -> Like .apply() but Applied to Entire DF
combined = combined.applymap(lambda cell: cell.strip() if type(cell)==str else cell)
combined = combined.applymap(lambda cell: cell.lstrip('b"') if type(cell)==str else cell)
combined.head(2)
# Sample Corpus from Combined DF
# -- Includes Only Top1 Headlines
corpus = combined[['Date', 'Label', 'Top1']]
print(corpus.shape)
corpus.head(3)
# +
# We will create a document-term matrix using a bag-of-words approach.
# -
# #### document-term matrix =
# - each **col** represents a word and
# - each **row** represents a documnent
# - the value in every cell can represent things:
# - Most traditionally, it is a count of how many times a word appears in a doc.
# - It can also be a boolean, does this word appear at all.
# - TF-IDF: term-frequency inverse document-frequency
#
# #### Bag of Words = Counting the Appearance of Words to try to predict something.
# Bag of words can be good for classification problems.
# - Why is bag of words good for classification?
# +
import spacy
# load NN sm, md, lg etc.
nlp = spacy.load("en_core_web_lg")
# +
tokens = []
# Tokenize Corpus
for doc in nlp.pipe(corpus['Top1'], batch_size=500):
doc_tokens = []
for token in doc:
if (token.is_stop == False) & (token.is_punct == False) & (token.is_space == False):
doc_tokens.append(token.text.lower())
tokens.append(doc_tokens)
# Create New Tokens Column in Corpus DF
corpus.insert(3, 'tokens', tokens)
# -
corpus.head(2)
# ## But... we **won't be using Tokenization** with the Vectorizors
# ### Count Vectorizor
# #### Vectorizor does NOT mean Word Embeddings
# -> Vectorizor are things that build Document Term Matricies
# - One of 3 BOW methods
# +
# Building a Document Term Matrix
from sklearn.feature_extraction.text import CountVectorizer
# --- Key Arguments/Parameters for Count Vectorizor ---
# max_features -> sets limit on vocabulary size aka Limit Features
# - Only most common words are made into features.
# max_df/min_df -> Limit Features by Document Frequency
# -> Ignores terms that show up in greater than 90% of docs
# --> Or less than 2% of docs for example
# ngram_range -> range of n-values for different n-gram
# or char n-grams to be extracted
# You can also override scikit-learn's default tokenization.
# Scikit-learns default tokenization is not as good as spaCy's.
# Instantiate the Transformer
vect = CountVectorizer(stop_words=nlp.Defaults.stop_words,
max_features=1000)
# Build Vocab
# We simply pass an iterable of docs...
# It 'tokenizes on its own...'
# AKA it Builds the Vocab on its own
# Tokenization -> One way to Build Vocab I think...
# ?- Can this be improved using lemmitization? -?
vect.fit(corpus['Top1'])
# transform text - Count Vocab
# Build the Matrix Using the Vocab Determined during the fit Command
dtm = vect.transform(corpus['Top1'])
# -
# Print Out Words in Vocabulary
print(vect.get_feature_names()[0:50])
# The DTM is Values and their Location in Coordinates
print('Type:', type(dtm))
print('Shape:', dtm.shape)
print('First 2 Values:')
print(dtm[2]) # These are coordinates of the matrix :)
# dtm sparce to df
# Get Word Counts for Each Document
dtm = pd.DataFrame(dtm.todense(), columns=vect.get_feature_names())
print(dtm.shape)
dtm.head()
dtm.head()
# Examine Distribution of Doc Lengths
# AKA Distrubution of Headline Lengths
doc_lengths = [len(doc) for doc in corpus['Top1']]
pd.Series(doc_lengths).hist()
# ### -> Think about it...
# #### Word counts are affected by document size.
# #### Enter -> TF-IDF
# # TF-IDF -> Term Frequency Inverse Documnet Frequency
# ### Helps Control for Different Document Lengths in the Same Corpus
# Term Frequency = Percentage Words in Document for each word.
#
# Document Frequency = A penalty for the word existing in a high number of documents.
#
# The purpose of TF-IDF is to find what is unique to each document. It penalizes the term frequencies of words that are common across all documents which will allow for each document's most different topics to rise to the top.
# ### Document Term Matrix with TF-IDF
# +
from sklearn.feature_extraction.text import TfidfVectorizer
# ?-- What percentage of overall corpus size
# sould max_features be? --?
# Instantiate Vectorizer Object
tfidf = TfidfVectorizer(stop_words='english', max_features=2500)
# Create a vocabulary and get word counts per document.
# Similar to fit_predict
dtm = tfidf.fit_transform(corpus['Top1'])
# Print Word Counts
# Get feature names to use as dataframe column headers.
dtm = pd.DataFrame(dtm.todense(), columns=tfidf.get_feature_names())
# View Feature Matrix as DataFrame
print(dtm.shape)
dtm.head()
# -
# ### We want to trim down our DTM by reducing noise while improving signal.
# #### We will use spaCy tokenization, stop-words, n-grams, and statistical trimming to help us refine the results of our dtm.
def tokenize(document):
# Return Lemmas
doc = nlp(document)
return [token.lemma_.strip() for token in doc if (token.is_stop != True) and (token.is_punct != True)]
# n-grams = multi-word phrases
# bi-gram = 2 words
# tri-gram = 3 words
# sklearn will search through all the valid combinations of words
# and return all those possible phrases as they appear in text....
# This results in WAYYY more words in a vocabulary as you have all the words
# singular ANDDD the words as combinations SO...
# When specifying ngram_range -> ALWAYS limit the features!
print(len(corpus['Top1']))
# +
# NOTE
# Tunning Parameters
# Instantiate Vectorizor Object
tfidf = TfidfVectorizer(stop_words='english',
ngram_range=(1, 2),
# Term or N-Gram Must Appear in no more than 97 docs.
max_df=97,
# Term or N-Gram Must Appear in at least 3 Docs
min_df=4,
max_features=2000,
# spaCy Tokenization from Func above
tokenizer=tokenize)
# Create a Vocabulary and Get Word Counts per Document
dtm = tfidf.fit_transform(corpus['Top1']) # Similar to Fit-Predict
# Print Word Counts
# Get Feature Names to Use ad DF Column Headers
dtm = pd.DataFrame(dtm.todense(), columns=tfidf.get_feature_names())
# View Feature Matrix as DataFrame
print(dtm.shape)
dtm.head()
# -
# ### Cosine Similarity
# +
# Calculate Distance of TF-IDF Vectors
# cosine similarity == the idea of measuring the Euclidian (straight-line)
# distance between two points and normalizing this
# cosine similarity == normalization of euclidean distance
from sklearn.metrics.pairwise import cosine_similarity
# We are finding the distance between each document's TF-IDF vector
dist_matrix = cosine_similarity(dtm)
# -
# Let's find a headline similar to the first row of Top1.
corpus['Top1'][0]
# +
# Turn the Dist Matrix into a Dataframe
dist = pd.DataFrame(dist_matrix)
# Similarity Accross DF
print(dist.shape)
dist.head()
# -
# Each row in the similarity of one document to all other documents.
dist[0][0:15]
# +
# Reccomendation Engine
# Find all rows that are not 1 (aka, identical to headline)
# Orderthem by most similar.
dist[dist[11] < 1][11].sort_values(ascending=False).index[0]
# -
# We can see the similarity.
print(corpus['Top1'][11])
print(' ')
print(corpus['Top1'][848])
def find_top_similarity(index_value):
print(corpus['Top1'][index_value])
# Find all rows that are not 1 (aka, identical to headline)
# Orderthem by most similar -> Get Top One
similar = dist[dist[index_value] < 1]\
[index_value].sort_values(ascending=False).index[0]
print('\n')
print(corpus['Top1'][similar])
find_top_similarity(20)
# #### Cosine Similarity is too computationally expensive to work in the majority of situations
#
# Besides cosine similarity euclidian distance there are 2 main ways to store similarity:
#
# - KD Trees
# - Ball Trees
#
# These are alternatives to storing the distance between every combination of vectors.
# Document Term Matrix with TF-IDF as Values
print(dtm.shape)
dtm.head(3)
# +
from sklearn.neighbors import NearestNeighbors
# Fit on DTM
# Specify we want 5 neighbors per doc
nn = NearestNeighbors(n_neighbors=5, algorithm='kd_tree')
nn.fit(dtm)
# -
dtm.iloc[0]
# We pass our model a doc vector to get the model to point us to it's
# K- Nearest Neighbors
nn.kneighbors([dtm.iloc[0]])
# Query Using Kneighbors
nn.kneighbors([dtm.iloc[14]])
corpus['Top1'][14]
# Looks Like Euclidean distance was more effective...
print(corpus['Top1'][14])
print(' ')
print(corpus['Top1'][606])
print(' ')
print(corpus['Top1'][385])
# +
# Let's try again with a random number:
from numpy import random
randint = random.randint(len(corpus['Top1']))
print("Random Integer", randint, "\n")
kneighbors_query = nn.kneighbors([dtm.iloc[randint]])
print(kneighbors_query, "\n")
print(corpus['Top1'][randint], "\n")
print(corpus['Top1'][kneighbors_query[1][0][1]], "\n")
print(corpus['Top1'][kneighbors_query[1][0][2]])
# From seeing a few examples I get the sense that the system is biased
# towards shorter headlines... I wonder why??
# ?-- TODO --? The Following Show Up Disproportionately:
# 'What a surprise...' [385]
# 'Scenes from the recession' [152]
# 'June 4th, 1989' [205]
# -
# ### Making a match with an outside text source.
new_doc = ["There is a New President."]
# Query for Sim of Random Doc to Our Reddit Headlines
# Create a DTM Row (With TDIF Values) for the new doc.
# -> Relative the Reddit Headline Corpus
new = tfidf.transform(new_doc)
new # LOOK -> Only 2 stored elements! Only 2 words from the sample new doc are
# included in the Term Set.. a good indicator that this headline might not
# work as well with the present model.
# We could decide on a threshold for `number of stored_elements` under-which we refuse to provide a reccomendation in order to reduce the amount of impractical, inaccurate results.
# Turn the Sparse Matrix into a Dense Matrix
new = new.todense()
# Now we have a new doc/row that is expressed in a dense TF-IDF Term Vector
# and we are going to plug that row into our KNN model to get an output of our
# K- nearest neighbors
nn.kneighbors(new)
corpus['Top1'][192]
# The limitation exists in that... docs that have few tokens that are included in the vocabulary of the Doc-Term Matrix will be extra sparse (aka. many zeros).
# --> Other vectors that also contain a high proportion of zeros will be identified by the KNN model as 'similar', even though the only similarity the 2 docs have in common is thier lack of tokens that are a part of the term set. --> Thats why the KNN model might be biased to favor shorter headlines when comparing a headline that does not include many high TF-IDF values.
# How would JC set a distance threshold for recommendations from my KNN model?
#
# 1. Select a sample size of query articles.
# 2. Then he would get the knn recommendations for those 10 articles ~5 recommendations each.
# 3. For each of the 50 recs, he would label them as useful or not useful.
# 4. Take one of the following values as the threshold:
# - Max Distance of the Useful Labelled recs
# - 75% percentile distance of the use labelled recs
# - Min of the not useful labelled recs
#
# Then after he has his model in production, he would run a similar experiment using A/B testing.
# ### Word Embeddings
# +
nlp = spacy.load('en_core_web_lg')
doc = nlp("Two bananas in pyjamas")
# Get the vector for the token "bananas"
# ?-- How does this spaCy vector differ from the vectors of the DTM? --?
# For each term-document pair, the TF-IDF DTM ascribes only 1 TF-IDF Value
# ?-- In this case we would see a matrix for each document-term pair..
# or a matrix for each term/token? --?
# -- SOLVED -> It is querying from a complex data-structure that represents
# the english language. The vector of a doc represents the location of the doc
# among this "english map"...
bananas_vector = doc.vector
# These values are un-interpretable.
# For ALL Docs of ANY Length, a vector of length 300 will be constructed to
# represent said vector.
# ?-- New Question -> How are the multiple tokens that form a doc aggregated
# to always have a vector of length 300? Are they averaged? --?
print(len(bananas_vector))
print(bananas_vector[:50])
# +
from sklearn.decomposition import PCA
def get_word_vectors(words):
# Converts a list of words into their word vectors.
# Returns a List of Word Vectors
return [nlp(word).vector for word in words]
words = ['car', 'truck', 'suv', 'race', 'elves', 'dragon', 'sword', 'king',
'queen', 'prince', 'horse', 'fish', 'lion', 'tiger', 'lynx', 'potato',
'baking', 'textbook', 'student', 'homework', 'studying', 'fear']
# Initalize pca model and tell it to project data down onto 2 dimensions.
pca = PCA(n_components=2)
# fit the pca model to our 300-D data, this will work out which is the best way
# to project the data down that will best maintain the relative distances
# between data points.
# It will store these instructions on how to transform the data.
pca.fit(get_word_vectors(words))
# Tell our (fitted) pca model to transform our 300D data down onto 2D using the
# instructions it learned during the fit phase.
word_vecs_2d = pca.transform(get_word_vectors(words))
# let's look at our new 2D word vectors
word_vecs_2d
# -
print("Length of Embeddings List: ", len(get_word_vectors(words)))
print("Size of Original Embedding Vector: ", len(get_word_vectors(words)[0]))
print("Size of Reduced Embedding Vector: ", len(word_vecs_2d[0]))
# +
import matplotlib.pyplot as plt
plt.figure(figsize=(20, 15))
# plot the catter plot of where the words will be
plt.scatter(word_vecs_2d[:, 0], word_vecs_2d[:,1])
# for each word and coordinate pair: draw the text on the plot
for word, coord in zip(words, word_vecs_2d):
x, y = coord
plt.text(x, y, word, size=15)
# show the plot
plt.show()
# +
# Most popular word embeddings today is 'Berts' or 'Elmo'?
# -
|
vector_repr.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <img style="float: left; padding-right: 10px; width: 45px" src="https://raw.githubusercontent.com/Harvard-IACS/2018-CS109A/master/content/styles/iacs.png"> CS109A Introduction to Data Science
#
# ## Lecture 3, Exercise 1: Web Scraping and Parsing Intro
#
#
# **Harvard University**<br/>
# **Fall 2020**<br/>
# **Instructors**: <NAME>, <NAME>, and <NAME>
#
# ---
# # Title
#
# **Exercise 1: Web Scraping and Parsing Intro**
#
# # Description
#
# **OVERVIEW**
#
# As we learned in class, the three most common sources of data used for Data Science are:
#
# - files (e.g, `.csv`, `.txt`) that already contain the dataset
# - APIs (e.g., Twitter or Facebook)
# - web scraping (e.g., Requests)
#
# Here, we get practice with web scraping by using **Requests**. Once we fetch the page contents, we will need to extract the information that we actually care about. We rely on <a href="https://www.crummy.com/software/BeautifulSoup/bs4/doc/" target="_blank">BeautifulSoup</a> to help with this.
import re
import requests
from bs4 import BeautifulSoup
# ## **NOTE**: After running every cell, be sure to auto-grade your work by clicking 'Mark' in the lower-right corner. Otherwise, no credit will be given.
#
# For this exercise, we will be grabbing data (the Top News stories) from [AP News](apnews.com), a not-for-profit news agency.
# the URL of the webpage that has the desired info
url = "https://apnews.com/hub/ap-top-news"
# # Web Scraping (Graded)
# Let's use [`requests`](https://requests.readthedocs.io/en/master/user/quickstart/) to fetch the contents. Specifically, the [`requests`](https://requests.readthedocs.io/en/master/user/quickstart/) library has a `.get()` function that returns a [Response object](https://www.w3schools.com/python/ref_requests_response.asp). A Response object contains the server's _response_ to the HTTP request, and thus contains all the information that we could want from the page.
#
# Below, fill in the blank to fetch AP News' Top News website.
### edTest(test_a) ###
home_page = requests.get(____)
home_page.status_code
# You should have received a status code of 200, which means the page was successfully found on the server and sent to receiver (aka client/user/you). [Again, you can click here](https://www.restapitutorial.com/httpstatuscodes.html) for a full list of status codes. Recall that sometimes, while browsing the Internet, webpages will report a 404 error, possibly with an entertaining graphic to ease your pain. That 404 is the status code, just like we are using here!
# `home_page` is now a [Response object](https://www.w3schools.com/python/ref_requests_response.asp). It contains many attributes, including the `.text`. Run the cell below and note that it's identical to if we were to visit the webpage in our browser and clicked 'View Page Source'.
home_page.text
# # Data Parsing Intro (Graded)
# The above `.text` property is atrocious to view and make sense of. Sure, we could write Regular Expressions to extract all of the contents that we're interested in. Instead, let's first use [`BeautifulSoup`](https://www.crummy.com/software/BeautifulSoup/bs4/doc/) to parse the content into more manageable chunks.
#
# Below, fill in the blank to construct an HTML-parsed [`BeautifulSoup`](https://www.crummy.com/software/BeautifulSoup/bs4/doc/) object from our website.
### edTest(test_b) ###
soup = BeautifulSoup(____, ____)
soup
# You'll notice that the `soup` object is better formatted than just looking at the entire text. It's still dense, but it helps.
#
# Below, fill in the blank to set `webpage_title` equal to the text of the webpage's title (no HTML tags included).
### edTest(test_c) ###
webpage_title = ____
# Again, our BeautifulSoup object allows for quick, convenient searching and access to the web page contents.
#
# # Data Parsing Examples (Not Graded)
#
# Anytime you wish to extract specific contents from a webpage, it is necessary to:
# - **Step 1**. While viewing the page in your browser, identify what contents of the page you're interested in.
# - **Step 2**. Look at the HTML returned from the BeautifulSoup object, and pinpoint the specific context that surrounds each of these items that you're interested in
# - **Step 3.** Devise a pattern using BeautifulSoup and/or RegularExpressions to extract said contents.
# For example:
# ### **Step 1:**
# Let's say, for every news article found on the AP's Top News page, you want to extract the link and associated title. In this screenshot
# <img src="https://github.com/Harvard-IACS/2020-CS109A/blob/master/content/lectures/lecture03/images/apnews_sample.png?raw=true">
#
# we can see one news article (there are many more below on the page). Its title is `"California fires bring more chopper rescues, power shutoffs"` and its link is to [/c0aa17fff978e9c4768ee32679b8555c](/c0aa17fff978e9c4768ee32679b8555c). Since the current page is stored at apnews.com, the article link's full address is [apnews.com/c0aa17fff978e9c4768ee32679b8555c](apnews.com/c0aa17fff978e9c4768ee32679b8555c).
#
#
# ### **Step 2:**
#
# After printing the `soup` object, we saw a huge mess of all of the HTML still. So, let's drill down into certain sections. As illustrated in the [official documentation here](https://www.crummy.com/software/BeautifulSoup/bs4/doc/#navigating-using-tag-names), we can retrieve all `<a>` links by running the cell below:
soup.find_all("a")
# This is still a ton of text (links). So, let's get more specific. I now search for the title text `California fires bring more chopper rescues, power shutoffs` within the output of the previous cell (the HTML of all links). I notice the following:
#
# `<a class="Component-headline-0-2-110" data-key="card-headline" href="/c0aa17fff978e9c4768ee32679b8555c"><h1 class="Component-h1-0-2-111">California fires bring more chopper rescues, power shutoffs</h1></a>`
#
# I also see that this is repeatable; every news article on the Top News page has such text! Great!
# ### **Step 3:**
#
# The pattern is that we want the value of the `href` attribute, along with the text of the link. There are many ways to get at this information. Below, I show just a few:
# +
# EXAMPLE 1
# returns all `a` links that also contain `Component-headline-0-2-110`
soup.find_all("a", "Component-headline-0-2-110")
# iterates over each link and extracts the href and title
for link in soup.find_all("a", "Component-headline-0-2-110"):
url = "www.apnews.com" + link['href']
title = link.text
# -
# As mentioned in the official documentation [here](https://www.crummy.com/software/BeautifulSoup/bs4/doc/#attributes) and [here](https://www.crummy.com/software/BeautifulSoup/bs4/doc/#the-keyword-arguments), a tag (such as `a`) may have many attributes, and you can search them by putting your terms in a dictionary.
# EXAMPLE 2
# this returns the same exact subset of links as the example above
# so, we could iterate through the list just like above
soup.find_all("a", attrs={"data-key": "card-headline"})
# Alternatively, we could use Regular Expressions if we were confident that our Regex pattern only matched on the relevant links.
# EXAMPLE 3
# instead of using the BeautifulSoup, we are handling all of the parsing
# ourselves, and working directly with the original Requests text
re.findall("<a class=\"Component-headline.*?href=\"(.+?)\"><h1.+?>(.+?)</h1></a>", home_page.text)
|
docs/lectures/lecture03/notebook/cs109a_L3_Ex1_solutions.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="Ff_Jv8Ptu4lP"
#
# # INSTALLATION
# + colab={"base_uri": "https://localhost:8080/"} id="q5qYRG8zufHw" executionInfo={"status": "ok", "timestamp": 1629766552146, "user_tz": -570, "elapsed": 14179, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhE0cXJBikzHY7xUavBkbPRseKZ_N-obrw0xIoLhQ=s64", "userId": "04127040763952829247"}} outputId="d132d22a-9464-4619-e261-d12f6e8231fb"
# !pip install aif360
# !pip install fairlearn
# + colab={"base_uri": "https://localhost:8080/"} id="TltW3iPkux0Q" executionInfo={"status": "ok", "timestamp": 1629766552967, "user_tz": -570, "elapsed": 842, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhE0cXJBikzHY7xUavBkbPRseKZ_N-obrw0xIoLhQ=s64", "userId": "04127040763952829247"}} outputId="68d0e48e-95ba-40b2-d75a-4f2ba37a8c10"
# !apt-get install -jre
# !java -version
# + colab={"base_uri": "https://localhost:8080/"} id="KssrNl8GvDYU" executionInfo={"status": "ok", "timestamp": 1629766585039, "user_tz": -570, "elapsed": 31759, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhE0cXJBikzHY7xUavBkbPRseKZ_N-obrw0xIoLhQ=s64", "userId": "04127040763952829247"}} outputId="728914ba-50ea-4a9f-82f8-4cbbb99e8eeb"
# !pip install h2o
# + colab={"base_uri": "https://localhost:8080/"} id="_NQn2JJ0uw6u" executionInfo={"status": "ok", "timestamp": 1629766588460, "user_tz": -570, "elapsed": 3440, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhE0cXJBikzHY7xUavBkbPRseKZ_N-obrw0xIoLhQ=s64", "userId": "04127040763952829247"}} outputId="ab30bf55-41ae-43af-e966-8a7674f2541c"
# !pip install xlsxwriter
# + [markdown] id="-Y_uQ6vdvN4a"
# #IMPORTS
# + colab={"base_uri": "https://localhost:8080/"} id="rf1aISz6vGfR" executionInfo={"status": "ok", "timestamp": 1629766598251, "user_tz": -570, "elapsed": 9810, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhE0cXJBikzHY7xUavBkbPRseKZ_N-obrw0xIoLhQ=s64", "userId": "04127040763952829247"}} outputId="c7c573c3-b810-4ca4-bcb9-3e77d8702323"
import numpy as np
from mlxtend.feature_selection import ExhaustiveFeatureSelector
from xgboost import XGBClassifier
# import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import openpyxl
import xlsxwriter
from openpyxl import load_workbook
import shap
#suppress setwith copy warning
pd.set_option('mode.chained_assignment',None)
from sklearn.feature_selection import VarianceThreshold
from sklearn.feature_selection import SelectKBest, SelectFwe, SelectPercentile,SelectFdr, SelectFpr, SelectFromModel
from sklearn.feature_selection import chi2, mutual_info_classif
# from skfeature.function.similarity_based import fisher_score
import aif360
import matplotlib.pyplot as plt
from aif360.metrics.classification_metric import ClassificationMetric
from aif360.algorithms.postprocessing import EqOddsPostprocessing
from aif360.metrics import BinaryLabelDatasetMetric
from aif360.datasets import StandardDataset , BinaryLabelDataset
from sklearn.preprocessing import MinMaxScaler
MM= MinMaxScaler()
import h2o
from h2o.automl import H2OAutoML
from h2o.estimators.glm import H2OGeneralizedLinearEstimator
import sys
sys.path.append("../")
import os
# + colab={"base_uri": "https://localhost:8080/", "height": 554} id="RcxQeeX7vUXz" executionInfo={"status": "ok", "timestamp": 1629766605765, "user_tz": -570, "elapsed": 7571, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhE0cXJBikzHY7xUavBkbPRseKZ_N-obrw0xIoLhQ=s64", "userId": "04127040763952829247"}} outputId="edb9f32a-f7d7-4e83-a0f8-8ca0b848f6e2"
h2o.init()
# + [markdown] id="RQVI-ISXvrZm"
# #**************************LOADING DATASET*******************************
# + colab={"base_uri": "https://localhost:8080/"} id="FEGPULDrvk3g" executionInfo={"status": "ok", "timestamp": 1629766875056, "user_tz": -570, "elapsed": 37471, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhE0cXJBikzHY7xUavBkbPRseKZ_N-obrw0xIoLhQ=s64", "userId": "04127040763952829247"}} outputId="105bc1c7-51c4-45e7-945d-31f16c7d4cd0"
from google.colab import drive
drive.mount('/content/gdrive', force_remount=True)
# + id="uN9VfZBAvxCj" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1629774371885, "user_tz": -570, "elapsed": 7277694, "user": {"displayName": "<NAME>", "photoUrl": "<KEY>", "userId": "04127040763952829247"}} outputId="1f7fe382-e0bf-42c8-8150-173c637b3645"
for i in range(1,51,1):
train_url=r'/content/gdrive/MyDrive/Datasets/SurveyData/DATASET/Law/Train'
train_path= os.path.join(train_url ,("Train"+ str(i)+ ".csv"))
train= pd.read_csv(train_path).drop(['region_first'], axis=1)
test_url=r'/content/gdrive/MyDrive/Datasets/SurveyData/DATASET/Law/Test'
test_path= os.path.join(test_url ,("Test"+ str(i)+ ".csv"))
test= pd.read_csv(test_path).drop(['region_first'], axis=1)
# normalization of train and test sets
Fitter= MM.fit(train)
transformed_train=Fitter.transform(train)
train=pd.DataFrame(transformed_train, columns= train.columns)
#test normalization
transformed_test=Fitter.transform(test)
test=pd.DataFrame(transformed_test, columns= test.columns)
# ********************SETTING TO H20 FRAME AND MODEL TRAINING*******************************
x = list(train.columns)
y = "first_pf"
x.remove(y)
Train=h2o.H2OFrame(train)
Test= h2o.H2OFrame(test)
Train[y] = Train[y].asfactor()
Test[y] = Test[y].asfactor()
aml = H2OAutoML(max_models=10, nfolds=10, include_algos=['GBM'] , stopping_metric='AUTO') #verbosity='info',,'GBM', 'DRF'
aml.train(x=x, y=y, training_frame=Train)
best_model= aml.leader
# a.model_performance()
#**********************REPLACE LABELS OF DUPLICATED TRAIN AND TEST SET WITH 0.5 THRESHOLDED PREDICT PROBA****************************
#predicted proba for train labels
gbm_Predictions_train= best_model.predict(Train)
gbm_Predictions_train= gbm_Predictions_train.as_data_frame()
train_label= (gbm_Predictions_train.p1>0.5).astype(int)
predicted_df_train= train.copy()
predicted_df_train['first_pf']= train_label
#predicted proba for test labels
gbm_Predictions_test= best_model.predict(Test)
gbm_Predictions_test= gbm_Predictions_test.as_data_frame()
test_label= (gbm_Predictions_test.p1>0.5).astype(int)
predicted_df_test= test.copy()
predicted_df_test['first_pf']= test_label
# *************CHECKING FAIRNESS IN DATASET**************************
## ****************CONVERTING TO BLD FORMAT******************************
'''the EO's fit method takes as input the original data (ground truth) and the predicted dataset with the predicted dataset having
same features but different labels (label for predicted dataset is the thresholded predict proba). as TPR and FPR requires \hat{Y} and Y
Converting the two required dataset to binary format as accepted by EO.
'''
#Transforming the Ground truth Train Test to BLD
advantagedGroup= [{'race':1}]
disadvantagedGroup= [{'race':0}]
class Train(StandardDataset):
def __init__(self,label_name= 'first_pf',
favorable_classes= [1],protected_attribute_names=['race'], privileged_classes=[[1]], ):
super(Train, self).__init__(df=train , label_name=label_name ,
favorable_classes=favorable_classes , protected_attribute_names=protected_attribute_names ,
privileged_classes=privileged_classes ,
)
BLD_Train= Train(protected_attribute_names= ['race'],
privileged_classes= [[1]])
class Test(StandardDataset):
def __init__(self,label_name= 'first_pf',
favorable_classes= [1],protected_attribute_names=['race'], privileged_classes=[[1]], ):
super(Test, self).__init__(df=test , label_name=label_name ,
favorable_classes=favorable_classes , protected_attribute_names=protected_attribute_names ,
privileged_classes=privileged_classes ,
)
BLD_Test= Test(protected_attribute_names= ['race'],
privileged_classes= [[1]])
#**************************************Predicted Train Test BLD*****************************************
class PreTrain(StandardDataset):
def __init__(self,label_name= 'first_pf',
favorable_classes= [1],protected_attribute_names=['race'], privileged_classes=[[1]], ):
super(PreTrain, self).__init__(df=predicted_df_train , label_name=label_name ,
favorable_classes=favorable_classes , protected_attribute_names=protected_attribute_names ,
privileged_classes=privileged_classes ,
)
BLD_PredictedTrain= PreTrain(protected_attribute_names= ['race'],
privileged_classes= [[1]])
class PreTest(StandardDataset):
def __init__(self,label_name= 'first_pf',
favorable_classes= [1],protected_attribute_names=['race'], privileged_classes=[[1]], ):
super(PreTest, self).__init__(df=predicted_df_test , label_name=label_name ,
favorable_classes=favorable_classes , protected_attribute_names=protected_attribute_names ,
privileged_classes=privileged_classes ,
)
BLD_PredictedTest= PreTest(protected_attribute_names= ['race'],
privileged_classes= [[1]])
#****************************************Equal Opportunity Instance*******************************
Equalizer= EqOddsPostprocessing(unprivileged_groups= disadvantagedGroup , privileged_groups= advantagedGroup )
#uses ground truth and unfair predictions of classifier to determine probabilities with which unfair output labels are changed to satisfy EO
Equalizer.fit(BLD_Train,BLD_PredictedTrain )
#predicting the new labels assigned by the EO engine
BLD_PredictedTest= Equalizer.predict(BLD_PredictedTest)
# ********************COMPUTE DISCRIMINATION*****************************
excelBook= load_workbook('/content/gdrive/MyDrive/Datasets/SurveyData/RESULTS/EqualOdds/EO_gbm.xlsx')
Law= excelBook['Law']
data= Law.values
# Get columns
columns = next(data)[0:]
10# Create a DataFrame based on the second and subsequent lines of data
OldDF = pd.DataFrame(data, columns=columns)
ClassifierBias = ClassificationMetric( BLD_Test,BLD_PredictedTest , unprivileged_groups= disadvantagedGroup, privileged_groups= advantagedGroup)
Accuracy= ClassifierBias.accuracy()
TPR= ClassifierBias.true_positive_rate()
TNR= ClassifierBias.true_negative_rate()
NPV= ClassifierBias.negative_predictive_value()
PPV= ClassifierBias.positive_predictive_value()
SP=ClassifierBias .statistical_parity_difference()
IF=ClassifierBias.consistency()
DI=ClassifierBias.disparate_impact()
EOP=ClassifierBias.true_positive_rate_difference()
EO=ClassifierBias.average_odds_difference()
FDR= ClassifierBias.false_discovery_rate(privileged=False)- ClassifierBias.false_discovery_rate(privileged=True)
NPV_diff=ClassifierBias.negative_predictive_value(privileged=False)-ClassifierBias.negative_predictive_value(privileged=True)
FOR=ClassifierBias.false_omission_rate(privileged=False)-ClassifierBias.false_omission_rate(privileged=True)
PPV_diff=ClassifierBias.positive_predictive_value(privileged=False) -ClassifierBias.positive_predictive_value(privileged=True)
BGE = ClassifierBias.between_group_generalized_entropy_index()
WGE = ClassifierBias.generalized_entropy_index()-ClassifierBias.between_group_generalized_entropy_index()
BGTI = ClassifierBias.between_group_theil_index()
WGTI = ClassifierBias.theil_index() -ClassifierBias.between_group_theil_index()
EDF= ClassifierBias.differential_fairness_bias_amplification()
newdf= pd.DataFrame(index = [0], data= { 'ACCURACY': Accuracy,'TPR': TPR, 'PPV':PPV, 'TNR':TNR,'NPV':NPV,'SP':SP,'CONSISTENCY':IF,'DI':DI,'EOP':EOP,'EO':EO,'FDR':FDR,'NPV_diff':NPV_diff,
'FOR':FOR,'PPV_diff':PPV_diff,'BGEI':BGE,'WGEI':WGE,'BGTI':BGTI,'WGTI':WGTI,'EDF':EDF})
newdf=pd.concat([OldDF,newdf])
pathway= r"/content/gdrive/MyDrive/Datasets/SurveyData/RESULTS/EqualOdds/EO_gbm.xlsx"
with pd.ExcelWriter(pathway, engine='openpyxl') as writer:
#load workbook base as for writer
writer.book= excelBook
writer.sheets=dict((ws.title, ws) for ws in excelBook.worksheets)
newdf.to_excel(writer, sheet_name='Law', index=False)
# newdf.to_excel(writer, sheet_name='Adult', index=False)
print('Accuracy', Accuracy)
# + [markdown] id="Hoc62jY7Olbt"
# #LOGISTIC REGRESSION
# + id="RZmY0q8iVY3O" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1629774490151, "user_tz": -570, "elapsed": 118300, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhE0cXJBikzHY7xUavBkbPRseKZ_N-obrw0xIoLhQ=s64", "userId": "04127040763952829247"}} outputId="5bfc4a01-df54-4d56-93c3-113ebba60fbf"
for i in range(1,51,1):
train_url=r'/content/gdrive/MyDrive/Datasets/SurveyData/DATASET/Law/Train'
train_path= os.path.join(train_url ,("Train"+ str(i)+ ".csv"))
train= pd.read_csv(train_path).drop(['region_first'], axis=1)
test_url=r'/content/gdrive/MyDrive/Datasets/SurveyData/DATASET/Law/Test'
test_path= os.path.join(test_url ,("Test"+ str(i)+ ".csv"))
test= pd.read_csv(test_path).drop(['region_first'], axis=1)
# normalization of train and test sets
Fitter= MM.fit(train)
transformed_train=Fitter.transform(train)
train=pd.DataFrame(transformed_train, columns= train.columns)
#test normalization
transformed_test=Fitter.transform(test)
test=pd.DataFrame(transformed_test, columns= test.columns)
# ********************SETTING TO H20 FRAME AND MODEL TRAINING*******************************
x = list(train.columns)
y = "first_pf"
x.remove(y)
Train=h2o.H2OFrame(train)
Test= h2o.H2OFrame(test)
Train[y] = Train[y].asfactor()
Test[y] = Test[y].asfactor()
LogReg = H2OGeneralizedLinearEstimator(family= "binomial", lambda_ = 0)
LogReg.train(x=x, y=y, training_frame=Train)
#**********************REPLACE LABELS OF DUPLICATED TRAIN AND TEST SET WITH 0.5 THRESHOLDED PREDICT PROBA****************************
#predicted proba for train labels
lr_Predictions_train= LogReg.predict(Train)
lr_Predictions_train= lr_Predictions_train.as_data_frame()
train_label= (lr_Predictions_train.p1>0.5).astype(int)
predicted_df_train= train.copy()
predicted_df_train['first_pf']= train_label
#predicted proba for test labels
lr_Predictions_test= LogReg.predict(Test)
lr_Predictions_test= lr_Predictions_test.as_data_frame()
test_label= (lr_Predictions_test.p1>0.5).astype(int)
predicted_df_test= test.copy()
predicted_df_test['first_pf']= test_label
# *************CHECKING FAIRNESS IN DATASET**************************
## ****************CONVERTING TO BLD FORMAT******************************
'''the EO's fit method takes as input the original data (ground truth) and the predicted dataset with the predicted dataset having
same features but different labels (label for predicted dataset is the thresholded predict proba). as TPR and FPR requires \hat{Y} and Y
Converting the two required dataset to binary format as accepted by EO.
'''
#Transforming the Ground truth Train Test to BLD
advantagedGroup= [{'race':1}]
disadvantagedGroup= [{'race':0}]
class Train(StandardDataset):
def __init__(self,label_name= 'first_pf',
favorable_classes= [1],protected_attribute_names=['race'], privileged_classes=[[1]], ):
super(Train, self).__init__(df=train , label_name=label_name ,
favorable_classes=favorable_classes , protected_attribute_names=protected_attribute_names ,
privileged_classes=privileged_classes ,
)
BLD_Train= Train(protected_attribute_names= ['race'],
privileged_classes= [[1]])
class Test(StandardDataset):
def __init__(self,label_name= 'first_pf',
favorable_classes= [1],protected_attribute_names=['race'], privileged_classes=[[1]], ):
super(Test, self).__init__(df=test , label_name=label_name ,
favorable_classes=favorable_classes , protected_attribute_names=protected_attribute_names ,
privileged_classes=privileged_classes ,
)
BLD_Test= Test(protected_attribute_names= ['race'],
privileged_classes= [[1]])
#**************************************Predicted Train Test BLD*****************************************
class PreTrain(StandardDataset):
def __init__(self,label_name= 'first_pf',
favorable_classes= [1],protected_attribute_names=['race'], privileged_classes=[[1]], ):
super(PreTrain, self).__init__(df=predicted_df_train , label_name=label_name ,
favorable_classes=favorable_classes , protected_attribute_names=protected_attribute_names ,
privileged_classes=privileged_classes ,
)
BLD_PredictedTrain= PreTrain(protected_attribute_names= ['race'],
privileged_classes= [[1]])
class PreTest(StandardDataset):
def __init__(self,label_name= 'first_pf',
favorable_classes= [1],protected_attribute_names=['race'], privileged_classes=[[1]], ):
super(PreTest, self).__init__(df=predicted_df_test , label_name=label_name ,
favorable_classes=favorable_classes , protected_attribute_names=protected_attribute_names ,
privileged_classes=privileged_classes ,
)
BLD_PredictedTest= PreTest(protected_attribute_names= ['race'],
privileged_classes= [[1]])
#****************************************Equal Opportunity Instance*******************************
Equalizer= EqOddsPostprocessing(unprivileged_groups= disadvantagedGroup , privileged_groups= advantagedGroup )
#uses ground truth and unfair predictions of classifier to determine probabilities with which unfair output labels are changed to satisfy EO
Equalizer.fit(BLD_Train,BLD_PredictedTrain )
#predicting the new labels based on the fitted EO engine
BLD_PredictedTest= Equalizer.predict(BLD_PredictedTest)
# ***************************COMPUTE DISCRIMINATION********************************
excelBook= load_workbook("/content/gdrive/MyDrive/Datasets/SurveyData/RESULTS/EqualOdds/EO_LogReg.xlsx")
Law= excelBook['Law']
data= Law.values
# Get columns
columns = next(data)[0:]
10# Create a DataFrame based on the second and subsequent lines of data
OldDF = pd.DataFrame(data, columns=columns)
ClassifierBias = ClassificationMetric( BLD_Test,BLD_PredictedTest , unprivileged_groups= disadvantagedGroup, privileged_groups= advantagedGroup)
Accuracy= ClassifierBias.accuracy()
TPR= ClassifierBias.true_positive_rate()
TNR= ClassifierBias.true_negative_rate()
NPV= ClassifierBias.negative_predictive_value()
PPV= ClassifierBias.positive_predictive_value()
SP=ClassifierBias .statistical_parity_difference()
IF=ClassifierBias.consistency()
DI=ClassifierBias.disparate_impact()
EOP=ClassifierBias.true_positive_rate_difference()
EO=ClassifierBias.average_odds_difference()
FDR= ClassifierBias.false_discovery_rate(privileged=False)- ClassifierBias.false_discovery_rate(privileged=True)
NPV_diff=ClassifierBias.negative_predictive_value(privileged=False)-ClassifierBias.negative_predictive_value(privileged=True)
FOR=ClassifierBias.false_omission_rate(privileged=False)-ClassifierBias.false_omission_rate(privileged=True)
PPV_diff=ClassifierBias.positive_predictive_value(privileged=False) -ClassifierBias.positive_predictive_value(privileged=True)
BGE = ClassifierBias.between_group_generalized_entropy_index()
WGE = ClassifierBias.generalized_entropy_index()-ClassifierBias.between_group_generalized_entropy_index()
BGTI = ClassifierBias.between_group_theil_index()
WGTI = ClassifierBias.theil_index() -ClassifierBias.between_group_theil_index()
EDF= ClassifierBias.differential_fairness_bias_amplification()
newdf= pd.DataFrame(index = [0], data= { 'ACCURACY': Accuracy,'TPR': TPR, 'PPV':PPV, 'TNR':TNR,'NPV':NPV,'SP':SP,'CONSISTENCY':IF,'DI':DI,'EOP':EOP,'EO':EO,'FDR':FDR,'NPV_diff':NPV_diff,
'FOR':FOR,'PPV_diff':PPV_diff,'BGEI':BGE,'WGEI':WGE,'BGTI':BGTI,'WGTI':WGTI,'EDF':EDF})
newdf=pd.concat([OldDF,newdf])
pathway= r"/content/gdrive/MyDrive/Datasets/SurveyData/RESULTS/EqualOdds/EO_LogReg.xlsx"
with pd.ExcelWriter(pathway, engine='openpyxl') as writer:
#load workbook base as for writer
writer.book= excelBook
writer.sheets=dict((ws.title, ws) for ws in excelBook.worksheets)
newdf.to_excel(writer, sheet_name='Law', index=False)
# newdf.to_excel(writer, sheet_name='Adult', index=False)
print('Accuracy', Accuracy)
# + id="YyfPmIRsIMuE" executionInfo={"status": "aborted", "timestamp": 1629774490967, "user_tz": -570, "elapsed": 831, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhE0cXJBikzHY7xUavBkbPRseKZ_N-obrw0xIoLhQ=s64", "userId": "04127040763952829247"}}
|
Fairness_Survey/ALGORITHMS/EO/LawSchool.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 划分方式
# +
# 12666 imgs:
# train: 9000
# val: 1666
# test: 2000
# -
import pandas as pd
server_df = pd.read_csv('server-train.txt', header=None)
server_df.describe()
server_df.head()
shuffled_df = server_df.sample(frac=1).reset_index(drop=True)
server_df.head()
shuffled_df.head()
shuffled_df.to_csv('shuffled-all.txt', header=None, index=None)
train_df = shuffled_df.iloc[0:9000]
val_df = shuffled_df.iloc[9000:10666]
test_df = shuffled_df.iloc[10666:12666]
train_df.to_csv('train_1.txt', header=None, index=None)
val_df.to_csv('val_1.txt', header=None, index=None)
test_df.to_csv('test_1.txt', header=None, index=None)
train_df.describe()
# # See
import pandas as pd
train_df = pd.read_csv('train_1.txt', header=None, delim_whitespace=True)
train_df.to_csv('train_1_9000.csv', header=None)
|
_baseline/pre/shuffle.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 課題4 テキストデータ分析
# 配点
# - Q1, 1P
# - Q2, 2P
# - Q3, 5P
# - Q4, 2P
#
# 必要なモジュールのimport
import json
import csv
import numpy as np
import pandas as pd
# ## Q1
#
# $n$-次元ベクトル空間における、任意の2つのベクトル、$ \vec{x}=(x_1,x_2,...,x_n)$、$\vec{y}=(y_1,y_2,...,y_n)$、の間のcos類似度 $cos( \vec{x}, \vec{y})$ は以下のように定義されます。
#
# $cos( \vec{x}, \vec{y}) = \frac{\vec{x}\cdot \vec{y}}{\|x\|_2\|y\|_2}=\frac{\Sigma^{n}_{i=1}{x_{i}y_{i}}}{\sqrt{\Sigma^{n}_{i=1}x_{i}^{2}}\sqrt{\Sigma^{n}_{i=1}y_{i}^{2}}}$
#
# 入力ベクトル $\vec{x}$, $\vec{y}$ をそれぞれ`NumPy` の配列として引数で受け取り、それらのベクトル間のcos類似度を計算して返す関数 `cos_sim` を完成させてください。
def cos_sim(vec1, vec2):
return # (vec1とvec2の内積)/np.sqrt(vec1の要素の二乗和*vec2の要素の二乗和)
# `cos_sim()`関数が完成したら、以下のセルを実行して動作を確認してください。`-1.0, 1.0, 0.0`がそれぞれのcos類似度です。
print(cos_sim(np.array([1,0]), np.array([-1,0])))
print(cos_sim(np.array([1,0]), np.array([1,0])))
print(cos_sim(np.array([1,0]), np.array([0,-1])))
assert cos_sim(np.array([1,0]), np.array([-1,0])) == -1.0
assert cos_sim(np.array([1,0]), np.array([1,0])) == 1.0
assert cos_sim(np.array([1,0]), np.array([0,-1])) == 0.0
# ## Q2
# "course_list.csv"ファイルには以下のように各行に授業名のテキストデータ(文字コードはUTF8)が含まれています。以下では、このファイルを読み込み、各授業のベクトルを作成し、授業間の類似度を求めるコードを実装します。
# ```Python
# ## course_list.csvファイル
# ...
# 計量経済学Ⅰ
# 数理統計
# 計量経済学
# 経済史Ⅱ
# ICTマネジメントⅠ
# 現代日本経済史
# 経済学史
# ...
# ```
# Colaboratoryでは以下を実行して必要なファイルをダウンロード
# !wget https://raw.githubusercontent.com/UTDataMining/2020A/master/ex4/course_list.csv
# ### Q2.1
# 引数`file`に与えられたファイル名のファイルを1行ずつ**順番に**読み込み、その各行を要素とするリストを作成して返す`create_list`関数を完成さてください。作成されたリストは変数`courses`で受け取ります。以降の処理では、リスト`courses`のインデックスをその要素(授業名)のIDとして扱います。
#
def create_list(file):
lst=[]
with open(file, 'r', encoding="utf-8") as f:
dataReader = csv.reader(f)
for row in dataReader:
# lstにrow[0]を追加
return lst
# `create_list`関数が完成したら、以下のセルを実行して動作を確認してください。リストの長さ(授業の数)は`4678`、"Pythonプログラミング入門"授業のID(リスト`courses`のインデックス)は`6`となります。
courses = create_list('course_list.csv')
print(len(courses))
print(courses.index('Pythonプログラミング入門'))
assert len(courses) == 4678
assert courses.index('Pythonプログラミング入門') == 6
# "keyword_list.csv"ファイルには以下のように各行に1単語が含まれています(文字コードはUTF8)。
# ```Python
# ## keyword_list.csvファイル
# ...
# 資源
# 広域
# フランス語
# 語学
# 教育法
# 環境学
# 相関
# 解析学
# 森林
# ...
# ```
# 上記と同様に`create_list`関数により、"keywor_list.csv"ファイルを1行ずつ**順番に**読み込み、その各行を要素とするリストを作成します。作成されたリストは変数`vocab`で受け取ります。以降の処理では、リスト`vocab`のインデックスをその要素(単語)のIDとして扱います。リスト`vocab`は以降の処理における語彙となります。
# Colaboratoryでは以下を実行して必要なファイルをダウンロード
# !wget https://raw.githubusercontent.com/UTDataMining/2020A/master/ex4/keyword_list.csv
# 以下のセルを実行して動作を確認してください。リストの長さ(単語の数)は`910`、単語"プログラミング"のID(リスト`vocab`のインデックス)は`113`、単語"英語"のID(リスト`vocab`のインデックス)は`15`となります。となります。
vocab = create_list('keyword_list.csv')
print(len(vocab))
print(vocab.index('プログラミング'))
print(vocab.index('英語'))
assert len(vocab) == 910
assert vocab.index('プログラミング') == 113
assert vocab.index('英語') == 15
# ### Q2.2
# リスト`courses`と`vocab`を引数で受け取り、単語のID(リスト`vocab`のその単語のインデックス)をキー、その単語のDF(Document Frequency)を値とする辞書を作成して返す`count_df`関数を作成してください。作成された辞書は変数`df`で受け取ります。この場合、ある単語のDFはその単語を授業名に含む授業数に対応します。**単語が授業名に複数回含まれる場合でも1回と数えます**
def count_df(courses, vocab):
dic={}
for i in range(len(vocab)):
for course in courses:
if # courseがvocab[i]を含むという条件:
# dic[i]を1つ増やす
# dicにiのキーがまだない場合に注意
return dic
# `count_df`関数が完成したら、以下のセルを実行して動作を確認してください。辞書の長さは`910`、単語"プログラミング"(IDは`113`)のDFは`23`、単語"英語"(IDは`15`)のDFは`145`となります。
df = count_df(courses, vocab)
print(len(df))
print(vocab.index('プログラミング'), df[vocab.index('プログラミング')])
print(vocab.index('英語'), df[vocab.index('英語')])
assert len(df) == 910
assert df[113] == 23
assert df[15] == 145
# ## Q3
# ### Q3.1
# リスト`vocab`の各単語を次元とする授業ベクトルを考えます。授業ベクトルの長さはリスト`vocab`の長さと等しく、リスト`vocab`のインデックス`i`の単語`vocab[i]`が授業名に含まれる時、授業ベクトルの`i`番目の要素は`1`、含まれなければ`0`とします。
#
# 以下では、リスト`courses`と`vocab`を引数で受け取り、リスト`courses`の各授業のベクトルを行、リスト`vocab`の各単語を列とした`NumPy`の行列を作成して返す`lec_word_matrix`関数を完成させてください。作成した授業-単語行列は、授業(行)の授業名に単語(列)が含まれていれば、その要素が1であるような行列です。
def lec_word_matrix(courses, vocab):
mat = np.zeros((# 授業数, 単語数)) # 授業数x単語数の0要素の行列を初期化
for i in range(len(courses)):
for j in range(len(vocab)):
if # courses[i]がvocab[j]を含むという条件:
# mat[i, j]に1を代入
return mat
# `lec_word_matrix`関数が完成したら、以下のセルを実行して動作を確認してください。作成した授業-単語行列の要素の値の和は`17406`となります。
bi_matrix = lec_word_matrix(courses, vocab)
np.sum(bi_matrix)
assert np.sum(lec_word_matrix(courses, vocab)) == 17406
# ### Q3.2
# Q3.1で作成した授業-単語行列の各要素を、その授業の授業名に単語が含まれるか否かの1or0ではなく、その授業の授業名に単語が何回含まれるか(TF: Term Frequency)で表した行列を作成して返す`lec_word_tf_matrix`関数を完成させてください。
def lec_word_tf_matrix(courses, vocab):
mat = np.zeros((# 授業数, 単語数)) # 授業数x単語数の0要素の行列を初期化
for i in range(len(courses)):
for j in range(len(vocab)):
if # courses[i]がvocab[j]を含むという条件:
# courses[i]に含まれるvocab[j]の数(ヒント count())をmat[i, j]に代入
return mat
# `lec_word_tf_matrix`関数が完成したら、以下のセルを実行して動作を確認してください。作成した授業-単語行列の要素の値の和は`17732`となります。
tf_matrix = lec_word_tf_matrix(courses, vocab)
np.sum(tf_matrix)
# 授業「総合社会科学高度教養(計量社会科学研究)」には「科学」が2回含まれるので対応する行列の要素の値は2となります。
tf_matrix[courses.index('総合社会科学高度教養(計量社会科学研究)'), vocab.index('科学')]
assert np.sum(lec_word_tf_matrix(courses, vocab)) == 17732
assert lec_word_tf_matrix(courses, vocab)[courses.index('総合社会科学高度教養(計量社会科学研究)'), vocab.index('科学')] == 2
# ### Q3.3
# Q3.2で作成した授業-単語行列の各要素(授業`i`の単語`j`の$TF_{ij}$)にその単語のIDF値を掛けたTFIDF値を要素とする行列を作成して返す`lec_word_tfidf_matrix`関数を完成させてください。作成した行列は変数`tfidf_matrix`で受け取ります。
#
# ここで、授業`i`、単語`j`のTFIDF値は以下のように定義されます。
# ```Python
# TFIDF=TF_ij*log(すべての授業数/単語jのDF)=TF_ij*log(len(courses)/df[j])
# ```
# `log`の計算には`np.log()`を使用してよいです。
def lec_word_tfidf_matrix(courses, vocab, df):
### 引数:
# courses: 授業リスト
# vocab: 単語リスト
# df: DF辞書
### Q3.2のmat[i,j]を"授業名iの単語jのTFIDF値"とした行列を作成するコード ###
return ### 上記で作成した行列を返す ###
# `lec_word_tfidf_matrix`関数が完成したら、以下のセルを実行して動作を確認してください。作成した授業-単語行列の要素の値の和の整数部分は`76994`となります。
tfidf_matrix = lec_word_tfidf_matrix(courses, vocab, df)
int(np.sum(tfidf_matrix))
# 授業「総合社会科学高度教養(計量社会科学研究)」には「科学」が`2`回含まれ、「科学」のDF値は`412`なので対応する行列の要素の値は$2*log(4678/412)\simeq4.85$となります。
print(df[vocab.index('科学')])
print(tfidf_matrix[courses.index('総合社会科学高度教養(計量社会科学研究)'), vocab.index('科学')])
assert int(np.sum(tfidf_matrix)) == 76994
assert df[vocab.index('科学')] == 412
assert int(tfidf_matrix[courses.index('総合社会科学高度教養(計量社会科学研究)'), vocab.index('科学')]*100) == 485
# ## Q4
# Q3.3で作成した授業-単語行列を元に、入力の授業に対してcos類似度に基づいて他のすべての授業との類似度を計算し、類似する授業名をキー、その類似度を値とした辞書を返す以下の`find_similar_course`関数を完成させてください。その際、**入力の授業および類似度が0の授業は辞書に含めない**ようにしてください。cos類似度の計算にはQ1で作成した関数を使ってもよいです。
def find_similar_course(target, tfidf_matrix, courses):
### 引数:
# target: 入力授業のID
# tfidf_matrix: Q3.3で作成した授業-単語行列
# courses: 授業リスト
### 入力授業のベクトルtfidf_matrix[target]と各授業iのベクトルtfidf_matrix[i]の類似度を計算するコード ###
return ### 類似する授業名をキー、その類似度を値とする辞書を返す ###
# `find_similar_course`関数が完成したら、以下のセルを実行して動作を確認してください。
#
# 入力の授業`再生可能エネルギーと公共政策`に類似した上位の授業は、`公共政策、金融市場と公共政策、文化人類学特殊演習(公共政策論)、エネルギー政策、河川流域の環境とその再生、...`となります。
#
# 類似した授業数は68となります。
# +
title="再生可能エネルギーと公共政策" # 入力の授業名
results=find_similar_course(courses.index(title), tfidf_matrix, courses) # 類似授業の辞書を受け取る
print(len(results))
ranking = pd.DataFrame(list(results.items()), columns=['title', "similarity"]) # 類似度が上位の授業を表示
ranking.sort_values(by='similarity', ascending=False).head(10)
# -
assert len(results) == 68
assert ranking.sort_values(by='similarity', ascending=False).iloc[0,0] == '公共政策'
assert int(ranking.sort_values(by='similarity', ascending=False).iloc[0,1]*100) == 64
assert ranking.sort_values(by='similarity', ascending=False).iloc[1,0] == '金融市場と公共政策'
assert int(ranking.sort_values(by='similarity', ascending=False).iloc[1,1]*100) == 48
|
ex4/ex4.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # "Galfitting" with lenstronomy
# An example of using **lenstronomy** without imposing a lensing deflector is the inference of galaxy structural parameters through the fitting of a parameterized surface brightness model (or many thereof).
#
# Specific software packages and algorithm that are designed to perform these tasks may well be faster and more optimized. However, we want to demonstrate the flexibility of **lenstronomy**. Matching the structural properties of lensing and source galaxy is an integral part of lens modelling. Where speed may not be the primary requirement, **lenstronomy** may provide an alternative in python.
#
#
# some standard python imports #
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# ## create mock image as a superposition of different profiles
# In the example below, we generate a galaxy image with a two component Sersic profile at the same center and a compagnion Hernquist light distribution. The data quality is comparable with wide field surveys.
# +
import lenstronomy.Util.simulation_util as sim_util
import lenstronomy.Util.image_util as image_util
from lenstronomy.Data.imaging_data import ImageData
from lenstronomy.Data.psf import PSF
# data specifics
background_rms = .05 # background noise per pixel
exp_time = 100 # exposure time (arbitrary units, flux per pixel is in units #photons/exp_time unit)
numPix = 50 # cutout pixel size
deltaPix = 0.3 # pixel size in arcsec (area per pixel = deltaPix**2)
fwhm = 0.8 # full width half max of PSF
kwargs_data = sim_util.data_configure_simple(numPix, deltaPix, exp_time, background_rms)
data_class = ImageData(**kwargs_data)
# PSF specification
kwargs_psf = {'psf_type': 'GAUSSIAN', 'fwhm': fwhm, 'pixel_size': deltaPix, 'truncation': 6}
psf_class = PSF(**kwargs_psf)
# create a model with three Sersic profiles
# all the models are part of 'lens_light_model_list', meaning that their surface brightness profile are not lensed
lens_light_model_list = ['SERSIC_ELLIPSE', 'SERSIC_ELLIPSE', 'HERNQUIST']
from lenstronomy.LightModel.light_model import LightModel
lightModel = LightModel(lens_light_model_list)
kwargs_1 = {'amp': 100, 'R_sersic': .5, 'n_sersic': 3, 'e1': 0.1, 'e2': 0, 'center_x': 0, 'center_y': 0}
kwargs_2 = {'amp': 100, 'R_sersic': 1.5, 'n_sersic': 1, 'e1': 0.2, 'e2': -0.2, 'center_x': 0, 'center_y': 0}
kwargs_3 = {'amp': 100, 'Rs': 0.3, 'center_x': 3.5, 'center_y': -0.5}
kwargs_light = [kwargs_1, kwargs_2, kwargs_3]
kwargs_numerics = {'supersampling_factor': 4}
from lenstronomy.ImSim.image_model import ImageModel
imageModel = ImageModel(data_class, psf_class, lens_light_model_class=lightModel, kwargs_numerics=kwargs_numerics)
image_sim = imageModel.image(kwargs_lens_light=kwargs_light)
poisson = image_util.add_poisson(image_sim, exp_time=exp_time)
bkg = image_util.add_background(image_sim, sigma_bkd=background_rms)
image_noisy = image_sim + bkg + poisson
data_class.update_data(image_noisy)
kwargs_data['image_data'] = image_noisy
plt.matshow(np.log10(image_noisy), origin='lower')
plt.show()
# +
kwargs_model = {'lens_light_model_list': lens_light_model_list}
kwargs_constraints = {}
kwargs_numerics_galfit = {'supersampling_factor': 2}
kwargs_likelihood = {'check_bounds': True}
image_band = [kwargs_data, kwargs_psf, kwargs_numerics_galfit]
multi_band_list = [image_band]
kwargs_data_joint = {'multi_band_list': multi_band_list, 'multi_band_type': 'multi-linear'}
# +
# lens light model choices
fixed_lens_light = []
kwargs_lens_light_init = []
kwargs_lens_light_sigma = []
kwargs_lower_lens_light = []
kwargs_upper_lens_light = []
# first Sersic component
fixed_lens_light.append({})
kwargs_lens_light_init.append({'R_sersic': .1, 'n_sersic': 4, 'e1': 0, 'e2': 0, 'center_x': 0, 'center_y': 0})
kwargs_lens_light_sigma.append({'n_sersic': 0.5, 'R_sersic': 0.2, 'e1': 0.1, 'e2': 0.1, 'center_x': 0.1, 'center_y': 0.1})
kwargs_lower_lens_light.append({'e1': -0.5, 'e2': -0.5, 'R_sersic': 0.01, 'n_sersic': 0.5, 'center_x': -10, 'center_y': -10})
kwargs_upper_lens_light.append({'e1': 0.5, 'e2': 0.5, 'R_sersic': 10, 'n_sersic': 8, 'center_x': 10, 'center_y': 10})
# second Sersic component
fixed_lens_light.append({})
kwargs_lens_light_init.append({'R_sersic': .5, 'n_sersic': 1, 'e1': 0, 'e2': 0, 'center_x': 0, 'center_y': 0})
kwargs_lens_light_sigma.append({'n_sersic': 0.5, 'R_sersic': 0.2, 'e1': 0.1, 'e2': 0.1, 'center_x': 0.1, 'center_y': 0.1})
kwargs_lower_lens_light.append({'e1': -0.5, 'e2': -0.5, 'R_sersic': 0.01, 'n_sersic': 0.5, 'center_x': -10, 'center_y': -10})
kwargs_upper_lens_light.append({'e1': 0.5, 'e2': 0.5, 'R_sersic': 10, 'n_sersic': 8, 'center_x': 10, 'center_y': 10})
# Hernquist component
fixed_lens_light.append({})
kwargs_lens_light_init.append({'Rs': 0.7, 'center_x': 3., 'center_y': 0.})
kwargs_lens_light_sigma.append({'Rs': 0.3, 'center_x': 0.5, 'center_y': 0.5})
kwargs_lower_lens_light.append({'Rs': 0.01, 'center_x': -10, 'center_y': -10})
kwargs_upper_lens_light.append({'Rs': 10, 'center_x': 10, 'center_y': 10})
lens_light_params = [kwargs_lens_light_init, kwargs_lens_light_sigma, fixed_lens_light, kwargs_lower_lens_light, kwargs_upper_lens_light]
kwargs_params = {'lens_light_model': lens_light_params}
from lenstronomy.Workflow.fitting_sequence import FittingSequence
fitting_seq = FittingSequence(kwargs_data_joint, kwargs_model, kwargs_constraints, kwargs_likelihood, kwargs_params)
fitting_kwargs_list = [['PSO', {'sigma_scale': 1., 'n_particles': 200, 'n_iterations': 200}]]
chain_list = fitting_seq.fit_sequence(fitting_kwargs_list)
lens_result, source_result, lens_light_result, ps_result, cosmo_result = fitting_seq.best_fit()
# +
from lenstronomy.Plots.output_plots import ModelPlot
import lenstronomy.Plots.output_plots as out_plot
modelPlot = ModelPlot(multi_band_list, kwargs_model, lens_result, source_result,
lens_light_result, ps_result, arrow_size=0.02, cmap_string="gist_heat")
for i in range(len(chain_list)):
out_plot.plot_chain_list(chain_list, i)
f, axes = plt.subplots(1, 3, figsize=(16, 8), sharex=False, sharey=False)
modelPlot.data_plot(ax=axes[0])
modelPlot.model_plot(ax=axes[1])
modelPlot.normalized_residual_plot(ax=axes[2], v_min=-6, v_max=6)
f.tight_layout()
#f.subplots_adjust(left=None, bottom=None, right=None, top=None, wspace=0., hspace=0.05)
plt.show()
f, axes = plt.subplots(1, 3, figsize=(16, 8), sharex=False, sharey=False)
modelPlot.decomposition_plot(ax=axes[0], text='Lens light', lens_light_add=True, unconvolved=True)
modelPlot.decomposition_plot(ax=axes[1], text='Lens light convolved', lens_light_add=True)
modelPlot.subtract_from_data_plot(ax=axes[2], text='Data - Lens Light', lens_light_add=True)
f.tight_layout()
#f.subplots_adjust(left=None, bottom=None, right=None, top=None, wspace=0., hspace=0.05)
plt.show()
print(lens_light_result)
# -
|
lenstronomy_extensions/Notebooks/galfitting.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# %matplotlib inline
import mxnet as mx
from mxnet import nd
from mxnet.contrib.ndarray import MultiBoxPrior
from mxnet.gluon.contrib import nn as nn_contrib
from mxnet.gluon import nn
ctx = mx.gpu()
# ### Predict classes
# - channel `i*(num_class+1)` store the scores for this box contains only background
# - channel `i*(num_class+1)+1+j` store the scores for this box contains an object from the *j*-th class
def class_predictor(num_anchors, num_classes):
return nn.Conv2D(num_anchors * (num_classes + 1), 3, padding=1)
# ### Predict anchor boxes
# - $t_x = (Y_x - b_x) / b_{width}$
# - $t_y = (Y_y - b_y) / b_{height}$
# - $t_{width} = (Y_{width} - b_{width}) / b_{width}$
# - $t_{height} = (Y_{height} - b_{height}) / b_{height}$
def box_predictor(num_anchors):
return nn.Conv2D(num_anchors * 4, 3, padding=1)
# ### Manage preditions from multiple layers
# +
def flatten_prediction(pred):
return nd.flatten(nd.transpose(pred, axes=(0, 2, 3, 1)))
def concat_predictions(preds):
return nd.concat(*preds, dim=1)
# -
# ### Down-sample features
def dp_layer(nfilters, stride, expension_constant):
out = nn.HybridSequential()
out.add(nn.Conv2D(nfilters, 3, strides=stride, padding=1, groups=nfilters, use_bias=False))
out.add(nn.BatchNorm(use_global_stats=False, epsilon=1e-05, momentum=0.9, axis=1))
out.add(nn.Activation('relu'))
out.add(nn.Conv2D(nfilters*expension_constant, 1, strides=1, padding=0, use_bias=False))
out.add(nn.BatchNorm(use_global_stats=False, epsilon=1e-05, momentum=0.9, axis=1))
out.add(nn.Activation('relu'))
return out
# ### Scale units
global alpha
alpha = 0.25
num_filters = int(32*alpha)
# ### Body network
# +
from mxnet import gluon
def s16():
out = nn.HybridSequential()
# conv_0 layer
out.add(nn.Conv2D(num_filters, 3, strides=2, padding=1, use_bias=False))
out.add(nn.BatchNorm(use_global_stats=False, epsilon=1e-05, momentum=0.9, axis=1))
out.add(nn.Activation('relu'))
# conv_1 layer
out.add(dp_layer(num_filters, 1, 2))
# conv_2 layer
out.add(dp_layer(num_filters*2, 2, 2))
# conv_3 layer
out.add(dp_layer(num_filters*4, 1, 1))
out.add(nn.Conv2D(num_filters*4, 3, strides=2, padding=1, groups=num_filters*4, use_bias=False))
out.load_parameters("weights/mobilenet_0_25_s16_dist.params", ctx=ctx)
out.hybridize()
return out
def s32():
out = nn.HybridSequential()
# from last layer
out.add(nn.BatchNorm(use_global_stats=False, epsilon=1e-05, momentum=0.9, axis=1))
out.add(nn.Activation('relu'))
out.add(nn.Conv2D(num_filters*8, 1, strides=1, padding=0, use_bias=False))
out.add(nn.BatchNorm(use_global_stats=False, epsilon=1e-05, momentum=0.9, axis=1))
out.add(nn.Activation('relu'))
# conv_4_layer
out.add(dp_layer(num_filters*8, 1, 1))
out.add(nn.Conv2D(num_filters*8, 3, strides=2, padding=1, groups=num_filters*8, use_bias=False))
out.load_parameters("weights/mobilenet_0_25_s32_dist.params", ctx=ctx)
out.hybridize()
return out
def b1():
out = nn.HybridSequential()
# from last layer
out.add(nn.BatchNorm(use_global_stats=False, epsilon=1e-05, momentum=0.9, axis=1))
out.add(nn.Activation('relu'))
out.add(nn.Conv2D(num_filters*16, 1, strides=1, padding=0, use_bias=False))
out.add(nn.BatchNorm(use_global_stats=False, epsilon=1e-05, momentum=0.9, axis=1))
out.add(nn.Activation('relu'))
# conv_6_layer
out.add(dp_layer(num_filters*16, 1, 1))
out.add(nn.Conv2D(num_filters*16, 3, strides=2, padding=1, groups=num_filters*16, use_bias=False))
out.initialize(mx.init.Xavier(magnitude=2), ctx=ctx)
out.hybridize()
return out
def b2():
out = nn.HybridSequential()
# from last layer
out.add(nn.BatchNorm(use_global_stats=False, epsilon=1e-05, momentum=0.9, axis=1))
out.add(nn.Activation('relu'))
out.add(nn.Conv2D(num_filters*16, 1, strides=1, padding=0, use_bias=False))
out.add(nn.BatchNorm(use_global_stats=False, epsilon=1e-05, momentum=0.9, axis=1))
out.add(nn.Activation('relu'))
# conv_7_layer
out.add(dp_layer(num_filters*16, 1, 1))
out.add(nn.Conv2D(num_filters*16, 3, strides=2, padding=1, groups=num_filters*16, use_bias=False))
out.initialize(mx.init.Xavier(magnitude=2), ctx=ctx)
out.hybridize()
return out
def b3():
out = nn.HybridSequential()
# from last layer
out.add(nn.BatchNorm(use_global_stats=False, epsilon=1e-05, momentum=0.9, axis=1))
out.add(nn.Activation('relu'))
out.add(nn.Conv2D(num_filters*16, 1, strides=1, padding=0, use_bias=False))
out.add(nn.BatchNorm(use_global_stats=False, epsilon=1e-05, momentum=0.9, axis=1))
out.add(nn.Activation('relu'))
# conv_8_layer
out.add(dp_layer(num_filters*16, 1, 1))
out.add(nn.Conv2D(num_filters*16, 3, strides=2, padding=1, groups=num_filters*16, use_bias=False))
out.initialize(mx.init.Xavier(magnitude=2), ctx=ctx)
out.hybridize()
return out
def b4():
out = nn.HybridSequential()
# from last layer
out.add(nn.BatchNorm(use_global_stats=False, epsilon=1e-05, momentum=0.9, axis=1))
out.add(nn.Activation('relu'))
out.add(nn.Conv2D(num_filters*16, 1, strides=1, padding=0, use_bias=False))
out.add(nn.BatchNorm(use_global_stats=False, epsilon=1e-05, momentum=0.9, axis=1))
out.add(nn.Activation('relu'))
# conv_9_layer
out.add(dp_layer(num_filters*16, 1, 1))
out.add(nn.Conv2D(num_filters*16, 3, strides=2, padding=1, groups=num_filters*16, use_bias=False))
out.initialize(mx.init.Xavier(magnitude=2), ctx=ctx)
out.hybridize()
return out
# -
# ### Create an SSD model
def ssd_model(num_anchors, num_classes):
class_preds = nn.Sequential()
box_preds = nn.Sequential()
for scale in range(6):
class_preds.add(class_predictor(num_anchors, num_classes))
box_preds.add(box_predictor(num_anchors))
class_preds.initialize(mx.init.Xavier(magnitude=2), ctx=ctx)
box_preds.initialize(mx.init.Xavier(magnitude=2), ctx=ctx)
return s16(), s32(), b1(), b2(), b3(), b4(), class_preds, box_preds
# ### Forward
def ssd_forward(x, s16, s32, b1, b2, b3, b4, class_preds, box_preds, sizes, ratios):
default_anchors = []
predicted_boxes = []
predicted_classes = []
x = s16(x)
default_anchors.append(MultiBoxPrior(x, sizes=sizes[0], ratios=ratios[0]))
predicted_boxes.append(flatten_prediction(box_preds[0](x)))
predicted_classes.append(flatten_prediction(class_preds[0](x)))
x = s32(x)
default_anchors.append(MultiBoxPrior(x, sizes=sizes[1], ratios=ratios[1]))
predicted_boxes.append(flatten_prediction(box_preds[1](x)))
predicted_classes.append(flatten_prediction(class_preds[1](x)))
x = b1(x)
default_anchors.append(MultiBoxPrior(x, sizes=sizes[2], ratios=ratios[2]))
predicted_boxes.append(flatten_prediction(box_preds[2](x)))
predicted_classes.append(flatten_prediction(class_preds[2](x)))
x = b2(x)
default_anchors.append(MultiBoxPrior(x, sizes=sizes[3], ratios=ratios[3]))
predicted_boxes.append(flatten_prediction(box_preds[3](x)))
predicted_classes.append(flatten_prediction(class_preds[3](x)))
x = b3(x)
default_anchors.append(MultiBoxPrior(x, sizes=sizes[4], ratios=ratios[4]))
predicted_boxes.append(flatten_prediction(box_preds[4](x)))
predicted_classes.append(flatten_prediction(class_preds[4](x)))
x = b4(x)
default_anchors.append(MultiBoxPrior(x, sizes=sizes[5], ratios=ratios[5]))
predicted_boxes.append(flatten_prediction(box_preds[5](x)))
predicted_classes.append(flatten_prediction(class_preds[5](x)))
return default_anchors, predicted_classes, predicted_boxes
# ### Put all things together
from mxnet import gluon
class SSD(gluon.Block):
def __init__(self, num_classes, **kwargs):
super(SSD, self).__init__(**kwargs)
self.anchor_sizes = [[0.04, 0.1],[0.1,0.26],[0.26,0.42],[0.42,0.58],[0.58,0.74],[0.74,0.9],[0.9,1.06]]
self.anchor_ratios = [[1, 2, .5]] * 6
self.num_classes = num_classes
with self.name_scope():
self.s16, self.s32, self.b1, self.b2, self.b3, self.b4, self.class_preds, self.box_preds = ssd_model(4, num_classes)
def forward(self, x):
default_anchors, predicted_classes, predicted_boxes = ssd_forward(x, self.s16, self.s32, self.b1, self.b2, self.b3, self.b4,
self.class_preds, self.box_preds, self.anchor_sizes, self.anchor_ratios)
anchors = concat_predictions(default_anchors)
box_preds = concat_predictions(predicted_boxes)
class_preds = concat_predictions(predicted_classes)
class_preds = nd.reshape(class_preds, shape=(0, -1, self.num_classes + 1))
return anchors, class_preds, box_preds
# ### Outputs of SSD
net = SSD(2)
#net.initialize(mx.init.Xavier(magnitude=2), ctx=ctx)
net.load_parameters("process/ssd_300.params",ctx=ctx)
x = nd.zeros((1, 3, 512, 512),ctx=ctx)
default_anchors, class_predictions, box_predictions = net(x)
print('Outputs:', 'anchors', default_anchors.shape, 'class prediction', class_predictions.shape, 'box prediction', box_predictions.shape)
# ### Load dataset
# +
from source.NACDDetection import NACDDetection
train_dataset = NACDDetection(splits=[('NACDwNegswAugCropped', 'train')])
test_dataset = NACDDetection(splits=[('NACDwNegswAugCropped', 'test')])
print('Training images:', len(train_dataset))
print('Test images:', len(test_dataset))
# -
from source import NACDTransform
width, height = 512, 512
train_transform = NACDTransform.NACDDefaultTransform(width, height, False)
test_transform = NACDTransform.NACDDefaultTransform(width, height, True)
from gluoncv.data.transforms import presets
from gluoncv import utils
from mxnet import nd
from matplotlib import pyplot as plt
from gluoncv.utils import viz
train_image, train_label = test_dataset[0]
bboxes = train_label[:, :4]
cids = train_label[:, 4:5]
print('image:', train_image.shape)
print('bboxes:', bboxes.shape, 'class ids:', cids.shape)
train_image2, train_label2 = train_transform(train_image, train_label)
print('tensor shape:', train_image2.shape)
# +
from gluoncv.data.batchify import Tuple, Stack, Pad
from mxnet.gluon.data import DataLoader
batch_size = 16
num_workers = 4
batchify_fn = Tuple(Stack(), Pad(pad_val=-1))
train_loader = DataLoader(train_dataset.transform(train_transform), batch_size, shuffle=True,
batchify_fn=batchify_fn, last_batch='rollover', num_workers=num_workers)
test_loader = DataLoader(test_dataset.transform(test_transform), batch_size, shuffle=False,
batchify_fn=batchify_fn, last_batch='keep', num_workers=num_workers)
for ib, batch in enumerate(test_loader):
if ib > 3:
break
print('data:', batch[0].shape, 'label:', batch[1].shape)
# -
train_image2 = train_image2.transpose((1, 2, 0)) * nd.array((0.229, 0.224, 0.225)) + nd.array((0.485, 0.456, 0.406))
train_image2 = (train_image2 * 255).clip(0, 255)
ax = viz.plot_bbox(train_image2.asnumpy(), train_label2[:, :4],
labels=train_label2[:, 4:5],
class_names=train_dataset.classes)
plt.show()
# ## Train
from mxnet.contrib.ndarray import MultiBoxTarget
def training_targets(default_anchors, class_predicts, labels):
class_predicts = nd.transpose(class_predicts, axes=(0, 2, 1))
z = MultiBoxTarget(anchor=default_anchors.as_in_context(mx.cpu()), label=labels.as_in_context(mx.cpu()), cls_pred=class_predicts.as_in_context(mx.cpu()))
box_target = z[0].as_in_context(ctx) # box offset target for (x, y, width, height)
box_mask = z[1].as_in_context(ctx) # mask is used to ignore box offsets we don't want to penalize, e.g. negative samples
cls_target = z[2].as_in_context(ctx) # cls_target is an array of labels for all anchors boxes
return box_target, box_mask, cls_target
def convertlbl(y):
mtrx = y[:,:,0:4]
mtrx = mtrx.asnumpy()
mtrx[mtrx == -1] = -width
mtrx = mtrx/512
return mx.nd.concat(nd.expand_dims(y[:,:,4],2),mx.nd.array(mtrx),dim=2)
# +
class FocalLoss(gluon.loss.Loss):
def __init__(self, axis=-1, alpha=0.25, gamma=2, batch_axis=0, **kwargs):
super(FocalLoss, self).__init__(None, batch_axis, **kwargs)
self._axis = axis
self._alpha = alpha
self._gamma = gamma
def hybrid_forward(self, F, output, label):
output = F.softmax(output)
pt = F.pick(output, label, axis=self._axis, keepdims=True)
loss = -self._alpha * ((1 - pt) ** self._gamma) * F.log(pt)
return F.mean(loss, axis=self._batch_axis, exclude=True)
# cls_loss = gluon.loss.SoftmaxCrossEntropyLoss()
cls_loss = FocalLoss()
print(cls_loss)
# +
class SmoothL1Loss(gluon.loss.Loss):
def __init__(self, batch_axis=0, **kwargs):
super(SmoothL1Loss, self).__init__(None, batch_axis, **kwargs)
def hybrid_forward(self, F, output, label, mask):
loss = F.smooth_l1((output - label) * mask, scalar=1.0)
return F.mean(loss, self._batch_axis, exclude=True)
box_loss = SmoothL1Loss()
print(box_loss)
# -
# ### Initialize parameters
import time
from mxnet import autograd as ag
from gluoncv.loss import SSDMultiBoxLoss
# +
# loop params
epochs = 100
start_epoch = 1
# initialize trainer
net.collect_params().reset_ctx(ctx)
trainer = gluon.Trainer(net.collect_params(), 'sgd', {'learning_rate': 1e-1, 'wd': 4e-5})
# evaluation metrics
cls_metric = mx.metric.Accuracy()
box_metric = mx.metric.MAE()
cls_metric_test = mx.metric.Accuracy()
box_metric_test = mx.metric.MAE()
# training loop
for epoch in range(start_epoch, epochs):
# reset iterator and tick
#train_data.reset()
cls_metric.reset()
box_metric.reset()
tic = time.time()
train_loss = 0
# iterate through all batch
for i, batch in enumerate(train_loader):
# record gradients
with ag.record():
x = batch[0].as_in_context(ctx)
y = batch[1].as_in_context(ctx)
lbl = convertlbl(batch[1])
default_anchors, class_predictions, box_predictions = net(x)
box_target, box_mask, cls_target = training_targets(default_anchors, class_predictions, lbl)
# losses
loss1 = cls_loss(class_predictions, cls_target)
loss2 = box_loss(box_predictions, box_target, box_mask)
# sum all losses
loss = loss1 + loss2
train_loss += nd.sum(loss).asscalar()
# backpropagate
loss.backward()
# apply
trainer.step(batch_size, ignore_stale_grad=True)
# update metrics
cls_metric.update([cls_target], [nd.transpose(class_predictions, (0, 2, 1))])
box_metric.update([box_target], [box_predictions * box_mask])
#if (i + 1) % log_interval == 0:
toc = time.time()
name1_train, val1_train = cls_metric.get()
name2_train, val2_train = box_metric.get()
cls_metric_test.reset()
box_metric_test.reset()
tic = time.time()
test_loss = 0
for i, batch in enumerate(test_loader):
# record gradients
x = batch[0].as_in_context(ctx)
y = batch[1].as_in_context(ctx)
lbl = convertlbl(batch[1])
default_anchors, class_predictions, box_predictions = net(x)
box_target, box_mask, cls_target = training_targets(default_anchors, class_predictions, lbl)
# losses
loss1 = cls_loss(class_predictions, cls_target)
loss2 = box_loss(box_predictions, box_target, box_mask)
# sum all losses
loss = loss1 + loss2
test_loss += nd.sum(loss).asscalar()
# update metrics
cls_metric_test.update([cls_target], [nd.transpose(class_predictions, (0, 2, 1))])
box_metric_test.update([box_target], [box_predictions * box_mask])
#if (i + 1) % log_interval == 0:
toc = time.time()
name1_test, val1_test = cls_metric_test.get()
name2_test, val2_test = box_metric_test.get()
print('epoch:%3d;\t train:%.6e;%f;%.6e;\t test:%.6e;%f;%.6e'
%(epoch, train_loss/len(train_dataset), val1_train, val2_train, test_loss/len(test_dataset), val1_test, val2_test))
net.save_parameters('process/ssd_%d.params' % epoch)
# + active=""
# # detached
# epoch: 1; train:1.142976e-02;0.982709;2.026818e-03; test:5.950928e-03;0.994906;1.982350e-03
# epoch: 2; train:5.194239e-03;0.996069;2.009641e-03; test:4.234332e-03;0.996789;1.951143e-03
# epoch: 3; train:4.158913e-03;0.996979;2.006346e-03; test:3.657099e-03;0.997238;1.947551e-03
# epoch: 4; train:3.677135e-03;0.997343;1.992840e-03; test:3.354415e-03;0.997519;1.946653e-03
# epoch: 5; train:3.401528e-03;0.997572;1.996442e-03; test:3.117369e-03;0.997727;1.929599e-03
# epoch: 6; train:3.208723e-03;0.997733;1.996436e-03; test:2.934627e-03;0.997840;1.941947e-03
# epoch: 7; train:3.043061e-03;0.997848;1.982956e-03; test:2.810119e-03;0.997902;1.946837e-03
# epoch: 8; train:2.934248e-03;0.997902;1.986833e-03; test:2.702097e-03;0.997948;1.948930e-03
# epoch: 9; train:2.830745e-03;0.997944;1.982301e-03; test:2.612207e-03;0.998003;1.936180e-03
# epoch: 10; train:2.762063e-03;0.997970;1.982813e-03; test:2.515140e-03;0.998042;1.909976e-03
# epoch: 11; train:2.703538e-03;0.997994;1.982351e-03; test:2.459528e-03;0.998043;1.915174e-03
# epoch: 12; train:2.639759e-03;0.998017;1.974574e-03; test:2.436586e-03;0.998037;1.936900e-03
# epoch: 13; train:2.602960e-03;0.998031;1.978068e-03; test:2.354643e-03;0.998098;1.884615e-03
# epoch: 14; train:2.561904e-03;0.998052;1.968779e-03; test:2.341381e-03;0.998092;1.909436e-03
# epoch: 15; train:2.534079e-03;0.998057;1.975983e-03; test:2.291877e-03;0.998124;1.888833e-03
# epoch: 16; train:2.511962e-03;0.998066;1.976526e-03; test:2.308066e-03;0.998085;1.939067e-03
# epoch: 17; train:2.472376e-03;0.998087;1.965343e-03; test:2.222756e-03;0.998167;1.863912e-03
# epoch: 18; train:2.446823e-03;0.998099;1.961130e-03; test:2.255968e-03;0.998114;1.925121e-03
# epoch: 19; train:2.441774e-03;0.998097;1.973160e-03; test:2.205283e-03;0.998153;1.892387e-03
# epoch: 20; train:2.409007e-03;0.998111;1.963464e-03; test:2.190169e-03;0.998156;1.897249e-03
# epoch: 21; train:2.388927e-03;0.998122;1.960642e-03; test:2.196097e-03;0.998152;1.914378e-03
# epoch: 22; train:2.383896e-03;0.998123;1.964754e-03; test:2.152177e-03;0.998178;1.888872e-03
# epoch: 23; train:2.350935e-03;0.998136;1.954015e-03; test:2.194683e-03;0.998135;1.946209e-03
# epoch: 24; train:2.346067e-03;0.998131;1.962783e-03; test:2.134656e-03;0.998188;1.898344e-03
# epoch: 25; train:2.323595e-03;0.998135;1.961664e-03; test:2.117964e-03;0.998200;1.878940e-03
# epoch: 26; train:2.311218e-03;0.998142;1.957463e-03; test:2.139356e-03;0.998164;1.924112e-03
# epoch: 27; train:2.299307e-03;0.998146;1.951310e-03; test:2.107378e-03;0.998181;1.900978e-03
# epoch: 28; train:2.291833e-03;0.998144;1.961066e-03; test:2.061676e-03;0.998218;1.863837e-03
# epoch: 29; train:2.292896e-03;0.998135;1.970330e-03; test:2.082349e-03;0.998193;1.895989e-03
# epoch: 30; train:2.280118e-03;0.998142;1.961631e-03; test:2.078096e-03;0.998178;1.903263e-03
# epoch: 31; train:2.260903e-03;0.998153;1.953065e-03; test:2.095484e-03;0.998155;1.931414e-03
# epoch: 32; train:2.245266e-03;0.998158;1.945520e-03; test:2.075902e-03;0.998171;1.919849e-03
# epoch: 33; train:2.241738e-03;0.998156;1.950018e-03; test:2.025873e-03;0.998217;1.870370e-03
# epoch: 34; train:2.236749e-03;0.998151;1.953592e-03; test:2.039586e-03;0.998205;1.889697e-03
# epoch: 35; train:2.229998e-03;0.998150;1.952954e-03; test:2.038359e-03;0.998191;1.898776e-03
# epoch: 36; train:2.207965e-03;0.998168;1.935480e-03; test:2.021230e-03;0.998181;1.890462e-03
# epoch: 37; train:2.212996e-03;0.998156;1.948612e-03; test:1.988168e-03;0.998223;1.857528e-03
# epoch: 38; train:2.200792e-03;0.998167;1.935721e-03; test:1.979053e-03;0.998227;1.852233e-03
# epoch: 39; train:2.204709e-03;0.998154;1.949604e-03; test:1.961526e-03;0.998240;1.840145e-03
# epoch: 40; train:2.201101e-03;0.998155;1.949461e-03; test:1.993912e-03;0.998193;1.878827e-03
# epoch: 41; train:2.189703e-03;0.998157;1.942740e-03; test:1.989849e-03;0.998205;1.878617e-03
# epoch: 42; train:2.179876e-03;0.998158;1.939266e-03; test:1.997957e-03;0.998173;1.895185e-03
# epoch: 43; train:2.182080e-03;0.998161;1.938708e-03; test:1.984803e-03;0.998196;1.882469e-03
# epoch: 44; train:2.184544e-03;0.998147;1.949377e-03; test:1.989881e-03;0.998181;1.893322e-03
# epoch: 45; train:2.185554e-03;0.998136;1.956369e-03; test:1.960152e-03;0.998200;1.862091e-03
# epoch: 46; train:2.164659e-03;0.998157;1.936824e-03; test:1.966413e-03;0.998174;1.885044e-03
# epoch: 47; train:2.153890e-03;0.998157;1.933145e-03; test:1.974376e-03;0.998161;1.894998e-03
# epoch: 48; train:2.150771e-03;0.998156;1.933808e-03; test:1.935181e-03;0.998191;1.860862e-03
# epoch: 49; train:2.151517e-03;0.998146;1.941098e-03; test:1.932593e-03;0.998188;1.860501e-03
# epoch: 50; train:2.139217e-03;0.998146;1.936553e-03; test:1.944881e-03;0.998170;1.879081e-03
# epoch: 51; train:2.135019e-03;0.998147;1.932027e-03; test:1.923505e-03;0.998177;1.860166e-03
# epoch: 52; train:2.126196e-03;0.998150;1.928114e-03; test:1.914599e-03;0.998184;1.862451e-03
# epoch: 53; train:2.120271e-03;0.998151;1.927391e-03; test:1.909548e-03;0.998175;1.858101e-03
# epoch: 54; train:2.116308e-03;0.998154;1.924875e-03; test:1.922235e-03;0.998154;1.878517e-03
# epoch: 55; train:2.117089e-03;0.998140;1.935076e-03; test:1.909469e-03;0.998174;1.861899e-03
# epoch: 56; train:2.111890e-03;0.998147;1.925975e-03; test:1.872411e-03;0.998188;1.833489e-03
# epoch: 57; train:2.089831e-03;0.998159;1.913233e-03; test:1.881727e-03;0.998178;1.851595e-03
# epoch: 58; train:2.100960e-03;0.998139;1.930461e-03; test:1.895495e-03;0.998137;1.860015e-03
# epoch: 59; train:2.090515e-03;0.998156;1.915678e-03; test:1.872900e-03;0.998178;1.848193e-03
# epoch: 60; train:2.089127e-03;0.998145;1.925394e-03; test:1.853789e-03;0.998173;1.833551e-03
# epoch: 61; train:2.083314e-03;0.998145;1.920246e-03; test:1.895247e-03;0.998151;1.876001e-03
# epoch: 62; train:2.079127e-03;0.998146;1.917338e-03; test:1.861901e-03;0.998168;1.846308e-03
# epoch: 63; train:2.076532e-03;0.998141;1.919341e-03; test:1.826680e-03;0.998202;1.811602e-03
# epoch: 64; train:2.057156e-03;0.998160;1.900993e-03; test:1.880602e-03;0.998147;1.871994e-03
# epoch: 65; train:2.059280e-03;0.998146;1.911460e-03; test:1.861245e-03;0.998140;1.861563e-03
# epoch: 66; train:2.052574e-03;0.998149;1.907208e-03; test:1.856796e-03;0.998171;1.857425e-03
# epoch: 67; train:2.054312e-03;0.998149;1.908779e-03; test:1.830579e-03;0.998162;1.831204e-03
# epoch: 68; train:2.052651e-03;0.998148;1.911483e-03; test:1.830642e-03;0.998190;1.828219e-03
# epoch: 69; train:2.054516e-03;0.998134;1.919294e-03; test:1.859446e-03;0.998129;1.862922e-03
# epoch: 70; train:2.051749e-03;0.998140;1.915174e-03; test:1.859713e-03;0.998121;1.870316e-03
# epoch: 71; train:2.047911e-03;0.998135;1.918965e-03; test:1.840182e-03;0.998153;1.855044e-03
# epoch: 72; train:2.034744e-03;0.998148;1.901537e-03; test:1.786083e-03;0.998187;1.790220e-03
# epoch: 73; train:2.030020e-03;0.998154;1.900152e-03; test:1.804264e-03;0.998194;1.815234e-03
# epoch: 74; train:2.022393e-03;0.998153;1.896800e-03; test:1.794604e-03;0.998169;1.820397e-03
# epoch: 75; train:2.022016e-03;0.998149;1.896774e-03; test:1.823751e-03;0.998152;1.845379e-03
# epoch: 76; train:2.010884e-03;0.998161;1.891553e-03; test:1.795239e-03;0.998185;1.818036e-03
# epoch: 77; train:2.024395e-03;0.998145;1.904379e-03; test:1.776688e-03;0.998192;1.799722e-03
# epoch: 78; train:2.022971e-03;0.998150;1.899538e-03; test:1.809304e-03;0.998147;1.835538e-03
# epoch: 79; train:2.014794e-03;0.998145;1.900646e-03; test:1.805820e-03;0.998141;1.841452e-03
# epoch: 80; train:2.000818e-03;0.998161;1.885137e-03; test:1.795081e-03;0.998174;1.828082e-03
# epoch: 81; train:1.995158e-03;0.998157;1.884493e-03; test:1.776539e-03;0.998177;1.811125e-03
# epoch: 82; train:1.994048e-03;0.998159;1.884103e-03; test:1.783530e-03;0.998168;1.821404e-03
# epoch: 83; train:2.004568e-03;0.998151;1.893677e-03; test:1.766683e-03;0.998176;1.797622e-03
# epoch: 84; train:1.997345e-03;0.998149;1.890405e-03; test:1.775889e-03;0.998187;1.813888e-03
# epoch: 85; train:1.987978e-03;0.998158;1.883820e-03; test:1.773353e-03;0.998164;1.810636e-03
# epoch: 86; train:1.999321e-03;0.998142;1.894635e-03; test:1.759538e-03;0.998166;1.808876e-03
# epoch: 87; train:1.997971e-03;0.998138;1.898382e-03; test:1.779130e-03;0.998151;1.819722e-03
# epoch: 88; train:1.980534e-03;0.998158;1.880547e-03; test:1.742700e-03;0.998196;1.791956e-03
# epoch: 89; train:1.978422e-03;0.998151;1.882145e-03; test:1.766175e-03;0.998152;1.817176e-03
# epoch: 90; train:1.975442e-03;0.998154;1.881371e-03; test:1.758351e-03;0.998176;1.795834e-03
# epoch: 91; train:1.984499e-03;0.998144;1.889861e-03; test:1.752967e-03;0.998124;1.793375e-03
# epoch: 92; train:1.973727e-03;0.998147;1.887329e-03; test:1.756537e-03;0.998158;1.804491e-03
# epoch: 93; train:1.969905e-03;0.998149;1.880354e-03; test:1.750057e-03;0.998147;1.799334e-03
# epoch: 94; train:1.970800e-03;0.998150;1.881868e-03; test:1.755878e-03;0.998159;1.810874e-03
# epoch: 95; train:1.962106e-03;0.998156;1.874172e-03; test:1.722353e-03;0.998183;1.777675e-03
# epoch: 96; train:1.973104e-03;0.998149;1.883841e-03; test:1.700250e-03;0.998205;1.753466e-03
# epoch: 97; train:1.961292e-03;0.998153;1.878947e-03; test:1.769909e-03;0.998138;1.836047e-03
# epoch: 98; train:1.952597e-03;0.998162;1.866912e-03; test:1.752998e-03;0.998167;1.816530e-03
# epoch: 99; train:1.956106e-03;0.998156;1.872133e-03; test:1.697247e-03;0.998205;1.750872e-03
#
# # attached
# epoch:100; train:1.941430e-03;0.998161;1.863020e-03; test:1.739691e-03;0.998193;1.808003e-03
# epoch:101; train:1.936580e-03;0.998168;1.857441e-03; test:1.714077e-03;0.998162;1.774645e-03
# epoch:102; train:1.947499e-03;0.998152;1.871150e-03; test:1.698287e-03;0.998197;1.758418e-03
# epoch:103; train:1.944474e-03;0.998151;1.874674e-03; test:1.702367e-03;0.998172;1.768250e-03
# epoch:104; train:1.924715e-03;0.998168;1.856019e-03; test:1.724661e-03;0.998179;1.801624e-03
# epoch:105; train:1.939456e-03;0.998152;1.870907e-03; test:1.714528e-03;0.998169;1.785630e-03
# epoch:106; train:1.927463e-03;0.998155;1.863835e-03; test:1.718806e-03;0.998184;1.800176e-03
# epoch:107; train:1.916820e-03;0.998167;1.850669e-03; test:1.685695e-03;0.998145;1.745721e-03
# epoch:108; train:1.913406e-03;0.998162;1.853086e-03; test:1.686859e-03;0.998219;1.774500e-03
# epoch:109; train:1.920340e-03;0.998162;1.857672e-03; test:1.684705e-03;0.998210;1.765106e-03
# epoch:110; train:1.909942e-03;0.998173;1.845541e-03; test:1.647863e-03;0.998279;1.733184e-03
# epoch:111; train:1.909420e-03;0.998166;1.852941e-03; test:1.686377e-03;0.998183;1.768925e-03
# epoch:112; train:1.893877e-03;0.998173;1.837463e-03; test:1.673676e-03;0.998234;1.759512e-03
# epoch:113; train:1.905638e-03;0.998164;1.849512e-03; test:1.654255e-03;0.998240;1.741051e-03
# epoch:114; train:1.904039e-03;0.998164;1.847921e-03; test:1.658849e-03;0.998243;1.751753e-03
# epoch:115; train:1.894228e-03;0.998170;1.840012e-03; test:1.690349e-03;0.998203;1.780372e-03
# epoch:116; train:1.887090e-03;0.998184;1.831074e-03; test:1.656691e-03;0.998231;1.743756e-03
# epoch:117; train:1.888524e-03;0.998175;1.838113e-03; test:1.672543e-03;0.998202;1.759107e-03
# epoch:118; train:1.881834e-03;0.998179;1.831527e-03; test:1.661895e-03;0.998232;1.754573e-03
# epoch:119; train:1.884258e-03;0.998177;1.836630e-03; test:1.649608e-03;0.998238;1.748253e-03
# epoch:120; train:1.873708e-03;0.998178;1.829041e-03; test:1.646181e-03;0.998264;1.757325e-03
# epoch:121; train:1.874073e-03;0.998181;1.825581e-03; test:1.636852e-03;0.998259;1.750387e-03
# epoch:122; train:1.868644e-03;0.998180;1.825202e-03; test:1.652274e-03;0.998227;1.747894e-03
# epoch:123; train:1.881636e-03;0.998174;1.835390e-03; test:1.623525e-03;0.998247;1.714875e-03
# epoch:124; train:1.864054e-03;0.998184;1.820850e-03; test:1.625001e-03;0.998273;1.718847e-03
# epoch:125; train:1.863748e-03;0.998184;1.821219e-03; test:1.609891e-03;0.998277;1.707407e-03
# epoch:126; train:1.870754e-03;0.998174;1.828245e-03; test:1.654340e-03;0.998249;1.765044e-03
# epoch:127; train:1.854250e-03;0.998184;1.813799e-03; test:1.626063e-03;0.998243;1.723711e-03
# epoch:128; train:1.850847e-03;0.998183;1.813095e-03; test:1.631441e-03;0.998225;1.730116e-03
# epoch:129; train:1.854354e-03;0.998186;1.814591e-03; test:1.622610e-03;0.998264;1.725540e-03
# epoch:130; train:1.858623e-03;0.998180;1.820084e-03; test:1.639518e-03;0.998227;1.747886e-03
# epoch:131; train:1.853384e-03;0.998186;1.811805e-03; test:1.577884e-03;0.998309;1.676688e-03
# epoch:132; train:1.849472e-03;0.998180;1.815238e-03; test:1.596427e-03;0.998271;1.706395e-03
# epoch:133; train:1.842326e-03;0.998188;1.809325e-03; test:1.600355e-03;0.998297;1.700598e-03
# epoch:134; train:1.849689e-03;0.998173;1.816845e-03; test:1.617905e-03;0.998268;1.727452e-03
# epoch:135; train:1.836732e-03;0.998187;1.801475e-03; test:1.599201e-03;0.998262;1.700689e-03
# epoch:136; train:1.847632e-03;0.998175;1.820610e-03; test:1.581050e-03;0.998304;1.693631e-03
# epoch:137; train:1.843884e-03;0.998186;1.807994e-03; test:1.590779e-03;0.998294;1.697701e-03
# epoch:138; train:1.826498e-03;0.998194;1.796218e-03; test:1.598732e-03;0.998272;1.693337e-03
# epoch:139; train:1.840782e-03;0.998177;1.812786e-03; test:1.607012e-03;0.998275;1.712516e-03
# epoch:140; train:1.822266e-03;0.998197;1.792532e-03; test:1.591641e-03;0.998290;1.700585e-03
# epoch:141; train:1.821947e-03;0.998196;1.791506e-03; test:1.585708e-03;0.998289;1.689861e-03
# epoch:142; train:1.823303e-03;0.998191;1.793924e-03; test:1.594933e-03;0.998285;1.715050e-03
# epoch:143; train:1.818241e-03;0.998198;1.788694e-03; test:1.591116e-03;0.998284;1.716457e-03
# epoch:144; train:1.814444e-03;0.998197;1.788319e-03; test:1.589159e-03;0.998285;1.703942e-03
# epoch:145; train:1.825136e-03;0.998188;1.799593e-03; test:1.580114e-03;0.998274;1.682870e-03
# epoch:146; train:1.826343e-03;0.998179;1.801926e-03; test:1.582201e-03;0.998280;1.701998e-03
# epoch:147; train:1.819998e-03;0.998194;1.792539e-03; test:1.614316e-03;0.998275;1.740828e-03
# epoch:148; train:1.824873e-03;0.998184;1.801563e-03; test:1.590007e-03;0.998258;1.707298e-03
# epoch:149; train:1.806517e-03;0.998200;1.783375e-03; test:1.567962e-03;0.998296;1.682338e-03
# epoch:150; train:1.803075e-03;0.998205;1.780478e-03; test:1.557305e-03;0.998298;1.669522e-03
# epoch:151; train:1.796908e-03;0.998200;1.776197e-03; test:1.561606e-03;0.998295;1.666160e-03
# epoch:152; train:1.814071e-03;0.998191;1.794864e-03; test:1.580231e-03;0.998275;1.692163e-03
# epoch:153; train:1.817135e-03;0.998181;1.794486e-03; test:1.573217e-03;0.998309;1.678252e-03
# epoch:154; train:1.797870e-03;0.998205;1.775585e-03; test:1.574518e-03;0.998282;1.683839e-03
# epoch:155; train:1.794358e-03;0.998200;1.773321e-03; test:1.571699e-03;0.998268;1.687480e-03
# epoch:156; train:1.790311e-03;0.998205;1.770916e-03; test:1.557074e-03;0.998321;1.672021e-03
# epoch:157; train:1.800432e-03;0.998186;1.783788e-03; test:1.552186e-03;0.998314;1.678471e-03
# epoch:158; train:1.793350e-03;0.998199;1.772712e-03; test:1.522993e-03;0.998340;1.649229e-03
# epoch:159; train:1.793456e-03;0.998195;1.781079e-03; test:1.543520e-03;0.998317;1.663595e-03
# epoch:160; train:1.781776e-03;0.998200;1.767049e-03; test:1.549995e-03;0.998320;1.663697e-03
# epoch:161; train:1.789008e-03;0.998209;1.770230e-03; test:1.542672e-03;0.998309;1.654156e-03
# epoch:162; train:1.789989e-03;0.998196;1.777983e-03; test:1.570431e-03;0.998293;1.691194e-03
# epoch:163; train:1.781384e-03;0.998203;1.766183e-03; test:1.538859e-03;0.998318;1.657582e-03
# epoch:164; train:1.781036e-03;0.998200;1.770582e-03; test:1.569136e-03;0.998285;1.696522e-03
# epoch:165; train:1.778529e-03;0.998203;1.771213e-03; test:1.549137e-03;0.998307;1.671316e-03
# epoch:166; train:1.773874e-03;0.998212;1.758400e-03; test:1.522223e-03;0.998329;1.641856e-03
# epoch:167; train:1.763386e-03;0.998219;1.749996e-03; test:1.544159e-03;0.998314;1.672284e-03
# epoch:168; train:1.787032e-03;0.998188;1.778918e-03; test:1.509532e-03;0.998347;1.631359e-03
# epoch:169; train:1.771837e-03;0.998206;1.763885e-03; test:1.523542e-03;0.998337;1.637520e-03
# epoch:170; train:1.767236e-03;0.998207;1.756558e-03; test:1.540684e-03;0.998303;1.667455e-03
# epoch:171; train:1.774860e-03;0.998201;1.764439e-03; test:1.534371e-03;0.998329;1.658521e-03
# epoch:172; train:1.766205e-03;0.998206;1.757632e-03; test:1.525448e-03;0.998333;1.650515e-03
# epoch:173; train:1.759938e-03;0.998205;1.756532e-03; test:1.514179e-03;0.998336;1.651046e-03
# epoch:174; train:1.763764e-03;0.998206;1.755375e-03; test:1.539898e-03;0.998294;1.661927e-03
# epoch:175; train:1.758914e-03;0.998213;1.754088e-03; test:1.514967e-03;0.998325;1.636344e-03
# epoch:176; train:1.754947e-03;0.998209;1.750761e-03; test:1.536947e-03;0.998301;1.660294e-03
# epoch:177; train:1.758797e-03;0.998209;1.753039e-03; test:1.522332e-03;0.998316;1.654774e-03
# epoch:178; train:1.748391e-03;0.998217;1.746467e-03; test:1.515855e-03;0.998317;1.632936e-03
# epoch:179; train:1.754528e-03;0.998216;1.747804e-03; test:1.530515e-03;0.998305;1.659779e-03
# epoch:180; train:1.754437e-03;0.998210;1.753479e-03; test:1.506400e-03;0.998335;1.636308e-03
# epoch:181; train:1.753712e-03;0.998211;1.750837e-03; test:1.500162e-03;0.998326;1.623050e-03
# epoch:182; train:1.746209e-03;0.998218;1.741456e-03; test:1.514605e-03;0.998313;1.652245e-03
# epoch:183; train:1.754528e-03;0.998205;1.752917e-03; test:1.537649e-03;0.998297;1.666600e-03
# epoch:184; train:1.747727e-03;0.998207;1.748563e-03; test:1.504216e-03;0.998327;1.636001e-03
# epoch:185; train:1.746592e-03;0.998218;1.744850e-03; test:1.501689e-03;0.998336;1.630552e-03
# epoch:186; train:1.747857e-03;0.998208;1.748528e-03; test:1.503047e-03;0.998320;1.627961e-03
# epoch:187; train:1.739748e-03;0.998218;1.738399e-03; test:1.488855e-03;0.998354;1.613676e-03
# epoch:188; train:1.742050e-03;0.998219;1.742508e-03; test:1.535039e-03;0.998290;1.670418e-03
# epoch:189; train:1.747617e-03;0.998206;1.752949e-03; test:1.510470e-03;0.998326;1.646046e-03
# epoch:190; train:1.748650e-03;0.998211;1.748507e-03; test:1.499574e-03;0.998325;1.632144e-03
# epoch:191; train:1.738402e-03;0.998214;1.744992e-03; test:1.514086e-03;0.998313;1.649120e-03
# epoch:192; train:1.729223e-03;0.998225;1.731975e-03; test:1.517066e-03;0.998315;1.646859e-03
# epoch:193; train:1.732172e-03;0.998217;1.738077e-03; test:1.487612e-03;0.998338;1.621335e-03
# epoch:194; train:1.730872e-03;0.998215;1.736637e-03; test:1.506796e-03;0.998338;1.641642e-03
# epoch:195; train:1.731216e-03;0.998217;1.737694e-03; test:1.507848e-03;0.998337;1.637851e-03
# epoch:196; train:1.738757e-03;0.998208;1.746222e-03; test:1.486031e-03;0.998343;1.614452e-03
# epoch:197; train:1.731191e-03;0.998214;1.737514e-03; test:1.518419e-03;0.998309;1.667245e-03
# epoch:198; train:1.726013e-03;0.998224;1.732500e-03; test:1.492273e-03;0.998320;1.626487e-03
# epoch:199; train:1.721657e-03;0.998221;1.732491e-03; test:1.477473e-03;0.998350;1.605542e-03
# epoch:200; train:1.726062e-03;0.998224;1.736643e-03; test:1.471630e-03;0.998368;1.594217e-03
# epoch:201; train:1.720155e-03;0.998217;1.733176e-03; test:1.500573e-03;0.998311;1.636563e-03
# epoch:202; train:1.720508e-03;0.998222;1.730439e-03; test:1.478474e-03;0.998353;1.611709e-03
# epoch:203; train:1.726319e-03;0.998214;1.737019e-03; test:1.488788e-03;0.998344;1.621275e-03
# epoch:204; train:1.720292e-03;0.998219;1.732199e-03; test:1.504794e-03;0.998317;1.642327e-03
# epoch:205; train:1.712551e-03;0.998234;1.719754e-03; test:1.496952e-03;0.998315;1.632780e-03
# epoch:206; train:1.708332e-03;0.998230;1.717224e-03; test:1.486048e-03;0.998347;1.619440e-03
# epoch:207; train:1.714243e-03;0.998227;1.724357e-03; test:1.479546e-03;0.998349;1.609096e-03
# epoch:208; train:1.708307e-03;0.998230;1.722784e-03; test:1.467099e-03;0.998358;1.609097e-03
# epoch:209; train:1.705388e-03;0.998234;1.716380e-03; test:1.472132e-03;0.998361;1.604432e-03
# epoch:210; train:1.705172e-03;0.998233;1.716448e-03; test:1.476092e-03;0.998344;1.614079e-03
# epoch:211; train:1.705084e-03;0.998232;1.718653e-03; test:1.480363e-03;0.998339;1.615023e-03
# epoch:212; train:1.695339e-03;0.998239;1.710009e-03; test:1.468863e-03;0.998367;1.592822e-03
# epoch:213; train:1.706923e-03;0.998230;1.716645e-03; test:1.460416e-03;0.998352;1.591525e-03
# epoch:214; train:1.708639e-03;0.998227;1.724184e-03; test:1.460314e-03;0.998360;1.590578e-03
# epoch:215; train:1.702107e-03;0.998230;1.717600e-03; test:1.484194e-03;0.998322;1.624894e-03
# epoch:216; train:1.687253e-03;0.998241;1.705793e-03; test:1.469905e-03;0.998351;1.608050e-03
# epoch:217; train:1.693541e-03;0.998235;1.709818e-03; test:1.448452e-03;0.998378;1.591549e-03
# epoch:218; train:1.701737e-03;0.998229;1.716765e-03; test:1.471089e-03;0.998345;1.605067e-03
# epoch:219; train:1.694545e-03;0.998237;1.711736e-03; test:1.462804e-03;0.998359;1.593139e-03
# epoch:220; train:1.706133e-03;0.998220;1.725733e-03; test:1.454398e-03;0.998359;1.587005e-03
# epoch:221; train:1.693068e-03;0.998241;1.711797e-03; test:1.483605e-03;0.998309;1.626177e-03
# epoch:222; train:1.692873e-03;0.998235;1.715485e-03; test:1.465294e-03;0.998345;1.598335e-03
# epoch:223; train:1.690965e-03;0.998232;1.712518e-03; test:1.467148e-03;0.998350;1.605152e-03
# epoch:224; train:1.695931e-03;0.998230;1.716884e-03; test:1.449940e-03;0.998352;1.590593e-03
# epoch:225; train:1.683786e-03;0.998244;1.702593e-03; test:1.417062e-03;0.998405;1.548839e-03
# epoch:226; train:1.696808e-03;0.998235;1.715830e-03; test:1.467455e-03;0.998353;1.607996e-03
# epoch:227; train:1.683992e-03;0.998242;1.705066e-03; test:1.450981e-03;0.998359;1.596111e-03
# epoch:228; train:1.690170e-03;0.998231;1.712123e-03; test:1.437502e-03;0.998375;1.581523e-03
# epoch:229; train:1.681747e-03;0.998243;1.702019e-03; test:1.459515e-03;0.998333;1.598052e-03
# epoch:230; train:1.679177e-03;0.998240;1.703009e-03; test:1.439953e-03;0.998371;1.566308e-03
# epoch:231; train:1.677801e-03;0.998242;1.699523e-03; test:1.459169e-03;0.998334;1.607019e-03
# epoch:232; train:1.682527e-03;0.998242;1.706159e-03; test:1.458047e-03;0.998336;1.596785e-03
# epoch:233; train:1.688471e-03;0.998233;1.714950e-03; test:1.436940e-03;0.998380;1.570862e-03
# epoch:234; train:1.681078e-03;0.998242;1.705019e-03; test:1.451352e-03;0.998361;1.590894e-03
# epoch:235; train:1.685255e-03;0.998236;1.710118e-03; test:1.435641e-03;0.998373;1.575659e-03
# epoch:236; train:1.680818e-03;0.998233;1.708146e-03; test:1.438214e-03;0.998363;1.582408e-03
# epoch:237; train:1.684345e-03;0.998229;1.711761e-03; test:1.441066e-03;0.998367;1.587458e-03
# epoch:238; train:1.675588e-03;0.998245;1.700659e-03; test:1.424159e-03;0.998386;1.564341e-03
# epoch:239; train:1.678522e-03;0.998243;1.704180e-03; test:1.429886e-03;0.998369;1.572588e-03
# epoch:240; train:1.676034e-03;0.998241;1.701212e-03; test:1.447198e-03;0.998365;1.590816e-03
# epoch:241; train:1.663700e-03;0.998253;1.691943e-03; test:1.409263e-03;0.998402;1.536137e-03
# epoch:242; train:1.669256e-03;0.998252;1.693386e-03; test:1.437979e-03;0.998369;1.594268e-03
# epoch:243; train:1.670016e-03;0.998245;1.698580e-03; test:1.429972e-03;0.998374;1.563333e-03
# epoch:244; train:1.675653e-03;0.998239;1.705092e-03; test:1.441554e-03;0.998365;1.587856e-03
# epoch:245; train:1.658718e-03;0.998256;1.685259e-03; test:1.426848e-03;0.998382;1.570084e-03
# epoch:246; train:1.668271e-03;0.998242;1.697738e-03; test:1.458589e-03;0.998336;1.610620e-03
# epoch:247; train:1.669507e-03;0.998247;1.699628e-03; test:1.428034e-03;0.998362;1.574192e-03
# epoch:248; train:1.669105e-03;0.998246;1.701488e-03; test:1.454480e-03;0.998341;1.599737e-03
# epoch:249; train:1.668724e-03;0.998248;1.698968e-03; test:1.413901e-03;0.998392;1.554457e-03
# epoch:250; train:1.658678e-03;0.998257;1.687435e-03; test:1.446051e-03;0.998354;1.594351e-03
# epoch:251; train:1.650839e-03;0.998259;1.682951e-03; test:1.438184e-03;0.998370;1.573033e-03
# epoch:252; train:1.662553e-03;0.998250;1.694015e-03; test:1.425274e-03;0.998358;1.573262e-03
# epoch:253; train:1.661471e-03;0.998249;1.691583e-03; test:1.421734e-03;0.998366;1.574520e-03
# epoch:254; train:1.665235e-03;0.998242;1.698188e-03; test:1.433295e-03;0.998368;1.580100e-03
# epoch:255; train:1.654626e-03;0.998260;1.682780e-03; test:1.415968e-03;0.998393;1.556242e-03
# epoch:256; train:1.650863e-03;0.998256;1.683942e-03; test:1.415063e-03;0.998395;1.554382e-03
# epoch:257; train:1.669139e-03;0.998239;1.705967e-03; test:1.447707e-03;0.998355;1.592035e-03
# epoch:258; train:1.664779e-03;0.998245;1.695097e-03; test:1.417150e-03;0.998390;1.557628e-03
# epoch:259; train:1.648794e-03;0.998257;1.679854e-03; test:1.454079e-03;0.998357;1.598937e-03
# epoch:260; train:1.646327e-03;0.998255;1.683553e-03; test:1.433468e-03;0.998359;1.592634e-03
# epoch:261; train:1.655534e-03;0.998253;1.687955e-03; test:1.432870e-03;0.998372;1.580501e-03
# epoch:262; train:1.659547e-03;0.998246;1.695548e-03; test:1.424367e-03;0.998355;1.575158e-03
# epoch:263; train:1.653136e-03;0.998258;1.685300e-03; test:1.445048e-03;0.998358;1.600755e-03
# epoch:264; train:1.647382e-03;0.998256;1.680752e-03; test:1.417185e-03;0.998378;1.565166e-03
# epoch:265; train:1.652929e-03;0.998250;1.689719e-03; test:1.410184e-03;0.998390;1.557093e-03
# epoch:266; train:1.648030e-03;0.998262;1.680720e-03; test:1.416665e-03;0.998379;1.566419e-03
# epoch:267; train:1.637248e-03;0.998263;1.673516e-03; test:1.427590e-03;0.998375;1.579131e-03
# epoch:268; train:1.643664e-03;0.998260;1.680615e-03; test:1.425493e-03;0.998361;1.572676e-03
# epoch:269; train:1.654446e-03;0.998252;1.688323e-03; test:1.425478e-03;0.998369;1.571201e-03
# epoch:270; train:1.652106e-03;0.998250;1.687494e-03; test:1.418610e-03;0.998376;1.567556e-03
# epoch:271; train:1.648791e-03;0.998253;1.687389e-03; test:1.424771e-03;0.998364;1.576824e-03
# epoch:272; train:1.641105e-03;0.998259;1.680395e-03; test:1.375966e-03;0.998405;1.523552e-03
# epoch:273; train:1.634332e-03;0.998267;1.671937e-03; test:1.449562e-03;0.998348;1.587081e-03
# epoch:274; train:1.640454e-03;0.998261;1.676861e-03; test:1.395459e-03;0.998389;1.547720e-03
# epoch:275; train:1.641510e-03;0.998256;1.681663e-03; test:1.407487e-03;0.998395;1.552096e-03
# epoch:276; train:1.640667e-03;0.998258;1.682638e-03; test:1.428221e-03;0.998342;1.587362e-03
# epoch:277; train:1.636366e-03;0.998266;1.672219e-03; test:1.416288e-03;0.998391;1.561464e-03
# epoch:278; train:1.641294e-03;0.998260;1.681747e-03; test:1.404862e-03;0.998382;1.550669e-03
# epoch:279; train:1.641776e-03;0.998255;1.680817e-03; test:1.375495e-03;0.998415;1.518481e-03
# epoch:280; train:1.638963e-03;0.998257;1.680382e-03; test:1.410448e-03;0.998375;1.559388e-03
# epoch:281; train:1.641177e-03;0.998253;1.682830e-03; test:1.419746e-03;0.998354;1.574072e-03
# epoch:282; train:1.638149e-03;0.998262;1.677556e-03; test:1.391718e-03;0.998406;1.539103e-03
# epoch:283; train:1.641591e-03;0.998254;1.682933e-03; test:1.402884e-03;0.998368;1.556622e-03
# epoch:284; train:1.630491e-03;0.998265;1.673363e-03; test:1.398663e-03;0.998375;1.550214e-03
# epoch:285; train:1.634652e-03;0.998259;1.676092e-03; test:1.381821e-03;0.998395;1.531392e-03
# epoch:286; train:1.633694e-03;0.998261;1.679053e-03; test:1.409044e-03;0.998364;1.558868e-03
# epoch:287; train:1.643351e-03;0.998250;1.684679e-03; test:1.396459e-03;0.998390;1.549610e-03
# epoch:288; train:1.629854e-03;0.998259;1.673962e-03; test:1.396006e-03;0.998385;1.543797e-03
# epoch:289; train:1.626640e-03;0.998265;1.670278e-03; test:1.399200e-03;0.998381;1.556042e-03
# epoch:290; train:1.634723e-03;0.998259;1.681324e-03; test:1.397704e-03;0.998368;1.553187e-03
# epoch:291; train:1.634319e-03;0.998257;1.681309e-03; test:1.384950e-03;0.998406;1.528719e-03
# epoch:292; train:1.624222e-03;0.998266;1.670621e-03; test:1.396271e-03;0.998381;1.547505e-03
# epoch:293; train:1.625333e-03;0.998264;1.671422e-03; test:1.393696e-03;0.998392;1.545345e-03
# epoch:294; train:1.631527e-03;0.998259;1.675935e-03; test:1.371522e-03;0.998395;1.529297e-03
# epoch:295; train:1.626665e-03;0.998265;1.670865e-03; test:1.359643e-03;0.998404;1.505784e-03
# epoch:296; train:1.616556e-03;0.998268;1.663102e-03; test:1.388109e-03;0.998389;1.539631e-03
# epoch:297; train:1.622301e-03;0.998266;1.667402e-03; test:1.391051e-03;0.998385;1.546056e-03
# epoch:298; train:1.614000e-03;0.998275;1.659016e-03; test:1.378988e-03;0.998395;1.530024e-03
# epoch:299; train:1.610867e-03;0.998273;1.658164e-03; test:1.394352e-03;0.998382;1.547077e-03
# epoch:300; train:1.626397e-03;0.998259;1.676995e-03; test:1.371228e-03;0.998397;1.527078e-03
# -
# ## Test
# ### Prepare the test data
test_image, test_label = test_dataset[0]
test_image2, test_label2 = train_transform(test_image, test_label)
test_image2 = nd.expand_dims(test_image2,0)
print('tensor shape:', test_image2.shape)
# ### Network inference
anchors, cls_preds, box_preds = net(test_image2.as_in_context(ctx))
# ### Convert predictions to real object detection results
from mxnet.contrib.ndarray import MultiBoxDetection
cls_probs = nd.SoftmaxActivation(nd.transpose(cls_preds, (0, 2, 1)), mode='channel')
output = MultiBoxDetection(cls_prob=cls_probs, loc_pred=box_preds, anchor=anchors, force_suppress=True, clip=True, nms_topk=250)
# ### Display results
# +
class_names = ('cluster')
def display(img, out, thresh=0.5):
import random
import matplotlib as mpl
import numpy as np
mpl.rcParams['figure.figsize'] = (10,10)
img = img.asnumpy()
img = np.transpose(img,(2,3,1,0))
img = np.squeeze(img)
plt.clf()
plt.imshow(img)
for det in out:
cid = int(det[0])
if cid == 0:
continue
score = det[1]
if score < thresh:
continue
scales = [img.shape[1], img.shape[0]] * 2
xmin, ymin, xmax, ymax = [int(p * s) for p, s in zip(det[2:6].tolist(), scales)]
rect = plt.Rectangle((xmin, ymin), xmax - xmin, ymax - ymin, fill=False,
edgecolor='red', linewidth=3)
plt.gca().add_patch(rect)
text = class_names[cid]
plt.gca().text(xmin, ymin-2, '{:s} {:.3f}'.format(text, score),
bbox=dict(facecolor='red', alpha=0.5),
fontsize=12, color='white')
display(test_image2, output[0].asnumpy(), thresh=0.52)
# -
|
experiments/ssd512-mobilenet_v1_alpha4_from_distilled.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] deletable=true editable=true
# ## 01 - 00 Introduction to Python
# We mentioned the pros of using Python as our tool for performing urban science in the Introduction to Notebook and Beyond. In this introductory module we will discuss the basic semantics of the language and get our hands dirty by actually typing and executing the code blocks.
# > ##### Remeber:
# > - To run a cell, you can press `Shift+Enter` (to execute and go to next cell) or `Ctrl+Enter` (to execute and stay on the same cell).
# > - To add an empty cell, press `Esc` to get out of edit mode and press either `b` key (to add it after the current cell) or `a` key (to add it before the current cell).
# > - You can make edits to the code and play around with it. However you cannot save the edits made to this notebook if you are running it on CDF.
#
# -
# Python has following 6 built-in Data-Types:
#
# | Type | Description | Example |
# |----------|-----------------------|---------|
# | int | Integer values | 123 |
# | float | Floating point values | 10.12 |
# | complex | Complex values | 1 + 3j |
# | bool | Boolean values | True |
# | str | String values | "Hello" |
# | NoneType | None value | None |
# and 4 Data-Structures
#
# | Type | Description | Example |
# |-------|---------------------------------------|----------------------|
# | list | Ordered collection of values | [1, 'abc', 3, 1] |
# | set | Unordered collection of unique values | {1, 'abc', 3} |
# | tuple | Immutable Ordered collection | (1, 'abc', 3) |
# | dict | Unordered key. value pairs | {'abc': 1, 'def': 2} |
# In this section we will learn about the above Python's built-in Data-Types and Data-Structures.
|
notebooks/01-00 Introduction to Python.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: LSST
# language: python
# name: lsst
# ---
# # Exploring a Data Repository
#
# <br>Owner: **<NAME>** ([@rmorgan10](https://github.com/LSSTScienceCollaborations/StackClub/issues/new?body=@rmorgan10)), **<NAME>** ([@drphilmarshall](https://github.com/LSSTScienceCollaborations/StackClub/issues/new?body=@drphilmarshall)), **<NAME>** ([@kadrlica](https://github.com/LSSTScienceCollaborations/StackClub/issues/new?body=@kadrlica))
# <br>Some minor updates by: <NAME> ([@douglasleetucker](https://github.com/LSSTScienceCollaborations/StackClub/issues/new?body=@douglasleetucker))
# <br>Last Verified to Run: **2021-03-12**
# <br>Verified Stack Release: **v21.0.0**
#
# This notebook examines the content of a data repository -- in this particular case, an HSC data repository -- and shows how to determine the inputs for each component. A related notebook is "Exploring a DC2 Data Repository," which explores the same basic concepts, but using a DC2 data repository; this other notebook covers matters in a slightly differ manner, though, and it is useful to explore both notebooks for a fuller understanding of data repositories.
#
# ### Learning Objectives:
# After working through and studying this notebook you should be able to understand how to use the Butler to figure out:
# 1. What a data repo is;
# 2. Which data types are present in a data repository;
# 3. If coadds have been made, what the available tracts are;
# 4. Which parts of the sky those tracts cover.
#
# ### Logistics
# This notebook is intended to be runnable on `lsst-lsp-stable.ncsa.illinois.edu` from a local git clone of https://github.com/LSSTScienceCollaborations/StackClub.
#
#
# ## Set Up
import os
import sys
import subprocess as sub
import warnings
import matplotlib.pyplot as plt
from IPython.display import display, Markdown
import numpy as np
import os, glob
# %matplotlib inline
# ## What is a Data Repo?
#
# A data repo is a directory containing raw images, calibration files, metadata and configuration information, defining an LSST-format dataset. Data repositories contain either a `_mapper` file or a `repositoryCfg.yaml` file, which record the "obs package" that was used to organize the data. The obs package gives the repository more structure and organization than an ordinary data directory. Let's take a look at this file structure in the HSC data repo.
# ### The HSC Data Repo: What's in there?
# We'll use the `hsc` data repository as our testing ground, and start by figuring out what it contains. In the `hsc` case, the `_mapper` file is in the top level folder, while the data repo for each field is a few levels down.
repo = '/datasets/hsc/repo'
# ! ls /datasets/hsc/repo/
# We can see the `_mapper` file here, and at contains one line giving the name of the `Mapper` object for the HSC repo:
# +
# ! cat /datasets/hsc/repo/_mapper
# Import the Mapper object once you know its name
from lsst.obs.hsc import HscMapper
# -
# You can get some more information on this object like this:
# +
#help(HscMapper)
# -
# The mapper defines a (large) number of different "dataset types". Some of these are specific to this particular data repo, others are more general. Even filtering out some intermediate dataset types, we are still left with a long list. But, once we figure out which dataset types we are interested in, we can start querying for information about those datasets.
# +
mapper = HscMapper(root=repo)
all_dataset_types = mapper.getDatasetTypes()
remove = ['_config', '_filename', '_md', '_sub', '_len', '_schema', '_metadata']
shortlist = []
for dataset_type in all_dataset_types:
keep = True
for word in remove:
if word in dataset_type:
keep = False
if keep:
shortlist.append(dataset_type)
print(shortlist)
# -
# The `Butler`, directed by the `Mapper`, will have access to all the above dataset types.
#
# Another important file in the repo parent folder is `registry.sqlite3`. This database contains metadata for the HSC **raw** images.
# ### Great, but where is the actual data, and how was it processed?
# The raw visit images are stored by field. In the HSC dataset the fields have names like `COSMOS` and `DEEPE09`. Within those field folders, there is a directory structure that eventually gets down to visit image FITS files whose names and paths contain the date/time and filter for that exposure. For example:
# ! \ls /datasets/hsc/repo/COSMOS/2015-01-18/01113/HSC-Y/HSC-0018476-00?.fits
# How do the pipeline tasks know which raw data to process? This information is captured in the "configs". In the HSC repo there is no config folder or files in the top level directory - in fact the only two files are `_mapper` and `registry.sqlite3`. So what's going on?
#
# It turns out that the provenance of the stack processing of the HSC raw images is captured in "rerun" folders, one for each time the science pipelines were run on the data. Let's do some detective work to find out what happened to the HSC data.
# ! ls /datasets/hsc/repo/rerun
# First, `DM-10404` looks like a run ID. What's in that folder?
# ! ls /datasets/hsc/repo/rerun/DM-10404
# `DEEP`, `WIDE`, and `UDEEP` are the names of the sub-surveys of the HSC survey. We might expect each to contain results from the processing of that sub-survey's images.
# ! ls /datasets/hsc/repo/rerun/DM-10404/UDEEP
# The numerically-named folders contain the generated catalog files, organized by sky tract. Note that a `config` folder is present, and also a `repositoryCfg.yaml` file - which means that this folder is itself a `repo`, from the Butler's point of view.
# ! cat /datasets/hsc/repo/rerun/DM-10404/WIDE/repositoryCfg.yaml
# What does it mean that `SFM` is the "parent" of this repo? Let's see what _that_ folder contains:
# ! ls /datasets/hsc/repo/rerun/DM-10404/SFM
# `SFM` seems to contain _all_ the tracts that have been produced - so is the entire HSC survey. It's `repositoryCfg.yaml` file shows that it's "parent" is the top level folder, `/datasets/hsc/repo/`.
# ! cat /datasets/hsc/repo/rerun/DM-10404/SFM/repositoryCfg.yaml
# Let's see what the `DM-10404/UDEEP` repo's `config` folder contains:
# ! ls -a /datasets/hsc/repo/rerun/DM-10404/UDEEP/config
# These are the configuration files that were used when the science pipelines were run on these data. While we don't know which versions of the software were used, we at least know which tasks were run.
#
# Here's what a config file looks like (ignoring the many import statements and just looking at a few example lines):
# ! cat /datasets/hsc/repo/rerun/DM-10404/UDEEP/config/forcedPhotCcd.py | grep -v import | head -15
# The next level to dig into here is the tract folders within one of the repos in this rerun. For example:
# ! ls /datasets/hsc/repo/rerun/DM-10404/UDEEP/00814/HSC-Y/tract9570 | head -10
# Those FITS files contain the forced source tables.
# ## Instantiating the Butler and looking for Dataset Types
#
# Now that we have an idea of the structure of the repo itself, let's use the Butler to explore the data within the repo. Here we will demonstrate a few useful `Butler` methods for learning about the data in a repo. Let's choose one of the rerun repos, and investigate its properties. We'll summon two butlers, one that is pointed at the parent repo, and another (an "under butler") that is asked to focus on a particular sub-survey in a particular re-run.
# +
parent_repo = '/datasets/hsc/repo'
# Choose a re-run repo:
rerun_id = 'DM-10404'
depth = 'UDEEP'
# Try a different one:
# rerun = 'DM-13666'
# depth = 'WIDE'
repo = parent_repo + '/rerun/' + rerun_id + '/' + depth
print(repo)
from lsst.daf.persistence import Butler
butler = Butler(parent_repo)
under_butler = Butler(repo)
# -
# The `butler` can check whether a datatype (like the source catalogs) actually exists or not, but it needs a specific dataset ID to check whether that specific part of the dataset exists.
#
# Note that the metadata being queried here is in the `registry.sqlite3` database in the _parent_ repo - and so refers to the _initial_ processing run, not the most recent rerun. We'll need to work carefully around this below.
# ## Obtaining Basic Dataset Properties Using the Butler
# Now we can start using Butler methods to query metadata for the repo. For this dataset, we can look at the filters used, number of visits, number of pointings, etc. by examining the Butler's keys and metadata. For these basic properties, we will look at the `calexp` and `src` tables. The contents of these tables are derived from the processing of individual sensors, and exist in the parent folder. (That means that we can use either of our two butlers to query for them.)
#
# Note that the metadata is created from the raw exposures loaded into the sqlite registry. The fact that we can get metadata for a specific datasetType and dataId **does not** imply that the data exist on disk (we will check this in a subsequent step).
# +
# This would be faster if only one query were issued...
visits = butler.queryMetadata('calexp', ['visit'])
pointings = butler.queryMetadata('calexp', ['pointing'])
ccds = butler.queryMetadata('calexp', ['ccd'])
fields = butler.queryMetadata('calexp', ['field'])
filters = butler.queryMetadata('calexp', ['filter'])
sources = butler.queryMetadata('src', ['id'])
# It is possible to specify multiple formats -- i.e., butler.queryMetadata('calexp', ['visit','ccd'])
metadata = butler.queryMetadata('calexp', ['visit','pointing','ccd','field','filter'])
# -
num_visits = len(visits)
num_pointings = len(pointings)
num_ccds = len(ccds)
num_fields = len(fields)
num_filters = len(filters)
num_sources = len(sources)
num_metadata = len(metadata)
print("The HSC {}/{} rerun contains {} visits.".format(rerun_id,depth,num_visits))
print("The HSC {}/{} rerun contains {} pointings.".format(rerun_id,depth,num_pointings))
print("The HSC {}/{} rerun contains {} ccds.".format(rerun_id,depth,num_ccds))
print("The HSC {}/{} rerun contains {} fields.".format(rerun_id,depth,num_fields))
print("The HSC {}/{} rerun contains {} filters.".format(rerun_id,depth,num_filters))
print("The HSC {}/{} rerun contains {} sources.".format(rerun_id,depth,num_sources))
# print("The HSC {}/{} rerun contains {} coadd sources.".format(rerun_id,depth,num_coadd_sources))
# print("The HSC {}/{} rerun contains {} forced sources.".format(rerun_id,depth,num_forced_sources))
# As a check, let's compare what our two butlers find when asked for the number of sources:
# +
alt_num_sources = len(under_butler.queryMetadata('src', ['id']))
print("The butler says that we have {:d} input sources.".format(num_sources))
print("The under butler says we have {:d} input sources.".format(alt_num_sources))
# -
# So, we could have done our query with the `under_butler` as well. In practice, it's best to specify a Butler for the rerun repo, because that Butler will also have access to the parent repo.
#
#
# ### So where are the Calexps and Source Catalogs?
#
# Notice that while we were able to get metadata for the `processCcd` outputs (the `calexp` and `src`), that **does not** guarantee that these products are on disk. The metadata is created from the raw inputs stored in the registry and a template for the derived data products. In this case, we have a reason to be suspicious: remember that when we examined the directory structure we did not see a directory for the `processCcd` products.
#
# To check the existence of the data requires the use of the `datasetExists` method of the `butler`. Let's give this a try.
# First we select the metadata for a specific calexp
dataId={'filter':'HSC-I','visit':872,'ccd':10}
butler.queryMetadata('calexp', ['visit','ccd','filter','field','pointing'], dataId=dataId)
# Ok, so we have the metadata, let's try to get the calexp...
# +
try:
calexp = butler.get('calexp',dataId=dataId)
except Exception as e:
print(str(e))
print("\nWhat? Does the data exist?")
# Explicitly check for existence
exists = butler.datasetExists('calexp',dataId=dataId)
print("\nbutler.datasetExists: " + str(exists))
# -
# So it appears that we don't have these intermediate products stored in this repo.
# ## Coadd Sky Area
#
# One may also be interested in the total sky area imaged for a particular coadd rerun/depth. We can estimate and visualize this from the coadd tract info that neither our `under_butler` nor our `butler` has access to. To collect all the tracts, we have to get them via the file structure. This operation will hopefully be `Butler`-ized with the Gen3 Butler.
# +
# Collect tract indices from file names
tracts = sorted([int(os.path.basename(x)) for x in
glob.glob(os.path.join(repo, 'deepCoadd-results', 'merged', '*'))])
num_tracts = len(tracts)
print("Found {} merged tracts in repo {}".format(num_tracts, repo))
# -
# A quick way of extimating the sky area covered is to sum the areas of the inner boxes of all the tracts. For more information on the properties of tracts, you can look at the [Documentation](http://doxygen.lsst.codes/stack/doxygen/x_masterDoxyDoc/classlsst_1_1skymap_1_1tract_info_1_1_tract_info.html).
# As a quick note, the file structure only tells us the names of the tracts in the particular rerun/depth to look at. The actual `TractInfo` objects are obtained by selecting the tracts we want from the `deepCoadd_skyMap` dataset in our particular rerun repo. Therefore, we will have to ask the `under_butler` to bring us this dataset for the particular rerun/depth.
# +
# Calculate area from all tracts
skyMap = under_butler.get('deepCoadd_skyMap')
total_area = 0.0 #deg^2
plotting_vertices = []
for test_tract in tracts:
# Get inner vertices for tract
tractInfo = skyMap[test_tract]
vertices = tractInfo._vertexCoordList
plotting_vertices.append(vertices)
#calculate area of box
av_dec = 0.5 * (vertices[2][1] + vertices[0][1])
av_dec = av_dec.asRadians()
delta_ra_raw = vertices[0][0] - vertices[1][0]
delta_ra = delta_ra_raw.asDegrees() * np.cos(av_dec)
delta_dec= vertices[2][1] - vertices[0][1]
area = delta_ra * delta_dec.asDegrees()
#combine areas
total_area += area
# Round off the total area for presentation purposes
rounded_total_area = round(total_area, 2)
print("Total area imaged (sq deg): ",rounded_total_area)
# -
# ## Displaying Dataset Characteristics
# Now let's print out a report of all the characteristcs we have found. We'll use the sky area from the rerun we chose, and the numbers common to all reruns.
# +
dataset_name = 'HSC'
display(Markdown('### %s' % repo))
# Make a table of the collected metadata
collected_data = [num_visits, num_pointings, num_ccds, num_fields, num_filters, num_sources,
num_tracts, rounded_total_area]
data_names = ("Number of Visits", "Number of Pointings", "Number of CCDs", "Number of Fields",
"Number of Filters", "Number of Sources", "Number of Tracts", "Total Sky Area (deg$^2$)")
# TODO: include coadd sources and forced sources
output_table = "| Metadata Characteristic | Value | \n | ---: | ---: | \n "
counter = 0
while counter < len(collected_data):
output_table += "| %s | %s | \n" %(data_names[counter], collected_data[counter])
counter += 1
display(Markdown(output_table))
# Show which fields and filters we're talking about:
display(Markdown('Fields: (%i total)' %num_fields))
print(fields)
display(Markdown('Filters: (%i total)' %num_filters))
print(filters)
# -
# ## Plotting the sky coverage
#
# For this we will need our list of merged `tracts` from above, and also the `skyMap` object. We can then extract the sky coordinates of the corners of each tract, and use them to draw a set of rectangles to illustrate the sky coverage, following <NAME>'s LSST DESC tutorial [dm_butler_skymap.ipynb](https://github.com/LSSTDESC/DC2-analysis/blob/master/tutorials/dm_butler_skymap.ipynb).
#
# In the future, we could imagine overlaying the focal plane and color the individual visits, using more of the code from Jim's notebook. Let's wait to see what functionality the Gen3 Butler provides first, and maybe return to visualization later.
# +
plt.figure()
for tract in tracts:
tractInfo = skyMap[tract]
corners = [(x[0].asDegrees(), x[1].asDegrees()) for x in tractInfo.getVertexList()]
x = [k[0] for k in corners] + [corners[0][0]]
y = [k[1] for k in corners] + [corners[0][1]]
plt.plot(x,y, color='b')
plt.xlabel('RA (deg)')
plt.ylabel('Dec (deg)')
plt.title('2D Projection of Sky Coverage')
plt.show()
# -
# We could imagine plotting the patches as well, to show which tracts were incomplete - but this gives us a rough idea of where our data is on the sky.
# ## Finding a Coadd Object Catalog
#
# Since we know we have `deepCoadd` images, we might want to find the sources detected in those images. These are the precursors of `Objects`. If forced photometry has been run on these sources, there should be `deepCoadd_forced_src` catalogs present. The `config` tells us which tasks have been run in the rerun, but metadata describing the results of the run is not propagated back into the registry database in the parent folder. So, we need to get data from the `under_butler`.
#
# Since the registry only knows about the raw exposures (and products that can be directly derived from templates using the raws), we can't get valid metadata from `queryMetadata` directly. Instead, we query the directory structure for a valid filter, tract, patch combination.
# +
dirname=os.path.join(repo,'deepCoadd-results')
print('filters: '+dirname)
# !ls $dirname
filter='HSC-I'
dirname = os.path.join(dirname, filter)
print('\ntracts: '+dirname)
# !ls $dirname
tract=8523
dirname = os.path.join(dirname,str(tract))
print('\npatches: '+dirname)
# !ls $dirname
patch='1,3'
dirname = os.path.join(dirname,patch)
print('\ndata products: '+dirname)
# !ls $dirname
# -
# We can also get some available tract, patches from the `skyMap` object. We need to feed it a list of coordinates defining a polygon to search for viable tract,patch info.
from lsst.geom import SpherePoint, Angle
# Create a list of corners of the polygon to search.
coordList = [SpherePoint(Angle(np.radians(0)),Angle(np.radians(-6))),
SpherePoint(Angle(np.radians(0)),Angle(np.radians(-2))),
SpherePoint(Angle(np.radians(40)),Angle(np.radians(-2))),
SpherePoint(Angle(np.radians(40)),Angle(np.radians(-6)))
]
# Print the tract,patch info
tractInfo = skyMap.findTractPatchList(coordList)
for _tract in tractInfo:
print(_tract[0])
for _patch in _tract[1]:
print(' ',_patch)
# Here we grab the coadd source catalog for a specific filter
dataId={'filter':'HSC-I','tract':tract,'patch':patch}
coadd_sources = under_butler.get('deepCoadd_forced_src',dataId=dataId)
coadd_sources.asAstropy()
# We can also get the merged detections
dataId={'tract':8523,'patch':'1,8'}
merged_sources = under_butler.get('deepCoadd_mergeDet',dataId=dataId)
# +
#One can get the available keys by running :
#coadd_sources.getSchema()
# -
# ## Visualizing Simple Measurements
#
# We'd like to know something about the sources that have been detected and measured, beyond a simple total number. We create a simple histogram of the Kron flux of sources in our HSC-I band tile. You can find more details on merging catalogs across bands, etc on the [Science Pipelines documentation](https://pipelines.lsst.io/getting-started/multiband-analysis.html).
plt.figure()
plt.hist(coadd_sources['ext_photometryKron_KronFlux_flux'],bins=np.linspace(0,100,50))
plt.xlabel("log10(Kron Flux)")
plt.title("{filter} {tract} {patch}".format(filter=filter,tract=tract,patch=patch))
# # Summary
#
# We have shown a few techniques for exploring a data repo. ~~To make this process straightforward, we are implementing all these techniques into methods of a `Taster` class, which is now a part of the `stackclub` library. The `Taster` will give you a taste of what the `Butler` delivers. we demonstrate the use of this class in the [DataInventory.ipynb](https://github.com/LSSTScienceCollaborations/StackClub/blob/dc2_gen2/Graveyard/DataInventory.ipynb) notebook.~~
#
# (Note: the `Taster` class and the [DataInventory.ipynb](https://github.com/LSSTScienceCollaborations/StackClub/blob/dc2_gen2/Graveyard/DataInventory.ipynb) notebook are now both deprecated. The DataInventory notebook is largely a demonstration of the `Taster` class, a useful tool that was created to augment the deficiencies of the Gen-2 butler for investigating and exploring data repos. Unfortunately, it depends on classes and repo structures that have evolved substantially over the past few years. Since the `Taster` resides outside of the DM Stack, it requires constant maintenance to be kept up-to-date with the Stack. With the impending release of the Gen-3 butler, the `Taster` and the DataInventory notebook have been deprecated.)
|
Graveyard/Exploring_An_HSC_Data_Repo.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# clear all variables
for i in list(globals().keys()):
if(i[0] != '_'):
exec('del {}'.format(i))
#suppress future warnings -- not really a good idea
from warnings import simplefilter
# ignore all future warnings
simplefilter(action='ignore', category=FutureWarning)
# suppress future warnings -- not really a good idea
import warnings
warnings.simplefilter(action='ignore', category=FutureWarning)
# ignore all warnings
import warnings
warnings.filterwarnings('ignore')
import os
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import xlrd
import csv
import seaborn as sns
from IPython.display import set_matplotlib_formats
set_matplotlib_formats('png', 'pdf') # uses vector figures in pdf exports
plt.style.use('seaborn-pastel')
# create a folder
def create_folder(folder):
import os
try:
os.mkdir(folder)
except FileExistsError:
print("Directory [ %s ] already exists"%folder)
#create folder to store results
create_folder('Figures')
def ageCat(row):
if row["age"] < 75:
return "-75"
elif row["age"] > 74 and row["age"] < 80:
return "75-79"
elif row["age"] > 79 and row["age"] < 85:
return "80-84"
elif row["age"] > 84 and row["age"] < 90:
return "85-89"
elif row["age"] > 89 and row["age"] < 95:
return "90-94"
elif row["age"] > 94 and row["age"] < 100:
return "95-99"
elif row["age"] > 99:
return "+100"
def brainwgt_Cat(row):
if row["brain weight"] >= 0 and row["brain weight"] < 270:
return "0-259"
elif row["brain weight"] > 259 and row["brain weight"] < 540:
return "270-539"
elif row["brain weight"] > 539 and row["brain weight"] < 810:
return "540-809"
elif row["brain weight"] > 809 and row["brain weight"] < 1080:
return "810-1079"
elif row["brain weight"] > 1079 and row["brain weight"] < 1350:
return "1080-1349"
elif row["brain weight"] > 1349 and row["brain weight"] < 1621:
return "1350-1620"
def CAATotalSev_Cat(row):
if row["CAATotalSev"] >= 0 and row["CAATotalSev"] < 5:
return "0-4"
elif row["CAATotalSev"] > 4 and row["CAATotalSev"] < 10:
return "5-9"
elif row["CAATotalSev"] > 9 and row["CAATotalSev"] < 15:
return "10-14"
elif row["CAATotalSev"] > 14 and row["CAATotalSev"] < 20:
return "15-19"
elif row["CAATotalSev"] > 19 and row["CAATotalSev"] < 25:
return "20-24"
def CAAAreas_Cat(row):
if row["CAAParenc"] < 1:
return "0"
elif row["CAAParenc"] > 0 and row["CAAParenc"] < 3:
return "1-2"
elif row["CAAParenc"] > 2 and row["CAAParenc"] < 5:
return "3-4"
elif row["CAAParenc"] > 4 and row["CAAParenc"] < 7:
return "5-6"
elif row["CAAParenc"] > 6 and row["CAAParenc"] < 9:
return "7-8"
elif row["CAAParenc"] > 8 and row["CAAParenc"] < 11:
return "9-10"
def CAAParenc_Cat(row):
if row["CAAParenc"] < 1:
return "0"
elif row["CAAParenc"] > 0 and row["CAAParenc"] < 3:
return "1-2"
elif row["CAAParenc"] > 2 and row["CAAParenc"] < 5:
return "3-4"
elif row["CAAParenc"] > 4 and row["CAAParenc"] < 7:
return "5-6"
elif row["CAAParenc"] > 6 and row["CAAParenc"] < 9:
return "7-8"
elif row["CAAParenc"] > 8 and row["CAAParenc"] < 11:
return "9-10"
elif row["CAAParenc"] > 10 and row["CAAParenc"] < 13:
return "11-12"
def CAAMeningeal_Cat(row):
if row["CAAMeningeal"] < 1:
return "0"
elif row["CAAMeningeal"] > 0 and row["CAAMeningeal"] < 3:
return "1-2"
elif row["CAAMeningeal"] > 2 and row["CAAMeningeal"] < 5:
return "3-4"
elif row["CAAMeningeal"] > 4 and row["CAAMeningeal"] < 7:
return "5-6"
elif row["CAAMeningeal"] > 6 and row["CAAMeningeal"] < 9:
return "7-8"
elif row["CAAMeningeal"] > 8 and row["CAAMeningeal"] < 11:
return "9-10"
elif row["CAAMeningeal"] > 10 and row["CAAMeningeal"] < 13:
return "11-12"
def frequencyChart(df,cat1,cat2):
bar = df[[cat1,cat2]]
bar = bar.groupby([cat1,cat2]).size().reset_index(name='Size')
dement = bar.loc[bar[cat2] == 'Dementia']
notDement = bar.loc[bar[cat2] == 'No dementia']
dement = dement.drop([cat2],axis=1)
notDement = notDement.drop([cat2],axis=1)
dement.columns = [cat1,'Dementia']
notDement.columns = [cat1,'noDementia']
dement.reset_index(inplace = True)
notDement.reset_index(inplace = True)
# set up frequency chart for dementia cases with thal stage
bar = pd.merge(dement, notDement, on=cat1, how='outer')
bar = bar.sort_values(cat1)
#bar = bar.fillna(0)
# bar = bar.dropna() #previously (0) but I think this was a bug
bar.drop(labels = ["index_x","index_y"],axis = 1,inplace = True)
bar.reset_index(inplace = True)
bar.drop(labels = ["index"],axis = 1,inplace = True)
# print(bar)
return bar
# -
# #### Extracting data from files loaded for two centers (Cambridge and New Castle) using both Neuropathology and Clinical datasets
# +
# load clinical and pathological dataframes with datasets
clin = pd.read_csv('../data/CFAS_ClinicalData.csv',header = 0) # read clinical dataset
patho = pd.read_excel('../data/CFAS_NeuropathologyData May 2018.xlsx', header = 21) # load pathological dataset
patho.dropna(subset=['Case'], inplace=True) # drop missing values
clin.rename(columns={'labno':'Case'}, inplace=True) # renaming a feature
# Create Seperate dataframes by centre
# Cambridge
# Clinical dataset
clinCambridge = clin['Case'].str.startswith('RH')
clinCambridge = clin[clinCambridge].copy()
# Pathological dataset
pathoCambridge = patho['Case'].str.startswith('RH')
pathoCambridge = patho[pathoCambridge].copy()
short = pd.to_numeric(clinCambridge.loc[:,'Case'].str[2:])
clinCambridge.loc[:,'Case'] = short
clinCambridge.sort_values(by=['Case'], inplace = True)
clinCambridge.head()
short = pathoCambridge.loc[:,'Case'].str[2:]
pathoCambridge.loc[:,'Case'] = pd.to_numeric(short)
pathoCambridge.sort_values(by=['Case'],inplace =True)
cambridge = pathoCambridge.merge(clinCambridge, how = 'left', on = ['Case'])
cambridge.head()
# Newcastle
clinNewcastle = clin['Case'].str.startswith('NA')
clinNewcastle = clin[clinNewcastle].copy()
pathoNewcastle = patho['Case'].str.startswith('NA')
pathoNewcastle = patho[pathoNewcastle].copy()
short = pd.to_numeric(clinNewcastle.loc[:,'Case'].str[2:].str.replace("/",""))
clinNewcastle.loc[:,'Case'] = short
clinNewcastle.sort_values(by=['Case'], inplace = True)
short = pathoNewcastle.loc[:,'Case'].str[2:].str.replace("/","")
pathoNewcastle.loc[:,'Case'] = pd.to_numeric(short)
pathoNewcastle.sort_values(by=['Case'],inplace =True)
newcastle = pathoNewcastle.merge(clinNewcastle, how = 'left', on = ['Case'])
# newcastle.head()
# Creates master data set containing both cambridge and newcastle data sets
master = cambridge.append(newcastle).copy()
master = master[(master[['dem_nver4']] != 0).all(axis=1)]
master.to_csv(r'../data/master.csv',mode = 'w',index=False)
patho.to_csv(r'../data/patho.csv',mode = 'w',index=False)
# count controls and dementia
# count cluster memberships
dementias=master.dem_nver4.value_counts()
print('No dementia : {} \nDementia : {}'.format(dementias[0], dementias[1]))
print('Samples : {} \nFeatures : {}'.format(master.shape[0], master.shape[1]))
# +
# extract neuropathology dataset
patho_features = pd.read_csv('../data/Neuropathology Features.csv',header = 0).Features
master = pd.read_csv('../data/master.csv',header = 0)
master.rename(columns={'aged':'age', 'brainwgt':'brain weight'}, inplace=True)
patho_data = master[patho_features].copy()
patho_data['dem_nver4'] = master['dem_nver4'].copy()
patho_data.columns
# -
# group features values within some different ranges by calling function defined above
patho_data["age"] = patho_data.apply(ageCat,axis=1)
patho_data["brain weight"] = patho_data.apply(brainwgt_Cat,axis=1)
patho_data["CAAAreas"] = patho_data.apply(CAAAreas_Cat,axis=1)
patho_data["CAATotalSev"] = patho_data.apply(CAATotalSev_Cat,axis=1)
patho_data["CAAParenc"] = patho_data.apply(CAAParenc_Cat,axis=1)
patho_data["CAAMeningeal"] = patho_data.apply(CAAMeningeal_Cat,axis=1)
from natsort import natsorted
df = patho_data.age.value_counts().sort_index().reset_index()
adf = df['index'].values
if adf.dtype != 'float64':
adf = natsorted(adf)
print(adf)
else:
print('good')
# +
patho_feature_order = pd.DataFrame()
patho_feature_order['Features'] = ('Braak stage',
'BrainNet tau stage',
'age',
'CAA type',
'CAA meningeal',
'brain weight',
'Thal phase',
'CAA parenchymal',
'CAA total severity',
'subpial TSA in mesial temporal lobe',
'subpial brainstem',
'CAA areas',
'TSA-any',
'CAA parietal',
'CAA hippocampus',
'CAA occipital',
'Subpial mesial temporal',
'CAA temporal',
'CAA frontal',
'subpial TSA in brainstem',
'CAA cerebellum',
'Aβ stage typical',
'hippocampal tau stage',
'Temporal microinfarct',
'frontal microinfarct',
'TSA-total',
'subcortical stage',
'PART-all',
'cortical stage',
'PART-definite',
'Occipital microinfarct',
'microinfarct stage',
'subpial TSA in expanded cortex',
'Argyrophilic grains',
'parietal microinfarct',
'Tufted astrocytes',
'dem_nver4')
# add the class label
patho_features = patho_feature_order.iloc[:-1,:]
# +
patho_data.rename(columns={'BraakStage':'Braak stage',
'BrainNetStage':'BrainNet tau stage',
# 'aged':'age',
'CAATotalSev':'CAA total severity',
'CAAMeningeal':'CAA meningeal',
# 'brainwgt':'brain weight',
'CAAType':'CAA type',
'ThalStage':'Thal phase',
'CAAAreas':'CAA areas',
'SubpialBrainstem':'subpial brainstem',
'CAAParenc':'CAA parenchymal',
'TSATotal':'TSA-total',
'TSAAny':'TSA-any',
'CAAParietal':'CAA parietal',
'CAAHippocampus':'CAA hippocampus',
'CAAFrontal':'CAA frontal',
'CAAOccipital':'CAA occipital',
'CAACerebellum':'CAA cerebellum',
'AbStageTypical':'Aβ stage typical',
'HippocTauStage':'hippocampal tau stage',
'CxSPETSA':'subpial TSA in expanded cortex',
'MTSPETSA':'subpial TSA in mesial temporal lobe',
'BSSPETSA':'subpial TSA in brainstem',
'SubcorticalStage':'subcortical stage',
'CorticalStage':'cortical stage',
'SubpialMesTemp':'Subpial mesial temporal',
'PARTall':'PART-all',
'MicroinfarctStage':'microinfarct stage',
'ArgyrGrains':'Argyrophilic grains',
'OccipMicroing':'Occipital microinfarct',
'ParMicrin':'parietal microinfarct',
'PARTdefinite':'PART-definite',
'FrontalMicroin':'frontal microinfarct',
'TempMicroinf':'Temporal microinfarct',
'CAATemp':'CAA temporal',
'TuftedAst':'Tufted astrocytes'} , errors="raise", inplace=True)
patho_features = patho_data.columns
patho_data = patho_data[patho_feature_order.Features]
# -
patho_features
# +
import numpy as np
import seaborn as sns
import pandas as pd
import matplotlib.pyplot as plt
from natsort import natsorted
import textwrap
sns.set_theme(style="ticks", color_codes=True)
x_axis=range(5)
fig, axes =plt.subplots(6,6, figsize=(16,18), sharey=False) #
axes = axes.flatten()
for ax, col in zip(axes, patho_data.columns):
x, y, hue = col, "proportion", "dem_nver4"
hue_order = ["No dementia", "Dementia"]
order = pd.DataFrame(patho_data[x].value_counts().sort_index().reset_index())
order= order['index'].values
if order.dtype != 'float64':
order = natsorted(order)
g = (patho_data[x]
.groupby(patho_data[hue])
.value_counts(normalize=True)#.sort_index()
.rename(y)
.reset_index()
.pipe((sns.barplot, "data"), x=x, y=y, hue=hue, order=order, ax=ax))
g.legend([],[], frameon=False)
plt.setp(ax.get_xticklabels(), rotation=45, size='14', horizontalalignment="right")
plt.setp(ax.get_yticklabels(), size='14')
ax.xaxis.get_label().set_fontsize(14)
ax.yaxis.get_label().set_fontsize(14)
for ax in axes:
ax.set_ylabel('')
axes[0].set_ylabel('proportion', size='16')
axes[6].set_ylabel('proportion', size='16')
axes[12].set_ylabel('proportion', size='16')
axes[18].set_ylabel('proportion', size='16')
axes[24].set_ylabel('proportion', size='16')
axes[30].set_ylabel('proportion', size='16')
plt.legend(loc='best');
l = plt.legend()
l.set_title('Status')
plt.setp(ax.get_legend().get_texts(), fontsize='14') # for legend text
plt.setp(ax.get_legend().get_title(), fontsize='14') # for legend title
plt.tight_layout()
plt.subplots_adjust(wspace=0.28, hspace=.7)
# plt.xticks(wrap=True)
fig.savefig('Figures/Neuropathology_Distrbution.png',dpi=300, bbox_inches="tight")
fig.savefig('Figures/Neuropathology_Distrbution.pdf',dpi=300, bbox_inches="tight")
plt.show()
# +
# load clinical and pathological dataframes with datasets
features = pd.read_csv('../data/neuropathology_features.csv',header = 0).features
feats= pd.DataFrame(features)
feats.replace({'features':{'aged':'age', 'brainwgt':'brain weight'}}, inplace=True)
feats = feats.squeeze()
patho = master[feats]
patho['age']=master.age
#For Preprocess and Analysis of Selected Features
patho_selected_features = patho.copy()
patho_with_AB = patho_selected_features
patho.rename(columns={'BraakStage':'Braak stage',
'BrainNetStage':'BrainNet tau stage',
'CAATotalSev':'CAA total severity',
'CAAMeningeal':'CAA meningeal',
'CAAType':'CAA type',
'ThalStage':'Thal phase',
'CAAAreas':'CAA areas',
'SubpialBrainstem':'subpial brainstem',
'CAAParenc':'CAA parenchymal',
'TSATotal':'TSA-total',
'TSAAny':'TSA-any',
'CAAParietal':'CAA parietal',
'CAAHippocampus':'CAA hippocampus',
'CAAFrontal':'CAA frontal',
'CAAOccipital':'CAA occipital',
'CAACerebellum':'CAA cerebellum',
'AbStageTypical':'Aβ stage typical',
'HippocTauStage':'hippocampal tau stage',
'CxSPETSA':'subpial TSA in expanded cortex',
'MTSPETSA':'subpial TSA in mesial temporal lobe',
'BSSPETSA':'subpial TSA in brainstem',
'SubcorticalStage':'subcortical stage',
'CorticalStage':'cortical stage',
'SubpialMesTemp':'Subpial mesial temporal',
'PARTall':'PART-all',
'MicroinfarctStage':'microinfarct stage',
'ArgyrGrains':'Argyrophilic grains',
'OccipMicroing':'Occipital microinfarct',
'ParMicrin':'parietal microinfarct',
'PARTdefinite':'PART-definite',
'FrontalMicroin':'frontal microinfarct',
'TempMicroinf':'Temporal microinfarct',
'CAATemp':'CAA temporal',
'TuftedAst':'Tufted astrocytes'} , errors="raise", inplace=True)
# compute correlations
corr = patho.corr('spearman')
# plot correlation
fig=plt.figure(figsize=(15,13))
# plt.figure(figsize=(15,10))
plt.rcParams['savefig.facecolor']='white'
sns.set_theme(style="ticks", color_codes=True)
ax = sns.heatmap(
corr,
vmin=-1, vmax=1,
center=0,
cmap=sns.diverging_palette(30, 245, n=100, as_cmap=True),#'RdBu_r', #
square=True,
cbar_kws={'label': 'Spearman Coefficients', "shrink": .9})
ax.figure.axes[-1].yaxis.label.set_size(14)
plt.xticks(fontsize=14)
plt.yticks(fontsize=14)
plt.tight_layout()
plt.savefig('Figures/spearman_correlation_heatmap.png',dpi=300, bbox_inches="tight")
plt.savefig('Figures/spearman_correlation_heatmap.pdf',dpi=300, bbox_inches="tight")
# -
(master.groupby(['dem_nver4', 'sex']).agg({'age': ['mean', 'count', 'median']}))
|
python/1. CFAS Distribution and Correlation Figures.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# I want to write an equation that recreates the RC-as-a-distance-ladder precision value reported in Hawkins+17, and then do the calculation for the updated spread in $K$ for this work.
# $m - M = 5log10(d) - 5 + A$
#
# Lets assume for these purposes that extinction is 0, or well known.
# Now, lets rearrange to solve for $d$.
#
# $d = 10^{(m - M + 5)/5}$
# Lets say we have a star which we know belongs to the Red Clump with a magnitude of $m_K = 13$. The RC parameters reported in Hawkins+17 are:
#
# $\mu_{RC} = -1.61$
#
# $\sigma_{RC} = 0.17$
#
# Lets treat $\mu_{RC}$ as the value for $M$, and $\sigma_{RC}$, the spread of the RC, as an uncertainty which we propagate. Lets assume there to be no uncertainty on apparent magnitude $m$.
# +
import numpy as np
from astropy import units as u
def d(m, M):
return 10**((m - M + 5)/5) * u.pc
# -
m = 13.
M = -1.61
s = 0.17
dist = d(m, M)
print('Distance : {:.2f}'.format(dist))
# Now to calculate the uncertainty incurred from the spread of the RC.
def err(m, M, s):
return np.sqrt((-2**(0.2*(m - M + 5)) * 5**((m - M)/5) * np.log(10))**2 * s**2) * u.pc
error = err(m, M, s)
print('Error : {:.2f}'.format(error))
print('Fractional Error : {:.2f}'.format(error/dist * 100 * u.percent))
# This is in agreement with the values reported in Hawkins+17. Nice!
# In our Hall+19 work, we have the following RC parameters:
#
# $\mu_{RC} = -1.638$
#
# $\sigma_{RC} = 0.03$
#
# So we find a fractional error of:
# +
m = 13.
M = -1.638
s = 0.03
dist = d(m, M)
error = err(m, M, s)
print('Distance : {:.2f}'.format(dist))
print('Error : {:.2f}'.format(error))
print('Fractional Error : {:.2f}'.format(error/dist * 100 * u.percent))
# -
# And in the G band:
# +
m = 13.
M = 0.546
s = 0.13
dist = d(m, M)
error = err(m, M, s)
print('Distance : {:.2f}'.format(dist))
print('Error : {:.2f}'.format(error))
print('Fractional Error : {:.2f}'.format(error/dist * 100 * u.percent))
# -
|
code/Scripts/clump-ladder.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data Collection Using Spotify Web API
#
# ## Spotify Web API
# Spotify has a number of [API endpoints](https://developer.spotify.com/documentation/web-api/reference-beta/) available to access the Spoitfy data. In this notebook, I use the following endpoints:
#
# + [search enpoint](https://developer.spotify.com/documentation/web-api/reference/search/search/) to get the track IDs
# + [audio features endpoint](https://developer.spotify.com/documentation/web-api/reference/tracks/get-several-audio-features/) to get the corresponding audio features.
#
# ## Purpose of Notebook
# The purpose of this notebook is to show how to collect and store audio features data for tracks from the [official Spotify Web API](https://developer.spotify.com/documentation/web-api/) for futher exploratory data analysis and machine learning.
# # 1. Setup
# The following code uses `spotipy` from the [Spotify](https://spotipy.readthedocs.io/en/latest/) library. Spotipy is a python library for accessing the Spotify web API.
import spotipy
from spotipy.oauth2 import SpotifyClientCredentials
import pandas as pd
import timeit
# +
cid ="c27d762046d144e48d9d7d929e9c2206"
secret = "<KEY>"
client_credentials_manager = SpotifyClientCredentials(client_id=cid, client_secret=secret)
sp = spotipy.Spotify(client_credentials_manager=client_credentials_manager)
# -
# # Data Collection
#
# Data collection is done in 3 parts: first we need the artist lineup, second the track IDs, and third the audio features for each track ID.
df_lineup = pd.read_csv("data/anjunadeep-explorations-artists.txt", sep=",", header=0)
#df_lineup = pd.DataFrame({'artist_name':lineup})
print(df_lineup.shape)
df_lineup.head()
print(df_lineup['artist_uri'][2])
#print(df_lineup.at[0,'artist_id'])
# +
#Pull all of the artist's albums
sp_albums = sp.artist_albums(df_lineup['artist_uri'][0], album_type='album')
sp_singles = sp.artist_albums(df_lineup['artist_uri'][0], album_type='single')
sp_appears_on = sp.artist_albums(df_lineup['artist_uri'][0], album_type='appears_on')
sp_compilations = sp.artist_albums(df_lineup['artist_uri'][0], album_type='compilation')
#Store artist's albums' names' and uris in separate lists
album_names = []
album_uris = []
for i in range(len(sp_albums['items'])):
album_names.append(sp_albums['items'][i]['name'])
album_uris.append(sp_albums['items'][i]['uri'])
single_names = []
single_uris = []
for i in range(len(sp_singles['items'])):
single_names.append(sp_singles['items'][i]['name'])
single_uris.append(sp_singles['items'][i]['uri'])
appears_on_names = []
appears_on_uris = []
for i in range(len(sp_appears_on['items'])):
single_names.append(sp_appears_on['items'][i]['name'])
single_uris.append(sp_appears_on['items'][i]['uri'])
compilation_names = []
compilation_uris = []
for i in range(len(sp_compilations['items'])):
single_names.append(sp_compilations['items'][i]['name'])
single_uris.append(sp_compilations['items'][i]['uri'])
print(df_lineup['artist_name'][0])
print('Albums:')
for i in range(len(album_names)):
print(album_names[i], '-', album_uris[i])
print('\nSingles:')
for i in range(len(single_names)):
print(single_names[i], '-', single_uris[i])
print('\nAppears On:')
for i in range(len(appears_on_names)):
print(appears_on_names[i], '-', appears_on_uris[i])
print('\nCompilations:')
for i in range(len(compilation_names)):
print(compilation_names[i], '-', compilation_uris[i])
#Keep names and uris in same order to keep track of duplicate albums
# -
def albumTracks(uri):
album = uri #assign album uri to a_name
spotify_albums[album] = {} #Creates dictionary for that specific album
#Create keys-values of empty lists inside nested dictionary for album
spotify_albums[album]['album'] = [] #create empty list
spotify_albums[album]['track_number'] = []
spotify_albums[album]['id'] = []
spotify_albums[album]['name'] = []
spotify_albums[album]['uri'] = []
tracks = sp.album_tracks(album) #pull data on album tracks
for n in range(len(tracks['items'])): #for each song track
spotify_albums[album]['album'].append(album_names[album_count]) #append album name tracked via album_count
spotify_albums[album]['track_number'].append(tracks['items'][n]['track_number'])
spotify_albums[album]['id'].append(tracks['items'][n]['id'])
spotify_albums[album]['name'].append(tracks['items'][n]['name'])
spotify_albums[album]['uri'].append(tracks['items'][n]['uri'])
spotify_albums = {}
album_count = 0
for i in album_uris: #each album
albumTracks(i)
print("Album " + str(album_names[album_count]) + " songs has been added to spotify_albums dictionary")
album_count+=1 #Updates album count once all tracks have been added
# +
start = timeit.default_timer()
# create empty lists where the results are going to be stored
artist_name = []
track_name = []
popularity = []
track_id = []
for k in range(len(df_lineup)):
lineup = 'artist:' + df_lineup['artist_name'][k]
for i in range(0,1000,50):
track_results = sp.search(q=lineup, type='track', limit=50,offset=i)
for i, t in enumerate(track_results['tracks']['items']):
artist_name.append(t['artists'][0]['name'])
track_name.append(t['name'])
track_id.append(t['id'])
popularity.append(t['popularity'])
progress = '(' + str(k) + '/' + str(len(df_lineup)) + ')'
print(progress, "Finished downloading songs for: ", df_lineup['artist_name'][k])
stop = timeit.default_timer()
print ('Time to run this code (in seconds):', stop - start)
# -
# # 3. EDA
# Now for some exploratory data analysis on the data we just collected.
# Checking the `track_id` list:
print('number of elements in track_id list:', len(track_id))
# Loading the lists into a dataframe
df_tracks = pd.DataFrame({'artist_name':artist_name,
'track_name':track_name,
'track_id':track_id,
'popularity':popularity})
print(df_tracks.shape)
df_tracks.head()
# Let's view some information about the data frame
df_tracks.info()
# ## Checking Our Data
# There can be duplicates of the same track under different `track_id`s. This is caused by the track being released in singles albums and full albums.
#
# Let's check how many duplicates there are by checking the `artist_name` and `track_name`.
# +
# group the entries by artist_name and track_name and check for duplicates
duplicates = df_tracks.groupby(['artist_name','track_name'], as_index=True).size()
print("Number of duplicate tracks: ", duplicates[duplicates > 1].count() )
# -
# There are a bunch of duplicate tracks in the dataframe. But which should be dropped and which should be kept? We can sort by the track `popularity`.
#
# From the official [Spotify docs](https://developer.spotify.com/documentation/web-api/reference/search/search/):
#
# > "The popularity of the track. The value will be between 0 and 100, with 100 being the most popular.
# The popularity of a track is a value between 0 and 100, with 100 being the most popular. The popularity is calculated by algorithm and is based, in the most part, on the total number of plays the track has had and how recent those plays are.
# Generally speaking, songs that are being played a lot now will have a higher popularity than songs that were played a lot in the past. Duplicate tracks (e.g. the same track from a single and an album) are rated independently. Artist and album popularity is derived mathematically from track popularity. Note that the popularity value may lag actual popularity by a few days: the value is not updated in real time."
#
# The `popularity` metric isn't based solely on 'number of plays' but also how recently those plays were.
df_tracks.sort_values(by=['artist_name', 'track_name', 'popularity'], ascending=False, inplace=True)
df_tracks.drop_duplicates(subset=['artist_name', 'track_name'], keep='first', inplace=True)
# Now we check again if there are duplicates:
# group the entries by artist_name and track_name and check for duplicates
duplicates = df_tracks.groupby(['artist_name','track_name'], as_index=True).size()
print("Number of duplicate tracks: ", duplicates[duplicates > 1].count() )
# Alternatively, we can check for duplicates via:
df_tracks[df_tracks.duplicated(subset=['artist_name','track_name'],keep=False)].count()
# Check the number of tracks after dropping duplicates:
df_tracks.shape
# # 4. Retrieve Audio Features Data
# Using the [audio features endpoint](https://developer.spotify.com/documentation/web-api/reference/tracks/get-several-audio-features/) we can retrieve the audio features data for the tracks we have collected.
#
# There is a 100 track ID limit per query for this endpoint. We can use anested for loop to pull track IDs in batches of size 100.
# +
# Measuring time
start = timeit.default_timer()
rows = []
batchsize = 100
None_counter = 0
for i in range(0, len(df_tracks['track_id']), batchsize):
batch = df_tracks['track_id'][i:i+batchsize]
feature_results = sp.audio_features(batch)
for i, t in enumerate(feature_results):
if t == None:
None_counter = None_counter + 1
else:
rows.append(t)
print('Number of tracks where no audio features were available:', None_counter)
stop = timeit.default_timer()
print ('Code runtime (sec):', stop - start)
# -
# # 5. EDA & Data Preparation
print('Number of elements in track_feature list:', len(rows))
# Loading audio features into a dataframe:
df_audio_features = pd.DataFrame.from_dict(rows, orient='columns')
print("Shape of dataset:", df_audio_features.shape)
df_audio_features.head()
df_audio_features.info()
# Renaming `id` to `track_id` to match the `df_tracks` dataframe:
df_audio_features.rename(columns = {'id': 'track_id'}, inplace=True)
df_audio_features.shape # checking our progress
df_track_meta_data = df_audio_features[['track_id', 'analysis_url', 'track_href', 'type', 'uri']]
df_track_meta_data.to_csv('data/anjunadeep-explorations-track-meta-data.csv')
df_track_meta_data.head()
# Dropping all variables (columns) not relevant to the analysis:
columns_to_drop = ['analysis_url', 'track_href', 'type', 'uri']
df_audio_features.drop(columns_to_drop, axis=1, inplace=True)
# To combine the two dataframes we do an `inner` merge to only keep track IDs that are in both datasets.
df = pd.merge(df_tracks, df_audio_features, on='track_id', how='inner')
print("Shape of dataset:", df_audio_features.shape)
df.head()
df.info()
# Checking again for duplicate tracks:
df[df.duplicated(subset=['artist_name', 'track_name'], keep=False)]
#df.hist(by=df['artist_name'])
duplicates = df_tracks.groupby(['artist_name'], as_index=True).size()
print(duplicates)
#print("Number of duplicate artists: ", duplicates[duplicates > 1].count() )
# So far everything looks good so let's save the dataframe as a .csv file.
df.to_csv('data/anjunadeep-explorations-track-audio-features.csv')
|
.ipynb_checkpoints/anjunadeep-explorations-playlists-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (conda:jupyterhub-tutorial)
# language: python
# name: python3
# ---
# # JupyterHub Spawners
#
# Let's peek at the base classes:
from jupyterhub.spawner import Spawner, LocalProcessSpawner
# +
# Spawner??
# -
# Start is the key method in a Spawner. It's how we decide how to start the process that will become the single-user server:
# +
# LocalProcessSpawner.start??
# -
# Here is an example of a spawner that allows specifying extra *arguments* to pass to a user's notebook server, via `.options_form`. It results in a form like this:
#
# 
# +
from traitlets import default
class DemoFormSpawner(LocalProcessSpawner):
@default('options_form')
def _options_form(self):
default_env = "YOURNAME=%s\n" % self.user.name
return """
<label for="args">Extra notebook CLI arguments</label>
<input name="args" placeholder="e.g. --debug"></input>
""".format(env=default_env)
def options_from_form(self, formdata):
"""Turn html formdata (always lists of strings) into the dict we want."""
options = {}
arg_s = formdata.get('args', [''])[0].strip()
if arg_s:
options['argv'] = shlex.split(arg_s)
return options
def get_args(self):
"""Return arguments to pass to the notebook server"""
argv = super().get_args()
if self.user_options.get('argv'):
argv.extend(self.user_options['argv'])
return argv
def get_env(self):
"""Return environment variable dict"""
env = super().get_env()
return env
# -
# ## Exercise:
#
# Write a custom Spawner that allows users to specify *environment variables* to load into their server.
#
#
# -------
from dockerspawner import DockerSpawner
# +
# DockerSpawner.start??
# -
# ## Exercise:
#
# Subclass DockerSpawner so that users can specify via `options_form` what docker image to use.
#
# Candidates from the [Jupyter docker-stacks repo](https://github.com/jupyter/docker-stacks) include:
#
# - jupyter/minimal-singleuser
# - jupyter/scipy-singleuser
# - jupyter/r-singleuser
# - jupyter/datascience-singleuser
# - jupyter/pyspark-singleuser
#
# Or, build your own images with
#
# FROM jupyterhub/singleuser
#
# The easiest version will assume that the images are fetched already.
# ## Extra credit:
#
# Subclass DockerSpawner so that users can specify via `options_form` a GitHub repository to clone and install, a la [binder](http://mybinder.org).
|
spawners.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from kh2lib.kh2lib import kh2lib
lib = kh2lib()
import os,json
if not os.path.isdir("workspace"):
os.mkdir("workspace")
DIR = os.path.join(os.environ["USE_KH2_GITPATH"], "KH2", "msn", "jp")
for fn in os.listdir(DIR):
lib.editengine.bar_extract(os.path.join(DIR,fn), os.path.join(os.getcwd(),"workspace","msns", fn.split(".")[0]))
records = []
for barfn in os.listdir(os.path.join(os.getcwd(),"workspace","msns")):
info = {}
BARDIR = os.path.join(os.getcwd(), "workspace", "msns", barfn)
info["name"] = barfn
info["namelen"] = len(barfn)
count = 0
for fn in os.listdir(BARDIR):
count += 1
ext = fn.split(".")[1]
if "ext-{}".format(ext) in info:
info["ext-{}".format(ext)] += 1
else:
info["ext-{}".format(ext)] = 1
if ext == "ai":
info["ai_os_name"] = fn.split(".")[0]
aifile = open(os.path.join(BARDIR,fn), "rb").read()
name = ""
for b in range(len(aifile)):
byte = aifile[b]
if byte == 0x0:
name = aifile[:b].decode('utf-8')
break
info["ai_hex_name"] = name
info["ai_len"] = len(aifile)
if ext == "list":
info["list_len"] = len(open(os.path.join(BARDIR,fn),"rb").read())
info["filecount"] = count
records.append(info)
c = ''
# get total list of ext- filetypes
filetypes = []
for record in records:
for k in record:
if k.startswith("ext-"):
filetypes.append(k)
c = 'name,namelen,ai_os_name,ai_hex_name,ai_len,list_len,filecount,'
c += ','.join(set(filetypes))
cols = c.split(',')
c += "\n"
for r in records:
for col in cols:
if col in r:
#print(col,r[col])
c += str(r[col])
c += ','
c+="\n"
c
open(os.path.join("workspace", "msns.csv"),"w").write(c)
for dr in os.listdir(os.path.join(os.getcwd(), "workspace", "msns")):
aicount = 0
listcount = 0
for fn in os.listdir(os.path.join(os.getcwd(),"workspace","msns",dr)):
ext = fn.split(".")[-1]
if ext == "ai":
aicount += 1
if ext == "list":
listcount += 1
if aicount != 1:
print("{} has {} ai files".format(dr, aicount))
if listcount != 1:
print("{} has {} list files".format(dr, listcount))
# loop through and see stuff
|
examples/Other/Parse MSNs For Some Stats/Parse MSNs For Some Stats.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] tags=["header"]
# <table width="100%">
# <tr style="border-bottom:solid 2pt #009EE3">
# <td class="header_buttons">
# <a href="prepare_jupyter.zip" download><img src="../../images/icons/download.png" alt="biosignalsnotebooks | download button"></a>
# </td>
# <td class="header_buttons">
# <a href="https://mybinder.org/v2/gh/biosignalsplux/biosignalsnotebooks/mybinder_complete?filepath=biosignalsnotebooks_environment%2Fcategories%2FInstall%2Fprepare_jupyter.dwipynb" target="_blank"><img src="../../images/icons/program.png" alt="biosignalsnotebooks | binder server" title="Be creative and test your solutions !"></a>
# </td>
# <td></td>
# <td class="header_icons">
# <a href="../MainFiles/biosignalsnotebooks.ipynb"><img src="../../images/icons/home.png" alt="biosignalsnotebooks | home button"></a>
# </td>
# <td class="header_icons">
# <a href="../MainFiles/contacts.ipynb"><img src="../../images/icons/contacts.png" alt="biosignalsnotebooks | contacts button"></a>
# </td>
# <td class="header_icons">
# <a href="https://github.com/biosignalsplux/biosignalsnotebooks" target="_blank"><img src="../../images/icons/github.png" alt="biosignalsnotebooks | github button"></a>
# </td>
# <td class="header_logo">
# <img src="../../images/ost_logo.png" alt="biosignalsnotebooks | project logo">
# </td>
# </tr>
# </table>
# + [markdown] tags=["intro_info_title"]
# <link rel="stylesheet" href="../../styles/theme_style.css">
# <!--link rel="stylesheet" href="../../styles/header_style.css"-->
# <link rel="stylesheet" href="https://cdnjs.cloudflare.com/ajax/libs/font-awesome/4.7.0/css/font-awesome.min.css">
#
# <table width="100%">
# <tr>
# <td id="image_td" width="15%" class="header_image_color_13"><div id="image_img"
# class="header_image_13"></div></td>
# <td class="header_text"> Download, Install and Execute Jupyter Notebook Environment </td>
# </tr>
# </table>
# + [markdown] tags=["intro_info_tags"]
# <div id="flex-container">
# <div id="diff_level" class="flex-item">
# <strong>Difficulty Level:</strong> <span class="fa fa-star checked"></span>
# <span class="fa fa-star"></span>
# <span class="fa fa-star"></span>
# <span class="fa fa-star"></span>
# <span class="fa fa-star"></span>
# </div>
# <div id="tag" class="flex-item-tag">
# <span id="tag_list">
# <table id="tag_list_table">
# <tr>
# <td class="shield_left">Tags</td>
# <td class="shield_right" id="tags">install☁jupyter☁notebook☁download</td>
# </tr>
# </table>
# </span>
# <!-- [OR] Visit https://img.shields.io in order to create a tag badge-->
# </div>
# </div>
# + [markdown] tags=["test"]
# In every journey we always need to prepare our toolbox with the needed resources !
#
# With <strong><span class="color1">biosignalsnotebooks</span></strong> happens the same, being <strong><span class="color4">Jupyter Notebook</span></strong> environment the most relevant application (that supports <strong><span class="color1">biosignalsnotebooks</span></strong>) to take the maximum advantage during your learning process.
#
# In the following sequence of instruction it will be presented the operations that should be completed in order to have <strong><span class="color4">Jupyter Notebook</span></strong> ready to use and to open our .ipynb files on local server.
#
# <table width="100%">
# <tr>
# <td style="text-align:left;font-size:12pt;border-top:dotted 2px #62C3EE">
# <span class="color1">☌</span> A journey can be done through multiple paths and at this point you can also start programming with diversified resources and in multiple ways. A complete Python toolbox (which includes <span class="color4"><strong>Jupyter Notebook</strong></span> environment) is available for world community and a quick installation guide can be accessed on <a href="../Install/prepare_anaconda.ipynb"><span class="color1"><strong>"Download, Install and Execute Anaconda" <img src="../../images/icons/link.png" width="10px" height="10px" style="display:inline"></strong></span></a>
# </td>
# </tr>
# </table>
# -
# <hr>
# <p class="steps">PR - <strong><span class="color4">Jupyter Notebook</span></strong> environment is able to run code in multiple programming languages, but his primordial format has origin on <a href="https://ipython.org/" target="_blank"><strong><span class="color7">IPython Project <img src="../../images/icons/link.png" width="10px" height="10px" style="display:inline"></span></strong></a>.</p>
# <i>Taking this info into consideration, there is only one prerequisite to install <strong><span class="color4">Jupyter Notebook</span></strong>, which is the installation of a <strong><span class="color1">Python</span></strong> compiler</i>
# <p class="steps">PR1 - Access to <span class="color1"><strong>Python</strong></span> main page at <a href="https://www.python.org/">https://www.python.org/</a></p>
# <img src="../../images/install/prepare_jupyter/python_page.png">
# <p class="steps">PR2 - Click on "Download" tab</p>
# <img src="../../images/install/prepare_jupyter/python_download_page.png">
# <p class="steps">PR3 - Select the download link accordingly with your operating system (for the current example we will use Microsoft Windows)</p>
# <img src="../../images/install/prepare_jupyter/python_download.png">
# Wait a few moments until the download finishes...
# <p class="steps">PR4 - Execute the downloaded file (with a double-click in the file icon)</p>
# <img src="../../images/install/prepare_jupyter/python_executable_click.gif">
# <span class="color13" style="font-size:30px">⚠</span>
# <p class="steps" style="margin-top:0px">PR5 - Activate "Add Python *.* to PATH" and click in "Install Now". After this, please, follow the sequential instructions presented in the installer</p>
# <i><span class="color7">"Add Python *.* to PATH"</span> option is essential for executing <span class="color1">Python</span> in the operating system command line, as will be seen while installing <span class="color4">Jupyter Notebook</span> in the next steps</i>
# <img src="../../images/install/prepare_jupyter/install_python.gif">
# When the installation was complete you finally fulfill the only <span class="color4"><strong>Jupyter Notebook</strong></span> prerequisite.<br>So, lets advance to the <span class="color4"><strong>Jupyter Notebook</strong></span> installation !!!
# <hr>
# <p class="steps">1 - Access to the <strong><span class="color4">Jupyter Notebook</span></strong> official page at <a href="https://jupyter.org/">https://jupyter.org/</a></p>
# <img src="../../images/install/prepare_jupyter/jupyter_notebook_page.png">
# <p class="steps">2 - Go to "Install" tab in order to see detailed installation instructions (with different possibilities)</p>
# <img src="../../images/install/prepare_jupyter/jupyter_install_page.gif">
# We will follow the third option <span class="color1"><strong>"Install Jupyter with pip"</strong></span>
# <img width="50%" src="../../images/install/prepare_jupyter/jupyter_install_page_step3.png">
# <br>This option is the most common one when programmers need to install new <strong>Python</strong> packages for use during development tasks.
# <p class="steps">3 - Open a "Command Prompt"</p>
# <i>If you are a Microsoft Windows native, just type click on Windows logo (bottom-left corner of the screen) and type "cmd". Then press "Enter".</i>
# <p class="steps">4 - Type <span class="color7">"python -m pip install --upgrade pip"</span> (or <span class="color7">"python3 -m pip install --upgrade pip"</span> if you had a previously installed version 2 of Python) to update a very practical <span class="color1">Python</span> tool (<a href="https://pypi.org/project/pip/" target="_blank">pip <img src="../../images/icons/link.png" width="10px" height="10px" style="display:inline"></a>) for installing new packages</p>
# <img src="../../images/install/prepare_jupyter/jupyter_install_cmd.gif">
# <p class="steps">5 - Type <span class="color7">"python -m pip install jupyter"</span> (or <span class="color7">"python3 -m pip install jupyter"</span> if you had a previously installed version 2 of Python) to install the amazing <span class="color4">Jupyter Notebook</span> environment</p>
# <img src="../../images/install/prepare_jupyter/jupyter_install_cmd_final.gif">
# Our first mission is now completed with the installation of <span class="color4">Jupyter Notebook</span> !
# <hr>
# <span class="color13" style="font-size:30px">⚠</span>
# <p class="steps" style="margin-top:0px">You can skip step 6 if you did not close the previously opened console !</p>
# <p class="steps">6 - For executing <span class="color4">Jupyter Notebook</span> environment you should open a <strong>console</strong> (in your operating system).</p>
# <i>If you are a Microsoft Windows native, just type click on Windows logo (bottom-left corner of the screen) and type "cmd". Then press "Enter".</i>
# <p class="steps">7 - Type <strong>"jupyter notebook"</strong> inside the opened console. A local <span class="color4"><strong>Jupyter Notebook</strong></span> server will be launched.</p>
# <img src="../../images/install/prepare_jupyter/open_jupyter.gif">
# <p class="steps">8 - Now, you should navigate through your directories until reaching the folder where you want to create or open a Notebook (as demonstrated in the following video)</p>
#
# <span class="color13" style="font-size:30px">⚠</span>
# <p style="margin-top:0px">You should note that your folder hierarchy is unique, so, the steps followed in the next image, will depend on your folder organization, being merely illustrative </p>
# <img src="../../images/install/prepare_jupyter/create_notebook_part1.gif">
# <p class="steps">9 - For creating a new Notebook, "New" button (top-right zone of Jupyter Notebook interface) should be pressed and <span class="color1"><strong>Python 3</strong></span> option selected.</p>
# <i>A blank Notebook will arise and now you just need to be creative and expand your thoughts to others persons!!!</i>
# <img src="../../images/install/prepare_jupyter/create_notebook_part2.gif">
# This can be the start of something great. Now you have all the software conditions to create and develop interactive tutorials, combining Python with HTML !
#
# <strong><span class="color7">We hope that you have enjoyed this guide. </span><span class="color2">biosignalsnotebooks</span><span class="color4"> is an environment in continuous expansion, so don't stop your journey and learn more with the remaining <a href="../MainFiles/biosignalsnotebooks.ipynb">Notebooks <img src="../../images/icons/link.png" width="10px" height="10px" style="display:inline"></a></span></strong> !
# + [markdown] tags=["footer"]
# <hr>
# <table width="100%">
# <tr>
# <td class="footer_logo">
# <img src="../../images/ost_logo.png" alt="biosignalsnotebooks | project logo [footer]">
# </td>
# <td width="40%" style="text-align:left">
# <a href="../MainFiles/aux_files/biosignalsnotebooks_presentation.pdf" target="_blank">☌ Project Presentation</a>
# <br>
# <a href="https://github.com/biosignalsplux/biosignalsnotebooks" target="_blank">☌ GitHub Repository</a>
# <br>
# <a href="https://pypi.org/project/biosignalsnotebooks/" target="_blank">☌ How to install biosignalsnotebooks Python package ?</a>
# <br>
# <a href="https://www.biosignalsplux.com/notebooks/Categories/MainFiles/signal_samples.ipynb">☌ Signal Library</a>
# </td>
# <td width="40%" style="text-align:left">
# <a href="https://www.biosignalsplux.com/notebooks/Categories/MainFiles/biosignalsnotebooks.ipynb">☌ Notebook Categories</a>
# <br>
# <a href="https://www.biosignalsplux.com/notebooks/Categories/MainFiles/by_diff.ipynb">☌ Notebooks by Difficulty</a>
# <br>
# <a href="https://www.biosignalsplux.com/notebooks/Categories/MainFiles/by_signal_type.ipynb">☌ Notebooks by Signal Type</a>
# <br>
# <a href="https://www.biosignalsplux.com/notebooks/Categories/MainFiles/by_tag.ipynb">☌ Notebooks by Tag</a>
# </td>
# </tr>
# </table>
# + tags=["hide_both"]
from biosignalsnotebooks.__notebook_support__ import css_style_apply
css_style_apply()
# + tags=["hide_both"] language="html"
# <script>
# // AUTORUN ALL CELLS ON NOTEBOOK-LOAD!
# require(
# ['base/js/namespace', 'jquery'],
# function(jupyter, $) {
# $(jupyter.events).on("kernel_ready.Kernel", function () {
# console.log("Auto-running all cells-below...");
# jupyter.actions.call('jupyter-notebook:run-all-cells-below');
# jupyter.actions.call('jupyter-notebook:save-notebook');
# });
# }
# );
# </script>
|
notebookToHtml/biosignalsnotebooks_html_publish/Categories/Install/prepare_jupyter.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: dvfx2
# language: python
# name: dvfx2
# ---
# +
import torch
import torch.nn as nn
import torchvision.datasets
import h5py
import zipfile
import imageio
import os
import numpy
from torch.utils.data import Dataset
import matplotlib.pyplot as plt
import random
import pandas
import time
import cv2
#mport pandas, numpy, random
# -
torch.__version__
torch.cuda_version
# +
if torch.cuda.is_available():
torch.set_default_tensor_type(torch.cuda.FloatTensor)
for i in range(torch.cuda.device_count()):
print("Using CUDA :", i+1, torch.cuda.get_device_name(i))
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
device
# +
def generate_random_image(size):
random_data = torch.rand(size)
return random_data
def generate_random_seed(size):
random_data = torch.randn(size)
return random_data
# +
# modified from https://github.com/pytorch/vision/issues/720
class View(nn.Module):
def __init__(self, shape):
super().__init__()
self.shape = shape,
def forward(self, x):
return x.view(*self.shape)
# -
class CelebADataset(Dataset):
def __init__(self, file):
self.file_object = h5py.File(file, 'r')
self.dataset = self.file_object['img_align_celeba']
pass
def __len__(self):
return len(self.dataset)
def __getitem__(self, index):
if (index >= len(self.dataset)):
raise IndexError()
img = numpy.array(self.dataset[str(index)+'.jpg'])
return torch.cuda.FloatTensor(img) / 255.0
def plot_image(self, index):
plt.imshow(numpy.array(self.dataset[str(index)+'.jpg']), interpolation='nearest')
pass
pass
import torchvision.datasets
import h5py
import zipfile
import imageio
import os
# +
import torchvision.datasets
import h5py
import zipfile
import imageio
import os
print("start!!")
# location of the HDF5 package, yours may be under /gan/ not /myo_gan/
hdf5_file = './celeba_dataset/celeba_aligned_small.h5py'
# how many of the 202,599 images to extract and package into HDF5
total_images = 30000
with h5py.File(hdf5_file, 'w') as hf:
count = 0
with zipfile.ZipFile('./celeba_dataset/celeba.zip', 'r') as zf:
print("start!!")
for i in zf.namelist():
print("start!!")
if (i[-4:] == '.jpg'):
# extract image
ofile = zf.extract(i)
img = imageio.imread(ofile)
os.remove(ofile)
# add image data to HDF5 file with new name
hf.create_dataset('img_align_celeba/'+str(count)+'.jpg', data=img, compression="gzip", compression_opts=9)
count = count + 1
if (count%1000 == 0):
print("images done .. ", count)
pass
# stop when total_images reached
if (count == total_images):
break
pass
pass
pass
# -
celeba_dataset = CelebADataset('./celeba_dataset/celeba_aligned_small.h5py')
#celeba_dataset.plot_image(30000)
# +
with h5py.File('./celeba_dataset/celeba_aligned_small.h5py', 'r') as file_object:
for group in file_object:
print(group)
pass
celeba_dataset = CelebADataset('./celeba_dataset/celeba_aligned_small.h5py')
celeba_dataset.plot_image(1000)
# +
def generate_random_image(size):
random_data = torch.rand(size)
return random_data
def generate_random_seed(size):
random_data = torch.randn(size)
return random_data
# -
class View(nn.Module):
def __init__(self, shape):
super().__init__()
self.shape = shape,
def forward(self, x):
return x.view(*self.shape)
class Discriminator(nn.Module):
def __init__(self):
# initialise parent pytorch class
super().__init__()
# define neural network layers
self.model = nn.Sequential(
View(128*128*3),
nn.Linear(3*128*128, 100),
nn.LeakyReLU(),
nn.LayerNorm(100),
nn.Linear(100, 1),
nn.Sigmoid()
)
# create loss function
self.loss_function = nn.BCELoss()
# create optimiser, simple stochastic gradient descent
self.optimiser = torch.optim.Adam(self.parameters(), lr=0.0001)
# counter and accumulator for progress
self.counter = 0;
self.progress = []
pass
def forward(self, inputs):
# simply run model
return self.model(inputs)
def train(self, inputs, targets):
# calculate the output of the network
outputs = self.forward(inputs)
# calculate loss
loss = self.loss_function(outputs, targets)
# increase counter and accumulate error every 10
self.counter += 1;
if (self.counter % 10 == 0):
self.progress.append(loss.item())
pass
if (self.counter % 1000 == 0):
#print("counter = ", self.counter)
pass
# zero gradients, perform a backward pass, update weights
self.optimiser.zero_grad()
loss.backward()
self.optimiser.step()
pass
def plot_progress(self):
df = pandas.DataFrame(self.progress, columns=['loss'])
df.plot(ylim=(0), figsize=(16,8), alpha=0.1, marker='.', grid=True, yticks=(0, 0.25, 0.5, 1.0, 5.0))
pass
pass
# +
# %%time
# test discriminator can separate real data from random noise
print("start1")
D = Discriminator()
# move model to cuda device
D.to(device)
for image_data_tensor in celeba_dataset:
# real data
print("start2")
D.train(image_data_tensor, torch.cuda.FloatTensor([1.0]))
# fake data
D.train(generate_random_image((128,128,3)), torch.cuda.FloatTensor([0.0]))
pass
# +
# generator class
class Generator(nn.Module):
def __init__(self):
# initialise parent pytorch class
super().__init__()
# define neural network layers
self.model = nn.Sequential(
nn.Linear(100, 3*10*10),
nn.LeakyReLU(),
nn.LayerNorm(3*10*10),
nn.Linear(3*10*10, 3*128*128),
nn.Sigmoid(),
View((128,128,3))
)
# create optimiser, simple stochastic gradient descent
self.optimiser = torch.optim.Adam(self.parameters(), lr=0.0001)
# counter and accumulator for progress
self.counter = 0;
self.progress = []
pass
def forward(self, inputs):
# simply run model
return self.model(inputs)
def train(self, D, inputs, targets):
# calculate the output of the network
g_output = self.forward(inputs)
# pass onto Discriminator
d_output = D.forward(g_output)
# calculate error
loss = D.loss_function(d_output, targets)
# increase counter and accumulate error every 10
self.counter += 1;
if (self.counter % 10 == 0):
self.progress.append(loss.item())
pass
# zero gradients, perform a backward pass, update weights
self.optimiser.zero_grad()
loss.backward()
self.optimiser.step()
pass
def plot_progress(self):
df = pandas.DataFrame(self.progress, columns=['loss'])
df.plot(ylim=(0), figsize=(16,8), alpha=0.1, marker='.', grid=True, yticks=(0, 0.25, 0.5, 1.0, 5.0))
pass
pass
# +
D = Discriminator()
G = Generator()
D.to(device)
G.to(device)
epochs = 500
print("start!")
a = time.time()
for epoch in range(epochs):
print ("epoch = ", epoch + 1)
# train Discriminator and Generator
for image_data_tensor in celeba_dataset:
# train discriminator on true
D.train(image_data_tensor, torch.cuda.FloatTensor([1.0]))
# train discriminator on false
# use detach() so gradients in G are not calculated
D.train(G.forward(generate_random_seed(100)).detach(), torch.cuda.FloatTensor([0.0]))
# train generator
G.train(D, generate_random_seed(100), torch.cuda.FloatTensor([1.0]))
# Util Visualizaing
output = G.forward(generate_random_seed(100))
img = output.detach().cpu().numpy()
cv2.imwrite("v1_vis/face_" + str(epoch+1) + ".jpg", cv2.cvtColor(img*255, cv2.COLOR_BGR2RGB))
b = time.time()
print(b-a)
# -
D.plot_progress()
G.plot_progress()
# +
# plot several outputs from the trained generator
# plot a 3 column, 2 row array of generated images
f, axarr = plt.subplots(2,3, figsize=(16,8))
for i in range(2):
for j in range(3):
output = G.forward(generate_random_seed(100))
img = output.detach().cpu().numpy()
axarr[i,j].imshow(img, interpolation='none', cmap='Blues')
pass
pass
# +
# current memory allocated to tensors (in Gb)
torch.cuda.memory_allocated(device) / (1024*1024*1024)
# +
# total memory allocated to tensors during program (in Gb)
torch.cuda.max_memory_allocated(device) / (1024*1024*1024)
|
face_generate_model_v1_failure_vis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Python for Environmental Science Day 7
# ## Topics
# * Making Nice Plots
# * Matplotlib Usage
# * Matplotlib Tips and Tricks
# ## Making Nice Plots
# This day is meant to cover the usage of matplotlib (and to some extension seaborn), but first I would like to get a concept across that will help you to make nicer plots. The [data-ink ratio](https://youtu.be/JIMUzJzqaA8). This concept helps you to focus your graphics on the most essential parts. The following two bar charts represent the most extreme examples for a very high and a very low data to ink ratio.
#
#
# 
# This does not mean that every graphic you make has to be as spartan as the right one, but you should always think about what parts of your graphic are essential to its message. If you want to learn a bit more about this and similar concepts take a look at [this article](https://medium.com/marax-ai/intelligent-signals-visualising-data-df9152c10b00). Also, you might wanna take a look at [this article](http://journals.plos.org/ploscompbiol/article?id=10.1371/journal.pcbi.1003833) for "10 simple rules for better figures", but this is optional.
#
# ### Practice Questions
# * Explaine the data-ink ratio in your own words. Can you think of a graphic you made yourself that had a bad data-ink ratio?
# * What is the lie-factor of a graphic (see article)?
#
# ### Exercise 1
# Try to find two graphics each, that have a good or bad data-ink ratio, respectively. A good starting point for your search are [data is beautiful](https://www.reddit.com/r/dataisbeautiful/) and [data is ugly](https://www.reddit.com/r/dataisugly/).
# ## Matplotlib Usage
# After this is out of the way, let us start with the real topic for today: **matplotlib**. matplotlib is the main package in Python for 2D graphics (though it can do a bit 3D as well) and allows you to create publication ready figures with relative ease. For starters take a look at [this video](https://youtu.be/V-OWkPCYa0s) (it is a bit hard to understand, but I could not find a better video that covers this topic; take a look at the next section for an outline of what you should have understood from the video; you can probably stop watching after ~ 20 minutes, as he mainly talks about maps after that) to get a general feel for the way matplotlib is structured and the cool things you will be capable of once you get the hang of it. And [this video](https://youtu.be/q7Bo_J8x_dw) for other basic information and simple plotting.
#
# ## Matplotlib Structure
# As we learned in the video matplotlib is relatively similar to matlab, but has a few differences. First of all, it is free and open source, which allows a much larger community to work on it. The second important thing is the anatomy of matplotlib.
#
# 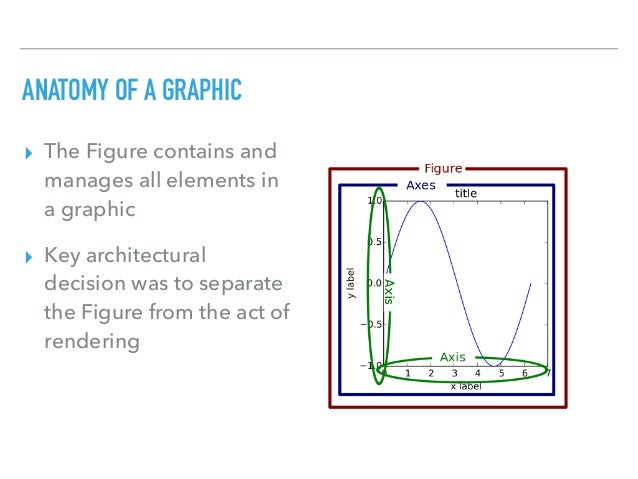
#
# As you can see in this picture matplotlib graphics subdivides intro three main things:
# * Figure: This is the overarching structure that contains everything else. If you work with several subplots, they are contained in one single figure, also.
# * Axes: This is where most of the real work happens, meaning that you will plot the things you want with the axes.
# * Axis: The x, and y axis which you can access and work with seperately.
#
# When using matplotlib it helps to be aware of the three layers of matplotlib:
# * Scripting Layer: This is the layer where you will usually do your actual coding using pyplot. Also, it is the only layer you are usually realizing you are working with.
# * Artist Layer: On this layer the real drawing of your graphics happens. This is usually an automated process when you call the pyplot functions, but you can also fiddle around here on your own if you need to.
# * Backend Layer: This is the part of matplotlib that interacts with your computer for saving and displaying files. When using matplotlib for creating scientific figures you can usually forget about it.
#
# Interfaces of matplotlib:
# * pylab: This is an old interface of matplotlib which is not used anymore, but you should know it existed if you see it in old code examples. It mainly has the same functionality as pyplot.
# * pyplot: This is the real deal. If you use matplotlib, you should access it over pyplot. It allows you to easily create figures and change parts of them.
# * object-oriented API: this is useful if you want to embed your matplotlib figure in something like a webpage. But when you use matplotlib for science you can probably forget about it. It allows you more control, but is also more complicated than pyplot.
#
# More good things to know about pyplot:
# * pyplot can work with several figures
# * if you call pyplot without having a figure, it will create one for you
# * pyplot.gca() gives you the current axes, pyplot.gcf() gives you the current figure
# * pyplot.close() clears everything, so you can avoid plotting in old figures
# * matplotlib works quite well with pandas dataframes
#
# An exhausting course on matplotlib can be found [here](https://github.com/zutn/oreilly-matplotlib-course) if you want to dive in deep.
#
# ### Practice Questions
# * What is the scripting layer of matplotlib and what is it used for?
# * What is the difference between a figure, axes and an axis?
#
# Have no worries, we will usually use pyplot to make things easier, but it can sometimes be helpful to know what happens behind the scenes of matplotlib.
#
# ### Most Basic Plotting
# So, after this introduction let us try a bit of plotting ourselves. As stated in the videos the most basic to plot things in matplotlib is the plot function, which allows you to draw dots or lines to represent your data. First let's start with a line.
import matplotlib.pyplot as plt
import random
# Make the plot reproducible
random.seed(1)
# Allow plotting in the notebook
# %matplotlib inline
# Create some fake data
data1 = [random.randint(0, 30) for i in range(10)]
plt.plot(data1, "-")
# But we can easily use the same data and represent them by little stars.
plt.plot(data1, "*")
# Or change the color.
plt.plot(data1, "-", color="black")
# As you can already see Python allows you to specify your figures quite a lot. This is often very helpful, as you can tailor it exactly to your needs, but will also lead you to long stackoverflow sessions to change one minor detail that you just could not figure out. If you take a look at [the documentation](https://matplotlib.org/api/_as_gen/matplotlib.pyplot.plot.html) you will see that plot() usually expects an x and y value. However, we did only provide one list. From this, plot() infers that this is a kind of series and simply plots the values against their index (which is in this case 0 to 9).
#
# **Note**: If we plot inside the notebook the figures are shown instantly. This will not happen if you execute the same code in Spyder. You will have to tell it explicitly that you want to see your figure by invoking **plt.show()**. But get in the habit of calling plt.show() here as well, so you get in the habit of doing so.
# ### Labelling Stuff
# When you create figures, especially scientific ones, you need to label your axes! This is relatively easy in matplotlib. A bit longer explanation can be found [here](https://youtu.be/aCULcv_IQYw). Basically you just tell pyplot to handle it.
plt.plot(data1, "-", color="black")
plt.xlabel("Index")
plt.ylabel("Data")
# Also label the figure as a whole
plt.title("A wonderful graph")
plt.show()
# When you have more than one dataset in a figure it becomes essential to label it, so the readers know what is what. As this is part of almost every figure you will make, matplotlib has an easy integration for this. Just specify a label for the plot you are creating (as a keyword in the function) and call **plt.legend()**. There are some cases where this approach will not give you the desired results, but stackoverflow will have a solution for you.
# Create a second dataset to plot also
import random
random.seed(0)
data2 = [random.randint(0, 30) for i in range(10)]
# Make the plotting
plt.plot(data1, "-", color="black", label="data1")
plt.plot(data2, "-", color="red", label="data2")
plt.xlabel("Index")
plt.ylabel("Data")
# Also label the figure as a whole
plt.title("Two wonderful graphs")
# And finally simply call the legend
plt.legend()
plt.show()
# ### Practice Questions
# * Can you include scientific notation in labels?
# ### Often used Kinds of Plots
# In this section we will look at the useful matplotlib figures. But first we need our pokemon dataset again.
import pandas as pd
pokemon = pd.read_csv("pokemon.csv")
# #### Bar Chart
# After this is out of the way we will take a look at [bar charts and histograms](https://youtu.be/ZyTO4SwhSeE). First let us prepare a part of a dataframe we want to plot. So let us get the max attack values for different 'Type 1' and sort them.
max_attack = pokemon.groupby("Type 1").max().loc[:,"Attack"]
max_attack
# And now we can create a barplot with this data. The barplot needs something to indicate where to plot the attack values on the x axis. Therefore, we also extract the types. And we change the rotation of the x-labels, so they are readable.
#
# Play around with the rotation to see how it influences the plot. Also try giving it a different color.
types = max_attack.index
plt.bar(x=types, height=max_attack)
plt.xticks(rotation=90)
plt.show()
# #### Histograms
# Histograms are often used to look at distributions of things. In our case, let us check out the distribution of defense values in water pokemon. First extract the data.
water_defense = pokemon.loc[pokemon["Type 1"] == "Water", "Defense"]
water_defense
# After we have the data we can simply call matplotlibs hist() function.
plt.hist(water_defense)
plt.show()
# This is again a bit rudimentary. So let us make this thing a bit more interesting, by adding labels and changing some properties of the histogram.
plt.hist(x=water_defense, histtype="step", linestyle=":", color="black", bins=15)
plt.xlabel("Defense Value")
plt.ylabel("Count")
plt.title("Distribution of Defense Values in Water Pokemon")
plt.show()
# Again, you can see that matplotlib allows you very easily to change your plots. You will learn to appreciate this feature very soon!
# #### Scatter Plots
# Another common plot is the scatter plot. As you might have already noticed the matplotlib functions all work very similarly. Therefore, I will keep this short. But if you need additional information, take a look at the [documentation of pyplot](https://matplotlib.org/api/_as_gen/matplotlib.pyplot.html#module-matplotlib.pyplot). First we need some data to plot against each other. Let us look at attack and speed of all pokemon.
attack = pokemon["Attack"]
speed = pokemon["Speed"]
plt.scatter(x=attack, y=speed, alpha=0.5)
plt.xlabel("Attack Value")
plt.ylabel("Speed Value")
plt.title("Scatter Plot Attack vs. Speed for all Pokemon")
plt.show()
# ### Practice Questions
# * What does the 'alpha' keyword do and what is its use?
# * What is the difference between a barplot and a histogram?
# * What are the advantages of a boxplot over a barplot?
# ### Exercise 2
# Write a Python program to draw a scatter plot taking a random distribution for the x and y values and plot them against each other. Make the scatter points black with an alpha of 0.5.
# ### Exercise 3
# Try to recreate [this figure](https://www.w3resource.com/w3r_images/matplotlib-basic-exercise-1.png). Also add a label to the line (with the keyword) and create a legend.
#
# Hint: It does not have to look exactly the same.
# ### Exercise 4
# Create a random sample of 30 values between 0 and 100 and create a boxplot of them.
# ### Exercise 5
# Create a random sample of 1000 values between 0 and 1. Use this to plot a histogram and set the histogram type to "step".
# ### Exercise 6
# Take a look at [the documentation](https://matplotlib.org/api/_as_gen/matplotlib.pyplot.boxplot.html#matplotlib.pyplot.boxplot) of the boxplot in matplotlib. Use this and your knowledge of pandas to create a figure that contains a boxplot for all meaningful attributes of the pokemon dataset.
# ### Working with Subplots
# Often those kinds of problems are better tackled with subplots. matplotlib allows you to plot several subplots in one figure. Take a look at [this short tutorial](https://matplotlib.org/gallery/recipes/create_subplots.html) and [this question](https://stackoverflow.com/questions/31726643/how-do-i-get-multiple-subplots-in-matplotlib) to get a first understanding. Then let us think about what kind of figure might profit from several subplots next to each other. My idea would be to make four subplots and compare attack to defense, speed, special attack and special defense respectively. First let us extract the data we need.
attack = pokemon["Attack"]
speed = pokemon["Speed"]
special_attack = pokemon["Sp. Atk"]
special_defense = pokemon["Sp. Def"]
defense = pokemon["Defense"]
# Next we create our subplots. Usually it is better to create all the subplots you need first and then plot, instead of creating one subplot at a time, as it makes for cleaner programming. As we want four subplots, two rows and two columns of subplots are needed.
fig, ax = plt.subplots(nrows=2, ncols=2, sharex=True, sharey=True)
ax[0,0].scatter(attack, defense, alpha=0.2)
ax[0,0].set_title("Attack vs. Defense")
ax[0,1].scatter(attack, speed, alpha=0.2)
ax[0,1].set_title("Attack vs Speed")
ax[1,0].scatter(attack, special_attack, alpha=0.2)
ax[1,0].set_title("Attack vs Special Attack")
ax[1,1].scatter(attack, special_defense, alpha=0.2)
ax[1,1].set_title("Attack vs Special Defense")
ax[1,1].set_xlabel("test")
plt.show()
# Now our four subbplots are stored in ax in the same way we see them. Meaning the left upper subplot is stored in ax[0,0], the upper right in ax[0,1], the lower left in ax[1,0] and the lower right in ax[1,1]. With this knowledge we can access ax to plot our scatterplots.
#
# In additon, we learned that a linear relationship of attack can be found for all other four attributes. However, it is not a very strong one.
# ### Practice Questions
# * What problems can occur when you do not use 'sharex' and 'sharey'?
# * How would you add an x and y label to the plot above?
# * Find out what plt.close() does and why it might be useful in the exercises.
#
# ### Saving your Figures
# Saving your files in matplotlib is easy. You simply have to call it with **plt.savefig()** and provide a name for the function. This will save the current figure in your working directory. Here are a few additional tips what savefig can do and what makes your saved figures nicer:
# +
plt.plot([1,0], [2,3], linestyle=":")
# Get the current figure, so you can alter its properties
fig = plt.gcf()
# Change the size of the figure to adjust it to your needs (play around)
fig.set_size_inches(1,2)
# Sometimes (especially with subplots) the default saved images are cropped off at the edges
# fig.tight_layout() corrects this
fig.tight_layout()
# Now let us save the figure. There are many keywords you can use but for me the most helpful
# are dpi (sets the resolution of your figure), bbox_inches (helps remove useless whitespace around the figure)
# and transparent (saves the figure without background)
plt.savefig("simple.png", dpi=300, bbox_inches="tight", transparent=True)
# -
# ### Exercise 7
# Download [this dataset](https://www.kaggle.com/dorbicycle/world-foodfeed-production) from kaggle and read it in.
#
#
# Do the following:
# * Let pandas describe the dataset (include="all") to give you a feeling for it and to determine if you read in everything correctly
# * Print all the unique values for the 'Item' column to see what's there (convert it to a list to see all)
# * Make a meaningful figure for each of the following descriptions:
# * shows the total amount of food and feed over the whole time period (for every year, not as a total).
# * compares amount of barley for food and for feed in Afghanistan in the year 1961.
# * shows the 50 countries that had the most food and feed combined for the whole time period (this is probably a bit easier if you use pandas plotting function).
# * explores if there is a linear correlation between food and feed amount for the whole time period in all countries.
# * explores if the soybean amount 2000 to 2009 is larger in the northern or southern hemisphere (keep the latitude in mind). Do this also for the time period 1990 to 1999. Plot both figures in a subplot next to each other.
# * shows the distribution of the amount of all food oils combined (by country, in one total per country) for the year 2000. [This](https://stackoverflow.com/questions/11350770/pandas-dataframe-select-by-partial-string) might be helpful.
#
# Hint: Some categorical data will be identified by pandas as numeric. Correct this misunderstanding.
#
# Hint: This will also require you to work with pandas a lot. Also, this exercise might take you a while, but will help you realize that the tools you learned are enough to explore a new dataset on your own.
#
# Hint: This is already quite similar to the final project, so make sure to ask a lot of questions.
#
# Hint: Reading the csv will probably raise a Unicode Error. Stackoverflow will help you!
#
# Hint: If you want something funny to cheer you up during this long exercise type the following in your console:
import antigravity
# ## Matplotlib Tips and Tricks
# During my usage of matplotlib I came across a few things that help me making figures that I want to share with you.
#
# ### Seaborn
# [Seaborn](https://seaborn.pydata.org/) is a package that builts upon matplotlib and allows you to make some very nice figures. For example this one here is created with seaborn:
#
# 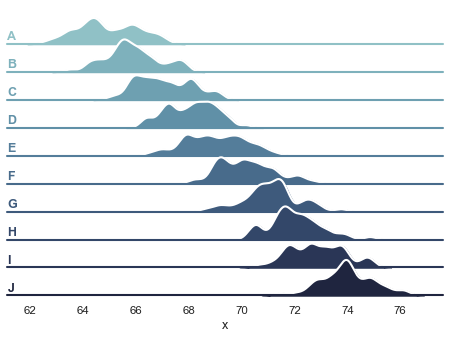
#
#
# In general I find the style and default color palette of seaborn much more appealing. If you want to have your matplotlib plots look more like seaborn, simply import seaborn at the beginning of your script. It will override some of the matplotlib defaults.
#
# ### Some Commands for a better Data-Ink Ratio
# When I introduced the concept of data-ink ratio at the beginning of the notebook, you might have wondered how you can include this concept in your coding. Here are a few tips:
# * Use the alpha keyword more often. With a clever usage you can highlight the most important parts of your plot
# * Get rid of gridlines
# * Get rid of borders
# * Use alpha also on labels
#
# So now let us try this with a simple figure:
#
#
#
plt.plot([1,0], [2,3], linestyle=":")
plt.grid(True)
plt.xlabel("X")
plt.ylabel("Y")
plt.title("X and Y")
plt.show()
# And now with all the advices mentioned above:
plt.plot([1,0], [2,3], linestyle=":")
# Remove grid
plt.grid(False)
# Apply alpha
alpha = 0.6
plt.xlabel("X", alpha=alpha)
plt.ylabel("Y", alpha=alpha)
plt.title("X and Y", alpha=alpha)
# Remove the borders
ax = plt.gca()
for spine in ax.spines.values():
spine.set_visible(False)
# Also use alpha on the ticklabels
plt.setp(ax.get_yticklabels(), alpha=alpha)
plt.setp(ax.get_xticklabels(), alpha=alpha)
plt.show()
# You do not have to make all your figures as barebone as this one, but I think it cannot hurt to know the tools.
#
# ### xkcd style
# You probably know [xkcd comics](https://xkcd.com/). If you like the style, you can simply activate it in your plots by calling **plt.xkcd()** at the beginning of your code. This makes your plots look like this:
#
#
# 
# ## Final Advice
#
# 
|
week_2/day_7_notebook.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="S-UXEXV-ifo6" colab_type="text"
# ### Read input data from Google Drive
# + id="CAxFJY6xlI0T" colab_type="code" colab={}
from pydrive.auth import GoogleAuth
from pydrive.drive import GoogleDrive
from google.colab import auth
from oauth2client.client import GoogleCredentials
# + id="-vt_4jPklI4f" colab_type="code" colab={}
# authenticate and create the PyDrive client.
auth.authenticate_user()
gauth = GoogleAuth()
gauth.credentials = GoogleCredentials.get_application_default()
drive = GoogleDrive(gauth)
# + id="myOIMc87lRuG" colab_type="code" colab={}
# get the raw data file
downloaded = drive.CreateFile({'id':"154ZPP7J54KPmq8TuFfX8kgAHb2ah0neo"})
downloaded.GetContentFile('design_thinking_data.csv')
# + id="xztilg9zlRxb" colab_type="code" colab={}
# read file as panda dataframe
import pandas as pd
raw_data = pd.read_csv('design_thinking_data.csv')
# + [markdown] id="vSSPld8jnqp2" colab_type="text"
# ### Clean up text
# + id="vEP-GX0tnpp-" colab_type="code" colab={}
# split lines by '\n' character
import os
LINE_SEP = os.linesep
raw_data['clean_text'] = raw_data['article_text'].apply(lambda s: str(s).split(LINE_SEP))
# + id="7dpfh4MAnpxQ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 102} outputId="a5aae2c9-20ea-4862-b398-aaf1cb4aeef7"
# download NLTK punctuations & stop words
import nltk
nltk.download('punkt')
nltk.download('stopwords')
# + id="8M8GcGk9npt_" colab_type="code" colab={}
# split the the text in the articles into sentences
from nltk.tokenize import sent_tokenize
sentences = []
for article in raw_data['clean_text']:
for paragraph in article:
sentences.append(sent_tokenize(paragraph)) if paragraph else None
# + id="bwffxhL_np0X" colab_type="code" colab={}
# flatten the list
sentences = [y for x in sentences for y in x]
# + id="pADAtHvenp3h" colab_type="code" colab={}
# remove punctuations, numbers and special characters
clean_sentences = pd.Series(sentences).str.replace("[^a-zA-Z]", " ")
# make alphabets lowercase
clean_sentences = [s.lower() for s in clean_sentences]
# + id="aQniQHZDnp66" colab_type="code" colab={}
# remove stop words
from nltk.corpus import stopwords
stop_words = stopwords.words('english')
def remove_stopwords(sen):
sen_new = " ".join([i for i in sen if i not in stop_words])
return sen_new
clean_sentences = [remove_stopwords(r.split()) for r in clean_sentences]
# + id="2XqpKMsVnp-e" colab_type="code" colab={}
# drop empty items in the list
clean_sentences = [clean_sentence for clean_sentence in clean_sentences if clean_sentence != '']
# + [markdown] id="9s3ODFwFszXD" colab_type="text"
# ### Embeddings
# + [markdown] id="LljC5Vs6q7as" colab_type="text"
# #### Word vectors
# + id="jngGGEw1nqHL" colab_type="code" colab={}
# get the Glove data file
downloaded = drive.CreateFile({'id':"1XlK7waXNOsGf3mdgMWiLVrDhCgmnDhr1"})
downloaded.GetContentFile('glove.6B.100d.txt')
# + id="jcs9efyviN8E" colab_type="code" colab={}
# extract word vectors from GloVe
from numpy import asarray
word_embeddings = {}
with open('glove.6B.100d.txt', encoding='utf-8') as f:
for line in f:
values = line.split()
word = values[0]
coefs = asarray(values[1:], dtype='float32')
word_embeddings[word] = coefs
# + [markdown] id="AyiT5sFftBUM" colab_type="text"
# #### Sentence vectors
# + id="9vl30p4MiN-U" colab_type="code" colab={}
# build sentence vectors
from numpy import zeros
sentence_vectors = []
for i in clean_sentences:
if len(i) != 0:
v = sum([word_embeddings.get(w, zeros((100,))) for w in i.split()])/(len(i.split())+0.001)
else:
v = zeros((100,))
sentence_vectors.append(v)
# + [markdown] id="YY9toPVXq_FN" colab_type="text"
# ### TextRank algorithm
# + id="tXsRqz2OrCde" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="aac86e33-4d53-4806-e372-d2f6ef6ca1d2"
# similarity matrix
import time
from scipy import sparse
from sklearn.metrics.pairwise import cosine_similarity
sentence_vectors_sparse = sparse.csr_matrix(sentence_vectors)
start_time = time.time()
similarities = cosine_similarity(sentence_vectors_sparse)
print(f"Similarity matrix calculated in {time.time() - start_time:.2f} seconds")
# + id="hCpiQN0frCfX" colab_type="code" colab={}
import networkx as nx
nx_graph = nx.from_numpy_array(similarities)
scores = nx.pagerank(nx_graph)
# + id="u2tkoYuyDrFN" colab_type="code" colab={}
# ranks
ranked_sentences = []
for i, s in enumerate(sentences[:100]):
ranked_sentences.append((scores[i], s))
ranked_sentences.sort(key=lambda x: x[1], reverse=True)
# + id="BtHTsvDEFtxL" colab_type="code" colab={}
# ranked_sentences = sorted(((scores[i], s) for i, s in enumerate(sentences)), reverse=True)
# + [markdown] id="UjuBU2K3rCsD" colab_type="text"
# ### Results
# + id="uH4n4xGAvujM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 547} outputId="177a6779-25eb-4bdb-c92d-1bfb67e64601"
# Specify number of sentences to form the summary
sn = 15
# Generate summary
for i in range(sn):
print(f"\n{ranked_sentences[i][1]}")
# + id="YPO93ux_rIqG" colab_type="code" colab={}
|
textrank_summerization.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Ranking document similarity using keywords and semantic matching
#
# #### 4OH4
# #### March 2020
#
# https://github.com/4OH4/doc-similarity
#
# This notebook contains examples of two methods for comparing content of text documents for similarity, such as might be used for search queries or content recommender systems. The first (TF-idf) scores document relationship based on the frequency of occurence of shared words. It is fast, and works well when documents are large and/or have lots of overlap. The second technique looks for shared words that address similar concepts, but does not require an exact match: for example, it links 'fruit and vegetables' with the word 'tomato'. This is slower, and gives less clear-cut results, but is good with shorter search queries or documents with low overlap.
#
# ## Contents
# 1. [TF-IDF to score shared key words](#sec1)
# 1b. [Using a lemmatizer](#sec1b)
# 1c. [Using the standalone module](#sec1c)
# 2. [Semantic matching using GloVe embeddings](#sec2)
# 2b. [Using the ready-made DocSim class](#sec2b)
#
# ## Requirements
# To install the required packages:
#
# pip install -r requirements.txt
#
# ## Known Issues
# - Warning generated by `gensim`: `RuntimeWarning: divide by zero encountered in true_divide` - I haven't been able to find the route cause of this, although does not appear to be producting eroneous results
# ### Load test data
import json
# +
# Load test data
with open('test_data.json') as in_file:
test_data = json.load(in_file)
titles = [item[0] for item in test_data['data']]
documents = [item[1] for item in test_data['data']]
print(f'{len(documents)} documents')
for idx in range(5):
print(idx, " \t ", titles[idx], " : \t", documents[idx][:100])
# -
# <a id="sec1"></a>
# ## 1. TF-IDF to score shared key words
# +
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.metrics.pairwise import linear_kernel
import nltk
from nltk.corpus import stopwords
nltk.download('punkt')
stop_words = set(stopwords.words('english'))
# +
search_terms = 'fruit and vegetables'
# search_terms = 'tomato'
# search_terms = 'sewing machine'
vectorizer = TfidfVectorizer(stop_words=stop_words)
vectors = vectorizer.fit_transform([search_terms] + documents)
# Calculate the word frequency, and calculate the cosine similarity of the search terms to the documents
cosine_similarities = linear_kernel(vectors[0:1], vectors).flatten()
document_scores = [item.item() for item in cosine_similarities[1:]] # convert back to native Python dtypes
# Print the top-scoring results and their titles
score_titles = [(score, title) for score, title in zip(document_scores, titles)]
for score, title in (sorted(score_titles, reverse=True, key=lambda x: x[0])[:5]):
print(f'{score:0.3f} \t {title}')
# -
# When using the search terms 'fruits and vegetables', only two documents have returned non-zero similarity scores - both contain the word 'fruit'. When searching for 'tomato', however, there are no matches; only the plural 'tomatoes' is present in the document corpus, and that does not match.
# <a id="sec1b"></a>
# ## 1b. Using a lemmatizer
# A lemmatizer reduces words down to their simplest 'lemma'. This is particularly helpful with dealing with plurals.
# +
# from: https://scikit-learn.org/stable/modules/feature_extraction.html
from nltk import word_tokenize
from nltk.stem import WordNetLemmatizer
class LemmaTokenizer:
"""
Interface to the WordNet lemmatizer from nltk
"""
ignore_tokens = [',', '.', ';', ':', '"', '``', "''", '`']
def __init__(self):
self.wnl = WordNetLemmatizer()
def __call__(self, doc):
return [self.wnl.lemmatize(t) for t in word_tokenize(doc) if t not in self.ignore_tokens]
# +
# Demonstrate the job of the tokenizer
tokenizer=LemmaTokenizer()
tokenizer('It was raining cats and dogs in FooBar')
# +
search_terms = 'fruit and vegetables'
# search_terms = 'tomato'
# search_terms = 'sewing machine'
# Initialise TfidfVectorizer with the LemmaTokenizer. Also need to lemmatize the stop words as well
token_stop = tokenizer(' '.join(stop_words))
vectorizer = TfidfVectorizer(stop_words=token_stop, tokenizer=tokenizer)
# Calculate the word frequency, and calculate the cosine similarity of the search terms to the documents
vectors = vectorizer.fit_transform([search_terms] + documents)
cosine_similarities = linear_kernel(vectors[0:1], vectors).flatten()
document_scores = [item.item() for item in cosine_similarities[1:]] # convert back to native Python dtypes
score_titles = [(score, title) for score, title in zip(document_scores, titles)]
for score, title in (sorted(score_titles, reverse=True, key=lambda x: x[0])[:5]):
print(f'{score:0.3f} \t {title}')
# -
# This gives better results - the document that contains the word 'tomatoes' is now scoring highly.
# <a id="sec1c"></a>
# ## 1c. Using the standalone module
#
# You can find the above functionality (TFidfVectorizer, stop_words, LemmaTokenizer, cosine_similarity) inside the `tfidf.py` module. This allows document scores to be calculated from a single function call:
from tfidf import rank_documents
document_scores = rank_documents(search_terms, documents)
# +
score_titles = [(score, title) for score, title in zip(document_scores, titles)]
for score, title in (sorted(score_titles, reverse=True, key=lambda x: x[0])[:5]):
print(f'{score:0.3f} \t {title}')
# -
# <a id="sec2"></a>
# ## 2. Semantic matching using GloVe embeddings
# This example and the class code for DocSim re-use and extend code from the Gensim tutorial notebook:
# https://github.com/RaRe-Technologies/gensim/blob/develop/docs/notebooks/soft_cosine_tutorial.ipynb
#
# The first part of this section runs though the individual steps in the process. This code is also available packaged in a ready-to-use class - scroll further down to see how it works.
# +
import json
import logging
from re import sub
from multiprocessing import cpu_count
import numpy as np
import gensim.downloader as api
from gensim.utils import simple_preprocess
from gensim.corpora import Dictionary
from gensim.models import TfidfModel
from gensim.models import WordEmbeddingSimilarityIndex
from gensim.similarities import SparseTermSimilarityMatrix
from gensim.similarities import SoftCosineSimilarity
# +
import logging
# Initialize logging.
logging.basicConfig(format='%(asctime)s : %(levelname)s : %(message)s', level=logging.WARNING) # DEBUG # INFO
# +
import nltk
# Import and download stopwords from NLTK.
nltk.download('stopwords') # Download stopwords list.
stopwords = set(nltk.corpus.stopwords.words("english"))
# +
# Support functions for pre-processing and calculation
# From: https://github.com/RaRe-Technologies/gensim/blob/develop/docs/notebooks/soft_cosine_tutorial.ipynb
def preprocess(doc):
# Tokenize, clean up input document string
doc = sub(r'<img[^<>]+(>|$)', " image_token ", doc)
doc = sub(r'<[^<>]+(>|$)', " ", doc)
doc = sub(r'\[img_assist[^]]*?\]', " ", doc)
doc = sub(r'http[s]?://(?:[a-zA-Z]|[0-9]|[$-_@.&+]|[!*\(\),]|(?:%[0-9a-fA-F][0-9a-fA-F]))+', " url_token ", doc)
return [token for token in simple_preprocess(doc, min_len=0, max_len=float("inf")) if token not in stopwords]
# -
# ### Prepare the data
# +
# Load test data
with open('test_data.json') as in_file:
test_data = json.load(in_file)
titles = [item[0] for item in test_data['data']]
documents = [item[1] for item in test_data['data']]
print(f'{len(documents)} documents')
# Print the first few document titles and intro text
# for idx in range(5):
# print(idx, "\t | \t", titles[idx], "\t | \t", documents[idx][:100])
# +
query_string = 'fruit and vegetables'
# Preprocess the documents, including the query string
corpus = [preprocess(document) for document in documents]
query = preprocess(query_string)
# query
# -
# ### Build the model
#
# The word embedding model is a large file, so loading is quite a long-running task.
# +
# %%time
# Download and/or load the GloVe word vector embeddings
if 'glove' not in locals(): # only load if not already in memory
glove = api.load("glove-wiki-gigaword-50")
similarity_index = WordEmbeddingSimilarityIndex(glove)
# +
# %%time
# Build the term dictionary, TF-idf model
# The search query must be in the dictionary as well, in case the terms do not overlap with the documents (we still want similarity)
# Lấy ra các từ điển và đánh index cho nó, và có 1 số feature khác nữa.
dictionary = Dictionary(corpus+[query])
tfidf = TfidfModel(dictionary=dictionary)
# Create the term similarity matrix.
# The nonzero_limit enforces sparsity by limiting the number of non-zero terms in each column.
# For my application, I got best results by removing the default value of 100
similarity_matrix = SparseTermSimilarityMatrix(similarity_index, dictionary, tfidf) # , nonzero_limit=None
# -
dictionary
# +
# Compute Soft Cosine Measure between the query and the documents.
query_tf = tfidf[dictionary.doc2bow(query)]
index = SoftCosineSimilarity(
tfidf[[dictionary.doc2bow(document) for document in corpus]],
similarity_matrix)
doc_similarity_scores = index[query_tf]
# -
index
# ### Output the document similarity results
# Output the similarity scores for top 15 documents
sorted_indexes = np.argsort(doc_similarity_scores)[::-1]
for idx in sorted_indexes[:15]:
print(f'{idx} \t {doc_similarity_scores[idx]:0.3f} \t {titles[idx]}')
# ### Find the most relevant terms in the documents
# For each term in the search query, what were the most similar words in each document?
doc_similar_terms = []
max_results_per_doc = 5
for term in query:
idx1 = dictionary.token2id[term]
for document in corpus:
results_this_doc = []
for word in set(document):
idx2 = dictionary.token2id[word]
score = similarity_matrix.matrix[idx1, idx2]
if score > 0.0:
results_this_doc.append((word, score))
results_this_doc = sorted(results_this_doc, reverse=True, key=lambda x: x[1]) # sort results by score
results_this_doc = results_this_doc[:min(len(results_this_doc), max_results_per_doc)] # take the top results
doc_similar_terms.append(results_this_doc)
# Output the results for the top 15 documents
for idx in sorted_indexes[:15]:
similar_terms_string = ', '.join([result[0] for result in doc_similar_terms[idx]])
print(f'{idx} \t {doc_similarity_scores[idx]:0.3f} \t {titles[idx]} : {similar_terms_string}')
# This shows which terms in each of the documents were most similar to terms in the search query. What it doesn't show, however, is the exact contribution of each of the terms to the document score, as each word similarity score will be weighted by the term frequency.
# <a id="sec2b"></a>
# ## 2b. Using the ready-made DocSim class
# The `DocSim` class wraps up functionality to prepare and compare data in a single object. It also persists the word embedding model to avoid having to reload it each time it is used. The word embedding model is loaded on initialisation, as this is quite a long-running task.
#
# `DocSim_threaded` has similar functionality, but loads the model in a separate thread. Similarity queries cannot be evaluated until the model is ready - check the status of the `model_ready` flag.
import json
import docsim
# +
# %%time
docsim_obj = docsim.DocSim(verbose=True)
# docsim_obj = docsim.DocSim_threaded(verbose=True)
# -
print(f'Model ready: {docsim_obj.model_ready}')
# +
# Load test data
with open('test_data.json') as in_file:
test_data = json.load(in_file)
titles = [item[0] for item in test_data['data']]
documents = [item[1] for item in test_data['data']]
print(f'{len(documents)} documents')
query_string = 'fruit and vegetables'
# +
# %%time
similarities = docsim_obj.similarity_query(query_string, documents)
# -
# Output the similarity scores for top 15 documents
for idx, score in (sorted(enumerate(similarities), reverse=True, key=lambda x: x[1])[:15]):
print(f'{idx} \t {score:0.3f} \t {titles[idx]}')
|
examples.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Homework 6, Part One: Lots and lots of questions about beer
# ### Do your importing and your setup
import pandas as pd
# ## Read in the file `craftcans.csv`, and look at the first first rows
df = pd.read_csv('craftcans.csv' , na_values = ['np.nan','Does not apply', 'NaN'] )
df.head()
# ## How many rows do you have in the data? What are the column types?
df.shape
df.dtypes
# # Checking out our alcohol
# ## What are the top 10 producers of cans of beer?
df.groupby('Beer').head(10)
# ## What is the most common ABV? (alcohol by volume)
df.ABV.describe()
# ## Oh, weird, ABV isn't a number. Convert it to a number for me, please.
#
# It's going to take a few steps!
#
# ### First, let's just look at the ABV column by itself
df.ABV
# ### Hm, `%` isn't part of a number. Let's remove it.
#
# When you're confident you got it right, save the results back into the `ABV` column.
#
# - *Tip: In programming the easiest way to remove something is to *replacing it with nothing*.
# - *Tip: "nothing" might seem like `NaN` sinc we talked about it a lot in class, but in this case it isn't! It's just an empty string, like ""*
# - *Tip: `.replace` is used for replacing ENTIRE cells, while `.str.replace` is useful for replacing PARTS of cells (see my New York example)*
df.ABV = df.ABV.str.replace('%' , '')
# ### Now let's turn `ABV` into a numeric data type
#
# Save the results back into the `ABV` column (again), and then check `df.dtypes` to make sure it worked.
#
# - *Tip: We used `.astype(int)` during class, but this has a decimal in it...*
df.ABV = df.ABV.astype(float)
# ## What's the ABV of the average beer look like?
#
# ### Show me in two different ways: one command to show the `median`/`mean`/etc, and secondly show me a chart
df.ABV.mean() , df.ABV.median()
df.ABV.hist()
# ### We don't have ABV for all of the beers, how many are we missing them from?
#
# - *Tip: You can use `isnull()` or `notnull()` to see where a column is missing data.*
# - *Tip: You just want to count how many `True`s and `False`s there are.*
# - *Tip: It's a weird trick involving something we usually use to count things in a column*
df.ABV.isna()
# # Looking at location
#
# Brooklyn used to produce 80% of the country's beer! Let's see if it's still true.
# ## What are the top 10 cities in the US for canned craft beer?
df.Location.head(10)
# ## List all of the beer from Brooklyn, NY
df[df.Location == 'Brooklyn, NY']
# ## What brewery in Brooklyn puts out the most cans of beer?
df[df.Location == 'Brooklyn, NY'].groupby('Brewery').Beer.count()
# ## What are the five most popular styles of beer produced by Sixpoint?
df[df.Brewery == 'Sixpoint Craft Ales'].value_counts('Style').head(5)
# ## List all of the breweries in New York state.
#
# - *Tip: We want to match *part* of the `Location` column, but not all of it.*
# - *Tip: Watch out for `NaN` values! You might be close, but you'll need to pass an extra parameter to make it work without an error.*
df[df.Location.str.contains("NY" , na=False)]
# ### Now *count* all of the breweries in New York state
df[df.Location.str.contains("NY" , na=False)].Location.count()
# # Measuring International Bitterness Units
#
# ## Display all of the IPAs
#
# Include American IPAs, Imperial IPAs, and anything else with "IPA in it."
#
# IPA stands for [India Pale Ale](https://www.bonappetit.com/story/ipa-beer-styles), and is probably the most popular kind of beer in the US for people who are drinking [craft beer](https://www.craftbeer.com/beer/what-is-craft-beer).
df[df.Style.str.contains("IPA" , na=False)]
# IPAs are usually pretty hoppy and bitter. IBU stands for [International Bitterness Unit](http://www.thebrewenthusiast.com/ibus/), and while a lot of places like to brag about having the most bitter beer (it's an American thing!), IBUs don't necessary *mean anything*.
#
# Let's look at how different beers have different IBU measurements.
# ## Try to get the average IBU measurement across all beers
df.IBUs.mean()
# ### Oh no, it doesn't work!
#
# It looks like some of those values *aren't numbers*. There are two ways to fix this:
#
# 1. Do the `.replace` and `np.nan` thing we did in class. Then convert the column to a number. This is boring.
# 2. When you're reading in your csv, there [is an option called `na_values`](http://pandas.pydata.org/pandas-docs/version/0.23/generated/pandas.read_csv.html). You can give it a list of **numbers or strings to count as `NaN`**. It's a lot easier than doing the `np.nan` thing, although you'll need to go add it up top and run all of your cells again.
#
# - *Tip: Make sure you're giving `na_values` a LIST, not just a string*
#
# ### Now try to get the average IBUs again
df.IBUs.mean()
# ## Draw the distribution of IBU measurements, but with *twenty* bins instead of the default of 10
#
# - *Tip: Every time I ask for a distribution, I'm looking for a histogram*
# - *Tip: Use the `?` to get all of the options for building a histogram*
# - *Tip: Make sure your `matplotlib` thing is set up right!*
df.IBUs.hist(bins=20)
# ## Hm, Interesting distribution. List all of the beers with IBUs above the 75th percentile
#
# - *Tip: There's a single that gives you the 25/50/75th percentile*
# - *Tip: You can just manually type the number when you list those beers*
df[df.IBUs > df.IBUs.quantile(0.75)]
# ## List all of the beers with IBUs below the 25th percentile
df[df.IBUs < df.IBUs.quantile(0.25)]
# ## List the median IBUs of each type of beer. Graph it.
#
# Put the highest at the top, and the missing ones at the bottom.
#
# - Tip: Look at the options for `sort_values` to figure out the `NaN` thing. The `?` probably won't help you here.
df_grouped_by_style = df.groupby('Style')
df_grouped_by_style[['IBUs']].median().sort_values(['IBUs'],ascending=False)
# +
df_grouped_by_style[['IBUs']].median().sort_values(['IBUs'],ascending=False).plot(kind="bar")
# -
# ## Hmmmm, it looks like they are generally different styles. What are the most common 5 styles of high-IBU beer vs. low-IBU beer?
#
# - *Tip: You'll want to think about it in three pieces - filtering to only find the specific beers beers, then finding out what the most common styles are, then getting the top 5.*
# - *Tip: You CANNOT do this in one command. It's going to be one command for the high and one for the low.*
# - *Tip: "High IBU" means higher than 75th percentile, "Low IBU" is under 25th percentile*
df[df.IBUs > df.IBUs.quantile(0.75)]['Style'].value_counts().head(5)
df[df.IBUs < df.IBUs.quantile(0.25)]['Style'].value_counts().head(5)
# ## Get the average IBU of "Witbier", "Hefeweizen" and "American Pale Wheat Ale" styles
#
# I'm counting these as wheat beers. If you see any other wheat beer categories, feel free to include them. I want ONE measurement and ONE graph, not three separate ones. And 20 to 30 bins in the histogram, please.
#
# - *Tip: I hope that `isin` is in your toolbox*
# +
#({'num_wings': [0, 3]})
df[df['Style'].isin(["Witbier", "Hefeweizen","American Pale Wheat Ale"])].IBUs.mean()
# -
# ## Draw a histogram of the IBUs of those beers
df[df['Style'].isin(["Witbier", "Hefeweizen","American Pale Wheat Ale"])].IBUs.hist(bins=25)
# ## Get the average IBU of any style with "IPA" in it (also draw a histogram)
#df.Narrative_1.str.contains("ALCOHOL", case=False)
df[df['Style'].str.contains("IPA", case=False , na=False)].IBUs.mean()
df[df['Style'].str.contains("IPA", case=False , na=False)].IBUs.hist(bins=25)
# ## Plot those two histograms on top of one another
#
# To plot two plots on top of one another, you do two steps.
#
# 1. First, you make a plot using `plot` or `hist`, and you save it into a variable called `ax`.
# 2. You draw your second graph using `plot` or `hist`, and send `ax=ax` to it as a parameter.
#
# It would look something like this:
#
# ```python
# ax = df.plot(....)
# df.plot(ax=ax, ....)
# ```
#
# (...except totally different)
ax = df[df['Style'].isin(["Witbier", "Hefeweizen","American Pale Wheat Ale"])].IBUs
ay = df[df['Style'].str.contains("IPA", case=False , na=False)].IBUs
import matplotlib.pyplot as plt
plt.hist([ax,ay], bins=25,label=['Wheat', 'IPA'])
plt.legend(loc='upper right')
# ## Compare the ABV of wheat beers vs. IPAs : their IBUs were really different, but how about their alcohol percentage?
#
# Wheat beers might include witbier, hefeweizen, American Pale Wheat Ale, and anything else you think is wheaty. IPAs probably have "IPA" in their name.
ax = df[df['Style'].isin(["Witbier", "Hefeweizen","American Pale Wheat Ale"])].ABV
ay = df[df['Style'].str.contains("IPA", case=False , na=False)].ABV
import matplotlib.pyplot as plt
plt.hist([ax,ay], bins=25,label=['Wheat', 'IPA'])
plt.legend(loc='upper right')
# ## Good work!
#
# If you made it this far you deserve a drink.
ay.head()
|
06-homework/beer/.ipynb_checkpoints/Dataset ONE - Beer cans-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_ds
# language: python
# name: conda_ds
# ---
# +
from nlp_adapter.train.training import set_up_exp
set_up_exp(
data_path='/home/apilkevich/notebooks/mipt/BD/data/input/train.tsv',
transformer_model_name='./pretrain_detox_s42/bs12_accum1_7500',
tokenizer_model_name='sberbank-ai/ruT5-base',
# pretrain_adapter_path='./adapter/adapter_1914.pt',
batch_size=64,
max_steps=2000,
dump_model_name='adapter_s42/bs64_accum3_lr3e3',
period_of_dump=174,
train_model=False,
train_adapter=True,
seed=42,
test_size=None,
device='1',
loss_mode='sum',
model_lr=3e-3,
num_accum_steps=1,
)
# -
|
notebooks/adapter_train.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pymysql
import random
from faker import Faker
fake =Faker('ko_KR')
Faker.seed(3);
db= pymysql.connect(host='localhost',user='root',password='<PASSWORD>',db='musinsa')
curs =db.cursor()
seq = 995
for _ in range(6):
while True:
ssn=fake.ssn()
if (ssn[2]=='0' and ssn[3]=='0') or (ssn[4]=='0' and ssn[5]=='0'):
continue
break
if ssn[7]=='1' or ssn[7]=='2':
birthday= '19'+ssn[0:2]+'-'+ssn[2:4]+'-'+ssn[4:6]
elif ssn[7]=='3' or ssn[7]=='4':
birthday= '20'+ssn[0:2]+'-'+ssn[2:4]+'-'+ssn[4:6]
if ssn[7]=='1' or ssn[7]=='3':
gender='M'
height=random.randint(160,190)
print(gender)
print(height)
if height >=190:
size='2XL'
elif height >=180:
size='XL'
elif height >=170:
size='L'
elif height >=170:
size='M'
elif height >=170:
size='S'
print(size)
elif ssn[7]=='2' or ssn[7]=='4':
gender = 'F'
height=random.randint(150,180)
print(gender)
print(height)
if height >=170:
size='L'
elif height >=160:
size='M'
elif height >=150:
size='S'
print(size)
query = "INSERT IGNORE INTO users(seq, id, password, name, cellphone,\
address, ssn, CI, DI, birthday, gender,height, size, join_date) \
VALUES("\
+str(seq)+",'"+fake.email() +"','"\
+fake.password(length=8, special_chars=False, upper_case=False)\
+"','"\
+fake.name()+"','"\
+fake.phone_number() +"','"\
+fake.address() +"','"\
+ssn +"','-','-','"\
+birthday+"','"\
+gender+"','"+str(height)+"','"+size+"', '"\
+str(fake.date_between(start_date='-10y', end_date='-2m'))+"');"
ret=curs.execute(query)
print(query)
if ret == 1:
seq += 1
else:
print("error",seq,"ret",ret)
db.commit()
db.close()
import pymysql
from faker import Faker
fake =Faker('ko_KR')
Faker.seed(1234);
import random
for _ in range(6):
while True:
ssn=fake.ssn()
if (ssn[2]=='0' and ssn[3]=='0') or (ssn[4]=='0' and ssn[5]=='0'):
continue
break
if ssn[7]=='1' or ssn[7]=='2':
birthday= '19'+ssn[0:2]+'-'+ssn[2:4]+'-'+ssn[4:6]
elif ssn[7]=='3' or ssn[7]=='4':
birthday= '20'+ssn[0:2]+'-'+ssn[2:4]+'-'+ssn[4:6]
if ssn[7]=='1' or ssn[7]=='3':
gender='M'
height=random.randint(160,190)
print(gender)
print(height)
if height >=190:
size='2XL'
elif height >=180:
size='XL'
elif height >=170:
size='L'
elif height >=170:
size='M'
elif height >=170:
size='S'
print(size)
elif ssn[7]=='2' or ssn[7]=='4':
gender = 'F'
height=random.randint(150,180)
print(gender)
print(height)
if height >=170:
size='L'
elif height >=160:
size='M'
elif height >=150:
size='S'
print(size)
|
Code/musinsa_users.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _uuid="2f0e779a6d98b9c9c06e2abe47f5428fa0e2c68c"
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.metrics import mean_squared_error
from sklearn.feature_extraction.text import TfidfVectorizer, CountVectorizer
from sklearn.decomposition import TruncatedSVD
from sklearn import preprocessing, model_selection, metrics
import lightgbm as lgb
from xgboost import XGBRegressor
from sklearn.model_selection import GridSearchCV
from lightgbm import LGBMRegressor
color = sns.color_palette()
# %matplotlib inline
pd.options.mode.chained_assignment = None
pd.options.display.max_columns = 9999
lgbr = LGBMRegressor()
# -
# %run '../00_rh_settings.ipynb'
# %run '../00_rh_load_processed.ipynb'
train_df = df
# +
import pandas as pd
import numpy as np
from sklearn.model_selection import GridSearchCV, RandomizedSearchCV
class EstimatorSelectionHelper:
def __init__(self, models, params):
if not set(models.keys()).issubset(set(params.keys())):
missing_params = list(set(models.keys()) - set(params.keys()))
raise ValueError("Some estimators are missing parameters: %s" % missing_params)
self.models = models
self.params = params
self.keys = models.keys()
self.grid_searches = {}
def fit(self, X, y, cv=3, n_jobs=3, verbose=1, scoring=None, refit=False):
for key in self.keys:
print("Running GridSearchCV for %s." % key)
model = self.models[key]
params = self.params[key]
gs = RandomizedSearchCV(model, params, cv=cv, n_jobs=n_jobs,
verbose=verbose, scoring=scoring, refit=refit, n_iter=30,
return_train_score=True)
gs.fit(X,y)
self.grid_searches[key] = gs
def score_summary(self, sort_by='mean_score'):
def row(key, scores, params):
d = {
'estimator': key,
'min_score': min(scores),
'max_score': max(scores),
'mean_score': np.mean(scores),
'std_score': np.std(scores),
}
return pd.Series({**params,**d})
rows = []
for k in self.grid_searches:
print(k)
params = self.grid_searches[k].cv_results_['params']
scores = []
for i in range(self.grid_searches[k].cv):
key = "split{}_test_score".format(i)
r = self.grid_searches[k].cv_results_[key]
scores.append(r.reshape(len(params),1))
all_scores = np.hstack(scores)
for p, s in zip(params,all_scores):
rows.append((row(k, s, p)))
df = pd.concat(rows, axis=1).T.sort_values([sort_by], ascending=False)
columns = ['estimator', 'min_score', 'mean_score', 'max_score', 'std_score']
columns = columns + [c for c in df.columns if c not in columns]
return df[columns]
# +
from catboost import CatBoostClassifier
models1 = {
'CatBoost': CatBoostClassifier(),}
params1 = {
'CatBoost': {
'iterations':[250,100,500,1000],
'depth': range(2,10),
'learning_rate': [0.001,0.05,0.1,0.2,0.3],
'l2_leaf_reg':[3,1,5,10,100],
'cat_features': [np.where(train_X.dtypes == np.object)[0]],
'logging_level': ['Silent'],
},
}
# -
df.drop(columns=['Unnamed: 0',"loanKey","rep_loan_date","first_loan","first_overdue_date"], inplace=True, errors="ignore")
df.dropna(inplace=True)
df = df.replace(r'\s+', np.nan, regex=True)
df = df.replace('nan', np.nan)
train_X = df.drop(columns="bad_flag")
train_y = df["bad_flag"]
helper1 = EstimatorSelectionHelper(models1, params1)
helper1.fit(train_X, train_y, scoring='f1', n_jobs=-1)
helper1.score_summary()
|
notebooks/model/baseline-gridsearch.ipynb
|