code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Input Distributions
#
# In this chapter, the default input distributions are presented.
# These are automatically seeded when a new database is created.
# They can be simply overwritten with your use-case specific assumptions.
# First, the required libraries are imported.
import datetime
import matplotlib.pyplot as plt
import conflowgen
# Next, an
# [in-memory SQLite database](https://www.sqlite.org/inmemorydb.html)
# is opened.
# This is a fresh database without any content.
# While creating the database, it is automatically seeded with the default values.
database_chooser = conflowgen.DatabaseChooser()
database_chooser.create_new_sqlite_database(":memory:")
# If that was too fast, you can switch on logging to have a look behind the scenes.
# The logger is registered under the name of the module.
import sys
import logging
logger = logging.getLogger("conflowgen")
logger.setLevel(logging.DEBUG)
handler = logging.StreamHandler(stream=sys.stdout)
handler.setLevel(logging.DEBUG)
logger.addHandler(handler)
# Now you can see the same output like in the demo scripts.
# + tags=[]
database_chooser.create_new_sqlite_database(":memory:")
# -
# Once the database is set up, we can use the different distribution managers to have a look at the distributions which were automatically seeded.
# ## Container Length Distribution
#
# For each container length, the container length distribution determines its frequency among the containers.
# The numbers have been determined by the following reasoning:
# + raw_mimetype="text/restructuredtext" tags=[] active=""
# .. note::
# .. autodata:: conflowgen.domain_models.distribution_seeders.container_length_distribution_seeder.DEFAULT_CONTAINER_LENGTH_FREQUENCIES
# -
# The container length distribution is obtained with the following lines of code:
# +
container_length_manager = conflowgen.ContainerLengthDistributionManager()
length_distribution = container_length_manager.get_container_length_distribution()
length_distribution
# + raw_mimetype="text/restructuredtext" active=""
# We can see that all the keys are enum values of the enum
# :class:`.ContainerLength`.
# For a nicer visualization, we can circumvent that by converting the enum values into strings.
# +
length_distribution_with_key_as_str = {
str(key): value
for (key, value) in length_distribution.items()
}
length_distribution_with_key_as_str
# -
# Now we can also plot the same information as a pie chart.
# +
length_distribution_with_key_as_str_without_zeros = {
key: value
for (key, value) in length_distribution_with_key_as_str.items()
if value > 0
}
plt.title("Frequency of container lengths")
plt.pie(
list(length_distribution_with_key_as_str_without_zeros.values()),
labels=list(length_distribution_with_key_as_str_without_zeros.keys())
)
plt.gcf().set_size_inches(5, 5)
plt.show()
# + raw_mimetype="text/restructuredtext" active=""
# More information on setting and getting the distribution can be found at
# :class:`.ContainerLengthDistributionManager`.
# -
# ## Container Weight Distribution
#
# The container weight of each container is drawn from a distribution.
# For each container length, a different weight distribution can be provided.
# + raw_mimetype="text/restructuredtext" active=""
# .. note::
# .. autodata:: conflowgen.domain_models.distribution_seeders.container_weight_distribution_seeder.DEFAULT_CONTAINER_WEIGHT_DISTRIBUTION
# +
container_weight_distribution_manager = conflowgen.ContainerWeightDistributionManager()
weight_distribution = container_weight_distribution_manager.get_container_weight_distribution()
weight_distribution
# -
# The container weight distributions can only be overwritten all at once.
# The values are automatically normalized by default.
container_weight_distribution_manager.set_container_weight_distribution(
{
conflowgen.ContainerLength.twenty_feet: {
10: 20,
20: 50,
30: 30
},
conflowgen.ContainerLength.forty_feet: {
10: 15,
20: 50,
30: 35
},
conflowgen.ContainerLength.forty_five_feet: {
10: 10,
20: 5,
30: 85
},
conflowgen.ContainerLength.other: {
10: 1,
20: 1,
30: 1
}
}
)
# From now on, ConFlowGen uses the new container weight distribution:
container_weight_distribution_manager.get_container_weight_distribution()
# + raw_mimetype="text/restructuredtext" active=""
# More information on setting and getting the distribution can be found at
# :class:`.ContainerWeightDistributionManager`.
# -
# ## Default Values
#
# In addition to the input distributions, also some default values are defined.
# All of them are currently some kind of minimum or maximum value.
# Thus, they directly influence other distributions.
# + raw_mimetype="text/restructuredtext" tags=[] active=""
# .. note::
# .. autodata:: conflowgen.domain_models.seeders.DEFAULT_MAXIMUM_DWELL_TIME_OF_IMPORT_CONTAINERS_IN_HOURS
# .. autodata:: conflowgen.domain_models.seeders.DEFAULT_MINIMUM_DWELL_TIME_OF_IMPORT_CONTAINERS_IN_HOURS
# .. autodata:: conflowgen.domain_models.seeders.DEFAULT_MAXIMUM_DWELL_TIME_OF_EXPORT_CONTAINERS_IN_HOURS
# .. autodata:: conflowgen.domain_models.seeders.DEFAULT_MINIMUM_DWELL_TIME_OF_EXPORT_CONTAINERS_IN_HOURS
# .. autodata:: conflowgen.domain_models.seeders.DEFAULT_MAXIMUM_DWELL_TIME_OF_TRANSSHIPMENT_CONTAINERS_IN_HOURS
# .. autodata:: conflowgen.domain_models.seeders.DEFAULT_MINIMUM_DWELL_TIME_OF_TRANSSHIPMENT_CONTAINERS_IN_HOURS
# .. autodata:: conflowgen.domain_models.seeders.DEFAULT_TRANSPORTATION_BUFFER
#
# The default values can be overwritten with the help of :meth:`.ContainerFlowGenerationManager.set_properties`.
# -
container_flow_generation_manager = conflowgen.ContainerFlowGenerationManager()
container_flow_generation_manager.get_properties()
# All default values are optional.
# They are only overwritten if provided.
# The parameters `start_date` and `end_date` are obligatory though.
container_flow_generation_manager.set_properties(
start_date=datetime.date(2021, 1, 15),
end_date=datetime.date(2021, 1, 31),
maximum_dwell_time_of_export_containers_in_hours=10*24
)
container_flow_generation_manager.get_properties()
# + raw_mimetype="text/restructuredtext" active=""
# More information on setting and getting the values can be found at
# :class:`.ContainerFlowGenerationManager`.
| docs/notebooks/input_distributions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Fred's Scratchpad
# +
import sys
import os
# Function that configures the notebook kernel by putting the lib directory
# onto the library path and changing the working directory to the top-level
# project dir. Idempotent.
def setup_kernel():
# Move to project root if we're not already there
if os.getcwd().endswith("notebooks"):
os.chdir("..")
# TODO: Verify that we're actually at the project root.
# Add the lib dir to the Python path if it's not already there.
lib_dir = os.getcwd() + "/lib"
if lib_dir not in sys.path:
sys.path.append(lib_dir)
setup_kernel()
# -
import reefer.simulator.domain.reefer_simulator as sim
data_gen = sim.ReeferSimulator()
df = data_gen.generateCo2()
df.head()
# +
import matplotlib.pyplot as plt
df[['Temperature(celsius)', 'Target_Temperature(celsius)']].plot()
# -
df.Power.plot()
df.PowerConsumption.plot()
df[["O2", "CO2"]].plot()
# +
df[["CO2", "Maintenance_Required"]].iloc[:100].plot(figsize=(20, 6))
# -
df.Defrost_Cycle.iloc[:20].plot()
from pandas.plotting import autocorrelation_plot
autocorrelation_plot(df.CO2)
| pipeline-samples/icp4d-demo/etl-source-code/notebooks/scratchpad.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction to command line
# Identify and open your command line interface.
#
# - Windows: we will choose to use **Git bash** (which was installed on your machine when you installed Git).
# - nix (another way to describe Mac OS and/or Linux machines): we will use the system **terminal**.
# # Finding your computer's name.
#
# Let us first let's find out the name of your computer by running:
#
# ```shell
# $ whoami
# ```
# # Finding your current location
#
# Now let's find out which directory (folder) we are currently in:
#
# ```shell
# $ pwd
# ```
#
# This stands for "present working directory"
#
# Type the command in and press enter. It should list where you are currently located in your command line interface.
# # Seeing what is in your current location
#
# To view the contents of the current directory:
#
#
# ```shell
# $ ls
# ```
#
# This stands for "list"
#
# Type the command in and press enter. You should see a list of the various files and directory in your current directory. Open your current directory in a graphical user interface and compare.
# # Moving to another location
#
# If you want to enter a directory that is in your current directory type:
#
# ```shell
# $ cd <directory>
# ```
#
# Try moving to your Desktop. It should be something like:
#
# ```shell
# $ cd Desktop
# ```
# # Creating a directory
#
# To create a directory:
#
# ```shell
# $ mkdir <directory_name>
# ```
#
# Experiment with creating a directory for this workshop:
#
# ```shell
# $ mkdir rsd-workshop
# ```
#
# If your directory structure looked like this:
#
# ```
# |--- home/
# |--- Desktop/
# |--- research
# |--- photos
# ```
# It will now look something like:
#
# ```
# |--- home/
# |--- Desktop/
# |--- research
# |--- photos
# |--- rsd-workshop
# ```
# As an exercise move into the directory we just created:
#
# ```shell
# $ cd rsd-workshop
# ```
#
# and create two further directories:
#
# ```
# |--- rsd-workshop
# |--- src
# |--- test
# ```
#
# If you now wanted to go back to the "parent" directory:
#
# ```shell
# $ cd ..
# ```
#
# Where `..` is short hand for a previous directory.
#
# Experiment with these, in combination with the command to find your current location as well as the command to list the contents of your directory.
# # Creating a file
#
# To create a directory:
#
# ```shell
# $ touch <file_name>
# ```
#
# Experiment with creating a file named `addition.py` in the directory `rsd-workshop`.
#
#
# ```shell
# $ touch addition.py
# ```
#
# If you type `ls` you will see that the file has been created.
# # Copying files
#
# To copy a file:
#
# ```shell
# $ cp <file> <new_file_directory_and_name>
# ```
# Experiment with copying any file.
# # Moving/renaming files
#
# To move a file:
#
# ```shell
# $ mv <file> <new_file_directory_and_name>
# ```
# Experiment with moving any file. Note that if you want to rename a file you can do this by passing the new name in the same directory.
#
# WARNING When using the command line interface you will not be prompted for confirmation if move/mv were to overwrite another file. Be careful.
# # Deleting files
#
# To delete a file:
#
# ```shell
# $ rm <file>
# ```
# # Copying and removing directories
#
# To copy a directory:
#
#
# ```shell
# $ cp -r <dir> <target>
# ```
#
# To remove a directory:
#
# ```shell
# $ rm -r <dir>
# ```
#
| Day I - Part I - Introduction to command line.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: qnet-dev
# language: python
# name: qnet-dev
# ---
# +
## generate pole separation across years
#creed2_/GSS/Qnets
import os
import matplotlib.pyplot as plt
import collections
import seaborn as sns
import pandas as pd
import numpy as np
from cognet.cognet import cognet as cg
from cognet.dataFormatter import dataFormatter
from cognet.model import model
# get years
features_by_year = pd.read_csv('examples_data/features_by_year_GSS.csv',
keep_default_na=True,
index_col=False)
POLEFILE='examples_data/polar_vectors.csv'
qnets_by_year = {x[-11:-7]: x for x in os.listdir('../../creed2_/GSS/Qnets/')}
data_by_year = {x[4:8]: x for x in os.listdir('../../creed2_/GSS/data/processed_data')}
polarDist_by_year = {}
# -
for year, qnet in qnets_by_year.items():
qnet = '../../creed2_/GSS/Qnets/' + qnet
data_obj=dataFormatter('../../creed2_/GSS/data/processed_data/'+data_by_year[year])
model_obj = model()
model_obj.load(qnet)
cognet_obj = cg()
cognet_obj.load_from_model(model_obj, data_obj, 'all')
# produce stats on how many column names actually match
stats = cognet_obj.set_poles(POLEFILE,"R","L",steps=120)
# compute polar distance matrix
dmatrix = cognet_obj.polar_separation(nsteps=0)
polarDist_by_year[year]=dmatrix[0][1]
polarDist_by_year
# # Change in Polar Separation over Time
# + jupyter={"source_hidden": true} tags=[]
sorted_dist = collections.OrderedDict(sorted(polarDist_by_year.items()))
df = pd.DataFrame.from_dict(sorted_dist, orient='index')
df.reset_index(level=0, inplace=True)
df.columns = ['year', 'dist']
sns.set(style='darkgrid', rc={'figure.figsize':(20, 8)})
sns.lineplot(x='year', y='dist', data=df)
# year = list(sorted_dist.keys())
# dist = list(sorted_dist.values())
# plt.plot(year, dist)
# plt.show
# -
| examples/Yearly_polarseparation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (mciso)
# language: python
# name: mciso
# ---
# + [markdown] id="11c70480"
# # Monte Carlo Stocking Optimization
# + id="68798b6f" jupyter={"source_hidden": true} tags=[]
import itertools as it
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import pyomo.environ as pyo
from mciso import model, utils, visualize
plt.style.use("seaborn")
RNG = np.random.default_rng()
# + executionInfo={"elapsed": 805, "status": "ok", "timestamp": 1627823349127, "user": {"displayName": "Lance", "photoUrl": "", "userId": "07981726938149847530"}, "user_tz": -480} id="be58c790" jupyter={"source_hidden": true} tags=[]
sheet_id = "1mPcE2lKxwxgNtohLG5Oj57YzsNV2rL0czuWTUPxHPg8"
sheets = ["Products", "Model_Spec"]
base_url = "https://docs.google.com/spreadsheets/d/{sheet_id}/gviz/tq?tqx=out:csv&sheet={sheet_name}"
prod_spec = pd.read_csv(base_url.format(sheet_id=sheet_id, sheet_name="Products")).set_index(["Product"])
model_spec = pd.read_csv(base_url.format(sheet_id=sheet_id, sheet_name="Model_Spec")).set_index(["parameter"])
prod_df = prod_spec.filter(regex="^(?!cust_)")
cust_df = prod_spec.filter(regex="^cust_").rename(
columns=lambda x: x.replace("cust_", "")
)
idx_labels = {
"i": [f"month_{i+1}" for i in range(model_spec.loc["no of months", "value"])],
"k": prod_df.index.tolist()
}
# + [markdown] id="a0b5dff8"
# ## Modeling Demand
# + [markdown] id="aa88cc5d-e0cc-41ee-a61d-0faf0a60d0b7"
# Monthly demand for each customer is estimated as follows:
# + colab={"base_uri": "https://localhost:8080/", "height": 162} executionInfo={"elapsed": 325, "status": "ok", "timestamp": 1627823358879, "user": {"displayName": "Lance", "photoUrl": "", "userId": "07981726938149847530"}, "user_tz": -480} id="1dae856c-565d-47bb-9198-eab144e30d56" jupyter={"source_hidden": true} outputId="729806f5-d884-4007-ac35-80351f8c86aa" tags=[]
cust_df.style.background_gradient(axis=None)
# + [markdown] id="333cbdfa-cfe0-4eba-8794-437dad71c8fe"
# However, in reality, there will be *some* level of uncertainty.
# Hence, demand is modelled as a [Poisson distribution](https://en.wikipedia.org/wiki/Poisson_distribution)
# + id="2ac353bc" jupyter={"source_hidden": true} outputId="81f2a30e-a37a-4608-b2c2-79f7df2e553f" tags=[]
n_samples = 100
fig, ax = plt.subplots(
*cust_df.shape, figsize=(15, 8), sharex=True, sharey=True
)
fig.suptitle("Monthly Demand Probability Distribution\n(client vs. product)")
for i, row in enumerate(cust_df.index):
for j, col in enumerate(cust_df.columns):
ax_i = ax[i, j]
if i == 0:
ax_i.set_title(col)
if j == 0:
ax_i.set_ylabel(row)
ax_i.hist(RNG.poisson(lam=cust_df.loc[row, col], size=n_samples), alpha=0.8)
fig.tight_layout()
# + [markdown] id="f0a05d8f"
# ## Model Parameters
# + [markdown] id="15a99cfa-97ec-4779-8d13-88c6182d8c56"
# We consider the following parameters:
# * working capital: maximum inventory value (i.e. capital tied in inventory)
# * no of months
# * no of scenarios: # of scenarios we consider to draw from random distribution
#
# For each product, the parameters are as follows:
# * Unit Price: buy price for each unit of product
# * Sell Price: sell price for each unit of product
# * MOQ: minimum order quantity
# * Min Stock: minimum stock
# * Storage Cost: can be thought of as forgone interest
# * Out of Stock Cost: can be thought of as "reputational damage"
#
# + id="b7f0fbd5" jupyter={"source_hidden": true} outputId="4d33075e-5260-4e9a-84ed-2f2b08a0068f" tags=[]
display(model_spec.style.format("{:,.0f}"))
display(
prod_df.drop(columns=["BI", "L"])
.rename(
columns={
"UP": "Unit Price",
"SP": "Sell Price",
"LB": "Min Stock",
"W": "Storage Cost",
"O": "Out of Stock Cost",
}
)
.style.format("{:,.2f}")
)
# + id="7d83dc21" jupyter={"source_hidden": true} tags=[]
# Generate demand scenarios
# D[n_month, n_scenario, n_sku]
D = RNG.poisson(
lam=cust_df.values,
size=[
model_spec.loc["no of months", "value"],
model_spec.loc["no of scenarios", "value"],
]
+ list(cust_df.shape),
).sum(axis=-1)
# + [markdown] id="98d9c051-6431-4b20-a3dc-793e086d3efa"
# Demand scenarios generated are show below:
# * Each gray line represents a scenario
# * The blue line represents the "average" of scenarios
# + id="c0568f68" jupyter={"source_hidden": true} outputId="4b71ad50-dc29-4533-a1ba-ad6e2c80a256" tags=[]
fig, ax = plt.subplots(D.shape[-1], 1, figsize=(8, D.shape[-1] * 2), sharex=True)
visualize.scenarios_by_product(D, idx_labels["i"], prod_df.index, ax=ax)
fig.suptitle("Demand Scenarios by Product across time")
fig.tight_layout()
# + [markdown] id="2c1cef1f"
# ## Model Optimization
# + id="dd7e0002" jupyter={"source_hidden": true} outputId="554e8db4-7417-430e-c331-b2ab49212a31" tags=[]
data = {
None: {
"n_months": {None: D.shape[0]},
"n_scenarios": {None: D.shape[1]},
"n_skus": {None: D.shape[2]},
"D": utils.array_to_dok(D),
"BI": utils.array_to_dok(prod_df["BI"].values),
"UP": utils.array_to_dok(prod_df["UP"].values),
"SP": utils.array_to_dok(prod_df["SP"].values),
"MOQ": utils.array_to_dok(prod_df["MOQ"].values),
"LB": utils.array_to_dok(prod_df["LB"].values),
"W": utils.array_to_dok(prod_df["W"].values),
"O": utils.array_to_dok(prod_df["O"].values),
"L": utils.array_to_dok(prod_df["L"].values),
"UB": {None: model_spec.loc["working capital", "value"]},
},
}
abs_model = model.create_model()
model_ins = abs_model.create_instance(data)
opt = pyo.SolverFactory("cbc")
print("Solving model...")
res = opt.solve(model_ins)
print("Done!")
# + [markdown] id="6653a893-093b-46dc-b227-e5f5c8a4af36"
# ## Results
# + id="205eaed0-b34e-4efe-9b14-5cb86ce52e9d" jupyter={"source_hidden": true} tags=[]
X = utils.dok_to_array(model_ins.X.get_values())
V = utils.dok_to_array(model_ins.V.get_values())
S = utils.dok_to_array(model_ins.S.get_values())
s = utils.dok_to_array(model_ins.s.get_values())
d = utils.dok_to_array(model_ins.d.get_values())
z = utils.dok_to_array(model_ins.z.get_values())
X_df = pd.DataFrame(X, index=idx_labels["i"], columns=idx_labels["k"])
V_df = pd.DataFrame(V.mean(axis=1), index=idx_labels["i"], columns=idx_labels["k"])
s_df = pd.DataFrame(s.mean(axis=1), index=idx_labels["i"], columns=idx_labels["k"])
d_df = pd.DataFrame(d.mean(axis=1), index=idx_labels["i"], columns=idx_labels["k"])
# + [markdown] id="58a9e407-be17-4e3c-afe6-1dadd98dd8a6"
# The optimal order schedule for each product is shown below:
# + id="f9ea1527-a073-4164-92c7-dc397230f82e" jupyter={"source_hidden": true} outputId="579dc754-8023-4daf-ad16-cd1610915ece" tags=[]
X_df.style.format("{:.0f}").background_gradient(axis=0)
# + [markdown] id="033707a0-feee-4c7e-9da8-230644c89014"
# ### Expected ending inventory
# + id="8b6cceba-75e9-4051-8a01-3ddcb01dddd8" jupyter={"source_hidden": true} outputId="92c76db9-e875-422b-8166-180088cb69ac" tags=[]
fig, ax = plt.subplots(s.shape[-1], 1, figsize=(8, s.shape[-1] * 2), sharex=True)
visualize.scenarios_by_product(s, idx_labels["i"], prod_df.index, ax=ax)
for ax_i in ax.flatten():
ax_i.set_ylim(0)
fig.suptitle("Expected Stock by Product across time")
fig.tight_layout()
# + [markdown] id="48700b51-964f-4402-af3f-f55a3e76c11d"
# ### Inventory Value
# + id="20b51fa9-791c-4f84-976c-ac4253e9eefe" jupyter={"source_hidden": true} outputId="1ef3914b-b63b-47c6-ee3f-bcd66c2e7eaf" tags=[]
inv_val = (S * prod_df["UP"].values).sum(axis=-1, keepdims=True)
fig, ax = plt.subplots(1,1)
visualize.scenarios_by_product(inv_val, idx_labels["i"], ["Total Inventory (PHP)"], ax=ax)
fig.suptitle("Expected Inventory Value across time")
fig.tight_layout()
# + [markdown] id="9c26bef4-8816-4e81-a7a2-90962b153344"
# ### Financial Results
# + id="ca092104-f1d4-4412-a1ae-86c27a7b41e4" jupyter={"source_hidden": true} outputId="b0b735e7-701d-4e4f-828a-3408752003b2" tags=[]
fin_s = pd.Series(dtype="float", name="value")
fin_s["Revenue"] = V_df.mul(prod_df["SP"]).values.sum()
fin_s["COGS"] = - V_df.mul(prod_df["UP"]).values.sum()
fin_s["Contribution Margin"] = fin_s["Revenue"] + fin_s["COGS"]
fin_s["Storage Cost"] = - s_df.mul(prod_df["W"]).values.sum()
fin_s["Out of Stock Cost"] = - d_df.mul(prod_df["O"]).values.sum()
fin_s["Gross Margin"] = fin_s["Contribution Margin"] + fin_s["Storage Cost"] + fin_s["Out of Stock Cost"]
fin_s.to_frame().style.format('PHP {:>10,.2f}')
# + id="1ce9367a-7405-4244-8192-a45850116620"
| Demo Notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys
import time
import cv2
import json
import decimal
import pytz
from pytz import timezone
import datetime
from kafka import KafkaProducer
from kafka.errors import KafkaError
import base64
topic = "testpico"
brokers = ["172.16.31.10:9092"]
def convert_ts(ts, config):
'''Converts a timestamp to the configured timezone. Returns a localized datetime object.'''
#lambda_tz = timezone('US/Pacific')
tz = timezone(config['timezone'])
utc = pytz.utc
utc_dt = utc.localize(datetime.datetime.utcfromtimestamp(ts))
localized_dt = utc_dt.astimezone(tz)
return localized_dt
def publish_camera():
"""
Publish camera video stream to specified Kafka topic.
Kafka Server is expected to be running on the localhost. Not partitioned.
"""
# Start up producer
producer = KafkaProducer(bootstrap_servers=brokers,
value_serializer=lambda v: json.dumps(v).encode('utf-8'))
camera_data = {'camera_id':"1","position":"frontspace","image_bytes":"123"}
camera = cv2.VideoCapture(0)
camera.
framecount = 0
try:
while(True):
success, frame = camera.read()
utc_dt = pytz.utc.localize(datetime.datetime.now())
now_ts_utc = (utc_dt - datetime.datetime(1970, 1, 1, tzinfo=pytz.utc)).total_seconds()
ret, buffer = cv2.imencode('.jpg', frame)
camera_data['image_bytes'] = base64.b64encode(buffer).decode('utf-8')
camera_data['frame_count'] = str(framecount)
camera_data['capture_time'] = str(now_ts_utc)
producer.send(topic, camera_data)
framecount = framecount + 1
# Choppier stream, reduced load on processor
time.sleep(0.2)
if framecount==20:
break
except Exception as e:
print((e))
print("\nExiting.")
sys.exit(1)
camera.release()
producer.close()
if __name__ == "__main__":
publish_camera()
# -
camera = cv2.VideoCapture(0)
# +
camera.set(cv2.CAP_PROP_FRAME_WIDTH,3840)
camera.set(cv2.CAP_PROP_FRAME_HEIGHT,2160)
camera.set(cv2.CAP_PROP_FPS,30)
# -
camera.get(cv2.CAP_PROP_FRAME_COUNT)
| projects/pico/testing/producer-test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import imaplib
M = imaplib.IMAP4_SSL('imap.gmail.com')
import getpass
email = getpass.getpass("Email : ")
password = getpass.getpass("Password: ")
M.login(email,password)
M.list()
M.select('inbox')
typ,data = M.search(None,'SUBJECT "Python"')
typ
data
data[0]
email_id = data[0]
email_id
result, data = M.uid('search', None, "ALL") # search all email and return uids
if result == 'OK':
for num in data[0].split():
result, data = M.uid('fetch', num, '(RFC822)')
if result == 'OK':
email_message = email.message_from_bytes(data[0][1]) # raw email text including headers
print('From:' + email_message['From'])
result , email_data= M.fetch(email_id,'(RFC822)')
em
| receive_mail.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ## Bring in Alignment for mapping
#
# This program will map TFBS using the Biopython's motif package.
#
# **Inputs**:
# 1. before alignment (fasta)
# 2. after alignment (fasta)
# 3. TFBS Position Frequency Matrix.
from Bio import motifs
from Bio import SeqIO
from Bio.Seq import Seq
from Bio.Alphabet import IUPAC, generic_dna, generic_protein
import pandas as pd
import numpy as np
import os, sys
# +
## Alignment Input
# read in alignment as a list of sequences
alignment = list(SeqIO.parse("../data/fasta/output_ludwig_eve-striped-2.fa", "fasta"))
# Check
print("Found %i records in alignment file" % len(alignment))
## Turn a sequences into a list of strings
alignment_string_list = []
for seq in alignment:
alignment_string_list.append(str(seq.seq))
alignment_id = []
for seq in alignment:
alignment_id.append(str(seq.id))
## Do I even need turn int a dataframe?
alignment_df = pd.DataFrame(
{'id': alignment_id,
'align_seq': alignment_string_list
})
## Check
print list(alignment_df)
print type(alignment_df)
##############################
# [x] I feel weird going ahead with out seq ids. Maybe at this point turn into a dictionary?
# [x] No, what I really need is to turn this into a dataframe
# [ ] Now I need to make sure I can use the seqences to continue on. Since I turned it into a dataframe.
# +
## Raw Sequences Input
raw_sequences = list(SeqIO.parse("../data/fasta/ludwig_eve-striped-2.fasta", "fasta"))
print("Found %i records in raw sequence file" % len(raw_sequences))
# make all IUPAC.IUPACUnambiguousDNA()
raw_sequences_2 = []
for seq in raw_sequences:
raw_sequences_2.append(Seq(str(seq.seq), IUPAC.IUPACUnambiguousDNA()))
# Check
#print raw_sequences_2
#print type(raw_sequences_2)
# Check
#for seq in raw_sequences_2:
#print(seq.alphabet)
#print(type(seq))
# +
## Motif Input
bcd = motifs.read(open("../data/PWM/transpose_fm/bcd_FlyReg.fm"),"pfm")
print(bcd.counts)
pwm = bcd.counts.normalize(pseudocounts=0.0)
pssm = pwm.log_odds()
print(pssm.alphabet)
print(type(raw_sequences_2))
# +
## Searching the Sequences
pssm_list = [ ]
for seq in raw_sequences_2:
pssm_list.append(pssm.calculate(seq))
## Check
#print(pssm_list)
#for seq in pssm_list:
#print("Background: %f" % bcd.pssm.mean(bcd.background))
################################
# [ ] Its the same background for all the sequences? That weird. Right?
################################
# Patser Threshold
distribution = pssm.distribution(background=bcd.background, precision=10**4)
threshold = distribution.threshold_patser()
print("Patser Threshold %5.3f" % threshold) #automatically calulate Paster threshold.
print("nothing")
# +
position_list = []
score_list = []
###################################
# [ ] Need to reiterate over raw_sequences_2
# [ ] When reiterating over raw_sequences_2, attach id
##################################
for position, score in pssm.search(raw_sequences_2[0], threshold=6):
position_list.append(position)
score_list.append(score)
# Change position to positive
position_list_pos = []
for x in position_list:
if x < 0:
position_list_pos.append(905 + x)
else:
position_list_pos.append(x)
#print(position_list_pos)
strand = []
for x in position_list:
if x < 0:
strand.append("negative")
else:
strand.append("positive")
# +
## get alignment position using `alignment_string_list`
remap_dict = {}
nuc_list = ['A', 'a', 'G', 'g', 'C', 'c', 'T', 't', 'N', 'n']
counter = 0
#######################
# [ ] Reiterate through all species?
# [ ] maybe create a list of dictionaries?
#######################
for xInd, x in enumerate(alignment_string_list[1]):
if x in nuc_list:
remap_dict[counter] = xInd
counter += 1
# Check
# print(remap_dict)
# Now find the value from the key??? Find the alignment posititon from raw position
align_pos = [remap_dict[x] for x in position_list_pos]
# check
print(align_pos)
# +
# Make dataframe that has everything
pos_df = pd.DataFrame(
{'raw_positition': position_list,
'raw_position_pos_only': position_list_pos,
'alignment_pos':position_list_pos,
'strand_direction': strand,
'score': score_list
})
print(pos_df)
# -
| py/.ipynb_checkpoints/TFBSmapping_draft-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Crater attributes
# Prototype analysis for a single detected crater. Extract crater attributes like best-fitting ellipse, eccentricity, gradient, etc.
# +
import warnings
from pathlib import Path
import numpy as np
import rasterio
from skimage.measure import EllipseModel, regionprops, label, find_contours, approximate_polygon
# Get path to test images
images_dir = Path.cwd().parent.parent / 'test/fixtures'
# Construct filenames to grayscale image and individual crater annotations
img_ctx_fpath = images_dir / 'img1.png'
img_annot_fpaths = [images_dir / f'img1_annot{num}.png' for num in range(3)]
grad_image_fpath = images_dir / 'diagonal_gradient.png'
# +
orig_image = rasterio.open(img_ctx_fpath).read().squeeze()
annot_images = [rasterio.open(img_annot_fpath).read().squeeze()
for img_annot_fpath in img_annot_fpaths]
grad_image = rasterio.open(img_ctx_fpath).read().squeeze()
label_objects = [label(annot_image) for annot_image in annot_images]
props = [regionprops(label_object) for label_object in label_objects]
# +
def calculate_crater_props(binary_label_img):
if binary_label_img.ndim != 2:
raise RuntimeError('Image must be 2d for calculating region properties.')
props = regionprops(binary_label_img)
print(f'Area = {props[0]["area"]}')
print(f'Major Axis = {props[0]["major_axis_length"]}')
print(f'Minor Axis = {props[0]["minor_axis_length"]}')
print(f'Eccentricity = {props[0]["eccentricity"]}')
print(f'Orientation = {props[0]["orientation"]}')
# Spatial moments?
# Euler number?
# TODO: If we use EllipseModel, need to grab only the edge pixels (and not use filled blob)
coords = np.stack(np.nonzero(binary_label_img), axis=1)
em = EllipseModel()
em.estimate(coords)
print(f'Ellipse model params: {em.params}')
def calculate_grad(image, good_inds):
"""Calculate the X and Y gradient for an image using numpy's gradient.
Parameters
==========
image: array-like
Image pixels.
good_inds: array-like
Bool mask for pixels to include when returninng mean gradient.
For example, pass the binary crater mask to only calculate the
gradient for pixels within the crater.
Returns
=======
mean_h: float
Mean horizontal gradient of pixels specified by `good_inds`. Positive
is to the right.
mean_v: float
Mean vertical gradient of pixels specified by `good_inds`. Positive is
downward.
"""
# TODO: could improve function to take a list of good_inds and avoid
# recomputing gradient repeatedly
if good_inds.dtype != bool:
raise ValueError('`good_inds` must be of type bool')
np_mask = np.invert(good_inds) # Pixels we want to exclude should be True
mean_h = np.ma.masked_array(np.gradient(image, axis=0),
mask=np_mask).mean()
mean_v = np.ma.masked_array(np.gradient(image, axis=1),
mask=np_mask).mean()
print(f'Grad horizontal:{mean_h}, Grad vertical:{mean_v}')
return mean_h, mean_v
def calculate_wkt_border(binary_label_img, contour_tol=2.5, pad=1):
"""Pull out the polygon representation of the crater border."""
# TODO: apply and remove padding properly; add test
print(f'orig shape: {binary_label_img.shape}')
padded_image = np.pad(binary_label_img, pad_width=1, mode='constant')
print(f'padded shape: {padded_image.shape}')
contours = find_contours(padded_image, 0.5)
if len(contours) > 1:
warnings.warn(f'Found {len(contours)} contour objects. Expected 1.')
coords = approximate_polygon(contours[0], tolerance=contour_tol)
closed = coords[0] == coords[-1]
return coords, closed
# -
# Get some values from first crater
for li, binary_label_img in enumerate(label_objects):
print (f'\nObject {li}')
calculate_crater_props(binary_label_img)
calculate_grad(orig_image, binary_label_img.astype(np.bool))
# +
import matplotlib.pyplot as plt
fig, axes = plt.subplots(ncols=3, figsize=(15, 7))
plt.gray()
for ii, ax in enumerate(axes):
ax.imshow(annot_images[ii])
contour, closed = calculate_wkt_border(annot_images[ii])
ax.plot(contour[:, 1], contour[:, 0], '-r', linewidth=3)
if ii == 1:
print(contour)
# -
| divdet/inference/crater_attributes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Sphinx教程
# ## 安装
#
# ### 使用pip安装
#
# 可以使用pip安装,如下:
#
# `pip install -U sphinx`
#
# ### 使用docker安装
#
# Sphinx的官方docker镜像有两个:
#
# * sphinxdoc/sphinx,常用的Sphinx镜像。
# * sphinxdoc/sphinx-latexpdf,比较大,主要使用LaTeX构建PDF的Sphinx镜像。
#
# 先安装好docker,然后直接使用docker run命令自动下载镜像并启动容器。如下::
#
# ```
# #从sphinxdoc/sphinx镜像启动一个名称为sphinx的容器并后台运行,挂载当前目录下的project/docs目录到容器docs目录。
# docker run -itd -v $PWD/project/docs:/docs --name sphinx sphinxdoc/sphinx /bin/bash
# #进入sphinx容器
# docker exec -it sphinx /bin/bash
# #运行sphinx-quickstartx脚本生成Sphinx默认模板,
# sphinx-quickstart
# #修改首页index.rst等
# #,仅需运行make html而不需要使用sphinx-build生成。
# make html
# ```
# ## 入门
# ## 扩展
# ### 常用扩展
# #### autodoc
# autodoc扩展能够提取源代码中的文档字符串(DocStrings)生成文档,文档字符串需要按reStructuredText格式编写,可以使用所有常用的Sphinx标记。一般搭配napoleon扩展一起使用。
#
# ```
# 在文档中先设定当前模块
# .. currentmodule:: dfdata
# autodoc的autofunction可以导入函数文档字符串,如:
# .. autofunction:: save_futures_contract
# ```
#
# 详细信息阅读:
# * https://www.sphinx-doc.org/en/master/usage/extensions/autodoc.html
# * https://www.sphinx.org.cn/usage/extensions/autodoc.html
| docs/source/development/document/sphinx.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import matplotlib.pyplot as plt
import numpy as np
# %matplotlib inline
# # Using Pipelines with Grid-Search
# ## Feature selection and regression without pipelines
# +
from sklearn.datasets import make_regression
X, y = make_regression(random_state=42, effective_rank=90)
print(X.shape)
# +
from sklearn.cross_validation import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=42, train_size=.5)
# +
from sklearn.feature_selection import SelectFpr, f_regression
from sklearn.linear_model import Ridge
fpr = SelectFpr(score_func=f_regression)
fpr.fit(X_train, y_train)
X_train_fpr = fpr.transform(X_train)
X_test_fpr = fpr.transform(X_test)
print(X_train_fpr.shape)
# -
ridge = Ridge()
ridge.fit(X_train_fpr, y_train)
ridge.score(X_test_fpr, y_test)
# ## With pipelines
# +
from sklearn.pipeline import make_pipeline
pipe = make_pipeline(SelectFpr(score_func=f_regression), Ridge())
pipe.fit(X_train, y_train)
pipe.score(X_test, y_test)
# -
# ## Grid-Searching alpha in Ridge
from sklearn.grid_search import GridSearchCV
# without pipeline:
param_grid_no_pipeline = {'alpha': 10. ** np.arange(-3, 5)}
pipe.named_steps.keys()
# with pipeline
param_grid = {'ridge__alpha': 10. ** np.arange(-3, 5)}
grid = GridSearchCV(pipe, param_grid, cv=10)
grid.fit(X_train, y_train)
grid.score(X_test, y_test)
grid.best_params_
# ## Selecting parameters of the preprocessing steps
param_grid = {'ridge__alpha': 10. ** np.arange(-3, 5),
'selectfpr__alpha': [0.01, 0.02, 0.05, 0.1, 0.3]}
grid = GridSearchCV(pipe, param_grid, cv=10)
grid.fit(X_train, y_train)
grid.score(X_test, y_test)
grid.best_params_
final_selectfpr = grid.best_estimator_.named_steps['selectfpr']
final_selectfpr.get_support()
| Chapter 3/Parameter Selection with Pipelines.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # fetch_emojis Discord Bot
# ### by riggedCoinflip
#
# ## raw Command Below
#
# Copyright: Do whatever you want with the Code. I would like it if you mention me.
#
# This Bot uses the Guild.emojis as described in:
# https://discordpy.readthedocs.io/en/latest/api.html#guild.emojis <br>
# to create a JSON-File containing these keywords as:
# <blockquote>
# { <br>
#   "emoji.name" = "str(emoji)", <br>
#   . <br>
#   . <br>
#   . <br>
# } <br>
# </blockquote>
#
# with str(emoji) containing name and id.
#
# for example:
# <blockquote>
# { <br>
#   "foo" = "<:foo:123456789012345678>", <br>
#   "bar" = "<:bar:987654321098765432>", <br>
#   "ack" = "<:ack:2582012840158349683>", <br>
#   "wibble" = "<:wibble:287394871261947134>" <br>
# } <br>
# </blockquote>
#
# Attention: This overwrites a (probably) existing file in path.
#
# To use the Bot, follow the instructions on https://discordpy.readthedocs.io/en/latest/discord.html
# and replace Token in the last line (bot.run('Token')) with your generated Token.
#
# Change the Path Variable to where you want your File saved.
# Then write !emojis_json in your Server to fetch the emotes.
# +
# !pip install nest_asyncio
# !pip install discord.py
import nest_asyncio #to allow nested event loop - else: RuntimeError: Cannot Close a running event loop
nest_asyncio.apply()
import discord
from discord.ext import commands
import json
description = '''Exports all Emojis of the Server into a JSON file with: "emoji.name" = "str(emoji)""'''
bot = commands.Bot(command_prefix='!', description=description)
@bot.event
async def on_ready():
print('Logged in as')
print(bot.user.name)
print(bot.user.id)
print('------')
'''Exports all Emojis of the Server into a JSON file with: "emoji.name" = "str(emoji)'''
@bot.command()
async def emojis_json(ctx):
emoji_dict = {emoji.name: str(emoji) for emoji in ctx.guild.emojis} # fill dictionary with values
path = r'C:\folder\emojis.json'
with open(path, 'w') as f:
json.dump(emoji_dict, f, sort_keys=True, indent=4)
print ("Emojis exported to", path)
await ctx.send('Fetching Emojis.....Done!')
bot.run('Token')
# -
# # Raw Command
# +
import json
'''Exports all Emojis of the Server into a JSON file with: "emoji.name" = "str(emoji)'''
@bot.command()
async def emojis_json(ctx):
emoji_dict = {emoji.name: str(emoji) for emoji in ctx.guild.emojis} # fill dictionary with values
path = r'C:\folder\emojis.json'
with open(path, 'w') as f:
json.dump(emoji_dict, f, sort_keys=True, indent=4)
print ("Emojis exported to", path)
await ctx.send('Fetching Emojis.....Done!')
# -
| code/emojis_json.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import lorem
import yaml
from pymongo import MongoClient
from random import choice
from models.todo import TodoSchema
from models.todo_dao import MongoDAO
from library.utils import replace_env, make_url
# +
with open("/config/todos/default_config.yml", "r") as f:
config = yaml.load(f, yaml.SafeLoader)
replace_env(config)
url = make_url(config["database"]["mongo"], include_db=False)
client = MongoClient(url)
collection = client.todos.todos_collection
dao = MongoDAO(collection, TodoSchema)
# +
schema = TodoSchema()
for _ in range(100):
todo = schema.load({
'user_id' : choice([0,1,2,3,4,5]),
'title': lorem.sentence(),
'completed': choice([True,False])
})
dao.add_item(todo)
# -
schema.validate(data={})
schema.validate({
'something':'else'
})
| distributed-todos/services/todos/src/notebooks/create_todos.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6 with Spark
# language: python3
# name: python36
# ---
# ## Regression
#
# TOTAL POINTS 5
#
# ### 1. Question 1
#
# ### The real added value of the author's research on residential real estate properties is quantifying people's preferences of different transport services.
#
# * True.
# * False.
#
# ### Ans: False
#
# ### 2. Question 2
#
# ### Regression is a statistical technique developed by <NAME>.
#
# * False.
# * True.
#
# ### Ans: False
#
# ### 3. Question 3
#
# ### What did the author's research discover about the impact of an additional washroom on the price of a housing unit?
#
# * A. The author found that an additional bedroom adds more to the housing prices than an additional washroom.
# * B. The author found that an additional washroom did not have any impact on the pricing of a housing unit.
# * C. The author found that an additional washroom adds more to the housing prices than an additional bedroom.
# * D. The author found that an additional bedroom adds the same to the housing prices than an additional washroom. In other words, any additional room results in an equal increase to the housing prices.
#
# ### Ans: C
#
# ### 4. Question 4
#
# ### The author discovered that, all else being equal, houses located less than 5 kms but more than 2 kms to shopping centres sold for more than the rest.
#
# * True.
# * False.
#
# ### Ans: True
#
# ### 5.Question 5
#
# ### Based on the reading, which of the following are questions that can be put to regression analysis?
#
# * A. What are typical land taxes in a house sale?
# * B. Do homes with brick exterior sell in rural areas?
# * C. Do homes with brick exterior sell for less than homes with stone exterior?
# * D. What is the impact of lot size on housing price?
#
# ### Ans: CD
| What-is-Data-Science?/Week-2/Quiz-Solutions/Quiz 4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import sys
import h5py
import numpy as np
import corner as DFM
# -- galpopfm --
from galpopfm import dustfm as dustFM
from galpopfm import dust_infer as dustInfer
from galpopfm import measure_obs as measureObs
# -- plotting --
import matplotlib as mpl
import matplotlib.pyplot as plt
mpl.rcParams['text.usetex'] = True
mpl.rcParams['font.family'] = 'serif'
mpl.rcParams['axes.linewidth'] = 1.5
mpl.rcParams['axes.xmargin'] = 1
mpl.rcParams['xtick.labelsize'] = 'x-large'
mpl.rcParams['xtick.major.size'] = 5
mpl.rcParams['xtick.major.width'] = 1.5
mpl.rcParams['ytick.labelsize'] = 'x-large'
mpl.rcParams['ytick.major.size'] = 5
mpl.rcParams['ytick.major.width'] = 1.5
mpl.rcParams['legend.frameon'] = False
dat_dir = os.environ['GALPOPFM_DIR']
# +
#########################################################################
# read in SDSS measurements
#########################################################################
r_edges, gr_edges, fn_edges, _ = dustInfer.sumstat_obs(name='sdss',
statistic='2d', return_bins=True)
dr = r_edges[1] - r_edges[0]
dgr = gr_edges[1] - gr_edges[0]
dfn = fn_edges[1] - fn_edges[0]
ranges = [(r_edges[0], r_edges[-1]), (-0.05, 1.5), (-1., 4.)]
fsdss = os.path.join(dat_dir, 'obs', 'tinker_SDSS_centrals_M9.7.valueadd.hdf5')
sdss = h5py.File(fsdss, 'r')
mr_complete = (sdss['mr_tinker'][...] < -20.)
x_obs = [-1.*sdss['mr_tinker'][...][mr_complete],
sdss['mg_tinker'][...][mr_complete] - sdss['mr_tinker'][...][mr_complete],
sdss['ABSMAG'][...][:,0][mr_complete] - sdss['ABSMAG'][...][:,1][mr_complete]]
# -
def _sim_observables(sim, theta):
''' read specified simulations and return data vector
'''
_sim_sed = dustInfer._read_sed(sim)
wlim = (_sim_sed['wave'] > 1e3) & (_sim_sed['wave'] < 8e3)
#downsample = np.zeros(len(_sim_sed['logmstar'])).astype(bool)
#downsample[::10] = True
downsample = np.ones(len(_sim_sed['logmstar'])).astype(bool)
f_downsample = 1.#0.1
cens = _sim_sed['censat'].astype(bool) & (_sim_sed['logmstar'] > 9.4) & downsample
sim_sed = {}
sim_sed['sim'] = sim
sim_sed['logmstar'] = _sim_sed['logmstar'][cens].copy()
sim_sed['logsfr.inst'] = _sim_sed['logsfr.inst'][cens].copy()
sim_sed['logsfr.100'] = _sim_sed['logsfr.100'][cens].copy()
sim_sed['wave'] = _sim_sed['wave'][wlim].copy()
sim_sed['sed_noneb'] = _sim_sed['sed_noneb'][cens,:][:,wlim].copy()
sim_sed['sed_onlyneb'] = _sim_sed['sed_onlyneb'][cens,:][:,wlim].copy()
x_mod = dustInfer.sumstat_model(theta, sed=sim_sed,
dem='slab_noll_msfr', f_downsample=f_downsample, statistic='2d',
return_datavector=True)
return x_mod, sim_sed
#########################################################################
# read in simulations without dust attenuation
#########################################################################
x_simba, simba = _sim_observables('simba', np.array([0. for i in range(9)]))
zero_sfr100 = simba['logsfr.100'] == -999
print('%i of %i galaxies with 0 SFR_100Myr' % (np.sum(zero_sfr100), len(zero_sfr100)))
zero_sfrinst = simba['logsfr.inst'] == -999
print('%i of %i galaxies with 0 SFR_inst' % (np.sum(zero_sfrinst), len(zero_sfrinst)))
# +
xs = [x_obs, x_simba]
names = ['SDSS', 'SIMBA (no dust)']
clrs = ['k', 'C1']
fig = plt.figure(figsize=(5*len(xs),10))
# R vs (G - R)
for i, _x, name, clr in zip(range(len(xs)), xs, names, clrs):
sub = fig.add_subplot(2,len(xs),i+1)
#sub.pcolormesh(r_edges, gr_edges, _x[1].T,
# vmin=1e-5, vmax=1e-2, norm=mpl.colors.LogNorm(), cmap=clr)
DFM.hist2d(_x[0], _x[1], levels=[0.68, 0.95],
range=[ranges[0], ranges[1]], bins=20, color=clrs[i],
plot_datapoints=True, fill_contours=False, plot_density=True, ax=sub)
if name != 'SDSS':
sub.scatter(_x[0][zero_sfr100], _x[1][zero_sfr100], c='k', s=1)
sub.scatter(_x[0][zero_sfrinst], _x[1][zero_sfrinst], c='C0', s=1)
sub.text(0.95, 0.95, name, ha='right', va='top', transform=sub.transAxes, fontsize=25)
sub.set_xlim(20., 23)
sub.set_xticks([20., 21., 22., 23])
sub.set_xticklabels([])
if i == 0:
sub.set_ylabel(r'$G-R$', fontsize=20)
else:
sub.set_yticklabels([])
sub.set_ylim(ranges[1])
sub.set_yticks([0., 0.5, 1.])
# R vs FUV-NUV
for i, _x, name, clr in zip(range(len(xs)), xs, names, clrs):
sub = fig.add_subplot(2,len(xs),i+len(xs)+1)
#h = sub.pcolormesh(r_edges, fn_edges, _x[2].T,
# vmin=1e-5, vmax=1e-2, norm=mpl.colors.LogNorm(), cmap=clr)
DFM.hist2d(_x[0], _x[2], levels=[0.68, 0.95],
range=[ranges[0], ranges[2]], bins=20, color=clrs[i],
plot_datapoints=True, fill_contours=False, plot_density=True, ax=sub)
if name != 'SDSS':
sub.scatter(_x[0][zero_sfr100], _x[2][zero_sfr100], c='k', s=1)
sub.scatter(_x[0][zero_sfrinst], _x[2][zero_sfrinst], c='C0', s=1)
sub.set_xlim(20., 23)
sub.set_xticks([20., 21., 22., 23])
sub.set_xticklabels([-20, -21, -22, -23])
if i == 0:
sub.set_ylabel(r'$FUV - NUV$', fontsize=20)
else:
sub.set_yticklabels([])
sub.set_ylim(ranges[2])
bkgd = fig.add_subplot(111, frameon=False)
bkgd.set_xlabel(r'$M_r$ luminosity', labelpad=10, fontsize=25)
bkgd.tick_params(labelcolor='none', top=False, bottom=False, left=False, right=False)
fig.subplots_adjust(wspace=0.1, hspace=0.1)
# -
# What happens if we remove the SFRinst = 0 galaxies?
# +
xs = [x_obs, x_simba]
names = ['SDSS', 'SIMBA (no dust)']
clrs = ['k', 'C1']
fig = plt.figure(figsize=(5*len(xs),10))
# R vs (G - R)
for i, _x, name, clr in zip(range(len(xs)), xs, names, clrs):
sub = fig.add_subplot(2,len(xs),i+1)
if name == 'SDSS':
DFM.hist2d(_x[0], _x[1], levels=[0.68, 0.95],
range=[ranges[0], ranges[1]], bins=20, color=clrs[i],
plot_datapoints=True, fill_contours=False, plot_density=True, ax=sub)
else:
sub.scatter(_x[0][zero_sfrinst], _x[1][zero_sfrinst], c='k', s=1)
DFM.hist2d(_x[0][~zero_sfrinst], _x[1][~zero_sfrinst], levels=[0.68, 0.95],
range=[ranges[0], ranges[1]], bins=20, color=clrs[i],
plot_datapoints=True, fill_contours=False, plot_density=True, ax=sub)
sub.text(0.95, 0.95, name, ha='right', va='top', transform=sub.transAxes, fontsize=25)
sub.set_xlim(20., 23)
sub.set_xticks([20., 21., 22., 23])
sub.set_xticklabels([])
if i == 0:
sub.set_ylabel(r'$G-R$', fontsize=20)
else:
sub.set_yticklabels([])
sub.set_ylim(ranges[1])
sub.set_yticks([0., 0.5, 1.])
# R vs FUV-NUV
for i, _x, name, clr in zip(range(len(xs)), xs, names, clrs):
sub = fig.add_subplot(2,len(xs),i+len(xs)+1)
if name == 'SDSS':
DFM.hist2d(_x[0], _x[2], levels=[0.68, 0.95],
range=[ranges[0], ranges[2]], bins=20, color=clrs[i],
plot_datapoints=True, fill_contours=False, plot_density=True, ax=sub)
else:
sub.scatter(_x[0][zero_sfrinst], _x[2][zero_sfrinst], c='k', s=1, label='inst. SFR = 0')
DFM.hist2d(_x[0][~zero_sfrinst], _x[2][~zero_sfrinst], levels=[0.68, 0.95],
range=[ranges[0], ranges[2]], bins=20, color=clrs[i],
plot_datapoints=True, fill_contours=False, plot_density=True, ax=sub)
sub.legend(loc='lower right', handletextpad=0.1, markerscale=5, fontsize=20)
sub.set_xlim(20., 23)
sub.set_xticks([20., 21., 22., 23])
sub.set_xticklabels([-20, -21, -22, -23])
if i == 0:
sub.set_ylabel(r'$FUV - NUV$', fontsize=20)
else:
sub.set_yticklabels([])
sub.set_ylim(ranges[2])
bkgd = fig.add_subplot(111, frameon=False)
bkgd.set_xlabel(r'$M_r$ luminosity', labelpad=10, fontsize=25)
bkgd.tick_params(labelcolor='none', top=False, bottom=False, left=False, right=False)
fig.subplots_adjust(wspace=0.1, hspace=0.1)
# -
# the SFR_inst = 0 population has a very distinct imprint on the observables: e.g. they are tightly clustered in the color-magnitude space.
def _sim_observables_resample(sim, theta):
''' read specified simulations and return data vector
'''
_sim_sed = dustInfer._read_sed(sim)
wlim = (_sim_sed['wave'] > 1e3) & (_sim_sed['wave'] < 8e3)
#downsample = np.zeros(len(_sim_sed['logmstar'])).astype(bool)
#downsample[::10] = True
downsample = np.ones(len(_sim_sed['logmstar'])).astype(bool)
f_downsample = 1.#0.1
cens = _sim_sed['censat'].astype(bool) & (_sim_sed['logmstar'] > 9.4) & downsample
sim_sed = {}
sim_sed['sim'] = sim
sim_sed['logmstar'] = _sim_sed['logmstar'][cens].copy()
sim_sed['logsfr.inst'] = _sim_sed['logsfr.inst'][cens].copy()
sim_sed['wave'] = _sim_sed['wave'][wlim].copy()
sim_sed['sed_noneb'] = _sim_sed['sed_noneb'][cens,:][:,wlim].copy()
sim_sed['sed_onlyneb'] = _sim_sed['sed_onlyneb'][cens,:][:,wlim].copy()
_zero = (sim_sed['logsfr.inst'] == -999)
sim_sed['logsfr.inst'][_zero] = np.random.uniform(0., 0.0004018745094072074, size=np.sum(_zero))
x_mod = dustInfer.sumstat_model(theta, sed=sim_sed,
dem='slab_noll_msfr', f_downsample=f_downsample, statistic='2d',
return_datavector=True)
return x_mod, sim_sed
#########################################################################
# read in simulations with some generic dust attenuation
#########################################################################
x_simba, _ = _sim_observables_resample('simba', np.array([1., 2., 2., 0., 0., -0.5, 0., 0., 2.]))
# +
xs = [x_obs, x_simba]
names = ['SDSS', 'SIMBA (some DEM)']
clrs = ['k', 'C1']
fig = plt.figure(figsize=(5*len(xs),10))
# R vs (G - R)
for i, _x, name, clr in zip(range(len(xs)), xs, names, clrs):
sub = fig.add_subplot(2,len(xs),i+1)
if name == 'SDSS':
DFM.hist2d(_x[0], _x[1], levels=[0.68, 0.95],
range=[ranges[0], ranges[1]], bins=20, color=clrs[i],
plot_datapoints=True, fill_contours=False, plot_density=True, ax=sub)
else:
sub.scatter(_x[0][zero_sfrinst], _x[1][zero_sfrinst], c='k', s=1)
DFM.hist2d(_x[0][~zero_sfrinst], _x[1][~zero_sfrinst], levels=[0.68, 0.95],
range=[ranges[0], ranges[1]], bins=20, color=clrs[i],
plot_datapoints=True, fill_contours=False, plot_density=True, ax=sub)
sub.text(0.95, 0.95, name, ha='right', va='top', transform=sub.transAxes, fontsize=25)
sub.set_xlim(20., 23)
sub.set_xticks([20., 21., 22., 23])
sub.set_xticklabels([])
if i == 0:
sub.set_ylabel(r'$G-R$', fontsize=20)
else:
sub.set_yticklabels([])
sub.set_ylim(ranges[1][0], 2.)
sub.set_yticks([0., 0.5, 1.])
# R vs FUV-NUV
for i, _x, name, clr in zip(range(len(xs)), xs, names, clrs):
sub = fig.add_subplot(2,len(xs),i+len(xs)+1)
if name == 'SDSS':
DFM.hist2d(_x[0], _x[2], levels=[0.68, 0.95],
range=[ranges[0], ranges[2]], bins=20, color=clrs[i],
plot_datapoints=True, fill_contours=False, plot_density=True, ax=sub)
else:
sub.scatter(_x[0][zero_sfrinst], _x[2][zero_sfrinst], c='k', s=1, label='inst. SFR = 0')
DFM.hist2d(_x[0][~zero_sfrinst], _x[2][~zero_sfrinst], levels=[0.68, 0.95],
range=[ranges[0], ranges[2]], bins=20, color=clrs[i],
plot_datapoints=True, fill_contours=False, plot_density=True, ax=sub)
sub.legend(loc='lower right', handletextpad=0.1, markerscale=5, fontsize=20)
sub.set_xlim(20., 23)
sub.set_xticks([20., 21., 22., 23])
sub.set_xticklabels([-20, -21, -22, -23])
if i == 0:
sub.set_ylabel(r'$FUV - NUV$', fontsize=20)
else:
sub.set_yticklabels([])
sub.set_ylim(ranges[2][0], 10.)
bkgd = fig.add_subplot(111, frameon=False)
bkgd.set_xlabel(r'$M_r$ luminosity', labelpad=10, fontsize=25)
bkgd.tick_params(labelcolor='none', top=False, bottom=False, left=False, right=False)
fig.subplots_adjust(wspace=0.1, hspace=0.1)
# -
# Even with attenuation, these galaxies are quite clustered in the observable space. This will likely affect the distance.
| nb/sfr_res_issue.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## _*Initializing next computation from prior result*_
#
# This notebook demonstrates using Qiskit Chemistry to plot graphs of the ground state energy of the Hydrogen (H2) molecule over a range of inter-atomic distances using VQE and TwoLocal. It is compared to the same energies as computed by the NumPyMinimumEigensolver and we also compare using the previous computed optimal solution as the starting initial point for the next distance.
#
# This notebook has been written to use the PYQUANTE chemistry driver. See the PYQUANTE chemistry driver readme if you need to install the external PyQuante2 library that this driver requires.
# +
import numpy as np
import pylab
import copy
from qiskit import BasicAer
from qiskit.aqua import aqua_globals, QuantumInstance
from qiskit.aqua.algorithms import NumPyMinimumEigensolver, VQE
from qiskit.aqua.components.optimizers import COBYLA
from qiskit.circuit.library import TwoLocal
from qiskit.chemistry.drivers import PyQuanteDriver, BasisType
from qiskit.chemistry.core import Hamiltonian, QubitMappingType
molecule = 'H .0 .0 -{0}; H .0 .0 {0}'
algorithms = [{'name': 'VQE'},
{'name': 'VQE'},
{'name': 'NumPyMinimumEigensolver'}]
titles= ['VQE Random Seed', 'VQE + Initial Point', 'NumPyMinimumEigensolver']
start = 0.5 # Start distance
by = 0.5 # How much to increase distance by
steps = 20 # Number of steps to increase by
energies = np.empty([len(algorithms), steps+1])
hf_energies = np.empty(steps+1)
distances = np.empty(steps+1)
eval_counts = np.zeros([len(algorithms), steps+1], dtype=np.intp)
aqua_globals.random_seed = 50
print('Processing step __', end='')
for i in range(steps+1):
print('\b\b{:2d}'.format(i), end='', flush=True)
d = start + i*by/steps
for j in range(len(algorithms)):
driver = PyQuanteDriver(molecule.format(d/2), basis=BasisType.BSTO3G)
qmolecule = driver.run()
operator = Hamiltonian(qubit_mapping=QubitMappingType.PARITY, two_qubit_reduction=True)
qubit_op, aux_ops = operator.run(qmolecule)
if algorithms[j]['name'] == 'NumPyMinimumEigensolver':
result = NumPyMinimumEigensolver(qubit_op).run()
else:
optimizer = COBYLA(maxiter=10000)
var_form = TwoLocal(qubit_op.num_qubits, ['ry', 'rz'], 'cz', reps=5, entanglement='linear')
algo = VQE(qubit_op, var_form, optimizer)
result = algo.run(QuantumInstance(BasicAer.get_backend('statevector_simulator'),
seed_simulator=aqua_globals.random_seed,
seed_transpiler=aqua_globals.random_seed))
eval_counts[j][i] = result.optimizer_evals
if j == 1:
algorithms[j]['initial_point'] = result.optimal_point.tolist()
result = operator.process_algorithm_result(result)
energies[j][i] = result.energy
hf_energies[i] = result.hartree_fock_energy
distances[i] = d
print(' --- complete')
print('Distances: ', distances)
print('Energies:', energies)
print('Hartree-Fock energies:', hf_energies)
print('VQE num evaluations:', eval_counts)
# -
# The plot of ground energies from VQE, whether starting from a random initial point or the optimal solution from the prior point are indistinguisable here.
pylab.plot(distances, hf_energies, label='Hartree-Fock')
for j in range(len(algorithms)):
pylab.plot(distances, energies[j], label=titles[j])
pylab.xlabel('Interatomic distance')
pylab.ylabel('Energy')
pylab.title('H2 Ground State Energy')
pylab.legend(loc='upper right');
for i in range(2):
pylab.plot(distances, np.subtract(energies[i], energies[2]), label=titles[i])
pylab.plot(distances, np.subtract(hf_energies, energies[2]), label='Hartree-Fock')
pylab.xlabel('Interatomic distance')
pylab.ylabel('Energy')
pylab.title('Energy difference from NumPyMinimumEigensolver')
pylab.legend(loc='upper left');
# Lets plot the difference of the VQE ground state energies from the NumPyMinimumEigensolver. They are both in the same ballpark and both very small.
for i in range(len(algorithms)-1):
pylab.plot(distances, np.subtract(energies[i], energies[2]), label=titles[i])
pylab.xlabel('Interatomic distance')
pylab.ylabel('Energy')
pylab.yscale('log')
pylab.title('H2 Ground State Energy')
pylab.legend(loc='upper right');
# Finally lets plot the number of evaluations taken at each point. Both start out at the same number since we start them the same. But we can see, as we step along small distances, that the prior solution is a better guess as the starting point for the next step leading to fewer evaluations.
for i in range(2):
pylab.plot(distances, eval_counts[i], '-o', label=titles[i])
pylab.xlabel('Interatomic distance')
pylab.ylabel('Evaluations')
pylab.title('VQE number of evaluations')
pylab.legend(loc='center left');
# +
for i in range(2):
print("Total evaluations for '{}' = {}".format(titles[i], np.sum(eval_counts[i])))
percent = np.sum(eval_counts[1])*100/np.sum(eval_counts[0])
print("\nTotal evaluations for '{}' are {:.2f}% of '{}'".format(titles[1], percent, titles[0]))
| chemistry/h2_vqe_initial_point.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="w_e5GXnWH9tO" colab_type="text"
# # Character-Level Text Generation 2
#
# Now it is your turn. Choose any text from the selection below and try to generate something interesting.
# + id="rwxnYkcOhgjB" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
import keras
import numpy as np
import re, collections # for text processing
from google.colab import files # for download files
# + [markdown] id="BLMS2qirKgCs" colab_type="text"
# ## Step 1) Choose the dataset
#
# Select one of the texts below by uncommenting the corresponding line:
# * Shakespeare's sonets (95kb)
# * Obama speaches (4.3Mb)
# * Skautské stanovy (482kb, in Czech)
# * Bible kralická (3.9Mb, in Czech)
# + id="Tzfhr5p1aFWD" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
# Uncomment one of the lines below:
# path = keras.utils.get_file('sonnet.txt', origin='https://raw.githubusercontent.com/michaelrzhang/Char-RNN/master/data/sonnet.txt')
# path = keras.utils.get_file('obama.txt', origin='https://raw.githubusercontent.com/michaelrzhang/Char-RNN/master/data/obama.txt')
# path = keras.utils.get_file('skautske_stanovy.txt', origin='https://drive.google.com/uc?export=download&id=0B2yuKzlrzs84SnhBZ2tYSTA3aG8')
# path = keras.utils.get_file('bible.txt', origin='https://drive.google.com/uc?export=download&id=0B3hE_6FIbbVWdy1FQTZOS29NZkk')
with open(path, encoding='utf-8') as f:
text = f.read()
print('corpus length:', len(text))
# + [markdown] id="iDGmLs5eK3NI" colab_type="text"
# ## Step 2) Text preprocessing
#
# Think carefully about text preprocessing. Which characters do you want to keep and which should be removed?
# + id="fbvOJk0JauHR" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
# You might want to apply one of the transformations below:
# text = text.lower()
# text = text.replace("\n", " ")
# text = re.sub('[ëéä]', '', text)
chars = sorted(list(set(text)))
num_chars = len(chars)
print('total characters in vocabulary:', num_chars)
charcounts = collections.Counter(list(text))
sorted(charcounts.items(), key=lambda i: i[1])
# + [markdown] id="aOT2fC_-MJZD" colab_type="text"
# ## Step 3) Cut the text in semi-redundant sequences
#
# For training, the test is cut into smaller pieces of the same length. Longer pieces enable better context but needs more time and memory for training.
# + id="su1RvGsRbLxc" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
SEQ_LENGTH = 40 # length of sequences
STEP = 10 # shift in cursor between sequences
DEPTH = 1 # number of hidden LSTM/GRU layers
UNIT_SIZE = 128 # number of units per LSTM
DROPOUT = 0.1 # dropout parameter
# + id="OBXopIx-nwU8" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
sentences = list()
targets = list()
for i in range(0, len(text) - SEQ_LENGTH - 1, STEP):
sentences.append(text[i: i + SEQ_LENGTH])
targets.append(text[i + 1: i + SEQ_LENGTH + 1])
print('number of sequences:', len(sentences))
# + [markdown] id="qcxSCl-0NUHF" colab_type="text"
# ## Step 4) Vectorization
#
# One reason to do this is that entering raw numbers into a RNN may not make sense
# because it assumes an ordering for catergorical variables.
# + id="aYavT4Lko2t-" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
# dictionaries to convert characters to numbers and vice-versa
char_to_indices = dict((c, i) for i, c in enumerate(chars))
indices_to_char = dict((i, c) for i, c in enumerate(chars))
X = np.zeros((len(sentences), SEQ_LENGTH, num_chars), dtype=np.bool)
y = np.zeros((len(sentences), SEQ_LENGTH, num_chars), dtype=np.bool)
for i in range(len(sentences)):
sentence = sentences[i]
target = targets[i]
for j in range(SEQ_LENGTH):
X[i][j][char_to_indices[sentence[j]]] = 1
y[i][j][char_to_indices[target[j]]] = 1
# + [markdown] id="O_fGbXpYP_3d" colab_type="text"
# ## Step 5) Model definition
#
# One, two (or three) layers of LSTM and dropout, followed by dense connected layer and softmax. You can experiment and modify this code: use GRU (keras.layers.GRU) instead of LSTM, try SGD or Adam optimizers instead of RMSprop and modify learning rate (lr parameter).
# + id="XOHedg-vboEs" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
model = keras.models.Sequential()
for _ in range(DEPTH):
model.add(keras.layers.LSTM(UNIT_SIZE, input_shape=(None, num_chars), return_sequences=True))
model.add(keras.layers.Dropout(DROPOUT))
model.add(keras.layers.wrappers.TimeDistributed(keras.layers.Dense(num_chars)))
model.add(keras.layers.Activation('softmax'))
# + id="lTXKpmFLdpG7" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
optimizer = keras.optimizers.RMSprop(lr=0.01)
model.compile(loss='categorical_crossentropy', optimizer=optimizer, metrics=['accuracy'])
# + [markdown] id="sS1LR_jlRiEd" colab_type="text"
# ## Helper functions: Generating text from the model
#
# The function **sample** takes the trained model and get you a sample of a text generated from it.
# You can set the beginning (`set.seed`) and `temperature`. Lower temperature makes text more confident (but also more conservative).
# + id="fKDuSeEpd8Fr" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
def multinomial_with_temperature(preds, temperature=1.0):
"""
Helper function to sample from a multinomial distribution (+adj. for temperature)
"""
preds = np.asarray(preds).astype('float64')
preds = np.log(preds + 1e-8) / temperature
exp_preds = np.exp(preds)
preds = exp_preds / np.sum(exp_preds)
probas = np.random.multinomial(1, preds, 1)
return np.argmax(probas)
def sample(model, char_to_indices, indices_to_char,
seed_string=" ", temperature=1.0, test_length=150):
"""
Generates text of test_length length from model starting with seed_string.
"""
num_chars = len(char_to_indices.keys())
for i in range(test_length):
test_in = np.zeros((1, len(seed_string), num_chars))
for t, char in enumerate(seed_string):
test_in[0, t, char_to_indices[char]] = 1
entire_prediction = model.predict(test_in, verbose=0)[0]
next_index = multinomial_with_temperature(entire_prediction[-1], temperature)
next_char = indices_to_char[next_index]
seed_string = seed_string + next_char
return seed_string
# + [markdown] id="B9aqL0FsTr1t" colab_type="text"
# ## Step 6) Model training
#
# Each time you run the code below, the model is trained for 10 epochs (each sequence is visited 10 times). If the quality of predictions is not sufficient, you can add another 10 epochs, etc.
# + id="-GWjeLVYeBz3" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
history = model.fit(X, y,
batch_size=1024,
epochs=10)
# + [markdown] id="XRPywzlTUyBj" colab_type="text"
# ## Step 7) Generate text
#
# Is it good? Congratulation! You can save and download the model with the code below. Does it need improvement? Either you need more training (Step 6) or you need to change your parameters or model definition (Steps 3 and 5).
# + id="UZnAKchWmigy" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
sample(model, char_to_indices=char_to_indices, indices_to_char=indices_to_char, seed_string="truth", temperature=0.8)
# + [markdown] id="zq5jWTRmV0vK" colab_type="text"
# ## Step 8) Saving the model
# + id="hLMURFCF7ncu" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
model_filename = 'nietzsche.loss{0:.2f}.h5'.format(history.history['loss'][-1])
model.save(model_filename)
files.download(model_filename)
# + [markdown] id="jQUKJwW2WEgs" colab_type="text"
# ## Acknowledgement
#
# This notebook was adapted from <NAME>'s [Char-RNN](https://github.com/michaelrzhang/Char-RNN) and [lstm_text_generation.py](https://github.com/keras-team/keras/blob/master/examples/lstm_text_generation.py) example in keras github repo. Both were inspired from <NAME>'s blog post [The Unreasonable Effectiveness of Recurrent Neural Networks](http://karpathy.github.io/2015/05/21/rnn-effectiveness/).
# + id="FzWOWh-haGfx" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
| 05-Your_text_generation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# =============================================================
# Copyright © 2020 Intel Corporation
#
# SPDX-License-Identifier: MIT
# =============================================================
# -
# # Daal4py K-Means Clustering Example for Distributed Memory Systems [SPMD mode]
# ## IMPORTANT NOTICE
# When using daal4py for distributed memory systems, the command needed to execute the program should be **executed
# in a bash shell**. In order to run this example, please download it as a .py file then run the following command (**the number 4 means that it will run on 4 processes**):
# mpirun -n 4 python ./daal4py_Distributed_Kmeans.py
# ## Importing and Organizing Data
# In this example we will be using K-Means clustering to **initialize centroids** and then **use them to cluster the synthetic dataset.**
#
# Let's start by **importing** all necessary data and packages.
##### daal4py K-Means Clustering example for Distributed Memory Systems [SPMD Mode] #####
import daal4py as d4p
import pickle
import pandas as pd
import numpy as np
# Now let's **load** in the dataset and **organize** it as necessary to work with our model. For distributed, every file has a unique ID.
#
# We will also **initialize the distribution engine**.
# +
d4p.daalinit() #initializes the distribution engine
# organizing variables used in the model for prediction
# each process gets its own data
infile = "./data/distributed_data/daal4py_Distributed_Kmeans_" + str(d4p.my_procid()+1) + ".csv"
# read data
X = pd.read_csv(infile)
# -
# ## Computing and Saving Initial Centroids
# Time to **initialize our centroids!**
# computing inital centroids
init_result = d4p.kmeans_init(nClusters = 3, method = "plusPlusDense").compute(X)
# To **get initial centroid information and save it** to a file:
# +
# retrieving and printing inital centroids
centroids = init_result.centroids
print("Here's our centroids:\n\n\n", centroids, "\n")
centroids_filename = './models/kmeans_clustering_initcentroids_'+ str(d4p.my_procid()+1) + '.csv'
# saving centroids to a file
pickle.dump(centroids, open(centroids_filename, "wb"))
# -
# Now let's **load up the centroids** and look at them.
# loading the initial centroids from a file
loaded_centroids = pickle.load(open(centroids_filename, "rb"))
print("Here is our centroids loaded from file:\n\n",loaded_centroids)
# # Assign The Data to Clusters and Save The Results
# Let's **assign the data** to clusters.
# compute the clusters/centroids
kmeans_result = d4p.kmeans(nClusters = 3, maxIterations = 5, assignFlag = True).compute(X, init_result.centroids)
# To **get Kmeans result objects** (assignments, centroids, goalFunction [deprecated], nIterations, and objectiveFunction):
# retrieving and printing cluster assignments
assignments = kmeans_result.assignments
print("Here is our cluster assignments for first 5 datapoints: \n\n", assignments[:5])
| AI-and-Analytics/Features-and-Functionality/IntelPython_daal4py_DistributedKMeans/IntelPython_daal4py_Distributed_Kmeans.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
folders = os.listdir("data_gray")
print(folders)
# +
count = 0
image_data =[]
labels = []
label_dict = {
"NonViolence" : 0,
"Violence" : 1
}
print(label_dict)
# -
for ix in folders:
path= os.path.join("./data_gray/",ix)
print(path)
for im in os.listdir(path):
print(im)
# +
from keras.preprocessing import image
import matplotlib.pyplot as plt
for ix in folders:
path= os.path.join("./data_gray",ix)
print(path)
for im in os.listdir(path):
#print(im)
img = image.load_img(os.path.join(path,im), target_size = ((224,224)))
count = count + 1
# print(img)
img_array = image.img_to_array(img)
image_data.append(img_array)
labels.append(label_dict[ix])
print(count)
print(count)
# -
print(labels)
print(img_array)
# +
# print(image_data)
# -
print(len(image_data),len(labels))
print(labels[-4227])
import random
combined = list(zip(image_data,labels))
random.shuffle(combined)
image_data[:], labels[:] = zip(*combined)
print(labels[:10])
import numpy as np
X_train = np.array(image_data)
Y_train = np.array(labels)
print(X_train.shape,Y_train.shape)
from keras.utils import np_utils
Y_train = np_utils.to_categorical(Y_train)
print(X_train.shape,Y_train.shape)
from keras.applications.resnet50 import ResNet50
from keras.optimizers import Adam
from keras.layers import *
from keras.models import Model
model = ResNet50(include_top =False, weights = "imagenet", input_shape = (224,224,3))
model.summary()
# +
av1 = GlobalAveragePooling2D()(model.output)
fc1 = Dense(256 , activation = "relu")(av1)
d1 = Dropout(0.5)(fc1)
fc2 = Dense(2 , activation= "softmax")(d1)
model_new = Model(input = model.input, output = fc2)
model_new.summary()
# -
adam = Adam(lr = 0.00003)
model_new.compile(loss = "categorical_crossentropy", optimizer ="adam", metrics = ["accuracy"] )
for ix in range(len(model_new.layers)):
print(ix , model_new.layers[ix])
for ix in range(169):
model_new.layers[ix].trainable = False
model_new.compile(loss = "categorical_crossentropy", optimizer ="adam", metrics = ["accuracy"] )
model_new.summary()
# +
hist = model_new.fit(X_train , Y_train, shuffle=True , batch_size = 128, epochs = 1, validation_split=0.20)
model_new.save("weights.h5")
print("model weights are save on disk")
# -
import cv2
import numpy as np
import matplotlib.pyplot as plt
# +
# img = cv2.imread("fight_test.jpg")
# # img = cv2.resize(224,224,3)
# img = cv2.cvtColor(img, cv2.COLOR_BGR2RGB)
# # img = cv2.cvtColor(img, cv2.COLOR_RGB2GRAY)
# img_array = np.array(img)
# plt.imshow(img_array)
# -
img_test = cv2.imread("b_non_fight6.jpg")
img_test = cv2.resize(img_test, (224, 224))
img_test = cv2.cvtColor(img_test, cv2.COLOR_BGR2RGB)
img_test_array = np.array(img_test)
plt.imshow(img_test_array)
img_test_array.shape = (1,224,224,3)
z = model_new.predict(img_test_array)
print(z)
| model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Drugmonizome-Consensus
# language: python
# name: drugmonizome-consensus
# ---
# ## Drug Set Consensus Enrichment
# Author : <NAME> | <EMAIL>¶
# appyter init
from appyter import magic
magic.init(lambda _=globals: _())
# +
import json
import csv
import time
import math
from IPython.display import display, IFrame, Markdown
from tqdm import tqdm
import pandas as pd
import seaborn as sns
import matplotlib
from matplotlib import pyplot as plt
import requests
# +
# %%appyter hide_code
{% do SectionField(
name='data',
title='Drugmonizome Consensus Terms',
subtitle='This appyter returns consensus terms from Drugmonizome signature search results using a set of drug sets',
img='drugmonizome_logo.ico'
) %}
{% set gmt_file = FileField(
constraint='.*\.gmt$',
name='gmt_files',
label='Drug Set',
default='example.gmt',
examples={
'example.gmt': 'https://appyters.maayanlab.cloud/storage/Drugmonizome_Consensus/example.gmt'
},
section='data'
) %}
# -
# ### Set Global Variables and Import Input
# +
# %%appyter code_exec
# Drugmonizome API URLs
entities_endpoint = 'https://maayanlab.cloud/drugmonizome/metadata-api/entities/find'
signatures_endpoint = 'https://maayanlab.cloud/drugmonizome/metadata-api/signatures/'
enrichment_endpoint = 'https://maayanlab.cloud/drugmonizome/data-api/api/v1/enrich/overlap'
# Figure & Table Labels
table = 1
figure = 1
# Drug set libraries
libraries = {{MultiCheckboxField(
name = 'datasets',
label = 'Drug set libraries from Drugmonizome',
choices = [
'L1000FWD_GO_Biological_Processes_drugsetlibrary_up',
'L1000FWD_GO_Biological_Processes_drugsetlibrary_down',
'L1000FWD_GO_Cellular_Component_drugsetlibrary_up',
'L1000FWD_GO_Cellular_Component_drugsetlibrary_down',
'L1000FWD_GO_Molecular_Function_drugsetlibrary_up',
'L1000FWD_GO_Molecular_Function_drugsetlibrary_down',
'L1000FWD_KEGG_Pathways_drugsetlibrary_up',
'L1000FWD_KEGG_Pathways_drugsetlibrary_down',
'L1000FWD_signature_drugsetlibrary_up',
'L1000FWD_signature_drugsetlibrary_down',
'L1000FWD_predicted_side_effects',
'Geneshot_associated_drugsetlibrary',
'Geneshot_predicted_coexpression_drugsetlibrary',
'Geneshot_predicted_autorif_drugsetlibrary',
'Geneshot_predicted_generif_drugsetlibrary',
'Geneshot_predicted_enrichr_drugsetlibrary',
'Geneshot_predicted_tagger_drugsetlibrary',
'KinomeScan_kinase_drugsetlibrary',
'ATC_drugsetlibrary',
'CREEDS_signature_drugsetlibrary_up',
'CREEDS_signature_drugsetlibrary_down',
'DrugRepurposingHub_moa_drugsetlibrary',
'DrugRepurposingHub_target_drugsetlibrary',
'Drugbank_smallmolecule_target_drugsetlibrary',
'Drugbank_smallmolecule_carrier_drugsetlibrary',
'Drugbank_smallmolecule_transporter_drugsetlibrary',
'Drugbank_smallmolecule_enzyme_drugsetlibrary',
'DrugCentral_target_drugsetlibrary',
'PharmGKB_OFFSIDES_side_effects_drugsetlibrary',
'PharmGKB_snp_drugsetlibrary',
'SIDER_side_effects_drugsetlibrary',
'SIDER_indications_drugsetlibrary',
'STITCH_target_drugsetlibrary',
'RDKIT_maccs_fingerprints_drugsetlibrary'
],
default = ['L1000FWD_GO_Biological_Processes_drugsetlibrary_down',
'L1000FWD_GO_Biological_Processes_drugsetlibrary_up'
],
section = 'data'
)}}
# User-defined parameters
alpha = {{FloatField(
name='alpha',
label='p-value cutoff',
default=0.05,
section='data'
)}}
top_results = {{IntField(
name = 'min_count',
label = 'Top results',
description = 'Number of top results to keep',
default = 20,
section = 'data'
)}}
# +
# Plotting functions
cmap = sns.cubehelix_palette(50, hue=0.05, rot=0, light=1, dark=0)
def heatmap(df, filename, width=20, height=10):
cg = sns.clustermap(df, cmap=cmap, figsize=(width, height), cbar_pos=(0.02, 0.65, 0.05, 0.18),)
cg.ax_row_dendrogram.set_visible(False)
cg.ax_col_dendrogram.set_visible(False)
display(cg)
plt.show()
cg.savefig(filename)
def stackedBarPlot(df, filename, width = 15, height = 10):
df['mean'] = df.mean(axis=1)
df_bar = df.sort_values(by = 'mean', ascending = False)[0:top_results]\
.sort_values(by = 'mean')\
.drop(['mean'], axis = 1)
df_bar.plot.barh(stacked = True, figsize = (width,height), fontsize = 20)
plt.legend(bbox_to_anchor=(1.25, 0.30), loc='lower right', prop={'size': 16})
plt.xlabel('-log(p)',labelpad = 20, fontsize = 'xx-large')
plt.savefig(filename, format = 'svg', bbox_inches='tight')
plt.show()
# +
# Drugmonizome enrichment functions
def get_entity_uuids(drug_list):
filter_body = {
"filter": {
"where": {
"meta.Name": {
"inq": drug_list
}
}
}
}
entities = requests.post(entities_endpoint, json=filter_body)
# create UUID dict matched to names
entity_lookup = {}
for item in entities.json():
entity_lookup[item['id']] = item['meta']['Name']
return entity_lookup
def enrich(entity_lookup, library, alpha):
output = []
payload = {'database': library ,'entities': list(entity_lookup.keys())}
res = requests.post(enrichment_endpoint, json = payload)
try:
for item in res.json()['results']:
if item['p-value'] < alpha:
signature = requests.get(signatures_endpoint + item['uuid']).json()
# Create output object with UUIDs for entities and signatures decoded
output.append({'term': signature['meta']['Term'][0]['Name'],
'p-value': item['p-value'],
'overlap': list(set([entity_lookup.get(x,x) for x in item['overlap']]))
})
except (json.decoder.JSONDecodeError, ValueError):
pass
return output
# -
# %%appyter code_exec
drug_sets = {}
with open({{gmt_file}}, 'r') as f:
reader = csv.reader(f, delimiter = '\t')
for row in reader:
drug_sets[row[0]] = {
"drug_list": list(set([str(drug).lower() for drug in row[2:]]))
}
# ### Query drug sets through Drugmonizome to retrieve enrichment results
for description, values in tqdm(drug_sets.items()):
drugs = values["drug_list"]
entity_uuids = get_entity_uuids(drugs)
drug_sets[description]["libraries"] = {}
for library in libraries:
results = enrich(entity_uuids, library, alpha)
drug_sets[description]["libraries"][library] = results
time.sleep(0.2)
enrichment_df = {}
num_sets = len(drug_sets)
for lib in libraries:
term_df = pd.DataFrame(columns=drug_sets.keys())
for k,v in drug_sets.items():
sigs = v["libraries"][lib]
for sig in sigs:
term = sig['term']
p = sig['p-value']
term_df.at[term, k] = -math.log(p)
term_df.fillna(0.0, inplace = True)
term_df.to_csv("%s_enrichment_table.tsv"%lib, sep="\t")
enrichment_df[lib] = term_df
display(term_df.head(10))
display(Markdown(" **Table %d** The table above shows the enrichment analysis results of %d drug sets \
with the **%s** library in Drugmonizome. Each score is computed by getting the negative logarithm of the p-value \
($-\ln{pval}$). [Download complete table](%s_enrichment_table.tsv)"%(table, num_sets, lib.replace("_"," "), lib)))
table += 1
# ## Analysis
# Heatmaps and stacked bar plots will be created for each library's enrichment results
for lib in libraries:
df = enrichment_df[lib]
consensus = df.sum(1).sort_values(ascending=False)[0:top_results].to_frame(name="scores")
consensus.to_csv("%s_consensus_table.tsv"%lib, sep="\t")
display(consensus.head(10))
display(Markdown("**Table %d** %s consensus terms. \
[Download top %d terms](%s_consensus_table.tsv)"%(table, lib.replace("_"," "), top_results, lib)))
table +=1
consensus_df = df.loc[consensus.index]
if (consensus_df.shape[1] > 0):
heatmap(consensus_df, "%s_consensus_heatmap.svg"%lib)
display(Markdown("**Figure %d** Heatmap for the top %d consensus terms for **%s**. [Download figure](%s_consensus_heatmap.svg)"%(figure, top_results, lib.replace("_"," "), lib)))
figure += 1
else:
print("No terms found")
if (df.shape[1] > 0):
stackedBarPlot(df, "%s_consensus_barplot.svg"%lib)
display(Markdown("**Figure %d** Stacked bar plot for the top %d consensus terms for **%s**. [Download figure](%s_consensus_barplot.svg)"%(figure, top_results, lib.replace("_"," "), lib)))
figure +=1
else:
print("No terms found")
| appyters/Drugmonizome_Consensus_Terms/Drugmonizome-Consensus.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# # Neural Networks for Regression with TensorFlow
# > Notebook demonstrates Neural Networks for Regression Problems with TensorFlow
#
# - toc: true
# - badges: true
# - comments: true
# - categories: [DeepLearning, NeuralNetworks, TensorFlow, Python, LinearRegression]
# - image: images/nntensorflow.png
# + [markdown] id="xJ8U39gZyokt"
# ## Neural Network Regression Model with TensorFlow
# + [markdown] id="0OYbsBIE_3YM"
# This notebook is continuation of the Blog post [TensorFlow Fundamentals](https://sandeshkatakam.github.io/My-Machine_learning-Blog/tensorflow/machinelearning/2022/02/09/TensorFlow-Fundamentals.html). **The notebook is an account of my working for the Tensorflow tutorial by <NAME> on Youtube**.
# **The Notebook will cover the following concepts:**
# * Architecture of a neural network regression model.
# * Input shapes and output shapes of a regression model(features and labels).
# * Creating custom data to view and fit.
# * Steps in modelling
# * Creating a model, compiling a model, fitting a model, evaluating a model.
# * Different evaluation methods.
# * Saving and loading models.
# + [markdown] id="ANnSp9X9yxVn"
# **Regression Problems**:
# A regression problem is when the output variable is a real or continuous value, such as “salary” or “weight”. Many different models can be used, the simplest is the linear regression. It tries to fit data with the best hyper-plane which goes through the points.
# Examples:
# * How much will this house sell for?
# * How many people will buy this app?
# * How much will my health insurace be?
# * How much should I save each week for fuel?
# + [markdown] id="-E8ExcAlyxS1"
# We can also use the regression model to try and predict where the bounding boxes should be in object detection problem. Object detection thus involves both regression and then classifying the image in the box(classification problem).
#
# + [markdown] id="16bWKZ3SBBgP"
# ### Regression Inputs and outputs
#
# + [markdown] id="nfBo8NKpBxN-"
# Architecture of a regression model:
# * Hyperparameters:
# * Input Layer Shape : same as shape of number of features.
# * Hidden Layrer(s): Problem specific
# * Neurons per hidden layer : Problem specific.
# * Output layer shape: same as hape of desired prediction shape.
# * Hidden activation : Usually ReLU(rectified linear unit) sometimes sigmoid.
# * Output acitvation: None, ReLU, logistic/tanh.
# * Loss function : MSE(Mean squared error) or MAE(Mean absolute error) or combination of both.
# * Optimizer: SGD(Stochastic Gradient Descent), Adam optimizer.
#
# **Source:** Adapted from page 239 of [Hands-On Machine learning with Scikit-Learn, Keras & TensorFlow](https://www.oreilly.com/library/view/hands-on-machine-learning/9781492032632/)
#
# + [markdown] id="6u4ISCkIBxKn"
# Example of creating a sample regression model in TensorFlow:
#
# ```
# # 1. Create a model(specific to your problem)
#
# model = tf.keras.Sequential([
# tf.keras.Input(shape = (3,)),
# tf.keras.layers.Dense(100, activation = "relu"),
# tf.keras.layers.Dense(100, activation = "relu"),
# tf.keras.layers.Dense(100, activation = "relu"),
# tf.keras.layers.Dense(1, activation = None)
# ])
#
# # 2. Compile the model
#
# model.compile(loss = tf.keras.losses.mae, optimizer = tf.keras.optimizers.Adam(lr = 0.0001), metrics = ["mae"])
#
# # 3. Fit the model
#
# model.fit(X_train, Y_train, epochs = 100)
#
# ```
# + [markdown] id="czVyjJ67BxH-"
# ### Introduction to Regression with Neural Networks in TensorFlow
# + colab={"base_uri": "https://localhost:8080/"} id="qL_9ZKOJBxFt" outputId="112f3215-1d2e-498b-be2c-2d63c9552dab"
# Import TensorFlow
import tensorflow as tf
print(tf.__version__)
# + colab={"base_uri": "https://localhost:8080/", "height": 287} id="QvD60FItBxDQ" outputId="3074e91c-0756-4820-fe40-b77ed0a28e7e"
## Creating data to view and fit
import numpy as np
import matplotlib.pyplot as plt
from matplotlib import style
style.use('dark_background')
# create features
X = np.array([-7.0,-4.0,-1.0,2.0,5.0,8.0,11.0,14.0])
# Create labels
y = np.array([3.0,6.0,9.0,12.0,15.0,18.0,21.0,24.0])
# Visualize it
plt.scatter(X,y)
# + colab={"base_uri": "https://localhost:8080/"} id="VRuW86z-Bw9I" outputId="ccf2533b-9b86-4780-c25a-29fa5cb33e80"
y == X + 10
# + [markdown] id="v0fKVUJ4Jj4W"
# Yayy.. we got the relation by just seeing the data. Since the data is small and the relation ship is just linear, it was easy to guess the relation.
# + [markdown] id="qJM7kbPXMbid"
# ### Input and Output shapes
# + colab={"base_uri": "https://localhost:8080/"} id="YKfTGR08Mhhe" outputId="6db1d2af-d5e1-47f1-f60e-7d47d95e03cf"
# Create a demo tensor for the housing price prediction problem
house_info = tf.constant(["bedroom","bathroom", "garage"])
house_price = tf.constant([939700])
house_info, house_price
# + colab={"base_uri": "https://localhost:8080/"} id="LgCFCb0pNDLq" outputId="d95aff15-65f3-4d1d-a37a-5fb65b6aa776"
X[0], y[0]
# + colab={"base_uri": "https://localhost:8080/"} id="8flytFXLNFo2" outputId="45640485-21f4-45ba-ba55-d6e5970a0f57"
X[1], y[1]
# + colab={"base_uri": "https://localhost:8080/"} id="i1L_iq5yMh4c" outputId="a509171d-2e7f-42f6-d637-bc14dc255236"
input_shape = X[0].shape
output_shape = y[0].shape
input_shape, output_shape
# + colab={"base_uri": "https://localhost:8080/"} id="v8ugEWGaMh-c" outputId="8009cf97-d1b5-45f4-a3ec-584fb69e4206"
X[0].ndim
# + [markdown] id="m1ouSDrpNcCP"
# we are specifically looking at scalars here. Scalars have 0 dimension
# + colab={"base_uri": "https://localhost:8080/"} id="lirayixlNlXB" outputId="2c27a76d-1dda-4dc4-9900-82c4cc15d6fa"
# Turn our numpy arrays into tensors
X = tf.cast(tf.constant(X), dtype = tf.float32)
y = tf.cast(tf.constant(y), dtype = tf.float32)
X.shape, y.shape
# + colab={"base_uri": "https://localhost:8080/"} id="lAyfbGYfNokD" outputId="19ea277a-8a2b-4f90-ee24-d5b5f951e463"
input_shape = X[0].shape
output_shape = y[0].shape
input_shape, output_shape
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="z8X-5kRWNohl" outputId="83a0b47b-5858-4087-c563-52bebbde0e1f"
plt.scatter(X,y)
# + [markdown] id="kWoD2QiaNofG"
# ### Steps in modelling with Tensorflow
#
# 1. **Creating a model** - define the input and output layers, as well as the hidden layers of a deep learning model.
#
# 2. **Compiling a model** - define the loss function(how wrong the prediction of our model is) and the optimizer (tells our model how to improve the partterns its learning) and evaluation metrics(what we can use to interpret the performance of our model).
#
# 3. Fitting a model - letting the model try to find the patterns between X & y (features and labels).
# + colab={"base_uri": "https://localhost:8080/"} id="mX6Oh1D5SGjM" outputId="15bed3d1-5c37-4927-a6b2-ddd0c980369b"
X,y
# + colab={"base_uri": "https://localhost:8080/"} id="DhALjpKxSiC5" outputId="1cd8e077-b423-46c8-e910-a5ec2bbd3860"
X.shape
# + colab={"base_uri": "https://localhost:8080/"} id="1KKH0VkwOxyF" outputId="09ac6e76-3771-46b5-8d98-b578cf1f9ae3"
# Set random seed
tf.random.set_seed(42)
# Create a model using the Sequential API
model = tf.keras.Sequential([
tf.keras.layers.Dense(1)
])
# Compile the model
model.compile(loss=tf.keras.losses.mae, # mae is short for mean absolute error
optimizer=tf.keras.optimizers.SGD(), # SGD is short for stochastic gradient descent
metrics=["mae"])
# Fit the model
# model.fit(X, y, epochs=5) # this will break with TensorFlow 2.7.0+
model.fit(tf.expand_dims(X, axis=-1), y, epochs=5)
# + colab={"base_uri": "https://localhost:8080/"} id="AJ42qfCaQy5a" outputId="97c2ebc5-56bf-4c2c-ced0-405091434221"
# Check out X and y
X, y
# + colab={"base_uri": "https://localhost:8080/"} id="zpgxqLAxQn4q" outputId="7ba43424-1b40-4f10-8a35-eda8035044f7"
# Try and make a prediction using our model
y_pred = model.predict([17.0])
y_pred
# + [markdown] id="AEvwvmTjgAzL"
# The output is very far off from the actual value. So, Our model is not working correctly. Let's go and improve our model in the next section.
# + [markdown] id="WgO7oEeZQnz4"
# ### Improving our Model
#
# Let's take a look about the three steps when we created the above model.
#
# We can improve the model by altering the steps we took to create a model.
#
# 1. **Creating a model** - here we might add more layers, increase the number of hidden units(all called neurons) within each of the hidden layers, change the activation function of each layer.
#
# 2. **Compiling a model** - here we might change the optimization function or perhaps the learning rate of the optimization function.
#
# 3. **Fitting a model** - here we might fit a model for more **epochs** (leave it for training longer) or on more data (give the model more examples to learn from)
#
# + colab={"base_uri": "https://localhost:8080/"} id="Vj9qGFDAQnxn" outputId="27c29447-bff7-4031-b64a-cf03238c4f2d"
# Let's rebuild our model with change in the epoch number
# 1. Create the model
model = tf.keras.Sequential([
tf.keras.layers.Dense(1)
])
# 2. Compile the model
model.compile(loss = tf.keras.losses.mae,
optimizer = tf.keras.optimizers.SGD(),
metrics = ["mae"])
# 3. Fit the model to our dataset
model.fit(tf.expand_dims(X, axis=-1), y, epochs=100, verbose = 0)
# + colab={"base_uri": "https://localhost:8080/"} id="rhcp3a9c3OYR" outputId="395cc812-b9b5-46eb-e3cd-63a97a2cd1c0"
# Our data
X , y
# + colab={"base_uri": "https://localhost:8080/"} id="NGeJdiXn3h0i" outputId="05b8de25-d5b8-42bd-a323-bf3f4872df3d"
# Let's see if our model's prediction has improved
model.predict([17.0])
# + [markdown] id="GFmSn7VY3rsM"
# We got so close the actual value is 27 we performed a better prediction than the last model we trained. But we need to improve much better.
# Let's see what more we change and how close can we get to our actual output
# + colab={"base_uri": "https://localhost:8080/"} id="TlmQFvsC4L1b" outputId="d5c971c3-0d68-4349-db64-c1ec63bbfbc5"
# Let's rebuild our model with changing the optimization function to Adam
# 1. Create the model
model = tf.keras.Sequential([
tf.keras.layers.Dense(1)
])
# 2. Compile the model
model.compile(loss = tf.keras.losses.mae,
optimizer = tf.keras.optimizers.Adam(lr = 0.0001), # lr stands for learning rate
metrics = ["mae"])
# 3. Fit the model to our dataset
model.fit(tf.expand_dims(X, axis=-1), y, epochs=100, verbose = 0)
# + colab={"base_uri": "https://localhost:8080/"} id="Pj2U8Kbr4L8-" outputId="7601fa2a-886f-401b-ab18-a03b7a3ebd24"
# Prediction of our newly trained model:
model.predict([17.0]) # we are going to predict for the same input value 17
# + [markdown] id="BGJtW6bk4MDL"
# Oh..god!! This result went really bad for us.
# + colab={"base_uri": "https://localhost:8080/"} id="HBKbv1MO5Eya" outputId="c9bbb271-7159-404d-e80a-78851814bbca"
# Let's rebuild our model by adding one extra hidden layer with 100 units
# 1. Create the model
model = tf.keras.Sequential([
tf.keras.layers.Dense(100, activation = "relu"), # only difference we made
tf.keras.layers.Dense(1)
])
# 2. Compile the model
model.compile(loss = "mae",
optimizer = tf.keras.optimizers.SGD(),
metrics = ["mae"])
# 3. Fit the model to our dataset
model.fit(tf.expand_dims(X, axis=-1), y, epochs=100, verbose = 0) # verbose will hide the output from epochs
# + colab={"base_uri": "https://localhost:8080/"} id="FZ7WgIEv56-Y" outputId="2e0d2bda-ce7d-44d6-a4c0-e54ccfe74324"
X , y
# + colab={"base_uri": "https://localhost:8080/"} id="rDEiL5Ji5Ev_" outputId="7e3f17df-ea2e-4b89-c895-d57c43b3bb2f"
# It's prediction time!
model.predict([17.0])
# + [markdown] id="09bdyxhC5EtX"
# Oh, this should be 27 but this prediction is very far off from our previous prediction.
# It seems that our previous model did better than this.
#
# Even though we find the values of our loss function are very low than that of our previous model. We still are far away from our label value.
# **Why is that so??**
# The explanation is our model is overfitting the dataset. That means it is trying to map a function that just fits the already provided examples correctly but it cannot fit the new examples that we are giving.
# So, the `mae` and `loss value` if not the ultimate metric to check for improving the model. because we need to get less error for new examples that the model has not seen before.
#
# + colab={"base_uri": "https://localhost:8080/"} id="mftXMLwG5Emc" outputId="c751a1b7-9086-4a27-e3ab-712162781078"
# Let's rebuild our model by using Adam optimizer
# 1. Create the model
model = tf.keras.Sequential([
tf.keras.layers.Dense(100, activation = "relu"), # only difference we made
tf.keras.layers.Dense(1)
])
# 2. Compile the model
model.compile(loss = "mae",
optimizer = tf.keras.optimizers.Adam(),
metrics = ["mae"])
# 3. Fit the model to our dataset
model.fit(tf.expand_dims(X, axis=-1), y, epochs=100, verbose = 0)# verbose will hide the epochs output
# + colab={"base_uri": "https://localhost:8080/"} id="Dcdpd0ih5Ekh" outputId="6179607d-7fbf-465e-ed5e-ed3edb86d80a"
model.predict([17.0])
# + [markdown] id="CEaaso5o7qUH"
# Still not better!!
# + colab={"base_uri": "https://localhost:8080/"} id="wTZtdEl27xCw" outputId="30943b3b-f8e5-4736-b83e-9ea34376c1fb"
# Let's rebuild our model by adding more layers
# 1. Create the model
model = tf.keras.Sequential([
tf.keras.layers.Dense(100, activation = "relu"),
tf.keras.layers.Dense(100, activation = "relu"),
tf.keras.layers.Dense(100, activation = "relu"),# only difference we made
tf.keras.layers.Dense(1)
])
# default value of lr is 0.001
# 2. Compile the model
model.compile(loss = "mae",
optimizer = tf.keras.optimizers.Adam(lr = 0.01), # lr stands for learning rate
metrics = ["mae"])
# 3. Fit the model to our dataset
model.fit(tf.expand_dims(X, axis=-1), y, epochs=100, verbose = 0) # verbose will hide the epochs output
# + [markdown] id="ME2Z56_k7xIs"
# The learning rate is the most important hyperparameter for all the Neural Networks
# + [markdown] id="_VFlk7eY7xOD"
# ### Evaluating our model
#
# In practice, a typical workflow you'll go through when building a neural network is:
#
# ```
# Build a model -> fit it -> evaluate it -> tweak a model -> fit it -> evaluate it -> tweak it -> fit it
# ```
# Common ways to improve a deep model:
# * Adding Layers
# * Increase the number of hidden units
# * Change the activation functions
# * Change the optimization function
# * Change the learning rate
# * Fitting on more data
# * Train for longer (more epochs)
#
# **Because we can alter each of these they are called hyperparameters**
# + [markdown] id="Y6eiPEWF8hzr"
# When it comes to evaluation.. there are 3 words you should memorize:
#
# > "Visualize, Visualize, Visualize"
#
# It's a good idea to visualize:
# * The data - what data are working with? What does it look like
# * The model itself - What does our model look like?
# * The training of a model - how does a model perform while it learns?
# * The predictions of the model - how does the prediction of the model line up against the labels(original value)
#
# + colab={"base_uri": "https://localhost:8080/"} id="dKkinwK-8hwo" outputId="59976505-7778-4225-bfa6-05dc4d31cf49"
# Make a bigger dataset
X_large = tf.range(-100,100,4)
X_large
# + colab={"base_uri": "https://localhost:8080/"} id="tmJs64Sb8huM" outputId="b0959bee-b2dc-4caf-a1ae-ca997c81c241"
y_large = X_large + 10
y_large
# + colab={"base_uri": "https://localhost:8080/", "height": 283} id="3qQq7v8p8hrc" outputId="8c50a653-05d2-4cad-edb3-db95d781cbe4"
import matplotlib.pyplot as plt
plt.scatter(X_large,y_large)
# + [markdown] id="6AhCjZ2Z8hoy"
# ### The 3 sets ...
#
# * **Training set** - The model learns from this data, which is typically 70-80% of the total data you have available.
#
# * **validation set** - The model gets tuned on this data, which is typically 10-15% of the data avaialable.
#
# * **Test set** - The model gets evaluated on this data to test what it has learned. This set is typically 10-15%.
#
# + colab={"base_uri": "https://localhost:8080/"} id="7b6UMaLA7xUp" outputId="8c398b2c-8eef-4c39-81b0-946c64270911"
# Check the length of how many samples we have
len(X_large)
# + colab={"base_uri": "https://localhost:8080/"} id="qDH1qyt7AfCL" outputId="e0691618-ed0c-4f21-d3e0-d13212d77358"
# split the data into train and test sets
# since the dataset is small we can skip the valdation set
X_train = X_large[:40]
X_test = X_large[40:]
y_train = y_large[:40]
y_test = y_large[40:]
len(X_train), len(X_test), len(y_train), len(y_test)
# + [markdown] id="sgQo2MWvBF2G"
# ### Visualizing the data
#
# Now we've got our data in training and test sets. Let's visualize it.
# + colab={"base_uri": "https://localhost:8080/", "height": 428} id="wUmGlF3jBb9S" outputId="d00ec668-0d90-47ac-fbcb-9ddc6aad3de6"
plt.figure(figsize = (10,7))
# Plot the training data in blue
plt.scatter(X_train, y_train, c= 'b', label = "Training data")
# Plot the test data in green
plt.scatter(X_test, y_test, c = "g", label = "Training data")
plt.legend();
# + id="GEYDrk10CimF"
# Let's have a look at how to build neural network for our data
# 1. Create the model
model = tf.keras.Sequential([
tf.keras.layers.Dense(1)
])
# default value of lr is 0.001
# 2. Compile the model
model.compile(loss = "mae",
optimizer = tf.keras.optimizers.SGD(), # lr stands for learning rate
metrics = ["mae"])
# 3. Fit the model to our dataset
#model.fit(tf.expand_dims(X_train, axis=-1), y_train, epochs=100)
# + [markdown] id="dkFPxPz2C_tL"
# Let's visualize it before fitting the model
# + colab={"base_uri": "https://localhost:8080/", "height": 288} id="HD22ognkDtvA" outputId="a46d526a-081e-4da8-a280-fec8c1f34731"
model.summary()
# + [markdown] id="riOKDo-0DElv"
# model.summary() doesn't work without building the model or fitting the model
# + colab={"base_uri": "https://localhost:8080/"} id="w5VweDySDod7" outputId="22379507-8978-41cf-9f89-31037d6f32de"
X[0], y[0]
# + id="Bq6XiEZnDE3w"
# Let's create a model which builds automatically by defining the input_shape arguments
tf.random.set_seed(42)
# Create a model(same as above)
model = tf.keras.Sequential([
tf.keras.layers.Dense(1, input_shape = [1]) # input_shape is 1 refer above code cell
])
# Compile the model
model.compile(loss= "mae",
optimizer = tf.keras.optimizers.SGD(),
metrics = ["mae"])
# + colab={"base_uri": "https://localhost:8080/"} id="wvUmRiuOEF2Q" outputId="ed9dc6f1-4aec-4b5f-815b-106a7247bcbb"
model.summary()
# + [markdown] id="5HXGJdgbEFz9"
# * **Total params** - total number of parameters in the model.
# * **Trainable parameters**- these are the parameters (patterns) the model can update as it trains.
# * **Non-Trainable parameters** - these parameters aren't updated during training(this is typical when you have paramters from other models during **transfer learning**)
# + id="fJl20PEXEFxS"
# Let's have a look at how to build neural network for our data
# 1. Create the model
model = tf.keras.Sequential([
tf.keras.layers.Dense(10, input_shape = [1], name= "input_layer"),
tf.keras.layers.Dense(1, name = "output_layer")
], name = "model_1")
# 2. Compile the model
model.compile(loss = "mae",
optimizer = tf.keras.optimizers.SGD(), # lr stands for learning rate
metrics = ["mae"])
# + id="S2pph2l4EFX_" colab={"base_uri": "https://localhost:8080/"} outputId="05cf03b2-d4cf-4671-e724-dedcbc5eb935"
model.summary()
# + [markdown] id="1Gtqtj6pI08o"
# We have changed the layer names and added our custom model name.
# + id="qg_zkUxyEFVi" colab={"base_uri": "https://localhost:8080/", "height": 312} outputId="99f6c155-3f0c-4859-f466-4fda0dad63ba"
from tensorflow.keras.utils import plot_model
plot_model(model = model, to_file = 'model1.png', show_shapes = True)
# + colab={"base_uri": "https://localhost:8080/", "height": 236} id="SWM3DMD9Bb6d" outputId="1f78baf3-eb1e-4c71-bdd4-d182c6e833ee"
# Let's have a look at how to build neural network for our data
# 1. Create the model
model = tf.keras.Sequential([
tf.keras.layers.Dense(100, activation = "relu"),
tf.keras.layers.Dense(100, activation = "relu"),
tf.keras.layers.Dense(100, activation = "relu"),# only difference we made
tf.keras.layers.Dense(1)
], name)
# default value of lr is 0.001
# 2. Compile the model
model.compile(loss = "mae",
optimizer = tf.keras.optimizers.Adam(lr = 0.01), # lr stands for learning rate
metrics = ["mae"])
# 3. Fit the model to our dataset
model.fit(tf.expand_dims(X_train, axis=-1), y_train, epochs=100, verbose = 0)
# + colab={"base_uri": "https://localhost:8080/"} id="Qh-xet_iBb4J" outputId="0fa0b7f7-d2cf-45e0-9ee9-f112452b8045"
model.predict(X_test)
# + [markdown] id="bB45dBitBb2B"
# wow, we are so close!!!
# + id="YvB6K2QXBbzK" colab={"base_uri": "https://localhost:8080/"} outputId="8b9b2fcb-f213-45df-d188-6c4e89153f6f"
model.summary()
# + id="Z571FPj4Bbww" colab={"base_uri": "https://localhost:8080/", "height": 312} outputId="224f72b7-8074-494b-904b-dfbdff3ad455"
from tensorflow.keras.utils import plot_model
plot_model(model = model, to_file = 'model.png', show_shapes = True)
# + [markdown] id="78BTt4eoBbuz"
# ### Visualizing our model's predictions
#
# To visualize predictions, it's a good idea to plot them against the ground truth labels.
#
# Often you'll see this in the form of `y_test` or `y_true` versus `y_pred`
# + id="RTTUIToTNxbV"
# Set random seed
tf.random.set_seed(42)
# Create a model (same as above)
model = tf.keras.Sequential([
tf.keras.layers.Dense(10, input_shape = [1], name = "input_layer"),
tf.keras.layers.Dense(1, name = "output_layer") # define the input_shape to our model
], name = "revised_model_1")
# Compile model (same as above)
model.compile(loss=tf.keras.losses.mae,
optimizer=tf.keras.optimizers.SGD(),
metrics=["mae"])
# + colab={"base_uri": "https://localhost:8080/"} id="frX0VbrWNxZD" outputId="e918288c-367a-4f04-cb83-32c2fcb2d1d9"
model.summary()
# + colab={"base_uri": "https://localhost:8080/"} id="mDiGxRlgNxWT" outputId="991add0b-3d51-4205-fc83-61425265f059"
model.fit(X_train, y_train, epochs=100, verbose=0)
# + colab={"base_uri": "https://localhost:8080/"} id="KPDQV8qRNxT-" outputId="286f6978-6acf-4e51-998f-007ba562d0be"
model.summary()
# + id="FzndWm_sBbrd" colab={"base_uri": "https://localhost:8080/"} outputId="6f4d0a52-4b38-49a1-8215-cd9105012bdc"
# Make some predictions
y_pred = model.predict(X_test)
tf.constant(y_pred)
# + [markdown] id="ryr1OIUcKO86"
# These are our predictions!
# + id="lpYAC6aPBbpN" colab={"base_uri": "https://localhost:8080/"} outputId="8d7a7944-08d9-401d-a544-ffd5c89c64a0"
y_test
# + [markdown] id="PqdZW6JFKS2W"
# These are the ground truth labels!
# + colab={"base_uri": "https://localhost:8080/", "height": 312} id="Q64Lq1OJOAkm" outputId="cd6975fb-672a-4d62-ee32-e0d0c7d2fccd"
plot_model(model, show_shapes=True)
# + [markdown] id="Iv-VrZlxKYOu"
# **Note:** IF you feel like you're going to reuse some kind of functionality in future,
# it's a good idea to define a function so that we can reuse it whenever we need.
# + id="xbnB1aGcKYMV"
#Let's create a plotting function
def plot_predictions(train_data= X_train,
train_labels = y_train,
test_data = X_test,
test_labels =y_test,
predictions = y_pred):
"""
Plots training data, test data and compares predictions to ground truth labels
"""
plt.figure(figsize = (10,7))
# Plot training data in blue
plt.scatter(train_data, train_labels, c= "b", label = "Training data")
# Plot testing data in green
plt.scatter(test_data, test_labels, c= "g", label = "Testing data")
# Plot model's predictions in red
plt.scatter(test_data, predictions, c= "r", label = "Predictions")
# Show legends
plt.legend();
# + colab={"base_uri": "https://localhost:8080/", "height": 428} id="wiEDqUVvKYJy" outputId="53e56e51-8fbc-4758-a94f-9269733c2139"
plot_predictions(train_data=X_train,
train_labels=y_train,
test_data=X_test,
test_labels=y_test,
predictions=y_pred)
# + [markdown] id="vS-qUdKZPPyG"
# We tuned our model very well this time. The predictions are really close to the actual values.
# + [markdown] id="Nm1812GKKM6k"
# ### Evaluating our model's predictions with regression evaluation metrics
#
# Depending on the problem you're working on, there will be different evaluation metrics to evaluate your model's performance.
#
# Since, we're working on a regression, two of the main metrics:
#
# * **MAE** - mean absolute error, "on average, how wrong id each of my model's predictions"
# * TensorFlow code: `tf.keras.losses.MAE()`
# * or `tf.metrics.mean_absolute_error()`
# $$ MAE = \frac{Σ_{i=1}^{n} |y_i - x_i| }{n} $$
# * **MSE** - mean square error, "square of the average errors"
# * `tf.keras.losses.MSE()`
# * `tf.metrics.mean_square_error()`
# $$ MSE = \frac{1}{n} Σ_{i=1}^{n}(Y_i - \hat{Y_i})^2$$
#
# $\hat{Y_i}$ is the prediction our model makes.
# $Y_i$ is the label value.
#
# * **Huber** - Combination of MSE and MAE, Less sensitive to outliers than MSE.
# * `tf.keras.losses.Huber()`
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="ynQfcSZMPaAE" outputId="20a7792c-f2b1-4f78-94b7-47ccfa4ded13"
# Evaluate the model on test set
model.evaluate(X_test, y_test)
# + colab={"base_uri": "https://localhost:8080/"} id="EqBHYGgIcdTW" outputId="4787381b-ac3d-4222-c7a8-f7d84a579c3c"
# calculate the mean absolute error
mae = tf.metrics.mean_absolute_error(y_true = y_test,
y_pred = tf.constant(y_pred))
mae
# + [markdown] id="2V6DO4_1d-NX"
# We got the metric values wrong..why did this happen??
# + colab={"base_uri": "https://localhost:8080/"} id="qGuglu_EPaGF" outputId="9fc2a28a-4574-4d24-b14c-50e899b96ea4"
tf.constant(y_pred)
# + colab={"base_uri": "https://localhost:8080/"} id="d6RH6OCjPaLa" outputId="424455fc-81e0-4b57-f5de-0292676708d6"
y_test
# + [markdown] id="4UIeXSPcdeSn"
# Notice that the shape of `y_pred` is (10,1) and the shape of `y_test` is (10,)
# They might seem the same but they are not of the same shape.
# Let's reshape the tensor to make the shapes equal.
# + colab={"base_uri": "https://localhost:8080/"} id="f7g-tuSRdy65" outputId="abaa0043-ec88-4590-862f-d40a62188f52"
tf.squeeze(y_pred)
# + colab={"base_uri": "https://localhost:8080/"} id="NeIz5ShCcb_t" outputId="50ab40c8-11dd-481b-85ee-53fcee2e0a8f"
# Calculate the mean absolute error
mae = tf.metrics.mean_absolute_error(y_true = y_test,
y_pred = tf.squeeze(y_pred))
mae
# + [markdown] id="i-KsRsLVeS1E"
# Now,we got our metric value. The mean absolute error of our model is 3.1969407.
# Now, let's calculate the mean squared error and see how that goes.
# + colab={"base_uri": "https://localhost:8080/"} id="VCJvzzGSeb0y" outputId="1a48d927-d04f-45a7-bbcd-06f2249c40ad"
# Calculate the mean squared error
mse = tf.metrics.mean_squared_error(y_true = y_test,
y_pred = tf.squeeze(y_pred))
mse
# + [markdown] id="W50t9j0tebyK"
# Our mean squared error is 13.070143. Remember, the mean squared error squares the error for every example in the test set and averages the values. So, generally, the mse is largeer than mae.
# When larger errors are more significant than smaller errors, then it is best to use mse.
# MAE can be used as a great starter metric for any regression problem.
# We can also try Huber and see how that goes.
# + colab={"base_uri": "https://localhost:8080/"} id="gis6u8heebv4" outputId="a600b96a-1653-4895-e7e8-8d83a132bb55"
# Calculate the Huber metric for our model
huber_metric = tf.losses.huber(y_true = y_test,
y_pred = tf.squeeze(y_pred))
huber_metric
# + id="ZaR4JBTTf2fs"
# Make some functions to reuse MAE and MSE and also Huber
def mae(y_true, y_pred):
return tf.metrics.mean_absolute_error(y_true = y_test,
y_pred = tf.squeeze(y_pred))
def mse(y_true, y_pred):
return tf.metrics.mean_squared_error(y_true = y_test,
y_pred = tf.squeeze(y_pred))
def huber(y_true, y_pred):
return tf.losses.huber(y_true = y_test,
y_pred = tf.squeeze(y_pred))
# + [markdown] id="hVNQgNsCebtO"
# ### Running experiments to improve our model
#
# ```
# Build a model -> fit it -> evaluate it -> tweak a model -> fit it -> evaluate it -> tweak it -> fit it
# ```
#
# 1. Get more data - get more examples for your model to train on(more oppurtunities to learn patterns or relationships between features and labels).
# 2. Make your mode larger(using a more complex model) - this might come in the form of more layeres or more hidden unites in each layer.
# 3. Train for longer - give your model more of a chance to find patterns in the data.
#
# Let's do a few modelling experiments:
# 1. `model_1` - same as original model, 1 layer, trained for 100 epochs.
# 2. `model_2` - 2 layers, trained for 100 epochs
# 3. `model_3` - 2 layers, trained for 500 epochs.
#
# You can design more experiments too to make the model more better
#
#
#
# **Build `Model_1`**
# + colab={"base_uri": "https://localhost:8080/"} id="TX_BcAQtiIma" outputId="8707eb53-06da-4cf0-e189-6aa585df95ac"
X_train, y_train
# + colab={"base_uri": "https://localhost:8080/"} id="dQc0qo_QiL18" outputId="2effd38a-bfc1-424a-b40c-14bbe4abd0db"
# Set random seed
tf.random.set_seed(42)
# 1. Create the model
model_1 = tf.keras.Sequential([
tf.keras.layers.Dense(1, input_shape = [1])
], name = "Model_1")
# 2. Compile the model
model_1.compile(loss = tf.keras.losses.mae,
optimizer = tf.keras.optimizers.SGD(),
metrics = ["mae"])
# 3. Fit the model
model_1.fit(X_train, y_train ,epochs = 100, verbose = 0)
# + colab={"base_uri": "https://localhost:8080/"} id="y6sKm8rEnbt7" outputId="c68480f9-cc61-43a4-f0b5-0e397bd377c4"
model_1.summary()
# + colab={"base_uri": "https://localhost:8080/", "height": 428} id="UeMx0tvxjUKd" outputId="fa6a9e57-fcf7-44fb-acc0-f27665735700"
# Make and plot the predictions for model_1
y_preds_1 = model_1.predict(X_test)
plot_predictions(predictions = y_preds_1)
# + colab={"base_uri": "https://localhost:8080/"} id="_xB_iZGukBON" outputId="10d097a4-4994-4c50-9c5a-936901669eba"
# Calculate model_1 evaluation metrics
mae_1 = mae(y_test, y_preds_1)
mse_1 = mse(y_test, y_preds_1)
mae_1, mse_1
# + [markdown] id="vH7Z1uZakkAU"
# **Build `Model_2`**
#
# * 2 dense layers, trained for 100 epochs
# + colab={"base_uri": "https://localhost:8080/"} id="fXkZAV43k58t" outputId="5fd10ef7-957c-4b10-b75b-67281e0c9037"
# Set random seed
tf.random.set_seed(42)
# 1. Create the model
model_2 = tf.keras.Sequential([
tf.keras.layers.Dense(10, input_shape =[1]),
tf.keras.layers.Dense(1)
], name = "model_2")
# 2. Compile the model
model_2.compile(loss = tf.keras.losses.mae,
optimizer = tf.keras.optimizers.SGD(),
metrics = ["mse"]) # Let's build this model with mse as eval metric.
# 3. Fit the model
model_2.fit(X_train, y_train ,epochs = 100, verbose = 0)
# + colab={"base_uri": "https://localhost:8080/"} id="MBwWqPdUlx2z" outputId="e6024c7d-8722-4db0-d255-97c0ec2869a8"
model_2.summary()
# + colab={"base_uri": "https://localhost:8080/", "height": 428} id="dyMYjrO5l0Xc" outputId="ba7b5df0-caed-49b0-ceee-773e24b4bc0e"
# Make and plot predictions of model_2
y_preds_2 = model_2.predict(X_test)
plot_predictions(predictions = y_preds_2)
# + [markdown] id="6kErmqkXmAEd"
# Yeah,we improved this model very much than the previous one.
# If you want to compare with previous one..scroll up and see the plot_predictions of
# previous one and compare it with this one.
# + colab={"base_uri": "https://localhost:8080/"} id="sd-XMSHymt5U" outputId="4e435e77-3bf1-431e-835b-95d066c49702"
# Calculate the model_2 evaluation metrics
mae_2 = mae(y_test, y_preds_2)
mse_2 = mse(y_test, y_preds_2)
mae_2, mse_2
# + [markdown] id="ri0xXdlXm8-s"
# **Build `Model_3`**
#
# * 2 layers, trained for 500 epochs
# + colab={"base_uri": "https://localhost:8080/"} id="3TlZzf93nIor" outputId="89c947c4-9498-458c-a66e-35449ee51354"
# Set random seed
tf.random.set_seed(42)
# 1. Create the model
model_3 = tf.keras.Sequential([
tf.keras.layers.Dense(10, input_shape =[1]),
tf.keras.layers.Dense(1)
], name = "model_3")
# 2. Compile the model
model_3.compile(loss = tf.keras.losses.mae,
optimizer = tf.keras.optimizers.SGD(),
metrics = ["mae"]) # Let's build this model with mse as eval metric.
# 3. Fit the model
model_2.fit(X_train, y_train ,epochs = 500, verbose = 0)
# + colab={"base_uri": "https://localhost:8080/", "height": 428} id="nICFgtpdnjgM" outputId="d1024a8f-57db-40d6-e9de-f8472dadb63a"
# Make and plot some predictions
y_preds_3 = model_3.predict(X_test)
plot_predictions(predictions = y_preds_3)
# + [markdown] id="x4FZmqhUn2-t"
# This is even terrible performance than the first model. we have actually made the model worse. WHY??
#
# We, overfitted the model too much because we trained it for much longer than we are supposed to.
# + colab={"base_uri": "https://localhost:8080/"} id="Hyfs1MgIoBO7" outputId="83144c31-d902-4d51-9523-8ef7ee718291"
# Calculate the model_3 evaluation metrics
mae_3 = mae(y_test, y_preds_3)
mse_3 = mse(y_test, y_preds_3)
mae_3, mse_3
# + [markdown] id="qO916PjgoXWk"
# whoaa, the error is extremely high. I think the best of our models is `model_2`
# + [markdown] id="R0d2altrogLk"
# The Machine Learning practitioner's motto:
#
# `Experiment, experiment, experiment`
# + [markdown] id="sF9hW503o-U7"
# **Note:** You want to start with small experiments(small models) and make sure they work and then increase their scale when neccessary.
# + [markdown] id="l27fhsBoogFy"
# ### Comparing the results of our experiments
# We've run a few experiments, let's compare the results now.
# + colab={"base_uri": "https://localhost:8080/", "height": 143} id="y9dY0p8NogC-" outputId="1b9b6fb3-9bf5-425b-e0d1-ed16ee0e8a50"
# Let's compare our models'c results using pandas dataframe:
import pandas as pd
model_results = [["model_1", mae_1.numpy(), mse_1.numpy()],
["model_2", mae_2.numpy(), mse_2.numpy()],
["model_3", mae_3.numpy(), mse_3.numpy()]]
all_results = pd.DataFrame(model_results, columns =["model", "mae", "mse"])
all_results
# + [markdown] id="4sLfbMRpp_10"
# It looks like model_2 performed done the best. Let's look at what is model_2
# + colab={"base_uri": "https://localhost:8080/"} id="_wE7lfkwqPcA" outputId="113d45d0-4791-4c27-e5f0-3146697bedf0"
model_2.summary()
# + [markdown] id="ZPC8ozETqPjN"
# This is the model that has done the best on our dataset.
# + [markdown] id="3dpYTBNDqPoN"
# **Note:** One of your main goals should be to minimize the time between your experiments. The more experiments you do, the more things you will figure out which don't work and in turn, get closer to figuring out what does work. Remeber, the machine learning pracitioner's motto : "experiment, experiment, experiment".
# + [markdown] id="t6z_prWIq0yq"
# ## Tracking your experiments:
#
# One really good habit of machine learning modelling is to track the results of your experiments.
#
# And when doing so, it can be tedious if you are running lots of experiments.
#
# Luckily, there are tools to help us!
#
# **Resources:** As you build more models, you'll want to look into using:
#
# * TensorBoard - a component of TensorFlow library to help track modelling experiments. It is integrated into the TensorFlow library.
#
# * Weights & Biases - A tool for tracking all kinds of machine learning experiments (it plugs straight into tensorboard).
# + [markdown] id="c3UeCEVDsT22"
# ## Saving our models
#
# Saving our models allows us to use them outside of Google Colab(or wherever they were trained) such as in a web application or a mobile app.
#
# There are two main formats we can save our model:
#
# 1. The SavedModel format
# 2. The HDF5 format
# + [markdown] id="Kw-ztpAYuxyb"
# `model.save()` allows us to save the model and we can use it again to do add things to the model after reloading it.
# + colab={"base_uri": "https://localhost:8080/"} id="WVmmA7sAslYw" outputId="d863426a-0c5f-4816-fea2-0ca9b82fbe29"
# Save model using savedmodel format
model_2.save("best_model_SavedModel_format")
# + [markdown] id="LWHI-OjEufw8"
# If we are planning to use this model inside the tensorflow framework. we will be better off using the `SavedModel` format. But if we are planning to export the model else where and use it outside the tensorflow framework use the HDF5 format.
# + id="BXDVAL3FslHj"
# Save model using HDF5 format
model_2.save("best_model_HDF5_format.h5")
# + [markdown] id="G26eDi2EvMvN"
# Saving a model with SavedModel format will give us a folder with some files regarding our model.
# Saving a model with HDF5 format will give us just one file with our model.
# + [markdown] id="dqPLapY-vHb1"
# ### Loading in a saved model
# + colab={"base_uri": "https://localhost:8080/"} id="27gq-doFwN0_" outputId="6bb8e7c2-2773-4e34-daae-97cfe8dc3500"
# Load in the SavedModel format model
loaded_SavedModel_format = tf.keras.models.load_model("/content/best_model_SavedModel_format")
loaded_SavedModel_format.summary()
# + colab={"base_uri": "https://localhost:8080/"} id="Girv6tubwOGl" outputId="1a7bc379-78ed-43a3-c145-c1efcb4470b5"
# Let's check is that the same thing as model_2
model_2.summary()
# + colab={"base_uri": "https://localhost:8080/"} id="UR27o5jxwOLk" outputId="5fabd27c-8b6b-406a-e0b8-2d552a3b6021"
# Compare the model_2 predictions with SavedModel format model predictions
model_2_preds = model_2.predict(X_test)
loaded_SavedModel_format_preds = loaded_SavedModel_format.predict(X_test)
model_2_preds == loaded_SavedModel_format_preds
# + colab={"base_uri": "https://localhost:8080/"} id="JRMvOEB9wOQl" outputId="5f00d5d3-c3ee-48e4-b2ce-f8738be74808"
mae(y_true = y_test, y_pred = model_2_preds) == mae(y_true = y_test, y_pred = loaded_SavedModel_format_preds)
# + colab={"base_uri": "https://localhost:8080/"} id="3gE17p40xx1F" outputId="cc7479c9-0a42-4807-a1a8-f6596c3d0c20"
# Load in a model using the .hf format
loaded_h5_model = tf.keras.models.load_model("/content/best_model_HDF5_format.h5")
loaded_h5_model.summary()
# + colab={"base_uri": "https://localhost:8080/"} id="eSLWsCJXyUzP" outputId="e2b1fec7-251d-4fa6-e9fc-a87bbf204cea"
model_2.summary()
# + [markdown] id="IJPRNyBRyXCE"
# Yeah the loading of .hf format model matched with our original mode_2 format.
# So, our model loading worked correctly.
# + colab={"base_uri": "https://localhost:8080/"} id="kKyVyssLyhk1" outputId="1e73ec49-6eaf-490b-c833-01723788381f"
# Check to see if loaded .hf model predictions match model_2
model_2_preds = model_2.predict(X_test)
loaded_h5_model_preds = loaded_h5_model.predict(X_test)
model_2_preds == loaded_h5_model_preds
# + [markdown] id="MiO4g2Fey0At"
# ### Download a model(or any other file) from google colab
#
# If you want to download your files from Google Colab:
#
# 1. you can go to the files tab and right click on the file you're after and click download.
#
# 2. Use code(see the cell below).
#
# 3. You can save it to google drive by connecting to google drive and copying it there.
#
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="fS_BhNylzwcN" outputId="fb713b82-390f-43a9-88c8-fbc6227bf36a"
# Download a file from Google Colab
from google.colab import files
files.download("/content/best_model_HDF5_format.h5")
# + id="OQjab5s4z7Ld"
# Save a file from Google Colab to Google Drive(requires mounting google drive)
# !cp /content/best_model_HDF5_format.h5 /content/drive/MyDrive/tensor-flow-deep-learning
# + colab={"base_uri": "https://localhost:8080/"} id="97zV7k0N0rAF" outputId="cd59f693-555e-493b-d2a4-fc3a14f8589c"
# !ls /content/drive/MyDrive/tensor-flow-deep-learning
# + [markdown] id="Jg-bIT5V1DoV"
# We have saved our model to our google drive !!!
# + [markdown] id="QY7iwl6X1Inl"
# ## A larger example
#
# We take a larger dataset to do create a regression model. The model we do is insurance forecast by using linear regression available from kaggle [Medical Cost Personal Datasets](https://www.kaggle.com/mirichoi0218/insurance)
# + id="kbMhE8RP1UF2"
# Import required libraries
import tensorflow as tf
import pandas as pd
import matplotlib.pyplot as plt
# + colab={"base_uri": "https://localhost:8080/", "height": 423} id="rlo7RC541UL3" outputId="ce258aef-1a64-472d-de03-2ba2db98275e"
# Read in the insurance data set
insurance = pd.read_csv("https://raw.githubusercontent.com/stedy/Machine-Learning-with-R-datasets/master/insurance.csv")
insurance
# + [markdown] id="vzY7jqM52UwR"
# This is a quite bigger dataset than the one we have previously worked with.
# + colab={"base_uri": "https://localhost:8080/", "height": 270} id="8VZKvJRAEsb-" outputId="dc2f7704-d268-44d9-ea84-23017a23d75a"
# one hot encoding on a pandas dataframe
insurance_one_hot = pd.get_dummies(insurance)
insurance_one_hot.head()
# + id="DLDYcJDREsZI"
# Create X & y values (features and labels)
X = insurance_one_hot.drop("charges", axis =1)
y = insurance_one_hot["charges"]
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="spUY12S1M3i6" outputId="c5be1144-2dda-4305-b678-ae476015048a"
# View X
X.head()
# + colab={"base_uri": "https://localhost:8080/"} id="ONex7TjxM8gW" outputId="aab2dab3-be32-4ffe-9803-365f024bf07a"
# View y
y.head()
# + colab={"base_uri": "https://localhost:8080/"} id="cyyDEgjkFwgz" outputId="47f19794-fe14-4864-f992-b4d65179b84b"
# Create training and test sets
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y,test_size = 0.2, random_state = 42)
len(X), len(X_train), len(X_test)
# + colab={"base_uri": "https://localhost:8080/", "height": 423} id="4O8YwaKbEr0H" outputId="30d10108-b5d4-4385-999e-6cf7e4d50b12"
X_train
# + colab={"base_uri": "https://localhost:8080/"} id="RmMaGlwrD6YW" outputId="9e189bb5-a820-41f9-ad15-2bdef7e2d8aa"
insurance["smoker"] , insurance["sex"]
# + colab={"base_uri": "https://localhost:8080/"} id="u7A6b3vfD6TA" outputId="a5a2ee56-2f91-48c6-c9f0-9f4215fe73bb"
# Build a neural network (sort of like model_2 above)
tf.random.set_seed(42)
# 1. Create a model
insurance_model = tf.keras.Sequential([
tf.keras.layers.Dense(10),
tf.keras.layers.Dense(1)
])
# 2. Compile the model
insurance_model.compile(loss = tf.keras.losses.mae,
optimizer = tf.keras.optimizers.SGD(),
metrics = ["mae"])
#3. Fit the model
insurance_model.fit(X_train, y_train,epochs = 100, verbose = 0)
# + colab={"base_uri": "https://localhost:8080/"} id="I5SXUKXcD6Kw" outputId="4a708449-d000-4806-c3bf-1c134b1e1df4"
# Check the results of the insurance model on the test data
insurance_model.evaluate(X_test,y_test)
# + colab={"base_uri": "https://localhost:8080/"} id="tUCGxlQi2Upc" outputId="6dbb1059-637f-42c7-f966-87693f4d1c51"
y_train.median(), y_train.mean()
# + [markdown] id="jCHK5okh2fYW"
# Right now it looks like our model is not performing well, lets try and improve it.
#
# To try and improve our model, we'll run 2 experiments:
# 1. Add an extra layer with more hidden units and use the Adam optimizer
# 2. Train for longer (like 200 epochs)
# 3. We can also do our custom experiments to improve it.
# + colab={"base_uri": "https://localhost:8080/"} id="0noQgs0h2fdG" outputId="aef5a34d-61c7-451c-92c9-2d461ec01539"
# Set random seed
tf.random.set_seed(42)
# 1. Create the model
insurance_model_2 = tf.keras.Sequential([
tf.keras.layers.Dense(100),
tf.keras.layers.Dense(10),
tf.keras.layers.Dense(1)
],name = "insurace_model_2")
# 2. Compile the model
insurance_model_2.compile(loss = tf.keras.losses.mae,
optimizer = tf.keras.optimizers.Adam(),
metrics = ["mae"])
# 3. Fit the model
insurance_model_2.fit(X_train, y_train, epochs = 100, verbose = 0)
# + colab={"base_uri": "https://localhost:8080/"} id="xvbwsSKi2fh0" outputId="8427f59b-5547-4362-ca95-bc5826b2897a"
insurance_model_2.evaluate(X_test, y_test)
# + id="KNmGz50G2fml"
# Set random seed
tf.random.set_seed(42)
# 1. Create the model
insurance_model_3 = tf.keras.Sequential([
tf.keras.layers.Dense(100),
tf.keras.layers.Dense(10),
tf.keras.layers.Dense(1)
],name = "insurace_model_2")
# 2. Compile the model
insurance_model_3.compile(loss = tf.keras.losses.mae,
optimizer = tf.keras.optimizers.Adam(),
metrics = ["mae"])
# 3. Fit the model
history = insurance_model_3.fit(X_train, y_train, epochs = 200, verbose = 0)
# + colab={"base_uri": "https://localhost:8080/"} id="l6Eoor542UTQ" outputId="6b2ca494-8489-4337-985e-7f53cbbe2571"
# Evaluate our third model
insurance_model_3.evaluate(X_test, y_test)
# + colab={"base_uri": "https://localhost:8080/", "height": 313} id="Hv0krA6L1UQ1" outputId="742bd938-2a0e-4f84-82de-c75c943a320c"
# Plot history (also known as a loss curve or a training curve)
pd.DataFrame(history.history).plot()
plt.ylabel("loss")
plt.xlabel("epochs")
plt.title("Training curve of our model")
# + [markdown] id="0gKwZS-T1UWN"
# **Question:** How long should you train for?
#
# It depends, It really depends on problem you are working on. However, many people have asked this question before, so TensorFlow has a solution!, It is called [EarlyStopping callback](https://www.tensorflow.org/api_docs/python/tf/keras/callbacks/EarlyStopping), which is a TensorFlow component you can add to your model to stop training once it stops improving a certain metric.
#
#
# + [markdown] id="N6XVGc5bU4j7"
# ## Preprocessing data (normalization and standardization)
# + [markdown] id="3AM6Zd-kVieF"
# Short review of our modelling steps in TensorFlow:
# 1. Get data ready(turn into tensors)
# 2. Build or pick a pretrained model (to suit your problem)
# 3. Fit the model to the data and make a prediction.
# 4. Evaluate the model.
# 5. Imporve through experimentation.
# 6. Save and reload your trained models.
#
# we are going to focus on the step 1 to make our data set more rich for training.
# some steps involved in getting data ready:
# 1. Turn all data into numbers(neural networks can't handle strings).
# 2. Make sure all of your tensors are the right shape.
# 3. Scale features(normalize or standardize, neural networks tend to prefer normalization) -- this is the one thing we haven't done while preparing our data.
#
# **If you are not sure on which to use for scaling, you could try both and see which perform better**
# + id="NTsaVDFmbq3v"
# Import required libraries
import tensorflow as tf
import pandas as pd
import matplotlib.pyplot as plt
# + colab={"base_uri": "https://localhost:8080/", "height": 423} id="8Dd11XPQbq1E" outputId="0acfad7d-7e0a-44f6-cf80-f5dcbb6f2519"
# Read in the insurance dataframe
insurance = pd.read_csv("https://raw.githubusercontent.com/stedy/Machine-Learning-with-R-datasets/master/insurance.csv")
insurance
# + [markdown] id="93OQY5mmbqyS"
# To prepare our data, we can borrow few classes from Scikit-Learn
# + id="uyI-ZyMvbqv1"
from sklearn.compose import make_column_transformer
from sklearn.preprocessing import MinMaxScaler, OneHotEncoder
from sklearn.model_selection import train_test_split
# + [markdown] id="nMarkBvpXm4v"
# **Feature Scaling**:
#
#
# | **Scaling type** | **what it does** | **Scikit-Learn Function** | **when to use** |
# | --- | --- | --- | --- |
# | scale(refers to as normalization) | converts all values to between 0 and 1 whilst preserving the original distribution | `MinMaxScaler` | Use as default scaler with neural networks |
# | Standarization | Removes the mean and divides each value by the standard deviation | `StandardScaler` | Transform a feature to have close to normal distribution |
#
# + id="bpWKk8umbqt2"
#Create a column transformer
ct = make_column_transformer(
(MinMaxScaler(), ["age", "bmi", "children"]), # Turn all values in these columns between 0 and 1
(OneHotEncoder(handle_unknown = "ignore"), ["sex", "smoker", "region"])
)
# Create our X and Y values
# because we reimported our dataframe
X = insurance.drop("charges", axis = 1)
y = insurance["charges"]
# Build our train and test set
X_train, X_test, y_train, y_test = train_test_split(X,y, test_size = 0.2, random_state = 42)
# Fit the column transformer to our training data (only training data)
ct.fit(X_train)
# Transform training and test data with normalization(MinMaxScaler) and OneHotEncoder
X_train_normal = ct.transform(X_train)
X_test_normal = ct.transform(X_test)
# + colab={"base_uri": "https://localhost:8080/"} id="0VMTxc3Fbqqm" outputId="cc20605e-045e-4438-8b65-1197a9e80b5f"
# What does our data look like now??
X_train.loc[0]
# + colab={"base_uri": "https://localhost:8080/"} id="kjtsAdpTevso" outputId="00803320-e9e9-4bc5-c15c-657a9f4b651d"
X_train_normal[0], X_train_normal[12], X_train_normal[78]
# we have turned all our data into numerical encoding and aso normalized the data
# + colab={"base_uri": "https://localhost:8080/"} id="wKKVwAVDevzO" outputId="c0921c94-c3fd-4a6b-853d-edad66315ea2"
X_train.shape, X_train_normal.shape
# + [markdown] id="DNfVDX3jfery"
# Beautiful! our data has been normalized and One hot encoded. Let's build Neural Network on it and see how it goes.
# + id="IDSFGWHybqn9"
# Build a neural network model to fit on our normalized data
tf.random.set_seed(42)
# 1. Create the model
insurance_model_4 = tf.keras.Sequential([
tf.keras.layers.Dense(100),
tf.keras.layers.Dense(10),
tf.keras.layers.Dense(1)
])
# 2. Compile the model
insurance_model_4.compile(loss = tf.keras.losses.mae,
optimizer = tf.keras.optimizers.Adam(),
metrics = ["mae"])
# 3. Fit the model
history = insurance_model_4.fit(X_train_normal, y_train, epochs= 100, verbose = 0)
# + colab={"base_uri": "https://localhost:8080/"} id="7j5HBOnsglgQ" outputId="84ffa97c-4aad-47c5-ff51-d1b35899a37d"
# Evaluate our insurance model trained on normalized data
insurance_model_4.evaluate(X_test_normal, y_test)
# + colab={"base_uri": "https://localhost:8080/"} id="YqNHTyOngldS" outputId="43f19c09-0b4d-4fca-e7a4-d90bd9342459"
insurance_model_4.summary()
# + colab={"base_uri": "https://localhost:8080/", "height": 313} id="SjNyIzg-glaj" outputId="0838fcff-f75f-4571-b387-95b35e9a19ef"
pd.DataFrame(history.history).plot()
plt.ylabel("loss")
plt.xlabel("epochs")
plt.title("Training curve of insurance_model_4")
# + [markdown] id="tKam_38Tfsd9"
# Let's just plot some graphs. Since we have use them the least in this notebook.
# + colab={"base_uri": "https://localhost:8080/", "height": 283} id="BhE-ytXuVibS" outputId="c734fd5d-3ea1-40fb-95b3-f7b9b0f2a7da"
X["age"].plot(kind = "hist")
# + colab={"base_uri": "https://localhost:8080/", "height": 283} id="u-4DHJ-uViYW" outputId="53f65642-394a-4919-ea65-499c4d460efe"
X["bmi"].plot(kind = "hist")
# + colab={"base_uri": "https://localhost:8080/"} id="cQ8osxbLXiuo" outputId="e4dcb3a4-1437-45d9-a668-0ca8bf5371da"
X["children"].value_counts()
# + [markdown] id="EuFKGN4Ui1wp"
# ## **External Resources:**
# * [MIT introduction deep learning lecture 1](https://youtu.be/njKP3FqW3Sk)
# * [Kaggle's datasets](https://www.kaggle.com/data)
# * [Lion Bridge's collection of datasets](https://lionbridge.ai/datasets/)
# + [markdown] id="zbEyF2ukZN9y"
# ## Bibliography:
#
# * [Learn TensorFlow and Deep Learning fundamentals with Python (code-first introduction) Part 1/2](https://www.youtube.com/watch?v=tpCFfeUEGs8&list=RDCMUCr8O8l5cCX85Oem1d18EezQ&start_radio=1&rv=tpCFfeUEGs8&t=3)
#
# * [Medical cost personal dataset](https://www.kaggle.com/mirichoi0218/insurance)
#
# * [TensorFlow documentation](https://www.tensorflow.org/api_docs/python/tf)
#
# * [TensorFlow and Deep learning Daniel Bourke GitHub Repo](https://github.com/mrdbourke/tensorflow-deep-learning)
| _notebooks/2022-02-11-Neural-Networks-Regression-with-TensorFlow.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
pha_mad_csv = "/media/andivanov/DATA/experiments/VideoMatte5x18/out/pha_mad.csv"
df = pd.read_csv(pha_mad_csv, index_col="clipname")
df
df['fgr'] = df.index.map(lambda x: x.split("_", 1)[0])
df['bgr'] = df.index.map(lambda x: "".join(x.split("_", 1)[1:]))
metrics_csv = "/media/andivanov/DATA/experiments/VideoMatte5x18/out/metrics.csv"
full_df = pd.read_csv(metrics_csv, index_col="clipname")
df = pd.merge(full_df['bgr_type'], df, left_index=True, right_index=True)
df
# ### Worst frame analysis
df['worst_frame'] = df.max(axis=1, skipna=True, numeric_only=True).values
df['worst_frame'].to_csv("/media/andivanov/DATA/experiments/VideoMatte5x18/out/worst_pha_mad.csv", index_label="clipname")
df['worst_frame']
df.groupby('bgr_type')['worst_frame'].mean()
df.groupby('bgr')['worst_frame'].mean()
df.groupby('fgr')['worst_frame'].mean()
# #### Single video worst frame analysis
df.loc['0001_00037', "worst_frame"]
# ### Intervals of frames analysis
df["mad(0-100)"] = df[list(map(lambda x: str(x), list(range(0, 100))))].mean(axis=1).values
df["mad(100-200)"] = df[list(map(lambda x: str(x), list(range(100, 200))))].mean(axis=1, skipna=True).values
df["mad(200-300)"] = df[list(map(lambda x: str(x), list(range(200, 300))))].mean(axis=1, skipna=True).values
df["mad(300-400)"] = df[list(map(lambda x: str(x), list(range(300, 400))))].mean(axis=1, skipna=True).values
df["mad(400-500)"] = df[list(map(lambda x: str(x), list(range(400, 500))))].mean(axis=1, skipna=True).values
df["mad(500-600)"] = df[list(map(lambda x: str(x), list(range(500, 600))))].mean(axis=1, skipna=True).values
df["mad(600-700)"] = df[list(map(lambda x: str(x), list(range(600, 700))))].mean(axis=1, skipna=True).values
df.groupby('bgr_type')[["mad(0-100)", "mad(100-200)", "mad(200-300)", "mad(300-400)", "mad(400-500)"]].mean()
df.groupby('bgr')[["mad(0-100)", "mad(100-200)", "mad(200-300)", "mad(300-400)", "mad(400-500)"]].mean()
df.loc["0001_00037"].filter(like='mad')
df.loc["0001_00037"][[x for x in range(200, 300)]].values
df_dict = {"clip_0": [1, 1, 1], "clip_1": [1, 1]}
df_dummy = pd.DataFrame.from_dict(df_dict, orient="index", columns=[0, 1, 2])
df_dummy
df_dummy.loc["mean"] = df.mean()
df_dummy
| evaluation/notebooks/per_frame_mad_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
def dydx(x,y):
#set the derivatives
#our equations is d^2y/dx^2
#so we can write
#dydx =z
#dzdx=-y
# We will set y = y[0]
# We will set z = y[1]
#declare an array
y_derivs=np.zeros(2)
#set dy/dx =z
y_derivs[0] = y[1]
#set dfdx = -y
y_derivs[1] = -1.*y[0]
return y_derivs
# ## Define the 4th order RK method
#nv number of varbiable
#xi x and time i
#yi the values of y and z
#h is a step
def rk4_mv_core(dydx,xi,yi,nv,h):
#declare k? array
k1 = np.zeros(nv)
k2 = np.zeros(nv)
k3 = np.zeros(nv)
k4 = np.zeros(nv)
#define x at 1/2 step
x_ipoh = xi +0.5*h
#define x at 1 step
x_ipo = xi +h
y_temp = np.zeros(nv)
y_derivs = dydx(xi,yi)
k1[:] = h*y_derivs[:]
y_temp[:] = yi[:] + 0.5*k1[:]
y_derivs = dydx(x_ipoh, y_temp)
k2[:] = h*y_derivs[:]
#get k3 values
y_temp[:] =yi[:] + 0.5*k2[:]
y_derivs = dydx(x_ipoh,y_temp)
k3[:] = h*y_derivs[:]
#get k4 values
y_temp[:] = yi[:] + k3[:]
y_derivs = dydx(x_ipo, y_temp)
k4[:]=h*y_derivs[:]
#advance y by a step h
yipo = yi+(k1 + 2*k2 + 2*k3 + k4)/6.0
return yipo
# ## Define an adaptive step size driver for RK4
def rk4_mv_ad(dydx,x_i,y_i,nv,h,tol):
#define safety scale
SAFETY = 0.9
H_NEW_FAC = 2.0
#set a max # of iterations
imax = 10000
#set a iteration variable
i=0
#create an error
Delta = np.full(nv,2*tol)
h_step = h
#adjust the step
while(Delta.max()/tol > 1.0):
#estimate our error by taking one step of size h
y_2 = rk4_mv_core(dydx, x_i,y_i,nv,h_step)
y_1 = rk4_mv_core(dydx,x_i,y_i,nv,0.5*h_step)
y_11 = rk4_mv_core(dydx,x_i+0.5*h_step,y_1,nv,0.5*h_step)
Delta = np.fabs(y_2 - y_11)
#if the error is too large, take a smaller step
if(Delta.max()/tol>1.0): # this means it to big
h_step *=SAFETY * (Delta.max()/tol)**(-0.25)
#check iterations
if(i>imax):
print("Too many iterations in rk4_mv_ad()")
raise StopIteration("Ending after i = ", i)
i +=1
#next time, try to take a bigger step
h_new = np.fmin(h_step * (Delta.max()/tol)**(-0.9), h_step * H_NEW_FAC)
#return the answer
return y_2,h_new,h_step
# ## Define a wrapper for RK4
# +
def rk4_mv(dydx,a,b,y_a,tol):
#dydx is the deriative wrt x
#a is the lower bound
#b is the uper bond
#y-a boundary Condution
xi = a
yi = y_a.copy()
#An initial step size == make very small
h = 1.0e-4 * (b-a)
#set a maximum number of iterations
imax = 10000
#set an iterations variable
i =0
#set the number of coupled odes to the size of y_a
nv = len(y_a) # y_a has the size of our boundery conditions
x=np.full(1,a)
y= np.full((1,nv),y_a)
#set a flag
flag = 1
#loop till we reach the right side
while(flag):
#calcualte y_i+1
yi_new, h_new,h_step = rk4_mv_ad(dydx, xi, yi,nv,h, tol)
#Update the step
h = h_new
#Prevent an over step
if(xi+h_step>b):
#Take a smaller step
h = b-xi
#retake the step
yi_new, h_new,h_step = rk4_mv_ad(dydx, xi, yi,nv,h, tol)
flag = 0
#update the values
xi += h_step
yi[:] = yi_new[:]
#add the step to the array
x = np.append(x,xi) #Add the answer into the new array
y_new = np.zeros((len(x),nv))
y_new[0:len(x)-1,:] = y
y_new[-1,:]=yi[:]
del y
y = y_new
#prevent too mant iterations
if(i>=imax):
print("Maximum iterations reached.")
raise StopIteration("Iteration number =",i)
i +=1
#output some function
s= "i = %3d\tx = %9.8f\th = %9.8f\tb =%9.8f" % (i,xi,h_step,b)
#\tb print a tab (makes space)
#% print a float size of 8 digits behind the ecimal
print (s)
#break if new xi is == b
if (xi==b):
flag=0
#return the answer
return x,y
# +
a = 0.0
b = 2.0 * np.pi
y_0 = np.zeros(2)
y_0[0] = 0.0
y_0[1] = 1.0
nv = 2
tolerance = 1.0e-6
#perform the integratoin
x,y = rk4_mv(dydx,a,b,y_0,tolerance)
# -
# ## Plot the result
# +
plt.plot(x,y[:,0],'o',label='y(x)')
plt.plot(x,y[:,1],'o',label='dydx(x)')
xx = np.linspace(0,2.0*np.pi,1000)
plt.plot(xx,np.sin(xx),label='sin(x)')
plt.plot(xx,np.cos(xx),label='cos(x)')
plt.xlabel('X')
plt.ylabel('y,dy/dx')
plt.legend(frameon=False)
plt.show()
# -
# +
sine = np.sin(x)
cosine = np.cos(x)
y_error = (y[:,0]-sine)
dydx_error = ( y[:,1]-cosine)
plt.plot(x,y_error,label='y(x) Error')
plt.plot(x, dydx_error,label = 'dydx(x) Error')
plt.legend(frameon = False)
plt.show()
# -
| rk4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
# %matplotlib inline
# -
# First of all, we need the dataset. We can download and it using the following commands.
# Now, we load the dataset. For this we use the special function, which reads the csv-files from the directory and puts the data in a convenient format.
# +
from alice_ml.preprocessing import load_dataset
path = 'dataset_Kids'
data_kids, raw_annotations_kids = load_dataset(path, preload=False)
# +
from alice_ml.preprocessing import load_dataset
path = 'dataset_mu_rhythm_1.1'
data_mu, raw_annotations_mu = load_dataset(path, preload=False)
# +
from alice_ml.preprocessing import load_dataset
path = 'dataset_Adults_1.0'
data_adults, raw_annotations_adults = load_dataset(path, preload=False)
# -
# Now, we calculate features for each ic in the dataset.
# +
from alice_ml.features import build_feature_df
features_adults = build_feature_df(data_adults, default=True, custom_features={})
features_adults
# +
from alice_ml.features import build_feature_df
features_mu = build_feature_df(data_mu, default=True, custom_features={})
features_mu
# -
features_kids = build_feature_df(data_kids, default=True, custom_features={})
features_kids
common_features = pd.concat([features_adults, features_kids, features_mu])
ann = pd.DataFrame(index = common_features.index)
ann['dataset'] = 'adults'
ann.loc[features_kids.index] = 'kids'
ann.loc[features_mu.index] = 'kids'
# Targets are calculated as the average of the answers of multiple experts for each ic.
for i, n in raw_annotations.groupby('user_hash'):
print(i)
for j in raw_annotations[raw_annotations.user_hash == i].drop(['ic_id', 'user_hash', 'comment'], axis=1).fillna(-1).columns:
print(raw_annotations[raw_annotations.user_hash == i][j].value_counts())
# +
def get_target_distribution(annotations):
return annotations.groupby('ic_id').mean().apply(lambda x: x.value_counts())
get_target_distribution(raw_annotations_mu)
# -
# For now we do not want to distinguish between eye movement types, so we unite them under `'flag_eyes'`. We also presume the positive label for `'flag_alpha'` or `'flag_mu'` implies the positive label for `'flag_brain'`. Finally, since we do not have sufficient data for distinguishing line noise, we will not consider this flag.
# +
def propagate_labels(annotations):
return annotations.assign(flag_brain=annotations[['flag_brain', 'flag_alpha', 'flag_mu']].any(axis=1)) \
.drop(columns=['flag_line_noise'])
annotations_mu = propagate_labels(raw_annotations_mu)
get_target_distribution(annotations_mu)
# +
def propagate_labels(annotations):
return annotations.assign(flag_brain=annotations[['flag_brain', 'flag_alpha', 'flag_mu']].any(axis=1)) \
.drop(columns=['flag_line_noise'])
annotations_kids = propagate_labels(raw_annotations_kids)
get_target_distribution(annotations_kids)
# +
def propagate_labels(annotations):
return annotations.assign(flag_eyes=annotations[['flag_eyes', 'flag_eyes_h', 'flag_eyes_v']].any(axis=1)).assign(flag_brain=annotations[['flag_brain', 'flag_alpha', 'flag_mu']].any(axis=1)) \
.drop(columns=['flag_line_noise'])
# annotations_kids = propagate_labels(raw_annotations_kids)
# get_target_distribution(annotations_kids)
# -
annotations_adults = propagate_labels(raw_annotations_adults)
annotations_kids = propagate_labels(raw_annotations_kids)
# annotations_kids = propagate_labels(raw_annotations_kids)
annotations_mu = propagate_labels(raw_annotations_mu)
annotations_adults = propagate_labels(raw_annotations_adults)
# In order to get binary labels we apply threshold to these averages. By default, the each label has weight $\frac{1}{n}$, where $n$ is the number of labels given to the ic by this expert. We set the threshold to $0.33$ For flags `alpha` and `mu` we choose a simpler approach: each label has weight $1$. The threshold in this case is equal to $0.5$.
# +
from alice_ml.preprocessing import build_target_df
targets_mu = build_target_df(annotations_mu, weights='uniform', strategy='mean', threshold=0.33)
targets_mu[['flag_alpha', 'flag_mu', 'flag_heart']] = build_target_df(annotations_mu, ['flag_alpha', 'flag_mu', 'flag_heart'], strategy='majority')
print(targets_mu.sum())
targets_mu = targets_mu.loc[:, (targets_mu.mean(axis=0) > 0.05) & (targets_mu.all(axis=0) < 0.95)]
# targets_mu.head()
# +
targets_kids = build_target_df(annotations_kids, weights='uniform', strategy='mean', threshold=0.33)
targets_adults = build_target_df(annotations_adults, weights='uniform', strategy='mean', threshold=0.33)
# -
targets_mu.shape, targets_kids.shape, targets_adults.shape
features_mu.shape, features_kids.shape, features_adults.shape
targets = pd.concat([targets_kids, targets_adults, targets_mu])
features = common_features.copy()
targets.to_csv('targets_common.csv')
features.to_csv('featrues_common.csv')
common_features.shape
def to_common_samples(df_list=()):
cs = set(df_list[0].index)
for i in range(1, len(df_list)):
cs = cs.intersection(df_list[i].index)
if len(cs) < 1:
warnings.warn('No common samples!')
return [df_list[i].loc[list(cs)] for i in range(len(df_list))]
features, targets = to_common_samples([features, targets])
features.shape, targets.shape
# +
from alice_ml.preprocessing import build_target_df
targets_kids = build_target_df(annotations_kids, weights='uniform', strategy='mean', threshold=0.33)
targets_kids[['flag_alpha', 'flag_mu', 'flag_heart']] = build_target_df(annotations_kids, ['flag_alpha', 'flag_mu', 'flag_heart'], strategy='majority')
print(targets_kids.sum())
targets_kids = targets_kids.loc[:, (targets_kids.mean(axis=0) > 0.05) & (targets_kids.all(axis=0) < 0.95)]
targets_kids.head()
# +
from alice_ml.preprocessing import build_target_df
targets_adults = build_target_df(annotations_adults, weights='uniform', strategy='mean', threshold=0.33)
targets_adults[['flag_alpha', 'flag_mu', 'flag_heart']] = build_target_df(annotations_adults, ['flag_alpha', 'flag_mu', 'flag_heart'], strategy='majority')
print(targets_adults.sum())
targets_adults = targets_adults.loc[:, (targets_adults.mean(axis=0) > 0.05) & (targets_adults.all(axis=0) < 0.95)]
targets_adults.shape
# -
features = pd.read_csv('featrues_common.csv', index_col=0)
targets = pd.read_csv('target_common.csv', index_col=0)
features.loc[targets.index].shape, targets.shape
# For training we use XGboost, LinearSVM, and Logistic Regression with default parameters. We do a stratified train-test split with test size = 0.3.
from sklearn.preprocessing import StandardScaler
# +
from sklearn.linear_model import LogisticRegression
from xgboost import XGBClassifier
from sklearn.svm import SVC
from sklearn.preprocessing import StandardScaler
models = {'LR': LogisticRegression(solver='liblinear', penalty='l2', C=1.0),
'XGB': XGBClassifier(learning_rate=0.3, max_depth=4, n_estimators=30, eval_metric='logloss', use_label_encoder=False),
'SVM': SVC(kernel='linear', probability=True)}
n_repeats = 50
# +
from sklearn.model_selection import train_test_split
def repeated_train_test(features, target, model, n_repeats, test_size=0.3, axes=None, random_state=57):
y_true = []
y_pred = []
y_proba = []
np.random.seed(random_state)
for _ in range(n_repeats):
X_train, X_test, y_train, y_test = train_test_split(features, target, test_size=test_size, stratify=target)
scaler = StandardScaler()
X_train = scaler.fit_transform(X_train)
X_test = scaler.transform(X_test)
model.fit(X_train, y_train)
y_true.append(y_test)
y_pred.append(model.predict(X_test))
y_proba.append(model.predict_proba(X_test)[:, 1])
return np.array(y_true), np.array(y_pred), np.array(y_proba)
# +
from alice_ml.vis import plot_aggregated_pr_curve, plot_aggregated_roc_curve
def plot_curves_grid(features, targets, models):
fig_roc, axes_roc = plt.subplots(len(models), len(targets.columns), figsize=(4 * len(targets.columns), 3 * len(models)))
fig_pr, axes_pr = plt.subplots(len(models), len(targets.columns), figsize=(4 * len(targets.columns), 3 * len(models)))
for (model_name, model), row_roc, row_pr in zip(models.items(), axes_roc, axes_pr):
for flag, ax_roc, ax_pr in zip(targets, row_roc, row_pr):
y_true, _, y_proba = repeated_train_test(features, targets[flag], model, n_repeats)
plot_aggregated_roc_curve(y_true, y_proba, ax_roc)
plot_aggregated_pr_curve(y_true, y_proba, ax_pr)
row_roc[0].set_ylabel(model_name, fontsize=28, labelpad=16)
row_pr[0].set_ylabel(model_name, fontsize=28, labelpad=16)
row_roc[-1].set_ylabel('True positive rate', fontsize=18)
row_pr[-1].set_ylabel('Precision', fontsize=18)
row_roc[-1].yaxis.set_label_position('right')
row_pr[-1].yaxis.set_label_position('right')
for target_name, ax_roc, ax_pr in zip(targets, axes_roc[0], axes_pr[0]):
ax_roc.set_title(target_name[5:].capitalize(), fontsize=28, pad=16)
ax_pr.set_title(target_name[5:].capitalize(), fontsize=28, pad=16)
for ax_roc, ax_pr in zip(axes_roc[-1], axes_pr[-1]):
ax_roc.set_xlabel('False positive rate', fontsize=18)
ax_pr.set_xlabel('Recall', fontsize=18)
fig_roc.subplots_adjust(wspace=0.1, hspace=0.1)
fig_pr.subplots_adjust(wspace=0.1, hspace=0.1)
for row_roc, row_pr in zip(axes_roc[:-1], axes_pr[:-1]):
for ax_roc, ax_pr in zip(row_roc, row_pr):
plt.setp(ax_roc.get_xticklabels(), visible=False)
plt.setp(ax_pr.get_xticklabels(), visible=False)
for row_roc, row_pr in zip(axes_roc, axes_pr):
for ax_roc, ax_pr in zip(row_roc[:-1], row_pr[:-1]):
plt.setp(ax_roc.get_yticklabels(), visible=False)
plt.setp(ax_pr.get_yticklabels(), visible=False)
return fig_roc, fig_pr
# -
i='kids'
for i in ['mu', 'adults', 'kids']:
print(i)
fig_roc, fig_pr = plot_curves_grid(common_features.loc[ann[ann.dataset!=i].index].sort_index(), targets.loc[ann[ann.dataset!=i].index][['flag_eyes', 'flag_brain']].sort_index(), models)
# + tags=[]
fig_roc, fig_pr = plot_curves_grid(common_features.sort_index(), targets[['flag_eyes', 'flag_brain']].sort_index(), models)
# +
from sklearn.metrics import auc, roc_curve, precision_recall_curve
def get_aucs(true, proba):
roc_aucs, pr_aucs = [], []
for true, proba in zip(y_true, y_proba):
fpr, tpr, _ = roc_curve(true, proba, drop_intermediate=False)
prec, rec, _ = precision_recall_curve(true, proba)
roc_aucs.append(auc(fpr, tpr))
pr_aucs.append(auc(rec, prec))
return roc_aucs, pr_aucs
# -
features
ann, features = to_common_samples([ann, features])
# + tags=[]
from itertools import product
from sklearn.metrics import f1_score
roc_aucs = pd.DataFrame(index=models, columns=targets.columns, dtype=float)
pr_aucs = pd.DataFrame(index=models, columns=targets.columns, dtype=float)
f1_scores = pd.DataFrame(index=models, columns=targets.columns, dtype=float)
for (model_name, model), flag in product(models.items(), targets[['flag_eyes', 'flag_brain']]):
y_true, y_pred, y_proba = repeated_train_test(common_features.sort_index().loc[ann[ann.dataset!='mu'].index], targets[flag].sort_index().loc[ann[ann.dataset!='mu'].index], model, n_repeats)
roc_auc_values, pr_auc_values = get_aucs(y_true, y_proba)
f1_score_values = [f1_score(true, pred) for true, pred in zip(y_true, y_pred)]
roc_aucs.loc[model_name, flag] = np.mean(roc_auc_values)
pr_aucs.loc[model_name, flag] = np.mean(pr_auc_values)
f1_scores.loc[model_name, flag] = np.mean(f1_score_values)
# -
roc_aucs
pr_aucs
pr_aucs
f1_scores
f1_scores['flag_eyes']
# For each flag we choose the model based on ROC-AUC if the classes are balanced and on F1 scores otherwise.
# Save pretrained models
# +
import joblib
scaler = StandardScaler()
X_train = scaler.fit_transform(features)
joblib.dump(scaler, 'scaler.joblib')
for flag in targets[['flag_brain', 'flag_eyes']]:
model_lr = LogisticRegression(solver='liblinear', penalty='l2', C=1.0)
model.fit(X_train, targets[flag])
joblib.dump(model, 'lr_' + flag + '.joblib')
# +
scaler = StandardScaler()
X_train = scaler.fit_transform(features)
joblib.dump(scaler, 'scaler.joblib')
for flag in targets[['flag_brain', 'flag_eyes']]:
model = XGBClassifier(base_score=0.5, booster='gbtree', colsample_bylevel=1,
colsample_bynode=1, colsample_bytree=1, enable_categorical=False,
eval_metric='logloss', gamma=0, gpu_id=-1, importance_type=None,
interaction_constraints='', learning_rate=0.3, max_delta_step=0,
max_depth=4, min_child_weight=1,
monotone_constraints='()', n_estimators=30, n_jobs=4,
num_parallel_tree=1, predictor='auto', random_state=0,
reg_alpha=0, reg_lambda=1, scale_pos_weight=1, subsample=1,
tree_method='exact', use_label_encoder=False,
validate_parameters=1, verbosity=None)
model.fit(X_train, targets[flag])
joblib.dump(model, 'xgb_' + flag + '.joblib')
# +
scaler = StandardScaler()
X_train = scaler.fit_transform(features)
joblib.dump(scaler, 'scaler.joblib')
for flag in targets[['flag_brain', 'flag_eyes']]:
model = SVC(kernel='linear', probability=True)
model.fit(X_train, targets[flag])
joblib.dump(model, 'svc_' + flag + '.joblib')
# -
# # bezdelushki
# +
y_train = targets['flag_eyes'].sort_index().loc[ann[ann.dataset!='adults'].index]
params = {
'max_depth': [2, 3, 4, 5],
'learning_rate': [0.001, 0.01, 0.1, 0.2, 0.3],
'subsample': [0.5, 0.6, 0.7, 0.8, 0.9, 1.0],
'colsample_bytree': [0.4, 0.5, 0.6, 0.7, 0.8, 0.9, 1.0],
# 'num_leaves': [6,8,12,16],
'subsample' : [0.7,0.75],
'reg_alpha' : [1,1.2],
# 'colsample_bylevel': [0.4, 0.5, 0.6, 0.7, 0.8, 0.9, 1.0],
'min_child_weight': [0.5, 1.0, 3.0, 5.0, 7.0, 10.0],
# 'gamma': [0, 0.25, 0.5, 1.0],
'reg_lambda': [0.1, 1.0, 5.0, 10.0, 50.0, 100.0],
'n_estimators': [40, 100, 160, 220, 280, 340, 400]}
model = XGBClassifier(use_label_encoder=False)
skf = StratifiedKFold(n_splits=3, shuffle=True)
random_search = RandomizedSearchCV(
model,
param_distributions=params,
n_iter=10,
scoring='f1_micro',
n_jobs=-1,
cv = skf.split(X_train, y_train))
random_search.fit(X_train, y_train)
best_params = random_search.best_params_
model_xgb = XGBClassifier(use_label_encoder=False, **best_params)
model_xgb.fit(X_train, y_train)
# -
best_params
# +
best_params= {
'subsample': 0.76,
'reg_lambda': 50.0,
'reg_alpha': 1.0,
'n_estimators': 4000,
'min_child_weight': 1.0,
'max_depth': 2,
'learning_rate': .0001,
'colsample_bytree': 0.8
}
model_xgb = XGBClassifier(use_label_encoder=False, **best_params)
model_xgb.fit(X_train, y_train)
f1_score(np.ravel(targets['flag_eyes'].sort_index().loc[ann[ann.dataset=='adults'].index]),
model_xgb.predict(scaler.transform(features.sort_index().loc[ann[ann.dataset=='adults'].index])))
# -
f1_score_values_lr = f1_score(np.ravel(targets['flag_eyes'].sort_index().loc[ann[ann.dataset=='adults'].index]),
model_lr.predict(scaler.transform(features.sort_index().loc[ann[ann.dataset=='adults'].index])))
# roc_auc_values_lr, pr_auc_values_lr = get_aucs(
# np.ravel(targets['flag_eyes'].sort_index().loc[ann[ann.dataset=='adults'].index]),
# model_lr.predict_proba(scaler.transform(features.sort_index().loc[ann[ann.dataset=='adults'].index])))
f1_score_values_svc = f1_score(np.ravel(targets['flag_eyes'].sort_index().loc[ann[ann.dataset=='adults'].index]),
model_svc.predict(scaler.transform(features.sort_index().loc[ann[ann.dataset=='adults'].index])))
# roc_auc_values_svc, pr_auc_values_svc = get_aucs(
# np.ravel(targets['flag_eyes'].sort_index().loc[ann[ann.dataset=='adults'].index]),
# model_svc.predict_proba(scaler.transform(features.sort_index().loc[ann[ann.dataset=='adults'].index])))
f1_score_values_xgb = f1_score(np.ravel(targets['flag_eyes'].sort_index().loc[ann[ann.dataset=='adults'].index]),
model_xgb.predict(scaler.transform(features.sort_index().loc[ann[ann.dataset=='adults'].index])))
# roc_auc_values_xgb, pr_auc_values_xgb = get_aucs(
# np.ravel(targets['flag_eyes'].sort_index().loc[ann[ann.dataset=='adults'].index]),
# model_xgb.predict_proba(scaler.transform(features.sort_index().loc[ann[ann.dataset=='adults'].index])))
f1_score_values_lr, f1_score_values_svc, f1_score_values_xgb
# +
scaler = StandardScaler()
X_train = scaler.fit_transform(features.sort_index().loc[ann[ann.dataset!='kids'].index])
model_lr = LogisticRegression(solver='liblinear', penalty='l2', C=1.0)
model_lr.fit(X_train, targets['flag_eyes'].sort_index().loc[ann[ann.dataset!='kids'].index])
# scaler = StandardScaler()
# X_train = scaler.fit_transform(features.sort_index().loc[ann[ann.dataset!='kids'].index])
model_svc = SVC(kernel='linear', probability=True)
model_svc.fit(X_train, targets['flag_eyes'].sort_index().loc[ann[ann.dataset!='kids'].index])
model_xgb = XGBClassifier(base_score=0.5, booster='gbtree', colsample_bylevel=1,
colsample_bynode=1, colsample_bytree=1, enable_categorical=False,
eval_metric='logloss', gamma=0, gpu_id=-1, importance_type=None,
interaction_constraints='', learning_rate=0.09, max_delta_step=0,
max_depth=4, min_child_weight=1,
monotone_constraints='()', n_estimators=30, n_jobs=4,
num_parallel_tree=1, predictor='auto', random_state=0,
reg_alpha=0, reg_lambda=1, scale_pos_weight=1, subsample=1,
tree_method='exact', use_label_encoder=False,
validate_parameters=1, verbosity=None)
model_xgb.fit(X_train, targets['flag_eyes'].sort_index().loc[ann[ann.dataset!='kids'].index])
# +
y_train = targets['flag_eyes'].sort_index().loc[ann[ann.dataset!='kids'].index]
params = {
'max_depth': [2, 3, 4, 5],
'learning_rate': [0.001, 0.01, 0.1, 0.2, 0.3],
'subsample': [0.5, 0.6, 0.7, 0.8, 0.9, 1.0],
'colsample_bytree': [0.4, 0.5, 0.6, 0.7, 0.8, 0.9, 1.0],
# 'num_leaves': [6,8,12,16],
'subsample' : [0.7,0.75],
'reg_alpha' : [1,1.2],
# 'colsample_bylevel': [0.4, 0.5, 0.6, 0.7, 0.8, 0.9, 1.0],
'min_child_weight': [0.5, 1.0, 3.0, 5.0, 7.0, 10.0],
# 'gamma': [0, 0.25, 0.5, 1.0],
'reg_lambda': [0.1, 1.0, 5.0, 10.0, 50.0, 100.0],
'n_estimators': [40, 100, 160, 220, 280, 340, 400]}
model = XGBClassifier(use_label_encoder=False)
skf = StratifiedKFold(n_splits=3, shuffle=True)
random_search = RandomizedSearchCV(
model,
param_distributions=params,
n_iter=10,
scoring='f1_micro',
n_jobs=-1,
cv = skf.split(X_train, y_train))
random_search.fit(X_train, y_train)
best_params = random_search.best_params_
model_xgb = XGBClassifier(use_label_encoder=False, **best_params)
model_xgb.fit(X_train, y_train)
# -
best_params
# +
best_params= {
'subsample': 0.8,
'reg_lambda': 50,
'reg_alpha': 5,
'n_estimators': 400,
'min_child_weight': 4.0,
'max_depth': 7,
'learning_rate': 0.9,
'colsample_bytree': 1
}
model_xgb = XGBClassifier(use_label_encoder=False, **best_params)
model_xgb.fit(X_train, y_train)
f1_score(np.ravel(targets['flag_eyes'].sort_index().loc[ann[ann.dataset=='kids'].index]),
model_xgb.predict(scaler.transform(features.sort_index().loc[ann[ann.dataset=='kids'].index])))
# -
f1_score_values_lr = f1_score(np.ravel(targets['flag_eyes'].sort_index().loc[ann[ann.dataset=='kids'].index]),
model_lr.predict(scaler.transform(features.sort_index().loc[ann[ann.dataset=='kids'].index])))
# roc_auc_values_lr, pr_auc_values_lr = get_aucs(
# np.ravel(targets['flag_eyes'].sort_index().loc[ann[ann.dataset=='kids'].index]),
# model_lr.predict_proba(scaler.transform(features.sort_index().loc[ann[ann.dataset=='kids'].index])))
f1_score_values_svc = f1_score(np.ravel(targets['flag_eyes'].sort_index().loc[ann[ann.dataset=='kids'].index]),
model_svc.predict(scaler.transform(features.sort_index().loc[ann[ann.dataset=='kids'].index])))
# roc_auc_values_svc, pr_auc_values_svc = get_aucs(
# np.ravel(targets['flag_eyes'].sort_index().loc[ann[ann.dataset=='kids'].index]),
# model_svc.predict_proba(scaler.transform(features.sort_index().loc[ann[ann.dataset=='kids'].index])))
f1_score_values_xgb = f1_score(np.ravel(targets['flag_eyes'].sort_index().loc[ann[ann.dataset=='kids'].index]),
model_xgb.predict(scaler.transform(features.sort_index().loc[ann[ann.dataset=='kids'].index])))
# roc_auc_values_xgb, pr_auc_values_xgb = get_aucs(
# np.ravel(targets['flag_eyes'].sort_index().loc[ann[ann.dataset=='kids'].index]),
# model_xgb.predict_proba(scaler.transform(features.sort_index().loc[ann[ann.dataset=='kids'].index])))
f1_score_values_lr, f1_score_values_svc, f1_score_values_xgb
# +
scaler = StandardScaler()
X_train = scaler.fit_transform(features.sort_index().loc[ann[ann.dataset!='mu'].index])
model_lr = LogisticRegression(solver='liblinear', penalty='l2', C=.1)
model_lr.fit(X_train, targets['flag_eyes'].sort_index().loc[ann[ann.dataset!='mu'].index])
# scaler = StandardScaler()
# X_train = scaler.fit_transform(features.sort_index().loc[ann[ann.dataset!='mu'].index])
model_svc = SVC(kernel='linear', probability=True)
model_svc.fit(X_train, targets['flag_eyes'].sort_index().loc[ann[ann.dataset!='mu'].index])
model_xgb = XGBClassifier(base_score=0.5, booster='gbtree', colsample_bylevel=1,
colsample_bynode=1, colsample_bytree=1, enable_categorical=False,
eval_metric='logloss', gamma=0, gpu_id=-1, importance_type=None,
interaction_constraints='', learning_rate=0.3, max_delta_step=0,
max_depth=4, min_child_weight=1,
monotone_constraints='()', n_estimators=30, n_jobs=4,
num_parallel_tree=1, predictor='auto', random_state=0,
reg_alpha=0, reg_lambda=1, scale_pos_weight=1, subsample=1,
tree_method='exact', use_label_encoder=False,
validate_parameters=1, verbosity=None)
model_xgb.fit(X_train, targets['flag_eyes'].sort_index().loc[ann[ann.dataset!='mu'].index])
# +
y_train = targets['flag_eyes'].sort_index().loc[ann[ann.dataset!='mu'].index]
params = {
'max_depth': [2, 3, 4, 5],
'learning_rate': [0.001, 0.01, 0.1, 0.2, 0.3],
'subsample': [0.5, 0.6, 0.7, 0.8, 0.9, 1.0],
'colsample_bytree': [0.4, 0.5, 0.6, 0.7, 0.8, 0.9, 1.0],
# 'num_leaves': [6,8,12,16],
'subsample' : [0.7,0.75],
'reg_alpha' : [1,1.2],
# 'colsample_bylevel': [0.4, 0.5, 0.6, 0.7, 0.8, 0.9, 1.0],
'min_child_weight': [0.5, 1.0, 3.0, 5.0, 7.0, 10.0],
# 'gamma': [0, 0.25, 0.5, 1.0],
'reg_lambda': [0.1, 1.0, 5.0, 10.0, 50.0, 100.0],
'n_estimators': [40, 100, 160, 220, 280, 340, 400]}
model = XGBClassifier(use_label_encoder=False)
skf = StratifiedKFold(n_splits=3, shuffle=True)
random_search = RandomizedSearchCV(
model,
param_distributions=params,
n_iter=10,
scoring='f1',
n_jobs=-1,
cv = skf.split(X_train, y_train))
random_search.fit(X_train, y_train)
best_params = random_search.best_params_
model_xgb = XGBClassifier(use_label_encoder=False, **best_params)
model_xgb.fit(X_train, y_train)
# +
best_params= {
'subsample': 0.75,
'reg_lambda': 0.15,
'reg_alpha': 1.2,
'n_estimators': 160,
'min_child_weight': 3.0,
'max_depth': 5,
'learning_rate': 0.9,
'colsample_bytree': 0.6
}
model_xgb = XGBClassifier(use_label_encoder=False, **best_params)
model_xgb.fit(X_train, y_train)
f1_score(np.ravel(targets['flag_eyes'].sort_index().loc[ann[ann.dataset=='mu'].index]),
model_xgb.predict(scaler.transform(features.sort_index().loc[ann[ann.dataset=='mu'].index])))
# -
f1_score_values_lr = f1_score(np.ravel(targets['flag_eyes'].sort_index().loc[ann[ann.dataset=='mu'].index]),
model_lr.predict(scaler.transform(features.sort_index().loc[ann[ann.dataset=='mu'].index])))
# roc_auc_values_lr, pr_auc_values_lr = get_aucs(
# np.ravel(targets['flag_eyes'].sort_index().loc[ann[ann.dataset=='mu'].index]),
# model_lr.predict_proba(scaler.transform(features.sort_index().loc[ann[ann.dataset=='mu'].index])))
f1_score_values_svc = f1_score(np.ravel(targets['flag_eyes'].sort_index().loc[ann[ann.dataset=='mu'].index]),
model_svc.predict(scaler.transform(features.sort_index().loc[ann[ann.dataset=='mu'].index])))
# roc_auc_values_svc, pr_auc_values_svc = get_aucs(
# np.ravel(targets['flag_eyes'].sort_index().loc[ann[ann.dataset=='mu'].index]),
# model_svc.predict_proba(scaler.transform(features.sort_index().loc[ann[ann.dataset=='mu'].index])))
f1_score_values_xgb = f1_score(np.ravel(targets['flag_eyes'].sort_index().loc[ann[ann.dataset=='mu'].index]),
model_xgb.predict(scaler.transform(features.sort_index().loc[ann[ann.dataset=='mu'].index])))
# roc_auc_values_xgb, pr_auc_values_xgb = get_aucs(
# np.ravel(targets['flag_eyes'].sort_index().loc[ann[ann.dataset=='mu'].index]),
# model_xgb.predict_proba(scaler.transform(features.sort_index().loc[ann[ann.dataset=='mu'].index])))
f1_score_values_lr, f1_score_values_svc, f1_score_values_xgb
f1_score_values_lr = f1_score(np.ravel(y_ho),
model_lr.predict(scaler.transform(X_ho)))
# roc_auc_values_lr, pr_auc_values_lr = get_aucs(
# np.ravel(targets['flag_eyes'].sort_index().loc[ann[ann.dataset=='kids'].index]),
# model_lr.predict_proba(scaler.transform(features.sort_index().loc[ann[ann.dataset=='kids'].index])))
f1_score_values_svc = f1_score(np.ravel(y_ho),
model_svc.predict(scaler.transform(X_ho)))
# roc_auc_values_svc, pr_auc_values_svc = get_aucs(
# np.ravel(targets['flag_eyes'].sort_index().loc[ann[ann.dataset=='kids'].index]),
# model_svc.predict_proba(scaler.transform(features.sort_index().loc[ann[ann.dataset=='kids'].index])))
f1_score_values_xgb = f1_score(np.ravel(y_ho),
model_xgb.predict(scaler.transform(X_ho)))
# roc_auc_values_xgb, pr_auc_values_xgb = get_aucs(
# np.ravel(targets['flag_eyes'].sort_index().loc[ann[ann.dataset=='kids'].index]),
# model_xgb.predict_proba(scaler.transform(features.sort_index().loc[ann[ann.dataset=='kids'].index])))
f1_score_values_lr, f1_score_values_svc, f1_score_values_xgb
# + code_folding=[]
X_train, X_ho, y_train, y_ho = train_test_split(features.sort_index(), targets.flag_eyes.sort_index(),
test_size=0.3, stratify=pd.concat([targets.flag_eyes, ann.dataset], axis=1)[['flag_eyes', 'dataset']],
random_state=42)
# +
# y_train = targets['flag_eyes'].sort_index().loc[ann[ann.dataset!='mu'].index]
params = {
'max_depth': [2, 3, 4, 5],
'learning_rate': [0.001, 0.01, 0.1, 0.2, 0.3],
'subsample': [0.5, 0.6, 0.7, 0.8, 0.9, 1.0],
'colsample_bytree': [0.4, 0.5, 0.6, 0.7, 0.8, 0.9, 1.0],
# 'num_leaves': [6,8,12,16],
'subsample' : [0.7,0.75],
'reg_alpha' : [1,1.2],
# 'colsample_bylevel': [0.4, 0.5, 0.6, 0.7, 0.8, 0.9, 1.0],
'min_child_weight': [0.5, 1.0, 3.0, 5.0, 7.0, 10.0],
# 'gamma': [0, 0.25, 0.5, 1.0],
'reg_lambda': [0.1, 1.0, 5.0, 10.0, 50.0, 100.0],
'n_estimators': [40, 100, 160, 220, 280, 340, 400]}
model = XGBClassifier(use_label_encoder=False)
skf = StratifiedKFold(n_splits=3, shuffle=True)
random_search = RandomizedSearchCV(
model,
param_distributions=params,
n_iter=10,
scoring='f1',
n_jobs=-1,
cv = skf.split(X_train, y_train))
random_search.fit(X_train, y_train)
best_params = random_search.best_params_
model_xgb = XGBClassifier(use_label_encoder=False, **best_params)
model_xgb.fit(X_train, y_train)
# +
scaler = StandardScaler()
X_train = scaler.fit_transform(X_train)
model_lr = LogisticRegression(solver='liblinear', penalty='l2', C=1.0)
model_lr.fit(X_train, y_train)
# scaler = StandardScaler()
# X_train = scaler.fit_transform(X_train)
model_svc = SVC(kernel='linear', probability=True)
model_svc.fit(X_train, y_train)
model_xgb = XGBClassifier(base_score=0.5, booster='gbtree', colsample_bylevel=1,
colsample_bynode=1, colsample_bytree=1, enable_categorical=False,
eval_metric='logloss', gamma=0, gpu_id=-1, importance_type=None,
interaction_constraints='', learning_rate=0.3, max_delta_step=0,
max_depth=4, min_child_weight=1,
monotone_constraints='()', n_estimators=30, n_jobs=4,
num_parallel_tree=1, predictor='auto', random_state=0,
reg_alpha=0, reg_lambda=1, scale_pos_weight=1, subsample=1,
tree_method='exact', use_label_encoder=False,
validate_parameters=1, verbosity=None)
model_xgb.fit(X_train, y_train)
# +
best_params= {
'subsample': 0.75,
'reg_lambda': 15.0,
'reg_alpha': 1.3,
'n_estimators': 220,
'min_child_weight': 1.0,
'max_depth': 3,
'learning_rate': .1,
'colsample_bytree': 0.7
}
model_xgb = XGBClassifier(use_label_encoder=False, **best_params)
model_xgb.fit(X_train, y_train)
f1_score(np.ravel(y_ho),
model_xgb.predict(scaler.transform(X_ho)))
# -
import shap
# +
explainer = shap.Explainer(model_xgb)
shap_values = explainer(X_ho)
# visualize the first prediction's explanation
shap.plots.waterfall(shap_values[0])
# -
shap.plots.beeswarm(shap_values)
l = ['flag_brain', 'flag_alpha', 'flag_mu', 'flag_eyes', 'flag_muscles', 'flag_heart', 'flag_ch_noise',
'flag_eyes_blinks', 'flag_muscles_and_movement', 'flag_movement',
'flag_noise', 'flag_uncertain', 'flag_other']
from xgboost import XGBClassifier
model = XGBClassifier(learning_rate=0.3, max_depth=4, n_estimators=30, eval_metric='logloss', use_label_encoder=False)
# 'SVM': SVC(kernel='linear', probability=True)}
model = SVC(kernel='linear', probability=True)
model.fit(X_train, targets[flag])
from sklearn.svm import SVC
# +
# import joblib
# scaler = StandardScaler()
# X_train = scaler.fit_transform(features)
# joblib.dump(scaler, 'scaler.joblib')
for flag in ['flag_brain','flag_eyes']:
# model = LogisticRegression(solver='liblinear', penalty='l2', C=1.0)
model.fit(X_train, targets[flag])
joblib.dump(model, 'svc_' + flag + '.joblib')
# -
| Basic_all_models.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="TGhrHUvK0nzy" colab_type="text"
# # Installation and neccessary imports
# + id="GU_WaXquzzfK" colab_type="code" colab={}
# !pip install torch
# !pip install torchvision
# !pip install Pillow
# !pip install -U -q PyDrive
# !pip install pretrainedmodels
# + id="4gA-Aj9a0SOu" colab_type="code" colab={}
import os
import cv2
import torch
import numpy as np
import pandas as pd
from PIL import Image
import torch.nn as nn
from zipfile import ZipFile
import torch.nn.functional as F
from torchvision import transforms
from torch.utils.data import Dataset, DataLoader
from torch.utils.data.sampler import SubsetRandomSampler
import matplotlib.pyplot as plt
# Ignore warnings
import warnings
warnings.filterwarnings("ignore")
from pydrive.auth import GoogleAuth
from pydrive.drive import GoogleDrive
from google.colab import auth
from oauth2client.client import GoogleCredentials
import torchvision
import torchvision.transforms.functional as TF
import random
# %matplotlib inline
from IPython import display
import pretrainedmodels
# + id="WKAk5dcngG_B" colab_type="code" colab={}
# from PIL import Image
# def register_extension(id, extension): Image.EXTENSION[extension.lower()] = id.upper()
# Image.register_extension = register_extension
# def register_extensions(id, extensions):
# for extension in extensions: register_extension(id, extension)
# Image.register_extensions = register_extensions
# not working :|
# + id="XAcI93TX0YCJ" colab_type="code" colab={}
auth.authenticate_user()
gauth = GoogleAuth()
gauth.credentials = GoogleCredentials.get_application_default()
drive = GoogleDrive(gauth)
# + [markdown] id="ami7WNZc1Erj" colab_type="text"
# ## List the files present in the drive and download the dataset
# + id="ITABPygY1NSv" colab_type="code" colab={}
listed = drive.ListFile().GetList()
for file in listed[:10]:
print('title {}, id {}'.format(file['title'], file['id']))
# + id="8_w2GF_C1WdY" colab_type="code" colab={}
downloaded = drive.CreateFile({'id': '1MwjE6MUY4OnkTytUMTZKa9WskD43_-Al'})
downloaded.GetContentFile('train.zip')
with ZipFile("train.zip", 'r') as z:
z.extractall()
os.remove("train.zip")
# + id="eeH39q0D5BeN" colab_type="code" colab={}
os.listdir()
# + [markdown] id="p51Hkx78Yeon" colab_type="text"
# ## Some utility functions
# + id="Z4QR3kGWcMSH" colab_type="code" colab={}
def normalize(df,cols,dtype = 'float16'):
data = df
for col in cols:
data[col] = ((data[col]-data[col].mean())/(data[col].max()-data[col].min())).astype(dtype)
return data
def display_tensor(x):
if type(x) is tuple:
img,msk = x
fig, axes = plt.subplots(ncols=2,nrows=1)
axes.ravel()[0].imshow(img[0,:,:].cpu().numpy(),cmap='gray')
axes.ravel()[0].set_title('Image')
axes.ravel()[1].imshow(msk[0,:,:].numpy(),cmap='gray')
axes.ravel()[1].set_title('Mask')
plt.show()
else:
plt.imshow(x[0,:,:].cpu().numpy(),cmap='gray')
def adjust_learning_rate(optimizerr,lr):
for param_group in optimizerr.param_groups:
param_group['lr'] = lr
def adjust_momentum(optimizerr,lr):
for param_group in optimizerr.param_groups:
param_group['momentum'] = lr
def save_checkpoint(state, filename='checkpoint.pth.tar'):
auth.authenticate_user()
gauth = GoogleAuth()
gauth.credentials = GoogleCredentials.get_application_default()
drive = GoogleDrive(gauth)
torch.save(state, filename)
uploaded = drive.CreateFile({'title': 'MixNetv2_epoch'+str(state['epoch'])+'.pt'})
uploaded.SetContentFile(filename)
uploaded.Upload()
def load_checkpoint(args):
if os.path.isfile(args):
print("=> loading checkpoint '{}'".format(args))
checkpoint = torch.load(args)
epoch = checkpoint['epoch']
model.load_state_dict(checkpoint['state_dict'])
optimizer.load_state_dict(checkpoint['optimizer'])
print("=> loaded checkpoint '{}' (epoch {})"
.format(args, checkpoint['epoch']))
else:
print("=> no checkpoint found at '{}'".format(args.resume))
# + [markdown] id="WQqHFGhDI3hB" colab_type="text"
# # Dataset Class
# + id="g7cGzjj-I56S" colab_type="code" colab={}
class TGS_dataset(Dataset):
'''
This is the dataset class that feeds the data to the model.
It has been created with a specific file organization in mind as given below:
test.csv
root/
|---train.csv
|---depths.csv
|--train/
| |---images/
| | -images...
| |---masks/
| -masks...
|--test/
| |---images/
| | -images...
| |---masks/
| -masks...
The transform method can be made more efficient while running on test by checking is_train=False and then skipping over mask
'''
def __init__(self,root='./',is_train=True):
depths = normalize(pd.read_csv(root+'depths.csv'), ['z'])
if(is_train):
self.input = pd.read_csv(root+'train.csv').merge(depths, on='id', how='left')
self.input.drop(['rle_mask'], axis=1, inplace=True)
self.image_dir = root+'train/images'
self.mask_dir = root+'train/masks'
else:
self.input = pd.read_csv('test.csv').merge(depths, on='id', how='left')
self.image_dir = root+'test/images'
self.is_train = is_train
def __len__(self):
return len(self.input)
def transform(self,image,mask,depth):
# image, mask = TF.to_grayscale(image), TF.to_grayscale(mask)
mask = TF.to_grayscale(mask)
resize = transforms.Resize((128,128))
image,mask = resize(image), resize(mask)
# Random horizontal flipping
if random.random() > 0.5:
image = TF.hflip(image)
mask = TF.hflip(mask)
# Random vertical flipping
if random.random() > 0.5:
image = TF.vflip(image)
mask = TF.vflip(mask)
# Transform to tensor
image = TF.to_tensor(image)
mask = TF.to_tensor(mask).view(128,128)
# append depth data to image
# d = torch.ones((1)).new_full((1,128,128),depth)
# image = torch.cat((image,d),0)
return image, mask
def __getitem__(self, idx):
img_name = os.path.join(self.image_dir,self.input.iloc[idx,0]+".png")
img = Image.open(img_name)
if self.is_train :
msk = Image.open(os.path.join(self.mask_dir,self.input.iloc[idx,0]+".png"))
else:
msk = img
depth = self.input.iloc[idx,1]
img,msk = self.transform(img,msk,float(depth))
if self.is_train: return img,msk
else : return img
# + id="ALv3mSmeXzkT" colab_type="code" colab={}
train_dataset = TGS_dataset(is_train = True)
# + [markdown] id="c77lFWMVXhEC" colab_type="text"
# # Model Area
# + id="HM2uWdH-Xotg" colab_type="code" colab={}
class Down(nn.Module):
def __init__(self,in_channels,out_channels):
super(Down, self).__init__()
self.down = nn.Sequential(nn.Conv2d(in_channels,out_channels,(3,3),stride=1, padding=1),
nn.AvgPool2d(kernel_size=(2,2)),
nn.BatchNorm2d(out_channels),
nn.ReLU()
)
def forward(self,x):
# print(x.shape)
return self.down(x)
# + id="lZMhEnGQdzoZ" colab_type="code" colab={}
class Up(nn.Module):
def __init__(self,in_channels, out_channels,size):
super(Up,self).__init__()
# self.upscale = nn.Upsample(scale_factor= 2)
self.upscale = nn.ConvTranspose2d(in_channels = in_channels, out_channels = out_channels, kernel_size = (2,2),stride=2)
self.bn = nn.BatchNorm2d(out_channels)
self.out_channels = out_channels
self.nonlin1 = nn.ReLU()
self.nonlin2 = nn.ReLU()
self.size = size
self.noe = size ** 2
# self.bilinear = nn.Bilinear(self.noe ,self.noe ,self.noe )
def forward(self,x1, x2):
x1 = self.upscale(x1)
x1 = self.bn(x1)
x1 = self.nonlin1(x1)
# print(x1.shape)
# print(x2.shape)
x = torch.cat((x1, x2),dim=1)
return x
# + id="7eteCM3sd1kR" colab_type="code" colab={}
class Decoder(nn.Module):
def __init__(self,no_features=2048,in_size=(4,4),out_size=(128,128),in_channels=3):
super(Decoder,self).__init__()
self.down1 = Down(in_channels,32)
self.down2 = Down(32,128)
self.down3 = Down(128,256)
self.down4 = Down(256,512)
self.up1 = Up(2048, 512, 8)
self.up2 = Up(1024, 256, 16)
self.up3 = Up(512, 128, 32)
self.up4 = Up(256, 32, 64)
self.up5 = Up(64,5,128)
def forward(self,img,feats):
x1 = self.down1(img)
x2 = self.down2(x1)
x3 = self.down3(x2)
x4 = self.down4(x3)
x = self.up1(feats, x4)
x = self.up2(x,x3)
x = self.up3(x, x2)
x = self.up4(x,x1)
x = self.up5(x,img)
return x
# + [markdown] id="-xULd5g7RtYE" colab_type="text"
# ### importing Se Resnext 50 trained on imagenet
# + id="a23h4ycMQrIc" colab_type="code" colab={}
class MixNet(nn.Module):
def __init__(self,in_channels = 3, out_channels=1, base= 'se_resnext50_32x4d'):
super(MixNet, self).__init__()
self.resnet = pretrainedmodels.__dict__[base](num_classes=1000, pretrained='imagenet')
self.decoder = Decoder()
self.conv1 = nn.Conv3d(1,1,kernel_size=(8,1,1),stride=(8,1,1))
self.conv2 = nn.Conv2d(1,2,kernel_size=3,padding=1)
def forward(self,x):
feats = self.resnet.features(x)
x = self.decoder(x,feats)
x.unsqueeze_(1)
x = self.conv1(x)
x.squeeze_(1)
x = self.conv2(x)
return F.softmax(x)
# + [markdown] id="zgcG3zpDT37R" colab_type="text"
# ## DataLoader and dataset splitting
# + id="HPrgWtu2wXtU" colab_type="code" colab={}
batch_size = 50
validation_split = 0.1
shuffle_dataset = True
random_seed= 42
# Creating data indices for training and validation splits:
dataset_size = len(train_dataset)
indices = list(range(dataset_size))
split = int(np.floor(validation_split * dataset_size))
if shuffle_dataset :
np.random.seed(random_seed)
np.random.shuffle(indices)
train_indices, val_indices = indices[split:], indices[:split]
# Creating PT data samplers and loaders:
train_sampler = SubsetRandomSampler(train_indices)
validation_sampler = SubsetRandomSampler(val_indices)
train_loader = torch.utils.data.DataLoader(train_dataset,batch_size=batch_size,sampler=train_sampler)
validation_loader = torch.utils.data.DataLoader(train_dataset, batch_size=batch_size,sampler=validation_sampler)
# + id="FBrXCu4jUBQB" colab_type="code" colab={}
# + [markdown] id="uCgMJb_eUByD" colab_type="text"
# ## Some evaluation metrics and Loss functions
# + id="ILpmwaDtwXth" colab_type="code" colab={}
class BinaryCrossEntropyLoss2d(nn.Module):
def __init__(self, weight=None, size_average=True):
"""
Binary cross entropy loss 2D
Args:
weight:
size_average:
"""
super(BinaryCrossEntropyLoss2d, self).__init__()
self.bce_loss = nn.BCELoss(weight, size_average)
if torch.cuda.is_available():
self.bce_loss = self.bce_loss.cuda()
self.threshold = torch.tensor(0.3,requires_grad=True).cuda()
def forward(self, pred, target):
# pred = F.sigmoid(pred)
threshold=0.3
pred1 = ((pred>(1-self.threshold)) * (pred<(1+self.threshold))).float().cuda()
pred1 = pred1.view(-1) # Flatten
target = target.view(-1) # Flatten
return self.bce_loss(pred1, target)
# + id="cLvCr3IQwXtm" colab_type="code" colab={}
class SoftDiceLoss(nn.Module):
def __init__(self):
super(SoftDiceLoss, self).__init__()
def forward(self, pred, target):
smooth = 1
num = target.size(0)
#pred = F.sigmoid(pred)
pred = pred.view(num, -1)
target = target.view(num, -1)
intersection = (pred * target)
score = 2. * (intersection.sum(1) + smooth) / (pred.sum(1) + target.sum(1) + smooth)
score = 1 - score.sum() / num
return score
# + id="9CU7PcdGwXtt" colab_type="code" colab={}
def dice_coeff(pred, target):
smooth = 1.
num = target.size(0)
pred = pred.view(num, -1) # Flatten
target = target.view(num, -1) # Flatten
intersection = (pred * target)
score = (2. * intersection.sum(1) + smooth).float() / (pred.sum(1) + target.sum(1) + smooth).float()
return score.sum()/num
# + id="QcxIxXq5wXtz" colab_type="code" colab={}
def bce_dice_loss(y_true, y_pred):
return 0.5*BinaryCrossEntropyLoss2d()(y_true, y_pred)-dice_coeff(y_true, y_pred)
# + id="lVCCgK0EUNZf" colab_type="code" colab={}
# + [markdown] id="oIV4nJsYUN3Y" colab_type="text"
# # Training
# + id="OL9OQNTawXt-" colab_type="code" colab={}
model=MixNet().float()
criterion = nn.CrossEntropyLoss()
optimizer = torch.optim.SGD(model.parameters(), lr = 0.1, momentum=0.9)
exp_lr_scheduler = torch.optim.lr_scheduler.StepLR(optimizer, step_size=30, gamma=0.1)
if torch.cuda.is_available():
model = model.cuda()
# + id="krdeJDd5wXuL" colab_type="code" colab={}
def validate():
total_loss = 0
total_accuracy = 0
model.eval()
for batch_idx, (data,target) in enumerate(validation_loader):
if torch.cuda.is_available():
data = data.cuda()
target = target.cuda()
target = target.long()
# forward
output = model(data)
predict =torch.argmax(output, dim=1)
# backward + optimize
loss = criterion(output, target)
# print statistics
accuracy = dice_coeff(predict,target)
total_accuracy+=accuracy.item()
total_loss+=loss.item()
print('Validation Loss: {:.5f} Validation Accuracy: {:.5f}'.format(total_loss*batch_size/len(val_indices),total_accuracy*batch_size/len(val_indices)))
return total_loss,total_accuracy
# + id="7ci5neK7wXuU" colab_type="code" colab={}
def train(epoch=1):
loss_train_data = []
accuracy_train_data = []
loss_test_data = []
accuracy_test_data = []
epoch_data=[]
while True:
total_loss = 0
total_accuracy = 0
model.train()
# if epoch%5==0:
# save_checkpoint({
# 'epoch': epoch + 1,
# 'state_dict': model.state_dict(),
# 'optimizer' : optimizer.state_dict(),
# })
exp_lr_scheduler.step()
print(exp_lr_scheduler.get_lr())
for batch_idx, data in enumerate(train_loader):
X,target = data
target = target.long()
if torch.cuda.is_available():
X = X.cuda()
target = target.cuda()
# forward
output = model(X)
predict =torch.argmax(output, dim=1)
# backward + optimize
loss = criterion(output, target)
optimizer.zero_grad()
loss.backward()
optimizer.step()
# print statistics
accuracy = dice_coeff(predict,target)
total_accuracy+=accuracy.item()
total_loss+=loss.item()
print('Epoch: {} [{}/{} ({:.0f}%)]\tLoss: {:.5f}\tAccuracy: {:.5f}'.format(epoch, (batch_idx + 1) * batch_size, len(train_indices),100*(batch_idx + 1)* batch_size / len(train_indices), loss.item(),accuracy))
print('Train Loss: {:.5f} Train Accuracy: {:.5f}'.format(total_loss*batch_size/len(train_indices),total_accuracy*batch_size/len(train_indices)))
loss,accuracy = validate()
# Data append
loss_train_data.append(total_loss*batch_size/len(train_indices))
loss_test_data.append(loss*batch_size/len(val_indices))
accuracy_train_data.append(total_accuracy*batch_size/len(train_indices))
accuracy_test_data.append(accuracy*batch_size/len(val_indices))
epoch_data.append(epoch)
# Visualize
plt.figure(figsize=(10,5))
plt.plot(epoch_data, loss_train_data,label="Train Loss {:.5f}".format(loss_train_data[-1]))
plt.plot(epoch_data, accuracy_train_data,label="Train Accuracy {:.5f}".format(accuracy_train_data[-1]))
plt.plot(epoch_data,loss_test_data, label="Validation Loss {:.5f}".format(loss_test_data[-1]))
plt.plot(epoch_data,accuracy_test_data,label="Validation Accuracy {:.5f}".format(accuracy_test_data[-1]))
display.clear_output(wait=False)
plt.legend()
plt.show()
epoch+=1
# + id="1XQF7jq5wXud" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1614} outputId="38dfb80d-129e-4ed6-b254-209d61fe871e"
train(epoch=84)
# + id="Gxraqy4nfoB_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="d3e4a136-b520-4656-80cb-de24b7e3c73b"
validate()
# + id="qYFODkVXiwGn" colab_type="code" colab={}
save_checkpoint({
'epoch': 70 + 1,
'state_dict': model.state_dict(),
'optimizer' : optimizer.state_dict(),
})
# + id="kXlxdyRI-zCw" colab_type="code" colab={}
adjust_learning_rate(optimizer,0.01)
adjust_momentum(optimizer, 0.1)
# + [markdown] id="DcDHfTFSU7sp" colab_type="text"
# # Training Afterparty (or is there anything to celebrate? )
# + [markdown] id="-N_vCv29Mzzw" colab_type="text"
# ## creating the test dataset
# + id="WvVbhpdIVuPC" colab_type="code" colab={}
#create test
fnames = os.listdir('./test/images')
data = {'id':[]}
for i in fnames:
data['id'].append(i[:-4])
test = pd.DataFrame(data)
test.to_csv('test.csv',index=False)
del data
del fnames
del test
# + id="JIyhBhHgWqfD" colab_type="code" colab={}
batch_size = 30
test_dataset = TGS_dataset(is_train=False)
test_loader = torch.utils.data.DataLoader(test_dataset, batch_size=batch_size)
# + [markdown] id="xi_UdsF8M9xi" colab_type="text"
# ## Transforms to get 101 * 101 image
# + id="QEDjPcUHM8Ne" colab_type="code" colab={}
def mask_from_tensor(x):
img = TF.to_pil_image(x.cpu().type(torch.uint8))
img = TF.to_grayscale(img)
img = TF.resize(img,(101,101))
img = TF.to_tensor(img)
img = img>0.0001
return img
# + id="By6QyI_iWs7x" colab_type="code" colab={}
def rle_encode(img):
'''
img: numpy array, 1 - mask, 0 - background
Returns run length as string formated
'''
pixels = img.flatten()
pixels = np.concatenate([[0], pixels, [0]])
runs = np.where(pixels[1:] != pixels[:-1])[0] + 1
runs[1::2] -= runs[::2]
return ' '.join(str(x) for x in runs)
def predict(test_loader):
rles=[]
model.eval()
for batch_idx,( data) in enumerate(test_loader):
if torch.cuda.is_available():
data = data.cuda()
output = model(data)
predict = torch.argmax(output, dim=1)
predict = predict.long().cpu()
for t in torch.chunk(predict,predict.shape[0]):
t = mask_from_tensor(t)
rles.append(rle_encode(t.squeeze(0).cpu().numpy()))
if batch_idx%15==0:
display.clear_output(wait=False)
print('Progress {:.5f} %'.format(((batch_idx+1)*batch_size*100)/len(test_dataset)))
return rles
# + id="B1gZjbUbKxiV" colab_type="code" colab={}
rle = predict(test_loader)
# + id="k-7ZEcu7L5Db" colab_type="code" colab={}
test_dataset.input['rle_mask'] = rle
# + id="LNBIotAHWwiJ" colab_type="code" colab={}
test_dataset.input.drop(["z"],axis=1,inplace=True)
test_dataset.input.to_csv("submission.csv",index=False)
uploaded = drive.CreateFile({'title': 'submissions_mixnetv2.csv'})
uploaded.SetContentFile('submission.csv')
uploaded.Upload()
# + [markdown] id="Tp6hWpoo0Gj_" colab_type="text"
# # Review
# Big improvement over last time(incorect implamentation)
# This shall serve as our baseline for future experiments
# The model starts to converge with about 0.88 dice score on train and 0.85 on validation
# submission score = 0.454566
#
# The hyperparameters are very much arbitrary and hence there is lot of scope to improve that
# ## Remarks
# - slight overfitting (expected more because of no dropout but random flips and resnext's features are quite balancing the absence of regularization)
#
# - the final conv2d with padding maybe giving wrong values for boundary pixels that make up (4*127)/2^14 or about 3%
#
# ## Future scope
# - lovasz loss
#
# - other pretrained models
#
#
# + [markdown] id="NRdqWVHsXSRm" colab_type="text"
# ## Misc code
# + id="GREExsDIXXdM" colab_type="code" colab={}
# download last saved model to resume training or predict
downloaded = drive.CreateFile({'id': '1uHwN6i_N99GNBdFtKgJI7__n34abrBFz'})
downloaded.GetContentFile('model.dict')
model=MixNet().float()
criterion = nn.CrossEntropyLoss()
optimizer = torch.optim.SGD(model.parameters(), lr = 0.1, momentum=0.9)
epoch=0
# + id="YPRsogSxXip2" colab_type="code" colab={}
load_checkpoint('model.dict')
model.cuda()
# + id="aZH6XcIcYyf7" colab_type="code" colab={}
| notebooks/colab/TGSDetection_baseline_with_submission.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] _uuid="9d7057b0f4ce0d2bb50bfea37c29ce0927cdf53f"
# # Introduction: Manual Feature Engineering
#
# If you are new to this competition, I highly suggest checking out [this notebook](https://www.kaggle.com/willkoehrsen/start-here-a-gentle-introduction/) to get started.
#
# In this notebook, we will explore making features by hand for the Home Credit Default Risk competition. In an earlier notebook, we used only the `application` data in order to build a model. The best model we made from this data achieved a score on the leaderboard around 0.74. In order to better this score, we will have to include more information from the other dataframes. Here, we will look at using information from the `bureau` and `bureau_balance` data. The definitions of these data files are:
#
# * bureau: information about client's previous loans with other financial institutions reported to Home Credit. Each previous loan has its own row.
# * bureau_balance: monthly information about the previous loans. Each month has its own row.
#
# Manual feature engineering can be a tedious process (which is why we use automated feature engineering with featuretools!) and often relies on domain expertise. Since I have limited domain knowledge of loans and what makes a person likely to default, I will instead concentrate of getting as much info as possible into the final training dataframe. The idea is that the model will then pick up on which features are important rather than us having to decide that. Basically, our approach is to make as many features as possible and then give them all to the model to use! Later, we can perform feature reduction using the feature importances from the model or other techniques such as PCA.
#
# The process of manual feature engineering will involve plenty of Pandas code, a little patience, and a lot of great practice manipulation data. Even though automated feature engineering tools are starting to be made available, feature engineering will still have to be done using plenty of data wrangling for a little while longer.
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5"
# pandas and numpy for data manipulation
import pandas as pd
import numpy as np
# matplotlib and seaborn for plotting
import matplotlib.pyplot as plt
import seaborn as sns
# Suppress warnings from pandas
import warnings
warnings.filterwarnings('ignore')
plt.style.use('fivethirtyeight')
# + [markdown] _uuid="b4e0980ff7f4d8d9f0661b5d5ebf07e66d304222"
# ## Example: Counts of a client's previous loans
#
# To illustrate the general process of manual feature engineering, we will first simply get the count of a client's previous loans at other financial institutions. This requires a number of Pandas operations we will make heavy use of throughout the notebook:
#
# * `groupby`: group a dataframe by a column. In this case we will group by the unique client, the `SK_ID_CURR` column
# * `agg`: perform a calculation on the grouped data such as taking the mean of columns. We can either call the function directly (`grouped_df.mean()`) or use the `agg` function together with a list of transforms (`grouped_df.agg([mean, max, min, sum])`)
# * `merge`: match the aggregated statistics to the appropriate client. We need to merge the original training data with the calculated stats on the `SK_ID_CURR` column which will insert `NaN` in any cell for which the client does not have the corresponding statistic
#
# We also use the (`rename`) function quite a bit specifying the columns to be renamed as a dictionary. This is useful in order to keep track of the new variables we create.
#
# This might seem like a lot, which is why we'll eventually write a function to do this process for us. Let's take a look at implementing this by hand first.
# + _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a"
# Read in bureau
bureau = pd.read_csv('../input/bureau.csv')
bureau.head()
# + _uuid="6665d87bd3a157c20fb5a383322aa153005cad2b"
# Groupby the client id (SK_ID_CURR), count the number of previous loans, and rename the column
previous_loan_counts = bureau.groupby('SK_ID_CURR', as_index=False)['SK_ID_BUREAU'].count().rename(columns = {'SK_ID_BUREAU': 'previous_loan_counts'})
previous_loan_counts.head()
# + _uuid="1adcc4acb891adae8646211db629fb263660c2bb"
# Join to the training dataframe
train = pd.read_csv('../input/application_train.csv')
train = train.merge(previous_loan_counts, on = 'SK_ID_CURR', how = 'left')
# Fill the missing values with 0
train['previous_loan_counts'] = train['previous_loan_counts'].fillna(0)
train.head()
# + [markdown] _uuid="53d83da3d8a28541c2dd49be8047e110034c10e6"
# Scroll all the way to the right to see the new column.
# + [markdown] _uuid="173b8548125c8a344f67d397d7c24af223af7254"
# ## Assessing Usefulness of New Variable with r value
#
# To determine if the new variable is useful, we can calculate the Pearson Correlation Coefficient (r-value) between this variable and the target. This measures the strength of a linear relationship between two variables and ranges from -1 (perfectly negatively linear) to +1 (perfectly positively linear). The r-value is not best measure of the "usefulness" of a new variable, but it can give a first approximation of whether a variable will be helpful to a machine learning model. The larger the r-value of a variable with respect to the target, the more a change in this variable is likely to affect the value of the target. Therefore, we look for the variables with the greatest absolute value r-value relative to the target.
#
# We can also visually inspect a relationship with the target using the Kernel Density Estimate (KDE) plot.
# + [markdown] _uuid="f5bc72f178a44d45399f620ee43483df2482c02f"
# ### Kernel Density Estimate Plots
#
# The kernel density estimate plot shows the distribution of a single variable (think of it as a smoothed histogram). To see the different in distributions dependent on the value of a categorical variable, we can color the distributions differently according to the category. For example, we can show the kernel density estimate of the `previous_loan_count` colored by whether the `TARGET` = 1 or 0. The resulting KDE will show any significant differences in the distribution of the variable between people who did not repay their loan (`TARGET == 1`) and the people who did (`TARGET == 0`). This can serve as an indicator of whether a variable will be 'relevant' to a machine learning model.
#
# We will put this plotting functionality in a function to re-use for any variable.
# + _uuid="4a5fab58ab8327c7f53361b584603d78df96c7a4"
# Plots the disribution of a variable colored by value of the target
def kde_target(var_name, df):
# Calculate the correlation coefficient between the new variable and the target
corr = df['TARGET'].corr(df[var_name])
# Calculate medians for repaid vs not repaid
avg_repaid = df.ix[df['TARGET'] == 0, var_name].median()
avg_not_repaid = df.ix[df['TARGET'] == 1, var_name].median()
plt.figure(figsize = (12, 6))
# Plot the distribution for target == 0 and target == 1
sns.kdeplot(df.ix[df['TARGET'] == 0, var_name], label = 'TARGET == 0')
sns.kdeplot(df.ix[df['TARGET'] == 1, var_name], label = 'TARGET == 1')
# label the plot
plt.xlabel(var_name); plt.ylabel('Density'); plt.title('%s Distribution' % var_name)
plt.legend();
# print out the correlation
print('The correlation between %s and the TARGET is %0.4f' % (var_name, corr))
# Print out average values
print('Median value for loan that was not repaid = %0.4f' % avg_not_repaid)
print('Median value for loan that was repaid = %0.4f' % avg_repaid)
# + [markdown] _uuid="783e2b23b473f5f61d17ba5b5b64349d3a1d60cf"
# We can test this function using the `EXT_SOURCE_3` variable which we [found to be one of the most important variables ](https://www.kaggle.com/willkoehrsen/start-here-a-gentle-introduction) according to a Random Forest and Gradient Boosting Machine.
# + _uuid="f9e51eb7f02c5612edef3c950ae89da3b8ececf4"
kde_target('EXT_SOURCE_3', train)
# + [markdown] _uuid="c4598cb54ad22b04646683832e2e4c4bc98ff8e9"
# Now for the new variable we just made, the number of previous loans at other institutions.
# + _uuid="bbae3d4e85f106c1e7b457ad71e1549c0e7b1267"
kde_target('previous_loan_counts', train)
# + [markdown] _uuid="a85c4e3f8ccb54bb4dbbd85a414744d301916fd8"
# From this it's difficult to tell if this variable will be important. The correlation coefficient is extremely weak and there is almost no noticeable difference in the distributions.
#
# Let's move on to make a few more variables from the bureau dataframe. We will take the mean, min, and max of every numeric column in the bureau dataframe.
# + [markdown] _uuid="11f1b0ab2146ec95a50055d2788b5f05c43e0afb"
# ## Aggregating Numeric Columns
#
# To account for the numeric information in the `bureau` dataframe, we can compute statistics for all the numeric columns. To do so, we `groupby` the client id, `agg` the grouped dataframe, and merge the result back into the training data. The `agg` function will only calculate the values for the numeric columns where the operation is considered valid. We will stick to using `'mean', 'max', 'min', 'sum'` but any function can be passed in here. We can even write our own function and use it in an `agg` call.
# + _uuid="582885d850fa1316ce869b856b9d886ad542b04b"
# Group by the client id, calculate aggregation statistics
bureau_agg = bureau.drop(columns = ['SK_ID_BUREAU']).groupby('SK_ID_CURR', as_index = False).agg(['count', 'mean', 'max', 'min', 'sum']).reset_index()
bureau_agg.head()
# + [markdown] _uuid="812f7b1e0d6016768d9276da790b0986181f9fdd"
# We need to create new names for each of these columns. The following code makes new names by appending the stat to the name. Here we have to deal with the fact that the dataframe has a multi-level index. I find these confusing and hard to work with, so I try to reduce to a single level index as quickly as possible.
# + _uuid="01acafad68d285c875018ae916c002e9dad1a2fa"
# List of column names
columns = ['SK_ID_CURR']
# Iterate through the variables names
for var in bureau_agg.columns.levels[0]:
# Skip the id name
if var != 'SK_ID_CURR':
# Iterate through the stat names
for stat in bureau_agg.columns.levels[1][:-1]:
# Make a new column name for the variable and stat
columns.append('bureau_%s_%s' % (var, stat))
# + _uuid="4f34abd311ffcb34e9feb0786998e478ffd90766"
# Assign the list of columns names as the dataframe column names
bureau_agg.columns = columns
bureau_agg.head()
# + [markdown] _uuid="13c6a8b69a3bdbd1bf2d420d984f78400fc969d3"
# Now we simply merge with the training data as we did before.
# + _uuid="2da1e6aa5651f4567813896d79d2e087fd5e7749"
# Merge with the training data
train = train.merge(bureau_agg, on = 'SK_ID_CURR', how = 'left')
train.head()
# + [markdown] _uuid="290cf9aaa44d462f0f987ef35bb22b0010bd01a0"
# ### Correlations of Aggregated Values with Target
#
# We can calculate the correlation of all new values with the target. Again, we can use these as an approximation of the variables which may be important for modeling.
# + _uuid="8cf94533bb8df06623f70d0cb01c24156f8af689"
# List of new correlations
new_corrs = []
# Iterate through the columns
for col in columns:
# Calculate correlation with the target
corr = train['TARGET'].corr(train[col])
# Append the list as a tuple
new_corrs.append((col, corr))
# + [markdown] _uuid="774fe7e4c3e2708de823d3f94f65137b56bc2c2a"
# In the code below, we sort the correlations by the magnitude (absolute value) using the `sorted` Python function. We also make use of an anonymous `lambda` function, another important Python operation that is good to know.
# + _uuid="2802a2568214396e919837353683c63867d38a17"
# Sort the correlations by the absolute value
# Make sure to reverse to put the largest values at the front of list
new_corrs = sorted(new_corrs, key = lambda x: abs(x[1]), reverse = True)
new_corrs[:15]
# + [markdown] _uuid="581f41a665ed3b23d8b28a6e3050a11bc8a68dcf"
# None of the new variables have a significant correlation with the TARGET. We can look at the KDE plot of the highest correlated variable, `bureau_DAYS_CREDIT_mean`, with the target in in terms of absolute magnitude correlation.
# + _uuid="be155ee8fd05290d11a205b95f4b553b8ade06e3"
kde_target('bureau_DAYS_CREDIT_mean', train)
# + [markdown] _uuid="58ca4b2852b486fa10e1bf31c2a7e61e9de52dd7"
# The definition of this column is: "How many days before current application did client apply for Credit Bureau credit". My interpretation is this is the number of days that the previous loan was applied for before the application for a loan at Home Credit. Therefore, a larger negative number indicates the loan was further before the current loan application. We see an extremely weak positive relationship between the average of this variable and the target meaning that clients who applied for loans further in the past potentially are more likely to repay loans at Home Credit. With a correlation this weak though, it is just as likely to be noise as a signal.
#
# #### The Multiple Comparisons Problem
#
# When we have lots of variables, we expect some of them to be correlated just by pure chance, a [problem known as multiple comparisons](https://towardsdatascience.com/the-multiple-comparisons-problem-e5573e8b9578). We can make hundreds of features, and some will turn out to be corelated with the target simply because of random noise in the data. Then, when our model trains, it may overfit to these variables because it thinks they have a relationship with the target in the training set, but this does not necessarily generalize to the test set. There are many considerations that we have to take into account when making features!
# + [markdown] _uuid="645ab51cb64cd20f122d61e17ef8ad1ed78443cc"
# ## Function for Numeric Aggregations
#
# Let's encapsulate all of the previous work into a function. This will allow us to compute aggregate stats for numeric columns across any dataframe. We will re-use this function when we want to apply the same operations for other dataframes.
# + _uuid="8203852b47164d7800f69e572dda2cbe83c50976"
def agg_numeric(df, group_var, df_name):
"""Aggregates the numeric values in a dataframe. This can
be used to create features for each instance of the grouping variable.
Parameters
--------
df (dataframe):
the dataframe to calculate the statistics on
group_var (string):
the variable by which to group df
df_name (string):
the variable used to rename the columns
Return
--------
agg (dataframe):
a dataframe with the statistics aggregated for
all numeric columns. Each instance of the grouping variable will have
the statistics (mean, min, max, sum; currently supported) calculated.
The columns are also renamed to keep track of features created.
"""
# Remove id variables other than grouping variable
for col in df:
if col != group_var and 'SK_ID' in col:
df = df.drop(columns = col)
group_ids = df[group_var]
numeric_df = df.select_dtypes('number')
numeric_df[group_var] = group_ids
# Group by the specified variable and calculate the statistics
agg = numeric_df.groupby(group_var).agg(['count', 'mean', 'max', 'min', 'sum']).reset_index()
# Need to create new column names
columns = [group_var]
# Iterate through the variables names
for var in agg.columns.levels[0]:
# Skip the grouping variable
if var != group_var:
# Iterate through the stat names
for stat in agg.columns.levels[1][:-1]:
# Make a new column name for the variable and stat
columns.append('%s_%s_%s' % (df_name, var, stat))
agg.columns = columns
return agg
# + _uuid="2784423d6e900b66f108cb2e6a425d63eb28743e"
bureau_agg_new = agg_numeric(bureau.drop(columns = ['SK_ID_BUREAU']), group_var = 'SK_ID_CURR', df_name = 'bureau')
bureau_agg_new.head()
# + [markdown] _uuid="171660155dd332d9df85f61fdd7004a9f2593e59"
# To make sure the function worked as intended, we should compare with the aggregated dataframe we constructed by hand.
# + _uuid="57dab1b1b23d0bd0167c9367688b56b43066a721"
bureau_agg.head()
# + [markdown] _uuid="af5a3d2ebc6850707c0bb021bc08df50b98d008b"
# If we go through and inspect the values, we do find that they are equivalent. We will be able to reuse this function for calculating numeric stats for other dataframes. Using functions allows for consistent results and decreases the amount of work we have to do in the future!
#
# ### Correlation Function
#
# Before we move on, we can also make the code to calculate correlations with the target into a function.
# + _uuid="f55cdb9e28c9fcdb16b7922ce90c99ddda590e67"
# Function to calculate correlations with the target for a dataframe
def target_corrs(df):
# List of correlations
corrs = []
# Iterate through the columns
for col in df.columns:
print(col)
# Skip the target column
if col != 'TARGET':
# Calculate correlation with the target
corr = df['TARGET'].corr(df[col])
# Append the list as a tuple
corrs.append((col, corr))
# Sort by absolute magnitude of correlations
corrs = sorted(corrs, key = lambda x: abs(x[1]), reverse = True)
return corrs
# + [markdown] _uuid="a86ad170924ebf66122d24083679d2a061a080ff"
# ## Categorical Variables
#
# Now we move from the numeric columns to the categorical columns. These are discrete string variables, so we cannot just calculate statistics such as mean
# and max which only work with numeric variables. Instead, we will rely on calculating value counts of each category within each categorical variable. As an example, if we have the following dataframe:
#
# | SK_ID_CURR | Loan type |
# |------------|-----------|
# | 1 | home |
# | 1 | home |
# | 1 | home |
# | 1 | credit |
# | 2 | credit |
# | 3 | credit |
# | 3 | cash |
# | 3 | cash |
# | 4 | credit |
# | 4 | home |
# | 4 | home |
#
# we will use this information counting the number of loans in each category for each client.
#
# | SK_ID_CURR | credit count | cash count | home count | total count |
# |------------|--------------|------------|------------|-------------|
# | 1 | 1 | 0 | 3 | 4 |
# | 2 | 1 | 0 | 0 | 1 |
# | 3 | 1 | 2 | 0 | 3 |
# | 4 | 1 | 0 | 2 | 3 |
#
#
# Then we can normalize these value counts by the total number of occurences of that categorical variable for that observation (meaning that the normalized counts must sum to 1.0 for each observation).
#
# | SK_ID_CURR | credit count | cash count | home count | total count | credit count norm | cash count norm | home count norm |
# |------------|--------------|------------|------------|-------------|-------------------|-----------------|-----------------|
# | 1 | 1 | 0 | 3 | 4 | 0.25 | 0 | 0.75 |
# | 2 | 1 | 0 | 0 | 1 | 1.00 | 0 | 0 |
# | 3 | 1 | 2 | 0 | 3 | 0.33 | 0.66 | 0 |
# | 4 | 1 | 0 | 2 | 3 | 0.33 | 0 | 0.66 |
#
# Hopefully, encoding the categorical variables this way will allow us to capture the information they contain. If anyone has a better idea for this process, please let me know in the comments!
# We will now go through this process step-by-step. At the end, we will wrap up all the code into one function to be re-used for many dataframes.
# + [markdown] _uuid="3a8c1a8a74741d727a17ec0e83c237abb931e89e"
# First we one-hot encode a dataframe with only the categorical columns (`dtype == 'object'`).
# + _uuid="cd9fee51331a8181b29867d04b568336aa2bd445"
categorical = pd.get_dummies(bureau.select_dtypes('object'))
categorical['SK_ID_CURR'] = bureau['SK_ID_CURR']
categorical.head()
# + _uuid="839ae477431e4131cfecc3bc0b2046ee964c6623"
categorical_grouped = categorical.groupby('SK_ID_CURR').agg(['sum', 'mean'])
categorical_grouped.head()
# + [markdown] _uuid="ed965c98dee32b0b9194a97c0babd1ade9896e52"
# The `sum` columns represent the count of that category for the associated client and the `mean` represents the normalized count. One-hot encoding makes the process of calculating these figures very easy!
#
# We can use a similar function as before to rename the columns. Again, we have to deal with the multi-level index for the columns. We iterate through the first level (level 0) which is the name of the categorical variable appended with the value of the category (from one-hot encoding). Then we iterate stats we calculated for each client. We will rename the column with the level 0 name appended with the stat. As an example, the column with `CREDIT_ACTIVE_Active` as level 0 and `sum` as level 1 will become `CREDIT_ACTIVE_Active_count`.
# + _uuid="d0348b245ddb0e1f7bfa1e72b5c2e5b04416f66b"
categorical_grouped.columns.levels[0][:10]
# + _uuid="dbfa21f67f3fa910bdb052f482c6cbf684a2e474"
categorical_grouped.columns.levels[1]
# + _uuid="41dccd044bedbf169b761daeff0e5d7b177d08ef"
group_var = 'SK_ID_CURR'
# Need to create new column names
columns = []
# Iterate through the variables names
for var in categorical_grouped.columns.levels[0]:
# Skip the grouping variable
if var != group_var:
# Iterate through the stat names
for stat in ['count', 'count_norm']:
# Make a new column name for the variable and stat
columns.append('%s_%s' % (var, stat))
# Rename the columns
categorical_grouped.columns = columns
categorical_grouped.head()
# + [markdown] _uuid="6ebc592c38c6d7664c7c57d3884ea80dfeabf760"
# The sum column records the counts and the mean column records the normalized count.
#
# We can merge this dataframe into the training data.
# + _uuid="69e0695167a8e793c1a1d3934f8a8e510343606d"
train = train.merge(categorical_grouped, left_on = 'SK_ID_CURR', right_index = True, how = 'left')
train.head()
# + _uuid="072b94cbbc2dba8df21d777e46bc693d731fd512"
train.shape
# + _uuid="28ba0118d6544d6d4fe436901ee9dc8834ae1b0e"
train.iloc[:10, 123:]
# + [markdown] _uuid="a30770ca960821a6c915f6acd54161a37bffb432"
# ### Function to Handle Categorical Variables
#
# To make the code more efficient, we can now write a function to handle the categorical variables for us. This will take the same form as the `agg_numeric` function in that it accepts a dataframe and a grouping variable. Then it will calculate the counts and normalized counts of each category for all categorical variables in the dataframe.
# + _uuid="ddc7decf9497eaa536255d601f76c3a6259fca6b"
def count_categorical(df, group_var, df_name):
"""Computes counts and normalized counts for each observation
of `group_var` of each unique category in every categorical variable
Parameters
--------
df : dataframe
The dataframe to calculate the value counts for.
group_var : string
The variable by which to group the dataframe. For each unique
value of this variable, the final dataframe will have one row
df_name : string
Variable added to the front of column names to keep track of columns
Return
--------
categorical : dataframe
A dataframe with counts and normalized counts of each unique category in every categorical variable
with one row for every unique value of the `group_var`.
"""
# Select the categorical columns
categorical = pd.get_dummies(df.select_dtypes('object'))
# Make sure to put the identifying id on the column
categorical[group_var] = df[group_var]
# Groupby the group var and calculate the sum and mean
categorical = categorical.groupby(group_var).agg(['sum', 'mean'])
column_names = []
# Iterate through the columns in level 0
for var in categorical.columns.levels[0]:
# Iterate through the stats in level 1
for stat in ['count', 'count_norm']:
# Make a new column name
column_names.append('%s_%s_%s' % (df_name, var, stat))
categorical.columns = column_names
return categorical
# + _uuid="d0f1629b07ec6e75e98da459f22611d275fcb282"
bureau_counts = count_categorical(bureau, group_var = 'SK_ID_CURR', df_name = 'bureau')
bureau_counts.head()
# + [markdown] _uuid="cecd442561b3a8405721875878bcbb90c94f2517"
# ### Applying Operations to another dataframe
#
# We will now turn to the bureau balance dataframe. This dataframe has monthly information about each client's previous loan(s) with other financial institutions. Instead of grouping this dataframe by the `SK_ID_CURR` which is the client id, we will first group the dataframe by the `SK_ID_BUREAU` which is the id of the previous loan. This will give us one row of the dataframe for each loan. Then, we can group by the `SK_ID_CURR` and calculate the aggregations across the loans of each client. The final result will be a dataframe with one row for each client, with stats calculated for their loans.
# + _uuid="b49d1c1553eb3c0a8887d967fe9c981b015a7e40"
# Read in bureau balance
bureau_balance = pd.read_csv('../input/bureau_balance.csv')
bureau_balance.head()
# + [markdown] _uuid="815ea915661653e3322e5cbf0f60614754411526"
# First, we can calculate the value counts of each status for each loan. Fortunately, we already have a function that does this for us!
# + _uuid="891d69c6cf3634d2fab71b0c8cc1eb212f433083"
# Counts of each type of status for each previous loan
bureau_balance_counts = count_categorical(bureau_balance, group_var = 'SK_ID_BUREAU', df_name = 'bureau_balance')
bureau_balance_counts.head()
# + [markdown] _uuid="4b1b1623a426efd45f7488636ffffc6b54c68d8e"
# Now we can handle the one numeric column. The `MONTHS_BALANCE` column has the "months of balance relative to application date." This might not necessarily be that important as a numeric variable, and in future work we might want to consider this as a time variable. For now, we can just calculate the same aggregation statistics as previously.
# + _uuid="da531864649dc2b8dab18c31019fc36d6ec6469d"
# Calculate value count statistics for each `SK_ID_CURR`
bureau_balance_agg = agg_numeric(bureau_balance, group_var = 'SK_ID_BUREAU', df_name = 'bureau_balance')
bureau_balance_agg.head()
# + [markdown] _uuid="50b714b2165463138c29715ad68919bcc47aac2d"
# The above dataframes have the calculations done on each _loan_. Now we need to aggregate these for each _client_. We can do this by merging the dataframes together first and then since all the variables are numeric, we just need to aggregate the statistics again, this time grouping by the `SK_ID_CURR`.
# + _uuid="dbd657d3cbe3cf9948ba8d7543f6653e61025bc6"
# Dataframe grouped by the loan
bureau_by_loan = bureau_balance_agg.merge(bureau_balance_counts, right_index = True, left_on = 'SK_ID_BUREAU', how = 'outer')
# Merge to include the SK_ID_CURR
bureau_by_loan = bureau_by_loan.merge(bureau[['SK_ID_BUREAU', 'SK_ID_CURR']], on = 'SK_ID_BUREAU', how = 'left')
bureau_by_loan.head()
# + _uuid="ae22ded2231b37858d9d1d0c3c5b0eb72202c6ab"
bureau_balance_by_client = agg_numeric(bureau_by_loan.drop(columns = ['SK_ID_BUREAU']), group_var = 'SK_ID_CURR', df_name = 'client')
bureau_balance_by_client.head()
# + [markdown] _uuid="fb037a66b921c2dbf837e8d6ddb5179be51e8446"
# To recap, for the `bureau_balance` dataframe we:
#
# 1. Calculated numeric stats grouping by each loan
# 2. Made value counts of each categorical variable grouping by loan
# 3. Merged the stats and the value counts on the loans
# 4. Calculated numeric stats for the resulting dataframe grouping by the client id
#
# The final resulting dataframe has one row for each client, with statistics calculated for all of their loans with monthly balance information.
#
# Some of these variables are a little confusing, so let's try to explain a few:
#
# * `client_bureau_balance_MONTHS_BALANCE_mean_mean`: For each loan calculate the mean value of `MONTHS_BALANCE`. Then for each client, calculate the mean of this value for all of their loans.
# * `client_bureau_balance_STATUS_X_count_norm_sum`: For each loan, calculate the number of occurences of `STATUS` == X divided by the number of total `STATUS` values for the loan. Then, for each client, add up the values for each loan.
# + [markdown] _uuid="e7c7be26bc22cacc8168f907d01c2c259ad6c867"
# We will hold off on calculating the correlations until we have all the variables together in one dataframe.
# + [markdown] _uuid="90aa8e57fbff00777a1e62b060c2c06cf7845381"
# # Putting the Functions Together
#
# We now have all the pieces in place to take the information from the previous loans at other institutions and the monthly payments information about these loans and put them into the main training dataframe. Let's do a reset of all the variables and then use the functions we built to do this from the ground up. This demonstrate the benefit of using functions for repeatable workflows!
# + _uuid="26848bd0252a31559e254c454b905305701ee522"
# Free up memory by deleting old objects
import gc
gc.enable()
del train, bureau, bureau_balance, bureau_agg, bureau_agg_new, bureau_balance_agg, bureau_balance_counts, bureau_by_loan, bureau_balance_by_client, bureau_counts
gc.collect()
# + _uuid="dc574b9486fb40f72146e6481b341931dd180e3d"
# Read in new copies of all the dataframes
train = pd.read_csv('../input/application_train.csv')
bureau = pd.read_csv('../input/bureau.csv')
bureau_balance = pd.read_csv('../input/bureau_balance.csv')
# + [markdown] _uuid="feba956fa449260cc71505cb9156887a62ca8337"
# ### Counts of Bureau Dataframe
# + _uuid="491f45986682beb4ee2bd62c32c622830b3b0e76"
bureau_counts = count_categorical(bureau, group_var = 'SK_ID_CURR', df_name = 'bureau')
bureau_counts.head()
# + [markdown] _uuid="c4c80230ee098fa39f4446b7161f105cd18965bc"
# ### Aggregated Stats of Bureau Dataframe
# + _uuid="5f9d2491e244da4811f78f0340fd9d42127e9076"
bureau_agg = agg_numeric(bureau.drop(columns = ['SK_ID_BUREAU']), group_var = 'SK_ID_CURR', df_name = 'bureau')
bureau_agg.head()
# + [markdown] _uuid="3ce2211679505e646121cc0b2f8b217ab9718ba6"
# ### Value counts of Bureau Balance dataframe by loan
# + _uuid="0e55cc0b41e66e77dfd19ac1b808edc4581a1206"
bureau_balance_counts = count_categorical(bureau_balance, group_var = 'SK_ID_BUREAU', df_name = 'bureau_balance')
bureau_balance_counts.head()
# + [markdown] _uuid="7c2378a4fc130bc7de954c454d50e26adda6f9f6"
# ### Aggregated stats of Bureau Balance dataframe by loan
# + _uuid="eb25294df7f998058e15e6ae24438c772d4ae86c"
bureau_balance_agg = agg_numeric(bureau_balance, group_var = 'SK_ID_BUREAU', df_name = 'bureau_balance')
bureau_balance_agg.head()
# + [markdown] _uuid="f7343f605c4bc38c0c3af1b53bc258699e480dcf"
# ### Aggregated Stats of Bureau Balance by Client
# + _uuid="9315370e38f25339af200a45809387f211464ce9"
# Dataframe grouped by the loan
bureau_by_loan = bureau_balance_agg.merge(bureau_balance_counts, right_index = True, left_on = 'SK_ID_BUREAU', how = 'outer')
# Merge to include the SK_ID_CURR
bureau_by_loan = bureau[['SK_ID_BUREAU', 'SK_ID_CURR']].merge(bureau_by_loan, on = 'SK_ID_BUREAU', how = 'left')
# Aggregate the stats for each client
bureau_balance_by_client = agg_numeric(bureau_by_loan.drop(columns = ['SK_ID_BUREAU']), group_var = 'SK_ID_CURR', df_name = 'client')
# + [markdown] _uuid="1f8f616b645ba974517be24adb30b4af4cf1afe3"
# ## Insert Computed Features into Training Data
# + _uuid="f61f183707d7ca8d363a344b073dd9122acfdcc5"
original_features = list(train.columns)
print('Original Number of Features: ', len(original_features))
# + _uuid="02f4310f7485d299c1588805ce24f3360afa6f42"
# Merge with the value counts of bureau
train = train.merge(bureau_counts, on = 'SK_ID_CURR', how = 'left')
# Merge with the stats of bureau
train = train.merge(bureau_agg, on = 'SK_ID_CURR', how = 'left')
# Merge with the monthly information grouped by client
train = train.merge(bureau_balance_by_client, on = 'SK_ID_CURR', how = 'left')
# + _uuid="f672226eee3059311d476c0eb68946988f9f5712"
new_features = list(train.columns)
print('Number of features using previous loans from other institutions data: ', len(new_features))
# + [markdown] _uuid="e16d0c10cf2a0cc778db576a4ad233960c70a028"
# # Feature Engineering Outcomes
#
# After all that work, now we want to take a look at the variables we have created. We can look at the percentage of missing values, the correlations of variables with the target, and also the correlation of variables with the other variables. The correlations between variables can show if we have collinear varibles, that is, variables that are highly correlated with one another. Often, we want to remove one in a pair of collinear variables because having both variables would be redundant. We can also use the percentage of missing values to remove features with a substantial majority of values that are not present. __Feature selection__ will be an important focus going forward, because reducing the number of features can help the model learn during training and also generalize better to the testing data. The "curse of dimensionality" is the name given to the issues caused by having too many features (too high of a dimension). As the number of variables increases, the number of datapoints needed to learn the relationship between these variables and the target value increases exponentially.
#
# Feature selection is the process of removing variables to help our model to learn and generalize better to the testing set. The objective is to remove useless/redundant variables while preserving those that are useful. There are a number of tools we can use for this process, but in this notebook we will stick to removing columns with a high percentage of missing values and variables that have a high correlation with one another. Later we can look at using the feature importances returned from models such as the `Gradient Boosting Machine` or `Random Forest` to perform feature selection.
# + [markdown] _uuid="ab576d2c245270ca843f825afab02583c091897d"
# ## Missing Values
#
# An important consideration is the missing values in the dataframe. Columns with too many missing values might have to be dropped.
# + _uuid="eef874f5edd28e7449e198264987b93566a5227a"
# Function to calculate missing values by column# Funct
def missing_values_table(df):
# Total missing values
mis_val = df.isnull().sum()
# Percentage of missing values
mis_val_percent = 100 * df.isnull().sum() / len(df)
# Make a table with the results
mis_val_table = pd.concat([mis_val, mis_val_percent], axis=1)
# Rename the columns
mis_val_table_ren_columns = mis_val_table.rename(
columns = {0 : 'Missing Values', 1 : '% of Total Values'})
# Sort the table by percentage of missing descending
mis_val_table_ren_columns = mis_val_table_ren_columns[
mis_val_table_ren_columns.iloc[:,1] != 0].sort_values(
'% of Total Values', ascending=False).round(1)
# Print some summary information
print ("Your selected dataframe has " + str(df.shape[1]) + " columns.\n"
"There are " + str(mis_val_table_ren_columns.shape[0]) +
" columns that have missing values.")
# Return the dataframe with missing information
return mis_val_table_ren_columns
# + _uuid="dba1e683347bbd9b07a162675a4f79e08c28db0a"
missing_train = missing_values_table(train)
missing_train.head(10)
# + [markdown] _uuid="f0defbc704277c67350cf2064b511800a3cdbad2"
# We see there are a number of columns with a high percentage of missing values. There is no well-established threshold for removing missing values, and the best course of action depends on the problem. Here, to reduce the number of features, we will remove any columns in either the training or the testing data that have greater than 90% missing values.
# + _uuid="5739842767f1a58e9c33a6683d5b0f32c97b69b3"
missing_train_vars = list(missing_train.index[missing_train['% of Total Values'] > 90])
len(missing_train_vars)
# + [markdown] _uuid="055b96cb462b6876d6ae7f0e0c348426ddfc5cca"
# Before we remove the missing values, we will find the missing value percentages in the testing data. We'll then remove any columns with greater than 90% missing values in either the training or testing data.
# Let's now read in the testing data, perform the same operations, and look at the missing values in the testing data. We already have calculated all the counts and aggregation statistics, so we only need to merge the testing data with the appropriate data.
# + [markdown] _uuid="dc3921576a8fb644b36f5119f169b88d30a51c01"
# ## Calculate Information for Testing Data
# + _uuid="48be68ba3403d8e3f208b436219003b1f6097fad"
# Read in the test dataframe
test = pd.read_csv('../input/application_test.csv')
# Merge with the value counts of bureau
test = test.merge(bureau_counts, on = 'SK_ID_CURR', how = 'left')
# Merge with the stats of bureau
test = test.merge(bureau_agg, on = 'SK_ID_CURR', how = 'left')
# Merge with the value counts of bureau balance
test = test.merge(bureau_balance_by_client, on = 'SK_ID_CURR', how = 'left')
# + _uuid="0ee6595328bd2eab6a89b542475500aaaa4b3947"
print('Shape of Testing Data: ', test.shape)
# + [markdown] _uuid="518da486b84f28f7ca9c1b366fdb2719f0e19f38"
# We need to align the testing and training dataframes, which means matching up the columns so they have the exact same columns. This shouldn't be an issue here, but when we one-hot encode variables, we need to align the dataframes to make sure they have the same columns.
# + _uuid="39d6e6a2b8af1bb15df0f98ff5eb601c2e828099"
train_labels = train['TARGET']
# Align the dataframes, this will remove the 'TARGET' column
train, test = train.align(test, join = 'inner', axis = 1)
train['TARGET'] = train_labels
# + _uuid="f3d3fcc7cf9a68208a6d6303565883cd58e7c531"
print('Training Data Shape: ', train.shape)
print('Testing Data Shape: ', test.shape)
# + [markdown] _uuid="77cd0cbc96158d555ca156b93906497364cbdabd"
# The dataframes now have the same columns (with the exception of the `TARGET` column in the training data). This means we can use them in a machine learning model which needs to see the same columns in both the training and testing dataframes.
#
# Let's now look at the percentage of missing values in the testing data so we can figure out the columns that should be dropped.
# + _uuid="ff8e3932b44097ae679ea7bb9b0464a077e07727"
missing_test = missing_values_table(test)
missing_test.head(10)
# + _uuid="1ecf51b6455fdef32ea7b50cdaf05eed211b15e7"
missing_test_vars = list(missing_test.index[missing_test['% of Total Values'] > 90])
len(missing_test_vars)
# + _uuid="caa333e30cf19898f4b902d64fbd790659c8f035"
missing_columns = list(set(missing_test_vars + missing_train_vars))
print('There are %d columns with more than 90%% missing in either the training or testing data.' % len(missing_columns))
# + _uuid="3ae7c69dc4112159c5e1356b06821ba169ac595d"
# Drop the missing columns
train = train.drop(columns = missing_columns)
test = test.drop(columns = missing_columns)
# + [markdown] _uuid="ba618f25fe2fb25681228d518cc56f88572cf7fd"
# We ended up removing no columns in this round because there are no columns with more than 90% missing values. We might have to apply another feature selection method to reduce the dimensionality.
# + [markdown] _uuid="91197da37fcd84538520f03b50e0b060a19cc71d"
# At this point we will save both the training and testing data. I encourage anyone to try different percentages for dropping the missing columns and compare the outcomes.
# + _uuid="a7d15638e419b84bf164b20e5195db37885e3e0d"
train.to_csv('train_bureau_raw.csv', index = False)
test.to_csv('test_bureau_raw.csv', index = False)
# + [markdown] _uuid="9425c558df0c882cffe4fc07da41394a6fb3b422"
# ## Correlations
#
# First let's look at the correlations of the variables with the target. We can see in any of the variables we created have a greater correlation than those already present in the training data (from `application`).
# + _uuid="87c47e087eaacb846572ffdd58c22ca1f0ce0010"
# Calculate all correlations in dataframe
corrs = train.corr()
# + _uuid="75c757ae81ff84d9e3453ca7c74ce59326e24b84"
corrs = corrs.sort_values('TARGET', ascending = False)
# Ten most positive correlations
pd.DataFrame(corrs['TARGET'].head(10))
# + _uuid="6774bdef9c97bc58191101414a0a067561f12160"
# Ten most negative correlations
pd.DataFrame(corrs['TARGET'].dropna().tail(10))
# + [markdown] _uuid="0d98ec8d33f097aa3702933cfb64d82bfbd62232"
# The highest correlated variable with the target (other than the `TARGET` which of course has a correlation of 1), is a variable we created. However, just because the variable is correlated does not mean that it will be useful, and we have to remember that if we generate hundreds of new variables, some are going to be correlated with the target simply because of random noise.
#
# Viewing the correlations skeptically, it does appear that several of the newly created variables may be useful. To assess the "usefulness" of variables, we will look at the feature importances returned by the model. For curiousity's sake (and because we already wrote the function) we can make a kde plot of two of the newly created variables.
# + _uuid="c92a7810a7e0e6fd5d92e1be50749e427fae7c11"
kde_target(var_name='client_bureau_balance_counts_mean', df=train)
# + [markdown] _uuid="b8c47b227c6c4130ea9d3454a789b82c422d20a5"
# This variable represents the average number of monthly records per loan for each client. For example, if a client had three previous loans with 3, 4, and 5 records in the monthly data, the value of this variable for them would be 4. Based on the distribution, clients with a greater number of average monthly records per loan were more likely to repay their loans with Home Credit. Let's not read too much into this value, but it could indicate that clients who have had more previous credit history are generally more likely to repay a loan.
# + _uuid="17e1cb7c492b1c122556ac16472fbbdee5a1030a"
kde_target(var_name='bureau_CREDIT_ACTIVE_Active_count_norm', df=train)
# + [markdown] _uuid="dd71a301ca0fe799b5f36ddd07cf3224c03e1cd9"
# Well this distribution is all over the place. This variable represents the number of previous loans with a `CREDIT_ACTIVE` value of `Active` divided by the total number of previous loans for a client. The correlation here is so weak that I do not think we should draw any conclusions!
# + [markdown] _uuid="e6ed8ce92d28415771fe4e4cc872117a18357181"
# ### Collinear Variables
#
# We can calculate not only the correlations of the variables with the target, but also the correlation of each variable with every other variable. This will allow us to see if there are highly collinear variables that should perhaps be removed from the data.
#
# Let's look for any variables that have a greather than 0.8 correlation with other variables.
# + _uuid="7330e7311864a70afa75b7e7c33ad4b3d77df040"
# Set the threshold
threshold = 0.8
# Empty dictionary to hold correlated variables
above_threshold_vars = {}
# For each column, record the variables that are above the threshold
for col in corrs:
above_threshold_vars[col] = list(corrs.index[corrs[col] > threshold])
# + [markdown] _uuid="8dcfa3892d84efa2e0db448b9542c9effad01e85"
# For each of these pairs of highly correlated variables, we only want to remove one of the variables. The following code creates a set of variables to remove by only adding one of each pair.
# + _uuid="0134543670e895436769c64a58bbdee7eb7a255f"
# Track columns to remove and columns already examined
cols_to_remove = []
cols_seen = []
cols_to_remove_pair = []
# Iterate through columns and correlated columns
for key, value in above_threshold_vars.items():
# Keep track of columns already examined
cols_seen.append(key)
for x in value:
if x == key:
next
else:
# Only want to remove one in a pair
if x not in cols_seen:
cols_to_remove.append(x)
cols_to_remove_pair.append(key)
cols_to_remove = list(set(cols_to_remove))
print('Number of columns to remove: ', len(cols_to_remove))
# + [markdown] _uuid="be37ec11c061291e143c01102703103c0915ffcb"
# We can remove these columns from both the training and the testing datasets. We will have to compare performance after removing these variables with performance keeping these variables (the raw csv files we saved earlier).
# + _uuid="0fedcbce9bce99809bf7d4559b6ad93e8ecca8ea"
train_corrs_removed = train.drop(columns = cols_to_remove)
test_corrs_removed = test.drop(columns = cols_to_remove)
print('Training Corrs Removed Shape: ', train_corrs_removed.shape)
print('Testing Corrs Removed Shape: ', test_corrs_removed.shape)
# + _uuid="2f4443a62f6e8987c0be2e3162589444ecd7d275"
train_corrs_removed.to_csv('train_bureau_corrs_removed.csv', index = False)
test_corrs_removed.to_csv('test_bureau_corrs_removed.csv', index = False)
# + [markdown] _uuid="be48572e88f64f00b4221da7782ac834444c96b0"
# # Modeling
#
# To actually test the performance of these new datasets, we will try using them for machine learning! Here we will use a function I developed in another notebook to compare the features (the raw version with the highly correlated variables removed). We can run this kind of like an experiment, and the control will be the performance of just the `application` data in this function when submitted to the competition. I've already recorded that performance,
# so we can list out our control and our two test conditions:
#
# __For all datasets, use the model shown below (with the exact hyperparameters).__
#
# * control: only the data in the `application` files.
# * test one: the data in the `application` files with all of the data recorded from the `bureau` and `bureau_balance` files
# * test two: the data in the `application` files with all of the data recorded from the `bureau` and `bureau_balance` files with highly correlated variables removed.
# + _uuid="58aac4a109bde812ca8b75b1c9271550dcf2d397"
import lightgbm as lgb
from sklearn.model_selection import KFold
from sklearn.metrics import roc_auc_score
from sklearn.preprocessing import LabelEncoder
import gc
import matplotlib.pyplot as plt
# + _uuid="ae2c8a505885482595f027e6139b55966a88861e" code_folding=[0]
def model(features, test_features, encoding = 'ohe', n_folds = 5):
"""Train and test a light gradient boosting model using
cross validation.
Parameters
--------
features (pd.DataFrame):
dataframe of training features to use
for training a model. Must include the TARGET column.
test_features (pd.DataFrame):
dataframe of testing features to use
for making predictions with the model.
encoding (str, default = 'ohe'):
method for encoding categorical variables. Either 'ohe' for one-hot encoding or 'le' for integer label encoding
n_folds (int, default = 5): number of folds to use for cross validation
Return
--------
submission (pd.DataFrame):
dataframe with `SK_ID_CURR` and `TARGET` probabilities
predicted by the model.
feature_importances (pd.DataFrame):
dataframe with the feature importances from the model.
valid_metrics (pd.DataFrame):
dataframe with training and validation metrics (ROC AUC) for each fold and overall.
"""
# Extract the ids
train_ids = features['SK_ID_CURR']
test_ids = test_features['SK_ID_CURR']
# Extract the labels for training
labels = features['TARGET']
# Remove the ids and target
features = features.drop(columns = ['SK_ID_CURR', 'TARGET'])
test_features = test_features.drop(columns = ['SK_ID_CURR'])
# One Hot Encoding
if encoding == 'ohe':
features = pd.get_dummies(features)
test_features = pd.get_dummies(test_features)
# Align the dataframes by the columns
features, test_features = features.align(test_features, join = 'inner', axis = 1)
# No categorical indices to record
cat_indices = 'auto'
# Integer label encoding
elif encoding == 'le':
# Create a label encoder
label_encoder = LabelEncoder()
# List for storing categorical indices
cat_indices = []
# Iterate through each column
for i, col in enumerate(features):
if features[col].dtype == 'object':
# Map the categorical features to integers
features[col] = label_encoder.fit_transform(np.array(features[col].astype(str)).reshape((-1,)))
test_features[col] = label_encoder.transform(np.array(test_features[col].astype(str)).reshape((-1,)))
# Record the categorical indices
cat_indices.append(i)
# Catch error if label encoding scheme is not valid
else:
raise ValueError("Encoding must be either 'ohe' or 'le'")
print('Training Data Shape: ', features.shape)
print('Testing Data Shape: ', test_features.shape)
# Extract feature names
feature_names = list(features.columns)
# Convert to np arrays
features = np.array(features)
test_features = np.array(test_features)
# Create the kfold object
k_fold = KFold(n_splits = n_folds, shuffle = False, random_state = 50)
# Empty array for feature importances
feature_importance_values = np.zeros(len(feature_names))
# Empty array for test predictions
test_predictions = np.zeros(test_features.shape[0])
# Empty array for out of fold validation predictions
out_of_fold = np.zeros(features.shape[0])
# Lists for recording validation and training scores
valid_scores = []
train_scores = []
# Iterate through each fold
for train_indices, valid_indices in k_fold.split(features):
# Training data for the fold
train_features, train_labels = features[train_indices], labels[train_indices]
# Validation data for the fold
valid_features, valid_labels = features[valid_indices], labels[valid_indices]
# Create the model
model = lgb.LGBMClassifier(n_estimators=10000, objective = 'binary',
class_weight = 'balanced', learning_rate = 0.05,
reg_alpha = 0.1, reg_lambda = 0.1,
subsample = 0.8, n_jobs = -1, random_state = 50)
# Train the model
model.fit(train_features, train_labels, eval_metric = 'auc',
eval_set = [(valid_features, valid_labels), (train_features, train_labels)],
eval_names = ['valid', 'train'], categorical_feature = cat_indices,
early_stopping_rounds = 100, verbose = 200)
# Record the best iteration
best_iteration = model.best_iteration_
# Record the feature importances
feature_importance_values += model.feature_importances_ / k_fold.n_splits
# Make predictions
test_predictions += model.predict_proba(test_features, num_iteration = best_iteration)[:, 1] / k_fold.n_splits
# Record the out of fold predictions
out_of_fold[valid_indices] = model.predict_proba(valid_features, num_iteration = best_iteration)[:, 1]
# Record the best score
valid_score = model.best_score_['valid']['auc']
train_score = model.best_score_['train']['auc']
valid_scores.append(valid_score)
train_scores.append(train_score)
# Clean up memory
gc.enable()
del model, train_features, valid_features
gc.collect()
# Make the submission dataframe
submission = pd.DataFrame({'SK_ID_CURR': test_ids, 'TARGET': test_predictions})
# Make the feature importance dataframe
feature_importances = pd.DataFrame({'feature': feature_names, 'importance': feature_importance_values})
# Overall validation score
valid_auc = roc_auc_score(labels, out_of_fold)
# Add the overall scores to the metrics
valid_scores.append(valid_auc)
train_scores.append(np.mean(train_scores))
# Needed for creating dataframe of validation scores
fold_names = list(range(n_folds))
fold_names.append('overall')
# Dataframe of validation scores
metrics = pd.DataFrame({'fold': fold_names,
'train': train_scores,
'valid': valid_scores})
return submission, feature_importances, metrics
# + _uuid="6b39fe9584e0308b7e940c11e6de04b262d123c7" code_folding=[0]
def plot_feature_importances(df):
"""
Plot importances returned by a model. This can work with any measure of
feature importance provided that higher importance is better.
Args:
df (dataframe): feature importances. Must have the features in a column
called `features` and the importances in a column called `importance
Returns:
shows a plot of the 15 most importance features
df (dataframe): feature importances sorted by importance (highest to lowest)
with a column for normalized importance
"""
# Sort features according to importance
df = df.sort_values('importance', ascending = False).reset_index()
# Normalize the feature importances to add up to one
df['importance_normalized'] = df['importance'] / df['importance'].sum()
# Make a horizontal bar chart of feature importances
plt.figure(figsize = (10, 6))
ax = plt.subplot()
# Need to reverse the index to plot most important on top
ax.barh(list(reversed(list(df.index[:15]))),
df['importance_normalized'].head(15),
align = 'center', edgecolor = 'k')
# Set the yticks and labels
ax.set_yticks(list(reversed(list(df.index[:15]))))
ax.set_yticklabels(df['feature'].head(15))
# Plot labeling
plt.xlabel('Normalized Importance'); plt.title('Feature Importances')
plt.show()
return df
# + [markdown] _uuid="52cc81edc0b29905d2325f201f7902e96c321e32"
# ### Control
#
# The first step in any experiment is establishing a control. For this we will use the function defined above (that implements a Gradient Boosting Machine model) and the single main data source (`application`).
# + _uuid="7b9f0e5faa3de9403f87032fe3fc339cb732ad17"
train_control = pd.read_csv('../../input/application_train.csv')
test_control = pd.read_csv('../../input/application_test.csv')
# + [markdown] _uuid="74c09643eee73b19d80acfca60dcd33df8747364"
# Fortunately, once we have taken the time to write a function, using it is simple (if there's a central theme in this notebook, it's use functions to make things simpler and reproducible!). The function above returns a `submission` dataframe we can upload to the competition, a `fi` dataframe of feature importances, and a `metrics` dataframe with validation and test performance.
# + _uuid="3711f26b43863952e4edba7c3fc33f877ed499b2"
submission, fi, metrics = model(train_control, test_control)
# + _uuid="fed9ad5478719de194aeabec9b151f21effb99f5"
metrics
# + [markdown] _uuid="4f3382baf72ccfc101660b2f062fee707e7cb041"
# The control slightly overfits because the training score is higher than the validation score. We can address this in later notebooks when we look at regularization (we already perform some regularization in this model by using `reg_lambda` and `reg_alpha` as well as early stopping).
#
# We can visualize the feature importance with another function, `plot_feature_importances`. The feature importances may be useful when it's time for feature selection.
# + _uuid="c09f30bd4fb6e12b55d5a288fab358ffd1fe6f4d"
fi_sorted = plot_feature_importances(fi)
# + _uuid="6fe65cc25940a40c9966101acf9ea8f8477d7b77"
submission.to_csv('../../output/control.csv', index = False)
# + [markdown] _uuid="e502ef191ea2201b05f85cece2202931744fa539"
# __The control scores 0.745 when submitted to the competition.__
# + [markdown] _uuid="392da17505a7feb3320242ff9785ac81346ee0b7"
# ### Test One
#
# Let's conduct the first test. We will just need to pass in the data to the function, which does most of the work for us.
# + _uuid="640bceff24f8c53aed68568c6ee1f70575adab69"
submission_raw, fi_raw, metrics_raw = model(train, test)
# + _uuid="b404e436782d6de379bfa4756d58bed0175396f8"
metrics_raw
# + [markdown] _uuid="0fcb737753ca178e32afda3fe29bf4c29262c379"
# Based on these numbers, the engineered features perform better than the control case. However, we will have to submit the predictions to the leaderboard before we can say if this better validation performance transfers to the testing data.
# + _uuid="3313e261d947645ca0e72f824de45827a43de9c1"
fi_raw_sorted = plot_feature_importances(fi_raw)
# + [markdown] _uuid="3bf21f18e6430cea006cdd6b0139c52cb760d6ee"
# Examining the feature improtances, it looks as if a few of the feature we constructed are among the most important. Let's find the percentage of the top 100 most important features that we made in this notebook. However, rather than just compare to the original features, we need to compare to the _one-hot encoded_ original features. These are already recorded for us in `fi` (from the original data).
# + _uuid="b5abb37ffe89b767a01f494801c4db5bc3abc6db"
top_100 = list(fi_raw_sorted['feature'])[:100]
new_features = [x for x in top_100 if x not in list(fi['feature'])]
print('%% of Top 100 Features created from the bureau data = %d.00' % len(new_features))
# + [markdown] _uuid="4ae9e0771495bdfff89f221f435c5320c8fd27f0"
# Over half of the top 100 features were made by us! That should give us confidence that all the hard work we did was worthwhile.
# + _uuid="0fe38b12afa98f44ad2a1e20da417f3e1f3728fd"
submission_raw.to_csv('test_one.csv', index = False)
# + [markdown] _uuid="9bff36a22e3ee286f046f819bcca121c14df27a8"
# __Test one scores 0.759 when submitted to the competition.__
# + [markdown] _uuid="9469d920abeeb1c912391b06c1f4bd0175c15780"
# ### Test Two
#
# That was easy, so let's do another run! Same as before but with the highly collinear variables removed.
# + _uuid="008b1859c010277339ac2b1c2b52c0516c001774"
submission_corrs, fi_corrs, metrics_corr = model(train_corrs_removed, test_corrs_removed)
# + _uuid="6f91e580802f054053d497495de1c7aa4f4047eb"
metrics_corr
# + [markdown] _uuid="c1c840d66cb195848d861133e24be78514624f30"
# These results are better than the control, but slightly lower than the raw features.
# + _uuid="ed8a71b2d72719418dcb9463dea983dfe2b1ade8"
fi_corrs_sorted = plot_feature_importances(fi_corrs)
# + _uuid="521e22fc1ba905c4851ca2d948260298a7527519"
submission_corrs.to_csv('test_two.csv', index = False)
# + [markdown] _uuid="5cca331fe96700aa942ef56f96ce2fa34ad1933a"
# __Test Two scores 0.753 when submitted to the competition.__
# + [markdown] _uuid="457467aab25e8c047a2c74a1392730b96203cfa2"
# # Results
#
# After all that work, we can say that including the extra information did improve performance! The model is definitely not optimized to our data, but we still had a noticeable improvement over the original dataset when using the calculated features. Let's officially summarize the performances:
#
# | __Experiment__ | __Train AUC__ | __Validation AUC__ | __Test AUC__ |
# |------------|-------|------------|-------|
# | __Control__ | 0.815 | 0.760 | 0.745 |
# | __Test One__ | 0.837 | 0.767 | 0.759 |
# | __Test Two__ | 0.826 | 0.765 | 0.753 |
#
#
# (Note that these scores may change from run to run of the notebook. I have not observed that the general ordering changes however.)
#
# All of our hard work translates to a small improvement of 0.014 ROC AUC over the original testing data. Removing the highly collinear variables slightly decreases performance so we will want to consider a different method for feature selection. Moreover, we can say that some of the features we built are among the most important as judged by the model.
#
# In a competition such as this, even an improvement of this size is enough to move us up 100s of spots on the leaderboard. By making numerous small improvements such as in this notebook, we can gradually achieve better and better performance. I encourage others to use the results here to make their own improvements, and I will continue to document the steps I take to help others.
#
# ## Next Steps
#
# Going forward, we can now use the functions we developed in this notebook on the other datasets. There are still 4 other data files to use in our model! In the next notebook, we will incorporate the information from these other data files (which contain information on previous loans at Home Credit) into our training data. Then we can build the same model and run more experiments to determine the effect of our feature engineering. There is plenty more work to be done in this competition, and plenty more gains in performance to be had! I'll see you in the next notebook.
# + _uuid="94bdea7d6198e5cf61f073c3c4d0386d5961b564"
| automaticFeatureEngineering/Introduction to Manual Feature Engineering.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
from matplotlib import pyplot as plt
# CSV pandas
df: pd.DataFrame = pd.DataFrame(np.array([["Lithium","Argon","Potassium","Calcium","Scandium","Vanadium","Strontium"],
[0.53,1.78,0.86,1.6,3.0,6.11,2.54],
[4.0753,6.6678,1.5177,3.6375,4.7243,9.0698,5.3002],
[np.nan,2.1328,3.6852,8.5389,10.1570,2.8739,4.4508]])).T
df.columns = ["Element","Density","Volume1","Volume2"]
# Save as CSV
df.to_csv("data/matlab_onroad_11_2_element.csv",index=False)
# load CSV to Dataframe
elements: pd.DataFrame = pd.read_csv("data/matlab_onroad_11_2_element.csv")
elements
# Task 1 Assign the contents of elements.Density to a column vector named d.
d: np.array = np.array(elements.Density)
d
# +
# Task 2
# Multiply each element of elements.Density with elements.Volume1. Remember to use elementwise multiplication with .*.
# Assign the result to elements.Mass.
elements["Mass"] = elements["Density"] * elements["Volume1"]
# Task 3 Sort the table by smallest to largest mass.
elements.sort_values("Mass")
# -
| notebooks/MATLAB OnRoad 11.2 Importing Data as a Table.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="QKNMsGnmoonF"
# # Comparación de más de dos medias
# # Vamos a comprarar los datos de ventas dependiendo de las estaciones
# + [markdown] id="ECkQFJfdoqn5"
# Importamos librerias
# + [markdown] id="bSrAunyLom8o"
#
# + id="UtTGWv6cnarr"
from google.colab import drive # For linking colab to Google Drive
import pandas as pd # Datasets
import numpy as np # Vectores, matrices
import matplotlib.pyplot as plt # Hacer gráficos
import scipy.stats as stats # Estadística
import seaborn as sns # Gráficos
from pandas.api.types import CategoricalDtype # Para variables ordinales
# + [markdown] id="pLmZOrBfo0AZ"
# Montamos nuestro Google Drive como una unidad para cargar y guardar archivos
# + colab={"base_uri": "https://localhost:8080/"} id="exQs36vwozcO" outputId="438dcb87-7df2-4841-9f58-0e6d5de42339"
drive.mount('mydrive')
# + [markdown] id="cip1ryJ-pJXJ"
# Cargamos el archivo con los datos de nuestro Google Drive y lo vemos
# + colab={"base_uri": "https://localhost:8080/", "height": 417} id="jc6Y_FQDpNkn" outputId="c437c751-9845-49a1-a069-c52bfb081328"
path = r'/content/mydrive/Othercomputers/portatil/Master/AnalisisDatos/Ejercicio_12nov_2/WBR_11_12_denormalized_temp.csv'
wbr= pd.read_csv(path, sep = ';', decimal = ',')
wbr
# + [markdown] id="i37IpN61qHxT"
# La columna con las estaciones está codificada con números, vamos a hacer una nueva pero usando texto
# + id="nLvnCU6apfeR"
wbr.loc[(wbr['season'] == 1), 'season_cat'] = 'Winter'
wbr.loc[(wbr['season'] == 2), 'season_cat'] = 'Spring'
wbr.loc[(wbr['season'] == 3), 'season_cat'] = 'Summer'
wbr.loc[(wbr['season'] == 4), 'season_cat'] = 'Autumn'
# + [markdown] id="8mFMbVtwp6-_"
# Comprobamos que se ha hecho bien el cambio
# + colab={"base_uri": "https://localhost:8080/", "height": 203} id="TOvFVisKp6i3" outputId="2fe014af-7709-483f-ccae-f7de17481e13"
pd.crosstab(wbr.season, wbr.season_cat)
# + [markdown] id="2ZFvon3rqT2t"
# A partir de esta columna creamos otra pero conviertiendo sus datos en ordinales.
# De esta manera, al hacer un gráfico se hará con el orden que marquemos
# + id="Cu37NO7kqWVb"
# Primero definimos la lista y orden de las categorias
my_categories = ['Spring', 'Summer', 'Autumn', 'Winter']
# Definimos el tipo de dato con la lista de categorias diciendole que están en orden
season_type = CategoricalDtype(categories = my_categories, ordered = True)
# Creamos una nueva columna del tiempo pero con variables de tipo ordinal
wbr['season_cat_ord'] = wbr.season_cat.astype(season_type)
# + [markdown] id="7F5Ik8WKqqpV"
# Dibujamos que porcentaje de días hay para cada estación
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="8ra5WLHQqtoq" outputId="d6152855-38a4-4f47-9f24-6cfc7bc5f893"
mytable = pd.crosstab(index = wbr["season_cat_ord"], columns="count")
n=mytable.sum()
mytable2 = (mytable/n)*100
plt.bar(mytable2.index, mytable2['count'])
plt.show()
# + id="t5K7zkHArAbC"
Agrupamos los datos de venta por estación
# + id="N8VsCAdGrDg5"
cnt_spring = wbr.loc[wbr.season_cat_ord == 'Spring', 'cnt']
cnt_summer = wbr.loc[wbr.season_cat_ord == 'Summer', 'cnt']
cnt_autumn = wbr.loc[wbr.season_cat_ord == 'Autumn', 'cnt']
cnt_winter = wbr.loc[wbr.season_cat_ord == 'Winter', 'cnt']
# + [markdown] id="yPglLq23rPdF"
# Hacemos un test anova para las medias de los datos por estación.
# Primero obtenemos el valor t
# + colab={"base_uri": "https://localhost:8080/"} id="iapietHhrO6x" outputId="7e9da0d5-095b-45a4-f52d-121c7296407c"
stats.f_oneway(cnt_spring, cnt_summer, cnt_autumn, cnt_winter)[0]
# + [markdown] id="keVd3wx1rW9v"
# Ahora obtenemos el p value
# + colab={"base_uri": "https://localhost:8080/"} id="anMnpmyTrZ90" outputId="1892eb75-136d-4adc-abef-a2ee728f85ff"
stats.f_oneway(cnt_spring, cnt_summer, cnt_autumn, cnt_winter)[1]
# + [markdown] id="Y1yoE9mAreI2"
# El pvalue siendo menos que 0.05 nos hace rechazar la hipotesis nula de que todas las medias son iguales. Hay por tanto diferencia de medias entre todas las medias o en una media con las otras
# + [markdown] id="01ScxzHMrma3"
# Vemos la media de todas las estaciones
# + colab={"base_uri": "https://localhost:8080/"} id="SxqMZPgirog6" outputId="56d6d52a-a0be-4be4-be34-adeb16028997"
wbr.groupby('season_cat_ord').cnt.mean()
# + [markdown] id="kjFhQFn6rzle"
# Vemos la media total para usarla en el gráfico
# + colab={"base_uri": "https://localhost:8080/"} id="hLD3QPDZr2V9" outputId="55007eda-ec02-4e27-c780-d76763360819"
wbr.cnt.mean()
# + [markdown] id="tKMa5zAPr3gQ"
# Vemos ahora también para el gráfico el total de casos
# + colab={"base_uri": "https://localhost:8080/"} id="YyNfthVFr7Xy" outputId="7f10bf64-47b9-4ad5-c53f-bd4c5a699715"
wbr.cnt.count()
# + [markdown] id="2HypwLa8sC9f"
# Ahora dibujamos un gráfico con las medias e intervalos de confianza para cada estación.
# + colab={"base_uri": "https://localhost:8080/", "height": 256} id="S4VADiHksMOe" outputId="5c600073-3df5-4c95-b995-31e19ed1baca"
# Elegimos el tamaño del gráfico
plt.figure(figsize=(5,3))
# Creamos el gráfico
ax = sns.pointplot(x="season_cat_ord", y="cnt", data=wbr,ci=99, join=0)
# Vamos a poner cuando poner los ticks del eje y y el rango de este
plt.yticks(np.arange(1000, 7000, step=500))
plt.ylim(1000,6200)
# Ponemos una línea horizontal en la media total
plt.axhline(y=wbr.cnt.mean(), linewidth=1, linestyle= 'dashed', color="blue")
# Vamos a añadir un cuadro con algunos datos: Ponemos la media total, el numero de datos, el pvalue y el valor de t que es otro dato de interés
props = dict(boxstyle = 'round', facecolor= 'white', lw=0.5)
plt.text(2.4,4500,'Mean:4504.3''\n''n:731' '\n' 't:1.601' '\n' 'Pval.:0.110', bbox=props)
# Nombre del eje x
plt.xlabel('Weather type')
# Título del gráfico
plt.title('Average rentals by season.''\n')
plt.show()
| Fundamentos/AnalisisDatos/Ejercicio_19nov_6/19_nov_6.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="fyiTgkiLIoc-" outputId="925175d1-dfd2-402f-978b-931f08f92bba"
import torch
import torch.nn.functional as F
import torch.nn as nn
import torchvision
import torchvision.datasets as datasets
import matplotlib.pyplot as plt
import numpy as np
import pickle
from torch import Tensor
import torch.optim as optim
from torch.utils.data import DataLoader, TensorDataset
import matplotlib.pyplot as plt
import pandas as pd
from sklearn import preprocessing
from sklearn.model_selection import train_test_split
from scipy.fft import rfft, rfftfreq, fft, fftfreq
import scipy
import time
import copy
from google.colab import drive
drive.mount('/content/drive')
# Load in Data
with open('/content/drive/MyDrive/CSE 481 Capstone/processed_data.npy', 'rb') as f:
data = np.load(f)
print(data.shape)
# + colab={"base_uri": "https://localhost:8080/"} id="anBhQSNuJIME" outputId="1ea3b3a1-4109-48dd-8214-4013cae4f352"
# Load in Labels
with open('/content/drive/MyDrive/CSE 481 Capstone/labels.npy', 'rb') as f:
labels = np.load(f)
sub_labels = labels
print(sub_labels.shape)
print(sub_labels)
# convert to windowed labels
data_labels = np.repeat(sub_labels, 14, axis=0)
print(data_labels.shape)
print(data_labels)
#convert to tensor
#ata = torch.tensor(data)
data = torch.from_numpy(data).float()
data_labels = torch.from_numpy(data_labels).float()
print(data.dtype)
print(data.shape)
dataset = TensorDataset(Tensor(data) , Tensor(data_labels))
print(data[29][1])
# + colab={"base_uri": "https://localhost:8080/"} id="RwaUThpnVp1T" outputId="96540012-3a74-4900-a0e7-e22eef6e6c41"
pre_train_size = int(0.9 * len(dataset))
test_size = len(dataset) - pre_train_size
pre_train_set, test_set = torch.utils.data.random_split(dataset, [pre_train_size, test_size])
train_size = int(0.8 * len(pre_train_set))
val_size = len(pre_train_set) - train_size
print(len(pre_train_set))
print(train_size)
print(val_size)
train_set, val_set = torch.utils.data.random_split(pre_train_set, [train_size, val_size])
batch_size = 128
print(len(train_set), len(val_set), len(test_set))
trainloader = torch.utils.data.DataLoader(train_set, batch_size=batch_size,
shuffle=True, num_workers=2)
valloader = torch.utils.data.DataLoader(val_set, batch_size=batch_size,
shuffle=True, num_workers=2)
testloader = torch.utils.data.DataLoader(test_set, batch_size=batch_size,
shuffle=False, num_workers=2)
dataloaders = {
'train': trainloader,
'val': valloader,
}
# + colab={"base_uri": "https://localhost:8080/"} id="6lDcUw-cbcuO" outputId="3ea1945c-adbc-4425-a58c-1eb33b6c54f6"
# with auto encoder
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
print(device)
# skipped autoencoder
model = nn.Sequential(
nn.Conv2d(4, 32, [3, 1]),
nn.ReLU(),
nn.Dropout(),
nn.Conv2d(32, 64, [3, 1]),
nn.ReLU(), # Maybe not sure
nn.Dropout(),
nn.MaxPool2d([3, 3]),
nn.Flatten(),
nn.Linear(5760, 2048),
nn.ReLU(),
nn.Linear(2048, 1024),
nn.ReLU(),
nn.Linear(1024, 512),
nn.ReLU(),
nn.Linear(512, 256),
nn.ReLU(),
nn.Linear(256, 128),
nn.ReLU(),
nn.Linear(128, 2)
)
model.to(device)
loss_func = nn.MSELoss()
optimizer = torch.optim.Adam(model.parameters(), lr=0.001)
print(model)
# + id="8f86YR4I89Km"
def accuracy(input):
output = np.zeros((len(input)))
for i in range(len(input)):
instance = input[i]
valence = instance[0]
arousal = instance[1]
if (valence < 5 and arousal < 5):
output[i] = 0
elif (valence < 5 and arousal >= 5):
output[i] = 1
elif (valence >= 5 and arousal < 5):
output[i] = 2
else:
output[i] = 3
return output
# + colab={"base_uri": "https://localhost:8080/"} id="r29Sozai8viu" outputId="8f15551e-5b73-4009-d379-0b6a4823fc84"
since = time.time()
best_model_wts = copy.deepcopy(model.state_dict())
best_loss = 10000.0
all_train_loss = []
all_val_loss = []
num_epochs = 351
for epoch in range(num_epochs):
print('Epoch {}/{}'.format(epoch, num_epochs - 1))
print('-' * 10)
# Each epoch has a training and validation phase
for phase in ['train', 'val']:
if phase == 'train':
model.train() # Set model to training mode
else:
model.eval() # Set model to evaluate mode
running_loss = 0.0
running_corrects = 0
# Iterate over data.
for inputs, labels in dataloaders[phase]:
inputs = inputs.to(device)
labels = labels.to(device)
#print(inputs)
#print(labels)
# zero the parameter gradients
optimizer.zero_grad()
# forward
# track history if only in train
with torch.set_grad_enabled(phase == 'train'):
outputs = model(inputs)
#print(outputs)
#_, preds = torch.max(outputs, 1)
#print(preds)
#print(outputs.dtype)
#print(labels.dtype)
loss = loss_func(outputs, labels)
#print(loss.item())
# backward + optimize only if in training phase
if phase == 'train':
loss.backward()
optimizer.step()
# statistics
running_loss += loss.item()
#print(running_loss)
#print(outputs)
#print(accuracy(outputs))
#print(accuracy(labels))
if (epoch % 50 == 0):
running_corrects += np.sum(accuracy(outputs) == accuracy(labels)) / inputs.size(0)
epoch_loss = running_loss / len(dataloaders[phase])
#epoch_acc = running_corrects / len(dataloaders[phase])
print('{} Loss: {:.4f}'.format(
phase, epoch_loss))
if (epoch % 50 == 0):
print('Acc: {:.4f}'.format(running_corrects / len(dataloaders[phase])))
if phase == 'train':
all_train_loss.append(epoch_loss)
else:
all_val_loss.append(epoch_loss)
# deep copy the model
# if phase == 'val' and epoch_acc > best_acc:
# best_acc = epoch_acc
# best_model_wts = copy.deepcopy(model.state_dict())
if phase == 'val' and epoch_loss < best_loss:
best_loss = epoch_loss
best_model_wts = copy.deepcopy(model.state_dict())
print()
time_elapsed = time.time() - since
print('Training complete in {:.0f}m {:.0f}s'.format(
time_elapsed // 60, time_elapsed % 60))
print('Best Val Loss: {:4f}'.format(best_loss))
# load best model weights
model.load_state_dict(best_model_wts)
# + colab={"base_uri": "https://localhost:8080/", "height": 506} id="Q54ZGAok6wzI" outputId="f456b65e-674c-4f7b-f06b-5ceb871331a8"
plt.plot(all_train_loss)
plt.plot(all_val_loss)
plt.legend('train', 'val')
# + colab={"base_uri": "https://localhost:8080/"} id="0oGxf1v3Y1ui" outputId="32968ecc-24d0-4bd4-eab4-a32ea90f815c"
running_loss = 0.0
running_corrects = 0
model.eval()
# Iterate over data.
for inputs, labels in testloader:
inputs = inputs.to(device)
labels = labels.to(device)
outputs = model(inputs)
loss = loss_func(outputs, labels)
# statistics
running_loss += loss.item()
running_corrects += np.sum(accuracy(outputs) == accuracy(labels)) / inputs.size(0)
epoch_loss = running_loss / len(testloader)
print('Test Loss: {:.4f}'.format(epoch_loss))
print('Test Acc: {:.4f}'.format(running_corrects / len(testloader)))
print()
# + colab={"base_uri": "https://localhost:8080/"} id="BKC3H70XJUMg" outputId="67817389-76a6-44a7-bd15-3a044af3afb8"
model.eval()
datatestiter = iter(testloader)
input_test, labels_test = datatestiter.next()
input_test = input_test.to(device)
labels_test = labels_test.to(device)
output_test = model(input_test)
print(labels_test.shape)
print('outputs',output_test[0:8])
print('labels', labels_test[0:8])
print(np.sum(accuracy(output_test) == accuracy(labels_test)) / len(labels_test))
# + id="eEvblcX9StSe"
torch.save(model.state_dict(), '/content/drive/MyDrive/CSE 481 Capstone/model.pth')
| dev/train_model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# name: python2
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/aswit3/Start_Your_NLP_Career/blob/master/word2vec_pretrained_gensim.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="yxf3vOkCVYgm" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="47e7a5c2-f69f-429a-9ea1-2f1ab534ed5d"
from google.colab import drive
drive.mount('/content/drive')
# + id="nzaCH2rXVDf1" colab_type="code" colab={}
from gensim.models import KeyedVectors
# + id="iaAg1YbiVqX5" colab_type="code" colab={}
# Load vectors directly from the file
model = KeyedVectors.load_word2vec_format('drive/My Drive/Pretrained_Word_Embeddings/GoogleNews-vectors-negative300.bin', binary=True)
# + id="MWC-1DLRVqki" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="2dca5e99-8eca-4361-b521-217e3ea391d6"
# Access vectors for specific words with a keyed lookup:
vector = model['easy']
# see the shape of the vector (300,)
print(vector.shape)
# + id="EHUHCJJSVqzs" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="2e98211a-f287-479c-9ef1-46e4a6d6eff2"
# Processing sentences is not as simple as with Spacy:
vectors = [model[x] for x in "This is some text I am processing with Spacy".split(' ')]
#print(vectors)
import numpy as np
vectors = np.array(vectors)
print(vectors.shape)
# + id="b2AT4HYBWiwf" colab_type="code" colab={}
| demo/word2vec_pretrained_gensim.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # BEST NEIGHBORHOOD IN PITTSBURGH
# ## Introduction
# Using the datasets provided by the WPRDC, we chose the following three factors for the best neighborhood in Pittsburgh:
# * Crime Rate (Arrest Records)
# * Fire Incidents
# * Locations of Art Work
# Our three metrics were chosen to delegate which Pittsburgh neighborhood would provide the best quality of life. Crime rate data was included because safety is always a number one priority when looking at any neighborhood that you may move into. Nobody willingly wants to live in a neighborhood where crime rate is high because it is simply not safe. We also chose to add fire incidents data because a fire can be very detrimental to your home and neighborhood. Frequency of fires in a particular neighborhood can indicate poor craftsmanship of the homes or carelessness of neighbors. Lastly, we selected the frequency of art in Pittsburgh because we believe this can increase the visual aspect of your neighborhood. Also, sculptures and monuments tend to be in and around parks.
# ## The Metric
# We determined our metric, quality of life, through the following three items:
# * Arrest Data - This metric is of safety in each neighborhood. We measured this using data of arrests in Pittsburgh from 2016 to the present day.
# * Art Data - This metric is of the enjoyability of living in a city and is measured by the amount of public art in each neighborhood. We believe a good neighborhood should consist of a variety of public art for people to enjoy as they go about their daily lives.
# * Fire Data - This metric is safety from fires. We measured this using the amount of fires per each neighborhood in Pittsburgh. We think a good neighborhood consists of a low amount of fires.
#
# We believe that these three data seta accurately represent key components to the best quality of life in your neighborhood.
# # Arrest Data
# This metric is of safety in each neighborhood. We measured this using data of arrests in Pittsburgh from 2016 to the present day.
#
# ----
# ### Scoring of data
# We have decided to measure our data by giving a certain amount of qulaity points to the top five neighborhoods in this metric. These Quality points will be combined with other metrics to give us a standardized score to determine the overall best neighborhood in Pittsburgh.
#
# ----
# ### Data Collection
import pandas as pd
arrestdata = pd.read_csv("https://data.wprdc.org/datastore/dump/e03a89dd-134a-4ee8-a2bd-62c40aeebc6f")
arrestdata
arrestdata.groupby('INCIDENTNEIGHBORHOOD')['INCIDENTNEIGHBORHOOD'].count().sort_values(ascending=True).head(15)
# ----
# ### Quality Point Scale:
# * 1st: 5 Points
# * 2nd: 4 Points
# * 3rd: 3 Points
# * 4th: 2 Points
# * 5th: 1 Point
#
# ----
# ### Results
# * Troy Hill-Herrs Island - 5 Quality Points
# * Mt. Oliver - 4 Quality Points
# * Central Northside - 3 Quality Points
# * Regent Square - 2 Quality Points
# * Ridgemont - 1 Quality Point
# # Art Data
# This metric is of the enjoyability of living in a city and is measured by the amount of public art in each neighborhood. We believe a good neighborhood should consist of a variety of public art for people to enjoy as they go about their daily lives.
#
# ----
# ## Scoring of Data
# We have decided to measure our data by giving a certain amount of quality points to the top five neighborhoods in this metric. These Quality points will be combined with other metrics to give us a standardized score to determine the overall best neighborhood in Pittsburgh.
#
# ----
# ## Data Collection
import pandas as pd
artdata = pd.read_csv("https://data.wprdc.org/datastore/dump/00d74e83-8a23-486e-841b-286e1332a151")
artdata.groupby('neighborhood')['neighborhood'].count().sort_values(ascending=False).head(15)
# ----
# ## Quality Point Scale:
#
# * 1st: 5 Points
# * 2nd: 4 Points
# * 3rd: 3 Points
# * 4th: 2 Points
# * 5th: 1 Point
#
# ----
# ## Results
# * Central Business District - 5 Quality Points
# * Squirrel Hill South - 4 Quality Points
# * Allegheny Center - 3 Quality Points
# * Highland Park - 2 Quality Points
# * South Side Flats - 1 Quality Point
#
# ----
# # Fire Data
# This metric is safety from fires. We measured this using the amount of fires per each neighborhood in Pittsburgh. We think a good neighborhood consists of a low amount of fires.
#
# ----
# ## Scoring Data
# We have decided to measure our data by giving a certain amount of quality points to the top five neighborhoods in this metric.These quality points will be combined with other metrics to give us a standardized score to determine the overall best neighborhood in Pittsburgh.
#
# -----
# ## Data Collection
import pandas as pd
firedata = pd.read_csv("https://data.wprdc.org/datastore/dump/8d76ac6b-5ae8-4428-82a4-043130d17b02")
firedata.groupby('neighborhood')['neighborhood'].count().sort_values(ascending=True).head(15)
# ----
# ## Quality Point Scale:
# * 1st: 5 Points
# * 2nd: 4 Points
# * 3rd: 3 Points
# * 4th: 2 Points
# * 5th: 1 Point
# -----
# ## Results
# * Mount Oliver Borough - 5 Quality Points
# * Regent Square - 4 Quality Points
# * East Carnegie - 3 Quality Points
# * <NAME> - 2 Quality Points
# * St. Clair - 1 Quality Point
#
# ----
# # The Best Neighborhood
# Listed is the quality points given to each neighboorhood that was in the top five of each metric:
#
# * Troy 5
# * <NAME> 4
# * Central North 4
# * Regent 7
# * Ridgemont 2
# * Central business 5
# * Squirell Hill 4
# * Allegehny 3
# * Highland 2
# * Southside 1
# * East Carnegie 4
# * St Clair 3
# * Arlington 2
# +
import matplotlib.pyplot as plot
data = {"Neighborhood":["Troy Hill-Herrs Island", "<NAME>", "Central Northside", "Regent Square", "Ridgemont", "Central Business District", "Squirrel Hill South", "Allegheny Center", "Highland Park", "South Side Flats", "East Carnegie", "St. Clair", "Arlington Heights"], "Quality Points":[5,4,4,7,2,5,4,3,2,1,4,3,2]};
dataFrame = pd.DataFrame(data=data)
# -
dataFrame.plot.bar(x="Neighborhood", y="Quality Points", rot=70, title="Quality Points by Neighborhood");
plot.show(block=True);
# According to our data the best neighborhood in Pittsburgh is Regent Square because it scored the highest on our quality-of-life scale. As you can see by the graph Regent Square has the highest amount of quality points with 7. We believe this is the best neighborhood in Pittsburgh because of its low number of arrests and fires. According to a quote by <NAME> on Onlyinyourstate.com, “Regent Square tops the list of the safest places to live in Pittsburgh in 2016. With 4,198 residents, Regent Square tallied no violent crimes - murder, assault, or robbery. The average rent (821 a month) in Regent Square is lower than the national average of 902 and home price averages (178,809) are slightly higher than the national average of 176,700.” This website accredited Regent Square with labeling it as the safest place to live in Pittsburgh, which we believe is the most important metric.
# # Conclusion
# Personally, I have never visited but based on pictures this does seem like a great neighborhood to move into. Safety is one of my top priorities when looking at a neighborhood, so I believe that Regent Square is the best neighborhood in Pittsburgh. Also, the location of the neighborhood is close to a lot of major and popular places in Pittsburgh.
# I have never personally been to Regent Square, however after it scored the highest out of our results I looked up the neighborhood and it looks like an overall good place to live. I think while our system of ranking the neighborhoods was simple, it was effective. Our three metrics worked well to create a balanced analysis on each neighborhood. And as we expected, our system returned a neighborhood that is well balanced, very low in crime rate, and overall is ranked very high among Pittsburgh cities in many articles.
| MAIN.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# # データサイエンス100本ノック(構造化データ加工編) - R
# + [markdown] tags=[]
# ## はじめに
# - 初めに以下のセルを実行してください
# - 必要なライブラリのインポートとデータベース(PostgreSQL)からのデータ読み込みを行います
# - 利用が想定されるライブラリは以下セルでインポートしています
# - その他利用したいライブラリがあれば適宜インストールしてください(!マークに続けてOSコマンドを入力することで、任意のubuntu Linuxコマンドが入力可能)
# - 名前、住所等はダミーデータであり、実在するものではありません
# +
require('RPostgreSQL')
require('tidyr')
require('dplyr')
require('stringr')
require('caret')
require('lubridate')
require('rsample')
require('recipes')
host <- 'db'
port <- Sys.getenv()["PG_PORT"]
dbname <- Sys.getenv()["PG_DATABASE"]
user <- Sys.getenv()["PG_USER"]
password <- Sys.getenv()["PG_PASSWORD"]
con <- dbConnect(PostgreSQL(), host=host, port=port, dbname=dbname, user=user, password=password)
df_customer <- dbGetQuery(con,"SELECT * FROM customer")
df_category <- dbGetQuery(con,"SELECT * FROM category")
df_product <- dbGetQuery(con,"SELECT * FROM product")
df_receipt <- dbGetQuery(con,"SELECT * FROM receipt")
df_store <- dbGetQuery(con,"SELECT * FROM store")
df_geocode <- dbGetQuery(con,"SELECT * FROM geocode")
# -
# # 演習問題
# ---
# > R-001: レシート明細データフレーム(df_receipt)から全項目の先頭10件を表示し、どのようなデータを保有しているか目視で確認せよ。
head(df_receipt, n = 10)
# ---
# > R-002: レシート明細データフレーム(df_receipt)から売上日(sales_ymd)、顧客ID(customer_id)、商品コード(product_cd)、売上金額(amount)の順に列を指定し、10件表示させよ。
head(df_receipt[c("sales_ymd", "customer_id", "product_cd", "amount")], n=10)
# ---
# > R-003: レシート明細データフレーム(df_receipt)から売上日(sales_ymd)、顧客ID(customer_id)、商品コード(product_cd)、売上金額(amount)の順に列を指定し、10件表示させよ。ただし、sales_ymdはsales_dateに項目名を変更しながら抽出すること。
head(rename(df_receipt[c("sales_ymd", "customer_id", "product_cd", "amount")],
sales_date = sales_ymd), n=10)
# ---
# > R-004: レシート明細データフレーム(df_receipt)から売上日(sales_ymd)、顧客ID(customer_id)、商品コード(product_cd)、売上金額(amount)の順に列を指定し、以下の条件を満たすデータを抽出せよ。
# > - 顧客ID(customer_id)が"CS018205000001"
df_receipt[c("sales_ymd", "customer_id", "product_cd", "amount")] %>%
filter(customer_id == "CS018205000001")
# ---
# > R-005: レシート明細データフレーム(df_receipt)から売上日(sales_ymd)、顧客ID(customer_id)、商品コード(product_cd)、売上金額(amount)の順に列を指定し、以下の条件を満たすデータを抽出せよ。
# > - 顧客ID(customer_id)が"CS018205000001"
# > - 売上金額(amount)が1,000以上
df_receipt[c("sales_ymd", "customer_id", "product_cd", "amount")] %>%
filter(customer_id == "CS018205000001" & amount >= 1000)
# ---
# > R-006: レシート明細データフレーム(receipt)から売上日(sales_ymd)、顧客ID(customer_id)、商品コード(product_cd)、売上数量(quantity)、売上金額(amount)の順に列を指定し、以下の条件を満たすデータを抽出せよ。
# > - 顧客ID(customer_id)が"CS018205000001"
# > - 売上金額(amount)が1,000以上または売上数量(quantity)が5以上
df_receipt[c("sales_ymd", "customer_id",
"product_cd", "quantity", "amount")] %>%
filter(customer_id == "CS018205000001" &
(amount >= 1000 | quantity >= 5))
# ---
# > R-007: レシート明細のデータフレーム(df_receipt)から売上日(sales_ymd)、顧客ID(customer_id)、商品コード(product_cd)、売上金額(amount)の順に列を指定し、以下の条件を満たすデータを抽出せよ。
# > - 顧客ID(customer_id)が"CS018205000001"
# > - 売上金額(amount)が1,000以上2,000以下
df_receipt[c("sales_ymd", "customer_id", "product_cd", "amount")] %>%
filter(customer_id == "CS018205000001" &
between(amount, 1000, 2000))
# ---
# > R-008: レシート明細データフレーム(df_receipt)から売上日(sales_ymd)、顧客ID(customer_id)、商品コード(product_cd)、売上金額(amount)の順に列を指定し、以下の条件を満たすデータを抽出せよ。
# > - 顧客ID(customer_id)が"CS018205000001"
# > - 商品コード(product_cd)が"P071401019"以外
df_receipt[c("sales_ymd", "customer_id", "product_cd", "amount")] %>%
filter(customer_id == "CS018205000001" &
product_cd != "P071401019")
# ---
# > R-009: 以下の処理において、出力結果を変えずにORをANDに書き換えよ。
#
# `
# df_store %>%
# filter(!(prefecture_cd == "13" | floor_area > 900))
# `
df_store %>%
filter(prefecture_cd != "13" & floor_area <= 900)
# ---
# > R-010: 店舗データフレーム(df_store)から、店舗コード(store_cd)が"S14"で始まるものだけ全項目抽出し、10件だけ表示せよ。
head(df_store %>% filter(grepl("^S14", store_cd)), n=10)
# ---
# > R-011: 顧客データフレーム(df_customer)から顧客ID(customer_id)の末尾が1のものだけ全項目抽出し、10件だけ表示せよ。
head(df_customer %>% filter(grepl("1$", customer_id)), n=10)
# ---
# > R-012: 店舗データフレーム(df_store)から横浜市の店舗だけ全項目表示せよ。
df_store %>% filter(grepl("横浜市", address))
# ---
# > R-013: 顧客データフレーム(df_customer)から、ステータスコード(status_cd)の先頭がアルファベットのA〜Fで始まるデータを全項目抽出し、10件だけ表示せよ。
head(df_customer %>% filter(grepl("^[A-F]", status_cd)), n=10)
# ---
# > R-014: 顧客データフレーム(df_customer)から、ステータスコード(status_cd)の末尾が数字の1〜9で終わるデータを全項目抽出し、10件だけ表示せよ。
head(df_customer %>% filter(grepl("[1-9]$", status_cd)), n=10)
# ---
# > R-015: 顧客データフレーム(df_customer)から、ステータスコード(status_cd)の先頭がアルファベットのA〜Fで始まり、末尾が数字の1〜9で終わるデータを全項目抽出し、10件だけ表示せよ。
head(df_customer %>% filter(grepl("^[A-F].*[1-9]$", status_cd)), n=10)
# ---
# > R-016: 店舗データフレーム(df_store)から、電話番号(tel_no)が3桁-3桁-4桁のデータを全項目表示せよ。
df_store %>% filter(grepl("^[0-9]{3}-[0-9]{3}-[0-9]{4}$", tel_no))
# ---
# > R-017: 顧客データフレーム(df_customer)を生年月日(birth_day)で高齢順にソートし、先頭10件を全項目表示せよ。
head(
df_customer[order(df_customer$birth_day
, decreasing = FALSE), ],
n=10
)
# ---
# > R-018: 顧客データフレーム(df_customer)を生年月日(birth_day)で若い順にソートし、先頭10件を全項目表示せよ。
head(
df_customer[order(df_customer$birth_day,
decreasing = TRUE), ],
n=10
)
# ---
# > R-019: レシート明細データフレーム(df_receipt)に対し、1件あたりの売上金額(amount)が高い順にランクを付与し、先頭10件を抽出せよ。項目は顧客ID(customer_id)、売上金額(amount)、付与したランクを表示させること。なお、売上金額(amount)が等しい場合は同一順位を付与するものとする。
df_receipt %>%
mutate(ranking = min_rank(desc(amount))) %>%
arrange(ranking) %>%
slice(1:10)
# ---
# > R-020: レシート明細データフレーム(df_receipt)に対し、1件あたりの売上金額(amount)が高い順にランクを付与し、先頭10件を抽出せよ。項目は顧客ID(customer_id)、売上金額(amount)、付与したランクを表示させること。なお、売上金額(amount)が等しい場合でも別順位を付与すること。
df_receipt %>%
mutate(ranking = row_number(desc(amount))) %>%
arrange(ranking) %>%
slice(1:10)
# ---
# > R-021: レシート明細データフレーム(df_receipt)に対し、件数をカウントせよ。
nrow(df_receipt)
# ---
# > R-022: レシート明細データフレーム(df_receipt)の顧客ID(customer_id)に対し、ユニーク件数をカウントせよ。
length(unique(df_receipt$customer_id))
# ---
# > R-023: レシート明細データフレーム(df_receipt)に対し、店舗コード(store_cd)ごとに売上金額(amount)と売上数量(quantity)を合計せよ。
df_receipt %>%
group_by(store_cd) %>%
summarise(amount=sum(amount),
quantity=sum(quantity), .groups = 'drop')
# ---
# > R-024: レシート明細データフレーム(df_receipt)に対し、顧客ID(customer_id)ごとに最も新しい売上日(sales_ymd)を求め、10件表示せよ。
df_receipt %>%
group_by(customer_id) %>%
summarise(max_ymd=max(sales_ymd), .groups = 'drop') %>%
slice(1:10)
# ---
# > R-025: レシート明細データフレーム(df_receipt)に対し、顧客ID(customer_id)ごとに最も古い売上日(sales_ymd)を求め、10件表示せよ。
df_receipt %>%
group_by(customer_id) %>%
summarise(min_ymd=min(sales_ymd), .groups = 'drop') %>%
slice(1:10)
# ---
# > R-026: レシート明細データフレーム(df_receipt)に対し、顧客ID(customer_id)ごとに最も新しい売上日(sales_ymd)と古い売上日を求め、両者が異なるデータを10件表示せよ。
df_receipt %>%
group_by(customer_id) %>%
summarise(max_ymd=max(sales_ymd)
,min_ymd=min(sales_ymd), .groups = 'drop') %>%
filter(max_ymd != min_ymd) %>%
slice(1:10)
# ---
# > R-027: レシート明細データフレーム(df_receipt)に対し、店舗コード(store_cd)ごとに売上金額(amount)の平均を計算し、降順でTOP5を表示せよ。
df_receipt %>%
group_by(store_cd) %>%
summarise(mean_amount=mean(amount), .groups = 'drop') %>%
arrange(desc(mean_amount)) %>%
slice(1:5)
# ---
# > R-028: レシート明細データフレーム(df_receipt)に対し、店舗コード(store_cd)ごとに売上金額(amount)の中央値を計算し、降順でTOP5を表示せよ。
df_receipt %>%
group_by(store_cd) %>%
summarise(median_amount=median(amount), .groups = 'drop') %>%
arrange(desc(median_amount)) %>%
slice(1:5)
# ---
# > R-029: レシート明細データフレーム(df_receipt)に対し、店舗コード(store_cd)ごとに商品コード(product_cd)の最頻値を求めよ。
# コード例1
df_receipt %>%
group_by(store_cd, product_cd) %>%
summarise(n = n(), .groups = 'drop_last') %>%
filter(n == n %>% max()) %>%
ungroup()
# +
# コード例2:whtch.max()を使ってこんな感じも(最頻値複数発生時、どれか一つに絞られる)
table_product <-table(c(df_receipt$store_cd), df_receipt$product_cd)
store <- names(table_product[,1])
mode_product <- c()
for (i in 1:length(store)){
mode_product[i] <- names(which.max(table_product[i,]))
}
data.frame(store_cd = store, store_cd = mode_product)
# -
# ---
# > R-030: レシート明細データフレーム(df_receipt)に対し、店舗コード(store_cd)ごとに売上金額(amount)の標本分散を計算し、降順でTOP5を表示せよ。
# +
var_sample <- function(x){ var(x)*(length(x)-1)/length(x) }
df_receipt %>%
group_by(store_cd) %>%
summarise(var_amount=var_sample(amount), .groups = 'drop') %>%
arrange(desc(var_amount)) %>%
slice(1:5)
# -
# ---
# > R-031: レシート明細データフレーム(df_receipt)に対し、店舗コード(store_cd)ごとに売上金額(amount)の標本標準偏差を計算し、降順でTOP5を表示せよ。
# +
var_sample <- function(x){ var(x)*(length(x)-1)/length(x) }
std_sample <- function(x){ sqrt(var_sample(x)) }
head(
df_receipt %>%
group_by(store_cd) %>%
summarise(std_amount=std_sample(amount), .groups = 'drop') %>%
arrange(desc(std_amount))
, n=5
)
# -
# ---
# > R-032: レシート明細データフレーム(df_receipt)の売上金額(amount)について、25%刻みでパーセンタイル値を求めよ。
df_receipt %>%
summarise(amount_25per=quantile(amount, 0.25),
amount_50per=quantile(amount, 0.5),
amount_75per=quantile(amount, 0.75),
amount_100per=quantile(amount, 1.0))
# ---
# > R-033: レシート明細データフレーム(df_receipt)に対し、店舗コード(store_cd)ごとに売上金額(amount)の平均を計算し、330以上のものを抽出せよ。
df_receipt %>%
group_by(store_cd) %>%
summarise(mean_amount=mean(amount), .groups = 'drop') %>%
filter(mean_amount >= 330)
# ---
# > R-034: レシート明細データフレーム(df_receipt)に対し、顧客ID(customer_id)ごとに売上金額(amount)を合計して全顧客の平均を求めよ。ただし、顧客IDが"Z"から始まるのものは非会員を表すため、除外して計算すること。
df_mean <- df_receipt %>%
filter(!grepl("^Z", customer_id)) %>%
group_by(customer_id) %>%
summarise(sum_amount=sum(amount), .groups = 'drop') %>%
summarise(mean_amount=mean(sum_amount))
df_mean$mean_amount
# ---
# > R-035: レシート明細データフレーム(df_receipt)に対し、顧客ID(customer_id)ごとに売上金額(amount)を合計して全顧客の平均を求め、平均以上に買い物をしている顧客を抽出せよ。ただし、顧客IDが"Z"から始まるのものは非会員を表すため、除外して計算すること。なお、データは10件だけ表示させれば良い。
df_sum <- df_receipt %>%
filter(!grepl("^Z", customer_id)) %>%
group_by(customer_id) %>%
summarise(sum_amount=sum(amount), .groups = 'drop')
df_mean <- df_sum %>%
summarise(mean_amount=mean(sum_amount))
df_sum %>%
filter(sum_amount > df_mean$mean_amount) %>%
slice(1:10)
# ---
# > R-036: レシート明細データフレーム(df_receipt)と店舗データフレーム(df_store)を内部結合し、レシート明細データフレームの全項目と店舗データフレームの店舗名(store_name)を10件表示させよ。
inner_join(df_receipt,
df_store[c('store_cd', 'store_name')], by = 'store_cd') %>%
slice(1:10)
# ---
# > R-037: 商品データフレーム(df_product)とカテゴリデータフレーム(df_category)を内部結合し、商品データフレームの全項目とカテゴリデータフレームの小区分名(category_small_name)を10件表示させよ。
inner_join(df_product,
df_category[c('category_small_cd','category_small_name')],
by = 'category_small_cd') %>%
slice(1:10)
# ---
# > R-038: 顧客データフレーム(df_customer)とレシート明細データフレーム(df_receipt)から、各顧客ごとの売上金額合計を求めよ。ただし、買い物の実績がない顧客については売上金額を0として表示させること。また、顧客は性別コード(gender_cd)が女性(1)であるものを対象とし、非会員(顧客IDが'Z'から始まるもの)は除外すること。なお、結果は10件だけ表示させれば良い。
# +
df_sum <- df_receipt %>%
filter(!grepl("^Z", customer_id)) %>%
group_by(customer_id) %>%
summarise(sum_amount=sum(amount), .groups = 'drop')
df_target <- df_customer %>%
filter(!grepl("^Z", customer_id) & gender_cd == '1')
left_join(df_target['customer_id'], df_sum, by = 'customer_id') %>%
replace_na(list(sum_amount=0)) %>%
slice(1:10)
# -
# ---
# > R-039: レシート明細データフレーム(df_receipt)から売上日数の多い顧客の上位20件と、売上金額合計の多い顧客の上位20件を抽出し、完全外部結合せよ。ただし、非会員(顧客IDが'Z'から始まるもの)は除外すること。
# +
# コード例1
df_sum <- df_receipt %>%
filter(!grepl("^Z", customer_id)) %>%
group_by(customer_id) %>%
summarise(sum_amount=sum(amount), .groups = 'drop') %>%
arrange(desc(sum_amount)) %>%
slice(1:20)
df_cnt <- df_receipt %>%
filter(!grepl("^Z", customer_id)) %>%
group_by(customer_id) %>%
summarise(come_days=n_distinct(sales_ymd), .groups = 'drop') %>%
arrange(desc(come_days), customer_id) %>%
slice(1:20)
full_join(df_sum, df_cnt, by = "customer_id")
# +
# コード例2
# slice_maxを使用したコード例
# with_ties = FALSEをwith_ties = TRUEに書き換えると同一順位を含める形に変更できる
df_sum <- df_receipt %>%
filter(!grepl("^Z", customer_id)) %>%
group_by(customer_id) %>%
summarise(sum_amount = sum(amount)) %>%
slice_max(sum_amount, n = 20, with_ties = FALSE)
df_cnt <- df_receipt %>%
filter(!grepl("^Z", customer_id)) %>%
group_by(customer_id) %>%
summarise(come_days = n_distinct(sales_ymd)) %>%
slice_max(come_days, n = 20, with_ties = FALSE)
df_original <- full_join(df_sum, df_cnt, by = "customer_id")
df_original
# -
# ---
# > R-040: 全ての店舗と全ての商品を組み合わせると何件のデータとなるか調査したい。店舗(df_store)と商品(df_product)を直積した件数を計算せよ。
df_store_tmp <- df_store
df_product_tmp <- df_product
df_store_tmp['key'] <- 0
df_product_tmp['key'] <- 0
nrow(full_join(df_store_tmp, df_product_tmp, by = "key"))
# ---
# > R-041: レシート明細データフレーム(df_receipt)の売上金額(amount)を日付(sales_ymd)ごとに集計し、前日からの売上金額増減を計算せよ。なお、計算結果は10件表示すればよい。
df_receipt %>%
group_by(sales_ymd) %>%
summarise(sum_amount=sum(amount), .groups = 'drop') %>%
mutate(lag_ymd = lag(sales_ymd),
lag_amount = lag(sum_amount),
diff_amount = sum_amount - lag_amount) %>%
slice(1:10)
# ---
# > R-042: レシート明細データフレーム(df_receipt)の売上金額(amount)を日付(sales_ymd)ごとに集計し、各日付のデータに対し、1日前、2日前、3日前のデータを結合せよ。結果は10件表示すればよい。
# +
# コード例1:縦持ちケース
df_sum <-df_receipt %>%
group_by(sales_ymd) %>%
summarise(sum_amount=sum(amount), .groups = 'drop')
for (i in 1:3){
if (i == 1){
df_lag <- df_sum %>%
mutate(lag_ymd = lag(sales_ymd, n=i),
lag_amount = lag(sum_amount, n=i))
}
else{
df_tmp <- df_sum %>%
mutate(lag_ymd = lag(sales_ymd, n=i),
lag_amount = lag(sum_amount, n=i))
df_lag <- rbind(df_lag, df_tmp)
}
}
drop_na(df_lag, everything()) %>%
arrange(sales_ymd, lag_ymd) %>%
slice(1:10)
# +
# コード例2:横持ちケース
df_sum <-df_receipt %>%
group_by(sales_ymd) %>%
summarise(sum_amount=sum(amount), .groups = 'drop')
for (i in 1:3){
if (i == 1){
df_lag <- df_sum %>%
mutate(lag_ymd_1 = lag(sales_ymd, n=i),
lag_amount_1 = lag(sum_amount, n=i))
}
else{
col_name_1 <- paste("lag_ymd", i , sep="_")
col_name_2 <- paste("lag_amount", i , sep="_")
print(col_name_1)
print(col_name_2)
df_tmp <- df_sum %>%
mutate(!!col_name_1 := lag(sales_ymd, n=i),
!!col_name_2 := lag(sum_amount, n=i))
df_lag <- cbind(df_lag, df_tmp[c(col_name_1, col_name_2)])
}
}
drop_na(df_lag, everything()) %>%
arrange(sales_ymd) %>%
slice(1:10)
# -
# ---
# > R-043: レシート明細データフレーム(df_receipt)と顧客データフレーム(df_customer)を結合し、性別(gender)と年代(ageから計算)ごとに売上金額(amount)を合計した売上サマリデータフレーム(df_sales_summary)を作成せよ。性別は0が男性、1が女性、9が不明を表すものとする。
# >
# > ただし、項目構成は年代、女性の売上金額、男性の売上金額、性別不明の売上金額の4項目とすること(縦に年代、横に性別のクロス集計)。また、年代は10歳ごとの階級とすること。
# +
df_sales_summary <- df_customer[c("customer_id",
"gender_cd", "birth_day" , "age")] %>%
mutate(era = trunc(age / 10) * 10) %>%
inner_join(df_receipt, by="customer_id") %>%
group_by(gender_cd, era) %>%
summarise(sum_amount=sum(amount), .groups = 'drop') %>%
spread(gender_cd, sum_amount, fill=0) %>%
rename(male='0', female='1', unknown='9')
df_sales_summary
# -
# ---
# > R-044: 前設問で作成した売上サマリデータフレーム(df_sales_summary)は性別の売上を横持ちさせたものであった。このデータフレームから性別を縦持ちさせ、年代、性別コード、売上金額の3項目に変換せよ。ただし、性別コードは男性を'00'、女性を'01'、不明を'99'とする。
gather(df_sales_summary, key = gender_cd,
value = sum_amount, male, female, unknown) %>%
mutate(gender_cd = case_when(
gender_cd == "male" ~ "00",
gender_cd == "female" ~ "01",
gender_cd == "unknown" ~ "99",
))
# ---
# > R-045: 顧客データフレーム(df_customer)の生年月日(birth_day)は日付型でデータを保有している。これをYYYYMMDD形式の文字列に変換し、顧客ID(customer_id)とともに抽出せよ。データは10件を抽出すれば良い。
# +
df_tmp <- cbind(df_customer['customer_id'],
strftime(df_customer$birth_day, format="%Y%m%d"))
colnames(df_tmp) <- c("customer_id","birth_day")
head(df_tmp,10)
# -
# ---
# > R-046: 顧客データフレーム(df_customer)の申し込み日(application_date)はYYYYMMDD形式の文字列型でデータを保有している。これを日付型に変換し、顧客ID(customer_id)とともに抽出せよ。データは10件を抽出すれば良い。
df_tmp <- cbind(df_customer["customer_id"],
strptime(df_customer$application_date, '%Y%m%d'))
colnames(df_tmp) <- c("customer_id","application_date")
head(df_tmp, 10)
# ---
# > R-047: レシート明細データフレーム(df_receipt)の売上日(sales_ymd)はYYYYMMDD形式の数値型でデータを保有している。これを日付型に変換し、レシート番号(receipt_no)、レシートサブ番号(receipt_sub_no)とともに抽出せよ。データは10件を抽出すれば良い。
df_tmp <- cbind(df_receipt[c('receipt_no', 'receipt_sub_no')],
strptime(as.character(df_receipt$sales_ymd), '%Y%m%d'))
colnames(df_tmp) <- c("receipt_no","receipt_sub_no", "sales_ymd")
head(df_tmp, 10)
# ---
# > R-048: レシート明細データフレーム(df_receipt)の売上エポック秒(sales_epoch)は数値型のUNIX秒でデータを保有している。これを日付型に変換し、レシート番号(receipt_no)、レシートサブ番号(receipt_sub_no)とともに抽出せよ。データは10件を抽出すれば良い。
df_tmp <- cbind(df_receipt[c('receipt_no', 'receipt_sub_no')],
as.POSIXct(df_receipt$sales_epoch, origin="1970-01-01"))
colnames(df_tmp) <- c("receipt_no","receipt_sub_no", "sales_ymd")
head(df_tmp, 10)
# ---
# > R-049: レシート明細データフレーム(df_receipt)の売上エポック秒(sales_epoch)を日付型に変換し、「年」だけ取り出してレシート番号(receipt_no)、レシートサブ番号(receipt_sub_no)とともに抽出せよ。データは10件を抽出すれば良い。
df_tmp <- cbind(df_receipt[c('receipt_no', 'receipt_sub_no')],
substring(as.POSIXct(df_receipt$sales_epoch,
origin="1970-01-01"), 1, 4))
colnames(df_tmp) <- c("receipt_no","receipt_sub_no", "sales_ymd")
head(df_tmp, 10)
# ---
# > R-050: レシート明細データフレーム(df_receipt)の売上エポック秒(sales_epoch)を日付型に変換し、「月」だけ取り出してレシート番号(receipt_no)、レシートサブ番号(receipt_sub_no)とともに抽出せよ。なお、「月」は0埋め2桁で取り出すこと。データは10件を抽出すれば良い。
df_tmp <- cbind(df_receipt[c('receipt_no', 'receipt_sub_no')],
substring(as.POSIXct(df_receipt$sales_epoch,
origin="1970-01-01"), 6, 7))
colnames(df_tmp) <- c("receipt_no","receipt_sub_no", "sales_ymd")
head(df_tmp, 10)
# ---
# > R-051: レシート明細データフレーム(df_receipt)の売上エポック秒を日付型に変換し、「日」だけ取り出してレシート番号(receipt_no)、レシートサブ番号(receipt_sub_no)とともに抽出せよ。なお、「日」は0埋め2桁で取り出すこと。データは10件を抽出すれば良い。
df_tmp <- cbind(df_receipt[c('receipt_no', 'receipt_sub_no')],
substring(as.POSIXct(df_receipt$sales_epoch,
origin="1970-01-01"), 9, 10))
colnames(df_tmp) <- c("receipt_no","receipt_sub_no", "sales_ymd")
head(df_tmp, 10)
# ---
# > R-052: レシート明細データフレーム(df_receipt)の売上金額(amount)を顧客ID(customer_id)ごとに合計の上、売上金額合計に対して2,000円以下を0、2,000円より大きい金額を1に2値化し、顧客ID、売上金額合計とともに10件表示せよ。ただし、顧客IDが"Z"から始まるのものは非会員を表すため、除外して計算すること。
df_receipt %>%
filter(!grepl("^Z", customer_id)) %>%
group_by(customer_id) %>%
summarise(sum_amount=sum(amount), .groups = 'drop') %>%
mutate(bit_amount = ifelse(sum_amount <= 2000, 0, 1)) %>%
slice(1:10)
# ---
# > R-053: 顧客データフレーム(df_customer)の郵便番号(postal_cd)に対し、東京(先頭3桁が100〜209のもの)を1、それ以外のものを0に2値化せよ。さらにレシート明細データフレーム(df_receipt)と結合し、全期間において買い物実績のある顧客数を、作成した2値ごとにカウントせよ。
inner_join(df_receipt, df_customer, by = "customer_id") %>%
mutate(postal_bit =
ifelse(100 <= as.integer(str_sub(postal_cd, start=1, end=3))
& as.integer(str_sub(postal_cd, start=1, end=3)) <= 209,
1, 0)) %>%
group_by(postal_bit) %>%
summarise(customer_cnt = n_distinct(customer_id), .groups = 'drop')
# ---
# > R-054: 顧客データフレーム(df_customer)の住所(address)は、埼玉県、千葉県、東京都、神奈川県のいずれかとなっている。都道府県毎にコード値を作成し、顧客ID、住所とともに抽出せよ。値は埼玉県を11、千葉県を12、東京都を13、神奈川県を14とすること。結果は10件表示させれば良い。
df_customer %>%
mutate(prefecture_cd =
case_when(
str_sub(address, start=1, end=3) == "埼玉県" ~ "11",
str_sub(address, start=1, end=3) == "千葉県" ~ "12",
str_sub(address, start=1, end=3) == "東京都" ~ "13",
str_sub(address, start=1, end=3) == "神奈川" ~ "14")) %>%
select(customer_id, address, prefecture_cd) %>%
slice(1:10)
# ---
# > R-055: レシート明細データフレーム(df_receipt)の売上金額(amount)を顧客ID(customer_id)ごとに合計し、その合計金額の四分位点を求めよ。その上で、顧客ごとの売上金額合計に対して以下の基準でカテゴリ値を作成し、顧客ID、売上金額合計とともに表示せよ。カテゴリ値は上から順に1〜4とする。結果は10件表示させれば良い。
# >
# > - 最小値以上第一四分位未満
# > - 第一四分位以上第二四分位未満
# > - 第二四分位以上第三四分位未満
# > - 第三四分位以上
df_receipt %>%
group_by(customer_id) %>%
summarise(sum_amount = sum(amount), .groups = 'drop') %>%
mutate(amount_cat = case_when(
sum_amount < quantile(sum_amount)[2] ~ "1",
sum_amount < quantile(sum_amount)[3] ~ "2",
sum_amount < quantile(sum_amount)[4] ~ "3",
quantile(sum_amount)[4] <= sum_amount ~ "4"
)) %>%
slice(1:10)
# ---
# > R-056: 顧客データフレーム(df_customer)の年齢(age)をもとに10歳刻みで年代を算出し、顧客ID(customer_id)、生年月日(birth_day)とともに抽出せよ。ただし、60歳以上は全て60歳代とすること。年代を表すカテゴリ名は任意とする。先頭10件を表示させればよい。
df_customer[c("customer_id", "birth_day","age")] %>%
mutate(era = trunc(age / 10) * 10) %>%
mutate(era = case_when(
era < 60 ~ as.factor(era),
era >= 60 ~ as.factor("60")
)) %>%
slice(1:10)
# ---
# > R-057: 前問題の抽出結果と性別(gender)を組み合わせ、新たに性別×年代の組み合わせを表すカテゴリデータを作成せよ。組み合わせを表すカテゴリの値は任意とする。先頭10件を表示させればよい。
# 性別コード1桁と年代コード2桁を連結した性年代コードを生成する
df_customer[c("customer_id", "gender_cd","birth_day", "age")] %>%
mutate(era = trunc(age / 10) * 10) %>%
mutate(era = case_when(
era < 60 ~ as.factor(era),
era >= 60 ~ as.factor("60"))) %>%
mutate(gender_era_cd = paste(gender_cd, era, sep="")) %>%
slice(1:10)
# ---
# > R-058: 顧客データフレーム(df_customer)の性別コード(gender_cd)をダミー変数化し、顧客ID(customer_id)とともに抽出せよ。結果は10件表示させれば良い。
dummy_gender_model <- dummyVars(~gender_cd,
data = df_customer, fullRank = FALSE)
dummy_dender <- predict(dummy_gender_model, df_customer)
head(cbind(df_customer['customer_id'], dummy_dender),10)
# ---
# > R-059: レシート明細データフレーム(df_receipt)の売上金額(amount)を顧客ID(customer_id)ごとに合計し、売上金額合計を平均0、標準偏差1に標準化して顧客ID、売上金額合計とともに表示せよ。標準化に使用する標準偏差は、不偏標準偏差と標本標準偏差のどちらでも良いものとする。ただし、顧客IDが"Z"から始まるのものは非会員を表すため、除外して計算すること。結果は10件表示させれば良い。
# Rのscaleを使用しているため不偏標準偏差で標準化される
df_receipt %>%
filter(!grepl("^Z", customer_id)) %>%
group_by(customer_id) %>%
summarise(sum_amount = sum(amount), .groups = 'drop') %>%
mutate(normalized_amount = scale(sum_amount,
center = TRUE, scale = TRUE)) %>%
slice(1:10)
# ---
# > R-060: レシート明細データフレーム(df_receipt)の売上金額(amount)を顧客ID(customer_id)ごとに合計し、売上金額合計を最小値0、最大値1に正規化して顧客ID、売上金額合計とともに表示せよ。ただし、顧客IDが"Z"から始まるのものは非会員を表すため、除外して計算すること。結果は10件表示させれば良い。
df_receipt %>%
filter(!grepl("^Z", customer_id)) %>%
group_by(customer_id) %>%
summarise(sum_amount = sum(amount), .groups = 'drop') %>%
mutate(
normalized_amount = scale(sum_amount,
center = min(sum_amount),
scale = max(sum_amount) - min(sum_amount)))%>%
slice(1:10)
# ---
# > R-061: レシート明細データフレーム(df_receipt)の売上金額(amount)を顧客ID(customer_id)ごとに合計し、売上金額合計を常用対数化(底=10)して顧客ID、売上金額合計とともに表示せよ(ただし、顧客IDが"Z"から始まるのものは非会員を表すため、除外して計算すること)。結果は10件表示させれば良い。
df_receipt %>%
filter(!grepl("^Z", customer_id)) %>%
group_by(customer_id) %>%
summarise(sum_amount = sum(amount), .groups = 'drop') %>%
mutate(log_amount = log((sum_amount + 1),10)) %>%
slice(1:10)
# ---
# > R-062: レシート明細データフレーム(df_receipt)の売上金額(amount)を顧客ID(customer_id)ごとに合計し、売上金額合計を自然対数化(底=e)して顧客ID、売上金額合計とともに表示せよ。ただし、顧客IDが"Z"から始まるのものは非会員を表すため、除外して計算すること。結果は10件表示させれば良い。
df_receipt %>%
filter(!grepl("^Z", customer_id)) %>%
group_by(customer_id) %>%
summarise(sum_amount = sum(amount), .groups = 'drop') %>%
mutate(log_amount = log(sum_amount + 1)) %>%
slice(1:10)
# ---
# > R-063: 商品データフレーム(df_product)の単価(unit_price)と原価(unit_cost)から、各商品の利益額を算出せよ。結果は10件表示させれば良い。
df_product %>%
mutate(unit_profit = unit_price - unit_cost) %>%
slice(1:10)
# ---
# > R-064: 商品データフレーム(df_product)の単価(unit_price)と原価(unit_cost)から、各商品の利益率の全体平均を算出せよ。 ただし、単価と原価にはNULLが存在することに注意せよ。
df_product %>%
mutate(unit_profit_rate = (unit_price - unit_cost)/ unit_price)%>%
summarise(total_mean = mean(unit_profit_rate, na.rm = TRUE))
# ---
# > R-065: 商品データフレーム(df_product)の各商品について、利益率が30%となる新たな単価を求めよ。ただし、1円未満は切り捨てること。そして結果を10件表示させ、利益率がおよそ30%付近であることを確認せよ。ただし、単価(unit_price)と原価(unit_cost)にはNULLが存在することに注意せよ。
df_product %>%
mutate(new_price = trunc(unit_cost / 0.7)) %>%
mutate(new_profit_rate = (new_price - unit_cost)/ new_price) %>%
slice(1:10)
# ---
# > R-066: 商品データフレーム(df_product)の各商品について、利益率が30%となる新たな単価を求めよ。今回は、1円未満を四捨五入すること(0.5については偶数方向の丸めで良い)。そして結果を10件表示させ、利益率がおよそ30%付近であることを確認せよ。ただし、単価(unit_price)と原価(unit_cost)にはNULLが存在することに注意せよ。
df_product %>%
mutate(new_price = round(unit_cost / 0.7)) %>%
mutate(new_profit_rate = (new_price - unit_cost)/ new_price) %>%
slice(1:10)
# ---
# > R-067: 商品データフレーム(df_product)の各商品について、利益率が30%となる新たな単価を求めよ。今回は、1円未満を切り上げること。そして結果を10件表示させ、利益率がおよそ30%付近であることを確認せよ。ただし、単価(unit_price)と原価(unit_cost)にはNULLが存在することに注意せよ。
df_product %>%
mutate(new_price = ceiling(unit_cost / 0.7)) %>%
mutate(new_profit_rate = (new_price - unit_cost)/ new_price) %>%
slice(1:10)
# ---
# > R-068: 商品データフレーム(df_product)の各商品について、消費税率10%の税込み金額を求めよ。1円未満の端数は切り捨てとし、結果は10件表示すれば良い。ただし、単価(unit_price)にはNULLが存在することに注意せよ。
head(cbind(df_product['product_cd'],trunc(df_product['unit_price'] * 1.1)),10)
# ---
# > R-069: レシート明細データフレーム(df_receipt)と商品データフレーム(df_product)を結合し、顧客毎に全商品の売上金額合計と、カテゴリ大区分(category_major_cd)が"07"(瓶詰缶詰)の売上金額合計を計算の上、両者の比率を求めよ。抽出対象はカテゴリ大区分"07"(瓶詰缶詰)の購入実績がある顧客のみとし、結果は10件表示させればよい。
inner_join(df_receipt,
df_product[c("product_cd","category_major_cd")], by="product_cd") %>%
filter(category_major_cd == "07") %>%
group_by(customer_id) %>%
summarise(amount_07 = sum(amount), .groups = 'drop') %>%
inner_join(df_receipt, by="customer_id") %>%
group_by(customer_id) %>%
summarise(sum_amount=sum(amount),
amount_07=max(amount_07), .groups = 'drop') %>%
mutate(rate_07 = amount_07 / sum_amount) %>%
slice(1:10)
# ---
# > R-070: レシート明細データフレーム(df_receipt)の売上日(sales_ymd)に対し、顧客データフレーム(df_customer)の会員申込日(application_date)からの経過日数を計算し、顧客ID(customer_id)、売上日、会員申込日とともに表示せよ。結果は10件表示させれば良い(なお、sales_ymdは数値、application_dateは文字列でデータを保持している点に注意)。
inner_join(df_receipt[c("customer_id", "sales_ymd")],
df_customer[c("customer_id","application_date")],
by="customer_id") %>%
distinct(.,.keep_all=TRUE) %>%
mutate(elapsed_days = strptime(as.character(sales_ymd), '%Y%m%d')
- strptime(application_date, '%Y%m%d')) %>%
select(customer_id, sales_ymd, application_date, elapsed_days) %>%
slice(1:10)
# ---
# > R-071: レシート明細データフレーム(df_receipt)の売上日(sales_ymd)に対し、顧客データフレーム(df_customer)の会員申込日(application_date)からの経過月数を計算し、顧客ID(customer_id)、売上日、会員申込日とともに表示せよ。結果は10件表示させれば良い(なお、sales_ymdは数値、application_dateは文字列でデータを保持している点に注意)。1ヶ月未満は切り捨てること。
(time_length(strptime('20171122', '%Y%m%d')
- strptime('20150623', '%Y%m%d'), "month"))
time_length(interval(strptime('20150623', '%Y%m%d'),
strptime('20171122', '%Y%m%d')), "month")
inner_join(df_receipt[c("customer_id", "sales_ymd")],
df_customer[c("customer_id","application_date")],
by="customer_id") %>%
distinct(.,.keep_all=TRUE) %>%
mutate(elapsed_months =
trunc(time_length(
interval(
strptime(application_date, '%Y%m%d'),
strptime(as.character(sales_ymd), '%Y%m%d')
), "month"))) %>%
select(customer_id, sales_ymd, application_date, elapsed_months) %>%
slice(1:10)
# ---
# > R-072: レシート明細データフレーム(df_receipt)の売上日(sales_ymd)に対し、顧客データフレーム(df_customer)の会員申込日(application_date)からの経過年数を計算し、顧客ID(customer_id)、売上日、会員申込日とともに表示せよ。結果は10件表示させれば良い(なお、sales_ymdは数値、application_dateは文字列でデータを保持している点に注意)。1年未満は切り捨てること。
inner_join(df_receipt[c("customer_id", "sales_ymd")],
df_customer[c("customer_id","application_date")],
by="customer_id") %>%
distinct(.,.keep_all=TRUE) %>%
mutate(elapsed_years =
trunc(time_length(interval(
strptime(application_date, '%Y%m%d'),
strptime(as.character(sales_ymd), '%Y%m%d')), "year")))%>%
select(customer_id, sales_ymd, application_date, elapsed_years) %>%
slice(1:10)
# ---
# > R-073: レシート明細データフレーム(df_receipt)の売上日(sales_ymd)に対し、顧客データフレーム(df_customer)の会員申込日(application_date)からのエポック秒による経過時間を計算し、顧客ID(customer_id)、売上日、会員申込日とともに表示せよ。結果は10件表示させれば良い(なお、sales_ymdは数値、application_dateは文字列でデータを保持している点に注意)。なお、時間情報は保有していないため各日付は0時0分0秒を表すものとする。
inner_join(df_receipt[c("customer_id", "sales_ymd")],
df_customer[c("customer_id","application_date")],
by="customer_id") %>%
distinct(.,.keep_all=TRUE) %>%
mutate(elapsed_epoch =
as.numeric(strptime(as.character(sales_ymd), '%Y%m%d'))
- as.numeric(strptime(application_date, '%Y%m%d'))) %>%
select(customer_id, sales_ymd, application_date, elapsed_epoch) %>%
slice(1:10)
# ---
# > R-074: レシート明細データフレーム(df_receipt)の売上日(sales_ymd)に対し、当該週の月曜日からの経過日数を計算し、顧客ID、売上日、当該週の月曜日付とともに表示せよ。結果は10件表示させれば良い(なお、sales_ymdは数値でデータを保持している点に注意)。
df_receipt[c("customer_id", "sales_ymd")] %>%
distinct(.,.keep_all=TRUE) %>%
# 以下では開始日が日曜日となるため1日前シフトさせることで日曜日を前週の最終日に変換している。
# その状態でfloor_dateをすると、週の開始日として前週の日曜日が取得できるため、開始日を月曜日にするために1を足している。
mutate(monday = as.Date(floor_date(
strptime(as.character(sales_ymd), '%Y%m%d') - 1 , unit="week")) + 1) %>%
mutate(elapse_weekday = as.Date(
strptime(as.character(sales_ymd), '%Y%m%d')) - monday) %>%
select(customer_id, sales_ymd, monday, elapse_weekday) %>%
slice(1:10)
# ---
# > R-075: 顧客データフレーム(df_customer)からランダムに1%のデータを抽出し、先頭から10件データを抽出せよ。
head(sample_frac(tbl = df_customer, 0.01),10)
# ---
# > R-076: 顧客データフレーム(df_customer)から性別(gender_cd)の割合に基づきランダムに10%のデータを層化抽出し、性別ごとに件数を集計せよ。
set.seed(71)
df_customer %>%
group_by(gender_cd) %>%
sample_frac(0.1) %>%
summarise(customer_num = n(), .groups = 'drop')
# ---
# > R-077: レシート明細データフレーム(df_receipt)の売上金額(amount)を顧客単位に合計し、合計した売上金額の外れ値を抽出せよ。ただし、顧客IDが"Z"から始まるのものは非会員を表すため、除外して計算すること。なお、ここでは外れ値を平均から3σ以上離れたものとする。結果は10件表示させれば良い。
df_receipt %>%
filter(!grepl("^Z", customer_id)) %>%
group_by(customer_id) %>%
summarise(sum_amount = sum(amount), .groups = 'drop') %>%
filter(abs(sum_amount - mean(sum_amount)) / sd(sum_amount) >= 3) %>%
slice(1:10)
# ---
# > R-078: レシート明細データフレーム(df_receipt)の売上金額(amount)を顧客単位に合計し、合計した売上金額の外れ値を抽出せよ。ただし、顧客IDが"Z"から始まるのものは非会員を表すため、除外して計算すること。なお、ここでは外れ値を第一四分位と第三四分位の差であるIQRを用いて、「第一四分位数-1.5×IQR」よりも下回るもの、または「第三四分位数+1.5×IQR」を超えるものとする。結果は10件表示させれば良い。
# +
# 確認用コード
df_tmp <- df_receipt %>%
filter(!grepl("^Z", customer_id)) %>%
group_by(customer_id) %>%
summarise(sum_amount = sum(amount), .groups = 'drop')
quantile(df_tmp$sum_amount)
# -
# 確認用コード
(3602-554) * 1.5 + 3602
df_receipt %>%
group_by(customer_id) %>%
filter(!grepl("^Z", customer_id)) %>%
summarise(sum_amount = sum(amount), .groups = 'drop') %>%
filter(
sum_amount < quantile(sum_amount)[2]
- 1.5 * (quantile(sum_amount)[4] - quantile(sum_amount)[2])
|
sum_amount > quantile(sum_amount)[4]
+ 1.5 * (quantile(sum_amount)[4] - quantile(sum_amount)[2])
) %>%
slice(1:10)
# ---
# > R-079: 商品データフレーム(df_product)の各項目に対し、欠損数を確認せよ。
sapply(df_product, function(x) sum(is.na(x)))
# ---
# > R-080: 商品データフレーム(df_product)のいずれかの項目に欠損が発生しているレコードを全て削除した新たなdf_product_1を作成せよ。なお、削除前後の件数を表示させ、前設問で確認した件数だけ減少していることも確認すること。
nrow(df_product)
df_product_1 <- na.omit(df_product)
nrow(df_product_1)
# ---
# > R-081: 単価(unit_price)と原価(unit_cost)の欠損値について、それぞれの平均値で補完した新たなdf_product_2を作成せよ。なお、平均値について1円未満は四捨五入とし、0.5については偶数寄せでかまわない。補完実施後、各項目について欠損が生じていないことも確認すること。
# +
price_mean <- round(mean(df_product$unit_price, na.rm=TRUE))
cost_mean <- round(mean(df_product$unit_cost, na.rm=TRUE))
df_product_2 <- df_product %>%
replace_na(list(unit_price = price_mean, unit_cost = cost_mean))
sapply(df_product_2, function(x) sum(is.na(x)))
# -
# ---
# > R-082: 単価(unit_price)と原価(unit_cost)の欠損値について、それぞれの中央値で補完した新たなdf_product_3を作成せよ。なお、中央値について1円未満は四捨五入とし、0.5については偶数寄せでかまわない。補完実施後、各項目について欠損が生じていないことも確認すること。
# +
price_median <- round(median(df_product$unit_price, na.rm=TRUE))
cost_median <- round(median(df_product$unit_cost, na.rm=TRUE))
df_product_3 <- df_product %>%
replace_na(list(unit_price = price_median, unit_cost = cost_median))
sapply(df_product_3, function(x) sum(is.na(x)))
# -
# ---
# > R-083: 単価(unit_price)と原価(unit_cost)の欠損値について、各商品の小区分(category_small_cd)ごとに算出した中央値で補完した新たなdf_product_4を作成せよ。なお、中央値について1円未満は四捨五入とし、0.5については偶数寄せでかまわない。補完実施後、各項目について欠損が生じていないことも確認すること。
# +
df_product_4 <- df_product %>%
group_by(category_small_cd) %>%
summarise(price_median = round(median(unit_price, na.rm=TRUE)),
cost_median=round(median(unit_cost, na.rm=TRUE)),
.groups = 'drop') %>%
inner_join(df_product, by="category_small_cd") %>%
mutate(unit_price = ifelse(is.na(unit_price), price_median, unit_price),
unit_cost = ifelse(is.na(unit_cost), cost_median, unit_cost))
sapply(df_product_4, function(x) sum(is.na(x)))
# -
df_product_4 %>%
filter(product_cd == 'P050103021')
# ---
# > R-084: 顧客データフレーム(df_customer)の全顧客に対し、全期間の売上金額に占める2019年売上金額の割合を計算せよ。ただし、販売実績のない場合は0として扱うこと。そして計算した割合が0超のものを抽出せよ。 結果は10件表示させれば良い。また、作成したデータにNAやNANが存在しないことを確認せよ。
# +
df_receipt_2019 <- df_receipt %>%
filter(20190101 <= sales_ymd & sales_ymd <= 20191231)
df_amount_all <-left_join(df_customer, df_receipt, by="customer_id") %>%
group_by(customer_id) %>%
summarise(amount_all = sum(amount), .groups = 'drop')
df_amount_2019 <-left_join(df_customer, df_receipt_2019, by="customer_id") %>%
group_by(customer_id) %>%
summarise(amount_2019 = sum(amount), .groups = 'drop') %>%
inner_join(df_amount_all, by="customer_id") %>%
replace_na(list(amount_2019 = 0, amount_all = 0)) %>%
mutate(amount_rate = ifelse(amount_all == 0, 0,
amount_2019 / amount_all))
df_amount_2019 %>%
filter(amount_rate > 0) %>%
slice(1:10)
# -
sapply(df_amount_2019, function(x) sum(is.na(x)))
# ---
# > R-085: 郵便番号(postal_cd)を用いて経度緯度変換用データフレーム(df_geocode)を紐付け、新たなdf_customer_1を作成せよ。ただし、複数紐づく場合は経度(longitude)、緯度(latitude)それぞれ平均を算出すること。
# +
df_customer_1 <- inner_join(df_customer[c("customer_id", "postal_cd")],
df_geocode[c("postal_cd", "longitude" ,"latitude")],
by="postal_cd") %>%
group_by(customer_id) %>%
summarise(m_longiture=mean(longitude),
m_latitude=mean(latitude), .groups = 'drop') %>%
inner_join(df_customer, by="customer_id")
head(df_customer_1, 5)
# -
# ---
# > R-086: 前設問で作成した緯度経度つき顧客データフレーム(df_customer_1)に対し、申込み店舗コード(application_store_cd)をキーに店舗データフレーム(df_store)と結合せよ。そして申込み店舗の緯度(latitude)・経度情報(longitude)と顧客の緯度・経度を用いて距離(km)を求め、顧客ID(customer_id)、顧客住所(address)、店舗住所(address)とともに表示せよ。計算式は簡易式で良いものとするが、その他精度の高い方式を利用したライブラリを利用してもかまわない。結果は10件表示すれば良い。
# $$
# 緯度(ラジアン):\phi \\
# 経度(ラジアン):\lambda \\
# 距離L = 6371 * arccos(sin \phi_1 * sin \phi_2
# + cos \phi_1 * cos \phi_2 * cos(\lambda_1 − \lambda_2))
# $$
# +
calc_distance <- function(x1, y1, x2, y2) {
distance <- 6371 * acos(
sin(y1 * pi / 180)
* sin(y2 * pi / 180)
+ cos(y1 * pi / 180)
* cos(y2 * pi / 180)
* cos((x1 * pi / 180)
- (x2 * pi / 180 )))
return(distance)
}
inner_join(df_customer_1, df_store,
by = c("application_store_cd" = "store_cd")) %>%
mutate(distance =
calc_distance(m_longiture, m_latitude, longitude, latitude)) %>%
select(customer_id, address.x, address.y, distance) %>%
slice(1:10)
# -
# ---
# > R-087: 顧客データフレーム(df_customer)では、異なる店舗での申込みなどにより同一顧客が複数登録されている。名前(customer_name)と郵便番号(postal_cd)が同じ顧客は同一顧客とみなし、1顧客1レコードとなるように名寄せした名寄顧客データフレーム(df_customer_u)を作成せよ。ただし、同一顧客に対しては売上金額合計が最も高いものを残すものとし、売上金額合計が同一もしくは売上実績の無い顧客については顧客ID(customer_id)の番号が小さいものを残すこととする。
# +
df_sales_amount <- df_receipt %>%
group_by(customer_id) %>%
summarise(sum_amount=sum(amount), .groups = 'drop')
df_customer_u <- left_join(df_customer, df_sales_amount, by="customer_id") %>%
arrange(desc(sum_amount), customer_id) %>%
distinct(customer_name, postal_cd, .keep_all=TRUE)
#減少件数をカウント
nrow(df_customer) - nrow(df_customer_u)
# -
# ---
# > R-088: 前設問で作成したデータを元に、顧客データフレームに統合名寄IDを付与したデータフレーム(df_customer_n)を作成せよ。ただし、統合名寄IDは以下の仕様で付与するものとする。
# > - 重複していない顧客:顧客ID(customer_id)を設定
# > - 重複している顧客:前設問で抽出したレコードの顧客IDを設定
df_customer_n = inner_join(df_customer,
df_customer_u[c("customer_id",
"customer_name",
"postal_cd")],
by=c("customer_name", "postal_cd")) %>%
rename(integration_id = customer_id.y)
# 確認
length(unique(df_customer_n$customer_id)) - length(unique(df_customer_n$integration_id))
# ---
# > R-089: 売上実績のある顧客に対し、予測モデル構築のため学習用データとテスト用データに分割したい。それぞれ8:2の割合でランダムにデータを分割せよ。
# +
#rsampleのinitial_splitを使った例
df_sales_amount <- df_receipt %>%
group_by(customer_id) %>%
summarise(sum_amount = sum(amount), .groups = 'drop')
df_customer_s <- inner_join(df_customer, df_sales_amount, by="customer_id")
df_split <- initial_split(df_customer_s, prop = 0.8)
df_customer_train <- training(df_split)
df_customer_test <- testing(df_split)
# 確認
print(nrow(df_customer_train))
print(nrow(df_customer_test))
# -
# ---
# > R-090: レシート明細データフレーム(df_receipt)は2017年1月1日〜2019年10月31日までのデータを有している。売上金額(amount)を月次で集計し、学習用に12ヶ月、テスト用に6ヶ月のモデル構築用データを3セット作成せよ。
# +
#caretのcreateTimeSlicesを使った例
df_ts_amount <- df_receipt %>%
group_by(substr(as.character(sales_ymd), 1, 6)) %>%
summarise(sum_amount = sum(amount), .groups = 'drop')
colnames(df_ts_amount) <- c("sales_ym","sales_amount")
timeSlice <- createTimeSlices(1:nrow(df_ts_amount),
initialWindow = 12, horizon = 6,
skip = 5, fixedWindow = TRUE)
df_train_1 <- df_ts_amount[timeSlice$train[[1]],]
df_train_2 <- df_ts_amount[timeSlice$train[[2]],]
df_train_3 <- df_ts_amount[timeSlice$train[[3]],]
df_test_1 <- df_ts_amount[timeSlice$test[[1]],]
df_test_2 <- df_ts_amount[timeSlice$test[[2]],]
df_test_3 <- df_ts_amount[timeSlice$test[[3]],]
# -
df_train_1
df_test_1
# ---
# > R-091: 顧客データフレーム(df_customer)の各顧客に対し、売上実績のある顧客数と売上実績のない顧客数が1:1となるようにアンダーサンプリングで抽出せよ。
# +
#recipesパッケージを使った例
#themis::step_downsample()を使うようワーニングが出るが、なるべくインストールパッケージを少なくするためrecipeのstep_downsample()を使用
df_sales_amount <- df_receipt %>%
group_by(customer_id) %>%
summarise(sum_amount = sum(amount), .groups = 'drop')
df_tmp <- left_join(df_customer, df_sales_amount, by="customer_id") %>%
replace_na(list(sum_amount = 0)) %>%
mutate(sum_amount = factor(ifelse(sum_amount > 0, 1, 0)))
set.seed(71)
df_down_train <- df_tmp %>%
recipe(~ .) %>%
step_downsample(sum_amount) %>%
prep() %>%
juice()
df_down_train %>%
group_by(sum_amount) %>%
summarise(cnt = n(), .groups = 'drop')
# -
head(df_down_train, 3)
# +
# #unbalancedのubUnderを使った例(unbalamncedの依存関係にあるxmlパッケージがR3.6.3でインストールできなくなっているためコメントアウト中※2020.07.29時点)
# #unbalancedも良く利用されるのでコードは残しておく
# df_sales_amount <- df_receipt %>%
# group_by(customer_id) %>%
# summarise(sum_amount = sum(amount))
# df_tmp <- left_join(df_customer, df_sales_amount, by="customer_id") %>%
# replace_na(list(sum_amount = 0)) %>%
# mutate(sum_amount = ifelse(sum_amount > 0, 1, 0))
# data_under<-ubUnder(X=df_tmp[,colnames(df_tmp) != "sum_amount"], Y= df_tmp$sum_amount, perc = 50, method = "percPos")
# df_down_sampling <- cbind(data_under$X, data_under$Y) %>%
# rename(sum_amount = 'data_under$Y')
# -
# ---
# > R-092: 顧客データフレーム(df_customer)では、性別に関する情報が非正規化の状態で保持されている。これを第三正規化せよ。
# +
df_gender = unique(df_customer[c("gender_cd", "gender")])
df_customer_s = df_customer[, colnames(df_customer) != "gender"]
# -
# ---
# > R-093: 商品データフレーム(df_product)では各カテゴリのコード値だけを保有し、カテゴリ名は保有していない。カテゴリデータフレーム(df_category)と組み合わせて非正規化し、カテゴリ名を保有した新たな商品データフレームを作成せよ。
df_product_full <- inner_join(df_product,
df_category[c("category_small_cd",
"category_major_name",
"category_medium_name",
"category_small_name")],
by="category_small_cd")
# ---
# > R-094: 先に作成したカテゴリ名付き商品データを以下の仕様でファイル出力せよ。なお、出力先のパスはdata配下とする。
# > - ファイル形式はCSV(カンマ区切り)
# > - ヘッダ有り
# > - 文字コードはUTF-8
write.csv(df_product_full, "../data/R_df_product_full_UTF-8_header.csv",
fileEncoding="UTF-8")
# ---
# > R-095: 先に作成したカテゴリ名付き商品データを以下の仕様でファイル出力せよ。なお、出力先のパスはdata配下とする。
# > - ファイル形式はCSV(カンマ区切り)
# > - ヘッダ有り
# > - 文字コードはCP932
write.csv(df_product_full, "../data/R_df_product_full_CP932_header.csv",
fileEncoding="CP932")
# ---
# > R-096: 先に作成したカテゴリ名付き商品データを以下の仕様でファイル出力せよ。なお、出力先のパスはdata
# 配下とする。
# > - ファイル形式はCSV(カンマ区切り)
# > - ヘッダ無し
# > - 文字コードはUTF-8
write.table(df_product_full, "../data/R_df_product_full_UTF-8_noh.csv",
col.names=FALSE, sep=",", fileEncoding="UTF-8")
# ---
# > R-097: 先に作成した以下形式のファイルを読み込み、データフレームを作成せよ。また、先頭10件を表示させ、正しくとりまれていることを確認せよ。
# > - ファイル形式はCSV(カンマ区切り)
# > - ヘッダ有り
# > - 文字コードはUTF-8
df_product_tmp <- read.csv('../data/R_df_product_full_UTF-8_header.csv',
fileEncoding = "UTF-8")
head(df_product_tmp, 10)
# ---
# > R-098: 先に作成した以下形式のファイルを読み込み、データフレームを作成せよ。また、先頭10件を表示させ、正しくとりまれていることを確認せよ。
# > - ファイル形式はCSV(カンマ区切り)
# > - ヘッダ無し
# > - 文字コードはUTF-8
df_product_tmp <- read.csv('../data/R_df_product_full_UTF-8_noh.csv',
header=FALSE, fileEncoding="UTF-8")
head(df_product_tmp, 10)
# ---
# > R-099: 先に作成したカテゴリ名付き商品データを以下の仕様でファイル出力せよ。なお、出力先のパスはdata配下とする。
# > - ファイル形式はTSV(タブ区切り)
# > - ヘッダ有り
# > - 文字コードはUTF-8
write.table(df_product_full, "../data/R_df_product_full_UTF-8_head.tsv",
sep="\t", fileEncoding="UTF-8")
# ---
# > R-100: 先に作成した以下形式のファイルを読み込み、データフレームを作成せよ。また、先頭10件を表示させ、正しくとりまれていることを確認せよ。
# > - ファイル形式はTSV(タブ区切り)
# > - ヘッダ有り
# > - 文字コードはUTF-8
df_product_tmp <- read.table("../data/R_df_product_full_UTF-8_head.tsv",
fileEncoding="UTF-8")
head(df_product_tmp,10)
# # これで100本終わりです。おつかれさまでした!
| docker/work/answer/ans_preprocess_knock_R.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7 fastai2
# language: python
# name: fastai2
# ---
# # Training a deep CNN to learn about galaxies in 15 minutes
# > Let's train a deep neural network from scratch! In this post, I provide a demonstration of how to optimize a model in order to predict galaxy metallicities using images, and I discuss some tricks for speeding up training and obtaining better results.
#
# - toc: true
# - badges: true
# - comments: true
# - author: <NAME>
# - categories: [galaxies, astrophysics, deep learning, computer vision, fastai]
# - image: images/train-cnn.png
#
# +
#hide
from fastai2.basics import *
from fastai2.vision.all import *
from mish_cuda import MishCuda
from matplotlib.colors import LogNorm
ROOT = Path('../').resolve()
# -
# # Predicting metallicities from pictures: obtaining the data
#
# In my [previous post](https://github.com/jwuphysics/blog/blob/master/_notebooks/2020-05-21-exploring-galaxies-with-deep-learning.ipynb), I described the problem that we now want to solve. To summarize, we want to train a convolutional neural network (CNN) to perform regression. The inputs are images of individual galaxies (although sometimes we're photobombed by other galaxies). The outputs are metallicities, $Z$, which usually take on a value between 7.8 and 9.4.
# The first step, of course, is to actually get the data. Galaxy images can be fetched using calls to the Sloan Digital Sky Survey (SDSS) SkyServer `getJpeg` cutout service via their [RESTful API](http://skyserver.sdss.org/dr16/en/help/docs/api.aspx#imgcutout). For instance, [this URL](http://skyserver.sdss.org/dr16/SkyserverWS/ImgCutout/getjpeg?ra=39.8486&dec=1.094&scale=1&width=224&height=224) grabs a three-channel, $224 \times 224$-pixel JPG image:
# 
#
#
# Galaxy metallicities can be obtained from the SDSS SkyServer using a [SQL query](http://skyserver.sdss.org/dr16/en/help/docs/sql_help.aspx) and a bit of `JOIN` magic. All in all, we use 130,000 galaxies with metallicity measurements as our training + validation data set.
# The code for the original published work ([Wu & Boada 2019](https://ui.adsabs.harvard.edu/abs/2019MNRAS.484.4683W/abstract)) can be found in my [Github repo](https://github.com/jwuphysics/galaxy-cnns). However, this code (from 2018) used `fastai` *version 0.7*, and I want to show an updated version using the new and improved `fastai` *version 2* codebase. Also, some of the "best practices" for deep learning and computer vision have evolved since then, so I'd like to highlight those updates as well!
# # Organizing the data using the `fastai` DataBlock API
# Suppose that we now have a directory full of galaxy images, and a `csv` file with the object identifier, coordinates, and metallcity for each galaxy. The `csv` table can be [read using Pandas](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.read_csv.html), so let's store that in a DataFrame `df`. We can take a look at five random rows of the table by calling `df.sample(5)`:
#hide_input
df = pd.read_csv(f'{ROOT}/data/master.csv', dtype={'objID': str}).rename({'oh_p50': 'metallicity'}, axis=1)
df[['objID', 'ra', 'dec', 'metallicity']].sample(5)
# This means that a galaxy with `objID` 1237654601557999788 is located at RA = 137.603036 deg, Dec = 3.508882 deg, and has a metallicity of $Z$ = 8.819281. Our directory structure is such that the corresponding image is stored in `{ROOT}/images/1237660634922090677.jpg`, where `ROOT` is the path to project repository.
#
# A tree-view from our `{ROOT}` directory might look like this:
# ```
# .
# ├── data
# │ └── master.csv
# ├── images
# │ ├── 1237654601557999788.jpg
# │ ├── 1237651067353956485.jpg
# │ └── [...]
# └── notebooks
# └── training-a-cnn.ipynb
# ```
# We are ready to set up our `DataBlock`, which is a core fastai construct for handling data. The process is both straightforward and extremely powerful, and comprises a few steps:
# - Define the inputs and outputs in the `blocks` argument
# - Specify how to get your inputs (`get_x`) and outputs (`get_y`)
# - Decide how to split the data into training and validation sets (`splitter`)
# - Define any CPU-level transformations (`item_tfms`) and GPU-level transformations (`batch_tfms`) used for preprocessing or augmenting your data.
#
# Before going into the details for each component, here is the code in action:
dblock = DataBlock(
blocks=(ImageBlock, RegressionBlock),
get_x=ColReader(['objID'], pref=f'{ROOT}/images/', suff='.jpg'),
get_y=ColReader(['metallicity']),
splitter=RandomSplitter(0.2),
item_tfms=[CropPad(144), RandomCrop(112)],
batch_tfms=aug_transforms(max_zoom=1., flip_vert=True, max_lighting=0., max_warp=0.) + [Normalize],
)
# Okay, now let's take a look at each part.
# ## Input and output blocks
# First, we want to make use of the handy `ImageBlock` class for handling our input images. Since we're using galaxy images in the JPG format, we can rely on the `PIL` backend of `ImageBlock` to open the images efficiently. If, for example, we instead wanted to use images in the astronomical `FITS` format, we could extend the `TensorImage` class and define the following bit of code:
# +
#collapse-hide
class FITSImage(TensorImage):
@classmethod
def create(cls, filename, chans=None, **kwargs) -> None:
"""Create FITS format image by using Astropy to open the file, and then
applying appropriate byte swaps and flips to get a Pytorch Tensor.
"""
return cls(
torch.from_numpy(
astropy.io.fits.getdata(fn).byteswap().newbyteorder()
)
.flip(0)
.float()
)
def show(self, ctx=None, ax=None, vmin=None, vmax=None, scale=True, title=None):
"""Plot using matplotlib or your favorite program here!"""
pass
FITSImage.create = Transform(FITSImage.create)
def FITSImageBlock():
"""A FITSImageBlock that can be used in the fastai DataBlock API.
"""
return TransformBlock(partial(FITSImage.create))
# -
# For our task, the vanilla `ImageBlock` will suffice.
#
# We also want to define an output block, which will be a `RegressionBlock` for our task (note that it handles both single- and multi-variable regression). If, for another problem, we wanted to do a categorization problem, then we'd intuitively use the `CategoryBlock`. Some other examples of the DataBlock API can be found in the [documentation](http://dev.fast.ai/data.block).
#
# We can pass in these arguments in the form of a tuple: `blocks=(ImageBlock, RegressionBlock)`.
# ## Input and output object getters
# Next, we want to be able to access the table, `df`, which contain the columns `objID` and `metallicity`. As we've discussed above, each galaxy's `objID` can be used to access the JPG image on disk, which is stored at `{ROOT}/images/{objID}.jpg`. Fortunately, this is easy to do with the fastai `ColumnReader` method! We just have to supply it with the column name (`objID`), a prefix (`{ROOT}/images/`), and a suffix (`.jpg`); since the prefix/suffix is only used for file paths, the function knows that the file needs to be opened (rather than interpreting it as a string). So far we have:
# ```python
# get_x=ColReader(['objID'], pref=f'{ROOT}/images/', suff='.jpg')
# ```
#
# The targets are stored in `metallicity`, so we can simply fill in the `get_y` argument:
# ```python
# get_y=ColReader(['metallicity'])
# ```
#
# (At this point, we haven't yet specified that `df` is the DataFrame we're working with. The `DataBlock` object knows how to handle the input/output information, but isn't able to load it until we provide it with `df` -- that will come later!)
# ## Splitting the data set
# For the sake of simplicity, we'll just randomly split our data set using the aptly named `RandomSplitter` function. We can provide it with a number between 0 and 1 (corresponding to the fraction of data that will become the validation set), and also a random seed if we wish. If we want to set aside 20% of the data for validation, we can use this:
#
# ```python
# splitter=RandomSplitter(0.2, seed=56)
# ```
# ## Transformations and data augmentation
# Next, I'll want to determine some data augmentation transformations. These are handy for varying our image data: crops, flips, and rotations can be applied at random using fastai's `aug_transforms()` in order to dramatically expand our data set. Even though we have >100,000 unique galaxy images, our CNN model will contain millions of trainable parameters. Augmenting the data set will be especially valuable for mitigating overfitting.
#
# Translations, rotations, and reflections to our images should not change the properties of our galaxies. However, we won't want to zoom in and out of the images, since that might impact CNN's ability to infer unknown (but possibly important) quantities such as the galaxies' intrinsic sizes. Similarly, color shifts or image warps may alter the star formation properties or stellar structures of the galaxies, so we don't want to mess with that.
#
# We will center crop the image to $144 \times 144$ pixels using `CropPad()`, which reduces some of the surrounding black space (and other galaxies) near the edges of the images. We will then apply a $112 \times 112$-pixel `RandomCrop()` for some more translational freedom. This first set of image crop transformations, `item_tfms`, will be performed on images one by one using a CPU. Afterwards, the cropped images (which should all be the same size) will be loaded onto the GPU. At this stage, data augmentation transforms will be performed along with image normalization, which rescales the intensities in each channel so that they have zero mean and unit variance. The second set of transformations, `batch_tfms`, will be applied one batch at a time on the GPU.
item_tfms=[CropPad(144), RandomCrop(112)]
batch_tfms=aug_transforms(max_zoom=1., flip_vert=True, max_lighting=0., max_warp=0.) + [Normalize]
# > Note: `Normalize` will pull the batch statistics from your images, and apply it any time you load in new data (see below). Sometimes this can lead to unintended consequences, for example, if you're loading in a test data set which is characterized by different image statistics. In that case, I recommend saving your batch statistics and then using them later, e.g., `Normalize.from_stats(*image_statistics)`.
# ## Putting it all together and loading the data
# We've now gone through each of the steps, but we haven't yet loaded the data! `ImageDataLoaders` has a class method called `from_dblock()` that loads everything in quite nicely if we give it a data source. We can pass along the `DataBlock` object that we've constructed, the DataFrame `df`, the file path `ROOT`, and a batch size. We've set the batch size `bs=128` because that fits on the GPU, and it ensures speedy training, but I've found that values between 32 and 128 often work well.
dls = ImageDataLoaders.from_dblock(dblock, df, path=ROOT, bs=128)
# Once this is functional, we can view our data set! As we can see, the images have been randomly cropped such that the galaxies are not always in the center of the image. Also, much of the surrounding space has been cropped out.
dls.show_batch(nrows=2, ncols=4)
# Pardon the excessive number of significant figures. We can fix this up by creating custom classes extending `Transform` and `ShowTitle`, but this is beyond the scope of the current project. Maybe I'll come back to this in a future post!
# # Neural network architecture and optimization
# 
# There's no way that I can describe all of the tweaks and improvements that machine learning researchers have made in the past couple of years, but I'd like to highlight a few that really help out our cause. We need to use some kind of residual CNNs (or resnets), introduced by [Kaiming He et al. (2015)](https://arxiv.org/abs/1512.03385). Resnets outperform previous CNNs such as the AlexNet or VGG architectures because they can leverage gains from "going deeper" (i.e., by extending the resnets with additional layers). The paper is quite readable and interesting, and there are plenty of other works explaining why resnets are so successful (e.g., a [blog post by <NAME>](http://teleported.in/posts/decoding-resnet-architecture/) and [a deep dive into residual blocks by He et al.](https://arxiv.org/abs/1603.05027)).
# 
# In `fastai`, we can instantiate a 34-layer *enhanced* resnet model by using `model = xresnet34()`. We could have created a 18-layer model with `model = xresnet18()`, or even defined our own custom 9-layer resnet using
# ```python
# xresnet9 = XResNet(ResBlock, expansion=1, layers=[1, 1, 1, 1])
# model = xresnet9()
# ```
# But first, we need to set the number of outputs. By default, these CNNs are suited for the ImageNet classification challenge, and so there are `1000` outputs. Since we're performing single-variable regression, the number of outputs (`n_out`) should be `1`. Our `DataLoaders` class, `dls`, already knows this and has stored the value `1` in `dls.c`.
# Okay, let's make our model for real:
model = xresnet34(n_out=dls.c, sa=True, act_cls=MishCuda)
# So why did I say that we're using an "enhanced" resnet -- an "xresnet"? And what does `sa=True` and `act_cls=MishCuda` mean? I'll describe these tweaks below.
# ## A more powerful resnet
# The ["bag of tricks" paper](https://arxiv.org/abs/1812.01187) by Tong He et al. (2018) summarizes many small tweaks that can be combined to dramatically improve the performance of a CNN. They describe several updates to the resnet model architecture in Section 4 of their paper. The fastai library takes these into account, and also implements a few other tweaks, in order to increase performance and speed. I've listed some of them below:
# - The CNN stem (first few layers) is updated using efficient $3 \times 3$ convolutions rather than a single expensive layer of $7\times 7$ convolutions.
# - Residual blocks are changed so that $1 \times 1$ convolutions don't skip over useful information. This is done by altering the order of convolution strides in one path of the downsampling block, and adding a pooling layer in the other path (see Figure 2 of He et al. 2018).
# - The model concatenates the outputs of both AveragePool and MaxPool layers (using `AdaptiveConcatPool2d`) rather than using just one.
#
# Some of these tweaks are described in greater detail in [Chapter 14](https://github.com/fastai/fastbook/blob/master/14_resnet.ipynb) of the fastai book, "Deep Learning for Coders with fastai and Pytorch" (which can be also be purchased on [Amazon](https://www.amazon.com/Deep-Learning-Coders-fastai-PyTorch/dp/1492045527)).
# ## Self-attention layers
# The concept of attention has gotten a lot of, well, *attention* in deep learning, particularly in natural language processing (NLP). This is because the attention mechanism is a core part of the [Transformer architecture](https://arxiv.org/abs/1706.03762), which has revolutionized our ability to learn from text data. I won't cover the Transformer architecture or NLP in this post, since it's way out of scope, but suffice it to say that lots of deep learning folks are interested in this idea.
# 
#
# The attention mechanism allows a neural network layer to encode interactions from inputs on scales larger than the size of a typical convolutional filter. Self-attention is simply when these relationships, encoded via a *query/key/value* system, are applied using the same input. As a concrete example, self-attention added to CNNs in our scenario -- estimating metallicity from galaxy images -- may allow the network to learn morphological features that often require long-range dependencies, such as the orientation and position angle of a galaxy.
#
# In fastai, we can set `sa=True` when initializing a CNN in order to get the [self-attention layers](https://github.com/fastai/fastai/blob/master/fastai/layers.py#L288)!
# Another way to let a CNN process global information is to use [Squeeze-and-Excitation Networks](https://arxiv.org/abs/1709.01507), which are also [included in fastai](https://github.com/fastai/fastai2/blob/44cc025d9e5e2823d6fd033b84245b0be0c5c9df/fastai2/layers.py#L562). Or, one could even [entirely replace convolutions with self-attention](https://arxiv.org/abs/1906.05909). But we're starting to get off-topic...
# ## The Mish activation function
# Typically, the Rectified Linear Unit (ReLU) is the non-linear activation function of choice for nearly all deep learning tasks. It is both cheap to compute and simple to understand: `ReLU(x) = max(0, x)`.
#
# That was all before <NAME> introduced us to the [Mish activation function](https://github.com/digantamisra98/Mish) -- as an undergraduate researcher! He also [wrote a paper](https://arxiv.org/abs/1908.08681) and summarizes some of the reasoning behind it in a [forum post](https://forums.fast.ai/t/meet-mish-new-activation-function-possible-successor-to-relu/53299). <NAME>, from the fastai community, shows that [it performs extremely well](https://medium.com/@lessw/meet-mish-new-state-of-the-art-ai-activation-function-the-successor-to-relu-846a6d93471f) in several image classification challenges. I've also found that Mish is perfect as a drop-in replacement for ReLU in regression tasks.
#
# The intuition behind the Mish activation function's success is similar to the reason why resnets perform so well: the loss landscape becomes smoother and thereby easier to explore. ReLU is non-differentiable at the origin, causing steep spikes in the loss. Mish resembles another activation function, [GELU (or SiLU)](https://openreview.net/pdf?id=Bk0MRI5lg), in that neither it nor its derivative is monotonic; this seems to lead to more complex and nuanced behavior during training. However, it's not clear (from a theoretical perspective) why such activation functions empirically perform so well.
#
# Although Mish is a little bit slower than ReLU, a [CUDA implementation](https://github.com/thomasbrandon/mish-cuda/) helps speed things up a bit. We need to `pip install` it and then import it with `from mish_cuda import MishCuda`. Then, we can substitute it into the model when initializing our CNN using `act_cls=MishCuda`.
# ## RMSE loss
# Next we want to select a loss function. The mean squared error (MSE) is suitable for training the network, but we can more easily interpret the root mean squared error (RMSE). We need to create a function to compute the RMSE loss between predictions `p` and true metalllicity values `y`.
#
# (Note that we use `.view(-1)` to flatten our Pytorch `Tensor` objects since we're only predicting a single variable.)
def root_mean_squared_error(p, y):
return torch.sqrt(F.mse_loss(p.view(-1), y.view(-1)))
# ## Ranger: a combined RAdam + LookAhead optimzation function
# Around mid-2019, we saw two new papers regarding the stability of training neural networks: [LookAhead](https://arxiv.org/abs/1907.08610) and [Rectified Adam (RAdam)](https://arxiv.org/abs/1908.03265). Both papers feature novel optimizers that address the problem of excess variance during training. LookAhead mitigates the variance problem by scouting a few steps ahead, and then choosing how to optimally update the model's parameters. RAdam adds a term while computing the adaptive learning rate in order to address training instabilities (see, e.g., the original [Adam optimizer](https://arxiv.org/abs/1412.6980)).
#
# Less Wright quickly realized that these two optimizers [could be combined](https://forums.fast.ai/t/meet-ranger-radam-lookahead-optimizer/52886). His [`ranger` optimizer](https://github.com/lessw2020/Ranger-Deep-Learning-Optimizer) is the product of these two papers (and now also includes a new tweak, [gradient centralization](https://arxiv.org/abs/2004.01461v2), by default). I have found `ranger` to give excellent results using empirical tests.
# So, now we'll put everything together in a fastai `Learner` object:
learn = Learner(
dls,
model,
opt_func=ranger,
loss_func=root_mean_squared_error
)
# ## Selecting a learning rate
# Fastai offers a nice feature for determining an optimal learning rate, taken from [<NAME> (2015)](https://arxiv.org/abs/1506.01186). All we have to do is call `learn.lr_find()`.
#
# The idea is to begin feeding your CNN batches of data, while exponentially increasing learning rates (i.e., step sizes) and monitoring the loss. At some point the loss will bottom out, and then begin to increase and diverge wildly, which is a sign that the learn rate is now too high.
#
# 
#
# Generally, before the loss starts to diverge, the learning rate will be suitable for the loss to steadily decrease. We can generally read an optimal learning rate off the plot -- the suggested learning rate is around $0.03$ (since that is about an order of magnitude below the learning rate at which the loss "bottoms out" and is also where the loss is decreasing most quickly). I tend to choose a slightly lower learning rate (here I'll select $0.01$), since that seems to work better for my regression problems.
learn.lr_find()
# ## Training the neural network with a "one-cycle" schedule
# Finally, now that we've selected a learning rate ($0.01$), we can train for a few epochs. Remember that an *epoch* is just a run-through using all of our training data (and we send in one batch of 64 images at a time). Sometimes, researchers simply train at a particular learning rate and wait until the results converge, and then lower the learning rate in order for the model to continue learning. This is because the model needs some serious updates toward the beginning of training (given that it has been initialized with random weights), and then needs to start taking smaller steps once its weights are in the right ballpark. However, the learning rate can't be too high in th beginning, or the loss will diverge! Traditionally, researchers will select a safe (i.e., low) learning rate in the beginning, which can take a long time to converge.
#
# Fastai offers a few optimization *schedules*, which involve altering the learning rate over the course of training. The two most promising are called [`fit_flat_cos`](https://dev.fast.ai/callback.schedule#Learner.fit_flat_cos) and [`fit_one_cycle`](https://dev.fast.ai/callback.schedule#Learner.fit_one_cycle) ([see more here](https://arxiv.org/abs/1708.07120)). I've found that `fit_flat_cos` tends to work better for classification tasks, while `fit_one_cycle` tends to work better for regression problems. Either way, the empirical results are fantastic -- especially coupled with the Ranger optimizer and all of the other tweaks we've discussed.
learn.fit_one_cycle(7, 1e-2)
# Here we train for only seven epochs, which took under 14 minutes of training on a single NVIDIA P100 GPU, and achieve a validation loss of 0.086 dex. In our published paper, we were able to reach a RMSE of 0.085 dex in under 30 minutes of training, but that wasn't from a randomly initialized CNN -- we were using transfer learning then! Here we can accomplish similar results, without pre-training, in only half the time.
# We can visualize the training and validation losses. The x-axis shows the number of training iterations (i.e., batches), and the y-axis shows the RMSE loss.
learn.recorder.plot_loss()
plt.ylim(0, 0.4);
# # Evaluating our results
# Finally, we'll perform another round of [data augmentation](#Transformations-and-data-augmentation) on the validation set in order to see if the results improve. This can be done using `learn.tta()`, where TTA stands for test-time augmentation.
preds, trues = learn.tta()
# Note that we'll want to flatten these `Tensor` objects and convert them to numpy arrays, e.g., `preds = np.array(preds.view(-1))`. At this point, we can plot our results. Everything looks good!
#hide
preds = to_np(preds.view(-1))
trues = to_np(trues.view(-1))
# +
#hide_input
plt.figure(figsize=(4, 4), dpi=150)
plt.hist2d(trues, preds, bins=30, range=[(8.2, 9.3), (8.2, 9.3)], cmap='Blues', norm=LogNorm(vmin=1, vmax=1000))
plt.xlim(8.2, 9.3)
plt.ylim(8.2, 9.3)
plt.gca().set_aspect('equal')
plt.text(8.65, 8.25, f'RMSE = {np.mean((preds-trues)**2)**0.5:.4f} dex', fontsize=12)
plt.xlabel(r'$Z_{\rm true}$', fontsize=12)
plt.ylabel(r'$Z_{\rm pred}$', fontsize=12)
# -
# It appears that we didn't get a lower RMSE using TTA, but that's okay. TTA is usually worth a shot after you've finished training, since evaluating the neural network is relatively quick.
# # Summary
# In summary, we were able to train a deep convolutional neural network to predict galaxy metallicity from three-color images in under 15 minutes. Our data set contained over 100,000 galaxies, so this was no easy feat! Data augmentation, neural network architecture design, and clever optimization tricks were essential for improving performance. With these tools in hand, we can quickly adapt our methodology to tackle many other kinds of problems!
#
# `fastai` version 2 is a powerful high-level library that extends Pytorch and is easy to use/customize. As of November 2020, the [documentation](https://docs.fast.ai/) is still a bit lacking, but hopefully will continue to mature. One big takeaway is that fastai, which is all about *democratizing AI*, makes deep learning more accessible than ever before.
# **Acknowledgments**: I want to thank fastai core development team, [<NAME>](https://twitter.com/jeremyphoward) and [<NAME>](https://twitter.com/GuggerSylvain), as well as other contributors and invaluable members of the community, including [Less Wright](https://github.com/lessw2020), [<NAME>](https://twitter.com/digantamisra1?lang=en), and [<NAME>](https://muellerzr.github.io/). I also want to acknowledge Google for their support via GCP credits for academic research. Finally, I want to give a shout out to [<NAME>](https://github.com/boada), my original collaborator and co-author on [our paper](https://arxiv.org/abs/1810.12913).
#
# **Last updated**: November 16, 2020
| _notebooks/2020-05-26-training-a-deep-cnn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Implementing Bag of Words from scratch
# +
documents=[]
for line in open(r"C:/Users/<NAME>/Desktop/Projects 2021/Pyrated/if.txt"):
documents.append(line.rstrip().strip().strip(',').strip(':').strip('!').strip('?').strip(';'))
# converting all the words to unique case to avoid redundancy of words with upper case and lower case.
# removing various unnecessary data which doesn't convey any special meaning such as , . ! ? etc.
updated_documents = []
for i in documents:
updated_documents.append(i.lower())
updated_documents
# -
preprocessed_documents = []
for i in updated_documents:
preprocessed_documents.append(i.split(' '))
print(preprocessed_documents)
# +
frequency_list = []
import pprint
from collections import Counter
for i in preprocessed_documents:
frequency_list.append(Counter(i))
frequency_list
# +
# Implementing Bag of Words in scikit-learn
# -
import pandas as pd
documents=[]
for line in open(r"C:/Users/<NAME>/Desktop/Projects 2021/Pyrated/if.txt"):
documents.append(line.rstrip().strip().strip(','))
documents
from sklearn.feature_extraction.text import CountVectorizer
count_vector = CountVectorizer(documents)
print(count_vector)
count_vector.fit(documents)
count_vector.get_feature_names()
doc_array = count_vector.transform(documents).toarray()
doc_array
frequency_matrix = pd.DataFrame(doc_array,index=documents,columns=count_vector.get_feature_names())
frequency_matrix
| BagOfWords.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# This notebook briefly shows capabilities of CardIO framework.
#
# For details refer to other [tutorials](https://github.com/analysiscenter/cardio/tree/master/tutorials) or [CardIO documentation](https://analysiscenter.github.io/cardio/index.html)
# You can also find use of article [CardIO framework for deep research of electrocardiograms](https://medium.com/data-analysis-center/cardio-framework-for-deep-research-of-electrocardiograms-2a38a0673b8e), which is closely related to this notebook.
# Some general imports
# +
import os
import sys
sys.path.append("..")
import cardio.dataset as ds
from cardio import EcgBatch
from cardio.models.metrics import classification_report
# %env CUDA_VISIBLE_DEVICES=0
import tensorflow as tf
# -
# Create indices from filenames
index = ds.FilesIndex(path="../cardio/tests/data/A*.hea", no_ext=True, sort=True)
# Check which indices are in list of indices
print(index.indices)
# Create dataset
eds = ds.Dataset(index, batch_class=EcgBatch)
# Generate batch
batch = eds.next_batch(batch_size=2)
# Fill batch with data
batch_with_data = batch.load(fmt="wfdb", components=["signal", "meta"])
# Plot short segment of ECG with index 'A00001'
batch_with_data.show_ecg('A00001', start=10, end=15)
# For detection of QRS intervals, P-waves and T-waves we need to train model first.
#
# Download the [QT Database](https://www.physionet.org/physiobank/database/qtdb/) with labeled ECGs. Let ```SIGNALS_PATH``` be the folder where database is saved.
# +
from cardio.pipelines import hmm_preprocessing_pipeline, hmm_train_pipeline
import warnings
warnings.filterwarnings('ignore')
SIGNALS_PATH = "path_to_QT_database" #set path to QT database
SIGNALS_MASK = os.path.join(SIGNALS_PATH, "*.hea")
index = ds.FilesIndex(path=SIGNALS_MASK, no_ext=True, sort=True)
dtst = ds.Dataset(index, batch_class=EcgBatch)
pipeline = hmm_preprocessing_pipeline()
ppl_inits = (dtst >> pipeline).run()
pipeline = hmm_train_pipeline(ppl_inits)
ppl_train = (dtst >> pipeline).run()
# -
# Save model
ppl_train.save_model("HMM", path="model_dump.dill")
# Let's make prediction with hmm model
# +
from cardio.pipelines import hmm_predict_pipeline
batch = (eds >> hmm_predict_pipeline("model_dump.dill", annot="hmm_annotation")).next_batch()
# -
# Plot ECG signal with detected QRS intervals, P-waves and T-waves
batch.show_ecg('A00001', start=10, end=15, annot="hmm_annotation")
# For detection of atrial fibrillation we need to train another model.
# Download the PhysioNet short single lead ECG database from [here](https://physionet.org/challenge/2017/). Let AF_SIGNALS_PATH be the folder where database is saved.
# +
from cardio.pipelines import dirichlet_train_pipeline
gpu_options = tf.GPUOptions(per_process_gpu_memory_fraction=0.33, allow_growth=True)
AF_SIGNALS_PATH = "path_to_PhysioNet_database" #set path to PhysioNet database
AF_SIGNALS_MASK = os.path.join(AF_SIGNALS_PATH, "*.hea")
AF_SIGNALS_REF = os.path.join(AF_SIGNALS_PATH, "REFERENCE.csv")
index = ds.FilesIndex(path=AF_SIGNALS_MASK, no_ext=True, sort=True)
afds = ds.Dataset(index, batch_class=EcgBatch)
pipeline = dirichlet_train_pipeline(AF_SIGNALS_REF, gpu_options=gpu_options)
train_ppl = (afds >> pipeline).run()
# -
# Save model
model_path = "af_model_dump"
train_ppl.save_model("dirichlet", path=model_path)
# +
from cardio.pipelines import dirichlet_predict_pipeline
pipeline = dirichlet_predict_pipeline(model_path, gpu_options=gpu_options)
res = (eds >> pipeline).run()
pred = res.get_variable("predictions_list")
# -
# Get predicted probalilities for atrial fibrillation
print(["{:.2f}".format(x["target_pred"]["A"]) for x in pred])
| examples/Getting_started.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Using ORF finder to analyze drosophila sequence
# ### Objective
# The goal is to determine if a recently-obtained genomic sequence from Drosophila yakuba (a relative of the model fruit fly Drosophila melanogaster) contains region(s) with sequence similarity to any known genes. The unknown sequence is an 11,000 base pair (bp) fragment of genomic DNA, and the objective of gene annotation is to find and precisely map the coding regions of any genes in this part of the genome.
# ### Overview about the ORF
# ORF finder searches for open reading frames (ORFs) in the DNA sequence you enter.
# The program returns the range of each ORF, along with its protein translation.
# Use ORF finder to search newly sequenced DNA for potential protein encoding segments,
# verify predicted protein using newly developed SMART BLAST or regular BLASTP.
# ### Select Drosophila yakuba blasted sequence
sequence = 'NC_011091.1'
# ### Find desired open reading frame using ORF viewer
# ORF viewer shows list of all possible ORF from desired sequences
# ### Mark down ORF
# Mark down single or multiple ORFs for blast search.
# ### Use blast to determine matching proteins
# orf sequence --> blast alignmnet --> matching proteins
#
| notebooks/Analyze_Unknown_Blast_Sequences_Using_NCBI_Databases.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import numpy as np
X = 2 * np.random.rand(100, 1)
y = 4 + 3 * X + np.random.randn(100, 1)
print(X.shape, y.shape)
# +
import matplotlib.pyplot as plt
plt.scatter(X, y)
plt.show()
# -
# compute theta-hat using Normal Equation
# (X_T * X)-1 X_T * y
X_b = np.c_[np.ones((100, 1)), X]
X_b.shape
theta_best = np.linalg.inv(X_b.T.dot(X_b)).dot(X_b.T).dot(y)
theta_best
X_new = np.array([[0], [2]])
X_new_b = np.c_[np.ones((2, 1)), X_new]
y_predict = X_new_b.dot(theta_best)
y_predict
plt.plot(X_new, y_predict, 'r-')
plt.plot(X, y, 'b.')
plt.axis([0, 2, 0, 15])
plt.show()
# +
# using sklearn
from sklearn.linear_model import LinearRegression
reg = LinearRegression()
reg.fit(X, y)
reg.intercept_, reg.coef_
# -
reg.predict(X_new)
# based on least squares
theta_best_svd, residuals, rank, s = np.linalg.lstsq(X_b, y, rcond=1e-6)
theta_best_svd
# psuedoinverse of X
np.linalg.pinv(X_b).dot(y)
# computational complexity of X_T * X is about O(n^2.4) to O(n^3)
# SVD approached used by LinearRegression is O(n^2)
# Both Normal Equation and SVD get slow when features > 100K
# However they are linear in the size of the training set (m) i.e. O(m)
#
# +
# Gradient descent
# not all costs functions are convex
# MSE cost function for linear reg is though => guaranteed to approach global minimum
# ensure all features have a similar scale (StandardScaler) to ensure speedy convergence
eta = 0.1 # learning rate
n_iter = 1000
m = 100 # batch size
theta = np.random.randn(2, 1)
for i in range(n_iter):
gradients = 2 / m * X_b.T.dot(X_b.dot(theta) - y)
theta -= eta * gradients
theta
# +
# stochastic gradient descent / simulated annealing
# learning schedule determines learning rate
n_epochs = 50
t0, t1 = 5, 50 # learning schedule hyperparameters
def learning_schedule(t):
return t0 / (t + t1)
theta = np.random.randn(2, 1)
for epoch in range(n_epochs):
for i in range(m):
r = np.random.randint(m)
xi = X_b[r: r + 1]
yi = y[r: r + 1]
gradients = 2 * xi.T.dot(xi.dot(theta) - yi)
eta = learning_schedule(epoch * m + i)
theta -= eta * gradients
theta
# +
from sklearn.linear_model import SGDRegressor
reg = SGDRegressor(max_iter=50, penalty=None, eta0=0.1)
reg.fit(X, y.ravel())
# -
print(reg.intercept_, reg.coef_)
# +
# Polynomial regression
m = 100
X = 6 * np.random.rand(m, 1) - 3
y = 0.5 * X**2 + X + 2 + np.random.randn(m, 1)
plt.scatter(X, y)
# -
from sklearn.preprocessing import PolynomialFeatures
poly_features = PolynomialFeatures(degree=2, include_bias=False)
X_poly = poly_features.fit_transform(X)
print(X_poly[0])
print(X[0])
# X_poly contains original feature of X plus feature squared
# now fit linear regression to extended training data
lin_reg = LinearRegression()
lin_reg.fit(X_poly, y)
print(lin_reg.intercept_, lin_reg.coef_)
y_predict = lin_reg.predict(X_poly)
plt.scatter(X, y_predict, color='red')
plt.scatter(X, y)
# warning PolynomialFeatures(degree=d) transforms an array containing n features into (n+d)!/n!d!
# Ridge Regression / L2 regularization
# objective = RSS + a * (sum of square of coefficients)
#
# a = 0 => linear regression
# a = inf => coeffs will be zero, result will be a flat line going thru mean
# 0 < a < inf => coeffs in (0, 1)
#
# regularization term: a Sum(1, N, theta^2) added to cost function, only during training
# J(theta) = MSE(theta) + alpha/2 Sum(1, N, theta^2)
# e.g. classifier trained using log loss is evaluted using F1
# important to scale the data (StandardScaler) before performing ridge regression
from sklearn.linear_model import Ridge
ridge_reg = Ridge(alpha=1, solver='cholesky')
ridge_reg.fit(X, y)
ridge_reg.predict([[1.5]])
sgd_reg = SGDRegressor(penalty='l2')
sgd_reg.fit(X, y.ravel())
sgd_reg.predict([[1.5]])
# Lasso Regression
# Least absolute shrinkage and selection operator
# uses l1 norm of weight vector instead of half the square of the l2 norm
#
# J(theta) = MSE(theta) + alpha * sum(1, n, |theta_i|)
#
# Tends to completely eliminate weights of the least important features (set to zero) (outputs a sparse model)
from sklearn.linear_model import Lasso
lasso_reg = Lasso(alpha=0.1)
lasso_reg.fit(X, y)
lasso_reg.predict([[1.5]])
# Elastic Net
#
# mix of ridge and lasso controlled by r.
# if you suspect only a few features matter, prefer Lasso or Elastic Net.
# Lasso may behave erractically when features > train instances or when feature are correlated
from sklearn.linear_model import ElasticNet
elastic_net = ElasticNet(alpha=0.1, l1_ratio=0.5)
elastic_net.fit(X, y)
elastic_net.predict([[1.5]])
| modules/linear_regression/Linear Regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import matplotlib.pyplot as plt
import pandas as pd
import os
import sys
sys.path.insert(0,'../satori')
from utils import annotate_motifs
# -
# ## Filter redundancy per motif: Human dataset
experiment_path = '../results/Human_Promoter_Analysis_euclidean_v8'
motif_dir = os.path.join(experiment_path,'Motif_Analysis')
motif_dir_neg = os.path.join(experiment_path,'Motif_Analysis_Negative')
df_pos = annotate_motifs(annotate_arg='default', motif_dir=motif_dir, store=False)
df_neg = annotate_motifs(annotate_arg='default', motif_dir=motif_dir_neg, store=False)
df_pos = pd.DataFrame(df_pos)
df_neg = pd.DataFrame(df_neg)
df_pos = df_pos[~df_pos[10].isna()]
df_neg = df_neg[~df_neg[10].isna()]
df_pos[10] = df_pos[10].apply(lambda x: x.split(','))
df_neg[10] = df_neg[10].apply(lambda x: x.split(','))
df_pos = df_pos.explode(10)
df_neg = df_neg.explode(10)
df_pos.shape, df_neg.shape
df_pos_redundancy = df_pos.groupby(10)[0].nunique()
df_pos_redundancy.argmax()
df_pos_redundancy.mean(),df_pos_redundancy.median()
# ## Filter reduandancy per motif: Arabidopsis dataset
experiment_path = '../results/Arabidopsis_GenomeWide_Analysis_euclidean_v8'
motif_dir = os.path.join(experiment_path,'Motif_Analysis')
motif_dir_neg = os.path.join(experiment_path,'Motif_Analysis_Negative')
df_pos = annotate_motifs(annotate_arg='default', motif_dir=motif_dir, store=False)
df_neg = annotate_motifs(annotate_arg='default', motif_dir=motif_dir_neg, store=False)
df_pos = pd.DataFrame(df_pos)
df_neg = pd.DataFrame(df_neg)
df_pos_redundancy = df_pos.groupby(1)[0].nunique()
df_pos_redundancy.mean(), df_pos_redundancy.median()
| analysis/Redundancy_Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: dev
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Dependencies
# Update sklearn to prevent version mismatches
# !pip install sklearn --upgrade
# install joblib. This will be used to save your model.
# Restart your kernel after installing
# !pip install joblib
import joblib
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import MinMaxScaler
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import classification_report
from sklearn.svm import SVC
# # Preprocess the raw data
# ### Read the CSV
df = pd.read_csv("Resources/exoplanet_data.csv")
# ### Perform Basic Data Cleaning
# Drop the null columns where all values are null
df = df.dropna(axis='columns', how='all')
# Drop the null rows
df = df.dropna()
df.head()
df.columns
# ### Select significant features (columns)
# Perform feature selection based upon physical characteristics of the exoplanet candidates. These will also be used as X values.<br>I will mainly select features related to the appearance and physical characteristics. They would be checked off in the following lists. Also, if there is a feature that might result from a linear combination of other features, then that would be unchecked off.
# KOI = Kepler Objects of Interest<br> It is a number used to identify and track a Kepler Object of Interest. A KOI is a target identified by the Kepler Project that displays at least one transit-like sequence within Kepler time-series photometry that appears to be of astrophysical origin and initially consistent with a planetary transit hypothesis.
# #### Project Disposition Columns
# Flags designate the most probable physical explanation of the KOI.
# - [x] koi_fpflag_nt: Not Transit-Like Flag - KOI whose light curve is not consistent with that of a transiting planet.
# - [x] koi_fpflag_ss: Stellar Eclipse Flag - A KOI that is observed to have a significant secondary event, transit shape, or out-of-eclipse variability.
# - [x] koi_fpflag_co: Centroid Offset Flag - The source of the signal is from a nearby star.
# - [x] koi_fpflag_ec: Ephemeris Match Indicates Contamination Flag - The KOI shares the same period and epoch as another object.
# #### Transit Properties
# Transit parameters delivered by the Kepler Project are typically best-fit parameters produced by a Mandel-Agol (2002) fit to a multi-quarter Kepler light curve, assuming a linear orbital ephemeris. Some of the parameters listed below are fit directly, other are derived from the best-fit parameters.
# - [x] koi_period: Orbital Period (days) - The interval between consecutive planetary transits.
# - [x] koi_period_err1: Orbital Period (days) - Uncertainties Column (positive +)
# - [ ] koi_period_err2: Orbital Period (days) - Uncertainties Column (negative -)
# - [x] koi_time0bk: Transit Epoch - The time corresponding to the center of the first detected transit in Barycentric Julian Day (BJD) minus a constant offset of 2,454,833.0 days.
# - [x] koi_time0bk_err1: Transit Epoch - Uncertainties Column (positive +)
# - [ ] koi_time0bk_err2: Transit Epoch - Uncertainties Column (negative -)
# - [x] koi_impact: Impact Parameter - The sky-projected distance between the center of the stellar disc and the center of the planet disc at conjunction, normalized by the stellar radius.
# - [x] koi_impact_err1: Impact Parameter - Uncertainties Column (positive +)
# - [ ] koi_impact_err2: Impact Parameter - Uncertainties Column (negative -)
# - [x] koi_duration: Transit Duration (hours) - The duration of the observed transits.
# - [x] koi_duration_err1: Transit Duration (hours) - Uncertainties Column (positive +)
# - [ ] koi_duration_err2: Transit Duration (hours) - Uncertainties Column (negative -)
# - [x] koi_depth: Transit Depth (parts per million) - The fraction of stellar flux lost at the minimum of the planetary transit.
# - [x] koi_depth_err1: Transit Depth (parts per million) - Uncertainties Column (positive +)
# - [ ] koi_depth_err2: Transit Depth (parts per million) - Uncertainties Column (negative -)
# - [x] koi_prad: Planetary Radius (Earth radii) - The radius of the planet. Planetary radius is the product of the planet star radius ratio and the stellar radius.
# - [x] koi_prad_err1: Planetary Radius (Earth radii) - Uncertainties Column (positive +)
# - [x] koi_prad_err2: Planetary Radius (Earth radii) - Uncertainties Column (negative -)
# - [x] koi_teq: Equilibrium Temperature (Kelvin) - Approximation for the temperature of the planet.
# - [x] koi_insol: Insolation Flux [Earth flux]
# - [x] koi_insol_err1: Insolation Flux [Earth flux] - Uncertainties Column (positive +)
# - [x] koi_insol_err2: Insolation Flux [Earth flux] - Uncertainties Column (negative -)
# #### Threshold-Crossing Event (TCE) Information
# The Transiting Planet Search (TPS) module of the Kepler data analysis pipeline performs a detection test for planet transits in the multi-quarter, gap-filled flux time series. The TPS module detrends each quarterly PDC light curve to remove edge effects around data gaps and then combines the data segments together, filling gaps with interpolated data so as to condition the flux time series for a matched filter.
# - [x] koi_model_snr: Transit Signal-to-Noise - Transit depth normalized by the mean uncertainty in the flux during the transits.
# - [x] koi_tce_plnt_num: TCE Planet Number - TCE Planet Number federated to the KOI.
# #### Stellar Parameters
# Stellar effective temperature, surface gravity, metallicity, radius, mass, and age should comprise a consistent set.
# - [x] koi_steff: Stellar Effective Temperature (Kelvin) - The photospheric temperature of the star.
# - [x] koi_steff_err1: Stellar Effective Temperature (Kelvin) - Uncertainties Column (positive +)
# - [x] koi_steff_err2: Stellar Effective Temperature (Kelvin) - Uncertainties Column (negative -)
# - [x] koi_slogg: Stellar Surface Gravity - The base-10 logarithm of the acceleration due to gravity at the surface of the star.
# - [x] koi_slogg_err1: Stellar Surface Gravity - Uncertainties Column (positive +)
# - [x] koi_slogg_err2: Stellar Surface Gravity - Uncertainties Column (negative -)
# - [x] koi_srad: Stellar Radius (solar radii) - The photospheric radius of the star.
# - [x] koi_srad_err1: Stellar Radius (solar radii) - Uncertainties Column (positive +)
# - [x] koi_srad_err2: Stellar Radius (solar radii) - Uncertainties Column (negative -)
# #### Kepler Input Catalog (KIC) Parameters
# - [x] ra: RA (deg) - KIC Right Ascension of the planetary system in decimal degrees
# - [x] dec: Dec (deg) - KIC Declination in decimal degrees
# - [x] koi_kepmag: Kepler-band (mag) - Kepler-band (mag), it is a magnitude computed according to a hierarchical scheme and depends on what pre-existing catalog source is available.
# Selected features
feature_names = ['koi_fpflag_nt',
'koi_fpflag_ss',
'koi_fpflag_co',
'koi_fpflag_ec',
'koi_period',
'koi_period_err1',
'koi_time0bk',
'koi_time0bk_err1',
'koi_impact',
'koi_impact_err1',
'koi_duration',
'koi_duration_err1',
'koi_depth',
'koi_depth_err1',
'koi_prad',
'koi_prad_err1',
'koi_prad_err2',
'koi_teq',
'koi_insol',
'koi_insol_err1',
'koi_insol_err2',
'koi_model_snr',
'koi_tce_plnt_num',
'koi_steff',
'koi_steff_err1',
'koi_steff_err2',
'koi_slogg',
'koi_slogg_err1',
'koi_slogg_err2',
'koi_srad',
'koi_srad_err1',
'koi_srad_err2',
'ra',
'dec',
'koi_kepmag'
]
X = df[feature_names]
X.head()
# Use `koi_disposition` for the y values
y = df["koi_disposition"]
y.head()
# ### Spliting the data into training and testing data.
# split data into training and testing
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=42)
X_train.head()
# ### Scaling the data
#
# Scaling the data using the MinMaxScaler
# Scale your data
X_scaler = MinMaxScaler().fit(X_train)
X_train_scaled = X_scaler.transform(X_train)
X_test_scaled = X_scaler.transform(X_test)
X.shape
y.shape
# # Original Model
# ### Create the model
# Create the SVC Models: I will compare rbf, poly and linear models.
model_rbf = SVC(kernel='rbf') # Radial Basis Function (RBF)
model_linear = SVC(kernel='linear')
model_poly = SVC(kernel='poly')
# ### Train the Model
# +
# Fit the Models with Trained Data
model_rbf.fit(X_train_scaled, y_train)
model_linear.fit(X_train_scaled, y_train)
model_poly.fit(X_train_scaled, y_train)
# Score the rbf Model with Test Data
model_training_score_rbf = round(model_rbf.score(X_train_scaled, y_train)*100,3)
model_accuracy_rbf = round(model_rbf.score(X_test_scaled, y_test)*100,3)
print(f"RBF Training Data Score: {model_training_score_rbf}%")
print(f"RBF Testing Data Score: {model_accuracy_rbf}%")
# Score the linear Model with Test Data
model_training_score_linear = round(model_linear.score(X_train_scaled, y_train)*100,3)
model_accuracy_linear = round(model_linear.score(X_test_scaled, y_test)*100,3)
print(f"LINEAR Training Data Score: {model_training_score_linear}%")
print(f"LINEAR Testing Data Score: {model_accuracy_linear}%")
# Score the poly Model with Test Data
model_training_score_poly = round(model_linear.score(X_train_scaled, y_train)*100,3)
model_accuracy_poly = round(model_linear.score(X_test_scaled, y_test)*100,3)
print(f"POLY Training Data Score: {model_training_score_poly}%")
print(f"POLY Testing Data Score: {model_accuracy_poly}%")
# -
# According to the score results, it seems that the linear model predicts more reliably.<br>
# I will use the linear kernel then, it looks like the most effective and inexpensive algorithm for the current data
model = model_linear
# # Hyperparameter Tuning
# ### Create the `GridSearchCV` model to find best/tuned parameters
# +
param_grid = {'C': [1, 5, 10, 50],
'gamma': [0.0001, 0.0005, 0.001, 0.005]}
grid_model = GridSearchCV(model, param_grid, verbose=3)
# Train the model with GridSearch
_ = grid_model.fit(X_train_scaled, y_train)
# -
# ### Find Tuned parameters
print(f"Best Parameters: {grid_model.best_params_}")
print(f"Best Score: {round(grid_model.best_score_*100,3)}%")
# # Tuned Model
# ### Create the model with specific parameters
# Tuned model based upon best parameters previously found
tuned_model = SVC(kernel='linear',
C=grid_model.best_params_['C'],
gamma=grid_model.best_params_['gamma'],
random_state=42)
# ### Train the Model
# +
# Fit and score the tuned model
tuned_model.fit(X_train_scaled, y_train)
tuned_training_score = round(tuned_model.score(X_train_scaled, y_train)*100,3)
tuned_accuracy = round(tuned_model.score(X_test_scaled, y_test)*100,3)
print(f"Training Data Score: {tuned_training_score} %")
print(f"Testing Data Score: {tuned_accuracy} %")
# -
# ### Classification report
# +
# Make predictions with the hypertuned model
predictions = tuned_model.predict(X_test_scaled)
# Calculate classification report
print(classification_report(y_test, predictions, target_names=["CANDIDATE","FALSE POSITIVE","CONFIRMED"]))
| svm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Dask
#
#
# <div class="alert-info">
#
# ### Overview
#
# * **teaching:** 20 minutes
# * **exercises:** 0
# * **questions:**
# * How does Dask parallelize computations in Python?
# </div>
# ### Table of contents
# 1. [**Dask primer**](#Dask-primer)
# 1. [**Dask clusters**](#Dask-Clusters)
# 1. [**Dask dataframe**](#Dask-Dataframe)
# 1. [**Dask arrays**](#Dask-Arrays)
# 1. [**Dask delayed**](#Dask-Delayed)
# ## Dask Primer
#
# <img src="http://dask.readthedocs.io/en/latest/_images/dask_horizontal.svg"
# width="30%"
# align=right
# alt="Dask logo">
#
#
# Dask is a flexible parallel computing library for analytic computing. Dask provides dynamic parallel task scheduling and high-level big-data collections like `dask.array` and `dask.dataframe`. More on dask here: https://docs.dask.org/en/latest/
#
# _Note: Pieces of this notebook comes from the following sources:_
#
# - https://github.com/rabernat/research_computing
# - https://github.com/dask/dask-examples
# ## Dask Clusters
#
# Dask needs a collection of computing resources in order to perform parallel computations. Dask Clusters have different names corresponding to different computing environments (for example, [LocalCluster](https://distributed.dask.org/en/latest/local-cluster.html) for your Laptop, [PBSCluster](http://jobqueue.dask.org/) for your HPC, or [Kubernetes Cluster](http://kubernetes.dask.org/) for machines on the Cloud). Each cluster has a certain number of computing resources called 'Workers', that each get allocated CPU and RAM. The dask scheduling system maps jobs to each worker on a cluster for you, so the syntax is mostly the same once you initialize a cluster!
# Let's start simple with a LocalCluster that makes use of all the cores and RAM we have on a single machine
from dask.distributed import Client, LocalCluster
cluster = LocalCluster()
# explicitly connect to the cluster we just created
client = Client(cluster)
client
# ## Dask Dataframe
#
# If you are working with a very large Pandas dataframe, you can consider parallizing computations by turning it into a Dask Dataframe. Dask Dataframes split a dataframe into partitions along an index. They support a large subset of the Pandas API. You can find additional details and examples here https://examples.dask.org/dataframe.html
#
# +
# Although this is small csv file, we'll reuse our same example from before!
# Load csv results from server into a Pandas DataFrame
import dask.dataframe as dd
server = 'https://webservices.volcano.si.edu/geoserver/GVP-VOTW/ows?'
query = 'service=WFS&version=2.0.0&request=GetFeature&typeName=GVP-VOTW:Smithsonian_VOTW_Holocene_Volcanoes&outputFormat=csv'
# blocksize=None means use a single partion
df = dd.read_csv(server+query, blocksize=None)
# -
# We only see the metadata, the actual data are only computed when requested.
df
# We can break up the table into 4 partions to map out to each core:
df = df.repartition(npartitions=4)
df
# Let's say we want to know the minimum last eruption year for all volcanoes
last_eruption_year_min = df.Last_Eruption_Year.min()
last_eruption_year_min
# Instead of getting the actual value we see dd.Scalar, which represents a recipe for actually calculating this value
last_eruption_year_min.visualize()
# To get the value call the 'compute method'
# NOTE: this was slower than using pandas directly,,, for small data you often don't need to use parallel computing!
last_eruption_year_min.compute()
# ## Dask Arrays
#
# A dask array looks and feels a lot like a numpy array.
# However, a dask array doesn't directly hold any data.
# Instead, it symbolically represents the computations needed to generate the data.
# Nothing is actually computed until the actual numerical values are needed.
# This mode of operation is called "lazy"; it allows one to build up complex, large calculations symbolically before turning them over the scheduler for execution.
#
# If we want to create a numpy array of all ones, we do it like this:
import numpy as np
shape = (1000, 4000)
ones_np = np.ones(shape)
ones_np
# This array contains exactly 32 MB of data:
print('%.1f MB' % (ones_np.nbytes / 1e6))
# Now let's create the same array using dask's array interface.
import dask.array as da
ones = da.ones(shape)
ones
# This works, but we didn't tell dask how to split up the array, so it is not optimized for distributed computation.
#
# A crucal difference with dask is that we must specify the `chunks` argument. "Chunks" describes how the array is split up over many sub-arrays.
#
# 
# _source: [Dask Array Documentation](http://dask.pydata.org/en/latest/array-overview.html)_
#
# There are [several ways to specify chunks](http://dask.pydata.org/en/latest/array-creation.html#chunks).
# In this lecture, we will use a block shape.
chunk_shape = (1000, 1000)
ones = da.ones(shape, chunks=chunk_shape)
ones
# Notice that we just see a symbolic represetnation of the array, including its shape, dtype, and chunksize.
# No data has been generated yet.
# When we call `.compute()` on a dask array, the computation is trigger and the dask array becomes a numpy array.
ones.compute()
# In order to understand what happened when we called `.compute()`, we can visualize the dask _graph_, the symbolic operations that make up the array
ones.visualize()
# Our array has four chunks. To generate it, dask calls `np.ones` four times and then concatenates this together into one array.
#
# Rather than immediately loading a dask array (which puts all the data into RAM), it is more common to reduce the data somehow. For example:
sum_of_ones = ones.sum()
sum_of_ones.visualize()
# Here we see dask's strategy for finding the sum. This simple example illustrates the beauty of dask: it automatically designs an algorithm appropriate for custom operations with big data.
#
# If we make our operation more complex, the graph gets more complex.
fancy_calculation = (ones * ones[::-1, ::-1]).mean()
fancy_calculation.visualize()
# ### A Bigger Calculation
#
# The examples above were toy examples; the data (32 MB) is nowhere nearly big enough to warrant the use of dask.
#
# We can make it a lot bigger!
bigshape = (200000, 4000)
big_ones = da.ones(bigshape, chunks=chunk_shape)
big_ones
print('%.1f MB' % (big_ones.nbytes / 1e6))
# This dataset is 6.4 GB, rather than 32 MB! This is probably close to or greater than the amount of available RAM than you have in your computer. Nevertheless, dask has no problem working on it.
#
# _Do not try to `.visualize()` this array!_
#
# When doing a big calculation, dask also has some tools to help us understand what is happening under the hood. Let's watch the dashboard again as we do a bigger computation.
# +
big_calc = (big_ones * big_ones[::-1, ::-1]).mean()
result = big_calc.compute()
result
# -
# ### Reduction
#
# All the usual numpy methods work on dask arrays.
# You can also apply numpy function directly to a dask array, and it will stay lazy.
big_ones_reduce = (np.cos(big_ones)**2).mean(axis=1)
big_ones_reduce
# Plotting also triggers computation, since we need the actual values
from matplotlib import pyplot as plt
# %matplotlib inline
plt.rcParams['figure.figsize'] = (12,8)
plt.plot(big_ones_reduce)
# ## Dask Delayed
#
# Dask.delayed is a simple and powerful way to parallelize existing code. It allows users to delay function calls into a task graph with dependencies. Dask.delayed doesn't provide any fancy parallel algorithms like Dask.dataframe, but it does give the user complete control over what they want to build.
#
# Systems like Dask.dataframe are built with Dask.delayed. If you have a problem that is paralellizable, but isn't as simple as just a big array or a big dataframe, then dask.delayed may be the right choice for you.
#
# ## Create simple functions
#
# These functions do simple operations like add two numbers together, but they sleep for a random amount of time to simulate real work.
# +
import time
def inc(x):
time.sleep(0.1)
return x + 1
def dec(x):
time.sleep(0.1)
return x - 1
def add(x, y):
time.sleep(0.2)
return x + y
# -
# We can run them like normal Python functions below
# %%time
x = inc(1)
y = dec(2)
z = add(x, y)
z
# These ran one after the other, in sequence. Note though that the first two lines `inc(1)` and `dec(2)` don't depend on each other, we *could* have called them in parallel had we been clever.
#
# ## Annotate functions with Dask Delayed to make them lazy
#
# We can call `dask.delayed` on our funtions to make them lazy. Rather than compute their results immediately, they record what we want to compute as a task into a graph that we'll run later on parallel hardware.
import dask
inc = dask.delayed(inc)
dec = dask.delayed(dec)
add = dask.delayed(add)
# Calling these lazy functions is now almost free. We're just constructing a graph
# %%time
x = inc(1)
y = dec(2)
z = add(x, y)
z
# ## Visualize computation
z.visualize(rankdir='LR')
# ## Run in parallel
#
# Call `.compute()` when you want your result as a normal Python object
#
# If you started `Client()` above then you may want to watch the status page during computation.
# %%time
z.compute()
# ## Parallelize Normal Python code
#
# Now we use Dask in normal for-loopy Python code. This generates graphs instead of doing computations directly, but still looks like the code we had before. Dask is a convenient way to add parallelism to existing workflows.
# +
# %%time
zs = []
for i in range(256):
x = inc(i)
y = dec(x)
z = add(x, y)
zs.append(z)
zs = dask.persist(*zs) # trigger computation in the background
| dask.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/", "height": 222} colab_type="code" executionInfo={"elapsed": 98926, "status": "ok", "timestamp": 1549193114654, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-xVEOvhAYCwI/AAAAAAAAAAI/AAAAAAAAAAc/m8NWBboPeYY/s64/photo.jpg", "userId": "17018506043059870013"}, "user_tz": -330} id="EwDVm3m-A2zc" outputId="1551a942-b314-4faa-b2f8-361973c242b0"
import os
print(os.getcwd())
# + colab={} colab_type="code" id="9w15dc6-BJhJ"
os.chdir('Dataset')
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 824, "status": "ok", "timestamp": 1549195084438, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-xVEOvhAYCwI/AAAAAAAAAAI/AAAAAAAAAAc/m8NWBboPeYY/s64/photo.jpg", "userId": "17018506043059870013"}, "user_tz": -330} id="jOHbh_IQBhCZ" outputId="9f31aa31-2cbb-43aa-ebe4-8f7c7bff6024"
print(os.getcwd())
# + colab={"base_uri": "https://localhost:8080/", "height": 162} colab_type="code" executionInfo={"elapsed": 1998, "status": "error", "timestamp": 1549195074943, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-xVEOvhAYCwI/AAAAAAAAAAI/AAAAAAAAAAc/m8NWBboPeYY/s64/photo.jpg", "userId": "17018506043059870013"}, "user_tz": -330} id="52jddDUmBn9C" outputId="c28e3ab7-1631-4c2a-983b-09222705ea5e"
# !ls
# + [markdown] colab={"base_uri": "https://localhost:8080/", "height": 162} colab_type="code" executionInfo={"elapsed": 1998, "status": "error", "timestamp": 1549195074943, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-xVEOvhAYCwI/AAAAAAAAAAI/AAAAAAAAAAc/m8NWBboPeYY/s64/photo.jpg", "userId": "17018506043059870013"}, "user_tz": -330} id="52jddDUmBn9C" outputId="c28e3ab7-1631-4c2a-983b-09222705ea5e"
# # Importing Libraries
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 861, "status": "ok", "timestamp": 1549195088664, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-xVEOvhAYCwI/AAAAAAAAAAI/AAAAAAAAAAc/m8NWBboPeYY/s64/photo.jpg", "userId": "17018506043059870013"}, "user_tz": -330} id="PpiCsU-aCb0b" outputId="63b98d60-50e0-414f-bb99-81021a711c32"
from __future__ import print_function
import keras
from keras.models import Sequential
from keras.layers import Dense, Dropout, Flatten, Activation
from keras.layers.normalization import BatchNormalization
from keras.layers import Conv1D, Conv2D, MaxPooling2D, MaxPooling1D
from keras import backend as K
import h5py
import numpy as np
import scipy.io as sio
import matplotlib.pyplot as plt
# + [markdown] colab={"base_uri": "https://localhost:8080/", "height": 101} colab_type="code" executionInfo={"elapsed": 2931, "status": "ok", "timestamp": 1549195092554, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-xVEOvhAYCwI/AAAAAAAAAAI/AAAAAAAAAAc/m8NWBboPeYY/s64/photo.jpg", "userId": "17018506043059870013"}, "user_tz": -330} id="84WiYC0FCeHJ" outputId="6f11a35b-3371-4ba5-fa1e-96451acc43c5"
# # Defining Constants
# + colab={"base_uri": "https://localhost:8080/", "height": 101} colab_type="code" executionInfo={"elapsed": 2931, "status": "ok", "timestamp": 1549195092554, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-xVEOvhAYCwI/AAAAAAAAAAI/AAAAAAAAAAc/m8NWBboPeYY/s64/photo.jpg", "userId": "17018506043059870013"}, "user_tz": -330} id="84WiYC0FCeHJ" outputId="6f11a35b-3371-4ba5-fa1e-96451acc43c5"
batch_size = 32
num_classes = 13
epochs = 225
# + [markdown] colab={"base_uri": "https://localhost:8080/", "height": 101} colab_type="code" executionInfo={"elapsed": 2931, "status": "ok", "timestamp": 1549195092554, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-xVEOvhAYCwI/AAAAAAAAAAI/AAAAAAAAAAc/m8NWBboPeYY/s64/photo.jpg", "userId": "17018506043059870013"}, "user_tz": -330} id="84WiYC0FCeHJ" outputId="6f11a35b-3371-4ba5-fa1e-96451acc43c5"
# # Loadin MATLAB File
# + colab={} colab_type="code" id="tbWa8HFBCfOD"
trainset = {}
trainset = sio.loadmat('chess_train_final.mat')
print("trainset size X:", trainset['X'].shape)
print("trainset size Y:", trainset['Y'].shape)
# + colab={} colab_type="code" id="jnjgvWdaCxwk"
trainset['X'] = np.rollaxis(trainset['X'], axis = 3)
# + colab={"base_uri": "https://localhost:8080/", "height": 50} colab_type="code" executionInfo={"elapsed": 2424, "status": "ok", "timestamp": 1549195099888, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-xVEOvhAYCwI/AAAAAAAAAAI/AAAAAAAAAAc/m8NWBboPeYY/s64/photo.jpg", "userId": "17018506043059870013"}, "user_tz": -330} id="BnbtkZPvDEUO" outputId="3497ff1a-1a3e-4d1c-db71-456eb7fb93f0"
print('Trainset size X:',trainset['X'].shape)
print('Trainset size Y:',trainset['Y'].shape)
# + colab={} colab_type="code" id="VVr0j79oDO63"
label = trainset['Y']
# + colab={"base_uri": "https://localhost:8080/", "height": 50} colab_type="code" executionInfo={"elapsed": 882, "status": "ok", "timestamp": 1549195101666, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-xVEOvhAYCwI/AAAAAAAAAAI/AAAAAAAAAAc/m8NWBboPeYY/s64/photo.jpg", "userId": "17018506043059870013"}, "user_tz": -330} id="0ej-9Wy2EF-X" outputId="43d51655-9527-42fc-d1db-262755387ffa"
# convert class vectors to binary class matrices
trainset['Y'] = keras.utils.to_categorical(trainset['Y'], num_classes)
# + colab={} colab_type="code" id="Nd3lje75DJnD"
print(trainset['Y'].shape)
# + colab={} colab_type="code" id="2E0mPHTSG8YI"
print(trainset['Y'].shape)
print(trainset['Y'][2606,:])
#2606 are the number of images starting from 0 to 2606 (A total of 2607 Number of Images if counted from 1)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 1130, "status": "ok", "timestamp": 1549195103988, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-xVEOvhAYCwI/AAAAAAAAAAI/AAAAAAAAAAc/m8NWBboPeYY/s64/photo.jpg", "userId": "17018506043059870013"}, "user_tz": -330} id="IixP2uReHCEI" outputId="39410e01-f5cc-4ed7-b60a-257f0a460551"
print(trainset['X'].shape[0], 'train samples')
# + [markdown] colab={} colab_type="code" id="QWVcwIrOHGH7"
# # Setting Model Input Image Dimensions
# + colab={} colab_type="code" id="QWVcwIrOHGH7"
# input image dimensions
img_rows, img_cols = 150, 150
input_shape = (img_rows, img_cols, 3)
# + [markdown] colab={"base_uri": "https://localhost:8080/", "height": 50} colab_type="code" executionInfo={"elapsed": 994, "status": "ok", "timestamp": 1549195104388, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-xVEOvhAYCwI/AAAAAAAAAAI/AAAAAAAAAAc/m8NWBboPeYY/s64/photo.jpg", "userId": "17018506043059870013"}, "user_tz": -330} id="YEUKrgI3HkHh" outputId="1245ed08-7dda-4719-87d6-e546f792ba82"
# # Testing Random Image
# + colab={"base_uri": "https://localhost:8080/", "height": 50} colab_type="code" executionInfo={"elapsed": 994, "status": "ok", "timestamp": 1549195104388, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-xVEOvhAYCwI/AAAAAAAAAAI/AAAAAAAAAAc/m8NWBboPeYY/s64/photo.jpg", "userId": "17018506043059870013"}, "user_tz": -330} id="YEUKrgI3HkHh" outputId="1245ed08-7dda-4719-87d6-e546f792ba82"
# Trainset image
#index=2606
index = np.random.randint(2606)
M1 = trainset['X'][index,:,:,:]
plt.imshow(M1)
plt.show()
y = label[index,:]
if y==0:
classes='E'
elif y==1:
classes='B'
elif y==2:
classes='K'
elif y==3:
classes='N'
elif y==4:
classes='P'
elif y==5:
classes='Q'
elif y==6:
classes='R'
elif y==7:
classes='b'
elif y==8:
classes='k'
elif y==9:
classes='n'
elif y==10:
classes='p'
elif y==11:
classes='q'
else:
classes='r'
print('Label:',classes)
# + [markdown] colab={"base_uri": "https://localhost:8080/", "height": 50} colab_type="code" executionInfo={"elapsed": 994, "status": "ok", "timestamp": 1549195104388, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-xVEOvhAYCwI/AAAAAAAAAAI/AAAAAAAAAAc/m8NWBboPeYY/s64/photo.jpg", "userId": "17018506043059870013"}, "user_tz": -330} id="YEUKrgI3HkHh" outputId="1245ed08-7dda-4719-87d6-e546f792ba82"
# # Model Layers
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 839, "status": "ok", "timestamp": 1549195104390, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-xVEOvhAYCwI/AAAAAAAAAAI/AAAAAAAAAAc/m8NWBboPeYY/s64/photo.jpg", "userId": "17018506043059870013"}, "user_tz": -330} id="2m6EB1waITkj" outputId="57bbe5fb-7c7a-4129-82b2-8a79b6b5ca3f"
model = Sequential()
model.add(BatchNormalization(input_shape = input_shape))
model.add(Conv2D(32, (3, 3), padding = 'valid'))
model.add(Activation('relu'))
model.add(Conv2D(32, (3, 3), padding = 'valid'))
model.add(Activation('relu'))
model.add(MaxPooling2D(pool_size=(2, 2), strides = None, padding = 'valid'))
model.add(Dropout(0.25))
model.add(Conv2D(64, (3, 3), padding = 'valid'))
model.add(Activation('relu'))
model.add(Conv2D(64, (3, 3), padding = 'valid'))
model.add(Activation('relu'))
model.add(MaxPooling2D(pool_size=(2, 2), strides = None, padding = 'valid'))
model.add(Dropout(0.25))
model.add(Conv2D(128, (3, 3), padding = 'valid'))
model.add(Activation('relu'))
model.add(Conv2D(128, (3, 3), padding = 'valid'))
model.add(Activation('relu'))
model.add(MaxPooling2D(pool_size=(2, 2), strides = None, padding = 'valid'))
model.add(Dropout(0.25))
model.add(Flatten())
model.add(Dense(128))
model.add(Activation('relu'))
model.add(Dropout(0.5))
model.add(Dense(num_classes))
model.add(Activation('softmax'))
# initiate RMSprop optimizer
opt = keras.optimizers.Adadelta(lr=1.0, rho=0.95, epsilon=None, decay=0.0)
# Let's train the model using RMSprop
model.compile(loss='categorical_crossentropy',
optimizer=opt,
metrics=['accuracy'])
model.summary()
# + [markdown] colab={} colab_type="code" id="orhtce1TIOzU"
# # Start Training the Model
# + colab={} colab_type="code" id="orhtce1TIOzU"
from keras.preprocessing.image import ImageDataGenerator
datagen = ImageDataGenerator(
rotation_range=40,
width_shift_range=0.2,
height_shift_range=0.2,
shear_range=0.2,
zoom_range=0.2,
horizontal_flip=True,
fill_mode='nearest')
datagen.fit(trainset['X'])
model.fit_generator(datagen.flow(trainset['X'], trainset['Y'], batch_size=32),
steps_per_epoch=len(trainset['X']) / 32, epochs=epochs)
# + [markdown] colab={"base_uri": "https://localhost:8080/", "height": 366} colab_type="code" executionInfo={"elapsed": 1018, "status": "ok", "timestamp": 1549195233216, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-xVEOvhAYCwI/AAAAAAAAAAI/AAAAAAAAAAc/m8NWBboPeYY/s64/photo.jpg", "userId": "17018506043059870013"}, "user_tz": -330} id="s0IY_XEUHlp2" outputId="7c4e7e57-85d9-4ac6-f493-45dc7dfced86"
# # Saving Trained Model to Directory
# + colab={"base_uri": "https://localhost:8080/", "height": 366} colab_type="code" executionInfo={"elapsed": 1018, "status": "ok", "timestamp": 1549195233216, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-xVEOvhAYCwI/AAAAAAAAAAI/AAAAAAAAAAc/m8NWBboPeYY/s64/photo.jpg", "userId": "17018506043059870013"}, "user_tz": -330} id="s0IY_XEUHlp2" outputId="7c4e7e57-85d9-4ac6-f493-45dc7dfced86"
save_dir = os.path.join(os.getcwd(), 'saved_models')
model_name = 'trained_model_final_bs32_225_dg_6.h5'
if not os.path.isdir(save_dir):
os.makedirs(save_dir)
model_path = os.path.join(save_dir, model_name)
model.save(model_path)
print('Saved trained model at %s ' % model_path)
| Model/Model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:py38] *
# language: python
# name: conda-env-py38-py
# ---
# # PMEL/FOCI ERDDAP for Moorings Example
#
# ## EcoFOCI Background
#
# EcoFOCI maintains an internal server (or two) that mooring/ctd and other data is served up on. downdraft/thundersnow is the test server (S.Bell's office) and Akutan is considered the production server (virtual linux system)
#
# ## ERDDAP References
#
# more info can be found on the erddapy webpage - https://ioos.github.io/erddapy/quick_intro.html#
#standard imports for this operation
from erddapy import ERDDAP
import pandas as pd
import numpy as np
# erddap has two types of data basically... gridded (foci data uses this for 1hr datasets and adcp data) and tabular. Tabular imports to pandas very nicely as csv files and gridded imports to xarray (via .nc files) nicely. Of course, more meta information is retained when importing a netcdf file than a csv file if this is important.
# +
server_url = 'http://akutan.pmel.noaa.gov:8080/erddap'
did = 'datasets_Mooring_17bsm2a_final'
d = ERDDAP(server=server_url,
protocol='tabledap',
response='csv',
)
#usally two data bases exist for each deployment, a preliminary and a final
#a gridded may also exist (build from the final tabular dataset) and an ADCP dataset may exist
d.dataset_id=did
#define the variables you want to get
d.variables = [
'timeseries_id',
'Temperature',
'Salinity',
'Chlorophyll',
'Turbidity',
'latitude',
'longitude',
'depth',
"time",
'Oxy_Conc',
'Oxy_Sat'
]
# useful if only wanting a small time portion
d.constraints = {
'time>=': '2017-01-01T00:00:00Z',
'time<=': '2020-10-10T00:00:00Z',
}
#data download below will fail unitl the variables are named apporpiatly
# -
# the following will get variable names by attribute "standard_name" where standard_name is not empty (useful as variables may have different names among datasets, but standard_names are pulled from the CF standard naming convention where available)
d.get_var_by_attr(dataset_id=did,standard_name='sea_water_temperature')
# +
variables = d.get_var_by_attr(
dataset_id=did,
standard_name=lambda v: (v is not None)
)
variables
# -
def show_iframe(src):
"""Helper function to show HTML returns."""
from IPython.display import HTML
iframe = f'<iframe src="{src}" width="100%" height="950"></iframe>'
return HTML(iframe)
# +
#watch out for deg E conversions - either >180 or <0
kw = {
'standard_name': 'seawater_temperature',
'min_lon': 180.0,
'max_lon': 220.0,
'min_lat': 50.0,
'max_lat': 85.0,
}
search_url = d.get_search_url(response='html', **kw)
show_iframe(search_url)
# -
# erddapy really is just a url builder, the following url can be passed into just about any RESTful service
# +
d.variables = variables
download_url = d.get_download_url()
from urllib.parse import urlencode
print(download_url) #not working?
# +
#download data
df = d.to_pandas(
index_col='time (UTC)',
parse_dates=True,
skiprows=(1,) # units information can be dropped.
)
df.sort_index(inplace=True)
df.columns = [x[1].split()[0] for x in enumerate(df.columns)]
df.head()
# +
import matplotlib as mpl
import matplotlib.pyplot as plt
import matplotlib.dates as mdates
from matplotlib.dates import YearLocator, WeekdayLocator, MonthLocator, DayLocator, HourLocator, DateFormatter
import matplotlib.ticker as ticker
import cmocean
### specify primary bulk figure parameters
fontsize = 10
labelsize = 10
#plotstyle = 'seaborn'
max_xticks = 10
plt.style.use('seaborn-ticks')
mpl.rcParams['svg.fonttype'] = 'none'
mpl.rcParams['ps.fonttype'] = 42 #truetype/type2 fonts instead of type3
mpl.rcParams['pdf.fonttype'] = 42 #truetype/type2 fonts instead of type3
mpl.rcParams['axes.grid'] = False
mpl.rcParams['axes.edgecolor'] = 'black'
mpl.rcParams['axes.linewidth'] = 1.5
mpl.rcParams['axes.labelcolor'] = 'black'
mpl.rcParams['grid.linestyle'] = '--'
mpl.rcParams['grid.linestyle'] = '--'
mpl.rcParams['xtick.major.size'] = 4
mpl.rcParams['xtick.minor.size'] = 2
mpl.rcParams['xtick.major.width'] = 2
mpl.rcParams['xtick.minor.width'] = 0.5
mpl.rcParams['ytick.major.size'] = 4
mpl.rcParams['ytick.minor.size'] = 2
mpl.rcParams['ytick.major.width'] = 2
mpl.rcParams['ytick.minor.width'] = 0.5
mpl.rcParams['ytick.direction'] = 'out'
mpl.rcParams['xtick.direction'] = 'out'
mpl.rcParams['ytick.color'] = 'black'
mpl.rcParams['xtick.color'] = 'black'
# -
fig, ax = plt.subplots(figsize=(11, 8))
for name,group in df.groupby(df.depth):
ax.plot(group.index,group.temperature)
# +
#not gridded
# %time
fig, ax = plt.subplots(figsize=(11, 8))
for name,group in df.groupby(df.depth):
cs = ax.scatter(group.index,group.depth,c=group.temperature,
vmin=-2, vmax=15, marker='o', edgecolor='none',
cmap=cmocean.cm.thermal)
ax.invert_yaxis()
cbar = fig.colorbar(cs, orientation='vertical', extend='both')
cbar.ax.set_ylabel('Temperature ($^\circ$C)')
ax.set_ylabel('Depth (m)')
# +
#the above is slow... is nc the way to go? doesn't seem so
d.response = 'nc'
download_url = d.get_download_url()
ds = d.to_xarray(decode_times=True)
ds
# -
# one long timeseries - need to grid it to make it more useful though... so nc not really - not from tabular anyways...
#
# so from the tabular data, I create a gridded product (easy to modify) set at 1hr intervals which are then hosted as grids on erddap with appropriate file modified for the name
# +
dataset = '1hr_gridded_datasets_Mooring_17bsm2a_final'
e = ERDDAP(server=server_url)
e.constraints = None
e.protocol = 'griddap'
opendap_url = e.get_download_url(
dataset_id=dataset,
response='opendap',
)
print(opendap_url)
# +
from netCDF4 import Dataset
with Dataset(opendap_url) as nc:
print(nc.summary)
# -
e.dataset_id=dataset
e.response = 'nc'
ds = e.to_xarray(decode_times=True)
ds
# much more useful as it is geophysically gridded along time/depth
for i in range(0,len(ds.temperature)):
ds.temperature[i].plot()
fig, (ax1) = plt.subplots(nrows=1, sharex=True, figsize=(8.5,5.5))
ds.plot.scatter(x='time',y='depth',hue='temperature', cmap=cmocean.cm.thermal, ax=ax1)
ax1.invert_yaxis()
# + tags=[]
#or better yet
fig, (ax1) = plt.subplots(nrows=1, sharex=True, figsize=(8.5,5.5))
ax1.pcolormesh(ds.time,ds.depth,ds.temperature,
cmap=cmocean.cm.thermal)
ax1.invert_yaxis()
xfmt = mdates.DateFormatter('%d-%b\n%y')
ax1.xaxis.set_major_formatter(xfmt)
ax1.xaxis.set_major_locator(MonthLocator())
ax1.xaxis.set_minor_locator(DayLocator())
# -
| EcoFOCI_Moorings/ERDDAP_Automated_Tools/FOCI_ERDDAP_Moorings_Example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# argv:
# - python
# - -m
# - ipykernel_launcher
# - -f
# - '{connection_file}'
# display_name: Python 3
# env: null
# interrupt_mode: signal
# language: python
# metadata: null
# name: python3
# ---
# # Lesson 04: Classification Performance ROCs
#
# - evaluating and comparing trained models is of extreme importance when deciding in favor/against
# + model architectures
# + hyperparameter sets
#
# - evaluating performance or quality of prediction is performed with a myriad of tests, figure-of-merits and even statistical hypothesis testing
# - in the following, the rather popular "Receiver Operating Characteristic" curve (spoken ROC curve)
# - the ROC was invented in WWII by radar engineers when seeking to detect enemy vessels and comparing different devices/techniques
#
# ## preface
# - two main ingredients to ROC:
#
# + TPR = True Positive Rate
# + FPR = False Positive Rate
#
#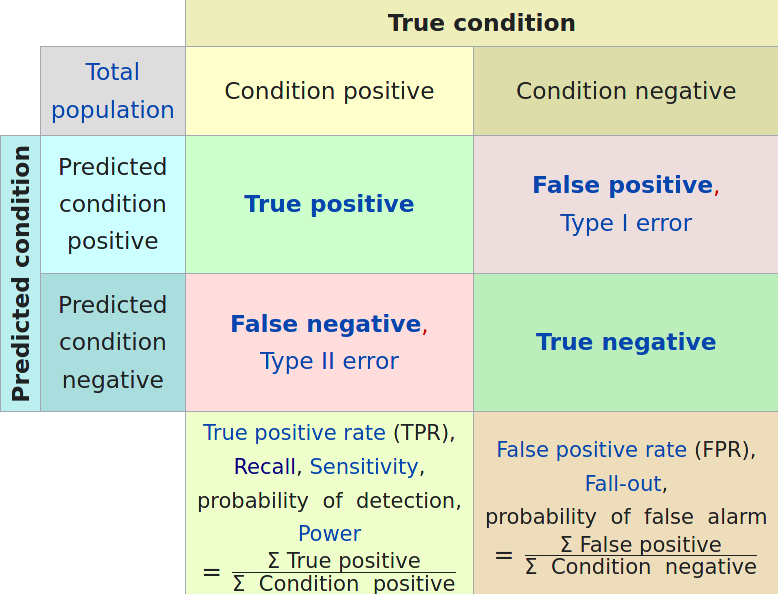 
#
# - $TPR = \frac{TP}{TP+FN}$ also known as `recall`, always within $[0,1]$
# - $FPR = \frac{FP}{FP+TN}$ also known as `fall-out`, always within $[0,1]$
# ## Data
#
# For the following, I will rely (again) on the Palmer penguin dataset obtained from [this repo](https://github.com/allisonhorst/palmerpenguins). To quote the repo:
#
# > Data were collected and made available by [Dr. <NAME>](https://www.uaf.edu/cfos/people/faculty/detail/kristen-gorman.php)
# > and the [Palmer Station, Antarctica LTER](https://pal.lternet.edu/), a member of the [Long Term Ecological Research Network](https://lternet.edu/).
#
import pandas as pd
import numpy as np
df = pd.read_csv("https://raw.githubusercontent.com/allisonhorst/palmerpenguins/master/inst/extdata/penguins.csv")
#let's remove the rows with NaN values
df = df[ df.bill_length_mm.notnull() ]
#convert species column to
df[["species_"]] = df[["species"]].astype("category")
# +
print(df.shape)
print((df.species_.cat.codes < 1).shape)
#create binary column
df["is_adelie"] = (df.species_.cat.codes < 1).astype(np.int8)
print(df.head())
# -
import matplotlib.pyplot as plt
plt.style.use('dark_background')
import seaborn as sns
print(f'seaborn version: {sns.__version__}')
# +
from sklearn.neighbors import KNeighborsClassifier as knn
from sklearn.model_selection import train_test_split
kmeans = knn(n_neighbors=5)
# +
#this time we train the knn algorithm, i.e. an unsupervised method is used in a supervised fashion
#prepare the data
X = np.stack((df.bill_length_mm, df.flipper_length_mm), axis=-1)
y = df.is_adelie
print(X.shape)
print(y.shape)
# +
X_train, X_test, y_train, y_test = train_test_split(X, y,
test_size = .15,
random_state = 20210303)
print(X_train.shape)
print(y_train.shape)
print(X_test.shape)
print(y_test.shape)
# -
kmeans = kmeans.fit(X_train, y_train)
# +
from sklearn.metrics import confusion_matrix, ConfusionMatrixDisplay
y_test_hat = kmeans.predict(X_test)
cm = confusion_matrix( y_test, y_test_hat )
print(cm)
# -
from sklearn.metrics import ConfusionMatrixDisplay
disp = ConfusionMatrixDisplay(confusion_matrix=cm)
disp.plot()
print(int(True))
# # Starting to ROC
#
# - let's take 4 samples of different size from our test set (as if we would conduct 4 experiments)
#
# +
n_experiments = 4
X_test_exp = np.split(X_test[:32,...],n_experiments,axis=0)
y_test_exp = np.split(y_test.values[:32,...],n_experiments,axis=0)
print(X_test_exp[0].shape)
print(y_test_exp[0].shape)
y_test_exp
# +
y_test_hat = kmeans.predict(X_test)
y_test_hat_exp = np.split(y_test_hat[:32,...],n_experiments,axis=0)
# +
#let's compute tpr and fpr for each
from sklearn.metrics import recall_score as tpr
def fpr(y_true, y_pred):
""" compute the false positive rate using the confusion_matrix"""
cm = confusion_matrix(y_true, y_pred)
assert cm.shape == (2,2), f"{y_true.shape, y_pred.shape} => {cm,cm.shape}"
cond_negative = cm[:,1].sum()
value = cm[0,1] / cond_negative
return value
tpr_ = []
fpr_ = []
for i in range(len(y_test_exp)):
tpr_.append(tpr(y_test_exp[i], y_test_hat_exp[i]))
fpr_.append(fpr(y_test_exp[i], y_test_hat_exp[i]))
print(tpr_)
print(fpr_)
# +
f, ax = plt.subplots(1)
ax.plot(fpr_, tpr_, 'ro', markersize=10)
ax.set_xlabel('False Positive Rate')
ax.set_ylabel('True Positive Rate')
ax.set_xlim(0,1)
ax.set_ylim(0,1)
# -
# # But how to get from single entries to a full curve?
#
# - in our case, we can employ the positive class prediction probabilities
# - for KNN, this is given by the amount of N(true label)/N in the neighborhood around a query point
kmeans.predict_proba(X_test[:10])
# 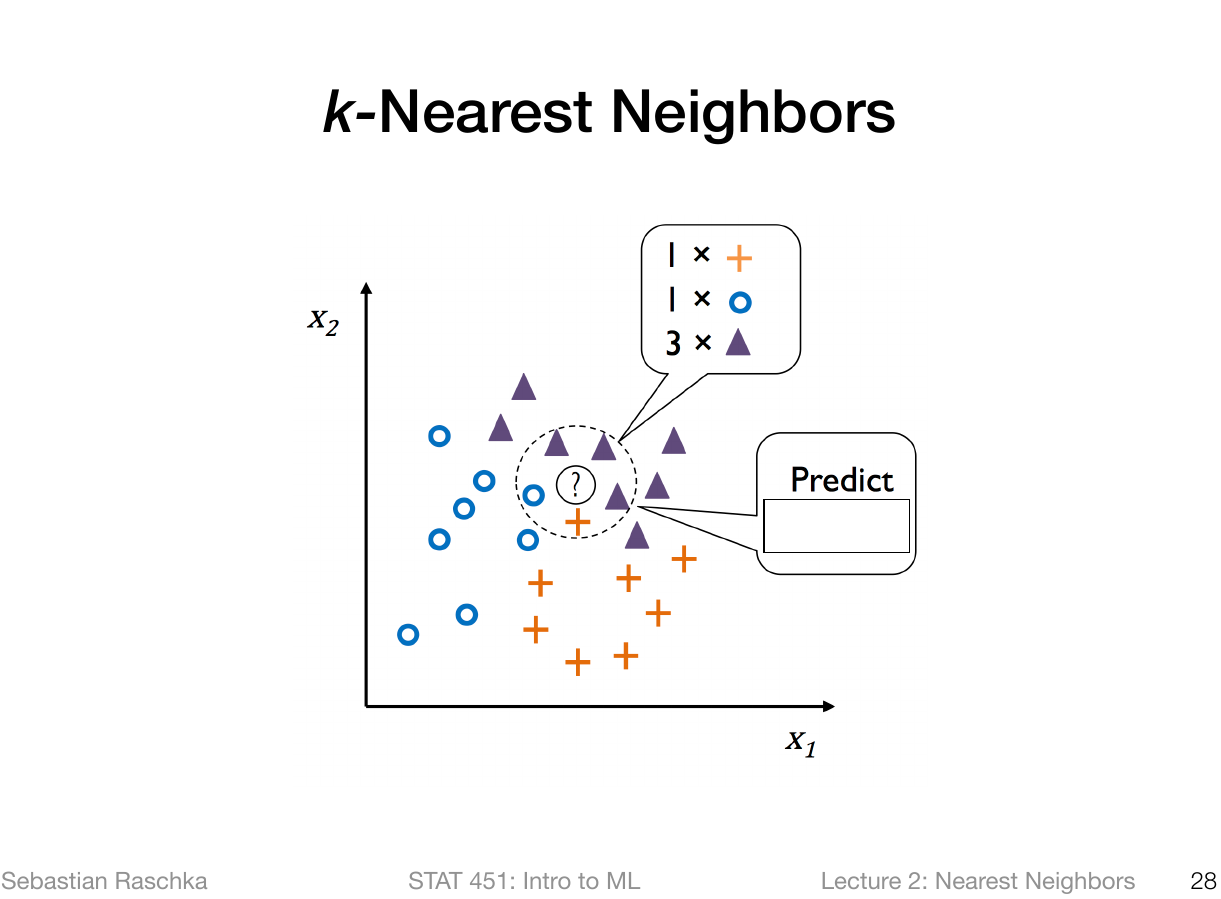
# - demonstrates how kNN classifyer is similar to `RandomForests`, `SVM`, ... :
# + spacial interpretation of the class prediction probability
# + the higher the probability for a sample, the more likely the sample belongs to `Adelie` in our case (i.e. the positive class in a binary classification setup)
#
# - relating this again to
#
# 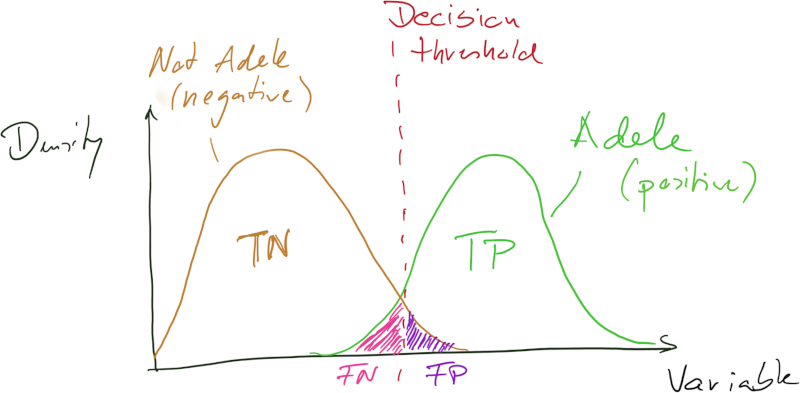
#
# the decision threshold for a `5`-neighborhood with a binary classification task is `0.6`, i.e. 3 of 5 neighbors have the positive class (then our query point will get the positive class assigned)
#
# - knowing these positive class prediction probabilities, I can now draw an envelope that gives me the ROC from the test set as with these probabilites and the theoretical threshold, we can compute FPR and TPR
#
#
# +
from sklearn.metrics import roc_curve
probs = kmeans.predict_proba(X_test)
pos_pred_probs = probs[:,-1]
fpr, tpr, thr = roc_curve(y_test, pos_pred_probs)
print('false positive rate\n',fpr)
print('true positive rate\n',tpr)
print('thresholds\n',thr)
# +
from sklearn.metrics import plot_roc_curve
roc = plot_roc_curve(kmeans, X_test, y_test)
# -
# - difference to plot with single entries? (size of test set -> only discrete values for single experiments, limited amount of samples)
# - summary of curve possible: AUC = area under curve
# - TPR and FPR used for ROC only -> similar plots possible with other variables, e.g. precision_recall_curve
# # Take-Aways
#
# <p><a href="https://commons.wikimedia.org/wiki/File:Roc-draft-xkcd-style.svg#/media/File:Roc-draft-xkcd-style.svg"><img src="https://upload.wikimedia.org/wikipedia/commons/thumb/3/36/Roc-draft-xkcd-style.svg/640px-Roc-draft-xkcd-style.svg.png" alt="Roc-draft-xkcd-style.svg"></a><br>By <a href="//commons.wikimedia.org/wiki/User:MartinThoma" title="User:MartinThoma">MartinThoma</a> - <span class="int-own-work" lang="en">Own work</span>, <a href="http://creativecommons.org/publicdomain/zero/1.0/deed.en" title="Creative Commons Zero, Public Domain Dedication">CC0</a>, <a href="https://commons.wikimedia.org/w/index.php?curid=70212136">Link</a></p>
#
# - nearest neighbor clustering algorithms are able to offer a probabilistic score for each predicted datum based on the neighborhood chosen
# - the ROC is an envelope that describes how well a classifyer performs given a fixed testset
# - ROC expresses the balance between true-positives and false positives
#
# # Further Reading
#
# - some parts of this material were inspired by [<NAME>](https://sebastianraschka.com)
# + [lecture 12.4, Receiver Operating Curve](https://youtu.be/GdSEkiArM3k)
#
# - a generally good resource
# + [Confusion_matrix](https://en.wikipedia.org/wiki/Confusion_matrix)
#
# - all of the above is nicely implemented and documented
# + [sklearn examples](https://scikit-learn.org/stable/auto_examples/model_selection/plot_roc.html#sphx-glr-auto-examples-model-selection-plot-roc-py)
# + [roc_curve API docs](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.roc_curve.html#sklearn.metrics.roc_curve)
#
# - [extensive discussion of ROC](https://stackabuse.com/understanding-roc-curves-with-python/)
#
| source/lesson04/script.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.6 64-bit (conda)
# metadata:
# interpreter:
# hash: b3ba2566441a7c06988d0923437866b63cedc61552a5af99d1f4fb67d367b25f
# name: python3
# ---
# # Kitti
import os
os.listdir()
# +
import pykitti
basedir = '2011_09_26_drive_0001_extract'
date = '2011_09_26'
drive = '0001'
# -
dataset = pykitti.raw(basedir,date,drive)
| kitti.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: cuml4
# language: python
# name: python3
# ---
# # Truncated Singular Value Decomposition (TSVD)
# The TSVD algorithm is a linear dimensionality reduction algorithm that works really well for datasets in which samples correlated in large groups. Unlike PCA, TSVD does not center the data before computation.
#
# The model can take array-like objects, either in host as NumPy arrays or in device (as Numba or cuda_array_interface-compliant), as well as cuDF DataFrames as the input.
#
# For information on converting your dataset to cuDF format, refer to the documentation: https://rapidsai.github.io/projects/cudf/en/latest/
#
# For information on cuML's TSVD implementation: https://rapidsai.github.io/projects/cuml/en/0.6.0/api.html#truncated-svd
# +
import os
import numpy as np
import pandas as pd
import cudf as gd
from cuml.datasets import make_blobs
from sklearn.decomposition import TruncatedSVD as skTSVD
from cuml.decomposition import TruncatedSVD as cumlTSVD
# +
n_samples = 2**15
n_features = 128
n_components = 2
random_state = 42
# -
# ## Generate Data
#
# ### GPU
# +
device_data, _ = make_blobs(
n_samples=n_samples, n_features=n_features, centers=1, random_state=7)
device_data = gd.DataFrame.from_gpu_matrix(device_data)
# -
# ### Host
host_data = device_data.to_pandas()
# ## Scikit-learn Model
# +
# %%time
tsvd_sk = skTSVD(n_components=n_components,
algorithm="arpack",
n_iter=5000,
tol=0.00001,
random_state=random_state)
result_sk = tsvd_sk.fit_transform(host_data)
# -
# ## cuML Model
# +
# %%time
tsvd_cuml = cumlTSVD(n_components=n_components,
algorithm="full",
n_iter=50000,
tol=0.00001,
random_state=random_state)
result_cuml = tsvd_cuml.fit_transform(device_data)
# -
# ## Evaluate Results
#
# ### Singular Values
passed = np.allclose(tsvd_sk.singular_values_,
tsvd_cuml.singular_values_,
atol=0.01)
print('compare tsvd: cuml vs sklearn singular_values_ {}'.format('equal' if passed else 'NOT equal'))
# ### Components
passed = np.allclose(tsvd_sk.components_,
np.asarray(tsvd_cuml.components_.as_gpu_matrix()),
atol=1e-2)
print('compare tsvd: cuml vs sklearn components_ {}'.format('equal' if passed else 'NOT equal'))
# ### Transform
# compare the reduced matrix
passed = np.allclose(result_sk, np.asarray(result_cuml.as_gpu_matrix()), atol=0.2)
# larger error margin due to different algorithms: arpack vs full
print('compare tsvd: cuml vs sklearn transformed results %s'%('equal'if passed else 'NOT equal'))
| cuml/tsvd_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#Data
revenue = [14574.49, 7606.46, 8611.41, 9175.41, 8058.65, 8105.44, 11496.28, 9766.09, 10305.32, 14379.96, 10713.97, 15433.50]
expenses = [12051.82, 5695.07, 12319.20, 12089.72, 8658.57, 840.20, 3285.73, 5821.12, 6976.93, 16618.61, 10054.37, 3803.96]
import numpy as np
profit_per_month = np.array(revenue) - np.array(expenses)
profit_per_month # Revenue - Expenses
profit_per_month_after_tax = profit_per_month * 0.7
profit_per_month_after_tax
profit_margin_per_month = profit_per_month_after_tax / revenue
profit_margin_per_month
mean_year_profit_after_tax = profit_per_month_after_tax.mean()
mean_year_profit_after_tax
filtered_months = [] # where the profit after tax was fgreater than the mean for the year
filtered_months = profit_per_month_after_tax > mean_year_profit_after_tax;
profit_good_months = profit_per_month_after_tax[filtered_months]
good_months
print(profit_per_month_after_tax)
filtered_months = profit_per_month_after_tax > mean_year_profit_after_tax;
profit_bad_months = profit_per_month_after_tax[filtered_months]
profit_bad_months
filtered_months = profit_per_month_after_tax == profit_per_month_after_tax.max();
profit_best_month = profit_per_month_after_tax[filtered_months]
best_month = np.where(profit_per_month_after_tax == profit_best_month)[0][0] + 1
best_month
filtered_months = profit_per_month_after_tax == profit_per_month_after_tax.min();
profit_worst_month = profit_per_month_after_tax[filtered_months]
worst_month = np.where(profit_per_month_after_tax == profit_worst_month)[0][0] + 1
worst_month
| Financial Statement Analysis Assignment Solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# <table style="width:100%; background-color: #D9EDF7">
# <tr>
# <td style="border: 1px solid #CFCFCF">
# <b>Weather data: Main notebook</b>
# <ul>
# <li>Main Notebook</li>
# <li><a href="download.ipynb"> Downloading Notebook</a></li>
# <li><a href="documentation.ipynb">Documentation</a></li>
# </ul>
# <br>This Notebook is part of the <a href="http://data.open-power-system-data.org/weather_data">Weather data Datapackage</a> of <a href="http://open-power-system-data.org">Open Power System Data</a>.
# </td>
# </tr>
# </table>
# # Table of Contents
# * [1. About Open Power System Data](#1.-About-Open-Power-System-Data)
# * [2. About Jupyter Notebooks and GitHub](#2.-About-Jupyter-Notebooks-and-GitHub)
# * [3. About this datapackage](#3.-About-this-datapackage)
# * [4. Data sources](#4.-Data-sources)
# * [5. License](#5.-License)
#
# ---
# # 1. About Open Power System Data
# This notebook is part of the project [Open Power System Data](http://open-power-system-data.org). Open Power System Data develops a platform for free and open data for electricity system modeling. We collect, check, process, document, and provide data that are publicly available but currently inconvenient to use.
# More info on Open Power System Data:
# - [Information on the project on our website](http://open-power-system-data.org)
# - [Data and metadata on our data platform](http://data.open-power-system-data.org)
# - [Data processing scripts on our GitHub page](https://github.com/Open-Power-System-Data)
# # 2. About Jupyter Notebooks and GitHub
# This file is a [Jupyter Notebook](http://jupyter.org/). A Jupyter Notebook is a file that combines executable programming code with visualizations and comments in markdown format, allowing for an intuitive documentation of the code. We use Jupyter Notebooks for combined coding and documentation. We use Python 3 as programming language. All Notebooks are stored on [GitHub](https://github.com/), a platform for software development, and are publicly available. More information on our IT-concept can be found [here](http://open-power-system-data.org/it). See also our [step-by-step manual](http://open-power-system-data.org/step-by-step) how to use the dataplatform.
# # 3. About this datapackage
# We provide data in different chunks, or [datapackages](http://frictionlessdata.io/data-packages/). The one you are looking at right now, [Weather data](http://data.open-power-system-data.org/weather_data/), contains scripts that allow the a download, subset and processing of [MERRA-2](http://gmao.gsfc.nasa.gov/reanalysis/MERRA-2/) datasets (provided by NASA Goddard Space Flight Center) and export them as CSV.
#
# **Weather data differ significantly from the other data types used resp. provided by OPSD** in that the sheer size of the data packages greatly exceeds OPSD's capacity to host them in a similar way as feed-in timeseries, power plant data etc. While the other data packages also offer a complete one-klick download of the bundled data packages with all relevant data this is impossible for weather datasets like MERRA-2 due to their size (variety of variables, very long timespan, huge geographical coverage etc.). It would make no sense to mirror the data from the NASA servers.
#
# Instead we choose to provide a **documented methodological script** (as a kind of tutorial). The method describes one way to automatically obtain the desired weather data from the MERRA-2 database and simplifies resp. unifies alternative manual data obtaining methods in a single script.
#
# To access, subset and download the MERRA-2 database we use the [OPeNDAP](https://www.opendap.org/) framework. The use of MERRA-2 is only exemplary for this method - through the use of OPenDAP it can be adapted to other datasets using the same protocol.
#
# This method/script is tailored to the needs of energy system modellers that a) do not want to downlad and haggle with the original MERRA-2 data manually, and those who on the other side also b) do not just want to take over ready-made feed-ins calculated by tools like [renewables.ninja](https://www.renewables.ninja/) but rather want to use their own feed-in tools with processed weather data.
#
# See the [OPSD wiki](https://github.com/Open-Power-System-Data/common/wiki/Information-on-weather-data) on Github for more information on MERRA-2 weather data, OPeNDAP and OPSD's approach.
# # 4. Data sources
# The data source is the [MERRA-2 dataset](http://gmao.gsfc.nasa.gov/reanalysis/MERRA-2/) provided by NASA Goddard Space Flight Center. Specifically we use the following datasets (Two–Dimensional, Hourly, Time‐averaged Assimilation and Forecast Fields)
# * [tavg1_2d_slv_Nx (M2T1NXSLV)](http://goldsmr4.sci.gsfc.nasa.gov/opendap/MERRA2/M2T1NXSLV.5.12.4/contents.html) (Single Level Diagnostics Collection)
# * [tavg1_2d_flx_Nx (M2T1NXFLX)](http://goldsmr4.sci.gsfc.nasa.gov/opendap/MERRA2/M2T1NXFLX.5.12.4/contents.html) (Turbulence Collection)
# * [tavg1_2d_rad_Nx (M2T1NXRAD)](http://goldsmr4.sci.gsfc.nasa.gov/opendap/MERRA2/M2T1NXRAD.5.12.4/contents.html) (Radiation Collection)
# # 5. License
# This notebook as well as all other documents in this repository is published under the [MIT License](https://opensource.org/licenses/MIT).
| main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# <script async src="https://www.googletagmanager.com/gtag/js?id=UA-59152712-8"></script>
# <script>
# window.dataLayer = window.dataLayer || [];
# function gtag(){dataLayer.push(arguments);}
# gtag('js', new Date());
#
# gtag('config', 'UA-59152712-8');
# </script>
#
# # `GiRaFFE_HO` C code library: Conservative-to-Primitive and Primitive-to-Conservative Solvers
#
# ## Author: <NAME>
#
# <a id='intro'></a>
#
# **Module Status:** <font color=orange><b> Self-Validated </b></font>
#
# **Validation Notes:** These codes are modified versions of the working code used by the original `GiRaFFE`.
#
# ## Introduction:
# This writes and documents the C code that `GiRaFFE_HO` uses in order to update the Valencia 3-velocity at each timestep. It also computes corrections to the densitized Poynting flux in order to keep the physical quantities from violating the GRFFE constraints.
#
# These algorithms are adapted from the original `GiRaFFE` code (see [arxiv:1704.00599v2](https://arxiv.org/abs/1704.00599v2)), based on the description in [arXiv:1310.3274v2](https://arxiv.org/pdf/1310.3274v2.pdf). They have been modified to work with the NRPy+ infrastructure instead of the Einstein Toolkit. They have also been modified to use the Valencia 3-velocity instead of the drift velocity.
# <a id='toc'></a>
#
# # Table of Contents
# $$\label{toc}$$
#
# This notebook is organized as follows
#
# 1. [Step 1](#c2p): The conservative-to-primitive solver
# 1. [Step 1.a](#definitions): Function definitions and inverse metric
# 1. [Step 1.b](#header_loop): The function header and loop parameters
# 1. [Step 1.c](#reading): Reading from memory
# 1. [Step 1.d](#ortho_s_b): Enforce the orthogonality of $\tilde{S}_i$ and $B^i$
# 1. [Step 1.e](#vel_cap): Rescale ${\tilde S}_i$ to limit the Lorentz factor and enforce the velocity cap
# 1. [Step 1.f](#update_vel): Recompute the velocities at the new timestep
# 1. [Step 1.g](#current_sheet): Enforce the Current Sheet prescription
# 1. [Step 2](#p2c): The primitive-to-conservative solver
# 1. [Step 3](#code_validation): Code Validation against original C code
# 1. [Step 4](#latex_pdf_output): Output this notebook to $\LaTeX$-formatted PDF file
import os
import cmdline_helper as cmd
outdir = "GiRaFFE_HO/GiRaFFE_Ccode_validation/"
cmd.mkdir(outdir)
# <a id='c2p'></a>
#
# # Step 1: The conservative-to-primitive solver \[Back to [top](#toc)\]
# $$\label{c2p}$$
#
# We start with the Conservative-to-Primitive solver. This function is called after the vector potential and Poynting vector have been evolved at a timestep and updates the velocities. The algorithm will be as follows:
#
# 1. Enforce the orthogonality of ${\tilde S}_i$ and $B^i$
# * ${\tilde S}_i \rightarrow {\tilde S}_i - ({\tilde S}_j {\tilde B}^j) {\tilde B}_i/{\tilde B}^2$
# 2. Rescale ${\tilde S}_i$ to limit the Lorentz factor and enforce the velocity cap
# * $f = \sqrt{(1-\gamma_{\max}^{-2}){\tilde B}^4/(16 \pi^2 \gamma {\tilde S}^2)}$
# * ${\tilde S}_i \rightarrow {\tilde S}_i \min(1,f)$
# 3. Recompute the velocities at the new timestep
# * $v^i = 4 \pi \gamma^{ij} {\tilde S}_j \gamma^{-1/2} B^{-2}$
# 4. Enforce the Current Sheet prescription
# * ${\tilde n}_i v^i = 0$
#
# We will begin simply by creating the file. We will also `#include` the header file `<sys/time.h>` and define $\pi$.
# +
# %%writefile $outdir/driver_conserv_to_prims_FFE.C
/* We evolve forward in time a set of functions called the
* "conservative variables" (magnetic field and Poynting vector),
* and any time the conserv's are updated, we must recover the
* primitive variables (velocities), before reconstructing & evaluating
* the RHSs of the MHD equations again.
*
* This file contains the routine for this algebraic calculation.
* The velocity is calculated with formula (85), arXiv:1310.3274v2
* $v^i = 4 \pi \alpha \gamma^{ij} {\tilde S}_j \gamma{-1/2} B^{-2} - \beta^i$
* The force-free condition: $B^2>E^2$ is checked before computing the velocity.
* and after imposing the constraint ${\tilde B}^i {\tilde S}_i = 0$
* The procedure is as described in arXiv:1310.3274v2:
* 1. ${\tilde S}_i ->{\tilde S}_i - ({\tilde S}_j {\tilde B}^j) {\tilde B}_i/{\tilde B}^2$
* 2. $f = \sqrt{(1-\gamma_{max}^{-2}){\tilde B}^4/(16 \pi^2 \gamma {\tilde S}^2)}$
* 3. ${\tilde S}_i -> {\tilde S}_i min(1,f)
* 4. $v^i = 4 \pi \alpha \gamma^{ij} {\tilde S}_j \gamma{-1/2} B^{-2} - \beta^i$
* 5. ${\tilde n}_i v^i = 0$
*
* All equations are from: http://arxiv.org/pdf/1310.3274.pdf (v2)
* */
//#include <iostream>
//#include <iomanip>
//#include <fstream>
#include <sys/time.h>
//#include <cmath>
//#include <ctime>
//#include <cstdlib>
//#include "Symmetry.h"
#ifndef M_PI
#define M_PI 3.141592653589793238463
#endif
# -
# <a id='definitions'></a>
#
# ## Step 1.a: Function definitions and inverse metric \[Back to [top](#toc)\]
# $$\label{definitions}$$
#
# The next order of business will be to write the function definitions for some functions that the main, conservative-to-primitive function will need as well as some useful macros. First, we write a basic macro to find the minimum. We write the function definition of the Primitive-to-Conservative solver, `GiRaFFE_HO_compute_conservatives` and include it from the next file we will write [below](#p2c). We define the standard indexing macros used in NRPy+ that map the values of each gridfunction at each point in three-dimensional space to the elements of a single one-dimensional array.
#
# Next, we write the function that calculates the metric determinant and inverse from the metric, `GiRaFFE_HO_update_metric_det_inverse()`. The included header file `"metric_quantities.h"` does most of that work for us by looping over the entire grid and performing the matrix inverse operation.
#
# Notice the use of the flag `-a` in the cell magic command below - this will append to the file instead of overwriting it.
# +
# %%writefile -a $outdir/driver_conserv_to_prims_FFE.C
#define MIN(a,b) ( ((a) < (b)) ? (a) : (b) )
void GiRaFFE_HO_compute_conservatives(const REAL gxxL,const REAL gxyL,const REAL gxzL,const REAL gyyL,const REAL gyzL,const REAL gzzL,
const REAL BxL, const REAL ByL, const REAL BzL, const REAL vxL, const REAL vyL, const REAL vzL,
//const REAL betaxL, const REAL betayL, const REAL betazL, const REAL alpL,
const REAL sqrtg,REAL *StildeD0L, REAL *StildeD1L, REAL *StildeD2L);
#include "compute_conservatives_FFE.C"
#define REAL double
#define IDX4(g,i,j,k) \
( (i) + Nxx_plus_2NGHOSTS[0] * ( (j) + Nxx_plus_2NGHOSTS[1] * ( (k) + Nxx_plus_2NGHOSTS[2] * (g) ) ) )
#define IDX3(i,j,k) ( (i) + Nxx_plus_2NGHOSTS[0] * ( (j) + Nxx_plus_2NGHOSTS[1] * (k) ) )
// Assuming idx = IDX3(i,j,k). Much faster if idx can be reused over and over:
#define IDX4pt(g,idx) ( (idx) + (Nxx_plus_2NGHOSTS[0]*Nxx_plus_2NGHOSTS[1]*Nxx_plus_2NGHOSTS[2]) * (g) )
void GiRaFFE_HO_update_metric_det_inverse(const int Nxx_plus_2NGHOSTS[3],const REAL dxx[3],REAL *xx[3],REAL *aux_gfs) {
#include "metric_quantities.h"
}
# -
# <a id='header_loop'></a>
#
# ## Step 1.b: The function header and loop parameters \[Back to [top](#toc)\]
# $$\label{header_loop}$$
#
# Now, we get into the meat of the Conservative-to-Primitive solver itself. Note that, in addition to the basic parameters defining the grid, we also pass the conservative variables `in_gfs` and primitive variables `aux_gfs`. Then, we define the boundaries over which we would like to recompute the primitive variables by setting `imin`, `imax`, `jmin`, `jmax`, `kmin`, and `kmax`. Currently, these are set such that we only loop over the interior of the grid.
# +
# %%writefile -a $outdir/driver_conserv_to_prims_FFE.C
void GiRaFFE_HO_conserv_to_prims_FFE(const int Nxx[3],const int Nxx_plus_2NGHOSTS[3],const REAL dxx[3],
REAL *xx[3], REAL *in_gfs, REAL *aux_gfs) {
//printf("Starting conservative-to-primitive solver...\n");
/*// We use proper C++ here, for file I/O later.
using namespace std;*/
const int imin=NGHOSTS,jmin=NGHOSTS,kmin=NGHOSTS;
const int imax=Nxx_plus_2NGHOSTS[0]-NGHOSTS,jmax=Nxx_plus_2NGHOSTS[1]-NGHOSTS,kmax=Nxx_plus_2NGHOSTS[2]-NGHOSTS;
const REAL dz = dxx[2];
REAL error_int_numer=0,error_int_denom=0;
int num_vel_limits=0,num_vel_nulls_current_sheet=0;
GiRaFFE_HO_update_metric_det_inverse(Nxx_plus_2NGHOSTS, dxx, xx,aux_gfs);
#pragma omp parallel for reduction(+:error_int_numer,error_int_denom,num_vel_limits,num_vel_nulls_current_sheet) schedule(static)
for(int k=kmin;k<kmax;k++)
for(int j=jmin;j<jmax;j++)
for(int i=imin;i<imax;i++) {
# -
# <a id='reading'></a>
#
# ## Step 1.c: Reading from memory \[Back to [top](#toc)\]
# $$\label{reading}$$
#
# Next, we will read in values from memory. Note the `if` statement - this allows us to disable the evolution of gridfunctions inside of a specifiable radius, which is especially useful for neutron star simulations.
#
# In addition to reading in values from memory, we also compute the densitized magnetic field $\tilde{B}^i = B^i \sqrt{\gamma}$ and the index-lowered form, $\tilde{B}_i = \gamma_{ij} \tilde{B}^j$.
# %%writefile -a $outdir/driver_conserv_to_prims_FFE.C
const int index = IDX3(i,j,k);
const REAL xx0 = xx[0][i];
const REAL xx1 = xx[1][j];
const REAL xx2 = xx[2][k];
const REAL rL = sqrt(xx0*xx0+xx1*xx1+xx2*xx2);
if(rL>min_radius_inside_of_which_conserv_to_prims_FFE_and_FFE_evolution_is_DISABLED) {
const REAL sqrtg = sqrt(aux_gfs[IDX4pt(GAMMADETGF, index)]); // Determinant of 3-metric
// \gamma_{ij}, computed from \tilde{\gamma}_{ij}
const REAL gxxL = aux_gfs[IDX4pt(GAMMADD00GF, index)];
const REAL gxyL = aux_gfs[IDX4pt(GAMMADD01GF, index)];
const REAL gxzL = aux_gfs[IDX4pt(GAMMADD02GF, index)];
const REAL gyyL = aux_gfs[IDX4pt(GAMMADD11GF, index)];
const REAL gyzL = aux_gfs[IDX4pt(GAMMADD12GF, index)];
const REAL gzzL = aux_gfs[IDX4pt(GAMMADD22GF, index)];
// \gamma^{ij} = psim4 * \tilde{\gamma}^{ij}
const REAL gupxxL = aux_gfs[IDX4pt(GAMMAUU00GF, index)];
const REAL gupxyL = aux_gfs[IDX4pt(GAMMAUU01GF, index)];
const REAL gupxzL = aux_gfs[IDX4pt(GAMMAUU02GF, index)];
const REAL gupyyL = aux_gfs[IDX4pt(GAMMAUU11GF, index)];
const REAL gupyzL = aux_gfs[IDX4pt(GAMMAUU12GF, index)];
const REAL gupzzL = aux_gfs[IDX4pt(GAMMAUU22GF, index)];
// Read in magnetic field and momentum variables once from memory, since memory access is expensive:
const REAL BU0L = aux_gfs[IDX4pt(BU0GF, index)];
const REAL BU1L = aux_gfs[IDX4pt(BU1GF, index)];
const REAL BU2L = aux_gfs[IDX4pt(BU2GF, index)];
// End of page 7 on http://arxiv.org/pdf/1310.3274.pdf
const REAL BtildexL = BU0L*sqrtg;
const REAL BtildeyL = BU1L*sqrtg;
const REAL BtildezL = BU2L*sqrtg;
const REAL Btilde_xL = gxxL*BtildexL + gxyL*BtildeyL + gxzL*BtildezL;
const REAL Btilde_yL = gxyL*BtildexL + gyyL*BtildeyL + gyzL*BtildezL;
const REAL Btilde_zL = gxzL*BtildexL + gyzL*BtildeyL + gzzL*BtildezL;
REAL StildeD0L = in_gfs[IDX4pt(STILDED0GF, index)];
REAL StildeD1L = in_gfs[IDX4pt(STILDED1GF, index)];
REAL StildeD2L = in_gfs[IDX4pt(STILDED2GF, index)];
const REAL StildeD0_orig = StildeD0L;
const REAL StildeD1_orig = StildeD1L;
const REAL StildeD2_orig = StildeD2L;
const REAL ValenciavU0_orig = aux_gfs[IDX4pt(VALENCIAVU0GF, index)];
const REAL ValenciavU1_orig = aux_gfs[IDX4pt(VALENCIAVU1GF, index)];
const REAL ValenciavU2_orig = aux_gfs[IDX4pt(VALENCIAVU2GF, index)];
//const REAL alpL = alp[index];
//const REAL fourpialpha = 4.0*M_PI*alpL;
const REAL fourpi = 4.0*M_PI;
//const REAL betaxL = betax[index];
//const REAL betayL = betay[index];
//const REAL betazL = betaz[index];
# <a id='ortho_s_b'></a>
#
# ## Step 1.d: Enforce the orthogonality of $\tilde{S}_i$ and $B^i$ \[Back to [top](#toc)\]
# $$\label{ortho_s_b}$$
#
# Now, we will enforce the orthogonality of the magnetic field and densitized poynting flux:
# $${\tilde S}_i \rightarrow {\tilde S}_i - ({\tilde S}_j {\tilde B}^j) {\tilde B}_i/{\tilde B}^2$$
# First, we compute the inner products ${\tilde S}_j {\tilde B}^j$ and ${\tilde B}^2 = \gamma_{ij} {\tilde B}^i {\tilde B}^j$; then, we subtract $({\tilde S}_j {\tilde B}^j) {\tilde B}_i/{\tilde B}^2$ from ${\tilde S}_i$. We thus guarantee that ${\tilde S}_i B^i=0$.
#
# Having fixed ${\tilde S}_i$, we will also compute the related quantities ${\tilde S}^i = \gamma^{ij} {\tilde S}_j$ and ${\tilde S}^2 = {\tilde S}_i {\tilde S}^i$.
#
# Note also the macro `APPLY_GRFFE_FIXES`; by commenting out this one line, we can easily disable the GRFFE fixes for testing purposes.
# +
# %%writefile -a $outdir/driver_conserv_to_prims_FFE.C
//* 1. Just below Eq 90: Enforce orthogonality of B^i & S^i, so that B^i S_i = 0
//* Correction ${\tilde S}_i ->{\tilde S}_i - ({\tilde S}_j {\tilde B}^j) {\tilde B}_i/{\tilde B}^2$
//* NOTICE THAT THE {\tilde B}_i IS LOWERED, AS IT SHOULD BE. THIS IS A TYPO IN PASCHALIDIS ET AL.
// First compute Btilde^i Stilde_i:
const REAL BtildeiSt_i = StildeD0L*BtildexL + StildeD1L*BtildeyL + StildeD2L*BtildezL;
//printf("xterm = %f ; yterm = %f ; zterm = %f\n",StildeD0L*BtildexL,StildeD1L*BtildeyL,StildeD2L*BtildezL);
// Then compute (Btilde)^2
const REAL Btilde2 = gxxL*BtildexL*BtildexL + gyyL*BtildeyL*BtildeyL + gzzL*BtildezL*BtildezL
+ 2.0*(gxyL*BtildexL*BtildeyL + gxzL*BtildexL*BtildezL + gyzL*BtildeyL*BtildezL);
#define APPLY_GRFFE_FIXES
// Now apply constraint: Stilde_i = Stilde_i - (Btilde^i Stilde_i) / (Btilde)^2
#ifdef APPLY_GRFFE_FIXES
StildeD0L -= BtildeiSt_i*Btilde_xL/Btilde2;
StildeD1L -= BtildeiSt_i*Btilde_yL/Btilde2;
StildeD2L -= BtildeiSt_i*Btilde_zL/Btilde2;
//printf("BtildeiSt_i = %f ; Btilde2 = %f\n",BtildeiSt_i,Btilde2);
#endif
// Now that tildeS_i has been fixed, let's compute tildeS^i:
REAL mhd_st_upx = gupxxL*StildeD0L + gupxyL*StildeD1L + gupxzL*StildeD2L;
REAL mhd_st_upy = gupxyL*StildeD0L + gupyyL*StildeD1L + gupyzL*StildeD2L;
REAL mhd_st_upz = gupxzL*StildeD0L + gupyzL*StildeD1L + gupzzL*StildeD2L;
// Just below Eq. 86 in http://arxiv.org/pdf/1310.3274.pdf:
REAL St2 = StildeD0L*mhd_st_upx + StildeD1L*mhd_st_upy + StildeD2L*mhd_st_upz;
# -
# <a id='vel_cap'></a>
#
# ## Step 1.e: Rescale ${\tilde S}_i$ to limit the Lorentz factor and enforce the velocity cap \[Back to [top](#toc)\]
# $$\label{vel_cap}$$
#
# The next fix that we will apply limits the Lorentz factor. That is, we define the factor $f$ as
# $$f = \sqrt{(1-\gamma_{\max}^{-2}){\tilde B}^4/(16 \pi^2 \gamma {\tilde S}^2)}.$$
# Note that $\gamma_\max$ here refers to the Lorentz factor, *not* the determinant of the metric.
#
# If $f<1$, (or if, as the code actually calculates, $\tilde{S}^2 > f^2 \tilde{S}^2$), we rescale the components of ${\tilde S}_i$ by $f$. That is, if
# $$\tilde{S}^2 > (1-\gamma_{\max}^{-2}){\tilde B}^4/(16 \pi^2 \gamma),$$
# we must then set
# $${\tilde S}_i \rightarrow {\tilde S}_i \min(1,f).$$
#
# We then double check that the cap was effective by checking if $\tilde{S}^2 > f^2 \tilde{S}^2$ for large $\gamma_\max$ and error out if it wasn't.
# **TODO:** Why do we check the case $\gamma_\max \rightarrow \infty$?
# +
# %%writefile -a $outdir/driver_conserv_to_prims_FFE.C
//* 2. Eq. 92: Factor $f = \sqrt{(1-\gamma_{max}^{-2}){\tilde B}^4/(16 \pi^2 \gamma {\tilde S}^2)}$
#ifdef APPLY_GRFFE_FIXES
const REAL gmax = GAMMA_SPEED_LIMIT;
if(St2 > (1.0 - 1.0/(gmax*gmax))*Btilde2*Btilde2/ (16.0*M_PI*M_PI*sqrtg*sqrtg)) {
const REAL fact = sqrt((1.0 - 1.0/(gmax*gmax))/St2)*Btilde2/(4.0*M_PI*sqrtg);
//* 3. ${\tilde S}_i -> {\tilde S}_i min(1,f)
StildeD0L *= MIN(1.0,fact);
StildeD1L *= MIN(1.0,fact);
StildeD2L *= MIN(1.0,fact);
// Recompute S^i
mhd_st_upx = gupxxL*StildeD0L + gupxyL*StildeD1L + gupxzL*StildeD2L;
mhd_st_upy = gupxyL*StildeD0L + gupyyL*StildeD1L + gupyzL*StildeD2L;
mhd_st_upz = gupxzL*StildeD0L + gupyzL*StildeD1L + gupzzL*StildeD2L;
/*
printf("%e %e %e | %e %e %e | %e %e %e | oldgamma: %e %e should be > %e vfix\n",x[index],y[index],z[index],
BU0L,BU1L,BU2L,
St2,(1.0 - 1.0/(gmax*gmax))*Btilde2*Btilde2/ (16.0*M_PI*M_PI*sqrtg*sqrtg),gmax,
sqrt(Btilde2 / (Btilde2 - 16*M_PI*M_PI*sqrtg*sqrtg * St2 / Btilde2) ) , Btilde2,16*M_PI*M_PI*sqrtg*sqrtg * St2 / Btilde2 );
//exit(1);
*/
// Recompute Stilde^2:
St2 = StildeD0L*mhd_st_upx + StildeD1L*mhd_st_upy + StildeD2L*mhd_st_upz;
if( St2 >= Btilde2*Btilde2/ (16.0*M_PI*M_PI*sqrtg*sqrtg) ) {
printf("ERROR: Velocity cap fix wasn't effective; still have B^2 > E^2\n"); exit(1);
}
num_vel_limits++;
}
#endif
# -
# <a id='update_vel'></a>
#
# ## Step 1.f: Recompute the velocities at the new timestep \[Back to [top](#toc)\]
# $$\label{update_vel}$$
#
# Finally, we can calculate the velocities. In the source used, the equation to compute the drift velocity is given as
# $$v^i = 4 \pi \alpha \gamma^{ij} {\tilde S}_j \gamma^{-1/2} B^{-2} - \beta^i.$$
# However, we wish to use the Valencia velocity instead. Since the Valencia velocity $\bar{v}^i = \frac{1}{\alpha} \left( v^i + \beta^i \right)$, we will code
# $$\bar{v}^i = 4 \pi \frac{\gamma^{ij} {\tilde S}_j}{\sqrt{\gamma} B^2}.$$
#
# %%writefile -a $outdir/driver_conserv_to_prims_FFE.C
//* 4. Eq. 85: $v^i = 4 pi \alpha \gamma^{ij} {\tilde S}_j \gamma{-1/2} B^{-2} - \beta^i$:
// See, e.g., Eq 71 in http://arxiv.org/pdf/1310.3274.pdf
// ... or end of page 7 on http://arxiv.org/pdf/1310.3274.pdf:
const REAL B2 = Btilde2/(sqrtg*sqrtg);
/*
Eq. 75:
v^i = \alpha \gamma^{ij} S_j / \mathcal{B}^2 - \beta^i
Eq. 7: \mathcal{B}^{\mu} = B^{\mu}/\sqrt{4 \pi}
-> v^i = 4 \pi \alpha \gamma^{ij} S_j / B^2 - \beta^i
Eq. 79: \tilde{S_i} = \sqrt{\gamma} S_i
-> v^i = 4 \pi \alpha \gamma^{ij} \tilde{S}_j / (\sqrt{\gamma} B^2) - \beta^i
*/
// Modified from the original GiRaFFE to use Valencia, not drift velocity
const REAL ValenciavU0L = fourpi*mhd_st_upx/(sqrtg*B2);
const REAL ValenciavU1L = fourpi*mhd_st_upy/(sqrtg*B2);
/* ValenciavU2L not necessarily const! See below. */
REAL ValenciavU2L = fourpi*mhd_st_upz/(sqrtg*B2);
# <a id='current_sheet'></a>
#
# ## Step 1.g: Enforce the Current Sheet prescription \[Back to [top](#toc)\]
# $$\label{current_sheet}$$
#
# Now, we seek to handle any current sheets (a physically important phenomenon) that might form. This algorithm will preserve current sheets that form in the xy-plane by preventing our numerical scheme from dissipating them. After fixing the z-component of the velocity, we recompute the conservative variables $\tilde{S}_i$ to be consistent with the new velocities.
#
# Thus, if we are within four gridpoints **(Why is it 4.01?)** of $z=0$, we set the component of the velocity perpendicular to the current sheet to zero by $n_i v^i = 0$, where $n_i = \gamma_{ij} n^j$ is a unit normal to the current sheet and $n^j = \delta^{jz} = (0\ 0\ 1)$. For drift velocity, this means we just set $$v^z = -\frac{\gamma_{xz} v^x + \gamma_{yz} v^y}{\gamma_{zz}}.$$ This reduces to $v^z = 0$ in flat space, as one would expect. **This should be checked for Valencia velocity.** The code also tracks the number of times this correction has been performed.
# %%writefile -a $outdir/driver_conserv_to_prims_FFE.C
//* 5. Eq. 94: ${\tilde n}_i v^i = 0$ in the current sheet region
// n^i is defined as the normal from the current sheet, which lies in the
// xy-plane (z=0). So n = (0,0,1)
#ifdef APPLY_GRFFE_FIXES
if(current_sheet_null_v) {
if (fabs(xx2) <= (4.0 + 1.0e-2)*dz ) {
//ValenciavU2L = 0.0;
ValenciavU2L = - (ValenciavU0L*gxzL + ValenciavU1L*gyzL) / gzzL;
// FIXME: This is probably not right, but also definitely not the problem.
// ValenciavU2L reset: TYPICALLY WOULD RESET CONSERVATIVES TO BE CONSISTENT. LET'S NOT DO THAT, TO AVOID MESSING UP B-FIELDS
if(1==1) {
GiRaFFE_HO_compute_conservatives(gxxL, gxyL, gxzL, gyyL, gyzL, gzzL,
BU0L, BU1L, BU2L, ValenciavU0L, ValenciavU1L, ValenciavU2L,
/*const REAL betaxL, const REAL betayL, const REAL betazL, const REAL alpL,*/
sqrtg, &StildeD0L, &StildeD1L, &StildeD2L);
}
num_vel_nulls_current_sheet++;
}
}
#endif
aux_gfs[IDX4pt(VALENCIAVU0GF, index)] = ValenciavU0L;
aux_gfs[IDX4pt(VALENCIAVU1GF, index)] = ValenciavU1L;
aux_gfs[IDX4pt(VALENCIAVU2GF, index)] = ValenciavU2L;
# To finish out this portion of the algorithm, we include some diagnostic code (commented out for now) that compares the velocities before and after the current sheet prescription. We also write the new values of $\tilde{S}_i$ to memory, since they may have been changed in the first or third of the GRFFE fixes.
# %%writefile -a $outdir/driver_conserv_to_prims_FFE.C
//Now we compute the difference between original & new conservatives, for diagnostic purposes:
//error_int_numer += fabs(StildeD0L - StildeD0_orig) + fabs(StildeD1L - StildeD1_orig) + fabs(StildeD2L - StildeD2_orig);
//error_int_denom += fabs(StildeD0_orig) + fabs(StildeD1_orig) + fabs(StildeD2_orig);
/*
if(fabs(ValenciavU0_orig) > 1e-13 && fabs(ValenciavU0L-ValenciavU0_orig)/ValenciavU0_orig > 1e-2) printf("BAD ValenciavU0: %e %e | %e %e %e\n",ValenciavU0L,ValenciavU0_orig,x[index],y[index],z[index]);
if(fabs(ValenciavU1_orig) > 1e-13 && fabs(ValenciavU1L-ValenciavU1_orig)/ValenciavU1_orig > 1e-2) printf("BAD ValenciavU1: %e %e | %e %e %e\n",ValenciavU1L,ValenciavU1_orig,x[index],y[index],z[index]);
if(fabs(ValenciavU2_orig) > 1e-13 && fabs(ValenciavU2L-ValenciavU2_orig)/ValenciavU2_orig > 1e-2) printf("BAD ValenciavU2: %e %e | %e %e %e\n",ValenciavU2L,ValenciavU2_orig,x[index],y[index],z[index]);
*/
error_int_numer += fabs(ValenciavU0L - ValenciavU0_orig) + fabs(ValenciavU1L - ValenciavU1_orig) + fabs(ValenciavU2L - ValenciavU2_orig);
error_int_denom += fabs(ValenciavU0_orig) + fabs(ValenciavU1_orig) + fabs(ValenciavU2_orig);
in_gfs[IDX4pt(STILDED0GF, index)] = StildeD0L;
in_gfs[IDX4pt(STILDED1GF, index)] = StildeD1L;
in_gfs[IDX4pt(STILDED2GF, index)] = StildeD2L;
}
}
}
# <a id='p2c'></a>
#
# # Step 2: The primitive-to-conservative solver \[Back to [top](#toc)\]
# $$\label{p2c}$$
#
# This function is used to recompute the conservatives $\tilde{S}_i$ after the 3-velocity is changed as part of the current sheet prescription. It implements the same equation used to compute the initial Poynting flux from the initial velocity: $$\tilde{S}_i = \gamma_{ij} \frac{v^j \sqrt{\gamma}B^2}{4 \pi}$$ in terms of the Valencia 3-velocity. In the implementation here, we first calculate $B^2 = \gamma_{ij} B^i B^j$, then $v_i = \gamma_{ij} v^j$ before we calculate the equivalent expression $$\tilde{S}_i = \frac{v_j \sqrt{\gamma}B^2}{4 \pi}.$$
# %%writefile $outdir/compute_conservatives_FFE.C
void GiRaFFE_HO_compute_conservatives(const REAL gxxL,const REAL gxyL,const REAL gxzL,const REAL gyyL,const REAL gyzL,const REAL gzzL,
const REAL BxL, const REAL ByL, const REAL BzL, const REAL vxL, const REAL vyL, const REAL vzL,
//const REAL betaxL, const REAL betayL, const REAL betazL, const REAL alpL,
const REAL sqrtg,REAL *StildeD0L, REAL *StildeD1L, REAL *StildeD2L) {
//const REAL fourpialpha_inv = 1.0/( 4.0*M_PI*(METRIC[LAPM1] + 1.0) );
const REAL fourpi_inv = 1.0/( 4.0*M_PI );
const REAL B2 = gxxL*BxL*BxL + gyyL*ByL*ByL + gzzL*BzL*BzL
+ 2.0*(gxyL*BxL*ByL + gxzL*BxL*BzL + gyzL*ByL*BzL);
// NOTE: SIGNIFICANTLY MODIFIED FROM ILLINOISGRMHD VERSION:
// velocities in GiRaFFE are defined to be "drift" velocity.
// cf. Eqs 47 and 85 in http://arxiv.org/pdf/1310.3274.pdf
// Modified again from the original GiRaFFE to use Valencia velocity
const REAL v_xL = gxxL*vxL + gxyL*vyL + gxzL*vzL;
const REAL v_yL = gxyL*vxL + gyyL*vyL + gyzL*vzL;
const REAL v_zL = gxzL*vxL + gyzL*vyL + gzzL*vzL;
/*
* Comments:
* Eq. 85 in https://arxiv.org/pdf/1310.3274.pdf:
* v^i = 4 pi alpha * (gamma^{ij} tilde{S}_j) / (sqrtgamma * B^2) - beta^i
* which implies that
* (v^i + beta^i)*(sqrtgamma * B^2)/(4 pi alpha) = gamma^{ij} tilde{S}_j
* Multiply both sides by gamma_{ik}:
* gamma_{ik} (v^i + beta^i)*(sqrtgamma * B^2)/(4 pi alpha) = gamma_{ik} gamma^{ij} tilde{S}_j
*
* -> tilde{S}_k = gamma_{ik} (v^i + beta^i)*(sqrtgamma * B^2)/(4 pi alpha)
*/
*StildeD0L = v_xL * sqrtg * B2 * fourpi_inv;
*StildeD1L = v_yL * sqrtg * B2 * fourpi_inv;
*StildeD2L = v_zL * sqrtg * B2 * fourpi_inv;
}
# <a id='code_validation'></a>
#
# # Step 3: Code Validation against original C code \[Back to [top](#toc)\]
# $$\label{code_validation}$$
#
# To validate the code in this tutorial we check for agreement between the files
#
# 1. that were written in this tutorial and
# 1. those that are stored in `GiRaFFE_HO/GiRaFFE_Ccode_library`
#
# +
import difflib
import sys
# Define the directory that we wish to validate against:
valdir = "GiRaFFE_HO/GiRaFFE_Ccode_library/"
print("Printing difference between original C code and this code...")
# Open the files to compare
files_to_check = ["driver_conserv_to_prims_FFE.C","compute_conservatives_FFE.C"]
for file in files_to_check:
print("Checking file " + file)
with open(os.path.join(valdir+file)) as file1, open(os.path.join(outdir+file)) as file2:
# Read the lines of each file
file1_lines = file1.readlines()
file2_lines = file2.readlines()
num_diffs = 0
for line in difflib.unified_diff(file1_lines, file2_lines, fromfile=os.path.join(valdir+file), tofile=os.path.join(outdir+file)):
sys.stdout.writelines(line)
num_diffs = num_diffs + 1
if num_diffs == 0:
print("No difference. TEST PASSED!")
else:
print("ERROR: Disagreement found with .py file. See differences above.")
# -
# <a id='latex_pdf_output'></a>
#
# # Step 4: Output this notebook to $\LaTeX$-formatted PDF file \[Back to [top](#toc)\]
# $$\label{latex_pdf_output}$$
#
# The following code cell converts this Jupyter notebook into a proper, clickable $\LaTeX$-formatted PDF file. After the cell is successfully run, the generated PDF may be found in the root NRPy+ tutorial directory, with filename
# [Tutorial-GiRaFFE_HO_C_code_library-C2P_P2C.pdf](Tutorial-GiRaFFE_HO_C_code_library-C2P_P2C.pdf) (Note that clicking on this link may not work; you may need to open the PDF file through another means.)
# !jupyter nbconvert --to latex --template latex_nrpy_style.tplx Tutorial-GiRaFFE_HO_C_code_library-C2P_P2C.ipynb
# !pdflatex -interaction=batchmode Tutorial-GiRaFFE_HO_C_code_library-C2P_P2C.tex
# !pdflatex -interaction=batchmode Tutorial-GiRaFFE_HO_C_code_library-C2P_P2C.tex
# !pdflatex -interaction=batchmode Tutorial-GiRaFFE_HO_C_code_library-C2P_P2C.tex
# !rm -f Tut*.out Tut*.aux Tut*.log
| notebook/Tutorial-GiRaFFE_HO_C_code_library-C2P_P2C.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import pandas
import re
import numpy as np
from random import randint
tract = pandas.read_csv("ACS_Median_HH_Income.csv")
# +
#### Extracting float values from cencus tract cells and converting them into similar household format
#### TypeError will keep coming up but won't effect output table (filled with NA)
for x in range(len(tract.all_tracts)):
tract.all_tracts[x] = float(re.sub(r'[^\d.]+', '', tract.all_tracts[x])) * 100
for i in range(len(tract.Hhsize_tracts)):
tract.Hhsize_tracts[i] = float(re.sub(r'[^\d.]+', '', tract.Hhsize_tracts[i])) * 100
# +
#### Genereated formatted cs data
tract.to_csv('NewIncome.csv')
#### This 'NewIncome' csv should be seperated into two seperate tables, HouseholdMedians.csv and TractMedians.csv if there are medians based off of household size.
# +
#### Requires three data tables (or two if no hhsize medians)
#### generated_household.csv: The generated household table from doppelganger, this should be the largest file out of the three
#### TractMedians.csv: ACS household median incomes based off of tract
#### HouseholdMedians.csv: ACS household median incomes based off of tract and household size
hh = pandas.read_csv("generated_households.csv")
hh_median = pandas.read_csv("HouseholdMedians.csv")
tract_median = pandas.read_csv("TractMedians.csv")
# +
#### Bulk of the acs median income replacements happen here.
for x in range(hh_median['Hhsize_tracts'].shape[0]):
hh.loc[ (hh.household_income=='<=0') & (hh['tract']==hh_median['Hhsize_tracts'][x]) & (hh.num_people=='1'),'household_income'] = hh_median['h1'][x]
hh.loc[ (hh.household_income=='<=0') & (hh['tract']==hh_median['Hhsize_tracts'][x]) & (hh.num_people=='2'),'household_income'] = hh_median['h2'][x]
hh.loc[ (hh.household_income=='<=0') & (hh['tract']==hh_median['Hhsize_tracts'][x]) & (hh.num_people=='3'),'household_income'] = hh_median['h3'][x]
hh.loc[ (hh.household_income=='<=0') & (hh['tract']==hh_median['Hhsize_tracts'][x]) & (hh.num_people=='4+'),'household_income'] = hh_median['h4'][x]
#### The following loop should be used it you only have median incomes by tract and not household size.
for x in range(tract_median['all_tracts'].shape[0]):
hh.loc[ (hh.household_income=='<=0') & (hh.tract==tract_median['all_tracts'][x]),'household_income'] = tract_median['Income'][x]
# +
#### .loc can be used for any salary that doesn't need randint()
hh.loc[ (hh.household_income=='100000+'),'household_income'] = 100001
# +
## Dataframe.loc method for replacing income ranges is much faster than the below looping method but gives the same random value for each range.
## hh.loc[ (hh.household_income=='0-10000'),'household_income'] = randint(0,10000)
# +
#### Run time ~30min. Very inefficient and could use updating (excel macro is much faster) but this does work!
# VB Macro for this process is located in this folder as 'IncomeRangeReplacement'
for x in range(hh['household_income'].shape[0]):
if hh.household_income[x]=='0-10000':
hh.household_income[x]=randint(0,10000)
elif hh.household_income[x]=='10000-20000':
hh.household_income[x]=randint(10000,20000)
elif hh.household_income[x]=='20000-30000':
hh.household_income[x]=randint(20000,30000)
elif hh.household_income[x]=='30000-40000':
hh.household_income[x]=randint(30000,40000)
elif hh.household_income[x]=='40000-50000':
hh.household_income[x]=randint(40000,50000)
elif hh.household_income[x]=='50000-60000':
hh.household_income[x]=randint(50000,60000)
elif hh.household_income[x]=='60000-70000':
hh.household_income[x]=randint(60000,70000)
elif hh.household_income[x]=='70000-80000':
hh.household_income[x]=randint(70000,80000)
elif hh.household_income[x]=='80000-90000':
hh.household_income[x]=randint(80000,90000)
elif hh.household_income[x]=='90000-100000':
hh.household_income[x]=randint(90000,100000)
# +
#### Finished updated table exported as csv
#### This csv file will be used in the household transition model. This specific file should be merged into the data folder in urbansim
hh.to_csv('households.csv')
| data/Household_Data_Process/ReplacingIncome.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
train_data.info()
test_data.info()
train_data
train_data['Survived'].value_counts()
train_data['Pclass'].value_counts()
train_data['Sex'].value_counts()
# + active=""
# train_data['SibSp'].value_counts()
# + active=""
# train_data['Parch'].value_counts()
# -
train_data['Embarked'].value_counts()
# + active=""
# train_data['CabinClass'].value_counts()
# -
sns.distplot(train_data['Age'], kde=False, rug=True);
sns.distplot(train_data['Fare'], kde=False, rug=True);
| p-titanic/titanic_visualize.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## This notebook is for the woodscape challenge
import torch, torchvision
import mmseg
from mmseg.apis import inference_segmentor, init_segmentor, show_result_pyplot
from mmseg.core.evaluation import get_palette
# +
import mmcv
import matplotlib.pyplot as plt
data_dir = "/media/user/Seagate Expansion Drive/rectified_images/dataset_rectified"
img_dir = 'images'
ann_dir = 'labels'
img = mmcv.imread(data_dir+'/images/00004_FVL.png')
plt.figure(figsize=(8, 6))
plt.imshow(mmcv.bgr2rgb(img))
plt.show()
# +
import matplotlib.patches as mpatches
from PIL import Image
import numpy as np
classes = ('void','road', 'lanemarks', 'curb', 'pedestrians', 'rider', 'vehicles', 'bicycle', 'motorcycle', 'traffic_sign')
palette = [[128, 128, 128], [129, 127, 38], [120, 69, 125], [53, 125, 34], [120, 134, 255],
[0, 11, 123], [118, 20, 12], [122, 81, 25], [241, 134, 51], [0, 125, 151]]
img = np.array(Image.open(data_dir+'/labels/00004_FVL.png'))
seg_img = Image.fromarray(img[:,:,1]).convert('P')
seg_img.putpalette(np.array(palette, dtype=np.uint8))
plt.figure(figsize=(8, 6))
plt.imshow(seg_img)
# create a patch (proxy artist) for every color
patches = [mpatches.Patch(color=np.array(palette[i])/255.,
label=classes[i]) for i in range(10)]
# put those patched as legend-handles into the legend
plt.legend(handles=patches, bbox_to_anchor=(1.05, 1), loc=2, borderaxespad=0.,
fontsize='large')
plt.show()
# +
from mmseg.datasets.builder import DATASETS
from mmseg.datasets.custom import CustomDataset
@DATASETS.register_module()
class Woodscape(CustomDataset):
CLASSES = classes
PALETTE = palette
def __init__(self, split, **kwargs):
super().__init__(img_suffix='.png', seg_map_suffix='_.png',
split=split, **kwargs)
assert osp.exists(self.img_dir) and self.split is not None
# -
from mmcv import Config
cfg = Config.fromfile('../configs/pspnet/pspnet_r50-d8_512x1024_40k_cityscapes.py')
# +
from mmseg.apis import set_random_seed
# Since we use ony one GPU, BN is used instead of SyncBN
cfg.norm_cfg = dict(type='BN', requires_grad=True)
cfg.model.backbone.norm_cfg = cfg.norm_cfg
cfg.model.decode_head.norm_cfg = cfg.norm_cfg
cfg.model.auxiliary_head.norm_cfg = cfg.norm_cfg
# modify num classes of the model in decode/auxiliary head
cfg.model.decode_head.num_classes = 11
cfg.model.auxiliary_head.num_classes = 11
# Modify dataset type and path
cfg.dataset_type = 'Woodscape'
cfg.data_root = data_dir
cfg.data.samples_per_gpu = 8
cfg.data.workers_per_gpu = 8
cfg.img_norm_cfg = dict(
mean=[123.675, 116.28, 103.53], std=[58.395, 57.12, 57.375], to_rgb=True)
cfg.crop_size = (256, 256)
cfg.train_pipeline = [
dict(type='LoadImageFromFile'),
dict(type='LoadAnnotations'),
dict(type='Resize', img_scale=(320, 240), ratio_range=(0.5, 2.0)),
dict(type='RandomCrop', crop_size=cfg.crop_size, cat_max_ratio=0.75),
dict(type='RandomFlip', flip_ratio=0.5),
dict(type='PhotoMetricDistortion'),
dict(type='Normalize', **cfg.img_norm_cfg),
dict(type='Pad', size=cfg.crop_size, pad_val=0, seg_pad_val=255),
dict(type='DefaultFormatBundle'),
dict(type='Collect', keys=['img', 'gt_semantic_seg']),
]
cfg.test_pipeline = [
dict(type='LoadImageFromFile'),
dict(
type='MultiScaleFlipAug',
img_scale=(320, 240),
# img_ratios=[0.5, 0.75, 1.0, 1.25, 1.5, 1.75],
flip=False,
transforms=[
dict(type='Resize', keep_ratio=True),
dict(type='RandomFlip'),
dict(type='Normalize', **cfg.img_norm_cfg),
dict(type='ImageToTensor', keys=['img']),
dict(type='Collect', keys=['img']),
])
]
cfg.data.train.type = cfg.dataset_type
cfg.data.train.data_root = cfg.data_root
cfg.data.train.img_dir = img_dir
cfg.data.train.ann_dir = ann_dir
cfg.data.train.pipeline = cfg.train_pipeline
cfg.data.train.split = 'splits/train.txt'
cfg.data.val.type = cfg.dataset_type
cfg.data.val.data_root = cfg.data_root
cfg.data.val.img_dir = img_dir
cfg.data.val.ann_dir = ann_dir
cfg.data.val.pipeline = cfg.test_pipeline
cfg.data.val.split = 'splits/val.txt'
cfg.data.test.type = cfg.dataset_type
cfg.data.test.data_root = cfg.data_root
cfg.data.test.img_dir = img_dir
cfg.data.test.ann_dir = ann_dir
cfg.data.test.pipeline = cfg.test_pipeline
cfg.data.test.split = 'splits/val.txt'
# We can still use the pre-trained Mask RCNN model though we do not need to
# use the mask branch
cfg.load_from = 'checkpoints/pspnet_r50-d8_512x1024_40k_cityscapes_20200605_003338-2966598c.pth'
# Set up working dir to save files and logs.
cfg.work_dir = './work_dirs/tutorial'
cfg.runner.max_iters = 200
cfg.log_config.interval = 10
cfg.evaluation.interval = 200
cfg.checkpoint_config.interval = 200
# Set seed to facitate reproducing the result
cfg.seed = 0
set_random_seed(0, deterministic=False)
cfg.gpu_ids = range(1)
# Let's have a look at the final config used for training
print(f'Config:\n{cfg.pretty_text}')
# +
from mmseg.datasets import build_dataset
from mmseg.models import build_segmentor
from mmseg.apis import train_segmentor
import os.path as osp
from ipywidgets import IntProgress
# Build the dataset
datasets = [build_dataset(cfg.data.train)]
# Build the detector
model = build_segmentor(
cfg.model, train_cfg=cfg.get('train_cfg'), test_cfg=cfg.get('test_cfg'))
# Add an attribute for visualization convenience
model.CLASSES = datasets[0].CLASSES
print(model)
# -
# Create work_dir
mmcv.mkdir_or_exist(osp.abspath(cfg.work_dir))
train_segmentor(model, datasets, cfg, distributed=False, validate=False,
meta=dict())
img = mmcv.imread(data_dir+'/images/00004_FVL.png')
model.cfg = cfg
result = inference_segmentor(model, img)
plt.figure(figsize=(8, 6))
show_result_pyplot(model, img, result, palette)
| demo/Woodscape.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Q#
# language: qsharp
# name: iqsharp
# ---
# # Multi-Qubit Systems Workbook
#
# **What is this workbook?**
# A workbook is a collection of problems, accompanied by solutions to them.
# The explanations focus on the logical steps required to solve a problem; they illustrate the concepts that need to be applied to come up with a solution to the problem, explaining the mathematical steps required.
#
# Note that a workbook should not be the primary source of knowledge on the subject matter; it assumes that you've already read a tutorial or a textbook and that you are now seeking to improve your problem-solving skills. You should attempt solving the tasks of the respective kata first, and turn to the workbook only if stuck. While a textbook emphasizes knowledge acquisition, a workbook emphasizes skill acquisition.
#
# This workbook describes the solutions to the problems offered in the [Multi-Quibt Systems tutorial](./MultiQubitSystems.ipynb).
#
#
# **What you should know for this workbook**
#
# 1. Basic single-qubit gates.
# 2. The concept of tensor product.
# To begin, first prepare this notebook for execution (if you skip this step, you'll get "Syntax does not match any known patterns" error when you try to execute Q# code in the next cells):
%package Microsoft.Quantum.Katas::0.11.2006.403
# > The package versions in the output of the cell above should always match. If you are running the Notebooks locally and the versions do not match, please install the IQ# version that matches the version of the `Microsoft.Quantum.Katas` package.
# > <details>
# > <summary><u>How to install the right IQ# version</u></summary>
# > For example, if the version of `Microsoft.Quantum.Katas` package above is 0.1.2.3, the installation steps are as follows:
# >
# > 1. Stop the kernel.
# > 2. Uninstall the existing version of IQ#:
# > dotnet tool uninstall microsoft.quantum.iqsharp -g
# > 3. Install the matching version:
# > dotnet tool install microsoft.quantum.iqsharp -g --version 0.1.2.3
# > 4. Reinstall the kernel:
# > dotnet iqsharp install
# > 5. Restart the Notebook.
# > </details>
#
# For an overview of all the gates have a look at the Quickref [here](../../quickref/qsharp-quick-reference.pdf).
# ### <span style="color:blue">Exercise 1</span>: Show that the state is separable
#
# $$\frac{1}{2} \begin{bmatrix} 1 \\ i \\ -i \\ 1 \end{bmatrix} =
# \begin{bmatrix} ? \\ ? \end{bmatrix} \otimes \begin{bmatrix} ? \\ ? \end{bmatrix}$$
#
# ### Solution
#
# To separate the state into a tensor product of two single-qubit states, we need to represent it in the following way:
#
# $$\begin{bmatrix} \alpha \color{red}\gamma \\ \alpha \color{red}\delta \\ \beta \color{red}\gamma \\ \beta \color{red}\delta \end{bmatrix} = \begin{bmatrix} \alpha \\ \beta \end{bmatrix} \otimes \begin{bmatrix} \color{red}\gamma \\ \color{red}\delta \end{bmatrix}$$
#
# This brings us to a system of equations:
#
# $$\begin{cases}
# \alpha\gamma = \frac{1}{2} \\
# \alpha\delta = \frac{i}{2} \\
# \beta \gamma = \frac{-i}{2} \\
# \beta \delta = \frac{1}{2} \\
# \end{cases}$$
#
# Solving this system of equations gives us the answer:
#
# $$\alpha = \frac{1}{\sqrt2}, \beta = \frac{-i}{\sqrt2}, \gamma = \frac{1}{\sqrt2}, \delta = \frac{i}{\sqrt2}$$
#
# $$\frac{1}{2} \begin{bmatrix} 1 \\ i \\ -i \\ 1 \end{bmatrix} = \frac{1}{\sqrt2}
# \begin{bmatrix} 1 \\ -i \end{bmatrix} \otimes \frac{1}{\sqrt2} \begin{bmatrix} 1 \\ i \end{bmatrix}$$
# [Return to task 1 of the Multi-Qubit Systems tutorial.](./MultiQubitSystems.ipynb#Exercise-1:-Show-that-the-state-is-separable)
# ### <span style="color:blue">Exercise 2</span>: Is this state separable?
#
# $$\frac{1}{\sqrt{2}}\begin{bmatrix} 1 \\ 0 \\ 0 \\ 1 \end{bmatrix}$$
#
# ### Solution
#
# Let's assume that this state is separable and write down the system of equations to determine the coefficients of individual qubit states in the tensor product, similar to what we did in the previous exercise:
#
# $$\begin{cases}
# \alpha\gamma = \frac{1}{\sqrt2} \\
# \alpha\delta = 0 \\
# \beta \gamma = 0 \\
# \beta \delta = \frac{1}{\sqrt2} \\
# \end{cases}$$
#
# Now let's multiply the first and the last equations, and the second and the third equations:
#
# $$\begin{cases}
# \alpha\beta\gamma\delta = \frac{1}{2} \\
# \alpha\beta\gamma\delta = 0
# \end{cases}$$
#
# We can see that this system of equations doesn't have a solution, which means that this state is **not separable**.
# [Return to task 2 of the Multi-Qubit Systems tutorial.](./MultiQubitSystems.ipynb#Exercise-2:-Is-this-state-separable?)
# ### <span style="color:blue">Exercise 3</span>: Prepare a basis state
#
# **Input:** A two-qubit system in the basis state $|00\rangle = \begin{bmatrix} 1 \\ 0 \\ 0 \\ 0 \end{bmatrix}$.
#
# **Goal:** Transform the system into the basis state $|11\rangle = \begin{bmatrix} 0 \\ 0 \\ 0 \\ 1 \end{bmatrix}$.
# ### Solution
#
# The starting state can be represented as follows:
# $$ \begin{bmatrix} 1 \\ 0 \\ 0 \\ 0 \end{bmatrix} = |0\rangle \otimes |0\rangle $$
#
# The goal state can be represented as follows:
# $$ \begin{bmatrix} 0 \\ 0 \\ 0 \\ 1 \end{bmatrix} = |1\rangle \otimes |1\rangle $$
#
# Applying an **X** gate to a qubit in the $|0\rangle$ state transforms the qubit state into the $|1\rangle$ state. So, if we apply the **X** gate on the first qubit and the second qubit, we get the desired state.
# +
%kata T1_PrepareState1_Test
operation PrepareState1 (qs : Qubit[]) : Unit is Adj+Ctl {
X(qs[0]);
X(qs[1]);
}
# -
# [Return to task 3 of the Multi-Qubit Systems tutorial.](./MultiQubitSystems.ipynb#Exercise-3:-Prepare-a-basis-state)
# ### <span style="color:blue">Exercise 4</span>: Prepare a superposition of two basis states
#
# **Input:** A two-qubit system in the basis state $|00\rangle = \begin{bmatrix} 1 \\ 0 \\ 0 \\ 0 \end{bmatrix}$.
#
# **Goal:** Transform the system into the state $\frac{1}{\sqrt2}\big(|00\rangle - |01\rangle\big) = \frac{1}{\sqrt2}\begin{bmatrix} 1 \\ -1 \\ 0 \\ 0 \end{bmatrix}$.
# ### Solution
#
# We begin in the same state as the previous excercise:
# $$ \begin{bmatrix} 1 \\ 0 \\ 0 \\ 0 \end{bmatrix} = \begin{bmatrix} 1 \\ 0 \end{bmatrix} \otimes \begin{bmatrix} 1 \\ 0 \end{bmatrix} = |0\rangle \otimes |0\rangle$$
#
# The goal state can be separated as follows:
# $$ \frac{1}{\sqrt2} \begin{bmatrix} 1 \\ -1 \\ 0 \\ 0 \end{bmatrix} = \begin{bmatrix} 1 \\ 0 \end{bmatrix} \otimes \frac{1}{\sqrt2}\begin{bmatrix} 1 \\ -1 \end{bmatrix} = |0\rangle \otimes \frac{1}{\sqrt2}\big(|0\rangle - |1\rangle\big)$$
#
# This means that the first qubit is already in the state we want it to be, but the second qubit needs to be transformed from the $ \begin{bmatrix} 1 \\ 0 \end{bmatrix} $ into $ \frac{1}{\sqrt{2}}\begin{bmatrix} 1 \\ -1\end{bmatrix}$ state.
#
# First, we apply the **X** gate to the second qubit; this performs the following transformation:
# $$ X |0\rangle = \begin{bmatrix}0 & 1 \\ 1 & 0 \end{bmatrix} \cdot \begin{bmatrix}1 \\ 0 \end{bmatrix} = \begin{bmatrix} 0 \\ 1 \end{bmatrix} = |1\rangle $$
#
# Second, we apply the **H** gate to the second qubit; this transforms its state into the desired one:
# $$ H|1\rangle = \frac{1}{\sqrt2}\begin{bmatrix} 1 & 1 \\ 1 & -1 \end{bmatrix} \cdot \begin{bmatrix} 0 \\ 1 \end{bmatrix} = \frac{1}{\sqrt2}\begin{bmatrix} 1 \\ -1 \end{bmatrix}$$
# +
%kata T2_PrepareState2_Test
operation PrepareState2 (qs : Qubit[]) : Unit is Adj+Ctl {
X(qs[1]);
H(qs[1]);
}
# -
# [Return to task 4 of the Multi-Qubit Systems tutorial.](./MultiQubitSystems.ipynb#Exercise-4:-Prepare-a-superposition-of-two-basis-states)
# ### <span style="color:blue">Exercise 5</span>: Prepare a superposition with real amplitudes
#
# **Input:** A two-qubit system in the basis state $|00\rangle = \begin{bmatrix} 1 \\ 0 \\ 0 \\ 0 \end{bmatrix}$.
#
# **Goal:** Transform the system into the state $\frac{1}{2}\big(|00\rangle - |01\rangle + |10\rangle - |11\rangle\big) = \frac{1}{2}\begin{bmatrix} 1 \\ -1 \\ 1 \\ -1 \end{bmatrix}$.
# ### Solution
#
# Again, to start we will represent the goal state as a tensor product of single-qubit states; this gives us the following representation:
#
# $$ \frac{1}{2}\big(|00\rangle - |01\rangle + |10\rangle - |11\rangle\big) = \frac{1}{2}\begin{bmatrix} 1 \\ -1 \\ 1 \\ -1 \end{bmatrix} = \frac{1}{\sqrt2} \begin{bmatrix} 1 \\ 1 \end{bmatrix} \otimes \frac{1}{\sqrt2}\begin{bmatrix} 1 \\ -1 \end{bmatrix} = \frac{1}{\sqrt2}\big(|0\rangle + |1\rangle\big) \otimes \frac{1}{\sqrt2}\big(|0\rangle - |1\rangle\big) $$
#
# This time we need to transform both the first and the second qubits. Let's start with the first qubit. Applying the **H** gate transforms its state as follows:
#
# $$ H|0\rangle = \frac{1}{\sqrt2}\begin{bmatrix} 1 & 1 \\ 1 & -1 \end{bmatrix} \cdot \begin{bmatrix} 1 \\ 0 \end{bmatrix} = \frac{1}{\sqrt2} \begin{bmatrix} 1 \\ 1 \end{bmatrix} = \frac{1}{\sqrt2}\big(|0\rangle + |1\rangle\big)$$
#
# For the second qubit we can use the same transformation we've seen in [exercise 4](#Exercise-4:-Prepare-a-superposition-of-two-basis-states); this will give the desired end state.
# +
%kata T3_PrepareState3_Test
operation PrepareState3 (qs : Qubit[]) : Unit is Adj+Ctl {
H(qs[0]);
X(qs[1]);
H(qs[1]);
}
# -
# [Return to task 5 of the Multi-Qubit Systems tutorial.](./MultiQubitSystems.ipynb#Exercise-5:-Prepare-a-superposition-with-real-amplitudes)
# ### <span style="color:blue">Exercise 6</span>: Prepare a superposition with complex amplitudes
#
# **Input:** A two-qubit system in the basis state $|00\rangle = \begin{bmatrix} 1 \\ 0 \\ 0 \\ 0 \end{bmatrix}$.
#
# **Goal:** Transform the system into the state $\frac{1}{2}\big(|00\rangle + e^{i\pi/4}|01\rangle + e^{i\pi/2}|10\rangle + e^{3i\pi/4}|11\rangle\big) = \frac{1}{2}\begin{bmatrix} 1 \\ e^{i\pi/4} \\ e^{i\pi/2} \\ e^{3i\pi/4} \end{bmatrix}$.
# ### Solution
#
# The start state is the same as the previous exercises:
# $$ \begin{bmatrix} 1 \\ 0 \\ 0 \\ 0 \end{bmatrix} = \begin{bmatrix} 1 \\ 0 \end{bmatrix} \otimes \begin{bmatrix} 1 \\ 0 \end{bmatrix} = |0\rangle \otimes |0\rangle $$
#
# The goal state, factored as a tensor product, looks like this (remember that $e^{3i\pi/4} = e^{i\pi/4} e^{i\pi/2}$):
#
# $$ \frac{1}{2}\begin{bmatrix} 1 \\ e^{i\pi/4} \\ e^{i\pi/2} \\ e^{3i\pi/4} \end{bmatrix}
# = \frac{1}{\sqrt2} \begin{bmatrix} 1 \\ e^{i\pi/2} \end{bmatrix} \otimes \frac{1}{\sqrt2}\begin{bmatrix} 1 \\ e^{i\pi/4} \end{bmatrix}
# = \frac{1}{\sqrt2}\big(|0\rangle + e^{i\pi/2}|1\rangle\big) \otimes \frac{1}{\sqrt2}\big(|0\rangle + e^{i\pi/4}|1\rangle\big) $$
#
# We will again need to adjust the states of both qubits independently.
#
# For the first qubit, we'll start by applying the **H** gate, getting the state $\frac{1}{\sqrt2} \begin{bmatrix} 1 \\ 1 \end{bmatrix}$, as we've seen in the previous task. Afterwards we'll apply the **S** gate with the following result:
#
# $$ \begin{bmatrix} 1 & 0 \\ 0 & i \end{bmatrix} \cdot \frac{1}{\sqrt2} \begin{bmatrix} 1 \\ 1 \end{bmatrix} = \frac{1}{\sqrt2} \begin{bmatrix} 1 \\ i \end{bmatrix}$$
#
# If we recall that $i = e^{i\pi/2}$, we can write the final state of the second qubit as:
# $$ \frac{1}{\sqrt2} \begin{bmatrix} 1 \\ e^{i\pi/2} \end{bmatrix} $$
#
# For the second qubit. we'll apply the **H** gate, followed by the **T** gate, with the following result:
# $$ \begin{bmatrix} 1 & 0 \\ 0 & e^{i\pi/4} \end{bmatrix} \cdot \frac{1}{\sqrt2} \begin{bmatrix} 1 \\ 1 \end{bmatrix} = \frac{1}{\sqrt2} \begin{bmatrix} 1 \\ e^{i\pi/4} \end{bmatrix} $$
# +
%kata T4_PrepareState4_Test
operation PrepareState4 (qs : Qubit[]) : Unit is Adj+Ctl {
H(qs[0]);
S(qs[0]);
H(qs[1]);
T(qs[1]);
}
# -
# [Return to task 6 of the Multi-Qubit Systems tutorial.](./MultiQubitSystems.ipynb#Exercise-6:-Prepare-a-superposition-with-complex-amplitudes)
# ## Conclusion
#
# As you've seen in the exercises, you can prepare separable multi-qubit states using only single-qubit gates.
# However, to prepare and manipulate entangled states you'll need more powerful tools.
# In the [next tutorial](../MultiQubitGates/MultiQubitGates.ipynb) you will learn about multi-qubit gates which give you access to all states of multi-qubit systems.
| tutorials/MultiQubitSystems/Workbook_MultiQubitSystems.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.7 ('myPyEnv')
# language: python
# name: python3
# ---
# ## Task
# * take password length as input (length must be between 6 & 12)
# * generate random password having digits, special characters, upper and lower case characters
#
# ###### Core Concepts:
# * lambda (anonymous) function
# * recursion
#
# ###### Used Modules:
# * string
# * random
#
# #### Data structure:
# * list
import string as st, random as r
# length of password
x= lambda length:length if length<=12 and length>=6 else x(int(input("Enter length between 6 & 12")))
length=x(int(input("Enter length of password")))
# dataset
dataset=[st.punctuation,st.ascii_letters, st.digits]
dataset=''.join(dataset)
# generate randomly from dataset
passwordList=r.sample(dataset[:],length)
password="".join(passwordList)
password
| Password Generator.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Q Learning 介绍
# 在增强学习中,有一种很有名的算法,叫做 q-learning,我们下面会从原理入手,然后通过一个简单的小例子讲一讲 q-learning。
#
# ## q-learning 的原理
# 我们使用一个简单的例子来导入 q-learning,假设一个屋子有 5 个房间,某一些房间之间相连,我们希望能够走出这个房间,示意图如下
#
# 
# 那么我们可以将其简化成一些节点和图的形式,每个房间作为一个节点,两个房间有门相连,就在两个节点之间连接一条线,可以得到下面的图片
#
# 
# 为了模拟整个过程,我们放置一个智能体在任何一个房间,希望它能够走出这个房间,也就是说希望其能够走到了 5 号节点。为了能够让智能体知道 5 号节点是目标房间,我们需要设置一些奖励,对于每一条边,我们都关联一个奖励值:直接连到目标房间的边的奖励值设置为 100,其他的边可以设置为 0,注意 5 号房间有一个指向自己的箭头,奖励值也设置为 100,其他直接指向 5 号房间的也设置为 100,这样当智能体到达 5 号房间之后,他就会选择一只待在 5 号房间,这也称为吸收目标,效果如下
#
# 
# 想想一下智能体可以不断学习,每次我们将其放在其中一个房间,然后它可以不断探索,根据奖励值走到 5 号房间,也就是走出这个屋子。比如现在这个智能体在 2 号房间,我们希望其能够不断探索走到 5 号房间。
#
# ### 状态和动作
# q-learning 中有两个重要的概念,一个是状态,一个是动作,我们将每一个房间都称为一个状态,而智能体从一个房间走到另外一个房间称为一个动作,对应于上面的图就是每个节点是一个状态,每一个箭头都是一种行动。假如智能体处在状态 4,从状态 4 其可以选择走到状态 0,或者状态 3 或者状态 5,如果其走到了状态 3,也可以选择走到状态 2 或者状态 1 或者 状态 4。
#
# 我们可以根据状态和动作得到的奖励来建立一个奖励表,用 -1 表示相应节点之间没有边相连,而没有到达终点的边奖励都记为 0,如下
#
# 
# 类似的,我们可以让智能体通过和环境的交互来不断学习环境中的知识,让智能体根据每个状态来估计每种行动可能得到的收益,这个矩阵被称为 Q 表,每一行表示状态,每一列表示不同的动作,对于状态未知的情景,我们可以随机让智能体从任何的位置出发,然后去探索新的环境来尽可能的得到所有的状态。刚开始智能体对于环境一无所知,所以数值全部初始化为 0,如下
#
# 
#
# 我们的智能体通过不断地学习来更新 Q 表中的结果,最后依据 Q 表中的值来做决策。
# ### Q-learning 算法
# 有了奖励表和 Q 表,我们需要知道智能体是如何通过学习来更新 Q 表,以便最后能够根据 Q 表进行决策,这个时候就需要讲一讲 Q-learning 的算法。
#
# Q-learning 的算法特别简单,状态转移公式如下
#
# $$Q(s, a) = R(s, a) + \gamma \mathop{max}_{\tilde{a}}\{ Q(\tilde{s}, \tilde{a}) \}$$
#
# 其中 s, a 表示当前的状态和行动,$\tilde{s}, \tilde{a}$ 分别表示 s 采取 a 的动作之后的下一个状态和该状态对应所有的行动,参数 $\gamma$ 是一个常数,$0 \leq \gamma \le 1 $表示对未来奖励的一个衰减程度,形象地比喻就是一个人对于未来的远见程度。
#
# 解释一下就是智能体通过经验进行自主学习,不断从一个状态转移到另外一个状态进行探索,并在这个过程中不断更新 Q 表,直到到达目标位置,Q 表就像智能体的大脑,更新越多就越强。我们称智能体的每一次探索为 episode,每个 episode 都表示智能体从任意初始状态到达目标状态,当智能体到达一个目标状态,那么当前的 episode 结束,进入下一个 episode。
# 下面给出 q-learning 的整个算法流程
# - step1 给定参数 $\gamma$ 和奖励矩阵 R
# - step2 令 Q:= 0
# - step3 For each episode:
# - 3.1 随机选择一个初始状态 s
# - 3.2 若未到达目标状态,则执行以下几步
# - (1)在当前状态 s 的所有可能行动中选取一个行为 a
# - (2)利用选定的行为 a,得到下一个状态 $\tilde{s}$
# - (3)按照前面的转移公式计算 Q(s, a)
# - (4)令 $s: = \tilde{s}$
# ### 单步演示
# 为了更好地理解 q-learning,我们可以示例其中一步。
#
# 首先选择 $\gamma = 0.8$,初始状态为 1,Q 初始化为零矩阵
#
# 
#
# 
#
# 因为是状态 1,所以我们观察 R 矩阵的第二行,负数表示非法行为,所以下一个状态只有两种可能,走到状态 3 或者走到状态 5,随机地,我们可以选择走到状态 5。
#
# 当我们走到状态 5 之后,会发生什么事情呢?观察 R 矩阵的第 6 行可以发现,其对应于三个可能采取的动作:转至状态 1,4 或者 5,根据上面的转移公式,我们有
#
# $$Q(1, 5) = R(1, 5) + 0.8 * max\{Q(5, 1), Q(5, 4), Q(5, 5)\} = 100 + 0.8 * max\{0, 0, 0\} = 100$$
#
# 所以现在 Q 矩阵进行了更新,变为了
#
# 
#
# 现在我们的状态由 1 变成了 5,因为 5 是最终的目标状态,所以一次 episode 便完成了,进入下一个 episode。
#
# 在下一个 episode 中又随机选择一个初始状态开始,不断更新 Q 矩阵,在经过了很多个 episode 之后,矩阵 Q 接近收敛,那么我们的智能体就学会了从任意状态转移到目标状态的最优路径。
# 从上面的原理,我们知道了 q-learning 最重要的状态转移公式,这个公式也叫做 Bellman Equation,通过这个公式我们能够不断地进行更新 Q 矩阵,最后得到一个收敛的 Q 矩阵。
#
# 下面我们通过代码来实现这个过程
#
# 我们定义一个简单的走迷宫过程,也就是
#
# 
# 初始位置随机在 state 0, state 1 和 state 2 上,然后希望智能体能够走到 state 3 获得宝藏,上面可行的行动路线已经用箭头标注了
import numpy as np
import random
# 下面定义奖励矩阵,一共是 4 行,5 列,每一行分别表示 state 0 到 state 3 这四个状态,每一列分别表示上下左右和静止 5 种状态,奖励矩阵中的 0 表示不可行的路线,比如第一个行,上走和左走都是不可行的路线,都用 0 表示,向下走会走到陷阱,所以使用 -10 表示奖励,向右走和静止都给与 -1 的奖励,因为既没有触发陷阱,也没有到达宝藏,但是过程中浪费了时间。
reward = np.array([[0, -10, 0, -1, -1],
[0, 10, -1, 0, -1],
[-1, 0, 0, 10, -10],
[-1, 0, -10, 0, 10]])
# 接下来定义一个初始化为 0 的 q 矩阵
q_matrix = np.zeros((4, 5))
# 然后定义一个转移矩阵,也就是从一个状态,采取一个可行的动作之后到达的状态,因为这里的状态和动作都是有限的,所以我们可以将他们存下来,比如第一行表示 state 0,向上和向左都是不可行的路线,所以给 -1 的值表示,向下走到达了 state 2,所以第二个值为 2,向右走到达了 state 1,所以第四个值是 1,保持不同还是在 state 0,所以最后一个标注为 0,另外几行类似。
transition_matrix = np.array([[-1, 2, -1, 1, 0],
[-1, 3, 0, -1, 1],
[0, -1, -1, 3, 2],
[1, -1, 2, -1, 3]])
# 最后定义每个状态的有效行动,比如 state 0 的有效行动就是下、右和静止,对应于 1,3 和 4
valid_actions = np.array([[1, 3, 4],
[1, 2, 4],
[0, 3, 4],
[0, 2, 4]])
# 定义 bellman equation 中的 gamma
gamma = 0.8
# 最后开始让智能体与环境交互,不断地使用 bellman 方程来更新 q 矩阵,我们跑 10 个 episode
for i in range(10):
start_state = np.random.choice([0, 1, 2], size=1)[0] # 随机初始起点
current_state = start_state
while current_state != 3: # 判断是否到达终点
action = random.choice(valid_actions[current_state]) # greedy 随机选择当前状态下的有效动作
next_state = transition_matrix[current_state][action] # 通过选择的动作得到下一个状态
future_rewards = []
for action_nxt in valid_actions[next_state]:
future_rewards.append(q_matrix[next_state][action_nxt]) # 得到下一个状态所有可能动作的奖励
q_state = reward[current_state][action] + gamma * max(future_rewards) # bellman equation
q_matrix[current_state][action] = q_state # 更新 q 矩阵
current_state = next_state # 将下一个状态变成当前状态
print('episode: {}, q matrix: \n{}'.format(i, q_matrix))
print()
# 可以看到在第一次 episode 之后,智能体就学会了在 state 2 的时候向下走能够得到奖励,通过不断地学习,在 10 个 episode 之后,智能体知道,在 state 0,向右走能得到奖励,在 state 1 向下走能够得到奖励,在 state 3 向右 走能得到奖励,这样在这个环境中任何一个状态智能体都能够知道如何才能够最快地到达宝藏的位置
#
# 从上面的例子我们简单的演示了 q-learning,可以看出自己来构建整个环境是非常麻烦的,所以我们可以通过一些第三方库来帮我们搭建强化学习的环境,其中最有名的就是 open-ai 的 gym 模块,下一章我们将介绍一下 gym。
| chapter7_RL/q-learning-intro.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# [](https://github.com/awslabs/aws-data-wrangler)
#
# # 8 - Redshift - COPY & UNLOAD
#
# `Amazon Redshift` has two SQL command that help to load and unload large amount of data staging it on `Amazon S3`:
#
# 1 - [COPY](https://docs.aws.amazon.com/redshift/latest/dg/r_COPY.html)
#
# 2 - [UNLOAD](https://docs.aws.amazon.com/redshift/latest/dg/r_UNLOAD.html)
#
# Let's take a look and how Wrangler can use it.
# +
import awswrangler as wr
engine = wr.catalog.get_engine("aws-data-wrangler-redshift")
# -
# ## Enter your bucket name:
import getpass
bucket = getpass.getpass()
path = f"s3://{bucket}/stage/"
# ## Enter your IAM ROLE ARN:
iam_role = getpass.getpass()
# ### Creating a Dataframe from the NOAA's CSV files
#
# [Reference](https://registry.opendata.aws/noaa-ghcn/)
# +
cols = ["id", "dt", "element", "value", "m_flag", "q_flag", "s_flag", "obs_time"]
df = wr.s3.read_csv(
path="s3://noaa-ghcn-pds/csv/1897.csv",
names=cols,
parse_dates=["dt", "obs_time"]) # ~127MB, ~4MM rows
df
# -
# ## Load and Unload with the regular functions (to_sql and read_sql_query)
# +
# %%time
wr.db.to_sql(
df,
engine,
schema="public",
name="regular",
if_exists="replace",
index=False
)
# +
# %%time
wr.db.read_sql_query("SELECT * FROM public.regular", con=engine)
# -
# ## Load and Unload with COPY and UNLOAD commands
# +
# %%time
wr.db.copy_to_redshift(
df=df,
path=path,
con=engine,
schema="public",
table="commands",
mode="overwrite",
iam_role=iam_role,
)
# +
# %%time
wr.db.unload_redshift(
sql="SELECT * FROM public.commands",
con=engine,
iam_role=iam_role,
path=path,
keep_files=True,
)
# -
| tutorials/008 - Redshift - Copy & Unload.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Python: Data analysis with numpy
# **Goal**: perform statistical computations and interprete the results!
# ## Goal
# The **goal** of this part is to analyze the data of the dataset world_alcohol with numpy. As a reminder, this dataset lists the alcohol consumption by country. We will look at which countries consume the most alcohol per population.
import numpy as np
world_alcohol = np.genfromtxt("world_alcohol.csv", delimiter=",", dtype="U75")
world_alcohol
# ## Perform comparisons
# In this section, we will learn how to perform comparisons with numpy. These last will return booleans (True or False).
# example for vector
vector = np.array([5, 10, 15, 20])
vector
vector == 5
# example for matrix
matrix = np.array([[5,10,15], [20,25,30], [35,40,45]])
matrix
matrix == 25
# ### Training
# In this practice, we will try to answer the following questions:
#
# * extract the 3rd column of world_alcohol and compare it to the country "Canada", assign the result to the variable countries_canada
#
# * extract the first column of world_alcohol and compare it to the string "1984", assign the result to the variable years_1984
world_alcohol = np.genfromtxt("world_alcohol.csv", delimiter=",", dtype="U75", skip_header=1)
world_alcohol
countries_canada = (world_alcohol[:,2] == "Canada")
countries_canada
years_1984 = (world_alcohol[:,0] == "1984")
years_1984
# ## Selecting items
# In this section we will have how to select specific elements from a numpy array. To do this, we will use the comparisons made above, these return vectors or matrix of booleans that we will use to condition the selection of elements from a numpy array.
# example with vector
print(vector)
is_equal_to_20 = (vector == 20)
vector_20 = vector[is_equal_to_20]
print(vector_20)
# example with matrix
print(matrix)
col_contain_25 = (matrix[:,1] == 25)
matrix_row_contain_25 = matrix[col_contain_25,:]
print(matrix_row_contain_25)
# ### Training
# In this practice, we will try to answer the following questions:
#
# * compare the 3rd column of world_alcohol to the string "Senegal"
#
# * assign the result to the variable country_is_senegal
#
# * select only the lines of world_alcohol for which country_is_senegal is true
#
# * assign the result to the variable country_senegal
#
# * display the results
#
# * do the same work to retrieve all rows corresponding to the year "1984" and assign the result to the years_1984 variable
country_is_senegal = (world_alcohol[:,2] == "Senegal")
country_senegal = world_alcohol[country_is_senegal]
print(country_senegal)
years_is_1984 = (world_alcohol[:,0] == "1984")
years_1984 = world_alcohol[years_is_1984]
print(years_1984)
# ## Perform comparisons with multiple conditions
# As a reminder, comparisons are the key concepts in numpy tables for selecting the desired elements. To realize multiple conditions, we use the ```&``` and ```|``` operators. Do not hesitate to use brackets to improve the reading of the code and to avoid mistakes.
# example with vector
vector
equal_to_5_and_10 = ((vector == 5) & (vector == 10))
equal_to_5_and_10
equal_to_5_or_10 = ((vector == 5) | (vector == 10))
equal_to_5_or_10
# ### Training
# In this practice, we will try to answer the following questions:
#
# * select the rows where the country is "Senegal" and the year is "1986"
# * create this double comparison and assign the result to the variable is_senegal_and_1986
# * use the variable is_senegal_and_1986 to select the corresponding rows in the table world_alcohol
# * assign the result to the variable rows_with_senegal_and_1986
# * display the result
is_senegal_and_1986 = ((world_alcohol[:,2] == "Senegal") & (world_alcohol[:,0] == "1984"))
rows_with_senegal_and_1986 = world_alcohol[is_senegal_and_1986]
print(rows_with_senegal_and_1986)
# ## Replace values in a numpy array
# example with vector
vector
equal_to_5_or_10
vector[equal_to_5_or_10] = 10
vector
# example for matrix
matrix
second_column_25 = (matrix[:,1] == 25)
second_column_25
matrix[second_column_25, 1] = 50
matrix
# ### Training
# In this practice, we will try to answer the following questions:
#
# * create a numpy array world_alcohol_2 equal world_alcohol to duplicate it under another name
# * replace all the years "1986" in the first column of world_alcohol_2 with "2018"
# * replace all "Wine" alcohols in the 4th column of world_alcohol_2 with "Beer"
world_alcohol_2 = world_alcohol.copy()
world_alcohol_2
world_alcohol_2[:,0][world_alcohol_2[:,0] == "1986"] = "2018"
world_alcohol_2
world_alcohol_2[:,3][world_alcohol_2[:,3] == "Wine"] = "Beer"
world_alcohol_2
# ## Replace empty strings
# In this practice, we will try to answer the following questions:
#
# * compare all the elements of the 5th column of world_alcohol with the empty string i.e. "
# * assign the result to the variable is_value_empty
# * select all the values of the 5th column of world_alcohol for which is_value_empty is equal to True and finally replace them by the string '0'
is_value_empty = (world_alcohol[:,4] == '')
world_alcohol[is_value_empty] = '0'
world_alcohol
# ## Converting data types
# example
string_vector = np.array(["1", "2", "3", "4", "5"])
string_vector
float_vector = string_vector.astype(float)
float_vector
int_vector = string_vector.astype(int)
int_vector
# ### Training
# In this practice, we will try to answer the following questions:
#
# * extract the 5th column of world alcohol and assign the result to the variable alcohol_consumption
# * use the astype() method to convert alcohol_consumption to decimal (float)
alcohol_consumption = world_alcohol[:,4]
alcohol_consumption = alcohol_consumption.astype(float)
alcohol_consumption
# ## Performing mathematical computations with numpy
# example with vector
vector
# sum()
vector.sum()
# mean()
vector.mean()
# max()
vector.max()
# example with matrix
matrix
# sum on the rows
matrix.sum(axis=1)
# sum on the columns
matrix.sum(axis=0)
# ### Training
# In this practice, we will try to answer the following questions:
#
# * use the sum() method to calculate the sum of the alcohol_consumption values and assign the result to the total_alcohol variable
# * use the method mean() to calculate the average of the values of alcohol_consumption and assign the result to the variable average_alcohol
# * display the results
total_alcohol = alcohol_consumption.sum()
total_alcohol
average_alcohol = alcohol_consumption.mean()
average_alcohol
# ## Calculate the total annual consumption
# In this practice, we will try to answer the following questions:
#
# * create a matrix named france_1986 which contains all the rows of world_alcohol corresponding to the year "1986" and the country "France"
# * extract the 5th column of france_1986, replace any empty string (") with '0' and convert the column to decimal (float) and assign the result to the variable france_alcohol
# * calculate the sum of france_alcohol and assign the result to the variable total_france_drinking
# * display the result
france_1986 = world_alcohol[(world_alcohol[:,0] == "1986") & (world_alcohol[:,2] == "France"),:]
france_1986
france_alcohol = france_1986[:,4]
france_alcohol
france_alcohol[france_alcohol == ''] = "0"
france_alcohol = france_alcohol.astype(float)
france_alcohol
total_france_drinking = france_alcohol.sum()
total_france_drinking
# ## Calculate the consumption for each country
# In this practice, we will try to answer the following questions:
#
# * first of all, we create an empty dictionary which will contain all the countries and their associated alcohol consumption, we will named it totals
# * then select the rows of world_alcohol corresponding to the given year, say 1989 and assign the result to the year variable
# * select all the countries in a list called countries
# * go through all the countries in the list using a loop and for each country:
# * select the lines of year corresponding to this country
# * assign the result to the variable country_consumption
# * extract the 5th column of country_consumption
# * replace any empty string in this column with 0
# * convert the column into a decimal (float)
# * calculate the sum of the column
# * add the sum to the totals dictionary, with the country name as key and the sum as value
# * display the dictionary totals
# +
totals = {}
year = world_alcohol[world_alcohol[:,0] == "1989",:]
countries = world_alcohol[:,2]
for country in countries:
country_consumption = year[year[:,2] == country,:]
alcohol_consumption = country_consumption[:,4]
alcohol_consumption[alcohol_consumption == ''] = "0"
alcohol_consumption = alcohol_consumption.astype(float)
totals[country] = alcohol_consumption.sum()
# -
print(totals)
# ## Find the country that consumes the most alcohol
# In this practice, we will try to answer the following questions:
#
# * create a variable highest_value which will keep in memory the highest value of the dictionary totals. We set it to 0 to start with
# * create a similar variable named highest_key which will keep in memory the name of the country associated with the highest value and set it to None
# * browse each country in the totals dictionary and if the value associated with the country is greater than highest_value, assign the value in question to the highest_value variable and assign the corresponding key (country name) to the highest_key variable
# * display the country that consumes the most alcohol (variable highest_key)
# +
highest_value = 0
highest_key = None
for country in totals:
consumption = totals[country]
if highest_value < consumption:
highest_value = consumption
highest_key = country
# -
print(highest_key, ':', highest_value)
| courses/2. Data analysis with numpy in Python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Predicting life of components: A simple Probabilistic Crack propagation example
#
# Predicting the life of a component that is prone to cracking is an age-old problem that has been studied ad-nauseam by the [fracture mechanics community](https://www.google.com/search?tbm=bks&q=fracture+mechanics). [Crack propagation models](https://en.wikipedia.org/wiki/Fracture_mechanics) are at the core of Prognostics and Health Management (PHM) solutions for engineering systems and the aptly titled book [Prognostics and Health Management of Engineering Systems: An Introduction](https://books.google.com/books?id=pv9RDQAAQBAJ&lpg=PR3&dq=prognostics%20and%20health%20management%20of%20engineering%20systems&pg=PR3#v=onepage&q=prognostics%20and%20health%20management%20of%20engineering%20systems&f=false) provides a great example of how real world data can be used to calibrate engineering models. With the example below, we would like to motivate the use of "Hybrid models" that combine probabilistic learning techniques and engineering domain models.
#
# The phenomenon of fatigue crack propagation can be modeled with the Paris law. Paris law relates the rate of crack growth $\left(da/dN\right)$ to the stress intensity factor $\left(\Delta K = \Delta\sigma\sqrt{\pi a}\right)$ through the equation below:
#
# $\frac{da}{dN}=C(\Delta\sigma\sqrt{\pi a})^m$
#
# Where $a$ is the crack length, $N$ is the number of loading cycles, $\sigma$ is the stress, and $C, m$ are material properties.
#
# Integrating the Paris law for a specific geometry and loading configuration, we arrive at the analytical formulation for the size of a crack as a function of the loading cycle as shown below:
#
# $a(N) = \left[ N C \left(1-\frac{m}{2}\right) \left(\Delta\sigma\sqrt{\pi}\right)^m + a_0^{1-\frac{m}{2}}\right]^\frac{2}{2-m}$
#
# where $a_0$ is the initial crack length.
#
# The parameters $C$ and $m$ need to be calibrated for each application with crack length $(a)$ vs loading cycles $(N)$ data. Such data is usually obtained during maintenance and inspections of engineering systems. In this example, we will use the sample dataset from the [PHM book by <NAME>](https://books.google.com/books?id=pv9RDQAAQBAJ&lpg=PR3&dq=prognostics%20and%20health%20management%20of%20engineering%20systems&pg=PR3#v=onepage&q=prognostics%20and%20health%20management%20of%20engineering%20systems&f=false) .
#
# We will demonstrate a probabilistic calibration of $C$ and $m$ using Tensorflow Probability.
#
# At BHGE Digital we leverage our [Depend-on-Docker](https://github.com/bhgedigital/depend-on-docker) project for automating analytics development. A sample of such automation is available [here](https://github.com/bhgedigital/bayesian_calibration), with the complete code of the example below.
#
# With the right automation for developing and deploying analytics in place, we start by importing the following libraries:
# +
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
import tensorflow_probability as tfp
import pandas as pd
tfd = tfp.distributions
tfb = tfp.bijectors
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
from tensorflow_probability.python.mcmc import util as mcmc_util
import time
import math
import matplotlib
matplotlib.rc('xtick', labelsize=20)
matplotlib.rc('ytick', labelsize=20)
font = {'family' : 'Dejavu Sans','size' : 20}
matplotlib.rc('font', **font)
# -
# ### Setting up the data
#
# Setting up the data for the calibration comes next. Here we leverage the dataset provided in the Table 4.2 of the [PHM book by <NAME>](https://books.google.com/books?id=pv9RDQAAQBAJ&lpg=PR3&dq=prognostics%20and%20health%20management%20of%20engineering%20systems&pg=PR3#v=onepage&q=prognostics%20and%20health%20management%20of%20engineering%20systems&f=false):
#
# true values of the parameters
t = np.arange(0,1600, 100) #cycles
y = [0.0100,0.0109,0.0101,0.0107,0.0110,0.0123,0.0099,0.0113,
0.0132,0.0138,0.0148,0.0156,0.0155,0.0141,0.0169,0.0168] # measured crack size data
# ### Priors
#
# For Bayesian calibration, we need to define the prior distributions for the calibration variables. In a real application, these priors can be informed by a subject matter expert. For this example, we will assume that both $C$ and $m$ are Gaussian and independent.
#
prio_par_C = [-23.0, 1.1] # [location, scale] for Normal Prior
prio_par_m = [4.0, 0.2] # [location, scale] for Normal Prior
rv_m = tfd.Normal(loc = 0.0, scale = 1.0, name = 'm_norm') # Random variable m definition
rv_C = tfd.Normal(loc = 0.0, scale = 1.0, name = 'C_norm') # Random variable logC definition
# ### Log-prob function
#
# We have defined external parameters and standard Normal distribution for both variables, just to sample from a normalized space. Therefore, we will need to de-normalize both random variables when computing the crack model.
#
# Now we define the joint log probability for the random variables being calibrated and the associated crack model:
#
def joint_logprob(cycles, observations, y0, C_norm, m_norm):
# Joint logProbability function for both random variables and observations.
# Some constants
dsig = 75.0
B = tf.constant(dsig*math.sqrt(math.pi), tf.float32)
# Computing m and logC on original space
C = C_norm*tf.sqrt(prio_par_C[1]) + prio_par_C[0] #
m = m_norm*tf.sqrt(prio_par_m[1]) + prio_par_m[0]
# Crack Propagation model
crack_model = tf.pow(cycles*tf.exp(C)*(1-m/2.0)*tf.pow(B,m) + tf.pow(y0, 1-m/2.0), 2.0/(2.0-m))
y_model = observations - crack_model
# Defining child model random variable
rv_model = tfd.Independent(tfd.Normal(loc = tf.zeros(observations.shape), scale = 0.001),
reinterpreted_batch_ndims=1, name = 'model')
# Sum of logProbabilities
sum_log_prob = rv_C.log_prob(C_norm) + rv_m.log_prob(m_norm) + rv_model.log_prob(y_model)
return sum_log_prob
# ### Sampler
#
# Finally, it is time to set up the sampler and run a Tensorflow session:
# +
# Number of samples and burnin for the MCMC sampler
samples = 20000
burnin = 10000
# Initial state for the HMC
initial_state = [0.0,0.0]
# Converting the data into tensors
cycles = tf.convert_to_tensor(t,tf.float32)
observations = tf.convert_to_tensor(y,tf.float32)
y0 = tf.convert_to_tensor(y[0], tf.float32)
# Setting up a target posterior for our joint logprobability
unormalized_target_posterior= lambda *args: joint_logprob(cycles, observations, y0, *args)
# And finally setting up the mcmc sampler
[C_samples,m_samples], kernel_results = tfp.mcmc.sample_chain(num_results= samples, num_burnin_steps= burnin,
current_state=initial_state,
kernel= tfp.mcmc.HamiltonianMonteCarlo(
target_log_prob_fn=unormalized_target_posterior,
step_size = 0.02,
num_leapfrog_steps=6))
# Tracking the acceptance rate for the sampled chain
acceptance_rate = tf.reduce_mean(tf.to_float(kernel_results.is_accepted))
# Actually running the sampler
with tf.Session() as sess:
[C_samples_, m_samples_, acceptance_rate_] = sess.run([C_samples, m_samples, acceptance_rate])
# Some initial results
print('acceptance_rate:', acceptance_rate_)
# -
# ### Plotting Results
#
# If everything has gone according to plan, at this point we will see a acceptance rate of the sampler of around 60%. Pretty good for our first bayesian hierarchical crack propagation model. A key metric for the HMC sampler are the sampled chains themselves, which should look "mixed" . In this case, the sampler did a decent job as seen below:
#
# +
# plotting the mcmc chains
plt.figure(figsize=(20,10))
plt.plot(np.arange(samples), C_samples_,color='#377eb8')
plt.title('C samples',fontsize=20)
plt.figure(figsize=(20,10))
plt.plot(np.arange(samples), m_samples_,color= '#e41a1c')
plt.title('m samples',fontsize=20)
# -
#
#
# If we collect our sample and rescale them to the original space, we can get some summary statistics on the posterior estimates of the calibrated parameters $\log(C)$ and $m$ and take a look at the distributions:
# +
# Converting to proper scale
C_samples_scale = C_samples_*np.sqrt(prio_par_C[1]) + prio_par_C[0]
m_samples_scale = m_samples_*np.sqrt(prio_par_m[1]) + prio_par_m[0]
df = pd.DataFrame(np.concatenate([m_samples_scale[:,None], C_samples_scale[:,None]], axis = 1), columns = ['m', 'logC'])
ax = pd.plotting.scatter_matrix(df,figsize=(7.2,7.2))
for axi in ax:
axi[0].tick_params(labelsize=16)
axi[1].tick_params(labelsize=16)
# -
# It is interesting to note that although we started off with two independent Gaussian distributions for $C$ and $m$, the posterior distributions are highly correlated. This is because, the solution space dictates that for high values for m, the only physically meaningful results lie at small values of C and vice-versa. If one were to have used any number of deterministic optimization techniques to find the "best fit" for $C$ and $m$ that fits this dataset, we could end up in any value that lies on the straight line depending on our starting point and constraints. Performing a probabilsitic optimization (a.k.a. Bayesian calibration) provides us with a global view of all possible solutions that can explain the dataset.
#
#
df.describe(percentiles=[.05, .95])
# ### Sampling the Posterior for Prognostic
#
# And now for the final act, we shall define a posterior function for our probabilistic crack propagation model, in order to finaly make the prognostic:
# +
def posterior(C_samples, m_samples, time):
n_s = len(C_samples)
n_inputs = len(time)
# Some Constants
dsig = 75.0
B = tf.constant(dsig*math.sqrt(math.pi), tf.float32)
# Crack Propagation model
y_model = tf.pow(time[:,None]*tf.exp(C_samples[None,:])*(1-m_samples[None,:]/2.0)*tf.pow(B,m_samples[None,:]) +
tf.pow(y0, 1-m_samples[None,:]/2.0), 2.0/(2.0-m_samples[None,:]))
noise = tfd.Normal(loc = 0.0, scale = 0.001)
#y_model = tf.convert_to_tensor(y_model, tf.float32)
# rv_model = tfd.Independent(tfd.Normal(loc = y_model, scale = 1e-3),
# reinterpreted_batch_ndims=1, name = 'model')
samples = y_model + noise.sample(n_s)[tf.newaxis,:]
# samples = rv_model.sample(1)
with tf.Session() as sess:
samples_ = sess.run(samples)
return samples_
# -
time = np.arange(0,3000,100)
y_samples = posterior(C_samples_scale, m_samples_scale, time)
print(y_samples.shape)
# +
lower_per = np.nanpercentile(y_samples,2.5, axis = 1)
upper_per = np.nanpercentile(y_samples,97.5, axis = 1)
plt.figure(figsize =(20,10))
for samps in range(y_samples.shape[1]):
if samps % 250 == 0:
plt.plot(time, y_samples[:,samps],color='blue', alpha = 0.05)
plt.plot(time, y_samples[:,-1],color='blue', label='Sample',alpha = 0.5)
plt.plot(time, np.nanmedian(y_samples,axis=1), 'r', label = 'Median',linewidth=2)
plt.plot(t,y,'kx', label = 'Data',markersize=12)
plt.xlabel('Cycles')
plt.ylabel('Crack size ')
plt.hlines(0.05, np.min(time), np.max(time), linestyles = '--', label = 'threshold')
plt.ylim([-0., 0.15])
plt.legend(loc=2)
# -
# The predicted mean with the 95% uncertainty bounds of crack length using the hybrid-physics probabilistic model is shown above. Clearly, the model captures both the mean behavior as well as provides an estimate of uncertainty (of the model prediction) for every time point.
# ## Next steps
#
# This is the first of a series of examples aimed at expanding the use of Probabilistic Deep Learning techniques for industrial applications. We hope you have found this useful. We would love to hear about your applications and look forward to seeing these methods used in ways that we cannot imagine. Stay tuned to this blog feed for more updates and examples on anomaly detection, missing data estimation and forecasting with variational inference.
| notebooks/.ipynb_checkpoints/Simple_Probabilistic_Crack_Growth_Model-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import pandas as pd
import spacy
from IPython.display import display, HTML, Markdown
import random
fname = 'dataset.xlsx'
df = pd.read_excel(fname)
display(HTML(df.sample(10).to_html()))
# +
def printmd(string, color="Black"):
colorstr = f"<span style='color:{color}'>{string}</span>"
display(Markdown(colorstr))
# -
# ### Count occurrences of specific words or types of words in both types of statements.
# ### Visualize the proportion
# + pycharm={"name": "#%%\n"}
from collections import Counter
right=0
wrong=0
nlp = spacy.load("en_core_web_sm")
def visualizeFeature(name, tcount, fcount):
labels = ['Truthful', 'Deceptive']
sizes = [tcount, fcount]
fig1, ax1 = plt.subplots()
ax1.pie(sizes, labels=labels, autopct='%1.1f%%',
shadow=True, startangle=90, textprops={'color': "black"})
printmd(
f"\n\n## **{name}:** {labels[0]} {tcount} {labels[1]} {fcount}")
plt.show()
# -
# # Significance tests
#
# H<sub>0</sub> : Words from wordlist are equally likely to occur in truthful and deceptive statements
#
# H<sub>A</sub> : Words from wordlist occur in truthful and deceptive statements with different probability
#
#
# **Sample**: measure the proportion truthful/deceptive in the data
#
# **Simulation**: Generate 1000 pairs of 160 docs, randomly inserting words of interest according to the probability with which they occur in the text according to H<sub>0</sub>. Of course, this kind of simulation is rather naïve, since actual language places a lot more constraints on the simulation. Theoretically, an advanced generative model could be used to generate better data. However, such simulation may be skewed, because generative models are typically trained on truthful data.
#
# **Assumption**: If less than 1% of the simulations show a truthful/deceptive proportion equal to or less (or greater, for proportions > 1) than the measured one, reject H<sub>0</sub>
#
# If H<sub>0</sub> is rejected, the feature can be used to predict the veracity of a statement
# +
#generate a set of "truthful" and "deceptive" "documents" with the same number of words
#count "occurrences" of a "word" that has a probability prob to appear in a "document"
def simstat(numdoc, numwords, prob):
tcount = 0
for i in range(numdoc):
for j in range(numwords):
if(random.uniform(0, 1) < prob):
tcount += 1
return tcount
#simulation: 1000 times generate 2 random documents, each containing words_per_doc
def simulation(numdoc, numwords, prob):
sim = []
for k in range(1000):
sim.append(simstat(numdoc, int(words_per_doc), prob) /
simstat(numdoc, int(words_per_doc), prob))
return sim
# +
import matplotlib.pyplot as plt
import numpy as np
# %matplotlib inline
def visualizeSimulation(sim, measured_proportion):
#sort sim
sim = sorted(sim)
bottom = sim[int(0.01*len(sim))]
top = sim[int(0.99*len(sim))]
#count members of sim that are below the measured proportion
below = sum(map(lambda x: x < measured_proportion, sim))
#above = sum(map(lambda x: x > 1/measured_proportion, sim))
alpha = 0.01
pfactor = below/len(sim)
plt.hist(sim, density=False, bins=30) # density=False would make counts
plt.ylabel('Frequency')
plt.xlabel('Proportion')
plt.show()
print(
f"Probability of getting a ratio at or below {measured_proportion:.2f}: {pfactor*100:.2f}%")
if pfactor < alpha:
printmd("Feature can be used for veracity assessment\n", color="green")
else:
printmd("Feature cannot be used for veracity assessment\n", color="red")
# -
# # Extract a few language features that Pennebaker claims can be used to assess veracity of written text.
#
# The built-in "count_by" method of spaCy cannot be used because we want the ability to count not just spacific POS, but specific POS that are also a part of a short list. For example, from the auxilliary verb group we only care about the modal verbs. As it turns out, further splitting that group into two yields a really good results in terms of distribution between deceptive and truthful statements.
#
# The idea is to count the occurences of members of each feature group in deceptive as well as truthful statements, and if they are unbalanced, perform a significance test.
# +
import matplotlib.pyplot as plt
from spacy.tokens import Doc
def count_words(doc, type, wordlist):
alloftype = [token.lower_ for token in doc if token.pos_ == type]
if wordlist:
alloftype = [x for x in alloftype if x in wordlist]
return sum(Counter(alloftype).values())
# +
featureTypes=[
{
'name':'i-words',
'POS':'PRON',
'wordlist': ['we','i', 'me', 'myself', 'my', 'mine'],
'tcount':0,
'fcount':0,
'indicates':None
},
{
'name': 'verbs',
'POS': 'VERB',
'wordlist': None,
'tcount': 0,
'fcount': 0,
'indicates': None
},
{
'name': 'articles',
'POS': 'DET',
'wordlist': ['a', 'an', 'the'],
'tcount':0,
'fcount':0,
'indicates':None
},
{
'name': 'modal verbs 1',
'POS': 'AUX',
'wordlist': ["could", "should"],
'tcount': 0,
'fcount': 0,
'indicates':None
},
{
'name': 'modal verbs 2',
'POS': 'AUX',
'wordlist': ["would", "may"],
'tcount': 0,
'fcount': 0,
'indicates':None
},
{
'name': 'cognitive verbs',
'POS': 'VERB',
'wordlist': ['realize' , 'think', 'understand', 'figure', 'derive', "know", "believe", "recognize", "appreciates"],
'tcount': 0,
'fcount': 0,
'indicates':None
},
{
'name': 'interjections',
'POS': 'INTJ',
'wordlist': None,
'tcount': 0,
'fcount': 0,
'indicates': None
}
]
twcount = 0
fwcount = 0
docs = []
labels = []
for index, row in df.iterrows():
text = row['Transcription']
doc = nlp(text)
docs.append(doc)
labels.append(row['Type'])
cdoc = Doc.from_docs(docs)
for doc, label in zip(docs, labels):
for feature in featureTypes:
if label == 'Truthful':
feature['tcount'] += count_words(doc, feature['POS'], feature['wordlist'])
else:
feature['fcount'] += count_words(doc, feature['POS'], feature['wordlist'])
if label == 'Truthful':
twcount += len(doc)
else:
fwcount += len(doc)
numdocs = len(docs)
total_wordcount = twcount + fwcount
words_per_doc = total_wordcount/len(docs)
for feature in featureTypes:
listlen = len(feature['wordlist']) if feature['wordlist'] else 1
global_occurences = feature['tcount'] + feature['fcount']
visualizeFeature(feature['name'], feature['tcount'], feature['fcount'])
prob = global_occurences/total_wordcount
sim = simulation(numdocs, words_per_doc, prob)
measured_proportion = feature['tcount']/feature['fcount']
if measured_proportion > 1. :
measured_proportion = 1./measured_proportion
feature['indicates'] = 'truthful'
else:
feature['indicates'] = 'deceptive'
visualizeSimulation(sim, measured_proportion)
# -
# Next steps: construct features by checking for the presence of multiple significant words in a statement.
#
# Perhaps add a score, either 1 for each significant word present, or assign different weight based on the calculated significance
# +
def pos_list(doc, pos):
pos_list = [token.lemma for token in doc if token.pos_ == pos]
return Counter(pos_list)
def rwratio(lieword, trueword, counter):
right = 0
wrong = 0
if counter[lieword] > counter[trueword] and label == 'Truthful':
wrong += 1
elif counter[lieword] < counter[trueword] and label == 'Deceptive':
wrong += 1
elif counter[lieword] != counter[trueword]:
right += 1
return right, wrong
# +
MINSCORE = 0
def veval(doc, features):
tscore = 0
fscore = 0
for feature in features:
fcount = count_words(doc, feature['POS'], feature['wordlist'])
if feature['indicates'] == 'truthful':
tscore += fcount
else:
fscore += fcount
#print(f"{feature['name']} {fcount}")
score = tscore - fscore/2 #compensate for the feature set being unbalanced
if score == 0:
return score
elif score > 0:
return 'Truthful'
else:
return 'Deceptive'
right = 0
wrong = 0
usable_features = ['modal verbs 1', 'modal verbs 2', 'cognitive verbs']
features = [x for x in featureTypes if x['name'] in usable_features]
for doc, label in zip(docs, labels):
score = veval(doc, features)
if score != 0:
if score == label:
right += 1
else:
wrong += 1
print(f"Right: {right}")
print(f"Wrong: {wrong}")
print(f"Accuracy: {right/(right+wrong)*100:.2f}%")
# -
| analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] _cell_guid="99c2be30-0fd8-4b60-885e-6bfd1b0246f6" _uuid="7945a7f9b578f6c3ad98d6066e036f918419fa5a"
# # Gini Coefficient - An Intuitive Explanation
#
# I was struggling a bit with the definition of the Scoring Metric. Googling "Gini Coefficient" gives you mostly economic explanations. Here is a descriptive explanation with regard to the challenge.
#
# First, let's define our predictions and their actual values:
# + _cell_guid="1acaec53-ac2c-4474-a2a2-629d41b161c5" _uuid="a1edc32cb536f27c798be8fc6cec932b19f409d9"
import numpy as np
import matplotlib.pyplot as plt
import scipy.interpolate
import scipy.integrate
predictions = [0.9, 0.3, 0.8, 0.75, 0.65, 0.6, 0.78, 0.7, 0.05, 0.4, 0.4, 0.05, 0.5, 0.1, 0.1]
actual = [1, 1, 1, 1, 1, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0]
# + [markdown] _cell_guid="9c76b2b1-7ee3-4092-9fbd-a185aba4c6f1" _uuid="717d9928be1af4b1303db6183b52140a0f96d102"
# We use the Python implementation from the [Gini coefficient discussion with code samples](https://www.kaggle.com/c/ClaimPredictionChallenge/discussion/703):
# + _cell_guid="017f3ca4-8b00-4ac8-80fd-dda0c714dd8d" _uuid="16fd6f021f05aa00705fac1b6d9b14c132e7271a"
def gini(actual, pred):
assert (len(actual) == len(pred))
all = np.asarray(np.c_[actual, pred, np.arange(len(actual))], dtype=np.float)
all = all[np.lexsort((all[:, 2], -1 * all[:, 1]))]
totalLosses = all[:, 0].sum()
giniSum = all[:, 0].cumsum().sum() / totalLosses
giniSum -= (len(actual) + 1) / 2.
return giniSum / len(actual)
def gini_normalized(actual, pred):
return gini(actual, pred) / gini(actual, actual)
# + [markdown] _cell_guid="d5d08469-ef0b-425f-9ce2-aee204e944ee" _uuid="75550b4a620a7da300aa509b3b6752f8290ff17b"
# We calculate the Gini coefficient for the predictions:
# + _cell_guid="f439a2c7-da20-4595-b151-23b9104a2f0e" _uuid="35fe46255a39081c4e8165893a51a0d1cc106e94"
gini_predictions = gini(actual, predictions)
gini_max = gini(actual, actual)
ngini= gini_normalized(actual, predictions)
print('Gini: %.3f, Max. Gini: %.3f, Normalized Gini: %.3f' % (gini_predictions, gini_max, ngini))
# + [markdown] _cell_guid="6ab2c51d-7871-4b88-bc6e-b810d0b44176" _uuid="3fce33a6cf4491ca1ba457c1610e82be79754ef1"
# **So, how do we get this Gini of 0.189 and the Normalized Gini of 0.630?**
#
# ## Economic Explanation
#
# The first figure on the "Gini Coefficient" Wikipedia article is this one:
#
# <img style="float:left" height="400" width="400" src="https://upload.wikimedia.org/wikipedia/commons/thumb/5/5b/Economics_Gini_coefficient.svg/500px-Economics_Gini_coefficient.svg.png"/>
# <div style="clear:both"/>
#
# They go through the population from poorest to richest and plot the running total / cumulative share of income, which gives them the Lorenz Curve. The Gini Coefficient is then defined as the blue area divided by the area of the lower triangle.
#
# ## Application to our challenge
#
# Instead of going through the population from poorest to richest, we go through our predictions from lowest to highest.
# + _cell_guid="0573d447-d308-4395-9b4d-6483396dfcff" _uuid="57c69007da514a299465e05e8925a8525c9f4723"
# Sort the actual values by the predictions
data = zip(actual, predictions)
sorted_data = sorted(data, key=lambda d: d[1])
sorted_actual = [d[0] for d in sorted_data]
print('Sorted Actual Values', sorted_actual)
# + [markdown] _cell_guid="3c72f2a1-e695-4375-b3fa-29eac9068bc7" _uuid="fb5531a044c31e6feb562357e6a31d5f705958c9"
# Instead of summing up the income, we sum up the actual values of our predictions:
# + _cell_guid="a6f4cbb0-123d-4b8a-a858-b11391e132d2" _uuid="1dca60a72fda9a101e5daed88aa73947cd35218a"
# Sum up the actual values
cumulative_actual = np.cumsum(sorted_actual)
cumulative_index = np.arange(1, len(cumulative_actual)+1)
plt.plot(cumulative_index, cumulative_actual)
plt.xlabel('Cumulative Number of Predictions')
plt.ylabel('Cumulative Actual Values')
plt.show()
# + [markdown] _cell_guid="92be5881-fecc-423b-9744-c854d8ab3274" _uuid="9ff032a20d9cefef1268ca8658539510c208747e"
# This corresponds to the Lorenz Curve in the diagram above.
#
# We normalize both axes so that they go from 0 to 100% like in the economic figure and display the 45° line for illustrating random guessing:
# + _cell_guid="e3da8bdb-6118-406e-b25d-7b7a06550309" _uuid="ffdb97781b685bc4634d8e5479af52734ca5287e"
cumulative_actual_shares = cumulative_actual / sum(actual)
cumulative_index_shares = cumulative_index / len(predictions)
# Add (0, 0) to the plot
x_values = [0] + list(cumulative_index_shares)
y_values = [0] + list(cumulative_actual_shares)
# Display the 45° line stacked on top of the y values
diagonal = [x - y for (x, y) in zip(x_values, y_values)]
plt.stackplot(x_values, y_values, diagonal)
plt.xlabel('Cumulative Share of Predictions')
plt.ylabel('Cumulative Share of Actual Values')
plt.show()
# + [markdown] _cell_guid="f0e3ee8c-245a-4dd9-a1b7-2538954fc0f3" _uuid="dd59403cb0747474216887177e9cbce653aafe55"
# Now, we calculate the orange area by integrating the curve function:
# + _cell_guid="fbce16e9-ee12-41d0-97e2-9c1263601d20" _uuid="7832cedb49a7cc7c5ff58818523a4f42f484c451"
fy = scipy.interpolate.interp1d(x_values, y_values)
blue_area, _ = scipy.integrate.quad(fy, 0, 1, points=x_values)
orange_area = 0.5 - blue_area
print('Orange Area: %.3f' % orange_area)
# + [markdown] _cell_guid="d6347ea7-81de-4662-b049-4296629c0788" _uuid="8ff8f72335376c8ea6492c7755687878e72b998e"
# So, the orange area is equal to the Gini Coefficient calcualted above with the `gini` function. We can do the same using the actual values as predictions to get the maximum possible Gini Coefficient.
# + _cell_guid="b23bc1ed-19c8-4f4b-b65b-7205138f5467" _uuid="325e65de5224d5850a29b1ca0c27dae620677252"
cumulative_actual_shares_perfect = np.cumsum(sorted(actual)) / sum(actual)
y_values_perfect = [0] + list(cumulative_actual_shares_perfect)
# Display the 45° line stacked on top of the y values
diagonal = [x - y for (x, y) in zip(x_values, y_values_perfect)]
plt.stackplot(x_values, y_values_perfect, diagonal)
plt.xlabel('Cumulative Share of Predictions')
plt.ylabel('Cumulative Share of Actual Values')
plt.show()
# Integrate the the curve function
fy = scipy.interpolate.interp1d(x_values, y_values_perfect)
blue_area, _ = scipy.integrate.quad(fy, 0, 1, points=x_values)
orange_area = 0.5 - blue_area
print('Orange Area: %.3f' % orange_area)
# + [markdown] _cell_guid="82bd2a05-48cb-4371-b3b9-217aa41e8c71" _uuid="c032f242a2b15103185fbae9af91de142bc61b4b"
# Dividing both orange areas gives us the Normalized Gini Coefficient:
#
# 0.189 / 0.3 = 0.630
# + [markdown] _cell_guid="ab3a0113-cae2-4d70-ade4-3d88a0711ad3" _uuid="03fcc0ce3db7b15ee5a96f6e7a2584a6a43a3878"
# ## Alternative explanation
#
# I also found another interpreation of the Gini Coefficient [here](http://www.rhinorisk.com/Publications/Gini%20Coefficients.pdf). Again, we take the predictions and actual values from above and sort them in descending order:
# + _cell_guid="e2238378-b157-4a33-96d5-44797eaa784d" _uuid="66cd9caee9275a63149cda7754a47ea1eaec7069"
print("Predictions", predictions)
print("Actual Values", actual)
print("Sorted Actual", list(reversed(sorted_actual)))
# + [markdown] _cell_guid="1fd79d9e-bd7f-4d50-8c1d-c457238db2b9" _uuid="61b70166cff94ebc224887428ce2f89db14ef6e3"
# Now, we count the number of swaps of adjacent digits (like in bubble sort) it would take to get from the "Sorted Actual" state to the "Actual Values" state. In this scenario, it would take 10 swaps.
#
# We also calculate the number of swaps it would take on average to get from a random state to the "Actual Values" state. With 6 ones and 9 zeros this results in
#
# $$\frac{6 \cdot 9}{2} = 27$$ swaps.
#
# The Normalized Gini-Coefficient is how far away our sorted actual values are from a random state measured in number of swaps:
#
# $$NGini = \frac{swaps_{random} - swaps_{sorted}}{swaps_{random}} = \frac{27 - 10}{27} = 63\%$$
#
# + [markdown] _cell_guid="a6153b13-832e-4b59-83ab-7eeb90ebdaea" _uuid="38b44b35ce1a4ca8b3f8bd74454dd63ba363f5b3"
# I hope I could give you a better feeling for the Gini coefficient.
| 10 poer sugero safe driver prediction/gini-coefficient-an-intuitive-explanation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Métricas de Regressão
import pandas as pd
import numpy as np
import sklearn.metrics as metricas
# +
X = [1, 3, 3]
y = [1, 4, 7]
y_pred = [3, 2, 4.5]
df = pd.DataFrame({ 'X': X, 'y': y, 'y_pred': y_pred })
df
# -
# ### MAE
metricas.mean_absolute_error(df['y'], df['y_pred'])
mae = (df['y'] - df['y_pred']).abs().sum() / df.shape[0]
mae
# ### MSE
metricas.mean_squared_error(df['y'], df['y_pred'])
df.shape[0]
mse = ((df['y'] - df['y_pred'])**2).sum() / df.shape[0]
mse
# ### RMSE
metricas.mean_squared_error(df['y'], df['y_pred'], squared=False)
rmse = np.sqrt(mse)
rmse
rmse = mse ** (1/2)
rmse
| 04-Metricas de Regressao.ipynb |
# ---
# jupyter:
# jupytext:
# formats: ipynb,py:percent
# text_representation:
# extension: .py
# format_name: percent
# format_version: '1.3'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %% [markdown]
#
# <p align="center">
# <img width="100%" src="../../../multimedia/mindstorms_51515_logo.png">
# </p>
#
# # `the_mvp_buggy`
# Python equivalent of the `The MVP Buggy` program. Make MVP Buggy go round.
#
# # Required robot
# * MVP (in its buggy form)
#
# <img src="../multimedia/mvp_buggy.jpg" width="50%" align="center">
#
# # Source code
# You can find the code in the accompanying [`.py` file](https://github.com/arturomoncadatorres/lego-mindstorms/blob/main/base/mvp/programs/the_mvp_buggy.py). To get it running, simply copy and paste it in a new Mindstorms project.
#
# # Imports
# %%
from mindstorms import MSHub, Motor, MotorPair, ColorSensor, DistanceSensor, App
from mindstorms.control import wait_for_seconds, wait_until, Timer
from mindstorms.operator import greater_than, greater_than_or_equal_to, less_than, less_than_or_equal_to, equal_to, not_equal_to
import math
# %% [markdown]
# # Initialization
# %%
print("-"*15 + " Execution started " + "-"*15 + "\n")
# %%
hub = MSHub()
# %%
hub.status_light.on('black')
# %% [markdown]
# # Configure motors
# %%
print("Configuring motors...")
motor_steer = Motor('A') # Front wheels (for steering)
motor_power = Motor('B') # Back wheels (for moving)
print("DONE!")
# %% [markdown]
# # Set steering motor to starting position
# %%
print("Setting steering motor to position 0...")
motor_steer.run_to_position(45, speed=100)
motor_steer.run_to_position(0, speed=100)
print("DONE!")
# %% [markdown]
# # Move MVP
# %%
print("Steering...")
motor_steer.run_to_position(50, speed=35)
print("DONE!")
# %%
print("Moving...")
motor_power.set_default_speed(80)
motor_power.run_for_rotations(-16)
print("DONE!")
# %%
print("-"*15 + " Execution ended " + "-"*15 + "\n")
| base/mvp/programs/the_mvp_buggy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Environment (conda_tensorflow_p36)
# language: python
# name: conda_tensorflow_p36
# ---
# + [markdown] colab_type="text" id="KFPcBuVFw61h"
# # DeepLab Demo
#
# This demo will demostrate the steps to run deeplab semantic segmentation model on sample input images.
# + cellView="code" colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="kAbdmRmvq0Je"
#@title Imports
import os
from io import BytesIO
import tarfile
import tempfile
from six.moves import urllib
from matplotlib import gridspec
from matplotlib import pyplot as plt
import numpy as np
from PIL import Image
import tensorflow as tf
# + cellView="code" colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="vN0kU6NJ1Ye5"
#@title Helper methods
class DeepLabModel(object):
"""Class to load deeplab model and run inference."""
INPUT_TENSOR_NAME = 'ImageTensor:0'
OUTPUT_TENSOR_NAME = 'SemanticPredictions:0'
INPUT_SIZE = 513
FROZEN_GRAPH_NAME = 'frozen_inference_graph'
def __init__(self, tarball_path):
"""Creates and loads pretrained deeplab model."""
self.graph = tf.Graph()
graph_def = None
# Extract frozen graph from tar archive.
tar_file = tarfile.open(tarball_path)
for tar_info in tar_file.getmembers():
if self.FROZEN_GRAPH_NAME in os.path.basename(tar_info.name):
file_handle = tar_file.extractfile(tar_info)
graph_def = tf.GraphDef.FromString(file_handle.read())
break
tar_file.close()
if graph_def is None:
raise RuntimeError('Cannot find inference graph in tar archive.')
with self.graph.as_default():
tf.import_graph_def(graph_def, name='')
self.sess = tf.Session(graph=self.graph)
logits = self.graph.get_tensor_by_name('ResizeBilinear_1:0')[:, :513, :513]
logits = tf.image.resize_images(logits, (256, 256), method=tf.image.ResizeMethod.BILINEAR, align_corners=True)
self.logits = logits
print(self.logits.shape)
def run(self, image, logits=True):
"""Runs inference on a single image.
Args:
image: A PIL.Image object, raw input image.
Returns:
resized_image: RGB image resized from original input image.
seg_map: Segmentation map of `resized_image`.
"""
width, height = image.size
resize_ratio = 1.0 * self.INPUT_SIZE / max(width, height)
target_size = (int(resize_ratio * width), int(resize_ratio * height))
resized_image = image.convert('RGB').resize(target_size, Image.ANTIALIAS)
batch_seg_map = self.sess.run(
self.logits,
feed_dict={self.INPUT_TENSOR_NAME: [np.asarray(resized_image)]})
seg_map = batch_seg_map[0]
return resized_image, seg_map
def create_pascal_label_colormap():
"""Creates a label colormap used in PASCAL VOC segmentation benchmark.
Returns:
A Colormap for visualizing segmentation results.
"""
colormap = np.zeros((256, 3), dtype=int)
ind = np.arange(256, dtype=int)
for shift in reversed(range(8)):
for channel in range(3):
colormap[:, channel] |= ((ind >> channel) & 1) << shift
ind >>= 3
return colormap
def label_to_color_image(label):
"""Adds color defined by the dataset colormap to the label.
Args:
label: A 2D array with integer type, storing the segmentation label.
Returns:
result: A 2D array with floating type. The element of the array
is the color indexed by the corresponding element in the input label
to the PASCAL color map.
Raises:
ValueError: If label is not of rank 2 or its value is larger than color
map maximum entry.
"""
if label.ndim != 2:
raise ValueError('Expect 2-D input label')
colormap = create_pascal_label_colormap()
if np.max(label) >= len(colormap):
raise ValueError('label value too large.')
return colormap[label]
def vis_segmentation(image, seg_map):
"""Visualizes input image, segmentation map and overlay view."""
plt.figure(figsize=(15, 5))
grid_spec = gridspec.GridSpec(1, 4, width_ratios=[6, 6, 6, 1])
plt.subplot(grid_spec[0])
plt.imshow(image)
plt.axis('off')
plt.title('input image')
plt.subplot(grid_spec[1])
seg_image = label_to_color_image(seg_map).astype(np.uint8)
plt.imshow(seg_image)
plt.axis('off')
plt.title('segmentation map')
plt.subplot(grid_spec[2])
plt.imshow(image)
plt.imshow(seg_image, alpha=0.7)
plt.axis('off')
plt.title('segmentation overlay')
unique_labels = np.unique(seg_map)
ax = plt.subplot(grid_spec[3])
plt.imshow(
FULL_COLOR_MAP[unique_labels].astype(np.uint8), interpolation='nearest')
ax.yaxis.tick_right()
plt.yticks(range(len(unique_labels)), LABEL_NAMES[unique_labels])
plt.xticks([], [])
ax.tick_params(width=0.0)
plt.grid('off')
plt.show()
LABEL_NAMES = np.asarray([
'background', 'aeroplane', 'bicycle', 'bird', 'boat', 'bottle', 'bus',
'car', 'cat', 'chair', 'cow', 'diningtable', 'dog', 'horse', 'motorbike',
'person', 'pottedplant', 'sheep', 'sofa', 'train', 'tv'
])
FULL_LABEL_MAP = np.arange(len(LABEL_NAMES)).reshape(len(LABEL_NAMES), 1)
FULL_COLOR_MAP = label_to_color_image(FULL_LABEL_MAP)
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="c4oXKmnjw6i_"
#@title Select and download models {display-mode: "form"}
MODEL_NAME = 'cityspace' # @param ['mobilenetv2_coco_voctrainaug', 'mobilenetv2_coco_voctrainval', 'xception_coco_voctrainaug', 'xception_coco_voctrainval']
_DOWNLOAD_URL_PREFIX = 'http://download.tensorflow.org/models/'
_MODEL_URLS = {
'mobilenetv2_coco_voctrainaug':
'deeplabv3_mnv2_pascal_train_aug_2018_01_29.tar.gz',
'mobilenetv2_coco_voctrainval':
'deeplabv3_mnv2_pascal_trainval_2018_01_29.tar.gz',
'xception_coco_voctrainaug':
'deeplabv3_pascal_train_aug_2018_01_04.tar.gz',
'xception_coco_voctrainval':
'deeplabv3_pascal_trainval_2018_01_04.tar.gz',
'cityspace':
'deeplab_cityscapes_xception71_trainfine_2018_09_08.tar.gz'
}
_TARBALL_NAME = 'deeplab_model.tar.gz'
model_dir = tempfile.mkdtemp()
tf.gfile.MakeDirs(model_dir)
download_path = os.path.join(model_dir, _TARBALL_NAME)
print('downloading model, this might take a while...')
urllib.request.urlretrieve(_DOWNLOAD_URL_PREFIX + _MODEL_URLS[MODEL_NAME],
download_path)
print('download completed! loading DeepLab model...')
# -
download_path='/tmp/tmpo9ecjb1s/deeplab_model.tar.gz'
MODEL = DeepLabModel(download_path)
print('model loaded successfully!')
# + [markdown] colab_type="text" id="SZst78N-4OKO"
# ## Run on sample images
#
# Select one of sample images (leave `IMAGE_URL` empty) or feed any internet image
# url for inference.
#
# Note that we are using single scale inference in the demo for fast computation,
# so the results may slightly differ from the visualizations in
# [README](https://github.com/tensorflow/models/blob/master/research/deeplab/README.md),
# which uses multi-scale and left-right flipped inputs.
# -
from tqdm import tqdm_notebook
# +
MT_PATH = '/data/pytorch-CycleGAN-and-pix2pix/results2/c16r8_all/test_latest/images/cityscapes_test_%d_fake_A.png'
ST_PATH = '/data/pytorch-CycleGAN-and-pix2pix/results2/cityscapes_baseline/test_latest/images/%d_fake_A.png'
import skimage.transform
def evaluate_folders(path, gts, n = 500, group=None):
#segs = []
for i in tqdm_notebook(range(1, n + 1)):
p = path % i
#print(p)
original_im = Image.open(p)
resized_im, seg_map = MODEL.run(original_im)
#segs.append(seg_map)
if '%d' % i not in group:
group.create_dataset('%d/accuracy' % i, data=accuracy(gts[i - 1], seg_map))
group.create_dataset('%d/nll' % i, data=log_prob(gts[i - 1], seg_map))
#return segs
def load_groundtruth(path):
segs = []
for i in tqdm_notebook(range(1, 500 + 1)):
p = path % i
p = p.replace('fake_A', 'real_B')
#print(p)
segs.append(np.array(Image.open(p)))
return segs
import scipy.ndimage
def accuracy(ground_truth, predictions):
predictions = np.argmax(predictions, axis=-1)
#predictions = skimage.transform.resize(predictions, (256, 256), order=0, preserve_range=True, anti_aliasing=False)
mask = (ground_truth <= 18)
return np.sum((ground_truth == predictions) * mask) / np.sum(mask)
def onehot(label, n):
s = label.shape
label = label.flatten()
y = np.zeros((label.shape[0], n))
y[np.arange(0, y.shape[0], dtype=np.int), label.astype(np.int)] = 1
return y.reshape(s + (n,))
def cross_entropy_with_logits(logits, y_true):
# y_i *log(e^x_i / \sum_i e^x_i)
logits = logits - np.max(logits)
return np.sum(logits * y_true, axis=-1) - np.log(np.sum(np.exp(logits), axis=-1))
def log_prob(ground_truth, predictions):
mask = (ground_truth <= 18)
gt = np.where(mask, ground_truth, 0)
nll = -np.sum(cross_entropy_with_logits(predictions, onehot(gt, 19)) * mask) / np.sum(mask)
return nll
# -
import h5py
import cityscapesscripts
from cityscapesscripts.helpers import labels
import numpy as np
from collections import defaultdict
class Label:
def __init__(self):
self.color_map = {}
self.id_map = defaultdict(lambda : -1)
for l in labels.labels:
self.color_map[l.trainId] = l.color
self.id_map[l.color] = l.trainId
self.color_map_arr = np.array([self.color_map[i] for i in range(19)])
def convert_image(self, label):
s = label.shape
mask = (label != 255)
label = np.where(mask, label, 0)
rgb = np.take(self.color_map_arr, label.flatten(), axis=0).reshape(s + (3,))
rgb = np.where(np.expand_dims(mask, 2), rgb, [[(0, 0, 0)]])
return rgb
def convert_label(self, rgb, th=10):
rgb = np.array(rgb)
s = rgb.shape[:2]
dist = np.linalg.norm(rgb.reshape((-1, 1, 3)) - self.color_map_arr.reshape(1, -1, 3), axis=-1)
cdist = np.min(dist, axis=-1)
label = np.argmin(dist, axis=-1)
label = np.where(cdist < th, label, 255)
return label.reshape(s)
l = Label()
gt_segs = load_groundtruth(MT_PATH)
gt_ids = [ l.convert_label(seg, th=40) for seg in gt_segs]
with h5py.File('/data/pytorch-CycleGAN-and-pix2pix/results2/eval-result.h5', 'w') as df:
ns = 500
evaluate_folders(MT_PATH, gt_ids, n = ns, group=df.require_group('mt'))
evaluate_folders(ST_PATH, gt_ids, n = ns, group=df.require_group('st'))
evaluate_folders(MT_PATH.replace('fake_A', 'real_A'), gt_ids, n = ns, group=df.require_group('real'))
import pandas as pd
with h5py.File('/data/pytorch-CycleGAN-and-pix2pix/results2/eval-result.h5', 'r') as df:
records = []
for method in df:
print(method)
for sample in df[method]:
entry = {k : v.value for k, v in df[method][sample].items()}
entry['method'] = method
entry['sample'] = sample
records.append(entry)
df = pd.DataFrame(records)
df.groupby('method').mean()
op = MODEL.graph.get_operation_by_name('ArgMax')
op
op.inputs[0]
MODEL.graph.get_tensor_by_name('ResizeBilinear_1:0')
MODEL.graph.get_tensor_by_name('ResizeBilinear:0')
MODEL.graph.get_tensor_by_name('logits/semantic/BiasAdd:0')
np.mean(list(map(accuracy, gt_ids, gt_preds)))
np.mean(list(map(accuracy, gt_ids, seg_mt))), np.mean(list(map(accuracy, gt_ids, seg_st)))
plt.figure(figsize=(10, 5))
plt.subplot(131)
path = '/data/pytorch-CycleGAN-and-pix2pix/datasets/cityscapes/testA/100_A.jpg'
path = '/data/pytorch-CycleGAN-and-pix2pix/results/c16r8_all/test_latest/images/cityscapes_testA_100_A_fake_A.png'
#path = '/data/pytorch-CycleGAN-and-pix2pix/results2/cityscapes_baseline/test_latest/images/101_A_real_A.png'
original_im = Image.open(path)
plt.imshow(original_im)
plt.subplot(132)
path = '/data/pytorch-CycleGAN-and-pix2pix/datasets/cityscapes/testB/100_B.jpg'
#path = '/data/pytorch-CycleGAN-and-pix2pix/results2/cityscapes_baseline/test_latest/images/101_A_real_B.png'
original_seg = Image.open(path)
plt.imshow(original_seg)
def run_visualization(original_im):
resized_im, seg_map = MODEL.run(original_im)
return resized_im, seg_map
#vis_segmentation(resized_im, seg_map)
rim, seg_map = run_visualization(original_im)
plt.subplot(133)
plt.imshow(l.convert_image(seg_map))
dt=np.array(Image.open('/data/pytorch-CycleGAN-and-pix2pix/datasets/cityscapes/evaluate_cityscape/frankfurt_000000_000294_leftImg8bit.jpg'))
plt.imshow(dt)
| deeplab_demo_report.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import findspark
findspark.init('/home/ubuntu/spark-2.1.1-bin-hadoop2.7')
import pyspark
from pyspark.sql import SparkSession
spark = SparkSession.builder.appName('opps').getOrCreate()
df = spark.read.csv('appl_stock.csv',inferSchema=True,header=True)
df.printSchema()
df.show()
df.head()
df.filter("Close<500").select('Open').show()
df.filter("Close<500").show()
df.filter("Close<500").select(['Open','Close']).show()
df.filter(df['Close']<500).select('Volume').show()
df.filter((df['CLose']<200) & (df['Open']>200)).show()#Two COnditions
df.filter((df['CLose']<200) & ~(df['Open']>200)).show() #Not condition ~
result= df.filter(df['Low']==197.16).collect()
result
result[0]
row=result[0]
row.asDict()
row.asDict()['Date']
| Spark_Dataframe/BasicOperation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Module 4 Practice
# ## Nested Conditionals
# <font size="5" color="#00A0B2" face="verdana"> <B>Student will be able to</B></font>
# - create nested conditional logic in code
# - print format print using escape sequence (**\**)
# #
# <font size="6" color="#B24C00" face="verdana"> <B>Tasks</B></font>
# +
# [ ] print a string that outputs the following exactly: The new line character is "\n"
# +
# [ ] print output that is exactly (with quotes): "That's how we escape!"
# +
# [ ] with only 1 print statement and using No Space Characters, output the text commented below
# 1 one
# 22 two
# 333 three
# -
# #
# ## Program: quote_me() Function
# quote_me takes a string argument and returns a string that will display surrounded with **added double quotes** if printed
# - check if passed string starts with a double quote (`"\""`), then surround string with single quotations
# - if the passed string starts with single quote, or if doesn't start with a quotation mark, then surround with double quotations
#
# Test the function code passing string input as the argument to quote_me()
# +
# [ ] create and test quote_me()
# -
# #
# ### Program: shirt order
# First get input for color and size
# - White has sizes L, M
# - Blue has sizes M, S
#
# print available or unavailable, then
# print the order confirmation of color and size
#
# *
# **hint**: set a variable "available = False" before nested if statements and
# change to True if color and size are avaliable*
# +
# [ ] create shirt order using nested if
# -
# #
# ## Program: str_analysis() Function
# Create the str_analysis() function that takes a string argument. In the body of the function:
# - Check `if` string is digits
# - if digits: convert to `int` and check `if` greater than 99
# - if greater than 99, print a message about a "big number"
# - if not greater than 99, print message about "small number"
# - if not digits: check if string isalpha
# - if isalpha print message about being all alpha
# - if not isalpha print a message about being neither all alpha nor all digit
#
# call the function with a string from user input
# +
# [ ] create and test str_analysis()
# -
# #
# ### Program: ticket_check() - finds out if a seat is available
# Call ticket_check() function with 2 arguments: *section* and *seats* requested and return True or False
# - **section** is a string and expects: general, floor
# - **seats** is an integer and expects: 1 - 10
#
# Check for valid section and seats
# - if section is *general* (or use startswith "g")
# - if seats is 1-10 return True
# - if section is *floor* (or use starts with "f")
# - if seats is 1-4 return True
#
# otherwise return False
# [ ] create and call ticket_check()
#Call ticket_check() function with 2 arguments: section and seats requested and return True or False
# # Module 4 Practice 2
# ## `while()` loops & increments
# <font size="5" color="#00A0B2" face="verdana"> <B>Student will be able to</B></font>
# - create forever loops using `while` and `break`
# - use incrementing variables in a while loop
# - control while loops using Boolean operators
# +
# [ ] use a "forever" while loop to get user input of integers to add to sum,
# until a non-digit is entered, then break the loop and print sum
sum = 0
# +
# [ ] use a while True loop (forever loop) to give 4 chances for input of a correct color in a rainbow
# rainbow = "red orange yellow green blue indigo violet"
# +
# [ ] Get input for a book title, keep looping while input is Not in title format (title is every word capitalized)
title = ""
# +
# [ ] create a math quiz question and ask for the solution until the input is correct
# -
# ### Fix the Error
# +
# [ ] review the code, run, fix the error
tickets = int(input("enter tickets remaining (0 to quit): "))
while int(tickets) > 0:
# if tickets are multiple of 3 then "winner"
if int(tickets/3) == tickets/3:
print("you win!")
else:
print("sorry, not a winner.")
tickets = int(input("enter tickets remaining (0 to quit): "))
print("Game ended")
# -
# #### create a function: quiz_item() that asks a question and tests if input is correct
# - quiz_item()has 2 parameter **strings**: question and solution
# - shows question, gets answer input
# - returns True if `answer == solution` or continues to ask question until correct answer is provided
# - use a while loop
#
# create 2 or more quiz questions that call quiz_item()
# **Hint**: provide multiple choice or T/F answers
# +
# Create quiz_item() and 2 or more quiz questions that call quiz_item()
#quiz_item()has 2 parameter strings: question and solution
def quiz_item(question, solution):
print("Question: ", question)
answer = input('Answer: ')
if answer == solution:
return True
return False
if __name__ == '__main__':
question1 = "Do you like cookies?"
answer1 = "yes"
question2 = "Do you like school?"
answer2 = "no"
question3 = "What is your favorite color?"
answer3 = "red"
# call the function for each pair
print(quiz_item(question1, answer1))
print(quiz_item(question2,answer2))
print(quiz_item(question3, answer3))
# -
# [Terms of use](http://go.microsoft.com/fwlink/?LinkID=206977) [Privacy & cookies](https://go.microsoft.com/fwlink/?LinkId=521839) © 2017 Microsoft
| Python Absolute Beginner/Module_4.1_Practice.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:dammit]
# language: python
# name: conda-env-dammit-py
# ---
# + language="bash"
#
# conda activate dammit
# dammit databases --help
# + language="bash"
#
# conda activate dammit
# dammit annotate --help
# -
# ls -lha ~/data/tick-genome/rna/pacbio-isoseq/2018-06-07/fake_genome_and_transcriptome/
# +
import glob
transcriptomes = glob.glob('~/data/tick-genome/rna/pacbio-isoseq/2018-06-07/fake_genome_and_transcriptome/*fake_ref_transcriptome.fasta')
# -
template = 'conda activate dammit && dammit annotate {transcriptome} --database-dir /mnt/data_sm/home/olga/dammit_databases/ --busco-group arthropoda --n_threads 64'
# + language="bash"
#
# BASE_DIR=$HOME/data/tick-genome/rna/pacbio-isoseq/2018-06-07/fake_genome_and_transcriptome/
#
# conda activate dammit
# dammit annotate \
# --database-dir /mnt/data_sm/home/olga/dammit_databases/ \
# --busco-group arthropoda \
# --n_threads 64
# -
| jupyter/.ipynb_checkpoints/001_run_dammit-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # CIFAR 10
#
# This example is a copy of [Convolutional Neural Network (CNN)](https://www.tensorflow.org/tutorials/images/cnn) exmample of Tensorflow.
# **It does NOT work with a Complex database** but uses this library Layers to test it's correct behaviour.
# ## Import stuff
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
from tensorflow.keras import datasets, models
import cvnn.layers as complex_layers # Ouw layers!
# ## Download and prepare the CIFAR10 dataset
#
# The CIFAR10 dataset contains 60,000 color images in 10 classes, with 6,000 images in each class. The dataset is divided into 50,000 training images and 10,000 testing images. The classes are mutually exclusive and there is no overlap between them.
(train_images, train_labels), (test_images, test_labels) = datasets.cifar10.load_data()
# Normalize pixel values to be between 0 and 1
train_images, test_images = train_images.astype(dtype=np.float32) / 255.0, test_images.astype(dtype=np.float32) / 255.0
# ## Verify the data
#
# To verify that the dataset looks correct, let's plot the first 25 images from the training set and display the class name below each image.
# +
class_names = ['airplane', 'automobile', 'bird', 'cat', 'deer',
'dog', 'frog', 'horse', 'ship', 'truck']
plt.figure(figsize=(10,10))
for i in range(25):
plt.subplot(5,5,i+1)
plt.xticks([])
plt.yticks([])
plt.grid(False)
plt.imshow(train_images[i], cmap=plt.cm.binary)
# The CIFAR labels happen to be arrays,
# which is why you need the extra index
plt.xlabel(class_names[train_labels[i][0]])
plt.show()
# -
# ## Create the convolutional base
#
# By default, complex layers have complex dtype. A difference with Tensorflow example is that we must spicitly tell the layer to be real (`float32`)
#
# The 6 lines of code below define the convolutional base using a common pattern: a stack of Conv2D and MaxPooling2D layers.
#
# As input, a CNN takes tensors of shape (image_height, image_width, color_channels), ignoring the batch size. If you are new to these dimensions, color_channels refers to (R,G,B). In this example, you will configure our CNN to process inputs of shape (32, 32, 3), which is the format of CIFAR images. You can do this by passing the argument input_shape to our first layer.
model = models.Sequential()
model.add(complex_layers.ComplexConv2D(32, (3, 3), activation='cart_relu', input_shape=(32, 32, 3), dtype=np.float32))
model.add(complex_layers.ComplexMaxPooling2D((2, 2), dtype=np.float32))
model.add(complex_layers.ComplexConv2D(64, (3, 3), activation='relu', dtype=np.float32)) # Either tensorflow ' relu' or 'cart_relu' will work
model.add(complex_layers.ComplexMaxPooling2D((2, 2), dtype=np.float32))
model.add(complex_layers.ComplexConv2D(64, (3, 3), activation='cart_relu', dtype=np.float32))
# Let's display the architecture of our model so far.
model.summary()
# Above, you can see that the output of every Conv2D and MaxPooling2D layer is a 3D tensor of shape (height, width, channels). The width and height dimensions tend to shrink as you go deeper in the network. The number of output channels for each Conv2D layer is controlled by the first argument (e.g., 32 or 64). Typically, as the width and height shrink, you can afford (computationally) to add more output channels in each Conv2D layer.
#
# ## Add Dense layers on top
#
# To complete our model, you will feed the last output tensor from the convolutional base (of shape (4, 4, 64)) into one or more Dense layers to perform classification. Dense layers take vectors as input (which are 1D), while the current output is a 3D tensor. First, you will flatten (or unroll) the 3D output to 1D, then add one or more Dense layers on top. CIFAR has 10 output classes, so you use a final Dense layer with 10 outputs.
model.add(complex_layers.ComplexFlatten())
model.add(complex_layers.ComplexDense(64, activation='cart_relu', dtype=np.float32))
model.add(complex_layers.ComplexDense(10, dtype=np.float32))
# Here's the complete architecture of our model.
model.summary()
# As you can see, our (4, 4, 64) outputs were flattened into vectors of shape (1024) before going through two Dense layers.
# ## Compile and train the model
# +
model.compile(optimizer='adam',
loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=['accuracy'])
history = model.fit(train_images, train_labels, epochs=10, validation_data=(test_images, test_labels))
# -
# ## Evaluate the model
# +
plt.plot(history.history['accuracy'], label='accuracy')
plt.plot(history.history['val_accuracy'], label = 'val_accuracy')
plt.xlabel('Epoch')
plt.ylabel('Accuracy')
plt.ylim([0.5, 1])
plt.legend(loc='lower right')
test_loss, test_acc = model.evaluate(test_images, test_labels, verbose=2)
# -
print(f"{test_acc:.2%}")
# Our simple CNN has achieved a test accuracy of over 70%. Not bad for a few lines of code!
| docs/code_examples/cifar10.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Semi-Gradient vs. Full-Gradient #
# ## DQN ##
# +
# %matplotlib inline
import numpy as np
import random
import torch
from torch import nn
import torch.nn.functional as F
from torch import optim
from tqdm import tqdm as _tqdm
def tqdm(*args, **kwargs):
return _tqdm(*args, **kwargs, mininterval=1) # Safety, do not overflow buffer
import matplotlib.pyplot as plt
import sys
import time
import gym
import datetime as datetime
import pandas as pd
assert sys.version_info[:3] >= (3, 6, 0), "Make sure you have Python 3.6 installed!"
# +
class QNetwork(nn.Module):
def __init__(self, num_hidden=128):
nn.Module.__init__(self)
self.l1 = nn.Linear(4, num_hidden)
self.l2 = nn.Linear(num_hidden, 2)
def forward(self, x):
out=self.l1(x)
out=F.relu(out)
out=self.l2(out)
return out
class ReplayMemory:
def __init__(self, capacity):
self.capacity = capacity
self.memory = []
def push(self, transition):
self.memory.append(transition)
if len(self.memory)>self.capacity:
self.memory=self.memory[1:]
def sample(self, batch_size):
return random.sample(self.memory,batch_size)
def __len__(self):
return len(self.memory)
def get_epsilon(it):
epsilon =np.maximum(1+ - .95*((it)/1000),.05)
return epsilon
class EpsilonGreedyPolicy(object):
"""
A simple epsilon greedy policy.
"""
def __init__(self, Q, epsilon):
self.Q = Q
self.epsilon = epsilon
def sample_action(self, obs):
"""
This method takes a state as input and returns an action sampled from this policy.
Args:
obs: current state
Returns:
An action (int).
"""
with torch.no_grad():
obs = torch.tensor(obs, dtype=torch.float)
q=self.Q(obs)
p=random.random()
if p<self.epsilon:
return int(len(q) * p/self.epsilon)
else:
return np.argmax(q).item()
def set_epsilon(self, epsilon):
self.epsilon = epsilon
def compute_q_vals(Q, states, actions):
try:
return torch.gather(Q.forward(states),1, actions)
except:
return Q_net.forward(states)[actions].flatten()
def compute_targets(Q, rewards, next_states, dones, discount_factor):
"""
This method returns targets (values towards which Q-values should move).
Args:
Q: Q-net
rewards: a tensor of actions. Shape: Shape: batch_size x 1
next_states: a tensor of states. Shape: batch_size x obs_dim
dones: a tensor of boolean done flags (indicates if next_state is terminal) Shape: batch_size x 1
discount_factor: discount
Returns:
A torch tensor filled with target values. Shape: batch_size x 1.
"""
dones=dones.squeeze()
ndmask=(1-dones.type(torch.FloatTensor))
targets= rewards.squeeze()+(discount_factor*torch.max(Q(next_states),1)[0])*ndmask.flatten()
return targets.reshape(len(dones),1)
def train(Q, memory, optimizer, batch_size, discount_factor,semi):
# DO NOT MODIFY THIS FUNCTION
loss_func = nn.MSELoss()
# don't learn without some decent experience
if len(memory) < batch_size:
return None
# random transition batch is taken from experience replay memory
transitions = memory.sample(batch_size)
# transition is a list of 4-tuples, instead we want 4 vectors (as torch.Tensor's)
state, action, reward, next_state, done = zip(*transitions)
state = torch.tensor(state, dtype=torch.float)
action = torch.tensor(action, dtype=torch.int64)[:, None] # Need 64 bit to use them as index
next_state = torch.tensor(next_state, dtype=torch.float)
reward = torch.tensor(reward, dtype=torch.float)[:, None]
done = torch.tensor(done, dtype=torch.uint8)[:, None] # Boolean
# compute the q value
q_val = compute_q_vals(Q, state, action)
if semi==True:
with torch.no_grad(): # Don't compute gradient info for the target (semi-gradient)
target = compute_targets(Q, reward, next_state, done, discount_factor)
else:
target = compute_targets(Q, reward, next_state, done, discount_factor)
loss = loss_func(q_val, target)
optimizer.zero_grad()
loss.backward()
optimizer.step()
return loss.item() # Returns a Python scalar, and releases history (similar to .detach())
def run_episodes(train, Q, policy, memory, env, num_episodes, batch_size, discount_factor, learn_rate, semi):
# print(episode_durations)
optimizer = optim.Adam(Q.parameters(), learn_rate)
global_steps = 0 # Count the steps (do not reset at episode start, to compute epsilon)
episode_durations = []
global_iter = 0
for i in range(num_episodes):
state = env.reset()
steps = 0
while True:
steps=steps+1
epsilon=get_epsilon(global_steps)
policy.set_epsilon(epsilon)
a = policy.sample_action(state)
ns, reward, done, _ = env.step(a)
memory.push((state, a, reward, ns, done))
train(Q, memory, optimizer, batch_size, discount_factor,semi)
state=ns
global_steps=global_steps+1
if done:
if i % 100 == 0:
print("{2} Episode {0} finished after {1} steps"
.format(i, steps, '\033[92m' if steps >= 195 else '\033[99m'))
episode_durations.append(steps)
break
return episode_durations
# +
# Let's run it!
num_episodes = 1000
batch_size = 64
#discount_factor = 0.8
learn_rate = 1e-3
memory = ReplayMemory(10000)
num_hidden = 128
env = gym.envs.make("CartPole-v1")
x=datetime.datetime.now()
for semib in [True, False]:
results={}
counter=0
for seed in range(10):
for epsilon in [0.0]:
for discount_factor in [1]:
# We will seed the algorithm (before initializing QNetwork!) for reproducibility
random.seed(seed)
torch.manual_seed(seed)
env.seed(seed)
Q_net = QNetwork(num_hidden)
policy = EpsilonGreedyPolicy(Q_net, epsilon)
episode_durations1 = run_episodes(train, Q_net, policy, memory, env, num_episodes, batch_size, discount_factor, learn_rate,semi=semib)
results[counter]={'EpisodeDuration':episode_durations1,'Seed':seed,'SemiB':semib,'Epsilon':epsilon,'DisFactor':discount_factor}
counter=counter+1
print(seed, datetime.datetime.now()-x)
results_df=pd.DataFrame(results).T
results_df.to_json('results_semi_{}.json'.format(semib))
# -
results_df
# ## Plot Results ##
# +
files = ['results_semi_True.json', 'results_semi_False.json']
n = 0
fig, axs = plt.subplots(2, sharey=True)
for file in files:
results_df = pd.read_json(file)
if results_df.shape[0] != 10:
results_df = results_df.loc[10:, :]
results_df.reset_index(drop=True, inplace=True)
tmp = []
for i in range(10):
tmp.append(results_df.loc[i, 'EpisodeDuration'])
durations_np = np.array(tmp)
print(durations_np.shape)
mean = durations_np.mean(axis=0)
std = durations_np.std(axis=0)
plt.rcParams.update({'font.size': 26})
axs[n].plot(range(len(mean)), mean, '-', color='blue')
axs[n].fill_between(range(len(mean)), mean-std, mean+std,
color='blue', alpha=0.2)
if n == 0:
axs[n].set_title('Semi-Gradient')
else:
axs[n].set_title('Full Gradient')
axs[1].set_xlabel('Episode')
axs[1].set_ylabel('Duration (ms)')
axs[n].grid(True)
axs[n].set_xlim([0, 1000])
n += 1
fig.set_size_inches(18.5, 10.5)
fig.tight_layout()
plt.savefig('cart_pole_semi.pdf', bbox_inches='tight')
# -
| DQN_polecart_cleaned.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:comoving-rv]
# language: python
# name: conda-env-comoving-rv-py
# ---
# Group:
# * <NAME>
# * <NAME>
# * <NAME>
# +
# Third-party
import matplotlib as mpl
import matplotlib.pyplot as plt
import numpy as np
plt.rcParams['axes.titlesize'] = 26
plt.rcParams['axes.labelsize'] = 22
plt.rcParams['xtick.labelsize'] = 18
plt.rcParams['ytick.labelsize'] = 18
# %matplotlib inline
from isochrones.mist import MISTModelGrid
# -
grid = MISTModelGrid(['J', 'H', 'Ks'])
# ### Goal:
#
# * Show that the color of helium-burning stars gets bluer with decreasing metallicity, redder with increased metallicity
# ## The "before" figure:
# +
fig,ax = plt.subplots(1, 1, figsize=(7,6))
iso = grid.df[(grid.df['feh'] >= -1.5) & (grid.df['feh'] <= 1) & (grid.df['log10_isochrone_age_yr'] == 10)]
c = ax.scatter(iso['J']-iso['Ks'], iso['J'], c=iso['feh'])
cb = fig.colorbar(c)
cb.set_label(r'$[{\rm Fe}/{\rm H}]$')
ax.xaxis.set_ticks(np.arange(-0.2, 1.2+0.1, 0.2))
ax.axvline(0.45, alpha=0.1)
ax.set_xlim(-0.21, 1.21)
ax.set_ylim(6, -6)
ax.set_xlabel('$J-K_s$')
ax.set_ylabel('$M_J$')
# -
# Problems with this initial figure:
#
# * Colorbar confusing / doesn't mean anything.
# * Isochrones are points instead of lines
# * Axis limits not set well
# * HB stars are not highlighted over other stars
from matplotlib.colors import Normalize, ListedColormap
iso = grid.df[(grid.df['feh'] >= -2) & (grid.df['feh'] <= 1) &
(grid.df['log10_isochrone_age_yr'] == 10) & (grid.df['phase'] < 6)]
fehs = np.unique(iso['feh'])
norm = Normalize(vmin=-2., vmax=0.5)
coolwarm = plt.get_cmap('coolwarm')
coolwarm_colors = coolwarm(np.linspace(0, 1, 256))
colors = (coolwarm_colors[:100][::1].tolist() +
[[0.8, 0.8, 0.8, 1.]] +
coolwarm_colors[-50:][::2].tolist())
mod_coolwarm = ListedColormap(colors, name='modified_coolwarm')
cmap = mod_coolwarm
# +
fig = plt.figure(figsize=(6,6))
ax = fig.add_axes([0.1, 0.1, 0.8, 0.8])
for feh in fehs:
# if feh == 0: continue
feh_iso = iso[iso['feh'] == feh]
under_iso = feh_iso #[feh_iso['phase'] != 2]
c = ax.plot(under_iso['J']-under_iso['Ks'], under_iso['J'], linewidth=2.,
marker='None', linestyle='-', color=cmap(norm(feh)), alpha=0.25)
# phase = 3 is core-helium burning
HB_feh_iso = feh_iso[feh_iso['phase'] == 3]
idx = np.array(HB_feh_iso['J']).argmax()
c = ax.plot((HB_feh_iso['J']-HB_feh_iso['Ks']).iloc[idx], HB_feh_iso['J'].iloc[idx],
marker='o', linestyle='none', color=cmap(norm(feh)),
markersize=10, markeredgecolor='k', markeredgewidth=1)
ax.xaxis.set_ticks(np.arange(-0.2, 1.2+0.1, 0.2))
ax_cb = fig.add_axes([0.1, 0.94, 0.8, 0.04])
cb = mpl.colorbar.ColorbarBase(ax_cb, cmap=cmap, norm=norm,
orientation='horizontal')
cb.set_label('[Fe/H]', labelpad=20, fontsize=18)
ax_cb.xaxis.set_ticks_position('top')
ax_cb.xaxis.set_label_position('top')
ax.set_xlim(0.1, 1.1)
ax.set_ylim(6, -6)
ax.set_xlabel('$J-K_s$')
ax.set_ylabel('$M_J$')
fig.set_facecolor('w')
| day4/Horizontal-Branch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: covid
# language: python
# name: covid
# ---
# # Metadata overview
# +
# %matplotlib inline
import matplotlib.pyplot as plt
# magics and warnings
# %load_ext autoreload
# %autoreload 2
import warnings; warnings.simplefilter('ignore')
import os, random, codecs, json
import pandas as pd
import numpy as np
seed = 99
random.seed(seed)
np.random.seed(seed)
import nltk, sklearn
import matplotlib.pyplot as plt
import seaborn as sns
sns.set(style="white")
sns.set_context("notebook", font_scale=1.2, rc={"lines.linewidth": 2.5})
# +
# load metadata
df_meta = pd.read_csv("datasets_output/df_pub.csv",compression="gzip")
df_cord = pd.read_csv("datasets_output/sql_tables/cord19_metadata.csv",sep="\t",header=None,names=[ 'cord19_metadata_id', 'source', 'license', 'full_text_file', 'ms_academic_id',
'who_covidence', 'sha', 'full_text', 'pub_id'])
df_meta.drop(columns="Unnamed: 0",inplace=True)
# -
df_meta.head()
df_meta.columns
# #### Publication years
# +
import re
def clean_year(s):
if pd.isna(s):
return np.nan
if not (s>1900):
return np.nan
elif s>2020:
return 2020
return s
df_meta["publication_year"] = df_meta["publication_year"].apply(clean_year)
# -
df_meta.publication_year.describe()
sns.distplot(df_meta.publication_year.tolist(), bins=30, kde=False)
sns.distplot(df_meta[(pd.notnull(df_meta.publication_year)) & (df_meta.publication_year > 2000)].publication_year.tolist(), bins=20, hist=True, kde=False)
# #### Null values
df_meta.shape
sum(pd.notnull(df_meta.abstract))
sum(pd.notnull(df_meta.doi))
sum(pd.notnull(df_meta.pmcid))
sum(pd.notnull(df_meta.pmid))
sum(pd.notnull(df_meta.journal))
# #### Journals
df_meta.journal.value_counts()[:30]
df_sub = df_meta[df_meta.journal.isin(df_meta.journal.value_counts()[:20].index.tolist())]
b = sns.countplot(y="journal", data=df_sub, order=df_sub['journal'].value_counts().index)
#b.axes.set_title("Title",fontsize=50)
b.set_xlabel("Count",fontsize=15)
b.set_ylabel("Journal",fontsize=15)
b.tick_params(labelsize=8)
# #### Sources and licenses
#
# For CORD19
df_cord.head()
# source
df_sub = df_cord[df_cord.source.isin(df_cord.source.value_counts()[:30].index.tolist())]
b = sns.countplot(y="source", data=df_sub, order=df_sub['source'].value_counts().index)
#b.axes.set_title("Title",fontsize=50)
b.set_xlabel("Count",fontsize=15)
b.set_ylabel("Source",fontsize=15)
b.tick_params(labelsize=15)
# license
df_sub = df_cord[df_cord.license.isin(df_cord.license.value_counts()[:30].index.tolist())]
b = sns.countplot(y="license", data=df_sub, order=df_sub['license'].value_counts().index)
#b.axes.set_title("Title",fontsize=50)
b.set_xlabel("Count",fontsize=15)
b.set_ylabel("License",fontsize=15)
b.tick_params(labelsize=15)
# #### Full text availability
df_cord["has_full_text"] = pd.notnull(df_cord.full_text)
df_cord = df_cord.merge(df_meta, how="left", left_on="pub_id", right_on="pub_id")
# full text x source
df_plot = df_cord.groupby(['has_full_text', 'source']).size().reset_index().pivot(columns='has_full_text', index='source', values=0)
df_plot.plot(kind='bar', stacked=True)
# full text x journal
df_sub = df_cord[df_cord.journal.isin(df_cord.journal.value_counts()[:30].index.tolist())]
df_plot = df_sub.groupby(['has_full_text', 'journal']).size().reset_index().pivot(columns='has_full_text', index='journal', values=0)
df_plot.plot(kind='bar', stacked=True)
# full text x year
df_sub = df_cord[(pd.notnull(df_cord.publication_year)) & (df_cord.publication_year > 2000)]
df_plot = df_sub.groupby(['has_full_text', 'publication_year']).size().reset_index().pivot(columns='has_full_text', index='publication_year', values=0)
df_plot.plot(kind='bar', stacked=True)
# ### Overlap of CORD19 with Dimensions and WHO
#
# This is still problematically low..
# +
# datasets
who_latest = "datasets_input/WHO_28_03_2020.csv"
dimensions_latest = "datasets_input/Dimensions_28_03_2020.csv"
df_who = pd.read_csv(who_latest)
df_dimensions = pd.read_csv(dimensions_latest)
# -
who_dois = df_who[pd.notnull(df_who["DOI"])]["DOI"].tolist()
dimensions_dois = df_dimensions[pd.notnull(df_dimensions["DOI"])]["DOI"].tolist()
cord_dois = df_cord[pd.notnull(df_cord["doi"])]["doi"].tolist()
len(cord_dois)
len(set(cord_dois).intersection(set(who_dois)))/len(set(who_dois))
len(set(cord_dois).intersection(set(dimensions_dois)))/len(set(dimensions_dois))
len(set(who_dois).intersection(set(dimensions_dois)))/len(set(dimensions_dois))
# ## Abstracts
abstracts = df_meta[pd.notnull(df_meta["abstract"])].abstract.tolist()
abstracts[1]
# #### Topic modelling
from tqdm import tqdm
import gensim, sklearn, spacy
import pyLDAvis.gensim
# !python -m spacy download en
nlp = spacy.load('en')
STOPWORDS = spacy.lang.en.stop_words.STOP_WORDS
# +
# %%time
processed_docs = list()
for doc in nlp.pipe(abstracts, n_threads=5, batch_size=10):
# Process document using Spacy NLP pipeline.
#ents = doc.ents # Named entities
# Keep only words (no numbers, no punctuation).
# Lemmatize tokens, remove punctuation and remove stopwords.
doc = [token.lemma_ for token in doc if token.is_alpha and not token.is_stop]
# Remove common words from a stopword list and keep only words of length 3 or more.
doc = [token for token in doc if token not in STOPWORDS and len(token) > 2]
# Add named entities, but only if they are a compound of more than one word.
#doc.extend([str(entity) for entity in ents if len(entity) > 1])
processed_docs.append(doc)
# +
import pickle
pickle.dump(processed_docs, open("datasets_output/processed_docs.pk", "wb"))
# +
import pickle
processed_docs = pickle.load(open("datasets_output/processed_docs.pk", "rb"))
# +
docs = processed_docs
del processed_docs
# Add bigrams
from gensim.models.phrases import Phrases
# Add bigrams to docs (only ones that appear several times or more). A better approach would be to use a chi_sq test.
bigram = Phrases(docs, min_count=50)
for idx in range(len(docs)):
for token in bigram[docs[idx]]:
if '_' in token:
# Token is a bigram, add to document.
docs[idx].append(token)
# -
# Remove rare and common tokens.
# Filter out words that occur too frequently or too rarely.
max_freq = 0.5
min_wordcount = 10
# +
# Create a dictionary representation of the documents, and filter out frequent and rare words.
from gensim.corpora import Dictionary
dictionary = Dictionary(docs)
dictionary.filter_extremes(no_below=min_wordcount, no_above=max_freq)
# Bag-of-words representation of the documents.
corpus = [dictionary.doc2bow(doc) for doc in docs]
#MmCorpus.serialize("models/corpus.mm", corpus)
print('Number of unique tokens: %d' % len(dictionary))
print('Number of docs: %d (%d)' % (len(corpus),len(abstracts)))
# +
# %%time
from gensim.models import LdaMulticore, LdaModel
params = {'num_topics': 15,'passes': 3, 'random_state': seed}
model = LdaModel(corpus=corpus, num_topics=params['num_topics'], id2word=dictionary, #workers=6,
passes=params['passes'], random_state=params['random_state'])
# -
model.show_topics(num_words=5, num_topics=params['num_topics'])
# plot topics (NOTE: the IDs here do not match those from the model)
data = pyLDAvis.gensim.prepare(model, corpus, dictionary)
pyLDAvis.display(data)
# +
# topics over time
# the topics of ALL the documents of our corpus
df_local = df_meta[pd.notnull(df_meta["abstract"])]
publication_years = df_local.publication_year.tolist()
dois = df_local.doi.tolist()
topics = np.zeros((len(docs),params['num_topics']))
for n,doc_topics in enumerate(model.get_document_topics(corpus)):
for t in doc_topics:
topics[n][t[0]] = t[1]
# -
topics[0,:]
model.get_document_topics(corpus[0])
df_topics = pd.DataFrame(topics)
df_topics["year"] = publication_years
df_topics["doi"] = dois
# +
from_which_year = 2000
grouped = df_topics.groupby('year')
df_grouped = grouped.aggregate(np.mean)
df_grouped = df_grouped[df_grouped.index >= from_which_year]
#df_grouped
# -
fig = plt.figure(figsize=(16, 12))
plt.pcolor(df_grouped.to_numpy(), norm=None, cmap='Blues')
plt.yticks(np.arange(df_grouped.to_numpy().shape[0]), df_grouped.index.values)
plt.xticks(np.arange(df_grouped.to_numpy().shape[1])+0.5, ["Topic #"+str(n) for n in range(model.num_topics)], rotation = 90)
plt.colorbar(cmap='Blues') # plot colorbar
plt.tight_layout() # fixes margins
plt.show()
# #### Language modelling
# +
from gensim.models import Word2Vec
import pickle
processed_docs = pickle.load(open("datasets_output/processed_docs.pk", "rb"))
model = Word2Vec(processed_docs, size=256, window=10, min_count=2, workers=6)
# -
model.wv.most_similar("coronavirus")
# #### Keyword extraction
#
# Using [RAKE](https://csurfer.github.io/rake-nltk/_build/html/index.html)
# +
from rake_nltk import Rake
# Uses stopwords for English from NLTK, and all puntuation characters by default
r = Rake(min_length=2)
# Extraction given the text
r.extract_keywords_from_text(abstracts[1])
# -
# To get keyword phrases ranked highest to lowest.
r.get_ranked_phrases()[:10]
# To get keyword phrases ranked highest to lowest with scores.
r.get_ranked_phrases_with_scores()[:10]
len(processed_docs)
r = Rake(min_length=2, max_length=3)
# Extraction given the list of strings where each string is a sentence.
r.extract_keywords_from_sentences([" ".join(d) for d in processed_docs])
# To get keyword phrases ranked highest to lowest with scores.
r.get_ranked_phrases_with_scores()[:50]
| Notebook_3_metadata_overview.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] graffitiCellId="id_w6aue5c"
# # Leafs on Binary Tree
# + [markdown] graffitiCellId="id_vtiso2s"
# Write a function to count the leafs on this binary tree.
#
#
# A
# / \
# B C
# / \
# D E
# \
# F
#
#
# + [markdown] graffitiCellId="id_c660jkr"
# #### Try finding the number of leafs on the Binary Tree
# + graffitiCellId="id_l94c0y1"
def Count_Binary_Tree_Leafs(tree):
"""
Find the number of leafs on a binary tree
Args:
tree(object): Input binary tree
Returns:
int: The number of leafs of the tree
"""
# TODO: Write function to find the number of leafs on a binary tree
pass
# + [markdown] graffitiCellId="id_1z41pj8"
# #### Test Cases
# + graffitiCellId="id_0oyn7m0"
class Tree:
def __init__(self, value, left = None, right = None):
self.left = left
self.right = right
self.value = value
def __str__(self):
return str(self.value)
f = Tree("F")
e = Tree("E", None, f)
d = Tree("D")
b = Tree("B", d, e)
c = Tree("C")
a = Tree("A", b, c)
my_tree = a
print ("Pass" if (3 == Count_Binary_Tree_Leafs(my_tree)) else "Fail")
# + [markdown] graffitiCellId="id_lbj0bd5"
# <span class="graffiti-highlight graffiti-id_lbj0bd5-id_vqtuk96"><i></i><button>Show Solution</button></span>
| trees/Leafs on Binary Tree.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # EDA
# * Pixel-level assessments of the imaging data for healthy & disease states of interest (e.g. histograms of intensity values) and compare distributions across diseases.
# * The patient demographic data such as gender, age.
# * The x-ray views taken (i.e. view position)
# * The number of cases including:
# * number of pneumonia cases,
# * number of non-pneumonia cases
# * The distribution of other diseases that are comorbid with pneumonia
# * Number of disease per patient
import os
from glob import glob
from skimage import io
from itertools import chain
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
# +
# load 'sample_labels.csv' data for pixel level assessments
sample_df = pd.read_csv('sample_labels.csv')
# build a dataframe that has 50% pneumonia cases
df1 = sample_df.loc[sample_df['Finding Labels'].str.contains('Pneumonia')].reset_index()[18:26]
df2 = sample_df.loc[sample_df['Finding Labels'].str.contains('No Finding')][:8]
df3 = pd.concat([df1, df2], axis=0)
# +
image_paths = {os.path.basename(x): x for x in glob(os.path.join('/data','images*', '*', '*.png'))}
# Visualize medical images
fig, m_axs = plt.subplots(4,4, figsize = (16, 16))
m_axs = m_axs.flatten()
imgs = df3['Image Index']
ind=0
for img, ax in zip(imgs, m_axs):
img = io.imread(image_paths[img])
ax.imshow(img,cmap='gray')
ax.set_title(df3.iloc[ind]['Finding Labels'])
ind=ind+1
# +
# visualize intensity distributions of images with different disease labels
# for normal lungs, there is usually a peak at around 150
# pneumonia together with comorbidities could exhibit a different intensity profile compared to pneumonia without comorbidities
# however, they are both different from the intensity profiles with no findings and can be used in trainings
fig, m_axs = plt.subplots(4,4, figsize = (16, 16))
m_axs = m_axs.flatten()
imgs = df3['Image Index'][-16:]
ind=0
for img, ax in zip(imgs, m_axs):
img = io.imread(image_paths[img])
ax.hist(img.ravel(), bins = 256, range=(0, 250));
ax.set_ylim(0, 20000)
ax.set_title(df3.iloc[ind]['Finding Labels'])
ind += 1
# +
# Load NIH data
all_xray_df = pd.read_csv('/data/Data_Entry_2017.csv')
# Create paths for each image
all_image_paths = {os.path.basename(x): x for x in
glob(os.path.join('/data','images*', '*', '*.png'))}
print('Scans found:', len(all_image_paths), ', Total Headers', all_xray_df.shape[0])
all_xray_df['Path'] = all_xray_df['Image Index'].map(all_image_paths.get)
# -
# OneHotCode the finding labels
all_labels = np.unique(list(chain(*all_xray_df['Finding Labels'].apply(lambda x: x.split('|')).tolist())))
all_labels = [x for x in all_labels if len(x)>0]
print('All Labels ({}): {}'.format(len(all_labels), all_labels))
for c_label in all_labels:
all_xray_df[c_label] = all_xray_df['Finding Labels'].map(lambda finding: 1 if c_label in finding else 0)
all_xray_df['Class'] = all_xray_df['Pneumonia'].apply(lambda x: 'Pneumonia' if x==1 else "Non-pneumonia")
all_xray_df.sample(3)
# examine data types and null values
all_xray_df.info()
# examine the statistics
all_xray_df.describe()
# drop the Unnamed: 11 column
all_xray_df = all_xray_df.drop(columns=['Unnamed: 11'])
# fix the column names
all_xray_df.columns = ['Image Index', 'Finding Labels', 'Follow-up #', 'Patient ID', 'Patient Age', 'Patient Gender', 'View Position', 'OriginalImageWidth', 'OriginalImageHeight', 'OriginalImagePixelSpacing_x', 'OriginalImagePixelSpacing_y' , 'Path', 'Atelectasis', 'Cardiomegaly', 'Consolidation', 'Edema','Effusion', 'Emphysema', 'Fibrosis', 'Hernia', 'Infiltration', 'Mass', 'No Finding', 'Nodule', 'Pleural_Thickening', 'Pneumonia','Pneumothorax', 'Class']
# examine and fix the data error in Patient Age
print(all_xray_df['Patient Age'].sort_values(ascending=False)[:20])
all_xray_df = all_xray_df[all_xray_df['Patient Age'] <= 120]
# gender distribution for the entire population
# the number of man is 20% more than the number of women in the population
base_color = sns.color_palette()[0]
sns.countplot(data = all_xray_df, x = 'Patient Gender', color=base_color, alpha=0.5);
# gender distribution for pneumonia patients
# the number of man is 33% more than the number of women in the population
sns.countplot(data = all_xray_df.loc[all_xray_df.Pneumonia==1], x = 'Patient Gender', color=base_color, alpha=0.5);
# age distribution for the entire population
# patient age peaks at about 60
base_color = sns.color_palette()[1]
sns.distplot(all_xray_df['Patient Age'], color=base_color);
# age distribution for pneumonia patients
# patient age peaks at about 60
sns.distplot(all_xray_df.loc[all_xray_df.Pneumonia==1, 'Patient Age'], color=base_color);
# view position distribution for the entire population
base_color = sns.color_palette()[2]
sns.countplot(data = all_xray_df, x = 'View Position', color=base_color, alpha=0.5);
# follow-up number distribution for the entire population
base_color = sns.color_palette()[3]
sns.distplot(all_xray_df['Follow-up #'], color=base_color);
# image width distribution for the entire population
base_color = sns.color_palette()[4]
sns.distplot(all_xray_df['OriginalImageWidth'], color=base_color);
# image height distribution for the entire population
base_color = sns.color_palette()[6]
sns.distplot(all_xray_df['OriginalImageHeight'], color=base_color);
# correlation between pneumonia and other diseases
# pneumonia has a relatively high correlation with Edema and Infiltration
plt.figure(figsize=(12,10))
comorbid = ['Pneumonia', 'Atelectasis', 'Cardiomegaly', 'Consolidation', 'Edema', 'Effusion', 'Emphysema', 'Fibrosis', 'Hernia', 'Infiltration', 'Mass', 'Nodule', 'Pleural_Thickening', 'Pneumothorax', 'No Finding', 'Class']
label_df = all_xray_df[comorbid]
corr_df = label_df.corr()
sns.heatmap(corr_df, cmap='coolwarm')
# disease number per patient
(label_df.drop(columns=['Class']).sum() / all_xray_df['Patient ID'].nunique()).plot.bar();
# the population has a small percentage of pneumonia patients
# which needs to be addressed when spliting the train and test samples
base_color = sns.color_palette()[8]
sns.countplot(data = label_df, x = 'Class', color = base_color);
# Pneumonia patients are likely to have edema and infiltration as comorbidities
group = label_df.groupby('Class')[[x for x in comorbid if x != 'Pneumonia']].mean()
group.plot.bar(stacked=True, figsize=(16, 10));
plt.legend(loc=0)
all_xray_df.to_csv('Data_Entry_2017_updated.csv', index=False)
| EDA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# <img src="./images/galvanize-logo.png" alt="galvanize-logo" align="center" style="width: 200px;"/>
#
# <hr />
#
# ### Introduction to High-Performance Computing
# + [markdown] slideshow={"slide_type": "slide"}
# ## Objectives
# + [markdown] slideshow={"slide_type": "fragment"}
# * Describe the right sequence to creating programs.
# + [markdown] slideshow={"slide_type": "fragment"}
# * Explain how to optimize your code.
# + [markdown] slideshow={"slide_type": "fragment"}
# * Describe commonly used techniques and tools for code optimization.
# + [markdown] slideshow={"slide_type": "slide"}
#
# * Explain why run code in parallel.
# + [markdown] slideshow={"slide_type": "fragment"}
# * Describe what is High Performance Computing (HPC), in particular paralell computing. Explain when you need HPC?
# + [markdown] slideshow={"slide_type": "slide"}
# ## The right sequence to creating programs
#
#
# + [markdown] slideshow={"slide_type": "fragment"}
# 1. Make it work
#
# 2. Ensure it is right
#
# 3. Make it fast i.e. code optimization + parallel computing
# + [markdown] slideshow={"slide_type": "notes"}
# Concentrating on the last step before the previous two can result in significantly more total work. Sometimes our programs are fast enough and we do not even need the last step.
# + [markdown] slideshow={"slide_type": "slide"}
# ## How to optimize your code
# + [markdown] slideshow={"slide_type": "fragment"}
# 1. To identify which part of you code need to be optimized. This is done using
# [profiling](https://en.wikipedia.org/wiki/Profiling_(computer_programming)) or more specifically [Python
# profilers](https://docs.python.org/3/library/profile.html).
#
# 2. look around for implementations before you build your own.
#
# >- For generic optimization: [scipy.optimize](https://docs.scipy.org/doc/scipy/reference/tutorial/optimize.html)
#
# >- For graph related problem: [algorithms implemented by NetworkX](https://networkx.github.io/documentation/stable/reference/algorithms/index.html)
#
# 3. When there is no existing implementation, consider optimizing code by some common tools.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Commonly used techniques and tools
# + [markdown] slideshow={"slide_type": "subslide"}
# - Use appropriate data containers
# For example, a Python set has a shorter look-up time than a Python list. Similarly, use dictionaries and NumPy
# arrays whenever possible.
# + [markdown] slideshow={"slide_type": "subslide"}
# - [Multiprocessing](https://docs.python.org/3/library/multiprocessing.html)
# This is a package in the standard Python library and it supports spawning processes (for each core) using an API
# similar to the threading module. The multiprocessing package offers both local and remote concurrency.
#
#
# + [markdown] slideshow={"slide_type": "subslide"}
# - [Threading](https://docs.python.org/3/library/threading.html#module-threading)
# Another package in the standard library that allows separate flows of execution at a lower level than
# multiprocessing.
#
#
# + [markdown] slideshow={"slide_type": "subslide"}
# - [Subprocessing](https://docs.python.org/3/library/subprocess.html)
# A module that allows you to spawn new processes, connect to their input/output/error pipes, and obtain their return codes.
# You may run **and control** non-Python processes like Bash or R with the subprocessing module.
#
#
# + [markdown] slideshow={"slide_type": "subslide"}
# - [Multiprocessing](https://docs.python.org/3/library/multiprocessing.html)
# This is a package in the standard Python library and it supports spawning processes (for each core) using an API
# similar to the threading module. The multiprocessing package offers both local and remote concurrency.
#
#
# + [markdown] slideshow={"slide_type": "subslide"}
# - [mpi4py](https://mpi4py.readthedocs.io/en/stable/)
# MPI for Python provides bindings of the Message Passing Interface (MPI) standard for the Python programming
# language, allowing any Python program to exploit multiple processors.
#
#
#
# + [markdown] slideshow={"slide_type": "subslide"}
# - [ipyparallel](https://ipyparallel.readthedocs.io/en/latest/)
# Parallel computing tools for use with Jupyter notebooks and IPython. Can be used with mpi4py.
#
#
# + [markdown] slideshow={"slide_type": "subslide"}
# - [Cython](https://cython.org/)
# An optimizing static compiler for both the Python programming language and the extended Cython programming language
# It is generally used to write C extensions for slow portions of code.
#
#
# + [markdown] slideshow={"slide_type": "subslide"}
# - [CUDA(Compute Unified Device Architecture)](https://en.wikipedia.org/wiki/CUDA)
# Parallel computing platform and API created by [Nvidia](https://www.nvidia.com/en-us/) for use with CUDA-enabled
# GPUs. CUDA in the Python environment is often run using the package [PyCUDA](https://documen.tician.de/pycuda/).
#
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Why run code in parallel?
#
# + [markdown] slideshow={"slide_type": "subslide"}
# * Modern computers have multiple cores and [hyperthreading](https://en.wikipedia.org/wiki/Hyper-threading)
#
# + [markdown] slideshow={"slide_type": "fragment"}
# * Graphics processing units (GPUs) have driven many of the recent advancements in data science
# + [markdown] slideshow={"slide_type": "fragment"}
# * Many of the newest *i7* processors have 8 cores
#
# + [markdown] slideshow={"slide_type": "fragment"}
# * The is a lot of **potential** but the overhead can be demanding for some problems
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## How many cores do I have?
#
# Parallel computing can help us make better use of the available hardware.
# + slideshow={"slide_type": "subslide"}
from multiprocessing import Pool, cpu_count
total_cores = cpu_count()
print('total cores: ', total_cores)
# + [markdown] slideshow={"slide_type": "slide"}
# ## High-performance computing
#
# The aggregation of compute resources to dramatically increase available compute resources is known as high-performance computing (HPC)
#
# In particular, [parallel computing](https://en.wikipedia.org/wiki/Parallel_computing)
# is what we enable by adding multiple GPUs to compuation tasks
# + [markdown] slideshow={"slide_type": "slide"}
# ## Two laws that constrain the maximum speed-up of computing:
#
# - [Amdahl's law](https://en.wikipedia.org/wiki/Amdahl%27s_law)
#
# - [Gustafson's law](https://en.wikipedia.org/wiki/Gustafson%27s_law)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Important terminologies
# + [markdown] slideshow={"slide_type": "subslide"}
# - [Symmetric multiprocessing](https://en.wikipedia.org/wiki/Symmetric_multiprocessing)
# Two or more identical processors connected to a single unit of memory.
# - [Distributed computing](https://en.wikipedia.org/wiki/Distributed_computing)
# Processing elements are connected by a network.
# - [Cluster computing](https://en.wikipedia.org/wiki/Computer_cluster)
# Group of loosely (or tightly) coupled computers that work together in a way that they can be viewed as a single system.
# - [Massive parallel processing](https://en.wikipedia.org/wiki/Massively_parallel)
# Many networked processors usually > 100 used to perform computations in parallel.
# - [Grid computing](https://en.wikipedia.org/wiki/Grid_computing)
# distributed computing making use of a middle layer to create a `virtual super computer`.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Questions to ask for data science scaling
#
# **data science scaling = code optimization + parallel computing**
#
# + [markdown] slideshow={"slide_type": "subslide"}
# * Does my service train in a reasonable amount of time given a lot more data?
#
# * Does my service predict in a reasonable amount of time given a lot more data?
#
# * Is my service ready to support additional request load?
#
# Here, **service** = **the deployed model** + **the infrastructure**
#
# + [markdown] slideshow={"slide_type": "notes"}
# It is important to think about what kind of scale is required by your model and business application in terms of which
# bottleneck is most likely going to be the problem associated with scale. These bottlenecks will depend heavily on
# available infrastructure, code optimizations, choice of model and type of business opportunity.
# + [markdown] slideshow={"slide_type": "slide"}
# #### data science scaling = code optimization + parallel computing.
#
# Example: We could use [Apache Spark](https://spark.apache.org/), a cluster-computing framework, to enable parallel computing.
| hpc/lectures/introduction-to-hpc.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (PES_AI_PIP)
# language: python
# name: pes_ai_pip
# ---
# ## WEB-SCRAPYING
# Web scraping is a computer software technique of extracting information from websites.
# This technique mostly focuses on the transformation of unstructured data (HTML format) on the web into structured data (database or spreadsheet).
# ### PROBLEM STATEMENT
# ### Crawl popular websites & create a database of Indian movie celebrities with their images and details.
# **Libraries used:**
#
# Requests library :https://requests.readthedocs.io/en/master/
# - It is a Python module which can be used for fetching URLs
#
# BeautifulSoup library:https://www.crummy.com/software/BeautifulSoup/bs4/doc/
# - Beautiful Soup is a Python library for pulling data out of HTML and XML files. It works with your favorite parser to provide idiomatic ways of navigating, searching, and modifying the parse tree.
# ### IMPORT LIBRARIES
""" lxml, re,numpy as np,pandas as pd,os, cv2,from bs4 import BeautifulSoup,from requests import get,
import matplotlib.pyplot as plt
%matplotlib inline"""
import lxml
import re
import numpy as np
import pandas as pd
import urllib
import os
import cv2
from bs4 import BeautifulSoup
from requests import get
import matplotlib.pyplot as plt
# %matplotlib inline
os.getcwd()
# ### Make the list of Urls
url1 = ["https://www.imdb.com/list/ls068010962/","https://www.imdb.com/list/ls025929404/",
"https://www.imdb.com/list/ls051665794/","https://www.imdb.com/list/ls038407562/",
"https://www.imdb.com/list/ls024256128/","https://www.imdb.com/list/ls062161896/",
"https://www.imdb.com/list/ls099354268/"]
url2 = ["https://www.imdb.com/list/ls024256128/","https://www.imdb.com/list/ls062161896/"]
# ### Create a class to extract the details from one website
class IMDB(object):
"""docstring for IMDB"""
def __init__(self, url):
super(IMDB, self).__init__()
"""fetch the webpage using the url provided using the requests library"""
page = get(url)
"""import the Beautiful soup functions to parse the data returned from the website using lxml parser"""
self.soup = BeautifulSoup(page.content, 'lxml')
def articleTitle(self):
"""return the title of the article by searching for class = **header** with h1 tag """
return self.soup.find("h1", class_="header").text.replace("\n","")
def bodyContent(self):
"""return the body of the article by searching for class **lister-item mode-detail** With div tag """
content = self.soup.find(id="main")
return content.find_all("div", class_="lister-item mode-detail")
def IndiancelebData(self):
"""fetch the body content of the page using self.bodyContent function"""
CelebFrame = self.bodyContent()
"""initialise the list of features for particular page"""
CelebName = []
CelebImageURL = []
Description = []
CelebImage = []
"""create the celeb_img folder to store folder if required
and change the directory to that particular folder to store image"""
try:
os.mkdir("celeb_img")
except:
pass
os.chdir("celeb_img")
"""searh for the box containing details of each celebrity and append the name,image Url,detail and image"""
for celeb in CelebFrame:
celebFirstLine = celeb.find("h3", class_="lister-item-header")
CelebName.append(celebFirstLine.find("a").text[:-1])
Description.append(celeb.find_all("p")[-1].text.lstrip())
for img in CelebFrame:
imgUrl = img.img['src'].split("imgurl=")[0]
CelebImageURL.append(imgUrl)
data=urllib.request.urlretrieve(imgUrl, os.path.basename(imgUrl))
image = cv2.imread(data[0])
CelebImage.append(image)
##CelebData = ["""list of celebrity features"""]
CelebData = [CelebName,CelebImageURL,Description,CelebImage]
os.chdir("../")
return CelebData
if __name__ == '__main__':
"""initialise a list for each feature and one new list to store the features"""
CelebName=[]
CelebImageURL=[]
Description = []
CelebImage = []
data = []
for i in range(len(url1)):
"""create an instance for each url in the url list and print the title of the page,
add the information extracted from the particular page onto the list initialised
"""
site1 = IMDB(url1[i])
print("Subject: ", site1.articleTitle())
newdata = site1.IndiancelebData()
CelebName = CelebName + newdata[0]
CelebImageURL =CelebImageURL + newdata[1]
Description = Description + newdata[2]
CelebImage = CelebImage + newdata[3]
data = [CelebName,CelebImageURL,Description,CelebImage]
# ### Create the dataframe
"""store the data (CelebName,CelebImageurl,description ,CelebImage) in the dataframe using pandas"""
df = pd.DataFrame({'CelebName': data[0],
'CelebImageURL': data[1],
'Description' : data[2],
'CelebImage' : data[3],
})
print(df.info())
df.head()
# ### Remove the duplicates
# +
"""print remove the duplicates based on name or even all the data"""
df['CelebName'].apply(tuple)
df = df[~df['CelebName'].apply(tuple).duplicated()]
# -
# ### Display the dataset
"""print actor's name
actors image url
about actor
image of the actor
"""
for i in range(0,5):
print("ACTOR NAME :",df.CelebName[i])
print("ACTOR IMAGE URL:",df.CelebImageURL[i])
print("ABOUT ACTOR : \n",df.Description[i])
plt.imshow(df.CelebImage[i])
plt.show()
# ### Store the dataframe in csv file
# +
"""store the dataframe in csv using pandas"""
df.to_csv('Celeb_list.csv')
# -
| IMDB WEB-SCRAPYING/Imdb web-scrapying_solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# # 📝 Exercise M5.02
#
# The aim of this exercise is to find out whether a decision tree
# model is able to extrapolate.
#
# By extrapolation, we refer to values predicted by a model outside of the
# range of feature values seen during the training.
#
# We will first load the regression data.
# +
import pandas as pd
penguins = pd.read_csv("../datasets/penguins_regression.csv")
feature_name = "Flipper Length (mm)"
target_name = "Body Mass (g)"
data_train, target_train = penguins[[feature_name]], penguins[target_name]
# -
# <div class="admonition note alert alert-info">
# <p class="first admonition-title" style="font-weight: bold;">Note</p>
# <p class="last">If you want a deeper overview regarding this dataset, you can refer to the
# Appendix - Datasets description section at the end of this MOOC.</p>
# </div>
# First, create two models, a linear regression model and a decision tree
# regression model, and fit them on the training data. Limit the depth at
# 3 levels for the decision tree.
# +
from sklearn.linear_model import LinearRegression
from sklearn.tree import DecisionTreeRegressor
linear_model = LinearRegression()
linear_model.fit(data_train, target_train)
tree = DecisionTreeRegressor(max_depth=3)
tree.fit(data_train, target_train)
# -
# Create a synthetic dataset containing all possible flipper length from
# the minimum to the maximum of the training dataset. Get the predictions of
# each model using this dataset.
# +
import numpy as np
data_test = pd.DataFrame(np.arange(data_train[feature_name].min(),
data_train[feature_name].max()),
columns=[feature_name])
# -
target_predicted_linear_regression = linear_model.predict(data_test)
target_predicted_tree = tree.predict(data_test)
# Create a scatter plot containing the training samples and superimpose the
# predictions of both models on the top.
# +
# solution
import matplotlib.pyplot as plt
import seaborn as sns
sns.scatterplot(data=penguins, x=feature_name, y=target_name,
color="black", alpha=0.5)
plt.plot(data_test[feature_name], target_predicted_linear_regression,
label="Linear regression")
plt.plot(data_test[feature_name], target_predicted_tree, label="Decision tree")
plt.legend()
_ = plt.title("Prediction of linear model and a decision tree")
# -
# Now, we will check the extrapolation capabilities of each model. Create a
# dataset containing a broader range of values than your previous dataset,
# in other words, add values below and above the minimum and the maximum of
# the flipper length seen during training.
# solution
offset = 30
data_test = pd.DataFrame(np.arange(data_train[feature_name].min() - offset,
data_train[feature_name].max() + offset),
columns=[feature_name])
# Finally, make predictions with both models on this new interval of data.
# Repeat the plotting of the previous exercise.
# solution
target_predicted_linear_regression = linear_model.predict(data_test)
target_predicted_tree = tree.predict(data_test)
sns.scatterplot(data=penguins, x=feature_name, y=target_name,
color="black", alpha=0.5)
plt.plot(data_test[feature_name], target_predicted_linear_regression,
label="Linear regression")
plt.plot(data_test[feature_name], target_predicted_tree, label="Decision tree")
plt.legend()
_ = plt.title("Prediction of linear model and a decision tree")
| notebooks/56 - trees_ex_02.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/coryroyce/code_assignments/blob/main/211104_K_Means_Shopping_Cory_Randolph.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="u8LmF1GL2kx8"
# # K Means Shopping
#
# CMPE 256
#
# <NAME>
#
# 11/4/2021
#
#
# + [markdown] id="AAYjnlUQ2xwg"
# # Prompt
# + [markdown] id="h4XFgkuw2zjV"
# Develop K Means clustering for the following dataset: This data set is to be grouped into two clusters.
#
# Please develop Python code to Cluster K = 2, K = 3 & K = 4
# + [markdown] id="ARgfSHA2Nod0"
# # Summary of Analysis
# + [markdown] id="h4hoeq2DNrAx"
# Kmeans is a relatively simple clustering algorithm to code manually (without sklearn library), and as part of the process I had to figure out a good way to graph the data using ploty and could reuse that code for the simple sklearn versions.
#
# The overall process is to randomly choose a starting center, map each point to the closest center, recalculate the centers and repeat until the centers are no longer changing.
#
# + [markdown] id="YRiHF4Bs3jg1"
# # Imports
# + id="uppRKHQf3zKa"
import numpy as np
import pandas as pd
import plotly.express as px
import plotly.graph_objects as go
from sklearn.metrics import pairwise_distances_argmin
# + [markdown] id="v_s7E8jG3KwA"
# # Data
# + [markdown] id="bUdPoJimOcKc"
# Input the data for the shopper Spending Index and Income Index as a list of pairs.
# + id="6d-GeVmZOo4M"
data = [
[1,3,5],
[2,3,4],
[3,5,6],
[4,2,6],
[5,4,5],
[6,6,8],
[7,6,2],
[8,6,3],
[9,5,6],
[10,6,7],
[11,7,2],
[12,8,5],
[13,9,1],
[14,8,2],
[15,9,6],
[16,9,1],
[17,8,3],
]
columns = ['shopper', 'spending_index', 'income_index']
# + [markdown] id="KtvBDJDheGfr"
# Convert the data into a Pandas Dataframe
# + id="mvhdPDNdOowb" colab={"base_uri": "https://localhost:8080/", "height": 175} outputId="d6559e37-787a-4754-9d20-dd261957eb9c"
df = pd.DataFrame(data = data, columns = columns)
# Set the index
df.set_index('shopper',inplace = True)
# Display the first few rows
df.head(3)
# + [markdown] id="QWcX4y2uSLYq"
# Plot the data for easy visualization
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="--p7aJAxSPmh" outputId="75032909-d9a2-458f-faa5-3427a050c54e"
fig = px.scatter(df,
x = 'spending_index',
y = 'income_index',
labels={
'spending_index' : 'Spending Index',
'income_index' : 'Income Index'},
title = {'text': 'Shopper Data',
'x': 0.5},
)
fig.show()
# + [markdown] id="ssEkcPTxTJFD"
# # Manual K Means Cluster
# + [markdown] id="3gpD61-NTkip"
# Create clusters from the data manually (i.e. without using a clustering package)
# + [markdown] id="Nvbl7xQ5ZedS"
# Convert the dataframe into a Numpy array for easier math formulas
# + id="Jo6JktfeZkct"
X = df.values
# + [markdown] id="gAdf-kXFZ7rL"
# Set the number of clusters
# + id="mdKmmyqLZ_kH"
n_clusters = 2
# + [markdown] id="RPPvaIZvCBmQ"
# 1) Randomly choos starting cluster centers
# + id="K1YrVu6IB7FH"
# Set random seed for repeatability
rng = np.random.RandomState(3)
i = rng.permutation(X.shape[0])[:n_clusters]
centers = X[i]
# + [markdown] id="Oa3dElrCaVJQ"
# 2) Assign cluster labels to each point, recalculate and repeat
# + colab={"base_uri": "https://localhost:8080/"} id="bP49uPMwaTWB" outputId="9b285a29-8e02-4154-854a-ae70c21a8c6f"
current_iteration = 1
while True:
# 2a. Assign labels based on closest center
labels = pairwise_distances_argmin(X, centers)
# 2b. Find new centers from means of points
new_centers = np.array([X[labels == i].mean(0)
for i in range(n_clusters)])
# Print out the number of itterations for reference
print(f'Currently on iteration #{current_iteration}')
current_iteration += 1
# 2c. Check for convergence
if np.all(centers == new_centers):
break
centers = new_centers
# + [markdown] id="xZrnDUgVbiPU"
# Now that the new cluster centers have been found and labels assigned, let's plot the data.
# + id="SWhnx7gVbu-h"
# Create a new Dataframe for the cluster labels
df_manual_cluster = df.copy()
df_manual_cluster['cluster_label'] = labels + 1
df_manual_cluster['cluster_label'] = df_manual_cluster['cluster_label'].apply(np.str)
# + [markdown] id="FRSMotx0cPOQ"
# Plot the cluster and overlay the cluster centers
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="nVdFsl8pcTPY" outputId="ca2a8718-5f39-4003-8c89-8656c1b5208d"
# Plot the inital data points labeled/colored by cluster
fig = px.scatter(df_manual_cluster,
x = 'spending_index',
y = 'income_index',
color = 'cluster_label',
labels={
'spending_index' : 'Spending Index',
'income_index' : 'Income Index'},
title = {'text': 'Shopper Data',
'x': 0.5},
)
# Add in the centers to the plot
fig.add_trace(go.Scatter(x = centers[:,0], y = centers[:,1],
name = 'Centers',
line= {'width':0},
marker={'size':10, 'symbol':'x', 'color':'rgb(0, 0, 0)'},
)
)
fig.show()
# + [markdown] id="TfCCj9iRjlx4"
# # K Means Clusters
# + [markdown] id="sNp82g6mjpBE"
# Create carious K-Means clusters with k = 2,3,4
# + id="CEnGyIlVi0m2"
from sklearn.cluster import KMeans
# + [markdown] id="bxkLKs7AkRHG"
# ## K = 2
# + [markdown] id="3DMh1lmgkkF6"
# Fit the kmeans clustering
# + id="ih69STJVi0ki"
kmeans = KMeans(n_clusters=2, random_state=3).fit(X)
# + [markdown] id="yydHaxJKkmo-"
# Plot the clusters
# + id="LLXAYS2Wku11"
# Create a new Dataframe for the cluster labels
df_manual_cluster = df.copy()
df_manual_cluster['cluster_label'] = kmeans.labels_ + 1
df_manual_cluster['cluster_label'] = df_manual_cluster['cluster_label'].apply(np.str)
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="qC3VlJQvkpDB" outputId="d5d55f33-bf28-4fe5-90c4-43baa76dac23"
# Plot the inital data points labeled/colored by cluster
fig = px.scatter(df_manual_cluster,
x = 'spending_index',
y = 'income_index',
color = 'cluster_label',
labels={
'spending_index' : 'Spending Index',
'income_index' : 'Income Index'},
title = {'text': 'Shopper Data',
'x': 0.5},
)
# Add in the centers to the plot
fig.add_trace(go.Scatter(x = kmeans.cluster_centers_[:,0], y = kmeans.cluster_centers_[:,1],
name = 'Centers',
line= {'width':0},
marker={'size':10, 'symbol':'x', 'color':'rgb(0, 0, 0)'},
)
)
fig.show()
# + [markdown] id="alphPo9Ql44t"
# ## K = 3
# + [markdown] id="MMSK99WMl441"
# Fit the kmeans clustering
# + id="n8pCOh9wl441"
kmeans = KMeans(n_clusters=3, random_state=3).fit(X)
# + [markdown] id="L31Xibtpl441"
# Plot the clusters
# + id="v1nCIDjal441"
# Create a new Dataframe for the cluster labels
df_manual_cluster = df.copy()
df_manual_cluster['cluster_label'] = kmeans.labels_ + 1
df_manual_cluster['cluster_label'] = df_manual_cluster['cluster_label'].apply(np.str)
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="keQIGIdDl441" outputId="6bbe2e11-ec28-4263-88e6-2c49382b0e0a"
# Plot the inital data points labeled/colored by cluster
fig = px.scatter(df_manual_cluster,
x = 'spending_index',
y = 'income_index',
color = 'cluster_label',
labels={
'spending_index' : 'Spending Index',
'income_index' : 'Income Index'},
title = {'text': 'Shopper Data',
'x': 0.5},
)
# Add in the centers to the plot
fig.add_trace(go.Scatter(x = kmeans.cluster_centers_[:,0], y = kmeans.cluster_centers_[:,1],
name = 'Centers',
line= {'width':0},
marker={'size':10, 'symbol':'x', 'color':'rgb(0, 0, 0)'},
)
)
fig.show()
# + [markdown] id="LC85fTIcl6E3"
# ## K = 4
# + [markdown] id="QTmaIlhIl6E4"
# Fit the kmeans clustering
# + id="_JOoG-7Ul6E4"
kmeans = KMeans(n_clusters=4, random_state=3).fit(X)
# + [markdown] id="xiuiZ-R0l6E4"
# Plot the clusters
# + id="FgA5N0tWl6E4"
# Create a new Dataframe for the cluster labels
df_manual_cluster = df.copy()
df_manual_cluster['cluster_label'] = kmeans.labels_ + 1
df_manual_cluster['cluster_label'] = df_manual_cluster['cluster_label'].apply(np.str)
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="vGIrqgeul6E4" outputId="791fd4ad-c41b-4996-981c-916a76cfde5d"
# Plot the inital data points labeled/colored by cluster
fig = px.scatter(df_manual_cluster,
x = 'spending_index',
y = 'income_index',
color = 'cluster_label',
labels={
'spending_index' : 'Spending Index',
'income_index' : 'Income Index'},
title = {'text': 'Shopper Data',
'x': 0.5},
)
# Add in the centers to the plot
fig.add_trace(go.Scatter(x = kmeans.cluster_centers_[:,0], y = kmeans.cluster_centers_[:,1],
name = 'Centers',
line= {'width':0},
marker={'size':10, 'symbol':'x', 'color':'rgb(0, 0, 0)'},
)
)
fig.show()
# + [markdown] id="m8BXz0dGi5OY"
# # Reference
# + [markdown] id="_Sb1_DIzi7RL"
# Example of manual clustering as [reference](https://jakevdp.github.io/PythonDataScienceHandbook/05.11-k-means.html)
#
# Example of Kmeans in [Scikit Learn](https://scikit-learn.org/stable/modules/generated/sklearn.cluster.KMeans.html?highlight=kmeans#sklearn.cluster.KMeans)
#
# [Plotly Scatter Plot](https://plotly.com/python/line-and-scatter/) examples
| 211104_K_Means_Shopping_Cory_Randolph.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import numpy as np
import vaex as vx
import pylab
# %matplotlib inline
#server = vx.server("localhost")
#names = server.list_datasets()
server = vx.server("localhost", port=9000)
list = server.datasets()
print list
ds = list[0]
reload(vx)
reload(vx.dataset)
server = vx.server("localhost", port=9000)
list = server.datasets()
ds = list[0]
subspace = ds("x", "y")
limits = subspace.limits_sigma(square=True)
print limits
grid = subspace.histogram(limits=limits, weight="10 + sqrt(x**2+y**2)")
subspace.figlarge()
subspace.plot(np.log(grid+1), limits=limits, cmap="afmhot")
# +
import ipy_autoreload
# %load_ext autoreload
# %autoreload 2
# -
# %aimport vaex
# %aimport vaex.dataset
# %autoreload?
import autoreloadload
| examples/example_server.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import random
from random import *
from collections import defaultdict
suits = ['h', 'd', 'c', 's']
values = ['2', '3', '4', '5', '6', '7', '8', '9', '10', 'j', 'q', 'k', 'a']
deck = [(value, suit) for suit in suits for value in values]
class Card:
def __init__(self, value, suit):
self.value = value
self.suit = suit
deck
def dealCards(numPlayers, numCardsForDeal):
hands = defaultdict(list)
shuffledDeck = sample(deck, 52)
i = 0
for x in range(numPlayers * numCardsForDeal):
hands[i].append(shuffledDeck[0])
shuffledDeck.pop(0)
i += 1
if i > (numPlayers - 1):
i = 0
print(hands)
dealCards(2,8)
# +
class Game:
def __init__(self, name, numPlayers, numCardsForDeal):
self.name = name
self.numPlayers = numPlayers
self.numCardsForDeal = numCardsForDeal
values = ['2', '3', '4', '5', '6', '7', '8', '9', '10', 'j', 'q', 'k', 'a']
suits = ['h', 'd', 'c', 's']
self.deck = [(value, suit) for suit in suits for value in values]
def shuffleDeck(self):
try:
self.shuffledDeck = sample(self.shuffledDeck, len(self.shuffledDeck))
return self.shuffledDeck
except:
self.shuffledDeck = sample(self.deck, 52)
return self.shuffledDeck
def dealCards(self):
self.hands = defaultdict(list)
i = 0
for x in range(self.numPlayers * self.numCardsForDeal):
self.hands[i].append(self.shuffledDeck[0])
self.shuffledDeck.pop(0)
i += 1
if i > (self.numPlayers - 1):
i = 0
return self.hands
def drawCard(self, playerPosition):
self.hands[playerPosition].append(self.shuffledDeck[0])
self.shuffledDeck.pop(0)
def playCards(self, playerPosition, cardValues):
self.inPlayCards = defaultdict(list)
for cardValue in cardValues:
if cardValue in hands[playerPosition]:
self.inPlayCards[playerPosition].append(cardValue)
self.hands.pop(r'{cardValue}')
return self.inPlayCards
# -
Golf1 = Game("Golf", 3, 8)
Golf1.dealCards()
Golf1.shuffleDeck()
Golf1.drawCard(2)
Golf1.hands[2]
Golf1.shuffledDeck
| .ipynb_checkpoints/cardDealWorking-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Assignment
import pandas as pd
import numpy as np
# Link for Datasets
# (https://drive.google.com/drive/folders/105ftuIwN9kqyPNEEm3E6IM7LqywjyvJa?usp=sharing)
# Q21. Write a pandas program to import three datasheets from a given
# excel data (coalpublic2013.xlsx ) in to a single dataframe.
#
# Note: Structure of three datasheets are same.
# +
df1 = pd.read_excel('coalpublic2013.xlsx', sheet_name='Sheet-1')
df2 = pd.read_excel('coalpublic2013.xlsx', sheet_name='Sheet-2')
df3 = pd.read_excel('coalpublic2013.xlsx', sheet_name='Sheet-3')
df = pd.concat([df1, df2, df3], axis = 0)
df
# -
# Q 22. Write a pandas program to import three datasheets from a given
# excel data (employee.xlsx ) into a single data frame and export the
# result into new Excel file.
#
# Note: Structure of three datasheets are same.
df1 = pd.read_excel('employee.xlsx', sheet_name='Sheet1')
df2 = pd.read_excel('employee.xlsx', sheet_name='Sheet2')
df3 = pd.read_excel('employee.xlsx', sheet_name='Sheet3')
df = pd.concat([df1, df2, df3])
df.to_excel('employee-1.xlsx', index = False)
# Q23. Write a pandas program to create the Pivot table with multiple
# indexes from the data set of the titanic.csv.
df = pd.read_csv('titanic.csv')
print(pd.pivot_table(df, index = ['sex', 'age'], aggfunc=np.sum))
# Q24. Write a Pandas program to create the Pivot table and find survival
# rate by gender?
df = pd.read_csv('titanic.csv')
pd.pivot_table(df, index=['sex'], values=['survived'],aggfunc=np.sum)
# Q25. Write a pandas program to make partition each of the passengers
# into 4 categories based on their age.
#
# Note: Age categories- (0, 10), (10, 30), (30, 60), (60, 80)
df = pd.read_csv('titanic.csv')
result = pd.cut(df['age'], bins= [0, 10, 30, 60, 80])
print(result)
# Q26. Write a pandas program to create the Pivot table and find survival
# rate by the gender, age of the different categories of various
# classes.
df = pd.read_csv('titanic.csv')
age = pd.cut(df['age'], bins = [0, 20, 55])
df1 = df.pivot_table('survived', index=['sex', age], columns='class')
df1
# Q27. Write a pandas program to create the Pivot table and calculate
# number of women and men were in a particular cabin class.
#
df = pd.read_csv('titanic.csv')
df1 = df.pivot_table(index=['sex'], columns=['pclass'], aggfunc='count')
df1
# Q28. Write a pandas program to create the Pivot table and separate
# the gender according to whether they travelled alone or not to get
# the probability of survival
#
df = pd.read_csv('titanic.csv')
df1 = df.pivot_table('survived', index=['sex', 'alone'])
df1
# Q29. Write a pandas program to create the Pivot table and find the
# probability of survival by class, gender, solo boarding, and the port
# of embarkation.
df = pd.read_csv('titanic.csv')
df1 = df.pivot_table('survived', index=['sex', 'alone'], columns=['embark_town', 'class'])
print(df1)
# Q30. Write a pandas program to get current date, oldest date and
# number of days between Current date and the oldest date of Ufo
# dataset.
df = pd.read_csv(r'ufo-1.csv')
df['Date_time'] = df['Date_time'].astype('datetime64[ns]')
print("Original Dataframe:")
print(df.head())
print("\nCurrent date of Ufo dataset:")
print(df.Date_time.max())
print("\nOldest date of Ufo dataset:")
print(df.Date_time.min())
print("\nNumber of days between Current date and oldest date of Ufo dataset:")
print((df.Date_time.max() - df.Date_time.min()).days)
# Q31. Write a pandas program to get all sighting days of the
# unidentified flying object (ufo) between 1950-10-10 and 1960-10-
# 10.
selected_period = df[(df['Date_time'] >= '01-01-1950 00:00:00') & (df['Date_time'] <= '31-12-1960 23:59:59')]
selected_period
# Q32. Write a Pandas program to extract the year, month, day, hour,
# minute, second, and weekday from unidentified flying object (UFO)
# reporting date.
print('Years')
print(pd.DatetimeIndex(df['Date_time']).year)
print('\nMonth')
print(pd.DatetimeIndex(df['Date_time']).month)
print('\nDay')
print(pd.DatetimeIndex(df['Date_time']).day)
print('\nHour')
print(pd.DatetimeIndex(df['Date_time']).hour)
print('\nMinute')
print(pd.DatetimeIndex(df['Date_time']).minute)
print('\nSecond')
print(pd.DatetimeIndex(df['Date_time']).second)
print('\nWeekday')
print(pd.DatetimeIndex(df['Date_time']).weekday_name)
# Q33. Write a pandas program to count year-country wise frequency of
# reporting dates of the unidentified flying object(UFO).
#
df['Year'] = pd.DatetimeIndex(df['Date_time']).year
result = df.groupby(['Year', 'country']).size()
print('count year-country wise frequency of reporting dates of the unidentified flying object(UFO)')
print(result)
# Q34. Write a pandas program to get the difference (in days) between
# documented date and reporting date of unidentified flying object
# (UFO).
df.head(1)
df['Date_time'] = pd.to_datetime(df['Date_time'])
df['date_documented'] = pd.to_datetime(df['date_documented'])
df['Difference'] = (df['date_documented'] - df['Date_time']).dt.days
print(df.head())
# Q35. Write a pandas program to generate sequences of fixedfrequency dates and time spans.
# +
dtr = pd.date_range('2019-01-01', periods=12, freq='H')
print('Hourly Frequence')
print(dtr)
dtr = pd.date_range('2019-01-01', periods=12, freq='min')
print('\nMinutely Frequence')
print(dtr)
dtr = pd.date_range('2019-01-01', periods=12, freq='S')
print('\nSecondly Frequence')
print(dtr)
# -
# Q36. Write a pandas program to manipulate and convert date times
# with timezone information.
dt = pd.date_range('2019-01-01', periods=12, freq='H')
dt = dt.tz_localize('UTC')
print(dt)
print("\nFrom UTC to Asia/India:")
dt = dt.tz_convert('Asia/Kolkata')
print(dt)
# Q37. Write a pandas program to create the graphical analysis of UFO
# (unidentified flying object) Sightings year.
# +
df['Date_time'] = pd.to_datetime(df['Date_time'], errors='coerce')
df['date_documented'] = pd.to_datetime(df['date_documented'])
df['year'] = pd.DatetimeIndex(df['Date_time']).year
years_data = df['year'].value_counts()
years_index = years_data.index
year_values = years_data.get_values()
# -
# %matplotlib inline
plt.figure(figsize=(15,8))
plt.xticks(rotation = 60)
plt.title('UFO Sightings by year')
plt.xlabel('Year')
plt.ylabel('Number of reports')
plt.bar(years_index, year_values)
# Q38. Write a pandas program to create a comparison of the top 10
# years in which the (UFO) was sighted VS each Month.
def is_top_years(year):
if year in most_sightings_years.index:
return year
# +
df = pd.read_csv(r'ufo-1.csv')
df['Date_time'] = pd.to_datetime(df['Date_time'], errors='coerce')
columns=df['Date_time'].dt.month
index=df['Date_time'].dt.year.apply(is_top_years)
aggfunc='count'
values='city'
columns = columns.get_values()# columns.reset_index(level=1, drop=False,inplace=False)
month_vs_year = df .pivot_table(columns=columns,index=index, aggfunc='count',values='city')
print("\nComparison of the top 10 years in which the UFO was sighted vs each month:")
print(month_vs_year.head(10))
# -
# Q39. Write a pandas program to create a heatmap (rectangular data as
# a colour-encoded matrix) for comparison of top 10 years in
# which (UFO ) was sighted VS each Month.
# +
df =pd.read_csv(r'ufo-1.csv')
df['Date_time'] = pd.to_datetime(df['Date_time'])
df['year'] = pd.DatetimeIndex(df['Date_time']).year
years_data = df['year'].value_counts()
most_sightings_years = years_data.head(10)
def is_top_years(year):
if year in most_sightings_years.index:
return year
columns=df['Date_time'].dt.month
index=df['Date_time'].dt.year.apply(is_top_years)
aggfunc='count'
values='city'
columns = columns.get_values()# columns.reset_index(level=1, drop=False,inplace=False)
month_vs_year = df .pivot_table(columns=columns,index=index, aggfunc='count',values='city')
month_vs_year.columns = month_vs_year.columns.astype(int)
print("\nHeatmap for comparison of the top 10 years in which the UFO was sighted vs each month:")
plt.figure(figsize=(10,8))
plt.imshow(month_vs_year, cmap='hot', interpolation='nearest')
plt.show()
# -
# Q40. Write a pandas program to create a Timewheel of Hour VS Year
# comparison of the top 10 years in which the (UFO) was sighted
import matplotlib.pyplot as plt
import matplotlib as mpl
import matplotlib.cm as cm
def is_top_years(year):
if year in most_sightings_years.index:
return year
# +
most_sightigs_yeara = df['Date_time'].dt.year.value_counts().head(10)
columns=df['Date_time'].dt.month
index=df['Date_time'].dt.year.apply(is_top_years)
columns = columns.get_values()
month_vs_year = df.pivot_table(columns=columns,
index=index,
aggfunc='count',
values='city'
)
month_vs_year.index = month_vs_year.index.astype(int)
month_vs_year.columns = month_vs_year.columns.astype(int)
# -
print("\nComparison of the top 10 years in which the UFO was sighted vs each month:")
def pie_heatmap(table, cmap='coolwarm_r', vmin=None, vmax=None,inner_r=0.25, pie_args={}):
n, m = table.shape
vmin= table.min().min() if vmin is None else vmin
vmax= table.max().max() if vmax is None else vmax
centre_circle = plt.Circle((0,0),inner_r,edgecolor='black',facecolor='white',fill=True,linewidth=0.25)
plt.gcf().gca().add_artist(centre_circle)
norm = mpl.colors.Normalize(vmin=vmin, vmax=vmax)
cmapper = cm.ScalarMappable(norm=norm, cmap=cmap)
for i, (row_name, row) in enumerate(table.iterrows()):
labels = None if i > 0 else table.columns
wedges = plt.pie([1] * m,radius=inner_r+float(n-i)/n, colors=[cmapper.to_rgba(x) for x in row.values],
labels=labels, startangle=90, counterclock=False, wedgeprops={'linewidth':-1}, **pie_args)
plt.setp(wedges[0], edgecolor='grey',linewidth=1.5)
wedges = plt.pie([1], radius=inner_r+float(n-i-1)/n, colors=['w'], labels=[row_name], startangle=-90, wedgeprops={'linewidth':0})
plt.setp(wedges[0], edgecolor='grey',linewidth=1.5)
plt.figure(figsize=(8,8))
plt.title("Timewheel of Hour Vs Year",y=1.08,fontsize=30)
pie_heatmap(month_vs_year, vmin=-20,vmax=80,inner_r=0.2)
# ## Great Job!
| Deep Learning/Subjective Assignment - 9 - Pandas 2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: me
# language: python
# name: me
# ---
# ## Model 1b: Single-country policy simulation
#
# The objective of this model-based simulation is to analyse the impact of policy, technology, and commodity changes on consumer price inflation in selected countries. The simulation environment is learnt from real data, after which simulations using synthetic data are used to do policy analysis by manipulating a number of selected variables such as government debt, cellular subscription, gdp growth, and real interest rates in the synthetic data. A secondary purpose of the simulation model is to identify and map the interactions between world-level and country-level indicator variables.
#
# #### Features
# ------------
#
# Human and technological development indicator timeseries for a country x.
#
# #### Labels
# ----------
#
# Consumer price inflation levels.
#
# #### Training
# ------------
#
# Training is done on a feature - single country basis.
# ### Load and prepare the data
# +
import warnings
import numpy as np
import pandas as pd
import seaborn as sns
import tensorflow as tf
import tensorflow_probability as tfp
import matplotlib.pyplot as plt
from tensorflow import keras
# %matplotlib inline
# -
warnings.filterwarnings('ignore')
pd.options.display.float_format = '{:20,.4f}'.format
sns.set_style("whitegrid")
sns.set_palette("colorblind")
tf.__version__
country = 'Switzerland'
# #### Load and combine the features and labels
features_df = pd.read_csv('features/m_one/%s_features.csv' % country, sep=';', header=0)
labels_df = pd.read_csv('features/m_one/labels_interpolated.csv', sep=';', header=0)
features_df.head()
labels_df.head()
combined_df = pd.concat([features_df, labels_df.drop(columns=['date'])], axis=1)
combined_df.head()
# +
fig, ax = plt.subplots(figsize=(15,7))
[sns.lineplot(x='date', y=c, markers=True, ax=ax, label=c, data=combined_df) for c in list([country, 'lending interest rate', 'real interest rate', 'inflation', 'gross domestic savings', 'government debt service'])]
xticks=ax.xaxis.get_major_ticks()
for i in range(len(xticks)):
if i % 12 == 1:
xticks[i].set_visible(True)
else:
xticks[i].set_visible(False)
ax.set_xticklabels(combined_df['date'], rotation=45);
# -
combined_df.columns
# ### Prepare the country features
base_feature_df = combined_df[['date', 'bank capital to assets ratio', 'bank nonperforming loans', 'cereal yield',
'energy imports', 'food exports', 'high-tech exports', 'inflation',
'lending interest rate', 'life expectancy', 'population density', 'real interest rate',
'broad money', 'exports of goods and services', 'gross domestic savings',
'high-tech value added', 'household consumption expenditure',
'imports of goods and services', 'listed companies', 'manufacturing value added',
'r and d spend', 'services trade', 'trade', 'government debt service',
'government interest payments external debt', 'government tax revenue', 'birth deaths',
'broadband subscriptions', 'electricity access', 'co2 emissions',
'electricity consumption', 'mobile subscriptions', 'newborns', 'overweight',
'rural population', 'urban population', country]]
base_feature_df.to_csv('features/m_one/combined_country_level_%s.csv' % country.lower(), sep=',', index=False)
base_feature_df['label'] = base_feature_df[country].shift(periods=1)
base_df = base_feature_df.drop(country, axis=1).fillna(0.00);
base_df.set_index('date')
num_obs = len(base_df)
num_cols = len(base_df.columns)
num_features = len(base_df.columns) - 1
# ### Model iterations
# ---------------------
# ### Exploration 0
#
# **ARIMA** fitted on the real data.
from statsmodels.tsa.arima_model import ARIMA
from sklearn.metrics import mean_squared_error
ar_params = {
'lag': 4,
'difference': 2,
'moving_average': 1
}
# ARIMA dataprep
# +
exo_cols = ['energy imports', 'broad money', 'real interest rate', 'household consumption expenditure',
'exports of goods and services']
ar_endo = base_df['label'].values
ar_exo = base_df[exo_cols].values
# +
fig, ax = plt.subplots(figsize=(15,7))
[sns.lineplot(x='date', y=c, markers=True, ax=ax, label='cpi in %s' % c, data=combined_df) for c in list([country])]
xticks=ax.xaxis.get_major_ticks()
for i in range(len(xticks)):
if i % 12 == 1:
xticks[i].set_visible(True)
else:
xticks[i].set_visible(False)
ax.set_xticklabels(combined_df['date'], rotation=45);
# -
len(ar_endo)
ar_endo_train, ar_endo_test = ar_endo[0:550], ar_endo[551:696]
ar_exo_train, ar_exo_test = ar_exo[0:550], ar_exo[551:696]
ar_exo_test[0]
# Fit the ARIMA model
arima = ARIMA(ar_endo, order=(ar_params['lag'], ar_params['difference'], ar_params['moving_average']), exog=ar_exo)
arima_fitted = arima.fit()
arima_fitted.summary()
# #### Evaluate the ARIMA predictions
# +
preds = []
obs = []
hist = [x for x in ar_endo_train]
exo_hist = [x for x in ar_exo_train]
for t in range(len(ar_endo_test)):
m = ARIMA(hist, order=(ar_params['lag'], ar_params['difference'], ar_params['moving_average']))
m_fit = m.fit()
yhat = m_fit.forecast()[0][0]
preds.append(yhat)
hist.append(ar_endo_test[t])
exo_hist.append(ar_exo_test[t])
if t % 50 == 0:
print('obs: %s, pred: %s' % (ar_endo_test[t], yhat))
# -
predictions = list(map(lambda x: 0.00 if np.isnan(x) else x, preds))
mean_squared_error(ar_endo_test, predictions)
plt.plot(ar_endo_test)
plt.plot(preds, color='green')
# ### Exploration 1
#
# **Multivariate LSTM** fitted on the real data, see https://machinelearningmastery.com/multivariate-time-series-forecasting-lstms-keras/
# - Activation function: Leaky ReLU.
# - Loss function: mean squared error.
# - Optimizer: adam.
# - Num observations source dataset: 684 (using lagshift, 1960-2016 inclusive monthly)
# - Num sequences (@ sequence length 6): 116.
# - Batch size: 4-8 sequences (although `size=48` would lead to more stable training)
lstm_params = {
'sequence_length': 4,
'batch_size': 8,
'num_epochs': 50,
'num_units': 512,
'lrelu_alpha': 0.35
}
# #### LSTM features
lstm_df = base_df[['bank nonperforming loans', 'lending interest rate', 'real interest rate',
'gross domestic savings', 'household consumption expenditure', 'energy imports',
'exports of goods and services', 'imports of goods and services', 'services trade', 'trade',
'broad money', 'government interest payments external debt', 'government tax revenue']]
num_lstm_cols = len(lstm_df.columns)
# +
features = []
labels = []
for i in range(int(num_obs - 1)):
labels_df = base_df['label']
labels.append(labels_df[i:(i+lstm_params['sequence_length'])].values[-1:])
features.append(lstm_df[i:(i+lstm_params['sequence_length'])].values)
# -
len(features[550:])
lstm_train_X = np.asarray(features[0:550])
lstm_train_X = lstm_train_X.reshape((lstm_train_X.shape[0], lstm_params['sequence_length'], num_lstm_cols))
lstm_train_y = np.asarray(labels[0:550])
lstm_train_y = lstm_train_y.reshape((lstm_train_y.shape[0]))
lstm_test_X = np.asarray(features[550:650])
lstm_test_X = lstm_test_X.reshape((lstm_test_X.shape[0], lstm_params['sequence_length'], num_lstm_cols))
lstm_test_y = np.asarray(labels[550:650])
lstm_test_y = lstm_test_y.reshape((lstm_test_y.shape[0]))
X = np.asarray(features[0:650])
X = X.reshape((X.shape[0], lstm_params['sequence_length'], num_lstm_cols))
y = np.asarray(labels[0:650])
y = y.reshape((y.shape[0], 1))
print('X: %s, y: %s' % (X.shape, y.shape))
# #### Model: LSTM
model = tf.keras.models.Sequential()
model.add(tf.keras.layers.LSTM(lstm_params['num_units'],
input_shape=(lstm_params['sequence_length'], num_lstm_cols)))
model.add(tf.keras.layers.Dense(1, activation=tf.keras.layers.LeakyReLU(alpha=lstm_params['lrelu_alpha'])))
model.compile(loss='mse', optimizer='adam')
model.summary()
early_stopping = tf.keras.callbacks.EarlyStopping(monitor='loss', mode='min', patience=2)
train_run = model.fit(lstm_train_X, lstm_train_y, epochs=lstm_params['num_epochs'],
batch_size=lstm_params['batch_size'], callbacks=[early_stopping])
plt.plot(train_run.history['loss'], label='train')
plt.legend()
plt.show()
# ##### Evaluate model performance
model.evaluate(lstm_test_X, lstm_test_y)
yhat = model.predict(lstm_test_X)
plt.figure(figsize=(15,7))
plt.plot(lstm_test_y, label='observed')
plt.plot(yhat, label='predicted')
plt.legend()
plt.title('Observed versus predicted values for consumer price inflation in %s' % country)
plt.show()
print('rmse: %s\nmean observed: %s\nmean predicted: %s' % (np.sqrt(mean_squared_error(lstm_test_y, yhat)),
np.mean(lstm_test_y), np.mean(yhat)))
# ## Exploration 2
# --------------------
#
# **GAN** to generate training data, **LSTM** trained on generated data validated on the real data.
# ### Conditional GAN for policy-constrained timeseries generation
#
# See https://arxiv.org/pdf/1706.02633.pdf.
from tensorflow.keras.models import Sequential, Model
from tensorflow.keras.layers import Input
from tensorflow.keras.optimizers import Adam
from sklearn.metrics import mean_squared_error
gan_df = base_df[['bank nonperforming loans', 'lending interest rate', 'real interest rate',
'gross domestic savings', 'household consumption expenditure', 'exports of goods and services',
'imports of goods and services', 'services trade', 'trade', 'broad money',
'government interest payments external debt', 'government tax revenue']]
gan_df.shape
gan_cols = gan_df.shape[1]
gan_params = {
'num_epochs': 100,
'save_interval': 100,
'sequence_length': 4,
'num_variables': gan_cols,
'batch_size': 8,
'lr': 0.001
}
generator_params = {
'noise_sigma': 0.2,
'lstm_units': 128,
'lstm_dropout': 0.3,
'gru_units': 64,
'gru_dropout': 0.2,
'lr': 0.001
}
discriminator_params = {
'bi_lstm_units': 64,
'dropout_rate': 0.4,
'lr': 0.001
}
# #### GAN input sequences
#
# The collated World Bank and IMF data used as input for the data generator and to validate the model trained on generated data.
# +
gan_features = []
gan_labels = []
for i in range(int(num_obs)):
gan_labels_df = base_df['label']
gan_labels.append(gan_labels_df[i:(i+gan_params['sequence_length'])].values[-1:])
gan_features.append(gan_df[i:(i+gan_params['sequence_length'])].values)
# -
real = np.asarray(gan_features[0:650])
real = real.reshape((real.shape[0], gan_params['sequence_length'], gan_cols))
real.shape
# #### Generator
def build_encoder(params):
gshape = params['sequence_length'], params['num_variables']
inputs = tf.keras.layers.Input(shape=(gshape))
e = tf.keras.models.Sequential(name='encoder')
e.add(tf.keras.layers.LSTM(params['lstm_units'], input_shape=(gshape), return_sequences=True))
e.add(tf.keras.layers.Dropout(params['lstm_dropout']))
e.add(tf.keras.layers.GaussianNoise(stddev=params['noise_sigma']))
e.add(tf.keras.layers.BatchNormalization(axis=2, momentum=0.8, epsilon=0.01))
e.add(tf.keras.layers.Dense(params['num_variables'], activation='relu'))
e.summary()
return tf.keras.models.Model(inputs, e(inputs))
encoder = build_encoder({**gan_params, **generator_params})
def build_generator(params):
gshape = params['sequence_length'], params['num_variables']
inputs = tf.keras.layers.Input(shape=(gshape))
g = tf.keras.models.Sequential(name='generator')
g.add(tf.keras.layers.GRU(params['gru_units'], input_shape=(gshape), return_sequences=True))
g.add(tf.keras.layers.Dropout(params['gru_dropout']))
g.add(tf.keras.layers.Dense(params['num_variables'], activation='tanh'))
g.add(tf.keras.layers.Reshape(target_shape=(gshape)))
g.summary()
return tf.keras.models.Model(inputs, g(inputs))
generator = build_generator({**gan_params, **generator_params})
# #### Discriminator
def build_discriminator(params):
dshape = params['sequence_length'], params['num_variables']
batch_shape = params['batch_size'], params['sequence_length'], params['num_variables']
real = tf.keras.layers.Input(shape=(dshape))
generated = tf.keras.layers.Input(shape=(dshape))
inputs = tf.keras.layers.concatenate([generated, real], axis=1)
d = tf.keras.models.Sequential(name='discriminator')
d.add(tf.keras.layers.Bidirectional(
tf.keras.layers.LSTM(params['bi_lstm_units']), batch_input_shape=(batch_shape)))
d.add(tf.keras.layers.Dropout(params['dropout_rate']))
d.add(tf.keras.layers.Dense(1, activation='sigmoid'))
d.summary()
return tf.keras.models.Model([generated, real], d(inputs))
discriminator = build_discriminator({**gan_params, **discriminator_params})
discriminator.compile(loss='binary_crossentropy',
optimizer=tf.keras.optimizers.Adam(lr=discriminator_params['lr']), metrics=['accuracy'])
# #### GAN
#
# Bidirectional generative adversarial network, viz https://arxiv.org/abs/1605.09782.
def build_gan(encoder, generator, discriminator, params):
ganshape = params['sequence_length'], params['num_variables']
discriminator.trainable = False
noise = tf.keras.layers.Input(shape=(ganshape))
generated = generator(noise)
data = tf.keras.layers.Input(shape=(ganshape))
encoded = encoder(data)
fake = discriminator([noise, generated])
real = discriminator([encoded, data])
gan = tf.keras.models.Model([noise, data], [fake, real], name='gan')
gan.summary()
return gan
gan = build_gan(encoder, generator, discriminator, gan_params)
gan.compile(loss=['binary_crossentropy', 'mean_squared_error'],
optimizer=tf.keras.optimizers.Adam(lr=generator_params['lr']), metrics=['accuracy', 'mse'])
def train_gan(real, batch_size, params):
g_metrics = []
d_real_metrics = []
d_synth_metrics = []
reals = np.ones(batch_size)
synths = np.zeros(batch_size)
for i in range(params['num_epochs']):
# create input of real and synthetic data
random_index = np.random.randint(0, len(real) - batch_size)
half_real = real[random_index:int(random_index + batch_size)]
half_synth = np.random.normal(-1.0, 1.0, size=[batch_size, params['sequence_length'], real.shape[2]])
# apply generator and encoder
generated = generator.predict(half_synth)
encoded = encoder.predict(half_real)
# train discriminator
d_real = discriminator.train_on_batch([encoded, half_real], reals)
d_synth = discriminator.train_on_batch([half_synth, generated], synths)
# train gan
gen_ = gan.train_on_batch([generated, encoded], [reals, synths])
if i % 10 == 0:
print('Epoch %s losses: discriminator real: %.4f%%, discriminator synth: %.4f%%, generator: %.4f%%' %
(i, d_real[0], d_synth[0], gen_[0]))
d_real_metrics.append(d_real)
d_synth_metrics.append(d_synth)
g_metrics.append(gen_)
return d_real_metrics, d_synth_metrics, g_metrics
d_r_metrics, d_s_metrics, g_metrics = train_gan(real, gan_params['batch_size'], gan_params)
plt.figure(figsize=(15,7))
plt.plot([metrics[0] for metrics in d_r_metrics], label='discriminator loss on reals')
plt.plot([metrics[0] for metrics in d_s_metrics], label='discriminator loss on synths')
plt.plot([metrics[0] for metrics in g_metrics], label='generator loss')
plt.legend()
plt.title('GAN losses')
plt.show()
plt.figure(figsize=(15,7))
plt.plot([metrics[1] for metrics in d_r_metrics], label='discriminator accuracy reals')
plt.plot([metrics[1] for metrics in d_s_metrics], label='discriminator accuracy synths')
plt.plot([metrics[1] for metrics in g_metrics], label='generator mean average error')
plt.legend()
plt.title('GAN performance metrics')
plt.show()
generated_y = generator.predict(np.random.rand(num_obs, gan_params['sequence_length'], gan_cols))[:,-1,-1]
gan_y = base_df['label'].values
plt.figure(figsize=(15,7))
plt.plot(gan_y, label='observed cpi')
plt.plot(generated_y, label='gan-generated cpi')
plt.legend()
plt.title('Observed versus GAN-generated values for consumer price inflation in %s' % country)
plt.show()
print('rmse: %s\nmean observed: %s\nmean generated: %s' % (np.sqrt(mean_squared_error(gan_y, generated_y)),
np.mean(gan_y), np.mean(generated_y)))
# ## Exploration 3
# --------------------
#
# **Sequence transformer network** to generate training data, **LSTM** trained on generated data validated on the real data. See https://arxiv.org/abs/1808.06725
| experiments/univariate_country_level_models.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# default_exp countrycodes
# -
# export
__doc__ = """Country codes module"""
import pandas as pd
from fastcore.test import test_eq
from nbdev.showdoc import show_doc
# # Country Codes
# > Class `CountryCodes()` to search and convert between `iso` and `fips` country codes.
# ## Solution to a problem
# Joshua Project uses **FIPS** codes (*US Federal Information Processing Standard)* for countries.
#
# >[FIPS 10-4](https://www.wikiwand.com/en/FIPS_10-4) (April 1995) -- Countries, Dependencies, Areas of Special Sovereignty, and Their
# Principal Administrative Divisions.
#
# However, FIPS 10-4 was [withdrawn by NIST](https://www.nist.gov/system/files/documents/itl/FIPSCodesReplacementChart2015.pdf) on September 2, 2008 in favor of the international [**ISO 3166** standard](https://www.iso.org/iso-3166-country-codes.html).
#
# In the meantime, as of Jan 2020 Joshua Project has no plans to switch onto ISO codes, I was notified. *sigh*
#
# Sadly, FIPS in **60%** cases differs from the ISO.
#
# This library provides a `CountryCodes()` class to workaround this discrepancy.
#
# Wherever the library uses the term `iso code`, **ISO 3166 alpha 2** country code is implied as defined here: https://www.iso.org/obp/ui/
from joshuaproject.countrycodes import CountryCodes
# export
class CountryCodes():
"""Returns FIPS and ISO 3166 alpha 2 country codes for `cname` and converts between."""
def __init__(self):
self.data = pd.read_csv('data/country_codes.csv', skiprows=3, names=['cname', 'FIPS', 'ISO'])
def __len__(self):
"""Returns lenth of `data`."""
return len(self.data)
def __repr__(self):
"""Returns a `str` representing the `data`."""
return f'{self.data.to_string(max_rows = None)}'
def head(self, rows=10):
"""Returns first `rows`."""
return self.data.head(rows)
def tail(self, rows=10):
"""Returns last `rows`."""
return self.data.tail(rows)
def __getitem__(self, idx):
"""Return `idx`th element of data."""
return self.data.iloc[idx]
def like(self, cname: str, mx: int = None)->list:
"""Returns a list of dict with `mx` entries with country name like `cname`."""
res = self.data[self.data['cname'].str.contains(cname)].to_dict(orient='record')
if not res: return []
if (mx is not None) and (len(res) > mx):
assert mx > 0, "CountryCodes().like: `mx` argument must be positive or `None`."
return res[:mx]
return res
def __call__(self, cname: str)->str:
"""Returns `fips` code of `cname`."""
return self.like(cname, mx=1)[0]['FIPS']
def fips(self, cname: str)->str:
"""Returns `fips` code of `cname`."""
return self.__call__(cname)
def iso(self, cname: str)->str:
"""Returns `iso` code of `cname`."""
return self.like(cname, mx=1)[0]['ISO']
def name(self, cname: str)->str:
"""Returns a `str` of the first of country name like `cname`."""
return self.like(cname, mx=1)[0]['cname']
show_doc(CountryCodes)
cc = CountryCodes()
test_eq(len(cc),len(cc.data))
len(cc)
show_doc(CountryCodes.head)
cc.head(5)
cc.data[cc.data.FIPS.isna()]
show_doc(CountryCodes.tail)
cc.tail(5)
show_doc(CountryCodes.__getitem__)
cc[101]
test_eq(cc[101].cname, 'Iceland')
test_eq(cc[101].FIPS,'IC')
test_eq(cc[101].ISO, 'IS')
# Also supports slicing.
cc[10:13]
show_doc(CountryCodes.like)
cc.like('United')
# +
test_eq(cc.like('Russia'),[{'cname': 'Russian Federation', 'FIPS': 'RS', 'ISO': 'RU'}])
test_eq(cc.like('United',2),
[{'cname': 'Tanzania, United Republic of', 'FIPS': 'TZ', 'ISO': 'TZ'},
{'cname': 'United Arab Emirates', 'FIPS': 'AE', 'ISO': 'AE'}])
test_eq(cc.like('Non-existing'),[])
# -
show_doc(CountryCodes.iso)
test_eq(cc.iso('Russia'), 'RU')
show_doc(CountryCodes.fips)
# Note that two forms of calls are possible:
test_eq(cc.fips('Russia'), 'RS')
test_eq(cc('Russia'), 'RS')
# ## Export -
#export
#hide
from nbdev.export import notebook2script
notebook2script()
| 01_countrycodes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import pickle
import pandas as pd
import numpy as np
from sklearn.preprocessing import LabelEncoder
import xgboost as xgb
# +
TEST_DATA_ROOT = '~/workspace/datarobot-user-models/tests/testdata'
BINARY_DATA = os.path.join(
TEST_DATA_ROOT, 'iris_binary_training.csv'
)
REGRESSION_DATA = os.path.join(
TEST_DATA_ROOT, 'boston_housing.csv'
)
MULTICLASS_DATA = os.path.join(
TEST_DATA_ROOT, 'skyserver_sql2_27_2018_6_51_39_pm.csv'
)
bin_X = pd.read_csv(BINARY_DATA)
bin_y = bin_X.pop('Species')
reg_X = pd.read_csv(REGRESSION_DATA)
reg_y = reg_X.pop('MEDV')
multi_X = pd.read_csv(MULTICLASS_DATA)
multi_y = multi_X.pop('class')
# +
bin_target_encoder = LabelEncoder()
bin_target_encoder.fit(bin_y)
bin_dtrain = xgb.DMatrix(bin_X, bin_target_encoder.transform(bin_y))
bin_model = xgb.train({'objective':'binary:logistic'}, bin_dtrain)
reg_dtrain = xgb.DMatrix(reg_X, reg_y)
reg_model = xgb.train({'objective':'reg:squarederror'}, reg_dtrain)
multi_target_encoder = LabelEncoder()
multi_target_encoder.fit(multi_y)
multi_dtrain = xgb.DMatrix(multi_X, multi_target_encoder.transform(multi_y))
multi_model = xgb.train({'objective':'multi:softprob', 'num_class': len(multi_target_encoder.classes_)}, multi_dtrain)
# -
bin_dtest = xgb.DMatrix(bin_X)
print(bin_model.predict(bin_dtest))
reg_dtest = xgb.DMatrix(reg_X)
print(reg_model.predict(reg_dtest))
multi_dtest = xgb.DMatrix(multi_X)
print(multi_model.predict(multi_dtest))
FIXTURE_ROOT = '~/workspace/datarobot-user-models/tests/fixtures/drop_in_model_artifacts'
with open(os.path.expanduser(os.path.join(FIXTURE_ROOT, 'xgb_bin.pkl')), 'wb') as picklefile:
pickle.dump(bin_model, picklefile)
with open(os.path.expanduser(os.path.join(FIXTURE_ROOT, 'xgb_reg.pkl')), 'wb') as picklefile:
pickle.dump(reg_model, picklefile)
with open(os.path.expanduser(os.path.join(FIXTURE_ROOT, 'xgb_multi.pkl')), 'wb') as picklefile:
pickle.dump(multi_model, picklefile)
| tests/fixtures/drop_in_model_artifacts/XGBoost.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# +
#remotes::install_github("coolbutuseless/ggpattern")
list.of.packages <- c("ggpubr","magick","ggpattern","tidyverse","stargazer","dplyr","ggplot2")
new.packages <- list.of.packages[!(list.of.packages %in% installed.packages()[,"Package"])]
if(length(new.packages)) install.packages(new.packages, repos = "http://cran.us.r-project.org")
invisible(lapply(list.of.packages, library, character.only = TRUE))
select <- dplyr::select
options(repr.matrix.max.rows=50, repr.matrix.max.cols=500)
# -
# ### Database one row each paper
path <- "../5_Final_databases/output/database_one_row_each_paper.csv"
df <- read_csv(path)
sprintf("%i x %i dataframe", nrow(df), ncol(df))
head(df,1)
df_pub <- df %>%
select(publication_year,Region)%>%
#for easier reading aggregate all pub year before 2000 to 2000
mutate(pub_year = ifelse(publication_year <= 2002, 2002, publication_year))%>%
mutate(Region=factor(Region, levels = c('Antarctica','Oceania','Africa','Latin America',
'North America','European Union','Europe','Asia'))) %>%
#calculate the sum of publi by region
group_by(pub_year,Region) %>%
summarise('number'=n()) %>%
ungroup() %>%
#calculate percentage for column labels
mutate('relative'=unlist(by(data = number, INDICES = pub_year,
FUN = function(x) round(x/sum(x)*100, digits = 0)))) %>%
mutate(Estimated = ifelse(pub_year == 2002, "aggregated", "yearly"))
# +
options(repr.plot.width=10, repr.plot.height=10)
plot <- ggplot(data = df_pub,aes(x=pub_year,y=number,fill=Region, pattern = Estimated)) +
geom_bar_pattern(stat="identity",
color = "black",
pattern_fill = "black",
pattern_angle = 45,
pattern_density = 0.1,
pattern_spacing = 0.02,
pattern_key_scale_factor = 0.2) +
scale_pattern_manual(values = c(aggregated = "stripe", yearly = "none")) +
labs(x = " \n Horizon Year", y = "Number of Papers \n ", fill = " Region", pattern = "Level") +
guides(pattern = FALSE, fill = guide_legend(override.aes = list(pattern = "none")))+
geom_vline(xintercept= 2002.5, linetype="dashed", size=0.5)+
annotate("text", x = 2002, y = 200, label = "Until 2002",angle = 90) +
xlab(" \n Publication Year")+
ylab("Number of Papers \n ")+
scale_fill_manual(values=c('Asia'='darkorange',
'European Union'='#7CAE00',
'Europe'='seagreen4',
'North America'='darkblue',
'Latin America'='dodgerblue2',
'Africa'='orchid',
'Oceania'='coral2',
'Antarctica'='#CAB2D6')) +
geom_text(data = subset(df_pub,pub_year ==2002 & relative >=15),
aes(x = pub_year, label = paste0(relative,'%')),
colour = 'black', position=position_stack(vjust=0.5))+
geom_text(data = subset(df_pub,pub_year >=2007 & pub_year <=2013 & relative >=15),
aes(x = pub_year, label = paste0(relative,'%')),
colour = 'black', position=position_stack(vjust=0.5))+
geom_text(data = subset(df_pub,pub_year >=2014 & pub_year <=2016 & relative >=5),
aes(x = pub_year, label = paste0(relative,'%')),
colour = 'black', position=position_stack(vjust=0.5))+
geom_text(data = subset(df_pub,pub_year >=2017 & pub_year <=2020 & relative >=2),
aes(x = pub_year, label = paste0(relative,'%')),
colour = 'black', position=position_stack(vjust=0.5))+
theme_minimal()+
theme(
legend.title = element_text(size = 16,face ="bold"),
legend.text = element_text(size = 16),
legend.position = 'top',
axis.text.x = element_text(size = 16),
axis.text.y = element_text(size = 16),
axis.title.x = element_text(size = 16, hjust = 0.5,face ="bold"),
axis.title.y = element_text(size = 16, hjust = 0.5,face ="bold")
)
plot
# -
ggsave('./output/Fig_SI1_pub_year_distrib.png', height=10, width=10, plot=plot)
| 6_Figures/R_Fig_SI1_Publi_Horizon_Years_Regions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <table width="100%"><tr style="background-color:white;">
# <td style="text-align:left;padding:0px;width:142px'">
# <a href="https://qworld.net" target="_blank">
# <img src="qworld/images/QWorld.png"></a></td>
# <td width="*"> </td>
# <!-- ############################################# -->
# <td style="padding:0px;width:90px;">
# <img align="right" src="qworld/images/follow_us.png" height="40px"></td>
# <td style="padding:0px;width:40px;">
# <a href="https://twitter.com/QWorld19" target="_blank">
# <img align="right" src="qworld/images/Twitter.png" width="40px"></a> </td>
# <td style="padding:0px;width:5px;"></td>
# <td style="padding:0px;width:40px;">
# <a href="https://www.facebook.com/qworld19/" target="_blank">
# <img align="right" src="qworld/images/Fb.png"></a></td>
# <td style="padding:0px;width:5px;"></td>
# <td style="padding:0px;width:40px;">
# <a href="https://www.linkedin.com/company/qworld19" target="_blank">
# <img align="right" src="qworld/images/LinkedIn.png"></a></td>
# <td style="padding:0px;width:5px;"></td>
# <td style="padding:0px;width:40px;">
# <a href="https://youtube.com/QWorld19?sub_confirmation=1" target="_blank">
# <img align="right" src="qworld/images/YT.png"></a></td>
# <!-- ############################################# -->
# <td style="padding:0px;width:60px;">
# <img align="right" src="qworld/images/join.png" height="40px"></td>
# <td style="padding:0px;width:40px;">
# <a href="https://discord.gg/akCvr7U87g"
# target="_blank">
# <img align="right" src="qworld/images/Discord.png"></a></td>
# <!-- ############################################# -->
# <td style="padding:0px;width:72px;">
# <img align="right" src="qworld/images/w3.png" height="40px"></td>
# <td style="padding:0px;width:40px;">
# <a href="https://qworld.net" target="_blank">
# <img align="right" src="qworld/images/www.png"></a></td>
# </tr></table>
# <h1 align="left" style="color: #cd7f32;"> Contents </h1>
# ### Prerequisites
#
# These notebooks are intended to be a Mathematical Introduction to Quantum Computing, mainly aimed to first years university students. The student is expected to have gone thru a first course on Linear Algebra and have some exposure to Trigonometry, Probability Theory and Complex Numbers. All exercises can be worked out with pen and paper, so no special software is required beyond the ability to run Jupyter Notebooks.
# ### Introduction
#
# [Motivation](1-intro/Motivation.ipynb) |
# [Logic Gates](1-intro/LogicGates.ipynb)
# ### Mathematics for Quantum Computing
#
# [Hilbert Space](2-math/HilbertSpace.ipynb) |
# [Bloch Sphere & Qubits](2-math/BlochSphere.ipynb) |
# [Tensor Product](2-math/TensorProduct.ipynb) |
# [Linear Operators](2-math/LinearOperators.ipynb) |
# [Spectral Theory](2-math/SpectralTheory.ipynb) |
# [Heisenberg Principle](2-math/HeisenbergPrinciple.ipynb) |
# [Pauli Operators](2-math/PauliOperators.ipynb)
# ### A (Very) Short Introduction to Quantum Mechanics
#
# [Quantum Mechanics Postulates](3-qm/Postulates.ipynb) |
# [Time Evolution Operator](3-qm/TimeEvolution.ipynb) |
# [Worked Example](3-qm/WorkedExample.ipynb) |
# [Projective Measurement](3-qm/ProjectiveMeasurement.ipynb)
# ### Quantum Entanglement
#
# [Einstein-Podolsky-<NAME>](4-entangle/EPR-Paradox.ipynb) |
# [Quantum Entanglement](4-entangle/Entanglement.ipynb) |
# [Bell's Inequality](4-entangle/Bell.ipynb)
# ### Quantum Gates
#
# [Quantum Logic Gates](5-gates/Gates.ipynb) |
# [Hadamard Gate](5-gates/Hadamard.ipynb) |
# [Rotation Gates](5-gates/Rotation.ipynb) |
# [Single-Qubit Gates](5-gates/SingleQubit.ipynb) |
# [Multi-Qubit Gates](5-gates/MultiQubit.ipynb) |
# [Solovay-Kitaev Theorem](5-gates/Solovay-Kitaev.ipynb)
| START.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# cd ..
# +
import src.plotting as plot
from src.helpers import read_video
import matplotlib.pyplot as plt
from src.plotting import plot_continues_line, plot_bodypart_lines, plot_bodypoint, stick_movie, \
BODYPART_LINES, DEFAULT_BODYPARTS, COLORS
from src.settings import LYING_VIDEOS_DATA_FOLDER, SITTING_VIDEOS_DATA_FOLDER
# %load_ext autoreload
# %autoreload 2
# -
df = read_video('006', LYING_VIDEOS_DATA_FOLDER)
# +
# set plot variables
size = (12,14)
bodypart_lines_frame = 25
# initialise plot
fig = plt.figure(figsize=size)
# plot the bodypart lines for one frame
plot_bodypart_lines(df, frame = bodypart_lines_frame)
# plot blue colors on the wrists
plot_bodypoint(df, 'wrist1', 'bo', alpha=1, frame = bodypart_lines_frame)
plot_bodypoint(df, 'wrist2', 'bo', alpha=1, frame = bodypart_lines_frame)
# plot all bodyparts from whole video
for bodypart, color in zip(DEFAULT_BODYPARTS, COLORS+COLORS):
for i in range(len(df)):
plot_bodypoint(df, bodypart, color, frame=i)
# plot the average bodypart lines, in dashed black
plot_bodypart_lines(df, bodypart_lines_colors = ['ko--']*12)
# plot continues lines between shoulders and hips
plot_continues_line(df, 'shoulder1', 'hip1')
plot_continues_line(df, 'shoulder2', 'hip2')
# invert y-axis
plt.gca().invert_yaxis()
# -
# %matplotlib tk
stick_movie(df[:100])
| notebooks/feature-visualisation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Exploring Kickoff Data
# <NAME>
#
# 1. [Introduction](#1)
# 2. [Exploratory Data Analysis](#2)
# - [Statistics](#2a)
# - [Issues](#2b)
# - [Grouping Metrics](#2c)
#
# <a id='1'>
# ### Introduction
#
# **A Foreword: generate_DF.py**
#
# First, I should mention how we got to the data we have here. Most of the data scraping that creates the kickoff dataframe can be seen in *pbp_boxscores.ipynb*. This *generate_DF.py* file is a Python script that runs the *make_DF* function over a directory of ten csv files that contain boxscores of NFL games for each year of the 2010's. As such, the code that is unique to that script is generally working in reading in the different files, with a few notable exceptions. First, when reading in a csv, I added a column to the dataframe that kept track of which season the kickoff occurred during. This information is easily obtained by searching the filename string for four consecutive numbers. There are a number of other small changes I made, feel free to check them out yourself!
#
# After running the generate_DF.py script, we now have a CSV file of around 28,000 kickoffs. This should be enough to make some basic observations about kickoffs over the past 10 years. In the next section, we'll use measures of central tendancy, graphs, and other measurements in order to come to some conclusions about kickoffs over the past decade.
# <a id='2'>
# ### Exploratory Data Analysis
#
# ***Statistics***
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
import csv
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import re
import nltk
df = pd.read_csv("kickoff_dataset.csv")
df.columns
print(f"We've tracked {len(df.columns)-1} statistics over {len(df)} kickoffs. Here's a glimpse of the dataset:")
df.head()
df.columns
#Kicking and Receiving Teams
len(set(df["kicking_team"]))
len(set(df["receiving_team"]))
set(df["kicking_team"]) == set(df["receiving_team"])
set(df["receiving_team"])
plot = sns.histplot(data=df, x="receiving_team")
plt.xticks(rotation=45)
#So, we have 35 teams. We know there are only 32. What's the anomaly? This data is over ten years, so we have leftover
#abbreviations like STL, LA, JAC, SD, OAK, etc. You can see this borne out in that the smallest bars are new name changes.
# +
#Kickers
len(set(df["kicker"]))
df["kicker"].value_counts()[:-20]
#What we learned from this is that our scraping of kickers is not very good! We need to go back and fix.
# +
#Returners
len(set(df["returner"]))
df["returner"].value_counts()[:-20]
#I'm no expert, but 1120 sounds abouuuut right? 1120 returns/32 teams/10 years of returns = 3-4 returners per team per year.
# +
#penaltyType
len(set(df["penaltyType"]))
df["penaltyType"].value_counts()
#looks good
# +
#Years
sns.countplot(data=df, x="year")
#This looks good! You wouldn't expect any one year to have more than another.
# +
#isTouchback, isOutOfBounds, isOnside, isFairCatch, isReturned, isTouchdown, isMuff, retainsMuff, isPenalty, isFumble, retainsFumble
touchback = len(df[df["isTouchback"]])/len(df)
print(f"{touchback*100}% of kicks were touchbacks.")
oob = len(df[df["isOutOfBounds"]])/len(df)
print(f"{oob*100}% of kicks were out of bounds.")
fc = len(df[df["isFairCatch"]])/len(df)
print(f"{fc*100}% of kicks were fair catches.")
returned = len(df[df["isReturned"]])/len(df)
print(f"{returned*100}% of kicks were returned.")
td = len(df[df["isTouchdown"]])/len(df)
print(f"{td*100}% of kicks were returned for a touchdown.")
muff = len(df[df["isMuff"]])/len(df)
print(f"{muff*100}% of kicks were muffed.")
cut = df[df["isMuff"]]
rm = len(cut[cut["retainsMuff"]])/len(cut)
print(f"{rm*100}% of muffs were retained.")
penalty = len(df[df["isPenalty"]])/len(df)
print(f"{penalty*100}% of kickoffs had an accepted penalty.")
fumble = len(df[df["isFumble"]])/len(df)
print(f"{fumble*100}% of kicks were fumbled.")
cut = df[df["isFumble"]]
rf = len(cut[cut["retainsFumble"]])/len(cut)
print(f"{rf*100}% of fumbles were retained.")
#These numbers look to be about right. One immediate comparison we can do is fumbles lost and muffs kicks vs. touchdowns.
# +
#kickYards, kickStart, kickLand, returnSpot, returnLand, finalSpot
np.mean(df["kickYards"])
np.std(df["kickYards"])
sns.displot(data=df, x="kickYards", kind="hist")
plt.show()
#almost every kick is between 60-70 yards
np.mean(df["kickStart"])
np.std(df["kickStart"])
sns.displot(data=df, x="kickStart", kind="hist")
plt.show()
#almost all kicks start at either 35 or 30. to be expected here. if you notice, seaborn actually used less bins b/c so many 0 values
np.mean(df["kickLand"])
np.std(df["kickLand"])
sns.displot(data=df, x="kickLand", kind="hist")
plt.show()
#this mirrors kickYards very well
np.mean(df["returnYards"])
np.std(df["returnYards"])
sns.displot(data=df, x="returnYards", kind="hist")
plt.show()
#mirrors return spot, good.
np.mean(df["returnSpot"])
np.std(df["returnSpot"])
sns.displot(data=df, x="returnSpot", kind="hist")
plt.show()
#looks good, you would expect this distribution with the exception of the huge number of 100's, which are all touchbacks.
#filter these out when you look at the yardage game
np.mean(df["finalSpot"])
np.std(df["finalSpot"])
sns.displot(data=df, x="finalSpot", kind="hist")
plt.show()
# +
len(df[df["kickYards"]==67])/len(df)
len(df[df["kickYards"]==66])/len(df)
len(df[df["kickYards"]==65])/len(df)
len(df[df["kickYards"]==64])/len(df)
len(df[df["kickYards"]==63])/len(df)
#There's something to suggest that the recording of boxscores is actually incorrect here. Why should kicks land on the end zone
#yard exactly 10x more likely than a yard before or after? This is something to consider in the yardage game.
# -
# <a id="2b">
# ***Issues***
#
# 1. The *kickYards* statistic is incorrect for a portion of the dataset. Since there is no line in boxscore descriptions that contains the final spotting of the ball, we have to construct this statistic. This calculation is sometimes very simple--for touchbacks, it's as easy as just saying "25 yards". On the other hand, it can get very complicated: consider an example where there is a penalty, fumbled ball, recovered ball, and then another penalty. I'm not sure what I'm going to do with this statistic at this time.
# 2. The *kickLand* statistic is also out of wack, but for reasons more unclear than *finalSpot*. About 80% of kicks land exactly on the goal line, according to our database. However, we know that this cannot possibly be. In fact, upon watching football and following along with the ESPN app, I saw a >65 yard kick be reported as "65 yards, Touchback". When we consider that we see the exact same pattern for all of the touchbacks in our data (which make up about half of the dataframe), we must assume that any touchback is reported as landing on the goal line. Let's test this theory.
# 3. The *kickers* statistic is scraped incorrectly. This is the first data I scraped, and for some reason I thought I could just take anything before the word "kick", so I used *box_score.split("kick")* as my rule. This is, and I can't understate this, bad. Will fix soon.
#
# All of these are yet to be addressed, but I'm a little sick of data wrangling. Let's do some analysis of game altering plays while I rebuild a tolerance for RE.
#
# <a id='2c'>
# ***Grouping Metrics***
# Here, I'm going to group the more intensive analysis into groups.
#
# 1. *Game Altering Plays* - These metrics include fumbles, touchdowns, and muffs. In this section, we should compare the rates of very impactful kickoff successes (touchdowns) with very impactful kickoff failures (lost possessions such as lost fumbles and muffs, as well as touchdowns scored by the kicking team off of fumbles). I have a suspicion that these plays will be more failures than successes.
# 2. *The Yardage Game* - This group is the largest and is made up of the meatiest metrics: kickLand, returnYards, finalSpot, and more. For this group, it will be important to answer where the ball was fielded from, and where the ball is spotted after the play. In this section, we should see how often it is that the returner made it back to the 25 yard line, where a touchback would have placed them. It is important to note, however, that kicks fielded right in front of the endzone and forward don't come with the option of a touchback or return.
# 3. *Players, Teams, Years* - This group doesn't explicitly answer whether kicks should be returned or not, but it may be worth exploring. One hugely important detail here is that the NFL changed the touchback rule from possession at the 20 to possession at the 25 in 2016. It may be worth examining how this affected the game.
| code/.ipynb_checkpoints/exploring_kickoff_data-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # PR-027 Glove
#
# 곽근봉 님의 [Glove 강의](https://www.youtube.com/watch?v=uZ2GtEe-50E&list=PLlMkM4tgfjnJhhd4wn5aj8fVTYJwIpWkS&index=28) 감사드립니다.
#
# word embedding이 처음이시라면 word2vector.ipynb 부터 보시는 걸 추천드립니다.
#
# Glove를 이용해 간단한 문장을 학습시킵니다.
#
# [tensorflow-glove](https://github.com/GradySimon/tensorflow-glove) 코드를 simple하게 옮겨보았습니다.
#
# 논문: https://nlp.stanford.edu/pubs/glove.pdf
# ## Glove 간단히 살펴보기
# 간단한 문장 하나를 학습시켜 보면서 glove를 간단히 살펴보겠습니다.
#
# <code>내가 그의 이름을 불러주었을 때, 그는 내게로 와 꽃이 되었다.</code>
#
# 데이터는 word2vec 처럼 가공을 합니다.
#
# window_size 가 1일 때의 예입니다.
#
# (word, context)의 형태
# ```
# (그의, 내가)
# (그의, 이름을)
# (이름을, 그의)
# (이름을, 불러주었을)
# (내게로, 그는)
# (내게로, 와)
# ...
# ```
# +
# 현재 케라스에는 glove에서 필요한 cooccurrence matrix를 구하며 깔끔하게 sampling 해주는 함수가 없습니다. (ㅠㅅㅠ)
from collections import defaultdict, Counter
txt = "내가 그의 이름을 불러주었을 때, 그는 내게로 와 꽃이 되었다."
datas = txt.split();
# hyperparameter
vocab_size = 1000 # 총 단어 개수
embed_size = 100 # 임베딩 할 사이즈
ws = 3 # window size
min_occurrences = 0 # 제외할 단어 빈도수
# 각 단어의 빈도수 체크
word_counts = Counter()
word_counts.update(datas)
#### co occurrence matrix 및 sample 구하기
co_occurrence_count = defaultdict(float)
data_length = len(datas)
for i, word in enumerate(datas):
# 완전 탐색을 하며 left context, word, right context를 얻어냅니다.
if i < ws:
l_context = datas[:i]
r_context = datas[i+1:i+ws+1]
elif i == data_length - 1:
l_context = datas[i-ws:i]
r_context = []
elif i >= data_length - ws:
l_context = datas[i-ws:i]
r_context = datas[i+1:]
else:
l_context = datas[i-ws:i]
r_context = datas[i+1:i+ws+1]
# co occurrence matrix 를 구합니다.
for i, context_word in enumerate(l_context):
co_occurrence_count[(word, context_word)] += 1 / (i + 1)
for i, context_word in enumerate(r_context):
co_occurrence_count[(word, context_word)] += 1 / (i + 1)
### 구한 count를 토대로 words 사전과 cooccurrence matrix를 만듭니다.
# min occurrences가 넘는 words 만 사용합니다.
words = [word for word, count in word_counts.most_common(vocab_size) if count >= min_occurrences]
word_to_id = {word: i for i, word in enumerate(words)}
co_occurrence_matrix = {
(word_to_id[words[0]], word_to_id[words[1]]): count
for words, count in co_occurrence_count.items()
if words[0] in word_to_id and words[1] in word_to_id}
print(words, '\n')
print(word_to_id, '\n')
print(co_occurrence_matrix, '\n')
# -
# 이제 모델을 짜 봅시다!
#
# 케라스의 [**Embedding layer**](https://keras.io/layers/embeddings/)는 int형 argument들을 vector화 시키는 layer 입니다.
#
# 예를 들어
# ```
# [[4], [20]]
# ```
# 이 입력되었다면 다음과 같이 output을 반환시켜 줄 수 있습니다.
# ```
# [[0.25, 0.1], [0.6, -0.2]]
# ```
#
# 4 라는 수가 \[0.25, 0.1\] 이라는 벡터로 변한 것이죠.
#
# 공식대로
# ``` python
# word_model_out * context_model_out + b1 + b2 == log(co_occurrence_count)
# ```
# 가 되도록 만들어 줍시다
# +
from keras.layers.merge import Dot, Add
from keras.layers.core import Dense, Reshape, Flatten
from keras.layers import Input
from keras.layers.embeddings import Embedding
from keras.models import Sequential, Model
import keras.backend as K
# word model
model_w = Sequential()
model_w.add(Embedding(vocab_size, embed_size, embeddings_initializer='glorot_uniform', input_length=1))
model_w.add(Flatten())
# word bias model
model_wb = Sequential()
model_wb.add(Embedding(vocab_size, 1, embeddings_initializer='glorot_uniform', input_length=1))
model_wb.add(Flatten())
# context model
model_c = Sequential()
model_c.add(Embedding(vocab_size, embed_size, embeddings_initializer='glorot_uniform', input_length=1))
model_c.add(Flatten())
# context bias model
model_cb = Sequential()
model_cb.add(Embedding(vocab_size, 1, embeddings_initializer='glorot_uniform', input_length=1))
model_cb.add(Flatten())
# 전체 모델 작성
input_w = Input(shape=(None,))
input_c = Input(shape=(None,))
output_w = model_w(input_w)
output_wb = model_wb(input_w)
output_c = model_c(input_c)
output_cb = model_cb(input_c)
# word model과 context model의 output들을 내적시킵니다.
x = Dot(axes=1)([output_w, output_c])
# bias들을 더합니다.
x = Add()([x, output_wb, output_cb])
model = Model([input_w, input_c], x)
model.compile(loss='mse', optimizer='adam')
model.summary()
# +
import numpy as np
import math
X1 = []
X2 = []
Y = []
for i in co_occurrence_matrix.items():
X1.append(i[0][0])
X2.append(i[0][1])
Y.append(math.log(i[1]))
X1 = np.array(X1)
X2 = np.array(X2)
Y = np.array(Y)
# -
# 학습시킵니다.
model.fit([X1, X2], Y,
epochs=100,
verbose=1)
# 실제로 사용하는 부분은 model_w의 embedding layer 입니다.
#
# 임베딩하는 방식이 중요한 것이지요.
#
# model_w 에 벡터화 시키고 싶은 수를 넣으면 결과가 나올 것입니다.
print(model_w.predict(np.array([3])))
# 사전 학습된 glove를 사용해 보고 싶으시다면 아래 링크를 참고하시면 좋겠습니다.
#
# https://blog.keras.io/using-pre-trained-word-embeddings-in-a-keras-model.html
# 코드에서 문제가 되는 부분이 있다면 꼭 연락 주세요
# ## Contact me
# 케라스를 사랑하는 개발자 입니다.
#
# 질문, 조언, contribtuion 등 소통은 언제나 환영합니다.
#
# <NAME>(김동현) : <EMAIL>
#
| PR-027_Glove/PR-027_Glove.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exploring Language Features
#
# This is related to processing the AirBnB reviews.
#
# - LangDetect, for detecting language
# - TextBlob, for Sentiment Analysis
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.linear_model import LinearRegression
from sklearn.model_selection import train_test_split
from sklearn.metrics import r2_score, mean_squared_error
import seaborn as sns
import folium
import math
'''
NLP tools
'''
from textblob import TextBlob
from langdetect import detect
# %matplotlib inline
review_df = pd.read_csv('./reviews.csv')
# -
review_df.head(20)
review_df['dtype'] = review_df['comments'].apply(lambda x: type(x))
sns.countplot(review_df['dtype'])
# +
def try_to_detect(txt):
ret = ''
try:
ret = detect(txt)
except:
ret = ''
return ret
review_df['lang'] = review_df['comments'].apply(lambda x: '' if isinstance(x, float) else try_to_detect(x) )
# -
review_df.groupby('lang').count()
# Aha! There are many non-English reviews that need to be disregarded for sentiment analysis.
review_df['sentiment'] = review_df.query('lang == "en"')['comments'].apply(lambda x: TextBlob(x).sentiment.polarity)
review_df['sentiment'].hist(bins=20)
# This shows that almost all reviews are positive, and the majority are moderately positive.
# ## Updates with 2018 Data
# Source: http://insideairbnb.com/get-the-data.html
#
#
review2_df = pd.read_csv('./reviews_2.csv')
review2_df['dtype'] = review2_df['comments'].apply(lambda x: type(x))
sns.countplot(review2_df['dtype'])
review2_df['lang'] = review2_df['comments'].apply(lambda x: '' if isinstance(x, float) else try_to_detect(x) )
review2_df.groupby('lang').count()
review2_df['sentiment'] = review2_df.query('lang == "en"')['comments'].apply(lambda x: TextBlob(x).sentiment.polarity)
review2_df['sentiment'].hist(bins=20)
| .ipynb_checkpoints/Sentiment_Analysis-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Modelado de sistemas
# ===
#
# **<NAME>**
# <EMAIL>
# Universidad Nacional de Colombia, Sede Medellín
# Facultad de Minas
# Medellín, Colombia
#
# ---
#
# Haga click [aquí](https://github.com/jdvelasq/deep-neural-nets/tree/master/) para acceder al repositorio online.
#
# Haga click [aquí](http://nbviewer.jupyter.org/github/jdvelasq/deep-neural-nets/tree/master/) para explorar el repositorio usando `nbviewer`.
# ---
# # Definición del problema
# Se tiene un sistema no lineal con dos entradas y una salida. Se desea construir un modelo que permita pronósticar la salida del sistema a partir de dichas entradas. Un esquema representativo es presentado en la siguiente figura.
#
#
# 
#
#
# # Metodología de Solución
# En el modelado de sistemas se desea obtener un modelo matemático que permita reproducir con cierta precisión la respuesta (variables de salida) de un sistema físico ante la evolución de unas variables de entrada. Ya que en muchos casos no existen modelos matemáticos que permitan describir las relaciones internas entre las variables de entrada y de salida, se suele recurrir a modelos adaptativos como las redes neuronales artificiales. En este caso, el modelo de redes neuronales es entrenado on-line, esto es, la corrección de los pesos sinápticos se realiza para cada patrón presentado, tal que permite la adaptación en el tiempo.
# 
| xx-MulticapaModeladoSis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="4SuL06w0jfZR"
# # Transformer from scratch in PyTorch
# + [markdown] id="PKbsUSTuiPQU"
# Coding the scaled dot-product attention is pretty straightforward — just a few matrix multiplications, plus a softmax function. For added simplicity, we omit the optional Mask operation.
# + id="fdvcDkbKiMA6"
from torch import Tensor
import torch.nn.functional as f
def scaled_dot_product_attention(query: Tensor, key: Tensor, value: Tensor) -> Tensor:
temp = query.bmm(key.transpose(1, 2))
scale = query.size(-1) ** 0.5
softmax = f.softmax(temp / scale, dim=-1)
return softmax.bmm(value)
# + [markdown] id="fiLDxyPJiQty"
# Note that MatMul operations are translated to torch.bmm in PyTorch. That’s because Q, K, and V (query, key, and value arrays) are batches of matrices, each with shape (batch_size, sequence_length, num_features). Batch matrix multiplication is only performed over the last two dimensions.
#
# From the diagram above, we see that multi-head attention is composed of several identical attention heads. Each attention head contains 3 linear layers, followed by scaled dot-product attention. Let’s encapsulate this in an AttentionHead layer:
# + id="5t097JfqiQrc"
import torch
from torch import nn
class AttentionHead(nn.Module):
def __init__(self, dim_in: int, dim_k: int, dim_v: int):
super().__init__()
self.q = nn.Linear(dim_in, dim_k)
self.k = nn.Linear(dim_in, dim_k)
self.v = nn.Linear(dim_in, dim_v)
def forward(self, query: Tensor, key: Tensor, value: Tensor) -> Tensor:
return scaled_dot_product_attention(self.q(query), self.k(key), self.v(value))
# + [markdown] id="e4V_iqTSiQno"
# Now, it’s very easy to build the multi-head attention layer. Just combine num_heads different attention heads and a Linear layer for the output.
# + id="2L_SNeYFiQlz"
class MultiHeadAttention(nn.Module):
def __init__(self, num_heads: int, dim_in: int, dim_k: int, dim_v: int):
super().__init__()
self.heads = nn.ModuleList(
[AttentionHead(dim_in, dim_k, dim_v) for _ in range(num_heads)]
)
self.linear = nn.Linear(num_heads * dim_v, dim_in)
def forward(self, query: Tensor, key: Tensor, value: Tensor) -> Tensor:
return self.linear(
torch.cat([h(query, key, value) for h in self.heads], dim=-1)
)
# + [markdown] id="KBaOoNjMiQjm"
# We need one more component before building the complete transformer: positional encoding. Notice that MultiHeadAttention has no trainable components that operate over the sequence dimension (axis 1). Everything operates over the feature dimension (axis 2), and so it is independent of sequence length. We have to provide positional information to the model, so that it knows about the relative position of data points in the input sequences.
# + id="vH2OGPkniQhH"
def position_encoding(
seq_len: int, dim_model: int, device: torch.device = torch.device("cpu"),
) -> Tensor:
pos = torch.arange(seq_len, dtype=torch.float, device=device).reshape(1, -1, 1)
dim = torch.arange(dim_model, dtype=torch.float, device=device).reshape(1, 1, -1)
phase = pos / 1e4 ** (dim // dim_model)
return torch.where(dim.long() % 2 == 0, torch.sin(phase), torch.cos(phase))
# + [markdown] id="JnmZrPpFid3w"
# We can code the encoder/decoder modules independently of one another, and then combine them at the end. But first we need a few more pieces of information, which aren’t included in the figure above. For example, how should we choose to build the feed forward networks?
# + id="RXG7YhX8ihQl"
def feed_forward(dim_input: int = 512, dim_feedforward: int = 2048) -> nn.Module:
return nn.Sequential(
nn.Linear(dim_input, dim_feedforward),
nn.ReLU(),
nn.Linear(dim_feedforward, dim_input),
)
# + [markdown] id="q-NWu_5uiuSn"
# The output of each sub-layer is LayerNorm(x + Sublayer(x)), where Sublayer(x) is the function implemented by the sub-layer itself. … We apply dropout to the output of each sub-layer, before it is added to the sub-layer input and normalized.
# + id="B-IOnfeyihNA"
class Residual(nn.Module):
def __init__(self, sublayer: nn.Module, dimension: int, dropout: float = 0.1):
super().__init__()
self.sublayer = sublayer
self.norm = nn.LayerNorm(dimension)
self.dropout = nn.Dropout(dropout)
def forward(self, *tensors: Tensor) -> Tensor:
# Assume that the "value" tensor is given last, so we can compute the
# residual. This matches the signature of 'MultiHeadAttention'.
return self.norm(tensors[-1] + self.dropout(self.sublayer(*tensors)))
# + [markdown] id="W5D8RZrqihK2"
# Time to dive in and create the encoder. Using the utility methods we just built, this is pretty easy.
# + id="tbCpBmFcihIk"
class TransformerEncoderLayer(nn.Module):
def __init__(
self,
dim_model: int = 512,
num_heads: int = 6,
dim_feedforward: int = 2048,
dropout: float = 0.1,
):
super().__init__()
dim_k = dim_v = dim_model // num_heads
self.attention = Residual(
MultiHeadAttention(num_heads, dim_model, dim_k, dim_v),
dimension=dim_model,
dropout=dropout,
)
self.feed_forward = Residual(
feed_forward(dim_model, dim_feedforward),
dimension=dim_model,
dropout=dropout,
)
def forward(self, src: Tensor) -> Tensor:
src = self.attention(src, src, src)
return self.feed_forward(src)
class TransformerEncoder(nn.Module):
def __init__(
self,
num_layers: int = 6,
dim_model: int = 512,
num_heads: int = 8,
dim_feedforward: int = 2048,
dropout: float = 0.1,
):
super().__init__()
self.layers = nn.ModuleList([
TransformerEncoderLayer(dim_model, num_heads, dim_feedforward, dropout)
for _ in range(num_layers)
])
def forward(self, src: Tensor) -> Tensor:
seq_len, dimension = src.size(1), src.size(2)
src += position_encoding(seq_len, dimension)
for layer in self.layers:
src = layer(src)
return src
# + [markdown] id="GZrsGydnihGL"
# The decoder module is extremely similar. Just a few small differences:
# - The decoder accepts two arguments (target and memory), rather than one.
# - There are two multi-head attention modules per layer, instead of one.
# - The second multi-head attention accepts memory for two of its inputs.
# + id="dxezkFJzihEL"
class TransformerDecoderLayer(nn.Module):
def __init__(
self,
dim_model: int = 512,
num_heads: int = 6,
dim_feedforward: int = 2048,
dropout: float = 0.1,
):
super().__init__()
dim_k = dim_v = dim_model // num_heads
self.attention_1 = Residual(
MultiHeadAttention(num_heads, dim_model, dim_k, dim_v),
dimension=dim_model,
dropout=dropout,
)
self.attention_2 = Residual(
MultiHeadAttention(num_heads, dim_model, dim_k, dim_v),
dimension=dim_model,
dropout=dropout,
)
self.feed_forward = Residual(
feed_forward(dim_model, dim_feedforward),
dimension=dim_model,
dropout=dropout,
)
def forward(self, tgt: Tensor, memory: Tensor) -> Tensor:
tgt = self.attention_1(tgt, tgt, tgt)
tgt = self.attention_2(memory, memory, tgt)
return self.feed_forward(tgt)
class TransformerDecoder(nn.Module):
def __init__(
self,
num_layers: int = 6,
dim_model: int = 512,
num_heads: int = 8,
dim_feedforward: int = 2048,
dropout: float = 0.1,
):
super().__init__()
self.layers = nn.ModuleList([
TransformerDecoderLayer(dim_model, num_heads, dim_feedforward, dropout)
for _ in range(num_layers)
])
self.linear = nn.Linear(dim_model, dim_model)
def forward(self, tgt: Tensor, memory: Tensor) -> Tensor:
seq_len, dimension = tgt.size(1), tgt.size(2)
tgt += position_encoding(seq_len, dimension)
for layer in self.layers:
tgt = layer(tgt, memory)
return torch.softmax(self.linear(tgt), dim=-1)
# + [markdown] id="qISz9Njni6Y_"
# Lastly, we need to wrap everything up into a single Transformer class. This requires minimal work, because it’s nothing new — just throw an encoder and decoder together, and pass data through them in the correct order.
# + id="mtdMH0rbi66X"
class Transformer(nn.Module):
def __init__(
self,
num_encoder_layers: int = 6,
num_decoder_layers: int = 6,
dim_model: int = 512,
num_heads: int = 6,
dim_feedforward: int = 2048,
dropout: float = 0.1,
activation: nn.Module = nn.ReLU(),
):
super().__init__()
self.encoder = TransformerEncoder(
num_layers=num_encoder_layers,
dim_model=dim_model,
num_heads=num_heads,
dim_feedforward=dim_feedforward,
dropout=dropout,
)
self.decoder = TransformerDecoder(
num_layers=num_decoder_layers,
dim_model=dim_model,
num_heads=num_heads,
dim_feedforward=dim_feedforward,
dropout=dropout,
)
def forward(self, src: Tensor, tgt: Tensor) -> Tensor:
return self.decoder(tgt, self.encoder(src))
# + [markdown] id="F_VtLkoci8zB"
# And we’re done! Let’s create a simple test, as a sanity check for our implementation. We can construct random tensors for src and tgt, check that our model executes without errors, and confirm that the output tensor has the correct shape.
# + colab={"base_uri": "https://localhost:8080/"} id="np60EZkci-JP" executionInfo={"status": "ok", "timestamp": 1634026193801, "user_tz": -330, "elapsed": 2353, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}} outputId="e53e7f16-7e37-4d8f-cdcd-96654401456c"
src = torch.rand(64, 16, 512)
tgt = torch.rand(64, 16, 512)
out = Transformer()(src, tgt)
print(out.shape)
# torch.Size([64, 16, 512])
| docs/T904848_Transformer_from_scratch_in_PyTorch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Capture Images from Webcam
# This script captures video from webcam, detects face area and shows it to user in a window. Video (of face area detected) is shown to user until user presses `ESC (or T/F/Q)` key. When an `ESC (or T/F/Q)` is pressed then video is exited and an image of current state is saved in current working directory. This is to show that when a user will answer the question by pressing some key/button then as soon as a button/key is pressed, user's image will be saved.
# +
import cv2
class VideoCamera:
def __init__(self):
#passing 0 to VideoCapture means fetch video from webcam
self.video_capture = cv2.VideoCapture(0)
#release resources like webcam
def __del__(self):
self.video_capture.release()
def read_image(self):
#get a single frame of video
ret, frame = self.video_capture.read()
#return the frame to user
return ret, frame
#method to release webcam manually
def release(self):
self.video_capture.release()
# +
def detect_face(img):
#load OpenCV face detector, I am using LBP which is fast
#there is also a more accurate but slow Haar classifier
face_cascade = cv2.CascadeClassifier('data/lbpcascade_frontalface.xml')
# img_copy = np.copy(colored_img)
#convert the test image to gray image as opencv face detector expects gray images
gray = cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)
#let's detect multiscale (some images may be closer to camera than others) images
#result is a list of faces
faces = face_cascade.detectMultiScale(gray, scaleFactor=1.1, minNeighbors=5);
#if no faces are detected then return original img
if (len(faces) == 0):
return img
#under the assumption that there will be only one face,
#extract the face area
(x, y, w, h) = faces[0]
#return only the face part of the image
return img[y:y+w, x:x+h]
# +
import matplotlib.pyplot as plt
# %matplotlib inline
#opencv loads an image into BGR color space,
#I use this method to convert it to RGB
def convert2RGB(img):
return cv2.cvtColor(img, cv2.COLOR_BGR2RGB)
#load a test image and display it
img = cv2.imread('data/test2.jpg')
plt.imshow(convert2RGB(img))
# -
#detect the face and get the face area
face = detect_face(img)
#display detected face
plt.imshow(cv2.cvtColor(face, cv2.COLOR_BGR2RGB))
# +
#create a Video camera instance
camera = VideoCamera()
cv2.namedWindow("Webcam feed", 0)
while (True):
#read a video frame
ret, frame = camera.read_image()
if(ret == False):
print('Video capture failed')
break
face = detect_face(frame)
#display the image to user
cv2.imshow("Webcam feed", face)
#wait for 100ms for user to press a key
key = cv2.waitKey(100)
#if user pressed an ESC (or T/F/Q) key then exit and save the img
if(key in [27, ord('T'), ord('F'), ord('t'), ord('f'), ord('q'), ord('Q')]):
cv2.imwrite('img.jpg', face)
break;
# release resources
camera.release()
cv2.waitKey(100)
cv2.destroyAllWindows()
cv2.waitKey(1)
cv2.destroyAllWindows()
# -
camera.release()
| Webcam-images-capture.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/TLeahSpencer/Loops_Lists_Wkshps/blob/master/9_1_20AdvLoops%26Lists.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="dQGprOi2OMjD" colab_type="text"
# ##Office Hours workshop with Anade (<NAME>)
# #Advanced Loops & Lists in Python
# + [markdown] id="gxqm_cSaDRm3" colab_type="text"
# Interger = Number
#
# Float = Numbers with Decimals
#
# String = Sequence of Characters (Words, Letters, Numbers, intergers, floats, etc)
#
# Object = All Characters (Strings and Objects have become synonymous)
#
# ##**Loops** = Perform a task over and over again
#
# ##**Range Function** = Ordered Sequence
#
# ##**List** = Ordered Sequence
#
# ##**Set** = Unordered Sequence
# + [markdown] id="bq-Vdt_hElUG" colab_type="text"
# **Conditional Statements**
#
# Equals: x==y
#
# Not Equals: x!=y
#
# Less than: x<y
#
# Less than or equal to: x<=y
#
# Greater than: x>y
#
# Greater than or equal to: x>=y
# + id="to1MxgTNE_T-" colab_type="code" colab={}
#define a list, Family is the name of the list here
Family = ['Husband', 'Sister', 'Child', 'Brother']
# + id="p5Yg0RR25rFy" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 84} outputId="e14ef1f8-8f9e-42ac-a375-fcd062b11275"
#function that acts on each item in the list
#in this case, it prints them
#A is a made-up thing, can be anything you make up
for A in Family: #this creates the Value A (blank variable) for values in Family (a real variable, a list)
print(A) #this tells to print the variable A, which is defined by the values in Family
# + id="H-j5Avh4OLNm" colab_type="code" colab={}
#define a new variable Math, a list
Math = [7,8,9,5,4,2,1]
# + id="YVqMaYvrFWMQ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 134} outputId="436e3956-6546-4151-aad4-8c19b472be35"
#this tells the variable to be multiplied by itself
for c in Math:
print(c*c)
# + id="wxn0_w1CFaQQ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 134} outputId="fda600eb-52fa-4caf-9bb4-e368133859e1"
#this tells the variable to be added to 12
for c in Math:
print(c+12)
# + id="dLiZWUCLF9Y2" colab_type="code" colab={}
#define a new variable with range function
#3 is position where range starts (so 4th because 1st is 0)
#200 is position where range ends (so 199 would be the highest number in range)
#21 is how many numbers to "skip" by in creating the range
Math2 = range(3,200,21)
# + id="ftWoYG6hvjXl" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 33} outputId="79cdd702-4096-4183-ded4-bcdc2a1b453b"
#this shows what Math2 is defined as
Math2
# + id="Z4seainavtsd" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 33} outputId="edeb730f-a06d-4774-8104-98d433261f35"
#this tells what type of object Math2 is
type(Math2)
# + id="quhdVlcowWWS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 33} outputId="d79d39c2-a70c-4672-d7f7-2422370a8be1"
#we can't get the values the range defines without the for statement below
print(Math2)
# + id="sKxFYPbXGKqn" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 184} outputId="f4930dc5-35d7-4a8f-e7b3-09a9664f098a"
#here is how we can pull the values we created in the range
for c in Math2:
print(c)
# + id="1va_St-GGNym" colab_type="code" colab={}
#define another range variable
Math3 = range(2, 50, 3)
# + id="PezDDAiBxc4r" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 284} outputId="23dbe644-ae06-4d90-b222-b3b9c4fa8733"
#see the values in our new range variable
for c in Math3:
print(c)
# + id="NFcsGdF2GZhq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 284} outputId="19532d48-d1c2-4839-f7d4-3c8d3af203eb"
#now do some math on the values in the variable
for c in Math3:
print(c*c+2) #2*2=4+2=6, 5*5=25+2=27, ets
# + id="iAbry7NFGd2D" colab_type="code" colab={}
#define another list as variable Math4
Math4 = [5,6,7,7,5,7,8,12,14]
# + id="YRLdQW_DGtdS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 33} outputId="0e365cc7-d8be-4bd0-8a8d-c0653281bedf"
#find out the length or amount of characters or items in our list
len(Math4)
# + id="4M0tRlEkGvb_" colab_type="code" colab={}
#define a variable based on the length of the other variable
NewMath=len(Math4)
# + id="WNULFtbAGzNR" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 33} outputId="ab261b10-4bdf-4afe-d6f5-5cb88b0b3513"
#view the value of the new variable
#notice the for function is not needed
NewMath
# + id="xGx3wGo4G0t1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 167} outputId="4627e0fb-4460-41f2-dfd7-8a7eb778bc02"
#create a range based on the new variable
for x in range(0,NewMath):
print(x)
# + id="Yi1GryixHK8B" colab_type="code" colab={}
#create a new variable which is a list
Family2 = ['Husband', 'Sister', 'Child', 'Brother', 'Mother', 'Father']
# + id="MPAzCiBWHW0K" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 117} outputId="acca16b1-843e-4eb5-9f50-191b98acdf26"
#view all the values in the variable
for b in Family2:
print(b)
# + id="tmSOY8qSHZn5" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 100} outputId="07da258e-ee83-462b-e79f-f706b0f6b007"
#break the loop after the == Conditional Statement, by placing print() function before it
for b in Family2:
print(b)
if b == 'Mother':
break
# + id="vwudH-xmHp7R" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 84} outputId="a923e613-3f7a-4650-ca02-49d8652a7736"
#break the loop before the == Conditional Statement, by placing the print() function after it
for b in Family2:
if b == 'Mother':
break
print(b)
# + id="2JFhPLhHH_Mk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 100} outputId="81ce2877-9d3d-4e56-d097-d646788ea9ec"
#this is how to ignore a value in the loop
#poor mama :( she always gets ignored
for b in Family2:
if b != 'Mother':
print(b)
# + [markdown] id="tWl3Gr8vzaMr" colab_type="text"
# ###If != is present, that means the loop remains True, green light to keep going, just don't include this output
# + id="vKh3-tlEIbQB" colab_type="code" colab={}
#define another variable
A = 16
# + id="IaZ-tzaQInms" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 217} outputId="679be7f2-f283-47da-df36-aa7685561990"
#a "While Loop" starts from the start of the variable
#and ends until our while is over, until A is no longer less than 40, in this case
while A < 40:
print(A)
A += 2
# + id="F4qCGU6SIul7" colab_type="code" colab={}
#define another variable
A2 = 60
# + id="sxVthxgxJJ-L" colab_type="code" colab={}
#WARNING: this code will keep running and running and running
#because we are adding, and the conditional statement of being greater than 40 will always be True
#to stop the code, right-click and select 'Interrupt Execution'
#no heads need to roll, at least not tonight
while A2 > 40:
print(A2)
A2 += 2
# + [markdown] id="E9wVFnnh01ah" colab_type="text"
# #Some final definitions from Anade
# ##While Loop: repeated as long as an expression is true
# ##Expression: a statement that has a value
# ##Do While Loop or Repeat Until Loop: repeats until an expression becomes False
# ##Infinite or Endless Loop: repeats indefinitely because it has no terminating condition, the exit condition is never met, or the loop is instructed to start over from the beginning
# ###Although it is possible for a programmer to intentionally use an infinite loop, they are often mistakes made by new programmers.
# ##Nested Loop: appears inside any other For, While, or Do While Loop
# + id="TI3NoJPhJRje" colab_type="code" colab={}
| 9_1_20AdvLoops&Lists.ipynb |
# +
# Copyright 2010 <NAME> <EMAIL>
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
"""
Kakuru puzzle in Google CP Solver.
http://en.wikipedia.org/wiki/Kakuro
'''
The object of the puzzle is to insert a digit from 1 to 9 inclusive
into each white cell such that the sum of the numbers in each entry
matches the clue associated with it and that no digit is duplicated in
any entry. It is that lack of duplication that makes creating Kakuro
puzzles with unique solutions possible, and which means solving a Kakuro
puzzle involves investigating combinations more, compared to Sudoku in
which the focus is on permutations. There is an unwritten rule for
making Kakuro puzzles that each clue must have at least two numbers
that add up to it. This is because including one number is mathematically
trivial when solving Kakuro puzzles; one can simply disregard the
number entirely and subtract it from the clue it indicates.
'''
This model solves the problem at the Wikipedia page.
For a larger picture, see
http://en.wikipedia.org/wiki/File:Kakuro_black_box.svg
The solution:
9 7 0 0 8 7 9
8 9 0 8 9 5 7
6 8 5 9 7 0 0
0 6 1 0 2 6 0
0 0 4 6 1 3 2
8 9 3 1 0 1 4
3 1 2 0 0 2 1
Compare with the following models:
* Comet : http://www.hakank.org/comet/kakuro.co
* MiniZinc: http://www.hakank.org/minizinc/kakuro.mzn
* SICStus : http://www.hakank.org/sicstus/kakuro.pl
* ECLiPSe: http://www.hakank.org/eclipse/kakuro.ecl
* Gecode: http://www.hakank.org/gecode/kenken2.cpp
This model was created by <NAME> (<EMAIL>)
Also see my other Google CP Solver models:
http://www.hakank.org/google_or_tools/
"""
from __future__ import print_function
import sys
from ortools.constraint_solver import pywrapcp
#
# Ensure that the sum of the segments
# in cc == res
#
def calc(cc, x, res):
solver = list(x.values())[0].solver()
# ensure that the values are positive
for i in cc:
solver.Add(x[i[0] - 1, i[1] - 1] >= 1)
# sum the numbers
solver.Add(solver.Sum([x[i[0] - 1, i[1] - 1] for i in cc]) == res)
# Create the solver.
solver = pywrapcp.Solver("Kakuro")
#
# data
#
# size of matrix
n = 7
# segments
# [sum, [segments]]
# Note: 1-based
problem = [[16, [1, 1], [1, 2]], [24, [1, 5], [1, 6], [1, 7]],
[17, [2, 1], [2, 2]], [29, [2, 4], [2, 5], [2, 6], [2, 7]],
[35, [3, 1], [3, 2], [3, 3], [3, 4], [3, 5]], [7, [4, 2], [4, 3]],
[8, [4, 5], [4, 6]], [16, [5, 3], [5, 4], [5, 5], [5, 6], [5, 7]],
[21, [6, 1], [6, 2], [6, 3], [6, 4]], [5, [6, 6], [6, 7]],
[6, [7, 1], [7, 2], [7, 3]], [3, [7, 6], [7, 7]],
[23, [1, 1], [2, 1], [3, 1]], [30, [1, 2], [2, 2], [3, 2], [4, 2]],
[27, [1, 5], [2, 5], [3, 5], [4, 5], [5, 5]], [12, [1, 6], [2, 6]],
[16, [1, 7], [2, 7]], [17, [2, 4], [3, 4]],
[15, [3, 3], [4, 3], [5, 3], [6, 3], [7, 3]],
[12, [4, 6], [5, 6], [6, 6], [7, 6]], [7, [5, 4], [6, 4]],
[7, [5, 7], [6, 7], [7, 7]], [11, [6, 1], [7, 1]],
[10, [6, 2], [7, 2]]]
num_p = len(problem)
# The blanks
# Note: 1-based
blanks = [[1, 3], [1, 4], [2, 3], [3, 6], [3, 7], [4, 1], [4, 4], [4, 7],
[5, 1], [5, 2], [6, 5], [7, 4], [7, 5]]
num_blanks = len(blanks)
#
# variables
#
# the set
x = {}
for i in range(n):
for j in range(n):
x[i, j] = solver.IntVar(0, 9, "x[%i,%i]" % (i, j))
x_flat = [x[i, j] for i in range(n) for j in range(n)]
#
# constraints
#
# fill the blanks with 0
for i in range(num_blanks):
solver.Add(x[blanks[i][0] - 1, blanks[i][1] - 1] == 0)
for i in range(num_p):
segment = problem[i][1::]
res = problem[i][0]
# sum this segment
calc(segment, x, res)
# all numbers in this segment must be distinct
segment = [x[p[0] - 1, p[1] - 1] for p in segment]
solver.Add(solver.AllDifferent(segment))
#
# search and solution
#
db = solver.Phase(x_flat, solver.INT_VAR_DEFAULT, solver.INT_VALUE_DEFAULT)
solver.NewSearch(db)
num_solutions = 0
while solver.NextSolution():
for i in range(n):
for j in range(n):
val = x[i, j].Value()
if val > 0:
print(val, end=" ")
else:
print(" ", end=" ")
print()
print()
num_solutions += 1
solver.EndSearch()
print()
print("num_solutions:", num_solutions)
print("failures:", solver.Failures())
print("branches:", solver.Branches())
print("WallTime:", solver.WallTime())
| examples/notebook/contrib/kakuro.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
os.environ['CUDA_VISIBLE_DEVICES'] = '0'
# -
import tensorflow as tf
# Simple demo neural network classifier:
# +
from tensorflow.python.keras.models import Sequential
from tensorflow.python.keras.layers import Dense
num_classes = 2
def build_classifier(num_classes, num_dimensions=1781):
classifier = Sequential([
Dense(1024, activation='relu', input_shape=(num_dimensions, )),
Dense(128, activation='relu'),
Dense(16, activation='relu'),
Dense(num_classes, activation='softmax')
])
classifier.compile(optimizer='adam',
loss='categorical_crossentropy',
metrics=['accuracy',])
return classifier
# +
import numpy as np
with open("counts/SRR_47normal_small.list") as normal_subjects_file:
healthy_subjects = [line.strip() for line in normal_subjects_file]
with open("counts/SRR_50aml_small.list") as diseased_subjects_file:
diseased_subjects = [line.strip() for line in diseased_subjects_file]
# -
print(len(healthy_subjects), len(diseased_subjects))
# +
def load_data(subjects_array):
a = []
for person in subjects_array:
try:
with open("counts/" + person + ".count") as person_file:
vector = [line.strip().split()[-1] for line in person_file]
a.append(np.array(vector))
except FileNotFoundError:
continue
return a
X_healthy = load_data(healthy_subjects)
y_healthy = [0. for _ in range(len(X_healthy))]
X_diseased = load_data(diseased_subjects)
y_diseased = [1. for _ in range(len(X_diseased))]
# -
print(len(X_healthy), len(X_diseased))
print(len(y_healthy), len(y_diseased))
X = np.array(X_healthy + X_diseased)
y = np.array(y_healthy + y_diseased)
num_people = len(X)
# One-Hot Encode Output Labels:
# +
from tensorflow.python.keras.utils import to_categorical
y1hot = to_categorical(y, num_classes=num_classes)
# -
# ## K-Fold Cross Validation
# Since we don't have many people represented in our data, we'll do leave-one-out cross validation:
# +
from sklearn.model_selection import KFold
kf = KFold(n_splits=num_people)
fold_number = 1
cvscores = []
for train_index, val_index in kf.split(X):
print("Fold " + str(fold_number))
X_train, X_val = X[train_index], X[val_index]
y_train, y_val = y1hot[train_index], y1hot[val_index]
classifier = build_classifier(num_classes=num_classes)
classifier.fit(X_train, y_train, batch_size=num_people, epochs=8, validation_data=(X_val, y_val), verbose=2)
scores = classifier.evaluate(X_val, y_val, verbose=0)
print(classifier.metrics_names)
print(scores)
cvscores.append(scores)
classifier.save_weights("fold{}.weights.hdf5".format(fold_number))
del classifier
fold_number += 1
# -
accuracies = np.array(cvscores)[:, 1]
print("%.2f (+/- %.2f)" % (np.mean(accuracies), np.std(accuracies)))
# ## Predict phenotype on test data using ensemble average of K models
# Generate dummy test set:
test_set = [(np.zeros(1700), 0) if np.random.choice(2) == 0 else (np.ones(1700), 1) for _ in range(10)]
X_test, y_test = zip(*test_set)
y_test1hot = to_categorical(y_test, num_classes=num_classes)
predictions = []
for fold in range(50):
model = build_classifier(num_classes=num_classes)
model.load_weights("fold{}.weights.hdf5".format(fold + 1))
predictions.append(model.predict(np.array(X_test)))
avg_predictions = np.mean(predictions, axis=0)
y_pred = np.argmax(avg_predictions, axis=1)
# +
# %matplotlib inline
# http://scikit-learn.org/stable/auto_examples/model_selection/plot_confusion_matrix.html
import itertools
import matplotlib.pyplot as plt
from sklearn.metrics import confusion_matrix
def plot_confusion_matrix(cm, classes,
normalize=False,
title='Confusion matrix',
cmap=plt.cm.Blues):
"""
This function prints and plots the confusion matrix.
Normalization can be applied by setting `normalize=True`.
"""
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
print("Normalized confusion matrix")
else:
print('Confusion matrix, without normalization')
print(cm)
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45)
plt.yticks(tick_marks, classes)
fmt = '.2f' if normalize else 'd'
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, format(cm[i, j], fmt),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
cnf_matrix = confusion_matrix(y_test, y_pred)
plt.figure()
plot_confusion_matrix(cnf_matrix, classes=["healthy", "diseased"],
title='Confusion matrix, without normalization')
| training.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Chapter 16: Data formats I (CSV and TSV)
#
# You probably have heard of (or are already quite familiar with) different data formats, such as plain text, tables (CSV/TSV), XML, JSON and RDF. These formats are simply the result of agreements that were made between people on how to organize and store data. Some of these formats, such as XML and RDF, have a high degree of structure, whereas plain text is a typical example of unstructured data. Structuring data according to predefined specifications allows information in the data to be easily ordered and processed by machines. You can compare highly structured data with a perfectly organized filing cabinet where everything is identified, labeled and easy to access.
#
# **In general, you can treat any of these data formats as a text file with certain regularities people have agreed on**. You can simply read in the file and use the tools you already to exploit these regularities and access information. For some dataformats with rather complex structures (such as xml), it is more convenient to use existing python packages to extract infromation.
#
# This notebook introduces tabular formats: CSV/TSV. TSV/CSV is a highly transparent way of structuring data in tables. It is a rather straight-forward structure and does not necessarily require specific python packages.
#
# **At the end of this chapter, you will be able to:**
# * read CSV/TSV data
# * manipulate CSV/TSV data
# * write CSV/TSV data
#
# **If you want to learn more about these topics, you might find the following links useful:**
# * [Tutorial: Reading and Manipulating CSV Files](https://newcircle.com/s/post/1572/python_for_beginners_reading_and_manipulating_csv_files)
#
# If you have **questions** about this chapter, please contact **Pia** (<EMAIL>).
# ## 1. Introduction to CSV and TSV (tables)
# The **table** is probably one of the most common and intuitive data formats. Undoubtedly, you have already worked with tabular data in Excel, Numbers or Google Sheets. A table represents a set of data points as a series of rows, with a column for each of the data points' properties. In other words: a table has vertical **columns** (often identifiable by name) and horizontal **rows**, and the **cells** are the unit where a row and column intersect. This is a very simple example:
#
# | name | house | patronus |
# |----------|------------|----------|
# | Harry | Gryffindor | stag |
# | Hermione | Gryffindor | otter |
# | Draco | Slytherin | unknown |
# | Hannah | Hufflepuff | unknown |
#
# Tabular data can be encoded as **CSV (comma-separated values) or TSV (tab-separated values)**. CSV and TSV files are simply plain text files in which each line represents a row and the columns are separated by a comma (for CSV) or a tab character (for TSV).
#
#
#
#
# You can find examples of CSV files in the folder `../Data/csv_data`. For instance, look at the files in `../Data/csv_data/baby_names/names_by_state/`. If you like, open them in a text editor (e.g. [Atom](https://atom.io/), [BBEdit](https://www.barebones.com/products/bbedit/download.html) or [Notepad++](https://notepad-plus-plus.org)) or Excel (convert text to columns by using the comma as delimiter) to see their content.
#
# For example, these are the first 10 rows of the CSV file `AK.csv`:
#
#
# 
#
# A TSV file would look like this (note that the tab separator is represented as '\t', which is not shown here):
#
#
# 
#
#
# The file `AK.csv` contains a list of names given to children in the state Alaska from 1910 to 2015 with their frequency. Each line in this file has five elements, which are separated by commas:
#
# - the state abbreviation (AK for Alaska)
# - gender (F/M)
# - year
# - name
# - frequency of that name in the given year and state
#
# Below, we will work towards representing this data in Python as a **list of lists** (i.e. nested list), or as a **list of dicts**. In both cases, the elements of the (first) list represent the complete rows. The individual rows, then, can be either represented as a list (without column names) or as a dictionary (with column names).
#
# | LIST OF LISTS | LIST OF DICTS |
# |:-------------------------:|:-------------------------:|
#  | 
# ## 2. Reading CSV files
#
# Because CSV/TSV files are essentially text files, we can open and read them in the same way as we have seen before:
# Read the file and print its content
filename = "../Data/csv_data/baby_names/names_by_state/AK.csv"
with open(filename, "r") as csvfile:
content = csvfile.read()
print(content)
# Please also have a close look at the internal representation of the file. Do you see how the columns are separated by commas, and the rows by newline characters `\n`?
print(repr(content))
# ### 2.1 Reading rows as lists
# Now, let's see how we can get to the 'list of lists' representation. We can do that by iterating over each line of this file (as we have seen before), and then split each row into columns using the `split()` method:
# +
# Read the file and get all lines
# create empty list to collect the rows:
csv_data = []
filename = "../Data/csv_data/baby_names/names_by_state/AK.csv"
with open(filename, "r") as csvfile:
for row in csvfile:
row = row.strip("\n") # remove all newlines
columns = row.split(",") # split the line into columns
csv_data.append(columns)
# Print only first 10 rows
print(csv_data[:10])
print()
# Iterate over first 10 rows
for row in csv_data[:10]:
print(row)
# -
# It worked! The variable `csv_data` now contains a list of all rows in the file. Now we can easily work with the data by using the indices of the lists to access cells. Have a look at the following examples:
# Example: print all information of the first 10 rows
for row in csv_data[:10]:
state = row[0]
gender = row[1]
year = row[2]
name = row[3]
frequency = row[4]
print(state, gender, year, name, frequency)
# Example: print all names given in 1912
for row in csv_data:
year = row[2]
name = row[3]
# note that the year is represented as a string (not an int)
if year == "1912":
print(name)
# ### 2.2 Reading rows as dicts
#
# We can also create a 'list of dicts'. We do this by first creating a dictionary for each row, and appending that dictionary to the list of rows. We can simply use the colum headers as keys, so we can easily access the corresponding values later on.
#
# Note: Do you see that we also have already converted the numerical values to `int`?
# +
# Read the file and get all lines
# again, create a list to collect the rows:
csv_data = []
filename = "../Data/csv_data/baby_names/names_by_state/AK.csv"
with open(filename, "r") as csvfile:
for row in csvfile:
row = row.strip("\n") # remove all newlines
columns = row.split(",") # split the line into columns
# Create a dictionary and add to list
dict_row = {"state": columns[0],
"gender": columns[1],
"year": int(columns[2]),
"name": columns[3],
"frequency": int(columns[4])}
csv_data.append(dict_row)
# First 10 rows
print(csv_data[:10])
print()
# Iterate over first 10 rows
for row in csv_data[:10]:
print(row)
# -
# Again, we can now easily work with the data, but now we use the *names* of the columns instead of indices to access the cells. This can make the code more readable. Have a look at the following examples:
# Example: print all information of the first 5 rows
for row in csv_data[:5]:
for column_name, column_value in row.items():
print(column_name, "=", column_value)
print()
# Example: print all names given in 1912
for row in csv_data:
if row["year"] == 1912:
print(row["name"])
# It does not really matter whether you choose for a 'list of lists' or a 'list of dicts'. Just use the one that you prefer.
# ## 3. Writing CSV files
#
# Let's say now we have a table in Python stored as a 'list of lists' or as a 'list of dicts' and we want to store our result in a CSV file. This is basically the inverse process of reading a CSV file.
# ### 3.1 Writing rows as lists
#
# In order to write a list of lists as a CSV file, we need to iterate over the rows and make a string out of them. Remember that we can concatenate strings in a list with any separator with the `join()` method:
a_list = ["John", "<EMAIL>", "555-1234"]
a_string = ",".join(a_list)
print(a_string)
# In addition, we should use the newline character `\n` to write each row on a line.
# +
# Create list of lists
address_book = [
["John", "<EMAIL>", "555-1234"],
["William", "<EMAIL>", "555-5678"],
["Jane", "<EMAIL>", "555-7777"]
]
# Write the list of lists to a CSV file
outfilename = "../Data/address_book.csv"
with open(outfilename, "w") as outfile:
for row in address_book:
line = ",".join(row) + '\n'
outfile.write(line)
# -
# ### 3.2 Writing rows as dicts
#
# In order to write a list of dicts to a CSV file, we need to first get all the values in each dictionary. The rest works exactly the same. In the following code, we use the tab separator `\t` and save it with the `.tsv` extension.
# +
# Create list of dicts
address_book = [
{"name":"John", "e-mail":"<EMAIL>", "phone":"555-1234"},
{"name":"William", "e-mail":"<EMAIL>", "phone":"555-5678"},
{"name":"Jane", "e-mail":"<EMAIL>", "phone":"555-7777"}
]
# Write the list of dicts to a TSV file
outfilename = "../Data/csv_data/address_book.tsv"
with open(outfilename, "w") as outfile:
# Write the rows using the values of the dictionaries
for row in address_book:
column_values = row.values()
line = "\t".join(column_values) + '\n'
outfile.write(line)
# -
# ## 4. Dealing with column headers
#
# Many csv/tsv files contain headers. This means that the first row contains the names of the columns. We easily read and write these kinds of files without having to type the headers manually.
#
#
# ### 4.1 Reading csv/tsv files with a header row
# Consider the file called Concreteness_ratings_Brysbaert_et_al_BRM.txt (we're reading it in and printing the first 5 lines below).
#
# The file has 10 columns, each of which have a column **header**. This means that all the values in the first row are not actual values. We need to treat them differently if we want to analyze the data (i.e. everything from the second row on). In addition, it would be convenient to use this information right away, without having to first inspect the file and manually type the headers as keys in dictionaries!
# +
with open('../Data/csv_data/Concreteness_ratings_Brysbaert_et_al_BRM.tsv') as infile:
lines = infile.read().split('\n')
for line in lines[:5]:
print(line)
# -
# Can you complete the example below?
# +
data_dict_list = []
with open('../Data/csv_data/Concreteness_ratings_Brysbaert_et_al_BRM.tsv') as infile:
lines = infile.read().split('\n')
# the first row is at index 0 and contains the headers
headers = lines[0]
# we consider everythin following the header row as data
for line in lines[1:3]:
line_list = line.split('\t')
header_list = headers.split('\t')
row_dict = dict()
# can you think of a way to fill the row dict?
# hint: check out the zip function
# zip for iterating over two lists at the same time
### YOUR CODE ####
data_dict_list.append(row_dict)
# -
# ### 4.2 Writing csv/tsv files with a header row
#
# You can also write files with headers:
#
# +
row1 = {'name': 'Harry', 'house' : 'Gryffindor'}
row2 = {'name': 'Ron', 'house' : 'Gryffindor'}
row3 = {'name': 'Hannah', 'house': 'Hufflepuff'}
row4 = {'name': 'Leta', 'house' : 'Slytherin'}
data_dict_list = [row1, row2, row2, row4]
for d in data_dict_list:
print(d)
header_row = ### your code here
with open('../Data/csv_data/hp_example.csv', 'w') as outfile:
# write header
for d in data_dict_list:
row_list = d.values()
row = ','.join(values)
outfile.write(row+'\n')
# -
# ## A note on csv/tsv files containing a lot of textual data
#
# If you're dealing with tables containing textual data (e.g. full sentences or tweets), you will realize that splitting on commas or tabs may result in confusion (as text often contains commas and may contain tabs). Consider for instances this file: `../Data/csv_data/debate.csv`.
#
# You may have ideas for a fix regarding this specific file and surely you will eventually find a good solution for it. However, for these more complex CSV files, we recommend that you make use of the **[csv module](https://docs.python.org/3/library/csv.html)**. Feel free to explore it yourself, but do not use it for the exercises and assignments in this course.
# ## Exercises
# ### Exercise 1:
# Have another look at the code below. Can you predict what would happen if you skip some of the steps? How will the data be different?
filename = "../Data/csv_data/baby_names/names_by_state/AK.csv"
with open(filename, "r") as csvfile:
csv_data = []
for row in csvfile:
#row = row.strip("\n") # what would happen if you skip this step?
columns = row.split(",")
csv_data.append(columns)
print(csv_data[0:2])
filename = "../Data/csv_data/baby_names/names_by_state/AK.csv"
with open(filename, "r") as csvfile:
csv_data = []
for row in csvfile:
row = row.strip("\n")
#columns = row.split(",") # what would happen if you skip this step?
csv_data.append(row) # replaced columns by row
print(csv_data[0:2])
# ### Exercise 2:
# Read the csv data in `AK.csv` and store it as a **list of lists**. Now print the following:
# - all names that started with an M given in 1990
# - all unique female names (hint: create a set)
# - all names that were given more than 30 times in a certain year (print name+year)
# - all unique names longer than 8 letters given between 1985 and 1990
# Read the csv data in `AK.csv` and store it as a **list of dicts**. Now print the following:
# - all names that started with an M given in 1990
# - all unique female names (hint: create a set)
# - all names that were given more than 30 times in a certain year (print name+year)
# - all unique names longer than 8 letters given between 1985 and 1990
# ### Exercise 3:
# Can you think of a way to add a header to the TSV file below? Hint: make use of the dictionary keys.
# +
# Create list of dicts
address_book = [
{"name":"John", "e-mail":"<EMAIL>", "phone":"555-1234"},
{"name":"William", "e-mail":"<EMAIL>", "phone":"555-5678"},
{"name":"Jane", "e-mail":"<EMAIL>", "phone":"555-7777"}
]
# Write the list of dicts to a TSV file
outfilename = "../Data/csv_data/address_book.tsv"
with open(outfilename, "w") as outfile:
# Write the header
# your code here
# Write the rows using the values of the dictionaries
for row in address_book:
column_values = row.values()
line = "\t".join(column_values) + '\n'
outfile.write(line)
# -
# ### Exercise 4:
# Now *append* information about Jennifer and Justin (stored as a list) at the bottom of this `address_book.tsv` file (attention: make sure you do not overwrite the contents of the file).
# +
more_people = [
["Jennifer", "<EMAIL>", "555-9876"],
["Justin", "<EMAIL>", "555-5555"]
]
# Append the data to the `address_book.tsv` file
# -
# ### Exercise 5:
# Create a function `load_tabular_data(filename, delimiter)` that receives the filename and a delimiter as input parameters, and returns the file content as a list of lists. Then we can call this function for CSV (with a comma as an argument) and for TSV (with a tabulator as an argument). Also make sure that `delimiter` is a keyword parameter, with a defaul value ','.
# +
# Create your function here
# Now let's test the function
csv_filename = "../Data/csv_data/baby_names/names_by_state/AK.csv"
| Chapters/Chapter 16 - Data formats I (CSV and TSV).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: finalproject
# language: python
# name: finalproject
# ---
from sktime.utils.load_data import load_from_tsfile_to_dataframe
import time
import pickle
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
sns.set()
sns.set(font_scale=1)
sns.set_style('whitegrid')
datasets = ['AtrialFibrillation',
'Epilepsy']
def ts_df_to_array(ts_df, index):
return np.array([ts_df.iloc[index].iloc[i].values for i in range(len(ts_df.iloc[0]))])
def ts_df_to_arrays(ts_df, swapaxes=False):
arrays=[]
for index in range(len(ts_df)):
array = ts_df_to_array(ts_df, index)
if swapaxes:
arrays.append(np.swapaxes(array,0,1))
else:
arrays.append(array)
return np.array(arrays)
def plot_multivariate_dataset_examples_with_seaborn(train_x, train_y, test_x, test_y, synthetic_x_train, synthetic_y_train, dataset_name, label):
num_dimensions = train_x.shape[2]
fig, ax = plt.subplots(nrows=num_dimensions, ncols=3, figsize=(12,2*num_dimensions), sharex=True, sharey=True)
for data_index, (x, y) in enumerate(zip([train_x, synthetic_x_train, test_x], [train_y, synthetic_y_train, test_y])):
label_data = x[y == label]
label_indices = np.arange(len(label_data))
random_index = np.random.choice(label_indices, size=1)
random_example = label_data[random_index]
for dimension_index in range(random_example.shape[2]):
ax[dimension_index,data_index].plot(random_example[:,:,dimension_index][0])
plt.subplots_adjust(wspace=0.1, hspace=0.1)
cols = ["Train", "Synthetic", "Test"]
rows = ['Dimension {}'.format(row) for row in range(1,num_dimensions+1)]
for one_ax, col in zip(ax[0], cols):
one_ax.set_title(col, size='large')
for one_ax, row in zip(ax[:,0], rows):
one_ax.set_ylabel(row)
fig.text(0.5, 0.08, "Time", ha='center', size='large')
fig.suptitle("Dataset: %s, Label: %s" % (dataset_name, label), fontsize=16, y=1)
plt.show()
for dataset_name in datasets:
num_synthetic_ts = 1000
dba_iters = 5
limit_N = False
print("-----------")
start = time.process_time()
print(dataset_name)
train_x, train_y = load_from_tsfile_to_dataframe("../data/%s/%s_TRAIN.ts" % (dataset_name, dataset_name))
test_x, test_y = load_from_tsfile_to_dataframe("../data/%s/%s_TEST.ts" % (dataset_name, dataset_name))
train_x = ts_df_to_arrays(train_x, swapaxes=True)
test_x = ts_df_to_arrays(test_x, swapaxes=True)
num_replicates = train_x.shape[0]
print("# replicates: %d" % (num_replicates))
num_dimensions = train_x.shape[2]
print("# dimensions: %d" % (num_dimensions))
len_series = train_x.shape[1]
print("length of series: %d" % (len_series))
num_classes = len(np.unique(train_y))
print("# classes: %d" % (num_classes))
total_size = num_replicates*num_dimensions*len_series
print("total 'size': %d" % (total_size))
synthetic_x_train = pickle.load(open("../syntheticdata/%s_softdtw_synthetic_x_train_%d_%d_%s.pkl" % (dataset_name, num_synthetic_ts, dba_iters, str(limit_N)), 'rb'))
synthetic_y_train = pickle.load(open("../syntheticdata/%s_softdtw_synthetic_y_train_%d_%d_%s.pkl" % (dataset_name, num_synthetic_ts, dba_iters, str(limit_N)), 'rb'))
labels = np.unique(train_y)
for label in labels:
plot_multivariate_dataset_examples_with_seaborn(train_x, train_y, test_x, test_y, synthetic_x_train, synthetic_y_train, dataset_name, label)
print("Time (s): %f" % (time.process_time() - start))
print("-----------")
| notebooks/rawtimeseriesvisualization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/hafeezjaan77/Code/blob/main/Actylcholinesterase_1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="oeuTF6GcREe0"
#
# ChEMBL Database
#
# The ChEMBL Database is a database that contains curated bioactivity data of more than 2 million compounds. It is compiled from more than 76,000 documents, 1.2 million assays and the data spans 13,000 targets and 1,800 cells and 33,000 indications. [Data as of March 25, 2020; ChEMBL version 26].
# Installing libraries
#
# Install the ChEMBL web service package so that we can retrieve bioactivity data from the ChEMBL Database.
#
# + colab={"base_uri": "https://localhost:8080/"} id="uQmY8tXIPN_Y" outputId="deafda1b-9c16-44fa-b86e-9bf0520ee052"
pip install chembl_webresource_client
# + id="LguPniMlQek6"
import pandas as pd
from chembl_webresource_client.new_client import new_client
# + [markdown] id="70IRCSo7Q2gb"
# Search for Target protein
# Target search for Acetylcholinesterase
#
#
# + id="Tk93iXyPRcPN" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="8a211153-f4d7-4cd0-9f46-0133573a40b2"
# Target search for SLP1
target = new_client.target
target_query = target.search('acetylcholinesterase')
targets = pd.DataFrame.from_dict(target_query)
targets
# + [markdown] id="sUyFJPBDg8ps"
#
# **Select and retrieve bioactivity data for Human Acetylcholinesterase (first entry)**
#
# We will assign the fifth entry (which corresponds to the target protein, Human Acetylcholinesterase) to the selected_target variable
#
# + colab={"base_uri": "https://localhost:8080/", "height": 36} id="Tedm1Dmiaqfu" outputId="930bb021-86b7-4773-ca36-c8afa69b2a04"
selected_target = targets.target_chembl_id[0]
selected_target
# + [markdown] id="sY4tCVH2hJGn"
# Here, we will retrieve only bioactivity data for Human Acetylcholinesterase (CHEMBL220) that are reported as pChEMBL values.
# + id="rsST5_E7axHa"
activity = new_client.activity
res = activity.filter(target_chembl_id=selected_target).filter(standard_type="IC50")
# + id="inQ4XjEta5An"
df = pd.DataFrame.from_dict(res)
# + colab={"base_uri": "https://localhost:8080/", "height": 791} id="SQ8w3gTCb3uD" outputId="c8dc603f-fdc4-45a3-959f-72bfbc61942b"
df
# + [markdown] id="mYToJZFShTlo"
# Finally we will save the resulting bioactivity data to a CSV file bioactivity_data.csv.
# + id="ZnWNjGZVe2Eu"
df.to_csv('acetylcholinesterase_01_bioactivity_data_raw.csv', index=False)
# + [markdown] id="SBJvCKlGha6g"
#
# **Handling missing data**
#
# If any compounds has missing value for the standard_value and canonical_smiles column then drop it.
#
# + colab={"base_uri": "https://localhost:8080/", "height": 828} id="DOGKb2cIfAaE" outputId="c5207738-bb6d-46df-a2c0-05c627ddb5a2"
df2 = df[df.standard_value.notna()]
df2 = df2[df.canonical_smiles.notna()]
df2
# + colab={"base_uri": "https://localhost:8080/"} id="hCScgs9vfJ4a" outputId="bd7c0098-3abe-490f-d217-399612bb1b3d"
len(df2.canonical_smiles.unique())
# + colab={"base_uri": "https://localhost:8080/", "height": 791} id="vS9nBjlRfOhI" outputId="bc1292c4-f4c4-4ad0-87e9-a8c36f891cc4"
df2_nr = df2.drop_duplicates(['canonical_smiles'])
df2_nr
# + [markdown] id="VuqXvnxshmTD"
#
# **Data pre-processing of the bioactivity data**
#
# Combine the 3 columns (molecule_chembl_id,canonical_smiles,standard_value) and bioactivity_class into a DataFrame
# + colab={"base_uri": "https://localhost:8080/", "height": 424} id="voi3mwYDfWc8" outputId="33379bda-e512-40cf-92da-82a982c82f4c"
selection = ['molecule_chembl_id','canonical_smiles','standard_value']
df3 = df2_nr[selection]
df3
# + [markdown] id="5JafTvXRhxoS"
# Saves dataframe to CSV file
# + id="GjEpbgp9fdE9"
df3.to_csv('acetylcholinesterase_02_bioactivity_data_preprocessed.csv', index=False)
# + [markdown] id="Yqpo8-Vvh2qp"
#
# **Labeling compounds as either being active, inactive or intermediate**
#
# The bioactivity data is in the IC50 unit. Compounds having values of less than 1000 nM will be considered to be active while those greater than 10,000 nM will be considered to be inactive. As for those values in between 1,000 and 10,000 nM will be referred to as intermediate.
#
# + id="gTZdHaPbfiKv"
df4 = pd.read_csv('acetylcholinesterase_02_bioactivity_data_preprocessed.csv')
# + id="aMOogCcGfoKW"
bioactivity_threshold = []
for i in df4.standard_value:
if float(i) >= 10000:
bioactivity_threshold.append("inactive")
elif float(i) <= 1000:
bioactivity_threshold.append("active")
else:
bioactivity_threshold.append("intermediate")
# + colab={"base_uri": "https://localhost:8080/", "height": 424} id="ZGlda54afuaJ" outputId="bc4583c9-6170-442d-e3f0-4f04c74de433"
bioactivity_class = pd.Series(bioactivity_threshold, name='class')
df5 = pd.concat([df4, bioactivity_class], axis=1)
df5
# + [markdown] id="5n4xaj4TiH7A"
# Saves dataframe to CSV file
# + id="yBQZlQu7f1QE"
df5.to_csv('acetylcholinesterase_03_bioactivity_data_curated.csv', index=False)
# + colab={"base_uri": "https://localhost:8080/", "height": 135} id="VIAecB4zf50R" outputId="6adca98a-c3ea-4faf-d1f7-af1876aa2853"
zip acetylcholinesterase.zip *.csv
# + colab={"base_uri": "https://localhost:8080/"} id="1fj1EENFgJ14" outputId="bc4cfd8e-1feb-4273-f453-ecdb4283cd5c"
# ls -l
| Actylcholinesterase_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/AGoyal0512/Madison-Crime/blob/main/Heat%20Maps-Time%20Exploration.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="g4vM5YApPujS"
import csv
import pandas as pd
import numpy as np
import datetime
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sb
# + id="mX5IVWv9P38R" outputId="b74676b5-aaf5-4f6e-fb43-c639306a6830" colab={"base_uri": "https://localhost:8080/"}
# !git clone https://github.com/AGoyal0512/Madison-Crime.git
# + id="yftFJ9CWPujV"
data = pd.read_csv('Madison-Crime/cleaned_df.csv.zip', compression='zip')
# + id="OMbQaVAQPujW" outputId="d09b1aeb-715b-4cb9-8bc6-0a407fdf2ea5" colab={"base_uri": "https://localhost:8080/", "height": 884}
data
# + id="OBowaj2bPujX"
data['my_dates'] = pd.to_datetime(data[['Year','Month', 'Day']])
data['day_of_week'] = data['my_dates'].dt.day_name()
# data['weekday'] = datetime.date(data['Year','Month', 'Day'])
# + id="6as_DSkaPujX" outputId="178ca214-05b5-4606-9724-21e7cebda44e" colab={"base_uri": "https://localhost:8080/", "height": 424}
is_rob=data['IncidentType']=="Robbery"
rob= data[is_rob]
rob=rob[['Hour','day_of_week','IncidentType']]
rob
# + id="4Y32BJ_pPujY" outputId="4550dc5c-4088-4865-8903-f809529c19a4" colab={"base_uri": "https://localhost:8080/", "height": 865}
new=rob.groupby(['Hour','day_of_week'])["IncidentType"].count().unstack()
new
# + id="1OtPmyvjPujY" outputId="93188e99-a3d3-4b10-b685-ce70ce90a05f" colab={"base_uri": "https://localhost:8080/", "height": 556}
fig, ax = plt.subplots(figsize=(11, 9))
sb.heatmap(new,cmap="Blues")
plt.show()
# + id="k7FxaFRcPujZ" outputId="f30b826b-9f6b-48d4-b51e-bc97038a5137" colab={"base_uri": "https://localhost:8080/", "height": 424}
is_vio=data['IncidentType']=="Weapons Violation"
vio= data[is_vio]
vio=vio[['Hour','day_of_week','IncidentType']]
vio
# + id="QEiEhHjWPuja" outputId="07d6fb94-c8fd-4556-e048-65d467016a09" colab={"base_uri": "https://localhost:8080/", "height": 865}
new1=vio.groupby(['Hour','day_of_week'])["IncidentType"].count().unstack()
new1
# + id="sL0SKWS_Pujb" outputId="5ca1fd6f-7323-401c-8186-8d0275962cd3" colab={"base_uri": "https://localhost:8080/", "height": 556}
fig, ax = plt.subplots(figsize=(11, 9))
sb.heatmap(new1,cmap="Blues")
plt.show()
# + id="xo0KgjNiPujc" outputId="25c6e97c-b680-4525-bb8e-99842bd11dd4" colab={"base_uri": "https://localhost:8080/", "height": 424}
is_bat=data['IncidentType']=="Battery"
bat= data[is_bat]
bat=bat[['Hour','day_of_week','IncidentType']]
bat
# + id="21wV8s6iPujc" outputId="fb7343f9-bdbb-4fd5-950d-d1f4e24ee12c" colab={"base_uri": "https://localhost:8080/", "height": 865}
new2=bat.groupby(['Hour','day_of_week'])["IncidentType"].count().unstack()
new2
# + id="U-BWr7c5Pujc" outputId="3068410a-e2d4-48a0-94aa-d2873a7350c8" colab={"base_uri": "https://localhost:8080/", "height": 552}
fig, ax = plt.subplots(figsize=(11, 9))
sb.heatmap(new2,cmap="Blues")
plt.show()
# + id="mvu-XqmbPujd" outputId="aafbf295-c67a-447c-89ee-62927a43cbfe" colab={"base_uri": "https://localhost:8080/", "height": 424}
is_thef=data['IncidentType']=="Theft"
thef= data[is_thef]
thef=thef[['Hour','day_of_week','IncidentType']]
thef
# + id="RMktBwcoPujd" outputId="43b61f50-47d0-4ff9-a4c2-3957a9c7f9cb" colab={"base_uri": "https://localhost:8080/", "height": 865}
new3=thef.groupby(['Hour','day_of_week'])["IncidentType"].count().unstack()
new3
# + id="4v5TatkIPujd" outputId="6cbdf8a1-3a02-403c-a806-24f089d5aae1" colab={"base_uri": "https://localhost:8080/", "height": 552}
fig, ax = plt.subplots(figsize=(11, 9))
sb.heatmap(new3,cmap="Blues")
plt.show()
# + id="pDjF655MPuje" outputId="db8e4626-96d7-4685-d771-02eb61c19c7f" colab={"base_uri": "https://localhost:8080/", "height": 424}
is_burg=data['IncidentType']=="Residential Burglary"
burg= data[is_burg]
burg=burg[['Hour','day_of_week','IncidentType']]
burg
# + id="Sts-hhYhPuje"
new4=burg.groupby(['Hour','day_of_week'])["IncidentType"].count().unstack()
new4
# + id="aLfkSsa0Puje" outputId="729f7796-fc23-4994-e313-aecb8e980753" colab={"base_uri": "https://localhost:8080/", "height": 556}
fig, ax = plt.subplots(figsize=(11, 9))
sb.heatmap(new4,cmap="Blues")
plt.show()
| Heat Maps-Time Exploration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 0. Dependências
# +
import numpy as np
import matplotlib.pyplot as plt
from sklearn.metrics import accuracy_score
# %matplotlib inline
# -
# # 1. Introdução
# **Boosting** é uma abordadem de Machine Learning baseada na ideia de criar regras de predição com alta acurácia pela combinação de regras fracas e imprecisas. Boosting também é uma das abordages de aprendizagem supervisionada mais populares e bem sucedidas.
#
# O **Adaboost** é a técnica mais popular de boosting. A ideia básica do algoritmo é, para cada iteração, treinar um classificador fraco (accurácia > 50%) dando preferência (maiores pesos) aos exemplos incorretamente classificados pelo classificador anterior. Ao final, um classificador forte é construído pelo voto ponderado de cada um dos classificadores.
#
# **Vantagens:**
# - Fácil implementação
# - Rápido
# - Apenas um parâmetro para tunning (número de estimadores)
# - Pode ser usado como seletor de características
# - Boa generalização
# - Versátil
#
# **Desvantagens:**
# - Não é robusto a presença de ruído uniforme
# - Muitos classificadores fracos podem acarretar em overfitting
#
#
# __Pseudo-algoritmo__
#
# 1. Para cada um dos n_estimadores:
# - treine um classificador binário $C_i$ com $y \in \{-1, +1\}$ de acordo com os pesos $w$
# - Calcula as predições do classificador
# - Calcule a taxa de erro ponderada
#
# $$e = w*1(y \neq y_{pred}) = 1 - w*1(y=y_{pred}) \tag{1}$$
#
# - Calcule os coeficientes
#
# $$\alpha_i = 0.5 * \log{\frac{1-e}{e}} \tag{2}$$
#
# - Atualize os pesos
#
# $$w = w*\exp^{-\alpha_i *y*y_{pred}} \tag{3}$$
#
# - Normalize os pesos
#
# $$w = \frac{w}{\sum w} \tag{4}$$
#
# O Classificador final será dado por:
#
# $$sign(\sum_i{\alpha_i*C_i.predict(x_{test})}) \tag{5}$$
# # 2. Dados
# Os dados são baseados na [aula sobre boosting](https://www.youtube.com/watch?v=u1MXf5N3wYU) do [curso de Machine Learning da Udacity](https://br.udacity.com/course/machine-learning--ud262). Eu tentei reproduzir, o mais fielmente possível, os dados do gráfico mostrado na aula.
# +
x = np.array([[1.0, 6.0],[1.5, 3.0],[3.2, 8.0],[5.2, 9.0],[7.0, 7.0],
[2.2, 2.0],[2.5, 6.5],[5.3, 5.5],[8.5, 8.5],[8.5, 3.2]])
y = np.array([1, 1, 1, 1, 1, -1, -1, -1, -1, -1])
plt.scatter(x[:5,0], x[:5,1], s=300, marker='+')
plt.scatter(x[5:,0], x[5:,1], s=300, marker='_', color='red')
plt.xlim(0, 10)
# -
# # 3. Implementação
# +
def weighted_accuracy(pred, y, weights):
return sum([w for p,t,w in zip(pred, y, weights) if p==t])/sum(weights)
def sign(x):
if x == 0: return 0
return 1 if x > 0 else -1
# -
# Como o Adaboost pode utilizar qualquer classificador como base, nós vamos implementar um classificador que só traça linhas em cada atributo (*feature*) tentando maximizar a acurácia.
class LineClassifier():
def __init__(self, weights=np.array([])):
self.weights = weights
self.value, self.col = 0, 0
def fit(self, x, y):
column_count = len(x[0])
if len(self.weights) == 0:
self.weights = [1.0/len(x)]*len(x)
best_accuracy = 0.0
for col in range(column_count):
column_values = [row[col] for row in x]
for value in np.arange(min(column_values), max(column_values), 0.1):
accuracy = weighted_accuracy(LineClassifier.predict_(x, col, value), y, self.weights)
if accuracy > best_accuracy:
best_accuracy = accuracy
self.col, self.value = col, value
def predict(self, x):
return LineClassifier.predict_(x, self.col, self.value)
@staticmethod
def predict_(x, col, value):
if col == 0:
return [1 if row[col] < value else -1 for row in x]
else:
return [1 if row[col] > value else -1 for row in x]
class AdaBoost():
def __init__(self, base_estimator=LineClassifier, minHitRate=0.5, n_estimators=50):
self.classifiers = []
self.alphas = []
self.weights = []
self.base_estimator = base_estimator
self.n_estimators = n_estimators
self.minHitRate = minHitRate
def fit(self, x, y):
from math import log, e
ln = lambda x: log(x)/log(e)
if len(self.weights) == 0:
self.weights = [1.0/len(x)]*len(x)
for n in range(self.n_estimators):
clf = self.base_estimator()
clf.weights = self.weights
clf.fit(x, y)
pred = clf.predict(x)
error = 1.0 - weighted_accuracy(pred, y, self.weights) # Eq. (1)
if(error < self.minHitRate):
alpha = 0.5 * ln((1-error)/error) # Eq. (2)
self.weights = [w*(e**(-alpha*p*t)) for p,t,w in zip(pred, y, self.weights)] # Eq. (3)
self.weights = [w/sum(self.weights) for w in self.weights] # Eq. (4)
self.classifiers.append(clf)
self.alphas.append(alpha)
print('[{}]: ε={:.2f} α={:.2f}'.format(n, error, alpha))
def predict(self, x):
predictions = []
for row in x:
weak_sum = 0.0
for alpha, clf in zip(self.alphas, self.classifiers):
weak_sum += alpha*clf.predict([row])[0]
predictions.append(sign(weak_sum)) # Eq. (5)
return np.array(predictions)
# # 4. Teste
# A gente pode conferir a implementação por esse [link](https://alliance.seas.upenn.edu/~cis520/wiki/index.php?n=lectures.boosting).
# +
ada = AdaBoost(base_estimator=LineClassifier, n_estimators=3, minHitRate=0.3)
ada.fit(x, y)
y_pred = ada.predict(x)
print('y_true:', y)
print('y_pred:', y_pred)
# +
xv, yv = np.meshgrid(np.linspace(0, 10, 100), np.linspace(0, 10, 100))
xyv = np.concatenate((xv, yv), axis=1)
pred_2 = []
for i in range(xv.shape[0]):
for j in range(xv.shape[1]):
pred_2.append(ada.predict([[xv[i,j], yv[i,j]]]))
pred_2 = np.array(pred_2).reshape(xv.shape)
plt.contourf(xv, yv, pred_2, cmap=plt.cm.Spectral, alpha=0.6)
plt.scatter(x[:5,0], x[:5,1], s=300, marker='+')
plt.scatter(x[5:,0], x[5:,1], s=300, marker='_', color='red')
plt.xlim(0,10)
plt.ylim(0,10)
# -
# # 5. Referências
# - [Machine Learning by Georgia Tech (Udacity)](https://br.udacity.com/course/machine-learning--ud262)
# - [Playlist about Adaboost on Youtube (same as above)](https://www.youtube.com/watch?v=w75WyRjRpAg)
# - [CIS 520 - Machine Learning 2018](https://alliance.seas.upenn.edu/~cis520/wiki/index.php?n=lectures.boosting)
| Adaboost.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Django Shell-Plus
# language: python
# name: django_extensions
# ---
import pandas as pd
file = "dedupe/output_names.csv"
df = pd.read_csv(file)
for x in df.groupby('Cluster ID'):
cur_df = x[1]
new_name = "/".join(set("/".join([f"{x}" for x in cur_df['Name'].unique()]).split('/')))
| dedupe_names.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/jesuspablo888/daa_2021_1/blob/master/13_enero.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="PntaktO6L4FY"
class NodoArbol:
def __init__(self, value, left=None, rigth=None):
self.data = value
self.left =left
self.rigth = rigth
# + [markdown] id="LPk1iW1aMSsl"
# # Árbol bninario de busqueda
# Los nodos a la izquierda son menores a la raiz y los nodos a la derecha son mayores a la raiz.
# Pueden ser recorridos en: pre.orden, in-orden y post-orden.
# + id="SMUJI7pxMLGN"
class BinarySearchTree:
def __init__(self):
self.__root = None
def insert(self, value):
if self.__root == None:
self.__root = NodoArbol(value,None,None)
else:
# preguntar si value es menor que root de ser el caso
#insertar ala isquierda PERO.. puede ser le caos que el
# sub arbol tenga muchos elementos
self.__insert_nodo__(self.__root, value)
def __insert_nodo__(self, nodo, value):
if nodo.data == value:
pass # no se admiten valores repetidos
elif value < nodo.data:# si es true va a la izq
if nodo.left == None: # si hay espacio lo colaoca ahi
nodo.left = NodoArbol(value)# inserta
else:
self.__insert_nodo__(nodo.left,value) # buscar en sub arbol izq
else:
if nodo.right == None:
nodo.right = TreeNode(value)
else:
self.__insert_nodo__(nodo.right, value) # buscar en sub arbol ider
# + id="2U9sBO5gclWM"
bst = BinarySearchTree()
bst.insert(50)
bst.insert(30)
bst.insert(20)
| 13_enero.ipynb |