code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Question 1: Given some sample data, write a program to answer the following: click here to access the required data set https://docs.google.com/spreadsheets/d/16i38oonuX1y1g7C_UAmiK9GkY7cS-64DfiDMNiR41LM/edit#gid=0
#
# On Shopify, we have exactly 100 sneaker shops, and each of these shops sells only one model of shoe. We want to do some analysis of the average order value (AOV). When we look at orders data over a 30 day window, we naively calculate an AOV of $3145.13. Given that we know these shops are selling sneakers, a relatively affordable item, something seems wrong with our analysis.
#
# a.Think about what could be going wrong with our calculation. Think about a better way to evaluate this data.
# b.What metric would you report for this dataset?
# c.What is its value?
#
#
import pandas as pd
url = "./data/2019 Winter Data Science Intern Challenge Data Set - Sheet1.csv"
df = pd.read_csv(url)
df.shape
df.head()
df.describe()
df['shop_id'].value_counts()
# +
import seaborn as sns
import matplotlib.pyplot as plt
df['order_amount'].plot(kind='box',figsize=(16,2),vert = False)
# -
df_outlier = df[df['order_amount'] > 200000]
df_outlier.head(17)
df_clean = df[df['order_amount'] < 200000]
df_clean.describe()
df_clean['order_amount'].plot(kind='box',figsize=(16,2),vert = False)
df_clean = df[df['order_amount'] < 20000]
df_clean['order_amount'].plot(kind='box',figsize=(16,2),vert = False)
df_clean.describe()
# +
SELECT count(orderid)
FROM orders as O
inner join shippers as S
on O.shipperid = S.ShipperID
where S.ShipperName == 'Speedy Express';
answer = 54
# +
SELECT O.employeeID, E.LastName, count(orderid) as tot_num
FROM orders as O
inner join employees as E
on O.employeeID = E.employeeID
group by O.employeeID
order by tot_num DESC
limit 1
LastName is Peacock
# +
SELECT * from customers
where Country ='Germany';
orders
-------
OrderID CustomerID EmployeeID OrderDate ShipperID
customers
---------
CustomerID
CustomerName
ContactName Address City PostalCode Country
products
--------
ProductID ProductName SupplierID CategoryID Unit Price
orderdetails
-----------
OrderDetailID OrderID ProductID Quantity
SELECT *, count(productID) as tot_num
from orderdetails
group by productID
order by tot_num DESC
limit 1
;
SELECT productID, sum(quantity) as tot_quant
from orderdetails
group by productID
order by tot_quant DESC
limit 1;
# +
SELECT OD.productid,count(OD.productid) as tot,productname,country from orders as O
inner join customers as C
on O.customerID = C.customerID
inner join orderdetails as OD
on O.orderid = OD.orderid
inner join products as P
on OD.productID = P.productID
where country = 'Germany'
group by OD.productid
order by tot DESC
limit 1;
| .ipynb_checkpoints/Dev_Intern-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Uniview module for of the asteroid explosion simulation
#
# ### From <NAME> from [this article](https://www.sciencedirect.com/science/article/pii/S001910351830349X?via%3Dihub).
#
#
# *<NAME>, 2019*
#
# ### Imports and function definitions
# +
#This directory contains all the data needed for the module. It should be in the same directory as the notebook
dataFolder = "data"
import sys, os, shutil, errno, string, urllib
sys.path.append(( os.path.abspath( os.path.join(os.path.realpath("__file__"), os.pardir, os.pardir) )))
import uvmodlib.v1 as uvmod
# -
# ### USES Conf Template
Template = """mesh
{
data asteroid ./modules/$folderName/asteroid.raw
dataHints asteroid disablePostprocessing
data quad ./modules/$folderName/quad.3ds
cullRadius $cr
glslVersion 330
propertyCollection
{
__objectName__
{
vec1f pSize 300 | public | desc "point size " | widget slider | range 0 1000
vec1f eventTime 0 | public | desc "time " | widget slider | range 0 130000
vec1f dMin 0.125 | public | desc "minimum damage"
vec1f dMax 0.23 | public | desc "maximum damage"
vec1f alpha 1.0 | public | desc "alpha" | widget slider | range 0 1
vec1f transitionLength 10 | public | desc "transition length in seconds"
bool jump true | public | desc "jump to time without transition"
}
}
############# to hold the time information
renderTexture
{
name stateTexture
width 1
height 1
numTextures 1
isPingPong true
isPersistent true
isFramePersistent true
internalTextureFormat GL_RGB32F
magnify GL_NEAREST
minify GL_NEAREST
}
############# set Transition State
pass
{
useDataObject quad
renderTarget
{
name stateTexture
enableColorClear false
}
shader
{
type defaultMeshShader
{
vertexShader ./modules/$folderName/pass0.vs
fragmentShader ./modules/$folderName/state.fs
textureFBO stateTexture stateTexture
stateManagerVar __objectName__.transitionLength transitionLength
stateManagerVar __objectName__.jump jump
stateManagerVar __objectName__.eventTime eventTime
parameter2f timeRange 0 133000
}
}
}
############# asteroid
pass
{
useDataObject asteroid
shader
{
type defaultMeshShader
{
geometryShader ./modules/$folderName/asteroid.gs
vertexShader ./modules/$folderName/asteroid.vs
fragmentShader ./modules/$folderName/asteroid.fs
textureFBO stateTexture stateTexture
texture cmap ./modules/$folderName/cmap.png
{
wrapModeS GL_CLAMP_TO_EDGE
wrapModeR GL_CLAMP_TO_EDGE
#colorspace linear
}
stateManagerVar __objectName__.pSize pSize
stateManagerVar __objectName__.dMin dMin
stateManagerVar __objectName__.dMax dMax
stateManagerVar __objectName__.alpha alpha
glState
{
UV_CULL_FACE_ENABLE false
UV_BLEND_ENABLE true
UV_DEPTH_ENABLE true
UV_WRITE_MASK_DEPTH true
UV_BLEND_FUNC GL_SRC_ALPHA GL_ONE_MINUS_SRC_ALPHA
}
}
}
}
}"""
# ### Asteroid class
class asteroid():
def __init__(self, object):
self.object = object
uvmod.Utility.ensurerelativepathexsists("asteroid.gs",dataFolder)
uvmod.Utility.ensurerelativepathexsists("asteroid.vs",dataFolder)
uvmod.Utility.ensurerelativepathexsists("asteroid.fs",dataFolder)
self.cr = 1000
self.Scale = 1
def generatemod(self):
self.object.setgeometry(self.object.name+"Mesh.usesconf")
return self.object.generatemod()
def generatefiles(self, absOutDir, relOutDir):
fileName = self.object.name+"Mesh.usesconf"
s = string.Template(Template)
f = open(absOutDir+"\\"+fileName, 'w')
if f:
f.write(s.substitute(folderName = relOutDir,
cr = self.cr,
Scale = self.Scale
))
f.close()
uvmod.Utility.copyfoldercontents(os.getcwd()+"\\"+dataFolder, absOutDir)
# ### Object Instantiation
# +
model = asteroid(uvmod.OrbitalObject())
generator = uvmod.Generator()
scene = uvmod.Scene()
scene.setname("AsteroidExplosion")
scene.setparent("Earth")
scene.setentrydist(100000) #not sure what this does
scene.setunit(1000)
scene.setstaticposition(-1e7,0,0)
modinfo = uvmod.ModuleInformation()
# -
# ### Specify Settings and generate the module
# +
model.object.setcameraradius(1) #I think smaller values here mean less clipping
model.object.setcoord(scene.name)
model.object.setname("AsteroidExplosion")
model.object.setguiname("/Adler/Universe Update/Asteroid Explosion")
model.object.settargetradius(100) #this is the location where the camera will fly to
model.object.showatstartup(False)
model.cr = 100 #I think this is clipping on the back end?
modinfo.setname("Asteroid Explosion")
modinfo.setauthor("<NAME><sup>1</sup> and <NAME><sup>2</sup>\
<br />(1) Adler Planetarium,<br />(2)Johns Hopkins University")
modinfo.cleardependencies()
modinfo.setdesc("Uniview module for the asteroid explosion simulation from <NAME>, et al. \
in this article : https://www.sciencedirect.com/science/article/pii/S001910351830349X?via%3Dihub")
#modinfo.setthumbnail("data/R0010133.JPG")
modinfo.setversion("1.0")
generator.generate("AsteroidExplosion",[scene],[model],modinfo)
uvmod.Utility.senduvcommand(model.object.name+".reload")
# -
# ## Helper Functions for modifing code
# *Reload Module and Shaders in Uniview*
uvmod.Utility.senduvcommand(model.object.name+".reload; system.reloadallshaders")
# *Copy modified Shader files and reload*
from config import Settings
uvmod.Utility.copyfoldercontents(os.getcwd()+"\\"+dataFolder, Settings.uvcustommodulelocation+'\\'+model.object.name)
uvmod.Utility.senduvcommand(model.object.name+".reload")
# ### Create colormap texture
import numpy as np
import matplotlib.pyplot as plt
from matplotlib import cm
# %matplotlib inline
# +
gradient = np.linspace(0, 1, 256)
gradient = np.vstack((gradient, gradient))
def plot_cmap(colormap):
fig=plt.imshow(gradient, aspect=1, cmap=colormap)
plt.axis('off')
fig.axes.get_xaxis().set_visible(False)
fig.axes.get_yaxis().set_visible(False)
plt.savefig("data/cmap.png", bbox_inches='tight',pad_inches=0)
plot_cmap('YlOrBr_r')
# -
| generate.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: SageMath 9.2
# language: sagemath
# metadata:
# cocalc:
# description: Open-source mathematical software system
# priority: 10
# url: https://www.sagemath.org/
# name: sage-9.2
# resource_dir: /ext/jupyter/kernels/sage-9.2
# ---
# +
n(pi, digits=4)
1+10
1/10 + 2/10 == 3/10
0.1 + 0.2 == 0.3
pi + 4*pi
show(x+x^2)
# -
show(x*(1+x))
100 // 7
100 % 7
sqrt(100)
factor(1024)
show(factor(1001))
divisors(1001)
vector([1,2,3])
| 20210407/materials/0-recap.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/tomfox1/DS-Unit-1-Sprint-2-Data-Wrangling/blob/master/DS_Unit_1_Sprint_Challenge_2_Data_Wrangling.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="4yMHi_PX9hEz" colab_type="text"
# # Data Science Unit 1 Sprint Challenge 2
#
# ## Data Wrangling
#
# In this Sprint Challenge you will use data from [Gapminder](https://www.gapminder.org/about-gapminder/), a Swedish non-profit co-founded by <NAME>. "Gapminder produces free teaching resources making the world understandable based on reliable statistics."
# - [Cell phones (total), by country and year](https://raw.githubusercontent.com/open-numbers/ddf--gapminder--systema_globalis/master/ddf--datapoints--cell_phones_total--by--geo--time.csv)
# - [Population (total), by country and year](https://raw.githubusercontent.com/open-numbers/ddf--gapminder--systema_globalis/master/ddf--datapoints--population_total--by--geo--time.csv)
# - [Geo country codes](https://github.com/open-numbers/ddf--gapminder--systema_globalis/blob/master/ddf--entities--geo--country.csv)
#
# These two links have everything you need to successfully complete the Sprint Challenge!
# - [Pandas documentation: Working with Text Data](https://pandas.pydata.org/pandas-docs/stable/text.html]) (one question)
# - [Pandas Cheat Sheet](https://github.com/pandas-dev/pandas/blob/master/doc/cheatsheet/Pandas_Cheat_Sheet.pdf) (everything else)
# + [markdown] id="wWEU2GemX68A" colab_type="text"
# ## Part 0. Load data
#
# You don't need to add or change anything here. Just run this cell and it loads the data for you, into three dataframes.
# + id="bxKtSi5sRQOl" colab_type="code" colab={}
import pandas as pd
import numpy as np
cell_phones = pd.read_csv('https://raw.githubusercontent.com/open-numbers/ddf--gapminder--systema_globalis/master/ddf--datapoints--cell_phones_total--by--geo--time.csv')
population = pd.read_csv('https://raw.githubusercontent.com/open-numbers/ddf--gapminder--systema_globalis/master/ddf--datapoints--population_total--by--geo--time.csv')
geo_country_codes = (pd.read_csv('https://raw.githubusercontent.com/open-numbers/ddf--gapminder--systema_globalis/master/ddf--entities--geo--country.csv')
.rename(columns={'country': 'geo', 'name': 'country'}))
# + [markdown] id="AZmVTeCsX9RC" colab_type="text"
# ## Part 1. Join data
# + [markdown] id="GLzX58u4SfEy" colab_type="text"
# First, join the `cell_phones` and `population` dataframes (with an inner join on `geo` and `time`).
#
# The resulting dataframe's shape should be: (8590, 4)
# + id="GVV7Hnj4SXBa" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="50a37d00-405e-4cb5-b4c1-ef31ea1f1c2e"
#merging data
phone_pop = pd.merge(cell_phones, population, how = "inner", on = ["geo", "time"])
phone_pop.shape
# + [markdown] id="xsXpDbwwW241" colab_type="text"
# Then, select the `geo` and `country` columns from the `geo_country_codes` dataframe, and join with your population and cell phone data.
#
# The resulting dataframe's shape should be: (8590, 5)
# + id="Q2LaZta_W2CE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="27876c50-72f5-4ea7-ea33-ba8ba24c5359"
#merging data
geo_pop = geo_country_codes[["geo", "country"]].merge(phone_pop)
geo_pop.shape
# + [markdown] id="oK96Uj7vYjFX" colab_type="text"
# ## Part 2. Make features
# + [markdown] id="AD2fBNrOYzCG" colab_type="text"
# Calculate the number of cell phones per person, and add this column onto your dataframe.
#
# (You've calculated correctly if you get 1.220 cell phones per person in the United States in 2017.)
# + id="wXI9nQthYnFK" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 80} outputId="02bbce8c-1a67-47dc-98c7-53bce64037bf"
#easy way to calculate cell phone per person
geo_pop["cell phones per person"] = geo_pop["cell_phones_total"] / geo_pop["population_total"]
geo_pop[geo_pop["country"] == "United States"].tail(1)
# + [markdown] id="S3QFdsnRZMH6" colab_type="text"
# Modify the `geo` column to make the geo codes uppercase instead of lowercase.
# + id="93ADij8_YkOq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="c01abb3e-7074-46f6-ae1f-05fee35f154f"
#converting our lowercase strings in "geo" to uppercase
geo_pop["geo"] = geo_pop["geo"].str.upper()
geo_pop.head(5)
# + [markdown] id="hlPDAFCfaF6C" colab_type="text"
# ## Part 3. Process data
# + [markdown] id="k-pudNWve2SQ" colab_type="text"
# Use the describe function, to describe your dataframe's numeric columns, and then its non-numeric columns.
#
# (You'll see the time period ranges from 1960 to 2017, and there are 195 unique countries represented.)
# + id="g26yemKre2Cu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 390} outputId="012dc4e5-f139-45b5-86b6-ab5b5aefa04a"
#first describing all data
geo_pop.describe(include = "all")
# + id="NH_4Df-fIoq8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 173} outputId="98edf7be-9b6f-49c4-a4c8-7e549d5a4b96"
#describing data excluding numeric columns
geo_pop.describe(exclude = [np.number])
# + [markdown] id="zALg-RrYaLcI" colab_type="text"
# In 2017, what were the top 5 countries with the most cell phones total?
#
# Your list of countries should have these totals:
#
# | country | cell phones total |
# |:-------:|:-----------------:|
# | ? | 1,474,097,000 |
# | ? | 1,168,902,277 |
# | ? | 458,923,202 |
# | ? | 395,881,000 |
# | ? | 236,488,548 |
#
#
# + id="JdlWvezHaZxD" colab_type="code" colab={}
# This optional code formats float numbers with comma separators
pd.options.display.float_format = '{:,}'.format
# + id="tmcikV9PTJfz" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 136} outputId="8e2f8a90-19fe-4f72-bde3-15bff6d4ac04"
#it is important to create a condition for the year 2017 before using the groupby function to sort for "cell_phones_total"
condition = (geo_pop["time"] == 2017)
geo_2017 = geo_pop[condition]
geo_2017.groupby(["country"])["cell_phones_total"].sum().sort_values(ascending = False).head()
# + [markdown] id="03V3Wln_h0dj" colab_type="text"
# 2017 was the first year that China had more cell phones than people.
#
# What was the first year that the USA had more cell phones than people?
# + id="k-tENcTgYkey" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 80} outputId="87e1597f-dcb0-463a-e222-d30c45935536"
#The year where number of cell phone surpasses people is the year where number of phones per person >= 1
#we create a condition to select the US as our country of choice
#after creating the condition we for values >= 1 and choose the first output
#since that's the first year (2014) where total cell phones surpassed population
countries = ["United States"]
condition = geo_pop['country'].isin(countries)
us = geo_pop[condition]
us[us["cell phones per person"] >= 1].head(1)
# + [markdown] id="6J7iwMnTg8KZ" colab_type="text"
# ## Part 4. Reshape data
# + [markdown] id="LP9InazRkUxG" colab_type="text"
# Create a pivot table:
# - Columns: Years 2007—2017
# - Rows: China, India, United States, Indonesia, Brazil (order doesn't matter)
# - Values: Cell Phones Total
#
# The table's shape should be: (5, 11)
# + id="JD7mXXjLj4Ue" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="5122b07d-2fe3-43e5-9ec4-e2d602114dac"
#creating 2 conditions to reduce the scope of our data to our specifications
years = [2007, 2008, 2009, 2010, 2011, 2012, 2013, 2014, 2015, 2016, 2017]
countries = ["China", "India", "United States", "Indonesia", "Brazil"]
condition = geo_pop['time'].isin(years)
condition2 = geo_pop['country'].isin(countries)
subset = geo_pop[condition]
subset = subset[condition2]
# + id="WazA_t9eQLvv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 252} outputId="d55f3912-b27c-483e-861e-8971ca340432"
#our pivot table looks right
sub_pivot = subset.pivot_table(index = "country", columns = "time", values = "cell_phones_total")
sub_pivot
# + id="5_yhF3UySR8J" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="fd24e1d8-5f11-46b8-92c9-91584fba4275"
#passsing shape to confirm our findings
sub_pivot.shape
# + [markdown] id="CNKTu2DCnAo6" colab_type="text"
# #### OPTIONAL BONUS QUESTION!
#
# Sort these 5 countries, by biggest increase in cell phones from 2007 to 2017.
#
# Which country had 935,282,277 more cell phones in 2017 versus 2007?
# + [markdown] id="7iHkMsa3Rorh" colab_type="text"
# If you have the time and curiosity, what other questions can you ask and answer with this data?
# + id="JZNb_t0TgmBz" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="55bdb71d-3be0-43ab-d31d-01e0fb6aa254"
#note: attempted to do bonus question, however, not finished
years1 = [2007, 2017]
countries1 = ["China", "India", "United States", "Indonesia", "Brazil"]
new_condition = geo_pop['time'].isin(years1)
new_condition2 = geo_pop['country'].isin(countries1)
new_subset = geo_pop[new_condition]
new_subset = new_subset[new_condition2]
# + id="9Cqt0Yd9hA1h" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 359} outputId="2b452bfe-7809-4450-834c-2e99af3ad60f"
#conceptually we would calculate percentage differences in total phones from 2017 and 2007 and then create a new column to sort in ascending order
new_subset
# + id="4wzZr3IphCCN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 142} outputId="67f7a5ce-865e-4c92-f593-f1c443ef471f"
#We would need to calculate 5 values, it could be done by hand and then inputed, however, that is not the task at hand
#with more time I would have enjoyed trying to calculate programatically
new_sub_pivot = new_subset.pivot_table(index = "time", columns = "country", values = "cell_phones_total")
new_sub_pivot
# + id="MLibQZ27kDlg" colab_type="code" colab={}
| DS_Unit_1_Sprint_Challenge_2_Data_Wrangling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import torch
import os
import pickle
import matplotlib.pyplot as plt
# # %matplotlib inline
# plt.rcParams['figure.figsize'] = (20, 20)
# plt.rcParams['image.interpolation'] = 'bilinear'
import sys
sys.path.append('../train/')
from torch.autograd import Variable
from torch.utils.data import DataLoader, Dataset
import torchvision.datasets as datasets
import torchvision
import torchvision.transforms as T
import torch.nn.functional as F
import torch.nn as nn
import collections
import numbers
import random
import math
from PIL import Image, ImageOps, ImageEnhance
import time
from torch.utils.data import Dataset
from networks.UNet_standard import UNet
import tool
from tqdm import tqdm
flip_index = ['16', '15', '14', '13', '12', '11', '10']
# -
NUM_CHANNELS = 3
NUM_CLASSES = 2
BATCH_SIZE = 8
W, H = 1918, 1280
STRIDE = 256
IMAGE_SIZE = 512
test_mask_path = '../../data/test_masks/UNet_same/'
weight_path = '../_weights/UNet_same-fold0-0.00555.pth'
def load_model(filename, model):
checkpoint = torch.load(filename)
model.load_state_dict(checkpoint['model_state'])
model = UNet(in_channels=NUM_CHANNELS, n_classes=NUM_CLASSES, padding=True)
model = model.cuda()
model.eval()
load_model(weight_path, model)
# +
test_path = '../../data/images/test/'
if not os.path.exists(test_mask_path):
os.makedirs(test_mask_path)
# -
test_names = os.listdir(test_path)
test_names = sorted(test_names)
with torch.no_grad():
batch_size = BATCH_SIZE
normalize_mean = [.485, .456, .406]
normalize_std = [.229, .224, .225]
test_names = sorted(os.listdir(test_path))
for image_pack in tqdm(range(len(test_names) // batch_size)):
images = np.zeros((batch_size, 3, H, W), dtype='float32')
test_masks = np.zeros((batch_size, 2, H, W), dtype='float32')
ifflip = [False] * batch_size
image_batch_names = test_names[image_pack * batch_size: image_pack * batch_size + batch_size]
mask_names = [input_name.split('.')[0] + '.png' for input_name in image_batch_names]
for idx, image_name in enumerate(image_batch_names):
image = Image.open(os.path.join(test_path, image_name))
angle = image_name.split('.')[0].split('_')[-1]
if angle in flip_index:
ifflip[idx] = True
image = ImageOps.mirror(image)
image = np.array(image).astype('float') / 255
image = image.transpose(2, 0, 1)
for i in range(3):
image[i] = (image[i] - normalize_mean[i]) / normalize_std[i]
images[idx] = image
for h_idx in range(int(math.ceil((H - STRIDE) / STRIDE))):
h_start = h_idx * STRIDE
h_end = h_start + IMAGE_SIZE
if h_end > H:
h_end = H
h_start = h_end - IMAGE_SIZE
for w_idx in range(int(math.ceil((W - STRIDE) / STRIDE))):
w_start = w_idx * STRIDE
w_end = w_start + IMAGE_SIZE
if w_end > W:
w_end = W
w_start = w_end - IMAGE_SIZE
input_batchs = images[:, :, h_start:h_end, w_start:w_end]
input_tensor = torch.from_numpy(input_batchs).cuda()
inputs = Variable(input_tensor, )
outputs = model(inputs)
ouputs = outputs.cpu().data.numpy()
test_masks[:, :, h_start:h_end, w_start:w_end] += ouputs
test_masks = np.argmax(test_masks, axis=1).astype('uint8')
for idx in range(batch_size):
output_PIL = Image.fromarray(test_masks[idx].astype('uint8')*255).convert('1')
if ifflip[idx]:
output_PIL = ImageOps.mirror(output_PIL)
mask_name = mask_names[idx]
output_PIL.save(test_mask_path + mask_name)
| get_test_masks/get_test_mask_UNet_same.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # Segmentation
# If you have Unet, all CV is segmentation now.
# ## Goals
#
# - train Unet on isbi dataset
# - visualize the predictions
# # Preparation
# Get the [data](https://www.dropbox.com/s/0rvuae4mj6jn922/isbi.tar.gz) and unpack it to `catalyst-examples/data` folder:
# ```bash
# catalyst-examples/
# data/
# isbi/
# train-volume.tif
# train-labels.tif
# ```
# # Data
import os
os.environ["CUDA_VISIBLE_DEVICES"] = "0"
# +
# # ! pip install tifffile
# +
import tifffile as tiff
images = tiff.imread('./data/isbi/train-volume.tif')
masks = tiff.imread('./data/isbi/train-labels.tif')
data = list(zip(images, masks))
train_data = data[:-4]
valid_data = data[-4:]
# +
import collections
import numpy as np
import torch
import torchvision
import torchvision.transforms as transforms
from catalyst.data.augmentor import Augmentor
from catalyst.dl.utils import UtilsFactory
bs = 4
n_workers = 4
data_transform = transforms.Compose([
Augmentor(
dict_key="features",
augment_fn=lambda x: \
torch.from_numpy(x.copy().astype(np.float32) / 255.).unsqueeze_(0)),
Augmentor(
dict_key="features",
augment_fn=transforms.Normalize(
(0.5, 0.5, 0.5),
(0.5, 0.5, 0.5))),
Augmentor(
dict_key="targets",
augment_fn=lambda x: \
torch.from_numpy(x.copy().astype(np.float32) / 255.).unsqueeze_(0))
])
open_fn = lambda x: {"features": x[0], "targets": x[1]}
loaders = collections.OrderedDict()
train_loader = UtilsFactory.create_loader(
train_data,
open_fn=open_fn,
dict_transform=data_transform,
batch_size=bs,
workers=n_workers,
shuffle=True)
valid_loader = UtilsFactory.create_loader(
valid_data,
open_fn=open_fn,
dict_transform=data_transform,
batch_size=bs,
workers=n_workers,
shuffle=False)
loaders["train"] = train_loader
loaders["valid"] = valid_loader
# -
# # Model
from catalyst.contrib.models.segmentation import UNet
# # Model, criterion, optimizer
# +
import torch
import torch.nn as nn
model = UNet(num_classes=1, in_channels=1, num_filters=64, num_blocks=4)
criterion = nn.BCEWithLogitsLoss()
optimizer = torch.optim.Adam(model.parameters(), lr=1e-3)
# scheduler = None # for OneCycle usage
scheduler = torch.optim.lr_scheduler.MultiStepLR(optimizer, milestones=[10, 20, 40], gamma=0.3)
# scheduler = torch.optim.lr_scheduler.ReduceLROnPlateau(optimizer, factor=0.5, patience=2, verbose=True)
# -
# # Callbacks
# +
import collections
from catalyst.dl.callbacks import (
LossCallback,
Logger, TensorboardLogger,
OptimizerCallback, SchedulerCallback, CheckpointCallback,
PrecisionCallback, OneCycleLR)
n_epochs = 50
logdir = "./logs/segmentation_notebook"
callbacks = collections.OrderedDict()
callbacks["loss"] = LossCallback()
callbacks["optimizer"] = OptimizerCallback()
# OneCylce custom scheduler callback
callbacks["scheduler"] = OneCycleLR(
cycle_len=n_epochs,
div=3, cut_div=4, momentum_range=(0.95, 0.85))
# Pytorch scheduler callback
# callbacks["scheduler"] = SchedulerCallback(
# reduce_metric="loss_main")
callbacks["saver"] = CheckpointCallback()
callbacks["logger"] = Logger()
callbacks["tflogger"] = TensorboardLogger()
# -
# # Train
# +
from catalyst.dl.runner import SupervisedModelRunner
runner = SupervisedModelRunner(
model=model,
criterion=criterion,
optimizer=optimizer,
scheduler=scheduler)
runner.train(
loaders=loaders,
callbacks=callbacks,
logdir=logdir,
epochs=n_epochs, verbose=True)
# -
# # Inference
from catalyst.dl.callbacks import InferCallback
# +
callbacks = collections.OrderedDict()
callbacks["saver"] = CheckpointCallback(
resume=f"{logdir}/checkpoint.best.pth.tar")
callbacks["infer"] = InferCallback()
# +
loaders = collections.OrderedDict()
loaders["infer"] = UtilsFactory.create_loader(
valid_data,
open_fn=open_fn,
dict_transform=data_transform,
batch_size=bs,
workers=n_workers,
shuffle=False)
# -
runner.infer(
loaders=loaders,
callbacks=callbacks,
verbose=True)
# # Predictions visualization
import matplotlib.pyplot as plt
plt.style.use("ggplot")
# %matplotlib inline
# +
sigmoid = lambda x: 1/(1 + np.exp(-x))
for i, (input, output) in enumerate(zip(
valid_data, callbacks["infer"].predictions["logits"])):
image, mask = input
threshold = 0.5
plt.figure(figsize=(10,8))
plt.subplot(1, 3, 1)
plt.imshow(image, 'gray')
plt.subplot(1, 3, 2)
output = sigmoid(output[0].copy())
output = (output > threshold).astype(np.uint8)
plt.imshow(output, 'gray')
plt.subplot(1, 3, 3)
plt.imshow(mask, 'gray')
plt.show()
# -
| examples/segmentation-example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Classical AES used in TLS
# +
#https://stackoverflow.com/a/21928790
import base64
import hashlib
from Crypto import Random
from Crypto.Cipher import AES
class AESCipher(object):
def __init__(self, key):
self.bs = AES.block_size
self.key = hashlib.sha256(key.encode()).digest()
def encrypt(self, raw):
raw = self._pad(raw)
iv = Random.new().read(AES.block_size) #initialization vector/seed
cipher = AES.new(self.key, AES.MODE_CBC, iv)
return base64.b64encode(iv + cipher.encrypt(raw.encode()))
def decrypt(self, enc):
enc = base64.b64decode(enc)
iv = enc[:AES.block_size]
cipher = AES.new(self.key, AES.MODE_CBC, iv)
return self._unpad(cipher.decrypt(enc[AES.block_size:])).decode('utf-8')
def _pad(self, s):
return s + (self.bs - len(s) % self.bs) * chr(self.bs - len(s) % self.bs)
@staticmethod
def _unpad(s):
return s[:-ord(s[len(s)-1:])]
# -
# ## Quantum resistant key-exchange
#
# NewHope specification: https://newhopecrypto.org/data/NewHope_2020_04_10.pdf
#
# Key exchange (page 5): https://eprint.iacr.org/2015/1092.pdf
# +
#https://github.com/nakov/PyNewHope
from pynewhope import newhope
def convert_key(sharedKey):
return ''.join([chr(byte) for byte in sharedKey])
# -
#Alice side part: 1/2
alicePrivKey, b_and_seed = newhope.keygen()
#Bob side
bobSharedKey, u_and_r = newhope.sharedB(b_and_seed)
#Alice side part: 2/2
aliceSharedKey = newhope.sharedA(u_and_r, alicePrivKey)
# ## Comunication
#Alice side
state=AESCipher(convert_key(aliceSharedKey))
message="Hello quantum world!"
secret=state.encrypt(message)
print(secret)
#Bob side
state=AESCipher(convert_key(bobSharedKey))
message=state.decrypt(secret)
print(message)
| Zad6/Example_newhope.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/wfwiggins/rad-ml-tutor/blob/master/NIIC21_Image_Classification.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] _uuid="b71bad4ea3f995104509072243b7837bb4ce8463" id="GO_D2Ne5ebYC"
# # **NIIC-RAD 2021: AI Mini-Lab**
#
# ## **Basics of Image Classification**
# In this demonstration, we will utilize techniques of _computer vision_, including deep _convolutional neural networks_ (CNNs), to train an image classifier model capable of classifying radiographs as either **chest** or **abdominal**.
#
# ### Code
# We will utilize the [fast.ai v2 library](https://docs.fast.ai/), written primarily by <NAME> and <NAME> (with help from many others). It is written in the [Python programming language](https://www.python.org/) and built on top of the [PyTorch deep learning library](https://www.pytorch.org/).
#
# The demonstration in this notebook relies heavily on examples from the `fast.ai` book, _Deep Learning for Coders with fastai and PyTorch: AI Applications without a PhD_ by <NAME> and <NAME>, which was written entirely in Jupyter notebooks, which are [freely available for download on GitHub](https://github.com/fastai/fastbook). A print copy of the book can be purchased from Amazon.
#
# ### Data
# This work is adapted from "[Hello World Deep Learning in Medical Imaging](https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5959832/)" (full reference below). The chest and abdominal radiographs were obtained from [Paras Lakhani's GitHub repository](https://github.com/paras42/Hello_World_Deep_Learning/tree/9921a12c905c00a88898121d5dc538e3b524e520).
#
# > _Reference:_ <NAME>, <NAME>, <NAME>, <NAME>, <NAME>. Hello World Deep Learning in Medical Imaging. J Digit Imaging. 2018 Jun; 31(3):283-289. Published online 2018 May 3. doi: 10.1007/s10278-018-0779-6
#
# ### Developers
# - <NAME>, MD, PhD - Duke University Hospital, Durham, NC, USA
# - <NAME>, MD, PhD, - University of California, San Francisco, CA, USA
# - <NAME>, MD, PhD - University of California, San Francisco, CA, USA
#
# ### Acknowledgements
# Other versions of this notebook implemented on the [Kaggle Notebooks platform](https://www.kaggle.com/wfwiggins203/hello-world-for-deep-learning-siim) were presented at the 2019 Society for Imaging Informatics in Medicine (SIIM) Annual Meeting, for the American College of Radiology (ACR) Residents & Fellows Section (RFS) [AI Journal Club](https://www.acr.org/Member-Resources/rfs/Journal-Club), and for the 2020 Radiological Society of North America (RSNA) AI Refresher Course.
#
# We would also like to acknowledge the following individuals for inspiring our transition to the Google Colab platform with their excellent notebook from the 2019 RSNA AI Refresher Course:
# - <NAME>, MD, PhD
# - <NAME>, MD, MSc
# - <NAME>, MD
# - <NAME>, MD
#
# + [markdown] _uuid="512cb129098ec3d6147e66be3e3c014aa6aeb80e" id="BDdzgQIpebYI"
# # System Setup & Downloading the Data
#
# **_FIRST_:** Save a copy of this notebook in your Google Drive folder by selecting _Save a Copy in Drive_ from the _File_ menu in the top left corner of this page.
#
# **_NEXT_:** Make sure you have the _runtime type_ set to **"GPU"**.
#
# 
#
# + _uuid="0c1e51dbcbe223c29555c5188b0df55b10ed8b06" id="N2yr5s5_ebYI" cellView="form"
#@title **Setting up the runtime environment...**
#@markdown Running this cell will install the necessary libraries and download the data.
import os
# !pip install fastai==2.1.4 >/dev/null
# !pip install fastcore==1.3.1 >/dev/null
# **Downloading the data...**
# !wget -q https://github.com/wfwiggins/RSNA-Image-AI-2020/blob/master/data.zip?raw=true
# !mkdir -p data
# !unzip -o data.zip?raw=true -d data >/dev/null
# !rm data.zip?raw=true
# + [markdown] _uuid="018a4a808153bc354041f7dc012c7c04ad00807e" id="ov62fFTuebYP"
# # Exploring the Data
# Let's take a look at the directory structure and contents, then create some variables to help us as we proceed.
# + _uuid="4cfcb87c72a542aefce13f8453097c0d1eb0c7b9" id="3Wyx5pQJebYQ" cellView="form" colab={"base_uri": "https://localhost:8080/"} outputId="9b2aa34b-301d-4561-bf41-9adbae9e0de1"
#@title Directory structure
from fastai.basics import *
from fastai.vision.all import *
import warnings
warnings.simplefilter('ignore')
# Set path variable to the directory where the data is located
path = Path('/content/data')
# Command line "magic" command to show directory contents
# !ls {path}/**/*
# + [markdown] _uuid="e7103f1d034e49ccd219cf1190fc8358eca18b1d" id="S09vTIX4ebYT"
# As you can see, the `data` directory contains subdirectories `train`, `val` and `test`, which contain the *training*, *validation* and *test* data for our experiment. `train` and `val` contain subdirectories `abd` and `chest` containing abdominal and chest radiographs for each data set. There are 65 training images and 10 validation images with *balanced distributions* over our *target classes* (i.e. approximately equal numbers of abdominal and chest radiographs in each data set and optimized for a classification problem).
# + [markdown] id="JfsLnJMWjPiK"
# # Model Training Setup
#
# Before we train the model, we have to get the data in a format such that it can be presented to the model for training.
#
# ## Data Loaders
# The first step is to load the data for the training and validation datasets into a `ImageDataLoaders` object from the `fastai` library. When training a model, the `ImageDataLoaders` will present training - and subsequently, validation - data to the model in _batches_.
#
# ## Data Augmentation
# In order to be sure that the model isn't simply "memorizing" the training data, we will _augment_ the data by randomly applying different _transformations_ to each image before it is sent to the model.
#
# Transformations can include rotation, translation, flipping, rescaling, etc.
# + _uuid="d1c24e4a78f57f12a42de3481db746fe0f170ee3" id="E4S9uHACebYT" cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 373} outputId="001a87c8-84d3-48c0-ce95-afd5dff719b9"
#@title Load the data into `ImageDataLoaders` with data augmentation
#@markdown When you run this cell, a batch of data will be shown with or without augmentation transforms applied.
#@markdown > 1. Run this cell once with the box next to `apply_transforms` unchecked to see a sample of the original images.
#@markdown > 2. Next, run the cell a few more times after checking the box next to `apply_transforms` to see what happens to the images when the transforms are applied.
apply_transforms = True #@param {type: 'boolean'}
if apply_transforms:
flip = True
max_rotate = 10.0
max_warp = 0.2
p_affine = 0.75
else:
flip = False
max_rotate, max_warp, p_affine = 0, 0, 0
tfms = aug_transforms(
do_flip=flip,
max_rotate=max_rotate,
max_warp=max_warp,
p_affine=p_affine,
size=224,
min_scale=0.75
)
dls = ImageDataLoaders.from_folder(path, valid='val', seed=42, item_tfms=Resize(460), batch_tfms=tfms, bs=16)
dls.show_batch(max_n=6)
# + id="KHOsqIn_c5oe" cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 349, "referenced_widgets": ["b80bfc8778714eb3a20c071c5539ee35", "<KEY>", "9566f729f4114e0f9b769c97282b5ee3", "<KEY>", "9c2d9b2b50fd4ef7817f169d2c53366d", "270305c6fde3439091d91f593796d242", "c6499a88d87140199d37fa10206bdbd2", "cb5d338c3f124ae99438401a6d84e707"]} outputId="7ee44316-1221-4c03-a516-6f608df98e2c"
#@title Find the optimal learning rate
#@markdown The learning rate is a hyperparameter that controls how much your model adjusts in response to percieved error after each training epoch. Choosing an optimal learning rate is an optimal step in model training.
#@markdown From the `fastai` [docs](https://docs.fast.ai/callback.schedule#Learner.lr_find):
#@markdown > First introduced by <NAME> in [Cyclical Learning Rates for Training Neural Networks](https://arxiv.org/pdf/1506.01186.pdf), the `LRFinder` trains the model with exponentially growing learning rates and stops in case of divergence.
#@markdown > The losses are then plotted against the learning rates with a log scale. <br><br>
#@markdown > A good value for the learning rates is then either:
#@markdown > - 1/10th of the minimum before the divergence
#@markdown > - where the slope is the steepest
#@markdown When you run this cell for the first time in a session, it will download a pretrained version of the model to your workspace before running the `LRFinder`.
dls = ImageDataLoaders.from_folder(path, valid='val', seed=42, item_tfms=Resize(460), batch_tfms=aug_transforms(size=224, min_scale=0.75), bs=16)
learn = cnn_learner(dls, resnet18, metrics=accuracy)
learn.lr_find();
# + [markdown] id="LzeZ8DNmndBH"
# # Transfer Learning
#
# Deep learning requires large amounts of training data to successfully train a model.
#
# When we don't have enough data to work with for the planned task, starting with a _pre-trained_ network that has been optimally trained on another task can be helpful. The concept of re-training a pre-trained network for a different task is called _transfer learning_.
#
# ## Fine-tuning
#
# In the process of re-training the model, we start by changing the final layers of the network to define the output or predictions our model will make. In order to avoid propagating too much error through the rest of the network during the initial training, we freeze the other layers of the network for the first cycle or _epoch_ of training. Next, we open up the rest of the network for training and train for a few more _epochs_. This process is called _fine-tuning_.
#
# ## Epochs and data augmentation
#
# During each epoch, the model will be exposed to the entire dataset. Each batch of data will have our data transformations randomly applied in order to provide data augmentation. This helps to ensure that our model never sees the exact same image twice. This is important because we wouldn't want our model to simply memorize the training dataset and not converge on a generalized solution, resulting in poor performance on the validation dataset.
#
# ## The loss function
#
# In a classification task, you're either right or wrong. This binary information doesn't give us much nuance to work with when training a model. A _loss function_ give us a numeric estimation of "how wrong" our model is. This gives us a target to optimize during the training process.
#
# When reviewing the results of successive epochs in training, the loss on your validation dataset should always be **decreasing**. When it starts to increase, that is a sign of your model _overfitting_ to the training dataset.
# + id="HBFmaUmvesW8" cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 267} outputId="5cc93c83-4a9b-473c-ace5-e4fbb3a30273"
#@title Fine-tune the model
#@markdown First, choose the number of epochs for which you will train your model.
#@markdown Then, choose a base learning rate based on the results in the `LRFinder` plot above.
#@markdown Finally, run the cell to train the model.
#@markdown After you've seen the results of your experiment, you can re-run this cell with different hyperparameters to see how they affect the result.
epochs = 5 #@param {type: "integer"}
base_lr = 2e-3 #@param {type: "number"}
learn = cnn_learner(dls, resnet18, metrics=accuracy)
learn.fine_tune(epochs, base_lr=base_lr)
# + id="iHD1vnazhUvd" cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 265} outputId="001b4a1a-4fec-42bc-e16d-2f3eb8be5633"
#@title Review training curves
#@markdown The visual representation of the training and validation losses are useful to evaluate how successfully you were able to train your model.
learn.recorder.plot_loss()
# + [markdown] id="e0efexR3jcYw"
# # Testing the Model
# + id="CfrLxQyuh4NY" cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 214} outputId="e2261570-7134-4ae8-fc30-4cd99c26f45f"
#@title **Test the model on the test dataset**
#@markdown When you run this cell, the first line shows the groundtruth for whether the radiograph is of the chest or abdomen.
#@markdown The second line is the model prediction for whether the image ia a chest or abdominal radiograph.
# # !mkdir data/test/abd
# # !mkdir data/test/chst
# # !mv data/test/abd_test.png data/test/abd
# # !mv data/test/chest_test.png data/test/chst
test_files = get_image_files(path/'test')
test_dl = learn.dls.test_dl(test_files, with_labels=True)
learn.show_results(dl=test_dl)
# + id="IY2G68YIi8aK" cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 210} outputId="30de69e4-32b7-4ee3-96fd-543033d45989"
#@title **A little more detail on the predictions**
#@markdown Running this cell will provide us with the loss on each image, as well as the model's predicted probability, which can be thought of as the model's confidence in its prediction.
#@markdown If the model is completely confident, the loss will be "0.00" and the probability will be "1.00".
interp = ClassificationInterpretation.from_learner(learn, dl=test_dl)
interp.plot_top_losses(k=2)
# + id="JdPI1HS079ZI" cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 277} outputId="8f8ec09a-adc5-4ae4-ad46-35401b3f537e"
#@title **Test the model on a surprise example**
#@markdown Here, we present the model with an unexpected image and see how it responds.
y = get_image_files(path, recurse=False)
test_dl = learn.dls.test_dl(y)
x, = first(test_dl)
res = learn.get_preds(dl=test_dl, with_decoded=True)
x_dec = TensorImage(dls.train.decode((x,))[0][0])
fig, ax = plt.subplots()
fig.suptitle('Prediction / Probability', fontsize=14, fontweight='bold')
x_dec.show(ctx=ax)
ax.set_title(f'{dls.vocab[res[2][0]]} / {max(res[0][0]):.2f}');
# + [markdown] id="DpgwiGVEuzkO"
# When presented with this radiograph of an elbow, the model makes a prediction but is less confident than with the other test images.
#
# This is an important point to consider for two reasons:
# 1. A deep learning model can only learn what we teach it to learn
# 2. In designed our model implementation, we might consider designing a pre-processing step in which the data (or metadata) is checked to ensure the input to the model is valid. This is an important practical consider for AI applications in radiology.
# + [markdown] id="KoL98c9Yj9Sn"
# # Visualizing Model Inferences
# + id="z6E9OLdNU7zd" cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 248} outputId="15f11ce4-5b9a-4535-c2d3-403790498b0d"
#@title **Class activation map (CAM)**
#@markdown CAM allows one to visualize which regions of the original image are heavily weighted in the prediction of the corresponding class.
#@markdown This technique provides a visualization of the activations in the **final** _convolutional_ block of a Convolutional Neural Network (CNN).
#@markdown CAM is also useful to determine if the model is "cheating" and looking somewhere it shouldn't be to make its prediction (i.e. radioopaque markers placed by the technologist).
#@markdown > Choose which of the two test images you would like to examine and run this cell to see the CAM output overlayed on the input image.
test_case = 'abd' #@param ['abd', 'chest']
cls = 0 if test_case == 'abd' else 1
label = test_case
y = get_image_files(path/'test'/label)
test_dl = learn.dls.test_dl(y, with_labels=True)
hook = hook_output(learn.model[0])
x, _ = first(test_dl)
with torch.no_grad(): output = learn.model.eval()(x)
act = hook.stored[0]
cam_map = torch.einsum('ck,kij->cij', learn.model[1][-1].weight, act)
x_dec = TensorImage(dls.train.decode((x,))[0][0])
_, ax = plt.subplots()
x_dec.show(ctx=ax)
ax.imshow(cam_map[cls].detach().cpu(), alpha=0.6, extent=(0,224,224,0),
interpolation='bilinear', cmap='magma');
hook.remove()
# + id="sDR9QDm8c7pr" cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 248} outputId="27a16845-7b7b-4e3a-c1c6-e609c34af7ae"
#@title **Grad-CAM**
#@markdown Gradient-weighted CAM (Grad-CAM) allows us to visualize the output from _any convolutional block_ in a CNN.
#@markdown By default, this cell is setup to show the Grad-CAM output from the final convolutional block in the CNN, for comparison to the CAM output.
#@markdown > Choose which of the two test images you would like to examine and run this cell to see the Grad-CAM output overlayed on the input image.
#@markdown >
#@markdown > Next, select a _different_ block and re-run the cell to see how the output changes for different blocks in the network.
test_case = 'abd' #@param ['abd', 'chest']
cls = 0 if test_case == 'abd' else 1
label = test_case
y = get_image_files(path/'test'/label)
test_dl = learn.dls.test_dl(y, with_labels=True)
x, _ = first(test_dl)
mod = learn.model[0]
block = -1 #@param {type: "slider", min: -8, max: -1, step: 1}
hook_func = lambda m,i,o: o[0].detach().clone()
with Hook(mod[block], hook_func, is_forward=False) as hookg:
with Hook(mod[block], hook_func) as hook:
output = learn.model.eval()(x.cuda())
act = hook.stored
output[0, cls].backward()
grad = hookg.stored
w = grad[0].mean(dim=[1,2], keepdim=True)
cam_map = (w * act[0]).sum(0)
x_dec = TensorImage(dls.train.decode((x,))[0][0])
_, ax = plt.subplots()
x_dec.show(ctx=ax)
ax.imshow(cam_map.detach().cpu(), alpha=0.6, extent=(0,224,224,0),
interpolation='bilinear', cmap='magma');
# + id="hjcrIiJ-zrGa"
| NIIC21_Image_Classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Objected-Oriented Programming in Python
#
# All materials are from the source below.
#
# Source: https://jeffknupp.com/blog/2014/06/18/improve-your-python-python-classes-and-object-oriented-programming/
class Customer(object):
"""A customer of ABC Bank with a checking account. Customers have the
following properties:
Attributes:
name: A string representing the customer's name.
balance: A float tracking the current balance of the customer's account.
"""
def __init__(self, name, balance=0.0):
"""Return a Customer object whose name is *name* and starting
balance is *balance*."""
self.name = name
self.balance = balance
def withdraw(self, amount):
"""Return the balance remaining after withdrawing *amount*
dollars."""
if amount > self.balance:
raise RuntimeError('Amount greater than available balance.')
self.balance -= amount
return self.balance
def deposit(self, amount):
"""Return the balance remaining after depositing *amount*
dollars."""
self.balance += amount
return self.balance
jeff = Customer('<NAME>', 1000.0)
print(jeff.name)
print(jeff.balance)
# ### Self?
#
# Self is the instance
# ### _ _init_ _
# Below two are the same:
# - jeff = Customer('<NAME>', 1000.0)
# - jeff = Customer(jeff, '<NAME>', 1000.0)
#
# ## Instance Attributes and Methods
#
#
# ### Static Methods and Class Methods
#
# To make it clear that this method should not receive the instance as the first parameter (i.e. self on "normal" methods), the @staticmethod decorator is used.
#
# Class methods may not make much sense right now, but that's because they're used most often in connection with our next topic: inheritance.
# +
class Car(object):
wheels = 4
def __init__(self, make, model):
self.make = make
self.model = model
mustang = Car('Ford', 'Mustang')
print (mustang.wheels)
# 4
print (Car.wheels)
# 4
# +
class Car(object):
...
@staticmethod
def make_car_sound():
print ('VRooooommmm!')
class Vehicle(object):
...
@classmethod
def is_motorcycle(cls):
return cls.wheels == 2
# -
# ### Inheritance
#
#
class Vehicle(object):
"""A vehicle for sale by <NAME>.
Attributes:
wheels: An integer representing the number of wheels the vehicle has.
miles: The integral number of miles driven on the vehicle.
make: The make of the vehicle as a string.
model: The model of the vehicle as a string.
year: The integral year the vehicle was built.
sold_on: The date the vehicle was sold.
"""
base_sale_price = 0
def __init__(self, wheels, miles, make, model, year, sold_on):
"""Return a new Vehicle object."""
self.wheels = wheels
self.miles = miles
self.make = make
self.model = model
self.year = year
self.sold_on = sold_on
def sale_price(self):
"""Return the sale price for this vehicle as a float amount."""
if self.sold_on is not None:
return 0.0 # Already sold
return 5000.0 * self.wheels
def purchase_price(self):
"""Return the price for which we would pay to purchase the vehicle."""
if self.sold_on is None:
return 0.0 # Not yet sold
return self.base_sale_price - (.10 * self.miles)
# +
class Car(Vehicle):
def __init__(self, wheels, miles, make, model, year, sold_on):
"""Return a new Car object."""
self.wheels = wheels
self.miles = miles
self.make = make
self.model = model
self.year = year
self.sold_on = sold_on
self.base_sale_price = 8000
class Truck(Vehicle):
def __init__(self, wheels, miles, make, model, year, sold_on):
"""Return a new Truck object."""
self.wheels = wheels
self.miles = miles
self.make = make
self.model = model
self.year = year
self.sold_on = sold_on
self.base_sale_price = 10000
| *Python_Basics/OOP/OOP-Object-Oriented Programming Intro.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6 - AzureML
# language: python
# name: python3-azureml
# ---
# <i>Copyright (c) Microsoft Corporation. All rights reserved.</i>
#
# <i>Licensed under the MIT License.</i>
# # Train SAR on MovieLens with Azure Machine Learning (Python, CPU)
# ---
# ## Introduction to Azure Machine Learning
# The **[Azure Machine Learning service (AzureML)](https://docs.microsoft.com/azure/machine-learning/service/overview-what-is-azure-ml)** provides a cloud-based environment you can use to prep data, train, test, deploy, manage, and track machine learning models. By using Azure Machine Learning service, you can start training on your local machine and then scale out to the cloud. With many available compute targets, like [Azure Machine Learning Compute](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-set-up-training-targets#amlcompute) and [Azure Databricks](https://docs.microsoft.com/en-us/azure/azure-databricks/what-is-azure-databricks), and with [advanced hyperparameter tuning services](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-tune-hyperparameters), you can build better models faster by using the power of the cloud.
#
# Data scientists and AI developers use the main [Azure Machine Learning Python SDK](https://docs.microsoft.com/en-us/python/api/overview/azure/ml/intro?view=azure-ml-py) to build and run machine learning workflows with the Azure Machine Learning service. You can interact with the service in any Python environment, including Jupyter Notebooks or your favorite Python IDE. The Azure Machine Learning SDK allows you the choice of using local or cloud compute resources, while managing and maintaining the complete data science workflow from the cloud.
# 
#
# This notebook provides an example of how to utilize and evaluate the Smart Adaptive Recommender (SAR) algorithm using the Azure Machine Learning service. It takes the content of the [SAR quickstart notebook](sar_movielens.ipynb) and demonstrates how to use the power of the cloud to manage data, switch to powerful GPU machines, and monitor runs while training a model.
#
# See the hyperparameter tuning notebook for more advanced use cases with AzureML.
#
# ### Advantages of using AzureML:
# - Manage cloud resources for monitoring, logging, and organizing your machine learning experiments.
# - Train models either locally or by using cloud resources, including GPU-accelerated model training.
# - Easy to scale out when dataset grows - by just creating and pointing to new compute target
#
# ---
# ## Details of SAR
# <details>
# <summary>Click to expand</summary>
#
# SAR is a fast scalable adaptive algorithm for personalized recommendations based on user transaction history. It produces easily explainable / interpretable recommendations and handles "cold item" and "semi-cold user" scenarios. SAR is a kind of neighborhood based algorithm (as discussed in [Recommender Systems by Aggarwal](https://dl.acm.org/citation.cfm?id=2931100)) which is intended for ranking top items for each user.
#
# SAR recommends items that are most ***similar*** to the ones that the user already has an existing ***affinity*** for. Two items are ***similar*** if the users who have interacted with one item are also likely to have interacted with another. A user has an ***affinity*** to an item if they have interacted with it in the past.
#
# ### Advantages of SAR:
# - High accuracy for an easy to train and deploy algorithm
# - Fast training, only requiring simple counting to construct matrices used at prediction time
# - Fast scoring, only involving multiplication of the similarity matric with an affinity vector
#
# ### Notes to use SAR properly:
# - SAR does not use item or user features, so cannot handle cold-start use cases
# - SAR requires the creation of an $mxm$ dense matrix (where $m$ is the number of items). So memory consumption can be an issue with large numbers of items.
# - SAR is best used for ranking items per user, as the scale of predicted ratings may be different from the input range and will differ across users.
#
# For more details see the deep dive notebook on SAR here: [SAR Deep Dive Notebook](../02_model/sar_deep_dive.ipynb)
# </details>
#
# ---
# ## Prerequisities
# - **Azure Subscription**
# - If you don’t have an Azure subscription, create a free account before you begin. Try the [free or paid version of Azure Machine Learning service today](https://azure.microsoft.com/en-us/free/services/machine-learning/).
# - You get credits to spend on Azure services, which will easily cover the cost of running this example notebook. After they're used up, you can keep the account and use [free Azure services](https://azure.microsoft.com/en-us/free/). Your credit card is never charged unless you explicitly change your settings and ask to be charged. Or [activate MSDN subscriber benefits](https://azure.microsoft.com/en-us/pricing/member-offers/credit-for-visual-studio-subscribers/), which give you credits every month that you can use for paid Azure services.
# ---
# +
# set the environment path to find Recommenders
import sys
sys.path.append("../../")
import os
import shutil
from tempfile import TemporaryDirectory
import azureml
from azureml.core import Workspace, Run, Experiment
from azureml.core.compute import ComputeTarget, AmlCompute
from azureml.train.estimator import Estimator
from azureml.widgets import RunDetails
from reco_utils.azureml.azureml_utils import get_or_create_workspace
from reco_utils.dataset import movielens
print("azureml.core version: {}".format(azureml.core.VERSION))
# + tags=["parameters"]
# top k items to recommend
TOP_K = 10
# Select Movielens data size: 100k, 1m, 10m, or 20m
MOVIELENS_DATA_SIZE = '1m'
# -
# ### Connect to an AzureML workspace
#
# An [AzureML Workspace](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.core.workspace.workspace?view=azure-ml-py) is an Azure resource that organizes and coordinates the actions of many other Azure resources to assist in executing and sharing machine learning workflows. In particular, an Azure ML Workspace coordinates storage, databases, and compute resources providing added functionality for machine learning experimentation, deployment, inferencing, and the monitoring of deployed models.
#
# The function below will get or create an AzureML Workspace and save the configuration to `aml_config/config.json`.
#
# It defaults to use provided input parameters or environment variables for the Workspace configuration values. Otherwise, it will use an existing configuration file (either at `./aml_config/config.json` or a path specified by the config_path parameter).
#
# Lastly, if the workspace does not exist, one will be created for you. See [this tutorial](https://docs.microsoft.com/en-us/azure/machine-learning/service/setup-create-workspace#portal) to locate information such as subscription id.
ws = get_or_create_workspace(
subscription_id="<SUBSCRIPTION_ID>",
resource_group="<RESOURCE_GROUP>",
workspace_name="<WORKSPACE_NAME>",
workspace_region="<WORKSPACE_REGION>"
)
# ### Create a Temporary Directory
# This directory will house the data and scripts needed by the AzureML Workspace
tmp_dir = TemporaryDirectory()
# ### Download dataset and upload to datastore
#
# Every workspace comes with a default [datastore](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-access-data) (and you can register more) which is backed by the Azure blob storage account associated with the workspace. We can use it to transfer data from local to the cloud, and access it from the compute target.
#
# The data files are uploaded into a directory named `data` at the root of the datastore.
# +
TARGET_DIR = 'movielens'
# download dataset
data = movielens.load_pandas_df(
size=MOVIELENS_DATA_SIZE,
header=['UserId','MovieId','Rating','Timestamp']
)
# upload dataset to workspace datastore
data_file_name = "movielens_" + MOVIELENS_DATA_SIZE + "_data.pkl"
data.to_pickle(os.path.join(tmp_dir.name, data_file_name))
ds = ws.get_default_datastore()
ds.upload(src_dir=tmp_dir.name, target_path=TARGET_DIR, overwrite=True, show_progress=True)
# -
# ### Create or Attach Azure Machine Learning Compute
#
# We create a cpu cluster as our **remote compute target**. If a cluster with the same name already exists in your workspace, the script will load it instead. You can read [Set up compute targets for model training](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-set-up-training-targets) to learn more about setting up compute target on different locations. You can also create GPU machines when larger machines are necessary to train the model.
#
# According to Azure [Pricing calculator](https://azure.microsoft.com/en-us/pricing/calculator/), with example VM size `STANDARD_D2_V2`, it costs a few dollars to run this notebook, which is well covered by Azure new subscription credit. For billing and pricing questions, please contact [Azure support](https://azure.microsoft.com/en-us/support/options/).
#
# **Note**:
# - 10m and 20m dataset requires more capacity than `STANDARD_D2_V2`, such as `STANDARD_NC6` or `STANDARD_NC12`. See list of all available VM sizes [here](https://docs.microsoft.com/en-us/azure/templates/Microsoft.Compute/2018-10-01/virtualMachines?toc=%2Fen-us%2Fazure%2Fazure-resource-manager%2Ftoc.json&bc=%2Fen-us%2Fazure%2Fbread%2Ftoc.json#hardwareprofile-object).
# - As with other Azure services, there are limits on certain resources (e.g. AzureML Compute quota) associated with the Azure Machine Learning service. Please read [these instructions](https://docs.microsoft.com/en-us/azure/azure-supportability/resource-manager-core-quotas-request) on the default limits and how to request more quota.
# ---
# #### Learn more about Azure Machine Learning Compute
# <details>
# <summary>Click to learn more about compute types</summary>
#
# [Azure Machine Learning Compute](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-set-up-training-targets#amlcompute) is managed compute infrastructure that allows the user to easily create single to multi-node compute of the appropriate VM Family. It is created within your workspace region and is a resource that can be used by other users in your workspace. It autoscales by default to the max_nodes, when a job is submitted, and executes in a containerized environment packaging the dependencies as specified by the user.
#
# Since it is managed compute, job scheduling and cluster management are handled internally by Azure Machine Learning service.
#
# You can provision a persistent AzureML Compute resource by simply defining two parameters thanks to smart defaults. By default it autoscales from 0 nodes and provisions dedicated VMs to run your job in a container. This is useful when you want to continously re-use the same target, debug it between jobs or simply share the resource with other users of your workspace.
#
# In addition to vm_size and max_nodes, you can specify:
# - **min_nodes**: Minimum nodes (default 0 nodes) to downscale to while running a job on AzureML Compute
# - **vm_priority**: Choose between 'dedicated' (default) and 'lowpriority' VMs when provisioning AzureML Compute. Low Priority VMs use Azure's excess capacity and are thus cheaper but risk your run being pre-empted
# - **idle_seconds_before_scaledown**: Idle time (default 120 seconds) to wait after run completion before auto-scaling to min_nodes
# - **vnet_resourcegroup_name**: Resource group of the existing VNet within which Azure MLCompute should be provisioned
# - **vnet_name**: Name of VNet
# - **subnet_name**: Name of SubNet within the VNet
# </details>
# ---
# +
# Remote compute (cluster) configuration. If you want to save the cost more, set these to small.
VM_SIZE = 'STANDARD_D2_V2'
# Cluster nodes
MIN_NODES = 0
MAX_NODES = 2
CLUSTER_NAME = 'cpucluster'
try:
compute_target = ComputeTarget(workspace=ws, name=CLUSTER_NAME)
print("Found existing compute target")
except:
print("Creating a new compute target...")
# Specify the configuration for the new cluster
compute_config = AmlCompute.provisioning_configuration(
vm_size=VM_SIZE,
min_nodes=MIN_NODES,
max_nodes=MAX_NODES
)
# Create the cluster with the specified name and configuration
compute_target = ComputeTarget.create(ws, CLUSTER_NAME, compute_config)
# Wait for the cluster to complete, show the output log
compute_target.wait_for_completion(show_output=True, min_node_count=None, timeout_in_minutes=20)
# -
# # Prepare training script
# ### 1. Create a directory
# Create a directory that will contain all the necessary code from your local machine that you will need access to on the remote resource. This includes the training script, and any additional files your training script depends on.
SCRIPT_DIR = os.path.join(tmp_dir.name, 'movielens-sar')
os.makedirs(SCRIPT_DIR, exist_ok=True)
TRAIN_FILE = os.path.join(SCRIPT_DIR, 'train.py')
# ### 2. Create a training script
# To submit the job to the cluster, first create a training script. Run the following code to create the training script called `train.py` in temporary directory. This training adds a regularization rate to the training algorithm, so produces a slightly different model than the local version.
#
# This code takes what is in the local quickstart and convert it to one single training script. We use run.log() to record parameters to the run. We will be able to review and compare these measures in the Azure Portal at a later time.
# +
# %%writefile $TRAIN_FILE
import argparse
import os
import numpy as np
import pandas as pd
import itertools
import logging
import time
from azureml.core import Run
from sklearn.externals import joblib
from reco_utils.dataset import movielens
from reco_utils.dataset.python_splitters import python_random_split
from reco_utils.evaluation.python_evaluation import map_at_k, ndcg_at_k, precision_at_k, recall_at_k
from reco_utils.recommender.sar.sar_singlenode import SARSingleNode
TARGET_DIR = 'movielens'
OUTPUT_FILE_NAME = 'outputs/movielens_sar_model.pkl'
MODEL_FILE_NAME = 'movielens_sar_model.pkl'
# get hold of the current run
run = Run.get_context()
# let user feed in 2 parameters, the location of the data files (from datastore), and the regularization rate of the logistic regression model
parser = argparse.ArgumentParser()
parser.add_argument('--data-folder', type=str, dest='data_folder', help='data folder mounting point')
parser.add_argument('--data-file', type=str, dest='data_file', help='data file name')
parser.add_argument('--top-k', type=int, dest='top_k', default=10, help='top k items to recommend')
parser.add_argument('--data-size', type=str, dest='data_size', default=10, help='Movielens data size: 100k, 1m, 10m, or 20m')
args = parser.parse_args()
data_pickle_path = os.path.join(args.data_folder, args.data_file)
data = pd.read_pickle(path=data_pickle_path)
# Log arguments to the run for tracking
run.log("top-k", args.top_k)
run.log("data-size", args.data_size)
train, test = python_random_split(data)
# instantiate the SAR algorithm and set the index
header = {
"col_user": "UserId",
"col_item": "MovieId",
"col_rating": "Rating",
"col_timestamp": "Timestamp",
}
logging.basicConfig(level=logging.DEBUG,
format='%(asctime)s %(levelname)-8s %(message)s')
model = SARSingleNode(
remove_seen=True, similarity_type="jaccard",
time_decay_coefficient=30, time_now=None, timedecay_formula=True, **header
)
# train the SAR model
start_time = time.time()
model.fit(train)
train_time = time.time() - start_time
run.log(name="Training time", value=train_time)
start_time = time.time()
top_k = model.recommend_k_items(test)
test_time = time.time() - start_time
run.log(name="Prediction time", value=test_time)
# TODO: remove this call when the model returns same type as input
top_k['UserId'] = pd.to_numeric(top_k['UserId'])
top_k['MovieId'] = pd.to_numeric(top_k['MovieId'])
# evaluate
eval_map = map_at_k(test, top_k, col_user="UserId", col_item="MovieId",
col_rating="Rating", col_prediction="prediction",
relevancy_method="top_k", k=args.top_k)
eval_ndcg = ndcg_at_k(test, top_k, col_user="UserId", col_item="MovieId",
col_rating="Rating", col_prediction="prediction",
relevancy_method="top_k", k=args.top_k)
eval_precision = precision_at_k(test, top_k, col_user="UserId", col_item="MovieId",
col_rating="Rating", col_prediction="prediction",
relevancy_method="top_k", k=args.top_k)
eval_recall = recall_at_k(test, top_k, col_user="UserId", col_item="MovieId",
col_rating="Rating", col_prediction="prediction",
relevancy_method="top_k", k=args.top_k)
run.log("map", eval_map)
run.log("ndcg", eval_ndcg)
run.log("precision", eval_precision)
run.log("recall", eval_recall)
# automatic upload of everything in ./output folder doesn't work for very large model file
# model file has to be saved to a temp location, then uploaded by upload_file function
joblib.dump(value=model, filename=MODEL_FILE_NAME)
run.upload_file(OUTPUT_FILE_NAME, MODEL_FILE_NAME)
# -
# # copy dependent python files
UTILS_DIR = os.path.join(SCRIPT_DIR, 'reco_utils')
shutil.copytree('../../reco_utils/', UTILS_DIR)
# # Run training script
# ### 1. Create an estimator
# An [estimator](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-train-ml-models) object is used to submit the run. You can create and use a generic Estimator to submit a training script using any learning framework you choose (such as scikit-learn) you want to run on any compute target, whether it's your local machine, a single VM in Azure, or a GPU cluster in Azure.
#
# Create your estimator by running the following code to define:
# * The name of the estimator object, `est`
# * The directory that contains your scripts. All the files in this directory are uploaded into the cluster nodes for execution.
# * The compute target. In this case you will use the AzureML Compute you created
# * The training script name, train.py
# * Parameters required from the training script
# * Python packages needed for training
# * Connect to the data files in the datastore
#
# In this tutorial, this target is AzureML Compute. All files in the script folder are uploaded into the cluster nodes for execution. `ds.as_mount()` mounts a datastore on the remote compute and returns the folder. See documentation [here](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-access-data#access-datastores-during-training).
# + tags=["configure estimator"]
script_params = {
'--data-folder': ds.as_mount(),
'--data-file': 'movielens/' + data_file_name,
'--top-k': TOP_K,
'--data-size': MOVIELENS_DATA_SIZE
}
est = Estimator(source_directory=SCRIPT_DIR,
script_params=script_params,
compute_target=compute_target,
entry_script='train.py',
conda_packages=['pandas'],
pip_packages=['sklearn', 'tqdm'])
# -
# ### 2. Submit the job to the cluster
# An [experiment](https://docs.microsoft.com/en-us/python/api/overview/azure/ml/intro?view=azure-ml-py#experiment) is a logical container in an AzureML Workspace. It hosts run records which can include run metrics and output artifacts from your experiments. We access an experiment from our AzureML workspace by name, which will be created if it doesn't exist.
#
# Then, run the experiment by submitting the estimator object.
# +
# create experiment
EXPERIMENT_NAME = 'movielens-sar'
exp = Experiment(workspace=ws, name=EXPERIMENT_NAME)
run = exp.submit(config=est)
# -
#
# ### 3. Monitor remote run
#
# #### Jupyter widget
#
# Jupyter widget can watch the progress of the run. Like the run submission, the widget is asynchronous and provides live updates every 10-15 seconds until the job completes.
RunDetails(run).show()
# ### 4. Viewing run results
# Azure Machine Learning stores all the details about the run in the Azure cloud. Let's access those details by retrieving a link to the run using the default run output. Clicking on the resulting link will take you to an interactive page.
run
# Above cell should output similar table as below.
# 
# After clicking "Link to Azure Portal", experiment run details tab looks like this with logged metrics.
# 
# run below after run is complete, otherwise metrics is empty
metrics = run.get_metrics()
print(metrics)
# # Deprovision compute resource
# To avoid unnecessary charges, if you created compute target that doesn't scale down to 0, make sure the compute target is deprovisioned after use.
# +
# delete () is used to deprovision and delete the AzureML Compute target.
# do not run below before experiment completes
# compute_target.delete()
# deletion will take a few minutes. You can check progress in Azure Portal / Computing tab
# -
# clean up temporary directory
tmp_dir.cleanup()
| notebooks/00_quick_start/sar_movielens_with_azureml.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/misabhishek/gcp-iam-recommender/blob/main/all_firewall_insights_in_an_org.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="W_rpQ2AiSPPK"
# #Problem Statement
#
# Get all firewall insights in an org
# + [markdown] id="3USCxX9ESZ5I"
# ### Imports
# + id="WkYIk28fShJw"
from google.colab import auth
import json
import subprocess
import concurrent
import logging
logging.basicConfig(format="%(levelname)s[%(asctime)s]:%(message)s")
# + [markdown] id="evqyijtaSjVU"
# ### Lets authenticate
# + id="hZGML0yrSmoS" outputId="39257e94-e04a-4a43-b178-44cc601ab410" colab={"base_uri": "https://localhost:8080/"}
auth.authenticate_user()
print('Authenticated')
# + [markdown] id="Xs_eM8aqSqnb"
# ### Helper functions
# + id="ItzbkPSXS1lx"
def execute_command(command):
return json.loads(subprocess.check_output(filter(lambda x: x, command.split(" "))).decode("utf-8"))
# + [markdown] id="7388TwhAS5KC"
# ### Enter your org details
# + id="_oJmktKHS8S9"
organization = "" # organizations/[ORGANIZATION-ID]
billing_project_id = "" # Billing project
# + [markdown] id="nQCho5C2S_Ub"
# ### Get All the projects
# + id="Jmf0amhLTUgs"
get_all_projects_command = f"""gcloud asset search-all-resources \
--asset-types=cloudresourcemanager.googleapis.com/Project \
--scope={organization} --format=json --project={billing_project_id}"""
# + id="kJLq-SteTeDs"
def get_all_projects():
projects = execute_command(get_all_projects_command)
return [p["additionalAttributes"]["projectId"] for p in projects]
project_ids = get_all_projects()
print("project-ids\n\n", "\n".join(project_ids[:10]))
# + [markdown] id="DpCmtf9cTxe6"
# ### Get All Firewall insights
# + id="z79MeXx2VcOT"
firewall_insight_command = """gcloud beta recommender insights list \
--project={} \
--location=global --insight-type=google.compute.firewall.Insight \
--format=json --billing-project=""" + billing_project_id
# + id="KhpNWxU-V450"
def get_all_firewall_insights():
def get_insights(project_id):
try:
return {"project_id":project_id,
"insights": execute_command(firewall_insight_command.format(project_id))}
except:
logging.warning(f"You don't have permissions to access project:`{project_id}`")
return {"project_id": project_id,
"insights": []}
with concurrent.futures.ThreadPoolExecutor(max_workers=100) as executor:
insights = {"all_insights": list(executor.map(get_insights, project_ids))}
return insights
# + id="JnhlB22rWFW-"
all_firewall_insights = get_all_firewall_insights()
# + id="M2Aof5DSXWJb"
| all_firewall_insights_in_an_org.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Análise de Dados com Python
#
#
# Neste notebook, utilizaremos dados de automóveis para analisar a influência das características de um carro em seu preço, tentando posteriormente prever qual será o preço de venda de um carro. Utilizaremos como fonte de dados um arquivo .csv com dados já tratados em outro notebook. Caso você tenha dúvidas quanto a como realizar o tratamento dos dados, dê uma olhada no meu repositório Learn-Pandas
import pandas as pd
import numpy as np
df = pd.read_csv('clean_auto_df.csv')
df.head()
# <h4> Utilizando visualização de dados para verificar padrões de características individuais</h4>
# Importando as bibliotecas "Matplotlib" e "Seaborn
# utilizando "%matplotlib inline" para plotar o gráfico dentro do notebook.
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
# <h4> Como escolher o método de visualização correto? </h4>
# <p> Ao visualizar variáveis individuais, é importante primeiro entender com que tipo de variável você está lidando. Isso nos ajudará a encontrar o método de visualização correto para essa variável. Por exemplo, podemos calcular a correlação entre variáveis do tipo "int64" ou "float64" usando o método "corr":</p>
df.corr()
# Os elementos diagonais são sempre um; (estudaremos isso, mais precisamente a correlação de Pearson no final do notebook)
#
# +
# se quisermos verificar a correlação de apenas algumas colunas
df[['bore', 'stroke', 'compression-ratio', 'horsepower']].corr()
# -
# <h2> Variáveis numéricas contínuas: </h2>
#
# <p> Variáveis numéricas contínuas são variáveis que podem conter qualquer valor dentro de algum intervalo. Variáveis numéricas contínuas podem ter o tipo "int64" ou "float64". Uma ótima maneira de visualizar essas variáveis é usando gráficos de dispersão com linhas ajustadas. </p>
#
# <p> Para começar a compreender a relação (linear) entre uma variável individual e o preço. Podemos fazer isso usando "regplot", que plota o gráfico de dispersão mais a linha de regressão ajustada para os dados. </p>
# <h4> Relação linear positiva </h4>
# Vamos encontrar o gráfico de dispersão de "engine-size" e "price"
# Engine size as potential predictor variable of price
sns.regplot(x="engine-size", y="price", data=df)
plt.ylim(0,)
# <p> Note que conforme o tamanho do motor aumenta, o preço sobe: isso indica uma correlação direta positiva entre essas duas variáveis. O tamanho do motor parece um bom preditor de preço, já que a linha de regressão é quase uma linha diagonal perfeita. </p>
# Podemos examinar a correlação entre 'engine-size' e 'price' e ver que é aproximadamente 0,87
df[["engine-size", "price"]].corr()
# <h4> Relação linear Negativa </h4>
#
# city-mpg também pode ser um bom preditor para a variável price:
sns.regplot(x="city-mpg", y="price", data=df)
# <p> À medida que o city-mpg sobe, o preço desce: isso indica uma relação inversa / negativa entre essas duas variáveis, podendo ser um indicador de preço. </p>
df[['city-mpg', 'price']].corr()
# <h4> Relação linear neutra (ou fraca) </h4>
#
sns.regplot(x="peak-rpm", y="price", data=df)
# <p> A variável peak-rpm não parece ser um bom preditor do preço, pois a linha de regressão está próxima da horizontal. Além disso, os pontos de dados estão muito dispersos e distantes da linha ajustada, apresentando grande variabilidade. Portanto, não é uma variável confiável. </p>
df[['peak-rpm','price']].corr()
# <h2> Variáveis categóricas: </h2>
#
#
# <p> Essas são variáveis que descrevem uma 'característica' de uma unidade de dados e são selecionadas a partir de um pequeno grupo de categorias. As variáveis categóricas podem ser do tipo "objeto" ou "int64". Uma boa maneira de visualizar variáveis categóricas é usar boxplots. </p>
sns.boxplot(x="body-style", y="price", data=df)
# Vemos que as distribuições de preço entre as diferentes categorias de body-style têm uma sobreposição significativa e, portanto, body-style não seria um bom preditor de preço. Vamos examinar a "engine-location" e o "price" do motor:
sns.boxplot(x="engine-location", y="price", data=df)
# <p> Aqui, vemos que a distribuição de preço entre essas duas categorias de localização do motor, dianteira e traseira, são distintas o suficiente para considerar a localização do motor como um bom indicador de preço em potencial. </p>
# drive-wheels
sns.boxplot(x="drive-wheels", y="price", data=df)
# <p> Aqui vemos que a distribuição de preço entre as diferentes categorias de drive-wheels difere e podem ser um indicador de preço. </p>
# <h2> Estatística Descritiva </h2>
#
# <p> Vamos primeiro dar uma olhada nas variáveis usando um método de descrição. </p>
#
# <p> A função <b> describe </b> calcula automaticamente estatísticas básicas para todas as variáveis contínuas. Quaisquer valores NaN são automaticamente ignorados nessas estatísticas. </p>
#
# Isso mostrará:
#
# <ul>
# <li> a contagem dessa variável </li>
# <li> a média </li>
# <li> o desvio padrão (std) </li>
# <li> o valor mínimo </li>
# <li> o IQR (intervalo interquartil: 25%, 50% e 75%) </li>
# <li> o valor máximo </li>
# <ul>
df.describe()
# A configuração padrão de "describe" ignora variáveis do tipo de objeto.
# Podemos aplicar o método "describe" nas variáveis do tipo 'objeto' da seguinte forma:
df.describe(include=['object'])
# <h3>Value Counts</h3>
#
# A contagem de valores é uma boa maneira de entender quantas unidades de cada característica / variável temos.
# Podemos aplicar o método "value_counts" na coluna 'drive-wheels'.
# Não se esqueça que o método "value_counts" só funciona na série Pandas, não nos Dataframes Pandas.
# Por isso, incluímos apenas um colchete "df ['drive-wheels']" e não dois colchetes "df [['drive-wheels']]".
#
df['drive-wheels'].value_counts()
# +
# nós podemos converter a série para um dataframe:
df['drive-wheels'].value_counts().to_frame()
# -
drive_wheels_counts = df['drive-wheels'].value_counts().to_frame()
drive_wheels_counts.rename(columns={'drive-wheels': 'value_counts'}, inplace=True)
drive_wheels_counts
# +
# vamos renomear o index para 'drive-wheels':
drive_wheels_counts.index.name = 'drive-wheels'
drive_wheels_counts
# -
# repetindo o processo para engine-location
engine_loc_counts = df['engine-location'].value_counts().to_frame()
engine_loc_counts.rename(columns={'engine-location': 'value_counts'}, inplace=True)
engine_loc_counts.index.name = 'engine-location'
engine_loc_counts.head()
# <h2>Agrupando</h2>
#
#
# <p> O método "groupby" agrupa os dados por categorias diferentes. Os dados são agrupados com base em uma ou várias variáveis e a análise é realizada nos grupos individuais. </p>
#
# <p> Por exemplo, vamos agrupar pela variável "drive-wheels". Vemos que existem 3 categorias diferentes de rodas motrizes. </p>
df['drive-wheels'].unique()
# <p> Se quisermos saber, em média, qual tipo de drive-wheels é mais valiosa, podemos agrupar "drive-wheels" e depois fazer a média delas. </p>
#
# <p> Podemos selecionar as colunas 'drive-wheels', 'body-style' e 'price' e, em seguida, atribuí-las à variável "df_group_one". </p>
df_group_one = df[['drive-wheels','body-style','price']]
# Podemos então calcular o preço médio para cada uma das diferentes categorias de dados
df_group_one = df_group_one.groupby(['drive-wheels'],as_index=False).mean()
df_group_one
# <p> Pelos nossos dados, parece que os veículos com tração traseira são, em média, os mais caros, enquanto as 4 rodas e as rodas dianteiras têm preços aproximadamente iguais. </p>
#
# <p> Você também pode agrupar com várias variáveis. Por exemplo, vamos agrupar por 'drive-wheels' e 'body-style'. Isso agrupa o dataframe pelas combinações exclusivas 'drive-wheels' e 'body-style'. Podemos armazenar os resultados na variável 'grouped_test1'. </p>
df_gptest = df[['drive-wheels','body-style','price']]
grouped_test1 = df_gptest.groupby(['drive-wheels','body-style'],as_index=False).mean()
grouped_test1
# Esses dados agrupados são muito mais fáceis de visualizar quando transformados em uma tabela dinâmica. Uma tabela dinâmica é como uma planilha do Excel, com uma variável ao longo da coluna e outra ao longo da linha. Podemos converter o dataframe em uma tabela dinâmica usando o método "pivô" para criar uma tabela dinâmica a partir dos grupos.
#
# Nesse caso, deixaremos a variável da drive-wheels como as linhas da tabela e giraremos no estilo do corpo para se tornar as colunas da tabela:
grouped_pivot = grouped_test1.pivot(index='drive-wheels',columns='body-style')
grouped_pivot
# As vezes não teremos dados para algumas das células pivô. Podemos preencher essas células ausentes com o valor 0, mas qualquer outro valor também pode ser usado. Deve ser mencionado que a falta de dados é um assunto bastante complexo...
grouped_pivot = grouped_pivot.fillna(0) #fill missing values with 0
grouped_pivot
df_gptest2 = df[['body-style','price']]
grouped_test_bodystyle = df_gptest2.groupby(['body-style'],as_index= False).mean()
grouped_test_bodystyle
# <h2>Visualização dos dados</h2>
#
# Vamos usar um mapa de calor para visualizar a relação entre body-style e price.
import matplotlib.pyplot as plt
# %matplotlib inline
plt.pcolor(grouped_pivot, cmap='RdBu')
plt.colorbar()
plt.show()
# <p> O mapa de calor representa a variável alvo (price) proporcional à cor em relação às variáveis 'drive-wheels' e 'body-style' nos eixos vertical e horizontal, respectivamente. Isso nos permite visualizar como o preço está relacionado a 'drive-wheels' e 'body-style'. </p>
#
# <p> Os rótulos padrão não transmitem informações úteis para nós. Vamos mudar isso: </p>
# +
fig, ax = plt.subplots()
im = ax.pcolor(grouped_pivot, cmap='RdBu')
#label names
row_labels = grouped_pivot.columns.levels[1]
col_labels = grouped_pivot.index
#move ticks and labels to the center
ax.set_xticks(np.arange(grouped_pivot.shape[1]) + 0.5, minor=False)
ax.set_yticks(np.arange(grouped_pivot.shape[0]) + 0.5, minor=False)
#insert labels
ax.set_xticklabels(row_labels, minor=False)
ax.set_yticklabels(col_labels, minor=False)
#rotate label if too long
plt.xticks(rotation=90)
fig.colorbar(im)
plt.show()
# -
# <p> A visualização é muito importante na ciência de dados e os pacotes de visualização oferecem grande liberdade</p>
#
# <p> A principal questão que queremos responder neste notebook é "Quais são as principais características que têm mais impacto no preço do carro?". </p>
#
# <p> Para obter uma melhor medida das características importantes, olhamos para a correlação dessas variáveis com o preço do carro, em outras palavras: como o preço do carro depende dessa variável? </p>
# <h2>Correlação e Causalidade</h2>
# <p> <b> Correlação </b>: uma medida da extensão da interdependência entre as variáveis. </p>
#
# <p> <b> Causalidade </b>: a relação entre causa e efeito entre duas variáveis. </p>
#
# <p> É importante saber a diferença entre os dois e que a correlação não implica causalidade. Determinar a correlação é muito mais simples do que determinar a causalidade, pois a causalidade pode exigir experimentação independente. </p>
# <p3> Correlação de Pearson </p>
#
# <p> A Correlação de Pearson mede a dependência linear entre duas variáveis X e Y. </p>
# <p> O coeficiente resultante é um valor entre -1 e 1 inclusive, onde: </p>
# <ul>
# <li> <b> 1 </b>: Correlação linear positiva total. </li>
# <li> <b> 0 </b>: Sem correlação linear, as duas variáveis provavelmente não se afetam. </li>
# <li> <b> -1 </b>: Correlação linear negativa total. </li>
# </ul>
#
# <p> Correlação de Pearson é o método padrão da função "corr". Como antes, podemos calcular a Correlação de Pearson das variáveis 'int64' ou 'float64'. </p>
df.corr()
# <b> P-value </b>:
#
# <p>P-value é o valor da probabilidade de que a correlação entre essas duas variáveis seja estatisticamente significativa. Normalmente, escolhemos um nível de significância de 0.05, o que significa que temos 95% de confiança de que a correlação entre as variáveis é significativa. </p>
#
# Por convenção, quando o
#
# <ul>
# <li> o valor de p é $ <$ 0.001: afirmamos que há fortes evidências de que a correlação é significativa. </li>
# <li> o valor p é $ <$ 0.05: há evidências moderadas de que a correlação é significativa. </li>
# <li> o valor p é $ <$ 0.1: há evidências fracas de que a correlação é significativa. </li>
# <li> o valor p é $> $ 0.1: não há evidências de que a correlação seja significativa. </li>
# </ul>
# +
# Podemos obter essas informações usando o módulo "stats" da biblioteca "scipy"
from scipy import stats
# -
# <h3>Wheel-base vs Price</h3>
#
# Vamos calcular o coeficiente de correlação de Pearson e o P-value entre 'wheel-base' e 'price'.
pearson_coef, p_value = stats.pearsonr(df['wheel-base'], df['price'])
print('Coeficiente de Pearson', pearson_coef)
print('P-value', p_value)
# A notacão científica do resultado indica que o valor é muito maior ou muito pequeno.
#
# No caso de 8.076488270733218e-20 significa:
#
#
# 8.076488270733218 vezes 10 elevado a menos 20 (o que faz andar a casa decimal 20 vezes para esquerda):
# 0,0000000000000000008076488270733218
# <h5> Conclusão: </h5>
# <p> Como o P-value é $ <$ 0.001, a correlação entre wheel-base e price é estatisticamente significativa, embora a relação linear não seja extremamente forte (~ 0,585) </p>
# <h3>Horsepower vs Price</h3>
#
pearson_coef, p_value = stats.pearsonr(df['horsepower'], df['price'])
print('Coeficiente de Pearson', pearson_coef)
print('P-value', p_value)
# <h5> Conclusão: </h5>
#
# <p> Como o P-value é $ <$ 0,001, a correlação entre a horsepower e price é estatisticamente significativa, e a relação linear é bastante forte (~ 0,809, próximo de 1) </p>
# <h3>Length vs Price</h3>
#
pearson_coef, p_value = stats.pearsonr(df['length'], df['price'])
print('Coeficiente de Pearson', pearson_coef)
print('P-value', p_value)
# <h5> Conclusão: </h5>
# <p> Como o valor p é $ <$ 0,001, a correlação entre length e price é estatisticamente significativa, e a relação linear é moderadamente forte (~ 0,691). </p>
# <h3>Width vs Price</h3>
#
pearson_coef, p_value = stats.pearsonr(df['width'], df['price'])
print('Coeficiente de Pearson', pearson_coef)
print('P-value', p_value)
# ##### Conclusão:
#
# Como o valor p é <0,001, a correlação entre largura e preço é estatisticamente significativa e a relação linear é bastante forte (~ 0,751).
# <h2>ANOVA</h2>
#
# <p> A Análise de Variância (ANOVA) é um método estatístico usado para testar se existem diferenças significativas entre as médias de dois ou mais grupos. ANOVA retorna dois parâmetros: </p>
#
# <p> <b> F-test score </b>: ANOVA assume que as médias de todos os grupos são iguais, calcula o quanto as médias reais se desviam da suposição e relata como a pontuação do F-test. Uma pontuação maior significa que há uma diferença maior entre as médias. </p>
#
# <p> <b> P-value </b>: P-value diz o quão estatisticamente significativo é nosso valor de pontuação calculado. </p>
#
# <p> Se nossa variável de preço estiver fortemente correlacionada com a variável que estamos analisando, espere que a ANOVA retorne uma pontuação considerável no F-test e um pequeno P-value. </p>
# <h3>Drive Wheels</h3>
#
# <p> Uma vez que ANOVA analisa a diferença entre diferentes grupos da mesma variável, a função groupby será útil. Como o algoritmo ANOVA calcula a média dos dados automaticamente, não precisamos tirar a média antes. </p>
#
# <p> Vamos ver se diferentes tipos de 'drive wheels' afetam o 'price', agrupamos os dados. </ p>
#
grouped_test2=df_gptest[['drive-wheels', 'price']].groupby(['drive-wheels'])
grouped_test2.head(2)
# Podemos obter os valores do grupo de métodos usando o método "get_group".
grouped_test2.get_group('4wd')['price']
# +
# podemos usar a função 'f_oneway' no módulo 'stats' para obter pontuação do test-F e o P-value
f_val, p_val = stats.f_oneway(grouped_test2.get_group('fwd')['price'], grouped_test2.get_group('rwd')['price'], grouped_test2.get_group('4wd')['price'])
print( "ANOVA: F=", f_val, ", P =", p_val)
# -
# Este é um ótimo resultado, com uma grande pontuação no test-F mostrando uma forte correlação e um P-value de quase 0 implicando em significância estatística quase certa. Mas isso significa que todos os três grupos testados são altamente correlacionados?
#### fwd e rwd
f_val, p_val = stats.f_oneway(grouped_test2.get_group('fwd')['price'], grouped_test2.get_group('rwd')['price'])
print( "ANOVA: F=", f_val, ", P =", p_val )
#### 4wd and rwd
f_val, p_val = stats.f_oneway(grouped_test2.get_group('4wd')['price'], grouped_test2.get_group('rwd')['price'])
print( "ANOVA: F=", f_val, ", P =", p_val)
#### 4wd and fwd
f_val, p_val = stats.f_oneway(grouped_test2.get_group('4wd')['price'], grouped_test2.get_group('fwd')['price'])
print("ANOVA: F=", f_val, ", P =", p_val)
# <h3>Conclusão</h3>
#
# <p> Agora temos uma ideia melhor de como são os nossos dados e quais variáveis são importantes levar em consideração ao prever o preço do carro.</p>
#
#
# <p> À medida que avançamos na construção de modelos de aprendizado de máquina para automatizar nossa análise, alimentar o modelo com variáveis que afetam significativamente nossa variável de destino melhorará o desempenho de previsão do nosso modelo. </p>
# # É isso!
#
# ### Este é apenas um exemplo de análise de dados com Python
# Este notebook faz parte de uma série de notebooks com conteúdos extraídos de cursos dos quais participei como aluno, ouvinte, professor, monitor... Reunidos para consulta futura e compartilhamento de idéias, soluções e conhecimento!
#
# ### Muito obrigado pela sua leitura!
#
#
# <h4><NAME></h4>
#
# Você pode encontrar mais conteúdo no meu Medium<br> ou então entrar em contato comigo :D
#
# <a href="https://www.linkedin.com/in/andercordeiro/" target="_blank">[LinkedIn]</a>
# <a href="https://medium.com/@andcordeiro" target="_blank">[Medium]</a>
#
| .ipynb_checkpoints/pandas-basico-analise-carros-Copy1-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # cobrapy compliance
#
# Notebook for confirming that every field is preserved when the model is used with cobrapy.
import cobra
model = cobra.io.read_sbml_model("../../model/yeast-GEM.xml")
# ## 1. Metabolites
model.metabolites[0]
model.metabolites[0].charge
model.metabolites[0].annotation
# ## 2. Reactions
model.reactions.get_by_id("r_2112")
model.reactions.get_by_id("r_2112").annotation
model.reactions.get_by_id("r_2112").notes
# ## 3. Genes
model.genes[0]
# # 4. Subsystems
model.groups[4].name
model.groups[4].members
| code/modelTests/cobrapy-compliance.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
import pickle
import numpy as np
import pandas as pd
import random
import plotnine
from plotnine import *
np.random.seed(1234)
random.seed(1234)
#load the data
entries=pickle.load(open('cnn_inputs.p','rb'))
X=entries['X']
y=entries['y']
#create train,test,validate splits
indices=list(X.keys())
print(len(indices))
print(len(y.keys()))
print(indices[0:10])
random.shuffle(indices)
num_items=len(indices)
#split 70% train, 15% validate, 15% test
num_train=int(round(0.7*num_items))
num_validate=int(round(0.15*num_items))
num_test=num_items-num_train-num_validate
print(num_train)
print(num_validate)
print(num_test)
train_indices=indices[0:num_train]
validate_indices=indices[num_train:num_train+num_validate]
test_indices=indices[num_train+num_validate::]
# +
train_X_seq=np.array([X[i][0] for i in train_indices])
train_X_struct=np.array([X[i][1] for i in train_indices])
train_X=[train_X_seq,train_X_struct]
validate_X_seq=np.array([X[i][0] for i in validate_indices])
validate_X_struct=np.array([X[i][1] for i in validate_indices])
validate_X=[validate_X_seq,validate_X_struct]
test_X_seq=np.array([X[i][0] for i in test_indices])
test_X_struct=np.array([X[i][1] for i in test_indices])
test_X=[test_X_seq,test_X_struct]
# -
train_y=np.asarray([y[i] for i in train_indices])
validate_y=np.asarray([y[i] for i in validate_indices])
test_y=np.asarray([y[i] for i in test_indices])
train_X_seq.shape
train_X_struct.shape
train_y.shape
# ## Create keras model
#import keras dependencies
import keras
from keras.models import Model
from keras.layers import Input, Concatenate
from keras.layers.core import Dropout, Reshape, Dense, Activation, Flatten
from keras.layers.convolutional import Conv1D, MaxPooling1D
from keras.optimizers import Adadelta, SGD, RMSprop;
import keras.losses;
from keras.constraints import maxnorm;
from keras.layers.normalization import BatchNormalization
from keras.regularizers import l1, l2
from keras import backend as K
from kerasAC.custom_losses import *
#define a keras model
K.set_image_data_format('channels_last')
padding='same'
seq_input=Input(shape=(166,4),name='seq_input')
struct_input=Input(shape=(166,6),name='struct_input')
x_seq=Conv1D(filters=1,kernel_size=6,input_shape=(166,4),padding=padding,name='conv_seq')(seq_input)
x_struct=Conv1D(filters=1,kernel_size=6,input_shape=(166,6),padding=padding,name='conv_struct')(struct_input)
x_seq=Activation('relu',name='activation_1')(x_seq)
x_struct=Activation('relu',name='activation_2')(x_struct)
x_seq=Flatten(name='flatten1')(x_seq)
x_struct=Flatten(name='flatten2')(x_struct)
aggregated=Concatenate(axis=-1)([x_struct,x_seq])
x=Dense(32,name='dense1')(aggregated)
x=Activation('relu',name='activation_3')(x)
outputs=Dense(1,name='final')(x)
model=Model(inputs=[seq_input,struct_input],outputs=outputs)
adam = keras.optimizers.Adam(lr=0.001, beta_1=0.9, beta_2=0.999, epsilon=1e-08)
model.compile(optimizer=adam,loss=ambig_mean_squared_error)
model.summary()
# ## Train the model
from keras.callbacks import *
from kerasAC.custom_callbacks import *
#define callbacks
checkpointer = ModelCheckpoint(filepath="model.hdf5", verbose=1, save_best_only=True)
earlystopper = EarlyStopping(monitor='val_loss', patience=3, verbose=1,restore_best_weights=True)
history=LossHistory("logs.batch",['loss','val_loss'])
csvlogger = CSVLogger("logs.epoch.csv", append = False)
reduce_lr = ReduceLROnPlateau(monitor='val_loss', factor=0.4,patience=2, min_lr=0.00000001)
cur_callbacks=[checkpointer,earlystopper,csvlogger,reduce_lr,history]
#fit the model
model.fit(x=train_X,
y=train_y,
batch_size=32,
epochs=40,
verbose=1,
callbacks=cur_callbacks,
validation_data=(validate_X,validate_y),
shuffle=True,
max_queue_size=100,
use_multiprocessing=True,
workers=12)
# ## plot the loss curves
## plot the losses
losses=pd.read_csv("logs.epoch.csv",header=0,sep=',')
losses.shape
batch_losses=pd.read_csv("logs.batch",header=0,sep='\t')
losses=pd.melt(losses,id_vars='epoch')
losses['variable'][losses['variable']=='loss']='TrainingLoss'
losses['variable'][losses['variable']=='val_loss']='ValidationLoss'
plotnine.options.figure_size = (6, 6)
(ggplot(losses,aes(x='epoch',
y='value',
group='variable',
color='variable'))+geom_line(size=2)+theme_bw(20))
# +
batch_losses['sequence']=batch_losses.index
plotnine.options.figure_size = (6, 6)
(ggplot(batch_losses,aes(x="sequence",y="loss"))+geom_line()+ylab("Training Loss MSE"))
# -
# ## get predictions
predictions_test=model.predict(test_X,batch_size=128,max_queue_size=100,use_multiprocessing=True,workers=4)
predictions_train=model.predict(train_X,batch_size=128,max_queue_size=100,use_multiprocessing=True,workers=4)
predictions_validate=model.predict(validate_X,batch_size=128,max_queue_size=100,use_multiprocessing=True,workers=4)
#get performance metrics
from kerasAC.performance_metrics.regression_performance_metrics import *
test_perf=get_performance_metrics_regression(predictions_test,np.expand_dims(test_y,axis=1))
test_perf
train_perf=get_performance_metrics_regression(predictions_train,np.expand_dims(train_y,axis=1))
train_perf
valid_perf=get_performance_metrics_regression(predictions_validate,np.expand_dims(validate_y,axis=1))
valid_perf
# +
#plot the predictions
# %matplotlib inline
import matplotlib.pyplot as plt
from matplotlib import cm
from matplotlib.colors import Normalize
from scipy.interpolate import interpn
def density_scatter( x , y, title, ax = None, sort = True, bins = 20, **kwargs ) :
"""
Scatter plot colored by 2d histogram
"""
nan_indices=[i[0] for i in np.argwhere(np.isnan(x))]
y= np.delete(y, nan_indices, 0)
x=np.delete(x,nan_indices,0)
if ax is None :
fig , ax = plt.subplots(figsize=(4, 3), dpi= 80, facecolor='w', edgecolor='k')
data , x_e, y_e = np.histogram2d( x, y, bins = bins, density = True )
z = interpn( ( 0.5*(x_e[1:] + x_e[:-1]) , 0.5*(y_e[1:]+y_e[:-1]) ) , data , np.vstack([x,y]).T , method = "splinef2d", bounds_error = False)
#To be sure to plot all data
z[np.where(np.isnan(z))] = 0.0
# Sort the points by density, so that the densest points are plotted last
if sort :
idx = z.argsort()
x, y, z = x[idx], y[idx], z[idx]
ax.scatter( x, y, c=z, **kwargs,s=1 )
lims = [
np.min([ax.get_xlim(), ax.get_ylim()]), # min of both axes
np.max([ax.get_xlim(), ax.get_ylim()]), # max of both axes
]
# now plot both limits against eachother
ax.plot(lims, lims, 'k-', alpha=0.75, zorder=0)
ax.set_aspect('equal')
ax.set_xlim(lims)
ax.set_ylim(lims)
plt.title(title)
plt.xlabel("Observed")
plt.ylabel("Predicted")
norm = Normalize(vmin = np.min(z), vmax = np.max(z))
cbar = fig.colorbar(cm.ScalarMappable(norm = norm), ax=ax)
cbar.ax.set_ylabel('Density')
return ax
# -
density_scatter(train_y,np.squeeze(predictions_train),'Joint Training: Training',bins=[30,30])
density_scatter(validate_y,np.squeeze(predictions_validate),'Joint Training: Validation',bins=[30,30])
density_scatter(test_y,np.squeeze(predictions_test),'Joint Training: Test',bins=[30,30])
import tensorflow as tf
from tensorflow.compat.v1.keras.backend import get_session
tf.compat.v1.disable_v2_behavior()
# select a set of background examples to take an expectation over
background=train_X
# +
import shap
# explain predictions of the model on four images
e = shap.DeepExplainer(model, background)
# -
shap_vals=e.shap_values(test_X)[0]
shap_vals[0][0].shape
| neural_net/.ipynb_checkpoints/train_predict_interpret_cnn_substrate_splits_train_neil1_ttyh2_test_ajuba-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.0 64-bit (''magic-touch-env-39'': pyenv)'
# language: python
# name: python3
# ---
# +
import pickle
from pprint import pprint
from dataclasses import dataclass
# +
@dataclass
class Position:
x: int
y: int
radius: int
@dataclass
class Contour:
x: int
y: int
radius: int
ss: float
@dataclass
class FrameData:
frame_number: int
analyzed: bool
frame_read_time: float = None
mask_creation_time: float = None
contour_find_time: float = None
num_contours: int = None
num_evaluated_contours: int = None
contour_evaluated_time: float = None
contour_evaluated_time_avg: float = None
ball_position: Position = None
# +
filename = 'spikeball3-with-ball.mov_framedata.p'
with open(filename, 'rb') as f:
framedata = pickle.load(f)
# +
avg_mask_creation_time = sum(f.mask_creation_time
for f in framedata) / len(framedata)
print(avg_mask_creation_time)
total_contour_find_time = sum(f.contour_find_time for f in framedata)
total_contours = sum(f.num_contours for f in framedata)
total_evaluated_contours = sum(f.num_evaluated_contours for f in framedata)
total_contour_evaluation_time = sum(f.contour_evaluated_time for f in framedata)
avg_contour_evaluated_time = total_contour_evaluation_time / total_evaluated_contours
print(avg_contour_evaluated_time)
print(total_evaluated_contours / len(framedata))
| analyze-data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import matplotlib.pyplot as plt
import numpy as np
import scipy.io
import time
# %matplotlib inline
# +
def diff(x,y):
return np.abs( np.mod( x - y + 90, 180) - 90 )
def G(x,y,sigma):
return np.exp(-1*diff(x,y)**2/(2*sigma**2))
def G2D(x_range, y_range, mean, sigma):
x0 = mean[0]
y0 = mean[1]
return np.exp( -1*( ( x_range-x0)**2 + (y_range-y0)**2) / (2*sigma**2) )
def mean_connections(W_ab):
total = 0.
for i in range(W_ab.shape[0]):
sub_mat = W_ab[i,:,:]
total = total + sub_mat[sub_mat != 0].size
return total / W_ab.shape[0]
def stimulus_size(x,length,sig_RF):
return (1.+np.exp(-(x + length/2.)/sig_RF) )**-1. * (1. - (1.+np.exp(-(x - length/2.)/sig_RF))**-1. )
# +
# Determine the connection probabilities
N_pairs = 75 # no. of E/I pairs to a side of a grid
field_size = 16. # size of field to a side (degrees)
dx = field_size / N_pairs
xy_range = np.linspace(0, field_size, N_pairs, False)
# xy_range = np.linspace(-field_size/2, field_size/2, N_pairs)
xv, yv = np.meshgrid(xy_range, xy_range) # x and y grid values (degrees)
# load from Bryan's code
data = scipy.io.loadmat('orientation-map.mat')
OP_map = data['map']
plt.imshow(OP_map)
plt.colorbar()
plt.title('Orientation Preferences')
# Connection weight parameters (from supp. materials S1.1.2):
kappa_E = 0.1
kappa_I = 0.5
J_EE = 0.1
J_IE = 0.38
J_EI = 0.089
J_II = 0.096
sig_EE = 8*dx
sig_IE = 12*dx
sig_EI = 4*dx
sig_II = 4*dx
sig_ori = 45
# calculate probability of no connections for each neuron (sparse connectivity)
G_EE = np.zeros((N_pairs**2, N_pairs, N_pairs))
G_IE = np.copy(G_EE)
# may not need these
G_EI = np.copy(G_EE)
G_II = np.copy(G_EE)
G_ori = np.copy(G_EE)
pW_EE = np.copy(G_EE)
pW_IE = np.copy(G_EE)
pW_EI = np.copy(G_EE)
pW_II = np.copy(G_EE)
rnd_EE = np.copy(G_EE)
rnd_IE = np.copy(G_EE)
rnd_EI = np.copy(G_EE)
rnd_II = np.copy(G_EE)
np.random.seed(1)
# iterate through each E/I pair:
for i in range(N_pairs):
for j in range(N_pairs):
G_EE[N_pairs*i+j, :, :] = G2D( xv, yv, (xv[0,i] , yv[j,0]), sig_EE)
G_IE[N_pairs*i+j, :, :] = G2D( xv, yv, (xv[0,i] , yv[j,0]), sig_IE)
G_EI[N_pairs*i+j, :, :] = G2D( xv, yv, (xv[0,i] , yv[j,0]), sig_EI)
G_II[N_pairs*i+j, :, :] = G2D( xv, yv, (xv[0,i] , yv[j,0]), sig_II)
G_ori[N_pairs*i+j,:,:] = G(OP_map[j,i], OP_map, sig_ori)
rnd_EE[N_pairs*i+j, :, :] = np.random.rand(N_pairs, N_pairs)
rnd_IE[N_pairs*i+j, :, :] = np.random.rand(N_pairs, N_pairs)
rnd_EI[N_pairs*i+j, :, :] = np.random.rand(N_pairs, N_pairs)
rnd_II[N_pairs*i+j, :, :] = np.random.rand(N_pairs, N_pairs)
for i in range(N_pairs**2):
pW_EE[i,:,:] = kappa_E * np.multiply(G_EE[i,:,:], G_ori[i,:,:])
pW_IE[i,:,:] = kappa_E * np.multiply(G_IE[i,:,:], G_ori[i,:,:])
pW_EI[i,:,:] = kappa_I * np.multiply(G_EI[i,:,:], G_ori[i,:,:])
pW_II[i,:,:] = kappa_I * np.multiply(G_II[i,:,:], G_ori[i,:,:])
# find zero-weighted connections:
W_EE = np.ones((N_pairs**2, N_pairs, N_pairs))
W_IE = np.copy(W_EE)
W_EI = np.copy(W_EE)
W_II = np.copy(W_EE)
W_EE[pW_EE<rnd_EE] = 0
W_IE[pW_IE<rnd_IE] = 0
W_EI[pW_EI<rnd_EI] = 0
W_II[pW_II<rnd_II] = 0
u_EE = mean_connections(W_EE)
u_IE = mean_connections(W_IE)
u_EI = mean_connections(W_EI)
u_II = mean_connections(W_II)
# -
# Print mean number of connections:
print u_EE, u_IE, u_EI, u_II
# +
# For non-zero connections, determine the weight
W_EE[W_EE != 0] = np.random.normal(J_EE, 0.25*J_EE, W_EE[W_EE!=0].size)
W_IE[W_IE != 0] = np.random.normal(J_IE, 0.25*J_IE, W_IE[W_IE!=0].size)
W_EI[W_EI != 0] = np.random.normal(J_EI, 0.25*J_EI, W_EI[W_EI!=0].size)
W_II[W_II != 0] = np.random.normal(J_II, 0.25*J_II, W_II[W_II!=0].size)
# W_EE[W_EE != 0] = np.random.normal(J_EE, (0.25*J_EE)**2, W_EE[W_EE!=0].size)
# W_IE[W_IE != 0] = np.random.normal(J_IE, (0.25*J_IE)**2, W_IE[W_IE!=0].size)
# W_EI[W_EI != 0] = np.random.normal(J_EI, (0.25*J_EI)**2, W_EI[W_EI!=0].size)
# W_II[W_II != 0] = np.random.normal(J_II, (0.25*J_II)**2, W_II[W_II!=0].size)
# Set negative weights to zero:
W_EE[W_EE < 0] = 0
W_IE[W_IE < 0] = 0
W_EI[W_EI < 0] = 0
W_II[W_II < 0] = 0
# W_EE = W_EE*1.15
# "Weights of a given type 'b' onto each unit
# are then scaled so that all units of a given type 'a' receive the same
# total type b synaptic weight, equal to Jab times the mean number of
# connections received under p (Wab(x, x′) ̸= 0)"
for i in range(N_pairs**2):
if np.all(W_EE[i,:,:] == np.zeros((N_pairs, N_pairs))) == False:
W_EE[i,:,:] = W_EE[i,:,:]*J_EE*u_EE/np.sum(W_EE[i,:,:])
if np.all(W_IE[i,:,:] == np.zeros((N_pairs, N_pairs))) == False:
W_IE[i,:,:] = W_IE[i,:,:]*J_IE*u_IE/np.sum(W_IE[i,:,:])
if np.all(W_EI[i,:,:] == np.zeros((N_pairs, N_pairs))) == False:
W_EI[i,:,:] = W_EI[i,:,:]*J_EI*u_EI/np.sum(W_EI[i,:,:])
if np.all(W_II[i,:,:] == np.zeros((N_pairs, N_pairs))) == False:
W_II[i,:,:] = W_II[i,:,:]*J_II*u_II/np.sum(W_II[i,:,:])
# +
# Model parameters (from supplementary methods)
sig_FF = 32
sig_RF = dx
k = np.random.normal(0.012, 0.05*0.012, (N_pairs, N_pairs))
n_E = np.random.normal(2.0, 0.05*2.0, (N_pairs, N_pairs))
n_I = np.random.normal(2.2, 0.05*2.2, (N_pairs, N_pairs))
tau_E = np.random.normal(0.02, 0.05*0.02, (N_pairs, N_pairs))
tau_I = np.random.normal(0.01, 0.05*0.01, (N_pairs, N_pairs))
# +
# From S.1.3.2: for strongest nonlinear behaviour, omega_E < 0 and omega_E < omega_I
# where omega_E = sum(W_II) - sum(W_EI), omega_I = sum(W_IE) - sum(W_EE)
# Verify here:
omega_E = np.sum(W_II) - np.sum(W_EI)
omega_I = np.sum(W_IE) - np.sum(W_EE)
print 'Omega_E: ', omega_E
print 'Omega_I: ', omega_I
if omega_E < 0 and omega_I > omega_E:
print "System should show strong nonlinear behaviour!"
else:
print "System may not show strong nonlinear behaviour."
# -
def generate_ext_stimulus(ori, size, centre, sig_RF=16./75, sig_FF = 32., fsize=16., full_frame=False):
G_FF = G(ori, OP_map, sig_FF)
v_range = np.linspace(0, fsize, N_pairs, False)
xv, yv = np.meshgrid( v_range, v_range )
if full_frame==True:
h = G_FF
else:
x_distance = np.abs(xv - centre[0])
y_distance = np.abs(yv - centre[1])
dist = np.sqrt(x_distance**2 + y_distance**2)
mask = stimulus_size(dist, size, sig_RF)
h = np.multiply( mask, G_FF )
return h
def run_simulation( dt, timesteps, c, h, k, n_E, n_I, tau_E, tau_I, init_cond=[np.zeros((N_pairs, N_pairs)),np.zeros((N_pairs, N_pairs))]):
r_E = np.zeros((timesteps, N_pairs, N_pairs))
r_I = np.copy(r_E)
# add initial conditions:
r_E[0,:,:] = init_cond[0]
r_I[0,:,:] = init_cond[1]
I_E = np.zeros((timesteps, N_pairs, N_pairs))
I_I = np.copy(I_E)
# rSS_E = np.copy(I_E)
# rSS_I = np.copy(I_I)
for t in range(1,timesteps):
# Input drive from external input and network
I_E[t,:,:] = c*h + np.sum( np.sum( W_EE * r_E[t-1,:,:],1 ), 1 ).reshape(N_pairs, N_pairs).T - np.sum( np.sum( W_EI * r_I[t-1,:,:],1 ), 1 ).reshape(N_pairs, N_pairs).T
I_I[t,:,:] = c*h + np.sum( np.sum( W_IE * r_E[t-1,:,:],1 ), 1 ).reshape(N_pairs, N_pairs).T - np.sum( np.sum( W_II * r_I[t-1,:,:],1 ), 1 ).reshape(N_pairs, N_pairs).T
# steady state firing rates - power law I/O
rSS_E = np.multiply(k, np.power(np.fmax(0,I_E[t,:,:]), n_E))
rSS_I = np.multiply(k, np.power(np.fmax(0,I_I[t,:,:]), n_I))
# set negative steady state rates to zero
rSS_E[rSS_E < 0] = 0
rSS_I[rSS_I < 0] = 0
# instantaneous firing rates approaching steady state
r_E[t,:,:] = r_E[t-1,:,:] + dt*(np.divide(-r_E[t-1,:,:]+rSS_E, tau_E))
r_I[t,:,:] = r_I[t-1,:,:] + dt*(np.divide(-r_I[t-1,:,:]+rSS_I, tau_I))
return [r_E, r_I, I_E, I_I]
# +
# run a single simulation for testing
r_units = np.floor(N_pairs*np.random.rand(5,2))
h = generate_ext_stimulus(45, 8, (8,8))
dt = 0.005
timesteps = 100
c = 40
start_t = time.time()
[r_E, r_I, I_E, I_I] = run_simulation(dt, timesteps, c, h, k, n_E, n_I, tau_E, tau_I)
end_t = time.time()
print "Elapsed simulation time: ", end_t - start_t
# plt.figure()
# plt.hold(True)
# for i in range(len(r_units)):
# plt.plot(np.linspace(0,dt*timesteps,timesteps), r_E[:,r_units[i,0],r_units[i,1]],'r')
# plt.title('Individual Excitatory Response')
# plt.xlabel('Time (seconds)')
# plt.ylabel('Firing Rate (Hz)')
plt.figure()
plt.imshow(r_E[-1,:,:])
plt.colorbar()
# +
# Plot results of single stimulus trial
plt.figure()
plt.hold(True)
rnd_units = np.floor( 75*np.random.rand( 10, 2 ) )
for i in range(10):
plt.plot(np.linspace(0,0.1,100), r_E[:,rnd_units[i,0],rnd_units[i,1]])
plt.title('Individual Excitatory Response')
plt.xlabel('Time (seconds)')
plt.ylabel('Firing Rate (Hz)')
print "Average E rate: ", np.mean(r_E)
print "Average I rate: ", np.mean(r_I)
print 'Maximum E Rate: ', np.max(r_E)
print 'Maximum I Rate: ', np.max(r_I)
plt.figure()
plt.hold(True)
for i in range(10):
plt.plot(np.linspace(0,0.1,100), r_I[:,rnd_units[i,0],rnd_units[i,1]])
plt.title('Individual Inhibitory Response')
plt.xlabel('Time (seconds)')
plt.ylabel('Firing Rate (Hz)')
plt.figure()
plt.imshow(OP_map)
print r_E[-1,:,:][np.floor(r_E[-1,:,:]) != 0].size
# +
# parameters for simulation to reproduce mean length tuning curves, figure 6E
size_range = np.linspace(1,16,10) # size
c = 40
stim_ori = 45 # degrees
centre = (8,8)
h_range = np.zeros((len(size_range), N_pairs, N_pairs))
for i in range(len(size_range)):
h_range[i,:,:] = generate_ext_stimulus(stim_ori, size_range[i], centre)
# plt.figure()
# plt.imshow(h_range[i,:,:])
# plt.title('Stimulus size: %d' % size_range[i])
# plt.colorbar()
dt = 0.005
timesteps = 100
# +
# run simulations to reproduce mean length tuning curves, figure 6E
# (this will take a long time to run)
# store all the firing rates for every trial
results_E = np.zeros((len(size_range), timesteps, N_pairs, N_pairs))
results_I = np.copy(results_E)
results_I_E = np.copy(results_E)
results_I_I = np.copy(results_E)
for i in range(len(size_range)):
[r_E, r_I, I_E, I_I] = run_simulation(dt, timesteps, c, h_range[i])
results_E[i,:,:,:] = r_E
results_I[i,:,:,:] = r_I
results_I_E[i,:,:,:] = I_E
results_I_I[i,:,:,:] = I_I
# +
# Find which units demonstrate SSI > 0.25:
# first, find max firing rates for stimuli shorter than 2/3 * 16 degrees:
trials = size_range[ size_range < 2./3*field_size ].size
# r_max - maximum firing rate to stimuli shorter tha (2/3)*16 degrees
# r_max_E = np.max( np.max( results_E[:trials, :, :, :], 1), 0 )
r_max_E = np.max( results_E[:trials, -1, :, :], 0)
# r_full - response to the largest stimulus
r_full_E = results_E[-1, -1, :, :]
SSI_E = (r_max_E - r_full_E) / r_max_E
print 'Number of E units showing strong surround suppression: ', SSI_E[SSI_E > 0.25].size
# r_max_I = np.max( np.max( results_I[:trials, :, :, :], 1), 0 )
r_max_I = np.max( results_I[:trials, -1, :, :], 0)
r_full_I = results_I[-1, -1, :, :]
SSI_I = (r_max_I - r_full_I) / r_max_I
print 'Number of I units showing strong surround suppression: ', SSI_I[SSI_I > 0.25].size
y_I, x_I = np.where(SSI_I > 0.25)
y_E, x_E = np.where(SSI_E > 0.25)
avg_r_E = np.zeros(len(size_range))
avg_r_I = np.copy(avg_r_E)
max_r_E = np.copy(avg_r_E)
max_r_I = np.copy(avg_r_E)
SS_r_E = np.copy(avg_r_E)
SS_r_I = np.copy(avg_r_E)
for i in range(len(size_range)):
avg_r_E[i] = np.mean( results_E[i, :, y_E, x_E] )
avg_r_I[i] = np.mean( results_I[i, :, y_I, x_I] )
max_r_E[i] = np.mean(np.max( results_E[i, :, y_E, x_E], 0 ))
max_r_I[i] = np.mean(np.max( results_I[i, :, y_I, x_I], 0 ))
SS_r_E[i] = np.mean( results_E[i, -1, y_E, x_E] )
SS_r_I[i] = np.mean( results_I[i, -1, y_I, x_I] )
# plt.figure()
# plt.plot(size_range, avg_r_E, 'r', size_range, avg_r_I, 'b')
# plt.ylabel('Average Firing Rate')
# plt.xlabel('Stimulus Size (degrees)')
# plt.figure()
# plt.plot(size_range, max_r_E, 'r', size_range, max_r_I, 'b')
# plt.ylabel('Max Firing Rate')
# plt.xlabel('Stimulus Size (degrees)')
plt.figure()
plt.plot(size_range, SS_r_E, 'r', size_range, SS_r_I, 'b')
plt.ylabel('SS Firing Rate')
plt.xlabel('Stimulus Size (degrees)')
plt.title('Responses of units showing surround suppression')
r_units = np.floor(N_pairs*np.random.rand(14,2))
plt.figure()
plt.hold(True)
for i in range(len(r_units)):
plt.plot(size_range, results_E[:, -1, r_units[i,0], r_units[i,1]], 'r')
plt.ylabel('SS Firing Rate')
plt.xlabel('Stimulus Size (degrees)')
plt.title('Responses of %d randomly selected E units' % len(r_units))
plt.figure()
plt.hold(True)
for i in range(len(r_units)):
plt.plot(size_range, results_I[:, -1, r_units[i,0], r_units[i,1]], 'b')
plt.ylabel('SS Firing Rate')
plt.xlabel('Stimulus Size (degrees)')
plt.title('Responses of %d randomly selected I units' % len(r_units))
plt.figure()
plt.title('Positions of selected units')
plt.scatter(r_units[:,1], r_units[:,0])
# +
# Reproduction of figure 6F - length tuning for various levels of stimulus strength:
dt = 0.005
timesteps = 100
# c_range = np.array([3,10,17,31,52])
c_range = np.array([31,52])
k = np.random.normal(0.012, 0.05*0.012, (N_pairs, N_pairs))
n_E = np.random.normal(2.0, 0.05*2.0, (N_pairs, N_pairs))
n_I = np.random.normal(2.2, 0.05*2.2, (N_pairs, N_pairs))
tau_E = np.random.normal(0.02, 0.05*0.02, (N_pairs, N_pairs))
tau_I = np.random.normal(0.01, 0.05*0.01, (N_pairs, N_pairs))
# select 14 random units:
n_units = 14
r_units = np.floor( N_pairs*np.random.rand(n_units,2) )
size_range = np.linspace(1,16,10)
h_range = np.zeros((n_units, len(size_range), N_pairs, N_pairs))
start_time = time.time()
for i in range(len(size_range)):
for j in range(n_units):
yi = r_units[j,0]
xi = r_units[j,1]
h_range[j,i,:,:] = generate_ext_stimulus(OP_map[yi,xi], size_range[i], (dx*xi,dx*yi))
print "Time to generate stimuli: ", time.time() - start_time
# store all the firing rates for every trial
results_E = np.zeros((n_units, len(c_range), len(size_range), timesteps, N_pairs, N_pairs))
results_I = np.copy(results_E)
results_I_E = np.copy(results_E)
results_I_I = np.copy(results_E)
start_time = time.time()
for m in range(n_units):
for i in range(len(c_range)):
c = c_range[i]
for j in range(len(size_range)):
[r_E, r_I, I_E, I_I] = run_simulation(dt, timesteps, c, h_range[m,j,:,:], k, n_E, n_I, tau_E, tau_I)
results_E[m,i,j,:,:,:] = r_E
results_I[m,i,j,:,:,:] = r_I
results_I_E[m,i,j,:,:,:] = I_E
results_I_I[m,i,j,:,:,:] = I_I
end_time = time.time()
print "Elapsed simulation time: ", end_time - start_time
# +
# plotting for figure 6F:
for i in range(len(c_range)):
plt.figure()
plt.title( "Stimulus strength: %d" % c_range[i] )
plt.xlabel( "Stimulus Size (degrees)" )
plt.ylabel( "Firing Rate (Hz)" )
for j in range(n_units):
plt.plot( size_range, results_E[j,i,:,-1,r_units[j,0],r_units[j,1] ], 'r' )
plt.plot( size_range, results_I[j,i,:,-1,r_units[j,0],r_units[j,1] ], 'b' )
plt.savefig('results/fig-6F-str-%d' % (c_range[i]) )
# +
# Reproducing figure 6G - showing summation field size shrinking with
# increasing stimulus strength
# first, randomly select 100 units
n_units = 100
r_units = np.floor( N_pairs*np.random.rand(n_units,2) )
# get steady-state results from simulation results:
SS_r_E = np.zeros( ( len(c_range), len(h_range), n_units ) )
SS_r_I = np.copy(SS_r_E)
for j in range(len(c_range)):
for i in range(len(h_range)):
for k in range(n_units):
SS_r_E[j,i,k] = results_E[j,i,-1,r_units[k,0], r_units[k,1]]
SS_r_I[j,i,k] = results_I[j,i,-1,r_units[k,0], r_units[k,1]]
def find_first_peak(data):
max_val = data[0]
for i in range(len(data)):
if data[i] >= max_val:
max_val = data[i]
else:
break
return max_val
peaks_E = np.zeros( ( len(c_range), n_units ) )
peaks_I = np.copy(peaks_E)
for j in range(len(c_range)):
for i in range(n_units):
peaks_E[j, i] = find_first_peak( SS_r_E[j, :, i ] )
peaks_I[j, i] = find_first_peak( SS_r_I[j, :, i ] )
avg_peak_E = np.zeros( len(c_range) )
avg_peak_I = np.copy(avg_peak_E)
sd_peak_E = np.copy(avg_peak_E)
sd_peak_I = np.copy(avg_peak_E)
avg_peak_E = np.mean(peaks_E, 1)
sd_peak_E = np.std(peaks_E, 1)
avg_peak_I = np.mean(peaks_I, 1)
sd_peak_I = np.std(peaks_I, 1)
plt.figure()
plt.errorbar(c_range, avg_peak_E, sd_peak_E, fmt='r')
plt.xlabel('Stimulus Strength')
plt.ylabel('Summation Field Size')
plt.title('Excitatory summation Field Size vs. Stim. Strength')
plt.figure()
plt.errorbar(c_range, avg_peak_I, sd_peak_I)
plt.xlabel('Stimulus Strength')
plt.ylabel('Summation Field Size')
plt.title('Inhibitory summation Field Size vs. Stim. Strength')
# +
# Recreate figure 6B - showing network transition from externally driven to network-driven
# with increasing stimulus strength
# First, run simulations with full field gratings and several contrast strengths:
c_range = np.linspace(1, 50, 10)
dt = 0.005
timesteps = 100
stim_pref = 45
ff_stim = generate_ext_stimulus(stim_pref, 0, 0, full_frame=True)
results_E = np.zeros((len(c_range), timesteps, N_pairs, N_pairs))
results_I = np.copy(results_E)
results_I_E = np.copy(results_E)
results_I_I = np.copy(results_E)
start_time = time.time()
for i in range(len(c_range)):
[r_E, r_I, I_E, I_I] = run_simulation(dt, timesteps, c_range[i], ff_stim, k, n_E, n_I, tau_E, tau_I)
results_E[i,:,:,:] = r_E
results_I[i,:,:,:] = r_I
results_I_E[i,:,:,:] = I_E
results_I_I[i,:,:,:] = I_I
end_time = time.time()
print "Elapsed simulation time: ", end_time-start_time
# Plotting
# f_I_E = np.sum(np.sum(np.abs(results_I_E[:,-1,:,:]), 2), 1)
# f_I_I = np.sum(np.sum(np.abs(results_I_I[:,-1,:,:]), 2), 1)
# plt.figure()
# ext_input = np.zeros(len(c_range))
# for i in range(len(c_range)):
# ext_input[i] = np.sum(c_range[i]*ff_stim)
# f_I_E[i] = f_I_E[i] - ext_input[i]
# f_I_I[i] = f_I_I[i] - ext_input[i]
# f_I_E = np.abs(f_I_E)
# f_I_I = np.abs(f_I_I)
# plt.plot( c_range, f_I_E/(ext_input+f_I_E), 'r')
# plt.plot( c_range, ext_input/(ext_input+f_I_E), 'r--')
# plt.plot( c_range, f_I_I/(ext_input+f_I_I), 'b')
# plt.plot( c_range, ext_input/(ext_input+f_I_I), 'b--')
# plt.title('Responses to full-field stimuli')
# plt.xlabel('Input Strength')
# plt.ylabel('Percent of Input')
# +
# Plotting for figure 6B
# Find all units with orientation preference within 10 degrees of the stimulus orientation
pref_units = np.where( np.logical_and(OP_map > stim_pref - 5, OP_map < stim_pref + 5) )
# randomly select 25 of those units to average over
r_units = np.floor( pref_units[0].size*np.random.rand( 25 ) )
r_units = r_units.astype(int)
unit_idx_x = pref_units[1][r_units]
unit_idx_y = pref_units[0][r_units]
# Sanity check for x,y ordering
# print OP_map[unit_idx_y, unit_idx_x]
# plt.figure()
# plt.imshow(np.logical_and(OP_map > stim_pref - 5, OP_map < stim_pref + 5))
# plt.scatter(unit_idx_x, unit_idx_y)
net_E = np.zeros((len(c_range), len(r_units)))
net_I = np.copy(net_E)
for i in range(len(c_range)):
for j in range(len(r_units)):
net_E[i,j] = np.sum(np.multiply(results_E[i,-1,:,:],W_EE[unit_idx_y[j] + N_pairs*unit_idx_x[j],:,:])) + np.sum(np.multiply(results_I[i,-1,:,:],W_EI[unit_idx_y[j] + N_pairs*unit_idx_x[j],:,:]))
net_I[i,j] = np.sum(np.multiply(results_E[i,-1,:,:],W_IE[unit_idx_y[j] + N_pairs*unit_idx_x[j],:,:])) + np.sum(np.multiply(results_I[i,-1,:,:],W_II[unit_idx_y[j] + N_pairs*unit_idx_x[j],:,:]))
external_drive = np.tile(ff_stim[unit_idx_y, unit_idx_x], (len(c_range), 1))*c_range[:, np.newaxis]
print external_drive.shape
net_percent_E = net_E / (net_E + external_drive)
net_percent_I = net_I / (net_I + external_drive)
ext_percent_E = external_drive / (net_E + external_drive)
ext_percent_I = external_drive / (net_I + external_drive)
plt.figure()
plt.plot( c_range, np.mean( net_percent_E, 1), 'r')
plt.plot( c_range, np.mean( ext_percent_E, 1), 'r--')
plt.plot( c_range, np.mean( net_percent_I, 1), 'b')
plt.plot( c_range, np.mean( ext_percent_I, 1), 'b--')
plt.title('Responses to full-field stimuli')
plt.xlabel('Input Strength')
plt.ylabel('Percent of Input')
# +
# Recreate figure 6C (run simulations for figure 6B above first)
# is this equal to the final firing rate * the appropriate weight?
E_n_input_E = np.zeros((len(c_range),len(r_units)))
I_input_E = np.copy(E_n_input_E)
E_n_input_I = np.copy(E_n_input_E)
I_input_I = np.copy(E_n_input_E)
for i in range(len(c_range)):
for j in range(len(r_units)):
E_n_input_E[i, j] = np.sum(results_E[i,-1,ry[j], rx[j]] * W_EE[ry[j]+N_pairs*rx[j], :, :])
E_n_input_I[i, j] = np.sum(results_E[i,-1,ry[j], rx[j]] * W_IE[ry[j]+N_pairs*rx[j], :, :])
I_input_E[i, j] = np.sum(results_I[i,-1,ry[j], rx[j] ] * W_EI[ry[j]+N_pairs*rx[j], :, :])
I_input_I[i, j] = np.sum(results_I[i,-1,ry[j], rx[j] ] * W_II[ry[j]+N_pairs*rx[j], :, :])
mean_E = np.zeros(len(c_range))
mean_I = np.copy(mean_E)
sd_E = np.copy(mean_E)
sd_I = np.copy(mean_E)
for i in range(len(c_range)):
mean_E[i] = np.mean(E_n_input_E[i,:] / ( I_input_E[i,:] + E_n_input_E[i,:] ))
mean_I[i] = np.mean(E_n_input_I[i,:] / ( I_input_I[i,:] + E_n_input_I[i,:] ))
sd_E = np.std(E_n_input_E[i,:] / ( I_input_E[i,:] + E_n_input_E[i,:] ))
sd_I = np.std(E_n_input_I[i,:] / ( I_input_I[i,:] + E_n_input_I[i,:] ))
plt.figure()
plt.ylabel('E_n/(E_n+I)')
plt.xlabel('Stimulus Strength')
plt.errorbar(c_range, mean_I, sd_I)
plt.errorbar(c_range, mean_E, sd_E, fmt='r')
plt.savefig('figure6c.png')
# -
print W_EE[ry[j]+N_pairs*rx[j], :, :]
# +
# Recreate figure 6D
c_range = np.linspace(1,90,10)
| mechanistic/old_stuff/SSN_2D_model-June-17.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.1 64-bit (''OED'': conda)'
# name: python3
# ---
# # predictive uncertainty analysis
# import packages
import numpy as np
import pandas as pd
import matplotlib as mpl
import matplotlib.pyplot as plt
import seaborn as sns
import math
from matplotlib.ticker import FuncFormatter
import matplotlib.ticker as mtick
mpl.rcParams['font.size'] = 16
mpl.rcParams['savefig.bbox'] = 'tight'
mpl.rcParams['savefig.format'] = 'pdf'
mpl.rcParams['axes.labelsize'] = 20
mpl.rcParams['xtick.labelsize'] = 20
mpl.rcParams['ytick.labelsize'] = 20
mpl.rcParams['legend.fontsize'] = 20
# import the annual loads
file_date = '20220116'
fpath = f'../../output/work_run_{file_date}/'
fn = '126001A.3.obs.csv'
fn_meas = '126001A.base.obs.csv'
log_load = True
# +
df = pd.read_csv(fpath + fn, index_col = 'real_name')
# select results of which the pbias is with 15%
# df = df[(df.din_pbias < 15) & (df.din_pbias > -15)]
df_meas = pd.read_csv(fpath + fn_meas, index_col = 'real_name')
if log_load:
df_meas.loc[:, 'din_2009':] = 10**(df_meas.loc[:, 'din_2009':])
df.loc[:, 'din_2009':] = 10**(df.loc[:, 'din_2009':])
df['average'] = df.loc[:, 'din_2009':'din_2017'].mean(axis=1).values
df_meas['average'] = df_meas.loc[:, 'din_2009':'din_2017'].mean(axis=1).values
# obs data
obs_annual = [52.093, 99.478, 44.064, 57.936, 53.449, 21.858, 38.561, 51.843, 14.176]
obs_annual.append(np.round(np.mean(obs_annual), 2))
obs_df = pd.DataFrame(data=obs_annual, index = [*np.arange(2009, 2018), 'average'], columns=['Annual loads'])
# +
# reorganize the dataframe for plotting
df_plot = pd.DataFrame(data = df.values[:, 1:].T.flatten(), columns=['Annual loads'])
year_col = np.repeat(np.arange(2009, 2019), df.shape[0], 0).T
df_plot['year'] = year_col
df_plot['type'] = 'Estimate'
df_meas_plot = pd.DataFrame(data = df_meas.values[:, 1:].T.flatten(), columns=['Annual loads'])
year_col = np.repeat(np.arange(2009, 2019), df_meas.shape[0], 0).T
df_meas_plot['year'] = year_col
df_meas_plot['type'] = 'Measurement realization'
df_plot = pd.concat([df_meas_plot, df_plot])
df_plot.reset_index().tail()
# -
# Plot the uncertainty of annual loads
sns.set_style('whitegrid')
fig = plt.figure(figsize=(10, 6))
ax = sns.violinplot(x='year', y='Annual loads', data=df_plot, hue='type', split=True);
ax.legend(handles=ax.legend_.legendHandles, labels=['Measurement', 'Estimate'])
# obs_df.plot(y=[0], linewidth=0, ax=ax, marker='d', markersize=5, color='r', legend=['Obs']);
ax.set_xticklabels([*np.arange(2009, 2018), 'average'], rotation=90);
plt.gca().yaxis.set_major_formatter(mtick.FormatStrFormatter('%.2e'))
ax.set_xlabel('Year')
ax.set_ylabel('Annual loads(KG)')
# plt.savefig(f'../../output/figs/predict_uncertainty_{file_date}.png', dpi=300, format='png')
sns.set_style('darkgrid', {"axes.facecolor": ".9"})
df_temp = df - df_meas
ax = df_temp.loc[:, 'din_2009':].T.plot(legend=False, figsize=(20, 6), use_index=True, xlabel='Year', ylabel='Annual loads(KG)')
ax.set_xticks(range(len(df.columns[1:])))
obs_df.plot(y=[0], linewidth=0, ax=ax, marker='d', markersize=5, color='r', legend=['Obs']);
ax.set_xticklabels([*np.arange(2009, 2018), 'average']);
# plt.savefig(f'../../output/figs/predict_uncertainty_lineplot_{file_date}.png', dpi=300, format='png')
| src/notebooks/predictive_uncertainty.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # DA0101 course
#
# ### "Can we estimate the price of a used car based on its characteristics?"
# 1st: Datas => https://archive.ics.uci.edu/ml/machine-learning-databases/autos/imports-85.data
# #### Describe of columns:
# ```
# Attribute: Attribute Range:
# ------------------ -----------------------------------------------
# 1. symboling: -3, -2, -1, 0, 1, 2, 3.
# 2. normalized-losses: continuous from 65 to 256.
# 3. make: alfa-romero, audi, bmw, chevrolet, dodge, honda,
# isuzu, jaguar, mazda, mercedes-benz, mercury,
# mitsubishi, nissan, peugot, plymouth, porsche,
# renault, saab, subaru, toyota, volkswagen, volvo
# 4. fuel-type: diesel, gas.
# 5. aspiration: std, turbo.
# 6. num-of-doors: four, two.
# 7. body-style: hardtop, wagon, sedan, hatchback, convertible.
# 8. drive-wheels: 4wd, fwd, rwd.
# 9. engine-location: front, rear.
# 10. wheel-base: continuous from 86.6 120.9.
# 11. length: continuous from 141.1 to 208.1.
# 12. width: continuous from 60.3 to 72.3.
# 13. height: continuous from 47.8 to 59.8.
# 14. curb-weight: continuous from 1488 to 4066.
# 15. engine-type: dohc, dohcv, l, ohc, ohcf, ohcv, rotor.
# 16. num-of-cylinders: eight, five, four, six, three, twelve, two.
# 17. engine-size: continuous from 61 to 326.
# 18. fuel-system: 1bbl, 2bbl, 4bbl, idi, mfi, mpfi, spdi, spfi.
# 19. bore: continuous from 2.54 to 3.94.
# 20. stroke: continuous from 2.07 to 4.17.
# 21. compression-ratio: continuous from 7 to 23.
# 22. horsepower: continuous from 48 to 288.
# 23. peak-rpm: continuous from 4150 to 6600.
# 24. city-mpg: continuous from 13 to 49.
# 25. highway-mpg: continuous from 16 to 54.
# 26. price: continuous from 5118 to 45400.
# ```
import pandas as pd
url_from = 'https://archive.ics.uci.edu/ml/machine-learning-databases/autos/imports-85.data'
df_cars = pd.read_csv(url_from,header=None)
df_cars.head()
df_cars.tail(3)
headres = [ 'symboling', 'normalized-losses', 'make', 'fuel-type' , ' aspiration',
'num-of-doors','body-style', 'drive-wheels', 'engine-location' , 'wheel-base',
'length','width', 'height', 'curb-weight', 'engine-type', 'num-of-cylinders', 'engine-size', 'fuel-system',
'bore', 'stroke', 'compression-ratio', 'horsepower', 'peak-rpm','city-mpg', 'highway-mpg','price']
df_cars.columns = headres
df_cars.head()
test_file_csv = 'df_cars_1985.csv'
test_file_jsn = 'df_cars_1985.json'
df_cars.to_csv(test_file_csv)
df_cars.to_json(test_file_jsn)
# sprawdzam typy danych
df_cars.dtypes
# średnie wartości...
df_cars.describe()
df_cars.describe(include='all')
# another info....
df_cars.info()
# top 30 and bottom 30 rows...
df_cars.info
# wycinanie z tabeli...
df_cars[['length','compression-ratio']].head()
df_cars[['length','compression-ratio']].describe()
# drop NaN elements... in place
df_cars.dropna(inplace=True)
df_cars.info()
| DA0101-01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # HackerRank challenges
import math
import os
import random
import re
import sys
from collections import Counter
# ## Warmup challenges
# #### 1. Sock Merchant (easy)
# John works at a clothing store. He has a large pile of socks that he must pair by color for sale. Given an array of integers representing the color of each sock, determine how many pairs of socks with matching colors there are.
#
# For example, there are socks with colors . There is one pair of color and one of color . There are three odd socks left, one of each color. The number of pairs is .
#
# **Function Description**
#
# Complete the sockMerchant function in the editor below. It must return an integer representing the number of matching pairs of socks that are available.
#
# **sockMerchant has the following parameter(s):**
#
# n: the number of socks in the pile
#
# ar: the colors of each sock
def sockMerchant(n, ar):
socks, pairs = Counter(map(int,ar)), 0
for sock in socks:
pairs += socks[sock]//2
print(pairs)
# #### 2. Counting Valleys
# Gary is an avid hiker. He tracks his hikes meticulously, paying close attention to small details like topography. During his last hike he took exactly steps. For every step he took, he noted if it was an uphill, , or a downhill, step. Gary's hikes start and end at sea level and each step up or down represents a unit change in altitude. We define the following terms:
#
# A mountain is a sequence of consecutive steps above sea level, starting with a step up from sea level and ending with a step down to sea level.
# A valley is a sequence of consecutive steps below sea level, starting with a step down from sea level and ending with a step up to sea level.
# Given Gary's sequence of up and down steps during his last hike, find and print the number of valleys he walked through.
#
# For example, if Gary's path is , he first enters a valley units deep. Then he climbs out an up onto a mountain units high. Finally, he returns to sea level and ends his hike.
#
# **Function Description**
#
# Complete the countingValleys function in the editor below. It must return an integer that denotes the number of valleys Gary traversed.
#
# **countingValleys has the following parameter(s):**
#
# n: the number of steps Gary takes
#
# s: a string describing his path
def countingValleys(n, s):
level = valleys = 0
for step in s:
level += 1 if step == "U" else -1
valleys += level == 0 and step == "U"
return valleys
# #### 3. Jumping on the clouds
# Emma is playing a new mobile game that starts with consecutively numbered clouds. Some of the clouds are thunderheads and others are cumulus. She can jump on any cumulus cloud having a number that is equal to the number of the current cloud plus or . She must avoid the thunderheads. Determine the minimum number of jumps it will take Emma to jump from her starting postion to the last cloud. It is always possible to win the game.
#
# For each game, Emma will get an array of clouds numbered if they are safe or if they must be avoided. For example, indexed from . The number on each cloud is its index in the list so she must avoid the clouds at indexes and . She could follow the following two paths: or . The first path takes jumps while the second takes .
#
# **Function Description**
#
# Complete the jumpingOnClouds function in the editor below. It should return the minimum number of jumps required, as an integer.
#
# **jumpingOnClouds has the following parameter(s):**
#
# c: an array of binary integers
def jumpingOnClouds(c):
i = count_jumps = 0
length = len(c)
while i < length - 1:
if i < length - 2 and c[i + 2] == 0:
i += 2
else:
i += 1
count_jumps += 1
return count_jumps
# #### 4. Repeated strings
# Lilah has a string, , of lowercase English letters that she repeated infinitely many times.
#
# Given an integer, , find and print the number of letter a's in the first letters of Lilah's infinite string.
#
# For example, if the string and , the substring we consider is , the first characters of her infinite string. There are occurrences of a in the substring.
#
# **Function Description**
#
# Complete the repeatedString function in the editor below. It should return an integer representing the number of occurrences of a in the prefix of length in the infinitely repeating string.
#
# **repeatedString has the following parameter(s):**
#
# s: a string to repeat
#
# n: the number of characters to consider
def repeatedString(s, n):
print(s.count("a") * (n // len(s)) + s[:n % len(s)].count("a"))
# ## Arrays
# #### 1. 2D array - DS
# Given a 2D Array, :
#
# 1 1 1 0 0 0
#
# 0 1 0 0 0 0
#
# 1 1 1 0 0 0
#
# 0 0 0 0 0 0
#
# 0 0 0 0 0 0
#
# 0 0 0 0 0 0
#
# We define an hourglass in to be a subset of values with indices falling in this pattern in 's graphical representation:
#
# a b c
#
# d
#
# e f g
#
# There are hourglasses in , and an hourglass sum is the sum of an hourglass' values. Calculate the hourglass sum for every hourglass in , then print the maximum hourglass sum.
#
# For example, given the 2D array:
#
# -9 -9 -9 1 1 1
#
# 0 -9 0 4 3 2
#
# -9 -9 -9 1 2 3
#
# 0 0 8 6 6 0
#
# 0 0 0 -2 0 0
#
# 0 0 1 2 4 0
#
# We calculate the following hourglass values:
#
# -63, -34, -9, 12,
# -10, 0, 28, 23,
# -27, -11, -2, 10,
# 9, 17, 25, 18
#
# Our highest hourglass value is from the hourglass:
#
# 0 4 3
#
# 1
#
# 8 6 6
#
# **Function Description**
#
# Complete the function hourglassSum in the editor below. It should return an integer, the maximum hourglass sum in the array.
#
# **hourglassSum has the following parameter(s):**
#
# arr: an array of integers
def hourglassSum(arr):
count = -64
row = 0
col = 0
while row < 4 :
temp = arr[row][col] + arr[row][col+1]+arr[row][col+2]+arr[row+1][col+1] + arr[row+2][col]+arr[row+2][col+1]+ arr[row+2][col+2]
if temp > count:
count = temp
col +=1
if col == 4:
col = 0
row +=1
return count
# #### 2. Left Rotation
# A left rotation operation on an array shifts each of the array's elements unit to the left. For example, if left rotations are performed on array , then the array would become .
#
# Given an array of integers and a number, , perform left rotations on the array. Return the updated array to be printed as a single line of space-separated integers.
#
# **Function Description**
#
# Complete the function rotLeft in the editor below. It should return the resulting array of integers.
#
# **rotLeft has the following parameter(s):**
#
# An array of integers .
# An integer , the number of rotations.
def rotLeft(a, k):
alist = list(a)
b = alist[k:]+alist[:k]
return b
# #### 3. New Year Chaos
# It's New Year's Day and everyone's in line for the Wonderland rollercoaster ride! There are a number of people queued up, and each person wears a sticker indicating their initial position in the queue. Initial positions increment by from at the front of the line to at the back.
#
# Any person in the queue can bribe the person directly in front of them to swap positions. If two people swap positions, they still wear the same sticker denoting their original places in line. One person can bribe at most two others. For example, if and bribes , the queue will look like this: .
#
# Fascinated by this chaotic queue, you decide you must know the minimum number of bribes that took place to get the queue into its current state!
#
# **Function Description**
#
# Complete the function minimumBribes in the editor below. It must print an integer representing the minimum number of bribes necessary, or Too chaotic if the line configuration is not possible.
#
# **minimumBribes has the following parameter(s):**
#
# q: an array of integers
def minimumBribes(Q):
#
# initialize the number of moves
moves = 0
#
# decrease Q by 1 to make index-matching more intuitive
# so that our values go from 0 to N-1, just like our
# indices. (Not necessary but makes it easier to
# understand.)
Q = [P-1 for P in Q]
#
# Loop through each person (P) in the queue (Q)
for i,P in enumerate(Q):
# i is the current position of P, while P is the
# original position of P.
#
# First check if any P is more than two ahead of
# its original position
if P - i > 2:
print("Too chaotic")
return
#
# From here on out, we don't care if P has moved
# forwards, it is better to count how many times
# P has RECEIVED a bribe, by looking at who is
# ahead of P. P's original position is the value
# of P.
# Anyone who bribed P cannot get to higher than
# one position in front if P's original position,
# so we need to look from one position in front
# of P's original position to one in front of P's
# current position, and see how many of those
# positions in Q contain a number large than P.
# In other words we will look from P-1 to i-1,
# which in Python is range(P-1,i-1+1), or simply
# range(P-1,i). To make sure we don't try an
# index less than zero, replace P-1 with
# max(P-1,0)
for j in range(max(P-1,0),i):
if Q[j] > P:
moves += 1
print(moves)
# #### 4. Minimum swaps 2
# You are given an unordered array consisting of consecutive integers [1, 2, 3, ..., n] without any duplicates. You are allowed to swap any two elements. You need to find the minimum number of swaps required to sort the array in ascending order.
#
# For example, given the array we perform the following steps:
#
# i arr swap (indices)
#
# 0 [7, 1, 3, 2, 4, 5, 6] swap (0,3)
#
# 1 [2, 1, 3, 7, 4, 5, 6] swap (0,1)
#
# 2 [1, 2, 3, 7, 4, 5, 6] swap (3,4)
#
# 3 [1, 2, 3, 4, 7, 5, 6] swap (4,5)
#
# 4 [1, 2, 3, 4, 5, 7, 6] swap (5,6)
#
# 5 [1, 2, 3, 4, 5, 6, 7]
#
# It took swaps to sort the array.
#
# **Function Description**
#
# Complete the function minimumSwaps in the editor below. It must return an integer representing the minimum number of swaps to sort the array.
#
# **minimumSwaps has the following parameter(s):**
#
# arr: an unordered array of integers
def minimumSwaps(arr):
res = 0
arr = [x-1 for x in arr]
value_idx = {x:i for i, x in enumerate(arr)}
for i, x in enumerate(arr):
if i != x:
to_swap_idx = value_idx[i]
arr[i], arr[to_swap_idx] = i, x
value_idx[i] = i
value_idx[x] = to_swap_idx
res += 1
print(res)
return res
# #### 5. Array Manipulation
# Starting with a 1-indexed array of zeros and a list of operations, for each operation add a value to each of the array element between two given indices, inclusive. Once all operations have been performed, return the maximum value in your array.
#
# For example, the length of your array of zeros . Your list of queries is as follows:
#
# a b k
#
# 1 5 3
#
# 4 8 7
#
# 6 9 1
#
# Add the values of between the indices and inclusive:
#
# index-> 1 2 3 4 5 6 7 8 9 10
#
# [0,0,0, 0, 0,0,0,0,0, 0]
#
# [3,3,3, 3, 3,0,0,0,0, 0]
#
# [3,3,3,10,10,7,7,7,0, 0]
#
# [3,3,3,10,10,8,8,8,1, 0]
#
# The largest value is after all operations are performed.
#
# **Function Description**
#
# Complete the function arrayManipulation in the editor below. It must return an integer, the maximum value in the resulting array.
#
# **arrayManipulation has the following parameters:**
#
# n - the number of elements in your array
#
# queries - a two dimensional array of queries where each queries[i] contains three integers, a, b, and k.
def arrayManipulation(n, queries):
array = [0] * (n + 1)
for query in queries:
a = query[0] - 1
b = query[1]
k = query[2]
array[a] += k
array[b] -= k
max_value = 0
running_count = 0
for i in array:
running_count += i
if running_count > max_value:
max_value = running_count
return max_value
# ## Dictionnaries and hashmaps
# #### 1. Ransom note
# Harold is a kidnapper who wrote a ransom note, but now he is worried it will be traced back to him through his handwriting. He found a magazine and wants to know if he can cut out whole words from it and use them to create an untraceable replica of his ransom note. The words in his note are case-sensitive and he must use only whole words available in the magazine. He cannot use substrings or concatenation to create the words he needs.
#
# Given the words in the magazine and the words in the ransom note, print Yes if he can replicate his ransom note exactly using whole words from the magazine; otherwise, print No.
#
# For example, the note is "Attack at dawn". The magazine contains only "attack at dawn". The magazine has all the right words, but there's a case mismatch. The answer is .
#
# **Function Description**
#
# Complete the checkMagazine function in the editor below. It must print if the note can be formed using the magazine, or .
#
# **checkMagazine has the following parameters:**
#
# magazine: an array of strings, each a word in the magazine
#
# note: an array of strings, each a word in the ransom note
def ransom_note(magazine, rasom):
return (Counter(rasom) - Counter(magazine)) == {}
# #### 2. Two Strings
# Given two strings, determine if they share a common substring. A substring may be as small as one character.
#
# For example, the words "a", "and", "art" share the common substring . The words "be" and "cat" do not share a substring.
#
# **Function Description**
#
# Complete the function twoStrings in the editor below. It should return a string, either YES or NO based on whether the strings share a common substring.
#
# **twoStrings has the following parameter(s):**
#
# s1, s2: two strings to analyze .
def twoStrings(s1, s2):
return 'YES' if set(s1) & set(s2) else 'NO'
# #### 3. Sherlock and Anagrams
# Two strings are anagrams of each other if the letters of one string can be rearranged to form the other string. Given a string, find the number of pairs of substrings of the string that are anagrams of each other.
#
# For example , the list of all anagrammatic pairs is at positions respectively.
#
# **Function Description**
#
# Complete the function sherlockAndAnagrams in the editor below. It must return an integer that represents the number of anagrammatic pairs of substrings in .
#
# **sherlockAndAnagrams has the following parameter(s):**
#
# s: a string .
def sherlockAndAnagrams(s):
n = len(s)
mp = dict()
# loop for length of substring
for i in range(n):
sb = ''
for j in range(i, n):
print(sb, "and", s[j])
sb = ''.join(sorted(sb + s[j]))
print(sb)
mp[sb] = mp.get(sb, 0)
# increase count corresponding
# to this dict array
mp[sb] += 1
print(mp)
anas = 0
# loop over all different dictionary
# items and aggregate substring count
for k, v in mp.items():
anas += (v*(v-1))//2
return anas
# #### 4. Count triplets
# You are given an array and you need to find number of tripets of indices such that the elements at those indices are in geometric progression for a given common ratio and .
#
# For example, . If , we have and at indices and .
#
# **Function Description**
#
# Complete the countTriplets function in the editor below. It should return the number of triplets forming a geometric progression for a given as an integer.
#
# **countTriplets has the following parameter(s):**
#
# arr: an array of integers
#
# r: an integer, the common ratio
# +
def is_geometric(arr, ratio):
for i in range(1, len(arr)):
if arr[i]/float(arr[i-1]) != ratio:
return False
return True
# Complete the countTriplets function below.
def countTriplets(arr, r):
sub_arr = [comb for comb in combinations(arr, 3)]
c = 0
for sub in sub_arr:
if is_geometric(sub,r) == True:
c += 1
else:
pass
return c
# -
# #### 5. Frequency queries
# You are given queries. Each query is of the form two integers described below:
#
# - : Insert x in your data structure.
# - : Delete one occurence of y from your data structure, if present.
# - : Check if any integer is present whose frequency is exactly . If yes, print 1 else 0.
#
# The queries are given in the form of a 2-D array of size where contains the operation, and contains the data element. For example, you are given array . The results of each operation are:
#
# Operation Array Output
#
# (1,1) [1]
#
# (2,2) [1]
#
# (3,2) 0
#
# (1,1) [1,1]
#
# (1,1) [1,1,1]
#
# (2,1) [1,1]
#
# (3,2) 1
#
# Return an array with the output: .
#
# **Function Description**
#
# Complete the freqQuery function in the editor below. It must return an array of integers where each element is a if there is at least one element value with the queried number of occurrences in the current array, or 0 if there is not.
#
# **freqQuery has the following parameter(s):**
#
# queries: a 2-d array of integers
def freqQuery(queries):
array = []
for order in queries:
print(order)
if order[0] == 1:
array.append(order[1])
counter = Counter(array)
elif order[0] == 2:
try:
array.remove(order[1])
counter = Counter(array)
except:
pass
elif order[0] == 3:
for el in counter:
if counter[el] == order[1]:
pres = True
break
else:
pres = None
if pres:
print("1")
else:
print("0")
print(order,array,counter)
# ## String Manipulation
# #### 1. Making Anagrams
# Alice is taking a cryptography class and finding anagrams to be very useful. We consider two strings to be anagrams of each other if the first string's letters can be rearranged to form the second string. In other words, both strings must contain the same exact letters in the same exact frequency For example, bacdc and dcbac are anagrams, but bacdc and dcbad are not.
#
# Alice decides on an encryption scheme involving two large strings where encryption is dependent on the minimum number of character deletions required to make the two strings anagrams. Can you help her find this number?
#
# Given two strings, and , that may or may not be of the same length, determine the minimum number of character deletions required to make and anagrams. Any characters can be deleted from either of the strings.
#
# For example, if and , we can delete from string and from string so that both remaining strings are and which are anagrams.
#
# **Function Description**
#
# Complete the makeAnagram function in the editor below. It must return an integer representing the minimum total characters that must be deleted to make the strings anagrams.
#
# **makeAnagram has the following parameter(s):**
#
# a: a string
#
# b: a string
def makeAnagram(a, b):
ct_a = Counter(a)
ct_b = Counter(b)
ct_a.subtract(ct_b)
return sum(abs(i) for i in ct_a.values())
# #### 2. Alternating Characters
# You are given a string containing characters and only. Your task is to change it into a string such that there are no matching adjacent characters. To do this, you are allowed to delete zero or more characters in the string.
#
# Your task is to find the minimum number of required deletions.
#
# For example, given the string , remove an at positions and to make in deletions.
#
# **Function Description**
#
# Complete the alternatingCharacters function in the editor below. It must return an integer representing the minimum number of deletions to make the alternating string.
#
# **alternatingCharacters has the following parameter(s):**
#
# s: a string
def alternatingCharacters(s):
c = 0
for i in range(1,len(s)):
if s[i] == s[i-1]:
c += 1
return c
# #### 3. Sherlock and the Valid String
# Sherlock considers a string to be valid if all characters of the string appear the same number of times. It is also valid if he can remove just character at index in the string, and the remaining characters will occur the same number of times. Given a string , determine if it is valid. If so, return YES, otherwise return NO.
#
# For example, if , it is a valid string because frequencies are . So is because we can remove one and have of each character in the remaining string. If however, the string is not valid as we can only remove occurrence of . That would leave character frequencies of .
#
# **Function Description**
#
# Complete the isValid function in the editor below. It should return either the string YES or the string NO.
#
# **isValid has the following parameter(s):**
#
# s: a string
def isValid(s):
#Create a list containing just counts of each distinct element
freq = [s.count(letter) for letter in set(s) ]
#If all values the same, then return 'YES'
if max(freq)-min(freq) == 0:
return 'YES'
#If difference between highest count and lowest count is 1
#and there is only one letter with highest count,
#then return 'YES' (because we can subtract one of these
#letters and max=min , i.e. all counts are the same)
elif freq.count(max(freq)) == 1 and max(freq) - min(freq) == 1:
return 'YES'
#If the minimum count is 1
#then remove this letter, and check whether all the other
#counts are the same
elif freq.count(min(freq)) == 1:
freq.remove(min(freq))
if max(freq)-min(freq) == 0:
return 'YES'
else:
return 'NO'
else:
return 'NO'
# #### 4. Special String Again
# A string is said to be a special string if either of two conditions is met:
#
# All of the characters are the same, e.g. aaa.
# All characters except the middle one are the same, e.g. aadaa.
# A special substring is any substring of a string which meets one of those criteria. Given a string, determine how many special substrings can be formed from it.
#
# For example, given the string , we have the following special substrings: .
#
# **Function Description**
#
# Complete the substrCount function in the editor below. It should return an integer representing the number of special substrings that can be formed from the given string.
#
# **substrCount has the following parameter(s):**
#
# n: an integer, the length of string s
#
# s: a string
def substrCount(n, s):
count = n
# tracks the last three sequences of unique characters
# seqx = length of the sequence
# seqx_c = character in that sequence
seq3, seq3_c = 0, ""
seq2, seq2_c = 0, ""
seq1, seq1_c = 1, s[0]
# note: because the slice starts at 1, i is one less than the index of char
for i, char in enumerate(s[1:]):
if char == s[i]:
count += seq1
seq1 += 1
else:
seq3, seq3_c = seq2, seq2_c
print(seq3, seq3_c)
seq2, seq2_c = seq1, seq1_c
print(seq2, seq2_c)
seq1, seq1_c = 1, char
print(seq1, seq1_c)
if seq2 == 1 and seq3 >= seq1 and seq3_c == seq1_c:
count += 1
return count
# #### 5. Common child
# A string is said to be a child of a another string if it can be formed by deleting 0 or more characters from the other string. Given two strings of equal length, what's the longest string that can be constructed such that it is a child of both?
#
# For example, ABCD and ABDC have two children with maximum length 3, ABC and ABD. They can be formed by eliminating either the D or C from both strings. Note that we will not consider ABCD as a common child because we can't rearrange characters and ABCD ABDC.
#
# **Function Description**
#
# Complete the commonChild function in the editor below. It should return the longest string which is a common child of the input strings.
#
# **commonChild has the following parameter(s):**
#
# s1, s2: two equal length strings
s1 = 'abcd'
s2 = 'abdc'
#s1 = set(s1)
#s2 = set(s2)
#print(len(s1.intersection(s2)))
def commonChild(s1, s2):
sub = 0
for i in range(len(s1)):
for j in range(len(s2)):
if s1[i] == s2[j] and i == j:
sub += 1
return sub
commonChild(s1,s2)
# ## Sorting
# 1. Bubble sort
# Consider the following version of Bubble Sort:
#
# for (int i = 0; i < n; i++) {
# for (int j = 0; j < n - 1; j++) {
# // Swap adjacent elements if they are in decreasing order
# if (a[j] > a[j + 1]) {
# swap(a[j], a[j + 1]);
# }
# }
# }
#
# Given an array of integers, sort the array in ascending order using the Bubble Sort algorithm above. Once sorted, print the following three lines:
#
# Array is sorted in numSwaps swaps., where is the number of swaps that took place.
#
# First Element: firstElement, where is the first element in the sorted array.
#
# Last Element: lastElement, where is the last element in the sorted array.
#
# Hint: To complete this challenge, you must add a variable that keeps a running tally of all swaps that occur during execution.
#
# For example, given a worst-case but small array to sort: we go through the following steps:
#
# swap a
#
# 0 [6,4,1]
#
# 1 [4,6,1]
#
# 2 [4,1,6]
#
# 3 [1,4,6]
#
# It took swaps to sort the array. Output would be
#
# Array is sorted in 3 swaps.
#
# First Element: 1
#
# Last Element: 6
#
# Function Description
#
# Complete the function countSwaps in the editor below. It should print the three lines required, then return.
#
# **countSwaps has the following parameter(s):**
#
# a: an array of integers .
def countSwaps(a):
issorted = False
swaps = 0
while not issorted:
issorted = True
for i in range(0, len(a) - 1):
if a[i] > a[i + 1]:
a[i], a[i + 1] = a[i + 1], a[i]
swaps += 1
issorted = False
print("Array is sorted in %d swaps." % swaps)
print("First Element: %d" % a[0])
print("Last Element: %d" % a[-1])
# #### 2. Mark and Toys
# Mark and Jane are very happy after having their first child. Their son loves toys, so Mark wants to buy some. There are a number of different toys lying in front of him, tagged with their prices. Mark has only a certain amount to spend, and he wants to maximize the number of toys he buys with this money.
#
# Given a list of prices and an amount to spend, what is the maximum number of toys Mark can buy? For example, if and Mark has to spend, he can buy items for , or for units of currency. He would choose the first group of items.
#
# **Function Description**
#
# Complete the function maximumToys in the editor below. It should return an integer representing the maximum number of toys Mark can purchase.
#
# **maximumToys has the following parameter(s):**
#
# prices: an array of integers representing toy prices
#
# k: an integer, Mark's budget
def maximumToys(prices, k):
prices.sort()
count = 0
for i in prices:
if (i <= k):
count += 1
k -= i
else:
break
return count
# #### 3. Comparator
# Comparators are used to compare two objects. In this challenge, you'll create a comparator and use it to sort an array. The Player class is provided in the editor below. It has two fields:
#
# Given an array of Player objects, write a comparator that sorts them in order of decreasing score. If or more players have the same score, sort those players alphabetically ascending by name. To do this, you must create a Checker class that implements the Comparator interface, then write an int compare(Player a, Player b) method implementing the Comparator.compare(T o1, T o2) method. In short, when sorting in ascending order, a comparator function returns if , if , and if .
#
# For example, given Player objects with values of , we want to sort the list as .
#
# **Function Description**
#
# Declare a Checker class that implements the comparator method as described. It should sort first descending by score, then ascending by name. The code stub reads the input, creates a list of Player objects, uses your method to sort the data, and prints it out properly.
class Player:
def __init__(self, name, score):
self.name = name
self.score = score
def __repr__(self):
pass
def comparator(a, b):
val = b.score - a.score
if val == 0:
return -1 if a.name < b.name else 1
return val
# #### 4. Fraudulent Activity Notifications
# HackerLand National Bank has a simple policy for warning clients about possible fraudulent account activity. If the amount spent by a client on a particular day is greater than or equal to the client's median spending for a trailing number of days, they send the client a notification about potential fraud. The bank doesn't send the client any notifications until they have at least that trailing number of prior days' transaction data.
#
# Given the number of trailing days and a client's total daily expenditures for a period of days, find and print the number of times the client will receive a notification over all days.
#
# For example, and . On the first three days, they just collect spending data. At day , we have trailing expenditures of . The median is and the day's expenditure is . Because , there will be a notice. The next day, our trailing expenditures are and the expenditures are . This is less than so no notice will be sent. Over the period, there was one notice sent.
#
# Note: The median of a list of numbers can be found by arranging all the numbers from smallest to greatest. If there is an odd number of numbers, the middle one is picked. If there is an even number of numbers, median is then defined to be the average of the two middle values. (Wikipedia)
#
# **Function Description**
#
# Complete the function activityNotifications in the editor below. It must return an integer representing the number of client notifications.
#
# **activityNotifications has the following parameter(s):**
#
# expenditure: an array of integers representing daily expenditures
#
# d: an integer, the lookback days for median spending
# +
from statistics import median
import bisect
def pop_then_insort(arr, x, y):
# Use bisect_left because item already exists in list, otherwise _right returns index+1
idx = bisect.bisect_left(arr, x)
# Remove existing item, pop should be faster than remove here
arr.pop(idx)
# Insort = insort_right, place item into sorted position ---> much faster than sorting array yourself
bisect.insort_right(arr, y)
return arr
def manual_median(a):
# Using built-in medians would sort the array themselves, that's too slow for us
num_items = len(a)
if num_items % 2 == 0:
m = (a[num_items//2] + a[(num_items//2)-1])/2
else:
# You don't need to do -1 but I left it as a lesson
m = a[(num_items-1)//2]
return m, a
def activityNotifications(expenditure, d):
notifs = 0
arr = []
for i in range(d, len(expenditure)):
if not arr:
# Array from trailing to one before current, remember slice ends before i here
arr = expenditure[i-d:i]
# Must be initially sorted once
arr.sort()
# Track trailing and head values as these are the only ones changing per iteration technically
old_val = expenditure[i - d]
new_val = expenditure[i]
# We don't need to do the process if old_val == new_val but there was a bug with implementing an "if !="
median_val, arr = manual_median(arr)
arr = pop_then_insort(arr, old_val, new_val)
if new_val >= 2 * median_val:
notifs += 1
return notifs
# -
# #### 5. Counting Inversion
# In an array, , the elements at indices and (where ) form an inversion if . In other words, inverted elements and are considered to be "out of order". To correct an inversion, we can swap adjacent elements.
#
# For example, consider the dataset . It has two inversions: and . To sort the array, we must perform the following two swaps to correct the inversions:
#
# Given datasets, print the number of inversions that must be swapped to sort each dataset on a new line.
#
# **Function Description**
#
# Complete the function countInversions in the editor below. It must return an integer representing the number of inversions required to sort the array.
#
# **countInversions has the following parameter(s):**
#
# arr: an array of integers to sort .
# +
def merge(arr, left_half, right_half):
i, j, k = 0, 0, 0
inversions = 0
left_len, right_len = len(left_half), len(right_half)
while i < left_len and j < right_len:
if left_half[i] <= right_half[j]:
arr[k] = left_half[i]
i += 1
else:
arr[k] = right_half[j]
j += 1
inversions += left_len - i
k += 1
while i < left_len:
arr[k] = left_half[i]
i, k = i+1, k+1
while j < right_len:
arr[k] = right_half[j]
j, k = j+1, k+1
return inversions
def merge_sort(arr):
if len(arr) > 1:
mid = len(arr)//2
left_half, right_half = arr[:mid], arr[mid:]
inversions = merge_sort(left_half) + merge_sort(right_half) + merge(arr, left_half, right_half)
return inversions
return 0
def countInversions(arr):
return merge_sort(arr)
# -
# ## Greedy Algorithms
# #### 1. Minimum absolute difference
# Consider an array of integers, . We define the absolute difference between two elements, and (where ), to be the absolute value of .
#
# Given an array of integers, find and print the minimum absolute difference between any two elements in the array. For example, given the array we can create pairs of numbers: and . The absolute differences for these pairs are , and . The minimum absolute difference is .
#
# **Function Description**
#
# Complete the minimumAbsoluteDifference function in the editor below. It should return an integer that represents the minimum absolute difference between any pair of elements.
#
# **minimumAbsoluteDifference has the following parameter(s):**
#
# n: an integer that represents the length of arr
#
# arr: an array of integers
def minimumAbsoluteDifference(arr):
arr = sorted(arr)
temp = 10000000000000000000000
for i in range(1,len(arr)):
local_diff = abs(arr[i-1] - arr[i])
if local_diff < temp:
temp = local_diff
return temp
# #### 2. Luck Balance
# Lena is preparing for an important coding competition that is preceded by a number of sequential preliminary contests. Initially, her luck balance is 0. She believes in "saving luck", and wants to check her theory. Each contest is described by two integers, and :
#
# is the amount of luck associated with a contest. If Lena wins the contest, her luck balance will decrease by ; if she loses it, her luck balance will increase by .
# denotes the contest's importance rating. It's equal to if the contest is important, and it's equal to if it's unimportant.
# If Lena loses no more than important contests, what is the maximum amount of luck she can have after competing in all the preliminary contests? This value may be negative.
#
# For example, and:
#
# Contest L[i] T[i]
#
# 1 5 1
#
# 2 1 1
#
# 3 4 0
#
# If Lena loses all of the contests, her will be . Since she is allowed to lose important contests, and there are only important contests. She can lose all three contests to maximize her luck at . If , she has to win at least of the important contests. She would choose to win the lowest value important contest worth . Her final luck will be .
#
# **Function Description**
#
# Complete the luckBalance function in the editor below. It should return an integer that represents the maximum luck balance achievable.
#
# **luckBalance has the following parameter(s):**
#
# k: the number of important contests Lena can lose
#
# contests: a 2D array of integers where each contains two integers that represent the luck balance and importance of the contest.
def luckBalance(k, contests):
luck = 0
arr_new = sorted(contests, key = lambda x: (-x[1],-x[0]))
for i in range(0,k):
luck += arr_new[i][0]
if len(arr_new) > k:
for j in range(k,len(arr_new)):
if arr_new[j][1] != 0:
luck -= arr_new[j][0]
else:
luck += arr_new[j][0]
return luck
# #### 3. Greedy Florist
# A group of friends want to buy a bouquet of flowers. The florist wants to maximize his number of new customers and the money he makes. To do this, he decides he'll multiply the price of each flower by the number of that customer's previously purchased flowers plus . The first flower will be original price, , the next will be and so on.
#
# Given the size of the group of friends, the number of flowers they want to purchase and the original prices of the flowers, determine the minimum cost to purchase all of the flowers.
#
# For example, if there are friends that want to buy flowers that cost each will buy one of the flowers priced at the original price. Having each purchased flower, the first flower in the list, , will now cost . The total cost will be .
#
# **Function Description**
#
# Complete the getMinimumCost function in the editor below. It should return the minimum cost to purchase all of the flowers.
#
# **getMinimumCost has the following parameter(s):**
#
# c: an array of integers representing the original price of each flower
#
# k: an integer, the number of friends
def getMinimumCost(k, c):
n = len(c)
c = sorted(c)
l = 1
index = k
if k == len(c):
return sum(c)
else:
temp = 0
temp += sum(c[(n-k):])
print(n-k,n,temp,"\n")
for i in range(n-k-1,-1,-1):
print(i, c[i], c[i] * (1 + l))
temp += c[i] * (1 + l)
index -= 1
if index == 0:
l += 1
index = k
return temp
# #### 4. Max Min
# You will be given a list of integers, , and a single integer . You must create an array of length from elements of such that its unfairness is minimized. Call that array . Unfairness of an array is calculated as
#
# Where:
# - max denotes the largest integer in
# - min denotes the smallest integer in
#
# As an example, consider the array with a of . Pick any two elements, test .
#
# Testing for all pairs, the solution provides the minimum unfairness.
#
# Note: Integers in may not be unique.
#
# **Function Description**
#
# Complete the maxMin function in the editor below. It must return an integer that denotes the minimum possible value of unfairness.
#
# **maxMin has the following parameter(s):**
#
# k: an integer, the number of elements in the array to create
# arr: an array of integers .
def maxMin(k, arr):
arr_n = sorted(arr)
temp = 10000000000000
n = len(arr_n)
for i in range(k,n):
slide = arr_n[(i-k):i]
balance = max(slide) - min(slide)
#print(slide, balance, temp)
if balance < temp:
temp = balance
return temp
# faster version :
def maxMin(k, a):
a.sort()
return(min(a[i + k - 1] - a[i] for i in range(len(a) - k + 1)))
# #### 5. Reverse Shuffle Merge
# Given a string, , we define some operations on the string as follows:
#
# a. denotes the string obtained by reversing string . Example:
#
#
# b. denotes any string that's a permutation of string . Example:
#
#
# c. denotes any string that's obtained by interspersing the two strings & , maintaining the order of characters in both. For example, & , one possible result of could be , another could be , another could be and so on.
#
# Given a string such that for some string , find the lexicographically smallest .
#
# For example, . We can split it into two strings of . The reverse is and we need to find a string to shuffle in to get . The middle two characters match our reverse string, leaving the and at the ends. Our shuffle string needs to be . Lexicographically , so our answer is .
#
# **Function Description**
#
# Complete the reverseShuffleMerge function in the editor below. It must return the lexicographically smallest string fitting the criteria.
#
# **reverseShuffleMerge has the following parameter(s):**
#
# s: a string
# +
# To be completed
| InterviewPrepKit.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/longqua69/practice-deep-learning/blob/44_tf_data_pipeline/44_tf_data_pipeline_exercise/44_tf_data_pipeline_exercise.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="vniRD-3tE70G"
import tensorflow as tf
# + [markdown] id="lKqVs68dFYI9"
# # Create dataset
# + [markdown] id="RucXvVuPQPv0"
# ## Clone the entire repository and navigate to the exercise directory
# + id="w2FWlQHwNja8" colab={"base_uri": "https://localhost:8080/"} outputId="c50d91ca-447f-47c1-cdc9-586e02e2a130"
# Clone the entire repo
# !git clone -l -s https://github.com/longqua69/practice-deep-learning.git working_dir
# %cd working_dir
# !ls -la
# + id="dz1J4fy9O8qy" colab={"base_uri": "https://localhost:8080/"} outputId="7e7602d2-a10b-4243-8366-d867f35c0371"
# Navigate to the exercise working directory
# %cd 44_tf_data_pipeline_exercise
# !ls -la
# + [markdown] id="ata9HaYrQlP0"
# ## Load files for creating dataset
# + id="xUM5yvQNI5H8" colab={"base_uri": "https://localhost:8080/"} outputId="fc77e3ff-c16d-426e-9284-780cdd32fc56"
dataset = tf.data.Dataset.list_files('reviews/*/*', shuffle=False)
dataset
# + id="YptbzqueP3Ti" colab={"base_uri": "https://localhost:8080/"} outputId="affdd737-8cdb-4537-b01b-a4643f47f68e"
# Take 2 samples
for file in dataset.take(2):
print(file.numpy())
# + id="UMjDdB5uSBZ3" outputId="5b8eb6d5-eab0-4c0d-9e7b-c0a9a9330858" colab={"base_uri": "https://localhost:8080/"}
len(dataset)
# + [markdown] id="4X2-Mf-GRGGa"
# # Create data input pipeline
# + [markdown] id="s1ZRaa7_YNUZ"
# ## 1. Read text review and generate a label from folder name. Dataset has review text and label as tuple
# + id="xn8FJlRoROEu" outputId="5601cec4-edfa-4619-92ef-de2844c8eb12" colab={"base_uri": "https://localhost:8080/"}
# Take all files
for file in dataset:
print(file.numpy())
# + id="WMgSWkfyaiHR" outputId="c43813ad-81e0-4d55-bc7e-633880f2a0ad" colab={"base_uri": "https://localhost:8080/"}
type(dataset)
# + id="wBrIgh6lX78C"
# Method to get labels from folder name
def get_label(file_path):
import os
parts = tf.strings.split(file_path, os.path.sep)
return parts[-2]
# + id="NnCGyw9fYys6" outputId="8656907e-d7a2-47c9-ea8a-b5b8d74ec530" colab={"base_uri": "https://localhost:8080/"}
# Create an empty set for storing labels of dataset
labels = set()
# Extract labels from folder names
for data in dataset:
label = get_label(data).numpy()
if label not in labels:
labels.add(label)
print(labels)
# + id="6OA_76ReeugY"
def text_review(filepath):
label = get_label(filepath)
txt_review = tf.io.read_file(filepath)
return txt_review , label
# + id="XzTI9Jdwg6LE" outputId="3b57c487-13c3-470f-e989-d4fc2fdc0916" colab={"base_uri": "https://localhost:8080/"}
# Read text review and generate a label from folder name.
# Dataset has review text and label as tuple
for data in dataset.take(1):
txt, label = text_review(data)
print(type(txt))
# print(label)
# + id="U7cWyOJrpift"
review_dataset = dataset.map(text_review)
# + [markdown] id="jHFB3epEjPcw"
# ## 2. Filter blank text review
# + id="3G_q8Krcps8Z"
filtered_dataset = review_dataset.filter(c)
# + id="Difu_fn0qCA5" outputId="ec9247bf-5923-4d22-9460-3b6c7a3f717a" colab={"base_uri": "https://localhost:8080/"}
# Verify the result
for file_name, label in filtered_dataset:
print("Text: ", file_name.numpy()[:50])
print("Label: ", label)
# + [markdown] id="WPqIn9BAsWnL"
# ## 3. Do all of the above transformations in single line of code.
# ## Also shuffle all the reviews
# + id="Tciw3Qz1slTJ"
# Tensorflow data pipeline
pipeline_dataset = tf.data.Dataset.list_files('reviews/*/*', shuffle=True).map(text_review).filter(lambda txt, label: txt!="")
# + id="VttRA5mntNhy" outputId="a7601607-43b4-44be-8e33-9e7b686bc858" colab={"base_uri": "https://localhost:8080/"}
# Verify the result
for text, label in pipeline_dataset:
print("Text: ", text.numpy()[:70])
print("Label", label)
print("*" * 70)
| 44_tf_data_pipeline_exercise/44_tf_data_pipeline_exercise.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Creating and Writing to file
# Creating a file and writing Hello World! to it
# +
with open('hello.txt', 'w') as f:
print("Hello World!", file=f)
# open file with open()method
# 1st argument is path
# 2nd argument is write but does not append. Means it over writes and removes the content already written
# as f is to ensure that file gets closed after writing
print(f.closed)
# When we had opened the file we could just print to it. The print is just like any other print,
# but we also need to specify that we want to print to the file we opened using file=f.
# -
# ## Reading from a file
#
# Reading a non existing file generates error
# +
lines = []
with open('hello.txt', 'r') as f:
for line in f:
lines.append(line)
print(lines)
# -
# Writing and reading multiple lines
# +
with open('hello.txt', 'w') as f:
print("Hello World!", file=f)
print("Hello two!", file=f)
print("Hello three!", file=f)
lines = []
with open('hello.txt', 'r') as f:
for line in f:
lines.append(line)
print(lines)
# +
# Strip \n
stripped = []
with open('hello.txt', 'r') as f:
for line in f:
stripped.append(line.rstrip('\n')) # read line by line
print(stripped)
# -
# For reading whole content at once!
# +
with open('hello.txt', 'r') as f:
full_content = f.read()
print(full_content)
print(type(full_content))
# +
path = '/home/hawk_pc/Python tutorial'
print(r'C:\some\name')
# -
# This program prints the contents of files:
while True:
filename = input("Filename or path, or nothing at all to exit: ")
if filename == '':
break
with open(filename, 'r') as f:
# We could read the whole file at once, but this is
# faster if the file is very large.
for line in f:
print(line.rstrip('\n'))
| Files in Python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
def create_intervals(data):
"""
Create a list of intervals out of set of ints.
"""
data=sorted(data)
if len(data) == 0:
return []
else:
answer =[-1]
for i in range(len(data)-1):
if data[i+1] - data[i] !=1:
answer.append(i)
answer.append(len(data)-1)
return [(data[answer[i]+1], data[answer[i+1]]) for i in range(len(answer)-1)]
data={1, 2, 3, 4, 5, 7, 8, 12}
data=sorted(data)
answer =[-1]
for i in range(len(data)-1):
if data[i+1] - data[i] !=1:
answer.append(i)
answer.append(len(data)-1)
[(data[answer[0]],data[answer[1]]),(data[answer[1]+1],data[answer[2]]),(data[answer[2]+1],data[answer[3]])]
# +
# others' answer
from itertools import groupby
def create_intervals(data):
return [(r[0], r[-1]) for r in (list(zip(*g))[1] for _, g in groupby(enumerate(sorted(data)), lambda i: i[1] - i[0]))]
| Creater Intervals.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Open initial condition file. Download from olympus if necessary.
# %matplotlib inline
import matplotlib.pyplot as plt
import holoviews as hv
hv.extension('bokeh', 'matplotlib')
url = "http://atmos.washington.edu/~nbren12/data/ic.nc"
![[ -e ic.nc ]] || wget {url}
ic = xr.open_dataset("ic.nc")
# +
from sam.case import InitialConditionCase
import tempfile, os, shutil
from os.path import join
def process_ngaqua(ic):
"""Diagnose the vertical velocity using SAM's pressure solver"""
path = tempfile.mkdtemp(dir=os.getcwd(), prefix='.tmp')
case = InitialConditionCase(ic=ic, path=path)
case.prm['parameters']['nstop'] = 0
case.prm['parameters']['dodamping'] = True
case.prm['parameters']['dosgs'] = True
case.save()
# !docker run -v {case.path}:/run -w /run nbren12/uwnet ./run.sh
# !docker run -v {case.path}:/run -w /run/OUT_3D nbren12/uwnet bin3D2nc CASE__1_0000000000.bin3D > /dev/null
processed = xr.open_dataset(join(case.path, 'OUT_3D', 'CASE__1_0000000000.nc'))
shutil.rmtree(case.path)
return processed
# -
# # Blurring the initial condition
#
# V is staggered to the bottom and U to the right.
v = ic.V.values
# +
from scipy.ndimage import gaussian_filter1d
def pad_south_v(v, n):
v[:,0,:] = 0
return np.pad(v, [(0,0), (n,0), (0,0)], mode='reflect', reflect_type='odd')
def pad_north_v(v, n):
v = np.pad(v, [(0,0), (0,1), (0,0)], mode='constant')
return np.pad(v, [(0,0), (0,n-1), (0,0)], mode='reflect', reflect_type='odd')
def pad_v(v,n):
return pad_north_v(pad_south_v(v, n), n)
def blur_staggered_y(v, sigma, n=None):
if n is None:
n = max(int(3*sigma), 3)
v_pad = pad_v(v, n)
return gaussian_filter1d(
gaussian_filter1d(v_pad, sigma, axis=-2)[:,n:-n,:],
sigma,
mode='wrap', axis=-1)
def blur_centered_y(f, sigma):
return gaussian_filter1d(
gaussian_filter1d(f, sigma, axis=-2, mode='nearest'),
sigma,
mode='wrap', axis=-1)
class BlurXarray:
def blur_staggered_y(V, sigma):
return xr.apply_ufunc(blur_staggered_y, V,
input_core_dims=[['z', 'y' ,'x']],
output_core_dims=[['z', 'y' ,'x']],
kwargs=dict(sigma=sigma))
def blur_centered_y(U, sigma):
return xr.apply_ufunc(blur_centered_y, U,
input_core_dims=[['z', 'y' ,'x']],
output_core_dims=[['z', 'y' ,'x']],
kwargs=dict(sigma=sigma))
@classmethod
def blur(cls, ds, sigma):
data = {}
for key in ds.data_vars:
if {'x','y'} < set(ds[key].dims):
if key == 'V':
data[key] = cls.blur_staggered_y(ds[key], sigma)
else:
data[key] = cls.blur_centered_y(ds[key], sigma)
else:
data[key] = ds[key]
return xr.Dataset(data)
# -
np.testing.assert_array_equal(pad_v(v, 10)[:,10:-10,:], v)
plt.pcolormesh(blur_staggered_y(v, sigma=1.0)[5])
plt.pcolormesh(blur_centered_y(ic.U.values, sigma=1.0)[5])
u_b = BlurXarray.blur_centered_y(ic.U, 1)
v_b = BlurXarray.blur_staggered_y(ic.V, 1)
# +
ic_b = BlurXarray.blur(ic, 1.0)
fig, (a,b) = plt.subplots(2, 1)
ic_b.V[5].plot(ax=a)
ic_b.U[5].plot(ax=b)
# -
ic_b_n = process_ngaqua(ic_b)
ic_b_n.W[0,12].plot()
# %%output size=200
# %%opts Image[width=300, height=150](cmap='RdBu_r') {-framewise -axiswise}
ds = hv.Dataset(ic_b_n.W[0])
ds.to.image(["x", "y"])
# + active=""
# %%output backend='bokeh', size=300
# %%opts QuadMesh[width=200, height=100, colorbar=True](cmap='RdBu_r') {-framewise}
# ds.to(hv.QuadMesh, ["x", "z"], dynamic=True).redim.range(W=(-.1, .1))
# -
# ## Run with blurred initial condition
86400 / 100
# +
# path = tempfile.mkdtemp(dir=os.getcwd(), prefix='.tmp')
case = InitialConditionCase(ic=ic_b, path=path)
case.prm['parameters']['dt'] = 100
case.prm['parameters']['nstop'] = 864
case.prm['parameters']['dodamping'] = True
case.prm['parameters']['dosgs'] = True
case.prm['parameters']['nstatfrq'] = 1
case.prm['parameters']['nstat'] = 36
case.prm['parameters']['nsave3d'] = 78
case.prm['parameters']['nsave2d'] = 78
case.prm['parameters']['khyp'] = 1e17
case.save()
# !docker run -v {case.path}:/run -w /run nbren12/uwnet ./run.sh
# # !docker run -v {case.path}:/run -w /run/OUT_3D nbren12/uwnet bin3D2nc CASE__1_0000000000.bin3D > /dev/null
# processed = xr.open_dataset(join(case.path, 'OUT_3D', 'CASE__1_0000000000.nc'))
# shutil.rmtree(case.path)
# !docker run -v {case.path}:/run -w /run nbren12/uwnet /opt/sam/docker/convert_files.sh > /dev/null
# -
ds_3d = xr.open_mfdataset(f"{path}/OUT_3D/*.nc")
# Here is the zonal mean meridional winds. The response looks similar to the simulations with non-blurred initial conditions.
ds_3d.V.mean('x').plot(col='time', col_wrap=4)
ds_3d.V.isel(x=0).plot(col='time', col_wrap=4)
# And for a single slice
# Here is the vertical velocity. It looks like there is some kind of wave which starts at the center of the domain and propagates outward very rapidly (at around 50 m /s). I am not sure if this is the kind of response one would expect from turning of the diabatic forcing suddenly.
ds_3d.W.mean('x').plot(col='time', col_wrap=4)
# This propogation is even more clear in this hovmoller diagram zonally averaged vertical vecloity at z=7763 m.
ds_3d.W.isel(z=16).mean('x').plot(y='y')
# Here is the zonally averaged temperature.
# +
levels = np.r_[:11]*10+273
ds_3d.SLI.mean('x').plot.contourf(col='time', col_wrap=4, levels=levels)
# -
# These are snapshots of $W$ at z = 6555.
ds_3d.W[:,15].plot(col='time', col_wrap=4)
# And V
ds_3d.V[:,15].plot(col='time', col_wrap=4)
# The hyperdiffusivity is fairly high.
# ! grep -i khyp {path}/CASE/*.nml
# # Conclusions
#
# Blurring the initial conditions does not make the meridional velocity fields behave better.
#
# $ b_t - N^2 v_y = S$
#
# $ v_t + b_y = 0$
#
| notebooks/2.4-smooth-init-cond.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Dependencies and Setup
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import requests
import time
import json
# Import API key
from api_keys import api_keys
# Incorporated citipy to determine city based on latitude and longitude
from citipy import citipy
# Output File (CSV)
output_data_file = "output_data/cities.csv"
# Range of latitudes and longitudes
lat_range = (-90, 90)
lng_range = (-180, 180)
# +
# List for holding lat_lngs and cities
lat_lngs = []
cities = []
# Create a set of random lat and lng combinations
lats = np.random.uniform(low=-90.000, high=90.000, size=1500)
lngs = np.random.uniform(low=-180.000, high=180.000, size=1500)
lat_lngs = zip(lats, lngs)
# Identify nearest city for each lat, lng combination
for lat_lng in lat_lngs:
city = citipy.nearest_city(lat_lng[0], lat_lng[1]).city_name
# If the city is unique, then add it to a our cities list
if city not in cities:
cities.append(city)
# Print the city count to confirm sufficient count
len(cities)
# -
url = "http://api.openweathermap.org/data/2.5/weather?units=Imperial&APPID="
print(request.get(url))
# +
print(requests.get(url)).json())
response = requests.get(url).json()
print(json.dump(response, indent=4 sort_keys=True))
# -
| starter_code/WeatherPy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.10.2 64-bit
# language: python
# name: python3
# ---
# +
from typing import Tuple
SAVINGS = 0.15
CHECKINGS = 0.85
def add_checkings(numbers, transactions, checkings):
for x in numbers:
checkings += transactions[x]
return checkings
def add_savings(numbers, transactions, checkings, savings):
for x in numbers:
checkings += transactions[x] * CHECKINGS
savings += transactions[x] * SAVINGS
return checkings, savings
def saturdays_bank_transactions(transactions) -> Tuple[float, float]:
savings = 1096.25
checking = 1590.80
checking = add_checkings([1,2,3,6,7,8,9,10], transactions, checking)
checking, savings = add_savings([0,4,5], transactions, checking, savings)
return checking, savings
if __name__ == "__main__":
transactions = (300.00, -50.00, -5.00, -20, 15.72, 2083.93, -1034.00, -420.00, -5.23, -15.93, -72.90)
new_balance = saturdays_bank_transactions(transactions)
print("Your new checking balance is:", '${:.2f}'.format(round(new_balance[0], 2)), "\nYour new savings balance is: ", '${:.2f}'.format(round(new_balance[1], 2)))
| JupyterNotebooks/Labs/Lab 5/dry_code.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/ashishpatel26/Pytorch-Learning/blob/main/Pytorch_Basics_Operations.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="0UejjJB1QXb8"
# ## Torch
# * Tensors are matrix-like data structures which are essential components in deep learning libraries and efficient computation.
# * **Graphical Processing Units (GPUs)** are especially effective at calculating operations between tensors, and this has spurred the surge in deep learning capability in recent times.
# * In **PyTorch**, tensors can be declared simply in a number of ways:
# 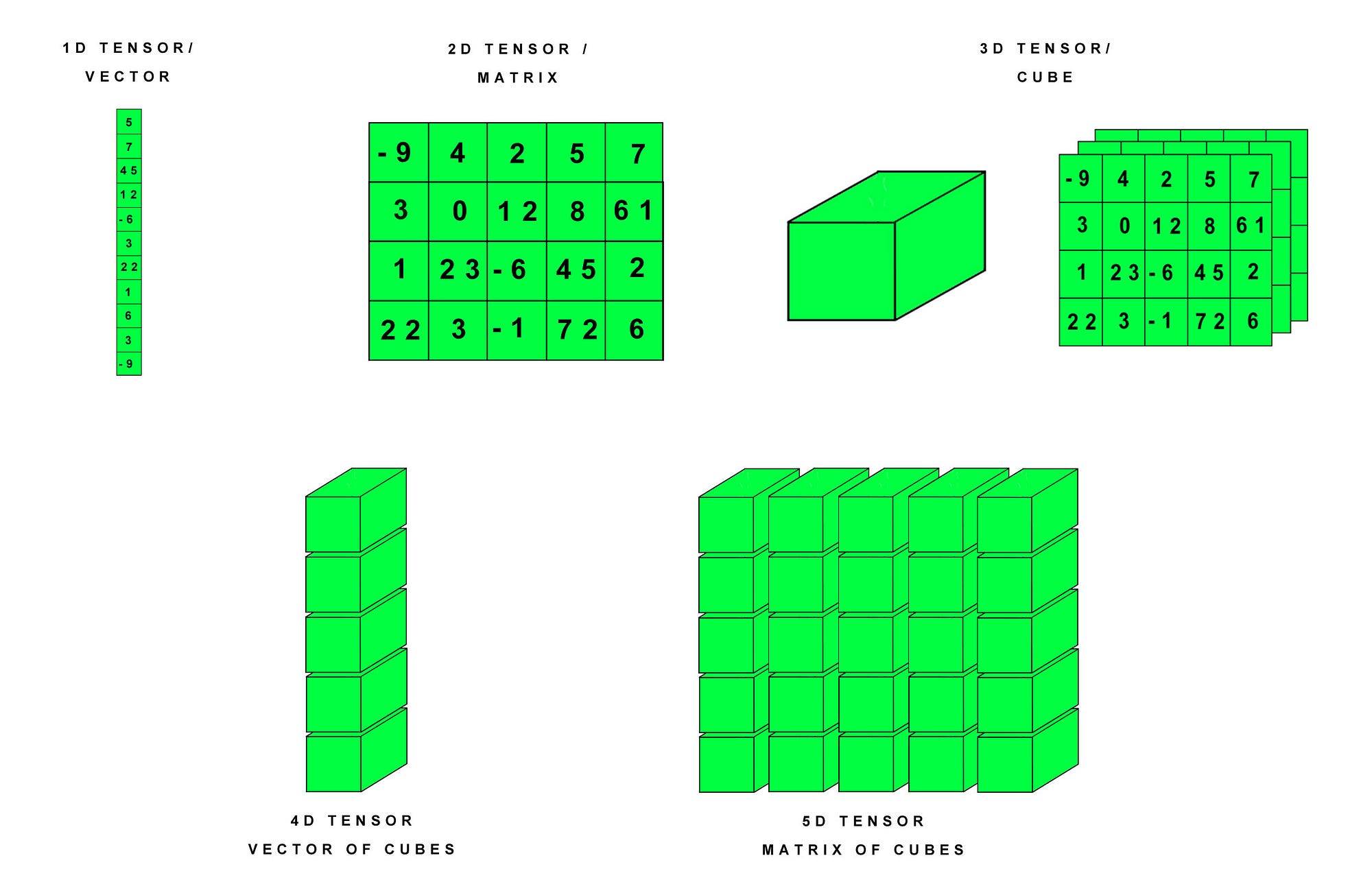
# + [markdown] id="Wdb44SCCQx1l"
# **Constuct the 4 X 4 Matrix Empty Matrix**
# + id="YvvA4u8zQN3b"
import torch
import numpy as np
# + colab={"base_uri": "https://localhost:8080/"} id="l7fyc6B2RSCI" outputId="7cbc24b2-a3bb-4804-ebd3-d1deacb072d8"
x = torch.empty(4,4)
x
# + [markdown] id="fnzJVRU7ROBI"
# **Convert to numpy**
# + colab={"base_uri": "https://localhost:8080/"} id="TCsRM2lQRDca" outputId="b9f9ddce-fc2d-470f-e2d9-bb561ef880b5"
x.numpy()
# + [markdown] id="oR5FcOwkRa1C"
# **Size of the Tensor**
# + colab={"base_uri": "https://localhost:8080/"} id="efk-DdDJRYWx" outputId="c9cdf108-a8b9-44a7-d1da-2635b032bcc6"
x.size()
# + [markdown] id="NTvy0tPXRf2z"
# From numpy to tensor
# + colab={"base_uri": "https://localhost:8080/"} id="6LD9n3ZaRdA_" outputId="38e1a1f9-67f8-472e-bdd8-e58b2a222d59"
a = np.array([[5,7],[7,5]])
b = torch.from_numpy(a)
b
# + [markdown] id="05NAafQ4Sft5"
# **Direct from Data**
# + colab={"base_uri": "https://localhost:8080/"} id="GkB09zoMSic8" outputId="9e0ab5fc-37b0-40e0-9269-f6eb7442c3c7"
data = [[1,5], [3,4]]
x_data = torch.tensor(data)
x_data
# + [markdown] id="9T2hEjM9S0Fk"
# **From another tensor:**
#
# The new tensor retains the properties (shape, datatype) of the argument tensor, unless explicitly overridden.
# + colab={"base_uri": "https://localhost:8080/"} id="J4b4FqZqS4X3" outputId="10970ec5-7f04-4afa-d30c-35024ab7afdb"
x_ones = torch.ones_like(x_data)
print(f"One Tensor : \n{x_ones}")
x_rand = torch.rand_like(x_data, dtype=torch.float)
print(f"Random Tensor : \n{x_rand}")
# + [markdown] id="2zrJ0Lb_Tg7p"
# **Random and Constant Values**
# * `shape` is a tuple of tensor dimensions.
# + colab={"base_uri": "https://localhost:8080/"} id="g-jXskKzTsOL" outputId="97560ad2-3b0a-49e4-c628-5e35a64d73ef"
shape = (2,3, )
print(f"Random Tensor : \n{torch.rand(shape)}")
print(f"Ones Tensor : \n{torch.ones(shape)}")
print(f"Zeros Tensor : \n{torch.zeros(shape)}")
# + [markdown] id="9IRu-22OUGwh"
# ### Tensor Attributes
# * Tensor attributes describe their shape, datatype, and the device on which they are stored.
# + colab={"base_uri": "https://localhost:8080/"} id="6M5mVTCrUOQ0" outputId="f26229c3-f922-44d6-e818-1c2b6483db11"
tensor = torch.rand(3,4)
print(f"Shape of the Tensor: {tensor.shape}")
print(f"Datatypes of the tensor: {tensor.dtype}")
print(f"Device tensor is stored on: {tensor.device}")
# + [markdown] id="qU2vYxseR7sL"
# ### Tensor Operation
# * Over 100 tensor operations, including transposing, indexing, slicing, mathematical operations, linear algebra, random sampling, and more are comprehensively described [here](https://pytorch.org/docs/stable/torch.html).
# + id="va24-cqQRvgu"
if torch.cuda.is_available():
tensor = tensor.to('cuda')
# + [markdown] id="Wtbyd6rcV4lS"
# **Basic Tensor Operation**
# 
#
# * Addition(`torch.add()`)
# * Substraction(`torch.sub()`)
# * Division(`torch.div()`)
# * Multiplication(`torch.mul()`)
# + colab={"base_uri": "https://localhost:8080/"} id="fXNbTx4aVcUp" outputId="ae1afcbe-e633-4db8-825a-8b9cdf1de4fd"
# Initialization
a = torch.rand(2,2)
b = torch.rand(2,2)
# Old Method
print(a + b)
# New Method
print(torch.add(a,b))
# + colab={"base_uri": "https://localhost:8080/"} id="-hxuCTXyXr1j" outputId="68c11552-f663-44de-c03e-d27bf0d26e48"
# Old Method
print(a - b)
# New Method
print(torch.sub(a,b))
# + colab={"base_uri": "https://localhost:8080/"} id="TzCwZV1vYIJa" outputId="a8da91c3-1ddc-4b58-ad36-e05c984ea5a3"
# Old Method
print(a * b)
# New Method
print(torch.mul(a,b))
# + colab={"base_uri": "https://localhost:8080/"} id="lCKV5GooYWir" outputId="0953384a-c1a8-4c9e-bc00-688f6fa3c296"
# Old Method
print(a / b)
# New Method
print(torch.div(a,b))
# + [markdown] id="cMwHWjtvYgIm"
# **Add x to y**
# + colab={"base_uri": "https://localhost:8080/"} id="nTcTry62Ydx0" outputId="1b24e07c-b532-4cc4-8bc8-7cae153248e6"
# a directly add to b and answer stored in b
print(f"b value before adding into a : \n{b}")
b.add_(a)
print(f"b value after adding into a : \n{b}")
# + [markdown] id="GWZRoA9xY5EQ"
# **Standard Numpy like Indexing**
# + colab={"base_uri": "https://localhost:8080/"} id="O-hpiL15Y1-D" outputId="14ace5a7-fd07-45b3-be1a-d14b99ab210f"
print(f"Column 0 : {a[:,0]}")
print(f"Column 1 : {a[:,1]}")
# + [markdown] id="Jtp1HXjLZ1j1"
# **Resizing**
# + colab={"base_uri": "https://localhost:8080/"} id="hO4G9voCY_F4" outputId="6cccab1d-673f-4b3a-a852-e29670ea6e89"
x = torch.rand(4,4)
print(f"x: \n{x}")
y = x.view(16) # View is used to flatten the tensor
print(f"y : \n{y}")
z = x.view(-1, 8) # the size -1 is inferred from other dimensions
print(f"z : \n{z}")
print(f"Size of x : {x.size()}\nSize of y : {y.size()}\nSize of z : {z.size()}")
# + [markdown] id="XRScor2SbdwR"
# ---
# Note : Any operation that mutates a tensor in-place is post-fixed with an . For example: `x.copy_(y)`, `x.t_()`, will change x.
# for more about tensor go on this link : https://pytorch.org/docs/stable/tensors.html
#
#
# ---
# + [markdown] id="B6kTlu2A5Zsi"
# **Joining Tensor**
# + colab={"base_uri": "https://localhost:8080/"} id="ZUbHHrWZ5oIH" outputId="06c953a5-4279-443b-a609-53633f3f4bb1"
tensor = torch.ones(4, 4)
tensor[:,1] = 0
print(tensor)
# + colab={"base_uri": "https://localhost:8080/"} id="yF8TJErz5XKK" outputId="3591dbbc-dce8-4b2c-96df-30c7313cf682"
t1 = torch.cat([tensor,tensor,tensor], dim=1)
print(t1)
# + [markdown] id="ElxJSXc05rLZ"
# **Multiplying Tensor**
# + colab={"base_uri": "https://localhost:8080/"} id="g-sDPqka5hti" outputId="f1bc56d5-5c76-45fa-a255-4fbcb48b0593"
# This is computes the element-wise product
print(f"tensor.mul(tensor) : \n {tensor.mul(tensor)}")
# Alternative syntax
print(f"tensor * tensor : \n {tensor * tensor}")
# + [markdown] id="Dt8C3A6G6ENL"
# **Matrix Multiplication**
# + colab={"base_uri": "https://localhost:8080/"} id="rYWqn5ZX5-Ea" outputId="ba9ba230-2449-4e02-c5d6-9714c5de2171"
print(f"tensor.matmul(tensor.T) \n {tensor.matmul(tensor.T)} \n")
# Alternative syntax:
print(f"tensor @ tensor.T \n {tensor @ tensor.T}")
# + [markdown] id="kt2QKHqZ6Tlf"
# **Bridge with Numpy**
# + colab={"base_uri": "https://localhost:8080/"} id="kXRpjVH36KMO" outputId="2c122e0d-6ca3-4a8c-af3e-93a855f4168b"
t = torch.ones(5)
print(f"t: {t}")
n = t.numpy()
print(f"n: {n}")
# + colab={"base_uri": "https://localhost:8080/"} id="1D3-GiBe6f0m" outputId="df3dc7eb-6a27-4a24-97a6-b20e052c07c3"
t.add_(1) #inplace with adding 1 to tensor
print(f"t: {t}")
print(f"n: {n}")
| Pytorch_Basics_Operations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# default_exp models.RNN_FCNPlus
# -
# # RNN_FCNPlus
#
# > This is an unofficial PyTorch implementation by <NAME> - <EMAIL> based on:
#
# * <NAME>., <NAME>., <NAME>., & <NAME>. (2017). LSTM fully convolutional networks for time series classification. IEEE Access, 6, 1662-1669.
# * Official LSTM-FCN TensorFlow implementation: https://github.com/titu1994/LSTM-FCN
#
# * <NAME>., <NAME>., & <NAME>. (2018). Deep Gated Recurrent and Convolutional Network Hybrid Model for Univariate Time Series Classification. arXiv preprint arXiv:1812.07683.
# * Official GRU-FCN TensorFlow implementation: https://github.com/NellyElsayed/GRU-FCN-model-for-univariate-time-series-classification
#export
from tsai.imports import *
from tsai.models.layers import *
# +
#export
class _RNN_FCN_BasePlus(Module):
def __init__(self, c_in, c_out, seq_len=None, hidden_size=100, rnn_layers=1, bias=True, cell_dropout=0, rnn_dropout=0.8, bidirectional=False, shuffle=True,
fc_dropout=0., conv_layers=[128, 256, 128], kss=[7, 5, 3], se=0):
if shuffle: assert seq_len is not None, 'need seq_len if shuffle=True'
# RNN - first arg is usually c_in. Authors modified this to seq_len by not permuting x. This is what they call shuffled data.
self.rnn = self._cell(seq_len if shuffle else c_in, hidden_size, num_layers=rnn_layers, bias=bias, batch_first=True,
dropout=cell_dropout, bidirectional=bidirectional)
self.rnn_dropout = nn.Dropout(rnn_dropout) if rnn_dropout else noop
self.shuffle = Permute(0,2,1) if not shuffle else noop # You would normally permute x. Authors did the opposite.
# FCN
assert len(conv_layers) == len(kss)
self.convblock1 = ConvBlock(c_in, conv_layers[0], kss[0])
self.se1 = SqueezeExciteBlock(conv_layers[0], se) if se != 0 else noop
self.convblock2 = ConvBlock(conv_layers[0], conv_layers[1], kss[1])
self.se2 = SqueezeExciteBlock(conv_layers[1], se) if se != 0 else noop
self.convblock3 = ConvBlock(conv_layers[1], conv_layers[2], kss[2])
self.gap = GAP1d(1)
# Common
self.concat = Concat()
self.fc_dropout = nn.Dropout(fc_dropout) if fc_dropout else noop
self.fc = nn.Linear(hidden_size * (1 + bidirectional) + conv_layers[-1], c_out)
def forward(self, x):
# RNN
rnn_input = self.shuffle(x) # permute --> (batch_size, seq_len, n_vars) when batch_first=True
output, _ = self.rnn(rnn_input)
last_out = output[:, -1] # output of last sequence step (many-to-one)
last_out = self.rnn_dropout(last_out)
# FCN
x = self.convblock1(x)
x = self.se1(x)
x = self.convblock2(x)
x = self.se2(x)
x = self.convblock3(x)
x = self.gap(x)
# Concat
x = self.concat([last_out, x])
x = self.fc_dropout(x)
x = self.fc(x)
return x
class RNN_FCNPlus(_RNN_FCN_BasePlus):
_cell = nn.RNN
class LSTM_FCNPlus(_RNN_FCN_BasePlus):
_cell = nn.LSTM
class GRU_FCNPlus(_RNN_FCN_BasePlus):
_cell = nn.GRU
class MRNN_FCNPlus(_RNN_FCN_BasePlus):
_cell = nn.RNN
def __init__(self, *args, se=16, **kwargs):
super().__init__(*args, se=16, **kwargs)
class MLSTM_FCNPlus(_RNN_FCN_BasePlus):
_cell = nn.LSTM
def __init__(self, *args, se=16, **kwargs):
super().__init__(*args, se=16, **kwargs)
class MGRU_FCNPlus(_RNN_FCN_BasePlus):
_cell = nn.GRU
def __init__(self, *args, se=16, **kwargs):
super().__init__(*args, se=16, **kwargs)
# -
bs = 16
n_vars = 3
seq_len = 12
c_out = 2
xb = torch.rand(bs, n_vars, seq_len)
test_eq(RNN_FCNPlus(n_vars, c_out, seq_len)(xb).shape, [bs, c_out])
test_eq(LSTM_FCNPlus(n_vars, c_out, seq_len)(xb).shape, [bs, c_out])
test_eq(MLSTM_FCNPlus(n_vars, c_out, seq_len)(xb).shape, [bs, c_out])
test_eq(GRU_FCNPlus(n_vars, c_out, shuffle=False)(xb).shape, [bs, c_out])
test_eq(GRU_FCNPlus(n_vars, c_out, seq_len, shuffle=False)(xb).shape, [bs, c_out])
LSTM_FCNPlus(n_vars, seq_len, c_out, se=8)
#hide
out = create_scripts()
beep(out)
| nbs/107b_models.RNN_FCNPlus.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="ZTmOqMz1LpFz"
from google.colab import drive
# + colab={"base_uri": "https://localhost:8080/"} id="5nUmyVUhYNbp" outputId="1c3e6d4b-525e-479b-d1d3-68bd4a28c69c"
drive.mount('/content/drive')
# + id="f34tBfdVYeJ5"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
sns.set()
# + id="jjCOL1waYkbo"
data = pd.read_csv('/content/drive/MyDrive/Data/Indian automoble buying behavour study 1.0.csv')
# + colab={"base_uri": "https://localhost:8080/", "height": 575} id="BrwH7x7VYzAu" outputId="0f8b42fb-86c8-4133-acca-c4fa828c7bb1"
data
# + id="O7HOC5IJY1nj"
sns.set_style('dark')
# + colab={"base_uri": "https://localhost:8080/"} id="G8yaXprhY66A" outputId="b0d91715-ef62-464f-b01d-fc1111e34df5"
data.isnull().sum()
# + colab={"base_uri": "https://localhost:8080/", "height": 356} id="4Pqk41xqY-1a" outputId="adff91db-0d79-4fad-8c04-c38e97759f19"
sns.distplot(data['Age'])
# + colab={"base_uri": "https://localhost:8080/", "height": 367} id="NO1JeoZgZNh1" outputId="4e15542e-4650-4ebe-eb97-3691547cc8b0"
sns.distplot(data['Total Salary'])
# + colab={"base_uri": "https://localhost:8080/", "height": 367} id="Uw-D4v1HaC1C" outputId="e6494712-37c9-4a5c-ece2-e13c7e793794"
sns.distplot(data['Price'])
# + colab={"base_uri": "https://localhost:8080/", "height": 301} id="cANTnjB9Zg4M" outputId="75149595-97ab-49f3-f1af-461e9eac82fe"
plt.scatter(data['Age'],data['Total Salary'])
plt.title('Age and Total Salary')
# + colab={"base_uri": "https://localhost:8080/", "height": 301} id="ysqZMWlWaRxh" outputId="36f4e4cc-3670-4d5d-e3b8-0ac2b82217f9"
plt.scatter(data['Age'],data['Price'])
plt.title('Age and Car Price')
| Customerbase.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from keras.layers import Dense, Flatten, Dropout, ZeroPadding3D
from keras.layers.recurrent import LSTM
from keras.models import Sequential, load_model
from keras.optimizers import Adam, RMSprop
from keras.layers.wrappers import TimeDistributed
from keras.layers.convolutional import (Conv2D, MaxPooling3D, Conv3D,
MaxPooling2D)
from collections import deque
import sys
class ResearchModels():
def __init__(self, nb_classes, model, seq_length,
saved_model=None, features_length=2048):
"""
`model` = one of:
lstm
lrcn
mlp
conv_3d
c3d
`nb_classes` = the number of classes to predict
`seq_length` = the length of our video sequences
`saved_model` = the path to a saved Keras model to load
"""
# Set defaults.
self.seq_length = seq_length
self.load_model = load_model
self.saved_model = saved_model
self.nb_classes = nb_classes
self.feature_queue = deque()
# Set the metrics. Only use top k if there's a need.
metrics = ['accuracy']
if self.nb_classes >= 10:
metrics.append('top_k_categorical_accuracy')
# Get the appropriate model.
if self.saved_model is not None:
print("Loading model %s" % self.saved_model)
self.model = load_model(self.saved_model)
elif model == 'lstm':
print("Loading LSTM model.")
self.input_shape = (seq_length, features_length)
self.model = self.lstm()
elif model == 'lrcn':
print("Loading CNN-LSTM model.")
self.input_shape = (seq_length, 80, 80, 3)
self.model = self.lrcn()
elif model == 'mlp':
print("Loading simple MLP.")
self.input_shape = (seq_length, features_length)
self.model = self.mlp()
elif model == 'conv_3d':
print("Loading Conv3D")
self.input_shape = (seq_length, 80, 80, 3)
self.model = self.conv_3d()
elif model == 'c3d':
print("Loading C3D")
self.input_shape = (seq_length, 80, 80, 3)
self.model = self.c3d()
else:
print("Unknown network.")
sys.exit()
# Now compile the network.
optimizer = Adam(lr=1e-5, decay=1e-6)
self.model.compile(loss='categorical_crossentropy', optimizer=optimizer,
metrics=metrics)
print(self.model.summary())
def lstm(self):
"""Build a simple LSTM network. We pass the extracted features from
our CNN to this model predomenently."""
# Model.
model = Sequential()
model.add(LSTM(2048, return_sequences=False,
input_shape=self.input_shape,
dropout=0.5))
model.add(Dense(512, activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(self.nb_classes, activation='softmax'))
return model
def lrcn(self):
"""Build a CNN into RNN.
Starting version from:
https://github.com/udacity/self-driving-car/blob/master/
steering-models/community-models/chauffeur/models.py
Heavily influenced by VGG-16:
https://arxiv.org/abs/1409.1556
Also known as an LRCN:
https://arxiv.org/pdf/1411.4389.pdf
"""
def add_default_block(model, kernel_filters, init, reg_lambda):
# conv
model.add(TimeDistributed(Conv2D(kernel_filters, (3, 3), padding='same',
kernel_initializer=init, kernel_regularizer=L2_reg(l=reg_lambda))))
model.add(TimeDistributed(BatchNormalization()))
model.add(TimeDistributed(Activation('relu')))
# conv
model.add(TimeDistributed(Conv2D(kernel_filters, (3, 3), padding='same',
kernel_initializer=init, kernel_regularizer=L2_reg(l=reg_lambda))))
model.add(TimeDistributed(BatchNormalization()))
model.add(TimeDistributed(Activation('relu')))
# max pool
model.add(TimeDistributed(MaxPooling2D((2, 2), strides=(2, 2))))
return model
initialiser = 'glorot_uniform'
reg_lambda = 0.001
model = Sequential()
# first (non-default) block
model.add(TimeDistributed(Conv2D(32, (7, 7), strides=(2, 2), padding='same',
kernel_initializer=initialiser, kernel_regularizer=L2_reg(l=reg_lambda)),
input_shape=self.input_shape))
model.add(TimeDistributed(BatchNormalization()))
model.add(TimeDistributed(Activation('relu')))
model.add(TimeDistributed(Conv2D(32, (3,3), kernel_initializer=initialiser, kernel_regularizer=L2_reg(l=reg_lambda))))
model.add(TimeDistributed(BatchNormalization()))
model.add(TimeDistributed(Activation('relu')))
model.add(TimeDistributed(MaxPooling2D((2, 2), strides=(2, 2))))
# 2nd-5th (default) blocks
model = add_default_block(model, 64, init=initialiser, reg_lambda=reg_lambda)
model = add_default_block(model, 128, init=initialiser, reg_lambda=reg_lambda)
model = add_default_block(model, 256, init=initialiser, reg_lambda=reg_lambda)
model = add_default_block(model, 512, init=initialiser, reg_lambda=reg_lambda)
# LSTM output head
model.add(TimeDistributed(Flatten()))
model.add(LSTM(256, return_sequences=False, dropout=0.5))
model.add(Dense(self.nb_classes, activation='softmax'))
return model
def mlp(self):
"""Build a simple MLP. It uses extracted features as the input
because of the otherwise too-high dimensionality."""
# Model.
model = Sequential()
model.add(Flatten(input_shape=self.input_shape))
model.add(Dense(512))
model.add(Dropout(0.5))
model.add(Dense(512))
model.add(Dropout(0.5))
model.add(Dense(self.nb_classes, activation='softmax'))
return model
def conv_3d(self):
"""
Build a 3D convolutional network, based loosely on C3D.
https://arxiv.org/pdf/1412.0767.pdf
"""
# Model.
model = Sequential()
model.add(Conv3D(
32, (3,3,3), activation='relu', input_shape=self.input_shape
))
model.add(MaxPooling3D(pool_size=(1, 2, 2), strides=(1, 2, 2)))
model.add(Conv3D(64, (3,3,3), activation='relu'))
model.add(MaxPooling3D(pool_size=(1, 2, 2), strides=(1, 2, 2)))
model.add(Conv3D(128, (3,3,3), activation='relu'))
model.add(Conv3D(128, (3,3,3), activation='relu'))
model.add(MaxPooling3D(pool_size=(1, 2, 2), strides=(1, 2, 2)))
model.add(Conv3D(256, (2,2,2), activation='relu'))
model.add(Conv3D(256, (2,2,2), activation='relu'))
model.add(MaxPooling3D(pool_size=(1, 2, 2), strides=(1, 2, 2)))
model.add(Flatten())
model.add(Dense(1024))
model.add(Dropout(0.5))
model.add(Dense(1024))
model.add(Dropout(0.5))
model.add(Dense(self.nb_classes, activation='softmax'))
return model
def c3d(self):
"""
Build a 3D convolutional network, aka C3D.
https://arxiv.org/pdf/1412.0767.pdf
With thanks:
https://gist.github.com/albertomontesg/d8b21a179c1e6cca0480ebdf292c34d2
"""
model = Sequential()
# 1st layer group
model.add(Conv3D(64, 3, 3, 3, activation='relu',
border_mode='same', name='conv1',
subsample=(1, 1, 1),
input_shape=self.input_shape))
model.add(MaxPooling3D(pool_size=(1, 2, 2), strides=(1, 2, 2),
border_mode='valid', name='pool1'))
# 2nd layer group
model.add(Conv3D(128, 3, 3, 3, activation='relu',
border_mode='same', name='conv2',
subsample=(1, 1, 1)))
model.add(MaxPooling3D(pool_size=(2, 2, 2), strides=(2, 2, 2),
border_mode='valid', name='pool2'))
# 3rd layer group
model.add(Conv3D(256, 3, 3, 3, activation='relu',
border_mode='same', name='conv3a',
subsample=(1, 1, 1)))
model.add(Conv3D(256, 3, 3, 3, activation='relu',
border_mode='same', name='conv3b',
subsample=(1, 1, 1)))
model.add(MaxPooling3D(pool_size=(2, 2, 2), strides=(2, 2, 2),
border_mode='valid', name='pool3'))
# 4th layer group
model.add(Conv3D(512, 3, 3, 3, activation='relu',
border_mode='same', name='conv4a',
subsample=(1, 1, 1)))
model.add(Conv3D(512, 3, 3, 3, activation='relu',
border_mode='same', name='conv4b',
subsample=(1, 1, 1)))
model.add(MaxPooling3D(pool_size=(2, 2, 2), strides=(2, 2, 2),
border_mode='valid', name='pool4'))
# 5th layer group
model.add(Conv3D(512, 3, 3, 3, activation='relu',
border_mode='same', name='conv5a',
subsample=(1, 1, 1)))
model.add(Conv3D(512, 3, 3, 3, activation='relu',
border_mode='same', name='conv5b',
subsample=(1, 1, 1)))
model.add(ZeroPadding3D(padding=(0, 1, 1)))
model.add(MaxPooling3D(pool_size=(2, 2, 2), strides=(2, 2, 2),
border_mode='valid', name='pool5'))
model.add(Flatten())
# FC layers group
model.add(Dense(4096, activation='relu', name='fc6'))
model.add(Dropout(0.5))
model.add(Dense(4096, activation='relu', name='fc7'))
model.add(Dropout(0.5))
model.add(Dense(self.nb_classes, activation='softmax'))
return model
# -
from keras.callbacks import TensorBoard, ModelCheckpoint, EarlyStopping, CSVLogger
from models import ResearchModels
from data import DataSet
import time
import os.path
| UCf101_run.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <center>Praca domowa 2</center>
# ### <center>Kodowanie zmiennych kategorycznych i uzupełnienie braków<br><NAME><br>21 marca 2020r.</center>
#
# ## 1. Wczytanie potrzebnych bibliotek oraz danych
# +
import pandas as pd
import numpy as np
import sklearn
# 68MB
data = pd.read_csv("~/Downloads/allegro-api-transactions.csv")
data.head()
# -
data.info()
print("Liczba unikalnych wartości pola it_location: " + str(len(data.it_location.unique())))
# #### Od razu widać jaka jest przewaga target end=coding nad one-hot - druga metoda spowodowałaby stworzenie ponad 10000 nowych kolumn
# ## 2. Target encoding zmiennej it_location
data.it_location
import category_encoders
te = category_encoders.target_encoder.TargetEncoder(data)
target_encoded = te.fit_transform(data.it_location, data.price)
data.it_location = target_encoded
data.head()
data.it_location.head()
data.info()
# #### Po wykonaniu target encodingu zmienna it_location jest typu float
# ## 3. One hot encoding kolumny main_category
print("Liczba unikalnych wartości kolumny main_category: " + str(len(data.main_category.unique())))
# +
from sklearn.preprocessing import OneHotEncoder
oe = OneHotEncoder(sparse=False)
mc = data["main_category"].to_numpy().reshape(len(data["main_category"]), 1)
new_part = pd.DataFrame(oe.fit_transform(mc), columns=(sorted("mc_" + data.main_category.unique())))
# Sprawdźmy popraność kodowania poszczególnych wartości main_category:
for vals in data.main_category.unique():
x = data.main_category == vals
y = new_part[["mc_"+vals]] == 1
print(vals + ": " + str((y[["mc_"+vals]].squeeze() == x).all()))
# -
new_part.head()
# Następnie należałoby odpowiednio połączyć ramki danych
# ## 4. CatBoost Encoder
# Przypomnijmy jak wygląda nasza ramka danych:
data.head()
cbe = category_encoders.cat_boost.CatBoostEncoder(cols = ["main_category"])
h = cbe.fit_transform(data, data.price)
h.head()
# Wydawać by się mogło, że wiele z wartości zmiennej main_category powtarza się wielokrotnie. Jest to jednak nieprawda, jak zobaczymy tylko niewielki ułamek to wartości dublujące się:
print("Wartości dublujące się stanowią " + str(100*(1 - float(len(h.main_category.unique()))/len(h))) + "%.")
# #### Krótki opis działania algorytmu:
#
# Idea działania algorytmu kodowania zmiennych kategorycznych CatBoost jest podobna do Target Encodingu, wartości są zastępowane odpowiednimi średnimi zmiennej objaśnianej, jednakże w celu ustrzeżenia się przed outlierami, nie jest uwzględniana wartość z danego wiersza. Dodatkowo, CatBoost encoder wykonuje te operacje w locie, dlatego te wartości stale się zmieniają. Jak się domyślam, w pierwszych wierszach brana jest średnia z wszystkich obserwacji:
print(data.price.mean())
#tak
# ## 5.Ordinal
ord = category_encoders.ordinal.OrdinalEncoder(cols = ["main_category"])
ord.fit(data)
ordinaled = ord.transform(data)
ordinaled
# #### Krótki opis działania algorytmu:
#
# Jest to jedna z prostszych metod. W przypadku niepodania konkretnego sposobu mapowania, kolejnym wartościom przypisywane są kolejne liczby całkowite. Dla braku danych również przypisana jest specjalna wartość.
# ## 6. Uzupełnianie braków
numdata = data[["price", "it_quantity", "it_seller_rating"]]
numdata.head()
# ## 7. Multivariate feature imputation
# #### Usunięte 10% wartości it_seller_rating
from sklearn.experimental import enable_iterative_imputer
from sklearn.impute import IterativeImputer
from sklearn.metrics import mean_squared_error
import math
itimp = IterativeImputer(max_iter=10)
import random
RMSE = []
for i in range(10):
numdata = data.copy()
numdata = numdata[["price", "it_quantity", "it_seller_rating"]]
assert ~pd.isna(numdata).any().any() # sprawdzenie czy nie ma brakujacych wartosci
indx = random.sample(range(len(numdata)), int(len(numdata.price)/10)) #losowanie 10% indeksów
assert len(np.unique(indx)) == len(indx) #czy na pewno losowanie odbywalo sie bez zwracania aby bylo 10%?
cut_values = numdata.loc[indx, "it_seller_rating"] # zapisanie wartosci ktore zostana usuniete
numdata.loc[indx, "it_seller_rating"] = np.nan #usuniecie wartosci
no_na = numdata[~pd.isna(numdata.it_seller_rating)]
only_na = numdata[pd.isna(numdata.it_seller_rating)]
itimp.fit(no_na) #imputacja - model
IterativeImputer()
i = itimp.transform(only_na)
RMSE.append(math.sqrt(mean_squared_error(cut_values, pd.DataFrame(i).iloc[:, 2])))
RMSE
# ## 8. Usunięcie 10% wartości it_seller_rating oraz it_quantity
RMSE2 = []
for i in range(10):
numdata = data[["price", "it_quantity", "it_seller_rating"]].copy()
assert ~pd.isna(numdata).any().any() # sprawdzenie czy nie ma brakujacych wartosci
indx = random.sample(range(len(numdata)), int(len(numdata.price)/10)) #losowanie 10% indeksów
indx2 = random.sample(range(len(numdata)), int(len(numdata.price)/10)) #losowanie 10% indeksów dla 2. kolumny
assert len(np.unique(indx)) == len(indx) #czy na pewno losowanie odbywalo sie bez zwracania aby bylo 10%?
assert len(np.unique(indx2)) == len(indx2)
cut_values = numdata.loc[indx, "it_seller_rating"] # zapisanie wartosci ktore zostana usuniete
numdata.loc[indx, "it_seller_rating"] = np.nan #usuniecie wartosci
numdata.loc[indx2, "it_quantity"] = np.nan #usuniecie wartosci
assert len(indx)>(0.05*len(numdata))
assert len(indx2)>(0.05*len(numdata))
assert sum(pd.isna(numdata.it_seller_rating)) == len(indx) #sprawdzenie czy usuwanie wartości się powiodło
assert sum(pd.isna(numdata.it_quantity)) == len(indx2)
no_na = numdata[~pd.isna(numdata.it_seller_rating)]
only_na = numdata[pd.isna(numdata.it_seller_rating)]
itimp.fit(no_na) #imputacja - model
IterativeImputer()
i = itimp.transform(only_na)
RMSE2.append(math.sqrt(mean_squared_error(cut_values, pd.DataFrame(i).iloc[:, 2])))
RMSE2
import seaborn as sns
rmses = pd.DataFrame({"Błąd średniokwadratowy": RMSE+RMSE2})
rmses.loc[0:10, "Próba"] = "1. metoda"
rmses.loc[10:20, "Próba"] = "2. metoda"
p1 = sns.boxplot(x = "Próba", y = "Błąd średniokwadratowy", data = rmses)
p1.set_title("Rozrzut błędów RMSE uzupełniania braków it_seller_rating")
p1
err = pd.DataFrame({"1. metoda": RMSE, "2. metoda": RMSE2})
err.describe()
from matplotlib import pyplot
err.plot()
# #### Powyższe wykresy wskazują, że trudno jednoznacznie określić czy usunięcie części wartości kolumny it_quantity negatywnie wpłynęło na szacowanie wartości it_seller_rating. Z pewnością większy był wówczas rozrzut błędów (odchylenie standardowe: 303 v 351), jednak minialny błąd wystąpił w drugim przypadku testowym, a maksymalny w pierwszym. Co ciekawe, średnia wartość błędu była zbliżona w obu przypadkach.
| Prace_domowe/Praca_domowa2/Grupa1/KozminskiPawel/PracaDomowa2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## In this notebook, we will demostrate how to run some local machine learning experiments and collect the performance measurements. These measurements will be later used to train the IRT models.
import sys; sys.path.insert(0, '..')
import numpy
import scipy.stats
import sklearn.datasets
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import RandomForestClassifier
from sklearn.naive_bayes import GaussianNB
import atml.measure
import atml.exp
# ## To set up the machine learning experiments, we need to first define the datasets and models. This toolbox requires the datasets and models to be indexed by python dictionaries.
data_dict = {0: 'iris',
1: 'digits',
2: 'wine'}
model_dict = {0: 'lr',
1: 'rf',
2: 'nb'}
# ## Furthermore, we also need to provide two functions to load the datasets and declare the models. We assume the datasets to be represented as numpy.ndarray, with x as features, y as target. The model should have the same format as sklearn.predictor, with fit() as the training function, and predict_proba() as the function to predict probability vectors.
# +
def get_data(ref):
if ref == 'iris':
x, y = sklearn.datasets.load_iris(return_X_y=True)
elif ref == 'digits':
x, y = sklearn.datasets.load_digits(return_X_y=True)
elif ref == 'wine':
x, y = sklearn.datasets.load_wine(return_X_y=True)
return x, y
def get_model(ref):
if ref == 'lr':
mdl = LogisticRegression()
elif ref == 'rf':
mdl = RandomForestClassifier()
elif ref == 'nb':
mdl = GaussianNB()
return mdl
# -
# ## For this example, we use the built-in measure of Brier score.
measure = atml.measure.BS()
# ## Now we can use the built-in function to perform an exhaustive testing, that is, to test all combinations of different datasets and models, and collect the corresponding performance measurements.
res = atml.exp.get_exhaustive_testing(data_dict, get_data, model_dict, get_model, measure)
# ## We can check the results with Pandas dataframe.
res
# ## Save the results (to be used later for IRT training)
res.to_csv('./res_base.csv')
| notebooks/base_testing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Programming Exercise 1: Linear Regression
#
# ## Introduction
#
# In this exercise, you will implement linear regression and get to see it work on data. Before starting on this programming exercise, we strongly recommend watching the video lectures and completing the review questions for the associated topics.
#
# All the information you need for solving this assignment is in this notebook, and all the code you will be implementing will take place within this notebook. The assignment can be promptly submitted to the coursera grader directly from this notebook (code and instructions are included below).
#
# Before we begin with the exercises, we need to import all libraries required for this programming exercise. Throughout the course, we will be using [`numpy`](http://www.numpy.org/) for all arrays and matrix operations, and [`matplotlib`](https://matplotlib.org/) for plotting.
#
# You can find instructions on how to install required libraries in the README file in the [github repository](https://github.com/dibgerge/ml-coursera-python-assignments).
# +
# used for manipulating directory paths
import os
# Scientific and vector computation for python
import numpy as np
# Plotting library
from matplotlib import pyplot as plt
from mpl_toolkits.mplot3d import Axes3D # needed to plot 3-D surfaces
# library written for this exercise providing additional functions for assignment submission, and others
import utils
# define the submission/grader object for this exercise
grader = utils.Grader()
# tells matplotlib to embed plots within the notebook
# %matplotlib inline
# -
# ## Submission and Grading
#
# After completing each part of the assignment, be sure to submit your solutions to the grader.
#
# For this programming exercise, you are only required to complete the first part of the exercise to implement linear regression with one variable. The second part of the exercise, which is optional, covers linear regression with multiple variables. The following is a breakdown of how each part of this exercise is scored.
#
# **Required Exercises**
#
# | Section | Part |Submitted Function | Points
# |---------|:- |:- | :-:
# | 1 | [Warm up exercise](#section1) | [`warmUpExercise`](#warmUpExercise) | 10
# | 2 | [Compute cost for one variable](#section2) | [`computeCost`](#computeCost) | 40
# | 3 | [Gradient descent for one variable](#section3) | [`gradientDescent`](#gradientDescent) | 50
# | | Total Points | | 100
#
# **Optional Exercises**
#
# | Section | Part | Submitted Function | Points |
# |:-------:|:- |:-: | :-: |
# | 4 | [Feature normalization](#section4) | [`featureNormalize`](#featureNormalize) | 0 |
# | 5 | [Compute cost for multiple variables](#section5) | [`computeCostMulti`](#computeCostMulti) | 0 |
# | 6 | [Gradient descent for multiple variables](#section5) | [`gradientDescentMulti`](#gradientDescentMulti) |0 |
# | 7 | [Normal Equations](#section7) | [`normalEqn`](#normalEqn) | 0 |
#
# You are allowed to submit your solutions multiple times, and we will take only the highest score into consideration.
#
# <div class="alert alert-block alert-warning">
# At the end of each section in this notebook, we have a cell which contains code for submitting the solutions thus far to the grader. Execute the cell to see your score up to the current section. For all your work to be submitted properly, you must execute those cells at least once. They must also be re-executed everytime the submitted function is updated.
# </div>
#
#
# ## Debugging
#
# Here are some things to keep in mind throughout this exercise:
#
# - Python array indices start from zero, not one (contrary to OCTAVE/MATLAB).
#
# - There is an important distinction between python arrays (called `list` or `tuple`) and `numpy` arrays. You should use `numpy` arrays in all your computations. Vector/matrix operations work only with `numpy` arrays. Python lists do not support vector operations (you need to use for loops).
#
# - If you are seeing many errors at runtime, inspect your matrix operations to make sure that you are adding and multiplying matrices of compatible dimensions. Printing the dimensions of `numpy` arrays using the `shape` property will help you debug.
#
# - By default, `numpy` interprets math operators to be element-wise operators. If you want to do matrix multiplication, you need to use the `dot` function in `numpy`. For, example if `A` and `B` are two `numpy` matrices, then the matrix operation AB is `np.dot(A, B)`. Note that for 2-dimensional matrices or vectors (1-dimensional), this is also equivalent to `A@B` (requires python >= 3.5).
# <a id="section1"></a>
# ## 1 Simple python and `numpy` function
#
# The first part of this assignment gives you practice with python and `numpy` syntax and the homework submission process. In the next cell, you will find the outline of a `python` function. Modify it to return a 5 x 5 identity matrix by filling in the following code:
#
# ```python
# A = np.eye(5)
# ```
# <a id="warmUpExercise"></a>
def warmUpExercise():
"""
Example function in Python which computes the identity matrix.
Returns
-------
A : array_like
The 5x5 identity matrix.
Instructions
------------
Return the 5x5 identity matrix.
"""
# ======== YOUR CODE HERE ======
A = np.identity(5) # modify this line
# ==============================
return A
# The previous cell only defines the function `warmUpExercise`. We can now run it by executing the following cell to see its output. You should see output similar to the following:
#
# ```python
# array([[ 1., 0., 0., 0., 0.],
# [ 0., 1., 0., 0., 0.],
# [ 0., 0., 1., 0., 0.],
# [ 0., 0., 0., 1., 0.],
# [ 0., 0., 0., 0., 1.]])
# ```
warmUpExercise()
# ### 1.1 Submitting solutions
#
# After completing a part of the exercise, you can submit your solutions for grading by first adding the function you modified to the grader object, and then sending your function to Coursera for grading.
#
# The grader will prompt you for your login e-mail and submission token. You can obtain a submission token from the web page for the assignment. You are allowed to submit your solutions multiple times, and we will take only the highest score into consideration.
#
# Execute the next cell to grade your solution to the first part of this exercise.
#
# *You should now submit your solutions.*
# +
# appends the implemented function in part 1 to the grader object
grader[1] = warmUpExercise
# send the added functions to coursera grader for getting a grade on this part
grader.grade()
# -
# ## 2 Linear regression with one variable
#
# Now you will implement linear regression with one variable to predict profits for a food truck. Suppose you are the CEO of a restaurant franchise and are considering different cities for opening a new outlet. The chain already has trucks in various cities and you have data for profits and populations from the cities. You would like to use this data to help you select which city to expand to next.
#
# The file `Data/ex1data1.txt` contains the dataset for our linear regression problem. The first column is the population of a city (in 10,000s) and the second column is the profit of a food truck in that city (in $10,000s). A negative value for profit indicates a loss.
#
# We provide you with the code needed to load this data. The dataset is loaded from the data file into the variables `x` and `y`:
# +
# Read comma separated data
data = np.loadtxt(os.path.join('Data', 'ex1data1.txt'), delimiter=',')
X, y = data[:, 0], data[:, 1]
m = y.size # number of training examples
# -
# ### 2.1 Plotting the Data
#
# Before starting on any task, it is often useful to understand the data by visualizing it. For this dataset, you can use a scatter plot to visualize the data, since it has only two properties to plot (profit and population). Many other problems that you will encounter in real life are multi-dimensional and cannot be plotted on a 2-d plot. There are many plotting libraries in python (see this [blog post](https://blog.modeanalytics.com/python-data-visualization-libraries/) for a good summary of the most popular ones).
#
# In this course, we will be exclusively using `matplotlib` to do all our plotting. `matplotlib` is one of the most popular scientific plotting libraries in python and has extensive tools and functions to make beautiful plots. `pyplot` is a module within `matplotlib` which provides a simplified interface to `matplotlib`'s most common plotting tasks, mimicking MATLAB's plotting interface.
#
# <div class="alert alert-block alert-warning">
# You might have noticed that we have imported the `pyplot` module at the beginning of this exercise using the command `from matplotlib import pyplot`. This is rather uncommon, and if you look at python code elsewhere or in the `matplotlib` tutorials, you will see that the module is named `plt`. This is used by module renaming by using the import command `import matplotlib.pyplot as plt`. We will not using the short name of `pyplot` module in this class exercises, but you should be aware of this deviation from norm.
# </div>
#
#
# In the following part, your first job is to complete the `plotData` function below. Modify the function and fill in the following code:
#
# ```python
# pyplot.plot(x, y, 'ro', ms=10, mec='k')
# pyplot.ylabel('Profit in $10,000')
# pyplot.xlabel('Population of City in 10,000s')
# ```
def plotData(x, y):
"""
Plots the data points x and y into a new figure. Plots the data
points and gives the figure axes labels of population and profit.
Parameters
----------
x : array_like
Data point values for x-axis.
y : array_like
Data point values for y-axis. Note x and y should have the same size.
Instructions
------------
Plot the training data into a figure using the "figure" and "plot"
functions. Set the axes labels using the "xlabel" and "ylabel" functions.
Assume the population and revenue data have been passed in as the x
and y arguments of this function.
Hint
----
You can use the 'ro' option with plot to have the markers
appear as red circles. Furthermore, you can make the markers larger by
using plot(..., 'ro', ms=10), where `ms` refers to marker size. You
can also set the marker edge color using the `mec` property.
"""
fig = pyplot.figure() # open a new figure
# ====================== YOUR CODE HERE =======================
plt.scatter(x, y, c='r', s=100, edgecolors='#000000')
plt.ylabel('Profit in $10,000')
plt.xlabel('Population of City in 10,000s')
# plt.show()
# =============================================================
# Now run the defined function with the loaded data to visualize the data. The end result should look like the following figure:
#
# 
#
# Execute the next cell to visualize the data.
plotData(X, y)
# To quickly learn more about the `matplotlib` plot function and what arguments you can provide to it, you can type `?pyplot.plot` in a cell within the jupyter notebook. This opens a separate page showing the documentation for the requested function. You can also search online for plotting documentation.
#
# To set the markers to red circles, we used the option `'or'` within the `plot` function.
# ?pyplot.plot
# <a id="section2"></a>
# ### 2.2 Gradient Descent
#
# In this part, you will fit the linear regression parameters $\theta$ to our dataset using gradient descent.
#
# #### 2.2.1 Update Equations
#
# The objective of linear regression is to minimize the cost function
#
# $$ J(\theta) = \frac{1}{2m} \sum_{i=1}^m \left( h_{\theta}(x^{(i)}) - y^{(i)}\right)^2$$
#
# where the hypothesis $h_\theta(x)$ is given by the linear model
# $$ h_\theta(x) = \theta^Tx = \theta_0 + \theta_1 x_1$$
#
# Recall that the parameters of your model are the $\theta_j$ values. These are
# the values you will adjust to minimize cost $J(\theta)$. One way to do this is to
# use the batch gradient descent algorithm. In batch gradient descent, each
# iteration performs the update
#
# $$ \theta_j = \theta_j - \alpha \frac{1}{m} \sum_{i=1}^m \left( h_\theta(x^{(i)}) - y^{(i)}\right)x_j^{(i)} \qquad \text{simultaneously update } \theta_j \text{ for all } j$$
#
# With each step of gradient descent, your parameters $\theta_j$ come closer to the optimal values that will achieve the lowest cost J($\theta$).
#
# <div class="alert alert-block alert-warning">
# **Implementation Note:** We store each example as a row in the the $X$ matrix in Python `numpy`. To take into account the intercept term ($\theta_0$), we add an additional first column to $X$ and set it to all ones. This allows us to treat $\theta_0$ as simply another 'feature'.
# </div>
#
#
# #### 2.2.2 Implementation
#
# We have already set up the data for linear regression. In the following cell, we add another dimension to our data to accommodate the $\theta_0$ intercept term. Do NOT execute this cell more than once.
# Add a column of ones to X. The numpy function stack joins arrays along a given axis.
# The first axis (axis=0) refers to rows (training examples)
# and second axis (axis=1) refers to columns (features).
X = np.stack([np.ones(m), X], axis=1)
# <a id="section2"></a>
# #### 2.2.3 Computing the cost $J(\theta)$
#
# As you perform gradient descent to learn minimize the cost function $J(\theta)$, it is helpful to monitor the convergence by computing the cost. In this section, you will implement a function to calculate $J(\theta)$ so you can check the convergence of your gradient descent implementation.
#
# Your next task is to complete the code for the function `computeCost` which computes $J(\theta)$. As you are doing this, remember that the variables $X$ and $y$ are not scalar values. $X$ is a matrix whose rows represent the examples from the training set and $y$ is a vector whose each elemennt represent the value at a given row of $X$.
# <a id="computeCost"></a>
def computeCost(X, y, theta):
"""
Compute cost for linear regression. Computes the cost of using theta as the
parameter for linear regression to fit the data points in X and y.
Parameters
----------
X : array_like
The input dataset of shape (m x n+1), where m is the number of examples,
and n is the number of features. We assume a vector of one's already
appended to the features so we have n+1 columns.
y : array_like
The values of the function at each data point. This is a vector of
shape (m, ).
theta : array_like
The parameters for the regression function. This is a vector of
shape (n+1, ).
Returns
-------
J : float
The value of the regression cost function.
Instructions
------------
Compute the cost of a particular choice of theta.
You should set J to the cost.
"""
# initialize some useful values
m = y.size # number of training examples
# You need to return the following variables correctly
J = 0
# ====================== YOUR CODE HERE =====================
ht = np.dot(X, theta)
ht_minus_y = ht - y # ht(x) - y
ht_minus_y = ht_minus_y ** 2
summation = np.sum(ht_minus_y)
J = (1 / (2*m)) * summation
# ===========================================================
return J
# Once you have completed the function, the next step will run `computeCost` two times using two different initializations of $\theta$. You will see the cost printed to the screen.
# +
J = computeCost(X, y, theta=np.array([0.0, 0.0]))
print('With theta = [0, 0] \nCost computed = %.2f' % J)
print('Expected cost value (approximately) 32.07\n')
# further testing of the cost function
J = computeCost(X, y, theta=np.array([-1, 2]))
print('With theta = [-1, 2]\nCost computed = %.2f' % J)
print('Expected cost value (approximately) 54.24')
# -
# *You should now submit your solutions by executing the following cell.*
grader[2] = computeCost
grader.grade()
# <a id="section3"></a>
# #### 2.2.4 Gradient descent
#
# Next, you will complete a function which implements gradient descent.
# The loop structure has been written for you, and you only need to supply the updates to $\theta$ within each iteration.
#
# The equation is -
# $$ \theta_j = \theta_j - \alpha \frac{1}{m} \sum_{i=1}^m \left( h_\theta(x^{(i)}) - y^{(i)}\right)x_j^{(i)} \qquad \text{simultaneously update } \theta_j \text{ for all } j$$
#
# As you program, make sure you understand what you are trying to optimize and what is being updated. Keep in mind that the cost $J(\theta)$ is parameterized by the vector $\theta$, not $X$ and $y$. That is, we minimize the value of $J(\theta)$ by changing the values of the vector $\theta$, not by changing $X$ or $y$. [Refer to the equations in this notebook](#section2) and to the video lectures if you are uncertain. A good way to verify that gradient descent is working correctly is to look at the value of $J(\theta)$ and check that it is decreasing with each step.
#
# The starter code for the function `gradientDescent` calls `computeCost` on every iteration and saves the cost to a `python` list. Assuming you have implemented gradient descent and `computeCost` correctly, your value of $J(\theta)$ should never increase, and should converge to a steady value by the end of the algorithm.
#
# <div class="alert alert-box alert-warning">
# **Vectors and matrices in `numpy`** - Important implementation notes
#
# A vector in `numpy` is a one dimensional array, for example `np.array([1, 2, 3])` is a vector. A matrix in `numpy` is a two dimensional array, for example `np.array([[1, 2, 3], [4, 5, 6]])`. However, the following is still considered a matrix `np.array([[1, 2, 3]])` since it has two dimensions, even if it has a shape of 1x3 (which looks like a vector).
#
# Given the above, the function `np.dot` which we will use for all matrix/vector multiplication has the following properties:
# - It always performs inner products on vectors. If `x=np.array([1, 2, 3])`, then `np.dot(x, x)` is a scalar.
# - For matrix-vector multiplication, so if $X$ is a $m\times n$ matrix and $y$ is a vector of length $m$, then the operation `np.dot(y, X)` considers $y$ as a $1 \times m$ vector. On the other hand, if $y$ is a vector of length $n$, then the operation `np.dot(X, y)` considers $y$ as a $n \times 1$ vector.
# - A vector can be promoted to a matrix using `y[None]` or `[y[np.newaxis]`. That is, if `y = np.array([1, 2, 3])` is a vector of size 3, then `y[None, :]` is a matrix of shape $1 \times 3$. We can use `y[:, None]` to obtain a shape of $3 \times 1$.
# <div>
# <a id="gradientDescent"></a>
def gradientDescent(X, y, theta, alpha, num_iters):
"""
Performs gradient descent to learn `theta`. Updates theta by taking `num_iters`
gradient steps with learning rate `alpha`.
Parameters
----------
X : array_like
The input dataset of shape (m x n+1).
y : arra_like
Value at given features. A vector of shape (m, ).
theta : array_like
Initial values for the linear regression parameters.
A vector of shape (n+1, ).
alpha : float
The learning rate.
num_iters : int
The number of iterations for gradient descent.
Returns
-------
theta : array_like
The learned linear regression parameters. A vector of shape (n+1, ).
J_history : list
A python list for the values of the cost function after each iteration.
Instructions
------------
Peform a single gradient step on the parameter vector theta.
While debugging, it can be useful to print out the values of
the cost function (computeCost) and gradient here.
"""
# Initialize some useful values
m = y.shape[0] # number of training examples
# make a copy of theta, to avoid changing the original array, since numpy arrays
# are passed by reference to functions
theta = theta.copy()
J_history = [] # Use a python list to save cost in every iteration
for i in range(num_iters):
# ==================== YOUR CODE HERE =================================
ht = np.dot(X,theta)
ht_minus_y = ht - y # ht(x) - y
s = np.dot(ht_minus_y , X)
delta = (1/m) * s
theta = theta - alpha * delta
# =====================================================================
# save the cost J in every iteration
J_history.append(computeCost(X, y, theta))
return theta, J_history
# After you are finished call the implemented `gradientDescent` function and print the computed $\theta$. We initialize the $\theta$ parameters to 0 and the learning rate $\alpha$ to 0.01. Execute the following cell to check your code.
# +
# initialize fitting parameters
theta = np.zeros(2)
# some gradient descent settings
iterations = 1500
alpha = 0.01
theta, J_history = gradientDescent(X ,y, theta, alpha, iterations)
print('Theta found by gradient descent: {:.4f}, {:.4f}'.format(*theta))
print('Expected theta values (approximately): [-3.6303, 1.1664]')
# -
# We will use your final parameters to plot the linear fit. The results should look like the following figure.
#
# 
# +
# plot the linear fit
plotData(X[:, 1], y)
plt.plot(X[:, 1], np.dot(X, theta), '-')
pyplot.legend([ 'Linear regression', 'Training data']);
# -
# Your final values for $\theta$ will also be used to make predictions on profits in areas of 35,000 and 70,000 people.
#
# <div class="alert alert-block alert-success">
# Note the way that the following lines use matrix multiplication, rather than explicit summation or looping, to calculate the predictions. This is an example of code vectorization in `numpy`.
# </div>
#
# <div class="alert alert-block alert-success">
# Note that the first argument to the `numpy` function `dot` is a python list. `numpy` can internally converts **valid** python lists to numpy arrays when explicitly provided as arguments to `numpy` functions.
# </div>
#
# +
# Predict values for population sizes of 35,000 and 70,000
predict1 = np.dot([1, 3.5], theta)
print('For population = 35,000, we predict a profit of {:.2f}\n'.format(predict1*10000))
predict2 = np.dot([1, 7], theta)
print('For population = 70,000, we predict a profit of {:.2f}\n'.format(predict2*10000))
# -
# *You should now submit your solutions by executing the next cell.*
grader[3] = gradientDescent
grader.grade()
# ### 2.4 Visualizing $J(\theta)$
#
# To understand the cost function $J(\theta)$ better, you will now plot the cost over a 2-dimensional grid of $\theta_0$ and $\theta_1$ values. You will not need to code anything new for this part, but you should understand how the code you have written already is creating these images.
#
# In the next cell, the code is set up to calculate $J(\theta)$ over a grid of values using the `computeCost` function that you wrote. After executing the following cell, you will have a 2-D array of $J(\theta)$ values. Then, those values are used to produce surface and contour plots of $J(\theta)$ using the matplotlib `plot_surface` and `contourf` functions. The plots should look something like the following:
#
# 
#
# The purpose of these graphs is to show you how $J(\theta)$ varies with changes in $\theta_0$ and $\theta_1$. The cost function $J(\theta)$ is bowl-shaped and has a global minimum. (This is easier to see in the contour plot than in the 3D surface plot). This minimum is the optimal point for $\theta_0$ and $\theta_1$, and each step of gradient descent moves closer to this point.
# +
# grid over which we will calculate J
theta0_vals = np.linspace(-10, 10, 100)
theta1_vals = np.linspace(-1, 4, 100)
# initialize J_vals to a matrix of 0's
J_vals = np.zeros((theta0_vals.shape[0], theta1_vals.shape[0]))
# Fill out J_vals
for i, theta0 in enumerate(theta0_vals):
for j, theta1 in enumerate(theta1_vals):
J_vals[i, j] = computeCost(X, y, [theta0, theta1])
# Because of the way meshgrids work in the surf command, we need to
# transpose J_vals before calling surf, or else the axes will be flipped
J_vals = J_vals.T
# surface plot
fig = pyplot.figure(figsize=(12, 5))
ax = fig.add_subplot(121, projection='3d')
ax.plot_surface(theta0_vals, theta1_vals, J_vals, cmap='viridis')
pyplot.xlabel('theta0')
pyplot.ylabel('theta1')
pyplot.title('Surface')
# contour plot
# Plot J_vals as 15 contours spaced logarithmically between 0.01 and 100
ax = pyplot.subplot(122)
pyplot.contour(theta0_vals, theta1_vals, J_vals, linewidths=2, cmap='viridis', levels=np.logspace(-2, 3, 20))
pyplot.xlabel('theta0')
pyplot.ylabel('theta1')
pyplot.plot(theta[0], theta[1], 'ro', ms=10, lw=2)
pyplot.title('Contour, showing minimum')
pass
# -
# ## Optional Exercises
#
# If you have successfully completed the material above, congratulations! You now understand linear regression and should able to start using it on your own datasets.
#
# For the rest of this programming exercise, we have included the following optional exercises. These exercises will help you gain a deeper understanding of the material, and if you are able to do so, we encourage you to complete them as well. You can still submit your solutions to these exercises to check if your answers are correct.
#
# ## 3 Linear regression with multiple variables
#
# In this part, you will implement linear regression with multiple variables to predict the prices of houses. Suppose you are selling your house and you want to know what a good market price would be. One way to do this is to first collect information on recent houses sold and make a model of housing prices.
#
# The file `Data/ex1data2.txt` contains a training set of housing prices in Portland, Oregon. The first column is the size of the house (in square feet), the second column is the number of bedrooms, and the third column is the price
# of the house.
#
# <a id="section4"></a>
# ### 3.1 Feature Normalization
#
# We start by loading and displaying some values from this dataset. By looking at the values, note that house sizes are about 1000 times the number of bedrooms. When features differ by orders of magnitude, first performing feature scaling can make gradient descent converge much more quickly.
# +
# Load data
data = np.loadtxt(os.path.join('Data', 'ex1data2.txt'), delimiter=',')
X = data[:, :2]
y = data[:, 2]
m = y.size
# print out some data points
print('{:>8s}{:>8s}{:>10s}'.format('X[:,0]', 'X[:, 1]', 'y'))
print('-'*26)
for i in range(10):
print('{:8.0f}{:8.0f}{:10.0f}'.format(X[i, 0], X[i, 1], y[i]))
# -
# Your task here is to complete the code in `featureNormalize` function:
# - Subtract the mean value of each feature from the dataset.
# - After subtracting the mean, additionally scale (divide) the feature values by their respective “standard deviations.”
#
# The standard deviation is a way of measuring how much variation there is in the range of values of a particular feature (most data points will lie within ±2 standard deviations of the mean); this is an alternative to taking the range of values (max-min). In `numpy`, you can use the `std` function to compute the standard deviation.
#
# For example, the quantity `X[:, 0]` contains all the values of $x_1$ (house sizes) in the training set, so `np.std(X[:, 0])` computes the standard deviation of the house sizes.
# At the time that the function `featureNormalize` is called, the extra column of 1’s corresponding to $x_0 = 1$ has not yet been added to $X$.
#
# You will do this for all the features and your code should work with datasets of all sizes (any number of features / examples). Note that each column of the matrix $X$ corresponds to one feature.
#
# <div class="alert alert-block alert-warning">
# **Implementation Note:** When normalizing the features, it is important
# to store the values used for normalization - the mean value and the standard deviation used for the computations. After learning the parameters
# from the model, we often want to predict the prices of houses we have not
# seen before. Given a new x value (living room area and number of bedrooms), we must first normalize x using the mean and standard deviation that we had previously computed from the training set.
# </div>
# <a id="featureNormalize"></a>
def featureNormalize(X):
"""
Normalizes the features in X. returns a normalized version of X where
the mean value of each feature is 0 and the standard deviation
is 1. This is often a good preprocessing step to do when working with
learning algorithms.
Parameters
----------
X : array_like
The dataset of shape (m x n).
Returns
-------
X_norm : array_like
The normalized dataset of shape (m x n).
Instructions
------------
First, for each feature dimension, compute the mean of the feature
and subtract it from the dataset, storing the mean value in mu.
Next, compute the standard deviation of each feature and divide
each feature by it's standard deviation, storing the standard deviation
in sigma.
Note that X is a matrix where each column is a feature and each row is
an example. You needto perform the normalization separately for each feature.
Hint
----
You might find the 'np.mean' and 'np.std' functions useful.
"""
# You need to set these values correctly
X_norm = X.copy()
mu = np.zeros(X.shape[1])
sigma = np.zeros(X.shape[1])
# =========================== YOUR CODE HERE =====================
mu = np.mean(X_norm, axis = 0)
# mu = np.sum(X_norm, axis = 0) / X.shape[0]
sigma = np.std(X_norm , axis = 0)
X_norm = (X_norm - mu) / sigma
# ================================================================
return X_norm, mu, sigma
# Execute the next cell to run the implemented `featureNormalize` function.
# +
# call featureNormalize on the loaded data
X_norm, mu, sigma = featureNormalize(X)
print('Computed mean:', mu)
print('Computed standard deviation:', sigma)
# -
# *You should now submit your solutions.*
grader[4] = featureNormalize
grader.grade()
# After the `featureNormalize` function is tested, we now add the intercept term to `X_norm`:
# Add intercept term to X
X = np.concatenate([np.ones((m, 1)), X_norm], axis=1)
# <a id="section5"></a>
# ### 3.2 Gradient Descent
#
# Previously, you implemented gradient descent on a univariate regression problem. The only difference now is that there is one more feature in the matrix $X$. The hypothesis function and the batch gradient descent update
# rule remain unchanged.
#
# You should complete the code for the functions `computeCostMulti` and `gradientDescentMulti` to implement the cost function and gradient descent for linear regression with multiple variables. If your code in the previous part (single variable) already supports multiple variables, you can use it here too.
# Make sure your code supports any number of features and is well-vectorized.
# You can use the `shape` property of `numpy` arrays to find out how many features are present in the dataset.
#
# <div class="alert alert-block alert-warning">
# **Implementation Note:** In the multivariate case, the cost function can
# also be written in the following vectorized form:
#
# $$ J(\theta) = \frac{1}{2m}(X\theta - \vec{y})^T(X\theta - \vec{y}) $$
#
# where
#
# $$ X = \begin{pmatrix}
# - (x^{(1)})^T - \\
# - (x^{(2)})^T - \\
# \vdots \\
# - (x^{(m)})^T - \\ \\
# \end{pmatrix} \qquad \mathbf{y} = \begin{bmatrix} y^{(1)} \\ y^{(2)} \\ \vdots \\ y^{(m)} \\\end{bmatrix}$$
#
# the vectorized version is efficient when you are working with numerical computing tools like `numpy`. If you are an expert with matrix operations, you can prove to yourself that the two forms are equivalent.
# </div>
#
# <a id="computeCostMulti"></a>
def computeCostMulti(X, y, theta):
"""
Compute cost for linear regression with multiple variables.
Computes the cost of using theta as the parameter for linear regression to fit the data points in X and y.
Parameters
----------
X : array_like
The dataset of shape (m x n+1).
y : array_like
A vector of shape (m, ) for the values at a given data point.
theta : array_like
The linear regression parameters. A vector of shape (n+1, )
Returns
-------
J : float
The value of the cost function.
Instructions
------------
Compute the cost of a particular choice of theta. You should set J to the cost.
"""
# Initialize some useful values
m = y.shape[0] # number of training examples
# You need to return the following variable correctly
J = 0
# ======================= YOUR CODE HERE ===========================
ht = np.dot(X, theta)
tem = (ht - y)**2
summ = np.sum(tem, axis = 0)
J = (1/ (2*m)) * summ
# ==================================================================
return J
# *You should now submit your solutions.*
grader[5] = computeCostMulti
grader.grade()
# <a id="gradientDescentMulti"></a>
def gradientDescentMulti(X, y, theta, alpha, num_iters):
"""
Performs gradient descent to learn theta.
Updates theta by taking num_iters gradient steps with learning rate alpha.
Parameters
----------
X : array_like
The dataset of shape (m x n+1).
y : array_like
A vector of shape (m, ) for the values at a given data point.
theta : array_like
The linear regression parameters. A vector of shape (n+1, )
alpha : float
The learning rate for gradient descent.
num_iters : int
The number of iterations to run gradient descent.
Returns
-------
theta : array_like
The learned linear regression parameters. A vector of shape (n+1, ).
J_history : list
A python list for the values of the cost function after each iteration.
Instructions
------------
Peform a single gradient step on the parameter vector theta.
While debugging, it can be useful to print out the values of
the cost function (computeCost) and gradient here.
"""
# Initialize some useful values
m = y.shape[0] # number of training examples
# make a copy of theta, which will be updated by gradient descent
theta = theta.copy()
J_history = []
for i in range(num_iters):
# ======================= YOUR CODE HERE ==========================
ht = np.dot(X, theta)
ht_minus_y = ht - y
s = np.dot(ht_minus_y , X)
theta = theta - alpha * (1/m) * s
# =================================================================
# save the cost J in every iteration
J_history.append(computeCostMulti(X, y, theta))
return theta, J_history
# +
# initialize fitting parameters
# some gradient descent settings
iterations = 1500
alpha = 0.01
theta, J_history = gradientDescentMulti(X ,y, theta = np.array([0, 0, 0]) , alpha = alpha,num_iters=iterations)
print(theta)
# -
# *You should now submit your solutions.*
grader[6] = gradientDescentMulti
grader.grade()
# #### 3.2.1 Optional (ungraded) exercise: Selecting learning rates
#
# In this part of the exercise, you will get to try out different learning rates for the dataset and find a learning rate that converges quickly. You can change the learning rate by modifying the following code and changing the part of the code that sets the learning rate.
#
# Use your implementation of `gradientDescentMulti` function and run gradient descent for about 50 iterations at the chosen learning rate. The function should also return the history of $J(\theta)$ values in a vector $J$.
#
# After the last iteration, plot the J values against the number of the iterations.
#
# If you picked a learning rate within a good range, your plot look similar as the following Figure.
#
# 
#
# If your graph looks very different, especially if your value of $J(\theta)$ increases or even blows up, adjust your learning rate and try again. We recommend trying values of the learning rate $\alpha$ on a log-scale, at multiplicative steps of about 3 times the previous value (i.e., 0.3, 0.1, 0.03, 0.01 and so on). You may also want to adjust the number of iterations you are running if that will help you see the overall trend in the curve.
#
# <div class="alert alert-block alert-warning">
# **Implementation Note:** If your learning rate is too large, $J(\theta)$ can diverge and ‘blow up’, resulting in values which are too large for computer calculations. In these situations, `numpy` will tend to return
# NaNs. NaN stands for ‘not a number’ and is often caused by undefined operations that involve −∞ and +∞.
# </div>
#
# <div class="alert alert-block alert-warning">
# **MATPLOTLIB tip:** To compare how different learning learning rates affect convergence, it is helpful to plot $J$ for several learning rates on the same figure. This can be done by making `alpha` a python list, and looping across the values within this list, and calling the plot function in every iteration of the loop. It is also useful to have a legend to distinguish the different lines within the plot. Search online for `pyplot.legend` for help on showing legends in `matplotlib`.
# </div>
#
# Notice the changes in the convergence curves as the learning rate changes. With a small learning rate, you should find that gradient descent takes a very long time to converge to the optimal value. Conversely, with a large learning rate, gradient descent might not converge or might even diverge!
# Using the best learning rate that you found, run the script
# to run gradient descent until convergence to find the final values of $\theta$. Next,
# use this value of $\theta$ to predict the price of a house with 1650 square feet and
# 3 bedrooms. You will use value later to check your implementation of the normal equations. Don’t forget to normalize your features when you make this prediction!
# +
"""
Instructions
------------
We have provided you with the following starter code that runs
gradient descent with a particular learning rate (alpha).
Your task is to first make sure that your functions - `computeCost`
and `gradientDescent` already work with this starter code and
support multiple variables.
After that, try running gradient descent with different values of
alpha and see which one gives you the best result.
Finally, you should complete the code at the end to predict the price
of a 1650 sq-ft, 3 br house.
Hint
----
At prediction, make sure you do the same feature normalization.
"""
# Choose some alpha value - change this
alpha = 0.1
num_iters = 400
# init theta and run gradient descent
theta = np.zeros(3)
theta, J_history = gradientDescentMulti(X, y, theta, alpha, num_iters)
# Plot the convergence graph
pyplot.plot(np.arange(len(J_history)), J_history, lw=2)
pyplot.xlabel('Number of iterations')
pyplot.ylabel('Cost J')
# Display the gradient descent's result
print('theta computed from gradient descent: {:s}'.format(str(theta)))
# Estimate the price of a 1650 sq-ft, 3 br house
# ======================= YOUR CODE HERE ===========================
# Recall that the first column of X is all-ones.
# Thus, it does not need to be normalized.
price = 0 # You should change this
# ===================================================================
print('Predicted price of a 1650 sq-ft, 3 br house (using gradient descent): ${:.0f}'.format(price))
# -
# *You do not need to submit any solutions for this optional (ungraded) part.*
# <a id="section7"></a>
# ### 3.3 Normal Equations
#
# In the lecture videos, you learned that the closed-form solution to linear regression is
#
# $$ \theta = \left( X^T X\right)^{-1} X^T\vec{y}$$
#
# Using this formula does not require any feature scaling, and you will get an exact solution in one calculation: there is no “loop until convergence” like in gradient descent.
#
# First, we will reload the data to ensure that the variables have not been modified. Remember that while you do not need to scale your features, we still need to add a column of 1’s to the $X$ matrix to have an intercept term ($\theta_0$). The code in the next cell will add the column of 1’s to X for you.
# Load data
data = np.loadtxt(os.path.join('Data', 'ex1data2.txt'), delimiter=',')
X = data[:, :2]
y = data[:, 2]
m = y.size
X = np.concatenate([np.ones((m, 1)), X], axis=1)
# Complete the code for the function `normalEqn` below to use the formula above to calculate $\theta$.
#
# <a id="normalEqn"></a>
def normalEqn(X, y):
"""
Computes the closed-form solution to linear regression using the normal equations.
Parameters
----------
X : array_like
The dataset of shape (m x n+1).
y : array_like
The value at each data point. A vector of shape (m, ).
Returns
-------
theta : array_like
Estimated linear regression parameters. A vector of shape (n+1, ).
Instructions
------------
Complete the code to compute the closed form solution to linear
regression and put the result in theta.
Hint
----
Look up the function `np.linalg.pinv` for computing matrix inverse.
"""
theta = np.zeros(X.shape[1])
# ===================== YOUR CODE HERE ============================
# =================================================================
return theta
# *You should now submit your solutions.*
grader[7] = normalEqn
grader.grade()
# Optional (ungraded) exercise: Now, once you have found $\theta$ using this
# method, use it to make a price prediction for a 1650-square-foot house with
# 3 bedrooms. You should find that gives the same predicted price as the value
# you obtained using the model fit with gradient descent (in Section 3.2.1).
# +
# Calculate the parameters from the normal equation
theta = normalEqn(X, y);
# Display normal equation's result
print('Theta computed from the normal equations: {:s}'.format(str(theta)));
# Estimate the price of a 1650 sq-ft, 3 br house
# ====================== YOUR CODE HERE ======================
price = 0 # You should change this
# ============================================================
print('Predicted price of a 1650 sq-ft, 3 br house (using normal equations): ${:.0f}'.format(price))
| Assignment-1/Given-Materials-Python/.ipynb_checkpoints/exercise1-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: thesis_language_change
# language: python
# name: thesis_language_change
# ---
# # KFA for the Chapter
#
# In this notebook, we will run a method based on Usage Fluctuation Analysis (UFA) that uses keywords as features rather than collocates.
# +
# %matplotlib inline
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.cm as cm
import seaborn as sns
import sys
import os
import json
import itertools
import sqlite3
import regex as re
from datetime import datetime
from collections import Counter
from sklearn.preprocessing import StandardScaler
from nltk import ngrams as make_ngrams
# +
GRAPH_DIR = "./Graphs"
out_dir = GRAPH_DIR
sys.path.insert(1, "../")
sys.path.insert(1, "../utilities")
from settings import DB_FP
from helper_functions import split_corpus
# -
sql_get_all_posts ="""
SELECT c.uid, m.name, m.PimsId, p.party, d.date, c.body, c.topic, c.section, s.tmay_deal, s.benn_act, s.ref_stance, s.constituency_leave, c.usas_file
FROM contributions as c
INNER JOIN members as m
ON m.PimsId = c.member
INNER JOIN debates as d
ON d.uid = c.debate
INNER JOIN member_party as p
ON p.PimsId = m.PimsId
INNER JOIN member_stances as s
ON s.PimsId = m.PimsId
WHERE (d.date BETWEEN date("2015-05-01") AND date("2019-12-11"))
AND (((d.date BETWEEN p.start AND p.end) AND NOT (p.end IS NULL))
OR ((d.date >= p.start) AND (p.end IS NULL)));""".strip()
from language_change_methods import vnc
from language_change_methods.utility_functions import tokenise
from language_change_methods.features import function_words
with open("../resources/speakers.json") as speaker_file:
speaker_list = json.load(speaker_file)
# +
# %%time
conn = sqlite3.connect(DB_FP)
curs = conn.cursor()
# Gets all the contributions and creates a nice dataframe
all_contributions = pd.read_sql_query(sql_get_all_posts, conn)
all_contributions.columns = ['uid', 'name', 'PimsId', 'party', 'date', 'text', 'topic', 'section', 'tmay_deal', 'benn_act', 'ref_stance', 'constituency_leave', 'usas_file']
all_contributions.set_index("uid", inplace=True)
convert_to_date = lambda x: datetime.strptime(x, "%Y-%m-%d %H:%M:%S")
all_contributions['date'] = all_contributions['date'].apply(convert_to_date)
all_contributions = all_contributions.query("PimsId not in @speaker_list")
all_contributions.sort_values("date", inplace=True)
# Tokenise the contributions
all_toks = all_contributions["text"].apply(tokenise)
# Get the EU and Non-EU mentions
eu_mentions, non_eu_mentions = split_corpus(all_contributions, "eu")
# -
# # Create the Keyword Feature Matrix
from language_change_methods.features import get_ngram_lr_and_ll, get_wordcounts_multiple_texts
# %%time
# Get key words for EU debate
kw = get_ngram_lr_and_ll(all_toks.loc[eu_mentions.index], all_toks.loc[non_eu_mentions.index], 1)
merge_lists = lambda x: list(itertools.chain.from_iterable(x))
# %%time
all_counts = Counter(merge_lists(all_toks))
kw_list = list(kw[kw["LR"] > 1].index)
kw_list = [k for k in kw_list if all_counts[k] > 100]
from language_change_methods.utility_functions import get_data_windows
def create_group_kw_matrices(contributions, tokens, group_indexes, win_size, win_step, w_list):
win_counts = {gname: dict() for gname in group_indexes}
n_words_per_window = {gname: dict() for gname in group_indexes}
for window, contribs in get_data_windows(contributions, win_size, win_step, time_column="date"):
for gname in group_indexes:
g_contribs = contribs.loc[contribs.index.isin(group_indexes[gname])]
curr_counts = get_wordcounts_multiple_texts(tokens.loc[g_contribs.index])
curr_counts = pd.Series(curr_counts)
curr_total = curr_counts.sum()
curr_counts = curr_counts[curr_counts.index.isin(w_list)]
curr_counts = curr_counts / curr_total
win_counts[gname][window] = curr_counts
win_counts = {g: pd.DataFrame(win_counts[g]).T.fillna(0) for g in group_indexes}
return win_counts
# +
con = all_contributions[all_contributions["party"]=="Conservative"]
lab = all_contributions[all_contributions["party"]=="Labour"]
print("{0:12}: {1:10}".format("Conservative", con.shape[0]))
print("{0:12}: {1:10}".format("Labour", lab.shape[0]))
# -
# %%time
g_bow = create_group_kw_matrices(contributions=all_contributions, tokens=all_toks,
group_indexes={"Conservative": con.index, "Labour": lab.index},
win_size=50000, win_step=10000, w_list=kw_list)
from language_change_methods.fluctuation_analysis import fluct_anal, calc_ac1, plot_gam_of_series
# This method calculates cosine distance between two vectors.
from scipy.spatial.distance import cosine as cosine_dist
# This method simply inverts it to get similarity.
cosine_sim = lambda x,y: 1 - cosine_dist(x,y)
# %%time
# Make a collocate matrix
fig, ax = plt.subplots(figsize=(10,6))
fluct = fluct_anal(g_bow["Conservative"], cosine_sim)
ax.scatter(fluct.index, fluct.values, alpha=0.4)
plot_gam_of_series(fluct, ax)
ax.grid()
plt.show()
# +
# %%time
# Make a collocate matrix
fig, ax = plt.subplots(figsize=(10,6))
df = g_bow["Conservative"]
df = df.applymap(lambda x: True if x > 0 else False)
fluct = fluct_anal(df, calc_ac1)
ax.scatter(fluct.index, fluct.values, alpha=0.4)
plot_gam_of_series(fluct, ax)
ax.grid()
plt.show()
# -
# # Get Keywords at each Window
def get_kw_per_window(contributions, tokens, group_indexes, subset, win_size, win_step, min_freq=10):
win_counts = {gname: dict() for gname in group_indexes}
n_words_per_window = {gname: dict() for gname in group_indexes}
for window, contribs in get_data_windows(contributions, win_size, win_step, time_column="date"):
for gname in group_indexes:
g_contribs = contribs.loc[contribs.index.isin(group_indexes[gname])]
g_toks = tokens.loc[g_contribs.index]
curr_counts = get_wordcounts_multiple_texts(g_toks[g_toks.index.isin(subset)])
# Get key words for EU debate
kw = get_ngram_lr_and_ll(g_toks[g_toks.index.isin(subset)], g_toks[~g_toks.index.isin(subset)], 1)
kw_list = list(kw[kw["LR"] > 1].index)
kw_list = [k for k in kw_list if curr_counts[k] > min_freq]
win_counts[gname][window] = pd.Series({w: curr_counts[w] for w in kw_list})
win_counts = {g: pd.DataFrame(win_counts[g]).T.fillna(0) for g in group_indexes}
return win_counts
# %%time
g_kw_over_time = get_kw_per_window(contributions=all_contributions, tokens=all_toks,
group_indexes={"Conservative": con.index, "Labour": lab.index},
subset=eu_mentions.index,
win_size=50000, win_step=10000, min_freq=50)
def plot_group_fluct(g_kw_over_time, g_colours, comp_method=calc_ac1, binary=True, ax=None):
# Make a collocate matrix
if ax is None:
fig, ax = plt.subplots(figsize=(10,6))
for g in g_kw_over_time:
df = g_kw_over_time[g]
if binary:
df = df.applymap(lambda x: True if x > 0 else False)
fluct = fluct_anal(df, comp_method)
ax.scatter(fluct.index, fluct.values, alpha=0.4, color=g_colours[g])
plot_gam_of_series(fluct, ax, line_colour=g_colours[g], label=g)
if ax is None:
ax.grid()
plt.show()
# +
from language_change_methods.fluctuation_analysis import comp_anal
def plot_group_comparison(g1, g2, comp_method=calc_ac1, binary=True, ax=None, colour=None, label=None):
# Make a collocate matrix
if ax is None:
fig, ax = plt.subplots(figsize=(10,6))
if binary:
df1 = g1.applymap(lambda x: True if x > 0 else False)
df2 = g2.applymap(lambda x: True if x > 0 else False)
else:
df1 = g1
df2 = g2
fluct = comp_anal(df1, df2, comp_method)
ax.scatter(fluct.index, fluct.values, alpha=0.4, color=colour)
plot_gam_of_series(fluct, ax, line_colour=colour, label=label)
if ax is None:
ax.grid()
plt.show()
# +
fig, ax = plt.subplots(figsize=(10,6))
plot_group_fluct(g_kw_over_time, {"Conservative": "blue", "Labour": "red"}, ax=ax)
ax.xaxis.set_tick_params(labelsize=14, rotation=0)
ax.yaxis.set_tick_params(labelsize=14)
ax.set_xlabel("Time", fontsize=16)
ax.set_ylabel("AC1 Agreement", fontsize=16)
ax.legend(fontsize=14)
plt.tight_layout(pad=0)
ax.grid()
fig.savefig(os.path.join(GRAPH_DIR, "kfa_fluctuation.pdf"))
plt.show()
# +
# # %%time
# for mf in [50]:
# g_kw_over_time = get_kw_per_window(contributions=all_contributions, tokens=all_toks,
# group_indexes={"Conservative": con.index, "Labour": lab.index},
# subset=eu_mentions.index,
# win_size=50000, win_step=10000, min_freq=mf)
# plot_group_fluct(g_kw_over_time, {"Conservative": "blue", "Labour": "red"})
# -
# ### Match Columns of Both DataFrames
from language_change_methods.fluctuation_analysis import add_missing_columns, make_dfs_comparable
# +
df1 = g_kw_over_time["Conservative"]
df2 = g_kw_over_time["Labour"]
df1, df2 = make_dfs_comparable(df1, df2)
df1 = df1 / df1.sum(axis=1).values[:,None]
df2 = df2 / df2.sum(axis=1).values[:,None]
# -
plot_group_fluct({"Conservative": df1, "Labour": df2}, {"Conservative": "blue", "Labour": "red"}, comp_method=calc_ac1, binary=True)
plot_group_fluct({"Conservative": df1, "Labour": df2}, {"Conservative": "blue", "Labour": "red"}, comp_method=cosine_sim, binary=False)
# +
fig, ax = plt.subplots(figsize=(10,6))
plot_group_comparison(df1, df2, comp_method=calc_ac1, binary=True, ax=ax, colour="purple")
ax.xaxis.set_tick_params(labelsize=14, rotation=0)
ax.yaxis.set_tick_params(labelsize=14)
ax.set_xlabel("Time", fontsize=16)
ax.set_ylabel("AC1 Agreement", fontsize=16)
plt.tight_layout(pad=0)
ax.grid()
fig.savefig(os.path.join(GRAPH_DIR, "kfa_comparison.pdf"))
plt.show()
# +
fig, ax = plt.subplots(figsize=(10,6))
plot_group_comparison(df1, df2, comp_method=cosine_sim, binary=False, colour="purple", ax=ax)
ax.xaxis.set_tick_params(labelsize=14, rotation=0)
ax.yaxis.set_tick_params(labelsize=14)
ax.set_xlabel("Time", fontsize=16)
ax.set_ylabel("Cosine Similarity", fontsize=16)
plt.tight_layout(pad=0)
ax.grid()
fig.savefig(os.path.join(GRAPH_DIR, "kfa_comparison_cosine.pdf"))
plt.show()
# +
# Make a collocate matrix
fig, (ax1, ax2) = plt.subplots(2, figsize=(10,6), sharex=True)
fluct = comp_anal(g_kw_over_time["Conservative"].applymap(lambda x: True if x > 0 else False),
g_kw_over_time["Labour"].applymap(lambda x: True if x > 0 else False),
calc_ac1)
ax.scatter(fluct.index, fluct.values, alpha=0.4)
plot_gam_of_series(fluct, ax1)
plot_gam_of_series(g_kw_over_time["Conservative"].sum(axis=1), ax2)
ax2.xaxis.set_tick_params(labelsize=14, rotation=0)
ax2.yaxis.set_tick_params(labelsize=14)
ax1.yaxis.set_tick_params(labelsize=14)
ax2.set_xlabel("Time", fontsize=16)
ax2.set_ylabel("Num Keywords", fontsize=16)
ax1.set_ylabel("AC1 Agreement", fontsize=16)
ax1.grid()
ax2.grid()
plt.tight_layout(pad=0)
fig.savefig(os.path.join(GRAPH_DIR, "kfa_comparison_to_num_keywords.pdf"))
plt.show()
# -
# ## Comparing to a Reference
def create_freq_matrices(contributions, tokens, indices, win_size, win_step, w_list):
win_counts = dict()
for window, contribs in get_data_windows(contributions, win_size, win_step, time_column="date"):
g_contribs = contribs.loc[contribs.index.isin(indices)]
curr_counts = get_wordcounts_multiple_texts(tokens.loc[g_contribs.index])
curr_counts = pd.Series(curr_counts)
curr_counts = curr_counts[curr_counts.index.isin(w_list)]
win_counts[window] = curr_counts
return pd.DataFrame(win_counts).T.fillna(0)
# %%time
ref_counts = create_freq_matrices(all_contributions, all_toks, non_eu_mentions.index, 50000, 10000, df1.columns)
ref_counts = make_dfs_comparable(ref_counts, df1)[0]
ref_counts = ref_counts / ref_counts.sum(axis=1).values[:,None]
fig, ax = plt.subplots(figsize=(10,6))
plot_group_comparison(df1, ref_counts, comp_method=calc_ac1, binary=True, ax=ax, colour="blue", label="Conservative to Reference")
plot_group_comparison(df2, ref_counts, comp_method=calc_ac1, binary=True, ax=ax, colour="red", label="Labour to Reference")
plt.legend()
ax.grid()
plt.show()
fig, ax = plt.subplots(figsize=(10,6))
plot_group_comparison(df1, ref_counts, comp_method=cosine_sim, binary=False, ax=ax, colour="blue", label="Conservative to Reference")
plot_group_comparison(df2, ref_counts, comp_method=cosine_sim, binary=False, ax=ax, colour="red", label="Labour to Reference")
ax.grid()
plt.legend()
plt.show()
# # KFA so both groups have the same features
def get_kw_per_window_same_kws(contributions, tokens, group_indexes, subset, win_size, win_step, min_freq=10):
win_counts = {gname: dict() for gname in group_indexes}
for window, contribs in get_data_windows(contributions, win_size, win_step, time_column="date"):
w_toks = tokens.loc[contribs.index]
w_counts = get_wordcounts_multiple_texts(w_toks[w_toks.index.isin(subset)])
# Get key words for EU debate
kw = get_ngram_lr_and_ll(w_toks[w_toks.index.isin(subset)], w_toks[~w_toks.index.isin(subset)], 1)
kw_list = list(kw[kw["LR"] > 1].index)
kw_list = [k for k in kw_list if w_counts[k] > min_freq]
for gname in group_indexes:
g_contribs = contribs.loc[contribs.index.isin(group_indexes[gname])]
g_toks = tokens.loc[g_contribs.index]
g_counts = get_wordcounts_multiple_texts(g_toks[g_toks.index.isin(subset)])
curr_total = sum(g_counts.values())
win_counts[gname][window] = pd.Series({w: g_counts[w] / curr_total for w in kw_list})
win_counts = {g: pd.DataFrame(win_counts[g]).T.fillna(0) for g in group_indexes}
return win_counts
# %%time
for mf in [50]:
g_kw_over_time = get_kw_per_window_same_kws(contributions=all_contributions, tokens=all_toks,
group_indexes={"Conservative": con.index, "Labour": lab.index},
subset=eu_mentions.index,
win_size=50000, win_step=10000, min_freq=mf)
plot_group_fluct(g_kw_over_time, {"Conservative": "blue", "Labour": "red"})
# %%time
# Make a collocate matrix
fig, ax = plt.subplots(figsize=(10,6))
for g, c in zip(["Conservative", "Labour"], ["blue", "red"]):
fluct = fluct_anal(g_kw_over_time[g], cosine_sim)
ax.scatter(fluct.index, fluct.values, alpha=0.4, color=c)
plot_gam_of_series(fluct, ax, line_colour=c)
ax.grid()
plt.show()
# ## Fixing it so only keywords of specific group are counted
from language_change_methods.utility_functions import get_log_ratio
def get_kw_per_window_same_kws(contributions, tokens, group_indexes, subset, win_size, win_step, min_freq=10):
win_counts = {gname: dict() for gname in group_indexes}
for window, contribs in get_data_windows(contributions, win_size, win_step, time_column="date"):
w_toks = tokens.loc[contribs.index]
w_counts = get_wordcounts_multiple_texts(w_toks[w_toks.index.isin(subset)])
# Get key words for EU debate
kw = get_ngram_lr_and_ll(w_toks[w_toks.index.isin(subset)], w_toks[~w_toks.index.isin(subset)], 1)
kw_list = list(kw[kw["LR"] > 1].index)
kw_list = [k for k in kw_list if w_counts[k] > min_freq]
for gname in group_indexes:
g_contribs = contribs.loc[contribs.index.isin(group_indexes[gname])]
g_toks = tokens.loc[g_contribs.index]
g_sub_counts = get_wordcounts_multiple_texts(g_toks[g_toks.index.isin(subset)])
g_oth_counts = get_wordcounts_multiple_texts(g_toks[~g_toks.index.isin(subset)])
curr_total = sum(g_sub_counts.values())
curr_kws = [w for w in kw_list if get_log_ratio(w, g_sub_counts, g_oth_counts) > 1]
win_counts[gname][window] = pd.Series({w: g_sub_counts[w] / curr_total if w in curr_kws else 0 for w in kw_list})
win_counts = {g: pd.DataFrame(win_counts[g]).T.fillna(0) for g in group_indexes}
return win_counts
# %%time
for mf in [50]:
g_kw_over_time = get_kw_per_window_same_kws(contributions=all_contributions, tokens=all_toks,
group_indexes={"Conservative": con.index, "Labour": lab.index},
subset=eu_mentions.index,
win_size=50000, win_step=10000, min_freq=mf)
plot_group_fluct(g_kw_over_time, {"Conservative": "blue", "Labour": "red"})
g_kw_over_time["Conservative"].applymap(lambda x: True if x > 0 else False)
g_kw_over_time["Conservative"].applymap(lambda x: True if x > 0 else False)
# ## Comparing Groups
plot_group_comparison(g_kw_over_time["Conservative"], g_kw_over_time["Conservative"])
plot_group_fluct({"Conservative": g_kw_over_time["Conservative"]}, {"Conservative": "blue"})
# +
df1 = g_kw_over_time["Conservative"].applymap(lambda x: True if x > 0 else False)
df2 = g_kw_over_time["Labour"].applymap(lambda x: True if x > 0 else False)
fluct = comp_anal(df1, df2, calc_ac1)
# -
fluct.plot()
fluct_anal(df1, calc_ac1).plot()
for d, currow in (df1 == df2).applymap(int).sum(axis=1).items():
print(d, currow, df1.shape[1], currow / df1.shape[1])
((df1 == df2).applymap(int).sum(axis=1) / df1.shape[1]).plot()
| 4.5 - Fluctuation Analysis/KFA for Chapter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda-clmm
# language: python
# name: conda-clmm
# ---
# # Fit halo mass to shear profile: 2. realistic data
#
# _the LSST-DESC CLMM team_
#
#
# This notebook demonstrates how to use `clmm` to estimate a WL halo mass from observations of a galaxy cluster. It uses several functionalities of the support `mock_data` module to produce datasets of increasing complexity. This notebook also demonstrates the bias introduced on the reconstructed mass by a naive fit, when the redshift distribution of the background galaxies is not properly accounted for in the model. Organization of this notebook goes as follows:
#
# - Setting things up, with the proper imports.
# - Generating 3 datasets: an ideal dataset (dataset1) similar to that of Example1 (single source plane); an ideal dataset but with source galaxies following the Chang et al. (2013) redshift distribution (dataset2); a noisy dataset where photoz errors and shape noise are also included (dataset3).
# - Computing the binned reduced tangential shear profile, for the 3 datasets, using logarithmic binning.
# - Setting up the "single source plane" model to be fitted to the 3 datasets. Only dataset1 has a single source plane, so we expect to see a bias in the reconstructed mass when using this model on datasets 2 and 3.
# - Perform a simple fit using `scipy.optimize.curve_fit` and visualize the results.
# ## Setup
# First, we import some standard packages.
# +
import sys
sys.path.append('./support')
try: import clmm
except:
import notebook_install
notebook_install.install_clmm_pipeline(upgrade=False)
import clmm
import matplotlib.pyplot as plt
import numpy as np
from numpy import random
from sampler import fitters
clmm.__version__
# -
# Next, we import `clmm`'s core modules.
import clmm.dataops as da
import clmm.galaxycluster as gc
import clmm.modeling as modeling
from clmm import Cosmology
# We then import a support modules for a specific data sets.
# `clmm` includes support modules that enable the user to generate mock data in a format compatible with `clmm`.
import mock_data as mock
# ## Making mock data
# For reproducibility:
np.random.seed(11)
# To create mock data, we need to define a true cosmology.
mock_cosmo = Cosmology(H0 = 70.0, Omega_dm0 = 0.27 - 0.045, Omega_b0 = 0.045, Omega_k0 = 0.0)
# We now set some parameters for a mock galaxy cluster.
cosmo = mock_cosmo
cluster_m = 1.e15 # M200,m [Msun]
cluster_z = 0.3
concentration = 4
ngals = 10000
Delta = 200
cluster_ra = 0.0
cluster_dec = 0.0
# Then we use the `mock_data` support module to generate 3 galaxy catalogs:
# - `ideal_data`: all background galaxies at the same redshift.
# - `ideal_data_z`: galaxies distributed according to the Chang et al. (2013) redshift distribution.
# - `noisy_data_z`: `ideal_data_z` + photoz errors + shape noise
ideal_data = mock.generate_galaxy_catalog(cluster_m, cluster_z, concentration, cosmo, 0.8, ngals=ngals)
ideal_data_z = mock.generate_galaxy_catalog(cluster_m, cluster_z, concentration, cosmo,'chang13', ngals=ngals)
noisy_data_z = mock.generate_galaxy_catalog(cluster_m, cluster_z, concentration, cosmo, 'chang13',
shapenoise=0.05,
photoz_sigma_unscaled=0.05, ngals=ngals)
# The galaxy catalogs are converted to a `clmm.GalaxyCluster` object and may be saved for later use.
# +
cluster_id = "CL_ideal"
gc_object = clmm.GalaxyCluster(cluster_id, cluster_ra, cluster_dec,
cluster_z, ideal_data)
gc_object.save('ideal_GC.pkl')
cluster_id = "CL_ideal_z"
gc_object = clmm.GalaxyCluster(cluster_id, cluster_ra, cluster_dec,
cluster_z, ideal_data_z)
gc_object.save('ideal_GC_z.pkl')
cluster_id = "CL_noisy_z"
gc_object = clmm.GalaxyCluster(cluster_id, cluster_ra, cluster_dec,
cluster_z, noisy_data_z)
gc_object.save('noisy_GC_z.pkl')
# -
# Any saved `clmm.GalaxyCluster` object may be read in for analysis.
# +
cl1 = clmm.GalaxyCluster.load('ideal_GC.pkl') # all background galaxies at the same redshift
cl2 = clmm.GalaxyCluster.load('ideal_GC_z.pkl') # background galaxies distributed according to Chang et al. (2013)
cl3 = clmm.GalaxyCluster.load('noisy_GC_z.pkl') # same as cl2 but with photoz error and shape noise
print("Cluster info = ID:", cl2.unique_id, "; ra:", cl2.ra, "; dec:", cl2.dec, "; z_l :", cl2.z)
print("The number of source galaxies is :", len(cl2.galcat))
# -
h = plt.hist(cl2.galcat['z'], bins=50)
# ## Deriving observables
# ### Computing shear
# `clmm.dataops.compute_tangential_and_cross_components` calculates the tangential and cross shears for each source galaxy in the cluster.
theta1, g_t1, g_x1 = cl1.compute_tangential_and_cross_components(geometry="flat")
theta2, g_t2, g_x2 = cl2.compute_tangential_and_cross_components(geometry="flat")
theta2, g_t3, g_x3 = cl3.compute_tangential_and_cross_components(geometry="flat")
# ### Radially binning the data
bin_edges = da.make_bins(0.7, 4, 15, method='evenlog10width')
# `clmm.dataops.make_radial_profile` evaluates the average shear of the galaxy catalog in bins of radius.
profile1 = cl1.make_radial_profile("Mpc", bins=bin_edges,cosmo=cosmo)
profile2 = cl2.make_radial_profile("Mpc", bins=bin_edges,cosmo=cosmo)
profile3 = cl3.make_radial_profile("Mpc", bins=bin_edges,cosmo=cosmo)
# After running `clmm.dataops.make_radial_profile` on a `clmm.GalaxyCluster` object, the object acquires the `clmm.GalaxyCluster.profile` attribute.
for n in cl1.profile.colnames: cl1.profile[n].format = "%6.3e"
cl1.profile.pprint(max_width=-1)
# We visualize the radially binned shear for the 3 configurations
# +
fig = plt.figure(figsize=(10, 6))
fsize = 14
fig.gca().errorbar(profile1['radius'], profile1['gt'], yerr=profile1['gt_err'], marker='o', label='z_src = 0.8')
fig.gca().errorbar(profile2['radius'], profile2['gt'], yerr=profile2['gt_err'], marker='o',
label='z_src = Chang et al. (2013)')
fig.gca().errorbar(profile3['radius'], profile3['gt'], yerr=profile3['gt_err'], marker='o',
label='z_src = Chang et al. (2013) + photoz err + shape noise')
plt.gca().set_title(r'Binned shear of source galaxies', fontsize=fsize)
plt.gca().set_xlabel(r'$r\;[Mpc]$', fontsize=fsize)
plt.gca().set_ylabel(r'$g_t$', fontsize=fsize)
plt.legend()
# -
# ## Create the halo model
#
# `clmm.modeling.predict_reduced_tangential_shear` supports various parametric halo profile functions, including `nfw`.
# Beware that the `clmm.modeling` module works in units of $Mpc/h$, whereas the data is cosmology-independent, with units of $Mpc$.
# model definition to be used with scipy.optimize.curve_fit
def shear_profile_model(r, logm, z_src):
m = 10.**logm
gt_model = clmm.predict_reduced_tangential_shear(r,
m, concentration,
cluster_z, z_src, cosmo,
delta_mdef=200,
halo_profile_model='nfw')
return gt_model
# ### Fitting a halo mass - highlighting bias when not accounting for the source redshift distribution in the model
# We estimate the best-fit mass using `scipy.optimize.curve_fit`.
#
# Here, to build the model we make the WRONG assumption that the average shear in bin $i$ equals the shear at the average redshift in the bin; i.e. we assume that $\langle g_t\rangle_i = g_t(\langle z\rangle_i)$. This should not impact `cluster 1` as all sources are located at the same redshift. However, this yields a bias in the reconstructed mass for `cluster 2` and `cluster 3`, where the sources followed the Chang et al. (2013) distribution.
# +
# Cluster 1: ideal data
popt1,pcov1 = fitters['curve_fit'](lambda r, logm:shear_profile_model(r, logm, profile1['z']),
profile1['radius'],
profile1['gt'],
profile1['gt_err'], bounds=[13.,17.])
#popt1,pcov1 = spo.curve_fit(lambda r, logm:shear_profile_model(r, logm, profile1['z']),
# profile1['radius'],
# profile1['gt'],
# sigma=profile1['gt_err'], bounds=[13.,17.])
m_est1 = 10.**popt1[0]
m_est_err1 = m_est1 * np.sqrt(pcov1[0][0]) * np.log(10) # convert the error on logm to error on m
# Cluster 2: ideal data with redshift distribution
popt2,pcov2 = fitters['curve_fit'](lambda r, logm:shear_profile_model(r, logm, profile2['z']),
profile2['radius'],
profile2['gt'],
profile2['gt_err'], bounds=[13.,17.])
m_est2 = 10.**popt2[0]
m_est_err2 = m_est2 * np.sqrt(pcov2[0][0]) * np.log(10) # convert the error on logm to error on m
# Cluster 3: noisy data with redshift distribution
popt3,pcov3 = fitters['curve_fit'](lambda r, logm:shear_profile_model(r, logm, profile3['z']),
profile3['radius'],
profile3['gt'],
profile3['gt_err'], bounds=[13.,17.])
m_est3 = 10.**popt3[0]
m_est_err3 = m_est3 * np.sqrt(pcov3[0][0]) * np.log(10) # convert the error on logm to error on m
print(f'Best fit mass for cluster 1 = {m_est1:.2e} +/- {m_est_err1:.2e} Msun')
print(f'Best fit mass for cluster 2 = {m_est2:.2e} +/- {m_est_err2:.2e} Msun')
print(f'Best fit mass for cluster 3 = {m_est3:.2e} +/- {m_est_err3:.2e} Msun')
# -
# As expected, the reconstructed mass is biased whenever the sources are not located at a single redshift as this was not accounted for in the model.
# ## Visualization of the results
#
# For visualization purpose, we calculate the reduced tangential shear predicted by the model when using the average redshift of the catalog.
# +
rr = np.logspace(-0.5, np.log10(5), 100)
gt_model1 = clmm.predict_reduced_tangential_shear(rr,
m_est1, concentration,
cluster_z, np.mean(cl1.galcat['z']), cosmo,
delta_mdef=200,
halo_profile_model='nfw')
gt_model2 = clmm.predict_reduced_tangential_shear(rr,
m_est2, concentration,
cluster_z, np.mean(cl2.galcat['z']), cosmo,
delta_mdef=200,
halo_profile_model='nfw')
gt_model3 = clmm.predict_reduced_tangential_shear(rr,
m_est3, concentration,
cluster_z, np.mean(cl3.galcat['z']), cosmo,
delta_mdef=200,
halo_profile_model='nfw')
# -
# We visualize that prediction of reduced tangential shear along with the data
# +
fig, axes = plt.subplots(nrows=1, ncols=2, figsize=(16, 6))
axes[0].errorbar(profile1['radius'], profile1['gt'],profile1['gt_err'], color='red',
label='ideal_data, M_input = %.3e Msun' % cluster_m, fmt='.')
axes[0].plot(rr, gt_model1,color='red',
label='best fit model 1, M_fit = %.2e +/- %.2e' % (m_est1, m_est_err1))
axes[0].errorbar(profile2['radius'], profile2['gt'],profile2['gt_err'], color='green',
label='ideal_data_z, M_input = %.3e Msun' % cluster_m, fmt='.')
axes[0].plot(rr, gt_model2, color='green',
label='best fit model 2, M_fit = %.2e +/- %.2e' % (m_est2, m_est_err2))
axes[0].set_title('Ideal data w/wo src redshift distribution',fontsize=fsize)
axes[0].semilogx()
axes[0].semilogy()
axes[0].legend(fontsize=fsize)
axes[0].set_xlabel('R [Mpc]', fontsize=fsize)
axes[0].set_ylabel('reduced tangential shear', fontsize=fsize)
axes[1].errorbar(profile3['radius'], profile3['gt'],profile3['gt_err'], color='red',
label='noisy_data_z, M_input = %.3e Msun' % cluster_m, fmt='.')
axes[1].plot(rr, gt_model3,color='red',
label='best fit model 3, M_fit = %.2e +/- %.2e' % (m_est3, m_est_err3))
axes[1].set_title('Noisy data with src redshift distribution',fontsize=fsize)
axes[1].semilogx()
axes[1].semilogy()
axes[1].legend(fontsize=fsize)
axes[1].set_xlabel('R [Mpc]', fontsize=fsize)
axes[1].set_ylabel('reduced tangential shear', fontsize=fsize)
fig.tight_layout()
| examples/Example2_Fit_Halo_Mass_to_Shear_Catalog.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="s-jEzGA-yRqe" colab_type="code" outputId="1caea6f4-16b1-4892-d181-bfcb9fba8d3e" colab={"base_uri": "https://localhost:8080/", "height": 136}
# !git clone https://github.com/abhishek-choudharys/PokemonGAN
# + id="TU1HY5gCycqQ" colab_type="code" outputId="d7c5291b-455c-4a31-d966-031915ccd154" colab={"base_uri": "https://localhost:8080/", "height": 34}
# cd ../content/PokemonGAN
# + id="FS-RKZ0KygXp" colab_type="code" colab={}
run resize.py
# + id="SJyoBZSoyusv" colab_type="code" colab={}
run convertRGBAtoRGB.py
# + id="EGFhJ-uIy3Uh" colab_type="code" outputId="44dce53f-4532-4ba3-d028-04cd67d78e2c" colab={"base_uri": "https://localhost:8080/", "height": 34}
# %matplotlib inline
import tensorflow as tf
from tensorflow import keras
import numpy as np
#import plot_utils
import matplotlib.pyplot as plt
from tqdm import tqdm
from IPython import display
print('Tensorflow version: ', tf.__version__)
# + id="7LHhhAHly7hA" colab_type="code" outputId="bd6f946e-2345-4cd2-baf6-69ba2a8af6d9" colab={"base_uri": "https://localhost:8080/", "height": 34}
import cv2
import os
def load_images_from_folder(folder):
images = []
for filename in os.listdir(folder):
img = cv2.imread(os.path.join(folder,filename))
img_rgb = img[:, :, [2, 1, 0]]
if img_rgb is not None:
images.append(img_rgb)
return images
images = load_images_from_folder('resized_RGB')
type(images)
# + id="RmqToCwqy-9K" colab_type="code" outputId="7c348aec-5033-4c2c-f425-4f856aa23bd4" colab={"base_uri": "https://localhost:8080/", "height": 51}
import random
def grab_batch(images, batch_size):
image_batch = np.zeros([32, 256, 256, 3])
#image_batchx = np.empty()
#print(np.asarray(random.sample(images, 1)).shape)
for i in range(0, batch_size):
image_batch[i,:,:,:] = np.asarray(random.sample(images, 1))[:,:,:]
#image_batchx = np.append(image_batchx, np.asarray(random.sample(images, 1))[:,:,:])
return image_batch*(1./255)
image_batch = np.asarray(grab_batch(images, 32))
print(type(image_batch))
print(image_batch.shape)
# + id="RFYnyZdGzBdM" colab_type="code" colab={}
def show_batch(image_batch):
print(type(image_batch))
plt.figure(figsize=(10,10))
for n in range(25):
ax = plt.subplot(5,5,n+1)
plt.imshow(image_batch[n])
plt.axis('off')
# + id="mHvrd30CzfvW" colab_type="code" colab={}
def generate_and_save_images(model, epoch, test_input):
predictions = model(test_input, training = False)
fig = plt.figure(figsize = (10,10))
for i in range(25):
plt.subplot(5, 5, i+1)
plt.imshow(predictions[i, :, :, :] * 127.5 + 127.5)
plt.axis('off')
plt.savefig('image_at_each_epoch_{:04d}.png'.format(epoch))
plt.show()
# + id="Si8Qg099zKrJ" colab_type="code" outputId="11249ba0-e311-4bcf-b51c-c120abe3bdbe" colab={"base_uri": "https://localhost:8080/", "height": 592}
show_batch(image_batch) #from custom grab
# + id="6gBjbjGjzQHR" colab_type="code" outputId="41e572ed-c696-420b-9935-ec49b8fc4ee8" colab={"base_uri": "https://localhost:8080/", "height": 1000}
num_features = 10000
generator = keras.models.Sequential([
keras.layers.Dense(8*8*256, input_shape = [num_features]), #dense layer
keras.layers.Reshape([8,8,256]), #reshaped to 7*7 of 128
keras.layers.BatchNormalization(),
keras.layers.Conv2DTranspose(128, (5,5), (2,2), padding='same', activation='selu'),
keras.layers.BatchNormalization(),
keras.layers.Conv2DTranspose(64, (5,5), (2,2), padding='same', activation='selu'),
keras.layers.BatchNormalization(),
keras.layers.Conv2DTranspose(32, (5,5), (2,2), padding='same', activation='selu'),
keras.layers.BatchNormalization(),
keras.layers.Conv2DTranspose(16, (5,5), (2,2), padding='same', activation='selu'),
keras.layers.BatchNormalization(),
keras.layers.Conv2DTranspose(3, (5,5), (2,2), padding='same', activation='tanh'),
])
generator.summary()
noise = tf.random.normal(shape = [1, num_features])
generated_image = generator(noise, training = False)
#show(generated_image, 1)
discriminator = keras.models.Sequential([
keras.layers.Conv2D(64, (5,5), (2,2), padding='same', input_shape = [256, 256, 3]),
keras.layers.LeakyReLU(0.2),
keras.layers.Dropout(0.3),
keras.layers.Conv2D(128, (5,5), (2,2), padding='same'),
keras.layers.LeakyReLU(0.2),
keras.layers.Dropout(0.3),
keras.layers.Conv2D(256, (5,5), (2,2), padding='same'),
keras.layers.LeakyReLU(0.2),
keras.layers.Dropout(0.3),
keras.layers.Flatten(),
keras.layers.Dense(1, activation = 'sigmoid'),
])
discriminator.summary()
discriminator_output = discriminator(generated_image, training=False)
print(discriminator_output)
discriminator.compile(loss = 'binary_crossentropy', optimizer = 'rmsprop')
discriminator.trainable = False
gan = keras.models.Sequential([generator, discriminator])
gan.compile(loss = 'binary_crossentropy', optimizer = 'rmsprop')
# + id="pFH5pr5bzT2k" colab_type="code" colab={}
batch_size = 32
seed = tf.random.normal(shape = [batch_size, num_features])
# + id="HvbnJuWJ93_K" colab_type="code" colab={}
def show(images, n_cols = None):
n_cols = n_cols or len(images)
n_rows = (len(images) - 1) // n_cols + 1
if(images.shape[-1] == 1):
images = np.squeeze(images, axis=-1)
plt.figure(figsize = (n_cols, n_rows))
for index, image in enumerate(images):
plt.subplot(n_cols, n_rows, index+1)
plt.imshow(image)
plt.axis("off")
# + id="MtdJRIO56ssc" colab_type="code" outputId="2131f0ea-2a47-440a-bd41-b2052b603108" colab={"base_uri": "https://localhost:8080/", "height": 170}
noise = tf.random.normal(shape = [1, num_features])
print("Noise : ", noise[0,:10])
generated_image = generator(noise, training = False)
show(generated_image, 1)
decision = discriminator(generated_image)
print("Decision: ", decision)
# + id="uAMTte5QzcVx" colab_type="code" colab={}
def train_dcgan2(gan, batch_size, num_features, epochs = 5):
generator, discriminator = gan.layers
for epoch in tqdm(range(epochs)):
print("Epoch : {}/{}".format(epoch+1, epochs))
for i in range(0,10):
X_batch = grab_batch(images, batch_size)
#show_batch(X_batch)
noise = tf.random.normal(shape = [batch_size, num_features])
generated_images = generator(noise)
X_fake_and_real = tf.concat([generated_images, X_batch], axis = 0)
show_batch(X_fake_and_real)
y1 = tf.constant([[0.]] * batch_size + [[1.]] * batch_size)
discriminator.trainable = True
discriminator.train_on_batch(X_fake_and_real, y1)
y2 = tf.constant([[1.]] * batch_size)
discriminator.trainable = False
gan.train_on_batch(noise, y2)
display.clear_output(wait = True)
generate_and_save_images(generator, epoch+1, seed)
generate_and_save_images(generator, epochs, seed)
# + id="tdwIvLjNzr1b" colab_type="code" colab={}
train_dcgan2(gan, batch_size, num_features, epochs=10)
| GAN/PokemonGAN_3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Métodos de ensamble aplicados a clasificación
#
#
# Los métodos de esnamble sirven para probar muchos estimadores, es decir, probamos diferentes algorimtos y luego le decimos que lleguen en común a una conclusión.
#
# Combinar diferentes métodos de ML con diferentes configuraciones y aplicar un método para logra un consenso.
#
# La diversidad es una muy buena opción
# Los métods de ensamble se han destacado por ganar muchas competencias de ML.
#
#
# ### Baggin:
#
# Viene de Boottrap Aggregation
#
# Consiste en consultar la opinion de varios expertos, el algotritmo lo que hace es que permtie que cada uno de los expertos decida en paralelo. Y atraves de una formula al final filtra el mejor.
#
# 1. Se parten los datos
# 2. Se entrenan los modelos
# 3. Se combinan por ejemplo las respeustas
#
#
# Algunos métodos reconocidos de Baggin son:
#
# 1. Random Forest
#
# 2. Voting Classifiers/Regressors
#
# 3. En general se peuden aplciar sobre cualquer modelo de mahcine learnign
#
#
# ### Boosting
#
# Boosting Viene de impulsar
# Trabaja no en paraleo si no en serie, le pedimos a un experto el criterio sobre un problema. Medimos su posible errir, y luego usando ese error calculado le pedimos a otr experto su juiocio sobre el mismo problema. Es como pasar los datos por diferentes filtros. E experto corrige basados en un error. Todo
# hasta lelgar a un consejo.
#
# Busca fortalecer gradualmente un modelo de aprendizaje usando siempre el error residual de las etapas anteriores.
#
# El resultado final también se consigue por consenso entre tods los modelos.
#
# Los modelos mas conocidos de ensamblado por boosting
#
# 1.AdaBoost
#
# 2.Gradiente Tree Boosting
#
# 3. XGBoost
#
#
#
#
# ## Preparación de datos para implementar métodos de ensamble
#
#
# El metaestimado Bugguin clasifiers significa que lo puedo adaptar para diferentes familais de estimadores y automaticamente se convierte en un método de ensamble.
#
# Haremos este ejerccio con el data set de afecciones cardiacas.
# +
import pandas as pd
from sklearn.neighbors import KNeighborsClassifier
from sklearn.ensemble import BaggingClassifier
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score
# -
dt_heart = pd.read_csv('./data/heart.csv')
dt_heart
dt_heart['target'].describe()
# Cargamso en los features todos los datasets excepto la variable target.
X = dt_heart.drop(['target'], axis =1)
y = dt_heart['target']
# Entreno mi modelo
X_train, X_test, y_train, y_test = train_test_split(X,y, test_size =0.35)
# ## Implementación de Bagging
# +
knn_class = KNeighborsClassifier().fit(X_train, y_train) # Ajustamos
knn_pred = knn_class.predict(X_test) # Predecimos
accuracy_only_clasifier = accuracy_score(knn_pred, y_test) #Comparamos contra lsodatos reales
print("This is the accuracy with Knn: "+ str(accuracy_only_clasifier) )
# -
# Ahora queremos comparar este clasificador con la implmentación del clasificador por ensamble
'''
Primer argumento:
** En qué está basado nuestro pimer estimaodor base_estimator
** Y un numero de estimadores
'''
bag_class = BaggingClassifier(base_estimator=KNeighborsClassifier(), n_estimators=50).fit(X_train, y_train)
bag_predictions = bag_class.predict(X_test)
accuracy_bag_ensamble = accuracy_score(bag_predictions, y_test)
print("This is the accuracy with Assemble: " + str(accuracy_bag_ensamble) )
# Con el método por ensamble se optiene un valro superior a l del clasificador solo. Tener meayor grado de confianza supone la vida de un paciente y al desición respecto al diagnostico.
#
# ¡Como interpreto este numero?Assemble: 0.7409470752089137 Que con un 74,% de confianza estamso predicioendo que un paciente tenga probelams cariacos o no. ¿?
# ## Implementación de Boosting
#
#
# Usaremos el gradientr tree boosting. Usaremos neuvamente target como la variable que uqeremso clasificar, e importaremos nuestros arboles de deicicon.
from sklearn.ensemble import GradientBoostingClassifier
# +
# Defnimos el clasificadors
'''
Los estimadores aqui son el numeor de arboles de desicion.
Se peude ajsutar según criterio o ealgo llamado Crosx Validation... que es?
Construye arboles pequeños profundo con pocas hojas, entrena uno de tras de otro para un resultado optimio de la sclasificacion.
'''
boost = GradientBoostingClassifier(n_estimators=50).fit(X_train, y_train)
boost_predictions = boost.predict(X_test)
accuracy_boost = accuracy_score(boost_predictions, y_test)
print("This is the accuracy with Boosting: " + str(accuracy_boost) )
# -
# La mejora es significativa entre los métodos baggin y boosting, es decir entre procesameinto en paralelo y en serie. Estos son los métodos de ensamble.
| NoteBooks/Curso de Scikit-learn/Jupyter Notes/Métodos de ensamble aplicados a clasificación.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Exploring the Dataframe
# Exploring the indexes (rows) of a certain dataframe
# Method 1:
list(dfObj.index.values)
# Method 2:
list(dfObj.index)
# Good!, now we can inspect this new dataframe:
# returns a tuple with number of rows/columns
DF.shape
# In order to have basic information the DataFrame:
DF.info()
# In order to have a more detailed report on the memory usage you do:
DF.info(memory_usage='deep')
# And we can also take a look to the first rows of the dataframe:
DF.head(3) #only the 3 first lines are shown
# In order to know whe column names:
DF.columns
# If we want to check a particular column from the dataframe ('RSI' for example):
RSI=DF[['RSI']]
# If we want to select 2 non consecutive columns:
a=DF[['RSI','Ranging']]
# ### Selecting using .iloc and .loc
# Extracted from:
# https://www.shanelynn.ie/select-pandas-dataframe-rows-and-columns-using-iloc-loc-and-ix/
#
# #### .iloc<br>
# Single selection:<br>
# * Rows:<br>
# data.iloc[0] # first row of data frame (<NAME>) - Note a Series data type output.<br>
# data.iloc[1] # second row of data frame (<NAME>)<br>
# data.iloc[-1] # last row of data frame (<NAME>)<br>
# * Columns:<br>
# data.iloc[:,0] # first column of data frame (first_name)<br>
# data.iloc[:,1] # second column of data frame (last_name)<br>
# data.iloc[:,-1] # last column of data frame (id)<br>
#
# Multiple selection:<br>
# <br>
# data.iloc[0:5] # first five rows of dataframe<br>
# data.iloc[:, 0:2] # first two columns of data frame with all rows<br>
# data.iloc[[0,3,6,24], [0,5,6]] # 1st, 4th, 7th, 25th row + 1st 6th 7th columns.<br>
# data.iloc[0:5, 5:8] # first 5 rows and 5th, 6th, 7th columns of data frame (county -> phone1)<br>
# data.iloc[:, [0,1]] <br>
#
# #### .loc<br>
# Single selection:<br>
# a=DF.loc[:,'Direction']<br>
#
# Multiple selection:<br>
# a=DF.loc[:,['Direction','RSI']]
# ### Objects returned by .iloc and .loc
#
# * If only one row is selected then we will get a Pandas series:<br>
# data.iloc[0]
# * If we use list selector then we get a Dataframe:<br>
# data.iloc[[0]]
# * If we select multiple rows then we get a Dataframe:<br>
# data.iloc[0:5]
# +
ix='RSI'
DF.loc[:,ix]
# -
# ### Setting the value of a certain cell in the dataframe
# #### By index:
# +
import pandas as pd
df=pd.DataFrame(index=['A','B','C'], columns=['x','y'])
df.at['C', 'x'] = 10
# -
# #### By position:
df = pd.DataFrame([[0, 2, 3], [0, 4, 1], [10, 20, 30]],columns=['A', 'B', 'C'])
df.iat[1, 2] = 10
# ### Logical selection
# And for example, if we want to select all records for which the 'Reversed' column is TRUE:
reversed_true=DF.loc[DF['Reversed']==True]
# And if we want to select based in either the value of one column or a different one:
DF.loc[(DF['Reversed']==True) | DF['Divergence']==True]
# Now, if we want the counts (frequencies) for a certain categorical variable we have to enter the following:
DF['Currency Pair'].value_counts()
# And if we want to have proportions instead of counts we do:
DF['Currency Pair'].value_counts(normalize=True)
# And if we want we have percentages we do:
DF['Currency Pair'].value_counts(normalize=True)*100
# Now, if we want to copy the entire dataframe:
# +
newDF = DF.copy()
newDF.head(3)
| PANDAS/DATAFRAMES/.ipynb_checkpoints/Exploring a dataframe-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: qawork
# language: python
# name: qawork
# ---
# +
from easyquant import MongoIo
from easyquant.indicator.base import *
import easyquotation
import math
import numpy as np
import pandas as pd
from easyquant import MongoIo
# from easyquant.qafetch import QATdx as tdx
from easyquant.indicator.base import *
import pandas as pd
from easyquant import EasyTime
from datetime import datetime, time
from ctypes import *
# import datetime
import math
import numpy as np
import pandas as pd
import talib
from ctypes import *
import QUANTAXIS as QA
# import time
# -
m=MongoIo()
source=easyquotation.use('sina')
table = 'st_orders-%s' % 'tdx_yhzc'
# +
def new_df(df_day, data, now_price):
code = data['code']
# now_vol = data['volume']
last_time = pd.to_datetime(data['datetime'][0:10])
# print("code=%s, data=%s" % (self.code, self._data['datetime']))
# df_day = data_buf_day[code]
# df_day.loc[last_time]=[0 for x in range(len(df_day.columns))]
df_day.at[(last_time, code), 'open'] = data['open']
df_day.at[(last_time, code), 'high'] = data['high']
df_day.at[(last_time, code), 'low'] = data['low']
df_day.at[(last_time, code), 'close'] = now_price
df_day.at[(last_time, code), 'volume'] = data['volume']
df_day.at[(last_time, code), 'amount'] = data['amount']
return df_day
# -
def tdx_dqe_cfc_A1(data, sort=False):
# 选择/排序
C = data.close
O = data.open
JC =IF(ISLASTBAR(C), O, C)
MC = (0.3609454219 * JC - 0.03309329629 * REF(C, 1) - 0.04241822779 * REF(C, 2) - 0.026737249 * REF(C, 3) \
- 0.007010041271 * REF(C, 4) - 0.002652859952 * REF(C, 5) - 0.0008415042966 * REF(C, 6) \
- 0.0002891931964 * REF(C, 7) - 0.0000956265934 * REF(C, 8) - 0.0000321286052 * REF(C, 9) \
- 0.0000106773454 * REF(C, 10) - 0.0000035457562 * REF(C, 11) -- 0.0000011670713 * REF(C, 12)) / (1 - 0.7522406533)
# 竞价涨幅 := (DYNAINFO(4) / DYNAINFO(3) - 1) * 100;
竞价涨幅 = (C / REF(C, 1) - 1) * 100
# ST := STRFIND(stkname, 'ST', 1) > 0;
# S := STRFIND(stkname, 'S', 1) > 0;
# 停牌 := (DYNAINFO(4)=0);
#
# 附加条件 := (not (ST) and not (S) and NOT(停牌)) * (竞价涨幅 < 9.85) * (竞价涨幅 > (0));
附加条件 = IFAND(竞价涨幅 < 9.85, 竞价涨幅 > 0, 1, 0)
# if sort:
刀 = (MC - JC) / JC * 1000
# else:
# 刀 = (MC - JC) / JC * 1000 * 附加条件
return 刀
def tdx_dqe_cfc_A12(data, sort=False):
# 选择/排序
C = data.close
O = data.open
JC = IF(ISLASTBAR(C), O, C)
MC = (0.3609454219 * JC - 0.03309329629 * REF(C, 1) - 0.04241822779 * REF(C, 2) - 0.026737249 * REF(C, 3) \
- 0.007010041271 * REF(C, 4) - 0.002652859952 * REF(C, 5) - 0.0008415042966 * REF(C, 6) \
- 0.0002891931964 * REF(C, 7) - 0.0000956265934 * REF(C, 8) - 0.0000321286052 * REF(C, 9) \
- 0.0000106773454 * REF(C, 10) - 0.0000035457562 * REF(C, 11) -- 0.0000011670713 * REF(C, 12)) / (1 - 0.7522406533)
# 竞价涨幅 := (DYNAINFO(4) / DYNAINFO(3) - 1) * 100;
竞价涨幅 = (O / REF(C, 1) - 1) * 100
# ST := STRFIND(stkname, 'ST', 1) > 0;
# S := STRFIND(stkname, 'S', 1) > 0;
# 停牌 := (DYNAINFO(4)=0);
#
# 附加条件 := (not (ST) and not (S) and NOT(停牌)) * (竞价涨幅 < 9.85) * (竞价涨幅 > (0));
附加条件 = IFAND(竞价涨幅 < 9.85, 竞价涨幅 > 0, 1, 0)
# if sort:
刀 = (MC - JC) / JC * 1000
# else:
# 刀 = (MC - JC) / JC * 1000 * 附加条件
return 刀
# +
def tdx_yhzc(data):
# 用户注册
# pass
# C = data.close
CLOSE = data.close
# HIGH = data.high
# H = data.high
# L = data.low
# LOW = data.low
# OPEN = data.open
# O = data.open
VOL = data.volume
# AMOUNT = data.amount
# 除业绩后退股 := FINANCE(30) >= REF(FINANCE(30), 130);
# D0 := 除业绩后退股;
# D2 := IF(NAMELIKE('S'), 0, 1);
# D3 := IF(NAMELIKE('*'), 0, 1);
# D4 := DYNAINFO(17) > 0;
# 去除大盘股 := CAPITAL / 1000000 < 20;
# 去高价 := C <= 60;
# 去掉 := D0 and D2 and D3 and D4 and 去除大盘股 and 去高价 and NOT(C >= REF(C, 1) * 1.097 and C = O and H = L);
TJ_V = VOL > 3 * MA(VOL,89)
DIF1 = (EMA(CLOSE, 12) - EMA(CLOSE, 26)) / EMA(CLOSE, 26) * 100
DEA1 = EMA(DIF1, 9)
AAA1 = (DIF1 - DEA1) * 100
# MA120 = REF(MA(C,120),1)
# MA5 = REF(MA(C, 120),1)
# MA10 = REF(MA(C, 120),1)
# PTGD = REF(HHV(C,120),1)
# XIN_GAO = IFAND(C > PTGD, C > MA120, True, False)
用 = 45
户 = AAA1 - REF(AAA1, 1)
注册 = CROSS(户, 用)
DIF = (EMA(CLOSE, 10) - EMA(CLOSE, 72)) / EMA(CLOSE, 72) * 100
DEA = EMA(DIF, 17)
AAA = (DIF - DEA) * 100
用户 = CROSS(AAA - REF(AAA, 1), 45)
# 用户注册 = IFAND4(注册 , 用户, TJ_V, XIN_GAO, 1, 0) #and 去掉;
用户注册 = IFAND3(注册 , 用户, TJ_V, 1, 0) #and 去掉;
return 用户注册
# -
# today
code='300787'
data=m.get_stock_day(code)
newdata=source.stocks(code)[code]
now_price = newdata['open']
data = new_df(data.copy(), newdata, now_price)
# data.iat[-1,3]=newdata['open']
data.tail()
C = data.close
O = data.open
V = data.volume
H = data.high
L = data.low
TJ1=IFAND(O > REF(MA(C, 5), 1), O > REF(MA(C, 10), 1), True, False)
TJ2=(COUNT(REF(V, 1) / REF(V, 2) > 6, 10) == 0)
ccc=tdx_dqe_cfc_A1(data)
yhzc=tdx_yhzc(data)
data['A1']=ccc
data['yhzc']=yhzc
data['tj1']=IFAND(O > REF(MA(C, 5), 1), O > REF(MA(C, 10), 1), True, False)
data['tj2']=(COUNT(REF(V, 1) / REF(V, 2) > 6, 10) == 0)
# result
data.tail()
# +
# back:
code='002242'
data=m.get_stock_day(code, st_end='2020-11-05')
C = data.close
O = data.open
V = data.volume
H = data.high
L = data.low
# data.tail()
# -
data=data.copy()
ccc=tdx_dqe_cfc_A12(data).tail()
data['A1']=ccc
data['tj1']=IFAND(O > REF(MA(C, 5), 1), O > REF(MA(C, 10), 1), True, False)
data['tj2']=(COUNT(REF(V, 1) / REF(V, 2) > 6, 10) == 0)
# result
data.tail()
code='300612'
dataInfo = m.get_stock_info(code)
dataInfo.jinglirun[0]
300364 7.62 8.49 7.38 7.92 787380.0 621797440.0 48.325356
300464 20.50 22.88 20.08 22.30 249674.0 533748384.0 0.000000 True True
300689 48.51 53.56 47.11 49.70 51398.0 258483824.0 25.845675
dataInfo = mongo.get_stock_info(code)
| test/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: ElastixEnv
# language: python
# name: elastixenv
# ---
# ### Napari Viewer
# Napari can be used as a GUI for registration with elastix.
# This can only be run locally, since napari opens an external window.
import enum
import napari
from napari.layers import Image
from magicgui import magicgui
from qtpy.QtGui import QFont
from itk import itkElastixRegistrationMethodPython
import itk
from itkwidgets import compare, checkerboard
import numpy as np
import imageio
# +
# Load images with itk floats (itk.F). Necessary for elastix
fixed_image = itk.imread('../examples/data/CT_2D_head_fixed.mha', itk.F)
moving_image = itk.imread('../examples/data/CT_2D_head_moving.mha', itk.F)
# Cast images to numpy arrays for Napari Viewer
fixed_image_np = np.asarray(fixed_image).astype(np.float32)
moving_image_np = np.asarray(moving_image).astype(np.float32)
# -
# Create parameterObject function
def parameterObject(transform, resolution, optimizer, imageSampler):
parameter_object = itk.ParameterObject.New()
default_rigid_parameter_map = parameter_object.GetDefaultParameterMap(transform, resolution)
default_rigid_parameter_map['Optimizer'] = [optimizer]
default_rigid_parameter_map['ImageSampler'] = [imageSampler]
parameter_object.AddParameterMap(default_rigid_parameter_map)
return parameter_object
# Create Napari Gui plugin
with napari.gui_qt():
viewer = napari.Viewer()
viewer.add_image(fixed_image_np, name=f"Fixed")
viewer.add_image(moving_image_np, name=f"Moving")
@magicgui(call_button="register",
transform = {"choices": ["rigid", "affine", "bspline"]},
resolutions = {"resolutions": int, "minimum":1, "maximum":10},
optimizer = {"choices":["AdaptiveStochasticGradientDescent","ConjugateGradient","FiniteDifferenceGradientDescent","Powell","PreconditionedStochasticGradientDescent","QuasiNewtonLBFGS","RegularStepGradientDescent", "StandardGradientDescent"]},
imageSampler = {"choices":["Full", "Grid", "RandomCoordinate", "RandomSparseMask", "Random"]})
def registration(transform= "rigid", resolution = 2, optimizer = "AdaptiveStochasticGradientDescent", imageSampler="Random") -> Image:
parameter_object = parameterObject(transform, resolution, optimizer, imageSampler)
result_image, result_transform_parameters = itk.elastix_registration_method(
fixed_image_np, moving_image_np,
parameter_object=parameter_object,
log_to_console=True)
result_image_np = np.asarray(result_image).astype(np.float32)
viewer.add_image(result_image_np, name=f"Result_"+transform+"_"+imageSampler)
return result_image_np
gui = registration.Gui()
gui.setFont(QFont('Arial',15))
viewer.window.add_dock_widget(gui)
viewer.layers.events.changed.connect(lambda x: gui.refresh_choices())
| napari/Napari_Registration_App.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Lab 01 : MLP -- solution
#
# # Understanding the training loop
# For Google Colaboratory
import sys, os
if 'google.colab' in sys.modules:
from google.colab import drive
drive.mount('/content/gdrive')
file_name = 'mlp_solution.ipynb'
import subprocess
path_to_file = subprocess.check_output('find . -type f -name ' + str(file_name), shell=True).decode("utf-8")
print(path_to_file)
path_to_file = path_to_file.replace(file_name,"").replace('\n',"")
os.chdir(path_to_file)
# !pwd
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from random import randint
import utils
# ### Download the data and print the sizes
from utils import check_fashion_mnist_dataset_exists
data_path=check_fashion_mnist_dataset_exists()
# +
train_data=torch.load(data_path+'fashion-mnist/train_data.pt')
print(train_data.size())
# +
train_label=torch.load(data_path+'fashion-mnist/train_label.pt')
print(train_label.size())
# +
test_data=torch.load(data_path+'fashion-mnist/test_data.pt')
print(test_data.size())
# -
# ### Make a ONE layer net class. The network output are the scores! No softmax needed! You have only one line to write in the forward function
class one_layer_net(nn.Module):
def __init__(self, input_size, output_size):
super(one_layer_net , self).__init__()
self.linear_layer = nn.Linear( input_size, output_size , bias=False)
def forward(self, x):
scores = self.linear_layer(x)
return scores
# ### Build the net
net=one_layer_net(28*28,10)
print(net)
# ### Choose the criterion, optimizer. Set the batchize and learning rate to be:
# ### batchize = 50
# ### learning rate = 0.01
# +
criterion = nn.CrossEntropyLoss()
optimizer=torch.optim.SGD( net.parameters() , lr=0.01 )
bs=50
# -
# ### Complete the training loop
for iter in range(1,5000):
# Set dL/dU, dL/dV, dL/dW to be filled with zeros
optimizer.zero_grad()
# create a minibatch
indices=torch.LongTensor(bs).random_(0,60000)
minibatch_data = train_data[indices]
minibatch_label= train_label[indices]
#reshape the minibatch
inputs = minibatch_data.view(bs,28*28)
# tell Pytorch to start tracking all operations that will be done on "inputs"
inputs.requires_grad_()
# forward the minibatch through the net
scores=net( inputs )
# Compute the average of the losses of the data points in the minibatch
loss = criterion( scores , minibatch_label)
# backward pass to compute dL/dU, dL/dV and dL/dW
loss.backward()
# do one step of stochastic gradient descent: U=U-lr(dL/dU), V=V-lr(dL/dU), ...
optimizer.step()
# ### Choose image at random from the test set and see how good/bad are the predictions
# +
# choose a picture at random
idx=randint(0, 10000-1)
im=test_data[idx]
# diplay the picture
utils.show(im)
# feed it to the net and display the confidence scores
scores = net( im.view(1,784))
probs= F.softmax(scores, dim=1)
utils.show_prob_fashion_mnist(probs)
# -
| codes/labs_lecture05/lab01_mlp/mlp_solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Ten Sigma Event Extension
# Statistical Consequences of Fat Tails (Page 53): [Link to ebook PDF](https://researchers.one/articles/statistical-consequences-of-fat-tails-real-world-preasymptotics-epistemology-and-applications/5f52699d36a3e45f17ae7e36)
import numpy as np
import pandas as pd
from scipy.stats import kurtosis as scipy_kurtosis
from sympy.stats import P, E, variance, std, Die, Normal, StudentT
from sympy import Eq, simplify
Z = Normal('Z', 0, 1) # Declare a Normal random variable with mean 0, std 1
T = StudentT('T', 2)
# # Replicate Page 53
# We want to find P(Gaussian|Event):
# \begin{equation*}
# \frac{P(Gaussian)*P(Event|Gaussian)}{\Bigl(1-P(Gaussian)\Bigr)*P(Event|NonGaussian)+P(Gaussian)*P(Event|Gaussian)}
# \end{equation*}
# +
p_gaussian_list = [0.5, 0.999, 0.9999, 0.99999, 0.999999, 1] # P(Gaussian) values to check for
p_if_gauss = P(Z>10).evalf()
p_if_nongauss = P(T>10).evalf()
# -
1/p_if_gauss # Should be 1.31x10^23
1/p_if_nongauss # # Should be 203
# Evaluate the equation for each value in p_gaussian_list
p_gauss_if_event_list = []
for p_gauss in p_gaussian_list:
numerator = p_gauss * p_if_gauss
denominator = (1-p_gauss)*p_if_nongauss+p_gauss*p_if_gauss
p_gauss_if_event = numerator/denominator
p_gauss_if_event_list.append(p_gauss_if_event)
p_gaussian_list
p_gauss_if_event_list
# Create DataFrame
d = {'P(Gaussian)':p_gaussian_list, 'P(Gaussian|Event)':p_gauss_if_event_list}
page_53_table = pd.DataFrame(d)
page_53_table
# # Extension:
# What if you fit a new normal distribution after observing the 10 sigma event. I'm not saying it makes sense but let's see what happens.
# Let's suppose the event is from one day in 50 years. Then it's a 1 in 365*50 event.
n = 365*50 # Our dataset holds this many points before the 10 sigma event happens
n
# +
normal_array = np.random.normal(size=n)
df = pd.DataFrame(normal_array, columns = ['normal_sample'])
df.head()
# -
df.describe().T
scipy_kurtosis(df.normal_sample, fisher=False)
# So before the tail event:
# * std=1
# * kurtosis=3
# * max observation ~= 4
# ### Add tail event
new_df = df.append({'normal_sample':10},ignore_index=True)
new_df.describe().T
scipy_kurtosis(new_df.normal_sample, fisher=False)
# After adding the tail event:
# * std=1
# * kurtosis=3.5
# * max=10
# Rough conclusion, If we add a 10 sigma event to 50 years of daily gaussian data, ...
# * The standard deviation doesn't go up much
# * The kurtosis goes up from 3 to 3.5.
#
# Which leaves us with:
# * New dataset has mu and sigma of a standard normal, but its kurtosis gives it away as being non-gaussian
| notebooks/Notebook-05 - Sympy 10 Sigma Event Extension.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + pycharm={"is_executing": false}
# First, I import some helpful packages
import matplotlib as mpl
import matplotlib.pyplot as plt
import numpy as np
# Also, I set some font formatting for latex text
#plt.rc("text", usetex=True)
#plt.rc("font", **{"family":"serif", "size":11})
new_rc_params = {"text.usetex": False,
"svg.fonttype": 'none',
"savefig.dpi": 300,
"font.size": 12,
"legend.fontsize": 12,
#"figure.titlesize": 12,
"lines.linewidth": 1.0,
}
mpl.rcParams.update(new_rc_params)
# + pycharm={"is_executing": false, "name": "#%%\n"}
# Here, I define a few helper functions that convert spike counts to probabilities and vice verca
def count_to_prob(c, steps):
# deterministic transformation from number of spikes to probability of activity
return c / steps
def prob_to_count(p, steps):
# deterministic transformation from probability of activity to number of spikes
return np.round_(p*steps).astype(int)
# + pycharm={"is_executing": false, "name": "#%%\n"}
# Here, I define three different functions to generate spike trains from summarized statistics
def poisson_encode(p, steps):
# generate spike train from firing rate / spike density via Poisson encoding
spikes = []
for t in np.arange(1, steps + 1):
if p > np.random.rand():
spikes.append(t)
return spikes
def uniform_encode(f, steps):
# generate spike train from fixed spike number via uniform distribution across time
return np.random.choice(np.arange(1, steps + 1), f, replace=False)
def linear_encode(f, steps):
# generate spike train from fixed spike number via linear distribution
return np.round_(np.linspace(1, steps, f))
# + pycharm={"is_executing": false, "name": "#%%\n"}
# Now, lets define how arbitrary incoming spike trains are integrated over time
# based on deterministic spiking neuron internal dynamics
def integrate_spike_trains(steps, spikes_in, weights_in, threshold, tau, u_rest, u0):
# integrates a number of incoming spike trains as list-like objects of spike timepoints
assert len(spikes_in) == len(weights_in)
x = np.arange(0, steps + 1)
v = u0
v_t = []
spikes_out = []
for t in x:
v = v * tau
for i in range(len(spikes_in)):
if t in spikes_in[i]:
v += weights_in[i]
v_t.append(v)
if v >= threshold:
v = u_rest
spikes_out.append(t)
if v < -threshold:
v = -threshold
V_t = [0.0]
for v in v_t[1:]:
V_t.append(tau*V_t[-1]+v)
return v_t, V_t, spikes_out
# + pycharm={"is_executing": false, "name": "#%%\n"}
# Now I give a function to display the integration process visually
def plot_spike_integration(steps, spikes_in, weights_in, u_t, U_t, spikes_out, u_rest, threshold,
title="", show=True, close=True, file="", fig=None, label=""):
x = np.arange(0, steps + 1)
if fig==None:
fig, axs = plt.subplots(3, 1, figsize=(5,6), sharex=True, gridspec_kw={'height_ratios': [5, 5, 10]})
plt.subplots_adjust(wspace=0, hspace=0)
fig.patch.set_facecolor('xkcd:white')
fig.suptitle(title + r" $\Sigma_t \phi_{out} = $" + f" {len(spikes_out)}", fontsize=12)
axs[0].hlines(u_rest, min(x), max(x), colors="black", linestyles="--", label="$u_{rest}$", lw=1, alpha=0.5)
axs[0].hlines(threshold, min(x), max(x), colors="red", linestyles="--", label="threshold", lw=1, alpha=0.5)
axs[0].set_yticks([u_rest, threshold])
axs[0].set_yticklabels([r"$u_{rest}$", r"$\omega$"])
else:
axs = fig.axes
axs[0].plot(x,u_t, label="$u(t)$" + label)
axs[1].plot(x,U_t, label="$U(t)$" + label)
sign_colors = ["g" if w > 0.0 else "r" for w in weights_in] + ["b"]
axs[2].eventplot(spikes_in + [spikes_out], colors=sign_colors, linelengths=0.8)
#axs[0].legend()
#axs[0].set_ylabel("membrane potential $u$")
axs[2].set_xlabel("$t$")
axs[2].set_yticks(np.arange(0, len(spikes_in)+1))
#ylabels = [r"$\phi^{+}$" if w > 0.0 else r"$\phi^{-}$" for w in weights_in] + [r"$\phi_{out}$"]
ylabels = ["" for w in weights_in] + [r"$\phi_{out}$"]
axs[2].set_yticklabels(ylabels)
axs[0].xaxis.set_tick_params(length=0)
axs[1].xaxis.set_tick_params(length=0)
axs[0].spines["top"].set_visible(False)
axs[0].spines["right"].set_visible(False)
axs[0].spines["bottom"].set_visible(False)
axs[1].spines["top"].set_visible(False)
axs[1].spines["right"].set_visible(False)
axs[1].spines["bottom"].set_visible(False)
axs[2].spines["top"].set_visible(False)
axs[2].spines["right"].set_visible(False)
axs[2].spines["left"].set_visible(False)
plt.tight_layout()
if file:
plt.savefig(file, dpi=300)
if show:
plt.show()
if close:
plt.close()
return fig
# -
def plot_multicell_spike_integration(steps, spikes_in, weights_ins, u_ts, U_ts, spikes_outs, u_rest, threshold,
title="", show=True, close=True, file="", labels=None):
x = np.arange(0, steps + 1)
fig, axs = plt.subplots(3, 1, figsize=(4,6), sharex=True, gridspec_kw={'height_ratios': [5, 5, 10]})
plt.subplots_adjust(wspace=0, hspace=0)
fig.patch.set_facecolor('xkcd:white')
fig.suptitle(title, fontsize=12)
axs[0].hlines(u_rest, min(x), max(x), colors="black", linestyles="--", label="$u_{rest}$", lw=1, alpha=0.5)
axs[0].hlines(threshold, min(x), max(x), colors="red", linestyles="--", label="threshold", lw=1, alpha=0.5)
axs[0].set_yticks([u_rest, threshold])
axs[0].set_yticklabels([r"$u_{rest}$", r"$\omega$"])
for i in range(len(u_ts)):
if labels:
label = labels[i]
else:
label = f"neuron {i+1}"
axs[0].plot(x,u_ts[i], label="$u(t)$" + label)
axs[1].plot(x,U_ts[i], label="$U(t)$" + label)
sign_colors = ["g" for w in weights_ins[0]] + ["b"] * len(u_ts)
axs[2].eventplot(spikes_in + spikes_outs, colors=sign_colors, linelengths=0.8)
#axs[0].legend()
#axs[0].set_ylabel("membrane potential $u$")
axs[2].set_xlabel("$t$")
axs[2].set_yticks(np.arange(0, len(spikes_in)+1))
#ylabels = [r"$\phi^{+}$" if w > 0.0 else r"$\phi^{-}$" for w in weights_in] + [r"$\phi_{out}$"]
ylabels = ["" for w in weights_ins[0]] + [(r" $\Sigma_t \phi_{out} = $" + f" {len(spikes_outs[i])}") for i in range(len(u_ts))]
print(ylabels)
axs[2].set_yticklabels(ylabels)
axs[0].xaxis.set_tick_params(length=0)
axs[1].xaxis.set_tick_params(length=0)
axs[0].spines["top"].set_visible(False)
axs[0].spines["right"].set_visible(False)
axs[0].spines["bottom"].set_visible(False)
axs[1].spines["top"].set_visible(False)
axs[1].spines["right"].set_visible(False)
axs[1].spines["bottom"].set_visible(False)
axs[2].spines["top"].set_visible(False)
axs[2].spines["right"].set_visible(False)
axs[2].spines["left"].set_visible(False)
plt.tight_layout()
if file:
plt.savefig(file, dpi=300)
if show:
plt.show()
if close:
plt.close()
return fig
# + pycharm={"is_executing": false, "name": "#%%\n"}
# A function that chooses which of the above defined functions to use based on parameter choices
def run_simulation():
for i in np.arange(repeats):
np.random.seed(i)
if encoding=="poisson":
spikes_in = [poisson_encode(p, steps) for p in probs]
elif encoding=="uniform":
frequencies = [prob_to_count(p, steps) for p in probs]
spikes_in = [uniform_encode(f, steps) for f in frequencies]
elif encoding=="linear":
frequencies = [prob_to_count(p, steps) for p in probs]
spikes_in = [linear_encode(f, steps) for f in frequencies]
u_t, U_t, spikes_out = integrate_spike_trains(steps,
spikes_in,
weights_in,
threshold,
tau,
u_rest,
u0
)
title = f"{encoding} encoding,"
plot_spike_integration(steps,
spikes_in,
weights_in,
u_t,
spikes_out,
u_rest,
threshold,
title=title,
)
# + pycharm={"is_executing": false, "name": "#%%\n"}
# Also, we need to define a bunch of parameter values of the simulation
repeats = 5 # how often to run the simulation
steps = 100 # number of time steps simulated
u_rest = 0.0 # resting membrane potential
u0 = u_rest # initial membrane potential
threshold = 1.0 # firing threshold
tau = 0.99 # decay term of membrane potential
# Weights of the incoming neurons
w1 = 0.8
w2 = -0.8
w3 = -0.8
weights_in = [w1, w2] #, w3]
# Firing probabilities of incoming neurons at each time step
p1 = 0.05
p2 = 0.03
p3 = 0.07
probs = [p1, p2] #, p3]
# + pycharm={"is_executing": false, "name": "#%%\n"}
# Finally, lets compare the different spike train generation methods used in simulation
# "linear" encoding is fully deterministic
# and spreads the expected number of spikes in equally spaced intervals
encoding = "linear"
run_simulation()
# + pycharm={"is_executing": false, "name": "#%%\n"}
# "uniform" encoding is deterministic about the number of spikes, but their spacing is uniform random
encoding = "uniform"
run_simulation()
# + pycharm={"is_executing": false, "name": "#%%\n"}
# "poisson" encoding uses a random process to determine the number of spikes and their timing
encoding = "poisson"
run_simulation()
# + pycharm={"name": "#%%\n"}
steps = 100
spikes_in = [
[10, 20],
[11, 21],
]
weights_in = [1.0, 1.0]
u_t, U_t, spikes_out = integrate_spike_trains(steps,
spikes_in,
weights_in,
threshold,
tau,
u_rest,
u0
)
title = f"Spike integration dynamics,"
plot_spike_integration(steps,
spikes_in,
weights_in,
u_t,
spikes_out,
u_rest,
threshold,
title=title,
file="integration_demo.svg"
)
# + pycharm={"name": "#%%\n"}
np.random.seed(3)
fan_in = 100
freq_in = 20
steps = 100
p = 0.1
weights_in = np.random.normal(scale=np.sqrt(2/fan_in) , size=fan_in)
weights_in = np.sort(weights_in)
for s in range(5):
#spikes_in = np.random.randint(1, 101, (fan_in, freq_in))
#spikes_in = np.sort(spikes_in)
#spikes_in = [l.tolist() for l in spikes_in]
spikes_in = [poisson_encode(np.random.normal(p, 0.25), steps) for _ in range(fan_in)]
print(np.mean([len(x) for x in spikes_in]))
weights_ins = []
u_ts = []
U_ts = []
spikes_outs = []
for d, i in enumerate([-0.02, 0.02, 0.15]):
weights_ins.append(weights_in + i)
u_t, U_t, spikes_out = integrate_spike_trains(steps,
spikes_in,
weights_in + i,
threshold,
tau,
u_rest,
u0
)
u_ts.append(u_t)
U_ts.append(U_t)
spikes_outs.append(spikes_out)
title = r"$\mu(\theta)=$" + f"{np.mean(weights_in + i):.3f}, "
"""
fig = plot_spike_integration(steps,
spikes_in,
weights_in + i,
u_t,
U_t,
spikes_out,
u_rest,
threshold,
title=title,
file=f"integration_demo_n{i}_e{s}.svg",
close=False,
show=False,
fig=fig,
)
"""
plot_multicell_spike_integration(
steps,
spikes_in,
weights_ins,
u_ts,
U_ts,
spikes_outs,
u_rest,
threshold,
title="",
show=True,
close=True,
file=f"integration_demo_e{s}.svg",
labels=None)
# + pycharm={"name": "#%%\n"}
for d, i in enumerate([-0.01, 0.02, 0.12]):
plt.plot(weights_in + i)
plt.hlines(0, xmin=0, xmax=100, linestyles="--", lw=1, alpha=0.5)
plt.savefig("weights.svg")
| notebooks/spike_integration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Note
# If you are finding this on github, you will need a local copy of the MAG, which is not an easy thing to create!
#
# ## Objective
# Create a simple process for making a list of Coronavirus-related papers using the Microsoft Academic Graph (MAG).
#
# If possible, also find papers on health-policy and health-education.
#
# ## Process
# 1. Start with a list of relevant papers after searching Microsoft Academic. Retrieve the PaperIds
# 2. Pull the details of the papers from the MAG
# 3. Find related FieldsOfStudy
# 4. Optionally, also find the _children_ of those FieldsOfStudy
# 5. Use those FieldsOfStudy to find more related papers
# 6. Filter those papers for SAGE papers.
# %matplotlib inline
import pandas as pd
import numpy as np
import json
import os
# # Retrieve the data
# 1. Peform a basic search
# > https://academic.microsoft.com/search?q=coronavirus%20covid-19%20pandemic&f=&orderBy=0&skip=0&take=10
# 2. Click the quote marks to build a reference list
# <br>
# <img src="images/untitled.png">
# 3. Download in MS Word format as an xml file
# <br>
# <img src = "images/untitled2.png">
# Read in the XML data and parse out the MAG PaperIds
from bs4 import BeautifulSoup as bs
with open('data/MSA_CV_refs.xml', 'r', encoding = 'utf-8') as f:
soup = bs(f.read(),'lxml')
pids= [x.text[x.text.rfind('/')+1:] for x in soup.find_all('url')]
len(pids)
# # now connect to sql
from sqlalchemy import create_engine
def retrieve_sql_data(sql):
dialect = 'mssql'
driver = 'pyodbc'
driver_string = "ODBC+Driver+17+for+SQL+Server"
username = os.environ['SQL_USER']
password = os.environ['<PASSWORD>']
host = os.environ['SQL_DATALAKE_SVR']
port = '1433'
database = os.environ['SQL_DATALAKE_DB']
con_s = f"{dialect}+{driver}://{username}:{password}@{host}:{port}/{database}?driver={driver_string}&trusted_connection=yes"
engine = create_engine(con_s)
connection = engine.connect()
df = pd.read_sql(sql,connection)
connection.close()
return df
# ## Find related fieldsofstudy
# Get all of the FieldsOfStudy associated with the papers (note that this just gets the Ids of the fieldsofstudy and their scores, but not their names)
# %%time
pids = papers['PaperId'].tolist()
s = "( "+ ", ".join(["'"+str(x)+"'" for x in pids]) + ")"
sql = """
SELECT * FROM [mag].[PaperFieldsOfStudy] WHERE PaperId IN {}
""".format(s)
pfos = retrieve_sql_data(sql)
pfos['FieldOfStudyId'].value_counts().hist(bins = 100)
# optionally filter fields of study to only include fields that appear more than n times
# in the input xml
vc = pd.DataFrame(pfos['FieldOfStudyId'].value_counts(ascending = False))
vc.columns = ['count_in_orig_ls']
# greater than n
# n = 1
# gtn = vc[vc['FieldOfStudyId']>n].index.tolist()
# Filter for fields with high scores only.
pfos['Score'].hist(bins=50)
# pick a cutoff score
cutoff = 0.4
pfos = pfos[(pfos['Score']>cutoff)]
pfos = pfos.merge(right = vc, left_on='FieldOfStudyId', right_index=True, how='left')
# pfos = pfos[pfos['count_in_orig_ls']>0]
# pfos = pfos[pfos['FieldOfStudyId'].isin(set(gtn))]
pfos.shape
best_scores = pfos.groupby('FieldOfStudyId').max()
best_scores = best_scores[['Score']]
best_scores.head()
fields = list(set(pfos['FieldOfStudyId'].tolist()))
len(fields)
# Now retrieve related data on the fields of study. Up until now, we've only had Ids. This will get us the names of the fieldsofstudy so that we can see how relevant they are.
#
# Fields of study have a 'level' associated with them. A level of '0' would be a broad topic, like 'physics' or 'medicine'. A level of '5' might be a very narrow topic like "human mortality from h5n1". So we set our cutoff to avoid taking too broad a swathe of topics.
# %%time
level_cutoff = 1
s = "( "+ ", ".join(["'"+str(x)+"'" for x in fields]) + ")"
sql = """
SELECT * FROM [mag].[FieldOfStudy] WHERE FieldOfStudyId IN {} AND Level >{}
""".format(s,level_cutoff)
fos = retrieve_sql_data(sql)
fos = fos.merge(best_scores, left_on='FieldOfStudyId', right_index=True, how='left')
fos = fos.rename(columns = {'Score':'Best Score'})
fos.shape
fos = fos.merge(right = vc, left_on='FieldOfStudyId', right_index=True, how='left')
fos = fos.sort_values(['Level','Best Score'], ascending = False)
fos.head(3)
# +
# optionally write out so that everyone can see what list of topics you're using
# fos.to_excel('fos.xlsx', index=False)
# -
# Do some further filtering here.
fieldset = set(pfos[pfos['Score'] > 0.6]['FieldOfStudyId'].tolist())
len(fieldset)
fos_subset = fos[(fos['Level']>=3) & (fos['FieldOfStudyId'].isin(fieldset))]
fos_subset.head(3)
# MAG's Fieldsofstudy all have 'children'. That is narrower topics which fit within another topic. So virology might be a child of medicine and coronavirus might be a child of virology.
#
# We probably want all or most of the children of the fieldsofstudy that we have chosen.
# +
# %%time
# add fos children
# now get all SAGE articles with these FOS from the last 3 years
fos_ls = fos_subset['FieldOfStudyId'].tolist()
s = "( "+ ", ".join(["'"+str(x)+"'" for x in fos_ls]) + ")"
sql = """
SELECT * FROM [mag].[FieldOfStudyChildren] WHERE FieldOfStudyId IN {}
""".format(s)
children = retrieve_sql_data(sql)
children.shape
# -
## Add children...
cfields = fields + list(set(children['ChildFieldOfStudyId'].tolist()))
len(cfields)
# +
# %%time
s = "( "+ ", ".join(["'"+str(x)+"'" for x in cfields]) + ")"
sql = """
SELECT * FROM [mag].[FieldOfStudy] WHERE FieldOfStudyId IN {}
""".format(s)
fosc = retrieve_sql_data(sql)
fosc = fosc.merge(best_scores, left_on='FieldOfStudyId', right_index=True, how='left')
fosc = fosc.rename(columns = {'Score':'Best Score'})
fosc.shape
# +
# fos = fos.merge(right = vc, left_on='FieldOfStudyId', right_index=True, how='left')
# -
# Now let's inspect the list.
fosc = fosc.sort_values(['Level'], ascending = False)
fosc.head(10)
fosc.tail(10)
# +
# write out thelist of fields and children that we are using
# fosc.to_excel('fieldsofstudy_coronavirus.xlsx', index = False)
# -
# # Get papers
# +
# %%time
# now get all SAGE articles with these FOS from the last X years
fos_ls = fosc[fosc['Level']>2]['FieldOfStudyId'].tolist()
s = "( "+ ", ".join(["'"+str(x)+"'" for x in fos_ls]) + ")"
sql = """
SELECT * FROM [mag].[PaperFieldsOfStudy] WHERE FieldOfStudyId IN {} AND Score > 0.6
""".format(s)
rel_pfos = retrieve_sql_data(sql)
rel_pfos.shape
# +
# rel_pfos = rel_pfos[rel_pfos['Score']>0.5]
# rel_pfos.shape
# -
years = [x for x in range(2010,2021)]
years
# %%time
# now get the Papers data for these
rel_pids = rel_pfos['PaperId']
s = "( "+ ", ".join(["'"+str(x)+"'" for x in rel_pids]) + ")"
s2 = "( "+ ", ".join([str(x) for x in years]) + ")"
sql = """
SELECT PaperId, OriginalTitle, Doi, Year, Date, Publisher, JournalId, CitationCount, EstimatedCitation
FROM [mag].[Papers]
WHERE PaperId IN {}
AND Year IN {}
AND DocType = 'Journal'
""".format(s, s2)
rel_papers = retrieve_sql_data(sql)
rel_papers.shape
rel_papers['Link'] = rel_papers['Doi'].map(lambda x: 'http:/doi/org/'+str(x) if (x!=None) else x)
rel_papers.to_csv('Related_papers.csv', encoding='utf-8-sig', index=False)
# There we are! Now we have a big list of Coronavirus-related papers from the Microsoft Academic Graph. We can then filter this list for relevant articles published in our subscription journals and make them free-to-read.
#
# This was obviously a super-rough process for finding coronavirus-related content. There are certainly a few articles which have nothing to do with coronavirus in the list. One of our goals was to find papers on health-policy and, while we certainly found them, it's clear that a lot of papers we found on health-policy do not have anything to do with the ongoing coronavirus outbreak. There are also surely papers that are relevant to the outbreak that have not been found by this process.
#
# That said, this process provided better overall coverage of content than a simple keyword search and also saved a huge amount of time compared with manual curation of a list of papers.
#
# ### Next steps
# If anyone is interested in developing this further, I think that there is an opportunity to create a coronavirus classifier using text data and potentially other MAG data, like reference-data, as well as the fieldsofstudy we used here.
#
# That classifier could be very useful for publishers of subscription content who want to drop the paywall on coronavirus research, but it would also be useful to create a feed of related research for anyone who's interested in keeping up with the latest research on the outbreak.
| MAG_00_SQL_keywords_and_papers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Processing of ARIS data for Benchmarking Script
# Looking at the Fairbanks script, these are the fields that are necessary to do the benchmarking:
#
# 'Site ID': This was a 4 letter code identifying the particular facility. It doesn't have to be a 4-letter code for ARIS, just some unique ID.
# 'From': The start date of the billing period. Any date format works.
# 'Thru': The end date of the billing period. This could be multiple months after the start date in the case of oil or propane.
# 'Service Name': This was ['Oil #1', 'Oil #2', 'Electricity', 'Steam', 'Natural Gas', 'Water', 'Sewer', 'Refuse'] for FNSB.
# 'Item Description': This was the description of the particular line item on the bill, like "Customer Charge", "Cost Adjustments", "Demand Charge" plus a very wide variety of other types of charges. In the script, I combined all the miscellaneous charges into "Other Charge". These were billing line items that had no Usage associated with them.
# 'Usage': For usage-based charges, like kWh, kW, gallons, etc, this contains the usage. For "Other Charges" (fixed charges), this is blank.
# 'Cost': $ cost of the line item.
# 'Units': For Usage based charges, the units. These are what was in the Fairbanks data: 'Gallons', 'kWh', 'klbs', 'lbs', 'MMBtu', 'kVAR', 'kW', 'CCF', 'kVARh', 'Loads', 'Cgallons', 'kGal', 'Tons'.
#
# So if we can get the ARIS data into this format, and if we could keep Service Name, Units, and some of the Item Descriptions the same, the script should run. As far as Item descriptions that need to stay the same, I think it is just peak Electric Demand that needs to be labeled like Fairbanks. It can be any one of the following: 'KW Charge', 'On peak demand', 'Demand Charge'
#
# ## To Do
#
# * Standardize values in each field to match FNSB values.
# * To be safe, change spelling of units to match (may not really be necessary although there are some specific tests for CCF)
# * Map 'Service Type' to categories and then only do processing based on those categories.
# * Think more about how long of a billing period is too long.
# * Document the Zip-to-DD_site file. Tell people that Zip code is needed to determine degree-day site. Have them look in Buildings.xlsx file to see if a Degree-day site was assigned.
# * Find degree-day sites for the missing zip codes.
# * Add a README.md in the Data directory to explain what the files are.
# * Update the main README.md.
# * Add a section to the Settings File to accomodate ARIS passwords/URL and other ARIS specific settings.
# * Incorporate new graphs into the `benchmark.py` script, including creation of new Template.
# * Labels in Existing Template Files
# * 'FNSB' appears in Energy Index Comparison.
# * No Propane or Wood present in headings.
# * Oil #2 and #1 probably should be combined and labeled "Fuel Oil", "Heating Oil", or "Oil"
# * Hot Water District Heat and Steam should probably be combined
# +
import time
from datetime import timedelta
import pandas as pd
import numpy as np
import requests
###################### Matplotlib #######################
# import matplotlib pyplot commands
from matplotlib.pyplot import *
# Show Plots in the Notebook
# %matplotlib inline
# 'style' the plot like fivethirtyeight.com website
style.use('bmh')
# -
rcParams['figure.figsize']= (10, 8) # set Chart Size
rcParams['font.size'] = 14 # set Font size in Chart
# ## Read Data from ARIS API
# URLs and Passwords
my_username = 'buildingenergyapp'
my_password = '<PASSWORD>!'
base_url = 'http://arisapi.test.ahfc.us/api/buildingenergy'
building_list = '/GetBuildingList'
building_energy_detail = '/GetBuildingEnergyDetail'
my_params = {'username': my_username,
'password':<PASSWORD>}
building_list_url = base_url + building_list
building_energy_url = base_url + building_energy_detail
results = requests.post(building_list_url, params=my_params).json()
df_bldgs = pd.DataFrame(results)
df_bldgs.head()
# Add a Degree-Day Site column by looking up via zip code
df_zip_to_dd = pd.read_excel('../data/Zip_to_DD_Site.xlsx', skiprows=4)
df_zip_to_dd['zip_code'] = df_zip_to_dd.zip_code.astype(str)
print(df_zip_to_dd.head())
zip_to_dd = dict(zip(df_zip_to_dd.zip_code, df_zip_to_dd.dd_site))
zip_to_dd.get('99645')
df_bldgs['dd_site'] = df_bldgs.BuildingZip.map(zip_to_dd)
df_bldgs.head()
# +
# Check that all zip codes produced a degree-day site; NO
print(sum(df_bldgs.BuildingZip.isna()), sum(df_bldgs.dd_site.isna()))
# So need to find the zip codes that don't map to a Degree-Day site
# 'dd_site != dd_site' is a hack for finding NaN values.
df_no_map = df_bldgs.query('(BuildingZip > "") and (dd_site != dd_site)')
df_no_map.BuildingZip.unique()
# -
col_map = [
('BuildingId', 'site_id'),
('BuildingName', 'site_name'),
('BuildingOwnerName', 'site_category'),
('BuildingStreet', 'address'),
('BuildingCity', 'city'),
('BuildingUsageName', 'primary_func'),
('YearBuilt', 'year_built'),
('SquareFeet', 'sq_ft'),
('dd_site', 'dd_site')
]
old_cols, new_cols = zip(*col_map)
df_bldgs2 = df_bldgs[list(old_cols)].copy()
df_bldgs2.columns = new_cols
df_bldgs2['onsite_gen'] = '' # not used
df_bldgs2.to_excel('data/Buildings.xlsx', startrow=3, index=False)
df_bldgs2.head()
# +
# Now work on the detailed records
print(len(df_bldgs2.site_id.unique()))
my_data = {'username': my_username,
'password':<PASSWORD>,
'buildingId':44}
detail = requests.post(building_energy_url, data=my_data).json()
df_detail = pd.DataFrame(detail['BuildingEnergyDetailList'])
df_detail['UsageDate'] = pd.to_datetime(df_detail.UsageDate)
df_detail.head()
# -
my_data = {'username': my_username,
'password':<PASSWORD>,
'buildingId': None}
dfd = None
next_prn = time.time()
for bldg_id in df_bldgs2.site_id.unique():
my_data['buildingId'] = bldg_id
detail = requests.post(building_energy_url, data=my_data).json()
if len(detail['BuildingEnergyDetailList']):
df_detail = pd.DataFrame(detail['BuildingEnergyDetailList'])
if dfd is not None:
dfd = dfd.append(df_detail, ignore_index=True)
else:
dfd = df_detail.copy()
if time.time() > next_prn:
print('{:,} records fetched'.format(len(dfd)))
next_prn += 10.0 # wait 10 seconds before printing
dfd = dfd.apply(pd.to_numeric, errors='ignore')
dfd[['UsageDate', 'MeterReadDate']] = dfd[['UsageDate', 'MeterReadDate']].apply(pd.to_datetime)
dfd.to_pickle('dfd.pkl')
dfd.head()
len(dfd)
dfd.to_pickle('dfd.pkl')
dfd.head()
dfd.dtypes
# ## Process ARIS API data into Form useable by FNSB Script
# All of the columns in the FNSB CSV file:
fnsb_cols = ["Site ID","Site Name","Vendor Code","Vendor Name","Account Number","Bill Date","Due Date","Entry Date","Invoice #","Voucher #","From","Thru","Service Name","Item Description","Meter Number","Usage","Cost","Units","Account Financial Code","Site Financial Code"]
fnsb_cols
# +
dfd = pd.read_pickle('dfd.pkl')
dfd = dfd.apply(pd.to_numeric, errors='ignore')
dfd[['UsageDate', 'MeterReadDate']] = dfd[['UsageDate', 'MeterReadDate']].apply(pd.to_datetime)
dfd.head()
# -
# Get rid of unneeded columns
dfd.drop(columns=['EnergyTypeId', 'EnergyUnitId', 'UsageYear'], inplace=True)
dfd.head()
dfd.query('EnergyTypeName=="Coal"').BuildingId.unique()
# The dictionary that renames the columns to names needed
# by the benchmarking script
col_map = {
'BuildingId': 'Site ID',
'EnergyTypeName': 'Service Name',
'EnergyUnitTypeName': 'Units',
'EnergyQuantity': 'Usage',
'DollarCost': 'Cost',
}
def add_to_final(df_to_add):
global df_final
df_add = df_to_add.copy()
df_add.rename(columns=col_map, inplace=True)
df_add.drop(columns=['DemandUse', 'DemandCost', 'UsageDate', 'MeterReadDate'], inplace=True)
df_final = df_final.append(df_add, ignore_index=True)
# For the usage end date, 'Thru', use the MeterReadDate if available, otherwise
# use the middle of the UsageDate month.
def thru_date(row):
if pd.isnull(row.MeterReadDate):
return row.UsageDate.replace(day=15)
else:
return row.MeterReadDate
dfd['Thru'] = dfd.apply(thru_date, axis=1)
dfd.head()
dfd.query('MeterReadDate > "2000-01-01"').head()
dfd.EnergyTypeName.value_counts()
dfd.query('EnergyTypeName=="Demand - Electric"')
# Change these to 'Electric'
dfd.loc[dfd.EnergyTypeName == 'Demand - Electric', 'EnergyTypeName'] = 'Electric'
dfd.EnergyTypeName.value_counts()
# +
# There are a number of records where the EnergyQuantity is 0 or NaN,
# which probably occurs because someone doesn't have the bill for that
# month or there was no fuel fill-up in that month. We will eliminate
# those records, because they distort the period over which fuel usage
# occurred for sporadically bought fuels like oil and wood. For
# monthly-billed fuels, we will later in the code make sure that the
# From - Thru billing period only covers 1 month.
# Start by converting 0s to NaN to make future tests easier.
dfd.loc[dfd.EnergyQuantity == 0.0, 'EnergyQuantity'] = np.NaN
dfd.loc[dfd.DemandUse == 0.0, 'DemandUse'] = np.NaN
# Also found that there were a bunch of -1.0 values for DemandUse that
# are very likely not valid.
dfd.loc[dfd.DemandUse == -1.0, 'DemandUse'] = np.NaN
dfd.query('(EnergyQuantity == 0.0) or (DemandUse == 0.0) or (DemandUse == -1.0)') # should be no records
# -
# Now filter down to just the records where we have a number for
# either EnergyQuantity or DemandUse.
print(len(dfd))
mask = ~(dfd.EnergyQuantity.isnull() & dfd.DemandUse.isnull())
dfd = dfd[mask].copy()
print(len(dfd))
# These have Demand but no energy
dfd.query('EnergyQuantity != EnergyQuantity').describe()
# Fill out the From date by using the Thru date from the prior bill
# for the building and for the particular fuel type
df_final = None
for gp, recs in dfd.groupby(['BuildingId', 'EnergyTypeName']):
recs = recs.sort_values(['Thru']).copy()
# Start date comes from prior record
recs['From'] = recs.Thru.shift(1)
recs['Item Description'] = 'Energy'
if df_final is None:
df_final = recs.copy()
else:
df_final = df_final.append(recs, ignore_index=True)
len(df_final)
# +
# For the services that are normally billed on a monthly basis, fill out
# any missing From dates (e.g. the first bill for a building) with a value
# 30 days prior to Thru. Also, restrict the Thru - From difference to 25 to 35 days.
# If it is outside that range, set to Thru - 30 days.
# Fuel types that are normally billed on a monthly basis
mo_fuels = ['Electric', 'Natural Gas', 'Steam District Ht', 'Hot Wtr District Ht']
mask_mo = df_final.EnergyTypeName.isin(mo_fuels)
# Find records of that type that have NaT for From date and
# set to 30 days prior to Thru
df_final.loc[mask_mo & df_final.From.isnull(), 'From'] = df_final.Thru - timedelta(days=30)
# -
# Now find any records where Thru - From is outside 25 - 35 window and fix those.
# Perhaps they are buildings where there are two separate electric bills.
bill_len = df_final.Thru - df_final.From
mask2 = mask_mo & ((bill_len < timedelta(days=25)) | (bill_len > timedelta(days=35)))
df_final.loc[mask2, 'From'] = df_final.Thru - timedelta(days=30)
print(len(df_final[mask2]))
df_final[mask2].head()
# Now work on the fuel types that are not billed monthly. Some of these records
# have NaT for the From date because they were the first record for the building
# and a particular fuel type. We will ultimately delete these. In this step
# find sporadically billed records that have a billing length of greater than 450
# days and put a NaT in for From, so that deleting all From==NaT records will catch
# them as well. A billing period more than 450 days probably indicates that a fuel
# fill was missed making the record invalid.
mask_sporadic = ~mask_mo
mask3 = mask_sporadic & (bill_len > timedelta(days=450))
df_final.loc[mask3, 'From'] = pd.NaT
len(df_final)
# Now eliminate all the sporadically billed records that have a From
# with a NaT
mask_elim = (mask_sporadic & df_final.From.isnull())
df_final = df_final[~mask_elim].copy()
len(df_final)
# +
# Now add the Electric Demand Charge records. The From-Thru dates on these
# have already been set. The demand quantity and cost
# appear in separate, dedicated columns, but we will move them to the 'EnergyQuantity'
# and 'DollarCost' columns.
df_demand = df_final.query('DemandUse > 0 and EnergyTypeName=="Electric"').copy()
df_demand['EnergyQuantity'] = df_demand.DemandUse
df_demand['DollarCost'] = df_demand.DemandCost
df_demand['EnergyUnitTypeName'] = 'kW'
df_demand['Item Description'] = 'Demand Charge'
# add these to the final DataFrame
df_final = df_final.append(df_demand, ignore_index=True)
len(df_final)
# -
df_final.columns
# Eliminate the columns that are not needed
df_final.drop(columns=['DemandCost', 'DemandUse', 'MeterReadDate', 'UsageDate'], inplace=True)
df_final.head()
df_final.info()
col_map = {
'BuildingId': 'Site ID',
'EnergyTypeName': 'Service Name',
'EnergyUnitTypeName': 'Units',
'EnergyQuantity': 'Usage',
'DollarCost': 'Cost',
}
df_final.rename(col_map, axis=1, inplace=True)
df_final.head()
df_final['Units'].value_counts()
df_final['Service Name'].value_counts()
df_final.dtypes
set(zip(df_final['Service Name'], df_final['Units']))
df_final['Item Description'].value_counts()
# These fields are used in the report summarizing vendors.
df_final['Account Number'] = ''
df_final['Vendor Name'] = ''
df_final.groupby('Service Name').sum()
df_final.to_pickle('df_final.pkl')
df_final.to_csv('df_final.csv')
# FNSB Processed Data
dfu3 = pd.read_pickle('../df_processed.pkl')
dfu3.head()
dfu3.service_type.value_counts()
dfu3.units.value_counts()
df_raw = pd.read_pickle('../df_raw.pkl')
df_raw[df_raw['Service Name']=='Electricity'].head()
df_mo = pd.read_pickle('../df_mo.pkl')
df_mo.head()
dfmf = None
for gp, recs in df_mo.groupby(['BuildingId', 'EnergyTypeName']):
recs = recs.query('(DollarCost > 0) or (EnergyQuantity > 0)').copy()
if len(recs) == 0:
continue
recs.sort_values(['Thru'], inplace=True)
# Start date comes from prior record
recs['From'] = recs.Thru.shift(1)
recs['Item Description'] = 'Energy'
recs.loc[recs.From.isnull(), 'From'] = recs.Thru - timedelta(days=30)
# If any of the billing periods are longer than 45 days, then probably
# a missing prior bill. Just make those periods 30 days long.
mask = ((recs.Thru - recs.From) > timedelta(days=45))
recs.loc[mask, 'From'] = recs.Thru - timedelta(days=30)
if dfmf is None:
dfmf = recs.copy()
else:
dfmf = dfmf.append(recs, ignore_index=True)
(dfmf.Thru - dfmf.From).value_counts()
dfmf.query('EnergyUnitTypeName == "kW"')
dfmf.head()
| testing/ARIS_data_processing_alan.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Data referência https://data.sandiegocounty.gov/Health/Prostate-Cancer/gpsr-f4mg
# -
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
prostate_cancer = pd.read_csv('DataPrincipal/Prostate_Cancer.csv')
prostate_cancer
prostate_cancer_principal = prostate_cancer[['CONDITION', 'OUTCOME', 'Year', 'Geography', 'GeoType','GeoName', 'GeoID', 'Region', 'District', 'Total_Male' ]]
prostate_cancer_principal.to_csv('prostate_cancer_principal.csv')
counting_age_prostate_cancer = prostate_cancer.iloc[:, 12:]
counting_age_prostate_cancer.head(5)
counting_age_prostate_cancer_without_values_columns_NAN = counting_age_prostate_cancer[counting_age_prostate_cancer.columns[~counting_age_prostate_cancer.isnull().all()]]
counting_age_prostate_cancer_without_values_columns_NAN
counting_age_prostate_cancer.columns.unique()
male = counting_age_prostate_cancer.iloc[:,0:10]
male = male[sorted(male.columns)]
male.head(3)
# +
male_plot = counting_age_prostate_cancer[['White_Age0_14_Male','White_Age15_24_Male','White_Age25_44_Male' ,'White_Age45_64_Male', 'White_Age65Plus_Male', 'White_Total_Male']]
male_plot.to_csv('people/male.csv')
male_plot_rate = counting_age_prostate_cancer[['White_Age0_14_MaleRate','White_Age15_24_MaleRate','White_Age25_44_MaleRate' ,'White_Age45_64_MaleRate', 'White_Age65Plus_MaleRate', 'White_Total_MaleRate']]
male_plot_rate.to_csv('people/male_rate.csv')
# +
white_male = counting_age_prostate_cancer[['White_Age0_14_Male' , 'White_Age15_24_Male' , 'White_Age25_44_Male', 'White_Age45_64_Male' , 'White_Age65Plus_Male']]
# white_male.sum().plot.bar(figsize=(20,12), rot=1,stacked=True)
white_male.to_csv('people/white_male.csv')
white_male_rate = counting_age_prostate_cancer[['White_Age0_14_MaleRate' , 'White_Age15_24_MaleRate' , 'White_Age25_44_MaleRate', 'White_Age45_64_MaleRate' , 'White_Age65Plus_MaleRate']]
# white_male_rate.sum().plot.bar(figsize=(20,12), rot=1,stacked=True)
white_male_rate.to_csv('people/white_male_rate.csv')
# +
black_male_plot = counting_age_prostate_cancer[['Black_Age0_14_Male', 'Black_Age15_24_Male', 'Black_Age25_44_Male' , 'Black_Age45_64_Male', 'Black_Age65Plus_Male', 'Black_Total_Male']]
black_male_plot.to_csv('people/black.csv')
black_male_plot_rate = counting_age_prostate_cancer[['Black_Age0_14_MaleRate', 'Black_Age15_24_MaleRate', 'Black_Age25_44_MaleRate' , 'Black_Age45_64_MaleRate', 'Black_Age65Plus_MaleRate','Black_Total_MaleRate']]
black_male_plot_rate.to_csv('people/black_rate.csv')
# black_male_plot.sum().plot.bar(figsize=(20,12), rot=1,stacked=True)
# +
hispanic_plot = counting_age_prostate_cancer[['Hispanic_Age0_14_Male' , 'Hispanic_Age15_24_Male' , 'Hispanic_Age25_44_Male' , 'Hispanic_Age45_64_Male', 'Hispanic_Age65Plus_Male']]
hispanic_plot.to_csv('people/hispanic.csv')
# hispanic_plot.sum().plot.bar(figsize=(20,12), rot=1,stacked=True)
hispanic_plot_rate = counting_age_prostate_cancer[['Hispanic_Age0_14_MaleRate' , 'Hispanic_Age15_24_MaleRate' , 'Hispanic_Age25_44_MaleRate' , 'Hispanic_Age45_64_MaleRate', 'Hispanic_Age65Plus_MaleRate']]
hispanic_plot_rate.to_csv('people/hispanic_rate.csv')
# +
api_plot = counting_age_prostate_cancer[['API_Age0_14_Male' , 'API_Age15_24_Male' , 'API_Age25_44_Male' , 'API_Age45_64_Male', 'API_Age65Plus_Male' , 'API_Total_Male']]
api_plot.to_csv('people/api.csv')
# hispanic_plot.sum().plot.bar(figsize=(20,12), rot=1,stacked=True)
api_plot_rate = counting_age_prostate_cancer[['Hispanic_Age0_14_MaleRate' , 'Hispanic_Age15_24_MaleRate' , 'Hispanic_Age25_44_MaleRate' , 'Hispanic_Age45_64_MaleRate', 'Hispanic_Age65Plus_MaleRate']]
api_plot_rate.to_csv('people/api_rate.csv')
# +
aian_plot = counting_age_prostate_cancer[['AIAN_Age0_14_Male' , 'AIAN_Age15_24_Male' , 'AIAN_Age25_44_Male' , 'AIAN_Age45_64_Male', 'AIAN_Age65Plus_Male' , 'AIAN_Total_Male']]
aian_plot.to_csv('people/aian.csv')
# hispanic_plot.sum().plot.bar(figsize=(20,12), rot=1,stacked=True)
aian_plot_rate = counting_age_prostate_cancer[['AIAN_Age0_14_MaleRate' , 'AIAN_Age15_24_MaleRate' , 'AIAN_Age25_44_MaleRate' , 'AIAN_Age45_64_MaleRate', 'AIAN_Age65Plus_MaleRate', 'AIAN_Total_MaleRate']]
aian_plot_rate.to_csv('people/aian_rate.csv')
# +
other_notaian_plot = counting_age_prostate_cancer[['Other_notAIAN_Age0_14_Male' , 'Other_notAIAN_Age15_24_Male' , 'Other_notAIAN_Age25_44_Male' , 'Other_notAIAN_Age45_64_Male', 'Other_notAIAN_Age65Plus_Male' , 'Other_notAIAN_Total_Male']]
other_notaian_plot.to_csv('people/other_notaian.csv')
# hispanic_plot.sum().plot.bar(figsize=(20,12), rot=1,stacked=True)
other_notaian_plot_rate = counting_age_prostate_cancer[['Other_notAIAN_Age0_14_MaleRate' , 'Other_notAIAN_Age15_24_MaleRate' , 'Other_notAIAN_Age25_44_MaleRate' , 'Other_notAIAN_Age45_64_MaleRate', 'Other_notAIAN_Age65Plus_MaleRate' , 'Other_notAIAN_Total_MaleRate']]
other_notaian_plot_rate.to_csv('people/other_notaian_rate.csv')
# -
other_male = counting_age_prostate_cancer.iloc[:, 71:]
other_male.head(3)
black_male = counting_age_prostate_cancer.iloc[:,23:34]
black_male.head(3)
# black_male.sum().plot.bar(figsize=(20,12))
hispanic =counting_age_prostate_cancer.iloc[:,34:46]
hispanic.head(3)
api_male = counting_age_prostate_cancer.iloc[:, 47:58]
api_male.head(3)
aian_male = counting_age_prostate_cancer.iloc[:, 58:70]
aian_male.head(3)
names_dataframe = list()
for people in types_of_people:
names_dataframe.append(f'dataframe_type_people_{people}')
for coluna in counting_age_prostate_cancer_without_valuecolumnsolumns_NAN.columns.unique():
counting_age_prostate_cancer_without_values_columns_NAN.groupby(coluna)[coluna].sum().plot.bar(figsize=(20,12),label=coluna)
death = prostate_cancer_principal[prostate_cancer_principal['CONDITION'] == 'Death']
outcome = prostate_cancer_principal.OUTCOME.unique()
outcome
# About San Diego
#https://mmasc.org/167/San-Diego-County-Region
geography = prostate_cancer_principal.Geography.unique()
geography
death = prostate_cancer_principal[prostate_cancer_principal['OUTCOME'] == 'Death']
death.head(5)
death.plot(x='Year', y ='Total_Male', figsize=(20,12))
| Data-Cleaning-San_diego.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + deletable=false editable=false
# Initialize Otter
import otter
grader = otter.Notebook("hw08.ipynb")
# -
# # Homework 8: Confidence Intervals
# **Reading**:
# * [Estimation](https://www.inferentialthinking.com/chapters/13/estimation.html)
# Please complete this notebook by filling in the cells provided.
#
# Directly sharing answers is not okay, but discussing problems with the course staff or with other students is encouraged. Refer to the policies page to learn more about how to learn cooperatively.
#
# For all problems that you must write our explanations and sentences for, you **must** provide your answer in the designated space. Moreover, throughout this homework and all future ones, please be sure to not re-assign variables throughout the notebook! For example, if you use `max_temperature` in your answer to one question, do not reassign it later on.
# +
# Don't change this cell; just run it.
import numpy as np
from datascience import *
# These lines do some fancy plotting magic.
import matplotlib
# %matplotlib inline
import matplotlib.pyplot as plt
plt.style.use('fivethirtyeight')
import warnings
warnings.simplefilter('ignore', FutureWarning)
# -
# ## 1. Thai Restaurants
#
# Ben and Frank are trying see what the best Thai restaurant in Berkeley is. They survey 1500 UC Berkeley students selected uniformly at random, and ask each student what Thai restaurant is the best (*Note: this data is fabricated for the purposes of this homework*). The choices of Thai restaurant are Lucky House, Imm Thai, Thai Temple, and Thai Basil. After compiling the results, Ben and Frank release the following percentages from their sample:
#
# |Thai Restaurant | Percentage|
# |:------------:|:------------:|
# |Lucky House | 8% |
# |Imm Thai | 52% |
# |Thai Temple | 25% |
# |Thai Basil | 15% |
#
# These percentages represent a uniform random sample of the population of UC Berkeley students. We will attempt to estimate the corresponding *parameters*, or the percentage of the votes that each restaurant will receive from the entire population (the entire population is all UC Berkeley students). We will use confidence intervals to compute a range of values that reflects the uncertainty of our estimates.
#
# The table `votes` contains the results of the survey.
# Just run this cell
votes = Table.read_table('votes.csv').sample(with_replacement = False)
votes
# + [markdown] deletable=false editable=false
# **Question 1.** Complete the function `one_resampled_percentage` below. It should return Imm Thai's **percentage** of votes after simulating one bootstrap sample of `tbl`.
#
# **Note:** `tbl` will always be in the same format as `votes`.
#
# <!--
# BEGIN QUESTION
# name: q1_1
# manual: false
# -->
# +
def one_resampled_percentage(tbl):
...
one_resampled_percentage(votes)
# + deletable=false editable=false
grader.check("q1_1")
# + [markdown] deletable=false editable=false
# **Question 2.** Complete the `percentages_in_resamples` function such that it returns an array of 2500 bootstrapped estimates of the percentage of voters who will vote for Imm Thai. You should use the `one_resampled_percentage` function you wrote above.
#
# *Note:* There are no public tests for this question, the autograder cell below will return 0.0% passed.
#
# <!--
# BEGIN QUESTION
# name: q1_2
# manual: false
# -->
# + for_assignment_type="solution"
def percentages_in_resamples():
percentage_imm = make_array()
...
# + deletable=false editable=false
grader.check("q1_2")
# -
# In the following cell, we run the function you just defined, `percentages_in_resamples`, and create a histogram of the calculated statistic for the 2,500 bootstrap estimates of the percentage of voters who voted for Imm Thai. Based on what the original Thai restaurant percentages were, does the graph seem reasonable? Talk to a friend or ask a TA if you are unsure!
resampled_percentages = percentages_in_resamples()
Table().with_column('Estimated Percentage', resampled_percentages).hist("Estimated Percentage")
# + [markdown] deletable=false editable=false
# **Question 3.** Using the array `resampled_percentages`, find the values at the two edges of the middle 95% of the bootstrapped percentage estimates. (Compute the lower and upper ends of the interval, named `imm_lower_bound` and `imm_upper_bound`, respectively.)
#
# <!--
# BEGIN QUESTION
# name: q1_3
# manual: false
# -->
# -
imm_lower_bound = ...
imm_upper_bound = ...
print("Bootstrapped 95% confidence interval for the percentage of Imm Thai voters in the population: [{:f}, {:f}]".format(imm_lower_bound, imm_upper_bound))
# + deletable=false editable=false
grader.check("q1_3")
# + [markdown] deletable=false editable=false
# **Question 4.** The survey results seem to indicate that Imm Thai is beating all the other Thai restaurants combined among voters. We would like to use confidence intervals to determine a range of likely values for Imm Thai's true lead over all the other restaurants combined. The calculation for Imm Thai's lead over Lucky House, Thai Temple, and Thai Basil combined is:
#
# $$\text{Imm Thai's % of the vote} - \text{(Lucky House's % of the vote + Thai Temple's % of the vote + Thai Basil's % of the vote)}$$
#
# Define the function `one_resampled_difference` that returns **exactly one value** of Imm Thai's percentage lead over Lucky House, Thai Temple, and Thai Basil combined from one bootstrap sample of `tbl`.
#
# <!--
# BEGIN QUESTION
# name: q1_4
# manual: false
# -->
# -
def one_resampled_difference(tbl):
bootstrap = ...
imm_percentage = ...
lh_percentage = ...
tt_percentage = ...
tb_percentage = ...
...
# + deletable=false editable=false
grader.check("q1_4")
# -
# HIDEEN TEST
np.random.seed(123)
-6 <= float(one_resampled_difference(votes)) <= 15
# + [markdown] deletable=false editable=false
# <!-- BEGIN QUESTION -->
#
# **Question 5.**
# Write a function called `leads_in_resamples` that finds 2,500 bootstrapped estimates (the result of calling `one_resampled_difference`) of Imm Thai's lead over Lucky House, Thai Temple, and Thai Basil combined. Plot a histogram of the resulting samples.
#
# **Note:** Imm Thai's lead can be negative.
#
# <!--
# BEGIN QUESTION
# name: q1_5
# manual: true
# -->
# + export_pdf=true for_assignment_type="solution"
def leads_in_resamples():
...
sampled_leads = leads_in_resamples()
Table().with_column('Estimated Lead', sampled_leads).hist("Estimated Lead")
# + [markdown] deletable=false editable=false
# <!-- END QUESTION -->
#
# **Question 6.** Use the simulated data from Question 5 to compute an approximate 95% confidence interval for Imm Thai's true lead over Lucky House, Thai Temple, and Thai Basil combined.
#
# <!--
# BEGIN QUESTION
# name: q1_6
# manual: false
# -->
# -
diff_lower_bound = ...
diff_upper_bound = ...
print("Bootstrapped 95% confidence interval for Imm Thai's true lead over Lucky House, Thai Temple, and Thai Basil combined: [{:f}, {:f}]".format(diff_lower_bound, diff_upper_bound))
# + deletable=false editable=false
grader.check("q1_6")
# -
# ## 2. Interpreting Confidence Intervals
#
# The staff computed the following 95% confidence interval for the percentage of Imm Thai voters:
#
# $$[49.40, 54.47]$$
#
# (Your answer may have been a bit different; that doesn't mean it was wrong!)
# + [markdown] deletable=false editable=false
# <!-- BEGIN QUESTION -->
#
# #### Question 1
# Can we say there is a 95% probability that the interval [49.40, 54.47] contains the true percentage of the population that votes for Imm Thai as the best Berkeley Thai restaurant? Answer "yes" or "no" and explain your reasoning.
#
# *Note:* ambiguous answers using language like "sometimes" or "maybe" will not receive credit.
#
# <!--
# BEGIN QUESTION
# name: q2_1
# manual: true
# -->
# -
# _Type your answer here, replacing this text._
# + [markdown] deletable=false editable=false
# <!-- END QUESTION -->
#
# <!-- BEGIN QUESTION -->
#
# **Question 2**
#
# The staff also created 70%, 90%, and 99% confidence intervals from the same sample, but we forgot to label which confidence interval represented which percentages! Match each confidence level (70%, 90%, 99%) with its corresponding interval in the cell below (e.g. __ % CI: [49.87, 54.0] $\rightarrow$ replace the blank with one of the three confidence levels). **Then**, explain your thought process and how you came up with your answers.
#
# The intervals are below:
#
# * [49.87, 54.00]
# * [50.67, 53.27]
# * [48.80, 55.40]
#
# <!--
# BEGIN QUESTION
# name: q2_2
# manual: true
# -->
# -
# _Type your answer here, replacing this text._
# + [markdown] deletable=false editable=false
# <!-- END QUESTION -->
#
# #### Question 3
# Suppose we produced 5,000 new samples (each one a uniform random sample of 1,500 voters/students) from the population and created a 95% confidence interval from each one. Roughly how many of those 5,000 intervals do you expect will actually contain the true percentage of the population?
#
# Assign your answer to `true_percentage_intervals`.
#
# <!--
# BEGIN QUESTION
# name: q2_3
# manual: false
# -->
# -
true_percentage_intervals = ...
# + deletable=false editable=false
grader.check("q2_3")
# + [markdown] deletable=false editable=false
# Recall the second bootstrap confidence interval you created, which estimated Imm Thai's lead over Lucky House, Thai Temple, and Thai Basil combined. Among
# voters in the sample, Imm Thai's lead was 4%. The staff's 95% confidence interval for the true lead (in the population of all voters) was
#
# $$[-0.80, 8.80]$$
#
# Suppose we are interested in testing a simple yes-or-no question:
#
# > "Is the percentage of votes for Imm Thai tied with the percentage of votes for Lucky House, Thai Temple, and Thai Basil combined?"
#
# Our null hypothesis is that the percentages are equal, or equivalently, that Imm Thai's lead is exactly 0. Our alternative hypothesis is that Imm Thai's lead is not equal to 0. In the questions below, don't compute any confidence interval yourself - use only the staff's 95% confidence interval.
#
#
# **Question 4**
#
# Say we use a 5% P-value cutoff. Do we reject the null, fail to reject the null, or are we unable to tell using our staff confidence interval?
#
# Assign `restaurants_tied` to the number corresponding to the correct answer.
#
# 1. Reject the null / Data is consistent with the alternative hypothesis
# 2. Fail to reject the null / Data is consistent with the null hypothesis
# 3. Unable to tell using our staff confidence interval
#
# *Hint:* If you're confused, take a look at [this chapter](https://www.inferentialthinking.com/chapters/13/4/using-confidence-intervals.html) of the textbook.
#
# <!--
# BEGIN QUESTION
# name: q2_4
# manual: false
# -->
# -
restaurants_tied = ...
# + deletable=false editable=false
grader.check("q2_4")
# + [markdown] deletable=false editable=false
# #### Question 5
# What if, instead, we use a P-value cutoff of 1%? Do we reject the null, fail to reject the null, or are we unable to tell using our staff confidence interval?
#
# Assign `cutoff_one_percent` to the number corresponding to the correct answer.
#
# 1. Reject the null / Data is consistent with the alternative hypothesis
# 2. Fail to reject the null / Data is consistent with the null hypothesis
# 3. Unable to tell using our staff confidence interval
#
# <!--
# BEGIN QUESTION
# name: q2_5
# manual: false
# -->
# -
cutoff_one_percent = ...
# + deletable=false editable=false
grader.check("q2_5")
# + [markdown] deletable=false editable=false
# #### Question 6
# What if we use a P-value cutoff of 10%? Do we reject, fail to reject, or are we unable to tell using our confidence interval?
#
# Assign `cutoff_ten_percent` to the number corresponding to the correct answer.
#
# 1. Reject the null / Data is consistent with the alternative hypothesis
# 2. Fail to reject the null / Data is consistent with the null hypothesis
# 3. Unable to tell using our staff confidence interval
#
# <!--
# BEGIN QUESTION
# name: q2_6
# manual: false
# -->
# -
cutoff_ten_percent = ...
# + deletable=false editable=false
grader.check("q2_6")
# + [markdown] deletable=false editable=false
# ---
#
# To double-check your work, the cell below will rerun all of the autograder tests.
# + deletable=false editable=false
grader.check_all()
# + [markdown] deletable=false editable=false
# ## Submission
#
# Make sure you have run all cells in your notebook in order before running the cell below, so that all images/graphs appear in the output. The cell below will generate a zip file for you to submit. **Please save before exporting!**
# + deletable=false editable=false
# Save your notebook first, then run this cell to export your submission.
grader.export()
# -
#
| hw/hw08/.ipynb_checkpoints/hw08-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (tabula-muris-env)
# language: python
# name: tabula-muris-env
# ---
# +
from anndata import read_h5ad
from scipy import stats, sparse
import numpy as np
import sys
from collections import Counter
from OnClass.OnClassModel import OnClassModel
from utils import read_ontology_file, read_data, run_scanorama_multiply_datasets
from config import ontology_data_dir, scrna_data_dir, model_dir, Run_scanorama_batch_correction, NHIDDEN, MAX_ITER
# -
# ## read data
# +
#Change train_file, test_file, train_label, test_label according to your datasets
#train_label is the key of labels in .h5ad train_file.
train_file = scrna_data_dir + '/Lemur/microcebusBernard.h5ad'
test_file = scrna_data_dir + '/Lemur/microcebusAntoine.h5ad'
train_label = 'cell_ontology_id'
test_label = 'cell_ontology_id'
model_path = model_dir + 'example_file_model'
print ('read ontology data and initialize training model...')
cell_type_nlp_emb_file, cell_type_network_file, cl_obo_file = read_ontology_file('cell ontology', ontology_data_dir)
OnClass_train_obj = OnClassModel(cell_type_nlp_emb_file = cell_type_nlp_emb_file, cell_type_network_file = cell_type_network_file)
print ('read training single cell data...')
train_feature, train_genes, train_label, _, _ = read_data(train_file, cell_ontology_ids = OnClass_train_obj.cell_ontology_ids,
exclude_non_leaf_ontology = False, tissue_key = 'tissue', AnnData_label_key = train_label, filter_key = {},
nlp_mapping = False, cl_obo_file = cl_obo_file, cell_ontology_file = cell_type_network_file, co2emb = OnClass_train_obj.co2vec_nlp)
#you can also replace it with your own data and make sure that:
#train_feature is a ncell by ngene matrix
#train_genes is a ngene long vector of gene names
#train_label is a ncell long vector
# -
# ## Embed cell types based on the Cell Ontology Graph
print ('embed cell types using the cell ontology...')
OnClass_train_obj.EmbedCellTypes(train_label)
# ## Training
# +
# print ('read test single cell data...')
x = read_h5ad(test_file)
test_label = x.obs[test_label].tolist()
test_feature = x.X.toarray()
test_genes = np.array([x.upper() for x in x.var.index])
# optional batch correction
if Run_scanorama_batch_correction:
train_feature, test_feature = run_scanorama_multiply_datasets([train_feature, test_feature], [train_genes, test_genes], scan_dim = 10)[1]
print (np.shape(train_feature), np.shape(test_feature))
print ('generate pretrain model. Save the model to $model_path...')
cor_train_feature, cor_test_feature, cor_train_genes, cor_test_genes = OnClass_train_obj.ProcessTrainFeature(train_feature, train_label, train_genes, test_feature = test_feature, test_genes = test_genes)
OnClass_train_obj.BuildModel(ngene = len(cor_train_genes), nhidden = NHIDDEN)
OnClass_train_obj.Train(cor_train_feature, train_label, save_model = model_path, max_iter = MAX_ITER)
# -
# ## Classify test cells
# +
print ('initialize test model. Load the model from $model_path...')
OnClass_test_obj = OnClassModel(cell_type_nlp_emb_file = cell_type_nlp_emb_file, cell_type_network_file = cell_type_network_file)
cor_test_feature = OnClass_train_obj.ProcessTestFeature(cor_test_feature, cor_test_genes, use_pretrain = model_path, log_transform = False)
OnClass_test_obj.BuildModel(ngene = None, use_pretrain = model_path)
#use_normalize=False will return a tree-based prediction, where parent node often has higher score than child node. use_normalize=True will normalize among child nodes and parent nodes
pred_Y_seen, pred_Y_all, pred_label = OnClass_test_obj.Predict(cor_test_feature, test_genes = cor_test_genes, use_normalize=True)
pred_label_str = [OnClass_test_obj.i2co[l] for l in pred_label]
#x.obs['OnClass_annotation_flat_based_ontology_ID'] = pred_label_str
pred_Y_seen, pred_Y_all, pred_label = OnClass_test_obj.Predict(cor_test_feature, test_genes = cor_test_genes, use_normalize=False)
pred_label_str = [OnClass_test_obj.i2co[l] for l in pred_label]
#x.obs['OnClass_annotation_tree_based_ontology_ID'] = pred_label_str
#x.write(scrna_data_dir + 'Pilot12.annotated.h5ad')
| run_OnClass_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img width="100" src="https://carbonplan-assets.s3.amazonaws.com/monogram/dark-small.png" style="margin-left:0px;margin-top:20px"/>
#
# # Forest Emissions Tracking - Phase I
#
# _by <NAME> and <NAME> (CarbonPlan)_
#
# March 29, 2020
#
# ## Introduction
#
# In general, greenhouse gasses (GHGs) arising from forest land use changes can be
# attributed to both natural factors (e.g. wildfire) and human activities (e.g.
# deforestation). Our approach is to build upon an existing body of research that
# has provided high-resolution satellite-based estimates of aboveground biomass
# (Spawn et al., 2020), forest cover change (Hansen et al., 2013), and change
# attribution (Curtis et al., 2018). While many of the necessary data products
# already exist, we can integrate, extend, or update these resources to provide
# global, current estimates that can be integrated with the other resources
# produced by the coalition.
#
# Specifically, for any given spatial extent and time duration ($t1$ to $t2$), we
# can use three quantities — existing biomass, forest cover change, and change
# attribution — to estimate the effective GHG emissions from land use changes. The
# simplest estimate is:
#
# $\Delta Biomass (t) = TotalBiomass (t) * \Delta ForestCover (\%)$
#
# $Emissions (tCO_2) = \Delta Biomass (t) * 0.5 (tC/t) * 3.67 (tC02 / tC)$
#
# where $\Delta ForestCover$ is the fraction of pixels within the given spatial
# extent that experienced a stand-replacement disturbance between $t1$ and $t2$.
# The $TotalBiomass$ is estimated as the aboveground biomass at time $t1$. This
# estimate can be further refined by attributing, for each pixel, the source of
# forest cover loss (e.g. wildfire, deforestation, etc.), and using those sources
# to express emissions fractionally and/or exclude certain categories from total
# estimates (e.g. rotational clear-cutting within tree plantations). Pixel-wise
# estimates can then be aggregated into province and country-wide estimates.
#
# ## Setup
#
# To begin, we'll import a handful of Python libraries and set a few constants.
#
# +
# %matplotlib inline
import dask
import matplotlib.pyplot as plt
import geopandas
import intake
import xarray as xr
import rioxarray
import numpy as np
import seaborn as sns
from dask.distributed import Client
from shapely.geometry import mapping
from sklearn.metrics import mean_squared_error
from data import open_hansen_2018_tile
# TODO: switch to carbonplan style theme
plt.rcParams["font.family"] = "monospace"
TC02_PER_TC = 3.67
TC_PER_TBM = 0.5
SQM_PER_HA = 10000
ORNL_SCALING = 0.1
# +
# start a local Dask cluster
# client = Client(n_workers=4)
# display(client)
# -
# Next we will open a handful of datasets that we'll use in our analysis. Note
# that these datasets are all included in an _Intake Catalog_ called
# `catalog.yaml`. Intake, along with a small helper script (`data.py`) will handle
# opening all of our data for us.
#
# data catalog
cat = intake.open_catalog("catalog.yaml")
# The first dataset we will open is the Hansen et al (2018) forest cover change
# dataset. We also open auxillary datasets like above ground biomass (`abg`) and
# two partial estimates of emissions (`emissions_ha` and `emissions_px`) that are
# all available on the same 30x30m grid. All of this data is distributed in
# 10x10degree tiles so we define which tile we'll be looking at for this
# demonstration. We also define a smaller bounding box (`box`) that we will use
# for a few analyses later on.
#
# +
# open a single 10x10degree tile of the Hansen 30x30m data
lat = "50N"
lon = "130W"
box = dict(lat=slice(41.5, 41.0), lon=slice(-123, -122.5))
ds = open_hansen_2018_tile(lat, lon)
display(ds)
# -
# Next, we open a few other datasets:
#
# 1. The Spawn and Gibbs (2020) Global Aboveground and Belowground Biomass Carbon
# Density Maps for the Year 2010: `ds_ornl`.
# 1. A shapefile including the each of the US states: `us_states`.
# 1. A data table of estimated state aggregated biomass for each US state:
# `df_fia_state`.
# 1. A data table of biomass for all the FIA plot locations in California:
# `df_ca_fia`.
#
# + jupyter={"outputs_hidden": true}
# open some data for benchmarking
# ORNL biomass
ds_ornl = cat.ornl_biomass.to_dask() * ORNL_SCALING
# shapefile for US states
states = geopandas.read_file(
"/home/jovyan/.local/share/cartopy/shapefiles/natural_earth/cultural/ne_50m_admin_1_states_provinces_lakes.shp"
)
us_states = states.query("admin == 'United States of America'").set_index(
"iso_3166_2"
)
us_states.crs = "EPSG:4326"
# State-wide Biomass aggregation from FIA
df_fia_state = cat.fia_by_state.read()
# California Biomass by FIA plot
df_ca_fia = cat.fia_ca.read()
# +
variables = ["treecover2000", "lossyear", "agb", "emissions_ha"]
titles = [
"Tree Cover Fraction (year: 2000)\n(Hanson et al. 2013) ",
"Year of Tree Cover Loss\n(Hanson et al. 2013)",
"Aboveground Biomass\n(Zarin et al. 2016)",
"Emissions",
]
units = ["[%]", "[year]", "[t/Ha]", "[tC02/Ha]"]
fig, axes = plt.subplots(
nrows=2, ncols=2, sharex=True, sharey=True, figsize=(8, 8)
)
ds_plot = ds.sel(**box)
kwargs = {}
for i, var in enumerate(variables):
ax = axes.flat[i]
if i == 2:
kwargs = dict(vmin=0, vmax=600, extend="max")
if i == 3:
kwargs = dict(vmin=0, vmax=1100, extend="max")
if i == 1:
kwargs = dict(vmin=2000, vmax=2018, levels=19)
da = ds_plot[var]
da.plot.imshow(
ax=ax, add_labels=False, cbar_kwargs={"label": units[i]}, **kwargs
)
ax.set_title(titles[i])
fig.tight_layout()
# -
# The Hansen tree cover and loss data is compressed into a initial treecover field
# and a lossyear field. In the cell below, we unpack this data into a
# 3-dimensional mask of losses by year. This will come in handy when we start
# analyzing emissions by year. We add this new field (`d_treecover`) to our
# dataset.
#
# calculate d_treecover
years = xr.DataArray(range(2001, 2019), dims=("year",), name="year")
loss_frac = []
for year in years:
loss_frac.append(xr.where((ds["lossyear"] == year), ds["treecover2000"], 0))
ds["d_treecover"] = xr.concat(loss_frac, dim=years)
display(ds)
# We can now sum over all the years in our data to see the change in treecover for
# each pixel in our tile.
#
ds["d_treecover"].sum("year").sel(**box).plot()
# ### Emissions Calculation
#
# As we discussed above, the equation to compute emissions from a change in above
# ground biomass is simply a conversion of lost biomass to an equivalent mass of
# C02. The function below represents how this is performed in our analysis.
#
def calc_emissions(ds):
d_biomass = ds["agb"] * ds["d_treecover"]
emissions = d_biomass * TC_PER_TBM * TC02_PER_TC
return emissions
ds["emissions"] = calc_emissions(ds)
display(ds["emissions"])
# Now that we have calculated the emissions at each point in our tile, we can
# visualize the results. The maps below show emissions, by year, for each pixel in
# our tile.
#
temp = ds["emissions"].sel(**box)
temp = temp.where(temp).persist()
fg = temp.plot(col="year", col_wrap=5, robust=True)
# We can now sum over our tile to compute the emissions from this region by year.
# To do this, we first need to compute the per-pixel area to perform an accurate
# unit conversion.
#
def compute_grid_area(da):
R = 6.371e6
total_area = 4 * np.pi * R ** 2
dϕ = np.radians((da["lat"][1] - da["lat"][0]).values)
dλ = np.radians((da["lon"][1] - da["lon"][0]).values)
dA = R ** 2 * np.abs(dϕ * dλ) * np.cos(np.radians(da["lat"]))
areacella = dA * (0 * da + 1)
return areacella / SQM_PER_HA
def scatter_style(ax, aspect=("equal", "box")):
"""helper function to style the next few scatter plots"""
if aspect:
ax.set_aspect(*aspect)
# Hide the right and top spines
ax.spines["right"].set_visible(False)
ax.spines["top"].set_visible(False)
# Only show ticks on the left and bottom spines
ax.yaxis.set_ticks_position("left")
ax.xaxis.set_ticks_position("bottom")
# +
area = compute_grid_area(temp)
fig, ax = plt.subplots(figsize=(5, 3))
(temp * area).sum(("lat", "lon")).plot(ax=ax)[0]
plt.ylabel("emissions (tCO2)")
plt.xticks(ticks=range(2000, 2020, 2))
scatter_style(ax, aspect=None)
# -
# ## Comparison
#
# We focus on comparative analysis on the biomass datasets used so far in this
# notebook. For the purposes of this demonstration we will take a look at a small
# tile, to qualatatively assess the Zarin and Spawn & Gibbs data.
#
(ds_ornl["aboveground_biomass_carbon_2010"]).sel(**box).plot(vmin=0, vmax=200)
(ds["agb"] * TC_PER_TBM).sel(**box).plot(vmin=0, vmax=300)
# ## Ground Truthing
#
# ### Part 1. US State Aggregation
#
# Our ground truthing has taken two initial forms. First we compare regional
# (state) aggregated biomass estimates from Spawn & Gibbs to FIA state aggregated
# data.
#
@dask.delayed
def calc_state_biomass(source, gdf):
"""lazy helper function to compute aggregated biomass by shape"""
import rioxarray
# open the dataset inside the task
obj = source.to_dask()["aboveground_biomass_carbon_2010"] * ORNL_SCALING
obj.rio.set_spatial_dims("lon", "lat")
obj.rio.set_crs(4326)
clipped = obj.rio.clip(gdf.geometry.apply(mapping), gdf.crs, drop=True)
cell_area = compute_grid_area(clipped)
return (clipped * cell_area).sum().values.item() / 1e6 # Tg C
# +
# aggregate the ORNL (Spawn and Gibbs) data by state.
from dask.diagnostics import ProgressBar
biomass = [
calc_state_biomass(cat.ornl_biomass, us_states.iloc[[i]])
for i in range(len(us_states))
]
with ProgressBar():
us_states["biomass_computed"] = dask.compute(
*biomass, scheduler="single-threaded"
)
# extract the FIA data and put it in our us_states dataframe
for st, val in df_fia_state.values:
us_states.loc[f"US-{st}", "biomass_fia"] = val
us_states.head(n=2)
# -
# Now that we've computed biomass estimates for each state, we can compare them to
# the FIA aggregated data.
#
# +
state_biomass = (
us_states[["biomass_fia", "biomass_computed", "geometry"]]
.drop(labels="US-AK")
.dropna(axis=0)
)
f, ax = plt.subplots(figsize=(5, 5))
sns.regplot(
x="biomass_fia",
y="biomass_computed",
data=state_biomass,
color="#7EB36A",
ax=ax,
)
scatter_style(ax)
skill = mean_squared_error(
state_biomass["biomass_fia"],
state_biomass["biomass_computed"],
squared=False,
)
ax.text(0.65, 0.02, f"rmse: {skill:.2f}", transform=ax.transAxes)
# -
# a few simple difference statistics
state_biomass["pdiff"] = (
(state_biomass["biomass_computed"] - state_biomass["biomass_fia"])
/ state_biomass["biomass_fia"]
) * 100
state_biomass["adiff"] = (
state_biomass["biomass_computed"] - state_biomass["biomass_fia"]
)
state_biomass.plot(
column="biomass_fia",
legend=True,
vmin=0,
vmax=1000,
cmap="Greens",
legend_kwds={"label": "Biomass (Tg C)", "orientation": "horizontal"},
)
state_biomass.plot(
column="pdiff",
legend=True,
vmin=-100,
vmax=100,
cmap="RdBu",
legend_kwds={"label": "Difference (% FIA)", "orientation": "horizontal"},
)
state_biomass.plot(
column="adiff",
legend=True,
vmin=-200,
vmax=200,
cmap="RdBu",
legend_kwds={"label": "Difference (Tg C)", "orientation": "horizontal"},
)
# ### Part 2. FIA Plot Comparison
#
# For the second form of ground truthing we compare biomass estimates to actual
# FIA plot data. We have extracted all FIA plot locations in the state of
# California (shown in figure below) and we compare those to Spawn and Gibbs data
# below.
#
df_ca_fia.plot.scatter("lon", "lat", c="carbon", cmap="viridis", vmax=120)
fia_da = df_ca_fia.to_xarray().rename({"index": "plot"})[["lat", "lon"]]
da = (
ds_ornl["aboveground_biomass_carbon_2010"]
.sel(lat=fia_da.lat, lon=fia_da.lon, method="nearest", tolerance=1.0)
.load()
)
df_ca_fia["ornl_biomass"] = da.to_series()
df_ca_fia.head()
# +
f, axes = plt.subplots(nrows=3, figsize=(5, 8), sharex=True, sharey=True)
sns.regplot(
x="carbon", y="ornl_biomass", data=df_ca_fia, color="#7EB36A", ax=axes[0]
)
skill = mean_squared_error(
df_ca_fia["carbon"], df_ca_fia["ornl_biomass"], squared=False
)
axes[0].annotate(
f"rmse: {skill:.2f}",
xy=(0.15, 0.8),
xycoords="data",
xytext=(0.75, 0.1),
textcoords="axes fraction",
)
axes[0].set_title(f"all plots (n={len(df_ca_fia)})")
df = df_ca_fia.query("year == 2010")
sns.regplot(x="carbon", y="ornl_biomass", data=df, color="#7EB36A", ax=axes[1])
skill = mean_squared_error(df["carbon"], df["ornl_biomass"], squared=False)
axes[1].annotate(
f"rmse: {skill:.2f}",
xy=(0.15, 0.8),
xycoords="data",
xytext=(0.75, 0.1),
textcoords="axes fraction",
)
axes[1].set_title(f"2010 only (n={len(df)})")
df = df.query("forested > 0.99")
sns.regplot(x="carbon", y="ornl_biomass", data=df, color="#7EB36A", ax=axes[2])
skill = mean_squared_error(df["carbon"], df["ornl_biomass"], squared=False)
axes[2].annotate(
f"rmse: {skill:.2f}",
xy=(0.15, 0.8),
xycoords="data",
xytext=(0.75, 0.1),
textcoords="axes fraction",
)
axes[2].set_title(f"2010 & 100% forested (n={len(df)})")
for ax in axes:
plt.sca(ax)
plt.ylabel("ORNL biomass \n(tC / ha)")
plt.xlabel("FIA biomass (tC / ha)")
scatter_style(ax)
fig.tight_layout()
# +
f, ax = plt.subplots(figsize=(6, 6))
sns.kdeplot(
df_ca_fia.carbon,
df_ca_fia.ornl_biomass,
ax=ax,
)
plt.ylabel("ORNL biomass \n(tC / ha)")
plt.xlabel("FIA biomass (tC / ha)")
plt.xlim(0, 150)
plt.ylim(0, 150)
scatter_style(ax)
# -
# ## References:
#
# - <NAME>. et al. (2018) ‘Classifying drivers of global forest loss’,
# Science, 361(6407), pp. 1108–1111. doi: 10.1126/science.aau3445.
# - <NAME> al. (2007) ‘A Project for Monitoring Trends in Burn
# Severity’, Fire Ecology, 3(1), pp. 3–21. doi: 10.4996/fireecology.0301003.
# - <NAME>. et al. (2013) ‘High-resolution global maps of 21st-century
# forest cover change’, Science, 342(6160), pp. 850–853. doi:
# 10.1126/science.1244693.
# - <NAME>., et al. (2020) ‘Asynchronous carbon sink saturation in African and
# Amazonian tropical forests’, Nature, 80–87(2020). doi:
# 10.1038/s41586-020-2035-0
# - <NAME> al. (2020) ‘Harmonized global maps of above and belowground
# biomass carbon density in the year 2010’, Scientific Data. doi:
# 10.1038/s41597-020-0444-4.
# - <NAME> al. (2016) ‘Can carbon emissions from tropical deforestation
# drop by 50% in 5 years?’, Global Change Biology, pp. 1336–1347. doi:
# 10.1111/gcb.13153.
#
| notebooks/single_tile.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
def hello_world():
print("Hello World!")
hello_world()
# -
# # A Hello World Function in Python
#
# This is the most simple function and typically used in every language as a first test.
# It's only purpose is, to print "**Hello World!**"
| Peter's-Hello-World.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Lecture 4: Programming a quantum algorithm with pyQuil
#
# This notebook is part of the course [CS 269Q: Quantum Computer Programming](https://cs269q.stanford.edu/), offered at Stanford in the Spring of 2019.
#
# It was created by [<NAME>](https://github.com/karalekas), Quantum Software Engineer at [Rigetti Computing](https://rigetti.com/) and Guest Lecturer for the course.
#
# ## Introduction
#
# In this lecture, we will build up the mathematical and software tools required to implement our very first quantum algorithm—[Deutsch's algorithm](https://en.wikipedia.org/wiki/Deutsch%E2%80%93Jozsa_algorithm)—using [pyQuil](https://github.com/rigetti/pyquil) and the [QVM](https://github.com/rigetti/qvm) to step through the protocol and inspect the wavefunction. The three things that we will need to understand before we get to the algorithm are:
#
# - [Part I: The wavefunction & quantum circuits](#wavefunction)
# - [Part II: Classical logic & function evaluation](#classical-logic)
# - [Part III: Quantum parallelism](#parallelism)
#
# Once we've worked through these concepts, we will then conclude with [Part IV: Deutsch's algorithm](#deutsch).
#
# ## Setup
#
# If you are running this from [Binder](https://mybinder.org/v2/gh/karalekas/stanford-cs269q/master?urlpath=lab/tree/Lecture4.ipynb), then you don't need to worry about any setup, but I do recommend that you click on "Kernel > Restart & Clear Output" before running through the notebook.
#
# If you've cloned this notebook locally, you'll need to get a QVM server running and install all the notebook dependencies. To run a QVM server on the standard port (5000), the command is simply `qvm -S`. This assumes that you have a correctly configured `~/.forest_config` file with `qvm_address` equal to `http://localhost:5000`. As for dependencies, this notebook uses `python3.6` and requires `pyquil`, `matplotlib`, and `qutip` to run. The `qutip` package in turn requires preinstalled `cython`, `numpy`, and `scipy`.
#
# <a id='wavefunction'></a>
# ## Part I: The wavefunction & quantum circuits
#
# We begin by doing our standard imports. We will explain below what each of these things means.
from pyquil import Program
from pyquil.api import QVMConnection
from pyquil.gates import I, H, X, CNOT, MEASURE
# ### Gross Plotting Code
#
# Understanding this code is not important at all for this lecture. I will eventually clean it up and add the useful bits to the mainline codebase for pyquil.
# +
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
from pyquil.wavefunction import get_bitstring_from_index, Wavefunction
from qutip import Bloch, basis
def plot_bloch(wf: Wavefunction, axes=None, fig=None):
if len(wf.amplitudes) > 2:
raise ValueError('Bloch sphere plotting only works with 1Q')
state = (wf.amplitudes[0] * basis(2,0) + wf.amplitudes[1] * basis(2,1))
b = Bloch(fig=fig, axes=axes)
b.add_states(state)
b.render(fig=fig, axes=axes)
def plot_probabilities(wf: Wavefunction, axes=None, qubit_subset=None):
prob_dict = wf.get_outcome_probs()
if qubit_subset:
sub_dict = {}
qubit_num = len(wf)
for i in qubit_subset:
if i > (2**qubit_num - 1):
raise IndexError("Index {} too large for {} qubits.".format(i, qubit_num))
else:
sub_dict[get_bitstring_from_index(i, qubit_num)] = prob_dict[get_bitstring_from_index(i, qubit_num)]
prob_dict = sub_dict
axes.set_ylim(0, 1)
axes.set_ylabel('Outcome probability', fontsize=16)
axes.set_xlabel('Bitstring outcome', fontsize=16)
axes.bar(range(len(prob_dict)), prob_dict.values(), align='center', color='#6CAFB7')
axes.set_xticks(range(len(prob_dict)))
axes.set_xticklabels(prob_dict.keys(), fontsize=14)
def plot_wf(wf: Wavefunction, wf0=None, wf1=None, bitstring_subset=None):
if len(wf.amplitudes) == 2:
fig = plt.figure(figsize=(12, 6))
wf_ax = fig.add_subplot(121)
plot_probabilities(wf, axes=wf_ax, qubit_subset=bitstring_subset)
bloch_ax = fig.add_subplot(122, projection='3d')
plot_bloch(wf, axes=bloch_ax, fig=fig)
fig.suptitle(f'$|\psi>$ = {wf}\n', fontsize=16)
elif len(wf.amplitudes) == 4 and wf0 is not None and wf1 is not None:
fig = plt.figure(figsize=(18, 6))
wf_ax = fig.add_subplot(131)
plot_probabilities(wf, axes=wf_ax, qubit_subset=bitstring_subset)
bloch1_ax = fig.add_subplot(132, projection='3d')
plot_bloch(wf1, axes=bloch1_ax, fig=fig)
bloch0_ax = fig.add_subplot(133, projection='3d')
plot_bloch(wf0, axes=bloch0_ax, fig=fig)
fig.suptitle(f'$|\psi>$ = {wf}\n', fontsize=18)
else:
fig = plt.figure(figsize=(6, 6))
wf_ax = fig.add_subplot(111)
plot_probabilities(wf, axes=wf_ax, qubit_subset=bitstring_subset)
fig.suptitle(f'$|\psi>$ = {wf}\n', fontsize=16)
# -
# ### Initialize our `QVMConnection` object
#
# The `QVMConnection` object contains everything we need to communicate with the QVM server process. It also has two methods that we will be concerned with for today—`wavefunction` and `run`.
#
# The `wavefunction` method takes a `Program` object as input, uses it to evolve the state of the QVM, and then returns the wavefunction (quantum memory) to the user, in the form of a complex array (state vector). We'll mostly be using the `wavefunction` method today because it is incredibly convenient and instructive, but it is important to note that there is no way to inspect the wavefunction on a real quantum computer.
#
# The `run` method also takes `Program` object as input and uses it to evolve the state of the QVM, but instead of returning the wavefunction, it returns the block of classical memory named "ro". Thus, `Program`s supplied to `run` need to have readout memory declared, and measurement at the end, to return something to the user. We'll learn more about this is part 2.
qvm = QVMConnection()
# ### Create a `Program` and add gates
#
# The `Program` object in pyquil is what we use to write Quil programs, which can then be sent to the QVM or a quantum computer. After constructing the `Program`, we can add gates to it by doing something like `p += X(0)`, which adds the `X` gate on qubit 0 to the `Program` stored in variable `p`.
p = Program()
p += X(0)
# ### Wavefunction initial state is $|0\rangle$
#
# Now that we understand the `QVMConnection` object, `Program` object, and gates, let's put the three of them together and run on the QVM. As we know, our QVM starts out entirely in the $|0\rangle$ state, which we verify in the next cell.
#
# $$|\psi\rangle = |0\rangle$$
#
# The plots you will see are (1) a bar plot of the probability distribution of the wavefunction, and (2) a Bloch sphere plot of the wavefunction. In addition, the full wavefunction is printed out at the top of the combined plot.
p = Program()
p += I(0)
wf = qvm.wavefunction(p)
plot_wf(wf)
# ### Run an $X$ gate to flip to $|1\rangle$
#
# $$X|0\rangle = |1\rangle$$
p = Program()
p += X(0)
wf = qvm.wavefunction(p)
plot_wf(wf)
# ### Create the 1Q superposition state $|+\rangle$ using the $H$ (Hadamard) gate
#
# $$H|0\rangle = \dfrac{|0\rangle + |1\rangle}{\sqrt{2}} = |+\rangle$$
p = Program()
p += H(0)
wf = qvm.wavefunction(p)
plot_wf(wf)
# ### Create the 2Q computational basis state $|10\rangle$
#
# $$(X \otimes I)|00\rangle = |10\rangle$$
# +
p = Program()
p += I(0)
p += X(1)
wf = qvm.wavefunction(p)
# this state is a product state
p0 = Program()
p0 += I(0)
wf0 = qvm.wavefunction(p0)
p1 = Program()
p1 += X(1)
wf1 = qvm.wavefunction(p1)
plot_wf(wf, wf0, wf1)
# -
# ### Create the 2Q superposition state $|+,+\rangle$ using two Hadamard gates
#
# $$(H \otimes H)|00\rangle = \left( \dfrac{|0\rangle + |1\rangle}{\sqrt{2}}\right) \otimes \left( \dfrac{|0\rangle + |1\rangle}{\sqrt{2}}\right) = |+,+\rangle$$
# +
p = Program()
p += H(0)
p += H(1)
wf = qvm.wavefunction(p)
# this state is a product state
p0 = Program()
p0 += H(0)
wf0 = qvm.wavefunction(p0)
p1 = Program()
p1 += H(1)
wf1 = qvm.wavefunction(p1)
plot_wf(wf, wf0, wf1)
# -
# ### Create the 2Q entangled (Bell) state $|\Phi^+\rangle$
#
# Note that, because we are in an entangled state, we can no longer factor our state into the tensor product of two individual qubit states (called a "product state"), and therefore cannot plot two independent Bloch spheres as we did above.
#
# $$\text{CNOT}_{0,1}(I \otimes H)|00\rangle = \text{CNOT}_{0,1}|0\rangle \otimes \left( \dfrac{|0\rangle + |1\rangle}{\sqrt{2}}\right) = \dfrac{|00\rangle + |11\rangle}{\sqrt{2}} = |\Phi^+\rangle$$
p = Program()
p += H(0)
p += CNOT(0, 1)
wf = qvm.wavefunction(p)
plot_wf(wf)
# <a id='classical-logic'></a>
# ## Part II: Classical logic & function evaluation
#
# In Computer Science, we learn about Boolean logic gates like `NOT`, `AND`, `OR`, and `XOR`. In quantum computing, we can implement these classical logic gates, but we must reimplement these gates in a way that respects the unitary requirements of quantum logic gates.
# ### Function for calculating the truth table of some circuit
#
# The contents of the first half of this function are not particularly important to understand. However, the loop at the bottom introduces some new concepts that are important for running on the QVM. Because we are using the `run` method of the QVM now instead of the `wavefunction` method, we need to declare our readout memory "ro" and measure our output qubit into the readout register.
#
# In the following code block, we take an existing `Program` and add a `DECLARE` statement to it. The first argument is the name of the block of memory ("ro"), and the second argument is the data type ("BIT"). There is an optional third argument for specifying the length of the block, but it defaults to 1 (which is what we want). This line looks different than the typical `+= INSTRUCTION` format because the return value of the `declare` method is important to us. This variable `ro` is then passed to the `MEASURE` instruction in the following code block.
#
# ```
# ro = p.declare('ro', 'BIT')
# ```
#
# In this code block we add a `MEASURE` instruction to the end of our `Program`. The first argument of the instruction is the qubit that we want to measure (in our case this is whatever qubit we declare to represent the "out" bit of our Boolean function). The second argument is the memory reference variable that corresponds to a particular readout register. Our readout memory is of length 1, and so we select the first (zero-indexed) address to use.
#
# ```
# p += MEASURE(out, ro[0])
# ```
# +
import itertools
gate_map = {'0': I, '1': X}
def calculate_truth_table(qvm: QVMConnection, circuit: Program, name: str, out: int, ancilla: int):
if ancilla:
qubits = circuit.get_qubits() - {ancilla}
else:
qubits = circuit.get_qubits()
num_qubits_set = set(range(len(qubits)))
if qubits != num_qubits_set:
raise ValueError('Please index the qubits of your circuit starting at 0')
bitstrings = itertools.product('01', repeat=len(qubits))
print(f'\n{name}\n')
print('in -> out\n---------')
for bitstring in bitstrings:
p = Program()
ro = p.declare('ro', 'BIT')
for idx, bit in enumerate(reversed(list(bitstring))):
p += gate_map[bit](idx)
p += circuit
p += MEASURE(out, ro[0])
result = qvm.run(p)
print(f"{''.join(bitstring)} -> {result[0][0]}")
# -
# ### Encoding Boolean functions of 1-bit domain in quantum circuits
#
# $$x \in \{0,1\} \hspace{1cm} f(x) \rightarrow \{0,1\}$$
#
# One-bit boolean functions represent the simplest classical logic we can implement on a quantum computer. There are four possible one-bit functions $f(x)$, and we will work through all of them.
#
# ### Balanced functions
#
# $$\text{Balanced-}I : (0 \rightarrow 0, 1 \rightarrow 1)
# \hspace{1 cm}
# \text{Balanced-}X : (0 \rightarrow 1, 1 \rightarrow 0)$$
#
# For the balanced 1-bit functions, it’s pretty easy to come up with a quantum circuit that works. If we use just an $I$ gate for Balanced-$I$ and just an $X$ gate for Balanced-$X$, we can produce a quantum circuit $U_f$ that maps $|x\rangle \rightarrow |f(x)\rangle$. Knowing the gate and and output, we can reproduce the input, which means our circuit satisfies the requirements of unitarity—it is length persevering and invertible.
#
# ### Constant functions
#
# $$\text{Constant-}0 : (0 \rightarrow 0, 1 \rightarrow 0)
# \hspace{1 cm}
# \text{Constant-}1 : (0 \rightarrow 1, 1 \rightarrow 1)$$
#
# However, coming up with the circuit for the constant functions seems less trivial. You can write down a matrix that maps the 0 state to the 0 state and the 1 state to the 0 state, but this matrix has some problems. It is not invertible (determinant 0) and it is not length preserving (superposition state changes length), and therefore it is not unitary. We can also see that it is not reversible simply from the truth table—knowing the output and the gate isn’t enough to get back to the input.
#
# ### Ancilla qubits
#
# $$|0, x\rangle \rightarrow |f(x), x\rangle$$
#
# In order to write this function as a quantum circuit, we need to introduce a new concept—the ancilla qubit. An ancilla qubit is an additional qubit used in a computation that we know the initial state of. Using an ancilla, we can produce a quantum circuit $U_f$ that maps $|0, x\rangle \rightarrow |f(x), x\rangle$. Now, we can come up with a unitary matrix (albeit a trivial one) that allows us to evaluate constant functions. For the Constant-$0$, we just simply do nothing to the ancilla, and its state encodes $f(x)$. And for the Constant-$1$, all we have to do is flip the ancilla with an $X$ gate, and we get $f(x)$ for all $x$.
# +
one_bit_functions = """
DEFCIRCUIT BALANCED_I in out:
CNOT in out
DEFCIRCUIT BALANCED_X in out:
X out
CNOT in out
DEFCIRCUIT CONSTANT_0 in out:
I in
I out
DEFCIRCUIT CONSTANT_1 in out:
I in
X out
"""
print('ONE-BIT FUNCTIONS\n=================')
for circuit in ['BALANCED_I 0 1', 'BALANCED_X 0 1', 'CONSTANT_0 0 1', 'CONSTANT_1 0 1']:
p = Program(one_bit_functions)
p += circuit
calculate_truth_table(qvm=qvm, circuit=p, name=circuit.split(' ')[0], out=1, ancilla=1)
# -
# ### The Quantum `XOR` gate
#
# The boolean function `XOR` (for "exclusive or") takes in two bits $x$ and $y$ and returns 1 if and only if the values of the bits are different from one another. Otherwise, it returns 0. The operation is written as $x \oplus y$, and although it is a two-bit function, we can implement it as a quantum circuit without an ancilla, by simply using the $\text{CNOT}$ gate.
#
# $$\textrm{CNOT}_{0,1}|y, x\rangle = |y \oplus x, x\rangle$$
#
# **Note**: We are using `DEFCIRCUIT` below for consistency, but you could just as easily not use it and instead replace all instances of `QXOR` with `CNOT` for the same effect.
# +
xor_circuit = """
DEFCIRCUIT QXOR x y:
CNOT x y
"""
print('QUANTUM XOR\n===========')
xor = Program(xor_circuit)
xor += 'QXOR 0 1'
calculate_truth_table(qvm=qvm, circuit=xor, name='QXOR', out=1, ancilla=None)
# -
# ### Deutsch Oracle
#
# In Deutsch's algorithm, you are given something called an oracle (referred to as $U_f$), which maps $|y, x\rangle \rightarrow |y \oplus f(x), x\rangle$, and the goal is to determine a global property of the function $f(x)$ with as few queries to the oracle as possible. We can combine the two concepts above (one-bit function evaluation with ancillas, and the `XOR` gate), to produce the four implementations of the Deutsch Oracle with one ancilla qubit.
#
# $$U_f : |y, 0, x\rangle \rightarrow |y \oplus f(x), 0, x\rangle$$
# +
deutsch_oracles_naive = """
DEFCIRCUIT DEUTSCH_BALANCED_I x y fx:
CNOT x fx
CNOT fx y
CNOT x fx
DEFCIRCUIT DEUTSCH_BALANCED_X x y fx:
X fx
CNOT x fx
CNOT fx y
CNOT x fx
X fx
DEFCIRCUIT DEUTSCH_CONSTANT_0 x y fx:
I x
I fx
CNOT fx y
DEFCIRCUIT DEUTSCH_CONSTANT_1 x y fx:
I x
X fx
CNOT fx y
X fx
"""
print('NAIVE DEUTSCH ORACLES\n=====================')
for circuit in ['DEUTSCH_BALANCED_I 0 1 2', 'DEUTSCH_BALANCED_X 0 1 2', 'DEUTSCH_CONSTANT_0 0 1 2', 'DEUTSCH_CONSTANT_1 0 1 2']:
p = Program(deutsch_oracles_naive)
p += circuit
calculate_truth_table(qvm=qvm, circuit=p, name=circuit.split(' ')[0], out=1, ancilla=2)
# -
# ### Optimized Deutsch Oracle
#
# For pedagogical reasons, it is nice to separate out the three steps in the Deutsch Oracle—evaluate $f(x)$, calculate $y \oplus f(x)$, and then return the ancilla to $|0\rangle$—but in practice we always want to implement our circuits in as few gates as possible (this is especially important when running on a real, noisy quantum computer!). Below, we show how we can rewrite each of the four Deutsch Oracle implementations (which we call $U_f$) without the need for an ancilla qubit.
#
# $$U_f : |y, x\rangle \rightarrow |y \oplus f(x), x\rangle$$
# +
deutsch_oracles = """
DEFCIRCUIT DEUTSCH_BALANCED_I x y:
CNOT x y
DEFCIRCUIT DEUTSCH_BALANCED_X x y:
X x
CNOT x y
X x
DEFCIRCUIT DEUTSCH_CONSTANT_0 x y:
I x
I y
DEFCIRCUIT DEUTSCH_CONSTANT_1 x y:
I x
X y
"""
print('OPTIMIZED DEUTSCH ORACLES\n=========================')
for circuit in ['DEUTSCH_BALANCED_I 0 1', 'DEUTSCH_BALANCED_X 0 1', 'DEUTSCH_CONSTANT_0 0 1', 'DEUTSCH_CONSTANT_1 0 1']:
p = Program(deutsch_oracles)
p += circuit
calculate_truth_table(qvm=qvm, circuit=p, name=circuit.split(' ')[0], out=1, ancilla=None)
# -
# <a id='parallelism'></a>
# ## Part III: Quantum Parallelism
#
# In the previous section, we showed that we could implement classical logic using quantum circuits. However, when using a computational basis state ($|0\rangle$ or $|1\rangle$), we don't do anything more interesting than a classical computer can do. If we instead feed a superposition state into one of these circuits, we can effectively evaluate a function $f(x)$ on multiple values of $x$ at once!
#
# $$U_f : |0,+\rangle \rightarrow \dfrac{|f(0), 0\rangle + |f(1), 1\rangle}{\sqrt{2}}$$
#
# $$U_f : |0,-\rangle \rightarrow \dfrac{|f(0), 0\rangle - |f(1), 1\rangle}{\sqrt{2}}$$
#
# It is important to note, that although this quantum parallelism concept is interesting, we are unable to learn about both $f(0)$ and $f(1)$ when the states above are in that form. This is due to the fact that we can only extract one classical bit's worth of information from a quantum computer (of 1 qubit) when we measure it. But, as we will find in Deutsch's algorithm below, we can cleverly take advantage of quantum parallelism to do things that a classical computer cannot, even with the constraint that measurement yields only one classical bit.
# ### Run Balanced-$I$ on state $|0, +\rangle$
#
# $$\text{CNOT}_{0,1}|0,+\rangle = \dfrac{|0, 0\rangle + |1, 1\rangle}{\sqrt{2}}$$
balanced_I = Program(one_bit_functions)
balanced_I += H(0)
balanced_I += 'BALANCED_I 0 1'
wf = qvm.wavefunction(balanced_I)
plot_wf(wf)
# ### Run Balanced-$I$ on state $|0, -\rangle$
#
# $$\text{CNOT}_{0,1}|0,-\rangle = \dfrac{|0, 0\rangle - |1, 1\rangle}{\sqrt{2}}$$
balanced_I = Program(one_bit_functions)
balanced_I += X(0)
balanced_I += H(0)
balanced_I += 'BALANCED_I 0 1'
wf = qvm.wavefunction(balanced_I)
plot_wf(wf)
# ### Run Balanced-$X$ on state $|0, +\rangle$
#
# $$(I \otimes X)\text{CNOT}_{0,1}(I \otimes X)|0, +\rangle = \dfrac{|1, 0\rangle + |0, 1\rangle}{\sqrt{2}}$$
balanced_X = Program(one_bit_functions)
balanced_X += H(0)
balanced_X += 'BALANCED_X 0 1'
wf = qvm.wavefunction(balanced_X)
plot_wf(wf)
# ### Run Balanced-$X$ on state $|0, -\rangle$
#
# $$(I \otimes X)\text{CNOT}_{0,1}(I \otimes X)|0, -\rangle = \dfrac{|1, 0\rangle - |0, 1\rangle}{\sqrt{2}}$$
balanced_X = Program(one_bit_functions)
balanced_X += X(0)
balanced_X += H(0)
balanced_X += 'BALANCED_X 0 1'
wf = qvm.wavefunction(balanced_X)
plot_wf(wf)
# <a id='deutsch'></a>
# ## Part IV: Deutsch's algorithm
#
# **Goal**: Determine if a one-bit function $f(x)$ is *constant* or *balanced*. We show that we can do this with only one query to the Deutsch Oracle, which is impossible on a classical computer, which would require two queries to the Deutsch Oracle to determine this global property of $f(x)$.
#
# As part of the algorithm, we are given a Deutsch Oracle and are unaware of which function $f(x)$ it implements. However, for the purposes of understanding exactly how the algorithm works, we will pick an instantiation of the oracle to use with the QVM. But, of course, the quantum speedup only makes sense if we don't know the contents of the oracle and instead treat it as a black box.
# ### Step 0: Initial state $|10\rangle$
#
# $$|\psi_0\rangle = |10\rangle$$
# +
step0 = Program(deutsch_oracles)
step0 += [I(0), X(1)]
wf = qvm.wavefunction(step0)
# this is a product state
p0 = Program()
p0 += I(0)
wf0 = qvm.wavefunction(p0)
p1 = Program()
p1 += X(1)
wf1 = qvm.wavefunction(p1)
plot_wf(wf, wf0, wf1)
# -
# ### Step 1: Prepare superpositions
#
# We can't do anything interesting with computational basis states, so to take advantage of quantum parallelism we put our qubits in superposition states.
#
# $$|\psi_1\rangle = (H \otimes H)|\psi_0\rangle = (H \otimes H)|10\rangle = \left( \dfrac{|0\rangle - |1\rangle}{\sqrt{2}}\right) \otimes \left( \dfrac{|0\rangle + |1\rangle}{\sqrt{2}}\right) = |-,+\rangle$$
# +
step1 = Program(H(0), H(1))
wf = qvm.wavefunction(step0 + step1)
# this is a product state still
p0 = Program(I(0), H(0))
wf0 = qvm.wavefunction(p0)
p1 = Program(X(0), H(0))
wf1 = qvm.wavefunction(p1)
plot_wf(wf, wf0, wf1)
# -
# ### Step 2: Apply the Deutsch Oracle
#
# We learned earlier that the action of the Deutsch Oracle on input state $|y,x\rangle$ is $U_f|y, x\rangle \rightarrow |y \oplus f(x), x\rangle$. So, what happens if we apply the Deutsch Oracle to the input state $|-,x\rangle$?
#
# $$U_f|-, x\rangle \rightarrow |-\rangle \otimes (-1)^{f(x)}|x\rangle$$
#
# We get a negative sign if $f(x) = 1$, and the state is unchanged if $f(x) = 0$. However, something interesting happens when we apply $U_f$ to the state $|-,+\rangle$, which is our state $|\psi_1\rangle$:
#
# $$|\psi_2\rangle = U_f|\psi_1\rangle = U_f|-, +\rangle = |-\rangle \otimes \left( \dfrac{(-1)^{f(0)}|0\rangle + (-1)^{f(1)}|1\rangle}{\sqrt{2}}\right) =
# \begin{cases}
# \pm |-\rangle \otimes \left( \dfrac{|0\rangle + |1\rangle}{\sqrt{2}}\right) = \pm |-,+\rangle \text{ if constant;}\\
# \pm |-\rangle \otimes \left( \dfrac{|0\rangle - |1\rangle}{\sqrt{2}}\right) = \pm |-,-\rangle \text{ if balanced.}
# \end{cases}$$
#
# If $f(x)$ is balanced, this has the effect of changing the relative phase between the $|0\rangle$ and $|1\rangle$ components of qubit 0's state, which flips it from $|+\rangle$ to $|-\rangle$. This is interesting, because the action of our oracle on the computational basis state $|y,x\rangle$ is to change the state of qubit 1 and leave qubit 0 alone. But, when our qubits are in superposition states, the balanced oracle actually changes the state of qubit 0, and leaves the state of qubit 1 alone.
#
# Although we can work through the math without choosing an implementation of the oracle, we can't use the QVM unless we give it an actual circuit. So, we will use the Deutsch Oracle `DEFCIRCUIT`s we defined in the previous section, to see what happens next. You can comment out any of the four `step2` lines to experiment with different oracle instantations!
step2 = Program('DEUTSCH_BALANCED_I 0 1')
# step2 = Program('DEUTSCH_BALANCED_X 0 1')
# step2 = Program('DEUTSCH_CONSTANT_0 0 1')
# step2 = Program('DEUTSCH_CONSTANT_1 0 1')
wf = qvm.wavefunction(step0 + step1 + step2)
plot_wf(wf)
# ### Step 3: Return to 2Q computational basis states
#
# We know that the outcome of step 3 is to produce one of two superposition product states, $|-, +\rangle$ or $|-, -\rangle$.
#
# $$|\psi_3\rangle = (H \otimes H)|\psi_2\rangle =
# \begin{cases}
# \pm (H \otimes H)|-, +\rangle = \pm |10\rangle \text{ if constant;}\\
# \pm (H \otimes H)|-, -\rangle = \pm |11\rangle \text{ if balanced.}
# \end{cases}$$
#
# Thus, we are in two distinct 2Q computational basis states, dependent on the nature of $f(x)$. We could then measure the state of qubit 0 one time, and we would immediately know the answer of whether $f(x)$ is constant or balanced.
step3 = Program(H(1), H(0))
wf = qvm.wavefunction(step0 + step1 + step2 + step3)
plot_wf(wf)
# ### Conclusions
#
# So, we were able to learn about a *global property* of the function $f(x)$ in just one query to the Deutsch Oracle, which is impossible on a classical computer. Although the problem statement for Deutsch's algorithm is a bit contrived, if you can suspend your judgment, you can imagine that we could take some of the non-classical concepts of this algorithm and apply them to a more complex scenario to actually produce an interesting quantum speedup. And, later in the course, you will do exactly this!
# ## Further Questions & Support
#
# Join the Rigetti-Forest Slack workspace [here](https://join.slack.com/t/rigetti-forest/shared_invite/enQtNTUyNTE1ODg3MzE2LWExZWU5OTE4YTJhMmE2NGNjMThjOTM1MjlkYTA5ZmUxNTJlOTVmMWE0YjA3Y2M2YmQzNTZhNTBlMTYyODRjMzA) and feel free to reach out to me (@peter) with any questions!
# ## Time to run on the QPU!
#
# If we have time left, we'll log into the Rigetti lab and try to run Deutsch's algortithm on a real quantum processor ([QPU](https://www.rigetti.com/qpu)) using our Quantum Cloud Services ([QCS](https://www.rigetti.com/qcs)) platform! Rigetti has allocated some QCS credits for this course, so for your final projects you will be able to run on the QPU as well, if you so choose.
| Lecture4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.4
# language: python
# name: python-374
# ---
# # MPI
#
# Often, a parallel algorithm requires moving data between the engines. One way is to push and pull over the `DirectView`. However, this is slow because all of the data has to get through the controller to the client and then back to the final destination.
#
# A much better option is to use the [Message Passing Interface (MPI)](https://de.wikipedia.org/wiki/Message_Passing_Interface). IPython's parallel computing architecture was designed from the ground up to integrate with MPI. This notebook gives a brief introduction to using MPI with IPython.
# ## Requirements
#
# * A standard MPI implementation like [OpenMPI](http://www.open-mpi.org/) or [MPICH](MPICH).
#
# For Debian/Ubuntu these can be installed with
#
# ```
# $ sudo apt install openmpi-bin
# ```
#
# or
#
# ```
# $ sudo apt install mpich
# ```
#
# Alternatively, OpenMPI or MPICH can also be installed with [Spack](../../../productive/envs/spack/use.rst): the packages are `openmpi` or `mpich`.
#
#
# * [mpi4py](https://mpi4py.readthedocs.io/)
# ## Starting the engines with activated MPI
#
# ### Automatic start with `mpiexec` and `ipcluster`
#
# This can be done with, e.g.
#
# ```
# $ pipenv run ipcluster start -n 4 --profile=mpi
# ```
#
# For this, however, a corresponding profile must first be created; see [configuration](config.rst).
#
# ### Automatic start with PBS and `ipcluster`
#
# The `ipcluster` command also offers integration in [PBS](https://www.pbspro.org/). You can find more information about this in [Using ipcluster in PBS mode](https://ipyparallel.readthedocs.io/en/latest/process.html#using-ipcluster-in-pbs-mode).
# ## Example
#
# The following notebook cell calls `psum.py` with the following content:
#
# ```
# from mpi4py import MPI
# import numpy as np
#
# def psum(a):
# locsum = np.sum(a)
# rcvBuf = np.array(0.0,'d')
# MPI.COMM_WORLD.Allreduce([locsum, MPI.DOUBLE],
# [rcvBuf, MPI.DOUBLE],
# op=MPI.SUM)
# return rcvBuf
# ```
# +
import ipyparallel as ipp
c = ipp.Client(profile='mpi')
view = c[:]
view.activate()
view.run('psum.py')
view.scatter('a',np.arange(16,dtype='float'))
view['a']
# -
# %px totalsum = psum(a)
view['totalsum']
| docs/refactoring/performance/ipyparallel/mpi.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <div align="center"><img src='http://ufq.unq.edu.ar/sbg/images/top.jpg' alt="SGB logo"> </div>
#
# <h1 align='center'> TALLER “PROGRAMACIÓN ORIENTADA A LA BIOLOGÍA”</h1>
# <h3 align='center'>(En el marco del II CONCURSO “BIOINFORMÁTICA EN EL AULA”)</h3>
#
# La bioinformática es una disciplina científica destinada a la aplicación de métodos computacionales al análisis de datos biológicos, para poder contestar numerosas preguntas. Las tecnologías computacionales permiten, entre otras cosas, el análisis en plazos cortos de gran cantidad de datos (provenientes de experimentos, bibliografía, bases de datos públicas, etc), así como la predicción de la forma o la función de las distintas moléculas, o la simulación del comportamiento de sistemas biológicos complejos como células y organismos.
#
# La bioinformática puede ser pensada como una herramienta en el aprendizaje de la biología: su objeto de trabajo son entidades biológicas (ADN, proteínas, organismos completos, sus poblaciones, etc.) que se analizan con métodos que permiten “visualizar” distintos procesos de la naturaleza. Así, la bioinformática aporta una manera clara y precisa de percibir los procesos biológicos, y acerca a los estudiantes herramientas que integran múltiples conocimientos (lógico-matemático, biológico, físico, estadístico) generando un aprendizaje significativo y envolvente.
# <p style="font-family: Arial; font-size:1.75em;color:#2462C0; font-style:bold"><br> RETO I: ¿Podés descubrir y anotar el orden en que se ha ejecutado cada operación? </p>
# #### ¡Un momento, cerebrito!
# Una variable es un espacio para almacenar datos modificables, en la memoria de tu computadora. Es decir, le damos un nombre a un conjunto de “cosas” y una vez declarada esa variable, Python recordará que contiene las cosas que le hayamos asignado. Las variables se utilizan en todos los lenguajes de programación. En Python, una variable se define con la sintaxis:
# - **nombre** de la variable = **valor** de la variable
# Definamos entonces algunas variables:
a = 5
b = 3
c = 'hola'
# Como verás, una variable puede contener números, textos, lista de cosas, etc. En el caso de las letras o palabras, siempre debemos escribirlas entre comillas para que Python las considere como texto.
# Ahora le preguntaremos a Python cuánto vale **b**:
print(b)
# Veremos que efectivamente se ha guardado en memoria que b es igual a 3. En Python podemos reescribir una variable, es decir asignarle un nuevo contenido, simplemente declarándola de nuevo:
b = 500
print(b)
# Además, podemos asignar a más de una variable el mismo valor:
d = 500
print(b)
print(d)
# <p style="font-family: Arial; font-size:1.75em;color:#2462C0; font-style:bold"><br>RETO II: Creá una variable llamada doble, que sea el doble de la suma entre a y b. </p>
# **¿Y qué con eso?**
#
# Muy interesante esto de las variables, pero ¿para qué sirve? Una utilidad obvia es almacenar en memoria datos que nosotros seguro olvidaremos, pero que podríamos llegar a necesitar más adelante. Una variable también nos podría ayudar a guardar datos que aún no conocemos, pero que querríamos modificar o utilizar una vez adquiridos.
#
# Por ejemplo, nos gustaría crear un programa que salude al que lo usa. Como no sabemos a priori quién va a usar nuestro programa, podemos consultarle primero su nombre y luego saludarlo. Para ello sería útil guardarnos el dato de quién es la persona:
input('Decime tu nombre, por favor! ')
# Esta función **input** le consulta al usuario algo, lo que escribamos entre paréntesis y comillas, y espera la respuesta. Verás que una vez ejecutada la línea no vuelve a aparecer el prompt hasta que no ingresemos alguna palabra y le demos **Enter**. Ahora bien ¿podrías armar un programa que salude a quien lo use? Pensemos juntos qué pasos debe tener el programa:
# 1) Lo primero que deberíamos hacer es preguntarle su nombre al usuario y almacenarlo en una variable. ¡Hagámoslo!
nombre_de_usuario = input('Decime tu nombre, por favor!' )
# 2) Luego podemos imprimir un mensaje en pantalla que contenga el mensaje deseado:
print ('Hola ' + nombre_de_usuario + '¡Bienvenido a mi programa!')
# Muy probablemente, si lo hiciste solo, tu programa no sea exactamente igual al mío. ¡Existen muchas formas de lograr un mismo resultado! Así pues, si tu programa saluda a la persona que lo usa lo has hecho bien, aún cuando no sea de lo más estético el saludo.
#
# En el ejemplo anterior usamos un “truco” para imprimir el saludo, que puede serte muy útil de ahora en más: en Python podemos unir palabras o textos para generar frases o textos más largos simplemente sumándolas (+).
# **Nada es mejor, nada es igual… Bueno igual puede ser!**
#
# Existen también formas de comparar dos variables, lo que se conoce como operadores relacionales. Podemos saber si dos variables son iguales (==), o si son distintas (!=), o si una es mayor que la otra (>). Por ejemplo:
edad_lola = 13
edad_ana = 32
print(edad_lola == edad_ana)
# **¿Qué resultado obtenés al comparar dos variables?**
#
# Sí, los operadores relacionales no devuelven valores numéricos, sino que nos afirman (True) o rechazan (False) la hipótesis que hayamos puesto a prueba. En nuestro ejemplo la hipótesis es que la edad de Ana es igual a la de Lola y por eso Python nos devuelve False.
#
# Los operadores relacionales que se pueden usar en Python son:
# Símbolo | Descripción
# ------------ | -------------
# == | Igual
# != | Distinto
# < | Menor
# > | Mayor
# **Una palabra no dice nada…**
#
# En programación al texto se le llama **‘string’**. Este tipo de datos no es más que una cadena de caracteres, así como una palabra se puede entender como una cadena de letras. Un string no necesariamente tiene que tener sentido.
#
# En Python las cadenas se definen escribiendo los caracteres entre comillas (simples o dobles, indistintamente). A una variable se le puede asignar una cadena de la siguiente forma:
cadena = 'este es un ejemplo de cadena'
print(cadena)
# Podemos imprimir una cadena junto con el valor de una variable (un número u otra cadena) utilizando el marcador **%s**. Este símbolo marca el lugar donde va a incorporarse el texto de la variable. Por ejemplo:
mi_texto = 'Hola %s'
print(mi_texto %'Ana')
# Probemos otro ejemplo, esta vez con números, y sin definir antes una variable
print('El resultado de la cuenta es %s' %5)
# Podemos operar sumando y multiplicando cadenas del mismo modo en que operamos con números.
a = 'Hola '
b = 'chicos'
print(a+b)
# ¿Qué pasa si a una cadena le sumamos un número?
#
# Probá hacer **print(a + 5)** a ver qué pasa
print(a+5)
#El error nos indica que no podemos sumar cadenas de textos con números enteros.
# Las cadenas pueden ser comparadas con los operadores relacionales que vimos antes. Así, podemos saber entonces si dos cadenas son distintas o no lo son (tené en cuenta que Python distingue entre mayúsculas y minúsculas):
palabra = 'si'
lo_mismo = 'si'
print(palabra == lo_mismo)
# ¿Qué pasa si el contenido de la variable lo_mismo comienza con mayúsculas?
palabra = 'si'
lo_mismo = 'Si'
print(palabra == lo_mismo)
# <p style="font-family: Arial; font-size:1.75em;color:#2462C0; font-style:bold"><br>RETO III: </p>
# Digamos que el **ADN** no es más que un mensaje en clave, que debe ser descifrado o interpretado para la síntesis de proteínas. El mensaje está escrito por una secuencia determinada de **4 nucleótidos** distintos representados por las letras **A, T, G y C**. Dentro de la célula, el mensaje es transportado por otra molécula, el **ARN**, muy similar al **ADN** pero con `U en vez de T. `
#
# En este mensaje, cada triplete o grupo de tres letras del **ARN** se denomina codón, y cada aminoácido de las proteínas está codificado por uno o varios codones. Así por ejemplo el codón **‘AUG’** codifica para el **aminoácido Metionina**, el codón **‘AAA’** para Lisina, el codón **‘CUA’** para **Leucina**, etc.
#
# ##### ¿Podrías escribir una cadena de ARN que codifique para el péptido (es decir, la cadena corta de aminoácidos) ‘Met-Lis-Lis-Lis-Leu-Leu-Met’ combinando las variables met = ‘AUG’ , lis = ‘AAA’ y leu = ‘CUA’ utilizando operadores matemáticos?
#
# ¡Compartinos tu código en nuestro grupo de Facebook [‘Talleres de programación Orientada a la Biologia - SBG_UNQ’!](https://www.facebook.com/groups/1854914101467222/)
# **Fetas de texto: dame doscientos!**
#
# En Python podemos saber qué caracteres o subpartes conforman una cadena o string. Python le asigna a cada caracter de una cadena un número de posición. El primer carácter es la posición cero (¡sí, cero!) y las posiciones aumentan de a una hasta el fin de la cadena. Por ejemplo en la cadena a = ‘hola’, la ‘h’ tiene asignada la posición cero, la ‘o’ la posición uno, la ‘l’ la dos y la ‘a’ la tres. Por ejemplo, si quisiéramos saber cuál es el primer caracter de la cadena, hacemos referencia al caracter de la posición cero de la misma escribiendo el nombre de la variable seguida de la posición que nos interesa, escrita entre corchetes:
a = 'Hola mundo'
print(a[0])
# Podríamos tomar solo un segmento de la cadena, indicando entre corchetes desde qué carácter hasta qué carácter, separado por dos puntos.
print(a[0:1])
# En este caso, ¿cuántos caracteres se imprimirán? ¿Cómo hacemos para imprimir la palabra ‘Hola’ completa?
print(a[:])
# **Chamuyo: el arte de manipular palabras/cadenas.**
#
# Existen muchas funciones útiles para manipular cadenas. Cómo dijimos antes, a Python no le dan lo mismo las mayúsculas que las minúsculas. Existen funciones muy útiles para interconvertir unas en otras:
apellido = 'velez'
apellido.upper()
# ¿Y si queremos minúsculas?
nombre = 'ANA'
nombre.lower()
# Podemos conocer la longitud de una cadena (es decir de cuántos caracteres está formada) utilizando la función len():
apellido = 'velez'
len(apellido)
# También podemos conocer la cantidad de veces que aparece un dado carácter en una cadena utilizando la función **count()**:
apellido = 'velez'
apellido.count('e')
# Otra función muy útil para la manipulación de cadenas es replace(). Esta función nos permite reemplazar un tipo de carácter por otro. Por ejemplo:
apellido = 'velez'
apellido.replace('e','a')
# Cómo podés ver, la forma de reemplazar un carácter por otro es escribir entre los paréntesis de la función **replace()** el carácter que queremos reemplazar, entre comillas y separado por coma del carácter por el cual lo queremos reemplazar, también entre comillas.
| 1_retos_taller_v1_2018.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="ftyfzMxYeLVM" outputId="a32dcf8b-d27c-4ca4-98f6-438b2c80cebb" colab={"base_uri": "https://localhost:8080/", "height": 86}
# import the necessary packages
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from tensorflow.keras.applications import MobileNetV2
from tensorflow.keras.layers import AveragePooling2D
from tensorflow.keras.layers import Dropout
from tensorflow.keras.layers import Flatten
from tensorflow.keras.layers import Dense
from tensorflow.keras.layers import Input
from tensorflow.keras.models import Model
from tensorflow.keras.optimizers import Adam
from tensorflow.keras.applications.mobilenet_v2 import preprocess_input
from tensorflow.keras.preprocessing.image import img_to_array
from tensorflow.keras.preprocessing.image import load_img
from tensorflow.keras.utils import to_categorical
from sklearn.preprocessing import LabelBinarizer
from sklearn.model_selection import train_test_split
from sklearn.metrics import classification_report
from imutils import paths
import matplotlib.pyplot as plt
import numpy as np
import os
# initialize the initial learning rate, number of epochs to train for,
# and batch size
INIT_LR = 1e-4
EPOCHS = 20
BS = 32
DIRECTORY = r"/content/mask detection"
CATEGORIES = ["mask", "unmask"]
# grab the list of images in our dataset directory, then initialize
# the list of data (i.e., images) and class images
print("[INFO] loading images...")
data = []
labels = []
for category in CATEGORIES:
path = os.path.join(DIRECTORY, category)
for img in os.listdir(path):
img_path = os.path.join(path, img)
image = load_img(img_path, target_size=(224, 224))
image = img_to_array(image)
image = preprocess_input(image)
data.append(image)
labels.append(category)
# perform one-hot encoding on the labels
lb = LabelBinarizer()
labels = lb.fit_transform(labels)
labels = to_categorical(labels)
data = np.array(data, dtype="float32")
labels = np.array(labels)
(trainX, testX, trainY, testY) = train_test_split(data, labels,
test_size=0.20, stratify=labels, random_state=42)
# construct the training image generator for data augmentation
aug = ImageDataGenerator(
rotation_range=20,
zoom_range=0.15,
width_shift_range=0.2,
height_shift_range=0.2,
shear_range=0.15,
horizontal_flip=True,
fill_mode="nearest")
# load the MobileNetV2 network, ensuring the head FC layer sets are
# left off
baseModel = MobileNetV2(weights="imagenet", include_top=False,
input_tensor=Input(shape=(224, 224, 3)))
# construct the head of the model that will be placed on top of the
# the base model
headModel = baseModel.output
headModel = AveragePooling2D(pool_size=(7, 7))(headModel)
headModel = Flatten(name="flatten")(headModel)
headModel = Dense(128, activation="relu")(headModel)
headModel = Dropout(0.5)(headModel)
headModel = Dense(2, activation="softmax")(headModel)
# place the head FC model on top of the base model (this will become
# the actual model we will train)
model = Model(inputs=baseModel.input, outputs=headModel)
# loop over all layers in the base model and freeze them so they will
# *not* be updated during the first training process
for layer in baseModel.layers:
layer.trainable = False
# compile our model
print("[INFO] compiling model...")
opt = Adam(lr=INIT_LR, decay=INIT_LR / EPOCHS)
model.compile(loss="binary_crossentropy", optimizer=opt,
metrics=["accuracy"])
# + id="u0KlSGH0eYGm" outputId="b44ed959-3972-4803-fcf6-b86e9eee2f57" colab={"base_uri": "https://localhost:8080/", "height": 916}
#
# train the head of the network
print("[INFO] training head...")
# H = model.fit(
# aug.flow(trainX, trainY, batch_size=BS),
#steps_per_epoch=len(trainX) // BS,
#validation_data=(testX, testY),
#validation_steps=len(testX) // BS,epochs= EPOCHS )
H = model.fit(
aug.flow(trainX, trainY, batch_size=BS),epochs= EPOCHS)
# make predictions on the testing set
print("[INFO] evaluating network...")
predIdxs = model.predict(testX, batch_size=BS)
# for each image in the testing set we need to find the index of the
# label with corresponding largest predicted probability
predIdxs = np.argmax(predIdxs, axis=1)
# show a nicely formatted classification report
print(classification_report(testY.argmax(axis=1), predIdxs,
target_names=lb.classes_))
# serialize the model to disk
print("[INFO] saving mask detector model...")
model.save("mask_detector.model", save_format="h5")
# plot the training loss and accuracy
N = EPOCHS
plt.style.use("ggplot")
plt.figure()
# + id="7Q1yLbk7howP" outputId="19efdf39-f731-4846-bbb8-d3dfb6ac3b80" colab={"base_uri": "https://localhost:8080/", "height": 298}
plt.plot(np.arange(0, N), H.history["loss"], label="train_loss")
# plt.plot(np.arange(0, N), H.history["val_loss"], label="val_loss")
plt.plot(np.arange(0, N), H.history["accuracy"], label="train_acc")
# plt.plot(np.arange(0, N), H.history["val_accuracy"],label="val_accuracy")
plt.title("Training Loss and Accuracy")
plt.xlabel("Epoch #")
plt.ylabel("Loss/Accuracy")
plt.legend(loc="lower left")
plt.savefig("plot.png")
# + id="42PPbL3FjTr3" outputId="d3ba10b2-d92a-4c4c-979d-f5309d2043c6" colab={"base_uri": "https://localhost:8080/", "height": 285}
from tensorflow.keras.preprocessing import image
x = image.load_img('/content/mask detection/unmask/0_0_anhu_0020.jpg',target_size=(trainX.shape[1:]))
plt.imshow(x)
# + id="bInTrnRYkHKx" outputId="7503d383-f2a4-4404-a045-1d938b54d546" colab={"base_uri": "https://localhost:8080/", "height": 285}
from tensorflow.keras.preprocessing import image
x = image.load_img('/content/mask detection/mask/0_0_≈˙◊¢ 2020-02-23 132400.png',target_size=(trainX.shape[1:]))
plt.imshow(x)
| mask.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Loading the data
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import json
# !pip install nltk
plt.rcParams["figure.figsize"] = (10,10)
data = json.load(open('xad_tags'))
len(data)
# -
# # How the raw data looks like ?
# +
for article in data[:2]:
print(article)
print(data[0].keys())
# -
# # Clean the data a little bit
# +
to_be_removed = ['welt_kompakt', 'die_welt', 'wams' 'print_politik',
'print' 'newsapp', 'deutschland', 'ausland', 'hamburg', 'wirtschaft']
for article in data:
categories_for_article = article['Categories']
text = article['Introtext']
if len(categories_for_article) == 1 and "." in categories_for_article[0]:
categories_for_article[0] = ''
if len(categories_for_article) >= 1:
for current_category in categories_for_article:
if current_category in to_be_removed:
categories_for_article.remove(current_category)
for index, category in enumerate(categories_for_article):
if "_" in category:
splitted = category.split("_")
categories_for_article[index] = splitted[1]
# +
from nltk.stem.snowball import SnowballStemmer
stopwords = {
"a": True,
"aber": True,
"ab": True,
"alle": True,
"allem": True,
"allen": True,
"aller": True,
"allerdings": True,
"alles": True,
"als": True,
"also": True,
"außerdem": True,
"am": True,
"an": True,
"ander": True,
"andere": True,
"anderem": True,
"anderen": True,
"anderer": True,
"anderes": True,
"anderm": True,
"andern": True,
"anders": True,
"auch": True,
"auf": True,
"aus": True,
"bei": True,
"beiden": True,
"bin": True,
"bis": True,
"bist": True,
"bald": True,
"beim": True,
"da": True,
"davon": True,
"damit": True,
"dann": True,
"das": True,
"damals": True,
"dass": True,
"dasselbe": True,
"dazu": True,
"daß": True,
"dein": True,
"dabei": True,
"deine": True,
"deinem": True,
"deinen": True,
"deiner": True,
"deines": True,
"dem": True,
"demselben": True,
"den": True,
"denn": True,
"denselben": True,
"der": True,
"derer": True,
"derselbe": True,
"derselben": True,
"dadurch": True,
"des": True,
"desselben": True,
"dessen": True,
"dich": True,
"die": True,
"dies": True,
"dafür": True,
"diese": True,
"denen": True,
"dieselbe": True,
"deswegen": True,
"dieselben": True,
"diesem": True,
"diesen": True,
"dieser": True,
"dieses": True,
"dir": True,
"doch": True,
"darf": True,
"datum": True,
"darunter": True,
"dort": True,
"du": True,
"durch": True,
"deutschen": True,
"ein": True,
"eine": True,
"einem": True,
"einen": True,
"einer": True,
"eines": True,
"einig": True,
"einige": True,
"einigem": True,
"einigen": True,
"einiger": True,
"einiges": True,
"erst": True,
"ende": True,
"einmal": True,
"eigentlich": True,
"er": True,
"es": True,
"etwas": True,
"etwa": True,
"euch": True,
"euer": True,
"eure": True,
"euro": True,
"eurem": True,
"endlich": True,
"euren": True,
"eurer": True,
"eures": True,
"für": True,
"gegen": True,
"gibt": True,
"ganz": True,
"gut": True,
"gewesen": True,
"gar": True,
"gerade": True,
"geworden": True,
"geht": True,
"hab": True,
"habe": True,
"haben": True,
"hat": True,
"hätte": True,
"hätten": True,
"hatte": True,
"hatten": True,
"hier": True,
"hin": True,
"hinter": True,
"ich": True,
"ihm": True,
"ihn": True,
"ihnen": True,
"ihr": True,
"ihre": True,
"ihrem": True,
"ihren": True,
"immer": True,
"ihrer": True,
"ihres": True,
"im": True,
"in": True,
"indem": True,
"ins": True,
"ist": True,
"jede": True,
"jedem": True,
"jeden": True,
"jeder": True,
"jedes": True,
"jene": True,
"jenem": True,
"jenen": True,
"jener": True,
"jenes": True,
"jetzt": True,
"kann": True,
"kein": True,
"keine": True,
"keinem": True,
"keinen": True,
"konnte": True,
"keiner": True,
"keines": True,
"können": True,
"könnte": True,
"klein": True,
"kürze": True,
"kurz": True,
"kürzem": True,
"mal": True,
"machen": True,
"man": True,
"manche": True,
"manchem": True,
"manchen": True,
"mancher": True,
"manches": True,
"mein": True,
"menschen": True,
"millionen": True,
"meine": True,
"meinem": True,
"meinen": True,
"meisten": True,
"meiner": True,
"müssen": True,
"mehr": True,
"meines": True,
"mann": True,
"mich": True,
"mir": True,
"mit": True,
"muss": True,
"musste": True,
"mutter": True,
"monat": True,
"land": True,
"trotzdem": True,
"liegt": True,
"nach": True,
"neuen": True,
"nicht": True,
"nichts": True,
"noch": True,
"nun": True,
"nur": True,
"pro": True,
"ob": True,
"oft": True,
"öfter": True,
"oder": True,
"ohne": True,
"sehr": True,
"sein": True,
"seit": True,
"seine": True,
"sagt": True,
"sagte": True,
"seinem": True,
"seinen": True,
"seiner": True,
"seines": True,
"selbst": True,
"schon": True,
"sich": True,
"sie": True,
"sind": True,
"so": True,
"solche": True,
"solchem": True,
"sogar": True,
"solchen": True,
"solcher": True,
"solches": True,
"soll": True,
"sollte": True,
"sollen": True,
"steht": True,
"sei": True,
"sondern": True,
"sonst": True,
"stadt": True,
"tagen": True,
"trainer": True,
"tages": True,
"rund": True,
"um": True,
"und": True,
"uns": True,
"unser": True,
"unsere": True,
"prozent": True,
"president": True,
"unserem": True,
"unseren": True,
"unserer": True,
"unseres": True,
"unter": True,
"viel": True,
"viele": True,
"vater": True,
"vom": True,
"von": True,
"vor": True,
"war": True,
"waren": True,
"warst": True,
"was": True,
"weg": True,
"weil": True,
"weiter": True,
"welche": True,
"welchem": True,
"welchen": True,
"welcher": True,
"welches": True,
"wenn": True,
"wenig": True,
"werde": True,
"werden": True,
"wie": True,
"wieder": True,
"will": True,
"wir": True,
"wird": True,
"wirst": True,
"warum": True,
"wegen": True,
"wurde": True,
"wo": True,
"wäre": True,
"wollen": True,
"wollte": True,
"während": True,
"wer": True,
"wem": True,
"würde": True,
"würden": True,
"zu": True,
"zum": True,
"zur": True,
"zwar": True,
"zwischen": True,
"über": True,
"jahr": True,
"jahre": True,
"jahren": True,
"frau": True,
"frauen": True,
}
stemmer = SnowballStemmer("german")
keys = ['Introtext', 'Categories']
df = pd.DataFrame(data, columns=keys)
df['Introtext'].replace('', np.nan, inplace=True)
df['Categories'].replace('', np.nan, inplace=True)
df.dropna(inplace=True)
def to_s(categories):
return ''.join(categories)
def to_lowercase(text):
return text.lower()
def remove_stopwords(text):
result = []
splitted = text.split(" ")
for word in splitted:
if word not in stopwords.keys():
result.append(" " + stemmer.stem(word))
return "".join(result)
df['Categories'] = df['Categories'].apply(to_s)
filtered = df.groupby('Categories').filter(lambda x: len(x) >= 250)
df['Introtext'] = df['Introtext'].apply(to_lowercase)
df['Introtext'] = df['Introtext'].apply(remove_stopwords)
pd.value_counts(filtered['Categories']).plot(kind='barh')
#g.apply(lambda x: x.sample(g.size().min()).reset_index(drop=True))
# -
# # Prepare training and test data
# +
train_size = int(len(filtered) * .8)
train_articles = filtered['Introtext'][:train_size]
train_categories = filtered['Categories'][:train_size]
test_articles = filtered['Introtext'][train_size:]
test_categories = filtered['Categories'][train_size:]
num_labels = pd.value_counts(train_categories).size
print(train_articles.shape)
print(train_articles.ndim)
# -
# # Tokenize and prepare to data to feed into the neural network
# +
from keras.preprocessing.text import Tokenizer
vocabulary_size = 20000
tokenize = Tokenizer(num_words=vocabulary_size, lower=True)
tokenize.fit_on_texts(train_articles)
x_train = tokenize.texts_to_matrix(train_categories)
# +
from sklearn import preprocessing
encoder = preprocessing.LabelBinarizer()
encoder.fit(train_categories)
y_train = encoder.transform(train_categories)
y_test = encoder.transform(test_categories)
# -
# # Train...
# +
from keras.models import Sequential
from keras.layers import Dense, Activation
model = Sequential()
model.add(Dense(512, input_shape=(vocabulary_size,)))
model.add(Activation('relu'))
model.add(Dense(num_labels))
model.add(Activation('softmax'))
batch_size = 64
model.compile(loss='categorical_crossentropy',
optimizer='adam',
metrics=['accuracy'])
history = model.fit(x_train, y_train,
batch_size=batch_size,
epochs=3,
verbose=1,
validation_split=0.1)
# -
# # Let's try it out
# +
text_labels = encoder.classes_
for ctr in range(25):
prediction = model.predict(np.array([x_train[ctr]]))
predicted_label = text_labels[np.argmax(prediction[0])]
print(test_articles.iloc[ctr][:80], "..")
print('Real category: ' + test_categories.iloc[ctr])
print("Predicted category: " + predicted_label)
# -
| 02_deep_learning/classify_articles.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import nltk
import pandas as pd
import numpy as np
import re
import gensim
news_df = pd.read_csv("dataset/Combined_News_DJIA.csv")
stocks_df = pd.read_csv("dataset/DJIA_table.csv").iloc[::-1] # reverse
news_stocks_df = news_df.merge(stocks_df, on="Date")
news_stocks_df.head(20)
stopwords = set(nltk.corpus.stopwords.words('english'))
def combine_news(row, tokenizer, remove_stopwords=True):
top_n_news = 25
all_titles = []
for i in range(1, top_n_news+1):
col_name = "Top{}".format(i)
title = row[col_name]
if type(title) != str:
continue
# Remove everything except a-z A-Z
title = re.sub("[^a-zA-Z]", " ", title)
title = title[1:].strip().lower()
# Tokenize
words = title.split()
if remove_stopwords:
words = title.split()
words = [w for w in words if not w in stopwords]
title = " ".join(words)
all_titles.append(title)
all_titles = " ".join(all_titles)
return all_titles
news_stocks_df["All_news"] = news_stocks_df.apply(
lambda row: combine_news(row, tokenizer),
axis=1)
news_stocks_df.to_csv("dataset/news_stocks.csv")
| .ipynb_checkpoints/Preprocess-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 「標準地域コード」の助けを借りつつ、「全国地方公共団体コード」の RDF を生成する。
import pandas as pd
import numpy as np
import rdflib
import datetime
import re
import lxml.html
import rdflib
import math
import os.path
import urllib.parse
import urllib.request
import jaconv
from distutils.version import LooseVersion
from rdflib.namespace import RDF, RDFS, DCTERMS, XSD, SKOS
SAC = rdflib.Namespace("http://data.e-stat.go.jp/lod/sac/")
SACS = rdflib.Namespace("http://data.e-stat.go.jp/lod/terms/sacs#")
JITI = rdflib.Namespace("http://hkwi.github.com/denshijiti/#")
JITIS = rdflib.Namespace("http://hkwi.github.com/denshijiti/terms#")
IC = rdflib.Namespace("http://imi.go.jp/ns/core/rdf#")
# e-stat「標準地域コード」を取り込む。http://data.e-stat.go.jp/lodw/rdfschema/downloadfile/
estat = rdflib.Graph()
estat.load("http://data.e-stat.go.jp/lodw/download/rdfschema/StandardAreaCode.ttl", format="turtle")
# 現在の excel を収集する。
# +
# xmlns:gcse="uri:google-did-not-provide-a-real-ns"
url = "http://www.soumu.go.jp/denshijiti/code.html"
p = lxml.html.parse(urllib.request.urlopen(url), base_url=url)
r = p.getroot()
r.make_links_absolute()
for h in r.xpath("//h3"):
if h.text.find("都道府県コード") >= 0:
for l in h.xpath("following-sibling::node()//li"):
if not l.text or l.text.find("都道府県") < 0:
continue
if l.text.find("改正一覧") >= 0:
for a in l.xpath("self::node()//a/@href"):
if a.lower().endswith(".xls"):
clist_hist = a
break
else:
for a in l.xpath("self::node()//a/@href"):
if a.lower().endswith(".xls"):
clist = a
break
assert clist
assert clist_hist
assert clist_hist != clist
# -
# まず過去履歴を集める
# +
x = pd.read_excel(clist_hist, skiprows=1, header=[0,1,2])
x = pd.read_excel("http://www.soumu.go.jp/main_content/000562731.xls", skiprows=1, header=[0,1,2])
def offset():
for i,c in enumerate(x.columns):
for j,e in enumerate(c):
if "都道府県" in e:
return i,j
x.columns=["a","b","c","d","e"][:offset()[0]] + [
"都道府県名",
"改正前コード","改正前市区町村名","改正前市区町村名ふりがな",
"改正区分","改正年月日",
"改正後コード","改正後市区町村名","改正後市区町村名ふりがな",
"事由等"
]
x.index = range(len(x.index))
# +
asub = x["改正前市区町村名"].str.extract(r"^[\((](.*)[\))]$", expand=False)
bsub = x["改正後市区町村名"].str.extract(r"^[\((](.*)[\))]$", expand=False)
ks = ["都道府県名", "改正前コード", "改正区分", "改正年月日", "改正後コード"]
x2 = x[asub.isnull() & bsub.isnull() & x[ks].notnull().any(axis=1)].assign(asub=None, bsub=None)
for i,d in asub[asub.notnull()].iteritems():
x2.loc[i-1, "asub"] = d
for i,d in asub[bsub.notnull()].iteritems():
x2.loc[i-1, "bsub"] = d
x2.index = range(len(x2.index))
# -
for ri,r in x2.iterrows():
for ci,c in r.iteritems():
if c=="〃":
x2.loc[ri, ci] = x2.loc[ri-1, ci]
cids = []
cid = 0
t = x2.assign(u=lambda o: o["都道府県名"].notnull())
for ri, r in x2.iterrows():
if not isinstance(r["都道府県名"], float):
cid += 1
cids.append(cid)
dts = []
t = pd.concat([x2, pd.Series(cids, name="cid"), pd.Series(None, name="date")], axis=1)
for i in range(cid):
u = t[(t["cid"]==i+1) & t["改正年月日"].notnull()]
assert len(set(u["改正年月日"]))==1, u.to_csv()
for d in set(u["改正年月日"]):
if isinstance(d, datetime.datetime):
dt = d.date()
elif isinstance(d, datetime.date):
dt = d
else:
m = re.match(r"H(\d{2})\.(\d+)\.(\d+)", d)
dt = datetime.date(2000+int(m.group(1))-12, int(m.group(2)), int(m.group(3)))
for ri,r in t[(t["cid"]==i+1)].iterrows():
t.loc[ri,"date"] = dt
# +
g = rdflib.Graph()
g.bind("ic", IC)
g.bind("sac", SAC)
g.bind("skos", SKOS)
g.bind("jiti", JITI)
g.bind("jitis", JITIS)
g.bind("dcterms", DCTERMS)
for cls in ("StandardAreaCode", "CodeSet", "CodeChangeEvent"):
g.add((JITIS[cls], RDF.type, RDFS["Class"]))
g.add((JITIS["StandardAreaCode"], RDFS.subClassOf, IC["住所"]))
def get_code(code_like):
code = "%06d" % int(code_like)
assert code_checksum(code) == code[5:], "code %s checksum error" % code
return code
def code_checksum(code):
s = np.array([int(s) for s in code[:5]]) * np.array([6,5,4,3,2])
return str(11 - s.sum() % 11)[-1]
def get_code_id(url_like):
m = re.search(r"C(?P<code>\d{5})-(?P<ymd>\d{8})$", url_like)
assert m, "code_id %s format error" % url_like
return m.group(0)
def sacq(code5):
return estat.query('''
PREFIX dcterms: <http://purl.org/dc/terms/>
PREFIX sacs: <http://data.e-stat.go.jp/lod/terms/sacs#>
PREFIX rdfs: <http://www.w3.org/2000/01/rdf-schema#>
SELECT ?s ?n ?d WHERE {
?s a sacs:StandardAreaCode ;
rdfs:label ?n ;
dcterms:identifier "%s" ;
dcterms:issued ?d .
FILTER ( LANG(?n) = "ja" )
}
ORDER BY DESC(?d)
''' % code5)
class Code(object):
codes = {}
@classmethod
def singleton(cls, code_id):
obj = cls.codes.get(code_id)
if obj is None:
cls.codes[code_id] = obj = cls(code_id)
return obj
def __init__(self, code_id):
m = re.match(r"C(?P<code>\d{5})-(?P<ymd>\d{8})$", code_id)
assert m, "code %s format error" % code_id
self.code = m.group("code")
self.csum = code_checksum(self.code)
self.ymd = datetime.datetime.strptime(m.group("ymd"), "%Y%m%d").date()
# e-Stat LOD ベースで採番する
self.sac = JITI[code_id]
g.add((self.sac, RDF["type"], JITIS["StandardAreaCode"]))
g.add((self.sac, SKOS["closeMatch"], SAC[code_id]))
g.add((self.sac, DCTERMS["identifier"], rdflib.Literal(self.code)))
g.add((self.sac, DCTERMS["issued"], rdflib.Literal(self.ymd.strftime("%Y-%m-%d"), datatype=XSD.date)))
# 6桁コードでリンクできるようにする
g.add((self.sac, JITIS["code"], rdflib.Literal(self.code+self.csum)))
# 共通語彙基盤 住所IEP : 現状は blank node での出力を強制する
pref = rdflib.BNode()
g.add((pref, RDF.type, IC["コード型"]))
g.add((pref, IC["識別値"], rdflib.Literal(self.code[:2], datatype=XSD.string)))
g.add((self.sac, IC["都道府県コード"], pref))
ward = rdflib.BNode()
g.add((ward, RDF.type, IC["コード型"]))
g.add((ward, IC["識別値"], rdflib.Literal(self.code[2:]+self.csum, datatype=XSD.string)))
g.add((self.sac, IC["市区町村コード"], ward))
def set_name(self, name, kana):
assert name != "同左"
assert kana != "同左"
name = name.strip()
kana = kana.strip()
# e-Stat LOD 準拠
g.add((self.sac, RDFS["label"], rdflib.Literal(name, lang="ja")))
g.add((self.sac, RDFS["label"], rdflib.Literal(kana, lang="ja-hrkt")))
ku = None
sub = int(self.code[2:])
if self.code[2:] == "000":
g.add((self.sac, JITIS["type"], JITIS["都道府県"]))
elif self.code[:3] == "131":
g.add((self.sac, JITIS["type"], JITIS["特別区"]))
elif self.code[2:] == "100":
g.add((self.sac, JITIS["type"], JITIS["指定都市"]))
elif self.code[2] == "1":
if name.endswith("市"):
g.add((self.sac, JITIS["type"], JITIS["指定都市"]))
else:
g.add((self.sac, JITIS["type"], JITIS["指定都市の区"]))
for i in range(sub // 10 * 10, 90, -10):
if ku:
break
for s,n,d in sacq("%s%03d" % (self.code[:2], i)):
if d.value <= self.ymd:
if n.endswith("市"):
ku = n
break
elif self.code[2] == "2":
g.add((self.sac, JITIS["type"], JITIS["市"]))
elif self.code[:2] == "01":
if sub % 30:
gr = "01%03d" % (sub // 30 * 30)
g.add((self.sac, JITIS["group"], rdflib.Literal(gr + code_checksum(gr))))
elif self.code[:2] == "47":
if 340 < sub and sub < 370:
gr = "47340"
g.add((self.sac, JITIS["group"], rdflib.Literal(gr + code_checksum(gr))))
elif 370 < sub and sub < 380:
gr = "47370"
g.add((self.sac, JITIS["group"], rdflib.Literal(gr + code_checksum(gr))))
elif sub % 20:
gr = "47%03d" % (sub // 20 * 20)
g.add((self.sac, JITIS["group"], rdflib.Literal(gr + code_checksum(gr))))
elif sub % 20:
gr = "%s%03d" % (self.code[:2], sub // 20 * 20)
g.add((self.sac, JITIS["group"], rdflib.Literal(gr + code_checksum(gr))))
if self.code[2:] == "000":
g.add((self.sac, IC["都道府県"], rdflib.Literal(name, lang="ja")))
else:
if ku:
g.add((self.sac, IC["市区町村"], ku))
g.add((self.sac, IC["区"], rdflib.Literal(name, lang="ja")))
else:
g.add((self.sac, IC["市区町村"], rdflib.Literal(name, lang="ja")))
for s,n,d in sacq(self.code[:2]+"000"):
if d.value <= self.ymd:
g.add((self.sac, IC["都道府県"], n))
break
code_ids = []
last_date = None
ev = None
cid = -1
for ri, r in t.sort_values(["date","cid"]).iterrows():
last_date = r["date"]
if cid != r["cid"]:
dtstr = r["date"].strftime("%Y-%m-%d")
ev = rdflib.BNode("_:ev%s-%02d" % (dtstr,cid))
g.add((ev, RDF["type"], JITIS["CodeChangeEvent"]))
g.add((ev, DCTERMS["issued"], rdflib.Literal(dtstr, datatype=XSD.date)))
cid = r["cid"]
if not math.isnan(r["改正前コード"]):
tc = get_code(r["改正前コード"])
cids = [cid for c,cid in code_ids if c==tc]
code_id = None
if cids:
code_id = cids[-1]
else:
for s,n,d in sacq(tc[:5]):
if d.value < r["date"]:
code_id = get_code_id(s)
break
assert code_id
g.add((ev, JITIS["old"], JITI[code_id]))
Code.singleton(code_id).set_name(r["改正前市区町村名"], r["改正前市区町村名ふりがな"])
code = None
code_id = None
if r["改正後コード"]=="削除" or r["改正区分"] == "欠番":
pass
elif r["改正後コード"] == "同左":
if math.isnan(r["改正前コード"]):
raise Exception(r.to_csv())
code = get_code(r["改正前コード"])
code_id = "C%s-%s" % (code[:5], r["date"].strftime("%Y%m%d"))
else:
try:
code = get_code(r["改正後コード"])
code_id = "C%s-%s" % (code[:5], r["date"].strftime("%Y%m%d"))
except:
raise Exception(r.to_csv())
if code and code_id:
for t in g.triples((JITI[code_id], RDFS["label"], None)):
g.remove(t)
code_ids.append((code, code_id))
g.add((ev, JITIS["new"], JITI[code_id]))
name = r["改正後市区町村名"]
if name.strip()=="同左":
name = r["改正前市区町村名"]
kana = r["改正後市区町村名ふりがな"]
if isinstance(kana, float) or kana.strip()=="同左":
kana = r["改正前市区町村名ふりがな"]
# errata in xls row 1319
if isinstance(kana, float) and name=="西海市":
kana = "さいかいし"
Code.singleton(code_id).set_name(name, kana)
# -
# コードセットの登録。
# +
cs = []
pq = '''
PREFIX dcterms: <http://purl.org/dc/terms/>
SELECT ?s WHERE {
?s a <%s> ;
dcterms:identifier "%s" ;
dcterms:issued ?d .
}
ORDER BY DESC(?d)
'''
x = pd.read_excel(clist)
for ri,r in x.iterrows():
code = get_code(r["団体コード"])
ident = rdflib.Literal(code[:5])
code_id = None
for s, in g.query(pq % (JITIS["StandardAreaCode"], ident)):
code_id = get_code_id(s)
break
if code_id is None:
for s, in estat.query(pq % (SACS["StandardAreaCode"], ident)):
code_id = get_code_id(s)
break
name = r["市区町村名\n(漢字)"]
if isinstance(name, float) or not name.strip():
name = r["都道府県名\n(漢字)"]
kana = r["市区町村名\n(カナ)"]
if isinstance(kana, float) or not kana.strip():
kana = r["都道府県名\n(カナ)"]
assert code_id, code
Code.singleton(code_id).set_name(name, jaconv.kata2hira(jaconv.h2z(kana)))
assert code_id, code
cs.append(code_id)
if LooseVersion(pd.__version__) >= LooseVersion("0.21.0"):
x = pd.read_excel(clist, sheet_name=1, header=None)
else:
x = pd.read_excel(clist, sheetname=1, header=None)
for ri,r in x.iterrows():
code = get_code(r[0])
ident = rdflib.Literal(code[:5])
code_id = None
for s, in g.query(pq % (JITIS["StandardAreaCode"], ident)):
code_id = get_code_id(s)
break
if code_id is None:
# 「札幌市中央区」などが入っている。
# e-Stat では「中央区」が使われていて、揃えるために estat の値を引く
#name = r[1]
#kana = r[2]
name = kana = None
for s, in estat.query(pq % (SACS["StandardAreaCode"], ident)):
code_id = get_code_id(s)
for n in estat.objects(s, RDFS["label"]):
if n.language=="ja":
name = n
elif n.language=="ja-hrkt":
kana = n
Code.singleton(code_id).set_name(name, jaconv.kata2hira(jaconv.h2z(kana)))
assert code_id, code
cs.append(code_id)
# -
# 過去履歴を逆向きに遡ってコードセットを登録する。
# +
date = last_date
b = JITI["CS-" + date.strftime("%Y-%m-%d")]
g.add((b, rdflib.RDF["type"], JITIS["CodeSet"]))
g.add((b, DCTERMS["issued"], rdflib.Literal(dtstr, datatype=XSD.date)))
for c in cs:
g.add((b, DCTERMS["hasPart"], JITI[c]))
pq = '''
PREFIX xsd: <http://www.w3.org/2001/XMLSchema#>
PREFIX jitis: <http://hkwi.github.com/denshijiti/terms#>
PREFIX dcterms: <http://purl.org/dc/terms/>
SELECT ?v WHERE {
?s a jitis:CodeChangeEvent ;
dcterms:issued "%s"^^xsd:date ;
<%s> ?v .
}
'''
dts = g.query('''
PREFIX xsd: <http://www.w3.org/2001/XMLSchema#>
PREFIX jitis: <http://hkwi.github.com/denshijiti/terms#>
PREFIX dcterms: <http://purl.org/dc/terms/>
SELECT DISTINCT ?d WHERE {
?s a jitis:CodeChangeEvent ;
dcterms:issued ?d .
}
ORDER BY DESC(?d)
''')
for dt, in dts:
if dt.value < date: # 履歴と一覧が同期していないときのための安全
dtstr = date.strftime("%Y-%m-%d")
b = JITI["CS-" + dtstr]
g.add((b, rdflib.RDF["type"], JITIS["CodeSet"]))
g.add((b, DCTERMS["issued"], rdflib.Literal(dtstr, datatype=XSD.date)))
for c in cs:
g.add((b, DCTERMS["hasPart"], JITI[c]))
date = dt.value
for old, in g.query(pq % (JITIS["old"], dt.value.strftime("%Y-%m-%d"))):
code_id = get_code_id(old)
cs.append(code_id)
for new, in g.query(pq % (JITIS["new"], dt.value.strftime("%Y-%m-%d"))):
code_id = get_code_id(new)
try:
cs.remove(code_id)
except:
raise Exception(code_id)
# -
with open("code.ttl", "wb") as f:
g.serialize(destination=f, format="turtle")
| notebook/code.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# ## Mutation analysis
#
# This is a Jupyter notebook.
#
# To run all cells in the notebook use `Cell --> Run All`.
#
# To run cells one at a time click into the first code cell and key `Shift-Enter` in each cell in sequence.
#
# More information on Jupyter notebooks can be found
# [here](http://jupyter-notebook.readthedocs.io/en/stable/examples/Notebook/Running%20Code.html).
## Set the plot window sizes within the notebook (in inches)
options(repr.plot.width=8, repr.plot.height=4)
## Load the libraries
library(ggplot2)
library(reshape2)
# +
## Minimum number of mutations per gene to be included in analysis
opt.num = 3
## File arguments
fn.hist <- './user_data/UserDogData_Phenotype.csv' # User-provided
fn.muts <- './user_data/mutations_genesOnly.csv' # Created by the mutations pipeline (see shell scripts)
fn.peps <- './user_data/CMT_peps.csv' # PEP lists created in the expression pipeline. This is also Supp Table 1 in the manuscript
fn.pam50 <- NULL #set to NULL if you don't want to run, filename otherwise
## Filenames that don't change between users
data.dir <- './data/' # Working directory with the provided data files
fn.cosmic <- paste0(data.dir,'genes_COSMIC.csv') # COSMIC genes list, should download most recent version instead of using included one?
fn.pam50.genes <- paste0(data.dir,'PAM50_genes.csv') # This list will never change, no need to have as input
# max ratio cutoff for genes of interest used in figure 4
max.ratios.cutoff <- 0.15
# Optional: Compare mutation rates in PAM50 genes and subtypes (predicted subtypes generated by PAM50 notebook)
fn.pam50 <- './user_data/output/PAM50_dog.csv'
run.pam50 <- !is.null(fn.pam50)
# -
# ## Please comment or uncomment the following declarations as appropriate for your run of this notebook:
# +
# Synthetic and "canned" data for testing this notebook (uncomment if needed)
fn.hist <- "./synthetic_data/User_SampleData_Phenotype.csv"
fn.muts <- "./synthetic_data//User_SampleData_Mutations.csv"
# use a lower cutoff for synthetic data
#max.ratios.cutoff <- 0.07
# +
# Create an output directory
output.dir <- './user_data/results'
## If the output directory doesn't exist, create it
if(!dir.exists(opt$outdir)) {
print(paste('Creating output directory',opt$outdir))
system(paste('mkdir -p',opt$outdir))
}
# output files
mut.rates.file <- paste(output.dir,'Sample_Mut_Rates.pdf', sep='/')
density.plot.file <- paste(output.dir,'Sample_Mutation_Counts_Density.pdf', sep='/')
cosmic.mutations.file <- paste(output.dir,'COSMIC_Genes_Mutations.pdf', sep='/')
mutations.consistency.file <- paste(output.dir,'MutationConsistency.pdf', sep='/')
freq.mutations.file <- paste(output.dir,'FreqMutatedGenes_ClinicalCorrelations.csv', sep='/')
# -
## Make sure all of the required files exist - quit if any are missing
for( fn in c(fn.hist, fn.muts, fn.peps, fn.cosmic, fn.pam50.genes) ) {
if(!file.exists(fn)) { print(paste('ERROR: Unable to locate',fn)); quit(save='no',status=1) }
}
## Load the clinical data, extract dog IDs
print('Loading script data')
dat.hist <- read.table(fn.hist, sep=',', header=TRUE, row.names=1)
## Alphabetic IDs for each dog instead of numeric
## Generally don't need this- only CMTGA changes up the names halfway through
dat.hist$Patient <- as.character(dat.hist$Patient) # Ensure patient names are character strings for plotting consistency
## Load the PAM50 subtypes
## This file is created by PAM50_refactored.R, should just port straight over (don't need user to specify)
if(run.pam50) {
pam50 <- read.table(fn.pam50, sep=',', row.names=1)
print('PAM50 subtype counts per patient:')
print(table(pam50[,1], dat.hist[rownames(pam50),'Patient']))
dat.hist$PAM50 <- pam50[rownames(dat.hist),1]
}
## Load the list of COSMIC genes
genes.cosmic <- rownames(read.table(fn.cosmic, sep=',', header=TRUE, row.names=1))
# +
## Load the mutations data, make 0/1 calls instead of # calls per gene
dat <- read.table(fn.muts, sep=',', header=TRUE, row.names=1, check.names=FALSE)
## Create matrix of binary mutation calls
dat.bin <- dat
dat.bin[dat.bin>0] <- 1
# +
## Create 2 matrices: Benign and Malignant samples
print('Creating the mutations matrices.')
dat.m <- dat.bin[,rownames(dat.hist)[dat.hist$Hist=='M']]
dat.b <- dat.bin[,rownames(dat.hist)[dat.hist$Hist=='B']]
## Aggregate mutation calls by patient rather than by sample
dat.m <- t(aggregate(t(dat.m), by=list(dat.hist[colnames(dat.m),'Patient']), FUN=sum))
colnames(dat.m) <- dat.m[1,]
dat.m <- dat.m[-1,]
dat.b <- t(aggregate(t(dat.b), by=list(dat.hist[colnames(dat.b),'Patient']), FUN=sum))
colnames(dat.b) <- dat.b[1,]
dat.b <- dat.b[-1,]
## Convert from character to numeric
class(dat.b) <- 'numeric'
class(dat.m) <- 'numeric'
# For now we don't care about # samples mutated in each gene per patient, just that at least 1 sample is mutated
# So set >1 values to 1
dat.m[dat.m>0] <- 1
dat.b[dat.b>0] <- 1
# +
#########################################
## Color Palette for manuscript
#####################################
print('Setting color palettes.')
cols <- c('#9DC7D8','#7FA1BE','#EBDA8C','#01B3CA','#4F6E35','#965354','#7DD1B9','#808040','#C6CBCE','#1D4E88','#C78C6C','#F69256','#D2B29E','#8B868C','#E38691','#B490B2') # All colors in palette
cols.hist <- c('#7DD1B9','#EBDA8C','#965354') # order = healthy, benign, malignant
cols.peps <- c('#7FA1BE','#F69256','#E38691') # order = tumor, adenoma, carcinoma
# +
## Create histogram of mutations per sample per histology.
print('Generating mutation histogram')
## Create data frame for patient summaries, converting to alphabet patient IDs instead of numeric
mut.rates <- data.frame(Muts=apply(dat.bin, 2, sum), Hist=dat.hist[colnames(dat.bin),'Hist'], Dog=dat.hist[colnames(dat.bin),'Patient'], Sample=colnames(dat.bin))
## Plot the subfigure
ggplot(mut.rates, aes(Dog, Muts)) +
geom_bar(aes(fill = Hist), position = "dodge", stat="identity") +
scale_fill_manual(values=cols.hist[2:3]) +
theme_minimal() +
coord_flip() +
theme(axis.text.x=element_text(angle = -325, hjust = 1), text = element_text(size=10))
ggsave(mut.rates.file,width=4,height=10)
# +
#########################################
## Figure 2a - red&blue density plot
#####################################
print('Generating density plot')
## Count mutations in each benign & malignant sample, create and save density plot
samples.freq <- data.frame(Mutations=apply(dat, 2, sum), Hist=dat.hist[colnames(dat),'Hist'])
ggplot(samples.freq) + geom_density(aes(Mutations,group=Hist,col=Hist),lwd=3) + scale_color_manual(values=cols.hist[2:3]) + theme_bw() + theme(text = element_text(size=20))
ggsave(density.plot.file,width=12, height=4)
# -
## Print the median number of mutated genes per histology
print( paste('Median mutations in benign samples:', median( samples.freq[samples.freq$Hist=='B','Mutations']) ))
print( paste('Median mutations in malignant samples:', median( samples.freq[samples.freq$Hist=='M','Mutations']) ))
# +
#########################################
## Figure 2b - navy&white dot plot
#####################################
print('Generating pooled mutations plot')
## Calculate most frequently mutated (by % of samples of each type) to get balanced frequently mutated genes
## Otherwise will give mostly benign mutations, since we have 2x benign samples
ids.benign <- rownames(dat.hist)[ dat.hist$Hist=='B' ]
ids.tumor <- rownames(dat.hist)[ dat.hist$Hist=='M' ]
benign.ratios <- apply(dat.bin[,ids.benign], 1, function(x){sum(x==1)/length(x)})
tumor.ratios <- apply(dat.bin[,ids.tumor], 1, function(x){sum(x==1)/length(x)})
max.ratios <- apply(cbind(benign.ratios,tumor.ratios), 1, max)
# +
## Pick 30 most frequently mutated genes in COSMIC to use in the plots
genes <- names(sort(apply(dat[rownames(dat) %in% genes.cosmic,], 1, function(x){sum(x>0)}),decreasing=TRUE))[1:30] # Use this for the COSMIC plot
# Another option - pick some cutoff of mutated ratios for benign/malignant
#genes <- names(max.ratios)[max.ratios>0.15]
print(c('Most frequently mutated COSMIC genes:',genes))
# +
## Melt the malignant sample matrix
dat.m.melted <- melt( as.matrix(dat.m[rownames(dat.m) %in% genes,]) )
dat.m.melted$value <- as.numeric(as.character(dat.m.melted$value))
dat.m.melted$value[dat.m.melted$value>0] <- 1
dat.m.melted$value <- as.factor(dat.m.melted$value) # For color scales
## Melt the benign sample matrix
dat.b.melted <- melt( as.matrix(dat.b[rownames(dat.b) %in% genes,]) )
dat.b.melted$value <- as.numeric(as.character(dat.b.melted$value))
dat.b.melted$value[dat.b.melted$value>0] <- 1
dat.b.melted$value <- as.factor(dat.b.melted$value) # For color scales
## Combine the 2 melted matrices
dat.melted <- cbind(dat.m.melted, dat.b.melted$value)
colnames(dat.melted) <- c('Gene','Dog','Tumor','Benign')
dat.melted$Dog <- as.character(dat.melted$Dog) # So the plot sorts them alphabetically
# -
## Plot the result
ggplot(dat.melted) + geom_point(aes(Gene, Dog, col=Tumor), size=8, pch=15) +
geom_point(aes(Gene,Dog,col=Benign), size=4, pch=16) +
theme(axis.text.x=element_text(angle = -325, hjust = 1)) +
scale_color_manual(values=c('white',cols[10]))
ggsave(cosmic.mutations.file, width=10, height=5.5)
# +
#########################################
## Frequently mutated genes
#####################################
print('Generating per-sample mutations plot')
## Number of samples per patient (used to plot vertical grey dividing lines on the plot)
s.counts <- table(dat.hist[colnames(dat.bin),'Patient'])
# For our dataset only, reorder the names (because we used numeric patient names :/
#s.counts <- s.counts[sort(paste0(names(s.counts),'A'),index=TRUE)$ix]
## Prepare the data for the plot
dat.bin.melted <- melt(as.matrix(dat.bin[genes,]))
dat.bin.melted <- dat.bin.melted[,c(2,1,3)]
colnames(dat.bin.melted) <- c('Sample','Gene','Alteration')
dat.bin.melted$Sample <- as.character(dat.bin.melted$Sample)
dat.bin.melted$Hist <- dat.hist[ dat.bin.melted$Sample, 'Hist' ]
## Samples by genes plots of mutations, colored by sample histology
ggplot(dat.bin.melted) +
geom_point(aes(Sample, Gene, color=interaction(factor(Alteration),Hist)),pch=15,size=3) +
scale_color_manual(values=c('white',cols.hist[2],'white',cols.hist[3])) +
theme_classic() +
theme(legend.position='none',axis.text.x=element_text(angle = -325, hjust = 1)) +
geom_vline(xintercept=cumsum(s.counts[-length(s.counts)])+0.5,col=cols[14],size=2)
ggsave(mutations.consistency.file,width=13,height=7)
# +
## Do the subtypes have different numbers of mutations (total, not just in PAM50 genes)
## For samples of each subtype, print median num mutations in the samples
num.muts <- apply(dat.bin, 2, sum)
### Are COSMIC genes more frequently mutated than non-COSMIC?
print(res.ttest <- t.test( apply(dat.bin, 1, sum) ~ factor(rownames(dat.bin) %in% genes.cosmic) ))
if( res.ttest$p.value < 0.05) {
print( paste('COSMIC genes are significantly more frequently mutated than non-COSMIC genes, p-value =', signif(res.ttest$p.value,digits=3)) )
} else {
print( paste('COSMIC genes are NOT significantly more frequently mutated than non-COSMIC genes, p-value =', signif(res.ttest$p.value,digits=3)) )
}
rm(res.ttest)
# +
## Are PAM50 genes more frequently mutated than non-PAM50?
## Print PAM50 subtype sample mutation counts
if(run.pam50) {
colnames(pam50)[1] <- 'PAM50'
pam50$Muts <- NA
pam50[colnames(dat.bin),'Muts'] <- num.muts
print('PAM50 sample counts:'); flush.console()
print(sapply( levels(pam50$PAM50), function(x) {median( pam50[pam50$PAM50==x,'Muts'], na.rm=TRUE )} ))
}
# Only run if enabled
if(run.pam50) {
genes.pam50 <- rownames(read.table(fn.pam50.genes, sep=',', row.names=1))
print( res.ttest <- t.test( apply(dat.bin, 1, sum) ~ factor(rownames(dat.bin) %in% genes.pam50) ) )
if( res.ttest$p.value < 0.05) {
print( paste('PAM50 genes are significantly more frequently mutated than non-PAM50 genes, p-value =', signif(res.ttest$p.value,digits=3)) )
} else {
print( paste('PAM50 genes are NOT significantly more frequently mutated than non-PAM50 genes, p-value =', signif(res.ttest$p.value,digits=3)) )
}
rm(res.ttest)
}
# +
## Correlate mutations w/clinical factors of interest- this will return a matrix of dat.hist columns by genes, filled with corrected pvals
print(paste('Calculating correlations between mutations and phenotype data (Patient, Location, Histology, etc) in genes with >',opt.num,'mutations in the cohort.'))
get.pvals <- function(id) {
phen.cols <- c('Patient','Location','Goldschmidt','Hist','SimHist','DetHist') # Which clinical factors we care about
phen.cols <- phen.cols[ phen.cols %in% colnames(dat.hist) ] # Make sure these are in the provided phenotype/clinical data
p.adjust(apply(dat.hist[colnames(dat.bin),phen.cols], 2, function(x) {try(chisq.test(table( factor(x), unlist(dat.bin[id,])))$p.value)}))
}
## Look at most frequently mutated genes, correlate these mutations with phenotypes of interest
genes <- names(which(apply(dat.bin, 1, sum)>opt.num)) # Only care about frequently mutated genes
print(paste(length(genes),'genes are mutated in at least',opt.num,'samples:'))
print(genes)
if( length(genes) > 0 ) {
genes.pvals <- sapply(genes, get.pvals)
write.table(signif(t(genes.pvals),digits=5), file=freq.mutations.file, sep=',', col.names=TRUE, row.names=TRUE, quote=FALSE)
print('Phenotype/Clinical correlations stored to file.')
} else {
print('WARNING: Not enough mutated genes for clinical factor correlation analysis, skipping this step.')
}
# +
## Are PEP list genes more frequently mutated than non-PEP genes?
## Load the PEPs & print PEP genes that are frequently mutated
print('Loading PEPs. PEP list lengths:')
peps <- read.table(fn.peps, sep=',', header=TRUE, stringsAsFactors=FALSE)
peps <- list(Adenoma=peps[peps$Adenoma_Expression_Pattern < 0.05,'HumanSymbol'],
Carcinoma=peps[peps$Carcinoma_Expression_Pattern < 0.05,'HumanSymbol'],
Tumor=peps[peps$Tumor_Expression_Pattern < 0.05,'HumanSymbol'])
print(sapply(peps, length))# Print num genes in each PEP
print(paste('Checking for frequently mutated PEP genes (>',opt.num,'mutations):'))
pep.mut.counts <- sapply(peps, function(x) { apply(dat.bin[rownames(dat.bin) %in% x,], 1, sum)} )
print(sapply(pep.mut.counts, function(x){ names(x)[which(x>opt.num)] }))
print(sapply(pep.mut.counts, summary))
flush.console()
print('Done with mutation analysis.')
| DataAnalysis/mutation_analysis_notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exercise Introduction
# We will return to the automatic rotation problem you worked on in the previous exercise. But we'll add data augmentation to improve your model.
#
# The model specification and compilation steps don't change when you start using data augmentation. The code you've already worked with for specifying and compiling a model is in the cell below. Run it so you'll be ready to work on data augmentation.
# + _kg_hide-output=true
from tensorflow.keras.applications import ResNet50
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Flatten, GlobalAveragePooling2D
num_classes = 2
resnet_weights_path = '../input/resnet50/resnet50_weights_tf_dim_ordering_tf_kernels_notop.h5'
my_new_model = Sequential()
my_new_model.add(ResNet50(include_top=False, pooling='avg', weights=resnet_weights_path))
my_new_model.add(Dense(num_classes, activation='softmax'))
my_new_model.layers[0].trainable = False
my_new_model.compile(optimizer='sgd', loss='categorical_crossentropy', metrics=['accuracy'])
# Set up code checking
from learntools.core import binder
binder.bind(globals())
from learntools.deep_learning.exercise_5 import *
print("Setup Complete")
# -
# # 1) Fit the Model Using Data Augmentation
#
# Here is some code to set up some ImageDataGenerators. Run it, and then answer the questions below about it.
# +
from tensorflow.keras.applications.resnet50 import preprocess_input
from tensorflow.keras.preprocessing.image import ImageDataGenerator
image_size = 224
# Specify the values for all arguments to data_generator_with_aug.
data_generator_with_aug = ImageDataGenerator(preprocessing_function=preprocess_input,
horizontal_flip = True,
width_shift_range = 0.1,
height_shift_range = 0.1)
data_generator_no_aug = ImageDataGenerator(preprocessing_function=preprocess_input)
# -
# Why do we need both a generator with augmentation and a generator without augmentation? After thinking about it, check out the solution below.
# Check your answer (Run this code cell to receive credit!)
q_1.solution()
# # 2) Choosing Augmentation Types
# ImageDataGenerator offers many types of data augmentation. For example, one argument is `rotation_range`. This rotates each image by a random amount that can be up to whatever value you specify.
#
# Would it be sensible to use automatic rotation for this problem? Why or why not?
# Check your answer (Run this code cell to receive credit!)
q_2.solution()
# # 3) Code
# Fill in the missing pieces in the following code. We've supplied some boilerplate. You need to think about what ImageDataGenerator is used for each data source.
# + tags=["raises-exception"]
# Specify which type of ImageDataGenerator above is to load in training data
train_generator = data_generator_with_aug.flow_from_directory(
directory = '../input/dogs-gone-sideways/images/train',
target_size=(image_size, image_size),
batch_size=12,
class_mode='categorical')
# Specify which type of ImageDataGenerator above is to load in validation data
validation_generator = data_generator_no_aug.flow_from_directory(
directory = '../input/dogs-gone-sideways/images/val',
target_size=(image_size, image_size),
class_mode='categorical')
my_new_model.fit_generator(
____, # if you don't know what argument goes first, try the hint
epochs = 3,
steps_per_epoch=19,
validation_data=____)
# Check your answer
q_3.check()
# +
# q_3.hint()
# q_3.solution()
# +
# #%%RM_IF(PROD)%%
train_generator = data_generator_with_aug.flow_from_directory(
directory = '../input/dogs-gone-sideways/images/train',
target_size=(image_size, image_size),
batch_size=12,
class_mode='categorical')
# Specify which type of ImageDataGenerator above is to load in validation data
validation_generator = data_generator_no_aug.flow_from_directory(
directory = '../input/dogs-gone-sideways/images/val',
target_size=(image_size, image_size),
class_mode='categorical')
my_new_model.fit_generator(
train_generator,
epochs = 3,
steps_per_epoch=19,
validation_data=validation_generator)
q_3.assert_check_passed()
# -
# # 4) Did Data Augmentation Help?
# How could you test whether data augmentation improved your model accuracy?
# Check your answer (Run this code cell to receive credit!)
q_4.solution()
# # Keep Going
# You are ready for **[a deeper understanding of deep learning](https://www.kaggle.com/dansbecker/a-deeper-understanding-of-deep-learning/)**.
#
| notebooks/deep_learning/raw/ex5_data_augmentation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Analysis of the 4x3 lexicons for the HLC RSA and Non-Ostensive simulations.
# Import packages for reading data and generating images
#imports
import os
import numpy as np
import pandas as pd
import seaborn as sns
from matplotlib import pyplot as plt
import scipy.stats as st
import re
import csv
# Below are three methods for reading the outputfiles as Pandas DataFrames
# config.csv:
# agentPairs;maxTurns;roundsPlayed;beta;entropyThreshold;order;costs;initiatorDistribution;responderDistribution
def read_config(location, config_title):
filename = location + "config" + config_title + ".csv"
with open(filename) as f:
config = pd.read_csv(filename, sep=";")
return config
# results_rounds.csv:
# pair;round;nrTurns;success
def read_results_rounds(location, config_title):
filename = location + "results_rounds" + config_title + ".csv"
with open(filename) as f:
results_rounds = pd.read_csv(filename, sep=";")
return results_rounds
# results_turns.csv:
# pair;round;turn;initiatorIntention;initiatorSignal;responderInference;responderSignal;entropyInitiatorListen;entropyResponderListen;entropyInitiatorLexicon;entropyResponderLexicon;KLDivItoR;KLDivRtoI
def read_results_turns(location, config_title):
filename = location + "results_turns" + config_title + ".csv"
with open(filename) as f:
results_turns = pd.read_csv(filename, sep=";")
return results_turns
# Defining the names and locations of files that I am going to compare. 4x3 lexicon, binomial distribution with X=0.5, alpha=5 and agents=500.
# +
# This is the baseline to compare against
non_ostensive_model_title = "_a500_b5.0_d0.5"
non_ostensive_model_location = "data/S4R3_non_ostensive_a500_b5.0_d0.5/"
# List of all titles and locations
# This list is sorted on neighborliness first
# hlc_rsa_model_title_list = ["_a5_b5.0_d0.5_f0_n0.0", "_a100_b5.0_d0.5_f1_n0.0", "_a100_b5.0_d0.5_f2_n0.0", "_a100_b5.0_d0.5_f3_n0.0",
# "_a100_b5.0_d0.5_f0_n0.5", "_a100_b5.0_d0.5_f1_n0.5", "_a100_b5.0_d0.5_f2_n0.5", "_a500_b5.0_d0.5_f3_n0.5",
# "_a500_b5.0_d0.5_f0_n0.7", "_a500_b5.0_d0.5_f1_n0.7", "_a500_b5.0_d0.5_f2_n0.7", "_a500_b5.0_d0.5_f3_n0.7",
# "_a500_b5.0_d0.5_f0_n0.9", "_a500_b5.0_d0.5_f1_n0.9", "_a100_b5.0_d0.5_f2_n0.9", "_a500_b5.0_d0.5_f3_n0.9"]
# This list is sorted on facts first
hlc_rsa_model_title_list = ["_a5_b5.0_d0.5_f0_n0.0", "_a100_b5.0_d0.5_f0_n0.5", "_a500_b5.0_d0.5_f0_n0.7", "_a500_b5.0_d0.5_f0_n0.9",
"_a100_b5.0_d0.5_f1_n0.0", "_a100_b5.0_d0.5_f1_n0.5", "_a500_b5.0_d0.5_f1_n0.7", "_a500_b5.0_d0.5_f1_n0.9",
"_a100_b5.0_d0.5_f2_n0.0", "_a100_b5.0_d0.5_f2_n0.5", "_a500_b5.0_d0.5_f2_n0.7", "_a100_b5.0_d0.5_f2_n0.9",
"_a100_b5.0_d0.5_f3_n0.0", "_a500_b5.0_d0.5_f3_n0.5", "_a500_b5.0_d0.5_f3_n0.7", "_a500_b5.0_d0.5_f3_n0.9"]
hlc_rsa_model_location = "data/S4R3"
# Dictionary for renaming the codelike parameters to meaningful descriptions
dictionary_to_rename_parameters = {
"a": "Agents",
"b": "Alpha",
"d": "X",
"f": "Facts",
"n": "Neighborliness",
}
# Looks up the codelike character in the dictionary and replaces that with the descriptive word
def make_human_readable(parameter):
parameter[0] = dictionary_to_rename_parameters[parameter[0]]
return parameter
# Splits the parameter string in letters and number. ('a500' becomes 'a', '500')
def split_parameter_string(parameters):
return [(re.split('(\d.*)', level)[:-1]) for level in parameters]
# Replaces the single letters with the descriptive words
def readable_string(parameters):
return [make_human_readable(level) for level in parameters]
# Split the title into the seperate parameters, using '_'. "_a500_b5" becomes "a500", "b5"
split_parameters_list = [title.split("_")[1:] for title in hlc_rsa_model_title_list]
# List comprehension that creates a list of lists that contain the names and values for the individual parameters
parameters_list = [readable_string(split_parameter_string(parameters)) for parameters in split_parameters_list]
# -
# Importing all the datasets
# +
# Do this only once, baseline is the same every time
non_ostensive_results_turns = read_results_turns(non_ostensive_model_location, non_ostensive_model_title)
# List comprehension to get the results of the turns stored in a list
hlc_rsa_results_turns_list = [read_results_turns(hlc_rsa_model_location + title + "/", title) for title in hlc_rsa_model_title_list]
# -
# The labelling of turns is off slightly because of how the models was built, the below code fixes that (do not run it extra without reloading the data, else the +1 row will throw everything off)
# +
# Run this only once after loading the data
for i, row in non_ostensive_results_turns.iterrows():
if row["turn"] == 0 and np.isnan(row["entropyInitiatorListen"]):
non_ostensive_results_turns.at[i,"turn"] = -1
non_ostensive_results_turns["turn"] = [x+1 for x in non_ostensive_results_turns["turn"]]
# Running this for all results in the list
for hlc_rsa_results_turns in hlc_rsa_results_turns_list:
for i, row in hlc_rsa_results_turns.iterrows():
if row["turn"] == 0 and np.isnan(row["entropyInitiatorListen"]):
hlc_rsa_results_turns.at[i,"turn"] = -1
hlc_rsa_results_turns["turn"] = [x+1 for x in hlc_rsa_results_turns["turn"]]
# -
# Merging the results of the rounds with that of the turns
# +
# Results of baseline dialogue
non_ostensive_results_rounds = read_results_rounds(non_ostensive_model_location, non_ostensive_model_title)
# List comprehension to get the results of the rounds stored in a list
hlc_rsa_results_rounds_list = [read_results_rounds(hlc_rsa_model_location + title + "/", title) for title in hlc_rsa_model_title_list]
# -
#
#
# +
# Baseline
non_ostensive_results_merged = non_ostensive_results_rounds.merge(non_ostensive_results_turns, left_on=['pair', 'round'], right_on=['pair', 'round'])
# List comprehension to get merged results
hlc_rsa_results_merged_list = [rounds.merge(turns, left_on=['pair', 'round'], right_on=['pair', 'round']) for rounds, turns in zip(hlc_rsa_results_rounds_list, hlc_rsa_results_turns_list)]
# -
# Adding disambiguation between the models and putting them in one dataframe for some of the image creation.
# +
# Baseline
non_ostensive_results_rounds['model'] = "Non-Ostensive Baseline"
# Combining hlc-rsa with baseline for image creation
for hlc_rsa_results_rounds, parameters in zip(hlc_rsa_results_rounds_list, parameters_list):
hlc_rsa_results_rounds['model'] = "HLC-RSA\n{}\n{}".format(" = ".join(parameters[3]), " = ".join(parameters[4]))
mixed_results_rounds_list = [pd.concat([hlc_rsa_results_rounds, non_ostensive_results_rounds]) for hlc_rsa_results_rounds in hlc_rsa_results_rounds_list]
# -
# Removing all cases where the agents gave up
for mixed_results_rounds in mixed_results_rounds_list:
mixed_results_rounds = mixed_results_rounds[mixed_results_rounds['nrTurns'] < 7]
# Creating the baseline image, this has only one model in it
colorDict = {"Non-Ostensive Baseline" : "#ff7f0e"}
plt.figure()
f = sns.lineplot(data=non_ostensive_results_rounds, x="round", y="nrTurns", hue="model", err_style="bars", palette=colorDict)
c = sns.lineplot(data=non_ostensive_results_rounds, x="round", y="nrTurns", hue="model", ci='sd', palette=colorDict, legend=False)
sns.despine()
f.set_ylim([0,7])
f.set_xlabel("Intention")
f.set_title("Reduction of turn sequence length")
#plt.savefig(fname="images/analysis 1/non-ostensive.jpg")
# Creating all images for analysis 1. The clarification sequence length that the agents needed
path = "images/analysis 1/"
for mixed_results_rounds, title in zip(mixed_results_rounds_list, hlc_rsa_model_title_list):
plt.figure()
f = sns.lineplot(data=mixed_results_rounds, x="round", y="nrTurns", hue="model", err_style="bars")
c = sns.lineplot(data=mixed_results_rounds, x="round", y="nrTurns", hue="model", ci='sd', legend=False)
sns.despine()
f.set_ylim([0,7])
f.set_xlabel("Intention")
f.set_title("Reduction of turn sequence length")
name = path + title + ".jpg"
#plt.savefig(fname=name)
# The images for analysis 2. The factual understanding for each model
# Baseline non-ostensive
plt.figure()
f = sns.catplot(x="nrTurns", hue="success", col="round", data=non_ostensive_results_rounds, kind="count", palette="Dark2_r")
(f.set_axis_labels("Turns", "Count")
.set(ylim=(0, 250))
.set(xlim=(-0.5, 7)))
f.fig.subplots_adjust(top=0.9)
f.fig.suptitle("Facts = 0\nNeighborliness = 0.0")
#plt.savefig(fname="images/analysis 2/non-ostensive.jpg")
# For loop to create all images. These images are compounded into two large images in my thesis
path = "images/analysis 2/adjusted/"
for hlc_rsa_results_rounds, parameters, title in zip(hlc_rsa_results_rounds_list, parameters_list, hlc_rsa_model_title_list):
plt.figure()
f = sns.catplot(x="nrTurns", hue="success", col="round", data=hlc_rsa_results_rounds, kind="count", palette="Dark2_r")
if(parameters[0][1] == "100"):
(f.set_axis_labels("Turns", "Count").set(ylim=(0, 50)).set(xlim=(-0.5, 7)))
else:
(f.set_axis_labels("Turns", "Count").set(ylim=(0, 250)).set(xlim=(-0.5, 7)))
f.fig.subplots_adjust(top=0.9)
f.fig.suptitle("{}\n{}".format(" = ".join(parameters[3]), " = ".join(parameters[4])))
name = path + title + ".jpg"
#plt.savefig(fname=name)
# Analysis 3 is for the percentual perceived understanding for all models. Only the final set of tables is relevant for the thesis
# Non-Ostensive baseline
non_ostensive_understanding_table = [[0, 0], [0, 0]]
perceived_understanding_non_ostensive = non_ostensive_results_rounds[non_ostensive_results_rounds.nrTurns <= 6]
perceived_misunderstanding_non_ostensive = non_ostensive_results_rounds[non_ostensive_results_rounds.nrTurns > 6]
non_ostensive_understanding_table[0][0] = perceived_understanding_non_ostensive[perceived_understanding_non_ostensive.success == True].count(1).size
non_ostensive_understanding_table[0][1] = perceived_understanding_non_ostensive[perceived_understanding_non_ostensive.success == False].count(1).size
non_ostensive_understanding_table[1][0] = perceived_misunderstanding_non_ostensive[perceived_misunderstanding_non_ostensive.success == True].count(1).size
non_ostensive_understanding_table[1][1] = perceived_misunderstanding_non_ostensive[perceived_misunderstanding_non_ostensive.success == False].count(1).size
print("Non-Ostensive\t\t | Perceived understanding | Give up")
print("-------------------------|-------------------------|--------")
print("Factual understanding\t | {}\t\t\t | {}".format(non_ostensive_understanding_table[0][0], non_ostensive_understanding_table[1][0]))
print("-------------------------|-------------------------|--------")
print("Factual misunderstanding | {}\t\t\t | {}\n".format(non_ostensive_understanding_table[0][1], non_ostensive_understanding_table[1][1]))
# And for the HLC-RSA case
combined_list = []
for hlc_rsa_results_rounds, parameters in zip(hlc_rsa_results_rounds_list, parameters_list):
temp_table = [[0, 0], [0, 0]]
perceived_understanding_hlc_rsa = hlc_rsa_results_rounds[hlc_rsa_results_rounds.nrTurns <= 6]
perceived_misunderstanding_hlc_rsa = hlc_rsa_results_rounds[hlc_rsa_results_rounds.nrTurns > 6]
temp_table[0][0] = perceived_understanding_hlc_rsa[perceived_understanding_hlc_rsa.success == True].count(1).size
temp_table[0][1] = perceived_understanding_hlc_rsa[perceived_understanding_hlc_rsa.success == False].count(1).size
temp_table[1][0] = perceived_misunderstanding_hlc_rsa[perceived_misunderstanding_hlc_rsa.success == True].count(1).size
temp_table[1][1] = perceived_misunderstanding_hlc_rsa[perceived_misunderstanding_hlc_rsa.success == False].count(1).size
params = "{}\n{}".format(" = ".join(parameters[3]), " = ".join(parameters[4]))
hlc_rsa_understanding_tables = temp_table
combined_list.append((params, hlc_rsa_understanding_tables))
# +
def print_combined_table(combined_table):
model, table = combined_table
print("{}\t | Perceived understanding | Give up".format(model))
print("-------------------------|-------------------------|--------")
print("Factual understanding\t | {}\t\t\t | {}".format(table[0][0], table[1][0]))
print("-------------------------|-------------------------|--------")
print("Factual misunderstanding | {}\t\t\t | {}\n".format(table[0][1], table[1][1]))
for combined_table in combined_list:
print_combined_table(combined_table)
# -
# In percentages:
# +
def table_to_percentage(table):
total = sum(sum(x) for x in table)
percentage_table = [[0, 0], [0, 0]]
percentage_table[0][0] = table[0][0] / total * 100.0
percentage_table[0][1] = table[0][1] / total * 100.0
percentage_table[1][0] = table[1][0] / total * 100.0
percentage_table[1][1] = table[1][1] / total * 100.0
return percentage_table
def print_percentage_table(combined_percentage_table):
model, table = combined_percentage_table
print("{}\t | Perceived understanding | Give up".format(model))
print("-------------------------|-------------------------|--------")
print("Factual understanding\t | {:.2f} %\t\t | {:.2f} %".format(table[0][0], table[1][0]))
print("-------------------------|-------------------------|--------")
print("Factual misunderstanding | {:.2f} %\t\t | {:.2f} %\n".format(table[0][1], table[1][1]))
# -
# Baseline
non_ostensive_percentage_table = table_to_percentage(non_ostensive_understanding_table)
print_percentage_table(("Non-Ostensive\nbaseline\t", non_ostensive_percentage_table))
# HLC-RSA
for combined_table in combined_list:
model, table = combined_table
combined_percentage_table = (model, table_to_percentage(table))
print_percentage_table(combined_percentage_table)
# Below is the final analysis 3: showing percentage of factual understanding split by whether the agents perceived understanding or gave up.
# +
def table_to_percentage_split(table):
result = [0, 0]
total_perceived = table[0][0] + table[0][1]
total_give_up = table[1][0] + table[1][1]
if(total_perceived != 0):
result[0] = table[0][0] / total_perceived * 100
if(total_give_up != 0):
result[1] = table[1][0] / total_give_up * 100
return result
non_ostensive_result = table_to_percentage_split(non_ostensive_understanding_table)
# Use this when the list is sorted on facts first
with open("percentage_csv_facts.csv", 'w') as f:
writer=csv.writer(f, delimiter=',',lineterminator='\n')
writer.writerow(["Facts", "Neighborliness", "Perceived understanding", "Give up"])
for (_, table), params in zip(combined_list, parameters_list):
_, table = (_, table)
row = []
result = table_to_percentage_split(table)
row.append(int(params[3][1]))
row.append(round(float(params[4][1]), 2))
row.append(str(round(result[0], 2)) + ' %')
row.append(str(round(result[1], 2)) + ' %')
writer.writerow(row)
# Use this when the list is sorted on neighborliness first
# with open("percentage_csv_neighborliness.csv", 'w') as f:
# writer=csv.writer(f, delimiter=',',lineterminator='\n')
# writer.writerow(["Neighborliness", "Facts", "Perceived understanding", "Give up"])
# for (_, table), params in zip(combined_list, parameters_list):
# _, table = (_, table)
# row = []
# result = table_to_percentage_split(table)
# row.append(round(float(params[4][1]), 2))
# row.append(int(params[3][1]))
# row.append(str(round(result[0], 2)) + ' %')
# row.append(str(round(result[1], 2)) + ' %')
# writer.writerow(row)
for model, table in combined_list:
result = table_to_percentage_split(table)
print("{}\t | Perceived understanding | Give up".format(model))
print("-------------------------|-------------------------|--------")
print("RSA-HLC\t\t\t | {:.2f} %\t\t | {:.2f} %".format(result[0], result[1]))
print("-------------------------|-------------------------|--------")
print("Non-Ostensive\t\t | {:.2f} %\t\t | {:.2f} %".format(non_ostensive_result[0], non_ostensive_result[1]))
print()
# -
print(non_ostensive_result)
| HLC-RSA/code/Human Language Constrained RSA Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
# ### Determinant of a square array
# +
arr_shape = int(input("Enter size of square array: "))
arr = list(map(int,input("Enter a square array in row-wise manner: ").split()))
if len(arr) == arr_shape**2 :
sq_arr = np.array(arr, dtype=int).reshape(arr_shape, arr_shape)
print("\nSUCCESS: Reshape operation completed")
print(f"Given square array is")
print(sq_arr)
print(f"\nThe approx. determinant of the above square array is {round(np.linalg.det(sq_arr),0)}")
print(f"The absolute determinant of the above square array is {np.linalg.det(sq_arr)}")
else:
print(f"ERROR: Cannot reshape array of size {len(arr)} into {(arr_shape, arr_shape)}")
# -
| bits_wilp/Ex2_Numpy_Q4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# %matplotlib inline
import mxnet as mx
from mxnet import nd
from mxnet.contrib.ndarray import MultiBoxPrior
from mxnet.gluon.contrib import nn as nn_contrib
from mxnet.gluon import nn
ctx = mx.gpu()
# ### Predict classes
# - channel `i*(num_class+1)` store the scores for this box contains only background
# - channel `i*(num_class+1)+1+j` store the scores for this box contains an object from the *j*-th class
def class_predictor(num_anchors, num_classes):
return nn.Conv2D(num_anchors * (num_classes + 1), 3, padding=1)
# ### Predict anchor boxes
# - $t_x = (Y_x - b_x) / b_{width}$
# - $t_y = (Y_y - b_y) / b_{height}$
# - $t_{width} = (Y_{width} - b_{width}) / b_{width}$
# - $t_{height} = (Y_{height} - b_{height}) / b_{height}$
def box_predictor(num_anchors):
return nn.Conv2D(num_anchors * 4, 3, padding=1)
# ### Manage preditions from multiple layers
# +
def flatten_prediction(pred):
return nd.flatten(nd.transpose(pred, axes=(0, 2, 3, 1)))
def concat_predictions(preds):
return nd.concat(*preds, dim=1)
# -
# ### Down-sample features
def dp_layer(nfilters, stride, expension_constant):
out = nn.HybridSequential()
out.add(nn.Conv2D(nfilters, 3, strides=stride, padding=1, groups=nfilters, use_bias=False))
out.add(nn.BatchNorm(use_global_stats=False, epsilon=1e-05, momentum=0.9, axis=1))
out.add(nn.Activation('relu'))
out.add(nn.Conv2D(nfilters*expension_constant, 1, strides=1, padding=0, use_bias=False))
out.add(nn.BatchNorm(use_global_stats=False, epsilon=1e-05, momentum=0.9, axis=1))
out.add(nn.Activation('relu'))
return out
# ### Scale units
global alpha
alpha = 0.5
num_filter = int(32*alpha)
# ### Body network
# +
def s16():
out = nn.HybridSequential()
with out.name_scope():
# conv2d
out.add(nn.Conv2D(num_filter, kernel_size=3, strides=2, padding=1, use_bias=False))
out.add(nn.BatchNorm(use_global_stats=False, epsilon=1e-05, momentum=0.9, axis=1))
out.add(nn.Activation('relu'))
# (3) LinearBottleneck
out.add(dp_layer(num_filter, 1, 1))
#out.add(nn.Conv2D(num_filter, kernel_size=3, strides=1, padding=1, use_bias=False))
out.add(nn.BatchNorm(use_global_stats=False, epsilon=1e-05, momentum=0.9, axis=1))
out.add(nn.Activation('relu'))
out.add(nn.Conv2D(num_filter, kernel_size=3, strides=1, padding=1, groups=num_filter, use_bias=False))
out.add(nn.BatchNorm(use_global_stats=False, epsilon=1e-05, momentum=0.9, axis=1))
out.add(nn.Activation('relu'))
out.add(nn.Conv2D(num_filter/2, kernel_size=1, strides=1, padding=0, use_bias=False))
out.add(nn.BatchNorm(use_global_stats=False, epsilon=1e-05, momentum=0.9, axis=1))
# (4) LinearBottleneck
out.add(dp_layer(num_filter/2, 1, 6))
#out.add(nn.Conv2D(num_filter*3, kernel_size=3, strides=1, padding=1, use_bias=False))
out.add(nn.BatchNorm(use_global_stats=False, epsilon=1e-05, momentum=0.9, axis=1))
out.add(nn.Activation('relu')) # conv2_2_linear_scale
out.add(nn.Conv2D(num_filter*3, kernel_size=3, strides=2, padding=1, groups=num_filter*3, use_bias=False))
#out.load_parameters("weights/mobilenet2_0_25_s16_org.params")
return out
def s32():
out = nn.HybridSequential()
with out.name_scope():
# (4) LinearBottleneck con't
out.add(nn.BatchNorm(use_global_stats=False, epsilon=1e-05, momentum=0.9, axis=1))
out.add(nn.Activation('relu'))
out.add(nn.Conv2D(num_filter*3/4, kernel_size=1, strides=1, padding=0, use_bias=False))
out.add(nn.BatchNorm(use_global_stats=False, epsilon=1e-05, momentum=0.9, axis=1)) # conv2_2_linear_scale
# (5) LinearBottleneck
out.add(dp_layer(num_filter*3/4, 1, 6))
#out.add(nn.Conv2D(num_filter*9/2, kernel_size=3, strides=1, padding=1, use_bias=False))
out.add(nn.BatchNorm(use_global_stats=False, epsilon=1e-05, momentum=0.9, axis=1))
out.add(nn.Activation('relu'))
out.add(nn.Conv2D(num_filter*9/2, kernel_size=3, strides=1, padding=1, groups=num_filter*9/2, use_bias=False))
out.add(nn.BatchNorm(use_global_stats=False, epsilon=1e-05, momentum=0.9, axis=1))
out.add(nn.Activation('relu'))
out.add(nn.Conv2D(num_filter*6/8, kernel_size=1, strides=1, padding=0, use_bias=False))
out.add(nn.BatchNorm(use_global_stats=False, epsilon=1e-05, momentum=0.9, axis=1)) # conv2_2_linear_scale concatenate
# (6) LinearBottleneck
out.add(dp_layer(num_filter*6/8, 1, 6))
#out.add(nn.Conv2D(num_filter*9/2, kernel_size=3, strides=1, padding=1, use_bias=False))
out.add(nn.BatchNorm(use_global_stats=False, epsilon=1e-05, momentum=0.9, axis=1))
out.add(nn.Activation('relu'))
out.add(nn.Conv2D(num_filter*9/2, kernel_size=3, strides=2, padding=1, groups=num_filter*9/2, use_bias=False))
#out.load_parameters("weights/mobilenet2_0_25_s32_org.params")
return out
def b1():
out = nn.HybridSequential()
with out.name_scope():
# (7) LinearBottleneck con't
out.add(nn.BatchNorm(use_global_stats=False, epsilon=1e-05, momentum=0.9, axis=1))
out.add(nn.Activation('relu'))
out.add(nn.Conv2D(int(num_filter*6/8), kernel_size=1, strides=1, padding=0, use_bias=False))
out.add(nn.BatchNorm(use_global_stats=False, epsilon=1e-05, momentum=0.9, axis=1))
# (8) LinearBottleneck
out.add(dp_layer(num_filter*3/4, 1, 6))
#out.add(nn.Conv2D(num_filter*9/2, kernel_size=3, strides=1, padding=1, use_bias=False))
out.add(nn.BatchNorm(use_global_stats=False, epsilon=1e-05, momentum=0.9, axis=1))
out.add(nn.Activation('relu'))
out.add(nn.Conv2D(int(num_filter*9/2), kernel_size=3, strides=1, padding=1, groups=int(num_filter*9/2), use_bias=False))
out.add(nn.BatchNorm(use_global_stats=False, epsilon=1e-05, momentum=0.9, axis=1))
out.add(nn.Activation('relu'))
out.add(nn.Conv2D(int(num_filter*6/8), kernel_size=1, strides=1, padding=0, use_bias=False))
out.add(nn.BatchNorm(use_global_stats=False, epsilon=1e-05, momentum=0.9, axis=1))
# (9) LinearBottleneck
out.add(dp_layer(num_filter*3/4, 1, 6))
#out.add(nn.Conv2D(num_filter*9/2, kernel_size=3, strides=1, padding=1, use_bias=False))
out.add(nn.BatchNorm(use_global_stats=False, epsilon=1e-05, momentum=0.9, axis=1))
out.add(nn.Activation('relu'))
out.add(nn.Conv2D(int(num_filter*9/2), kernel_size=3, strides=2, padding=1, groups=int(num_filter*9/2), use_bias=False))
return out
def b2():
out = nn.HybridSequential()
with out.name_scope():
# (7) LinearBottleneck con't
out.add(nn.BatchNorm(use_global_stats=False, epsilon=1e-05, momentum=0.9, axis=1))
out.add(nn.Activation('relu'))
out.add(nn.Conv2D(int(num_filter*6/8), kernel_size=1, strides=1, padding=0, use_bias=False))
out.add(nn.BatchNorm(use_global_stats=False, epsilon=1e-05, momentum=0.9, axis=1))
# (8) LinearBottleneck
out.add(dp_layer(num_filter*3/4, 1, 6))
#out.add(nn.Conv2D(num_filter*9/2, kernel_size=3, strides=1, padding=1, use_bias=False))
out.add(nn.BatchNorm(use_global_stats=False, epsilon=1e-05, momentum=0.9, axis=1))
out.add(nn.Activation('relu'))
out.add(nn.Conv2D(int(num_filter*9/2), kernel_size=3, strides=1, padding=1, groups=int(num_filter*9/2), use_bias=False))
out.add(nn.BatchNorm(use_global_stats=False, epsilon=1e-05, momentum=0.9, axis=1))
out.add(nn.Activation('relu'))
out.add(nn.Conv2D(int(num_filter*6/8), kernel_size=1, strides=1, padding=0, use_bias=False))
out.add(nn.BatchNorm(use_global_stats=False, epsilon=1e-05, momentum=0.9, axis=1))
# (9) LinearBottleneck
out.add(dp_layer(num_filter*3/4, 1, 6))
#out.add(nn.Conv2D(num_filter*9/2, kernel_size=3, strides=1, padding=1, use_bias=False))
out.add(nn.BatchNorm(use_global_stats=False, epsilon=1e-05, momentum=0.9, axis=1))
out.add(nn.Activation('relu'))
out.add(nn.Conv2D(int(num_filter*9/2), kernel_size=3, strides=2, padding=1, groups=int(num_filter*9/2), use_bias=False))
return out
# -
# ### Create an SSD model
def ssd_model(num_anchors, num_classes):
class_preds = nn.Sequential()
box_preds = nn.Sequential()
for scale in range(4):
class_preds.add(class_predictor(num_anchors, num_classes))
box_preds.add(box_predictor(num_anchors))
#class_preds.initialize(mx.init.Xavier(magnitude=2), ctx=ctx)
#box_preds.initialize(mx.init.Xavier(magnitude=2), ctx=ctx)
return s16(), s32(), b1(), b2(), class_preds, box_preds
# ### Forward
def ssd_forward(x, s16, s32, b1, b2, class_preds, box_preds, sizes, ratios):
default_anchors = []
predicted_boxes = []
predicted_classes = []
x = s16(x)
default_anchors.append(MultiBoxPrior(x, sizes=sizes[0], ratios=ratios[0]))
predicted_boxes.append(flatten_prediction(box_preds[0](x)))
predicted_classes.append(flatten_prediction(class_preds[0](x)))
x = s32(x)
default_anchors.append(MultiBoxPrior(x, sizes=sizes[1], ratios=ratios[1]))
predicted_boxes.append(flatten_prediction(box_preds[1](x)))
predicted_classes.append(flatten_prediction(class_preds[1](x)))
x = b1(x)
default_anchors.append(MultiBoxPrior(x, sizes=sizes[2], ratios=ratios[2]))
predicted_boxes.append(flatten_prediction(box_preds[2](x)))
predicted_classes.append(flatten_prediction(class_preds[2](x)))
x = b2(x)
default_anchors.append(MultiBoxPrior(x, sizes=sizes[3], ratios=ratios[3]))
predicted_boxes.append(flatten_prediction(box_preds[3](x)))
predicted_classes.append(flatten_prediction(class_preds[3](x)))
return default_anchors, predicted_classes, predicted_boxes
# ### Put all things together
from mxnet import gluon
class SSD(gluon.Block):
def __init__(self, num_classes, **kwargs):
super(SSD, self).__init__(**kwargs)
self.anchor_sizes = [[0.04, 0.26],[0.26,0.58],[0.58,0.9],[0.9,1.06]]
self.anchor_ratios = [[1, 2, .5]] * 4
self.num_classes = num_classes
with self.name_scope():
self.s16, self.s32, self.b1, self.b2, self.class_preds, self.box_preds = ssd_model(4, num_classes)
def forward(self, x):
default_anchors, predicted_classes, predicted_boxes = ssd_forward(x, self.s16, self.s32, self.b1, self.b2,
self.class_preds, self.box_preds, self.anchor_sizes, self.anchor_ratios)
anchors = concat_predictions(default_anchors)
box_preds = concat_predictions(predicted_boxes)
class_preds = concat_predictions(predicted_classes)
class_preds = nd.reshape(class_preds, shape=(0, -1, self.num_classes + 1))
return anchors, class_preds, box_preds
# ### Outputs of SSD
net = SSD(2)
net.initialize(mx.init.Xavier(magnitude=2), ctx=ctx)
#net.load_parameters("process/ssd_99.params",ctx=ctx)
x = nd.zeros((1, 3, 512, 512),ctx=ctx)
default_anchors, class_predictions, box_predictions = net(x)
print('Outputs:', 'anchors', default_anchors.shape, 'class prediction', class_predictions.shape, 'box prediction', box_predictions.shape)
# ### Load dataset
# +
from source.NACDDetection import NACDDetection
train_dataset = NACDDetection(splits=[('NACDwNegswAugCropped', 'train')])
test_dataset = NACDDetection(splits=[('NACDwNegswAugCropped', 'test')])
print('Training images:', len(train_dataset))
print('Test images:', len(test_dataset))
# -
from source import NACDTransform
width, height = 512, 512
train_transform = NACDTransform.NACDDefaultTransform(width, height, False)
test_transform = NACDTransform.NACDDefaultTransform(width, height, True)
from gluoncv.data.transforms import presets
from gluoncv import utils
from mxnet import nd
from matplotlib import pyplot as plt
from gluoncv.utils import viz
train_image, train_label = test_dataset[0]
bboxes = train_label[:, :4]
cids = train_label[:, 4:5]
print('image:', train_image.shape)
print('bboxes:', bboxes.shape, 'class ids:', cids.shape)
train_image2, train_label2 = train_transform(train_image, train_label)
print('tensor shape:', train_image2.shape)
# +
from gluoncv.data.batchify import Tuple, Stack, Pad
from mxnet.gluon.data import DataLoader
batch_size = 16
num_workers = 4
batchify_fn = Tuple(Stack(), Pad(pad_val=-1))
train_loader = DataLoader(train_dataset.transform(train_transform), batch_size, shuffle=True,
batchify_fn=batchify_fn, last_batch='rollover', num_workers=num_workers)
test_loader = DataLoader(test_dataset.transform(test_transform), batch_size, shuffle=False,
batchify_fn=batchify_fn, last_batch='keep', num_workers=num_workers)
for ib, batch in enumerate(test_loader):
if ib > 3:
break
print('data:', batch[0].shape, 'label:', batch[1].shape)
# -
train_image2 = train_image2.transpose((1, 2, 0)) * nd.array((0.229, 0.224, 0.225)) + nd.array((0.485, 0.456, 0.406))
train_image2 = (train_image2 * 255).clip(0, 255)
ax = viz.plot_bbox(train_image2.asnumpy(), train_label2[:, :4],
labels=train_label2[:, 4:5],
class_names=train_dataset.classes)
plt.show()
# ## Train
from mxnet.contrib.ndarray import MultiBoxTarget
def training_targets(default_anchors, class_predicts, labels):
class_predicts = nd.transpose(class_predicts, axes=(0, 2, 1))
z = MultiBoxTarget(anchor=default_anchors.as_in_context(mx.cpu()), label=labels.as_in_context(mx.cpu()), cls_pred=class_predicts.as_in_context(mx.cpu()))
box_target = z[0].as_in_context(ctx) # box offset target for (x, y, width, height)
box_mask = z[1].as_in_context(ctx) # mask is used to ignore box offsets we don't want to penalize, e.g. negative samples
cls_target = z[2].as_in_context(ctx) # cls_target is an array of labels for all anchors boxes
return box_target, box_mask, cls_target
def convertlbl(y):
mtrx = y[:,:,0:4]
mtrx = mtrx.asnumpy()
mtrx[mtrx == -1] = -width
mtrx = mtrx/width
return mx.nd.concat(nd.expand_dims(y[:,:,4],2),mx.nd.array(mtrx),dim=2)
# +
class FocalLoss(gluon.loss.Loss):
def __init__(self, axis=-1, alpha=0.25, gamma=2, batch_axis=0, **kwargs):
super(FocalLoss, self).__init__(None, batch_axis, **kwargs)
self._axis = axis
self._alpha = alpha
self._gamma = gamma
def hybrid_forward(self, F, output, label):
output = F.softmax(output)
pt = F.pick(output, label, axis=self._axis, keepdims=True)
loss = -self._alpha * ((1 - pt) ** self._gamma) * F.log(pt)
return F.mean(loss, axis=self._batch_axis, exclude=True)
# cls_loss = gluon.loss.SoftmaxCrossEntropyLoss()
cls_loss = FocalLoss()
print(cls_loss)
# +
class SmoothL1Loss(gluon.loss.Loss):
def __init__(self, batch_axis=0, **kwargs):
super(SmoothL1Loss, self).__init__(None, batch_axis, **kwargs)
def hybrid_forward(self, F, output, label, mask):
loss = F.smooth_l1((output - label) * mask, scalar=1.0)
return F.mean(loss, self._batch_axis, exclude=True)
box_loss = SmoothL1Loss()
print(box_loss)
# -
# ### Initialize parameters
import time
from mxnet import autograd as ag
from gluoncv.loss import SSDMultiBoxLoss
# +
# loop params
epochs = 350
start_epoch = 0
# initialize trainer
net.collect_params().reset_ctx(ctx)
trainer = gluon.Trainer(net.collect_params(), 'sgd', {'learning_rate': 1e-1, 'wd': 4e-5})
# evaluation metrics
cls_metric = mx.metric.Accuracy()
box_metric = mx.metric.MAE()
cls_metric_test = mx.metric.Accuracy()
box_metric_test = mx.metric.MAE()
# training loop
for epoch in range(start_epoch, epochs):
# reset iterator and tick
#train_data.reset()
cls_metric.reset()
box_metric.reset()
#tic = time.time()
train_loss = 0
# iterate through all batch
for i, batch in enumerate(train_loader):
tic = time.time()
# record gradients
with ag.record():
x = batch[0].as_in_context(ctx)
y = batch[1].as_in_context(ctx)
lbl = convertlbl(batch[1])
default_anchors, class_predictions, box_predictions = net(x)
box_target, box_mask, cls_target = training_targets(default_anchors, class_predictions, lbl)
# losses
loss1 = cls_loss(class_predictions, cls_target)
loss2 = box_loss(box_predictions, box_target, box_mask)
# sum all losses
loss = loss1 + loss2
train_loss += nd.sum(loss).asscalar()
# backpropagate
loss.backward()
# apply
trainer.step(batch_size, ignore_stale_grad=True)
# update metrics
cls_metric.update([cls_target], [nd.transpose(class_predictions, (0, 2, 1))])
box_metric.update([box_target], [box_predictions * box_mask])
#if (i + 1) % log_interval == 0:
print(time.time()-tic)
#toc = time.time()
name1_train, val1_train = cls_metric.get()
name2_train, val2_train = box_metric.get()
cls_metric_test.reset()
box_metric_test.reset()
tic = time.time()
test_loss = 0
for i, batch in enumerate(test_loader):
# record gradients
x = batch[0].as_in_context(ctx)
y = batch[1].as_in_context(ctx)
lbl = convertlbl(batch[1])
default_anchors, class_predictions, box_predictions = net(x)
box_target, box_mask, cls_target = training_targets(default_anchors, class_predictions, lbl)
# losses
loss1 = cls_loss(class_predictions, cls_target)
loss2 = box_loss(box_predictions, box_target, box_mask)
# sum all losses
loss = loss1 + loss2
test_loss += nd.sum(loss).asscalar()
# update metrics
cls_metric_test.update([cls_target], [nd.transpose(class_predictions, (0, 2, 1))])
box_metric_test.update([box_target], [box_predictions * box_mask])
#if (i + 1) % log_interval == 0:
toc = time.time()
name1_test, val1_test = cls_metric_test.get()
name2_test, val2_test = box_metric_test.get()
print('epoch:%3d;\t train:%.6e;%f;%.6e;\t test:%.6e;%f;%.6e'
%(epoch, train_loss/len(train_dataset), val1_train, val2_train, test_loss/len(test_dataset), val1_test, val2_test))
net.save_parameters('process/ssd_%d.params' % epoch)
# -
# ## Test
# ### Prepare the test data
test_image, test_label = test_dataset[0]
test_image2, test_label2 = train_transform(test_image, test_label)
test_image2 = nd.expand_dims(test_image2,0)
print('tensor shape:', test_image2.shape)
# ### Network inference
anchors, cls_preds, box_preds = net(test_image2.as_in_context(ctx))
# ### Convert predictions to real object detection results
from mxnet.contrib.ndarray import MultiBoxDetection
cls_probs = nd.SoftmaxActivation(nd.transpose(cls_preds, (0, 2, 1)), mode='channel')
output = MultiBoxDetection(cls_prob=cls_probs, loc_pred=box_preds, anchor=anchors, force_suppress=True, clip=True, nms_topk=250)
# ### Display results
# +
class_names = ('cluster')
def display(img, out, thresh=0.5):
import random
import matplotlib as mpl
import numpy as np
mpl.rcParams['figure.figsize'] = (10,10)
img = img.asnumpy()
img = np.transpose(img,(2,3,1,0))
img = np.squeeze(img)
plt.clf()
plt.imshow(img)
for det in out:
cid = int(det[0])
if cid == 0:
continue
score = det[1]
if score < thresh:
continue
scales = [img.shape[1], img.shape[0]] * 2
xmin, ymin, xmax, ymax = [int(p * s) for p, s in zip(det[2:6].tolist(), scales)]
rect = plt.Rectangle((xmin, ymin), xmax - xmin, ymax - ymin, fill=False,
edgecolor='red', linewidth=3)
plt.gca().add_patch(rect)
text = class_names[cid]
plt.gca().text(xmin, ymin-2, '{:s} {:.3f}'.format(text, score),
bbox=dict(facecolor='red', alpha=0.5),
fontsize=12, color='white')
display(test_image2, output[0].asnumpy(), thresh=0.52)
# -
| experiments/ssd512-mobilenet_v2_alpha2_pretrained_from_strach.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Customize your Processing Pipeline
# While *NeuroKit* is designed to be beginner-friendly, experts who desire to have more control over their own processing pipeline are also offered the possibility to tune functions to their specific usage. This example shows how to use NeuroKit to customize your own processing pipeline for advanced users taking ECG processing as an example.
# Load NeuroKit and other useful packages
import neurokit2 as nk
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
# %matplotlib notebook
# %matplotlib inline
sns.set_style('darkgrid')
plt.rcParams['figure.figsize'] = [8, 5] # Bigger images
# ## The Default NeuroKit processing pipeline
# *NeuroKit* provides a very useful set of functions, `*_process()` (e.g. `ecg_process()`, `eda_process()`, `emg_process()`, ...), which are all-in-one functions that cleans, preprocesses and processes the signals. It includes good and sensible defaults that should be suited for most of users and typical use-cases. That being said, in some cases, you might want to have more control over the processing pipeline.
#
# This is how `ecg_process()` is typically used:
# +
# Simulate ecg signal (you can use your own one)
ecg = nk.ecg_simulate(duration=15, sampling_rate=1000, heart_rate=80)
# Default processing pipeline
signals, info = nk.ecg_process(ecg, sampling_rate=1000)
# Visualize
plot = nk.ecg_plot(signals)
# -
# ## Building your own `process()` function
# Now, if you look at the code of [`ecg_process()`](https://github.com/neuropsychology/NeuroKit/blob/master/neurokit2/ecg/ecg_process.py#L49) (see [here](https://neurokit2.readthedocs.io/en/latest/tutorials/understanding.html) for how to explore the code), you can see that it is in fact very simple.
#
# It uses what can be referred to as "mid-level functions", such as `ecg_clean()`, `ecg_peaks()`, `ecg_rate()` etc.
#
# This means that you can basically **re-create** the `ecg_process()` function very easily by calling these mid-level functions:
# Define a new function
def my_processing(ecg_signal):
# Do processing
ecg_cleaned = nk.ecg_clean(ecg, sampling_rate=1000)
instant_peaks, rpeaks, = nk.ecg_peaks(ecg_cleaned, sampling_rate=1000)
rate = nk.ecg_rate(rpeaks, sampling_rate=1000, desired_length=len(ecg_cleaned))
# Prepare output
signals = pd.DataFrame({"ECG_Raw": ecg_signal,
"ECG_Clean": ecg_cleaned,
"ECG_Rate": rate})
signals = pd.concat([signals, instant_peaks], axis=1)
info = rpeaks
return signals, info
# You can now use this function as you would do with `ecg_process()`.
# +
# Process the signal using previously defined function
signals, info = my_processing(ecg)
# Visualize
plot = nk.ecg_plot(signals)
# -
# ## Changing the processing parameters
# Now, you might want to ask, why would you re-create the processing function? Well, it allows you to **change the parameters** of the inside as you please. Let's say you want to use a specific **cleaning** method.
#
# First, let's look at the [documentation for `ecg_clean()`](https://neurokit2.readthedocs.io/en/latest/functions.html#neurokit2.ecg_clean), you can see that they are several different methods for cleaning which can be specified. The default is the **Neurokit** method, however depending on the quality of your signal (and several other factors), other methods may be more appropriate. It is up to you to make this decision.
#
# You can now change the methods as you please for each function in your custom processing function that you have written above:
# Define a new function
def my_processing(ecg_signal):
# Do processing
ecg_cleaned = nk.ecg_clean(ecg, sampling_rate=1000, method="engzeemod2012")
instant_peaks, rpeaks, = nk.ecg_peaks(ecg_cleaned, sampling_rate=1000)
rate = nk.ecg_rate(rpeaks, sampling_rate=1000, desired_length=len(ecg_cleaned))
# Prepare output
signals = pd.DataFrame({"ECG_Raw": ecg_signal,
"ECG_Clean": ecg_cleaned,
"ECG_Rate": rate})
signals = pd.concat([signals, instant_peaks], axis=1)
info = rpeaks
return signals, info
# Similarly, you can select a different method for the peak detection.
# ## Customize even more!
# It is possible that none of these methods suit your needs, or that you want to test a new method. Rejoice yourself, as *NeuroKit* allows you to do that by providing what can be referred to as "low-level" functions.
#
# For instance, you can rewrite the **cleaning** procedure by using the [signal processsing tools](https://neurokit2.readthedocs.io/en/latest/functions.html#general-signal-processing) offered by NeuroKit:
def my_cleaning(ecg_signal, sampling_rate):
detrended = nk.signal_detrend(ecg_signal, order=1)
cleaned = nk.signal_filter(detrended, sampling_rate=sampling_rate, lowcut=2, highcut=9, method='butterworth')
return cleaned
# You can use this function inside your custom processing written above:
# Define a new function
def my_processing(ecg_signal):
# Do processing
ecg_cleaned = my_cleaning(ecg, sampling_rate=1000)
instant_peaks, rpeaks, = nk.ecg_peaks(ecg_cleaned, sampling_rate=1000)
rate = nk.ecg_rate(rpeaks, sampling_rate=1000, desired_length=len(ecg_cleaned))
# Prepare output
signals = pd.DataFrame({"ECG_Raw": ecg_signal,
"ECG_Clean": ecg_cleaned,
"ECG_Rate": rate})
signals = pd.concat([signals, instant_peaks], axis=1)
info = rpeaks
return signals, info
# Congrats, you have created your own processing pipeline! Let's see how it performs:
signals, info = my_processing(ecg)
plot = nk.ecg_plot(signals)
# This doesn't look bad :) **Can you do better?**
| docs/examples/custom.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .fs
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: F#
// language: fsharp
// name: ifsharp
// ---
// WARNING: experimental FSI + .net standard
#load "Paket.fsx"
Paket.Version [
"Microsoft.ML", "0.5"
"NETStandard.Library", "2.0.3"]
// +
//Patch location for native dependencies
System.Environment.SetEnvironmentVariable("Path",
System.Environment.GetEnvironmentVariable("Path") + ";" + Paket.RootPath + @"/packages/Microsoft.ML/runtimes/win-x64/native/")
System.Environment.SetEnvironmentVariable("Path",
System.Environment.GetEnvironmentVariable("Path") + ";" + Paket.RootPath + @"/packages/Microsoft.ML.CpuMath/runtimes/win-x64/native/")
// -
#r "packages/NETStandard.Library/build/netstandard2.0/ref/netstandard"
#load "Paket.Generated.Refs.fsx"
open Microsoft.ML
open Microsoft.ML.Runtime.Api
open Microsoft.ML.Transforms
open Microsoft.ML.Trainers
open System
open Microsoft.ML.Data
Util.Url "https://github.com/dotnet/machinelearning/blob/master/test/data/wikipedia-detox-250-line-data.tsv"
let testDataPath = @"wikipedia-detox-250-line-data.tsv"
// +
type SentimentData() =
[<Column(ordinal = "0", name = "Label"); DefaultValue>]
val mutable Sentiment : float32
[<Column(ordinal = "1"); DefaultValue>]
val mutable SentimentText : string
type SentimentPrediction() =
[<ColumnName "PredictedLabel"; DefaultValue>]
val mutable Sentiment : bool
// -
let pipeline = LearningPipeline()
pipeline.Add(TextLoader(testDataPath).CreateFrom<SentimentData>())
pipeline.Add(TextFeaturizer("Features", "SentimentText"))
pipeline.Add(FastTreeBinaryClassifier(NumLeaves = 5, NumTrees = 5, MinDocumentsInLeafs = 2))
let model = pipeline.Train<SentimentData, SentimentPrediction>()
// +
//let evaluator = new BinaryClassificationEvaluator();
// +
let predictions =
[ SentimentData(SentimentText = "Contoso's 11 is a wonderful experience")
SentimentData(SentimentText = "Oooooh thank you Mr. DietLimeCola. Once again, nice job trying to pretend you have some authority over anybody here. You are a wannabe admin, which is even sadder than a real admin") ]
|> List.map model.Predict
predictions
|> Seq.iter(fun p -> printfn "%b" p.Sentiment)
// +
let predictions =
[ SentimentData(SentimentText = "SOMETHING AWFUL IS DEAD DEAD DEAD DEAD DEAD")]
|> List.map model.Predict
predictions
|> Seq.iter(fun p -> printfn "%b" p.Sentiment)
// -
| OpenFSharpComplete/08_mlnet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Combining node and constraint detection
#
# In this notebook the advances made in the previous chapters are combined to not only show the nodes, but also the constraints of an image.
#
# At first both models have to be loaded.
# +
from os import path
import cv2
import tensorflow as tf
from matplotlib import pyplot as plt
# +
symbol_detector_path = path.join('models', 'devel', 'fcn_sym_det.h5')
symbol_detector = tf.keras.models.load_model(symbol_detector_path)
symbol_detector.summary()
# +
crop_detector_path = path.join('models', 'devel', 'crop_detector.h5')
crop_detector = tf.keras.models.load_model(crop_detector_path)
crop_detector.summary()
# -
# Then images have to be loaded, which are used to test the models.
#
# The second step of the interim data contains images of nodes which are connected with constraints, so 5 images of that dataset are loaded.
# +
image_dir = path.join('data', 'sep_interim_02')
image_paths = [path.join(image_dir, str(i)+'.png') for i in range(5)]
images = [cv2.imread(image, cv2.IMREAD_GRAYSCALE) for image in image_paths]
plt.figure(figsize=(20,20))
for idx, image in enumerate(images):
plt.subplot(1, 5, idx + 1)
plt.set_cmap('gray')
plt.axis('off')
plt.imshow(image)
plt.show()
# +
def get_bounding_boxes_nms(predictions):
sqz = tf.squeeze
max_idx = tf.math.argmax(sqz(predictions), -1)
node_idx = tf.where(tf.equal(max_idx, 1))
base_idx = tf.where(tf.equal(max_idx, 2))
all_idx = tf.concat([node_idx, base_idx], 0)
max_val = tf.math.reduce_max(sqz(predictions), -1)
y, x = tf.split(all_idx * 4, 2, -1)
coords = sqz(tf.stack([y, x, y + 32, x + 32], -1))
all_boxes = tf.cast(coords / 360, tf.float32)
scores = tf.gather_nd(max_val, all_idx)
eps = tf.keras.backend.epsilon()
nms_idx = tf.image.non_max_suppression(all_boxes, scores, 99, eps, 0.8)
limit = tf.cast(tf.math.count_nonzero(node_idx, 0)[0], tf.int32)
mask = tf.less(nms_idx, limit)
node_mask = tf.boolean_mask(nms_idx, mask)
base_mask = tf.boolean_mask(nms_idx,~mask)
node_boxes = tf.gather(all_boxes, node_mask)
base_boxes = tf.gather(all_boxes, base_mask)
return node_boxes, base_boxes
image_tensor = tf.convert_to_tensor(images)
image_tensor = tf.cast(image_tensor / 255, tf.float32)
image_tensor = tf.expand_dims(image_tensor, -1)
node_pred = symbol_detector(image_tensor)
boxes = [get_bounding_boxes_nms(pred) for pred in node_pred]
image_rgb = tf.image.grayscale_to_rgb(image_tensor)
plt.figure(figsize=(20,20))
save_images = []
for idx, image in enumerate(image_rgb):
image = tf.expand_dims(image, 0)
image = tf.image.draw_bounding_boxes(
image, [boxes[idx][0]], [[0, 1, 0]])
image = tf.image.draw_bounding_boxes(
image, [boxes[idx][1]], [[1, 0, 0]])
plt.subplot(1, 5, idx + 1)
plt.set_cmap('hsv')
plt.axis('off')
image = tf.squeeze(image)
plt.imshow(image)
save_images.append(image.numpy())
plt.show()
# -
# The accuracy is okay.
# Considering the fact that there are a lot of fragments in the form of constraints in these images, only one node is detected at a point where no node should be (second image, the left (wrong) node detection).
# The node in the fourth image is actually detected, but the bounding box is not visible for some reason...
#
# The next step is to crop the image between the nodes.
#
# This code has a lot of parts of `2.2.3-extracting_crops.ipynb`, which is not surprising, considering the fact that the same operations have to be performed.
#
# The information needed at this point are the crop images taken from the images, which are in a list of lists, assigning the crops to the image by their index and a corresponding list containing the coordinates of the crops to be able to identify the position of the prediction later on.
#
# To be able to perform the cropping in the first place the position of the nodes has to be identified.
# We already have the information needed to generate bounding boxes.
# The middle point of each bounding box is therefore considered to be the center of the nodes.
# +
box_coords = []
for box in boxes:
blob = tf.concat([box[0], box[1]], 0)
blob *= 360
blob = tf.cast(blob, tf.int32)
box_coords.append(
[[int((i[0]+i[2])/2),int((i[1]+i[3])/2)] for i in blob])
crops = []
crops_info = []
for idx, coords in enumerate(box_coords):
image_crop = []
image_info = []
for node1 in coords:
for node2 in coords:
x1 = min(node1[1], node2[1])
y1 = min(node1[0], node2[0])
x2 = max(node1[1], node2[1])
y2 = max(node1[0], node2[0])
# Remove images with area 0
if x1 == x2 or y1 == y2:
continue
crop = images[idx][y1:y2, x1:x2]
crop = cv2.resize(crop, (96, 96))
if x1 == node2[1]:
x1 = node1[1]
x2 = node2[1]
crop = cv2.flip(crop, 1)
if y1 == node2[0]:
y1 = node1[0]
y2 = node2[0]
crop = cv2.flip(crop, 0)
crop = tf.expand_dims(crop, -1)
crop = tf.cast(crop / 255, tf.float32)
image_crop.append(crop)
image_info.append([x1, y1, x2, y2])
crops.append(image_crop)
crops_info.append(image_info)
crop_pred = [crop_detector(tf.convert_to_tensor(crop)) for crop in crops]
# +
for idx, pred in enumerate(crop_pred):
for jdx, argmax in enumerate(tf.argmax(pred, -1)):
if argmax == 0:
continue
x1, y1, x2, y2 = crops_info[idx][jdx]
color = (0, int(argmax - 1), 1)
cv2.arrowedLine(save_images[idx], (x1, y1), (x2, y2),color , 3)
plt.figure(figsize=(20,20))
save_path = path.join('reports', 'sep', 'images')
for idx, image in enumerate(save_images):
plt.subplot(1, 5, idx + 1)
plt.set_cmap('hsv')
plt.axis('off')
plt.imshow(image)
image_path = path.join(save_path, '225_' + str(idx) + '.png')
cv2.imwrite(image_path, 255 - image * 255)
plt.show()
# -
# The results do seem to be okay, actually.
# Most constraints are detected and where they are not, the environment is often cluttered.
# The performance is not perfect though, but as a first proof of concept this should suffice.
#
# A take away is that crop detection relies heavily on the performance of the node detection, because nodes which are not detected are not taken into account when creating the data for the `crop_detector`.
#
# Falsely predicted nodes are most likely not going to result in a pair of nodes connected by constraints, but they are slowing the process down significantly, since the number of crops grows exponentially to the number of nodes.
# By taking the coordinates of the nodes and subsequentially pairs of nodes which share a constraint, all necessary information is needed to be able to create a model of a mechanism.
| reports/sep/notebooks/2.2.5-combining_node_constraints.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="TA21Jo5d9SVq"
#
#
# 
#
# [](https://colab.research.google.com/github/JohnSnowLabs/spark-nlp-workshop/blob/master/tutorials/streamlit_notebooks/healthcare/NER_LEGAL_DE.ipynb)
#
#
#
# + [markdown] id="CzIdjHkAW8TB"
# # **Detect legal entities in German**
# + [markdown] id="6uDmeHEFW7_h"
# To run this yourself, you will need to upload your license keys to the notebook. Otherwise, you can look at the example outputs at the bottom of the notebook. To upload license keys, open the file explorer on the left side of the screen and upload `workshop_license_keys.json` to the folder that opens.
# + [markdown] id="wIeCOiJNW-88"
# ## 1. Colab Setup
# + [markdown] id="HMIDv74CYN0d"
# Import license keys
# + id="ttHPIV2JXbIM" executionInfo={"status": "ok", "timestamp": 1601207660130, "user_tz": -300, "elapsed": 1351, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10508284328555930330"}} outputId="91f738d9-7e66-4144-d236-a5da9117e038" colab={"base_uri": "https://localhost:8080/", "height": 51}
import os
import json
with open('/content/spark_nlp_for_healthcare.json', 'r') as f:
license_keys = json.load(f)
license_keys.keys()
secret = license_keys['SECRET']
os.environ['SPARK_NLP_LICENSE'] = license_keys['SPARK_NLP_LICENSE']
os.environ['AWS_ACCESS_KEY_ID'] = license_keys['AWS_ACCESS_KEY_ID']
os.environ['AWS_SECRET_ACCESS_KEY'] = license_keys['AWS_SECRET_ACCESS_KEY']
sparknlp_version = license_keys["PUBLIC_VERSION"]
jsl_version = license_keys["JSL_VERSION"]
print ('SparkNLP Version:', sparknlp_version)
print ('SparkNLP-JSL Version:', jsl_version)
# + [markdown] id="rQtc1CHaYQjU"
# Install dependencies
# + id="CGJktFHdHL1n" executionInfo={"status": "ok", "timestamp": 1601207726887, "user_tz": -300, "elapsed": 66146, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10508284328555930330"}} outputId="678fab45-2d1c-4b96-f55e-5a1f2b499fb8" colab={"base_uri": "https://localhost:8080/", "height": 326}
# Install Java
# ! apt-get update -qq
# ! apt-get install -y openjdk-8-jdk-headless -qq > /dev/null
# ! java -version
# Install pyspark
# ! pip install --ignore-installed -q pyspark==2.4.4
# Install Spark NLP
# ! pip install --ignore-installed spark-nlp==$sparknlp_version
# ! python -m pip install --upgrade spark-nlp-jsl==$jsl_version --extra-index-url https://pypi.johnsnowlabs.com/$secret
# + [markdown] id="Hj5FRDV4YSXN"
# Import dependencies into Python and start the Spark session
# + id="sw-t1zxlHTB7" executionInfo={"status": "ok", "timestamp": 1601207746177, "user_tz": -300, "elapsed": 83597, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10508284328555930330"}}
os.environ['JAVA_HOME'] = "/usr/lib/jvm/java-8-openjdk-amd64"
os.environ['PATH'] = os.environ['JAVA_HOME'] + "/bin:" + os.environ['PATH']
import pandas as pd
from pyspark.ml import Pipeline
from pyspark.sql import SparkSession
import pyspark.sql.functions as F
import sparknlp
from sparknlp.annotator import *
from sparknlp_jsl.annotator import *
from sparknlp.base import *
import sparknlp_jsl
spark = sparknlp_jsl.start(secret)
# + [markdown] id="9RgiqfX5XDqb"
# ## 2. Construct the pipeline
#
# For more details: https://github.com/JohnSnowLabs/spark-nlp-models#pretrained-models---spark-nlp-for-healthcare
# + id="LLuDz_t40be4" executionInfo={"status": "ok", "timestamp": 1601207886961, "user_tz": -300, "elapsed": 140775, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10508284328555930330"}} outputId="44bea217-d857-4abd-ca07-6261b5063b90" colab={"base_uri": "https://localhost:8080/", "height": 119}
document_assembler = DocumentAssembler() \
.setInputCol('text')\
.setOutputCol('document')
sentence_detector = SentenceDetector() \
.setInputCols(['document'])\
.setOutputCol('sentence')
tokenizer = Tokenizer()\
.setInputCols(['sentence']) \
.setOutputCol('token')
# German word embeddings
word_embeddings = WordEmbeddingsModel.pretrained('w2v_cc_300d','de', 'clinical/models') \
.setInputCols(["sentence", 'token'])\
.setOutputCol("embeddings")
# German NER model
clinical_ner = NerDLModel.pretrained('ner_legal','de', 'clinical/models') \
.setInputCols(["sentence", "token", "embeddings"]) \
.setOutputCol("ner")
ner_converter = NerConverter()\
.setInputCols(['sentence', 'token', 'ner']) \
.setOutputCol('ner_chunk')
nlp_pipeline = Pipeline(stages=[
document_assembler,
sentence_detector,
tokenizer,
word_embeddings,
clinical_ner,
ner_converter])
empty_df = spark.createDataFrame([['']]).toDF("text")
pipeline_model = nlp_pipeline.fit(empty_df)
light_pipeline = LightPipeline(pipeline_model)
# + [markdown] id="2Y9GpdJhXIpD"
# ## 3. Create example inputs
# + id="vBOKkB2THdGI" executionInfo={"status": "ok", "timestamp": 1601207886963, "user_tz": -300, "elapsed": 140772, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10508284328555930330"}}
# Enter examples as strings in this array
input_list = [
"""Dementsprechend hat der Bundesgerichtshof mit Beschluss vom 24 August 2017 ( - III ZA 15/17 - ) das bei ihm von der Antragstellerin anhängig gemachte „ Prozesskostenhilfeprüfungsverfahre“ an das Bundesarbeitsgericht abgegeben. 2 Die Antragstellerin hat mit Schriftsatz vom 21 März 2016 und damit mehr als sechs Monate vor der Anbringung des Antrags auf Gewährung von Prozesskostenhilfe für die beabsichtigte Klage auf Entschädigung eine Verzögerungsrüge iSv § 198 Abs 5 Satz 1 GVG erhoben. 3 Nach § 198 Abs 1 Satz 1 GVG wird angemessen entschädigt , wer infolge unangemessener Dauer eines Gerichtsverfahrens als Verfahrensbeteiligter einen Nachteil erleidet. a ) Die Angemessenheit der Verfahrensdauer richtet sich gemäß § 198 Abs 1 Satz 2 GVG nach den Umständen des Einzelfalls , insbesondere nach der Schwierigkeit und Bedeutung des Verfahrens sowie nach dem Verhalten der Verfahrensbeteiligten und Dritter. Hierbei handelt es sich um eine beispielhafte , nicht abschließende Auflistung von Umständen , die für die Beurteilung der Angemessenheit besonders bedeutsam sind ( BT-Drs 17/3802 S 18 ). Weitere gewichtige Beurteilungskriterien sind die Verfahrensführung durch das Gericht sowie die zur Verfahrensbeschleunigung gegenläufigen Rechtsgüter der Gewährleistung der inhaltlichen Richtigkeit von Entscheidungen , der Beachtung der richterlichen Unabhängigkeit und des gesetzlichen Richters.""",
]
# + [markdown] id="mv0abcwhXWC-"
# ## 4. Use the pipeline to create outputs
# + id="TK1DB9JZaPs3" executionInfo={"status": "ok", "timestamp": 1601207887485, "user_tz": -300, "elapsed": 141292, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10508284328555930330"}}
df = spark.createDataFrame(pd.DataFrame({"text": input_list}))
result = pipeline_model.transform(df)
# + [markdown] id="UQY8tAP6XZJL"
# ## 5. Visualize results
# + [markdown] id="hnsMLq9gctSq"
# Visualize outputs as data frame
# + id="Ar32BZu7J79X" executionInfo={"status": "ok", "timestamp": 1601207893799, "user_tz": -300, "elapsed": 147599, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10508284328555930330"}} outputId="c06e4003-6610-40b9-d49a-83f51dd8104a" colab={"base_uri": "https://localhost:8080/", "height": 221}
exploded = F.explode(F.arrays_zip('ner_chunk.result', 'ner_chunk.metadata'))
select_expression_0 = F.expr("cols['0']").alias("chunk")
select_expression_1 = F.expr("cols['1']['entity']").alias("ner_label")
result.select(exploded.alias("cols")) \
.select(select_expression_0, select_expression_1).show(truncate=False)
result = result.toPandas()
# + [markdown] id="1wdVmoUcdnAk"
# Functions to display outputs as HTML
# + id="tFeu7loodcQQ" executionInfo={"status": "ok", "timestamp": 1601208025823, "user_tz": -300, "elapsed": 1275, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10508284328555930330"}}
from IPython.display import HTML, display
import random
def get_color():
r = lambda: random.randint(128,255)
return "#%02x%02x%02x" % (r(), r(), r())
def annotation_to_html(full_annotation):
ner_chunks = full_annotation[0]['ner_chunk']
text = full_annotation[0]['document'][0].result
label_color = {}
for chunk in ner_chunks:
label_color[chunk.metadata['entity']] = get_color()
html_output = "<div>"
pos = 0
for n in ner_chunks:
if pos < n.begin and pos < len(text):
html_output += f"<span class=\"others\">{text[pos:n.begin]}</span>"
pos = n.end + 1
html_output += f"<span class=\"entity-wrapper\" style=\"color: black; background-color: {label_color[n.metadata['entity']]}\"> <span class=\"entity-name\">{n.result}</span> <span class=\"entity-type\">[{n.metadata['entity']}]</span></span>"
if pos < len(text):
html_output += f"<span class=\"others\">{text[pos:]}</span>"
html_output += "</div>"
display(HTML(html_output))
# + [markdown] id="-piHygJ6dpEa"
# Display example outputs as HTML
# + id="AtbhE24VeG_C" executionInfo={"status": "ok", "timestamp": 1601208028802, "user_tz": -300, "elapsed": 2087, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10508284328555930330"}} outputId="3f23d0b6-7f95-401f-c3d7-c10c4ad0ffae" colab={"base_uri": "https://localhost:8080/", "height": 187}
for example in input_list:
annotation_to_html(light_pipeline.fullAnnotate(example))
| tutorials/streamlit_notebooks/healthcare/NER_LEGAL_DE.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# + [markdown] language="text"
# # # GWAS exploration
# #
# # Polynote notebook for manual exploratory analysis of VEP annotations and vcf files
# #
# #
# + cell.metadata.exec_info={"endTs": 1609327031758, "startTs": 1609327031634} language="scala"
# import org.apache.spark._
# import org.apache.spark.sql.{DataFrame, Encoders, SparkSession}
# import org.apache.spark.sql.types.StructType
# import scala.reflect.runtime.universe._
# import org.apache.spark.storage.StorageLevel
# import org.apache.spark.rdd._
# import org.apache.spark.sql.functions._
# import org.apache.spark.sql.ColumnName
# + cell.metadata.exec_info={"endTs": 1609327033378, "startTs": 1609327033333} language="scala"
# import better.files._
# import File._
# import java.io.{File => JFile}
# + cell.metadata.exec_info={"endTs": 1609327034449, "startTs": 1609327034405} language="scala"
# import org.apache.spark.sql.expressions._
# import group.research.aging.spark.extensions._
# import group.research.aging.spark.extensions.functions._
# + cell.metadata.exec_info={"endTs": 1609327036436, "startTs": 1609327036380} language="scala"
# def display(f: File) = f.children.foreach(println)
# + cell.metadata.exec_info={"endTs": 1609327044971, "startTs": 1609327038605} language="scala"
# val data = File("/data")
# val gwas_path = data / "gwas" / "anton"
# display(gwas_path)
# + cell.metadata.exec_info={"endTs": 1609327045216, "startTs": 1609327044974} language="scala"
# val dante = gwas_path / "dante"
# display(dante)
# + [markdown] cell.metadata.exec_info={"endTs": 1609215130874, "startTs": 1609215130858} language="text"
# # Exploring variants
# # ==================
# + cell.metadata.exec_info={"endTs": 1609327045459, "startTs": 1609327045218} language="scala"
# val variants = gwas_path / "variants"
# display(variants)
# + cell.metadata.exec_info={"endTs": 1609327047397, "startTs": 1609327047353} language="scala"
# import org.apache.spark.SparkContext
# import org.bdgenomics.adam.rdd.ADAMContext._
# import org.bdgenomics.adam.rdd.read.AlignmentDataset
#
# + [markdown] language="text"
# # # DANTE
# #
# # let's check DANTE variants first
# #
# #
# #
# + [markdown] language="text"
# # ## SNP variants
# #
# #
# + cell.metadata.exec_info={"endTs": 1609218317052, "startTs": 1609218316739} language="scala"
# val dante_variants = dante / "result_variation"
# display(dante_variants)
# + cell.metadata.exec_info={"endTs": 1609327148572, "startTs": 1609327144157} language="scala"
# val dante_snp_annotations = spark.readTSV( (dante_variants / "snp" / "750018002018_WGZ.snp.annot.csv").toString, sep=",", header = true)
# dante_snp_annotations.show(10,1000)
# + language="scala"
# //TODO: do some exploration
# + [markdown] language="text"
# # # Our Variants
# #
# # Now let's check variants computed by our pipeline
# #
# #
# #
# + cell.metadata.exec_info={"endTs": 1609327111714, "startTs": 1609327110699} language="scala"
# val smoove = variants / "smoove" / "antonkulaga-smoove.vcf.gz"
# val cnv = spark.sparkContext.loadVcf(smoove.toString)
# cnv
# + cell.metadata.exec_info={"endTs": 1609327117868, "startTs": 1609327114414} language="scala"
# cnv.dataset.show(10,1000)
# + [markdown] language="text"
# # # Exploring annotations
# #
# # exploring VEP annotations
# #
# #
# #
# + cell.metadata.exec_info={"endTs": 1609327170524, "startTs": 1609327169399} language="scala"
# val gene_names: DataFrame = spark.readTSV( "/data/sources/yspecies/data/input/genes/reference_genes.tsv", header = true)
# .withColumnRenamed("gene", "Gene")
# println(gene_names.count)
# gene_names.show(10,1000)
# + cell.metadata.exec_info={"endTs": 1609327176093, "startTs": 1609327176022} language="scala"
# def with_names(df: DataFrame, how: String="inner") = df.join(gene_names, Seq("Gene"), how)
# + cell.metadata.exec_info={"endTs": 1609243961510, "startTs": 1609243961197} language="scala"
# val anno_path = gwas_path / "vep"
# display(anno_path)
# + [markdown] language="text"
# # # SNPs
# #
# #
# + cell.metadata.exec_info={"endTs": 1609327182804, "startTs": 1609327182736} language="scala"
# display(anno_path / "strelka" / "annotations")
# + cell.metadata.exec_info={"endTs": 1609327196209, "startTs": 1609327196059} language="scala"
# val columns = List("Uploaded_variation", "Location", "Allele", "Gene", "Feature", "Feature_type", "Consequence", "cDNA_position", "CDS_position", "Protein_position", "Amino_acids", "Codons", "Existing_variation", "IND", "ZYG", "IMPACT", "DISTANCE", "STRAND", "FLAGS", "VARIANT_CLASS", "SYMBOL", "SYMBOL_SOURCE", "HGNC_ID", "BIOTYPE", "CANONICAL", "MANE", "TSL", "APPRIS", "CCDS", "ENSP", "SWISSPROT", "TREMBL", "UNIPARC", "UNIPROT_ISOFORM", "GENE_PHENO", "SIFT", "PolyPhen", "EXON", "INTRON", "DOMAINS", "miRNA", "HGVSc", "HGVSp", "HGVS_OFFSET", "AF", "AFR_AF", "AMR_AF", "EAS_AF", "EUR_AF", "SAS_AF", "AA_AF", "EA_AF", "gnomAD_AF", "gnomAD_AFR_AF", "gnomAD_AMR_AF", "gnomAD_ASJ_AF", "gnomAD_EAS_AF", "gnomAD_FIN_AF", "gnomAD_NFE_AF", "gnomAD_OTH_AF", "gnomAD_SAS_AF", "MAX_AF", "MAX_AF_POPS", "CLIN_SIG", "SOMATIC", "PHENO", "PUBMED", "MOTIF_NAME", "MOTIF_POS", "HIGH_INF_POS", "MOTIF_SCORE_CHANGE", "TRANSCRIPTION_FACTORS", "G2P_complete", "G2P_flag", "G2P_gene_req")
# + cell.metadata.exec_info={"endTs": 1609327222956, "startTs": 1609327197241} language="scala"
# val snp_path = "/data/cromwell-executions/annotations/c2db854a-0256-4598-8497-bb5a0e69d528/call-vep_annotation/execution/antonkulaga_variant_annotations.tsv"
# val annotations = spark.readTSV(snp_path).toDF(columns:_*)
# annotations.show(10,10000)
# + cell.metadata.exec_info={"endTs": 1609257282983, "startTs": 1609257282791} language="scala"
# annotations.select("G2P_flag").distinct
# + [markdown] language="text"
# # ## STRUCTURAL
# #
# #
# #
# + cell.metadata.exec_info={"endTs": 1609219125282, "startTs": 1609219122181} language="scala"
# val annotations = spark.readTSV( (anno_path / "test" / "antonkulaga_variant_annotations.tsv").toString, header = false).toDF(cols:_*)
# annotations.show(10)
# + cell.metadata.exec_info={"endTs": 1609262552412, "startTs": 1609262551742} language="scala"
# annotations.where($"gnomAD_AMR_AF" =!= "-").show(10,1000)
# + cell.metadata.exec_info={"endTs": 1609111180794, "startTs": 1609111179211} language="scala"
#
# val counts = annotations.select("Consequence", "Gene").groupBy("Consequence").agg(count($"Gene").as("count"))
#
# counts.sort($"count".desc).show(1000,1000)
# + [markdown] language="text"
# # GET SNP ids
# + language="scala"
# annotations
# + cell.metadata.exec_info={"endTs": 1609219718336, "startTs": 1609219715889} language="scala"
# val frameshift = annotations.where($"Consequence".contains("frameshift"))
# println(frameshift.count)
# frameshift.show(1000,1000)
# + language="scala"
# frameshift
# + [markdown] language="text"
# # Clinical
# + cell.metadata.exec_info={"endTs": 1609111554983, "startTs": 1609111553781} language="scala"
# val clinical = annotations.where($"CLIN_SIG" =!= "-")
# println(clinical.count)
# clinical.show(10,1000)
# + language="scala"
#
| gwas-notebooks/gwas_exploration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="tlG7SA7wnwnW" colab_type="text"
# Lambda School Data Science, Unit 2: Predictive Modeling
#
# # Regression & Classification, Module 2
#
# ## Objectives
# - Go from simple regression (1 feature) to multiple regression (2+ features)
# - Use regression metrics: MAE (Mean Absolute Error), RMSE (Root Mean Squared Error), and $R^2$ Score
# - Understand how ordinary least squares regression minimizes the sum of squared errors
# - Get and plot coefficients
# - Explain why overfitting is a problem. Do train/test split
# + [markdown] id="VJKr75uEnwnb" colab_type="text"
# ## Setup
# + id="kH-AoJbTnwne" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="db440528-2551-43e0-c885-9e1d855b1554"
# If you're in Colab...
import os, sys
in_colab = 'google.colab' in sys.modules
if in_colab:
# Install required python packages:
# pandas-profiling, version >= 2.0
# plotly, version >= 4.0
# !pip install --upgrade pandas-profiling plotly
# Pull files from Github repo
os.chdir('/content')
# !git init .
# !git remote add origin https://github.com/LambdaSchool/DS-Unit-2-Regression-Classification.git
# !git pull origin master
# Change into directory for module
os.chdir('module2')
# + id="wS565BWtnwnu" colab_type="code" colab={}
# Ignore this Numpy warning when using Plotly Express:
# FutureWarning: Method .ptp is deprecated and will be removed in a future version. Use numpy.ptp instead.
import warnings
warnings.filterwarnings(action='ignore', category=FutureWarning, module='numpy')
# + id="Bt3BVNYTnwn2" colab_type="code" colab={}
# Use these functions later
# %matplotlib inline
from matplotlib.patches import Rectangle
import matplotlib.pyplot as plt
import numpy as np
import plotly.express as px
import plotly.graph_objs as go
from sklearn.linear_model import LinearRegression
from sklearn.metrics import mean_absolute_error, mean_squared_error, r2_score
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import PolynomialFeatures
from sklearn.linear_model import LinearRegression
def regression_3d(df, x, y, z, **kwargs):
"""
Visualize linear regression in 3D: 2 features + 1 target
df : Pandas DataFrame
x : string, feature 1 column in df
y : string, feature 2 column in df
z : string, target column in df
"""
# Plot data
fig = px.scatter_3d(df, x, y, z, **kwargs)
# Fit Linear Regression
features = [x, y]
target = z
model = LinearRegression()
model.fit(df[features], df[target])
# Define grid of four points in the feature space
xmin, xmax = df[x].min(), df[x].max()
ymin, ymax = df[y].min(), df[y].max()
coords = [[xmin, ymin],
[xmin, ymax],
[xmax, ymin],
[xmax, ymax]]
# Make predictions for the grid
Z = model.predict(coords).reshape((2,2), order='F')
# Plot predictions as a 3D surface (plane)
fig.add_trace(go.Surface(x=[xmin,xmax], y=[ymin,ymax], z=Z))
return fig
def regression_residuals(df, feature, target, m, b):
"""
Visualize linear regression, with residual errors,
in 2D: 1 feature + 1 target.
Use the m & b parameters to "fit the model" manually.
df : Pandas DataFrame
feature : string, feature column in df
target : string, target column in df
m : numeric, slope for linear equation
b : numeric, intercept for linear requation
"""
# Plot data
df.plot.scatter(feature, target)
# Make predictions
x = df[feature]
y = df[target]
y_pred = m*x + b
# Plot predictions
plt.plot(x, y_pred)
# Plot residual errors
for x, y1, y2 in zip(x, y, y_pred):
plt.plot((x, x), (y1, y2), color='grey')
# Print regression metrics
mae = mean_absolute_error(y, y_pred)
r2 = r2_score(y, y_pred)
print('Mean Absolute Error:', mae)
print('R^2:', r2)
def regression_squared_errors(df, feature, target, m, b):
"""
Visualize linear regression, with squared errors,
in 2D: 1 feature + 1 target.
Use the m & b parameters to "fit the model" manually.
df : Pandas DataFrame
feature : string, feature column in df
target : string, target column in df
m : numeric, slope for linear equation
b : numeric, intercept for linear requation
"""
# Plot data
fig = plt.figure(figsize=(7,7))
ax = plt.axes()
df.plot.scatter(feature, target, ax=ax)
# Make predictions
x = df[feature]
y = df[target]
y_pred = m*x + b
# Plot predictions
ax.plot(x, y_pred)
# Plot squared errors
xmin, xmax = ax.get_xlim()
ymin, ymax = ax.get_ylim()
scale = (xmax-xmin)/(ymax-ymin)
for x, y1, y2 in zip(x, y, y_pred):
bottom_left = (x, min(y1, y2))
height = abs(y1 - y2)
width = height * scale
ax.add_patch(Rectangle(xy=bottom_left, width=width, height=height, alpha=0.1))
# Print regression metrics
mse = mean_squared_error(y, y_pred)
rmse = np.sqrt(mse)
mae = mean_absolute_error(y, y_pred)
r2 = r2_score(y, y_pred)
print('Mean Squared Error:', mse)
print('Root Mean Squared Error:', rmse)
print('Mean Absolute Error:', mae)
print('R^2:', r2)
# Credit: <NAME>, Python Data Science Handbook, Chapter 5.3
# https://jakevdp.github.io/PythonDataScienceHandbook/05.03-hyperparameters-and-model-validation.html#Validation-curves-in-Scikit-Learn
def PolynomialRegression(degree=2, **kwargs):
return make_pipeline(PolynomialFeatures(degree),
LinearRegression(**kwargs))
# + [markdown] id="PcY0iIBSnwn8" colab_type="text"
# # Predict Elections! 🗳️
# + id="8AtWb3Eynwn9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 617} outputId="aa2db861-d02d-45c7-f0bb-740d5b7d63a2"
import pandas as pd
import plotly.express as px
df = pd.read_csv('../data/bread_peace_voting.csv')
px.scatter(
df,
x='Average Recent Growth in Personal Incomes',
y='Incumbent Party Vote Share',
text='Year',
title='US Presidential Elections, 1952-2016',
trendline='ols',
)
# + [markdown] id="M2nMwiEknwoB" colab_type="text"
# #### <NAME>, [Background Information on the ‘Bread and Peace’ Model of Voting in Postwar US Presidential Elections](https://douglas-hibbs.com/background-information-on-bread-and-peace-voting-in-us-presidential-elections/)
#
# > Aggregate two-party vote shares going to candidates of the party holding the presidency during the postwar era are well explained by just two fundamental determinants:
#
# > (1) Positively by weighted-average growth of per capita real disposable personal income over the term.
# > (2) Negatively by cumulative US military fatalities (scaled to population) owing to unprovoked, hostile deployments of American armed forces in foreign wars.
#
# #### Data sources
# - 1952-2012: <NAME>, [2014 lecture at Deakin University Melbourne](http://www.douglas-hibbs.com/HibbsArticles/HIBBS-PRESVOTE-SLIDES-MELBOURNE-Part1-2014-02-26.pdf), Slide 40
# - 2016, Vote Share: [The American Presidency Project](https://www.presidency.ucsb.edu/statistics/elections)
# - 2016, Recent Growth in Personal Incomes: [The 2016 election economy: the "Bread and Peace" model final forecast](https://angrybearblog.com/2016/11/the-2016-election-economy-the-bread-and-peace-model-final-forecast.html)
# - 2016, US Military Fatalities: Assumption that Afghanistan War fatalities in 2012-16 occured at the same rate as 2008-12
#
# > Fatalities denotes the cumulative number of American military fatalities per millions of US population the in Korea, Vietnam, Iraq and Afghanistan wars during the presidential terms preceding the 1952, 1964, 1968, 1976 and 2004, 2008 and 2012 elections. —[Hibbs](http://www.douglas-hibbs.com/HibbsArticles/HIBBS-PRESVOTE-SLIDES-MELBOURNE-Part1-2014-02-26.pdf), Slide 33
# + id="XUMDm5OGnwoD" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 593} outputId="f1290928-1dd6-411e-dd24-165e459db4fb"
df
# + [markdown] id="FjP9pj8zd4Sn" colab_type="text"
# What's the average incumbent Party Vote Share?
# + id="pAduKp3Ydk0a" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="3d47d03b-d596-4355-e811-3b3addf9d651"
# More likely to win as incumbent
df['Incumbent Party Vote Share'].mean()
# + id="BE2t6_sPeOQd" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 617} outputId="26c57d46-84cd-46ca-cccb-c2c26f41912e"
px.scatter_3d(
df,
x = 'Average Recent Growth in Personal Incomes',
y = 'US Military Fatalities per Million',
z = 'Incumbent Party Vote Share',
text = 'Year'
)
# + id="T6JZ3qOjfbUm" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 617} outputId="0e6c99a7-6d81-460a-8b1d-90b2b2822237"
import itertools
import numpy as np
import plotly.express as px
import plotly.graph_objs as go
from sklearn.linear_model import LinearRegression
def regression_3d(df, x, y, z, num=100, **kwargs):
"""
Visualize linear regression in 3D: 2 features + 1 target
df : Pandas DataFrame
x : string, feature 1 column in df
y : string, feature 2 column in df
z : string, target column in df
num : integer, number of quantiles for each feature
"""
# Plot data
fig = px.scatter_3d(df, x, y, z, **kwargs)
# Fit Linear Regression
features = [x, y]
target = z
model = LinearRegression()
model.fit(df[features], df[target])
# Define grid of coordinates in the feature space
xmin, xmax = df[x].min(), df[x].max()
ymin, ymax = df[y].min(), df[y].max()
xcoords = np.linspace(xmin, xmax, num)
ycoords = np.linspace(ymin, ymax, num)
coords = list(itertools.product(xcoords, ycoords))
# Make predictions for the grid
predictions = model.predict(coords)
Z = predictions.reshape(num, num).T
# Plot predictions as a 3D surface (plane)
fig.add_trace(go.Surface(x=xcoords, y=ycoords, z=Z))
return fig
regression_3d(
df,
x = 'Average Recent Growth in Personal Incomes',
y = 'US Military Fatalities per Million',
z = 'Incumbent Party Vote Share'
)
# + id="AB_gFS4agH_z" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="72ee61c7-d2ab-4c08-b4ae-df4318440a5f"
# 1. Import the appropriate estimator class from Scikit-Learn
from sklearn.linear_model import LinearRegression
# 2. Instantiate this class
model = LinearRegression()
# 3. Arrange X features matrix & y target vector
features = ['Average Recent Growth in Personal Incomes', 'US Military Fatalities per Million']
target = 'Incumbent Party Vote Share'
X = df[features]
y = df[target]
# 4. Fit the model
model.fit(X, y)
# + id="wwElDGk6heGN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="1fff74ef-958b-4874-ad71-652402e77045"
model.intercept_
# + id="Ndma4RiwiSms" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="7f4fd3c5-d568-4bb9-d5fc-ce5dab8ff946"
model.coef_
# + [markdown] id="TuXAePSjidLO" colab_type="text"
# y= coef1* x1 + coef2* x2 + b
# + id="wuTpUyUtiVbp" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="72be4948-6a1b-45aa-a02c-260ae9ac4546"
model.predict([[0,0]])
# + id="7EKDiiYxiwRS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="e1fa2f18-6f61-4ff6-f9a7-2cd86a8d3a6c"
model.predict([[1,0]])
# + id="eBUK63rii1S7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="219ab25a-ab35-4704-b26a-c6259c68f67a"
model.predict([[1,0]]) - model.predict([[0,0]])
# + id="c2LSN5jqi-Eh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="41116348-c3b2-4c48-ff1d-4978c9e492e1"
model.predict([[2,0]]) - model.predict([[1,0]])
# + id="MuWHlAlKjEu9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 610} outputId="fa078eb9-ca72-4911-d3a4-18faba6e799a"
y_pred = model.predict(X)
df['Predicted'] = y_pred
df['Error'] = y_pred - y
df
# + id="xLbaIyrtjk6P" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="eb5cfa5c-f481-46d7-ee97-484097551c31"
df['Error'].mean()
# + id="7-Y7uq-uj3cG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="0a43ba82-f6a0-458c-cfc2-cac8c1f826ad"
df['Error'].abs().mean()
# + id="HMGBliNdkKMM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="08383c0f-e745-4623-e51e-b7145e403312"
# Mean Absolute Error(MAE)
from sklearn.metrics import mean_absolute_error
mean_absolute_error(y, y_pred)
# + id="ZFp6AZugkbVW" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="c198f601-06d0-4054-f9bf-a866dd40f79f"
# Average vote share
y.mean()
# + id="lYH2Bsk5koPK" colab_type="code" colab={}
# Guess this for each election
#[42] * 10
guesses = [y.mean()] * len(y)
# + id="iYiK4PPRk998" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="4886c97c-6ecd-440e-8610-06f7cb0b37e6"
# How far off is this?
mean_absolute_error(y, guesses)
# + id="9N0vd0_DlS86" colab_type="code" colab={}
import matplotlib.pyplot as plt
from sklearn.metrics import mean_absolute_error, r2_score
def regression_residuals(df, feature, target, m, b):
"""
Visualize linear regression, with residual errors,
in 2D: 1 feature + 1 target.
Use the m & b parameters to "fit the model" manually.
df : Pandas DataFrame
feature : string, feature column in df
target : string, target column in df
m : numeric, slope for linear equation
b : numeric, intercept for linear requation
"""
# Plot data
df.plot.scatter(feature, target)
# Make predictions
x = df[feature]
y = df[target]
y_pred = m*x + b
# Plot predictions
plt.plot(x, y_pred)
# Plot residual errors
for x, y1, y2 in zip(x, y, y_pred):
plt.plot((x, x), (y1, y2), color='grey')
# Print regression metrics
mae = mean_absolute_error(y, y_pred)
r2 = r2_score(y, y_pred)
print('Mean Absolute Error:', mae)
print('R^2:', r2)
# + id="pBgz3Tw6lUpy" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 317} outputId="1242c342-0ba8-4e6f-8656-e00697832b65"
feature = 'Average Recent Growth in Personal Incomes'
regression_residuals(df, feature,target,m=0, b=y.mean())
# + id="g4BP_yq6mL6w" colab_type="code" colab={}
# + id="EZF26k9Tlmw0" colab_type="code" colab={}
from matplotlib.patches import Rectangle
import matplotlib.pyplot as plt
import numpy as np
from sklearn.metrics import mean_absolute_error, mean_squared_error, r2_score
def regression_squared_errors(df, feature, target, m, b):
"""
Visualize linear regression, with squared errors,
in 2D: 1 feature + 1 target.
Use the m & b parameters to "fit the model" manually.
df : Pandas DataFrame
feature : string, feature column in df
target : string, target column in df
m : numeric, slope for linear equation
b : numeric, intercept for linear requation
"""
# Plot data
fig = plt.figure(figsize=(7,7))
ax = plt.axes()
df.plot.scatter(feature, target, ax=ax)
# Make predictions
x = df[feature]
y = df[target]
y_pred = m*x + b
# Plot predictions
ax.plot(x, y_pred)
# Plot squared errors
xmin, xmax = ax.get_xlim()
ymin, ymax = ax.get_ylim()
scale = (xmax-xmin)/(ymax-ymin)
for x, y1, y2 in zip(x, y, y_pred):
bottom_left = (x, min(y1, y2))
height = abs(y1 - y2)
width = height * scale
ax.add_patch(Rectangle(xy=bottom_left, width=width, height=height, alpha=0.1))
# Print regression metrics
mse = mean_squared_error(y, y_pred)
rmse = np.sqrt(mse)
mae = mean_absolute_error(y, y_pred)
r2 = r2_score(y, y_pred)
print('Mean Squared Error:', mse)
print('Root Mean Squared Error:', rmse)
print('Mean Absolute Error:', mae)
print('R^2:', r2)
# + id="4NClSHxumvBN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 514} outputId="d606f7b5-8579-4312-ac1e-abbbaf458dd0"
regression_squared_errors(df, feature, target, m=3, b=46)
# + id="DpAGTsWqnM65" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 17} outputId="438c3c4d-4dbc-4c86-f048-431677542b79"
from ipywidgets import interact, fixed
interact(
regression_squared_errors,
df=fixed(df),
feature=fixed(feature),
target=fixed(target),
m=(-5,-5,0.5),
b=(40,60,0.5)
);
# + [markdown] id="VQUPU_3Uh6vc" colab_type="text"
# ## Follow Along
# + [markdown] id="Kv9ihhAah86r" colab_type="text"
# What's the equation for the hyperplane?
#
# Can you relate the intercept and coefficients to what you see in the plot?
# + id="QIRzYL3diAsL" colab_type="code" colab={}
# + [markdown] id="sDofh8lRiBnl" colab_type="text"
# What if ...
#
# Income growth = 0%, fatalities = 0
# + id="A9pT_eoiiFKY" colab_type="code" colab={}
# + [markdown] id="kqh8SabriGPQ" colab_type="text"
# Income growth = 1% (fatalities = 0)
# + id="mit_WTuxiJY2" colab_type="code" colab={}
# + [markdown] id="Ch6GPjbZiKN2" colab_type="text"
# The difference between these predictions = ?
# + id="wOzJKxBoiObb" colab_type="code" colab={}
# + [markdown] id="F2XMTNJ2nwoG" colab_type="text"
# ## Linear Algebra
#
# The same result that is found by minimizing the sum of the squared errors can be also found through a linear algebra process known as the "Least Squares Solution:"
#
# \begin{align}
# \hat{\beta} = (X^{T}X)^{-1}X^{T}y
# \end{align}
#
# Before we can work with this equation in its linear algebra form we have to understand how to set up the matrices that are involved in this equation.
#
# ### The $\beta$ vector
#
# The $\beta$ vector represents all the parameters that we are trying to estimate, our $y$ vector and $X$ matrix values are full of data from our dataset. The $\beta$ vector holds the variables that we are solving for: $\beta_0$ and $\beta_1$
#
# Now that we have all of the necessary parts we can set them up in the following equation:
#
# \begin{align}
# y = X \beta + \epsilon
# \end{align}
#
# Since our $\epsilon$ value represents **random** error we can assume that it will equal zero on average.
#
# \begin{align}
# y = X \beta
# \end{align}
#
# The objective now is to isolate the $\beta$ matrix. We can do this by pre-multiplying both sides by "X transpose" $X^{T}$.
#
# \begin{align}
# X^{T}y = X^{T}X \beta
# \end{align}
#
# Since anything times its transpose will result in a square matrix, if that matrix is then an invertible matrix, then we should be able to multiply both sides by its inverse to remove it from the right hand side. (We'll talk tomorrow about situations that could lead to $X^{T}X$ not being invertible.)
#
# \begin{align}
# (X^{T}X)^{-1}X^{T}y = (X^{T}X)^{-1}X^{T}X \beta
# \end{align}
#
# Since any matrix multiplied by its inverse results in the identity matrix, and anything multiplied by the identity matrix is itself, we are left with only $\beta$ on the right hand side:
#
# \begin{align}
# (X^{T}X)^{-1}X^{T}y = \hat{\beta}
# \end{align}
#
# We will now call it "beta hat" $\hat{\beta}$ because it now represents our estimated values for $\beta_0$ and $\beta_1$
#
# ### Lets calculate our $\beta$ coefficients with numpy!
# + id="OpJ9lQQpnwoH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 646} outputId="5bd92fe7-a608-474d-a93d-7c1f3738d29e"
# This is NOT an objective you'll be tested on. It's just a demo.
# X is a matrix. Add constant for the intercept.
from statsmodels.api import add_constant
X = add_constant(df['Average Recent Growth in Personal Incomes'].values)
print('X')
print(X)
# y is a column vector
y = df['Incumbent Party Vote Share'].values[:, np.newaxis]
print('y')
print(y)
# Least squares solution in code
X_transpose = X.T
X_transpose_X = X_transpose @ X
X_transpose_X_inverse = np.linalg.inv(X_transpose_X)
X_transpose_y = X_transpose @ y
beta_hat = X_transpose_X_inverse @ X_transpose_y
print('Beta Hat')
# + id="sqP8MLVmnwoL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 617} outputId="a8c80604-547d-4944-bf58-194aa3d1d08e"
px.scatter(
df,
x='Average Recent Growth in Personal Incomes',
y='Incumbent Party Vote Share',
text='Year',
title='US Presidential Elections, 1952-2016',
trendline='ols',
)
# + [markdown] id="6dtFnUepnwoO" colab_type="text"
# ## Overfitting Demo
# + id="gvjk6-K8nwoR" colab_type="code" colab={}
# Read New York City property sales data, from first 4 months of 2019.
# Dataset has 23040 rows, 21 columns.
df = pd.read_csv('../data/NYC_Citywide_Rolling_Calendar_Sales.csv')
assert df.shape == (23040, 21)
# Change column names. Replace spaces with underscores
df.columns = [col.replace(' ', '_') for col in df]
# Remove symbols from SALE_PRICE string, convert to integer
df['SALE_PRICE'] = (
df['SALE_PRICE']
.str.replace('$','')
.str.replace('-','')
.str.replace(',','')
.astype(int)
)
# Keep subset of rows:
# Tribeca neighborhood, Condos - Elevator Apartments,
# 1 unit, sale price more than $1, less than $35 million
mask = (
(df['NEIGHBORHOOD'].str.contains('TRIBECA')) &
(df['BUILDING_CLASS_CATEGORY'] == '13 CONDOS - ELEVATOR APARTMENTS') &
(df['TOTAL_UNITS'] == 1) &
(df['SALE_PRICE'] > 0) &
(df['SALE_PRICE'] < 35000000)
)
df = df[mask]
# Data now has 90 rows, 21 columns
assert df.shape == (90, 21)
# Convert SALE_DATE to datetime
df['SALE_DATE'] = pd.to_datetime(df['SALE_DATE'], infer_datetime_format=True)
from ipywidgets import interact
import pandas as pd
from sklearn.linear_model import LinearRegression
# Read New York City property sales data, from first 4 months of 2019.
# Dataset has 23040 rows, 21 columns.
df = pd.read_csv('../data/NYC_Citywide_Rolling_Calendar_Sales.csv')
assert df.shape == (23040, 21)
# Change column names. Replace spaces with underscores
df.columns = [col.replace(' ', '_') for col in df]
# Remove symbols from SALE_PRICE string, convert to integer
df['SALE_PRICE'] = (
df['SALE_PRICE']
.str.replace('$','')
.str.replace('-','')
.str.replace(',','')
.astype(int)
)
# Keep subset of rows:
# Tribeca neighborhood, Condos - Elevator Apartments,
# 1 unit, sale price more than $1, less than $35 million
mask = (
(df['NEIGHBORHOOD'].str.contains('TRIBECA')) &
(df['BUILDING_CLASS_CATEGORY'] == '13 CONDOS - ELEVATOR APARTMENTS') &
(df['TOTAL_UNITS'] == 1) &
(df['SALE_PRICE'] > 0) &
(df['SALE_PRICE'] < 35000000)
)
df = df[mask]
# Data now has 90 rows, 21 columns
assert df.shape == (90, 21)
# Convert SALE_DATE to datetime
df['SALE_DATE'] = pd.to_datetime(df['SALE_DATE'], infer_datetime_format=True)
# Arrange X features matrix & y target vector
features = ['GROSS_SQUARE_FEET']
target = 'SALE_PRICE'
X = df[features]
y = df[target]
# + [markdown] id="8Wdo1Uv5VSRQ" colab_type="text"
# Do random test/train - Polynomial
# + id="9yg81hvCnwoT" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="dc532a51-616e-48db-d861-b035afcfb9a0"
from IPython.display import display, HTML
from sklearn.model_selection import train_test_split
# Train/Test Split
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=11)
# Repeatedly fit increasingly complex models, and keep track of the scores
polynomial_degrees = range(1, 10, 2)
train_r2s = []
test_r2s = []
for degree in polynomial_degrees:
model = PolynomialRegression(degree)
display(HTML(f'Polynomial degree={degree}'))
model.fit(X_train, y_train)
train_r2 = model.score(X_train, y_train)
test_r2 = model.score(X_test, y_test)
display(HTML(f'<b style="color: blue">Train R2 {train_r2:.2f}</b>'))
display(HTML(f'<b style="color: red">Test R2 {test_r2:.2f}</b>'))
plt.scatter(X_train, y_train, color='blue', alpha=0.5)
plt.scatter(X_test, y_test, color='red', alpha=0.5)
plt.xlabel(features)
plt.ylabel(target)
x_domain = np.linspace(X.min(), X.max())
curve = model.predict(x_domain)
plt.plot(x_domain, curve, color='blue')
plt.show()
display(HTML('<hr/>'))
train_r2s.append(train_r2)
test_r2s.append(test_r2)
display(HTML('Validation Curve'))
plt.plot(polynomial_degrees, train_r2s, color='blue', label='Train')
plt.plot(polynomial_degrees, test_r2s, color='red', label='Test')
plt.xlabel('Model Complexity (Polynomial Degree)')
plt.ylabel('R^2 Score')
plt.legend()
plt.show()
# + [markdown] id="exL02cMYnwoV" colab_type="text"
# #### <NAME>, [_Python Data Science Handbook,_ Chapter 5.3](https://jakevdp.github.io/PythonDataScienceHandbook/05.03-hyperparameters-and-model-validation.html#The-Bias-variance-trade-off)
#
# 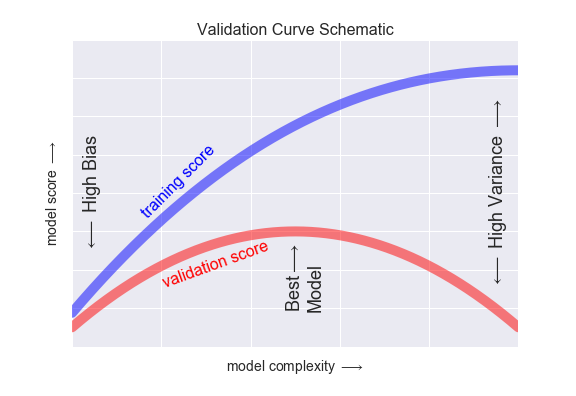
| module2/lesson_regression_classification_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/dkapitan/jads-nhs-proms/blob/master/notebooks/1.1-collect-initial-data.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="Ezut5pDim-JZ"
# # 1.1 - Collect initial data
#
# ## Background
# Read up on the documentation of the NHS PROMs data to prepare for this lesson:
# - [PROMs Guide](https://github.com/dkapitan/jads-nhs-proms-hko/blob/master/references/nhs/proms_guide_v12.pdf?raw=true)
# - [Data Dictionairy](https://github.com/dkapitan/jads-nhs-proms-hko/blob/master/references/nhs/proms_data_dictionary.pdf?raw=true?)
#
# ## Objectives
# - How to collect data?
# - How to handle missing values?
# - How to manage memory usage?
# - How to rename columns and variables?
#
# ## Python skills
# - Use `request` library to download files from the internet
# - Use `zipfile` to work with zipfiles
# - Know difference between getting a `bytes`, `string` and a `file` object for reading and writing data
# - How to do string manipulation for replacing columns names
# - How to use [`toolz.functoolz.compose`](https://toolz.readthedocs.io/en/latest/api.html#toolz.functoolz.compose) to compose functions to operate in series
# - How to use pandas:
# - for combining downloaded data into one dataframe: [`pd.concat`](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.concat.html)
# - do some memory optimization: [`df.select_dtypes`](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.select_dtypes.html), [`df.astype`](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.astype.html), [`pd.to_numeric`](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.to_numeric.html)
# - use [`df.rename`](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.rename.html) for renaming
# + colab={} colab_type="code" id="SNElYlTTm-Jc"
from io import BytesIO
import requests
import re
from zipfile import ZipFile
import pandas as pd
from toolz.functoolz import compose
CONDITION = 'Knee'
ORG = 'Provider'
def get_zipped_data_urls(condition, org):
"""
Get zipped datafiles for 3 years for condition and org
condition: choice of ['Knee', 'Hip']
org: choice of ['Provider', 'CCG']
"""
assert condition in ['Knee', 'Hip'], 'Condition is one of ["Knee", "Hip"]'
assert org in ['Provider', 'CCG'], 'Condition is one of ["Provider", "CCG"]'
return [
f'https://github.com/dkapitan/jads-nhs-proms-hko/blob/master/data/external/data-pack-2018-19/{condition}%20Replacement%20{org}%201819.csv.zip?raw=true',
f'https://github.com/dkapitan/jads-nhs-proms-hko/blob/master/data/external/data-pack-2017-18/{condition}%20Replacements%20{org}%201718.csv.zip?raw=true',
f'https://github.com/dkapitan/jads-nhs-proms-hko/blob/master/data/external/data-pack-2016-17/{condition}%20Replacement%20{org}%201617.csv.zip?raw=true',
]
def clean_python_name(s):
"""
https://gist.github.com/dkapitan/89ff20eeed38e6d9757fef9e09e23c3d
Method to convert string to clean string for use
in dataframe column names such :
i) it complies to python 2.x object name standard:
(letter|'_')(letter|digit|'_')
ii) my preference to use lowercase and adhere
to practice of case-insensitive column names for data
Based on
https://stackoverflow.com/questions/3303312/how-do-i-convert-a-string-to-a-valid-variable-name-in-python
"""
import re
# Remove leading characters until we find a letter or underscore, and remove trailing spaces
s = re.sub('^[^a-zA-Z_]+', '', s.strip())
# Replace invalid characters with underscores
s = re.sub('[^0-9a-zA-Z_]', '_', s)
return s.lower()
# + colab={} colab_type="code" id="w_qcyzmZm-Jf" outputId="0fdfab2d-2503-41f3-ba91-73fc654f729b"
# exmample: getting a single file
urls = get_zipped_data_urls(CONDITION, ORG)
get_ = requests.get(urls[0])
with ZipFile(BytesIO(get_.content)) as zipfile:
# print(file_.namelist())
df_ = pd.read_csv(zipfile.open(zipfile.namelist()[0]),
na_values=['*'])
df_.head()
# + colab={} colab_type="code" id="OjTHBmKTm-Jk"
# assignment: write a loop to get all files
dfs = []
for url in get_zipped_data_urls(CONDITION, ORG):
get_ = requests.get(url)
with ZipFile(BytesIO(get_.content)) as zipfile:
dfs.append(
pd.read_csv(zipfile.open(zipfile.namelist()[0]), na_values=['*']))
df = pd.concat(dfs)
# reset index since we have duplicate integer indexes from different years
df.reset_index(drop=True, inplace=True)
# + colab={} colab_type="code" id="NSPWLDvym-Jn" outputId="5e659061-abb4-4516-9ec8-4eda0efec0ec"
# check we have 3 years of data
df['Year'].unique()
# + colab={} colab_type="code" id="vh2k-WEDm-Jq" outputId="1685f06c-4a86-4300-99d2-35979ab01606"
# inspect memory usage: orignal is 85.3 MB
df.info(verbose=False)
# + colab={} colab_type="code" id="RxvLaxRHm-Js" outputId="9026d70b-b8f0-4802-963f-db2cadf16c43"
# reduce memory: strings to categories
# see: https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.select_dtypes.html
# see: https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.astype.html
for col in df.select_dtypes(include='object').columns:
df[col] = df[col].astype('category')
# int64 to smallest, unsigned int
for col in df.select_dtypes(include='int64').columns:
df[col] = pd.to_numeric(df[col], downcast='unsigned')
# float64 to smallest, unsigned int
for col in df.select_dtypes(include='float64').columns:
df[col] = pd.to_numeric(df[col], downcast='float')
# memory footprint now less than a fifth: 16 MB
df.info(verbose=False)
# + colab={} colab_type="code" id="AP4Q0pF9m-Jw" outputId="45c476d0-6744-4285-c62a-d563c68d792e"
# example: common string operations
def shorten_name(s):
return (s.replace('pre_op_q', 't0')
.replace('post_op_q', 't1')
.replace('knee_replacement', 'oks')
.replace('hip_replacement', 'ohs')
)
# Note: toolz.functoolz.compose applies functions from right to left
# use compose_left if you want left to right
df.rename(columns=compose(shorten_name, clean_python_name, shorten_name), inplace=True)
df.columns
# + colab={} colab_type="code" id="r2J5egEmm-Jz"
# explain how you can use parquet. File 6.3 MB
df.to_parquet(f'../data/interim/{CONDITION.lower()}-{ORG.lower()}.parquet', compression='gzip')
| notebooks/1.1-collect-initial-data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
## Extract info from OSM files
# -
import hashlib
import json
import pandas as pd
names = pd.read_csv('data/firstnames.csv', delimiter=';')
female_names = set(names[(names['gender'].str.contains('F')) |
(names['gender'] == '?') |
(names['gender'].str.contains('1'))]['name'])
male_names = set(names[(names['gender'].str.contains('M')) |
(names['gender'] == '?') |
(names['gender'].str.contains('1'))]['name'])
df = pd.read_csv('data/berlin-streets.csv', delimiter='\t', header=0, names=['oname', 'objectid', 'type', 'street'])
df.dropna(subset=['street'], inplace=True)
df['id'] = df.apply(lambda x: hashlib.sha1(x['street'].encode('utf-8')).hexdigest(), axis=1)
df.drop_duplicates(subset=['id'], inplace=True)
# +
df['extracted_name'] = df.apply(lambda x: ' '.join(x['street'].split('-')[0:-1]) or None, axis=1)
df['maybe_person'] = df.apply(lambda x: True if x['extracted_name'] else None, axis=1)
df['maybe_woman'] = df.apply(lambda x: (True if set(x['extracted_name'].split(' ')).intersection(female_names)
else None) if x['extracted_name'] else None, axis=1)
df['maybe_man'] = df.apply(lambda x: (True if set(x['extracted_name'].split(' ')).intersection(male_names)
else None) if x['extracted_name'] else None, axis=1)
df['is_person'] = None
df['is_woman'] = None
df['is_man'] = None
json_str = json.dumps(json.loads(df.to_json(orient='records')), indent=2)
with open('streets.json', 'w') as outfile:
outfile.write(json_str)
# +
## Extract street lists alphabetically
# -
import locale
# locale.getlocale()
# help(locale)
locale.setlocale(locale.LC_ALL, 'de_DE.UTF-8')
df.head()
# +
from IPython.display import display, Markdown
initial_chars = list(filter(lambda x: x.isalpha(),
sorted(set([n[0].upper() for n in df['street'].dropna()]),
key=locale.strxfrm)))
# [(letter, len(df[df['street'].str.startswith(letter)])) for letter in initial_chars]
def gen_markdown_table(c):
def border(row):
return '|' + row + '|\n'
def rpad(val, target_len):
return val + ' ' * (1 + target_len - len(str(val)))
fields = ['extracted_name',
'street',
'type',
'maybe_person',
'maybe_woman',
'maybe_man',
'is_person',
'is_woman',
'is_man']
length = {'street': 0,
'extracted_name': 0,
'type': 0,
'maybe_person': 12,
'maybe_woman': 11,
'maybe_man': 9,
'is_person': 9,
'is_woman': 8,
'is_man': 7
}
subset = df[df['street'].str.startswith(c)].sort_values(by='street', inplace=False)
for f in fields:
length[f] = max((length[f],
max(subset.apply(lambda x: len(str(x[f])), axis=1))
)
)
md = border('|'.join([' ' + rpad(f, length[f]) for f in fields]))
md += border('|'.join(['-' * (length[f] + 2) for f in fields]))
for idx, row in subset.iterrows():
md += border('|'.join([' ' + rpad(str(row[f]), length[f]) if row[f] else '' for f in fields]))
display(Markdown(md))
print(md)
gen_markdown_table('A')
# -
df[df['street'].str.startswith('7')]
streets.to_json('streets.json',orient='records')
# +
## From here we'll work on extracting street nodes
## This point and below is a WIP.
# -
import json
import numpy as np
from scipy.spatial import ConvexHull
import wikipedia
street_nodes = df[df['type'].isin(['primary', 'secondary', 'tertiary'
'residential', 'living_street', 'unclassified',
'trunk', 'motorway', 'pedestrian', 'cycleway'])][['id', 'type', 'node', 'lat', 'lon']].groupby(by=['id'], axis=0)
# json_array = street_nodes.get_group('a65f18281e38ff7a43eb605e2a06e86c6649a337')[['lat', 'lon']].to_json(orient='values')
points = street_nodes.get_group('a65f18281e38ff7a43eb605e2a06e86c6649a337')[['lat', 'lon']].values
# points = json.loads(json_array)
# hull = ConvexHull(points)
# a = [points[x] for x in hull.vertices]
# a.append(points[hull.vertices[0]])
# pd.Series(a).to_json(orient='values')
street_nodes.get_group('a65f18281e38ff7a43eb605e2a06e86c6649a337')
nodes = street_nodes.get_group('a65f18281e38ff7a43eb605e2a06e86c6649a337')[['lat', 'lon']].values
nodes
wikipedia.set_lang('de')
print(women.iloc[0])
wikipedia.page(wikipedia.search(women.iloc[0])[0]).title
results = women.apply(lambda x : wikipedia.search(x))
for idx, r in results[results.apply(lambda x : len(x)) < 10].iteritems():
print({women.loc[idx]: r})
women.loc[230352]
df2 = pd.read_csv('data/berlin-streets.csv', delimiter='\t', header=0, names=['node', 'id', 'lat', 'long', 'country', 'city', 'street'])
df2.head()
bystreet = df2.groupby(by=["street", "city", "country"], axis=0)
bystreet.groups.keys()
df[df['city'] != 'Berlin']
bystreet.get_group(('Pestalozzistraße', 'Berlin', 'DE')).head()
| explore-street-names.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Dimensionality Reduction with the Shogun Machine Learning Toolbox
# #### *By <NAME> ([lisitsyn](https://github.com/lisitsyn)) and <NAME> ([iglesias](https://github.com/iglesias)).*
# This notebook illustrates <a href="http://en.wikipedia.org/wiki/Unsupervised_learning">unsupervised learning</a> using the suite of dimensionality reduction algorithms available in Shogun. Shogun provides access to all these algorithms using [Tapkee](http://tapkee.lisitsyn.me/), a C++ library especialized in <a href="http://en.wikipedia.org/wiki/Dimensionality_reduction">dimensionality reduction</a>.
# ## Hands-on introduction to dimension reduction
# First of all, let us start right away by showing what the purpose of dimensionality reduction actually is. To this end, we will begin by creating a function that provides us with some data:
# +
import numpy
import os
SHOGUN_DATA_DIR=os.getenv('SHOGUN_DATA_DIR', '../../../data')
def generate_data(curve_type, num_points=1000):
if curve_type=='swissroll':
tt = numpy.array((3*numpy.pi/2)*(1+2*numpy.random.rand(num_points)))
height = numpy.array((numpy.random.rand(num_points)-0.5))
X = numpy.array([tt*numpy.cos(tt), 10*height, tt*numpy.sin(tt)])
return X,tt
if curve_type=='scurve':
tt = numpy.array((3*numpy.pi*(numpy.random.rand(num_points)-0.5)))
height = numpy.array((numpy.random.rand(num_points)-0.5))
X = numpy.array([numpy.sin(tt), 10*height, numpy.sign(tt)*(numpy.cos(tt)-1)])
return X,tt
if curve_type=='helix':
tt = numpy.linspace(1, num_points, num_points).T / num_points
tt = tt*2*numpy.pi
X = numpy.r_[[(2+numpy.cos(8*tt))*numpy.cos(tt)],
[(2+numpy.cos(8*tt))*numpy.sin(tt)],
[numpy.sin(8*tt)]]
return X,tt
# -
# The function above can be used to generate three-dimensional datasets with the shape of a [Swiss roll](http://en.wikipedia.org/wiki/Swiss_roll), the letter S, or an helix. These are three examples of datasets which have been extensively used to compare different dimension reduction algorithms. As an illustrative exercise of what dimensionality reduction can do, we will use a few of the algorithms available in Shogun to embed this data into a two-dimensional space. This is essentially the dimension reduction process as we reduce the number of features from 3 to 2. The question that arises is: what principle should we use to keep some important relations between datapoints? In fact, different algorithms imply different criteria to answer this question.
# Just to start, lets pick some algorithm and one of the data sets, for example lets see what embedding of the Swissroll is produced by the Isomap algorithm. The Isomap algorithm is basically a slightly modified Multidimensional Scaling (MDS) algorithm which finds embedding as a solution of the following optimization problem:
#
# $$
# \min_{x'_1, x'_2, \dots} \sum_i \sum_j \| d'(x'_i, x'_j) - d(x_i, x_j)\|^2,
# $$
#
# with defined $x_1, x_2, \dots \in X~~$ and unknown variables $x_1, x_2, \dots \in X'~~$ while $\text{dim}(X') < \text{dim}(X)~~~$,
# $d: X \times X \to \mathbb{R}~~$ and $d': X' \times X' \to \mathbb{R}~~$ are defined as arbitrary distance functions (for example Euclidean).
#
# Speaking less math, the MDS algorithm finds an embedding that preserves pairwise distances between points as much as it is possible. The Isomap algorithm changes quite small detail: the distance - instead of using local pairwise relationships it takes global factor into the account with shortest path on the neighborhood graph (so-called geodesic distance). The neighborhood graph is defined as graph with datapoints as nodes and weighted edges (with weight equal to the distance between points). The edge between point $x_i~$ and $x_j~$ exists if and only if $x_j~$ is in $k~$ nearest neighbors of $x_i$. Later we will see that that 'global factor' changes the game for the swissroll dataset.
#
# However, first we prepare a small function to plot any of the original data sets together with its embedding.
# +
# %matplotlib inline
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
# %matplotlib inline
def plot(data, embedded_data, colors='m'):
fig = plt.figure()
fig.set_facecolor('white')
ax = fig.add_subplot(121,projection='3d')
ax.scatter(data[0],data[1],data[2],c=colors,cmap=plt.cm.Spectral)
plt.axis('tight'); plt.axis('off')
ax = fig.add_subplot(122)
ax.scatter(embedded_data[0],embedded_data[1],c=colors,cmap=plt.cm.Spectral)
plt.axis('tight'); plt.axis('off')
plt.show()
# +
from shogun import Isomap, features, MultidimensionalScaling
# wrap data into Shogun features
data, colors = generate_data('swissroll')
feats = features(data)
# create instance of Isomap converter and configure it
isomap = Isomap()
isomap.put('target_dim', 2)
# set the number of neighbours used in kNN search
isomap.put('k', 20)
# create instance of Multidimensional Scaling converter and configure it
mds = MultidimensionalScaling()
mds.put('target_dim', 2)
# embed Swiss roll data
embedded_data_mds = mds.embed(feats).get_feature_matrix()
embedded_data_isomap = isomap.embed(feats).get_feature_matrix()
plot(data, embedded_data_mds, colors)
plot(data, embedded_data_isomap, colors)
# -
# As it can be seen from the figure above, Isomap has been able to "unroll" the data, reducing its dimension from three to two. At the same time, points with similar colours in the input space are close to points with similar colours in the output space. This is, a new representation of the data has been obtained; this new representation maintains the properties of the original data, while it reduces the amount of information required to represent it. Note that the fact the embedding of the Swiss roll looks good in two dimensions stems from the *intrinsic* dimension of the input data. Although the original data is in a three-dimensional space, its intrinsic dimension is lower, since the only degree of freedom are the polar angle and distance from the centre, or height.
# Finally, we use yet another method, Stochastic Proximity Embedding (SPE) to embed the helix:
# +
from shogun import StochasticProximityEmbedding
# wrap data into Shogun features
data, colors = generate_data('helix')
features = features(data)
# create MDS instance
converter = StochasticProximityEmbedding()
converter.put('target_dim', 2)
# embed helix data
embedded_features = converter.embed(features)
embedded_data = embedded_features.get_feature_matrix()
plot(data, embedded_data, colors)
# -
# ## References
# - <NAME>., <NAME>., <NAME>, <NAME>: An Efficient Dimension Reduction Library. ([Link to paper in JMLR](http://jmlr.org/papers/v14/lisitsyn13a.html#!).)
# - <NAME>., <NAME>. and <NAME>. A Global Geometric Framework for Nonlinear Dimensionality Reduction. ([Link to Isomap's website](http://isomap.stanford.edu/).)
| doc/ipython-notebooks/converter/Tapkee.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
# +
s = pd.Series([0,1,4,9,16, 25], name = 'squeares')
print(s.index)
print(s.values, s.index)
print(s[2:4])
# +
pop2014 = pd.Series([100,93.3,99.1],
index = ['Java', 'C', 'C++'])
print(pop2014) #index sort
#exit()
pop2015 = pd.Series({'Java': 100, 'C':99.3, 'C++':99.5})
print(pop2015)
# -
print(pop2014.index)
print(pop2014.iloc[0:2])#dataframe er actuial index from 0
print(pop2014.loc[:'C++']) # je index amra push korbo seta
#C++ er age prjnto
#ix loc er moto kaj kore kintu index na dile iloc er moto kaj kore
twoyears = pd.DataFrame({'2014': pop2014, '2015': pop2015})
print(twoyears)
twoyears['Average'] = 0.5*(twoyears['2014'] + twoyears['2015'])
print(twoyears)
test_data = pd.DataFrame(np.random.choice(['a','b', 'c', 'd'],(3,3)), index = [1,2,3], columns = ['AA', 'BB', 'CC', ])
print(test_data)
# # Pandas aggregation
open('tips.csv','r').readlines()[:10]
tips = pd.read_csv('tips.csv')
tips.head(10)
#print("comes")
#tips.tail(10)
#print(tips)
#tips.head(10)
tips.shape
#tips.mean
tips.dtypes
tips.describe()
#tips.describe
tips.groupby('gender').mean()
tips.groupby(['gender', 'smoker']).mean()
pd.pivot_table(tips,'total_bill','gender','smoker')
pd.pivot_table(tips,'total_bill',['gender','smoker'], ['day','time'])
# gender, smoker - row
# # data frame creation and visulatizion
import pandas as pd
from matplotlib import pyplot as plt
url = 'http://archive.ics.uci.edu/ml/machine-learning-databases/iris/iris.data'
df = pd.read_csv(url)
df.head()
df.columns = ['sepal_length','sepal_width','petal_length','petal_width','flower_type',]
df['flower_type'] = df['flower_type'].astype('category')
df.flower_type = df.flower_type.cat.rename_categories([0,1,2])
df.head()
df['flower_type'].describe()
df.hist()
plt.show()
pd.scatter_matrix(df, diagonal = 'kde')
plt.show()
# +
df.to_csv('iris_normalized>csv')
new_df
#write csv
#DEAL WITH MISSING DATA
#df.fillna(0)
#df.dropna()
# -
#feature extraction
from sklearn.feature_extraction.text import TfidfVectorizer
import numpy as np
def latlong_distance(loc, p):
return lambda x : x+p
loc_lat = [1.123, 2.34, 3.56]
loc_lon = [1.06, 2.11, 3.12]
#loc_latlon = loc_lat * loc_lon
loc1 = np.array(loc_lat)
loc2 = np.array(loc_lon)
loc_final = np.arange(2)
print(loc_final)
# ##
#
| lab 3/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
class Solution:
def kInversePairs(self, n: int, k: int) -> int:
mod = pow(10, 9) + 7
dp = [[0] * 1002 for _ in range(1002)]
dp[1][0] = 1
for i in range(2, n+1):
dp[i][0] = 1
for j in range(1, k+1):
dp[i][j] = (dp[i][j-1] + dp[i-1][j]) % mod;
if j >= i:
dp[i][j]=(dp[i][j] - dp[i-1][j-i] + mod) % mod;
return dp[n][k] % mod
class Solution:
def kInversePairs(self, n: int, k: int) -> int:
mod = pow(10, 9) + 7
dp = [[0] * (1 + k) for _ in range(n + 1)]
for i in range(0, n + 1):
dp[i][0] = 1
for i in range(1, n + 1):
for j in range(1, k + 1):
dp[i][j] = dp[i - 1][j] + dp[i][j - 1] - (dp[i - 1][j - i] if j - i >= 0 else 0)
return dp[-1][-1] % mod
solution = Solution()
solution.kInversePairs(3, 1)
# 1、dp[n][k] = dp[n - 1][k] + dp[n - 1][k-1] + ... + dp[n - 1][k - n + 1]
#
# 我们可以用 k+1 代替 k,得到:
#
# 2、dp[n][k+1] = dp[n - 1][k+1] + dp[n - 1][k] + ... + dp[n - 1][k + 1 - n + 1]
#
# 用第二个等式减去第一个等式可以得到:
#
# 3、dp[n][k+1] = dp[n][k] + dp[n - 1][k+1] - dp[n - 1][k - n + 1]
#
# 将 k+1 换回成 k,可以得到:
#
# 4、dp[n][k] = dp[n][k-1] + dp[n - 1][k] - dp[n - 1][k - n]
# +
from collections import Counter
class Solution:
def frequencySort(self, nums):
counter = Counter(nums)
counter = sorted(counter.items(), key=lambda x: (x[1], -x[0]))
res = []
for x, y in counter:
x = [x] * y
res.extend(x)
return res
# -
solution = Solution()
solution.frequencySort([2,3,1,3,2])
| Dynamic Programming/1202/629. K Inverse Pairs Array.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="3sbN5ESqh6QK"
# Initialize Google Earth Engine API
# + colab={"base_uri": "https://localhost:8080/"} id="Th6366lYh2Kp" outputId="13220b83-9551-411b-96b7-90246ce7639b"
import ee
# trigger the authentication flow
ee.Authenticate()
# initialize the library
ee.Initialize()
# + [markdown] id="na39Be12iAYq"
# Import some python modules and enable inline graphics
# + id="zvT-tCCViBwq"
import numpy as np
import matplotlib.pyplot as plt
from scipy.stats import norm, gamma, f, chi2
import IPython.display as disp
# %matplotlib inline
# + [markdown] id="aRIa85igmmoJ"
# Make use of interactive maps with the package Folium
# + id="M2R8eOuumpNi"
# import the Folium library.
import folium
# define a method for displaying Earth Engine image tiles to folium map.
def add_ee_layer(self, ee_image_object, vis_params, name):
map_id_dict = ee.Image(ee_image_object).getMapId(vis_params)
folium.raster_layers.TileLayer(
tiles = map_id_dict['tile_fetcher'].url_format,
attr = 'Map Data © <a href="https://earthengine.google.com/">Google Earth Engine</a>',
name = name,
overlay = True,
control = True
).add_to(self)
# add EE drawing method to folium.
folium.Map.add_ee_layer = add_ee_layer
# + [markdown] id="Mz89CT9wiVgS"
# Define the region of interest with GeoJSON
# + id="MsPyt-zgiIzd"
coords = [-8.49606, 41.49750, -8.36868, 41.59050]
geoJSON = {
"type": "Feature",
"properties": {},
"geometry": {
"type": "Polygon",
"coordinates": coords
}
}
aoi = ee.Geometry.Rectangle(coords)
# + [markdown] id="uyvde1ociZS_"
# Get the image collection from the Google Earth Engine archives
# + id="qcsoexsuib_p"
coll_fl = (ee.ImageCollection('COPERNICUS/S1_GRD_FLOAT') # specify image archive
.filterBounds(aoi) # specify area on interest
.filterDate(ee.Date('2014-01-01'),ee.Date('2021-01-01')) # specify time period
.filter(ee.Filter.eq('orbitProperties_pass', 'ASCENDING')) # specify orbit pass
#.filter(ee.Filter.eq('relativeOrbitNumber_start', 15)) # specify relative orbit number
.sort('system:time_start')) # sort by date
# log-scaled collection
coll_ls = (ee.ImageCollection('COPERNICUS/S1_GRD') # specify image archive
.filterBounds(aoi) # specify area on interest
.filterDate(ee.Date('2014-01-01'),ee.Date('2021-01-01')) # specify time period
.filter(ee.Filter.eq('orbitProperties_pass', 'ASCENDING')) # specify orbit pass
#.filter(ee.Filter.eq('relativeOrbitNumber_start', 15)) # specify relative orbit number
.sort('system:time_start')) # sort by date
# + [markdown] id="DA-elgzhilWL"
# Get collection as a list and clip all images to the area of interest
# + colab={"base_uri": "https://localhost:8080/"} id="80toKrniil7z" outputId="381ea2f1-0106-4e86-9478-d5135a1961b9"
# get the image collection as a list
list_fl = coll_fl.toList(coll_fl.size())
list_ls = coll_ls.toList(coll_ls.size())
print('Number of images available:', list_fl.length().getInfo())
# clip an image to the area of interest
def clip_img(img):
return ee.Image(img).clip(aoi)
# clip all images to the area of interest
list_fl = ee.List(list_fl.map(clip_img))
list_ls = ee.List(list_ls.map(clip_img))
# + [markdown] id="sG7gpVnMm7xq"
# Display first image
# + colab={"base_uri": "https://localhost:8080/", "height": 799} id="1kzCZn8zm8PY" outputId="cf3c0dca-85db-453c-cc5c-a36fcc94930f"
url = ee.Image(list_ls.get(0)).select('VV').getThumbURL({'min': -20, 'max': 0})
disp.Image(url=url, width=800)
# + [markdown] id="lUlMWCA7mzB9"
# Display first image as a RGB composite
# + id="ktli6hDsm0qP" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="4f4aa320-d52b-44be-eff0-c90ed20eefac"
def displayImage(img):
url = img.select('VV').getThumbURL({'min': -20, 'max': 0})
disp.Image(url=url, width=800)
location = aoi.centroid().coordinates().getInfo()[::-1]
# Make an RGB color composite image (VV,VH,VV/VH).
rgb = ee.Image.rgb(img.select('VV'),
img.select('VH'),
img.select('VV').divide(img.select('VH')))
# Create the map object.
m = folium.Map(location=location, zoom_start=12)
# Add the S1 rgb composite to the map object.
m.add_ee_layer(rgb, {'min': [-20, -20, 0], 'max': [0, 0, 2]}, 'FFA')
# Add a layer control panel to the map.
m.add_child(folium.LayerControl())
# Display the map.
display(m)
displayImage(ee.Image(list_ls.get(0)))
# + [markdown] id="UGL8R13Enwfq"
# Build the ratio of the VV bands of the first two images and display it
# + id="o1ECbdBbnyDW" colab={"base_uri": "https://localhost:8080/", "height": 799} outputId="f3fb5338-3ed7-441d-d8cc-cc23ed137061"
img1 = ee.Image(list_fl.get(0)).select('VV')
img2 = ee.Image(list_fl.get(1)).select('VV')
ratio = img1.divide(img2)
url = ratio.getThumbURL({'min': 0, 'max': 10})
disp.Image(url=url, width=800)
| Notebooks/sentinel1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="eU0knuKH7QKA" colab_type="code" colab={}
# !pip install keras==2.2.4
# + id="BdNqCnGF7t1B" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="dd09abb8-a8ef-4f59-e605-baecd4323c8b"
# %tensorflow_version 1.x
# + id="9z_0ayoS7t3U" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 72} outputId="9766974f-2d61-475e-96b0-2a4238f16fe4"
import keras
print(keras.__version__)
import tensorflow
print(tensorflow.__version__)
# + id="eFqYGDTx7t5p" colab_type="code" colab={}
import os
import numpy as np
import pandas as pd
import tensorflow as tf
import keras.backend as K
from keras import callbacks
from keras.utils import to_categorical
from keras.utils.vis_utils import plot_model
from keras.utils.data_utils import get_file
from keras import initializers, layers, models
import matplotlib.pyplot as plt
import warnings
warnings.filterwarnings('ignore')
# + id="yz-OKO7J7t9A" colab_type="code" colab={}
class Length(layers.Layer):
"""
Compute the length of vectors. This is used to compute a Tensor that has the same shape with y_true in margin_loss
inputs: shape=[dim_1, ..., dim_{n-1}, dim_n]
output: shape=[dim_1, ..., dim_{n-1}]
"""
def call(self, inputs, **kwargs):
return K.sqrt(K.sum(K.square(inputs), -1))
def compute_output_shape(self, input_shape):
return input_shape[:-1]
class Mask(layers.Layer):
"""
Mask a Tensor with shape=[None, d1, d2] by the max value in axis=1.
Output shape: [None, d2]
"""
def call(self, inputs, **kwargs):
# use true label to select target capsule, shape=[batch_size, num_capsule]
if type(inputs) is list: # true label is provided with shape = [batch_size, n_classes], i.e. one-hot code.
assert len(inputs) == 2
inputs, mask = inputs
else: # if no true label, mask by the max length of vectors of capsules
x = inputs
# Enlarge the range of values in x to make max(new_x)=1 and others < 0
x = (x - K.max(x, 1, True)) / K.epsilon() + 1
mask = K.clip(x, 0, 1) # the max value in x clipped to 1 and other to 0
# masked inputs, shape = [batch_size, dim_vector]
inputs_masked = K.batch_dot(inputs, mask, [1, 1])
return inputs_masked
def compute_output_shape(self, input_shape):
if type(input_shape[0]) is tuple: # true label provided
return tuple([None, input_shape[0][-1]])
else:
return tuple([None, input_shape[-1]])
def squash(vectors, axis=-1):
"""
The non-linear activation used in Capsule. It drives the length of a large vector to near 1 and small vector to 0
:param vectors: some vectors to be squashed, N-dim tensor
:param axis: the axis to squash
:return: a Tensor with same shape as input vectors
"""
s_squared_norm = K.sum(K.square(vectors), axis, keepdims=True)
scale = s_squared_norm / (0.5 + s_squared_norm) / K.sqrt(s_squared_norm)
return scale * vectors
class CapsuleLayer(layers.Layer):
"""
The capsule layer. It is similar to Dense layer. Dense layer has `in_num` inputs, each is a scalar, the output of the
neuron from the former layer, and it has `out_num` output neurons. CapsuleLayer just expand the output of the neuron
from scalar to vector. So its input shape = [None, input_num_capsule, input_dim_vector] and output shape = \
[None, num_capsule, dim_vector]. For Dense Layer, input_dim_vector = dim_vector = 1.
:param num_capsule: number of capsules in this layer
:param dim_vector: dimension of the output vectors of the capsules in this layer
:param num_routings: number of iterations for the routing algorithm
"""
def __init__(self, num_capsule, dim_vector, num_routing=3,
kernel_initializer='glorot_uniform',
bias_initializer='zeros',
**kwargs):
super(CapsuleLayer, self).__init__(**kwargs)
self.num_capsule = num_capsule
self.dim_vector = dim_vector
self.num_routing = num_routing
self.kernel_initializer = initializers.get(kernel_initializer)
self.bias_initializer = initializers.get(bias_initializer)
def build(self, input_shape):
assert len(input_shape) >= 3, "The input Tensor should have shape=[None, input_num_capsule, input_dim_vector]"
self.input_num_capsule = input_shape[1]
self.input_dim_vector = input_shape[2]
# Transformation matrix/Weight matrix
self.W = self.add_weight(shape=[self.input_num_capsule, self.num_capsule, self.input_dim_vector, self.dim_vector],
initializer=self.kernel_initializer,
name='W')
# Coupling coefficient. The redundant dimensions are just to facilitate subsequent matrix calculation.
self.bias = self.add_weight(shape=[1, self.input_num_capsule, self.num_capsule, 1, 1],
initializer=self.bias_initializer,
name='bias',
trainable=False)
self.built = True
def call(self, inputs, training=None):
# inputs.shape=[None, input_num_capsule, input_dim_vector]
# Expand dims to [None, input_num_capsule, 1, 1, input_dim_vector]
inputs_expand = K.expand_dims(K.expand_dims(inputs, 2), 2)
# Replicate(tile) num_capsule dimension to prepare being multiplied by W
# Now it has shape = [None, input_num_capsule, num_capsule, 1, input_dim_vector]
inputs_tiled = K.tile(inputs_expand, [1, 1, self.num_capsule, 1, 1])
"""
# Compute `inputs * W` by expanding the first dim of W. More time-consuming and need batch_size.
# Now W has shape = [batch_size, input_num_capsule, num_capsule, input_dim_vector, dim_vector]
w_tiled = K.tile(K.expand_dims(self.W, 0), [self.batch_size, 1, 1, 1, 1])
# Transformed vectors, inputs_hat.shape = [None, input_num_capsule, num_capsule, 1, dim_vector]
inputs_hat = K.batch_dot(inputs_tiled, w_tiled, [4, 3])
"""
# inputs_hat.shape = [None, input_num_capsule, num_capsule, 1, dim_vector]
inputs_hat = tf.scan(lambda ac, x: K.batch_dot(x, self.W, [3, 2]),
elems=inputs_tiled,
initializer=K.zeros([self.input_num_capsule, self.num_capsule, 1, self.dim_vector]))
# DYNAMIC ROUTING
assert self.num_routing > 0, 'The num_routing should be > 0.'
for i in range(self.num_routing):
c = tf.nn.softmax(self.bias, dim=2) # dim=2 is the num_capsule dimension
outputs = squash(K.sum(c * inputs_hat, 1, keepdims=True))
# last iteration needs not compute bias which will not be passed to the graph any more anyway.
if i != self.num_routing - 1:
# update the raw weights for the next routing iteration
# by adding the agreement to the previous raw weights
self.bias += K.sum(inputs_hat * outputs, -1, keepdims=True)
return K.reshape(outputs, [-1, self.num_capsule, self.dim_vector])
def compute_output_shape(self, input_shape):
return tuple([None, self.num_capsule, self.dim_vector])
def PrimaryCap(inputs, dim_vector, n_channels, kernel_size, strides, padding):
"""
Apply Conv2D `n_channels` times and concatenate all capsules
:param inputs: 4D tensor, shape=[None, width, height, channels]
:param dim_vector: the dim of the output vector of capsule
:param n_channels: the number of types of capsules
:return: output tensor, shape=[None, num_capsule, dim_vector]
"""
output = layers.Conv2D(filters=dim_vector*n_channels, kernel_size=kernel_size, strides=strides, padding=padding)(inputs)
outputs = layers.Reshape(target_shape=[-1, dim_vector])(output)
return layers.Lambda(squash)(outputs)
# + id="bd1zazIf7t_a" colab_type="code" colab={}
def CapsNet(input_shape, n_class, num_routing, digit_caps_dim_vector=16):
"""
A Capsule Network on MNIST.
:param input_shape: data shape, 4d, [None, width, height, channels]
:param n_class: number of classes
:param num_routing: number of routing iterations
:return: A Keras Model with 2 inputs and 2 outputs
"""
x = layers.Input(shape=input_shape)
# Layer 1: Conventional Conv2D layer
conv1 = layers.Conv2D(filters=256, kernel_size=9, strides=1, padding='valid', activation='relu', name='conv1')(x)
conv1 = layers.BatchNormalization(name='batch_norm1')(conv1)
# Layer 2: Conv2D layer with `squash` activation, then reshape to [None, num_capsule, dim_vector]
primarycaps = PrimaryCap(conv1, dim_vector=8, n_channels=32, kernel_size=9, strides=2, padding='valid')
# Layer 3: Capsule layer. Dynamic Routing algorithm works here.
digitcaps = CapsuleLayer(num_capsule=n_class, dim_vector=digit_caps_dim_vector, num_routing=num_routing, name='digit_caps')(primarycaps)
# Layer 4: This is an auxiliary layer to replace each capsule with its length. Just to match the true label's shape.
out_caps = Length(name='out_caps')(digitcaps)
y = layers.Input(shape=(n_class,))
masked = Mask()([digitcaps, y]) # The true label is used to mask the output of capsule layer.
"""
Novelty:
As opposed to using the conventional dense network as a reconstruction/decoder network,
this model architecture leverages "Deconvolution"(which actually is the Transposed Convolution)
layer for reconstructing the input image from the learned features.
Using such a decoder architecture makes intuitive sense over a dense network as similar decoder
architectures have been deployed in other computer vision tasks such as Image/Instance segmentation.
NOTE: the deconvolution name is a misnomer and the layer should more appropriately be called the
transposed convolution layer, something which has been fixed in the later versions of keras and
tensorflow.
"""
# Decoder Network
decoder = models.Sequential(name='decoder')
decoder.add(layers.Dense(input_dim=digit_caps_dim_vector, activation="relu", output_dim=7*7*digit_caps_dim_vector, name='dense1'))
decoder.add(layers.Reshape((7, 7, digit_caps_dim_vector)))
decoder.add(layers.BatchNormalization(name='batch_norm2'))
decoder.add(layers.Deconvolution2D(64, 3, 3, subsample=(1, 1), border_mode='same', name='deconv1'))
decoder.add(layers.Deconvolution2D(32, 3, 3, subsample=(2, 2), border_mode='same', name='deconv2'))
decoder.add(layers.Deconvolution2D(16, 3, 3, subsample=(2, 2), border_mode='same', name='deconv3'))
decoder.add(layers.Deconvolution2D(1, 3, 3, subsample=(1, 1), border_mode='same', name='deconv4'))
decoder.add(layers.Activation("relu"))
decoder.add(layers.Reshape(target_shape=input_shape, name='out_recon'))
# plot the decoder/reconstruction network
try:
plot_model(decoder, to_file='decoder.png', show_shapes=True)
except Exception as e:
print('No fancy plot {}'.format(e))
# create model used for training/inference/prediction
# two-input-two-output keras Model
train_model = models.Model([x, y], [out_caps, decoder(masked)])
# create model to manipulate fed digit to visualise the effect of a caps dimension on the reconstruction
# three-input-one-output keras Model; input: (image, mask, noise)
noise = layers.Input(shape=(n_class, digit_caps_dim_vector))
noise_plus_digitcaps = layers.Add()([digitcaps, noise])
masked_noised_y = Mask()([noise_plus_digitcaps, y]) # masked the noised digitcaps with input
manip_model = models.Model([x, y, noise], decoder(masked_noised_y))
return train_model, manip_model
# + id="vUmhfMRo7uET" colab_type="code" colab={}
def margin_loss(y_true, y_pred):
"""
Margin loss for Eq.(4). When y_true[i, :] contains not just one `1`, this loss should work too. Not test it.
:param y_true: [None, n_classes]
:param y_pred: [None, num_capsule]
:return: a scalar loss value.
"""
L = y_true * K.square(K.maximum(0., 0.9 - y_pred)) + \
0.5 * (1 - y_true) * K.square(K.maximum(0., y_pred - 0.1))
return K.mean(K.sum(L, 1))
# + id="yLb8-11d7uIh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 53} outputId="71b378cf-2909-4799-c404-a0260472c3c9"
(train_images, train_labels), (test_images, test_labels) = tf.keras.datasets.mnist.load_data()
# + id="R-USwZ3g7uLb" colab_type="code" colab={}
[train_images, test_images] = [x.reshape((-1, 28, 28, 1)).astype('float32')/255. for x in [train_images, test_images]]
[train_labels, test_labels] = [to_categorical(x.astype('float32')) for x in [train_labels, test_labels]]
# + id="5hi3jPAZ7uNc" colab_type="code" colab={}
val_images = train_images[54000:]
val_labels = train_labels[54000:]
train_images = train_images[:54000]
train_labels = train_labels[:54000]
# + id="VJrisF187uQG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 72} outputId="52103119-35a9-4b71-cd84-18de275f65ff"
print("Training images: ", train_images.shape)
# print(type(train_images))
print("Validation images: ", val_images.shape)
# print(type(val_images))
print("Testing images: ", test_images.shape)
# print(type(test_images))
# Aliasing for conventions
x_train = train_images
y_train = train_labels
x_val = val_images
y_val = val_labels
x_test = test_images
y_test = test_labels
# + id="F_AnP7DI7uUM" colab_type="code" colab={}
# custom util function for visualisation of the input images
def input_viz(data, title, num_row=5, num_col=10, facecolor='xkcd:ecru'):
x, y = data
num = num_row*num_col
images = x[:num]
labels = y[:num]
fig, axes = plt.subplots(num_row, num_col, figsize=(1.5*num_col,2*num_row), facecolor=facecolor)
fig.suptitle(title, fontsize=18, fontweight='bold')
for i in range(num):
ax = axes[i//num_col, i%num_col]
ax.imshow(images[i].reshape((28, 28)), cmap='gray')
ax.set_title(f'Number: {np.argmax(labels[i])}', fontsize=12)
ax.axis('Off')
plt.tight_layout(pad=2.0)
plt.show()
# + id="gRxzLK_D7uYY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 697} outputId="9cedfc6c-bb82-4d23-9645-8dcbf1837730"
input_viz((x_train, y_train), 'Train set')
# + id="TDAWw4Pu7uc2" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 697} outputId="88c3555c-26ec-44f5-9a5b-b95f468ca79b"
input_viz((x_val, y_val), 'Validation set')
# + id="hIdjrLY27uf5" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 697} outputId="e08af168-51bf-487e-da84-95f70f9dc909"
input_viz((x_test, y_test), 'Test set')
# + id="trr1gsns7ubM" colab_type="code" colab={}
class args:
routings = 3 # number of iterations of the dynamic routing algorithm between 2 capsule layers
digit_caps_dim_vector = 32 # number of capsules in the "digit-caps" layer OR the dimensionality of the output vectors of the digit caps layer
batch_size = 128
epochs = 75
recon_loss_weight = 0.0005
lr = 0.001
augmentation = True
# + id="4TfRQEMS7uWy" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 562} outputId="3339b06f-9691-47ba-a62e-475374da95bd"
input_shape = x_train.shape[1:] # for kannada mnist, mnist: (28, 28, 1)
num_classes = len(np.unique(np.argmax(y_train, 1))) # for kannada mnist, mnist: 10
# define model
model, manip_model = CapsNet(input_shape=input_shape,
n_class=num_classes,
num_routing=args.routings,
digit_caps_dim_vector=args.digit_caps_dim_vector)
model.summary()
try:
plot_model(model, to_file='model.png', show_shapes=True)
except Exception as e:
print('No fancy plot {}'.format(e))
# + id="yOSx1Lk_7uR-" colab_type="code" colab={}
"""
the original research paper proposes using a SSE loss with a weight of 0.0005
that is equivalent to using the mean loss (MSE) and then multiply it by the total
values being output in the final layer
(sum = number of instances * mean)
"""
recon_lambda = args.recon_loss_weight*np.prod(input_shape)
# + id="i2_xJ-3I7uG9" colab_type="code" colab={}
from keras.preprocessing.image import ImageDataGenerator
def train(model, data):
"""
Training a CapsuleNet
:param model: the CapsuleNet model
:param data: a tuple containing training and testing data, like `((x_train, y_train), (x_test, y_test))`
:param args: arguments
:return: The trained model
"""
# unpacking the data
(x_train, y_train), (x_val, y_val) = data
# callbacks
log = callbacks.CSVLogger('log.csv')
checkpoint = callbacks.ModelCheckpoint('weights-{epoch:02d}.h5', save_best_only=True, save_weights_only=True, verbose=1)
lr_decay = callbacks.LearningRateScheduler(schedule=lambda epoch: args.lr * np.exp(-epoch / 10.))
# compile the model
opt = 'nadam'
model.compile(optimizer=opt,
loss=[margin_loss, 'mse'], # use MSE loss metric for the reconstruction network
loss_weights=[1., recon_lambda],
metrics={'out_caps': 'accuracy'})
if(args.augmentation == False):
print("Training the model without applying image data augmentation")
# Training without data augmentation:
history = model.fit([x_train, y_train], [y_train, x_train], batch_size=args.batch_size, epochs=args.epochs,
validation_data=[[x_val, y_val], [y_val, x_val]], callbacks=[lr_decay, checkpoint, log])
else:
print("Training the model with image data augmentation")
# Begin: Training with data augmentation ---------------------------------------------------------------------#
def train_generator(x, y, batch_size, shift_fraction=0.2):
train_datagen = ImageDataGenerator(width_shift_range=shift_fraction,
height_shift_range=shift_fraction, # shift up to 2 pixel for MNIST
zca_whitening=True,
rotation_range=20,
zoom_range=0.2,
shear_range=0.2,
featurewise_center=True,
featurewise_std_normalization=True)
generator = train_datagen.flow(x, y, batch_size=args.batch_size)
while 1:
x_batch, y_batch = generator.next()
yield ([x_batch, y_batch], [y_batch, x_batch])
# Training with data augmentation. If shift_fraction=0., also no augmentation.
model.fit_generator(generator=train_generator(x_train, y_train, args.batch_size, 0.2),
steps_per_epoch=int(y_train.shape[0]/args.batch_size),
epochs=args.epochs,
validation_data=[[x_val, y_val], [y_val, x_val]], callbacks=[lr_decay, checkpoint, log])
# End: Training with data augmentation -----------------------------------------------------------------------#
model.save_weights('trained_model.h5')
print('Trained model saved to \'trained_model.h5\'')
return model
# + id="GGHBEi907uCz" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="4bf8ff98-3c8f-4db3-a4f4-d4e30ed2da2e"
train(model=model, data=((x_train, y_train), (x_val, y_val)))
# + id="P4Wr7yEzGUcA" colab_type="code" colab={}
model.load_weights('trained_model.h5')
# + id="DOK-JwSJGUYY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="5af7ee80-83c1-42dd-c7a2-500e21997fcb"
y_pred, x_recon = model.predict([x_test, y_test], batch_size=args.batch_size)
print('Test acc:', np.sum(np.argmax(y_pred, 1) == np.argmax(y_test, 1)) / y_test.shape[0])
# + [markdown] id="j-s_bNOtG2Tp" colab_type="text"
# **Load the best weights**
# + id="Qce-1o54Acyp" colab_type="code" colab={}
model.load_weights('weights.h5')
# + id="sFrSru91PHl-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="017d3d45-8de1-400c-c22a-df0e89984cd3"
y_pred, x_recon = model.predict([x_test, y_test], batch_size=args.batch_size)
print('Test acc:', np.sum(np.argmax(y_pred, 1) == np.argmax(y_test, 1)) / y_test.shape[0])
# + id="GFRB73dg8WaJ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 90} outputId="4fa9b490-3480-4b61-b568-7c932eac3adc"
pd.set_option('display.max_columns', None)
df = pd.read_csv('log.csv')
print(df.columns)
# + id="uQ1c1Y12ConK" colab_type="code" colab={}
# loss
loss = df['loss'].to_list()
val_loss = df['val_loss'].to_list()
# Break-down the losses into components
out_caps_loss = df['out_caps_loss'].to_list()
val_out_caps_loss = df['val_out_caps_loss'].to_list()
decoder_loss = df['decoder_loss'].to_list()
val_decoder_loss = df['val_decoder_loss'].to_list()
# accuracy
out_caps_acc = df['out_caps_acc'].to_list()
val_out_caps_acc = df['val_out_caps_acc'].to_list()
# + id="P_j1FicWCqxm" colab_type="code" colab={}
from pylab import rcParams
rcParams['figure.figsize'] = 7, 5
plt.style.use('fivethirtyeight')
plt.rc('grid', color='k', linestyle='--')
plt.rc('xtick', direction='out', color='black')
plt.rc('ytick', direction='out', color='black')
plt.rc('axes', facecolor='#E6E6E6', edgecolor='gray', axisbelow=True, grid=True)
# + id="niDgPftkCyAK" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 807} outputId="cf770401-6bb0-45cf-8a24-1677037065af"
epochs = range(len(loss))
params = {'left' : 1,
'right' : 3,
'bottom' : 1,
'top' : 3,
'wspace' : 1,
'hspace' : 0.2}
plt.subplots_adjust(**params)
# Plot retrieved data : accuracy
plt.subplot(221)
plt.plot(epochs, out_caps_acc)
plt.plot(epochs, val_out_caps_acc)
plt.title("Training and Validation Accuracy")
# Plot retrieved data : loss
plt.subplot(222)
plt.plot(epochs, loss)
plt.plot(epochs, val_loss)
plt.title("Training and Validation Loss")
# Plot retrieved data : capsule loss
plt.subplot(223)
plt.plot(epochs, out_caps_loss)
plt.plot(epochs, val_out_caps_loss)
plt.title("Training and Validation Capsule Loss")
# Plot retrieved data : reconstruction loss
plt.subplot(224)
plt.plot(epochs, decoder_loss)
plt.plot(epochs, val_decoder_loss)
plt.title("Training and Validation Reconstruction Loss")
# + id="5k8EQXwUC0NH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 449} outputId="a4a20e5a-1cc2-470b-8039-7f0dac99da32"
import random
index = random.randint(1, 10000)
print(f'For the sample number {index} in the test set')
f, (ax1, ax2) = plt.subplots(1, 2, sharey=True, figsize=(12, 12))
ax1.set_title(f'Actual number is {np.argmax(y_test[index])}', fontsize=12, fontweight='bold')
ax1.imshow(x_test[index].reshape(28, 28), cmap=plt.cm.binary)
ax2.set_title(f'Reconstructed number is {np.argmax(y_pred[index])}', fontsize=12, fontweight='bold')
ax2.imshow(x_recon[index].reshape(28, 28), cmap=plt.cm.binary)
# + id="2LAbbzJ_DEDH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 659} outputId="b9417897-b73e-401f-f610-11fe0025504d"
from sklearn.metrics import confusion_matrix
import seaborn as sn
labels = []
preds = []
for i in range(y_test.shape[0]):
labels.append(np.argmax(y_test[i]))
preds.append(np.argmax(y_pred[i]))
plt.figure(figsize=(10, 10))
plt.title("Test Set Confusion Matrix", fontsize=18, fontweight="bold")
cm = confusion_matrix(labels, preds)
sn.set(font_scale=1.4) # for label size
sn.heatmap(cm, annot=True, fmt='g', cmap="gist_earth", annot_kws={"size": 12}) # flag_r, copper, gist_earth, icefire
plt.show()
# + id="-k0N5pLKDHFk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 217} outputId="be3ed780-4a35-4a3b-9cd8-2a1ce7093e2a"
print("For the MNIST test set:")
num = 0
for row in cm:
print(f'Classification accuracy for the number {num} is {round((max(row)/sum(row)*100), 2)}%')
num += 1
# + id="bUvRE-kLDRQk" colab_type="code" colab={}
| mnist/capsnet mnist.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import doctest
import unittest
# ## ABC Tree
class Tree:
"""Abstract base class representing a tree structure"""
#---------------------------- nested Position class ----------------------------
class Position:
"""An abstraction representing the location of a single element."""
def element(self):
"""Return the element stored at this Position."""
raise NotImplementedError("Must be implemented by subclass")
def __eq__(self, other):
"""Return True if other Position represents the same location."""
raise NotImplementedError("Must be implemented by subclass")
def __ne__(self, other):
"""Return True if other does not represent the same location."""
return not (self == other)
# --------- abstract methods that concrete subclass must support -------------
def root(self):
"""Return the position representing the tree's root(or None if empty)"""
raise NotImplementedError("Must be implemented by subclass")
def parent(self, p):
"""Return Position representing p's parent(or None if p is root)."""
raise NotImplementedError("Must be implemented by subclass")
def num_children(self, p):
"""Return the number of children that Position p has."""
raise NotImplementedError("Must be implemented by subclass")
def children(self, p):
"""Generate an iteration of Positions representing p's children."""
raise NotImplementedError("Must be implemented by subclass")
def __len__(self):
"""Return the total number of elements in the tree."""
raise NotImplementedError("Must be implemented by subclass")
# --------- concrete methods that concrete subclass must support -------------
def is_root(self, p):
"""Return True if Position p represents the root of the tree."""
return self.root() == p
def is_leaf(slef, p):
"""Return True if Position p does not have any children."""
return self.num_children() == 0
def is_empty(self):
"""Return True if the tree is empty."""
return len(self) == 0
def depth(self, p):
"""Return the number of levels separating Position p from the root."""
if self.is_root(p):
return 0
return self.depth(self.parent(p)) + 1
def _height1(self):
"""Return the height of the tree."""
return max(self.depth(p) for p in self.positions() if self.is_leaf(p))
def _height2(self):
"""Return the height of the subtree rooted at Position p."""
if self.is_leaf(p):
return 0
return 1 + max(self._height2(c) for c in self.children(p))
def height(self, p):
if p is None:
p = self.root()
return self._height2(p)
# ### Binary Trees
class BinaryTree(Tree):
"""Abstract base class represent a binary tree."""
# --------------------- additional abstract methods ---------------------
def left(self, p):
"""Return the position that represents the left child of p.
Return None if p has no left child. """
raise NotImplementedError("Must be implemented by subclass")
def right(self, p):
"""Return the position that represents the right child of p.
Return None if p has no right child. """
raise NotImplementedError("Must be implemented by subclass")
# ---------- concrete methods implemented in this class ----------
def sibling(self, p):
"""Return a Position representing ps sibling (or None if no sibling)."""
parent = self.parent(p)
if not parent: # p must be the root
return None # root has no sibling
else:
if p == slef.left(parent):
return self.right(parent)
else:
return self.left(parent)
def children(self, p):
"""Generate an iteration of Positions representing ps children."""
if self.left(p):
yield self.left(p)
if self.right(p):
yield self.right(p)
| Data_Structures_and_Algorithms_in_Python/Trees.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Quality Control CTD data with PySeabird
# ### Author: <NAME>
# This is a minimalist example on how to use the Python Seabird package to read and apply a quality control in a CTD output file. For more details, please check the [documentation](https://seabird.readthedocs.io/en/latest/).
#
# ### Requirements
#
# This notebook requires the packages seabird, supportdata, and cotede. You can install those using pip as following:
#
# ```shell
# pip install seabird[QC]
# ```
# +
# #%matplotlib inline
from seabird.cnv import fCNV
from seabird.qc import fProfileQC
# -
# Let's first download an example file with some CTD data
# !wget https://raw.githubusercontent.com/castelao/seabird/master/sampledata/CTD/dPIRX003.cnv
profile = fCNV('dPIRX003.cnv')
print("Header: %s" % profile.attributes.keys())
print("Data: %s" % profile.keys())
# Let's apply the quality control procedure recommended by GTSPP
profile = fProfileQC('dPIRX003.cnv', cfg='gtspp')
# The QC flags are groupped for each variable. On this example there are temperature, salinity and the respective secondary sensors.
profile.flags.keys()
# Let's check which tests were performed, hence which flags are available, on the the primary temperature sensor
profile.flags['TEMP'].keys()
# The flagging standard is described in [CoTeDe's manual](https://cotede.readthedocs.io/en/latest/) . The one used here is 0 for no QC performed, 1 for approved data, and 9 for missing data.
#
# Note that the overall flag is the combined result from all tested flags. In the example above it considers the other 7 flags and takes the highest value, therefore, if the overall is equal to 1 means that all possible tests approved that measurement, while a value of 4 means that at least one tests suggests its a bad measurement.
profile.flags['TEMP']['spike']
idx = profile.flags['TEMP']['overall'] <= 2
# +
from matplotlib import pyplot as plt
plt.figure(figsize=(12,8))
plt.plot(profile['TEMP'][idx], profile['PRES'][idx],'b')
plt.plot(profile['TEMP'][~idx], profile['PRES'][~idx],'ro')
plt.gca().invert_yaxis()
# plt.plot(profile['TEMP2'], profile['PRES'],'g')
plt.xlabel('Temperature')
plt.ylabel('Pressure')
plt.title(profile.attributes['filename'])
# -
# Other pre defined quality control procedures are available, please check [CoTeDe's manual](https://cotede.readthedocs.io/en/latest/) to learn the details of the tests and what is available. For instance, to apply the EuroGOOS recommendations change the cfg argument
profile = fProfileQC('dPIRX003.cnv', cfg='eurogoos')
profile.flags['TEMP'].keys()
# If not defined, the default configuration is a collection of tests resulted for our work on [IQuOD](http://www.iquod.org/), and is equivalent to define `cfg='cotede'`.
profile = fProfileQC('dPIRX003.cnv')
profile.flags['TEMP'].keys()
| docs/notebooks/QualityControl.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a id='start'></a>
# # Transforming
#
# During the data gathering phase, your goal was to record as much data about your observations as possible since you never know which features are going to end up being the golden ones that allow your machine learning algorithm to succeed. Due to this, there usually are a few redundant or even poor features in your dataset. <br>
# *To be effective, many machine learning algorithms need their input data to be discerning, discriminating and independent.* <br>
# In this notebook, we are going to discover methods to get your data behaving like that using transformers. This will help improve your own knowledge of your data, as well as improve your machine learning algorithm's performance.
#
# This notebook will present the [Principal Component Analysis - PCA](#section1)<a href='#section1'></a>.
#
# <a id='section1'></a>
# ## 1) Principal Component Analysis - PCA
# PCA and other dimensionality reduction methods, have three main uses: <br>
# 1) Reducing the dimensionality and thus complexity of your dataset; <br>
# 2) Pre-process your data in preparation for other supervised learning tasks, such as regression and classification; <br>
# 3) To make visualizing your data esaier.
#
#
# Principal Component Analysis (PCA), is a transformation that attempts to convert possibly correlated features into a set of linearly uncorrelated ones. <br>
# In many real-world datasets, we aren't aware of what specifically needs to be measured to address the issue driving our data collection. <br>
# So instead, we simply collect any feature we can measure, usually resulting in a higher dimensionality (*a lot of features of the dataset*) than what is truly needed. This is undesirable, but it's the only reliable way to ensure we capture the relationship modeled by our data.
# <br>
# <br>
# If we have reason to believe the question we want solved can be answered using a subset of our collected features, or if the features we've collected are actually many indirect observations of some inherent property we either cannot or do not know how to directly measure, then dimensionality reduction might work for us. <br>
# **PCA's approach to dimensionality reduction is to derive a set of degrees of freedom that can then be used to reproduce most of the variability of our data.** <br>
# By PCA, first find the center of our data, based off its numeric features. Next, it would search for the direction that has the most variance or widest spread of values. That direction is the principal component vector, so it is added to a list. By searching for more directions of maximal variance that are orthogonal to all previously computed vectors, more principal component can then be added to the list. <br>
# This set of vectors form a new feature space that can represent our samples with.<br>
# PCA ensures that each newly computed view (feature) is orthogonal or linearly independent to all previously computed ones, minimizing these overlaps. PCA also orders the features by importance, assuming that the more variance expressed in a feature, the more important said feature is.<br>
# With the newly computed features ordered by importance, dropping the least important features on the list intelligently reduces the number of dimensions needed to represent your dataset, with minimal loss of information. <br>
# The remain group of features are the *principal components*, and they are the best possible, linearly independent combination of features that we can use to describe your data. <br>
#
#
# Try to better understand PCA with an example:
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import matplotlib
import math
from sklearn import preprocessing
plt.style.use('ggplot')
# +
# Defining a function for scaling the features of a dataset
def scaleFeaturesDF(df):
# Feature scaling is a type of transformation that only changes the
# scale, but not number of features. Because of this, we can still
# use the original dataset's column names... so long as we keep in
# mind that the _units_ have been altered:
scaled = preprocessing.StandardScaler(with_mean = False).fit_transform(df)
scaled = pd.DataFrame(scaled, columns=df.columns)
print("New Variances:\n", scaled.var())
print("New Describe:\n", scaled.describe())
return scaled
# -
# SKLearn contains many methods for transforming our features by scaling them, a type of [pre-processing](http://scikit-learn.org/stable/modules/preprocessing.html#preprocessing-scaler)):
# - [`RobustScaler`](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.RobustScaler.html)
# - [`Normalizer`](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.Normalizer.html#sklearn.preprocessing.Normalizer)
# - [`MinMaxScaler`](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.MinMaxScaler.html)
# - [`MaxAbsScaler`](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.MaxAbsScaler.html)
# - [`StandardScaler`](http://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.StandardScaler.html#sklearn.preprocessing.StandardScaler)
# <br>
#
# However in order to be effective at PCA, there are a few requirements that must be met, and which will drive the selection of your scaler. **PCA requires our data is standardized**, in other words, *it's mean should equal 0, and it should have unit variance*.
#
# SKLearn's regular `Normalizer()` doesn't zero out the mean of your data, it only clamps it, so it could be inappropriate to use depending on your data. `MinMaxScaler` and `MaxAbsScaler` both fail to set a unit variance, so you won't be using them here either. `RobustScaler` can work, again depending on your data (watch for outliers!). So for this assignment, we're going to use the `StandardScaler`.
# +
# Defining a function for visualizations
def drawVectors(transformed_features, components_, columns, plt, scaled):
if not scaled:
return plt.axes() # No cheating ;-)
num_columns = len(columns)
# This funtion will project your *original* feature (columns)
# onto your principal component feature-space, so that you can
# visualize how "important" each one was in the
# multi-dimensional scaling
# Scale the principal components by the max value in
# the transformed set belonging to that component
xvector = components_[0] * max(transformed_features[:,0])
yvector = components_[1] * max(transformed_features[:,1])
## visualize projections
# Sort each column by it's length. These are your *original*
# columns, not the principal components.
important_features = { columns[i] : math.sqrt(xvector[i]**2 + yvector[i]**2) for i in range(num_columns) }
important_features = sorted(zip(important_features.values(), important_features.keys()), reverse=True)
print("Features by importance:\n", important_features)
ax = plt.axes()
for i in range(num_columns):
# Use an arrow to project each original feature as a
# labeled vector on your principal component axes
plt.arrow(0, 0, xvector[i], yvector[i], color='b', width=0.0005, head_width=0.02, alpha=0.75)
plt.text(xvector[i]*1.2, yvector[i]*1.2, list(columns)[i], color='b', alpha=0.75)
return ax
# -
scaleFeatures = True
# Load up the dataset for doing a PCA analysis. <br>
# We will use the dataset of [Chronic Kidney Disease](https://archive.ics.uci.edu/ml/datasets/Chronic_Kidney_Disease)
kidn_dis = pd.read_csv('kidney_disease.csv')
kidn_dis.head()
kidn_dis.describe()
# +
# Investigating the main characteristic of the dataset
print("Shape of the dataset:", kidn_dis.shape)
# -
kidn_dis.columns
kidn_dis.dtypes
kidn_dis.describe()
# Count na
kidn_dis.isnull().sum()
# +
# Drop the id column
kidn_dis = kidn_dis.drop(['id'], axis = 1)
# Drop any na
kidn_dis = kidn_dis.dropna(axis=0)
kidn_dis.head()
# -
# **Pay attention to the index rows!** After a dropna some rows has been deleted, it could be better to [reset_index](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.reset_index.html) the dataframe:
kidn_dis.reset_index(inplace = True)
kidn_dis.head()
kidn_dis.isnull().sum()
print("After dropping na, the rows of the dataset are:", kidn_dis.shape[0])
# The actual label feature will be removed prior to executing PCA, since it's unsupervised.<br>
# We're only labeling by color so we can see the effects of PCA:
# We define color red if the record is 'ckd' for the features classification, while green if it is 'notckd'
labels = ['red' if i=='ckd' else 'green' for i in kidn_dis.classification]
# For semplicity of calculation, we will remove maintain only numeric columns
kidn_dis_2 = kidn_dis.loc[:, ['age', 'bp', 'sg', 'al', 'su', 'bgr', 'bu', 'sc', 'sod', 'pot', 'hemo']]
kidn_dis_2.head()
kidn_dis_2.dtypes
# PCA Operates based on variance. The variable with the greatest variance will dominate.
kidn_dis_2.describe()
# Each Standard deviation is a lot of different from each other; so it is necessary scaling the datas:
df = kidn_dis_2
if scaleFeatures: df = scaleFeaturesDF(df)
# To get started, **import PCA from sklearn.decomposition** and then create a new instance of the model setting the *n_components* parameter to the number of dimensions you wish to keep. This value has to be less than or equal to the number of features in our original dataset, since each computed component is a linear combination of our original features. <br>
# The second parameter, *svd_solver*, dictates if a full singular value decomposition should be preformed on our data, or a randomized truncated one. If we decide to use randomized, be sure to seed the random_state variable whenever if we intend on producing replaceable results.
# Once we've fit the model against our dataframe, we can use it to transform our dataset's observatios (or any other observation that share its feature space) into the newly computed, principal component feature space with the **.transform()** method. <br>
# This transformation is bidirectional, so we can recover our original feature values using **.inverse_transform()** so long as we don't drop any components. If even one component was removed, then after performing the inverse transformation back to the regular feature space, there will be some signs of information loss proportional to which component was dropped.
# Run [PCA](https://scikit-learn.org/stable/modules/generated/sklearn.decomposition.PCA.html) on our dataset, reducing it to 2 principal components.
# +
from sklearn.decomposition import PCA
pca = PCA(n_components=2, svd_solver='full')
pca.fit(df)
PCA(copy=True, n_components=2, whiten=False)
T = pca.transform(df)
# -
# There are a few other interesting model attribute that SciKit-Learn exposes to us after we've trained our PCA model with the .fit() method: <br>
# - **components_**: These are our principal component vectors and are linear combinations of our original features. As such, they exist within the feature space of our original dataset.
# - **explained_variance_**: This is the calculated amount of variance which exists in the newly computed principal components.
# - **explained_variance_ratio_**: Normalized version of *explained_variance_* for when our interest is with probabilities.
pca.explained_variance_ratio_
# +
# Since we transformed via PCA, we no longer have column names; but we know we
# are in `principal-component` space, so we'll just define the coordinates accordingly:
ax = drawVectors(T, pca.components_, df.columns.values, plt, scaleFeatures)
T = pd.DataFrame(T)
T.columns = ['component1', 'component2']
T.plot.scatter(x='component1', y='component2', marker='o', c=labels, alpha=0.75, ax=ax)
plt.show()
# -
T.head()
# After doing a PCA, we'll lost the header of the dataframe, so it could be difficult understand which features of the dataset could describe hypotetical linear relationship between the datas. <br>
# To interpret each component, we must compute the correlations between the original data and each principal component. <br>
# These correlations are obtained using the correlation procedure. In the variable statement we include the two principal components, "component1 and component2", in addition to all eleven of the original variables.
#
# We will note that if we look at the principal components themselves, then there is zero correlation between the components.
# Concating PCA's dataframe and starting dataset
df_corr = pd.concat([T, df], axis = 1)
df_corr.head()
corr_matrix = df_corr.corr()
corr_matrix
plt.imshow(df_corr.corr(), cmap = plt.cm.Blues, interpolation='nearest')
plt.colorbar()
tick_marks = [i for i in range(len(df_corr.columns))]
plt.xticks(tick_marks, df_corr.columns, rotation='vertical')
plt.yticks(tick_marks, df_corr.columns)
# Interpretation of the principal components is based on finding which variables are most strongly correlated with each component, i.e., which of these numbers are large in magnitude, the farthest from zero in either direction. Which numbers we consider to be large or small is of course is a subjective decision. You need to determine at what level the correlation is of importance
# From the visualization it is possible notice that component 1 is more correlated with 'sc', 'bu', 'hemo' and 'al' feature, while component 2 is more correlated with 'bgr'.
# #### Weaknesses of PCA <br>
# 1) PCA is sensitive to the scaling of our features. PCA maximizes variability based off of variance, and then projects our original data on these directions of maximal variances.<br><br>
# 2) PCA is fast, but for very large datasets it might take a while to train. If you're willing to sacrifice a bit of accuracy for computational efficiency, SciKit-Learn allows us to solve PCA using a offers an approximate matrix solver called *RandomizedPCA*. <br> <br>
# 3) PCA will only, therefore, be able to capture the underlying linear shapes and variance within our data and cannot discern any complex, nonlinear intricacies. For such cases, we will have to make use different dimensionality reduction algorithms, such as [*Isomap*](https://scikit-learn.org/stable/modules/generated/sklearn.manifold.Isomap.html).
# **Some useful links:**
# - [Interpreting PCA](https://newonlinecourses.science.psu.edu/stat505/node/54/); <br>
# - [Another method for interpreting PCA](http://www.stat.cmu.edu/~cshalizi/350/lectures/10/lecture-10.pdf); <br>
# - [Interactive PCA Demo](http://setosa.io/ev/principal-component-analysis/); <br>
# - [A tutorial on Principal Components Analysis](http://www.cs.otago.ac.nz/cosc453/student_tutorials/principal_components.pdf); <br>
# <br>
# With this paragraph ends the notebook "Transforming", the next notebook will be "Modeling".
# <br><br>
# [Click here to return to the top of the page](#start)<a href='#start'></a>
# <br>
# <br>
# If you have any doubts, you can write to us on Teams!<br>
# See you soon!
| 4_Transforming/Transforming.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import json
import random
from pathlib import Path
import pandas as pd
# %load_ext autoreload
# %autoreload 2
Path.ls = lambda x: list(x.iterdir())
# from urllib.parse import quote_plus
# -
pd.read_html("https://www.hindigeetmala.net/geetmala/binaca_geetmala_1953.php")[10]
| 50_Years_Bollywood/01_Data_Extraction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/agemagician/ProtTrans/blob/master/Embedding/PyTorch/Advanced/ProtBert.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="3PubGNU0dZmr" colab_type="text"
# <h3> Extracting protein sequences' features using ProtBert pretrained-model <h3>
# + [markdown] id="SAV0QHxfdZmt" colab_type="text"
# <b>1. Load necessry libraries including huggingface transformers<b>
# + id="C34oYIVxdeab" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 102} outputId="c992d658-aa06-4359-98ab-e79f4a5bc6d7"
# !pip install -q transformers
# + id="3NJPeESYdZmw" colab_type="code" colab={}
import torch
from transformers import BertModel, BertTokenizer
import re
import os
import requests
from tqdm.auto import tqdm
# + [markdown] id="JftgWQNZeGv3" colab_type="text"
# <b>2. Load the vocabulary and ProtBert Model</b>
# + id="70TQTgjadZm-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 66, "referenced_widgets": ["<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "373e1f37e53144f38d36c33c11998427", "9710ec2ec1264f4f8f994fcf7a198cdb", "<KEY>"]} outputId="2084ce7a-9906-4db3-af36-ed536b1ef4bc"
tokenizer = BertTokenizer.from_pretrained("Rostlab/prot_bert", do_lower_case=False)
# + id="2GcmM3pGdZnE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 115, "referenced_widgets": ["ac320877b54b4a229f754236fd9f8b89", "<KEY>", "535f6fb89b924ec19170812992c33360", "<KEY>", "<KEY>", "b5b979c9e4f64e37a476a8675ec62e36", "<KEY>", "4a4e488ea35f457aa538f6178f2374f7", "<KEY>", "bc3e358be1c442db944f0409e57e39d5", "<KEY>", "9c131285edd949978ea36a787ccbd9dd", "c1f60ee8b8834519961bde539af7a9d1", "8926483848c44a7491a45575f9ec4a74", "700ac1af0d83495f96beefa9ae4393ab", "<KEY>"]} outputId="21f43ebd-fef7-4f39-a2bf-f9eafec6bff8"
model = BertModel.from_pretrained("Rostlab/prot_bert")
# + [markdown] id="eM-12RxodZnK" colab_type="text"
# <b>3. Load the model into the GPU if avilabile and switch to inference mode<b>
# + id="xxElo34RdZnL" colab_type="code" colab={}
device = torch.device('cuda:0' if torch.cuda.is_available() else 'cpu')
# + id="YyQf6mwQdZnP" colab_type="code" colab={}
model = model.to(device)
model = model.eval()
# + [markdown] id="ZkqAotTcdZnW" colab_type="text"
# <b>4. Create or load sequences and map rarely occured amino acids (U,Z,O,B) to (X)<b>
# + id="a0zwKinIdZnX" colab_type="code" colab={}
sequences_Example = ["A E T C Z A O","S K T Z P"]
# + id="EkINwL9DdZna" colab_type="code" colab={}
sequences_Example = [re.sub(r"[UZOB]", "X", sequence) for sequence in sequences_Example]
# + [markdown] id="66BZEB3MdZnf" colab_type="text"
# <b>5. Tokenize, encode sequences and load it into the GPU if possibile<b>
# + id="xt5uYuu7dZnf" colab_type="code" colab={}
ids = tokenizer.batch_encode_plus(sequences_Example, add_special_tokens=True, pad_to_max_length=True)
# + id="Grl3ieUhdZnj" colab_type="code" colab={}
input_ids = torch.tensor(ids['input_ids']).to(device)
attention_mask = torch.tensor(ids['attention_mask']).to(device)
# + [markdown] id="Zylf1HyBdZnl" colab_type="text"
# <b>6. Extracting sequences' features and load it into the CPU if needed<b>
# + id="i8CVGPRFdZnm" colab_type="code" colab={}
with torch.no_grad():
embedding = model(input_ids=input_ids,attention_mask=attention_mask)[0]
# + id="dvgfP2h3dZnq" colab_type="code" colab={}
embedding = embedding.cpu().numpy()
# + [markdown] id="R6oeRZ7xdZns" colab_type="text"
# <b>7. Remove padding ([PAD]) and special tokens ([CLS],[SEP]) that is added by Bert model<b>
# + id="1XXoVSPDdZns" colab_type="code" colab={}
features = []
for seq_num in range(len(embedding)):
seq_len = (attention_mask[seq_num] == 1).sum()
seq_emd = embedding[seq_num][1:seq_len-1]
features.append(seq_emd)
# + id="swOQ_7r9dZnw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 391} outputId="dfe62e60-673f-4f06-cf09-7abc0298c845"
print(features)
| Embedding/PyTorch/Advanced/ProtBert.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data Visualisation
# +
## Data Visualisation
### importing libraries
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# +
## columns to be open
columns = ['sentiment','id','date','query_string','user','text']
filePath_r = 'data/original_data.csv'
## dataframe
df = pd.read_csv(filePath_r,header=None, names=columns, encoding = "cp1252")
print(df.head())
# -
## Printing the values counts
df.sentiment.value_counts()
## Drop the Meta data
df.drop(['id','date','query_string','user'],axis=1,inplace=True)
print(df.head())
## Print the rows with sentiment value is 0
df[df.sentiment == 0].head()
## print the rows where the sentiment value is 4
df[df.sentiment == 4].head()
| phase_1_visualisation/df_visualisation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19"
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python Docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
# Input data files are available in the read-only "../input/" directory
# For example, running this (by clicking run or pressing Shift+Enter) will list all files under the input directory
import os
for dirname, _, filenames in os.walk('/kaggle/input'):
for filename in filenames:
print(os.path.join(dirname, filename))
# You can write up to 20GB to the current directory (/kaggle/working/) that gets preserved as output when you create a version using "Save & Run All"
# You can also write temporary files to /kaggle/temp/, but they won't be saved outside of the current session
# -
from sklearn.linear_model import LogisticRegression
df = pd.read_csv('../data/train.csv')
test = pd.read_csv('../data/test.csv')
test
df.describe().T
df = df.drop(df.iloc[:, 1:5], axis=1)
test = test.drop(test.iloc[:, 1:5], axis=1)
target = df['revenue']
train = df.drop('revenue', axis=1)
train_dum = pd.get_dummies(train, drop_first=True)
test_dum = pd.get_dummies(test,drop_first=True)
test_dum = test_dum.loc[:,test_dum.columns.isin(train_dum.columns)]
train_dum = train_dum.loc[:,train_dum.columns.isin(test_dum.columns)]
log_model = LogisticRegression(solver='liblinear')
log_model.fit(train_dum, target)
# + pycharm={"name": "#%%\n"}
import pickle
pickle.dump(log_model, open('../models/logistic.pkt', 'wb'))
# -
predicted_test = log_model.predict(test_dum)
predicted_test
out = test['Id'].to_frame().copy()
out['Prediction'] = predicted_test
out.to_csv('my_submission.csv', index=False)
| notebooks/LogisticRegression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: test-pytorch-hooks
# language: python
# name: test-pytorch-hooks
# ---
# # Loading the model
# +
import torch
from torchvision.models import resnet34
device = torch.device('cuda') if torch.cuda.is_available() else torch.device('cpu')
model = resnet34(pretrained=True)
model = model.to(device)
# -
# # Defining the hook
class SaveOutput:
def __init__(self):
self.outputs = []
def __call__(self, module, module_in, module_out):
self.outputs.append(module_out)
def clear(self):
self.outputs = []
# # Registering the hook
# We register the hook to each convolutional layer.
# +
save_output = SaveOutput()
hook_handles = []
for layer in model.modules():
if isinstance(layer, torch.nn.modules.conv.Conv2d):
handle = layer.register_forward_hook(save_output)
hook_handles.append(handle)
# -
# # Forward pass
#
# The hook is not called yet, so `save_output.outputs` should be empty.
len(save_output.outputs)
# +
from PIL import Image
from torchvision import transforms as T
image = Image.open('cat.jpg')
transform = T.Compose([T.Resize((224, 224)), T.ToTensor()])
X = transform(image).unsqueeze(dim=0).to(device)
out = model(X)
# -
# The hook was called by each convolutional layer during the forward pass, so the layer outputs are saved.
len(save_output.outputs)
# We can visualize the intermediate results.
# +
import matplotlib.pyplot as plt
def module_output_to_numpy(tensor):
return tensor.detach().to('cpu').numpy()
images = module_output_to_numpy(save_output.outputs[0])
with plt.style.context("seaborn-white"):
plt.figure(figsize=(20, 20), frameon=False)
for idx in range(16):
plt.subplot(4, 4, idx+1)
plt.imshow(images[0, idx])
plt.setp(plt.gcf().get_axes(), xticks=[], yticks=[]);
| hooks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Pyteomics ms data reader
#
# The tool is adapted from [Link to the Github](https://github.com/pymzml/pymzML)
#
# The whole workflow should cite
#
# ```
# <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>; pymzML v2.0: introducing a highly compressed and seekable gzip format, Bioinformatics, doi: https://doi.org/10.1093/bioinformatics/bty046
# ```
#
# The Introduction is [Here](https://pyteomics.readthedocs.io/en/latest/data.html#mzml-and-mzxml)
# +
import pyteomics
from pyteomics import mzml, auxiliary
import matplotlib.pyplot as plt
import numpy as np
import math
import plotly.graph_objects as go
import re
from scipy.integrate import simps
import pandas as pd
from progressbar import ProgressBar
pbar = ProgressBar()
import peakutils
from peakutils.plot import plot as pplot
from matplotlib import pyplot
# -
# %matplotlib inline
#Read mzml files
f=mzml.MzML('../example_data/20191210_MSpos_tl_frac_B7_F5_narrow_2_noSPE.mzML')
# +
#The data structure is a mzml list contains scans as dictionary and titles
#Could iterate for the MS1/MS2 spectrums
#f[i] #where i is the index
#Get the scan time--Data structure is complex
f[0]['scanList']['scan'][0]['scan start time']
# -
#TIC plot
def tic_plot(spectrum, interactive=True):
'''
Static tic plot function
'''
time=[]
TIC=[]
for i in range(len(spectrum)):
time.append(spectrum[i]['scanList']['scan'][0]['scan start time'])
TIC.append(spectrum[i]['total ion current'])
if interactive == True:
fig = go.Figure([go.Scatter(x=time, y=TIC,
hovertemplate = 'Int: %{y}' + '<br>RT: %{x}minute<br>')])
fig.update_layout(
template = 'simple_white',
width = 1000,
height = 600,
xaxis = {'title':'Retention Time (min)'},
yaxis = dict(
showexponent = 'all',
exponentformat = 'e',
title = 'Intensity'))
fig.show()
elif interactive == False:
plt.figure(figsize=(10,6))
plt.plot(time,TIC)
plt.ticklabel_format(style='sci', axis='y', scilimits=(0,0))
plt.xlabel('RT (min)')
plt.ylabel('TIC')
plt.title('TIC spectrum')
plt.show()
return
tic_plot(f, False)
def ms_plot(spectrum, time, interactive=False):
'''
Interactive spectrum plot with nearest retention time from the given time
'''
for i in range(len(spectrum)):
if spectrum[i]['scanList']['scan'][0]['scan start time'] >= time:
mz = f[i]['m/z array']
ints = f[i]['intensity array']
rt = spectrum[i]['scanList']['scan'][0]['scan start time']
break
if interactive == True:
fig = go.Figure([go.Bar(x=mz, y=ints, marker_color = 'red', width = 0.5,
hovertemplate =
'Int: %{y}'+
'<br>m/z: %{x}<br>')])
fig.update_layout(
title_text=str(round(rt, 3)) + ' min MS1 spectrum, input '+ str(time) + ' min',
template = 'simple_white',
width = 1000,
height = 600,
xaxis = {'title':'m/z ratio'},
yaxis = dict(
showexponent = 'all',
exponentformat = 'e',
title = 'Intensity'))
fig.show()
elif interactive == False:
plt.figure(figsize=(10,5))
plt.bar(mz, ints, width = 1.0)
plt.ticklabel_format(style='sci', axis='y', scilimits=(0,0))
plt.xlabel('m/z')
plt.ylabel('Intensity')
plt.title('MS1 spectrum')
return
ms_plot(f, 15, True)
#Quick trail for try except
test = [1,2,3,'wow',5]
for i in test:
try:
i += 1
print(i)
except:
print('wrong type')
def mz_locator(input_list, mz, error):
'''
Find specific mzs from given mz and error range
input list: mz list
'''
target_mz = []
target_index = []
lower_mz = mz - error
higher_mz = mz + error
for i, mzs in enumerate(input_list):
if mzs < lower_mz:
continue
elif mzs >= lower_mz:
if mzs <= higher_mz:
target_mz.append(mzs)
target_index.append(i)
elif mzs > higher_mz:
target_mz = 0
target_index = 'NA'
break
return target_mz, target_index
def formula_mass(input_formula, mode = 'pos'):
'''
sudo code:
convert input string into a list with element:number structure
convert all the element into upper case
match the string list into a given list of element weight
add adduct/delete H according to mode -- also have neutral mode
'''
#Define a list
elist = {'C': 12,
'H':1.00782,
'N':14.0031,
'O':15.9949,
'S':31.9721,
'P':30.973763,
'e':0.0005485799}
mol_weight = 0
parsed_formula = re.findall(r'([A-Z][a-z]*)(\d*)', input_formula)
for element_count in parsed_formula:
element = element_count[0]
count = element_count[1]
if count == '':
count = 1
mol_weight += elist[element]*float(count)
if mode == 'pos':
mol_weight += elist['e'] + elist['H']
elif mode == 'neg':
mol_weight -= elist['e'] + elist['H']
else:
pass
return mol_weight
def ms_chromatogram(ms_file, input_mz, error, smooth=False, mode='pos', interactive=True):
'''
Interactive chromatogram for selected m/z
'''
if type(input_mz) == float:
pass
elif type(input_mz) == int:
pass
elif type(input_mz) == str:
input_mz = formula_mass(input_mz, mode)
else:
print('Cant recognize input type!')
retention_time = []
intensity = []
for i in range(len(ms_file)):
#print(i)
retention_time.append(ms_file[i]['scanList']['scan'][0]['scan start time'])
target_mz, target_index = mz_locator(ms_file[i]['m/z array'], input_mz, error)
if target_index == 'NA':
intensity.append(0)
else:
intensity.append(sum(ms_file[i]['intensity array'][target_index]))
def peak_smooth(input_list, baseline=500):
for i, int_ in enumerate(input_list):
if i > 1 and i < len(input_list)-3:
if int_ > baseline:
for index in np.arange(i+1,i+3):
if input_list[index] == 0:
input_list[index] = (input_list[index-1]+input_list[index+1])/2
else:
continue
if smooth == True:
peak_smooth(intensity)
if interactive == True:
fig = go.Figure([go.Scatter(x=retention_time, y=intensity,
hovertemplate = 'Int: %{y}' + '<br>RT: %{x}minute<br>')])
fig.update_layout(
title_text=str(round(input_mz, 2)) + ' chromatogram, error '+ str(error),
template = 'simple_white',
width = 1000,
height = 600,
xaxis = {'title':'Retention Time (min)'},
yaxis = dict(
showexponent = 'all',
exponentformat = 'e',
title = 'Intensity'))
fig.show()
elif interactive == False:
plt.figure(figsize=(20,10))
plt.plot(retention_time, intensity)
plt.ticklabel_format(style='sci', axis='y', scilimits=(0,0))
plt.xlabel('m/z')
plt.ylabel('Intensity')
plt.title('MS1 spectrum')
plt.xlim(0,retention_time[-1])
plt.ylim(0,)
plt.show()
return
ms_chromatogram(f, 'C18H22N2O2', 0.002, False, 'pos',False)
def ms_chromatogram_list(ms_file, input_mz, error, baseline = 5000):
'''
Generate a peak list for specific input_mz over whole rt period from the mzml file
***Most useful function!
'''
retention_time = []
intensity = []
for i in range(len(ms_file)):
#print(i)
retention_time.append(ms_file[i]['scanList']['scan'][0]['scan start time'])
target_mz, target_index = mz_locator(ms_file[i]['m/z array'], input_mz, error)
if target_index == 'NA':
intensity.append(0)
else:
intensity.append(sum(ms_file[i]['intensity array'][target_index]))
for i, ints in enumerate(intensity):
if ints < baseline:
intensity[i] = 0
return retention_time, intensity
def peak_pick(rt, intensity, peak_base = 5000, thr = 0.02, min_d = 1, rt_window = 1.5, peak_area_thres = 1e5, min_scan = 7, max_scan = 200):
'''
rt, ints from ms_chromatogram_list
rt_window now set up for minutes
'''
#Get rt_window corresponded scan number
scan_window = int((rt_window / (rt[int(len(intensity) / 2)] - rt[int(len(intensity) / 2) - 1])) / 2)
#Get peak index
indexes = peakutils.indexes(intensity, thres=thr, min_dist = min_d)
result_dict = {}
for index in indexes:
h_range = index
l_range = index
base_intensity = peak_base
#Get the higher and lower boundary
while intensity[h_range] >= base_intensity:
h_range += 1
if h_range > len(intensity)-2:
break
while intensity[l_range] >= base_intensity:
l_range -= 1
#Output a range from the peak list
peak_range = []
if h_range - l_range >= min_scan:
if rt[h_range] - rt[l_range] <= rt_window:
peak_range = intensity[l_range:h_range]
else:
l_range = index - scan_window
h_range = index + scan_window
peak_range = intensity[l_range:h_range]
#print(index + scan_window)
#Intergration based on the simps function
if len(peak_range) >= min_scan:
integration_result = simps(peak_range)
if integration_result >= peak_area_thres:
result_dict.update({index : [l_range, h_range, integration_result]})
return result_dict
def integration_plot(rt, ints, peak_base = 0.005, thr = 0.02, min_d = 1, rt_window = 2, peak_area_thres = 1e5):
result_dict = peak_pick(rt, ints)
plt.figure(figsize=(20,10))
plt.plot(rt, ints)
plt.ticklabel_format(style='sci', axis='y', scilimits=(0,0))
plt.xlabel('m/z')
plt.ylabel('Intensity')
plt.title('Integration result')
plt.xlim(0,rt[-1])
plt.ylim(0,)
for index in result_dict:
print('Peak retention time: {:0.2f} minute, Peak area: {:0.1f}'.format(rt[index], result_dict[index][2]))
plt.fill_between(rt[result_dict[index][0] : result_dict[index][1]], ints[result_dict[index][0] : result_dict[index][1]])
return
rt, ints = ms_chromatogram_list(f, 299.5, 0.5, 5000)
peak_pick(rt,ints)
integration_plot(rt, ints)
test_mz = []
for i in range(len(f_5000)):
test_mz.append(f_5000[i]['m/z array'])
from pandas.core.common import flatten
test_mz1 = list(flatten(test_mz))
len(test_mz1)
test_mz2 = list(set([test_mz1[i] for i in range(len(test_mz1))]))
len(test_mz2)
plt.figure(figsize=(20,15))
plt.hist(test_mz1)
min_mz = f_5000[0]['m/z array'].min()
max_mz = f_5000[0]['m/z array'].max()
for i in f_5000:
if min_mz > i['m/z array'].min():
min_mz = i['m/z array'].min()
if max_mz < i['m/z array'].max():
max_mz = i['m/z array'].max()
error = 0.002
i_list = []
i = min_mz + error
while i <= max_mz:
i_list.append(i)
i += 2 * error
df = pd.DataFrame(columns = ['1','2','3'])
f_5000[0]['rt']
f_5000[0]
def ms1_baseline(spectrum, baseline):
'''
Remove noises, then remove void features from the whole list
Note that this function now only works on MS1 data, no filter is applied on MS2 data, code needs to be updated in the future
'''
#Make a copy of spectrum
spec = spectrum
rt_dict = []
for i in range(len(spec)):
int_list = spec[i]['intensity array']
mz_list = spec[i]['m/z array']
idxs = [(index, ints) for index, ints in enumerate(int_list) if ints < baseline]
drop_idxs = [idx[0] for idx in idxs]
int_list = [i for j, i in enumerate(int_list) if j not in drop_idxs]
mz_list = [i for j, i in enumerate(mz_list) if j not in drop_idxs]
int_array = np.array(int_list)
mz_array = np.array(mz_list)
rt = spec[i]['scanList']['scan'][0]['scan start time']
current_dict = {'rt' : rt, 'm/z array' : mz_array, 'int array' : int_array}
rt_dict.append(current_dict)
#peaklist=list(zip(mz_list, int_list))
#rt_dict.update({spec[i]['scanList']['scan'][0]['scan start time'] : peaklist})
return rt_dict
f_5000 = ms1_baseline(f, 5000)
#Main dev issue
def peak_list(baseline_list, mz_error, peak_base = 0.005, thr = 0.02, min_d = 1, rt_window = 1.5, peak_area_thres = 1e5, min_scan = 7, scan_thres = 7):
'''
input from ms1_baseline function
Q to solve: how to correctly select mz slice?? see mz_locator
'''
#Get m/z range
def mz_list(spec):
min_mz = spec[0]['m/z array'].min()
max_mz = spec[0]['m/z array'].max()
for i in spec:
if min_mz > i['m/z array'].min():
min_mz = i['m/z array'].min()
if max_mz < i['m/z array'].max():
max_mz = i['m/z array'].max()
mz_list = []
mz = min_mz + mz_error
while mz <= max_mz:
mz_list.append(mz)
mz += 2 * mz_error
return mz_list
def ms_chromatogram_list(ms_file, input_mz, error, baseline = 5000):
retention_time = []
intensity = []
for scan in ms_file:
#print(i)
retention_time.append(scan['rt'])
target_mz, target_index = mz_locator(scan['m/z array'], input_mz, error)
if target_index == 'NA':
intensity.append(0)
else:
intensity.append(sum(scan['int array'][target_index]))
for i, ints in enumerate(intensity):
if ints < baseline:
intensity[i] = 0
return retention_time, intensity
mzlist = mz_list(baseline_list)
result_dict = {}
for mz in pbar(mzlist):
try:
rt, intensity = ms_chromatogram_list(baseline_list, mz, mz_error)
if len(intensity) >= scan_thres:
peak_dict = peak_pick(rt, intensity, peak_base = 0.005, thr = 0.02, min_d = 1, rt_window = 1.5, peak_area_thres = 1e5, min_scan = 7)
except:
pass
if len(peak_dict) != 0:
#Note: next to solve : how to deal with replicate items, i,e #3809
#Note2: how to deal with baseline noise i,e ms_chromatogram_list1(f_5000, 102.01360767939408, 0.002)
for index in peak_dict:
result_dict.update({'m/z' : mz,
'rt' : rt[index],
'peak area' : peak_dict[index][2]})
d_result = pd.DataFrame(result_dict)
return d_result
peak_list(f_5000, 0.002)
def ms_chromatogram_list1(ms_file, input_mz, error, baseline = 5000):
retention_time = []
intensity = []
for scan in ms_file:
#print(i)
retention_time.append(scan['rt'])
target_mz, target_index = mz_locator(scan['m/z array'], input_mz, error)
if target_index == 'NA':
intensity.append(0)
else:
intensity.append(sum(scan['int array'][target_index]))
for i, ints in enumerate(intensity):
if ints < baseline:
intensity[i] = 0
return retention_time, intensity
rt, ints = ms_chromatogram_list1(f_5000, 102.01360767939408, 0.002)
plt.plot(rt, ints)
# Formula input module
| dev/.ipynb_checkpoints/MSdata-reader-checkpoint.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .sh
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Bash
# language: bash
# name: bash
# ---
# ChainerMNで[MNIST](https://en.wikipedia.org/wiki/MNIST_database)(手書き数字認識)のサンプルを動かします。<br>
#
# まず、ChainerMNのGitHubからソースコードをダウンロードします。
curl -L https://github.com/chainer/chainermn/archive/v1.0.0b2.tar.gz | tar zx
# chainermn-1.0.0b2というディレクトリが作成されます。
ls
# MNISTのサンプルはchainermn-1.0.0b2/examples/mnist以下にあります。
ls chainermn-1.0.0b2/examples/mnist
cat chainermn-1.0.0b2/examples/mnist/train_mnist.py
# Chainerを用いたプログラムの作成方法の詳細については下記を参照してください。<br>
# http://docs.chainer.org/en/latest/tutorial/index.html<br>
# ChaiinerMNを用いたプログラムの作成方法の詳細については下記を参照してください。<br>
# https://chainermn.readthedocs.org/en/latest/tutorial<br>
#
#
# ひとまず、ここでは、どのようなオプションがあるか確認してみましょう。
python chainermn-1.0.0b2/examples/mnist/train_mnist.py -h
# "-g"オプションでGPUを使った実行をすることができます。<br>
# "-e 数値"オプションでepch数を指定することができます。<br>
# "-b 数値"オプションでミニバッチサイズを指定することができます。このミニバッチサイズに並列に実行するプロセス数を掛けたものが全体のバッチサイズとなります。<br>
# このtrain_mnist.pyをMPIを使って実行します。<br>
# MPIを使って並列にプロセスを実行する場合にはmpirun (またはmpiexec)を使用します。<br>
# どのようなオプションがあるか確認してみましょう。
mpirun -h
# "-n 数値"オプションで並列に実行するプロセス数を指定することができます。<br>
# 例えば、AAICは1ノードあたり8GPU搭載しているので、1ノードで実行する場合は"-n 8"とします。
# 2ノードで実行する場合は16GPU(2ノード×8GPU)使うことになるとで"-n 16"とします。
#
# MPIでPythonプログラムを実行する場合は、<br>
# mpirun [MPIのオプション] python [Pythonのプログラム名] [Pythonのプログラムのオプション]<br>
# という順で引数を記述します。
# ここでは、GPUを8台用いて、Epoch数を20、ミニバッチサイズを32(全体のバッチサイズは256)として実行します。
#
# インタラクティブノードにはGPUが搭載されていないため、以下のサンプルは、
# ```
# $ ssh gpx001.aaic.hpcc.jp
# ```
# とGPUを搭載した異なる計算ノード上で実行することができます。
mpirun -np 8 python chainermn-1.0.0b2/examples/mnist/train_mnist.py --epoch 20 --batchsize 100 -g
# 以下のようなエラーが出力される場合がありますが多くの場合実行に問題ありません。<br>
# mpirunのオプションに”--mca mpi_warn_on_fork 0”を追加することで出力を抑制することができます。<br>
# 詳細は下記のサイトを参照してください。<br>
# https://github.com/chainer/chainermn/blob/master/docs/source/tutorial/tips_faqs.rst
#
# ```
# --------------------------------------------------------------------------
# A process has executed an operation involving a call to the
# "fork()" system call to create a child process. Open MPI is currently
# operating in a condition that could result in memory corruption or
# other system errors; your job may hang, crash, or produce silent
# data corruption. The use of fork() (or system() or other calls that
# create child processes) is strongly discouraged.
#
# The process that invoked fork was:
#
# Local host: [[25170,1],1] (PID 29290)
#
# If you are *absolutely sure* that your application will successfully
# and correctly survive a call to fork(), you may disable this warning
# by setting the mpi_warn_on_fork MCA parameter to 0.
# --------------------------------------------------------------------------
# ```
# 無事実行されると、resultというディレクトリにtrain_mnist.pyの実行結果が出力されます。
ls result
# cg.dotはDOT言語で記述されたネットワーク構造のファイル、logはJSONで記述された実行時間、エポック数、反復回数、精度などを記述したファイルになります。
| 2/5_chainermn_run.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
data=pd.read_csv("https://archive.ics.uci.edu/ml/machine-learning-databases/abalone/abalone.data")
data.head()
names=['Sex' ,'Length','Diameter','Height','Whole weight','Shucked weight','Viscera weight','Shell weight','Rings']
data.columns=names
data.head()
data.tail()
data.to_csv('Abalone_DataSet.csv')
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
import warnings
warnings.filterwarnings('ignore')
data1=pd.read_csv("/home/manikanta/Documents/ML/classification/Navie_bayes/Abalone_DataSet.csv")
data1.isna().sum()
data1.describe()
data1.info()
data1.shape
import seaborn as sns
sns.heatmap(data1.isnull(), cbar=False)
# find categorical variables
categorical = [var for var in data.columns if data1[var].dtype=='O']
print('There are {} categorical variables\n'.format(len(categorical)))
print('The categorical variables are :\n\n', categorical)
# find numerical variables
numerical = [var for var in data.columns if data1[var].dtype!='O']
print('There are {} numerical variables\n'.format(len(numerical)))
print('The numerical variables are :', numerical)
# view frequency counts of values in categorical variables
for var in categorical:
print(data1[var].value_counts())
# find numerical variables
numerical = [var for var in data.columns if data1[var].dtype!='O']
print('There are {} numerical variables\n'.format(len(numerical)))
print('The numerical variables are :', numerical)
# view the numerical variables
data1[numerical].head()
data1.columns
# +
col_names = ['Unnamed: 0', 'Sex', 'Length', 'Diameter', 'Height', 'Whole weight', 'Shucked weight', 'Viscera weight', 'Shell weight', 'Rings']
data1.columns = col_names
data1.columns
# -
# Import label encoder
from sklearn import preprocessing
# label_encoder object knows how to understand word labels.
label_encoder = preprocessing.LabelEncoder()
# Encode labels in column 'species'.
data1['Sex']= label_encoder.fit_transform(data1['Sex'])
data1['Sex'].unique()
data1.head()
x=data1.drop(['Unnamed: 0','Sex'],axis=1)
y=data1['Sex']
# split X and y into training and testing sets
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(x, y, test_size = 0.30, random_state = 1)
print(X_train.shape)
print(X_test.shape)
print(y_train.shape)
print(y_test.shape)
cols = X_train.columns
# Feature Scalling
from sklearn.preprocessing import RobustScaler
scaler = RobustScaler()
X_train = scaler.fit_transform(X_train)
X_test = scaler.transform(X_test)
X_train = pd.DataFrame(X_train, columns=[cols])
X_test = pd.DataFrame(X_test, columns=[cols])
X_train.head(2)
# +
# train a Gaussian Naive Bayes classifier on the training set
from sklearn.naive_bayes import GaussianNB
# instantiate the model
gnb = GaussianNB()
# fit the model
gnb.fit(X_train, y_train)
# -
# print the scores on training and test set
print('Training set score: {:.4f}'.format(gnb.score(X_train, y_train)))
print('Test set score: {:.4f}'.format(gnb.score(X_test, y_test)))
y_pred = gnb.predict(X_test)
y_pred
from sklearn.metrics import accuracy_score
print('Model accuracy score: {0:0.4f}'. format(accuracy_score(y_test, y_pred)))
# +
# Print the Confusion Matrix and slice it into four pieces
from sklearn.metrics import confusion_matrix
cm = confusion_matrix(y_test, y_pred)
print('Confusion matrix\n\n', cm)
print('\nTrue Positives(TP) = ', cm[0,0])
print('\nTrue Negatives(TN) = ', cm[1,1])
print('\nFalse Positives(FP) = ', cm[0,1])
print('\nFalse Negatives(FN) = ', cm[1,0])
# -
from sklearn.metrics import classification_report
print(classification_report(y_test, y_pred))
TP = cm[0,0]
TN = cm[1,1]
FP = cm[0,1]
FN = cm[1,0]
# print classification accuracy
classification_accuracy = (TP + TN) / float(TP + TN + FP + FN)
print('Classification accuracy : {0:0.4f}'.format(classification_accuracy))
# print classification error
classification_error = (FP + FN) / float(TP + TN + FP + FN)
print('Classification error : {0:0.4f}'.format(classification_error))
# print precision score
precision = TP / float(TP + FP)
print('Precision : {0:0.4f}'.format(precision))
recall = TP / float(TP + FN)
print('Recall or Sensitivity : {0:0.4f}'.format(recall))
specificity = TN / (TN + FP)
print('Specificity : {0:0.4f}'.format(specificity))
true_positive_rate = TP / float(TP + FN)
print('True Positive Rate : {0:0.4f}'.format(true_positive_rate))
# print the first 10 predicted probabilities of three classes- 0 and 1 and 2
y_pred_prob = gnb.predict_proba(X_test)[0:10]
y_pred_prob
# store the probabilities in dataframe
y_pred_prob_df = pd.DataFrame(data=y_pred_prob, columns=['0','1','2'])
y_pred_prob_df
# print the first 10 predicted probabilities for classes
gnb.predict_proba(X_test)[0:10, 1]
y_pred1 = gnb.predict_proba(X_test)[:, 1]
# +
# Applying 10-Fold Cross Validation
from sklearn.model_selection import cross_val_score
scores = cross_val_score(gnb, X_train, y_train, cv = 10, scoring='accuracy')
print('Cross-validation scores:{}'.format(scores))
# -
# compute Average cross-validation score
print('Average cross-validation score: {:.4f}'.format(scores.mean()))
| NaiveBayes/Naive_Bayes_abalone_dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/hieuza/fun/blob/main/Edit_distance_%2B_visualization.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="9guB09LLUQes"
import numpy as np
import pandas as pd
from typing import Tuple
def min_edit_distance(
source: str, target: str, ins_cost: int = 1, del_cost: int = 1,
rep_cost: int = 1) -> Tuple[np.ndarray, int]:
nrows = len(source)
ncols = len(target)
D = np.zeros((nrows + 1, ncols + 1), dtype=int)
D[:, 0] = range(nrows + 1)
D[0, :] = range(ncols + 1)
for row in range(1, nrows + 1):
for col in range(1, ncols + 1):
real_rep_cost = rep_cost * (source[row - 1] != target[col - 1])
D[row,col] = min(
D[row - 1, col] + del_cost,
D[row, col - 1] + ins_cost,
D[row - 1, col - 1] + real_rep_cost)
return D, D[nrows, ncols]
# + colab={"base_uri": "https://localhost:8080/"} id="cUTUYKKTV8M1" outputId="60c6fe2d-6ca4-4b04-b9cc-d2066f441061"
source = 'cat'
target = 'match'
matrix, dist = min_edit_distance(source, target, 1, 1, 1)
df = pd.DataFrame(matrix, index=list('#' + source), columns=list('#' + target))
print(df)
# + colab={"base_uri": "https://localhost:8080/"} id="Q5qQU0nNWaRm" outputId="8520a78e-249f-4d7a-e2ac-1bdda09988d3"
def visualize_min_edit_distance(source: str, target: str, ins_cost: int, del_cost: int, rep_cost: int):
"""Visualize the transformation from a given source to a target."""
D, _ = min_edit_distance(source, target, ins_cost, del_cost, rep_cost)
r = len(source)
c = len(target)
print(f'Edit distance: {D[r][c]}')
ops = []
while r != 0 or c != 0:
is_ins = c > 0 and D[r, c] == D[r, c - 1] + ins_cost
is_del = r > 0 and D[r, c] == D[r - 1, c] + del_cost
r_cost = rep_cost * (r > 0 and c > 0 and source[r - 1] != target[c - 1])
is_rep = r > 0 and c > 0 and D[r, c] == D[r - 1, c - 1] + r_cost
assert is_ins or is_del or is_rep
op = ''
if is_ins:
op = ('I', c-1, target[c-1])
c -= 1
elif is_del:
op = ('D', r-1, source[r-1])
r -= 1
elif is_rep:
if r_cost > 0:
op = ('R', r-1, target[c-1])
c -= 1
r -= 1
if op:
ops.insert(0, op)
edit = source[:]
offset = 0
steps = np.arange(len(ops)) + 1
pre_edits = []
op_desc = []
edits = []
for step, op in enumerate(ops):
pre_edit = edit[:]
pos, char = op[1], op[2]
if op[0] == 'I':
edit = edit[:pos] + char + edit[pos:]
offset += 1
elif op[0] == 'D':
pos += offset
edit = edit[:pos] + edit[pos+1:]
offset -= 1
elif op[0] == 'R':
pos += offset
edit = edit[:pos] + char + edit[pos+1:]
pre_edits.append(pre_edit)
op_desc.append(f'{op[0]},{pos},{char}')
edits.append(edit)
df = pd.DataFrame([pre_edits, op_desc, edits], columns=steps,
index=['[source]', '[op]', '[target]']).transpose()
print(df)
source = 'zz_mode'
target = 'code_y'
visualize_min_edit_distance(source, target, 1, 1, 1)
# + id="9DXVKjb9ZQmo"
| Edit_distance_+_visualization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## MNIST CNN
# %reload_ext autoreload
# %autoreload 2
# %matplotlib inline
from fastai.vision import *
# ### Data
# Untar the data for the MNIST dataset; save the path to `path`.
# Show the contents of the `path` directory.
# Create an `ImageList` from the folder `path`. Set `convert_mode` to `L`. Save it to variable `il`. What does this do?
# Show the first element in the `items` attribute in `il`.
# Set `defaults.cmap` to `'binary'`. What does this do?
# Take a look at `il`.
# Show the image in the first element of `il`. What's the difference in `il[0]` and `il.items[0]`?
# Split the imagelist by folder with training folder `training` and validation folder `testing`. Save to variable `sd`.
# Show `sd`.
# Show the contents of `path/'training'`.
# Label the examples from the folder they're in, and save the resulting `LabelLists` to `ll`. How do we call `label_from_folder` on an `ItemLists` when it's not actually a method on `ItemLists` (and `ItemLists` doesn't inherit from anything)?
# Show `ll`.
# Unpack the first item in the `train` property of `ll` into `x, y`.
# Show `x`. Print the shapes of `x` and `y`.
# Can you explain the relationships between `ItemList`, `ItemLists`, `LabelList`, and `LabelLists`?
# Create a 2-tuple of lists of transformations, one for training and one for validation. The training transformations should be `[*rand_pad(padding=3, size=28, mode='zeros')]` and the validation ones should be empty.
# Overwrite `ll` with the `transform`ed version of `ll` using `tfms`.
# Set `bs=128`.
# Set `data` to a `databunch` from `ll`, using `bs`, and normalize it. Note that we're not using `imagenet_stats` here because we're not using a pretrained model. What's the difference between a databunch and a `LabelLists`?
# Set `x` and `y` to the values in the first element in `data.train_ds`.
# Show x. Print y.
# Define a function `_plot` that takes a row `i` a column `j` and a matplotlib axis `ax` and the first x value in `train_ds` with the cmap `gray`.
# Assign a batch of data to `xb` and `yb`.
# Show the batch over three rows with a figsize of (5,5).
# ### Basic CNN with batchnorm
# Define a function `conv` with arguments `ni` and `nf` that creates a `nn.Conv2d` filter with `ni` inputs, `nf` filters, a kernel size of 3, stride of 2, and padding of 1.
# create a `nn.Sequential` that takes a 28x28 and starts with `ni` and `nf` of 1 and 8 respectively and adds blocks of (conv, batchnorm2d, relu) until there are 10 filters of size (1,1), and then flatten that.
# Instantiate a learner object that takes our data, model, a loss func of `nn.CrossEntropyLoss` and the `accuracy` metric.
# Print a summary of your model and look it over.
# Put `xb` on the GPU.
# Do `model(xb).shape`.
# Find an lr (end_lr=100; what does this do?)
# Plot the learning rates. You shoudl land somewhere around 1e-2.
# Fit a cycle with 3 epochs and a max_lr of 0.1.
# ### Refactor
# Take a look at the documentation for `conv_layer`.
# Define a new function `conv2` that takes in `ni`, the number of inputs, and `nf`, the number of filters, and creates a new conv layer with `stride=2`.
# Rewrite the model above in terms of `conv2`. For each layer, add a comment with the length/width of the square image.
# Create a learner (named `learn`) from the new model, same specs as above.
# Show `learn.summary`.
# Fit a cycle with 10 epochs, max_lr=0.1
# ### Resnet-ish
# Get help with `res_block`.
# Show the documentation for `res_block`.
# Create a model thaat alternates a conv2 with a res_block until you get down to a vector of outputs.
# Show the documentation for `res_block`.
# Create a function `conv_and_res` that takes `ni` and `nf` and returns a sequential model with a `conv2` with ni, nf and a res_block with `nf`.
# Rewrite the model in terms of `conv_and_res` and `conv2`.
# Create a `learn`er.
# Show a summary of the learner.
# Find the learning rate (Hint: `end_lr=100`)
# Fit a 12-epoch cycle with max_lr=0.05 (or whatever you just found for learning rate)
# Show the learner summary one more time.
# ## fin
| nbs/dl1/lesson7-resnet-mnist-ex.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="Ssib8PW0ZswG"
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# + colab={"base_uri": "https://localhost:8080/", "height": 368} id="TseQlxJanGTF" outputId="5dbf5bf5-3d4a-4ef3-d6f8-d0eff8a57469"
url='https://raw.githubusercontent.com/alura-cursos/agendamento-hospitalar/main/dados/A160324189_28_143_208.csv'
dados = pd.read_csv(url, encoding="ISO-8859-1", engine='python',
skiprows = 3, sep=";", skipfooter=12,
thousands=".", decimal=",", index_col=0).drop('Total', axis=1)
dados.head()
# + colab={"base_uri": "https://localhost:8080/"} id="EIqd-uJxnS47" outputId="e1073fd2-eb5e-443f-c638-ea31e18a7754"
dados.isnull().sum().sum()
# + colab={"base_uri": "https://localhost:8080/"} id="XmvtShcNnkKR" outputId="867fd0a1-6bd2-4a72-edb8-785bd6985017"
dados.info()
# + colab={"base_uri": "https://localhost:8080/", "height": 546} id="ZLtYQXiYnpNv" outputId="346496e3-0c2d-4db6-ca8f-75d75e390c26"
dados.loc['35 São Paulo':].select_dtypes('object')
# + colab={"base_uri": "https://localhost:8080/", "height": 368} id="u-F0oUZ1nyJS" outputId="fdb2d0be-56e8-462e-a85f-7d33978e27b8"
dados = dados.replace('-', np.nan)
dados.head()
# + colab={"base_uri": "https://localhost:8080/"} id="E2NOO58no83w" outputId="dd658f23-ff74-4971-bc59-d496299ba942"
dados.T.info()
# + colab={"base_uri": "https://localhost:8080/"} id="fqJDFTmGpG2x" outputId="f3f69145-a1ce-4783-bb05-36858b1ca068"
dados.isnull().sum().sum()
# + colab={"base_uri": "https://localhost:8080/", "height": 368} id="oAi-p2Cxpbk3" outputId="9c38e738-5281-4abe-8293-14868458e1e9"
dados = dados.astype('float64')
dados.head()
# + colab={"base_uri": "https://localhost:8080/"} id="EUIQ276hpg7i" outputId="88caa78b-b330-4870-965c-96fd517953d6"
dados.T.info()
# + colab={"base_uri": "https://localhost:8080/", "height": 402} id="mUA0sD5UpM9t" outputId="6ccf0682-8fbb-4c51-8960-5e8216c2051d"
dados.loc['35 São Paulo'].plot(figsize=(10, 6))
# + colab={"base_uri": "https://localhost:8080/", "height": 402} id="rrbqdlVWpm-r" outputId="ea51729f-d782-440c-9ae4-aea4c7c21e90"
dados.loc['50 Mato Grosso do Sul'].plot(figsize=(10, 6))
# + colab={"base_uri": "https://localhost:8080/"} id="1bLiLA82qIQB" outputId="839625c5-236c-4054-a144-e312833a422f"
dados_estado = dados.loc['35 São Paulo', '2004/Mai':'2007/Jun']
dados_estado
# + colab={"base_uri": "https://localhost:8080/"} id="HipzXO_pqMxi" outputId="9c765c30-3ec0-47da-85e1-03656f3028ec"
dados_estado.isnull().sum().sum()
# + colab={"base_uri": "https://localhost:8080/"} id="9pXCvuPPqX_V" outputId="b990e5c5-62cc-4341-96d3-23922ef9cb4e"
dados_estado.dropna().isnull().sum().sum()
# + colab={"base_uri": "https://localhost:8080/", "height": 411} id="XqVeYJiaqcUI" outputId="9e6036c2-8892-446f-c215-7aa4e28b6d87"
plt.figure(figsize=(20,6))
plt.subplot(121)
dados_estado.plot(marker='o')
plt.subplot(122)
dados_estado.dropna().plot(marker='o')
# + colab={"base_uri": "https://localhost:8080/"} id="ggYk2ugqqmsY" outputId="297cbe67-b5bb-42a5-d09a-84e11a5af35b"
dados_estado.fillna(value=dados_estado.mean())
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="cBf_wdcYq9h3" outputId="6ba9374d-0d7f-4060-a985-c6aff1e0433c"
pd.merge(dados_estado,dados_estado.fillna(value=dados_estado.mean()), left_index=True, right_index=True)
# + colab={"base_uri": "https://localhost:8080/", "height": 391} id="oWnofjVUrEES" outputId="21fb63bc-c36d-48f7-dcb7-4a3b2114e78b"
dados_estado.fillna(value=dados_estado.mean()).plot(marker='o',figsize=(10,6))
dados_estado.plot(marker='o')
plt.legend(['média', 'original'])
# + colab={"base_uri": "https://localhost:8080/"} id="WLe0cHdQrMBe" outputId="0a09add2-3974-4178-f3ca-e56f1488a7e9"
dados_estado.fillna(method='backfill')
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="0q1U-9tYrVhp" outputId="0b758e4f-3ff3-42ca-bd51-1b1c868cd712"
pd.merge(dados_estado,dados_estado.fillna(method='backfill'), left_index=True, right_index=True)
# + colab={"base_uri": "https://localhost:8080/", "height": 391} id="ECpMqqAxrkS0" outputId="74d7f7c1-596f-493c-936e-e630ccca4e73"
dados_estado.fillna(method='backfill').plot(marker='o',figsize=(10,6))
dados_estado.plot(marker='o')
plt.legend(['backfill', 'original'])
# + colab={"base_uri": "https://localhost:8080/"} id="AD53wImfuKid" outputId="2c2fb741-f508-4eb8-b701-0e2abb4ae595"
dados_estado.interpolate()
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="InZL3-QhuQoe" outputId="99a6faf8-1f3c-4b86-f8ed-a0c206d3aaad"
pd.merge(dados_estado,dados_estado.interpolate(), left_index=True, right_index=True)
# + colab={"base_uri": "https://localhost:8080/", "height": 391} id="JR_MbblkuXQQ" outputId="f69be723-2e52-4b03-c1a6-a6b4c86c19ef"
dados_estado.interpolate().plot(marker='o',figsize=(10,6))
dados_estado.plot(marker='o')
plt.legend(['interpolate', 'original'])
# + colab={"base_uri": "https://localhost:8080/", "height": 391} id="G05nEZljudF-" outputId="1f44b619-19fe-4dd5-a0cf-d72ecf1bc96b"
dados_estado.fillna(value=dados_estado.mean()).plot(marker='o',figsize=(10,6))
dados_estado.fillna(method='backfill').plot(marker='o',figsize=(10,6))
dados_estado.interpolate().plot(marker='o',figsize=(10,6))
dados_estado.plot(marker='o')
plt.legend(['media', 'backfill', 'interpolate', 'original'])
| notebooks/modulo_01/resolucoes_modulo_01/.ipynb_checkpoints/notebook_live_monitoria_1_bootcamp_2-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Chapter 8 - Local Realism
# ## with Alice and Bob
from numpy import sin,cos,pi,sqrt,angle,exp,deg2rad,arange,rad2deg
import matplotlib.pyplot as plt
from qutip import *
# %matplotlib inline
H = Qobj([[1],[0]])
V = Qobj([[0],[1]])
# First define the projection operator for a state at angle $\theta$:
def P(theta):
"""The projection operator for a state at angle theta"""
theta_ket = cos(theta)*H + sin(theta)*V
return theta_ket*theta_ket.dag()
# Create the projection operators for each of the angles, two for Alice, two for Bob
Pa1 = P(deg2rad(19))
Pa2 = P(deg2rad(-35))
Pb1 = P(deg2rad(-19))
Pb2 = P(deg2rad(35))
# Create the state $\big|\psi\big\rangle = \sqrt{0.2} \big|H,H\big\rangle + \sqrt{0.8} \big|V,V\big\rangle$:
psi=sqrt(0.2)*tensor(H,H) + sqrt(0.8)*tensor(V,V)
# Now, find the joint probability that Alice measures A1 and Bob measures B1. We do this by finding the expectation value of the projection operator for the joint state $\big|\theta_{A1},\theta_{B1}\big\rangle$. This is formed as the tensor product of the two appropriate projection operators. **In these tensor products, be sure to put Alice's operator first, then Bob's (just like we did for the signal and idler photons).** Each operator acts on the photon corresponding to the order in the `tensor()` function.
#
# Notice we'll be using a new function `expect()`. This is equivalent to putting the operator in between the state bra and ket:
P1 = expect(tensor(Pa1,Pb1),psi) # joint for A1, B1 (expect 0.09)
P2 = psi.dag()*tensor(Pa1,Pb1)*psi
P1 == P2.data[0,0] # The only difference is that we have to pull out the value
# from the Qobj using the .data[0,0] method so we can compare it to result from `expect`
P1
# Find the conditional probability $P(\theta_{B2}|\theta_{A1}) = \frac{P(\theta_{B2},\theta_{A1})}{P(\theta_{A1})}$
# B2 conditioned on A1 (expect 1)
Prob_b2_a1 = expect(tensor(Pa1,Pb2),psi)
#(psi.dag()*tensor(Pa1,Pb2)*psi).data[0,0] # the joint probability
Prob_a1 = expect(tensor(Pa1,qeye(2)),psi)
#(psi.dag()*tensor(Pa1,qeye(2))*psi).data[0,0] # the singular probability
Prob_b2a1 = Prob_b2_a1 / Prob_a1 # the conditional probability
Prob_b2a1
# Find the conditional probability $P(\theta_{A2}|\theta_{B1}) = \frac{P(\theta_{A2},\theta_{B1})}{P(\theta_{B1})}$
# A2 conditioned on B1 (expect 1)
# can do it all on one line:
expect(tensor(Pa2,Pb1),psi) / expect(tensor(qeye(2),Pb1),psi)
expect(tensor(Pa2,Pb2),psi) # joint for A2, B2 (classically expect 0.09, QM says 0)
# This is what we described in class.
# ## What if the state was just $|H,H\rangle$?
psi2=tensor(H,H)
expect(tensor(Pa1,Pb1),psi2) # joint for A1, B1 (expect 0.09)
# B2 conditioned on A1:
expect(tensor(Pa1,Pb2),psi2) / expect(tensor(Pa1,qeye(2)),psi2)
# A2 conditioned on B1
expect(tensor(Pa2,Pb1),psi2) / expect(tensor(qeye(2),Pb1),psi2)
# joint for A2, B2
expect(tensor(Pa2,Pb2),psi2)
# This is harder to interpret, but we clearly have different probabilities. Finally, check if we had used a mixed state:
# ## A mixed state instead of the pure (entangled state).
# Here we have to use the density matrix (since a `ket` cannot describe a mixed state). First some background:
# QuTiP has a function that gives the density matrix from a `ket` state: `ket2dm`.
rho_mix = 0.2 * ket2dm(tensor(H,H)) + 0.8 * ket2dm(tensor(V,V))
rho_mix
# joint for A1, B1
expect(tensor(Pa1,Pb1),rho_mix)
# B2 conditioned on A1
expect(tensor(Pa1,Pb2),rho_mix) / expect(tensor(Pa1,qeye(2)),rho_mix)
# A2 conditioned on B1
expect(tensor(Pa2,Pb1),rho_mix) / expect(tensor(Pb1,qeye(2)),rho_mix)
# joint for A2, B2:
expect(tensor(Pa2,Pb2),rho_mix)
# We see that $P(\theta_{B2},\theta_{A2}) > P(\theta_{B1},\theta_{A1})$ as we said in class for a state that obeys realism.
# ## Now repeat with the pure state but using density matrix techniques.
# This isn't going to tell us anything new, but it shows how to work with the density matrix if you already know the `ket` state.
rho_pure = ket2dm(psi) # convert from a ket to a density matrix (dm)
rho_pure
# The calculations are actually the same in QuTiP, the `expect` function takes either a `ket` state or a density matrix.
# joint for A1, B1
expect(tensor(Pa1,Pb1),rho_pure)
# B2 conditioned on A1
expect(tensor(Pa1,Pb2),rho_pure) / expect(tensor(Pa1,qeye(2)),rho_pure)
# A2 conditioned on B1
expect(tensor(Pa2,Pb1),rho_pure) / expect(tensor(Pb1,qeye(2)),rho_pure)
# joint for A2, B2:
expect(tensor(Pa2,Pb2),rho_pure)
# These all agree (as they should).
# ## Explore the angles in more detail:
# Why these angles, 19 and 35?
psi=sqrt(0.2)*tensor(H,H) + sqrt(0.8)*tensor(V,V)
angles = arange(1,90,1)
rads = deg2rad(angles)
# Make a list of the probability of joint measurements for a pair of angles:
# +
out = []
for r in rads:
out.append(expect(tensor(P(-r),P(r)),psi))
plt.plot(angles,out,".") # plot in units of pi
# -
# We see that the joint probabilities have a zero at 35˚. Now plug that in to one of the conditional probabilities and see what angle for the conditional probability gives 1:
# +
out = []
for r in rads:
out.append(expect(tensor(P(r),P(deg2rad(35))),psi) / expect(tensor(P(r),qeye(2)),psi))
plt.plot(angles,out,".")
# -
# So only 19 and 35 work. Now, can you derive 19 and 35 given only the state $|\psi\rangle$? Try the first plot, i.e. calculate the joint probability $P(\theta_A,\theta_B)$
# # Solution
# Using the state, write the projection operators for a two photon state with angles $\theta_A$ and $\theta_B$. First, recall $$\big|\theta_i\big\rangle = \cos\theta_i\big|H\big\rangle + \sin\theta_i\big|V\big\rangle.$$ Next, form the two-photon state: $$\big|\theta_A,\theta_B\big\rangle = \big|\theta_A\big\rangle \otimes \big|\theta_B\big\rangle = \left(\cos\theta_A\big|H\big\rangle + \sin\theta_A\big|V\big\rangle\right) \otimes \left(\cos\theta_B\big|H\big\rangle + \sin\theta_B\big|V\big\rangle\right)$$
# which we can reduce to:
# $$=\cos\theta_A\cos\theta_B\big|H,H\big\rangle + \cos\theta_A\sin\theta_B\big|H,V\big\rangle + \sin\theta_A\cos\theta_B\big|V,H\big\rangle + \sin\theta_A\sin\theta_B\big|V,V\big\rangle.$$
# Find the probability of a joint measurement of polarizations $\theta_A$ and $\theta_B$:
# $$P(\theta_A,\theta_B) = \big|\big\langle\psi\big|\theta_A,\theta_B\big\rangle\big|^2$$
# Since $\big|\psi\big\rangle$ only has $\big|H,H\big\rangle$ and $\big|V,V\big\rangle$ terms, this probability only has two terms:
# $$P(\theta_A,\theta_B) = \left|\sqrt{0.2}\cos\theta_A\cos\theta_B + \sqrt{0.8}\sin\theta_A\sin\theta_B\right|^2$$
# Plot is shown below for $\theta_A = -\theta_B$ and it agrees perfectly with our model above.
# +
# Solution:
# For the first plot, we can show the joint probability for two angles is given by:
plt.plot(rad2deg(rads),(sqrt(0.2)*cos(-rads)*cos(rads) + sqrt(0.8)*sin(-rads)*sin(rads))**2)
# -
# ## Challenge:
# If we change the state to $\big|\psi\big\rangle = \sqrt{0.8} \big|H,H\big\rangle + \sqrt{0.2} \big|V,V\big\rangle$, the two angles that work for this state.
# +
# Solution
psi3=sqrt(0.8)*tensor(H,H) + sqrt(0.2)*tensor(V,V)
out = []
for r in rads:
out.append(expect(tensor(P(-r),P(r)),psi3))
plt.plot(angles,out,".") # plot in units of pi
# +
# Solution
out = []
for r in rads:
out.append(expect(tensor(P(r),P(deg2rad(55))),psi3) / expect(tensor(P(r),qeye(2)),psi3))
plt.plot(angles,out,".")
| Chapter 8 - Alice and Bob.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Задание №1
#
# ## Теория
#
# Этот ноутбук содержит вспомогательный материал для выполнения первого задания, в котором надо предсказать целевую переменную по двум признакам с помощью линейной регрессии. Хочу сразу предупредить, что здесь собрана вводная информация, причем в некоторых аспектах я намеренно привел упрощенное или не до конца полное описание. В дальнейшем мы отдельно размерем все эти аспекты.
#
# Для начала, вспомним, что такое линейная регрессия и как выглядит алгоритм подбора оптимальных параметров предсказания.
#
# Назовем множество имеющихся примеров **обучающей выборкой** или **тренировочными данными**. Обучающая выборка — это множество пар $D = \{(\vec{x^{(i)}},y^{(i)})\}_{i=1,2,...,n}$, где $n$ - размер выборки. В каждой паре через $y \in R$ обозначена целевая переменная (или ответ), который соответствует вектору признаков $\vec{x} \in R^{m\times1}$. Необходимо научиться предсказывать значение $y$ по $\vec{x}$. Обозначим предсказанное значение через $\hat{y}$. В линейной регрессии предсказание делается с помощью следующей формулы:
#
# \begin{equation}
# \hat{y} = h(\vec{x},\vec{\theta}) = \vec{x}^T\vec{\theta} = \langle \vec{x}, \vec{\theta} \rangle = \sum_{j=1}^{m} x_j\theta_j,
# \end{equation}
#
# где через $h(\vec{x},\vec{\theta})$ обозначена функция предсказания, а $\vec{\theta} \in R^{m\times1}$ — вектор параметров линейной регрессии. **Обучение** модели на основе линейной регрессии заключается в подборе $\vec{\theta}$ так, чтобы предсказанные значения $\hat{y}^{(i)}$ были как можно ближе к целевым переменным $y^{(i)}$ из обучающей выборки. Термин "ближе" значит, что есть способ измерения похожести $\hat{y}^{(i)}$ и $y^{(i)}$. В рассматриваемой задаче в качестве критерия похожести можно воспользоваться квадратичным отклонением:
#
# \begin{equation}
# e^{(i)} = \left( y^{(i)} - \hat{y}^{(i)} \right)^2.
# \end{equation}
#
# Значение $e^{(i)}$, или ошибка предсказания, показывает насколько сильно предсказанное значение целевой переменной отличается от требуемого значения для $i$-го примера обучающей выборки. Средняя ошибка предсказания, или **эмпирический риск**, на обучающей выборке определяется как
#
# \begin{equation}
# E(D, \vec{\theta}) = \frac{1}{n} \sum_{i=1}^{n} e^{(i)} = \frac{1}{n} \sum_{i=1}^{n} \left( y^{(i)} - h(\vec{x^{(i)}},\vec{\theta}) \right)^2 = \frac{1}{n}\sum_{i=1}^{n} \left(y^{(i)} - \sum_{j=1}^{m} x^{(i)}_j\theta_j \right)^2.
# \end{equation}
#
# Таким образом, задача обучения формулируется как "найти такой вектор параметров линейной регрессии, при котором минимальна средняя ошибка предсказания". Для поиска оптимального $\vec{\theta}$ можно воспользоваться методом градиентного спуска.
#
# Рассмотрим идею этого метода. "Градиент" функции в точке показывает направление роста функции в этой точке. Фактически, градиент -- это обобщение понятия производной на многомерный случай. Для рассмотренной нами функции эмпирического риска:
#
# \begin{equation}
# \nabla E(D, \vec{\theta}) \big\vert_{\vec{\theta}=\vec{t}} =
# \left(
# \begin{aligned}
# & \frac{\partial E(D, \vec{\theta})}{\partial \theta_0} \big\vert_{\vec{\theta}=\vec{t}} \\
# & \frac{\partial E(D, \vec{\theta})}{\partial \theta_1} \big\vert_{\vec{\theta}=\vec{t}} \\
# & ... \\
# & \frac{\partial E(D, \vec{\theta})}{\partial \theta_m} \big\vert_{\vec{\theta}=\vec{t}} \\
# \end{aligned}
# \right).
# \end{equation}
#
# Нас интересует минимизация эмпирического риска. Это значит, что можно выбрать некоторое, например случайное, значение вектора $\vec{\theta}$, посчитать в нем значение градиента, а потом изменить $\vec{\theta}$ так, чтобы значение фунцкии средней ошибки уменьшилось, т.е. изменить $\vec{\theta}$ в направлении, обратном направлению градиента. Далее повторить эту процедуру для измененного $\vec{\theta}$. Если функция эмпирического риска является выпуклой и унимодальной (имеет один оптимум), то подобный итеративный пересчет через достаточное число итераций (спусков) приведет к значению параметров $\vec{\theta}$, в которых достигается минимум средней ошибки предсказания.
#
# Опишем теперь алгоритм поиска оптимальных параметров линейной регрессии с помощью градиентного спуска.
#
# **Инициализация:**
# - задать $\alpha$ - скорость градиентного спуска;
# - задать начальные значения переменных: $\vec{\theta}^{(0)}$ = random, k = 0.
#
# **Повторять, пока не выполнятся условия остановки**
#
# \begin{equation}
# \vec{\theta}^{(k+1)} = \vec{\theta}^{(k)} - \alpha E(D, \vec{\theta}) \big\vert_{\vec{\theta}=\vec{\theta^{(k)}}} \\
# k = k + 1
# \end{equation}
#
# Условия остановки:
# - достигнуто максимальное число итераций градиентного спуска;
# - значения параметров $\theta$ почти не меняются от итерации к итерации;
# - значение градиента близко у нулю.
#
# ## Практика
#
# Перейдем теперь к практической части, где нужно реализовать линейную регрессию. Далее приведен "скелет" программы, в которую Вам необходимо встроить недостающие фрагменты. Места для Вашег кода помечены "TODO".
#
# Программа состоит из следующих блоков.
#
# 1. Подключение всех необходимых библиотек.
# 2. Чтение и вывод тренировочных данных.
# 3. Прототипы функций, которые Вам надо заполнить.
# 4. Цикл обучения.
# ### Подключение библиотек.
#
# В python есть много готовых библиотек для работы с данными. Наши тренировочные данные хранятся в csv файлах и для их парсинга и отображения можно воспользоваться удобной библиотекой pandas.
#
# Удобнее всего работать с данными, представленными в формате матриц и векторов. Для работы с ними в python есть библиотека numpy.
#
# Отображение графиков можно сделать с помощью matplotlib.
#
# Удобный вывод прогресса - tqdm.
# +
# Step 1. Import all the necessary packages
import sys
import tqdm
import pandas as pd
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
from matplotlib import cm
import numpy as np
# -
# ### Чтение и вывод тренировочных данных.
#
# Для чтения данных воспользуемся методом read_csv из pandas. Эта функция возвращает специальную структуру данных DataFrame, которая содержит много методов для вывода данных и расчета различных статистик по ним. Так как данные разбиты на 2 файла, перед выводом на экран первых десяти сэмплов, сконкатенируем их в один DataFrame.
#
# Затем преобразуем данные из DataFrame в матрицы numpy.
# +
# Step 2. Parse and visualize data
# parse train data: read CSV files with train features (train_x) and train targets (train_y)
x_train = pd.read_csv("train_x.csv", header=None)
y_train = pd.read_csv("train_y.csv", header=None)
# show first 10 samples
pd.concat([x_train, y_train], axis=1).head(10)
# +
# convert pandas dataframe to numpy arrays and matrices and diplay the dimensions of train dataset
x_train = x_train.to_numpy()
y_train = y_train.to_numpy()
print("Shape of train features:", x_train.shape)
print("Shape of train targets:", y_train.shape)
# -
# ### Прототипы функций.
#
# * predict_fn - предсказание с помощью линейной регрессии;
# * loss_fn - расчет среднего значения ошибки предсказания;
# * gradient_fn - расчет градиента в точке.
#
# Вам необходимо реализовать predict_fn и gradient_fn.
# +
# Step 3. Prototypes.
# In this demo we will use linear regression to predict targets from features.
# In linear regression model with parameters thetas
# the prediction y is calculated from features x using linear combination of x and thetas.
# For example, for the case of 2 features:
# y = theta_0 * x_o + theta_1 * x_1
# Let's define some helper functions
def predict_fn(x, thetas):
'''
Predict target from features x using parameters thetas and linear regression
param x: input features, shape NxM, N - number of samples to predict, M - number of features
param thetas: vector of linear regression parameters, shape Mx1
return y_hat: predicted scalar value for each input samples, shape Nx1
'''
# TODO: calculate y_hat using linear regression
y_hat = np.zeros((x.shape[0], 1))
return y_hat
def loss_fn(x_train, y_train, thetas):
'''
Calculate average loss value for train dataset (x_train, y_train).
param x_train: input features, shape NxM, N - number of samples to predict, M - number of features
param y_train: input tagrets, shape Nx1
param thetas: vector of linear regression parameters, shape Mx1
return loss: predicted scalar value for each input samples, shape Mx1
'''
y_predicted = predict_fn(x_train, thetas)
loss = np.mean(np.power(y_train - y_predicted, 2))
return loss
def gradient_fn(x_train, y_train, thetas):
'''
Calculate gradient value for linear regression.
param x_train: input features, shape NxM, N - number of samples to predict, M - number of features
param y_train: input tagrets, shape Nx1
param thetas: vector of linear regression parameters, shape Mx1
return g: predicted scalar value for each input samples, shape Mx1
'''
# TODO: calculate vector gradient
g = np.zeros_like(thetas)
return g
# -
# ### Цикл обучения
#
# Ниже дана упрощенная реализация градиентного спуска для поиска оптимальных параметров линейной регрессии. Вам необходимо подобрать значения скорости (alpha), максимальное число итераций (MAX_ITER) и условия остановки.
# +
# Step 4. Gradient descent.
# now let's find optimal parameters using gradient descent
MAX_ITER = 100000
thetas = np.random.randn(2, 1)
alpha = 1e-3
progress = tqdm.tqdm(range(MAX_ITER), "Training", file=sys.stdout)
loss_val = loss_fn(x_train, y_train, thetas)
progress.set_postfix(loss_val=loss_val)
for iter in progress:
gradient = gradient_fn(x_train, y_train, thetas)
thetas = thetas - alpha*gradient
# TODO: add stop conditions
# if stop_condition is True:
# progress.close()
# loss_val = loss_fn(x_train, y_train, thetas)
# print("Stop condition detected")
# print("Final loss:", loss_val)
# break
if iter % 100 == 0:
loss_val = loss_fn(x_train, y_train, thetas)
progress.set_postfix(loss_val=f"{loss_val:8.4f}", thetas=f"{thetas[0][0]:5.4f} {thetas[1][0]:5.4f}")
progress.close()
# -
# Выведем несколько предсказаний для примера.
for i in range(10):
y_hat = predict_fn(x_train[i], thetas)
print("Target: ", y_train[i][0], ", predicted:", y_hat[0][0])
# Скорее всего, в результате получится довольно большое значение ошибки и предсказания будут не очень точными. Как поднять точность, мы рассмотрим далее :)
| topic_1_linear_regression/task1_todo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.6 64-bit (''theegg_env'': venv)'
# language: python
# name: python3
# ---
# ## Flow Diagram
# 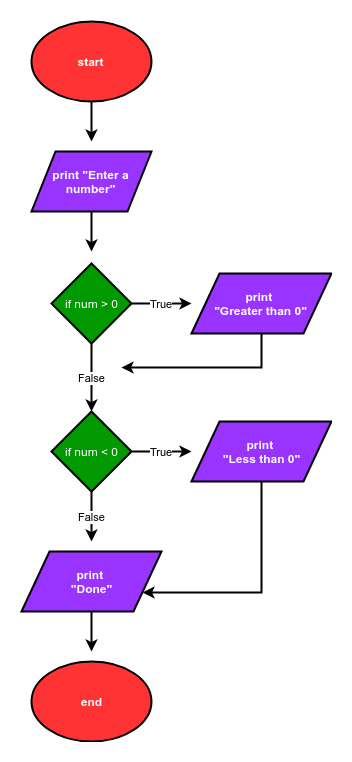
# start
num = input('Enter a number: ')
num = float(num)
if num>0:
print('num greater than zero')
if num<0:
print('num less than zero')
print('Done')
# end
| tarea_80/flow_diagram.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import tensorflow as tf
tf.enable_eager_execution()
import tensorflow.contrib.eager as tfe
# ## Simple CNN
# Here we create a simple convolutional neural network (CNN)to recognize hand-written digits (MNIST). We start by creating a simple alexnet CNN model.
#
# 
from tensorflow.keras.layers import Dense, Convolution2D, MaxPooling2D, Flatten, BatchNormalization, Dropout
from tensorflow.keras.models import Sequential
def create_model():
model = Sequential()
model.add(Convolution2D(filters = 16, kernel_size = 3, padding = 'same', input_shape = [28, 28, 1], activation = 'relu'))
model.add(MaxPooling2D(pool_size = (2,2)))
model.add(BatchNormalization())
model.add(Convolution2D(filters = 32, kernel_size = 3, padding = 'same', activation = 'relu'))
model.add(MaxPooling2D(pool_size = (2,2)))
model.add(BatchNormalization())
model.add(Flatten())
model.add(Dense(units = 100, activation = 'relu'))
model.add(Dropout(0.5))
model.add(Dense(units = 10 , activation = 'softmax'))
return model
model = create_model()
model.summary()
import numpy as np
model(np.zeros((10,28,28,1), np.float32))
# ## Load Mnist Data
(x_train, y_train), (x_test, y_test) = tf.keras.datasets.mnist.load_data()
# ## Look data
import matplotlib.pyplot as plt
print('The label is ',y_train[0])
plt.imshow(x_train[0])
plt.show()
# ## Preprocessing the dataset
# +
import numpy as np
N = x_train.shape[0]
#normalization and convert to batch input
x_train = tf.expand_dims(np.float32(x_train)/ 255., 3)
x_test = tf.expand_dims(np.float32(x_test )/ 255., 3)
#one hot encoding
y_train = tf.one_hot(y_train, 10)
y_test = tf.one_hot(y_test , 10)
# -
# ## Get Random Batch
import numpy as np
def get_batch(batch_size = 32):
r = np.random.randint(0, N-batch_size)
return x_train[r: r + batch_size], y_train[r: r + batch_size]
# ## Design the loss, gradient and accuract metric
# +
#evaluate the loss
def loss(model, x, y):
prediction = model(x)
return tf.losses.softmax_cross_entropy(y, logits=prediction)
#record the gradient with respect to the model variables
def grad(model, x, y):
with tf.GradientTape() as tape:
loss_value = loss(model, x, y)
return tape.gradient(loss_value, model.variables)
#calcuate the accuracy of the model
def accuracy(model, x, y):
#prediction
yhat = model(x)
#get the labels of the predicted values
yhat = tf.argmax(yhat, 1).numpy()
#get the labels of the true values
y = tf.argmax(y , 1).numpy()
return np.sum(y == yhat)/len(y)
# -
# ## Init
# +
i = 1
batch_size = 64
epoch_length = N // batch_size
epoch = 0
epochs = 5
#use Adam optimizer
optimizer = tf.train.AdamOptimizer()
#record epoch loss and accuracy
loss_history = tfe.metrics.Mean("loss")
accuracy_history = tfe.metrics.Mean("accuracy")
# -
# ## training
if tfe.num_gpus() > 0:
with tf.device("/gpu:0"):
while epoch < epochs:
#get next batch
x, y = get_batch(batch_size = batch_size)
# Calculate derivatives of the input function with respect to its parameters.
grads = grad(model, x, y)
# Apply the gradient to the model
optimizer.apply_gradients(zip(grads, model.variables),
global_step=tf.train.get_or_create_global_step())
#record the current loss and accuracy
loss_history(loss(model, x, y))
accuracy_history(accuracy(model, x, y))
if i % epoch_length == 0:
print("epoch: {:d} Loss: {:.3f}, Acc: {:.3f}".format(epoch, loss_history.result(), accuracy_history.result()))
#clear the history
loss_history.init_variables()
accuracy_history.init_variables()
epoch += 1
i += 1
print("train complete")
# ## Testing
accuracy(model, x_test, y_test)
| eager_excution/13.simpleCNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # 01 - Estimating metrics
# Often, the data we are provided does not (yet) contain the metrics we need. In these first exercises, we are going to load in some data (similar to the data we used before) and estimate some useful metrics.
# ## Loading and understanding data
# Let's import `pandas`, load in our dataset, and `print` the columns in the dataset.
#
# (Note: the dataset is named `wk3_listings_sample.csv` becuase it is slightly different from before)
import pandas as pd
df_listings = pd.read_csv('../data/wk3_listings_sample.csv')
print(df_listings.columns)
# As you can see, there are lots of columns in this dataset. If you would like to understand the columns better, then you can look at the [data dictionary](https://docs.google.com/spreadsheets/d/1iWCNJcSutYqpULSQHlNyGInUvHg2BoUGoNRIGa6Szc4/edit#gid=982310896) of the dataset.
# ### Exercise-01: Understanding columns
# **Questions** What do the values in the `minimum_maximum_nights` column represent?
# ## Price per person
# You are asked to estimate the "price per person" for each listing in the dataset. To do this, you first need to format the `price` column to contain values you can perform calculations on (like we did last week). Let's do this below. Create a `price_$` column with the price of the listing in \$'s in `float` format, and show the `head` of the `price` and `price_$` columns to check it worked OK.
# +
# (SOLUTION)
def format_price(price):
return(float(price.replace('$','').replace(',','')))
df_listings['price_$'] = df_listings['price'].apply(format_price)
df_listings[['price','price_$']].head()
# -
# ### Exercise-02: Estimate price-per-person
# With the price values in a `float` format, we can use the `/` (division) operator to divide the values in the `price_$` column by the values in the `accommodates` column, and create a new column named `price_$/person`. Run the code shown below to do this and show the `head` of the relevant columns to check it's working OK.
df_listings['price_$/person'] = df_listings['price_$'] / df_listings['accommodates']
df_listings[['price_$','accommodates','price_$/person']].head()
# We can also sort the rows of `df_listings` by the values in column `price_$/person` to identify which listing has the highest `price_$/person` of all the listings in the dataset. Run the code below to show the neighbourhood of the listing with the highest `price_$/person` in the dataset.
df_listings.sort_values(by='price_$/person', ascending=False).head(1)['neighbourhood_cleansed']
# **Question:** What neighbourhood is it in?
# Look again in the [data dictionary](https://docs.google.com/spreadsheets/d/1iWCNJcSutYqpULSQHlNyGInUvHg2BoUGoNRIGa6Szc4/edit#gid=982310896) to understand what values in the `accommodates` column represent.
#
# **Question:** Do you think using `accomodates` is a good way to estimate the price per person for each listing? If not, why not?
#
# **Question:** Why might someone want to estimate the price per person for each listing?
# ## Forecasting income
# You are asked to forecast how much money (in \$'s) each listing is likely to receive over the next 30 days. To do this, you decide to use the `availability_30` column to calculate how many nights of the next available 30 nights are booked and then multiple this number by the `price_$` using the `*` (multiplication) operator.
#
# ### Exercise-03: Forecast 30-day income
# Complete the code below to estimate the income for each listing over the next 30 days. Look at Exercise-02, to start.
# (SOLUTION)
df_listings['estimated_income_30'] = df_listings['price_$'] * (30 - df_listings['availability_30'])
df_listings[['price_$','availability_30', 'estimated_income_30']].head()
# In the code block below, sort the values of `df_listings` by the values in `esimated_income_30` to identify the listing with highest forecasted income for the next 30 days.
# (SOLUTION)
df_listings.sort_values(by='estimated_income_30', ascending=False).head(1)['neighbourhood_cleansed']
# **Question:** What's the value of `neighbourhood_cleansed` for this listing?
#
# **Question:** Why might someone want to forecast the next 30 days income for each listing?
# ## Further work
# If you want to explore the data further, please do, and think about what other metrics you might be able to estimate from the data and how they might be used.
| solutions/01 - Calculating metrics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # mean, median, mode
#
# #### https://github.com/SelcukDE
import numpy as np
from scipy import stats
age = [20, 22, 25, 25, 27, 27, 27, 29, 30, 31, 121]
np.mean(age)
np.median(age)
stats.mode(age)
stats.mode(age)[0]
stats.mode(age)[0][0]
age = [20, 22, 25, 25, 27, 27, 27, 29, 30, 31]
np.mean(age)
np.median(age)
age_2=[18,19,17,20,21,22]
stats.mode(age_2)
type(age)
# # mean in arrays
age_new=np.array(age)
type(age_new)
np.mean(age)
age_new.mean() # array oluşturduğunuzda böyle de kullanabilirsiniz
# +
a = np.array([[6, 8, 3, 0],
[3, 2, 1, 7],
[8, 1, 8, 4],
[4, 2, 0, 2],
[4, 7, 5, 9]])
stats.mode(a)
# -
stats.mode(a, axis=None)
stats.mode(a, axis=1)
# # range, sd, variance
age = [20, 22, 25, 25, 27, 27, 27, 29, 30, 31, 121]
range = np.max(age)-np.min(age)
print(range)
var = np.var(age)
print(var)
std = np.std(age)
print(std)
age = [20, 22, 25, 25, 27, 27, 27, 29, 30, 31]
std = np.std(age)
print(std)
# +
# IQR
# -
x=[8, 10, 5, 24, 8, 3, 11, 3, 40, 7, 6, 12, 4]
q75, q25 = np.percentile(x, [75, 25])
q75
q25
sorted(x)
len(x)
q50=np.percentile(x, 50)
q50
iqr = q75-q25
iqr
from scipy.stats import iqr
iqr(x)
import matplotlib.pyplot as plt
mu, sigma = 100, 40
s = np.random.normal(mu,sigma, 100000)
plt.hist(s, 100)
plt.xlim(0,200);
mu, sigma = 100, 10
s = np.random.normal(mu,sigma, 100000)
plt.hist(s, 100)
plt.xlim(0,200);
kurtosis(s)
s_new=s*10
kurtosis(s_new)
# # additional
x=[7,9,9,10,10,10,11,12,12,14]
q75, q25 = np.percentile(x, [75, 25])
q75
q25
iqr = q75-q25
iqr
from scipy.stats import iqr
iqr(x)
| Statistics_S3_mean median mode.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Titanic - Machine Learning from Disaster
# ### Predict survival on the Titanic and get familiar with ML basics
# <br />
#
# ## Data Description
# **Survival:** 0 = No, 1 = Yes
#
# **pclass (Ticket class):** 1 = 1st, 2 = 2nd, 3 = 3rd
#
# **sex:** Sex
#
# **Age:** Age in years
#
# **sibsp:** number of siblings/spouses aboard the Titanic
#
# **parch:** number of parents/children aboard the Titanic
#
# **ticket:** Ticket number
#
# **fare:** Passenger fare
#
# **cabin:** Cabin number
#
# **embarked:** Port of Embarkation, C = Cherbourg, Q = Queenstown, S = Southampton
# <br />
# <br />
#
# ## Goal
# * Predict who survives on the test dataset
# +
# import Dependecies
# python imports
import math, time, random, datetime
# data analysis imports
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import missingno # plots graph of missing values
# %matplotlib inline
#plt.style.use('seaborn-whitegrid')
# ignore warnings
import warnings
warnings.filterwarnings('ignore')
# -
# import train and test data
train = pd.read_csv('dataset/train.csv')
test = pd.read_csv('dataset/test.csv')
# ## What's the data?
# View and understand the data
# view training data
train.head()
len(train)
# view test data
test.head()
len(test)
train.describe()
# # Data Wrangling
# ## What are the missing values?
# Identfy the rows and columns that contain missing data.
train.info()
train.isnull().sum()
missingno.matrix(train, figsize=(30,10))
# Observed tha **Age**, **Cabin** and **Embarked** columns contain missing values. <br /> The **missingno** library was used to visualise missing data.
# <br />knowing this will help us figure out what data cleaning and and preprocessing is required.
# <br />
# <br />
# <br />
#
#
# ## To perform our data analysis, let's create two new dataframes
# We'll create one for exploring discretised continuous variables (continuous variables which have been sorted into some kind of category) and another for exploring continuous variables.
df_bin = pd.DataFrame() # for discretised continuous variables
df_con = pd.DataFrame() # for continuous variables
# ## What datatypes are in the dataframe?
# As a general rule of thumb, features with a datatype of object could be considered categorical features. And those which are floats or ints (numbers) could be considered numerical features.
#
# However, as we dig deeper, we might find features which are numerical may actually be categorical.
#
# The goal for the next few steps is to figure out how best to process the data so our machine learning model can learn from it.
#
# Ideally, all the features will be encoded into a numerical value of some kind.
train.dtypes
# ## Explore each feature individualy
# We'll go through each column iteratively and see which ones to use in our first models. Some may need more preprocessing than others to get ready.
# <br />
# <br />
# <br />
#
# ### Target Feature: Survived
# Description: Whether the passenger survived or not.
#
# Key: 0 = did not survive, 1 = survived
#
# This is the variable we want our machine learning model to predict based off all the others.
| titanic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import time
import numpy as np
from tqdm import tqdm
import pandas as pd
import scipy as sp
from theano import tensor as tt
import pymc3 as pm
import matplotlib.pyplot as plt
import jax.numpy as jnp
from jax import jit
from jax.scipy.stats import norm
from run_scripts.load_data import load_traintest_hier
from conformal_bayes import conformal_Bayes_functions as cb
from conformal_bayes import Bayes_MCMC_functions as bmcmc
# %reload_ext autoreload
# %autoreload 2
# -
# # Load Data
x,y,x_test,y_test,y_plot,n,d = load_traintest_hier(1,"radon",100)
K = np.shape(np.unique(x[:,1]))[0]
print(K)
# # Conformal
beta_post = jnp.array(np.load("samples/beta_post_hier_{}.npy".format('radon')))[0]
intercept_post = jnp.array(np.load("samples/intercept_post_hier_{}.npy".format('radon')))[0]
sigma_post = jnp.array(np.load("samples/sigma_post_hier_{}.npy".format('radon')))[0]
# ### Define x_test
# +
x_test = np.zeros((2*K,2))
for k in range(K):
x_test[2*k:2*k + 2,1] = k
x_test[2*k,0] = 0
x_test[2*k+1,0]= 1
n_test = np.shape(x_test)[0]
pi_cb = np.zeros((n_test,np.shape(y_plot)[0]))
region_cb = np.zeros((n_test,np.shape(y_plot)[0]))
length_cb = np.zeros(n_test)
band_bayes = np.zeros((n_test,2))
length_bayes = np.zeros(n_test)
dy = y_plot[1]- y_plot[0]
# -
# ### Compute rank plots
# +
groups_train = np.unique(x[:,1]).astype('int')
K_train = np.size(groups_train)
n_groups = np.zeros(K_train)
for k in (range(K_train)):
ind_group = (x[:,1] == groups_train[k])
n_groups[k] =np.sum(ind_group)
alpha_k = 1.1*(1/(n_groups + 1))
print(alpha_k)
# +
#Define likelihood from posterior samples
start = time.time()
@jit
def normal_likelihood_cdf(y,x):
group = x[:,-1].astype('int32')
x_0 = x[:,0]
return norm.cdf(y,loc = beta_post[:,group] * x_0.transpose()+ intercept_post[:,group]\
,scale = sigma_post) #compute likelihood samples
#Precompute cdfs
cdf_test = normal_likelihood_cdf(y_plot.reshape(-1,1,1),x_test)
#Compute conformal regions
for i in tqdm(range(n_test)):
group_ind = x_test[i,1].astype('int')
band_bayes[i] = bmcmc.compute_bayes_band_MCMC(alpha_k[group_ind],y_plot,cdf_test[:,:,i])
length_bayes[i] = np.abs(band_bayes[i,1]- band_bayes[i,0])
end = time.time()
print('Bayes took {}'.format(end - start))
start = time.time()
@jit
def normal_loglikelihood(y,x):
group = x[:,-1].astype('int32')
x_0 = x[:,0]
return norm.logpdf(y,loc = beta_post[:,group]* x_0.transpose() + intercept_post[:,group]\
,scale = sigma_post) #compute likelihood samples
#Compute loglikelihood across groups
logp_samp_n = []
for k in (range(K_train)):
ind_group = (x[:,1] == groups_train[k])
logp_samp_n.append(normal_loglikelihood(y[ind_group],x[ind_group]))
logwjk = normal_loglikelihood(y_plot.reshape(-1,1,1),x_test)
#Compute conformal regions
for i in tqdm(range(n_test)):
group_ind = x_test[i,1].astype('int')
pi_cb[i] = cb.compute_rank_IS(logp_samp_n[group_ind],logwjk[:,:,i])
region_cb[i] = cb.compute_cb_region_IS(alpha_k[group_ind],logp_samp_n[group_ind],logwjk[:,:,i])
length_cb[i] = np.sum(region_cb[i])*dy
end = time.time()
print('Conformal took {}'.format(end-start))
# -
length_bayes_grp = np.zeros(K_train)
length_cb_grp = np.zeros(K_train)
for k in range(K_train):
length_bayes_grp[k] = np.mean(length_bayes[2*k:2*k+2])
length_cb_grp[k] = np.mean(length_cb[2*k:2*k+2])
print(np.mean(length_bayes_grp))
print(np.mean(length_cb_grp))
# ### Analyze rank plots
np.where(n_groups ==1)
# +
f = plt.figure(figsize = (12,3))
plt.subplot(1,2,1)
k = 0
i = 2*k
county = x_test[i,1].astype('int')
n_k = n_groups[county].astype('int')
plt.plot(y_plot,pi_cb[i]/(n_k+1),label = r'$x ={}$'.format(x_test[i,0].astype('int')))
county = x_test[i+1,1].astype('int')
n_k = n_groups[county].astype('int')
plt.plot(y_plot,pi_cb[i+1]/(n_k+1),label = r'$x ={}$'.format(x_test[i+1,0].astype('int')))
plt.xlabel('$y$',fontsize = 12)
plt.ylabel(r'$\pi_j(y)$')
plt.ylim(-0.05,1.1)
plt.title('County {}: $n_j = {}$'.format(county,n_k))
plt.legend()
plt.subplot(1,2,2)
k = 1
i = 2*k
county = x_test[i,1].astype('int')
n_k = n_groups[county].astype('int')
plt.plot(y_plot,pi_cb[i]/(n_k+1),label = r'$x ={}$'.format(x_test[i,0].astype('int')))
county = x_test[i+1,1].astype('int')
n_k = n_groups[county].astype('int')
plt.plot(y_plot,pi_cb[i+1]/(n_k+1),label = r'$x ={}$'.format(x_test[i+1,0].astype('int')))
plt.xlabel('$y$',fontsize = 12)
plt.ylabel(r'$\pi_j(y)$')
plt.ylim(-0.05,1.1)
plt.title('County {}: $n_j = {}$'.format(county,n_k))
f.savefig('plots/radon_big.pdf', bbox_inches='tight')
# +
f = plt.figure(figsize = (12,3))
plt.subplot(1,2,1)
k = 41
i = 2*k
county = x_test[i,1].astype('int')
n_k = n_groups[county].astype('int')
plt.plot(y_plot,pi_cb[i]/(n_k+1),label = r'$x ={}$'.format(x_test[i,0].astype('int')))
county = x_test[i+1,1].astype('int')
n_k = n_groups[county].astype('int')
plt.plot(y_plot,pi_cb[i+1]/(n_k+1),label = r'$x ={}$'.format(x_test[i+1,0].astype('int')))
plt.xlabel('$y$',fontsize = 12)
plt.ylabel(r'$\pi_j(y)$')
plt.ylim(-0.05,1.1)
plt.title('County {}: $n_j = {}$'.format(county,n_k))
plt.legend()
plt.subplot(1,2,2)
k = 49
i = 2*k
county = x_test[i,1].astype('int')
n_k = n_groups[county].astype('int')
plt.plot(y_plot,pi_cb[i]/(n_k+1),label = r'$x ={}$'.format(x_test[i,0].astype('int')))
county = x_test[i+1,1].astype('int')
n_k = n_groups[county].astype('int')
plt.plot(y_plot,pi_cb[i+1]/(n_k+1),label = r'$x ={}$'.format(x_test[i+1,0].astype('int')))
plt.xlabel('$y$',fontsize = 12)
plt.ylabel(r'$\pi_j(y)$')
plt.ylim(-0.05,1.1)
plt.title('County {}: $n_j = {}$'.format(county,n_k))
f.savefig('plots/radon_small.pdf', bbox_inches='tight')
# -
# ### LMER
# +
import statsmodels.formula.api as smf
d = {'log_radon': y, 'floor': x[:,0],'county':x[:,1]}
data = pd.DataFrame(data=d)
md = smf.mixedlm("log_radon ~ floor", data, groups=data["county"], re_formula="~floor")
# %time mdf = md.fit(method=["lbfgs"])
print(mdf.summary())
| Radon.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# <h1>Batch Transform Using R with Amazon SageMaker</h1>
#
# **Read before running this notebook:**
#
# - This sample notebook has been updated for SageMaker SDK v2.0.
# - If you are using SageMaker Notebook instances, select `R` kernel for the notebook. If you are using SageMaker Studio notebooks, you will need to create a custom R kernel for your studio domain. Follow the instructions in this blog post to create and attach a custom R kernel.
# - [Bringing your own R environment to Amazon SageMaker Studio](https://aws.amazon.com/blogs/machine-learning/bringing-your-own-r-environment-to-amazon-sagemaker-studio/)
#
# **Summary:**
#
# This sample Notebook describes how to do batch transform to make predictions for an abalone's age, which is measured by the number of rings in the shell. The notebook will use the public [abalone dataset](https://archive.ics.uci.edu/ml/datasets/abalone) originally from [UCI Machine Learning Repository](https://archive.ics.uci.edu/ml/index.php).
#
# You can find more details about SageMaker's Batch Transform here:
# - [Batch Transform](https://docs.aws.amazon.com/sagemaker/latest/dg/batch-transform.html) using a Transformer
#
# We will use `reticulate` library to interact with SageMaker:
# - [`Reticulate` library](https://rstudio.github.io/reticulate/): provides an R interface to use the [Amazon SageMaker Python SDK](https://sagemaker.readthedocs.io/en/latest/index.html) to make API calls to Amazon SageMaker. The `reticulate` package translates between R and Python objects, and Amazon SageMaker provides a serverless data science environment to train and deploy ML models at scale.
#
# Table of Contents:
# - [Reticulating the Amazon SageMaker Python SDK](#Reticulating-the-Amazon-SageMaker-Python-SDK)
# - [Creating and Accessing the Data Storage](#Creating-and-accessing-the-data-storage)
# - [Downloading and Processing the Dataset](#Downloading-and-processing-the-dataset)
# - [Preparing the Dataset for Model Training](#Preparing-the-dataset-for-model-training)
# - [Creating a SageMaker Estimator](#Creating-a-SageMaker-Estimator)
# - [Batch Transform using SageMaker Transformer](#Batch-Transform-using-SageMaker-Transformer)
# - [Download the Batch Transform Output](#Download-the-Batch-Transform-Output)
#
#
# **Note:** The first portion of this notebook focused on data ingestion and preparing the data for model training is inspired by the data preparation section outlined in the ["Using R with Amazon SageMaker"](https://github.com/awslabs/amazon-sagemaker-examples/blob/master/advanced_functionality/r_kernel/using_r_with_amazon_sagemaker.ipynb) notebook on AWS SageMaker Examples Github repository with some modifications.
# <h3>Reticulating the Amazon SageMaker Python SDK</h3>
#
# First, load the `reticulate` library and import the `sagemaker` Python module. Once the module is loaded, use the `$` notation in R instead of the `.` notation in Python to use available classes.
# Turn warnings off globally
options(warn=-1)
# Install reticulate library and import sagemaker
library(reticulate)
sagemaker <- import('sagemaker')
# <h3>Creating and Accessing the Data Storage</h3>
#
# The `Session` class provides operations for working with the following [boto3](https://boto3.amazonaws.com/v1/documentation/api/latest/index.html) resources with Amazon SageMaker:
#
# * [S3](https://boto3.readthedocs.io/en/latest/reference/services/s3.html)
# * [SageMaker](https://boto3.readthedocs.io/en/latest/reference/services/sagemaker.html)
#
# Let's create an [Amazon Simple Storage Service](https://aws.amazon.com/s3/) bucket for your data.
session <- sagemaker$Session()
bucket <- session$default_bucket()
prefix <- 'r-batch-transform'
# **Note** - The `default_bucket` function creates a unique Amazon S3 bucket with the following name:
#
# `sagemaker-<aws-region-name>-<aws account number>`
#
# Specify the IAM role's [ARN](https://docs.aws.amazon.com/general/latest/gr/aws-arns-and-namespaces.html) to allow Amazon SageMaker to access the Amazon S3 bucket. You can use the same IAM role used to create this Notebook:
role_arn <- sagemaker$get_execution_role()
# <h3>Downloading and Processing the Dataset</h3>
#
# The model uses the [abalone dataset](https://archive.ics.uci.edu/ml/datasets/abalone) originally from the [UCI Machine Learning Repository](https://archive.ics.uci.edu/ml/index.php). First, download the data and start the [exploratory data analysis](https://en.wikipedia.org/wiki/Exploratory_data_analysis). Use tidyverse packages to read, plot, and transform the data into ML format for Amazon SageMaker:
library(readr)
data_file <- 's3://sagemaker-sample-files/datasets/tabular/uci_abalone/abalone.csv'
abalone <- read_csv(file = sagemaker$s3$S3Downloader$read_file(data_file, sagemaker_session=session),
col_names = FALSE)
names(abalone) <- c('sex', 'length', 'diameter', 'height', 'whole_weight', 'shucked_weight', 'viscera_weight', 'shell_weight', 'rings')
head(abalone)
# The output above shows that `sex` is a factor data type but is currently a character data type (F is Female, M is male, and I is infant). Change `sex` to a factor and view the statistical summary of the dataset:
abalone$sex <- as.factor(abalone$sex)
summary(abalone)
# The summary above shows that the minimum value for `height` is 0.
#
# Visually explore which abalones have height equal to 0 by plotting the relationship between `rings` and `height` for each value of `sex`:
library(ggplot2)
options(repr.plot.width = 5, repr.plot.height = 4)
ggplot(abalone, aes(x = height, y = rings, color = sex)) + geom_point() + geom_jitter()
# The plot shows multiple outliers: two infant abalones with a height of 0 and a few female and male abalones with greater heights than the rest. Let's filter out the two infant abalones with a height of 0.
library(dplyr)
abalone <- abalone %>%
filter(height != 0)
# <h3>Preparing the Dataset for Model Training</h3>
#
# The model needs three datasets: one for training, testing, and validation. First, convert `sex` into a [dummy variable](https://en.wikipedia.org/wiki/Dummy_variable_(statistics)) and move the target, `rings`, to the first column. Amazon SageMaker algorithm require the target to be in the first column of the dataset.
abalone <- abalone %>%
mutate(female = as.integer(ifelse(sex == 'F', 1, 0)),
male = as.integer(ifelse(sex == 'M', 1, 0)),
infant = as.integer(ifelse(sex == 'I', 1, 0))) %>%
select(-sex)
abalone <- abalone %>%
select(rings:infant, length:shell_weight)
head(abalone)
# Next, sample 70% of the data for training the ML algorithm. Split the remaining 30% into two halves, one for testing and one for validation:
abalone_train <- abalone %>%
sample_frac(size = 0.7)
abalone <- anti_join(abalone, abalone_train)
abalone_test <- abalone %>%
sample_frac(size = 0.5)
abalone_valid <- anti_join(abalone, abalone_test)
# Upload the training and validation data to Amazon S3 so that you can train the model. First, write the training and validation datasets to the local filesystem in `.csv` format. Then, upload the two datasets to the Amazon S3 bucket into the `data` key:
# +
write_csv(abalone_train, 'abalone_train.csv', col_names = FALSE)
write_csv(abalone_valid, 'abalone_valid.csv', col_names = FALSE)
# Remove target from test
write_csv(abalone_test[-1], 'abalone_test.csv', col_names = FALSE)
# +
s3_train <- session$upload_data(path = 'abalone_train.csv',
bucket = bucket,
key_prefix = paste(prefix,'data', sep = '/'))
s3_valid <- session$upload_data(path = 'abalone_valid.csv',
bucket = bucket,
key_prefix = paste(prefix,'data', sep = '/'))
s3_test <- session$upload_data(path = 'abalone_test.csv',
bucket = bucket,
key_prefix = paste(prefix,'data', sep = '/'))
# -
# Finally, define the Amazon S3 input types for the Amazon SageMaker algorithm:
s3_train_input <- sagemaker$inputs$TrainingInput(s3_data = s3_train,
content_type = 'csv')
s3_valid_input <- sagemaker$inputs$TrainingInput(s3_data = s3_valid,
content_type = 'csv')
# <h3>Hyperparameter Tuning for the XGBoost Model</h3>
#
# Amazon SageMaker algorithms are available via a [Docker](https://www.docker.com/) container. To train an [XGBoost](https://en.wikipedia.org/wiki/Xgboost) model, specify the training containers in [Amazon Elastic Container Registry](https://aws.amazon.com/ecr/) (Amazon ECR) for the AWS Region. We will use the `latest` version of the algorithm.
container <- sagemaker$image_uris$retrieve(framework='xgboost', region= session$boto_region_name, version='latest')
cat('XGBoost Container Image URL: ', container)
# Define an Amazon SageMaker [Estimator](http://sagemaker.readthedocs.io/en/latest/estimators.html), which can train any supplied algorithm that has been containerized with Docker. When creating the Estimator, use the following arguments:
# * **image_uri** - The container image to use for training
# * **role** - The Amazon SageMaker service role
# * **train_instance_count** - The number of Amazon EC2 instances to use for training
# * **train_instance_type** - The type of Amazon EC2 instance to use for training
# * **train_volume_size** - The size in GB of the [Amazon Elastic Block Store](https://aws.amazon.com/ebs/) (Amazon EBS) volume to use for storing input data during training
# * **train_max_run** - The timeout in seconds for training
# * **input_mode** - The input mode that the algorithm supports
# * **output_path** - The Amazon S3 location for saving the training results (model artifacts and output files)
# * **output_kms_key** - The [AWS Key Management Service](https://aws.amazon.com/kms/) (AWS KMS) key for encrypting the training output
# * **base_job_name** - The prefix for the name of the training job
# * **sagemaker_session** - The Session object that manages interactions with Amazon SageMaker API
# Model artifacts and batch output
s3_output <- paste('s3:/', bucket, prefix,'output', sep = '/')
# Estimator
estimator <- sagemaker$estimator$Estimator(image_uri = container,
role = role_arn,
train_instance_count = 1L,
train_instance_type = 'ml.m5.4xlarge',
train_volume_size = 30L,
train_max_run = 3600L,
input_mode = 'File',
output_path = s3_output,
output_kms_key = NULL,
base_job_name = NULL,
sagemaker_session = NULL)
# **Note** - The equivalent to `None` in Python is `NULL` in R.
#
# Next, we Specify the [XGBoost hyperparameters](https://docs.aws.amazon.com/sagemaker/latest/dg/xgboost_hyperparameters.html) for the estimator.
#
# Once the Estimator and its hyperparamters are specified, you can train (or fit) the estimator.
# Set Hyperparameters
estimator$set_hyperparameters(eval_metric='rmse',
objective='reg:linear',
num_round=100L,
rate_drop=0.3,
tweedie_variance_power=1.4)
# +
# Create a training job name
job_name <- paste('sagemaker-r-xgboost', format(Sys.time(), '%H-%M-%S'), sep = '-')
# Define the data channels for train and validation datasets
input_data <- list('train' = s3_train_input,
'validation' = s3_valid_input)
# train the estimator
estimator$fit(inputs = input_data, job_name = job_name)
# -
# <hr>
#
# <h3> Batch Transform using SageMaker Transformer </h3>
# For more details on SageMaker Batch Transform, you can visit this example notebook on [Amazon SageMaker Batch Transform](https://github.com/awslabs/amazon-sagemaker-examples/blob/master/sagemaker_batch_transform/introduction_to_batch_transform/batch_transform_pca_dbscan_movie_clusters.ipynb).
#
# In many situations, using a deployed model for making inference is not the best option, especially when the goal is not to make online real-time inference but to generate predictions from a trained model on a large dataset. In these situations, using Batch Transform may be more efficient and appropriate.
#
# This section of the notebook explains how to set up the Batch Transform Job and generate predictions.
#
# To do this, we need to identify the batch input data path in S3 and specify where generated predictions will be stored in S3.
# Define S3 path for Test data
s3_test_url <- paste('s3:/', bucket, prefix, 'data','abalone_test.csv', sep = '/')
# Then we create a `Transformer`. [Transformers](https://sagemaker.readthedocs.io/en/stable/transformer.html#transformer) take multiple paramters, including the following. For more details and the complete list visit the [documentation page](https://sagemaker.readthedocs.io/en/stable/transformer.html#transformer).
#
# - **model_name** (str) – Name of the SageMaker model being used for the transform job.
# - **instance_count** (int) – Number of EC2 instances to use.
# - **instance_type** (str) – Type of EC2 instance to use, for example, ‘ml.c4.xlarge’.
#
# - **output_path** (str) – S3 location for saving the transform result. If not specified, results are stored to a default bucket.
#
# - **base_transform_job_name** (str) – Prefix for the transform job when the transform() method launches. If not specified, a default prefix will be generated based on the training image name that was used to train the model associated with the transform job.
#
# - **sagemaker_session** (sagemaker.session.Session) – Session object which manages interactions with Amazon SageMaker APIs and any other AWS services needed. If not specified, the estimator creates one using the default AWS configuration chain.
#
# Once we create a `Transformer` we can transform the batch input.
# Define a transformer
transformer <- estimator$transformer(instance_count=1L,
instance_type='ml.m4.xlarge',
output_path = s3_output)
# Do the batch transform
transformer$transform(s3_test_url,
wait = TRUE)
# <hr>
# <h3> Download the Batch Transform Output </h3>
# Download the file from S3 using S3Downloader to local SageMaker instance 'batch_output' folder
sagemaker$s3$S3Downloader$download(paste(s3_output,"abalone_test.csv.out",sep = '/'),
"batch_output")
# Read the batch csv from sagemaker local files
library(readr)
predictions <- read_csv(file = 'batch_output/abalone_test.csv.out', col_names = 'predicted_rings')
head(predictions)
# Column-bind the predicted rings to the test data:
# Concatenate predictions and test for comparison
abalone_predictions <- cbind(predicted_rings = predictions,
abalone_test)
# Convert predictions to Integer
abalone_predictions$predicted_rings = as.integer(abalone_predictions$predicted_rings);
head(abalone_predictions)
# Define a function to calculate RMSE
rmse <- function(m, o){
sqrt(mean((m - o)^2))
}
# Calucalte RMSE
abalone_rmse <- rmse(abalone_predictions$rings, abalone_predictions$predicted_rings)
cat('RMSE for Batch Transform: ', round(abalone_rmse, digits = 2))
| r_examples/r_batch_transform/r_xgboost_batch_transform.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # QuTiP example: Bloch-Redfield Master Equation
# <NAME> and <NAME>
#
# For more information about QuTiP see [http://qutip.org](http://qutip.org)
import numpy as np
# %pylab inline
from qutip import *
# ## Single qubit dynamics
def qubit_integrate(w, theta, gamma1, gamma2, psi0, tlist):
# Hamiltonian
sx = sigmax()
sy = sigmay()
sz = sigmaz()
sm = sigmam()
H = w * (np.cos(theta) * sz + np.sin(theta) * sx)
# Lindblad master equation
c_op_list = []
n_th = 0.0 # zero temperature
rate = gamma1 * (n_th + 1)
if rate > 0.0:
c_op_list.append(np.sqrt(rate) * sm)
rate = gamma1 * n_th
if rate > 0.0:
c_op_list.append(np.sqrt(rate) * sm.dag())
lme_results = mesolve(H, psi0, tlist, c_op_list, [sx, sy, sz]).expect
# Bloch-Redfield tensor
#ohmic_spectrum = lambda w: gamma1 * w / (2*pi)**2 * (w > 0.0)
def ohmic_spectrum(w):
if w == 0.0:
# dephasing inducing noise
return gamma1/2
else:
# relaxation inducing noise
return gamma1/2 * w / (2*np.pi) * (w > 0.0)
brme_results = brmesolve(H, psi0, tlist, [[sx, ohmic_spectrum]], [sx, sy, sz]).expect
# alternative:
#R, ekets = bloch_redfield_tensor(H, [sx], [ohmic_spectrum])
#brme_results = bloch_redfield_solve(R, ekets, psi0, tlist, [sx, sy, sz])
return lme_results, brme_results
w = 1.0 * 2 * np.pi # qubit angular frequency
theta = 0.05 * np.pi # qubit angle from sigma_z axis (toward sigma_x axis)
gamma1 = 0.5 # qubit relaxation rate
gamma2 = 0.0 # qubit dephasing rate
# initial state
a = 0.8
psi0 = (a* basis(2,0) + (1-a)*basis(2,1))/(np.sqrt(a**2 + (1-a)**2))
tlist = np.linspace(0,15,5000)
lme_results, brme_results = qubit_integrate(w, theta, gamma1, gamma2, psi0, tlist)
# +
fig = figure(figsize=(12,12))
ax = fig.add_subplot(2,2,1)
title('Lindblad master equation')
ax.plot(tlist, lme_results[0], 'r')
ax.plot(tlist, lme_results[1], 'g')
ax.plot(tlist, lme_results[2], 'b')
ax.legend(("sx", "sy", "sz"))
ax = fig.add_subplot(2,2,2)
title('Bloch-Redfield master equation')
ax.plot(tlist, brme_results[0], 'r')
ax.plot(tlist, brme_results[1], 'g')
ax.plot(tlist, brme_results[2], 'b')
ax.legend(("sx", "sy", "sz"))
sphere=Bloch(axes=fig.add_subplot(2,2,3, projection='3d'))
sphere.add_points([lme_results[0],lme_results[1],lme_results[2]], meth='l')
sphere.vector_color = ['r']
sphere.add_vectors([sin(theta),0,cos(theta)])
sphere.make_sphere()
sphere=Bloch(axes=fig.add_subplot(2,2,4, projection='3d'))
sphere.add_points([brme_results[0],brme_results[1],brme_results[2]], meth='l')
sphere.vector_color = ['r']
sphere.add_vectors([sin(theta),0,cos(theta)])
sphere.make_sphere()
# -
# ## Coupled qubits
def qubit_integrate(w, theta, g, gamma1, gamma2, psi0, tlist):
#
# Hamiltonian
#
sx1 = tensor(sigmax(),qeye(2))
sy1 = tensor(sigmay(),qeye(2))
sz1 = tensor(sigmaz(),qeye(2))
sm1 = tensor(sigmam(),qeye(2))
sx2 = tensor(qeye(2),sigmax())
sy2 = tensor(qeye(2),sigmay())
sz2 = tensor(qeye(2),sigmaz())
sm2 = tensor(qeye(2),sigmam())
H = w[0] * (np.cos(theta[0]) * sz1 + np.sin(theta[0]) * sx1) # qubit 1
H += w[1] * (np.cos(theta[1]) * sz2 + np.sin(theta[1]) * sx2) # qubit 2
H += g * sx1 * sx2 # interaction
#
# Lindblad master equation
#
c_op_list = []
n_th = 0.0 # zero temperature
rate = gamma1[0] * (n_th + 1)
if rate > 0.0: c_op_list.append(np.sqrt(rate) * sm1)
rate = gamma1[1] * (n_th + 1)
if rate > 0.0: c_op_list.append(np.sqrt(rate) * sm2)
lme_results = mesolve(H, psi0, tlist, c_op_list, [sx1, sy1, sz1]).expect
#
# Bloch-Redfield tensor
#
def ohmic_spectrum1(w):
if w == 0.0:
# dephasing inducing noise
return gamma1[0]/2
else:
# relaxation inducing noise
return gamma1[0] * w / (2*np.pi) * (w > 0.0)
def ohmic_spectrum2(w):
if w == 0.0:
# dephasing inducing noise
return gamma1[1]/2
else:
# relaxation inducing noise
return gamma1[1] * w / (2*np.pi) * (w > 0.0)
brme_results = brmesolve(H, psi0, tlist, [[sx1,ohmic_spectrum1], [sx2,ohmic_spectrum2]], [sx1, sy1, sz1]).expect
# alternative:
#R, ekets = bloch_redfield_tensor(H, [sx1, sx2], [ohmic_spectrum1, ohmic_spectrum2])
#brme_results = brmesolve(R, ekets, psi0, tlist, [sx1, sy1, sz1])
return lme_results, brme_results
# +
w = array([1.0, 1.0]) * 2 * np.pi # qubit angular frequency
theta = array([0.15, 0.45]) * 2 * np.pi # qubit angle from sigma_z axis (toward sigma_x axis)
gamma1 = [0.25, 0.35] # qubit relaxation rate
gamma2 = [0.0, 0.0] # qubit dephasing rate
g = 0.1 * 2 * np.pi
# initial state
a = 0.8
psi1 = (a*basis(2,0) + (1-a)*basis(2,1))/(np.sqrt(a**2 + (1-a)**2))
psi2 = ((1-a)*basis(2,0) + a*basis(2,1))/(np.sqrt(a**2 + (1-a)**2))
psi0 = tensor(psi1, psi2)
tlist = np.linspace(0,15,5000)
# -
lme_results, brme_results = qubit_integrate(w, theta, g, gamma1, gamma2, psi0, tlist)
# +
fig = figure(figsize=(12,12))
ax = fig.add_subplot(2,2,1)
title('Lindblad master equation')
ax.plot(tlist, lme_results[0], 'r')
ax.plot(tlist, lme_results[1], 'g')
ax.plot(tlist, lme_results[2], 'b')
ax.legend(("sx", "sy", "sz"))
ax = fig.add_subplot(2,2,2)
title('Bloch-Redfield master equation')
ax.plot(tlist, brme_results[0], 'r')
ax.plot(tlist, brme_results[1], 'g')
ax.plot(tlist, brme_results[2], 'b')
ax.legend(("sx", "sy", "sz"))
sphere=Bloch(axes=fig.add_subplot(2,2,3, projection='3d'))
sphere.add_points([lme_results[0],lme_results[1],lme_results[2]], meth='l')
sphere.vector_color = ['r']
sphere.add_vectors([sin(theta[0]),0,cos(theta[0])])
sphere.make_sphere()
sphere=Bloch(axes=fig.add_subplot(2,2,4, projection='3d'))
sphere.add_points([brme_results[0],brme_results[1],brme_results[2]], meth='l')
sphere.vector_color = ['r']
sphere.add_vectors([sin(theta[0]),0,cos(theta[0])])
sphere.make_sphere()
# -
# ## Versions
# +
from qutip.ipynbtools import version_table
version_table()
| qutip-notebooks-master/examples/bloch-redfield.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/JSJeong-me/OpenCV_Practitioner_Guide_2/blob/main/count-1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="HWp8tHdPtVNH"
# low_red = [0, 142, 79]
# high_red = [255, 255, 255]
# + id="LTPRYuP2tJPZ"
import cv2
from google.colab.patches import cv2_imshow
# + id="KjyrDl3stmzY"
src = cv2.imread('./antivirus.jpg', cv2.COLOR_BGR2RGB)
# + colab={"base_uri": "https://localhost:8080/", "height": 467} id="tz2OcLAotxNM" outputId="5d1e5e81-02d3-4838-e96f-8adeb8bcc472"
cv2_imshow(src)
# + id="SLNtpYw0tzcw"
hsv_src = cv2.cvtColor(src, cv2.COLOR_RGB2HSV)
# + id="xiD7QYtBuHHq"
import numpy as np
# + id="LqN82gAhuE28"
low_red = np.array([0, 142, 79])
high_red = np.array([255, 255, 255])
# + id="F1HbDaOFuRDZ"
mask = cv2.inRange(hsv_src,low_red, high_red)
# + id="PE0cNr8mua8U"
result_src = cv2.bitwise_and(src, src, mask = mask)
# + colab={"base_uri": "https://localhost:8080/", "height": 467} id="unQPSuQpulTI" outputId="a7917cfb-2f59-46af-d284-3636a8255fd4"
cv2_imshow(result_src)
| count-1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import seaborn as sns
import matplotlib.pyplot as plt
import pandas as pd
df = pd.read_csv('C:\\Users\\<NAME>\\Documents\\Knime\\Data\\EDA_binned.csv')
df.head()
y = df["Churn"].value_counts()
sns.barplot(y.index, y.values)
df.groupby(["City", "Churn"]).size().unstack().plot(kind='bar', stacked=True, figsize=(30,10))
df_pivot = df.pivot_table(values='Phone', index=['City'], columns='Churn', aggfunc='count', margins=1)
df_ratio = df_pivot.iloc[:,1]/df_pivot.iloc[:,2]*100
#print(df_ratio)
df_ratio.plot(kind='bar')
df.groupby(["VMail Message", "Churn"]).size().unstack().plot(kind='bar', stacked=True, figsize=(5,5))
df.groupby(["Day Mins", "Churn"]).size().unstack().plot(kind='bar', stacked=True, figsize=(5,5))
df.groupby(["Eve Mins", "Churn"]).size().unstack().plot(kind='bar', stacked=True, figsize=(5,5))
df.groupby(["Night Mins", "Churn"]).size().unstack().plot(kind='bar', stacked=True, figsize=(5,5))
df_pivot = df.pivot_table(values='Phone', index=['VMail Message'], columns='Churn', aggfunc='count', margins=1)
df_ratio = df_pivot.iloc[:,1]/df_pivot.iloc[:,2]*100
#print(df_ratio)
df_ratio.plot(kind='bar')
df_pivot = df.pivot_table(values='Phone', index=['Day Mins'], columns='Churn', aggfunc='count', margins=1)
df_ratio = df_pivot.iloc[:,1]/df_pivot.iloc[:,2]*100
#print(df_ratio)
df_ratio.plot(kind='bar')
df_pivot = df.pivot_table(values='Phone', index=['Eve Mins'], columns='Churn', aggfunc='count', margins=1)
df_ratio = df_pivot.iloc[:,1]/df_pivot.iloc[:,2]*100
df_ratio.plot(kind='bar')
df_pivot = df.pivot_table(values='Phone', index=['Night Mins'], columns='Churn', aggfunc='count', margins=1)
df_ratio = df_pivot.iloc[:,1]/df_pivot.iloc[:,2]*100
df_ratio.plot(kind='bar')
df_pivot = df.pivot_table(values='Phone', index=['Intl Mins'], columns='Churn', aggfunc='count', margins=1)
df_ratio = df_pivot.iloc[:,1]/df_pivot.iloc[:,2]*100
df_ratio.plot(kind='bar')
# +
#df.groupby(["CustServ Calls", "Churn"]).size().unstack().plot(kind='bar', stacked=True, figsize=(5,5))
#df.groupby(["VMail Plan", "Churn"]).size().unstack().plot(kind='bar', stacked=True, figsize=(5,5))
#df.groupby(["Int'l Plan", "Churn"]).size().unstack().plot(kind='bar', stacked=True, figsize=(5,5))
#df.groupby(["Account Length", "Churn"]).size().unstack().plot(kind='bar', stacked=True, figsize=(5,5))
# -
df_pivot = df.pivot_table(values='Phone', index=['CustServ Calls'], columns='Churn', aggfunc='count', margins=1)
df_ratio = df_pivot.iloc[:,1]/df_pivot.iloc[:,2]*100
df_ratio.plot(kind='bar')
df_pivot = df.pivot_table(values='Phone', index=['VMail Plan'], columns='Churn', aggfunc='count', margins=1)
df_ratio = df_pivot.iloc[:,1]/df_pivot.iloc[:,2]*100
df_ratio.plot(kind='bar')
df_pivot = df.pivot_table(values='Phone', index=['Int\'l Plan'], columns='Churn', aggfunc='count', margins=1)
df_ratio = df_pivot.iloc[:,1]/df_pivot.iloc[:,2]*100
df_ratio.plot(kind='bar')
df_pivot = df.pivot_table(values='Phone', index=['Account Length'], columns='Churn', aggfunc='count', margins=1)
df_ratio = df_pivot.iloc[:,1]/df_pivot.iloc[:,2]*100
df_ratio.plot(kind='bar')
| Churn-prediction/jupyter-notebooks/EDA_binned_ChurnRatios.ipynb |