code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + nbsphinx="hidden"
# HIDDEN CELL
import sys, os
# Importing argopy in dev mode:
on_rtd = os.environ.get('READTHEDOCS', None) == 'True'
if not on_rtd:
p = "/Users/gmaze/git/github/euroargodev/argopy"
sys.path.insert(0, p)
import git
import argopy
from argopy.options import OPTIONS
print("argopy:", argopy.__version__,
"\nsrc:", argopy.__file__,
"\nbranch:", git.Repo(path=p, search_parent_directories=True).active_branch.name,
"\noptions:", OPTIONS)
else:
sys.path.insert(0, os.path.abspath('..'))
import xarray as xr
# xr.set_options(display_style="html");
xr.set_options(display_style="text");
# + raw_mimetype="text/restructuredtext" active=""
# .. _data_fetching:
# + [markdown] raw_mimetype="text/restructuredtext"
# # Fetching Argo data
# -
# To access Argo data, you need to use a data fetcher. You can import and instantiate the default argopy data fetcher
# like this:
from argopy import DataFetcher as ArgoDataFetcher
argo_loader = ArgoDataFetcher()
# Then, you can request data for a specific **space/time domain**, for a given **float** or for a given vertical **profile**.
# + [markdown] raw_mimetype="text/restructuredtext"
# ## For a space/time domain
# + raw_mimetype="text/restructuredtext" active=""
# Use the fetcher access point :meth:`argopy.DataFetcher.region` to specify a domain and chain with the :meth:`argopy.DataFetcher.to_xarray` to get the data returned as :class:`xarray.Dataset`.
#
# For instance, to retrieve data from 75W to 45W, 20N to 30N, 0db to 10db and from January to May 2011:
# -
ds = argo_loader.region([-75, -45, 20, 30, 0, 10, '2011-01-01', '2011-06']).to_xarray()
print(ds)
# Note that:
#
# - the constraints on time is not mandatory: if not specified, the fetcher will return all data available in this region.
# - the last time bound is exclusive: that's why here we specifiy June to retrieve data collected in May.
# ## For one or more floats
# + raw_mimetype="text/restructuredtext" active=""
# If you know the Argo float unique identifier number called a `WMO number <https://www.wmo.int/pages/prog/amp/mmop/wmo-number-rules.html>`_ you can use the fetcher access point :meth:`argopy.DataFetcher.float` to specify the float WMO platform number and chain with the :meth:`argopy.DataFetcher.to_xarray` to get the data returned as :class:`xarray.Dataset`.
#
# For instance, to retrieve data for float WMO *6902746*:
# -
ds = argo_loader.float(6902746).to_xarray()
print(ds)
# To fetch data for a collection of floats, input them in a list:
ds = argo_loader.float([6902746, 6902755]).to_xarray()
print(ds)
# ## For one or more profiles
# + raw_mimetype="text/restructuredtext" active=""
# Use the fetcher access point :meth:`argopy.DataFetcher.profile` to specify the float WMO platform number and the profile cycle number to retrieve profiles for, then chain with the :meth:`argopy.DataFetcher.to_xarray` to get the data returned as :class:`xarray.Dataset`.
#
# For instance, to retrieve data for the 12th profile of float WMO 6902755:
# -
ds = argo_loader.profile(6902755, 12).to_xarray()
print(ds)
# To fetch data for more than one profile, input them in a list:
ds = argo_loader.profile(6902755, [3, 12]).to_xarray()
print(ds)
| docs/data_fetching.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Variational inference in Bayesian neural networks
#
# This article demonstrates how to implement and train a Bayesian neural network with Keras following the approach described in [Weight Uncertainty in Neural Networks](https://arxiv.org/abs/1505.05424) (*Bayes by Backprop*). The implementation is kept simple for illustration purposes and uses Keras 2.2.4 and Tensorflow 1.12.0. For more advanced implementations of Bayesian methods for neural networks consider using [Tensorflow Probability](https://www.tensorflow.org/probability), for example.
#
# Bayesian neural networks differ from plain neural networks in that their weights are assigned a probability distribution instead of a single value or point estimate. These probability distributions describe the uncertainty in weights and can be used to estimate uncertainty in predictions. Training a Bayesian neural network via variational inference learns the parameters of these distributions instead of the weights directly.
#
# ## Probabilistic model
#
# A neural network can be viewed as probabilistic model $p(y \lvert \mathbf{x},\mathbf{w})$. For classification, $y$ is a set of classes and $p(y \lvert \mathbf{x},\mathbf{w})$ is a categorical distribution. For regression, $y$ is a continuous variable and $p(y \lvert \mathbf{x},\mathbf{w})$ is a Gaussian distribution.
#
# Given a training dataset $\mathcal{D} = \left\{\mathbf{x}^{(i)}, y^{(i)}\right\}$ we can construct the likelihood function $p(\mathcal{D} \lvert \mathbf{w}) = \prod_i p(y^{(i)} \lvert \mathbf{x}^{(i)}, \mathbf{w})$ which is a function of parameters $\mathbf{w}$. Maximizing the likelihood function gives the maximimum likelihood estimate (MLE) of $\mathbf{w}$. The usual optimization objective during training is the negative log likelihood. For a categorical distribution this is the *cross entropy* error function, for a Gaussian distribution this is proportional to the *sum of squares* error function. MLE can lead to severe overfitting though.
#
# Multiplying the likelihood with a prior distribution $p(\mathbf{w})$ is, by Bayes theorem, proportional to the posterior distribution $p(\mathbf{w} \lvert \mathcal{D}) \propto p(\mathcal{D} \lvert \mathbf{w}) p(\mathbf{w})$. Maximizing $p(\mathcal{D} \lvert \mathbf{w}) p(\mathbf{w})$ gives the maximum a posteriori (MAP) estimate of $\mathbf{w}$. Computing the MAP estimate has a regularizing effect and can prevent overfitting. The optimization objectives here are the same as for MLE plus a regularization term coming from the log prior.
#
# Both MLE and MAP give point estimates of parameters. If we instead had a full posterior distribution over parameters we could make predictions that take weight uncertainty into account. This is covered by the posterior predictive distribution $p(y \lvert \mathbf{x},\mathcal{D}) = \int p(y \lvert \mathbf{x}, \mathbf{w}) p(\mathbf{w} \lvert \mathcal{D}) d\mathbf{w}$ in which the parameters have been marginalized out. This is equivalent to averaging predictions from an ensemble of neural networks weighted by the posterior probabilities of their parameters $\mathbf{w}$.
#
# ## Variational inference
#
# Unfortunately, an analytical solution for the posterior $p(\mathbf{w} \lvert \mathcal{D})$ in neural networks is untractable. We therefore have to approximate the true posterior with a variational distribution $q(\mathbf{w} \lvert \boldsymbol{\theta})$ of known functional form whose parameters we want to estimate. This can be done by minimizing the [Kullback-Leibler divergence](https://en.wikipedia.org/wiki/Kullback%E2%80%93Leibler_divergence) between $q(\mathbf{w} \lvert \boldsymbol{\theta})$ and the true posterior $p(\mathbf{w} \lvert \mathcal{D})$. As shown in [Appendix](#Appendix), the corresponding optimization objective or cost function is
#
# $$
# \mathcal{F}(\mathcal{D},\boldsymbol{\theta}) =
# \mathrm{KL}(q(\mathbf{w} \lvert \boldsymbol{\theta}) \mid\mid p(\mathbf{w})) -
# \mathbb{E}_{q(\mathbf{w} \lvert \boldsymbol{\theta})} \log p(\mathcal{D} \lvert \mathbf{w})
# \tag{1}
# $$
#
# This is known as the *variational free energy*. The first term is the Kullback-Leibler divergence between the variational distribution $q(\mathbf{w} \lvert \boldsymbol{\theta})$ and the prior $p(\mathbf{w})$ and is called the *complexity cost*. The second term is the expected value of the likelihood w.r.t. the variational distribution and is called the *likelihood cost*. By re-arranging the KL term, the cost function can also be written as
#
# $$
# \mathcal{F}(\mathcal{D},\boldsymbol{\theta}) =
# \mathbb{E}_{q(\mathbf{w} \lvert \boldsymbol{\theta})} \log q(\mathbf{w} \lvert \boldsymbol{\theta}) -
# \mathbb{E}_{q(\mathbf{w} \lvert \boldsymbol{\theta})} \log p(\mathbf{w}) -
# \mathbb{E}_{q(\mathbf{w} \lvert \boldsymbol{\theta})} \log p(\mathcal{D} \lvert \mathbf{w})
# \tag{2}
# $$
#
# We see that all three terms in equation $2$ are expectations w.r.t. the variational distribution $q(\mathbf{w} \lvert \boldsymbol{\theta})$. The cost function can therefore be approximated by drawing samples $\mathbf{w}^{(i)}$ from $q(\mathbf{w} \lvert \boldsymbol{\theta})$.
#
# $$
# \mathcal{F}(\mathcal{D},\boldsymbol{\theta}) \approx {1 \over N} \sum_{i=1}^N \left[
# \log q(\mathbf{w}^{(i)} \lvert \boldsymbol{\theta}) -
# \log p(\mathbf{w}^{(i)}) -
# \log p(\mathcal{D} \lvert \mathbf{w}^{(i)})\right]
# \tag{3}
# $$
#
#
# In the following example, we'll use a Gaussian distribution for the variational posterior, parameterized by $\boldsymbol{\theta} = (\boldsymbol{\mu}, \boldsymbol{\sigma})$ where $\boldsymbol{\mu}$ is the mean vector of the distribution and $\boldsymbol{\sigma}$ the standard deviation vector. The elements of $\boldsymbol{\sigma}$ are the elements of a diagonal covariance matrix which means that weights are assumed to be uncorrelated. Instead of parameterizing the neural network with weights $\mathbf{w}$ directly we parameterize it with $\boldsymbol{\mu}$ and $\boldsymbol{\sigma}$ and therefore double the number of parameters compared to a plain neural network.
#
# ## Network training
#
# A training iteration consists of a forward-pass and and backward-pass. During a forward pass a single sample is drawn from the variational posterior distribution. It is used to evaluate the approximate cost function defined by equation $3$. The first two terms of the cost function are data-independent and can be evaluated layer-wise, the last term is data-dependent and is evaluated at the end of the forward-pass. During a backward-pass, gradients of $\boldsymbol{\mu}$ and $\boldsymbol{\sigma}$ are calculated via backpropagation so that their values can be updated by an optimizer.
#
# Since a forward pass involves a stochastic sampling step we have to apply the so-called *re-parameterization trick* for backpropagation to work. The trick is to sample from a parameter-free distribution and then transform the sampled $\boldsymbol{\epsilon}$ with a deterministic function $t(\boldsymbol{\mu}, \boldsymbol{\sigma}, \boldsymbol{\epsilon})$ for which a gradient can be defined. Here, $\boldsymbol{\epsilon}$ is drawn from a standard normal distribution i.e. $\boldsymbol{\epsilon} \sim \mathcal{N}(\mathbf{0}, \mathbf{I})$ and function $t(\boldsymbol{\mu}, \boldsymbol{\sigma}, \boldsymbol{\epsilon}) = \boldsymbol{\mu} + \boldsymbol{\sigma} \odot \boldsymbol{\epsilon}$ shifts the sample by mean $\boldsymbol{\mu}$ and scales it with $\boldsymbol{\sigma}$ where $\odot$ is element-wise multiplication.
#
# For numeric stability we will parameterize the network with $\boldsymbol{\rho}$ instead of $\boldsymbol{\sigma}$ directly and transform $\boldsymbol{\rho}$ with the softplus function to obtain $\boldsymbol{\sigma} = \log(1 + \exp(\boldsymbol{\rho}))$. This ensures that $\boldsymbol{\sigma}$ is always positive. As prior, a scale mixture of two Gaussians is used $p(\mathbf{w}) = \pi \mathcal{N}(\mathbf{w} \lvert 0,\sigma_1^2) + (1 - \pi) \mathcal{N}(\mathbf{w} \lvert 0,\sigma_2^2)$ where $\sigma_1$, $\sigma_2$ and $\pi$ are hyper-parameters i.e. they are not learned during training.
#
# ## Uncertainty characterization
#
# Uncertainty in predictions that arise from the uncertainty in weights is called [epistemic uncertainty](https://en.wikipedia.org/wiki/Uncertainty_quantification). This kind of uncertainty can be reduced if we get more data. Consequently, epistemic uncertainty is higher in regions of no or little training data and lower in regions of more training data. Epistemic uncertainty is covered by the variational posterior distribution. Uncertainty coming from the inherent noise in training data is an example of [aleatoric uncertainty](https://en.wikipedia.org/wiki/Uncertainty_quantification). It cannot be reduced if we get more data. Aleatoric uncertainty is covered by the probability distribution used to define the likelihood function.
#
# ## Implementation example
#
# Variational inference of neural network parameters is now demonstrated on a simple regression problem. We therefore use a Gaussian distribution for $p(y \lvert \mathbf{x},\mathbf{w})$. The training dataset consists of 32 noisy samples `X`, `y` drawn from a sinusoidal function.
# +
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
def f(x, sigma):
epsilon = np.random.randn(*x.shape) * sigma
return 10 * np.sin(2 * np.pi * (x)) + epsilon
train_size = 32
noise = 1.0
X = np.linspace(-0.5, 0.5, train_size).reshape(-1, 1)
y = f(X, sigma=noise)
y_true = f(X, sigma=0.0)
plt.scatter(X, y, marker='+', label='Training data')
plt.plot(X, y_true, label='Truth')
plt.title('Noisy training data and ground truth')
plt.legend();
# -
# The noise in training data gives rise to aleatoric uncertainty. To cover epistemic uncertainty we implement the variational inference logic in a custom `DenseVariational` Keras layer. The complexity cost (`kl_loss`) is computed layer-wise and added to the total loss with the `add_loss` method. Implementations of `build` and `call` directly follow the equations defined above.
# +
from keras import backend as K
from keras import activations, initializers
from keras.layers import Layer
import tensorflow as tf
import tensorflow_probability as tfp
class DenseVariational(Layer):
def __init__(self,
units,
kl_weight,
activation=None,
prior_sigma_1=1.5,
prior_sigma_2=0.1,
prior_pi=0.5, **kwargs):
self.units = units
self.kl_weight = kl_weight
self.activation = activations.get(activation)
self.prior_sigma_1 = prior_sigma_1
self.prior_sigma_2 = prior_sigma_2
self.prior_pi_1 = prior_pi
self.prior_pi_2 = 1.0 - prior_pi
self.init_sigma = np.sqrt(self.prior_pi_1 * self.prior_sigma_1 ** 2 +
self.prior_pi_2 * self.prior_sigma_2 ** 2)
super().__init__(**kwargs)
def compute_output_shape(self, input_shape):
return input_shape[0], self.units
def build(self, input_shape):
self.kernel_mu = self.add_weight(name='kernel_mu',
shape=(input_shape[1], self.units),
initializer=initializers.normal(stddev=self.init_sigma),
trainable=True)
self.bias_mu = self.add_weight(name='bias_mu',
shape=(self.units,),
initializer=initializers.normal(stddev=self.init_sigma),
trainable=True)
self.kernel_rho = self.add_weight(name='kernel_rho',
shape=(input_shape[1], self.units),
initializer=initializers.constant(0.0),
trainable=True)
self.bias_rho = self.add_weight(name='bias_rho',
shape=(self.units,),
initializer=initializers.constant(0.0),
trainable=True)
super().build(input_shape)
def call(self, inputs, **kwargs):
kernel_sigma = tf.math.softplus(self.kernel_rho)
kernel = self.kernel_mu + kernel_sigma * tf.random.normal(self.kernel_mu.shape)
bias_sigma = tf.math.softplus(self.bias_rho)
bias = self.bias_mu + bias_sigma * tf.random.normal(self.bias_mu.shape)
self.add_loss(self.kl_loss(kernel, self.kernel_mu, kernel_sigma) +
self.kl_loss(bias, self.bias_mu, bias_sigma))
return self.activation(K.dot(inputs, kernel) + bias)
def kl_loss(self, w, mu, sigma):
variational_dist = tfp.distributions.Normal(mu, sigma)
return self.kl_weight * K.sum(variational_dist.log_prob(w) - self.log_prior_prob(w))
def log_prior_prob(self, w):
comp_1_dist = tfp.distributions.Normal(0.0, self.prior_sigma_1)
comp_2_dist = tfp.distributions.Normal(0.0, self.prior_sigma_2)
return K.log(self.prior_pi_1 * comp_1_dist.prob(w) +
self.prior_pi_2 * comp_2_dist.prob(w))
# -
# Our model is a neural network with two `DenseVariational` hidden layers, each having 20 units, and one `DenseVariational` output layer with one unit. Instead of modeling a full probability distribution $p(y \lvert \mathbf{x},\mathbf{w})$ as output the network simply outputs the mean of the corresponding Gaussian distribution. In other words, we do not model aleatoric uncertainty here and assume it is known. We only model epistemic uncertainty via the `DenseVariational` layers.
#
# Since the training dataset has only 32 examples we train the network with all 32 examples per epoch so that the number of batches per epoch is 1. For other configurations, the complexity cost (`kl_loss`) must be weighted by $1/M$ as described in section 3.4 of the paper where $M$ is the number of mini-batches per epoch. The hyper-parameter values for the mixture prior, `prior_params`, have been chosen to work well for this example and may need adjustments in another context.
# +
import warnings
warnings.filterwarnings('ignore')
from keras.layers import Input
from keras.models import Model
batch_size = train_size
num_batches = train_size / batch_size
kl_weight = 1.0 / num_batches
prior_params = {
'prior_sigma_1': 1.5,
'prior_sigma_2': 0.1,
'prior_pi': 0.5
}
x_in = Input(shape=(1,))
x = DenseVariational(20, kl_weight, **prior_params, activation='relu')(x_in)
x = DenseVariational(20, kl_weight, **prior_params, activation='relu')(x)
x = DenseVariational(1, kl_weight, **prior_params)(x)
model = Model(x_in, x)
# -
# The network can now be trained with a Gaussian negative log likelihood function (`neg_log_likelihood`) as loss function assuming a fixed standard deviation (`noise`). This corresponds to the *likelihood cost*, the last term in equation $3$.
# +
from keras import callbacks, optimizers
def neg_log_likelihood(y_obs, y_pred, sigma=noise):
dist = tfp.distributions.Normal(loc=y_pred, scale=sigma)
return K.sum(-dist.log_prob(y_obs))
model.compile(loss=neg_log_likelihood, optimizer=optimizers.Adam(lr=0.08), metrics=['mse'])
model.fit(X, y, batch_size=batch_size, epochs=1500, verbose=0);
# -
# When calling `model.predict` we draw a random sample from the variational posterior distribution and use it to compute the output value of the network. This is equivalent to obtaining the output from a single member of a hypothetical ensemble of neural networks. Drawing 500 samples means that we get predictions from 500 ensemble members. From these predictions we can compute statistics such as the mean and standard deviation. In our example, the standard deviation is a measure of epistemic uncertainty.
# +
import tqdm
X_test = np.linspace(-1.5, 1.5, 1000).reshape(-1, 1)
y_pred_list = []
for i in tqdm.tqdm(range(500)):
y_pred = model.predict(X_test)
y_pred_list.append(y_pred)
y_preds = np.concatenate(y_pred_list, axis=1)
y_mean = np.mean(y_preds, axis=1)
y_sigma = np.std(y_preds, axis=1)
plt.plot(X_test, y_mean, 'r-', label='Predictive mean');
plt.scatter(X, y, marker='+', label='Training data')
plt.fill_between(X_test.ravel(),
y_mean + 2 * y_sigma,
y_mean - 2 * y_sigma,
alpha=0.5, label='Epistemic uncertainty')
plt.title('Prediction')
plt.legend();
# -
# We can clearly see that epistemic uncertainty is much higher in regions of no training data than it is in regions of existing training data. The predictive mean could have also been obtained with a single forward pass i.e. a single `model.predict` call by using only the mean of the variational posterior distribution which is equivalent to sampling from the variational posterior with $\boldsymbol{\sigma}$ set to $\mathbf{0}$. The corresponding implementation is omitted here but is trivial to add.
#
# For an example how to model both epistemic and aleatoric uncertainty I recommend reading [Regression with Probabilistic Layers in TensorFlow Probability](https://medium.com/tensorflow/regression-with-probabilistic-layers-in-tensorflow-probability-e46ff5d37baf) which uses probabilistic Keras layers from the upcoming Tensorflow Probability 0.7.0 release. Their approach to variational inference is similar to the approach described here but differs in some details. For example, they compute the complexity cost analytically instead of estimating it from samples, among other differences.
# ## Appendix
#
# ### Optimization objective
#
# The KL divergence between the variational distribution $q(\mathbf{w} \lvert \boldsymbol{\theta})$ and the true posterior $p(\mathbf{w} \lvert \mathcal{D})$ is defined as
#
# $$
# \begin{align*}
# \mathrm{KL}(q(\mathbf{w} \lvert \boldsymbol{\theta}) \mid\mid p(\mathbf{w} \lvert \mathcal{D})) &=
# \int{q(\mathbf{w} \lvert \boldsymbol{\theta}) \log
# {{q(\mathbf{w} \lvert \boldsymbol{\theta})} \over {p(\mathbf{w} \lvert \mathcal{D})}} d\mathbf{w}} \\\\ &=
# \mathbb{E}_{q(\mathbf{w} \lvert \boldsymbol{\theta})} \log
# {{q(\mathbf{w} \lvert \boldsymbol{\theta})} \over {p(\mathbf{w} \lvert \mathcal{D})}}
# \end{align*}
# $$
#
# Applying Bayes' rule to $p(\mathbf{w} \lvert \mathcal{D})$ we obtain
#
# $$
# \begin{align*}
# \mathrm{KL}(q(\mathbf{w} \lvert \boldsymbol{\theta}) \mid\mid p(\mathbf{w} \lvert \mathcal{D})) &=
# \mathbb{E}_{q(\mathbf{w} \lvert \boldsymbol{\theta})} \log
# {{q(\mathbf{w} \lvert \boldsymbol{\theta})} \over
# {p(\mathcal{D} \lvert \mathbf{w}) p(\mathbf{w})}} p(\mathcal{D}) \\\\ &=
# \mathbb{E}_{q(\mathbf{w} \lvert \boldsymbol{\theta})} \left[
# \log q(\mathbf{w} \lvert \boldsymbol{\theta}) -
# \log p(\mathcal{D} \lvert \mathbf{w}) -
# \log p(\mathbf{w}) +
# \log p(\mathcal{D})
# \right] \\\\ &=
# \mathbb{E}_{q(\mathbf{w} \lvert \boldsymbol{\theta})} \left[
# \log q(\mathbf{w} \lvert \boldsymbol{\theta}) -
# \log p(\mathcal{D} \lvert \mathbf{w}) -
# \log p(\mathbf{w})
# \right] + \log p(\mathcal{D}) \\\\ &=
# \mathrm{KL}(q(\mathbf{w} \lvert \boldsymbol{\theta}) \mid\mid p(\mathbf{w})) -
# \mathbb{E}_{q(\mathbf{w} \lvert \boldsymbol{\theta})} \log p(\mathcal{D} \lvert \mathbf{w}) +
# \log p(\mathcal{D})
# \end{align*}
# $$
#
# using the fact that the *log marginal likelihood* $\log p(\mathcal{D})$ doesn't depend on $\mathbf{w}$. The first two terms on the RHS are the *variational free energy* $\mathcal{F}(\mathcal{D},\boldsymbol{\theta})$ as defined in Eq. $(1)$. We obtain
#
# $$
# \mathrm{KL}(q(\mathbf{w} \lvert \boldsymbol{\theta}) \mid\mid p(\mathbf{w} \lvert \mathcal{D})) =
# \mathcal{F}(\mathcal{D},\boldsymbol{\theta}) + \log p(\mathcal{D})
# $$
#
# In order to minimize $\mathrm{KL}(q(\mathbf{w} \lvert \boldsymbol{\theta}) \mid\mid p(\mathbf{w} \lvert \mathcal{D}))$, we only need to minimize $\mathcal{F}(\mathcal{D},\boldsymbol{\theta})$ w.r.t. $\boldsymbol{\theta}$ as $p(\mathcal{D})$ doesn't depend on $\boldsymbol{\theta}$. The negative variational free energy is also known as *evidence lower bound* $\mathcal{L}(\mathcal{D},\boldsymbol{\theta})$ (ELBO).
#
# $$
# \mathrm{KL}(q(\mathbf{w} \lvert \boldsymbol{\theta}) \mid\mid p(\mathbf{w} \lvert \mathcal{D})) =
# -\mathcal{L}(\mathcal{D},\boldsymbol{\theta}) + \log p(\mathcal{D})
# $$
#
# It is a lower bound on $\log p(\mathcal{D})$ because the Kullback-Leibler divergence is always non-negative.
#
# $$
# \begin{align*}
# \mathcal{L}(\mathcal{D},\boldsymbol{\theta}) &=
# \log p(\mathcal{D}) - \mathrm{KL}(q(\mathbf{w} \lvert \boldsymbol{\theta}) \mid\mid p(\mathbf{w} \lvert \mathcal{D})) \\\\
# \mathcal{L}(\mathcal{D},\boldsymbol{\theta}) &\leq
# \log p(\mathcal{D})
# \end{align*}
# $$
#
# Therefore, the KL divergence between the variational distribution $q(\mathbf{w} \lvert \boldsymbol{\theta})$ and the true posterior $p(\mathbf{w} \lvert \mathcal{D})$ is also minimized by maximizing the evidence lower bound.
| bayesian_neural_networks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # ALeRCE classes
#
# https://github.com/ZwickyTransientFacility/ztf-avro-alert
#
# 1. **AGN:** Active Galactic Nuclei
# 1. **Blazar:** Blazar
# 1. **CV/Nova:** Cataclysmic Variable Star/Nova
# 1. **Ceph: Cepheid Variable Star**
# 1. **DSCT: Delta Scuti Star**
# 1. **EA:** Eclipsing Algol
# 1. **EB/EW: Eclipsing Binaries/Eclipsing W Ursa Majoris**
# 1. **LPV: Long Period Variable**
# 1. **Periodic-Other:** Periodic-Other
# 1. **QSO:** Quasi-Stellar Object
# 1. **RRL: RRLyrae Variable Star**
# 1. **RSCVn:** RS Canum Venaticorum
# 1. **SLSN:** Super Luminous Supernova
# 1. **SNII:** Supernova II
# 1. **SNIIb:** Supernova IIb
# 1. **SNIIn:** Supernova IIn
# 1. **SNIa:** Supernova Ia
# 1. **SNIbc:** Supernova Ibc
# 1. **TDE:** Tidal disruption event (to remove)
# 1. **YSO:** Young Stellar Object
# 1. **ZZ:** ZZ Ceti Stars (to remove)
# +
import numpy as np
import pandas as pd
def subset_df_columns(df, subset_cols):
df_cols = list(df.columns)
return df[[c for c in subset_cols if c in df_cols]]
def set_index(df, index_name):
if not df.index.name is None and df.index.name==index_name:
return df
df_cols = list(df.columns)
assert index_name in df_cols
return df.set_index([index_name])
def df_to_float32(df):
for c in df.columns:
if df[c].dtype=='float64':
df[c] = df[c].astype(np.float32)
# +
import numpy as np
import pandas as pd
#survey_name = 'alerceZTF_v5.1'
survey_name = 'alerceZTF_v7.1' # use this dataset
df_index_names = {
'oid':'oid', # object id
'label':'classALeRCE', # object class name
'ra':'ra',
'dec':'dec',
'band':'fid', # band
'obs_day':'mjd', # days
'obs':'magpsf_corr', # observations
'obs_error':'sigmapsf_corr', # observation errors
}
### load files
load_root_dir = f'data/{survey_name}'
labels_df = pd.read_csv(f'{load_root_dir}/labels_vs.csv')
print(f'labels_df - columns: {list(labels_df.columns)} - id: {labels_df.index.name}')
labels_df = set_index(labels_df, df_index_names['oid'])
detections_df = pd.read_parquet(f'{load_root_dir}/detections_vs.parquet')
print(f'detections_df - columns: {list(detections_df.columns)} - id: {detections_df.index.name}')
detections_df = set_index(detections_df, df_index_names['oid'])
df_to_float32(detections_df)
# -
print(labels_df.info())
labels_df[:20]
print(detections_df.info())
detections_df[:20]
example_oid = 'ZTF18aaakigd'
print(labels_df.loc[example_oid])
print(detections_df.loc[example_oid])
| .ipynb_checkpoints/load_data-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python36
# ---
# # Create and connect to an Azure Machine Learning Workspace
#
# Run the following cell to create a new Azure Machine Learning **Workspace**.
#
# **Important Note**: You will be prompted to login in the text that is output below the cell. Be sure to navigate to the URL displayed and enter the code that is provided. Once you have entered the code, return to this notebook and wait for the output to read `Workspace configuration succeeded`.
# +
from azureml.core import Workspace, Experiment, Run
ws = Workspace.from_config()
print('Workspace configuration succeeded')
# -
# # Get the Model Training Run
#
# **Load the run_info.json file that has the run id for the model training run**
# +
import os
import json
output_path = './outputs'
run_info_filepath = os.path.join(output_path, 'run_info.json')
try:
with open(run_info_filepath) as f:
run_info = json.load(f)
print('run_info.json loaded')
print(run_info)
except:
print("Cannot open: ", run_info_filepath)
print("Please fix output_path before proceeding!")
# -
# **Get the Run object from the run id**
experiment_name = 'deep-learning'
run = Run(Experiment(ws, experiment_name), run_info['id'])
# # Register Model
# **Register the Model with Azure Model Registry**
# +
model_name = 'compliance-classifier'
model_description = 'Deep learning model to classify the descriptions of car components as compliant or non-compliant.'
model_run = run.register_model(model_name=model_name,
model_path="outputs/model/model.h5",
tags={"type": "classification", "description": model_description, "run_id": run.id})
print("Model Registered: {} \nModel Description: {} \nModel Version: {}".format(model_run.name,
model_run.tags["description"], model_run.version))
# -
| Hands-on lab/notebooks/Register Model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Day 17
#
# The elves bought too much eggnog again - 150 liters this time. To fit it all into your refrigerator, you'll need to move it into smaller containers. You take an inventory of the capacities of the available containers.
#
# For example, suppose you have containers of size 20, 15, 10, 5, and 5 liters. If you need to store 25 liters, there are four ways to do it:
#
# 15 and 10
# 20 and 5 (the first 5)
# 20 and 5 (the second 5)
# 15, 5, and 5
#
# Filling all containers entirely, how many different combinations of containers can exactly fit all 150 liters of eggnog?
# ## Puzzle 1
# +
from itertools import combinations
from typing import List
def find_combinations(containers:List[int], total:int):
"""Return combinations of containers whose sizes sum to the total
:param containers: list of container sizes
:param total: total size to be contained
"""
valid = []
for count in range(1, len(containers) + 1):
valid.extend([_ for _ in combinations(containers, count) if sum(_) == total])
return valid
# +
containers = (20, 15, 10, 5, 5)
total = 25
find_combinations(containers, total)
# -
# ### Solution
# +
total = 150
with open("day17.txt", "r") as ifh:
containers = [int(_.strip()) for _ in ifh.readlines()]
valid = find_combinations(containers, total)
len(valid)
# -
# ## Puzzle 2
#
# While playing with all the containers in the kitchen, another load of eggnog arrives! The shipping and receiving department is requesting as many containers as you can spare.
#
# Find the minimum number of containers that can exactly fit all 150 liters of eggnog. How many different ways can you fill that number of containers and still hold exactly 150 litres?
#
# In the example above, the minimum number of containers was two. There were three ways to use that many containers, and so the answer there would be 3.
valid_sizes = [(len(_), _) for _ in valid]
len([_ for _ in valid_sizes if _[0] == 4])
| day17.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
from IPython.display import Image
# %matplotlib inline
# # Exploratory Data Analysis with Seaborn and Pandas
# ## Seaborn
#
# Seaborn is a library for making attractive and informative statistical graphics in Python. It is built on top of matplotlib and tightly integrated with the Python Data Science stack, including support for numpy and pandas data structures and statistical routines from scipy and statsmodels.
#
# Some of the features that seaborn offers are
#
# * Several built-in themes that improve on the default matplotlib aesthetics
# * Tools for choosing color palettes to make beautiful plots that reveal patterns in your data
# * Functions for visualizing univariate and bivariate distributions or for comparing them between subsets of data
# * Tools that fit and visualize linear regression models for different kinds of independent and dependent variables
# * Functions that visualize matrices of data and use clustering algorithms to discover structure in those matrices
# * A function to plot statistical timeseries data with flexible estimation and representation of uncertainty around the estimate
# * High-level abstractions for structuring grids of plots that let you easily build complex visualizations
#
# Seaborn aims to make visualization a central part of exploring and understanding data. The plotting functions operate on dataframes and arrays containing a whole dataset and internally perform the necessary aggregation and statistical model-fitting to produce informative plots. If matplotlib “tries to make easy things easy and hard things possible”, seaborn tries to make a well-defined set of hard things easy too.
#
# Seaborn should be thought of as a complement to matplotlib, not a replacement for it. When using seaborn, it is likely that you will often invoke matplotlib functions directly to draw simpler plots already available through the pyplot namespace. Further, while the seaborn functions aim to make plots that are reasonably “production ready” (including extracting semantic information from Pandas objects to add informative labels), full customization of the figures will require a sophisticated understanding of matplotlib objects.
#
# standard import statement for seaborn
import seaborn as sns
# ## What is Exploratory Data Analysis (EDA)
#
# Exploratory data analysis is an approach to analyzing data sets to summarize their main characteristics, often with visual methods. It is used to understand the data, get context about it, understand the variables and the relashionship between them, and formulate hypotheses that could be useful when building predictive models.
# # Doing Exploratory Data Analysis
# ## Loading the data
housing = pd.read_csv('house_train.csv')
housing.shape
housing.info()
# ## The questions that will guide our analysis
#
# All data analysis must be guided by some key questions or objectives that conduct all that we do. As example here are the two objectives that will guide our exploration of the data set:
#
# > **1. Understand the individual variables in the dataset**
#
# > **2. Understand how the variables in this dataset relate with the SalePrice of the house**
# **Understand the data and the problem**: the more context you have, the better. It is a good idea to take a look at each variable and think about their meaning and importance for this problem.
# For each variable in the dataset you should know it's **Type**. In general there are two possible types of variables: 'numerical' or 'categorical'. By 'numerical' we mean variables for which the values are numbers, and by 'categorical' we mean variables for which the values are categories.
# +
#Image("img/variable_types.png", width=500)
# -
# **Numerical variables**
#
# - **SalePrice**
# - **LotArea:** Lot size in square feet
# - **OverallQual:** Rates the overall material and finish of the house
# - **OverallCond:** Rates the overall condition of the house
# - **1stFlrSF:** First Floor square feet
# - **2ndFlrSF:** Second floor square feet
# - **BedroomAbvGr:** Bedrooms above grade (does NOT include basement bedrooms)
# - **YearBuilt:** Original construction date (this is not technically a numeric variable but we will use it to produce another variable called Age)
#
# **Categorical variables**
#
# - **MSZoning:** Identifies the general zoning classification of the sale.
# - **LotShape:** General shape of property
# - **Neighborhood:** Physical locations within Ames city limits
# - **CentralAir:** Central air conditioning
# - **SaleCondition:** Condition of sale
# - **MoSold:** Month Sold (MM)
# - **YrSold:** Year Sold (YYYY)
numerical_vars = ['SalePrice','LotArea', 'OverallQual', 'OverallCond',
'YearBuilt', '1stFlrSF', '2ndFlrSF', 'BedroomAbvGr']
categorical_vars = ['MSZoning', 'LotShape', 'Neighborhood', 'CentralAir', 'SaleCondition', 'MoSold', 'YrSold']
housing = housing[numerical_vars+categorical_vars]
housing.shape
# ## Understanding the main variable
#descriptive statistics summary
housing['SalePrice'].describe()
housing['SalePrice'].hist(edgecolor='black', bins=20);
#skewness and kurtosis
print("Skewness: {:0.3f}".format(housing['SalePrice'].skew()))
print("Kurtosis: {:0.3f}".format(housing['SalePrice'].kurt()))
# ## Numerical variables
housing[numerical_vars].describe()
# ## Relationships between numerical variables
#
# The seaborn library excels when we want to investigate relationship between variables, with very few lines of code we can get very informative plots and discover patterns and relationships between our variables.
housing.plot.scatter(x='1stFlrSF', y='SalePrice');
housing[numerical_vars].corr()
housing[numerical_vars].corr()['SalePrice'].sort_values(ascending=False)
| 09-Sep-2018/EDA 1/EDA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/ava6700/tensorflow-101/blob/master/rps_game.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="CUXVXcbe6-5M"
# + id="wXjuJfkQ7ZFu"
# + colab={"base_uri": "https://localhost:8080/"} id="Hs7hioxo7m8m" outputId="de16f10d-23e3-4b01-8c69-4860140fd8d7"
# !apt-get install -y xvfb # Install X Virtual Frame Buffer
import os
os.system('Xvfb :1 -screen 0 1600x1200x16 &') # create virtual display with size 1600x1200 and 16 bit color. Color can be changed to 24 or 8
os.environ['DISPLAY']=':1.0' # tell X clients to use our virtual DISPLAY :1.0
# + id="i7QFhuLvCbHn"
# + id="v1xjht_38FoQ"
# + [markdown] id="NCqZTZ9R8tiC"
# # Import libraries
# + colab={"base_uri": "https://localhost:8080/"} id="ha2LX8l42g4r" outputId="3ea1c33d-87fa-45df-e38f-be8cbab53f79"
# !pip install win32gui
# + id="tSnnDXfu8LPy"
from tkinter import*
import random
# + id="iNQRXigH8LSV"
# + [markdown] id="hUyz6Sxn9L8a"
# # initialize window:
# + id="sPr346R5UrYP"
# + colab={"base_uri": "https://localhost:8080/", "height": 380} id="b2esuCfL8LUw" outputId="05c0c555-39a0-4317-b04e-2ff898a5e2d9"
game = Tk()#use to initialize TKinter to create windows
game.geometry("400x200")#sets the windows heightand weight
game.resizable(0,0)#sie of the window
game.title('Rock paper Scissor')#title of the window
game.config(bg = '#856ff8')#backgorund color
# + id="DLNwblc_HTOs"
Label(game, text = 'Rock, Paper ,Scissors' , font='arial 20 bold', bg = 'seashell2').pack()
# + id="Kj6O36dkHpJG"
# + [markdown] id="zGKN0yifHqbP"
# # For user choice
# + id="GG6WPz6n8LXb"
user_take =StringVar()#stores the choice that the users enters
Label(game, text = 'What you prefer? rock, paper, scissor',
font='arial 16 bold', bg = 'green').place(x = 20, y = 70)
Entry(game, font = 'arial 15', textvariable = user_take , bg = 'antiquewhite2').place(x=90 , y = 130)
# + id="n62QwNWG8LZ_"
# + [markdown] id="8TYgVSmKHxIm"
# #For computer choice
# + id="GWiFbXwJ8LdW"
comp_pick = random.randint(1,3)#random.randint function will randomly take any nuber from the given number
if comp_pick == 1:
comp_pick = 'rock'
elif comp_pick == 2:
comp_pick = 'paper'
else:
comp_pick =='scissor'
# + id="0vsPeUu48JOI"
# + [markdown] id="smHlUNUyI2nk"
# # Starting game:
#
# + id="PlGUA_l2I7bK"
Result = StringVar()
def game_to_play():
user_pick = user_take.get()
if user_pick == comp_pick:
Result.set('Try Again,you both pick same')
elif user_pick =='rock'and comp_pick =='paper':
Result.set('ohho! you lose,computer win')
elif user_pick == 'rock' and comp_pick == 'scissors':
Result.set('you win,computer select scissors')
elif user_pick == 'paper' and comp_pick == 'scissors':
Result.set('ohho!you lose,computer select scissors')
elif user_pick == 'paper' and comp_pick == 'rock':
Result.set('you win,computer select rock')
elif user_pick == 'scissors' and comp_pick == 'rock':
Result.set('ohho! you lose,computer select rock')
elif user_pick == 'scissors' and comp_pick == 'paper':
Result.set('you win ,computer select paper')
else:
Result.set('invalid: choose any one -- rock, paper, scissors')
# + id="ctDZA3i5OCFA"
# + [markdown] id="d0lUlZYKOaU6"
# # Function to reset
# + id="tC9FhPiPOjcE"
def Reset():
Result.set("")
user_take.set("")
# + id="2EkLzuMaOxdN"
# + [markdown] id="cGIjjwNcOy5P"
# # End Game
# + id="_Aney_NHO2mP"
def Exit():
game.destroy()#game.destroy() will quit the game by stopping the mainloop()
# + id="yxoS7EzUPC3u"
# + [markdown] id="plqk6sicPP3k"
# # Game Buttons
# + colab={"base_uri": "https://localhost:8080/", "height": 296} id="yRrQ4LK0Pb-Q" outputId="5ba28c50-92fc-499b-85bf-b4c00d86eb9b"
Entry(game, font = 'arial 10 bold', textvariable = Result, bg ='antiquewhite2',width = 50,).place(x=25, y = 250)
Button(game, font = 'arial 13 bold', text = 'PLAY' ,padx =5,bg ='seashell4' ,command = game_to_play).place(x=150,y=190)
Button(game, font = 'arial 13 bold', text = 'RESET' ,padx =5,bg ='seashell4' ,command = Reset).place(x=70,y=310)
Button(game, font = 'arial 13 bold', text = 'EXIT' ,padx =5,bg ='seashell4' ,command = Exit).place(x=230,y=310)
game.mainloop()
# + id="MTWmjAs6PiuU"
| rps_game.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "slide"}
# # Title
#
# _Brief abstract/introduction/motivation. State what the chapter is about in 1-2 paragraphs._
# _Then, have an introduction video:_
# + slideshow={"slide_type": "skip"}
from bookutils import YouTubeVideo
YouTubeVideo("w4u5gCgPlmg")
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"}
# **Prerequisites**
#
# * _Refer to earlier chapters as notebooks here, as here:_ [Earlier Chapter](Debugger.ipynb).
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "skip"}
import bookutils
# + [markdown] slideshow={"slide_type": "slide"}
# ## Synopsis
# <!-- Automatically generated. Do not edit. -->
#
# To [use the code provided in this chapter](Importing.ipynb), write
#
# ```python
# >>> from debuggingbook.Template import <identifier>
# ```
#
# and then make use of the following features.
#
#
# _For those only interested in using the code in this chapter (without wanting to know how it works), give an example. This will be copied to the beginning of the chapter (before the first section) as text with rendered input and output._
#
# You can use `int_fuzzer()` as:
#
# ```python
# >>> print(int_fuzzer())
# 76.5
#
# ```
#
# + [markdown] button=false new_sheet=true run_control={"read_only": false} slideshow={"slide_type": "slide"}
# ## _Section 1_
#
# \todo{Add}
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "slide"}
# ## _Section 2_
#
# \todo{Add}
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Excursion: All the Details
# + [markdown] slideshow={"slide_type": "fragment"}
# This text will only show up on demand (HTML) or not at all (PDF). This is useful for longer implementations, or repetitive, or specialized parts.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### End of Excursion
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"}
# ## _Section 3_
#
# \todo{Add}
#
# _If you want to introduce code, it is helpful to state the most important functions, as in:_
#
# * `random.randrange(start, end)` - return a random number [`start`, `end`]
# * `range(start, end)` - create a list with integers from `start` to `end`. Typically used in iterations.
# * `for elem in list: body` executes `body` in a loop with `elem` taking each value from `list`.
# * `for i in range(start, end): body` executes `body` in a loop with `i` from `start` to `end` - 1.
# * `chr(n)` - return a character with ASCII code `n`
# + slideshow={"slide_type": "skip"}
import random
# + slideshow={"slide_type": "subslide"}
def int_fuzzer() -> float:
"""A simple function that returns a random float"""
return random.randrange(1, 100) + 0.5
# + button=false code_folding=[] new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"}
# More code
pass
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "slide"}
# ## _Section 4_
#
# \todo{Add}
# + [markdown] slideshow={"slide_type": "slide"}
# ## Synopsis
# + [markdown] slideshow={"slide_type": "fragment"}
# _For those only interested in using the code in this chapter (without wanting to know how it works), give an example. This will be copied to the beginning of the chapter (before the first section) as text with rendered input and output._
# + [markdown] slideshow={"slide_type": "fragment"}
# You can use `int_fuzzer()` as:
# + slideshow={"slide_type": "fragment"}
print(int_fuzzer())
# + [markdown] button=false new_sheet=true run_control={"read_only": false} slideshow={"slide_type": "slide"}
# ## Lessons Learned
#
# * _Lesson one_
# * _Lesson two_
# * _Lesson three_
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "slide"}
# ## Next Steps
#
# _Link to subsequent chapters (notebooks) here, as in:_
#
# * [use _assertions_ to check conditions at runtime](Assertions.ipynb)
# * [reduce _failing inputs_ for efficient debugging](DeltaDebugger.ipynb)
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## Background
#
# _Cite relevant works in the literature and put them into context, as in:_
#
# The idea of ensuring that each expansion in the grammar is used at least once goes back to Burkhardt \cite{Burkhardt1967}, to be later rediscovered by Paul Purdom \cite{Purdom1972}.
# + [markdown] button=false new_sheet=true run_control={"read_only": false} slideshow={"slide_type": "slide"}
# ## Exercises
#
# _Close the chapter with a few exercises such that people have things to do. To make the solutions hidden (to be revealed by the user), have them start with_
#
# ```
# **Solution.**
# ```
#
# _Your solution can then extend up to the next title (i.e., any markdown cell starting with `#`)._
#
# _Running `make metadata` will automatically add metadata to the cells such that the cells will be hidden by default, and can be uncovered by the user. The button will be introduced above the solution._
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"}
# ### Exercise 1: _Title_
#
# _Text of the exercise_
# + cell_style="center" slideshow={"slide_type": "fragment"}
# Some code that is part of the exercise
pass
# + [markdown] slideshow={"slide_type": "fragment"} solution2="hidden" solution2_first=true
# _Some more text for the exercise_
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden"
# **Solution.** _Some text for the solution_
# + cell_style="split" slideshow={"slide_type": "skip"} solution2="hidden"
# Some code for the solution
2 + 2
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden"
# _Some more text for the solution_
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"} solution="hidden" solution2="hidden" solution2_first=true solution_first=true
# ### Exercise 2: _Title_
#
# _Text of the exercise_
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "skip"} solution="hidden" solution2="hidden"
# **Solution.** _Solution for the exercise_
| docs/notebooks/Template.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# # Introduction to Node.js
#
# ## Installation of Node.js
#
# [https://nodejs.org](https://nodejs.org/en/)
#
# ## Installation of Integrated Development Environment (IDE) (Visual Studio Code)
#
#
# [https://code.visualstudio.com/](https://code.visualstudio.com/)
#
#
# ## Using the REPL
#
| advance/21.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from jiamtrader.trader.constant import Exchange,Interval
from data_analysis import DataAnalysis
from datetime import datetime
import matplotlib.pyplot as plt
herramiento = DataAnalysis()
herramiento.load_history(
symbol="XBTUSD",
exchange=Exchange.BITMEX,
interval=Interval.MINUTE,
start=datetime(2019, 9, 1),
end=datetime(2019, 10, 30),
rate = 8/10000,
index_3to1 = ["ATR","CCI"],
index_1to1 = ["STDDEV","SMA"],
index_2to2 = ["AROON"],
index_2to1 = ["AROONOSC"],
index_4to1 = ["BOP"],
window_index=30,
)
data = herramiento.base_analysis()
herramiento.show_chart(data[:1500], boll_wide=2.8)
# 多时间周期分析
intervals = ["5min","15min","30min","1h","2h","4h"]
herramiento.multi_time_frame_analysis(intervals=intervals)
| examples/data_analysis/data_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + deletable=true editable=true
import random,math
def numa(m,k):
n = int(input('请输入您想求的数的个数'))
i=0
total=0
if i<=n:
total=total+random.randint(m,k)
i+=1
return total/n
m=int(input('请输入一个整数'))
k=int(input('请输入另一个整数'))
print('n个随机整数均值的平方根是',math.sqrt(numa(m,k)))
# + deletable=true editable=true
import random,math
def xigema(m,k):
n = int(input('请输入您想求的数的个数'))
total1=0
total2=0
i=0
if i<=n:
total1=total1+math.log2(random.randint(m,k))
total2=total2+1/math.log2(random.randint(m,k))
i+=1
return total1,total2
m=int(input('请输入一个整数'))
k=int(input('请输入另一个整数'))
print('西格玛log(随机整数),西格玛1/log(随机整数)分别是',xigema(m,k))
# + deletable=true editable=true
import random,math
def main():
n=int(input('请输入数相加的个数'))
a=math.ceil(random.randint(1,9))
i=0
sum=0
while i<=n:
sum=sum+a
a=a*10+a
i=i+1
print('s=a+aa+aaa+aaaa+aa...a=',sum)
main()
# + deletable=true editable=true
import random, math
def win():
print('Win!')
def lose():
print('Lose!')
def guess_game():
n = int(input('请输入一个大于0的整数,作为神秘整数的上界,回车结束。'))
number = int(input('请输入一个随机的整数。'))
max_times = math.ceil(math.log2(n))
guess_times = 0
while guess_times <= max_times:
guess = random.randint(1, n)
guess_times += 1
print('一共可以猜', max_times, '次')
print('你已经猜了', guess_times, '次')
if guess == number:
win()
print('神秘数字是:', guess)
print('你比标准次数少', max_times-guess_times, '次')
break
elif guess > number:
print('抱歉,你猜大了')
else:
print('抱歉,你猜小了')
else:
print('神秘数字是:', number)
lose()
guess_game()
| chapter2/homework/computer/4-5/201611691031-Copy1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="mmM6yMcdYGFL"
#
# # Transferencia de Aprendizaje
#
# Una de las ventajas del DL es el uso de modelos preentrenados para resolver tareas nuevas, tareas para las cuales la ANN no fue entrenada inicialmente.
# Esto es posible dado que en cada capa de la ANN se aprenden diferentes características con diferentes niveles de abstracción. La ANN se puede cortar en cualquier capa y utilizar los pesos aprendidos hasta esa capa. A la capa cortada se le pueden anexar nuevas capas. Posteriormente se entrena el modelo con los nuevos datos. Este entrenamiento es más rápido pues inicia desde un punto avanzado y no desde cero. Esta técnica se conoce como transferencia de aprendizaje (**transfer learning**).
#
# 
# <NAME>, "Transfer Learning - Machine Learning's Next Frontier". http://ruder.io/transfer-learning/, 2017.
#
# <br>
#
# El *Transfer Learning* es un método de optimización, un atajo para ahorrar tiempo y obtener mejores resultados, pues se suelen utilizar modelos entrenados con grandes datasets, que continen alta cantidad de clases y variabilidad de datos.
#
# <br>
#
# Este método tiene tres grandes ventajas:
#
# * **Inicio avanzado**. La capacidad inicial en el modelo de origen (antes de refinar el modelo) es mayor de lo que sería iniciando desde cero.
# * **Pendiente más alta**. La tasa de precisión durante el entrenamiento del modelo es más pronunciada.
# * **Asíntota superior**. La tasa de convergencia del modelo entrenado es mejor.
#
# + [markdown] id="tcEaYyzaLt5n"
# # Ejercicio
#
# En este notebook utilizaremos el mecanismo de *Transferencia de Aprendizaje* para clasificar imágenes.
# + [markdown] id="yLV4a82oK5ct"
# # Cargando un modelo pre-entrenado
#
# Para nuestro ejercicio usaremos ResNet18, una red Red Neuronal Convolucional con 18 capas de profundidad. Recordemos que el nombre de **Residual-Net** proviene de la estrategia de usar conexiones de salto residualmente dentro de bloques (llamados bloques residuales, ver Figura), donde la entrada *x* se agrega directamente a la salida del bloque, es decir, *F(x) + x* garantizando el aumento en la profundidad de la red al omitir ciertas capas utilizando conexiones de omisión o bloques residuales. Un gran avance para el problema de optimización/degradación con redes profundas.
# [Paper de presentación de ResNet](https://arxiv.org/pdf/1512.03385.pdf).
#
# 
#
# <br>
#
# El modelo pre-entrenado puede clasificar imágenes en al menos 1000 categorías diferentes, tal como teclado, mouse, lápiz y diferentes clases de animales. Dada la variedad de objetos en el dataset de entrenamieto el modelo es rico en representación de features en un alto rango de clases. ResNet fue entrenado con [Imagenet](https://image-net.org/) un gigantesco dataset visual diseñado para el uso de proyectos en reconocimiento de objetos. Imagenet tiene al menos 14 millones de imágenes anotadas. La entrada de la red tiene un tamaño de 224x224x3.
#
#
# + [markdown] id="jAXsFFGuhC57"
# # Cargando los Datos
# + id="JRJDFkxOiktD"
#-- Descomprimimos el dataset
# # !rm -r mnist
# # !unzip /content/drive/MyDrive/Colab/IntroDeepLearning_202102/mnist.zip
# + id="D3WE5CFBlK_h" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="0a43745f-e980-488d-9a0b-97fc83227485"
#--- Buscamos las direcciones de cada archivo de imagen
from glob import glob
train_files = glob('./mnist/train/*/*.png')
valid_files = glob('./mnist/valid/*/*.png')
test_files = glob('./mnist/test/*/*.png')
#--- Ordenamos los datos de forma aleatoria para evitar sesgos
import numpy as np
np.random.shuffle(train_files)
np.random.shuffle(valid_files)
np.random.shuffle(test_files)
# + [markdown] id="Ymx3UK9uq1ey"
# Al momento de cargar los datos es necesario reescalarlos al tamaño de entrada del modelo reciclado en este caso (224,224,3). Es necesario reescalar y normalizar las imágenes (La normalización se hace usando la media y la desviación estándar). La normalización ayuda a la red a converger más rápido.
# + id="8-CKtsWBeSUh"
import torchvision.transforms as transforms
#--- Transformamos los datos para adaptarlos a la entrada de ResNet 224x224 px
data_transform = transforms.Compose([
transforms.Resize((224, 224)),
transforms.Grayscale(3), #Dado que MNIST tiene un solo canal, lo cambiamos a 3 para no tener que modificar más capas en el modelo
transforms.ToTensor(),
transforms.Normalize(mean=[0.485, 0.456, 0.406],std=[0.229, 0.224, 0.225])
])
# + id="2WZ6U1F0W6JS"
#--- Cargamos los datos de entrenamiento en listas
from PIL import Image
N_train = len(train_files)
X_train = []
Y_train = []
for i, train_file in enumerate(train_files):
Y_train.append( int(train_file.split('/')[3]) )
X_train.append( np.array(data_transform(Image.open(train_file) )))
# + id="ohJqowaMW-in"
#--- Cargamos los datos de testeo en listas
N_test = len(test_files)
X_test = []
Y_test = []
for i, test_file in enumerate(test_files):
Y_test.append( int(test_file.split('/')[3]) )
X_test.append( np.array(data_transform(Image.open(test_file)) ))
# + colab={"base_uri": "https://localhost:8080/", "height": 551} id="X4TFrZLmXB1E" outputId="34a03070-43d0-4282-f3d8-f8f1e37cce5c"
#-- Visualizamos los datos
import matplotlib.pyplot as plt
fig = plt.figure(figsize=(8,8))
for i in range(4):
plt.subplot(2,2,i+1)
plt.imshow(X_test[i*15].reshape(224,224,3))
plt.title(Y_test[i*15])
plt.axis(False)
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="J90slXqUcZfI" outputId="403e15c2-3076-47d6-ea63-b05cdf6d6567"
#--- Convetimos las listas con los datos a tensores de torch
import torch
from torch.autograd import Variable
X_train = Variable(torch.from_numpy(np.array(X_train))).float()
Y_train = Variable(torch.from_numpy(np.array(Y_train))).long()
X_test = Variable(torch.from_numpy(np.array(X_test))).float()
Y_test = Variable(torch.from_numpy(np.array(Y_test))).long()
X_train.data.size()
# + id="kJayWpqAYZI7"
#-- Creamos el DataLoader
batch_size = 32
train_ds = torch.utils.data.TensorDataset(X_train, Y_train)
train_dl = torch.utils.data.DataLoader(train_ds, batch_size=batch_size, shuffle=True)
# + [markdown] id="km5718N3gdl8"
# # Entrenando el modelo
# + colab={"base_uri": "https://localhost:8080/"} id="fP68YXpSKLhI" outputId="db5e3297-1331-4b43-efa8-1f3a3935df53"
#--- Seleccionamos y cargamos el modelo
import torch
model = torch.hub.load('pytorch/vision', 'resnet18', pretrained=True)
model
# + colab={"base_uri": "https://localhost:8080/"} id="-3AssIKykzg2" outputId="c438a1bf-dc4c-40e6-9ce2-c1cbf4cb1767"
#--- Congelamos los pesos en las capaz del modelo para que no se actualicen
for p in model.parameters():
p.requires_grad = False
#--- Definimos el número de clases
out_dim = 10
#--- Reescribimos la nueva capa de salida con el nuevo dataset
model.fc = torch.nn.Sequential(
torch.nn.Linear(model.fc.in_features, out_dim)
)
model.load_state_dict(model.state_dict())
model
# + colab={"base_uri": "https://localhost:8080/"} id="dIeGbvYyafpd" outputId="5cea9c8e-4f69-402c-b077-1c4a8519a46c"
# !pip install hiddenlayer
# + colab={"base_uri": "https://localhost:8080/", "height": 327} id="4FA4ZfwYaiPc" outputId="a1b9a439-cc2c-4f56-e508-27c78dcf159a"
#--- Visualizamos la estructura de nuestra CNN
import hiddenlayer as hl
hl.build_graph(model, torch.zeros([64,3,264,264]))
# + colab={"base_uri": "https://localhost:8080/", "height": 585} id="24ZJYjixjquT" outputId="27f6f52e-504f-4192-feb3-a8aaa4c5dcc1"
#--- Creamos variables para almacenar los scores en cada época
model = model.cuda()
model.train()
#--- Definimos nuestro criterio de evaluación y el optimizador
optimizer = torch.optim.SGD(model.parameters(), lr=0.001, weight_decay=0.1)
criterion = torch.nn.CrossEntropyLoss()
#--- Entrenamos el modelo usando únicamente 5 épocas
n_epochs = 5
history = hl.History()
canvas = hl.Canvas()
iter = 0
for epoch in range(n_epochs):
for batch_idx, (X_train_batch, Y_train_batch) in enumerate(train_dl):
# Pasamos os datos a 'cuda'
X_train_batch = X_train_batch.cuda()
Y_train_batch = Y_train_batch.cuda()
# Realiza una predicción
Y_pred = model(X_train_batch)
# Calcula el loss
loss = criterion(Y_pred, Y_train_batch)
Y_pred = torch.argmax(Y_pred, 1)
# Calcula el accuracy
acc = sum(Y_train_batch == Y_pred)/len(Y_pred)
# Backpropagation
optimizer.zero_grad()
loss.backward()
optimizer.step()
if iter%10 == 0:
#-- Visualizamos la evolución de los score loss y accuracy
history.log((epoch+1, iter), loss=loss, accuracy=acc)
with canvas:
canvas.draw_plot(history["loss"])
canvas.draw_plot(history["accuracy"])
iter += 1
del X_train_batch, Y_train_batch, Y_pred
# + colab={"base_uri": "https://localhost:8080/"} id="Traw3qusmhyH" outputId="9e187024-8d01-43c7-cda2-dcd4010ed96f"
#-- Validamos el modelo
from sklearn.metrics import f1_score
model.cpu()
model.eval()
Y_pred = model(X_test)
loss = criterion(Y_pred,Y_test)
Y_pred = torch.argmax(Y_pred, 1)
f1 = f1_score(Y_test, Y_pred, average='macro')
acc = sum(Y_test == Y_pred)/len(Y_pred)
print( 'Loss:{:.2f}, F1:{:.2f}, Acc:{:.2f}'.format(loss.item(), f1, acc ) )
# + id="0Yzd30Yv0yvh"
#--- Guardamos el nuevo Modelo
torch.save(model,open('./ResNet_MNIST.pt','wb'))
# + id="0fN9qjv8nKSq"
from sklearn.metrics import confusion_matrix
def CM(Y_true, Y_pred, classes, lclasses=None):
fig = plt.figure(figsize=(10, 10))
cm = confusion_matrix(Y_true, Y_pred)
if lclasses == None:
lclasses = np.arange(0,classes)
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
cmap=plt.cm.Blues
ax = fig.add_subplot(1,1,1)
im = ax.imshow(cm, interpolation='nearest', cmap=cmap)
ax.figure.colorbar(im, ax=ax, pad=0.01, shrink=0.86)
ax.set(xticks=np.arange(cm.shape[1]), yticks=np.arange(cm.shape[0]),xticklabels=lclasses, yticklabels=lclasses)
ax.set_xlabel("Predicted",size=20)
ax.set_ylabel("True",size=20)
ax.set_ylim(classes-0.5, -0.5)
plt.setp(ax.get_xticklabels(), size=12)
plt.setp(ax.get_yticklabels(), size=12)
fmt = '.2f'
thresh = cm.max()/2.
for i in range(cm.shape[0]):
for j in range(cm.shape[1]):
ax.text(j, i, format(cm[i, j], fmt),ha="center", va="center",size=15 , color="white" if cm[i, j] > thresh else "black")
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 543} id="0iEXPKLTsc_b" outputId="2113b102-315d-4bfb-e659-94e4e3d12f25"
CM(Y_test, Y_pred, 10)
# + id="JZX7rq8PsiXQ"
| notebooks/15_Reutilizando_CNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import time
import tqdm
import inceptionv3
import numpy as np
import tensorflow as tf
import utils
import defense
# -
config = tf.ConfigProto(gpu_options=tf.GPUOptions(allow_growth=True))
sess = tf.Session(config=config)
data_path = "./data"
output_path = "./finalresults"
if not os.path.exists(output_path):
os.makedirs(output_path)
cleandata = np.load(os.path.join(data_path, "clean100data.npy"))
cleanlabel = np.load(os.path.join(data_path, "clean100label.npy"))
targets = np.load(os.path.join(data_path, "random_targets.npy"))
# +
xs = tf.placeholder(tf.float32, (299, 299, 3))
l2_x = tf.placeholder(tf.float32, (299, 299, 3))
l2_orig = tf.placeholder(tf.float32, (299, 299, 3))
label = tf.placeholder(tf.int32, ())
one_hot = tf.expand_dims(tf.one_hot(label, 1000), axis=0)
lam = 1e-6
epsilon = 0.05
max_steps = 1000
LR = 0.1
label = tf.placeholder(tf.int32, ())
one_hot = tf.expand_dims(tf.one_hot(label, 1000), axis=0)
logits, preds = inceptionv3.model(sess, tf.expand_dims(xs, axis=0))
l2_loss = tf.sqrt(2 * tf.nn.l2_loss(l2_x - l2_orig) / (299, 299, 3))
labels = tf.tile(one_hot, (logits.shape[0], 1))
xent = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits_v2(logits=logits, labels=labels))
loss = xent + lam * tf.maximum(l2_loss - epsilon, 0)
grad, = tf.gradients(loss, xs)
# -
# # Without defense
adv = np.copy(data)
for index in range(data.shape[0]):
adv_bpda = np.copy(adv[index])
for i in tqdm.tqdm(range(max_steps)):
p, l2 = sess.run([preds, l2_loss], {xs: adv_bpda, l2_x: adv_bpda, l2_orig: data[index]})
if p == targets[index] and l2 < epsilon:
print("Found AE. Iter: {}. L2: {}.".format(i, l2))
break
elif l2 > epsilon:
print("Can't find AE under l2-norm 0.05.")
break
g = sess.run(grad, {xs: adv_def[0], label: targets[index]})
adv_bpda -= lr * g
adv_bpda = np.clip(adv_bpda, 0, 1)
adv[index] = adv_bpda
# # Adopt the BPDA over RDDfense
adv = np.copy(data)
for index in range(data.shape[0]):
adv_bpda = np.copy(adv[index])
for i in tqdm.tqdm(range(max_steps)):
adv_def = defense.defend_FD_sig(adv_bpda)
adv_def = defense.defended(defense_func, np.expand_dims(adv_def, axis=0))
p, l2 = sess.run([preds, l2_loss], {xs: adv_def[0], l2_x: adv_bpda, l2_orig: data[index]})
if p == targets[index] and l2 < epsilon:
print("Found AE. Iter: {}. L2: {}.".format(i, l2))
break
elif l2 > epsilon:
print("Can't find AE under l2-norm 0.05.")
break
g = sess.run(grad, {xs: adv_def[0], label: targets[index]})
adv_bpda -= lr * g
adv_bpda = np.clip(adv_bpda, 0, 1)
adv[index] = adv_bpda
np.save(path+'/NoDAcc.npy', AccList)
np.save(path+'/NoDSuc.npy', SuccessList)
np.save(path+'/NoDL2.npy', L2List)
np.save(path+'/NoDLinf.npy', LinfList)
# +
# RDD (RDG+FD)
funcname = 'GD'
LAM = 1.0
AccList_BPDA = []
L2List_BPDA = []
LinfList_BPDA = []
SuccessList_BPDA = []
LR = 0.1
totalbatch = 5
batchsize = 20
totalrounds = 50
adv_bpda = np.copy(cleandata)
start = time.time()
for i in tqdm.tqdm(range(50)):
fdadv_bpda = defend_FD(adv_bpda)
defendadv_bpda = defended(funcname, fdadv_bpda)
p_B= sess.run(preds,{x: defendadv_bpda})
Accuracy_B = np.mean((p_B == cleanlabel).astype(int))
SuccessRate_B = np.mean((p_B == TARGET).astype(int))
AccList_BPDA.append(Accuracy_B)
L2 = l2_distortion(adv_bpda, cleandata)
Linf = linf_distortion(adv_bpda, cleandata)
L2List_BPDA.append(L2)
LinfList_BPDA.append(Linf)
SuccessList_BPDA.append(SuccessRate_B)
for numbatch in range(totalbatch):
cleanbatch,_ = getabatch(cleandata,cleanlabel,numbatch,batchsize)
adv_bpda_batch,labelbatch = getabatch(adv_bpda,cleanlabel,numbatch,batchsize)
fdadv_bpda_batch = defend_FD(adv_bpda_batch)
adv_defbatch = defended(funcname, fdadv_bpda_batch)
g_bpda = sess.run(grad, {x: adv_defbatch})
adv_bpda_batch -= LR * g_bpda
adv_bpda_batch = np.clip(adv_bpda_batch, 0, 1)
adv_bpda[numbatch*batchsize:(numbatch*batchsize+batchsize)] = adv_bpda_batch
end = time.time()
print('total time: ' + str(end - start))
# -
np.save(path+'/RDD_BPDA_Acc.npy', AccList_BPDA)
np.save(path+'/RDD_BPDA_Suc.npy', SuccessList_BPDA)
np.save(path+'/RDD_BPDA_L2.npy', L2List_BPDA)
np.save(path+'/RDD_BPDA_Linf.npy', LinfList_BPDA)
# +
# RANDONLY
funcname = 'onlyrand'
LAM = 1.0
AccList_RAND_BPDA = []
L2List_RAND_BPDA = []
LinfList_RAND_BPDA = []
SuccessList_RAND_BPDA = []
LR = 0.1
totalbatch = 5
batchsize = 20
totalrounds = 50
adv_bpda = np.copy(cleandata)
start = time.time()
for i in tqdm.tqdm(range(50)):
defendadv_bpda = defended(funcname, adv_bpda)
p_B,l2_B = sess.run(preds,{x: defendadv_bpda, lam: LAM,l2_x: adv_bpda, l2_orig: cleandata})
Accuracy_B = np.mean((p_B == cleanlabel).astype(int))
SuccessRate_B = np.mean((p_B == TARGET).astype(int))
AccList_RAND_BPDA.append(Accuracy_B)
L2 = l2_distortion(adv_bpda, cleandata)
Linf = linf_distortion(adv_bpda, cleandata)
L2List_RAND_BPDA.append(L2)
LinfList_RAND_BPDA.append(Linf)
SuccessList_RAND_BPDA.append(SuccessRate_B)
for numbatch in range(totalbatch):
cleanbatch,_ = getabatch(cleandata,cleanlabel,numbatch,batchsize)
adv_bpda_batch,labelbatch = getabatch(adv_bpda,cleanlabel,numbatch,batchsize)
adv_defbatch = defended(funcname, adv_bpda_batch)
g_bpda, p_bpda = sess.run([grad, preds], {x: adv_defbatch,lam: LAM,l2_x: adv_bpda_batch, l2_orig: cleanbatch})
adv_bpda_batch -= LR * g_bpda
adv_bpda_batch = np.clip(adv_bpda_batch, 0, 1)
adv_bpda[numbatch*batchsize:(numbatch*batchsize+batchsize)] = adv_bpda_batch
end = time.time()
print('total time: ' + str(end - start))
# -
np.save(path+'/RAND_BPDA_Acc.npy', AccList_RAND_BPDA)
np.save(path+'/RAND_BPDA_Suc.npy', SuccessList_RAND_BPDA)
np.save(path+'/RAND_BPDA_L2.npy', L2List_RAND_BPDA)
np.save(path+'/RAND_BPDA_Linf.npy', LinfList_RAND_BPDA)
# +
# RAND+FD
funcname = 'onlyrand'
LAM = 1.0
AccList_RAND_FD_BPDA = []
L2List_RAND_FD_BPDA = []
LinfList_RAND_FD_BPDA = []
SuccessList_RAND_FD_BPDA = []
LR = 0.1
totalbatch = 5
batchsize = 20
totalrounds = 50
adv_bpda = np.copy(cleandata)
start = time.time()
for i in tqdm.tqdm(range(50)):
fdadv_bpda = defend_FD(adv_bpda)
defendadv_bpda = defended(funcname, fdadv_bpda)
p_B,l2_B = sess.run([preds,normalized_l2_loss],{x: defendadv_bpda, lam: LAM,l2_x: adv_bpda, l2_orig: cleandata})
Accuracy_B = np.mean((p_B == cleanlabel).astype(int))
SuccessRate_B = np.mean((p_B == TARGET).astype(int))
AccList_RAND_FD_BPDA.append(Accuracy_B)
L2 = l2_distortion(adv_bpda, cleandata)
Linf = linf_distortion(adv_bpda, cleandata)
L2List_RAND_FD_BPDA.append(L2)
LinfList_RAND_FD_BPDA.append(Linf)
SuccessList_RAND_FD_BPDA.append(SuccessRate_B)
for numbatch in range(totalbatch):
cleanbatch,_ = getabatch(cleandata,cleanlabel,numbatch,batchsize)
adv_bpda_batch,labelbatch = getabatch(adv_bpda,cleanlabel,numbatch,batchsize)
fdadv_bpda_batch = defend_FD(adv_bpda_batch)
adv_defbatch = defended(funcname, fdadv_bpda_batch)
g_bpda, p_bpda = sess.run([grad, preds], {x: adv_defbatch,lam: LAM,l2_x: adv_bpda_batch, l2_orig: cleanbatch})
adv_bpda_batch -= LR * g_bpda
adv_bpda_batch = np.clip(adv_bpda_batch, 0, 1)
adv_bpda[numbatch*batchsize:(numbatch*batchsize+batchsize)] = adv_bpda_batch
end = time.time()
print('total time: ' + str(end - start))
# -
np.save(path+'/RAND_FD_BPDA_Acc.npy', AccList_RAND_FD_BPDA)
np.save(path+'/RAND_FD_BPDA_Suc.npy', SuccessList_RAND_FD_BPDA)
np.save(path+'/RAND_FD_BPDA_L2.npy', L2List_RAND_FD_BPDA)
np.save(path+'/RAND_FD_BPDA_Linf.npy', LinfList_RAND_FD_BPDA)
# +
# FD
funcname = 'FD'
LAM = 1.0
AccList_FD_BPDA = []
SuccessList_FD_BPDA = []
LR = 0.1
totalbatch = 5
batchsize = 20
totalrounds = 50
adv_bpda = np.copy(cleandata)
start = time.time()
for i in tqdm.tqdm(range(50)):
defendadv_bpda = cropresult(defend_FD(padresult(adv_bpda)))
p_B,l2_B = sess.run([preds,normalized_l2_loss],{x: defendadv_bpda, lam: LAM,l2_x: adv_bpda, l2_orig: cleandata})
Accuracy_B = np.mean((p_B == cleanlabel).astype(int))
SuccessRate_B = np.mean((p_B == TARGET).astype(int))
AccList_FD_BPDA.append(Accuracy_B)
SuccessList_FD_BPDA.append(SuccessRate_B)
for numbatch in range(totalbatch):
cleanbatch,_ = getabatch(cleandata,cleanlabel,numbatch,batchsize)
adv_bpda_batch,labelbatch = getabatch(adv_bpda,cleanlabel,numbatch,batchsize)
adv_defbatch = cropresult(defend_FD(padresult(adv_bpda_batch)))
g_bpda, p_bpda = sess.run([grad, preds], {x: adv_defbatch,lam: LAM,l2_x: adv_bpda_batch, l2_orig: cleanbatch})
adv_bpda_batch -= LR * g_bpda
adv_bpda_batch = np.clip(adv_bpda_batch, 0, 1)
adv_bpda[numbatch*batchsize:(numbatch*batchsize+batchsize)] = adv_bpda_batch
end = time.time()
print('total time: ' + str(end - start))
# -
np.save(path+'/FD_BPDA_Acc.npy', AccList_FD_BPDA)
np.save(path+'/FD_BPDA_Suc.npy', SuccessList_FD_BPDA)
# +
# PD
funcname = 'pixel_deflection'
LAM = 1.0
AccList_PD_BPDA = []
SuccessList_PD_BPDA = []
LR = 0.1
totalbatch = 5
batchsize = 20
totalrounds = 50
adv_bpda = np.copy(cleandata)
start = time.time()
for i in tqdm.tqdm(range(50)):
defendadv_bpda = defended(funcname, adv_bpda)
p_B,l2_B = sess.run([preds,normalized_l2_loss],{x: defendadv_bpda, lam: LAM,l2_x: adv_bpda, l2_orig: cleandata})
Accuracy_B = np.mean((p_B == cleanlabel).astype(int))
SuccessRate_B = np.mean((p_B == TARGET).astype(int))
AccList_PD_BPDA.append(Accuracy_B)
SuccessList_PD_BPDA.append(SuccessRate_B)
for numbatch in range(totalbatch):
cleanbatch,_ = getabatch(cleandata,cleanlabel,numbatch,batchsize)
adv_bpda_batch,labelbatch = getabatch(adv_bpda,cleanlabel,numbatch,batchsize)
adv_defbatch = defended(funcname, adv_bpda_batch)
g_bpda, p_bpda = sess.run([grad, preds], {x: adv_defbatch,lam: LAM,l2_x: adv_bpda_batch, l2_orig: cleanbatch})
adv_bpda_batch -= LR * g_bpda
adv_bpda_batch = np.clip(adv_bpda_batch, 0, 1)
adv_bpda[numbatch*batchsize:(numbatch*batchsize+batchsize)] = adv_bpda_batch
end = time.time()
print('total time: ' + str(end - start))
# -
np.save(path+'/PD_BPDA_Acc.npy', AccList_PD_BPDA)
np.save(path+'/PD_BPDA_Suc.npy', SuccessList_PD_BPDA)
# +
# SHIELD
funcname = 'SHIELD'
LAM = 1.0
AccList_SHIELD_BPDA = []
SuccessList_SHIELD_BPDA = []
LR = 0.1
totalbatch = 5
batchsize = 20
totalrounds = 50
adv_bpda = np.copy(cleandata)
start = time.time()
for i in tqdm.tqdm(range(50)):
defendadv_bpda = defended(funcname, adv_bpda)
p_B,l2_B = sess.run([preds,normalized_l2_loss],{x: defendadv_bpda, lam: LAM,l2_x: adv_bpda, l2_orig: cleandata})
Accuracy_B = np.mean((p_B == cleanlabel).astype(int))
SuccessRate_B = np.mean((p_B == TARGET).astype(int))
AccList_SHIELD_BPDA.append(Accuracy_B)
SuccessList_SHIELD_BPDA.append(SuccessRate_B)
for numbatch in range(totalbatch):
cleanbatch,_ = getabatch(cleandata,cleanlabel,numbatch,batchsize)
adv_bpda_batch,labelbatch = getabatch(adv_bpda,cleanlabel,numbatch,batchsize)
adv_defbatch = defended(funcname, adv_bpda_batch)
g_bpda, p_bpda = sess.run([grad, preds], {x: adv_defbatch,lam: LAM,l2_x: adv_bpda_batch, l2_orig: cleanbatch})
adv_bpda_batch -= LR * g_bpda
adv_bpda_batch = np.clip(adv_bpda_batch, 0, 1)
adv_bpda[numbatch*batchsize:(numbatch*batchsize+batchsize)] = adv_bpda_batch
end = time.time()
print('total time: ' + str(end - start))
# -
np.save(path+'/SHIELD_BPDA_Acc.npy', AccList_SHIELD_BPDA)
np.save(path+'/SHIELD_BPDA_Suc.npy', SuccessList_SHIELD_BPDA)
# +
# Bit-depth Reduction
funcname = 'BitReduct'
LAM = 1.0
AccList_BitR_BPDA = []
SuccessList_BitR_BPDA = []
LR = 0.1
totalbatch = 5
batchsize = 20
totalrounds = 50
adv_bpda = np.copy(cleandata)
start = time.time()
for i in tqdm.tqdm(range(50)):
defendadv_bpda = defended(funcname, adv_bpda)
p_B,l2_B = sess.run([preds,normalized_l2_loss],{x: defendadv_bpda, lam: LAM,l2_x: adv_bpda, l2_orig: cleandata})
Accuracy_B = np.mean((p_B == cleanlabel).astype(int))
SuccessRate_B = np.mean((p_B == TARGET).astype(int))
AccList_BitR_BPDA.append(Accuracy_B)
SuccessList_BitR_BPDA.append(SuccessRate_B)
for numbatch in range(totalbatch):
cleanbatch,_ = getabatch(cleandata,cleanlabel,numbatch,batchsize)
adv_bpda_batch,labelbatch = getabatch(adv_bpda,cleanlabel,numbatch,batchsize)
adv_defbatch = defended(funcname, adv_bpda_batch)
g_bpda, p_bpda = sess.run([grad, preds], {x: adv_defbatch,lam: LAM,l2_x: adv_bpda_batch, l2_orig: cleanbatch})
adv_bpda_batch -= LR * g_bpda
adv_bpda_batch = np.clip(adv_bpda_batch, 0, 1)
adv_bpda[numbatch*batchsize:(numbatch*batchsize+batchsize)] = adv_bpda_batch
end = time.time()
print('total time: ' + str(end - start))
# -
np.save(path+'/BitR_BPDA_Acc.npy', AccList_BitR_BPDA)
np.save(path+'/BitR_BPDA_Suc.npy', SuccessList_BitR_BPDA)
# +
# Jpeg
funcname = 'FixedJpeg'
LAM = 1.0
AccList_Jpeg_BPDA = []
SuccessList_Jpeg_BPDA = []
LR = 0.1
totalbatch = 5
batchsize = 20
totalrounds = 50
adv_bpda = np.copy(cleandata)
start = time.time()
for i in tqdm.tqdm(range(50)):
defendadv_bpda = defended(funcname, adv_bpda)
p_B,l2_B = sess.run([preds,normalized_l2_loss],{x: defendadv_bpda, lam: LAM,l2_x: adv_bpda, l2_orig: cleandata})
Accuracy_B = np.mean((p_B == cleanlabel).astype(int))
SuccessRate_B = np.mean((p_B == TARGET).astype(int))
AccList_Jpeg_BPDA.append(Accuracy_B)
SuccessList_Jpeg_BPDA.append(SuccessRate_B)
for numbatch in range(totalbatch):
cleanbatch,_ = getabatch(cleandata,cleanlabel,numbatch,batchsize)
adv_bpda_batch,labelbatch = getabatch(adv_bpda,cleanlabel,numbatch,batchsize)
adv_defbatch = defended(funcname, adv_bpda_batch)
g_bpda, p_bpda = sess.run([grad, preds], {x: adv_defbatch,lam: LAM,l2_x: adv_bpda_batch, l2_orig: cleanbatch})
adv_bpda_batch -= LR * g_bpda
adv_bpda_batch = np.clip(adv_bpda_batch, 0, 1)
adv_bpda[numbatch*batchsize:(numbatch*batchsize+batchsize)] = adv_bpda_batch
end = time.time()
print('total time: ' + str(end - start))
# -
np.save(path+'/Jpeg_BPDA_Acc.npy', AccList_Jpeg_BPDA)
np.save(path+'/Jpeg_BPDA_Suc.npy', SuccessList_Jpeg_BPDA)
# +
****************
# Total Varience
# takes too much time to run
funcname = 'TotalVarience'
LAM = 1.0
AccList_TV_BPDA = []
SuccessList_TV_BPDA = []
LR = 0.1
totalbatch = 5
batchsize = 20
totalrounds = 50
adv_bpda = np.copy(cleandata)
start = time.time()
for i in tqdm.tqdm(range(50)):
defendadv_bpda = defended(funcname, adv_bpda)
p_B,l2_B = sess.run([preds,normalized_l2_loss],{x: defendadv_bpda, lam: LAM,l2_x: adv_bpda, l2_orig: cleandata})
Accuracy_B = np.mean((p_B == cleanlabel).astype(int))
SuccessRate_B = np.mean((p_B == TARGET).astype(int))
AccList_TV_BPDA.append(Accuracy_B)
SuccessList_TV_BPDA.append(SuccessRate_B)
for numbatch in range(totalbatch):
cleanbatch,_ = getabatch(cleandata,cleanlabel,numbatch,batchsize)
adv_bpda_batch,labelbatch = getabatch(adv_bpda,cleanlabel,numbatch,batchsize)
adv_defbatch = defended(funcname, adv_bpda_batch)
g_bpda, p_bpda = sess.run([grad, preds], {x: adv_defbatch,lam: LAM,l2_x: adv_bpda_batch, l2_orig: cleanbatch})
adv_bpda_batch -= LR * g_bpda
adv_bpda_batch = np.clip(adv_bpda_batch, 0, 1)
adv_bpda[numbatch*batchsize:(numbatch*batchsize+batchsize)] = adv_bpda_batch
end = time.time()
print('total time: ' + str(end - start))
# -
np.save(path+'/TV_BPDA_Acc.npy', AccList_TV_BPDA)
np.save(path+'/TV_BPDA_Suc.npy', SuccessList_TV_BPDA)
defend_quilt = make_defend_quilt(sess)
def defended_quilt(adv):
defendadv = np.zeros((adv.shape[0],299,299,3))
for i in range(adv.shape[0]):
defendadv[i] = defend_quilt(adv[i])
return defendadv
# +
****************
# Image Quilting
# GPU memory might explode
LAM = 1.0
AccList_IQ_BPDA = []
SuccessList_IQ_BPDA = []
LR = 0.1
totalbatch = 5
batchsize = 20
totalrounds = 50
adv_bpda = np.copy(cleandata)
start = time.time()
for i in tqdm.tqdm(range(50)):
defendadv_bpda = defended_quilt(adv_bpda)
p_B,l2_B = sess.run([preds,normalized_l2_loss],{x: defendadv_bpda, lam: LAM,l2_x: adv_bpda, l2_orig: cleandata})
Accuracy_B = np.mean((p_B == cleanlabel).astype(int))
SuccessRate_B = np.mean((p_B == TARGET).astype(int))
AccList_IQ_BPDA.append(Accuracy_B)
SuccessList_IQ_BPDA.append(SuccessRate_B)
for numbatch in range(totalbatch):
cleanbatch,_ = getabatch(cleandata,cleanlabel,numbatch,batchsize)
adv_bpda_batch,labelbatch = getabatch(adv_bpda,cleanlabel,numbatch,batchsize)
adv_defbatch = defended_quilt(adv_bpda_batch)
g_bpda, p_bpda = sess.run([grad, preds], {x: adv_defbatch,lam: LAM,l2_x: adv_bpda_batch, l2_orig: cleanbatch})
adv_bpda_batch -= LR * g_bpda
adv_bpda_batch = np.clip(adv_bpda_batch, 0, 1)
adv_bpda[numbatch*batchsize:(numbatch*batchsize+batchsize)] = adv_bpda_batch
end = time.time()
print('total time: ' + str(end - start))
# -
np.save(path+'/IQ_BPDA_Acc.npy', AccList_IQ_BPDA)
np.save(path+'/IQ_BPDA_Suc.npy', SuccessList_IQ_BPDA)
# +
path = 'finalresults'
AccList = np.load(path+'/NoDAcc.npy')
SuccessList = np.load(path+'/NoDSuc.npy')
L2List = np.load(path+'/NoDL2.npy')
LinfList = np.load(path+'/NoDLinf.npy')
AccList_BPDA = np.load(path+'/RDD_BPDA_Acc.npy')
SuccessList_BPDA = np.load(path+'/RDD_BPDA_Suc.npy')
L2List_BPDA = np.load(path+'/RDD_BPDA_L2.npy')
LinfList_BPDA = np.load(path+'/RDD_BPDA_Linf.npy')
AccList_SHIELD_BPDA = np.load(path+'/SHIELD_BPDA_Acc.npy')
SuccessList_SHIELD_BPDA = np.load(path+'/SHIELD_BPDA_Suc.npy')
AccList_PD_BPDA = np.load(path+'/PD_BPDA_Acc.npy')
SuccessList_PD_BPDA = np.load(path+'/PD_BPDA_Suc.npy')
AccList_FD_BPDA = np.load(path+'/FD_BPDA_Acc.npy')
SuccessList_FD_BPDA = np.load(path+'/FD_BPDA_Suc.npy')
AccList_BitR_BPDA = np.load(path+'/BitR_BPDA_Acc.npy')
SuccessList_BitR_BPDA = np.load(path+'/BitR_BPDA_Suc.npy')
AccList_Jpeg_BPDA = np.load(path+'/Jpeg_BPDA_Acc.npy')
SuccessList_Jpeg_BPDA = np.load(path+'/Jpeg_BPDA_Suc.npy')
AccList_RAND_BPDA = np.load(path+'/RAND_BPDA_Acc.npy')
SuccessList_RAND_BPDA = np.load(path+'/RAND_BPDA_Suc.npy')
AccList_RAND_FD_BPDA = np.load(path+'/RAND_FD_BPDA_Acc.npy')
SuccessList_RAND_FD_BPDA = np.load(path+'/RAND_FD_BPDA_Suc.npy')
L2List_RAND_FD_BPDA = np.load(path+'/RAND_FD_BPDA_L2.npy')
LinfList_RAND_FD_BPDA = np.load(path+'/RAND_FD_BPDA_Linf.npy')
AccList_TV_BPDA = np.load(path+'/TV_BPDA_Acc.npy')
SuccessList_TV_BPDA = np.load(path+'/TV_BPDA_Suc.npy')
# # AccList_IQ_BPDA = np.load(path+'/IQ_BPDA_Acc.npy')
# # SuccessList_IQ_BPDA = np.load(path+'/IQ_BPDA_Suc.npy')
# # L2List_IQ_BPDA = np.load(path+'/IQ_BPDA_L2.npy')
# +
totalrounds = 50
plt.figure(figsize=(10,6))
plt.plot(np.arange(totalrounds),AccList_BPDA, color="b",label='GD+FD attacked by BPDA', linestyle="-", marker="x", linewidth=1.0)
plt.plot(np.arange(totalrounds),AccList_SHIELD_BPDA, color="purple",label='SHIELD attacked by BPDA', linestyle="-.", marker="o", linewidth=1.0)
plt.plot(np.arange(totalrounds),AccList_PD_BPDA, color="y",label='PD attacked by BPDA', linestyle="-.", marker="o", linewidth=1.0)
plt.plot(np.arange(totalrounds),AccList_FD_BPDA, color="r",label='FD attacked by BPDA', linestyle="-.", marker="o", linewidth=1.0)
plt.plot(np.arange(totalrounds),AccList_BitR_BPDA, color="deeppink",label='BitR attacked by BPDA', linestyle="-.", marker="o", linewidth=1.0)
plt.plot(np.arange(totalrounds),AccList_Jpeg_BPDA, color="maroon",label='Jpeg attacked by BPDA', linestyle="-.", marker="o", linewidth=1.0)
plt.plot(np.arange(totalrounds),AccList_TV_BPDA, color="goldenrod",label='TV attacked by BPDA', linestyle="-.", marker="o", linewidth=1.0)
plt.plot(np.arange(totalrounds),AccList_RAND_BPDA, color="sienna",label='RAND attacked by BPDA', linestyle="-.", marker="o", linewidth=1.0)
plt.plot(np.arange(totalrounds),AccList_RAND_FD_BPDA, color="salmon",label='RAND_FD attacked by BPDA', linestyle="-.", marker="o", linewidth=1.0)
plt.plot(np.arange(totalrounds),AccList, color="g", label='Without Defence',linestyle="--", marker="*", linewidth=1.0)
plt.xlabel("Purturbation Rounds",fontsize=20)
plt.ylabel("Model Accuracy",fontsize=20)
plt.xticks(fontsize=15)
plt.yticks(fontsize=15)
plt.legend(fontsize=13)
plt.grid(color="k", linestyle=":")
plt.savefig(path+'/BPDA_ModelAcc.png',bbox_inches='tight')
plt.show()
plt.figure(figsize=(10,6))
plt.plot(np.arange(totalrounds),SuccessList_BPDA, color="b", label='GD+FD attacked by BPDA',linestyle="-", marker="x", linewidth=1.0)
plt.plot(np.arange(totalrounds),SuccessList_SHIELD_BPDA, color="purple", label='SHIELD attacked by BPDA',linestyle="-.", marker="o", linewidth=1.0)
plt.plot(np.arange(totalrounds),SuccessList_PD_BPDA, color="y", label='PD attacked by BPDA',linestyle="-.", marker="o", linewidth=1.0)
plt.plot(np.arange(totalrounds),SuccessList_FD_BPDA, color="r", label='FD attacked by BPDA',linestyle="-.", marker="o", linewidth=1.0)
plt.plot(np.arange(totalrounds),SuccessList_BitR_BPDA, color="deeppink", label='BitR attacked by BPDA',linestyle="-.", marker="o", linewidth=1.0)
plt.plot(np.arange(totalrounds),SuccessList_Jpeg_BPDA, color="maroon", label='Jpeg attacked by BPDA',linestyle="-.", marker="o", linewidth=1.0)
plt.plot(np.arange(totalrounds),SuccessList_TV_BPDA, color="goldenrod", label='TV attacked by BPDA',linestyle="-.", marker="o", linewidth=1.0)
plt.plot(np.arange(totalrounds),SuccessList_RAND_BPDA, color="sienna", label='RAND attacked by BPDA',linestyle="-.", marker="o", linewidth=1.0)
plt.plot(np.arange(totalrounds),SuccessList_RAND_FD_BPDA, color="salmon", label='RAND_FD attacked by BPDA',linestyle="-.", marker="o", linewidth=1.0)
plt.plot(np.arange(totalrounds),SuccessList, color="g", label='Without Defence',linestyle="--", marker="*", linewidth=1.0)
plt.xlabel("Purturbation Rounds",fontsize=20)
plt.ylabel("Adversarial Success Rate",fontsize=20)
plt.xticks(fontsize=15)
plt.yticks(fontsize=15)
plt.legend(fontsize=13)
plt.grid(color="k", linestyle=":")
plt.savefig(path+'/BPDA_AtkSucc.png',bbox_inches='tight')
plt.show()
plt.figure(figsize=(10,6))
plt.plot(np.arange(totalrounds),L2List_BPDA, color="b", label='GD+FD attacked by BPDA',linestyle="-", marker="x", linewidth=1.0)
plt.plot(np.arange(totalrounds),L2List_RAND_FD_BPDA, color="salmon", label='RAND_FD attacked by BPDA',linestyle="-.", marker="o", linewidth=1.0)
plt.plot(np.arange(totalrounds),L2List, color="g", label='Without Defence',linestyle="--", marker="*", linewidth=1.0)
plt.xlabel("Purturbation Rounds",fontsize=20)
plt.ylabel("Normalized L2 Norm",fontsize=20)
plt.xticks(fontsize=15)
plt.yticks(fontsize=15)
plt.legend(fontsize=13)
plt.grid(color="k", linestyle=":")
plt.savefig(path+'/BPDA_AEL2.png',bbox_inches='tight')
plt.show()
plt.figure(figsize=(10,6))
plt.plot(np.arange(totalrounds),LinfList_BPDA, color="b", label='GD+FD attacked by BPDA',linestyle="-", marker="x", linewidth=1.0)
plt.plot(np.arange(totalrounds),LinfList_RAND_FD_BPDA, color="salmon", label='RAND_FD attacked by BPDA',linestyle="-.", marker="o", linewidth=1.0)
plt.plot(np.arange(totalrounds),LinfList, color="g", label='Without Defence',linestyle="--", marker="*", linewidth=1.0)
plt.xlabel("Purturbation Rounds",fontsize=20)
plt.ylabel("Linf Norm",fontsize=20)
plt.xticks(fontsize=15)
plt.yticks(fontsize=15)
plt.legend(fontsize=13)
plt.grid(color="k", linestyle=":")
plt.savefig(path+'/BPDA_AELinf.png',bbox_inches='tight')
plt.show()
| BPDA_compare.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import numpy as np
import h5py
from sklearn.model_selection import train_test_split
from keras.models import Sequential
from keras.layers import Dense, Dropout, Activation, Flatten
from keras.layers import Convolution2D, MaxPooling2D
from keras.optimizers import SGD
from keras.callbacks import ModelCheckpoint
from sklearn.metrics import matthews_corrcoef
from sklearn.metrics import hamming_loss
from keras import backend as K
K.set_image_dim_ordering('th')
# -
x_train, x_test, y_train, y_test = load()
# +
x_train = x_train.astype('float32')
x_test = x_test.astype('float32')
x_train /= 255
x_test /= 255
# +
model = Sequential()
model.add(Convolution2D(32, kernel_size=(3, 3),padding='same',input_shape=(3 , 100, 100)))
model.add(Activation('relu'))
model.add(Convolution2D(32, (3, 3)))
model.add(Activation('relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(0.25))
model.add(Convolution2D(64,(3, 3), padding='same'))
model.add(Activation('relu'))
model.add(Convolution2D(64, (3, 3)))
model.add(Activation('relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(0.25))
model.add(Flatten())
model.add(Dense(512))
model.add(Activation('relu'))
model.add(Dropout(0.5))
model.add(Dense(5))
model.add(Activation('sigmoid'))
# +
sgd = SGD(lr=0.01, decay=1e-6, momentum=0.9, nesterov=True)
model.compile(loss='binary_crossentropy', optimizer=sgd, metrics=['accuracy'])
model.load_weights("weights.hdf5")
# -
model.summary()
out = model.predict_proba(x_test)
out = np.array(out)
out
# +
threshold = np.arange(0.1,0.9,0.1)
acc = []
accuracies = []
best_threshold = np.zeros(out.shape[1])
for i in range(out.shape[1]):
y_prob = np.array(out[:,i])
for j in threshold:
y_pred = [1 if prob>=j else 0 for prob in y_prob]
acc.append( matthews_corrcoef(y_test[:,i],y_pred))
acc = np.array(acc)
index = np.where(acc==acc.max())
accuracies.append(acc.max())
best_threshold[i] = threshold[index[0][0]]
acc = []
# -
best_threshold
y_pred = np.array([[1 if out[i,j]>=best_threshold[j] else 0 for j in range(y_test.shape[1])] for i in range(len(y_test))])
y_pred #predicted labels
y_test #actual labels
hamming_loss(y_test,y_pred) #the loss should be as low as possible and the range is from 0 to 1
total_correctly_predicted = len([i for i in range(len(y_test)) if (y_test[i]==y_pred[i]).sum() == 5])
total_correctly_predicted/400. #exact accuracy for eg y_pred = [0,0,1,1,1] and y_test=[0,0,1,1,1]
total_correctly_predicted
from IPython.display import Image
Image(filename='test_image.jpg')
import cv2
img = cv2.imread("test_image.jpg")
img.shape
img = cv2.resize(img,(100,100))
img.shape
img = img.transpose((2,0,1))
img.shape
img = img.astype('float32')
img = img/255
img = np.expand_dims(img,axis=0)
img.shape
pred = model.predict(img)
pred
y_pred = np.array([1 if pred[0,i]>=best_threshold[i] else 0 for i in range(pred.shape[1])])
y_pred
classes = ['desert','mountains','sea','sunset','trees']
[classes[i] for i in range(5) if y_pred[i]==1 ] #extracting actual class name
# +
#import matplotlib.pyplot as plt
#import matplotlib.image as mpimg
#import numpy as np
#img_load = mpimg.imread('test_image2.jpg')
#imgplot = plt.imshow(img_load)
# +
#img_load.shape
img = cv2.imread('test_image2.jpg')
img = cv2.resize(img,(100,100))
# -
#imgplot = plt.imshow(img)
Image(filename='test_image2.jpg')
img = img.transpose((2,0,1))
img.shape
img = img.astype('float32')
img = img/255
img = np.expand_dims(img,axis=0)
img.shape
pred = model.predict(img)
pred
y_pred = np.array([1 if pred[0,i]>=best_threshold[i] else 0 for i in range(pred.shape[1])])
y_pred
classes = ['desert','mountains','sea','sunset','trees']
[classes[i] for i in range(5) if y_pred[i]==1 ] #extracting actual class name
| .ipynb_checkpoints/04 - Multi Label Image Classification-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Rede convolucional tipo LeNet - dataset CIFAR-10
# Este experimento utiliza uma rede com duas camadas convolucionais e duas camadas densas
# similar à rede LeNet introduzida em 1998 para classificar 3 classes do dataset CIFAR-10.
#
# O notebook é organizado da seguinte forma:
# - importação das bibliotecas
# - carregamento do dataset e criação do data_loader
# - criação da rede
# - parâmetros de treinamento
# - avaliação no conjunto de validação
# - acurácia
# - amostras certas com maior probabilidade, etc.
# - visualização das camadas convolucionais
# ## Dataset CIFAR-10
# CIFAR-10: https://www.cs.toronto.edu/~kriz/cifar.html
#
# O conjunto de dados CIFAR-10 possui 60.000 imagens coloridas de dimensões $32\times 32$. Os dados são divididos em 10 classes com 6.000 imagens por classe.
#
# No exemplo abaixo nós utilizamos apenas 3 das 10 classes. A saída das diferentes camadas da CNN são mostradas.
# ## Importando as bibliotecas
# +
# %matplotlib inline
import os, sys
import numpy as np
import matplotlib.pylab as plt
from collections import OrderedDict
# from torch
import torch
from torch import nn
from torch.autograd import Variable
from torch.optim import lr_scheduler
# from torchvision
import torchvision
from torchvision import transforms
from torchvision.datasets import CIFAR10
# from course libs
import lib.pytorch_trainer as ptt
from lib.cifar import CIFARX, CIFAR_CLASSES
# -
# verifica se a GPU esta disponivel
use_gpu = torch.cuda.is_available()
print("Usando GPU:", use_gpu)
# ## Carregando e mostrando os dados (seleciona apenas as classes 0, 1 e 2)
# +
images_dir = 'data/datasets/CIFAR10/'
# Transformara os dados em tensores no intervalo [0.0, 1.0] (Os dados serão normalizados)
data_transform = transforms.ToTensor()
# carrega somente as classes 0, 1 e 2 do CIFAR dos dados de treinamento e de teste
image_datasets = {
'train': CIFARX(images_dir, [0, 1, 2], train=True, transform=data_transform, download=True),
'val' : CIFARX(images_dir, [0, 1, 2], train=False, transform=data_transform, download=True),
}
print('amostras para treinamento:', len(image_datasets['train']))
print('amostras para validação:', len(image_datasets['val']))
# -
# ## Mostrando algumas imagens do conjunto de treinamento
# +
n_samples = 40
tensor2pil = transforms.ToPILImage()
# cria um DataLoader temporario para pegar um batch de 'n_samples' imagens de treinamento
temp_dataloader = torch.utils.data.DataLoader(image_datasets['train'],
batch_size = n_samples,
shuffle=True, num_workers=4)
# pega um batch de imagens
image_batch, labels = next(iter(temp_dataloader))
# cria um grid com as imagens
grid = torchvision.utils.make_grid(image_batch, normalize=True, pad_value=1.0, padding=1)
img_pil = tensor2pil(grid)
img_pil
# -
# ## Teste inicial com pouquíssimas amostras
testing = True
if testing:
n_samples = 800
image_datasets['train'].train_data = image_datasets['train'].train_data[:n_samples]
image_datasets['train'].train_labels = image_datasets['train'].train_labels[:n_samples]
n_samples_test = 200
image_datasets['val'].test_data = image_datasets['val'].test_data[:n_samples_test]
image_datasets['val'].test_labels = image_datasets['val'].test_labels[:n_samples_test]
# ## Cria o DataLoader para os dados
# +
batch_size = 100
dataloaders = {
'train': torch.utils.data.DataLoader(image_datasets['train'], batch_size = batch_size,
shuffle=True, num_workers=4),
'val' : torch.utils.data.DataLoader(image_datasets['val'], batch_size = batch_size,
shuffle=False, num_workers=4)
}
dataset_sizes = {
'train': len(image_datasets['train']),
'val' : len(image_datasets['val'])
}
class_names = image_datasets['train'].classes
print('Tamanho do conjunto de treinamento:', dataset_sizes['train'])
print('Tamanho do conjunto de validacao:', dataset_sizes['val'])
print('Classes:', class_names)
# -
# # Construíndo a CNN com o PyTorch
# <img src = '../figures/Rede_LeNet_Cifar.png', width=600pt></img>
class MyModel(nn.Module):
def __init__(self):
super(MyModel, self).__init__()
# Camadas convolucionais
self.conv_layer = nn.Sequential(OrderedDict([
('conv1', nn.Conv2d( 3, 32, kernel_size=3)),
('relu1', nn.ReLU()),
('conv2', nn.Conv2d(32, 32, kernel_size=3)),
('relu2', nn.ReLU()),
('max_pool', nn.MaxPool2d(2)),
('drop', nn.Dropout(p=0.25))
]))
# Camadas densas
self.dense_layer = nn.Sequential(OrderedDict([
('dense1', nn.Linear(14*14*32, 128)),
('relu1', nn.ReLU()),
('drop1', nn.Dropout(p=0.5)),
('dense2', nn.Linear(128, 3)),
]))
def forward(self, x):
x = self.conv_layer(x)
x = x.view(-1, 14*14*32) # Transforma a imagem em vetor
x = self.dense_layer(x)
return x
# +
model = MyModel()
if use_gpu:
model = model.cuda()
model
# -
# ## Testando o modelo com uma amostra
# +
# imagem de entrada: 1 amostra, com 3 bandas (RGB), 32 linhas e 32 colunas
xin = Variable(torch.zeros(1, 3, 32, 32))
if use_gpu:
xin = xin.cuda()
# predict da rede: saída 1 linha (amostra) com 3 colunas (scores), (1 de cada classe)
yout = model(xin)
print('Saída:', yout.cpu().data.numpy())
# -
# ## Treinamento da rede
# +
optimizer = torch.optim.Adam(model.parameters())
criterion = nn.CrossEntropyLoss()
savebest = ptt.ModelCheckpoint('../../models/cifar_cnn',reset=True, verbose=1)
# Cria o objeto para treinar a rede
trainer = ptt.DeepNetTrainer(
model = model,
criterion = criterion,
optimizer = optimizer,
callbacks = [ptt.AccuracyMetric(),ptt.PrintCallback(),savebest]
)
# -
trainer.fit_loader(5, train_data=dataloaders['train'], valid_data=dataloaders['val'])
# ## Avaliação
# ### Gráfico da perda
train_loss = trainer.metrics['train']['losses']
valid_loss = trainer.metrics['valid']['losses']
epochs = np.arange(len(train_loss))
plt.plot(epochs, train_loss, label='train')
plt.plot(epochs, valid_loss, label='valid')
plt.legend()
plt.title('Losses')
plt.show()
# ### Gráfico da acurácia
train_acc = trainer.metrics['train']['acc']
valid_acc = trainer.metrics['valid']['acc']
epochs = np.arange(len(train_acc))
plt.plot(epochs, train_acc, label='train')
plt.plot(epochs, valid_acc, label='valid')
plt.legend()
plt.title('Acurácia')
plt.show()
# ## Avaliação do treinamento no conjunto de testes
y_hat_v = trainer.predict_classes_loader(dataloaders['val'])
y_hat = y_hat_v.numpy()
y_proba_v = trainer.predict_probas_loader(dataloaders['val']) # prob das classes
y_prob = y_proba_v.numpy().max(axis=1) # probabilidade máxima da classe
# ### Matriz de confusão
X_test = image_datasets['val'].test_data
y_test = np.array(image_datasets['val'].test_labels)
from pandas import crosstab
crosstab(y_test, y_hat)
X_test.shape,y_test.shape
# ## Predições certas com maior probabilidade
i_ok = np.where(y_hat==y_test)[0]
top_most_ok = np.argsort(y_prob[i_ok])[-5:][::-1]
y_top5 = i_ok[top_most_ok]
fig, raxis = plt.subplots(1,5)
for k,i in enumerate(y_top5):
raxis[k].imshow(X_test[i])
raxis[k].set_title('{}:{}:{:0.3f}'.format(y_test[i],y_hat[i],y_prob[i]))
raxis[k].axis('off')
# ## Predições certas com menor probabilidade
i_ok = np.where(y_hat==y_test)[0]
top_least_ok = np.argsort(y_prob[i_ok])[:5]
y_bot5 = i_ok[top_least_ok]
fig, raxis = plt.subplots(1,5)
for k,i in enumerate(y_bot5):
raxis[k].imshow(X_test[i])
raxis[k].set_title('{}:{}:{:0.3f}'.format(y_test[i],y_hat[i],y_prob[i]))
raxis[k].axis('off')
# ## Predições erradas com maior probabilidade
i_not_ok = np.where(y_hat!=y_test)[0]
top_most_not_ok = np.argsort(y_prob[i_not_ok])[-5:][::-1]
y_most_not_ok_top5 = i_not_ok[top_most_not_ok]
fig, raxis = plt.subplots(1,5)
for k,i in enumerate(y_most_not_ok_top5):
raxis[k].imshow(X_test[i])
raxis[k].set_title('{}:{}:{:0.3f}'.format(y_test[i],y_hat[i],y_prob[i]))
raxis[k].axis('off')
# ## Predições erradas com menor probabilidade
i_not_ok = np.where(y_hat!=y_test)[0]
top_least_not_ok = np.argsort(y_prob[i_not_ok])[:5]
y_least_not_ok_top5 = i_not_ok[top_least_not_ok]
fig, raxis = plt.subplots(1,5)
for k,i in enumerate(y_least_not_ok_top5):
raxis[k].imshow(X_test[i])
raxis[k].set_title('{}:{}:{:0.3f}'.format(y_test[i],y_hat[i],y_prob[i]))
raxis[k].axis('off')
# ## Visualização de uma amostra
sample_number = 70 # amostra de número 70
plt.figure()
plt.imshow(X_test[sample_number])
#plt.axis('off')
plt.title("Original");
# ## Visualização das camadas internas
# Observe que são 32 canais na primeira camada convolucional
# +
ncols = 8
H,W = 14,30
xin, _ = image_datasets['val'][sample_number]
xin = xin.view(1, 3, 32, 32)
x = Variable(xin)
if use_gpu:
x = x.cuda()
model.train(mode=False) # qual é a diferença quando em modo treinamento ou avaliação?
#Mostrando a saida das camadas convolucionais
for name, layer in model.conv_layer.named_children():
x = layer(x)
grid = torchvision.utils.make_grid(torch.transpose(x.data, 0, 1), normalize=True,
pad_value=1.0, padding=1).cpu().numpy()
if name == 'max_pool':
H /= 2
W /= 2
fig = plt.figure(figsize=(H,W))
plt.imshow(grid.transpose((1,2,0)))
plt.title(name)
plt.axis('off')
plt.show()
# -
# ## Exercícios
# 1. Quantos parâmetros são treinados nesta rede? Calcule o número de parâmetros de cada camada, não se esquecendo do *bias*.
# 2. Se as duas camadas convolucionais fossem densas, qual seria o número de parâmetros a serem treinados?
# ## Atividades
# 1. Troque a amostra que está sendo visualizada por outra
# 2. O que acontece na visualização dos valores nas camadas convolucionais, quando
# a rede é colocada no modo de treinamento ou no modo de avaliação? Faça o experimento
# e comprove.
# 3. Fazer uma função que calcule o número total de parâmetros de um objeto modelo de rede:
# ```
# def n_parameters(model):
# # calcula n. parâmetros
# return
# ```
# ## Aprendizados com este experimento
# 1.
| PyTorch/cifar10-CNN-features.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Python 🐍 Data Science with the TCLab
#
# Welcome to this data science course on Python! This course is intended to help you develop data science and machine learning skills in Python. A [beginning Python course](https://github.com/APMonitor/begin_python) is available for programmers with no prior programming experience. As with the beginning course, this course has video tutorials for each exercise if you have questions along the way. One of the unique things about this course is that you work on basic elements and then test your knowledge with data from hardware that controls heaters and measures temperatures. You will see your Python code have a real impact by designing the materials for a new product.
#
# [](https://www.youtube.com/watch?v=pAgW_bZVo88&list=PLLBUgWXdTBDg1Qgmwt4jKtVn9BWh5-zgy "Python Data Science")
#
# One of the best ways to start or review a programming language is to work on a project. These exercises are designed to teach data science Python programming skills. Data science applications are found across almost all industries where raw data is transformed into actionable information that drives scientific discovery, business innovations, and development.
#
# ### Topics
#
# There are 12 lessons to help you with the objective of learning data science in Python. The first thing that you will need is to [install Anaconda](https://github.com/APMonitor/data_science/blob/master/00.%20Introduction.ipynb) to open and run the IPython notebook files in Jupyter. Any Python distribution or Integrated Development Environment (IDE) can be used (IDLE (python.org), Spyder, PyCharm, and others) but Jupyter notebook is required to open and run the IPython notebook (`.ipynb`) files. All of the IPython notebook (`.ipynb`) files can be [downloaded at this link](https://github.com/APMonitor/data_science/archive/master.zip). Don't forget to unzip the folder (extract the archive) and copy it to a convenient location before starting.
#
# 1. [Overview](https://github.com/APMonitor/data_science/blob/master/01.%20Overview.ipynb)
# 2. [Data Import and Export](https://github.com/APMonitor/data_science/blob/master/02.%20Import_Export.ipynb)
# 3. [Data Analysis](https://github.com/APMonitor/data_science/blob/master/03.%20Analyze.ipynb)
# 4. [Visualize Data](https://github.com/APMonitor/data_science/blob/master/04.%20Visualize.ipynb)
# 5. [Prepare (Cleanse, Scale, Divide) Data](https://github.com/APMonitor/data_science/blob/master/05.%20Prepare_data.ipynb)
# 6. [Regression](https://github.com/APMonitor/data_science/blob/master/06.%20Regression.ipynb)
# 7. [Features](https://github.com/APMonitor/data_science/blob/master/07.%20Features.ipynb)
# 8. [Classification](https://github.com/APMonitor/data_science/blob/master/08.%20Classification.ipynb)
# 9. [Interpolation](https://github.com/APMonitor/data_science/blob/master/09.%20Interpolation.ipynb)
# 10. [Solve Equations](https://github.com/APMonitor/data_science/blob/master/10.%20Solve_Equations.ipynb)
# 11. [Differential Equations](https://github.com/APMonitor/data_science/blob/master/11.%20Differential_Equations.ipynb)
# 12. [Time Series](https://github.com/APMonitor/data_science/blob/master/12.%20Time_Series.ipynb)
#
# ### Install Python
#
# [Download and install Anaconda to use Jupyter](https://docs.anaconda.com/anaconda/install/) or [watch a video on how to install Anaconda](https://youtu.be/LrMOrMb8-3s).
#
# [](https://www.youtube.com/watch?v=LrMOrMb8-3s "Install Anaconda")
#
# There are additional instructions on how to [install Python and manage modules](https://apmonitor.com/pdc/index.php/Main/InstallPython).
#
# ### Get TCLab
#
# You will need a [TCLab kit](https://apmonitor.com/heat.htm) to complete the exercises. The TCLab is available for [purchase on Amazon](https://www.amazon.com/TCLab-Temperature-Control-Lab/dp/B07GMFWMRY).
#
# 
#
# ### Support
#
# We would love to hear any feedback or problems you would like to send us! We are always trying to improve this course and would like to hear about your experience. We can be contacted at <EMAIL> for issues related to the course or [getting started with the TCLab](https://apmonitor.com/pdc/index.php/Main/ArduinoSetup).
#
# ### Additional Resources
#
# - [Begin Python Course](https://github.com/APMonitor/begin_python)
# - [Engineering Programming Course](https://apmonitor.com/pdc) with [Source Code](https://github.com/APMonitor/learn_python)
# - [Temperature Control Lab (TCLab) Kit](http://apmonitor.com/pdc/index.php/Main/ArduinoTemperatureControl)
# - [Jupyter as interactive environment for Python, Julia, R](https://jupyter.org/)
| 00. Introduction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Бэггинг и случайный лес
# +
# %matplotlib inline
import matplotlib.pyplot as plt
# +
from sklearn.datasets import load_digits
from sklearn.model_selection import cross_val_score, validation_curve
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import BaggingClassifier, RandomForestClassifier
import numpy as np
import seaborn as sns
# -
# Загрузка датасета digits с помощью функции load_digits из sklearn.datasets и подготовка матрицы признаков X и ответов на обучающей выборке y (потребуются поля data и target в объекте, который возвращает load_digits).
digits = load_digits()
print(digits.data.shape)
print(digits.DESCR)
plt.gray()
plt.plot(digits.images[0])
plt.show()
X = digits.data
y = digits.target
# Для оценки качества далее нужно использовать cross_val_score из sklearn.cross_validation с параметром cv=10. Эта функция реализует k-fold cross validation c k равным значению параметра cv. Предлагается использовать k=10, чтобы полученные оценки качества имели небольшой разброс, и было проще проверить полученные ответы. На практике же часто хватает и k=5. Функция cross_val_score будет возвращать numpy.ndarray, в котором будет k чисел - качество в каждом из k экспериментов k-fold cross validation. Для получения среднего значения (которое и будет оценкой качества работы) вызовите метод .mean() у массива, который возвращает cross_val_score.
# С небольшой вероятностью можно натолкнуться на случай, когда полученное качество в каком-то из пунктов не попадет в диапазон, заданный для правильных ответов - в этом случае необходимо попробовать перезапустить ячейку с cross_val_score несколько раз и выбрать наиболее «типичное» значение.
# Если захочется ускорить вычисление cross_val_score - можно попробовать использовать параметр n_jobs, но нужно быть осторожным: в одной из старых версий sklearn была ошибка, которая приводила к неверному результату работы cross_val_score при задании n_jobs отличным от 1. Сейчас такой проблемы возникнуть не должно, но проверить, что все в порядке, не будет лишним.
# 1.
#
# Создадим DecisionTreeClassifier с настройками по умолчанию и измерим качество его работы с помощью cross_val_score. Эта величина и будет ответом в пункте 1.
dt_classifier = DecisionTreeClassifier()
cvs = cross_val_score(dt_classifier, X, y, cv=10)
print(cvs)
print('Mean model quality value: ' + str(cvs.mean()))
with open("answer1.txt", "w") as fout:
fout.write(str(cvs.mean()))
# 2.
#
# Воспользуемся BaggingClassifier из sklearn.ensemble, чтобы обучить бэггинг над DecisionTreeClassifier. Используем в BaggingClassifier параметры по умолчанию, задав только количество деревьев равным 100.
#
# Качество классификации новой модели - ответ в пункте 2. Обратите внимание, как соотносится качество работы композиции решающих деревьев с качеством работы одного решающего дерева.
bagging = BaggingClassifier(dt_classifier, n_estimators=100)
cvs = cross_val_score(bagging, X, y, cv=10)
print(cvs)
print('Mean model quality value: ' + str(cvs.mean()))
with open("answer2.txt", "w") as fout:
fout.write(str(cvs.mean()))
# 3.
#
# Теперь изучим параметры BaggingClassifier и выберем их такими, чтобы каждый базовый алгоритм обучался не на всех d признаках, а на sqrt(d) случайных признаков. Качество работы получившегося классификатора - ответ в пункте 3. Корень из числа признаков - часто используемая эвристика в задачах классификации, в задачах регрессии же часто берут число признаков, деленное на три. Но в общем случае ничто не мешает вам выбирать любое другое число случайных признаков.
n_features = digits.data.shape[1]
bagging = BaggingClassifier(dt_classifier, n_estimators=100, max_features=int(np.sqrt(n_features)))
cvs = cross_val_score(bagging, X, y, cv=10)
print(cvs)
print('Mean model quality value: ' + str(cvs.mean()))
with open("answer3.txt", "w") as fout:
fout.write(str(cvs.mean()))
# 4.
#
# Наконец, попробуем выбрать случайные признаки не один раз на все дерево, а при построении каждой вершины дерева. Сделать это несложно: нужно убрать выбор случайного подмножества признаков в BaggingClassifier и добавить его в DecisionTreeClassifier. Будем выбирать опять же sqrt(d) признаков. Качество полученного классификатора на контрольной выборке и будет ответом в пункте 4.
dt_classifier = DecisionTreeClassifier(max_features=int(np.sqrt(n_features)))
bagging = BaggingClassifier(dt_classifier, n_estimators=100)
cvs = cross_val_score(bagging, X, y, cv=10)
print(cvs)
print('Mean model quality value: ' + str(cvs.mean()))
with open("answer4.txt", "w") as fout:
fout.write(str(cvs.mean()))
# 5.
#
# Полученный в пункте 4 классификатор - бэггинг на рандомизированных деревьях (в которых при построении каждой вершины выбирается случайное подмножество признаков и разбиение ищется только по ним). Это в точности соответствует алгоритму Random Forest, поэтому почему бы не сравнить качество работы классификатора с RandomForestClassifier из sklearn.ensemble. Затем изучите, как качество классификации на данном датасете зависит от количества деревьев, количества признаков, выбираемых при построении каждой вершины дерева, а также ограничений на глубину дерева. Для наглядности лучше построить графики зависимости качества от значений параметров, но для сдачи задания это делать не обязательно.
rf_classifier = RandomForestClassifier()
bagging = BaggingClassifier(rf_classifier, n_estimators=100)
cvs = cross_val_score(bagging, X, y, cv=10)
print(cvs)
print('Mean model quality value: ' + str(cvs.mean()))
with open("answer5.txt", "w") as fout:
answer = str(2) + ' ' + str(3) + ' ' + str(4) + ' ' + str(7)
fout.write(answer)
param_range = np.array([10, 50, 100, 150])
train_scores, test_scores = validation_curve(bagging, X, y, param_name="n_estimators", param_range=param_range, cv=10, scoring="accuracy")
train_scores_mean = np.mean(train_scores, axis=1)
train_scores_std = np.std(train_scores, axis=1)
test_scores_mean = np.mean(test_scores, axis=1)
test_scores_std = np.std(test_scores, axis=1)
print(train_scores_mean, test_scores_mean)
param_range = np.array([5, 10, 20, 40])
train_scores, test_scores = validation_curve(bagging, X, y, param_name="max_features", param_range=param_range, cv=10, scoring="accuracy")
train_scores_mean = np.mean(train_scores, axis=1)
train_scores_std = np.std(train_scores, axis=1)
test_scores_mean = np.mean(test_scores, axis=1)
test_scores_std = np.std(test_scores, axis=1)
print(train_scores_mean, test_scores_mean)
param_range = np.array([5, 10, 50, 100])
train_scores, test_scores = validation_curve(bagging, X, y, param_name="base_estimator__max_depth", param_range=param_range, cv=10, scoring="accuracy")
train_scores_mean = np.mean(train_scores, axis=1)
train_scores_std = np.std(train_scores, axis=1)
test_scores_mean = np.mean(test_scores, axis=1)
test_scores_std = np.std(test_scores, axis=1)
print(train_scores_mean, test_scores_mean)
| 2 Supervised learning/Homework/6 bagging and random forest/Bagging and random forest.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
from r2_summary import _r2_intra, _r2_inter, _r2_other
# # Seaborn parameters
sns.set_theme(style="whitegrid")
sns.set(font_scale=1.5)
# # File names: average R2 in the cortex, FWHM=5
# +
path_results = '/data/cisl/pbellec/cneuromod_embeddings/xp_202012/r2_friends-s01_cortex/'
fwhm = 5
# DYPAC
dypac60 = os.path.join(path_results, f'r2_fwhm-intra_fwhm-{fwhm}_cluster-20_state-60.p')
dypac120 = os.path.join(path_results, f'r2_fwhm-intra_fwhm-{fwhm}_cluster-20_state-120.p')
dypac150 = os.path.join(path_results, f'r2_fwhm-intra_fwhm-{fwhm}_cluster-50_state-150.p')
dypac300 = os.path.join(path_results, f'r2_fwhm-intra_fwhm-{fwhm}_cluster-50_state-300.p')
dypac900 = os.path.join(path_results, f'r2_fwhm-intra_fwhm-{fwhm}_cluster-300_state-900.p')
# DYPAC inter-subject
inter60 = os.path.join(path_results, f'r2_fwhm-inter_fwhm-{fwhm}_cluster-20_state-60.p')
inter120 = os.path.join(path_results, f'r2_fwhm-inter_fwhm-{fwhm}_cluster-20_state-120.p')
inter150 = os.path.join(path_results, f'r2_fwhm-inter_fwhm-{fwhm}_cluster-50_state-150.p')
inter300 = os.path.join(path_results, f'r2_fwhm-inter_fwhm-{fwhm}_cluster-50_state-300.p')
inter900 = os.path.join(path_results, f'r2_fwhm-inter_fwhm-{fwhm}_cluster-300_state-900.p')
# DIFUMO
difumo256 = os.path.join(path_results, f'r2_fwhm-difumo256_fwhm-{fwhm}.p')
difumo512 = os.path.join(path_results, f'r2_fwhm-difumo512_fwhm-{fwhm}.p')
difumo1024 = os.path.join(path_results, f'r2_fwhm-difumo1024_fwhm-{fwhm}.p')
# MIST
mist197 = os.path.join(path_results, f'r2_fwhm-mist197_fwhm-{fwhm}.p')
mist444 = os.path.join(path_results, f'r2_fwhm-mist444_fwhm-{fwhm}.p')
# Schaefer
schaefer = os.path.join(path_results, f'r2_fwhm-schaefer_fwhm-{fwhm}.p')
# Smith
smith70 = os.path.join(path_results, f'r2_fwhm-smith_fwhm-{fwhm}.p')
# -
# # DYPAC intra vs DIFUMO
# Comparing R2 quality (average in the cortex) between individual dypac900 and difumoXX (256, 512, 1024). The difumo parcellations are really impressive for group parcellations, but dypac individual has a systematic edge.
# +
val_r2 = pd.read_pickle(difumo256)
val_r2 = val_r2.append(pd.read_pickle(difumo512))
val_r2 = val_r2.append(pd.read_pickle(difumo1024))
val_r2 = val_r2.append(pd.read_pickle(dypac900))
fig = plt.figure(figsize=(20, 15))
sns.boxenplot(data=val_r2, x='subject', y='r2', hue='params', scale='area')
plt.ylabel('R2 embedding quality')
plt.title(f'FWHM={fwhm}')
# -
# # DYPAC intra vs MIST
# When comparing dypac (cluster-300_state-900) with more traditional approaches such as low dimensional ICA (Smith70) or static group parcellations (MIST, Schaefer) the gains are massive.
# +
val_r2 = pd.read_pickle(smith70)
val_r2 = val_r2.append(pd.read_pickle(mist197))
val_r2 = val_r2.append(pd.read_pickle(mist444))
val_r2 = val_r2.append(pd.read_pickle(schaefer))
val_r2 = val_r2.append(pd.read_pickle(dypac900))
fig = plt.figure(figsize=(20, 15))
sns.boxenplot(data=val_r2, x='subject', y='r2', hue='params', scale='area')
plt.ylabel('R2 embedding quality')
plt.title(f'FWHM={fwhm}')
# -
# # MIST multi resolution
# This figure directly investigates the impart of cluster and stateon the dypac R2. It makes clear that the final number of states is the primary driver of R2. But even with 60 states, dypac is competitive with the best static group atlases (with hundreds of parcels), and 120 states already outperforms them. But it takes 900 individuals dypac states to outperform difumo1024.
# +
val_r2 = pd.read_pickle(dypac60)
val_r2 = val_r2.append(pd.read_pickle(dypac120))
val_r2 = val_r2.append(pd.read_pickle(dypac150))
val_r2 = val_r2.append(pd.read_pickle(dypac300))
val_r2 = val_r2.append(pd.read_pickle(dypac900))
fig = plt.figure(figsize=(20, 15))
sns.boxenplot(data=val_r2, x='subject', y='r2', hue='params', scale='area')
plt.ylabel('R2 embedding quality')
plt.title(f'FWHM={fwhm}')
# -
# # intra vs inter subject R2
# This figure compares intra-subject embedding quality (parcellation and data come from the same subject) vs inter-subject embedding quality (parcellation and data come from different subject). Average R2 in the cortex is systematically higher intra-subject than inter-subject.
# +
val_r2 = pd.read_pickle(dypac60)
val_r2 = val_r2.append(pd.read_pickle(dypac120))
val_r2 = val_r2.append(pd.read_pickle(dypac150))
val_r2 = val_r2.append(pd.read_pickle(dypac300))
val_r2 = val_r2.append(pd.read_pickle(dypac900))
val_r2 = val_r2.append(pd.read_pickle(inter60))
val_r2 = val_r2.append(pd.read_pickle(inter120))
val_r2 = val_r2.append(pd.read_pickle(inter150))
val_r2 = val_r2.append(pd.read_pickle(inter300))
val_r2 = val_r2.append(pd.read_pickle(inter900))
fig = plt.figure(figsize=(20, 15))
sns.boxenplot(data=val_r2, x='params', y='r2', hue='type', scale='area')
plt.ylabel('R2 embedding quality')
plt.title(f'FWHM={fwhm}')
# -
# # File names: average R2 in the cortex, FWHM=8
# +
path_results = '/data/cisl/pbellec/cneuromod_embeddings/xp_202012/r2_friends-s01_cortex/'
fwhm = 8
# DYPAC
dypac60 = os.path.join(path_results, f'r2_fwhm-intra_fwhm-{fwhm}_cluster-20_state-60.p')
dypac120 = os.path.join(path_results, f'r2_fwhm-intra_fwhm-{fwhm}_cluster-20_state-120.p')
dypac150 = os.path.join(path_results, f'r2_fwhm-intra_fwhm-{fwhm}_cluster-50_state-150.p')
dypac300 = os.path.join(path_results, f'r2_fwhm-intra_fwhm-{fwhm}_cluster-50_state-300.p')
dypac900 = os.path.join(path_results, f'r2_fwhm-intra_fwhm-{fwhm}_cluster-300_state-900.p')
# DYPAC inter-subject
inter60 = os.path.join(path_results, f'r2_fwhm-inter_fwhm-{fwhm}_cluster-20_state-60.p')
inter120 = os.path.join(path_results, f'r2_fwhm-inter_fwhm-{fwhm}_cluster-20_state-120.p')
inter150 = os.path.join(path_results, f'r2_fwhm-inter_fwhm-{fwhm}_cluster-50_state-150.p')
inter300 = os.path.join(path_results, f'r2_fwhm-inter_fwhm-{fwhm}_cluster-50_state-300.p')
inter900 = os.path.join(path_results, f'r2_fwhm-inter_fwhm-{fwhm}_cluster-300_state-900.p')
# DIFUMO
difumo256 = os.path.join(path_results, f'r2_fwhm-difumo256_fwhm-{fwhm}.p')
difumo512 = os.path.join(path_results, f'r2_fwhm-difumo512_fwhm-{fwhm}.p')
difumo1024 = os.path.join(path_results, f'r2_fwhm-difumo1024_fwhm-{fwhm}.p')
# MIST
mist197 = os.path.join(path_results, f'r2_fwhm-mist197_fwhm-{fwhm}.p')
mist444 = os.path.join(path_results, f'r2_fwhm-mist444_fwhm-{fwhm}.p')
# Schaefer
schaefer = os.path.join(path_results, f'r2_fwhm-schaefer_fwhm-{fwhm}.p')
# Smith
smith70 = os.path.join(path_results, f'r2_fwhm-smith_fwhm-{fwhm}.p')
# -
# # DYPAC intra vs DIFUMO
# When repeating the experiment with `fwhm=8` the qualitative conclusions on dypac vs difumo are identical with `fwhm=5`. But a striking difference is a huge boost in R2 (almost doubling) for both parcelation. This shows that the R2 metric is very sensitive to the level of smoothness in the data.
# +
val_r2 = pd.read_pickle(difumo256)
val_r2 = val_r2.append(pd.read_pickle(difumo512))
val_r2 = val_r2.append(pd.read_pickle(difumo1024))
val_r2 = val_r2.append(pd.read_pickle(dypac900))
fig = plt.figure(figsize=(20, 15))
sns.boxenplot(data=val_r2, x='subject', y='r2', hue='params', scale='area')
plt.ylabel('R2 embedding quality')
plt.title(f'FWHM={fwhm}')
# -
# # DYPAC intra vs MIST
# The exact same observations hold for traditional parcellations: same qualitative conclusion for `fwhm=8` and `fwhm=5`, but near doubling of R2 with increased smoothing.
# +
val_r2 = pd.read_pickle(smith70)
val_r2 = val_r2.append(pd.read_pickle(mist197))
val_r2 = val_r2.append(pd.read_pickle(mist444))
val_r2 = val_r2.append(pd.read_pickle(schaefer))
val_r2 = val_r2.append(pd.read_pickle(dypac900))
fig = plt.figure(figsize=(20, 15))
sns.boxenplot(data=val_r2, x='subject', y='r2', hue='params', scale='area')
plt.ylabel('R2 embedding quality')
plt.title(f'FWHM={fwhm}')
# -
# # MIST multi resolution
# Once again with `fwhm=8` we see the number of `state` being a huge driver of R4. But we can also note that modest numbers of states (120, 150) are enough to reach high levels of R2 (0.5), while 900 states provide very high R2 (0.7). The low resolution solutions are thus an accurate summary of fluctuations at low spatial resolution. So even if the R2 of 120 and 150 states is comparatively lower with `fwhm=5` they still capture important characteristics of the data, and should be investigated in parallel to a granular and high precision solution (`cluster-300_state-900`).
# +
val_r2 = pd.read_pickle(dypac60)
val_r2 = val_r2.append(pd.read_pickle(dypac120))
val_r2 = val_r2.append(pd.read_pickle(dypac150))
val_r2 = val_r2.append(pd.read_pickle(dypac300))
val_r2 = val_r2.append(pd.read_pickle(dypac900))
fig = plt.figure(figsize=(20, 15))
sns.boxenplot(data=val_r2, x='subject', y='r2', hue='params', scale='area')
plt.ylabel('R2 embedding quality')
plt.title(f'FWHM={fwhm}')
# -
# ## Intra vs inter subject R2
# Same conclusion for `fwhm=8` and `fwhm=5`: intra-subject R2 is markedly superior to inter-subject R2. However the gap tightens with `fwhm=8`.
# +
val_r2 = pd.read_pickle(dypac60)
val_r2 = val_r2.append(pd.read_pickle(dypac120))
val_r2 = val_r2.append(pd.read_pickle(dypac150))
val_r2 = val_r2.append(pd.read_pickle(dypac300))
val_r2 = val_r2.append(pd.read_pickle(dypac900))
val_r2 = val_r2.append(pd.read_pickle(inter60))
val_r2 = val_r2.append(pd.read_pickle(inter120))
val_r2 = val_r2.append(pd.read_pickle(inter150))
val_r2 = val_r2.append(pd.read_pickle(inter300))
val_r2 = val_r2.append(pd.read_pickle(inter900))
fig = plt.figure(figsize=(20, 15))
sns.boxenplot(data=val_r2, x='params', y='r2', hue='type', scale='area')
plt.ylabel('R2 embedding quality')
plt.title(f'FWHM={fwhm}')
| notebooks/fig_r2_friends-s01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.8 64-bit (''conda'': virtualenv)'
# name: python388jvsc74a57bd098b0a9b7b4eaaa670588a142fd0a9b87eaafe866f1db4228be72b4211d12040f
# ---
# # Working with LAS and PCD Files
#
# It's possible to read pointcloud files with the format .las and .pcd. In order to read them just the submodule PointCloud.from_file():
import pointcloudset as pcs
from pathlib import Path
import numpy as np
import laspy
from laspy.file import File
testpcd = Path().cwd().parent.joinpath("../../../tests/testdata/las_files/test_tree.pcd")
testlas = testpcd = Path().cwd().parent.joinpath("../../../tests/testdata/las_files/test_tree.las")
las_pc = pcs.PointCloud.from_file(testlas)
pcd_pc = pcs.PointCloud.from_file(testpcd)
laspy_las = File(testlas, mode = "rw")
print ("Scale-data of pointcloud: ", laspy_las.header.scale)
print ("Offset-data of pointcloud: ", laspy_las.header.offset)
# ### Note:
# Coordinates might not be correct yet, since the offset and scale values that are stored within the .las-file are not applied.
# But now you can use the data as a pcs.PointCloud and analyze + edit it.
#
las_pc.data
las_pc.plot(color = "z")
# +
def treetop(frame: pcs.PointCloud) -> pcs.PointCloud:
return frame.limit("z", 8, 19)
tip = treetop(las_pc)
# -
tip.plot(color = "z")
| doc/sphinx/source/tutorial_notebooks/reading_las_pcd.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Why is knowing the variance of WBO with respect to conformation important?
# 1. WBO is a function of the electronic density which is a function of the conformation. But we want to use the WBO as a criterion for fragmenting molecules and we don't want to generate many conformations for this step because it's too expensive.
# 2. It seems like the variance is higher for more conjugated bonds which we are interested in. However, if the distribution of those bond orders are around the cutoff value, how do we choose the cuttoff? While in general the higher variance bonds also have higher WBO (above the cutoff), I found that certain bonds are ambiguous. The variance is on the hihger side and the WBO is distributed around the cutoff value.
#
#
# Turns out, while the standard deviations are pretty low (0.08 is at the higher end), some of the more ambigous bonds
# that have high bond order correlations with neighboring rings or other bonds, have a slightly higher variance (not as high as the aromatics in the rings). The most troubling thing is that the distribution is over where the WBO is. (1.05-1.15). So if we continue with a hard cutoff in that range, we might miss some of these because of the conformation.
# Examples of this:
# (These are not obvious ones like amides which we'll just tag)
# Most of these are nitrogens next to aromatic rings. Ethers next to aromatic rings tend to have lower WBO but also higher variance
#
#
# * Abmaciclib bond (12, 13)
# * Afitinib (16, 20)
# * Bosutinib (13, 14), (7,13)
# * Brigatinib (11, 14), (20, 21), (25, 27)
# * Ceritinib (18, 19), (23, 25) ?? (18,4), (5, 14) (these are lower BO but high variance)
# * Desatinib (4, 6), (9, 14) ?? (6, 7) (low WBO but high variance)
# * Erlotinib (20, 21), ?? (11, 12), (6, 5) (low WBO but high variance - ether next to aromatic ring)
# * Gefitinib (19, 23) ?? (3, 2), (8, 9) (low WBO but high variance - ether next to aromatic rings)
# - This is a good example for figure / illustration
# * Ibrutinib (5, 3) technically an amide
# * Idelalisib (23, 24)
# * Imatinib (10, 11)
# - Decent example for the N attached to aromatic ring, aromatic rings and amide.
# * Lapatinib (20, 24) ?? (24, 25) lower WBO but higher variance
# * Neratinib (17, 18)
# * Nilotininb is a mess - clean up the data
# * Nintedanib (19, 20),
# * Osimertinib (13, 17), (17, 18), (22, 24),
# * Palbociclib (25, 28), (21, 6)
# * Pazopanib (15, 13) ?? (8, 9), (5, 8), (28, 31) lower WBO but higher variance
# * Regorafenib ?? (15, 18)
# * Ribociclib (20, 11), (24, 27), ?? (20, 21) lower WBO but higher variance
# * Sunitinib (13, 17)
# - This is a good example for figure / illustration for a conjugated bond next to a doubl bond.
# * Tofacitinib (10, 12), (6, 21 ) - amide attached to a ring
# * Vandetanib (18, 22) ?? (22, 23) lower WBO but higher variance, (9, 10) (15, 16) (ethers attached to aromatic rings)
# * Vemurafenib ?? (8, 7), (7, 4) lower WBO but higher variance.
#
#
# I only looked at psi4 optimized WBO. I also want to look at:
# * Psi4 Mayer unoptimized
# * Psi4 Mayer optimized goemetry
# * Psi4 Wiberg unoptimized
# * OE Wiberg bond orders
#
#
# %matplotlib inline
import matplotlib.pyplot as plt
from fragmenter import chemi
from openeye import oechem
import oenotebook as oenb
import json
import numpy as np
import glob
import cmiles
import os
from matplotlib.backends.backend_pdf import PdfPages
import seaborn as sbn
import pandas as pd
import matplotlib.image as mpimg
directories = [x[0] for x in os.walk('../conjugation/geometry_opt/')][1:]
output = {}
for kinase_inhibitor in directories:
ki_key = kinase_inhibitor.split('/')[-1]
output[ki_key] = []
output_files = glob.glob('../conjugation/geometry_opt/{}/*.bo.json'.format(ki_key))
for file in output_files:
with open(file, 'r') as f:
data = json.load(f)
error = data['error']
if error:
print(error)
continue
data['bond_orders'] = chemi.bond_order_from_psi4_raw_output(data['raw_output'])
data.pop('raw_output')
output[ki_key].append(data)
def collect_all_std(data, hydrogen=False, rings=False, only_rings=False, halogens=False, carbonyls=False, nitriles=False):
bond_order_std = {}
for ki in data:
if len(data[ki]) == 0:
continue
bond_order_std[ki] = {}
conformations = len(data[ki])
mol = cmiles.utils.load_molecule(data[ki][0]['tagged_smiles'])
n_atoms = mol.NumAtoms()
bond_order_wiberg = {}
bond_order_mayer = {}
for bond in mol.GetBonds():
if only_rings and not rings:
raise RuntimeError("If only rings is True, rings must be true")
if only_rings:
if not bond.IsInRing():
continue
if not rings:
if bond.IsInRing():
continue
atom_1 = bond.GetBgn()
atom_2 = bond.GetEnd()
if not hydrogen:
if atom_1.IsHydrogen() or atom_2.IsHydrogen():
continue
if not halogens:
if atom_1.IsHalogen() or atom_2.IsHalogen():
continue
if not nitriles:
if bond.GetOrder() == 3:
continue
if not carbonyls:
if bond.GetOrder() == 2 and (atom_1.IsOxygen() or atom_2.IsOxygen()):
continue
map_1 = atom_1.GetMapIdx()
map_2 = atom_2.GetMapIdx()
bond_order_wiberg[(map_1, map_2)] = np.zeros(conformations)
bond_order_mayer[(map_1, map_2)] = np.zeros(conformations)
# Populate array
for k, d in enumerate(data[ki]):
wiberg = d['bond_orders']['Wiberg_psi4']
mayer = d['bond_orders']['Mayer_psi4']
for i, j in bond_order_wiberg:
bond_order_wiberg[(i, j)][k] = wiberg[i-1][j-1]
bond_order_mayer[(i, j)][k] = mayer[i-1][j-1]
bond_order_std[ki]['wiberg_bo'] = bond_order_wiberg
bond_order_std[ki]['mayer_bo'] = bond_order_mayer
# calculate variance
bonds = []
mayer_std = []
wiberg_std = []
for bond in bond_order_mayer:
bonds.append(bond)
mayer_std.append(np.std(bond_order_mayer[bond]))
wiberg_std.append(np.std(bond_order_wiberg[bond]))
bond_order_std[ki]['wiberg_std'] = wiberg_std
bond_order_std[ki]['mayer_std'] = mayer_std
bond_order_std[ki]['bonds'] = bonds
return bond_order_std
bond_order_std_of_interest = collect_all_std(output)
bond_order_std_rings = collect_all_std(output, only_rings=True, rings=True)
# all bond orders
bond_order_std = collect_all_std(output, rings=True, hydrogen=True, halogens=True, nitriles=True, carbonyls=True)
bond_order_std_no_rings = collect_all_std(output, rings=False, hydrogen=False, halogens=True, nitriles=True, carbonyls=True)
bond_order_std_no_h = collect_all_std(output, rings=True, hydrogen=False, halogens=True, nitriles=True, carbonyls=True)
# write out xyz files on confmorations for viewing in Pymol
for ki in output:
if len(output[ki]) == 0:
continue
mol = cmiles.utils.load_molecule(output[ki][0]['tagged_smiles'])
n_atoms = mol.NumAtoms()
xyz = ""
for o in output[ki]:
molecule = o['molecule']
xyz += '{}\n'.format(n_atoms)
xyz+= '{}\n'.format(ki)
xyz += molecule
xyz += '\n'
with open('optimized_conformations/{}.xyz'.format(ki), 'w') as f:
f.write(xyz)
# plot std of bond orders for each molecule
with PdfPages('Bond_order_std.pdf') as pdf:
for ki in bond_order_std:
plt.figure()
plt.hist(bond_order_std[ki]['wiberg_std'], alpha=0.5, label='Wiberg, optimized');
plt.hist(bond_order_std[ki]['mayer_std'], alpha=0.5, label='mayer, optimized');
plt.legend()
bond = bond_order_std[ki]['bonds'][0]
confs = len(bond_order_std[ki]['wiberg_bo'][bond])
plt.title('{}, {} conformations'.format(ki, confs));
plt.xlabel('Standard Deviation')
plt.ylabel('counts')
pdf.savefig()
plt.close()
for ki in bond_order_std:
with PdfPages('std/{}_std_box_plots.pdf'.format(ki)) as pdf:
plt.figure()
plt.hist(bond_order_std[ki]['wiberg_std'], alpha=0.5, label='Wiberg, optimized');
plt.hist(bond_order_std[ki]['mayer_std'], alpha=0.5, label='mayer, optimized');
plt.legend()
bond = bond_order_std[ki]['bonds'][0]
confs = len(bond_order_std[ki]['wiberg_bo'][bond])
plt.title('{}, {} conformations'.format(ki, confs));
plt.xlabel('Standard Deviation')
plt.ylabel('counts')
pdf.savefig()
plt.close()
img = mpimg.imread('../conjugation/bond_order_without_geomopt/{}_mapped.png'.format(ki))
plt.figure()
imgplot = plt.imshow(img, interpolation='none')
plt.xticks([])
plt.yticks([])
pdf.savefig()
plt.close()
df = pd.DataFrame(bond_order_std_of_interest[ki]['wiberg_bo'])
plt.figure()
#sbn.boxplot(data=df, orient='h', fliersize=0.8, linewidth=0.8);
ax = sbn.boxplot(data=df, orient='h',fliersize=0.9, linewidth=0.8)
sbn.stripplot(data=df, size=0.8, jitter=True, orient='h', color='black')
plt.title('{}, rotatable bonds, {} conformations'.format(ki, confs));
plt.yticks(fontsize=7)
plt.xlabel('Bond order')
plt.ylabel('Bond indices')
pdf.savefig(dpi=400, bbox_inches='tight')
plt.close()
df = pd.DataFrame(bond_order_std_rings[ki]['wiberg_bo'])
plt.figure()
ax = sbn.boxplot(data=df, orient='h', fliersize=0.9, linewidth=0.8);
#ax = sbn.boxplot(data=df, orient='h', linewidth=0.8)
sbn.stripplot(data=df, size=0.8, jitter=True, orient='h', color='black')
plt.title('{}, ring bonds, {} conformations'.format(ki, confs));
plt.yticks(fontsize=7)
plt.xlabel('Bond order')
plt.ylabel('Bond indices')
pdf.savefig(dpi=400, bbox_inches='tight')
plt.close()
all_std = []
all_bo_rings = []
for ki in bond_order_std_rings:
for bond in bond_order_std_rings[ki]['wiberg_bo']:
all_bo_rings.extend(bond_order_std_rings[ki]['wiberg_bo'][bond])
#all_std_rings.extend(bond_order_std_rings[ki]['wiberg_std'])
all_bo_no_rings = []
for ki in bond_order_std_no_rings:
for bond in bond_order_std_no_rings[ki]['wiberg_bo']:
all_bo_no_rings.extend(bond_order_std_no_rings[ki]['wiberg_bo'][bond])
#all_std_no_rings.extend(bond_order_std_no_rings[ki]['wiberg_std'])
for ki in bond_order_std_no_h:
all_std.extend(bond_order_std_no_h[ki]['wiberg_std'])
import seaborn as sbn
plt.figure(dpi=200)
sbn.distplot(all_bo_rings, label='rings')
sbn.distplot(all_bo_no_rings, label='rotatable')
#plt.hist(all_bo_rings, alpha=0.5, label='rings');
#plt.hist(all_bo_no_rings, alpha=0.5, label='rotatable');
plt.legend()
plt.title('Histogram of WBO (psi4)')
plt.xlabel('Wiberg Bond Oder')
plt.ylabel('Counts');
plt.savefig('Wiberg_psi4_hist.pdf')
plt.figure(dpi=400)
#plt.hist(all_std);
sbn.distplot(all_std)
plt.title('Histogram of WBO std (psi4) ~3,500 conformations')
plt.xlabel('standard deviation')
plt.ylabel('Counts');
plt.savefig('wiberg_psi4_all_std.pdf')
# Look at variance of the non optimized molecules
directories = [x[0] for x in os.walk('../conjugation/bond_order_without_geomopt/')][1:]
output_no_opt = {}
for kinase_inhibitor in directories:
ki_key = kinase_inhibitor.split('/')[-1]
output_no_opt[ki_key] = []
output_files = glob.glob('../conjugation/bond_order_without_geomopt/{}/*.output.json'.format(ki_key))
for file in output_files:
with open(file, 'r') as f:
data = json.load(f)
error = data['error']
if error:
print(error)
continue
data['bond_orders'] = chemi.bond_order_from_psi4_raw_output(data['raw_output'])
data.pop('raw_output')
output_no_opt[ki_key].append(data)
confs = 0
for ki in output:
confs += (len(output[ki]))
confs
means = []
stds = []
for ki in bond_order_std_no_h:
for bond in bond_order_std_no_h[ki]['bonds']:
mean = (bond_order_std_no_h[ki]['wiberg_bo'][bond].mean())
if mean > 2.8 or mean < 1.0:
continue
means.append(mean)
stds.append(np.std(bond_order_std_no_h[ki]['wiberg_bo'][bond].mean()))
sbn.scatterplot(means, stds)
#plt.scatter(means, stds, markersize=0.1);
#plt.ylim(-0.0000001, 0.0000001)
np.corrcoef(means, stds)
plt.hist(stds)
max(all_std)
| bond_order_variance/bond_order_std.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
# # 592B, Class 8.1 (03/20). Midterm Review: how do you compute a spectrum/spectrogram?
import numpy as np
import matplotlib.pyplot as plt
import librosa
from scipy import fftpack #new
from scipy import signal
from ipywidgets import interactive
from IPython.display import Audio, display
# ## Intro: spectrogram tutorial
#
# Clone the [Github repository here](https://github.com/drammock/spectrogram-tutorial) and work through it. Try using a different wave file than the one used as a sample. Also compute a spectrogram/spectrograms using Praat for comparison. And try computing a narrow-band spectrogram, too.
#
# Some things to think about and answer:
# - What library and function is being used to compute the spectrogram here and how do you find out more about it?
# - What choices involved in computing a spectrogram also come up for computing a spectrum? What choices only come up for computing a spectrogram and not the spectrum?
# - What would ringing look like in the spectrum?
# - How do you compute a wide-band spectrogram? How do you compute a narrow-band spectrogram?
#
# ---
# ## Intro 2: windowing/leakage tutorial
#
# Work through the Elegant SciPy tutorial section on [windowing](https://www.safaribooksonline.com/library/view/elegant-scipy/9781491922927/ch04.html#windowing). (You can copy and paste code from there into this notebook.)
#
# Some things to think about/answer:
# - What happens to the spectrum when you change the width of the rectangular pulse?
# - Can you write a function so you can easily vary the width of the rectangular pulse and make the two plots?
# - Why do we do windowing?
# - How can you window with a Gaussian window rather than a Kaiser window?
# - How is windowing for computing the spectrum related to windowing in computing the spectrogram?
# ## Intro 2.5: Convolution. What really is windowing anyway?
#
# Last time we said that windowing, filtering, and smoothing, are the same operations from a mathematical perspective. That operation is called **convolution**. The convolution of two functions $f(t)$ and $g(t)$ is defined as:
#
# $ f * g = \int_{-\infty}^{\infty} f(\tau)g(t-\tau)d\tau $
#
# The motto that goes with this is: flip and shift and compute the overlapping area.
#
# Here are some examples:
#
# A rectangular pulse with itself:
# 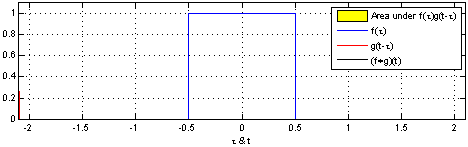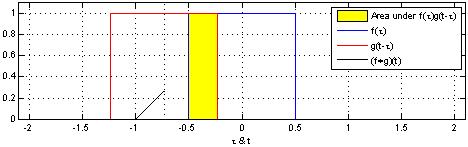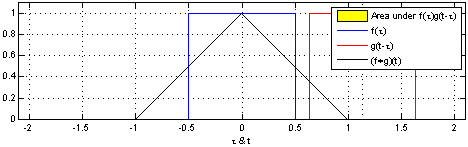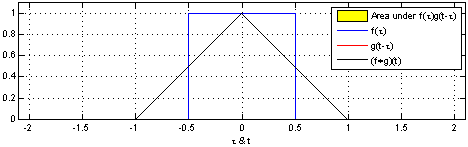
#
# 
#
# A rectangular pulse with a spiky function:
# 
#
# 
#
#
# And a Gaussian with a Gaussian:
#
# 
# ## Intro 3: Time-limited signals and window length
#
# Do the exercises I found [here](https://www.gaussianwaves.com/2011/01/fft-and-spectral-leakage-2/). Note that the code is in Matlab; you'll need to port to Python and modify as necessary.
#
# Things to think about and answer:
#
# - What causes spectral leakage?
# ## Intro 4: theoretical underpinnings
#
# Remember that we ended up with the Fourier series of $g(t)$ defined as $T \rightarrow \infty$, expressed as a double integral:
#
# \begin{equation}
# g(t) = \int_{-\infty}^{\infty} \int_{-\infty}^{\infty}e^{-2\pi ift}g(t)dt e^{2\pi ift}df
# \end{equation}
#
# and then we derived $\mathcal{F}g$, where $\mathcal{F}g$ is defined as the Fourier transform of a function $g(t)$:
#
# \begin{equation}
# \mathcal{F}g = \int_{-\infty}^{\infty}e^{-2\pi ift} g(t)dt
# \end{equation}
#
# and the inverse Fourier transform $\mathcal{F}^{-1}(t)$ of a function $\mathcal{F}g(f)$ is:
#
# \begin{equation}
# g(t) = \int_{-\infty}^{\infty}e^{2\pi ift} \mathcal{F}gdf
# \end{equation}
#
# Then we briefly introduced the **discrete Fourier transform (DFT)**: this is what we use when we're computing the Fourier Transform in python because we are working with *digital* signals.
#
#
# ## The Discrete Fourier Transform
#
# Today we're going to work on understanding the DFT a little more, because it will help us understand what's going on when we invoke `fftpack.fft` and have a deeper understanding of the answers to the review questions above. The derivation here follows Osgood Chapter 6.
#
# Our goal is to find a discrete version of $\mathcal{F}g(f)$, the Fourier transform of a function $g(t)$. (Note: by writing $\mathcal{F}g(f)$, we mean that the function $\mathcal{F}g$, the Fourier transform of $g(t)$, is a function of frequency, $f$. We start with $g(t)$, which is a function of time $t$, but once we take the Fourier transform of $g(t)$, $\mathcal{F}g$, we have a function of frequency $f$.).
#
# To do this, we need to find three things:
# - A discrete version of $g(t)$ that reasonably approximates $g(t)$
# - A discrete version of $\mathcal{F}g(f)$ that reasonably approximates $\mathcal{F}g(f)$
# - A way in which these two discrete versions are related, which approximates the relation between the continuous versions
#
# We start by assuming that $g(t)$ is:
# - *time-limited*, meaning that $g(t)$ is zero outside of $0\leq t \leq L$, where $L$ is the length of the signal (in time)
# - *band-limited*, meaning that $\mathcal{F}g(f)$ vanishes outside of $0 \lt f \lt 2B$, where $B$ is the *bandwith* of the signal.
#
#
# By the Sampling Theorem, if we sample $g(t)$ at the Nyquist rate of $2B$ samples/second, we can reconstruct $g(t)$ perfectly. This sampled version of $g(t)$, call it $g_{discrete}(t)$, is just a list of $N$ sampled values:
#
# $$ g(t_0), g(t_1), \ldots, g(t_{N-1}) $$,
#
# where $N=2BL$ and the timepoints are evenly spaced apart by $\frac{1}{2B}$.
#
# ***In-class exercise: Why is $N=2BL$ and why does $\Delta t_n = \frac{1}{2B}$?***
#
# ---
#
# ### The Dirac comb
#
# We can re-express $g_{discrete}(t)$ using the Dirac comb $III(t)$, defined as:
#
# $$III(t) = \displaystyle\sum_{n=0}^{N-1} \delta(t-t_n) $$
#
# All this is is a "train" of pulses, a comb of "lollipops" with amplitude 1, where the pulses occur exactly at the sampled points $t_0, t_1, \ldots, t_{N-1}$.
#
# Here's an image of a Dirac comb from Wikipedia. In our case, $T = 1/2B$.
#
# <img alt = "Dirac comb plot" src="https://upload.wikimedia.org/wikipedia/commons/4/49/Dirac_comb.svg" width="300" />
#
#
# And here's an example of [one way to define a Dirac comb function](https://scipython.com/book/chapter-6-numpy/examples/a-comb-function/), from [scipython.com](https://scipython.com).
#
# +
N, n = 101, 5
def f(i):
return (i % n == 0) * 1
comb = np.fromfunction(f, (N,), dtype=int)
print(comb)
# -
# ***In-class discussion: Does the Dirac comb form an orthogonal basis set?***
# ---
#
# Using $III(t)$, we can now express $g_{discrete}(t)$ as:
#
# $$g_{discrete}(t) = g(t) \displaystyle\sum_{n=0}^{N-1} \delta(t-t_n) = \displaystyle\sum_{n=0}^{N-1} g(t) \delta(t-t_n) $$
#
# And the Fourier transform of $g_{discrete}(t)$ is:
#
# \begin{equation}
# \mathcal{F}g_{discrete}(f) = \displaystyle\sum_{n=0}^{N-1} = \mathcal{F}g(t_n) \delta(t-t_n) = \displaystyle\sum_{n=0}^{N-1} g(t_n) e^{-2\pi ift_n}
# \end{equation}
#
# This gives us the continuous Fourier transform of the sampled version of $g(t)$.
#
# Now let's think about $g(t)$ in the frequency domain. Remember by assumption that $g(t)$ is time-limited so $g(t)$ is zero outside of $0\leq t \leq L$, where $L$ is the length of the signal (in time). So we can apply the Sampling Theorem to reconstruct $\mathcal{F}g(f)$ in the frequency domain. The sampling rate we need (the Nyquist rate) for perfect reconstruction is $L$ samples/Hz and the spacing between sampling points is $1/L$.
#
# Since $\mathcal{F}g(f)$ is band-limited by assumption and vanishes outside of $0 \lt f \lt 2B$, we sample $\mathcal{F}g(f)$ over $0 \lt f \lt 2B$, with points $1/L$ Hz apart.
#
# ---
#
# ***In-class exercise: Why is the sampling rate $L$ samples/Hz and why is the interval between sampling points $1/L$ Hz? What is the total number of sampling points, $N$***
#
# ---
#
# This sampled version of $\mathcal{F}g(f)$, call it $\mathcal{F}g_{discrete}(f)$, is just a list of $N$ sampled values, of the form $m/L$, where $m$ is a non-negative integer:
#
# $$ f_0=0,\, f_1 = \frac{1}{L},\, \ldots, f_{N-1} = \frac{N-1}{L} $$,
#
# And if we want the discrete version of $g_{discrete}(t)$, then we want to take $[\mathcal{F}(g_{discrete})](f)$, call this $F(f)$ for short.
#
# Taking our definition of the Fourier transform of $g_{discrete}(t)$,
#
# $$\mathcal{F}g(t_n) \delta(t-t_n) = \displaystyle\sum_{n=0}^{N-1} g(t_n) e^{-2\pi ift_n}$$
#
# this will give us the list:
#
# $$ F(f_0) = \displaystyle\sum_{n=0}^{N-1} g(t_n) e^{-2\pi if_0t_n}, \ldots, F(f_{N-1})=\displaystyle\sum_{n=0}^{N-1} g(t_n) e^{-2\pi if_{N-1}t_n} $$
#
# And so now we have a way to go from $g_{discrete}(t)$ to $\mathcal{F}g_{discrete}(f)$, for each $m$ from $m=0$ to $m=N-1$:
#
# $$F(f_m) = \displaystyle\sum_{n=0}^{N-1} g(t_n) e^{-2\pi if_mt_n} $$
#
# Recalling that $t_n = \frac{n}{2B}$ and $f_m = \frac{m}{L}$ and $N=2BL$, we can re-write this as:
#
# $$F(f_m) = \displaystyle\sum_{n=0}^{N-1} g(t_n) e^{-2\pi inm/N} $$
#
# ***In-class exercise: derive our final expression of $F(f_m)$.***
#
# ---
#
# At this point, let's come back to one of our starting questions and discuss. You should have more insight on this now! What is the "grid" spacing in the time-domain? The frequency domain? How are they related?
#
# > Why is the computation of the spectrum affected by the "window length" over which it is computed, and how is it affected?
#
# ---
# ## Positive and negative frequencies (Osgood 2010, p. 260)
#
# Given our discrete Fourier transform $[\mathcal{F}(g_{discrete})](f)$, call this $F(f)$:
#
# $$F(f_m) = \displaystyle\sum_{n=0}^{N-1} g(t_n) e^{-2\pi inm/N} $$
#
# it turns out that the spectrum *splits* at $N/2$. See Osgood (2010) for the derivation, but due to some periodicity relations:
#
# $$ F[\frac{N}{2} + 1] = \overline{F[\frac{N}{2} - 1]}$$
# $$ F[\frac{N}{2} + 2] = \overline{F[\frac{N}{2} - 2]}$$
# $$ \vdots$$
#
# ***In-class exercise: What is F[0]? What do the periodicity relations mean geometrically?***
#
# So because of this, the convention is to say, for a spectrum indexed from 0 to $N-1$:
# - The frequencies from $m=1$ to $m= N/2-1$ are the "positive" frequencies
# - The frequencies from $m=N/2+1$ to $m= N-1$ are the "negative" frequencies
#
# For a real signal, all the information you need is in the positive frequencies and the first component $F[0]$.
#
#
| 592B Spring 2018 Class 8.1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] papermill={"duration": 0.015389, "end_time": "2021-04-19T18:06:07.686676", "exception": false, "start_time": "2021-04-19T18:06:07.671287", "status": "completed"} tags=[]
# # Classifier Training
# + _kg_hide-output=true papermill={"duration": 153.045874, "end_time": "2021-04-19T18:08:40.747195", "exception": false, "start_time": "2021-04-19T18:06:07.701321", "status": "completed"} tags=[]
# ! rsync -a /kaggle/input/mmdetection-v280/mmdetection /
# ! pip install /kaggle/input/mmdetection-v280/src/mmpycocotools-12.0.3/mmpycocotools-12.0.3/
# ! pip install /kaggle/input/hpapytorchzoo/pytorch_zoo-master/
# ! pip install /kaggle/input/hpacellsegmentation/HPA-Cell-Segmentation/
# ! pip install /kaggle/input/iterative-stratification/iterative-stratification-master/
# ! cp -r /kaggle/input/kgl-humanprotein-data/kgl_humanprotein_data /
# ! cp -r /kaggle/input/humanpro/kgl_humanprotein /
import sys
sys.path.append('/kgl_humanprotein/')
# + papermill={"duration": 4.781661, "end_time": "2021-04-19T18:08:45.553844", "exception": false, "start_time": "2021-04-19T18:08:40.772183", "status": "completed"} tags=[]
import os
import time
from pathlib import Path
import shutil
import zipfile
import functools
import multiprocessing
import numpy as np
import pandas as pd
import cv2
from sklearn.model_selection import KFold,StratifiedKFold
from iterstrat.ml_stratifiers import MultilabelStratifiedKFold
import torch
from torch.backends import cudnn
from torch.utils.data import Dataset, DataLoader, RandomSampler, SequentialSampler
from torch.nn import DataParallel
import matplotlib.pyplot as plt
from tqdm import tqdm
from kgl_humanprotein.utils.common_util import *
from kgl_humanprotein.config.config import *
from kgl_humanprotein.data_process import *
from kgl_humanprotein.datasets.tool import image_to_tensor
from kgl_humanprotein.networks.imageclsnet import init_network
from kgl_humanprotein.layers.loss import *
from kgl_humanprotein.layers.scheduler import *
from kgl_humanprotein.utils.augment_util import train_multi_augment2
from kgl_humanprotein.utils.log_util import Logger
from kgl_humanprotein.run.train import *
# + papermill={"duration": 0.036392, "end_time": "2021-04-19T18:08:45.614532", "exception": false, "start_time": "2021-04-19T18:08:45.578140", "status": "completed"} tags=[]
# %cd /kaggle
# + [markdown] papermill={"duration": 0.025439, "end_time": "2021-04-19T18:08:45.665953", "exception": false, "start_time": "2021-04-19T18:08:45.640514", "status": "completed"} tags=[]
# ## Combine subsets' meta data
# + papermill={"duration": 0.034671, "end_time": "2021-04-19T18:08:45.727353", "exception": false, "start_time": "2021-04-19T18:08:45.692682", "status": "completed"} tags=[]
dir_data = Path('/kaggle/input')
dir_mdata = Path('/kaggle/mdata')
# + papermill={"duration": 4.054318, "end_time": "2021-04-19T18:08:49.807026", "exception": false, "start_time": "2021-04-19T18:08:45.752708", "status": "completed"} tags=[]
# %%time
# df_cells = combine_subsets_metadata(dir_data, n_subsets)
df_cells = pd.read_feather('/kaggle/input/humanpro-data-multilabel-cells-meta/train.feather')
# + papermill={"duration": 1.871073, "end_time": "2021-04-19T18:08:51.704200", "exception": false, "start_time": "2021-04-19T18:08:49.833127", "status": "completed"} tags=[]
dir_mdata_raw = dir_mdata/'raw'
dir_mdata_raw.mkdir(exist_ok=True, parents=True)
df_cells.to_feather(dir_mdata_raw/'train.feather')
# + papermill={"duration": 0.203269, "end_time": "2021-04-19T18:08:51.937914", "exception": false, "start_time": "2021-04-19T18:08:51.734645", "status": "completed"} tags=[]
del df_cells
# + [markdown] papermill={"duration": 0.025573, "end_time": "2021-04-19T18:08:51.989690", "exception": false, "start_time": "2021-04-19T18:08:51.964117", "status": "completed"} tags=[]
# ## Filter samples
# + papermill={"duration": 1.432776, "end_time": "2021-04-19T18:08:53.447671", "exception": false, "start_time": "2021-04-19T18:08:52.014895", "status": "completed"} tags=[]
# Remove samples whose target is ''
df_cells = pd.read_feather(dir_mdata_raw/'train.feather')
df_cells = df_cells[df_cells['Target'] != ''].reset_index(drop=True)
# + papermill={"duration": 0.039772, "end_time": "2021-04-19T18:08:53.514481", "exception": false, "start_time": "2021-04-19T18:08:53.474709", "status": "completed"} tags=[]
df_cells.shape
# + papermill={"duration": 0.036578, "end_time": "2021-04-19T18:08:53.577983", "exception": false, "start_time": "2021-04-19T18:08:53.541405", "status": "completed"} tags=[]
# Limit number of samples per label
def cap_number_per_label(df_cells, cap=10_000, idx_start=0):
df_cells_cap = pd.DataFrame()
for label in df_cells.Target.unique():
df = df_cells[df_cells.Target==label]
if len(df) > cap:
df = df.iloc[idx_start:idx_start + cap]
df_cells_cap = df_cells_cap.append(df, ignore_index=True)
return df_cells_cap
# + papermill={"duration": 9.891073, "end_time": "2021-04-19T18:09:03.496276", "exception": false, "start_time": "2021-04-19T18:08:53.605203", "status": "completed"} tags=[]
df_cells = cap_number_per_label(df_cells, cap=5_000, idx_start=0)
# + papermill={"duration": 0.065361, "end_time": "2021-04-19T18:09:03.595803", "exception": false, "start_time": "2021-04-19T18:09:03.530442", "status": "completed"} tags=[]
df_cells.Target.value_counts()
# + papermill={"duration": 1.513232, "end_time": "2021-04-19T18:09:05.140622", "exception": false, "start_time": "2021-04-19T18:09:03.627390", "status": "completed"} tags=[]
df_cells.to_feather(dir_mdata_raw/'train.feather')
# + [markdown] papermill={"duration": 0.029036, "end_time": "2021-04-19T18:09:05.199246", "exception": false, "start_time": "2021-04-19T18:09:05.170210", "status": "completed"} tags=[]
# ## One-hot encode labels
# + papermill={"duration": 23.020097, "end_time": "2021-04-19T18:09:28.248702", "exception": false, "start_time": "2021-04-19T18:09:05.228605", "status": "completed"} tags=[]
# %%time
generate_meta(dir_mdata, 'train.feather')
# + [markdown] papermill={"duration": 0.028548, "end_time": "2021-04-19T18:09:28.306138", "exception": false, "start_time": "2021-04-19T18:09:28.277590", "status": "completed"} tags=[]
# ## Split generation
# + papermill={"duration": 7.075065, "end_time": "2021-04-19T18:09:35.410308", "exception": false, "start_time": "2021-04-19T18:09:28.335243", "status": "completed"} tags=[]
# %%time
train_meta = pd.read_feather(dir_mdata/'meta'/'train_meta.feather')
create_random_split(dir_mdata, train_meta, n_splits=5, alias='random')
del train_meta
# + [markdown] papermill={"duration": 0.029681, "end_time": "2021-04-19T18:09:35.472968", "exception": false, "start_time": "2021-04-19T18:09:35.443287", "status": "completed"} tags=[]
# ## Training
# + papermill={"duration": 0.080825, "end_time": "2021-04-19T18:09:35.583655", "exception": false, "start_time": "2021-04-19T18:09:35.502830", "status": "completed"} tags=[]
from kgl_humanprotein.datasets.protein_dataset import ProteinDataset
from torch.utils.data.sampler import WeightedRandomSampler
def main_training(dir_data, dir_mdata, dir_results, out_dir, gpu_id='0',
arch='class_densenet121_dropout', pretrained=None, model_multicell=None,
num_classes=19, in_channels=4, loss='FocalSymmetricLovaszHardLogLoss',
scheduler='Adam45', epochs=55, img_size=768, crop_size=512, batch_size=32,
workers=3, pin_memory=True, split_name='random_ext_folds5', fold=0,
clipnorm=1, resume=None):
'''
PyTorch Protein Classification. Main training function.
Args:
dir_data (str, Path): Directory where training subsets are.
dir_mdata (str, Path): Directory where training meta data is.
dir_results (std, Path): Directory to save training results.
out_dir (str):
Name/label for this training run. Will be used to create
directory under `dir_results`.
gpu_id (str): GPU id used for training. Default: '0'
arch (str): Model architecture.
Default: ``'class_densenet121_dropout'``
pretrained (Path, str): Path to a pretrained model. These are
the parameters just before training starts.
model_multicell (Path, str): Path to multi-cell model
to start training from. Default: None
num_classes (int): Number of classes. Default: 19
in_channels (int): In channels. Default: 4
loss (str, optional): Loss function.
One of ``'FocalSymmetricLovaszHardLogLoss'``.
Default: ``'FocalSymmetricLovaszHardLogLoss'``
scheduler (str): Scheduler name. Default: ``'Adam45'``
epochs (int): Number of total epochs to run. Default: 55
img_size (int): Image size. Default: 768
crop_size (int): Crop size. Default: 512
batch_size (int): Train mini-batch size. Default: 32
workers (int): Number of data loading workers. Default: 3
pin_memory (bool): DataLoader's ``pin_memory`` argument.
split_name (str, optional): Split name.
One of: ``'random_ext_folds5'``,
or ``'random_ext_noleak_clean_folds5'``.
Default: ``'random_ext_folds5'``
fold (int): Index of fold. Default: 0
clipnorm (int): Clip grad norm'. Default: 1
resume (str): Name of the latest checkpoint. Default: None
'''
log_out_dir = opj(dir_results, 'logs', out_dir, 'fold%d' % fold)
if not ope(log_out_dir):
os.makedirs(log_out_dir)
log = Logger()
log.open(opj(log_out_dir, 'log.train.txt'), mode='a')
model_out_dir = opj(dir_results, 'models', out_dir, 'fold%d' % fold)
log.write(">> Creating directory if it does not exist:\n>> '{}'\n".format(model_out_dir))
if not ope(model_out_dir):
os.makedirs(model_out_dir)
# set cuda visible device
os.environ['CUDA_VISIBLE_DEVICES'] = gpu_id
cudnn.benchmark = True
# set random seeds
torch.manual_seed(0)
torch.cuda.manual_seed_all(0)
np.random.seed(0)
model_params = {}
model_params['architecture'] = arch
model_params['num_classes'] = num_classes
model_params['in_channels'] = in_channels
model = init_network(model_params, model_multicell=model_multicell)
if pretrained:
print(f'Loading pretrained model {pretrained}')
checkpoint = torch.load(pretrained)
model.load_state_dict(checkpoint['state_dict'])
# move network to gpu
model = DataParallel(model)
model.to(DEVICE)
# define loss function (criterion)
try:
criterion = eval(loss)().to(DEVICE)
except:
raise(RuntimeError("Loss {} not available!".format(loss)))
start_epoch = 0
best_loss = 1e5
best_epoch = 0
best_focal = 1e5
# define scheduler
try:
scheduler = eval(scheduler)()
except:
raise (RuntimeError("Scheduler {} not available!".format(scheduler)))
optimizer = scheduler.schedule(model, start_epoch, epochs)[0]
# optionally resume from a checkpoint
if resume:
resume = os.path.join(model_out_dir, resume)
if os.path.isfile(resume):
# load checkpoint weights and update model and optimizer
log.write(">> Loading checkpoint:\n>> '{}'\n".format(resume))
checkpoint = torch.load(resume)
start_epoch = checkpoint['epoch']
best_epoch = checkpoint['best_epoch']
best_focal = checkpoint['best_score']
model.module.load_state_dict(checkpoint['state_dict'])
optimizer_fpath = resume.replace('.pth', '_optim.pth')
if ope(optimizer_fpath):
log.write(">> Loading checkpoint:\n>> '{}'\n".format(optimizer_fpath))
optimizer.load_state_dict(torch.load(optimizer_fpath)['optimizer'])
log.write(">>>> loaded checkpoint:\n>>>> '{}' (epoch {})\n".format(resume, checkpoint['epoch']))
else:
log.write(">> No checkpoint found at '{}'\n".format(resume))
# Data loading code
train_transform = train_multi_augment2
train_split_file = dir_mdata / 'split'/ split_name / f'random_train_cv{fold}.feather'
train_dataset = ProteinDataset(
dir_data,
train_split_file,
img_size=img_size,
is_trainset=True,
return_label=True,
in_channels=in_channels,
transform=train_transform,
crop_size=crop_size,
random_crop=True)
label_weights = get_label_weights(train_dataset.split_df)
weights = torch.from_numpy(train_dataset.split_df['Target']
.apply(lambda o: label_weights[o]).values)
sampler = WeightedRandomSampler(weights, len(weights))
train_loader = DataLoader(
train_dataset,
sampler=sampler,
batch_size=batch_size,
drop_last=True,
num_workers=workers,
pin_memory=pin_memory)
valid_split_file = (dir_mdata/'split'/split_name/
f'random_valid_cv{fold}.feather')
valid_dataset = ProteinDataset(
dir_data,
valid_split_file,
img_size=img_size,
is_trainset=True,
return_label=True,
in_channels=in_channels,
transform=None,
crop_size=crop_size,
random_crop=False)
valid_loader = DataLoader(
valid_dataset,
sampler=SequentialSampler(valid_dataset),
batch_size=batch_size,
drop_last=False,
num_workers=workers,
pin_memory=pin_memory)
focal_loss = FocalLoss().to(DEVICE)
log.write('** start training here! **\n')
log.write('\n')
log.write('epoch iter rate | train_loss/acc | valid_loss/acc/focal/kaggle |best_epoch/best_focal| min \n')
log.write('-----------------------------------------------------------------------------------------------------------------\n')
start_epoch += 1
for epoch in range(start_epoch, epochs + 1):
end = time.time()
# set manual seeds per epoch
np.random.seed(epoch)
torch.manual_seed(epoch)
torch.cuda.manual_seed_all(epoch)
# adjust learning rate for each epoch
lr_list = scheduler.step(model, epoch, epochs)
lr = lr_list[0]
# train for one epoch on train set
iter, train_loss, train_acc = train(train_loader, model, criterion, optimizer, epoch, clipnorm=clipnorm, lr=lr)
with torch.no_grad():
valid_loss, valid_acc, valid_focal_loss, kaggle_score = validate(valid_loader, model, criterion, epoch, focal_loss)
# remember best loss and save checkpoint
is_best = valid_focal_loss < best_focal
best_loss = min(valid_focal_loss, best_loss)
best_epoch = epoch if is_best else best_epoch
best_focal = valid_focal_loss if is_best else best_focal
print('\r', end='', flush=True)
log.write('%5.1f %5d %0.6f | %0.4f %0.4f | %0.4f %6.4f %6.4f %6.4f | %6.1f %6.4f | %3.1f min \n' % \
(epoch, iter + 1, lr, train_loss, train_acc, valid_loss, valid_acc, valid_focal_loss, kaggle_score,
best_epoch, best_focal, (time.time() - end) / 60))
save_model(model, is_best, model_out_dir, optimizer=optimizer, epoch=epoch, best_epoch=best_epoch, best_focal=best_focal)
# + papermill={"duration": 0.04128, "end_time": "2021-04-19T18:09:35.656028", "exception": false, "start_time": "2021-04-19T18:09:35.614748", "status": "completed"} tags=[]
model_multicell = (
'../../kgl_humanprotein_data/result/models/'
'external_crop512_focal_slov_hardlog_class_densenet121_dropout_i768_aug2_5folds/'
'fold0/final.pth')
pretrained = Path(
'/kaggle/input/humanpro-classifier-crop/results/models/'
'external_crop256_focal_slov_hardlog_class_densenet121_dropout_i384_aug2_5folds/'
'fold0/final.pth')
gpu_id = '0' # '0,1,2,3'
arch = 'class_densenet121_dropout'
num_classes = len(LABEL_NAME_LIST)
scheduler = 'Adam55'
epochs = 10 #55
resume = None
sz_img = 384
crop_size = 256 #512
batch_size = 76
split_name = 'random_folds5'
fold = 0
workers = 3
pin_memory = True
dir_results = Path('results')
dir_results.mkdir(exist_ok=True, parents=True)
out_dir = Path(f'external_crop{crop_size}_focal_slov_hardlog_class_densenet121_dropout_i384_aug2_5folds')
# + papermill={"duration": 10141.818533, "end_time": "2021-04-19T20:58:37.505326", "exception": false, "start_time": "2021-04-19T18:09:35.686793", "status": "completed"} tags=[]
main_training(dir_data, dir_mdata, dir_results, out_dir,
split_name=split_name, fold=fold,
arch=arch, pretrained=pretrained, model_multicell=model_multicell, scheduler=scheduler,
epochs=epochs, resume=resume,
img_size=sz_img, crop_size=crop_size, batch_size=batch_size,
gpu_id=gpu_id, workers=workers, pin_memory=pin_memory)
# + papermill={"duration": 6.376078, "end_time": "2021-04-19T20:58:46.173237", "exception": false, "start_time": "2021-04-19T20:58:39.797159", "status": "completed"} tags=[]
# ! cp -r results/ /kaggle/working/.
# + papermill={"duration": 8.530096, "end_time": "2021-04-19T20:58:57.521771", "exception": false, "start_time": "2021-04-19T20:58:48.991675", "status": "completed"} tags=[]
| kaggle_notebooks/humanpro-classifier-training-multilabel.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
from sklearn import datasets
iris = datasets.load_iris()
X = iris.data
y = iris.target
from sklearn.model_selection import train_test_split
from sklearn.grid_search import GridSearchCV
from sklearn.cross_validation import cross_val_score , StratifiedKFold
from sklearn import metrics
from sklearn.model_selection import learning_curve
from sklearn.svm import SVC
from sklearn.ensemble import RandomForestClassifier
from sklearn.neighbors import KNeighborsClassifier
import numpy as np
from sklearn.linear_model import LogisticRegression
from sklearn.naive_bayes import GaussianNB
from sklearn.cross_validation import KFold
import xgboost as XGB
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.3, random_state =42)
# +
classifiers = []
logreg_tuned_parameters = [{'C': np.logspace(-1, 2, 4),'penalty':['l1','l2']}]
classifiers.append(["Logistic Regression", LogisticRegression(random_state = 0), logreg_tuned_parameters])
svm_tuned_parameters = [{'kernel': ['linear','rbf'],
'C': np.logspace(-1, 2, 4),
'gamma': np.logspace(-4, 0, 5)
}]
classifiers.append(["SVM", SVC(random_state = 0), svm_tuned_parameters])
rf_tuned_parameters = [{"criterion": ["gini"]}]
classifiers.append(["RandomForest", RandomForestClassifier(random_state = 0, n_jobs=-1), rf_tuned_parameters])
knn_tuned_parameters = [{"n_neighbors": [1, 3, 5, 10, 20]}]
classifiers.append(["kNN", KNeighborsClassifier(),knn_tuned_parameters])
classifiers.append(["gnb", GaussianNB(),{}])
classifiers.append(['xgb', XGB.XGBClassifier(objective='binary:logistic',), {}])
# -
def gsCV_accuracy(name,classifier, params, train, target):
print (name+":")
gs= GridSearchCV(classifier, params, n_jobs=-1, cv=5,scoring="accuracy")
gs.fit(train, target)
#print (gs.best_params_, gs.best_score_)
predict = gs.best_estimator_.predict(train)
print(metrics.classification_report(target,predict))
print(metrics.confusion_matrix(target, predict))
print(cross_val_score(gs.best_estimator_, train,target,cv= 5).mean())
for i in range(len(classifiers)):
gsCV_accuracy(classifiers[i][0],classifiers[i][1], classifiers[i][2], X_train, y_train)
# +
class Stacking(object):
def __init__(self, seed, n_fold, base_learners, meta_learner):
self.seed = seed
self.n_fold = n_fold
self.base_learners = base_learners
self.meta_learner = meta_learner
self.T = len(base_learners) # num of base learners
def generateBaseLearner(self, X_tr, y_tr, X_te, y_te):
n1 = X_tr.shape[0]
n2 = X_te.shape[0]
kf = KFold(n1, n_folds= self.n_fold, random_state= self.seed)
#constructing data for meta learner
meta_train = np.zeros((n1, self.T))
meta_test = np.zeros((n2, self.T))
for i, clf in enumerate(self.base_learners):
meta_test_i = np.zeros((n2, self.n_fold))
for j, (train_index, test_index) in enumerate(kf):
X_train = X_tr[train_index]
y_train = y_tr[train_index]
X_holdout = X_tr[test_index]
y_holdout = y_tr[test_index]
clf[1].fit(X_train, y_train)
y_pred = clf[1].predict(X_holdout)[:]
print 'Base Learner:%s accuracy = %s' % (clf[0], metrics.accuracy_score(y_holdout, y_pred))
# filling predicted X_holdout into meta training set
meta_train[test_index, i] = y_pred
meta_test_i[:, j] = clf[1].predict(X_te)[:]
meta_test[:, i] = meta_test_i.mean(1)
self.meta_learner.fit(meta_train, y_tr)
y_result_pred = self.meta_learner.predict(meta_test)
print metrics.classification_report(y_te, y_result_pred)
print(metrics.confusion_matrix(y_te, y_result_pred))
print 'Final accuracy = %s' % (metrics.accuracy_score(y_te, y_result_pred))
return y_result_pred
# +
#baseLearner Default
lg = LogisticRegression(random_state= 0)
svm = SVC(random_state= 0)
rf = RandomForestClassifier( random_state= 0, n_jobs=-1)
knn = KNeighborsClassifier()
gnb = GaussianNB()
xgb = XGB.XGBClassifier()
lg2 = LogisticRegression(penalty = 'l1', C = 10 ,random_state= 0)
svm2 = SVC(kernel= 'rbf', C= 100.0, gamma= 0.01,random_state= 0)
rf2 = RandomForestClassifier( criterion = 'gini',random_state= 0, n_jobs=-1)
knn2 = KNeighborsClassifier(n_neighbors = 1)
base_learner2 = [['SVM', svm2], ['Random Forest', rf2], ['KNN',knn2]]
# -
base_learner = [['SVM', svm], ['Random Forest', rf], ['KNN',knn]]
stackingD = Stacking(0, 3, base_learner, lg)
stackingD.generateBaseLearner(X_train, y_train, X_test, y_test)
# +
stacking2 = Stacking(0, 5, base_learner2, lg2)
stacking2.generateBaseLearner(X_train, y_train, X_test, y_test)
# +
''' X_tr = np.array(X_tr)
y_tr = np.array(y_tr)
X_te = np.array(X_te)
y_te = np.array(y_te)'''
base_learner3 = [['SVM', svm], ['Logistic regression', lg], ['KNN',knn]]
stacking3 = Stacking(0, 3, base_learner3, rf)
stacking3.generateBaseLearner(X_train, y_train, X_test, y_test)
# +
base_learner4 = [['Random Forest', rf], ['Logistic regression', lg], ['KNN',knn]]
stacking4 = Stacking(0, 3, base_learner4, svm)
stacking4.generateBaseLearner(X_train, y_train, X_test, y_test)
# +
base_learner5 = [['Random Forest', rf], ['Logistic regression', lg], ['SVM', svm]]
stacking5 = Stacking(0, 3, base_learner5, knn)
stacking5.generateBaseLearner(X_train, y_train, X_test, y_test)
# +
base_learner6 = [['Random Forest', rf], ['Logistic regression', lg], ['SVM', svm], ['knn', knn]]
stacking6 = Stacking(0, 3, base_learner6, gnb)
stacking6.generateBaseLearner(X_train, y_train, X_test, y_test)
# +
lg = LogisticRegression(random_state= 13)
rf = RandomForestClassifier(random_state= 13)
base_learner3 = [ ['lg', lg], ['knn', knn], ['svm', svm]]
stacking3 = Stacking(0, 3, base_learner3, gnb)
stacking3.generateBaseLearner(X_train, y_train, X_test, y_test)
# +
lg = LogisticRegression(random_state= 0)
rf = RandomForestClassifier(random_state= 0)
base_learner3 = [ ['lg', lg], ['knn', knn], ['svm', svm]]
stacking3 = Stacking(0, 3, base_learner3, gnb)
stacking3.generateBaseLearner(X_train, y_train, X_test, y_test)
# -
| Stacking.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
# +
import random
import json
import torch
from model import NeuralNet
from nltk_utils import bag_of_words, tokenize
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
with open('intents.json', 'r') as json_data:
intents = json.load(json_data)
FILE = "data.pth"
data = torch.load(FILE)
input_size = data["input_size"]
hidden_size = data["hidden_size"]
output_size = data["output_size"]
all_words = data['all_words']
tags = data['tags']
model_state = data["model_state"]
model = NeuralNet(input_size, hidden_size, output_size).to(device)
model.load_state_dict(model_state)
model.eval()
bot_name = "Sam"
print("Let's chat! (type 'quit' to exit)")
while True:
# sentence = "do you use credit cards?"
sentence = input("You: ")
if sentence == "quit":
break
sentence = tokenize(sentence)
X = bag_of_words(sentence, all_words)
X = X.reshape(1, X.shape[0])
X = torch.from_numpy(X).to(device)
output = model(X)
_, predicted = torch.max(output, dim=1)
tag = tags[predicted.item()]
probs = torch.softmax(output, dim=1)
prob = probs[0][predicted.item()]
if prob.item() > 0.75:
for intent in intents['intents']:
if tag == intent["tag"]:
print(f"{bot_name}: {random.choice(intent['responses'])}")
else:
print(f"{bot_name}: I do not understand...")
# -
import nltk
nltk.download('punkt')
| chat.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Stack Overflow Data - 2017 Survey
# ## Business Understanding
#
# Stack OverFlow is a website used by the global development community. Through an online survey, it was possible to obtain a large base of information about the wishes of the developer community. Through this information I will be looking to understand how the feeling of the community is in relation to the programming language, salary and work.
# ## Question
#
# Question 1. Happiness
#
# Question 2. Happiness and salary
#
# Question 3. Professional and happiness
#
# Question 4. Language and happiness
# # Data Understanding
# ## Imports necessary for first analisys
# +
import seaborn as sns
sns.set_theme(style="darkgrid")
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
pd.options.display.max_columns = None
pd.options.display.max_rows = None
# %config IPCompleter.greedy=True
# -
# ## Gather
#
# #### Access and Explore
df = pd.read_csv('survey_results_public.csv')
df.head()
# ## Analysis
# +
# Lets see how is our data
# -
df.shape
df.isna().sum()
# ### So many null data
df.describe()
# ### Only five numerical data
df.dtypes
# #### Many object data, with many missing values.
# # Analyse for Question
#
# For this analysis we are looking for Jobsatisfaction. For our case, is necessary to use the colum JobSatisfaction.
df.JobSatisfaction.unique()
# ## The range is so long, let made cut for have only tree categorical
# ### We are use only tree categorical values
# sad -> <=4
#
# neutral -> >4 && <= 7
#
# happy -> >7
df['JobSatisfactionCategorical'] = np.where(df.JobSatisfaction > 7, 'happy',
(np.where(df.JobSatisfaction > 4, 'neutral', 'sad')))
df[['JobSatisfactionCategorical','JobSatisfaction']].head(10)
# +
## NAN values turned sad, is necessary remove NaN for not up the sad values
# -
df = df.dropna(subset=['JobSatisfaction'], axis=0)
df[['JobSatisfactionCategorical','JobSatisfaction']].head(10)
# ## Question 1. Happiness
# +
## Analisys
# -
df.JobSatisfactionCategorical.value_counts()
# +
## Visualize
# -
sns.countplot(x="JobSatisfactionCategorical",
data=df,
);
# +
# Brief explanation for visualisation
# -
# ### Many happy employers, huhuuuuu
# ## Question 2. Happiness and salary
# ### In describe the colum Salary have many values, and diference values. For our case and made one more easy analyse we are use mean for Salary
# Lets made a division, salary mean
#
# salary_mean > acima
#
# salary_mean <= abaixo
# +
# Analyse
# -
df.Salary.mean()
df['SalaryCategorical'] = np.where(df.Salary > df.Salary.mean(), 1, 0)
df.SalaryCategorical.value_counts()
# +
# Visualize
# -
g = sns.catplot(x="JobSatisfactionCategorical", col="SalaryCategorical",
data=df, kind="count",
height=4, aspect=.7);
# +
# Brief explanation for visualisation
# -
# ## The salary does not interfere so much in the happiness of the employees
# # Question 3. Professional and happiness
# +
# Analyse
# -
g = sns.catplot(x="JobSatisfactionCategorical", col="Professional",
data=df, kind="count",
height=4, aspect=.9, col_wrap=3);
# +
# Brief explanation for visualisation
# -
# ### The professional is happy too
# # Question 4. Language and happiness
# +
# Analyse
# -
cols = ['HaveWorkedLanguage', 'WantWorkLanguage', 'JobSatisfactionCategorical', 'JobSatisfaction']
df2 = df[cols]
df2.shape
# ### for this analyse, we use count only. For this, no have a problem remove nan values
df2 = df2.dropna()
df2.shape
df2.head()
def worked_language_happiness(df, top, col_list_language, col_job_satisfaction):
"""
INPUT
df -> dataframe for analysis with columns [Workedlanguage, Jobsatisfaction]
top -> int for numbers of data show in end
col_list_language -> col from df, with string sep ; language
col_job_satisfaction -> col with float of jobstisfaction
OUTPUT
dataframe top languages
"""
dict_professional = {}
for index, row in df2.iterrows():
temp_value = row[col_list_language].split(";")
list_temp = [x.strip(' ') for x in temp_value]
for value in list_temp:
if value in dict_professional.keys():
mean = dict_professional[value]
mean = (mean + row[col_job_satisfaction]) / 2
else:
dict_professional[value] = row[col_job_satisfaction]
data = {'Language': dict_professional.keys(), 'Mean': dict_professional.values()}
df_3 = pd.DataFrame.from_dict(data)
df_3['Mean'] = df_3['Mean'].astype(int)
df_3['meanCategorical'] = np.where(df_3['Mean'] > 7, 'happy',
(np.where(df_3['Mean'] > 4, 'neutral', 'sad')))
return df_3
# +
# Visualise
# analise for HaveWorkedLanguage
# -
df_3 = worked_language_happiness(df2, 10, 'HaveWorkedLanguage', 'JobSatisfaction')
df_3.sort_values('Mean', ascending = False).head(10)
# ## Seriously? Julia, what a thing. But the calculation takes into account only the average.
# +
# Visualise
# analise for HaveWorkedLanguage
# -
df_3 = worked_language_happiness(df2, 10, 'WantWorkLanguage', 'JobSatisfaction')
df_3.sort_values('Mean', ascending = False).head(10)
# ## C, Python is in everywhere
# ## Conclusion
# ### The number of people satisfied with working with development is very large, it is not possible to see any very expressive bar in the graphics. This is good, it would be better if happiness was always greater than sad.
# ### About the programming language, the values 10 were curious, but the values 9 are well spread out.
#
# ### Check my blog post for this analysis: https://andrezio.medium.com/happiness-and-programming-language-fe52c0bcdd03
# # Acknowledgments
#
# For all Stackoverflow developers who have contributed to this incredible database.
| data_science/Stack_Overflow_Developer_Survey/Stack Overflow Data - 2017 Survey - Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # HW1 : Regression, Cross-Validation, and Regularization
# +
import os
import numpy as np
import warnings
import sklearn.preprocessing
import sklearn.pipeline
import sklearn.linear_model
import sklearn.neighbors
import sklearn.model_selection
# +
from matplotlib import pyplot as plt
# %matplotlib inline
import seaborn as sns
sns.set('notebook', font_scale=1.25, style='whitegrid')
# -
# # Set random seed to make all computations reproducible
SEED = 12345
# # Load the dataset
#
# Predefined 'x' and 'y' arrays for train/valid/test
DATA_DIR = '../data_auto/'
x_tr_MF = np.loadtxt(os.path.join(DATA_DIR, 'x_train.csv'), delimiter=',', skiprows=1)
x_va_NF = np.loadtxt(os.path.join(DATA_DIR, 'x_valid.csv'), delimiter=',', skiprows=1)
x_te_PF = np.loadtxt(os.path.join(DATA_DIR, 'x_test.csv'), delimiter=',', skiprows=1)
y_tr_M = np.loadtxt(os.path.join(DATA_DIR, 'y_train.csv'), delimiter=',', skiprows=1)
y_va_N = np.loadtxt(os.path.join(DATA_DIR, 'y_valid.csv'), delimiter=',', skiprows=1)
y_te_P = np.loadtxt(os.path.join(DATA_DIR, 'y_test.csv'), delimiter=',', skiprows=1)
# # Load completed code
from cross_validation import train_models_and_calc_scores_for_n_fold_cv
from performance_metrics import calc_mean_squared_error
# # Define useful plotting functions
def plot_train_and_valid_error_vs_degree(
degree_list, err_tr_list=None, err_va_list=None):
''' Plot provided errors versus degrees on a new figure
'''
if err_va_list is not None:
plt.plot(degree_list, err_va_list, 'rs-', label='valid');
if err_tr_list is not None:
plt.plot(degree_list, err_tr_list, 'bd:', label='train');
plt.ylim([0, 74]); # Do NOT change this! Helps all reports look the same.
plt.legend(loc='upper right'); # Always include a legend
# # Define methods for building pipelines
#
# Remember, we discussed pipelines in the lab from day04 on "Feature Engineering"
def make_poly_linear_regr_pipeline(degree=1):
pipeline = sklearn.pipeline.Pipeline(
steps=[
('rescaler', sklearn.preprocessing.MinMaxScaler()),
('poly_transformer', sklearn.preprocessing.PolynomialFeatures(degree=degree, include_bias=False)),
('linear_regr', sklearn.linear_model.LinearRegression()),
])
# Return the constructed pipeline
# We can treat it as if it has a 'regression' API
# e.g. a fit and a predict method
return pipeline
def make_poly_ridge_regr_pipeline(degree=1, alpha=1.0):
pipeline = sklearn.pipeline.Pipeline(
steps=[
('rescaler', sklearn.preprocessing.MinMaxScaler()),
('poly_transformer', sklearn.preprocessing.PolynomialFeatures(degree=degree, include_bias=False)),
('ridge_regr', sklearn.linear_model.Ridge(alpha=alpha)),
])
# Return the constructed pipeline
# We can treat it as if it has a 'regression' API
# e.g. a fit and a predict method
return pipeline
# # Problem 1: Polynomial Degree Selection on Fixed Validation Set
# +
degree_list = [1, 2, 3, 4, 5, 6, 7]
fv_err_tr_list = []
fv_err_va_list = []
pipeline_list = []
for degree in degree_list:
# TODO create a pipeline using features with current degree value
# TODO train this pipeline on provided training data
# Compute training error
yhat_tr_M = 0.0 # TODO fixme
err_tr = 23.0 # TODO fixme
# Compute validation error
yhat_va_N = 0.0 # TODO fixme
err_va = 25.0 # TODO fixme
fv_err_tr_list.append(err_tr)
fv_err_va_list.append(err_va)
# -
# ### Figure 1: Error vs degree
plot_train_and_valid_error_vs_degree(degree_list, fv_err_tr_list, fv_err_va_list);
plt.title('Model Selection with Fixed Validation Set: Error vs. Degree');
plt.savefig('figure1-err_vs_degree-fv.pdf')
# ### Prediction 1: Score on the test set using the chosen model
# +
# TODO compute score on test set for later
# -
print("Selected Parameters:")
print("TODO")
print("Fixed validation set estimate of heldout error:")
print("TODO")
print("Error on the test-set:")
print("TODO")
# # Problem 2: Cross Validation for Polynomial Feature Regression
x_trva_LF = x_tr_MF.copy() # TODO fix concat your train and validation set x values
y_trva_L = y_tr_M.copy() # TODO fix concat your train and validation set y values
# +
K = 10 # num folds
degree_list = [1, 2, 3, 4, 5, 6, 7]
cv_err_tr_list = []
cv_err_va_list = []
for degree in degree_list:
# TODO create a pipeline using features with current degree value
# TODO call your function to train a separate model for each fold and return train and valid errors
# Don't forget to pass random_state = SEED (where SEED is defined above) so its reproducible
# tr_error_K, valid_error_K = train_models_and_calc_scores_for_n_fold_cv() # TODO
err_tr = 20.0 # TODO fixme, compute average error across all train folds
err_va = 25.0 # TODO fixme, compute average error across all heldout folds
cv_err_tr_list.append(err_tr)
cv_err_va_list.append(err_va)
# -
# ### Figure 2: Error vs degree
plot_train_and_valid_error_vs_degree(degree_list, cv_err_tr_list, cv_err_va_list)
plt.title('Linear Regr. Model Selection with 10-fold Cross-Validation: Error vs. Degree');
plt.savefig('figure2-err_vs_degree-cv-seed=%d.pdf' % SEED)
# ### Prediction 2: Score on the test set using the chosen model
#
# Use the chosen hyperparameters, retrain ONE model on the FULL train+valid set.
# Then make predictions on the heldout test set.
# +
# TODO compute score on test set for later
# -
print("Selected Parameters:")
print("TODO")
print("10-fold CV estimate of heldout error:")
print("TODO")
print("Error on the test-set:")
print("TODO")
# # Problem 3: Cross Validation for Ridge Regression
# +
alpha_grid = np.logspace(-6, 6, 13) # 10^-6, 10^-5, 10^-4, ... 10^-1, 10^0, 10^1, ... 10^6
degree_list = [1, 2, 3, 4, 5, 6, 7]
K = 10 # num folds
ridge_cv_err_tr_list = []
ridge_cv_err_va_list = []
ridge_param_list = list()
for degree in degree_list:
for alpha in alpha_grid:
ridge_param_list.append(dict(alpha=alpha, degree=degree))
# TODO create a pipeline using features with current degree value
# TODO call your function to train a separate model for each fold and return train and valid errors
# Don't forget to pass random_state = SEED (where SEED is defined above) so its reproducible
# tr_error_K, valid_error_K = train_models_and_calc_scores_for_n_fold_cv() # TODO
err_tr = 20.0 # TODO fixme, compute average error across all train folds
err_va = 25.0 # TODO fixme, compute average error across all heldout folds
ridge_cv_err_tr_list.append(err_tr)
ridge_cv_err_va_list.append(err_va)
# -
# ### Figure 3: Error vs degree at alpha = 10^-3, alpha = 1, alpha = 1000
# +
fig, ax_grid = plt.subplots(nrows=1, ncols=3, sharex=True, sharey=True, figsize=(16, 5.5))
for aa, alpha in enumerate([0.00001, 0.1, 1000.0]):
# Find the elements of the param list that correspond to setting alpha to specific value
match_ids = [pp for pp in range(len(ridge_param_list)) if np.allclose(alpha, ridge_param_list[pp]['alpha'])]
train_err = np.asarray(ridge_cv_err_tr_list)[match_ids]
test_err = np.asarray(ridge_cv_err_va_list)[match_ids]
# Select which panel (of the 3 in figure) to be current active axis
cur_ax = ax_grid[aa]
plt.sca(cur_ax);
# Set the title of the active axis
cur_ax.set_title('alpha = %.5g' % alpha)
# Draw line plot in active axis
plot_train_and_valid_error_vs_degree(degree_list, train_err, test_err)
plt.suptitle('Ridge Model Selection with 10-fold Cross-Validation: Error vs. Degree');
plt.savefig('figure3-3_panels_by_alpha-err_vs_degree-seed=%d.pdf' % SEED, pad_inches=0, bbox_inches='tight')
# -
# ### Prediction 3: Score on the test set using the chosen model
#
print("Selected Parameters (alpha and degree):")
print("TODO")
print("10-fold CV estimate of heldout error:")
print("TODO")
print("Error on the test-set:")
print("TODO")
| hw1/hw1_notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # A Python Tour of Data Science: Data Visualization
#
# [<NAME>](http://deff.ch), *PhD student*, [EPFL](http://epfl.ch) [LTS2](http://lts2.epfl.ch)
# # Exercise
#
# Data visualization is a key aspect of exploratory data analysis.
# During this exercise we'll gradually build more and more complex vizualisations. We'll do this by replicating plots. Try to reproduce the lines but also the axis labels, legends or titles.
#
# * Goal of data visualization: clearly and efficiently communicate information through visual representations. While tables are generally used to look up a specific measurement, charts are used to show patterns or relationships.
# * Means: mainly statistical graphics for exploratory analysis, e.g. scatter plots, histograms, probability plots, box plots, residual plots, but also [infographics](https://en.wikipedia.org/wiki/Infographic) for communication.
#
# *Data visualization is both an art and a science. It should combine both aesthetic form and functionality.*
# # 1 Time series
#
# To start slowly, let's make a static line plot from some time series. Reproduce the plots below using:
# 1. The procedural API of [matplotlib](http://matplotlib.org), the main data visualization library for Python. Its procedural API is similar to matlab and convenient for interactive work.
# 2. [Pandas](http://pandas.pydata.org), which wraps matplotlib around his DataFrame format and makes many standard plots easy to code. It offers many [helpers for data visualization](http://pandas.pydata.org/pandas-docs/version/0.19.1/visualization.html).
#
# **Hint**: to plot with pandas, you first need to create a DataFrame, pandas' tabular data format.
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
# Random time series.
n = 1000
rs = np.random.RandomState(42)
data = rs.randn(n, 4).cumsum(axis=0)
# -
plt.figure(figsize=(15,5))
plt.plot(data[:, 0], label='A')
plt.plot(data[:, 1], '.-k', label='B')
plt.plot(data[:, 2], '--m', label='C')
plt.plot(data[:, 3], ':', label='D')
plt.legend(loc='upper left')
plt.xticks(range(0, 1000, 50))
plt.ylabel('Value')
plt.xlabel('Day')
plt.grid()
idx = pd.date_range('1/1/2000', periods=n)
df = pd.DataFrame(data, index=idx, columns=list('ABCD'))
df.plot(figsize=(15,5));
# # 2 Categories
#
# Categorical data is best represented by [bar](https://en.wikipedia.org/wiki/Bar_chart) or [pie](https://en.wikipedia.org/wiki/Pie_chart) charts. Reproduce the plots below using the object-oriented API of matplotlib, which is recommended for programming.
#
# **Question**: What are the pros / cons of each plot ?
#
# **Tip**: the [matplotlib gallery](http://matplotlib.org/gallery.html) is a convenient starting point.
data = [10, 40, 25, 15, 10]
categories = list('ABCDE')
# +
fig, axes = plt.subplots(1, 2, figsize=(15, 5))
axes[1].pie(data, explode=[0,.1,0,0,0], labels=categories, autopct='%1.1f%%', startangle=90)
axes[1].axis('equal')
pos = range(len(data))
axes[0].bar(pos, data, align='center')
axes[0].set_xticks(pos)
axes[0].set_xticklabels(categories)
axes[0].set_xlabel('Category')
axes[0].set_title('Allotment');
# -
# # 3 Frequency
#
# A frequency plot is a graph that shows the pattern in a set of data by plotting how often particular values of a measure occur. They often take the form of an [histogram](https://en.wikipedia.org/wiki/Histogram) or a [box plot](https://en.wikipedia.org/wiki/Box_plot).
#
# Reproduce the plots with the following three libraries, which provide high-level declarative syntax for statistical visualization as well as a convenient interface to pandas:
# * [Seaborn](http://seaborn.pydata.org) is a statistical visualization library based on matplotlib. It provides a high-level interface for drawing attractive statistical graphics. Its advantage is that you can modify the produced plots with matplotlib, so you loose nothing.
# * [ggplot](http://ggplot.yhathq.com) is a (partial) port of the popular [ggplot2](http://ggplot2.org) for R. It has his roots in the influencial book [the grammar of graphics](https://www.cs.uic.edu/~wilkinson/TheGrammarOfGraphics/GOG.html). Convenient if you know ggplot2 already.
# * [Vega](https://vega.github.io/) is a declarative format for statistical visualization based on [D3.js](https://d3js.org), a low-level javascript library for interactive visualization. [Vincent](https://vincent.readthedocs.io/en/latest/) (discontinued) and [altair](https://altair-viz.github.io/) are Python libraries to vega. Altair is quite new and does not provide all the needed functionality yet, but it is promising !
#
# **Hints**:
# * Seaborn, look at `distplot()` and `boxplot()`.
# * ggplot, we are interested by the [geom_histogram](http://ggplot.yhathq.com/docs/geom_histogram.html) geometry.
import seaborn as sns
import os
df = sns.load_dataset('iris', data_home=os.path.join('..', 'data'))
# +
fig, axes = plt.subplots(1, 2, figsize=(15, 5))
g = sns.distplot(df['petal_width'], kde=True, rug=False, ax=axes[0])
g.set(title='Distribution of petal width')
g = sns.boxplot('species', 'petal_width', data=df, ax=axes[1])
g.set(title='Distribution of petal width by species');
# +
import ggplot
ggplot.ggplot(df, ggplot.aes(x='petal_width', fill='species')) + \
ggplot.geom_histogram() + \
ggplot.ggtitle('Distribution of Petal Width by Species')
# +
import altair
altair.Chart(df).mark_bar(opacity=.75).encode(
x=altair.X('petal_width', bin=altair.Bin(maxbins=30)),
y='count(*)',
color=altair.Color('species')
)
# -
# # 4 Correlation
#
# [Scatter plots](https://en.wikipedia.org/wiki/Scatter_plot) are very much used to assess the correlation between 2 variables. Pair plots are then a useful way of displaying the pairwise relations between variables in a dataset.
#
# Use the seaborn `pairplot()` function to analyze how separable is the iris dataset.
sns.pairplot(df, hue="species");
# # 5 Dimensionality reduction
#
# Humans can only comprehend up to 3 dimensions (in space, then there is e.g. color or size), so [dimensionality reduction](https://en.wikipedia.org/wiki/Dimensionality_reduction) is often needed to explore high dimensional datasets. Analyze how separable is the iris dataset by visualizing it in a 2D scatter plot after reduction from 4 to 2 dimensions with two popular methods:
# 1. The classical [principal componant analysis (PCA)](https://en.wikipedia.org/wiki/Principal_component_analysis).
# 2. [t-distributed stochastic neighbor embedding (t-SNE)](https://en.wikipedia.org/wiki/T-distributed_stochastic_neighbor_embedding).
#
# **Hints**:
# * t-SNE is a stochastic method, so you may want to run it multiple times.
# * The easiest way to create the scatter plot is to add columns to the pandas DataFrame, then use the Seaborn `swarmplot()`.
from sklearn.decomposition import PCA
from sklearn.manifold import TSNE
# +
pca = PCA(n_components=2)
X = pca.fit_transform(df.values[:, :4])
df['pca1'] = X[:, 0]
df['pca2'] = X[:, 1]
tsne = TSNE(n_components=2)
X = tsne.fit_transform(df.values[:, :4])
df['tsne1'] = X[:, 0]
df['tsne2'] = X[:, 1]
# -
fig, axes = plt.subplots(1, 2, figsize=(15, 5))
sns.swarmplot(x='pca1', y='pca2', data=df, hue='species', ax=axes[0])
sns.swarmplot(x='tsne1', y='tsne2', data=df, hue='species', ax=axes[1]);
# # 6 Interactive visualization
#
# For interactive visualization, look at [bokeh](http://bokeh.pydata.org) (we used it during the [data exploration exercise](http://nbviewer.jupyter.org/github/mdeff/ntds_2016/blob/with_outputs/toolkit/01_demo_acquisition_exploration.ipynb#4-Interactive-Visualization)) or [VisPy](http://vispy.org).
# # 7 Geographic map
#
# If you want to visualize data on an interactive map, look at [Folium](https://github.com/python-visualization/folium).
| toolkit/04_sol_visualization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys
sys.path.insert(0, '..')
# %matplotlib inline
import gluonbook as gb
import mxnet as mx
from mxnet import autograd, gluon, image, init, nd
from mxnet.gluon import data as gdata, loss as gloss, utils as gutils
import sys
from time import time
from matplotlib.pyplot import savefig
# -
gb.set_figsize()
img = image.imread('/home/cad488/recognitioned_images_new/110032.jpg')
gb.plt.imshow(img.asnumpy())
# 本函数已保存在 gluonbook 包中方便以后使用。
def show_images(imgs, num_rows, num_cols, scale=2):
figsize = (12, 12)
_, axes = gb.plt.subplots(num_rows, num_cols, figsize=figsize)
for i in range(num_rows):
for j in range(num_cols):
axes[i][j].imshow(imgs[i * num_cols + j].asnumpy())
axes[i][j].axes.get_xaxis().set_visible(False)
axes[i][j].axes.get_yaxis().set_visible(False)
return axes
def apply(img, aug, num_rows=2, num_cols=4, scale=1.5):
Y = [aug(img) for _ in range(num_rows * num_cols)]
show_images(Y, num_rows, num_cols, scale)
savefig("1.jpg")
apply(img, gdata.vision.transforms.RandomFlipLeftRight())
apply(img, gdata.vision.transforms.RandomFlipTopBottom())
shape_aug = gdata.vision.transforms.RandomResizedCrop(
(128, 32), scale=(0.2, 1), ratio=(0.5, 2))
apply(img, shape_aug)
| imageaugmentation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Imitate GaSe
#
# パラメーター
# https://arxiv.org/ftp/arxiv/papers/1707/1707.01288.pdf
# $m_v = -2.16$
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
# %run ./main.py
import matplotlib.pyplot as plt
who
PC = parameter_class()
PC.Nk = 200
PC.mv = -2.16
PC.mc = -PC.mv/4.0 #Rough estimation from https://materialsproject.org/materials/mp-1943/
PC.Eg = 2.0/Hartree
PC.Make_kspace()
ev = k2ev(PC,PC.k)
ec = k2ec(PC,PC.k)
plt.figure()
plt.ylabel('Energy [eV]')
plt.plot(PC.k,ev*Hartree)
plt.plot(PC.k,ec*Hartree)
plt.grid()
plt.show()
#
vv = k2vv(PC,PC.k)
vc = k2vc(PC,PC.k)
plt.figure()
plt.plot(PC.k,vv)
plt.plot(PC.k,vc)
plt.grid()
plt.show()
# +
PC.Nt = 100000
PC.Ncycle = 1000
PC.omegac = 0.62/Hartree
PC.E0 = 1.0/Atomfield
t, A, E= Make_fields(PC)
print('tpi/dt',tpi/(t[1]-t[0]), ' a.u.')
print('Eg',PC.Eg, ' a.u.')
kpAt = get_kpAt(PC,A)
evkt = k2ev(PC,kpAt)
eckt = k2ec(PC,kpAt)
print(evkt.shape)
#
thetavkt = ekt2thetakt(PC, t, evkt)
thetackt = ekt2thetakt(PC, t, eckt)
#
dcckt = get_dcckt(PC, thetavkt, thetackt, evkt, eckt, E)
cckt = dcckt2cckt(PC, t, dcckt)
#
J = get_J(PC, cckt,thetavkt,thetackt)
plt.figure()
plt.xlabel('Time [optical cycle]')
plt.plot(t*PC.omegac/tpi,J)
plt.grid()
plt.show()
#
plt.figure()
plt.xlim(PC.Ncycle/2, PC.Ncycle/2+5.0)
plt.xlabel('Time [optical cycle]')
plt.plot(t*PC.omegac/tpi,J)
plt.fill_between(t*PC.omegac/tpi,E/np.amax(E)*0.8*np.amax(J),0.0*E,color='k',alpha=0.2)
plt.grid()
plt.show()
# +
dt = t[1] - t[0]
omega = np.fft.fftfreq(PC.Nt)*(tpi/dt)
JF = np.fft.fft(J)
plt.figure()
plt.xlim(0.0,10.0)
plt.xlabel('Harmonic order ['+str(np.round(PC.omegac*Hartree, decimals=3))+' eV]')
plt.yscale('log')
plt.plot(omega[:PC.Nt//2]/PC.omegac,np.abs(JF[:PC.Nt//2]))
plt.grid()
plt.show()
a = F2f(t,J)
aF = np.fft.fft(a)
plt.xlim(0.0,10.0)
plt.xlabel('Harmonic order ['+str(np.round(PC.omegac*Hartree, decimals=3))+' eV]')
plt.yscale('log')
plt.plot(omega[:PC.Nt//2]/PC.omegac,np.abs(aF[:PC.Nt//2]))
plt.grid()
plt.show()
# -
def f_Fourier_filter(t, f, ef_center, ef_width):
eps = 1.0e-15
dt = t[1] - t[0]
omega = np.fft.fftfreq(PC.Nt)*(tpi/dt)
fF = np.fft.fft(f)
filt = np.exp(-(np.abs(omega) - ef_center)**2/ef_width**2) + eps
#debub
plt.figure()
#plt.yscale('log')
plt.plot(omega*Hartree,np.abs(fF))
plt.plot(omega*Hartree,np.abs(filt*fF))
plt.show()
f_filter = np.fft.ifft(filt*fF)
#
plt.figure()
plt.plot(t,np.real(f_filter),label='real')
plt.plot(t,np.imag(f_filter),label='imag')
plt.legend()
plt.grid()
plt.show()
return np.real(f_filter)
ef_center = 1.0*PC.omegac
ef_width = 1.0*PC.omegac
J1 = f_Fourier_filter(t, J, ef_center, ef_width)
ef_center = 3.0*PC.omegac
J3 = f_Fourier_filter(t, J, ef_center, ef_width)
ef_center = 5.0*PC.omegac
J5 = f_Fourier_filter(t, J, ef_center, ef_width)
ef_center = 7.0*PC.omegac
J7 = f_Fourier_filter(t, J, ef_center, ef_width)
ef_center = 9.0*PC.omegac
J9 = f_Fourier_filter(t, J, ef_center, ef_width)
P1 = f2F(t,J1)
P1 = P1 - np.average(P1)
P3 = f2F(t,J3)
P3 = P3 - np.average(P3)
P5 = f2F(t,J5)
P5 = P5 - np.average(P5)
P7 = f2F(t,J7)
P7 = P7 - np.average(P7)
P9 = f2F(t,J9)
P9 = P9 - np.average(P9)
plt.figure()
plt.xlabel('Time [optical cycle]')
plt.plot(t*PC.omegac/tpi,P1/np.amax(P1),label='1')
plt.plot(t*PC.omegac/tpi,P3/np.amax(P3),label='3')
plt.plot(t*PC.omegac/tpi,P5/np.amax(P5),label='5')
plt.plot(t*PC.omegac/tpi,P7/np.amax(P7),label='7')
plt.plot(t*PC.omegac/tpi,P9/np.amax(P9),label='9')
plt.grid()
plt.show()
#
plt.figure()
plt.xlim(PC.Ncycle/2, PC.Ncycle/2+2.0)
plt.xlabel('Time [optical cycle]')
plt.plot(t*PC.omegac/tpi,P1/np.amax(P1),label='1')
plt.plot(t*PC.omegac/tpi,P3/np.amax(P3)+1.0,label='3')
plt.plot(t*PC.omegac/tpi,P5/np.amax(P5)+2.0,label='5')
plt.plot(t*PC.omegac/tpi,P7/np.amax(P7)+3.0,label='7')
plt.plot(t*PC.omegac/tpi,P9/np.amax(P9)+4.0,label='9')
plt.fill_between(t*PC.omegac/tpi,E/np.amax(E)*1.2,0.0*E,color='k',alpha=0.2)
plt.grid()
plt.show()
| src/sandbox_1.0Vnm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/powderflask/cap-comp215/blob/main/examples/week1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="WBMC0GrjNRTM"
# # Welcome to Comp 215!
# This is our week 1 examples notebook and will be available on Github from the powderflask/cap-comp215 repository.
#
# As usual, the first code block just imports the modules we will use.
# + id="C1a6aDXAM0WM"
import matplotlib.pyplot as plt
from pprint import pprint
# + [markdown] id="k70NJ-JTNwYN"
# ## Exponential Growth
# How can we use code to better understand exponential growth?
# + colab={"base_uri": "https://localhost:8080/"} id="HZSz8IftqJ72" outputId="7cab3814-86b9-4ca0-b664-495dc4eb7e76"
initial_duck_weed = 2
duck_weed = [initial_duck_weed, ]
for day in range(1, 31):
duck_weed.append( duck_weed[-1] * 2 )
print("There are now {num} duck weed plants.".format(num=duck_weed[-1]))
print(duck_weed)
# + [markdown] id="b6w3F2yErxuB"
# ## Plot the Growth
# we'll use matplotlib not because it is awesome, but because it is the standard tool most scientists use
# + colab={"base_uri": "https://localhost:8080/", "height": 318} id="i_5lOF51r63i" outputId="47fe3e2c-64de-4c17-ca09-825dc3ced101"
pct_cover = [dw/duck_weed[-1]*100 for dw in duck_weed]
fig, ax = plt.subplots()
ax.set_title("Duck Weed Growth over 30 days")
ax.plot(range(0, 31), pct_cover, label="% pond coveraged")
ax.legend()
| examples/week1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#Write a program to identify sublist [1,1,5] is there in given list in same order .If yes print "its a match" if no then print "its gone" in function
# -
lst= [5,4,1,6,2]
sublst=[5,4,1]
for i in range(len(lst)):
if lst[i] == sublst[0]:
for j in range(i+1, len(lst)):
if lst[j] == sublst[1]:
for k in range(j+1,len(lst)):
if lst[k] == sublst[2]:
print("its a match")
break
break
break
else:
print("its gone")
# +
#Function for prime number and use filter to filter out all prime numbers from 1-2500
# -
def isPrime(x):
for n in range(2,x):
if x%n==0:
return False
else:
return True
fltrObj=filter(isPrime, range(1,2500))
print ('Prime numbers are:', list(fltrObj))
# +
#Make a Lambda function for capitalizing whole sentence passed using arguments and map all sentences in the list with lambda function
# +
lst = [input()]
lst_arg = map(lambda x: x.title(),lst)
print(list(lst_arg))
# -
| python-day5-assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import keras
import os
from deepsense import neptune
from keras_retinanet.bin.train import create_models
from keras_retinanet.callbacks.eval import Evaluate
from keras_retinanet.models.resnet import download_imagenet, resnet_retinanet as retinanet
from keras_retinanet.preprocessing.detgen import DetDataGenerator
from keras_retinanet.preprocessing.csv_generator import CSVGenerator
from keras_retinanet.utils.transform import random_transform_generator
from keras_retinanet.preprocessing.detgen import DetDataGenerator
from detdata import DetGen
from detdata.augmenters import crazy_augmenter
from keras_retinanet.utils.eval import evaluate
import matplotlib.pyplot as plt
plt.style.use('ggplot')
# %matplotlib inline
import warnings
warnings.filterwarnings("ignore")
ctx = neptune.Context()
detg= DetGen('/home/i008/malaria_data/dataset_train.mxrecords',
'/home/i008/malaria_data/dataset_train.csv',
'/home/i008/malaria_data/dataset_train.mxindex', batch_size=4)
train_generator = DetDataGenerator(detg, augmenter=crazy_augmenter)
train_generator.image_max_side = 750
train_generator.image_min_side = 750
weights = download_imagenet('resnet50')
model_checkpoint = keras.callbacks.ModelCheckpoint('mod-{epoch:02d}_loss-{loss:.4f}.h5',
monitor='loss',
verbose=2,
save_best_only=False,
save_weights_only=False,
mode='auto',
period=1)
callbacks = []
class LossHistory(keras.callbacks.Callback):
def on_train_begin(self, logs={}):
self.losses = []
def on_batch_end(self, batch, logs={}):
ctx.channel_send("loss", logs.get('loss'))
def on_epoch_end(self, epoch, logs={}):
ctx.channel_send("val_loss", logs.get('val_loss'))
callbacks.append(keras.callbacks.ReduceLROnPlateau(
monitor='loss',
factor=0.1,
patience=2,
verbose=1,
mode='auto',
epsilon=0.0001,
cooldown=0,
min_lr=0
))
# callbacks.append(LossHistory())
callbacks.append(model_checkpoint)
model, training_model, prediction_model = create_models(
backbone_retinanet=retinanet,
backbone='resnet50',
num_classes=train_generator.num_classes(),
weights=weights,
multi_gpu=0,
freeze_backbone=True
)
# training_model.fit_generator(
# generator=train_generator,
# steps_per_epoch=5000,
# epochs=100,
# verbose=1,
# callbacks=callbacks,
# )
# +
# training_model.fit_generator(
# generator=train_generator,
# steps_per_epoch=5000,
# epochs=100,
# verbose=1,
# callbacks=callbacks,
# )
# -
model.load_weights('mod-40_loss-2.0113.h5')
import pandas as pd
df = pd.read_csv('/home/i008/malaria_data/dataset_train.csv')
df = df[['fname','xmin','ymin','xmax','ymax','class_name']]
df['fname'] = '/home/i008/googledrive/Projects/malaria/malaria_dataset/' + df.fname
df.to_csv('csvitertrain.csv', header=False, index=False)
# %%writefile cls.csv
Malaria, 0
g = CSVGenerator('csviterval.csv', 'cls.csv')
x, y =g.next()
model.predict(x)
# average_precisions, recall, precision, true_positives, false_positives = evaluate(g, model, score_threshold=0.1)
average_precisions
# +
plt.plot(recall, precision)
plt.xlabel('recall')
plt.ylabel('precision')
plt.style.use('ggplot')
import seaborn as sns
# -
true_positives / max(true_positives)
plt.plot(false_positives/max(false_positives), true_positives/max(true_positives))
plt.xlabel("false alarmrate (false positives)")
plt.ylabel("detection rate (true positives)")
# +
precisions = [1, 1, 1, 0.75, 0.6, 0.5, 0.42]
recalls = [0.1, 0.2, 0.3, 0.3, 0.3, 0.3, 0.3]
plt.plot(recalls, precisions, '*')
# +
import numpy as np
sorted_detections = np.array([0.6, 0.5, 0.49, 0.3, 0.2, 1.5, 0.1, 0.09, 0.05, 0.01, 0.01] )
sorted_detections_simplify = [1] * 11
positives = np.array([1,1,1,0,0,0,0,0,0,0,1])
assert len(sorted_detections_simplify) == len(positives)
all_true = sum(positives)
correct_detectio = 0
R = []
P = []
for i, (det, true) in enumerate(zip(sorted_detections_simplify, positives)):
i+=1
i=i * 1.0
if det == true:
correct_detectio+=1
precision = correct_detectio / i
recall = correct_detectio / len(positives)
print("top {} recall={} precision={}".format(i, recall, precision))
R.append(recall)
P.append(precision)
# -
plt.plot(R, P)
| train_retina_aiscope.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import scipy
import matplotlib.pyplot as plt
# %matplotlib inline
# Functions are input and output machines. Functions have a domain and a range. Functions can be composed together to create another composite function.
# +
def compose_functions(f1,f2,domain):
composition=f2(f1(domain))
plt.plot(domain,composition)
plt.show()
domain=np.arange(0,1,0.001)
f1=lambda x: 1-x**2
f2=lambda x: np.sqrt(x)
compose_functions(f1,f2,domain)
# -
# Functions can be inverted by flipping them along the diagonal line giving the inverse function.
# +
def invert(f1,domain):
y=f1(domain)
plt.plot(domain,y)
plt.plot(y,domain)
plt.plot(domain,domain)
domain=np.arange(-1,1,0.001)
f1=lambda x: x**3
invert(f1,domain)
# -
# Polynomials are special functions that are used in power series.
# +
def polynomial(coef,domain):
pol=np.polynomial.polynomial.Polynomial(coef=coef,domain=domain)
x,y=pol.linspace()
plt.plot(x,y)
coef=[1,2,3]
domain=[-100,100]
polynomial(coef,domain)
# -
# Rational functions consist of a polynomial divided by a polynomial.
# +
def rational_function(coef1,coef2,domain):
quotient,remainder=np.polydiv(coef1,coef2)
#x,y=pol.linspace(rat)
pol1=np.polynomial.polynomial.Polynomial(coef=coef1,domain=domain)
x1,y1=pol1.linspace()
pol2=np.polynomial.polynomial.Polynomial(coef=coef2,domain=domain)
x2,y2=pol2.linspace()
rat=y1/y2
plt.plot(x1,rat)
coef1=(1,2,3)
coef2=(4,5,6,10)
domain=[-100,100]
rational_function(coef1,coef2,domain)
# -
# Trigonometric functions and inverse trigonometric functions model geometric relationships.
domain=np.arange(-2*np.pi,2*np.pi,0.001)
sin=np.sin(domain)
cos=np.cos(domain)
tan=np.clip(sin/cos,a_min=-10,a_max=10)
cot=1./tan
sec=1./cos
csc=1./sin
plt.plot(domain,sin)
plt.plot(domain,cos)
plt.legend(('sin(x)','cos(x)'))
plt.show()
plt.plot(domain,tan)
plt.legend('tan(x)')
plt.show()
domain=np.arange(-1,1,0.001)
domain2=np.arange(-10,10,0.001)
arcsin=np.arcsin(domain)
arccos=np.arccos(domain)
arctan=np.arctan(domain2)
plt.plot(domain,arcsin)
plt.plot(domain,arccos)
plt.legend(('arcsin(x)','arccos(x)'))
plt.show()
plt.plot(domain2,arctan)
plt.legend('arctan(x)')
plt.show()
# Exponential functions and natural logarithms are reciprocals of each other.
domain=np.arange(-10,10,0.001)
e_x=np.e**domain
domain2=np.arange(0,10,0.001)
ln_x=np.log(domain2)
plt.plot(domain,e_x)
plt.legend(('e**x'))
plt.show()
plt.plot(domain2,ln_x)
plt.legend(('ln(x)'))
plt.show()
# Complex exponentials in the Euler formula connect imaginary numbers and trigonometric functions.
domain=np.arange(-10,10,0.001)
complex_exp=np.e**((1j)*domain)
plt.plot(domain,complex_exp.imag)
plt.plot(domain,complex_exp.real)
plt.legend(('Imaginary component','Real component'))
| Functions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Amazon Stock Analysis
# # Fama French 3 Factor Model
#
# The Fama and French model has three factors: size of firms, book-to-market values and excess return on the market. In other words, the three factors used are SMB (small minus big), HML (high minus low) and the portfolio's return less the risk free rate of return.
# +
#Importing Packages
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import scipy.stats as ss
from sklearn.metrics import mean_squared_error
from sklearn.metrics import r2_score
# -
df_pfe = pd.read_csv ('pfe_fama3_new.csv')
df_pfe
df_pfe.shape
# +
df_pfe['stock_return'] = (df_pfe['Adj Close']-df_pfe['Yest_Close'])/(df_pfe['Yest_Close'])
T = df_pfe.shape[0]
Y = df_pfe['stock_return'].values
columns = ['MktRF','SMB','HML']
X=df_pfe[columns]
X = np.column_stack([np.ones((len(X),1)),X])
N = X.shape
Y=np.asarray(Y)
Y = Y.reshape(T,1)
'REGRESSION STARTS:'
'Linear Regression of Y: T x 1 on'
'Regressors X: T x N'
invXX = np.linalg.inv(X.transpose()@X)
'OLS estimator beta: N x 1'
beta_hat = invXX@X.transpose()@Y
'Predictive value of Y_t using OLS'
y_hat = X@beta_hat;
'Residuals from OLS: Y - X*beta'
residuals = Y - y_hat;
'variance of Y_t or residuals'
sigma2 = (1/T)*(residuals.transpose()@residuals)
'standard deviation of Y_t or residuals'
sig = np.sqrt(sigma2)
'variance-covariance matrix of beta_hat'
'N x N: on-diagnal variance(beta_j)'
'N x N: off-diagnal cov(beta_i, beta_j)'
varcov_beta_hat = (sigma2)*invXX
var_beta_hat = np.sqrt(T*np.diag(varcov_beta_hat))
'Calculate R-square'
R_square = 1 - residuals.transpose()@residuals/(T*np.var(Y))
adj_R_square = 1-(1-R_square)*(T-1)/(T-N[1])
'Test Each Coefficient: beta_i'
't-test stat: N x 1'
t_stat = beta_hat.transpose()/var_beta_hat
' t-test significance level: N x 1'
p_val_t = 1-ss.norm.cdf(t_stat)
'Test of Joint Significance of Model'
F_stat = beta_hat.transpose()@varcov_beta_hat@beta_hat/\
(residuals.transpose()@residuals)
'size: (1 x N)*(N x N)*(N x 1)/((1 x T) * (T x 1)) = 1 x 1'
p_val_F = 1-ss.f.cdf(F_stat,N[1]-1,T-N[1])
print(' REGRESSION STATISTICS \n')
print('------------------------\n')
print('\n Joint significance of all coefficients\n',[F_stat,p_val_F])
print('Beta Values \n',beta_hat)
print('P values \n',p_val_t)
print('R-Square is \n',R_square)
print('Adjusted R Square \n',adj_R_square)
print('Standard Error \n',sig)
print('Observations \n',T)
print('-------------------------\n')
plt.plot(y_hat, color='blue')
plt.plot(Y, color = 'red')
plt.show()
pred = pd.DataFrame(y_hat)
act = pd.DataFrame(Y)
plot_df = pd.DataFrame({"actual": act[0], "predictions": pred[0]})
plot_df.plot(figsize=(20, 6), title='Predictions using FF3 using Linear Regression')
# -
mse = mean_squared_error(Y,y_hat)
rmse = np.sqrt(mse)
print('RMSE-------',rmse)
print('R-Squared--',r2_score(Y,y_hat))
| AmazonStockAnalysis-main/Pfizer_FamaFrench3_Model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Useful Built-in Functions
# ---
#
# These functions that we will learn are already programmed into the python language. We will learn how to use these functions to augment our code.
#
# ### Maximum and Minimum Functions
#
# ```max()``` and ```min()``` are built-in functions that help us find the extremes of its arguments.
#
# __Single Argument Use:__ When a function only has one argument.
#
# For max and min, the single argument must be either string or list. (Technically, iterable & comparable ... grade 12 definition)
# +
# Single Argument Example
word = '<PASSWORD>!'
print('Max of', word, 'is:', max(word))
print('Min of', word, 'is:', min(word))
# -
# __Explanation:__
#
# - When using max or min on a list or a string, we compare each individual item or character
# - For strings, we compare them by using the character's [ASCII value](http://www.asciitable.com/)
#
# __Multi-Argument Use:__ When a function has multiple arguments.
#
# For max and min, the arguments themselves will be compared to each other and output the appropriate extreme.
#
# ```max()``` and ```min()``` are special functions that can take any number of arguments.
# +
# Multi-Argument Example
print(max(7, 18, 15, 9, 18, 6, 21, 9, 4, 8))
print(min(9, 18, 6, 21, 9, 4, 8))
# -
# __NOTE:__
#
# We can assign the result of max() and min() function to a variable as well.
# example
result = max('hello')
print(result) # should be o
# ### Length Function
#
# The ```len()``` is the length function which is a __single argument__ function.
#
# The ```len()``` function is used to find the length of collection; moreover, the number of items in the collection.
#
# The only collection type data types in Grade 10 and 11 are strings and lists.
#
# We cannot find the length of non-collection based data types (int, float)
# len() examples
print(len('abcdefg')) # outputs 7 because there are 7 characters in the string
print(len([1,2,3,4,5,6])) # Will return 6 because there are 6 items in the list
| 5 Useful Functions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Training a neural network on QM9
#
# This tutorial will explain how to use SchNetPack for training a model
# on the QM9 dataset and how the trained model can be used for further.
#
# First, we import the necessary modules and create a new directory for the data and our model.
# +
import os
import schnetpack as spk
qm9tut = './qm9tut'
if not os.path.exists('qm9tut'):
os.makedirs(qm9tut)
# -
# ## Loading the data
#
# As explained in the [previous tutorial](tutorial_01_preparing_data.ipynb), datasets in SchNetPack are loaded with the `AtomsLoader` class or one of the sub-classes that are specialized for common benchmark datasets.
# The `QM9` dataset class will download and convert the data. We will only use the inner energy at 0K `U0`, so all other properties do not need to be loaded:
# +
from schnetpack.datasets import QM9
qm9data = QM9('./qm9.db', download=True, load_only=[QM9.U0], remove_uncharacterized=True)
# -
# ### Splitting the data
#
# Next, we split the data into training, validation and test set. Here, we choose to use 1000 training examples, 1000 examples for validation and the remaining data as test set.
# The corresponding indices are stored in the `split.npz` file.
train, val, test = spk.train_test_split(
data=qm9data,
num_train=1000,
num_val=500,
split_file=os.path.join(qm9tut, "split.npz"),
)
# Next, we will create an `AtomsLoader` for each split.
# This will take care of shuffling, batching and asynchronously loading the data during training and evaluation.
train_loader = spk.AtomsLoader(train, batch_size=100, shuffle=True)
val_loader = spk.AtomsLoader(val, batch_size=100)
# ### Dataset statistics
#
# Before building the model, we need some statistic about our target property for good initial conditions. We will get this from the training examples.
# For QM9, we also have single-atom reference values stored in the metadata:
atomrefs = qm9data.get_atomref(QM9.U0)
print('U0 of hyrogen:', '{:.2f}'.format(atomrefs[QM9.U0][1][0]), 'eV')
print('U0 of carbon:', '{:.2f}'.format(atomrefs[QM9.U0][6][0]), 'eV')
print('U0 of oxygen:', '{:.2f}'.format(atomrefs[QM9.U0][8][0]), 'eV')
# These can be used together with the mean and standard deviation of the energy per atom to initialize the model with a good guess of the energy of a molecule. When calculating these statistics, we pass the atomref to take into account, that the model will add these atomrefs to the predicted energy later, so that this part of the energy does not have to be considered in the statistics, i.e.
# \begin{equation}
# \mu_{U_0} = \frac{1}{n_\text{train}} \sum_{n=1}^{n_\text{train}} \left( U_{0,n} - \sum_{i=1}^{n_{\text{atoms},n}} U_{0,Z_{n,i}} \right)
# \end{equation}
# for the mean and analog for the standard deviation. In this case, this corresponds to the mean and std. dev of the *atomization energy* per atom.
means, stddevs = train_loader.get_statistics(
QM9.U0, get_atomwise_statistics=True, single_atom_ref=atomrefs
)
print('Mean atomization energy / atom:', means[QM9.U0])
print('Std. dev. atomization energy / atom:', stddevs[QM9.U0])
# ## Building the model
# The next step is to build the neural network model.
# This consists of 2 parts:
#
# 1. The representation part which either constructs atom-wise features, e.g. with SchNet, or build a fixed descriptor such as atom-centered symmetry functions.
# 2. One or more output modules for property prediction.
#
# We will use a `SchNet` module with 3 interaction layers, a 5 Angstrom cosine cutoff with pairwise distances expanded on 20 Gaussians and 50 atomwise features and convolution filters here, since we only have a few training examples (click [here](../modules/representation.rst#module-schnetpack.representation.schnet) for details on SchNet). Then, we will use one `Atomwise` modules to predict the energy, which takes mean and standard deviation per atom of the property for initialization. Both modules are then combined to an `AtomisticModel`.
# +
schnet = spk.representation.SchNet(
n_atom_basis=30, n_filters=30, n_gaussians=20, n_interactions=5,
cutoff=4., cutoff_network=spk.nn.cutoff.CosineCutoff
)
output_U0 = spk.atomistic.Atomwise(n_in=30, atomref=atomrefs[QM9.U0], property=QM9.U0,
mean=means[QM9.U0], stddev=stddevs[QM9.U0])
model = spk.AtomisticModel(representation=schnet, output_modules=output_U0)
# -
# ## Training the model
#
# To train the model, we will use the `Trainer` class that comes with SchNetPack.
# For this, we need to first define a loss function and select an optimizer.
# As the loss function, we will use the mean squared error of the energy
# $\ell(E_\text{ref}, E_\text{pred}) = \frac{1}{n_\text{train}} \sum_{n=1}^{n_\text{train}} (E_\text{ref} - E_\text{pred})^2$
#
# This will be minimized using the Adam optimizer from PyTorch.
# +
from torch.optim import Adam
# loss function
def mse_loss(batch, result):
diff = batch[QM9.U0]-result[QM9.U0]
err_sq = torch.mean(diff ** 2)
return err_sq
# build optimizer
optimizer = Adam(model.parameters(), lr=1e-2)
# -
# We can give the `Trainer` hooks, that are called at certain points during the training loop.
# This is useful to customize the training process, e.g. with logging, learning rate schedules or stopping criteria.
# Here, we set up a basis logging as well as a learning rate schedule that reduces the learning rate by factor 0.8 after 5 epochs without improvement of the validation loss.
#
# The logger takes a list of validation metrics that specify what is going to be stored.
# In this example, we want to log the mean absolute and root mean squared error of the $U_0$ energy prediction.
# +
# before setting up the trainer, remove previous training checkpoints and logs
# %rm -r ./qm9tut/checkpoints
# %rm -r ./qm9tut/log.csv
import schnetpack.train as trn
loss = trn.build_mse_loss([QM9.U0])
metrics = [spk.metrics.MeanAbsoluteError(QM9.U0)]
hooks = [
trn.CSVHook(log_path=qm9tut, metrics=metrics),
trn.ReduceLROnPlateauHook(
optimizer,
patience=5, factor=0.8, min_lr=1e-6,
stop_after_min=True
)
]
trainer = trn.Trainer(
model_path=qm9tut,
model=model,
hooks=hooks,
loss_fn=loss,
optimizer=optimizer,
train_loader=train_loader,
validation_loader=val_loader,
)
# -
# We can run the training for a given number of epochs. If we don't give a number, the trainer runs until a stopping criterion is met. For the purpose of this tutorial, we let it train for 200 epochs (on GPU this should take about x minutes).
device = "cuda" # change to 'cpu' if gpu is not available
n_epochs = 300 # takes about 10 min on a notebook GPU. reduces for playing around
trainer.train(device=device, n_epochs=n_epochs)
# You can also call this method several times to continue training. For the training to run until convergence, remove the n_epochs argument or set it to a very large number.
#
# Let us finally have a look at the CSV log:
# +
import numpy as np
import matplotlib.pyplot as plt
from ase.units import kcal, mol
results = np.loadtxt(os.path.join(qm9tut, 'log.csv'), skiprows=1, delimiter=',')
time = results[:,0]-results[0,0]
learning_rate = results[:,1]
train_loss = results[:,2]
val_loss = results[:,3]
val_mae = results[:,4]
print('Final validation MAE:', np.round(val_mae[-1], 2), 'eV =',
np.round(val_mae[-1] / (kcal/mol), 2), 'kcal/mol')
plt.figure(figsize=(14,5))
plt.subplot(1,2,1)
plt.plot(time, val_loss, label='Validation')
plt.plot(time, train_loss, label='Train')
plt.yscale('log')
plt.ylabel('Loss [eV]')
plt.xlabel('Time [s]')
plt.legend()
plt.subplot(1,2,2)
plt.plot(time, val_mae)
plt.ylabel('mean abs. error [eV]')
plt.xlabel('Time [s]')
plt.show()
# -
# Of course, the prediction can be improved by letting the training run longer, increasing the patience, the number of neurons and interactions or using regularization techniques.
# ## Using the model
#
# Having trained a model for QM9, we are going to use it to obtain some predictions.
# First, we need to load the model. The `Trainer` stores the best model in the model directory which can be loaded using PyTorch:
import torch
best_model = torch.load(os.path.join(qm9tut, 'best_model'))
# To evaluate it on the test data, we create a data loader for the test split:
# +
test_loader = spk.AtomsLoader(test, batch_size=100)
err = 0
print(len(test_loader))
for count, batch in enumerate(test_loader):
# move batch to GPU, if necessary
batch = {k: v.to(device) for k, v in batch.items()}
# apply model
pred = best_model(batch)
# calculate absolute error
tmp = torch.sum(torch.abs(pred[QM9.U0]-batch[QM9.U0]))
tmp = tmp.detach().cpu().numpy() # detach from graph & convert to numpy
err += tmp
# log progress
percent = '{:3.2f}'.format(count/len(test_loader)*100)
print('Progress:', percent+'%'+' '*(5-len(percent)), end="\r")
err /= len(test)
print('Test MAE', np.round(err, 2), 'eV =',
np.round(err / (kcal/mol), 2), 'kcal/mol')
# -
# If your data is not already in SchNetPack format, a more convenient way is to use ASE atoms objects with the provided `AtomsConverter`.
# +
converter = spk.data.AtomsConverter(device=device)
at, props = test.get_properties(idx=153)
inputs = converter(at)
print('Keys:', list(inputs.keys()))
print('Truth:', props[QM9.U0].cpu().numpy()[0])
pred = model(inputs)
print('Prediction:', pred[QM9.U0].detach().cpu().numpy()[0,0])
# -
# Alternatively, one can use the `SpkCalculator` as an interface to ASE:
calculator = spk.interfaces.SpkCalculator(model=model, device=device, energy=QM9.U0)
at.set_calculator(calculator)
print('Prediction:', at.get_total_energy())
# ## Summary
# In this tutorial, we have trained and evaluated a SchNet model on a small subset of QM9.
# A full training script with is available [here](../../src/examples/qm9_tutorial.py).
#
# In the next tutorial, we will show how to learn potential energy surfaces and forces field as well as performing molecular dynamics simulations with SchNetPack.
| docs/tutorials/tutorial_02_qm9.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#get Iris test dataset using sklearn library
from sklearn import datasets
import numpy as np
iris = datasets.load_iris()
X = iris.data[:, [2, 3]] #take only petal lengh and petal width parameters from the Iris dataset
y = iris.target
#split test data into 30% testing and 70% training data
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=0)
#feature scaling (standardize) using the StandardScaler
from sklearn.preprocessing import StandardScaler
sc = StandardScaler()
sc.fit(X_train)
X_train_std = sc.transform(X_train)
X_test_std = sc.transform(X_test)
#train Perceptron using One-vs-Rest method (all 3 flower classes at once)
from sklearn.linear_model import Perceptron
ppn = Perceptron(n_iter=40, eta0=0.1, random_state=0)
ppn.fit(X_train_std, y_train)
| ch03/.ipynb_checkpoints/01-scikit-learn-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: nlp
# language: python
# name: nlp
# ---
# +
# default_exp data
# -
# # Data
#
# > How to prepare dataset for our experiments?
#hide
from nbdev.showdoc import *
# +
#export
import torchtext
from inspect import signature
from fastai.text.all import *
from sklearn.feature_extraction.text import CountVectorizer
# -
# ### How to generate text pairs using Fast.AI library?
# - We want to show how we could use the low-level API provided by Fast.AI to build a custom dataset for our task.
# - We want to identify if a pair of text are duplicate of each other or not.
# - The data format would be something like ((t1, t2), label), where t1 and t2 represent the text and label represents a boolean variable indicating whether they are duplicate or not.
# - We will make use of the transforms provided by Fast.AI to build the dataset and dataloaders required by our model to process.
class SentenceSimTuple(fastuple):
def show(self, ctx=None, **kwargs):
df = pd.DataFrame({'a': [self[0]],
'b': [self[1]]
})
display_df(df)
class TextPairGetter(ItemTransform):
def __init__(self, s1='a', s2='b',target='target'):
store_attr('s1,s2,target')
def encodes(self, o):
return o[self.s1], o[self.s2]
class BOWVectorizer(ItemTransform):
def __init__(self, vec):
store_attr('vec')
def encodes(self, o):
ftok = self.vec.transform(np.array([o[0]]))
stok = self.vec.transform(np.array([o[1]]))
return ftok.toarray() * 1., stok.toarray() * 1.
def decodes(self, o):
forig = self.vec.inverse_transform(o[0])
sorig = self.vec.inverse_transform(o[1])
return SentenceSimTuple((TitledStr(' '.join(forig[0])), TitledStr(' '.join(sorig[0]))))
sample = pd.DataFrame({'a': ['this is a good life', 'slow life', 'am i good', 'waiting for'],
'b': ['take it easy', 'I am on the moon, rythm is right.', 'truely madly', 'for waiting'],
'target': [0, 1, 0, 1]
})
vec = CountVectorizer()
vec = vec.fit(sample['a'].tolist() + sample['b'].tolist())
dset = Datasets(sample, [[TextPairGetter(), BOWVectorizer(vec)], [ItemGetter('target'), Categorize()]])
x, y = dset.decode(dset[0])
show_at(dset, 1)
dls = dset.dataloaders(bs=2)
dls._types
x, y = dls.one_batch()
# ### How to override show_batch method for our dataset?
@typedispatch
def show_batch(x:tuple, y, samples, ctxs=None, max_n=10, trunc_at=150, **kwargs):
if ctxs is None: ctxs = get_empty_df(min(len(samples), max_n))
if isinstance(samples[0][0], tuple):
samples = L((*s[0], *s[1:]) for s in samples)
if trunc_at is not None: samples = L((s[0].truncate(trunc_at), s[1].truncate(trunc_at), *s[2:]) for s in samples)
if trunc_at is not None: samples = L((s[0].truncate(trunc_at),*s[1:]) for s in samples)
ctxs = show_batch[object](x, y, samples, max_n=max_n, ctxs=ctxs, **kwargs)
display_df(pd.DataFrame(ctxs))
dls.show_batch()
# ## Quora Questions Pair Dataset
# +
#slow
BASE_DIR = Path('~/data/dl_nlp')
RAW_DATA_PATH = BASE_DIR / 'data' / 'quodup'
train = pd.read_csv(RAW_DATA_PATH / 'train.csv')
train = train.sample(frac=1.)
train.index = np.arange(len(train))
# fill empty questions with ''
train.loc[:, 'question1'] = train.question1.fillna('')
train.loc[:, 'question2'] = train.question2.fillna('')
train.head()
# +
#slow
# %%time
splits = IndexSplitter(np.arange(len(train)-int(.2 * len(train)), len(train)))(train)
combined_df = pd.DataFrame({'text': list(train.iloc[splits[0]]['question1'].unique()) + list(train.iloc[splits[0]]['question2'].unique())})
_, cnt = tokenize_df(combined_df, text_cols='text')
# -
#export
class NumericalizePair(Numericalize):
def encodes(self, o):
return TensorText(tensor([self.o2i [o_] for o_ in o['q1']])), TensorText(tensor([self.o2i [o_] for o_ in o['q2']]))
#slow
dset = Datasets(train, [[Tokenizer.from_df('question1', tok_text_col='q1'), Tokenizer.from_df('question2', tok_text_col='q2'),
NumericalizePair(vocab=list(cnt.keys()))], [ItemGetter('is_duplicate'), Categorize()]], splits=splits)
#slow
dset[0]
#slow
dset.decode(dset[0])
#export
class Pad_Chunk_Pair(ItemTransform):
"Pad `samples` by adding padding by chunks of size `seq_len`"
def __init__(self, pad_idx=1, pad_first=True, seq_len=72,decode=True,**kwargs):
store_attr('pad_idx, pad_first, seq_len,seq_len')
super().__init__(**kwargs)
def before_call(self, b):
"Set `self.max_len` before encodes"
xas, xbs = [], []
for xs in b:
xa, xb = xs[0]
if isinstance(xa, TensorText):
xas.append(xa.shape[0])
if isinstance(xb, TensorText):
xbs.append(xb.shape[0])
self.max_len_a = max(xas)
self.max_len_b = max(xbs)
def __call__(self, b, **kwargs):
self.before_call(b)
return super().__call__(tuple(b), **kwargs)
def encodes(self, batch):
texts = ([s[0][0] for s in batch], [s[0][1] for s in batch])
labels = default_collate([s[1:] for s in batch])
inps = {}
pa = default_collate([pad_chunk(ta,pad_idx=self.pad_idx, pad_first=self.pad_first, seq_len=self.seq_len, pad_len=self.max_len_a) for ta in texts[0]])
pb = default_collate([pad_chunk(tb,pad_idx=self.pad_idx, pad_first=self.pad_first, seq_len=self.seq_len, pad_len=self.max_len_b) for tb in texts[1]])
inps['pa'] = pa
inps['pb'] = pb
if len(labels):
inps['labels'] = labels[0]
res = (inps, )
return res
#export
class Undict(Transform):
def decodes(self, x:dict):
if 'pa' in x and 'pb' in x: res = (x['pa'], x['pb'], x['labels'])
return res
# +
#slow
seq_len = 72
dls_kwargs = {
'before_batch': Pad_Chunk_Pair(seq_len=seq_len),
'after_batch': Undict(),
'create_batch': fa_convert
}
dls = dset.dataloaders(bs=128, seq_len=seq_len, **dls_kwargs)
# -
#slow
x = dls.one_batch()
#slow
x
#slow
xd = dls.decode(x)
xd
#export
def load_dataset():
BASE_DIR = Path('~/data/dl_nlp')
RAW_DATA_PATH = BASE_DIR / 'data' / 'quodup'
train = pd.read_csv(RAW_DATA_PATH / 'train.csv')
train = train.sample(frac=1.)
train.index = np.arange(len(train))
# fill empty questions with ''
train.loc[:, 'question1'] = train.question1.fillna('')
train.loc[:, 'question2'] = train.question2.fillna('')
return train
| 00_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # A Network Tour of Data Science
#
# [<NAME>](http://deff.ch), *PhD student*, [<NAME>](https://people.epfl.ch/pierre.vandergheynst), *Full Professor*, [EPFL](http://epfl.ch) [LTS2](http://lts2.epfl.ch).
#
# # Exercise 5: Graph Signals and Fourier Transform
#
# The goal of this exercise is to experiment with the notions of graph signals, graph Fourier transform and smoothness and illustrate these concepts in the light of clustering.
import numpy as np
import scipy.spatial
import matplotlib.pyplot as plt
# %matplotlib inline
# ## 1 Graph
#
# **Goal**: compute the combinatorial Laplacian $L$ of a graph formed with $c=2$ clusters.
#
# **Step 1**: construct and visualize a fabricated data matrix $X = [x_1, \ldots, x_n]^t \in \mathbb{R}^{n \times d}$ whose lines are $n$ samples embedded in a $d$-dimensional Euclidean space.
# +
d = 2 # Dimensionality.
n = 100 # Number of samples.
c = 1 # Number of communities.
# Data matrix, structured in communities.
X = np.random.uniform(0, 1, (n, d))
X += np.linspace(0, 2, c).repeat(n//c)[:, np.newaxis]
fig, ax = plt.subplots(1, 1, squeeze=True)
ax.scatter(X[:n//c, 0], X[:n//c, 1], c='b', s=40, linewidths=0, label='class 0');
ax.scatter(X[n//c:, 0], X[n//c:, 1], c='r', s=40, linewidths=0, label='class 1');
lim1 = X.min() - 0.5
lim2 = X.max() + 0.5
ax.set_xlim(lim1, lim2)
ax.set_ylim(lim1, lim2)
ax.set_aspect('equal')
ax.legend(loc='upper left');
# -
# **Step 2**: compute all $n^2$ pairwise euclidean distances $\operatorname{dist}(i, j) = \|x_i - x_j\|_2$.
#
# Hint: you may use the function `scipy.spatial.distance.pdist()` and `scipy.spatial.distance.squareform()`.
# +
# Pairwise distances.
dist = scipy.spatial.distance.pdist(X, metric='euclidean')
dist = scipy.spatial.distance.squareform(dist)
plt.figure(figsize=(15, 5))
plt.hist(dist.flatten(), bins=40);
# -
# Step 3: order the distances and, for each sample, solely keep the $k=10$ closest samples to form a $k$ nearest neighbor ($k$-NN) graph.
#
# Hint: you may sort a numpy array with `np.sort() or np.argsort()`.
# +
k = 10 # Miminum number of edges per node.
idx = np.argsort(dist)[:, 1:k+1]
dist.sort()
dist = dist[:, 1:k+1]
assert dist.shape == (n, k)
# -
# Step 4: compute the weights using a Gaussian kernel, i.e. $$\operatorname{weight}(i, j) = \exp\left(-\frac{\operatorname{dist}(i,j)^2}{\sigma^2}\right) = \exp\left(-\frac{\|x_i - x_j\|_2^2}{\sigma^2}\right).$$
#
# Hint: you may use the below definition of $\sigma^2$.
# +
# Scaling factor.
sigma2 = np.mean(dist[:, -1])**2
# Weights with Gaussian kernel.
dist = np.exp(- dist**2 / sigma2)
plt.figure(figsize=(15, 5))
plt.hist(dist.flatten(), bins=40);
# -
# Step 5: construct and visualize the sparse weight matrix $W_{ij} = \operatorname{weight}(i, j)$.
#
# Hint: you may use the function `scipy.sparse.coo_matrix()` to create a sparse matrix.
# +
# Weight matrix.
I = np.arange(0, n).repeat(k)
J = idx.reshape(n*k)
V = dist.reshape(n*k)
W = scipy.sparse.coo_matrix((V, (I, J)), shape=(n, n))
# No self-connections.
W.setdiag(0)
# Non-directed graph.
bigger = W.T > W
W = W - W.multiply(bigger) + W.T.multiply(bigger)
assert type(W) == scipy.sparse.csr_matrix
print('n = |V| = {}, k|V| < |E| = {}'.format(n, W.nnz))
plt.spy(W, markersize=2, color='black');
import scipy.io
import os.path
scipy.io.mmwrite(os.path.join('datasets', 'graph_inpainting', 'embedding.mtx'), X)
scipy.io.mmwrite(os.path.join('datasets', 'graph_inpainting', 'graph.mtx'), W)
# -
# Step 6: compute the combinatorial graph Laplacian $L = D - W$ where $D$ is the diagonal degree matrix $D_{ii} = \sum_j W_{ij}$.
# +
# Degree matrix.
D = W.sum(axis=0)
D = scipy.sparse.diags(D.A.squeeze(), 0)
# Laplacian matrix.
L = D - W
fig, axes = plt.subplots(1, 2, squeeze=True, figsize=(15, 5))
axes[0].spy(L, markersize=2, color='black');
axes[1].plot(D.diagonal(), '.');
# -
# ## 2 Fourier Basis
#
# Compute the eigendecomposition $L=U \Lambda U^t$ of the Laplacian, where $\Lambda$ is the diagonal matrix of eigenvalues $\Lambda_{\ell\ell} = \lambda_\ell$ and $U = [u_1, \ldots, u_n]^t$ is the graph Fourier basis.
#
# Hint: you may use the function `np.linalg.eigh()`.
# +
lamb, U = np.linalg.eigh(L.toarray())
#print(lamb)
plt.figure(figsize=(15, 5))
plt.plot(lamb, '.-');
# -
# 1. Visualize the eigenvectors $u_\ell$ corresponding to the first eight non-zero eigenvalues $\lambda_\ell$.
# 2. Can you explain what you observe and relate it to the structure of the graph ?
# +
def scatter(ax, x):
ax.scatter(X[:, 0], X[:, 1], c=x, s=40, linewidths=0)
ax.set_xlim(lim1, lim2)
ax.set_ylim(lim1, lim2)
ax.set_aspect('equal')
fig, axes = plt.subplots(2, 4, figsize=(15, 6))
for i, ax in enumerate(axes.flatten()):
u = U[:, i+1]
scatter(ax, u)
ax.set_title('u_{}'.format(i+1))
# -
# ## 3 Graph Signals
#
# 1. Let $f(u)$ be a positive and non-increasing function of $u$.
# 2. Compute the graph signal $x$ whose graph Fourier transform satisfies $\hat{x}(\ell) = f(\lambda_\ell)$.
# 3. Visualize the result.
# 4. Can you interpret it ? How does the choice of $f$ influence the result ?
# +
def f1(u, a=2):
y = np.zeros(n)
y[:a] = 1
return y
def f2(u):
return f1(u, a=3)
def f3(u):
return f1(u, a=n//4)
def f4(u):
return f1(u, a=n)
def f5(u, m=4):
return np.maximum(1 - m * u / u[-1], 0)
def f6(u):
return f5(u, 2)
def f7(u):
return f5(u, 1)
def f8(u):
return f5(u, 1/2)
def f9(u, a=1/2):
return np.exp(-u / a)
def f10(u):
return f9(u, a=1)
def f11(u):
return f9(u, a=2)
def f12(u):
return f9(u, a=4)
def plot(F):
plt.figure(figsize=(15, 5))
for f in F:
plt.plot(lamb, eval(f)(lamb), '.-', label=f)
plt.xlim(0, lamb[-1])
plt.legend()
F = ['f{}'.format(i+1) for i in range(12)]
plot(F[0:4])
plot(F[4:8])
plot(F[8:12])
# -
fig, axes = plt.subplots(3, 4, figsize=(15, 9))
for f, ax in zip(F, axes.flatten()):
xhat = eval(f)(lamb)
x = U.dot(xhat) # U @ xhat
#x = U.dot(xhat * U.T[:,2])
scatter(ax, x)
ax.set_title(f)
| algorithms/07_sol_graph_fourier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#Setup for Sympy
import sympy as sp
import numpy as np
# So far we have seen Trapezoidal Rule and Simpson Rule. In general, we may approximate the integral $$I = \int_a^b w(x) f(x) dx$$
# where $w \in C^0([a,b]), w>0$ and $w \in L^1((a,b))$ is a weight function, and $f \in L^1((a,b),w)$, by approximating $f$ as a polynomial.
#
# We may approximate $f$ with $n$-th order Largrange polynomial (sampling points $(x_i)_{i=0}^{k+1}$):
# \begin{equation}
# f(x) \approx \sum_{i=0}^n f(x_i) \ell_i(x) \quad \ell_i(x) = \prod_{0\leq j \leq i, \, j \neq i} \frac{x-x_j}{x_i-x_j}
# \end{equation}
#
# Then
# \begin{equation}
# \int_a^b w(x)f(x)dx \approx I_n = \sum_{i=0}^n f(x_i) \left(\int_a^b w(x) \ell_i(x) dx \right) = \sum_{i=0}^n w_i f(x_i)
# \end{equation}
#
# We want to choose $(x_i)_{i=0}^n$ such that $I_n(f) = I(f)$ for $f$ polynomial of degree at most $m$, $m>n$ is maximised.
# Turns out that if we choose roots of $\phi_{n+1}(x)$ (with $n+1$ degree), so that it belongs to the orthogonal basis of $L^2((a,b))$ (w.r.t. $\langle f,g \rangle = \int_a^b w(x) f(x)g(x) dx$, then $I_n(f) = I(f)$, $f$ any polynomial of degree at most $2n+1$. <br>
# Moreover, we can check by substituting $f = (\ell_i)^2$ that $w_i > 0$. We can find $w_i$ by such substitution, but this is usually tedious. Instead, we find $w_i$ by substituting $f = 1, x, x^2, ..., x_n$ and solve the simultaneous equation.
# # Example 1
# Let $f \in L^2([-1,1])$ and $w(x) = 1+4|x|$. Construct $I_2(f)$.
# 1. Construct Orthogonal Basis: Recall the following formula:
# $$\phi_j(x) = \bigg(x - \frac{\langle x\phi_{j-1}(x),\phi_{j-1}(x) \rangle}{\|\phi_{j-1}(x)\|^2} \bigg)\phi_{j-1}(x) - \frac{\|\phi_{j-1}(x)\|^2}{\|\phi_{j-2}(x)\|^2}\phi_{j-2}(x), j \geq 1$$
# where $\phi_{-1}(x)=0$ and $\phi_{0}(x)=1$.
# Define Symbols for inner product and norm
x = sp.symbols('x')
w = sp.Function('w')
f = sp.Function('f')
g = sp.Function('g')
# +
# inner product and norms
def inner(w,a,b,f,g):
output = sp.integrate(w*f*g, (x,a,b))
return output
def norm(w,a,b,f):
return sp.sqrt(inner(w,a,b,f,f))
# -
# Define symbols for basis
phi = sp.Function('phi')
phiminus1 = sp.Function('phiminus1')
phiminus2 = sp.Function('phiminus2')
# Iteration
def ortho(w,a,b,r):
phiminus2 = 1
phiminus1 = x - inner(w,a,b,x,1)/(norm(w,a,b,1))**2
if r == 0:
return [phiminus2]
elif r == 1:
return [phiminus2,phiminus1]
else:
philist = [phiminus2,phiminus1]
for i in range(r-1):
phi = (x - inner(w,a,b,x*phiminus1,phiminus1)/(norm(w,a,b,phiminus1))**2)*phiminus1 - ((norm(w,a,b,phiminus1)/norm(w,a,b,phiminus2))**2)*phiminus2
phi = sp.simplify(phi)
philist.append(phi)
phiminus2 = phiminus1
phiminus1 = phi
return philist
# Compute the basis, up to order 2+1 = 3
olist = ortho(1+4*sp.Abs(x),-1,1,3)
olist
# 2. Compute the roots of $\phi_3(x)$, as $x_0, x_1, x_2$.
rootlist = sp.solve(olist[3],x)
rootlist
# 3. Compute $b_0 = I(1), \, b_1 = I(x), \, b_2 = I(x^2)$
w = 1+4*sp.Abs(x)
b0 = sp.integrate(w,(x,-1,1))
b1 = sp.integrate(x*w,(x,-1,1))
b2 = sp.integrate((x**2)*w,(x,-1,1))
b = sp.Matrix([b0,b1,b2])
b
# 4. Compute $w_0, w_1, w_2$ by solving
# \begin{equation}
# \begin{cases}
# w_0 + w_1 + w_2 = b_0 \\
# w_0 x_0 + w_1 x_1 + w_2 x_2 = b_1 \\
# w_0 x_0^2 + w_1 x_1^2 + w_2 x_2^2 = b_2
# \end{cases}
# \end{equation}
A = sp.Matrix([[1,1,1],[rootlist[0],rootlist[1],rootlist[2]], [rootlist[0]**2,rootlist[1]**2,rootlist[2]**2]])
A
w0, w1, w2 = sp.symbols('w0,w1,w2')
sp.linsolve((A,b),[w0,w1,w2])
# # Please edit below
w = 1+x**2
a = 0
b = 2
n = 0
# Compute the basis, up to order n+1
olist = ortho(w,a,b,n+1)
olist
# Compute root of phi(n+1)
rootlist = sp.solve(olist[1],x)
rootlist
# Compute actual value of I(f) for f = 1, x,...
b0 = sp.integrate(w,(x,0,2))
b0
# Compute weight by solving equation
w0 = b0
w0
| M2AA3/M2AA3-Polynomials/Lesson 04 - Quadrature/Gaussian Quadrature.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Analysis of FSL, FF, and Metallcity in Nearby Galaxies
# Based on "analysis_nii_ff_bration.py" from 2015 and updated July 2017
# ## Setup Workbook
# +
# %matplotlib inline
import os.path
import numpy as np
import numpy.ma as ma
import matplotlib.pylab as plt
from astropy.io import fits
from astropy.io import ascii
from astropy.table import Table,join
from astropy import constants as const
import matplotlib.patches as mpatches
from astropy.cosmology import WMAP9
from astropy.cosmology import FlatLambdaCDM
from astropy import units as u
import matplotlib.patheffects as PathEffects
from astropy import constants
from matplotlib.patches import Rectangle
from matplotlib import colors
import astro_tools as at
# -
#Define a cosmology
cosmoWMAP=WMAP9
cosmoFLAT=FlatLambdaCDM(H0=70,Om0=0.3)
#define working directories
home = os.path.expanduser("~")
science_folder = os.path.join(home,'Dropbox','Projects')
active_folder = os.path.join(science_folder,'ff_fir_metalicity')
outfolder = os.path.join(active_folder,'outputs/')
#load the data on the local galaxies
#oiii_nii_ff_metal_location=os.path.join(active_folder,'oiii_nii_ff_metallicity.csv')
#L_FIR is lensing corrected for lensed galaxies
data_table = ascii.read("oiii_nii_ff_metal_location.csv",data_start=2)
for column in data_table.columns[2,3,5,6,8,9,11,12,14,15,17,18,20,21]:
data_table['%s' %column].unit = '1e-18 W m^-2'
for column in data_table.columns[24,25]:
data_table['%s' %column].unit='12+log(O/H)'
for column in data_table.columns[27,28,32,33]:
data_table['%s' %column].unit='mJy'
for column in data_table.columns[30,34]:
data_table['%s' %column].unit='GHz'
data_table.columns[30,34].unit = 'GHz'
data_table.columns[37].unit = 'Mpc'
# +
#Scientifically Useful Constants:
ff_fraction = 0.3 #The faction of the 5GHz Continuum that is attributable to free-free emission for star forming systems
def thermal_fraction(freq1,alpha):
freq2= 1.7 #GHz
alpha_nt=0.85 #murphy 2012
alpha_ff=0.1
upper = ((freq2/freq1)**-alpha-(freq2/freq1)**-alpha_nt)
lower = ((freq2/freq1)**-alpha_ff-(freq2/freq1)**-alpha_nt)
return upper/lower
# -
thermal_fraction(5,.9)
# ## Calculate the expected ratios of FSL/FF
# +
def free_free_specific_lum(frequency,temperature,gaunt_ff,electron_den,volume):
''' from Rybicki and Lightman 1985, assuminges hydrogen,i.e. charge_num, Z, is 1 and electron_den = ion_density
temperature must be in kelvin, freuquency in hertz, electron_den in cm^-3, and volume in cm^3
will return the luminosity in ergs/s/Hz
'''
specific_lum = 6.8e-38*electron_den**2*temperature**(-0.5)*np.exp(-const.h.value*frequency/(const.k_B.value*temperature))
return specific_lum
def fsl_luminosity_lowDen(frequency,electron_den,ion_fraction,abundance,collision_rate,volume):
'''
freuquency in hertz, ion fraction and abundnace as a decimal fraction, electron_den in cm^-3,
collisional rate coefficient in cm^3/s and volume in cm^3
will return the luminosity in J/s
'''
line_luminosity = const.h.to("erg s").value*frequency*electron_den**2*ion_fraction*abundance*collision_rate*volume
return line_luminosity
def collision_rate_coeff(temperature,stat_weight,collision_strength,frequency):
'''See Osterbrock & Ferland 2006
temperature in kelvin, freuquency in hertz,
returns collisional rate coefficient in cm^3/s
'''
collision_rate = 8.6e-6/temperature**(0.5)*collision_strength/stat_weight*np.exp(const.h.value*frequency/(const.k_B.value*temperature))
return collision_rate
line_frequencies = np.array([at.FSL_line_frequency('[NII] 122'),at.FSL_line_frequency('[OIII] 88'),
at.FSL_line_frequency('[CII] 158')])*10**9
radio_frequency = 5e9 #Hertz
gaunt_ff_5Ghz = 4.86
temperature = 8000 #Kelvin
volume = 1 #as long as it is the same then it is OK
electron_den = 1 #as long as it is the same then it is OK
collision_strengths_weights =np.array([('[NII] 122',0.25, 1), #from 3P0 to 3P2, the state weight for lower level, Hudson & Bell 2004
('[OIII] 88',0.55, 1), #from 3P0 to #3P1, the state weight for lower level
('[CII] 158',2.15, 2)], #from 1P1/2 to 1P3/2, the state weight for lower level
dtype={'names' :["Line","collision_strength", 'stat_weight'],
'formats' :['|S10', 'float64', 'float64']})
nii_q=collision_rate_coeff(8000.,collision_strengths_weights['stat_weight'][0],collision_strengths_weights["collision_strength"][0],line_frequencies[0])
oiii_q=collision_rate_coeff(8000.,collision_strengths_weights['stat_weight'][1],collision_strengths_weights["collision_strength"][1],line_frequencies[1])
cii_q=collision_rate_coeff(8000.,collision_strengths_weights['stat_weight'][2],collision_strengths_weights["collision_strength"][2],line_frequencies[2])
collision_rates =np.array([('[NII] 122', nii_q, 'cm^3/s'),
('[OIII] 88', oiii_q, 'cm^3/s') ,
('[CII] 158', cii_q, 'cm^3/s')],
dtype={'names' :["Line","collision_rate","rate_unit"],
'formats':['|S10', 'float64', '|S10']})
def nii_ff_metallicity(o_h_ratio,ion_fraction,o_n_ratio):
fsl = fsl_luminosity_lowDen(line_frequencies[0],1,ion_fraction,o_h_ratio/o_n_ratio,collision_rates["collision_rate"][0],1)
ff = free_free_specific_lum(radio_frequency,8000,gaunt_ff_5Ghz,1,1)
return fsl/ff
def oiii_ff_metallicity(o_h_ratio,ion_fraction,o_o_ratio):
fsl = fsl_luminosity_lowDen(line_frequencies[1],1,ion_fraction,o_h_ratio/o_o_ratio,collision_rates["collision_rate"][1],1)
ff = free_free_specific_lum(radio_frequency,8000,gaunt_ff_5Ghz,1,1)
return fsl/ff
def cii_ff_metallicity(o_h_ratio,ion_fraction,o_c_ratio):
fsl = fsl_luminosity_lowDen(line_frequencies[2],1,ion_fraction,o_h_ratio/o_c_ratio,collision_rates["collision_rate"][2],1)
ff = free_free_specific_lum(radio_frequency,8000,gaunt_ff_5Ghz,1,1)
return fsl/ff
# -
fsl_luminosity_lowDen(line_frequencies[0],1,1,1,collision_rates["collision_rate"][0],1)
(6.8e-38*8000**-0.5*4.86)
free_free_specific_lum(radio_frequency,8000,gaunt_ff_5Ghz,1,1)
fsl_luminosity_lowDen(line_frequencies[0],1,1,1,collision_rates["collision_rate"][0],1)/(6.8e-38*1**2*8000**(-0.5)*np.exp(-const.h.value*5e9/(const.k_B.value*8000)))
gaunt_ff_5Ghz
nii_ff_metallicity(1.,1.,1.)*.1*.13/10**15
oiii_ff_metallicity(1.,1.,1.)*.1*1
cii_ff_metallicity(1.,1.,1.)*.1*.67
#savage & semback 1996 HII nebular abundances
n_h = 7.76E-05
o_h = 5.89E-04
c_h = 3.98E-04
o_n_ratio = o_h/n_h
o_o_ratio = 1
o_c_ratio = o_h/c_h
nii_fraction = 0.1
oiii_fraction = 0.1
cii_fraction = 0.1
1/o_n_ratio
1/o_c_ratio
# ## Define Some Utility Functions
# +
def make_masked_quantities(data_table,convert_unit=False,desired_unit=None):
#print unit
if convert_unit == False:
values = (data_table.data.data.astype(np.float)*data_table.unit)
else:
values = (data_table.data.data.astype(np.float)*data_table.unit).to(desired_unit)
mask = ma.copy(data_table.mask)
return ma.array(values,mask=mask)
def linear_to_log_errors(data_table,linear_errors):
upper_bound = data_table+linear_errors
lower_bound = data_table-linear_errors
log_data = np.log10(data_table)
log_upper_bound = np.log10(upper_bound)
log_lower_bound = np.log10(lower_bound)
positive_error = log_upper_bound-log_data
negative_error = log_data-log_lower_bound
return positive_error,negative_error
def linear_to_log_errors_log_data(log_data_table,linear_errors):
upper_bound = 10**log_data_table+linear_errors
lower_bound = 10**log_data_table-linear_errors
log_data = log_data_table
log_upper_bound = np.log10(upper_bound)
log_lower_bound = np.log10(lower_bound)
positive_error = log_upper_bound-log_data
negative_error = log_data-log_lower_bound
return positive_error,negative_error
# -
# ## Define plotting variables
# +
#cii = make_masked_quantities(data_table['CII'],True,'W m^-2')
cii_data = make_masked_quantities(data_table['CII'],True,'W m^-2')
ERRcii = make_masked_quantities(data_table['errCII'],True,'W m^-2')
ULcii = np.zeros(len(cii_data))
ULcii[np.where(data_table['ulCII']==1)[0]]=True
#nii = make_masked_quantities(data_table['N122'],True,'W m^-2')
nii_data = make_masked_quantities(data_table['N122'],True,'W m^-2')
ERRnii = make_masked_quantities(data_table['errN122'],True,'W m^-2')
ULnii = np.zeros(len(nii_data))
ULnii[np.where(data_table['ulN122']==1)[0]]=True
#oiii= make_masked_quantities(data_table['OIII88'],True,'W m^-2')
oiii_data= make_masked_quantities(data_table['OIII88'],True,'W m^-2')
ERRoiii = make_masked_quantities(data_table['errOIII88'],True,'W m^-2')
ULoiii = np.zeros(len(oiii_data))
ULoiii[np.where(data_table['ulOIII88']==1)[0]]=True
ff = make_masked_quantities(data_table['FreeFree'])*ff_fraction
ERRff = make_masked_quantities(data_table['errFreeFree'])*ff_fraction
#For Free Free values without an uncertainty I assume an error 30% (i.e they are ~3 sigma detections)
ERRff[np.where(data_table['errFreeFree'].data.data==0.)[0]]=0.3*ff[np.where(data_table['errFreeFree'].data.data==0.)[0]].data*ff_fraction
ULff = np.zeros(len(ff))
ULff[np.where(data_table['ulFreeFree']==1)[0]]=True
metallicity = ma.array(np.copy(data_table['Metallicity'].data.data),mask=ma.copy(data_table['Metallicity'].data.mask))
ERRmetallicity = data_table['errMetallicity'].data.data
#For the metallicity values without a given uncertainty I give a 0.1 dex error
ERRmetallicity[np.where(data_table['errMetallicity'].data.data<0.1)[0]]=0.1
# -
# ## Plot Data -- Line, FF, Metallicity -- Individual Plots
# The code below makes plots of NII/FF, OIII/FF and CII/FF of the local galaxies. The points are color coded by the OIII/NII ratio. The free-free values are taken to be the 5GHz radio continuum assuming that the free-free is 30% at that point. Also plotted are the expected. C,N to O ratios are taken from Savage & Sembach b. The NII, CII and OIII ionization fractions are all assumed to be 0.1
# +
fig, axes = plt.subplots(nrows=1,ncols=3,figsize=(9,4.375),dpi=150)
fig.subplots_adjust(wspace=0.001)
logOH_12_range = np.arange(7.0,10,.1)
linearOH = 10**(logOH_12_range-12)
#----------------NII/FF versus Metalicity ------------------------------------#
axes[0].set_ylabel(r'$log(F_{FSL}/S_{5\: GHz}^{ff})\quad [Hz]$')
#axes[0].set_ylabel(r'$log(F_{FSL}/S_{\nu}^{radio})$')
#axes[0].set_xlabel(r'$12+log[O/H]$')
ydata = ma.array(np.log10((nii_data/ff).data.value*10.0**29),mask=ma.copy((nii_data/ff).mask))
xdata = metallicity
zdata = ma.array(np.log10((oiii_data/nii_data).data.value),mask=ma.copy((oiii_data/nii_data).mask))
#cmap = plt.cm.jet
#norm = colors.Normalize(zdata.min(),zdata.max())
#ec_colors = plt.cm.jet(norm(zdata))
yerr_linear = np.sqrt((ERRnii/nii_data).data.value**2+(ERRff/ff).data.value**2)*(nii_data/ff).data.value*10.0**29
yerr_log_pos,yerr_log_neg=linear_to_log_errors_log_data(ydata,yerr_linear)
yuplims = ULnii
#yerr_log_pos.mask = yuplims
#yerr_log_neg.mask = yuplims
where_UL=np.where(yuplims==1.0)[0]
yerr_log_pos[where_UL]=0
yerr=np.vstack([yerr_log_neg,yerr_log_pos])
#yerr.mask=yuplims
xerr = ERRmetallicity
axes[0].scatter(xdata,ydata,s=40,c=zdata,cmap='jet',alpha=0.7,zorder=1.0,linewidth=0.5)
axes[0].errorbar(xdata,ydata,xerr=xerr,yerr=yerr,uplims=yuplims,
ecolor='k',elinewidth=1,capthick=1,fmt='none',zorder=-1.0,alpha=0.5)
axes[0].plot(logOH_12_range,np.log10(nii_ff_metallicity(linearOH,nii_fraction,o_n_ratio)),color='black',linewidth=2,linestyle='--')
axes[0].annotate(r'$\frac{F_{[NII]}}{S_{5\: GHz}^{ff}} = $',
(.24,.67), xycoords='axes fraction', ha='right',va='center',fontsize=10)
axes[0].annotate(r'$6.80\times10^{15}\left( \frac{N{+}/N}{0.1} \right) \left( \frac{N/O}{0.13} \right) \left( \frac{O}{H} \right)$',
(.9,.75), xycoords='axes fraction', ha='right',va='center',fontsize=10)
axes[0].annotate(r"$[NII]\: 122\: \mu m$",(.05,.9),xycoords='axes fraction', ha='left',va='center',fontsize=14)
#----------------OIII/FF versus Metalicity ------------------------------------#
#axes[1].set_ylabel(r'$log([OIII]/S_{\nu}^{radio})$')
axes[1].set_xlabel(r'$12+log[O/H]$')
ydata = ma.array(np.log10((oiii_data/ff).data.value*10.0**29),mask=ma.copy((nii_data/ff).mask))
xdata = metallicity
zdata = ma.array(np.log10((oiii_data/nii_data).data.value),mask=ma.copy((oiii_data/nii_data).mask))
yerr_linear = np.sqrt((ERRoiii/oiii_data).data.value**2+(ERRff/ff).data.value**2)*(oiii_data/ff).data.value*10.0**29
yerr_log_pos,yerr_log_neg=linear_to_log_errors_log_data(ydata,yerr_linear)
yerr_log_pos.mask = ydata.mask
yerr_log_neg.mask = ydata.mask
yuplims = ULoiii
xerr = ERRmetallicity
axes[1].scatter(xdata,ydata,s=40,c=zdata,cmap='jet',alpha=0.7,zorder=1.0,linewidth=0.5)
axes[1].errorbar(xdata,ydata,xerr=xerr,yerr=[yerr_log_neg,yerr_log_pos],uplims=yuplims,
ecolor='k',elinewidth=1,capthick=1,fmt='none',zorder=-1.0,alpha=0.5)
axes[1].plot(logOH_12_range,np.log10(oiii_ff_metallicity(linearOH,oiii_fraction,o_o_ratio)),color='black',linewidth=2,linestyle='--')
axes[1].annotate(r'$\frac{F_{[OIII]}}{S_{5\: GHz}^{ff}} = $',
(.02,.45), xycoords='axes fraction', ha='left',va='center',fontsize=10)
axes[1].annotate(r'$1.60\times10^{17}\left( \frac{O{+}{+}/O}{0.1} \right) \left( \frac{O}{H} \right)$',
(.12,.3), xycoords='axes fraction', ha='left',va='bottom',fontsize=10)
axes[1].annotate(r"$[OIII]\: 88\: \mu m$",(.05,.15),xycoords='axes fraction', ha='left',va='bottom',fontsize=14)
#----------------CII/FF versus Metalicity ------------------------------------#
#axes[2].set_ylabel(r'$log([CII]/S_{\nu}^{radio})$')
#axes[2].set_xlabel(r'$12+log[O/H]$')
ydata = ma.array(np.log10((cii_data/ff).data.value*10.0**29),mask=ma.copy((nii_data/ff).mask))
xdata = metallicity
zdata = ma.array(np.log10((oiii_data/nii_data).data.value),mask=ma.copy((oiii_data/nii_data).mask))
yerr_linear = np.sqrt((ERRcii/cii_data).data.value**2+(ERRff/ff).data.value**2)*(cii_data/ff).data.value*10.0**29
yerr_log_pos,yerr_log_neg=linear_to_log_errors_log_data(ydata,yerr_linear)
yerr_log_pos.mask = ydata.mask
yerr_log_neg.mask = ydata.mask
yuplims = ULcii
xerr = ERRmetallicity
cii_plot=axes[2].scatter(xdata,ydata,s=40,c=zdata,cmap='jet',alpha=0.7,zorder=1.0,linewidth=0.5)
cii_error=axes[2].errorbar(xdata,ydata,xerr=xerr,yerr=[yerr_log_neg,yerr_log_pos],uplims=yuplims,
ecolor='k',elinewidth=1,capthick=1,fmt='none',zorder=-1.0,alpha=0.5)
axes[2].plot(logOH_12_range,np.log10(cii_ff_metallicity(linearOH,cii_fraction,o_c_ratio)),color='black',linewidth=2,linestyle='--')
axes[2].annotate(r'$\frac{F_{[CII]}}{S_{5\: GHz}^{ff}} = $',
(.24,.52), xycoords='axes fraction', ha='right',va='center',fontsize=10)
axes[2].annotate(r'$1.16\times10^{17}\left( \frac{C{+}/C}{0.1} \right) \left( \frac{C/O}{0.67} \right) \left( \frac{O}{H} \right)$',
(.95,.45), xycoords='axes fraction', ha='right',va='center',fontsize=10)
axes[2].annotate(r"$[CII]\: 158\: \mu m$",(.95,.3),xycoords='axes fraction', ha='right',va='bottom',fontsize=14)
#cb = plt.colorbar(ax=[ax1,ax2,ax3],orientation="horizontal",fraction=0.04,pad=0.07,
# anchor)
CBposition=fig.add_axes([0.67,0.2,0.2,0.02])
cb = fig.colorbar(cii_plot,cax=CBposition, ticks=[0.25,.75,1.25,1.75,2.25],orientation="horizontal")
cbax = cb.ax
cbax.text(0.0,1.5,r'$log([OIII]/[NII])$',horizontalalignment='left')
#cb.set_label(r'$[OIII]/[NII]$')
xticklabels = axes[1].get_yticklabels() + axes[2].get_yticklabels()
plt.setp(xticklabels, visible=False)
for ax in axes.flat:
ax.set_ylim([11.0,14.0])
ax.set_xlim([7.5,9.5])
ax.set_yticks([11,11.5,12.0,12.5,13.0,13.5,14])
ax.set_xticks([8.0,8.5,9.0,9.5])
ax.grid(True,alpha=0.3)
# -
fig.savefig(os.path.join(outfolder,'png','nii_oii_cii_ff_OH_relation.png'),dpi=300,bbox_inches='tight',pad_inches=0.5)
fig.savefig(os.path.join(outfolder,'pdf','nii_oii_cii_ff_OH_relation.pdf'),dpi=300,bbox_inches='tight',pad_inches=0.5)
# ## Plot Data -- Line, FF, Metallicity -- Individual Plots (cloudy abundances)
# The code below makes plots of NII/FF, OIII/FF and CII/FF of the local galaxies. The points are color coded by the OIII/NII ratio. The free-free values are taken to be the 5GHz radio continuum assuming that the free-free is 30% at that point. Also plotted are the expected. N to O ratios are taken from CLOUDY while C to O is from SAVAGE & SEMBACH
# +
cloudy_o_solar_abundance = 8.69
cloudy_n_solar_abundance = 7.93
#abundance = metallicity - cloudy_o_solar_abundance
abundance_data = metallicity - cloudy_o_solar_abundance
cloudy_o_n_ratio = 10**(cloudy_o_solar_abundance-12)/10**(cloudy_n_solar_abundance-12)
# +
fig, axes = plt.subplots(nrows=1,ncols=3,figsize=(9,4.375),dpi=150)
fig.subplots_adjust(wspace=0.001)
logOH_12_range = np.arange(7.0,10,.1)
linearOH = 10**(logOH_12_range-12)
#----------------NII/FF versus Metalicity ------------------------------------#
axes[0].set_ylabel(r'$log(F_{FSL}/S_{5\: GHz}^{ff})\quad [Hz]$')
#axes[0].set_ylabel(r'$log(F_{FSL}/S_{\nu}^{radio})$')
#axes[0].set_xlabel(r'$12+log[O/H]$')
ydata = ma.array(np.log10((nii_data/ff).data.value*10.0**29),mask=ma.copy((nii_data/ff).mask))
xdata = metallicity
zdata = ma.array(np.log10((oiii_data/nii_data).data.value),mask=ma.copy((oiii_data/nii_data).mask))
#cmap = plt.cm.jet
#norm = colors.Normalize(zdata.min(),zdata.max())
#ec_colors = plt.cm.jet(norm(zdata))
yerr_linear = np.sqrt((ERRnii/nii_data).data.value**2+(ERRff/ff).data.value**2)*(nii_data/ff).data.value*10.0**29
yerr_log_pos,yerr_log_neg=linear_to_log_errors_log_data(ydata,yerr_linear)
yuplims = ULnii
#yerr_log_pos.mask = yuplims
#yerr_log_neg.mask = yuplims
where_UL=np.where(yuplims==1.0)[0]
yerr_log_pos[where_UL]=0
yerr=np.vstack([yerr_log_neg,yerr_log_pos])
#yerr.mask=yuplims
xerr = ERRmetallicity
axes[0].scatter(xdata,ydata,s=40,c=zdata,cmap='jet',alpha=0.7,zorder=1.0,linewidth=0.5)
axes[0].errorbar(xdata,ydata,xerr=xerr,yerr=yerr,uplims=yuplims,
ecolor='k',elinewidth=1,capthick=1,fmt='none',zorder=-1.0,alpha=0.5)
axes[0].plot(logOH_12_range,np.log10(nii_ff_metallicity(linearOH,nii_fraction,o_n_ratio)),color='black',linewidth=2,linestyle='--')
axes[0].annotate(r"$[NII]\: 122\: \mu m$",(.05,.8),xycoords='axes fraction', ha='left',va='center',fontsize=14)
#----------------OIII/FF versus Metalicity ------------------------------------#
#axes[1].set_ylabel(r'$log([OIII]/S_{\nu}^{radio})$')
axes[1].set_xlabel(r'$12+log[O/H]$')
ydata = ma.array(np.log10((oiii_data/ff).data.value*10.0**29),mask=ma.copy((nii_data/ff).mask))
xdata = metallicity
zdata = ma.array(np.log10((oiii_data/nii_data).data.value),mask=ma.copy((oiii_data/nii_data).mask))
yerr_linear = np.sqrt((ERRoiii/oiii_data).data.value**2+(ERRff/ff).data.value**2)*(oiii_data/ff).data.value*10.0**29
yerr_log_pos,yerr_log_neg=linear_to_log_errors_log_data(ydata,yerr_linear)
yerr_log_pos.mask = ydata.mask
yerr_log_neg.mask = ydata.mask
yuplims = ULoiii
xerr = ERRmetallicity
axes[1].scatter(xdata,ydata,s=40,c=zdata,cmap='jet',alpha=0.7,zorder=1.0,linewidth=0.5)
axes[1].errorbar(xdata,ydata,xerr=xerr,yerr=[yerr_log_neg,yerr_log_pos],uplims=yuplims,
ecolor='k',elinewidth=1,capthick=1,fmt='none',zorder=-1.0,alpha=0.5)
axes[1].plot(logOH_12_range,np.log10(oiii_ff_metallicity(linearOH,oiii_fraction,o_o_ratio)),color='black',linewidth=2,linestyle='--')
axes[1].annotate(r"$[OIII]\: 88\: \mu m$",(.5,.3),xycoords='axes fraction', ha='right',va='bottom',fontsize=14)
#----------------CII/FF versus Metalicity ------------------------------------#
#axes[2].set_ylabel(r'$log([CII]/S_{\nu}^{radio})$')
#axes[2].set_xlabel(r'$12+log[O/H]$')
ydata = ma.array(np.log10((cii_data/ff).data.value*10.0**29),mask=ma.copy((nii_data/ff).mask))
xdata = metallicity
zdata = ma.array(np.log10((oiii_data/nii_data).data.value),mask=ma.copy((oiii_data/nii_data).mask))
yerr_linear = np.sqrt((ERRcii/cii_data).data.value**2+(ERRff/ff).data.value**2)*(cii_data/ff).data.value*10.0**29
yerr_log_pos,yerr_log_neg=linear_to_log_errors_log_data(ydata,yerr_linear)
yerr_log_pos.mask = ydata.mask
yerr_log_neg.mask = ydata.mask
yuplims = ULcii
xerr = ERRmetallicity
cii_plot=axes[2].scatter(xdata,ydata,s=40,c=zdata,cmap='jet',alpha=0.7,zorder=1.0,linewidth=0.5)
cii_error=axes[2].errorbar(xdata,ydata,xerr=xerr,yerr=[yerr_log_neg,yerr_log_pos],uplims=yuplims,
ecolor='k',elinewidth=1,capthick=1,fmt='none',zorder=-1.0,alpha=0.5)
axes[2].plot(logOH_12_range,np.log10(cii_ff_metallicity(linearOH,cii_fraction,o_c_ratio)),color='black',linewidth=2,linestyle='--')
axes[2].annotate(r"$[CII]\: 158\: \mu m$",(.95,.3),xycoords='axes fraction', ha='right',va='bottom',fontsize=14)
#cb = plt.colorbar(ax=[ax1,ax2,ax3],orientation="horizontal",fraction=0.04,pad=0.07,
# anchor)
CBposition=fig.add_axes([0.67,0.2,0.2,0.02])
cb = fig.colorbar(cii_plot,cax=CBposition, ticks=[0.25,.75,1.25,1.75,2.25],orientation="horizontal")
cbax = cb.ax
cbax.text(0.0,1.5,r'$log([OIII]/[NII])$',horizontalalignment='left')
#cb.set_label(r'$[OIII]/[NII]$')
xticklabels = axes[1].get_yticklabels() + axes[2].get_yticklabels()
plt.setp(xticklabels, visible=False)
for ax in axes.flat:
ax.set_ylim([11.0,14.0])
ax.set_xlim([7.5,9.5])
ax.set_yticks([11,11.5,12.0,12.5,13.0,13.5,14])
ax.set_xticks([8.0,8.5,9.0,9.5])
ax.grid(True,alpha=0.3)
# -
fig.savefig(os.path.join(outfolder,'png','nii_oii_cii_ff_OH_relation_noEqn.png'),dpi=300,bbox_inches='tight',pad_inches=0.5)
fig.savefig(os.path.join(outfolder,'pdf','nii_oii_cii_ff_OH_relation_noEqn.pdf'),dpi=300,bbox_inches='tight',pad_inches=0.5)
# ## Plot Data in a Square
# +
fig, axes = plt.subplots(nrows=2,ncols=2,figsize=(6,8.75),dpi=150)
fig.subplots_adjust(hspace=.175,wspace=.375)
#----------------NII/FF versus Metalicity ------------------------------------#
axes[0,0].set_ylabel(r'$log([NII]/S_{5\: GHz}^{ff})$')
axes[0,0].set_xlabel(r'$12+log[O/H]$')
ydata = ma.array(np.log10((nii_data/ff).data.value*10.0**29),mask=ma.copy((nii_data/ff).mask))
xdata = metallicity
zdata = ma.array(np.log10((oiii_data/nii_data).data.value),mask=ma.copy((oiii_data/nii_data).mask))
#cmap = plt.cm.jet
#norm = colors.Normalize(zdata.min(),zdata.max())
#ec_colors = plt.cm.jet(norm(zdata))
yerr_linear = np.sqrt((ERRnii/nii_data).data.value**2+(ERRff/ff).data.value**2)*(nii_data/ff).data.value*10.0**29
yerr_log_pos,yerr_log_neg=linear_to_log_errors_log_data(ydata,yerr_linear)
yuplims = ULnii
#yerr_log_pos.mask = yuplims
#yerr_log_neg.mask = yuplims
where_UL=np.where(yuplims==1.0)[0]
yerr_log_pos[where_UL]=0
yerr=np.vstack([yerr_log_neg,yerr_log_pos])
#yerr.mask=yuplims
xerr = ERRmetallicity
axes[0,0].scatter(xdata,ydata,s=40,c=zdata,cmap='jet',alpha=0.7,zorder=1.0,linewidth=0.5)
axes[0,0].errorbar(xdata,ydata,xerr=xerr,yerr=yerr,uplims=yuplims,
ecolor='k',elinewidth=1,capthick=1,fmt='none',zorder=-1.0,alpha=0.5)
#----------------OIII/FF versus Metalicity ------------------------------------#
axes[0,1].set_ylabel(r'$log([OIII]/S_{5\: GHz}^{ff})$')
axes[0,1].set_xlabel(r'$12+log[O/H]$')
ydata = ma.array(np.log10((oiii_data/ff).data.value*10.0**29),mask=ma.copy((nii_data/ff).mask))
xdata = metallicity
zdata = ma.array(np.log10((oiii_data/nii_data).data.value),mask=ma.copy((oiii_data/nii_data).mask))
yerr_linear = np.sqrt((ERRoiii/oiii_data).data.value**2+(ERRff/ff).data.value**2)*(oiii_data/ff).data.value*10.0**29
yerr_log_pos,yerr_log_neg=linear_to_log_errors_log_data(ydata,yerr_linear)
yerr_log_pos.mask = ydata.mask
yerr_log_neg.mask = ydata.mask
yuplims = ULoiii
xerr = ERRmetallicity
axes[0,1].scatter(xdata,ydata,s=40,c=zdata,cmap='jet',alpha=0.7,zorder=1.0,linewidth=0.5)
axes[0,1].errorbar(xdata,ydata,xerr=xerr,yerr=[yerr_log_neg,yerr_log_pos],uplims=yuplims,
ecolor='k',elinewidth=1,capthick=1,fmt='none',zorder=-1.0,alpha=0.5)
#----------------CII/FF versus Metalicity ------------------------------------#
axes[1,0].set_ylabel(r'$log([CII]/S_{5\: GHz}^{ff})$')
axes[1,0].set_xlabel(r'$12+log[O/H]$')
ydata = ma.array(np.log10((cii_data/ff).data.value*10.0**29),mask=ma.copy((nii_data/ff).mask))
xdata = metallicity
zdata = ma.array(np.log10((oiii_data/nii_data).data.value),mask=ma.copy((oiii_data/nii_data).mask))
yerr_linear = np.sqrt((ERRcii/cii_data).data.value**2+(ERRff/ff).data.value**2)*(cii_data/ff).data.value*10.0**29
yerr_log_pos,yerr_log_neg=linear_to_log_errors_log_data(ydata,yerr_linear)
yerr_log_pos.mask = ydata.mask
yerr_log_neg.mask = ydata.mask
yuplims = ULcii
xerr = ERRmetallicity
cii_plot=axes[1,0].scatter(xdata,ydata,s=40,c=zdata,cmap='jet',alpha=0.7,zorder=1.0,linewidth=0.5)
cii_error=axes[1,0].errorbar(xdata,ydata,xerr=xerr,yerr=[yerr_log_neg,yerr_log_pos],uplims=yuplims,
ecolor='k',elinewidth=1,capthick=1,fmt='none',zorder=-1.0,alpha=0.5)
#cb = plt.colorbar(ax=[ax1,ax2,ax3],orientation="horizontal",fraction=0.04,pad=0.07,
# anchor)
CBposition=fig.add_axes([0.15,0.16,0.28,0.0125])
cb = fig.colorbar(cii_plot,cax=CBposition, ticks=[0.25,.75,1.25,1.75,2.25],orientation="horizontal")
cbax = cb.ax
cbax.text(0.0,1.5,r'$log([OIII]/[NII])$',horizontalalignment='left')
#cb.set_label(r'$[OIII]/[NII]$')
for ax in axes.flat:
ax.set_ylim([11,14])
ax.set_xlim([7.5,9.5])
ax.set_yticks([11,11.5,12.0,12.5,13.0,13.5,14.0])
ax.set_xticks([8.0,8.5,9.0,9.5])
ax.grid(True,alpha=0.3)
# -
# ### Save Figures
fig.savefig(os.path.join(outfolder,'png','nii_oii_cii_ff_OH_SQUARE.png'),dpi=300,bbox_inches='tight',pad_inches=0.4)
fig.savefig(os.path.join(outfolder,'pdf','nii_oii_cii_ff_OH_SQUARE.pdf'),dpi=300,bbox_inches='tight',pad_inches=0.3)
| PHY460/nii_ff_analysis_201707 .ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
# -
import numpy as np
import pandas as pd
from pathlib import Path
import torch
import random
import os
import torch.nn as nn
from torch.utils.data import Dataset, DataLoader
from tqdm.notebook import tqdm
from sklearn.model_selection import KFold
from sklearn.metrics import mean_absolute_error
import matplotlib.pyplot as plt
import torch.nn.functional as F
from lib import common
pd.options.mode.chained_assignment = None
# ### Path
path = Path('/kaggle/osic_pulmonary')
assert path.exists()
model_path = Path('/kaggle/osic_pulmonary/model')
if os.path.isdir(model_path) == False:
os.makedirs(model_path)
assert model_path.exists()
# ### Read Data
train_df, test_df, submission_df = common.read_data(path)
# #### Feature generation
len(train_df)
submission_df = common.prepare_submission(submission_df, test_df)
submission_df[((submission_df['Patient'] == 'ID00419637202311204720264') & (submission_df['Weeks'] == 6))].head(25)
def adapt_percent_in_submission():
previous_match = None
for i, r in submission_df.iterrows():
in_training = train_df[(train_df['Patient'] == r['Patient']) & (train_df['Weeks'] == r['Weeks'])]
if(len(in_training) > 0):
previous_match = in_training['Percent'].item()
submission_df.iloc[i, submission_df.columns.get_loc('Percent')] = previous_match
elif previous_match is not None:
submission_df.iloc[i, submission_df.columns.get_loc('Percent')] = previous_match
adapt_percent_in_submission()
test_df
test_df[test_df['Patient'] == 'ID00419637202311204720264']
train_df[train_df['Patient'] == 'ID00419637202311204720264']
submission_df[submission_df['Patient'] == 'ID00419637202311204720264'].head(10)
# Adding missing values
train_df['WHERE'] = 'train'
test_df['WHERE'] = 'val'
submission_df['WHERE'] = 'test'
data = train_df.append([test_df, submission_df])
data['min_week'] = data['Weeks']
data.loc[data.WHERE=='test','min_week'] = np.nan
data['min_week'] = data.groupby('Patient')['min_week'].transform('min')
base = data.loc[data.Weeks == data.min_week]
base = base[['Patient','FVC', 'Percent']].copy()
base.columns = ['Patient','min_FVC', 'min_Percent']
base['nb'] = 1
base['nb'] = base.groupby('Patient')['nb'].transform('cumsum')
base = base[base.nb==1]
base
data = data.merge(base, on='Patient', how='left')
data['base_week'] = data['Weeks'] - data['min_week']
data['base_week'] = data['base_week']
del base
data[data['Patient'] == 'ID00421637202311550012437']
COLS = ['Sex','SmokingStatus'] #,'Age', 'Sex_SmokingStatus'
FE = []
for col in COLS:
for mod in data[col].unique():
FE.append(mod)
data[mod] = (data[col] == mod).astype(int)
data[data['Patient'] == 'ID00421637202311550012437']
np.mean(np.abs(data['Age'] - data['Age'].mean())), data['Age'].mad()
# +
def normalize(df:pd.DataFrame, cont_names, target_names):
"Compute the means and stds of `self.cont_names` columns to normalize them."
means, stds = {},{}
for n, t in zip(cont_names, target_names):
means[n], stds[n] = df[n].mean(), df[n].std()
# means[n], stds[n] = df[n].mean(), df[n].mad()
df[t] = (df[n]-means[n]) / (1e-7 + stds[n])
normalize(data, ['Age','min_FVC','base_week','Percent', 'min_Percent', 'min_week'], ['age','BASE','week','percent', 'min_percent', 'min_week'])
FE += ['age','week','BASE', 'percent']
# -
data['base_week'].min()
train_df = data.loc[data.WHERE=='train']
test_df = data.loc[data.WHERE=='val']
submission_df = data.loc[data.WHERE=='test']
del data
train_df.sort_values(['Patient', 'Weeks']).head(15)
X = train_df[FE]
X.head(15)
y = train_df['FVC']
y.shape
SEQ_LENGTH = 8
def create_input_sequences(input_data, target_data, tw, features=FE):
inout_features_seq = []
inout_target_seq = []
L = len(input_data)
for i in range(L-tw):
train_seq = input_data[features][i:i+tw].values
train_label = target_data[i+tw:i+tw+1].values
inout_features_seq.append(train_seq)
inout_target_seq.append(train_label)
return torch.tensor(np.array(inout_features_seq), dtype=torch.float32), torch.tensor(np.array(inout_target_seq), dtype=torch.float32)
train_features, train_target = create_input_sequences(train_df, train_df['FVC'], SEQ_LENGTH)
train_features.shape, train_target.shape
# #### Seed
# +
def seed_everything(seed=2020):
random.seed(seed)
os.environ['PYTHONHASHSEED'] = str(seed)
np.random.seed(seed)
torch.manual_seed(42)
torch.cuda.manual_seed(42)
seed_everything(42)
# -
# ### Create Dataset
class ArrayDataset(Dataset):
def __init__(self, x, y):
self.x, self.y = x, y
assert(len(self.x) == len(self.y))
def __len__(self):
return len(self.x)
def __getitem__(self, i):
return self.x[i], self.y[i]
def __repr__(self):
return f'x: {self.x.shape} y: {self.y.shape}'
def create_dl(df, target_data, batch_size=128, num_workers=10):
train_features, train_target = create_input_sequences(df, target_data, SEQ_LENGTH)
ds = ArrayDataset(train_features, train_target)
return DataLoader(ds, batch_size, shuffle=True, num_workers=num_workers)
sample_dl = create_dl(train_df, train_df['FVC'])
x_sample, y_sample = next(iter(sample_dl))
x_sample.shape, y_sample.shape
# ### Prepare neural network
# +
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
def move_to_dev(x, y):
x = x.to(device)
y = y.to(device)
return x, y
# +
C1, C2 = torch.tensor(70, dtype=torch.float32), torch.tensor(1000, dtype=torch.float32)
C1, C2 = move_to_dev(C1, C2)
q = torch.tensor([0.2, 0.50, 0.8]).float().to(device)
def score(y_true, y_pred):
sigma = y_pred[:, 2] - y_pred[:, 0]
fvc_pred = y_pred[:, 1]
#sigma_clip = sigma + C1
sigma_clip = torch.max(sigma, C1)
delta = torch.abs(y_true[:, 0] - fvc_pred)
delta = torch.min(delta, C2)
sq2 = torch.sqrt(torch.tensor(2.))
metric = (delta / sigma_clip)*sq2 + torch.log(sigma_clip* sq2)
return torch.mean(metric)
def qloss(y_true, y_pred):
# Pinball loss for multiple quantiles
e = y_true - y_pred
v = torch.max(q*e, (q-1)*e)
return torch.mean(v)
def mloss(_lambda):
def loss(y_true, y_pred):
y_true = y_true.unsqueeze(1)
return _lambda * qloss(y_true, y_pred) + (1 - _lambda)*score(y_true, y_pred)
return loss
# -
train_features.shape
# +
class OsicLSTM(nn.Module):
def __init__(self, input_size=train_features.shape[2], rnn_size = train_features.shape[1], hidden_layer_size=50, output_size=3, nh2=100):
super().__init__()
self.hidden_layer_size = hidden_layer_size
self.rnn_size = rnn_size
self.lstm1 = nn.LSTM(input_size, hidden_layer_size, bidirectional=True)
nh1 = hidden_layer_size * 2 * rnn_size // 2
self.l1 = nn.Linear(hidden_layer_size * 2 * rnn_size, nh1)
self.l1_bn = nn.BatchNorm1d(nh1, momentum=0.1)
self.l2 = nn.Linear(nh1, nh2)
self.p1 = nn.Linear(nh2, output_size)
self.p2 = nn.Linear(nh2, output_size)
self.relu = nn.ReLU()
self.create_hidden()
def create_hidden(self):
# Expected hidden[0] size (2, this.rnn_size, 50)
self.hidden_cell_a = torch.zeros(2, self.rnn_size, self.hidden_layer_size).to(device)
self.hidden_cell_b = torch.zeros(2, self.rnn_size, self.hidden_layer_size).to(device)
def forward(self, input_seq):
lstm_out, hidden = self.lstm1(input_seq, (self.hidden_cell_a, self.hidden_cell_b))
self.hidden_cell_a = hidden[0].detach()
self.hidden_cell_b = hidden[1].detach()
x = self.relu(self.l1(lstm_out.reshape([lstm_out.shape[0], -1])))
x = self.l1_bn(x)
x = self.relu(self.l2(x))
p1 = self.p1(x)
p2 = self.relu(self.p2(x))
preds = p1 + torch.cumsum(p2, axis=1)
return preds
class OsicModel(torch.nn.Module):
def __init__(self, ni, nh1, nh2):
super(OsicModel, self).__init__()
self.l1 = nn.Linear(ni, nh1)
self.l1_bn = nn.BatchNorm1d(nh1, momentum=0.1)
self.l2 = nn.Linear(nh1, nh2)
self.relu = nn.ReLU()
self.p1 = nn.Linear(nh2, 3)
self.p2 = nn.Linear(nh2, 3)
def forward(self, x):
x = self.relu(self.l1(x))
x = self.l1_bn(x)
x = self.relu(self.l2(x))
p1 = self.p1(x)
p2 = self.relu(self.p2(x))
preds = p1 + torch.cumsum(p2, axis=1)
return preds
# -
def create_model():
model = OsicLSTM()
model = model.to(device)
return model
sample_model = create_model()
sample_model
x_sample.shape, train_features.shape[1]
x_sample, y_sample = move_to_dev(x_sample, y_sample)
sample_model.create_hidden()
x, hidden = sample_model.lstm1(x_sample, (sample_model.hidden_cell_a, sample_model.hidden_cell_b))
hidden = (hidden[0].detach(), hidden[1].detach())
x.shape, x.reshape([x.shape[0], -1]).shape
x = sample_model.l1(x.reshape([x.shape[0], -1]))
x.shape, hidden[0].shape, hidden[1].shape
criterion=mloss(0.8)
# Test model
y_sample, x_sample = move_to_dev(y_sample, x_sample)
output = sample_model(x_sample)
output.shape, y_sample.shape
# loss = criterion(y_sample, output)
# loss.backward()
# #### Training functions
LR=1e-3
def get_lr(optimizer):
for param_group in optimizer.param_groups:
return param_group['lr']
def eval_loop(valid_dl, model):
with torch.no_grad():
model.eval()
total_eval_loss = 0
total_eval_score = 0
for x, y in valid_dl:
x, y = move_to_dev(x, y)
output = model(x)
loss = criterion(y, output)
total_eval_loss += loss.item()
total_eval_score += score(y.unsqueeze(1), output)
avg_val_loss = total_eval_loss / len(valid_dl)
avg_val_score = total_eval_score / len(valid_dl) * -1
return {
'avg_val_loss': avg_val_loss,
'avg_val_score': avg_val_score
}
def train_loop(epochs, train_dl, valid_dl, model, lr = LR, print_score=False, model_name='test'):
steps = len(train_dl) * epochs
optimizer = torch.optim.Adam(model.parameters(), lr=lr)
scheduler = torch.optim.lr_scheduler.OneCycleLR(optimizer, max_lr=lr, steps_per_epoch=len(train_dl), epochs=epochs)
avg_train_losses = []
avg_val_losses = []
avg_val_scores = []
lr = []
best_avg_val_score = -1000
for epoch in tqdm(range(epochs), total=epochs):
model.train()
total_train_loss = 0.0
for i, (x, y) in enumerate(train_dl):
x, y = move_to_dev(x, y)
model.zero_grad()
output = model(x)
loss = criterion(y, output)
total_train_loss += loss.item()
# Backward Pass and Optimization
loss.backward()
optimizer.step()
scheduler.step()
lr.append(get_lr(optimizer))
avg_train_loss = total_train_loss / len(train_dl)
avg_train_losses.append(avg_train_loss)
eval_res = eval_loop(valid_dl, model)
avg_val_loss = eval_res['avg_val_loss']
avg_val_score = eval_res['avg_val_score']
avg_val_losses.append(avg_val_loss)
avg_val_scores.append(avg_val_score.item())
if best_avg_val_score < avg_val_score:
best_avg_val_score = avg_val_score
# save best model
torch.save(model.state_dict(), model_path/f'best_model_simple_{model_name}.pt')
if print_score:
print(f'{epoch}: avg_val_score: {avg_val_score}')
return pd.DataFrame({'avg_train_losses': avg_train_losses, 'avg_val_losses': avg_val_losses, 'avg_val_scores': avg_val_scores}), pd.DataFrame({'lr': lr})
res_df, lr_df = train_loop(300, sample_dl, sample_dl, sample_model)
res_df[['avg_train_losses', 'avg_val_losses']].plot()
res_df[['avg_val_scores']].plot()
lr_df.plot()
res_df[['avg_val_scores']].max()
# #### Training
NFOLD = 5
kf = KFold(n_splits=NFOLD)
EPOCHS=100
def convert_to_tensor(df):
return torch.tensor(df.values, dtype=torch.float32).to(device)
submission_patients = submission_df['Patient']
submission_df['dummy_FVC'] = 0.0
submission_dl = create_dl(submission_df[FE], pd.Series(np.zeros(submission_df[FE].shape[0])))
x_sample, y_sample = next(iter(submission_dl))
x_sample.shape, y_sample.shape
pe = np.zeros((submission_df.shape[0], 3))
pred = np.zeros((train_df.shape[0], 3))
pred.shape
test_values = convert_to_tensor(submission_df[FE])
test_values.shape
def predict(features, model):
predict_features, _ = create_input_sequences(features, pd.Series(np.zeros(features[FE].shape[0])), SEQ_LENGTH)
predict_features = predict_features.to(device)
return model(predict_features).detach().cpu().numpy()
# +
# %%time
res_dfs = []
for cnt, (tr_idx, val_idx) in tqdm(enumerate(kf.split(X)), total=NFOLD):
X_train, y_train = X.loc[tr_idx], y[tr_idx]
X_valid, y_valid = X.loc[val_idx], y[val_idx]
print(f"FOLD {cnt}", X_train.shape, y_train.shape, X_valid.shape, y_valid.shape)
model = create_model()
train_dl = create_dl(X_train, y_train)
valid_dl = create_dl(X_valid, y_valid)
res_df, _ = train_loop(EPOCHS, train_dl, valid_dl, model, model_name=str(cnt), lr=LR)
res_dfs.append(res_df)
predicted = predict(X_valid, model)
print(len(val_idx), predicted.shape)
pred[val_idx] = predicted
# +
from matplotlib.pyplot import figure
def plot_results(cols=['avg_train_losses', 'avg_val_losses']):
nrows = len(res_dfs) // 2 + 1
ncols = 2
fig, axes = plt.subplots(nrows, ncols, figsize=(20, 10))
figure(num=None, figsize=(10, 6), dpi=80, facecolor='w', edgecolor='k')
for r in range(nrows):
for c in range(ncols):
index = r * 2 + c
if index < len(res_dfs):
res_dfs[r * 2 + c][cols].plot(ax=axes[r,c])
plot_results()
# -
plot_results(['avg_val_scores'])
print("Mean validation score", np.mean([res_dfs[i]['avg_val_scores'][len(res_dfs[0]) - 1] for i in range(NFOLD)]))
def load_best_model(i):
model_file = model_path/f'best_model_simple_{i}.pt'
model = create_model()
model.load_state_dict(torch.load(model_file))
model.to(device)
model.eval()
return model
# Using best models for prediction
pe = np.zeros((submission_df.shape[0], 3))
for i in range(NFOLD):
model = load_best_model(i)
pe += predict(test_values, model)
pe = pe / NFOLD
# #### Prediction
sigma_opt = mean_absolute_error(y, pred[:, 1])
unc = pred[:,2] - pred[:, 0]
sigma_mean = np.mean(unc)
sigma_opt, sigma_mean
submission_df['FVC1'] = pe[:,1]
submission_df['Confidence1'] = pe[:, 2] - pe[:, 0]
submission_df.head(15)
subm = submission_df[['Patient_Week','FVC','Confidence','FVC1','Confidence1']].copy()
subm.loc[~subm.FVC1.isnull()].shape, subm.shape
subm.loc[~subm.FVC1.isnull(),'FVC'] = subm.loc[~subm.FVC1.isnull(),'FVC1']
if sigma_mean<70:
subm['Confidence'] = sigma_opt
else:
subm.loc[~subm.FVC1.isnull(),'Confidence'] = subm.loc[~subm.FVC1.isnull(),'Confidence1']
subm.describe().T
# +
def replace_with_existing(df):
for i in range(len(df)):
patient_week_filter = subm['Patient_Week']==df.Patient[i]+'_'+str(df.Weeks[i])
subm.loc[patient_week_filter, 'FVC'] = df.FVC[i]
subm.loc[patient_week_filter, 'Confidence'] = 0.1
train_df = pd.read_csv(path/'train.csv', dtype = common.TRAIN_TYPES)
test_df = pd.read_csv(path/'test.csv', dtype = common.TRAIN_TYPES)
replace_with_existing(train_df)
replace_with_existing(test_df)
# -
subm[subm['Patient_Week'].str.find('ID00419637202311204720264') > -1].head(30)
subm[["Patient_Week","FVC","Confidence"]].to_csv("submission.csv", index=False)
submission_final_df = pd.read_csv('submission.csv')
submission_final_df[submission_final_df['Patient_Week'].str.find('ID00419637202311204720264') == 0]['FVC'].plot()
submission_final_df[submission_final_df['Patient_Week'].str.find('ID00421637202311550012437') == 0]['FVC'].plot()
submission_final_df[submission_final_df['Patient_Week'].str.find('ID00423637202312137826377') == 0]['FVC'].plot()
train_df[train_df['Patient'].str.find('ID00419637202311204720264') == 0][['Weeks', 'FVC']].plot(x='Weeks', y='FVC')
# !cat submission.csv
| nbs_gil/osic_pulmonary/10_training_pytorch_rnn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="fzH3L7yUkCA3"
#@title Set-up
import numpy as np
import matplotlib.pyplot as plt
from dataclasses import dataclass
# + id="-_kx3sSw7X4Z" cellView="form"
#@title Figure settings
# %config InlineBackend.figure_format = 'retina'
plt.style.use("https://raw.githubusercontent.com/rmaffulli/LIF_tutorial/main/figures.mplstyle")
# + [markdown] id="E4DTmOf1k40x"
# # Leaky Integrate and Fire (LIF) neurons
# During this tutorial you will learn how several classes of LIF work.
#
# We will start by building, step by step, a simple model of a leaky integrate-and-fire neuron. The model of the neuron is made of two key ingredients:
# 1. A leak conductance (hence the *leaky*): this part of the model ensures that the membrane potential of the neuron will tend to the equilibrium potential once the input stimulation ceases
# 2. A firing mechanism: point neuron models (i.e. models where the biochemical details of the firing mechanism are omitted) cannot generate spikes. We have to define a, biophysically plausible, firing mechanism and implement it.
#
# After this first section we will discuss the limitations of the LIF model and have a look at how to modify it to allow for more rich dynamics.
#
# All the models are partially incomplete. You will have to complete all the functions, according to the definition of the model. We will do part of the coding during the lecture. You should complete the remaining part at home. Completion of all the coding exercises is mandatory to pass the exam. You can submit the final results to me via email by sharing your version of the google colab.
#
# # References
# Below is a *non exaustive* list of textbooks that are a great starting point to explore the vastity of computational neuroscience!
# - <NAME>. An introductory course in computational neuroscience. MIT Press, 2018.
# - Gerstner, Wulfram, et al. Neuronal dynamics: From single neurons to networks and models of cognition. Cambridge University Press, 2014.
# - Churchland, <NAME>, and <NAME>. The computational brain. MIT press, 1994.
# + [markdown] id="3SiV5L8LsMdz"
# # The *Leaky Integrate* part
# A leaky integrator neuron takes the integral of an input, but gradually leaks a small amount of input over time. The state of the integrator, over time, is ruled by the following differential equation:
# $$ C_m\frac{dV_m(t)}{dt} = G_L(E_L-V_m(t)) + I_{in} $$
# Where: $C_m$ is the membrane capacitance, $V_m$ is the membrane potential, $G_L$ is the leak conductance, $E_L$ is the leak potential and $I_{in}$ is the input current to the neuron.
# In this section you will have to implement a neuron that behaves according to the aboce differential equation. Before jumping to the implementation take a look at the differential equation and try to address the following questions:
# - What happens once $E_L = V_m$?
# - What do you expect to happen if $I_{in} = 0$?
# - Can you give a hypothesis of what to expect from the membrane potential $V_m$ of a leaky integrator if $I_{in} > 0$ for a sustained period of time? Do you think this is a realistic description of a neuron? You will have the chance to test this hypothesis once implemented the leaky integrator
# + id="tHP7w18_k1iJ"
def leaky_int(t,i_in,model_constants):
### Inputs:
### t -> time (numpy array of doubles marking the time points where the simulation will be run)
### i_in -> input current (numpy array having the same dimension of t)
### model_constants -> structure containing all the constants for the model
### Output:
### v_m -> values of membrane potential over time (numpy array having the same dimension of t)
c_m = model_constants.c_m
g_l = model_constants.g_l
e_l = model_constants.e_l
dt = t[1]-t[0]
v_m = np.empty_like(t)
v_m[0] = e_l
if np.isscalar(i_in):
i_in_vec = np.full_like(t,i_in)
else:
i_in_vec = i_in
n_t = len(t)
for i in range(1,n_t):
v_m[i] = v_m[i-1] + dt*(g_l/c_m*(e_l-v_m[i-1]) + i_in_vec[i]/c_m)
return v_m
# + colab={"base_uri": "https://localhost:8080/", "height": 428} id="FbOQsXf0Dc7W" outputId="1ff64ad7-0d5f-476f-873e-4f05459d672e" cellView="form"
#@title Run leaky integrator with constant input current
dt = 0.0001 #@param {type:"number", default:0.1}
t_end = 0.5 #@param {type:"number"}
c_m = 100E-12 #@param {type:"number"}
g_l = 10E-9 #@param {type:"number"}
e_l = -70E-3 #@param {type:"number"}
i_in = 100E-12 #@param {type:"number"}
t = np.arange(0,t_end,dt)
@dataclass
class Constants:
c_m: float
g_l: float
e_l: float
model_constants = Constants(c_m,g_l,e_l)
v_m = leaky_int(t,i_in,model_constants)
_ = plt.plot(t,v_m*1000)
_ = plt.xlabel("t [s]")
_ = plt.ylabel("$V_m$ [mV]")
# + [markdown] id="Ynkbhh8XuYta"
# # Introduce spike generation: the Leaky Integrate and Fire
# So far we have an integrator that does not produce anything close to an action potential.
#
# - Can we simulate spikes using a LIF? Are we modelling all the mechanisms that are responsible of spike generation?
# - How can we overcome this?
#
# A simple way is to simply impose the membrane potential to drop to a specified value $V_{reset}$ once the membrane potential reaches a threshold $V_{th}$.
# + id="FIJkl3gQjZvq"
def leaky_int_and_fire(t,i_in,model_constants):
### Inputs:
### t -> time (numpy array of doubles marking the time points where the simulation will be run)
### i_in -> input current (numpy array having the same dimension of t)
### model_constants -> structure containing all the constants for the model
### Output:
### v_m -> values of membrane potential over time (numpy array having the same dimension of t)
### sp -> spike train
c_m = model_constants.c_m
g_l = model_constants.g_l
e_l = model_constants.e_l
v_th = model_constants.v_th
v_reset = model_constants.v_reset
dt = t[1]-t[0]
v_m = np.empty_like(t)
v_m[0] = e_l
if np.isscalar(i_in):
i_in_vec = np.full_like(t,i_in)
else:
i_in_vec = i_in
n_t = len(t)
sp = np.zeros(t.size)
for i in range(1,n_t):
v_m[i] = v_m[i-1] + dt*(g_l/c_m*(e_l-v_m[i-1]) + i_in_vec[i]/c_m)
# add firing
if v_m[i] >= v_th:
v_m[i] = v_reset
sp[i] = 1
return v_m, sp
# + colab={"base_uri": "https://localhost:8080/", "height": 428} id="7NqYv2GykHtW" outputId="ed2bba38-cf02-4ace-ce3b-344ca8c898e7" cellView="form"
#@title Run LIF with constant input current
dt = 0.0001 #@param {type:"number", default:0.1}
t_end = 0.5 #@param {type:"number"}
c_m = 100E-12 #@param {type:"number"}
g_l = 10E-9 #@param {type:"number"}
e_l = -70E-3 #@param {type:"number"}
i_in = 240E-12 #@param {type:"number"}
v_th = -50E-3 #@param {type:"number"}
v_reset = -80E-3 #@param {type:"number"}
t = np.arange(0,t_end,dt)
@dataclass
class Constants:
c_m: float
g_l: float
e_l: float
v_th: float
v_reset: float
model_constants = Constants(c_m,g_l,e_l,v_th, v_reset)
v_m, sp = leaky_int_and_fire(t,i_in,model_constants)
_ = plt.plot(t,v_m*1000,label="Membrane potential")
_ = plt.xlabel("t [s]")
_ = plt.ylabel("$V_m$ [mV]")
_ = plt.eventplot(t[sp == 1], color='orchid',linelengths=10, label="Spikes")
_ = plt.legend(loc='best')
# + [markdown] id="-mnaaGr7w2Jw"
# # Limitations of the LIF
# We have a spiking neuron but can we do better than that? There are many aspects relative to neuronal dynamics that are neglected with using a simple LIF as we have done so far.
# One issue is that the input current is integrated linearly, in a way that is not dependent from the state of the neuron. Another issue is that the membrane potential is reset every time there is a spike, losing all memory of the previous spikes.
#
# For the reasons above, a simple LIF cannot produce *adaption*, *bursting* and *inhibitory rebound* (see lecture notes), as all those depend on the spiking history of the neuron, which is erased at every spike for a LIF. A LIF cannot account also for the observation that the shape of postsynaptic potentials depends on the time elapsed since the last spike, nor can give us information about the spatial structure of inputs and their effects on the membrane potential (e.g. synapses that are located far away from the soma are expected to evoke a smaller postsynaptic response at the soma).
#
# But, if that's so bad, why bothering using a LIF? Despite its simplifications, it is very accurate in simulating spike generation in the soma, at least if we incorporate a way to account for adaptation and refractoriness.
# + [markdown] id="Sh4nYzN2y-eQ"
# # The refractory period
# As we have seen during the lecture, there are multiple biophysical events that happen inside the neuron before and after an action potential (charges accumulating on the membrane wall, opening/closing of membrane channels, entrance of ions in the cell, membrane pumps extracting the excess charges from the cell...). All those events have their own dynamics and cannot happen instantly and for a sustained amount of time. As a result a neuron cannot realistically fire with a very high frequency forever!
#
# The *refractory period* is a period of time, after an action potential, where a neuron cannot spike anymore. This is due to the fact that, during this time, the ion channels causing depolarization of the membrane cannot open anymore while they return to their baseline state.
#
# There are two ways to insert a refractory period in a LIF:
# 1. **Forced voltage clamp:** with this method we fix the membrane potential to its reset value for a set amount of time after an action potential. It's simple to implement and representative of real neurons at low firing rates. When firing rates are high, instead, the neuron will spend a greater proportion of its time with the membrane potential at its minimum. This means that the mean membrane potential will decrease when increasing the excitation current, which does not match experimental observations.
# 2. **Refractory conductance:** alternatively the refractory period can be simulated by addition of a large conductance that produces an hyperpolarizing potassium current. This will cause the membrane potential to decrease, making it harder for the neuron to generate an action potential, under the same excitation current. The refractory conductance increases at the time of each spike and decays between spike times with a short time constant $\tau_{ref}$.
# In this way the equations for the LIF neuron will become two, to be solved simultaneously:
# $$ C_m\frac{dV_m(t)}{dt} = G_L(E_L-V_m(t)) + I_{in} + G_{ref}(t)[E_k-V_m(t)]$$
#
# $$ \frac{dG_{ref}(t)}{dt} = \frac{-G_{ref}(t)}{\tau_{ref}};\; G_{ref} → G_{ref} + \Delta G_{ref} \;(after \:a \:spike)$$
#
# When the $G_{ref}$ term is much greater than the leak conductance, this term clamps the membrane potential at the Nernst potential for poassium ions $E_k$. With this approach we can omit the reset of the membrane potential after a spike as the increase in $G_{ref}$ causes the desired decrease of the membrane potential. Unlike the previous method, this has the advantage that the time spent at the reset value depends on the strength of the input current. The larger the currents, the more quickly they can overcome the hyperpolarizing current, which is also observed in real neurons.
#
# 3. **Voltage threshold increase:** The voltage threshold for producing a spike can be raised immediately following a spike and allowed to decay back with a short time constant.
# $$ \frac{dV_{th}}{dt} = \frac{V_{th}^{(0)} - V_{th}(t)}{\tau_{ref}};\; V_{th} → V_{th} + \Delta V_{th} \;(after \:a \:spike)$$
#
# As with the addition of a refractory conductance, this method has the advantage that the refractory period is not absolute, but depends on the input current. This method can be combined with the hard reset of the membrane potential after a spike, or with the addition of a hyperpolarizing potassium conductance, as for method 2 above to simulate action potentials.
#
# In the next section you will implement refractory period using a refractory conductance.
# + id="mLucUr3Rw1On"
def leaky_int_and_fire_ref_cond(t,i_in,model_constants):
### Inputs:
### t -> time (numpy array of doubles marking the time points where the simulation will be run)
### i_in -> input current (numpy array having the same dimension of t)
### model_constants -> structure containing all the constants for the model
### Output:
### v_m -> values of membrane potential over time (numpy array having the same dimension of t)
### sp -> spike train
### g_Ref -> refractory conductance
c_m = model_constants.c_m
g_l = model_constants.g_l
e_l = model_constants.e_l
e_k = model_constants.e_k
v_th = model_constants.v_th
tau_ref = model_constants.tau_ref
delta_g = model_constants.delta_g
dt = t[1]-t[0]
v_m = np.empty_like(t)
g_ref = np.empty_like(t)
v_m[0] = e_l
g_ref[0] = 0
if np.isscalar(i_in):
i_in_vec = np.full_like(t,i_in)
else:
i_in_vec = i_in
n_t = len(t)
sp = np.zeros(t.size)
for i in range(1,n_t):
v_m[i] = v_m[i-1] + dt/c_m*(g_l*(e_l-v_m[i-1]) + i_in_vec[i] + g_ref[i-1]*(e_k - v_m[i-1]))
g_ref[i] = g_ref[i-1] - dt*g_ref[i-1]/tau_ref
# add firing
if v_m[i] >= v_th:
g_ref[i] = g_ref[i] + delta_g
sp[i] = 1
return v_m, sp, g_ref
# + colab={"base_uri": "https://localhost:8080/", "height": 428} id="xsCweWEKIDiO" outputId="d0467133-6305-49f2-8aed-8c1207587534" cellView="form"
#@title Run LIF with refractory period
dt = 0.0001 #@param {type:"number"}
t_end = 0.5 #@param {type:"number"}
c_m = 100E-12 #@param {type:"number"}
g_l = 10E-9 #@param {type:"number"}
e_l = -70E-3 #@param {type:"number"}
e_k = -80E-3 #@param {type:"number"}
i_in = 240E-12 #@param {type:"number"}
v_th = -50E-3 #@param {type:"number"}
tau_ref = 2E-3 #@param {type:"number"}
delta_g = 500E-9 #@param {type:"number"}
t = np.arange(0,t_end,dt)
@dataclass
class Constants:
c_m: float
g_l: float
e_l: float
e_k: float
v_th: float
tau_ref: float
delta_g: float
model_constants = Constants(c_m,g_l,e_l,e_k,v_th,tau_ref,delta_g)
v_m, sp, g_ref = leaky_int_and_fire_ref_cond(t,i_in,model_constants)
fig,axs = plt.subplots(2,1)
axs[0].plot(t,v_m*1000,label="Membrane potential")
axs[0].eventplot(t[sp == 1], color='orchid',linelengths=10, label="Spikes")
axs[0].legend(loc='lower right')
axs[1].plot(t,g_ref*1E6,label="Refractory conductance")
_ = axs[0].set_xlabel("t [s]")
axs[0].set_ylabel("$V_m$ [mV]")
axs[1].set_ylabel("$G_{ref}$ [$\mu$S]")
_ =axs[1].set_xlabel("t [s]")
# + [markdown] id="CSsAUI7K17DJ"
# # Spike-rate-adaptation (SRA)
# Many neurons respond to a pulse of current with spikes that are progressively rarer over time.
# - Can you think of a reason why the brain does so? Can spike-rate adaptation have a 'meaning' in the brain?
#
# SRA can be modelled, as for the refractory period, with a refractory conductance. There are, though some differences between a refractory period and spike rate adapation. As you have seen in our previous tutorial exercise, our LIF neuron was showing a refractory period *wihthout* SRA! What did we miss?
#
# For the case of refractory conductance, each spike was causing a surge in $G_{ref}$, followed by an exponential decay. This was causing the membrane potential to drop and stay at small values for a specific amount of time, defined by $\tau_{ref}$. The refractory conductance was allowing us to keep trace of when a spike happened and causing the membrane potential to drop after a spike. Since the method was designed to cause a spike, its time scale had to be, by definition, the one of a spike! Try to see what happens if you modify $\tau_{ref}$ in the previous exercise.
#
# What would happen if we replaced the firing mechanism used for the refractory period with a hard threshold for generating spikes (like in the first implementation of the LIF), and keep the refractory conductance to simulate SRA?
# - How would you implement SRA?
# - How would you change the properties of the refractory conductance ($G_{ref}$ and $\tau_{ref}$) for this scope? Why?
#
# In the next section you will implement, from scratch, a SRA mechanism based on a refractory conductance plus a hard reset for the membrane potential as a firin mechanism.
# + id="MgdqWyzrcAwc"
def leaky_int_and_fire_sra(t,i_in,model_constants):
### Inputs:
### t -> time (numpy array of doubles marking the time points where the simulation will be run)
### i_in -> input current (numpy array having the same dimension of t)
### model_constants -> structure containing all the constants for the model
### Output:
### v_m -> values of membrane potential over time (numpy array having the same dimension of t)
### sp -> spike train
### g_ref -> refractory conductance
c_m = model_constants.c_m
g_l = model_constants.g_l
e_l = model_constants.e_l
e_k = model_constants.e_k
v_th = model_constants.v_th
v_reset = model_constants.v_reset
tau_sra = model_constants.tau_sra
delta_g = model_constants.delta_g
dt = t[1]-t[0]
v_m = np.empty_like(t)
g_ref = np.empty_like(t)
v_m[0] = e_l
g_ref[0] = 0
if np.isscalar(i_in):
i_in_vec = np.full_like(t,i_in)
else:
i_in_vec = i_in
n_t = len(t)
sp = np.zeros(t.size)
for i in range(1,n_t):
v_m[i] = v_m[i-1] + dt/c_m*(g_l*(e_l-v_m[i-1]) + i_in_vec[i] + g_ref[i-1]*(e_k - v_m[i-1]))
g_ref[i] = g_ref[i-1] - dt*g_ref[i-1]/tau_sra
# add firing
if v_m[i] >= v_th:
v_m[i] = v_reset
g_ref[i] = g_ref[i] + delta_g
sp[i] = 1
return v_m, sp, g_ref
# + colab={"base_uri": "https://localhost:8080/", "height": 428} id="KmxWoMJw7EZG" outputId="3dcc1626-3221-4d93-d5eb-f6cb3323805f" cellView="form"
#@title Run LIF with Spike Rate Adaptation
dt = 0.0001 #@param {type:"number"}
t_end = 0.5 #@param {type:"number"}
c_m = 100E-12 #@param {type:"number"}
g_l = 10E-9 #@param {type:"number"}
e_l = -70E-3 #@param {type:"number"}
e_k = -80E-3 #@param {type:"number"}
i_in = 400E-12 #@param {type:"number"}
v_th = -50E-3 #@param {type:"number"}
v_reset = -80E-3 #@param{type:"number"}
tau_sra = 0.2#@param {type:"number"}
delta_g = 1E-9 #@param {type:"number"}
t = np.arange(0,t_end,dt)
@dataclass
class Constants:
c_m: float
g_l: float
e_l: float
e_k: float
v_th: float
v_reset: float
tau_sra: float
delta_g: float
model_constants = Constants(c_m,g_l,e_l,e_k,v_th,v_reset,tau_sra,delta_g)
v_m, sp, g_ref = leaky_int_and_fire_sra(t,i_in,model_constants)
fig,axs = plt.subplots(2,1)
axs[0].plot(t,v_m*1000,label="Membrane potential")
axs[0].eventplot(t[sp == 1], color='orchid',linelengths=10, label="Spikes")
axs[0].legend(loc='lower right')
axs[1].plot(t,g_ref*1E6,label="Refractory conductance")
_ = axs[0].set_xlabel("t [s]")
axs[0].set_ylabel("$V_m$ [mV]")
axs[1].set_ylabel("$G_{ref}$ [$\mu$S]")
_ =axs[1].set_xlabel("t [s]")
# + [markdown] id="qY2PQD0EhzLi"
# # The Exponential Leaky Integrate and Fire with Adaptation (EALIF)
# The most striking limitation of a LIF neuron is the absence of a spiking mechanism.
# In all model neurons developed so far there is no actual action potential generated. On top of that, the slope of $V_m$ decreses as we get close to the threshold, which does not match what is observed in reality.
#
# Another simplification of the LIF is the definition of a fixed threshold for firing. In reality the firing threshold is not fixed, and depends on the state of the neuron (hence on prior inputs).
#
# The EALIF ([<NAME>., & <NAME>. (2005)](https://doi.org/10.1152/jn.00686.2005)) tries to solve those inconsistencies by introducing a non-linear spike-generating term in the differential equation for the membrane potential. On top of that, the model includes a second differential equation for a depolarizing current $I_{sra}$ that allows to account for spike-rate adaptation.
#
# $$ C_m\frac{dV_m(t)}{dt} = G_L\biggr[E_L-V_m(t) + \Delta_{th} e^{\frac{V_m(t) - V_{th}}{\Delta_{th}}}\biggr] + I_{app} - I_{sra}$$
# $$ \tau_{sra}\frac{dI_{sra}}{dt} = a(V_m - E_L) - I_{sra}$$
#
# the equations above are combined with the two reset rules if $V_m > V_{max}$:
# $$ V_m \mapsto V_{reset}$$
# $$I_{sra} \mapsto I_{sra} + b$$
#
# This model neuron is quite versatile as it generates a wide range of dynamics in its response. After implementing correctly the code you should be able to test it with the code cell below the next one. The code has already implemented some specific values of the model constants that generate very different neuronal responses.
# + id="H-ImqSTM6M6f"
def ealif(t,i_in,model_constants):
### Inputs:
### t -> time (numpy array of doubles marking the time points where the simulation will be run)
### i_in -> input current (numpy array having the same dimension of t)
### model_constants -> structure containing all the constants for the model
### Output:
### v_m -> values of membrane potential over time (numpy array having the same dimension of t)
### sp -> spike train
c_m = model_constants.c_m
g_l = model_constants.g_l
e_l = model_constants.e_l
delta_th = model_constants.delta_th
v_th = model_constants.v_th
v_reset = model_constants.v_reset
v_max = model_constants.v_max
tau_sra = model_constants.tau_sra
a = model_constants.a
b = model_constants.b
dt = t[1]-t[0]
v_m = np.empty_like(t)
i_sra = np.empty_like(t)
v_m[0] = e_l
i_sra[0] = 0
if np.isscalar(i_in):
i_in_vec = np.full_like(t,i_in)
else:
i_in_vec = i_in
n_t = len(t)
sp = np.zeros(t.size)
for i in range(1,n_t):
i_sra[i] = i_sra[i-1] + dt/tau_sra*(a*(v_m[i-1]-e_l) - i_sra[i-1])
v_m[i] = v_m[i-1] + dt/c_m*(g_l*(e_l-v_m[i-1] + delta_th*np.exp((v_m[i-1]-v_th)/delta_th)) + i_in_vec[i] - i_sra[i])
# reset conditions
if v_m[i] >= v_max:
v_m[i] = v_reset
i_sra[i] = i_sra[i] + b
sp[i] = 1
return v_m, sp, i_sra
# + colab={"base_uri": "https://localhost:8080/", "height": 428} cellView="form" id="bS00YDrX-7PZ" outputId="0cca774f-fe12-4d10-b13a-b8d216d25c6a"
#@title Run EALIF
@dataclass
class Constants:
c_m: float
g_l: float
e_l: float
delta_th: float
v_th: float
v_reset: float
v_max: float
tau_sra: float
a: float
b: float
model_constants = "delayed" #@param ["adapting", "tonic", "bursting","delayed"]
if model_constants == "adapting":
c_m = 100E-12
g_l = 10E-9
e_l = -70E-3
delta_th = 2E-3
v_th = -50E-3
v_reset = -80E-3
v_max = 50E-3
tau_sra = 0.2
a = 2E-9
b = 20E-12
i_in = 0.3E-9
elif model_constants == "tonic":
c_m = 100E-12
g_l = 10E-9
e_l = -70E-3
delta_th = 2E-3
v_th = -50E-3
v_reset = -80E-3
v_max = 50E-3
tau_sra = 0.2
a = 0
b = 0
i_in = 0.221E-9
elif model_constants == "bursting":
c_m = 10E-12
g_l = 2E-9
e_l = -70E-3
delta_th = 2E-3
v_th = -50E-3
v_reset = -46E-3
v_max = 50E-3
tau_sra = 0.1
a = -0.5E-9
b = 7E-12
i_in = 65E-12
elif model_constants == "delayed":
c_m = 10E-12
g_l = 2E-9
e_l = -70E-3
delta_th = 2E-3
v_th = -50E-3
v_reset = -60E-3
v_max = 50E-3
tau_sra = 0.1
a = -1E-9
b = 10E-12
i_in = 25E-12
dt = 0.001
t_end = 1
t = np.arange(0,t_end,dt)
model_constants = Constants(c_m,g_l,e_l,delta_th,v_th,v_reset,v_max,tau_sra,a,b)
v_m, sp, i_sra = ealif(t,i_in,model_constants)
fig,axs = plt.subplots(2,1)
axs[0].plot(t,v_m*1000,label="Membrane potential")
axs[0].eventplot(t[sp == 1], color='orchid',linelengths=10, label="Spikes")
axs[0].legend(loc='lower right')
_ = axs[0].set_xlabel("t [s]")
axs[0].set_ylabel("$V_m$ [mV]")
axs[1].plot(t,i_sra*1E9)
axs[0].set_ylabel("$V_m$ [mV]")
axs[1].set_ylabel("$I_{sra}$ [nA]")
_ =axs[1].set_xlabel("t [s]")
| LeakyIntegrateAndFireTutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#Selenium es una libreria de Python para automatizar scrapping, requiere del uso del chrome_driver
#sacar el xpath: con el navegador en las herramientas de desarrollo. Hay que cambiar las comillas dobles por simples
#DROPDOWN - <select> - 7.6
selec = Select(driver.find_element_by_xpath("//*[@id='main']/div[3]/div[1]/select"))
selec.select_by_value("5")
#CSS - 7.7
contenedor = css.find_element_by_css_selector('a.w3-blue')
contenedor.click()
#Tablas - <table> - 7.8
fila = len(tablas.find_elements_by_xpath("//*[@id='customers']/tbody/tr"))
colamnas = len(tablas.find_elements_by_xpath("//*[@id='customers']/tbody/tr[1]/th"))
#Scroll - 7.9
scroll.execute_script("window.scrollTo(0,document.body.scrollHeight)")
#Switch - 7.10
switch = driver.find_element_by_xpath("//*[@id='main']/label[3]/div")
switch.click()
#Radio Button - <input type="radio"> 7.11
boton = scraping.find_element_by_xpath("//*[@id='main']/div[3]/div[1]/input[4]")
boton.click()
#Posicionarnos en un enlace - 7.12
hipervinculo = enlace.find_element_by_link_text("Lo que pone en el enlace")
posicionar = ActionChains(enlace).move_to_element(hipervinculo)
posicionar.perform()
#Cookies - 7.13
all_cookie = cookie.get_cookies()
print(all_cookie)
#Capturar pantalla - 7.14
captura.get_screenshot_as_file("C:\\Users\\sergio\\Desktop\\Nueva carpeta (2)\\ejemplo.png")
#Subir archivos - <input type="file"> - 7.15
cargar.find_element_by_id("input-file-now").send_keys("C:\\Users\\sergio\\Desktop\\foto perfil\\Screenshot_2.png")
# -
| 0. INDICE de referencia.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **Download** (right-click, save target as ...) this page as a jupyterlab notebook from:
#
# [Laboratory 1](https://atomickitty.ddns.net/engr-1330-webroot/8-Labs/Lab01/Lab01.ipynb)
#
# ___
#
# # <font color=darkblue>Laboratory 1: A Notebook Like No Other!</font>
#
# **LAST NAME, FIRST NAME**
#
# **R00000000**
#
# ENGR 1330 Laboratory 1 - In-Lab
#
# Welcome to your first (or second) <span style='background:yellow'>Jupyter Notebook</span>. This is the medium that we will be using throughout the semester.
# ___
#
# **Why is this called a notebook?**
# Because you can write stuff in it!
#
# **Is that it?**
# Nope! you can **write** and **run** <span style='background:yellow'>CODE</span> in this notebook! Plus a bunch of other cool stuff such as making graphs, running tests and simulations, adding images, and building documents (such as this one!).
# ## <font color=purple>The Environment - Let's have a look around this window!</font>
#
#  <br>
# <font color=gray>*<NAME> in Mr. Robot*</font>
#
#
# - The tabs:
# - File
# - Edit
# - View
# - Insert
# - Cell
# - Kernel
#
# - The Icons:
# - Save
# - Insert Cell Below
# - Cut
# - Copy
# - Paste Cells Below
# - Move Up
# - Move Down
# - Run
# - Intruppt Kernel
# - Restart Kernel
# - Cell Type Selector (Dropdown list)
#
# ___
#
# The notebook consists of a sequence of cells. A cell is a multiline text input field, and its contents can be executed by using Shift-Enter, or by clicking Run in the menu bar. The execution behavior of a cell is determined by the cell’s type.
#
# There are three types of cells: code cells, markdown cells, and raw cells. Every cell starts off being a code cell, but its type can be changed by using a drop-down on the toolbar (which will be “Code”, initially).
#
#
# ## Code Cells:
#
# A code cell allows you to edit and write new code, with full syntax highlighting and tab completion. The programming language you use depends on the kernel. What we will use for this course and the default kernel IPython runs, is Python code.
#
# When a code cell is executed, code that it contains is sent to the kernel associated with the notebook. The results that are returned from this computation are then displayed in the notebook as the cell’s output. The output is not limited to text, with many other possible forms of output are also possible, including matplotlib figures and HTML tables. This is known as IPython’s rich display capability.
#
# ## Markdown Cells:
#
# You can document the computational process in a literate way, alternating descriptive text with code, using rich text. In IPython this is accomplished by marking up text with the Markdown language. The corresponding cells are called Markdown cells. The Markdown language provides a simple way to perform this text markup, that is, to specify which parts of the text should be emphasized (italics), bold, form lists, etc. In fact, markdown cells allow a variety of cool modifications to be applied:
#
#
# If you want to provide structure for your document, you can use markdown headings. Markdown headings consist of 1 to 5 hash # signs followed by a space and the title of your section. (The markdown heading will be converted to a clickable link for a section of the notebook. It is also used as a hint when exporting to other document formats, like PDF.) Here is how it looks:
#
# # # title
# ## ## major headings
# ### ### subheadings
# #### #### 4th level subheadings
# ##### ##### 5th level subheadings
#
#
# ### These codes are also quite useful:
# - Use triple " * " before and after a word (without spacing) to make the word bold and italic <br>
# B&I: ***string*** <br>
#
# - __ or ** before and after a word (without spacing) to make the word bold <br>
# Bold: __string__ or **string** <br>
#
# - _ or * before and after a word (without spacing to make the word italic <br>
# Italic: _string_ or *string* <br>
#
# - Double ~ before and after a word (without spacing to make the word scratched <br>
# Scratched: ~~string~~ <br>
#
# - For line breaks use "br" in the middle of <> <br>
#
# - For colors use this code:
# ### change this to a merkdown cell and run it!
# <font color=blue>Text</font> <br>
# <font color=red>Text</font> <br>
# <font color=orange>Text</font> <br>
# - For indented quoting, use a greater than sign (>) and then a space, then type the text. The text is indented and has a gray horizontal line to the left of it until the next carriage return.
# > here is an example of how it works!
#
# - For bullets, use the dash sign (- ) with a space after it, or a space, a dash, and a space ( - ), to create a circular bullet. To create a sub bullet, use a tab followed a dash and a space. You can also use an asterisk instead of a dash, and it works the same.
#
# - For numbered lists, start with 1. followed by a space, then it starts numbering for you. Start each line with some number and a period, then a space. Tab to indent to get subnumbering. <br>
#
# 1. first
# 2. second
# 3. third
# 4. ...
#
# - For horizontal lines: Use three asterisks: ***
# ***
# ***
# ***
#
# - For graphics, you can attach image files directly to a notebook only in Markdown cells. Drag and drop your images to the Mardown cell to attach it to the notebook.
#
#
# 
#
# - You can also use images from online sources be using this format:
#
#  <br>
#
#
# ## Raw Cells:
#
# Raw cells provide a place in which you can write output directly. Raw cells are not evaluated by the notebook.
# + active=""
# Thi$ is a raw ce11
# -
# ## <font color=purple>Let's meet world's most popular python!</font>
#
#  <br>
#
# ### What is python?
# > "Python is an interpreted, high-level and general-purpose programming language. Python's design philosophy emphasizes code readability with its notable use of significant whitespace." - Wikipedia @ [https://en.wikipedia.org/wiki/Python_(programming_language)](https://en.wikipedia.org/wiki/Python_(programming_language))
#
# ### How to have access to it?
#
# There are plenty of ways, from online compilers to our beloved Jupyter Notebook on your local machines. Here are a few examples of online compilers:
#
# a. https://www.programiz.com/python-programming/online-compiler/
#
# b. https://www.onlinegdb.com/online_python_compiler
#
# c. https://www.w3schools.com/python/python_compiler.asp
#
# d. https://repl.it/languages/python3
# We can do the exact same thing in this notebook. But we need a CODE cell.
print("Hello World")
# This is the classic "first program" of many languages! The script input is quite simple, we instruct the computer to print the literal string "hello world" to standard input/output device which is the console. Let's change it and see what happens:
print("This is my first notebook!")
# ### How to save a notebook?
#
# - __As a notebook file (.ipynb):__ <br>
# Go to File > Download As > Notebook (.ipynb)
#
# - __As an HTML file (.html):__ <br>
# Go to File > Download As > HTML (.html)
#
# - __As a Pdf (.pdf):__ <br>
# Go to File > Download As > PDF via LaTex (.pdf)
# or
# Save it as an HTML file and then convert that to a pdf via a website such as https://html2pdf.com/
#
# *Unless stated otherwise, we want you to submit your lab assignments in PDF and your exam and project deliverables in both PDF and .ipynb formats.*
# ___
# ## Readings
#
# *This notebook was inspired by several blogposts including:*
#
# - __"Markdown for Jupyter notebooks cheatsheet"__ by __<NAME>__ available at *https://medium.com/@ingeh/markdown-for-jupyter-notebooks-cheatsheet-386c05aeebed <br>
# - __"Jupyter Notebook: An Introduction"__ by __<NAME>__ available at *https://realpython.com/jupyter-notebook-introduction/ <br>
#
#
# *Here are some great reads on this topic:*
# - __"Jupyter Notebook Tutorial: The Definitive Guide"__ by __<NAME>__ available at *https://www.datacamp.com/community/tutorials/tutorial-jupyter-notebook <br>
# - __"Introduction to Jupyter Notebooks"__ by __<NAME>, <NAME>, and <NAME>__ available at *https://programminghistorian.org/en/lessons/jupyter-notebooks <br>
# - __"12 Things to know about Jupyter Notebook Markdown"__ by __<NAME>__ available at *https://medium.com/game-of-data/12-things-to-know-about-jupyter-notebook-markdown-3f6cef811707 <br>
#
#
# *Here are some great videos on these topics:*
# - __"Jupyter Notebook Tutorial: Introduction, Setup, and Walkthrough"__ by __<NAME>__ available at *https://www.youtube.com/watch?v=HW29067qVWk <br>
# - __"Quick introduction to Jupyter Notebook"__ by __<NAME>__ available at *https://www.youtube.com/watch?v=jZ952vChhuI <br>
# - __"What is Jupyter Notebook?"__ by __codebasics__ available at *https://www.youtube.com/watch?v=q_BzsPxwLOE <br>
# ___
# ## Exercise: Let's see who you are! <br>
#
# Similar to the hello world example, use a code cell and print a paragraph about you. You can introduce yourselves and write about interesting things to and about you! A few lines bekow can get you started, replace the ... parts and the other parts to make your paragraph.
print('my name is <NAME>')
print('my favorite food is tuna fish')
print('I am currently studying to be an biological transmutation engineer')
print('I speak 3 languages, they are: cat, english for cats, and of course profanity')
#
| 8-Labs/Lab01/dev_src/Lab01-WS.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/heros-lab/colaboratory/blob/master/verfication_results2.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="_W92yPHiMz83" colab_type="code" colab={}
# # +++ Import Packages for Experiments +++
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib.ticker as ptick
from google.colab import drive
drive.mount('/content/drive')
work_path = "/content/drive/My Drive/Colab Notebooks"
# + id="EPTDyY3XNCdo" colab_type="code" colab={}
class DatasetClass:
def __init__(self, data_path):
self.path = data_path
def __reflect_index(self, data, index):
if index != None:
data = data[:, index]
return data
def __load_df(self, data_label):
data_x = pd.read_csv(f"{self.path}/{data_label}_x.csv", index_col=0)
data_y = pd.read_csv(f"{self.path}/{data_label}_y.csv", index_col=0)
return data_x, data_y
def __load_data(self, data_label, x_index, y_index):
data_x, data_y = self.__load_df(data_label)
data_x = self.__reflect_index(data_x.values, x_index)
data_y = self.__reflect_index(data_y.values, y_index)
return data_x, data_y
def __load_stack(self, dataset_list, x_index, y_index):
for label in dataset_list:
tmp_x, tmp_y = self.__load_data(label, x_index, y_index)
if dataset_list.index(label) == 0:
data_x = tmp_x
data_y = tmp_y
else:
data_x = np.vstack((data_x, tmp_x))
data_y = np.vstack((data_y, tmp_y))
return data_x, data_y
def __load_dict(self, dataset_list, x_index, y_index):
data_x, data_y = {}, {}
for label in dataset_list:
tmp_x, tmp_y = self.__load_data(label, x_index, y_index)
data_x[label] = tmp_x
data_y[label] = tmp_y
return data_x, data_y
def get_data(self, dataset_label, x_index=None, y_index=None, dict_type:bool=False):
if not dict_type:
if type(dataset_label) == str:
data_x, data_y = self.__load_data(dataset_label, x_index, y_index)
else:
data_x, data_y = self.__load_stack(dataset_label, x_index, y_index)
else:
data_x, data_y = self.__load_dict(dataset_label, x_index, y_index)
return data_x, data_y
def get_dataframe(self, dataset_list):
data_x = {}
data_y = {}
for label in dataset_list:
tmp_x, tmp_y = self.__load_df(label)
data_x[label] = tmp_x
data_y[label] = tmp_y
return data_x, data_y
class NormsDatasetClass:
def __init__(self, data_path):
self.path = data_path
def __reflect_index(self, data, index):
if index != None:
data = data[:, index]
return data
def __load_df(self, data_label):
data_x = pd.read_csv(f"{self.path}/{data_label}_nx.csv", index_col=0)
data_y = pd.read_csv(f"{self.path}/{data_label}_ny.csv", index_col=0)
return data_x, data_y
def __load_data(self, data_label, x_index, y_index):
data_x, data_y = self.__load_df(data_label)
data_x = self.__reflect_index(data_x.values, x_index)
data_y = self.__reflect_index(data_y.values, y_index)
return data_x, data_y
def __load_stack(self, dataset_list, x_index, y_index):
for label in dataset_list:
tmp_x, tmp_y = self.__load_data(label, x_index, y_index)
if dataset_list.index(label) == 0:
data_x = tmp_x
data_y = tmp_y
else:
data_x = np.vstack((data_x, tmp_x))
data_y = np.vstack((data_y, tmp_y))
return data_x, data_y
def __load_dict(self, dataset_list, x_index, y_index):
data_x, data_y = {}, {}
for label in dataset_list:
tmp_x, tmp_y = self.__load_data(label, x_index, y_index)
data_x[label] = tmp_x
data_y[label] = tmp_y
return data_x, data_y
def get_data(self, dataset_label, x_index=None, y_index=None, dict_type:bool=False):
if not dict_type:
if type(dataset_label) == str:
data_x, data_y = self.__load_data(dataset_label, x_index, y_index)
else:
data_x, data_y = self.__load_stack(dataset_label, x_index, y_index)
else:
data_x, data_y = self.__load_dict(dataset_label, x_index, y_index)
return data_x, data_y
def get_dataframe(self, dataset_list):
data_x = {}
data_y = {}
for label in dataset_list:
tmp_x, tmp_y = self.__load_df(label)
data_x[label] = tmp_x
data_y[label] = tmp_y
return data_x, data_y
# + id="V4OzYPp9QdWn" colab_type="code" colab={}
def filtering(data_list):
pd_series = pd.Series(data_list)
q1 = pd_series.quantile(.25)
q3 = pd_series.quantile(.75)
iqr = q3 - q1
lim_upper = q3 + iqr*1.5
lim_lower = q1 - iqr*1.5
#return pd_series[pd_series.apply(lambda x:lim_lower < x < lim_upper)]
return pd_series.apply(lambda x:lim_lower < x < lim_upper)
# + id="93xaBZymPBy1" colab_type="code" colab={}
def norms(x_vec, x_max:float, x_min:float):
norm = np.vectorize(lambda x:(x - x_min)/(x_max - x_min))
return norm(x_vec)
def restore(x_vec, x_max:float, x_min:float):
rest = np.vectorize(lambda x: (x_max - x_min)*x+x_min)
return rest(x_vec)
x_max = [2.79251417013236, 1001.76434281299, 11.1129894774574, 50.878307668365]
x_min = [-1.68539431728585, -8.00252854145341, -14.5050341893581, -46.6201668706979]
# + id="zuxf0Z7NO2rg" colab_type="code" colab={}
model_c = "Conv.2"
model_p = "Prop.2"
test_name = ["ms1a", "free", "step"]
type_id = 0
dataset = DatasetClass(f"{work_path}/data")
dataset_norm = NormsDatasetClass(f"{work_path}/data/norms")
plot_x, plot_y = dataset.get_data(test_name, y_index=[type_id], dict_type=True)
test_x, test_y = dataset_norm.get_data(test_name, y_index=[type_id], dict_type=True)
df_msrs_c = pd.read_csv(f"{work_path}/estimated_response/{model_c}_msrs.csv", index_col=0)
df_free_c = pd.read_csv(f"{work_path}/estimated_response/{model_c}_free.csv", index_col=0)
df_step_c = pd.read_csv(f"{work_path}/estimated_response/{model_c}_step.csv", index_col=0)
df_msrs_p = pd.read_csv(f"{work_path}/estimated_response/{model_p}_msrs.csv", index_col=0)
df_free_p = pd.read_csv(f"{work_path}/estimated_response/{model_p}_free.csv", index_col=0)
df_step_p = pd.read_csv(f"{work_path}/estimated_response/{model_p}_step.csv", index_col=0)
# + id="VRXGdT9HfyfC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 236} outputId="3483aee4-f541-47af-cb49-e94b38ff0a5a"
# 外れ値を含むサンプルの検出
error_msrs_c = df_msrs_c.values - test_y[test_name[0]]
error_msrs_p = df_msrs_p.values - test_y[test_name[0]]
rms_msrs_c = np.sqrt((error_msrs_c**2).mean(axis=0))
rms_msrs_p = np.sqrt((error_msrs_p**2).mean(axis=0))
index_c = filtering(rms_msrs_c)
index_p = filtering(rms_msrs_p)
# 外れ値を除いた推定値の算出
resp_msrs_c = df_msrs_c.values[:,index_c].mean(axis=1)
resp_free_c = df_free_c.values[:,index_c].mean(axis=1)
resp_step_c = df_step_c.values[:,index_c].mean(axis=1)
resp_msrs_p = df_msrs_p.values[:,index_p].mean(axis=1)
resp_free_p = df_free_p.values[:,index_p].mean(axis=1)
resp_step_p = df_step_p.values[:,index_p].mean(axis=1)
print(f"Conv. model's sample is {df_msrs_c.values[:, index_c].shape[1]}/101")
print(f"Prop. model's sample is {df_msrs_p.values[:, index_p].shape[1]}/101")
# 正規化した推定値の復元
resp_msrs_c = restore(resp_msrs_c, x_max[type_id], x_min[type_id])
resp_free_c = restore(resp_free_c, x_max[type_id], x_min[type_id])
resp_step_c = restore(resp_step_c, x_max[type_id], x_min[type_id])
resp_msrs_p = restore(resp_msrs_p, x_max[type_id], x_min[type_id])
resp_free_p = restore(resp_free_p, x_max[type_id], x_min[type_id])
resp_step_p = restore(resp_step_p, x_max[type_id], x_min[type_id])
# 推定誤差の算出
error_msrs_c = plot_y[test_name[0]][:, 0] - resp_msrs_c
error_free_c = plot_y["free"][:, 0] - resp_free_c
error_step_c = plot_y["step"][:, 0] - resp_step_c
error_msrs_p = plot_y[test_name[0]][:, 0] - resp_msrs_p
error_free_p = plot_y["free"][:, 0] - resp_free_p
error_step_p = plot_y["step"][:, 0] - resp_step_p
# + id="FjDNMnupq6tY" colab_type="code" colab={}
def rms(x_vec):
return np.sqrt((x_vec**2).mean())
# + id="Kw-Zok_Pms5v" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="34f405ca-64ef-42a7-d81c-f937a2b6038b"
rms_c = rms(error_msrs_c)
rms_p = rms(error_msrs_p)
print(f"ms2a: conv->{rms_c:.4e}, prop->{rms_p:.4e}")
rms_c = rms(error_free_c)
rms_p = rms(error_free_p)
print(f"free: conv->{rms_c:.4e}, prop->{rms_p:.4e}")
rms_c = rms(error_step_c)
rms_p = rms(error_step_p)
print(f"step: conv->{rms_c:.4e}, prop->{rms_p:.4e}")
# + id="w6mfPZHucWxy" colab_type="code" colab={}
time10 = np.arange(0,10,0.001)
time20 = np.arange(0,20,0.001)
# + id="DPCJci6-TwgN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 898} outputId="5fc3ba76-b77a-49df-e4d8-3b2addb6a10f"
fig, ax = plt.subplots(2, 1, figsize=(5, 5), dpi=200)
plot_y_ms = plot_y[test_name[0]][:,0]
ax[0].plot(time10, plot_y_ms, "r:")
ax[0].plot(time10, resp_msrs_c)
ax[0].plot(time10, resp_msrs_p)
ax[0].set_ylabel("Estimated response")
ax[0].legend(["Reference", "Conventional Network", "Proposed Network"], loc=1)
ax[0].grid()
ax[1].plot(time10, error_msrs_c)
ax[1].plot(time10, error_msrs_p)
ax[1].set_ylabel("Estimated error")
ax[1].set_xlabel("Time [sec]")
ax[1].grid()
# + id="ZGwCIW4UaNCy" colab_type="code" colab={}
fig, ax = plt.subplots(2, 1, figsize=(5, 5), dpi=300)
plot_y_free = plot_y["free"][:,0]
ax[0].plot(time20, plot_y_free, "r:")
ax[0].plot(time20, resp_free_c)
ax[0].plot(time20, resp_free_p)
ax[0].set_ylabel("Estimated response")
ax[0].legend(["Reference", "Conventional Network", "Proposed Network"], loc=1)
ax[0].grid()
ax[1].plot(time20, error_free_c)
ax[1].plot(time20, error_free_p)
ax[1].set_ylabel("Estimated error")
ax[1].set_xlabel("Time [sec]")
ax[1].grid()
ax3 = plt.axes([0.28, 0.275, 0.6, 0.15])
ax3.plot(time20[5000:], error_free_c[5000:])
ax3.plot(time20[5000:], error_free_p[5000:])
ax3.yaxis.set_major_formatter(ptick.ScalarFormatter(useMathText=True))
ax3.ticklabel_format(style="sci", axis="y", scilimits=(0,0))
ax3.yaxis.offsetText.set_fontsize(9)
ax3.grid()
# + id="MdDqPpFgnv02" colab_type="code" colab={}
fig, ax = plt.subplots(2, 1, figsize=(5, 5), dpi=300)
plot_y_step = plot_y["step"][:,0]
ax[0].plot(time20, plot_y_step, "r:")
ax[0].plot(time20, resp_step_c)
ax[0].plot(time20, resp_step_p)
ax[0].set_ylabel("Estimated response")
ax[0].legend(["Reference", "Conventional Network", "Proposed Network"], loc=1)
ax[0].grid()
ax[1].plot(time20, error_step_c)
ax[1].plot(time20, error_step_p)
ax[1].set_ylabel("Estimated error")
ax[1].set_xlabel("Time [sec]")
ax[1].grid()
# + id="B214hmLzqLsj" colab_type="code" colab={}
| verfication_results2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# + [markdown] nbgrader={"grade": false, "grade_id": "q2_prompt", "locked": true, "solution": false}
# # Q2
#
# More loops, this time with generators.
# + [markdown] nbgrader={"grade": false, "grade_id": "q2a_prompt", "locked": true, "solution": false}
# ### A
#
# Print out the corresponding elements of the three lists, `list1`, `list2`, and `list3`, simultaneously using a loop e.g. `list1[0]`, `list2[0]`, and `list3[0]`.
# + nbgrader={"grade": true, "grade_id": "q2a", "locked": false, "points": 5, "solution": true}
import numpy as np
np.random.seed(85473)
list1 = np.random.randint(100, size = 10).tolist()
list2 = np.random.randint(100, size = 10).tolist()
list3 = np.random.randint(100, size = 10).tolist()
### BEGIN SOLUTION
### END SOLUTION
# + [markdown] nbgrader={"grade": false, "grade_id": "q2b_prompt", "locked": true, "solution": false}
# ### B
#
# Print out the *indices* where the corresponding elements of all four lists--`list1`, `list2`, `list3`, and `list4`--are exactly the same.
# + nbgrader={"grade": true, "grade_id": "q2b", "locked": false, "points": 5, "solution": true}
import numpy as np
np.random.seed(5894292)
list1 = np.random.randint(3, size = 100).tolist()
list2 = np.random.randint(3, size = 100).tolist()
list3 = np.random.randint(3, size = 100).tolist()
list4 = np.random.randint(3, size = 100).tolist()
### BEGIN SOLUTION
### END SOLUTION
# + [markdown] nbgrader={"grade": false, "grade_id": "q2c_prompt", "locked": true, "solution": false}
# ### C
#
# Write a loop that iterates through both `list1` and `list2` simultaneously, and which halts immediately once the sum of the two corresponding *elements* of the lists is greater than 15.
# + nbgrader={"grade": true, "grade_id": "q2c", "locked": false, "points": 5, "solution": true}
import numpy as np
np.random.seed(447388)
list1 = np.random.randint(10, size = 10).tolist()
list2 = np.random.randint(10, size = 10).tolist()
### BEGIN SOLUTION
### END SOLUTION
# + [markdown] nbgrader={"grade": false, "grade_id": "q2d_prompt", "locked": true, "solution": false}
# ### D
#
# Create a new dictionary where the keys and values of `original_dictionary` are flipped.
# + nbgrader={"grade": true, "grade_id": "q2d", "locked": false, "points": 5, "solution": true}
original_dictionary = {1:2, 3:4, 5:6, 7:8, 9:10}
print(original_dictionary)
new_dictionary = {}
### BEGIN SOLUTION
### END SOLUTION
| assignments/A3/A3_Q2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + tags=[]
import time
start = time.time()
# + tags=[]
import pyspark
import os
# + tags=[]
# %%local
import pandas as pd
import plotly.graph_objects as go
# + tags=[]
file = 's3://millionsongs-data/new_g4g.csv'
data = sc.textFile(file)
# + tags=[]
header = data.flatMap(lambda x: x.split('\n')).map(
lambda x: x.split('\t')).first()
# + tags=[]
header_dict = {head: i for i, head in enumerate(header)}
# -
input_rdd = data.flatMap(lambda x: x.split('\n')).map(
lambda x: x.split('\t')).filter(lambda x: x != header)
input_rdd.cache()
# + tags=[]
popularity_rdd = input_rdd.map(lambda x: (x[header_dict['get_artist_name']], x[header_dict['get_artist_latitude']], x[header_dict['get_artist_longitude']],
x[header_dict['get_artist_hotttnesss']])).filter(lambda x: x[1] != '').map(lambda x: (x[0], float(x[1]), float(x[2]), float(x[3]))).filter(lambda x: x[3] > 0.5)
# + tags=[]
popularity_df = popularity_rdd.toDF()
popularity_df.createOrReplaceTempView('popularity_view')
# + tags=[] magic_args="-o df_popularity" language="sql"
# SELECT * FROM popularity_view
# -
# %%local
import plotly.graph_objects as go
fig = go.Figure(data=go.Scattergeo(
lon=df_popularity['_3'],
lat=df_popularity['_2'],
text=df_popularity['_4'],
mode='markers',
marker=dict(
size=4,
opacity=0.7,
reversescale=False,
autocolorscale=False,
symbol='square',
line=dict(
width=1,
color='rgba(102, 102, 102)'
),
colorscale='Blues',
cmin=0,
color=df_popularity['_4'],
cmax=df_popularity['_4'].max(),
colorbar_title="Artist popularity"
)))
fig.show()
artist_count = input_rdd.map(lambda x: (
x[header_dict['get_artist_name']], x[header_dict['get_artist_location']])).groupByKey().mapValues(lambda x: len(list(x)))
artist_location = input_rdd.map(lambda x: (x[header_dict['get_artist_name']], x[header_dict['get_artist_location']], float(
x[header_dict['get_artist_hotttnesss']]))).filter(lambda x: x[2] > 0.5).distinct()
# + tags=[]
artist_ct_loc = artist_location.join(artist_count, numPartitions=7).distinct().map(lambda x: (x[1][0], (x[0], x[1][1])))\
.filter(lambda x: x[0] != '').map(lambda x: (x[0], 1))\
.reduceByKey(lambda x, y: x+y).sortBy(lambda x: x[1], ascending=False, numPartitions=7)
# + tags=[]
artist_df = artist_ct_loc.toDF()
artist_df.createOrReplaceTempView('artist_view')
# + tags=[] magic_args="-o df_artist" language="sql"
# SELECT * FROM artist_view limit 10
# -
# %%local
import plotly.express as px
fig = px.bar(df_artist, x='_1', y='_2')
fig.update_layout(
height=400,
title_text=' #Location count of popular artist by city',
yaxis_title="#Number of songs",
xaxis_title="Year"
)
fig.show()
stop = time.time()
stop-start
sc.stop()
| EDA_EMR_notebooks/artist_location_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/agemagician/CodeTrans/blob/main/prediction/multitask/pre-training/function%20documentation%20generation/ruby/small_model.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="c9eStCoLX0pZ"
# **<h3>Predict the documentation for ruby code using codeTrans multitask training model</h3>**
# <h4>You can make free prediction online through this
# <a href="https://huggingface.co/SEBIS/code_trans_t5_small_code_documentation_generation_ruby_multitask">Link</a></h4> (When using the prediction online, you need to parse and tokenize the code first.)
# + [markdown] id="6YPrvwDIHdBe"
# **1. Load necessry libraries including huggingface transformers**
# + colab={"base_uri": "https://localhost:8080/"} id="6FAVWAN1UOJ4" outputId="38618b37-19c0-439d-c2f4-574c0125a35c"
# !pip install -q transformers sentencepiece
# + id="53TAO7mmUOyI"
from transformers import AutoTokenizer, AutoModelWithLMHead, SummarizationPipeline
# + [markdown] id="xq9v-guFWXHy"
# **2. Load the token classification pipeline and load it into the GPU if avilabile**
# + colab={"base_uri": "https://localhost:8080/", "height": 316, "referenced_widgets": ["3c3e7bfc3cda4b018174b879815d0e97", "4527ece993c84c07b6e9d1cf18d6d0cc", "4546fd274db9470ca040160f994513e1", "e213889b71f345a299b1b94d3a430885", "<KEY>", "<KEY>", "8e5cedb0fd6642258986eaf142986df5", "c6aa3d77e50b4a329ffc27f40a9900ed", "<KEY>", "<KEY>", "8f4f52e19eb14a5f89971ed4b0c9f4e7", "10d3e3e8dc18486fbb636e5f972e125d", "<KEY>", "9cda2dd721e84a629b6d05f294668a7d", "2f213988efea4f3e88f6ad5ca725b5d6", "1a78a02f9fd2405a9736d12a1f05880a", "<KEY>", "8a7125b459ce4d5dadfd2320e29a5188", "56fd002622734a4f96fe79ed19b9777c", "<KEY>", "408c9a202b3844868721158f6fb327e9", "<KEY>", "7b15bed71ee2402b9bd1e90aabe9582f", "045ad1a1daec40e597559e1e6c15c5b3", "<KEY>", "a4aada90817844e48761a26312d8b073", "573c2eabff3a43d0b105e6ed5df7e1f2", "f1002e0e301e40f6b0eee7f397f1f31c", "3051c7708ad2481e9fed47dd953626c2", "147f0211e93d46449a3b1a000afe1990", "594e57e244684c819e46a244f81d970a", "ef3a99e764194cdd8fa080bfe5fac2d7", "e3b630c64a0544cb84e65845712bf071", "<KEY>", "<KEY>", "549a2a79826c4ec89fbef61207bec77e", "<KEY>", "<KEY>", "<KEY>", "63675e3b1a7c42f59a7de306fbfe5225"]} id="5ybX8hZ3UcK2" outputId="c4e2aad3-30a0-4d5c-93da-8d01bf8e0fe5"
pipeline = SummarizationPipeline(
model=AutoModelWithLMHead.from_pretrained("SEBIS/code_trans_t5_small_code_documentation_generation_ruby_multitask"),
tokenizer=AutoTokenizer.from_pretrained("SEBIS/code_trans_t5_small_code_documentation_generation_ruby_multitask", skip_special_tokens=True),
device=0
)
# + [markdown] id="hkynwKIcEvHh"
# **3 Give the code for summarization, parse and tokenize it**
# + id="nld-UUmII-2e"
code = "def add(severity, progname, &block)\n return true if io.nil? || severity < level\n message = format_message(severity, progname, yield)\n MUTEX.synchronize { io.write(message) }\n true\n end" #@param {type:"raw"}
# + id="cJLeTZ0JtsB5" colab={"base_uri": "https://localhost:8080/"} outputId="26e27014-e5c0-4576-9611-b34b1aaefd49"
# !pip install tree_sitter
# !git clone https://github.com/tree-sitter/tree-sitter-ruby
# + id="hqACvTcjtwYK"
from tree_sitter import Language, Parser
Language.build_library(
'build/my-languages.so',
['tree-sitter-ruby']
)
RUBY_LANGUAGE = Language('build/my-languages.so', 'ruby')
parser = Parser()
parser.set_language(RUBY_LANGUAGE)
# + id="LLCv2Yb8t_PP"
def get_string_from_code(node, lines):
line_start = node.start_point[0]
line_end = node.end_point[0]
char_start = node.start_point[1]
char_end = node.end_point[1]
if line_start != line_end:
code_list.append(' '.join([lines[line_start][char_start:]] + lines[line_start+1:line_end] + [lines[line_end][:char_end]]))
else:
code_list.append(lines[line_start][char_start:char_end])
def my_traverse(node, code_list):
lines = code.split('\n')
if node.child_count == 0:
get_string_from_code(node, lines)
elif node.type == 'string':
get_string_from_code(node, lines)
else:
for n in node.children:
my_traverse(n, code_list)
return ' '.join(code_list)
# + id="BhF9MWu1uCIS" colab={"base_uri": "https://localhost:8080/"} outputId="2d641e36-9136-4173-cb95-98f425877ce0"
tree = parser.parse(bytes(code, "utf8"))
code_list=[]
tokenized_code = my_traverse(tree.root_node, code_list)
print("Output after tokenization: " + tokenized_code)
# + [markdown] id="sVBz9jHNW1PI"
# **4. Make Prediction**
# + colab={"base_uri": "https://localhost:8080/"} id="KAItQ9U9UwqW" outputId="65db7af1-4835-4007-9a43-803a834ad66a"
pipeline([tokenized_code])
| prediction/multitask/pre-training/function documentation generation/ruby/small_model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img style="float: right; margin: 0px 0px 15px 15px;" src="https://encrypted-tbn0.gstatic.com/images?q=tbn:ANd9GcSQt6eQo8JPYzYO4p6WmxLtccdtJ4X8WR6GzVVKbsMjyGvUDEn1mg" width="300px" height="100px" />
#
# # Trabajando con opciones
# Una opción puede negociarse en el mercado secundario por lo que es importante determinar su valor $V_t$ para cada tiempo $t\in [0, T]$. La ganancia que obtiene quién adquiere la opción se llama función de pago o "payoff" y claramente depende del valor del subyacente.
#
# Hay una gran variedad de opciones en el mercado y éstas se clasiflcan según su función de pago y la forma en que pueden ejercerse. Las opciones que tienen como función de pago a
# $$ P(S(t),t)=max\{S(T)-K,0\} \rightarrow \text{En el caso de Call}$$
# $$ P(S(t),t)=max\{K-S(T),0\} \rightarrow \text{En el caso de Put}$$
# se llaman opciones **Vainilla**, con $h:[0,\infty) \to [0,\infty)$.
#
# La opción se llama **europea** si puede ejercerse sólo en la fecha de vencimiento.
#
# Se dice que una opción es **americana** si puede ejercerse en cualquier momento antes o en la fecha de vencimiento.
#
# Una opción compleja popular son las llamadas **opciones asiáticas** cuyos pagos dependen de todas las trayectorias del precio de los activos subyacentes. Las opciones cuyos pagos dependen de las trayectorias de los precios de los activos subyacentes se denominan opciones dependientes de la ruta.
#
# Principalmente, se puede resumir que las dos razones con más peso de importancia para utilizar opciones son el **aseguramiento** y la **especulación**.
#
# ## Opciones Plan Vainilla: opción de compra y opción de venta europea
#
# Una opción vainilla o estándar es una opción normal de compra o venta que no tiene características especiales o inusuales. Puede ser para tamaños y vencimientos estandarizados, y negociarse en un intercambio.
# En comparación con otras estructuras de opciones, las opciones de vanilla no son sofisticadas o complicadas.
#
# ## 1. ¿Cómo descargar datos de opciones?
#importar los paquetes que se van a usar
import pandas as pd
import pandas_datareader.data as web
import numpy as np
import datetime
import matplotlib.pyplot as plt
import scipy.stats as st
import seaborn as sns
# %matplotlib inline
#algunas opciones para Pandas
pd.set_option('display.notebook_repr_html', True)
pd.set_option('display.max_columns', 6)
pd.set_option('display.max_rows', 10)
pd.set_option('display.width', 78)
pd.set_option('precision', 3)
# Usando el paquete `pandas_datareader` también podemos descargar datos de opciones. Por ejemplo, descarguemos los datos de las opciones cuyo activo subyacente son las acciones de Apple
aapl = web.YahooOptions('AAPL')
aapl_opt = aapl.get_all_data().reset_index()
aapl_opt.set_index('Expiry')
# aapl
aapl_opt.loc[0]
aapl_opt.loc[0, 'JSON']
# ### Conceptos claves
# - El precio de la oferta ('bid') se refiere al precio más alto que un comprador pagará por un activo.
# - El precio de venta ('ask') se refiere al precio más bajo que un vendedor aceptará por un activo.
# - La diferencia entre estos dos precios se conoce como 'spread'; cuanto menor es el spread, mayor es la liquidez de la garantía dada.
# - Liquidez: facilidad de convertir cierta opción en efectivo.
# - La volatilidad implícita es el pronóstico del mercado de un probable movimiento en el precio de un valor.
# - La volatilidad implícita aumenta en los mercados bajistas y disminuye cuando el mercado es alcista.
# - El último precio ('lastprice') representa el precio al que ocurrió la última operación, de una opción dada.
# Una vez tenemos la información, podemos consultar de qué tipo son las opciones
aapl_opt.loc[:, 'Type']
# o en que fecha expiran
pd.set_option('display.max_rows', 10)
aapl_opt.loc[:, 'Expiry']
# Por otra parte, podríamos querer consultar todas las opciones de compra (call) que expiran en cierta fecha (2020-06-19)
fecha1 = '2020-06-19'
fecha2 = '2021-01-15'
call06 = aapl_opt.loc[(aapl_opt.Expiry== fecha1) & (aapl_opt.Type=='call')]
call06
# ## 2. ¿Qué es la volatilidad implícita?
# **Volatilidad:** desviación estándar de los rendimientos.
# - ¿Cómo se calcula?
# - ¿Para qué calcular la volatilidad?
# - **Para valuar derivados**, por ejemplo **opciones**.
# - Método de valuación de riesgo neutral (se supone que el precio del activo $S_t$ no se ve afectado por el riesgo de mercado).
#
# Recorderis de cuantitativas:
# 1. Ecuación de Black-Scholes
# $$ dS(t) = \mu S(t) + \sigma S(t)dW_t$$
# 2. Solución de la ecuación
#
# El valor de una opción Europea de vainilla $V_t$ puede obtenerse por:
# $$V_t = F(t,S_t)$$ donde
# 
# 3. Opción de compra europea, suponiendo que los precios del activo son lognormales
# 4. Opción de venta europea, suponiendo que los precios del activo son lognormales
# Entonces, ¿qué es la **volatilidad implícita**?
#
# La volatilidad es una medida de la incertidumbre sobre el comportamiento futuro de un activo, que se mide habitualmente como la desviación típica de la rentabilidad de dicho activo.
# ## Volatility smile
# - Cuando las opciones con la misma fecha de vencimiento y el mismo activo subyacente, pero diferentes precios de ejercicio, se grafican por la volatilidad implícita, la tendencia es que ese gráfico muestre una sonrisa.
# - La sonrisa muestra que las opciones más alejadas 'in- or out-of-the-money' tienen la mayor volatilidad implícita.
# - No todas las opciones tendrán una sonrisa de volatilidad implícita. Las opciones de acciones a corto plazo y las opciones relacionadas con la moneda tienen más probabilidades de tener una sonrisa de volatilidad
#
# 
#
# > Fuente: https://www.investopedia.com/terms/v/volatilitysmile.asp
# > ### Validar para la `fecha = '2020-01-17'` y para la fecha `fecha = '2021-01-15'`
ax = call06.set_index('Strike').loc[:, 'IV'].plot(figsize=(8,6))
ax.axvline(call06.Underlying_Price.iloc[0], color='g');
# call06.Underlying_Price
put06 = aapl_opt.loc[(aapl_opt.Expiry==fecha1) & (aapl_opt.Type=='put')]
put06
ax = put06.set_index('Strike').loc[:, 'IV'].plot(figsize=(8,6))
ax.axvline(put06.Underlying_Price.iloc[0], color='g')
# Con lo que hemos aprendido, deberíamos ser capaces de crear una función que nos devuelva un `DataFrame` de `pandas` con los precios de cierre ajustados de ciertas compañías en ciertas fechas:
# - Escribir la función a continuación
# Función para descargar precios de cierre ajustados:
def get_adj_closes(tickers, start_date=None, end_date=None):
# Fecha inicio por defecto (start_date='2010-01-01') y fecha fin por defecto (end_date=today)
# Descargamos DataFrame con todos los datos
closes = web.DataReader(name=tickers, data_source='yahoo', start=start_date, end=end_date)
# Solo necesitamos los precios ajustados en el cierre
closes = closes['Adj Close']
# Se ordenan los índices de manera ascendente
closes.sort_index(inplace=True)
return closes
# - Obtener como ejemplo los precios de cierre de Apple del año pasado hasta la fecha. Graficar...
# +
ticker = ['AAPL']
start_date = '2017-01-01'
closes_aapl = get_adj_closes(ticker, start_date)
closes_aapl.plot(figsize=(8,5));
plt.legend(ticker);
# -
# - Escribir una función que pasándole el histórico de precios devuelva los rendimientos logarítmicos:
def calc_daily_ret(closes):
return np.log(closes/closes.shift(1)).iloc[1:]
# - Graficar...
ret_aapl = calc_daily_ret(closes_aapl)
ret_aapl.plot(figsize=(8,6));
# También, descargar datos de opciones de Apple:
aapl = web.YahooOptions('AAPL')
aapl_opt = aapl.get_all_data().reset_index()
aapl_opt.set_index('Expiry').sort_index()
indice_opt = aapl_opt.loc[(aapl_opt.Type=='call') & (aapl_opt.Strike==250) & (aapl_opt.Expiry=='2021-01-15')]
indice_opt
i_opt= indice_opt.index
opcion_valuar = aapl_opt.loc[i_opt[0]]
opcion_valuar['JSON']
print('Precio del activo subyacente actual = ',opcion_valuar.Underlying_Price)
# # Simulación de precios usando rendimiento simple y logarítmico
# * Comenzaremos por suponer que los rendimientos son un p.e. estacionario que distribuyen $\mathcal{N}(\mu,\sigma)$.
# +
# Descargamos los precios de apple
ticker = ['AAPL']
start_date = '2017-01-01'
closes_aapl = get_adj_closes(ticker, start_date)
closes_aapl
# -
# - **Rendimiento Simple**
# Obtenemos el rendimiento simple
Ri = closes_aapl.pct_change(1).iloc[1:]
# Obtenemos su media y desviación estándar de los rendimientos
mu_R = Ri.mean()[0]
sigma_R = Ri.std()[0]
Ri
from datetime import date
Hoy = date.today()
# ndays = 109
nscen = 10
dates = pd.date_range(start=Hoy, end='2021-01-15') #periods = ndays)
ndays = len(dates)
dates
dt = 1; # Rendimiento diario
Z = np.random.randn(ndays,nscen) # Z ~ N(0,1)
# Simulación normal de los rendimientos
Ri_dt = pd.DataFrame(mu_R*dt+Z*sigma_R*np.sqrt(dt),index=dates)
Ri_dt.cumprod()
# +
# Simulación del precio
S_0 = closes_aapl.iloc[-1,0]
S_T = S_0*(1+Ri_dt).cumprod()
# Se muestran los precios simulados con los precios descargados
pd.concat([closes_aapl,S_T]).plot(figsize=(8,6));
plt.title('Simulación de precios usando rendimiento simple');
# -
# - **Rendimiento logarítmico**
# +
ri = calc_daily_ret(closes_aapl)
# Usando la media y desviación estándar de los rendimientos logarítmicos
mu_r = ri.mean()[0]
sigma_r = ri.std()[0]
# # Usando la equivalencia teórica
# mu_r2 = mu_R - (sigma_R**2)/2
sim_ret_ri = pd.DataFrame(mu_r*dt+Z*sigma_r*np.sqrt(dt), index=dates)
# Simulación del precio
S_0 = closes_aapl.iloc[-1,0]
S_T2 = S_0*np.exp(sim_ret_ri.cumsum())
# Se muestran los precios simulados con los precios descargados
pd.concat([closes_aapl,S_T2]).plot(figsize=(8,6));
plt.title('Simulación de precios usando rendimiento logarítmico');
# from sklearn.metrics import mean_absolute_error
e1 = np.abs(S_T-S_T2).mean().mean()
e1
# -
print('Las std usando rendimientos logarítmicos y simples son iguales')
sigma_R,sigma_r
# ## 2. Valuación usando simulación: modelo normal para los rendimientos
# - Hallar media y desviación estándar muestral de los rendimientos logarítmicos
mu = ret_aapl.mean()[0]
sigma = ret_aapl.std()[0]
mu, sigma
# No se toma la media sino la tasa libre de riesgo
# > Referencia: https://www.treasury.gov/resource-center/data-chart-center/interest-rates/Pages/TextView.aspx?data=yield
# Tasa de bonos de 1 yr de fecha 11/01/19 -> 1.53%
r = 0.0153/360 # Tasa diaria
# - Simularemos el tiempo de contrato (días=109) desde 2019-11-12 hasta 2020-02-29, 10 escenarios:
# > Calculador de fechas: https://es.calcuworld.com/calendarios/calculadora-de-tiempo-entre-dos-fechas/
#
# - Generar fechas
ndays = 108
nscen = 10
dates = pd.date_range(start='2019-11-14', periods = ndays)
dates
# - Generamos 10 escenarios de rendimientos simulados y guardamos en un dataframe
sim_ret = pd.DataFrame(sigma*np.random.randn(ndays,nscen)+r, index=dates)
sim_ret.cumsum()
# Las columnas son los escenarios y las filas son las días de contrato
# - Con los rendimientos simulados, calcular los escenarios de precios respectivos:
S0 = closes_aapl.iloc[-1,0] # Condición inicial del precio a simular
sim_closes = S0*np.exp(sim_ret.cumsum())
sim_closes
# - Graficar:
# +
# sim_closes.plot(figsize=(8,6));
# -
# Se muestran los precios simulados con los precios descargados
pd.concat([closes_aapl,sim_closes]).plot(figsize=(8,6));
opcion_valuar['JSON']
sigma = 0.27719839019775383/np.sqrt(252)
sigma
# +
from datetime import date
Hoy = date.today()
K=240 # strike price
ndays = 108
nscen = 100000
dates = pd.date_range(start= Hoy, periods = ndays)
S0 = closes_aapl.iloc[-1,0] # Condición inicial del precio a simular
sim_ret = pd.DataFrame(sigma*np.random.randn(ndays,nscen)+r,index=dates)
sim_closes = S0*np.exp(sim_ret.cumsum())
#strike = pd.DataFrame({'Strike':K*np.ones(ndays)}, index=dates)
#simul = pd.concat([closes_aapl.T,strike.T,sim_closes.T]).T
#simul.plot(figsize=(8,6),legend=False);
# -
strike = pd.DataFrame(K*np.ones([ndays,nscen]), index=dates)
call = pd.DataFrame({'Prima':np.exp(-r*ndays) \
*np.fmax(sim_closes-strike,np.zeros([ndays,nscen])).mean(axis=1)}, index=dates)
call.plot();
# La valuación de la opción es:
call.iloc[-1]
# Intervalo de confianza del 99%
confianza = 0.99
sigma_est = sim_closes.iloc[-1].sem()
mean_est = call.iloc[-1].Prima
i1 = st.t.interval(confianza,nscen-1, loc=mean_est, scale=sigma_est)
i2 = st.norm.interval(confianza, loc=mean_est, scale=sigma_est)
print(i1)
print(i1)
# ## Precios simulados usando técnicas de reducción de varianza
# +
# Usando muestreo estratificado----> #estratros = nscen
U = (np.arange(0,nscen)+np.random.rand(ndays,nscen))/nscen
Z = st.norm.ppf(U)
sim_ret2 = pd.DataFrame(sigma*Z+r,index=dates)
sim_closes2 = S0*np.exp(sim_ret.cumsum())
# Función de pago
strike = pd.DataFrame(K*np.ones([ndays,nscen]), index=dates)
call = pd.DataFrame({'Prima':np.exp(-r*ndays) \
*np.fmax(sim_closes2-strike,np.zeros([ndays,nscen])).T.mean()}, index=dates)
call.plot();
# -
# La valuación de la opción es:
call.iloc[-1]
# Intervalo de confianza del 99%
confianza = 0.99
sigma_est = sim_closes2.iloc[-1].sem()
mean_est = call.iloc[-1].Prima
i1 = st.t.interval(confianza,nscen-1, loc=mean_est, scale=sigma_est)
i2 = st.norm.interval(confianza, loc=mean_est, scale=sigma_est)
print(i1)
print(i1)
# ### Análisis de la distribución de los rendimientos
# ### Ajustando norm
# +
ren = calc_daily_ret(closes_aapl) # rendimientos
y,x,des = plt.hist(ren['AAPL'],bins=50,density=True,label='Histograma rendimientos')
mu_fit,sd_fit = st.norm.fit(ren) # Se ajustan los parámetros de una normal
# Valores máximo y mínimo de los rendiemientos a generar
ren_max = max(x);ren_min = min(x)
# Vector de rendimientos generados
ren_gen = np.arange(ren_min,ren_max,0.001)
# Generación de la normal ajustado con los parámetros encontrados
curve_fit = st.norm.pdf(ren_gen,loc=mu_fit,scale=sd_fit)
plt.plot(ren_gen,curve_fit,label='Distribución ajustada')
plt.legend()
plt.show()
# -
# ### Ajustando t
# +
ren = calc_daily_ret(closes_aapl) # rendimientos
y,x,des = plt.hist(ren['AAPL'],bins=50,density=True,label='Histograma rendimientos')
dof,mu_fit,sd_fit = st.t.fit(ren) # Se ajustan los parámetros de una normal
# Valores máximo y mínimo de los rendiemientos a generar
# ren_max = max(x);ren_min = min(x)
# Vector de rendimientos generados
ren_gen = np.arange(ren_min,ren_max,0.001)
# Generación de la normal ajustado con los parámetros encontrados
curve_fit = st.t.pdf(ren_gen,df=dof,loc=mu_fit,scale=sd_fit)
plt.plot(ren_gen,curve_fit,label='Distribución ajustada')
plt.legend()
plt.show()
# -
st.probplot(ren['AAPL'],sparams= dof, dist='t', plot=plt);
# ## 3. Valuación usando simulación: uso del histograma de rendimientos
#
# Todo el análisis anterior se mantiene. Solo cambia la forma de generar los números aleatorios para la simulación montecarlo.
#
# Ahora, generemos un histograma de los rendimientos diarios para generar valores aleatorios de los rendimientos simulados.
# - Primero, cantidad de días y número de escenarios de simulación
ndays = 109
nscen = 10
# - Del histograma anterior, ya conocemos las probabilidades de ocurrencia, lo que se llamó como variable `y`
prob = y/np.sum(y)
values = x[1:]
# - Con esto, generamos los números aleatorios correspondientes a los rendimientos (tantos como días por número de escenarios).
ret = np.random.choice(values, ndays*nscen, p=prob)
dates = pd.date_range(start=Hoy,periods=ndays)
sim_ret_hist = pd.DataFrame(ret.reshape((ndays,nscen)),index=dates)
sim_ret_hist
sim_closes_hist = (closes_aapl.iloc[-1,0])*np.exp(sim_ret_hist.cumsum())
sim_closes_hist
sim_closes_hist.plot(figsize=(8,6),legend=False);
pd.concat([closes_aapl,sim_closes_hist]).plot(figsize=(8,6),legend=False);
plt.title('Simulación usando el histograma de los rendimientos')
K=240
ndays = 109
nscen = 10000
freq, values = np.histogram(ret_aapl+r-mu, bins=2000)
prob = freq/np.sum(freq)
ret=np.random.choice(values[1:],ndays*nscen,p=prob)
dates=pd.date_range('2018-10-29',periods=ndays)
sim_ret_hist = pd.DataFrame(ret.reshape((ndays,nscen)),index=dates)
sim_closes_hist = (closes_aapl.iloc[-1,0])*np.exp(sim_ret_hist.cumsum())
strike = pd.DataFrame(K*np.ones(ndays*nscen).reshape((ndays,nscen)), index=dates)
call_hist = pd.DataFrame({'Prima':np.exp(-r*ndays)*np.fmax(sim_closes_hist-strike,np.zeros(ndays*nscen).reshape((ndays,nscen))).T.mean()}, index=dates)
call_hist.plot();
call_hist.iloc[-1]
opcion_valuar['JSON']
# Intervalo de confianza del 95%
confianza = 0.95
sigma_est = sim_closes_hist.iloc[-1].sem()
mean_est = call_hist.iloc[-1].Prima
i1 = st.t.interval(confianza,nscen-1, loc=mean_est, scale=sigma_est)
i2 = st.norm.interval(confianza, loc=mean_est, scale=sigma_est)
print(i1)
print(i1)
# # <font color = 'red'> Tarea: </font>
#
# Replicar el procedimiento anterior para valoración de opciones 'call', pero en este caso para opciones tipo 'put'.
# <script>
# $(document).ready(function(){
# $('div.prompt').hide();
# $('div.back-to-top').hide();
# $('nav#menubar').hide();
# $('.breadcrumb').hide();
# $('.hidden-print').hide();
# });
# </script>
#
# <footer id="attribution" style="float:right; color:#808080; background:#fff;">
# Created with Jupyter by <NAME> and modify by <NAME>.
# </footer>
| TEMA-3/Clase19_ValuacionOpciones.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Why Python is Slow: Looking Under the Hood
# *This notebook originally appeared as a blog post by <NAME> on [Pythonic Perambulations](http://jakevdp.github.io). The content is BSD-licensed.*
# We've all heard it before: Python is slow.
#
# When I teach courses on Python for scientific computing, I make this point [very early](http://nbviewer.ipython.org/github/jakevdp/2013_fall_ASTR599/blob/master/notebooks/11_EfficientNumpy.ipynb) in the course, and tell the students why: it boils down to Python being a dynamically typed, interpreted language, where values are stored not in dense buffers but in scattered objects. And then I talk about how to get around this by using NumPy, SciPy, and related tools for vectorization of operations and calling into compiled code, and go on from there.
#
# But I realized something recently: despite the relative accuracy of the above statements, the words "dynamically-typed-interpreted-buffers-vectorization-compiled" probably mean very little to somebody attending an intro programming seminar. The jargon does little to enlighten people about what's actually going on "under the hood", so to speak.
#
# So I decided I would write this post, and dive into the details that I usually gloss over. Along the way, we'll take a look at using Python's standard library to introspect the goings-on of CPython itself. So whether you're a novice or experienced programmer, I hope you'll learn something from the following exploration.
# ## Why Python is Slow
# Python is slower than Fortran and C for a variety of reasons:
# ### 1. **Python is Dynamically Typed rather than Statically Typed**.
#
# What this means is that at the time the program executes, the interpreter doesn't know the type of the variables that are defined. The difference between a C variable (I'm using C as a stand-in for compiled languages) and a Python variable is summarized by this diagram:
#
# 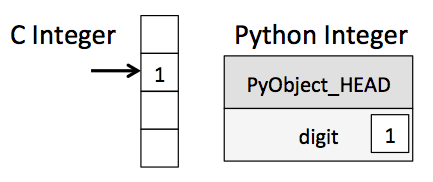
#
# For a variable in C, the compiler knows the type by its very definition. For a variable in Python, all you know at the time the program executes is that it's some sort of Python object.
#
# So if you write the following in C:
#
# ``` C
# /* C code */
# int a = 1;
# int b = 2;
# int c = a + b;
# ```
#
# the C compiler knows from the start that ``a`` and ``b`` are integers: they simply can't be anything else! With this knowledge, it can call the routine which adds two integers, returning another integer which is just a simple value in memory. As a rough schematic, the sequence of events looks like this:
#
# #### C Addition
#
# 1. Assign ``<int> 1`` to ``a``
# 2. Assign ``<int> 2`` to ``b``
# 3. call ``binary_add<int, int>(a, b)``
# 4. Assign the result to c
#
# The equivalent code in Python looks like this:
#
# ``` python
# # python code
# a = 1
# b = 2
# c = a + b
# ```
#
# here the interpreter knows only that ``1`` and ``2`` are objects, but not what type of object they are. So the The interpreter must inspect ``PyObject_HEAD`` for each variable to find the type information, and then call the appropriate summation routine for the two types. Finally it must create and initialize a new Python object to hold the return value. The sequence of events looks roughly like this:
#
# #### Python Addition
#
# 1. Assign ``1`` to ``a``
#
# - **1a.** Set ``a->PyObject_HEAD->typecode`` to integer
# - **1b.** Set ``a->val = 1``
#
# 2. Assign ``2`` to ``b``
#
# - **2a.** Set ``b->PyObject_HEAD->typecode`` to integer
# - **2b.** Set ``b->val = 2``
#
# 3. call ``binary_add(a, b)``
#
# - **3a.** find typecode in ``a->PyObject_HEAD``
# - **3b.** ``a`` is an integer; value is ``a->val``
# - **3c.** find typecode in ``b->PyObject_HEAD``
# - **3d.** ``b`` is an integer; value is ``b->val``
# - **3e.** call ``binary_add<int, int>(a->val, b->val)``
# - **3f.** result of this is ``result``, and is an integer.
#
# 4. Create a Python object ``c``
#
# - **4a.** set ``c->PyObject_HEAD->typecode`` to integer
# - **4b.** set ``c->val`` to ``result``
#
# The dynamic typing means that there are a lot more steps involved with any operation. This is a primary reason that Python is slow compared to C for operations on numerical data.
# ### 2. Python is interpreted rather than compiled.
#
# We saw above one difference between interpreted and compiled code. A smart compiler can look ahead and optimize for repeated or unneeded operations, which can result in speed-ups. Compiler optimization is its own beast, and I'm personally not qualified to say much about it, so I'll stop there. For some examples of this in action, you can take a look at my [previous post](http://jakevdp.github.io/blog/2013/06/15/numba-vs-cython-take-2/) on Numba and Cython.
# ### 3. Python's object model can lead to inefficient memory access
#
# We saw above the extra type info layer when moving from a C integer to a Python integer. Now imagine you have many such integers and want to do some sort of batch operation on them. In Python you might use the standard ``List`` object, while in C you would likely use some sort of buffer-based array.
#
# A NumPy array in its simplest form is a Python object build around a C array. That is, it has a pointer to a *contiguous* data buffer of values. A Python list, on the other hand, has a pointer to a contiguous buffer of pointers, each of which points to a Python object which in turn has references to its data (in this case, integers). This is a schematic of what the two might look like:
#
# 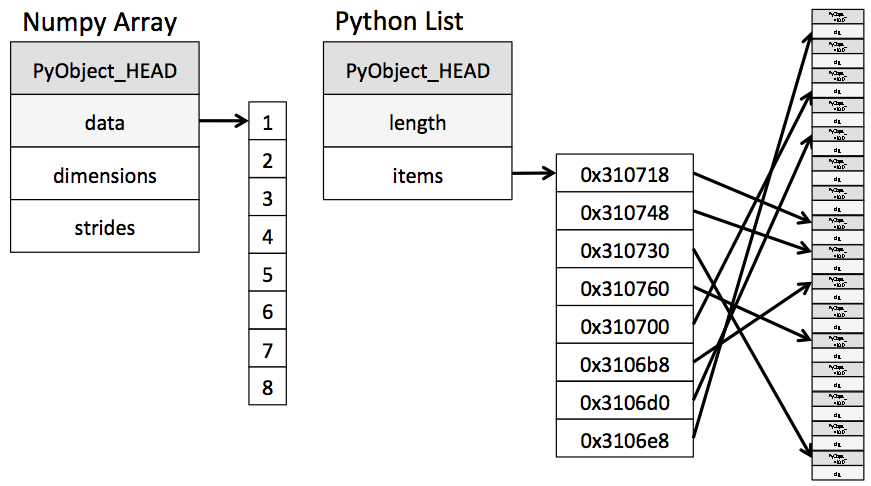
#
# It's easy to see that if you're doing some operation which steps through data in sequence, the numpy layout will be much more efficient than the Python layout, both in the cost of storage and the cost of access.
# ### So Why Use Python?
#
# Given this inherent inefficiency, why would we even think about using Python? Well, it comes down to this: Dynamic typing makes Python **easier to use** than C. It's extremely **flexible and forgiving**, this flexibility leads to **efficient use of development time**, and on those occasions that you really need the optimization of C or Fortran, **Python offers easy hooks into compiled libraries**. It's why Python use within many scientific communities has been continually growing. With all that put together, Python ends up being an extremely efficient language for the overall task of doing science with code.
# ## Python meta-hacking: Don't take my word for it
# Above I've talked about some of the internal structures that make Python tick, but I don't want to stop there. As I was putting together the above summary, I started hacking around on the internals of the Python language, and found that the process itself is pretty enlightening.
#
# In the following sections, I'm going to *prove* to you that the above information is correct, by doing some hacking to expose Python objects using Python itself. Please note that everything below is written using **Python 3.4**. Earlier versions of Python have a slightly different internal object structure, and later versions may tweak this further. Please make sure to use the correct version! Also, most of the code below assumes a 64-bit CPU. If you're on a 32-bit platform, some of the C types below will have to be adjusted to account for this difference.
import sys
print("Python version =", sys.version[:5])
# ### Digging into Python Integers
# Integers in Python are easy to create and use:
x = 42
print(x)
# But the simplicity of this interface belies the complexity of what is happening under the hood. We briefly discussed the memory layout of Python integers above. Here we'll use Python's built-in ``ctypes`` module to introspect Python's integer type from the Python interpreter itself. But first we need to know exactly what a Python integer looks like at the level of the C API.
#
# The actual ``x`` variable in CPython is stored in a structure which is defined in the CPython source code, in [Include/longintrepr.h](http://hg.python.org/cpython/file/3.4/Include/longintrepr.h/#l89)
#
# ``` C
# struct _longobject {
# PyObject_VAR_HEAD
# digit ob_digit[1];
# };
# ```
#
# The ``PyObject_VAR_HEAD`` is a macro which starts the object off with the following struct, defined in [Include/object.h](http://hg.python.org/cpython/file/3.4/Include/object.h#l111):
#
# ``` C
# typedef struct {
# PyObject ob_base;
# Py_ssize_t ob_size; /* Number of items in variable part */
# } PyVarObject;
# ```
#
# ... and includes a ``PyObject`` element, which is also defined in [Include/object.h](http://hg.python.org/cpython/file/3.4/Include/object.h#l105):
#
# ``` C
# typedef struct _object {
# _PyObject_HEAD_EXTRA
# Py_ssize_t ob_refcnt;
# struct _typeobject *ob_type;
# } PyObject;
# ```
#
# here ``_PyObject_HEAD_EXTRA`` is a macro which is not normally used in the Python build.
#
# With all this put together and typedefs/macros unobfuscated, our integer object works out to something like the following structure:
#
# ``` C
# struct _longobject {
# long ob_refcnt;
# PyTypeObject *ob_type;
# size_t ob_size;
# long ob_digit[1];
# };
# ```
#
# The ``ob_refcnt`` variable is the reference count for the object, the ``ob_type`` variable is a pointer to the structure containing all the type information and method definitions for the object, and the ``ob_digit`` holds the actual numerical value.
#
# Armed with this knowledge, we'll use the ``ctypes`` module to start looking into the actual object structure and extract some of the above information.
#
# We start with defining a Python representation of the C structure:
# +
import ctypes
class IntStruct(ctypes.Structure):
_fields_ = [("ob_refcnt", ctypes.c_long),
("ob_type", ctypes.c_void_p),
("ob_size", ctypes.c_ulong),
("ob_digit", ctypes.c_long)]
def __repr__(self):
return ("IntStruct(ob_digit={self.ob_digit}, "
"refcount={self.ob_refcnt})").format(self=self)
# -
# Now let's look at the internal representation for some number, say 42. We'll use the fact that in CPython, the ``id`` function gives the memory location of the object:
num = 42
IntStruct.from_address(id(42))
# The ``ob_digit`` attribute points to the correct location in memory!
#
# But what about ``refcount``? We've only created a single value: why is the reference count so much greater than one?
#
# Well it turns out that Python uses small integers *a lot*. If a new ``PyObject`` were created for each of these integers, it would take a lot of memory. Because of this, Python implements common integer values as **singletons**: that is, only one copy of these numbers exist in memory. In other words, every time you create a new Python integer in this range, you're simply creating a reference to the singleton with that value:
x = 42
y = 42
id(x) == id(y)
# Both variables are simply pointers to the same memory address. When you get to much bigger integers (larger than 255 in Python 3.4), this is no longer true:
x = 1234
y = 1234
id(x) == id(y)
# Just starting up the Python interpreter will create a lot of integer objects; it can be interesting to take a look at how many references there are to each:
# %matplotlib inline
import matplotlib.pyplot as plt
import sys
plt.loglog(range(1000), [sys.getrefcount(i) for i in range(1000)])
plt.xlabel('integer value')
plt.ylabel('reference count')
# We see that zero is referenced several thousand times, and as you may expect, the frequency of references generally decreases as the value of the integer increases.
#
# Just to further make sure that this is behaving as we'd expect, let's make sure the ``ob_digit`` field holds the correct value:
all(i == IntStruct.from_address(id(i)).ob_digit
for i in range(256))
# If you go a bit deeper into this, you might notice that this does not hold for numbers larger than 256: it turns out that some bit-shift gymnastics are performed in [Objects/longobject.c](http://hg.python.org/cpython/file/3.4/Objects/longobject.c#l232), and these change the way large integers are represented in memory.
#
# I can't say that I fully understand why exactly that is happening, but I imagine it has something to do with Python's ability to efficiently handle integers past the overflow limit of the long int data type, as we can see here:
2 ** 100
# That number is much too long to be a ``long``, which can only hold 64 bits worth of values (that is, up to $\sim2^{64}$)
# ### Digging into Python Lists
# Let's apply the above ideas to a more complicated type: Python lists. Analogously to integers, we find the definition of the list object itself in [Include/listobject.h](http://hg.python.org/cpython/file/3.4/Include/listobject.h#l23):
#
# ``` C
# typedef struct {
# PyObject_VAR_HEAD
# PyObject **ob_item;
# Py_ssize_t allocated;
# } PyListObject;
# ```
#
# Again, we can expand the macros and de-obfuscate the types to see that the structure is effectively the following:
#
# ``` C
# typedef struct {
# long ob_refcnt;
# PyTypeObject *ob_type;
# Py_ssize_t ob_size;
# PyObject **ob_item;
# long allocated;
# } PyListObject;
# ```
#
# Here the ``PyObject **ob_item`` is what points to the contents of the list, and the ``ob_size`` value tells us how many items are in the list.
class ListStruct(ctypes.Structure):
_fields_ = [("ob_refcnt", ctypes.c_long),
("ob_type", ctypes.c_void_p),
("ob_size", ctypes.c_ulong),
("ob_item", ctypes.c_long), # PyObject** pointer cast to long
("allocated", ctypes.c_ulong)]
def __repr__(self):
return ("ListStruct(len={self.ob_size}, "
"refcount={self.ob_refcnt})").format(self=self)
# Let's try it out:
L = [1,2,3,4,5]
ListStruct.from_address(id(L))
# Just to make sure we've done things correctly, let's create a few extra references to the list, and see how it affects the reference count:
tup = [L, L] # two more references to L
ListStruct.from_address(id(L))
# Now let's see about finding the actual elements within the list.
#
# As we saw above, the elements are stored via a contiguous array of ``PyObject`` pointers. Using ``ctypes``, we can actually create a compound structure consisting of our ``IntStruct`` objects from before:
# +
# get a raw pointer to our list
Lstruct = ListStruct.from_address(id(L))
# create a type which is an array of integer pointers the same length as L
PtrArray = Lstruct.ob_size * ctypes.POINTER(IntStruct)
# instantiate this type using the ob_item pointer
L_values = PtrArray.from_address(Lstruct.ob_item)
# -
# Now let's take a look at the values in each of the items:
[ptr[0] for ptr in L_values] # ptr[0] dereferences the pointer
# We've recovered the ``PyObject`` integers within our list! You might wish to take a moment to look back up to the schematic of the List memory layout above, and make sure you understand how these ``ctypes`` operations map onto that diagram.
# ### Digging into NumPy arrays
# Now, for comparison, let's do the same introspection on a numpy array. I'll skip the detailed walk-through of the NumPy C-API array definition; if you want to take a look at it, you can find it in [numpy/core/include/numpy/ndarraytypes.h](https://github.com/numpy/numpy/blob/maintenance/1.8.x/numpy/core/include/numpy/ndarraytypes.h#L646)
#
# Note that I'm using NumPy version 1.8 here; these internals may have changed between versions, though I'm not sure whether this is the case.
import numpy as np
np.__version__
# Let's start by creating a structure that represents the numpy array itself. This should be starting to look familiar...
#
# We'll also add some custom properties to access Python versions of the shape and strides:
class NumpyStruct(ctypes.Structure):
_fields_ = [("ob_refcnt", ctypes.c_long),
("ob_type", ctypes.c_void_p),
("ob_data", ctypes.c_long), # char* pointer cast to long
("ob_ndim", ctypes.c_int),
("ob_shape", ctypes.c_voidp),
("ob_strides", ctypes.c_voidp)]
@property
def shape(self):
return tuple((self.ob_ndim * ctypes.c_int64).from_address(self.ob_shape))
@property
def strides(self):
return tuple((self.ob_ndim * ctypes.c_int64).from_address(self.ob_strides))
def __repr__(self):
return ("NumpyStruct(shape={self.shape}, "
"refcount={self.ob_refcnt})").format(self=self)
# Now let's try it out:
x = np.random.random((10, 20))
xstruct = NumpyStruct.from_address(id(x))
xstruct
# We see that we've pulled out the correct shape information. Let's make sure the reference count is correct:
L = [x,x,x] # add three more references to x
xstruct
# Now we can do the tricky part of pulling out the data buffer. For simplicity we'll ignore the strides and assume it's a C-contiguous array; this could be generalized with a bit of work.
# +
x = np.arange(10)
xstruct = NumpyStruct.from_address(id(x))
size = np.prod(xstruct.shape)
# assume an array of integers
arraytype = size * ctypes.c_long
data = arraytype.from_address(xstruct.ob_data)
[d for d in data]
# -
# The ``data`` variable is now a view of the contiguous block of memory defined in the NumPy array! To show this, we'll change a value in the array...
x[4] = 555
[d for d in data]
# ... and observe that the data view changes as well. Both ``x`` and ``data`` are pointing to the same contiguous block of memory.
#
# Comparing the internals of the Python list and the NumPy ndarray, it is clear that NumPy's arrays are **much, much** simpler for representing a list of identically-typed data. That fact is related to what makes it more efficient for the compiler to handle as well.
# ## Just for fun: a few "never use these" hacks
# Using ``ctypes`` to wrap the C-level data behind Python objects allows you to do some pretty interesting things. With proper attribution to my friend <NAME>, I'll say it here: [seriously, don't use this code](http://seriously.dontusethiscode.com/). While nothing below should actually be used (ever), I still find it all pretty interesting!
# ### Modifying the Value of an Integer
# Inspired by [this Reddit post](http://www.reddit.com/r/Python/comments/2441cv/can_you_change_the_value_of_1/), we can actually modify the numerical value of integer objects! If we use a common number like ``0`` or ``1``, we're very likely to crash our Python kernel. But if we do it with less important numbers, we can get away with it, at least briefly.
#
# Note that this is a *really, really* bad idea. In particular, if you're running this in an IPython notebook, you might corrupt the IPython kernel's very ability to run (because you're screwing with the variables in its runtime). Nevertheless, we'll cross our fingers and give it a shot:
# +
# WARNNG: never do this!
id113 = id(113)
iptr = IntStruct.from_address(id113)
iptr.ob_digit = 4 # now Python's 113 contains a 4!
113 == 4
# -
# But note now that we can't set the value back in a simple manner, because the true value ``113`` no longer exists in Python!
113
112 + 1
# One way to recover is to manipulate the bytes directly. We know that $113 = 7 \times 16^1 + 1 * 16^0$, so **on a little-endian 64-bit system running Python 3.4**, the following should work:
ctypes.cast(id113, ctypes.POINTER(ctypes.c_char))[3 * 8] = b'\x71'
112 + 1
# and we're back!
#
# Just in case I didn't stress it enough before: **never do this.**
# ### In-place Modification of List Contents
# Above we did an in-place modification of a value in a numpy array. This is easy, because a numpy array is simply a data buffer. But might we be able to do the same thing for a list? This gets a bit more tricky, because lists store *references* to values rather than the values themselves. And to not crash Python itself, you need to be very careful to keep track of these reference counts as you muck around. Here's how it can be done:
# +
# WARNING: never do this!
L = [42]
Lwrapper = ListStruct.from_address(id(L))
item_address = ctypes.c_long.from_address(Lwrapper.ob_item)
print("before:", L)
# change the c-pointer of the list item
item_address.value = id(6)
# we need to update reference counts by hand
IntStruct.from_address(id(42)).ob_refcnt -= 1
IntStruct.from_address(id(6)).ob_refcnt += 1
print("after: ", L)
# -
# Like I said, you should never use this, and I honestly can't think of any reason why you would want to. But it gives you an idea of the types of operations the interpreter has to do when modifying the contents of a list. Compare this to the NumPy example above, and you'll see one reason why Python lists have more overhead than Python arrays.
# ### Meta Goes Meta: a self-wrapping Python object
# Using the above methods, we can start to get even stranger. The ``Structure`` class in ``ctypes`` is itself a Python object, which can be seen in [Modules/_ctypes/ctypes.h](http://hg.python.org/cpython/file/3.4/Modules/_ctypes/ctypes.h#l46). Just as we wrapped ints and lists, we can wrap structures themselves as follows:
class CStructStruct(ctypes.Structure):
_fields_ = [("ob_refcnt", ctypes.c_long),
("ob_type", ctypes.c_void_p),
("ob_ptr", ctypes.c_long), # char* pointer cast to long
]
def __repr__(self):
return ("CStructStruct(ptr=0x{self.ob_ptr:x}, "
"refcnt={self.ob_refcnt})").format(self=self)
# Now we'll attempt to make a structure that wraps itself. We can't do this directly, because we don't know at what address in memory the new structure will be created. But what we can do is create a *second* structure wrapping the first, and use this to modify its contents in-place!
#
# We'll start by making a temporary meta-structure and wrapping it:
# +
tmp = IntStruct.from_address(id(0))
meta = CStructStruct.from_address(id(tmp))
print(repr(meta))
# -
# Now we add a third structure, and use it to adjust the memory value of the second in-place:
# +
meta_wrapper = CStructStruct.from_address(id(meta))
meta_wrapper.ob_ptr = id(meta)
print(meta.ob_ptr == id(meta))
print(repr(meta))
# -
# We now have a self-wrapping Python structure!
#
# Again, I can't think of any reason you'd ever want to do this. And keep in mind there is noting groundbreaking about this type of self-reference in Python – due to its dynamic typing, it is realatively straightforward to do things like this without directly hacking the memory:
L = []
L.append(L)
print(L)
# ## Conclusion
# Python is slow. And one big reason for that, as we've seen, is the type indirection under the hood which makes Python quick, easy, and fun for the developer. And as we've seen, Python itself offers tools that can be used to hack into the Python objects themselves.
#
# I hope that this was made more clear through this exploration of the differences between various objects, and some liberal mucking around in the internals of CPython itself. This exercise was extremely enlightening for me, and I hope it was for you as well... Happy hacking!
# *This blog post was written entirely in the IPython Notebook. The full notebook can be downloaded
# [here](http://jakevdp.github.io/downloads/notebooks/WhyPythonIsSlow.ipynb),
# or viewed statically
# [here](http://nbviewer.ipython.org/url/jakevdp.github.io/downloads/notebooks/WhyPythonIsSlow.ipynb).*
| output/downloads/notebooks/WhyPythonIsSlow.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Linear regression with Professor Mittens, a.k.a. recipe for linear regression.
#
# ## Overview
#
# In this notebook we will learn how to use regression to study the factors that affect the number of pats cats will recieve. This will start with a visual inspection of the data, followed by the development of a linear model to explain the data. Along the way we will answer a few questions such as: does coat colour influence the number of pats, is a long coat better than a short coat, and how important is the volume of a meow.
#
# ## Specifying regression models
#
# A very popular way to describe regression models is with "formulae" as popularised by R. The [R documentation on formulae](https://cran.r-project.org/doc/manuals/R-intro.html#Formulae-for-statistical-models) is a good place to learn how to use these properly. For example, here is the syntax we will use today,
#
# - `y ~ x1 + x2` will make a linear model with the predictors $x_1$ and $x_2$.
# - `y ~ x1 * x2` includes the terms $x_1 + x_2 + x_1x_2$
# - `y ~ x1 : x2` includes *just* the interaction term $x_1x_2$
# - `y ~ C(x)` specifies that $x$ is a catagorical variable **NOTE** this is not necessary in R.
# %matplotlib inline
import pandas as pd
import numpy as np
import statsmodels.api as sm
import statsmodels.formula.api as smf
import altair as alt
from functools import reduce
# ## Helping cats get more pats
#
# Professor Mittens in interested in helping cats optimise the number of pats they can get. To learn more about this, he has interviewed 1000 cats and taken measurements of their behaviour and appearance. The data in `cat-pats.csv` contains measurments of the following:
#
# - `time_outdoors` is the number of hours that the cat is out of their primary dwelling,
# - `coat_colour` is either tortoiseshell, white, or "other" encoded as integers 1, 2, and 3 respectively,
# - `weight` is the weight of the cat in kilograms,
# - `height` is their height in centimeters,
# - `loudness` is a measure of how loud their meow is, the units are not known,
# - `whisker_length` is the length of their whiskers in centimeters,
# - `is_longhaired` is a Boolean variable equal to 1 if the cat is of a longhaired breed and 0 if it is of a shorthaired breed,
# - `coat_length` is the length of their fur in centimeters,
# - and `num_pats` is the number of pats they received on the day they were interviewed.
#
# The variable we are interested in explaining is `num_pats`. Although this is a discrete variable, we will ignore this aspect of the data and consider it as a continuous value. This is a useful simplifying assumption, as you learn more about regression, in particular generalized linear models, you will see additional ways to handle this. For this example, you can consider it a continuous variable though.
#
# The types of questions that Professor Mittens is interested in answering are as follows:
#
# 1. Do any of the variables correlate with the number of pats that the cats recieve?
# 2. Under a naive model, how much of the variability in pats can they explain? Do all the variables need to be included?
# 3. Does the coat colour matter?
# 4. Among short-haired breeds they say longer hair is better, among long-haired breeds they say short hair is better, who is correct?
# 5. **If a cat can choose to spend more time outdoors, or practise meowing louder, which will get them more pats?**
# ### Read in the data and generate some scatter plots to see if there are any good predictors of the number of pats
#
# The data is in the file `cat-pats.csv` so read this into a data frame using `pd.read_csv` and go from there. I have used altair to generate my scatter plots based on [this example](https://altair-viz.github.io/gallery/scatter_matrix.html) but you can use whatever you feel most comfortable with. It might be useful to use colour to see if `coat_colour` and `is_longhaired` are important.
#
# ### Question
#
# Based on these figures, what variables appear to relate to the number of pats? What do you notice about the catagorical variables `coat_colour` and `is_longhaired`?
cat_df = pd.read_csv("cat-pats.csv")
cat_df
# +
col_names = cat_df.columns.tolist()
predictor_names = col_names.copy()
predictor_names.remove("num_pats")
alt.Chart(cat_df).mark_circle().encode(
alt.X(alt.repeat("column"), type='quantitative', scale=alt.Scale(zero=False)),
alt.Y(alt.repeat("row"), type='quantitative', scale=alt.Scale(zero=False)),
color = "is_longhaired:O" #O here stands for Ordinal since longhaired is ordinal; N would be Nominal
).properties(
width=100,
height=100
).repeat(
row=["num_pats"],
column=predictor_names
)
# When you have continuous variables, plot here like this, but with categorical, good to use colour, which is why he did that
# +
# import matplotlib.pyplot as plt
# import seaborn as sns
# sns.FacetGrid(cat_df,col=predictor_names,row='num_pats')
# -
# ### Compute the correlation between each variable and the number of pats, what looks important
#
# ### Question
#
# Does the the correlation matrix raise any further questions? Does it handle the catagorical variables correctly?
# +
cat_df.corr(method='pearson')
# just focus on num_pats; good way to pre-select variables since it's a pain to make a million plots to see
# -
# Yennie: Also, can do heat map of this to visualize this!
sns.heatmap(cat_df.corr())
# +
import seaborn as sns
#Hannah: If anyone’s interested, seaboarn has a nice easy function for plotting correlation maps
# I posted some code above with some additional arguments, copied from this source (https://seaborn.pydata.org/examples/many_pairwise_correlations.html). Has some nice additional functionality if you need it in other work :)
# sns.heatmap(corr, vmax=.3, center=0,
# square=True, linewidths=.5,
# cbar_kws={"shrink": .5})
corr = cat_df.corr()
sns.heatmap(corr, annot = True, vmax=.3, center=0,
square=True, linewidths=.5,
cbar_kws={"shrink": .5})
# -
#Cameron: Pairplot in seaboarn can be done with
sns.pairplot(cat_df,hue="is_longhaired",y_vars='num_pats',x_vars=predictor_names)
# ### What is $R^2$?
#
# Sometimes called the *coefficient of determination*, this statistic measures the proportion of the variance in the response variable that is explained by the regression model. In the case of simple linear regression it is just the correlation squared, it can also be calculated as the ratio of the regression sum of squares and the total sum of squares.
#
# $$
# R^2 = \frac{\text{RegSS}}{\text{TSS}}
# $$
#
# It can be thought of as the proportion of the total variance that is explained by the regression model.
#
# ### What is an *adjusted* $R^2$?
#
# For a fixed number of observations, as the number of covariates increases you can get explain as much of the variability as you want! The adjusted $R^2$ is a way to penalise using too many covariates. The adjusted $R^2$ for a model with $n$ observations and $p$ coefficients is given by the following:
#
# $$
# \tilde{R}^2 = 1 - \frac{n - 1}{n - p}\left(1 - R^2\right)
# $$
#
# ### Under a naive model, how much of the variability in pats can they explain?
#
# Run an ordinary linear regression with all of the variables and see what percentage of the variability in the number of pats is explained. Make sure that you have used the catagorical variables correctly. Can be be confident in rejecting the null hypothesis that none of these variables is associated with the number of pats received?
cat_df
lm_xw = smf.ols("num_pats ~ time_outdoors + weight + height + loudness + C(coat_colour) + coat_length + whisker_length + C(is_longhaired)",cat_df).fit()
print(lm_xw.summary())
lm_1 = smf.ols("num_pats ~ time_outdoors + C(coat_colour) + weight + height + loudness + whisker_length + C(is_longhaired) + coat_length", cat_df).fit()
print(lm_1.summary())
# **Note:** f-statistic is used to test the null hypothesis that none of the variables are related to the num_pats and all coefficients should be zero
#
# Prob f statistic is the p-value of the null model being no relationship between the variables and response
# So in this case we reject the null hypothesis that none of the variables are related to the response
#
# --
#
# If you have coat color 2 you get on avg 2+ more pats than coat color 1, if you have coat color 3 you have on avg 8 more than coat color 1, if you have coat color 2 you
# for coat color 1, it becomes the default value, so the coef for cc1 falls into the intercept
#
# keeping all other things constant if you sampled cats and get two exactly the same except cc, then would be 2 more pats on avg...
#
# @Yennie - I think one can differentiate between impact and significance with the latter being the first sanity check to do. Once you have 2 variables that are significant, you can see which one is bigger to assess their respective contributions to the y variable
#
# coef of coat color is gamma in the lecture.. two different intercepts with the same gradient
#
# PATRICK:
# WE should look at both the correlation and regression output to see if there is an issue of colinearity... don't expect 1 test to reveal all problems
#
# If you reindexed the coat color, the intercept might change but the coef will not
# but if you had a tiny interaction variable, it will change
#
# VIF is a good default minimum test but it is not necessarily going to catch all concerning relationships
#
# Paulius:
# By the way, if you would like to get a separate dummy variable in your pandas dataframe (and just plug in that to the regression equation), such as: coat_color1, coat_color2, coat_color3, you can use this code:
# cat_df2 = pd.concat([cat_df, pd.get_dummies(cat_df['coat_colour'], prefix = 'coat_color')], axis = 1)
# Principle of Marginality -
# when you interpret these variables you have to consider all other things held constant
sm.graphics.influence_plot(lm_1, criterion="cooks")
sm.graphics.plot_fit(lm_1, "coat_length")
cat_df[543:546]
cat_df.describe()
# The variance (of the coefficients) is inflated when there is colinearity...
# ### Question: Is colinearity an issue in this model? Do all of the variables need to be included?
#
# Compute the VIF to see if there is a concerning amount of colinearity between any of the covariates.
# +
col_names = cat_df.copy().columns.tolist()
col_names.remove("num_pats") # we don't use the response variable in VIF calc
def join_strings(xs, sep):
return reduce(lambda a, b: a + sep + b, xs)
for v in col_names:
cns = col_names.copy()
cns.remove(v)
formula = v + " ~ " + join_strings(cns, " + ")
coef_det = smf.ols(formula, data = cat_df).fit()
vif = 1 / (1 - coef_det.rsquared)
if vif > 3:
print("\n" + 80 * "=")
print(v)
print(vif)
# +
# Yacopo for VIF
from statsmodels.stats.outliers_influence import variance_inflation_factor
from statsmodels.tools.tools import add_constant
X = add_constant(df.iloc[:, :-1])
pd.Series([variance_inflation_factor(X.values, i)
for i in range(X.shape[1])],
index=X.columns)
#Yen: Jacopo, what is add_constant?
#Yacopo: apparently you need a column of 1 at the beginning of your dataframe. It's the intercept pretty much
# Goda for VIF
# variables = fitted_naive.model.exog
# names = fitted_naive.model.exog_names
# vif = [variance_inflation_factor(variables, i) for i in range(variables.shape[1])]
# pd.DataFrame(zip(names, vif), columns=["names", 'vif'])
# -
lm_3 = smf.ols("num_pats ~ time_outdoors + C(coat_colour) + weight + height + loudness + C(is_longhaired) + coat_length", cat_df).fit()
print(lm_3.summary())
# pruned away a variable that is not informative because it's been informed by other variables
# can see p-value of loudness and coat_length have decreased....
# ### Does coat colour matter?
#
# 1. Make a box plot of the number of pats by coat colour to see this pattern.
# 2. Fit an additional linear model without the coat colour as a covariate to see how much of the explained variability comes from the inclusion of coat colour in the model.
coat_df = cat_df.loc[:,["coat_colour", "num_pats"]].copy()
alt.Chart(coat_df).mark_boxplot().encode(
x='coat_colour:O',
y='num_pats:Q'
).properties(
width=600,
height=300
)
# +
lm_with_colour = smf.ols("num_pats ~ time_outdoors + C(coat_colour) + weight + height + loudness + C(is_longhaired) + coat_length", cat_df).fit()
lm_without_colour = smf.ols("num_pats ~ time_outdoors + weight + height + loudness + C(is_longhaired) + coat_length", cat_df).fit()
# print(lm_with_colour.summary())
# print(lm_without_colour.summary())
a = (lm_with_colour.resid ** 2).sum()
b = (lm_without_colour.resid ** 2).sum()
print('a ' + str(a))
print('b ' + str(b))
(b - a) / b
# -
# 20% of the explained variance comes from the coat colour...
# +
df = sm.datasets.get_rdataset("Guerry", "HistData").data
df = df[['Lottery', 'Literacy', 'Wealth', 'Region']].dropna()
df['Region'].unique()
# -
# ### Among short-haired breeds they say longer hair is better, among long-haired breeds they say short hair is better, who is correct?
#
# Since in the figures above we saw that the breed longhaired/shorthaired appears to separate the data, it may be useful to consider different models on each subset. Fit a linear model to each subset of the data and see that the effect of the coat length is in each case.
lm_4a = smf.ols("num_pats ~ time_outdoors + C(coat_colour) + weight + height + loudness + coat_length", cat_df[cat_df["is_longhaired"] == 1]).fit()
lm_4b = smf.ols("num_pats ~ time_outdoors + C(coat_colour) + weight + height + loudness + coat_length", cat_df[cat_df["is_longhaired"] == 0]).fit()
print(lm_4a.summary())
print(lm_4b.summary())
# ### Fit a model with *only* the interacion term between the coat length and "is longhaired" and the other covariates
#
# What does this tell us about the age old debate about cat hair length? Why are we ignoring the [Principle of Marginality](https://en.wikipedia.org/wiki/Principle_of_marginality) in this example?
lm_4c = smf.ols("num_pats ~ time_outdoors + C(coat_colour) + weight + height + loudness + C(is_longhaired) : coat_length", cat_df).fit()
print(lm_4c.summary())
# ### How else could we handle coat length?
#
# We could instead have included quadratic terms for coat length to see if this was a better way to explain the non-linear effect.
# ### Shouldn't we check for influential points?
#
# We can generate a plot of the studentized residuals and the leverage to check if there are any influential points.
#
# If there is a potential outlier, does removing it change anything?
# ### Should a cat practise meowing or just spend more time outdoors to get more pats?
#
# We can just look at the coefficients to see that a much more efficient way to get pats is to be outside, the relationship between loudness and number of pats is not supported by this data set.
# You have to interpret this in the units the data was collected
# outliers not necessarily a problem w respect to model fit unless they dominate the value of the coef
| example-2/example-2-questions_EDIT_XW.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **Source of the materials**: Biopython cookbook (adapted)
#Lets load notebook's Image
from IPython.core.display import Image
from reportlab.lib import colors
from reportlab.lib.units import cm
from Bio.Graphics import GenomeDiagram
from Bio import SeqIO
# # Graphics including GenomeDiagram
# The Bio.Graphics module depends on the third party Python library ReportLab. Although focused on producing PDF files, ReportLab can also create encapsulated postscript (EPS) and (SVG) files. In addition to these vector based images, provided certain further dependencies such as the Python Imaging Library (PIL) are installed, ReportLab can also output bitmap images (including JPEG, PNG, GIF, BMP and PICT formats).
# ## GenomeDiagram
# ### Introduction
# The **Bio.Graphics.GenomeDiagram** module was added to Biopython 1.50, having previously been available as a separate Python module dependent on Biopython.
#
# As the name might suggest, GenomeDiagram was designed for drawing whole genomes, in particular prokaryotic genomes, either as linear diagrams (optionally broken up into fragments to fit better) or as circular wheel diagrams. It proved also well suited to drawing quite detailed figures for smaller genomes such as phage, plasmids or mitochrondia.
#
# This module is easiest to use if you have your genome loaded as a SeqRecord object containing lots of SeqFeature objects - for example as loaded from a GenBank file.
# ### Diagrams, tracks, feature-sets and features
# GenomeDiagram uses a nested set of objects. At the top level, you have a diagram object representing a sequence (or sequence region) along the horizontal axis (or circle). A diagram can contain one or more tracks, shown stacked vertically (or radially on circular diagrams). These will typically all have the same length and represent the same sequence region. You might use one track to show the gene locations, another to show regulatory regions, and a third track to show the GC percentage.
#
# The most commonly used type of track will contain features, bundled together in feature-sets. You might choose to use one feature-set for all your CDS features, and another for tRNA features. This isn’t required - they can all go in the same feature-set, but it makes it easier to update the properties of just selected features (e.g. make all the tRNA features red).
#
# There are two main ways to build up a complete diagram. Firstly, the top down approach where you create a diagram object, and then using its methods add track(s), and use the track methods to add feature-set(s), and use their methods to add the features. Secondly, you can create the individual objects separately (in whatever order suits your code), and then combine them.
# ### A top down example
# We’re going to draw a whole genome from a SeqRecord object read in from a GenBank file. This example uses the pPCP1 plasmid from Yersinia pestis biovar Microtus (<a href="data/NC_005816.gb">NC_005816.gb</a>)
record = SeqIO.read("data/NC_005816.gb", "genbank")
# We’re using a top down approach, so after loading in our sequence we next create an empty diagram, then add an (empty) track, and to that add an (empty) feature set:
gd_diagram = GenomeDiagram.Diagram("Yersinia pestis biovar Microtus plasmid pPCP1")
gd_track_for_features = gd_diagram.new_track(1, name="Annotated Features")
gd_feature_set = gd_track_for_features.new_set()
# Now the fun part - we take each gene SeqFeature object in our SeqRecord, and use it to generate a feature on the diagram. We’re going to color them blue, alternating between a dark blue and a light blue.
for feature in record.features:
if feature.type != "gene":
#Exclude this feature
continue
if len(gd_feature_set) % 2 == 0:
color = colors.blue
else:
color = colors.lightblue
gd_feature_set.add_feature(feature, color=color, label=True)
# Now we come to actually making the output file. This happens in two steps, first we call the draw method, which creates all the shapes using ReportLab objects. Then we call the write method which renders these to the requested file format. Note you can output in multiple file formats:
gd_diagram.draw(format="linear", orientation="landscape", pagesize='A4',
fragments=4, start=0, end=len(record))
gd_diagram.write("data/plasmid_linear.png", "png")
# Lets have a look at the previous one:
# <img src="plasmid_linear.png">
# Notice that the fragments argument which we set to four controls how many pieces the genome gets broken up into.
#
# If you want to do a circular figure, then try this:
gd_diagram.draw(format="circular", circular=True, pagesize=(20*cm,20*cm),
start=0, end=len(record), circle_core=0.7)
gd_diagram.write("data/plasmid_circular.png", "PNG")
Image("data/plasmid_circular.png")
# These figures are not very exciting, but we’ve only just got started.
# ### A bottom up example
# Now let’s produce exactly the same figures, but using the bottom up approach. This means we create the different objects directly (and this can be done in almost any order) and then combine them.
# +
record = SeqIO.read("data/NC_005816.gb", "genbank")
#Create the feature set and its feature objects,
gd_feature_set = GenomeDiagram.FeatureSet()
for feature in record.features:
if feature.type != "gene":
#Exclude this feature
continue
if len(gd_feature_set) % 2 == 0:
color = colors.blue
else:
color = colors.lightblue
gd_feature_set.add_feature(feature, color=color, label=True)
#(this for loop is the same as in the previous example)
#Create a track, and a diagram
gd_track_for_features = GenomeDiagram.Track(name="Annotated Features")
gd_diagram = GenomeDiagram.Diagram("Yersinia pestis biovar Microtus plasmid pPCP1")
#Now have to glue the bits together...
gd_track_for_features.add_set(gd_feature_set)
gd_diagram.add_track(gd_track_for_features, 1)
# -
# You can now call the draw and write methods as before to produce a linear or circular diagram, using the code at the end of the top-down example above. The figures should be identical.
# ### Features without a SeqFeature
# In the above example we used a SeqRecord’s SeqFeature objects to build our diagram. Sometimes you won’t have SeqFeature objects, but just the coordinates for a feature you want to draw. You have to create minimal SeqFeature object, but this is easy:
from Bio.SeqFeature import SeqFeature, FeatureLocation
my_seq_feature = SeqFeature(FeatureLocation(50,100),strand=+1)
# For strand, use +1 for the forward strand, -1 for the reverse strand, and None for both. Here is a short self contained example:
# +
gdd = GenomeDiagram.Diagram('Test Diagram')
gdt_features = gdd.new_track(1, greytrack=False)
gds_features = gdt_features.new_set()
#Add three features to show the strand options,
feature = SeqFeature(FeatureLocation(25, 125), strand=+1)
gds_features.add_feature(feature, name="Forward", label=True)
feature = SeqFeature(FeatureLocation(150, 250), strand=None)
gds_features.add_feature(feature, name="Strandless", label=True)
feature = SeqFeature(FeatureLocation(275, 375), strand=-1)
gds_features.add_feature(feature, name="Reverse", label=True)
gdd.draw(format='linear', pagesize=(15*cm,4*cm), fragments=1,
start=0, end=400)
gdd.write("data/GD_labels_default.png", "png")
Image("data/GD_labels_default.png")
# -
# The top part of the image in the next subsection shows the output (in the default feature color, pale green).
#
# Notice that we have used the name argument here to specify the caption text for these features. This is discussed in more detail next.
# ### Feature captions
# Recall we used the following (where feature was a SeqFeature object) to add a feature to the diagram:
gd_feature_set.add_feature(feature, color=color, label=True)
# In the example above the SeqFeature annotation was used to pick a sensible caption for the features. By default the following possible entries under the SeqFeature object’s qualifiers dictionary are used: gene, label, name, locus_tag, and product. More simply, you can specify a name directly:
gd_feature_set.add_feature(feature, color=color, label=True, name="My Gene")
# In addition to the caption text for each feature’s label, you can also choose the font, position (this defaults to the start of the sigil, you can also choose the middle or at the end) and orientation (for linear diagrams only, where this defaults to rotated by 45 degrees):
# +
#Large font, parallel with the track
gd_feature_set.add_feature(feature, label=True, color="green",
label_size=25, label_angle=0)
#Very small font, perpendicular to the track (towards it)
gd_feature_set.add_feature(feature, label=True, color="purple",
label_position="end",
label_size=4, label_angle=90)
#Small font, perpendicular to the track (away from it)
gd_feature_set.add_feature(feature, label=True, color="blue",
label_position="middle",
label_size=6, label_angle=-90)
# -
# Combining each of these three fragments with the complete example in the previous section should give something like this:
gdd.draw(format='linear', pagesize=(15*cm,4*cm), fragments=1,
start=0, end=400)
gdd.write("data/GD_labels.png", "png")
Image("data/GD_labels.png")
# We’ve not shown it here, but you can also set label_color to control the label’s color.
#
# You’ll notice the default font is quite small - this makes sense because you will usually be drawing many (small) features on a page, not just a few large ones as shown here.
# ### Feature sigils
# The examples above have all just used the default sigil for the feature, a plain box, which was all that was available in the last publicly released standalone version of GenomeDiagram. Arrow sigils were included when GenomeDiagram was added to Biopython 1.50:
# +
#Default uses a BOX sigil
gd_feature_set.add_feature(feature)
#You can make this explicit:
gd_feature_set.add_feature(feature, sigil="BOX")
#Or opt for an arrow:
gd_feature_set.add_feature(feature, sigil="ARROW")
#Box with corners cut off (making it an octagon)
gd_feature_set.add_feature(feature, sigil="OCTO")
#Box with jagged edges (useful for showing breaks in contains)
gd_feature_set.add_feature(feature, sigil="JAGGY")
#Arrow which spans the axis with strand used only for direction
gd_feature_set.add_feature(feature, sigil="BIGARROW")
# -
# These are shown below. Most sigils fit into a bounding box (as given by the default BOX sigil), either above or below the axis for the forward or reverse strand, or straddling it (double the height) for strand-less features. The BIGARROW sigil is different, always straddling the axis with the direction taken from the feature’s stand.
# ### Arrow sigils
# We introduced the arrow sigils in the previous section. There are two additional options to adjust the shapes of the arrows, firstly the thickness of the arrow shaft, given as a proportion of the height of the bounding box:
#Full height shafts, giving pointed boxes:
gd_feature_set.add_feature(feature, sigil="ARROW", color="brown",
arrowshaft_height=1.0)
#Or, thin shafts:
gd_feature_set.add_feature(feature, sigil="ARROW", color="teal",
arrowshaft_height=0.2)
#Or, very thin shafts:
gd_feature_set.add_feature(feature, sigil="ARROW", color="darkgreen",
arrowshaft_height=0.1)
# The results are shown below:
# Secondly, the length of the arrow head - given as a proportion of the height of the bounding box (defaulting to 0.5, or 50%):
#Short arrow heads:
gd_feature_set.add_feature(feature, sigil="ARROW", color="blue",
arrowhead_length=0.25)
#Or, longer arrow heads:
gd_feature_set.add_feature(feature, sigil="ARROW", color="orange",
arrowhead_length=1)
#Or, very very long arrow heads (i.e. all head, no shaft, so triangles):
gd_feature_set.add_feature(feature, sigil="ARROW", color="red",
arrowhead_length=10000)
# The results are shown below:
# Biopython 1.61 adds a new BIGARROW sigil which always stradles the axis, pointing left for the reverse strand or right otherwise:
#A large arrow straddling the axis:
gd_feature_set.add_feature(feature, sigil="BIGARROW")
# All the shaft and arrow head options shown above for the ARROW sigil can be used for the BIGARROW sigil too.
# ### A nice example
# Now let’s return to the pPCP1 plasmid from Yersinia pestis biovar Microtus, and the top down approach used above, but take advantage of the sigil options we’ve now discussed. This time we’ll use arrows for the genes, and overlay them with strand-less features (as plain boxes) showing the position of some restriction digest sites.
# +
record = SeqIO.read("data/NC_005816.gb", "genbank")
gd_diagram = GenomeDiagram.Diagram(record.id)
gd_track_for_features = gd_diagram.new_track(1, name="Annotated Features")
gd_feature_set = gd_track_for_features.new_set()
for feature in record.features:
if feature.type != "gene":
#Exclude this feature
continue
if len(gd_feature_set) % 2 == 0:
color = colors.blue
else:
color = colors.lightblue
gd_feature_set.add_feature(feature, sigil="ARROW",
color=color, label=True,
label_size = 14, label_angle=0)
#I want to include some strandless features, so for an example
#will use EcoRI recognition sites etc.
for site, name, color in [("GAATTC","EcoRI",colors.green),
("CCCGGG","SmaI",colors.orange),
("AAGCTT","HindIII",colors.red),
("GGATCC","BamHI",colors.purple)]:
index = 0
while True:
index = record.seq.find(site, start=index)
if index == -1 : break
feature = SeqFeature(FeatureLocation(index, index+len(site)))
gd_feature_set.add_feature(feature, color=color, name=name,
label=True, label_size = 10,
label_color=color)
index += len(site)
gd_diagram.draw(format="linear", pagesize='A4', fragments=4,
start=0, end=len(record))
gd_diagram.write("data/plasmid_linear_nice.png", "png")
Image("data/plasmid_linear_nice.png")
# +
gd_diagram.draw(format="circular", circular=True, pagesize=(20*cm,20*cm),
start=0, end=len(record), circle_core = 0.5)
gd_diagram.write("data/plasmid_circular_nice.png", "png")
Image("data/plasmid_circular_nice.png")
# -
# ### Multiple tracks
# All the examples so far have used a single track, but you can have more than one track – for example show the genes on one, and repeat regions on another. In this example we’re going to show three phage genomes side by side to scale, inspired by Figure 6 in Proux et al. (2002). We’ll need the GenBank files for the following three phage:
#
# - NC_002703 – Lactococcus phage Tuc2009, complete genome (38347 bp)
# - AF323668 – Bacteriophage bIL285, complete genome (35538 bp)
# - NC_003212 – Listeria innocua Clip11262, complete genome, of which we are focussing only on integrated prophage 5 (similar length).
#
# You can download these using Entrez if you like. For the third record we’ve worked out where the phage is integrated into the genome, and slice the record to extract it, and must also reverse complement to match the orientation of the first two phage:
A_rec = SeqIO.read("data/NC_002703.gbk", "gb")
B_rec = SeqIO.read("data/AF323668.gbk", "gb")
# The figure we are imitating used different colors for different gene functions. One way to do this is to edit the GenBank file to record color preferences for each feature - something Sanger’s Artemis editor does, and which GenomeDiagram should understand. Here however, we’ll just hard code three lists of colors.
#
# Note that the annotation in the GenBank files doesn’t exactly match that shown in Proux et al., they have drawn some unannotated genes.
# +
from reportlab.lib.colors import red, grey, orange, green, brown, blue, lightblue, purple
A_colors = [red]*5 + [grey]*7 + [orange]*2 + [grey]*2 + [orange] + [grey]*11 + [green]*4 \
+ [grey] + [green]*2 + [grey, green] + [brown]*5 + [blue]*4 + [lightblue]*5 \
+ [grey, lightblue] + [purple]*2 + [grey]
B_colors = [red]*6 + [grey]*8 + [orange]*2 + [grey] + [orange] + [grey]*21 + [green]*5 \
+ [grey] + [brown]*4 + [blue]*3 + [lightblue]*3 + [grey]*5 + [purple]*2
# -
# Now to draw them – this time we add three tracks to the diagram, and also notice they are given different start/end values to reflect their different lengths.
# +
name = "data/Proux Fig 6"
gd_diagram = GenomeDiagram.Diagram(name)
max_len = 0
for record, gene_colors in zip([A_rec, B_rec], [A_colors, B_colors]):
max_len = max(max_len, len(record))
gd_track_for_features = gd_diagram.new_track(1,
name=record.name,
greytrack=True,
start=0, end=len(record))
gd_feature_set = gd_track_for_features.new_set()
i = 0
for feature in record.features:
if feature.type != "gene":
#Exclude this feature
continue
gd_feature_set.add_feature(feature, sigil="ARROW",
color=gene_colors[i], label=True,
name = str(i+1),
label_position="start",
label_size = 6, label_angle=0)
i+=1
gd_diagram.draw(format="linear", pagesize='A4', fragments=1,
start=0, end=max_len)
gd_diagram.write(name + ".png", "png")
Image(name + ".png")
# -
# I did wonder why in the original manuscript there were no red or orange genes marked in the bottom phage. Another important point is here the phage are shown with different lengths - this is because they are all drawn to the same scale (they are different lengths).
#
# The key difference from the published figure is they have color-coded links between similar proteins – which is what we will do in the next section.
# ### Cross-Links between tracks
# Biopython 1.59 added the ability to draw cross links between tracks - both simple linear diagrams as we will show here, but also linear diagrams split into fragments and circular diagrams.
#
# Continuing the example from the previous section inspired by Figure 6 from Proux et al. 2002, we would need a list of cross links between pairs of genes, along with a score or color to use. Realistically you might extract this from a BLAST file computationally, but here I have manually typed them in.
#
# My naming convention continues to refer to the three phage as A, B and C. Here are the links we want to show between A and B, given as a list of tuples (percentage similarity score, gene in A, gene in B).
#Tuc2009 (NC_002703) vs bIL285 (AF323668)
A_vs_B = [
(99, "Tuc2009_01", "int"),
(33, "Tuc2009_03", "orf4"),
(94, "Tuc2009_05", "orf6"),
(100,"Tuc2009_06", "orf7"),
(97, "Tuc2009_07", "orf8"),
(98, "Tuc2009_08", "orf9"),
(98, "Tuc2009_09", "orf10"),
(100,"Tuc2009_10", "orf12"),
(100,"Tuc2009_11", "orf13"),
(94, "Tuc2009_12", "orf14"),
(87, "Tuc2009_13", "orf15"),
(94, "Tuc2009_14", "orf16"),
(94, "Tuc2009_15", "orf17"),
(88, "Tuc2009_17", "rusA"),
(91, "Tuc2009_18", "orf20"),
(93, "Tuc2009_19", "orf22"),
(71, "Tuc2009_20", "orf23"),
(51, "Tuc2009_22", "orf27"),
(97, "Tuc2009_23", "orf28"),
(88, "Tuc2009_24", "orf29"),
(26, "Tuc2009_26", "orf38"),
(19, "Tuc2009_46", "orf52"),
(77, "Tuc2009_48", "orf54"),
(91, "Tuc2009_49", "orf55"),
(95, "Tuc2009_52", "orf60"),
]
# For the first and last phage these identifiers are locus tags, for the middle phage there are no locus tags so I’ve used gene names instead. The following little helper function lets us lookup a feature using either a locus tag or gene name:
def get_feature(features, id, tags=["locus_tag", "gene"]):
"""Search list of SeqFeature objects for an identifier under the given tags."""
for f in features:
for key in tags:
#tag may not be present in this feature
for x in f.qualifiers.get(key, []):
if x == id:
return f
raise KeyError(id)
# We can now turn those list of identifier pairs into SeqFeature pairs, and thus find their location co-ordinates. We can now add all that code and the following snippet to the previous example (just before the gd_diagram.draw(...) line – see the finished example script <a href="data/Proux_et_al_2002_Figure_6.py">Proux_et_al_2002_Figure_6.py</a> included in the Doc/examples folder of the Biopython source code) to add cross links to the figure:
from Bio.Graphics.GenomeDiagram import CrossLink
from reportlab.lib import colors
#Note it might have been clearer to assign the track numbers explicitly...
for rec_X, tn_X, rec_Y, tn_Y, X_vs_Y in [(A_rec, 2, B_rec, 1, A_vs_B)]:
track_X = gd_diagram.tracks[tn_X]
track_Y = gd_diagram.tracks[tn_Y]
for score, id_X, id_Y in X_vs_Y:
feature_X = get_feature(rec_X.features, id_X)
feature_Y = get_feature(rec_Y.features, id_Y)
color = colors.linearlyInterpolatedColor(colors.white, colors.firebrick, 0, 100, score)
link_xy = CrossLink((track_X, feature_X.location.start, feature_X.location.end),
(track_Y, feature_Y.location.start, feature_Y.location.end),
color, colors.lightgrey)
gd_diagram.cross_track_links.append(link_xy)
gd_diagram.draw(format="linear", pagesize='A4', fragments=1,
start=0, end=max_len)
gd_diagram.write("data/cross.png", "png")
Image("data/cross.png")
# There are several important pieces to this code. First the GenomeDiagram object has a cross_track_links attribute which is just a list of CrossLink objects. Each CrossLink object takes two sets of track-specific co-ordinates (here given as tuples, you can alternatively use a GenomeDiagram.Feature object instead). You can optionally supply a colour, border color, and say if this link should be drawn flipped (useful for showing inversions).
#
# You can also see how we turn the BLAST percentage identity score into a colour, interpolating between white (0%) and a dark red (100%). In this example we don’t have any problems with overlapping cross-links. One way to tackle that is to use transparency in ReportLab, by using colors with their alpha channel set. However, this kind of shaded color scheme combined with overlap transparency would be difficult to interpret. The result:
# There is still a lot more that can be done within Biopython to help improve this figure. First of all, the cross links in this case are between proteins which are drawn in a strand specific manor. It can help to add a background region (a feature using the ‘BOX’ sigil) on the feature track to extend the cross link. Also, we could reduce the vertical height of the feature tracks to allocate more to the links instead – one way to do that is to allocate space for empty tracks. Furthermore, in cases like this where there are no large gene overlaps, we can use the axis-straddling BIGARROW sigil, which allows us to further reduce the vertical space needed for the track. These improvements are demonstrated in the example script <a href="data/Proux_et_al_2002_Figure_6.py">Proux_et_al_2002_Figure_6.py</a>.
# Beyond that, finishing touches you might want to do manually in a vector image editor include fine tuning the placement of gene labels, and adding other custom annotation such as highlighting particular regions.
#
# Although not really necessary in this example since none of the cross-links overlap, using a transparent color in ReportLab is a very useful technique for superimposing multiple links. However, in this case a shaded color scheme should be avoided.
# ## Chromosomes
# The Bio.Graphics.BasicChromosome module allows drawing of chromosomes. There is an example in Jupe et al. (2012) (open access) using colors to highlight different gene families.
# ### Simple Chromosomes
# **Important**: To continue this example you have first to download a few chromosomes from Arabidopsis thaliana, the code to help you is here:
#
# **Very important**: This is slow and clogs the network, you only need to do this once (even if you close the notebook as the download will be persistent)
from ftplib import FTP
ftp = FTP('ftp.ncbi.nlm.nih.gov')
print("Logging in")
ftp.login()
ftp.cwd('genomes/archive/old_genbank/A_thaliana/OLD/')
print("Starting download - This can be slow!")
for chro, name in [
("CHR_I", "NC_003070.fna"), ("CHR_I", "NC_003070.gbk"),
("CHR_II", "NC_003071.fna"), ("CHR_II", "NC_003071.gbk"),
("CHR_III", "NC_003072.fna"), ("CHR_III", "NC_003072.gbk"),
("CHR_IV", "NC_003073.fna"), ("CHR_IV", "NC_003073.gbk"),
("CHR_V", "NC_003074.fna"), ("CHR_V", "NC_003074.gbk")]:
print("Downloading", chro, name)
ftp.retrbinary('RETR %s/%s' % (chro, name), open('data/%s' % name, 'wb').write)
ftp.quit()
print('Done')
# Here is a very simple example - for which we’ll use Arabidopsis thaliana.
#
# You can skip this bit, but first I downloaded the five sequenced chromosomes from the NCBI’s FTP site (per the code above) and then parsed them with Bio.SeqIO to find out their lengths. You could use the GenBank files for this, but it is faster to use the FASTA files for the whole chromosomes:
from Bio import SeqIO
entries = [("Chr I", "NC_003070.fna"),
("Chr II", "NC_003071.fna"),
("Chr III", "NC_003072.fna"),
("Chr IV", "NC_003073.fna"),
("Chr V", "NC_003074.fna")]
for (name, filename) in entries:
record = SeqIO.read("data/" + filename, "fasta")
print(name, len(record))
# This gave the lengths of the five chromosomes, which we’ll now use in the following short demonstration of the BasicChromosome module:
# +
from reportlab.lib.units import cm
from Bio.Graphics import BasicChromosome
entries = [("Chr I", 30432563),
("Chr II", 19705359),
("Chr III", 23470805),
("Chr IV", 18585042),
("Chr V", 26992728)]
max_len = 30432563 #Could compute this
telomere_length = 1000000 #For illustration
chr_diagram = BasicChromosome.Organism(output_format="png")
chr_diagram.page_size = (29.7*cm, 21*cm) #A4 landscape
for name, length in entries:
cur_chromosome = BasicChromosome.Chromosome(name)
#Set the scale to the MAXIMUM length plus the two telomeres in bp,
#want the same scale used on all five chromosomes so they can be
#compared to each other
cur_chromosome.scale_num = max_len + 2 * telomere_length
#Add an opening telomere
start = BasicChromosome.TelomereSegment()
start.scale = telomere_length
cur_chromosome.add(start)
#Add a body - using bp as the scale length here.
body = BasicChromosome.ChromosomeSegment()
body.scale = length
cur_chromosome.add(body)
#Add a closing telomere
end = BasicChromosome.TelomereSegment(inverted=True)
end.scale = telomere_length
cur_chromosome.add(end)
#This chromosome is done
chr_diagram.add(cur_chromosome)
chr_diagram.draw("data/simple_chrom.png", "Arabidopsis thaliana")
Image("data/simple_chrom.png")
# -
# This example is deliberately short and sweet. The next example shows the location of features of interest.
# Continuing from the previous example, let’s also show the tRNA genes. We’ll get their locations by parsing the GenBank files for the five Arabidopsis thaliana chromosomes. You’ll need to download these files from the NCBI FTP site.
# +
entries = [("Chr I", "NC_003070.gbk"),
("Chr II", "NC_003071.gbk"),
("Chr III", "NC_003072.gbk"),
("Chr IV", "NC_003073.gbk"),
("Chr V", "NC_003074.gbk")]
max_len = 30432563 #Could compute this
telomere_length = 1000000 #For illustration
chr_diagram = BasicChromosome.Organism(output_format="png")
chr_diagram.page_size = (29.7*cm, 21*cm) #A4 landscape
for index, (name, filename) in enumerate(entries):
record = SeqIO.read("data/" + filename,"genbank")
length = len(record)
features = [f for f in record.features if f.type=="tRNA"]
#Record an Artemis style integer color in the feature's qualifiers,
#1 = Black, 2 = Red, 3 = Green, 4 = blue, 5 =cyan, 6 = purple
for f in features: f.qualifiers["color"] = [index+2]
cur_chromosome = BasicChromosome.Chromosome(name)
#Set the scale to the MAXIMUM length plus the two telomeres in bp,
#want the same scale used on all five chromosomes so they can be
#compared to each other
cur_chromosome.scale_num = max_len + 2 * telomere_length
#Add an opening telomere
start = BasicChromosome.TelomereSegment()
start.scale = telomere_length
cur_chromosome.add(start)
#Add a body - again using bp as the scale length here.
body = BasicChromosome.AnnotatedChromosomeSegment(length, features)
body.scale = length
cur_chromosome.add(body)
#Add a closing telomere
end = BasicChromosome.TelomereSegment(inverted=True)
end.scale = telomere_length
cur_chromosome.add(end)
#This chromosome is done
chr_diagram.add(cur_chromosome)
chr_diagram.draw("data/tRNA_chrom.png", "Arabidopsis thaliana")
Image("data/tRNA_chrom.png")
# -
| notebooks/17 - Graphics including GenomeDiagram.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
# %load_ext autoreload
# %autoreload 2 """Reloads all functions automatically"""
# %matplotlib notebook
from irreversible_stressstrain import StressStrain as strainmodel
import test_suite as suite
import graph_suite as plot
import numpy as np
model = strainmodel('ref/HSRS/22').get_experimental_data()
slopes = suite.get_slopes(model)
second_deriv_slopes = suite.get_slopes(suite.combine_data(model[:-1,0],slopes))
# -- we think that yield occurs where the standard deviation is decreasing AND the slopes are mostly negative
def findYieldInterval(slopes, numberofsections):
def numneg(val):
return sum((val<0).astype(int))
# -- divide into ten intervals and save stddev of each
splitslopes = np.array_split(slopes,numberofsections)
splitseconds = np.array_split(second_deriv_slopes,numberofsections)
# -- displays the number of negative values in a range (USEFUL!!!)
for section in splitslopes:
print numneg(section), len(section)
print "-------------------------------"
for section in splitseconds:
print numneg(section), len(section)
divs = [np.std(vals) for vals in splitslopes]
# -- stddev of the whole thing
stdev = np.std(slopes)
interval = 0
slopesect = splitslopes[interval]
secondsect = splitseconds[interval]
print divs, stdev
# -- the proportion of slope values in an interval that must be negative to determine that material yields
cutoff = 3./4.
while numneg(slopesect)<len(slopesect)*cutoff and numneg(secondsect)<len(secondsect)*cutoff:
interval = interval + 1
"""Guard against going out of bounds"""
if interval==len(splitslopes): break
slopesect = splitslopes[interval]
secondsect = splitseconds[interval]
print
print interval
return interval
numberofsections = 15
interval_length = len(model)/numberofsections
"""
Middle of selected interval
Guard against going out of bounds
"""
yield_interval = findYieldInterval(slopes,numberofsections)
yield_index = min(yield_interval*interval_length + interval_length/2,len(model[:])-1)
yield_value = np.array(model[yield_index])[None,:]
print
print yield_value
# -
# ## Make these estimates more reliable and robust
# +
model = strainmodel('ref/HSRS/326').get_experimental_data()
strain = model[:,0]
stress = model[:,1]
slopes = suite.get_slopes(model)
second_deriv = suite.get_slopes(suite.combine_data(model[:-1,0],slopes))
"""Now what if we have strain vs slope"""
strainvslope = suite.combine_data(strain,slopes)
strainvsecond = suite.combine_data(strain,second_deriv)
plot.plot2D(strainvsecond,'Strain','Slope',marker="ro")
plot.plot2D(model,'Strain','Stress',marker="ro")
# +
model = strainmodel('ref/HSRS/326').get_experimental_data()
strain = model[:,0]
stress = model[:,1]
slopes = suite.get_slopes(model)
second_deriv = suite.get_slopes(suite.combine_data(model[:-1,0],slopes))
num_intervals = 80
interval_length = len(second_deriv)/num_intervals
split_2nd_derivs = np.array_split(second_deriv,num_intervals)
print np.mean(second_deriv)
down_index = 0
for index, section in enumerate(split_2nd_derivs):
if sum(section)<np.mean(slopes):
down_index = index
break
yield_index = down_index*interval_length
print strain[yield_index], stress[yield_index]
# +
model = strainmodel('ref/HSRS/326').get_experimental_data()
strain = model[:,0]
stress = model[:,1]
first_deriv = suite.get_slopes(model)
second_deriv = suite.get_slopes(suite.combine_data(model[:-1,0],first_deriv))
plot1 = suite.combine_data(strain,first_deriv)
plot2 = suite.combine_data(strain,second_deriv)
plot.plot2D(model)
plot.plot2D(plot1)
plot.plot2D(plot2)
# -
# ### See when standard deviation of second derivative begins to decrease
# +
model = strainmodel('ref/HSRS/222').get_experimental_data()
strain = model[:,0]
stress = model[:,1]
first_deriv = suite.get_slopes(model)
second_deriv = suite.get_slopes(suite.combine_data(model[:-1,0],first_deriv))
ave_deviation = np.std(second_deriv)
deviation_second = [np.std(val) for val in np.array_split(second_deriv,30)]
yielding = 0
for index,value in enumerate(deviation_second):
if value != 0.0 and value<ave_deviation and index!=0:
yielding = index
break
print "It seems to yield at index:", yielding
print "These are all of the standard deviations, by section:", deviation_second, "\n"
print "The overall standard deviation of the second derivative is:", ave_deviation
# -
# ## The actual yield values are as follows (These are approximate):
# ### ref/HSRS/22: Index 106 [1.3912797535, 900.2614980977]
# ### ref/HSRS/222: Index 119 [0, 904.6702299]
# ### ref/HSRS/326: Index 150 [6.772314989, 906.275032]
# ### Index of max standard deviation of the curve
# +
model = strainmodel('ref/HSRS/22').get_experimental_data()
strain = model[:,0]
stress = model[:,1]
first_deriv = suite.get_slopes(model)
second_deriv = suite.get_slopes(suite.combine_data(model[:-1,0],first_deriv))
print second_deriv;
return;
chunks = 20
int_length = len(model[:])/chunks
deriv2spl = np.array_split(second_deriv,chunks)
deviation_second = [abs(np.mean(val)) for val in deriv2spl]
del(deviation_second[0])
print deviation_second
print np.argmax(deviation_second)
#print "The standard deviation of all the second derivatives is", np.std(second_deriv)
# -
# ### If our data dips, we can attempt to find local maxima
# +
# -- climbs a discrete dataset to find local max
def hillclimber(data, guessindex = 0):
x = data[:,0]
y = data[:,1]
curx = x[guessindex]
cury = y[guessindex]
guessleft = max(0,guessindex-1)
guessright = min(len(x)-1,guessindex+1)
done = False
while not done:
left = y[guessleft]
right = y[guessright]
difleft = left-cury
difright = right-cury
if difleft<0 and difright<0 or (difleft==0 and difright==0):
done = True
elif difleft>difright:
cur = left
guessindex = guessleft
elif difright>difleft or difright==difleft:
cur = right
guessindex = guessright
return guessindex
func = lambda x: x**2
xs = np.linspace(0.,10.,5)
ys = func(xs)
data = suite.combine_data(xs,ys)
print hillclimber(data)
| .ipynb_checkpoints/Try to Find Yield Strength Automatically-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
try:
os.environ["QT_API"] = "pyqt"
from mayavi import mlab
except:
try:
os.environ["QT_API"] = "pyside"
from mayavi import mlab
except:
print("Unable to import mayavi")
from TricubicInterpolation import TriCubic
import numpy as np
def plotTCI(neTCI,rays=None,filename=None,show=False):
xmin = neTCI.xvec[0]
xmax = neTCI.xvec[-1]
ymin = neTCI.yvec[0]
ymax = neTCI.yvec[-1]
zmin = neTCI.zvec[0]
zmax = neTCI.zvec[-1]
X,Y,Z = np.mgrid[xmin:xmax:len(neTCI.xvec)*1j,
ymin:ymax:len(neTCI.yvec)*1j,
zmin:zmax:len(neTCI.zvec)*1j]
#reshape array
data = neTCI.getShapedArray()
import pylab as plt
xy = np.mean(data,axis=2)
yz = np.mean(data,axis=0)
zx = np.mean(data,axis=1)
fig,(ax1,ax2,ax3) = plt.subplots(1,3)
ax1.imshow(xy,origin='lower',aspect='auto')
ax2.imshow(yz,origin='lower',aspect='auto')
ax3.imshow(zx,origin='lower',aspect='auto')
plt.show()
#print(np.mean(data),np.max(data),np.min(data))
#mlab.close()
l = mlab.pipeline.volume(mlab.pipeline.scalar_field(X,Y,Z,data))#,vmin=min, vmax=min + .5*(max-min))
l._volume_property.scalar_opacity_unit_distance = min((xmax-xmin)/4.,(ymax-ymin)/4.,(zmax-zmin)/4.)
#l._volume_property.shade = False
mlab.contour3d(X,Y,Z,data,contours=10,opacity=0.2)
mlab.colorbar()
if rays is not None:
#[Na, Nt, Nd, 4, N]
i = 0
while i < rays.shape[0]:
j = 0
k = 0
while k < rays.shape[2]:
x,y,z = rays[i,0,k,0,:],rays[i,0,k,1,:],rays[i,0,k,2,:]
mlab.plot3d(x,y,z,tube_radius=0.75)
k += 1
i += 1
if filename is not None:
mlab.savefig('{}.png'.format(filename))#,magnification = 2)#size=(1920,1080))
if show:
mlab.show()
mlab.close()
def animateSolutions(datafolder,template):
'''animate the ne solutions'''
import os
import glob
try:
os.makedirs("{}/tmp".format(datafolder))
except:
pass
files = glob.glob("{}/{}".format(datafolder,template))
neTCIs = []
for filename in files:
print("plotting {}".format(filename))
neTCIs.append(TriCubic(filename=filename))
rays = np.load("{}/rays.npy".format(datafolder)).item(0)
idx = 0
for neTCI in neTCIs:
plotTCI(neTCI,rays=rays[0]+rays[1]+rays[2],filename="{0}/tmp/fig{1:4d}.png".format(datafolder,idx),show=True)
idx += 1
#os.system('ffmpeg -r 10 -f image2 -s 1900x1080 -i {0}/tmp/fig%04d.png -vcodec libx264 -crf 25 -pix_fmt yuv420p {0}/solution_animation.mp4'.format(datafolder))
def plotWavefront(neTCI,rays,save=False,saveFile=None,animate=False):
if saveFile is None:
saveFile = "figs/wavefront.png"
print("Saving to: {0}".format(saveFile))
xmin = neTCI.xvec[0]
xmax = neTCI.xvec[-1]
ymin = neTCI.yvec[0]
ymax = neTCI.yvec[-1]
zmin = neTCI.zvec[0]
zmax = neTCI.zvec[-1]
X,Y,Z = np.mgrid[xmin:xmax:len(neTCI.xvec)*1j,
ymin:ymax:len(neTCI.yvec)*1j,
zmin:zmax:len(neTCI.zvec)*1j]
#reshape array
data = neTCI.getShapedArray()
#print(np.mean(data),np.max(data),np.min(data))
#mlab.close()
#l = mlab.pipeline.volume(mlab.pipeline.scalar_field(X,Y,Z,data))#,vmin=min, vmax=min + .5*(max-min))
#l._volume_property.scalar_opacity_unit_distance = min((xmax-xmin)/4.,(ymax-ymin)/4.,(zmax-zmin)/4.)
#l._volume_property.shade = False
mlab.contour3d(X,Y,Z,data,contours=10,opacity=0.2)
mlab.colorbar()
def getWave(rays,idx):
xs = np.zeros(len(rays))
ys = np.zeros(len(rays))
zs = np.zeros(len(rays))
ridx = 0
while ridx < len(rays):
xs[ridx] = rays[ridx]['x'][idx]
ys[ridx] = rays[ridx]['y'][idx]
zs[ridx] = rays[ridx]['z'][idx]
ridx += 1
return xs,ys,zs
if rays is not None:
for datumIdx in range(len(rays)):
ray = rays[datumIdx]
mlab.plot3d(ray["x"],ray["y"],ray["z"],tube_radius=0.75)
if animate:
plt = mlab.points3d(*getWave(rays,0),color=(1,0,0),scale_mode='vector', scale_factor=10.)
#mlab.move(-200,0,0)
view = mlab.view()
@mlab.animate(delay=100)
def anim():
nt = len(rays[0]["s"])
f = mlab.gcf()
save = False
while True:
i = 0
while i < nt:
#print("updating scene")
xs,ys,zs = getWave(rays,i)
plt.mlab_source.set(x=xs,y=ys,z=zs)
#mlab.view(*view)
if save:
#mlab.view(*view)
mlab.savefig('figs/wavefronts/wavefront_{0:04d}.png'.format(i))#,magnification = 2)#size=(1920,1080))
#f.scene.render()
i += 1
yield
save = False
anim()
if save and animate:
import os
os.system('ffmpeg -r 10 -f image2 -s 1900x1080 -i figs/wavefronts/wavefront_%04d.png -vcodec libx264 -crf 25 -pix_fmt yuv420p figs/wavefronts/wavefront.mp4')
else:
if save:
mlab.savefig(saveFile,figure=mlab.gcf())
else:
mlab.show()
def plotModel(neTCI,save=False):
'''Plot the model contained in a tricubic interpolator (a convienient container for one)'''
plotWavefront(neTCI,None,save=save)
def animateResults(files):
from TricubicInterpolation import TriCubic
import os
images = []
index = 0
for file in files:
abspath = os.path.abspath(file)
print("Plotting: {0}".format(abspath))
if os.path.isfile(abspath):
dir = os.path.dirname(abspath)
froot = os.path.split(abspath)[1].split('.')[0]
else:
continue
data = np.load(abspath)
xvec = data['xvec']
yvec = data['yvec']
zvec = data['zvec']
M = data['M']
Kmu = data['Kmu'].item(0)
rho = data['rho']
Krho = data['Krho'].item(0)
if 'rays' in data.keys():
rays = data['rays'].item(0)
else:
rays = None
TCI = TriCubic(xvec,yvec,zvec,M)
TCI.clearCache()
images.append("{0}/frame-{1:04d}.png".format(dir,index))
plotWavefront(TCI,rays,save=True,saveFile=images[-1],animate=False)
index += 1
os.system('ffmpeg -r 10 -f image2 -s 1900x1080 -i {0}/frame-%04d.png -vcodec libx264 -crf 25 -pix_fmt yuv420p {1}/solution_animation.mp4'.format(dir,dir))
print("Saved to {1}/solution_animation.mp4".format(dir))
if __name__ == '__main__':
#plotTCI(neTCI,rays=None,filename=None,show=False)
#animateSolutions("output/simulatedInversion_2","initial_neModel.npy")
neTCI = TriCubic(filename='output/test/bfgs_3_1/nePriori.hdf5')
if True:
from RealData import DataPack
from CalcRays import calcRays
from AntennaFacetSelection import selectAntennaFacets
datapack = DataPack(filename="output/test/simulate/simulate_3/datapackSim.hdf5")
antIdx = -1
timeIdx = [0]
dirIdx = -1
i0 = 0
datapack = selectAntennaFacets(5, datapack, antIdx=-1, dirIdx=-1, timeIdx = np.arange(1))
antennas,antennaLabels = datapack.get_antennas(antIdx = -1)
patches, patchNames = datapack.get_directions(dirIdx = -1)
times,timestamps = datapack.get_times(timeIdx=[0])
datapack.setReferenceAntenna(antennaLabels[i0])
#plotDataPack(datapack,antIdx = antIdx, timeIdx = timeIdx, dirIdx = dirIdx,figname='{}/dobs'.format(outputfolder))
dobs = datapack.get_dtec(antIdx = antIdx, timeIdx = timeIdx, dirIdx = dirIdx)
Na = len(antennas)
Nt = len(times)
Nd = len(patches)
fixtime = times[Nt>>1]
phase = datapack.getCenterDirection()
arrayCenter = datapack.radioArray.getCenter()
rays = calcRays(antennas,patches,times, arrayCenter, fixtime, phase, neTCI, datapack.radioArray.frequency,
True, 1000, neTCI.nz)
from TricubicInterpolation import TriCubic
#print(neTCI.m)
K_ne = np.mean(neTCI.m)
#print(K_ne)
TCI = TriCubic(filename='output/test/bfgs_3_1/m_25.hdf5')
TCI.m = K_ne*np.exp(TCI.m) - neTCI.m
plotTCI(TCI,rays=rays,filename="output/test/model",show=True)
| src/ionotomo/notebooks/AnimateSolutions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# -
# configs
pd.set_option('max_rows', 20)
df_vgs = pd.read_csv(r'datasets\vgsales.csv')
df_vgs.info()
df_vgs.shape
# ##### Entendendo os dados
df_vgs.head()
df_vgs.describe()
df_vgs.isnull().sum()
df_vgs.dtypes
# ### Tratamentos
df_vgs.set_index('Rank', inplace = True)
df_vgs['Year'] = df_vgs['Year'].fillna(df_vgs['Year'].mean())
# Removed null values in 'Publisher'
df_vgs.dropna(axis=0, inplace=True)
df_vgs.isnull().sum()
df_vgs['Platform'].unique()
df_vgs['Genre'].unique()
all_publisher = df_vgs['Publisher'].value_counts()
df_vgs['Publisher'] = df_vgs['Publisher'].apply(lambda x: 'Others' \
if all_publisher[x] < 50 else x)
df_vgs['Publisher'].value_counts()
# ### Análises
df_vgs.head()
| MachineLearning/VideoGameSales/DataAnalysisML.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_braket
# language: python
# name: python3
# ---
# # Robust randomness generation on quantum processing units
# Random numbers are a ubiquitous resource in computation and cryptography. For example, in security, random numbers are crucial to creating keys for encryption. Quantum random number generators (QRNGs), that make use of the inherent unpredictability in quantum physics, promise enhanced security compared to standard cryptographic pseudo-random number generators (CPRNGs) based on classical technologies.
#
# In this notebook, we implement our own QRNG. Namely, we program two separate quantum processor units (QPUs) from different suppliers in Amazon Braket to supply two streams of weakly random bits. We then show how to generate physically secure randomness from these two weak sources by means of classical post-processing based on randomness extractors. The prerequisites for the tutorial are a basic understanding of quantum states, quantum measurements, and quantum channels. For a detailed explanation of these concepts we refer to the Amazon Braket notebook [Simulating noise on Amazon Braket](https://github.com/aws/amazon-braket-examples/blob/main/examples/braket_features/Simulating_Noise_On_Amazon_Braket.ipynb).
#
# We believe that randomness generation is a practical application of nowadays available noisy and intermediate scale quantum (NISQ) technologies.
#
# ## Table of contents
#
# * [Amazon Braket](#amazon_braket)
# * [Quantum circuit for randomness generation](#quantum_circuit)
# * [Quick implementation](#quick_implementation)
# * [Critical assessment](#critical_assessment)
# * [Interlude randomness extractors](#interlude)
# * [Quantum information](#quantum_information)
# * [Extractor construction](#extractor_construction)
# * [Example](#example)
# * [Implementation](#implementation)
# * [Unpredictability of physical sources](#physical_sources)
# * [Noise as leakage](#noise_leakage)
# * [Numerical evaluation](#numerical_evaluation)
# * [Putting everything together](#putting-together)
# * [Beyond current implementation](#beyond_current)
# * [Literature](#literature)
# ## Amazon Braket <a name="amazon_braket"></a>
#
# We start out with some general Amazon Braket imports, as well as some mathemtical tools needed.
# +
# AWS imports: Import Braket SDK modules
from braket.circuits import Circuit
from braket.devices import LocalSimulator
from braket.aws import AwsDevice, AwsQuantumTask
# set up local simulator device
device = LocalSimulator()
# general math imports
import math, random
import numpy as np
from scipy.fft import fft, ifft
# magic word for producing visualizations in notebook
# %matplotlib inline
# import convex solver
import cvxpy as cp
# +
# set up Rigetti quantum device
rigetti = AwsDevice("arn:aws:braket:us-west-1::device/qpu/rigetti/Aspen-M-1")
# set up IonQ quantum device
ionq = AwsDevice("arn:aws:braket:::device/qpu/ionq/ionQdevice")
# simulator alternative: set up the managed simulator SV1
simulator = AwsDevice("arn:aws:braket:::device/quantum-simulator/amazon/sv1")
# -
# ## Quantum circuit for randomness generation <a name="quantum_circuit"></a>
#
# Arguably the simplest way of generating a random bit on a quantum computer is as follows:
# * Prepare the basis state vector $|0\rangle$
# * Apply the Hadamard gate $H=\frac{1}{\sqrt{2}}\begin{pmatrix} 1 & 1\\ 1 & 1 \end{pmatrix}$ leading to the state vector $|H\rangle=\frac{1}{\sqrt{2}}\big(|0\rangle+|1\rangle\big)$
# * Measure in the computational basis $\big\{|0\rangle\langle0|,|1\rangle\langle1|\big\}$.
#
# By the laws of quantum physics, the post-measurement probability distribution is then the uniformly distributed $(1/2,1/2)$ and leads to one random bit $0/1$.
#
# In the following, we discuss how above protocol is conceptually different from randomness obtained from classical sources and show in detail how it is implemented reliable even when the underlying quantum processing units employed are noisy. By the end of this tutorial, you will be able to create your own random bits from the quantum processing units available on Amazon Braket.
#
# ## Quick implementation <a name="quick_implementation"></a>
#
# The Hadmard gate based quantum circuit for generating one random bit can be repeated or run in parallel $n$ times, leading to a random bit string of length $n$. The corresponding circuit is easily implemented in Amazon Braket:
# +
# function for Hadamard cirquit
def hadamard_circuit(n_qubits):
"""
function to apply Hadamard gate on each qubit
input: number of qubits
"""
# instantiate circuit object
circuit = Circuit()
# apply series of Hadamard gates
for i in range(n_qubits):
circuit.h(i)
return circuit
# define circuit
n_qubits = 5
state = hadamard_circuit(n_qubits)
# print circuit
print(state)
# -
# Let us run this Hadamard circuit with $n=5$ qubits in the local quantum simulator for $m=1$ shots:
#
# (Note: We will work on actual QPUs towards the end of this tutorial.)
# +
# run circuit
m_shots = 1
result = device.run(state, shots = m_shots).result()
# get measurement shots
counts = result.measurement_counts.keys()
# print counts
list_one = list(counts)[0]
array_one = np.array([list_one])
print("The output bit string is: ",array_one)
# -
# ## Critical assessment <a name="critical_assessment"></a>
#
# The advantage of such quantum random number generators over implementations based on classical technologies is that the outcomes are intrinsically random. That is, according to the laws of quantum physics, the outcome of the measurement is not only hard to predict, but rather impossible to know before the measurement has taken place.
#
# However, since current quantum processing units are noisy to a certain degree, there are at least three potential problems that need to be addressed:
# * First, the noise acting on all states and operations performed might lead to a systematic bias towards the probability of getting the measurement outcomes $0$ or $1$, respectively.
# * Second, even if aforementioned noise is not biased towards certain measurement outcomes, the generated randomness is no longer solely based on intrinsically random quantum effects, but rather partly on the noise present.
# * Third, whereas by the laws of quantum physics a pure quantum state cannot be correlated to the outside world, any noisy acting on the system corresponds to information leaking to the environment. This is because no information is destroyed in quantum physics and hence, a malicious third party knowing about the noise occurring will be able to guess the generated bits (at least up to a certain degree).
#
# When the noise model acting on the quantum processor units is characterized to some degree (e.g., by means of previous benchmarking), these shortcomings can be overcome by employing two independent quantum processor units, together with an appropriate classical post-processing. The latter is based on classical algorithms from the theory of pseudo-randomness, so-called two-source extractors. This is what we discuss next.
#
# In the following, we refer a few times to the theory paper [1] that features formal cryptographic security definitions, together with mathematical proofs, as well as some statistical methods tailored to intermediate scale quantum devices. These pointers can be safely ignored when only interested in the implementation of our QRNG.
#
# ## Interlude randomness extractors <a name="interlude"></a>
#
# Two-source extractors allow distillation of physically secure random bits from two independent weak sources of randomness whenever they are sufficiently unpredictable to start with. The relevant measure of unpredictability is thereby given by the min-entropy of the respective sources, defined for the probability distribution $\{p_x\}_{x\in X}$ as
#
# $$ \text{$H_{\min}(X)=-\log_2p_{\text{guess}(X)}$ with $p_{\text{guess}(X)}=\max_{x\in X}p_x$.} $$
#
# That is, the min-entropy exactly quantifies how well we can guess the value of the source, or in other words, how unpredictable the source is. For example, for $n$-bit distributions $X$ we have $H_{\min}(X)\in[0,n]$, where $0$ corresponds to a deterministic distribution containing no randomness and $n$ to the perfectly random, uniform distribution.
#
# A two-source extractor is a function $\text{Ext}:\{0,1\}^{n_1}\times\{0,1\}^{n_2}\to\{0,1\}^m$ such that for any two independent sources with min-entropy at least $H_{\min}(X_1)\geq k_1$ and $H_{\min}(X_2)\geq k_2$, respectively, the output of length $m$ is $\epsilon\in[0,1]$ close in variational distance to the perfectly random, uniform distribution $U_M$ of size $m$:
#
# $$ \frac{1}{2}\left\|\text{Ext}(X_1,X_2)-U_M\right\|_1\leq\epsilon. $$
#
# So, two inpedendent sources that are only weakly random get condensed by these algorithms to one output that is (nearly) perfectly random! Importantly, the output becomes truly physically random with no computational assumptions introduced.
#
# ### Quantum information <a name="quantum_information"></a>
#
# For our setting we need an extension of this concept, as a potentially malicious third party can collect quantum information $Q$ about the weak source of randomness $X$. The corresponding conditional min-entropy is defined as
#
# $$ H_{\min}(X|Q)=-\log_2p_{\text{guess}(X|Q)}, $$
#
# where $p_{\text{guess}(X|Q)}$ denotes the maximal probability allowed by quantum physics to guess the classical value $X$ by applying any measurements on the quantum information $Q$. We notice that even though $p_{\text{guess}(X|Q)}$ does not have a closed form expression, it is efficiently computed by means of a semidefinite program (see the theory notes [1] for details). This is also how we will evaluate the conditional min-entropy quantity later on.
#
# Accordingly, a quantum-proof two-source extractor is then function $\text{Ext}:\{0,1\}^{n_1}\times\{0,1\}^{n_2}\to\{0,1\}^m$ such that for any two independent sources with quantum conditional min-entropy at least $H_{\min}(X_1|Q_1)\geq k_1$ and $H_{\min}(X_2|Q_2)\geq k_2$, respectively, we have for $\epsilon\in[0,1]$ in quantum variational distance
#
# $$ \frac{1}{2}\left\|\rho_{\text{Ext}(X_1,X_2)Q_1Q_2}-\tau_{M}\otimes\rho_{Q_1}\otimes\rho_{Q_2}\right\|_1\leq\epsilon,$$
#
# where $\tau_M$ denotes the fully mixed $m$ qubit state. That is, the extractor should not only make the $m$ output bits perfectly random, but also decouple them from any outside correlations initially present - up to the security parameter $\epsilon\in[0,1]$.
#
# For more details about these concepts, we refer to the theory notes [1] and references therein. All that is important to us here, is that there exist quantum-proof two-source extractors with good parameters. Next, we discuss one particular such construction that we subsequently implement in an efficient manner.
#
# ## Extractor construction <a name="extractor_construction"></a>
#
# In this paragraph, we provide an explicit construction of a quantum-proof two-source extractor that efficiently provides non-zero output $M$ for a wide range of sizes of the inputs $X_1$ and $X_2$. Namely, we employ a Toeplitz matrices based construction originally discussed in [2]:
#
# For the security parameter $\epsilon\in(0,1]$ and inputs $X_1,X_2$ of size $n$ and $n-1$, respectively, the function $\text{Ext}:\{0,1\}^n\times\{0,1\}^{n-1}\to\{0,1\}^m$ defined below is a quantum-proof two-source randomness extractor with output size
#
# $$ m=\left\lfloor(k_1+k_2-n)+1-2\log\left(1/\epsilon\right)\right\rfloor. $$
#
# The function is explicitly given via the vector-matrix multiplication
#
# $$ (x,y)\mapsto \text{Ext}(x,y)=x\cdot(T(y)|1_m)^T \mod{2}$$
#
# featuring the Toeplitz matrix
#
# $$ T(y)=\begin{pmatrix}
# y_0 & y_1 & \ldots & y_{n-m-1}\\
# y_{-1} & y_0 & \ldots & y_{n-m-2}\\
# \vdots & \vdots & \vdots & \vdots\\
# y_{1-m} & y_{2-m} & \ldots & y_{n-2m}
# \end{pmatrix}\;\text{from}\;y=(y_{1-m},\ldots,y_0,\ldots,y_{n-m-1})\in\{0,1\}^{n-1}, $$
#
# The quantum-proof property of this construction, as well as its complexity, is explicitly disccused in the theory notes [1].
#
# For our setting, we will have sources with linear min-entropy rates $k_i=\alpha_i\cdot n$ for $i=1,2$. The Toeplitz construction then works whenever $\alpha_1+\alpha_2-1>0$ and we can compute the required input size for fixed output size as
#
# $$ n=\left\lfloor\frac{m-1+2\log(1/\epsilon)}{\alpha_1+\alpha_2-1}\right\rfloor.$$
#
# A simple example shows that these numbers work well in practice, even for very small input sizes around $n\approx100$.
#
# ### Example <a name="example"></a>
#
# Let us set the security parameter to $\epsilon=10^{-8}$, say that we have min-entropy sources with linear rates $k_1=n\cdot0.8$ and $k_2=(n-1)\cdot0.8$, and ask for $m=100$ fully random bits. According to the formulae above, $n=253$ together with $n-1=252$ weakly random bits can be condensed into $100$ fully random bits (up to the security parameter $\epsilon=10^{-8}$).
#
# Next, we give an efficient implementation of this Toeplitz based construction.
#
# ### Implementation <a name="implementation"></a>
#
# The vector-matrix multiplication $x\cdot(T(y)|1_m)^T$ a priori has asymptotic complexity $O(n^2)$ in big O-notation, which is prohibitive for larger input sizes $n\geq10^4$. However, we discuss in the theory notes [1] that the operation is actually implemented in asymptotic complexity $O(n\log n)$ by first embedding the problem into circulant matrices and then making use of the Fast Fourier Transform (FFT). The corresponding code then performs well up to input sizes $n\geq10^7$. The following example demonstrates this implementation:
# +
# work with local simulator for testing Toeplitz construction
device = LocalSimulator()
# set security parameter
power = 8
eps = 10**(-power)
print(f"Security parameter: {eps}.")
# set number of output bits
m = 10
print(f"Desired output length: {m} bits.")
# set min-entropy rates for sources
k_1 = 0.8
print(f"Min-entropy rate of first source: {k_1}.")
k_2 = 0.8
print(f"Min-entropy rate of second source: {k_2}")
# required number of input bits (for each source)
n = math.floor((m-1-2*math.log2(eps))
/(k_1+k_2-1))
print(f"Required length of each input source: {n} bits.")
# quantum circuit for generating weakly random bit string one
n1_qubits = 1
m1_shots = n
state1 = hadamard_circuit(n1_qubits)
result1 = device.run(state1, shots=m1_shots).result()
array_one = result1.measurements.reshape(1,m1_shots*n1_qubits)
# print(array_one)
# quantum circuit for generating weakly random bit string two
n2_qubits = 1
m2_shots = n
state2 = hadamard_circuit(n2_qubits)
result2 = device.run(state2, shots=m2_shots).result()
array_two = result2.measurements.reshape(1,m2_shots*n2_qubits)
# print(array_two)
###
# alternative for generating two bit strings when no quantum source is available:
# create first list of pseudo-random bits
# alternative when no quantum source is available
# list_one = []
# for number in range(n):
# b = int(random.randint(0, 1))
# list_one.append(b)
# array_one = np.array([list_one])
# create second list of pseudo-random bits
# list_two = []
# for number in range(n):
# b = int(random.randint(0, 1))
# list_two.append(b)
# array_two = np.array([list_two])
###
# computing output of Toeplitz extractor by vector-matrix multiplication
# via efficient Fast Fourier Transform (FFT) as discussed in [1]
# setting up arrays for FFT implementation of Toeplitz
array_two_under = np.array(array_two[0,0:n-m])[np.newaxis]
zero_vector = np.zeros((1,n+m-3), dtype=int)
array_two_zeros = np.hstack((array_two_under,zero_vector))
array_two_over = array_two[0,n-m:n][np.newaxis]
array_one_merged = np.zeros((1,2*n-3), dtype=int)
for i in range(m):
array_one_merged[0,i] = array_one[0,m-1-i]
for j in range(n-m-1):
array_one_merged[0,n+m-2+j] = array_one[0,n-2-j]
# FFT multplication output of Toeplitz
output_fft = np.around(ifft(fft(array_one_merged)*fft(array_two_zeros)).real)
output_addition = output_fft[0,0:m] + array_two_over
output_final = output_addition.astype(int) % 2
print(f"The {m} random output bits are:\n{output_final}.")
# -
# As an alternative, we note that efficient implementations of other quantum-proof two-source extractors are discussed in [3].
#
# ## Unpredictability of physical sources <a name="physical_sources"></a>
#
# Given above methods on randomness extraction, the next step is to give lower bounds on the min-entropy present in the output distributions generated from our $n$-fold Hadamard circuit. For that, we need to model the noise present in the quantum processing units.
#
# Generally, for any given quantum processing unit, the supplier typically publishes some type of noise specification with it. This includes both, the noise characterization of state preparation, as well as the read-out measurements. In case such specifications are not available, or if one wants to double check them, it is in principle possible to benchmark the device. We refer to the theory notes [1] for more details on this and just mention here that we do not need a full characterization of the device, but rather only conservative upper bounds on the noise strength. This then translates into lower bounds on the min-entropy present in the system.
#
# For our case, since we are only applying single qubit gates for our Hadamard circuit, the noise is captured well by single qubit noise models. Moreover, for the state preparation step via Hadamard gates, the typical noise in quantum architectures is uniform, depolarizing noise of some strength $\lambda\in[0,1]$. That is, all possible single qubit errors such as bit flip or phase flip errors are equally likely, leading to the effective evolution
#
# $$ \psi=|\psi\rangle\langle\psi|\mapsto\text{Dep}^\lambda(\psi)=(1-\lambda)\cdot\psi+\lambda\cdot\frac{|0\rangle\langle0|+|1\rangle\langle1|}{2}, $$
#
# mapping any input qubit state $\psi$ onto a linear combination of itself and the maximally mixed qubit state $\frac{|0\rangle\langle0|+|1\rangle\langle1|}{2}$. Note that we now work with general mixed states in order to model classical statistical uncertainty coming from the noise model.
#
# So effectively, before the measurement step, instead of the perfect pure state $|H\rangle\langle H|_A$ as defined by the vector $|H\rangle_A=\frac{1}{\sqrt{2}}\big(|0\rangle_A+|1\rangle_A\big)$, we have the mixed state
#
# $$ \rho_A^\lambda=\frac{1}{2}\big(|0\rangle\langle0|_A+(1-\lambda)|0\rangle\langle1|_A+(1-\lambda)|1\rangle\langle0|_A+|1\rangle\langle1|_A\big) $$
#
# at hand.
#
# Next, we note that instead of the ideal measurement device given by $\mathcal{M}=\big\{|0\rangle\langle0|_A,|1\rangle\langle1|_A\big\}$, the typical noisy measurement device is described by
#
# $$ \mathcal{N}^\mu=\big\{1_A-\mu|1\rangle\langle1|_A,\mu|1\rangle\langle1|_A\big\} $$
#
# with some bias $\mu\in(0,1)$ towards reading-out the ground state $|0\rangle\langle0|_A$ over $|1\rangle\langle1|_A$. The post-measurement probability distribution is then given as
#
# $$ Q^{\lambda,\mu}=(q_0=1-\mu/2,q_1=\mu/2) $$
#
# instead of the perfectly uniform distribution $P=(p_0=1/2,p_1=1/2)$. Note that the former distribution has non-maximal min-entropy
#
# $$ H_{\min}(X)_{Q^{\lambda,\mu}}=1-\log\mu\leq1=H_{\min}(X)_P. $$
#
# More generally, as measurement devices are the most sensitive element of quantum randomness generation, we discuss in the theory notes [1] methods for benchmarking them (even when only noisy state preparation is available).
#
# ### Noise as leakage <a name="noise_leakage"></a>
#
# It is, however, crucial to realize that $H_{\min}(X)_{Q^{\lambda,\mu}}$ is not yet the quantity relevant for secure quantum randomness generation. As information is never lost in quantum mechanics, all the noise carries information to the environment, where it can in principle be picked up by an attacker. That is, we need to estimate the conditional min-entropy of the post-measurement probability distribution $X$ given any complementary information that leaked into the environment [5].
#
# This is worked out in detail in the theory notes [1] by means of so-called purifications of both, the noisy qubit state $\rho_A^\lambda$, as well as the noisy measurement device $\mathcal{N}^\mu$, leading to the additional qubit registers $A'$ and $E_2$, respectively. The relevant so-called classical-quantum state to consider then takes the form
#
# $$ \omega_{XA'E_2}^{\lambda,\mu}= q_0\cdot|0\rangle\langle0|_X\otimes\omega_{A'E_2}^{\lambda,\mu}(0) + q_1\cdot|1\rangle\langle1|_X\otimes\omega_{A'E_2}^{\lambda,\mu}(1) $$
#
# with a classing part $X$ and the quantum parts $A'E_2$ depending on both noise parameters $\lambda,\mu$. The corresponding conditional min-entropy is
#
# $$ H_{\min}(X|A'E_2)_{\omega^{\lambda,\mu}}\leq H_{\min}(X)_{Q^{\lambda,\mu}} $$
#
# with the gap between the conditional and the unconditional term typically being strict.
#
# We reiterate that the reason for mathematically introducing the purification registers $AE_2$ and working with the conditional min-entropy, is to make sure that the output bits generated are random even conditioned on the knowledge of the noise. This ensures that the randomness created is from a purely quantum origin and is secure against any eavesdropping from a malicious third party.
#
# We mention that a more detailed discussion of this point is given in the work [5].
#
# ### Numerical evaluation <a name="numerical_evaluation"></a>
#
# In the following, we use the noise parameters $\lambda=0.02$ and $\mu=0.98$ for the quantum processing units and this immediately gives
#
# $$H_{\min}(X)\approx0.944. $$
#
# The next step is to numerically evaluate the conditional min-entropy
#
# $$ H_{\min}(X|A'E_2)=-\log p_{\text{guess}}(X|A'E_2). $$
#
# This is done by means of a standard semidefinite program (sdp) solver as follows:
# +
# fix noise parameters
lamb = 0.02
mu = 0.98
# purification of rho input state
rho = 0.5*np.array([[1,1-lamb],[1-lamb,1]])
eigvals, eigvecs = np.linalg.eig(rho)
rho_vector =\
math.sqrt(eigvals[0])*np.kron(eigvecs[:,0],eigvecs[:,0])[np.newaxis]\
+math.sqrt(eigvals[1])*np.kron(eigvecs[:,1],eigvecs[:,1])[np.newaxis]
rho_pure = np.kron(rho_vector,rho_vector.T)
# sigma state of noisy measurement device
sigma_vector = np.array([[math.sqrt(1-mu),0,0,math.sqrt(mu)]])
sigma_pure = np.kron(sigma_vector,sigma_vector.T)
# omega state relevant for conditional min-entropy
rho_sigma = np.kron(rho_pure,sigma_pure)
id_2 = np.identity(2)
zero = np.array([[1,0]])
one = np.array([[0,1]])
zerozero = np.kron(np.kron(zero,id_2),np.kron(zero,id_2))
zeroone = np.kron(np.kron(zero,id_2),np.kron(one,id_2))
onezero = np.kron(np.kron(one,id_2),np.kron(zero,id_2))
oneone = np.kron(np.kron(one,id_2),np.kron(one,id_2))
omega_0 = zerozero@rho_sigma@zerozero.T+zeroone@rho_sigma@zeroone.T+onezero@rho_sigma@onezero.T
omega_1 = oneone@rho_sigma@oneone.T
omega = []
omega.append(omega_0)
omega.append(omega_1)
# sdp solver
m = 4 # dimension of quantum side information states
c = 2 # number of classical measurement outcomes
sigma = cp.Variable((m,m), complex=True) # complex variable
constraints = [sigma >> 0] # positive semi-definite
constraints += [sigma >> omega[i] for i in range(c)] # min-entropy constraints
obj = cp.Minimize(cp.real(cp.trace(sigma))) # objective function
prob = cp.Problem(obj,constraints) # set up sdp problem
prob.solve(solver=cp.SCS, verbose=True) # solve sdp problem using splitting conic solver (SCS)
guess = prob.value
qmin_entropy = (-1)*math.log2(guess)
min_entropy = 1-math.log2(2-mu*(1-lamb))
print("\033[1m" + "The coditional min-entropy is: ", qmin_entropy)
print("As a comparison, the unconditional min-entropy is: ", min_entropy)
# -
# That is, for the chosen noise parameters $\lambda=0.02$ and $\mu=0.98$ we find
#
# $$ H_{\min}(X|A'E_2)\approx0.719<0.944\approx H_{\min}(X).$$
#
# By varying the noise parameter to other values, one also sees by inspection that the conditional min-entropy is monotone in both $\lambda$ and $\mu$. Importantly, this ensures that the outputted randomness will be safe to use whenever we put a conservative estimate on the noisy strength, even in the absence of an exact characterization of the underlying quantum processing units.
#
# Finally, whenever we run our Hadamard circuit $n$ times or on $n$ qubits in parallel, the overall conditional min-entropy is just given by
#
# $$ H_{\min}(X|RE_2)_{\omega^{\otimes n}} = n\cdot H_{\min}(X|RE_2)_\omega. $$
#
# This reason is that we only consider single qubit product noise together with the min-entropy being additive on product states. This then leads to the promised linear min-entropy rates $k_i=\alpha_i\cdot n$ for $i=1,2$ going into the Toeplitz two-source extractor.
#
# As an added bonus, the Toeplitz two-source extractor used has the so-called strong property. That is, even conditioned on the knowledge of one input string of weakly random bits, the output bits are still fully random. We refer to the technical notes [1] for discussion, a consequence being that if one provider builds in some backdoors in the provided unit, the random generation scheme is still not broken unless the two providers come together to cooperate.
#
#
# ## Putting everything together <a name="putting-together"></a>
#
# Now that we have determined the conditional min-entropy of our physical sources and have an efficient quantum-proof two-source extractor in place, all that remains is to put the two pieces together.
#
# 1. First, we specify how many random bits we want to generate, the desired security parameter, and the conditional min-entropy of our weak sources of randomness from the quantum processing units:
# +
# set security parameter
power = 8
eps = 10**(-power)
print(f"Security parameter: {eps}.")
# set number of output bits
m = 10
print(f"Desired output length: {m} bits.")
# set min-entropy rates for sources - qmin_entropy from above
k_one = qmin_entropy
print(f"Min-entropy rate of first source: {k_1}.")
k_two = qmin_entropy
print(f"Min-entropy rate of second source: {k_2}.")
# required number of input bits (for each source)
n = math.floor((m-1-2*math.log2(eps))
/(k_one+k_two-1))
print(f"Required length of each input source: {n} bits.")
# -
# 2. At the beginning of the notebook, we loaded two separate QPUs as available in Amazon Braket. We now run on the respective QPUs the Hadamard circuit followed by measurements in the computational basis:
#
# (Note: If preferred, one can alternatively use the pre-loaded managed simulator SV1. In this case, please comment out the QPU code in the cell below and instead uncomment the provided SV1 code.)
# +
# Rigetti: quantum circuit for generating weakly random bit string one
n1_q = 1 # alternatively use more than one qubit (attention: 32 max + lower bounds on number of shots)
m1_s = int(math.ceil(n/n1_q))
state_rigetti = hadamard_circuit(n1_q)
rigetti_task = rigetti.run(state_rigetti, shots=m1_s, poll_timeout_seconds=5*24*60*60)
rigetti_task_id = rigetti_task.id
rigetti_status = rigetti_task.state()
print("Status of Rigetti task:", rigetti_status)
# IonQ: quantum circuit for generating weakly random bit string two
n2_q = 1 # alternatively use more than one qubit (attention: 11 max + lower bounds on number of shots)
m2_s = int(math.ceil(n/n2_q))
state_ionq = hadamard_circuit(n2_q)
ionq_task = ionq.run(state_ionq, shots=m2_s, poll_timeout_seconds=5*24*60*60)
ionq_task_id = ionq_task.id
ionq_status = ionq_task.state()
print("Status of IonQ task:", ionq_status)
###
# alternative via managed simulator SV1
# quantum circuit for generating weakly random bit string one (simulate Rigetti source)
# n1_q = 1 # alternatively run multiple qubits in parallel
# m1_s = int(math.ceil(n/n1_q))
# state1 = hadamard_circuit(n1_q)
# result1 = simulator.run(state1, shots=m1_s).result()
# array_rigetti = result1.measurements.reshape(1,m1_s*n1_q)
# print("The first raw bit string is: ",array_one)
# quantum circuit for generating weakly random bit string two (simulate IonQ source)
# n2_q = 1 # alternatively run multiple qubits in parallel
# m2_s = int(math.ceil(n/n2_q))
# state2 = hadamard_circuit(n2_q)
# result2 = simulator.run(state2, shots=m2_s).result()
# array_ionq = result2.measurements.reshape(1,m2_s*n2_q)
# print("The second raw bit string is: ",array_two)
###
# -
# 3. The tasks have now been sent to the respective QPUs and we can recover the results at any point in time once the status is completed:
#
# (Note: In case you opted to use the managed simulator SV1 instead of the QPUs, please do not run the following cell.)
# +
# recover Rigetti task
task_load_rigetti = AwsQuantumTask(arn=rigetti_task_id)
# print status
status_rigetti = task_load_rigetti.state()
print('Status of Rigetti task:', status_rigetti)
# wait for job to complete
# terminal_states = ['COMPLETED', 'FAILED', 'CANCELLED']
if status_rigetti == 'COMPLETED':
# get results
rigetti_results = task_load_rigetti.result()
# array
array_rigetti = rigetti_results.measurements.reshape(1,m1_s*n1_q)
print("The first raw bit string is: ",array_rigetti)
elif status_rigetti in ['FAILED', 'CANCELLED']:
# print terminal message
print('Your Rigetti task is in terminal status, but has not completed.')
else:
# print current status
print('Sorry, your Rigetti task is still being processed and has not been finalized yet.')
# recover IonQ task
task_load_ionq = AwsQuantumTask(arn=ionq_task_id)
# print status
status_ionq = task_load_ionq.state()
print('Status of IonQ task:', status_ionq)
# wait for job to complete
# terminal_states = ['COMPLETED', 'FAILED', 'CANCELLED']
if status_ionq == 'COMPLETED':
# get results
ionq_results = task_load_ionq.result()
# array
#print(m1_shots,n1_qubits)
#print(m2_shots,n2_qubits)
#print(ionq_results.measurements)
array_ionq = ionq_results.measurements.reshape(1,m2_s*n2_q)
print("The second raw bit string is: ",array_ionq)
elif status_ionq in ['FAILED', 'CANCELLED']:
# print terminal message
print('Your IonQ task is in terminal status, but has not completed.')
else:
# print current status
print('Sorry, your IonQ task is still being processed and has not been finalized yet.')
# -
# 4. We run the Toeplitz two-source extractor on the two sequences of raw random input bits:
# setting up arrays for fft implementation of Toeplitz
if status_ionq == 'COMPLETED':
array_two_under = np.array(array_ionq[0,0:n-m])[np.newaxis]
zero_vector = np.zeros((1,n+m-3), dtype=int)
array_two_zeros = np.hstack((array_two_under,zero_vector))
array_two_over = array_ionq[0,n-m:n][np.newaxis]
array_one_merged = np.zeros((1,2*n-3), dtype=int)
if status_rigetti == 'COMPLETED':
for i in range(m):
array_one_merged[0,i] = array_rigetti[0,m-1-i]
for j in range(n-m-1):
array_one_merged[0,n+m-2+j] = array_rigetti[0,n-2-j]
# fft multplication output of Toeplitz
output_fft = np.around(ifft(fft(array_one_merged)*fft(array_two_zeros)).real)
output_addition = output_fft[0,0:m] + array_two_over
output_final = output_addition.astype(int) % 2
print(f"The {m} random output bits are:\n{output_final}.")
else:
print(f"Your Rigetti task is in {status_rigetti} state.")
else:
print(f"Your IonQ task is in {status_ionq} state.")
# 5. That is it, above bits are random up to the chosen security parameter!
#
# If one of the two QPUs works better or provides more immediate results, you can also run the quantum circuit twice on QPUs from the same provider (while being aware of the potentially violated independence assumption).
#
# ## Beyond current implementation <a name="beyond_current"></a>
#
# Different quantum processing units are potentially governed by different noise models. Correspondingly, this will lead to different conditional min-entropy rates for the raw sources of randomness. In this notebook and following the theory notes [1], we can change the noise model for the numerical evaluation of the min-entropy (with all other parts remaining the same).
#
# From a cryptographic viewpoint, the quantum hardware providers as well as Amazon Braket have to be trusted in order to end up with secure random numbers. More broadly, we note that there are more intricate ways of generating randomness from quantum sources than presented here, in particular for the use case when one is not ready to trust the underlying quantum hardware because of potential backdoors inbuilt. Such a so-called (semi) device-independent scheme is for example given in [3], with a corresponding implementation in Amazon Braket [7]. For a general in-depth discussion of QRNGs, we refer to the review article [4], as well as the extensive security report [6].
#
#
# ## Literature <a name="literature"></a>
#
# [1] <NAME> and <NAME>, Robust randomness generation on quantum computers, [available online](http://marioberta.info/wp-content/uploads/2021/07/randomness-theory.pdf).
#
# [2] <NAME> and <NAME>, More Efficient Privacy Amplification With Less Random Seeds via Dual Universal Hash Function, IEEE Transactions on Information Theory, [10.1109/TIT.2016.2526018](https://ieeexplore.ieee.org/document/7399404).
#
# [3] <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, Practical randomness and privacy amplification, [arXiv:2009.06551](https://arxiv.org/abs/2009.06551).
#
# [4] <NAME> and <NAME>, Quantum random number generators, Review of Modern Physics, [10.1103/RevModPhys.89.015004](https://doi.org/10.1103/RevModPhys.89.015004).
#
# [5] <NAME>, <NAME>, <NAME>, True randomness from realistic quantum devices, [arXiv:1311.4547](https://arxiv.org/abs/1311.4547).
#
# [6] <NAME>, <NAME>, <NAME>, Quantum random-number generators: practical considerations and use cases, [evolutionQ](https://evolutionq.com/quantum-safe-publications/qrng-report-2021-evolutionQ.pdf).
#
# [7] Quantum-Proof Cryptography with IronBridge, TKET and Amazon Braket, <NAME>, <NAME>, <NAME>, [Cambridge Quantum Computing](https://medium.com/cambridge-quantum-computing/quantum-proof-cryptography-with-ironbridge-tket-and-amazon-braket-e8e96777cacc).
| examples/advanced_circuits_algorithms/Randomness/Randomness_Generation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # NumPy
#
# ## What is NumPy?
#
# NumPy is a library for the Python programming language, adding support for large, multi-dimensional arrays and matrices, along with a large collection of high-level mathematical functions to operate on these arrays. The ancestor of NumPy, Numeric, was originally created by <NAME> with contributions from several other developers. In 2005, <NAME> created NumPy by incorporating features of the competing Numarray into Numeric, with extensive modifications. NumPy is open-source software and has many contributors.
#
# ## How to install NumPy?
#
# The only prerequisite for NumPy is Python itself. If you don’t have Python yet and want the simplest way to get started, we recommend you use the Anaconda Distribution - it includes Python, NumPy, and other commonly used packages for scientific computing and data science. NumPy can be installed with conda, with pip, or with a package manager on macOS and Linux.
#
# If you use pip, you can install it with:
#
# ```bash
# pip install numpy
# ```
#
# If you use conda, you can install it with:
#
# ```bash
# conda install numpy
# ```
#
# ## Numpy basics
#
# NumPy’s main object is the homogeneous multidimensional array. It is a table of elements (usually numbers), all of the same type, indexed by a tuple of non-negative integers. In NumPy dimensions are called axes.
#
# For example, the coordinates of a point in 3D space `[1, 2, 1]` has one axis. That axis has 3 elements in it, so we say it has a length of 3. In the example pictured below, the array has 2 axes. The first axis has a length of 2, the second axis has a length of 3.
#
# ```python
# [[ 1., 0., 0.],
# [ 0., 1., 2.]]
# ```
#
# NumPy’s array class is called `ndarray`. It is also known by the alias array. Note that `numpy.array` is not the same as the Standard Python Library class `array.array`, which only handles one-dimensional arrays and offers less functionality. The more important attributes of an ndarray object are:
#
# - `ndarray.ndim` the number of axes (dimensions) of the array.
# - `ndarray.shape` the dimensions of the array. This is a tuple of integers indicating the size of the array in each dimension. For a matrix with n rows and m columns, shape will be (n,m). The length of the shape tuple is therefore the number of axes, ndim.
# - `ndarray.size` the total number of elements of the array. This is equal to the product of the elements of shape.
# - `ndarray.dtype` an object describing the type of the elements in the array. One can create or specify dtype’s using standard Python types. Additionally NumPy provides types of its own. numpy.int32, numpy.int16, and numpy.float64 are some examples.
# - `ndarray.itemsize` the size in bytes of each element of the array. For example, an array of elements of type `float64` has itemsize 8 (=64/8), while one of type `complex32` has itemsize 4 (=32/8). It is equivalent to `ndarray.dtype.itemsize`.
# - `ndarray.data` the buffer containing the actual elements of the array. Normally, we won’t need to use this attribute because we will access the elements in an array using indexing facilities.
#
# ```python
# >>> import numpy as np
# >>> a = np.arange(15).reshape(3, 5)
# >>> a
# array([[ 0, 1, 2, 3, 4],
# [ 5, 6, 7, 8, 9],
# [10, 11, 12, 13, 14]])
#
# >>> a.shape
# (3, 5)
#
# >>> a.ndim
# 2
#
# >>> a.dtype.name
# 'int64'
#
# >>> a.itemsize
# 8
#
# >>> a.size
# 15
#
# >>> type(a)
# <class 'numpy.ndarray'>
#
# >>> b = np.array([6, 7, 8])
# >>> b
# array([6, 7, 8])
#
# >>> type(b)
# <class 'numpy.ndarray'>
# ```
#
# ## Array Creation
#
# There are several ways to create arrays.
#
# For example, you can create an array from a regular Python list or tuple using the array function. The type of the resulting array is deduced from the type of the elements in the sequences.
#
# ```python
# >>> import numpy as np
# >>> a = np.array([2,3,4])
# >>> a
# array([2, 3, 4])
#
# >>> a.dtype
# dtype('int64')
#
# >>> b = np.array([1.2, 3.5, 5.1])
# >>> b.dtype
# dtype('float64')
# ```
#
# A frequent error consists in calling array with multiple arguments, rather than providing a single sequence as an argument.
#
# ```python
# >>> a = np.array(1,2,3,4) # WRONG
# Traceback (most recent call last):
# ...
# TypeError: array() takes from 1 to 2 positional arguments but 4 were given
#
# >>> a = np.array([1,2,3,4]) # RIGHT
# ```
#
# array transforms sequences of sequences into two-dimensional arrays, sequences of sequences of sequences into three-dimensional arrays, and so on.
#
# ```python
# >>> b = np.array([(1.5,2,3), (4,5,6)])
# >>> b
# array([[1.5, 2. , 3. ],
# [4. , 5. , 6. ]])
# ```
#
# The type of the array can also be explicitly specified at creation time:
#
# ```python
# >>> c = np.array( [ [1,2], [3,4] ], dtype=complex )
# >>> c
# array([[1.+0.j, 2.+0.j],
# [3.+0.j, 4.+0.j]])
# ```
#
# Often, the elements of an array are originally unknown, but its size is known. Hence, NumPy offers several functions to create arrays with initial placeholder content. These minimize the necessity of growing arrays, an expensive operation.
#
# The function zeros creates an array full of zeros, the function ones creates an array full of ones, and the function empty creates an array whose initial content is random and depends on the state of the memory. By default, the dtype of the created array is float64.
#
# ```python
# >>> np.zeros((3, 4))
# array([[0., 0., 0., 0.],
# [0., 0., 0., 0.],
# [0., 0., 0., 0.]])
#
# >>> np.ones( (2,3,4), dtype=np.int16 ) # dtype can also be specified
# array([[[1, 1, 1, 1],
# [1, 1, 1, 1],
# [1, 1, 1, 1]],
#
# [[1, 1, 1, 1],
# [1, 1, 1, 1],
# [1, 1, 1, 1]]], dtype=int16)
#
# >>> np.empty( (2,3) ) # uninitialized
# array([[ 3.73603959e-262, 6.02658058e-154, 6.55490914e-260],
# [ 5.30498948e-313, 3.14673309e-307, 1.00000000e+000]])
# ```
#
# To create sequences of numbers, NumPy provides the arange function which is analogous to the Python built-in range, but returns an array.
#
# ```python
# >>> np.arange( 10, 30, 5 )
# array([10, 15, 20, 25])
#
# >>> np.arange( 0, 2, 0.3 ) # it accepts float arguments
# array([0. , 0.3, 0.6, 0.9, 1.2, 1.5, 1.8])
# ```
#
# When arange is used with floating point arguments, it is generally not possible to predict the number of elements obtained, due to the finite floating point precision. For this reason, it is usually better to use the function linspace that receives as an argument the number of elements that we want, instead of the step:
#
# ```python
# >>> from numpy import pi
# >>> np.linspace( 0, 2, 9 ) # 9 numbers from 0 to 2
# array([0. , 0.25, 0.5 , 0.75, 1. , 1.25, 1.5 , 1.75, 2. ])
#
# >>> x = np.linspace( 0, 2*pi, 100 ) # useful to evaluate function at lots of points
# >>> f = np.sin(x)
# ```
#
# ## Basic operations
#
# Arithmetic operators on arrays apply elementwise. A new array is created and filled with the result.
#
# ```python
# >>> a = np.array( [20,30,40,50] )
# >>> b = np.arange( 4 )
# >>> b
# array([0, 1, 2, 3])
#
# >>> c = a-b
# >>> c
# array([20, 29, 38, 47])
#
# >>> b**2
# array([0, 1, 4, 9])
#
# >>> 10*np.sin(a)
# array([ 9.12945251, -9.88031624, 7.4511316 , -2.62374854])
#
# >>> a<35
# array([ True, True, False, False])
# ```
#
# Unlike in many matrix languages, the product operator `*` operates elementwise in NumPy arrays. The matrix product can be performed using the `@` operator (in python >=3.5) or the dot function or method:
#
# ```python
# >>> A = np.array( [[1,1],
# ... [0,1]] )
# >>> B = np.array( [[2,0],
# ... [3,4]] )
# >>> A * B # elementwise product
# array([[2, 0],
# [0, 4]])
#
# >>> A @ B # matrix product
# array([[5, 4],
# [3, 4]])
#
# >>> A.dot(B) # another matrix product
# array([[5, 4],
# [3, 4]])
# ```
#
# Some operations, such as `+=` and `*=`, act in place to modify an existing array rather than create a new one.
#
# ```python
# >>> rg = np.random.default_rng(1) # create instance of default random number generator
# >>> a = np.ones((2,3), dtype=int)
# >>> b = rg.random((2,3))
# >>> a *= 3
# >>> a
# array([[3, 3, 3],
# [3, 3, 3]])
#
# >>> b += a
# >>> b
# array([[3.51182162, 3.9504637 , 3.14415961],
# [3.94864945, 3.31183145, 3.42332645]])
#
# >>> a += b # b is not automatically converted to integer type
# Traceback (most recent call last):
# ...
# numpy.core._exceptions.UFuncTypeError: Cannot cast ufunc 'add' output from dtype('float64') to dtype('int64') with casting rule 'same_kind'
# ```
#
# When operating with arrays of different types, the type of the resulting array corresponds to the more general or precise one (a behavior known as upcasting).
#
#
# ```python
# >>> a = np.ones(3, dtype=np.int32)
# >>> b = np.linspace(0,pi,3)
# >>> b.dtype.name
# 'float64'
#
# >>> c = a+b
# >>> c
# array([1. , 2.57079633, 4.14159265])
#
# >>> c.dtype.name
# 'float64'
#
# >>> d = np.exp(c*1j)
# >>> d
# array([ 0.54030231+0.84147098j, -0.84147098+0.54030231j,
# -0.54030231-0.84147098j])
#
# >>> d.dtype.name
# 'complex128'
# ```
#
# Many unary operations, such as computing the sum of all the elements in the array, are implemented as methods of the ndarray class.
#
# ```python
# >>> a = rg.random((2,3))
# >>> a
# array([[0.82770259, 0.40919914, 0.54959369],
# [0.02755911, 0.75351311, 0.53814331]])
#
# >>> a.sum()
# 3.1057109529998157
#
# >>> a.min()
# 0.027559113243068367
#
# >>> a.max()
# 0.8277025938204418
# ```
#
# By default, these operations apply to the array as though it were a list of numbers, regardless of its shape. However, by specifying the axis parameter you can apply an operation along the specified axis of an array:
#
# ```python
# >>> b = np.arange(12).reshape(3,4)
# >>> b
# array([[ 0, 1, 2, 3],
# [ 4, 5, 6, 7],
# [ 8, 9, 10, 11]])
#
#
# >>> b.sum(axis=0) # sum of each column
# array([12, 15, 18, 21])
#
#
# >>> b.min(axis=1) # min of each row
# array([0, 4, 8])
#
# >>> b.cumsum(axis=1) # cumulative sum along each row
# array([[ 0, 1, 3, 6],
# [ 4, 9, 15, 22],
# [ 8, 17, 27, 38]])
# ```
#
# ## Indexing and slicing
#
# One-dimensional arrays can be indexed, sliced and iterated over, much like lists and other Python sequences.
#
# ```python
# >>> a = np.arange(10)**3
# >>> a
# array([ 0, 1, 8, 27, 64, 125, 216, 343, 512, 729])
#
# >>> a[2]
# 8
#
# >>> a[2:5]
# array([ 8, 27, 64])
#
# # equivalent to a[0:6:2] = 1000;
# # from start to position 6, exclusive, set every 2nd element to 1000
# >>> a[:6:2] = 1000
# >>> a
#
# array([1000, 1, 1000, 27, 1000, 125, 216, 343, 512, 729])
#
# >>> a[ : :-1] # reversed a
# array([ 729, 512, 343, 216, 125, 1000, 27, 1000, 1, 1000])
# ```
# +
# This notebook has no exercises
# You can test out numpy by yourself after this comment
| 03 Numpy/notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import cv2
import numpy as np
from matplotlib import pyplot as plt
def preprocess(file_name, grayscale=False):
file = cv2.imread(file_name)
if grayscale:
file = cv2.cvtColor(file, cv2.COLOR_BGR2GRAY)
data = cv2.resize(file, (400, 400), interpolation=cv2.INTER_CUBIC)
# ret,th1 = cv2.threshold(data, 0, 255,cv2.THRESH_TRUNC)
# ret, data = cv2.threshold(data, 0, 255, cv2.THRESH_OTSU)
return data
def plot(image_data, title=None, color=True):
image_data = np.float32(image_data/255)
if color:
color = ('b','g','r')
for i, col in enumerate(color):
histr = cv2.calcHist([image_data], [i], None, [256], [0, 1])
plt.plot(histr, color = col)
plt.xlim([0, 256])
plt.title(title)
plt.show()
else:
histr = cv2.calcHist([image_data], [0], None, [256], [0, ])
plt.plot(histr)
plt.xlim([0, 256])
plt.title(title)
plt.show()
images = {
"pivot": "test_image1.jpg",
"compare1": "test_image2.jpg",
"rose": "rose.jpg",
"yellow_car": "yellow_car.jpg",
"red_rose": "red_rose.jpg",
"yellow_lily": "yellow_lily.jpg"
}
for title, image in images.items():
image_data = preprocess(image)
plot(image_data, title)
| histogram_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
target_composer_names = ['<NAME>', '<NAME>']
embeddings_path = '../data/embeddings/composer-embeddings-c2v-dbow-5000-10000.h5'
# +
from difflib import SequenceMatcher
import glob
import re
import h5py
import numpy as np
import pandas as pd
import scipy
from scipy.spatial import distance
# -
all_composers = [(i, *c) for i, c in enumerate(pd.read_csv('../data/composers.csv', sep='|').values)]
# +
def similar(a, b):
return SequenceMatcher(None, a, b).ratio()
def name_to_composer_id(name):
composer = max(all_composers, key=lambda c: similar(c[1], name))
composer_id = composer[0]
print('Assuming {}: born {}; died {}; composer_id: {}'.format(composer[1], composer[2], composer[3], composer[0]))
return composer_id
target_ids = [name_to_composer_id(name) for name in target_composer_names]
# -
def path_to_embedding(path):
with h5py.File(path, 'r') as f:
return f.get('doc_embeddings').value
embeddings = path_to_embedding(embeddings_path)
distances = distance.cdist(embeddings, embeddings, metric='cosine')
closest = distances.argsort()
for t_id, t_name in zip(target_ids, target_composer_names):
print('Most similar to {}:'.format(t_name))
for c_id in closest[t_id, 1:6]:
print((all_composers[c_id][1], all_composers[c_id][-1]))
| notebooks/similarity-examples-dbow.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.0.2
# language: julia
# name: julia-1.0
# ---
# **Please note this jupyter notebook is in Julia.**
# ### Diversity Trumps Ability
#
#
#
# In 2004, <NAME> and <NAME> published a game-changing paper on diversity. They used agent-based model to prove that a diverse group of people is better than a group of best-performing individuals. These days, diversity is a hot topic as it falls into the spectrum of ESG. Don't get me wrong. Diversity here doesn't stand for gender or ethnicity. Diversity refers to the differences in problem solvers' perspectives and heuristics (Hong and Page, 1998). It is the variation in how people encode and search for solutions to problems. Despite the high citations of this paper, plenty of scholars dispute the idea which makes the theory rather controversial. The caveat is the experiment of solving problems on lattice is too simple and straight forward whereas in real life the situation is more dynamic. The diversity theory is completely inapplicable to social science. So the objective here is to convert lattice problem into the context of avant-garde random graphs. This random graph should satisfy two conditions. First, it should have as many of the properties of a real life problem as possible and second it should be simple with respect to our ability to simulate the performance of different agents. Thus, let's run an agent-based simulation on Waxman model.
import Plots
import Random
import LinearAlgebra
import Distributions
import Statistics
# ### Functions
#create waxman model
#which is a more generalized random geometric graph
#return the adjacency matrix
#the raison d'être of waxman model is
#its similarity with real geographic features
#the connectivity should be negatively correlated with distance
function create_waxman_model(coordinates,α=0.1,β=0.1)
#create euclidean distance matrix
distance_matrix=[LinearAlgebra.norm(coordinates[i]-coordinates[j]) for i in eachindex(coordinates),j in eachindex(coordinates)];
#create adjacency matrix
adjacency=zeros(length(coordinates),length(coordinates));
#only connect vertices when the probability is larger than the threshold
L=maximum(distance_matrix)
for i in 1:length(coordinates)
for j in i+1:length(coordinates)
prob=β*ℯ^(-distance_matrix[i,j]/L*α)
if Random.rand()>1-prob
adjacency[i,j]=distance_matrix[i,j]
adjacency[j,i]=distance_matrix[i,j]
end
end
end
return adjacency,distance_matrix
end
# +
#draw graph without graphplot.jl
#basically scatter plot to show vertices
#line plot to show edges
function viz(adjacency,coordinates)
pic=Plots.plot(legend=false,grid=false,axis=false,
title="Waxman Model"
)
#plot edges
for i in 1:length(coordinates)
for j in i+1:length(coordinates)
if adjacency[i,j]!=0
Plots.plot!([coordinates[i][1],coordinates[j][1]],
[coordinates[i][2],coordinates[j][2]],color="grey")
end
end
end
#plot vertices
for i in 1:length(coordinates)
Plots.scatter!([i[1] for i in coordinates],
[i[2] for i in coordinates],
markersize=10)
end
pic
end
# -
#agent based modelling
function agent_based_simulation(subgroup,adjacency,distance_matrix,
agents,max_path=100,current_vertex=1)
path=[]
stop=false
while !stop
for agent_id in subgroup
#create agent perspective+heuristic
d=Distributions.Normal(agents[agent_id]["perspective"],
agents[agent_id]["heuristic"],)
td=Distributions.truncated(d,-Inf,Inf)
#find neighbors of current vertex and their distance to the terminal
neighbors_key=[i for i in eachindex(adjacency[current_vertex,:]) if adjacency[current_vertex,i]!=0]
neighbors_val=[distance_matrix[i,length(adjacency[1,:])] for i in eachindex(adjacency[current_vertex,:]) if adjacency[current_vertex,i]!=0]
neighbors=Dict(zip(neighbors_key,neighbors_val))
#update the distance with perspective+heuristic
#the way it works is similar to a* algorithm in graph theory
# https://github.com/je-suis-tm/graph-theory/blob/master/a_star%20maze.ipynb
for neighbor_id in keys(neighbors)
ϵ=Random.rand(td)
neighbors[neighbor_id]+=ϵ
end
#select next vertex based upon the shortest subjective distance to the terminal
push!(path,current_vertex)
selected_neighbor=sort(collect(neighbors),by=x->x[2])[1][1]
current_vertex=selected_neighbor
#when maximum path length is reached
#terminate the while loop
if length(path)==max_path
stop=true
push!(path,current_vertex)
break
end
#when destination is reached
#terminate the while loop
if current_vertex==length(adjacency[1,:])
stop=true
push!(path,current_vertex)
break
end
end
end
return path
end
#create animated path
function animation(coordinates,vertex_status,filename)
anim=Plots.@animate for ind in 0:length(path)-1
pic=Plots.plot(legend=false,grid=false,axis=false,
title="$filename\nt=$ind")
#create edges
for i in 1:length(coordinates)
for j in i+1:length(coordinates)
if adjacency[i,j]!=0
Plots.plot!([coordinates[i][1],coordinates[j][1]],
[coordinates[i][2],coordinates[j][2]],
color="grey")
end
end
end
#create vertices
#path is showed via different colors
for i in 1:length(coordinates)
Plots.scatter!([i[1] for i in coordinates],
[i[2] for i in coordinates],
markersize=10,color=[i for i in vertex_status[ind+1]])
end
pic
end
Plots.gif(anim,filename*".gif",fps=0.5)
end
# ### Waxman Model Demo
#
#
#
# Waxman model was created by <NAME> to test how the algorithms of minimum spanning tree perform on random graphs. The model per se is a reflection of the reality where connectivity is negatively correlated with distance. The probability of creating an edge between vertex $u$ and $v$ is written below.
#
# $$P({u,v})=\beta \times \mathrm{exp} (\frac {-d(u,v)}{L \alpha})$$
#
# where
#
# $P({u,v})$ denotes the probability of creating an undirected edge from vertex $u$ to $v$
#
# $d(u,v)$ denotes Euclidean distance from vertex $u$ to $v$.
#
# $L$ denotes the largest distance inside the rectangular coordinate grid.
#
# $\mathrm{exp}$ denotes Euler's number.
#
# $\alpha$ denotes the density of short edges relative to longer ones.
#
# $\beta$ denotes overall graph edge density.
#
# There are something interesting about Waxman model. $\frac {d(u,v)} {L}$ creates a relative ratio to assess the distance within the rectangular coordinate grid regardless of the actual scale. A smaller $\alpha$ implies more short edges than long edges. A larger $\beta$ indicates more edges take place in general.
# +
#initialize graph order
graph_order=48
#create random coordinates
coordinate_range=20:80
X=Random.rand(coordinate_range,graph_order)
Y=Random.rand(coordinate_range,graph_order)
coordinates=Any[Int32[X[i],Y[i]] for i in 1:length(X)]
#add start and end
startpoint=Int32[1,1]
endpoint=Int32[100,100]
insert!(coordinates,1,startpoint)
insert!(coordinates,length(coordinates)+1,endpoint);
# -
#create waxman model and viz
adjacency,distance_matrix=create_waxman_model(coordinates,0.5,0.1)
viz(adjacency,coordinates)
# ### Agent-based Model Demo
#
#
#
# In the original paper (<NAME>, 2004), the authors proposed an agent-based model for a group of agents to find the largest possible value on a lattice. Here, we slightly modify the problem. A group formed with $N$ number of agents are required to travel from $start$ to $end$ on a random graph called Waxman model. Agents collaborate in a group sequentially. At each unit time $t$, agent $i$ pick the next vertex $v$ based upon its perception of the distance from current vertex $u$ to the destination $end$ which is called $f_i(u,end)$. The perception consists of two components, the actual Euclidean distance $d(u,end)$ and random disturbance $\epsilon$.
#
# $$ f_i(u,end)=d(u,end)+\epsilon_{i}$$
#
# Each agent's $\epsilon$ follows Gaussian distribution with mean $\mu$ and $\sigma$. Because the probability of connecting two vertices $P({u,v})$ is negatively correlated with their Euclidean distance $d(u,v)$, a good instinct for agent is to select a vertex closer to the destination which creates a higher chance of connecting to $end$. In that sense, a good agent should have relatively small $\epsilon$ so its perception of the distance $f_i(u,v)$ is closer to Euclidean distance $d(u,v)$.
#
# $$\epsilon_{i} \sim \mathcal{N} (\mu_i,\sigma_i)$$
#
# $\mu$ refers to the agent's perspective and $\sigma$ refers to the agent's heuristic. Both $\mu$ and $\sigma$ follow uniform distribution. $\sigma$ has one-sided interval due to its non-negativity constraint. An agent's perspective reflects how the world is perceived by the agent which is fixed by $\mu_i$. A good agent should have $\mu_i$ closer to zero which reflects its objectivity. An agent's heuristic reflects how consistent the agent's perspective is. A good agent should have $\sigma_i$ closer to zero which reflects its subjectivity.
#
# $$\mu_i \sim U [-5,5]$$
# $$\sigma_i \sim U [1,5]$$
#
# +
num_of_agents=50
group_size=5
#create random agents
#their perspectives follow uniform distribution [-5,5]
#their heuristics follow uniform distribution [1,5]
agents=Dict()
for i in 1:num_of_agents
agents[i]=Dict()
agents[i]["perspective"]=Random.rand(-5:5)
agents[i]["heuristic"]=Random.rand(1:5)
end
# +
#create a random group of size 3
randomgroup=Random.randperm(num_of_agents)[1:group_size]
#create path based upon group selection
path=agent_based_simulation(randomgroup,adjacency,distance_matrix,agents)
# +
#agent status determines which agent is making vertex selection at time t
agent_status=append!(repeat([i for i in 1:group_size],length(path)÷group_size),
[i for i in 1:group_size][1:length(path)%group_size])
#vertex status determines which vertex is being occupied by which agent
#this will be used as vertex color mapping for visualization
vertex_status=[]
push!(vertex_status,[4 for _ in 1:length(coordinates)])
vertex_status[end][1]=1
for itr in 1:length(path)-1
push!(vertex_status,[i for i in vertex_status[end]])
vertex_status[end][path[itr+1]]=agent_status[itr]
end
# -
#create animation with simple demo
#different colors show the choices by different agents
filename="Diversity Trumps Ability"
animation(coordinates,vertex_status,filename)
# ### Diversity and Ability
#
#
#
# The previous one was a simplified demonstration. To replicate the experiment in the paper, we need to expand the order and the size of our random graph. We create a random group which draws agents randomly from the pool to form the democracy. We also create an elite group which draws agents on the top percentile to form the technocracy. For comparison, we examine the mean distance two groups have traveled, the mean length of the paths and the diversity of two groups. The diversity is measured by $\rho$, which is defined by the aggregation of an agent's perspective and heuristic. The diversity metric $\Delta$ is merely the variance of $\rho$ inside the group.
#
# $$ \rho_i=|\mu_i|+\sigma_i$$
# $$ \Delta =\frac {1}{N} \sum_{i=1}^{N} (\rho_i-\bar\rho)^2$$
# +
#initialize graph order
graph_order=998
#create random coordinates
coordinate_range=3000:7000
X=Random.rand(coordinate_range,graph_order)
Y=Random.rand(coordinate_range,graph_order)
coordinates=Any[Int32[X[i],Y[i]] for i in 1:length(X)]
#add start and end
startpoint=Int32[1,1]
endpoint=Int32[10000,10000]
insert!(coordinates,1,startpoint)
insert!(coordinates,length(coordinates)+1,endpoint);
#create waxman model
adjacency,distance_matrix=create_waxman_model(coordinates,0.2,0.2);
# +
#we should limit the group size
#as the group size grows
#the diversity of elite group converges towards random group
num_of_agents=500
group_size=10
#create random agents
#their perspectives follow uniform distribution [-500,500]
#their heuristics follow uniform distribution [1,500]
agents=Dict()
for i in 1:num_of_agents
agents[i]=Dict()
agents[i]["perspective"]=Random.rand(-500:500)
agents[i]["heuristic"]=Random.rand(1:500)
end
# -
#a fine balance between speed and effectiveness
num_of_simulations=500;
#agent based simulation of random group
distance_arr=[]
path_arr=[]
subgroup_diversity_arr=[]
for _ in 1:num_of_simulations
subgroup=Random.randperm(num_of_agents)[1:group_size]
path=agent_based_simulation(subgroup,adjacency,distance_matrix,agents)
distance=sum([distance_matrix[path[i],path[i+1]] for i in 1:(length(path)-1)])
subgroup_diversity=Statistics.var([abs(agents[agent_id]["perspective"])+agents[agent_id]["heuristic"] for agent_id in subgroup])
push!(distance_arr,distance)
push!(path_arr,length(path))
push!(subgroup_diversity_arr,subgroup_diversity)
end
println("Distance travelled:",Statistics.mean(distance_arr))
println("Length of the path:",Statistics.mean(path_arr))
println("Group diversity:",Statistics.mean(subgroup_diversity_arr))
#random group quickly finds the path
#even though sometimes it looks like a random walk
path
#agent based simulation of elite group
distance_arr=[]
path_arr=[]
elitegroup_diversity_arr=[]
elitegroup=[i[1] for i in (sort(collect(agents),by=x->abs(x[2]["perspective"])+x[2]["heuristic"])[1:group_size])]
for _ in 1:num_of_simulations
path=agent_based_simulation(elitegroup,adjacency,distance_matrix,agents)
distance=sum([distance_matrix[path[i],path[i+1]] for i in 1:(length(path)-1)])
elitegroup_diversity=Statistics.var([abs(agents[agent_id]["perspective"])+agents[agent_id]["heuristic"] for agent_id in elitegroup])
push!(distance_arr,distance)
push!(path_arr,length(path))
push!(elitegroup_diversity_arr,elitegroup_diversity)
end
println("Distance travelled:",Statistics.mean(distance_arr))
println("Length of the path:",Statistics.mean(path_arr))
println("Group diversity:",Statistics.mean(elitegroup_diversity_arr))
#apparently the elite group suffers from elite bias
#an elite agent has firm heuristic which makes it not very open to other options
#thus elite group spends a lot of time back and forth making no progress at all
path
# ### Discussion
#
#
#
# I was quite a bit shocked to learn that diversity trumps ability in this experiment! This is no cherry picking. If you run enough times of simulations, you will see the random group beats the elite group for most of the times.
#
# The problem with the technocracy is elite bias. As we have observed from the example, an elite has relatively objective perspective and fairly stable heuristic which is good for a deterministic problem. When facing a stochastic problem with random twists and turns, an elite agent's may not be very open to other available options. Now how do we tackle the bias issue? **Diversity!** Elites think alike. A democratic representation of the agents is no panacea but it cancels out the random noise caused by each agent's own perspective and heuristic. In short, **diversity trumps ability!**
# ### Further Reading
#
#
#
# 1. <NAME>, Page SE (1998)
#
# Diversity And Optimality
#
# https://citeseerx.ist.psu.edu/viewdoc/download?doi=10.1.1.80.8046&rep=rep1&type=pdf
#
# *This paper gives a clear definition of diversity and its relationship with local optima.*
#
#
# 2. Hong L, Page SE (2004)
#
# Groups Of Diverse Problem Solvers Can Outperform Groups Of High-ability Problem Solvers
#
# https://www.pnas.org/content/101/46/16385
#
# *One of the most influential papers on diversity. Our experiment is based upon this paper.*
#
#
# 3. <NAME>, Page SE (2001)
#
# Problem Solving By Heterogenous Agents
#
# https://sites.lsa.umich.edu/scottepage/wp-content/uploads/sites/344/2015/11/jet.pdf
#
# *This paper illustrates the application of diverse problem solving agents in the public funding problems*
#
#
# 4. Hong L, Page SE (2012)
#
# Some Microfoundations Of Collective Wisdom
#
# http://sites.lsa.umich.edu/wp-content/uploads/sites/344/2015/11/CollectWisdomParissubmitted.pdf
#
# *This paper proves the diversity prediction theorem from the perspective of statistics. The greater the diversity, the better the prediction*
#
#
# 5. <NAME> (2017)
#
# Collective Wisdom, Diversity And Misuse Of Mathematics
#
# https://www.cairn-int.info/publications-of-Houlou-Garcia-Antoine--66972.htm
#
# *This paper argues the experiment done by Hong and Page in 2004 is inapplicable to political science. The experiment made a poor mathematical inference.*
#
#
# 6. <NAME> (2014)
#
# Does Diversity Trump Ability?
#
# https://www.ams.org/notices/201409/rnoti-p1024.pdf
#
# *The author provides counter-example to challenge the computational experiment done by Hong and Page in 2004. The attempt to equate mathematical quantities with human attributes is inappropriate.*
| diversity trumps ability.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
import pandas as pd
df=pd.read_csv('ex/data/days-simulated-v2.tsv')
#target structure
df.head()
df=pd.read_html('ex/3.html')
import matplotlib.pyplot as plt
# %matplotlib inline
df[0].head()
df1=pd.read_csv('ex/1.csv')
df2=pd.read_csv('ex/2.csv')
df3=pd.read_csv('ex/3.csv')
df=pd.concat([df1,df2[1:],df3[1:]]) #no need for headers twice, df headers completely identical
hkoz=df[df.columns[9:489]].reset_index()
hetv=df[df.columns[489:969]].reset_index()
desc=df[df.columns[969:]].reset_index()
time=df[df.columns[2:4]].reset_index()
#top 16 activities
activities=[[u'Alvás'],
[u'Zuhany / Mosdó'],
[u'Étkezés', u'Étterem/Vendéglő'],
[u'Munka (irodai)', u'Munka (kétkezi)'],
[u'Internet', u'Telefon/Chat/Facebook'],
[u'Vásárlás'],
[u'Vallásgyakorlás'],
[u'TV/Film', u'Mozi'],
[u'Olvasás', u'Újság/Keresztrejtvény'],
[u'Házimunka/Gyerekfelügyelet'],
[u'Hivatalos elintéznivalók'],
[u'Sport', u'Edzőterem/Szépségszalon'],
[u'Utazás/Vezetés'],
[u'Tanulás', u'Magánóra'],
[u'Szórakozóhely/Kávézó/Pub'],
[u'Séta/Kutyasétáltatás', u'Természet/Kirándulás'],
[u'Egyéb Hobby', u'PC játék', u'Önkéntesség', u'Kertészkedés/Barkácsolás', u'Rokonlátogatás', u'Más']]
'Alv\xc3\xa1s',
'Edz\xc5\x91terem/Sz\xc3\xa9ps\xc3\xa9gszalon',
'Egy\xc3\xa9b Hobby',
'Hivatalos elint\xc3\xa9znival\xc3\xb3k',
'H\xc3\xa1zimunka/Gyerekfel\xc3\xbcgyelet',
'Internet',
'Kert\xc3\xa9szked\xc3\xa9s/Bark\xc3\xa1csol\xc3\xa1s',
'Mag\xc3\xa1n\xc3\xb3ra',
'Mozi',
'Munka (irodai)',
'Munka (k\xc3\xa9tkezi)',
'M\xc3\xa1s',
'Olvas\xc3\xa1s',
'PC j\xc3\xa1t\xc3\xa9k',
'Rokonl\xc3\xa1togat\xc3\xa1s',
'Sport',
'Sz\xc3\xb3rakoz\xc3\xb3hely/K\xc3\xa1v\xc3\xa9z\xc3\xb3/Pub',
'S\xc3\xa9ta/Kutyas\xc3\xa9t\xc3\xa1ltat\xc3\xa1s',
'TV/Film',
'Tanul\xc3\xa1s',
'Telefon/Chat/Facebook',
'Term\xc3\xa9szet/Kir\xc3\xa1ndul\xc3\xa1s',
'Utaz\xc3\xa1s/Vezet\xc3\xa9s',
'Vall\xc3\xa1sgyakorl\xc3\xa1s',
'V\xc3\xa1s\xc3\xa1rl\xc3\xa1s',
'Zuhany / Mosd\xc3\xb3',
'\xc3\x89tkez\xc3\xa9s',
'\xc3\x89tterem/Vend\xc3\xa9gl\xc5\x91',
'\xc3\x96nk\xc3\xa9ntess\xc3\xa9g',
'\xc3\x9ajs\xc3\xa1g/Keresztrejtv\xc3\xa9ny'
actidict={}
for i in range(len(activities)):
for j in range(len(activities[i])):
actidict[activities[i][j]]=i
actidict
# run only once
hkoz.columns=hkoz.loc[0].values
hkoz=hkoz[1:].drop(0,axis=1)
for i in hkoz.columns[:10]:
print i
activity=i[:i.find('-')-1]
timeslice=i[i.find('-')+2:]
print activity,timeslice
hkozdata={}
for i in hkoz.index:
index=hkoz.loc[i].index
values=hkoz.loc[i].values
helper=[]
for j in range(len(values)):
if str(values[j]).lower()!='nan':
helper.append(index[j])
hkozdata[i]=helper
hkozdata[1]
j=1
timematrix={}
for i in hkozdata[j]:
activity=i[:i.find('-')-1]
timeslice=i[i.find('-')+2:]
if timeslice not in timematrix:timematrix[timeslice]=[]
timematrix[timeslice].append(actidict[activity])
set([i[:i.find('-')-1] for i in hkoz.columns])
actidict
timematrix
| rutin/Untitled-Copy1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
from pyvista import set_plot_theme
set_plot_theme('document')
# Compute Gradients of a Field
# ============================
#
# Estimate the gradient of a scalar or vector field in a data set.
#
# The ordering for the output gradient tuple will be {du/dx, du/dy, du/dz,
# dv/dx, dv/dy, dv/dz, dw/dx, dw/dy, dw/dz} for an input array {u, v, w}.
#
# Showing the
# `pyvista.DataSetFilters.compute_derivative`{.interpreted-text
# role="func"} filter.
#
# +
# sphinx_gallery_thumbnail_number = 1
import pyvista as pv
from pyvista import examples
import numpy as np
# A vtkStructuredGrid - but could be any mesh type
mesh = examples.download_carotid()
mesh
# -
# Now compute the gradients of the `vectors` vector field in the point
# data of that mesh. This is as simple as calling
# `pyvista.DataSetFilters.compute_derivative`{.interpreted-text
# role="func"}.
#
mesh_g = mesh.compute_derivative(scalars="vectors")
mesh_g["gradient"]
# ::: {.note}
# ::: {.admonition-title}
# Note
# :::
#
# You can also use
# `pyvista.DataSetFilters.compute_derivative`{.interpreted-text
# role="func"} for computing other derivative based quantities, such as
# divergence, vorticity, and Q-criterion. See function documentation for
# options.
# :::
#
# `mesh_g["gradient"]` is an `N` by 9 NumPy array of the gradients, so we
# could make a dictionary of NumPy arrays of the gradients like:
#
# +
def gradients_to_dict(arr):
"""A helper method to label the gradients into a dictionary."""
keys = np.array(["du/dx", "du/dy", "du/dz", "dv/dx", "dv/dy", "dv/dz", "dw/dx", "dw/dy", "dw/dz"])
keys = keys.reshape((3,3))[:,:arr.shape[1]].ravel()
return dict(zip(keys, mesh_g["gradient"].T))
gradients = gradients_to_dict(mesh_g["gradient"])
gradients
# -
# And we can add all of those components as individual arrays back to the
# mesh by:
#
mesh_g.point_arrays.update(gradients)
mesh_g
# +
keys = np.array(list(gradients.keys())).reshape(3,3)
p = pv.Plotter(shape=keys.shape)
for i in range(keys.shape[0]):
for j in range(keys.shape[1]):
name = keys[i,j]
p.subplot(i,j)
p.add_mesh(mesh_g.contour(scalars=name), scalars=name, opacity=0.75)
p.add_mesh(mesh_g.outline(), color="k")
p.link_views()
p.view_isometric()
p.show()
# -
# And there you have it, the gradients for a vector field! We could also
# do this for a scalar field like for the `scalars` field in the given
# dataset.
#
# +
mesh_g = mesh.compute_derivative(scalars="scalars")
gradients = gradients_to_dict(mesh_g["gradient"])
gradients
# +
mesh_g.point_arrays.update(gradients)
keys = np.array(list(gradients.keys())).reshape(1,3)
p = pv.Plotter(shape=keys.shape)
for i in range(keys.shape[0]):
for j in range(keys.shape[1]):
name = keys[i,j]
p.subplot(i,j)
p.add_mesh(mesh_g.contour(scalars=name), scalars=name, opacity=0.75)
p.add_mesh(mesh_g.outline(), color="k")
p.link_views()
p.view_isometric()
p.show()
| examples/01-filter/gradients.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Dependencies
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _kg_hide-input=true _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5"
import json, warnings, shutil
from tweet_utility_scripts import *
from tweet_utility_preprocess_roberta_scripts import *
from transformers import TFRobertaModel, RobertaConfig
from tokenizers import ByteLevelBPETokenizer
from tensorflow.keras.models import Model
from tensorflow.keras import optimizers, metrics, losses, layers
from tensorflow.keras.callbacks import EarlyStopping, ModelCheckpoint, LearningRateScheduler
SEED = 0
seed_everything(SEED)
warnings.filterwarnings("ignore")
# + _kg_hide-input=true
class DecoupledWeightDecayExtension(object):
"""This class allows to extend optimizers with decoupled weight decay.
It implements the decoupled weight decay described by Loshchilov & Hutter
(https://arxiv.org/pdf/1711.05101.pdf), in which the weight decay is
decoupled from the optimization steps w.r.t. to the loss function.
For SGD variants, this simplifies hyperparameter search since it decouples
the settings of weight decay and learning rate.
For adaptive gradient algorithms, it regularizes variables with large
gradients more than L2 regularization would, which was shown to yield
better training loss and generalization error in the paper above.
This class alone is not an optimizer but rather extends existing
optimizers with decoupled weight decay. We explicitly define the two
examples used in the above paper (SGDW and AdamW), but in general this
can extend any OptimizerX by using
`extend_with_decoupled_weight_decay(
OptimizerX, weight_decay=weight_decay)`.
In order for it to work, it must be the first class the Optimizer with
weight decay inherits from, e.g.
```python
class AdamW(DecoupledWeightDecayExtension, tf.keras.optimizers.Adam):
def __init__(self, weight_decay, *args, **kwargs):
super(AdamW, self).__init__(weight_decay, *args, **kwargs).
```
Note: this extension decays weights BEFORE applying the update based
on the gradient, i.e. this extension only has the desired behaviour for
optimizers which do not depend on the value of'var' in the update step!
Note: when applying a decay to the learning rate, be sure to manually apply
the decay to the `weight_decay` as well. For example:
```python
step = tf.Variable(0, trainable=False)
schedule = tf.optimizers.schedules.PiecewiseConstantDecay(
[10000, 15000], [1e-0, 1e-1, 1e-2])
# lr and wd can be a function or a tensor
lr = 1e-1 * schedule(step)
wd = lambda: 1e-4 * schedule(step)
# ...
optimizer = tfa.optimizers.AdamW(learning_rate=lr, weight_decay=wd)
```
"""
def __init__(self, weight_decay, **kwargs):
"""Extension class that adds weight decay to an optimizer.
Args:
weight_decay: A `Tensor` or a floating point value, the factor by
which a variable is decayed in the update step.
**kwargs: Optional list or tuple or set of `Variable` objects to
decay.
"""
wd = kwargs.pop('weight_decay', weight_decay)
super(DecoupledWeightDecayExtension, self).__init__(**kwargs)
self._decay_var_list = None # is set in minimize or apply_gradients
self._set_hyper('weight_decay', wd)
def get_config(self):
config = super(DecoupledWeightDecayExtension, self).get_config()
config.update({
'weight_decay':
self._serialize_hyperparameter('weight_decay'),
})
return config
def minimize(self,
loss,
var_list,
grad_loss=None,
name=None,
decay_var_list=None):
"""Minimize `loss` by updating `var_list`.
This method simply computes gradient using `tf.GradientTape` and calls
`apply_gradients()`. If you want to process the gradient before
applying then call `tf.GradientTape` and `apply_gradients()` explicitly
instead of using this function.
Args:
loss: A callable taking no arguments which returns the value to
minimize.
var_list: list or tuple of `Variable` objects to update to
minimize `loss`, or a callable returning the list or tuple of
`Variable` objects. Use callable when the variable list would
otherwise be incomplete before `minimize` since the variables
are created at the first time `loss` is called.
grad_loss: Optional. A `Tensor` holding the gradient computed for
`loss`.
decay_var_list: Optional list of variables to be decayed. Defaults
to all variables in var_list.
name: Optional name for the returned operation.
Returns:
An Operation that updates the variables in `var_list`. If
`global_step` was not `None`, that operation also increments
`global_step`.
Raises:
ValueError: If some of the variables are not `Variable` objects.
"""
self._decay_var_list = set(decay_var_list) if decay_var_list else False
return super(DecoupledWeightDecayExtension, self).minimize(
loss, var_list=var_list, grad_loss=grad_loss, name=name)
def apply_gradients(self, grads_and_vars, name=None, decay_var_list=None):
"""Apply gradients to variables.
This is the second part of `minimize()`. It returns an `Operation` that
applies gradients.
Args:
grads_and_vars: List of (gradient, variable) pairs.
name: Optional name for the returned operation. Default to the
name passed to the `Optimizer` constructor.
decay_var_list: Optional list of variables to be decayed. Defaults
to all variables in var_list.
Returns:
An `Operation` that applies the specified gradients. If
`global_step` was not None, that operation also increments
`global_step`.
Raises:
TypeError: If `grads_and_vars` is malformed.
ValueError: If none of the variables have gradients.
"""
self._decay_var_list = set(decay_var_list) if decay_var_list else False
return super(DecoupledWeightDecayExtension, self).apply_gradients(
grads_and_vars, name=name)
def _decay_weights_op(self, var):
if not self._decay_var_list or var in self._decay_var_list:
return var.assign_sub(
self._get_hyper('weight_decay', var.dtype) * var,
self._use_locking)
return tf.no_op()
def _decay_weights_sparse_op(self, var, indices):
if not self._decay_var_list or var in self._decay_var_list:
update = (-self._get_hyper('weight_decay', var.dtype) * tf.gather(
var, indices))
return self._resource_scatter_add(var, indices, update)
return tf.no_op()
# Here, we overwrite the apply functions that the base optimizer calls.
# super().apply_x resolves to the apply_x function of the BaseOptimizer.
def _resource_apply_dense(self, grad, var):
with tf.control_dependencies([self._decay_weights_op(var)]):
return super(DecoupledWeightDecayExtension,
self)._resource_apply_dense(grad, var)
def _resource_apply_sparse(self, grad, var, indices):
decay_op = self._decay_weights_sparse_op(var, indices)
with tf.control_dependencies([decay_op]):
return super(DecoupledWeightDecayExtension,
self)._resource_apply_sparse(grad, var, indices)
def extend_with_decoupled_weight_decay(base_optimizer):
"""Factory function returning an optimizer class with decoupled weight
decay.
Returns an optimizer class. An instance of the returned class computes the
update step of `base_optimizer` and additionally decays the weights.
E.g., the class returned by
`extend_with_decoupled_weight_decay(tf.keras.optimizers.Adam)` is
equivalent to `tfa.optimizers.AdamW`.
The API of the new optimizer class slightly differs from the API of the
base optimizer:
- The first argument to the constructor is the weight decay rate.
- `minimize` and `apply_gradients` accept the optional keyword argument
`decay_var_list`, which specifies the variables that should be decayed.
If `None`, all variables that are optimized are decayed.
Usage example:
```python
# MyAdamW is a new class
MyAdamW = extend_with_decoupled_weight_decay(tf.keras.optimizers.Adam)
# Create a MyAdamW object
optimizer = MyAdamW(weight_decay=0.001, learning_rate=0.001)
# update var1, var2 but only decay var1
optimizer.minimize(loss, var_list=[var1, var2], decay_variables=[var1])
Note: this extension decays weights BEFORE applying the update based
on the gradient, i.e. this extension only has the desired behaviour for
optimizers which do not depend on the value of 'var' in the update step!
Note: when applying a decay to the learning rate, be sure to manually apply
the decay to the `weight_decay` as well. For example:
```python
step = tf.Variable(0, trainable=False)
schedule = tf.optimizers.schedules.PiecewiseConstantDecay(
[10000, 15000], [1e-0, 1e-1, 1e-2])
# lr and wd can be a function or a tensor
lr = 1e-1 * schedule(step)
wd = lambda: 1e-4 * schedule(step)
# ...
optimizer = tfa.optimizers.AdamW(learning_rate=lr, weight_decay=wd)
```
Note: you might want to register your own custom optimizer using
`tf.keras.utils.get_custom_objects()`.
Args:
base_optimizer: An optimizer class that inherits from
tf.optimizers.Optimizer.
Returns:
A new optimizer class that inherits from DecoupledWeightDecayExtension
and base_optimizer.
"""
class OptimizerWithDecoupledWeightDecay(DecoupledWeightDecayExtension,
base_optimizer):
"""Base_optimizer with decoupled weight decay.
This class computes the update step of `base_optimizer` and
additionally decays the variable with the weight decay being
decoupled from the optimization steps w.r.t. to the loss
function, as described by Loshchilov & Hutter
(https://arxiv.org/pdf/1711.05101.pdf). For SGD variants, this
simplifies hyperparameter search since it decouples the settings
of weight decay and learning rate. For adaptive gradient
algorithms, it regularizes variables with large gradients more
than L2 regularization would, which was shown to yield better
training loss and generalization error in the paper above.
"""
def __init__(self, weight_decay, *args, **kwargs):
# super delegation is necessary here
super(OptimizerWithDecoupledWeightDecay, self).__init__(
weight_decay, *args, **kwargs)
return OptimizerWithDecoupledWeightDecay
class AdamW(DecoupledWeightDecayExtension, tf.keras.optimizers.Adam):
"""Optimizer that implements the Adam algorithm with weight decay.
This is an implementation of the AdamW optimizer described in "Decoupled
Weight Decay Regularization" by Loshchilov & Hutter
(https://arxiv.org/abs/1711.05101)
([pdf])(https://arxiv.org/pdf/1711.05101.pdf).
It computes the update step of `tf.keras.optimizers.Adam` and additionally
decays the variable. Note that this is different from adding L2
regularization on the variables to the loss: it regularizes variables with
large gradients more than L2 regularization would, which was shown to yield
better training loss and generalization error in the paper above.
For further information see the documentation of the Adam Optimizer.
This optimizer can also be instantiated as
```python
extend_with_decoupled_weight_decay(tf.keras.optimizers.Adam,
weight_decay=weight_decay)
```
Note: when applying a decay to the learning rate, be sure to manually apply
the decay to the `weight_decay` as well. For example:
```python
step = tf.Variable(0, trainable=False)
schedule = tf.optimizers.schedules.PiecewiseConstantDecay(
[10000, 15000], [1e-0, 1e-1, 1e-2])
# lr and wd can be a function or a tensor
lr = 1e-1 * schedule(step)
wd = lambda: 1e-4 * schedule(step)
# ...
optimizer = tfa.optimizers.AdamW(learning_rate=lr, weight_decay=wd)
```
"""
def __init__(self,
weight_decay,
learning_rate=0.001,
beta_1=0.9,
beta_2=0.999,
epsilon=1e-07,
amsgrad=False,
name="AdamW",
**kwargs):
"""Construct a new AdamW optimizer.
For further information see the documentation of the Adam Optimizer.
Args:
weight_decay: A Tensor or a floating point value. The weight decay.
learning_rate: A Tensor or a floating point value. The learning
rate.
beta_1: A float value or a constant float tensor. The exponential
decay rate for the 1st moment estimates.
beta_2: A float value or a constant float tensor. The exponential
decay rate for the 2nd moment estimates.
epsilon: A small constant for numerical stability. This epsilon is
"epsilon hat" in the Kingma and Ba paper (in the formula just
before Section 2.1), not the epsilon in Algorithm 1 of the
paper.
amsgrad: boolean. Whether to apply AMSGrad variant of this
algorithm from the paper "On the Convergence of Adam and
beyond".
name: Optional name for the operations created when applying
gradients. Defaults to "AdamW".
**kwargs: keyword arguments. Allowed to be {`clipnorm`,
`clipvalue`, `lr`, `decay`}. `clipnorm` is clip gradients by
norm; `clipvalue` is clip gradients by value, `decay` is
included for backward compatibility to allow time inverse decay
of learning rate. `lr` is included for backward compatibility,
recommended to use `learning_rate` instead.
"""
super(AdamW, self).__init__(
weight_decay,
learning_rate=learning_rate,
beta_1=beta_1,
beta_2=beta_2,
epsilon=epsilon,
amsgrad=amsgrad,
name=name,
**kwargs)
# -
# # Load data
# + _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" _kg_hide-input=true _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a"
database_base_path = '/kaggle/input/tweet-dataset-split-roberta-base-96/'
k_fold = pd.read_csv(database_base_path + '5-fold.csv')
display(k_fold.head())
# Unzip files
# !tar -xvf /kaggle/input/tweet-dataset-split-roberta-base-96/fold_1.tar.gz
# !tar -xvf /kaggle/input/tweet-dataset-split-roberta-base-96/fold_2.tar.gz
# !tar -xvf /kaggle/input/tweet-dataset-split-roberta-base-96/fold_3.tar.gz
# !tar -xvf /kaggle/input/tweet-dataset-split-roberta-base-96/fold_4.tar.gz
# !tar -xvf /kaggle/input/tweet-dataset-split-roberta-base-96/fold_5.tar.gz
# -
# # Model parameters
# +
vocab_path = database_base_path + 'vocab.json'
merges_path = database_base_path + 'merges.txt'
base_path = '/kaggle/input/qa-transformers/roberta/'
config = {
"MAX_LEN": 96,
"BATCH_SIZE": 32,
"EPOCHS": 5,
"LEARNING_RATE": 3e-5,
"ES_PATIENCE": 1,
"question_size": 4,
"N_FOLDS": 5,
"base_model_path": base_path + 'roberta-base-tf_model.h5',
"config_path": base_path + 'roberta-base-config.json'
}
with open('config.json', 'w') as json_file:
json.dump(json.loads(json.dumps(config)), json_file)
# -
# # Tokenizer
# + _kg_hide-output=true
tokenizer = ByteLevelBPETokenizer(vocab_file=vocab_path, merges_file=merges_path,
lowercase=True, add_prefix_space=True)
tokenizer.save('./')
# -
# ## Learning rate schedule
# + _kg_hide-input=true
LR_MIN = 1e-6
LR_MAX = config['LEARNING_RATE']
lr_start=1e-4
warmup_epochs=0
hold_max_epochs=0
num_cycles=0.5
total_epochs = config['EPOCHS']
@tf.function
def lrfn(epoch):
if epoch < warmup_epochs:
lr = (LR_MAX - lr_start) / (warmup_epochs * epoch + lr_start)
elif epoch < warmup_epochs + hold_max_epochs:
lr = LR_MAX
else:
progress = (epoch - warmup_epochs - hold_max_epochs) / (total_epochs - warmup_epochs - hold_max_epochs)
lr = LR_MAX * (0.5 * (1.0 + tf.math.cos(np.pi * num_cycles * 2.0 * progress)))
# if LR_MIN is not None:
# lr = max(LR_MIN, lr)
return lr
rng = [i for i in range(config['EPOCHS'])]
y = [lrfn(x) for x in rng]
fig, ax = plt.subplots(figsize=(20, 6))
plt.plot(rng, y)
print("Learning rate schedule: {:.3g} to {:.3g} to {:.3g}".format(y[0], max(y), y[-1]))
# -
# # Model
# +
module_config = RobertaConfig.from_pretrained(config['config_path'], output_hidden_states=False)
def model_fn(MAX_LEN):
input_ids = layers.Input(shape=(MAX_LEN,), dtype=tf.int32, name='input_ids')
attention_mask = layers.Input(shape=(MAX_LEN,), dtype=tf.int32, name='attention_mask')
base_model = TFRobertaModel.from_pretrained(config['base_model_path'], config=module_config, name="base_model")
last_hidden_state, _ = base_model({'input_ids': input_ids, 'attention_mask': attention_mask})
x = layers.Dropout(.1)(last_hidden_state)
x_start = layers.Dense(1)(last_hidden_state)
x_start = layers.Flatten()(x_start)
y_start = layers.Activation('softmax', dtype='float32', name='y_start')(x_start)
x_end = layers.Dense(1)(last_hidden_state)
x_end = layers.Flatten()(x_end)
y_end = layers.Activation('softmax', dtype='float32', name='y_end')(x_end)
model = Model(inputs=[input_ids, attention_mask], outputs=[y_start, y_end])
return model
# -
# # Train
# + _kg_hide-input=true _kg_hide-output=true
AUTO = tf.data.experimental.AUTOTUNE
strategy = tf.distribute.get_strategy()
history_list = []
for n_fold in range(config['N_FOLDS']):
n_fold +=1
print('\nFOLD: %d' % (n_fold))
# Load data
base_data_path = 'fold_%d/' % (n_fold)
x_train = np.load(base_data_path + 'x_train.npy')
y_train = np.load(base_data_path + 'y_train.npy')
x_valid = np.load(base_data_path + 'x_valid.npy')
y_valid = np.load(base_data_path + 'y_valid.npy')
step_size = x_train.shape[1] // config['BATCH_SIZE']
valid_step_size = x_valid.shape[1] // config['BATCH_SIZE']
### Delete data dir
# shutil.rmtree(base_data_path)
# Build TF datasets
train_dist_ds = strategy.experimental_distribute_dataset(get_training_dataset(x_train, y_train, config['BATCH_SIZE'], AUTO, seed=SEED))
valid_dist_ds = strategy.experimental_distribute_dataset(get_validation_dataset(x_valid, y_valid, config['BATCH_SIZE'], AUTO, repeated=True, seed=SEED))
train_data_iter = iter(train_dist_ds)
valid_data_iter = iter(valid_dist_ds)
# Step functions
@tf.function
def train_step(data_iter):
def train_step_fn(x, y):
with tf.GradientTape() as tape:
probabilities = model(x, training=True)
loss_start = loss_fn_start(y['y_start'], probabilities[0], label_smoothing=0.2)
loss_end = loss_fn_end(y['y_end'], probabilities[1], label_smoothing=0.2)
loss = tf.math.add(loss_start, loss_end)
grads = tape.gradient(loss, model.trainable_variables)
optimizer.apply_gradients(zip(grads, model.trainable_variables))
# update metrics
train_acc_start.update_state(y['y_start'], probabilities)
train_acc_end.update_state(y['y_end'], probabilities)
train_loss.update_state(loss)
train_loss_start.update_state(loss_start)
train_loss_end.update_state(loss_end)
for _ in tf.range(step_size):
strategy.experimental_run_v2(train_step_fn, next(data_iter))
@tf.function
def valid_step(data_iter):
def valid_step_fn(x, y):
probabilities = model(x, training=False)
loss_start = loss_fn_start(y['y_start'], probabilities[0])
loss_end = loss_fn_end(y['y_end'], probabilities[1])
loss = tf.math.add(loss_start, loss_end)
# update metrics
valid_acc_start.update_state(y['y_start'], probabilities)
valid_acc_end.update_state(y['y_end'], probabilities)
valid_loss.update_state(loss)
valid_loss_start.update_state(loss_start)
valid_loss_end.update_state(loss_end)
for _ in tf.range(valid_step_size):
strategy.experimental_run_v2(valid_step_fn, next(data_iter))
# Train model
model_path = 'model_fold_%d.h5' % (n_fold)
model = model_fn(config['MAX_LEN'])
# optimizer = optimizers.Adam(learning_rate=lambda: lrfn(tf.cast(optimizer.iterations, tf.float32)//step_size))
optimizer = optimizers.Adam(learning_rate=lambda: lrfn(tf.cast(optimizer.iterations, tf.float32)//step_size))
optimizer = AdamW(learning_rate=lambda: lrfn(tf.cast(optimizer.iterations, tf.float32)//step_size),
weight_decay=lambda : lrfn(tf.cast(optimizer.iterations, tf.float32)//step_size))
optimizer.weight_decay = lambda : lrfn(tf.cast(optimizer.iterations, tf.float32)//step_size) # Making sure wdecay is used
loss_fn_start = losses.categorical_crossentropy
loss_fn_end = losses.categorical_crossentropy
train_acc_start = metrics.CategoricalAccuracy()
valid_acc_start = metrics.CategoricalAccuracy()
train_acc_end = metrics.CategoricalAccuracy()
valid_acc_end = metrics.CategoricalAccuracy()
train_loss = metrics.Sum()
valid_loss = metrics.Sum()
train_loss_start = metrics.Sum()
valid_loss_start = metrics.Sum()
train_loss_end = metrics.Sum()
valid_loss_end = metrics.Sum()
metrics_dict = {'loss': train_loss, 'loss_start': train_loss_start, 'loss_end': train_loss_end,
'acc_start': train_acc_start, 'acc_end': train_acc_end,
'val_loss': valid_loss, 'val_loss_start': valid_loss_start, 'val_loss_end': valid_loss_end,
'val_acc_start': valid_acc_start, 'val_acc_end': valid_acc_end}
history = custom_fit(model, metrics_dict, train_step, valid_step, train_data_iter, valid_data_iter,
step_size, valid_step_size, config['BATCH_SIZE'], config['EPOCHS'], config['ES_PATIENCE'], model_path)
history_list.append(history)
model.load_weights(model_path)
# Make predictions
train_preds = model.predict(get_test_dataset(x_train, config['BATCH_SIZE']))
valid_preds = model.predict(get_test_dataset(x_valid, config['BATCH_SIZE']))
k_fold.loc[k_fold['fold_%d' % (n_fold)] == 'train', 'start_fold_%d' % (n_fold)] = train_preds[0].argmax(axis=-1)
k_fold.loc[k_fold['fold_%d' % (n_fold)] == 'train', 'end_fold_%d' % (n_fold)] = train_preds[1].argmax(axis=-1)
k_fold.loc[k_fold['fold_%d' % (n_fold)] == 'validation', 'start_fold_%d' % (n_fold)] = valid_preds[0].argmax(axis=-1)
k_fold.loc[k_fold['fold_%d' % (n_fold)] == 'validation', 'end_fold_%d' % (n_fold)] = valid_preds[1].argmax(axis=-1)
k_fold['end_fold_%d' % (n_fold)] = k_fold['end_fold_%d' % (n_fold)].astype(int)
k_fold['start_fold_%d' % (n_fold)] = k_fold['start_fold_%d' % (n_fold)].astype(int)
k_fold['end_fold_%d' % (n_fold)].clip(0, k_fold['text_len'], inplace=True)
k_fold['start_fold_%d' % (n_fold)].clip(0, k_fold['end_fold_%d' % (n_fold)], inplace=True)
k_fold['prediction_fold_%d' % (n_fold)] = k_fold.apply(lambda x: decode(x['start_fold_%d' % (n_fold)], x['end_fold_%d' % (n_fold)], x['text'], config['question_size'], tokenizer), axis=1)
k_fold['prediction_fold_%d' % (n_fold)].fillna(k_fold["text"], inplace=True)
k_fold['jaccard_fold_%d' % (n_fold)] = k_fold.apply(lambda x: jaccard(x['selected_text'], x['prediction_fold_%d' % (n_fold)]), axis=1)
# -
# # Model loss graph
# + _kg_hide-input=true
sns.set(style="whitegrid")
for n_fold in range(config['N_FOLDS']):
print('Fold: %d' % (n_fold+1))
plot_metrics(history_list[n_fold])
# -
# # Model evaluation
# + _kg_hide-input=true
display(evaluate_model_kfold(k_fold, config['N_FOLDS']).style.applymap(color_map))
# -
# # Visualize predictions
# + _kg_hide-input=true
display(k_fold[[c for c in k_fold.columns if not (c.startswith('textID') or
c.startswith('text_len') or
c.startswith('selected_text_len') or
c.startswith('text_wordCnt') or
c.startswith('selected_text_wordCnt') or
c.startswith('fold_') or
c.startswith('start_fold_') or
c.startswith('end_fold_'))]].head(15))
| Model backlog/Train/122-tweet-train-5fold-roberta-base-adamw-tf.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data Preparation
# +
#Import packages
import pandas as pd
import csv
import matplotlib.pyplot as plt
import seaborn as sns
import warnings
import numpy as np
import folium
#from PIL import Image
from sklearn.preprocessing import MinMaxScaler
from scipy import stats
from scipy.stats import anderson
from scipy.stats import norm
from matplotlib import pylab
pd.set_option('display.max_rows', None)
pd.set_option('display.max_columns', None)
warnings.filterwarnings('ignore')
# %matplotlib inline
# -
#import dataset created in Data Understanding
accidents = pd.read_csv(r"C:\Users\DETCAO03\V-Case study\02_Dataset\Used\Accidents.csv",low_memory=False, encoding='utf-8')
accidents.head()
# ## Select features clean and construct data
accidents.drop(['Location_Easting_OSGR', 'Location_Northing_OSGR', 'Accident_Index', 'LSOA_of_Accident_Location', 'Police_Force', 'Local_Authority_(District)',
'Local_Authority_(Highway)', 'Junction_Detail', '2nd_Road_Class', '2nd_Road_Number',
'Did_Police_Officer_Attend_Scene_of_Accident', '1st_Road_Number',
'Pedestrian_Crossing-Physical_Facilities'], axis=1, inplace=True)
# +
#date
#accidents['Date_time'] = accidents['Date'] +' '+ accidents['Time']
#accidents['Date_time'] = pd.to_datetime(accidents["Date_time"])
accidents.drop(['Date','Time'],axis =1 , inplace=True)
#coordinates
#accidents["LatLon"] = list(zip(accidents["Latitude"], accidents["Longitude"]))
#accidents.drop(['Latitude','Longitude'],axis=1 , inplace=True)
#fill missing region values with "mode" and binary encoding of region
accidents["Region"] = accidents["Region"].fillna(accidents["Region"].mode()[0])
accidents = pd.get_dummies(accidents, columns=['Region'], drop_first=True)
#drop missing values (151)
accidents.dropna(inplace=True)
#Normalisation
scaler = MinMaxScaler()
accidents[["Latitude", "Longitude", "Number_of_Vehicles","Number_of_Casualties","Day_of_Week","1st_Road_Class","Road_Type",
"Speed_limit","Junction_Control","Pedestrian_Crossing-Human_Control","Light_Conditions",
"Weather_Conditions","Road_Surface_Conditions","Special_Conditions_at_Site","Carriageway_Hazards",
"Urban_or_Rural_Area"]] = scaler.fit_transform(accidents[["Latitude", "Longitude", "Number_of_Vehicles","Number_of_Casualties","Day_of_Week","1st_Road_Class","Road_Type","Speed_limit","Junction_Control","Pedestrian_Crossing-Human_Control","Light_Conditions","Weather_Conditions","Road_Surface_Conditions","Special_Conditions_at_Site","Carriageway_Hazards","Urban_or_Rural_Area"]])
# -
accidents.head()
accidents.shape
accidents.dtypes
#final dataset for Modeling
accidents.to_csv(r"C:\Users\DETCAO03\V-Case study\02_Dataset\Used\Cleaned_dataset_accidents.csv", index=False, encoding='utf-8')
# # Modeling
from sklearn.model_selection import train_test_split
from sklearn.naive_bayes import MultinomialNB
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import accuracy_score
from sklearn.metrics import classification_report
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.svm import SVC, LinearSVC
from sklearn.metrics import log_loss
from sklearn.metrics import accuracy_score, f1_score, precision_score, recall_score, classification_report, confusion_matrix, cohen_kappa_score
from sklearn.neighbors import KNeighborsClassifier
accidents = pd.read_csv(r"C:\Users\DETCAO03\V-Case study\02_Dataset\Used\Cleaned_dataset_accidents.csv",low_memory=False, encoding='utf-8')
#define influencing and response variable
X = accidents.drop("Accident_Severity", axis=1)
y = accidents["Accident_Severity"]
# Split the data into a training and test set.
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.30, random_state=0)
# ## 1st Round
# ### Decision Tree
# +
dt = DecisionTreeClassifier(criterion = 'gini', min_samples_split = 30, splitter='best')
dt = dt.fit(X_train, y_train)
y_pred = dt.predict(X_test)
acc_decision = round(dt.score(X_test, y_test) * 100, 2)
sk_report = classification_report(digits = 6, y_true = y_test, y_pred = y_pred)
print("Cohen Kappa: "+str(cohen_kappa_score(y_test,y_pred)))
print("Accuracy", acc_decision)
print("\n")
print(sk_report)
### Confusion Matrix
pd.crosstab(y_test, y_pred, rownames=['Actual'], colnames=['Predicted'], margins=True)
# -
# ### Logistic Regression
# +
clf = LogisticRegression().fit(X_train, y_train)
y_pred = clf.predict(X_test)
sk_report = classification_report(digits = 6, y_true = y_test, y_pred = y_pred)
print("Accuracy", round(accuracy_score(y_test, y_pred) * 100,2))
print("Cohen Kappa: "+str(cohen_kappa_score(y_test,y_pred)))
print("\n")
print(sk_report)
### Confusion Matrix
pd.crosstab(y_test, y_pred, rownames=['Actual'], colnames=['Predicted'], margins=True)
# -
# ### KNN
# +
knn = KNeighborsClassifier()
knn.fit(X_train, y_train)
y_pred = knn.predict(X_test)
sk_report = classification_report(digits = 6, y_true = y_test, y_pred = y_pred)
print("Accuracy", round(accuracy_score(y_test, y_pred) * 100,2))
print("Cohen Kappa: "+str(cohen_kappa_score(y_test,y_pred)))
print("\n")
print(sk_report)
### Confusion Matrix
pd.crosstab(y_test, y_pred, rownames=['Actual'], colnames=['Predicted'], margins=True)
# -
# ### Support Vector Machines
# +
svm = SVC()
svm.fit(X_train, y_train)
y_pred = svm.predict(X_test)
sk_report = classification_report(digits = 6, y_true = y_test, y_pred = y_pred)
print("Accuracy", round(accuracy_score(y_test, y_pred) * 100,2))
print("Cohen Kappa: "+str(cohen_kappa_score(y_test,y_pred)))
print("\n")
print(sk_report)
### Confusion Matrix
pd.crosstab(y_test, y_pred, rownames=['Actual'], colnames=['Predicted'], margins=True)
| 03_DU+DP+M/Data Preparation and Modeling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Amazon Customer Data Wrangling
# Data from
# > Justifying recommendations using distantly-labeled reviews and fined-grained aspects
# <NAME>, <NAME>, <NAME>
# Empirical Methods in Natural Language Processing (EMNLP), 2019
# +
# import packages
import spacy
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
# load English tokenizer, tagger, parser, NER, and word vectors
nlp = spacy.load('en_core_web_sm')
# koalas
#import databricks.koalas as ks
# spark tools
from pyspark import SparkConf, SparkContext
from pyspark.ml.stat import Summarizer
from pyspark.sql import Row, SQLContext
from pyspark.sql.functions import isnan, when, count, col, countDistinct
# -
# initiate spark context
conf = SparkConf().setMaster('local[*]').setAppName('AmazonDataWrangling')
sc = SparkContext.getOrCreate(conf=conf)
sqlContext = SQLContext(sc)
# ## Skip to Spark DataFrame Section
# +
# read in the data to a koalas dataframe
#bks_df = ks.read_json('../Amazon_Data/Books.json.gz', compression='gzip')
# +
# inspect the column
#bks_df.columns
# +
# inspect the shape
#bks_df.shape
# +
#bks_df.groupby('overall')['asin'].count()
# +
#bks_chunks = ks.read_json('../Amazon_Data/Books.json.gz', compression='gzip', chunksize=1000)
# +
#pds_chunks = pd.read_json('../Amazon_Data/Books.json.gz', lines=True,
# compression='gzip', chunksize=1000)
# +
#help(pds_chunks)
# -
# +
# convert the star ratings to a numpy array
# getting memory/space errors
# same issues with running .head()
#overall_np = bks_df.overall.to_numpy()
# -
# +
# read the data into a spark rdd
#books_rdd = sc.textFile('../Amazon_Data/Books.json.gz')
# +
# inspect the rdd
#books_rdd.take(3)
# +
# get the headers from the json
#headers = books_rdd.map()
# +
# parse the json in the rdd
#books_mapped = books_rdd.map()
# -
# ## Spark DataFrame
# #### This is working the best right now
# read in data to spark df
books_df = spark.read.json('../Amazon_Data/Books.json.gz')
# review the structure of the spark dataframe
books_df.printSchema()
# #### Books Data Notes
# - 'reviewerID' is the customer ID
# - 'overall' is the star rating (ranges 0 to 5)
# - 'asin' is the ID of the product
# - 'title' of the product is located in the metadata file, join by 'asin'
books_df.take(5)[0]
# +
# books_df.dtypes
# +
# register df as table
#sqlContext.registerDataFrameAsTable(books_df, 'books_df')
# -
# get count of missing values
reviews_ratings = books_df.select('reviewText', 'overall')
reviews_ratings.select([count(when(isnan(c), c)).alias(c) for c in reviews_ratings.columns]).show()
# split into labels and features
reviews = reviews_ratings.select('reviewText')
ratings = reviews_ratings.select('overall')
# inspect the reviews
reviews.take(2)
# inspect the ratings
ratings.take(2)
# display the number of ratings
ratings.count()
# display the number of reviews
reviews.count()
# store counts of ratings by number of stars and show
ratings_df = reviews_ratings.groupby('overall').count().sort(col('overall'))
ratings_df.show()
# convert to pandas df for easy plotting
ratings_hist = ratings_df.toPandas()
# visualize the number of ratings for each level
_ = sns.barplot(ratings_hist['overall'], ratings_hist['count'], color='blue')
_ = plt.title('Number of Reviews by Star Rating')
_ = plt.xlabel('Number of Stars')
_ = plt.ylabel('Number of Occurrences (Tens of Millions)')
# map the ratings to positive, neutral, and negative
ratings_hist.loc[:,'sentiment'] = ratings_hist.overall.map({0.0: 'negative', 1.0: 'negative', 2.0: 'negative',
3.0: 'neutral', 4.0: 'positive', 5.0: 'positive'})
# visualize the number of positive, neutral, and negative ratings
_ = sns.barplot(ratings_hist['sentiment'].unique(), ratings_hist.groupby('sentiment')['count'].sum())
_ = plt.title('Number of Reviews by Star Rating')
_ = plt.xlabel('Number of Stars')
_ = plt.ylabel('Number of Occurrences (Tens of Millions)')
# map the spark dataframe target for multiclass evaluation
ratings_map_neg = ratings.withColumn('negative', ratings.overall < 3.0).select('overall', 'negative')
ratings_map_neut = ratings_map_neg.withColumn('neutral', ratings_map_neg.overall == 3.0).select('overall', 'negative', 'neutral')
ratings_mapped = ratings_map_neut.withColumn('positive', ratings_map_neut.overall > 3.0).select('overall', 'negative', 'neutral', 'positive')
# inspect the results
ratings_mapped.show(100)
# convert true/false values to numerical values
sa_target = ratings_mapped.select([col(c).cast('integer') for c in ['negative', 'neutral', 'positive']])
# inspect the results
sa_target.show(100)
# #### Text Processing
# inspect nlp object
type(nlp)
# import and set additional nlp tools
import string
punctuations = string.punctuation
from spacy.lang.en import English
parser = English()
# define a tokenizer
def my_tokenizer(review):
mytokens = parser(review)
mytokens = [word.lemma_.lower().strip() if word.lemma_ != "-PRON-" else word.lower_ for word in mytokens]
mytokens = [word for word in mytokens if word not in punctuations]
return mytokens
# ML Packages
from sklearn.feature_extraction.text import CountVectorizer,TfidfVectorizer
from sklearn.base import TransformerMixin
from sklearn.pipeline import Pipeline
# +
#Custom transformer using spaCy
class predictors(TransformerMixin):
def transform(self, X, **transform_params):
return [clean_text(text) for text in X]
def fit(self, X, y, **fit_params):
return self
def get_params(self, deep=True):
return {}
# Basic function to clean the text
def clean_text(text):
return text.strip().lower()
# +
# Vectorization
vectorizer = CountVectorizer(tokenizer = my_tokenizer, ngram_range=(1,1))
# Using Tfidf
tfvectorizer = TfidfVectorizer(tokenizer = my_tokenizer)
# -
# Splitting Data Set
from pyspark.ml.tuning import ParamGridBuilder, TrainValidationSplit
# out of memory exceptions trying to examine after split
train, test = reviews_ratings.randomSplit([0.8, 0.2], seed = 42)
# +
# split the data
# x data reviews.review_test
# y data sa_target (.negative, .neutral, .positive)
# def split_data(X, y, test_size, chunksize):
# x_vals = []
# x_columns = ['reviews']
# y_vals = []
# y_columns = ['negative', 'neutral', 'positive']
# x_df = spark.createDataFrame(x_vals, x_columns)
# y_df = spark.createDataFrame(y_vals, y_columns)
# train_test_split(, sa_target)
# -
# ## Recommendation Matrix
# +
# CODE BELOW ATTEMPTED TO GENERATE A CUSTOMER LIST
# +
#cust = books_df.select('reviewerID').distinct()
# +
#cust.show(30)
# -
# isolate the items purchased by each customer
books_df.createOrReplaceTempView('books')
items_by_cust = spark.sql('SELECT reviewerID, asin FROM books ORDER BY reviewerID').toDF('reviewerID', 'itemID')
# need to transform this into sparse matrix style
items_by_cust.show(5)
# +
# only need the info below to get the titles of the books
# the items and customers are in the reviews df
# -
# read in data to spark df
meta_df = spark.read.json('../Amazon_Data/meta_Books.json.gz')
# review the structure of the spark dataframe
meta_df.printSchema()
meta_df.show(2)
# #### Metadata Notes
# - 'asin' is the item number
# - 'also_buy' lists item numbers for other purchases ('people also bought')
# - 'title' is the title of the book (not found in books_df)
| .ipynb_checkpoints/1_ACD_Data_Wrangling-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# <img src="http://cfs22.simplicdn.net/ice9/new_logo.svgz "/>
#
# # Assignment 02: Evaluate the Sentiment Dataset
#
# *The comments/sections provided are your cues to perform the assignment. You don't need to limit yourself to the number of rows/cells provided. You can add additional rows in each section to add more lines of code.*
#
# *If at any point in time you need help on solving this assignment, view our demo video to understand the different steps of the code.*
#
# **Happy coding!**
#
# * * *
# #### 1: Import the dataset
#Import the required libraries
import pandas as pd
#Import the sentiment dataset
df_sentiment = pd.read_csv('//Users//sudhanmaharjan//Desktop//DATA SCIENCE//DS_using_Python//imdb_labelled.txt',sep='\t',names=['comment','label'])
# #### 2: Analyze the dataset
#View the first 10 observations (Note: 1 indicates positive sentiment and 0 indicates negative sentiment)
df_sentiment.head()
#View statistical information about the sentiment dataset
df_sentiment.describe()
#View a quantitative description of the dataset
df_sentiment.info()
#View the data based on the sentiment type (negative or positive) (Hint: group the data by the different labels, then apply the describe() function)
df_sentiment.groupby('label').describe()
# #### 3: Find the length of the messages and add it as a new column
#Calculate the length of the messages and add the values as a new column in the dataset
df_sentiment['length'] = df_sentiment['comment'].apply(len)
#Verify if the dataset is updated with the additional column
df_sentiment.head()
#View the first message which contains more than 50 characters
df_sentiment[df_sentiment['length']>50]['comment'].iloc[0]
# #### 4: Apply a transformer and fit the data in the Bag of Words
#Process the text with a vectorizer
from sklearn.feature_extraction.text import CountVectorizer
vectorizer = CountVectorizer()
# +
#Define a function to get rid of stopwords present in the messages
def message_text_process(mess):
# Check characters to see if there are punctuations
no_punctuation = [char for char in mess if char not in string.punctuation]
# Form the sentence
no_punctuation = ''.join(no_punctuation)
# Eliminate any stopwords
return[word for word in no_punctuation.split() if word.lower() not in stopwords.words('english')]
# -
#Apply the Bag of Words and fit the data (comments) into it
import string
from nltk.corpus import stopwords
bag_of_words = CountVectorizer(analyzer=message_text_process).fit(df_sentiment['comment'])
#Apply the transform method to the Bag of Words
comment_bagofwords = bag_of_words.transform(df_sentiment['comment'])
#Apply tf-idf transformer and fit the Bag of Words into it (transformed version)
from sklearn.feature_extraction.text import TfidfTransformer
tfidf_transformer = TfidfTransformer().fit(comment_bagofwords)
# #### 5: Print the shape of the transformer
#Print the shape
comment_tfidf = tfidf_transformer.transform(comment_bagofwords)
print comment_tfidf.shape
# #### 6: Create and check the Naïve Bayes model for predicted and expected values
#Create a Naïve Bayes model to detect the sentiment and fit the tf-idf data into it
from sklearn.naive_bayes import MultinomialNB
sentiment_detection_model = MultinomialNB().fit(comment_tfidf,df_sentiment['label'])
# +
#Evaluate the model's accuracy by comparing the predicted and expected values for, say, comment# 1 and comment#5
comment = df_sentiment['comment'][4]
bag_of_words_for_comment = bag_of_words.transform([comment])
tfidf = tfidf_transformer.transform(bag_of_words_for_comment)
print 'predicted sentiment label', sentiment_detection_model.predict(tfidf)[0]
print 'expected sentiment label', df_sentiment.label[4]
# -
| Assignment 02.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Image Classification using LeNet CNN
# ## CIFAR-10 Dataset - Animals and Objects (10 classes)
#
# 
# +
# import tensorflow module. Check API version.
import tensorflow as tf
import numpy as np
print (tf.__version__)
# required for TF to run within docker using GPU (ignore otherwise)
gpu = tf.config.experimental.list_physical_devices('GPU')
tf.config.experimental.set_memory_growth(gpu[0], True)
# -
# ## Load the data
# grab the CIFAR-10 dataset (may take time the first time)
print("[INFO] downloading CIFAR-10...")
((trainData, trainLabels), (testData, testLabels)) = tf.keras.datasets.cifar10.load_data()
# ## Prepare the data
# parameters for CIFAR-10 data set
num_classes = 10
image_width = 32
image_height = 32
image_channels = 3 # CIFAR data is RGB color
# define human readable class names
class_names = ['airplane', 'automobile', 'bird', 'cat',
'deer', 'dog', 'frog', 'horse', 'ship', 'truck']
# shape the input data using "channels last" ordering
# num_samples x rows x columns x depth
trainData = trainData.reshape(
(trainData.shape[0], image_height, image_width, image_channels))
testData = testData.reshape(
(testData.shape[0], image_height, image_width, image_channels))
# convert to floating point and scale data to the range of [0.0, 1.0]
trainData = trainData.astype("float32") / 255.0
testData = testData.astype("float32") / 255.0
# display data dimentions
print ("trainData:", trainData.shape)
print ("trainLabels:", trainLabels.shape)
print ("testData:", testData.shape)
print ("testLabels:", testLabels.shape)
# ## Define Model
#
# 
# +
# import the necessary packages
from tensorflow.keras import backend
from tensorflow.keras import models
from tensorflow.keras import layers
# define the model as a class
class LeNet:
# INPUT => CONV => TANH => AVG-POOL => CONV => TANH => AVG-POOL => FC => TANH => FC => TANH => FC => SMAX
@staticmethod
def init(numChannels, imgRows, imgCols, numClasses, weightsPath=None):
# if we are using "channels first", update the input shape
if backend.image_data_format() == "channels_first":
inputShape = (numChannels, imgRows, imgCols)
else: # "channels last"
inputShape = (imgRows, imgCols, numChannels)
# initialize the model
model = models.Sequential()
# define the first set of CONV => ACTIVATION => POOL layers
model.add(layers.Conv2D(filters=6, kernel_size=(5, 5), strides=(1, 1),
padding="valid", activation=tf.nn.tanh, input_shape=inputShape))
model.add(layers.AveragePooling2D(pool_size=(2, 2), strides=(2, 2)))
# define the second set of CONV => ACTIVATION => POOL layers
model.add(layers.Conv2D(filters=16, kernel_size=(5, 5), strides=(1, 1),
padding="valid", activation=tf.nn.tanh))
model.add(layers.AveragePooling2D(pool_size=(2, 2), strides=(2, 2)))
# flatten the convolution volume to fully connected layers
model.add(layers.Flatten())
# define the first FC => ACTIVATION layers
model.add(layers.Dense(units=120, activation=tf.nn.tanh))
# define the second FC => ACTIVATION layers
model.add(layers.Dense(units=84, activation=tf.nn.tanh))
# lastly, define the soft-max classifier
model.add(layers.Dense(units=numClasses, activation=tf.nn.softmax))
# if a weights path is supplied (inicating that the model was
# pre-trained), then load the weights
if weightsPath is not None:
model.load_weights(weightsPath)
# return the constructed network architecture
return model
# -
# ## Compile Model
# +
# initialize the model
print("[INFO] compiling model...")
model = LeNet.init(numChannels=image_channels,
imgRows=image_height, imgCols=image_width,
numClasses=num_classes,
weightsPath=None)
# compile the model
model.compile(optimizer=tf.keras.optimizers.Adam(), # Adam optimizer
loss="sparse_categorical_crossentropy",
metrics=["accuracy"])
# print model summary
model.summary()
# -
# ## Train Model
# +
# define callback function for training termination criteria
#accuracy_cutoff = 0.99
class myCallback(tf.keras.callbacks.Callback):
def on_epoch_end(self, epoch, logs=None):
if(logs.get('accuracy') > 0.90):
print("\nReached 90% accuracy so cancelling training!")
self.model.stop_training = True
# initialize training config
batch_size = 256
epochs = 100
# run training
print("[INFO] training...")
history = model.fit(x=trainData, y=trainLabels, batch_size=batch_size,
validation_data=(testData, testLabels), epochs=epochs, verbose=1, callbacks=[myCallback()])
# -
# ## Evaluate Training Performance
#
# ### Expected Output
#
#  
# +
# %matplotlib inline
import matplotlib.pyplot as plt
# retrieve a list of list results on training and test data sets for each training epoch
acc = history.history['accuracy']
val_acc = history.history['val_accuracy']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(len(acc)) # get number of epochs
# plot training and validation accuracy per epoch
plt.plot(epochs, acc, label='train accuracy')
plt.plot(epochs, val_acc, label='val accuracy')
plt.xlabel('epochs')
plt.ylabel('accuracy')
plt.legend(loc="lower right")
plt.title('Training and validation accuracy')
plt.figure()
# plot training and validation loss per epoch
plt.plot(epochs, loss, label='train loss')
plt.plot(epochs, val_loss, label='val loss')
plt.xlabel('epochs')
plt.ylabel('loss')
plt.legend(loc="upper right")
plt.title('Training and validation loss')
# -
# show the accuracy on the testing set
print("[INFO] evaluating...")
(loss, accuracy) = model.evaluate(testData, testLabels,
batch_size=batch_size, verbose=1)
print("[INFO] accuracy: {:.2f}%".format(accuracy * 100))
model.save_weights("weights/LeNetCIFAR10.temp.hdf5", overwrite=True)
# ## Evaluate Pre-trained Model
# +
# init model and load the model weights
print("[INFO] compiling model...")
model = LeNet.init(numChannels=image_channels,
imgRows=image_height, imgCols=image_width,
numClasses=num_classes,
weightsPath="weights/LeNetCIFAR10.hdf5")
# compile the model
model.compile(optimizer=tf.keras.optimizers.SGD(lr=0.01), # Stochastic Gradient Descent
loss="sparse_categorical_crossentropy",
metrics=["accuracy"])
# -
# show the accuracy on the testing set
print("[INFO] evaluating...")
batch_size = 128
(loss, accuracy) = model.evaluate(testData, testLabels,
batch_size=batch_size, verbose=1)
print("[INFO] accuracy: {:.2f}%".format(accuracy * 100))
# ## Model Predictions
# +
# %matplotlib inline
import numpy as np
import cv2
import matplotlib.pyplot as plt
# set up matplotlib fig, and size it to fit 3x4 pics
nrows = 3
ncols = 4
fig = plt.gcf()
fig.set_size_inches(ncols*4, nrows*4)
# randomly select a few testing digits
num_predictions = 12
test_indices = np.random.choice(np.arange(0, len(testLabels)), size=(num_predictions,))
test_images = np.stack(([testData[i] for i in test_indices]))
test_labels = np.stack(([testLabels[i] for i in test_indices]))
# compute predictions
predictions = model.predict(test_images)
for i in range(num_predictions):
# select the most probable class
prediction = np.argmax(predictions[i])
# rescale the test image
image = (test_images[i] * 255).astype("uint8")
# resize the image from a 32 X 32 image to a 96 x 96 image so we can better see it
image = cv2.resize(image, (96, 96), interpolation=cv2.INTER_CUBIC)
# select prediction text color
if prediction == test_labels[i]:
rgb_color = (0, 255, 0) # green for correct predictions
else:
rgb_color = (255, 0, 0) # red for wrong predictions
# show the image and prediction
cv2.putText(image, str(class_names[prediction]), (0, 10),
cv2.FONT_HERSHEY_SIMPLEX, 0.5, rgb_color, 1)
# set up subplot; subplot indices start at 1
sp = plt.subplot(nrows, ncols, i + 1, title="label: %s" % class_names[test_labels[i][0]])
sp.axis('Off') # don't show axes (or gridlines)
plt.imshow(image)
# show figure matrix
plt.show()
# -
| LeNet-CIFAR10.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import numpy as np
import powerlaw
edges= np.array([[1,2],[0,2],[0,3],[2,3],[3,4],[4,1]])
class karpatiGraphSolution:
def __init__(self,edges):
assert type(edges)==np.ndarray, "input is not an edge list"
self.edgeList=edges
self.numNodes=np.amax(edges)+1
def give_me_matrix(self):
res=[[0] * self.numNodes for i in range(self.numNodes)]
for edge in self.edgeList:
res[edge[0]][edge[1]]=1
self.adjMat=res
return res
def isConnected(self):
rowSums=np.asarray(self.adjMat).sum(0)
colSums=np.asarray(self.adjMat).sum(1)
print(rowSums)
print(colSums)
total=rowSums+colSums
res=0 not in total
return res
def isStronglyConnected(self):
rowSums=np.asarray(self.adjMat).sum(0)
colSums=np.asarray(self.adjMat).sum(1)
print(rowSums)
print(colSums)
res=0 not in rowSums & 0 not in colSums
return res
def MST(self):
assert self.isConnected, "Sorry, your graph is not connected"
treeMST=set()
nodeInMST=set()
nodeInMST.add(self.edgeList[0][0])
print(nodeInMST)
for edge in self.edgeList:
if (edge[1] in nodeInMST and edge[0] not in nodeInMST):
print("LOL")
treeMST.add((edge[0],edge[1]))
nodeInMST.add(edge[0])
print(nodeInMST)
elif (edge[0] in nodeInMST and edge[1] not in nodeInMST):
print("LOL2")
nodeInMST.add(edge[1])
treeMST.add((edge[1],edge[0]))
print(nodeInMST)
#nodeInMST.add(edge[1])
if len(nodeInMST)==self.numNodes:
print("BREAKING")
break
return(treeMST)
def fitPowerLaw(self):
#get degree distribution
rowSums=np.asarray(self.adjMat).sum(0)
colSums=np.asarray(self.adjMat).sum(1)
total=rowSums+colSums
results=powerlaw.Fit(total)
print("LOL")
return(results.power_law.alpha,results.power_law.xmin)
sol=karpatiGraphSolution(edges)
cucc=sol.give_me_matrix()
cucc3=sol.MST()
print(cucc3)
cucc4=sol.fitPowerLaw()
print(cucc4)
# +
var = 100
if var == 200:
print "1 - Got a true expression value"
print var
elif var == 150:
print "2 - Got a true expression value"
print var
elif var == 100:
print "3 - Got a true expression value"
print var
else:
print "4 - Got a false expression value"
print var
print "Good bye!"
# -
| Laci_1_graph.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="aTOLgsbN69-P"
# # Some Math Background
# + [markdown] id="yqUB9FTRAxd-"
# Manipulating numbers, scalars, vectors, and matrices lies at the heart of most machine learning approaches.
# + [markdown] id="d4tBvI88BheF"
# In this pairs of introductory notebooks, you need some basic algebra and geometry background. You will add to what you know by:
#
# * Learning what are and how to represent scalars, vectors, and matrices
#
# * Developing a geometric intuition of what’s going on beneath the hood of machine learning algorithms.
#
# * Grasping how numpy works and how we can use it in the design of machine learning algorithms
# + [markdown] id="Z68nQ0ekCYhF"
# Through the course of these 2 notebooks, you will come to know what is meant by tensors -- from whose name Google's Tensorflow is derived.
# -
# <a href="https://colab.research.google.com/github/philmui/study-algorithmic-bias/blob/main/notebooks/01a_numpy.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="2khww76J5w9n"
# ## 1: Representing Algebra
#
#
# + [markdown] id="niG_MgK-iV6K"
# ### Graphing algebra
#
# Let's start with a familiar algebra problem.
#
# > After robbing a bank, a robber took off in a car that travels at 2.5 kilometer per minute. 5 minutes after the robber took off, the police arrived at the bank and instantly chased after the robber at 3 kilometer per minute.
# > Would the sheriff be able to catch up with the robber? If so, how many minutes after the robber took off would the sheriff caught up, and at what kilometer distance away from the bank would both of them be at that time?
#
# Before we continue, please feel free to solve this problem on paper -- ideally with 2 line graphs showing the intersections.
#
# * Let `x` be the time (`t`) by minutes passed since the robber took off.
# * Let `y` be the distance (`d`) that either has traveled away from the bank.
# * `d_r` = distance traveled by the robber
# * `d_s` = distance traveled by the sheriff
# + id="LApX90aliab_"
import numpy as np
import matplotlib.pyplot as plt
# + id="E4odh9Shic1S"
t = np.linspace(0, 40, 1000) # start, finish, n points
# + [markdown] id="N-tYny12nIyO"
# Distance travelled by robber: $d = 2.5t$
# -
t.shape
t[:10]
# + id="e_zDOxgHiezz"
d_r = 2.5 * t
# + [markdown] id="djVjXZy-nPaR"
# Distance travelled by sheriff: $d = 3(t-5)$
# + id="JtaNeYSCifrI"
d_s = 3 * (t-5)
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="SaaIjJSEigic" outputId="46aa6132-c721-4453-b937-1a70b1ee84b6"
fig, ax = plt.subplots()
plt.title('A Bank Robber Caught')
plt.xlabel('time (in minutes)')
plt.ylabel('distance (in km)')
ax.set_xlim([0, 40])
ax.set_ylim([0, 100])
ax.plot(t, d_r, c='green')
ax.plot(t, d_s, c='brown')
plt.axvline(x=30, color='purple', linestyle='--')
_ = plt.axhline(y=75, color='purple', linestyle='--')
# + [markdown] id="kpwZw64EYfs6"
# So now it is clear by the graph intersection that 30 minutes after the robber took off, he will be caught by the sheriff a distance nearly 80 km away.
# + [markdown] id="NgGMhK4B51oe"
# ### Python Types
# + colab={"base_uri": "https://localhost:8080/"} id="ZXnTHDn_EW6b" outputId="b4619155-d8a7-42cc-8af1-eec886a46ec5"
x = 25
x
# + colab={"base_uri": "https://localhost:8080/"} id="VF8Jam76R4KJ" outputId="e31092f3-86b4-47ea-8c51-cf9403ce4d24"
type(x)
# + id="ZBzYlL0mRd-P"
y = 3
# + colab={"base_uri": "https://localhost:8080/"} id="1i-hW0bcReyy" outputId="8981c789-37d5-4176-a34e-29391039b21b"
py_sum = x + y
py_sum
# + colab={"base_uri": "https://localhost:8080/"} id="CpyUxB6XRk6y" outputId="25db8c52-df25-442d-cd8f-9d5f51f7d084"
type(py_sum)
# + colab={"base_uri": "https://localhost:8080/"} id="V2UiLj-JR8Ij" outputId="4cf38ebb-a628-4784-be37-25ceb2a54f23"
x_float = 25.0
float_sum = x_float + yfloat_sum
# + colab={"base_uri": "https://localhost:8080/"} id="ikOwjp6ASCaf" outputId="fc22a306-1065-44de-fcf4-001e7f09a632"
type(float_sum)
# -
# ### Scalars, Vectors, Matrices, Tensors
#
# Scalars are numbers or single variables
#
# Vectors are multivariate representation of an ordered set of numbers or variables
#
# Matrics are 2-dimensional vectors
#
# A tensor is an algebraic object that describes a (multilinear) relationship between sets of algebraic objects related to a vector space.
#
# PyTorch and TensorFlow are the two most popular ML libraries in Python, itself the most popular programming language in ML
# Both support a generalization of "arrays" called "tensors" (arrays in multiple dimensions very well). Tensors are very common data structures used for training deep learning neural networks.
# * scalars : tensor of rank 0
# * 1D arrays or vectors : tensor of rank 1
# * 2D arrays or matrics : tensor of rank 2
# * ...
# + [markdown] id="SgUvioyUz8T2"
# ### PyTorch
#
# * PyTorch tensors are designed to be pythonic, i.e., to feel and behave like NumPy arrays
# * The advantage of PyTorch tensors relative to NumPy arrays is that they easily be used for operations on GPU (see [here](https://pytorch.org/tutorials/beginner/examples_tensor/two_layer_net_tensor.html) for example)
# * Documentation on PyTorch tensors, including available data types, is [here](https://pytorch.org/docs/stable/tensors.html)
# + id="A9Hhazt2zKeD"
import torch
# + colab={"base_uri": "https://localhost:8080/"} id="a211IRW_0-iY" outputId="3fa02f13-5bf6-43ea-9e00-380aa416d51c"
x_pt = torch.tensor(25) # type specification optional, e.g.: dtype=torch.float16
x_pt
# + colab={"base_uri": "https://localhost:8080/"} id="LvxzMa_HhUNB" outputId="861450df-31ef-4c03-a326-5ddc5ea42631"
x_pt.shape
# -
vec = torch.tensor([1, 2, 3])
vec
vec.shape
mat = torch.tensor([[1,2,3], [1,2,3], [1,2,3], [1,2,3]])
mat.shape
# + [markdown] id="eUyuZXlWS8T9"
# ### TensorFlow
#
# Tensors created with a wrapper, all of which [you can read about here](https://www.tensorflow.org/guide/tensor):
#
# * `tf.Variable`
# * `tf.constant`
# * `tf.placeholder`
# * `tf.SparseTensor`
#
# Most widely-used is `tf.Variable`, which we'll use here.
#
# As with TF tensors, in PyTorch we can similarly perform operations, and we can easily convert to and from NumPy arrays
#
# Also, a full list of tensor data types is available [here](https://www.tensorflow.org/api_docs/python/tf/dtypes/DType).
# + id="CHBYse_MEqZM"
import tensorflow as tf
# + colab={"base_uri": "https://localhost:8080/"} id="sDv92Nh-NSOU" outputId="6f0da27c-6628-499b-e78a-908abcc56b68"
x_tf = tf.Variable(25, dtype=tf.int16) # dtype is optional
x_tf
# + colab={"base_uri": "https://localhost:8080/"} id="EmPMBIV9RQjS" outputId="372d2778-ea2f-4912-c5b6-93629561c476"
x_tf.shape
# + id="mEILtO9pPctO"
y_tf = tf.Variable(3, dtype=tf.int16)
# + colab={"base_uri": "https://localhost:8080/"} id="dvvWuaw6Ph_D" outputId="a343988f-241c-444b-828c-b88590c20b45"
x_tf + y_tf
# + colab={"base_uri": "https://localhost:8080/"} id="JZVhRnX9RUGW" outputId="8b0a8690-9636-4aa6-efce-070e00a748e8"
tf_sum = tf.add(x_tf, y_tf)
tf_sum
# + colab={"base_uri": "https://localhost:8080/"} id="sVbMxT1Ey6Y3" outputId="08bb9964-74f2-4318-f121-06d1f291a143"
tf_sum.numpy() # note that NumPy operations automatically convert tensors to NumPy arrays, and vice versa
# + colab={"base_uri": "https://localhost:8080/"} id="LXpv69t0y-f6" outputId="f46934d6-15d4-4352-85b2-31ee20a0f941"
type(tf_sum.numpy())
# + colab={"base_uri": "https://localhost:8080/"} id="VszuTUAg1uXk" outputId="273832c6-b4e9-4895-9558-295f05d1acf2"
tf_float = tf.Variable(25., dtype=tf.float16)
tf_float
# + [markdown] id="B5VRGo1H6010"
# Let's add a few higher (rank) dimensional tensors.
# + [markdown] id="4CURG9Er6aZI"
# ### Vectors (Rank 1 Tensors) in NumPy
# + colab={"base_uri": "https://localhost:8080/"} id="T9ME4kBr4wg0" outputId="6b354e27-c09d-4ae1-fad6-089ab96b0575"
x = np.array([25, 2, 5]) # type argument is optional, e.g.: dtype=np.float16
x
# + colab={"base_uri": "https://localhost:8080/"} id="ZuotxmlZL2wp" outputId="84f6dab8-f247-426e-d6da-f7ba856143e4"
len(x)
# + colab={"base_uri": "https://localhost:8080/"} id="OlPYy6GOaIVy" outputId="70509a9b-cfaa-4ed6-e5b3-d983cec675d3"
x.shape
# + colab={"base_uri": "https://localhost:8080/"} id="sWbYGwObcgtK" outputId="82d989e4-d140-452f-e998-75a997f712cd"
type(x)
# + colab={"base_uri": "https://localhost:8080/"} id="ME_xuvD_oTPg" outputId="b5d3b9f6-3838-49c4-cbd8-99b3300a615f"
x[0] # zero-indexed
# + colab={"base_uri": "https://localhost:8080/"} id="hXmBHZQ-nxFw" outputId="e143a337-35ec-4978-8ca8-a9eeae586a8d"
type(x[0])
# + [markdown] id="NiEofCzYZBrQ"
# ### Vector Transposition
# + colab={"base_uri": "https://localhost:8080/"} id="hxGFNDx6V95l" outputId="7ec7e531-d95f-40d6-8a31-01814ef6c84f"
# Transposing a regular 1-D array has no effect...
x_t = x.T
x_t
# + colab={"base_uri": "https://localhost:8080/"} id="_f8E9ExDWw4p" outputId="51dd64e7-e85e-418a-b046-3065e18d0023"
x_t.shape
# + colab={"base_uri": "https://localhost:8080/"} id="AEd8jB7YcgtT" outputId="ab5bb50b-c057-449d-d498-5c4fdf6464cc"
# ...but it does we use nested "matrix-style" brackets:
y = np.array([[25, 2, 5]])
y
# + colab={"base_uri": "https://localhost:8080/"} id="UHQd92oRcgtV" outputId="3a401b4b-9b3d-49df-b4c6-3def25d16077"
y.shape
# + colab={"base_uri": "https://localhost:8080/"} id="SPi1JqGEXXUc" outputId="2cb64c0d-2faf-420a-a7e0-0900d12a21a4"
# ...but can transpose a matrix with a dimension of length 1, which is mathematically equivalent:
y_t = y.T
y_t
# + colab={"base_uri": "https://localhost:8080/"} id="6rzUv762Yjis" outputId="2950eeda-49e7-4de3-8719-9561184c3fd4"
y_t.shape # this is a column vector as it has 3 rows and 1 column
# + colab={"base_uri": "https://localhost:8080/"} id="xVnQMLOrYtra" outputId="fdd09561-116a-492f-db1a-bcb7395862f1"
# Column vector can be transposed back to original row vector:
y_t.T
# + colab={"base_uri": "https://localhost:8080/"} id="QIAA2NLRZIXC" outputId="ab428a9e-13f6-45af-9e30-70d1ebc220eb"
y_t.T.shape
# + [markdown] id="Voj26mSpZLuh"
# ### Zero Vectors
#
# Have no effect if added to another vector
# + colab={"base_uri": "https://localhost:8080/"} id="-46AbOdkZVn_" outputId="bc21d3dc-8816-4803-d5b3-57232930f59f"
z = np.zeros(3)
z
# -
z.shape
z1 = np.array([np.zeros(3)])
z1.shape
z1
z1.T
z1.shape
# ### Vector and Matrix
# What is the shape of the array (or vector)? `np.array([1,2,3])`
v = np.array([1,2,3])
v
v.shape
# This is a vector of length 3. But as a 2D matrix, the 2nd dimension is not explicitly defined and has no value according to numpy.
#
# We can explicitly create a matrix with well defined rows and columns:
m = np.array([[1,2,3]])
m
# The interesting question to ask is: is this a "row vector" or a "column vector"? For numpy: this additional bracket creates a "row vector"
m.shape
# To convert this row vector to a column vector of shape (3,1), we need to do a matrix _transposition_.
m.T
m.T.shape
# ### Vector and Matrix Concaternation
# If we want to concaternate to the 1st dimension of the vector v, here is a conveience function to do that: `np.r_`
np.r_[np.array([1,2,3]), np.array([4,5,6])]
np.r_[np.array([1,2,3]), 0, 0, np.array([4,5,6])]
# By default, np.r_ adds vectors along their "first axis" -- which in this case is the only axis for all vectors. Scalars are converted to a 1x1 vector. Arrays are convered to 3x1 vectors. So np.r_ adds the vectors along the ***first dimension*** : which is the "row" dimension or explicitly as 'r':
m = np.r_['r', np.array([1,2,3]), 0, 0, np.array([4,5,6])]
m
m.shape
# We can also specify the concaternation to be done along the ***last dimension*** : which is the "column" dimension or 'c'. In this case, all of the np.arrays are convered to nx1 vectors:
mr = np.r_['c', np.array([1,2,3]), 0, 0, np.array([4,5,6])]
mr
mr.shape
# If we want to conceternate the array / vectors along the ***last dimension***. Arrays have only 1 dimension. So we mentally add one: so shape of both array becomes (3,1).
#
# So resultant shape would be (3,1+1) which is (3,2). which is the shape of result.
#
# There is a convenience method np.c_ that does the trick.
mc = np.c_[np.array([1,2,3]), np.array([4,5,6])]
mc
mc.shape
# Another example:
#
# `np.c_[np.array([[1,2,3]]), 0, 0, np.array([[4,5,6]])]`
#
# shapes:
#
# `np.array([[1,2,3]])` = 1,3
#
# `np.array([[4,5,6]])` = 1,3
#
# For scalars such as 0 so we can think of it as [[0]] = 1,1
#
# So result 1,3+1+1+3 = 1,8
#
# which should be the shape of result : array([[1, 2, 3, 0, 0, 4, 5, 6]])
#
np.c_[np.array([[1,2,3]]), 0, 0, np.array([[4,5,6]])]
np.c_[np.array([[1,2,3]]), 0, 0, np.array([[4,5,6]])].shape
# So, how do we stack "rows" of vectors along the first dimension of the rows? We need to specific the array conversion to vectors as extending along the 2nd column dimension: which is 1x2 but arrays have no first dimension, and this is the same as '0x2' or '0,2':
mr2 = np.r_['0,2', np.array([1,2,3]), np.array([4,5,6])]
mr2
mr2.shape
# + [markdown] id="c6xyYiwSnSGC"
# ### Vectors in PyTorch and TensorFlow
# + colab={"base_uri": "https://localhost:8080/"} id="s2TGDeqXnitZ" outputId="b7266589-224d-46a3-efdc-f0772dc1acf4"
x_pt = torch.tensor([25, 2, 5])
x_pt
# -
x_pt.T
x1_pt = torch.tensor([[25, 2, 5]])
x1_pt
x1_pt.T
# + colab={"base_uri": "https://localhost:8080/"} id="-0jbHgc5iijG" outputId="62ae4a45-8b8b-420f-e7d0-242d21549b49"
x_tf = tf.Variable([25, 2, 5])
x_tf
# -
x_tf.shape
x1_tf = tf.Variable([[25, 2, 5]])
x1_tf
x1_tf.shape
# + [markdown] id="8fU5qVTI6SLD"
# ### $L^2$ Norm
# + colab={"base_uri": "https://localhost:8080/"} id="lLc2FbGG6SLD" outputId="61005d0b-5ea3-4162-9109-4904b37c6684"
x
# + colab={"base_uri": "https://localhost:8080/"} id="AN43hsl86SLG" outputId="8eb00d73-1dae-4b96-f532-025ab499f375"
(25**2 + 2**2 + 5**2)**(1/2)
# + colab={"base_uri": "https://localhost:8080/"} id="D9CyWo-l6SLI" outputId="c15f5bbe-4e0f-40ad-d5fc-e44e747a8906"
np.linalg.norm(x)
# + [markdown] id="TNEMRi926SLK"
# So, if units in this 3-dimensional vector space are meters, then the vector $x$ has a length of 25.6m
# + [markdown] id="PwiRlMuC6SLK"
# ### $L^1$ Norm
# + colab={"base_uri": "https://localhost:8080/"} id="lcYKyc5H6SLL" outputId="6df30f51-49ef-48b6-839c-cf1ca534027c"
x
# + colab={"base_uri": "https://localhost:8080/"} id="8jNb6nYl6SLM" outputId="de08ed57-e3d7-40f6-acfa-990d2ea7f873"
np.abs(25) + np.abs(2) + np.abs(5)
# + [markdown] id="lQP73B916SLP"
# ### Squared $L^2$ Norm
# + colab={"base_uri": "https://localhost:8080/"} id="Qv1ouJ8r6SLP" outputId="c16a9a74-b980-4a33-b675-6eb829245331"
x
# + colab={"base_uri": "https://localhost:8080/"} id="eG3WSB5R6SLT" outputId="dcac7e6f-1950-4a46-f3ca-3f06a08765d7"
(25**2 + 2**2 + 5**2)
# + colab={"base_uri": "https://localhost:8080/"} id="bXwzSudS6SLV" outputId="1c635cf8-a62b-497c-a7de-0e73eb39eb35"
# we'll cover tensor multiplication more soon but to prove point quickly:
np.dot(x, x)
# + [markdown] id="BHWxVPFC6SLX"
# ### Max Norm
# + colab={"base_uri": "https://localhost:8080/"} id="vO-zfvDG6SLX" outputId="11ca2293-00ae-4339-a87c-a92091c7496f"
x
# + colab={"base_uri": "https://localhost:8080/"} id="vXXLgbyW6SLZ" outputId="74458574-db7c-4ef9-e17e-7676991040b9"
np.max([np.abs(25), np.abs(2), np.abs(5)])
# + [markdown] id="JzKlIpYZcgt9"
# ### Orthogonal Vectors
# -
# Vectors x and y are "orthogonal" if and only if (iff) x^T y = 0.
#
# Orthogonal vectors x and y are at 90 degree to each other.
#
# n-dimensional space has max n mutually orthogonal vectors.
#
# *Orthonormal* vectors are orthogonal and all have unit norm.
# + colab={"base_uri": "https://localhost:8080/"} id="4jHg9La-cgt9" outputId="6449ab56-0ae7-4c14-b183-6e398d57f2e4"
i = np.array([1, 0])
i
# + colab={"base_uri": "https://localhost:8080/"} id="3FyLhPK3cguA" outputId="5b2eefc4-2dfd-4be9-de34-60ab1badfb55"
j = np.array([0, 1])
j
# + colab={"base_uri": "https://localhost:8080/"} id="7eQtKhaDcguC" outputId="f5b7978c-ad97-4325-d9ea-befe8f3686e7"
np.dot(i, j)
# + [markdown] id="mK3AZH53o8Br"
# ### Matrices in NumPy
#
# Matrices are Rank 2 Tensors, and are denoted here as upper case variables such as X or Y.
# + colab={"base_uri": "https://localhost:8080/"} id="stk57cmaESW1" outputId="9990ddb4-5d91-4147-e0af-66eae9d3b40d"
# Use array() with nested brackets:
X = np.array([[25, 2], [5, 26], [3, 7]])
X
# + colab={"base_uri": "https://localhost:8080/"} id="IhDL4L8S6SLc" outputId="57dbb7c8-012a-4f05-ba14-6e7abc4c267d"
X.shape
# + colab={"base_uri": "https://localhost:8080/"} id="q3oyaAK36SLe" outputId="1d67cfe8-aa58-4393-9f5c-0064c00f5fe6"
X.size
# + colab={"base_uri": "https://localhost:8080/"} id="YN9CHzja6SLg" outputId="53059362-54ae-4077-dc00-58a37ae3b608"
# Select left column of matrix X (zero-indexed)
X[:,0]
# + colab={"base_uri": "https://localhost:8080/"} id="ih7nh4qC6SLi" outputId="120c343e-7b2f-4ead-e4cd-b204c53de655"
# Select middle row of matrix X:
X[1,:]
# + colab={"base_uri": "https://localhost:8080/"} id="pg7numxP6SLl" outputId="d026484d-805b-4a70-8408-34091017108c"
# Another slicing-by-index example:
X[0:2, 0:2]
# + [markdown] id="HGEfZiBb6SLt"
# ### Matrices in PyTorch
# + colab={"base_uri": "https://localhost:8080/"} id="-bibT9ye6SLt" outputId="de70f48d-71bb-4fdd-ea5a-582cda50f04e"
X_pt = torch.tensor([[25, 2], [5, 26], [3, 7]])
X_pt
# + colab={"base_uri": "https://localhost:8080/"} id="TBPu1L7P6SLv" outputId="622c74c0-4d34-4d2c-ab50-ee3efc900e69"
X_pt.shape # more pythonic
# + colab={"base_uri": "https://localhost:8080/"} id="4mTj56M16SLw" outputId="56ae6826-0c27-4b0f-fecc-2571a40f1187"
X_pt[1,:]
# -
# Another slicing-by-index example:
X_pt[0:2, 0:2]
# + [markdown] id="E026fQlD6SLn"
# ### Matrices in TensorFlow
# + colab={"base_uri": "https://localhost:8080/"} id="1gtGH6oA6SLn" outputId="52d4345c-de79-4164-a459-e4a328393d8b"
X_tf = tf.Variable([[25, 2], [5, 26], [3, 7]])
X_tf
# + colab={"base_uri": "https://localhost:8080/"} id="4CV_KiTP6SLp" outputId="9e9bcc51-6299-4598-ed5c-a458f8591d16"
tf.rank(X_tf)
# + colab={"base_uri": "https://localhost:8080/"} id="vUsce8tC6SLq" outputId="d224b13e-28c2-4d15-afc3-919da18c6301"
tf.shape(X_tf)
# + colab={"base_uri": "https://localhost:8080/"} id="QNpfvNPj6SLr" outputId="735453ed-d2dc-479b-b734-ef8dc245ff4a"
X_tf[1,:]
# -
X_tf[0:2, 0:2]
# + [markdown] id="iSHGMCxd6SL4"
# ### Vector & Matrix Transposition
# + colab={"base_uri": "https://localhost:8080/"} id="1YN1narR6SL4" outputId="3d300f4d-8554-4cce-a1ac-e4bec7c79af4"
X
# + colab={"base_uri": "https://localhost:8080/"} id="5hf3M_NL6SL5" outputId="fc2d5536-fef3-4124-a273-f43e19c574c0"
X.T
# + colab={"base_uri": "https://localhost:8080/"} id="vyBFN_4g6SL9" outputId="e5b524e0-ee0c-4077-9c7d-e2c31aa74cb6"
X_pt.T
# + colab={"base_uri": "https://localhost:8080/"} id="K2DuDJc_6SL6" outputId="7f3c5565-31c1-4539-e42f-ddca5c1bb282"
tf.transpose(X_tf) # less Pythonic
# + [markdown] id="Hp9P1jx76SL_"
# ### Basic Arithmetical Properties
# + [markdown] id="WxaImEUc6SMA"
# Adding or multiplying with scalar applies operation to all elements and tensor shape is retained:
# + colab={"base_uri": "https://localhost:8080/"} id="yhXGETii6SMA" outputId="6898a091-64bb-4905-9cb7-5c23cb29fd24"
X*2
# + colab={"base_uri": "https://localhost:8080/"} id="KnPULtDO6SMC" outputId="f22f6077-c9ba-4424-a426-8bea20f59ffb"
X+2
# + colab={"base_uri": "https://localhost:8080/"} id="MkfC0Gsb6SMD" outputId="41a225e6-62ce-427a-9a4b-e4506782ab3f"
X*2+2
# + colab={"base_uri": "https://localhost:8080/"} id="04bIDpGj6SMH" outputId="79062d4b-be68-4399-f601-21cf6a836cc7"
X_pt*2+2 # Python operators are overloaded; could alternatively use torch.mul() or torch.add()
# + colab={"base_uri": "https://localhost:8080/"} id="2oRBSmRL6SMI" outputId="c040caeb-db4f-44cf-8c92-e02c1b7adba1"
torch.add(torch.mul(X_pt, 2), 2)
# + colab={"base_uri": "https://localhost:8080/"} id="OMSb9Otd6SMF" outputId="a0d0a0dd-b18d-4816-d7bc-68d1582014ae"
X_tf*2+2 # Operators likewise overloaded; could equally use tf.multiply() tf.add()
# + colab={"base_uri": "https://localhost:8080/"} id="5ya2xZ4u6SMG" outputId="0a18847d-ce61-40b3-8032-10ee7c808259"
tf.add(tf.multiply(X_tf, 2), 2)
# + [markdown] id="wt8Ls4076SMK"
# If two matrices have the same size, operations are often by default applied element-wise. This is **not matrix multiplication**, but is rather called the **Hadamard product** or simply the **element-wise product**.
#
# The mathematical notation is $A \odot X$
# + colab={"base_uri": "https://localhost:8080/"} id="KUMyU1t46SMK" outputId="c7e1117d-be67-463e-d53b-2dad56547730"
X
# + colab={"base_uri": "https://localhost:8080/"} id="UNIbp0P36SML" outputId="31dd1a80-f76a-4dd8-ac3a-bfd90767ec7f"
A = X+2
A
# + colab={"base_uri": "https://localhost:8080/"} id="HE9xPWPdcgu4" outputId="98d314e9-72d4-446e-ae4e-1d576b0f2af8"
A + X
# + colab={"base_uri": "https://localhost:8080/"} id="xKyCwGia6SMP" outputId="ec290553-dd16-4b66-f213-8d6b2f3e74bc"
A * X
# + id="B5jXGIBp6SMT"
A_pt = X_pt + 2
# + colab={"base_uri": "https://localhost:8080/"} id="A7k6yxu36SMU" outputId="29ce5692-9808-42fa-df99-ae65c9b6942f"
A_pt + X_pt
# + colab={"base_uri": "https://localhost:8080/"} id="r8vOul0m6SMW" outputId="286d0736-697b-4fc3-f867-4907f55e60e9"
A_pt * X_pt
# + id="rQcBMSb76SMQ"
A_tf = X_tf + 2
# + colab={"base_uri": "https://localhost:8080/"} id="x6s1wtNj6SMR" outputId="7f4ce761-2b4d-4d1e-c0c0-d2cf3b2d720b"
A_tf + X_tf
# + colab={"base_uri": "https://localhost:8080/"} id="J1D7--296SMS" outputId="9a894404-ad84-4673-b12d-537ffabb628f"
A_tf * X_tf
# + [markdown] id="FE5f-FEq6SMY"
# ### Reduction
# + [markdown] id="WPJ9FVQF6SMY"
# Calculating the sum across all elements of a tensor is a common operation. For example:
#
# * For vector ***x*** of length *n*, we calculate $\sum_{i=1}^{n} x_i$
# * For matrix ***X*** with *m* by *n* dimensions, we calculate $\sum_{i=1}^{m} \sum_{j=1}^{n} X_{i,j}$
# + colab={"base_uri": "https://localhost:8080/"} id="rXi2stvz6SMZ" outputId="0f8e0e4a-41ec-4e61-e0e1-8c2ba6d813ad"
X
# -
X.shape
# + colab={"base_uri": "https://localhost:8080/"} id="W9FKaJbf6SMZ" outputId="f6f80b20-b729-41fe-98b3-0869d2da2a43"
X.sum()
# + colab={"base_uri": "https://localhost:8080/"} id="3y9aw7t66SMc" outputId="6fe55519-d388-4bd7-93cf-d015ed83f1bd"
torch.sum(X_pt)
# + colab={"base_uri": "https://localhost:8080/"} id="wcjRtFml6SMb" outputId="a5412adc-4d2b-49e6-9be6-2fbf8ad115d1"
tf.reduce_sum(X_tf)
# + colab={"base_uri": "https://localhost:8080/"} id="awjH9bOz6SMc" outputId="6a6e021a-0517-4384-fb21-78752023a267"
# Can also be done along one specific axis alone, e.g.:
X.sum(axis=0) # summing all rows
# + colab={"base_uri": "https://localhost:8080/"} id="n2SASjsn6SMd" outputId="f47594a4-0b2b-4b6b-9c9e-6d68f2267e31"
X.sum(axis=1) # summing all columns
# + colab={"base_uri": "https://localhost:8080/"} id="uVnSxvSJ6SMh" outputId="d0c31372-0f93-4612-f5ad-dcf0a5c71f23"
torch.sum(X_pt, 0)
# + colab={"base_uri": "https://localhost:8080/"} id="IO8drxz36SMe" outputId="96277393-de24-41fe-d136-3e03c266f21a"
tf.reduce_sum(X_tf, 1)
# + [markdown] id="gdAe8S4A6SMj"
# Many other operations can be applied with reduction along all or a selection of axes, e.g.:
#
# * maximum
# * minimum
# * mean
# * product
#
# They're fairly straightforward and used less often than summation, so you're welcome to look them up in library docs if you ever need them.
# + [markdown] id="r2eW8S_46SMj"
# ### The Dot Product
# + [markdown] id="LImETgD76SMj"
# If we have two vectors (say, ***x*** and ***y***) with the same length *n*, we can calculate the dot product between them. This is annotated several different ways, including the following:
#
# * $x \cdot y$
# * $x^Ty$
# * $\langle x,y \rangle$
#
# Regardless which notation you use (I prefer the first), the calculation is the same; we calculate products in an element-wise fashion and then sum reductively across the products to a scalar value. That is, $x \cdot y = \sum_{i=1}^{n} x_i y_i$
#
# The dot product is ubiquitous in deep learning: It is performed at every artificial neuron in a deep neural network, which may be made up of millions (or orders of magnitude more) of these neurons.
# + colab={"base_uri": "https://localhost:8080/"} id="HveIE3IDcgvP" outputId="b3492930-1884-47c2-fe92-252014423e14"
x
# + colab={"base_uri": "https://localhost:8080/"} id="3ZjkZcvVcgvQ" outputId="7cf7ffb7-4bb0-469d-fa7e-96a763788f58"
y = np.array([0, 1, 2])
y
# + colab={"base_uri": "https://localhost:8080/"} id="Xu8z0QB0cgvR" outputId="15db93f0-52ee-458e-cf77-a9f3b786d4b4"
25*0 + 2*1 + 5*2
# + colab={"base_uri": "https://localhost:8080/"} id="ThehRrr8cgvS" outputId="5e49b22e-b624-4e59-e038-e42f6b6c1335"
np.dot(x, y)
# + colab={"base_uri": "https://localhost:8080/"} id="J5Zdua4xcgvT" outputId="6d0a6105-a22c-4093-b981-4e1e3f20146f"
x_pt
# + colab={"base_uri": "https://localhost:8080/"} id="b3vEdroXcgvU" outputId="e6c250f0-61ff-4ef4-c293-86fab908b12c"
y_pt = torch.tensor([0, 1, 2])
y_pt
# + colab={"base_uri": "https://localhost:8080/"} id="F741E5imcgvV" outputId="4a8eb07c-c966-48e1-a4d2-dcc3b27cd152"
np.dot(x_pt, y_pt)
# + colab={"base_uri": "https://localhost:8080/"} id="-W5loHc8cgvX" outputId="f085e416-fb3d-4770-b12e-e11d83449541"
torch.dot(torch.tensor([25, 2, 5.]), torch.tensor([0, 1, 2.]))
# + colab={"base_uri": "https://localhost:8080/"} id="jUwKBiqzcgvY" outputId="04269f70-f5b7-4c05-916c-b9ae661d2a9b"
x_tf
# + colab={"base_uri": "https://localhost:8080/"} id="Xqt3Rac7cgvZ" outputId="ab42de2c-42af-488d-b215-ec4586808b56"
y_tf = tf.Variable([0, 1, 2])
y_tf
# + colab={"base_uri": "https://localhost:8080/"} id="x4pgc5JEcgvc" outputId="f60a18b9-1766-437b-cfd9-062e4a10428c"
tf.reduce_sum(tf.multiply(x_tf, y_tf))
| notebooks/01a_numpy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
#
# So far we have worked on EDA. This lab will focus on data cleaning and wrangling from everything we noticed before.
#
# We will start with removing outliers. So far, we have discussed different methods to remove outliers. Use the one you feel more comfortable with, define a function for that. Use the function to remove the outliers and apply it to the dataframe.
# Create a copy of the dataframe for the data wrangling.
# Normalize the continuous variables. You can use any one method you want.
# Encode the categorical variables
# The time variable can be useful. Try to transform its data into a useful one. Hint: Day week and month as integers might be useful.
# Since the model will only accept numerical data, check and make sure that every column is numerical, if some are not, change it using encoding.
import numpy as np
import pandas as pd
import datetime
pd.set_option('display.max_columns', None)
import warnings
warnings.filterwarnings('ignore')
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
data = pd.read_csv("we_fn_use_c_marketing_customer_value_analysis.csv")
data.head()
data.shape
numericals = data.select_dtypes(np.number)
numericals.head()
data = data.drop(['month'],axis=1)
numericals.isnull().sum()/len(data)
sns.boxplot(x ='Customer Lifetime Value', data=data)
plt.show()
sns.boxplot(x ='Monthly Premium Auto', data=data)
plt.show()
sns.boxplot(x ='Number of Policies', data=data)
plt.show()
sns.boxplot(x ='Total Claim Amount', data=data)
plt.show()
# Dealing with outliers
iqr = np.percentile(data['Customer Lifetime Value'],75) - np.percentile(data['Customer Lifetime Value'],25)
upper_limit = np.percentile(data['Customer Lifetime Value'],75) + 1.5*iqr
lower_limit = np.percentile(data['Customer Lifetime Value'],25) - 1.5*iqr
# Normalization
data = data[(data['Customer Lifetime Value']>lower_limit) & (data['Customer Lifetime Value']<upper_limit)]
# Dealing with outliers
iqr = np.percentile(data['Monthly Premium Auto'],75) - np.percentile(data['Monthly Premium Auto'],25)
upper_limit = np.percentile(data['Monthly Premium Auto'],75) + 1.5*iqr
lower_limit = np.percentile(data['Monthly Premium Auto'],25) - 1.5*iqr
# Normalization
data = data[(data['Monthly Premium Auto']>lower_limit) & (data['Monthly Premium Auto']<upper_limit)]
# Dealing with outliers
iqr = np.percentile(data['Number of Policies'],75) - np.percentile(data['Number of Policies'],25)
upper_limit = np.percentile(data['Number of Policies'],75) + 1.5*iqr
lower_limit = np.percentile(data['Number of Policies'],25) - 1.5*iqr
# Normalization
data = data[(data['Number of Policies']>lower_limit) & (data['Number of Policies']<upper_limit)]
# Dealing with outliers
iqr = np.percentile(data['Total Claim Amount'],75) - np.percentile(data['Total Claim Amount'],25)
upper_limit = np.percentile(data['Total Claim Amount'],75) + 1.5*iqr
lower_limit = np.percentile(data['Total Claim Amount'],25) - 1.5*iqr
# Normalization
data = data[(data['Total Claim Amount']>lower_limit) & (data['Total Claim Amount']<upper_limit)]
data_new = data
# # Normalization
from sklearn.preprocessing import Normalizer
X = data_new.select_dtypes(include=np.number)
transformer = Normalizer().fit(X)
X_normalised= transformer.transform(X)
X_normalised = pd.DataFrame(X_normalised)
# # Transforming time variables (day, week and month) to integers.
data_new['day'] = pd.DatetimeIndex(data_new['Effective To Date']).day
data_new['month'] = pd.DatetimeIndex(data_new['Effective To Date']).month
data_new['year'] = pd.DatetimeIndex(data_new['Effective To Date']).year
data_new.head()
# Droping Effective To date column
data_new = data_new.drop(['Effective To Date'], axis=1)
# df_new['effective_to_date'] = df_new['day'].map(str) + df_new['month'].map(str) + df_new['year'].map(str)
data_new
# data_new = data_new.drop(['day'], axis=1)
# data_new = data_new.drop(['month'], axis=1)
# data_new = data_new.drop(['year'], axis=1)
# data_new = data_new.drop(['customer'], axis=1) # customer id is not a value
# df_new['Effective To date'] = df_new['effective_to_date'].astype('int')
data_new['day'] = data_new['day'].astype('int')
data_new['month'] = data_new['month'].astype('int')
data_new['month'] = data_new['month'].astype('int')
data_new
data_new.info()
# # Since the model will only accept numerical data, check and make sure that every column is numerical, if some are not, change it using encoding.
#
cat = data_new.select_dtypes(include = np.object)
# cat
col = cat.columns
col
categorical=pd.get_dummies(cat, columns =['State', 'Response', 'Coverage', 'Education',
'Gender', 'Location Code',
'Policy Type', 'Policy', 'Renew Offer Type',
'Sales Channel', 'Vehicle Class', 'Vehicle Size'],drop_first=False)
categorical.head()
| data-cleaning-and-wrangling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Problem Definition
# A gold seeker needs to find at least 2 kg of bronze and 3 kg of copper to pay his gambling debts. There are two rivers in
# which the miner can find these precious metals. Each day the miner spends in river A, he finds 0.3 kg of bronze and 0.3
# kg of copper while each day he spends in river B, he finds 0.2 kg of bronze and 0.4 kg of copper.
#
# *Can you help this gold seeker decide how many days to spend in each day to pay his debts as soon as possible?*
# ### Problem Model
# **Decision variables**
#
# $x_{A}$ Days spent in river A
#
# $x_{B}$ Days spent in river B
#
# **Objective function**
#
# min $z = x_{A} + x_{B}$
#
# **Constraints**
#
# $0.3*x_{A} + 0.2*x_{B} \geq 2$ Bronze constrarint
#
# $0.3*x_{A} + 0.4*x_{B} \geq 3$ Copper constraint
| docs/source/CLP/solved/All Gold (Model Solved).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python
# language: python3
# name: python3
# ---
# # Exercise List from section2 (all)
#
# This file should have all exercises, even ones not defined in section2
#
#
# <a id='exerciselist-0'></a>
# **Exercise 1 (../exercise_list_all)**
#
# This is a new exercise. It should be repeated again in the list below, in addition to exercises from other files
#
# ([*back to text*](exercise_list_all.ipynb#exercise-0))
#
# **Exercise 1 (../exercise_list_labels)**
#
# This is an exercise from the `exercise_list_labels` file
#
# ([*back to text*](exercise_list_labels.ipynb#exercise-0))
#
# **Exercise 1 (../exercises)**
#
# This is a note that has some _italic_ and **bold** embedded
#
# - list
# - in
# - exercise
# + [markdown] hide-output=false
# ```python
# def foobar(x, y, z):
# print(x, y, z)
# ```
#
# -
# And text after the code block
#
# below is something that should be a real code block
# + hide-output=false
def foobar(x, y, z):
print(x, y, z)
# -
# And text to follow
#
# ([*back to text*](exercises.ipynb#exercise-0))
#
# **Exercise 2 (../exercises)**
#
# This is a normal exercise
#
# ([*back to text*](exercises.ipynb#exercise-1))
#
# **Exercise 3 (../exercises)**
#
# I'm a function with a label and a solution
#
# Define a function named `var` that takes a list (call it `x`) and
# computes the variance. This function should use the mean function that we
# defined earlier.
#
# Hint: $ \text{variance} = \frac{1}{N} \sum_i (x_i - \text{mean}(x))^2 $
# + hide-output=false
# your code here
# -
# ([*back to text*](exercises.ipynb#exercise-2))
#
# **Exercise 4 (../exercises)**
#
# This is another function with a label
#
# - and
# - *a*
# - **list**!
#
#
# ([*back to text*](exercises.ipynb#exercise-3))
#
# **Exercise 1 (exercise_list_sec2)**
#
# I am defined in the exercise_list_sec2 file
#
# ([*back to text*](exercise_list_sec2.ipynb#exercise-0))
#
# **Exercise 1 (exercises_section2)**
#
# Hello, I am an exercise from section2/exercises_section2.rst
#
# ([*back to text*](exercises_section2.ipynb#exercise-0))
#
# **Exercise 2 (exercises_section2)**
#
# Hi, I am also an exercise from section2/exercises_section2.rst
#
# ([*back to text*](exercises_section2.ipynb#exercise-1))
| tests/no_inline_exercises/ipynb/section2/.ipynb_checkpoints/exercise_list_sec2_all-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
niaaa_path = "../resources/niaaa_data_txt_to_csv.csv"
niaaa_df = pd.read_csv(niaaa_path,header=None)
niaaa_df
niaaa_col_path = "../resources/NIH NIAAA CSV FILE - COLUMN HEADERS.csv"
niaaa_col_df = pd.read_csv(niaaa_col_path,header=None)
niaaa_col_df
niaaa_columns = niaaa_col_df[1]
niaaa_columns
niaaa_df.columns = [niaaa_columns]
niaaa_df
niaaa_df_norm = niaaa_df.rename(columns = {"Geographic ID code (FIPS code, see specification below)" : "Geo Code"})
niaaa_df_norm
niaaa_geo_code_path = "../resources/niaaa_data_geographic_id_code.csv"
niaaa_geo_code_df = pd.read_csv(niaaa_geo_code_path,header=None)
niaaa_geo_code_df.columns = ["code","state_region"]
niaaa_geo_code_df
niaaa_df_geo = pd.merge(niaaa_df_norm, niaaa_geo_code_df, left_on ="Geo Code",
right_on="code", how="left")
niaaa_df_geo
niaaa_df['state_region'] = niaaa_df.lookup(df['best'])
| alec-work/normalizing niaaa data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="uD9ATtjvfV73" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 419} outputId="793f667d-d6f9-4c3a-caac-7ae446b2fadd"
import pandas as pd
import numpy as np
import plotly.express as px
# praying_df = pd.read_csv(r"C:\Users\Z Dubs\lambda\labs_week1\pop_data\historical_pop_final.csv", encoding='utf-8')
df = pd.read_csv('historical_pop_final.csv', encoding='utf-8')
df
# + id="V2_jm3OyMMqB" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 419} outputId="cee060f4-e06a-4a4b-f78f-28f45b6b10d1"
df['city_state'] = (df['city'] + "," + " " + df['state'])
df
# + id="_WRimcPp5wxF" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 272} outputId="ec03aaa4-d792-4193-ee6b-95f3b5444dff"
df.dtypes
# + id="c5H9RwgnsC0h" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 221} outputId="0b078a41-2bd8-4250-b8b5-27c2d4c9b125"
df['city']
# + id="IsGxq-HO5b7L" colab_type="code" colab={}
sample = df[(df.city == city) & (df.state == state) & (df.year == 2018)]
sample = sample.to_dict()
# + id="gqIkUfkRknqJ" colab_type="code" colab={}
sample2 = df[(df.city == city) & (df.state == state) & (df.year == 2018)]
sample2 = sample2.to_numpy()
# + id="cihKdO57k1em" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="684b1fff-33ee-45d9-a10d-84419968eea4"
sample2
# + id="cYA3JJq76C7g" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 255} outputId="495ea89b-1ddb-449c-b56c-dea7e1a58649"
sample
# + id="2uTQfk_5gFWq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 542} outputId="6864dbdb-b009-40c5-8357-d9e464225d5a"
city = 'Seattle'
state = 'WA'
metric = 'total_pop'
subset = df[(df.city == city) & (df.state == state)]
fig = px.line(subset, x='year', y=metric, title=f'{metric} in {city},{state}')
fig.show()
# + id="aPVgeNK_is5s" colab_type="code" colab={}
def citypopviz(metric = 'total_pop', city, state):
"""
Visualize historical population metrics from 2010 to 2018 for one city
### Query Parameters:
- `metric`: 'total_pop', 'male_pop', 'female_pop', 'age_under_20',
'age_20-29', 'age_30-39', 'age_40-49', 'age_50-59', or 'age_above_60';
default='total_pop',case sensitive, total/male/female pop in thousands,
age demographics in percentages
- `city`: [city name], case sensitive(ex: Birmingham)
- `state `: [state abbreviation], 2-letters; case sensitive (ex: AL)
### Response
JSON string to render with react-plotly.js
"""
df = pd.read_csv('9yr_city_pop_data.csv', encoding='utf-8')
subset = df[(df.city == city) & (df.state == state)]
fig = px.line(subset, x='year', y=metric, title=f'{metric} in {city}')
return fig.to_json()
# + id="Imy362A5lX3V" colab_type="code" colab={}
@router.get('/historic_pop_data'/{city_id})
def pop_to_dict(city_id):
"""
Pull demographic data for specific city, state, and year
### Query Parameters:
- `city_id`: [city_id], unique numeric mapping (ex: 0 returns Anchorage, AK)
### Response
Dictionary object
"""
rt_dict = {}
rt_data_dict = {}
df = pd.read_csv('100city_population_data_2018.csv', encoding='utf-8')
rt_data = df[df['city_id'] == city_id]
rt_data_dict["total_pop"] = rt_data[0][4]
rt_data_dict["male_pop"] = rt_data[0][5]
rt_data_dict["female_pop"] = rt_data[0][6]
rt_data_dict["age_under_20"] = rt_data[0][7]
rt_data_dict["age_20-29"] = rt_data[0][8]
rt_data_dict["age_30-39"] = rt_data[0][9]
rt_data_dict["age_40-49"] = rt_data[0][10]
rt_data_dict["age_50-59"] = rt_data[0][11]
rt_data_dict["age_above_60"] = rt_data[0][12]
rt_dict["data"] = rt_data_dict
rt_dict["viz"] = citypopviz(city= rt_data[0][1], state= rt_data[0][2])
return rt_dict
# + id="vPlcbeQL2OZz" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 419} outputId="107b543c-bd32-40ff-ba50-cfa4d8c7b070"
df = pd.read_csv('100city_population_data_2018.csv', encoding='utf-8')
df
# + id="cYQl7RTXI3Oq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 419} outputId="94bbbaa7-e6f5-4fb3-8261-f7f6f30769d8"
df2 = pd.read_csv('cities.csv', encoding='utf-8')
df2.columns = ['city_state']
df2
# + id="9nCp0meBKDy7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 419} outputId="94ff3033-1b71-4418-b846-06167e0c1fc9"
merged = pd.merge(df, df2, on=['city_state'], how='right')
merged
# + id="i0pDoD1xyz8N" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 419} outputId="62e7c3e3-23ed-4612-91a6-a326f0253c92"
merged = merged.drop(['city_id'], axis=1)
merged
# + id="H5JZOqJJM0Hn" colab_type="code" colab={}
len(merged['city'].unique())
# + id="Um9wcucSNeOL" colab_type="code" colab={}
pd.DataFrame.to_csv(merged, '9yr_city_pop_data.csv', sep=',', na_rep='NaN', index=False)
# + id="Qt7mo0R1IeUI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 419} outputId="e5e84673-e59a-4e6f-a41b-270cf846879e"
df18 = df[(df['year'] == 2018)]
df18 = df18.drop(['city_id'], axis=1)
df18
# + id="zU7nBXyoI0B5" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 419} outputId="4c94d525-5ce1-4af3-c544-789af4c1ac60"
merged18 = pd.merge(df18, df2, on=['city_state'], how='right')
merged18
# + id="CA9dJwvTKFjS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 419} outputId="8cd5ad50-0f78-47a0-c077-5f8663774164"
merged18 = merged18.sort_values(["state", "city"], ascending = (True, True))
merged18
# + id="1PVDiZMMKvGQ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 419} outputId="77b9c9a9-bc63-4947-da1c-48cdcc1d3cbf"
# merged18.insert(0, 'city_id', range(0, 0 + len(merged18)))
merged18
# + id="YznjeqOElaHS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 80} outputId="422a63a3-1443-4452-e991-2096e8eb4278"
sub_merged18 = merged18[(merged18.city == city) & (merged18.state == state)]
# sub_merged18 = sub_merged18.to_numpy()
sub_merged18
# + id="SWnnfQ5dwKeB" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 170} outputId="2b158e8a-28cb-4867-96e3-01a70eee69a3"
rt_dict = {"total_pop" : sub_merged18[0][3],
"male_pop" : sub_merged18[0][4],
"female_pop" : sub_merged18[0][5],
"age_under_20" : sub_merged18[0][6],
"age_20-29" : sub_merged18[0][7],
"age_30-39" : sub_merged18[0][8],
"age_40-49" : sub_merged18[0][9],
"age_50-59" : sub_merged18[0][10],
"age_above_60" : sub_merged18[0][11],
}
rt_dict
# + id="bugRR_FElsHZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 80} outputId="2c658a38-78c2-4485-df44-1773242184da"
sub_merged18
# + id="gKKF441vNSx_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="83d78405-6ea1-4361-ca0c-a63e4f22a987"
json_df = sub_merged18.to_json()
print(json_df)
# + id="mze7C-qcg1S5" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 323} outputId="a52c74d2-c3df-495f-bab9-1430732d28d5"
clean_merged18 = merged18[['city_id','city','state','city_state']]
clean_merged18
# + id="GY7kk-cGamjR" colab_type="code" colab={}
pd.DataFrame.to_csv(clean_merged18, '100city_state_data.csv', sep=',', na_rep='NaN', index=False)
# + id="V7m5R7Ssyu0b" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 419} outputId="385dc311-aea9-4b09-d883-5eadf7f87199"
new_df = pd.read_csv('100city_state_data.csv', encoding='utf-8')
new_df
| notebooks/Citrics_Historical_Population_Analysis_&_Visualization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# %matplotlib inline
#
# .. _tut_stats_cluster_methods:
#
# # Permutation t-test on toy data with spatial clustering
#
#
# Following the illustrative example of Ridgway et al. 2012,
# this demonstrates some basic ideas behind both the "hat"
# variance adjustment method, as well as threshold-free
# cluster enhancement (TFCE) methods in mne-python.
#
# This toy dataset consists of a 40 x 40 square with a "signal"
# present in the center (at pixel [20, 20]) with white noise
# added and a 5-pixel-SD normal smoothing kernel applied.
#
# For more information, see:
# Ridgway et al. 2012, "The problem of low variance voxels in
# statistical parametric mapping; a new hat avoids a 'haircut'",
# NeuroImage. 2012 Feb 1;59(3):2131-41.
#
# Smith and Nichols 2009, "Threshold-free cluster enhancement:
# addressing problems of smoothing, threshold dependence, and
# localisation in cluster inference", NeuroImage 44 (2009) 83-98.
#
# In the top row plot the T statistic over space, peaking toward the
# center. Note that it has peaky edges. Second, with the "hat" variance
# correction/regularization, the peak becomes correctly centered. Third,
# the TFCE approach also corrects for these edge artifacts. Fourth, the
# the two methods combined provide a tighter estimate, for better or
# worse.
#
# Now considering multiple-comparisons corrected statistics on these
# variables, note that a non-cluster test (e.g., FDR or Bonferroni) would
# mis-localize the peak due to sharpness in the T statistic driven by
# low-variance pixels toward the edge of the plateau. Standard clustering
# (first plot in the second row) identifies the correct region, but the
# whole area must be declared significant, so no peak analysis can be done.
# Also, the peak is broad. In this method, all significances are
# family-wise error rate (FWER) corrected, and the method is
# non-parametric so assumptions of Gaussian data distributions (which do
# actually hold for this example) don't need to be satisfied. Adding the
# "hat" technique tightens the estimate of significant activity (second
# plot). The TFCE approach (third plot) allows analyzing each significant
# point independently, but still has a broadened estimate. Note that
# this is also FWER corrected. Finally, combining the TFCE and "hat"
# methods tightens the area declared significant (again FWER corrected),
# and allows for evaluation of each point independently instead of as
# a single, broad cluster.
#
# Note that this example does quite a bit of processing, so even on a
# fast machine it can take a few minutes to complete.
#
#
# +
# Authors: <NAME> <<EMAIL>>
# License: BSD (3-clause)
import numpy as np
from scipy import stats
from functools import partial
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D # noqa; this changes hidden mpl vars
from mne.stats import (spatio_temporal_cluster_1samp_test,
bonferroni_correction, ttest_1samp_no_p)
try:
from sklearn.feature_extraction.image import grid_to_graph
except ImportError:
from scikits.learn.feature_extraction.image import grid_to_graph
print(__doc__)
# -
# Set parameters
# --------------
#
#
width = 40
n_subjects = 10
signal_mean = 100
signal_sd = 100
noise_sd = 0.01
gaussian_sd = 5
sigma = 1e-3 # sigma for the "hat" method
threshold = -stats.distributions.t.ppf(0.05, n_subjects - 1)
threshold_tfce = dict(start=0, step=0.2)
n_permutations = 1024 # number of clustering permutations (1024 for exact)
# Construct simulated data
# ------------------------
#
# Make the connectivity matrix just next-neighbor spatially
#
#
# +
n_src = width * width
connectivity = grid_to_graph(width, width)
# For each "subject", make a smoothed noisy signal with a centered peak
rng = np.random.RandomState(42)
X = noise_sd * rng.randn(n_subjects, width, width)
# Add a signal at the dead center
X[:, width // 2, width // 2] = signal_mean + rng.randn(n_subjects) * signal_sd
# Spatially smooth with a 2D Gaussian kernel
size = width // 2 - 1
gaussian = np.exp(-(np.arange(-size, size + 1) ** 2 / float(gaussian_sd ** 2)))
for si in range(X.shape[0]):
for ri in range(X.shape[1]):
X[si, ri, :] = np.convolve(X[si, ri, :], gaussian, 'same')
for ci in range(X.shape[2]):
X[si, :, ci] = np.convolve(X[si, :, ci], gaussian, 'same')
# -
# Do some statistics
# ------------------
#
# .. note::
# X needs to be a multi-dimensional array of shape
# samples (subjects) x time x space, so we permute dimensions:
#
#
X = X.reshape((n_subjects, 1, n_src))
# Now let's do some clustering using the standard method.
#
# .. note::
# Not specifying a connectivity matrix implies grid-like connectivity,
# which we want here:
#
#
# +
T_obs, clusters, p_values, H0 = \
spatio_temporal_cluster_1samp_test(X, n_jobs=1, threshold=threshold,
connectivity=connectivity,
tail=1, n_permutations=n_permutations)
# Let's put the cluster data in a readable format
ps = np.zeros(width * width)
for cl, p in zip(clusters, p_values):
ps[cl[1]] = -np.log10(p)
ps = ps.reshape((width, width))
T_obs = T_obs.reshape((width, width))
# To do a Bonferroni correction on these data is simple:
p = stats.distributions.t.sf(T_obs, n_subjects - 1)
p_bon = -np.log10(bonferroni_correction(p)[1])
# Now let's do some clustering using the standard method with "hat":
stat_fun = partial(ttest_1samp_no_p, sigma=sigma)
T_obs_hat, clusters, p_values, H0 = \
spatio_temporal_cluster_1samp_test(X, n_jobs=1, threshold=threshold,
connectivity=connectivity,
tail=1, n_permutations=n_permutations,
stat_fun=stat_fun)
# Let's put the cluster data in a readable format
ps_hat = np.zeros(width * width)
for cl, p in zip(clusters, p_values):
ps_hat[cl[1]] = -np.log10(p)
ps_hat = ps_hat.reshape((width, width))
T_obs_hat = T_obs_hat.reshape((width, width))
# Now the threshold-free cluster enhancement method (TFCE):
T_obs_tfce, clusters, p_values, H0 = \
spatio_temporal_cluster_1samp_test(X, n_jobs=1, threshold=threshold_tfce,
connectivity=connectivity,
tail=1, n_permutations=n_permutations)
T_obs_tfce = T_obs_tfce.reshape((width, width))
ps_tfce = -np.log10(p_values.reshape((width, width)))
# Now the TFCE with "hat" variance correction:
T_obs_tfce_hat, clusters, p_values, H0 = \
spatio_temporal_cluster_1samp_test(X, n_jobs=1, threshold=threshold_tfce,
connectivity=connectivity,
tail=1, n_permutations=n_permutations,
stat_fun=stat_fun)
T_obs_tfce_hat = T_obs_tfce_hat.reshape((width, width))
ps_tfce_hat = -np.log10(p_values.reshape((width, width)))
# -
# Visualize results
# -----------------
#
#
# +
fig = plt.figure(facecolor='w')
x, y = np.mgrid[0:width, 0:width]
kwargs = dict(rstride=1, cstride=1, linewidth=0, cmap='Greens')
Ts = [T_obs, T_obs_hat, T_obs_tfce, T_obs_tfce_hat]
titles = ['T statistic', 'T with "hat"', 'TFCE statistic', 'TFCE w/"hat" stat']
for ii, (t, title) in enumerate(zip(Ts, titles)):
ax = fig.add_subplot(2, 4, ii + 1, projection='3d')
ax.plot_surface(x, y, t, **kwargs)
ax.set_xticks([])
ax.set_yticks([])
ax.set_title(title)
p_lims = [1.3, -np.log10(1.0 / n_permutations)]
pvals = [ps, ps_hat, ps_tfce, ps_tfce_hat]
titles = ['Standard clustering', 'Clust. w/"hat"',
'Clust. w/TFCE', 'Clust. w/TFCE+"hat"']
axs = []
for ii, (p, title) in enumerate(zip(pvals, titles)):
ax = fig.add_subplot(2, 4, 5 + ii)
plt.imshow(p, cmap='Purples', vmin=p_lims[0], vmax=p_lims[1])
ax.set_xticks([])
ax.set_yticks([])
ax.set_title(title)
axs.append(ax)
plt.tight_layout()
for ax in axs:
cbar = plt.colorbar(ax=ax, shrink=0.75, orientation='horizontal',
fraction=0.1, pad=0.025)
cbar.set_label('-log10(p)')
cbar.set_ticks(p_lims)
cbar.set_ticklabels(['%0.1f' % p for p in p_lims])
plt.show()
| 0.12/_downloads/plot_stats_cluster_methods.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.1.0
# language: julia
# name: julia-1.1
# ---
# # Disease Spread Simulation
# ## Summary
# This model simulates the process of epidemic disease spread. In particular, it raises the problem of **herd immunity**, which is a form of defence against epidemic diseases, which protects people that can't (or won't) be vaccinated, because when the majority of population is vaccinated (immune), then the disease can't spread.
#
# The greater the proportion of immune ones (either in natural way or vaccinated), the lower the chance that those endangered will have contact with the disease. In case of, for example measles or smallpox, the *herd immunity threshold* is estimated to be 92-95% for the first one and 80-86% for the latter.
# The contact with infected person is necessary for these diseases to spread.
#
# The model performs a simulation of a hypothetical disease, that can be spread by contact with infected ones.
#
# __Sources__
# * <NAME>., <NAME>., <NAME>.. "Herd immunity": A rough guide. „Clinical Infectious Diseases”. 52 (7), s. 911–6, April 2011. https://academic.oup.com/cid/article/52/7/911/299077
# * https://ourworldindata.org/vaccination#how-vaccines-work-herd-immunity-and-reasons-for-caring-about-broad-vaccination-coverage
# ## Description
#
# The model simulates how multiple heterogeneous agents move around a map. There are 3 types of agents:
# 1. vaccinated [green]
# 2. unvaccinated [blue]
# 3. infected [red]
#
# Firstly, individuals' coordinates are generated, according to provided information about number of infected and vaccinated. During each iteration, individuals change their position on the map.
# At a new location, if infected one will be within a provided distance from not infected ones, it will have a chance (small in case of vaccinated, and high in case of unvaccinated ones) to spread the disease.
# In reality, even vaccinated people have very small chance to catch the disease.
# After every iteration the map with current situation is generated, showing what percentage of the whole population is infected. All the maps are saved in the working directory.
# ## Functions
using Random, PyPlot, DataFrames, Distributions
import Distributions: Uniform
function generate_data(population, square_side, perc_infected, perc_vaccinated, perc_unvaccinated)
"""
Generates each individual's coordinates on a map.
population - number of people
square_side - defines map size
perc_infected - % of population infected
perc_vaccinated - % of population vaccinated
perc_unvaccinated - % of population not vaccinated (or can't be)
"""
perc_list = [perc_infected, perc_vaccinated, perc_unvaccinated]
@assert sum(perc_list) == 1
types = ["infected", "vaccinated", "unvaccinated"]
df = DataFrame(x = Float64[], y = Float64[], typ = String[])
for (index, value) in enumerate(perc_list)
n_people = Int(round(population*value))
x = rand(Uniform(1, square_side), n_people)
y = rand(Uniform(1, square_side), n_people)
typ = types[index]
df_temp = DataFrame(x = x, y = y, typ = typ)
df = join(df, df_temp, kind = :outer, on = intersect(names(df), names(df_temp)))
end
return df
end
function make_plot(df, Title="")#, podtytul="")
"""
Makes map with data generated previously
data - df created previously
colors:
infected - red
vaccinated - green
unvaccinated - blue
"""
for person in unique(df[:typ])
data = df[(df[:typ] .== person), :]
if person == "infected"
color = "red"
elseif person == "vaccinated"
color = "green"
elseif person == "unvaccinated"
color = "blue"
end
PyPlot.scatter(data.x, data.y, alpha=0.4, c=color)
end
title(Title)
end
function distance(x_axis_1, y_axis_1, x_axis_2, y_axis_2)
"""
Calculates distance between 2 points (Pythagorean theorem)
x_axis_1, y_axis_1 - coordinates of the first point
x_axis_2, y_axis_2 - coordinates of the second point
"""
a=abs(x_axis_1-x_axis_2)
b=abs(y_axis_1-y_axis_2)
c = sqrt(a^2 + b^2)
return c
end
function new_positions(df, square_side)
"""
Gets new random positons for individuals
"""
df.x = rand(Uniform(1, square_side), nrow(df))
df.y = rand(Uniform(1, square_side), nrow(df))
return df
end
function simulation(
population, square_side, perc_infected, perc_vaccinated, perc_unvaccinated,
infection_distance, infection_prob_vaccinated, infection_prob_perc_unvaccinated,
simulation_time, when_stop)
data = generate_data(
population,
square_side,
perc_infected,
perc_vaccinated,
perc_unvaccinated)
# loop over time
for time in 1:simulation_time
# number of infected / the whole population
n_people = by(data, :typ, Sum = :typ => length)
n_all = sum(n_people.Sum)
n_infected = n_people[(n_people[:typ] .== "infected"), :Sum]
info_beginning = string(n_infected[1])*" / "*string(n_all)
title = "Start - time: "*string(time)*" | infected: "*info_beginning
make_plot(data, title)
plot_name="plot"*string(time)*"-beginning.png"
savefig(plot_name)
clf()
# 2 sets: infected i not_infected
not_infected = data[(data[:typ] .!= "infected"), :]
infected = data[(data[:typ] .== "infected"), :]
# loop over infected - because they may infect others
for ch in 1:nrow(infected)
# loop over not infected - because they may become infected
for nch in 1:nrow(not_infected)
# calculate distance between infected and others
dist_between = distance(
infected.x[ch],
infected.y[ch],
not_infected.x[nch],
not_infected.y[nch])
# if it's smaller than arbitrary set value (by us) - there is a chance for catching infection
if (dist_between <= infection_distance) & (not_infected.typ[nch] != "infected")
# vaccinated have very small chance to catch infection
if not_infected.typ[nch] == "vaccinated"
not_infected.typ[nch] = rand() < infection_prob_vaccinated ? "infected" : "vaccinated"
# not vaccinated have great chance to catch infection
elseif not_infected.typ[nch] == "unvaccinated"
not_infected.typ[nch] = rand() < infection_prob_perc_unvaccinated ? "infected" : "unvaccinated"
end
end
end
end
# merging two tables
data_after = join(not_infected, infected, kind = :outer, on = intersect(names(not_infected), names(infected)))
# number of infected compared to the whole population
n_people = by(data, :typ, Sum = :typ => length)
n_all = sum(n_people.Sum)
n_infected = n_people[(n_people[:typ] .== "infected"), :Sum]
info_end = string(n_infected[1])*" / "*string(n_all)
title = "End - time: "*string(time)*" | infected: "*info_end
make_plot(data_after, title)
plot_name="plot"*string(time)*"-end.png"
savefig(plot_name)
clf()
# current situation
infected_perc = n_infected[1]/n_all
# returns map after last iteration
if time == simulation_time
end_state = make_plot(data_after, title)
println("Percentage infected: "*string(infected_perc*100))
return (end_state, infected_perc)
end
# if `when_stop` is set to some value, function stops when % of infected reaches this value
if infected_perc >= when_stop
println("Percentage infected ("*string(round(infected_perc*100), 1)*") is over threshold: "*string(when_stop*100)*"%."*" Iteration: "*string(time))
end_state = make_plot(data_after, title)
return (end_state, infected_perc)
end
data = new_positions(data_after, square_side)
end
end
# ## Results
# ### Example
# +
res_1 = Float64[]
for i in 1:20
result = simulation(
500, # population
50, # square_side
0.01, # perc_infected
0.92, # perc_vaccinated
0.07, # perc_unvaccinated
1, # infection_distance
0.01, # infection_prob_vaccinated
0.8, # infection_prob_unvaccinated
50, # simulation_time
0.4) # when_stop
append!(res_1, result[2])
end
println(mean(res_1))
# +
res_2 = Float64[]
for i in 1:20
result = simulation(
500, # population
50, # square_side
0.01, # perc_infected
0.5, # perc_vaccinated
0.49, # perc_unvaccinated
1, # infection_distance
0.01, # infection_prob_vaccinated
0.8, # infection_prob_unvaccinated
50, # simulation_time
1) # when_stop
append!(res_2, result[2])
end
println(mean(res_2))
# -
# ### Analysis
# Let's take a look how does a model perform with the whole range of `perc_vaccinated`.
# +
x = [0.05:0.1:1;]
percentages = DataFrame(
perc_vaccinated = x,
perc_unvaccinated = 0.98 .- x,
perc_infected = 1 .- (x + y))
percentages
# +
final_result = Float64[]
res_tmp = Float64[]
for j in 1:nrow(percentages)
for i in 1:10
result = simulation(
500, # population
50, # square_side
percentages.perc_infected[j], # perc_infected
percentages.perc_vaccinated[j], # perc_vaccinated
percentages.perc_unvaccinated[j], # perc_unvaccinated
1, # infection_distance
0.01, # infection_prob_vaccinated
0.8, # infection_prob_unvaccinated
50, # simulation_time
1) # when_stop
append!(res_tmp, result[2])
end
append!(final_result, mean(res_tmp))
end
println(mean(res_tmp))
# -
percentages.result = final_result
percentages
plot(percentages.perc_vaccinated, percentages.result)
cor(percentages.perc_vaccinated, percentages.result)
# Results weren't surprising - the lower the percentage of vaccinated people, the lower final percentage of infected.
# Correlation: -0.99
# That'll be it. I'd appreciate all feedback :)
| disease_spread.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
data =pd.read_csv("cities.csv")
cities_df=pd.DataFrame(data)
cities_df =cities_df.set_index("City_ID")
cities_df.head()
cities_html=cities_df.to_html(buf=None, columns=None, col_space=None, header=True, index=True, na_rep='NaN', formatters=None, float_format=None, sparsify=None, index_names=True, justify=None, max_rows=None, max_cols=None, show_dimensions=False, decimal='.', bold_rows=True, classes=None, escape=True, notebook=False, border=None, table_id=None, render_links=False, encoding=None)
# + tags=[]
print(cities_html)
# -
| data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: basilcluster
# language: python
# name: basilcluster
# ---
from meshparty import trimesh_vtk, trimesh_io, skeleton_io, skeletonize, mesh_filters
import pandas as pd
import numpy as np
from scipy import sparse
import os
from itkwidgets import view
# %matplotlib notebook
cv_path = 'precomputed://gs://microns-seunglab/minnie65/seg_minnie65_0'
mesh_folder = 'minnie_meshes/'
skeleton_folder = 'minnie_skeletons/'
neuron_id = 103530771121793958
mm = trimesh_io.MeshMeta(cv_path=cv_path, disk_cache_path=mesh_folder)
mesh = mm.mesh(seg_id = neuron_id )
# +
# step 1
# convert your actors to vtkpolydata objects
poly_data = trimesh_vtk.trimesh_to_vtk(mesh.vertices, mesh.faces, None)
# step 2
# then create a viewer with this view function
# pass in polydata objects, what colors you want
# see docstring for more options
viewer=view(geometries=[poly_data],
geometry_colors=['m'],
ui_collapsed=True)
# -
| index.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.5 64-bit
# metadata:
# interpreter:
# hash: 2db524e06e9f5f4ffedc911c917cb75e12dbc923643829bf417064a77eb14d37
# name: python3
# ---
# libraries
import pandas as panda
import numpy as np
# Variables
exoplanetCatalogFileURL = 'data/exoplanet-catalog.csv'
earthMass = 0.003145701
earthRadius = 0.091130294
# +
# Prepare data and cleaning
CSVCatalog = panda.read_csv(exoplanetCatalogFileURL)
CSVCatalog = CSVCatalog.astype({'name': np.str_, 'binaryflag': np.int8, 'mass': np.float64, 'radius': np.float64 })
df = panda.DataFrame(data=CSVCatalog, columns=['name', 'binaryflag', 'mass', 'radius'])
# Filter null values
df = df.query('binaryflag == 0 and mass.notnull() > 0 and radius.notnull() > 0')
# convert mass in terms of earth mass
df.mass = df.mass.div(earthMass)
# convert radius in terms of earth radius
df.radius = df.radius.div(earthRadius)
# +
# Classification
# Filter by planet size, Criteria: mass 0.5 to 2 earth mass and radius
df = df.query('mass >= 0.5 and mass <= 2 and radius >= 0.5 and radius <= 1.5')
ax = df.plot(kind='scatter', title='Distribucion de planetas Masa vs Radio en terminos de la tierra', x='mass', y='radius', figsize=(10,10))
df[['mass','radius','name']].apply(lambda row: ax.text(*row),axis=1)
# Habitable Zone
| analisis-jupiter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Developer: <NAME>
# Implement a Max Heap
import math
# Max Heap
class Heap:
def __init__(self):
self.arr = [None]
# Insert value into heap
def insert(self, value):
# Add value to end of heap
self.arr.append(value)
# Sift it into place
self.sift_up(len(self.arr)-1)
# Sift value up through heap, needed for insert
def sift_up(self, index):
# If we're at the root, stop sifting
if index == 1:
return None
parent = math.floor(index/2)
# Swap values if greater
if self.arr[index] > self.arr[parent]:
temp = self.arr[parent]
self.arr[parent] = self.arr[index]
self.arr[index] = temp
return self.sift_up(parent)
else:
return None
# Return max item without removing it
def get_max(self):
# If we have an element return the first one since we are a max heap
if len(self.arr) > 1:
return self.arr[1]
return 'Error: empty heap'
# Return number of elements stored
def get_size(self):
return len(self.arr) - 1
# Return True if heap contains no elements
def is_empty(self):
# True if array is empty
return len(self.arr) < 2
# Returns max item and removes it
def extract_max(self):
if len(self.arr) < 2:
return 'Error: empty heap'
max_val = self.arr[1]
# Replace top of heap with the last element and sift down
self.arr[1] = len(self.arr) - 1
self.sift_down(1)
return max_val
# Sifts value down, needed for extract_max
def sift_down(self, index):
# We've reached the end of the heap. Stop sifting
if index*2 > len(self.arr):
return None
left = index * 2
right = index * 2 + 1
left_val = self.arr[left]
right_val = None
# Check that there's a right child
if right <= len(self.arr) - 1:
right_val = self.arr[right]
# Case 1: Right child is greater
if right_val and right_val > left_val:
self.arr[right] = self.arr[index]
self.arr[index] = right_val
return self.sift_down(right)
# Case 2: Left child is greater or equal
else:
self.arr[left] = self.arr[index]
self.arr[index] = left_val
return self.sift_down(left)
# Remove item at index x
def remove(self, i):
self.arr[i+1] = self.arr[-1]
self.arr.pop()
if i < len(self.arr) - 1:
self.sift_up(i+1)
self.sift_down(i+1)
# Create heap from array of elements. Needed for heap_sort
def heapify(self, array, n, i=0):
largest = i
left = i * 2 + 1
right = i * 2 + 2
print('ARRAY', array)
# If left child is larger
if left < n and array[left] > array[largest]:
largest = left
# If right child is largest
if right < n and array[right] > array[largest]:
largest = right
# If root isn't the largest, swap with the larger child
if largest != i:
array[i], array[largest] = array[largest], array[i]
return self.heapify(array, n, i=largest)
# Take unsorted array and turn into sorted array in-place
def heap_sort(self, array):
n = len(array)
print(array)
for i in range(n//2-1, -1, -1):
self.heapify(array, n, i)
for i in range(n-1, 0, -1):
array[i], array[0] = array[0], array[i]
self.heapify(array, i, 0)
# +
# Test the Max Heap and its methods
h = Heap()
# Test insert
print('##########')
print('Build heap')
h.insert(1)
h.insert(2)
h.insert(5)
h.insert(3)
h.insert(100)
h.insert(15)
h.insert(11)
print(h.arr)
print('##########\n')
# Test get_max
print('get_max', h.get_max())
# Test get_size
print('get_size', h.get_size())
# Test is_empty
print('is_empty', h.is_empty())
# Test extract_max
print('\nextract_max', h.extract_max())
print(h.arr)
# Test remove
print('\nRemoving element at index 3')
h.remove(3)
print(h.arr)
# Test heapify
H = [21,1,45,78,3,5]
print('\nHeapify array', H)
h.heapify(H, len(H))
print(H)
# Test heapsort
h.heap_sort(H)
# +
from heapq import heapify
H = [21,1,45,78,3,5]
print(H)
heapify(H)
print(H)
| datastructures/MaxHeap.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/pj-mathematician/gintama-ost-recognizer/blob/main/gintamaostfinder.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="28edfX_CbCnT"
# ##Choose the episode from the dropdown and select the time in Minute:Seconds
#
# (for example: to search 15:45 of Episode 4, select 4 in the drop down, slide to 15 in the *Minutes* slider and 45 in the *Seconds* slider
# + id="les96frxV2Z7" colab={"base_uri": "https://localhost:8080/"} outputId="a16a773b-cdd2-4d38-ecb9-5bc4d097eee0"
#@title Search OST by Episode number and time stamp { run: "auto", vertical-output: true, display-mode: "form" }
Episode = 4 #@param ['1','2','3','4','5','6','7','8', '9', '10', '11', '12', '13', '14', '15', '16', '17', '18', '19', '20', '21', '22', '23', '24', '25', '26', '27', '28', '29', '30', '31', '32', '33', '34', '35', '36', '37', '38', '39', '40', '41', '42', '43', '44', '45', '46', '47', '48', '49', '50', '51', '52', '53', '54', '55', '56', '57', '58', '59', '60', '61', '62', '63', '64', '65', '66', '67', '68', '69', '70', '71', '72', '73', '74', '75', '76', '77', '78', '79', '80', '81', '82', '83', '84', '85', '86', '87', '88', '89', '90', '91', '92', '93', '94', '95', '96', '97', '98', '99', '100', '101', '102', '103', '104', '105', '106', '107', '108', '109', '110', '111', '112', '113', '114', '115', '116', '117', '118', '119', '120', '121', '122', '123', '124', '125', '126', '127', '128', '129', '130', '131', '132', '133', '134', '135', '136', '137', '138', '139', '140', '141', '142', '143', '144', '145', '146', '147', '148', '149', '150', '151', '152', '153', '154', '155', '156', '157', '158', '159', '160', '161', '162', '163', '164', '165', '166', '167', '168', '169', '170', '171', '172', '173', '174', '175', '176', '177', '178', '179', '180', '181', '182', '183', '184', '185', '186', '187', '188', '189', '190', '191', '192', '193', '194', '195', '196', '197', '198', '199', '200', '201', '202', '203', '204', '205', '206', '207', '208', '209', '210', '211', '212', '213', '214', '215', '216', '217', '218', '219', '220', '221', '222', '223', '224', '225', '226', '227', '228', '229', '230', '231', '232', '233', '234', '235', '236', '237', '238', '239', '240', '241', '242', '243', '244', '245', '246', '247', '248', '249', '250', '251', '252', '253', '254', '255', '256', '257', '258', '259', '260', '261', '262', '263', '264', '265', '266', '267', '268', '269', '270', '271', '272', '273', '274', '275', '276', '277', '278', '279', '280', '281', '282', '283', '284', '285', '286', '287', '288', '289', '290', '291', '292', '293', '294', '295', '296', '297', '298', '299', '300', '301', '302', '303', '304', '305', '306', '307', '308', '309', '310', '311', '312', '313', '314', '315', '316', '317', '318', '319', '320', '321', '322', '323', '324', '325', '326', '327', '328', '329', '330', '331', '332', '333', '334', '335', '336', '337', '338', '339', '340', '341', '342', '343', '344', '345', '346', '347', '348', '349', '350', '351', '352', '353', '354', '355', '356', '357', '358', '359', '360', '361', '362', '363', '364', '365', '366', '367', '368', '369'] {type:"raw"}
#@markdown ---
Minutes = 15 #@param {type:"slider", min:0, max:25, step:1}
Seconds = 38 #@param {type:"slider", min:0, max:60, step:1}
st="ok"
#@markdown
print(Episode,str(Minutes)+":"+str(Seconds))
# + [markdown] id="oJglvsMfclHi"
# ###After selecting, click on the play button in the next cell (top left)
# + id="se1QmoDNdCnj" colab={"base_uri": "https://localhost:8080/"} outputId="91b2eab8-44da-4ae7-f22d-90b1f1b5c5f1"
#@title Start! { vertical-output: true, form-width: "20%", display-mode: "form" }
from IPython.utils import io
print("Starting the program...")
with io.capture_output() as captured:
#if True:
# !pip install -q kurby
# !pip install -q pydub
# !apt install sox
# !wget https://github.com/tsurumeso/vocal-remover/releases/download/v4.0.0/vocal-remover-v4.0.0.zip
# !unzip -u /content/vocal-remover-v4.0.0.zip
# !cd vocal-remover && pip install -r requirements.txt
# !git clone https://github.com/dpwe/audfprint
# !cd audfprint && pip install -r requirements.txt
# !wget -O ostp.pklz https://github.com/pj-mathematician/gintama-ost-recognizer/blob/main/ostp.pklz?raw=true
print("loading episode {}...".format(Episode))
with io.capture_output() as captured:
#if True:
# !kurby download --nfrom $Episode --nto $Episode gintama
print("seeking the time stamp and getting the audio...")
with io.capture_output() as captured:
#if True:
seconds=int(Minutes)*60 + int(Seconds)
if Episode < 10:
path = "/content/Gintama/Gintama-S00-E00{}.mp4".format(Episode)
elif Episode >= 10 and Episode <100:
path = "/content/Gintama/Gintama-S00-E0{}.mp4".format(Episode)
else:
path = "/content/Gintama/Gintama-S00-E{}.mp4".format(Episode)
# !ffmpeg -i $path master.wav
# !ffmpeg -ss $seconds -i master.wav -t 20 masterclip.wav
# !rm $path
# !rm master.wav
print("audioclip loaded. removing vocals...")
with io.capture_output() as captured:
#if True:
# !cd vocal-remover && python inference.py --input /content/masterclip.wav --tta --postprocess
# !rm masterclip.wav
print("vocals removed, matching the track in the database...")
with io.capture_output() as captured:
#if True:
# result = !cd audfprint && python audfprint.py match --dbase /content/ostp.pklz "/content/vocal-remover/masterclip_Instruments.wav"
# !rm "/content/vocal-remover/masterclip_Instruments.wav"
if result[3].startswith("Matched"):
name=result[3][result[3].index(" as ")+13:result[3].index(" at ")]
else:
pass
if result[3].startswith("Matched"):
print("Success!!!! the soundtrack is :")
print(name)
else:
print("sorry couldnt identify :/")
| gintamaostfinder.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (cvxpy)
# language: python
# name: cvxpy
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Квазиньютоновские методы: между двух огней
# + [markdown] slideshow={"slide_type": "slide"}
# ## Сравнительный анализ метода Ньютона и градиентного спуска
#
# Метод | Скорость сходимости | Сложность | Аффинная инвариантность | Требования к $f(x)$
# :---: | :---: | :---: | :---: | :---:
# Градиентный спуск | Глобально линейная | $O(n) + $ определение шага | Нет | Дифференцируема; градиент липшицев
# Метод Ньютона | Локально квадратичная | $O(n^3) + $ определение шага | Да | Дважды диференцируема; гессиан липшицев, положительно определён
# + [markdown] slideshow={"slide_type": "slide"}
# ## Как уменьшить сложность хранения и вычисления?
#
# - Сложность вычисления можно уменьшить с помощью
# - Квазиньютоновские методы, они же методы переменной метрики
# - Требуется хранение матрицы $n \times n$
#
# - Сложность вычисления и хранения можно уменьшить
# - квазиньютоновские методы с ограниченной памятью, например [L-BFGS](https://en.wikipedia.org/wiki/Limited-memory_BFGS) (Limited Broyden-Fletcher-Goldfarb-Shanno)
# - НЕ требуется хранить матрицу
# - вместо этого хранятся $k \ll n$ векторов из $\mathbb{R}^n$
# + [markdown] slideshow={"slide_type": "slide"}
# ## Единообразный способ получения метода Ньютона и градиентного спуска
#
# - градиентный метод получен из аппроксимации первого порядка:
#
# $$
# f_G(x) \approx f(y) + \langle f'(y), x - y \rangle + \frac{1}{2}(x-y)^{\top} \frac{1}{\alpha}I(x - y)
# $$
#
# причём при $\alpha \in (0, 1/L], f(x) \leq f_G(x)$, то есть $f_G$ - глобальная оценка $f(x)$
# - метод Ньютона получен из аппроксимации второго порядка
#
# $$
# f_N(x) \approx f(y) + \langle f'(y), x - y \rangle + \frac{1}{2} (x-y)^{\top}f''(y)(x-y)
# $$
# + [markdown] slideshow={"slide_type": "slide"}
# **Идея:** использовать промежуточную аппроксимацию вида
#
# $$
# f_q(x) \approx f(y) + \langle f'(y), x - y \rangle + \frac{1}{2} (x-y)^{\top}{\color{red}{B(y)}}(x-y),
# $$
#
# которая даёт переход к следующей точке:
#
# $$
# x_{k+1} = x_k - \alpha_k B^{-1}_k f'(x_k) = x_k - \alpha_k H_k f'(x_k)
# $$
# + [markdown] slideshow={"slide_type": "slide"}
# ## Немного истории...
# - Первый квазиньютоновский метод придумал физик <NAME> в середине 1950-х для ускорения своих вычислений на ненадёжных компьютерах
# - Его статью с описанием предложенного метода не приняли к публикации, и она оставалась техническим отчётом <br></br> более 30 лет
# - [Опубликована](http://epubs.siam.org/doi/abs/10.1137/0801001) в 1991 году в первом выпуске [SIAM Journal on Optimization](https://www.siam.org/journals/siopt.php)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Общая схема квазиньютоновских методов
#
# ```python
# def QuasiNewtonMethod(f, x0, epsilon, **kwargs):
#
# x = x0
#
# H = I
#
# while True:
#
# h = -H.dot(grad_f(x))
#
# if StopCriterion(x, f, h, **kwargs) < epsilon:
#
# break
#
# alpha = SelectStepSize(x, h, f, **kwargs)
#
# x = x + alpha * h
#
# H = UpdateH(H, f(x), grad_f(x))
#
# return x
# ```
# + [markdown] slideshow={"slide_type": "slide"}
# ## Как искать $B_{k+1}$?
#
# В точке $x_{k+1}$ имеем следующую аппрокисмацию:
#
# $$
# f_q(h) \approx f(x_{k+1}) + \langle f'(x_{k+1}), h \rangle + \frac{1}{2}h^{\top}B_{k+1}h
# $$
#
# Из определения, очевидно, что $B_{k+1} \in \mathbb{S}^n_{++}$.
# Какие требования естественно наложить на $f_q(h)$?
# + [markdown] slideshow={"slide_type": "slide"}
# $$
# f_q'(-\alpha_k h_k) = f'(x_k) \qquad f'_q(0) = f'(x_{k+1}),
# $$
#
# где первое условие даёт
#
# $$
# f'(x_{k+1}) - \alpha_k B_{k+1}h_k = f'(x_k),
# $$
#
# а второе выполняется автоматически.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Квазиньютоновское уравнение (Secant equation)
#
# Из первого условия получаем
#
# $$
# B_{k+1}s_k = y_k,
# $$
#
# где $s_k = x_{k+1} - x_k$ и $y_k = f'(x_{k+1}) - f'(x_k)$.
#
# Это уравнение будет иметь решение только при $s^{\top}_k y_k > 0$. Почему?
# + [markdown] slideshow={"slide_type": "fragment"}
# **Вопрос:** всегда ли выполнено такое соотношение
#
# между разностью градиентов и точек?
#
# **Hint**: вспомините условие Вольфа
# + [markdown] slideshow={"slide_type": "fragment"}
# **Вопрос:** единственным ли образом определено $B_{k+1}$?
# + [markdown] slideshow={"slide_type": "slide"}
# ### Как однозначно определить $B_{k+1}$?
#
# \begin{align*}
# & \min_B \| B_k - B \| \\
# \text{s.t. } & B = B^{\top}\\
# & Bs_k = y_k
# \end{align*}
# + [markdown] slideshow={"slide_type": "slide"}
# ## DFP (Davidon-Fletcher-Powell)
#
# $$
# B_{k+1} = (I - \rho_k y_k s^{\top}_k)B_k(I - \rho_k s_ky^{\top}_k) + \rho_k y_k y^{\top}_k,
# $$
#
# где $\rho_k = \dfrac{1}{y^{\top}_k s_k}$,
#
# или с помощью формулы Шермана-Морисона-Вудбери
#
# $$
# B^{-1}_{k+1} = H_{k+1} = H_k - \dfrac{H_ky_k y_k^{\top}H_k}{y^{\top}_kH_ky_k} + \dfrac{s_ks^{\top}_k}{y^{\top}_ks_k}
# $$
#
# **Вопрос:** какой ранг у разности матриц $B_{k+1} (H_{k+1})$ и $B_{k} (H_{k})$?
# + [markdown] slideshow={"slide_type": "slide"}
# ### Вывод
#
# Общая идея квазиньютоновских методов:
#
# вместо полного пересчёта гессиана на каждой итерации обновлять
#
# текущую его аппроксимацию с помощью легко вычислимого
#
# преобразования
# + [markdown] slideshow={"slide_type": "slide"}
# ## BFGS
# <img src="./bfgs.png" width=500>
#
# **Вопрос:** какая естественная модификация метода DFP?
# + [markdown] slideshow={"slide_type": "slide"}
# \begin{align*}
# & \min_H \| H_k - H \| \\
# \text{s.t. } & H = H^{\top}\\
# & Hy_k = s_k
# \end{align*}
# + [markdown] slideshow={"slide_type": "slide"}
# Формула пересчёта для метода BFGS:
#
# $$
# H_{k+1} = (I - \rho_k s_ky^{\top}_k)H_k(I - \rho_k y_k s^{\top}_k) + \rho_k s_k s^{\top}_k,
# $$
#
# где $\rho_k = \dfrac{1}{y^{\top}_k s_k}$
# + [markdown] slideshow={"slide_type": "slide"}
# ### Детали реализации
#
# - Не должно быть операций сложностью $O(n^3)$, то есть никаких матричных умножений и решений линейных систем (cf. [реализацию в SciPy](https://github.com/scipy/scipy/blob/v0.18.1/scipy/optimize/optimize.py#L874-L976))
# - Только правило Вольфа гарантирует соблюдения условия кривизны $y_k^{\top}s_k > 0$
# - Параметры в правиле Вольфа обычно следующие
# - $\alpha_0 = 1$ необходим для сверхлинейной скорости
# - $\beta_1 = 10^{-4}$, $\beta_2 = 0.9$
# - Способы инициализации $H_0$
# - единичная матрица
# - $H_0 = \frac{y_0^{\top}s_0}{y_0^{\top}y_0}I$ **после** первого шага, но до вычисления $H_1$.При вычислении $x_1$ используется $H_0 = I$
# - $H_0 = \delta \|g_0\|^{-1}_2 I$, параметр $\delta$ необходимо заранее задать
# - При использовании $B$ вместо $H$ нужно хранить $B$ в виде $LDL^{\top}$ разложения и обновлять не саму матрицу $B$, а её разложение. Это явно делается за $O(n^2)$. Вычисление $h_k$ - это решение линейной системы с предвычисленным раздложением матрицы, следовательно сложность также $O(n^2)$. Этот подход позволяет контролировать устройчивость в диагонали матрицы $D$. На практике предпочтительнее работать с матрицей $H$
#
# + [markdown] slideshow={"slide_type": "slide"}
# ### Сходимость
#
# **Теорема**
#
# Пусть $f$ дважды непрерывно дифференцируема и её гесс<NAME>, также пусть последовательность генерируемая методом BFGS сходится к точке $x^*$ так что $\sum_{k=1}^{\infty} \|x_k - x^*\| < \infty$. Тогда $x_k \to x^*$ сверхлинейно.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Самокоррекция
#
# - Если BFGS на некоторой итерации даёт плохую оценку обратного гессиана, то через несколько итераций это недоразумение будет **автоматически** исправлено, то есть метод сам скорректирует грубую оценку гессиана
# - Это свойство появляется только при правильном способе выбора шага, например при использовании правила Вольфа
# - Метод DFP существенно хуже корректирует неточные оценки обратного гессиана
# - Всё это будет ниже проиллюстрировано на примерах
# + [markdown] slideshow={"slide_type": "slide"}
# ## BFGS с ограниченной памятью (L-BFGS)
#
# - В методе BFGS нужна не сама матрица $H$, а только функция умножения её на вектор
# - Поскольку требуется локальная оценка гессиана, старые значения векторов $s$ и $y$ могут портить текущую оценку
#
# **Идея**
#
# - Хранить $k \ll n$ последних векторов $s$ и $y$ - снижение требуемой памяти с $n^2$ до $kn$
# - Выполнение умножения на вектор рекурсивно, без явного формирования матрицы $H$
# + [markdown] slideshow={"slide_type": "slide"}
# ### Сравнение с нелинейным методом сопряжённых градиентов
#
# - В методе Хестенса-Штифеля
#
# $$
# h_{k+1} = -f'(x_{k+1}) + \beta_{k+1} h_{k}, \quad \beta_{k+1} = \frac{y_k^{\top}f'(x_{k+1})}{y_k^{\top} h_k}
# $$
#
# или
#
# $$
# h_{k+1} = -\left(I - \frac{s_k y_k^{\top}}{y_k^{\top}s_k}\right)f'(x_{k+1}) = -\hat{H}_{k+1} f'(x_{k+1})
# $$
#
# - Матрица $\hat{H}_{k+1}$ несимметрична и неположительно определённая, однако матрица
#
# $$
# H_{k+1} = \left(I - \frac{s_k y_k^{\top}}{y_k^{\top}s_k}\right)\left(I - \frac{y_k s_k^{\top}}{y_k^{\top}s_k}\right) + \frac{s_ks_k^{\top}}{y_k^{\top}s_k}
# $$
#
# удовлетворяет всем требованиям к матрице в методе BFGS и совпадает с формулой для обновления $H_k$, если $H_k = I$, то есть $k=1$ в методе LBFGS и $H_0 = I$
# - Более того, при выборе шага по правилу наискорейшего спуска, формулы для метода Хестенса Штифеля и LBFGS с $k = 1$ в точности совпадают
# + [markdown] slideshow={"slide_type": "slide"}
# ## Barzilai-Borwein method
#
# - Первая [статья](http://pages.cs.wisc.edu/~swright/726/handouts/barzilai-borwein.pdf) об этом методе опубликована в 1988, в журнале IMA Journal of Numerical Analysis
# - [Статья](http://papers.nips.cc/paper/6286-barzilai-borwein-step-size-for-stochastic-gradient-descent.pdf) на NIPS 2016 о модификации этого метода в случае использования стохастической оценки градиента
# - Идея: комбинация идеи наискорейшего спуска и квазиньютоновского метода
# + [markdown] slideshow={"slide_type": "slide"}
# ### Идея метода
#
# - Наискорейший спуск: $x_{k+1} = x_k - \alpha_k f'(x_k)$, $\alpha_k = \arg \min\limits_{\alpha > 0} f(x_{k+1})$
# - Метод Ньютона: $x_{k+1} = x_k - (f''(x_k))^{-1} f'(x_k)$
# - Аппроксимация гессиана диагональной матрицей:
#
# $$
# \alpha_k f'(x_k) = \alpha_k I f'(x_k) = \left( \frac{1}{\alpha_k} I \right)^{-1} f'(x_k) \approx f''(x_k))^{-1} f'(x_k)
# $$
#
# - Как найти $\alpha_k$?
# + [markdown] slideshow={"slide_type": "slide"}
# ### Снова квазиньютоновское уравнение (Secant equation)
# - Для точного гессиана
# $$
# f''(x_{k})(x_{k} - x_{k-1}) = f'(x_{k}) - f'(x_{k-1})
# $$
# - Для приближения
#
# $$
# \alpha_k^{-1} s_{k-1} \approx y_{k-1}
# $$
#
# - Задача аппроксимации одного вектора с помощью масштабирования другого
# - Простейший квазиньютоновский метод вырождается в поиск оптимального шага
# + [markdown] slideshow={"slide_type": "slide"}
# ### Три способа найти $\alpha_k$
#
# - Первый способ
# - Задача
#
# $$
# \min_{\beta} \|\beta s_{k-1} - y_{k-1} \|^2_2
# $$
#
# - Решение
#
# $$
# \alpha = \frac{1}{\beta} = \frac{s^{\top}_{k-1} s_{k-1}}{s^{\top}_{k-1} y_{k-1}}
# $$
#
# - Второй способ
# - Задача
#
# $$
# \min_{\alpha} \| s_{k-1} - \alpha y_{k-1} \|^2_2
# $$
#
# - Решение
#
# $$
# \alpha = \frac{s^{\top}_{k-1} y_{k-1}}{y^{\top}_{k-1} y_{k-1}}
# $$
# - Третий способ называется немонотонный линейный поиск: специальная модификация правил Армихо, учитывающая историю изменений значения функции, [статья](https://www.math.lsu.edu/~hozhang/papers/nonmonotone.pdf) 2004 г. в SIAM Journal on Optimization
# + [markdown] slideshow={"slide_type": "slide"}
# ## Эксперименты
#
# ### Поиск аналитического центра системы неравенств
#
# $$
# f(x) = - \sum_{i=1}^m \log(1 - a_i^{\top}x) - \sum\limits_{i = 1}^n \log (1 - x^2_i) \to \min_x
# $$
# + slideshow={"slide_type": "slide"}
import numpy as np
import liboptpy.unconstr_solvers as methods
import liboptpy.step_size as ss
# %matplotlib inline
import matplotlib.pyplot as plt
import scipy.optimize as scopt
plt.rc("text", usetex=True)
# + slideshow={"slide_type": "slide"}
n = 3000
m = 100
x0 = np.zeros(n)
max_iter = 100
tol = 1e-5
A = np.random.rand(m, n) * 10
# + slideshow={"slide_type": "slide"}
f = lambda x: -np.sum(np.log(1 - A.dot(x))) - np.sum(np.log(1 - x*x))
grad_f = lambda x: np.sum(A.T / (1 - A.dot(x)), axis=1) + 2 * x / (1 - np.power(x, 2))
# + slideshow={"slide_type": "slide"}
def bb_method(f, gradf, x0, tol=1e-6, maxiter=100, callback=None, alpha_type=1):
it = 0
x_prev = x0.copy()
current_tol = np.linalg.norm(gradf(x_prev))
alpha = 1e-4
while current_tol > tol and it < maxiter:
it += 1
current_grad = gradf(x_prev)
if it != 1:
g = current_grad - prev_grad
if alpha_type == 1:
alpha = g.dot(s) / g.dot(g)
elif alpha_type == 2:
alpha = s.dot(s) / g.dot(s)
if callback:
callback(x_prev)
x_next = x_prev - alpha * current_grad
current_tol = np.linalg.norm(gradf(x_next))
prev_grad = current_grad
s = x_next - x_prev
x_prev = x_next
if callback:
callback(x_prev)
return x_next
# + slideshow={"slide_type": "slide"}
method = {
"BB 1": methods.fo.BarzilaiBorweinMethod(f, grad_f, init_alpha=1e-4, type=1),
"BFGS": methods.fo.BFGS(f, grad_f),
"DFP": methods.fo.DFP(f, grad_f),
"LBFGS": methods.fo.LBFGS(f, grad_f),
}
# + slideshow={"slide_type": "slide"}
for m in method:
print("\t Method {}".format(m))
_ = method[m].solve(x0=x0, tol=tol, max_iter=max_iter, disp=True)
print("\t Method BFGS Scipy")
scopt_conv = []
scopt_res = scopt.minimize(f, x0, method="BFGS", jac=grad_f, callback=lambda x: scopt_conv.append(x),
tol=tol, options={"maxiter": max_iter})
print("Result: {}".format(scopt_res.message))
if scopt_res.success:
print("Convergence in {} iterations".format(scopt_res.nit))
print("Function value = {}".format(f(scopt_res.x)))
# + slideshow={"slide_type": "slide"}
plt.figure(figsize=(8, 6))
for m in method:
plt.semilogy([np.linalg.norm(grad_f(x)) for x in method[m].get_convergence()], label=m)
plt.semilogy([np.linalg.norm(grad_f(x)) for x in [x0] + scopt_conv], label="BFGS SciPy")
plt.ylabel("$\|f'(x_k)\|_2$", fontsize=18)
plt.xlabel("Number of iterations, $k$", fontsize=18)
plt.legend(fontsize=18)
plt.xticks(fontsize=18)
_ = plt.yticks(fontsize=18)
# + slideshow={"slide_type": "slide"}
for m in method:
print("\t Method {}".format(m))
# %timeit method[m].solve(x0=x0, tol=tol, max_iter=max_iter)
# %timeit scopt.minimize(f, x0, method="BFGS", jac=grad_f, tol=tol, options={"maxiter": max_iter})
# + [markdown] slideshow={"slide_type": "slide"}
# ### Плохо обусловленная задача
# + slideshow={"slide_type": "slide"}
n = 50
D = np.arange(1, n+1)
U = np.random.randn(n, n)
U, _ = np.linalg.qr(U)
A = U.dot(np.diag(D)).dot(U.T)
b = np.random.randn(n)
eig_vals = np.linalg.eigvals(A)
print("Condition number = {}".format(np.max(eig_vals) / np.min(eig_vals)))
# + slideshow={"slide_type": "slide"}
f = lambda x: 0.5 * x.T.dot(A.dot(x)) - b.dot(x)
gradf = lambda x: A.dot(x) - b
x0 = np.random.randn(n)
# + slideshow={"slide_type": "slide"}
method = {
"BB 1": methods.fo.BarzilaiBorweinMethod(f, gradf, init_alpha=1e-4, type=1),
"BB 2": methods.fo.BarzilaiBorweinMethod(f, gradf, init_alpha=1e-4, type=2),
"BFGS": methods.fo.BFGS(f, gradf),
"DFP": methods.fo.DFP(f, gradf),
"GD": methods.fo.GradientDescent(f, gradf, ss.ExactLineSearch4Quad(A, b)),
"LBFGS": methods.fo.LBFGS(f, gradf, hist_size=10),
}
# + slideshow={"slide_type": "slide"}
for m in method:
print("\t Method {}".format(m))
_ = method[m].solve(x0=x0, tol=tol, max_iter=max_iter, disp=True)
print("\t Method BFGS Scipy")
scopt_conv = []
scopt_res = scopt.minimize(f, x0, method="BFGS", jac=gradf, callback=lambda x: scopt_conv.append(x),
tol=tol, options={"maxiter": max_iter})
print("Result: {}".format(scopt_res.message))
if scopt_res.success:
print("Convergence in {} iterations".format(scopt_res.nit))
print("Function value = {}".format(f(scopt_res.x)))
# + slideshow={"slide_type": "slide"}
plt.figure(figsize=(12, 8))
fontsize = 26
for m in method:
plt.semilogy([np.linalg.norm(gradf(x)) for x in method[m].get_convergence()], label=m)
plt.semilogy([np.linalg.norm(gradf(x)) for x in [x0] + scopt_conv], label='BFGS SciPy')
plt.legend(fontsize=fontsize)
plt.ylabel("$\|f'(x_k)\|_2$", fontsize=fontsize)
plt.xlabel("Number of iterations, $k$", fontsize=fontsize)
plt.xticks(fontsize=fontsize)
_ = plt.yticks(fontsize=fontsize)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Pro & Contra
#
# Pro:
# 1. Вместо **точного** вычисления гессиана используется его **оценка**, полученная с помощью градиента и оценки гессиана в предыдущей точке
# 2. Вместо решения систем линейных уравнений используется текущаю информация о функции и градиенте для аналитического вычисления приближения обращённого гессиана
# 3. Сложность одной итерации $O(n^2) + ...$ по сравнению с $O(n^3) + ...$ в методе Ньютона
# 4. Для метода L-BFGS требуется линейное количество памяти по размерности задачи
# 5. Свойство самокоррекции метода BFGS: если на некоторой итерации обратный гессиан оценен очень грубо, то следующие несколько итераций улучшат оценку
# 6. Сверхлинейная сходимость к решению задачи минимизации $f$ (подробнее см. [[1]](http://www.bioinfo.org.cn/~wangchao/maa/Numerical_Optimization.pdf))
#
# Contra:
# 1. Нет универсального рецепта выбора начального приближения $B_0$ или $H_0$
# 2. Нет разработанной теории сходимости и оптимальности
# 3. Не любое условие на линейный поиск шага гарантирует выполнения условия кривизны $y^{\top}_ks_k > 0$
| 09-Newton/Seminar9b.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + run_control={"frozen": false, "read_only": false}
import math
import warnings
from IPython.display import display
from matplotlib import pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
from sklearn import linear_model
import statsmodels.formula.api as smf
# Display preferences.
# %matplotlib inline
pd.options.display.float_format = '{:.3f}'.format
# Suppress annoying harmless error.
warnings.filterwarnings(
action="ignore",
module="scipy",
message="^internal gelsd"
)
# + [markdown] run_control={"frozen": false, "read_only": false}
# ## The Extraordinary Power of Explanatory Power
#
# The strength of multiple linear regression lies in its ability to provide straightforward and interpretable solutions that not only predict future outcomes, but also provide insight into the underlying processes that create these outcomes. For example, after fitting the following model:
#
# $$HourlyWidgetProduction = \alpha + \beta_1WorkerAgeFrom18+ \beta_2WorkerYearsinJob + \beta_3IsRoundWidget$$
#
# we get these parameters:
# $$\alpha = 2$$
# $$\beta_1 = .1$$
# $$\beta_2 = .2$$
# $$\beta_3 = 4$$
#
# Using those parameters, we learn that round widgets are twice as fast to produce as non-round widgets. We can tell because $\alpha$ represents the intercept, the hourly rate of production for widgets that are not round (2 an hour) and $\beta_3$ represents the difference between the intercept and the hourly rate of production for round widgets (also 2 an hour, for a total of 4 round widgets an hour).
#
# We also learn that for every year a worker ages after the age of 18, their hourly production-rate goes up by .1 ($\beta_1$). In addition, for every year a worker has been in that job, their hourly production-rate goes up by .2 ($\beta_2$).
#
# Furthermore, using this model, we can predict that a 20-year-old worker who has been in the job for a year and is making only round widgets will make $2 + .1*2 + .2*1 + 4 = 6.3$ round widgets an hour.
#
# Finally, and probably of greatest interest, we get an **R-Squared** value. This is a proportion (between 0 and 1) that expresses how much variance in the outcome variable our model was able to explain. Higher $R^2$ values are better to a point-- a low $R^2$ indicates that our model isn't explaining much information about the outcome, which means it will not give very good predictions. However, a very high $R^2$ is a warning sign for overfitting. No dataset is a perfect representation of reality, so a model that perfectly fits our data ($R^2$ of 1 or close to 1) is likely to be biased by quirks in the data, and will perform less well on the test-set.
#
# Here's an example using a toy advertising dataset:
#
# + run_control={"frozen": false, "read_only": false}
# Acquire, load, and preview the data.
data = pd.read_csv('https://tf-curricula-prod.s3.amazonaws.com/data-science/Advertising.csv')
display(data.head())
# Instantiate and fit our model.
regr = linear_model.LinearRegression()
Y = data['Sales'].values.reshape(-1, 1)
X = data[['TV','Radio','Newspaper']]
regr.fit(X, Y)
# Inspect the results.
print('\nCoefficients: \n', regr.coef_)
print('\nIntercept: \n', regr.intercept_)
print('\nR-squared:')
print(regr.score(X, Y))
# + [markdown] run_control={"frozen": false, "read_only": false}
# The model where the outcome Sales is predicted by the features TV, Radio, and Newspaper explains 89.7% of the variance in Sales. Note that we don't know from these results how much of that variance is explained by each of the three features. Looking at the coefficients, there appears to be a base rate of Sales that happen even with no ads in any medium (intercept: 2.939) and sales have the highest per-unit increase when ads are on the radio (0.189).
# + [markdown] run_control={"frozen": false, "read_only": false}
# ## Assumptions of Multivariable Linear Regression
#
# For regression to work its magic, inputs to the model need to be consistent with four assumptions:
#
#
# ### Assumption one: linear relationship
#
# As mentioned earlier, features in a regression need to have a linear relationship with the outcome. If the relationship is non-linear, the regression model will try to find any hint of a linear relationship, and only explain that – with predictable consequences for the validity of the model.
#
# Sometimes this can be fixed by applying a non-linear transformation function to a feature. For example, if the relationship between feature and outcome is quadratic and all feature scores are > 0, we can take the square root of the features, resulting in a linear relationship between the outcome and sqrt(feature).
#
# + run_control={"frozen": false, "read_only": false}
# Sample data.
outcome = [1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12]
feature = [3, 4, 10, 16, 25, 33, 49, 60, 85, 100, 130, 140]
# Plot the data as-is. Looks a mite quadratic.
plt.scatter(outcome, feature)
plt.title('Raw values')
plt.show()
# Create a feature using a non-linear transformation.
sqrt_feature = [math.sqrt(x) for x in feature]
# Well now isn't that nice.
plt.scatter(outcome, sqrt_feature)
plt.title('Transformed values')
plt.show()
# + [markdown] run_control={"frozen": false, "read_only": false}
# When interpreting features with non-linear transformations, it is important to keep the transformation in mind. For example, in the equation $y = 2log({x})$, y increases by one unit for every two-unit increase in $log({x})$. The relationship between y and x, however, is non-linear, and the amount of change in y varies based on the absolute value of x:
#
# |x |log(x)| y|
# |--|--|--|
# |1 |0 |0|
# |10 |1 |2|
# |100 |2 |4|
# |1000| 3 |6|
#
# So a one-unit change in x from 1 to 2 will result in a much greater change in y than a one-unit change in x from 100 to 101.
#
# There are many variable transformations. For a deep dive, check out the Variable Linearization section of [Fifty Ways to Fix Your Data](https://statswithcats.wordpress.com/2010/11/21/fifty-ways-to-fix-your-data/).
#
# ### Assumption two: multivariate normality
#
# The error from the model (calculated by subtracting the model-predicted values from the real outcome values) should be normally distributed. Since ordinary least squares regression models are fitted by choosing the parameters that best minimize error, skewness or outliers in the error can result in serious miss-estimations.
#
# Outliers or skewness in error can often be traced back to outliers or skewness in data.
# + run_control={"frozen": false, "read_only": false}
# Extract predicted values.
predicted = regr.predict(X).ravel()
actual = data['Sales']
# Calculate the error, also called the residual.
residual = actual - predicted
# This looks a bit concerning.
plt.hist(residual)
plt.title('Residual counts')
plt.xlabel('Residual')
plt.ylabel('Count')
plt.show()
# + [markdown] run_control={"frozen": false, "read_only": false}
#
# ### Assumption three: homoscedasticity
#
# The distribution of your error terms (its "scedasticity"), should be consistent for all predicted values, or **homoscedastic**.
#
# For example, if your error terms aren't consistently distributed and you have more variance in the error for large outcome values than for small ones, then the confidence interval for large predicted values will be too small because it will be based on the average error variance. This leads to overconfidence in the accuracy of your model's predictions.
#
# Some fixes to heteroscedasticity include transforming the dependent variable and adding features that target the poorly-estimated areas. For example, if a model tracks data over time and model error variance jumps in the September to November period, a binary feature indicating season may be enough to resolve the problem.
# + run_control={"frozen": false, "read_only": false}
plt.scatter(predicted, residual)
plt.xlabel('Predicted')
plt.ylabel('Residual')
plt.axhline(y=0)
plt.title('Residual vs. Predicted')
plt.show()
# Hm... looks a bit concerning.
# + [markdown] run_control={"frozen": false, "read_only": false}
# ### Assumption four: low multicollinearity
#
# Correlations among features should be low or nonexistent. When features are correlated, they may both explain the same pattern of variance in the outcome. The model will attempt to find a solution, potentially by attributing half the explanatory power to one feature and half to the other. This isn’t a problem if our only goal is prediction, because then all that matters is that the variance gets explained. However, if we want to know which features matter most when predicting an outcome, multicollinearity can cause us to underestimate the relationship between features and outcomes.
#
# Multicollinearity can be fixed by PCA or by discarding some of the correlated features.
# + run_control={"frozen": false, "read_only": false}
correlation_matrix = X.corr()
display(correlation_matrix)
# -
# ## Drill: fixing assumptions
#
# Judging from the diagnostic plots, your data has a problem with both heteroscedasticity and multivariate non-normality. Use the cell(s) below to see what you can do to fix it.
# + run_control={"frozen": false, "read_only": false}
f, axs = plt.subplots(3, 2, sharex=False)
f.set_size_inches(20,15)
# Your code here.
for i,col in enumerate(['TV','Radio','Newspaper']):
sns.regplot(x=data[col],y=Y.ravel(), color="g", ax=axs[i][0])
axs[i][1].hist(data[col])
# +
from scipy.stats import boxcox
data["Newspaper_mod"],new_coef = boxcox(data.Newspaper+0.1)
for i,col in enumerate(['TV','Radio','Newspaper']):
data[col+"_mod"] = boxcox(data[col]+0.1)[0]
f, axs = plt.subplots(3, 2, sharex=False)
f.set_size_inches(20,15)
# Your code here.
for i,col in enumerate(['TV','Radio','Newspaper']):
sns.regplot(x=data[col],y=Y.ravel(), color="g", ax=axs[i][0])
axs[i][1].hist(data[col+"_mod"])
# +
#data["TV_mod"] = boxcox(data.TV)[0]
Y = data['Sales'].values.reshape(-1, 1)
X = data[['TV_mod','Radio_mod','Newspaper_mod']]
regr.fit(X, Y)
# Inspect the results.
print('\nCoefficients: \n', regr.coef_)
print('\nIntercept: \n', regr.intercept_)
print('\nR-squared:')
print(regr.score(X, Y))
predicted = regr.predict(X).ravel()
actual = data['Sales']
# Calculate the error, also called the residual.
residual = actual - predicted
plt.scatter(predicted, residual)
plt.xlabel('Predicted')
plt.ylabel('Residual')
plt.axhline(y=0)
plt.title('Residual vs. Predicted')
plt.show()
# +
data['Sales_log'] = np.log10(data.Sales)
data['TV_log'] = np.log(data.TV)
data['Radio_log'] = np.log(data.Radio+1)
data['Newspaper_log'] = np.log(data.Newspaper+1)
data["predicted"] = predicted
data["residual"] = residual
# +
#data["TV_mod"] = boxcox(data.TV)[0]
Y = data['Sales_log'].values.reshape(-1, 1)
X = data[['TV','Radio','Newspaper_mod']]
regr.fit(X, Y)
# Inspect the results.
print('\nCoefficients: \n', regr.coef_)
print('\nIntercept: \n', regr.intercept_)
print('\nR-squared:')
print(regr.score(X, Y))
predicted = regr.predict(X).ravel()
actual =data['Sales_log']
# Calculate the error, also called the residual.
residual = actual - predicted
plt.scatter(predicted, residual)
plt.xlabel('Predicted')
plt.ylabel('Residual')
plt.axhline(y=0)
plt.title('Residual vs. Predicted')
plt.show()
# +
from sklearn.model_selection import cross_val_score
cross_val_score(regr, X, Y, cv=5)
| 2.4.3+The+Extraordinary+Power+of+Explanatory+Power.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="ItXfxkxvosLH"
# # 使用 Keras 和 Tensorflow Hub 对电影评论进行文本分类
# + [markdown] colab_type="text" id="Eg62Pmz3o83v"
#
# 此笔记本(notebook)使用评论文本将影评分为*积极(positive)*或*消极(nagetive)*两类。这是一个*二元(binary)*或者二分类问题,一种重要且应用广泛的机器学习问题。
#
# 本教程演示了使用 Tensorflow Hub 和 Keras 进行迁移学习的基本应用。
#
# 我们将使用来源于[网络电影数据库(Internet Movie Database)](https://www.imdb.com/)的 [IMDB 数据集(IMDB dataset)](https://www.tensorflow.org/api_docs/python/tf/keras/datasets/imdb),其包含 50,000 条影评文本。从该数据集切割出的 25,000 条评论用作训练,另外 25,000 条用作测试。训练集与测试集是*平衡的(balanced)*,意味着它们包含相等数量的积极和消极评论。
#
# 此笔记本(notebook)使用了 [tf.keras](https://www.tensorflow.org/guide/keras),它是一个 Tensorflow 中用于构建和训练模型的高级API,此外还使用了 [TensorFlow Hub](https://www.tensorflow.org/hub),一个用于迁移学习的库和平台。有关使用 `tf.keras` 进行文本分类的更高级教程,请参阅 [MLCC文本分类指南(MLCC Text Classification Guide)](https://developers.google.com/machine-learning/guides/text-classification/)。
# + colab={} colab_type="code" id="2ew7HTbPpCJH"
from __future__ import absolute_import, division, print_function, unicode_literals
import numpy as np
try:
# Colab only
# %tensorflow_version 2.x
except Exception:
pass
import tensorflow as tf
import tensorflow_hub as hub
import tensorflow_datasets as tfds
print("Version: ", tf.__version__)
print("Eager mode: ", tf.executing_eagerly())
print("Hub version: ", hub.__version__)
print("GPU is", "available" if tf.config.experimental.list_physical_devices("GPU") else "NOT AVAILABLE")
# + [markdown] colab_type="text" id="iAsKG535pHep"
# ## 下载 IMDB 数据集
# IMDB数据集可以在 [Tensorflow 数据集](https://github.com/tensorflow/datasets)处获取。以下代码将 IMDB 数据集下载至您的机器(或 colab 运行时环境)中:
# + colab={} colab_type="code" id="zXXx5Oc3pOmN"
# 将训练集按照 6:4 的比例进行切割,从而最终我们将得到 15,000
# 个训练样本, 10,000 个验证样本以及 25,000 个测试样本
train_validation_split = tfds.Split.TRAIN.subsplit([6, 4])
(train_data, validation_data), test_data = tfds.load(
name="imdb_reviews",
split=(train_validation_split, tfds.Split.TEST),
as_supervised=True)
# + [markdown] colab_type="text" id="l50X3GfjpU4r"
# ## 探索数据
#
# 让我们花一点时间来了解数据的格式。每一个样本都是一个表示电影评论和相应标签的句子。该句子不以任何方式进行预处理。标签是一个值为 0 或 1 的整数,其中 0 代表消极评论,1 代表积极评论。
#
# 我们来打印下前十个样本。
# + colab={} colab_type="code" id="QtTS4kpEpjbi"
train_examples_batch, train_labels_batch = next(iter(train_data.batch(10)))
train_examples_batch
# + [markdown] colab_type="text" id="IFtaCHTdc-GY"
# 我们再打印下前十个标签。
# + colab={} colab_type="code" id="tvAjVXOWc6Mj"
train_labels_batch
# + [markdown] colab_type="text" id="LLC02j2g-llC"
# ## 构建模型
#
# 神经网络由堆叠的层来构建,这需要从三个主要方面来进行体系结构决策:
#
# * 如何表示文本?
# * 模型里有多少层?
# * 每个层里有多少*隐层单元(hidden units)*?
#
# 本示例中,输入数据由句子组成。预测的标签为 0 或 1。
#
# 表示文本的一种方式是将句子转换为嵌入向量(embeddings vectors)。我们可以使用一个预先训练好的文本嵌入(text embedding)作为首层,这将具有三个优点:
#
# * 我们不必担心文本预处理
# * 我们可以从迁移学习中受益
# * 嵌入具有固定长度,更易于处理
#
# 针对此示例我们将使用 [TensorFlow Hub](https://www.tensorflow.org/hub) 中名为 [google/tf2-preview/gnews-swivel-20dim/1](https://tfhub.dev/google/tf2-preview/gnews-swivel-20dim/1) 的一种**预训练文本嵌入(text embedding)模型** 。
#
# 为了达到本教程的目的还有其他三种预训练模型可供测试:
#
# * [google/tf2-preview/gnews-swivel-20dim-with-oov/1](https://tfhub.dev/google/tf2-preview/gnews-swivel-20dim-with-oov/1) ——类似 [google/tf2-preview/gnews-swivel-20dim/1](https://tfhub.dev/google/tf2-preview/gnews-swivel-20dim/1),但 2.5%的词汇转换为未登录词桶(OOV buckets)。如果任务的词汇与模型的词汇没有完全重叠,这将会有所帮助。
# * [google/tf2-preview/nnlm-en-dim50/1](https://tfhub.dev/google/tf2-preview/nnlm-en-dim50/1) ——一个拥有约 1M 词汇量且维度为 50 的更大的模型。
# * [google/tf2-preview/nnlm-en-dim128/1](https://tfhub.dev/google/tf2-preview/nnlm-en-dim128/1) ——拥有约 1M 词汇量且维度为128的更大的模型。
# + [markdown] colab_type="text" id="In2nDpTLkgKa"
# 让我们首先创建一个使用 Tensorflow Hub 模型嵌入(embed)语句的Keras层,并在几个输入样本中进行尝试。请注意无论输入文本的长度如何,嵌入(embeddings)输出的形状都是:`(num_examples, embedding_dimension)`。
#
# + colab={} colab_type="code" id="_NUbzVeYkgcO"
embedding = "https://tfhub.dev/google/tf2-preview/gnews-swivel-20dim/1"
hub_layer = hub.KerasLayer(embedding, input_shape=[],
dtype=tf.string, trainable=True)
hub_layer(train_examples_batch[:3])
# + [markdown] colab_type="text" id="dfSbV6igl1EH"
# 现在让我们构建完整模型:
# + colab={} colab_type="code" id="xpKOoWgu-llD"
model = tf.keras.Sequential()
model.add(hub_layer)
model.add(tf.keras.layers.Dense(16, activation='relu'))
model.add(tf.keras.layers.Dense(1, activation='sigmoid'))
model.summary()
# + [markdown] colab_type="text" id="6PbKQ6mucuKL"
# 层按顺序堆叠以构建分类器:
#
# 1. 第一层是 Tensorflow Hub 层。这一层使用一个预训练的保存好的模型来将句子映射为嵌入向量(embedding vector)。我们所使用的预训练文本嵌入(embedding)模型([google/tf2-preview/gnews-swivel-20dim/1](https://tfhub.dev/google/tf2-preview/gnews-swivel-20dim/1))将句子切割为符号,嵌入(embed)每个符号然后进行合并。最终得到的维度是:`(num_examples, embedding_dimension)`。
# 2. 该定长输出向量通过一个有 16 个隐层单元的全连接层(`Dense`)进行管道传输。
# 3. 最后一层与单个输出结点紧密相连。使用 `Sigmoid` 激活函数,其函数值为介于 0 与 1 之间的浮点数,表示概率或置信水平。
#
# 让我们编译模型。
# + [markdown] colab_type="text" id="L4EqVWg4-llM"
# ### 损失函数与优化器
#
# 一个模型需要损失函数和优化器来进行训练。由于这是一个二分类问题且模型输出概率值(一个使用 sigmoid 激活函数的单一单元层),我们将使用 `binary_crossentropy` 损失函数。
#
# 这不是损失函数的唯一选择,例如,您可以选择 `mean_squared_error` 。但是,一般来说 `binary_crossentropy` 更适合处理概率——它能够度量概率分布之间的“距离”,或者在我们的示例中,指的是度量 ground-truth 分布与预测值之间的“距离”。
#
# 稍后,当我们研究回归问题(例如,预测房价)时,我们将介绍如何使用另一种叫做均方误差的损失函数。
#
# 现在,配置模型来使用优化器和损失函数:
# + colab={} colab_type="code" id="Mr0GP-cQ-llN"
model.compile(optimizer='adam',
loss='binary_crossentropy',
metrics=['accuracy'])
# + [markdown] colab_type="text" id="35jv_fzP-llU"
# ## 训练模型
#
# 以 512 个样本的 mini-batch 大小迭代 20 个 epoch 来训练模型。 这是指对 `x_train` 和 `y_train` 张量中所有样本的的 20 次迭代。在训练过程中,监测来自验证集的 10,000 个样本上的损失值(loss)和准确率(accuracy):
# + colab={} colab_type="code" id="tXSGrjWZ-llW"
history = model.fit(train_data.shuffle(10000).batch(512),
epochs=20,
validation_data=validation_data.batch(512),
verbose=1)
# + [markdown] colab_type="text" id="9EEGuDVuzb5r"
# ## 评估模型
#
# 我们来看下模型的表现如何。将返回两个值。损失值(loss)(一个表示误差的数字,值越低越好)与准确率(accuracy)。
# + colab={} colab_type="code" id="zOMKywn4zReN"
results = model.evaluate(test_data.batch(512), verbose=2)
for name, value in zip(model.metrics_names, results):
print("%s: %.3f" % (name, value))
# + [markdown] colab_type="text" id="z1iEXVTR0Z2t"
# 这种十分朴素的方法得到了约 87% 的准确率(accuracy)。若采用更好的方法,模型的准确率应当接近 95%。
# + [markdown] colab_type="text" id="5KggXVeL-llZ"
# ## 进一步阅读
#
# 有关使用字符串输入的更一般方法,以及对训练期间准确率(accuracy)和损失值(loss)更详细的分析,请参阅[此处](https://www.tensorflow.org/tutorials/keras/basic_text_classification)。
# -
| chapter2/2.3.1-text_classification_with_hub.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Dhaxy/OOP58002/blob/main/FUNDAMENTALS_OF_PYTHON.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="FbqDXyiVjNBQ"
# Python Variables
# + colab={"base_uri": "https://localhost:8080/"} id="Ri6cpT9AjR3u" outputId="50125ca4-3fd8-40c6-c017-db7d5e09af09"
x = float(1)
a, b = 0, -1
a, b, c = "nik", "pau", "np"
print('This a sample')
print(a)
print(c)
# + [markdown] id="0JhCjy0rjjjW"
# Casting
# + colab={"base_uri": "https://localhost:8080/"} id="8rl-UEY4jmmh" outputId="9e295e00-3d1a-4166-9bb1-814df4536fc4"
print (x)
# + [markdown] id="LXp2SvwHjrJT"
# Type() Function
# + colab={"base_uri": "https://localhost:8080/"} id="e3iWE9rMjud9" outputId="f2c40068-6925-4d6f-c4db-231832db9737"
y = "<NAME>"
print(type(y))
print(type(x))
# + [markdown] id="DjvazLXnj64t"
# Double quotes and Single quotes
# + colab={"base_uri": "https://localhost:8080/"} id="XVuRK8zyj9Cm" outputId="00650be5-f1e0-4944-d987-8928fc9f1e2e"
h= "Bopis"
v= 1
V= 2
print(h)
print(v)
print(v+1)
# + [markdown] id="HcNprdgIkKCz"
# Multiple Variables
# + colab={"base_uri": "https://localhost:8080/"} id="AQpu8wWckOkj" outputId="b457345b-e0da-4f3a-cf69-388652b99229"
x,y,z="i", "miss", "you"
print(x)
print(y)
print(z)
print(x,y,z)
# + [markdown] id="JH7q_poPkYay"
# One Value to Multiple Variables
# + colab={"base_uri": "https://localhost:8080/"} id="0mX2CXVrkZ7h" outputId="a0a4cf44-b4b8-4ac2-c1d1-36b0a595dde9"
x = y = z ="dhax"
print(x,y,z)
# + [markdown] id="sQmqXY9ykeka"
# Output Variables
# + colab={"base_uri": "https://localhost:8080/"} id="M31QUredkg2P" outputId="7cbea085-81cd-4234-b55b-5a1f9085ff70"
x= "singing"
print("I am " + x)
x = "Adamson"
y = "Hymn"
print(x+""+" "+y)
# + [markdown] id="9VmMIMRDk0BW"
# Arithmetic Operations
# + colab={"base_uri": "https://localhost:8080/"} id="Baghvf0qk1Xl" outputId="eae52e0a-9dc2-49cb-cc02-5763c2970e40"
f = 3
g = 6
i = 9
print(f+g)
print(f-g)
print(f*i)
print(int(i/g))
print(3/g)
print(3%g)
print(3//g)
print(3**6)
# + [markdown] id="B7b3ryublKqu"
# Assignment Operators
# + colab={"base_uri": "https://localhost:8080/"} id="85qM8AoElM0T" outputId="d8346185-ff00-4713-c928-51e48035da27"
k = 2
l=3
k+=3 #same as k=k+3
print(k)
print(l>>1)
# + [markdown] id="h8R7vjLll3K0"
# Boolean Operators
# + colab={"base_uri": "https://localhost:8080/"} id="gANJZnial4Sl" outputId="26519f43-2ddd-4fb9-8969-103ad1e7aba7"
k=4
l=3
print(k>>2)#shift right twice
print(k<<2)#shift left twice
# + [markdown] id="mMVbLE2cmS_v"
# Relational Operators
# + colab={"base_uri": "https://localhost:8080/"} id="W7-VMNmJmUR0" outputId="334d9b1e-9d3a-4c95-a4dd-895618b9faab"
print(v>k) #v=1, k=2
print(v==k)
# + [markdown] id="20wZbcG8mrcU"
# Logical Operators
# + colab={"base_uri": "https://localhost:8080/"} id="LvgeinFxmtQe" outputId="459a7af8-d3f4-4905-d4af-d41ed09a6476"
print(v<k and k==k)
print(v<k or k==v)
print(not (v<k or k==v))
# + [markdown] id="n04clxWTmxr8"
# Identity Operators
# + colab={"base_uri": "https://localhost:8080/"} id="UBttEeSqmz-H" outputId="5ba69045-a77e-4a33-d2ab-10abaa89c3fb"
print(v is k)
print(v is not k)
| FUNDAMENTALS_OF_PYTHON.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <small><small><i>
# All of these python notebooks are available at [https://gitlab.erc.monash.edu.au/andrease/Python4Maths.git]
# </i></small></small>
# ## Strings
# Strings have already been discussed in Chapter 02, but can also be treated as collections similar to lists and tuples.
# For example
# + jupyter={"outputs_hidden": false}
S = '<NAME> is beautiful'
print([x for x in S if x.islower()]) # list of lower case charactes
words=S.split() # list of words
print("Words are:",words)
print("--".join(words)) # hyphenated
" ".join(w.capitalize() for w in words) # capitalise words
# -
# String Indexing and Slicing are similar to Lists which was explained in detail earlier.
# + jupyter={"outputs_hidden": false}
print(S[4])
print(S[4:])
# -
# ## Dictionaries
# Dictionaries are mappings between keys and items stored in the dictionaries. Alternatively one can think of dictionaries as sets in which something stored against every element of the set. They can be defined as follows:
# To define a dictionary, equate a variable to { } or dict()
# + jupyter={"outputs_hidden": false}
d = dict() # or equivalently d={}
print(type(d))
d['abc'] = 3
d[4] = "A string"
print(d)
# -
# As can be guessed from the output above. Dictionaries can be defined by using the `{ key : value }` syntax. The following dictionary has three elements
# + jupyter={"outputs_hidden": false}
d = { 1: 'One', 2 : 'Two', 100 : 'Hundred'}
len(d)
# -
# Now you are able to access 'One' by the index value set at 1
# + jupyter={"outputs_hidden": false}
print(d[1])
# -
# There are a number of alternative ways for specifying a dictionary including as a list of `(key,value)` tuples.
# To illustrate this we will start with two lists and form a set of tuples from them using the **zip()** function
# Two lists which are related can be merged to form a dictionary.
# + jupyter={"outputs_hidden": false}
names = ['One', 'Two', 'Three', 'Four', 'Five']
numbers = [1, 2, 3, 4, 5]
[ (name,number) for name,number in zip(names,numbers)] # create (name,number) pairs
# -
# Now we can create a dictionary that maps the name to the number as follows.
# + jupyter={"outputs_hidden": false}
a1 = dict((name,number) for name,number in zip(names,numbers))
print(a1)
# -
# Note that the ordering for this dictionary is not based on the order in which elements are added but on its own ordering (based on hash index ordering). It is best never to assume an ordering when iterating over elements of a dictionary.
#
# By using tuples as indexes we make a dictionary behave like a sparse matrix:
# + jupyter={"outputs_hidden": false}
matrix={ (0,1): 3.5, (2,17): 0.1}
matrix[2,2] = matrix[0,1] + matrix[2,17]
print(matrix)
# -
# Dictionary can also be built using the loop style definition.
# + jupyter={"outputs_hidden": false}
a2 = { name : len(name) for name in names}
print(a2)
# -
# ### Built-in Functions
# The **len()** function and **in** operator have the obvious meaning:
# + jupyter={"outputs_hidden": false}
print("a1 has",len(a1),"elements")
print("One is in a1",'One' in a1,"but not Zero", 'Zero' in a1)
# -
# **clear( )** function is used to erase all elements.
# + jupyter={"outputs_hidden": false}
a2.clear()
print(a2)
# -
# **values( )** function returns a list with all the assigned values in the dictionary. (Acutally not quit a list, but something that we can iterate over just like a list to construct a list, tuple or any other collection):
# + jupyter={"outputs_hidden": false}
[ v for v in a1.values() ]
# -
# **keys( )** function returns all the index or the keys to which contains the values that it was assigned to.
# + jupyter={"outputs_hidden": false}
{ k for k in a1.keys() }
# -
# **items( )** is returns a list containing both the list but each element in the dictionary is inside a tuple. This is same as the result that was obtained when zip function was used - except that the ordering has been 'shuffled' by the dictionary.
# + jupyter={"outputs_hidden": false}
", ".join( "%s = %d" % (name,val) for name,val in a1.items())
# -
# **pop( )** function is used to get the remove that particular element and this removed element can be assigned to a new variable. But remember only the value is stored and not the key. Because the is just a index value.
# + jupyter={"outputs_hidden": false}
val = a1.pop('Four')
print(a1)
print("Removed",val)
| 001-Jupyter/001-Tutorials/005-Python4Maths/04.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from model import *
from data import *
import tensorflow as tf
tf.config.experimental.list_physical_devices('GPU')
import tensorflow as tf
os.environ["CUDA_VISIBLE_DEVICES"] = "0"
gpu_options =tf.compat.v1.GPUOptions(per_process_gpu_memory_fraction=0.5)
gpu_options = tf.compat.v1.GPUOptions(allow_growth=True)
sess = tf.compat.v1.Session(config=tf.compat.v1.ConfigProto(gpu_options=gpu_options))
tf.compat.v1.keras.backend.set_session(sess)
from numpy.random import seed
import random
seed(42)
tf.random.set_seed(42)
random.seed(42)
# ## Train your Unet with membrane data
# membrane data is in folder membrane/, it is a binary classification task.
#
# The input shape of image and mask are the same :(batch_size,rows,cols,channel = 1)
# ### Train with data generator
TIMES = 50
TRAIN_SHAPE = 30
BATCH_SIZE = 2
STEP_PER_EPOCH = TIMES*TRAIN_SHAPE/BATCH_SIZE
EPOCHS = 5
IMG_SIZE = 256
TARGET_SIZE = (IMG_SIZE,IMG_SIZE)
data_gen_args = dict(rotation_range=0.2,
width_shift_range=0.05,
height_shift_range=0.05,
shear_range=0.05,
zoom_range=0.05,
horizontal_flip=True,
fill_mode='nearest')
myGene = trainGenerator(BATCH_SIZE,'data/membrane/train',
image_folder='image',
mask_folder='label',
aug_dict = data_gen_args,
flag_multi_class = False,
num_class = 2,
save_to_dir = None,
target_size = TARGET_SIZE,
seed = 42)
model = unet()
model_checkpoint = ModelCheckpoint('unet_membrane.hdf5', monitor='loss',verbose=1, save_best_only=True)
model.fit_generator(myGene,steps_per_epoch=STEP_PER_EPOCH,epochs=EPOCHS,callbacks=[model_checkpoint])
# ### Train with npy file
# +
#imgs_train,imgs_mask_train = geneTrainNpy("data/membrane/train/aug/","data/membrane/train/aug/")
#model.fit(imgs_train, imgs_mask_train, batch_size=2, nb_epoch=10, verbose=1,validation_split=0.2, shuffle=True, callbacks=[model_checkpoint])
# -
# ### test your model and save predicted results
testGene = testGenerator("data/membrane/test",num_image=30)
model = unet()
model.load_weights("unet_membrane.hdf5")
results = model.predict_generator(testGene,verbose=1)
saveResult("data/membrane/predict",results)
| trainUnet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Reading and plotting experimental time series data
import microval.experiment as mexp
import matplotlib.pyplot as plt
import numpy as np
import imp
imp.reload(mexp)
# Parameters to adjust.
# Paths to experimental *.mat files
binaraydata_mat_file_name = r"image_time_series_red.mat"
segmenteddata_mat_file_name = r"foreground_mask.mat"
fatiguedata_mat_file_name = r"fatigue_data_red.mat"
# Scale factor for conversion of image data to physical domain
scale_factor = 0.6
# Interesting segment for plotting
interesting_region = [660,720,118,200]
# ## Binary data
# Get the number of frames for binary data.
bindat = mexp.BinaryData(binaraydata_mat_file_name=binaraydata_mat_file_name, fatiguedata_mat_file_name=fatiguedata_mat_file_name)
num_bindat = bindat.get_num_binary_data()
print("Number of time steps:", num_bindat)
# Select a timestep.
selected_timestep = num_bindat-1
# Plot binary data (as difference image wrt to initial image).
fig=plt.figure(figsize=(16,9))
bindat.imshow(scale_factor = scale_factor, time_step_number = selected_timestep, subtract_initial_state = True)
# Plot binary data in interesting region.
fig=plt.figure(figsize=(16,9))
bindat.imshow(scale_factor = scale_factor, time_step_number = selected_timestep, region = interesting_region)
# Linking frame id's to cycle numbers.
#bindat = mexp.BinaryData(mat_file_name=mat_file_name)
cyclenumbers = bindat.cyclenumbers()
# Frame id -> cycle number
print("Number of cycles at frame {}: {:e}".format(selected_timestep,cyclenumbers[selected_timestep]))
# Cycle number -> approx. frame id
def frame_at_cycles(lt):
fr = np.argmin(np.abs(cyclenumbers-lt))
return fr
print("Frame at approximately {:e} cycles: {}".format(1e8,frame_at_cycles(1e8)))
# ## Segmented data
# Selecting a frame is not necessary, since segemented data only exists for the last timestep.
# Plot segmented data.
segdat = mexp.SegmentedData(segmenteddata_mat_file_name=segmenteddata_mat_file_name,fatiguedata_mat_file_name=fatiguedata_mat_file_name)
fig=plt.figure(figsize=(16,9))
segdat.imshow(scale_factor = scale_factor, invert_blackwhite = True)
# Plot segmented data in interesting region.
fig=plt.figure(figsize=(16,9))
segdat.imshow(scale_factor = scale_factor, invert_blackwhite = True, region = interesting_region)
| script/example/Experimental.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="iKRcnc5xmmfL"
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# + colab={"base_uri": "https://localhost:8080/", "height": 592} id="LF1cqH8umyCN" outputId="160feb72-1a87-4ba8-c753-7787a38c0e41"
data=pd.read_csv("data.csv")
plt.figure(figsize=(15,10))
plt.plot(data['N'],data['Connection'],label="Examination")
x=np.linspace(0,5000,100)
y=4.5*x
plt.plot(x,y,'r',label="y=4.5x")
plt.legend();
| Assignments/Assignment 3/Assignment3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="iUYJADCcFtZX"
# ## Importing all the required libraries
# + id="mMwQ9mplFiAt"
import numpy as np # for linear algebra
import pandas as pd # for data processing and CSV file input/output (like pd.read_csv, pd.DataFrame ...)
import seaborn as sns # another library
import matplotlib.pyplot as plt # to make plots from the data
sns.set(style="white", color_codes=True) # using white style with color codes in seaborn
import warnings # to ignore any version related warnings generated by seaborn
warnings.filterwarnings("ignore")
# + id="or55EWt_F0--"
from sklearn.datasets import load_iris
# + id="FuWqh5XEHcBF"
iris = load_iris() #loading dataset
# + id="iWNTotQVHeY3"
X = iris.data[:,:4]
y = iris.target #updating species data into y
# + colab={"base_uri": "https://localhost:8080/"} id="dJfxqfb8H3QF" outputId="45b6e1ac-823d-4917-b621-64459262b5a6"
X
# + colab={"base_uri": "https://localhost:8080/"} id="MZz2TalUKDYp" outputId="835941e8-1870-47d6-b31d-54c09d2a0d29"
y
# + id="vv5NkY36H-ks"
# transform the raw data from the dataset into a dataframe
df = pd.DataFrame(iris.data, columns = iris.feature_names)
df['species'] = iris.target
# + colab={"base_uri": "https://localhost:8080/", "height": 417} id="Bssu6rkwXjc_" outputId="bd76f6c6-b512-4b28-d6b2-1f6d1ee40c39"
df.head(100) #printing first few records
# + colab={"base_uri": "https://localhost:8080/"} id="CJYHggFFXjRh" outputId="555ffcad-9eb4-4c09-9072-a5e4e10db37d"
df.shape
# + colab={"base_uri": "https://localhost:8080/"} id="RAj1WqGccTDY" outputId="9350a4df-96e8-496b-cf93-b6c1c2ad88ba"
df.isnull().sum()
# + colab={"base_uri": "https://localhost:8080/", "height": 203} id="HhnqazvbcWz1" outputId="e4a34c5c-e9dd-4b05-8f8d-a36edff86b6e"
df.corr() #identifying correlation
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="nxnQRrhCcX4V" outputId="3af0f493-84d4-43c6-bf84-316baec92750"
df.describe() #description of data
# + colab={"base_uri": "https://localhost:8080/"} id="znTeESjvcbeL" outputId="8a8882fe-ed32-488a-cee8-1bd749fc50e0"
df.info()
# + [markdown] id="PDkVjSQZ5sgM"
# ## Data Visualization
# + colab={"base_uri": "https://localhost:8080/", "height": 820} id="FYfvYcK_5qNw" outputId="56b4cda7-019e-4dc8-cc17-bc882886b9de"
## from sklearn docucmentation
from mpl_toolkits.mplot3d import Axes3D
from sklearn.decomposition import PCA
x_min, x_max = X[:, 0].min() - .5, X[:, 0].max() + .5
y_min, y_max = X[:, 1].min() - .5, X[:, 1].max() + .5
plt.figure(2, figsize=(8, 6))
plt.clf()
# Plot the training points
plt.scatter(X[:, 0], X[:, 1], c=y, cmap=plt.cm.Set1,
edgecolor='k')
plt.xlabel('Sepal length')
plt.ylabel('Sepal width')
plt.xlim(x_min, x_max)
plt.ylim(y_min, y_max)
plt.xticks(())
plt.yticks(())
# To getter a better understanding of interaction of the dimensions
# plot the first three PCA dimensions
fig = plt.figure(1, figsize=(8, 6))
ax = Axes3D(fig, elev=-150, azim=110)
X_reduced = PCA(n_components=3).fit_transform(iris.data)
ax.scatter(X_reduced[:, 0], X_reduced[:, 1], X_reduced[:, 2], c=y,
cmap=plt.cm.Set1, edgecolor='k', s=40)
ax.set_title("First three PCA directions")
ax.set_xlabel("1st eigenvector")
ax.w_xaxis.set_ticklabels([])
ax.set_ylabel("2nd eigenvector")
ax.w_yaxis.set_ticklabels([])
ax.set_zlabel("3rd eigenvector")
ax.w_zaxis.set_ticklabels([])
plt.show()
# + id="phH08pzz7iYb"
# + colab={"base_uri": "https://localhost:8080/", "height": 341} id="ze5vRF297olC" outputId="d71cbfb9-5cf3-4d8a-efc5-8e75106a4fc6"
# a scatter plot for IRIS features
df.plot(kind="scatter", x="sepal length (cm)", y="sepal width (cm)")
# + colab={"base_uri": "https://localhost:8080/", "height": 598} id="aGLb_PcP7tPj" outputId="86fcaf36-903d-4cc6-8991-09917d565644"
# making use of seaborn jointplot to make bivariate scatterplots and univariate histograms and join them in the same figure
sns.jointplot(x="sepal length (cm)", y="sepal width (cm)", data=df, size=8)
# + [markdown] id="AIzLD8x9ce9Y"
# ### Seperating the data into X and y
# + colab={"base_uri": "https://localhost:8080/", "height": 203} id="tl59t_WVXjD7" outputId="2e26210b-7890-409a-9a1b-967bc4aa4115"
X=df.iloc[:, :-1]
X.head()
# + colab={"base_uri": "https://localhost:8080/"} id="qAa-i3VJXi-Z" outputId="70e677da-70c0-44f8-e5df-b6cd0287b8c2"
y=df.iloc[:,-1]
y.head()
# + id="v2FtdT_Ra3xj"
# (Standardizing) Scaling the data to be between -1 and 1
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
scaler.fit(X)
X = scaler.transform(X)
# + [markdown] id="WnrQztxlbRtf"
# ## Train-test split on the data
# + id="I1Yv0j4ma3t6"
from sklearn.model_selection import train_test_split as tts
X_train, X_test, y_train, y_test = tts(X, y, test_size=0.3, random_state=0)
# + [markdown] id="KGucICJGeB9X"
# ### Using a KNN model to fit and then make predictions
# + id="7mN63sJBa3oq"
from sklearn.neighbors import KNeighborsClassifier as knc
from sklearn.metrics import confusion_matrix, accuracy_score
from sklearn.model_selection import cross_val_score
# + id="-V-6qhela3lZ"
# Instantiate learning model (k = 9)
classifier = knc(n_neighbors=9)
# Fitting the model
classifier.fit(X_train, y_train)
# Predicting the Test set results
y_pred = classifier.predict(X_test)
# + colab={"base_uri": "https://localhost:8080/"} id="wRXZvlJEa3hu" outputId="ac7f5686-237f-418c-d4de-e5e2c2002bd2"
con_matrix = confusion_matrix(y_test, y_pred)
con_matrix
# + colab={"base_uri": "https://localhost:8080/"} id="9b4umEysa3e3" outputId="1d5e03b6-ec83-4345-d25e-bde9d62fbd11"
accuracy = accuracy_score(y_test, y_pred)*100
print('The model accurace is ' + str(round(accuracy, 2)) + ' %.')
# + [markdown] id="ACJ0Qkmefenb"
# # Cross validation for model validation
# + id="Xt0-u4qXfc4n"
# creating list of K for KNN
k_list = list(range(1,50,2))
# creating list of cv scores
cv_scores = []
# perform 10-fold cross validation
for k in k_list:
knn = knc(n_neighbors=k)
scores = cross_val_score(knn, X_train, y_train, cv=10, scoring='accuracy')
cv_scores.append(scores.mean())
# + colab={"base_uri": "https://localhost:8080/", "height": 651} id="umReoEmNfqra" outputId="936cdb70-f6e4-4a33-c3f6-73b4d20e319e"
# changing to misclassification error
MSE = [1 - x for x in cv_scores]
plt.figure()
plt.figure(figsize=(15,10))
plt.title('The optimal number of neighbors', fontsize=20, fontweight='bold')
plt.xlabel('Number of Neighbors K', fontsize=15)
plt.ylabel('Misclassification Error', fontsize=15)
sns.set_style("whitegrid")
plt.plot(k_list, MSE)
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="_3hQckXAf3G_" outputId="f33799ce-56b1-47a9-819e-677b920dec8e"
# finding best k
best_k = k_list[MSE.index(min(MSE))]
print("The optimal number of neighbors is %d." % best_k)
# + id="rcOASoK9f29n"
from sklearn.model_selection import RandomizedSearchCV
from sklearn.ensemble import RandomForestClassifier
from scipy.stats import uniform, truncnorm, randint
# + [markdown] id="dkPMiXmHznVa"
# ## Randomized Search CV in KNN Classifier
# + colab={"base_uri": "https://localhost:8080/"} id="v4CUbkaozEuR" outputId="fff3062b-a9bc-4b94-e387-e6477c7ada11"
from sklearn.model_selection import RandomizedSearchCV
from sklearn.datasets import load_iris
iris = load_iris()
X=iris.data
y=iris.target
k_range=list(range(1,31))
options=['uniform', 'distance']
param_dist = dict(n_neighbors=k_range, weights=options)
knn = knc(n_neighbors = 1)
rand = RandomizedSearchCV(knn, param_dist, cv=10, scoring='accuracy', n_iter=10, random_state=5)
rand.fit(X, y)
print(rand.best_score_)
print(rand.best_params_)
best_scores=[]
for i in range(20):
rand = RandomizedSearchCV(knn, param_dist, cv=10, scoring='accuracy', n_iter=10)
rand.fit(X, y)
best_scores.append(round(rand.best_score_,3))
print(best_scores)
# + [markdown] id="9fTl8Cso5_WN"
# ## SVM Classifier to make predictions
# + id="3c-7RlEa1Mvq"
from sklearn.metrics import mean_squared_error
from math import sqrt
# + colab={"base_uri": "https://localhost:8080/"} id="7b6NAhW71Msl" outputId="0abd786c-dc09-45a7-b851-20b42166ab7a"
from sklearn.svm import SVC
s1 = SVC(kernel='rbf', random_state=0, gamma=.10, C=1.0)
s1.fit(X_train,y_train)
# + colab={"base_uri": "https://localhost:8080/"} id="zPPS1RyP1cw-" outputId="321e7140-71ae-4f98-c46f-8a0d4d2efa8e"
from sklearn.model_selection import cross_val_score
svc=SVC(kernel='rbf',gamma=0.01)
scores = cross_val_score(svc, X, y, cv = 10, scoring = 'accuracy')
print(scores)
print(scores.mean())
# + colab={"base_uri": "https://localhost:8080/"} id="jQMDdUlZ1ct0" outputId="9015fb48-531f-4730-8010-ed27a08a96b4"
svc=SVC(kernel='rbf')
scores = cross_val_score(svc, X, y, cv=10, scoring='accuracy') #cv is cross validation
print(scores)
print(scores.mean())
# + colab={"base_uri": "https://localhost:8080/"} id="3tGokuwn1cqp" outputId="5aa53d03-443a-47b3-93ba-4b8fe0132806"
print(s1.score(X_train,y_train)) #accuracy of training data
# + colab={"base_uri": "https://localhost:8080/"} id="m6CuNW6A1cno" outputId="d61e085c-a172-4d99-9ebc-ba8d7b9cc747"
print(s1.score(X_test,y_test)) #accuracy of test data
# + id="kcgdcP-V1clF"
p=s1.predict(X_test)
# + colab={"base_uri": "https://localhost:8080/"} id="VJB82TY91ciQ" outputId="90c82847-61fe-40aa-bc03-bb25f4bb49cb"
#calculation of root mean square error
rmse= sqrt(mean_squared_error(p, y_test))
print(rmse)
# + colab={"base_uri": "https://localhost:8080/"} id="uHu4M02Z1cf5" outputId="45923bce-821c-47a4-b3f6-6c8ce73c4df9"
from sklearn.metrics import classification_report
print(classification_report(y_test,p))
# + colab={"base_uri": "https://localhost:8080/"} id="9MYL-V4B1cdG" outputId="6a5386bf-21a8-4fa6-c9d5-d074af973169"
from sklearn.metrics import confusion_matrix
print(confusion_matrix(y_test,p))
# + [markdown] id="gSqi8DW16Hx0"
# ## Grid Search CV in SVM Classifier for hyperparamter optimization
# + colab={"base_uri": "https://localhost:8080/"} id="GOPSH1oL1cai" outputId="8607b578-fa4c-4eb8-db1d-38dfd22c884f"
#Tuning SVM hyperparameters by using GridSearchCV with cross validation
from sklearn.model_selection import GridSearchCV
param_grid = {'C':[0.1,1,10,100], 'gamma':[1,0.1,0.01,0.001]}
g1 = GridSearchCV(SVC(), param_grid, refit = True, verbose=3)
g1.fit(X_train, y_train)
# + colab={"base_uri": "https://localhost:8080/"} id="-S2So3Bf1cYA" outputId="30902839-bc38-4ed5-c720-0f880de11b44"
p2= g1.predict(X_test)
print(classification_report(y_test, p2))
# + colab={"base_uri": "https://localhost:8080/"} id="S71OJfd51cVm" outputId="6981d37f-e6d8-4957-86ef-4af88f7b96fd"
#default RBF kernel
from sklearn import metrics
svc=SVC(kernel='rbf')
svc.fit(X_train,y_train)
p3=svc.predict(X_test)
print('Accuracy Score:')
print(metrics.accuracy_score(y_test,p3))
# + colab={"base_uri": "https://localhost:8080/"} id="rdTHp8X_27re" outputId="470afc57-7472-4779-8517-d367b7eccd85"
#taking different gamma values for rbf kernel
gamma_range=[0.0001,0.001,0.01,0.1,1,10,100]
acc_score=[]
for g in gamma_range:
svc = SVC(kernel='rbf', gamma=g)
scores = cross_val_score(svc, X, y, cv=10, scoring='accuracy')
acc_score.append(scores.mean())
print(acc_score)
# + id="4i6xwgvJ27hY"
| Assignments/KNN_and_SVM_with_IRIS_dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Train a simple deep neural network on the MNIST dataset
#
# #### Gets to 98.18% test accuracy after 20 epochs (there is a lot of margin for parameter tuning).
# +
from __future__ import print_function
import tensorflow.keras as keras
from tensorflow.keras.datasets import mnist
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Dropout, Flatten
from tensorflow.keras.optimizers import RMSprop
# +
batch_size = 128
num_classes = 10
epochs = 20
# the data, shuffled and split between train and test sets
(x_train, y_train), (x_test, y_test) = mnist.load_data()
x_train = x_train.reshape(60000, 784)
x_test = x_test.reshape(10000, 784)
x_train = x_train.astype('float32')
x_test = x_test.astype('float32')
x_train /= 255
x_test /= 255
print(x_train.shape, 'train samples')
print(x_test.shape, 'test samples')
# -
# convert class vectors to binary class matrices
y_train = keras.utils.to_categorical(y_train, num_classes)
y_test = keras.utils.to_categorical(y_test, num_classes)
print(y_train.shape, y_test.shape)
print(y_train)
# +
model = Sequential()
model.add(Dense(512, activation='relu', input_shape=(784,)))
model.add(Dropout(0.2))
model.add(Dense(512, activation='relu'))
model.add(Dropout(0.2))
model.add(Dense(num_classes, activation='softmax'))
model.compile(loss='categorical_crossentropy',
optimizer=RMSprop(),
metrics=['accuracy'])
# -
model.summary()
# +
# #some learners constantly reported 502 errors in Watson Studio.
# #This is due to the limited resources in the free tier and the heavy resource consumption of Keras.
# #This is a workaround to limit resource consumption
# from keras import backend as K
# K.set_session(K.tf.Session(config=K.tf.ConfigProto(intra_op_parallelism_threads=1, inter_op_parallelism_threads=1)))
# -
history = model.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
verbose=1,
validation_data=(x_test, y_test))
score = model.evaluate(x_test, y_test, verbose=0)
print('Test loss:', score[0])
print('Test accuracy:', score[1])
| 03_Applied_AI_DeepLearning/notebooks/week_3_3_dnn_mnist_classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.9 64-bit (''workflow-calcium-imaging'': conda)'
# name: python379jvsc74a57bd01a512f474e195e32ad84236879d3bb44800a92b431919ef0b10d543f5012a23c
# ---
# # Interactively run workflow calcium imaging
#
# + This notebook walks you through the steps in detail to run the `workflow-calcium-imaging`.
#
# + The workflow requires the calcium imaging acquired data from ScanImage or Scanbox and processed data from Suite2p or CaImAn.
#
# + If you haven't configured the paths, refer to [01-configure](01-configure.ipynb).
#
# + To overview the schema structures, refer to [02-workflow-structure](02-workflow-structure.ipynb).
#
# + If you need a more automatic approach to run the workflow, refer to [03-automate](03-automate-optional.ipynb).
# Let's change the directory to the package root directory to load the local configuration (`dj_local_conf.json`).
import os
os.chdir('..')
import numpy as np
# ## `Pipeline.py`
#
# + This script `activates` the DataJoint `elements` and declares other required tables.
from workflow_calcium_imaging.pipeline import *
# ## Schema diagrams
dj.Diagram(subject.Subject) + dj.Diagram(session.Session) + dj.Diagram(scan) + dj.Diagram(imaging.Processing)
# ## Insert an entry into `subject.Subject`
subject.Subject.heading
subject.Subject.insert1(dict(subject='subject3',
sex='F',
subject_birth_date='2020-01-01',
subject_description='Scanbox acquisition. Suite2p processing.'))
# ## Insert an entry into `lab.Equipment`
Equipment.insert1(dict(scanner='Scanbox'))
# ## Insert an entry into `session.Session`
session.Session.describe();
session.Session.heading
# +
session_key = dict(subject='subject3', session_datetime='2021-04-30 12:22:15.032')
session.Session.insert1(session_key)
session.Session()
# -
# ## Insert an entry into `session.SessionDirectory`
#
# + The `session_dir` is the relative path to the `imaging_root_data_dir` for the given session, in POSIX format with `/`.
#
# + Instead of a relative path, `session_dir` could be an absolute path but it is not recommended as the absolute path would have to match the `imaging_root_data_dir` in `dj_local_conf.json`.
session.SessionDirectory.describe();
session.SessionDirectory.heading
# +
session.SessionDirectory.insert1(dict(subject='subject3',
session_datetime='2021-04-30 12:22:15.032',
session_dir='subject3/210107_run00_orientation_8dir'))
session.SessionDirectory()
# -
# ## Insert an entry into `scan.Scan`
scan.Scan.heading
scan.Scan.insert1(dict(subject='subject3',
session_datetime='2021-04-30 12:22:15.032',
scan_id=0,
scanner='Scanbox',
acq_software='Scanbox',
scan_notes=''))
scan.Scan()
# ## Populate `scan.ScanInfo`
#
# + This imported table stores information about the acquired image (e.g. image dimensions, file paths, etc.).
# + `populate` automatically calls `make` for every key for which the auto-populated table is missing data.
# + `populate_settings` passes arguments to the `populate` method.
# + `display_progress=True` reports the progress bar
scan.ScanInfo.describe();
scan.ScanInfo.heading
populate_settings = {'display_progress': True}
scan.ScanInfo.populate(**populate_settings)
scan.ScanInfo()
# ## (Optional) Insert a new entry into `imaging.ProcessingParamSet` for Suite2p or CaImAn
#
# + Define and insert the parameters that will be used for the Suite2p or CaImAn processing.
#
# + This step is not needed if you are using an existing ProcessingParamSet.
#
# ### Define Suite2p parameters
params_suite2p = {'look_one_level_down': 0.0,
'fast_disk': [],
'delete_bin': False,
'mesoscan': False,
'h5py': [],
'h5py_key': 'data',
'save_path0': [],
'subfolders': [],
'nplanes': 1,
'nchannels': 1,
'functional_chan': 1,
'tau': 1.0,
'fs': 10.0,
'force_sktiff': False,
'preclassify': 0.0,
'save_mat': False,
'combined': True,
'aspect': 1.0,
'do_bidiphase': False,
'bidiphase': 0.0,
'do_registration': True,
'keep_movie_raw': False,
'nimg_init': 300,
'batch_size': 500,
'maxregshift': 0.1,
'align_by_chan': 1,
'reg_tif': False,
'reg_tif_chan2': False,
'subpixel': 10,
'smooth_sigma': 1.15,
'th_badframes': 1.0,
'pad_fft': False,
'nonrigid': True,
'block_size': [128, 128],
'snr_thresh': 1.2,
'maxregshiftNR': 5.0,
'1Preg': False,
'spatial_hp': 50.0,
'pre_smooth': 2.0,
'spatial_taper': 50.0,
'roidetect': True,
'sparse_mode': False,
'diameter': 12,
'spatial_scale': 0,
'connected': True,
'nbinned': 5000,
'max_iterations': 20,
'threshold_scaling': 1.0,
'max_overlap': 0.75,
'high_pass': 100.0,
'inner_neuropil_radius': 2,
'min_neuropil_pixels': 350,
'allow_overlap': False,
'chan2_thres': 0.65,
'baseline': 'maximin',
'win_baseline': 60.0,
'sig_baseline': 10.0,
'prctile_baseline': 8.0,
'neucoeff': 0.7,
'xrange': np.array([0, 0]),
'yrange': np.array([0, 0])}
# ### Insert Suite2p parameters
#
# + A method of the class `ProcessingParamset` called `insert_new_params` is a helper function to insert the Suite2p or CaIman parameters and ensures that the parameter set inserted is not duplicated.
imaging.ProcessingParamSet.insert_new_params(
processing_method='suite2p',
paramset_idx=0,
params=params_suite2p,
paramset_desc='Calcium imaging analysis with Suite2p using default Suite2p parameters')
# ## Insert new ProcessingTask to trigger ingestion of motion correction and segmentation results
#
# + Motion correction and segmentation are performed for each scan in Suite2p or CaImAn.
#
# + An entry in `ProcessingTask` indicates a set of motion correction and segmentation results (generated from Suite2p or CaImAn outside of `workflow-calcium-imaging`) are ready to be ingested. In a future release, an entry in `ProcessingTask` can also indicate a new processing job (using Suite2p or CaImAn) is to be triggered.
#
# + Two pieces of information need to be specified:
#
# + The `paramset_idx` is the parameter set stored in `imaging.ProcessingParamSet` that is used for the Suite2p or CaImAn processing job.
#
# + The `processing_output_dir` stores the directory of the processing results (relative to the imaging root data directory).
imaging.ProcessingTask.insert1(dict(subject='subject3',
session_datetime='2021-04-30 12:22:15.032',
scan_id=0,
paramset_idx=0,
processing_output_dir='subject3/210107_run00_orientation_8dir/suite2p'))
# ## Populate `imaging.Processing`
imaging.Processing.populate(**populate_settings)
# ## Insert new Curation following the ProcessingTask
#
# + The next step in the pipeline is the curation of motion corection and segmentation results.
#
# + If a manual curation was implemented, an entry needs to be manually inserted into the table `imaging.Curation`, which specifies the directory to the curated results in `curation_output_dir`.
#
# + If we would like to use the processed outcome directly, an entry is also needed in `imaging.Curation`. A method `create1_from_processing_task` was provided to help this insertion. It copies the `processing_output_dir` in `imaging.ProcessingTask` to the field `curation_output_dir` in the table `imaging.Curation` with a new `curation_id`.
#
# + In this example, we create/insert one `imaging.Curation` for each `imaging.ProcessingTask`, specifying the same output directory.
#
# + To this end, we could also make use of a convenient function `imaging.Curation().create1_from_processing_task()`
imaging.Curation.insert1(dict(subject='subject3',
session_datetime='2021-04-30 12:22:15.032',
scan_id=0,
paramset_idx=0,
curation_id=0,
curation_time='2021-04-30 12:22:15.032',
curation_output_dir='subject3/210107_run00_orientation_8dir/suite2p',
manual_curation=False,
curation_note=''))
# ## Populate `imaging.MotionCorrection`
#
# + This table contains the rigid or non-rigid motion correction data including the shifts and summary images.
#
imaging.MotionCorrection.populate(**populate_settings)
# ## Populate `imaging.Segmentation`
#
# + This table contains the mask coordinates, weights, and centers.
imaging.Segmentation.populate(**populate_settings)
# ## Add another set of results from a new round of curation
#
# If you performed curation on an existing processed results (i.e. motion correction or segmentation) then:
#
# + Add an entry into `imaging.Curation` with the directory of the curated results and a new `curation_id`.
#
# + Populate the `imaging.MotionCorrection` and `imaging.Segmentation` tables again.
# ## Populate `imaging.MaskClassification`
#
# + This table contains the classification of the segmented masks and the confidence of classification.
imaging.MaskClassification.populate(**populate_settings)
# ## Populate `imaging.Fluorescence`
#
# + This table contains the fluorescence traces prior to filtering and spike extraction.
imaging.Fluorescence.populate(**populate_settings)
# ## Populate `imaging.Activity`
# + This table contains the inferred neural activity from the fluorescence traces.
imaging.Activity.populate(**populate_settings)
# ## Next steps
#
# + Proceed to the [05-explore](05-explore.ipynb) to learn how to query, fetch, and visualize the imaging data.
| notebooks/03-process.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from colubridae.category import Category,Functor,NaturalTransformation
from colubridae.sets import SetObject,Function,Relation
# ### Defining a cyclic group
X=SetObject(["a","b","c","d","e","f","g","h"])
t = Function(X,X,{"a":"b","b":"c","c":"d","d":"e","e":"f","f":"g","g":"h","h":"a"})
Z8 = Category([X],[t])
print(len(Z8.morphisms))
# +
# Determining the automorphism group of Z8
for x in list(Z8.morphisms):
print(Category([X],[x])==Z8)
print(x.mapping)
# -
F = Functor(Z8,Z8,{X:X},{t:t*t*t},from_generators=True)
print(F(t*t).mapping)
G = Functor(Z8,Z8,{X:X},{t:t*t*t*t*t},from_generators=True)
AutZ8 = Category([Z8],[F,G])
print(len(AutZ8.morphisms))
# ### More complex functors
A=SetObject(["a","b"])
f = Function(A,A,{"a":"b","b":"a"})
Z2 = Category([A],[f])
print(len(Z2.morphisms))
B=SetObject(["p","q","r","s"])
g = Relation(B,B,{"p":{"q","r"},"q":{"p","s"},"r":{"r"},"s":{"r"}})
M = Category([B],[g])
for x in M.morphisms:
print(x.mapping)
F=Functor(M,Z2,{B:A},{g:f},from_generators=True)
F(g*g*g).mapping
# ### Natural transformations
# +
U=SetObject(["u"])
V=SetObject(["v"])
W=SetObject(["w"])
fUV = Function(U,V,{"u":"v"})
fVW = Function(V,W,{"v":"w"})
cat1 = Category([U,V,W],[fUV,fVW])
F1 = Functor(cat1,Z8,{U:X,V:X,W:X},{fUV:t,fVW:t*t},from_generators=True)
F2 = Functor(cat1,Z8,{U:X,V:X,W:X},{fUV:t*t,fVW:t*t},from_generators=True)
N1 = NaturalTransformation(F1,F2,{U:t,V:t*t,W:t*t})
N2 = NaturalTransformation(F2,F1,{U:t*t,V:t,W:t})
func_cat = Category([F1,F2],[N1,N2])
print(len(func_cat.morphisms))
| examples.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/vksriharsha/Human-Cancer-Prediction/blob/main/TCGA_HumanCancer_prediction.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="6abikZSwXFbJ" outputId="1e649f3f-fccd-47c4-9ba2-302bd30fa33e"
from google.colab import drive
drive.mount('/content/drive')
# + id="ZiGTUKClaxz9"
import pandas as pd
from sklearn.preprocessing import LabelEncoder
from sklearn.preprocessing import OneHotEncoder
from sklearn import preprocessing
from sklearn.feature_selection import VarianceThreshold
import numpy as np
from sklearn.model_selection import train_test_split
import matplotlib.pyplot as plt
import keras
from keras.models import Sequential
from keras.layers import Dense
from keras.layers import Conv2D, MaxPooling2D, Dense, Dropout, Activation, Flatten, Input
from keras.callbacks import EarlyStopping, ModelCheckpoint
from keras.layers.advanced_activations import LeakyReLU
from sklearn.metrics import precision_recall_curve, roc_curve, auc, average_precision_score
from sklearn.model_selection import StratifiedKFold
# + colab={"base_uri": "https://localhost:8080/", "height": 830} id="3W4sBJKYY9kB" outputId="7c85a668-1e18-48cc-9df7-69b88075a4e2"
tcga_data_df = pd.read_csv('/content/drive/MyDrive/Thesis/Human-Cancer-Prediction/TCGA_GTEX_Data_18212_7142.tsv', delimiter='\t')
tcga_metadata_df = pd.read_csv('/content/drive/MyDrive/Thesis/Human-Cancer-Prediction/TCGA_GTEX_MetaData_7142_23.tsv', delimiter='\t')
tcga_data_df = tcga_data_df.drop(['NCBI_description','NCBI_other_designations','NCBI_chromosome', 'NCBI_map_location', 'NCBI_OMIM', 'CGC_Tumour Types(Somatic)', 'CGC_Tumour Types(Germline)', 'CGC_Role in Cancer', 'CGC_Translocation Partner', 'CGC_Somatic', 'CGC_Germline', 'CGC_Mutation Types', 'CGC_Molecular Genetics', 'CGC_Tissue Type', 'CGC_Cancer Syndrome', 'CGC_Other Syndrome', 'OMIM_Comments', 'OMIM_Phenotypes', 'Hugo_RefSeq IDs', 'Hugo_Ensembl gene ID', 'Hugo_Enzyme IDs', 'Hugo_Pubmed IDs', 'Hugo_Locus group', 'Hugo_Gene group name'],axis=1)
tcga_data_df = tcga_data_df.T
tcga_data_df.columns = tcga_data_df.iloc[0]
tcga_data_df = tcga_data_df.drop(tcga_data_df.index[0])
def x(a):
return np.log2(a.astype('float32') + 1)
tcga_data_df = tcga_data_df.apply(x, axis = 1)
tcga_data_df
# + colab={"base_uri": "https://localhost:8080/", "height": 450} id="uiO7BuWEzNJB" outputId="c5105000-d829-4bae-f12f-ef195ed9072f"
tcga_metadata_df = tcga_metadata_df[['portions.analytes.aliquots.submitter_id', 'age', 'tissue', 'clinical.disease']]
tcga_metadata_df['clinical.disease'] = tcga_metadata_df['clinical.disease'].fillna('normal')
tcga_metadata_df = tcga_metadata_df.set_index('portions.analytes.aliquots.submitter_id')
tcga_metadata_df
# + colab={"base_uri": "https://localhost:8080/", "height": 297} id="UMJOtHL85bjY" outputId="8114709b-e18e-45c5-afbf-facc337ba333"
counts = tcga_metadata_df['clinical.disease'].value_counts()
plt.rcParams["figure.figsize"] = (20,3)
plt.plot(counts)
plt.xticks(rotation = 90)
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="5SxfwxZz76Qt" outputId="d89b97a8-f140-4ee3-993d-84a907ce09fa"
counts
# + colab={"base_uri": "https://localhost:8080/", "height": 255} id="5G6yru5rMTKe" outputId="caf64aa1-8fdb-42cb-c0aa-39992c707e44"
tissue_counts = tcga_metadata_df['tissue'].value_counts()
plt.rcParams["figure.figsize"] = (20,3)
plt.plot(tissue_counts)
plt.xticks(rotation = 90)
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="RouD5ZR2Mx5C" outputId="808e3364-8b10-45a8-deca-68c8a87a39ba"
tissue_counts
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="puLHeCwn85KE" outputId="9774b4b4-062a-4e5b-f83c-63715c5dc798"
tcga_data_df = pd.merge(tcga_data_df, tcga_metadata_df, left_index=True, right_index=True)
tcga_data_df
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="j5xe4nXyVb6l" outputId="0e6bdb75-37a4-4ee6-e959-df3d6ce1f933"
le = preprocessing.LabelEncoder()
tcga_data_df['tissue'] = le.fit_transform(tcga_data_df.tissue.values)
tcga_data_df['clinical.disease'] = le.fit_transform(tcga_data_df['clinical.disease'].values)
tcga_data_df
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="OtyiJTVzgF1q" outputId="eafe26d1-feff-42d7-a51a-e8cd8f328531"
tcga_data_df = tcga_data_df.drop(['age'],axis=1)
tcga_data_df
# + colab={"base_uri": "https://localhost:8080/"} id="X2dgmaWnZ_h-" outputId="addc41b2-4f1b-483b-d613-f0a656608c71"
X = tcga_data_df.iloc[:,:18213].values
Y = tcga_data_df.iloc[:,18213:18214].values
X = np.asarray(X).astype('float32')
X
# + colab={"base_uri": "https://localhost:8080/"} id="QBA46eUspI11" outputId="f6fbac15-a732-496c-8868-1c7dc062ad8b"
X.shape
# + colab={"base_uri": "https://localhost:8080/"} id="YcGrTo4oAh5A" outputId="7760fbfb-e64d-4604-8c40-179347335502"
Y
# + id="Z5JNWalrQsLf"
v_threshold = VarianceThreshold(threshold=3.3580)
v_threshold.fit(X)
result = v_threshold.get_support()
bin = np.bincount(result)
indices = []
for idx, i in enumerate(result):
if i == False:
indices.append(idx)
X = np.delete(X, indices, 1)
# + colab={"base_uri": "https://localhost:8080/"} id="wmGjN0UOpmgp" outputId="0a2ca2e1-ce80-4227-8b56-0d3b7b2e2e29"
X.shape
# + colab={"base_uri": "https://localhost:8080/"} id="fvyheiXUDyBQ" outputId="06c1f2e7-f578-4fa9-89d2-7051dbd4d953"
ohe = OneHotEncoder()
Y = Y.reshape(len(Y), 1)
Y = ohe.fit_transform(Y).toarray()
Y.shape
# + colab={"base_uri": "https://localhost:8080/"} id="d8S3veZa6N5e" outputId="e4bbb9aa-4875-496f-c83c-b17234f7aedd"
X = np.reshape(X, (-1, 35, 100))
X.shape
# + colab={"base_uri": "https://localhost:8080/"} id="R-ggiWpSaucn" outputId="3e1a8516-d4bc-4110-f74f-4065103d5b26"
X_train,X_test,y_train,y_test = train_test_split(X,Y,stratify=Y,test_size = 0.25, random_state=42)
X_train
# + colab={"base_uri": "https://localhost:8080/"} id="LGZZFpMhrU4E" outputId="8f36d4ab-9a67-4f7d-c4aa-f4adf938035b"
img_rows, img_cols = len(X_test[0]), len(X_test[0][0])
num_classes = len(y_train[0])
batch_size = 128
epochs = 20
seed = 7
np.random.seed(seed)
input_shape = [img_rows, img_cols, 1]
X_train = X_train.reshape(X_train.shape[0], img_rows, img_cols, 1)
X_train.shape
# + id="1iYhGzdq-kAw"
X_test = X_test.reshape(X_test.shape[0], img_rows, img_cols, 1)
# + colab={"base_uri": "https://localhost:8080/"} id="YerlISooziz1" outputId="455eb00a-500e-4304-db37-bf6827ccf33c"
model = Sequential()
## *********** First layer Conv
model.add(Conv2D(32, kernel_size=(1, 35), strides=(1, 1),
input_shape=input_shape))
model.add(Activation('relu'))
model.add(MaxPooling2D(1, 2))
## ********* Classification layer
model.add(Flatten())
model.add(Dense(128, activation='relu'))
model.add(Dense(num_classes, activation='softmax'))
model.compile(loss='categorical_crossentropy',
optimizer='adam',
metrics=['categorical_accuracy'])
callbacks = [EarlyStopping(monitor='categorical_accuracy', patience=3, verbose=0)]
model.summary()
# + colab={"base_uri": "https://localhost:8080/"} id="TCUxlXRj9gli" outputId="409d21ab-92e4-4995-d7bd-451ba6fbdfce"
history = model.fit(X_train, y_train,
batch_size=batch_size,
epochs=epochs, callbacks=callbacks)
# + id="KiO8EVv7bGyq" colab={"base_uri": "https://localhost:8080/"} outputId="cd44af8e-37a8-4773-f381-f32c2df2db21"
model.evaluate(X_test, y_test)
# + colab={"base_uri": "https://localhost:8080/", "height": 443} id="VvSwVUvlcUwv" outputId="176c9cdb-d44f-4e56-e5e9-ff0f3967189d"
history = model.fit(X_train, y_train, epochs=100, batch_size=64)
# + colab={"base_uri": "https://localhost:8080/"} id="Mc_vsvxkdLOD" outputId="4fd8afdf-e3de-429c-e2bf-da52d86804a3"
_, accuracy = model.evaluate(X_test, y_test)
print('Accuracy: %.2f' % (accuracy*100))
| TCGA_HumanCancer_prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/JCesar-M/daa_2021_1/blob/master/7_Octubre.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="31SY48eySg_J"
# #Busqueda lineal
#
# Dado un conjunto de datos no ordenados, la busqueda lineal consiste en el recorrer el conjunto de datos desde el inicio al final, moviendose de unos hasta encontrar el elemento o llegar al final del conjunto.
#
# datos = [ 4,18,47,2,34,14,78,12,48,21,31,19,1,3,5]
#
# #Busqueda binaria
# Funciona sobre un conjunto de datos lineal ordenado.
#
# Consiste en dividir el conjuto en mitades y buscar en la mitad, preguntas si el elemento buscado no esta en la mitad, preguntas si el elemento esta a la derecha o a la izq. Haces la lista igual a la mitad correspondiente y repites el proceso.
#
#
# L = [1,2,3,4,5,12,14,18,19,21,31,34,47,48,78]
# DER = longitud(L) -1
# IZQ =0
#
# MID apuntara a la mitad del segmento de busqueda
#
# Buscado: es el valor a buscar
#
# 1.Hacer DER = longitud (L) -1
#
# 2.Hacer IZQ = 0
#
# 3.Si IZQ > DER significa que hay un error
#
# 4.en los datos
#
# 5.Calcular MID = int(IZQ + DER/2)
#
# 6.Mientras L(MID) != buscado hacer
#
# - preguntar L(MiD) > buscado
# - hacer DER = MID
# - De los contrario
# - HAcer IZQ = MID
# - preguntar (DER -IZQ) % 2
# - MID = (IZQ + (DER - IZQ)/2)
# - de lo contrario
# - MID = (IZQ +((DER -IZQ) /2)
# 7. Reurn MID
#
#
# + id="n0xy7gkYSW8U" outputId="a225e068-e7b4-4d3a-87ee-2396a7049f25" colab={"base_uri": "https://localhost:8080/"}
"""Busqueda lineal
regresa la posicion del elemmento 'buscado' si se encuentra dentro de la lista
regresa -1 si el elemento buscado no existe dentro de la lista
"""
def busq_lineal ( L , buscado ):# Pasamos la lista y el elemento que se busca
indice = -1
contador = 0
for idx in range(len(L)):
if L[idx] == buscado:
indice = idx
break
print(f"Numero de comparaciones realizadas = {contador}")
return indice
"""
Busqueda Binario
"""
def buqueda_Binaria(L, buscado):
IZQ = 0
DER = len(L)-1
MID = int((IZQ + DER)/2)
if len(L) % 2 == 0:
MID = (DER //2)+1
else:
MID = DER //2
while (L[MID] != buscado ):
if L[MID] > buscado:
DER = MID
else:
IZQ = MID
if (DER - IZQ) % 2 == 0:
MID = IZQ + ((DER -IZQ)//2)
else:
MID = IZQ + ((DER - IZQ)//2)
return MID
def main():
datos = [4,18,47,2,34,14,78,12,48,21,31,19,1,3,5]
dato = int(input("Que valor quieres buscar: "))
resultado = busq_lineal(datos, datos )
print("Resultado : ",resultado)
print("Busqueda lienal en una lista ordenada")
datos.sort()
print(datos)
resultado = busq_lineal(datos, dato)
print("Resultado: ",resultado)
print("Busqueda binaria")
main()
| 7_Octubre.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# ## <center><strong>Metode Numerik</strong><br />
# <img alt="" src="images/MetNum.png" style="height: 200px;" /></center>
#
# ## <center><font color="blue">Curve Fitting</font> <br>(C) <NAME> - 2022<br> <center><font color="blue">taudata Analytics</font> </center>
#
# ### <center><a href="https://taudata.blogspot.com/2020/04/mfdsnm-06.html" target="_blank"><span style="color: #0009ff;">https://taudata.blogspot.com/2020/04/mfdsnm-06.html</span></a></center>
# -
import numpy as np, matplotlib as mpl, matplotlib.pyplot as plt
from numpy import polyfit, poly1d
# +
# %matplotlib inline
x = np.linspace(-5, 5, 100)
y = 4 * x + 1.5
noise_y = y + np.random.randn(y.shape[-1]) * 2.5
p = plt.plot(x, noise_y, 'rx')
p = plt.plot(x, y, 'b:')
# -
coeff = polyfit(x, noise_y, 1)
coeff
X = [-1, 0, 1, 2, 3, 4, 5, 6]
Y = [10, 9, 7, 5, 4, 3, 0, -1]
polyfit(X, Y, 1)
# +
p = plt.plot(x, noise_y, 'rx')
p = plt.plot(x, coeff[0] * x + coeff[1], 'k-')
p = plt.plot(x, y, 'b--')
# -
# Lebih sederhana
f = poly1d(coeff)
p = plt.plot(x, noise_y, 'rx')
p = plt.plot(x, f(x))
print(f)
# +
tk = [0.2, 0.4, 0.6, 0.8, 1.0]
dk = [0.196, 0.785, 1.766, 3.14, 4.907]
dktk2 = [d*t**2 for d,t in zip(dk, tk)]
tk4 = [t**4 for t in tk]
A = sum(dktk2)/sum(tk4)
2*A
# + [markdown] slideshow={"slide_type": "slide"}
# ## Curve Fitting - Data Linearization Method for $y = C e^{Ax}$
# +
import numpy as np
from scipy.optimize import minimize
def f(x):
A, C = x[0], x[1] # agar mudah dimengerti
return (C-1.5)**2 + (C*np.exp(A) - 2.5)**2 + (C*np.exp(2*A) - 3.5)**2 + (C*np.exp(3*A) - 5.0)**2 + (C*np.exp(4*A) - 7.5)**2
# -
x0 = [1,1]
res = minimize(f, x0)
res.x
# + [markdown] slideshow={"slide_type": "slide"}
# <h3>End of Module</h3>
# <hr />
| MFDSNM-06-Code.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Building the model - manually
#
# While TPOT runs on my other PC, lets see what kind of results we can get by manually constructing a classifier.
# +
import pandas as pd
from tqdm import tqdm_notebook
data_dir = "../data/2018-08-10_AV_Innoplexus/"
train_df = pd.read_csv(data_dir+'train_df_tfidf.csv')
train_df.head()
# +
#Encode tags
tags = train_df['Tag'].unique().tolist()
tags.sort()
tag_dict = {key: value for (key, value) in zip(tags,range(len(tags)))}
train_df['Tag_encoded'] = train_df['Tag'].map(tag_dict)
train_df_encoded = train_df.drop('Tag',axis=1)
# -
# # Which model? Why?
#
# So some good classic models for this type of task include all the typical characters;
#
# NaiveBayes, SVM, Random Forest, Logistic Regression, NeuralNet to name a few. I'll start with those with default settings and see how things go.
#
# Other strategies I might want to employ include;
#
# 1. Extraction of further words from the html
# 2. Over/Undersampling to achieve parity between classes in the training data.
# 3. Hyperparameter optimization
# +
from sklearn.linear_model import LogisticRegression
from sklearn.naive_bayes import GaussianNB
from sklearn.svm import SVC
from sklearn.ensemble import RandomForestClassifier, ExtraTreesClassifier
from sklearn.neural_network import MLPClassifier
logreg = LogisticRegression(penalty='l2',)
naiveB = GaussianNB()
svc = SVC()
rforest = RandomForestClassifier(n_estimators=20, n_jobs=2)
#From TPOT
etrees = ExtraTreesClassifier(bootstrap=True, criterion="entropy", max_features=0.7500000000000001, min_samples_leaf=3, min_samples_split=2, n_estimators=100, n_jobs=2)
neural = MLPClassifier()
model_names = ["LogReg","NaiveB","SVC","RForest","ETrees","Neural"]
# +
from sklearn.model_selection import cross_val_score, train_test_split
from sklearn.metrics import f1_score, confusion_matrix
x_cols = [x for x in train_df_encoded.columns if x != "Tag_encoded"]
X_train, X_test, y_train, y_test = train_test_split(train_df_encoded, train_df_encoded['Tag_encoded'], test_size=0.33)
# +
fit_models = []
model_preds = []
cross_val_scores = []
f1_scores = []
conf_matrices = []
for name, model in tqdm_notebook(zip(model_names,[logreg, naiveB, svc, rforest, etrees, neural])):
print("Working on ",name,"...")
cross_score = cross_val_score(model, X_train, y_train, cv=5, n_jobs=2)
model.fit(X_train, y_train)
fit_models.append(model)
preds = model.predict(X_test)
model_preds.append(preds)
model_f1 = f1_score(y_test, preds, average='micro')
model_conf = confusion_matrix(y_test, preds)
print("Done!")
cross_val_scores.append(cross_score)
f1_scores.append(model_f1)
conf_matrices.append(model_conf)
# -
import plotly.plotly as py
import plotly.graph_objs as go
from plotly.offline import init_notebook_mode, iplot
init_notebook_mode(True)
# +
#Plot cv_scores
cv_scores_data = go.Bar(
x = model_names,
y = [x.mean() for x in cross_val_scores],
name='Cross Validation Score'
)
f1_scores_data = go.Bar(
x = model_names,
y = [x.tolist() for x in f1_scores],
name="F1-micro Score"
)
cv_scores_layout = go.Layout(
title="Cross Validation & F1-micro Scores"
)
cv_scores_fig = go.Figure(
data=[cv_scores_data, f1_scores_data],
layout=cv_scores_layout
)
iplot(cv_scores_fig)
# -
# ## Lets look at the confusion matrices.
# +
import itertools
import numpy as np
import matplotlib.pyplot as plt
def plot_confusion_matrix(cm, classes, name,
normalize=False,
cmap=plt.cm.Blues):
"""
This function prints and plots the confusion matrix.
Normalization can be applied by setting `normalize=True`.
"""
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
print("Normalized confusion matrix")
else:
print('Confusion matrix, without normalization')
title= str(name+' Confusion matrix'),
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45)
plt.yticks(tick_marks, classes)
fmt = '.2f' if normalize else 'd'
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, format(cm[i, j], fmt),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
plt.show()
# -
for m_name, conf_matrix in zip(model_names, conf_matrices):
print(m_name)
plot_confusion_matrix(conf_matrix, classes=tag_dict.keys(), name=m_name,
normalize=True)
# Huh, something doesn't seem right - are they really that good? I'm going to go ahead and try to generate a first answer just to get something into the competition.
# Because extended trees keeps getting supported by my TPOT instance (that's still running) - I'm going to build my first submission with that model.
| 2018-08-10_AV_Innoplexus/05. Model Building.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="XzVNaHDCCnty"
# Notebook inicial
# + executionInfo={"elapsed": 493, "status": "ok", "timestamp": 1634370923368, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj9dCF4KCwTZ8Ez_yH2VH32AcPXn41BgIz0ue8PxoA=s64", "userId": "10603174789748071316"}, "user_tz": -120} id="W8CBDLnIFrbE"
# + executionInfo={"elapsed": 9, "status": "ok", "timestamp": 1634370923369, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj9dCF4KCwTZ8Ez_yH2VH32AcPXn41BgIz0ue8PxoA=s64", "userId": "10603174789748071316"}, "user_tz": -120} id="eQrfcmhhCqip"
import datetime
import time
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 10223, "status": "ok", "timestamp": 1634370933585, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj9dCF4KCwTZ8Ez_yH2VH32AcPXn41BgIz0ue8PxoA=s64", "userId": "10603174789748071316"}, "user_tz": -120} id="fa9AdrpECsfn" outputId="549416a6-c2a5-47c5-823e-fca48536f18a"
i = 0
while i < 10:
now = datetime.datetime.now()
print(now.year, now.month, now.day, now.hour+2, now.minute, now.second)
time.sleep(1)
i = i + 1
| Fundamentos/Notebooks/Notebooks_Prueba.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Python for Scientific Computing
# -
#
# #### <NAME> <br /> <br /> Ecology and Evolutionary Biology <br /> <br /> <EMAIL> <br /> <br /> @QuentinCAUDRON
# + [markdown] slideshow={"slide_type": "slide"}
# # Princeton University Python Community
# -
# ### **`princetonpy.com`** <br /> <br /> @PrincetonPy
# - Discussion Forums
# - Events Calendar
# - All past events
# - All code and slides
# + [markdown] slideshow={"slide_type": "subslide"}
# <img src="depts.png" width=800px />
# + [markdown] slideshow={"slide_type": "slide"}
# ## Today's Workshop
# -
# **Slides**
#
# - Slides are available on **`princetonpy.com/course`**
# - All code is provided as static or dynamic Notebooks
# - Notes and slides are designed to be read later, so may be wordy at times
# - ( by the way ) **they're entirely written in Python**
# <br />
# <br />
#
# **Session**
#
# - Interrupt, shout, scream, hurl something if you need
# <br />
# <br />
#
# **Aims :**
#
# - Provide a very quick refresher / introduction to Python
# - Demonstrate Python's core scientific stack
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Helping me out...
#
# - **<NAME>**, postdoc, Geosciences
# - **<NAME>**, Linux sysadmin, MolBio
#
# Thanks !
# + [markdown] slideshow={"slide_type": "subslide"}
# ### A note on Python 2 and Python 3
#
# Python 2 is still predominantly used in most academic circles.
#
# Python 3 came out in 2008, but isn't completely backwards-compatible, so it's been slow to adopt.
#
# There are **very few** differences for our purposes. All code presented here is compatible with either version.
# + [markdown] slideshow={"slide_type": "subslide"}
# Not everything said today will interest everyone. Not all aspects of "scientific computing" interest everybody.
#
# If you're interested in something specific, **please ask**.
# + [markdown] slideshow={"slide_type": "slide"}
# ## What is Python ?
# -
# - general-purpose
# - object-oriented
# - dynamically typed
# - interpreted
#
# It has a thriving community of developers, especially in science.
# <br />
# **The Zen of Python**
#
# - Beautiful is better than ugly.
# - Explicit is better than implicit.
# - Simple is better than complex.
# - Complex is better than complicated.
# - Readability counts.
# + [markdown] slideshow={"slide_type": "skip"}
# ## Context
# + [markdown] slideshow={"slide_type": "skip"}
# **C, C++, and Fortran**
#
# These languages are very fast, and great for heavy computations. However, they're slow and painful to write - there's no interactivity, the syntax gets complicated, and have manual memory management.
# + [markdown] slideshow={"slide_type": "skip"}
# **R**
#
# A tool for advanced statistics, but the language is exactly that : a *tool* aimed at stats. It's not very good for general-purpose coding. I have a strange aversion to it. Hey, at least it's free and open-source.
# + [markdown] slideshow={"slide_type": "skip"}
# **Matlab and Octave**
#
# Matlab has a great development environment, and a huge number of optimised, implemented toolsets. It's very expensive though. Octave is a great free clone, but it's not as pleasant to use.
# + [markdown] slideshow={"slide_type": "skip"}
# **So, Python ?**
#
# Huge range of scientific tools - nonlinear function fitting, MCMC, spectral analysis, ODEs and PDEs, signal and image processing, great data science tools. Vast community, active development, and very high quality due to the way the language is developed and the way we code it. It's **batteries-included**. Downsides are that the IDE isn't as shiny as Matlab, but I'm well over it - IPython Notebooks ( we'll see later ) are awesome.
# + [markdown] slideshow={"slide_type": "slide"}
# ## The Scientific Python Stack
# + [markdown] slideshow={"slide_type": "skip"}
# Python's standard library is huge. Still, as scientists, we require some fairly specific things that pure programmers might not immediately need : reasonable vector notation and manipulation, matrix and linear algebra, optimisation, interpolation, random numbers and statistical functions, plotting, etc.. The "standard Python stack" puts together a few modules and extensions to Python to give us these.
# -
# - **Numpy** : arrays, matrices and their operations, random numbers, ...
# - **Scipy** : linear algebra, symbolic operations, signal tools, optimisation, ...
# - **Matplotlib** : plotting
# - **Pandas** : data analysis and manipulation
# <br />
#
# Also very awesome, not covered today :
# - **IPython** : interactivity, Notebooks ( like this one ), ...
# - **Sympy** : symbolic mathematics
# + [markdown] slideshow={"slide_type": "skip"}
# Then, you may need more specific tools, like a good MCMC sampler ( PyMC ), or constrained, nonlinear function fitting ( lmfit ), or machine learning algorithms ( scikit-learn ), image processing ( scikit-image ), etc.. The list goes on !
# -
# In this workshop, we'll get set up with the basic Python stack and explore how they work. Then we'll demo some other packages for fun.
# + [markdown] slideshow={"slide_type": "skip"}
# ## Recommended Installs - Distribution
# + [markdown] slideshow={"slide_type": "skip"}
# I find that **Anaconda** is a great distribution. It comes will most of the packages you'll need, and a great command-line package manager to help keep them up to date and install others. If you want to grab the faster distro ( free for academics ), head to **`store.continuum.io/cshop/academicanaconda`** and register with an @blah**.edu** address to grab an academic license. If you can't be bothered with that, it's at **`continuum.com/downloads`**.
#
# 1. Linux : just run the .sh
# 2. OSX : just run the .dmg. Might need to select "install for me only" if you don't have admin rights.
# 3. Windows : just run the .exe
#
# If you don't want Anaconda, you can install Python on its own, and then add packages and modules as you need them. I won't be covering this in the interest of time. If you're under Linux, use your package manager; if you're under OSX, then HomeBrew has what you need. If you're running Windows and are interested in installing everything yourself, cry a little and then head to **`python.org/getit`**.
#
# A note on what we're installing : Anaconda comes with Python 2.7. This is by far the most widespread version of Python. It goes by Python 2 for short. There has been, for years, a Python 3, but it isn't backwards compatible, and whilst many of the main scientific packages are working to fix that, then the vast majority of scientists use Python 2 ( actually, just about everyone : the Python dudes themselves recommend starting with 2.7, due to compatibility; this is changing however, as more packages move towards Py3 compatibility ).
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Recommended Installs - Frontend
# + [markdown] slideshow={"slide_type": "skip"}
# **Front-End / Interface**
#
# Here, we have several options. With Python now installed, you just need an environment to write it in. You could use a standard text editor. I recommend **Atom**, for all platforms.
#
# Then, you could go with something more advanced, like an IDE ( integrated development environment ). An IDE is a one-stop shop to write and run your code. For Python, I like Spyder. It's available for all three of the above OSs. If you've installed Anaconda, you already have **Spyder**. Linux and OSX users, call `spyder` from the command line. Windows users can call the Anaconda Launcher, and you'll have it there.
#
# Finally, there's my favourite way to write Python *for development* : the **IPython Notebook**. If you installed Anaconda, you already have IPython. Otherwise, go get it, you won't regret it. The IPython Notebook concept will be familiar to you if you've used Mathematica, and some aspects of Matlab ( though in Matlab, it's not done so well ). You have *cells* in which you write code, and you can execute cells independently. With a quick command-line hack, you can even get plots inline, such that all plots show up under the relevant cells. To call IPython Notebooks, OSX and Linux users can just call `ipython notebook` from the command line, and Windows users with Anaconda have an IPython Notebook shortcut in their Start Menu ( in theory ). For inline plots and sexiness all around, I prefer calling `ipython notebook --script`. Here, `--script` tells IPython to also save a `.py` as well as the `.ipynb` extension, so you can just run your code from the command line or on a remote computer if you want to. Linux and OSX users can write this as an alias if they want : drop the line
#
# `alias pynb='ipython notebook --script`
#
# in `~/.bashrc`, and Windows users can edit the shortcut to their IPython Notebook to get the same result.
# -
# Take the time to consider your workflow and select an option.
#
# - **IPython Notebook** for code / algorithm development
# - **Spyder** if you prefer an integrated development environment *a la* R-Studio or Matlab
# - **IPython / Python terminal** for old-school play
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Summary
# -
# 1. Lightning refresher of Python syntax
# 2. Numpy
# 3. Pandas
# 4. Matplotlib
# 5. Scipy
# 6. Demos
#
# We'll be doing a few exercises throughout.
# + [markdown] slideshow={"slide_type": "fragment"}
# ## Grab these slides and the data from `princetonpy.com/course` to follow along.
| 0.Intro-and-Setup.ipynb |