code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import cv2
import numpy as np
import copy
import math
import time
import webbrowser
import os
import pyautogui as py
import glob
import time
# +
def nothing(x):
pass
image_x, image_y = 64,64
s=''
cap_region_x_begin=0.6 # start point/total width
cap_region_y_end=0.4 # start point/total width
threshold = 60 # BINARY threshold
blurValue = 41 # GaussianBlur parameter
bgSubThreshold = 50
learningRate = 0
val=50.0
image_x, image_y = 64, 64
# variables
isBgCaptured = 0 # bool, whether the background captured
triggerSwitch = False
# -
import pickle
clf = pickle.load(open('modelhand6.sav', 'rb'))
def ellipse_variance(contour):
M=cv2.moments(contour)
cX=int(M['m10']/M['m00'])
cY=int(M['m01']/M['m00'])
v=np.zeros((1,2,1))
for cord in contour:
x=np.array([[[cord[0][0]-cX],[cord[0][1]-cY]]])
v=np.vstack((v,x))
# print(v)
# print(x)
# print(cord)
# print(cord[0][0])
# print(v.shape)
c_ellipse=np.zeros((2,2))
for i in v:
c_ellipse+=np.dot(i,np.transpose(i))
c_ellipse/=contour.shape[0]
# print(c_ellipse)
c_ellipse_inv=np.linalg.inv(c_ellipse)
d=np.zeros((1,1))
for i in v:
d=np.vstack((d,np.sqrt(np.dot(np.dot(np.transpose(i),c_ellipse_inv),i))))
d=d[1:]
mu=1/contour.shape[0]*np.sum(d)
sigma=np.sqrt(1/contour.shape[0]*np.sum(np.square(d-mu)))
return mu/sigma
# +
t=[]
ma=0
isBgCaptured=0
cou=0
cv2.namedWindow("Trackbars")
cv2.createTrackbar("L - H", "Trackbars", 0, 179, nothing)
cv2.createTrackbar("L - S", "Trackbars", 0, 255, nothing)
cv2.createTrackbar("L - V", "Trackbars", 0, 255, nothing)
cv2.createTrackbar("U - H", "Trackbars", 179, 179, nothing)
cv2.createTrackbar("U - S", "Trackbars", 255, 255, nothing)
cv2.createTrackbar("U - V", "Trackbars", 50, 255, nothing)
def printThreshold(thr):
print("! Changed threshold to "+str(thr))
def removeBG(frame):
fgmask = bgModel.apply(frame,learningRate=learningRate)
#kernel = cv2.getStructuringElement(cv2.MORPH_ELLIPSE, (3, 3))
#res = cv2.morphologyEx(fgmask, cv2.MORPH_OPEN, kernel)
kernel = np.ones((4, 4), np.uint8)
fgmask = cv2.erode(fgmask, kernel, iterations=1)
kernel = np.ones((1, 1), np.uint8)
#fgmask = cv2.dilate(fgmask, kernel, iterations=6)
res = cv2.bitwise_and(frame, frame, mask=fgmask)
return res
camera = cv2.VideoCapture(0)
camera.set(10,200)
cv2.namedWindow('threshold')
cv2.createTrackbar('trh1', 'threshold', threshold, 100, printThreshold)
ret, fra = camera.read()
fra=cv2.cvtColor(fra,cv2.COLOR_BGR2GRAY)
for i in range(480):
for j in range(640):
fra[i][j]=1
while camera.isOpened():
b=0
k = cv2.waitKey(1)
lower=np.array([90,90,90])
upper=np.array([240,240,240])
ret, frame = camera.read()
alpha = 1
beta = -70
frame = cv2.addWeighted(frame, alpha, np.zeros(frame.shape, frame.dtype),0, beta)
#print(frame.shape)
blur = cv2.GaussianBlur(frame, (25, 25), 0)
lower_blue = np.array([0, 12, 92])
upper_blue = np.array([179, 255, 230])
hsv = cv2.cvtColor(frame, cv2.COLOR_BGR2HSV)
mask = cv2.inRange(hsv, lower_blue, upper_blue)
#frame = cv2.bitwise_and(frame, frame, mask=mask)
t=[]
#frame = cv2.cvtColor(frame, cv2.COLOR_BGR2RGB)
#threshold = cv2.getTrackbarPos('trh1', 'threshold')
threshold = 60
frame = cv2.bilateralFilter(frame, 5, 50, 100) # smoothing filter
frame = cv2.flip(frame, 1) # flip the frame horizontally
cv2.rectangle(frame, (int(cap_region_x_begin * frame.shape[1]), 0),
(frame.shape[1], int(cap_region_y_end * frame.shape[0])), (255, 0, 0), 2)
cv2.imshow('original', frame)
if isBgCaptured == 1:
cou+=1
# this part wont run until background captured
#mas = cv2.inRange(frame, lower, upper)
#img = cv2.bitwise_and(frame, frame, mask=mas)
img = removeBG(frame)
img = img[0:int(cap_region_y_end * frame.shape[0]),
int(cap_region_x_begin * frame.shape[1]):frame.shape[1]] # clip the ROI
# for i in range(img.shape[0]):
# for j in range(img.shape[1]):
# if img[i][j][0]<140 or img[i][j][0]>180:
# img[i][j][0]=0
# img[i][j][1]=0
# img[i][j][2]=0
# l_h = cv2.getTrackbarPos("L - H", "Trackbars")
# l_s = cv2.getTrackbarPos("L - S", "Trackbars")
# l_v = cv2.getTrackbarPos("L - V", "Trackbars")
# u_h = cv2.getTrackbarPos("U - H", "Trackbars")
# u_s = cv2.getTrackbarPos("U - S", "Trackbars")
# u_v = cv2.getTrackbarPos("U - V", "Trackbars")
# lower_blue = np.array([0, 23, 92])
# upper_blue = np.array([179, 255, 225])
#hsv = cv2.cvtColor(img, cv2.COLOR_BGR2HSV)
#mask = cv2.inRange(hsv, lower_blue, upper_blue)
#cv2.imshow("m", hsv)
cv2.imshow('mask', img)
# convert the image into binary image
gray = cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)
blur = cv2.GaussianBlur(gray, (blurValue, blurValue), 0)
cv2.imshow('blur', blur)
#ret, thresh = cv2.threshold(blur, threshold, 255, cv2.THRESH_BINARY)
ret3,thresh = cv2.threshold(blur,0,255,cv2.THRESH_BINARY+cv2.THRESH_OTSU)
kernel = np.ones((5,5),np.uint8)
opening = cv2.morphologyEx(thresh, cv2.MORPH_OPEN, kernel)
kernel = np.ones((1,1),np.uint8)
dilation = cv2.dilate(opening,kernel,iterations = 4)
im2,contours,hierarchy=cv2.findContours(dilation,cv2.RETR_EXTERNAL,cv2.CHAIN_APPROX_SIMPLE)
# print(len(contours))
# for cnt in contours:
# a = cv2.contourArea(cnt)
# if a>ma:
# c=cnt;
if len(contours)>0:
req=max(contours,key=cv2.contourArea)
# print(req.shape)
#req=req[0]
#req=c
area=cv2.contourArea(req)
#eccentricity
dim_rect=cv2.minAreaRect(req)[1]
para1=dim_rect[1]/dim_rect[0]
#circularity
perimeter=cv2.arcLength(req,True)
para2=4*np.pi*area/(perimeter**2)
#rectangularity
para3=area/(dim_rect[1]*dim_rect[0])
#convexity
hull=cv2.convexHull(req)
para4=cv2.arcLength(hull,True)/perimeter
#solidity
para5=area/cv2.contourArea(hull)
#ellipse variance
para6=ellipse_variance(req)
#np.vstack((data,np.array([para1,para2,para3,para4,para5,para6,ord(label)])))
# for cnt in contours:
# a = cv2.contourArea(cnt)
# if a>ma:
# c=cnt;
# x,y,w,h = cv2.boundingRect(c)
# imcrop = thresh[y:y+h, x:x+w]
# cv2.imshow('ori', thresh)
cv2.imshow('final', dilation)
# cv2.imshow('final', imcrop)
# img = cv2.resize(imcrop, (image_x, image_y), interpolation = cv2.INTER_AREA)
if cou%50 == 0:
time.sleep(2)
g=[para1,para2,para3,para4,para5,para6]
t.append(g)
predicted = clf.predict(t)
print(predicted)
if predicted == 1:
#time.sleep(2)
py.FAILSAFE = False
py.typewrite([' '])
print(0)
elif predicted == 2:
py.FAILSAFE = False
py.typewrite(['f'])
print(2)
elif predicted == 3:
count=0
cv2.waitKey(10)
cv2.destroyAllWindows()
while True:
count+=1
ret, fram = camera.read()
blur = cv2.GaussianBlur(fram, (25, 25), 0)
hsv = cv2.cvtColor(fram, cv2.COLOR_BGR2HSV)
mask = cv2.inRange(hsv, lower_blue, upper_blue)
fram = cv2.bitwise_and(fram, fram, mask=mask)
fram = cv2.bilateralFilter(fram, 5, 50, 100) # smoothing filter
fram = cv2.flip(fram, 1) # flip the frame horizontally
cv2.rectangle(fram, (int(cap_region_x_begin * fram.shape[1]), 0),
(fram.shape[1], int(cap_region_y_end * fram.shape[0])), (255, 0, 0), 2)
cv2.imshow('original', fram)
if count>=200:
cv2.waitKey(10)
cv2.destroyAllWindows()
break
if count>=50:
imgi = removeBG(fram)
imgi = imgi[0:int(cap_region_y_end * fram.shape[0]),
int(cap_region_x_begin * fram.shape[1]):fram.shape[1]]
cv2.imshow('mask', imgi)
gray = cv2.cvtColor(imgi, cv2.COLOR_BGR2GRAY)
blur = cv2.GaussianBlur(gray, (blurValue, blurValue), 0)
#cv2.imshow('blur', blur)
#ret, thresh = cv2.threshold(blur, threshold, 255, cv2.THRESH_BINARY)
ret3,thresh = cv2.threshold(blur,0,255,cv2.THRESH_BINARY+cv2.THRESH_OTSU)
kernel = np.ones((5,5),np.uint8)
opening = cv2.morphologyEx(thresh, cv2.MORPH_OPEN, kernel)
kernel = np.ones((5,5),np.uint8)
dilation = cv2.dilate(opening,kernel,iterations = 1)
im2,contours,hierarchy=cv2.findContours(dilation,cv2.RETR_EXTERNAL,cv2.CHAIN_APPROX_SIMPLE)
if len(contours)>0:
req=max(contours,key=cv2.contourArea)
x,y,w,h = cv2.boundingRect(req)
# print(req.shape)
#req=req[0]
#req=c
#area=cv2.contourArea(req)
area=w*h
print(area)
a=area/50000
if a<b:
py.hotkey('ctrl', 'f2')
elif a>b:
py.hotkey('ctrl', 'f3')
b=a
continue
else:
cv2.waitKey(10)
cv2.destroyAllWindows()
break
elif predicted == 4:
count=0
#cv2.waitKey(10)
#cv2.destroyAllWindows()
while True:
count+=1
ret, fram = camera.read()
blur = cv2.GaussianBlur(fram, (25, 25), 0)
hsv = cv2.cvtColor(fram, cv2.COLOR_BGR2HSV)
mask = cv2.inRange(hsv, lower_blue, upper_blue)
fram = cv2.bitwise_and(fram, fram, mask=mask)
fram = cv2.bilateralFilter(fram, 5, 50, 100) # smoothing filter
fram = cv2.flip(fram, 1) # flip the frame horizontally
cv2.rectangle(fram, (int(cap_region_x_begin * fram.shape[1]), 0),
(fram.shape[1], int(cap_region_y_end * fram.shape[0])), (255, 0, 0), 2)
cv2.imshow('original', fram)
if count>=200:
cv2.waitKey(10)
cv2.destroyAllWindows()
break
if count>=50:
imgi = removeBG(fram)
imgi = imgi[0:int(cap_region_y_end * fram.shape[0]),
int(cap_region_x_begin * fram.shape[1]):fram.shape[1]]
#cv2.imshow('mask', imgi)
gray = cv2.cvtColor(imgi, cv2.COLOR_BGR2GRAY)
blur = cv2.GaussianBlur(gray, (blurValue, blurValue), 0)
#cv2.imshow('blur', blur)
#ret, thresh = cv2.threshold(blur, threshold, 255, cv2.THRESH_BINARY)
ret3,thresh = cv2.threshold(blur,0,255,cv2.THRESH_BINARY+cv2.THRESH_OTSU)
kernel = np.ones((5,5),np.uint8)
opening = cv2.morphologyEx(thresh, cv2.MORPH_OPEN, kernel)
kernel = np.ones((5,5),np.uint8)
dilation = cv2.dilate(opening,kernel,iterations = 1)
im2,contours,hierarchy=cv2.findContours(dilation,cv2.RETR_EXTERNAL,cv2.CHAIN_APPROX_SIMPLE)
if len(contours)>0:
req=max(contours,key=cv2.contourArea)
x,y,w,h = cv2.boundingRect(req)
# print(req.shape)
#req=req[0]
#req=c
#area=cv2.contourArea(req)
area=w*h
print(area)
a=area/50000
if a<b:
py.hotkey('ctrl', '2')
elif a>b:
py.hotkey('ctrl', '1')
b=a
continue
else:
cv2.waitKey(10)
cv2.destroyAllWindows()
break
elif predicted == 5:
count=0
cv2.waitKey(10)
cv2.destroyAllWindows()
while True:
count+=1
ret, fram = camera.read()
blur = cv2.GaussianBlur(fram, (25, 25), 0)
hsv = cv2.cvtColor(fram, cv2.COLOR_BGR2HSV)
mask = cv2.inRange(hsv, lower_blue, upper_blue)
frame = cv2.bitwise_and(fram, fram, mask=mask)
fram = cv2.bilateralFilter(fram, 5, 50, 100) # smoothing filter
fram = cv2.flip(fram, 1) # flip the frame horizontally
cv2.rectangle(fram, (int(cap_region_x_begin * fram.shape[1]), 0),
(fram.shape[1], int(cap_region_y_end * fram.shape[0])), (255, 0, 0), 2)
#cv2.imshow('original', fram)
if count>=200:
cv2.waitKey(10)
cv2.destroyAllWindows()
break
if count>=50:
imgi = removeBG(fram)
imgi = imgi[0:int(cap_region_y_end * fram.shape[0]),
int(cap_region_x_begin * fram.shape[1]):fram.shape[1]]
#cv2.imshow('mask', imgi)
gray = cv2.cvtColor(imgi, cv2.COLOR_BGR2GRAY)
blur = cv2.GaussianBlur(gray, (blurValue, blurValue), 0)
#cv2.imshow('blur', blur)
#ret, thresh = cv2.threshold(blur, threshold, 255, cv2.THRESH_BINARY)
ret3,thresh = cv2.threshold(blur,0,255,cv2.THRESH_BINARY+cv2.THRESH_OTSU)
kernel = np.ones((5,5),np.uint8)
opening = cv2.morphologyEx(thresh, cv2.MORPH_OPEN, kernel)
kernel = np.ones((5,5),np.uint8)
dilation = cv2.dilate(opening,kernel,iterations = 1)
im2,contours,hierarchy=cv2.findContours(dilation,cv2.RETR_EXTERNAL,cv2.CHAIN_APPROX_SIMPLE)
if len(contours)>0:
req=max(contours,key=cv2.contourArea)
x,y,w,h = cv2.boundingRect(req)
# print(req.shape)
#req=req[0]
#req=c
#area=cv2.contourArea(req)
area=w*h
print(area)
a=area/50000
# py.keyDown('ctrl')
if a>b:
py.hotkey('ctrl', 'right')
elif a<b:
py.hotkey('ctrl', 'left')
# py.keyUp('ctrl')
b=a
continue
else:
cv2.waitKey(10)
cv2.destroyAllWindows()
break
elif predicted==6:
py.FAILSAFE = False
#close window
time.sleep(0.5)
py.hotkey('alt', 'f4')
elif predicted==7:
count=0
cv2.waitKey(10)
cv2.destroyAllWindows()
while True:
count+=1
ret, fram = camera.read()
blur = cv2.GaussianBlur(fram, (25, 25), 0)
hsv = cv2.cvtColor(fram, cv2.COLOR_BGR2HSV)
mask = cv2.inRange(hsv, lower_blue, upper_blue)
fram = cv2.bitwise_and(fram, fram, mask=mask)
fram = cv2.bilateralFilter(fram, 5, 50, 100) # smoothing filter
fram = cv2.flip(fram, 1) # flip the frame horizontally
cv2.rectangle(fram, (int(cap_region_x_begin * fram.shape[1]), 0),
(fram.shape[1], int(cap_region_y_end * fram.shape[0])), (255, 0, 0), 2)
#cv2.imshow('original', fram)
if count>=200:
cv2.waitKey(10)
cv2.destroyAllWindows()
break
if count>=50:
imgi = removeBG(fram)
imgi = imgi[0:int(cap_region_y_end * fram.shape[0]),
int(cap_region_x_begin * fram.shape[1]):fram.shape[1]]
#cv2.imshow('mask', imgi)
gray = cv2.cvtColor(imgi, cv2.COLOR_BGR2GRAY)
blur = cv2.GaussianBlur(gray, (blurValue, blurValue), 0)
#cv2.imshow('blur', blur)
#ret, thresh = cv2.threshold(blur, threshold, 255, cv2.THRESH_BINARY)
ret3,thresh = cv2.threshold(blur,0,255,cv2.THRESH_BINARY+cv2.THRESH_OTSU)
kernel = np.ones((5,5),np.uint8)
opening = cv2.morphologyEx(thresh, cv2.MORPH_OPEN, kernel)
kernel = np.ones((5,5),np.uint8)
dilation = cv2.dilate(opening,kernel,iterations = 1)
im2,contours,hierarchy=cv2.findContours(dilation,cv2.RETR_EXTERNAL,cv2.CHAIN_APPROX_SIMPLE)
if len(contours)>0:
req=max(contours,key=cv2.contourArea)
x,y,w,h = cv2.boundingRect(req)
# print(req.shape)
#req=req[0]
#req=c
#area=cv2.contourArea(req)
area=w*h
print(area)
a=area/50000
#py.keyDown('shift')
if a<b:
py.typewrite(['['])
elif a>b:
py.typewrite([']'])
#py.keyUp('shift')
b=a
continue
else:
cv2.waitKey(10)
cv2.destroyAllWindows()
break
elif predicted==8:
py.FAILSAFE = False
time.sleep(0.5)
py.hotkey('ctrl', 'alt', 'm')
#minimize
# webbrowser.open("https://www.youtube.com/watch?v=du7paNLQN9M")
# time.sleep(5)
if k == 27: # press ESC to exit
break
elif k == ord('b'): # press 'b' to capture the background
bgModel = cv2.createBackgroundSubtractorMOG2(0, bgSubThreshold)
isBgCaptured = 1
print( '!!!Background Captured!!!')
elif k == ord('r'): # press 'r' to reset the background
bgModel = None
triggerSwitch = False
isBgCaptured = 0
print ('!!!Reset BackGround!!!')
elif k == ord('n'):
triggerSwitch = True
print ('!!!Trigger On!!!')
camera.release()
cv2.destroyAllWindows()
# +
import glob
t=[]
y_test=[]
l=[]
y_train=[]
ma=0
labels=['A','B','G','L','Y']
def ellipse_variance(contour):
M=cv2.moments(contour)
cX=int(M['m10']/M['m00'])
cY=int(M['m01']/M['m00'])
v=np.zeros((1,2,1))
for cord in contour:
x=np.array([[[cord[0][0]-cX],[cord[0][1]-cY]]])
v=np.vstack((v,x))
# print(v)
# print(x)
# print(cord)
# print(cord[0][0])
# print(v.shape)
c_ellipse=np.zeros((2,2))
for i in v:
c_ellipse+=np.dot(i,np.transpose(i))
c_ellipse/=contour.shape[0]
# print(c_ellipse)
c_ellipse_inv=np.linalg.inv(c_ellipse)
d=np.zeros((1,1))
for i in v:
d=np.vstack((d,np.sqrt(np.dot(np.dot(np.transpose(i),c_ellipse_inv),i))))
d=d[1:]
mu=1/contour.shape[0]*np.sum(d)
sigma=np.sqrt(1/contour.shape[0]*np.sum(np.square(d-mu)))
return mu/sigma
count=0
#data=np.zeros((1,7))
for label in labels:
count+=1
path = "training_set/"+label+"/"
cou=0
for filename in os.listdir(path):
cou+=1
#print(filename,'1')
input_path = os.path.join(path, filename)
img=cv2.imread(input_path,0)
ret,thresh=cv2.threshold(img,10,255,cv2.THRESH_BINARY)
im2,contours,hierarchy=cv2.findContours(img,cv2.RETR_TREE,cv2.CHAIN_APPROX_SIMPLE)
# print(len(contours))
# for cnt in contours:
# a = cv2.contourArea(cnt)
# if a>ma:
# c=cnt;
req=max(contours,key=cv2.contourArea)
# print(req.shape)
#req=req[0]
#req=c
area=cv2.contourArea(req)
#eccentricity
dim_rect=cv2.minAreaRect(req)[1]
para1=dim_rect[1]/dim_rect[0]
#circularity
perimeter=cv2.arcLength(req,True)
para2=4*np.pi*area/(perimeter**2)
#rectangularity
para3=area/(dim_rect[1]*dim_rect[0])
#convexity
hull=cv2.convexHull(req)
para4=cv2.arcLength(hull,True)/perimeter
#solidity
para5=area/cv2.contourArea(hull)
#ellipse variance
para6=ellipse_variance(req)
#np.vstack((data,np.array([para1,para2,para3,para4,para5,para6,ord(label)])))
g=[para1,para2,para3,para4,para5,para6]
l.append(g)
y_train.append(count)
# -
# + active=""
#
# + active=""
#
# -
print(predicted)
firefox http://www.youtube.com
import webbrowser
webbrowser.open("https://www.youtube.com/watch?v=du7paNLQN9M")
| specialprediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
import numpy as np
import pysut
import pandas as pd
import json
import pickle
import os
V = np.array([[1,0,0],[0.5,1,0],[0,0,1]])
U = np.array([[0,0.1,0.4],[0,0,1],[0.5,0.5,0]])
g = V.sum(axis=0)
q = V.sum(axis=1)
print("V:")
print(V)
print("U:")
print(U)
print("g:")
print(g)
print("q:")
print(q)
# $A_\mathrm{ITC-cxc} = U \cdot \hat{g}^{-1} \cdot V' \cdot \hat{q}^{-1}$
U.dot(np.diag(1/g)).dot((V.T).dot(np.diag(1/q)))
SUT = pysut.SupplyUseTable(V=V, U=U)
SUT.build_mr_Gamma()
SUT.build_E_bar()
A, __, __, __, Z, __ = SUT.aac_agg(keep_size=False)
print(Z)
SUT.E_bar
SUT.Gamma
# +
V_A = np.array([[1,0,0],[0.6,0,0],[0,0,1]])
V_B = np.array([[1,0,0],[0,1,0],[0,0,1]])
U_A = np.array([[0,0,0.5],[0,0,0.5],[0.5,0,0]])
U_B = np.array([[0,0.3,0],[1,0,0],[0.5,0,0]])
V_AB = np.array([[0,0,0],[0,0,0],[0,0,0]])
V_BA = np.array([[0,0,0],[0,0,0],[0,0,0]])
U_AB = np.array([[0,0,0],[0,0,0],[0.5,0,0]])
U_BA = np.array([[0.1,0,0],[0,0,0],[0,0,0]])
# -
V_total = np.concatenate((np.concatenate((V_A,V_AB), axis=1),np.concatenate((V_BA,V_B),axis=1)),axis=0)
U_total = np.concatenate((np.concatenate((U_A,U_AB), axis=1),np.concatenate((U_BA,U_B),axis=1)),axis=0)
V_total
U_total
SUT_MR = pysut.SupplyUseTable(V = V_total, U=U_total, regions=2)
SUT_MR.build_mr_Gamma()
SUT_MR.build_E_bar()
SUT_MR.build_mr_Xi()
#A_MR,_,_,_,_,_,Z_MR,_ = SUT_MR.psc_agg()
print(SUT_MR.Xi)
SUT_MR.aac_agg(keep_size=False)
print("V")
print(V_total)
print("U")
print(U_total)
SUT_MR.Gamma
SUT_MR.E_bar
SupplyDiag_Eval = SUT_MR.supply_diag_check()
print(SupplyDiag_Eval)
#A_MR,_,_,_,_,_,Z_MR,_ =
SUT_MR.psc_agg()
# +
print("V")
print(V_total)
print("U")
print(U_total)
SUT_MR.btc()
# -
np.dot(SUT_MR.Xi,V_total-np.diag(np.diag(V_total)))
np.dot(SUT_MR.Xi,U_total)
# +
print("V")
print(V_total)
print("U")
print(U_total)
np.dot(SUT_MR.Xi,U_total)-np.dot(SUT_MR.Xi,V_total-np.diag(np.diag(V_total)))
# -
SUT_MR.psc_agg()[0]-np.dot(SUT_MR.Xi,U_total)-np.dot(SUT_MR.Xi,V_total-np.diag(np.diag(V_total)))
exio_v = pd.read_csv('/home/jakobs/Documents/IndEcol/BONSAI/data/exiobase_hsut_3317/MR_HSUP_2011_v3_3_17_bonsai.csv', header=[0,1,2,3], index_col=[0,1,2,3,4])
exio_u = pd.read_csv('/home/jakobs/Documents/IndEcol/BONSAI/data/exiobase_hsut_3317/MR_HUSE_2011_v3_3_17_bonsai.csv', header=[0,1,2,3], index_col=[0,1,2,3,4])
exio_u.values.shape
exio_v.values.shape
exio_u.loc[exio_u.index.get_level_values(2) == 'p40.2.b']
np.where(exio_u.loc[exio_u.index.get_level_values(2) == 'p40.2.b'].values >0)
(exio_v.loc[exio_v.index.get_level_values(1) == 'Biogas'].values >0)
exioSUT = pysut.SupplyUseTable(V=exio_v.values, U=exio_u.values)
calval = pd.read_csv('/home/jakobs/Documents/IndEcol/BONSAI/data/exiobase_hsut_3317/Calorific_values.csv', header=[4], index_col=[0,1,2,3])
calval.fillna(1, inplace=True)
aggregation_matrix = pd.read_csv('/home/jakobs/Documents/IndEcol/BONSAI/data/exiobase_hsut_3317/aggregation_matrix_exiobase.csv', header=[0,1,2], index_col=[0,1,2,3])
aggregation_matrix.ix[aggregation_matrix.iloc[:,109]==1,109]
# ### Below is a litte function to document the aggregation
# +
N_regs = 48
N_sec = 164
N_prod = 200
nr_of_products = []
testfile = '/home/jakobs/Desktop/testfile.csv'
with open(testfile, 'w') as f:
f.write('#Agrgegation report \n')
f.write('|'.join(['Country code','Industry name','Industry code 1','Industry code 2', 'Number of products to be aggregated', 'Product name', 'Product code 1','Product code 2', 'Aggregation Value', '\n']))
for c in range(N_regs):
for i in range(aggregation_matrix.shape[1]):
n_prods = aggregation_matrix.iloc[:,i].sum()
if n_prods > 1:
prod_inds = np.where(aggregation_matrix.iloc[:,i]==1)[0]
#print(prod_inds)
aggregated_prods = [aggregation_matrix.index[x] for x in prod_inds]
aggregation_values = new_aggregation_matrix[prod_inds+c*N_prod, i+c*N_sec]
#print(aggregation_values)
nr_of_products.append(n_prods)
#print(*aggregation_matrix.columns[i], n_prods)
for j,prod in enumerate(aggregated_prods):
#if j == 0:
# print(prod[1:])
f.write('|'.join([country_list[c],'|'.join(aggregation_matrix.columns[i]), '{}'.format(n_prods), '|'.join(prod[1:]), '{}'.format(aggregation_values[j]), '\n']))
#else:
# f.write('|'.join(['','','','','', '|'.join(prod[1:]), '{}'.format(aggregation_values[j]), '\n']))
df = pd.read_csv('/home/jakobs/Desktop/testfile.csv', delimiter='|', skiprows=1, index_col=[0,1])
df
# -
#aggregation_matrix[aggregation_matrix['Manufacture of gas;']>0]
aggregation_matrix.columns[109][0][:-1]
calval.iloc[141:146]
calval.loc[calval.index.get_level_values(3) == 'C_GASE']
Natural_gas_calval = calval.loc[calval.index.get_level_values(3) == 'C_GASE'].values[0] #natural gas production has the same caloric value for now but we keep the a specific value for now
print(Natural_gas_calval)
calval.iloc[141:146]/Natural_gas_calval
new_aggregation_matrix = np.zeros((48*200,48*164))
for i in range(48):
new_aggregation_matrix[i*200:i*200+200,i*164:i*164+164] = aggregation_matrix
new_aggregation_matrix[200,164]
for i in range(48):
#print(calval.iloc[141:146,i]/Natural_gas_calval[i])
new_aggregation_matrix[i*200+141:i*200+146,i*164+109] = calval.iloc[141:146,i]/Natural_gas_calval[i]
j=10
new_aggregation_matrix[141+j*200,109+164*j]
exio_vagg = new_aggregation_matrix.T.dot(exio_v.values)
j=1
print(exio_vagg[24+j*164,24+j*164])
exio_vagg.shape
exio_v.loc[exio_v.index.get_level_values(3)=='C_IRON','AU'].values
aggregation_matrix
exio_uagg = new_aggregation_matrix.T.dot(exio_u.values)
exio_SUTagg_Obj = pysut.SupplyUseTable(V=exio_vagg, U=exio_uagg, regions=48)
# +
iot_names = pd.read_csv('/home/jakobs/data/EXIOBASE/EXIOBASE_3.3.17_hsut_2011/Classifications_v_3_3_17_IOTnames.csv')
country_list = iot_names['Country code'].unique()
country_dic = {}
for i,c in enumerate(country_list):
country_dic[c] = i
products = iot_names[['Product name', 'Product code 1']]
products.drop_duplicates(inplace=True)
prod_dic = {}
for i,prod in enumerate(products['Product name'].values):
prod_dic[prod] = i
# -
products[products['Product name'].str.contains('Distribution')]
exio_vagg.shape
supply_diag_check_eval = exio_SUTagg_Obj.supply_diag_check()
supply_diag_check_eval.shape
prod_names = iot_names.values[supply_diag_check_eval[:,2]==1,:]
print(prod_names)
def create_market_and_product_names(prod_names, N_reg, Reg_list):
"""Create market names, and product names.
Input:
prod_names : array of exclusive byproducts (this should include at
at least one electricity byproduct e.g. electricity
from coal (as byproduct from heat from coal).
N_reg : number of regions in the MRIO system
Output:
excl_byproducts : array of exclusive byproducts (excl electricity)
market_names : array of market names corresponding to the excl byprods
grid_electricity: array of 'Electricity from grid' names for the different
regions. i.e. 1 entry for each electricity market
elec_markets : Like grid_electricity but now "Market for electricity"
All outputs have the format [['Region', 'Name', 'code 1', 'code 2', 'unit']]
where the unit only exists for the products not for the markets.
"""
#get a unique list of the product names to create global markets
unique_prod_indices = np.unique(prod_names[:,1], return_index=True)[-1]
excl_byproducts = prod_names[unique_prod_indices]
#print(market_names)
market_names = excl_byproducts.copy()
market_names[:,0] = 'GLO' #set the region to global
#now change the names of the products to Market for 'product', incl codes.
#also remove the electricity from the excl_byproducts
rm_index = []
for i,prod in enumerate(excl_byproducts):
market_names[i,1] = 'Market for ' + market_names[i,1]
market_names[i,2] = market_names[i,2].replace('p','m')
market_names[i,3] = market_names[i,3].replace('C_','M_')
#there are multiple electricity byproducts. they supplyt the same market
#so give them the same code/market name
if 'm40.11' in market_names[i,2]:
market_names[i,2] = 'm40.11'
market_names[i,1] = 'Market for electricity'
market_names[i,3] = 'M_ELEC'
if 'p40.11.' in prod[2]:
rm_index.append(i)
#remove electricity from excl byproducts
excl_byproducts = np.delete(excl_byproducts, np.array(rm_index), axis=0)
#drop the duplicate electricity markets
market_name_indices = np.unique(market_names[:,1], return_index=True)[-1]
market_names = market_names[market_name_indices]
#split the markets into exclusive byproduct markets and electricity markets
elec_markets = np.array([x for x in market_names if x[2]=='m40.11']*N_reg).reshape(N_reg,5)
elec_markets[:,0] = Reg_list
market_names = np.delete(market_names,
np.where(market_names[:,2]=='m40.11')[0], axis=0)
#create electricity market products ('Electricity from the grid')
grid_electricity = elec_markets.copy()
for i in range(len(elec_markets)):
grid_electricity[i,1] = 'Electricity from the grid'
grid_electricity[i,2] = grid_electricity[i,2].replace('m','p')
grid_electricity[i,3] = grid_electricity[i,3].replace('M_','C_',)
#drop units for markets:
elec_markets = elec_markets[:,:-1]
market_names = market_names[:,:-1]
return excl_byproducts, market_names, grid_electricity, elec_markets
create_market_and_product_names(prod_names, 48, country_list)
# +
market_name_indices = np.unique(prod_names[:,1], return_index=True)[-1]
market_names = prod_names[market_name_indices]
#print(market_names)
market_names[:,0] = 'GLO'
for i in range(len(market_names)):
market_names[i,1] = 'Market for ' + market_names[i,1]
market_names[i,2] = market_names[i,2].replace('p','m')
market_names[i,3] = market_names[i,3].replace('C_','M_')
if 'm40.11' in market_names[i,2]:
market_names[i,2] = 'm40.11'
market_names[i,1] = 'Market for electricity'
market_names[i,3] = 'M_ELEC'
market_name_indices = np.unique(market_names[:,1], return_index=True)[-1]
market_names = market_names[market_name_indices]
elec_markets = np.array([x for x in market_names if x[2]=='m40.11']*48).reshape(48,5)
elec_markets[:,0] = iot_names['Country code'].unique()
market_names = np.delete(market_names, np.where(market_names[:,2]=='m40.11')[0], axis=0)
#np.append(market_names, elec_markets, axis=0)
print(elec_markets)
market_names
# +
U_without_elec = exio_uagg.copy()
V_without_elec = exio_vagg.copy()
diag_dummy_indices = np.arange(V_without_elec.shape[0])
V_without_elec[diag_dummy_indices, diag_dummy_indices] = 0 #set to zero for principle production so we can sum off diagonal electricity
V_elecmarkets = np.zeros(len(elec_markets)) #These are the totals of the electricity used in a country, as this is the total that a national grid will provide. Format is (N_countries x 1), will be diagonalized in final V' table
U_elecmarkets = np.zeros((exio_uagg.shape[0],len(elec_markets))) #This defines the electricity mix in a national grid. This will be updated with entso data. Format (N_products*N_countries x N_countries)
elec_indices = iot_names['Product code 1'].str.contains('p40.11').values #boolean array with the electricity commodities
elec_martket_product_use = np.zeros((len(elec_markets), exio_uagg.shape[0])) #The input vector for grid electricity for the different activities, as they now draw from the grid instead of directly from producers. Format (N_countries, N_countries*N_products+N_countries)
elec_martket_product_supply = np.zeros((len(elec_markets), exio_uagg.shape[0]))
for i in range(len(elec_markets)): #loop over countries
U_elecmarkets[elec_indices,i] = exio_uagg[elec_indices,i*164:i*164+164].sum(axis=1)
V_elecmarkets[i] = U_elecmarkets[elec_indices,i].sum()
elec_martket_product_use[i,i*164:i*164+164] = exio_uagg[elec_indices,i*164:i*164+164].sum(axis=0)
elec_martket_product_supply[i,i*164:i*164+164] = V_without_elec[elec_indices,i*164:i*164+164].sum(axis=0)
V_without_elec[elec_indices,i*164:i*164+164] = 0
V_without_elec[diag_dummy_indices, diag_dummy_indices] = exio_vagg[diag_dummy_indices, diag_dummy_indices]
U_without_elec[elec_indices,:] = 0 #is the new partial Use table where electricity use from producers has been set to 0
# -
elec_martket_product_supply[1,96+15+28+164]
unique_prod_indices = np.unique(prod_names[:,1], return_index=True)[-1]
excl_byproducts = prod_names[unique_prod_indices]
rm_index = []
for i,prod in enumerate(excl_byproducts):
if 'p40.11.' in prod[2]:
rm_index.append(i)
excl_byproducts = np.delete(excl_byproducts, np.array(rm_index), axis=0)
print(market_names)
excl_byproducts
# +
v_market_excl_byproduct = np.zeros(len(excl_byproducts)) #create a vector for the market supply
u_market_excl_byproduct = np.zeros((U_without_elec.shape[0], len(excl_byproducts)))
excl_market_products_use = np.zeros((len(excl_byproducts),U_without_elec.shape[0]))
excl_market_products_supply = np.zeros((len(excl_byproducts),U_without_elec.shape[0]))
U_markets = U_without_elec.copy() #this now will become the final use table (without markets)
V_markets = V_without_elec.copy() #this now will become the final supply table without markets
for i,excl_prod in enumerate(excl_byproducts):
prod_ind = prod_dic[excl_prod[1]]
excl_prod_indices = np.where(iot_names.as_matrix()[:,2]==excl_byproducts[i,2])[0]
u_market_excl_byproduct[excl_prod_indices,i] = exio_vagg[excl_prod_indices,excl_prod_indices]
v_market_excl_byproduct[i] = u_market_excl_byproduct[:,i].sum()
excl_byprod_countries = prod_names[np.where(prod_names[:,2] == excl_prod[2]),0][0]
c_index = np.array([country_dic[x] for x in excl_byprod_countries])
excl_market_products_use[i,:] = exio_uagg[c_index*164+prod_ind,:].sum(axis=0) #every acticity that normally buys the product from a country where this is only a by product now needs to buy this from the market.
U_markets[c_index*164+prod_ind,:] = 0 #We then set the use from the particular countries to 0 as they only buy from the market.
excl_market_products_supply[i,:] = exio_vagg[c_index*164+prod_ind,:].sum(axis=0) #We now need to move the byproduction in these countries to the market. We can simpy sum over the columns as the production of the other countries in this country will always be 0 and won't affect the sum
V_markets[c_index*164+prod_ind,:] = 0 #as above for the use but now for the byproduct supply
# +
#here we assemble everything into one Use and one Supply table
exio_dim = exio_uagg.shape[0]
n_elecmarket = len(elec_markets)
n_excl_bp = len(excl_byproducts)
full_dimension =exio_dim + n_elecmarket + n_excl_bp
U_full = np.zeros((full_dimension,full_dimension))
V_full = np.zeros((full_dimension,full_dimension))
#first insert the main use and supply tables
U_full[:exio_dim,:exio_dim] = U_markets
V_full[:exio_dim,:exio_dim] = V_markets
#now insert_electricity markets
U_full[exio_dim:exio_dim+n_elecmarket,:exio_dim] = elec_martket_product_use
U_full[:exio_dim,exio_dim:exio_dim+n_elecmarket] = U_elecmarkets
V_full[exio_dim:exio_dim+n_elecmarket,:exio_dim] = elec_martket_product_supply
V_full[exio_dim:exio_dim+n_elecmarket,exio_dim:exio_dim+n_elecmarket] = np.diag(V_elecmarkets)
#now insert exclusive byproduct markets
U_full[exio_dim+n_elecmarket:,:exio_dim] = excl_market_products_use
U_full[:exio_dim,exio_dim+n_elecmarket:] = u_market_excl_byproduct
V_full[exio_dim+n_elecmarket:,:exio_dim] = excl_market_products_supply
V_full[exio_dim+n_elecmarket:,exio_dim+n_elecmarket:] = np.diag(v_market_excl_byproduct)
# -
print(elec_markets)
print(market_names)
#create market electricity product names
# $Z = U = $
def make_IOT(U_full, V_full):
Z_full = U_full - V_full + np.diag(np.diag(V_full))
x_dummy = np.diag(V_full).copy()
x_dummy[x_dummy == 0] = 1
A_full = Z_full/x_dummy
return Z_full, A_full
Z_full, A_full = make_IOT(U_full, V_full)
# * check for every non zero entry in entsoe if it has principle production exiobase
# * update information
# +
#Z_e_update, A_e_update = update_electricity_mix()
def update_electricity_mix(entso_u, entso_v, entso_act, entso_prod, U,V):
country_ind = country_dic[country_code]
return U_update, V_update
# -
bentso_prods = json.load(open('/home/jakobs/Documents/IndEcol/BONSAI/bentso-data/entsoe-beebee-2016/entsoe-products.json'))
bentso_acts = json.load(open('/home/jakobs/Documents/IndEcol/BONSAI/bentso-data/entsoe-beebee-2016/entsoe-activities.json'))
bentso_use = np.load('/home/jakobs/Documents/IndEcol/BONSAI/bentso-data/entsoe-beebee-2016/entsoe-use.npy')
bentso_supply = np.load('/home/jakobs/Documents/IndEcol/BONSAI/bentso-data/entsoe-beebee-2016/entsoe-supply.npy')
bentso_acts
print(32*12)
print(672/32.)
pd.DataFrame(columns=np.array(bentso_acts).T.tolist(), index=np.array(bentso_prods).T.tolist(), data=bentso_use)
set(country_concord['bentso_Country_code'].unique()).difference(set(europe_exio))
country_concord
prod_cor = pd.read_csv('/home/jakobs/Documents/IndEcol/BONSAI/Correspondence-tables/final_tables/tables/exiobase_to_bentso_activities.csv')
len(prod_cor['Bentso technology'].dropna())
# +
ACTIVITY_MAPPING = {
'Fossil Hard coal': 'Production of electricity by coal',
'Fossil Brown coal/Lignite': 'Production of electricity by coal',
'Fossil Gas': 'Production of electricity by gas',
'Fossil Coal-derived gas': 'Production of electricity by gas',
'Nuclear': 'Production of electricity by nuclear',
'Hydro Pumped Storage': 'Production of electricity by hydro',
'Hydro Run-of-river and poundage': 'Production of electricity by hydro',
'Hydro Water Reservoir': 'Production of electricity by hydro',
'Wind Offshore': 'Production of electricity by wind',
'Wind Onshore': 'Production of electricity by wind',
'Fossil Oil': 'Production of electricity by petroleum and other oil derivatives',
'Fossil Oil shale': 'Production of electricity by petroleum and other oil derivatives',
'Biomass': 'Production of electricity by biomass and waste',
'Fossil Peat': 'Production of electricity by coal',
'Waste': 'Production of electricity by biomass and waste',
'Solar': 'Production of electricity by solar thermal',
'Other renewable': 'Production of electricity by tide, wave, ocean',
'Geothermal': 'Production of electricity by Geothermal',
'Other': 'Production of electricity nec',
'Marine': 'Production of electricity nec',
'Grid': "Market for Electricity",
}
FLOW_MAPPING = {
'Fossil Hard coal': 'Electricity by coal',
'Fossil Brown coal/Lignite': 'Electricity by coal',
'Fossil Gas': 'Electricity by gas',
'Fossil Coal-derived gas': 'Electricity by gas',
'Nuclear': 'Electricity by nuclear',
'Hydro Pumped Storage': 'Electricity by hydro',
'Hydro Run-of-river and poundage': 'Electricity by hydro',
'Hydro Water Reservoir': 'Electricity by hydro',
'Wind Offshore': 'Electricity by wind',
'Wind Onshore': 'Electricity by wind',
'Fossil Oil': 'Electricity by petroleum and other oil derivatives',
'Fossil Oil shale': 'Electricity by petroleum and other oil derivatives',
'Biomass': 'Electricity by biomass and waste',
'Fossil Peat': 'Electricity by coal',
'Waste': 'Electricity by biomass and waste',
'Solar': 'Electricity by solar thermal',
'Other renewable': 'Electricity by tide; wave; ocean',
'Geothermal': 'Electricity by Geothermal',
'Other': 'Electricity nec',
'Marine': 'Electricity nec',
'Grid': "Electricity",
}
# -
np.unique([ACTIVITY_MAPPING[x] for x in ACTIVITY_MAPPING.keys()])
np.unique([x for x in ACTIVITY_MAPPING.keys()])
def my_function(x,y):
x[5] = 100
y[0:2] = 1000
return x,y
# +
a = np.arange(10)
b = a.copy()
c,d = my_function(a,b)
print(a)
print(b)
print(c)
print(d)
print(a==c)
print(b==d)
b[5] = 0
print(b)
print(a)
# -
exio_u.shape
1*3e5/60/60/24.
| notebooks/BONSAI system model development.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
plt.rcParams['figure.figsize'] = [10, 6]
# %matplotlib inline
# +
# loading 'tips' dataset
tips = sns.load_dataset('tips')
tips.shape
tips.head()
# -
tips.groupby(['sex', 'day']).size()
# Basic box plot
plt.boxplot(tips['tip'])
plt.show()
# setting outlier symbol, title, xlabel
plt.boxplot(tips['tip'], sym="bo")
plt.title('Box plot of tip')
plt.xticks([1], ['tip'])
plt.show()
# Horizontal Box plot with notched box & red color outliers
plt.boxplot(tips['tip'],
notch=1, # if 'True' then notched box plot
sym='rs', # symbol: red square
vert=0 # vertical : if 'False' then horizontal box plot
)
plt.show()
# Multiple box plots on one Axes
fig, ax = plt.subplots()
ax.boxplot([tips['total_bill'], tips['tip']], sym="b*")
plt.title('Multiple box plots of tips on one Axes')
plt.xticks([1, 2], ['total_bill', 'tip'])
plt.show()
| gaze_project/visualization_system/data_processing/IQR.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Datathon 2019
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
path = './data/Dataset'
df_scores = pd.read_csv(path + '/Scores.csv', index_col = 0)
df_scores.head()
df_pre_contest = pd.read_csv(path + '/Pre_Contest.csv', index_col = 0)
df_pre_contest.head()
df_post_contest = pd.read_csv(path + '/Post_Contest.csv', index_col = 0)
df_post_contest.head()
df_pre_contest.describe()
df_pre_contest.info()
df_pre_contest.sum()
df_pre_contest.mean()
fig = plt.figure(figsize = (15,20))
ax = fig.gca()
df_pre_contest.hist(ax = ax)
plt.savefig('histogram_2.png', bbox_inches='tight')
df_pre_contest.groupby(['Which language are you most comfortable coding in?']).count()[['Rate your problem solving ability on the scale of 1 to 5', 'Rate your coding proficiency on the scale of 1 to 5']]
df_pre_contest.groupby(['Which language are you most comfortable coding in?']).mean()[['Rate your problem solving ability on the scale of 1 to 5', 'Rate your coding proficiency on the scale of 1 to 5']]
df_pre_contest.groupby(['Years of coding experience']).count()[['Rate your problem solving ability on the scale of 1 to 5', 'Rate your coding proficiency on the scale of 1 to 5']]
df_pre_contest['Department'] = df_pre_contest['Department'].replace({'MCA':'M.Tech Computer Science / M.C.A'})
df_pre_contest.groupby(['Department']).count()
df_years_exp_group = df_pre_contest.groupby(['Years of coding experience']).mean()[['Rate your problem solving ability on the scale of 1 to 5', 'Rate your coding proficiency on the scale of 1 to 5']]
df_years_exp_group
df_years_exp_group.index = ['1-2 years', '2-3 years', '3-4 years', '1 year', '4 years']
df_years_exp_group = df_years_exp_group.sort_index()
fig = plt.figure(figsize = (20,15))
ax = fig.gca()
df_years_exp_group.plot(ax = ax)
plt.title('Years of Experience and Ratings, Proficiency')
plt.savefig('plot_1.png', bbox_inches='tight')
| datathon2019-target-2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import pandas as pd
import numpy as np
from sklearn import preprocessing
from sklearn.tree import DecisionTreeClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.svm import SVC
from sklearn.svm import LinearSVC
from sklearn.calibration import CalibratedClassifierCV
from sklearn.feature_selection import RFE
from sklearn.model_selection import GridSearchCV
from sklearn import metrics
from sklearn.model_selection import cross_val_score
import matplotlib.pylab as plt
# %matplotlib inline
plt.rcParams['figure.figsize'] = 10, 8
# -
## Load cleaned approved loan data
df = pd.read_csv('approved_loan_2015_clean.csv',low_memory=False)
p_optimal='l1'
df.head()
c=0.0820849986238988
df2 = df.copy()
df2 = df2.ix[:,1:]
df2.head()
# +
from sklearn.model_selection import train_test_split
# Split training(labeled) and test(unlabled)
#df_train = df_bi[df_chin_bi['cuisine_Chinese'] != 2]
#df_test = df_chin_bi[df_chin_bi['Loan_status'] == 2]
# reduce data volumn by randomly selecting instances
np.random.seed(99)
ind = np.random.randint(0, len(df2), 100000)
df_reduced = df2.ix[ind, :]
df_labeled = df_reduced[df_reduced['Target']!=2]
df_unlabeled = df_reduced[df_reduced['Target']==2]
X = df_labeled.drop('Target',axis=1)
Y = df_labeled['Target']
X_train_val, X_test, Y_train_val, Y_test = train_test_split(X, Y, test_size=0.2, random_state=42)
X_train, X_vali , Y_train , Y_vali = train_test_split(X_train_val, Y_train_val, test_size=0.25, random_state=42)
# -
X.shape
X_test.shape
X_train.shape
X_train_val.shape
lr = LogisticRegression(C=c, penalty=p_optimal, random_state=99)
lr.fit(X_train, Y_train)
cost_matrix = pd.DataFrame([[0.101, -0.647], [0, 0]], columns=['p', 'n'], index=['Y', 'N'])
print ("Cost matrix")
print (cost_matrix)
probabilities = lr.predict_proba(X_test)[:, 1]
# +
prediction = probabilities > 0.5
# Build and print a confusion matrix
confusion_matrix_large = pd.DataFrame(metrics.confusion_matrix(Y_test, prediction, labels=[1, 0]).T,
columns=['p', 'n'], index=['Y', 'N'])
print (confusion_matrix_large)
# +
#MANUAL ROC CALCULATION
n=[1,2,3,4,5,6,7,8,9]
TPR=[]
FPR=[]
for i in n:
prediction = probabilities > ((i*1.0)/10)
confusion_matrix_large = pd.DataFrame(metrics.confusion_matrix(Y_test, prediction, labels=[1, 0]).T,columns=['p', 'n'], index=['Y', 'N'])
TPR.append(1.0*confusion_matrix_large['p']['Y']/(1.0*(confusion_matrix_large['p']['Y']+confusion_matrix_large['p']['N'])))
FPR.append(1.0*confusion_matrix_large['n']['Y']/(1.0*(confusion_matrix_large['n']['Y']+confusion_matrix_large['n']['N'])))
# -
fpr, tpr, thresholds = metrics.roc_curve(Y_test, lr.predict_proba(X_test)[:,1])
thresholds
profits=[]
for i in thresholds:
prediction = probabilities > i
confusion_matrix_large = pd.DataFrame(metrics.confusion_matrix(Y_test, prediction, labels=[1, 0]).T,columns=['p', 'n'], index=['Y', 'N'])
profits.append((1.0*confusion_matrix_large['p']['Y'])*cost_matrix['p']['Y']+(1.0*confusion_matrix_large['n']['Y'])*cost_matrix['n']['Y'])
profits
plt.plot(thresholds,profits)
plt.xlabel("Threshold")
plt.ylabel("Profit")
plt.title("Profits")
profits
max(profits)
profits.index(max(profits))
profits(250)
profits[250]
Topt=thresholds[250]
Topt
prediction
probabilities
NewStatus = probabilities>Topt
NewStatus
a,b=np.unique(NewStatus,return_counts='True')
b
a
# +
confusion_matrix_large = pd.DataFrame(metrics.confusion_matrix(Y_test, NewStatus, labels=[1, 0]).T,columns=['p', 'n'], index=['Y', 'N'])
# -
confusion_matrix_large
# +
probabilities = lr.predict_proba(X)[:, 1]
profits2=[]
for i in thresholds:
prediction = probabilities > i
confusion_matrix_large = pd.DataFrame(metrics.confusion_matrix(Y, prediction, labels=[1, 0]).T,columns=['p', 'n'], index=['Y', 'N'])
profits2.append((1.0*confusion_matrix_large['p']['Y'])*cost_matrix['p']['Y']+(1.0*confusion_matrix_large['n']['Y'])*cost_matrix['n']['Y'])
# -
plt.plot(thresholds,profits2)
plt.xlabel("Threshold")
plt.ylabel("Profit")
plt.title("Profits")
len(probabilities)
len(prediction)
max(profits2)
profits.index(max(profits))
thresholds.item(250)
OptPred=probabilities>thresholds.item(250)
OPTMatrix = pd.DataFrame(metrics.confusion_matrix(Y, OptPred, labels=[1, 0]).T,columns=['p', 'n'], index=['Y', 'N'])
OPTMatrix
| Predictive_Modeling/Profit Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# # FeatureHasher and DictVectorizer Comparison
#
#
# Compares FeatureHasher and DictVectorizer by using both to vectorize
# text documents.
#
# The example demonstrates syntax and speed only; it doesn't actually do
# anything useful with the extracted vectors. See the example scripts
# {document_classification_20newsgroups,clustering}.py for actual learning
# on text documents.
#
# A discrepancy between the number of terms reported for DictVectorizer and
# for FeatureHasher is to be expected due to hash collisions.
#
#
# +
# Author: <NAME>
# License: BSD 3 clause
from collections import defaultdict
import re
import sys
from time import time
import numpy as np
from sklearn.datasets import fetch_20newsgroups
from sklearn.feature_extraction import DictVectorizer, FeatureHasher
def n_nonzero_columns(X):
"""Returns the number of non-zero columns in a CSR matrix X."""
return len(np.unique(X.nonzero()[1]))
def tokens(doc):
"""Extract tokens from doc.
This uses a simple regex to break strings into tokens. For a more
principled approach, see CountVectorizer or TfidfVectorizer.
"""
return (tok.lower() for tok in re.findall(r"\w+", doc))
def token_freqs(doc):
"""Extract a dict mapping tokens from doc to their frequencies."""
freq = defaultdict(int)
for tok in tokens(doc):
freq[tok] += 1
return freq
categories = [
'alt.atheism',
'comp.graphics',
'comp.sys.ibm.pc.hardware',
'misc.forsale',
'rec.autos',
'sci.space',
'talk.religion.misc',
]
# Uncomment the following line to use a larger set (11k+ documents)
# categories = None
print(__doc__)
print("Usage: %s [n_features_for_hashing]" % sys.argv[0])
print(" The default number of features is 2**18.")
print()
try:
n_features = int(sys.argv[1])
except IndexError:
n_features = 2 ** 18
except ValueError:
print("not a valid number of features: %r" % sys.argv[1])
sys.exit(1)
print("Loading 20 newsgroups training data")
raw_data = fetch_20newsgroups(subset='train', categories=categories).data
data_size_mb = sum(len(s.encode('utf-8')) for s in raw_data) / 1e6
print("%d documents - %0.3fMB" % (len(raw_data), data_size_mb))
print()
print("DictVectorizer")
t0 = time()
vectorizer = DictVectorizer()
vectorizer.fit_transform(token_freqs(d) for d in raw_data)
duration = time() - t0
print("done in %fs at %0.3fMB/s" % (duration, data_size_mb / duration))
print("Found %d unique terms" % len(vectorizer.get_feature_names()))
print()
print("FeatureHasher on frequency dicts")
t0 = time()
hasher = FeatureHasher(n_features=n_features)
X = hasher.transform(token_freqs(d) for d in raw_data)
duration = time() - t0
print("done in %fs at %0.3fMB/s" % (duration, data_size_mb / duration))
print("Found %d unique terms" % n_nonzero_columns(X))
print()
print("FeatureHasher on raw tokens")
t0 = time()
hasher = FeatureHasher(n_features=n_features, input_type="string")
X = hasher.transform(tokens(d) for d in raw_data)
duration = time() - t0
print("done in %fs at %0.3fMB/s" % (duration, data_size_mb / duration))
print("Found %d unique terms" % n_nonzero_columns(X))
| 01 Machine Learning/scikit_examples_jupyter/text/plot_hashing_vs_dict_vectorizer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data Viz
#
# - toc:true
# - branch: master
# - badges: true
# - comments: true
# - author: <NAME>
# - categories: [data viz, posters, data science, Thomas Simm]
# 
# Some data visualizations from my work in metallurgy. Most of my work was line plots so tried to keep these to a minimum here.
#
# <h1 id="#EBSD"> EBSD </h1>
# Electron backscatter diffraction (EBSD) is a scanning electron microscope–based microstructural-crystallographic characterization technique commonly used in the study of crystalline or polycrystalline materials.[1][2] The technique can provide information about the structure,[3] crystal orientation ,[3]phase,[3] or strain[4] in the material.
#
#
# <i>[1] <NAME>; <NAME> (2000). Introduction to texture analysis: macrotexture, microtexture and orientation mapping (Digital printing 2003 ed.). Boca Raton: CRC Press. ISBN 978-9056992248.</i>
#
# <i>[2] <NAME>.; <NAME>.; <NAME>.; <NAME>. (2000). Electron backscatter diffraction in materials science. New York: Kluwer Academic.</i>
#
# <i>[3] Electron backscatter diffraction in materials science (2nd ed.). Springer Science+Business Media. 2009. p. 1. ISBN 978-0-387-88135-5.</i>
#
# <i>[4] <NAME>.; <NAME>; <NAME>. (2011). "A review of strain analysis using electron backscatter diffraction". Microscopy and Microanalysis. 17. 17 (3): 316–329. Bibcode:2011MiMic..17..316W. doi:10.1017/S1431927611000055. PMID 21418731.</i>
#
# From:
# https://en.wikipedia.org/wiki/Electron_backscatter_diffraction
# 
#
#
# A matenistic steel showing prior austenite grains. The top image has undergone an additional rolling regime resulting in smaller grain size.
# This is your classic EBSD orientation map with a twist in that the orientations are predicted. The colours represent different orientations given by the legend on the left- they are a vector [phi1,Phi,phi2] (MTEX) with each spatial point having it’s own vector. The colours are reconstructed using ARPGE to give a prediction on what the grains would be prior to cooling, based on the rotation matrix between adjacent elements. There are two problems here, defining the orientation relationships (or rotations) between neighbours and reconstructing grains. In the next case we can ignore the first.
#
# <a href='https://www.mdpi.com/1996-1944/10/7/730'> **Paper**: The Influence of Lath, Block and Prior Austenite Grain (PAG) Size on the Tensile, Creep and Fatigue Properties of Novel Maraging Steel </a>
#
#
# 
# An austenitic stainless steel that transforms to martensite under load.
# This is similar to the above example, but easier to solve as we only need to look at the rotation across one boundary at a time. In the top figure the coloured regions represent a different phase (martensite) with austenite grains and the particular colour what the orientation relationship (OR) is. The lines show the slip systems and the maximum Schmid factor. Basically, some OR are preferred, the direction of the martensite relates to the particular OR and the Schmid factors.
#
# <a href='https://www.sciencedirect.com/science/article/abs/pii/S026412751631231X?via%3Dihub'> **Paper**: In situ observation of strain and phase transformation in plastically deformed 301 austenitic stainless steel </a>
# 
# The change in various EBSD maps of an austenitic stainless steel (see above) before and after being pulled to 10% strain. See also the image below. Each set of maps (on horizontal axis and below) give a different measure of plastic deformation. But if we look closely we can see some areas with high values of one parameter can have low values of another. Even if we average over a grain it can be difficult to predict behaviour. These observations illustrate aspects of plastic deformation such as the chaotic nature and the none unique definition of plastic deformation.
#
# <a href='https://www.sciencedirect.com/science/article/abs/pii/S1044580319328256'> **Paper**: The τ-plot, a multicomponent 1-D pole figure plot, to quantify the heterogeneity of plastic deformation </a>
# 
# This is a localised strain map of the sample shown above but a slightly bigger region. This is produced by digital image correlation (DIC) by comparing two surface images. The data is then combined with the EBSD data allowing us to visualise the grain boundaries (black lines).
# <font size=6> So how do we take account of orientation differences in grains (they matter) AND the chaotic nature of deformation and differences in parameters? </font>
# <br>
#
# <font size=4 color=blue> We do some averageing </font>
# 
# This is a classic plot in metallurgy called the inverse pole figure (IPF) plot. Simply put each point on the triangle represents a different group of orientations. We are averageing based on an orientation criteria. In the top figures are two models that relate to plastic deformation (Schmid factor left and Taylor model right) plotted on to this IPF plot. The bottom two figures represent experimental data of a ‘Damage parameter’ (number of un-indexed points on boundary) of grains after creep deformation of an austenitic stainless steel. Clearly (hopefully), we can see a transition from type with increasing stress and overall plastic deformation. {Some adjustments could be done on the algorithm to create the plots but the main points remain}
# 
#
#
# In a similar manner we can average details of a sample based on orientation in a different manner, as shown here. The reason for this averaging is so we can combine EBSD with powder diffraction (X-ray and neutron). Powder diffraction provides useful insights on some parameters connected to EBSD such as texture, plastic deformation and phases. Furthermore, the information can be from larger volumes of the sample (and not just the surface) than EBSD. However, the way it is measured means we have to modify how we combine the data.
# In the figure crystal plasticity models are compared with experimental data for EBSD and powder diffraction on the same axis.
#
# <a href='https://www.sciencedirect.com/science/article/abs/pii/S1044580319328256'> **Paper**: The τ-plot, a multicomponent 1-D pole figure plot, to quantify the heterogeneity of plastic deformation </a>
#
# <h1 id='APT'>APT</h1>
# The atom probe was introduced at the 14th Field Emission Symposium in 1967 by <NAME> and <NAME>. It combined a field ion microscope with a mass spectrometer having a single particle detection capability and, for the first time, an instrument could “… determine the nature of one single atom seen on a metal surface and selected from neighboring atoms at the discretion of the observer”.[1]
#
# https://en.wikipedia.org/wiki/Atom_probe
# 
# The image is an APT tip (a 3D cylinder type shape) showing iso-surfaces for two different elements (i.e. inside these surfaces the composition of an element {Ni and Mo here} is higher than a set value). This image is typical of APT analysis and produced from a designated software package.
# 
# This transforms data similar to shown above, first the APT data was cut into slices- this makes it easier to visualize the density and details and compares better with other techniques such as TEM. Secondly, I adjusted colours and contrast to improve the visual feel.
# 
# The above maps are pretty but very qualitative. Some extra details can be found by extracting details from the data as shown here.
# <h1 id='posters'> Posters </h1>
# 
# 
# 
# 
| _notebooks/2021-10-24-Data-Viz.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Errors, or bugs, in your software
#
# Today we'll cover dealing with errors in your Python code, an important aspect of writing software.
#
# #### What is a software bug?
#
# According to [Wikipedia](https://en.wikipedia.org/wiki/Software_bug) (accessed 16 Oct 2018), a software bug is an error, flaw, failure, or fault in a computer program or system that causes it to produce an incorrect or unexpected result, or behave in unintended ways.
#
# #### Where did the terminology come from?
#
# Engineers have used the term well before electronic computers and software. Sometimes Thomas Edison is credited with the first recorded use of bug in that fashion. [[Wikipedia](https://en.wikipedia.org/wiki/Software_bug#Etymology)]
# #### If incorrect code is never executed, is it a bug?
#
# This is the software equivalent to "If a tree falls and no one hears it, does it make a sound?".
# ## Three classes of bugs
#
# Let's discuss three major types of bugs in your code, from easiest to most difficult to diagnose:
#
# 1. **Syntax errors:** Errors where the code is not written in a valid way. (Generally easiest to fix.)
# 1. **Runtime errors:** Errors where code is syntactically valid, but fails to execute. Often throwing exceptions here. (Sometimes easy to fix, harder when in other's code.)
# 1. **Semantic errors:** Errors where code is syntactically valid, but contain errors in logic. (Can be difficult to fix.)
import numpy as np
# ### Syntax errors
print ("This should only work in Python 2.x, not 3.x used in this class.")
x = 1; y = 2
b = x == y # Boolean variable that is true when x & y have the same value
b = 1 = 2
# ### Runtime errors
# invalid operation
try:
a = 0
5/a # Division by zero
# invalid operation
input = '40'
input/11 # Incompatiable types for the operation
# ### Semantic errors
#
# Say we're trying to confirm that a trigonometric identity holds. Let's use the basic relationship between sine and cosine, given by the Pythagorean identity"
#
# $$
# \sin^2 \theta + \cos^2 \theta = 1
# $$
#
# We can write a function to check this:
# +
import math
'''Checks that Pythagorean identity holds for one input, theta'''
def check_pythagorean_identity(theta):
return math.sin(theta)**2 + math.cos(theta) == 1
# -
check_pythagorean_identity(0)
# Is our code correct?
# ## How to find and resolve bugs?
#
# Debugging has the following steps:
#
# 1. **Detection** of an exception or invalid results.
# 2. **Isolation** of where the program causes the error. This is often the most difficult step.
# 3. **Resolution** of how to change the code to eliminate the error. Mostly, it's not too bad, but sometimes this can cause major revisions in codes.
#
# ### Detection of Bugs
#
# The detection of bugs is too often done by chance. While running your Python code, you encounter unexpected functionality, exceptions, or syntax errors. While we'll focus on this in today's lecture, you should never leave this up to chance in the future.
#
# Software testing practices allow for thoughtful detection of bugs in software. We'll discuss more in the lecture on testing.
# ### Isolation of Bugs
#
# There are three main methods commonly used for bug isolation:
# 1. The "thought" method. Think about how your code is structured and so what part of your could would most likely lead to the exception or invalid result.
# 2. Inserting ``print`` statements (or other logging techniques)
# 3. Using a line-by-line debugger like ``pdb``.
#
# Typically, all three are used in combination, often repeatedly.
# ### Using `print` statements
#
# Say we're trying to compute the **entropy** of a set of probabilities. The
# form of the equation is
#
# $$
# H = -\sum_i p_i \log(p_i)
# $$
#
# We can write the function like this:
import numpy as np
def entropy(p):
"""
arg p: list of float
"""
items = p * np.log(p)
return -np.sum(items)
entropy([0.5, 0.5])
# Next steps:
# - Other inputs
# - Determine reason for errors by looking at details of codes
# ### Using Python's debugger, `pdb`
#
# Python comes with a built-in debugger called [pdb](http://docs.python.org/2/library/pdb.html). It allows you to step line-by-line through a computation and examine what's happening at each step. Note that this should probably be your last resort in tracing down a bug. I've probably used it a dozen times or so in five years of coding. But it can be a useful tool to have in your toolbelt.
#
# You can use the debugger by inserting the line
# ``` python
# import pdb; pdb.set_trace()
# ```
# within your script. To leave the debugger, type "exit()". To see the commands you can use, type "help".
#
# Let's try this out:
def entropy(p):
items = p * np.log(p)
import pdb; pdb.set_trace()
return -np.sum(items)
# This can be a more convenient way to debug programs and step through the actual execution.
p = [.1, -.2, .3]
entropy(p)
| week_4/Debugging.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Reading and Writing to a File
#
# Sometimes you may want to read and write data from and to a file, for example like a .csv or .txt file.
#
# While, we'll learn some fancier ways to do this later on, it can also be useful to know how to do it using base python (i.e. not having to install/import fancy packages).
#
# In this short notebook we'll introduce `file` objects in python, if you're interested, here's a link to the python documentation on `file` objects, <a href="https://docs.python.org/3/tutorial/inputoutput.html#reading-and-writing-files">https://docs.python.org/3/tutorial/inputoutput.html#reading-and-writing-files</a>.
#
#
# #### A Note
#
# Before moving forward I want to remind you to be mindful of the order you run your code chunks in inside of `jupyter notebooks`. As we'll see in the rest of the notebook, some code needs to be run before/after other code to work properly.
# ## `file` Objects
#
# A `file` object in python is a way for you to read and write a file on your computer. You can either read a file, or write a file all from the comfort of your `jupyter notebook`!
#
# ### Writing a `file` Object
#
# We'll start by seeing how to write to a file object.
## we'll store the object in the variable file
## we open a file by calling open(file_name, open_method)
## for example here we call open("new_file.txt", "w+")
file = open("new_file.txt", "w+")
# Go to your repository. After running that code you should see the file, `new_file.txt`, show up. Why? When we call `open("new_file.txt, "w+")` it tells our computer to open the file, `new_file.txt`, and the `w+` tells python that we want to write on it. Specifically the `+` tells python that if this file doesn't exist we want to create it.
#
# If we left off the `+` we would have received a file not found error, but now that we've created the file we can open it whenever we want. Let's try below.
## first we must CLOSE the file
## this is an important step anytime
## you're done with a file, if you neglect
## to close the file your compute will just keep
## all of the files open until you close the
## jupyter notebook
file.close()
## now we'll reopen the file, but without the "+"
file = open("new_file.txt", "w")
# +
## once a file is open in "w"rite mode
## we can write to it with file.write(string)
file.write("Now my file has a line in it.\n")
## the \n at the end of that line tells python to
## make a "newline" in the file
## let's write a second line
file.write("This is the second line in the file.\n")
## I'm done with the file for now,
## let's close it.
file.close()
# -
# Before moving on to the next code chunk open up the file and check to see if it wrote what we told it to.
# +
## You code
## open the file in write mode
## write the line "A third line, might be fine"
## close the file when you're done
file = open("new_file.txt","w")
file.write("A third line, might be fine\n")
file.close()
# -
# Now open your file again. What happened?
#
# The `w` command tells your computer to overwrite any existing data that was there.
#
# If we want to append to our file after it has been created we must open with `a`, which stands for "a"ppend.
# +
## opening the file with "a" instead of "w"
## tells python we want to append lines to the
## file instead of overwrite the existing content
file = open("new_file.txt", "a")
file.write("A new line, we won't overwrite the old lines this time!")
file.close()
# -
# Check it one more time! Did I lie to you?
# +
## You code
## Create a .csv file called "my_first_data.csv"
## write a line of column names
## "x","y","z"
## don't close the file yet
## Hint: don't forget the \n!
file = open("my_first_data.csv","w+")
# column names
file.write("x,y,z\n")
# +
## You code
## use a loop to write the following corresponding data
## in the correct order,
## you can close it when you're done
## also separate the x, y and z values with commas
x = [1,2,3,4]
y = [2,4,6,8]
z = [1,8,27,64]
## code here
## hint you'll have to cast the contents of x, y and z
## as strs before writing them to file
## don't forget the \n!
for i in range(len(x)):
file.write(str(x[i]) + "," + str(y[i]) + "," + str(z[i]) + "\n")
file.close()
# -
# Open your file and compare it with `check_data_file.csv` to make sure it matches.
# ### Reading a `file` Object
#
# Now let's suppose that we have a file that contains data you would like to read in. Instead of `open(file_name,"w")` or `open(file_name,"a")`, you write `open(file_name,"r")`. You can then read the content with `.read()`. Let's see.
# +
## open the file to read it with "r"
file = open("new_file.txt", "r")
print(file.read())
# +
## You code
## try to reread the file's contents using .read()
print(file.read())
# -
# What happened?
#
# When we called `read()` on our file object the cursor of the object went through all the text and returned it to us, but this process left the cursor at the end of the file's text. Here we can think of the cursor as our own eyeballs. Once you've read through all of the text on a page, your eyes will be looking at the end of the page.
#
# In order to call `read()` again and return all the text we'll need to return our cursor back to the begining of the document (i.e. point our eyes back at the top of the page). This is done with a `seek()` call.
## Go back to the 0th item in the string
## aka the beginning of the file's contents
file.seek(0)
# +
## You code
## let's try this again store the output of file.read()
## in a variable called file_text
file_text = file.read()
# +
## You code
## print file_text with a print statement to check it worked
print(file_text)
# -
## close the file
file.close()
## You code
## open your file "my_first_data.csv" in "r"ead mode
file = open("my_first_data.csv", "r")
## You code
## see what the readlines() command does
file.readlines()
## close the file
file.close()
# That's all we'll need to know about reading and writing files using base python for the boot camp, I hope you enjoyed it!
# This notebook was written for the Erdős Institute Cőde Data Science Boot Camp by <NAME>, Ph. D., 2021.
#
# Redistribution of the material contained in this repository is conditional on acknowledgement of <NAME>, Ph.D.'s original authorship and sponsorship of the Erdős Institute as subject to the license (see License.md)
| python prep/4. Reading and Writing to a File - Complete.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import numpy as np
from sklearn.svm import SVC
from sklearn.metrics import accuracy_score
from sklearn.metrics import f1_score
from mlxtend.plotting import plot_confusion_matrix
import matplotlib.pyplot as plt
from sklearn.metrics import confusion_matrix
# +
#==============================================================
#======= Training and Testing Set declaration =================
#==============================================================
filename = '2021_Valence_Arousal_Class_emo.csv'
data = pd.read_csv(filename, header = None)
print(data.shape)
X = data.iloc[:,range(2,98)]
val_y = data.iloc[:,-2]
aro_y =data.iloc[:,-1]
# +
from sklearn.model_selection import StratifiedKFold
kf = StratifiedKFold(n_splits=10, shuffle=True)
i = 1
scr = []
for train, test in kf.split(X,val_y):
# print("%s %s" % (train, test))
# X_train, X_test, y_train, y_test = X[train], X[test], y[train], y[test]
#==============================================================
#============= Arousal Emotion train test======================
#==============================================================
aX_train, aX_test, ay_train, ay_test = X.iloc[train,:], X.iloc[test,:], val_y[train], val_y[test]
#==============================================================
#============= Arousal Emotion train test======================
#==============================================================
vX_train, vX_test, vy_train, vy_test = X.iloc[train,:], X.iloc[test,:], aro_y[train], aro_y[test]
#===========================================================================
#========================SVM Model Declaration for Valence =================
#===========================================================================
clf_val = SVC(gamma='auto')
#==============================================================
#==================Valence classification======================
#==============================================================
clf_val.fit(vX_train, vy_train) #SVM model train
vy_prdt = clf_val.predict(vX_test) #valence class prediction
#valence Accuracy
vAcc = accuracy_score(vy_test, vy_prdt)
vf1 = f1_score(vy_test, vy_prdt)
print('Fold:',i)
print('SVM Valence classification Accuracy :',vAcc)
print('SVM Valence classification F1 Score:',vf1)
# class_names = ['low valence','high valence']
# ## Plot Confusion matric Valence
# ## ================================
# fig1, ax1 = plot_confusion_matrix(conf_mat=cm_val, show_absolute=True,
# show_normed=True,
# colorbar=True,
# class_names=class_names)
# plt.figure(1)
# plt.show()
#==============================================================
#==================Arousal classification======================
#==============================================================
clf_aro = SVC(gamma='auto')
clf_aro.fit(aX_train, ay_train) #SVM model train
ay_prdt = clf_aro.predict(aX_test) #arousal class prediction
#arousal Accuracy
aAcc = accuracy_score(ay_test, ay_prdt)
af1 = f1_score(ay_test, ay_prdt)
print('SVM Arousal classification Accuracy :',aAcc)
print('SVM Arousal classification F1 Score:',af1)
# class_names = ['low arousal','high arousal']
# ## Plot Confusion matric Valence
# ## ================================
# fig1, ax1 = plot_confusion_matrix(conf_mat=cm_aro, show_absolute=True,
# show_normed=True,
# colorbar=True,
# class_names=class_names)
# plt.figure(1)
# plt.show()
print('-------------------------------------------------------------')
scr.append([i,vAcc,vf1, aAcc,af1])
i = i+1
flname = '09_JAN_2021_All_person'+'_SVM_results_10_Fold.csv'
np.savetxt(flname,scr, delimiter= ',')
| SVM-emotion-Deap-10Fold-_09-JAN-2021-Valence_Arousal_Class.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PySpark - Local
# language: python
# name: pyspark-local
# ---
sc
# # RDD API Examples
# ## Word Count
# In this example, we use a few transformations to build a dataset of (String, Int) pairs called counts and then save it to a file.
# ```
# sc.textFile(name, minPartitions=None, use_unicode=True)
# Read a text file from HDFS, a local file system (available on all nodes), or any Hadoop-supported file system URI, and return it as an RDD of Strings.
# ```
#
# +
import os
#text_file = sc.textFile(os.getcwd()+"/../datasets/quijote.txt")
# To avoid copying a local file to all workers
lines = []
with open('../datasets/quijote.txt') as my_file:
for line in my_file:
lines.append(line)
text_file = sc.parallelize(lines)
counts = text_file.flatMap(lambda line: line.split(" ")) \
.map(lambda word: (word, 1)) \
.reduceByKey(lambda a, b: a + b)
counts = counts.sortBy(lambda a: a[1], ascending=False)
#NOTE: sortBy is not as efficient as sortByKey since it involves keying by the values,
#sorting by the keys, and then grabbing the values
counts.take(50)
#counts.saveAsTextFile(os.path.join("/notebooks/","quixote-counts.txt"))
# -
# ## Pi Estimation
#
# Spark can also be used for compute-intensive tasks. This code estimates pi by "throwing darts" at a circle. We pick random points in the unit square ((0, 0) to (1,1)) and see how many fall in the unit circle. The fraction should be pi / 4, so we use this to get our estimate.
# +
import random
NUM_SAMPLES=12000000
def inside(p):
x, y = random.random(), random.random()
return x*x + y*y < 1
count = sc.parallelize(range(0, NUM_SAMPLES)) \
.filter(inside).count()
print ("Pi is roughly {}".format(4.0 * count / NUM_SAMPLES))
# -
# # DataFrame API Examples
# ### Testing Conversion to/from Pandas with arrow
# +
# Enable Arrow-based columnar data transfers
spark.conf.set("spark.sql.execution.arrow.enabled", "true")
# Generate a Pandas DataFrame
pdf = pd.DataFrame(np.random.rand(100, 3))
# Create a Spark DataFrame from a Pandas DataFrame using Arrow
df = spark.createDataFrame(pdf)
# Convert the Spark DataFrame back to a Pandas DataFrame using Arrow
result_pdf = df.select("*").toPandas()
# -
# In this example, we count al quijote lines mentioning Dulcinea.
# +
from pyspark.sql import SparkSession, Row
from pyspark.sql.functions import col
# Creates a DataFrame having a single column named "line"
df = text_file.map(lambda r: Row(r)).toDF(["line"])
dulcinea_lines = df.filter(col("line").like("%Dulcinea%"))
# Counts all the Dulcinea lines
print("There are {} lines with 'Dulcinea'".format(dulcinea_lines.count()))
# Counts lines mentioning Dulcinea and Quijote
print("There are {} lines with 'Dulcinea' and 'Quijote'".format(
dulcinea_lines.filter(col("line").like("%Quijote%")).count()))
# Fetches the lines as an array of strings
dulcinea_lines.filter(col("line").like("%Quijote%")).collect()
# -
# ### Exploring the superheroes dataset
# +
from pyspark.sql.types import *
# To avoid copying a local file to all workers we create pandas dataframe at driver and convert to spark dataframe
# To Enable Arrow-based columnar data transfers
spark.conf.set("spark.sql.execution.arrow.enabled", "true")
superhero_pdf = pd.read_csv("../datasets/superheroes_info.csv",index_col='Index')
# We explicitly set schema to avoid problems with mapping pandas NaN Strings to SparkDataframe
# If not set, Spark will try to convert NaN to DoubleType wiht error -> Can not merge type <class 'pyspark.sql.types.StringType'> and <class 'pyspark.sql.types.DoubleType'>
mapping = {'object': StringType, 'float64': FloatType}
superhero_df = spark.createDataFrame(superhero_pdf, schema= StructType( [StructField(name, mapping[dtype.name]()) for name,dtype in superhero_pdf.dtypes.iteritems() ]))
superhero_df.show(10)
# -
from pyspark.sql.functions import isnan, when, count, col
df=superhero_df
publisher_df = superhero_df.groupby("Publisher").count().show()
# ### Spark SQL Example
superhero_df.createOrReplaceTempView("superhero_table")
spark.sql("select Name,Gender,Status from superhero_table").show()
| example-pnda-apps/jupyter-spark-integration-0.1.0/jupyter/spark/python/Introduction to PySpark.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
bpmn = open("fleamarket-submission.bpmn").read()
# +
from IPython.display import display
import json
data = json.dumps({
"style": {
"height": "400px"
}
})
# + tags=[]
display_id = "my-display-id"
handle = display({"application/bpmn+xml": bpmn, "application/bpmn+json": data}, raw=True, display_id=display_id)
# +
from IPython.display import update_display
tasks = [
"robot_search",
"robot_count",
"robot_save_item",
"robot_review",
"robot_items_excel",
"robot_create_json"
]
data = json.dumps({
"style": {
"height": "400px"
},
"colors": {
task: {
"stroke": "#ffffff",
"fill": "#000000",
} for task in tasks
}
})
update_display({"application/bpmn+xml": bpmn, "application/bpmn+json": data}, raw=True, display_id=display_id)
| examples/example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import statistics
# +
er_500_50_0012 = pd.read_csv('proc_er_500_50_0012.csv')
del er_500_50_0012['Unnamed: 0']
# -
er_500_50_0012
# +
er_500_50_0012_rgg_rgg_data = er_500_50_0012.copy()
er_500_50_0012_rgg_rand_data = er_500_50_0012.copy()
er_500_50_0012_rand_rgg_data = er_500_50_0012.copy()
er_500_50_0012_rand_rand_data = er_500_50_0012.copy()
rgg_rgg_drop_list = []
rgg_rand_drop_list = []
rand_rgg_drop_list = []
rand_rand_drop_list = []
for i in range(400):
if i % 4 == 0:
rgg_rand_drop_list.append(i)
rand_rgg_drop_list.append(i)
rand_rand_drop_list.append(i)
elif i % 4 == 1:
rgg_rgg_drop_list.append(i)
rand_rgg_drop_list.append(i)
rand_rand_drop_list.append(i)
elif i % 4 == 2:
rgg_rgg_drop_list.append(i)
rgg_rand_drop_list.append(i)
rand_rand_drop_list.append(i)
elif i % 4 == 3:
rgg_rgg_drop_list.append(i)
rgg_rand_drop_list.append(i)
rand_rgg_drop_list.append(i)
rgg_rgg_data = rgg_rgg_data.drop(rgg_rgg_drop_list)
rgg_rand_data = rgg_rand_data.drop(rgg_rand_drop_list)
rand_rgg_data = rand_rgg_data.drop(rand_rgg_drop_list)
rand_rand_data = rand_rand_data.drop(rand_rand_drop_list)
rgg_rgg_data = rgg_rgg_data.reset_index(drop=True)
rgg_rand_data = rgg_rand_data.reset_index(drop=True)
rand_rgg_data = rand_rgg_data.reset_index(drop=True)
rand_rand_data = rand_rand_data.reset_index(drop=True)
# -
# ## ---------------------------------------------------------------------------------------------------
plt.plot(rgg_rgg_2_dict['intra_thres'], rgg_rgg_2_dict['alive_nodes'])
plt.plot(rgg_rgg_2_dict['intra_thres'], rgg_rand_2_dict['alive_nodes'])
plt.plot(rgg_rgg_2_dict['intra_thres'], rand_rgg_2_dict['alive_nodes'])
plt.plot(rgg_rgg_2_dict['intra_thres'], rand_rand_2_dict['alive_nodes'])
plt.legend(['RGG-RGG', 'RGG-Rand', 'Rand-RGG', 'Rand-Rand'])
plt.title('Mean Alive nodes')
plt.show()
# +
p = 0.8
plt.plot([p * i for i in rgg_rgg_2_dict['init_mean_deg']], rgg_rgg_2_dict['alive_nodes'])
plt.plot([p * i for i in rgg_rgg_2_dict['init_mean_deg']], rgg_rand_2_dict['alive_nodes'])
plt.plot([p * i for i in rgg_rgg_2_dict['init_mean_deg']], rand_rgg_2_dict['alive_nodes'])
plt.plot([p * i for i in rgg_rgg_2_dict['init_mean_deg']], rand_rand_2_dict['alive_nodes'])
plt.legend(['RGG-RGG', 'RGG-Rand', 'Rand-RGG', 'Rand-Rand'])
plt.title('Mean Alive nodes')
plt.show()
# +
p = 0.9
plt.plot(rgg_rgg_2_dict['init_mean_deg'], rgg_rgg_2_dict['alive_nodes'])
plt.plot(rgg_rgg_2_dict['init_mean_deg'], rgg_rand_2_dict['alive_nodes'])
plt.plot(rgg_rgg_2_dict['init_mean_deg'], rand_rgg_2_dict['alive_nodes'])
plt.plot(rgg_rgg_2_dict['init_mean_deg'], rand_rand_2_dict['alive_nodes'])
plt.legend(['RGG-RGG', 'RGG-Rand', 'Rand-RGG', 'Rand-Rand'])
plt.title('Mean Alive nodes')
plt.show()
# +
step_nums = []
step_nums.append(statistics.mean(rgg_rgg_data['cas_steps'].values.tolist()))
step_nums.append(statistics.mean(rgg_rand_data['cas_steps'].values.tolist()))
step_nums.append(statistics.mean(rand_rgg_data['cas_steps'].values.tolist()))
step_nums.append(statistics.mean(rand_rand_data['cas_steps'].values.tolist()))
index = np.arange(4)
graph_types = ['RGG-RGG', 'RGG-Rand', 'Rand-RGG', 'Rand-Rand']
plt.bar(index, step_nums, width=0.3, color='gray')
plt.xticks(index, graph_types)
plt.title('Number of steps')
plt.savefig('The number of steps.png')
plt.show()
# -
rgg_rgg_isol = []
rgg_rgg_unsupp = []
rgg_rand_isol = []
rgg_rand_unsupp = []
rand_rgg_isol = []
rand_rgg_unsupp = []
rand_rand_isol = []
rand_rand_unsupp =[]
# +
index = 1
for col_name in rgg_rgg_data:
if col_name == ('step%d_isol' % index):
rgg_rgg_isol.append(statistics.mean(rgg_rgg_data[col_name].values.tolist()))
if col_name == ('step%d_unsupp' % index):
rgg_rgg_unsupp.append(statistics.mean(rgg_rgg_data[col_name].values.tolist()))
index += 1
index = 1
for col_name in rgg_rand_data:
if col_name == ('step%d_isol' % index):
rgg_rand_isol.append(statistics.mean(rgg_rand_data[col_name].values.tolist()))
if col_name == ('step%d_unsupp' % index):
rgg_rand_unsupp.append(statistics.mean(rgg_rand_data[col_name].values.tolist()))
index += 1
index = 1
for col_name in rand_rgg_data:
if col_name == ('step%d_isol' % index):
rand_rgg_isol.append(statistics.mean(rand_rgg_data[col_name].values.tolist()))
if col_name == ('step%d_unsupp' % index):
rand_rgg_unsupp.append(statistics.mean(rand_rgg_data[col_name].values.tolist()))
index += 1
index = 1
for col_name in rand_rand_data:
if col_name == ('step%d_isol' % index):
rand_rand_isol.append(statistics.mean(rand_rand_data[col_name].values.tolist()))
if col_name == ('step%d_unsupp' % index):
rand_rand_unsupp.append(statistics.mean(rand_rand_data[col_name].values.tolist()))
index += 1
# -
print(len(rgg_rgg_isol))
print(len(rgg_rgg_unsupp))
print(len(rgg_rand_isol))
print(len(rgg_rand_unsupp))
print(len(rand_rgg_isol))
print(len(rand_rgg_unsupp))
print(len(rand_rand_isol))
print(len(rand_rand_unsupp))
# +
cum_rgg_rgg_isol = []
cum_rgg_rgg_unsupp = []
cum_rgg_rand_isol = []
cum_rgg_rand_unsupp = []
cum_rand_rgg_isol = []
cum_rand_rgg_unsupp = []
cum_rand_rand_isol = []
cum_rand_rand_unsupp = []
total = []
for i in range(len(rgg_rgg_isol)):
if i == 0:
total.append(rgg_rgg_isol[i])
total.append(rgg_rgg_unsupp[i])
else:
total[0] += rgg_rgg_isol[i]
total[1] += rgg_rgg_unsupp[i]
cum_rgg_rgg_isol.append(total[0])
cum_rgg_rgg_unsupp.append(total[1])
total = []
for i in range(len(rgg_rand_isol)):
if i == 0:
total.append(rgg_rand_isol[i])
total.append(rgg_rand_unsupp[i])
else:
total[0] += rgg_rand_isol[i]
total[1] += rgg_rand_unsupp[i]
cum_rgg_rand_isol.append(total[0])
cum_rgg_rand_unsupp.append(total[1])
total = []
for i in range(len(rand_rgg_isol)):
if i == 0:
total.append(rand_rgg_isol[i])
total.append(rand_rgg_unsupp[i])
else:
total[0] += rand_rgg_isol[i]
total[1] += rand_rgg_unsupp[i]
cum_rand_rgg_isol.append(total[0])
cum_rand_rgg_unsupp.append(total[1])
total = []
for i in range(len(rand_rand_isol)):
if i == 0:
total.append(rand_rand_isol[i])
total.append(rand_rand_unsupp[i])
else:
total[0] += rand_rand_isol[i]
total[1] += rand_rand_unsupp[i]
cum_rand_rand_isol.append(total[0])
cum_rand_rand_unsupp.append(total[1])
# -
# ## Isolation vs Unsupport
plt.plot(range(len(cum_rgg_rgg_isol)), cum_rgg_rgg_isol)
plt.plot(range(len(cum_rgg_rgg_isol)), cum_rgg_rgg_unsupp)
plt.legend(['rgg_rgg_isol','rgg_rgg_unsupp'])
plt.title('Isolation vs Unsupport: RGG-RGG')
plt.savefig('Isolation vs Unsupport_RGG-RGG.png')
plt.show()
plt.plot(range(len(cum_rgg_rand_isol)), cum_rgg_rand_isol)
plt.plot(range(len(cum_rgg_rand_isol)), cum_rgg_rand_unsupp)
plt.legend(['rgg_rand_isol','rgg_rand_unsupp'])
plt.title('Isolation vs Unsupport: RGG-Rand')
plt.savefig('Isolation vs Unsupport_RGG-Rand.png')
plt.show()
plt.plot(range(len(cum_rand_rgg_isol)), cum_rand_rgg_isol)
plt.plot(range(len(cum_rand_rgg_isol)), cum_rand_rgg_unsupp)
plt.legend(['rand_rgg_isol','rand_rgg_unsupp'])
plt.title('Isolation vs Unsupport: Rand-RGG')
plt.savefig('Isolation vs Unsupport_Rand-RGG.png')
plt.show()
plt.plot(range(len(cum_rand_rand_isol)), cum_rand_rand_isol)
plt.plot(range(len(cum_rand_rand_isol)), cum_rand_rand_unsupp)
plt.legend(['rand_rand_isol','rand_rand_unsupp'])
plt.title('Isolation vs Unsupport: Rand-Rand')
plt.savefig('Isolation vs Unsupport_Rand-Rand.png')
plt.show()
# +
df_len = []
df_len.append(list(rgg_rgg_isol))
df_len.append(list(rgg_rand_isol))
df_len.append(list(rand_rgg_isol))
df_len.append(list(rand_rand_isol))
max_df_len = max(df_len, key=len)
x_val = list(range(len(max_df_len)))
# +
proc_isol = []
proc_unsupp = []
proc_isol.append(cum_rgg_rgg_isol)
proc_isol.append(cum_rgg_rand_isol)
proc_isol.append(cum_rand_rgg_isol)
proc_isol.append(cum_rand_rand_isol)
proc_unsupp.append(cum_rgg_rgg_unsupp)
proc_unsupp.append(cum_rgg_rand_unsupp)
proc_unsupp.append(cum_rand_rgg_unsupp)
proc_unsupp.append(cum_rand_rand_unsupp)
for x in x_val:
if len(rgg_rgg_isol) <= x:
proc_isol[0].append(cum_rgg_rgg_isol[len(rgg_rgg_isol) - 1])
proc_unsupp[0].append(cum_rgg_rgg_unsupp[len(rgg_rgg_isol) - 1])
if len(rgg_rand_isol) <= x:
proc_isol[1].append(cum_rgg_rand_isol[len(rgg_rand_isol) - 1])
proc_unsupp[1].append(cum_rgg_rand_unsupp[len(rgg_rand_isol) - 1])
if len(rand_rgg_isol) <= x:
proc_isol[2].append(cum_rand_rgg_isol[len(rand_rgg_isol) - 1])
proc_unsupp[2].append(cum_rand_rgg_unsupp[len(rand_rgg_isol) - 1])
if len(rand_rand_isol) <= x:
proc_isol[3].append(cum_rand_rand_isol[len(rand_rand_isol) - 1])
proc_unsupp[3].append(cum_rand_rand_unsupp[len(rand_rand_isol) - 1])
# -
plt.plot(x_val, proc_isol[0])
plt.plot(x_val, proc_isol[1])
plt.plot(x_val, proc_isol[2])
plt.plot(x_val, proc_isol[3])
plt.legend(['rgg_rgg_isol','rgg_rand_isol', 'rand_rgg_isol', 'rand_rand_isol'])
plt.title('Isolation trend')
plt.show()
plt.plot(x_val, proc_unsupp[0])
plt.plot(x_val, proc_unsupp[1])
plt.plot(x_val, proc_unsupp[2])
plt.plot(x_val, proc_unsupp[3])
plt.legend(['rgg_rgg_unsupp','rgg_rand_unsupp', 'rand_rgg_unsupp', 'rand_rand_unsupp'])
plt.title('Unsupport trend')
plt.show()
# ## Pie Chart
# +
init_death = 150
labels = ['Alive nodes', 'Initial death', 'Dead nodes from isolation', 'Dead nodes from unsupport']
alive = []
alive.append(statistics.mean(rgg_rgg_data['alive_nodes']))
alive.append(statistics.mean(rgg_rand_data['alive_nodes']))
alive.append(statistics.mean(rand_rgg_data['alive_nodes']))
alive.append(statistics.mean(rand_rand_data['alive_nodes']))
tot_isol = []
tot_isol.append(statistics.mean(rgg_rgg_data['tot_isol_node']))
tot_isol.append(statistics.mean(rgg_rand_data['tot_isol_node']))
tot_isol.append(statistics.mean(rand_rgg_data['tot_isol_node']))
tot_isol.append(statistics.mean(rand_rand_data['tot_isol_node']))
tot_unsupp = []
tot_unsupp.append(statistics.mean(rgg_rgg_data['tot_unsupp_node']))
tot_unsupp.append(statistics.mean(rgg_rand_data['tot_unsupp_node']))
tot_unsupp.append(statistics.mean(rand_rgg_data['tot_unsupp_node']))
tot_unsupp.append(statistics.mean(rand_rand_data['tot_unsupp_node']))
# +
deaths = [alive[0], init_death, tot_isol[0], tot_unsupp[0]]
plt.pie(deaths, labels=labels, autopct='%.1f%%')
plt.title('RGG-RGG death trend')
plt.show()
# +
deaths = [alive[1], init_death, tot_isol[1], tot_unsupp[1]]
plt.pie(deaths, labels=labels, autopct='%.1f%%')
plt.title('RGG-Rand death trend')
plt.show()
# +
deaths = [alive[2], init_death, tot_isol[2], tot_unsupp[2]]
plt.pie(deaths, labels=labels, autopct='%.1f%%')
plt.title('Rand-RGG death trend')
plt.show()
# +
deaths = [alive[3], init_death, tot_isol[3], tot_unsupp[3]]
plt.pie(deaths, labels=labels, autopct='%.1f%%')
plt.title('Rand-Rand death trend')
plt.show()
# -
# ## Compute the number of nodes
# +
x_val = np.arange(4)
labels = ['initial', 'final']
plt.bar(x_val, alive)
plt.xticks(x_val, graph_types)
plt.title('Alive nodes')
plt.savefig('alive nodes.png')
plt.show()
# -
# ## Compare the number of edges
# +
init_intra = []
init_intra.append(statistics.mean(rgg_rgg_data['init_intra_edge']))
init_intra.append(statistics.mean(rgg_rand_data['init_intra_edge']))
init_intra.append(statistics.mean(rand_rgg_data['init_intra_edge']))
init_intra.append(statistics.mean(rand_rand_data['init_intra_edge']))
init_inter = []
init_inter.append(statistics.mean(rgg_rgg_data['init_inter_edge']))
init_inter.append(statistics.mean(rgg_rand_data['init_inter_edge']))
init_inter.append(statistics.mean(rand_rgg_data['init_inter_edge']))
init_inter.append(statistics.mean(rand_rand_data['init_inter_edge']))
init_supp = []
init_supp.append(statistics.mean(rgg_rgg_data['init_supp_edge']))
init_supp.append(statistics.mean(rgg_rand_data['init_supp_edge']))
init_supp.append(statistics.mean(rand_rgg_data['init_supp_edge']))
init_supp.append(statistics.mean(rand_rand_data['init_supp_edge']))
fin_intra = []
fin_intra.append(statistics.mean(rgg_rgg_data['fin_intra_edge']))
fin_intra.append(statistics.mean(rgg_rand_data['fin_intra_edge']))
fin_intra.append(statistics.mean(rand_rgg_data['fin_intra_edge']))
fin_intra.append(statistics.mean(rand_rand_data['fin_intra_edge']))
fin_inter = []
fin_inter.append(statistics.mean(rgg_rgg_data['fin_inter_edge']))
fin_inter.append(statistics.mean(rgg_rand_data['fin_inter_edge']))
fin_inter.append(statistics.mean(rand_rgg_data['fin_inter_edge']))
fin_inter.append(statistics.mean(rand_rand_data['fin_inter_edge']))
fin_supp = []
fin_supp.append(statistics.mean(rgg_rgg_data['fin_supp_edge']))
fin_supp.append(statistics.mean(rgg_rand_data['fin_supp_edge']))
fin_supp.append(statistics.mean(rand_rgg_data['fin_supp_edge']))
fin_supp.append(statistics.mean(rand_rand_data['fin_supp_edge']))
# -
plt.bar(x_val-0.1, init_intra, width=0.2)
plt.bar(x_val+0.1, fin_intra, width=0.2)
plt.legend(labels)
plt.xticks(x_val, graph_types)
plt.title('Initial_intra_edge vs Final_intra_edge')
plt.show()
plt.bar(x_val-0.1, init_inter, width=0.2)
plt.bar(x_val+0.1, fin_inter, width=0.2)
plt.legend(labels)
plt.xticks(x_val, graph_types)
plt.title('Initial_inter_edge vs Final_inter_edge')
plt.show()
plt.bar(x_val-0.1, init_supp, width=0.2)
plt.bar(x_val+0.1, fin_supp, width=0.2)
plt.legend(labels)
plt.xticks(x_val, graph_types)
plt.title('Initial_support_edge vs Final_support_edge')
plt.show()
# ## Network Analysis
# +
init_far = []
init_far.append(statistics.mean(rgg_rgg_data['init_far_node']))
init_far.append(statistics.mean(rgg_rand_data['init_far_node']))
init_far.append(statistics.mean(rand_rgg_data['init_far_node']))
init_far.append(statistics.mean(rand_rand_data['init_far_node']))
fin_far = []
fin_far.append(statistics.mean(rgg_rgg_data['fin_far_node']))
fin_far.append(statistics.mean(rgg_rand_data['fin_far_node']))
fin_far.append(statistics.mean(rand_rgg_data['fin_far_node']))
fin_far.append(statistics.mean(rand_rand_data['fin_far_node']))
# -
plt.bar(x_val-0.1, init_far, width=0.2)
plt.bar(x_val+0.1, fin_far, width=0.2)
plt.legend(labels)
plt.xticks(x_val, graph_types)
plt.title('Initial_far_node vs Final_far_node')
plt.show()
# +
init_clust = []
init_clust.append(statistics.mean(rgg_rgg_data['init_clust']))
init_clust.append(statistics.mean(rgg_rand_data['init_clust']))
init_clust.append(statistics.mean(rand_rgg_data['init_clust']))
init_clust.append(statistics.mean(rand_rand_data['init_clust']))
fin_clust = []
fin_clust.append(statistics.mean(rgg_rgg_data['fin_clust']))
fin_clust.append(statistics.mean(rgg_rand_data['fin_clust']))
fin_clust.append(statistics.mean(rand_rgg_data['fin_clust']))
fin_clust.append(statistics.mean(rand_rand_data['fin_clust']))
# -
plt.bar(x_val-0.1, init_clust, width=0.2)
plt.bar(x_val+0.1, fin_clust, width=0.2)
plt.legend(labels)
plt.xticks(x_val, graph_types)
plt.title('Initial_clustering_coefficient vs Final_clustering_coefficient')
plt.show()
# +
init_mean_deg = []
init_mean_deg.append(statistics.mean(rgg_rgg_data['init_mean_deg']))
init_mean_deg.append(statistics.mean(rgg_rand_data['init_mean_deg']))
init_mean_deg.append(statistics.mean(rand_rgg_data['init_mean_deg']))
init_mean_deg.append(statistics.mean(rand_rand_data['init_mean_deg']))
fin_mean_deg = []
fin_mean_deg.append(statistics.mean(rgg_rgg_data['fin_mean_deg']))
fin_mean_deg.append(statistics.mean(rgg_rand_data['fin_mean_deg']))
fin_mean_deg.append(statistics.mean(rand_rgg_data['fin_mean_deg']))
fin_mean_deg.append(statistics.mean(rand_rand_data['fin_mean_deg']))
# -
plt.bar(x_val-0.1, init_mean_deg, width=0.2)
plt.bar(x_val+0.1, fin_mean_deg, width=0.2)
plt.legend(labels)
plt.xticks(x_val, graph_types)
plt.title('Initial_mean_degree vs Final_mean_degree')
plt.show()
# +
init_larg_comp = []
init_larg_comp.append(statistics.mean(rgg_rgg_data['init_larg_comp']))
init_larg_comp.append(statistics.mean(rgg_rand_data['init_larg_comp']))
init_larg_comp.append(statistics.mean(rand_rgg_data['init_larg_comp']))
init_larg_comp.append(statistics.mean(rand_rand_data['init_larg_comp']))
fin_larg_comp = []
fin_larg_comp.append(statistics.mean(rgg_rgg_data['fin_larg_comp']))
fin_larg_comp.append(statistics.mean(rgg_rand_data['fin_larg_comp']))
fin_larg_comp.append(statistics.mean(rand_rgg_data['fin_larg_comp']))
fin_larg_comp.append(statistics.mean(rand_rand_data['fin_larg_comp']))
# -
plt.bar(x_val-0.1, init_larg_comp, width=0.2)
plt.bar(x_val+0.1, fin_larg_comp, width=0.2)
plt.legend(labels)
plt.xticks(x_val, graph_types)
plt.title('Initial_largest_component_size vs Final_largest_component_size')
plt.show()
# +
deg_assort = []
a = rgg_rgg_data['deg_assort'].fillna(0)
b = rgg_rand_data['deg_assort'].fillna(0)
c = rand_rgg_data['deg_assort'].fillna(0)
d = rand_rand_data['deg_assort'].fillna(0)
deg_assort.append(statistics.mean(a))
deg_assort.append(statistics.mean(b))
deg_assort.append(statistics.mean(c))
deg_assort.append(statistics.mean(d))
# -
plt.bar(x_val, deg_assort)
plt.xticks(x_val, graph_types)
plt.title('Degree Assortativity')
plt.show()
dist_deg_cent = []
dist_deg_cent.append(statistics.mean(rgg_rgg_data['dist_deg_cent']))
dist_deg_cent.append(statistics.mean(rgg_rand_data['dist_deg_cent']))
dist_deg_cent.append(statistics.mean(rand_rgg_data['dist_deg_cent']))
dist_deg_cent.append(statistics.mean(rand_rand_data['dist_deg_cent']))
plt.bar(x_val, dist_deg_cent)
plt.xticks(x_val, graph_types)
plt.title('Distance to degree centre from the attack point')
plt.show()
dist_bet_cent = []
dist_bet_cent.append(statistics.mean(rgg_rgg_data['dist_bet_cent']))
dist_bet_cent.append(statistics.mean(rgg_rand_data['dist_bet_cent']))
dist_bet_cent.append(statistics.mean(rand_rgg_data['dist_bet_cent']))
dist_bet_cent.append(statistics.mean(rand_rand_data['dist_bet_cent']))
plt.bar(x_val, dist_bet_cent)
plt.xticks(x_val, graph_types)
plt.title('Distance to betweenes centre from the attack point')
plt.show()
| data analysis/Buldyrev_vs_RGG/.ipynb_checkpoints/Ploting Graph_ER-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Import required packages
import quandl, math
import numpy as np
import pandas as pd
from sklearn import preprocessing, svm
from sklearn.model_selection import cross_validate
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LinearRegression
# Get dataframe from quandl
df = quandl.get("WIKI/GOOGL")
# show first lines of data
df.head()
# -
# See columns names
df.columns
# Pick some columns as the data
df = df[['Adj. Open', 'Adj. High', 'Adj. Low', 'Adj. Close', 'Adj. Volume']]
df.columns
# Add a couple of columns
df['HL_PCT'] = (df['Adj. High'] - df['Adj. Low']) / df['Adj. Close'] * 100.0
df['PCT_change'] = (df['Adj. Close'] - df['Adj. Open']) / df['Adj. Open'] * 100.0
# Rearrange the dataframe
df = df[['Adj. Close', 'HL_PCT', 'PCT_change', 'Adj. Volume']]
df.head()
# +
# Picking the label colum.
# Adjusted Close seems interesting, so
forecast_col = 'Adj. Close'
# Fill missing values with -99 999. The value is recommended
df.fillna(value=-99999, inplace=True)
# Pick a range for a valid forecast.
#with forecasting, you want to forecast out a certain number of datapoints.
#We're saying we want to forecast out 1% of the entire length of the dataset.
#Thus, if our data is 100 days of stock prices,
#we want to be able to predict the price 1 day out into the future.
#Choose whatever you like.
#If you are just trying to predict tomorrow's price,
#then you would just do 1 day out, and the forecast would be just one day out.
#If you predict 10 days out, we can actually generate a forcast for every day,
#for the next week and a half.
# Picking 1% of the data set lenght
forecast_out = int(math.ceil(0.01 * len(df)))
forecast_out
# -
# Add the label colum with name "label"
df['label'] = df[forecast_col].shift(-forecast_out)
df.head()
# +
# Visually inspecting the data
import matplotlib.pyplot as plt
# %matplotlib inline
df['label'].plot()
df['Adj. Close'].plot()
plt.show()
# -
#Drop out any NaNs
df.dropna(inplace=True)
X = np.array(df.drop(['label'], 1))
y = np.array(df['label'])
plt.plot(X,y)
df.corr()
from pandas.plotting import scatter_matrix
df.hist()
scatter_matrix(df)
# Generally, you want your features in machine learning to be in a range of -1 to 1.
#This may do nothing, but it usually speeds up processing and can also help with accuracy.
#Because this range is so popularly used, it is included in the preprocessing module of Scikit-Learn.
#To utilize this, you can apply preprocessing.scale to your X variable:
X = preprocessing.scale(X)
y = np.array(df['label'])
plt.plot(X,y)
#The way this works is you take, for example, 75% of your data,
#and use this to train the machine learning classifier.
#Then you take the remaining 25% of your data, and test the classifier.
#Since this is your sample data, you should have the features and known labels.
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2)
#The return here is the training set of features,
#testing set of features, training set of labels,
#and testing set of labels. Now, we're ready to plt.plot(X_test,y_test)
#define our classifier. There are many classifiers
#in general available through Scikit-Learn, and even
#a few specifically for regression. We'll show a
#couple in this example, but for now, let's use
#Support Vector Regression from Scikit-Learn's svm package:
clf = svm.SVR()
clf.fit(X_train, y_train)
confidence = clf.score(X_test, y_test)
print(confidence)
clf = LinearRegression()
clf.fit(X_train, y_train)
confidence = clf.score(X_test, y_test)
print(confidence)
for k in ['linear','poly','rbf','sigmoid']:
clf = svm.SVR(kernel=k)
clf.fit(X_train, y_train)
confidence = clf.score(X_test, y_test)
print(k,confidence)
# +
X = np.array(df.drop(['label'], 1))
X = preprocessing.scale(X)
dsos_ = X.size
X_lately = X[-forecast_out:]
X = X[:-forecast_out]
dshs_ = X.size
dsts_ = X_lately.size
df.dropna(inplace=True)
y = np.array(df['label'])
dslos_ = y.size
y = y[:-forecast_out]
dslns_ = y.size
print("Data sets sizes")
print("---------------")
print("head + tail = original size")
print(X.size,"+",X_lately.size,"=",X.size+X_lately.size)
print("Label sizes")
print(dslos_,dslns_)
# -
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2)
clf = LinearRegression(n_jobs=-1)
clf.fit(X_train, y_train)
confidence = clf.score(X_test, y_test)
print(confidence)
forecast_set = clf.predict(X_lately)
print(forecast_set, confidence, forecast_out)
# +
import datetime
import matplotlib.pyplot as plt
from matplotlib import style
style.use('ggplot')
df['Forecast'] = np.nan
last_date = df.iloc[-1].name
last_unix = last_date.timestamp()
one_day = 86400
next_unix = last_unix + one_day
for i in forecast_set:
next_date = datetime.datetime.fromtimestamp(next_unix)
next_unix += one_day
df.loc[next_date] = [np.nan for _ in range(len(df.columns)-1)]+[i]
# -
df['Adj. Close'].plot()
df['Forecast'].plot()
plt.legend(loc=2)
plt.xlabel('Date')
plt.ylabel('Price')
plt.xlim([datetime.date(2017,6,1).toordinal(),datetime.date(2017,8,22).toordinal()])
plt.ylim([910,1030])
plt.show()
| Pandas_foundations/Pandas-Quandl-ScikitLearn.ipynb |
# ---
# title: "Pandas Exercises"
# author: "Vaishnavi"
# date: 2020-08-10
# description: "-"
# type: technical_note
# draft: false
#
#
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.3 64-bit (''py38'': conda)'
# language: python
# name: python38364bitpy38conda4b9668195d454c7a8d87f7eb83269b8a
# ---
# +
# 1. Loading and finding info of data
import pandas as pd
df = pd.read_excel(open('kapi.xlsx', 'rb'),sheet_name='ActorScore')
df.info()
print("----------------")
print(df.describe())
# +
# 2. Finding shape of data
df.shape
# -
# 3. Seeing the first and last data
print(df.head(5))
print("________________________________________________________________________________________________")
print(df.tail(5))
# +
# 4. Looking null values in dataset
print(df.isna().sum())
print("______________")
print(df.isna().sum().sum())
# +
# 5. Dropping columns
df.drop(['Updated By'],axis=1,inplace=True)
df
# +
# 6. Dropping null value rows
df.dropna(how='any',inplace=True)
print(df)
print(df.shape)
# +
# 7. Checking whether null values are present
df.isna().sum()
# +
# 8. Prints the datatypes of all columns
print(df.dtypes)
# +
# 9. Finding unique actor names and year
act=list(df['Movie'].unique())
print(act)
print("_____________________________________________")
y=list(df['Year'].unique())
print(y)
# +
# 10. Finding avg in each score of the artist
c=list(df.iloc[::,3:6])
for i in c:
s=df.groupby(["Artist Name"])[i].sum()
av=s/(df.groupby(["Artist Name"])[i].count())
print(av)
# +
# 11. Data bucketing or data binning on box office score
op_labels = ['poor','moderate', 'good', 'excellent']
category = [0.,3.,5.,7.,10.]
df['avg boxoffice rating'] = pd.cut(df['Box Office Score'], labels=op_labels, bins=category, include_lowest=False)
df
# +
# 12. Renaming column names
df.rename(columns = {'Artist Name':'artistname', 'Audience Score':'audiencescore', 'Box Office Score':'boxofficescore'}, inplace = True)
df
# +
# 13. Max rating in a year with movie name
df.groupby(["Year","Movie"])["audiencescore"].max()
# +
# 14. Calculates mean
scores=df.iloc[:,3:]
scores.mean()
# +
# 15. If condition
numbers = {'set_of_numbers': [1,2,3,4,5,6,7,8,9,10]}
df = pd.DataFrame(numbers,columns=['set_of_numbers'])
df.loc[df['set_of_numbers'] <= 4, 'equal_or_lower_than_4?'] = 'True'
df.loc[df['set_of_numbers'] > 4, 'equal_or_lower_than_4?'] = 'False'
print (df)
# +
# 16. Strings - If condition
names = {'First_name': ['Jon','Bill','Maria','Emma']}
df = pd.DataFrame(names,columns=['First_name'])
df.loc[df['First_name'] == 'Bill', 'name_match'] = 'Match'
df.loc[df['First_name'] != 'Bill', 'name_match'] = 'Mismatch'
print (df)
# +
# 17. & and | condition
df.loc[(df['First_name'] == 'Bill') | (df['First_name'] == 'Emma'), 'name_match'] = 'Match'
df.loc[(df['First_name'] != 'Bill') & (df['First_name'] != 'Emma'), 'name_match'] = 'Mismatch'
print (df)
# +
# 18. To set values
numbers = {'set_of_numbers': [1,2,3,4,5,6,7,8,9,10,0,0]}
df = pd.DataFrame(numbers,columns=['set_of_numbers'])
print (df)
df.loc[df['set_of_numbers'] == 0, 'set_of_numbers'] = 999
df.loc[df['set_of_numbers'] == 5, 'set_of_numbers'] = 555
print (df)
# +
# 19. Change NAN values to 0.0
import numpy as np
numbers = {'set_of_numbers': [1,2,3,4,5,6,7,8,9,10,np.nan,np.nan]}
df = pd.DataFrame(numbers,columns=['set_of_numbers'])
print (df)
df.loc[df['set_of_numbers'].isnull(), 'set_of_numbers'] = 0
print (df)
# +
# 20. Copy function
df2 = df.copy()
df2
# +
# 21. String methods
s = pd.Series(['A', 'B', 'C', 'Aaba', 'Baca', np.nan, 'CABA', 'dog', 'cat'])
s.str.lower()
# +
# 22. Merge
left = pd.DataFrame({'key': ['foo', 'bar'], 'lval': [1, 2]})
right = pd.DataFrame({'key': ['foo', 'bar'], 'rval': [4, 5]})
print("left\n",left)
print('---------------------------------------')
print("right\n",right)
print('---------------------------------------')
print(pd.merge(left, right, on='key'))
# +
# 23. Drop duplicate
df = pd.DataFrame({'A': [1, 2, 2, 3, 4, 5, 5, 5, 6, 7, 7]})
df.drop_duplicates(subset='A')
# +
# 24. Append
sr1 = pd.Series(['New York', 'Chicago', 'Toronto', 'Lisbon', 'Rio'])
# Create the first Index
index_1 = ['City 1', 'City 2', 'City 3', 'City 4', 'City 5']
sr1.index = index_1
# Creating the second Series
sr2 = pd.Series(['Chicage', 'Shanghai', 'Beijing', 'Jakarta', 'Seoul'])
# Create the second Index
index_2 = ['City 6', 'City 7', 'City 8', 'City 9', 'City 10']
sr2.index = index_2
print(sr1)
print('---------------------------------------')
print(sr2)
print('---------------------------------------')
# append sr2 at the end of sr1
result = sr1.append(sr2)
print(result)
# +
# 25. Empty series NAN count
# min_count = 0 is the default
pd.Series([]).sum()
# When passed min_count = 1,
# sum of an empty series will be NaN
pd.Series([]).sum(min_count = 1)
# +
# 26. Rank function
df = pd.DataFrame(data={'Animal': ['cat', 'penguin', 'dog',
'spider', 'snake'],
'Number_legs': [4, 2, 4, 8, np.nan]})
df['default_rank'] = df['Number_legs'].rank()
df
# +
# 27. Setting a column a with the sum value
data = {'name': ['John', 'Peter', 'Karl'],
'age' : [23, 42, 19]}
val = pd.DataFrame(data)
# sum of all salary
val['total'] = val['age'].sum()
val
# +
# 28. Date and Time function
df = pd.DataFrame({'year': [2015, 2016],
'month': [2, 3],
'day': [4, 5]})
pd.to_datetime(df)
# +
# 29. Slicing
df[:2]
# +
# 30. Concat function
s1 = pd.Series(['a', 'b'])
s2 = pd.Series(['c', 'd'])
pd.concat([s1, s2])
# -
| docs/python/pandas/Pandas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Find the neighborhood a coordinate falls within
# #### Author: <NAME>
# #### Nov 6th, 2019
#
# #### Note:
# 1. The coordinate points need to be formatted as (longitude, latitude).
# ## Function
# Import following packages.
import fiona
from shapely.geometry import shape,mapping, Point, Polygon, MultiPolygon
import pandas as pd
import numpy as np
# +
def find_neighborhood(neighborhood, lat, long):
"""
Find the neighborhood where each coordinate in a list falls within. Remember to import
packages above first.
Note: Point in this funciton is formatted as (long, lat).
Parameters:
--------------------
neighborhood: string
String of data path and file name. The file should be a shape file containing
neighborhood name and boundary coordinates. Check which dictionary keys to
call for shaping polygon and neighborhood name.
lat: pd.Series
The latitude column of a panda dataframe which we need to find the neighborhood for.
long: pd.Series
The longitude column of a panda dataframe which we need to find the neighborhood for.
Return:
--------------------
neigh_names: list
The names of neighborhoods where each pair of lat and long lies within.
"""
neigh_names = []
multipol = fiona.open(neighborhood)
for i in range(0,len(lat)):
# Point in this case is formatted as (long, lat).
point = Point(long[i], lat[i])
counter = 0
for feat in multipol:
if point.within(shape(feat['geometry'])):
neigh_names.append(feat['properties']['pri_neigh'])
counter +=1
# If point does not find any neighborhood, return nan
if counter == 0:
neigh_names.append(np.nan)
return(neigh_names)
# -
# ## Find neighborhoods for datasets
# ### Station
station = pd.read_csv('stations.csv')
station['neighborhood'] = find_neighborhood("geo_export_cdc4661f-83bd-40ef-9034-c4bdb778cf15.shp", station['lat'], station['long'])
# Some bike stations are out of scope in terms of searching neighborhood.
sum(station['neighborhood_id'].isna())
# +
# Substitue neighborhood name with id.
neighborhood = pd.read_csv('neighborhood.csv')
station = station.merge(neighborhood, left_on = 'neighborhood', right_on = 'name', how='left')
station.drop(['name', 'neighborhood'], axis = 1, inplace=True)
# -
station['neighborhood_id'] = station['neighborhood_id'].astype('Int64')
station.to_csv('stations.csv', index=False)
# ### Segment
#
# 1. Data has start neighborhood and end neighborhood. 1046 rows
# 2. If start and/or end neighborhoods are out of scope, then delete entire row. 963 rows
# 3. Keep segments where start_neigh equals to end_neigh. 609 rows
seg = pd.read_csv('segments.csv')
seg['start_neigh'] = find_neighborhood("geo_export_cdc4661f-83bd-40ef-9034-c4bdb778cf15.shp", seg['START_LATITUDE'], seg['START_LONGITUDE'])
seg['end_neigh'] = find_neighborhood("geo_export_cdc4661f-83bd-40ef-9034-c4bdb778cf15.shp", seg['END_LATITUDE'], seg['END_LONGITUDE'])
# +
seg = seg.merge(neighborhood, left_on = 'start_neigh', right_on = 'name', how = 'left')
seg.rename(columns = {'neighborhood_id': 'start_neigh_id'}, inplace=True)
seg.drop(['start_neigh', 'name'], axis = 1, inplace=True)
seg = seg.merge(neighborhood, left_on = 'end_neigh', right_on = 'name', how = 'left')
seg.rename(columns = {'neighborhood_id': 'end_neigh_id'}, inplace = True)
seg.drop(['end_neigh', 'name'], axis=1, inplace=True)
# -
seg.to_csv('segments.csv', index=False)
# ### Crash
crash = pd.read_csv('/Users/Jenny/Downloads/Clean_Crash.csv')
crash.drop(['Unnamed: 0'], axis=1, inplace=True)
crash['neighborhood'] = find_neighborhood("geo_export_cdc4661f-83bd-40ef-9034-c4bdb778cf15.shp", crash['LATITUDE'], crash['LONGITUDE'])
crash.to_csv('crash_backup.csv', index=False)
# +
neighborhood = pd.read_csv('neighborhood.csv')
crash = crash.merge(neighborhood, left_on = 'neighborhood', right_on = 'name', how='left')
crash.drop(['name', 'neighborhood'], axis = 1, inplace=True)
# -
crash['neighborhood_id'] = crash['neighborhood_id'].astype('Int64')
crash.to_csv('crash.csv', index=False)
# ## Export all neighborhoods as csv with its id.
df = fiona.open("/Users/Jenny/Desktop/MSCA/Data Engineering/Project/Boundaries - Neighborhoods/Boundaries - Neighborhoods/geo_export_c0182ca0-1f35-4cb2-b1bd-75b05d26853c.shp")
for feat in df:
print(feat['properties']['pri_neigh'])
df.next()
# +
neighborhood = pd.DataFrame({'neighborhood_id':[], 'name':[]})
for feat in df:
neighborhood = neighborhood.append({'neighborhood_id':feat['id'], 'name':feat['properties']['pri_neigh']}
, ignore_index=True)
# -
neighborhood.to_csv('neighborhood.csv', index=False)
neighborhood=pd.read_csv('/Users/Jenny/Desktop/MSCA/Data Engineering/Project/Boundaries - Neighborhoods/neighborhood.csv')
for i in range(0,len(neighborhood)):
print("UPDATE neighborhood SET NAME = '{}' WHERE neighborhood_id = {};".format(neighborhood['name'][i], neighborhood['neighborhood_id'][i]))
neighborhood
| wrangle02-geocode-neighborhoods.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # sudire.py example notebook
# The aim of this notebook is to show how to perform Sufficient Dimension Reduction using the direpack package. The data we will use is the [auto-mpg dataset](http://archive.ics.uci.edu/ml/datasets/Auto+MPG). We wil show how the dimension of the central subspace and a basis for the central subspace can be estimated using Sufficient Dimension Reduction via Ball covariance and by using a user defined function.
import matplotlib.pyplot as plt
import seaborn as sns
import numpy as np
import pandas as pd
from scipy.stats import norm
from scipy import stats
from direpack import sudire, estimate_structural_dim
from direpack import sudire_plot
import warnings
from sklearn.model_selection import train_test_split
warnings.filterwarnings('ignore')
plt.rcParams["figure.figsize"] = [16,13]
plt.rcParams['figure.constrained_layout.use'] = True
from matplotlib import rcParams
rcParams.update({'figure.autolayout': True})
# ## Data preprocessing
auto_data = pd.read_csv('..\\Data\\auto-mpg.csv', index_col='car name')
display(auto_data.head())
print('dataset shape is',auto_data.shape)
print(auto_data.dtypes)
# Looking at the data, we see that the horsepower variable should be a numeric variable but is displayed as type object. This is because missing values are coded as '?'. We thus remove those missing values. After this step, there are no more missing values into the data.
auto_data = auto_data[auto_data.horsepower != '?']
auto_data.horsepower = auto_data.horsepower.astype('float')
print('data types \n', auto_data.dtypes)
print('any missing values \n',auto_data.isnull().any())
X = auto_data.copy()
y = X['mpg']
X.drop('mpg', axis=1, inplace=True)
X.drop('origin', axis = 1, inplace = True)
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.3, random_state=42)
# # Estimating a basis of the central subspace
# First let us suppose that we know the dimension of the central subspace to be 2. We will then see how to estimate a basis for the central subspaces using the various options.
struct_dim = 2
# # via distance covariance
dcov_auto = sudire('dcov-sdr', center_data= True, scale_data=True,n_components=struct_dim)
dcov_auto.fit(X_train.values, y_train.values)
dcov_auto.x_loadings_
# ## via Martingale Difference Divergence
mdd_auto = sudire('mdd-sdr', center_data= True, scale_data=True,n_components=struct_dim)
mdd_auto.fit(X_train.values, y_train.values)
mdd_auto.x_loadings_
# ## User defined functions
# Here we show how user can optimize their own functions as is done for Distance Covariance and Martingale Difference Divergence.
# For this example we will use Ball covariance. There is a python package : [Ball](https://pypi.org/project/Ball/) available on PyPi which computes the Ball covariance between random variables. We follow the development of the article [Robust sufficient Dimension Reduction Via Ball covariance](https://www.sciencedirect.com/science/article/pii/S0167947319301380). The process is similar to using scipy.optimize.minimize function.
import Ball
# First we define the objective function to be optimized. Here, beta is the flattened array representing the basis of the central subpace. A series of arguments can be passed to this function, including the X and y data as well as the dimension of the central subspace.
def ballcov_func(beta, *args):
X= args[0]
Y= args[1]
h=args[2]
beta = np.reshape(beta,(-1,h),order = 'F')
X_dat = np.matmul(X, beta)
res = Ball.bcov_test(X_dat,Y,num_permutations=0)[0]
return(-10*res)
# Next we define the contraints and additional optimization arguments. both the constraints and arguments are assumed to be dicts or tuples.
# +
def optim_const(beta, *args):
X= args[0]
h= args[1]
i = args[2]
j = args[3]
beta = np.reshape(beta,(-1,h),order = 'F')
covx = np.cov(X, rowvar=False)
ans = np.matmul(np.matmul(beta.T,covx), beta) - np.identity(h)
return(ans[i,j])
ball_const= []
for i in range(0, struct_dim):
for j in range(0,struct_dim):
ball_const.append({'type': 'eq', 'fun' : optim_const,
'args':(X_train,struct_dim,i,j)})
ball_const =tuple(ball_const)
optim_args = (X_train,y_train, struct_dim)
# -
bcov_auto = sudire(ballcov_func, center_data= True, scale_data=True,n_components=struct_dim)
bcov_auto.fit(X_train.values, y_train.values)
bcov_auto.x_loadings_
# ## Estimating the dimension of the central subspace
# The dimension of the central subspace can be estimated using the bootstrap method proposed in [Sufficient Dimension Reduction via Distance Covariance](https://www.tandfonline.com/doi/abs/10.1080/10618600.2015.1026601). All the implemented sdr methods can be used. Here we present the method using Directional Regression.
# +
central_dim, diff_vec = estimate_structural_dim('dr',X_train.values, y_train.values, B=100, n_slices=4)
central_dim
# -
# ## Plots
# Once the sufficient Dimension Reduction has been done, an OLS regression is fitted using the reduced subset of variables. we can visualise the predicted response values using the plot functions from sudire_plots.
sdr_plot=sudire_plot(dcov_auto,['w','w','g','y','m'])
sdr_plot.plot_yyp(label='mpg',title='fitted vs true mpg')
# The projections of the data can also be visualised
sdr_plot=sudire_plot(dcov_auto,['w','w','g','y','m'])
sdr_plot.plot_projections(label='mpg', title='projected data')
| examples/.ipynb_checkpoints/sudire_example-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Tasks:
#
# * Creating a scatter plot that shows how the tumor volume changes over time for each treatment.
# * Creating a scatter plot that shows how the number of [metastatic](https://en.wikipedia.org/wiki/Metastasis) (cancer spreading) sites changes over time for each treatment.
# * Creating a scatter plot that shows the number of mice still alive through the course of treatment (Survival Rate)
# * Creating a bar graph that compares the total % tumor volume change for each drug across the full 45 days.
# * Include 3 observations about the results of the study. Use the visualizations you generated from the study data as the basis for your observations.
# +
# Dependencies and Setup
# %matplotlib inline
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
from scipy import stats
# Hide warning messages in notebook
import warnings
warnings.filterwarnings('ignore')
# File to Load (Remember to Change These)
mouse_drug_data_to_load = "data/mouse_drug_data.csv"
clinical_trial_data_to_load = "data/clinicaltrial_data.csv"
# Read the Mouse and Drug Data and the Clinical Trial Data
mouse_trial = pd.read_csv(mouse_drug_data_to_load)
clinical_trial = pd.read_csv(clinical_trial_data_to_load)
# Combine the data into a single dataset
clinical_mouse_trial = mouse_trial.merge(clinical_trial, on='Mouse ID')
# Display the data table for preview
clinical_mouse_trial.head()
# -
# ## Tumor Response to Treatment
# +
# Store the Mean Tumor Volume Data Grouped by Drug and Timepoint & Convert to DataFrame
mean_tumor_volume_df = clinical_mouse_trial.groupby(by=(['Drug', 'Timepoint'])).mean()
# Preview DataFrame
mean_tumor_volume_df.head(20)
mean_tumor_volume_df = mean_tumor_volume_df.drop(columns=('Metastatic Sites'))
mean_tumor_volume_df = mean_tumor_volume_df.reset_index(drop=False)
mean_tumor_volume_df.head()
# +
# Store the Standard Error of Tumor Volumes Grouped by Drug and Timepoint & Convert to DataFrame
std_error_tumor_volume_df = clinical_mouse_trial.groupby(by=(clinical_mouse_trial['Drug'], clinical_mouse_trial['Timepoint'])).sem()
# Preview DataFrame
std_error_tumor_volume_df.head(20)
std_error_tumor_volume_df = std_error_tumor_volume_df.drop(columns=(['Metastatic Sites', 'Mouse ID']))
std_error_tumor_volume_df = std_error_tumor_volume_df.reset_index(drop=False)
std_error_tumor_volume_df.head()
# -
# Minor Data Munging to Re-Format the Data Frames
tumor_volume = mean_tumor_volume_df.pivot(index='Timepoint', columns='Drug', values='Tumor Volume (mm3)')
# Preview that Reformatting worked
tumor_volume = tumor_volume.drop(columns=(['Ceftamin', 'Naftisol', 'Propriva', 'Ramicane', 'Stelasyn', 'Zoniferol']))
tumor_volume.head()
# +
# Generate the Plot (with Error Bars)
plt.grid()
plt.plot(tumor_volume['Capomulin'], markersize=5, color='red', marker='o', linestyle=':', linewidth=1, label='Capomulin')
plt.plot(tumor_volume['Infubinol'], markersize=5, color='blue', marker='^', linestyle=':', linewidth=1, label='Infubinol')
plt.plot(tumor_volume['Ketapril'], markersize=5, color='green', marker='s', linestyle=':', linewidth=1, label='Ketapril')
plt.plot(tumor_volume['Placebo'], markersize=5, color='black', marker='d', linestyle=':', linewidth=1, label='Placebo')
# plt.errorbar(tumor_volume['Capomulin'], tumor_volume['Capomulin'].sem)
# Create a legend for our chart
plt.title('Tumor Response to Treatment')
plt.ylabel('Tumor Volume (mm3)')
plt.xlabel('Time (Days)')
plt.legend()
# Show the chart
plt.show()
# Save the Figure
# -
# ## Metastatic Response to Treatment
# +
# Store the Mean Met. Site Data Grouped by Drug and Timepoint
mean_met_site_df = clinical_mouse_trial.groupby(by=['Drug', 'Timepoint']).mean()
mean_met_site_df = mean_met_site_df.drop(columns='Tumor Volume (mm3)')
# Preview DataFrame
mean_met_site_df.head()
# -
mean_met_site_pivot = mean_met_site_df.reset_index(drop=False)
mean_met_site_pivot = mean_met_site_pivot.pivot(index='Timepoint', columns='Drug', values='Metastatic Sites')
mean_met_site_pivot = mean_met_site_pivot.drop(columns=(['Ceftamin', 'Naftisol', 'Propriva', 'Ramicane', 'Stelasyn', 'Zoniferol']))
mean_met_site_pivot.head()
# +
# Store the Standard Error associated with Met. Sites Grouped by Drug and Timepoint
standard_error_met_df = clinical_mouse_trial.groupby(by=['Drug', 'Timepoint']).sem()
standard_error_met_df = standard_error_met_df.drop(columns=['Mouse ID', 'Tumor Volume (mm3)'])
# Preview DataFrame
standard_error_met_df.head()
# -
# Minor Data Munging to Re-Format the Data Frames
standard_error_met_df = standard_error_met_df.reset_index(drop=False)
met_pivot = standard_error_met_df.pivot(index='Timepoint', columns='Drug', values='Metastatic Sites')
met_pivot = met_pivot.drop(columns=(['Ceftamin', 'Naftisol', 'Propriva', 'Ramicane', 'Stelasyn', 'Zoniferol']))
# Preview that Reformatting worked
met_pivot.head()
# +
# Generate the Plot (with Error Bars)
plt.plot(mean_met_site_pivot['Capomulin'], markersize=5, color='red', marker='o', linestyle=':', linewidth=1, label='Capomulin')
plt.plot(mean_met_site_pivot['Infubinol'], markersize=5, color='blue', marker='^', linestyle=':', linewidth=1, label='Infubinol')
plt.plot(mean_met_site_pivot['Ketapril'], markersize=5, color='green', marker='s', linestyle=':', linewidth=1, label='Ketapril')
plt.plot(mean_met_site_pivot['Placebo'], markersize=5, color='black', marker='d', linestyle=':', linewidth=1, label='Placebo')
plt.errorbar(met_pivot['Capomulin'], mean_met_site_pivot['Capomulin'])
plt.errorbar(mean_met_site_pivot['Infubinol'], met_pivot['Infubinol'])
plt.errorbar(mean_met_site_pivot['Ketapril'], met_pivot['Ketapril'])
plt.errorbar(mean_met_site_pivot['Placebo'], met_pivot['Placebo'])
# Save the Figure
plt.ylabel('Metastatic Sites')
plt.xlabel('Treatment Over Time (Days)')
plt.title('Metastatic Spread During Treatment')
plt.legend()
plt.grid()
# Show the Figure
plt.show()
# -
# ## Survival Rates
# +
# Store the Count of Mice Grouped by Drug and Timepoint (W can pass any metric)
survival = clinical_mouse_trial.groupby(by=['Drug', 'Timepoint']).count()
survival = survival.drop(columns=['Tumor Volume (mm3)', 'Metastatic Sites'])
# Convert to DataFrame
# Preview DataFrame
survival.head()
# +
# Minor Data Munging to Re-Format the Data Frames
survival = ((survival/25) * 100)
survival_pivot = survival.reset_index(drop=False)
survival_pivot = survival_pivot.pivot(index='Timepoint', columns='Drug', values='Mouse ID')
survival_pivot = survival_pivot.drop(columns=['Ceftamin', 'Naftisol', 'Propriva', 'Ramicane', 'Stelasyn', 'Zoniferol'])
# Preview the Data Frame
survival_pivot.head(15)
# +
# Generate the Plot (Accounting for percentages)
plt.plot(survival_pivot['Capomulin'], markersize=5, color='red', marker='o', linestyle=':', linewidth=1, label='Capomulin')
plt.plot(survival_pivot['Infubinol'], markersize=5, color='blue', marker='^', linestyle=':', linewidth=1, label='Infubinol')
plt.plot(survival_pivot['Ketapril'], markersize=5, color='green', marker='s', linestyle=':', linewidth=1, label='Ketapril')
plt.plot(survival_pivot['Placebo'], markersize=5, color='black', marker='d', linestyle=':', linewidth=1, label='Placebo')
plt.legend()
plt.grid()
plt.title('Survival Rate (by percent)')
plt.ylabel('Survival Percentage')
plt.xlabel('Timepoint (Days)')
# Show the Figure
plt.show()
# -
# ## Summary Bar Graph
# +
# Calculate the percent changes for each drug
tumor_volume = tumor_volume.reset_index(drop=True)
tumor_volume_change = tumor_volume.iloc[-1] - tumor_volume.iloc[0]
tumor_volume_percent_change = (tumor_volume_change / tumor_volume.iloc[0]) * 100
# # Display the data to confirm
percent_change_list = tumor_volume_percent_change.tolist()
percent_change_list_index = tumor_volume_percent_change.index.tolist()
percent_values = []
for i in percent_change_list:
value = round(i, 2)
percent_values.append(str(value)+ '%')
percent_values
# +
# Store all Relevant Percent Changes into a Tuple
colors = ["green", "red", "red", "red"]
x_axis = np.arange(0, 4)
tick_locations = []
for x in x_axis:
tick_locations.append(x)
ax.set_title("Metastatic Change by Treatment")
ax.set_xlabel("Treatment")
ax.set_ylabel("% Tumor Volume Change")
ax.set_xticklabels(percent_values)
ax.set_xlim(-0.75, len(percent_change_list_index)-.25)
ax.set_ylim(-30, 60)
plt.bar(x_axis, percent_change_list, color=colors, align="center")
plt.xticks(tick_locations, percent_change_list_index)
plt.tight_layout()
plt.grid()
# Show the Figure
fig.show()
# -
| Pymaceuticals/pymaceuticals.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction to ipydatawidgets
# This example assumes you have installed ipydatawidgets (and the appropriate frontend extension) according to the README.
#
# We start with some setup code:
from ipydatawidgets import (
NDArray, NDArrayWidget, DataUnion, array_serialization, shape_constraints,
create_constrained_arraywidget
)
from ipywidgets import Widget, widget_serialization
from traitlets import Instance, observe
import numpy as np
np.random.seed(0)
# Let's create some random data to use as an example:
raw_data = 255 * np.random.rand(4, 4, 4)
data_widget = NDArrayWidget(raw_data)
# Create a widget that will use some data
class MyWidget(Widget):
# Add a trait that only accepts raw numpy arrays. Note that we have
# to add the serializers explicitly:
array_only = NDArray(np.zeros(0)).tag(sync=True, **array_serialization)
# Add a trait that only accpets a reference to a data widget:
widget_only = Instance(NDArrayWidget, allow_none=True).tag(
sync=True, **widget_serialization)
# Add a trait that accepts either an array or a data widget. Note that
# this sets default serializers for itself. This is the recommended way
# of adding array traits to widgets.
data_union = DataUnion(np.zeros(0)).tag(sync=True)
# ## Simple use:
#
# With a widget and some example data setup, these are the ways it can be used:
# Since we have set valid default values for all traits, we can do an empty init:
w = MyWidget()
# First, we assign some valid data to the traits that only accept one type:
w.array_only = raw_data
w.array_only.shape
w.widget_only = data_widget
w.widget_only.array.shape
# Then, we try assigning some invalid data:
# + tags=["raises-exception"]
w.widget_only = raw_data
# + tags=["raises-exception"]
w.array_only = data_widget
# -
# Note that array-traits will coerce anything numpy can coerce to an array. Note that for integer inputs on non-Windows platforms, numpy defaults to using 64-bit ints, which will cause a warning, as JavaScript arrays does not support 64-bit integers.
w.array_only = [0., 1., 2., 3.]
w.array_only
# Next, we try to assign some data to the union field:
w.data_union = raw_data
w.data_union.shape
w.data_union = data_widget
w.data_union.array.shape
# ## Constraints:
#
# Now, we will add shape and dtype constraints:
# +
color_image_shape_constraint = shape_constraints(None, None, 3)
ColorImageDataWidget = create_constrained_arraywidget(color_image_shape_constraint, dtype=np.uint8)
# Create a widget that can hold color image data, in various forms:
class ColorImageWidget(Widget):
array_only = NDArray(dtype=np.uint8)\
.tag(sync=True, **array_serialization)\
.valid(color_image_shape_constraint)
widget_only = Instance(ColorImageDataWidget)\
.tag(sync=True, **widget_serialization)
data_union = DataUnion(dtype=np.uint8, shape_constraint=color_image_shape_constraint)\
.tag(sync=True)
# -
color_data = raw_data[:, :, :3].astype(np.uint8)
color_data.shape
color_data_widget = ColorImageDataWidget(array=color_data)
# Initialize with valid data:
wc = ColorImageWidget(
array_only=color_data,
widget_only = color_data_widget,
data_union = color_data_widget
)
# Now, try to set various invalid data, that will either fail, or be coerced:
# + tags=["raises-exception", "nbval-ignore-output"]
wc.array_only = raw_data # Fails, since raw_data has wrong size of last axis
# + tags=["raises-exception", "nbval-ignore-output"]
wc.array_only = raw_data[:, :, 0] # Fails, since data has wrong number of dimensions
# + tags=["nbval-ignore-output"]
# This will coerce the float data to uint8 (this creates a copy of the data)
wc.array_only = raw_data[:, :, :3]
wc.array_only[:2, :2, :] # Preview a few values
# + tags=["raises-exception"]
wc.widget_only = data_widget # Fails, since type of widget is wrong
# + tags=["raises-exception"]
wc.data_union = data_widget # Fails, since it cannot coerce data in a widget reference
# + tags=["raises-exception", "nbval-ignore-output"]
data_widget.array = color_data
wc.data_union = data_widget # Works, even if data_widget itself is unconstrained
data_widget.array = raw_data # Now not allowed, as our DataUnion trait are constraining data_widget
# -
| examples/introduction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/silro/bigvlaue/blob/master/MNIST.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="mfRURrD4TaE-" colab_type="code" colab={}
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
# %matplotlib inline
# + [markdown] id="0eW7T1rhVZY_" colab_type="text"
# #Load datasets
# MNIST
# + id="-WGykFkeVLKv" colab_type="code" colab={}
from tensorflow.keras import datasets
# + id="WeHgV4rrVVle" colab_type="code" colab={}
mnist = datasets.mnist
# + id="xQPjiZDqVgdW" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 53} outputId="ddb6aaf5-979e-4e3d-8eae-c3d7da51de45"
(train_x, train_y), (test_x, test_y) = mnist.load_data()
# + id="6_64eWN1VkrF" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="ae7adad3-dd8c-41e7-fd8e-4ef0b7b4f5ab"
train_x.shape
# + [markdown] id="00LwSUe7Wcnu" colab_type="text"
# #gray는 1차원, RGB는 3차원
# + [markdown] id="p9vANZksVznG" colab_type="text"
# Image Dataset 들여다보기
#
# + id="YqJGf5vcVxys" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="a428b0aa-f04b-4da3-c396-73a58f57cd5c"
image = train_x[0]
image.shape
# + id="fazpjxKhV821" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 265} outputId="4a4d6990-e78f-4b9d-d075-a27d0325dfcd"
plt.imshow(image,'gray')
plt.show()
# + [markdown] id="1oAEuLuxWBrV" colab_type="text"
# #Chaneel 관련
# 차원 수 늘리기(3차원으로 만들어주기 RGB)
# + id="rfAW6yWRWhgk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="86600b2a-36d5-4e63-e817-93457875fab2"
new_train_x = tf.expand_dims(train_x, -1)
new_train_x.shape
# + id="498zBzODWH-X" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="a16d0f30-d188-4832-effe-e1f601af573c"
exp_data = np.expand_dims(train_x, -1)
exp_data.shape
# + id="PTUd3Xa-WS1v" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="b57801ce-9fbf-453c-e8d4-6a2645f91ee8"
train_x[..., tf.newaxis].shape
#TF 공식 홈페이지 제공 방법
# + id="6vCWZFz9WueE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="e4d28654-e862-44d4-b8cb-0eb956498752"
new_train_x = train_x[..., tf.newaxis]
new_train_x.shape
# + id="udueVHIBXFEF" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 265} outputId="8c2cf444-a1ba-470c-c207-1681f6a9b370"
disp = new_train_x[15,:,:,0]
plt.imshow(disp, 'gray')
plt.show()
# + id="DHOrSNF5XUod" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="99aa8de2-be36-414f-89af-4df0a1b7b38e"
disp = np.squeeze(new_train_x[0])
disp.shape
# + [markdown] id="DM7awxBVX6vA" colab_type="text"
# #Label data 보기 (Y)
# + id="Hn_zF6dzX8lH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="527216de-b416-46a4-b2f7-f399271452af"
train_y.shape
# + id="sLtFssIqX9yk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="99810854-9ce7-4249-820d-94642601ac70"
train_y[0], train_x[0]
# + id="LSf3-VRuX-pV" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 279} outputId="08886daa-2b6c-4744-9d7f-37fbdd3f8459"
plt.title(train_y[0],fontsize=11)
plt.imshow(train_x[0],'gray')
plt.show()
# + [markdown] id="8RZsv_gpYikJ" colab_type="text"
# # OneHot Encoding
#
# * 단 하나의 값만 True이고 나머지는 False인 인코딩
# * 컴퓨터가 이해할 수 있는 형태로 변환해서 Label을 주도록 함
# * [0,1,0,0,0,0,0,0,0]
#
# + id="13AONeUOY3l_" colab_type="code" colab={}
#5
[0,0,0,0,0,1,0,0,0]
#9
[0,0,0,0,0,0,0,0,1]
# + id="eCx4_3NXYomP" colab_type="code" colab={}
# + [markdown] id="csUKa-7TZKRK" colab_type="text"
# tensorflow.keras.utils.to_categorical
#
#
# + id="EQujV21UZNsd" colab_type="code" colab={}
from tensorflow.keras.utils import to_categorical
# + id="v-4cs9foZQE5" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="f2e4cbbc-3d8f-4b75-aa50-d96b33705007"
to_categorical(1, 10)
# + id="oBltdOxiZUbu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="8dcab057-601f-4081-8a8e-ed6963034557"
label = train_y[0]
label
# + id="Ux6Q_L6lZhZW" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="5321455f-720c-4abf-e720-7bda8b024715"
label_onehot = to_categorical(label, num_classes=10)
label_onehot
# + id="bn6h4f7xZmmO" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 317} outputId="2a0a5b33-a63f-493d-e4f3-9c4e349b82fd"
plt.title(label_onehot)
plt.imshow(train_x[0],'gray')
plt.show()
# + id="mRE6_ureZwat" colab_type="code" colab={}
| MNIST.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # API quickstart
# +
# %matplotlib inline
import numpy as np
import theano.tensor as tt
import pymc3 as pm
import seaborn as sns
import matplotlib.pyplot as plt
sns.set_context('notebook')
plt.style.use('seaborn-darkgrid')
print('Running on PyMC3 v{}'.format(pm.__version__))
# -
# ## 1. Model creation
#
# Models in PyMC3 are centered around the `Model` class. It has references to all random variables (RVs) and computes the model logp and its gradients. Usually, you would instantiate it as part of a `with` context:
with pm.Model() as model:
# Model definition
pass
# We discuss RVs further below but let's create a simple model to explore the `Model` class.
with pm.Model() as model:
mu = pm.Normal('mu', mu=0, sd=1)
obs = pm.Normal('obs', mu=mu, sd=1, observed=np.random.randn(100))
model.basic_RVs
model.free_RVs
model.observed_RVs
model.logp({'mu': 0})
# **Warning**
# It's worth highlighting one of the counter-intuitive design choices with logp.
# The API makes the `logp` look like an attribute, when it actually puts together a function based on the current state of the model.
#
# The current design is super maintainable, does terrible if the state stays constant, and great if the state keeps changing, for reasons of design we assume that `Model` isn't static, in fact it's best in our experience and avoids bad results.
#
# If you need to use `logp` in an inner loop and it needs to be static, simply use something like `logp = model.logp` below. You can see the caching effect with the speed up below.
# %timeit model.logp({mu: 0.1})
logp = model.logp
# %timeit logp({mu: 0.1})
# ## 2. Probability Distributions
#
# Every probabilistic program consists of observed and unobserved Random Variables (RVs). Observed RVs are defined via likelihood distributions, while unobserved RVs are defined via prior distributions. In PyMC3, probability distributions are available from the main module space:
help(pm.Normal)
# In the PyMC3 module, the structure for probability distributions looks like this:
#
# [pymc3.distributions](../api/distributions.rst)
# - [continuous](../api/distributions/continuous.rst)
# - [discrete](../api/distributions/discrete.rst)
# - [timeseries](../api/distributions/timeseries.rst)
# - [mixture](../api/distributions/mixture.rst)
#
dir(pm.distributions.mixture)
# ### Unobserved Random Variables
# Every unobserved RV has the following calling signature: name (str), parameter keyword arguments. Thus, a normal prior can be defined in a model context like this:
with pm.Model():
x = pm.Normal('x', mu=0, sd=1)
# As with the model, we can evaluate its logp:
x.logp({'x': 0})
# ### Observed Random Variables
# Observed RVs are defined just like unobserved RVs but require data to be passed into the `observed` keyword argument:
with pm.Model():
obs = pm.Normal('x', mu=0, sd=1, observed=np.random.randn(100))
# `observed` supports lists, `numpy.ndarray`, `theano` and `pandas` data structures.
# ### Deterministic transforms
# PyMC3 allows you to freely do algebra with RVs in all kinds of ways:
with pm.Model():
x = pm.Normal('x', mu=0, sd=1)
y = pm.Gamma('y', alpha=1, beta=1)
plus_2 = x + 2
summed = x + y
squared = x**2
sined = pm.math.sin(x)
# While these transformations work seamlessly, their results are not stored automatically. Thus, if you want to keep track of a transformed variable, you have to use `pm.Deterministic`:
with pm.Model():
x = pm.Normal('x', mu=0, sd=1)
plus_2 = pm.Deterministic('x plus 2', x + 2)
# Note that `plus_2` can be used in the identical way to above, we only tell PyMC3 to keep track of this RV for us.
# ### Automatic transforms of bounded RVs
#
# In order to sample models more efficiently, PyMC3 automatically transforms bounded RVs to be unbounded.
with pm.Model() as model:
x = pm.Uniform('x', lower=0, upper=1)
# When we look at the RVs of the model, we would expect to find `x` there, however:
model.free_RVs
# `x_interval__` represents `x` transformed to accept parameter values between -inf and +inf. In the case of an upper and a lower bound, a `LogOdd`s transform is applied. Sampling in this transformed space makes it easier for the sampler. PyMC3 also keeps track of the non-transformed, bounded parameters. These are common determinstics (see above):
model.deterministics
# When displaying results, PyMC3 will usually hide transformed parameters. You can pass the `include_transformed=True` parameter to many functions to see the transformed parameters that are used for sampling.
#
# You can also turn transforms off:
# +
with pm.Model() as model:
x = pm.Uniform('x', lower=0, upper=1, transform=None)
print(model.free_RVs)
# -
# Or specify different transformation other than the default:
# +
import pymc3.distributions.transforms as tr
with pm.Model() as model:
# use the default log transformation
x1 = pm.Gamma('x1', alpha=1, beta=1)
# sepcified a different transformation
x2 = pm.Gamma('x2', alpha=1, beta=1, transform=tr.log_exp_m1)
print('The default transformation of x1 is: ' + x1.transformation.name)
print('The user specified transformation of x2 is: ' + x2.transformation.name)
# -
# ### Transformed distributions and changes of variables
# PyMC3 does not provide explicit functionality to transform one distribution to another. Instead, a dedicated distribution is usually created in consideration of optimising performance. However, users can still create transformed distribution by passing the inverse transformation to `transform` kwarg. Take the classical textbook example of LogNormal: $log(y) \sim \text{Normal}(\mu, \sigma)$
# +
class Exp(tr.ElemwiseTransform):
name = "exp"
def backward(self, x):
return tt.log(x)
def forward(self, x):
return tt.exp(x)
def jacobian_det(self, x):
return -tt.log(x)
with pm.Model() as model:
x1 = pm.Normal('x1', 0., 1., transform=Exp())
x2 = pm.Lognormal('x2', 0., 1.)
lognorm1 = model.named_vars['x1_exp__']
lognorm2 = model.named_vars['x2']
_, ax = plt.subplots(1, 1, figsize=(5, 3))
x = np.linspace(0., 10., 100)
ax.plot(
x,
np.exp(lognorm1.distribution.logp(x).eval()),
'--',
alpha=.5,
label='log(y) ~ Normal(0, 1)')
ax.plot(
x,
np.exp(lognorm2.distribution.logp(x).eval()),
alpha=.5,
label='y ~ Lognormal(0, 1)')
plt.legend();
# -
# Notice from above that the named variable `x1_exp__` in the `model` is Lognormal distributed.
# Using similar approach, we can create ordered RVs following some distribution. For example, we can combine the `ordered` transformation and `logodds` transformation using `Chain` to create a 2D RV that satisfy $x_1, x_2 \sim \text{Uniform}(0, 1) \space and \space x_1< x_2$
# +
Order = tr.Ordered()
Logodd = tr.LogOdds()
chain_tran = tr.Chain([Logodd, Order])
with pm.Model() as m0:
x = pm.Uniform(
'x', 0., 1., shape=2,
transform=chain_tran,
testval=[0.1, 0.9])
trace = pm.sample(5000, tune=1000, progressbar=False)
_, ax = plt.subplots(1, 2, figsize=(10, 5))
for ivar, varname in enumerate(trace.varnames):
ax[ivar].scatter(trace[varname][:, 0], trace[varname][:, 1], alpha=.01)
ax[ivar].set_xlabel(varname + '[0]')
ax[ivar].set_ylabel(varname + '[1]')
ax[ivar].set_title(varname)
plt.tight_layout()
# -
# ### Lists of RVs / higher-dimensional RVs
#
# Above we have seen how to create scalar RVs. In many models, you want multiple RVs. There is a tendency (mainly inherited from PyMC 2.x) to create list of RVs, like this:
with pm.Model():
x = [pm.Normal('x_{}'.format(i), mu=0, sd=1) for i in range(10)] # bad
# However, even though this works it is quite slow and not recommended. Instead, use the `shape` kwarg:
with pm.Model() as model:
x = pm.Normal('x', mu=0, sd=1, shape=10) # good
# `x` is now a random vector of length 10. We can index into it or do linear algebra operations on it:
with model:
y = x[0] * x[1] # full indexing is supported
x.dot(x.T) # Linear algebra is supported
# ### Initialization with test_values
#
# While PyMC3 tries to automatically initialize models it is sometimes helpful to define initial values for RVs. This can be done via the `testval` kwarg:
# +
with pm.Model():
x = pm.Normal('x', mu=0, sd=1, shape=5)
x.tag.test_value
# +
with pm.Model():
x = pm.Normal('x', mu=0, sd=1, shape=5, testval=np.random.randn(5))
x.tag.test_value
# -
# This technique is quite useful to identify problems with model specification or initialization.
# ## 3. Inference
#
# Once we have defined our model, we have to perform inference to approximate the posterior distribution. PyMC3 supports two broad classes of inference: sampling and variational inference.
#
# ### 3.1 Sampling
#
# The main entry point to MCMC sampling algorithms is via the `pm.sample()` function. By default, this function tries to auto-assign the right sampler(s) and auto-initialize if you don't pass anything.
with pm.Model() as model:
mu = pm.Normal('mu', mu=0, sd=1)
obs = pm.Normal('obs', mu=mu, sd=1, observed=np.random.randn(100))
trace = pm.sample(1000, tune=500)
# As you can see, on a continuous model, PyMC3 assigns the NUTS sampler, which is very efficient even for complex models. PyMC3 also runs variational inference (i.e. ADVI) to find good starting parameters for the sampler. Here we draw 1000 samples from the posterior and allow the sampler to adjust its parameters in an additional 500 iterations. These 500 samples are discarded by default:
len(trace)
# You can also run multiple chains in parallel using the `cores` kwarg:
with pm.Model() as model:
mu = pm.Normal('mu', mu=0, sd=1)
obs = pm.Normal('obs', mu=mu, sd=1, observed=np.random.randn(100))
trace = pm.sample(cores=4)
# Note, that we are now drawing 2000 samples, 500 samples for 4 chains each. The 500 tuning samples are discarded by default.
trace['mu'].shape
trace.nchains
trace.get_values('mu', chains=1).shape # get values of a single chain
# PyMC3, offers a variety of other samplers, found in `pm.step_methods`.
list(filter(lambda x: x[0].isupper(), dir(pm.step_methods)))
# Commonly used step-methods besides NUTS are `Metropolis` and `Slice`. **For almost all continuous models, `NUTS` should be preferred.** There are hard-to-sample models for which `NUTS` will be very slow causing many users to use `Metropolis` instead. This practice, however, is rarely successful. NUTS is fast on simple models but can be slow if the model is very complex or it is badly initialized. In the case of a complex model that is hard for NUTS, Metropolis, while faster, will have a very low effective sample size or not converge properly at all. A better approach is to instead try to improve initialization of NUTS, or reparameterize the model.
#
# For completeness, other sampling methods can be passed to sample:
with pm.Model() as model:
mu = pm.Normal('mu', mu=0, sd=1)
obs = pm.Normal('obs', mu=mu, sd=1, observed=np.random.randn(100))
step = pm.Metropolis()
trace = pm.sample(1000, step=step)
# You can also assign variables to different step methods.
with pm.Model() as model:
mu = pm.Normal('mu', mu=0, sd=1)
sd = pm.HalfNormal('sd', sd=1)
obs = pm.Normal('obs', mu=mu, sd=sd, observed=np.random.randn(100))
step1 = pm.Metropolis(vars=[mu])
step2 = pm.Slice(vars=[sd])
trace = pm.sample(10000, step=[step1, step2], cores=4)
# ### 3.2 Analyze sampling results
#
# The most common used plot to analyze sampling results is the so-called trace-plot:
pm.traceplot(trace);
# Another common metric to look at is R-hat, also known as the Gelman-Rubin statistic:
pm.gelman_rubin(trace)
# These are also part of the `forestplot`:
pm.forestplot(trace);
# Finally, for a plot of the posterior that is inspired by the book [Doing Bayesian Data Analysis](http://www.indiana.edu/~kruschke/DoingBayesianDataAnalysis/), you can use the:
pm.plot_posterior(trace);
# For high-dimensional models it becomes cumbersome to look at all parameter's traces. When using `NUTS` we can look at the energy plot to assess problems of convergence:
# +
with pm.Model() as model:
x = pm.Normal('x', mu=0, sd=1, shape=100)
trace = pm.sample(cores=4)
pm.energyplot(trace);
# -
# For more information on sampler stats and the energy plot, see [here](sampler-stats.ipynb). For more information on identifying sampling problems and what to do about them, see [here](Diagnosing_biased_Inference_with_Divergences.ipynb).
# ### 3.3 Variational inference
#
# PyMC3 supports various Variational Inference techniques. While these methods are much faster, they are often also less accurate and can lead to biased inference. The main entry point is `pymc3.fit()`.
with pm.Model() as model:
mu = pm.Normal('mu', mu=0, sd=1)
sd = pm.HalfNormal('sd', sd=1)
obs = pm.Normal('obs', mu=mu, sd=sd, observed=np.random.randn(100))
approx = pm.fit()
# The returned `Approximation` object has various capabilities, like drawing samples from the approximated posterior, which we can analyse like a regular sampling run:
approx.sample(500)
# The `variational` submodule offers a lot of flexibility in which VI to use and follows an object oriented design. For example, full-rank ADVI estimates a full covariance matrix:
mu = pm.floatX([0., 0.])
cov = pm.floatX([[1, .5], [.5, 1.]])
with pm.Model() as model:
pm.MvNormal('x', mu=mu, cov=cov, shape=2)
approx = pm.fit(method='fullrank_advi')
# An equivalent expression using the object-oriented interface is:
with pm.Model() as model:
pm.MvNormal('x', mu=mu, cov=cov, shape=2)
approx = pm.FullRankADVI().fit()
plt.figure()
trace = approx.sample(10000)
sns.kdeplot(trace['x'][:, 0], trace['x'][:, 1]);
# Stein Variational Gradient Descent (SVGD) uses particles to estimate the posterior:
w = pm.floatX([.2, .8])
mu = pm.floatX([-.3, .5])
sd = pm.floatX([.1, .1])
with pm.Model() as model:
pm.NormalMixture('x', w=w, mu=mu, sd=sd)
approx = pm.fit(method=pm.SVGD(n_particles=200, jitter=1.))
plt.figure()
trace = approx.sample(10000)
sns.distplot(trace['x']);
# For more information on variational inference, see [these examples](http://pymc-devs.github.io/pymc3/examples.html#variational-inference).
# ## 4. Posterior Predictive Sampling
#
# The `sample_ppc()` function performs prediction on hold-out data and posterior predictive checks.
data = np.random.randn(100)
with pm.Model() as model:
mu = pm.Normal('mu', mu=0, sd=1)
sd = pm.HalfNormal('sd', sd=1)
obs = pm.Normal('obs', mu=mu, sd=sd, observed=data)
trace = pm.sample()
with model:
post_pred = pm.sample_ppc(trace, samples=500, size=len(data))
# `sample_ppc()` returns a dict with a key for every observed node:
post_pred['obs'].shape
plt.figure()
ax = sns.distplot(post_pred['obs'].mean(axis=1), label='Posterior predictive means')
ax.axvline(data.mean(), color='r', ls='--', label='True mean')
ax.legend();
# ## 4.1 Predicting on hold-out data
#
# In many cases you want to predict on unseen / hold-out data. This is especially relevant in Probabilistic Machine Learning and Bayesian Deep Learning. While we plan to improve the API in this regard, this can currently be achieved with a `theano.shared` variable. These are theano tensors whose values can be changed later. Otherwise they can be passed into PyMC3 just like any other numpy array or tensor.
#
# This distinction is significant since internally all models in PyMC3 are giant symbolic expressions. When you pass data directly into a model, you are giving Theano permission to treat this data as a constant and optimize it away as it sees fit. If you need to change this data later you might not have a way to point at it in the symbolic expression. Using `theano.shared` offers a way to point to a place in that symbolic expression, and change what is there.
# +
import theano
x = np.random.randn(100)
y = x > 0
x_shared = theano.shared(x)
y_shared = theano.shared(y)
with pm.Model() as model:
coeff = pm.Normal('x', mu=0, sd=1)
logistic = pm.math.sigmoid(coeff * x_shared)
pm.Bernoulli('obs', p=logistic, observed=y_shared)
trace = pm.sample()
# -
# Now assume we want to predict on unseen data. For this we have to change the values of `x_shared` and `y_shared`. Theoretically we don't need to set `y_shared` as we want to predict it but it has to match the shape of `x_shared`.
# +
x_shared.set_value([-1, 0, 1.])
y_shared.set_value([0, 0, 0]) # dummy values
with model:
post_pred = pm.sample_ppc(trace, samples=500)
# -
post_pred['obs'].mean(axis=0)
| docs/source/notebooks/api_quickstart.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import tellurium as te
import numpy as np
import lmfit # Fitting lib
import math
import random
import matplotlib.pyplot as plt
# +
rr = te.loada("""
A -> B + C; k1*A
A = 5;
B = 0;
C = 0;
k1 = 0.15
""")
timeToSimulate = 30
nDataPoints = 10
# +
# Create the experimental data
# Create some 'experimental' data
x_data = data[:, 0]
y_data = data[:,1]
for i in range (nDataPoints):
y_data[i] = max(y_data[i] + np.random.normal (0, 0.5), 0) # standard deviation of noise
# Plot it to see what it looks like
plt.plot (x_data, y_data, marker='*', linestyle='None')
plt.show()
# -
# # Running a minimization
# 1. Define parameters present
# 1. Define python function that calculates residuals given parameter values
# 1. Construct the minimizer
# 1. Run the minimizer
# Define the parameters present
parameters = lmfit.Parameters()
parameters.add('k1', value=1, min=0, max=10)
# Define a python function that computes residuals from parameter values
def residuals(p):
rr.reset()
pp = p.valuesdict()
rr.k1 = pp['k1']
data = rr.simulate (0, timeToSimulate, nDataPoints, ['A'])
return (y_data - data[:,0])
# Create the minimizer
fitter = lmfit.Minimizer(residuals, parameters)
# +
# Run the minimizer and examine the results
result = fitter.minimize (method='leastsq')
print (result.message)
# You can also print
print (result.chisqr)
print (result.params)
# -
# ## Exercises
# 1. Run the fit 5 times and compute the standard deviation of the parameter estimate. How does it compare with the +/- values reported by the minimizer?
# 1. Add the reaction B -> C; k2\*B. Redo the fit.
import numpy as np
np.std([.169, .109, .139, .147, .162])
| archived_lectures/Fall_2019/lecture_7/Lecture_7_Model_Fitting.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <!--BOOK_INFORMATION-->
# <img align="left" style="padding-right:10px;" src="fig/cover-small.jpg">
# *This notebook contains an excerpt from the [Whirlwind Tour of Python](http://www.oreilly.com/programming/free/a-whirlwind-tour-of-python.csp) by <NAME>; the content is available [on GitHub](https://github.com/jakevdp/WhirlwindTourOfPython).*
#
# *The text and code are released under the [CC0](https://github.com/jakevdp/WhirlwindTourOfPython/blob/master/LICENSE) license; see also the companion project, the [Python Data Science Handbook](https://github.com/jakevdp/PythonDataScienceHandbook).*
#
# <!--NAVIGATION-->
# < [Errors and Exceptions](09-Errors-and-Exceptions.ipynb) | [Contents](Index.ipynb) | [List Comprehensions](11-List-Comprehensions.ipynb) >
# # Iterators
# Often an important piece of data analysis is repeating a similar calculation, over and over, in an automated fashion.
# For example, you may have a table of names that you'd like to split into first and last, or perhaps of dates that you'd like to convert to some standard format.
# One of Python's answers to this is the *iterator* syntax.
# We've seen this already with the ``range`` iterator:
for i in range(10):
print(i, end=' ')
# Here we're going to dig a bit deeper.
# It turns out that in Python 3, ``range`` is not a list, but is something called an *iterator*, and learning how it works is key to understanding a wide class of very useful Python functionality.
# ## Iterating over lists
# Iterators are perhaps most easily understood in the concrete case of iterating through a list.
# Consider the following:
for value in [2, 4, 6, 8, 10]:
# do some operation
print(value + 1, end=' ')
# The familiar "``for x in y``" syntax allows us to repeat some operation for each value in the list.
# The fact that the syntax of the code is so close to its English description ("*for [each] value in [the] list*") is just one of the syntactic choices that makes Python such an intuitive language to learn and use.
#
# But the face-value behavior is not what's *really* happening.
# When you write something like "``for val in L``", the Python interpreter checks whether it has an *iterator* interface, which you can check yourself with the built-in ``iter`` function:
iter([2, 4, 6, 8, 10])
# It is this iterator object that provides the functionality required by the ``for`` loop.
# The ``iter`` object is a container that gives you access to the next object for as long as it's valid, which can be seen with the built-in function ``next``:
I = iter([2, 4, 6, 8, 10])
print(next(I))
print(next(I))
print(next(I))
# What is the purpose of this level of indirection?
# Well, it turns out this is incredibly useful, because it allows Python to treat things as lists that are *not actually lists*.
# ## ``range()``: A List Is Not Always a List
# Perhaps the most common example of this indirect iteration is the ``range()`` function in Python 3 (named ``xrange()`` in Python 2), which returns not a list, but a special ``range()`` object:
range(10)
# ``range``, like a list, exposes an iterator:
iter(range(10))
# So Python knows to treat it *as if* it's a list:
for i in range(10):
print(i, end=' ')
# The benefit of the iterator indirection is that *the full list is never explicitly created!*
# We can see this by doing a range calculation that would overwhelm our system memory if we actually instantiated it (note that in Python 2, ``range`` creates a list, so running the following will not lead to good things!):
N = 10 ** 12
for i in range(N):
if i >= 10: break
print(i, end=', ')
# If ``range`` were to actually create that list of one trillion values, it would occupy tens of terabytes of machine memory: a waste, given the fact that we're ignoring all but the first 10 values!
#
# In fact, there's no reason that iterators ever have to end at all!
# Python's ``itertools`` library contains a ``count`` function that acts as an infinite range:
# +
from itertools import count
for i in count():
if i >= 10:
break
print(i, end=', ')
# -
# Had we not thrown-in a loop break here, it would go on happily counting until the process is manually interrupted or killed (using, for example, ``ctrl-C``).
# ## Useful Iterators
# This iterator syntax is used nearly universally in Python built-in types as well as the more data science-specific objects we'll explore in later sections.
# Here we'll cover some of the more useful iterators in the Python language:
# ### ``enumerate``
# Often you need to iterate not only the values in an array, but also keep track of the index.
# You might be tempted to do things this way:
L = [2, 4, 6, 8, 10]
for i in range(len(L)):
print(i, L[i])
# Although this does work, Python provides a cleaner syntax using the ``enumerate`` iterator:
for i, val in enumerate(L):
print(i, val)
# This is the more "Pythonic" way to enumerate the indices and values in a list.
# ### ``zip``
# Other times, you may have multiple lists that you want to iterate over simultaneously.
# You could certainly iterate over the index as in the non-Pythonic example we looked at previously, but it is better to use the ``zip`` iterator, which zips together iterables:
L = [2, 4, 6, 8, 10]
R = [3, 6, 9, 12, 15]
for lval, rval in zip(L, R):
print(lval, rval)
# Any number of iterables can be zipped together, and if they are different lengths, the shortest will determine the length of the ``zip``.
# ### ``map`` and ``filter``
# The ``map`` iterator takes a function and applies it to the values in an iterator:
# find the first 10 square numbers
square = lambda x: x ** 2
for val in map(square, range(10)):
print(val, end=' ')
# The ``filter`` iterator looks similar, except it only passes-through values for which the filter function evaluates to True:
# find values up to 10 for which x % 2 is zero
is_even = lambda x: x % 2 == 0
for val in filter(is_even, range(10)):
print(val, end=' ')
# The ``map`` and ``filter`` functions, along with the ``reduce`` function (which lives in Python's ``functools`` module) are fundamental components of the *functional programming* style, which, while not a dominant programming style in the Python world, has its outspoken proponents (see, for example, the [pytoolz](https://toolz.readthedocs.org/en/latest/) library).
# ### Iterators as function arguments
#
# We saw in [``*args`` and ``**kwargs``: Flexible Arguments](#*args-and-**kwargs:-Flexible-Arguments). that ``*args`` and ``**kwargs`` can be used to pass sequences and dictionaries to functions.
# It turns out that the ``*args`` syntax works not just with sequences, but with any iterator:
print(*range(10))
# So, for example, we can get tricky and compress the ``map`` example from before into the following:
print(*map(lambda x: x ** 2, range(10)))
# Using this trick lets us answer the age-old question that comes up in Python learners' forums: why is there no ``unzip()`` function which does the opposite of ``zip()``?
# If you lock yourself in a dark closet and think about it for a while, you might realize that the opposite of ``zip()`` is... ``zip()``! The key is that ``zip()`` can zip-together any number of iterators or sequences. Observe:
L1 = (1, 2, 3, 4)
L2 = ('a', 'b', 'c', 'd')
z = zip(L1, L2)
print(*z)
# zip objects are iterators and iterators can be only used once. If we were to print `z` again it would be an empty iterator:
print(*z)
# ## Specialized Iterators: ``itertools``
#
# We briefly looked at the infinite ``range`` iterator, ``itertools.count``.
# The ``itertools`` module contains a whole host of useful iterators; it's well worth your while to explore the module to see what's available.
# As an example, consider the ``itertools.permutations`` function, which iterates over all permutations of a sequence:
from itertools import permutations
p = permutations(range(3))
print(*p)
# Similarly, the ``itertools.combinations`` function iterates over all unique combinations of ``N`` values within a list:
from itertools import combinations
c = combinations(range(4), 2)
print(*c)
# Somewhat related is the ``product`` iterator, which iterates over all sets of pairs between two or more iterables:
from itertools import product
p = product('ab', range(3))
print(*p)
# Many more useful iterators exist in ``itertools``: the full list can be found, along with some examples, in Python's [online documentation](https://docs.python.org/3.5/library/itertools.html).
#
# <!--NAVIGATION-->
# < [Errors and Exceptions](09-Errors-and-Exceptions.ipynb) | [Contents](Index.ipynb) | [List Comprehensions](11-List-Comprehensions.ipynb) >
| Lectures/11-Iterators.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
df_missing = pd.read_excel("../datasets/Sample - Superstore.xls",sheet_name="Missing")
df_missing.head()
# -
df_missing.fillna('FILL')
df_missing[['Customer','Product']].fillna('FILL')
df_missing['Sales'].fillna(method='ffill')
df_missing['Sales'].fillna(method='bfill')
df_missing['Sales'].fillna(df_missing.mean()['Sales'])
| Exercise05/Exercise05.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
from collections import namedtuple
import pathlib
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
import numpy as np
import pandas as pd
import scipy
import toolz.curried as toolz
# -
import orchid
# project_names = ('bakken', 'permian', 'montney')
# project_filenames = ('frankNstein_Bakken_UTM13_FEET.ifrac',
# 'Project_frankNstein_Permian_UTM13_FEET.ifrac',
# 'Project-frankNstein_Montney_UTM13_METERS.ifrac')
project_names = ('bakken', 'montney')
project_filenames = ('frankNstein_Bakken_UTM13_FEET.ifrac',
'Project-frankNstein_Montney_UTM13_METERS.ifrac')
projects = toolz.pipe(
dict(zip(project_names, project_filenames)),
toolz.valmap(lambda fn: str(pathlib.Path(r'c:\src\Orchid.IntegrationTestData').joinpath(fn))),
toolz.valmap(toolz.curry(orchid.load_project)))
projects
# bakken_project, permian_project, montney_project = toolz.map(lambda pn: projects[pn], projects)
bakken_project, montney_project = toolz.map(lambda pn: projects[pn], projects)
wells = toolz.pipe(projects,
toolz.valmap(lambda project: project.wells()),
toolz.valmap(lambda well_map: [(w.name, w) for w in well_map]),
toolz.valmap(dict))
wells
def stages(project_name, well_name):
return toolz.pipe(wells,
toolz.get_in([project_name, well_name]),
lambda w: w.stages(),
toolz.map(lambda stage: (stage.display_stage_number, stage)),
dict)
StageDetails = namedtuple('StageDetails', ['stage_number', 'display_name_with_well', 'md_top', 'md_bottom', 'cluster_count'])
def stage_details(project_name, stage_list, stage_number):
project = projects[project_name]
length_unit = project.project_units.LENGTH
stage = stage_list[stage_number]
return StageDetails(stage_number, stage.display_name_with_well,
stage.md_top(length_unit), stage.md_bottom(length_unit), stage.cluster_count)
bakken_demo_1h_stage_details = toolz.partial(stage_details, 'bakken', stages('bakken', 'Demo_1H'))
bakken_demo_2h_stage_details = toolz.partial(stage_details, 'bakken', stages('bakken', 'Demo_2H'))
bakken_demo_3h_stage_details = toolz.partial(stage_details, 'bakken', stages('bakken', 'Demo_3H'))
bakken_demo_4h_stage_details = toolz.partial(stage_details, 'bakken', stages('bakken', 'Demo_4H'))
def print_stage_details(details_func, indices):
for n in indices:
print(details_func(n))
def print_bakken_stages_details():
for header, details_func, indices in [('Bakken Demo_1H', bakken_demo_1h_stage_details, [1, 50, 9, 33]),
('Bakken Demo_2H', bakken_demo_2h_stage_details, [1, 50, 21, 8]),
('Bakken Demo_3H', bakken_demo_3h_stage_details, []), # no stages for Demo_3H
('Bakken Demo_4H', bakken_demo_4h_stage_details, [1, 35, 7, 26]),
]:
print(f'\n{header}')
print_stage_details(details_func, indices)
print_bakken_stages_details()
montney_hori_01_stage_details = toolz.partial(stage_details, 'montney', stages('montney', 'Hori_01'))
montney_hori_02_stage_details = toolz.partial(stage_details, 'montney', stages('montney', 'Hori_02'))
montney_hori_03_stage_details = toolz.partial(stage_details, 'montney', stages('montney', 'Hori_03'))
montney_vert_01_stage_details = toolz.partial(stage_details, 'montney', stages('montney', 'Vert_01'))
def print_montney_stages_details():
for header, details_func, indices in [('Montney Hori_01', montney_hori_01_stage_details, [1, 15, 8, 2]),
('Montney Hori_02', montney_hori_02_stage_details, [1, 29, 8, 14]),
('Montney Hori_03', montney_hori_03_stage_details, [1, 28, 9, 20]),
('Montney Vert_01', montney_vert_01_stage_details, [1, 2, 3, 4]),
]:
print(f'\n{header}')
print_stage_details(details_func, indices)
print_montney_stages_details()
| features/notebooks/explore.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
import gym
import numpy as np
from collections import defaultdict
import pdb
import matplotlib.pyplot as plt
from Environment import *
# +
Map = np.array([['s','h','h','h','h','h','h'],
['f','h','h','h','h','h','h'],
['f','h','h','h','h','h','h'],
['f','h','h','h','h','h','h'],
['f','h','h','h','h','h','g'],
['f','h','h','h','h','h','f'],
['f','f','f','f','f','f','f']])
env = Environment(Map,False)
# -
Q_values = defaultdict(lambda: np.random.randint(0,4,4).astype(np.float32))
# +
action_to_num = {
'u':0,
'd':1,
'r':2,
'l':3
}
num_to_action = {
'0':'u',
'1':'d',
'2':'r',
'3':'l'
}
# -
def QLearning(env,Q_values,num_iter,num_episodes,alpha, gamma=0.9):
for it in range(num_iter):
state = env.reset()
for eps in range(num_episodes):
if np.random.rand() < 0.85:
action = num_to_action[str(np.argmax(Q_values[state]))]
new_state,reward,flag = env.step(action)
else:
action = num_to_action[str(np.random.randint(0,4))]
new_state,reward,flag = env.step(action)
Q_values[(state[0],state[1])][action_to_num[action]] += alpha*(reward+gamma*max(Q_values[(env.current_position[0],env.current_position[1])])-Q_values[(state[0],state[1])][action_to_num[action]])
state = new_state
if env.getState() == 'g':
break
return Q_values
Q_star = QLearning(env,Q_values,50000,200,0.9,0.9)
def QvalueToPolicy(Q,Map):
policy = np.empty(Map.shape,dtype=object)
for i in range(Map.shape[0]):
for j in range(Map.shape[1]):
policy[i,j] = num_to_action[str(np.argmax(Q[(i,j)]))]
return policy
policy = QvalueToPolicy(Q_star,Map)
policy
| Temporal difference learning/.ipynb_checkpoints/Q_learning-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # IA Notebook #3 - Programando una Red Neuronal desde Cero
# El problema a resolver está planteado en el [este enlace](https://youtu.be/uwbHOpp9xkc?t=500), básicamente, lo que se quiere es separar en dos grupos de puntos un conjunto de puntos. Un problema de clasificación.
#
# Para ello, primero haremos nuestro datasets y luego la red neuronal como tal. ;3
# +
import numpy as np
import scipy as sc
import matplotlib.pyplot as plt
from sklearn.datasets import make_circles
# +
#************Creamos el dataset************
#Número de registos que tenemos en nuestros datos:
n=500
#Número de características que tenemos sobre cada uno de nuestros datos:
p=2
#Para este ejercicio que se está desarrollando a modo de ejemplo, solo trabajamos con dos características
#para poder dibujarlas en un plano 2D, a mayor número de características, mayor las dimensiones.
#Para consultar los parametros de la función hacemos:
# #make_circle?
#Nuestro dataset estará conformada por una entrada "x" y una salida "y":
x, y = make_circles(n_samples=n, noise=0.05, factor=0.5)
# Aquí corregimos la variable "y" que viene a ser de la forma (500, )
y = y[:, np.newaxis] #Y ahora lo convertimos en un vector como tal -> (500, 1)
#print(y.shape) #Para verificarlo podemos ejecutar esto.
#Para visualizar como van quedando los datos, dibujamos:
#plt.scatter(x[:, 0], x[:, 1], c="orange")
#plt.scatter(x[:, 1], x[:, 0], c="red") #Solo lo puse a modo de prueba :3
#plt.show()
#Pero continuando con el ejercicio: (Y haciendo la correción de la "y")
#plt.scatter(x[y==0, 0], x[y==0, 1], c="red") #Con esto le decimos que solo dibuje uno de los circulos
plt.scatter(x[y[:, 0]==0, 0], x[y[:, 0]==0, 1], c="red")
#plt.scatter(x[y==1, 0], x[y==1, 1], c="orange")
plt.scatter(x[y[:, 0]==1, 0], x[y[:, 0]==1, 1], c="orange")
plt.axis("equal") #Con esto le decimos que esten en la misma proporción ambos ejes
plt.show()
# -
# ---
# ##### Otra vez, haciendo aquí unas pruebas:
#print(y[:, 0])
#print(y)
plt.scatter(x[:, 0], y+0.25, c="orange")
plt.scatter(x[:, 1], y-0.25, c="gray")
plt.show()
# ---
# ## Continuando...
# Ahora si, a programar la red neuronal! :D
#***********Estructura de datos que contiene nuestros parámetros de la red neuronal***********
#No es necesario heredar de la clase "object" porque ya lo hace automáticamente:
class neural_layer(): #Clase de la capa de la red neuronal
#Aquí podría inicializar los atributos:
n_conn=n_neur=act_f=0.0
b=w=np.zeros(1) #Definiendo a "b" y "w" como matrices o vectores unitarios con valor 0
#Y podría usar este constructor:
def __init__(self, numConn, numNeur, funcAct): #El cual SI FUNCIONA!
#def __init__(self, n_conn, n_neur, act_f):
#n_conn: número de conexiones que vienen de una capa anterior.
#n_neur: número de neuronas en nuestra capa
#act_f: función de activación en nuestra capa
self.act_f = funcAct
#self.act_f = act_f #Pero hacer las cosas de esta manera es más sencillo
self.n_conn=numConn #Sin embargo, así creo que es más formal.
self.n_neur=numNeur #Y lo es, pero Python es tan versátil que ni es necesario hacerlo.
#Así que estás líneas solo fueron opcinoales, podría hacerlo mucho más directo.
#Claro, solo con Python, otros lenguajes son más estrictos.
#Vector de vallas:
self.b = np.random.rand(1, numNeur)*2 -1
#self.b = np.random.rand(1, n_neur)*2 -1 #Esto es solo por si acaso falla.
#rand varía de 0 a 1 por eso se multiplica y resta para trabajar con red normalizada y estandarizada.
#Ya que estos serán los parámetros con los que inicializará la red neuronal
#(No lo apunté pero...) Este sería el vector de pesos:
self.w = np.random.rand(numConn, numNeur)*2 -1
#self.w = np.random.rand(numConn, numNeur)*2 -1 #Esto es solo por si acaso falla.
# +
# Funciones de activación (Si uno quiere puede definir más para hacer las pruebas)
sigmoide = (#Función sigmoide
lambda x: 1/(1+np.e**(-x)),
#Derivada de la función sigmoide
lambda x: x*(1-x))
relu = lambda x: np.maximum(0, x)
#Vector o matriz con 100 valores de -5 a 5
_x=np.linspace(-5, 5, 100)
#plt.plot(_x, relu(_x), c="red")
plt.plot(_x, sigmoide[0](_x), c="skyblue")
#plt.plot(_x, sigmoide[1](_x), c="blue")
plt.show()
# +
#Se podría crear capa por capa de manera manual:
#Capa 0
#l0 = neural_layer(p, 4, sigmoide)
#El primer parámetro es "p" porque es la cantidad de características que tenemos sobre nuestros datos.
#Capa 1
#l1 = neural_layer(4, 8, sigmoide)
#La cantidad es ahora 4 porque el número de neuronas de la anterior capa es igual
#Y el número de conexiones debe ser mínimo a esa cantidad.
#---------------------------------------------------------
#O hacerlo iterativo y para ello:
#Creamos este vector o lista o matriz, que definirá el número de neuronas por cada capa:
#topologia = [p, 4, 8, 16, 8, 4, 1] #El número ha sido arbitrario para este caso.
#Y se comentá aquí solo para mantener una estética.
#No necesariamente todas las capas deben tener la misma función de activación
#Para este ejemplo si, por simplicidad:
def crear_redNeuronal(topology, activation_function): #O "create_nn" en inglés.
#red_neuronal -> rn o en inglés: neural_network -> nn
nn=[]
#Ahora un bucle para recorrer todos lo valores y crear la red:
for indiceCapa, capa in enumerate(topology[:-1]): #Con el -1 le digo que recorra todo menos el último.
nn.append(neural_layer(topology[indiceCapa], topology[indiceCapa+1], activation_function))
return nn
#Aquí lo colocamos más ordenado: (pero nuevamente lo movemos y ahora al siguiente segmento)
#topologia = [p, 4, 8, 16, 8, 4, 1]
#Y creamos red neuronal:
#crear_redNeuronal(topologia, sigmoide)
# +
#Se define la topología:
#topologia = [p, 4, 8, 16, 8, 4, 1]
#Creamos la red neuronal:
#red_neuronal=crear_redNeuronal(topologia, sigmoide) #red_neuronal -> rn
#Todo esto anterior se usó para probar la función "entrenar".
#Definimos dos funciones de coste (vamos a usar el error cuadrático medio)
#La primera será la función como tal y la segunda será su derivada:
i2_cost = (
#Error cuadrático medio:
lambda Ypredicha, Yreal: np.mean((Ypredicha-Yreal)**2), #Opera y luego cálcula la media.
#Derivada del error cuadrático medio:
lambda Ypredicha, Yreal: Ypredicha-Yreal
)
#Recordar que el "lr" es el largo del paso que damos en el "Descenso del gradiente"
def entrenar(rn, _x, _y, func_Coste, lr=0.5, train=True):
#En este vector guardaremos esta información:
#out = [(z0, a0), (z1, a1), etc]
#Esta información representa lo procesado en cada capa.
out = [(None, _x)] #Es el caso inicial, ya que en la primera capa no hay iteración.
#Forward pass, básicamente ir ejecutando capa por capa nuestra red, pasando el vector de entrada
for indiceL, L in enumerate(rn):
#Y recordar que el hacer esto, es aplicar una suma ponderada seguida de la función de activación
#Dicha suma ponderada, la llamaremos "z":
z = out[-1][1] @ rn[indiceL].w + rn[indiceL].b
#z = _x @ rn[0].w + rn[0].b #Así sería de manera individual con "_x".
#z = out[0][1] @ rn[0].w + rn[0].b #Y así con "out".
#Es un producto matricial: "_x" @ "w"
#Activación:
a = rn[indiceL].act_f[0](z) #Recordar que nuestra act_f es una tupla con dos valores
#a = rn[0].act_f[0](z) #Así sería de manera individual.
#Todo lo guardamos en out:
out.append((z, a))
#print(out)
#print("---------------------------------------------")
#print(out[-1][1]) #Con esto vemos el primera elemento del úlitmo resultado
#y dicho elemento es justamente la última matriz generada ya con todos los datos.
#print(func_Coste[0](out[-1][1], _y)) #Con esto podemos visualizar cuanto es el error promedio.
if train: #Esto parte si es el entrenamiento como tal, lo anterior es solo para obtener un resultado.
#Backward pass
deltas = [] #Este será el vector con los errores calculados a partir de derivadas parciales.
for iL in reversed(range(0, len(rn))):
_z=out[iL+1][0]
_a=out[iL+1][1]
#print(_z.shape)
#print(_a.shape)
#Ya que para la última capa se le aplica una derivada un poco diferente a las demás
#Es que se hace la siguiente condición:
if iL == len(rn)-1:
#Calcular delta (derivada) en la última capa:
deltas.insert(0, func_Coste[1](_a, _y)*rn[iL].act_f[1](_a))
#_w = rn[iL].w
else:
#Calcular delta (derivada) respecto a capa previa:
deltas.insert(0, deltas[0]@_w.T * rn[iL].act_f[1](_a))
#Se inserta siempre en 0 para ir desplazando los demás resultados.
#deltas.insert(0, _w@deltas[0] * rn[iL].act_f[1](_a))
_w = rn[iL].w #Con esto vamos actualizando nuestros pesos.
#Gradient descent (Recordar que el gradiente es el vector que contiene a las pendientes)
#np.mean: calcula medias | axis: le dice que eje, del (500, 1) el 500
#keepdims: para que no lo convierta en serie, ejemplo de serie: (500, )
#y "lr" el cual es los pasos que daremos por cada iteración, ya lo vimos en el capítulo anterior.
rn[iL].b = rn[iL].b - np.mean(deltas[0], axis=0, keepdims=True) * lr #Básicamente: theta=theta-gradiente*lr
#Y de manera similar con "w":
rn[iL].w = rn[iL].w - out[iL][1].T@deltas[0] * lr #Con esto comenzamos a distribuir el error a las demás capas.
return out[-1][1] #Retornamos la última matriz calculada (u operada) que corresponde a la función de activación.
#entrenar(red_neuronal, x, y, i2_cost, 0.5, train=True) #Línea que comprueba que está funcionando la función "entrenar".
# -
# ## A graficar!
# Ahora lo que haremos será instanciar una nueva red e ir graficando cada cierta cantidad de iteraciones, para así ir viendo como evoluciona nuestra red neuronal.
# +
import time
from IPython.display import clear_output
#p definido al principio de este documento, al igual que x e y
topologia = [p, 4, 1]
#Función de activación definida desupués de la clase "neural_layer":
redNeuronal = crear_redNeuronal(topologia, sigmoide)
#Matriz que almacena cada variación entre la salida de la red y lo que esperamos.
loss=[]
for it in range(10000):
#Entrenamos a la red:
pY = entrenar(redNeuronal, x, y, i2_cost, lr=0.03)
#Con "pY" podremos comparar la salida de nuestra Red Neuronal con los valores que esperamos realmemente.
if it%25 == 0: #Cada 25 iteraciones o ciclos, vemos:
loss.append(i2_cost[0](pY, y))
#Resolución de nuestro gráfico:
res=50
#Valores aleatorios para hacer la gráfica base:
_x0 = np.linspace(-1.5, 1.5, res)
_x1 = np.linspace(-1.5, 1.5, res)
_Y = np.zeros((res, res))
#Ahora guardaremos
for ix0, x0 in enumerate(_x0):
for ix1, x1 in enumerate(_x1):
_Y[ix0, ix1] = entrenar(redNeuronal, np.array([[x0, x1]]), y, i2_cost, train=False)[0][0] #Aquí no entrenamos la red.
#Solo es para hacer predicciones de como la red neuronal está clasificando ambos conjuntos de puntos.
#Aquí comenzamos creando la gráfica, como en el anterior ejercicio:
plt.pcolormesh(_x0, _x1, _Y, cmap="coolwarm") #Coloreando superficies.
plt.axis("equal") #Proporcionamos los ejes.
#Esto está al principio y es para visualizar nuestro conjunto de datos:
plt.scatter(x[y[:, 0]==0, 0], x[y[:, 0]==0, 1], c="skyblue")
plt.scatter(x[y[:, 0]==1, 0], x[y[:, 0]==1, 1], c="salmon")
#No explico mucho pero básicamente borra y vuelve a hacer el dibujo dandole un efecto como de animado:
clear_output(wait=True)
plt.show() #Mostramos el primer gráfico con las superficies y nuestros puntos.
plt.plot(range(len(loss)), loss)
plt.show() #Mostramos un segundo gráfico de como el error se va comportando.
time.sleep(0.1) #Y un pequeño tiempo de descanso para la CPU xD
| IA Notebook #4 - Programando una Red Neuronal desde Cero.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: SageMath 9.2
# language: sagemath
# metadata:
# cocalc:
# description: Open-source mathematical software system
# priority: 10
# url: https://www.sagemath.org/
# name: sage-9.2
# resource_dir: /ext/jupyter/kernels/sage-9.2
# ---
# # Lecture 01: Welcome to Math 157!!
#
# ### Please note: This lecture will be recorded and made available for viewing online. If you do not wish to be recorded, please adjust your camera settings accordingly.
# ## What is Math 157?
#
# Math 157 is a upper level class which explains how mathematicians use software and programming to help understand abstract ideas.
#
# ### It *is not*
# - Abstract/theoretical (any abstract results will be used as a black box)
# - A computer science class
# - Proof based
# - Going to develop one large coherent theory over three months
#
# ### It *will hopefully*
# - Be very concrete, encouraging hands-on student experimentation
# - Cover a wide range of topics, each for about 1 week
# - Get you interested in programming as a tool for mathematics
# - Be fun!
# # Announcements:
# - If you do not have a Cocalc account, please sign up ASAP!
# - If you have a Cocalc account, but do not have a Math 157 Project, please check your email or email me/a TA ASAP!
# - Week 1 Section will be optional "Tech Support." There will be a modified schedule, *for this week only*:
# - Tuesday, 4-4:50 PM
# - Wednesday, 11 AM - 1 PM.
# - Starting Week 2, Sections will run on Tuesdays at 2-2:50 PM, 3-3:50 PM, and 4-4:50 PM. Sections will reinforce material learned and answer questions related to homeworks.
# - If you only recently joined the course you might not have Canvas access just yet. If this is not resolved by tomorrow, please email me! Thanks for your patience :D
# ## Course Logistics (a brief rundown)
#
# For more info, see the course webpage: https://www.math.ucsd.edu/~tgrubb/math157/
#
# #### Lectures can be viewed in two ways:
# - Concurrently: Lectures will be held over Zoom on Mondays, Wednesdays, and Fridays at 4:00 - 4:50 PM Pacific.
# - After the fact: Lectures will be made available on Canvas.
# #### Additionally, the lecture files will be made available to you through your Cocalc account (more on this later)
# - PLEASE NOTE! Viewing lectures is a critical part of the learning process in this course. As such, a small portion of the grade (8%) will be dedicated to "virtual participation." This will be measured through your interaction with the shared lecture files (more on this later)
# #### Grading Setup
# - The majority of your grade in this class will be determined by "homework", with the exception of two quizzes.
# - Homework, 60%:
# - 8 assignments due at the end of Weeks 2 - 9. Each assignment will cover the material discussed over the past week. Your best 6 assignments are scored (each worth 10%).
# - Virtual participation, 8%:
# - Approximately 25 lectures will have participation components, meant to ensure students are keeping up with the weekly material. These are meant to be very easy (and time will be dedicated in lectures to doing them). We will keep at most your top 20 scores to compute your grade.
# - Quizzes, 12%:
# - 2 "open internet" quizzes, equally weighted, on Jan 25 and Feb 22.
# - Final project, 20%:
# - This will consist of two parts. One part is a "cumulative" homework assignment. The other part consists of you constructing a "Lecture" based on a topic of your choosing, which you will then present in small groups (~4 students).
# ## Intermission: Introduction to Cocalc
# 
# ## Course Logistics: Zulip Chatroom
#
# Zulip Chat is similar to Piazza but, in my opinion, it is much better.
#
# You can access the Zulip Chat for this class via this link: https://ucsd157winter2021.zulipchat.com/
#
# You may post about essentially anything you'd like on the Zulip Chat, but *please be respectful to your fellow classmates and to the course staff.*
# ## Course Logistics: Homeworks/Assignments
#
# #### Purely Logistically, Homework is very easy to turn in for this class
#
# - Each week a new folder will appear in your project. This folder will contain the week's homework assignment, as well as any relevant files.
# - You will enter your homework solutions (code, text, etc.) *directly* into the homework assignment file that is given to you.
# - The homework file will automatically save as you work on it (think of a Google Doc).
# - At the due date, each of your files will automatically be collected by a script and sent to the grader.
# - You don't have to "submit" anything; Cocalc does it all for you!
#
# #### IMPORTANT: Do not change the filename or file location of the homework assignment that is given to you!
#
# - You are free to do scratch work anywhere you want (for instance, on your local machine) but at the due date, all of your solutions must be pasted into the original homework file
# - The first homework (Assignment 0) will be a practice run. The process is very easy once you get the hang of it!
# ## Course Logistics: Lectures and Participation Checks
#
# #### You will be given copies of each of the lecture files. They will appear in the "Lectures" folder of your project.
# - The lectures will be made available to you *before* I give them over Zoom. I recommend having the file open on your computer as I talk about it over Zoom/as you watch the recording of the lecture!
# - You are free to edit/play around with the lecture files in your project (I recommend it!). It will not change anyone else's file (the changes you make are local to your project)
# - You can undo these edits at any time via Time Travel
#
# #### IMPORTANT: In (most of) the lecture files, you will see several Participation Checks throughout the file (example below)
# - These are *for you* to play around with, so that you can gain experience with the topics *as I lecture about them* (or afterwards!)
# - You *will be graded* on the participation checks.
# - Time will be dedicated in the lecture for you to do the checks. These are not meant to be difficult or time consuming.
# - Your grade on the participation checks is determined by checking if your local file was edited during a specific time period (i.e. there are no right or wrong answers).
# - The participation checks will be graded on Sundays, to allow for asynchronous viewing of the lectures.
# - Several participation checks will be dropped, to allow for off days/weeks.
# - There will be "practice runs" to make sure everyone understands before it counts.
# # *** Participation Check ***
# *This is an example of what a participation check will look like and how it will work in the lecture. It will not be graded.*
#
# Over the next ~120 seconds: double click on this markdown cell so that you are in "edit mode." Type in the following (if you do not know how to do something, you can double click the above markdown cell to see how I formatted that text!):
#
# Your name:
#
# Something in italics:
#
# A list of something (with bullet points):
#
# # ****************************
# ## (Time Permitting) A taste of what is to come!
#
# We will cover many distinct topics in this class. Hopefully at least one of them will interest you!
# ### Combinatorics
show(graphs.PetersenGraph())
show(graphs.RandomGNP(10,.3))
# ### Natural Language Processing
import nltk
sentence = "I didn't like the new Avengers movie."
tokens = nltk.word_tokenize(sentence)
tokens
taggedTokens = nltk.pos_tag(tokens)
taggedTokens
# ### Cryptography/Number Theory
C = EllipticCurve([0,0,0,-2,0]).plot(xmin=-4, xmax=4, ymin=-3, ymax=3)
show(C)
# ### And much much more!
| Lectures/Lecture01/Lecture01_Jan04.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from tqdm import tqdm
from scipy import stats
plt.rcParams['font.sans-serif']=['Songti SC'] #用来正常显示中文标签
train_data = pd.read_csv('C:/ml_data/kdxf/train/train.csv')
test_data = pd.read_csv('C:/ml_data/kdxf/test/test.csv')
train_data.columns = ['timestamp', 'year', 'month', 'day', 'hour', 'min', 'sec', 'outtemp', 'outhum', 'outatmo',
'inhum', 'inatmo', 'temperature']
test_data.columns = ['timestamp', 'year', 'month', 'day', 'hour', 'min', 'sec', 'outtemp', 'outhum', 'outatmo',
'inhum', 'inatmo']
# + active=""
# 下面是增加temp_change的代码,暂时不要上线,试验证明特征效果不好。
# + active=""
# train_data = pd.read_csv('C:/ml_data/kdxf/train_data_csdn_4.csv')
# test_data = pd.read_csv('C:/ml_data/kdxf/test_data_csdn_4.csv')
# + active=""
# train_data['temp_change'] = train_data['outatmo']
# + active=""
# for i in tqdm(range(1,len(train_data['temp_change']))):
# train_data['temp_change'][i] = train_data['outtemp'][i] - train_data['outtemp'][i-1]
# + active=""
# train_data['temp_change'][0] = train_data['temp_change'][1]
# + active=""
# test_data['temp_change'] = test_data['outatmo']
# for i in tqdm(range(1,len(test_data['temp_change']))):
# test_data['temp_change'][i] = test_data['outtemp'][i] - test_data['outtemp'][i-1]
# + active=""
# test_data['temp_change'][0] = test_data['temp_change'][1]
# + active=""
# del train_data['Unnamed: 0']
# del test_data['Unnamed: 0']
# + active=""
# train_data.to_csv('C:/ml_data/kdxf/train_data_csdn_7.csv',index = False)
# + active=""
# test_data.to_csv('C:/ml_data/kdxf/test_data_csdn_7.csv',index = False)
# -
# + active=""
# 训练集缺失值处理
# -
tempa_miss = [x for x in range(len(train_data['temperature'].isnull())) if train_data['temperature'].isnull()[x] == True]
# +
#预测值存在缺失,直接删除
# -
len_0= train_data.shape[0]
train_data = train_data.drop(axis=0,index = tempa_miss).reset_index()
len_1 = train_data.shape[0]
print('remain_ratio :',len_1/len_0)
del train_data['index']
train_features_with_missing = ['outtemp','outhum','outatmo']
# +
#这些特征使用上下时间点,进行线性融合填充
# -
for feature_single in tqdm(train_features_with_missing):
miss_index = [x for x in range(len(train_data[feature_single].isnull())) if train_data[feature_single].isnull()[x] == True]
for index in miss_index:
value_last = train_data[feature_single][index - 1]
j = 1
while True:
if train_data[feature_single][index +j] > 0:
break
j += 1
ratio_ = (train_data['timestamp'][index] - train_data['timestamp'][index-1])/(train_data['timestamp'][index+j] - train_data['timestamp'][index-1])
train_data[feature_single][index] = ratio_*(train_data[feature_single][index +j] - train_data[feature_single][index - 1] ) + train_data[feature_single][index - 1]
# + active=""
# 测试集缺失值填充
# -
test_features_with_missing = ['outtemp','outhum','outatmo','inhum','inatmo']
# +
#特征还是按照时间融合填充
# -
for feature_single in tqdm(test_features_with_missing):
miss_index = [x for x in range(len(test_data[feature_single].isnull())) if test_data[feature_single].isnull()[x] == True]
for index in miss_index:
value_last = test_data[feature_single][index - 1]
j = 1
while True:
if test_data[feature_single][index +j] > 0:
break
j += 1
ratio_ = (test_data['timestamp'][index] - test_data['timestamp'][index-1])/(test_data['timestamp'][index+j] - test_data['timestamp'][index-1])
test_data[feature_single][index] = ratio_*(test_data[feature_single][index +j] - test_data[feature_single][index - 1] ) + test_data[feature_single][index - 1]
#timestamp_start = train_data['timestamp'][0]
#train_data['timestamp'] = train_data['timestamp'] - timestamp_start
#test_data['timestamp'] = test_data['timestamp'] - timestamp_start
del train_data['timestamp']
del test_data['timestamp']
train_data['day'] = (train_data['month'] - 3) * 31 + train_data['day']
test_data['day'] = (test_data['month'] - 3) * 31 + test_data['day']
# +
del train_data['year']
del train_data['month']
del test_data['year']
del test_data['month']
# +
#预测目标从室内温度变更为室内外温差
# -
train_data['gaptemp'] = train_data['temperature'] - train_data['outtemp']
del train_data['temperature']
train_data['min'] = train_data['hour'] * 60 + train_data['min']
test_data['min'] = test_data['hour'] * 60 + test_data['min']
train_data['sec'] = train_data['min'] * 60 + train_data['sec']
test_data['sec'] = test_data['min'] * 60 + test_data['sec']
all_data = pd.concat([train_data,test_data], axis=0, ignore_index=True)
# +
#新建特征up_wave
# -
all_data['up_wave'] = 0
temp_value = [x for x in all_data['outhum']]
all_data['up_wave'][0] = 0
tt = [x for x in all_data['outhum']]
plt.plot(all_data['up_wave'][k*1000:k*1000+1000])
k = 0
plt.plot( tt[k*1000:k*1000+5000])
plt.savefig('C:/ml_data/kdxf/qwe.png')
for i in tqdm(range(k*1000,k*1000 + 1000)):
ratio =(tt[i] - tt[i-1])/tt[i] * 100
if ratio < 2 and ratio > -2:
continue
else:
print(i)
print(ratio)
continue
for i in tqdm(range(23831,23840)):
all_data['up_wave'][i] = (1-(i-23831)/(23840-23831)) * (11.39240506329114)
all_data.isnull().sum()
# + active=""
# 人工判定异常值处理
# -
features_useful = ['inatmo','inhum','outatmo','outhum', 'outtemp']
#该算法无法处理第一个和最后一个数据,人工鉴定不存在这种情况
for v in [0,2]:
feature_single = features_useful[v]
for i in tqdm(range(1,len(all_data[feature_single])-1)):
if 20 * abs(all_data[feature_single][i] - all_data[feature_single][i-1]) > all_data[feature_single][i-1]:
all_data[feature_single][i] = (all_data[feature_single][i-1] +all_data[feature_single][i+1] )/2
for v in [0,2]:
feature_single = features_useful[v]
for i in (range(1,len(all_data[feature_single])-1)):
if 20 * abs(all_data[feature_single][i] - all_data[feature_single][i-1]) > all_data[feature_single][i-1]:
print(i)
# +
#正态化程度还行,不进行正态化处理了
#事后诸葛亮:其实还是应该做的
# -
train_data['gaptemp'].skew()
train_data['gaptemp'].kurt()
all_data.columns
# + active=""
# 新建特征:室内外差值,差值的比例
# -
all_data['gapatmo'] = all_data['inatmo'] - all_data['outatmo']
all_data['gaphum'] = all_data['inhum'] - all_data['outhum']
all_data['gapatmo_ratio'] = all_data['gapatmo'].values/all_data['outatmo'].values * 10000
all_data['gaphum_ratio'] = all_data['gaphum'].values/all_data['outhum'].values * 100
# +
#聚合特征
# -
group_features = []
for f in tqdm(['outtemp', 'outhum', 'outatmo', 'inhum', 'inatmo']):
all_data['MDH_{}_medi'.format(f)] = all_data.groupby(['day', 'hour'])[f].transform('median')
all_data['MDH_{}_mean'.format(f)] = all_data.groupby(['day', 'hour'])[f].transform('mean')
all_data['MDH_{}_max'.format(f)] = all_data.groupby(['day', 'hour'])[f].transform('max')
all_data['MDH_{}_min'.format(f)] = all_data.groupby(['day', 'hour'])[f].transform('min')
all_data['MDH_{}_std'.format(f)] = all_data.groupby(['day', 'hour'])[f].transform('std')
group_features.append('MDH_{}_medi'.format(f))
group_features.append('MDH_{}_mean'.format(f))
all_data = all_data.fillna(method='bfill')
# +
for f in tqdm(['gapatmo','gaphum','gapatmo_ratio','gaphum_ratio']):
all_data['MDH_{}_medi'.format(f)] = all_data.groupby(['day', 'hour'])[f].transform('median')
all_data['MDH_{}_mean'.format(f)] = all_data.groupby(['day', 'hour'])[f].transform('mean')
all_data['MDH_{}_max'.format(f)] = all_data.groupby(['day', 'hour'])[f].transform('max')
all_data['MDH_{}_min'.format(f)] = all_data.groupby(['day', 'hour'])[f].transform('min')
all_data['MDH_{}_std'.format(f)] = all_data.groupby(['day', 'hour'])[f].transform('std')
all_data = all_data.fillna(method='bfill')
# +
for f1 in tqdm(['outtemp', 'outhum', 'outatmo', 'inhum', 'inatmo'] + group_features):
for f2 in ['outtemp', 'outhum', 'outatmo', 'inhum', 'inatmo'] + group_features:
if f1 != f2:
colname = '{}_{}_ratio'.format(f1, f2)
all_data[colname] = all_data[f1].values / all_data[f2].values
all_data = all_data.fillna(method='bfill')
# + active=""
# for f1 in tqdm(['gapatmo','gaphum'] ):
#
# for f2 in ['gapatmo','gaphum'] :
# if f1 != f2:
# colname = '{}_{}_ratio'.format(f1, f2)
# all_data[colname] = all_data[f1].values / all_data[f2].values
#
# all_data = all_data.fillna(method='bfill')
# +
for f in ['outtemp', 'outhum', 'outatmo', 'inhum', 'inatmo','gapatmo','gaphum','gapatmo_ratio','gaphum_ratio']:
tmp_df = pd.DataFrame()
for t in tqdm(range(15, 45)):
tmp = all_data[all_data['day'] < t].groupby(['hour'])[f].agg({'mean'}).reset_index()
tmp.columns = ['hour', 'hit_{}_mean'.format(f)]
tmp['day'] = t
tmp_df = tmp_df.append(tmp)
all_data = all_data.merge(tmp_df, on=['day', 'hour'], how='left')
all_data = all_data.fillna(method='bfill')
# -
for f in ['outtemp', 'outhum', 'outatmo', 'inhum', 'inatmo','gapatmo','gaphum','gapatmo_ratio','gaphum_ratio']:
all_data[f + '_20_bin'] = pd.cut(all_data[f], 20, duplicates='drop').apply(lambda x: x.left).astype(int)
all_data[f + '_50_bin'] = pd.cut(all_data[f], 50, duplicates='drop').apply(lambda x: x.left).astype(int)
all_data[f + '_100_bin'] = pd.cut(all_data[f], 100, duplicates='drop').apply(lambda x: x.left).astype(int)
all_data[f + '_200_bin'] = pd.cut(all_data[f], 200, duplicates='drop').apply(lambda x: x.left).astype(int)
# +
for i in tqdm(['outtemp', 'outhum', 'outatmo', 'inhum', 'inatmo','gapatmo','gaphum','gapatmo_ratio','gaphum_ratio']):
f1 = i + '_20_bin'
for f2 in ['outtemp', 'outhum', 'outatmo', 'inhum', 'inatmo','gapatmo','gaphum','gapatmo_ratio','gaphum_ratio']:
all_data['{}_{}_medi'.format(f1, f2)] = all_data.groupby([f1])[f2].transform('median')
all_data['{}_{}_mean'.format(f1, f2)] = all_data.groupby([f1])[f2].transform('mean')
all_data['{}_{}_max'.format(f1, f2)] = all_data.groupby([f1])[f2].transform('max')
all_data['{}_{}_min'.format(f1, f2)] = all_data.groupby([f1])[f2].transform('min')
f1 = i + '_20_bin'
for f2 in ['outtemp', 'outhum', 'outatmo', 'inhum', 'inatmo','gapatmo','gaphum','gapatmo_ratio','gaphum_ratio']:
all_data['{}_{}_medi'.format(f1, f2)] = all_data.groupby([f1])[f2].transform('median')
all_data['{}_{}_mean'.format(f1, f2)] = all_data.groupby([f1])[f2].transform('mean')
all_data['{}_{}_max'.format(f1, f2)] = all_data.groupby([f1])[f2].transform('max')
all_data['{}_{}_min'.format(f1, f2)] = all_data.groupby([f1])[f2].transform('min')
f1 = i + '_100_bin'
for f2 in ['outtemp', 'outhum', 'outatmo', 'inhum', 'inatmo','gapatmo','gaphum','gapatmo_ratio','gaphum_ratio']:
all_data['{}_{}_medi'.format(f1, f2)] = all_data.groupby([f1])[f2].transform('median')
all_data['{}_{}_mean'.format(f1, f2)] = all_data.groupby([f1])[f2].transform('mean')
all_data['{}_{}_max'.format(f1, f2)] = all_data.groupby([f1])[f2].transform('max')
all_data['{}_{}_min'.format(f1, f2)] = all_data.groupby([f1])[f2].transform('min')
f1 = i + '_200_bin'
for f2 in ['outtemp', 'outhum', 'outatmo', 'inhum', 'inatmo','gapatmo','gaphum','gapatmo_ratio','gaphum_ratio']:
all_data['{}_{}_medi'.format(f1, f2)] = all_data.groupby([f1])[f2].transform('median')
all_data['{}_{}_mean'.format(f1, f2)] = all_data.groupby([f1])[f2].transform('mean')
all_data['{}_{}_max'.format(f1, f2)] = all_data.groupby([f1])[f2].transform('max')
all_data['{}_{}_min'.format(f1, f2)] = all_data.groupby([f1])[f2].transform('min')
# -
all_data.shape
train_data = all_data[0:24807]
test_data = all_data[24807:25213]
train_data.to_csv('C:/ml_data/kdxf/train_data_csdn_8.csv',index = False)
del test_data['gaptemp']
test_data.to_csv('C:/ml_data/kdxf/test_data_csdn_8.csv',index = False)
| kdxf_eda_master.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## MNIST Adversarial defense using SymDNN - This notebook has limited number of vizualizations
# +
# Basic imports & definitions for MNIST inference
import torch
import random
import torchvision
from torchvision import datasets, transforms
import torch.nn as nn
import torch.nn.functional as F
import matplotlib.pyplot as plt
import numpy as np
import math
import faiss
import sys
torch.manual_seed(0)
np.random.seed(0)
random.seed(0)
import warnings
warnings.filterwarnings('ignore')
plt.rcParams['figure.dpi'] = 100
sys.path.insert(1, '../core')
from patchutils import symdnn_purify, fm_to_symbolic_fm as sym_mnist
# +
# Basic model for MNIST inference
# From my training code
torch.manual_seed(0)
np.random.seed(0)
random.seed(0)
apply_transform = transforms.Compose([transforms.Resize(32),transforms.ToTensor(),
transforms.Normalize((0.1309,), (0.2893,))])
# Change the dataset folder to the proper location in a new system
testset = datasets.MNIST(root='../../../dataset', train=False, download=True, transform=apply_transform)
testloader = torch.utils.data.DataLoader(testset, batch_size=1, shuffle=False)
testloader_std = torch.utils.data.DataLoader(testset, batch_size=64, shuffle=False)
random_indices = list(range(0, len(testset), 5))
testset_subset = torch.utils.data.Subset(testset, random_indices)
testloader_subset = torch.utils.data.DataLoader(testset_subset, batch_size=1, shuffle=False)
class CNN_LeNet(nn.Module):
def __init__(self):
super(CNN_LeNet, self).__init__()
# Define the net structure
# This is the input layer first Convolution
self.conv1 = nn.Conv2d(1, 6, kernel_size=5)
self.pool1 = nn.MaxPool2d(kernel_size=2,stride=2)
self.conv2 = nn.Conv2d(6, 16, kernel_size=5)
self.pool2 = nn.MaxPool2d(kernel_size=2,stride=2)
self.fc1 = nn.Linear(400,120)
self.fc2 = nn.Linear(120,84)
self.fc3 = nn.Linear(84, 10)
def forward(self, x):
x = F.relu(self.conv1(x))
x = self.pool1(x)
x = F.relu(self.conv2(x))
x = self.pool2(x)
x = x.view(-1, 400)
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = self.fc3(x)
x = F.softmax(x,dim=1)
return x
# -
classes = ('0', '1', '2', '3', '4', '5', '6', '7', '8', '9')
# ### The standard model "pretrained_model" is trained on normalized MNIST data
# ### The model "pretrained_nr_model" is trained on data clamped between 0 & 1
# ### The model "pretrained_sc_model" is trained on data scaled between 0 & 1
pretrained_sc_model = "./mnist_sc_v0_best.pt"
mnist_sc_model = CNN_LeNet()
mnist_sc_model.load_state_dict(torch.load("mnist_sc_v0_best.pt"))
mnist_sc_model.eval()
# Define a custom function that will clamp the images between 0 & 1 , without being too harsh as torch.clamp
def softclamp01(image_tensor):
image_tensor_shape = image_tensor.shape
image_tensor = image_tensor.view(image_tensor.size(0), -1)
image_tensor -= image_tensor.min(1, keepdim=True)[0]
image_tensor /= image_tensor.max(1, keepdim=True)[0]
image_tensor = image_tensor.view(image_tensor_shape)
return image_tensor
# Lets check the kind of prediction the model is doing - the standard non symbolic infrence
def mnist_test_base_acc(model, nr=0):
base_clampled_clean = 0
total = 0
# Clean base inference
for images, labels in testloader_std:
with torch.no_grad():
if nr == 0:
X = images
elif nr == 1:
X = softclamp01(images)
else:
X = torch.clamp(images, 0,1)
y = labels
output = model.forward(X)
for idx, i in enumerate(output):
if torch.argmax(i) == y[idx]:
base_clampled_clean += 1
#else:
# # Whenever there is an error, print the image
# print("Misclassification: Model: clean base gradinit. Test Image #: {}, Mispredicted label: {}".format(total+1, torch.argmax(i)))
total +=1
return float(base_clampled_clean / total)
# Test accuracy of symbolic inference
def mnist_test_sym_acc(model, nr, sym_mnist,n_clusters, index, patch_size, stride, channel_count, instr=False):
correct = 0
total = 0
centroid_lut = index.reconstruct_n(0, n_clusters)
if instr:
pdf = np.zeros((n_clusters,), dtype=int)
model.eval()
with torch.no_grad():
for data in testloader:
X, y = data
if nr == 0:
X = X
elif nr == 1:
X = softclamp01(X)
else:
X = torch.clamp(X, 0,1)
if instr:
Xsym_, pdf = sym_mnist(X.squeeze(), n_clusters, index, centroid_lut, patch_size, stride, channel_count, ana=False, multi=False, instr=True, randomize=False, rlevel=None, rbalance=True, pdf=pdf)
else:
Xsym_ = sym_mnist(X.squeeze(), n_clusters, index, centroid_lut, patch_size, stride, channel_count, ana=False, multi=False, instr=False, randomize=False, rlevel=None, rbalance=True, pdf=None)
Xsym = torch.from_numpy(Xsym_)
Xsym = Xsym.unsqueeze(0)
output = model.forward(Xsym.float())
for idx, i in enumerate(output):
if torch.argmax(i) == y[idx]:
correct += 1
total += 1
if instr:
return round(correct/total, 4), pdf
else:
return round(correct/total, 4)
# ### Lets now try the final combination, which is both model & index trained with same scaled data - we also try different symbol sizes to come up with the best clean accuracy
index_32 = faiss.read_index("./kmeans_img_mnist_k2_s0_c32_sc_v0.index")
n_clusters_32 = 32
patch_size = (2, 2)
channel_count = 1
repeat = 2
location=False
stride = 0
acc = mnist_test_sym_acc(mnist_sc_model, 1, sym_mnist, n_clusters_32, index_32, patch_size, stride, channel_count)
print("Symbolic test accuracy (codebook 32):{}% ".format(100*acc))
index = faiss.read_index("./kmeans_img_mnist_k4_s0_c32_sc_v0.index")
n_clusters=32
patch_size = (4, 4)
channel_count = 1
repeat = 2
location=False
stride = 0
acc = mnist_test_sym_acc(mnist_sc_model, 1, sym_mnist, n_clusters, index, patch_size, stride, channel_count)
print("Symbolic test accuracy (codebook 32):{}% ".format(100*acc))
index = faiss.read_index("./kmeans_img_mnist_k8_s0_c32_sc_v0.index")
n_clusters=32
patch_size = (8, 8)
channel_count = 1
repeat = 2
location=False
stride = 0
acc = mnist_test_sym_acc(mnist_sc_model, 1, sym_mnist, n_clusters, index, patch_size, stride, channel_count)
print("Symbolic test accuracy (codebook 32):{}% ".format(100*acc))
index_64 = faiss.read_index("./kmeans_img_mnist_k2_s0_c64_sc_v0.index")
n_clusters_64=64
patch_size = (2, 2)
channel_count = 1
repeat = 2
location=False
stride = 0
centroid_lut = index.reconstruct_n(0, n_clusters)
acc = mnist_test_sym_acc(mnist_sc_model, 1, sym_mnist, n_clusters_64, index_64, patch_size, stride, channel_count)
print("Symbolic test accuracy (codebook 64):{}% ".format(100*acc))
index = faiss.read_index("./kmeans_img_mnist_k4_s0_c64_sc_v0.index")
n_clusters=64
patch_size = (4, 4)
channel_count = 1
repeat = 2
location=False
stride = 0
acc = mnist_test_sym_acc(mnist_sc_model, 1, sym_mnist, n_clusters, index, patch_size, stride, channel_count)
print("Symbolic test accuracy (codebook 64):{}% ".format(100*acc))
index = faiss.read_index("./kmeans_img_mnist_k8_s0_c64_sc_v0.index")
n_clusters=64
patch_size = (8, 8)
channel_count = 1
repeat = 2
location=False
stride = 0
centroid_lut = index.reconstruct_n(0, n_clusters)
acc = mnist_test_sym_acc(mnist_sc_model, 1, sym_mnist, n_clusters, index, patch_size, stride, channel_count)
print("Symbolic test accuracy (codebook 64):{}% ".format(100*acc))
index = faiss.read_index("./kmeans_img_mnist_k2_s0_c128_sc_v0.index")
n_clusters=128
patch_size = (2, 2)
channel_count = 1
repeat = 2
location=False
stride = 0
centroid_lut = index.reconstruct_n(0, n_clusters)
acc = mnist_test_sym_acc(mnist_sc_model, 1, sym_mnist, n_clusters, index, patch_size, stride, channel_count)
print("Symbolic test accuracy (codebook 128):{}% ".format(100*acc))
index = faiss.read_index("./kmeans_k4_s0_c128_sc_mnist.index")
n_clusters=128
patch_size = (4, 4)
channel_count = 1
repeat = 2
location=False
stride = 0
centroid_lut = index.reconstruct_n(0, n_clusters)
acc = mnist_test_sym_acc(mnist_sc_model, 1, sym_mnist, n_clusters, index, patch_size, stride, channel_count)
print("Symbolic test accuracy (codebook 128):{}% ".format(100*acc))
index = faiss.read_index("./kmeans_k8_s0_c128_sc_mnist.index")
n_clusters=128
patch_size = (8, 8)
channel_count = 1
repeat = 2
location=False
stride = 0
centroid_lut = index.reconstruct_n(0, n_clusters)
acc = mnist_test_sym_acc(mnist_sc_model, 1, sym_mnist, n_clusters, index, patch_size, stride, channel_count)
print("Symbolic test accuracy (codebook 128):{}% ".format(100*acc))
patterns = [ "/" , "\\" , "|" , "-" , "+" , "x", "o", "O", ".", "*" ]
# +
# Small hack in Jupyter
plt.rcParams['figure.figsize'] = [6,4]
plt.rcParams['figure.dpi'] = 120 # 200 e.g. is really fine, but slower
plt.rcParams.update({'font.size': 14})
N = 3
ind = np.arange(N)
labels = ('32 symbols', '64 symbols', '128 symbols')
k_2 = (99.11, 99.09, 99.14)
k_4 = (98.6, 98.93, 99.06)
k_8 = (89.3, 93.76, 95.69)
fig = plt.figure()
ax = fig.add_subplot(111)
rects1 = ax.plot(ind, k_2, marker="*", alpha=0.7, color='darkblue')
rects2 = ax.plot(ind, k_4, marker="^", color='darkred', alpha=0.9)
rects3 = ax.plot(ind, k_8, marker="o", linestyle="--", color='black', alpha=0.9)
# add some
ax.set_ylabel('Accuracy', fontsize=16)
ax.set_xlabel('Number of symbols used for abstraction', fontsize=16)
ax.set_xticks(ind)
ax.set_xticklabels(labels, fontsize=14)
ax.legend( (rects1[0], rects2[0], rects3[0]), ('2x2 patch size', '4x4 patch size', '8x8 patch size'),loc='upper center', bbox_to_anchor=(0.75, 0.4))
plt.tight_layout()
plt.show()
#plt.savefig('../patch_size_accuracy_mnist.png', format='png', dpi=1000)
# -
index_128 = faiss.read_index("./kmeans_img_mnist_k2_s0_c128_sc_v0.index")
n_clusters_128=128
patch_size = (2, 2)
channel_count = 1
repeat = 2
location=False
stride = 0
acc = mnist_test_sym_acc(mnist_sc_model, 0, sym_mnist, n_clusters_128, index_128, patch_size, stride, channel_count)
print("Symbolic test accuracy (codebook 128):{}% ".format(100*acc))
index_256 = faiss.read_index("./kmeans_img_mnist_k2_s0_c256_sc_v0.index")
n_clusters_256=256
patch_size = (2, 2)
channel_count = 1
repeat = 2
location=False
stride = 0
acc = mnist_test_sym_acc(mnist_sc_model, 0, sym_mnist, n_clusters_256, index_256, patch_size, stride, channel_count)
print("Symbolic test accuracy (codebook 256):{}% ".format(100*acc))
# +
import torchvision.utils
from torchvision import models
import torchattacks
from torchattacks import *
print("PyTorch", torch.__version__)
print("Torchvision", torchvision.__version__)
print("Torchattacks", torchattacks.__version__)
print("Numpy", np.__version__)
# +
class UnNormalize(object):
def __init__(self, mean, std):
self.mean = mean
self.std = std
def __call__(self, tensor):
"""
Args:
tensor (Tensor): Tensor image of size (C, H, W) to be normalized.
Returns:
Tensor: Normalized image.
"""
for t, m, s in zip(tensor, self.mean, self.std):
t.mul_(s).add_(m)
# The normalize code -> t.sub_(m).div_(s)
return tensor
unorm = UnNormalize(mean=(0.1309), std=(0.2893))
# -
def plot_all(img, ttl, norm=False):
# create figure (fig), and array of axes (ax)
fig = plt.figure(figsize=(5, 2))
columns = 4
rows = 1
for i in range(1, columns*rows +1):
j = i-1
if norm:
norm_img = unorm(img[j])
else:
norm_img = img[j]
#npimg = norm_img.numpy()
npimg = norm_img
fig.add_subplot(rows, columns, i)
plt.imshow(npimg, cmap="gray")
# write title for identification
plt.rcParams.update({'font.size': 8})
plt.rcParams["font.weight"] = "bold"
plt.title(ttl[j])
#axi.set_axis_off()
plt.axis('off')
plt.tight_layout(True)
plt.show()
def viz_adversarial_attack(atk, model, index, n_clusters , num_try, sc, norm=False):
counter = 0
correct_std = 0
correct_std_atk = 0
correct_sym = 0
correct_sym_atk = 0
total =0
plot_count = 500
patch_size = (2, 2)
centroid_lut = index.reconstruct_n(0, n_clusters)
for images, labels in testloader_subset:
if sc:
# Soft clamp image to make the attack effective
images_sc = softclamp01(images)
else:
images_sc = images
if counter > num_try:
break
#print("For image #",counter)
counter += 1
img_arr = []
ttl_arr = []
outputs_orig = model(images_sc)
_, pre_orig = torch.max(outputs_orig.data, 1)
if (pre_orig == labels): # We do further testing if the original model is right
images_std = images_sc
title_text = "Orig: "+str(classes[labels])
if counter % plot_count == 0:
plot_img = images_std.squeeze()
plot_img = images.squeeze()
img_arr.append(plot_img)
ttl_arr.append(title_text)
correct_std += (pre_orig == labels).sum()
# Symbolic inference
Xsym_ = sym_mnist(images_sc.squeeze(), n_clusters, index, centroid_lut, patch_size, stride, channel_count)
Xsym = torch.from_numpy(Xsym_)
Xsym = Xsym.unsqueeze(0)
outputs_sym = model.forward(Xsym.float())
_, pre_sym = torch.max(outputs_sym.data, 1)
if (pre_sym == labels):
correct_sym += (pre_sym == labels).sum()
if counter % plot_count == 0:
plot_img = Xsym.float().squeeze()
img_arr.append(plot_img)
title_text = "Sym: "+str(classes[pre_sym])
ttl_arr.append(title_text)
# Attack on vanila inference
adv_images = atk(images_sc, labels)
outputs_std = model(adv_images)
_, pre_std = torch.max(outputs_std.data, 1)
if (pre_std == labels):
correct_std_atk += (pre_std == labels).sum()
if counter % plot_count == 0:
plot_img_adv = adv_images.squeeze()
plot_img = plot_img_adv.data.cpu().numpy().copy()
img_arr.append(plot_img)
title_text = "Orig (perturb) :"+str(classes[pre_std])
ttl_arr.append(title_text)
# Attack on symbolic inference
pfm = adv_images.data.cpu().numpy().copy()
# Re-classify the perturbed image
Xsym_ = sym_mnist(pfm.squeeze(), n_clusters, index, centroid_lut, patch_size, stride, channel_count)
Xsym = torch.from_numpy(Xsym_)
Xsym = Xsym.unsqueeze(0)
outputs_sym_atk = model.forward(Xsym.float())
_, pre_sym_atk = torch.max(outputs_sym_atk.data, 1)
if (pre_sym_atk == labels):
correct_sym_atk += (pre_sym_atk == labels).sum()
if counter % plot_count == 0:
plot_img = Xsym.float().squeeze()
img_arr.append(plot_img)
title_text = "Sym (perturb):"+str(classes[pre_sym_atk])
ttl_arr.append(title_text)
if counter % plot_count == 0:
plot_all(img_arr, ttl_arr, norm)
total += 1
print('Attack on model: {}'.format(atk))
print('Standard accuracy: %.2f %%' % (100 * float(correct_std) / total))
print('Symbolic accuracy: %.2f %%' % (100 * float(correct_sym) / total))
print('Attacked standard accuracy: %.2f %%' % (100 * float(correct_std_atk) / total))
print('Attacked Symbolic accuracy: %.2f %%' % (100 * float(correct_sym_atk) / total))
atks = [
FGSM(mnist_sc_model, eps=8/255),
BIM(mnist_sc_model, eps=8/255, alpha=2/255, steps=100),
RFGSM(mnist_sc_model, eps=8/255, alpha=2/255, steps=100),
CW(mnist_sc_model, c=1, lr=0.01, steps=100, kappa=0),
PGD(mnist_sc_model, eps=8/255, alpha=2/225, steps=100, random_start=True),
PGDL2(mnist_sc_model, eps=1, alpha=0.2, steps=100),
EOTPGD(mnist_sc_model, eps=8/255, alpha=2/255, steps=100, eot_iter=2),
FFGSM(mnist_sc_model, eps=8/255, alpha=10/255),
TPGD(mnist_sc_model, eps=8/255, alpha=2/255, steps=100),
MIFGSM(mnist_sc_model, eps=8/255, alpha=2/255, steps=100, decay=0.1),
VANILA(mnist_sc_model),
GN(mnist_sc_model, sigma=0.1),
APGD(mnist_sc_model, eps=8/255, steps=100, eot_iter=1, n_restarts=1, loss='ce'),
APGD(mnist_sc_model, eps=8/255, steps=100, eot_iter=1, n_restarts=1, loss='dlr'),
APGDT(mnist_sc_model, eps=8/255, steps=100, eot_iter=1, n_restarts=1),
FAB(mnist_sc_model, eps=8/255, steps=100, n_classes=10, n_restarts=1, targeted=False),
FAB(mnist_sc_model, eps=8/255, steps=100, n_classes=10, n_restarts=1, targeted=True),
Square(mnist_sc_model, eps=8/255, n_queries=5000, n_restarts=1, loss='ce'),
AutoAttack(mnist_sc_model, eps=8/255, n_classes=10, version='standard'),
OnePixel(mnist_sc_model, pixels=5, inf_batch=50),
DeepFool(mnist_sc_model, steps=100),
DIFGSM(mnist_sc_model, eps=8/255, alpha=2/255, steps=100, diversity_prob=0.5, resize_rate=0.9)
]
# Lets choose one attack. How about EOTPGD? doesnt take much time. Will also try longer ones - Autoattck.
attack_name_level = EOTPGD(mnist_sc_model, eps=8/255, alpha=2/255, steps=100, eot_iter=2)
viz_adversarial_attack(attack_name_level, mnist_sc_model, index_32, n_clusters_32, 2000, True, norm=False)
# Lets choose one attack. How about EOTPGD? doesnt take much time. Will also try longer ones - Autoattck.
attack_name_level = EOTPGD(mnist_sc_model, eps=8/255, alpha=2/255, steps=100, eot_iter=2)
viz_adversarial_attack(attack_name_level, mnist_sc_model, index_64, n_clusters_64, 2000, True, norm=False)
# Lets choose one attack. How about EOTPGD? doesnt take much time. Will also try longer ones - Autoattck.
attack_name_level = EOTPGD(mnist_sc_model, eps=8/255, alpha=2/255, steps=100, eot_iter=2)
viz_adversarial_attack(attack_name_level, mnist_sc_model, index_128, n_clusters_128, 2000, True, norm=False)
# Lets choose one attack. How about EOTPGD? doesnt take much time. Will also try longer ones - Autoattck.
attack_name_level = EOTPGD(mnist_sc_model, eps=8/255, alpha=2/255, steps=100, eot_iter=2)
viz_adversarial_attack(attack_name_level, mnist_sc_model, index_256, n_clusters_256, 2000, True, norm=False)
# +
# Lets choose one attack. How about EOTPGD? doesnt take much time. Will also try longer ones - Autoattck.
attack_name_level = EOTPGD(mnist_sc_model, eps=16/255, alpha=2/255, steps=100, eot_iter=2)
viz_adversarial_attack(attack_name_level, mnist_sc_model, index_32, n_clusters_32, 2000, True, norm=False)
viz_adversarial_attack(attack_name_level, mnist_sc_model, index_64, n_clusters_64, 2000, True, norm=False)
viz_adversarial_attack(attack_name_level, mnist_sc_model, index_128, n_clusters_128, 2000, True, norm=False)
viz_adversarial_attack(attack_name_level, mnist_sc_model, index_256, n_clusters_256, 2000, True, norm=False)
attack_name_level = EOTPGD(mnist_sc_model, eps=32/255, alpha=2/255, steps=100, eot_iter=2)
viz_adversarial_attack(attack_name_level, mnist_sc_model, index_32, n_clusters_32, 2000, True, norm=False)
viz_adversarial_attack(attack_name_level, mnist_sc_model, index_64, n_clusters_64, 2000, True, norm=False)
viz_adversarial_attack(attack_name_level, mnist_sc_model, index_128, n_clusters_128, 2000, True, norm=False)
viz_adversarial_attack(attack_name_level, mnist_sc_model, index_256, n_clusters_256, 2000, True, norm=False)
# +
# Small hack in Jupyter
plt.rcParams['figure.figsize'] = [6,4]
plt.rcParams['figure.dpi'] = 120 # 200 e.g. is really fine, but slower
plt.rcParams.update({'font.size': 14})
N = 3
ind = np.arange(N)
labels = ('8/255', '16/255', '32/255')
c_32 = (98.4, 93.25, 73.85)
c_64 = (98.05, 93.75, 69.10)
c_128 = (97.55, 93.15, 68.50)
c_256 = (97.85, 93.00, 63.35)
c_std = (95.75, 85.80 ,29.60)
fig = plt.figure()
ax = fig.add_subplot(111)
rects1 = ax.plot(ind, c_32, linestyle=":", alpha=0.7, color='darkblue')
rects2 = ax.plot(ind, c_64, linewidth="4", linestyle="-.", color='red', alpha=0.9)
rects3 = ax.plot(ind, c_128, linestyle="-", color='black')
rects4 = ax.plot(ind, c_256, linewidth="4", linestyle="--", color='lightcoral', alpha=0.9)
rects5 = ax.plot(ind, c_std, color='yellow', alpha=0.9)
# add some
ax.set_ylabel('Accuracy', fontsize=16)
ax.set_xlabel('Attack strength: maximum change per pixel', fontsize=16)
ax.set_xticks(ind)
ax.set_xticklabels(labels, fontsize=14)
ax.legend( (rects1[0], rects2[0], rects3[0], rects4[0], rects5[0]), ('Symbols:32', 'Symbols:64', 'Symbols:128', 'Symbols:256' , 'Non-symbolic'),loc='upper center', bbox_to_anchor=(0.3, 0.6))
plt.tight_layout()
#plt.show()
plt.savefig('../strength_accuracy_symbols_mnist.png', format='png', dpi=1000)
| mnist/ablation-experiments-patch-sizes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.5 64-bit (''dyn'': conda)'
# name: python3
# ---
# +
import gym
import numpy as np
import torch
import torchkit.pytorch_utils as ptu
# import environments
import envs.pomdp
# import recurrent model-free RL (separate architecture)
from policies.models.policy_rnn import ModelFreeOffPolicy_Separate_RNN as Policy_RNN
# import the replay buffer
from buffers.seq_replay_buffer_vanilla import SeqReplayBuffer
from utils import helpers as utl
# -
# ## Build a POMDP environment: Pendulum-V (only observe the velocity)
# +
cuda_id = 0 # -1 if using cpu
ptu.set_gpu_mode(torch.cuda.is_available() and cuda_id >= 0, cuda_id)
env_name = "Pendulum-V-v0"
env = gym.make(env_name)
max_trajectory_len = env._max_episode_steps
act_dim = env.action_space.shape[0]
obs_dim = env.observation_space.shape[0]
print(env, obs_dim, act_dim, max_trajectory_len)
# -
# ## Build a recurent model-free RL agent: separate architecture, `lstm` encoder, `oar` policy input space, `td3` RL algorithm (context length set later)
agent = Policy_RNN(
obs_dim=obs_dim,
action_dim=act_dim,
encoder="lstm",
algo="td3",
action_embedding_size=8,
state_embedding_size=32,
reward_embedding_size=8,
rnn_hidden_size=128,
dqn_layers=[128, 128],
policy_layers=[128, 128],
lr=0.0003,
gamma=0.9,
tau=0.005,
).to(ptu.device)
# ## Define other training parameters such as context length and training frequency
# +
num_updates_per_iter = 1.0 # training frequency
sampled_seq_len = 64 # context length
buffer_size = 1e6
batch_size = 32
num_iters = 150
num_init_rollouts_pool = 5
num_rollouts_per_iter = 1
total_rollouts = num_init_rollouts_pool + num_iters * num_rollouts_per_iter
n_env_steps_total = max_trajectory_len * total_rollouts
_n_env_steps_total = 0
print("total env episodes", total_rollouts, "total env steps", n_env_steps_total)
# -
# ## Define key functions: collect rollouts and policy update
# +
@torch.no_grad()
def collect_rollouts(
num_rollouts, random_actions=False, deterministic=False, train_mode=True
):
"""collect num_rollouts of trajectories in task and save into policy buffer
:param
random_actions: whether to use policy to sample actions, or randomly sample action space
deterministic: deterministic action selection?
train_mode: whether to train (stored to buffer) or test
"""
if not train_mode:
assert random_actions == False and deterministic == True
total_steps = 0
total_rewards = 0.0
for idx in range(num_rollouts):
steps = 0
rewards = 0.0
obs = ptu.from_numpy(env.reset())
obs = obs.reshape(1, obs.shape[-1])
done_rollout = False
# get hidden state at timestep=0, None for mlp
action, reward, internal_state = agent.get_initial_info()
if train_mode:
# temporary storage
obs_list, act_list, rew_list, next_obs_list, term_list = (
[],
[],
[],
[],
[],
)
while not done_rollout:
if random_actions:
action = ptu.FloatTensor([env.action_space.sample()]) # (1, A)
else:
# policy takes hidden state as input for rnn, while takes obs for mlp
(action, _, _, _), internal_state = agent.act(
prev_internal_state=internal_state,
prev_action=action,
reward=reward,
obs=obs,
deterministic=deterministic,
)
# observe reward and next obs (B=1, dim)
next_obs, reward, done, info = utl.env_step(env, action.squeeze(dim=0))
done_rollout = False if ptu.get_numpy(done[0][0]) == 0.0 else True
# update statistics
steps += 1
rewards += reward.item()
# early stopping env: such as rmdp, pomdp, generalize tasks. term ignores timeout
term = (
False
if "TimeLimit.truncated" in info or steps >= max_trajectory_len
else done_rollout
)
if train_mode:
# append tensors to temporary storage
obs_list.append(obs) # (1, dim)
act_list.append(action) # (1, dim)
rew_list.append(reward) # (1, dim)
term_list.append(term) # bool
next_obs_list.append(next_obs) # (1, dim)
# set: obs <- next_obs
obs = next_obs.clone()
if train_mode:
# add collected sequence to buffer
policy_storage.add_episode(
observations=ptu.get_numpy(torch.cat(obs_list, dim=0)), # (L, dim)
actions=ptu.get_numpy(torch.cat(act_list, dim=0)), # (L, dim)
rewards=ptu.get_numpy(torch.cat(rew_list, dim=0)), # (L, dim)
terminals=np.array(term_list).reshape(-1, 1), # (L, 1)
next_observations=ptu.get_numpy(
torch.cat(next_obs_list, dim=0)
), # (L, dim)
)
print(
"Mode:",
"Train" if train_mode else "Test",
"env_steps",
steps,
"total rewards",
rewards,
)
total_steps += steps
total_rewards += rewards
if train_mode:
return total_steps
else:
return total_rewards / num_rollouts
def update(num_updates):
rl_losses_agg = {}
# print(num_updates)
for update in range(num_updates):
# sample random RL batch: in transitions
batch = ptu.np_to_pytorch_batch(policy_storage.random_episodes(batch_size))
# RL update
rl_losses = agent.update(batch)
for k, v in rl_losses.items():
if update == 0: # first iterate - create list
rl_losses_agg[k] = [v]
else: # append values
rl_losses_agg[k].append(v)
# statistics
for k in rl_losses_agg:
rl_losses_agg[k] = np.mean(rl_losses_agg[k])
return rl_losses_agg
# -
# ## Train and Evaluate the agent: only costs < 20 min
# +
policy_storage = SeqReplayBuffer(
max_replay_buffer_size=int(buffer_size),
observation_dim=obs_dim,
action_dim=act_dim,
sampled_seq_len=sampled_seq_len,
sample_weight_baseline=0.0,
)
env_steps = collect_rollouts(
num_rollouts=num_init_rollouts_pool, random_actions=True, train_mode=True
)
_n_env_steps_total += env_steps
# evaluation parameters
last_eval_num_iters = 0
log_interval = 5
eval_num_rollouts = 10
learning_curve = {
"x": [],
"y": [],
}
while _n_env_steps_total < n_env_steps_total:
env_steps = collect_rollouts(num_rollouts=num_rollouts_per_iter, train_mode=True)
_n_env_steps_total += env_steps
train_stats = update(int(num_updates_per_iter * env_steps))
current_num_iters = _n_env_steps_total // (
num_rollouts_per_iter * max_trajectory_len
)
if (
current_num_iters != last_eval_num_iters
and current_num_iters % log_interval == 0
):
last_eval_num_iters = current_num_iters
average_returns = collect_rollouts(
num_rollouts=eval_num_rollouts,
train_mode=False,
random_actions=False,
deterministic=True,
)
learning_curve["x"].append(_n_env_steps_total)
learning_curve["y"].append(average_returns)
print(_n_env_steps_total, average_returns)
# -
# ## Draw the learning curve
# +
import matplotlib.pyplot as plt
plt.plot(learning_curve["x"], learning_curve["y"])
plt.xlabel("env steps")
plt.ylabel("return")
plt.show()
# -
| example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Linear Regression with Scikit-Learn
# ### importing libraries
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
from scipy import stats
from sklearn import preprocessing
from sklearn import datasets, linear_model
from sklearn.model_selection import train_test_split, cross_val_score,KFold
# -
# ### Reading Data
column_names = ['CRIM', 'ZN', 'INDUS', 'CHAS', 'NOX', 'RM', 'AGE', 'DIS', 'RAD', 'TAX', 'PTRATIO', 'B', 'LSTAT', 'MEDV']
dataset = pd.read_csv('housing.csv', header=None, delimiter=r"\s+", names=column_names)
dataset = dataset.sample(frac=1).reset_index(drop=True)
dataset.head(10)
dataset.describe()
# ### Preprocessing
# ### Plot Data Statistics
fig,axs = plt.subplots(ncols = 7, nrows = 2, figsize = (20,10))
index = 0
axs = axs.flatten()
for k,v in dataset.items():
sns.boxplot(y=k,data=dataset,ax=axs[index])
index += 1
plt.tight_layout(pad=0.4, w_pad = 0.5, h_pad=0.5)
# ### Checking Outliers
for k, v in dataset.items():
q1 = v.quantile(0.25)
q3 = v.quantile(0.75)
irq = q3 - q1
v_col = v[(v <= q1 - 1.5 * irq) | (v >= q3 + 1.5 * irq)]
perc = np.shape(v_col)[0] * 100.0 / np.shape(dataset)[0]
print("Column %s outliers = %.2f%%" % (k, perc))
# ### Removing Outliers
dataset = dataset[~(dataset['MEDV'] >= 40.0)]
print(np.shape(dataset))
fig, axs = plt.subplots(ncols=7, nrows=2, figsize=(20, 10))
index = 0
axs = axs.flatten()
for k,v in dataset.items():
sns.histplot(v, ax=axs[index])
index += 1
plt.tight_layout(pad=0.4, w_pad=0.5, h_pad=5.0)
# ### Print Correlation
plt.figure(figsize=(20, 10))
sns.heatmap(dataset.corr().abs(), annot=True, cmap = 'YlGnBu')
# ### Scaling
std_scaler = preprocessing.StandardScaler()
column_sels = ['LSTAT', 'INDUS', 'NOX', 'PTRATIO', 'RM', 'TAX', 'DIS', 'AGE']
colors = ['navy','green','orange','deeppink','grey','olive','blueviolet','firebrick']
x = dataset.loc[:,column_sels]
y = dataset['MEDV']
x = pd.DataFrame(data=std_scaler.fit_transform(x), columns=column_sels)
fig, axs = plt.subplots(ncols=4, nrows=2, figsize=(20, 10))
index = 0
axs = axs.flatten()
for i, k in enumerate(column_sels):
sns.regplot(y=y, x=x[k], ax=axs[i], color = colors[i])
plt.tight_layout(pad=0.4, w_pad=0.5, h_pad=5.0)
x_train, x_test, y_train, y_test = train_test_split(x,y)
y_train.values.shape
# ### Simple Linear Regression
linear_regressor = linear_model.LinearRegression(normalize=True)
linear_regressor.fit(x_train.values, y_train)
linear_regressor.coef_
pred_y = linear_regressor.predict(x_test.values)
# plt.plot(x_test,pred_y,label='Linear Regression')
plt.scatter(range(len(pred_y)),pred_y,color='orange',label='prediction')
plt.scatter(range(len(y_test)),y_test,color='navy',label='true test label')
plt.legend()
MSE = (((pred_y - y_test)**2).mean())
MSE
LR = linear_model.LinearRegression()
kf = KFold(n_splits=5)
scores = cross_val_score(LR,x,y,cv=kf,scoring='neg_mean_squared_error')
print("MSE: %0.2f (+/- %0.2f)" % (scores.mean(), scores.std()))
pred_y = linear_regressor.predict(x_test.values)
# plt.plot(x_test,pred_y,label='Linear Regression')
plt.scatter(range(len(pred_y)),pred_y,color='orange',label='prediction')
plt.scatter(range(len(y_test)),y_test,color='navy',label='true test label')
plt.legend()
# ### Ridge Regression
linear_regressor = linear_model.Ridge(True)
linear_regressor.fit(x_train.values, y_train)
linear_regressor.coef_
pred_y = linear_regressor.predict(x_test.values)
# plt.plot(x_test,pred_y,label='Linear Regression')
plt.scatter(range(len(pred_y)),pred_y,color='orange',label='prediction')
plt.scatter(range(len(y_test)),y_test,color='navy',label='true test label')
plt.legend()
MSE = (((pred_y - y_test)**2).mean())
MSE
# ### Lasso Regression
linear_regressor = linear_model.Lasso()
linear_regressor.fit(x_train.values, y_train)
linear_regressor.coef_
pred_y = linear_regressor.predict(x_test.values)
# plt.plot(x_test,pred_y,label='Linear Regression')
plt.scatter(range(len(pred_y)),pred_y,color='orange',label='prediction')
plt.scatter(range(len(y_test)),y_test,color='navy',label='true test label')
plt.legend()
MSE = (((pred_y - y_test)**2).mean())
MSE
# ### Elastic Net
linear_regressor = linear_model.ElasticNet()
linear_regressor.fit(x_train.values, y_train)
linear_regressor.coef_
pred_y = linear_regressor.predict(x_test.values)
# plt.plot(x_test,pred_y,label='Linear Regression')
plt.scatter(range(len(pred_y)),pred_y,color='orange',label='prediction')
plt.scatter(range(len(y_test)),y_test,color='navy',label='true test label')
plt.legend()
MSE = (((pred_y - y_test)**2).mean())
MSE
| Assignment_5/Assignment_5.1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Python 3.6+
# We assume you are using Python 3.6+ in this course
# # Question 1
#
# Review list comprehension if needed: https://realpython.com/list-comprehension-python/
#
# The formula for list comprehension is: `new_list = [expression for member in iterable (if conditional)]`
#
# You need to do the following:
# 1. use a loop to create a list of 5 cube numbers and print the list:[0, 1, 8, 27, 64]
# 2. use list comprehension to create the same list
# 3. find the postions of all vowels in a sentence using list comprehension
# +
# use a loop to create a list
cubes = []
a = 0
for i in range(5):
b=(a**3)
cubes.append(b)
a+=1
print(cubes)
# +
# use list comprehension to create the same list
cubes = [0, 1, 8, 27, 64]
# +
# find the positions of all vowels in the following sentence using list comprehension
sentence = "Talk is cheap. Show me the code - <NAME>"
vowels = ("aeiouAEIOU")
position = 0
for char in sentence:
if char in vowels:
print(char, position)
position += 1
# -
# # Question 2
# You need to use to format strings
#
# Write a program using the "f-strings" (https://realpython.com/python-f-strings/), conditional statements, user input function to convert temperatures to and from Celsius, Fahrenheit. [Formula: Celsius/5 = (Fahrenheit – 32)/9]
#
# Hint: you may need int() function
#
# An example program output:
# ```
# Please enter the temperature: 60
# Is this Celsius or Fahrenheit? C
# 60C is 140 in Fahrenheit
# ```
#
# Another example:
# ```
# Please enter the temperature: 45
# Is this Celsius or Fahrenheit? F
# 45F is 7 in Celsius
# ```
# an example of f-strings and user input function
username = input('What\'s your name?')
print(f'Welcome! {username}')
# +
temperature = int(input('Please enter the temperature: '))
type = input('Is this Celsius or Fahrenheit? ')
if type == 'C':
new = int(temperature*(9/5)+32)
other = 'Fahrenheit'
round(new,2)
else:
new = int((temperature-32)*(5/9))
other = 'Celcius'
print(f'{temperature}{type} is {new} in {other}')
# -
| assignment1_python_finished.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.0 64-bit (''mun'': conda)'
# name: python3
# ---
# # Branching using Conditional Statements and Loops in Python
#
# 
#
# ### Part 3 of "Data Analysis with Python: Zero to Pandas"
# 본 자습서는 다음 주제를 다룹니다.:
#
# - `if`, `else`, `elif` 을 이용한 분기
# - 중첩 조건문과 `if` 표현식
# - `while` 반복문
# - `for` 반복문
# - 중첩 반복문, `break` 과 `continue` 설명
# ### How to run the code
#
# This tutorial is an executable [Jupyter notebook](https://jupyter.org). You can _run_ this tutorial and experiment with the code examples in a couple of ways: *using free online resources* (recommended) or *on your computer*.
#
# #### Option 1: Running using free online resources (1-click, recommended)
#
# The easiest way to start executing the code is to click the **Run** button at the top of this page and select **Run on Binder**. You can also select "Run on Colab" or "Run on Kaggle", but you'll need to create an account on [Google Colab](https://colab.research.google.com) or [Kaggle](https://kaggle.com) to use these platforms.
#
#
# #### Option 2: Running on your computer locally
#
# To run the code on your computer locally, you'll need to set up [Python](https://www.python.org), download the notebook and install the required libraries. We recommend using the [Conda](https://docs.conda.io/projects/conda/en/latest/user-guide/install/) distribution of Python. Click the **Run** button at the top of this page, select the **Run Locally** option, and follow the instructions.
#
# > **Jupyter Notebooks**: This tutorial is a [Jupyter notebook](https://jupyter.org) - a document made of _cells_. Each cell can contain code written in Python or explanations in plain English. You can execute code cells and view the results, e.g., numbers, messages, graphs, tables, files, etc., instantly within the notebook. Jupyter is a powerful platform for experimentation and analysis. Don't be afraid to mess around with the code & break things - you'll learn a lot by encountering and fixing errors. You can use the "Kernel > Restart & Clear Output" menu option to clear all outputs and start again from the top.
# ## `if`, `else` ,`elif` 이란
#
# 프로그래밍 언어의 가장 강력한 기능 중 하나는 하나 이상의 조건이 참인지 여부에 따라 의사 결정을 내리고 문장을 실행하는 능력인 *분기*입니다.
#
# ### The `if` statement
#
# Python 에서는 `if`문을 사용하여 분기를 구현합니다.
#
# ```
# if condition:
# statement1
# statement2
# ```
#
# `조건`은 값, 변수, 표현식이 될 수 있다. 조건이 `True`로 평가되면 **if block**내에서 코드가 실행됩니다.`statement1` ,`statement2`등의 앞에 4개의 공백이 있습니다. 공백은 Python 에서 이러한 문장이 위의 `if`문과 연관되어 있음을 알려줍니다. 공백을 추가하여 코드를 구성하는 이 기술을 **인덱션** 이라고 합니다.
#
# > **Indentation**: Python 은 코드 구조를 정의하기위 **인덱션**에 크게 의존합니다. 라서 Python 코드를 쉽게 읽고 이해할 수 있습니다. 덴트를 제대로 사용하지 않으면 문제가 발생할 수 있습니다. 줄의 시작 부분에 커서를 놓고 'Tab' 키를 한 번 눌러 4개의 공백을 추가하여 코드를 입력하세요. Tab을 다시 누르면 코드가 4칸 더 들어가고 Shift+Tab을 누르면 4칸 더 들어갑니다.
#
#
# 예를 들어, 특정 숫자가 짝수이면 메시지를 확인하고 인쇄하기 위한 코드를 작성하겠습니다.
a_number = 34
if a_number % 2 == 0:
print("We're inside an if block")
print('The given number {} is even.'.format(a_number))
# `a_number`를 2로 나눈 나머지를 계산하려면 계수 연산자 `%`를 사용합니다. 그럼 다음 나머지가 0인지 비교 연산자 `==` 체그를 사용하여 짝수인지 여부를 판단합니다.
#
# `34`는 2로 나누기 때문에 `a_number % 2 == 0`이라는 표현은 `True` 평가되므로 `if`문 내부가 코드가 실행된다
# 또한 문자열 `format`메서드를 사용하여 메시지에 숫자를 포함한다.
#
# 위의 내용을 홀수로 다시 한 번 시도해 봅시다.
another_number = 33
if another_number % 2 == 0:
print('The given number {} is even.'.format(a_number))
# 'another_number %2 == 0' 조건이 'False'로 평가되므로 메시지가 인쇄되지 않습니다.
#
# ### `else` 문
#
# 위 예에 없는 경우 다른 메시지를 출력할 수 있습니다. 이것은 `else` 문을 통해 구현할 수 있다.:
#
# ```
# if condition:
# statement1
# statement2
# else:
# statement4
# statement5
#
# ```
#
# `조건`이 `True`가 되면, `if` 내부 코드가 실행된다. `False`가 될 경우, `else`문이 실행된다.
a_number = 34
if a_number % 2 == 0:
print('The given number {} is even.'.format(a_number))
else:
print('The given number {} is odd.'.format(a_number))
another_number = 33
if another_number % 2 == 0:
print('The given number {} is even.'.format(another_number))
else:
print('The given number {} is odd.'.format(another_number))
# 아래는 `in` 연산자를 사용하여 tuple의 값들은 확인하는 예제 코드
the_3_musketeers = ('Athos', 'Porthos', 'Aramis')
a_candidate = "D'Artagnan"
if a_candidate in the_3_musketeers:
print("{} is a musketeer".format(a_candidate))
else:
print("{} is not a musketeer".format(a_candidate))
# ### `elif`문
# Python은 연쇄적인 조건문을 위해 `elif`문을 제공한다. 조건은 하나씩 평가된다.
# 첫번째 조건은 `True`이면 해당 조건문은 실행이되나 나머지 조건영역은 실행되지 않는다. `if`,`elif`,`elif` ... 연쇄적으로 있는 상황에서 조건이 `True`가 되는 첫번째 조건문만 실행된다.
today = 'Wednesday'
if today == 'Sunday':
print("Today is the day of the sun.")
elif today == 'Monday':
print("Today is the day of the moon.")
elif today == 'Tuesday':
print("Today is the day of Tyr, the god of war.")
elif today == 'Wednesday':
print("Today is the day of Odin, the supreme diety.")
elif today == 'Thursday':
print("Today is the day of Thor, the god of thunder.")
elif today == 'Friday':
print("Today is the day of Frigga, the goddess of beauty.")
elif today == 'Saturday':
print("Today is the day of Saturn, the god of fun and feasting.")
# 위의 예제에서는 처음 3개의 조건이 `False `로 메세지가 출력되지 않는다. 4번째 조건은 `True`가 되어 해당 조건문에 해당되는 메세지가 실행됩니다. 남은 조건문은 무시됩니다.
# 실제로 나머지 조건들이 무시되는지 확인하기 위하여 다른 예들을 살펴봅시다.
a_number = 15
if a_number % 2 == 0:
print('{} is divisible by 2'.format(a_number))
elif a_number % 3 == 0:
print('{} is divisible by 3'.format(a_number))
elif a_number % 5 == 0:
print('{} is divisible by 5'.format(a_number))
elif a_number % 7 == 0:
print('{} is divisible by 7'.format(a_number))
# 이전 `a_number % 3 == 0` 조건이 참이되어 `a_number % 5 == 0` 조건이 무시되고 이로인해 `15 is divisible by 5` 는 출력되지 않습니다.
# `if`문만 사용했을 때, 각 조건이 독립적으로 평가되는 것과 달리 `if` ,`elif`문에서는 선행 조건이 뒤 조건을 판정할지 영향을 줍니다.
if a_number % 2 == 0:
print('{} is divisible by 2'.format(a_number))
if a_number % 3 == 0:
print('{} is divisible by 3'.format(a_number))
if a_number % 5 == 0:
print('{} is divisible by 5'.format(a_number))
if a_number % 7 == 0:
print('{} is divisible by 7'.format(a_number))
# ### Using `if`, `elif`, and `else` together
#
# `else`문은 `if` , `elif`...문의 마지막에 사용된다. `else` 블럭은 선행조건문에서 `True` 판정이 없을 때 실행됩니다.
#
a_number = 49
if a_number % 2 == 0:
print('{} is divisible by 2'.format(a_number))
elif a_number % 3 == 0:
print('{} is divisible by 3'.format(a_number))
elif a_number % 5 == 0:
print('{} is divisible by 5'.format(a_number))
else:
print('All checks failed!')
print('{} is not divisible by 2, 3 or 5'.format(a_number))
# 조건은 논리 연산자 `and`, `or` , `not` 와 같이 결합될 수 있다. 논리 연산자에 대한 설명은 다음을 참조하세요. [first tutorial](https://jovian.ml/aakashns/first-steps-with-python/v/4#C49).
a_number = 12
if a_number % 3 == 0 and a_number % 5 == 0:
print("The number {} is divisible by 3 and 5".format(a_number))
elif not a_number % 5 == 0:
print("The number {} is not divisible by 5".format(a_number))
# ### Non-Boolean Conditions
#
# 조건이 꼭 boolean형일 필요는 없다. 실제로, 어떤 값이든 조건이 될 수 잇다. 조건은 `bool`함수를 통해 자동으로 boolean 타입으로 변환된다. **falsy** 값인 `0`,`0`, `''`, `{}`, `[]`,`False` 는 `False`로 나머지는 `True`가 됩니다.
if '':
print('The condition evaluted to True')
else:
print('The condition evaluted to False')
if 'Hello':
print('The condition evaluted to True')
else:
print('The condition evaluted to False')
if { 'a': 34 }:
print('The condition evaluted to True')
else:
print('The condition evaluted to False')
if None:
print('The condition evaluted to True')
else:
print('The condition evaluted to False')
# ### Nested conditional statements
#
# `if` 문 안에 다시 `if`문이 포함될 수 있다. 이러한 방식을 `nesting` 이라고 부르며, 특정 조건이 `True`인 상황에서 조건을 확인하기 위하여 사용됩니다.
a_number = 15
if a_number % 2 == 0:
print("{} is even".format(a_number))
if a_number % 3 == 0:
print("{} is also divisible by 3".format(a_number))
else:
print("{} is not divisibule by 3".format(a_number))
else:
print("{} is odd".format(a_number))
if a_number % 5 == 0:
print("{} is also divisible by 5".format(a_number))
else:
print("{} is not divisibule by 5".format(a_number))
# > `if`,`else` 문 중첩은 종종 사람들이 혼란을 주므로. 가능한한 중첩을 피하고 1~2단계정도만 제한하는 것이 좋습니다.
# ### Shorthand `if` conditional expression
#
# `if` 문은 조건을 확인하고 조건에 따라 변수값을 설정하는데 자주 사용됩니다\.
# +
a_number = 13
if a_number % 2 == 0:
parity = 'even'
else:
parity = 'odd'
print('The number {} is {}.'.format(a_number, parity))
# -
# Python 은 한줄에 조건을 작성하기 위하여, 축약형을 제공 한다. 이것을 **조건표현식**이라고 하며, 이를 위한 **삼항 조건 연산자** . 조건표현식은 아래 문법을 따른다.
#
# ```
# x = true_value if condition else false_value
# ```
# 이것을 `if` - `else`로 구현하면 다음과 같다.
#
# ```
# if condition:
# x = true_value
# else:
# x = false_value
# ```
parity = 'even' if a_number % 2 == 0 else 'odd'
print('The number {} is {}.'.format(a_number, parity))
# ### Statements and Expressions
#
# Python에서 조건식은 **명령문** 과 **표현식**사이의 차이를 강조합니다.
#
# > **명령문**: 명령문은 실행이 가능한 함수이다. 지금가지 작성한 모든 코드는 변수 할당, 함수 호출, if, elif, for, while 등을 사용한 조건문, 반복문등이 명령문입니다.
#
# > **표현식**: 표현식은 값을 평가하는 코드이다. 예로 다양한 데이터 유형의 값, 산술 식, 조건, 변수, 함수 호출, 조건식 등이 있습니다.
#
# 대부분 표현식은 명령문으로 실행될수 있으나, 모든 명령문이 표현식이 되지는 않는다. 예로들어, 값으로 평가되지 않는 정규 `if` 명령문은 표현식이 아니다.
# 이것은 코드안에 일부분이 실행될 뿐이다. 마찬가지로 루프는 함수 정의는 식이 아닙니다.
#
# 표현식은 `=` 연산자 우측에 들어갈 수 있는 모든 것들을 말합니다. 이 방식으로 표현식의 여부를 확인할 수 있습니다.
# 표현식이 아닌 것을 할당하려고 하면 구문 오류가 발생합니다.
# if statement
result = if a_number % 2 == 0:
'even'
else:
'odd'
# if expression
result = 'even' if a_number % 2 == 0 else 'odd'
# ### `pass` 문
#
# `if`문은 빈값이 들어갈 수 없고 최소한 하나 이상의 명령문이 들어가야 합니다. `pass`문을 사용하여 어떠한 명령도 수행하지 않고도 오류가 발생하지 않을 수 있습니다.
a_number = 9
if a_number % 2 == 0:
elif a_number % 3 == 0:
print('{} is divisible by 3 but not divisible by 2')
if a_number % 2 == 0:
pass
elif a_number % 3 == 0:
print('{} is divisible by 3 but not divisible by 2'.format(a_number))
# ## Iteration with `while` loops
#
# 조건문이 사용가능한 프로그래밍 언어들의 큰 강점은 하나 이상의 명령문을 여러 번 실행시킬 수 있다는 것입니다.
# 이러한 특징을 **반복문** 이라고 하며, Python에서는 `while` 반복문과 `for` 반복문이 있습니다.
#
# `while`반복문에 사용법은 아래와 같습니다:
#
# ```
# while condition:
# statement(s)
# ```
# `while`문 안에 명령은 반복문의 `조건`이 `True`인한 반복해서 실행된다. 일반적으로, `while`문안에는 특정 반복횟수에 도달하면 조건상태를 `False`로 만드는 명령문이 존재합니다.
#
# `while`문을 이용해서 팩토리얼 100을 구해봅시다.
# +
result = 1
i = 1
while i <= 100:
result = result * i
i = i+1
print('The factorial of 100 is: {}'.format(result))
# -
# 위 코드의 작동 방식은 다음과 같습니다.<br>
#
# * 두 변수인 `result`와 `i`를 초기화합니다. `result`에는 최종계산결과값이 들어갑니다. `i`는 다음 숫자에 `result`를 곱할 때 사용됩니다.
#
# * 조건 `i <= 100`은 `True` 이다. (`i`의 초기값이 `1` 이므로) 그러므로 `while`문 내부 코드는 실행된다.
#
# * `result`는 `result * i`로 업데이트 되고, `i` 값은 1이 증가하여 `2`가 된다.
#
# * 이 시점에서, 조건 `i <= 100`가 판단된다. 조건은 `True` 상태이므로, `result`값은 `result * i`값으로 업데이트 되고 , `i`는 1이 증가하여 `3`이 된다.
#
# * 이러한 과정이 `i`가 `101`이 되어 조건이 `False`가 될 떼 까지 반복된다. 반복문이 끝나면 `print` 명령문이 실행된다.
#
# * `result`가 왜 마지막에 100의 요인 값을 포함하는지 알겠습니까?
#
# 셀 상단에 *command '%%time'을 추가하여 셀 실행 시간을 확인할 수 있습니다. `100`, `1000`, `10000`, `10000` 등의 요인을 계산하는 데 얼마나 걸리는지 확인해 보십시오.
# +
# %%time
result = 1
i = 1
while i <= 1000:
result *= i # same as result = result * i
i += 1 # same as i = i+1
print(result)
# -
# `while` 문을 사용한 예제
# +
line = '*'
max_length = 10
while len(line) <= max_length:
print(line)
line += "*"
while len(line) > 0:
print(line)
line = line[:-1]
# -
# 연습 삼아 잠시 동안 `while` 루프를 사용하여 다음 패턴을 인쇄해 보십시오:
#
# ```
# *
# **
# ***
# ****
# *****
# ******
# *****
# ****
# ***
# **
# *
# ```
#
# 여기 또 하나가 있는데, 둘을 합치면 아래와 같습니다:
#
#
# ```
# *
# ***
# *****
# *******
# *********
# ***********
# *********
# *******
# *****
# ***
# *
# ```
# ### Infinite Loops
#
# `while` 루프의 조건이 항상 `True`일 경우, 루프가 무한히 반복되어 코드가 완료되지않는다. 이것을 `무한루프` 라고 합니다.
#
# 코드가 무한루프에 빠진경우, 도구 모음의 `stop` 버튼을 누르거나 메뉴 모음에서 `Kernel > Interrupt` 를 선택합니다. 그러면 코드 실행이 중단됩니다.
# +
# INFINITE LOOP - INTERRUPT THIS CELL
result = 1
i = 1
while i <= 100:
result = result * i
# forgot to increment i
# +
# INFINITE LOOP - INTERRUPT THIS CELL
result = 1
i = 1
while i > 0 : # wrong condition
result *= i
i += 1
# -
# ### `break` , `continue` 명령문
#
# 반복문 안에 `break`명렴문을 사용하여 반복문을 강제로 탈출할 수 있습니다.
# +
i = 1
result = 1
while i <= 100:
result *= i
if i == 42:
print('Magic number 42 reached! Stopping execution..')
break
i += 1
print('i:', i)
print('result:', result)
# -
#
# `continue`명령문을 통하여 현재 루프를 건너띄고, 바로 다음 루프로 넘어갈 수 있습니다.
# +
i = 1
result = 1
while i < 20:
i += 1
if i % 2 == 0:
print('Skipping {}'.format(i))
continue
print('Multiplying with {}'.format(i))
result = result * i
print('i:', i)
print('result:', result)
# -
#
# > **Logging**: 코드 안에 `print`문을 추가하여 단계의 변화에 따른 변수값을 검사하는 과정을 **Loagging** 이라고 합니다.
# ## `for` 문을 통한 반복문
# `for` 문을 통하여 순차적 자료형(list,tuples,dictionaries,string 등)을 반복하는데 사용된다. 예제는 다음과 같습니다:
#
# ```
# for value in sequence:
# statement(s)
# ```
#
# 반복문 안에 명령문은 순차적 자료형의 각 요소에 대해 한번 씩 수행됩니다.
# +
days = ['Monday', 'Tuesday', 'Wednesday', 'Thursday', 'Friday']
for day in days:
print(day)
# -
# 다른 예제를 예로 들겠습니다.
# Looping over a string
for char in 'Monday':
print(char)
# Looping over a tuple
for fruit in ['Apple', 'Banana', 'Guava']:
print("Here's a fruit:", fruit)
# +
# Looping over a dictionary
person = {
'name': '<NAME>',
'sex': 'Male',
'age': 32,
'married': True
}
for key in person:
print("Key:", key, ",", "Value:", person[key])
# -
# Dictionary에 경우 `for`반복문에서 키에 대한 반복문이 발생합니다. `.values` 메서드를 사용하여, 값에 직접 전근할 수 있습니다.
# `.items` 메서드를 사용하여 키-값에 대해 직접 반복할 수 있습니다.
for value in person.values():
print(value)
for key_value_pair in person.items():
print(key_value_pair)
# tuple은 키-값 쌍으로 되어 있으므로, 키와 값을 분리해서 변수로 가져올 수 있습니다.
for key, value in person.items():
print("Key:", key, ",", "Value:", value)
# ### Iterating using `range` and `enumerate`
#
# `range` 함수는 `for`문에서 연속적인 수를 생성하는데 사용됩니다.
# 해당 함수는 3가지 방식으로 사용 될 수 있습니다 :
#
# * `range(n)` - `0` 부터 `n-1` 를 생성합니다.
# * `range(a, b)` - `a` to `b-1` 를 생성합니다.
# * `range(a, b, step)` - `a` 부터 `b-1` 까지 `step`만큼 건너띄면 생성합니다.
for i in range(7):
print(i)
for i in range(3, 10):
print(i)
for i in range(3, 14, 4):
print(i)
# 반복문이 진행되는 동안 요소의 인덱스를 추적해야 하는 경우 list 반복에서 `range` 를 사용합니다.
# +
a_list = ['Monday', 'Tuesday', 'Wednesday', 'Thursday', 'Friday']
for i in range(len(a_list)):
print('The value at position {} is {}.'.format(i, a_list[i]))
# -
# `enumerate`를 사용하여 위와 같은 내용을 구현할 수 있습니다.
for i, val in enumerate(a_list):
print('The value at position {} is {}.'.format(i, val))
# ### `break`, `continue` and `pass` statements
# `while`문에서와 같이, `for`의 반복문에서도 `break`,`continue`,`pass` 가 동일하게 작동합니다.
weekdays = ['Monday', 'Tuesday', 'Wednesday', 'Thursday', 'Friday']
for day in weekdays:
print('Today is {}'.format(day))
if (day == 'Wednesday'):
print("I don't work beyond Wednesday!")
break
for day in weekdays:
if (day == 'Wednesday'):
print("I don't work on Wednesday!")
continue
print('Today is {}'.format(day))
# `if` 문처럼 반복문에서 내부 명령문이 없을 수 없으므로 반복 내에서 명령문을 실행하지 않으려면 pass 문을 사용할 수 있습니다.
for day in weekdays:
pass
# ### Nested `for` and `while` loops
#
# 조건문과 마찬가지로 반복문은 다른 반복문 내부에 선언될 수 있습니다. 이 기능은 lists, dictionaries등을 반복시킬 때 유용합니다.
# +
persons = [{'name': 'John', 'sex': 'Male'}, {'name': 'Jane', 'sex': 'Female'}]
for person in persons:
for key in person:
print(key, ":", person[key])
print(" ")
# +
days = ['Monday', 'Tuesday', 'Wednesday']
fruits = ['apple', 'banana', 'guava']
for day in days:
for fruit in fruits:
print(day, fruit)
# -
# With this, we conclude our discussion of branching and loops in Python.
# ## Further Reading and References
#
# 반복문에 대해 좀 더 자세히 알고 싶다면 아래에 자료를 참조하길 바랍니다:
#
# * Python Tutorial at W3Schools: https://www.w3schools.com/python/
# * Practical Python Programming: https://dabeaz-course.github.io/practical-python/Notes/Contents.html
# * Python official documentation: https://docs.python.org/3/tutorial/index.html
#
# You are now ready to move on to the next tutorial: [Writing Reusable Code Using Functions in Python](https://jovian.ai/aakashns/python-functions-and-scope)
#
# ## Questions for Revision
#
# Try answering the following questions to test your understanding of the topics covered in this notebook:
#
# 1. What is branching in programming languages?
# 2. What is the purpose of the `if` statement in Python?
# 3. What is the syntax of the `if` statement? Give an example.
# 4. What is indentation? Why is it used?
# 5. What is an indented block of statements?
# 6. How do you perform indentation in Python?
# 7. What happens if some code is not indented correctly?
# 8. What happens when the condition within the `if` statement evaluates to `True`? What happens if the condition evaluates for `false`?
# 9. How do you check if a number is even?
# 10. What is the purpose of the `else` statement in Python?
# 11. What is the syntax of the `else` statement? Give an example.
# 12. Write a program that prints different messages based on whether a number is positive or negative.
# 13. Can the `else` statement be used without an `if` statement?
# 14. What is the purpose of the `elif` statement in Python?
# 15. What is the syntax of the `elif` statement? Give an example.
# 16. Write a program that prints different messages for different months of the year.
# 17. Write a program that uses `if`, `elif`, and `else` statements together.
# 18. Can the `elif` statement be used without an `if` statement?
# 19. Can the `elif` statement be used without an `else` statement?
# 20. What is the difference between a chain of `if`, `elif`, `elif`… statements and a chain of `if`, `if`, `if`… statements? Give an example.
# 21. Can non-boolean conditions be used with `if` statements? Give some examples.
# 22. What are nested conditional statements? How are they useful?
# 23. Give an example of nested conditional statements.
# 24. Why is it advisable to avoid nested conditional statements?
# 25. What is the shorthand `if` conditional expression?
# 26. What is the syntax of the shorthand `if` conditional expression? Give an example.
# 27. What is the difference between the shorthand `if` expression and the regular `if` statement?
# 28. What is a statement in Python?
# 29. What is an expression in Python?
# 30. What is the difference between statements and expressions?
# 31. Is every statement an expression? Give an example or counterexample.
# 32. Is every expression a statement? Give an example or counterexample.
# 33. What is the purpose of the pass statement in `if` blocks?
# 34. What is iteration or looping in programming languages? Why is it useful?
# 35. What are the two ways for performing iteration in Python?
# 36. What is the purpose of the `while` statement in Python?
# 37. What is the syntax of the `white` statement in Python? Give an example.
# 38. Write a program to compute the sum of the numbers 1 to 100 using a while loop.
# 39. Repeat the above program for numbers up to 1000, 10000, and 100000. How long does it take each loop to complete?
# 40. What is an infinite loop?
# 41. What causes a program to enter an infinite loop?
# 42. How do you interrupt an infinite loop within Jupyter?
# 43. What is the purpose of the `break` statement in Python?
# 44. Give an example of using a `break` statement within a while loop.
# 45. What is the purpose of the `continue` statement in Python?
# 46. Give an example of using the `continue` statement within a while loop.
# 47. What is logging? How is it useful?
# 48. What is the purpose of the `for` statement in Python?
# 49. What is the syntax of `for` loops? Give an example.
# 50. How are for loops and while loops different?
# 51. How do you loop over a string? Give an example.
# 52. How do you loop over a list? Give an example.
# 53. How do you loop over a tuple? Give an example.
# 54. How do you loop over a dictionary? Give an example.
# 55. What is the purpose of the `range` statement? Give an example.
# 56. What is the purpose of the `enumerate` statement? Give an example.
# 57. How are the `break`, `continue`, and `pass` statements used in for loops? Give examples.
# 58. Can loops be nested within other loops? How is nesting useful?
# 59. Give an example of a for loop nested within another for loop.
# 60. Give an example of a while loop nested within another while loop.
# 61. Give an example of a for loop nested within a while loop.
# 62. Give an example of a while loop nested within a for loop.
#
#
| 3) python-branching-and-loops/python-branching-and-loops.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="gpGPl3rxD5bt" executionInfo={"status": "ok", "timestamp": 1625279906158, "user_tz": -330, "elapsed": 11, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11106137188993670892"}}
import numpy as np
import matplotlib.pyplot as plt
from IPython.display import Image
# %matplotlib inline
# + colab={"base_uri": "https://localhost:8080/", "height": 339} id="CMmNelT6-rAq" executionInfo={"status": "ok", "timestamp": 1624672409145, "user_tz": -330, "elapsed": 716, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04194286040197059483"}} outputId="11f2481a-6132-4f6f-a4f8-bd6bcb084d20"
Image('djikstra.png')
# + [markdown] id="xjRRn2qs-hse"
# Consider the example problem we discussed in the theory session (easy to verify!). We will implement from scratch Djikstra algorithm to find the cost of traversal from a source node to all other nodes in a given connected graph. Note that we are not finding the shortest path itself. But that will follow
# + id="rkAm7KZ5D5by" executionInfo={"status": "ok", "timestamp": 1625297292159, "user_tz": -330, "elapsed": 1138, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11106137188993670892"}}
class Graph(object):
def __init__(self, nodes, edges): #, edge_cost
self.nodes = nodes
self.adjacency = -np.ones([nodes, nodes])
self.shortest_path_set = [False] * nodes
#populate the adjacency matrix from edges
# format of edges = (node1, node2, edge_cost)
for node1, node2, cost in edges:
self.adjacency[node1, node2] = cost
# dist = 1D array of all distances to source
# check if node is not already in the shortest path set
# output = closest node
def min_distance(self, dist):
return np.argmin(np.array(dist) + 1000*np.array(self.shortest_path_set))
#min = edge_cost.maxsize
# process loop
#for v in range(self.nodes):
# if dist[v] < min:
# min = dist[v]
# min_index = v
def dijkstra(self, src):
#initialize distance array
dist = [1000] * self.nodes
dist[src] = 0
for _ in range(self.nodes):
#dist = [0, 1000, 1000, ..., 1000] -- first iteration
#dist = [0, 8, 5, 2, 1000, 1000, 1000] -- second iteration
# shortest_path_set = [1, 0, 0, 1, 0, 0, 0]
i = self.min_distance(dist)
# Store min distance vertex in shortest path tree
self.shortest_path_set[i] = True
# Update dist value of the neighbors of selected node
# Two conditions to check for each neighbor
# (a) not in shortest path tree (b) cost is now lowered
# first get neighbor list from adjacency matrix
all_nodes = self.adjacency[i,:]
# loop over neighbor list to check for other 2 conditions
# if satisfied, change dist[j]
for j, edge_cost in enumerate(all_nodes):
if edge_cost > 0 and not self.shortest_path_set[j]: # valid neighbor
if dist[i] + edge_cost < dist[j]:
dist[j] = dist[i] + edge_cost
return dist
# + colab={"base_uri": "https://localhost:8080/"} id="H9mxydyT4J4R" executionInfo={"status": "ok", "timestamp": 1625290870879, "user_tz": -330, "elapsed": 600, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11106137188993670892"}} outputId="c98a42bb-34eb-4222-c722-7a6d5354435e"
def min_cost(dist1, dist2):
return np.argmin(dist1 + 1000*dist2)
dist1 = np.array([0, 8, 5, 2, 1000])
dist2 = np.array([1, 0, 0, 0, 0])
min_cost(dist1, dist2)
# + colab={"base_uri": "https://localhost:8080/"} id="2uRghXv4gZbE" executionInfo={"status": "ok", "timestamp": 1625287754532, "user_tz": -330, "elapsed": 442, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11106137188993670892"}} outputId="86d5b578-32ed-4885-bdbb-696fce1141c9"
## reference snippet
edges = [(0, 1, 8), (0, 2, 5), (0, 3, 2), (1, 4, 2), (2, 5, 3), \
(3, 5, 8), (4, 5, 7), (5, 4, 7), (5, 6, 1), (6, 5, 4) ]
#for node1, node2, cost in edges:
# print(f"cost of going from {node1} to {node2} is {cost}")
cost = [lis[2] for lis in edges]
print(cost)
#adjacency = -np.ones([7,7])
#adjacency[2,3] = 14
#print(adjacency)
# + id="QFtApVYQJ9oQ" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1625297298615, "user_tz": -330, "elapsed": 1338, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11106137188993670892"}} outputId="dcae4cea-4933-47aa-bb84-71193b292d2b"
nodes = 7
# (node_A, node_B, edge_cost)
edges = [(0, 1, 8), (0, 2, 5), (0, 3, 2), (1, 4, 2), (2, 1, 1), (2, 5, 3), \
(3, 5, 8), (4, 5, 7), (5, 4, 7), (4, 6, 1), (6, 5, 4) ]
#edge_cost = [lis[2] for lis in edges]
#g = Graph(nodes, edges, edge_cost)
g = Graph(nodes, edges)
#print(g.nodes)
#print(g.adjacency)
dist_to_source = g.dijkstra(0)
#for node, dist in enumerate(g.dijkstra(0)[1:]):
for node, dist in enumerate(dist_to_source):
print(f"Node {node} is at distance {dist}")
# + colab={"base_uri": "https://localhost:8080/", "height": 403} id="MsUN5o6N_nUI" executionInfo={"status": "ok", "timestamp": 1624672400021, "user_tz": -330, "elapsed": 804, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04194286040197059483"}} outputId="92afe440-a5c2-46d8-a35a-75a649e2546a"
Image('astar.png')
# + [markdown] id="UDxJrADc3qoP"
# ## A*
# Let us now modify the graph to accept the 2D co-ordinates of the node. We will use Euclidean distance as the heuristic
# + id="EvoT3Ah0NB1Z"
node_coords = [(0, 0),(2,2),(1,2),
(1,0),(3,3),(3,2),
(4,2)]
# + id="75PNiEgOSzQz" executionInfo={"status": "ok", "timestamp": 1625299353895, "user_tz": -330, "elapsed": 714, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11106137188993670892"}}
# Function to calculate euclidean distance
# (x1, y1), (x2, y2) given
def euclidean(node1, node2):
#write code here
x1, y1 = node1
x2, y2 = node2
return np.sqrt((x1-x2)**2+(y1-y2)**2)
class Graph(object):
def __init__(self, nodes, edges, coords, weight=1.0, heuristic=euclidean):
self.nodes = nodes
self.adjacency = np.zeros([nodes, nodes])
self.shortest_path_set = [False] * nodes
self.heuristic = heuristic
self.coords = coords
self.weight = weight # weight of heuristic
#populate the adjacency matrix from edges
# edges = (node1, node2, edge_cost)
# Input: 1-D distance array to source, destination (x, y)
# output: next node to be selected
# remember criteria is source_cost + weight * heuristic_destination
# node should not be in shortest_path_set
def min_astar_distance(self, dist, dest_coords):
heuristic_cost = np.array([euclidean(n, dest, dest_coords) for n in self.coords])
src_cost = np.array(dist)
costs = src_cost + self.weight*heuristic_cost + 1000*np.array(self.shortest_path_set)
return np.argmin(costs)
def astar(self, src, dest):
#initialize distance array
dist = [1000] * self.nodes
dist[src] = 0
#get the destination (x,y)
dest_coords = self.coords[dest]
for _ in range(self.nodes):
i = self.min_distance(dist, dest_coords)
# Store min distance vertex in shortest path tree
self.shortest_path_set[i] = True
# Update dist value of the neighbors of selected node
# Two conditions to check for each neighbor
# (a) not in shortest path tree (b) cost is now lowered
# first get neighbor list from adjacency matrix
neighbors = self.adjacency[i,:]
# loop over neighbor list to check for other 2 conditions
# if satisfied, change dist[j]
for j, edge_cost in enumerate(neighbors):
if edge_cost > 0 and not self.shortest_path_set[j]: # valid neighbor
if dist[i] + edge_cost < dist[j]:
dist[j] = dist[i] + edge_cost
# find heuristic cost from all nodes to destination
# use list comprehension
heuristic_cost = abs(dest_coords.x) + abs(dest_coords.y)
return dist, heuristic_cost
# + colab={"base_uri": "https://localhost:8080/"} id="kBzF2BKHCQi4" executionInfo={"status": "ok", "timestamp": 1624673993723, "user_tz": -330, "elapsed": 389, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04194286040197059483"}} outputId="0f67c09c-794e-485b-f3c2-87f59e891326"
nodes = 7
# (node_A, node_B, edge_cost)
edges = [(0, 1, 8), (0, 2, 5), (0, 3, 2), (1, 4, 2), (2, 5, 3), \
(3, 5, 8), (4, 5, 7), (5, 4, 7), (5, 6, 1), (6, 5, 4) ]
node_coords = [(0, 0),(2,2),(1,2),(1,0),(3,3),(3,2),(4,2)]
g = Graph(nodes, edges, node_coords)
cost, heuristic = g.astar(0, 6)
for node, (dist, heur) in enumerate(zip(cost, heuristic)):
print(f"Node {node} is at distance {dist}")
print(f"Node {node} heuristic is {heur}")
# + [markdown] id="Fy-u1lwa75pt"
# Notice that this is a very simple implementation to get the costs of all nodes to the source node. We can make 2 changes
#
# 1. We did not get the predecessors of each node.
# Predecessors list is what will help us determine the path. Can you change the code to print out the predecessors as well?
# 2. In general we have to calculate only the path to the destination (not all nodes) as it is computationally expensive. What do you think should be the convergence criteria? Use it to find the shortest path to Node 5 instead of Node
#
# Feel free to experiment with other heuristics like (a) L-1 norm (b) number of edges
#
#
# + id="SKSHmljmI5Ln"
| week2/mayank_h/Q2 - Q/Attempt1_filesubmission_search_based_planning_scratch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
import sys,pprint
import os
sys.path.append(os.path.join(os.getcwd(),'..'))
import watson_developer_cloud
DISCOVERY_USERNAME='CHANGE_ME'
DISCOVERY_PASSWORD='<PASSWORD>'
pp = pprint.PrettyPrinter(indent=4)
# +
discovery = watson_developer_cloud.DiscoveryV1(
'2016-11-07',
username=DISCOVERY_USERNAME,
password=<PASSWORD>)
environments = discovery.get_environments()
pp.pprint(environments)
news_environments = [x for x in environments['environments'] if
x['name'] == 'Watson News Environment']
news_environment_id = news_environments[0]['environment_id']
pp.pprint(news_environment_id)
collections = discovery.list_collections(news_environment_id)
news_collections = [x for x in collections['collections']]
pp.pprint(collections)
# -
pp.pprint(discovery.list_configurations(environment_id=news_environment_id))
default_config_id = discovery.get_default_configuration_id(environment_id=news_environment_id)
pp.pprint(default_config_id)
default_config = discovery.get_configuration(environment_id=news_environment_id, configuration_id=default_config_id)
pp.pprint(default_config)
new_environment = discovery.create_environment(name="new env", description="bogus env")
# +
pp.pprint(new_environment)
if (discovery.get_environment(environment_id=new_environment['environment_id'])['status'] == 'active'):
writable_environment_id = new_environment['environment_id']
new_collection = discovery.create_collection(environment_id=writable_environment_id,
name='Example Collection',
description="just a test")
pp.pprint(new_collection)
#pp.pprint(discovery.get_collections(environment_id=writable_environment_id))
#res = discovery.delete_collection(environment_id='10b733d0-1232-4924-a670-e6ffaed2e641',
# collection_id=new_collection['collection_id'])
# pp.pprint(res)
# -
collections = discovery.list_collections(environment_id=writable_environment_id)
pp.pprint(collections)
with open(os.path.join(os.getcwd(),'..','resources','simple.html')) as fileinfo:
pp.pprint(discovery.test_document(environment_id=writable_environment_id, fileinfo=fileinfo))
with open(os.path.join(os.getcwd(),'..','resources','simple.html')) as fileinfo:
res = discovery.add_document(environment_id=writable_environment_id,
collection_id=collections['collections'][0]['collection_id'],
fileinfo=fileinfo)
pp.pprint(res)
res = discovery.get_collection(environment_id=writable_environment_id,
collection_id=collections['collections'][0]['collection_id'])
pp.pprint(res['document_counts'])
res = discovery.delete_environment(environment_id=writable_environment_id)
pp.pprint(res)
| examples/discovery_v1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="YNXhHEg0yRY2"
# ##**<NAME>**
#
# ##**MSBI - Spring 2021**
# + [markdown] id="6juf4DYYjnhf"
# This notebook focuses on processing CoV2 variants data and building a Convolutional Neural Network model to classify different Covid19 variants.
#
# Signature spike protein was identified by NCBI and virus variants were retrieved from GISAID website using corresponding spike protein.
# + id="2kmaJzMyyMWH"
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from sklearn.preprocessing import LabelEncoder, OneHotEncoder
# + colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY>zZWxmKTsK", "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": ""}}, "base_uri": "https://localhost:8080/", "height": 73} id="0zHz6KExzdK4" outputId="656a3f7d-7077-4ed0-fb2c-f7aa0f30a087"
from google.colab import files
file = files.upload()
# + colab={"base_uri": "https://localhost:8080/"} id="kezNd_Al8ngT" outputId="dc929ed6-7a8f-4e73-a1ec-f76452e906bc"
pip install Bio
# + id="nAyUWhSKycbI"
from Bio import SeqIO
# + colab={"base_uri": "https://localhost:8080/"} id="19rKCx6n7ker" outputId="50a75f3f-616c-4716-be4d-876c78038a60"
# read CoV2 variant caller ID and sequences
variant_seqs = []
variant_callerID = []
variant_labels = []
for record in SeqIO.parse('CoV2variant_proteins_labels3.fasta', 'fasta'):
variant_callerID.append(' '.join(record.id.split('-')[:2]))
variant_labels.append(record.id.split('-')[-1])
variant_seqs.append(str(record.seq[:-1]))
#print(record.seq)
print(variant_callerID[0])
print(variant_labels[0])
print(variant_seqs[0])
# + colab={"base_uri": "https://localhost:8080/"} id="b1r2Rvd5UH6n" outputId="b84887d1-38a7-4eaf-c486-d9264a043f93"
len(variant_seqs)
# + colab={"base_uri": "https://localhost:8080/", "height": 175} id="Q_9kxvwZXwQL" outputId="ab963530-a045-470e-cf7b-7122c6387008"
# create dataframe for variant sequences
variant_df = pd.DataFrame({'Caller ID': variant_callerID, 'Sequences': variant_seqs, 'Levels': variant_labels})
variant_df.head(4)
# + colab={"base_uri": "https://localhost:8080/", "height": 308} id="VZf6DMDzrrMi" outputId="3c46fdab-5001-4525-e824-dee12fc0215d"
"""trim sequences to the same length"""
min_seq_length = min(variant_df.Sequences.apply(lambda x: len(x)))
# min_seq_length
# trim variant sequences to the minimum length
def trimming(seq):
# adding X to the end of each seq
trimmed_seq = seq[:min_seq_length] + 'X'
return trimmed_seq
variant_df['Trim Sequences'] = variant_df.Sequences.apply(trimming)
variant_df.head(4)
# + colab={"base_uri": "https://localhost:8080/"} id="mgzWjvLD5fhh" outputId="ac76c8a8-7125-468f-c47d-bda2333935c3"
variant_df.Levels.unique()
# + colab={"base_uri": "https://localhost:8080/"} id="EVW1OIVn1jZv" outputId="53933e73-175b-45f6-ba18-a48762a65d49"
variant_df['Trim Sequences'].apply(lambda x: len(x)).unique()
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="HgdWktwT6gbj" outputId="128adf46-29a4-4e9b-bf6c-fb386b0967ef"
# view the end of the sequence
variant_df['Trim Sequences'][0][1170:]
# + colab={"base_uri": "https://localhost:8080/"} id="HxKNaPPBva-z" outputId="2e5d5b3b-947c-4e2f-9216-759581dc15fd"
# extract trim sequences
training_seqs = variant_df['Trim Sequences']
training_seqs.tail(4)
# + [markdown] id="xgUHp73PlA9Q"
# ##**Data Preprocessing**
# + colab={"base_uri": "https://localhost:8080/"} id="LZkcGK_QwPGK" outputId="bc1c98fd-95b3-4c18-f5d6-48cbff85b632"
"""generate one-hot encoder for input features"""
# encodes a sequence of bases as a sequence of integers
int_encoder = LabelEncoder()
# converts an array of integers to a sparse matrix in which each sub-list presents an amino acid letter
one_hot_encoder = OneHotEncoder(categories='auto')
input_features = []
for seq in training_seqs:
int_encoded = int_encoder.fit_transform(list(seq))
int_encoded = np.array(int_encoded).reshape(-1, 1)
one_hot_encoded = one_hot_encoder.fit_transform(int_encoded)
input_features.append(one_hot_encoded.toarray())
# stack sub-lists into an array
input_features = np.stack(input_features)
print('Sequence 0: ', training_seqs[0][:15])
print('One-hot encoder for the first 15 amino acids in sequence 0: ')
print(input_features)
# + colab={"base_uri": "https://localhost:8080/"} id="V67wOClIJEGQ" outputId="5cd35302-5a2a-40c2-da89-53f095b2f7ed"
input_features.shape
# + colab={"base_uri": "https://localhost:8080/"} id="ID8tKakPzBSp" outputId="5a469122-e13d-47b1-895e-191a84ee737e"
"""generate one-hot encoder for input labels"""
one_hot_encoder = OneHotEncoder(categories='auto')
labels = np.array(variant_df['Levels']).reshape(-1,1)
# labels
input_labels = one_hot_encoder.fit_transform(labels).toarray()
input_labels[:5]
# + colab={"base_uri": "https://localhost:8080/"} id="ckaaf88Az7Gp" outputId="fbf7ff6d-a404-4d8c-e4d1-2d386d19a235"
input_labels.shape
# + [markdown] id="lMgcfopTlG2q"
# ##**Building Convolutional Neural Network Model**
# + id="Fa41mji0dkK6"
from sklearn.model_selection import train_test_split
# + id="iNQxxDr3ZnBi"
# create training and testing sets
train_seqs, test_seqs, train_labels, test_labels = train_test_split(input_features, input_labels,
test_size=0.20, random_state=42)
# + id="j_vq-duUZAH8" colab={"base_uri": "https://localhost:8080/"} outputId="359a4651-e94f-486e-abd9-d3bb7c8c67c8"
print(train_seqs.shape)
train_seqs[:2]
# + colab={"base_uri": "https://localhost:8080/"} id="OZGoTdTeuiBM" outputId="9464be05-c917-4269-b6e4-1a1ffcbdd58b"
print(train_labels.shape)
train_labels[:2]
# + colab={"base_uri": "https://localhost:8080/"} id="dLZsMRVSkBQ5" outputId="84dd7ed8-2b2d-4c81-ea14-4d4e13b8a738"
test_seqs.shape
# + colab={"base_uri": "https://localhost:8080/"} id="mLdmDKEqkD7K" outputId="661ccd94-905e-4930-a66f-d2f21d6d20f9"
test_labels.shape
# + id="xmIwLmhovUXJ"
from tensorflow.keras.layers import Conv1D, MaxPooling1D, Flatten, Dropout, Dense
from tensorflow.keras.models import Sequential
# + id="6QOM4DpKupN3" colab={"base_uri": "https://localhost:8080/"} outputId="30d1fbdb-59d5-4ea8-8990-88f58df7d93f"
"""build CNN model to classify variant sequences"""
# generate a keras sequential model
model = Sequential()
# add different layers to the model
model.add(Conv1D(filters=128, kernel_size=40, input_shape=(train_seqs.shape[1], train_seqs.shape[2])))
model.add(Flatten())
model.add(Dense(210, activation='relu'))
model.add(Dropout(0.25))
model.add(Dense(100, activation='relu'))
model.add(Dense(3, activation='softmax'))
model.summary()
# + id="hc6ezDYSxM7A"
# compile the keras model
model.compile(loss='binary_crossentropy', optimizer='adam', metrics=['binary_accuracy'])
# train the model with a fixed number of epochs (number of iterations on a dataset)
cnn_model = model.fit(train_seqs, train_labels, epochs=28, verbose=2, validation_split=0.25)
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="K7vhwB3PcunC" outputId="61b0d7b8-e782-40b5-eed9-d000e911cf04"
# visualizing the training and validation loss
plt.figure()
plt.plot(cnn_model.history['loss'])
plt.plot(cnn_model.history['val_loss'])
plt.title('Plot of model loss for training and validating datasets', y=1.07)
plt.xlabel('epoch')
plt.ylabel('loss')
plt.legend(['train', 'validation'])
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 311} id="FNAZcIIfdqY8" outputId="1e41307c-c644-4fac-ea59-accd6b88c425"
# visualizing the training and validation accuracy
plt.figure()
plt.plot(cnn_model.history['binary_accuracy'])
plt.plot(cnn_model.history['val_binary_accuracy'])
plt.title('Plot of model accuracy for training and validating datasets', y=1.07)
plt.xlabel('epoch')
plt.ylabel('accuracy')
plt.legend(['train', 'validation'])
plt.show()
# + [markdown] id="9iAAYTjtk3KC"
# ##**Model Performance Evaluation**
# + id="pv-Ly_PKma0P"
from sklearn import metrics
import itertools
# + colab={"base_uri": "https://localhost:8080/"} id="xPUs2s44kpQL" outputId="2c14f7cb-5f35-4d30-ee7f-61a3c8fa6bb3"
# compute the accuracy of the model
predicted_labels = model.predict(np.stack(test_seqs))
percent_accuracy = metrics.accuracy_score(np.argmax(test_labels, axis=1), np.argmax(predicted_labels, axis=1))
print('percent accuracy of the CNN model: ', percent_accuracy)
# + colab={"base_uri": "https://localhost:8080/", "height": 516} id="VhmEWEJu8iI7" outputId="a82b3853-dd97-4a76-e2ce-ad1b3117c703"
# confusion matrix
cnn_cm = metrics.confusion_matrix(np.argmax(test_labels, axis=1),
np.argmax(predicted_labels, axis=1))
print('Confusion matrix:\n',cnn_cm)
# normalized confusion matrix
cm_normalized = cnn_cm.astype('float') / cnn_cm.sum(axis = 1)[:, np.newaxis]
print('Normalized Confusion matrix:\n',cm_normalized)
#cm_normalized[0]
df = pd.DataFrame({'Interest': cm_normalized[:, 0], 'Concern': cm_normalized[:, 1], 'Other': cm_normalized[:, 2]})
#df['Label'] = df.
df.insert(0,'Label',['Interest','Concern','Other'],True)
print(df)
# plot normalized confusion matrix
sns.heatmap(df.set_index('Label'), annot=True, cmap='GnBu', linewidths=0.1)
plt.title('Normalized confusion matrix')
plt.xlabel('Predicted label')
plt.ylabel('True label')
| COVID19_CNN_seqclassifier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
import os.path
import numpy as np
import matplotlib.pyplot as plt
from sdhk import SDHKHarvester
# +
sdhk_path = os.path.expanduser("~/Data/SDHK")
assert os.path.exists(sdhk_path)
lowdef_path = os.path.join(sdhk_path, "LowDef")
highdef_path = os.path.join(sdhk_path, "HighDef_png")
savefile = os.path.join(sdhk_path, "metadata.json.gz")
harvester = SDHKHarvester(savefile)
# -
# #### Popluate
dl_keys = list(set(range(1, 50000)).difference(set(harvester.keys())))
harvester.download(dl_keys)
print("%i good ids in database" % (len(harvester.get_good_ids())))
print("%i text entries in database" %
(np.sum([harvester[n]['textcontent'] is not None
and len(harvester[n]['textcontent']) > 50
for n in harvester.get_good_ids()])))
harvester.save()
# #### Clean up some paths
# +
print("%i bad paths removed form database" % harvester.remove_bad_paths())
if os.path.exists(lowdef_path):
harvester.scan_lowdef_path(lowdef_path)
print("%i low def images in database" %
(np.sum(['lowdefpath' in harvester[n]
for n in harvester.get_good_ids()])))
if os.path.exists(highdef_path):
harvester.scan_highdef_path(highdef_path)
print("%i high def images in database" %
(np.sum(['highdefpath' in harvester[n]
for n in harvester.get_good_ids()])))
harvester.save()
# https://lbiiif.riksarkivet.se/sdhk!1094/manifest
# -
# #### Write a CSV version of the harvested data
# +
csvfile = savefile.replace(".json.gz", ".csv.gz")
keys = list()
for n in harvester.keys():
for k in harvester[n].keys():
if k not in keys:
keys.append(k)
import gzip
with gzip.open(csvfile, 'wt') as f:
for i, k in enumerate(keys):
if i > 0:
f.write(", ")
f.write(k)
f.write('\n')
for n in harvester.keys():
for i, k in enumerate(keys):
if k in harvester[n].keys():
if i > 0:
f.write(", ")
data = harvester[n][k]
if type(data) == str:
f.write('\"' + data + '\"')
else:
f.write(repr(data))
f.write('\n')
# -
# #### Plot histogram over text length and entries per decade
# +
fig = plt.figure(figsize=(12, 4), dpi=100)
ax = fig.subplots(1, 2)
#plt.rc('text', usetex=True)
#plt.rc('font', family='serif')
text_ids = [n for n in harvester.get_good_ids()
if harvester[n]['textcontent'] is not None]
text_lengths = [len(harvester[n]['textcontent']) for n in text_ids]
ax[0].set_title("Length of transcribed texts")
ax[0].hist(text_lengths, 100)
ax[0].set_xlabel('Length of character')
ax[0].set_ylabel('Number of documents')
ax[0].set_xlim(np.min(text_lengths), np.max(text_lengths))
dated_ids = [n for n in harvester.get_good_ids()
if 0 < harvester[n]['year'] and
harvester[n]['year'] <= 1661]
years = [harvester[n]['year'] for n in dated_ids]
ax[1].set_title("SHDK charters per decade")
ax[1].hist(years, list(range(np.min(years)//10*10, np.max(years)//10*10, 10)))
ax[1].set_xlabel('Year')
ax[1].set_ylabel('Number of charters')
ax[1].set_xlim(np.min(years), np.max(years))
#ax[1].set_xlim(1135, 1546)
fig.show()
# -
# #### Plot months
# +
dates_as_text = [harvester[n]['date_as_text'].lower() for n in harvester.keys() if 'date_as_text' in harvester[n]]
months = ['januari', 'februari', 'mars', 'april', 'maj', 'juni', 'juli',
'augusti', 'september', 'oktober', 'november', 'december']
def get_date_tuple(text):
try:
t = text.split(" ")
assert len(t) == 3
year = int(t[0])
day = int(t[2])
for i, m in enumerate(months):
if t[1].find(m) >= 0:
return (year, i+1, day)
return None
except:
return None
assert get_date_tuple('1373 oktober 25')[0] == 1373
assert get_date_tuple('1373 oktober 25')[1] == 10
assert get_date_tuple('1373 oktober 25')[2] == 25
assert get_date_tuple('1409 november 5')[0] == 1409
assert get_date_tuple('1409 november 5')[1] == 11
assert get_date_tuple('1409 november 5')[2] == 5
assert get_date_tuple('1373utan dag') is None
month = [date[1] for date in list(map(get_date_tuple, dates_as_text)) if date is not None]
month_histogram = np.zeros(len(months))
for i, m in enumerate(months):
month_histogram[i] = np.sum(np.asarray(month)==(i+1))
fig = plt.figure(figsize=(12, 4), dpi=100)
ax = fig.subplots(1, 1)
#plt.rc('text', usetex=True)
#plt.rc('font', family='serif')
bars = ax.bar(list(range(12)), month_histogram)
ax.set_title('SHDK charters per month')
ax.set_ylabel('Number of charters')
ax.set_xticks(list(range(12)))
ax.set_xticklabels(months)
plt.show()
# +
#pip install convertdate
from convertdate import julian
from datetime import date
def to_gregorian_weekday(date_tuple):
try:
j = julian.to_gregorian(date_tuple[0], date_tuple[1], date_tuple[2])
ret = date(j[0], j[1], j[2]).weekday()
return ret
except:
return None
date_tuples = [date_tuple for date_tuple in list(map(get_date_tuple, dates_as_text)) if date_tuple is not None]
weekdays = [weekday for weekday in list(map(to_gregorian_weekday, date_tuples)) if weekday is not None]
weekday_names = ['Monday', 'Tuesday', 'Wednesday', 'Thursday', 'Friday', 'Saturday', 'Sunday']
weekday_histogram = np.zeros(len(weekday_names))
for weekday in weekdays:
weekday_histogram[weekday] += 1
fig = plt.figure(figsize=(12, 4), dpi=100)
ax = fig.subplots(1, 1)
#plt.rc('text', usetex=True)
#plt.rc('font', family='serif')
ax.bar(list(range(7)), weekday_histogram)
ax.set_title('SHDK charters per weekday')
ax.set_ylabel('Number of charters')
ax.set_xticks(list(range(7)))
ax.set_xticklabels(weekday_names)
plt.show()
| SDHK.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
import numpy as np
import progressbar
# results in this format from the output of the multi-threaded c++ code.
resultsFinal = np.asarray([
0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0,
3e-08, 0, 0, 0, 0, 0,
6e-08, 0, 0, 0, 0, 0,
2.5e-07, 0, 0, 0, 0, 0,
7.1e-07, 0, 0, 0, 0, 0,
1.65e-06, 0, 0, 0, 0, 0,
4.9e-06, 0, 0, 0, 0, 0,
1.178e-05, 0, 0, 0, 0, 0,
2.797e-05, 0, 0, 0, 0, 0,
6.254e-05, 0, 0, 0, 0, 0,
0.00013422, 2e-08, 0, 0, 0, 0,
0.00027849, 7e-08, 0, 0, 0, 0,
0.00054056, 1.5e-07, 0, 0, 0, 0,
0.00101865, 8.4e-07, 0, 0, 0, 0,
0.00185053, 3.71e-06, 0, 0, 0, 0,
0.00323235, 1.331e-05, 0, 0, 0, 0,
0.0054424, 3.902e-05, 0, 0, 0, 0,
0.00889944, 0.00011233, 1e-08, 0, 0, 0,
0.0140104, 0.00029576, 8e-08, 0, 0, 0,
0.0214406, 0.00073342, 1.19e-06, 0, 0, 0,
0.0318141, 0.00168044, 6.31e-06, 2e-08, 0, 0,
0.0459994, 0.00364552, 3.162e-05, 2e-08, 0, 0,
0.0646293, 0.00743669, 0.0001314, 4.6e-07, 0, 0,
0.0885937, 0.0142396, 0.00049523, 4.14e-06, 2e-08, 0,
0.118352, 0.0257663, 0.00162757, 3.233e-05, 3.6e-07, 0,
0.154433, 0.0440788, 0.00472106, 0.00020428, 3.66e-06, 1e-08,
0.196837, 0.0715445, 0.0122646, 0.00105628, 4.37e-05, 9.2e-07,
0.245502, 0.110379, 0.0283845, 0.00443653, 0.00041492, 2.269e-05,
0.299761, 0.162142, 0.0591246, 0.0153717, 0.00283166, 0.00037416,
0.358843, 0.227416, 0.111237, 0.0440746, 0.0141348, 0.00365242,
0.421386, 0.305263, 0.189755, 0.105458, 0.0524408, 0.0232268,
0.485932, 0.392829, 0.295232, 0.212378, 0.146406, 0.0962258,
0.550785, 0.486158, 0.421551, 0.364836, 0.314247, 0.268755,])
resultsFinal = np.reshape(resultsFinal, (40,6))
# +
# honest network delay over next n blocks.
def vectorDelayHonest(ps, es, init_endorsers, delay_priority, delay_endorse):
return (60 * len(ps)
+ delay_priority * sum(ps)
+ sum([delay_endorse * max(init_endorsers - e, 0) for e in es]))
# attacking network delay over next n blocks.
def vectorDelayAttacker(ps, es, init_endorsers, delay_priority, delay_endorse):
return (60 * len(ps)
+ delay_priority * sum(ps)
+ sum([delay_endorse * max(init_endorsers - e, 0) for e in es[1:]]))
# efficient sample generation
def getAH(alpha):
x = np.random.geometric(1-alpha)
if x == 1:
h = 0
a = np.random.geometric(alpha)
else:
a = 0
h = x - 1
return [a, h]
def rewardBlock(p, e):
if p == 0:
return e * 1.25
return e * 0.1875
def rewardEndorsement(p):
if p == 0:
return 1.25
return 0.8333333
def calcHonestSingle(p, e):
if p == 0:
return rewardBlock(0, 32) + e * rewardEndorsement(0)
return e * rewardEndorsement(0)
def calcAttackSingle(p, e):
return rewardBlock(p, e) + e * rewardEndorsement(p)
def vectorRewardHonest(ps, es):
totalReward = 0
for i in range(len(ps)):
totalReward += calcHonestSingle(ps[i], es[i])
return totalReward
def vectorRewardAttack(ps, es):
totalReward = calcAttackSingle(ps[0], 32)
for i in range(1,len(ps)):
totalReward += calcAttackSingle(ps[i], es[i])
return totalReward
def calcCosts(ps, es):
return vectorRewardHonest(ps, es) - vectorRewardAttack(ps, es)
# +
def getProbReorg(alpha, length, init_endorsers, delay_priority, delay_endorse, sample_size = int(1e3)):
feasible_count = 0
for _ in range(sample_size):
aVals = []
hVals = []
for i in range(length):
a, h = getAH(alpha)
aVals.append(a)
hVals.append(h)
eVals = np.random.binomial(32, alpha, size = length)
honest_delay = vectorDelayHonest(hVals, 32 - eVals, init_endorsers, delay_priority, delay_endorse)
selfish_delay = vectorDelayAttacker(aVals, eVals, init_endorsers, delay_priority, delay_endorse)
if selfish_delay <= honest_delay:
feasible_count += 1
return feasible_count / sample_size
def getProbSelfish(alpha, length, init_endorsers, delay_priority, delay_endorse, sample_size = int(1e3)):
feasible_count = 0
for _ in range(sample_size):
aVals = []
hVals = []
for i in range(length):
a, h = getAH(alpha)
aVals.append(a)
hVals.append(h)
eVals = np.random.binomial(32, alpha, size = length)
honest_delay = vectorDelayHonest(hVals, 32 - eVals, init_endorsers, delay_priority, delay_endorse)
selfish_delay = vectorDelayAttacker(aVals, eVals, init_endorsers, delay_priority, delay_endorse)
if (selfish_delay <= honest_delay) and (calcCosts(aVals, eVals) < 0):
feasible_count += 1
return feasible_count / sample_size
# -
length_20_probs = resultsFinal[:,2]
length_20_probs_nonzero = length_20_probs[24:]
length_20_probs_nonzero
alphas_total= np.arange(0.34, 0.50, 0.01)
for i in range(16):
print(alphas_total[i], length_20_probs_nonzero[i])
alphas = [0.41, 0.45, 0.49]
tezos_probs = [length_20_probs_nonzero[7], length_20_probs_nonzero[11], length_20_probs_nonzero[15]]
for i in range(3):
print(alphas[i], tezos_probs[i])
weights = 1 / np.asarray(tezos_probs)
weights
init_endorsers_range = list(range(33))
delay_priority_range = list(range(20, 61))
delay_endorse_range = list(range(4, 13))
results_grid = np.zeros(shape=(
len(init_endorsers_range),
len(delay_priority_range),
len(delay_endorse_range)))
alphas, weights
prob1 = getProbReorg(
alpha=0.45,
length=20,
init_endorsers=24,
delay_priority=40,
delay_endorse=8,
sample_size=int(1e5))
prob2 = getProbSelfish(
alpha=0.45,
length=3,
init_endorsers=24,
delay_priority=40,
delay_endorse=8,
sample_size=int(1e5))
print(prob1, prob2)
bar = progressbar.ProgressBar()
for i in bar(range(len(init_endorsers_range))):
for j in range(len(delay_priority_range)):
for k in range(len(delay_endorse_range)):
prob1 = getProbReorg(
alpha=0.45,
length=20,
init_endorsers=init_endorsers_range[i],
delay_priority=delay_priority_range[j],
delay_endorse=delay_endorse_range[k],
sample_size=int(1e3))
prob2 = getProbSelfish(
alpha=0.45,
length=3,
init_endorsers=init_endorsers_range[i],
delay_priority=delay_priority_range[j],
delay_endorse=delay_endorse_range[k],
sample_size=int(1e3))
# prob2 = getProbReorg(
# alphas[1],
# length=20,
# init_endorsers=init_endorsers_range[i],
# delay_priority=delay_priority_range[j],
# delay_endorse=delay_endorse_range[k],
# sample_size=int(1e3))
# prob3 = getProbReorg(
# alphas[2],
# length=20,
# init_endorsers=init_endorsers_range[i],
# delay_priority=delay_priority_range[j],
# delay_endorse=delay_endorse_range[k],
# sample_size=int(1e3))
# val = weights[0] * prob1 + weights[1] * prob2 + weights[2] * prob3
results_grid[i, j, k] = prob1 + prob2
list(results_grid)
np.savetxt('grid45_cost.txt', results_grid.flatten())
| grid.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Atul's Brooklyn housing price predictions
#
# _September 20, 2017_
#
# This is just some experimenting I did after taking the second week of Coursera's [Machine Learning](https://www.coursera.org/learn/machine-learning) class, which covers linear regression.
#
# The example <NAME> uses to motivate linear regression is predicting housing prices in Portland, so I figured I would do the same thing, but with Brooklyn.
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
# -
# I found the [NYC Department of Finance’s Rolling Sales data](http://www1.nyc.gov/site/finance/taxes/property-rolling-sales-data.page), which was in excel format, and converted it to CSV. Here's what it looks like:
# +
# Column name for square footage.
SQFT = 'GROSS SQUARE FEET'
# Column name for price.
PRICE = 'SALE PRICE'
# Maximum square footage that we care about.
MAX_SQFT = 4500
sa = pd.read_csv('rollingsales_brooklyn.csv')
for colname in [SQFT, PRICE]:
sa[colname] = pd.to_numeric(sa[colname].str.replace(',', ''), errors='coerce')
sa = sa[pd.notnull(sa[colname])]
sa = sa[sa[SQFT] < MAX_SQFT]
sa[:3]
# -
# I wanted to be able to easily drill down by neighborhood, so here's a list of all the available neighborhoods...
sa['NEIGHBORHOOD'].drop_duplicates().values
# And here's a function that returns only the dataframe rows that correspond to a given neighborhood...
def hood(name):
return sa[sa['NEIGHBORHOOD'] == name]
# Now let's do some plotting!
hood('GOWANUS').plot(x=SQFT, y=PRICE, kind='scatter')
# Neato. Now let's create an _unvectorized_ cost function, given a dataframe and linear regression parameters.
# +
def unvectorized_cost(df, theta_0, theta_1):
X = df[SQFT].values
Y = df[PRICE].values
m = len(X)
total = 0
for i in range(m):
total += np.power((theta_0 + theta_1 * X[i]) - Y[i], 2)
return total / m / 2
unvectorized_cost(hood('GOWANUS'), 1, 2)
# -
# Now let's make a vectorized version of the same function.
# +
def cost(df, theta_0, theta_1):
X = df[SQFT].values
Y = df[PRICE].values
m = len(X)
predicted_Y = theta_0 + theta_1 * X
total = np.sum(np.power(predicted_Y - Y, 2))
return total / m / 2
# Make sure it produces the same output as our unvectorized version.
assert unvectorized_cost(hood('GOWANUS'), 1, 2) == cost(hood('GOWANUS'), 1, 2)
# -
# Now let's create an unvectorized gradient-following function. Obviously using an analytical solution would be more efficient here, given our features, but I want to make sure I understand the general concept of gradient descent.
def unvectorized_follow_gradient(df, theta_0, theta_1, learning_rate):
X = df[SQFT].values
Y = df[PRICE].values
m = len(X)
total_theta_0 = 0
total_theta_1 = 0
for i in range(m):
predicted_y = theta_0 + theta_1 * X[i]
total_theta_0 += (predicted_y - Y[i]) * 1
total_theta_1 += (predicted_y - Y[i]) * X[i]
d_theta_0 = total_theta_0 / m
d_theta_1 = total_theta_1 / m
return (theta_0 - learning_rate * d_theta_0,
theta_1 - learning_rate * d_theta_1)
# Cool, now let's use the function to fit a line to our data!
# +
theta_0 = 0
theta_1 = 0
df = hood('GOWANUS')
for i in range(1000):
if i % 100 == 0:
print(f"cost at iteration {i} when theta_0={theta_0}, theta_1={theta_1} is {cost(df, theta_0, theta_1)}")
theta_0, theta_1 = unvectorized_follow_gradient(df, theta_0, theta_1, learning_rate=0.000000001)
# +
square_footage = np.arange(0, MAX_SQFT)
predicted_prices = theta_0 + theta_1 * square_footage
plt.plot(df[SQFT], df[PRICE], '.')
plt.plot(square_footage, predicted_prices, '-')
# -
| bk-housing/BkHousing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Centralised Logistic Regression Classifier
# Author: **<NAME>**
import math
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn import preprocessing
def read_file(filename):
read_data = pd.read_csv(filename, header=None)
return read_data
df = read_file("../data/spam.csv")
df.head()
# +
def get_x(df):
return df.iloc[:, 0:-1].values
def get_y(df):
return df.iloc[:, -1].values
def standardize(df):
x = get_x(df)
df_y = df.iloc[:, 57]
standard_scaler = preprocessing.StandardScaler()
x_scaled = standard_scaler.fit_transform(x)
df_x = pd.DataFrame(x_scaled)
df_scaled = df_x.join(df_y)
return df_scaled
# -
df = standardize(df)
df.head()
# +
def sigmoid(z):
return 1 / (1 + np.exp(-z))
def loss(p, y, w, lambda_reg):
epsilon = 1e-10
error_loss = -np.average(
y * np.log(p + epsilon) + (1 - y) * np.log(1 - p + epsilon)
)
reg_loss = lambda_reg * np.sum(np.square(w)) / (2 * y.size)
return error_loss + reg_loss
def predict_probability(x, b, w):
z = np.dot(x, w) + b
return sigmoid(z)
# -
def train(df, iterations=1000, learning_rate=10, lambda_reg=0.01, verbose=True):
x = get_x(df)
y = get_y(df)
w_train = np.zeros(x.shape[1])
b = 0
loss_array = []
for it in range(iterations):
predictions = predict_probability(x, b, w_train)
loss_a = loss(predictions, y, w_train, lambda_reg)
gradient_w = np.dot(x.T, (predictions - y))
gradient_b = np.average(predictions - y)
regularization = lambda_reg * w_train
w_train -= learning_rate * (gradient_w + regularization) / y.size
b -= learning_rate * gradient_b
if verbose and (it % (iterations/5) == 0 or it+1 == iterations):
print("It. %4d\t|\tLoss: %0.4f" %
(it, loss_a)
)
if it % 10 == 0:
loss_array.append(
loss_a
)
return b, w_train
# +
def predict(x, b, w, threshold):
prob = predict_probability(x, b, w)
return prob >= threshold
def accuracy(df, b, w, threshold=0.5):
x = get_x(df)
y = get_y(df)
predictions = predict(x, b, w, threshold=threshold)
acc = np.average(predictions == y)
return acc
def get_block_data(df, fold, tot_folds):
fold_size = math.floor(df.shape[0] / tot_folds)
start_index = fold_size * fold
end_index = start_index + fold_size
df_test = df.loc[start_index:end_index]
df.drop(df.loc[start_index:end_index].index, inplace=True)
return df, df_test
def shuffle(df):
return df.sample(frac=1).reset_index(drop=True)
# -
def cross_validation(df, iterations=1000, learning_rate=10, threshold=0.5,
lambda_reg=0.1, folds=10, verbose_train=False):
avg_acc = 0
df = shuffle(df)
for i in range(folds):
print("Fold number " + str(i+1))
tr_data, test_data = get_block_data(df.copy(), i, folds)
b, w = train(
tr_data,
learning_rate=learning_rate,
lambda_reg=lambda_reg,
verbose=verbose_train
)
avg_acc += accuracy(test_data, b, w, threshold=threshold)
avg_acc /= folds
print("\nAVG acc: %0.4f" % avg_acc)
return avg_acc
cross_validation(df,
folds=10,
verbose_train=False,
iterations=1000,
learning_rate=10,
lambda_reg=0.1,
threshold=0.5
)
b, w = train(df,
iterations=1000,
learning_rate=10,
lambda_reg=0.1
)
print(b)
accuracy(df, b, w)
# ### Skopt analysis
# Not required.
# +
import skopt
SPACE = [skopt.space.Real(
0.001,
10,
name = 'lambda_reg',
prior='log_uniform'
),
skopt.space.Real(
0.01,
10,
name='learning_rate',
prior='log_uniform'
)
]
@skopt.utils.use_named_args(SPACE)
def objective(**params):
all_params = {**params}
return -1.0 * cross_validation(df, all_params)
random_results = skopt.dummy_minimize(objective, SPACE)
forest_results = skopt.forest_minimize(objective, SPACE)
gbrt_results = skopt.gbrt_minimize(objective, SPACE)
gp_results = skopt.gp_minimize(objective, SPACE)
results = [('random_results', random_results),
('forest_results', forest_results),
('gbrt_results', gbrt_results),
('gp_results', gp_results)]
# -
# End.
| notebooks/Centralised_LR.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from keras import layers
def res_block(y, nb_channels, _strides = (1,1), _project_shortcut=False):
shortcut = y
y = layers.Conv2D(nb_channels, kernel_size=(3, 3), strides=_strides, padding='same')(y)
y = layers.BatchNormalization()(y)
y = layers.LeakyReLU()(y)
y = layers.Conv2D(nb_channels, kernel_size=(3, 3), strides=(1, 1), padding='same')(y)
y = layers.BatchNormalization()()
if _project_shortcut or _strides != (1, 1):
shortcut = layers.Conv2D(nb_channels, kernel_size=(1, 1), strides=_strides, padding='same')(shortcut)
shortcut = layers.BatchNormalization()(shortcut)
y = layers.add([shortcut, y])
y = layers.LeakyReLU()(y)
return y
| .ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas
import re
import os
import matplotlib.pyplot as plt
import numpy as np
from keras.models import Sequential, load_model
from keras.layers import InputLayer, Masking, LSTM, TimeDistributed, Dense
from math import ceil
from keras.callbacks import Callback, ModelCheckpoint
csvs = list(filter(lambda file: re.match(r'^(?!61706006)\d{8} Test Data\.txt$', file) is not None, os.listdir('./DynamometerData')))
dfs = [pandas.read_csv('./DynamometerData/' + csv, sep='\t', header=0) for csv in csvs]
max_length = (ceil(max([len(df) for df in dfs]) / 100)) * 100
max_length
# +
# Padding with invalid value -1.
X = np.full([len(dfs), max_length, 10], -1.)
Y = np.full([len(dfs), max_length, 8], -1.)
for i, df in enumerate(dfs):
# Current
X[i,:len(df)-1,0] += df['Brake_pressure_applied_PCM[]'].values[1:] + 1
X[i,:len(df)-1,1] += df['Pedal_accel_pos_CAN[per]'].values[1:] + 1
# Previous
X[i,:len(df)-1,2] += df['Dyno_Spd[mph]'].values[:-1] + 1
X[i,:len(df)-1,3] += df['Eng_throttle_electronic_control_actual_PCM[deg]'].values[:-1] + 1
X[i,:len(df)-1,4] += df['Eng_throttle_position_PCM[per]'].values[:-1] + 1
X[i,:len(df)-1,5] += df['Trans_gear_engaged_CAN[]'].values[:-1] + 1
X[i,:len(df)-1,6] += df['Eng_load_PCM[per]'].values[:-1] + 1
X[i,:len(df)-1,7] += df['Eng_speed_PCM[rpm]'].values[:-1] + 1
X[i,:len(df)-1,8] += df['Trans_gear_ratio_measured_TCM[]'].values[:-1] + 1
X[i,:len(df)-1,9] += df['Trans_output_shaft_speed_raw_TCM[rpm]'].values[:-1] + 1
# Outputs
Y[i,:len(df)-1,0] += df['Dyno_Spd[mph]'].values[1:] + 1
Y[i,:len(df)-1,1] += df['Eng_throttle_electronic_control_actual_PCM[deg]'].values[1:] + 1
Y[i,:len(df)-1,2] += df['Eng_throttle_position_PCM[per]'].values[1:] + 1
Y[i,:len(df)-1,3] += df['Trans_gear_engaged_CAN[]'].values[1:] + 1
Y[i,:len(df)-1,4] += df['Eng_load_PCM[per]'].values[1:] + 1
Y[i,:len(df)-1,5] += df['Eng_speed_PCM[rpm]'].values[1:] + 1
Y[i,:len(df)-1,6] += df['Trans_gear_ratio_measured_TCM[]'].values[1:] + 1
Y[i,:len(df)-1,7] += df['Trans_output_shaft_speed_raw_TCM[rpm]'].values[1:] + 1
# -
del dfs
# +
NEW_MIN = 0.25
NEW_MAX = 0.75
OLD_PAD_VAL = -1.
NEW_PAD_VAL = 0.
X_mins, X_maxs = [], []
for k in range(X.shape[2]):
X_mins.append(X[:,:,k][X[:,:,k] != OLD_PAD_VAL].min())
X_maxs.append(X[:,:,k][X[:,:,k] != OLD_PAD_VAL].max())
X_std = np.full(X.shape, NEW_PAD_VAL)
for i in range(X.shape[0]):
for k in range(X.shape[2]):
indices = np.where(X[i,:,k] != OLD_PAD_VAL)
X_std[i,indices,k] += ((X[i,indices,k] - X_mins[k]) / (X_maxs[k] - X_mins[k])) * (NEW_MAX - NEW_MIN) + NEW_MIN - NEW_PAD_VAL
Y_mins, Y_maxs = [], []
for k in range(Y.shape[2]):
Y_mins.append(Y[:,:,k][Y[:,:,k] != OLD_PAD_VAL].min())
Y_maxs.append(Y[:,:,k][Y[:,:,k] != OLD_PAD_VAL].max())
Y_std = np.full(Y.shape, NEW_PAD_VAL)
for i in range(Y.shape[0]):
for k in range(Y.shape[2]):
indices = np.where(Y[i,:,k] != OLD_PAD_VAL)
Y_std[i,indices,k] += ((Y[i,indices,k] - Y_mins[k]) / (Y_maxs[k] - Y_mins[k])) * (NEW_MAX - NEW_MIN) + NEW_MIN - NEW_PAD_VAL
# +
SPLIT = X_std.shape[0] - X_std.shape[0] // 10
indices = np.arange(0, X_std.shape[0])
np.random.shuffle(indices)
X_shuffled = X_std[indices,:,:]
Y_shuffled = Y_std[indices,:,:]
X_train, X_test = X_shuffled[:SPLIT,:,:], X_shuffled[SPLIT:,:,:]
Y_train, Y_test = Y_shuffled[:SPLIT,:,:], Y_shuffled[SPLIT:,:,:]
# +
test_model = Sequential()
test_model.add(InputLayer(batch_input_shape=(1, 1, X_std.shape[2])))
test_model.add(LSTM(200, stateful=True))
test_model.add(Dense(Y_std.shape[2], activation='linear'))
test_model.summary()
# -
test_model.load_weights('Models/MaximalStateLSTM/E200L9.6744E-06.hdf5')
test_model.reset_states()
# +
i = 2
Y_pred = np.zeros([1, Y_test.shape[1], Y_test.shape[2]])
inputs = X_train[i:i+1,0:1,:].copy()
j = 0
while X_train[i,j,0] != 0.:
Y_pred[0,j:j+1,:] += test_model.predict(inputs, batch_size=1)
inputs[0,0,:2] = X_train[i:i+1,j+1:j+2,:2]
inputs[0,0,2:] = Y_pred[0,j:j+1,:]
j += 1
test_model.reset_states()
# +
fig, ax = plt.subplots()
ax.scatter(range(max_length), Y_train[i,:,0], s=2)
ax.scatter(range(max_length), Y_pred[0,:,0], s=2)
# Zoom in
plt.xlim(0, 10000)
plt.ylim(0, 1)
plt.tight_layout()
plt.show()
# -
| archive/MaximalStateLSTM-Test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
DATA_DIRECTORY = '\data\eng.train'
DATA_TESTA_DIRECTORY = '\data\eng.testa'
DATA_TESTB_DIRECTORY = '\data\eng.testb'
TRAIN_FILENAME = '\data\\train.words.txt'
TRAIN_TAGS_FILENAME = '\data\\train.tags.txt'
TESTA_FILENAME = '\data\\testa.words.txt'
TESTA_TAGS_FILENAME = '\data\\testa.tags.txt'
TESTB_FILENAME = '\data\\testb.words.txt'
TESTB_TAGS_FILENAME = '\data\\testb.tags.txt'
SENTENCE_DELIMITER = '. . O O'
if os.path.isfile(os.getcwd() + DATA_TESTB_DIRECTORY):
with open(os.getcwd() + DATA_TESTB_DIRECTORY) as fp:
line = fp.readline()
cnt = 1
outfile = open(os.getcwd() + TESTB_FILENAME, 'w')
sentence = ''
while line:
line = line.strip()
if line == SENTENCE_DELIMITER:
outfile.write(sentence)
outfile.write('\n')
sentence = ''
else:
sentence = sentence + ' ' + ''.join(line.split()[:-3])
line = fp.readline()
cnt += 1
outfile.close()
if os.path.isfile(os.getcwd() + DATA_TESTB_DIRECTORY):
with open(os.getcwd() + DATA_TESTB_DIRECTORY) as fp:
line = fp.readline()
cnt = 1
outfile = open(os.getcwd() + TESTB_TAGS_FILENAME, 'w')
sentence_tags = ''
while line:
line = line.strip()
if line == SENTENCE_DELIMITER:
outfile.write(sentence_tags)
outfile.write('\n')
sentence_tags = ''
else:
if len(line.split()) > 0:
if ''.join(line.split()[-1]) == 'O' and ''.join(line.split()[-2]) == 'I-NP':
sentence_tags = sentence_tags + ' ' + ''.join(line.split()[-2])
else:
sentence_tags = sentence_tags + ' ' + ''.join(line.split()[-1])
line = fp.readline()
cnt += 1
outfile.close()
| sequence_tagging_ner/build_training_test_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Tradução de Textos - Experimento
# ## Utilização do modelo [MarianMT](https://huggingface.co/transformers/model_doc/marian.html) para tradução.
#
#
# * Neste exempo a tradução é feito do inglês para o português, mas ela pode ser feita em qualquer uma das línguas suportadas pelo MarianMT.
# * Para adaptar para traduções em outras línguas é necessário verificacar se há o modelo pré treinado disponível no MarianMT e adaptar o truncamento de strings do [spacy](https://spacy.io/usage/models) para o idioma desejado
# * A métrica computada é o [sacrebleu](https://https://github.com/mjpost/sacrebleu)
#
#
# ### **Em caso de dúvidas, consulte os [tutoriais da PlatIAgro](https://platiagro.github.io/tutorials/).**
# ## Declaração de parâmetros e hiperparâmetros
#
# Declare parâmetros com o botão <img src="data:image/png;base64,iVBORw0KGgoAAAANSUhEUgAAABQAAAAUCAYAAACNiR0NAAABhWlDQ1BJQ0MgcHJvZmlsZQAAKJF9kT1Iw0AcxV9TtaIVBzuIOASpThb8QhylikWwUNoKrTqYXPohNGlIUlwcBdeCgx+LVQcXZ10dXAVB8APEydFJ0UVK/F9SaBHjwXE/3t173L0DhFqJqWbbGKBqlpGMRcVMdkUMvKID3QhiCOMSM/V4aiENz/F1Dx9f7yI8y/vcn6NHyZkM8InEs0w3LOJ14ulNS+e8TxxiRUkhPiceNeiCxI9cl11+41xwWOCZISOdnCMOEYuFFpZbmBUNlXiKOKyoGuULGZcVzluc1VKFNe7JXxjMacsprtMcRAyLiCMBETIq2EAJFiK0aqSYSNJ+1MM/4PgT5JLJtQFGjnmUoUJy/OB/8LtbMz854SYFo0D7i21/DAOBXaBete3vY9uunwD+Z+BKa/rLNWDmk/RqUwsfAb3bwMV1U5P3gMsdoP9JlwzJkfw0hXweeD+jb8oCfbdA16rbW2Mfpw9AmrpaugEODoGRAmWveby7s7W3f880+vsBocZyukMJsmwAAAAGYktHRAD/AP8A/6C9p5MAAAAJcEhZcwAADdcAAA3XAUIom3gAAAAHdElNRQfkBgsMIwnXL7c0AAACDUlEQVQ4y92UP4gTQRTGf29zJxhJZ2NxbMBKziYWlmJ/ile44Nlkd+dIYWFzItiNgoIEtFaTzF5Ac/inE/urtLWxsMqmUOwCEpt1Zmw2xxKi53XitPO9H9978+aDf/3IUQvSNG0450Yi0jXG7C/eB0cFeu9viciGiDyNoqh2KFBrHSilWstgnU7nFLBTgl+ur6/7PwK11kGe5z3n3Hul1MaiuCgKDZwALHA7z/Oe1jpYCtRaB+PxuA8kQM1aW68Kt7e3zwBp6a5b1ibj8bhfhQYVZwMRiQHrvW9nWfaqCrTWPgRWvPdvsiy7IyLXgEJE4slk8nw+T5nDgDbwE9gyxryuwpRSF5xz+0BhrT07HA4/AyRJchUYASvAbhiGaRVWLIMBYq3tAojIszkMoNRulbXtPM8HwV/sXSQi54HvQRDcO0wfhGGYArvAKjAq2wAgiqJj3vsHpbtur9f7Vi2utLx60LLW2hljEuBJOYu9OI6vAzQajRvAaeBLURSPlsBelA+VhWGYaq3dwaZvbm6+m06noYicE5ErrVbrK3AXqHvvd4bD4Ye5No7jSERGwKr3Pms2m0pr7Rb30DWbTQWYcnFvAieBT7PZbFB1V6vVfpQaU4UtDQetdTCZTC557/eA48BlY8zbRZ1SqrW2tvaxCvtt2iRJ0i9/xb4x5uJRwmNlaaaJ3AfqIvKY/+78Av++6uiSZhYMAAAAAElFTkSuQmCC" /> na barra de ferramentas.<br>
# A variável `dataset` possui o caminho para leitura do arquivos importados na tarefa de "Upload de dados".<br>
# Você também pode importar arquivos com o botão <img src="data:image/png;base64,iVBORw0KGgoAAAANSUhEUgAAABQAAAAUCAYAAACNiR0NAAABhWlDQ1BJQ0MgcHJvZmlsZQAAKJF9kT1Iw0AcxV9TtaIVBzuIOASpThb8QhylikWwUNoKrTqYXPohNGlIUlwcBdeCgx+LVQcXZ10dXAVB8APEydFJ0UVK/F9SaBHjwXE/3t173L0DhFqJqWbbGKBqlpGMRcVMdkUMvKID3QhiCOMSM/V4aiENz/F1Dx9f7yI8y/vcn6NHyZkM8InEs0w3LOJ14ulNS+e8TxxiRUkhPiceNeiCxI9cl11+41xwWOCZISOdnCMOEYuFFpZbmBUNlXiKOKyoGuULGZcVzluc1VKFNe7JXxjMacsprtMcRAyLiCMBETIq2EAJFiK0aqSYSNJ+1MM/4PgT5JLJtQFGjnmUoUJy/OB/8LtbMz854SYFo0D7i21/DAOBXaBete3vY9uunwD+Z+BKa/rLNWDmk/RqUwsfAb3bwMV1U5P3gMsdoP9JlwzJkfw0hXweeD+jb8oCfbdA16rbW2Mfpw9AmrpaugEODoGRAmWveby7s7W3f880+vsBocZyukMJsmwAAAAGYktHRAD/AP8A/6C9p5MAAAAJcEhZcwAADdcAAA3XAUIom3gAAAAHdElNRQfkBgsOBy6ASTeXAAAC/0lEQVQ4y5WUT2gcdRTHP29m99B23Uiq6dZisgoWCxVJW0oL9dqLfyhCvGWY2YUBI95MsXgwFISirQcLhS5hfgk5CF3wJIhFI7aHNsL2VFZFik1jS1qkiZKdTTKZ3/MyDWuz0fQLc/m99/vMvDfv+4RMlUrlkKqeAAaBAWAP8DSgwJ/AXRG5rao/WWsvTU5O3qKLBMD3fSMiPluXFZEPoyj67PGAMzw83PeEMABHVT/oGpiamnoAmCcEWhH5tFsgF4bh9oWFhfeKxeJ5a+0JVT0oImWgBPQCKfAQuAvcBq67rltX1b+6ApMkKRcKhe9V9QLwbavV+qRer692Sx4ZGSnEcXw0TdP3gSrQswGYz+d/S5IkVtXTwOlCoZAGQXAfmAdagAvsAErtdnuXiDy6+023l7qNRsMODg5+CawBzwB9wFPA7mx8ns/KL2Tl3xCRz5eWlkabzebahrHxPG+v4zgnc7ncufHx8Z+Hhoa29fT0lNM03Q30ikiqqg+ttX/EcTy3WTvWgdVqtddaOw/kgXvADHBHROZVNRaRvKruUNU+EdkPfGWM+WJTYOaSt1T1LPDS/4zLWWPMaLVaPWytrYvIaBRFl/4F9H2/JCKvGmMu+76/X0QOqGoZKDmOs1NV28AicMsYc97zvFdc1/0hG6kEeNsY83UnsCwivwM3VfU7YEZE7lhr74tIK8tbnJiYWPY8b6/ruleAXR0ftQy8boyZXi85CIIICDYpc2ZgYODY3NzcHmvt1eyvP64lETkeRdE1yZyixWLx5U2c8q4x5mIQBE1g33/0d3FlZeXFR06ZttZesNZejuO4q1NE5CPgWVV9E3ij47wB1IDlJEn+ljAM86urq7+KyAtZTgqsO0VV247jnOnv7/9xbGzMViqVMVX9uANYj6LonfVtU6vVkjRNj6jqGeCXzGrPAQeA10TkuKpOz87ONrayhnIA2Qo7BZwKw3B7kiRloKSqO13Xja21C47jPNgysFO1Wi0GmtmzQap6DWgD24A1Vb3SGf8Hfstmz1CuXEIAAAAASUVORK5CYII=" /> na barra de ferramentas.
# + tags=["parameters"]
# dataset = "/tmp/data/paracrawl_en_pt_test.csv" #@param {type:"string"}
dataset = "pt_en_testing_texts.csv"
text = "text_english" #@param {type:"string", label:"Atributo do texto", description:"Este atributo será traduzido e apresentado o resultado."}
target = "text_portuguese" #@param {type:"string", label:"Atributo alvo", description:"Seu modelo será validado com os valores do alvo."}
input_language = "Português" #@param ["Alemão", "Catalão", "Espanhol", "Francês", "Inglês", "Italiano", "Latim", "Português", "Romeno"] {type:"string", label:"Idioma de entrada"}
target_language = "Inglês" #@param ["Alemão", "Catalão", "Espanhol", "Francês", "Inglês", "Italiano", "Latim", "Português", "Romeno"] {type:"string", label:"Idioma de saída"}
#Hyperparams
seed = 42 #@param {type:"integer",label"Semente de aleatoriedade"}
max_length = 256 #@param {type:"integer",label"Tamanho máximo da sentença de entrada que entrará no MarianMT. Se a sentença for maior ela será quebrada"}
#model_name = 'Helsinki-NLP/opus-mt-ROMANCE-en' #@param ["Helsinki-NLP/opus-mt-ROMANCE-en","Helsinki-NLP/opus-mt-en-ROMANCE"] {type:"integer",label:"Helsinki/NLP model name"}
inference_batch_size = 2 #@param {type:"integer",label"Tamanho do Batch de inferência"}
# -
if input_language == target_language:
raise Exception('Idioma de entrada e de saída não podem ser iguais')
# ## Acesso ao conjunto de dados
#
# O conjunto de dados utilizado nesta etapa será o mesmo carregado através da plataforma.<br>
# O tipo da variável retornada depende do arquivo de origem:
# - [pandas.DataFrame](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.html) para CSV e compressed CSV: .csv .csv.zip .csv.gz .csv.bz2 .csv.xz
# - [Binary IO stream](https://docs.python.org/3/library/io.html#binary-i-o) para outros tipos de arquivo: .jpg .wav .zip .h5 .parquet etc
# +
import pandas as pd
df = pd.read_csv(dataset)
# -
# ## Formatar dados
#
# Dados nulos serão removidos e serão separados em colunas de teste e validação.
# +
df = df.dropna()
X = df[text].to_numpy()
y = None
if target in df.columns:
y = df[target].to_numpy()
# -
# ## Verificando as configurações do MarianMT
#
# - Verificando disponibilidade de GPU e status de hardware
# - Instanciando modelo e tokenizador
# - Opções de tradução de idiomas
# - Modelos pré treinados disponíveis
# +
from multiprocessing import cpu_count
import torch
dev = "cuda:0" if torch.cuda.is_available() else "cpu"
device = torch.device(dev)
if dev == "cpu":
print(f"number of CPU cores: {cpu_count()}")
else:
print(f"GPU: {torch.cuda.get_device_name(0)}, number of CPU cores: {cpu_count()}")
# -
# ## Chamada da Classe MarianMT
if input_language=='Inglês' and target_language!='Inglês':
models_list = ["Helsinki-NLP/opus-mt-en-ROMANCE"]
elif input_language!='Inglês' and target_language=='Inglês':
models_list = ["Helsinki-NLP/opus-mt-ROMANCE-en"]
else:
models_list = ["Helsinki-NLP/opus-mt-ROMANCE-en","Helsinki-NLP/opus-mt-en-ROMANCE"]
# +
from marianmt_model import MarianMTTranslator
hyperparams = {'max_length': max_length,
'inference_batch_size': inference_batch_size,
'target_language':target_language,
'models_list':models_list,
'seed':seed
}
marianmt_model = MarianMTTranslator(hyperparams)
# -
if y is not None:
aux = marianmt_model.get_result_dataframe(X,y)
else:
aux = marianmt_model.predict(X)
aux
# ## Salva métricas
#
# Utiliza a função `save_metrics` do [SDK da PlatIAgro](https://platiagro.github.io/sdk/) para salvar métricas. Por exemplo: `accuracy`, `precision`, `r2_score`, `custom_score` etc.<br>
# +
from platiagro import save_metrics
if y is not None:
save_metrics(avg_bleu=marianmt_model.avg_bleu)
# -
# ## Salva resultados da tarefa
#
# A plataforma guarda o conteúdo de `/tmp/data/` para as tarefas subsequentes.
# +
from joblib import dump
artifacts = {
"model": marianmt_model
}
dump(artifacts, "/tmp/data/translate.joblib")
| tasks/nlp-marianmt-translator/Experiment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (crypto_predict)
# language: python
# name: crypto_predict
# ---
# # What's the data like?
# +
# %load_ext autoreload
# %autoreload 2
p = print
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
import datetime
import seaborn as sns
import os, dotenv
coin = 'BTC'
project_path = os.path.dirname(dotenv.find_dotenv())
raw_data_path = os.path.join(project_path, 'data', 'raw', coin + '.csv')
# -
"""
load data
"""
df = pd.read_csv(raw_data_path, index_col=0)
p('data shape is: ', df.shape)
df.head()
"""
descriptive stats
"""
df.describe()
"""
Confirm that no NA values are present
"""
p('nan values in data: ', df.dropna().shape != df.shape)
"""
Take a small sample for plotting
"""
sample = df.sample(n=1000, replace=False)
"""
Check relative distribution of data features
"""
sns.boxplot(data=sample[['close', 'high', 'low', 'open', 'volumefrom']])
# +
"""
Plot data features against time. Making sure that the data set was created correctly / sanity check.
"""
pltdf = sample.copy(). \
melt(id_vars='time')
fig, ax = plt.subplots(3, sharex=True, figsize=(10, 14))
sns.pointplot(ax=ax[0], data=pltdf[pltdf.variable.isin(['low', 'high', 'close'])],
x='time', y='value', hue='variable', linestyles='-', markers='')
sns.pointplot(ax=ax[1], data=pltdf[pltdf.variable.isin(['volumefrom'])],
x='time', y='value', linestyles='-', markers='')
sns.pointplot(ax=ax[2], data=pltdf[pltdf.variable.isin(['volumeto'])],
x='time', y='value', linestyles='-', markers='')
| notebooks/data_exploration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
from scipy import *
import os
import h5py
import keras as k
from keras.layers import Dense, Activation
from keras.optimizers import SGD
from keras.models import Sequential
from keras.losses import categorical_crossentropy
from sklearn.utils import shuffle
from keras.utils import np_utils
import matplotlib.pyplot as plt
from astropy.visualization import *
# +
# Setting directory paths
root_dir = os.getcwd()
# Loading data
data = h5py.File(r'C:\\Users\\fires\\Desktop\\CSIRO\\thursday\\Logistic Regression\\data.h5', 'r')
images = np.asarray(data['images'])
labels = data['labels']
# -
# Define function to crop off 'cut' number of pixels on both sides on both axixs
def crop_center(img,cut):
y = np.shape(img)[1]
x = np.shape(img)[2]
if x != y:
print ("The image is not a perfect sqaure. This is bad. Fix it ")
start = cut
end = x-cut
return img[:, start:end,start:end]
# Reducing Image Size
images_cut = crop_center(images,66)
images_cut = np.asarray(images_cut)
# +
# Reshape data
img = np.reshape(images_cut, (-1, 18225))
labels = np.reshape(labels, (-1,1))
# +
##### This section is horrendous, I know #####
# Normalising the number of examples for each class
nb_FRI = int(np.asarray((np.where(labels == False))).size/2)
nb_FRII = int(np.asarray((np.where(labels == True))).size/2)
# Defining the maximum number of samples for each class
if nb_FRI >= nb_FRII:
cutoff = nb_FRII
elif nb_FRI < nb_FRII:
cutoff = nb_FRI
# seperating images and labels by class
FRI = img[:nb_FRI, :]
FRII = img[nb_FRI:, :]
FRI_labels = labels[:nb_FRI]
FRII_labels = labels[nb_FRI:]
# Slicing off excess samples
FRI = FRI[:cutoff, :]
FRI_labels = FRI_labels[:cutoff]
# Putting classes back together
normalized_img = np.vstack((FRI, FRII))
normalized_labels = np.vstack((FRI_labels, FRII_labels ))
# Randomizing the order of the rows
normalized_img, normalized_labels = shuffle(normalized_img, normalized_labels)
# +
# Stretch Images
count = 0
for i, image in enumerate(normalized_img):
img = PowerStretch(image)
if count == 0:
stretched_img = np.expand_dims(img, axis=0)
else:
stretched_img = np.vstack((stretched_img, np.expand_dims(img, axis=0) ))
count += 1
print (stretched_img)
# +
imgs = np.reshape(stretched_img, (-1, 135, 135))
for n in range(0, stretched_img.shape[0]):
plt.imshow(stretched_img[n,:,:], cmap='gray', shape=(135, 135))
print (n)
print (normalized_labels[n])
plt.show()
# +
# Split data into training and testing sets
ratio = 0.8
def format_data(images, labels):
split = round(ratio*images.shape[0])
train_x = images[ :split, :]
test_x = images[split:, :]
train_y = labels[ :split]
test_y = labels[split:]
return train_x, test_x, train_y, test_y
train_x, test_x, train_y, test_y = format_data(normalized_img, normalized_labels)
# +
# Convert class vectors to binary class matrices
train_y = np_utils.to_categorical(train_y, 2)
test_y = np_utils.to_categorical(test_y, 2)
print (np.shape(train_x))
# +
# Define model attributes
batch_size = 5
nb_classes = output_dims = 2
nb_epoch = 1000
input_dim = 18225
model = Sequential()
model.add(Dense(output_dims, input_dim = input_dim, activation='sigmoid'))
# -
sgd = SGD(lr=1)
model.compile(optimizer=sgd, loss='categorical_crossentropy',
metrics=['accuracy'])
model.fit(train_x, train_y, epochs=nb_epoch, batch_size=batch_size,
validation_data=(test_x, test_y))
| Legacy Machine Learning Methods/Logistic Regression/Logistic Regression Stretched - Keras.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import random
import numpy as np
import pandas as pd
import time
from redis import Redis
from redis.commands.search.field import VectorField
from redis.commands.search.field import TextField
from redis.commands.search.field import TagField
from redis.commands.search.query import Query
from PIL import Image
from img2vec_pytorch import Img2Vec
class color:
PURPLE = '\033[95m'
CYAN = '\033[96m'
DARKCYAN = '\033[36m'
BLUE = '\033[94m'
GREEN = '\033[92m'
YELLOW = '\033[93m'
RED = '\033[91m'
BOLD = '\033[1m'
UNDERLINE = '\033[4m'
END = '\033[0m'
# + [markdown] tags=[]
# # Load Amazon Product and Image metadata
#
#
# -
# ?TagField
#Load Product data and truncate long text fields
all_prods_df = pd.read_csv('data/product_image_data.csv')
all_prods_df['primary_key'] = all_prods_df['item_id'] + '-' + all_prods_df['domain_name']
all_prods_df.shape
all_prods_df.head(5)
# + [markdown] tags=[]
# # Connect to Redis
# -
host = 'vecsim'
port = 6379
redis_conn = Redis(host = host, port = port)
redis_conn.ping()
print ('Connected to redis')
# + [markdown] tags=[]
# # Generate Embeddings
#
# We will use 'Img2Vec' to generate embeddings (vectors) for 1K product images
#
# https://github.com/christiansafka/img2vec
# -
img2vec = Img2Vec(cuda=False)
# + [markdown] tags=[]
#
# By Default, Img2Vect uses **'resnet-18'** as the neural network architecture to generate embeddings. In particular, each image is run through this network and the output at the 'avgpool' layer will be returned
#
# The output of the 'avgpool' layer in **'resnet-18' has 512 dimensions** so a single 512-float vector will be generated for every image converted
# +
NUMBER_PRODUCTS = 1000
subset_df = all_prods_df.head(NUMBER_PRODUCTS)
# -
subset_df.head()
subset_df.shape
#transform the df into a dictionary
product_metadata = subset_df.to_dict(orient='index')
# Check one of the products
product_metadata[0]
# # Some Utility Functions to Generate Vectors from Images
#
# Product images are stored under the 'data/small' folder
#
# Every product has metadata indicating the full path to the main product image
#
#
# The 'generate_img2vec_dict' function below simply takes:
# * A dataframe with product metadata
# * The folder where images are stored
# * A batch size to generate image vectors for a batch of products in one call
#
# The output will be a dictionary mapping 'full image path' to its corresponding vector generated
# +
def chunker(seq, size):
return (seq[pos:pos + size] for pos in range(0, len(seq), size))
def generate_img2vec_dict(df,image_path, batch_size=100):
output_dict={}
for batch in chunker(df, batch_size):
image_filenames=batch['path'].values.tolist()
images=[]
converted=[]
for img_fn in image_filenames:
try:
img = Image.open(image_path + img_fn)
images.append(img)
converted.append(img_fn)
except:
#unable_to_convert -> skip to the next image
continue
#Generate vectors for all images in this batch
vec_list = img2vec.get_vec(images)
#update the dictionary to be returned
batch_dict= dict(zip(converted, vec_list))
output_dict.update(batch_dict)
return output_dict
# -
# ### Time to generate the vectors!
#
# This may take 30-60 seconds depending on your set up
# +
# %%time
IMAGE_PATH= './data/images/small/'
img2vec_dict = generate_img2vec_dict(subset_df,IMAGE_PATH,batch_size=250)
# +
#img2vec_dict['30/3079540e.jpg']
# + [markdown] tags=[]
# # Utility Functions to Load Product metadata and image data
# Each product will be stored in a redis hash
# * **Hash Key** = **product:primary_key**
# * **Hash Fields:**
# * Item Id
# * Item Name
# * Product Image vector = 512-float vector
#
# -
def load_vectors(client:Redis, product_metadata, vector_dict, vector_field_name):
p = client.pipeline(transaction=False)
for index in product_metadata.keys():
#hash key
key='product:'+ product_metadata[index]['primary_key']
#hash values
item_metadata = product_metadata[index]
item_path = item_metadata['path']
if item_path in vector_dict:
#retrieve vector for product image
product_image_vector = vector_dict[item_path].astype(np.float32).tobytes()
item_metadata[vector_field_name]=product_image_vector
# HSET
#p.hset(key,mapping=product_data_values)
p.hset(key,mapping=item_metadata)
p.execute()
# # Utility Functions to Create Indexes on Vector field
# +
def create_flat_index (redis_conn,vector_field_name,number_of_vectors, vector_dimensions=512, distance_metric='L2'):
redis_conn.ft().create_index([
VectorField(vector_field_name, "FLAT", {"TYPE": "FLOAT32", "DIM": vector_dimensions, "DISTANCE_METRIC": distance_metric, "INITIAL_CAP": number_of_vectors, "BLOCK_SIZE":number_of_vectors }),
TagField("product_type"),
TextField("item_name"),
TagField("country")
])
def create_hnsw_index (redis_conn,vector_field_name,number_of_vectors, vector_dimensions=512, distance_metric='L2',M=40,EF=200):
redis_conn.ft().create_index([
VectorField(vector_field_name, "HNSW", {"TYPE": "FLOAT32", "DIM": vector_dimensions, "DISTANCE_METRIC": distance_metric, "INITIAL_CAP": number_of_vectors, "M": M, "EF_CONSTRUCTION": EF}),
TagField("product_type"),
TextField("item_name"),
TagField("country")
])
# + [markdown] tags=[]
# # FLAT - Load and Index 1000 Products
#
# Let's create a FLAT index for the image vectors and load 1000 hashes
#
# A FLAT index is used to perform an exact nearest neighbors search.
#
# The query vector will be compared against all other image vectors in the database
# +
# %%time
PRODUCT_IMAGE_VECTOR_FIELD='product_image_vector'
IMAGE_VECTOR_DIMENSION=512
print ('Loading and Indexing + ' + str(NUMBER_PRODUCTS) + ' products')
#flush all data
redis_conn.flushall()
#create flat index & load vectors
create_flat_index(redis_conn, PRODUCT_IMAGE_VECTOR_FIELD,NUMBER_PRODUCTS,IMAGE_VECTOR_DIMENSION,'COSINE')
load_vectors(redis_conn,product_metadata,img2vec_dict,PRODUCT_IMAGE_VECTOR_FIELD)
# -
# # FLAT index - FIND The Top K MOST VISUALLY Similar Products
# Let's use the FLAT index to find the exact top K nearest neighbors of a mobile phone cover available in the catalogue
#
#
pos=0
print (product_metadata[pos]['item_name'])
print (product_metadata[pos]['path'])
queryImage = Image.open(IMAGE_PATH + product_metadata[pos]['path'])
queryImage
# +
# %%time
topK=5
query_vector = img2vec.get_vec(queryImage).astype(np.float32).tobytes()
#prepare the query
q = Query(f'*=>[KNN {topK} @{PRODUCT_IMAGE_VECTOR_FIELD} $vec_param AS vector_score]').sort_by('vector_score').paging(0,topK).return_fields('vector_score','item_name','item_id','path').dialect(2)
params_dict = {"vec_param": query_vector}
#Execute the query
results = redis_conn.ft().search(q, query_params = params_dict)
docs = redis_conn.ft().search(q,params_dict).docs
#Print similar products found
for product in results.docs:
print ('***************Product found ************')
print (color.BOLD + 'hash key = ' + color.END + product.id)
print (color.YELLOW + 'Item Name = ' + color.END + product.item_name)
print (color.YELLOW + 'Item Id = ' + color.END + product.item_id)
print (color.YELLOW + 'Score = ' + color.END + product.vector_score)
result_img= Image.open(IMAGE_PATH + product.path)
display(result_img)
# -
# ## Examine Search Results
#
# You can see the redis hash fields projected in the query (e.g item_name, item_path,item_id).
#
# The score field returns the distance between the query vector to each of the vectors in the result
results.docs
# # HNSW - Load and Index Product Data
#
# Let's repeat the exercise of loading and indexing 1000 products but this time using an HNSW index
#
# This HNSW index is used to calculate Approximate Nearest Neighbors (ANN) of a given vector image.
#
# It speeds up query times but requires more memory to store the vector index
# %%time
print ('Loading and Indexing + ' + str(NUMBER_PRODUCTS) + ' products')
#flush all data
redis_conn.flushall()
#create HNSW index & load vectors
create_hnsw_index(redis_conn,PRODUCT_IMAGE_VECTOR_FIELD,NUMBER_PRODUCTS,IMAGE_VECTOR_DIMENSION,'COSINE',M=40,EF=200)
load_vectors(redis_conn,product_metadata,img2vec_dict,PRODUCT_IMAGE_VECTOR_FIELD)
# # HNSW - Query the top 5 most visually similar products
# Let's repeat the similarity search but this time using the HNSW index.
#
# Let's see the image we're sending in for visual similarity
#
#
#
queryImage
# +
# %%time
topK=5
query_vector = img2vec.get_vec(queryImage).astype(np.float32).tobytes()
EF_RUNTIME=10
#prepare the query
q = Query(f'*=>[KNN {topK} @{PRODUCT_IMAGE_VECTOR_FIELD} $vec_param EF_RUNTIME {EF_RUNTIME} AS vector_score]').sort_by('vector_score').paging(0,topK).return_fields('vector_score','item_name','item_id','path').dialect(2)
params_dict = {"vec_param": query_vector}
#Execute the query
results = redis_conn.ft().search(q, query_params = params_dict)
docs = redis_conn.ft().search(q,params_dict).docs
#Print similar products found
for product in results.docs:
print ('***************Product found ************')
print (color.BOLD + 'hash key = ' + color.END + product.id)
print (color.YELLOW + 'Item Name = ' + color.END + product.item_name)
print (color.YELLOW + 'Item Id = ' + color.END + product.item_id)
print (color.YELLOW + 'Score = ' + color.END + product.vector_score)
result_img= Image.open(IMAGE_PATH + product.path)
display(result_img)
# -
results.docs
# # HNSW - Hybrid Query the top 5 most visually similar products ONLY in selected markets
#
# Let's repeat our Top 5 search but this time limit to products that meet the following criteria:
# * **Listed on** Amazon Germany (DE), United States (US) or Italy (IT) **AND**
# * **Product type** = CELLULAR_PHONE_CASE
#
#
# This RediSearch query has this form:
#
# **(@country:{{DE|US|IT}} @product_type:{{CELLULAR_PHONE_CASE}})=> [KNN 5 vector_field_name $query_vector EF_RUNTIME 10 AS vector_score])**
#
#
# Note that there is only 1 matching this criteria
# +
# %%time
topK=5
query_vector = img2vec.get_vec(queryImage).astype(np.float32).tobytes()
EF_RUNTIME=10
#prepare the query
q = Query(f'(@country:{{DE|US|IT}} @product_type:{{CELLULAR_PHONE_CASE}})=>[KNN {topK} @{PRODUCT_IMAGE_VECTOR_FIELD} $vec_param EF_RUNTIME {EF_RUNTIME} AS vector_score]').sort_by('vector_score').paging(0,topK).return_fields('vector_score','item_name','item_id','path','country').dialect(2)
params_dict = {"vec_param": query_vector}
#Execute the query
results = redis_conn.ft().search(q, query_params = params_dict)
docs = redis_conn.ft().search(q,params_dict).docs
#Print similar products found
for product in results.docs:
print ('***************Product found ************')
print (color.BOLD + 'hash key = ' + color.END + product.id)
print (color.YELLOW + 'Item Name = ' + color.END + product.item_name)
print (color.YELLOW + 'Item Id = ' + color.END + product.item_id)
print (color.YELLOW + 'Score = ' + color.END + product.vector_score)
print (color.YELLOW + 'Country = ' + color.END + product.country)
result_img= Image.open(IMAGE_PATH + product.path)
display(result_img)
# -
docs
| VisualSearch1k.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # MALARIA DETECTION
#
# ## Motivation to work on the Malaria Detection problem
#
# Malaria is caused by Plasmodium parasites. The parasites are spread to people through the bites of infected female Anopheles mosquitoes, called "malaria vectors." There are 5 parasite species that cause malaria in humans, and 2 of these species – P. falciparum and P. vivax – pose the greatest threat.
#
# Malaria is an acute febrile illness. In a non-immune individual, symptoms usually appear 10–15 days after the infective mosquito bite. The first symptoms – fever, headache, and chills – may be mild and difficult to recognize as malaria. If not treated within 24 hours, P. falciparum malaria can progress to severe illness, often leading to death.
#
# 
#
# According to WHO protocol, diagnosis typically involves intensive examination of the blood smear at 100X magnification. Trained people manually count how many red blood cells contain parasites out of 5,000 cells. Thus, the detection of Malaria should be automized for the fast detection and treatment to prevent life losses.
#
# ## Dataset
#
# The dataset for Malaria detection was carefully collected and annotated by the researchers at the Lister Hill National Center for Biomedical Communications (LHNCBC), part of the National Library of Medicine (NLM). The dataset is [publicly available](https://ceb.nlm.nih.gov/repositories/malaria-datasets/). The dataset contains a total of 27,558 cell images with equal instances of parasitized and uninfected cells.
#
import dl
import numpy as np
import os.path
import matplotlib.pyplot as plt
import cv2
import h5py
from random import shuffle
from math import ceil
# +
from malaria_detection import read_images, train_test_set
from collections import Counter
# in the cell_images.zip file, we have 27558 cell images. Half of them are parasitized and the other half are
# the images of uninfected cells.
# Firstly, randomly mix the data one file. Below function will mix all the images in infected and healthy image files
# into one file.
files_df = read_images()
# Now, let's create train, validation, and the test splits of data.
# %30 of all data will be the test set.
# The rest %70 of data will be divided into %90 train and %10 validation data.
train_files, train_labels, val_files, val_labels, test_files, test_labels = train_test_set(files_df)
print("Train data shape : ", train_files.shape)
print("Validation data shape : ", val_files.shape)
print("Test data shape : ", test_files.shape)
print('Train:', Counter(train_labels), '\nVal:', Counter(val_labels), '\nTest:', Counter(test_labels))
# +
from malaria_detection import discover_dataset
# After making train, validation and test splits, we need to resize the images, since all of them have different sizes.
# To find the best size, we will go through all the train set and will find maximum, minimum, median and average
# image sizes.
train_img_dims = discover_dataset(train_files)
print('Min Dimensions:', np.min(train_img_dims, axis=0))
print('Avg Dimensions:', np.mean(train_img_dims, axis=0))
print('Median Dimensions:', np.median(train_img_dims, axis=0))
print('Max Dimensions:', np.max(train_img_dims, axis=0))
# +
# This operation might take a while.
from malaria_detection import load_images
from sklearn.preprocessing import LabelEncoder
IMG_DIM = (100, 100) # Here you can change the size.
hdf5_file = None
###################################################
hdf5_datapath = '../Malaria_Dataset/data_28.hdf5' # Here you can directly define the path to your hdf5 data file.
# It can only be used ../Malaria_Dataset/data_28.hdf5 or ../Malaria_Dataset/data_64.hdf5
###################################################
if not os.path.isfile(hdf5_datapath):
train_shape = (len(train_files), 100, 100 , 3)
val_shape = (len(val_files), 100, 100, 3)
test_shape = (len(test_files), 100, 100, 3)
# In the label files, all images anotated either as healthy or malaria.
# By using the label encoder, we will encode all of them as 0-healthy and 1-malaria
le = LabelEncoder()
le.fit(train_labels)
train_labels_enc = le.transform(train_labels)
val_labels_enc = le.transform(val_labels)
test_labels_enc = le.transform(test_labels)
print(train_labels[:6], train_labels_enc[:6])
hdf5_file = h5py.File(hdf5_datapath, mode='w')
hdf5_file.create_dataset("train_img", train_shape, np.float32)
hdf5_file.create_dataset("val_img", val_shape, np.float32)
hdf5_file.create_dataset("test_img", test_shape, np.float32)
hdf5_file.create_dataset("train_mean", train_shape[1:], np.float32)
hdf5_file.create_dataset("train_labels", (len(train_labels),), np.int8)
hdf5_file["train_labels"][...] = train_labels_enc
hdf5_file.create_dataset("val_labels", (len(val_labels),), np.int8)
hdf5_file["val_labels"][...] = val_labels_enc
hdf5_file.create_dataset("test_labels", (len(test_labels),), np.int8)
hdf5_file["test_labels"][...] = test_labels_enc
mean = np.zeros(train_shape[1:], np.float32)
# Here all the images are loaded into hdf5 file.
load_images(train_files, train_labels, val_files, test_files, mean, hdf5_file)
hdf5_file.close() # After loading of all images hdf5 file is closed.
# +
# Let's plot some random samples from our training samples, to see our data format properly.
# The hdf5 file is opened in the read format.
hdf5_file = h5py.File(hdf5_datapath, "r")
# the keys to reach the correct data in hdf5 file.
# hdf5_file["train_img"] --> training images
# hdf5_file["train_labels"] --> training labels
# hdf5_file["val_img"] --> validation images
# hdf5_file["val_labels"] --> validation labels
# hdf5_file["test_img"] --> test images
# hdf5_file["test_labels"] --> test labels
# hdf5_file["train_mean"] --> the mean of all training data.
plt.figure(1 , figsize = (8 , 8))
n = 0
for i in range(16):
n += 1
r = np.random.randint(0 , hdf5_file["train_img"].shape[0] , 1)
plt.subplot(4 , 4 , n)
plt.subplots_adjust(hspace = 0.5 , wspace = 0.5)
plt.imshow(hdf5_file["train_img"][r[0]]/255.)
plt.title('{}'.format(train_labels[r[0]]))
plt.xticks([]) , plt.yticks([])
plt.show()
# -
# ## Mini Batch Calculation
# +
BATCH_SIZE = 10 # You can change the batch size to something else.
EPOCH_SIZE = 3 # You can change the number of epochs.
data_num = hdf5_file["train_img"].shape[0] # Total number of samples
validation_set_num = hdf5_file["val_img"].shape[0]
# create list of batches to shuffle the data
batches_list = list(range(int(ceil(float(data_num) / BATCH_SIZE))))
shuffle(batches_list)
# create list of validation batches to shuffle the data
val_batches_list = list(range(int(ceil(float(validation_set_num) / BATCH_SIZE))))
shuffle(val_batches_list)
# -
# ## Building LeNet Architecture
#
# There are two different architectures which is used.
# First of all LeNet is used with the input size 28x28x3 images first, later with the image size 64x64x3.
# +
net = dl.Network()
# LeNet Architecture
output_size_conv1, num_of_outputs_conv1 = net.conv(hdf5_file['train_img'][0].shape[0],
hdf5_file['train_img'][0].shape[2],
5, # filter size
3, # filter depth
6, # number of filters
1, # stride
0) # padding
output_size_relu1_h, output_size_relu1_w, num_of_outputs_relu1 = net.relu(
num_of_outputs_conv1, # number of the feature maps
output_size_conv1, #height of one of the feature map
output_size_conv1 # width of one of the feature map
)
output_size_pooling1, num_of_outputs_pooling_1 = net.maxpool(
output_size_relu1_h,
num_of_outputs_relu1,
2, # filter size
2, # stride
0 # padding
)
output_size_conv2, num_of_outputs_conv2 = net.conv(
output_size_pooling1,
num_of_outputs_pooling_1,
5, # filter size
6, # filter dept
16, # number of filters
1, #stride
0 # padding
)
output_size_relu2_h, output_size_relu2_w, num_of_outputs_relu2 = net.relu(
num_of_outputs_conv2, # number of the feature maps
output_size_conv2, #height of one of the feature map
output_size_conv2 # width of one of the feature map
)
output_size_pooling2, num_of_outputs_pooling_2 = net.maxpool(
output_size_relu2_h,
num_of_outputs_relu2,
2, # filter size
2, # stride
0 # padding
)
output_size_fully1_h, output_size_fully1_w, num_of_outputs_fully1 = net.fullyConnected(
output_size_pooling2,
output_size_pooling2,
num_of_outputs_pooling_2,
32
)
output_size_relu4_h, output_size_relu4_w, num_of_outputs_relu4 = net.relu(
num_of_outputs_fully1, # number of the feature maps
output_size_fully1_h, # height of one of the feature map
output_size_fully1_w # width of the feature map
)
output_size_fully2_h, output_size_fully2_w, num_of_outputs_fully2 = net.fullyConnected(
output_size_relu4_h,
output_size_relu4_w, # 1
num_of_outputs_relu4,
2
)
output_size_softmax5_h, output_size_softmax5_w, num_of_outputs_sigmoid5 = net.softmax(
num_of_outputs_fully2, # number of the feature maps
output_size_fully2_h, #height of one of the feature map,
output_size_fully2_w # 1
)
# -
# ## Testing of LeNet Network Architecture with SGD by Using 28x28x3 images
# +
net.loadWeights("../Malaria_Dataset/training_sgd_normal_28/training_sgd_28.txt")
n_values = 2
test_data_num = hdf5_file["test_labels"].shape[0]
#####################################################
# Since we don't want to load all the data at once, #
# we need to divide it into batches. From the below #
# line you can change the batch size for the path. #
#####################################################
TEST_BATCH_SIZE = 10
test_batches_list = list(range(int(ceil(float(test_data_num) / TEST_BATCH_SIZE))))
shuffle(test_batches_list)
num_of_correct_guesses = 0
num_of_correct_guesses_10 = 0
for n, i in enumerate(test_batches_list):
i_s = i * TEST_BATCH_SIZE # index of the first image in this batch
i_e = min([(i + 1) * TEST_BATCH_SIZE, test_data_num]) # index of the last image in this batch
# read batch images and remove training mean
test_images = hdf5_file["test_img"][i_s:i_e, ...]
test_images = test_images.reshape(test_images.shape[0], test_images.shape[3],
test_images.shape[1], test_images.shape[2]) # reshaped to size 22046, 3, 64, 64
# read labels and convert to one hot encoding
test_labels = hdf5_file["test_labels"][i_s:i_e]
test_labels_one_hot = np.eye(n_values)[test_labels]
temp = net.test(test_images / 255., test_labels_one_hot)
num_of_correct_guesses += temp
num_of_correct_guesses_10 += temp
if n % 10 == 9: # print every 10 batches
print('Batch: {}, Accuracy: {}'.format(n+1,num_of_correct_guesses_10 * 100 / float(TEST_BATCH_SIZE * 10)))
num_of_correct_guesses_10 = 0
print('Total test Accuracy : {}'.format(num_of_correct_guesses / float(test_data_num)))
# -
# ## Visual Test Results of LeNet Network Architecture with SGD
# Below part is for picking a random 16 images, you can run it again to see the test results with different images.
plt.figure(1 , figsize = (8 , 8))
n = 0
mask = []
for i in range(16):
n += 1
r = np.random.randint(0 , hdf5_file["train_img"].shape[0] , 1)
mask.append(r[0])
plt.subplot(4 , 4 , n)
plt.subplots_adjust(hspace = 0.5 , wspace = 0.5)
plt.imshow(hdf5_file["train_img"][r[0]]/255.)
plt.title('{}'.format(train_labels[r[0]]))
plt.xticks([]) , plt.yticks([])
plt.show()
predictions = []
for sample_idx in mask:
sample = hdf5_file["train_img"][sample_idx]/ 255.
sample = sample.reshape(sample.shape[2], sample.shape[0], sample.shape[1])
label = hdf5_file["train_labels"][sample_idx]
predicted_label = net.predict(sample)
if predicted_label == 0:
predictions.append("healthy")
else:
predictions.append("malaria")
# ## Predictions
plt.figure(1 , figsize = (8 , 8))
n = 0
for i, sample_idx in enumerate(mask):
n += 1
plt.subplot(4 , 4 , n)
plt.subplots_adjust(hspace = 0.5 , wspace = 0.5)
plt.imshow(hdf5_file["train_img"][sample_idx]/255.)
plt.title('{}'.format(predictions[i]))
plt.xticks([]) , plt.yticks([])
plt.show()
# ## Testing of LeNet Network Architecture with Momentum Gradient Descent by Using 28x28x3 images
# +
net.loadWeights("../Malaria_Dataset/training_momentum_normal_28/training_normal_28_epoch_3.txt")
n_values = 2
test_data_num = hdf5_file["test_labels"].shape[0]
#####################################################
# Since we don't want to load all the data at once, #
# we need to divide it into batches. From the below #
# line you can change the batch size for the path. #
#####################################################
TEST_BATCH_SIZE = 10
test_batches_list = list(range(int(ceil(float(test_data_num) / TEST_BATCH_SIZE))))
shuffle(test_batches_list)
num_of_correct_guesses = 0
num_of_correct_guesses_10 = 0
for n, i in enumerate(test_batches_list):
i_s = i * TEST_BATCH_SIZE # index of the first image in this batch
i_e = min([(i + 1) * TEST_BATCH_SIZE, test_data_num]) # index of the last image in this batch
# read batch images and remove training mean
test_images = hdf5_file["test_img"][i_s:i_e, ...]
test_images = test_images.reshape(test_images.shape[0], test_images.shape[3],
test_images.shape[1], test_images.shape[2]) # reshaped to size 22046, 3, 64, 64
# read labels and convert to one hot encoding
test_labels = hdf5_file["test_labels"][i_s:i_e]
test_labels_one_hot = np.eye(n_values)[test_labels]
temp = net.test(test_images / 255., test_labels_one_hot)
num_of_correct_guesses += temp
num_of_correct_guesses_10 += temp
if n % 10 == 9: # print every 10 batches
print('Batch: {}, Accuracy: {}'.format(n+1,num_of_correct_guesses_10 * 100 / float(TEST_BATCH_SIZE * 10)))
num_of_correct_guesses_10 = 0
print('Total test Accuracy : {}'.format(num_of_correct_guesses / float(test_data_num)))
# -
# ## Visual Test Results of LeNet Network Architecture with Momentum
predictions = []
for sample_idx in mask:
sample = hdf5_file["train_img"][sample_idx]/ 255.
sample = sample.reshape(sample.shape[2], sample.shape[0], sample.shape[1])
label = hdf5_file["train_labels"][sample_idx]
predicted_label = net.predict(sample)
if predicted_label == 0:
predictions.append("healthy")
else:
predictions.append("malaria")
# ## Predictions
plt.figure(1 , figsize = (8 , 8))
n = 0
for i, sample_idx in enumerate(mask):
n += 1
plt.subplot(4 , 4 , n)
plt.subplots_adjust(hspace = 0.5 , wspace = 0.5)
plt.imshow(hdf5_file["train_img"][sample_idx]/255.)
plt.title('{}'.format(predictions[i]))
plt.xticks([]) , plt.yticks([])
plt.show()
# ## Testing of LeNet Network Architecture with SGD by Using 64x64x3 images
# Before running this part, don't forget to change the hdf5_datapath to ../Malaria_Dataset/data_64.hdf5
#
# +
net.loadWeights("../Malaria_Dataset/training_sgd_normal_64/training_sgd_normal_64.txt")
n_values = 2
test_data_num = hdf5_file["test_labels"].shape[0]
#####################################################
# Since we don't want to load all the data at once, #
# we need to divide it into batches. From the below #
# line you can change the batch size for the path. #
#####################################################
TEST_BATCH_SIZE = 10
test_batches_list = list(range(int(ceil(float(test_data_num) / TEST_BATCH_SIZE))))
shuffle(test_batches_list)
num_of_correct_guesses = 0
num_of_correct_guesses_10 = 0
for n, i in enumerate(test_batches_list):
i_s = i * TEST_BATCH_SIZE # index of the first image in this batch
i_e = min([(i + 1) * TEST_BATCH_SIZE, test_data_num]) # index of the last image in this batch
# read batch images and remove training mean
test_images = hdf5_file["test_img"][i_s:i_e, ...]
test_images = test_images.reshape(test_images.shape[0], test_images.shape[3],
test_images.shape[1], test_images.shape[2]) # reshaped to size 22046, 3, 64, 64
# read labels and convert to one hot encoding
test_labels = hdf5_file["test_labels"][i_s:i_e]
test_labels_one_hot = np.eye(n_values)[test_labels]
temp = net.test(test_images / 255., test_labels_one_hot)
num_of_correct_guesses += temp
num_of_correct_guesses_10 += temp
if n % 10 == 9: # print every 10 batches
print('Batch: {}, Accuracy: {}'.format(n+1,num_of_correct_guesses_10 * 100 / float(TEST_BATCH_SIZE * 10)))
num_of_correct_guesses_10 = 0
print('Total test Accuracy : {}'.format(num_of_correct_guesses / float(test_data_num)))
# -
# ## Visual Test Results of LeNet Network Architecture with SGD by using 64x64x3 images
# Below part is for picking a random 16 images, you can run it again to see the test results with different images.
plt.figure(1 , figsize = (8 , 8))
n = 0
mask = []
for i in range(16):
n += 1
r = np.random.randint(0 , hdf5_file["train_img"].shape[0] , 1)
mask.append(r[0])
plt.subplot(4 , 4 , n)
plt.subplots_adjust(hspace = 0.5 , wspace = 0.5)
plt.imshow(hdf5_file["train_img"][r[0]]/255.)
plt.title('{}'.format(train_labels[r[0]]))
plt.xticks([]) , plt.yticks([])
plt.show()
predictions = []
for sample_idx in mask:
sample = hdf5_file["train_img"][sample_idx]/ 255.
sample = sample.reshape(sample.shape[2], sample.shape[0], sample.shape[1])
label = hdf5_file["train_labels"][sample_idx]
predicted_label = net.predict(sample)
if predicted_label == 0:
predictions.append("healthy")
else:
predictions.append("malaria")
# ## Predictions
plt.figure(1 , figsize = (8 , 8))
n = 0
for i, sample_idx in enumerate(mask):
n += 1
plt.subplot(4 , 4 , n)
plt.subplots_adjust(hspace = 0.5 , wspace = 0.5)
plt.imshow(hdf5_file["train_img"][sample_idx]/255.)
plt.title('{}'.format(predictions[i]))
plt.xticks([]) , plt.yticks([])
plt.show()
# ## LeNet by using the receptive field property
# +
net = dl.Network()
# receptive field LeNet
output_size_conv1, num_of_outputs_conv1 = net.conv(hdf5_file['train_img'][0].shape[0],
hdf5_file['train_img'][0].shape[2],
3, # filter size
3, # filter depth
8, # number of filters
1, # stride
0) # padding
output_size_relu1_h, output_size_relu1_w, num_of_outputs_relu1 = net.relu(
num_of_outputs_conv1, # number of the feature maps
output_size_conv1, #height of one of the feature map
output_size_conv1 # width of one of the feature map
)
output_size_conv2, num_of_outputs_conv2 = net.conv(
output_size_relu1_h,
num_of_outputs_relu1,
3, # filter size
8, # filter dept
8, # number of filters
1, #stride
0 # padding
)
output_size_relu2_h, output_size_relu2_w, num_of_outputs_relu2 = net.relu(
num_of_outputs_conv2, # number of the feature maps
output_size_conv2, #height of one of the feature map
output_size_conv2 # width of one of the feature map
)
output_size_pooling2, num_of_outputs_pooling_2 = net.maxpool(
output_size_relu2_h,
num_of_outputs_relu2,
2, # filter size
2, # stride
0 # padding
)
output_size_conv3, num_of_outputs_conv3 = net.conv(
output_size_pooling2,
num_of_outputs_pooling_2,
3, # filter size
8, # filter dept
16, # number of filters
1, #stride
0 # padding
)
output_size_relu3_h, output_size_relu3_w, num_of_outputs_relu3 = net.relu(
num_of_outputs_conv3, # number of the feature maps
output_size_conv3, #height of one of the feature map
output_size_conv3 # width of one of the feature map
)
output_size_conv4, num_of_outputs_conv4 = net.conv(
output_size_relu3_h,
num_of_outputs_relu3,
3, # filter size
16, # filter dept
16, # number of filters
1, #stride
0 # padding
)
output_size_relu4_h, output_size_relu4_w, num_of_outputs_relu4 = net.relu(
num_of_outputs_conv4, # number of the feature maps
output_size_conv4, #height of one of the feature map
output_size_conv4 # width of one of the feature map
)
output_size_pooling4, num_of_outputs_pooling_4 = net.maxpool(
output_size_relu4_h,
num_of_outputs_relu4,
2, # filter size
2, # stride
0 # padding
)
output_size_fully1_h, output_size_fully1_w, num_of_outputs_fully1 = net.fullyConnected(
output_size_pooling4,
output_size_pooling4,
num_of_outputs_pooling_4,
32
)
output_size_relu5_h, output_size_relu5_w, num_of_outputs_relu5 = net.relu(
num_of_outputs_fully1, # number of the feature maps
output_size_fully1_h, #height of one of the feature map,
output_size_fully1_w
)
output_size_fully2_h, output_size_fully2_w, num_of_outputs_fully2 = net.fullyConnected(
output_size_relu5_h,
output_size_relu5_w,
num_of_outputs_relu5,
2
)
output_size_sigmoid5_h, output_size_sigmoid5_w, num_of_outputs_sigmoid5 = net.softmax(
num_of_outputs_fully2, # number of the feature maps
output_size_fully2_h, #height of one of the feature map,
output_size_fully2_w
)
# -
# ## Testing of the LeNet with Receptive Field Property with SGD by Using 28x28x3 images
# +
net.loadWeights("../Malaria_Dataset/training_sgd_receptive_28/training_sgd_receptive_28.txt")
n_values = 2
test_data_num = hdf5_file["test_labels"].shape[0]
#####################################################
# Since we don't want to load all the data at once, #
# we need to divide it into batches. From the below #
# line you can change the batch size for the path. #
#####################################################
TEST_BATCH_SIZE = 10
test_batches_list = list(range(int(ceil(float(test_data_num) / TEST_BATCH_SIZE))))
shuffle(test_batches_list)
num_of_correct_guesses = 0
num_of_correct_guesses_10 = 0
for n, i in enumerate(test_batches_list):
i_s = i * TEST_BATCH_SIZE # index of the first image in this batch
i_e = min([(i + 1) * TEST_BATCH_SIZE, test_data_num]) # index of the last image in this batch
# read batch images and remove training mean
test_images = hdf5_file["test_img"][i_s:i_e, ...]
test_images = test_images.reshape(test_images.shape[0], test_images.shape[3],
test_images.shape[1], test_images.shape[2]) # reshaped to size 22046, 3, 64, 64
# read labels and convert to one hot encoding
test_labels = hdf5_file["test_labels"][i_s:i_e]
test_labels_one_hot = np.eye(n_values)[test_labels]
temp = net.test(test_images / 255., test_labels_one_hot)
num_of_correct_guesses += temp
num_of_correct_guesses_10 += temp
if n % 10 == 9: # print every 10 batches
print('Batch: {}, Accuracy: {}'.format(n+1,num_of_correct_guesses_10 * 100 / float(TEST_BATCH_SIZE * 10)))
num_of_correct_guesses_10 = 0
print('Total test Accuracy : {}'.format(num_of_correct_guesses / float(test_data_num)))
# -
# ## Visual Test Results of LeNet Network Architecture with Receptive Field Properties by using SGD
predictions = []
for sample_idx in mask:
sample = hdf5_file["train_img"][sample_idx]/ 255.
sample = sample.reshape(sample.shape[2], sample.shape[0], sample.shape[1])
label = hdf5_file["train_labels"][sample_idx]
predicted_label = net.predict(sample)
if predicted_label == 0:
predictions.append("healthy")
else:
predictions.append("malaria")
# ## Predictions
plt.figure(1 , figsize = (8 , 8))
n = 0
for i, sample_idx in enumerate(mask):
n += 1
plt.subplot(4 , 4 , n)
plt.subplots_adjust(hspace = 0.5 , wspace = 0.5)
plt.imshow(hdf5_file["train_img"][sample_idx]/255.)
plt.title('{}'.format(predictions[i]))
plt.xticks([]) , plt.yticks([])
plt.show()
# ## Testing of the LeNet with Receptive Field Property with Momentum by Using 28x28x3 images
# +
net.loadWeights("../Malaria_Dataset/training_momentum_receptive_net_28/training_2recptive_28_epoch_3.txt")
n_values = 2
test_data_num = hdf5_file["test_labels"].shape[0]
#####################################################
# Since we don't want to load all the data at once, #
# we need to divide it into batches. From the below #
# line you can change the batch size for the path. #
#####################################################
TEST_BATCH_SIZE = 10
test_batches_list = list(range(int(ceil(float(test_data_num) / TEST_BATCH_SIZE))))
shuffle(test_batches_list)
num_of_correct_guesses = 0
num_of_correct_guesses_10 = 0
for n, i in enumerate(test_batches_list):
i_s = i * TEST_BATCH_SIZE # index of the first image in this batch
i_e = min([(i + 1) * TEST_BATCH_SIZE, test_data_num]) # index of the last image in this batch
# read batch images and remove training mean
test_images = hdf5_file["test_img"][i_s:i_e, ...]
test_images = test_images.reshape(test_images.shape[0], test_images.shape[3],
test_images.shape[1], test_images.shape[2]) # reshaped to size 22046, 3, 64, 64
# read labels and convert to one hot encoding
test_labels = hdf5_file["test_labels"][i_s:i_e]
test_labels_one_hot = np.eye(n_values)[test_labels]
temp = net.test(test_images / 255., test_labels_one_hot)
num_of_correct_guesses += temp
num_of_correct_guesses_10 += temp
if n % 10 == 9: # print every 10 batches
print('Batch: {}, Accuracy: {}'.format(n+1,num_of_correct_guesses_10 * 100 / float(TEST_BATCH_SIZE * 10)))
num_of_correct_guesses_10 = 0
print('Total test Accuracy : {}'.format(num_of_correct_guesses / float(test_data_num)))
# -
# ## Visual Test Results of LeNet Network Architecture with Receptive Field Properties by using Momentum
predictions = []
for sample_idx in mask:
sample = hdf5_file["train_img"][sample_idx]/ 255.
sample = sample.reshape(sample.shape[2], sample.shape[0], sample.shape[1])
label = hdf5_file["train_labels"][sample_idx]
predicted_label = net.predict(sample)
if predicted_label == 0:
predictions.append("healthy")
else:
predictions.append("malaria")
# ## Predictions
plt.figure(1 , figsize = (8 , 8))
n = 0
for i, sample_idx in enumerate(mask):
n += 1
plt.subplot(4 , 4 , n)
plt.subplots_adjust(hspace = 0.5 , wspace = 0.5)
plt.imshow(hdf5_file["train_img"][sample_idx]/255.)
plt.title('{}'.format(predictions[i]))
plt.xticks([]) , plt.yticks([])
plt.show()
| src/MalariaDetection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from datetime import timedelta
# -
# !pip install squarify
import squarify
df = pd.read_csv('D_sample_data.csv', encoding = 'ISO-8859-1')
df.head()
# # RFM
df.info()
df['InvoiceDate'] = pd.to_datetime(df['InvoiceDate'])
df.info()
df.shape
min(df['InvoiceDate'])
max(df['InvoiceDate'])
df = df.dropna()
len(df)
df['total_price'] = df['Quantity'] * df['UnitPrice']
reference_date = max(df['InvoiceDate']) + timedelta(days=1)
reference_date
df_processed = df.groupby(['CustomerID']).agg({'InvoiceDate': lambda x: (reference_date - x.max()).days,
'InvoiceNo': 'count',
'total_price': 'sum'})
df_processed
df_processed.columns = ['Recency', 'Frequency', 'Monetary']
df_processed.head()
# +
import seaborn as sns
sns.distplot(df_processed['Recency'])
# -
sns.distplot(df_processed['Frequency'])
sns.distplot(df_processed['Monetary'])
r = range(4, 0, -1)
f = range(1, 5)
m = range(1, 5)
r
for i in f:
print(i)
r_g = pd.qcut(df_processed['Recency'], q=4, labels=r)
f_g = pd.qcut(df_processed['Frequency'], q=4, labels=f)
m_g = pd.qcut(df_processed['Monetary'], q=4, labels=m)
final_df = df_processed.assign(R = r_g.values, F = f_g.values, M = m_g.values)
final_df
def combine_rfm(x): return str(x['R']) + str(x['F']) + str(x['M'])
final_df['combined'] = final_df.apply(combine_rfm, axis=1)
rfm = final_df
rfm.head()
rfm['sum_val'] = rfm[['R', 'F', 'M']].sum(axis=1)
rfm
rfm['combined'].nunique()
def rfm_level(df):
if df['sum_val'] >= 9:
return 'Can\'t Loose Them'
elif ((df['sum_val'] >= 8) and (df['sum_val'] < 9)):
return 'Champions'
elif ((df['sum_val'] >= 7) and (df['sum_val'] < 8)):
return 'Loyal'
elif ((df['sum_val'] >= 6) and (df['sum_val'] < 7)):
return 'Potential'
elif ((df['sum_val'] >= 5) and (df['sum_val'] < 6)):
return 'Promising'
elif ((df['sum_val'] >= 4) and (df['sum_val'] < 5)):
return 'Needs Attention'
else:
return 'Require Activation'
rfm['rfm_level'] = rfm.apply(rfm_level, axis=1)
rfm
rfm_level_agg = rfm.groupby('rfm_level').agg({
'Recency': 'mean',
'Frequency': 'mean',
'Monetary': ['mean', 'count']
}).round(1)
rfm_level_agg
rfm_level_agg.columns = rfm_level_agg.columns.droplevel()
rfm_level_agg.columns = ['Recency_Mean', 'Frequency_Mean', 'Monetary_Mean', 'Count']
rfm_level_agg
fig = plt.gcf()
ax = fig.add_subplot()
fig.set_size_inches(16, 9)
squarify.plot(sizes=rfm_level_agg['Count'],
label=['Can\'t Loose Them',
'Champions',
'Loyal',
'Needs Attention',
'Potential',
'Promising',
'Require Activation'], alpha=.6 )
plt.title('RFM Segments')
plt.axis('off')
plt.show()
| RFM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data Collection, Pre-Processing & Visualization using Python
#
# #### Author: <NAME>. | Email Id: <EMAIL>
# ## Introduction
#
# <b>The objective of this project is to collect a dataset from one or more open web APIs and use Python to pre-process and analyse the collected data. In this project, we collect, pre-process and analyze the data related to movies. The data is extracted using multiple web-APIs, and pre-processed and analyzed using the python pandas framework.</b>
# Import the relevant packages required
from IPython.display import display
import csv, json, os, time
import urllib.request
import pandas as pd
import config
# ## [1.] Selection of Relevant Web-APIs
# __We have used two web-APIs in this project.__
# <b>(a) TMDb API - https://developers.themoviedb.org/3/getting-started/introduction<br>
# - This is the official API of the The Movie Database (TMDb). For the freely available version, an API key is required, which is provided upon signup and a small amount of rate limiting is enforced. The current limits are 40 requests every 10 seconds and are limited by IP address, not API key. The only format supported by this API is JSON.</b>
# <b>(b) OMDb API - http://www.omdbapi.com/<br>
# - Although unofficial, the OMDb (The Open Movie Database) API is a RESTful web service to obtain movie information from IMDb. An API key is provided after signup, to fetch the movie data and it supports both XML and JSON formats. In our case, we have used the JSON format. For the free version, the rate limit is 1000 requests per day.</b>
# +
# Form the request URLs to fetch the movie data from TMDb
tmdb_latest_movie_url = "https://api.themoviedb.org/3/movie/popular?api_key="+config.tmdb_api_key+"&language=en-US&page="
tmdb_get_imdb_id_url_head = "https://api.themoviedb.org/3/movie/"
tmdb_get_imdb_id_url_tail = "/external_ids?api_key="+config.tmdb_api_key
# Form the request URL to fetch the movie data from OMDb
omdb_get_movie_details_url_head = "http://www.omdbapi.com/?i="
omdb_get_movie_details_url_tail = "&apikey="+config.omdb_api_key
# -
# __Now that we have chosen the respective web-APIs, we will use them to extract the details of different movies to form our dataset for analysis.__
# ## [2.] Data Collection using Web-APIs
# <b>First, define a generic function that takes in a URL (API URL request) as an argument, sends the request to the database and retrieves the data. As mentioned earlier, in our project, all the APIs return the data in JSON format. This function returns the JSON object containing the relevant data.</b>
# Function to fetch the data in a JSON format,using the url passed to it
def getApiURLJsonData(url):
response = urllib.request.urlopen(url)
raw_json = response.read().decode('utf-8')
movie_json = json.loads(raw_json)
return movie_json
# <b>The TMDb API does not return the IMDb ID of the movies, along with the other details. The IMDb ID of the corresponding movie has to be retrieved using a different end-point of the same API. A function is defined below, that returns the IMDb ID of the corresponding movie, by accepting the TMDb ID of the movie as a parameter.</b>
# Function to retrieve the IMDb ID of a movie
def getIMDbMovieID(tmdb_id):
tmdb_get_imdb_id_url=tmdb_get_imdb_id_url_head+str(tmdb_id)+tmdb_get_imdb_id_url_tail
tmdb_external_id_json = getApiURLJsonData(tmdb_get_imdb_id_url)
imdb_id = tmdb_external_id_json['imdb_id']
return imdb_id
# <b>The below function is to get the ID of the movie in IMDb using the TMDb ID of the movie passed as first parameter, and then use this ID with the OMDb API to retrieve the values for the list of parameters passed as the second argument. It returns a list of values corresponding to the parameters.</b>
# Function to get all the required details of a movie from IMDb.
def getIMDbMovieDetails(tmdb_id, imdbParamList):
imdb_details_list = []
# Retrieve the id of the movie in IMDb
imdb_id = getIMDbMovieID(tmdb_id)
if(imdb_id is None):
# If no IMDb ID is obtained (or it is of None type), then set 'N/A' as the default value for all the parameters.
for param in imdbParamList:
imdb_details_list.append("N/A")
else:
# Form the URL, fetch the details of the movie from IMDb and append them to the list.
omdb_get_movie_details_url = omdb_get_movie_details_url_head+imdb_id+omdb_get_movie_details_url_tail
imdb_details_json = getApiURLJsonData(omdb_get_movie_details_url)
if(imdb_details_json['Response']=="True"):
for param in imdbParamList:
imdb_details_list.append(imdb_details_json[param])
else:
for param in imdbParamList:
imdb_details_list.append("N/A")
return imdb_details_list
# <b>For every movie, certain details associated with it are fetched from TMDb (using TMDb API) and certain other details from IMDb (using OMDb API). The following function is to fetch the details of a movie from the TMDb and also invoke the other functions to retrieve the corresponding details from IMDb.</b>
# Function to fetch all the relevant movie details from TMDb API.
def getMovieParamData(movie_data, movieParamList, imdbParamList):
movieDataList=[]
for param in movieParamList:
if(param=='id'):
# If the parameter is id, then use it to get the IMDb ID and other details from IMDB.
movieDataList.append(movie_data[param])
imdb_details_list = getIMDbMovieDetails(movie_data[param], imdbParamList)
movieDataList[2:1] = imdb_details_list
else:
# Append all values to a list
movieDataList.append(movie_data[param])
return movieDataList
# <b>We have now completed the process of collecting the relevant data from multiple web-APIs (in JSON format). The next step is parsing them and saving them to a file for subsequent analysis.</b>
# ### [3.] Data Parsing & Storage in CSV Format
# <b>Define a function to create a CSV file with the required headers. This function takes in a two arguments, the name of the file and the headers to be used in it. Since, we use multiple APIs, the data needs to be combined and placed at a common source, which is the CSV file. Note that we would have two CSV files generated as part of the project, one as the main source of data and the other, is used to get an entry point to the possible future enhancements of this project.</b>
# Function to create a CSV file with the relevant headers.
def create_file (file_name, headerList):
# Create (open) the file. The parameter newline = '', in order to prevent blank lines being written.
with open(file_name, "w", newline='') as write_file:
f = csv.writer(write_file)
# Write the headers to the file
f.writerow(headerList)
write_file.close()
# <b>Create a function to keep appending the final set of movie data identified for our analysis, to the CSV file. The function takes the file name and the list of movie data to be written to the CSV file.</b>
# Function to append the required movie details to the CSV file.
def writeMovieData(fileName, movieDataList):
# Open the CSV file. The parameter newline = '', in order to prevent blank lines being written.
with open(fileName, "a", newline='', encoding='utf-8') as write_file:
f = csv.writer(write_file)
# Write the row of movie details identified for analysis
f.writerow(movieDataList)
write_file.close()
# <b>In the below section, we first define the name of the CSV file and invoke the function to create the same. Next, we define two separate lists of parameters to be retrieved through the TMDb and OMDb APIs, respectively. Finally, we define the number of pages to be retrieved from TMDb results (using the URL and TMDb API), iterate through each of them and form the list comprising of all values related to a particular movie. This list is then written as a single row to the CSV file.<br>At the moment we are collecting just <i>40 movie records</i>. To increase the amount of data retreieved, we may change the <i>'pageLimit'</i> parameter accordingly.</b>
# +
# The final CSV file with the combined data from all APIs
fileName="Extracted_Movie_Data.csv"
# Create the CSV file
headerList = ['MovieTitle', 'Id', 'Production', 'Director', 'Runtime', 'Genre', 'Revenue', 'Rated', 'Language', 'ReleaseDate', 'Popularity', 'VoteCount', 'VoteAverage', 'Adult']
create_file(fileName, headerList)
# The list of keys in the resulting JSON file, to be retrieved from the TMDb
movieParamList = ['original_title', 'id', 'release_date', 'popularity', 'vote_count', 'vote_average', 'adult']
# The list of keys in the resulting JSON file, to be retrieved from the IMDb
imdbParamList = ['Production', 'Director', 'Runtime', 'Genre', 'BoxOffice', 'Rated', 'Language']
# Set the no. of result-pages to be retrieved through the TMDb API
pageLimit = 3
for pageNum in range(1, pageLimit):
# Form the URL string by passing the api_key and the page number
tmdb_latest_movie_url_new = tmdb_latest_movie_url+str(pageNum)
# Get the details in JSON format
movie_json = getApiURLJsonData(tmdb_latest_movie_url_new)
# Iterate through the JSON object and fetch all the relevant movie details
for movie_data in movie_json['results']:
movieDataList=getMovieParamData(movie_data, movieParamList, imdbParamList)
# Write the final list of values as one row to the CSV file
writeMovieData(fileName, movieDataList)
time.sleep(0.300)
# -
# <b>Here, we complete the task of parsing the data collected from different APIs and storing them at a single source(a CSV file). We now proceed to loading this data to a dataframe usind pandas and pre-processing them subsequently.</b>
# ### [4.] Loading Data to DataFrame & Applying Pre-Processing
# ### [4.1.] Loading the Data into DataFrame
# <b>Using pandas, the movie data is loaded into a dataframe, directly from the CSV file we generated before. The number of rows and columns in the dataframe, along with a few rows is displayed on the console (using 'shape').</b>
# +
# Create the dataframe by loading the data from the CSV file, formed earlier.
movieDetailsDataFrame = pd.read_csv(fileName, index_col='Id')
# Get the count of rows & columns in the dataframe, and display them
totalRowCount = movieDetailsDataFrame.shape[0]
totalColCount = movieDetailsDataFrame.shape[1]
# Display the no. of rows and columns in the dataframe
display("Shape of DataFrame:"+str(movieDetailsDataFrame.shape))
# Display first 3 elements of the dataframe to verify the columns and their values
movieDetailsDataFrame.head(5)
# -
# ### [4.2.] Data Pre-Processing
# #### [4.2.1.] Drop the Columns that Add No Value
# <b>Now that we have our dataframe, we must pre-process and normalize its contents to extract the best possible insights. In our first step, we take a look at all the columns and figure out which of them would actualy be useful in our analysis.</b>
# <b>We can see from our dataframe that all the values in the column <i>'Adult'</i> are <i>'False'</i>. As such, this column is not useful in our analysis. Hence, we can drop it from the dataframe. To confirm this, we first take a count of the no. of rows in the dataframe, where the value in <i>'Adult'</i> column is <i>'False'</i> and then, compare it with the total rowcount of the dataframe. If both are the same, then we drop the column.</b>
# +
# Get the count of rows in the dataframe where the value in 'Adult' column is 'False'
adultFalseValRowCount = movieDetailsDataFrame.loc[movieDetailsDataFrame['Adult'] == False]['Adult'].count()
if totalRowCount==adultFalseValRowCount:
# Drop the 'Adult' column from the dataframe since all the values in the column are the same
movieDetailsDataFrame = movieDetailsDataFrame.drop(['Adult'], axis=1)
# Display first few elements of the dataframe to verify the columns and their values
movieDetailsDataFrame.head(3)
# -
# <b>We can see above that the column <i>'Adult'</i>, no longer exists in the dataframe.</b>
# #### [4.2.2.] Handle Missing Values
# <b>In the second step, we figure out all the null or empty values in our dataframe and replace them accordingly.</b>
# Get the different columns and the no. of missing values in them.
movieDetailsDataFrame.isnull().sum()
# <b>We can observe that, the columns <i>'Production', 'Director', 'Runtime', 'Genre', 'Revenue', 'Rated' and 'Language'</i> contain missing values.<br> - For the columns, <i>'Production', 'Director', 'Genre'</i>, we cannot set any calculated values, as these are all of type 'string'. Hence, we set a default value <i>'Unknown'</i> to all the columns.<br> - For the column <i>'Rated'</i>, we replace the missing values with one of its own values <i>'NOT RATED'</i>, whcih matches the situation.<br>- For the column <i>'Language'</i>, we replace the missing values with the default value <i>'English'</i>.<br> - For the columns <i>'Runtime' and 'Revenue'</i>, we set the values <i>'0 min' and '$0'</i>, respectively. <i>But these two values are temporary and would be replaced in further steps.</i><br>In the below section we fix them as mentioned.</b>
# +
# Replace all missing values in the columns 'Production', 'Director', 'Genre' & 'Rated'
movieDetailsDataFrame["Production"] = movieDetailsDataFrame["Production"].fillna("Unknown")
movieDetailsDataFrame["Director"] = movieDetailsDataFrame["Director"].fillna("Unknown")
movieDetailsDataFrame["Genre"] = movieDetailsDataFrame["Genre"].fillna("Unknown")
movieDetailsDataFrame["Rated"] = movieDetailsDataFrame["Rated"].fillna("NOT RATED")
# Replace all missing values in the column 'Language'
movieDetailsDataFrame["Language"] = movieDetailsDataFrame["Language"].fillna("English")
# Replace all missing values in the column 'Runtime' (temporary replacement)
movieDetailsDataFrame["Runtime"] = movieDetailsDataFrame["Runtime"].fillna("0 min")
# Replace all missing values in the column 'Revenue' (temporary replacement)
movieDetailsDataFrame["Revenue"] = movieDetailsDataFrame["Revenue"].fillna("$0")
# -
# #### [4.2.3.] Convert Column Types
# <b>Now let us convert the types of certain columns that are currently of type 'string'.<br> - The column <i>'ReleaseDate'</i>, needs to be converted from string to datetime, as it contains date values.<br> - Similarly, the columns <i>'Runtime' and 'Revenue'</i> need to be converted from string to int and float, respectively.<br>We will also rename the columns <i>'Runtime' & 'Revenue'</i> accordingly, so that it is easily readable and understandable.</b>
# +
# Convert the type of the column 'ReleaseDate' from string to datetime
movieDetailsDataFrame['ReleaseDate'] = pd.to_datetime(movieDetailsDataFrame['ReleaseDate'])
# Convert the type of the columns 'Runtime' & 'Revenue' from string to numeric
movieDetailsDataFrame['Runtime'] = movieDetailsDataFrame['Runtime'].replace('[ min]', '', regex=True)
movieDetailsDataFrame['Runtime'] = pd.to_numeric(movieDetailsDataFrame['Runtime'])
movieDetailsDataFrame['Revenue'] = movieDetailsDataFrame['Revenue'].replace('[\$,]', '', regex=True)
movieDetailsDataFrame['Revenue'] = pd.to_numeric(movieDetailsDataFrame['Revenue'])
# Rename the columns 'Runtime' & 'Revenue', for easy readability
movieDetailsDataFrame = movieDetailsDataFrame.rename(columns={'Revenue': 'Revenue($)', 'Runtime': 'Runtime(mins)'})
# -
# <b>Now that we have converted the types of the columns <i>'Runtime' and 'Revenue'</i> and also renamed them, let us complete the pending task of replacing the missing values in these columns. Recall that earlier, we had set temporary values in these columns.<br>Since, these are numeric fields, let us calculate the mean of these two columns and then set the same as the default value for both these columns.</b>
# +
# Calculate the mean of columns 'Runtime' & 'Revenue' and round them off accordingly.
mean_runtime = round(movieDetailsDataFrame['Runtime(mins)'].mean(), 2)
mean_revenue = round(movieDetailsDataFrame['Revenue($)'].mean(), 3)
# Now replace the temporary values set earlier, in these columns
movieDetailsDataFrame["Runtime(mins)"] = movieDetailsDataFrame["Runtime(mins)"].replace(0, mean_runtime)
movieDetailsDataFrame["Revenue($)"] = movieDetailsDataFrame["Revenue($)"].replace(0, mean_revenue)
# -
# #### [4.2.4.] Normalize/Scale the Numeric Values
# <b>All the numeric values in the dataframe must be normalized or brought to the same range, for effective results to be obtained as a result of the analysis.</b>
# <b>We can see that, all the values in the columns <i>'Runtime(mins)', 'Revenue($)', 'Popularity', 'VoteCount' and 'VoteAverage'</i> are not in the same range. Hence, we apply <i>min-max normalization</i> on the values in all these columns, except <i>'VoteAverage'</i>. To do this, we take the minimum and maximum values in each of these columns and, then apply the formula <i>(val-min)/(max-min)</i> to normalize the values.<br>In case of the column <i>'VoteAverage'</i>, we just divide each of its values by 10, since it is already in the range of 0-10.</b>
# +
# Normalize the values of the columns 'Runtime(mins)', 'Revenue($), 'Popularity' & 'VoteCount' using min-max normalization
movieDetailsDataFrame['Runtime(mins)'] = (movieDetailsDataFrame['Runtime(mins)'] - movieDetailsDataFrame['Runtime(mins)'].min())/(movieDetailsDataFrame['Runtime(mins)'].max() - movieDetailsDataFrame['Runtime(mins)'].min())
movieDetailsDataFrame['Revenue($)'] = (movieDetailsDataFrame['Revenue($)'] - movieDetailsDataFrame['Revenue($)'].min())/(movieDetailsDataFrame['Revenue($)'].max() - movieDetailsDataFrame['Revenue($)'].min())
movieDetailsDataFrame['Popularity'] = (movieDetailsDataFrame['Popularity'] - movieDetailsDataFrame['Popularity'].min())/(movieDetailsDataFrame['Popularity'].max() - movieDetailsDataFrame['Popularity'].min())
movieDetailsDataFrame['VoteCount'] = (movieDetailsDataFrame['VoteCount'] - movieDetailsDataFrame['VoteCount'].min())/(movieDetailsDataFrame['VoteCount'].max() - movieDetailsDataFrame['VoteCount'].min())
# Scale the values of the column 'VoteAverage' to bring it to the range(0,1)
movieDetailsDataFrame['VoteAverage'] = movieDetailsDataFrame['VoteAverage']/10
# -
# #### [4.2.5.] Create Any New Columns Required
# <b>Once pre-processing is done, we will create two new columns <i>'ReleaseMonth'</i> and <i>'ReleaseYear'</i> from the existing <i>'ReleaseDate'</i> column, which will further help our analysis using plots.<b>
# +
# Import the datetime package
from datetime import datetime as dt
# Create two new columns in the dataframe to represent the Month & Year of Release, respectively.
movieDetailsDataFrame['ReleaseMonth'] = movieDetailsDataFrame['ReleaseDate'].dt.strftime('%b')
movieDetailsDataFrame['ReleaseYear'] = movieDetailsDataFrame['ReleaseDate'].dt.strftime('%Y')
# Display first 3 elements of the dataframe to verify the columns and their values
movieDetailsDataFrame[0:3]
# -
# <b>With this, we completed loading the data from CSV into dataframe and data pre-processing. We dropped the columns that do not add any value to the analysis, handled/replaced missing values, converted column types and normalized/scaled the numeric values.</b>
# ### [5.] Summarizing the Analysis
# <b>We now summarize our analysis using different plots and graphical representations. All the interpretations, insights and conclusions drawn from this dataset would be brought to the table, in this section.</b>
# #### [5.1.] No. of Movies Released Per Month
# <b>Our objective here, is to plot a bar graph that displays the number of movies released per month, irrespective of the year.</b>
# +
# Define a list containing the names/abbreviations of months in a year
uniqueMonthList = ['Jan', 'Feb', 'Mar', 'Apr','May','Jun', 'Jul', 'Aug','Sep', 'Oct', 'Nov', 'Dec']
# Create a new smaller dataframe with the 'ReleaseMonth' and the no. of movies released per month
movieReleaseMonthDataFrame = pd.DataFrame(movieDetailsDataFrame['ReleaseMonth'].value_counts())
movieReleaseMonthDataFrame.index = pd.CategoricalIndex(movieReleaseMonthDataFrame.index, categories=uniqueMonthList, sorted=True)
movieReleaseMonthDataFrame = movieReleaseMonthDataFrame.sort_index()
# +
# Import the required libraries to plot the graphs
import matplotlib, pylab
import matplotlib.pyplot as plt
# %matplotlib inline
# Set the corresponding positions on the y-axis
y_pos = range(len(movieReleaseMonthDataFrame.index.values))
# Create a new figure & set the canvas size
plt.figure(figsize=(9,4))
# Plot the bar chart
bars = plt.bar(y_pos, movieReleaseMonthDataFrame['ReleaseMonth'], align='center', color='green')
plt.xticks(y_pos, movieReleaseMonthDataFrame.index.values)
plt.xlabel("Month of Release", fontsize=16)
plt.ylabel("No. of Movies Released", fontsize=16)
plt.title("No. of Movies Released Per Month", fontsize=18)
plt.show()
# -
# <b>We can see that most number of movies are released in <i>_July_</i>. Similarly, we can also create graphs for determining the year, day of month or day of week, when maximum number of movies were released.</b>
# #### [5.2.] Distribution of 'Runtime' Feature across the Dataset
# <b>This histogram plotted below, portrays the distribution of the attribute <i>'Runtime(mins)'</i> using its frequencies across our dataset.</b>
obj = movieDetailsDataFrame["Runtime(mins)"].plot(kind="hist", title="Distribution of Runtime across the Dataset", bins=8, color='black')
# #### [5.3.] Relationship between Popularity & Revenue
# <b>Here, we try to understand if there is a <i>directly proportional</i> relationship between the popularity of a movie among the audience and its box-office collection.</b>
# +
# Create a new smaller dataframe containing the 'ReleaseDate', 'ReleaseMonth', 'ReleaseYear', 'Popularity' & 'Revenue($)'
moviePopularityRevenueDf = movieDetailsDataFrame[['Popularity', 'Revenue($)']]
# Sort the dataframe based on the column 'Popularity'
moviePopularityRevenueDf = moviePopularityRevenueDf.sort_values(by='Popularity', ascending=False)[0:50]
# Plot a line graph with the values
plt.figure(figsize=(8,4))
p = plt.scatter(moviePopularityRevenueDf['Revenue($)'], moviePopularityRevenueDf['Popularity'], color="blue", s=65)
# Set the labels for both the axes and also set the title of the graph
plt.xlabel('Movie Revenue', fontsize=16)
plt.ylabel('Movie Popularity', fontsize=16)
plt.title("Relationship between a Movie's Popularity & Revenue", fontsize=18)
# -
# <b>We can infer from the plotted graph that the movies with huge popularity may not necessarily be the ones with the most revenue at the box-office. Hence, we can conclude that there is no <i>directly proportional</i> relationship between the poularity and revenue of movies.</b>
# #### [5.4.] Variations in Revenue and Vote Average across Months
# <b>We try to figure out how the box office collections and the average of audience votes for a movie, vary across the different months of a year.</b>
# +
# Create a smaller dataframe containing the mean revenue and vote average, grouped by the month of release
yearly_Group = movieDetailsDataFrame[['ReleaseMonth', 'Revenue($)', 'VoteAverage']].groupby('ReleaseMonth').agg('mean')
yearly_Group.index = pd.CategoricalIndex(yearly_Group.index, categories=uniqueMonthList, sorted=True)
yearly_Group = yearly_Group.sort_index()
# Plot a dual-axis graph representing how the mean revenue and vote average change with months
revVotePlt = yearly_Group.plot(secondary_y=['VoteAverage'])
revVotePlt.set_ylabel("Movie Revenue ($)", fontsize=16)
revVotePlt.set_xlabel("Release Month of Movie", fontsize=16)
revVotePlt.right_ax.set_ylabel("Average of Votes", fontsize=16)
plt.title("Revenue and Vote Average of Movies over Months of a Year\n", fontsize=18)
plt.show()
# -
# <b>From the dual-axis graph generated, we can observe that there are significant variations in the values of revenue and votes for the movies. <i>The overall trends seem to be similar, except for the fact that some of the movies released towards the end of an year, have very high revenue.</i></b>
# <b>The above graph can be split into sub-plots as shown below.</b>
yearly_Group.plot(subplots=True, figsize=(5, 5))
# #### [5.5.] Revenue Distribution across Months, for every Year of Release
# <b>We try to figure out the box office collections of each month in each year of release, for all the movies. This is represented using a heatmap, as shown below.</b>
# +
import seaborn as sns
from matplotlib.colors import ListedColormap
# Create a copy of the main dataframe that contains only the columns 'ReleaseMonth', 'ReleaseYear','Revenue($)'
monthYearGroupDf = movieDetailsDataFrame[['ReleaseMonth', 'ReleaseYear','Revenue($)']].copy()
# Convert the columns 'ReleaseMonth' & 'ReleaseYear' to categorical variables
monthYearGroupDf['ReleaseMonth'] = pd.CategoricalIndex(monthYearGroupDf['ReleaseMonth'], categories=uniqueMonthList, sorted=True)
monthYearGroupDf['ReleaseYear'] = monthYearGroupDf['ReleaseYear'].astype('category')
# Plot a heatmap using seaborn
fig,ax = plt.subplots(figsize=(12,12))
my_cmap = ListedColormap(sns.cubehelix_palette(8, start=2, rot=0, dark=0, light=.95))
sns.heatmap(monthYearGroupDf.pivot_table(index='ReleaseYear', columns='ReleaseMonth', values='Revenue($)'),
cmap=my_cmap, ax=ax, linewidth=0.5)
# -
# <b>It can be inferred that the highest revenue was obtained for the movies released in <i>April, 2012 and December, 2017</i>.</b>
# #### [5.6.] Popular Movies & their Directors, with Certification
# <b>Similar to the above plot, the below heatmap represents the popularity of movies directed by a director along with its certification.</b>
# +
# Create a copy of the main dataframe that contains only the columns 'ReleaseMonth', 'ReleaseYear','Revenue($)'
directorGroupDf = movieDetailsDataFrame[['Rated', 'Director', 'Popularity']].copy()
directorGroupDf = directorGroupDf.sort_values(by='Popularity', ascending=False)[0:10]
# Convert the columns 'ReleaseMonth' & 'ReleaseYear' to categorical variables
for ctg in ['Rated', 'Director']:
directorGroupDf[ctg] = directorGroupDf[ctg].astype('category')
my_cmap = ListedColormap(sns.cubehelix_palette(8, start=.5, rot=-.75))
fig,ax = plt.subplots(figsize=(12,12))
sns.heatmap(directorGroupDf.pivot_table(index='Director', columns='Rated', values='Popularity'),
cmap=my_cmap, ax=ax, linewidth=0.5, annot=True)
# -
# <b>From the top 10 popular movies we selected from the dataframe, we see that the highest popularity was obtained for the movie directed by <i>_J.A.Bayona_</i> which was certified/rated as <i>PG-13</i></b>.
# #### [5.7.] Connecting the Different Features in the Dataset
# <b>The below representation involves plotting our attributes <i>'Popularity', 'VoteAverage', 'Revenue($)' and 'Runtime(mins)'</i> on a different column and then connecting them using lines for each row (data point) in the dataframe. This is done to understand the relationship between these attributes and a movie's rating/certification.</b>
# +
# Copy the columns 'Popularity', 'VoteAverage', 'Revenue($)', 'Runtime(mins)' & 'Rated', to a new dataframe
tempDataFrame = movieDetailsDataFrame[['Popularity', 'VoteAverage', 'Revenue($)', 'Runtime(mins)', 'Rated']].copy()
# Plot the values using parallel co-ordinates
from pandas.plotting import parallel_coordinates
parCord = parallel_coordinates(tempDataFrame, "Rated")
# -
# <b>The lines in the below figure are coloured based on the values in the <i>'Rated'</i> column. We understand that there is a high convariance between these features.</b>
# #### [5.8.] No. of Movies Per Rating (Certification)
# <b>The below representation involves plotting each of our attributes on a different column and then connecting them using lines for each row (data point) in the dataframe.</b>
# +
ratedDataFrame = movieDetailsDataFrame[['MovieTitle', 'Rated']].copy()
ratedDataFrame = ratedDataFrame.groupby('Rated').count()
p = ratedDataFrame["MovieTitle"].plot(title="Number Of Movies Per Rating", figsize=(8,4), color='red')
# -
# <b>The lines in the below figure are coloured based on the values in the <i>'Rated'</i> column. We see that most no. of movies in our dataset are rated <i>'PG-13'</i>.</b>
# ## [6.] Conclusion
#
# <b>In the above steps, we used multiple web-APIs to retrieve the data related to movies, parsed and combined them to a single CSV file, pre-processed the data and finally, created some visualizations based on our analysis.</b>
# <b>
# Through a series of steps, we were able to analyze and understand the following:
# - In which month of a year, the most number of movies are released.
# - Identify the pattern of distribution of the runtime of a movie across our dataset.
# - Nature of relationship between the Popularity and Revenue of a movie.
# - How the Revenue and Average of Votes, vary across different months in a year.
# - Identify the pattern of Revenue Distribution across different months, for every Year of Release.
# - The top-10 Directors and Ratings for their movies, based on the Popularity of the movies.
# - How the different attributes of our dataset are connected to each other, based on the Ratings(certifications) of the movies.
# - The number of movies per Rating/Certification.
# </b>
# <b>The point to note here is that, these conclusions are tentative and may be vary (or) remain same/smiliar when analyzed on huge datasets. At the moment, these results are based on 40 popular movies collected using multiple APIs.</b>
# ## [7.] References
# <b>
# 1. https://developers.themoviedb.org/3
# 2. http://www.omdbapi.com/
# 3. https://pandas.pydata.org/pandas-docs/stable/
# 4. https://matplotlib.org/faq/usage_faq.html
# 5. https://seaborn.pydata.org/tutorial/color_palettes.html
# 6. https://seaborn.pydata.org/generated/seaborn.heatmap.html
# 7. COMP41680 - Lecture Slides, by Prof. <NAME>
# 8. COMP41680 - Lab Tutorial Notebooks, by Prof. <NAME> </b>
| Data_PreProcessing_Visualization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:.conda-oof] *
# language: python
# name: conda-env-.conda-oof-py
# ---
import pubchempy as pcp
import numpy as np
import pandas as pd
import xlrd
import ast
import csv
import opentrons
import datetime
import importlib
import setup_functions
importlib.reload(setup_functions)
# +
component_df = pd.read_excel('chemical_inventory.xlsx')
component_df
stock_spec_filepath = 'stock_specification.csv'
stock_dict = setup_functions.get_stock_dfs(stock_spec_filepath)
stock_dict['hba-stock']
# -
experiment_plan_filepath = 'experiment_plan.csv'
experiment_plan_dict = setup_functions.get_experiment_plan(experiment_plan_filepath)
experiment_plan_dict
conc_df = setup_functions.generate_candidate_samples(experiment_plan_dict)
# +
def cull_candidate_samples(experiment_plan_dict, stock_dict):
"""
Attempt to take care of components appearing in more than one set.
Pretty, but difficult to execute.
"""
start_df = setup_functions.generate_candidate_samples(experiment_plan_dict)
all_component_names_set = set()
# Generate set of all unique names
for k,v in stock_dict.items():
all_component_names_set.update(v['Chemical Abbreviation'].values)
print(all_component_names_set,'\n')
# Specify which componennt is unique to each stock.
for k,v in stock_dict.items():
name_set = set(v['Chemical Abbreviation'].values)
not_in_set = all_component_names_set-name_set
# print(name_set)
# print(not_in_set)
# print("\n")
def cull_candidate_samples(experiment_plan_dict, stock_dict):
start_df = setup_functions.generate_candidate_samples(experiment_plan_dict)
for stock_name in experiment_plan_dict['Stock order']:
stock_df = stock_dict[stock_name]
print(stock_name)
print(stock_df['Chemical Abbreviation'].values)
print('\n')
cull_candidate_samples(experiment_plan_dict, stock_dict)
# -
| code-collab/database/.ipynb_checkpoints/database-interactions-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.8 64-bit (''base'': conda)'
# name: python3
# ---
from Integrais import Integrais as it
from EDO import EDO
from sympy import var, Lambda, cos, sin, tan, exp, log, ln, sqrt, solve
x = var('x')
y = var('y')
# Questão 01
x0 = 0
y0 = 1
dy = Lambda((x, y), x*y**2 - y)
print('f(x, y) = ', dy(x,y))
I = (0, 2)
N = 8
Y = EDO(dy, y0, x0)
y1, y_steps, x_steps = Y.euler(I, N, dp=6)
print(f'Solução Aproximada: y(2) = {y1}')
# Questão 02
x0 = 0
y0 = 0
dy = Lambda((x, y), 2*x + y)
print('f(x, y) = ', dy(x,y))
I = (0, 0.6)
N = 3
Y = EDO(dy, y0, x0)
y1, y_steps, x_steps = Y.runge_kutta2(I, N, dp=7)
print(f'Solução Aproximada: y(0.6) = {y1}')
# +
# Questão 3
f = Lambda(x, sqrt(4 + x**2))
a = 0
b = 3
display(f)
Q1 = it(f, a, b)
Q1.simpson3_8(n=6)
# +
# Questão 4
f = Lambda(x, x**3 * exp(2*x))
a = 0
b = 2
display(f)
Q1 = it(f, a, b)
Q1.simpson1_3(n=8)
| MAT 271/P3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # for using range
# # for using iteration variable
l1 = [1, 3, 4, 7, 3, 9, 3]
# for using range (using index)
for i in range(len(l1)): # index
print('index:', i, 'value', l1[i])
# iteration variable
for number in l1:
print(number)
# finite loop
i = 5
while i >= 1:
print(i)
i -= 1
print('done')
# finite loop
i = 0
while i <= 5:
print(i)
i += 1
print('done')
| 01_revision/2019/loops/loop101.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Booleans
# +
# Let's declare some bools
spam = True
print spam
print type(spam)
eggs = False
print eggs
print type(eggs)
# -
# ### Python truth value testing
# - Any object can be tested for truth value
# - Truth value testing is used in flow control or in Boolean operations
# - All objects are evaluated as True except:
# - None (aka. null)
# - False
# - Zero of any numeric type: 0, 0L, 0.0, 0j, 0x0, 00
# - Any empty sequence or mapping: '', [], (), {}
# - Instances of user-defined classes implementing __nonzero__ or __len__ method
# and returning 0 or False
# +
# Let's try boolean operations
print True or True
print True or False
print False or True # Boolean or. Short-circuited, so it only evaluates the second argument if the first one is False
# -
print True and True
print True and False
print False and True # Boolean or. Short-circuited, so it only evaluates the second argument if the first one is True
print not True
print not False
# +
# So, if all objects can be tested for truth, let's try something different
spam = [1.2345, 7, "x"]
eggs = ("a", 0x07, True)
fooo = "aeiou"
# -
print spam or eggs
# Did you expect it to print True?
print fooo or []
print "" or eggs
print spam and eggs
print fooo and ()
print [] and eggs
print not spam
print not ""
print spam and eggs or "abcd" and False
print (spam and eggs) or ("abcd" and False)
print spam and (eggs or "abcd") and False
print spam and (eggs or "abcd" and False)
# ### Python boolean operands:
# - ALWAYS return one of the incoming arguments!
# - x or y => if x is false, then y, else x
# - x and y => if x is false, then x, else y
# - not x => if x is false, then True, else False
# - They are short-circuited, so second argument is not always evaluated
# - Can take any object type as arguments
# - Even function calls, so boolean operands are used for flow control
# - Parentheses may be used to change order of boolean operands or comparissons
#
# ### What about comparisons?
# +
spam = 2
eggs = 2.5
print spam == 2 # equal
print spam != eggs # not equal
# +
print spam >= eggs # greater than or equal
print spam > eggs # strictly greater than
print spam <= eggs # less than or equal
print spam < eggs # strictly less than
# +
print spam is 2 # object identity, useful to compare with None (discussed latter)
print spam is not None # negated object identity
# -
# # Flow Control
# ### Let's start with the conditional execution
# +
spam = [1, 2, 3] # True
eggs = "" # False
if spam:
print "spam is True"
else:
print "spam is False"
print "outside the conditional" # Notice that theres is no closing fi statement
# -
if spam:
print "spam is True"
else:
print "spam is False"
print "still inside the conditional"
# ### REMEMBER:
# - Indentation is Python's way of grouping statements!!
# - Typically four spaces per indentation level
# - No curly brackets { } or semicolons ; used anywhere
# - This enforces a more readable code
if eggs:
print "eggs is True"
elif spam:
print "eggs is False and spam is True"
else:
print "eggs and spam are False"
if eggs:
print "eggs is True"
elif max(spam) > 5:
print "eggs is False and second condition is True"
elif len(spam) == 3 and not eggs is None:
print "third condition is true"
else:
print "everything is False"
# ### Let's see the ternary operator
# +
spam = [1, 2, 3] # True
eggs = "" # False
print "first option" if spam else "second option"
# -
print "first option" if eggs else "second option"
print "first option" if eggs else "second option" if spam else "last option" # We can even concatenate them
print "first option" if eggs else ("second option" if spam else "last option")
# ### Time for the while loop
spam = [1, 2, 3]
while len(spam) > 0:
print spam.pop(0)
spam = [1, 2, 3]
idx = 0
while idx < len(spam):
print spam[idx]
idx += 1
# ### What about the for loop?
spam = [1, 2, 3]
for item in spam: # The for loop only iterates over the items of a sequence
print item
spam = [1, 2, 3]
for item in spam[::-1]: # As we saw, slicing may be slow. Keep it in mind
print item
eggs = "eggs"
for letter in eggs: # It can loop over characters of a string
print letter
spam = {"one": 1,
"two": 2,
"three": 3}
for key in spam: # Or even it can interate through a dictionary
print spam[key] # Note that it iterates over the keys of the dictionary
# ### Let's see how to interact with loops iterations
spam = [1, 2, 3]
for item in spam:
if item == 2:
break
print item
# - **break** statement halts a loop execution (inside while or for)
# - Only affects the closer inner (or smallest enclosing) loop
# A bit more complicated example
spam = ["one", "two", "three"]
for item in spam: # This loop is never broken
for letter in item:
if letter in "wh": # Check if letter is either 'w' or 'h'
break # Break only the immediate inner loop
print letter
print # It prints a break line (empty line)
# A bit different example
spam = ["one", "two", "three"]
for item in spam:
for letter in item:
if letter in "whe": # Check if letter is either 'w', 'h' or 'e'
continue # Halt only current iteration, but continue the loop
print letter
print
# - **continue** statement halts current iteration (inside while or for)
# - loops continue its normal execution
spam = [1, 2, 3, 4, 5, 6, 7, 8]
eggs = 5
while len(spam) > 0:
value = spam.pop()
if value == eggs:
print "Value found:", value
break
else: # Note that else has the same indentation than while
print "The right value was not found"
spam = [1, 2, 3, 4, 6, 7, 8]
eggs = 5
while len(spam) > 0:
value = spam.pop()
if value == eggs:
print "Value found:", value
break
else:
print "The right value was not found"
# **else** clause after a loop is executed if all iterations were run without break statement called
spam = [1, 2, 3]
for item in spam:
pass
# **pass** statement is Python's noop (does nothing)
# ### Let's check exception handling
spam = [1, 2, 3]
try:
print spam[5]
except: # Use try and except to capture exceptions
print "Failed"
spam = {"one": 1, "two": 2, "three": 3}
try:
print spam[5]
except IndexError as e: # Inside the except clause 'e' will contain the exception instance
print "IndexError", e
except KeyError as e: # Use several except clauses for different types of exceptions
print "KeyError", e
try:
print 65 + "spam"
except (IndexError, KeyError) as e: # Or even group exception types
print "Index or Key Error", e
except TypeError as e:
print "TypeError", e
try:
print 65 + 2
except (IndexError, KeyError), e:
print "Index or Key Error", e
except TypeError, e:
print "TypeError", e
else:
print "No exception" # Use else clause to run code in case no exception was raised
try:
print 65 + "spam"
raise AttributeError # Use 'raise' to launch yourself exceptions
except (IndexError, KeyError), e:
print "Index or Key Error", e
except TypeError, e:
print "TypeError", e
else:
print "No exception"
finally:
print "Finally we clean up" # Use finally clause to ALWAYS execute clean up code
try:
print 65 + 2
except (IndexError, KeyError), e:
print "Index or Key Error", e
raise # Use 'raise' without arguments to relaunch the exception
except TypeError, e:
print "TypeError", e
else:
print "No exception"
finally:
print "Finally we clean up" # Use finally clause to ALWAYS execute clean up code
# ### Let's see another construction
try:
f = open("tmp_file.txt", "a")
except:
print "Exception opening file"
else:
try:
f.write("I'm writing to a file...\n")
except:
print "Can not write to a file"
finally:
f.close()
# ### Not pythonic, too much code for only three real lines
try:
with open("tmp_file.txt", "a") as f:
f.write("I'm writing to a file...\n")
except:
print "Can not open file for writing"
# #### Where is the file closed? What happens if an exception is raised?
#
# ### Python context managers
# - Encapsulate common patterns used wrapping code blocks where real runs the program logic
# - Usually try/except/finally patterns
# - Several uses:
# - Automatic cleanup, closing files or network or DB connections when exiting the context block
# - Set temporary environment, like enable/disable logging, timing, profiling...
# - Use the 'with' and optionally the 'as' statements to open a context manager
# - It is automatically closed when code execution goes outside the block
# # Comprehension
# +
spam = [0, 1, 2, 3, 4]
eggs = [0, 10, 20, 30]
fooo = []
for s in spam:
for e in eggs:
if s > 1 and e > 1:
fooo.append(s * e)
print fooo
# -
# ### Short code, right?
spam = [0, 1, 2, 3, 4]
eggs = [0, 10, 20, 30]
fooo = [s * e for s in spam for e in eggs if s > 1 and e > 1]
print fooo
# ### What about now?
fooo = [s * s for s in spam] # This is the most basic list comprehension construction
print fooo
fooo = [s * s for s in spam if s > 1] # We can add 'if' clauses
print fooo
spam = [1, 2, 3, 4]
eggs = [0, -1, -2, -3]
fooo = [l.upper() * (s + e) for s in spam
for e in eggs
for l in "SpaM aNd eGgs aNd stuFf"
if (s + e) >= 1
if l.islower()
if ord(l) % 2 == 0] # We can add lots of 'for' and 'if' clauses
print fooo
spam = [1, 2, 3, 4]
eggs = [10, 20, 30, 40]
fooo = [[s * e for s in spam] for e in eggs] # It is possible to nest list comprehensions
print fooo
# - List comprehension is faster than standard loops (low level C optimizations)
# - However, built-in functions are still faster (see Functional and iterables tools module)
# ### There is also dict comprehension (2.7 or higher)
spam = ['monday', 'tuesday',
'wednesday', 'thursday',
'friday']
fooo = {s: len(s) for s in spam} # The syntax is a merge of list comprehension and dicts
print fooo
spam = [(0, 'monday'), (1, 'tuesday'),
(2, 'wednesday'), (3, 'thursday'),
(4, 'friday')]
fooo = {s: idx for idx, s in spam} # Tuple unpacking is useful here
print fooo
spam = ['monday', 'tuesday',
'wednesday', 'thursday',
'friday']
fooo = {s: len(s) for s in spam if s[0] in "tm"} # Ofc, you can add more 'for' and 'if' clauses
print fooo
# # Sources
# - http://docs.python.org/2/library/stdtypes.html#boolean-operations-and-or-not
# - http://docs.python.org/2/tutorial/controlflow.html#if-statements
# - http://docs.python.org/2/reference/compound_stmts.html
# - http://docs.python.org/2/reference/expressions.html#conditional-expressions
# - http://docs.python.org/2/reference/simple_stmts.html
# - http://www.python.org/dev/peps/pep-0343/
# - http://docs.python.org/2/reference/compound_stmts.html#the-with-statement
# - http://docs.python.org/2/tutorial/classes.html#iterators
# - http://docs.python.org/2/library/stdtypes.html#iterator-types
# - http://docs.python.org/2/tutorial/datastructures.html#list-comprehensions
# - http://www.python.org/dev/peps/pep-0274/
| basic/3_Booleans_Flow_Control_and_Comprehension.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
### multi_variable_linear_regression_01 코드의 data 플롯을 보여준다
(prediction)이 코드의 목표다.
import tensorflow as tf
import numpy as np
from matplotlib import pyplot as plt
# 한글 폰트를 설정해야 matplotlib에서 한글이 정상 출력이 된다.
from matplotlib.pyplot import rc
rc('font',family='New Gulim')
# +
x1_data = [73., 93., 89., 96., 73.]
x2_data = [80., 88., 91., 98., 66.]
x3_data = [75., 93., 90., 100., 70.]
y_data = [152., 185., 180., 196., 142.]
# -
plt.plot(x1_data, y_data, 'ro')
plt.plot(x2_data, y_data, 'go')
plt.plot(x3_data, y_data, 'bo')
plt.show()
# #### loss 함수에 해당하는 plot을 보면 gradient descent를 어떻게 해야 할 지 좀 더 잘 알게 될 것이다.
# 이런 류의 loss 함수를 MSE(Mean Square Error), LS(Least Square), RMS(Root Mean Square) 등으로 취급한다.
# 이 용어만 알아둬도 수식을 이해 할 때 아주 좋다.
# # loss
# loss = tf.reduce_sum(tf.square(y_hat - y)) # sum of the squares
#
# #### W, b의 초기값으로 임의로 설정한 상태의 loss 값을 관찰해보자
# 실제 학습시에는 대개 W는 0~1 사이의 normal distribution의 값을 가져오고, b는 0으로 설정한다.
# Weight initilization 문서 참고
W = 2.4
b = -8.0
y_hat = np.multiply(W, y_groundtruth) + b
plt.plot(x_train, y_hat, 'ro')
plt.plot(x_train, y_hat, 'r-')
plt.plot(x_train, y_groundtruth, 'bo')
plt.plot(x_train, y_groundtruth, 'b-')
plt.title('y_groundtruth : 파란색, y_hat : 빨강색')
plt.xlabel('x')
plt.ylabel('y')
plt.show()
# #### cost 값으로만 plot을 그려보자.
# W = 1.0, b = 0.0 이라고 정답을 알려준 상태에서의 W 변화에 따른 loss 값의 추세를 관찰해 보자.
# gradient descent를 하면 될 것 같지 않은가?
# +
W = 1.0
b = 0.0
number_of_W_range = 40
W_range = np.linspace(W - 2.0, W + 2.0, num=number_of_W_range)
cost_list = np.zeros([number_of_W_range], dtype=np.float32)
for i, W_value in enumerate(W_range):
y_hat = np.multiply(W_value, y_groundtruth) + b
cost_list[i] = np.sum(np.square(y_hat - y_groundtruth))
# -
plt.plot(W_range, cost_list, 'b-')
plt.plot(W_range, cost_list, 'ro')
plt.title('정답 근처에서의 cost 값 변화를 통한 Gradient descent 필요성 파악(b = 0)')
plt.xlabel('W')
plt.ylabel('cost')
plt.show()
| code/multi_variable_linear_regression_00_plot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Reading CSVs
import pandas as pd
# ## Reading the entire CSV
df = pd.read_csv("data/titantic-train.csv")
df.shape
# ## Reading in batches
for batch in pd.read_csv("data/titantic-train.csv", chunksize=100):
print(batch.shape)
| Reading CSVs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Transformada de Fourier de alguns sinais
#
# Neste notebook avaliaremos a transformada de Fourier de alguns sinais. A TF é definida por
#
# \begin{equation}
# X(\mathrm{j} \omega)=\int^{\infty}_{-\infty}x(t)e^{-\mathrm{j}\omega t}\mathrm{d}t
# \tag{1}
# \end{equation}
#
# E a TF inversa por:
#
# \begin{equation}
# x(t)=\frac{1}{2 \pi}\int^{\infty}_{-\infty}X(\mathrm{j} \omega)e^{\mathrm{j}\omega t}\mathrm{d}\omega
# \tag{2}
# \end{equation}
# importar as bibliotecas necessárias
import numpy as np # arrays
import matplotlib.pyplot as plt # plots
plt.rcParams.update({'font.size': 14})
# ### Exemplo 1
#
# $x(t) = \delta(t)$
#
# neste caso
#
# $X(\mathrm{j} \omega)=1$
#
# +
freq = np.linspace(-50, 50, 1000)
w = 2*np.pi*freq
Xjw = np.ones(len(w))
plt.figure(figsize=(15, 5))
plt.subplot(1,2,1)
plt.plot(freq, np.abs(Xjw), '-b', linewidth = 2)
plt.grid(linestyle = '--', which='both')
plt.xlabel('Frequência [Hz]')
plt.ylabel(r'$|X(\mathrm{j}\omega)|$ [-]')
plt.xticks(np.arange(freq[0], freq[-1]+10, 10))
plt.xlim((freq[0], freq[-1]))
plt.subplot(1,2,2)
plt.plot(freq, np.angle(Xjw), '-b', linewidth = 2)
plt.grid(linestyle = '--', which='both')
plt.xlabel('Frequência [Hz]')
plt.ylabel(r'$\angle X(\mathrm{j}\omega)$ [rad]')
plt.xticks(np.arange(freq[0], freq[-1]+10, 10))
plt.xlim((freq[0], freq[-1]))
plt.tight_layout()
plt.show()
# -
# ### Exemplo 2
#
# $x(t) = \mathrm{e}^{-at}u(t)$, $a>0$
#
# neste caso
#
# $X(\mathrm{j} \omega)=\frac{1}{a+\mathrm{j}\omega}$
# +
time = np.linspace(-1, 1, 1000)
xt = np.zeros(len(time))
a = 500
xt[time>=0] = np.exp(-a*time[time>=0])
# Time domain
plt.figure(figsize=(20, 2.5))
plt.plot(time, xt, '-b', linewidth = 2)
plt.grid(linestyle = '--', which='both')
plt.xlabel('Tempo [s]')
plt.ylabel(r'$x(t)$ [-]')
plt.xticks(np.arange(time[0], time[-1]+0.1, 0.1))
plt.xlim((time[0], time[-1]))
# Freq domain
freq = np.linspace(-5, 5, 10000)
w = 2*np.pi*freq
Xjw = 1/(a+1j*w)
plt.figure(figsize=(15, 5))
plt.subplot(1,2,1)
plt.plot(freq, np.abs(Xjw)/np.amax(np.abs(Xjw)), '-b', linewidth = 2)
plt.grid(linestyle = '--', which='both')
plt.xlabel('Frequência [Hz]')
plt.ylabel(r'$|X(\mathrm{j}\omega)|$ [-]')
plt.xticks(np.arange(freq[0], freq[-1]+1, 1))
plt.xlim((freq[0], freq[-1]))
plt.ylim((0, 1.2))
plt.subplot(1,2,2)
plt.plot(freq, np.angle(Xjw), '-b', linewidth = 2)
plt.grid(linestyle = '--', which='both')
plt.xlabel('Frequência [Hz]')
plt.ylabel(r'$\angle X(\mathrm{j}\omega)$ [rad]')
plt.xticks(np.arange(freq[0], freq[-1]+1, 1))
plt.xlim((freq[0], freq[-1]))
plt.tight_layout()
plt.show()
# -
# # Exercício
# +
time = np.linspace(-5, 5, 1000)
xt = np.ones(len(time))
Tp = 5
xt[time<=-Tp/2] = 0
xt[time>=Tp/2] = 0
# Time domain
plt.figure(figsize=(20, 2.5))
plt.plot(time, xt, '-b', linewidth = 2)
plt.grid(linestyle = '--', which='both')
plt.xlabel('Tempo [s]')
plt.ylabel(r'$x(t)$ [-]')
plt.xticks(np.arange(time[0], time[-1]+1, 1))
plt.xlim((time[0], time[-1]))
# -
# # Dualidade
# +
time = np.linspace(-5, 5, 1000)
xt1 = np.ones(len(time))
Tp = 2
xt1[time<=-Tp/2] = 0
xt1[time>=Tp/2] = 0
freq = np.linspace(-5, 5, 1000)
w=2*np.pi*freq
Xw1 = 2*np.sin(w*Tp/2)/(w*Tp/2)
w0 = 5
xt2 = 2*np.sin((w0/2)*time)/((w0/2)*time) #np.sinc(time)
Xw2 = np.ones(len(freq))
Xw2[freq<=-w0/2] = 0
Xw2[freq>=w0/2] = 0
plt.figure(figsize=(15, 10))
plt.subplot(2,2,1)
plt.plot(time, xt1, '-b', linewidth = 2)
plt.grid(linestyle = '--', which='both')
plt.xlabel('Tempo [s]')
plt.ylabel(r'$x(t)$ [-]')
plt.xticks(np.arange(time[0], time[-1]+1, 1))
plt.xlim((time[0], time[-1]))
plt.subplot(2,2,2)
plt.plot(freq, np.abs(Xw1)/len(Xw1), '-b', linewidth = 2)
plt.grid(linestyle = '--', which='both')
plt.xlabel('Frequência [Hz]')
plt.ylabel(r'$|X(\mathrm{j}\omega)|$ [-]')
plt.xticks(np.arange(freq[0], freq[-1]+1, 1))
plt.xlim((freq[0], freq[-1]))
plt.subplot(2,2,3)
plt.plot(time, xt2, '-b', linewidth = 2)
plt.grid(linestyle = '--', which='both')
plt.xlabel('Tempo [s]')
plt.ylabel(r'$x(t)$ [-]')
plt.xticks(np.arange(time[0], time[-1]+1, 1))
plt.xlim((time[0], time[-1]))
plt.subplot(2,2,4)
plt.plot(freq, np.abs(Xw2), '-b', linewidth = 2)
plt.grid(linestyle = '--', which='both')
plt.xlabel('Frequência [Hz]')
plt.ylabel(r'$|X(\mathrm{j}\omega)|$ [-]')
plt.xticks(np.arange(freq[0], freq[-1]+1, 1))
plt.xlim((freq[0], freq[-1]))
plt.tight_layout()
# +
time = np.linspace(-5, 5, 1000)
xt1 = np.ones(len(time))
Tp = 2
xt1[time<=-Tp/2] = 0
xt1[time>=Tp/2] = 0
freq = np.linspace(-5000, 5000, 10000)
w=2*np.pi*freq
Xw1 = 2*np.sin(w*Tp/2)/(w*Tp/2)
w0 = 1000
xt2 = 2*np.sin((w0/2)*time)/((w0/2)*time) #np.sinc(time)
Xw2 = np.ones(len(freq))
Xw2[freq<=-w0/2] = 0
Xw2[freq>=w0/2] = 0
plt.figure(figsize=(15, 5))
plt.subplot(1,2,1)
plt.plot(time, xt2, '-b', linewidth = 2)
plt.grid(linestyle = '--', which='both')
plt.xlabel('Tempo [s]')
plt.ylabel(r'$x(t)$ [-]')
plt.xticks(np.arange(time[0], time[-1]+1, 1))
plt.xlim((time[0], time[-1]))
plt.subplot(1,2,2)
plt.plot(freq, np.abs(Xw2), '-b', linewidth = 2)
plt.grid(linestyle = '--', which='both')
plt.xlabel('Frequência [Hz]')
plt.ylabel(r'$|X(\mathrm{j}\omega)|$ [-]')
plt.xticks(np.arange(freq[0], freq[-1]+1, 1))
plt.xlim((freq[0], freq[-1]))
plt.tight_layout()
| Aula 24 - TF de alguns sinais/TF alguns espectros.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Experiment Template
# ## Starting a new experiment
# * Activate virtualenv by `workon pamogk` or create one with `mkvirtualenv pamogk`
# * Run `jupyter notebook notebooks/template.ipynb`
# * Inside the notebook click `Cell > Run All` (only the first time)
# * After it finishes click `File > Make a Copy...`
# * Restart jupyter server (only the first time)
# * You can remove anything except for first 2 initialization cells, they do magic stuff such as installing requirements or setting the project root for imports
# ## Importing/Using project libraries
# First two cells do most of the job so you should be able to import them easily e.g:
# ```python
# import config
# print('Data dir:', config.data_dir)
# ```
#
# If you want to reload changed modules you can do
# ```python
# # %load_ext autoreload
# # %autoreload 2
# ```
# See [here](https://ipython.org/ipython-doc/3/config/extensions/autoreload.html) for more information on reloading modules.
#
# ## Extensions/Widgets
# Jupyter has nice extensions/widgets to make it easier to work with it. Again first 2 cells should enable some useful
# extensions for you. See [here](https://jupyter-contrib-nbextensions.readthedocs.io/en/latest/) for more information on them.
# + init_cell=true
# # %Template%
# detect virtual env
import sys
import os
# install needed libs and extensions manually
user_param = '--user'
if hasattr(sys, 'real_prefix') or (hasattr(sys, 'base_prefix') and sys.base_prefix != sys.prefix):
user_param = ''
# !pip install -q $user_param jupyter_contrib_nbextensions ipywidgets yapf rootpath
# !jupyter contrib nbextension install --InstallContribNbextensionsApp.log_level=ERROR --user
# !jupyter nbextension enable --py widgetsnbextension
# append project root to python path
import rootpath
_, ROOT_PATH = rootpath.append()
# change current directory to root
os.chdir(ROOT_PATH)
# !pip install -q -r requirements.txt
print('Initialization completed!')
# + init_cell=true
extensions = {
'notebook': ['contrib_nbextensions_help_item/main', 'scroll_down/main', 'code_prettify/code_prettify', 'splitcell/splitcell', 'execute_time/ExecuteTime', 'gist_it/main', 'python-markdown/main', 'init_cell/main', 'hinterland/hinterland', 'snippets/main', 'toc2/main', 'varInspector/main', 'jupyter-js-widgets/extension', 'move_selected_cells/main', 'snippets_menu/main'],
'edit': ['codefolding/edit'],
'tree': ['nbextensions_configurator/tree_tab/main'],
}
for sec, exts in extensions.items():
for ext in exts:
# !jupyter nbextension enable $ext --section=$sec --user --InstallContribNbextensionsApp.log_level=ERROR
# -
import config
print('Python path sanity check:', config.data_dir)
# +
# example plotly plot
import plotly.express as px
import numpy as np
import pandas as pd
data = pd.DataFrame(np.random.randn(100, 2), columns=list('xy'))
px.scatter(data, x='x', y='y', title='Random Scatter Plot Example').show()
# -
data['z'] = data['x'] - data['y']
px.scatter(data, x='x', y='y', color='z', title='Random Scatter Plot Example with Color').show()
# + [markdown] cell_style="center"
# You can check for some useful tips and tricks at (add more links as you see fit):
# * [this medium post](https://towardsdatascience.com/bringing-the-best-out-of-jupyter-notebooks-for-data-science-f0871519ca29)
| notebooks/template.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Importing Libraries
import itertools
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.ticker import NullFormatter
import pandas as pd
import numpy as np
import matplotlib.ticker as ticker
from sklearn import preprocessing
# %matplotlib inline
# ## Importing the Dataset
df = pd.read_csv('teleCust1000t.csv')
df.head()
df['custcat'].value_counts()
df.hist(column='income', bins=50)
y = df['custcat'].values
y[0:5]
X = preprocessing.StandardScaler().fit(X).transform(X.astype(float))
X[0:5]
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split( X, y, test_size=0.2, random_state=4)
print ('Train set:', X_train.shape, y_train.shape)
print ('Test set:', X_test.shape, y_test.shape)
from sklearn.neighbors import KNeighborsClassifier
k = 4
#Train Model and Predict
neigh = KNeighborsClassifier(n_neighbors = k).fit(X_train,y_train)
neigh
yhat = neigh.predict(X_test)
yhat[0:5]
from sklearn import metrics
print("Train set Accuracy: ", metrics.accuracy_score(y_train, neigh.predict(X_train)))
print("Test set Accuracy: ", metrics.accuracy_score(y_test, yhat))
| Supervised Learning/Classification/KNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introducing Scikit-Learn
# ### Basics of the API
#
# The steps of one Scikit-Learn estimator:
#
# 1. Choose a model and import it (The models are class).
# 2. Make an instance and choose the hyperparameters.
# 3. Arrange data into a features matrix and target vector.
# 4. Fit the model to your data by calling the ``fit()`` method of the model instance.
# 5. Apply the Model to new data:
# - For supervised learning, often we predict labels for unknown data using the ``predict()`` method.
# - For unsupervised learning, we often transform or infer properties of the data using the ``transform()`` or ``predict()`` method.
#
#
class Estimator(object):
def fit(self, X, y=None):
"""Fits estimator to data."""
# set state of ``self``
return self
# %matplotlib inline
import numpy as np
import pylab as plt
import pandas as pd
from sklearn import svm
from sklearn.metrics import classification_report,confusion_matrix,accuracy_score
np.random.seed(0)
X = np.r_[np.random.randn(20, 2) - [2, 2], np.random.randn(20, 2) + [2, 2]]
Y = [0] * 20 + [1] * 20
plt.scatter(X[:, 0], X[:, 1], c=Y, cmap=plt.cm.coolwarm, s=25)
# +
fig, ax = plt.subplots()
clf2 = svm.LinearSVC(C=1).fit(X, Y)
# get the separating hyperplane
w = clf2.coef_[0]
a = -w[0] / w[1]
xx = np.linspace(-5, 5)
yy = a * xx - (clf2.intercept_[0]) / w[1]
# create a mesh to plot in
x_min, x_max = X[:, 0].min() - 1, X[:, 0].max() + 1
y_min, y_max = X[:, 1].min() - 1, X[:, 1].max() + 1
xx2, yy2 = np.meshgrid(np.arange(x_min, x_max, .2),
np.arange(y_min, y_max, .2))
Z = clf2.predict(np.c_[xx2.ravel(), yy2.ravel()])
Z = Z.reshape(xx2.shape)
ax.contourf(xx2, yy2, Z, cmap=plt.cm.coolwarm, alpha=0.3)
ax.scatter(X[:, 0], X[:, 1], c=Y, cmap=plt.cm.coolwarm, s=25)
ax.plot(xx,yy)
ax.axis([x_min, x_max,y_min, y_max])
plt.show()
# -
# **Supervised learning**:
#
# - Linear models (Ridge, Lasso, Elastic Net, ...)
# - Support Vector Machines
# - Tree-based methods (Random Forests, Bagging, GBRT, ...)
# - Nearest neighbors
# - Neural networks (basics)
# **Unsupervised learning**:
#
# - Clustering (KMeans, Ward, ...)
# - Outlier detection
# ## Accuracy and precision
#
# 1. precision:
# The fraction of relevant instances among the retrieved instances,
# 1. recall:
# The fraction of relevant instances that have been retrieved over the total amount of relevant instances.
# 1. F-score
#
#
# * The precision is the ratio tp / (tp + fp) where tp is the number of true positives and fp the number of false positives. The precision is intuitively the ability of the classifier not to label as positive a sample that is negative.
#
# * The recall is the ratio tp / (tp + fn) where tp is the number of true positives and fn the number of false negatives. The recall is intuitively the ability of the classifier to find all the positive samples.
#
# * The F-beta score can be interpreted as a weighted harmonic mean of the precision and recall, where an F-beta score reaches its best value at 1 and worst score at 0. The F-beta score weights recall more than precision by a factor of beta. beta == 1.0 means recall and precision are equally important.
#
# * The support is the number of occurrences of each class in y_true.
#
# Take a look at [HERE](https://en.wikipedia.org/wiki/F1_score) or [HERE](https://en.wikipedia.org/wiki/Precision_and_recall).
# +
# Generate data
from sklearn.datasets import make_blobs,make_circles,make_moons
X, y = make_blobs(n_samples=1000, centers=2,
cluster_std=1.5,
center_box=(-4.0, 4.0))
# X, y = make_circles(n_samples=1000, noise=.05, factor=.5)
# X,y = make_moons(n_samples=1000, noise=.05)
plt.scatter(X[:,0],X[:,1],c=y,)
# -
print(X[:5])
print(y[:5])
# +
# X is a 2 dimensional array, with 1000 rows and 2 columns
print(X.shape)
# y is a vector of 1000 elements
print(y.shape)
# -
X_train, y_train = X[:700], y[:700]
X_test, y_test = X[700:], y[700:]
# ## K-Nearest Neighbours
# +
# K-Nearest Neighbours
from sklearn.neighbors import KNeighborsClassifier
Model = KNeighborsClassifier(n_neighbors=8)
Model.fit(X_train, y_train)
y_pred = Model.predict(X_test)
# -
y_pred[:5],y_test[:5]
# +
# Summary of the predictions made by the classifier
print(classification_report(y_test, y_pred))
# print(confusion_matrix(y_test, y_pred))
# Accuracy score
print('accuracy is',accuracy_score(y_pred,y_test))
# -
# Compute (approximate) class probabilities
print(Model.predict_proba(X_test[:5]))
# ## Radius Neighbors Classifier
from sklearn.neighbors import RadiusNeighborsClassifier
Model=RadiusNeighborsClassifier(radius=8.0)
Model.fit(X_train,y_train)
y_pred=Model.predict(X_test)
#summary of the predictions made by the classifier
print(classification_report(y_test,y_pred))
# print(confusion_matrix(y_test,y_pred))
#Accouracy score
print('accuracy is ', accuracy_score(y_test,y_pred))
# ## Naive Bayes
# \begin{align}\begin{aligned}P(y \mid x_1, \dots, x_n) \propto P(y) \prod_{i=1}^{n} P(x_i \mid y)\\\Downarrow\\\hat{y} = \arg\max_y P(y) \prod_{i=1}^{n} P(x_i \mid y),\end{aligned}\end{align}
# +
# Naive Bayes
from sklearn.naive_bayes import GaussianNB
Model = GaussianNB()
Model.fit(X_train, y_train)
y_pred = Model.predict(X_test)
# Summary of the predictions made by the classifier
print(classification_report(y_test, y_pred))
# print(confusion_matrix(y_test, y_pred))
# Accuracy score
print('accuracy is',accuracy_score(y_pred,y_test))
# -
# ## SVM
# +
# Support Vector Machine
from sklearn.svm import SVC
Model = SVC(kernel='linear')
# Model = svm.LinearSVC(C=1)
Model.fit(X_train, y_train)
y_pred = Model.predict(X_test)
# Summary of the predictions made by the classifier
print(classification_report(y_test, y_pred))
# print(confusion_matrix(y_test, y_pred))
# Accuracy score
print('accuracy is',accuracy_score(y_pred,y_test))
# -
plt.scatter(X_test[:,0],X_test[:,1],c=y_pred,)
# ## Decision Tree
# +
# Decision Tree's
from sklearn.tree import DecisionTreeClassifier
Model = DecisionTreeClassifier()
Model.fit(X_train, y_train)
y_pred = Model.predict(X_test)
# Summary of the predictions made by the classifier
print(classification_report(y_test, y_pred))
print(confusion_matrix(y_test, y_pred))
# Accuracy score
print('accuracy is',accuracy_score(y_pred,y_test))
# -
# ## RandomForest
from sklearn.ensemble import RandomForestClassifier
Model= RandomForestClassifier(max_depth=2)
Model.fit(X_train,y_train)
y_pred=Model.predict(X_test)
print(classification_report(y_test,y_pred))
print(confusion_matrix(y_pred,y_test))
#Accuracy Score
print('accuracy is ',accuracy_score(y_pred,y_test))
# ## Neural network
from sklearn.neural_network import MLPClassifier
Model=MLPClassifier()
Model.fit(X_train,y_train)
y_pred=Model.predict(X_test)
# Summary of the predictions
print(classification_report(y_test,y_pred))
print(confusion_matrix(y_test,y_pred))
#Accuracy Score
print('accuracy is ',accuracy_score(y_pred,y_test))
| notebooks/15-sklearn-ML.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: auto-sklearn
# language: python
# name: auto-sklearn
# ---
# + [markdown] pycharm={}
# ## 05. Implement a Simple AutoML System
# + [markdown] pycharm={}
# ### Top Level Design of AutoML System
# + [markdown] pycharm={}
# 我们在上个教程中,学习到了UltraOpt的设计哲学是将优化器与评价器分离,如图所示:
# + pycharm={}
from graphviz import Digraph; g = Digraph()
g.node("config space", shape="ellipse"); g.node("optimizer", shape="box")
g.node("config", shape="ellipse"); g.node("loss", shape="circle"); g.node("evaluator", shape="box")
g.edge("config space", "optimizer", label="initialize"); g.edge("optimizer", "config", label="<<b>ask</b>>", color='blue')
g.edge("config","evaluator" , label="send to"); g.edge("evaluator","loss" , label="evaluate")
g.edge("config", "optimizer", label="<<b>tell</b>>", color='red'); g.edge("loss", "optimizer", label="<<b>tell</b>>", color='red')
g.graph_attr['rankdir'] = 'LR'; g
# + [markdown] pycharm={}
# 而当我们要解决AutoML问题时,我们可以这样定义AutoML系统结构:
# + [markdown] pycharm={}
# 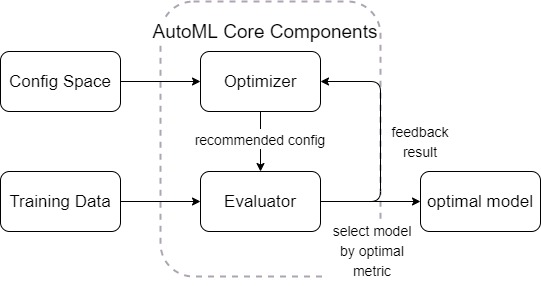
# + [markdown] pycharm={}
# ### Design a Simple AutoML Config Space by HDL
# + [markdown] pycharm={}
# 在`03. Conditional Parameter`教程中,我们知道了AutoML问题的优化可以视为一个CASH问题,不仅涉及算法选择,还涉及超参优化。我们用HDL定义一个简单的CASH问题的配置空间:
# + pycharm={}
from ultraopt.hdl import hdl2cs, plot_hdl, layering_config, plot_layered_dict
HDL = {
'classifier(choice)':{
"LinearSVC": {
"max_iter": {"_type": "int_quniform","_value": [300, 3000, 100], "_default": 600},
"penalty": {"_type": "choice", "_value": ["l1", "l2"],"_default": "l2"},
"dual": {"_type": "choice", "_value": [True, False],"_default": False},
"loss": {"_type": "choice", "_value": ["hinge", "squared_hinge"],"_default": "squared_hinge"},
"C": {"_type": "loguniform", "_value": [0.01, 10000],"_default": 1.0},
"multi_class": "ovr",
"random_state": 42,
"__forbidden": [
{"penalty": "l1","loss": "hinge"},
{"penalty": "l2","dual": False,"loss": "hinge"},
{"penalty": "l1","dual": False},
{"penalty": "l1","dual": True,"loss": "squared_hinge"},
]
},
"RandomForestClassifier": {
"n_estimators": {"_type": "int_quniform","_value": [10, 200, 10], "_default": 100},
"criterion": {"_type": "choice","_value": ["gini", "entropy"],"_default": "gini"},
"max_features": {"_type": "choice","_value": ["sqrt","log2"],"_default": "sqrt"},
"min_samples_split": {"_type": "int_uniform", "_value": [2, 20],"_default": 2},
"min_samples_leaf": {"_type": "int_uniform", "_value": [1, 20],"_default": 1},
"bootstrap": {"_type": "choice","_value": [True, False],"_default": True},
"random_state": 42
},
"KNeighborsClassifier": {
"n_neighbors": {"_type": "int_loguniform", "_value": [1,100],"_default": 3},
"weights" : {"_type": "choice", "_value": ["uniform", "distance"],"_default": "uniform"},
"p": {"_type": "choice", "_value": [1, 2],"_default": 2},
},
}
}
# + pycharm={}
CS = hdl2cs(HDL)
g = plot_hdl(HDL)
g.graph_attr['size'] = "15,8"
g
# + [markdown] pycharm={}
# ### Design a Simple AutoML Evaluator
# + [markdown] pycharm={}
# 在定义好了配置空间之后,现在我们来看评价器。
#
# 在UltraOpt的设计哲学中,评价器可以是一个类(实现`__call__`魔法方法),也可以是个函数。但其必须满足:
#
# - 接受 `dict` 类型参数 `config`
# - 返回 `float` 类型参数 `loss`
#
# 在AutoML问题中,评价器的工作流程如下:
# 1. 将config转化为一个机器学习模型
# 2. 在训练集上对机器学习模型进行训练
# 3. 在验证集上得到相应的评价指标
# 4. 对评价指标进行处理,使其`越小越好`,返回`loss`
#
# 在了解了这些知识后,我们来开发 **AutoML 评价器**
# + [markdown] pycharm={}
# 为了方便同学们理解,我们先顺序地将之前提到的**AutoML 评价器**工作流程跑一遍:
#
# #### Step 1. 将config转化为一个机器学习模型
# + pycharm={}
# 引入 sklearn 的分类器
from sklearn.svm import LinearSVC
from sklearn.ensemble import RandomForestClassifier
from sklearn.neighbors import KNeighborsClassifier
# + [markdown] pycharm={}
# 假设评价器传入了一个 `config` 参数,我们先通过获取配置空间默认值(也可以对配置空间 `CS` 对象进行采样)得到 `config`
# + pycharm={}
config = CS.get_default_configuration().get_dictionary()
config
# + [markdown] pycharm={}
# 我们用**配置分层函数** `ultraopt.hdl.layering_config` 处理这个配置
# + pycharm={}
layered_dict = layering_config(config)
layered_dict
# + pycharm={}
plot_layered_dict(layered_dict)
# + [markdown] pycharm={}
# 我们需要获取这个配置的如下信息:
#
# - 算法选择的结果
# - 被选择算法对应的参数
# + pycharm={}
AS_HP = layered_dict['classifier'].copy()
AS, HP = AS_HP.popitem()
AS # 算法选择的结果
# + pycharm={}
HP # 被选择算法对应的参数
# + [markdown] pycharm={}
# 根据 **算法选择结果** + **对应的参数** 实例化一个 **机器学习对象**
# + pycharm={}
ML_model = eval(AS)(**HP)
ML_model # 实例化的机器学习对象
# + [markdown] pycharm={}
# #### Step 2. 在训练集上对机器学习模型进行训练
# + [markdown] pycharm={}
# 我们采用MNIST手写数字数据集的一个子集来作为训练数据:
# + pycharm={}
from sklearn.datasets import load_digits
import seaborn as sns
import warnings
warnings.filterwarnings("ignore")
X, y = load_digits(return_X_y=True)
# + pycharm={}
digit = X[0].reshape([8, 8])
sns.heatmap(digit, annot=True);
# + pycharm={}
y[0]
# + [markdown] pycharm={}
# 我们需要划分一个训练集与验证集,在训练集上训练 `Step 1`得到的机器学习模型,在验证集上预测,预测值会被处理为返回的损失值 `loss` 。首先我们要对原数据进行切分以得到训练集和验证集:
# + pycharm={}
from sklearn.model_selection import train_test_split
# + pycharm={}
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.33, random_state=42)
# + [markdown] pycharm={}
# 然后我们需要在训练集`X_train`,`y_train`上对机器学习模型`ML_model`进行训练:
# + pycharm={}
ML_model.fit(X_train, y_train)
# + [markdown] pycharm={}
# #### Step 3. 在验证集上得到相应的评价指标
# + [markdown] pycharm={}
# 为了保证我们的机器学习模型具有对未知数据的泛化能力,所以需要用未参与训练的验证集来评价之前得到的机器学习模型。
#
# 所以,我们需要一个评价指标。这里我们选用最简单的 `accuracy_score`
# + pycharm={}
from sklearn.metrics import accuracy_score
# + pycharm={}
y_pred = ML_model.predict(X_test)
# + pycharm={}
accuracy = accuracy_score(y_test, y_pred)
accuracy
# + [markdown] pycharm={}
# 我们注意到本示例的 `Step 2`与`Step 3`的训练与验证部分可能存在偏差。比如如果数据集划分得不均匀,可能会在验证集上过拟合。
#
# 我们可以用**交叉验证**(Cross Validation, CV)的方法解决这个问题,交叉验证的原理如图所示:
# + [markdown] pycharm={}
# <img src="https://scikit-learn.org/stable/_images/grid_search_cross_validation.png" width=500></img>
# + pycharm={}
from sklearn.model_selection import StratifiedKFold # 采用分层抽样
from sklearn.model_selection import cross_val_score
# + [markdown] pycharm={}
# 因为这只是一个简单的示例,所以采用3-Folds交叉验证以减少计算时间。
#
# 在实践中建议使用5-Folds交叉验证或10-Folds交叉验证。
# + pycharm={}
cv = StratifiedKFold(n_splits=3, shuffle=True, random_state=0)
# + pycharm={}
scores = cross_val_score(ML_model, X, y, cv=cv)
scores
# + [markdown] pycharm={}
# 我们采用交叉验证的平均值作为最后的得分:
# + pycharm={}
score = scores.mean()
score
# + [markdown] pycharm={}
# #### Step 4. 对评价指标进行处理,使其越小越好,返回loss
# + [markdown] pycharm={}
# 我们注意到,评价指标正确率的取值范围是0-1,且是越大越好的,所以我们需要将score处理为越小越好的loss:
# + pycharm={}
loss = 1 - score
loss
# + [markdown] pycharm={}
# #### 用一个函数实现AutoML评价器
# + pycharm={}
metric = "accuracy"
def evaluate(config: dict) -> float:
layered_dict = layering_config(config)
display(plot_layered_dict(layered_dict)) # 用于在jupyter notebook中可视化,实践中可以删除次行
AS_HP = layered_dict['classifier'].copy()
AS, HP = AS_HP.popitem()
ML_model = eval(AS)(**HP)
# 注意到: X, y, cv, metric 都是函数外的变量
scores = cross_val_score(ML_model, X, y, cv=cv, scoring=metric)
score = scores.mean()
print(f"accuracy: {score:}") # 用于在jupyter notebook中调试,实践中可以删除次行
return 1 - score
# + pycharm={}
evaluate(CS.get_default_configuration())
# + [markdown] pycharm={}
# #### 用一个类实现AutoML评价器
# + [markdown] pycharm={}
# 我们注意到,`evaluate`函数的 X, y, cv, metric 都是函数外的变量,不利于管理,所以实践中我们一般采用定义**类**来实现评价器。
# + [markdown] pycharm={}
# 首先我们定义一个类,这个类需要根据 训练数据、评价指标和交叉验证方法 等条件进行初始化:
# + pycharm={}
default_cv = StratifiedKFold(n_splits=3, shuffle=True, random_state=0)
class Evaluator():
def __init__(self,
X, y,
metric="accuracy",
cv=default_cv):
# 初始化
self.X = X
self.y = y
self.metric = metric
self.cv = cv
# + [markdown] pycharm={}
# 然后我们需要实现`__call__`魔法方法,在该方法中实现整个评价过程:
# + pycharm={}
default_cv = StratifiedKFold(n_splits=3, shuffle=True, random_state=0)
class Evaluator():
def __init__(self,
X, y,
metric="accuracy",
cv=default_cv):
# 初始化
self.X = X
self.y = y
self.metric = metric
self.cv = cv
def __call__(self, config: dict) -> float:
layered_dict = layering_config(config)
AS_HP = layered_dict['classifier'].copy()
AS, HP = AS_HP.popitem()
ML_model = eval(AS)(**HP)
scores = cross_val_score(ML_model, self.X, self.y, cv=self.cv, scoring=self.metric)
score = scores.mean()
return 1 - score
# + [markdown] pycharm={}
# 实例化一个评价器对象:
# + pycharm={}
evaluator = Evaluator(X, y)
# + [markdown] pycharm={}
# 用配置空间的一个采样样本测试这个AutoML评价器:
# + pycharm={}
evaluator(CS.sample_configuration())
# + [markdown] pycharm={}
# ### Implement a Simple AutoML System Based on Above Knowledge
# + [markdown] pycharm={}
# **配置空间** 、**评价器**我们都有了,UltraOpt已经实现了成熟的**优化器**,所以我们只需要用`ultraopt.fmin`函数将这些组件串起来
# + pycharm={}
from ultraopt import fmin
# + pycharm={}
result = fmin(evaluator, HDL, optimizer="ETPE", n_iterations=40)
result
# + [markdown] pycharm={}
# 我们可以对AutoML得到的结果进行数据分析:
#
# 首先, 我们可以绘制拟合曲线:
# + pycharm={}
result.plot_convergence();
# + [markdown] pycharm={}
# 然后,我们想用`hiplot`绘制高维交互图。我们先整理出数据:
# + pycharm={}
data = result.plot_hi(return_data_only=True, target_name="accuracy", loss2target_func=lambda loss: 1 - loss)
# + [markdown] pycharm={}
# 如果直接绘图的话,您会发现由于算法选择的存在,会导致高维交互图非常的杂乱。我们简单编写一个函数,对不同的算法选择结果的超参进行可视化:
# + pycharm={}
import hiplot as hip
# + pycharm={}
def viz_subset(model_name, data):
data_filtered = []
for datum in data:
if datum["classifier:__choice__"] == model_name:
datum = datum.copy()
score = datum.pop("accuracy")
AS, HP = layering_config(datum)["classifier"].popitem()
HP["accuracy"] = score
data_filtered.append(HP)
hip.Experiment.from_iterable(data_filtered).display()
# + [markdown] pycharm={}
# 我们简单地对随机森林 `RandomForestClassifier` 进行可视化:
# -
# 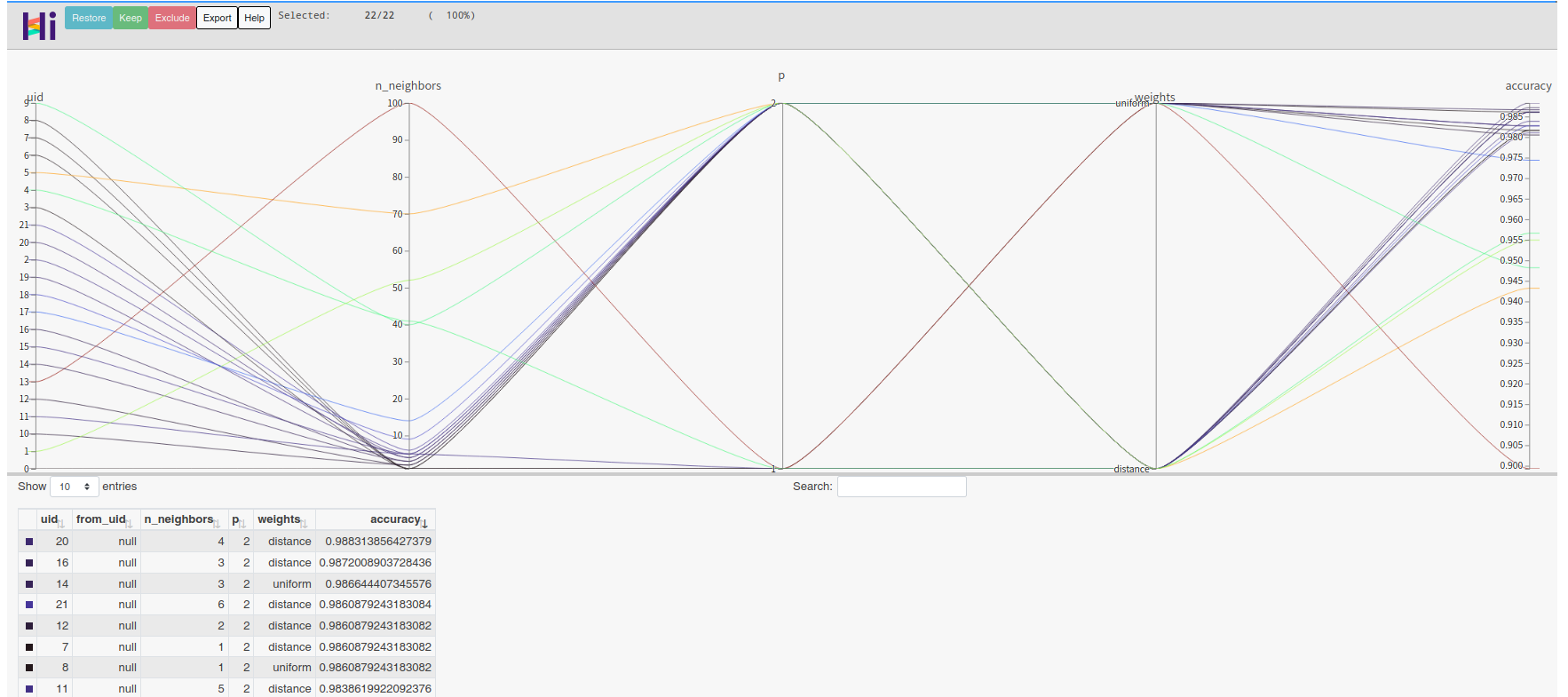
# 因为绘制`hiplot`高维交互图后`jupyter notebook`的文件大小会大量增加,所以我们以截图代替。您可以在自己的`notebook`中取消注释并执行以下代码:
# +
#viz_subset("RandomForestClassifier", data)
# +
#viz_subset("LinearSVC", data)
# +
#viz_subset("KNeighborsClassifier", data)
# + [markdown] pycharm={}
# ---
# + [markdown] pycharm={}
# 我们将所有代码整理为了 `05. Implement a Simple AutoML System.py` 脚本。
#
# 您可以在这个脚本中更直接地学习一个简单AutoML系统的搭建方法。
| tutorials/05. Implement a Simple AutoML System.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Your assignment is to get the last line to print without changing any
# of the code below. Instead, wrap each line that throws an error in a
# try/exept block.
try:
print("Infinity looks like + " + str(10 / 0) + ".")
except:
print("Woops. Can't do that.")
try:
print("I think her name was + " + name + "?")
except NameError:
print("Oh, I forgot to define 'name'. D'oh.")
try:
print("Your name is a nonsense number. Look: " + int("Gabriel"))
except ValueError:
print("Drat. 'Gabriel' isn't a number?")
print("I made it through the gauntlet. The message survived!")
# -
| 06-Python-APIs/2/Activities/08-Stu_MakingExceptions/Solved/Stu_MakingExceptions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
top_directory = '/Users/iaincarmichael/Dropbox/Research/law/law-net/'
import os
import sys
import time
from math import *
import copy
import cPickle as pickle
import re
import datetime
from collections import Counter
# data
import numpy as np
import pandas as pd
# viz
import matplotlib.pyplot as plt
# graph
import igraph as ig
# our code
sys.path.append(top_directory + 'code/')
from load_data import load_and_clean_graph, case_info
from pipeline.download_data import download_bulk_resource
from pipeline.make_clean_data import *
from viz import print_describe
from pipeline.make_raw_case_metadata import *
sys.path.append(top_directory + 'explore/vertex_metrics_experiment/code/')
from make_case_text_files import *
# directory set up
data_dir = top_directory + 'data/'
court_name = 'scotus'
# jupyter notebook settings
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
# +
# load scdb
scdb_modern = pd.read_csv(data_dir + 'scdb/SCDB_2016_01_caseCentered_Citation.csv', index_col=0)
scdb_legacy = pd.read_csv(data_dir + 'scdb/SCDB_Legacy_03_caseCentered_Citation.csv', index_col=0)
scdb = scdb_legacy.append(scdb_modern)
# -
# %time case_metadata = get_raw_case_metadata_from_court(court_name, data_dir)
# +
cert_cases = pd.DataFrame(False,
index=case_metadata.index,
columns=['denied', 'certiorari', 'zero_degree', 'scdb_link', 'in_scdb', 'text_length'])
cert_cases['text_length'] = 0
# add year
cert_cases['year'] = case_metadata['date'].apply(lambda d: d.year)
# -
# # find cases with bad words or short text
# +
# string search text for the words certiorari or denied
op_dir = data_dir + 'raw/' + court_name + '/opinions/'
# words we want to identify
bad_words = ['denied', 'certiorari']
i = 0
# check each opinion
for op_id in case_metadata.index:
i += 1
if int(log(i, 2)) == log(i, 2):
current_time = datetime.datetime.now().strftime('%H:%M:%S')
print '(%d/%d) at %s' % (i, len(case_metadata.index), current_time)
# grab the opinion file
op_path = op_dir + str(op_id) + '.json'
opinion = json_to_dict(op_path)
# get the lower case text
text = get_text_from_json(opinion)
text = text.lower()
# check each word in the text file
for word in bad_words:
if word in text:
cert_cases.loc[op_id, word] = True
# check if the text is really short
cert_cases.loc[op_id,'text_length'] = len(text)
# -
# # find cases with zero degree
# +
master_edgelist = pd.read_csv(data_dir + 'raw/edgelist_master_r.csv')
# dict keyed by case indicated if case is mentioned in the edgelist
max_id = max(master_edgelist['citing'].max(), master_edgelist['cited'].max())
mentions = {str(op_id): False for op_id in range(1, max_id + 1)}
i = 0
for index, edge in master_edgelist.iterrows():
i += 1
if int(log(i, 2)) == log(i, 2):
current_time = datetime.datetime.now().strftime('%H:%M:%S')
print '(%d/%d) at %s' % (i, len(master_edgelist), current_time)
# citing opinion mentioned
ing_op_id = str(edge[0])
ed_op_id = str(edge[1])
# cited opinion mentioned
mentions[ed_op_id] = True
# citing mentioned only if cited is not detroit lumber
if ed_op_id != 'g':
mentions[ing_op_id] = True
# +
case_ids = set(case_metadata.index)
zero_deg_cases = [op_id for op_id in mentions.keys() if (not mentions[op_id]) and (op_id in case_ids)]
cert_cases.loc[zero_deg_cases, 'zero_degree'] = True
# -
# # find cases without scdb links
# +
# scdb ids
scdb_ids = set(scdb.index)
for index, row in case_metadata.iterrows():
# check if case has link to SCDB id
if len(row['scdb_id']) > 0:
cert_cases.loc[index, 'scdb_link'] = True
# check if SCDB id is in SCDB ids
if row['scdb_id'] in scdb_ids:
cert_cases.loc[index, 'in_scdb'] = True
# -
cert_cases[~cert_cases['scdb_link']].index
# # save cert_cases file
#
# +
# cert_cases.to_csv('cert_cases_data.csv', index=True)
# cert_cases = pd.read_csv('cert_cases_data.csv', index_col=0)
# -
# # compare year counts between CL and scdb
# +
# initialize pandas series
CL_year_counts = pd.Series(0, index=range(1754, 2017))
scdb_year_counts = pd.Series(0, index=range(1754, 2017))
# count cases per year
CL_year_counter = Counter(cert_cases['year'])
scdb_year_counter = Counter(scdb['dateDecision'].apply(lambda d: d.split('/')[2]))
# make fill series
for y in CL_year_counts.index:
CL_year_counts[y] = CL_year_counter[y]
scdb_year_counts[y] = scdb_year_counter[y]
difference = CL_year_counts - scdb_year_counts
#years that have a lot of extra cases
bad_years = difference[difference > 400].index.tolist()
# plot difference
difference.plot()
plt.ylabel('difference')
plt.title('yearly case counts')
# -
# # analyze cert_cases
cert_cases.mean(axis=0)
# +
cases_denied = cert_cases['denied']
cases_certiorari = cert_cases['certiorari']
cases_zero_degree = cert_cases['zero_degree']
cases_no_scdb_link = ~cert_cases['scdb_link']
cases_notin_scdb = ~cert_cases['in_scdb']
cases_bad_years = cert_cases['year'].apply(lambda y: y in bad_years)
# 2 cases with scdb links but that don't registar in scdb
# cert_cases[~cases_no_scdb_link & cases_notin_scdb]
# +
# cert_cases[cases_zero_degree & cases_denied & cases_certiorari & cases_no_scdb_link]
# cert_cases[~cases_bad_years & cases_notin_scdb]
cert_cases[cases_certiorari & cases_denied & cases_zero_degree & ~cases_no_scdb_link]
# +
op_id = 106306
print case_metadata.loc[str(op_id)]
print
print case_info(op_id)
# -
# # cases missing SCDB links
cases_no_scdb = pd.Series(cert_cases[~cert_cases['scdb_link']].index)
cases_no_scdb.to_csv('no_scdb_link.csv', index=False)
| explore/make_data/expolore_certorari.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# name: ir
# ---
# + [markdown] id="W_tvPdyfA-BL"
# ##### Copyright 2019 The TensorFlow Authors.
# + cellView="form" id="0O_LFhwSBCjm"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="9-3Pry4jh1-E"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/rses-dl-course/rses-dl-course.github.io/blob/master/notebooks/R/R06_saving_and_loading_models.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/rses-dl-course/rses-dl-course.github.io/blob/master/notebooks/R/R06_saving_and_loading_models.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# </table>
# + [markdown] id="NxjpzKTvg_dd"
# # Lab 05: Saving and Loading Models
#
# In this lab we will learn how we can take a trained model, save it, and then load it back to keep training it or use it to perform inference. In particular, we will use transfer learning to train a classifier to classify images of cats and dogs, just like we did in the previous lesson. We will then take our trained model and save it as an HDF5 file, which is the format used by Keras. We will then load this model, use it to perform predictions, and then continue to train the model. Finally, we will save our trained model as a TensorFlow SavedModel and then we will download it to a local disk, so that it can later be used for deployment in different platforms.
# + [markdown] id="crU-iluJIEzw"
# ## Concepts that will be covered in this Colab
#
# 1. Saving models in HDF5 format for Keras
# 3. Loading models
#
# Before starting this Colab, you should reset the Colab environment by selecting `Runtime -> Reset all runtimes...` from menu above.
# + [markdown] id="7RVsYZLEpEWs"
# # Install and load dependencies
#
# + [markdown] id="ZUCEcRdhnyWn"
# First, you'll need to install and load R package Keras which will also install TensorFlow. We'll also install package fs which has useful functionality for working with our filesystem.
# + id="z0rPA7CkF5l8"
install.packages(c("keras", "fs"))
library(keras)
# + [markdown] id="vU8P2MUDrIUu"
# # Part 1: Load the Cats vs. Dogs Dataset
# + [markdown] id="9crYcwgzrH9A"
# ## Dataset
#
# We download the dataset again. The dataset we are using is a filtered version of <a href="https://www.kaggle.com/c/dogs-vs-cats/data" target="_blank">Dogs vs. Cats</a> dataset from Kaggle (ultimately, this dataset is provided by Microsoft Research).
# + id="NFpoMDvrTTLN"
URL <- "https://storage.googleapis.com/mledu-datasets/cats_and_dogs_filtered.zip"
zip_dir <- get_file('cats_and_dogs_filtered.zip', origin = URL, extract = TRUE)
# + [markdown] id="rKNVlt1MTbEf"
# The dataset we have downloaded has the following directory structure.
#
# <pre style="font-size: 10.0pt; font-family: Arial; line-height: 2; letter-spacing: 1.0pt;" >
# <b>cats_and_dogs_filtered</b>
# |__ <b>train</b>
# |______ <b>cats</b>: [cat.0.jpg, cat.1.jpg, cat.2.jpg ...]
# |______ <b>dogs</b>: [dog.0.jpg, dog.1.jpg, dog.2.jpg ...]
# |__ <b>validation</b>
# |______ <b>cats</b>: [cat.2000.jpg, cat.2001.jpg, cat.2002.jpg ...]
# |______ <b>dogs</b>: [dog.2000.jpg, dog.2001.jpg, dog.2002.jpg ...]
# </pre>
#
# We can list the directories with the following terminal command:
# + id="ZR3B2TqRV_Pr"
zip_dir_base <- dirname(zip_dir)
fs::dir_tree(zip_dir_base, recurse = 2)
# + [markdown] id="7KnMK35BWGqC"
# We'll now assign variables with the proper file path for the training and validation sets. We'll also create some variables that hold information about the size of our datasets
# + id="KlP-iXwoWHRO"
base_dir <- fs::path(zip_dir_base, "cats_and_dogs_filtered")
train_dir <- fs::path(base_dir, "train")
validation_dir <- fs::path(base_dir, "validation")
train_cats_dir <- fs::path(train_dir, "cats")
train_dogs_dir <- fs::path(train_dir, "dogs")
validation_cats_dir <- fs::path(validation_dir, "cats")
validation_dogs_dir <- fs::path(validation_dir, "dogs")
num_cats_tr <- length(fs::dir_ls(train_cats_dir))
num_dogs_tr <- length(fs::dir_ls(train_dogs_dir))
num_cats_val <- length(fs::dir_ls(validation_cats_dir))
num_dogs_val <- length(fs::dir_ls(validation_dogs_dir))
total_train <- num_cats_tr + num_dogs_tr
total_val <- num_cats_val + num_dogs_val
# + [markdown] id="tsRhwiQ8rkQv"
# Lets create training and validation image generators to read our images from their directories which rescales our image to values from 0 to 1. Let's also create a flow from our training directories which also resizes our images to the resolution (224 x 224) expected by our MobileNetV2 model we'll be using for transfer learning. Let's also set the batch size to 32.
# + id="FEtvEPm6r8hO"
batch_size <- 32
image_res <- 224
# training generators
train_image_generator <- image_data_generator(rescale = 1/255,
rotation_range = 45,
width_shift_range = 0.2,
height_shift_range = 0.2,
shear_range = 0.2,
zoom_range = 0.2,
horizontal_flip = TRUE,
fill_mode = 'nearest')
train_data_gen <- flow_images_from_directory(directory = train_dir,
generator = train_image_generator,
target_size = c(image_res, image_res),
class_mode = "binary",
batch_size = batch_size)
# validation generators
val_image_generator <- image_data_generator(rescale = 1/255)
val_data_gen <- flow_images_from_directory(directory = validation_dir,
generator = val_image_generator,
target_size = c(image_res, image_res),
class_mode = "binary",
batch_size = batch_size)
# + [markdown] id="41zDiZdKsYvo"
# # Part 2: Transfer Learning with TensorFlow Hub
#
#
# Let's now use the [MobileNet V2 model architecture](https://keras.rstudio.com/reference/application_mobilenet_v2.html) to do Transfer Learning.
# + id="kAc_Viz0siWC"
feature_extractor <- application_mobilenet_v2(input_shape = c(image_res, image_res, 3),
include_top = FALSE, pooling = "avg")
# + [markdown] id="Rn3zRO2Cslpq"
# Freeze the variables in the feature extractor layer, so that the training only modifies the final classifier layer.
# + id="HqMWwpNzsqv9"
freeze_weights(feature_extractor)
# + [markdown] id="Dk_WOWlms0Vy"
# Now let's use are feature extractor as part of a keras sequential model, and add a new classification layer of 2 units with softmax activation.
# + id="od6ur8sdswqO"
model <- keras_model_sequential() %>%
feature_extractor() %>%
layer_dense(units = 2, activation = "softmax")
model
# + [markdown] id="FhT9p4viV_tX"
# ## Train the model
#
# We now train this model like any other using an image generator, by first calling `compile` followed by `fit`.
# + id="MfZYgHQrT6vD"
model %>% compile(optimizer="adam",
loss = 'sparse_categorical_crossentropy',
metrics = "accuracy")
epochs = 3
history <- model %>%
fit(x = train_data_gen,
epochs = epochs,
validation_data = val_data_gen)
cat('Validation loss:', format(tail(history$metrics$val_loss, 1), digits = 2), "\n")
cat('Validation accuracy:', format(tail(history$metrics$val_accuracy, 1), digits = 2), "\n")
# + [markdown] id="oNwjmjaItOM7"
# ## Check the predictions
#
# To check our model, let's use it to make some predictions on the images from a single batch of our validation image generator.
#
# Let's compile our predictions into a data.frame. Let's also add the actual labels for each row to the data.frame and have a look at our predictions. We can see that predictions using our transfer learning very accurate with very high confidence in each prediciton!
# + id="C5Hv2-9hteps"
pred_batch <- val_data_gen[1]
pred_mat <- model %>%
predict(pred_batch[[1]])
predictions <- data.frame(class_description = names(val_data_gen$class_indices)[apply(pred_mat, 1, which.max)],
score = apply(pred_mat, 1, max),
label = names(val_data_gen$class_indices)[pred_batch[[2]] + 1])
predictions
# + [markdown] id="mNgGWQLKIowr"
# Let's now plot the images from our Dogs vs Cats dataset and put the predicted labels above them and their actual labels below. First we create a plotting function:
# + id="KMSo5SUpD0wE"
plot_rgb_image_ttl <- function(image_array, prediction){
image_array %>%
array_reshape(dim = c(dim(.)[1:3])) %>%
as.raster(max = 1) %>%
plot()
title(main = paste0(prediction$class_description,
" (", format(prediction$score * 100, digits = 2), "%)"),
sub = prediction$label)
}
# + [markdown] id="m9O6RHnsu3up"
# Then we loop the function over the first 10 images:
# + id="euKNnitcu8ul"
options(repr.plot.width = 16, repr.plot.height = 8)
# Set the layout
layout(matrix(1:10, ncol = 5))
# Loop plotting over the batch of images
for(i in 1:10){
plot_rgb_image_ttl(pred_batch[[1]][i,,,], predictions[i,])
}
# + [markdown] id="mmPQYQLx3cYq"
# # Part 3: Save as Keras `.h5` model
#
# Now that we've trained the model, we can save it as an HDF5 file, which is the format used by Keras using function [`save_model_hdf5()`](https://keras.rstudio.com/reference/save_model_hdf5.html). Our HDF5 file will have the extension '.h5', and it's name will correpond to the current time stamp.
# + id="tCnNWTkZ3Ckz"
export_path_keras = paste0("./", as.integer(Sys.time()) ,".h5")
export_path_keras
# + id="jgL20byNv6Ii"
model %>% save_model_hdf5(filepath = export_path_keras)
# + id="hi3YWnl51vMU"
fs::dir_tree(recurse = 0)
# + [markdown] id="3u6BiynOxs6z"
# You can later recreate the same model from this file, even if you no longer have access to the code that created the model.
#
# This file includes:
#
# - The model's architecture
# - The model's weight values (which were learned during training)
# - The model's training config (what you passed to `compile`), if any
# - The optimizer and its state, if any (this enables you to restart training where you left off)
# + [markdown] id="3QMVDmgAx3-L"
# # Part 4: Load the Keras `.h5` Model
#
# We will now load the model we just saved into a new model called `reloaded`.
# + id="CXFwZsFQyCfy"
reloaded_model <- load_model_hdf5(export_path_keras)
reloaded_model
# + [markdown] id="LX_EiYB42s_k"
# We can check that the reloaded model and the previous model give the same result. Let's make some predictions using the reloaded model on the batch of images we had previously used.
# + id="xcIS51Vx2RUc"
pred_mat_reloaded <- reloaded_model %>%
predict(pred_batch[[1]])
# + [markdown] id="rBmRjIdN279b"
# Predicted classes from both models should be equal so the following statement should return `TRUE`
# + id="6C-lVrNi2bru"
all(apply(pred_mat, 1, which.max) == apply(pred_mat_reloaded, 1, which.max))
# + [markdown] id="AGqic8G33KtH"
# # Keep Training
#
# Besides making predictions, we can also take our `reloaded_model` and keep training it. To do this, you can just train the `reloaded_model` as usual, using the `fit` method.
# + id="T38yNY0i3VKn"
epochs = 3
history <- model %>%
fit(x = train_data_gen,
epochs = epochs,
validation_data = val_data_gen)
cat('Validation loss:', format(tail(history$metrics$val_loss, 1), digits = 2), "\n")
cat('Validation accuracy:', format(tail(history$metrics$val_accuracy, 1), digits = 2), "\n")
| notebooks/R/R05_saving_and_loading_models.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="aklJxkHBD5aR"
# # Predicting babyweight using BigQuery ML
#
# This notebook illustrates:
# <ol>
# <li> Machine Learning using BigQuery
# <li> Make a Prediction with BQML using the Model
# </ol>
#
# Please see [this notebook](1_explore.ipynb) for more context on this problem and how the features were chosen.
# -
# !sudo chown -R jupyter:jupyter /home/jupyter/training-data-analyst
# Ensure the right version of Tensorflow is installed.
# !pip freeze | grep tensorflow==2.1
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="BhUiclqCD5aT"
# change these to try this notebook out
PROJECT = 'cloud-training-demos'
REGION = 'us-central1'
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="F10_KsX7D5aX"
import os
os.environ['PROJECT'] = PROJECT
os.environ['REGION'] = REGION
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="YrqwSv6vD5aZ" outputId="402ed11b-ab4a-4480-a1aa-f72741cd2cc1" language="bash"
# gcloud config set project $PROJECT
# gcloud config set compute/region $REGION
# + [markdown] colab_type="text" id="j782YaAzD5ae"
# ## Exploring the Data
#
# Here, we will be taking natality data and training on features to predict the birth weight.
#
# The CDC's Natality data has details on US births from 1969 to 2008 and is available in BigQuery as a public data set. More details: https://bigquery.cloud.google.com/table/publicdata:samples.natality?tab=details
#
# Lets start by looking at the data since 2000 with useful values > 0!
# -
# %%bigquery
SELECT
*
FROM
publicdata.samples.natality
WHERE
year > 2000
AND gestation_weeks > 0
AND mother_age > 0
AND plurality > 0
AND weight_pounds > 0
LIMIT 10
# + [markdown] colab_type="text" id="ep02J20pD5ai"
# ## Define Features
#
# Looking over the data set, there are a few columns of interest that could be leveraged into features for a reasonable prediction of approximate birth weight.
#
# Further, some feature engineering may be accomplished with the BigQuery `CAST` function -- in BQML, all strings are considered categorical features and all numeric types are considered continuous ones.
#
# The hashmonth is added so that we can repeatably split the data without leakage -- we want all babies that share a birthday to be either in training set or in test set and not spread between them (otherwise, there would be information leakage when it comes to triplets, etc.)
# -
# %%bigquery
SELECT
weight_pounds, -- this is the label; because it is continuous, we need to use regression
CAST(is_male AS STRING) AS is_male,
mother_age,
CAST(plurality AS STRING) AS plurality,
gestation_weeks,
FARM_FINGERPRINT(CONCAT(CAST(YEAR AS STRING), CAST(month AS STRING))) AS hashmonth
FROM
publicdata.samples.natality
WHERE
year > 2000
AND gestation_weeks > 0
AND mother_age > 0
AND plurality > 0
AND weight_pounds > 0
LIMIT 10
# + [markdown] colab_type="text" id="H-d7C8KcD5am"
# ## Train Model
#
# With the relevant columns chosen to accomplish predictions, it is then possible to create (train) the model in BigQuery. First, a dataset will be needed store the model. (if this throws an error in Datalab, simply create the dataset from the BigQuery console).
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="bWMZJQdKD5an" language="bash"
# bq --location=US mk -d demo
# + [markdown] colab_type="text" id="T9JZIQ9nD5ap"
# With the demo dataset ready, it is possible to create a linear regression model to train the model.
#
# This will take approximately **4 minutes** to run and will show **Done** when complete.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="V-UJOX0tD5aq" outputId="63aa219c-a14f-4db5-a271-2e614759fe1a"
# %%bigquery
CREATE or REPLACE MODEL demo.babyweight_model_asis
OPTIONS
(model_type='linear_reg', labels=['weight_pounds']) AS
WITH natality_data AS (
SELECT
weight_pounds,-- this is the label; because it is continuous, we need to use regression
CAST(is_male AS STRING) AS is_male,
mother_age,
CAST(plurality AS STRING) AS plurality,
gestation_weeks,
FARM_FINGERPRINT(CONCAT(CAST(YEAR AS STRING), CAST(month AS STRING))) AS hashmonth
FROM
publicdata.samples.natality
WHERE
year > 2000
AND gestation_weeks > 0
AND mother_age > 0
AND plurality > 0
AND weight_pounds > 0
)
SELECT
weight_pounds,
is_male,
mother_age,
plurality,
gestation_weeks
FROM
natality_data
WHERE
ABS(MOD(hashmonth, 4)) < 3 -- select 75% of the data as training
# + [markdown] colab_type="text" id="mJYtGxWyD5at"
# ## Training Statistics
# + [markdown] colab_type="text" id="0L92s-gtD5au"
# During the model training (and after the training), it is possible to see the model's training evaluation statistics.
#
# For each training run, a table named `<model_name>_eval` is created. This table has basic performance statistics for each iteration.
#
# While the new model is training, review the training statistics in the BigQuery UI to see the below model training: https://bigquery.cloud.google.com/
#
# Since these statistics are updated after each iteration of model training, you will see different values for each refresh while the model is training.
#
# The training details may also be viewed after the training completes from this notebook.
# -
# %%bigquery
SELECT * FROM ML.TRAINING_INFO(MODEL demo.babyweight_model_asis);
# + [markdown] colab_type="text" id="CATWu60vD5ay"
# Some of these columns are obvious although what do the non-specific ML columns mean (specific to BQML)?
#
# **training_run** - Will be zero for a newly created model. If the model is re-trained using warm_start, this will increment for each re-training.
#
# **iteration** - Number of the associated `training_run`, starting with zero for the first iteration.
#
# **duration_ms** - Indicates how long the iteration took (in ms).
#
# Note: You can also see these stats by refreshing the BigQuery UI window, finding the `<model_name>` table, selecting on it, and then the Training Stats sub-header.
#
# Observe the training and evaluation loss and see if the model has an overfit.
# + [markdown] colab_type="text" id="9QPwMA_ZD5a3"
# ## Make a Prediction with BQML using the Model
# + [markdown] colab_type="text" id="9QBZxMb6D5a5"
# With a trained model, it is now possible to make a prediction on the values. The only difference from the second query above is the reference to the model. The data has been limited (`LIMIT 100`) to reduce amount of data returned.
#
# When the `ml.predict` function is leveraged, output prediction column name for the model is `predicted_<label_column_name>`.
# -
# %%bigquery
SELECT
*
FROM
ml.PREDICT(MODEL demo.babyweight_model_asis,
(SELECT
weight_pounds,
CAST(is_male AS STRING) AS is_male,
mother_age,
CAST(plurality AS STRING) AS plurality,
gestation_weeks
FROM
publicdata.samples.natality
WHERE
year > 2000
AND gestation_weeks > 0
AND mother_age > 0
AND plurality > 0
AND weight_pounds > 0
))
LIMIT 100
# + [markdown] colab_type="text" id="0DZorSI-D5a9"
# # More advanced...
#
# In the original example, we were taking into account the idea that if no ultrasound has been performed, some of the features (e.g. is_male) will not be known. Therefore, we augmented the dataset with such masked features and trained a single model to deal with both these scenarios.
#
# In addition, during data exploration, we learned that the data size set for mothers older than 45 was quite sparse, so we will discretize the mother age.
# -
# %%bigquery
SELECT
weight_pounds,
CAST(is_male AS STRING) AS is_male,
IF(mother_age < 18, 'LOW',
IF(mother_age > 45, 'HIGH',
CAST(mother_age AS STRING))) AS mother_age,
CAST(plurality AS STRING) AS plurality,
CAST(gestation_weeks AS STRING) AS gestation_weeks,
FARM_FINGERPRINT(CONCAT(CAST(YEAR AS STRING), CAST(month AS STRING))) AS hashmonth
FROM
publicdata.samples.natality
WHERE
year > 2000
AND gestation_weeks > 0
AND mother_age > 0
AND plurality > 0
AND weight_pounds > 0
LIMIT 25
# + [markdown] colab_type="text" id="ZwetSzIeD5bC"
# On the same dataset, will also suppose that it is unknown whether the child is male or female (on the same dataset) to simulate that an ultrasound was not been performed.
# -
# %%bigquery
SELECT
weight_pounds,
'Unknown' AS is_male,
IF(mother_age < 18, 'LOW',
IF(mother_age > 45, 'HIGH',
CAST(mother_age AS STRING))) AS mother_age,
IF(plurality > 1, 'Multiple', 'Single') AS plurality,
CAST(gestation_weeks AS STRING) AS gestation_weeks,
FARM_FINGERPRINT(CONCAT(CAST(YEAR AS STRING), CAST(month AS STRING))) AS hashmonth
FROM
publicdata.samples.natality
WHERE
year > 2000
AND gestation_weeks > 0
AND mother_age > 0
AND plurality > 0
AND weight_pounds > 0
LIMIT 25
# + [markdown] colab_type="text" id="MmY2chPyD5bI"
# Bringing these two separate data sets together, there is now a dataset for male or female children determined with ultrasound or unknown if without.
# +
# %%bigquery
WITH with_ultrasound AS (
SELECT
weight_pounds,
CAST(is_male AS STRING) AS is_male,
IF(mother_age < 18, 'LOW',
IF(mother_age > 45, 'HIGH',
CAST(mother_age AS STRING))) AS mother_age,
CAST(plurality AS STRING) AS plurality,
CAST(gestation_weeks AS STRING) AS gestation_weeks,
FARM_FINGERPRINT(CONCAT(CAST(YEAR AS STRING), CAST(month AS STRING))) AS hashmonth
FROM
publicdata.samples.natality
WHERE
year > 2000
AND gestation_weeks > 0
AND mother_age > 0
AND plurality > 0
AND weight_pounds > 0
),
without_ultrasound AS (
SELECT
weight_pounds,
'Unknown' AS is_male,
IF(mother_age < 18, 'LOW',
IF(mother_age > 45, 'HIGH',
CAST(mother_age AS STRING))) AS mother_age,
IF(plurality > 1, 'Multiple', 'Single') AS plurality,
CAST(gestation_weeks AS STRING) AS gestation_weeks,
FARM_FINGERPRINT(CONCAT(CAST(YEAR AS STRING), CAST(month AS STRING))) AS hashmonth
FROM
publicdata.samples.natality
WHERE
year > 2000
AND gestation_weeks > 0
AND mother_age > 0
AND plurality > 0
AND weight_pounds > 0
),
preprocessed AS (
SELECT * from with_ultrasound
UNION ALL
SELECT * from without_ultrasound
)
SELECT
weight_pounds,
is_male,
mother_age,
plurality,
gestation_weeks
FROM
preprocessed
WHERE
ABS(MOD(hashmonth, 4)) < 3
LIMIT 25
# + [markdown] colab_type="text" id="hNnZxDw9D5bN"
# ## Create a new model
#
# With a data set which has been feature engineered, it is ready to create model with the `CREATE or REPLACE MODEL` statement
#
# This will take **5-10 minutes** and will show **Done** when complete.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="aKPqXqicD5bP"
# %%bigquery
CREATE or REPLACE MODEL demo.babyweight_model_fc
OPTIONS
(model_type='linear_reg', labels=['weight_pounds']) AS
WITH with_ultrasound AS (
SELECT
weight_pounds,
CAST(is_male AS STRING) AS is_male,
IF(mother_age < 18, 'LOW',
IF(mother_age > 45, 'HIGH',
CAST(mother_age AS STRING))) AS mother_age,
CAST(plurality AS STRING) AS plurality,
CAST(gestation_weeks AS STRING) AS gestation_weeks,
FARM_FINGERPRINT(CONCAT(CAST(YEAR AS STRING), CAST(month AS STRING))) AS hashmonth
FROM
publicdata.samples.natality
WHERE
year > 2000
AND gestation_weeks > 0
AND mother_age > 0
AND plurality > 0
AND weight_pounds > 0
),
without_ultrasound AS (
SELECT
weight_pounds,
'Unknown' AS is_male,
IF(mother_age < 18, 'LOW',
IF(mother_age > 45, 'HIGH',
CAST(mother_age AS STRING))) AS mother_age,
IF(plurality > 1, 'Multiple', 'Single') AS plurality,
CAST(gestation_weeks AS STRING) AS gestation_weeks,
FARM_FINGERPRINT(CONCAT(CAST(YEAR AS STRING), CAST(month AS STRING))) AS hashmonth
FROM
publicdata.samples.natality
WHERE
year > 2000
AND gestation_weeks > 0
AND mother_age > 0
AND plurality > 0
AND weight_pounds > 0
),
preprocessed AS (
SELECT * from with_ultrasound
UNION ALL
SELECT * from without_ultrasound
)
SELECT
weight_pounds,
is_male,
mother_age,
plurality,
gestation_weeks
FROM
preprocessed
WHERE
ABS(MOD(hashmonth, 4)) < 3
# + [markdown] colab_type="text" id="mJYtGxWyD5at"
# ## Training Statistics
# + [markdown] colab_type="text" id="0L92s-gtD5au"
# While the new model is training, review the training statistics in the BigQuery UI to see the below model training: https://bigquery.cloud.google.com/
#
# The training details may also be viewed after the training completes from this notebook.
# -
# %%bigquery
SELECT * FROM ML.TRAINING_INFO(MODEL demo.babyweight_model_fc);
# + [markdown] colab_type="text" id="1jlHzJqfD5bU"
# ## Make a prediction with the new model
#
#
# + [markdown] colab_type="text" id="EGo8fj9qD5bU"
# Perhaps it is of interest to make a prediction of the baby's weight given a number of other factors: Male, Mother is 28 years old, Mother will only have one child, and the baby was born after 38 weeks of pregnancy.
#
# To make this prediction, these values will be passed into the SELECT statement.
# -
# %%bigquery
SELECT
*
FROM
ml.PREDICT(MODEL demo.babyweight_model_fc,
(SELECT
'True' AS is_male,
'28' AS mother_age,
'1' AS plurality,
'38' AS gestation_weeks
))
# + [markdown] colab_type="text" id="PK_-WNGUD5bX"
# <br>
# <br>
# <br>
# <br>
# Copyright 2020 Google Inc. Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0 Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License
| courses/machine_learning/deepdive/06_structured/5_train_bqml.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Flickr30k to Features
#
# * <NAME>, <NAME>, <NAME>, and <NAME>. _From image description to visual denotations: New similarity metrics for semantic inference over event descriptions._ Transactions of the Association for Computational Linguistics (to appear).
#
#
# +
import os
import tensorflow.contrib.keras as keras
import numpy as np
import datetime
t_start=datetime.datetime.now()
import pickle
# -
image_folder_path = './data/Flickr30k/flickr30k-images'
# +
output_dir = './data/cache'
output_filepath = os.path.join(output_dir,
'FEATURES_%s_%s.pkl' % (
image_folder_path.replace('./', '').replace('/', '_'),
t_start.strftime("%Y-%m-%d_%H-%M"),
), )
output_filepath
# -
from tensorflow.contrib.keras.api.keras.applications.inception_v3 import decode_predictions
from tensorflow.contrib.keras.api.keras.preprocessing import image as keras_preprocessing_image
# +
from tensorflow.contrib.keras.api.keras.applications.inception_v3 import InceptionV3, preprocess_input
BATCHSIZE=16
# -
model = InceptionV3(weights='imagenet', include_top=False, pooling='avg')
print("InceptionV3 loaded")
# #### Plan
#
# * Form a list of every file in the image directory
# * Run InceptionV3 over the list
# * Save off features to an easy-to-load filetype
# +
import re
good_image = re.compile( r'\.(jpg|png|gif)$', flags=re.IGNORECASE )
img_arr = [ f for f in os.listdir(image_folder_path) if good_image.search(f) ]
', '.join( img_arr[:3] ), ', '.join( img_arr[-3:] )
# +
# Create a generator for preprocessed images
def preprocessed_image_gen():
#target_size=model.input_shape[1:]
target_size=(299, 299, 3)
print("target_size", target_size)
for img_name in img_arr:
#print("img_name", img_name)
img_path = os.path.join(image_folder_path, img_name)
img = keras_preprocessing_image.load_img(img_path, target_size=target_size)
yield keras.preprocessing.image.img_to_array(img)
#x = np.expand_dims(x, axis=0) # This is to make a single image into a suitable array
def image_batch(batchsize=BATCHSIZE):
while True: # This needs to run 'for ever' for Keras input, even if only a fixed number are required
preprocessed_image_generator = preprocessed_image_gen()
start = True
for img in preprocessed_image_generator:
if start:
arr, n, start = [], 0, False
arr.append(img)
n += 1
if n>=batchsize:
stack = np.stack( arr, axis=0 )
#print("stack.shape", stack.shape)
preprocessed = preprocess_input( stack )
#print("preprocessed.shape", preprocessed.shape)
yield preprocessed
start=True
if len(arr)>0:
stack = np.stack( arr, axis=0 )
print("Final stack.shape", stack.shape)
preprocessed = preprocess_input( stack )
print("Final preprocessed.shape", preprocessed.shape)
yield preprocessed
# -
if False:
image_batcher = image_batch()
batch = next(image_batcher)
features = model.predict_on_batch(batch)
features.shape
# +
# This should do the batch creation on the CPU and the analysis on the GPU asynchronously.
import math # for ceil
t0=datetime.datetime.now()
features = model.predict_generator(image_batch(), steps = math.ceil( len(img_arr)/BATCHSIZE) ) #, verbose=1
features.shape, (datetime.datetime.now()-t0)/len(img_arr)*1000.
# +
# Save the data into a useful structure
save_me = dict(
features = features,
img_arr = img_arr,
)
if not os.path.exists(output_dir):
os.makedirs(output_dir)
with open( output_filepath, 'wb') as f:
pickle.dump(save_me, f)
print("Features saved to '%s'" %(output_filepath,))
# -
| notebooks/2-CNN/7-Captioning/1-folder-images-to-features.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Indeterminate Beam calculator
### Table of Contents
1. Initialisation
2. Basic usage (Readme example)
- 2(b). Additional features (Readme example)
- 2(c). Changing units (Readme example)
3. Support class breakdown
4. Load classes breakdown
5. Statically determinate beam (Ex 12.14 Hibbeler)
6. Statically Indeterminate beam (Ex 12.21 Hibbeler)
7. Spring supported beam deflection (Ex 12.16 Hibbeler)
8. Axial Loaded Indeterminate Beam (Ex 4.13 Hibbeler)
Note:
You must always run the initialization cell first.
Then you are free to move to any of the other sections as per the table of contents.
# -
# ## 1. INITIALIZATION
#
# +
# RUN THIS CELL FIRST TO INITIALISE GOOGLE NOTEBOOK!!!!
# !pip install indeterminatebeam
# %matplotlib inline
# import beam and supports
from indeterminatebeam import Beam, Support
# import loads (all load types imported for reference)
from indeterminatebeam import (
PointTorque,
PointLoad,
PointLoadV,
PointLoadH,
UDL,
UDLV,
UDLH,
TrapezoidalLoad,
TrapezoidalLoadV,
TrapezoidalLoadH,
DistributedLoad,
DistributedLoadV,
DistributedLoadH
)
# Note: load ending in V are vertical loads
# load ending in H are horizontal loads
# load not ending in either takes angle as an input (except torque)
# +
# NOT FOR USE ONLINE.
# Use this instead for initialization if running directly from package folder,
import sys, os
sys.path.insert(0, os.path.abspath('../../'))
# import beam and supports
from indeterminatebeam import Beam, Support
# import loads (all load types imported for reference)
from indeterminatebeam.loading import (
PointTorque,
PointLoad,
PointLoadV,
PointLoadH,
UDL,
UDLV,
UDLH,
TrapezoidalLoad,
TrapezoidalLoadV,
TrapezoidalLoadH,
DistributedLoad,
DistributedLoadV,
DistributedLoadH
)
# Note: load ending in V are vertical loads
# load ending in H are horizontal loads
# load not ending in either takes angle as an input (except torque)
# -
# ## 2. Basic Usage (Readme example)
# +
# Arbritrary example defined in README.md
beam = Beam(7) # Initialize a Beam object of length 7 m with E and I as defaults
beam_2 = Beam(9,E=2000, I =100000) # Initialize a Beam specifying some beam parameters
a = Support(5,(1,1,0)) # Defines a pin support at location x = 5 m
b = Support(0,(0,1,0)) # Defines a roller support at location x = 0 m
c = Support(7,(1,1,1)) # Defines a fixed support at location x = 7 m
beam.add_supports(a,b,c)
load_1 = PointLoadV(1000,2) # Defines a point load of 1000 N acting up, at location x = 2 m
load_2 = DistributedLoadV(2000,(1,4)) # Defines a 2000 N/m UDL from location x = 1 m to x = 4 m
load_3 = PointTorque(2*10**3, 3.5) # Defines a 2*10**3 N.m point torque at location x = 3.5 m
beam.add_loads(load_1,load_2,load_3) # Assign the support objects to a beam object created earlier
beam.analyse()
fig_1 = beam.plot_beam_external()
fig_1.show()
fig_2 = beam.plot_beam_internal()
fig_2.show()
# -
# ## 2(b). Additional Features (Readme example)
# +
# Run section 2 (prior to running this example)
# query for the data at a specfic point (note units are not provided)
print("bending moments at 3 m: " + str(beam.get_bending_moment(3)))
print("shear forces at 1,2,3,4,5m points: " + str(beam.get_shear_force(1,2,3,4,5)))
print("normal force absolute max: " + str(beam.get_normal_force(return_absmax=True)))
print("deflection max: " + str(beam.get_deflection(return_max = True)))
##add a query point to a plot (adds values on plot)
beam.add_query_points(1,3,5)
beam.remove_query_points(5)
## plot the results for the beam
fig = beam.plot_beam_internal()
fig.show()
# -
# ## 2(c). Changing units (Readme example)
# +
# Readme example as a demonstration for changing units
# used and presented in following example. Note that
# the example is conceptually identical only with
# different units.
# update units usings the update_units() function.
# use the command below for more information.
# help(beam.update_units)
# note: initialising beam with the anticipation that units will be updated
beam = Beam(7000, E = 200 * 10 **6, I = 9.05 * 10 **6)
beam.update_units(key='length', unit='mm')
beam.update_units('force', 'kN')
beam.update_units('distributed', 'kN/m')
beam.update_units('moment', 'kN.m')
beam.update_units('E', 'kPa')
beam.update_units('I', 'mm4')
beam.update_units('deflection', 'mm')
a = Support(5000,(1,1,0)) # Defines a pin support at location x = 5 m (x = 5000 mm)
b = Support(0,(0,1,0)) # Defines a roller support at location x = 0 m
c = Support(7000,(1,1,1)) # Defines a fixed support at location x = 7 m (x = 7000 mm)
beam.add_supports(a,b,c)
load_1 = PointLoadV(1,2000) # Defines a point load of 1000 N (1 kN) acting up, at location x = 2 m
load_2 = DistributedLoadV(2,(1000,4000)) # Defines a 2000 N/m (2 kN/m) UDL from location x = 1 m to x = 4 m
load_3 = PointTorque(2, 3500) # Defines a 2*10**3 N.m (2 kN.m) point torque at location x = 3.5 m
beam.add_loads(load_1,load_2,load_3) # Assign the support objects to a beam object created earlier
beam.analyse()
fig_1 = beam.plot_beam_external()
fig_1.show()
fig_2 = beam.plot_beam_internal()
fig_2.show()
# -
# ## 3. Support class breakdown
# +
# The parameters for a support class are as below, taken from the docstring
# for the Support class __init__ method.
# Parameters:
# -----------
# coord: float
# x coordinate of support on a beam in m (default 0)
# fixed: tuple of 3 booleans
# Degrees of freedom that are fixed on a beam for movement in
# x, y and bending, 1 represents fixed and 0 represents free
# (default (1,1,1))
# kx :
# stiffness of x support (N/m), if set will overide the
# value placed in the fixed tuple. (default = None)
# ky : (positive number)
# stiffness of y support (N/m), if set will overide the
# value placed in the fixed tuple. (default = None)
# Lets define every possible degree of freedom combination for
# supports below, and view them on a plot:
support_0 = Support(0, (1,1,1)) # conventional fixed support
support_1 = Support(1, (1,1,0)) # conventional pin support
support_2 = Support(2, (1,0,1))
support_3 = Support(3, (0,1,1))
support_4 = Support(4, (0,0,1))
support_5 = Support(5, (0,1,0)) # conventional roller support
support_6 = Support(6, (1,0,0))
# Note we could also explicitly define parameters as follows:
support_0 = Support(coord=0, fixed=(1,1,1))
# Now lets define some spring supports
support_7 = Support(7, (0,0,0), kx = 10) #spring in x direction only
support_8 = Support(8, (0,0,0), ky = 5) # spring in y direction only
support_9 = Support(9, (0,0,0), kx = 100, ky = 100) # spring in x and y direction
# Now lets define a support which is fixed in one degree of freedom
# but has a spring stiffness in another degree of freedom
support_10 = Support(10, (0,1,0), kx = 10) #spring in x direction, fixed in y direction
support_11 = Support(11, (0,1,1), kx = 10) #spring in x direction, fixed in y and m direction
# Note we could also do the following for the same result since the spring
# stiffness overides the fixed boolean in respective directions
support_10 = Support(10, (1,1,0), kx =10)
# Now lets plot all the supports we have created
beam = Beam(11)
beam.add_supports(
support_0,
support_1,
support_2,
support_3,
support_4,
support_5,
support_6,
support_7,
support_8,
support_9,
support_10,
support_11,
)
fig = beam.plot_beam_diagram()
fig.show()
# -
# ## 4. Load classes breakdown
# ### 4.1 Point Torque
# +
# defined using a force (technically a moment, however force is used to maintain consistenct for all load classes) and a coordinate. An anti-clockwise moment is positive by convention of this package.
load_1 = PointTorque(force=1000, coord=1)
load_2 = PointTorque(force=-1000, coord=2)
# Plotting the loads
beam = Beam(3)
beam.add_loads(
load_1,
load_2
)
fig = beam.plot_beam_diagram()
fig.show()
# -
# ### 4.2 Point Load
# +
# defined by force, coord and angle (0)
load_1 = PointLoad(force=1000, coord=1, angle=0)
load_2 = PointLoad(force=1000, coord=2, angle=45)
load_3 = PointLoad(force=1000, coord=3, angle=90)
# Plotting the loads
beam = Beam(4)
beam.add_loads(
load_1,
load_2,
load_3
)
fig = beam.plot_beam_diagram()
fig.show()
# -
# ### 4.3 Uniformly Distributed Load (UDL)
# +
# defined by force, span (tuple with start and end point)
# and angle of force
load_1 = UDL(force=1000, span=(1,2), angle = 0)
load_2 = UDL(force=1000, span=(3,4), angle = 45)
load_3 = UDL(force=1000, span=(5,6), angle = 90)
# Plotting the loads
beam = Beam(7)
beam.add_loads(
load_1,
load_2,
load_3
)
fig = beam.plot_beam_diagram()
fig.show()
# -
# ### 4.4 Trapezoidal Load
# +
# defined by force (tuple with start and end force),
# span (tuple with start and end point) and angle of force
load_1 = TrapezoidalLoad(force=(1000,2000), span=(1,2), angle = 0)
load_2 = TrapezoidalLoad(force=(-1000,-2000), span=(3,4), angle = 45)
load_3 = TrapezoidalLoad(force=(-1000,2000), span=(5,6), angle = 90)
# Plotting the loads
beam = Beam(7)
beam.add_loads(
load_1,
load_2,
load_3
)
fig = beam.plot_beam_diagram()
fig.show()
# -
# ### 4.5 Distributed Load
# +
# defined with Sympy expression of the distributed load function
# expressed using variable x which represents the beam x-coordinate.
# Requires quotation marks around expression. As with the UDL and
# Trapezoidal load classes other parameters to express are the span
# (tuple with start and end point) and angle of force.
# NOTE: where UDL or Trapezoidal load classes can be spefied (linear functions)
# they should be used for quicker analysis times.
load_1 = DistributedLoad(expr= "2", span=(1,2), angle = 0)
load_2 = DistributedLoad(expr= "2*(x-6)**2 -5", span=(3,4), angle = 45)
load_3 = DistributedLoad(expr= "cos(5*x)", span=(5,6), angle = 90)
# Plotting the loads
beam = Beam(7)
beam.add_loads(
load_1,
load_2,
load_3
)
fig = beam.plot_beam_diagram()
fig.show()
# -
# ### 4.6 Vertical and Horizontal load child classes
# +
# for all loads except the point torque an angle is specified for the
# direction of the load. If the load to be specified is to be completely
# vertical or completely horizontal a V (vertical) or a H (horizontal)
# can be added at the end of the class name, and the angle does then
# not need to be spefied.
# The following two loads are equivalent horizontal loads
load_1 = PointLoad(force=1000, coord=1, angle = 0)
load_2 = PointLoadH(force=1000, coord=2)
# The following two loads are equivalent vertical loads
load_3 = PointLoad(force=1000, coord=3, angle = 90)
load_4 = PointLoadV(force=1000, coord=4)
# The following two loads are also equivalent (a negative sign
# esentially changes the load direction by 180 degrees).
load_5 = PointLoad(force=1000, coord=5, angle = 0)
load_6 = PointLoad(force=-1000, coord=6, angle = 180)
# Plotting the loads
beam = Beam(7)
beam.add_loads(
load_1,
load_2,
load_3,
load_4,
load_5,
load_6
)
fig = beam.plot_beam_diagram()
fig.show()
# -
# ## 5. STATICALLY DETERMINATE BEAM (Ex 12.14 Hibbeler)
# +
# Statically Determinate beam (Ex 12.14 Hibbeler)
# Determine the displacement at x = 8m for the following structure
# 8 m long fixed at A (x = 0m)
# A trapezoidal load of - 4000 N/m at x = 0 m descending to 0 N/m at x = 6 m.
beam = Beam(8, E=1, I = 1) ##EI Defined to be 1 get the deflection as a function of EI
a = Support(0, (1,1,1)) ##explicitly stated although this is equivalent to Support() as the defaults are for a cantilever on the left of the beam.
load_1 = TrapezoidalLoadV((-4000,0),(0,6))
beam.add_supports(a)
beam.add_loads(load_1)
beam.analyse()
print(f"Deflection is {beam.get_deflection(8)} N.m3 / EI (N.mm2)")
fig = beam.plot_beam_internal()
fig.show()
# Note: all plots are correct, deflection graph shape is correct but for actual deflection values will need real EI properties.
##save the results as a pdf (optional)
# Can save figure using `fig.write_image("./results.pdf")` (can change extension to be
# png, jpg, svg or other formats as reired). Requires pip install -U kaleido
# -
# ## 6. STATICALLY INDETERMINATE BEAM (Ex 12.21 Hibbeler)
# +
# Statically Indeterminate beam (Ex 12.21 Hibbeler)
# Determine the reactions at the roller support B of the beam described below:
# 3 m long, fixed at A (x = 0 m), roller support at B (x=3 m),
# vertical point load at midpan of 8000 N, UDL of 6000 N/m, EI constant.
beam = Beam(3)
a = Support(0,(1,1,1))
b = Support(3,(0,1,0))
load_1 = PointLoadV(-8000,1.5)
load_2 = UDLV(-6000, (0,3))
beam.add_supports(a,b)
beam.add_loads(load_1,load_2)
beam.analyse()
print(f"The beam has an absolute maximum shear force of: {beam.get_shear_force(return_absmax=True)} N")
print(f"The beam has an absolute maximum bending moment of: {beam.get_bending_moment(return_absmax=True)} N.mm")
print(f"The beam has a vertical reaction at B of: {beam.get_reaction(3,'y')} N")
fig1 = beam.plot_beam_external()
fig1.show()
fig2 = beam.plot_beam_internal()
fig2.show()
# fig1.write_image("./example_1_external.png")
# fig2.write_image("./example_1_internal.png")
# -
# ## 7. SPRING SUPPORTED BEAM (Ex 12.16 Hibbeler)
# +
# Spring Supported beam (Ex 12.16 Hibbeler)
# Determine the vertical displacement at x = 1 m for the beam detailed below:
# 3 m long, spring of ky = 45 kN/m at A (x = 0 m) and B (x = 3 m), vertical point load at x = 1 m of 3000 N, E = 200 GPa, I = 4.6875*10**-6 m4.
# when initializing beam we should specify E and I. Units should be expressed in MPa (N/mm2) for E, and mm4 for I
beam = Beam(3, E=(200)*10**3, I=(4.6875*10**-6)*10**12)
# creating supports, note that an x support must be specified even when there are no x forces. This will not affect the accuracy or reliability of results.
# Also note that ky units are kN/m in the problem but must be in N/m for the program to work correctly.
a = Support(0, (1,1,0), ky = 45000)
b = Support(3, (0,1,0), ky = 45000)
load_1 = PointLoadV(-3000, 1)
beam.add_supports(a,b)
beam.add_loads(load_1)
beam.analyse()
beam.get_deflection(1)
fig1 = beam.plot_beam_external()
fig1.show()
fig2 = beam.plot_beam_internal()
fig2.show()
##results in 38.46 mm deflection ~= 38.4mm specified in textbook (difference only due to their rounding)
##can easily check reliability of answer by looking at deflection at the spring supports. Should equal F/k.
## ie at support A (x = 0 m), the reaction force is 2kN by equilibrium, so our deflection is F/K = 2kn / 45*10-3 kN/mm = 44.4 mm (can be seen in plot)
# -
# ## 8. AXIAL LOADED INDETERMINATE BEAM (Ex 4.13 Hibbeler)
# +
##AXIAL LOADED INDETERMINATE BEAM (Ex 4.13 Hibbeler)
## A rod with constant EA has a force of 60kN applied at x = 0.1 m, and the beam has fixed supports at x=0, and x =0.4 m. Determine the reaction forces.
beam = Beam(0.4)
a = Support()
b = Support(0.4)
load_1 = PointLoadH(-60000, 0.1)
beam.add_supports(a,b)
beam.add_loads(load_1)
beam.analyse()
beam.plot_normal_force()
| docs/examples/.ipynb_checkpoints/simple_demo-Copy1-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + deletable=true editable=true
# 8 --- N(5.73, 0.28)
# + deletable=true editable=true
import numpy as np
from scipy.stats.mstats import normaltest
import matplotlib.pyplot as plt
# %matplotlib inline
# + deletable=true editable=true
mu, sigma = 5.73, np.sqrt(0.28)
# + [markdown] deletable=true editable=true
# Сгенерируем $10^4$ выборок размера $10^4$ и посмотрим на сходимость $T_i$ - для этого изобразим 10 из последовательностей $T_i$
# + deletable=true editable=true
n = int(1e4)
plt.figure(figsize=(15, 10))
Ts = []
for k in xrange(int(1e4)):
X = np.random.normal(mu, sigma, n)
sums = np.cumsum(X)
rng = np.arange(1, sums.shape[0] + 1)
T = (sums - rng * mu) / np.sqrt(rng) / sigma
Ts.append(T)
if k < 10:
plt.plot(T)
Ts = np.array(Ts)
plt.show()
# -
# Видно, что ни при одной из реализаций сходимости нет в силу ЦПТ.
# Проверим, что ЦПТ выполняется. Для этого построим гистограмму значений $T_n$ ($T_n$ отличается от случ. величины из ЦПТ на константный множитель)
plt.hist(Ts[:,-1], bins=30);
# Очень похоже на нормальное, что и требовалось доказать.
normaltest(Ts[:,-1])
# Также гипотеза о нормальности $T_n$ не отвергается с уровнем значимости 0.31
| 01_statistics_and_estimators/8.1.5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PaddlePaddle 2.0.0b0 (Python 3.5)
# language: python
# name: py35-paddle1.2.0
# ---
# # 用N-Gram模型在莎士比亚文集中训练word embedding
#
# **作者:** [PaddlePaddle](https://github.com/PaddlePaddle) <br>
# **日期:** 2021.05 <br>
# **摘要:**
# N-gram 是计算机语言学和概率论范畴内的概念,是指给定的一段文本中N个项目的序列。N=1 时 N-gram 又称为 unigram,N=2 称为 bigram,N=3 称为 trigram,以此类推。实际应用通常采用 bigram 和 trigram 进行计算。本示例在莎士比亚文集上实现了trigram。
# ## 一、环境配置
#
# 本教程基于Paddle 2.1 编写,如果你的环境不是本版本,请先参考官网[安装](https://www.paddlepaddle.org.cn/install/quick) Paddle 2.1 。
import paddle
paddle.__version__
# ## 二、数据集&&相关参数
# ### 2.1 数据集下载
# 训练数据集采用了莎士比亚文集,[点击下载](https://ocw.mit.edu/ans7870/6/6.006/s08/lecturenotes/files/t8.shakespeare.txt)后,保存为txt格式即可。<br>
# context_size设为2,意味着是trigram。embedding_dim设为256。
# !wget https://ocw.mit.edu/ans7870/6/6.006/s08/lecturenotes/files/t8.shakespeare.txt
# +
# 文件路径
path_to_file = './t8.shakespeare.txt'
test_sentence = open(path_to_file, 'rb').read().decode(encoding='utf-8')
# 文本长度是指文本中的字符个数
print ('Length of text: {} characters'.format(len(test_sentence)))
# -
# ### 2.2 数据预处理
# 因为标点符号本身无实际意义,用`string`库中的punctuation,完成英文符号的替换。
# +
from string import punctuation
process_dicts={i:'' for i in punctuation}
print(process_dicts)
punc_table = str.maketrans(process_dicts)
test_sentence = test_sentence.translate(punc_table)
# -
# 由于词表的的长尾,会降低模型训练的速度与精度。因此取词频前2500的单词作为词表,如果不在词表中的单词都用 '<pad>' 替换。
# +
test_sentence_list = test_sentence.lower().split()
word_dict_count = {}
for word in test_sentence_list:
word_dict_count[word] = word_dict_count.get(word, 0) + 1
word_list = []
soted_word_list = sorted(word_dict_count.items(), key=lambda x: x[1], reverse=True)
for key in soted_word_list:
word_list.append(key[0])
word_list = word_list[:2500]
print(len(word_list))
# -
# ### 2.3 模型参数设置
# 设置模型训练常用的参数。
# 设置参数
hidden_size = 1024 # Linear层 参数
embedding_dim = 256 # embedding 维度
batch_size = 256 # batch size 大小
context_size = 2 # 上下文长度
vocab_size = len(word_list) + 1 # 词表大小
epochs = 2 # 迭代轮数
# ## 三、数据加载
# ### 3.1 数据格式
# 将文本被拆成了元组的形式,格式为(('第一个词', '第二个词'), '第三个词');其中,第三个词就是目标。
# +
trigram = [[[test_sentence_list[i], test_sentence_list[i + 1]], test_sentence_list[i + 2]]
for i in range(len(test_sentence_list) - 2)]
word_to_idx = {word: i+1 for i, word in enumerate(word_list)}
word_to_idx['<pad>'] = 0
idx_to_word = {word_to_idx[word]: word for word in word_to_idx}
# 看一下数据集
print(trigram[:3])
# -
# ### 3.2 构建`Dataset`类 加载数据
# 用`paddle.io.Dataset`构建数据集,然后作为参数传入到`paddle.io.DataLoader`,完成数据集的加载。
# +
import numpy as np
class TrainDataset(paddle.io.Dataset):
def __init__(self, tuple_data):
self.tuple_data = tuple_data
def __getitem__(self, idx):
data = self.tuple_data[idx][0]
label = self.tuple_data[idx][1]
data = np.array(list(map(lambda word: word_to_idx.get(word, 0), data)))
label = np.array(word_to_idx.get(label, 0))
return data, label
def __len__(self):
return len(self.tuple_data)
train_dataset = TrainDataset(trigram)
# 加载数据
train_loader = paddle.io.DataLoader(train_dataset, return_list=True, shuffle=True,
batch_size=batch_size, drop_last=True)
# -
# ## 四、模型组网
# 这里用paddle动态图的方式组网。为了构建Trigram模型,用一层 `Embedding` 与两层 `Linear` 完成构建。`Embedding` 层对输入的前两个单词embedding,然后输入到后面的两个`Linear`层中,完成特征提取。
# +
import paddle.nn.functional as F
class NGramModel(paddle.nn.Layer):
def __init__(self, vocab_size, embedding_dim, context_size):
super(NGramModel, self).__init__()
self.embedding = paddle.nn.Embedding(num_embeddings=vocab_size, embedding_dim=embedding_dim)
self.linear1 = paddle.nn.Linear(context_size * embedding_dim, hidden_size)
self.linear2 = paddle.nn.Linear(hidden_size, vocab_size)
def forward(self, x):
x = self.embedding(x)
x = paddle.reshape(x, [-1, context_size * embedding_dim])
x = self.linear1(x)
x = F.relu(x)
x = self.linear2(x)
return x
# -
# ## 五、 方式1:基于高层API,完成模型的训练与预测
# ### 5.1 自定义Callback
# 在训练过程中,有时需要根据模型训练过程中loss,打印loss下降曲线来调参。为了保存训练时每个batch的loss信息,需要自己定义Callback函数,完成模型训练时loss信息的记录。具体的方式如下:
# +
# 自定义Callback 需要继承基类 Callback
class LossCallback(paddle.callbacks.Callback):
def __init__(self):
self.losses = []
def on_train_begin(self, logs={}):
# 在fit前 初始化losses,用于保存每个batch的loss结果
self.losses = []
def on_train_batch_end(self, step, logs={}):
# 每个batch训练完成后调用,把当前loss添加到losses中
self.losses.append(logs.get('loss'))
loss_log = LossCallback()
# -
# ### 5.2 模型训练
# 完成组网与自定义Callback后,将模型用` Model` 封装后,就可以用 `Model.prepare()、Model.fit()` 开始训练。
# +
n_gram_model = paddle.Model(NGramModel(vocab_size, embedding_dim, context_size)) # 用 Model封装 NGramModel
# 模型配置
n_gram_model.prepare(optimizer=paddle.optimizer.Adam(learning_rate=0.01,
parameters=n_gram_model.parameters()),
loss=paddle.nn.CrossEntropyLoss())
# 模型训练
n_gram_model.fit(train_loader,
epochs=epochs,
batch_size=batch_size,
callbacks=[loss_log],
verbose=1)
# -
# ### 5.3 loss可视化
# 利用 `matplotlib` 工具,完成loss的可视化
# 可视化 loss
import matplotlib.pyplot as plt
import matplotlib.ticker as ticker
# %matplotlib inline
log_loss = [loss_log.losses[i] for i in range(0, len(loss_log.losses), 500)]
plt.figure()
plt.plot(log_loss)
# ## 六、方式2:基于基础API,完成模型的训练与预测
# ### 6.1 自定义 `train` 函数
# 通过基础API,自定义 `train` 函数,完成模型的训练。
import paddle.nn.functional as F
losses = []
def train(model):
model.train()
optim = paddle.optimizer.Adam(learning_rate=0.01, parameters=model.parameters())
for epoch in range(epochs):
for batch_id, data in enumerate(train_loader()):
x_data = data[0]
y_data = data[1]
predicts = model(x_data)
loss = F.cross_entropy(predicts, y_data)
loss.backward()
if batch_id % 500 == 0:
losses.append(loss.numpy())
print("epoch: {}, batch_id: {}, loss is: {}".format(epoch, batch_id, loss.numpy()))
optim.step()
optim.clear_grad()
model = NGramModel(vocab_size, embedding_dim, context_size)
train(model)
# ### 6.2 loss可视化
# 通过可视化loss的曲线,可以看到模型训练的效果。
# +
import matplotlib.pyplot as plt
import matplotlib.ticker as ticker
# %matplotlib inline
plt.figure()
plt.plot(losses)
# -
# ### 6.3 预测
# 用训练好的模型进行预测。
import random
def test(model):
model.eval()
# 从最后10组数据中随机选取1个
idx = random.randint(len(trigram)-10, len(trigram)-1)
print('the input words is: ' + trigram[idx][0][0] + ', ' + trigram[idx][0][1])
x_data = list(map(lambda word: word_to_idx.get(word, 0), trigram[idx][0]))
x_data = paddle.to_tensor(np.array(x_data))
predicts = model(x_data)
predicts = predicts.numpy().tolist()[0]
predicts = predicts.index(max(predicts))
print('the predict words is: ' + idx_to_word[predicts])
y_data = trigram[idx][1]
print('the true words is: ' + y_data)
test(model)
| docs/practices/n_gram_model/n_gram_model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="http://image.yes24.com/goods/64625506/800x0" width="200" height="200"><br>
# # 11 - 1 데이터 집계
#
# ### groupby 메서드로 평균값 구하기
# 1.먼저 갭마인더 데이터 집합을 불러옵니다.
import pandas as pd
df = pd.read_csv('data/gapminder.tsv', sep='\t')
# 2.다음은 year 열을 기준으로 데이터를 그룹화한 다음 lifeExp 열의 평균을 구한 것입니다.
avg_life_exp_by_year = df.groupby('year').lifeExp.mean()
print(avg_life_exp_by_year)
# ### 분할 - 반영 - 결합 과정 살펴보기 -groupby 메서드
#
# 앞에서 groupby 메서드를 사용해 lifeExp 열의 연도별 평균값을 구했습니다. 그러면 실제로 groupby 메서드는 어떤 과정을 통해 데이터 집계 할까요? groupby 메서드 자체를 분해하여 살펴보는 것은 불가능하기 때문에 비슷한 연산을 수행하는 메서드를 순서대로 실행하면 알아보자
#
# #### 분할 - 반영 - 결합 과정 살펴보기
# 1.실제로 groupby 메서드에 life 열을 전달하면 가장 먼저 연도별로 데이터를 나누는 과정이 진행됩니다. 다음은 year 열의 데이터를 중복 없이 추출한 것입니다. groupby 메서드에 열 이름을 전달하면 이런 '분할'작업이 먼저 일어난다고 이해하세요.
years = df.year.unique()
print(years)
# 2.그런 다음에는 연도별로 평균값을 구합니다. 그러려면 일단 각 연도별로 데이터를 추출해야겠죠? 다음은 1952년의 데이터를 추출한 것입니다. 이과정을 '반영'작업의 한 부분이라고 이해하세요.
y1952 = df.loc[df.year == 1952, :]
print(y1952.head())
# 3.아직 lifeExp 열의 평균값을 구하지 않았습니다. 다음은 과정 2에서 추출한 1952년의 데이터에서 lifeExp 열의 평균값을 구한 것입니다. 이과정도 '반영'작업의 한 부분입니다.
#
y1952_mean = y1952.lifeExp.mean()
print(y1952_mean)
# 4.과정 2~3을 반복하여 남은 연도의 평균값을 구하면 비로소 '반영' 작업이 끝납니다.
# +
y1957 = df.loc[df.year == 1957, :]
y1957_mean = y1957.lifeExp.mean( )
print(y1957_mean)
y1962 = df.loc[df.year == 1962, :]
y1962_mean = y1962.lifeExp.mean( )
print(y1962_mean)
y2007 = df.loc[df.year == 2007, :]
y2007_mean = y2007.lifeExp.mean( )
print(y2007_mean)
# -
# 5.마지막으로 연도별로 계산한 lifeExp의 평균값을 합칩니다. 바로 이 과정을 '결합'작업 입니다.
df2 = pd.DataFrame({"year":[1952, 1957, 1962, 2007],
"":[y1952_mean, y1957_mean, y1962_mean, y2007_mean]})
print(df2)
# ### groupby 메서드와 함께 사용하는 집계 메서드
#
#
# #### 집계 메서드
#
# 메서드 | 설명
# --- | ---
# count | 누락값을 제외한 데이터 수를 반환
# size | 누락값을 포함한 데이터 수를 반환
# mean | 평균값 반환
# std | 표준편차 반환
# min | 최솟값 반환
# quantile(q=0.25) | 백분위수 25%
# quantile(q=0.50) | 백분위수 50%
# quantile(q=0.75) | 백분위수 75%
# max | 최댓값 반환
# sum | 전체 합 반환
# var | 분산 반환
# sem | 평균의 표준편차 반환
# describe | 데이터 수, 평균, 표준편차, 최소값, 백분위수(25,50,75%),최댓값을 모두 반환
# first | 첫 번째 행 반환
# last | 마지막 행 반환
# nth | n번째 행 반환
# ### agg 메서드로 사용자 함수와 groupby 메서드 조합하기
#
# 라이브러리에서 제공하는 집계 메서드로 원하는 값을 계산할 수 없는 경우에는 직접 함수를 만들어서 사용해야 합니다. 이번에는 사용자 함수와 groupby 메서드를 조합해서 사용해 보겠습니다. 사용자 함수와 groupby 메서드를 조합하려면 agg 메서드를 이용해야 합니다.
#
# ### 평균값을 구하는 사용자 함수와 groupby 메서드
# 1.다음은 입력받은 열의 평균값을 구하는 함수입니다.
def my_mean(values):
n = len(values)
sum = 0
for value in values:
sum += value
return sum / n
agg_my_mean = df.groupby('year').lifeExp.agg(my_mean)
print(agg_my_mean)
# ### 2개의 인잣값을 받아 처리하는 사용자 함수와 groupby 메서드
#
# 1.이번에는 2개의 인잣값을 받아 처리하는 사용자 정의 함수(my_mean_diff)를 만들어보겠습니다. 다음은 첫 번째 인자로 받은 열의 평균값을 구하여 두 번째 인자로 받은 값과의 차이를 계산한 다음 반환하는 함수입니다.
def my_mean_diff(values, diff_value):
n = len(values)
sum = 0
for value in values:
sum += value
mean = sum / n
return mean - diff_value
# 2.다음은 연도별 평균 수명에서 전체 평균 수명을 뺀 값을 구한 것입니다. agg 메서드의 첫 번째 인자에 my_mean_diff 함수를 전달하고 두 번째 인자에 전체 평균 수명값을 전달 했습니다.
global_mean = df.lifeExp.mean()
print(global_mean)
agg_mean_diff = df.groupby('year').lifeExp.agg(my_mean_diff, diff_value=global_mean)
print(agg_mean_diff)
# ### 집계 메서드를 리스트, 딕셔너리에 담아 전달하기
# 1.다음은 연도별로 그룹화한 lifeExp 열의 0이아닌 값의 개수, 평균, 표준편차를 한 번에 계산하여 출력한 것입니다. 넘파이 메서드인 count_nonzero, mean, std를 리스트에 담아 agg 메서드에 전달했습니다.
import numpy as np
gdf = df.groupby('year').lifeExp.agg([np.count_nonzero, np.mean, np.std])
print(gdf)
# 2.이번에는 집계 메서드를 딕셔너리에 담아 agg 메서드에 전달해 보겠습니다. 딕셔너리의 키로 집계 메서드를 적용할 열 이름을 전달하고 딕셔너리의 값으로 집계 메서드를 전달하면 됩니다.
gdf_dict = df.groupby('year').agg({ 'lifeExp': 'mean', 'pop': 'median', 'gdpPercap': 'median'})
print(gdf_dict)
# # 11 - 2 데이터 변환
#
# ### 표준점수 계산하기
# 통계 분야에서는 데이터 평균과 표준편차의 차이를 표준점수라고 부릅니다. 표준점수를 구하면 변환한 데이터의 평균값이 0이 되고 표준편차는 1이됩니다. 그러면 데이터가 표준화되어 서로 다른 데이터를 쉽게 비교할 수 있게되죠. 표준점수는 통계에서 자주 사용하는 지표입니다.
#
# #### 표준점수 계산하기
# 1.다음은 표준점수를 계산하는 함수입니다.
def my_zscore(x):
return (x - x.mean()) / x.std()
# 2.다음은 각 연도별 lifeExp 열의 표준점수를 계산한 것입니다. my_zscore 함수를 적용하기 위해 transform 메서드를 사용했습니다.
# +
transform_z = df.groupby('year').lifeExp.transform(my_zscore)
print(transform_z.head())
# -
# 3.my_zscore 함수는 데이터를 표준화할 뿐 집계는 하지 않습니다. 즉, 데이터의 양이 줄어들지 않습니다. 다음은 원본 데이터프레임(df)의 데이터 크기와 변환한 데이터프레임(transform_z)의 데이터 크기를 비교한 것입니다.
print(df.shape)
print(transform_z.shape)
# ### 누락값을 평균값으로 처리하기
#
# 1.다음은 seaborn 라이브러리의 tips 데이터 집합에서 10개의 행 데이터만 가져온 다음 total_bill 열의 값 4개를 임의로 선택하여 누락값으로 바꾼 것입니다.
# +
import seaborn as sns
import numpy as np
np.random.seed(42)
# -
tips_10 = sns.load_dataset('tips').sample(10)
tips_10.loc[np.random.permutation(tips_10.index)[:4], 'total_bill'] = np.NaN
print(tips_10)
# 2.그런데 total_bill 열의 누락값을 단순히 total_bill 열의 평균값으로 채우면 안됩니다. 무슨 말 일까요? 현재 tips_10의 데이터는 여성보다 남성이 더 많습니다. 즉, 여성과 남성을 구분하여 total_bill 열의 평균값을 구하지 않으면 여성 데이터가 남성 데이터의 영향을 많이 받아 여성의 데이터가 훼손될 수 있습니다. 다음은 성별로 그룹화한 다음 각 열의 데이터 수를 구한 것입니다. total_bill 열을 살펴보면 남성의 누락값은 3개, 여성의 누락값은 1개라는 것을 알 수 있습니다.
count_sex = tips_10.groupby('sex').count()
print(count_sex)
# 3.다음은 성별을 구분하여 total_bill 열의 데이터를 받아 평균값을 구하는 함수입니다.
def fill_na_mean(x):
avg = x.mean()
return x.fillna(avg)
# 4.다음은 성별을 구분한 totall_bill 열의 데이터를 fill_na_mean 함수에 전달하여 평균값을 구한 다음 tips_10에 새로운 열로 추가한 것입니다. 남성과 여성의 누락값을 고려하여 계산한 평균값으로 잘 채워져 있는 것을 알 수 있습니다.
total_bill_group_mean = tips_10.groupby('sex').total_bill.transform(fill_na_mean)
tips_10['fill_total_bill'] = total_bill_group_mean
print(tips_10)
# # 11 - 3 데이터 필터링
#
# 만약 그룹화한 데이터에서 원하는 데이터를 걸러내고 싶다면 어떻게 해야 할까? 그럴 때는 데이터 필터링을 사용하면 됩니다. 데이터 필터링을 사용하면 기준에 맞는 데이터를 걸러낼 수 있습니다.
#
# ### 데이터 필터링 사용하기 - filter 메서드
# 1.다음과 같이 tips 데이터 집합을 불러와 데이터 크기를 확인합니다.
tips = sns.load_dataset('tips')
print(tips.shape)
# 2.size 열의 데이터 수를 확인해 보면 1, 5, 6테이블의 주문이 매우 적다는 것을 알 수 있습니다.
print(tips['size'].value_counts())
# 3.상황에 따라 이런 데이터는 제외하기도 합니다. 만약 30번 이상의 주문이 있는 테이블만 추려 데이터 분석을 하려면 어떻게 해야 할까? 다음은 30번 이상의 주문이 있는 테이블만 그룹화하여 변수 tips_filtered에 저장한 것입니다.
tips_filtered = tips.\
groupby('size').\
filter(lambda x: x['size'].count() >= 30)
# 4.과정 3을 거치고 나면 1,5,6 테이블의 데이터가 제외되었다는 것을 알 수 있습니다.
print(tips_filtered.shape)
print(tips_filtered['size'].value_counts())
# # 11 - 4 그룹 오브젝트
#
# ### 그룹 오브젝트 저장하여 살펴보기
# 1.다음은 tips 데이터 집합에서 임의로 10개의 데이터를 추출한 것입니다.
tips_10 = sns.load_dataset('tips').sample(10, random_state=42)
print(tips_10)
# 2.groupby 메서드의 결괏값을 출력하면 자료형이 그룹 오브젝트라는 것을 확인할 수 있습니다.
grouped = tips_10.groupby('sex')
print(grouped)
# 3.그룹 오브젝트에 포함된 그룹을 보려면 groups 속성을 출력하면 됩니다. 그러면 sex 열로 그룹화한 데이터프레임의 인덱스를 확인할 수 있습니다. 이 그룹 오브젝트로 집계, 변환, 필터 작업을 수행하면 되는 것이죠.
print(grouped.groups)
# # 한 번에 그룹 오브젝트 계산하기
#
# ### 그룹 오브젝트의 평균 구하기
# 1.일단 앞에서 만든 그룹 오브젝트(grouped)를 이용하여 평균을 구해 보겠습니다. 그러면 tips 데이터 집합의 모든 열의 평균을 구한 것이 아니라 total_bill, tip, size 열의 평균을 구했다는 것을 알 수 있습니다.
avgs = grouped.mean()
print(avgs)
# 2.tips 데이터 집합의 열을 확인해 보면 평균값을 계산할 수 없는 열인 smoker, day, time 열은 그룹 연산에서 제외되었다는 것을 알 수 있습니다. 이처럼 파이썬은 그룹 연산에 적합한 열을 알아서 골라줍니다.
print(tips_10.columns)
# ## 그룹 오브젝트 활용하기
#
#
# 그룹 오브젝트를 활용하는 방법은 아주 많습니다. 그룹 오브젝트에서 '데이터 추출하기'와 '반복문 사용하기'를 실습을 해봅시다.
#
# ### 그룹 오브젝트에서 데이터 추출하고 반복하기
# 1.만약 그룹 오브젝트에서 특정 데이터만 추출하려면 get_group 메서드를 사용하면 됩니다. 다음은 sex 열로 그룹화한 그룹 오브젝트에 get_group 메서드를 사용하여 성별이 여성인 데이터만 추출한 것입니다.
female = grouped.get_group('Female')
print(female)
# 2.이번에는 그룹 오브젝트를 반복문에 사용해 보겠습니다. sex열을 기준으로 그룹화한 tips 데이터 집합은 여성 그룹과 남성 그룹으로 나누어져 있습니다. 이 특징을 이용하여 반복문을 사용하면 됩니다. 다음은 각 성별 그룹의 데이터를 반복문을 이용하여 출력한 것입니다.
for sex_group in grouped:
print(sex_group)
# 3.그런데 과정 2의 결과를 자세히 살펴보면 sex_group으로 넘어온 값이 튜플이라는 것을 알 수 있습니다. 다음은 sex_group의 자세한 정보를 출력한 것입니다.
for sex_group in grouped:
print('the type is: {}\n'.format(type(sex_group)))
print('the length is: {}\n'.format(len(sex_group)))
first_element = sex_group[0]
print('the first element is: {}\n'.format(first_element))
print('it has a type of: {}\n'.format(type(sex_group[0])))
second_element = sex_group[1]
print('the second element is:\n{}\n'.format(second_element))
print('it has a type of: {}\n'.format(type(second_element)))
print('what we have:')
print(sex_group)
break
# ## 여러 열을 사용해 그룹 오브젝트 만들고 계산하기
#
# 지금까지는 하나의 열을 사용하여 그룹 오브젝트를 만들고 연산을 수행했습니다. 하지만 여러 열을 사용하여 그룹 오브젝트를 맏들고 평균값을 구하는 등의 계산도 할 수 있습니다.
#
# ### 그룹 오브젝트 계산하고 살펴보기
# 1.여러 열을 사용하여 데이터를 그룹화하려면 리스트에 열 이름을 담아 groupby 메서드에 전달하면 됩니다. 다음은 sex, time 열을 기준으로 데이터를 그룹화하고 평균값을 구한것입니다.
# +
bill_sex_time = tips_10.groupby(['sex', 'time'])
group_avg = bill_sex_time.mean()
print(group_avg)
# -
# 2.과정 1을 거친 group_avg의 자료형을 확인해 보면 데이터프레임이라는 것을 알 수 있습니다. 그리고 변수 group_avg에 포함된 열은 total_bill, tip, size라는 것도 알 수 있습니다.
print(type(group_avg))
print(group_avg.columns)
# 3.group_avg의 자료형은 데이터프레임이라고 했습니다. 그러면 인덱스는 어떻게 구성되어 있을까요? 다음은 group_avg의 인덱스를 출력한 것입니다.
print(group_avg.index)
# 4.과정 3과 같이 데이터프레임의 인덱스가 MultiIndex인 경우에는 reset_index 메서드를 사용하여 데이터프레임의 인덱스를 새로 부여할 수도 있습니다.
group_method = tips_10.groupby(['sex', 'time']).mean().reset_index()
print(group_method)
# 5.reset_index 메서드 대신 as_index 인자를 False로 설정해도 과정 4와 같은 결과를 얻을 수 있습니다.
group_param = tips_10.groupby(['sex', 'time'], as_index=False).mean()
print(group_param)
# ### 마무리하며
#
# 이번에는 groupby 메서드의 분리 - 반영 = 결합 과정을 구체적으로 살펴보고 데이터를 집계, 변환, 필터링하는 방법을 알아보았습니다.
# 또 여러 가지 방법으로 그룹 오브젝트를 만들고 계산하는 방법도 알아보았습니다. 데이터를 그룹화하여 계산하는 작업은 실무에서 자주 사용하므로 반드시 알아두어야 합니다.
| pandas_11(Group_Calculation).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas
import matplotlib as mpl
import xarray as xr
import numpy as np
import datetime as dt
dir_cmc='F:/data/sst/cmc/CMC0.2deg/v2/'
dir_cmc_clim='F:/data/sst/cmc/CMC0.2deg/v2/climatology/'
def get_filename(lyr,idyjl):
d = dt.date(lyr,1,1) + dt.timedelta(idyjl - 1)
dir_cmc='F:/data/sst/cmc/CMC0.2deg/v2/'
syr=str(d.year).zfill(4)
smon=str(d.month).zfill(2)
sdym=str(d.day).zfill(2)
sjdy=str(idyjl).zfill(3)
fname_tem=syr + smon + sdym + '120000-CMC-L4_GHRSST-SSTfnd-CMC0.2deg-GLOB-v02.0-fv02.0.nc'
cmc_filename = dir_cmc + syr + '/' + sjdy + '/' + fname_tem
return cmc_filename
def get_filename_v2a(lyr,idyjl):
d = dt.date(lyr,1,1) + dt.timedelta(idyjl - 1)
dir_cmc='F:/data/sst/cmc/CMC0.2deg/v2/'
syr=str(d.year).zfill(4)
smon=str(d.month).zfill(2)
sdym=str(d.day).zfill(2)
sjdy=str(idyjl).zfill(3)
fname_tem=syr + smon + sdym + '120000-CMC-L4_GHRSST-SSTfnd-CMC0.2deg-GLOB-v02.0-fv03.0.nc'
cmc_filename = dir_cmc + syr + '/' + sjdy + '/' + fname_tem
return cmc_filename
def get_filename_v2a_dir(lyr,idyjl):
d = dt.date(lyr,1,1) + dt.timedelta(idyjl - 1)
dir_cmc='F:/data/sst/cmc/CMC0.2deg/v2/'
syr=str(d.year).zfill(4)
smon=str(d.month).zfill(2)
sdym=str(d.day).zfill(2)
sjdy=str(idyjl).zfill(3)
cmc_filename = dir_cmc + syr + '/' + sjdy + '/'
return cmc_filename
def get_filename_v3(lyr,idyjl):
d = dt.date(lyr,1,1) + dt.timedelta(idyjl - 1)
dir_cmc='F:/data/sst/cmc/CMC0.1deg/v3/'
syr=str(d.year).zfill(4)
smon=str(d.month).zfill(2)
sdym=str(d.day).zfill(2)
sjdy=str(idyjl).zfill(3)
fname_tem=syr + smon + sdym + '120000-CMC-L4_GHRSST-SSTfnd-CMC0.1deg-GLOB-v02.0-fv03.0.nc'
cmc_filename = dir_cmc + syr + '/' + sjdy + '/' + fname_tem
return cmc_filename
#make cmc monthly average ssts
for lyr in range(1992,2017): #2017):
ds_mnth=[]
for imon in range(1,13):
init = 0
for idyjl in range(1,366):
d = dt.date(lyr,1,1) + dt.timedelta(idyjl - 1)
if d.month!=imon:
continue
cmc_filename = get_filename(lyr,idyjl)
print(cmc_filename)
ds = xr.open_dataset(cmc_filename,drop_variables=['analysis_error','sea_ice_fraction'])
ds_masked = ds.where(ds['mask'] == 1.)
ds.close()
ds_masked['sq_sst']=ds_masked.analysed_sst**2
if init==0:
ds_sum = ds_masked
init = 1
else:
ds_sum = xr.concat([ds_sum,ds_masked],dim = 'time')
print(idyjl,ds_sum.dims)
ds_clim2 = ds_sum.resample(time='M').mean()
# ds_clim2 = ds_sum.groupby('time.month').mean('time')
#ds_sum = ds_sum.mean('time',skipna=True)
ds_mnth.append(ds_clim2)
combined = xr.concat(ds_mnth, dim='time')
fname_tem='monthly/' + str(lyr) + 'monthly_average_120000-CMC-L4_GHRSST-SSTfnd-CMC0.2deg-GLOB-v02.0-fv02.0.nc'
cmc_filename_out = dir_cmc + fname_tem
combined.to_netcdf(cmc_filename_out)
# +
#make cmc monthly average ssts FOR 0.1 deg data
#first read in 0.2 deg grid to interpolate onto
#testing 0.1 CMC
filename = 'F:/data/sst/cmc/CMC0.2deg/v2/data/1992/011/19920111120000-CMC-L4_GHRSST-SSTfnd-CMC0.2deg-GLOB-v02.0-fv02.0.nc'
ds_v2 = xr.open_dataset(filename)
new_lat = np.linspace(ds_v2.lat[0], ds_v2.lat[-1], ds_v2.dims['lat'])
new_lon = np.linspace(ds_v2.lon[0], ds_v2.lon[-1], ds_v2.dims['lon'])
for lyr in range(2019,2021): #2017):
ds_mnth=[]
for imon in range(1,13):
init = 0
if (lyr==2020) & (imon>1):
continue
for idyjl in range(1,366):
d = dt.date(lyr,1,1) + dt.timedelta(idyjl - 1)
if d.month!=imon:
continue
cmc_filename = get_filename_v3(lyr,idyjl)
print(cmc_filename)
ds = xr.open_dataset(cmc_filename,drop_variables=['analysis_error','sea_ice_fraction'])
ds_masked = ds.where(ds['mask'] == 1.)
ds_masked['sq_sst']=ds_masked.analysed_sst**2
ds_interp = ds_masked.interp(lat = new_lat,lon = new_lon)
ds.close()
if init==0:
ds_sum = ds_interp
init = 1
else:
ds_sum = xr.concat([ds_sum,ds_interp],dim = 'time')
#print(idyjl,ds_sum.dims)
ds_clim2 = ds_sum.resample(time='M').mean()
# ds_clim2 = ds_sum.groupby('time.month').mean('time')
#ds_sum = ds_sum.mean('time',skipna=True)
ds_mnth.append(ds_clim2)
combined = xr.concat(ds_mnth, dim='time')
fname_tem='monthly/' + str(lyr) + 'monthly_average_120000-CMC-L4_GHRSST-SSTfnd-CMC0.2deg-GLOB-v02.0-fv02.0.nc'
cmc_filename_out = dir_cmc + fname_tem
combined.to_netcdf(cmc_filename_out)
# +
#need to convert 2017 and 2018 to float 32 otherwise they don't combine with other index nicely
#testing data files
file1='F:/data/sst/cmc/CMC0.2deg/v2/monthly/2016monthly_average_120000-CMC-L4_GHRSST-SSTfnd-CMC0.2deg-GLOB-v02.0-fv02.0.nc'
file2='F:/data/sst/cmc/CMC0.2deg/v2/monthly/2017monthly_average_120000-CMC-L4_GHRSST-SSTfnd-CMC0.2deg-GLOB-v02.0-fv02.0.nc'
ds1=xr.open_dataset(file1)
ds2=xr.open_dataset(file2)
ds1.close()
ds2.close()
ds3 = ds2.astype('float32')
ds3['lon']=ds1.lon
ds3['lat']=ds1.lat
file2='F:/data/sst/cmc/CMC0.2deg/v2/monthly/2017monthly_average_120000-CMC-L4_GHRSST-SSTfnd-CMC0.2deg-GLOB-v02.0-fv02.0.nc'
ds3.to_netcdf(file2)
#need to convert 2017 and 2018 to float 32 otherwise they don't combine with other index nicely
#testing data files
file1='F:/data/sst/cmc/CMC0.2deg/v2/monthly/2016monthly_average_120000-CMC-L4_GHRSST-SSTfnd-CMC0.2deg-GLOB-v02.0-fv02.0.nc'
file2='F:/data/sst/cmc/CMC0.2deg/v2/monthly/2018monthly_average_120000-CMC-L4_GHRSST-SSTfnd-CMC0.2deg-GLOB-v02.0-fv02.0.nc'
ds1=xr.open_dataset(file1)
ds2=xr.open_dataset(file2)
ds1.close()
ds2.close()
ds3 = ds2.astype('float32')
ds3['lon']=ds1.lon
ds3['lat']=ds1.lat
file2='F:/data/sst/cmc/CMC0.2deg/v2/monthly/2018monthly_average_120000-CMC-L4_GHRSST-SSTfnd-CMC0.2deg-GLOB-v02.0-fv02.0.nc'
ds3.to_netcdf(file2)
# +
#need to convert 2019 to float 32 otherwise they don't combine with other index nicely
#testing data files
file1='F:/data/sst/cmc/CMC0.2deg/v2/monthly/2016monthly_average_120000-CMC-L4_GHRSST-SSTfnd-CMC0.2deg-GLOB-v02.0-fv02.0.nc'
file2='F:/data/sst/cmc/CMC0.2deg/v2/monthly/2019monthly_average_120000-CMC-L4_GHRSST-SSTfnd-CMC0.2deg-GLOB-v02.0-fv02.0.nc'
ds1=xr.open_dataset(file1)
ds2=xr.open_dataset(file2)
ds1.close()
ds2.close()
ds3 = ds2.astype('float32')
ds3['lon']=ds1.lon
ds3['lat']=ds1.lat
file2='F:/data/sst/cmc/CMC0.2deg/v2/monthly/2019monthly_average_120000-CMC-L4_GHRSST-SSTfnd-CMC0.2deg-GLOB-v02.0-fv02.0.nc'
ds3.to_netcdf(file2)
# -
#need to convert 2020 to float 32 otherwise they don't combine with other index nicely
#testing data files
file1='F:/data/sst/cmc/CMC0.2deg/v2/monthly/2016monthly_average_120000-CMC-L4_GHRSST-SSTfnd-CMC0.2deg-GLOB-v02.0-fv02.0.nc'
file2='F:/data/sst/cmc/CMC0.2deg/v2/monthly/2020monthly_average_120000-CMC-L4_GHRSST-SSTfnd-CMC0.2deg-GLOB-v02.0-fv02.0.nc'
ds1=xr.open_dataset(file1)
ds2=xr.open_dataset(file2)
ds1.close()
ds2.close()
ds3 = ds2.astype('float32')
ds3['lon']=ds1.lon
ds3['lat']=ds1.lat
file2='F:/data/sst/cmc/CMC0.2deg/v2/monthly/2020monthly_average_120000-CMC-L4_GHRSST-SSTfnd-CMC0.2deg-GLOB-v02.0-fv02.0.nc'
ds3.to_netcdf(file2)
#testing data files
file1='F:/data/sst/cmc/CMC0.2deg/v2/monthly/2016monthly_average_120000-CMC-L4_GHRSST-SSTfnd-CMC0.2deg-GLOB-v02.0-fv02.0.nc'
file2='F:/data/sst/cmc/CMC0.2deg/v2/monthly/2019monthly_average_120000-CMC-L4_GHRSST-SSTfnd-CMC0.2deg-GLOB-v02.0-fv02.0.nc'
ds1=xr.open_dataset(file1)
ds2=xr.open_dataset(file2)
ds3 = ds2.astype('float32')
ds3['lon']=ds1.lon
ds3['lat']=ds1.lat
#print(ds3)
ds1.close()
ds2.close()
ds3 = xr.concat([ds3,ds1],dim = 'time')
print(ds3)
#make cmc climatology from monthly files
for icase in range(0,3):
if icase==0:
iyr1,iyr2 = 1992,2000
if icase==1:
iyr1,iyr2 = 2000,2010
if icase==2:
iyr1,iyr2 = 2010,2019
init = 0
for lyr in range(iyr1,iyr2):
fname_tem='monthly/' + str(lyr) + 'monthly_average_120000-CMC-L4_GHRSST-SSTfnd-CMC0.2deg-GLOB-v02.0-fv02.0.nc'
cmc_filename_out = dir_cmc + fname_tem
ds = xr.open_dataset(cmc_filename_out)
ds_masked = ds.where(ds['mask'] == 1.)
if init==0:
ds_sum = ds_masked
init = 1
else:
ds_sum = xr.concat([ds_sum,ds_masked],dim = 'time')
print(lyr,ds_sum.dims)
ds_sum2 = ds_sum.groupby('time.month').mean('time')
fname_tem='monthly/monthly_climatology_'+str(iyr1)+'_'+str(iyr2-1)+'_120000-CMC-L4_GHRSST-SSTfnd-CMC0.2deg-GLOB-v02.0-fv02.0.nc'
cmc_filename_out = dir_cmc + fname_tem
ds_sum2.to_netcdf(cmc_filename_out)
num_year = 2018-1992+1
num_year_file1 = 1999 - 1992 +1
num_year_file2 = 2009 - 2000 +1
num_year_file3 = 2018 - 2010 +1
frac_file1 = num_year_file1 / num_year
frac_file2 = num_year_file2 / num_year
frac_file3 = num_year_file3 / num_year
fname_tem=dir_cmc + 'monthly/monthly_climatology_1992_1999_120000-CMC-L4_GHRSST-SSTfnd-CMC0.2deg-GLOB-v02.0-fv02.0.nc'
print(fname_tem)
ds = xr.open_dataset(fname_tem)
ds_masked = ds.where(ds['mask'] == 1.)
ds.close()
fname_tem=dir_cmc + 'monthly/monthly_climatology_2000_2009_120000-CMC-L4_GHRSST-SSTfnd-CMC0.2deg-GLOB-v02.0-fv02.0.nc'
print(fname_tem)
ds = xr.open_dataset(fname_tem)
ds_masked2 = ds.where(ds['mask'] == 1.)
#ds_sum = xr.concat([ds_masked2,ds_masked],dim = 'time')
ds.close()
fname_tem=dir_cmc + 'monthly/monthly_climatology_2010_2018_120000-CMC-L4_GHRSST-SSTfnd-CMC0.2deg-GLOB-v02.0-fv02.0.nc'
print(fname_tem)
ds = xr.open_dataset(fname_tem)
ds_masked3 = ds.where(ds['mask'] == 1.)
#ds_sum = xr.concat([ds_masked2,ds_masked],dim = 'time')
ds.close()
ds_ave = frac_file1*ds_masked + frac_file2*ds_masked2 + frac_file3*ds_masked3
#ds_sum = ds_sum.mean('time',skipna=True)
print(frac_file1+frac_file2+frac_file3,frac_file1,frac_file2,frac_file3)
fname_tem=dir_cmc + 'monthly/monthly_climatology_1992_2018_120000-CMC-L4_GHRSST-SSTfnd-CMC0.2deg-GLOB-v02.0-fv02.0.nc'
print(fname_tem)
ds_ave.to_netcdf(fname_tem)
num_year = 2018-2000+1
#num_year_file1 = 1999 - 1992 +1
num_year_file2 = 2009 - 2000 +1
num_year_file3 = 2018 - 2010 +1
#frac_file1 = num_year_file1 / num_year
frac_file2 = num_year_file2 / num_year
frac_file3 = num_year_file3 / num_year
#fname_tem=dir_cmc + 'monthly/monthly_climatology_1992_1999_120000-CMC-L4_GHRSST-SSTfnd-CMC0.2deg-GLOB-v02.0-fv02.0.nc'
#print(fname_tem)
#ds = xr.open_dataset(fname_tem)
#ds_masked = ds.where(ds['mask'] == 1.)
#ds.close()
fname_tem=dir_cmc + 'monthly/monthly_climatology_2000_2009_120000-CMC-L4_GHRSST-SSTfnd-CMC0.2deg-GLOB-v02.0-fv02.0.nc'
print(fname_tem)
ds = xr.open_dataset(fname_tem)
ds_masked2 = ds.where(ds['mask'] == 1.)
#ds_sum = xr.concat([ds_masked2,ds_masked],dim = 'time')
ds.close()
fname_tem=dir_cmc + 'monthly/monthly_climatology_2010_2018_120000-CMC-L4_GHRSST-SSTfnd-CMC0.2deg-GLOB-v02.0-fv02.0.nc'
print(fname_tem)
ds = xr.open_dataset(fname_tem)
ds_masked3 = ds.where(ds['mask'] == 1.)
#ds_sum = xr.concat([ds_masked2,ds_masked],dim = 'time')
ds.close()
ds_ave = frac_file2*ds_masked2 + frac_file3*ds_masked3
#ds_sum = ds_sum.mean('time',skipna=True)
print(frac_file2+frac_file3,frac_file2,frac_file3)
fname_tem=dir_cmc + 'monthly/monthly_climatology_2000_2018_120000-CMC-L4_GHRSST-SSTfnd-CMC0.2deg-GLOB-v02.0-fv02.0.nc'
print(fname_tem)
ds_ave.to_netcdf(fname_tem)
# +
#make daily 0.2 deg cmc files
#make cmc monthly average ssts FOR 0.1 deg data
#first read in 0.2 deg grid to interpolate onto
#20170101120000-CMC-L4_GHRSST-SSTfnd-CMC0.2deg-GLOB-v02.0-fv02.0.nc
import os
#testing 0.1 CMC
filename = 'F:/data/sst/cmc/CMC0.2deg/v2/1992/011/19920111120000-CMC-L4_GHRSST-SSTfnd-CMC0.2deg-GLOB-v02.0-fv02.0.nc'
ds_v2 = xr.open_dataset(filename)
new_lat = np.linspace(ds_v2.lat[0], ds_v2.lat[-1], ds_v2.dims['lat'])
new_lon = np.linspace(ds_v2.lon[0], ds_v2.lon[-1], ds_v2.dims['lon'])
for lyr in range(2018,2019): #2017):
for idyjl in range(1,366):
cmc_filename = get_filename_v3(lyr,idyjl)
print(cmc_filename)
ds = xr.open_dataset(cmc_filename,drop_variables=['analysis_error','sea_ice_fraction'])
ds_masked = ds.where(ds['mask'] == 1.)
ds_interp = ds_masked.interp(lat = new_lat,lon = new_lon)
ds.close()
cmc_filename_out = get_filename_v2a(lyr,idyjl)
cmc_dir = get_filename_v2a_dir(lyr,idyjl)
if not os.path.exists(cmc_dir):
os.mkdir(cmc_dir)
ds.to_netcdf(cmc_filename_out)
# -
| climatologies/make_monthly_cmc_ssts_and_monthly_climatology.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] toc=true
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"></ul></div>
# -
import requests
import json
API_ENDPOINT='https://api.wit.ai/utterances?v=20200513'
wit_access_token='<KEY>'
def createUtterances():
#ut=['Tell me about the sulfuric acid','I want to know the characteristics of the sulfuric acid','Tell me about the properties of the sulfuric acid','Tell me what you know about the sulfuric acid','Precautions with the sulfuric acid','Chemical reactions with sulfuric acid','chemical compound sulfuric acid','Warnings of the sulfuric acid','Properties of the sulfuric acid','Advance information about the sulfuric acid', 'Basic information about the sulfuric acid']
ut=['Is the benzene toxic and irritant?','Is the hexane a fammable liquid?']
chemical_compound=['benzene','hexane']
for i in range(len(ut)):
uterance=ut[i]
chemical=chemical_compound[i]
starting=uterance.find(chemical)
ending=starting+len(chemical)
body=uterance
body=body.replace(chemical,"")
print(uterance,starting,ending, body)
'''
d1={'text':'Tell me the benzene'}
d2={'intent':'info_storage_compatibility'}
d3={'entities':
[
{
'entity':'chemical_substance',
'start':12,
'end':19,
'body':'benzene',
'entities':[]
}
]
}
d4={'traits':[]}
#dat={**d1,**d2,**d3,**d4}
'''
'''
dat=[{
'text':str(ut[i]),
'intent':'ghs_classification',
'entities':
[
{
'entity':'chemical_substance:chemical_substance',
'start':int(len(ut[i])-13),
'end':int(len(ut[i])),
'body':str(ut[i][-13:]),
'entities':[]
}
],
'traits':[]
}]
'''
dat=[{
'text':str(ut[i]),
'intent':'get_ghs_classification',
'entities':
[
{
'entity':'chemical_substance:chemical_substance',
'start':starting,
'end':ending,
'body':chemical,
'entities':[]
}
],
'traits':[]
}]
headers = {'authorization': 'Bearer ' + wit_access_token,'Content-Type': 'application/json'}
resp=requests.post(API_ENDPOINT,headers=headers,json=dat)
data=json.loads(resp.content)
print(data)
return data
if __name__ == '__main__':
#intent=input()
textt=createUtterances()
if textt is None:
print("\n Result: Intent Saved on Wit.ai Success (200){}")
# +
import re
uterance='Can I mix water and sulfuric acid'
uterance2='Can I mix water and sulfuric acid'
chem_data=["water","sulfuric acid"]
for data in chem_data:
start=uterance.find(data,0)
final=start+len(data)
print("beginning:",start)
print("final:",final)
if data in uterance2:
uterance2=uterance2.replace(data,'')
body=uterance2
print('body:',body)
# -
| Training/utterance-post.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.2 64-bit (''new'': conda)'
# name: python3
# ---
# # D3net
# > Densely connected multidilated convolutional networks for dense prediction tasks
# - toc: true
# - badges: true
# - comments: true
# - sticky_rank: 8
# - author: Bowen
# - categories: [pytorch, basics]
#
# +
import mmcv
import matplotlib.pyplot as plt
from fastcore.basics import *
from fastai.vision.all import *
from fastai.torch_basics import *
import warnings
warnings.filterwarnings("ignore")
import kornia
from kornia.constants import Resample
from kornia.color import *
from kornia import augmentation as K
# import kornia.augmentation as F
import kornia.augmentation.random_generator as rg
from torchvision.transforms import functional as tvF
from torchvision.transforms import transforms
from torchvision.transforms import PILToTensor
from functools import partial
from timm.models.layers import trunc_normal_, DropPath
from timm.data import IMAGENET_DEFAULT_MEAN, IMAGENET_DEFAULT_STD
from timm.models.vision_transformer import _cfg
from einops import rearrange
from timm.models.registry import register_model
set_seed(105)
# -
# code borrowed from https://github.com/tky823/DNN-based_source_separation/blob/main/src/models/d2net.py
# +
EPS = 1e-12
def choose_layer_norm(name, num_features, n_dims=2, eps=EPS, **kwargs):
if name in ['BN', 'batch', 'batch_norm']:
if n_dims == 1:
layer_norm = nn.BatchNorm1d(num_features, eps=eps)
elif n_dims == 2:
layer_norm = nn.BatchNorm2d(num_features, eps=eps)
else:
raise NotImplementedError("n_dims is expected 1 or 2, but give {}.".format(n_dims))
else:
raise NotImplementedError("Not support {} layer normalization.".format(name))
return layer_norm
def choose_nonlinear(name, **kwargs):
if name == 'relu':
nonlinear = nn.ReLU()
else:
raise NotImplementedError("Invalid nonlinear function is specified. Choose 'relu' instead of {}.".format(name))
return nonlinear
# -
from torch.nn.modules.utils import _pair
_pair(1)
class ConvBlock2d(nn.Module):
def __init__(self, in_channels, out_channels, kernel_size, stride=1, dilation=1, norm=True, nonlinear='relu', eps=EPS):
super().__init__()
assert stride == 1, "`stride` is expected 1"
self.kernel_size = _pair(kernel_size)
self.dilation = _pair(dilation)
self.norm = norm
self.nonlinear = nonlinear
if self.norm:
if type(self.norm) is bool:
name = 'BN'
else:
name = self.norm
self.norm2d = choose_layer_norm(name, in_channels, n_dims=2, eps=eps)
if self.nonlinear is not None:
self.nonlinear2d = choose_nonlinear(self.nonlinear)
self.conv2d = nn.Conv2d(in_channels, out_channels, kernel_size=kernel_size, stride=stride, dilation=dilation)
def forward(self, input):
"""
Args:
input (batch_size, in_channels, H, W)
Returns:
output (batch_size, out_channels, H, W)
"""
Kh, Kw = self.kernel_size
Dh, Dw = self.dilation
padding_height = (Kh - 1) * Dh
padding_width = (Kw - 1) * Dw
padding_up = padding_height // 2
padding_bottom = padding_height - padding_up
padding_left = padding_width // 2
padding_right = padding_width - padding_left
x = input
if self.norm:
x = self.norm2d(x)
if self.nonlinear:
x = self.nonlinear2d(x)
x = F.pad(x, (padding_left, padding_right, padding_up, padding_bottom))
output = self.conv2d(x)
return output
temp = ConvBlock2d(3,128,3,1,1,norm=False,nonlinear=None)
temp(torch.randn(1,3,32,32)).shape
# +
EPS = 1e-12
class D2BlockFixedDilation(nn.Module):
def __init__(self, in_channels, growth_rate, kernel_size, dilation=1, norm=True, nonlinear='relu', depth=None, eps=EPS):
"""
Args:
in_channels <int>: # of input channels
growth_rate <int> or <list<int>>: # of output channels
kernel_size <int> or <tuple<int>>: Kernel size
dilation <int>: Dilation od dilated convolution.
norm <bool> or <list<bool>>: Applies batch normalization.
nonlinear <str> or <list<str>>: Applies nonlinear function.
depth <int>: If `growth_rate` is given by list, len(growth_rate) must be equal to `depth`.
"""
super().__init__()
if type(growth_rate) is int:
assert depth is not None, "Specify `depth`"
growth_rate = [growth_rate] * depth
elif type(growth_rate) is list:
if depth is not None:
assert depth == len(growth_rate), "`depth` is different from `len(growth_rate)`"
depth = len(growth_rate)
else:
raise ValueError("Not support growth_rate={}".format(growth_rate))
if not type(dilation) is int:
raise ValueError("Not support dilated={}".format(dilated))
if type(norm) is bool:
assert depth is not None, "Specify `depth`"
norm = [norm] * depth
elif type(norm) is list:
if depth is not None:
assert depth == len(norm), "`depth` is different from `len(norm)`"
depth = len(norm)
else:
raise ValueError("Not support norm={}".format(norm))
if type(nonlinear) is bool or type(nonlinear) is str:
assert depth is not None, "Specify `depth`"
nonlinear = [nonlinear] * depth
elif type(nonlinear) is list:
if depth is not None:
assert depth == len(nonlinear), "`depth` is different from `len(nonlinear)`"
depth = len(nonlinear)
else:
raise ValueError("Not support nonlinear={}".format(nonlinear))
self.growth_rate = growth_rate
self.depth = depth
net = []
_in_channels = in_channels - sum(growth_rate)
for idx in range(depth):
if idx == 0:
_in_channels = in_channels
else:
_in_channels = growth_rate[idx - 1]
_out_channels = sum(growth_rate[idx:])
conv_block = ConvBlock2d(_in_channels, _out_channels, kernel_size=kernel_size, stride=1, dilation=dilation, norm=norm[idx], nonlinear=nonlinear[idx], eps=eps)
net.append(conv_block)
self.net = nn.Sequential(*net)
def forward(self, input):
"""
Args:
input: (batch_size, in_channels, H, W)
Returns:
output: (batch_size, out_channels, H, W), where out_channels = growth_rate[-1].
"""
growth_rate, depth = self.growth_rate, self.depth
x_residual = 0
for idx in range(depth):
if idx == 0:
x = input
else:
_in_channels = growth_rate[idx - 1]
sections = [_in_channels, sum(growth_rate[idx:])]
x, x_residual = torch.split(x_residual, sections, dim=1)
x = self.net[idx](x)
x_residual = x_residual + x
output = x_residual
return output
class D2Block(nn.Module):
def __init__(self, in_channels, growth_rate, kernel_size, dilated=True, norm=True, nonlinear='relu', depth=None, eps=EPS):
"""
Args:
in_channels <int>: # of input channels
growth_rate <int> or <list<int>>: # of output channels
kernel_size <int> or <tuple<int>>: Kernel size
dilated <bool> or <list<bool>>: Applies dilated convolution.
norm <bool> or <list<bool>>: Applies batch normalization.
nonlinear <str> or <list<str>>: Applies nonlinear function.
depth <int>: If `growth_rate` is given by list, len(growth_rate) must be equal to `depth`.
"""
super().__init__()
if type(growth_rate) is int:
assert depth is not None, "Specify `depth`"
growth_rate = [growth_rate] * depth
elif type(growth_rate) is list:
if depth is not None:
assert depth == len(growth_rate), "`depth` is different from `len(growth_rate)`"
depth = len(growth_rate)
else:
raise ValueError("Not support growth_rate={}".format(growth_rate))
if type(dilated) is bool:
assert depth is not None, "Specify `depth`"
dilated = [dilated] * depth
elif type(dilated) is list:
if depth is not None:
assert depth == len(dilated), "`depth` is different from `len(dilated)`"
depth = len(dilated)
else:
raise ValueError("Not support dilated={}".format(dilated))
if type(norm) is bool:
assert depth is not None, "Specify `depth`"
norm = [norm] * depth
elif type(norm) is list:
if depth is not None:
assert depth == len(norm), "`depth` is different from `len(norm)`"
depth = len(norm)
else:
raise ValueError("Not support norm={}".format(norm))
if type(nonlinear) is bool or type(nonlinear) is str:
assert depth is not None, "Specify `depth`"
nonlinear = [nonlinear] * depth
elif type(nonlinear) is list:
if depth is not None:
assert depth == len(nonlinear), "`depth` is different from `len(nonlinear)`"
depth = len(nonlinear)
else:
raise ValueError("Not support nonlinear={}".format(nonlinear))
self.growth_rate = growth_rate
self.depth = depth
net = []
_in_channels = in_channels - sum(growth_rate)
for idx in range(depth):
if idx == 0:
_in_channels = in_channels
else:
_in_channels = growth_rate[idx - 1]
_out_channels = sum(growth_rate[idx:])
if dilated[idx]:
dilation = 2**idx
else:
dilation = 1
conv_block = ConvBlock2d(_in_channels, _out_channels, kernel_size=kernel_size, stride=1, dilation=dilation, norm=norm[idx], nonlinear=nonlinear[idx], eps=eps)
net.append(conv_block)
self.net = nn.Sequential(*net)
def forward(self, input):
"""
Args:
input: (batch_size, in_channels, H, W)
Returns:
output: (batch_size, out_channels, H, W), where out_channels = growth_rate[-1].
"""
growth_rate, depth = self.growth_rate, self.depth
for idx in range(depth):
if idx == 0:
x = input
x_residual = 0
else:
_in_channels = growth_rate[idx - 1]
sections = [_in_channels, sum(growth_rate[idx:])]
x, x_residual = torch.split(x_residual, sections, dim=1)
x = self.net[idx](x)
x_residual = x_residual + x
output = x_residual
return output
def _test_d2block():
batch_size = 4
n_bins, n_frames = 64, 64
in_channels = 3
kernel_size = (3, 3)
depth = 4
input = torch.randn(batch_size, in_channels, n_bins, n_frames)
print("-"*10, "D2 Block when `growth_rate` is given as int and `dilated` is given as bool.", "-"*10)
growth_rate = 2
dilated = True
model = D2Block(in_channels, growth_rate, kernel_size=kernel_size, dilated=dilated, depth=depth)
print("-"*10, "D2 Block", "-"*10)
print(model)
output = model(input)
print(input.size(), output.size())
print()
# print("-"*10, "D2 Block when `growth_rate` is given as list and `dilated` is given as bool.", "-"*10)
# growth_rate = [3, 4, 5, 6] # depth = 4
# dilated = False
# model = D2Block(in_channels, growth_rate, kernel_size=kernel_size, dilated=dilated)
# print(model)
# output = model(input)
# print(input.size(), output.size())
# print()
# print("-"*10, "D2 Block when `growth_rate` is given as list and `dilated` is given as list.", "-"*10)
# growth_rate = [3, 4, 5, 6] # depth = 4
# dilated = [True, False, False, True] # depth = 4
# model = D2Block(in_channels, growth_rate, kernel_size=kernel_size, dilated=dilated)
# print(model)
# output = model(input)
# print(input.size(), output.size())
# -
print("="*10, "D2 Block", "="*10)
_test_d2block()
| _notebooks/2021-12-08-d3net.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # The Magic of SHD
# > A simple yet fast and powerful forecasting algorithm
#
# - toc: true
# - badges: true
# - comments: true
# - categories: [pandas, numpy, data-cleaning]
# - hide: false
# ### What's SHD?
#
# SHD stands for (S)ingle Exponential Smoothing, (H)olt's, (D)amped forecasting algorithm. It's not often that you can describe the entire algorithm in one single sentence but I just did that. And this simple algorithm often outperforms some of the most complex forecasting algorithms including DNNs and FB Prophet on univariate low frequency time series. I have used it on many projects successfully with great results. I am sharing it because the great [Spyros Makridakis](https://www.insead.edu/faculty-research/faculty/spyros-makridakis) reminded on twitter that SHD was found superior in all M (M5 would be an exception) competitions.
# >twitter: https://twitter.com/spyrosmakrid/status/1368972398498824193?s=20
# Not many know about this gem so I thought I would share my code. It's a reminder that you don't always need complex algorithms to create forecast predictions. Use what's simple and parsimonious.
#
# **How does it work?**
#
# Just take arithmatic mean of forecast from SES, Holt's and Damped
# 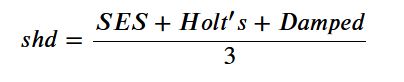
# **How does it stack against other algorithms?**
#
# [Read it yourself](https://flora.insead.edu/fichiersti_wp/inseadwp1999/99-70.pdf). It worked as good and even better than most other algorithms in the M3 competition. It works particularly well with low frequency time series (Yearly, monthly). It works well because we are ensembling three different algorithms. It's been shown that forecast combinations often outperform single best models.
#
# I will demonstrate it using an example below. This is the same dataset I used in my two previous [blogs](https://pawarbi.github.io/blog/forecasting/r/python/rpy2/altair/fbprophet/ensemble_forecast/uncertainty/simulation/2020/04/21/timeseries-part2.html).
#
#
# +
import pandas as pd
import numpy as np
import statsmodels.api as sm
from statsmodels.tsa.api import ExponentialSmoothing, SimpleExpSmoothing, Holt
import statsmodels
from statsmodels.tsa.statespace.exponential_smoothing import ExponentialSmoothing
import scipy
from scipy.stats import boxcox
from scipy.special import inv_boxcox
from statsmodels.tools.eval_measures import rmse
# -
print('Pandas:', pd.__version__)
print('Statsmodels:', sm.__version__)
print('Scipy:', scipy.__version__)
print('Numpy:', np.__version__)
# #### SHD
# +
def combshd(train,horizon,seasonality, init):
# Author: <NAME>
# Date: 8/30/2020
# version: 1.1
'''
params
----------
:train numpy array or Pandas series with univariate data
:horizon forecast horizon (int)
:seasonality For monthly 12, yearly 1, quarerly 4 (int)
:init initialization ('heuristic','concentrated')
output
------------
numpy array if length equal to specified horizon
'''
train_x,lam = boxcox (train)
ses=(sm.tsa.statespace.ExponentialSmoothing(train_x,
trend=True,
seasonal=None,
initialization_method= init,
damped_trend=False).fit())
fc1 = inv_boxcox(ses.forecast(horizon),lam)
holt=(sm.tsa.statespace.ExponentialSmoothing(train_x,
trend=True,
seasonal=seasonality,
initialization_method= init,
damped_trend=False).fit())
fc2 = inv_boxcox(holt.forecast(horizon),lam)
damp=(sm.tsa.statespace.ExponentialSmoothing(train_x,
trend=True,
seasonal=seasonality,
initialization_method= init,
damped_trend=True).fit())
fc3 = inv_boxcox(damp.forecast(horizon),lam)
fc = (fc1+fc2+fc3)/3
return fc
# +
data = pd.read_csv("https://raw.githubusercontent.com/pawarbi/datasets/master/timeseries/ts_frenchretail.csv")
data['Date']= pd.to_datetime(data['Date'])
data.set_index('Date', inplace= True)
train = data.iloc[:-4]
test = data.iloc[-4:]
data.head()
# -
data.plot();
# +
print("Train legth:", len(train), "\nTest legth:",len(test))
assert len(data)==len(train) + len(test)
# -
# #### Create forecast
shd_pred = combshd(train = train['Sales'].values,horizon=len(test),seasonality = 4, init = 'heuristic')
rmse(test['Sales'].values,shd_pred ).round(0)
# RMSE using SHD is 63734. For comparison, [FB Prophet](https://pawarbi.github.io/blog/forecasting/r/python/rpy2/altair/fbprophet/ensemble_forecast/uncertainty/simulation/2020/04/21/timeseries-part2.html#Facebook-Prophet) gave ~66,000 and [SARIMA](https://pawarbi.github.io/blog/forecasting/r/python/rpy2/altair/fbprophet/ensemble_forecast/uncertainty/simulation/2020/04/21/timeseries-part2.html#SARIMA2---(Using-Logged-value)) was ~82,000. I was able to further improve this by ensembling many algorithms but still it's impressive that with just few lines of code you can create a sophisticated algorithm !
#
# A reminder, **always start with baseline simple algorithms**. In practice, [fast and frugal](https://arxiv.org/abs/2102.13209) wins the long race.
| _notebooks/2021-03-08-shd-forecasting-ses-exponential-damped.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="https://raw.githubusercontent.com/Qiskit/qiskit-tutorials/master/images/qiskit-heading.png" alt="Note: In order for images to show up in this jupyter notebook you need to select File => Trusted Notebook" width="500 px" align="left">
# ## _*Two-qubit Quantum Random Access Coding (QRAC)*_
#
# The latest version of this notebook is available on https://github.com/QISKit/qiskit-tutorial.
#
# ***
# ### Contributors
# <NAME>, <NAME>
#
# ### Qiskit Package Versions
import qiskit
qiskit.__qiskit_version__
# ## Introduction
#
# As seen [here](single-qubit_quantum_random_access_coding.ipynb), thanks to the superposition of quantum states, by QRAC a single qubit can be used to encode up to 3 bits $x_1x_2x_3$ of information so that any one bit $x_i$ can be recovered with probability at least $0.78$. What happens if we have more qubits? As more qubits are available, we can go beyond a simple combination of QRAC of individual qubit by leveraging the entanglement. With two qubits, we can clearly use $(3,1)$-QRAC on each qubit and obtain a $(6,2)$-QRAC. Can we encode more bits? The answer is yes. Notice that two classical bits can be used to encode only up to 3 bits of information in the random access coding.
#
# We will show a $(7,2)$-QRAC following Example 4 in the paper [here](http://iopscience.iop.org/article/10.1088/1367-2630/8/8/129/meta). The success probability is $0.54$, which is slightly better than random guessing. The $(7,2)$-QRAC in the example uses [*quantum mixed states*](https://en.wikipedia.org/wiki/Quantum_state#Mixed_states) to mix two $(3,1)$-QRACs, for encoding the first six bits $x_1x_2x_3x_4x_5x_6$, with the encoding of the seventh bit $x_7$. Namely, the mixed state contains, with probability $\alpha \equiv \frac{6}{7+\sqrt{3}}$, two $(3,1)$-QRACs encoding the first six bits, and the encoding of $x_7$ that entangles the first qubit and the second qubit (if $x_7 = 0$, the two qubits become $|00\rangle + |11\rangle$, or $|01\rangle+|10\rangle$ otherwise) with probability $1 - \alpha$.
#
# The *quantum mixed states* used in the $(7,2)$-QRAC can be created by the sender using a series of unitary operations on three qubits by using the first qubit to control the mixing probability. The sender then sends the last two qubits (the second and third ones) that are used by the receiver to decode his choice of bit.
#
# The procedure is as follows. First, the sender prepares three qubits $|000\rangle$ and transforms them to obtain
#
# \begin{eqnarray}
# |000\rangle &\xrightarrow{U(1.187, 0, 0)}& \sqrt{\alpha} |000\rangle + \sqrt{1 - \alpha} |100\rangle,
# \end{eqnarray}
#
# where $U(1.187,0,0)$ is the single-qubit rotation gate applied to the first qubit. Notice that $\cos(1.187/2) \approx \sqrt\alpha$.
#
# Next, the sender uses her first qubit to control the value of the second and third qubits. When, the first qubit is $0$, the sender encodes the $x_1x_2x_3$ into the second qubit, and the $x_4x_5x_6$ into the third qubit. This means, she performs the following transformation (note that $V$ is conditioned on $x_1x_2x_3x_4x_5x_6$),
#
# \begin{eqnarray}
# |000\rangle &\xrightarrow{V}& |0\rangle |\phi\left(x_1x_2x_3\right)\rangle |\phi\left(x_4x_5x_6\right)\rangle,
# \end{eqnarray}
#
# where $|\phi\left(xyz\right)\rangle$ is the $(3,1)$-QRAC of $xyz$. Otherwise, when the second qubit is $1$, the sender encodes the $x_7$ by using the Bell states. Namely, by entangling the second and the third qubit so that when $x_7 = 0$ the second and third qubits are the same, and when $x_7 = 1$ they are different, as follows.
#
# \begin{eqnarray}
# |100\rangle & \xrightarrow{\mbox{if}~x_7 = 0} & |1\rangle\left( \frac{1}{\sqrt{2}}|00\rangle + \frac{1}{\sqrt{2}} |11\rangle\right) \equiv |1\rangle|\xi(0)\rangle \\
# |100\rangle & \xrightarrow{\mbox{if}~x_7 = 1} & |1\rangle\left(\frac{1}{\sqrt{2}}|01\rangle + \frac{1}{\sqrt{2}} |10\rangle\right) \equiv |1\rangle|\xi(1)\rangle
# \end{eqnarray}
#
# The second and third qubits are then sent to the receiver, who performs measurement to obtain his desired bit. Notice that because the receiver does not have the first qubit, the state of his two qubits is the mixed state as below
#
# $$
# \rho\left(x_1x_2\ldots x_7\right) = \alpha |\phi\left(x_1x_2x_3\right)\rangle\langle\phi\left(x_1x_2x_3\right)| \otimes |\phi\left(x_4x_5x_6\right)\rangle\langle\phi\left(x_4x_5x_6\right)| + (1-\alpha) |\xi\left(x_7\right)\rangle\langle\xi\left(x_7\right)|
# $$
#
# Now, the receiver can recover any one bit of $x_1x_2\ldots x_7$ as follows. If he wants to obtain any one bit of $x_1x_2x_3$, he measures his first qubit using the measurements of $(3,1)$-QRAC. Similarly, if he wants to obtain any one bit of $x_4x_5x_6$, he measures his second qubit using the measurements of $(3,1)$-QRAC. Finally, if he wants to obtain the seventh bit $x_7$, he measures both qubits and concludes that $x_7 = 0$ if the outcome of the measurement is the same (they are both $0$ or $1$), or $x_7 = 1$ otherwise.
#
# Now, we show how to perform the experiment on the above $(7,2)$-QRAC using the IBM Q Experience. We first prepare the environment.
# +
# useful math functions
from math import pi, cos, acos, sqrt
# importing QISKit
from qiskit import Aer, IBMQ
from qiskit import QuantumCircuit, ClassicalRegister, QuantumRegister, execute
from qiskit.tools.visualization import plot_histogram
from qiskit.tools.monitor import job_monitor
from qiskit.providers.ibmq import least_busy
# -
IBMQ.load_account()
# We prepare the controlled-Hadamard and controlled-u3 gates that are required in the encoding as below.
# +
def ch(qProg, a, b):
""" Controlled-Hadamard gate """
qProg.h(b)
qProg.sdg(b)
qProg.cx(a, b)
qProg.h(b)
qProg.t(b)
qProg.cx(a, b)
qProg.t(b)
qProg.h(b)
qProg.s(b)
qProg.x(b)
qProg.s(a)
return qProg
def cu3(qProg, theta, phi, lambd, c, t):
""" Controlled-u3 gate """
qProg.u1((lambd-phi)/2, t)
qProg.cx(c, t)
qProg.u3(-theta/2, 0, -(phi+lambd)/2, t)
qProg.cx(c, t)
qProg.u3(theta/2, phi, 0, t)
return qProg
# -
# ## Encoding 7 bits into 2 qubits with $(7,2)$-QRAC
#
# The sender prepares the state to be sent by working on 3 qubits and uses the first one to control the mixture. She needs controlled unitary gates of $(3,1)$-QRAC encoding for her first 6 bits. When the first qubit is zero, she encodes $x_1x_2x_3$ into the second qubit, and $x_4x_5x_6$ into the third qubit. This can be realized with NOT on the first qubit followed by controlled $(3,1)$-QRAC gates with the first qubit as control and the second and third qubits as targets.
#
# To encode $x_7$, when $x_7 = 0$ the sender applies a controlled Hadamard on the second qubit (with the first qubit as control), and apply Toffoli gate on the third qubit using the first and second qubits as controls. When $x_7 = 1$, the sender flips the second and third qubits before applying the same operations as when $x_7 = 0$.
#
# The decoding operations are the same as those of $(3,1)$-QRAC, and for encoding $x_7$, the receiver just measures the value of the second and third qubit on the computational basis.
#
# Below is the quantum circuits for encoding `"0101010"` and decoding any one bit with $(7,2)$-QRAC.
# +
#CHANGE THIS 7BIT 0-1 STRING TO PERFORM EXPERIMENT ON ENCODING 0000000, ..., 1111111
x1234567 = "0101010"
if len(x1234567) != 7 or not("1" in x1234567 or "0" in x1234567):
raise Exception("x1234567 is a 7-bit 0-1 pattern. Please set it to the correct pattern")
#compute the value of rotation angle theta of (3,1)-QRAC
theta = acos(sqrt(0.5 + sqrt(3.0)/6.0))
#to record the u3 parameters for encoding 000, 010, 100, 110, 001, 011, 101, 111
rotationParams = {"000":(2*theta, pi/4, -pi/4), "010":(2*theta, 3*pi/4, -3*pi/4),
"100":(pi-2*theta, pi/4, -pi/4), "110":(pi-2*theta, 3*pi/4, -3*pi/4),
"001":(2*theta, -pi/4, pi/4), "011":(2*theta, -3*pi/4, 3*pi/4),
"101":(pi-2*theta, -pi/4, pi/4), "111":(pi-2*theta, -3*pi/4, 3*pi/4)}
# Creating registers
# qubits for encoding 7 bits of information with qr[0] kept by the sender
qr = QuantumRegister(3)
# bits for recording the measurement of the qubits qr[1] and qr[2]
cr = ClassicalRegister(2)
encodingName = "Encode"+x1234567
encodingCircuit = QuantumCircuit(qr, cr, name=encodingName)
#Prepare superposition of mixing QRACs of x1...x6 and x7
encodingCircuit.u3(1.187, 0, 0, qr[0])
#Encoding the seventh bit
seventhBit = x1234567[6]
if seventhBit == "1": #copy qr[0] into qr[1] and qr[2]
encodingCircuit.cx(qr[0], qr[1])
encodingCircuit.cx(qr[0], qr[2])
#perform controlled-Hadamard qr[0], qr[1], and toffoli qr[0], qr[1] , qr[2]
encodingCircuit = ch(encodingCircuit, qr[0], qr[1])
encodingCircuit.ccx(qr[0], qr[1], qr[2])
#End of encoding the seventh bit
#encode x1...x6 with two (3,1)-QRACS. To do that, we must flip q[0] so that the controlled encoding is executed
encodingCircuit.x(qr[0])
#Encoding the first 3 bits 000, ..., 111 into the second qubit, i.e., (3,1)-QRAC on the second qubit
firstThreeBits = x1234567[0:3]
#encodingCircuit.cu3(*rotationParams[firstThreeBits], qr[0], qr[1])
encodingCircuit = cu3(encodingCircuit, *rotationParams[firstThreeBits], qr[0], qr[1])
#Encoding the second 3 bits 000, ..., 111 into the third qubit, i.e., (3,1)-QRAC on the third qubit
secondThreeBits = x1234567[3:6]
#encodingCircuit.cu3(*rotationParams[secondTreeBits], qr[0], qr[2])
encodingCircuit = cu3(encodingCircuit, *rotationParams[secondThreeBits], qr[0], qr[2])
#end of encoding
encodingCircuit.barrier()
# dictionary for decoding circuits
decodingCircuits = {}
# Quantum circuits for decoding the 1st to 6th bits
for i, pos in enumerate(["First", "Second", "Third", "Fourth", "Fifth", "Sixth"]):
circuitName = "Decode"+pos
decodingCircuits[circuitName] = QuantumCircuit(qr, cr, name=circuitName)
if i < 3: #measure 1st, 2nd, 3rd bit
if pos == "Second": #if pos == "First" we can directly measure
decodingCircuits[circuitName].h(qr[1])
elif pos == "Third":
decodingCircuits[circuitName].u3(pi/2, -pi/2, pi/2, qr[1])
decodingCircuits[circuitName].measure(qr[1], cr[1])
else: #measure 4th, 5th, 6th bit
if pos == "Fifth": #if pos == "Fourth" we can directly measure
decodingCircuits[circuitName].h(qr[2])
elif pos == "Sixth":
decodingCircuits[circuitName].u3(pi/2, -pi/2, pi/2, qr[2])
decodingCircuits[circuitName].measure(qr[2], cr[1])
#Quantum circuits for decoding the 7th bit
decodingCircuits["DecodeSeventh"] = QuantumCircuit(qr, cr, name="DecodeSeventh")
decodingCircuits["DecodeSeventh"].measure(qr[1], cr[0])
decodingCircuits["DecodeSeventh"].measure(qr[2], cr[1])
#combine encoding and decoding of (7,2)-QRACs to get a list of complete circuits
circuitNames = []
circuits = []
k1 = encodingName
for k2 in decodingCircuits.keys():
circuitNames.append(k1+k2)
circuits.append(encodingCircuit+decodingCircuits[k2])
print("List of circuit names:", circuitNames) #list of circuit names
# for circuit in circuits: #list qasms codes
# print(circuit.qasm())
# -
# Below are plots of the experimental results of extracting the first to sixth bit, that results in observing the-$i$th bit with probability at least $0.54$
# +
# Use the qasm simulator
backend = Aer.get_backend("qasm_simulator")
# Use the IBM Quantum Experience
#backend = least_busy(IBMQ.backends(simulator=False))
shots = 1000
job = execute(circuits, backend=backend, shots=shots)
#job_monitor(job)
# -
results = job.result()
for k in ["DecodeFirst", "DecodeSecond", "DecodeThird", "DecodeFourth", "DecodeFifth", "DecodeSixth"]:
print("Experimental Result of ", encodingName+k)
plot_histogram(results.get_counts(circuits[circuitNames.index(encodingName+k)]))
# The seventh bit is obtained by looking at the content of classical registers. If they are the same, i.e., both are `1` or `0`, then we conclude that it is `0`, or otherwise `1`. For the encoding of `0101010`, the seventh bit is `0`, so the total probability of observing `00` and `11` must exceed that of observing `01` and `10`.
print("Experimental result of ", encodingName+"DecodeSeventh")
plot_histogram(results.get_counts(circuits[circuitNames.index(encodingName+"DecodeSeventh")]))
# We can experiment with other seven-bit encodings by changing the value `x1234567` in the code above, and see that in any case we can decode the correct bit with probability strictly better than random guessing.
# ## About Quantum Random Access Coding
# The success probability $(7,2)$-QRAC shown above is not high due to the use of mixed states. There exists a better code using pure states as shown [here](https://arxiv.org/abs/1607.02667), where the success probability becomes $0.68$. [The paper](https://arxiv.org/abs/1607.02667) also shows the construction of $(n,2)$-QRACs with pure states for $n=7,\ldots,12$. It is still open if there is a coding scheme with only pure states for $n=13, 14, 15$. Notice that $(n, m)$-QRACs exist for $ n \le 2^{2m}-1$ that can be realized with mixed states as shown [here](http://link.springer.com/chapter/10.1007/978-3-540-73420-8_12).
| terra/qis_adv/two-qubit_state_quantum_random_access_coding.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import tensorflow as tf
mnist = tf.keras.datasets.fashion_mnist
(training_images, training_labels), (test_images, test_labels) = mnist.load_data()
training_images=training_images.reshape(60000, 28, 28, 1)
training_images=training_images / 255.0
test_images = test_images.reshape(10000, 28, 28, 1)
test_images=test_images/255.0
model = tf.keras.models.Sequential([
tf.keras.layers.Conv2D(64, (3,3), activation='relu', input_shape=(28, 28, 1)),
tf.keras.layers.MaxPooling2D(2, 2),
tf.keras.layers.Conv2D(64, (3,3), activation='relu'),
tf.keras.layers.MaxPooling2D(2,2),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(128, activation='relu'),
tf.keras.layers.Dense(10, activation='softmax')
])
model.compile(optimizer='adam', loss='sparse_categorical_crossentropy', metrics=['accuracy'])
model.summary()
model.fit(training_images, training_labels, epochs=5)
test_loss = model.evaluate(test_images, test_labels)
| jupyter/Basic_example/MNIST/MNIST_example_CNN&POOLING(not include validation).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: ML
# language: python
# name: ml
# ---
import unicodedata
# # Normalization
#
# Unicode normalization is used to *normalize* different but similiar characters. For example the following unicode characters (and character combinations) are equivalent:
#
# **Canonical Equivalence**
#
# | | | Equivalence Reason |
# | --- | --- | --- |
# | Ç | C◌̧ | Combined character sequences |
# | 가 | ᄀ ᅡ | Conjoined Korean characters |
#
# **Compatibility equivalence**
#
# | | | Equivalence Reason |
# | --- | --- | --- |
# | ℌ | H | Font variant |
# | \[NBSP\] | \[SPACE\] | Both are linebreak sequences |
# | ① | 1 | Circled variant |
# | x² | x2 | Superscript |
# | xⱼ | xj | Subscript |
# | ½ | 1/2 | Fractions |
#
# We have mentioned two different types of equivalence here, canonical and compatibility equivalence.
#
# **Canonical equivalence** means both forms are fundamentally the same and when rendered are indistinguishable. For example we can take the unicode for `'Ç' \u00C7` or the unicode for `'C' \u0043` and `'̧' \u0327`, when the latter two characters are rendered together they look the same as the first character:
print("\u00C7", "\u0043"+"\u0327")
# However, if we print these characters seperately, we can see very clearly that they are not the same:
print("\u00C7", "\u0043", "\u0327")
# These are examples of canonical equivalence, but we also have compatibility equivalence.
#
# **Compatibility equivalence** refers to the formatting differences between characters, which includes (but is not limited to):
#
# * font variants
# * cursive forms
# * circled characters
# * width variation
# * size changes
# * rotation
# * superscript and subscript
# * fractions
#
# In this case we can see a difference between the rendered characters, for example between `ℌ` and `H`, or `½` and `1 ⁄ 2`.
#
# For many of these examples which are either canonical equivalents (Ç ↔ C ̧ ) or compatibility equivalents (½ → 1 ⁄ 2), if we compare if these different forms are equal, we will find that they are not:
"Ç" == "Ç"
"ℌ" == "H"
"½" == "1⁄2" # note that 1⁄2 are the characters 1 ⁄ 2 placed together (they are automatically formatted)
# So it is in these cases that we use unicode normalization to *normalize* our characters into matching pairs. As there are different forms of equivalence, there are also different forms of normalization. These are all called **N**ormal **F**orm, and there are four different methods:
#
# | Name | Abbreviation | Description | Example |
# | --- | --- | --- | --- |
# | Form D | NFD | *Canonical* decomposition | `Ç` → `C ̧` |
# | Form C | NFC | *Canoncial* decomposition followed by *canonical* composition | `Ç` → `C ̧` → `Ç` |
# | Form KD | NFKD | *Compatibility* decomposition | `ℌ ̧` → `H ̧` |
# | Form KC | NFKC | *Compatibility* decomposition followed by *canonical* composition | `ℌ ̧` → `H ̧` → `Ḩ` |
#
# Let's take a look at each of these forms in action. Our C with cedilla character Ç can be represented in two ways, as a single character called *Latin capital C with cedilla* (*\u00C7*), or as two characters called *Latin capital C* (*\u0043*) and *combining cedilla* (*\u0327*):
c_with_cedilla = "\u00C7" # Latin capital C with cedilla (single character)
c_with_cedilla
c_plus_cedilla = "\u0043\u0327" # \u0043 = Latin capital C, \u0327 = 'combining cedilla' (two characters)
c_plus_cedilla
# And we will find that these two version do not match when compared:
c_with_cedilla == c_plus_cedilla
# If we perform **NFD** on our C with cedilla character `\u00C7`, we **decompose** the character into it's smaller components, which are the *Latin capital C* character, and *combining cedilla* character `\u0043` + `\u0327`. This means that if we compare an **NFD** normalized C with cedilla character to both the C character and the cedilla character, we will return true:
unicodedata.normalize('NFD', c_with_cedilla) == c_plus_cedilla
# However, if we perform **NFC** on our C with cedilla character `\u00C7`, we **decompose** the character into the smaller components `\u0043` + `\u0327`, and then **compose** them back to `\u00C7`, and so they will not match:
unicodedata.normalize('NFC', c_with_cedilla) == c_plus_cedilla
# But if we switch the **NFC** encoding to instead be performed on our two characters `\u0043` + `\u0327`, they will first be **decomposed** (which will do nothing as they are already decomposed), then compose them into the single `\u00C7` character:
c_with_cedilla == unicodedata.normalize('NFC', c_plus_cedilla)
# The **NFK** encodings do not decompose characters into smaller components, they decompose characters into their *normal* versions. For example if we take the fancy format *ℌ* `\u210B`, we cannot decompose this into multiple characters and so *NFD* or *NFC* encoding will do nothing. However, if we apply **NFKD**, we will find that our fancy *ℌ* `\u210B` becomes a plain, boring *H* `\u0048`:
unicodedata.normalize('NFKD', 'ℌ')
# But, what if we merge our fancy ℌ `\u210B` with a combining cedilla `\u0328` character?
"\u210B\u0327"
# Applying our compatibility decomposition normalization (*NFKD*) gives us a capital H character, and a combining cedilla character as two seperate encodings:
unicodedata.normalize('NFKD', "\u210B\u0327").encode('utf-8')
# But if we apply **NFKC**, we first perform compatibility decomposition, into the two seperate characters, before merging them during *canonical* composition:
unicodedata.normalize('NFKC', "\u210B\u0327").encode('utf-8')
# Because the only difference between these two methods is a canonical composition, we see no difference between the two character sets when they are rendered:
unicodedata.normalize('NFKC', "\u210B\u0327"), unicodedata.normalize('NFKD', "\u210B\u0327"),
# Okay, that's it for unicode normalization. When it comes down to it, all you really need to remember is the table we saw above, all of this stuff is quite abstract and not the easiest thing to grasp or remember. So if it seems confusing, that's normal, you'll get by with our normal forms table.
#
# ## Further Reading
#
# [UAX #15: Unicode Normalization Forms](https://unicode.org/reports/tr15/)
#
# [UTR #15: Unicode Normalization](https://unicode.org/reports/tr15/tr15-18.html)
| course/preprocessing/04_normalization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # EDA King County Housing Data
# +
# import libraries
import random
import pandas as pd
import numpy as np
import scipy.stats as stats
# %matplotlib inline
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LinearRegression
from sklearn.metrics import mean_squared_error, r2_score
from collections import Counter
import seaborn as sns
import math
# color blind accessible colors
COLORS = ["#D81B60", "#1E88E5", "#FFC107"]
# -
# Reading Data into a DataFrame
houses = pd.read_csv('King_County_House_prices_dataset.csv')
# Displaying the data table in a more readable format
pd.set_option('float_format', '{:.2f}'.format)
pd.set_option("display.max_columns", None)
houses.head()
# Displaying the size of the datasets and observed variables
houses.shape
# Displaying the list of variables, i.e columns
houses.columns
# Displaying a summary of the dataset
houses.info()
# display which variables have missing values
houses.isnull().sum()
# replacing missing values with the mean of the respective variables
houses["waterfront"].fillna((houses["waterfront"].mean()), inplace=True)
houses["yr_renovated"].fillna((houses["yr_renovated"].mean()), inplace=True)
houses.isnull().sum()
# display basic statistical data
houses.describe()
# dropping variables that are not interesting for the regression model
houses.drop(["id", "view", "lat", "long", "sqft_lot15", "sqft_basement"], axis=1, inplace=True)
houses.head()
# +
houses['waterfront'] = np.round(houses['waterfront'])
houses['bathrooms'] = np.round(houses['bathrooms'])
houses['yr_renovated'] = np.round(houses['yr_renovated'])
houses['floors'] = np.round(houses['floors'])
houses.head(3)
# -
# display interdependencies and correlations between the variables
houses.corr()
# a better display interdependencies and correlations between the variables
fig, ax = plt.subplots(figsize=(25, 15))
sns.heatmap(houses.corr(), annot=True, ax=ax)
# +
# first plot of the dependant variable price
plt.figure(figsize=(9, 8))
#plt.plot(houses['price'], color='#E66100')
#sns.distplot(houses['price'], color='#E66100', hist_kws={'alpha': 0.4})
chart = sns.histplot(data=houses, x='price', kde=True)
chart.set(xlim = (100000,2000000))
# format the ticks to read the axes better
ticks = chart.axes.get_xticks()
xlabels = ['$' + '{:,.0f}'.format(x) for x in ticks]
chart.set_xticklabels(xlabels)
chart
# -
# display the distribution of all variables
houses.hist(bins=100, figsize=(24, 22))
sns.pairplot(houses,x_vars=['sqft_living','bathrooms', 'bedrooms', 'grade', 'sqft_living15', 'waterfront', 'yr_renovated', 'condition'],y_vars=['price']);
# dropping variables that are not interesting for the regression model
houses.drop(['waterfront', 'yr_renovated'], axis=1, inplace=True)
# # Simple Linear Regression
# import an additional library
import statsmodels.formula.api as smf
# ## Model 1
# create first regression model with price as the dependant and sqft_living as independent variable
results1 = smf.ols(formula='price ~ sqft_living', data=houses).fit()
# assigned the parameters of the model to variables
intercept1, slope1 = results1.params
intercept1, slope1
# Plotting the results of our model
fig, ax = plt.subplots(figsize=(8, 4))
x = houses.sqft_living
y = houses.price
# add data points
ax.scatter(x, y, alpha=0.5, color="#D81B60")
fig.suptitle('Relationship between price and sqft_living')
# plotting regression line
ax.plot(x, x*slope1 +intercept1, '-', color=COLORS[1], linewidth=2);
ax.set_ylabel("price");
ax.set_xlabel("sqft_living")
# summary of the models data
results1.summary()
# ## Model 2
# create another regression model with price as the dependant and sqft_lot as independent variable
results2 = smf.ols(formula='price ~ sqft_lot', data=houses).fit()
intercept2, slope2 = results2.params
print("Intercept:", intercept2, " ", "Slope:", slope2)
# Plotting the results of our model
fig, ax = plt.subplots(figsize=(8, 4))
x = houses.sqft_lot
# add data points
ax.scatter(x, y, alpha=0.5, color="#D81B60")
fig.suptitle('Relationship between price and sqft_lot')
# plotting regression line
ax.plot(x, x*slope2 +intercept2, '-', color=COLORS[2], linewidth=2);
ax.set_ylabel("price");
ax.set_xlabel("sqft_lot");
# summary of the models data
results2.summary()
# ## Model 3
# import another library
from sklearn.linear_model import LinearRegression
import plotly.express as px
import plotly.graph_objects as go
# +
#get one X variable and our target(y)
X = houses['sqft_living15'].values.reshape(-1,1)
y = houses['price'].values.reshape(-1,1)
#splitting Train and Test
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.33, random_state=0)
#Liner Regression
from sklearn.linear_model import LinearRegression
regressor = LinearRegression()
regressor.fit(X_train, y_train)
# -
#evaluate the model (intercept and slope)
print(regressor.intercept_)
print(regressor.coef_)
#predicting the test set result
y_pred = regressor.predict(X_test)
#compare actual output values with predicted values
new_df = pd.DataFrame({'Actual': y_test.flatten(), 'Predicted': y_pred.flatten()})
# visualize comparison result as a bar graph
df1 = new_df.head(20)
df1.plot(kind='bar',figsize=(16,10))
plt.grid(which='major', linestyle='-', linewidth='0.5', color='green')
plt.grid(which='minor', linestyle=':', linewidth='0.5', color='black')
plt.show()
#prediction vs test set
plt.scatter(X_test, y_test, color='blue')
plt.plot(X_test, y_pred, color='red', linewidth=2)
plt.show()
# evaluate the performance of the algorithm
from sklearn import metrics
#(MAE):mean of the absolute value of the errors
print('MAE:', metrics.mean_absolute_error(y_test, y_pred))
#(MSE) is the mean of the squared errors
print('MSE:', metrics.mean_squared_error(y_test, y_pred))
#(RMSE): square root of the mean of the squared errors
print('RMSE:', np.sqrt(metrics.mean_squared_error(y_test, y_pred)))
# # Multiple Linear Regression
# ## Model 1
# a different and simpler regression model constructor to contruct a multivariable regression model
model1 = smf.ols(formula='price ~ sqft_living + bathrooms + bedrooms + condition + grade + sqft_lot + sqft_above + sqft_living15 + yr_built + zipcode', data=houses).fit()
# display rsquared
model1.rsquared
# +
from statsmodels.tools.eval_measures import rmse
# fit your model which you have already done
# now generate predictions
ypred = model1.predict(houses)
ypred
# calc rmse
rmse = rmse(houses['price'], ypred)
rmse
# -
# ## Model 2
# +
#training the simple Linear Regression model on the training set
X_1 = houses[['sqft_living', 'bathrooms', 'bedrooms', 'condition', 'grade', 'sqft_lot', 'sqft_above', 'sqft_living15', 'yr_built', 'zipcode']]
y_1 = houses['price']
X_train_1, X_test_1, y_train_1, y_test_1 = train_test_split(X_1, y_1, test_size=0.25, random_state=42, shuffle=True)
# Merge datasets after test split for formula notation
X_train_1 = X_train_1.merge(y_train_1, left_index = True, right_index=True)
# -
# adjusted previous regression model after transforming zipcode to dummies
adj_model1 = smf.ols(formula='price ~ sqft_living + bathrooms + bedrooms + condition + grade + sqft_lot + sqft_above + sqft_living15 + yr_built + C(zipcode)', data=X_train_1).fit()
adj_model1.rsquared
# +
from statsmodels.tools.eval_measures import rmse
# fit your model which you have already done
# now generate predictions
ypred = adj_model1.predict(X_test_1)
ypred
# calc rmse
rmse = rmse(y_test_1, ypred)
rmse
# -
#compare actual output values with predicted values
new2_df = pd.DataFrame({'Actual': y_test_1, 'Predicted': ypred})
df1 = new2_df.head(10)
df1.head()
# ## Polynomial Regression
# PolynomialFeatures
from sklearn.preprocessing import PolynomialFeatures
# +
#training the simple Linear Regression model on the training set
poly = PolynomialFeatures (degree = 2)
X_poly = poly.fit_transform(X_1)
X_train,X_test,y_train,y_test = train_test_split(X_poly,y, test_size = 0.25, random_state = 42)
#X_train = X_train.merge(y_train, left_index = True, right_index=True)
# +
#adj_model2 = smf.ols(formula='price ~ sqft_living + bathrooms + bedrooms + condition + grade + sqft_lot + sqft_above + sqft_living15 + yr_built + C(zipcode)', data=X_train).fit()
# -
#fit and predict model
poly_lr = LinearRegression().fit(X_train,y_train)
y_pred = poly_lr.predict(X_test)
#checking accuracy of Polynomial Regression Model
print('Polynomial Regression Model:')
print("Train Score {:.2f}".format(poly_lr.score(X_train,y_train)))
print("Test Score {:.2f}".format(poly_lr.score(X_test, y_test)))
#evaluate the model - Coefficient and constant
print(poly_lr.intercept_)
print(poly_lr.coef_)
#compare actual output values with predicted values
df = pd.DataFrame({'Actual': y_test, 'Predicted': y_pred})
df2 = df.head(10)
df2
# evaluate the performance of the algorithm (MAE - MSE - RMSE)
from sklearn import metrics
print('MAE:', metrics.mean_absolute_error(y_test, y_pred))
print('MSE:', metrics.mean_squared_error(y_test, y_pred))
print('RMSE:', np.sqrt(metrics.mean_squared_error(y_test, y_pred)))
| kc_house_prices/Housing_Prices_Linear_Regression_Project.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
from numpy import *
from PIL import *
import pickle
from pylab import *
import os
from scipy.misc import *
from matplotlib.pyplot import *
import cv2
import imtools
imtools = reload(imtools)
def compute_feature(im):
""" Returns a feature vector for an
ocr image patch. """
# resize and remove border
norm_im = imresize(im, (30, 30))
norm_im = norm_im[3:-3, 3:-3]
m = cv2.moments(norm_im)
hu = cv2.HuMoments(m)
hu = hu
return hu.flatten()
def load_ocr_data(path):
""" Return labels and ocr features for all images in path. """
# create list of all files ending in .jpg
imlist = [os.path.join(path, f) for f in os.listdir(path) if f.endswith('.jpg')]
labels = [int(imfile.split('/')[-1][0]) for imfile in imlist]
features = []
for imname in imlist:
im = array(Image.open(imname).convert('L'))
features.append(compute_feature(im))
return array(features), labels
from svmutil import *
# +
features, labels = load_ocr_data('sudoku_images/ocr_data/training/')
test_features, test_labels = load_ocr_data('sudoku_images/ocr_data/testing/')
# freatures = array([f/linalg.norm(f) for f in features.T if linalg.norm(f)>0]).T
features = map(list, features)
test_features = map(list, test_features)
# -
prob = svm_problem(labels, features)
param = svm_parameter('-t 0')
m = svm_train(prob, param)
res = svm_predict(labels, features, m)
res = svm_predict(test_labels, test_features, m)
# +
# Not very good. Any way to improve?
| Chapter-10/CV Book Chapter 10 Exercise 6.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="W5sq-RKaxriS"
import json
import tensorflow as tf
import tensorflow.keras as keras
import json
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
# + id="_rmpsz8o6FKv"
file ="data.json"
def load_data(filename=file):
with open(filename, "r") as f:
data = json.load(f)
inputs = np.array(data["mffc"])
targets = np.array(data["labels"])
return inputs , targets
def prepared_dataset(test_size, validation_size):
X,y = load_data()
X_train , X_test , y_train , y_test = train_test_split(X,y , test_size=test_size)
#Here we create the validation
X_train , X_validation , y_train , y_validation = train_test_split(X_train ,y_train , test_size=validation_size)
X_train = X_train[..., np.newaxis]
X_validation = X_validation[..., np.newaxis]
X_test = X_test[..., np.newaxis]
return X_train , X_validation ,X_test , y_train ,y_validation , y_test
def build_model(input_shape):
model = keras.Sequential()
model.add(keras.layers.Conv2D(32, (3, 3), activation='relu', input_shape=input_shape))
model.add(keras.layers.MaxPooling2D((3, 3), strides=(2, 2), padding='same'))
model.add(keras.layers.BatchNormalization())
model.add(keras.layers.Conv2D(32, (2, 2), activation='relu'))
model.add(keras.layers.MaxPooling2D((2, 2), strides=(2, 2), padding='same'))
model.add(keras.layers.BatchNormalization())
model.add(keras.layers.Flatten())
model.add(keras.layers.Dense(64, activation='relu'))
model.add(keras.layers.Dropout(0.3))
model.add(keras.layers.Dense(1, activation='sigmoid'))
return model
def predict(model, X, y):
X = X[np.newaxis, ...] # array shape (1, 130, 13, 1)
# perform prediction
prediction = model.predict(X)
# get index with max value
predicted_index = np.argmax(prediction, axis=1)
print("Target: {}, Predicted label: {}".format(y, predicted_index))
# + id="UIDynoA_uWEg"
X,y = load_data()
# + colab={"base_uri": "https://localhost:8080/"} id="xP-3naBVx4cm" outputId="d4e5b043-ac47-4527-9410-c4d42c63681a"
X_train , X_validation ,X_test , y_train ,y_validation , y_test= prepared_dataset(0.25 , 0.2)
input_shape = (X_train.shape[1], X_train.shape[2], 1)
model = build_model(input_shape)
# compile model
optimiser = keras.optimizers.RMSprop(learning_rate=0.0001)
model.compile(optimizer=optimiser, loss='binary_crossentropy', metrics=['accuracy'])
model.summary()
# train model
history = model.fit(X_train, y_train, validation_data=(X_validation, y_validation), batch_size=32, epochs=30)
model.save("model.h5")
# + id="oSOaiXYmz6KZ"
def plot_graph(history , string):
plt.plot(history.history[string])
plt.plot(history.history["val_"+string])
plt.xlabel(string)
plt.ylabel("val_"+string)
plt.legend([string , "val_"+string])
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 541} id="v5kued0I2J9q" outputId="8178e154-4b9c-422b-a59e-bce35394cd04"
plot_graph(history , "accuracy")
plot_graph(history , "loss")
| notebooks/CNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Geographical Original of Music
# all required imports
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import numpy as np
from scipy.stats import norm
from sklearn.preprocessing import StandardScaler
from scipy import stats
import warnings
import statistics
import math
warnings.filterwarnings('ignore')
# %matplotlib inline
pd.set_option('display.max_rows', 500)
pd.options.display.max_columns = None
# +
# adding coloumn names
collist = ['col_1', 'col_2', 'col_3', 'col_4', 'col_5', 'col_6', 'col_7', 'col_8', 'col_9', 'col_10', 'col_11', 'col_12', 'col_13', 'col_14', 'col_15', 'col_16', 'col_17', 'col_18', 'col_19', 'col_20', 'col_21', 'col_22', 'col_23', 'col_24', 'col_25', 'col_26', 'col_27', 'col_28', 'col_29', 'col_30', 'col_31', 'col_32', 'col_33', 'col_34', 'col_35', 'col_36', 'col_37', 'col_38', 'col_39', 'col_40', 'col_41', 'col_42', 'col_43', 'col_44', 'col_45', 'col_46', 'col_47', 'col_48', 'col_49', 'col_50', 'col_51', 'col_52', 'col_53', 'col_54', 'col_55', 'col_56', 'col_57', 'col_58', 'col_59', 'col_60', 'col_61', 'col_62', 'col_63', 'col_64', 'col_65', 'col_66', 'col_67',"col_68","latitude" , "longitude"]
data = pd.read_csv("./dataset/default_features_1059_tracks (copy).csv",names = collist)
# -
data.info()
# +
data.head()
# -
data.describe()
statistics.mean(data["col_1"])
df1 = pd.DataFrame(data.describe())
F= pd.Series(df1['col_1'])
F[1]
df1
df1['col_1']
df1["col_1"][3]
# +
# data.isnull()
# -
data.isnull().sum()
print(data["latitude"].skew())
print(data["latitude"].kurtosis())
print(data["longitude"].skew())
print(data["longitude"].kurtosis())
x = math.cos(data["latitude"][0]) * math.cos(data["longitude"][0])
print(x)
y = math.cos(data["latitude"][0]) * math.sin(data["longitude"][0])
z= math.sin(data["latitude"][0])
print(x,y,z)
x=[]
y=[]
z=[]
for i in range(len(data["latitude"])):
x.append(math.cos(data["latitude"][i]) * math.cos(data["longitude"][i]))
y.append(math.cos(data["latitude"][i]) * math.sin(data["longitude"][i]))
z.append(math.sin(data["latitude"][i]))
print(x)
# +
num_cols = data._get_numeric_data().columns
print("Numerical Columns",num_cols)
cat_cols=list(set(data.columns) - set(num_cols))
print("Categorical Columns:",cat_cols)
# -
z_scores = np.abs(stats.zscore(data))
threshold =3
print(np.where(z_scores > 3))
print(z_scores)
print(z_scores[0][0])
fig, ax = plt.subplots(figsize=(16,8))
ax.scatter(data["latitude"], data["longitude"])
ax.set_xlabel('Latitude')
ax.set_ylabel('Longitude')
sns.boxplot(x=data["col_1"])
refined_data = data[(z_scores < 3).all(axis=1)]
fig, ax = plt.subplots(figsize=(16,8))
ax.scatter(refined_data["latitude"], refined_data["longitude"])
ax.set_xlabel('Latitude')
ax.set_ylabel('Longitude')
refined_data.shape
Q1 = data.quantile(0.25)
Q3 = data.quantile(0.75)
IQR = Q3 - Q1
print(IQR)
print(data < (Q1 - 1.5 * IQR)) |(data > (Q3 + 1.5 * IQR))
data_iqr = data[~((data < (Q1 - 1.5 * IQR)) |(data > (Q3 + 1.5 * IQR))).any(axis=1)]
data_iqr.shape
fig, ax = plt.subplots(figsize=(16,8))
ax.scatter(data_iqr["latitude"], data_iqr["longitude"])
ax.set_xlabel('Latitude')
ax.set_ylabel('Longitude')
# # Building Model
# +
X = data.iloc[:, 0:68].values
y = data.iloc[:,68:70].values
# -
import matplotlib
from sklearn.model_selection import train_test_split
from sklearn.cluster import KMeans
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.20)
print(X[0 ,1])
print(y[0 ,0:2])
# print(y)
# ### kmeans
#
# +
n_clusters = 33
kmeans = KMeans(n_clusters=n_clusters, max_iter = 500, algorithm='full')
kmeans = kmeans.fit(y)
labels = kmeans.predict(y)
centroids = kmeans.cluster_centers_
print(centroids)
print(labels)
# +
label_color = [matplotlib.cm.nipy_spectral(float(l) /n_clusters) for l in labels]
plt.scatter(y[:, 1], y[:, 0], c = label_color, s=25)
plt.title("Clustered regions by coordinates")
plt.xlabel("Longitude")
plt.ylabel("Latitude")
plt.figure()
# -
print(y)
# +
from sklearn.cluster import DBSCAN
plt.rcParams["figure.figsize"] = (20,5)
dbs = DBSCAN(eps=3, min_samples=2).fit(y)
core_samples_mask = np.zeros_like(dbs.labels_, dtype=bool)
core_samples_mask[dbs.core_sample_indices_] = True
dbs_labels = dbs.labels_
n_clusters_ = len(set(labels)) - (1 if -1 in labels else 0)
n_noise_ = list(labels).count(-1)
print(n_clusters)
# +
unique_labels = set(labels)
colors = [plt.cm.Spectral(each)
for each in np.linspace(0, 1, len(unique_labels))]
for k, col in zip(unique_labels, colors):
if k == -1:
# Black used for noise.
col = [0, 0, 0, 1]
class_member_mask = (labels == k)
xy = X[class_member_mask & core_samples_mask]
plt.plot(xy[:, 0], xy[:, 1], 'o', markerfacecolor=tuple(col),
markeredgecolor='k', markersize=14)
xy = X[class_member_mask & ~core_samples_mask]
plt.plot(xy[:, 0], xy[:, 1], 'o', markerfacecolor=tuple(col),
markeredgecolor='k', markersize=6)
plt.title('Estimated number of clusters: %d' % n_clusters_)
plt.show()
# +
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
scaler.fit(X_train)
X_train = scaler.transform(X_train)
X_test = scaler.transform(X_test)
# +
# from sklearn.neighbors import KNeighborsClassifier
# classifier = KNeighborsClassifier(n_neighbors=5)
# classifier.fit(y_train)
# -
from sklearn.metrics import classification_report, confusion_matrix
# print(confusion_matrix(y_test, y_pred))
# print(classification_report(y_test, y_pred))
# +
error = []
# Calculating error for K values between 1 and 40
for i in range(1, 40):
kmeans = KMeans(n_clusters=i, max_iter = 500, algorithm='full')
kmeans = kmeans.fit(y)
labels = kmeans.predict(y)
# error.append(np.mean(labels != ))
# +
# plt.figure(figsize=(12, 6))
# plt.plot(range(1, 40), error, color='red', linestyle='dashed', marker='o',
# markerfacecolor='blue', markersize=10)
# plt.title('Error Rate K Value')
# plt.xlabel('K Value')
# plt.ylabel('Mean Error')
# +
# scores = cross_val_score(estimator_iter, Xtrain, ytrain, cv = 2, scoring='accuracy')
# -
y = labels
from sklearn.svm import SVC
from sklearn.model_selection import StratifiedKFold
from sklearn.feature_selection import RFECV
from sklearn.feature_selection import RFE
from sklearn.datasets import make_classification
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
from sklearn.grid_search import ParameterGrid
from sklearn.decomposition import PCA
from sklearn.linear_model import LogisticRegression
from sklearn.preprocessing import StandardScaler
# +
Xtrain, Xtest, ytrain, ytest = train_test_split(X, y, test_size=0.8, random_state=1234)
X_scaler = StandardScaler().fit(X)
scaled_Xtrain = X_scaler.transform(Xtrain)
scaler = StandardScaler()
scaled_Xtest = X_scaler.transform(Xtest)
# +
PCA_var = PCA(n_components = X_scaler.transform(X).shape[1])
PCA_var.fit(X_scaler.transform(X))
PCA_var_exp = [1 - x/sum(PCA_var.explained_variance_) for x in PCA_var.explained_variance_]
PCA_var_exp.insert(0, 0)
plt.figure()
plt.title("PCA variance explained over number of PCs")
plt.xlabel("Number of PCs")
plt.ylabel("Ratio of variance explained")
plt.plot(range(0, len(PCA_var_exp), 1), PCA_var_exp, c = 'r')
plt.show()
# +
estimators = {}
estimators['svc'] = SVC
estimators['rfc'] = RandomForestClassifier
estimators['logreg'] = LogisticRegression
params = {}
params['svc'] = {'kernel': ['linear'], 'C': [10**x for x in range(-1, 3, 1)],
'gamma': [10**x for x in range(-1, 2, 1)],
'random_state': [1234]}
params['rfc'] = {'n_estimators': [5*x for x in range(3, 5, 1)]}
params['logreg'] = {'C': [10**x for x in range(-1, 3, 1)], 'penalty': ['l1', 'l2']}
rfe_best_model = {}
pca_best_model = {}
for i, estimator in enumerate(estimators):
model_params = ParameterGrid(params[estimator])
grid = model_params
rfe_best_model[estimator] = []
pca_best_model[estimator] = []
for n_comps in range(1, scaled_Xtrain.shape[1], 1):
rfe_best_model[estimator].append(0)
pca_best_model[estimator].append(0)
for params_combo in grid:
estimator_iter = estimators[estimator]
estimator_iter = estimator_iter(**params_combo)
rfe = RFE(estimator = estimator_iter, n_features_to_select=n_comps)
rfe.fit(scaled_Xtrain, ytrain)
if (rfe.score(scaled_Xtest, ytest) > rfe_best_model[estimator][n_comps - 1]):
rfe_best_model[estimator][n_comps - 1] = rfe.score(scaled_Xtest, ytest)
PCA_model = PCA(n_components = n_comps)
PCA_model.fit(scaled_Xtrain)
PCA_Xtrain = PCA_model.transform(scaled_Xtrain)
PCA_Xtest = PCA_model.transform(scaled_Xtest)
estimator_iter.fit(PCA_Xtrain, ytrain)
if (estimator_iter.score(PCA_Xtest, ytest) > pca_best_model[estimator][n_comps - 1]):
pca_best_model[estimator][n_comps - 1] = estimator_iter.score(PCA_Xtest, ytest)
plt.figure()
plt.xlabel("%s - Number of features selected" % estimator)
plt.ylabel("Accuracy score")
plt.plot(range(1, len(pca_best_model[estimator]) + 1, 1), pca_best_model[estimator], c = 'r')
plt.plot(range(1, len(rfe_best_model[estimator]) + 1, 1), rfe_best_model[estimator], c = 'b')
plt.show()
# -
from factor_analyzer import FactorAnalyzer
# https://www.datacamp.com/community/tutorials/introduction-factor-analysis
from factor_analyzer.factor_analyzer import calculate_bartlett_sphericity
chi_square_value,p_value=calculate_bartlett_sphericity(data)
chi_square_value, p_value
from factor_analyzer.factor_analyzer import calculate_kmo
kmo_all,kmo_model=calculate_kmo(data)
kmo_model
# Create factor analysis object and perform factor analysis
fa = FactorAnalyzer(rotation = None)
fa.fit(data)
# Check Eigenvalues
ev, v = fa.get_eigenvalues()
ev
# +
# Create factor analysis object and perform factor analysis
fa = FactorAnalyzer( n_factors=25,rotation = None)
fa.fit(data)
# Check Eigenvalues
ev, v = fa.get_eigenvalues()
ev
# -
df = data
# Create scree plot using matplotlib
plt.scatter(range(1,df.shape[1]+1),ev)
plt.plot(range(1,df.shape[1]+1),ev)
plt.title('Scree Plot')
plt.xlabel('Factors')
plt.ylabel('Eigenvalue')
plt.grid()
plt.show()
# +
fa = FactorAnalyzer(n_factors = 20,rotation = "varimax")
fa.fit(data)
# -
fa.loadings_
fa.get_factor_variance()
# Pearson Correlation Test
#
# +
from sklearn.model_selection import train_test_split
data = data.drop(data.index[1])
X = data.iloc[:, 0:68].values
y = data.iloc[:,68:70].values
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.50)
# +
# X_train.shape
# X_test.shape
# +
# X_test.shape
# +
# sample1 = data.iloc[:529 , :]
# sample2 = data.iloc[529:1058 , :]
# -
https://machinelearningmastery.com/statistical-hypothesis-tests-in-python-cheat-sheet/
# +
from scipy.stats import pearsonr
corr , p = pearsonr(sample1 ,sample2)
# -
sample1 = data.sample(frac=0.5, replace=True, random_state=1)
sample1
sample2 = data.sample(frac=0.5, replace=True, random_state=2)
sample2
corr , p = pearsonr(sample1["col_1"] ,sample2["col_2"])
corr
# T-Test
from scipy.stats import ttest_ind
stat, p = ttest_ind(sample1, sample2)
if p.all() > 0.05:
print('Probably the same distribution')
else:
print('Probably different distributions')
stat
from scipy.stats import ttest_rel
stat, p = ttest_rel(sample1, sample2)
if p.all() > 0.05:
print('Probably the same distribution')
else:
print('Probably different distributions')
stat
p
# Analysis of Variance Test (ANOVA)
from scipy.stats import f_oneway
stat, p = f_oneway(sample1, sample2)
# print('stat=%.3f, p=%.3f' % (stat, p))
if p.all() > 0.05:
print('Probably the same distribution')
else:
print('Probably different distributions')
# Test for Equal variances
# +
from scipy.stats import levene
levene(sample1["col_1"],sample2["col_1"])
# -
# Homoscedasticity (equal variance) of residuals
# +
import seaborn as sns
import matplotlib.pyplot as plt
import statsmodels.stats.api as sms
sns.set_style('darkgrid')
sns.mpl.rcParams['figure.figsize'] = (15.0, 9.0)
from sklearn.linear_model import LinearRegression
lin_reg = LinearRegression()
lin_reg.fit(X, y)
def homoscedasticity_test(model):
'''
Function for testing the homoscedasticity of residuals in a linear regression model.
It plots residuals and standardized residuals vs. fitted values and runs Breusch-Pagan and Goldfeld-Quandt tests.
Args:
* model - fitted OLS model from statsmodels
'''
fitted_vals = model.predict(X)
resids = model.resid
resids_standardized = model.get_influence().resid_studentized_internal
fig, ax = plt.subplots(1,2)
sns.regplot(x=fitted_vals, y=resids, lowess=True, ax=ax[0], line_kws={'color': 'red'})
ax[0].set_title('Residuals vs Fitted', fontsize=16)
ax[0].set(xlabel='Fitted Values', ylabel='Residuals')
sns.regplot(x=fitted_vals, y=np.sqrt(np.abs(resids_standardized)), lowess=True, ax=ax[1], line_kws={'color': 'red'})
ax[1].set_title('Scale-Location', fontsize=16)
ax[1].set(xlabel='Fitted Values', ylabel='sqrt(abs(Residuals))')
bp_test = pd.DataFrame(sms.het_breuschpagan(resids, model.model.exog),
columns=['value'],
index=['Lagrange multiplier statistic', 'p-value', 'f-value', 'f p-value'])
gq_test = pd.DataFrame(sms.het_goldfeldquandt(resids, model.model.exog)[:-1],
columns=['value'],
index=['F statistic', 'p-value'])
print('\n Breusch-Pagan test ----')
print(bp_test)
print('\n Goldfeld-Quandt test ----')
print(gq_test)
print('\n Residuals plots ----')
homoscedasticity_test(lin_reg)
# -
| Project Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Setup
# +
# %reload_ext nb_black
import pandas as pd
import numpy as np
from scipy import stats
import statsmodels.api as sm
from statsmodels.formula.api import ols
from statsmodels.stats.multicomp import pairwise_tukeyhsd
from statsmodels.graphics.gofplots import qqplot
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
# -
# Function for determining Confidence Intervals
def ci_95(x1, x2):
signal = x1.mean() - x2.mean()
noise = np.sqrt(x1.var() / x1.size + x2.var() / x2.size)
ci_lo = signal - 1.96 * noise
ci_hi = signal + 1.96 * noise
return ci_lo, ci_hi
# Function for merging Dataframes by year
def merge_fun(year_df, year):
df = pd.DataFrame()
add_columns = []
for i in merge_columns:
add_columns.extend(x for x in year_df.columns if i in x)
df[merge_columns] = year_df[add_columns]
df["Happiness Score"] = year_df[[x for x in year_df.columns if "Score" in x]]
df["Year"] = year
df = df.set_index(["Country", "Year"])
return df
# # Exploring the data
# Load in 2019 DataFrame
url = "https://raw.githubusercontent.com/WoodyBurns44/Happiness_index_Analysis/main/2019.csv"
df_2019 = pd.read_csv(url)
df_2019.head()
# See what data we're working with
df_2019.info()
df_2019.shape
df_2019.index
df_2019.columns
df_2019.describe()
# # Is there a correlation between the GDP per capita in 2019 for the happiest and less happy countries? If so, how strong is the correlation?
# ## Divide the dataset into two groups: happy countries and less happy Countries based on the median happiness score.
# Split by median to get as close to equal sized data as possible
median = df_2019["Score"].median()
# Select the "Happiest" countries, above the median
happy_2019 = df_2019[df_2019["Score"] > median]
happy_2019.shape
# Select the less happy countries, below the median
unhappy_2019 = df_2019[df_2019["Score"] <= median]
unhappy_2019.shape
# In order to determine which test will most accurately find whether there is a correlation or not, I must look into the distribution of the data. First, I will test both variables for normality.
# ## Test for normality
# Find the mean and medians of the data sets to see if they tell us anything abou ttheir normality
happy_2019["GDP per capita"].mean()
happy_2019["GDP per capita"].median()
unhappy_2019["GDP per capita"].mean()
unhappy_2019["GDP per capita"].median()
# The means and medians of both samples are fairly close to eachother, which is an indication
# that the data might be normally distributed.
# Check the Kurtosis and Skewness
stats.describe(unhappy_2019["GDP per capita"])
stats.describe(happy_2019["GDP per capita"])
# Histogram of the GDP per capita of happy countries, with a black line showing the mean
# and an orange line showing the median.
happy_2019["GDP per capita"].hist()
plt.axvline(x=happy_2019["GDP per capita"].median(), c="orange", linestyle="solid")
plt.axvline(x=happy_2019["GDP per capita"].mean(), c="black", linestyle="solid")
plt.show()
# Histogram of the GDP per capita of less happy countries, with a black line showing the mean
# and an orange line showing the median.
unhappy_2019["GDP per capita"].hist()
plt.axvline(x=unhappy_2019["GDP per capita"].median(), c="orange", linestyle="solid")
plt.axvline(x=unhappy_2019["GDP per capita"].mean(), c="black", linestyle="solid")
plt.show()
## QQ plot to visualize happy countries GDP per capita relation to normal distribution
qqplot(happy_2019["GDP per capita"], line="s")
plt.show()
## QQ plot to visualize less happy countries Happiness scores relation to normal distribution
qqplot(unhappy_2019["GDP per capita"], line="s")
plt.show()
# Check normality with a Violin plot
sns.violinplot(x="GDP per capita", data=unhappy_2019, color="orange")
sns.violinplot(x="GDP per capita", data=happy_2019)
plt.show()
j, p = stats.jarque_bera(unhappy_2019["GDP per capita"])
j
p < 0.05
# The result of the Jarque-Bera test indicates that the distribution is not perectly normal. However, since the sample size is small and all of the other tests indicate normality, I will treat it as normal.
# Since both happy and less happy countries GDP per capita appear to be normal, I will perform a Students T-test to determine if there is variance between the groups.
# ## Student T-Test
# The Students T-test is used to detect if the means are different between two groups.
# * $H_o$ : Both developing and developed countries have the same mean of GDP per capita
# * $H_a$ : The mean of GDP per capita differs between developing and developed countries
ttest_score, ttest_p = stats.ttest_ind(
happy_2019["GDP per capita"], unhappy_2019["GDP per capita"]
)
ttest_score, ttest_p
ttest_p < 0.05
# The Students T-test indicates that the null hypothesis can be rejected and that the distribution of GDP per capita differs between happy and less happy countries.
# ## How significant is the difference in GDP per capita between the happiest and less happy countries?
# In order to determine the difference in means between happy and less happy countries' GDP per Capital I will calculate a confidence interval and then bootstrap to test that calculation.
# Calculating low and high confidence intervals using function defined above
ci_95(happy_2019["GDP per capita"], unhappy_2019["GDP per capita"])
# I can say with 95% confidence that there is a .45 and .63 difference in the GDP per Capita of happy and less happy countries, in favor of the more happy countries.
# +
# Testing confidence interval with a bootstrap
mean_diffs = []
for i in range(10000):
control_sample = happy_2019["GDP per capita"].sample(frac=1.0, replace=True)
treatment_sample = unhappy_2019["GDP per capita"].sample(frac=1.0, replace=True)
mean_diff = control_sample.mean() - treatment_sample.mean()
mean_diffs.append(mean_diff)
# -
low_ci = np.percentile(mean_diffs, 2.5)
high_ci = np.percentile(mean_diffs, 97.5)
low_ci, high_ci
# The sample bootstrapping confirms my calculations that there is, with 95% certainty, a .45 and .63 difference in the GDP per Capita of happy and less happy countries, in favor of the more happy countries.
# # Has the happiness of the world changed from 2015 to 2019? If so, in what way and how much?
# ## Investigate the DataFrame for 2015 data
# Use Pandas to import the 2015 Dataframe
url_2015 = "https://raw.githubusercontent.com/WoodyBurns44/Happiness_index_Analysis/main/2015.csv"
df_2015 = pd.read_csv(url_2015)
df_2015.head()
df_2015.info()
df_2015.describe()
df_2015.columns
# ## Isolate the Happiness columns for 2015 and 2019
# Create Dataframes for the 2015 and 2019 data
happy15 = df_2015["Happiness Score"]
# happy15.head()
happy15.index
happy19 = df_2019["Score"]
# happy19.head()
happy19.index
# ## Test for normality of happiness in 2015 and 2019
# ### 2015
happy15.mean()
happy15.hist()
plt.axvline(x=happy15.mean(), c="orange")
plt.axvline(x=happy15.median(), c="black")
plt.show()
qqplot(happy15, line="s")
plt.show()
# Use the Jarque-Bera test for normality
# * $H_o$ : The data comes from a normally distributed set.
# * $H_a$ : The data does not come form a normally distributed set.
j15, p15 = stats.jarque_bera(happy15)
j15
p15
p < 0.05
# Since we cannot reject the null, it can be assumed that the data comes from a normal distribution.
# ### 2019
happy19.mean()
happy19.hist()
plt.axvline(x=happy19.mean(), c="orange")
plt.axvline(x=happy19.median(), c="black")
plt.show()
qqplot(happy19, line="s")
plt.show()
j19, p19 = stats.jarque_bera(happy19)
j19
p19
# Since we cannot reject the null, it can be assumed that the data comes from a normal distribution.
# ## Perform an independent T-test
t_15_to_19, p_15_to_19 = stats.ttest_ind(happy15, happy19)
t_15_to_19
p_15_to_19
# ## Result
# There does not appear to be a significant diffance in the overall happiness of the world in 2019 as compared with 2015.
# # Which factors are most strongly correlated to the overall happiness score in 2019?
# Make a Spearman Correlatoin matrix to test for correlations between all numeric categories
spearman_correlations = df_2019.corr(method="spearman")
spearman_correlations
# Translate matrix into a heatmap for better visualization of the correlations.
#
# * code for heatmap inspired by <NAME> on Kaggle (https://www.kaggle.com/jesperdramsch/the-reason-we-re-happy).
fig, ax = plt.subplots(ncols=2, figsize=(24, 8))
sns.heatmap(
spearman_correlations,
vmin=-1,
vmax=1,
ax=ax[0],
center=0,
cmap="viridis",
annot=True,
)
sns.heatmap(
spearman_correlations,
vmin=-0.25,
vmax=1,
ax=ax[1],
center=0,
cmap="Accent",
annot=True,
linecolor="white",
)
# The heat map gives us a lot of information, including:
# * Validates that there is a strong correlation between GDP per capita and happiness score
# * Shows that there is very strong correlation between the following fields and the happiness score:
# * Social Support
# * Healthy Life Expectancy
# * Freedom to make Life choices
# # Appendix: Further Exploration
url_2016 = "https://raw.githubusercontent.com/WoodyBurns44/Happiness_index_Analysis/main/2016.csv"
url_2017 = "https://raw.githubusercontent.com/WoodyBurns44/Happiness_index_Analysis/main/2017.csv"
url_2018 = "https://raw.githubusercontent.com/WoodyBurns44/Happiness_index_Analysis/main/2018.csv"
df_2016 = pd.read_csv(url_2016)
df_2017 = pd.read_csv(url_2017)
df_2018 = pd.read_csv(url_2018)
df_2016.head(1)
df_2015.head(1)
# +
# Template Columns for merging table
merge_columns = [
"Country",
"GDP",
"Family",
"Life",
"Freedom",
"Generosity",
"Trust",
]
# -
# Format 2015 Dataframe
df = merge_fun(df_2015, 2015)
# Format and merge 2016 Dataframe
df = df.append(merge_fun(df_2016, 2016), sort=False)
# Format and Merge 2017 Dataframe
df = df.append(merge_fun(df_2017, 2017), sort=False)
# Rename columns in 2018 Datframe so that they work with merge function
df_2018 = df_2018.rename(
columns={
"Healthy life expectancy": "Life",
"Perceptions of corruption": "Trust",
"Social support": "Family",
},
)
# Merge 2018 Dataframe
df = df.append(merge_fun(df_2018, 2018), sort=False)
# Rename 2019 columns so that they work with merge function
df_2019 = df_2019.rename(
columns={
"Social support": "Family",
"Healthy life expectancy": "Life",
"Perceptions of corruption": "Trust",
}
)
# Merge 2019 Dataframe
df = df.append(merge_fun(df_2019, 2019), sort=False)
df.head()
df = df.rename(columns={"Happiness Score": "Happiness_Score"})
# Reset Index for plotability
df_test = df.reset_index()
df_test.head()
# Plot yearly change of Happiness Score for insights
sns.set(rc={"figure.figsize": (11.7, 8.27)})
happy_plot = sns.lineplot(
x="Year",
y="Happiness_Score",
hue="Country",
legend="brief",
data=df_test,
)
# happy_plot.legend(loc=10)
| happiness_index_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Surface defect pattern that can cause failure.
# Generate defect samples.
# import the required libraries
import math
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
# +
#sample preparation
#27.5 mm radius, perimeter = 2*pi*r
#generate a random array with 0 or 1 to represent the defect
# x= =100, y = 2*pi()*100
radius = 28
perimeter = int(2*math.pi*radius)
limit = 1001
threshold = limit - 1
#generating a 2D random array
X = np.random.randint(0,limit, size=(100, radius,perimeter)) #0 or 1, exluding 2
X = np.where((X<threshold), 0, X)
X = np.where((X!=0),1, X)
#print(sample)
plt.show()
X.shape
# +
# failed defect patterns
radius = 28
perimeter = int(2*math.pi*radius)
limit = 1001
threshold = limit - 10
#generating a 2D random array
F = np.random.randint(0,limit, size=(100, radius,perimeter)) #0 or 1, exluding 2
F = np.where((F<threshold), 0, F)
F = np.where((F!=0),1, F)
#print(sample)
plt.show()
F.shape
# +
#visualize the defect pattern
for i in range(1,2):
plt.figure(figsize=(20, 4))
print('sample:', i,F[i].sum())
plt.imshow(F[i], cmap=plt.cm.binary)
plt.show()
# -
sample = np.concatenate((X, F), axis=0) # combine all the samples
sample.shape
sample_results = []
for i in range(1,101):
sample_results.append(0)
for i in range(101,201):
sample_results.append(1)
sample_results = np.array(sample_results)
print(type(sample_results))
print(sample_results.shape)
type(sample_results)
# +
# Shuffle the sample
shuffler = np.random.permutation(len(sample))
sample_shuffled = sample[shuffler]
sample_results_shuffled = sample_results[shuffler]
# -
i=100
sample_shuffled[i]
sample_results_shuffled[i]
plt.figure(figsize=(20, 4))
plt.imshow(sample_shuffled[i], cmap=plt.cm.binary)
plt.show()
sample_results_shuffled.shape
x_train = sample[:100]
x_test = sample[100:]
y_train = sample_results[:100]
y_test = sample_results[100:]
print(x_train.shape)
print(x_test.shape)
print(y_train.shape)
print(y_test.shape)
x_train1 = x_train.reshape(100,4900)
y_train1 = y_train.reshape(100,1)
x_test1 = x_test.reshape(100,4900)
y_test1 = y_test.reshape(100,1)
testing.shape
# +
from tensorflow.keras import models
from tensorflow.keras import layers
network = models.Sequential()
network.add(layers.Dense(16, activation='relu', input_shape=(28*175,)))
network.add(layers.Dense(1, activation='sigmoid'))
network.summary()
# -
network.compile(optimizer='rmsprop',
loss='categorical_crossentropy',
metrics=['accuracy'])
network.fit(x_train1, y_train1, epochs=5, batch_size=128)
test_loss, test_acc = network.evaluate(x_test1, y_test1, verbose=2)
print('test_acc:', test_acc)
| DefectPattern.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import re
# negative altitudes
# regular expressions for numbers (x.0 !!)
#np.where()
# -
random_numbers=pd.read_csv('data4.txt',header=None,nrows=1)
bingo_tables=pd.read_csv('data4.txt',header=None,delimiter='\s+',skiprows=1)
random_numbers.shape,bingo_tables.shape
bingo_tables=np.array(bingo_tables)
bingo_tables=bingo_tables.reshape(100,5,5)
bingo_tables_long=np.ndarray.flatten(bingo_tables)
random_numbers=np.array(random_numbers)
random_numbers=random_numbers.reshape(100)
random_numbers=pd.Series(random_numbers)
np.sum([[1,2],[3,4]],axis=1)
test=[[1,2,,4,5],[5,6,7,8,9]]
# +
bingo_tables=np.array(bingo_tables)
bingo_tables=bingo_tables.reshape(100,5,5)
bingo_tables_long=np.ndarray.flatten(bingo_tables)
tensor=np.array([np.zeros(5*5*100)])
record_tensor=tensor.reshape(100,5,5)
record_tensor_long=np.ndarray.flatten(record_tensor)
stop=[]
for draw in random_numbers:
for j in range(0,2500):
if draw == bingo_tables_long[j]:
record_tensor_long[j]=1
for k in range(0,100):
if 5 in np.sum(record_tensor_long.reshape(100,5,5)[k],axis=1) or 5 in np.sum(record_tensor_long.reshape(100,5,5)[k],axis=0):
#print('tensor:',k,'draw:',draw)
#print(record_tensor_long.reshape(100,5,5)[k])
winner_record=record_tensor_long.reshape(100,5,5)[k]
winner_table=bingo_tables_long.reshape(100,5,5)[k]
stop=1
if stop==1:
for j in range(0,5):
for k in range(0,5):
if winner_record[k,j]==1:
winner_table[k,j]=0
print('result for first winner:',sum(winner_table).sum()*draw)
break
# +
bingo_tables=np.array(bingo_tables)
bingo_tables=bingo_tables.reshape(100,5,5)
bingo_tables_long=np.ndarray.flatten(bingo_tables)
tensor=np.array([np.zeros(5*5*100)])
record_tensor=tensor.reshape(100,5,5)
record_tensor_long=np.ndarray.flatten(record_tensor)
stop=[]
order_of_winners=np.array(np.zeros(100))
score_list=[]
counter=0
for p,draw in enumerate(random_numbers):
for j in range(0,2500-counter*25):
if draw == bingo_tables_long[j]:
record_tensor_long[j]=1
for k in range(0,100):
if 5 in np.sum(record_tensor_long.reshape(100-counter,5,5)[k],axis=1) or 5 in np.sum(record_tensor_long.reshape(100-counter,5,5)[k],axis=0):
#print('tensor:',k,'draw:',draw)
#print(record_tensor_long.reshape(100,5,5)[k])
winner_record=record_tensor_long.reshape(100,5,5)[k]
winner_table=bingo_tables_long.reshape(100,5,5)[k]
counter=1
bingo_tables=np.delete(bingo_tables,k,0)
record_tenser=np.delete(record_tenser,k,0)
record_tensor_long=np.ndarray.flatten(record_tensor)
order_of_winners[k]=1
stop=1
if stop==1:
for j in range(0,5):
for k in range(0,5):
if winner_record[k,j]==1:
winner_table[k,j]=0
score=sum(winner_table).sum()*draw
score_list.append(score)
stop=0
print('result for first winner:',score)
# -
order_of_winners=np.array(np.zeros(100))
score
bingo_tables=np.delete(bingo_tables,0,0)
bingo_tables
bingo_tables[49]
bingo_tables[51][1,0]
result=np.array([[1,0,1,0,0],[1,1,0,0,0],[1,1,1,1,1],[0,1,0,0,0],[0,0,0,0,0]])
result[1,1]
sum(bingo_tables[49]).sum()*41
| Challenge4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import torch
from torch.autograd import Variable
#
# # Tensor VS Variable
# +
tensor = torch.rand(2,2,)
x = Variable(tensor, requires_grad=True)
tensor, x
# -
x.sum(), x.sqrt(), x**2, x
#
# # Variable
x.data
x.requires_grad, x.grad, x.grad_fn, x.volatile
# +
x = Variable(torch.FloatTensor([3]), requires_grad=True)
y = x**3 + 3
z = Variable(torch.FloatTensor([5]))
x.requires_grad, y.requires_grad, z.requires_grad, y
# -
y.grad_fn.next_functions[0][0]
y.backward()
x, x.grad
x.grad, x.grad.data.zero_(), x.grad
x1 = Variable(torch.ones(2), requires_grad = True)
x2 = Variable(torch.ones(2))
y = x1 * 2 + x2
y
y.backward(torch.Tensor([1,1]))
x1.grad
x2.grad
x1.grad.data.zero_()
| 02_variable_autograd.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/hhk54250/20MA573-HHK/blob/master/Untitled7.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="D8HTeE2DB_i2" colab_type="code" colab={}
import numpy.matlib
import numpy as np
# + id="4IemwsySCLhV" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="5306c98c-6f99-4140-fdde-d0818554ef58"
import random
A=0
B=2*np.pi
n= 0
for i in range(1000000):
c=[]
a= random.uniform(A,B)
b= random.uniform(A,B)
if a>b:
c1=b
c2=a-b
c3=B-a
if b>a:
c1=a
c2=b-a
c3=B-b
c.append(c1)
c.append(c2)
c.append(c3)
l=np.max(c)
if l<np.pi:
n=n+1
print(str(n/1000000))
| Untitled7.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Importing modules
# +
import requests
import urllib.request
import time
from bs4 import BeautifulSoup
import pandas as pd
import os
from tqdm import tqdm_notebook as tqdm
import numpy as np
import string
dir_path = os.getcwd()
data_dir = "data/"
# -
# # Scraping urls and video_ids
# +
# Retrieve all partij urls
videos = ["/watch?v=0vFTfoOavRM&list=UUt7fwAhXDy3oNFTAzF2o8Pw&index=1"]
index = ["ex=1"]
title = ["Perfume Genius - Set My Heart on Fire Immediately ALBUM REVIEW"]
while len(videos) < 3238:
response = requests.get(base_url + videos[-1])
soup = BeautifulSoup(response.text, "html.parser")
next_video = soup.find_all('a', {"class": "spf-link playlist-video clearfix yt-uix-sessionlink spf-link"})
for nv in next_video:
if nv["href"][-4:] not in index:
videos.append(nv["href"])
index.append(nv["href"][-4:])
title.append(nv.find("h4").text.split("\n")[1].replace(" ", ""))
print(len(videos), end="\r")
# -
df_url = pd.DataFrame(columns = ["url", "id", "title"])
df_url.url = videos
df_url.id = df_url.url.str.split("&").apply(lambda x: x[0].replace("/watch?v=", ""))
df_url.title = title
df_url.to_csv(data_dir + "urls.csv", index = None, sep = ";")
# +
df_url = pd.read_csv(data_dir + "urls.csv", sep = ";")
df_url.title = df_url.title.str.lower()
filter1 = df_url.title.str.contains("review")
filter2 = df_url.title.str.contains("-")
df_title = df_url.loc[(filter1 & filter2), ["id", "title"]]
df_title.title = df_title.title.str.replace("album", "")
df_title.title = df_title.title.str.replace("review", "")
filter3 = df_url.title.str.contains("not good")
df_notgood = df_url.loc[filter3, ["id", "title"]]
df_notgood.title = df_notgood.title.str.replace("not good", "")
df_notgood.title = df_notgood.title.str.replace(": ", "")
# -
df_artist_album = pd.DataFrame(columns = ["id", "artist", "album"])
df_artist_album.id = df_title.id
df_artist_album.artist = df_title.title.str.split("-").apply(lambda x: x[0]).str.strip()
df_artist_album.album = df_title.title.str.split("-").apply(lambda x: x[-1]).str.strip()
df_notgood_artist_album = pd.DataFrame(columns = ["id", "artist", "album"])
df_notgood_artist_album.id = df_notgood.id
df_notgood_artist_album.artist = df_notgood.title.str.split("'s").apply(lambda x: x[0]).str.strip()
df_notgood_artist_album.album = df_notgood.title.str.split("'s").apply(lambda x: x[-1]).str.strip()
df_artist_album = df_artist_album.append(df_notgood_artist_album).sort_index()
df_artist_album.to_csv(data_dir + "albums.csv", index = False, sep = ";")
| url_webscraping.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
# %cd ..
from dev.retina_copie import Retina, RetinaWhiten
#args = init(filename='../data/2019-07-23')
from easydict import EasyDict
args = EasyDict({'w': 28,
'minibatch_size': 10,
'train_batch_size': 1000,
'test_batch_size': 126,
'noise_batch_size': 1000,
'mean': 0.1307,
'std': 0.3081,
'N_pic': 1718,
'N_X': 1718,
'N_Y': 2444,
'N_X2': 98,
'N_Y2': 140,
'offset_std': 30,
'offset_max': 34,
'noise': 0.75,
'contrast': 0.7,
'sf_0': 0.1,
'B_sf': 0.1,
'N_theta': 6,
'N_azimuth': 48,
'N_eccentricity': 80,
'N_phase': 2,
'rho': 1.41,
'bias_deconv': True,
'p_dropout': 0.5,
'dim1': 1000,
'dim2': 1000,
'lr': 0.005,
'do_adam': 'adam',
'bn1_bn_momentum': 0,
'bn2_bn_momentum': 0,
'momentum': 0,
'epochs': 60,
'num_processes': 1,
'no_cuda': True,
'log_interval': 20,
'verbose': 1,
'filename': '../data/2019-07-23',
'seed': 2019,
'N_cv': 10,
'do_compute': True,
'save_model': True,
'zoomW': 300})
print('N_X :', args.N_X)
print('N_Y :', args.N_Y)
print('N_theta :', args.N_theta)
print('N_azimuth :', args.N_azimuth)
print('N_eccentricity :', args.N_eccentricity)
print('N_phase :', args.N_phase)
args.N_X, args.N_Y = 768, 1024
args_N_pic = args.N_Y
retina = Retina(args)
retina.retina_dico.keys() # theta
retina.retina_dico[0].keys() # phi
retina.retina_dico[0][0].keys() # eccentricity
shape_10 = int(np.sqrt(retina.retina_dico[0][0][10].shape[0])) # central filter
shape_23 = int(np.sqrt(retina.retina_dico[0][0][23].shape[0])) # periphery filter
plt.imshow(retina.retina_dico[2][0][10].reshape((shape_10, shape_10)))
plt.imshow(retina.retina_dico[0][0][23].reshape((shape_23, shape_23)))
plt.imshow(retina.retina_dico[2][0][23].reshape((shape_23, shape_23)))
plt.imshow(retina.retina_dico[2][1][23].reshape((shape_23, shape_23)))
from PIL import Image
impath = 'data/i05june05_static_street_boston_p1010808.jpeg'
im = Image.open(impath)
im
im_color_npy = np.asarray(im)
im_color_npy
im_bw = im.convert("L")
im_bw
im_npy = np.asarray(im_bw)
im_npy
args.N_X, args.N_Y = im_npy.shape
args_N_pic = args.N_Y
# +
plt.figure(figsize=(20, 20))
plt.imshow(im_bw)
ecc_max = .8
rho = 1.05
args.N_eccentricity = 80
N_X, N_Y = args.N_X, args.N_Y
plt.plot(N_Y / 2 ,N_X / 2 , '+r')
for i_eccentricity in range(args.N_eccentricity):
for i_azimuth in range(args.N_azimuth):
ecc = ecc_max * (1 / rho) ** ((args.N_eccentricity - i_eccentricity) )
N_min = min(N_X, N_Y)
r = np.sqrt(N_min ** 2 + N_min ** 2) / 2 * ecc #- 30 # radius
# r = np.sqrt(N_X ** 2 + N_Y ** 2) / 2 * ecc - 30 # radius
psi = (i_azimuth + (i_eccentricity % 2) * .5) * np.pi * 2 / args.N_azimuth
x = int(N_X / 2 + r * np.cos(psi))
y = int(N_Y / 2 + r * np.sin(psi))
plt.plot(y, x , '.r')
# -
whiten_transform = RetinaWhiten(args)
im_whiten = whiten_transform(im_npy)
plt.figure(figsize=(20,20))
plt.imshow(im_whiten, cmap='gray')
retina_features = retina.transform(im_whiten)
retina_features.shape
plt.plot(retina_features)
img_trans = retina.inverse_transform(retina_features)
plt.figure(figsize=(20,20))
plt.imshow(img_trans, cmap='gray')
plt.figure(figsize=(10,20))
plt.subplot(121)
plt.imshow(im_whiten[args.N_X//2-50:args.N_X//2 + 50, args.N_Y//2-50:args.N_Y//2 + 50], cmap='gray')
plt.subplot(122)
plt.imshow(img_trans[args.N_X//2-50:args.N_X//2 + 50, args.N_Y//2-50:args.N_Y//2 + 50], cmap='gray')
# +
from LogGabor import LogGabor
N_theta=6
rho = 1.41
ecc_max = .8 # self.args.ecc_max
sf_0_r = 0.03 # self.args.sf_0_r
sf_0_max = 0.45
B_theta = np.pi / N_theta / 2 # self.args.B_theta
B_sf = .4
pe = {'N_image': 100, 'seed': None, 'N_X': 512, 'N_Y': 512, 'noise':
0.1, 'do_mask': True, 'mask_exponent': 3.0, 'do_whitening': True,
'white_name_database': 'kodakdb', 'white_n_learning': 0, 'white_N':
0.07, 'white_N_0': 0.0, 'white_f_0': 0.4, 'white_alpha': 1.4,
'white_steepness': 4.0, 'white_recompute': False, 'base_levels':
1.618, 'n_theta': 24, 'B_sf': 0.4, 'B_theta': 0.17453277777777776,
'use_cache': True, 'figpath': 'results', 'edgefigpath':
'results/edges', 'matpath': 'cache_dir', 'edgematpath':
'cache_dir/edges', 'datapath': 'database/', 'ext': '.pdf', 'figsize':
14.0, 'formats': ['pdf', 'png', 'jpg'], 'dpi': 450, 'verbose': 0}
lg = LogGabor(pe=pe)
i_eccentricity = 15
N_eccentricity = 48
ecc = ecc_max * (1 / rho) ** (N_eccentricity - i_eccentricity)
i_theta = 5
N_theta = 6
theta_ref = i_theta * np.pi / N_theta
sf_0 = 0.5 * sf_0_r / ecc
sf_0 = np.min((sf_0, sf_0_max))
N = 200
dimension_filtre = int( N /2 * ecc)
#if dimension_filtre % 2 == 1:
# dimension_filtre += 1
#dimension_filtre = 100
# print("dimension_filtre", dimension_filtre)
lg.set_size((dimension_filtre, dimension_filtre))
i_phase = 1
phase = i_phase * np.pi / 2
x = dimension_filtre // 2
y = dimension_filtre // 2
params = {'sf_0': sf_0,
'B_sf': B_sf,
'theta': theta_ref,
'B_theta': B_theta}
filter = lg.normalize(lg.invert(lg.loggabor(x, y, **params) * np.exp(-1j * phase)))
# -
plt.imshow(filter)
plt.plot(ecc_max * (1 / rho) ** (N_eccentricity - np.arange(N_eccentricity)))
plt.plot(ecc_max * (1 / 1.21) ** (N_eccentricity - np.arange(N_eccentricity)))
plt.plot(ecc_max * (1 / 1.08) ** (N_eccentricity - np.arange(N_eccentricity)))
plt.plot(ecc_max * (1 / rho) ** (N_eccentricity - np.arange(10)))
| dev/2019-12-09_retina-tests.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Automating Visualization:
# # Python script to find All of the source and destination Ports
# +
import re
import string
import glob
import os
import pandas as pd
import numpy as np
from pathlib import Path
pd.set_option('display.max_colwidth', 1500)
pd.set_option('display.max_rows', 3000)
# -
#Data is a CSV document prepared from parsed EDI files, containing product commodity code, source and destination ports and vessel names.
df = pd.read_csv('vis.csv')
c = df['HS_Codes'].isnull().sum(axis=0)
r = len(df.index)
t = df['file_name'].nunique()
print("Number of files parsed: ",t)
print("Total Number of transactions processed: ",r)
print("Total Number of transactions without commodity codes: ",c)
ports = pd.unique(df[['source', 'destination']].values.ravel('K'))
print(ports)
list_ports=[]
for i in ports:
list_ports.append(str(i))
print(list_ports)
# # USING GEOPY TO AUTOMATICALLY FIND THE PORT COORDINATES ON THE MAP
import geopy
from geopy.geocoders import Nominatim
locator = Nominatim(user_agent="myGeocoder")
rows=[]
for i in list_ports:
location = locator.geocode(str(i))
print(i)
coords = location.latitude, location.longitude
rows.append({"ports":i,"coords":coords})
cols=({"ports","coords"})
df2 = pd.DataFrame(rows, columns = cols)
df2
export_csv = df2.to_csv (r'~path\ports&coords.csv', index = None, header=True)
# +
df = pd.read_csv('vis.csv')
dfirstmerge = df.merge(df2, how='inner', left_on='source', right_on='ports')
new_df = dfirstmerge.drop(['file_name','file_type','transaction_no','raw_text','ports'], axis=1)
new_df.columns = ['vessel_identificatio','source','destination','HS_Codes','source_coords']
dsecondmerge = new_df.merge(df2, how='inner', left_on='destination', right_on='ports')
final_df = dsecondmerge.drop(['ports'], axis=1)
final_df.columns = ['vessel_identification','source','destination','HS_Codes','source_coords','dest_coords']
#Created new dataframe to sensor the output from sensitive data
final_df_output = final_df.drop(columns=['vessel_identification'])
final_df_output
# -
export_csv = final_df.to_csv (r'~path\source&dest.csv', index = None, header=True)
# +
#converting certain values to string to prepare a javascript file of source and destination coordinates for visualization
#cleared output due to sensitivity of the data
final_df['vessel_identification'] = df['vessel_identification'].apply(lambda x: "'" + str(x) + "'")
final_df['HS_Codes'] = final_df['HS_Codes'].apply(lambda x: "'" + str(x) + "'")
# -
# # Writing all of the filtered data such as Vessel Identification details, Source and destination points for the Vessel, Commodity codes, Source & destination coordinates and the in-between transition points for animated path
# +
index_size = (len(final_df.index))
print(index_size)
file = open("Prepared data\prepared_data.js","w")
for x in range(index_size):
path = []
x1 = final_df['source_coords'][x][0]
y1 = final_df['source_coords'][x][1]
start = (y1,x1)
path.append(list(start))
x2 = final_df['dest_coords'][x][0]
y2 = final_df['dest_coords'][x][1]
while((x1-x2)>=2 or (y1-y2)>=2):
if (x1>x2):
x1 = (x1 - 0.9)
else:
x1 = x1 + 0.9
if (y1 > y2):
y1 = y1 - 0.2
else:
y1 = y1 + 0.2
T = (y1,x1)
path.append(list(T))
T2 = (y2,x2)
path.append(list(T2))
L = ["var orthodroma"+str(x)+"={'type':'FeatureCollection','features':[{'type':'Feature','properties':{'hs_code':"+str(final_df['HS_Codes'][x])+",'name':"+str(final_df['vessel_identification'][x])+"},'geometry':{'type':'LineString','coordinates':"+str(path)+"}}]};"]
file.write("\n")
file.writelines(L)
file.close()
# -
# We obtain 3 important files from this automated script to later help visualization using D3.JS :
# 1. ports&coords.csv
# 2. source&dest.csv and finally
# 3. prepared_data.js
| Notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/", "height": 367} executionInfo={"elapsed": 3084, "status": "error", "timestamp": 1621623969006, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "06448352823627466388"}, "user_tz": 240} id="GeHaCVpDSVwU" outputId="befbe507-52aa-42d1-d507-9822b38861ed"
import numpy as np
import matplotlib.pyplot as plt
from qutip import *
from AFP_all import sech, AFP_rf
from AHP_all import AHP_rf
from AM_pulses import AM_rf
from BIR4 import BIR4_rf
from CHIRP import chirp_rf
from multi_freq import multi_freq
# %matplotlib notebook
# -
# # OPTION 1: Directly generate the RF pulse to simulate
"""
Available shaped pulse functions:
Amplitude Modulated (AM_rf): 'sinc', 'gaussian', 'hermite', 'square'
Adiabatic Full/Half Passage (AFP_rf/AHP_rf): 'HSn', 'tanh/tan'
B1 Insensitive Rotation (BIR4): 'tanh/tan', 'sech/tanh'
CHIRP Linear frequency sweep: chirp_rf
"""
pulse_length = 5000 #us
shape_pts = 256
rf, fm, pm, time = AFP_rf(shape_pts=shape_pts, pulse_length=pulse_length, func='HSn', n=8, sweep_bw=8000)
# # OPTION 2: Load a previously saved pulse from one of the other pulse functions (except SLR)
# ### Don't use if using OPTION 1
# +
BASE_RF_FILE = "C:/Users/RudrakshaMajumdar/Documents/GitHub/rf-bloch-simulator/saved_rf_pulses/sinc3_2ms/rf_pulse_file.npz"
BASE_RF_PULSE = np.load(BASE_RF_FILE)
rf = BASE_RF_PULSE['arr_0']
pm = BASE_RF_PULSE['arr_1']
fm = BASE_RF_PULSE['arr_2']
time = BASE_RF_PULSE['arr_5']
# RF pulse parameters
pulse_length = 2000 #us
shape_pts = len(rf) # points in shaped pulse
# -
# # Plot the shape
# +
# Convert degrees to radians for complex plot
pm_rad = np.deg2rad(pm)
fig = plt.figure(figsize=[6,6])
plt_rfamp = fig.add_subplot(221)
plt_fm = fig.add_subplot(222)
plt_pm = fig.add_subplot(223)
plt_complex = fig.add_subplot(224)
plt_rfamp.plot(time, rf, 'b')
plt_rfamp.set_ylabel("RF amplitude")
plt_rfamp.set_xlabel("Time (us)")
plt_rfamp.grid()
plt_fm.plot(time,fm, 'b')
plt_fm.set_ylabel("Frequency (Hz)")
plt_fm.set_xlabel("Time (us)")
plt_fm.grid()
plt_pm.plot(time,pm, 'b')
plt_pm.set_ylabel("Phase (deg.)")
plt_pm.set_xlabel("Time (us)")
plt_pm.grid()
plt_complex.plot(time, rf * np.cos(pm_rad), 'b', label = "B1x")
plt_complex.plot(time, rf * np.sin(pm_rad), 'r', label = "B1y.")
plt_complex.set_ylabel("RF amplitude")
plt_complex.set_xlabel("Time (us)")
plt_complex.set_ylim(-1,1)
plt_complex.legend()
plt_complex.grid()
plt.tight_layout()
plt.show()
# -
# # Set up the simulation conditions
# +
# Gyromagnetic ratio in MHz/T
gamma = 42.57
# Declare initial magnetization [Mx,My,Mz]
init_mag = np.array([0,0,1])
# Set Simulation parameters
sim_points = 200
rf_min = 0
rf_max = 5000 # Hz
freq_offset=0
freq_min = -10000
freq_max = 10000 # All in Hz
# -
# # Simulate the pulse amplitude profile
# + colab={"base_uri": "https://localhost:8080/", "height": 295} executionInfo={"elapsed": 13268, "status": "ok", "timestamp": 1621612653473, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "06448352823627466388"}, "user_tz": 240} id="wClJmrPRSVwc" outputId="9afec1be-157e-4d39-acb8-1a3ef43f1f95"
def RF_AMP_SIM(
pulse_length=pulse_length,
sim_points = sim_points,
rf_min = rf_min,
rf_max = rf_max,
freq_offset=freq_offset,
init_mag = init_mag
):
"""
A function to simulate magnetization as a function
of RF amplitude. In the current version, the pulse
needs to be pre-loaded outside the function.
"""
rf_amp = rf
rf_dwelltime = 1e-6 * (pulse_length/rf.size) # convert to seconds
# Convert rf phase from degrees to radians
rf_phase = (np.pi/180) * pm
# Convert freq offset from Hz to radians/sec
rf_offset = 2 * np.pi * freq_offset
# Initial magnetizations
Mx_0 = init_mag[0]
My_0 = init_mag[1]
Mz_0 = init_mag[2]
# x-axis step size
rf_step = (rf_max - rf_min)/sim_points
# rf amplitude range for x-axis
rf_range = np.arange(rf_min,rf_max,rf_step)
# Scale Scale RF pulse over the entire RF range and convert from Hz to rad/s
rf_amp = 2 * np.pi * rf_range.reshape(-1,1) * rf_amp.reshape(1,-1) #can also use np.outer
# Memory allocation
Mx = np.zeros(sim_points)
My = np.zeros(sim_points)
Mz = np.zeros(sim_points)
Rx = np.identity(3)
Ry = np.identity(3)
Ry2 = np.identity(3)
Rz = np.identity(3)
Rz2 = np.identity(3)
M = np.zeros([3,1])
# Start Simulation
for rf_range_counter in range(sim_points):
M[0,0] = Mx_0
M[1,0] = My_0
M[2,0] = Mz_0
for rf_pulse_counter in range(rf.size):
term_0 = rf_amp[rf_range_counter, rf_pulse_counter] ** 2
term_1 = rf_offset ** 2
#B_effective
Be = np.sqrt(term_0 + term_1) * rf_dwelltime
alpha = np.arctan2(rf_offset, rf_amp[rf_range_counter,rf_pulse_counter])
# Precalculate various sin/cos terms for increased speed
cosBe = np.cos(Be)
sinBe = np.sin(Be)
cosalpha = np.cos(alpha)
sinalpha = np.sin(alpha)
cosphi = np.cos(rf_phase[rf_pulse_counter])
sinphi = np.sin(rf_phase[rf_pulse_counter])
# Construct the total rotation matrix
Rx[1,1] = cosBe
Rx[1,2] = sinBe
Rx[2,1] = -1.0 * sinBe
Rx[2,2] = cosBe
Ry[0,0] = cosalpha
Ry[0,2] = -1.0 * sinalpha
Ry[2,0] = sinalpha
Ry[2,2] = cosalpha
Ry2[0,0] = cosalpha
Ry2[0,2] = sinalpha
Ry2[2,0] = -1.0 * sinalpha
Ry2[2,2] = cosalpha
Rz[0,0] = cosphi
Rz[0,1] = sinphi
Rz[1,0] = -1.0 * sinphi
Rz[1,1] = cosphi
Rz2[0,0] = cosphi
Rz2[0,1] = -1.0 * sinphi
Rz2[1,0] = sinphi
Rz2[1,1] = cosphi
M = np.linalg.multi_dot([Rz, Ry, Rx, Ry2, Rz2, M])
Mx[rf_range_counter] = M[0,0]
My[rf_range_counter] = M[1,0]
Mz[rf_range_counter] = M[2,0]
Mxy = np.sqrt(Mx**2 + My**2)
return(Mx, My, Mz, Mxy, rf_range, rf_amp)
Mx, My, Mz, Mxy, rf_range, rf_amp = RF_AMP_SIM()
def hz_2_uT(value):
return value * 2 * np.pi / gamma
def uT_2_hz(value):
return value * gamma / 2 * np.pi
fig = plt.figure(figsize=[8,4])
plt_all = fig.add_subplot(121)
plt_Mxy = fig.add_subplot(122)
plt.title("Pulse length: " + str(pulse_length) + " us", loc='center')
plt_all.plot(rf_range,Mx, 'r', label = "Mx", alpha=0.3)
plt_all.plot(rf_range,My, 'b', label = "My", alpha=0.3)
plt_all.plot(rf_range,Mz, 'g', label = "Mz")
plt_all.legend()
plt_all.set_xlabel("RF Amplitude (Hz)")
plt_all.set_ylabel("Mx, My, Mz")
plt_all.grid()
second_axis_all = plt_all.secondary_xaxis('top', functions = (hz_2_uT, uT_2_hz))
second_axis_all.set_xlabel("RF Amplitude (uT)")
plt_Mxy.plot(rf_range, Mxy, 'b', label = "Mxy")
plt_Mxy.legend()
plt_Mxy.set_xlabel("RF Amplitude (Hz)")
plt_Mxy.set_ylabel("Mxy")
plt_Mxy.grid()
second_axis_Mxy = plt_Mxy.secondary_xaxis('top', functions = (hz_2_uT, uT_2_hz))
second_axis_Mxy.set_xlabel("RF Amplitude (uT)")
# + [markdown] colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 26010, "status": "ok", "timestamp": 1621623942744, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "06448352823627466388"}, "user_tz": 240} id="06Uv5-RZUTVU" outputId="9df293e8-896b-4e07-9462-8b9e24305524"
# # Simulate the pulse frequency profile
# ### Use an appropriate amplitude from the amplitude profile
# ### usually corresponding to a 90 (Mz = 0) or 180 (Mz=-1) pulse
# + colab={"base_uri": "https://localhost:8080/", "height": 310} executionInfo={"elapsed": 13632, "status": "ok", "timestamp": 1621612843139, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "06448352823627466388"}, "user_tz": 240} id="-wK0i29RSVwf" outputId="bb6a599e-0de1-4848-8d89-2a1d74345d14"
rf_amplitude = 1300 # Hz, use appropriate vales from the amplitude plot above
def RF_FREQ_SIM(
pulse_length = pulse_length,
sim_points = sim_points,
freq_min = freq_min,
freq_max = freq_max,
rf_amplitude = rf_amplitude,
freq_offset=freq_offset,
init_mag = init_mag
):
"""
A function to simulate magnetization as a function
of Frequency Offset. In the current version, the pulse
needs to be pre-loaded outside the function.
"""
rf_amp = rf
rf_dwelltime = 1e-6 * (pulse_length/rf.size) # convert to seconds
# Convert rf phase from degrees to radians
rf_phase = (np.pi/180) * pm
# Initial magnetizations
Mx_0 = init_mag[0]
My_0 = init_mag[1]
Mz_0 = init_mag[2]
# x-axis step size
freq_step = (freq_max - freq_min)/sim_points
# rf frequency offset range for x-axis
freq_range = np.arange(freq_min,freq_max,freq_step)
# Scale Scale RF pulse over the entire RF range and convert from Hz to rad/s
rf_amp = 2 * np.pi * rf_amplitude * rf_amp.reshape(-1,1)
# Memory allocation
Mx = np.zeros(sim_points)
My = np.zeros(sim_points)
Mz = np.zeros(sim_points)
Rx = np.identity(3)
Ry = np.identity(3)
Ry2 = np.identity(3)
Rz = np.identity(3)
Rz2 = np.identity(3)
M = np.zeros([3,1])
# Start Simulation
for freq_range_counter in range(sim_points):
M[0,0] = Mx_0
M[1,0] = My_0
M[2,0] = Mz_0
# Convert frequency offset from Hz to rad/s
rf_offset = 2 * np.pi * freq_range[freq_range_counter]
for rf_pulse_counter in range(rf.size):
term_0 = rf_amp[rf_pulse_counter] ** 2
term_1 = rf_offset ** 2
#B_effective
Be = np.sqrt(term_0 + term_1) * rf_dwelltime
alpha = np.arctan2(rf_offset, rf_amp[rf_pulse_counter])
# Precalculate various sin/cos terms for increased speed
cosBe = np.cos(Be)
sinBe = np.sin(Be)
cosalpha = np.cos(alpha)
sinalpha = np.sin(alpha)
cosphi = np.cos(rf_phase[rf_pulse_counter])
sinphi = np.sin(rf_phase[rf_pulse_counter])
# Construct the total rotation matrix
Rx[1,1] = cosBe
Rx[1,2] = sinBe
Rx[2,1] = -1.0 * sinBe
Rx[2,2] = cosBe
Ry[0,0] = cosalpha
Ry[0,2] = -1.0 * sinalpha
Ry[2,0] = sinalpha
Ry[2,2] = cosalpha
Ry2[0,0] = cosalpha
Ry2[0,2] = sinalpha
Ry2[2,0] = -1.0 * sinalpha
Ry2[2,2] = cosalpha
Rz[0,0] = cosphi
Rz[0,1] = sinphi
Rz[1,0] = -1.0 * sinphi
Rz[1,1] = cosphi
Rz2[0,0] = cosphi
Rz2[0,1] = -1.0 * sinphi
Rz2[1,0] = sinphi
Rz2[1,1] = cosphi
M = np.linalg.multi_dot([Rz, Ry, Rx, Ry2, Rz2, M])
Mx[freq_range_counter] = M[0,0]
My[freq_range_counter] = M[1,0]
Mz[freq_range_counter] = M[2,0]
Mxy = np.sqrt(Mx**2 + My**2)
return(Mx, My, Mz, Mxy, freq_range)
Mx, My, Mz, Mxy, freq_range = RF_FREQ_SIM()
fig = plt.figure(figsize=[8,4])
plt_all = fig.add_subplot(121)
plt_Mxy = fig.add_subplot(122)
plt.title(
"Pulse Amplitude: "
+ str(rf_amplitude)
+ " Hz"
+ "\nPulse length: "
+ str(pulse_length) +
" us"
)
plt_all.plot(freq_range,Mx, 'r', label = "Mx", alpha=0.3)
plt_all.plot(freq_range,My, 'b', label = "My", alpha=0.3)
plt_all.plot(freq_range,Mz, 'g', label = "Mz")
plt_all.set_ylim(-1,1)
plt_all.legend()
plt_all.set_xlabel("RF Frequency Offset (Hz)")
plt_all.set_ylabel("Mx, My, Mz")
plt_all.grid()
plt_Mxy.plot(freq_range, Mxy, 'b', label = "Mxy")
plt_Mxy.set_ylim(0,1)
plt_Mxy.legend()
plt_Mxy.set_xlabel("RF Frequency Offset (Hz)")
plt_Mxy.set_ylabel("Mxy")
plt_Mxy.grid()
# -
# # Simulate the magnetization trajectory over time
# + colab={"base_uri": "https://localhost:8080/", "height": 657} executionInfo={"elapsed": 1203, "status": "ok", "timestamp": 1621610671311, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "06448352823627466388"}, "user_tz": 240} id="ItWEMJWfSVwm" outputId="cc86e5b1-d430-40ba-9b2e-e9bd61d7c409"
def RF_TIME_SIM(
pulse_length = pulse_length,
rf_amplitude = rf_amplitude,
freq_offset = freq_offset,
init_mag = init_mag
):
"""
A function to simulate magnetization as a function
of time and on the Bloch sphere. In the current version, the pulse
needs to be pre-loaded outside the function.
"""
rf_amp = rf
rf_dwelltime = 1e-6 * (pulse_length/rf.size) # convert to seconds
time_axis = np.arange(0, pulse_length, pulse_length/rf.size)
# Convert rf phase from degrees to radians
rf_phase = (np.pi/180) * pm
# Initial magnetizations
Mx_0 = init_mag[0]
My_0 = init_mag[1]
Mz_0 = init_mag[2]
# Scale Scale RF pulse over the entire RF range and convert from Hz to rad/s
rf_amp = 2 * np.pi * rf_amplitude * rf_amp.reshape(-1,1)
# Memory allocation
Mx = np.zeros(rf.size)
My = np.zeros(rf.size)
Mz = np.zeros(rf.size)
Rx = np.identity(3)
Ry = np.identity(3)
Ry2 = np.identity(3)
Rz = np.identity(3)
Rz2 = np.identity(3)
M = np.zeros([3,1])
# Start Simulation
M[0,0] = Mx_0
M[1,0] = My_0
M[2,0] = Mz_0
# Convert frequency offset from Hz to rad/s
rf_offset = 2 * np.pi * freq_offset
for rf_pulse_counter in range(rf.size):
term_0 = rf_amp[rf_pulse_counter] ** 2
term_1 = rf_offset ** 2
#B_effective
Be = np.sqrt(term_0 + term_1) * rf_dwelltime
alpha = np.arctan2(rf_offset, rf_amp[rf_pulse_counter])
# Precalculate various sin/cos terms for increased speed
cosBe = np.cos(Be)
sinBe = np.sin(Be)
cosalpha = np.cos(alpha)
sinalpha = np.sin(alpha)
cosphi = np.cos(rf_phase[rf_pulse_counter])
sinphi = np.sin(rf_phase[rf_pulse_counter])
# Construct the total rotation matrix
Rx[1,1] = cosBe
Rx[1,2] = sinBe
Rx[2,1] = -1.0 * sinBe
Rx[2,2] = cosBe
Ry[0,0] = cosalpha
Ry[0,2] = -1.0 * sinalpha
Ry[2,0] = sinalpha
Ry[2,2] = cosalpha
Ry2[0,0] = cosalpha
Ry2[0,2] = sinalpha
Ry2[2,0] = -1.0 * sinalpha
Ry2[2,2] = cosalpha
Rz[0,0] = cosphi
Rz[0,1] = sinphi
Rz[1,0] = -1.0 * sinphi
Rz[1,1] = cosphi
Rz2[0,0] = cosphi
Rz2[0,1] = -1.0 * sinphi
Rz2[1,0] = sinphi
Rz2[1,1] = cosphi
M = np.linalg.multi_dot([Rz, Ry, Rx, Ry2, Rz2, M])
Mx[rf_pulse_counter] = M[0,0]
My[rf_pulse_counter] = M[1,0]
Mz[rf_pulse_counter] = M[2,0]
return(Mx, My, Mz, time_axis)
Mx, My, Mz, time_axis = RF_TIME_SIM()
# Plotting
fig = plt.figure(figsize=[8,4])
plt_Mx_y = fig.add_subplot(121)
plt_Mz = fig.add_subplot(122)
plt_Mx_y.plot(time_axis,Mx, 'r', label = "Mx")
plt_Mx_y.plot(time_axis,My, 'b', label = "My")
plt_Mx_y.set_ylim(-1,1)
plt_Mx_y.set_xlabel("Time (us)")
plt_Mx_y.set_ylabel("Mx, My")
plt_Mx_y.legend()
plt_Mx_y.grid()
plt_Mz.plot(time_axis,Mz, 'b', label = "Mz")
plt_Mz.set_ylim(-1,1)
plt_Mz.set_xlabel("Time (us)")
plt_Mz.set_ylabel("Mz")
plt_Mz.legend()
plt_Mz.grid()
# Bloch sphere
bloch = Bloch()
pnts = [Mx, My, Mz]
bloch.add_points(pnts)
bloch.point_size = [2, 2, 2, 2]
bloch.xlabel = ['$x$', '$-x$']
bloch.ylabel = ['$y$', '$-y$']
bloch.zlabel = ['$z$', '$-z$']
bloch.show()
# -
| .ipynb_checkpoints/Bloch_simulator-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import torch
import torch.nn as nn
import numpy as np
import queue
# ### Momentum Contrastive learning
#
# This notebook contains stand-alone implmentation of Momentum contrastive learning.
# Things to keep in mind, should not depend on the dataset or the network
#
# Still need to figure out the batch norm thing
class MOC(nn.Module):
def __init__(self, dict_size=10000, update_momentum=0.999):
super(MOC, self).__init__()
self.moc_queue = queue.Queue()
self.moc_momentum = update_momentum
self.loss =
def forward(self, x_q, x_k, k_encoder, q_encoder):
# the algorithm is basically copied from the paper here
q = q_encoder(x_q) ## N x C
k = k_encoder(x_k) ## N x C
k = k.detach() # no gradient to keys
# do the dot product for the positives, for me I want to
# minimize the distance of positives and maximize the
# distance to the negatives
l_pos = d
l_neg =
# logits
| pytorch_disco/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Vector quantisation of one grey scale image
# This tutorial will show an example of vector quantisation applied to a grey scale image (*cameraman.tif*) where pairs of pixels are vector quantised with 8 bits for each pair, roughly amounting to 4 bits per single pixel. The coding efficiency of such quantisation scheme will be compared with a uniform scalar quantisation scheme which quantises each pixel with the same amount of bits (i.e. 4). Given that both quantisers (i.e. scalar and vector) operate at the same bits per pixel, we'll measure the distortion in terms of Peak-Signal-to-Noise-Ratio (PSNR) and comment the objective and subjetive visual quality. The main goal of this training is to provide the reader with a practical example of vector quantisation, most notably how the generalised Lloyd-Max algorithm could be implemented. For a thorough treatment of the fundamentals of vector quantisation, the interested reader is referred to the following textbooks:
# * <NAME> and <NAME>. Vector Quantization and Signal Compression. Kluwer Academic Press, 732 pages, 1992.
# * <NAME> and <NAME>, "JPEG2000: Image compression fundamentals, standards and practice", Kluwer Academic Press, 773 pages, 2002.
#
# For vector quantisation, our pairs are constituted by the pixels belonging to two consecutive rows. This is shown in the following figure.
#
# <img src="vectors.png" width="400">
# + [markdown] slideshow={"slide_type": "slide"}
# The overall processing can be summarised with the following three main steps:
# * **Step 1**: Select a subset of vectors which will constitute the so-called Training Set (*TS*) and use it to design the reproduction levels for all vectors to be quantised (the so-called codebook).
# * **Step 2**: Derive the reproduction levels $l_i$ using the generalised Lloyd-Max algorithm over the TS found earlier.
# * **Step 3**: Perform the actual vector quantisation.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Step 1: Selection of the training set
# This step receives as input a gray scale image and returns the aforementioned Training Set (*TS*), constituted by a subset of the pairs of pixels associated with the whole image (i.e. our vectors). More precisely, we'll subsample all pairs of adjacent image pixels by a factor of 4 and insert such pairs in *TS*. Note that the subsampling factor is arbitrary but its value leads to a trade-off between coding efficiency and complexity. In fact, large subsampling factors will speed up the design of the reproduction levels (i.e. **Step 2**) but will result in lower coding efficiency as the levels have been designed on a set of pixel pairs which may not be representative of the image statistics. Conversely, a smaller subsampling factor, will increase coding efficiency given that now more pixels are included in the design of the codebook. The price to pay for this is an increase in the encoder's complexity. The following image depicts the selection of vectors to be included in the training set, while the code cell below implements such selection.
#
# <img src="training-set.png" width="650">
# + slideshow={"slide_type": "slide"}
import cv2
import numpy as np
import random as rnd
import matplotlib.pyplot as plt
# Vector quantiser bits
vq_bits = 8
vq_levels = 2**vq_bits
image = cv2.imread('../../input-data/cameraman.tif', cv2.IMREAD_UNCHANGED)
rows, cols = image.shape[0:2]
sampling_ratio, vector_height, vector_width = (4, 2, 1)
total_training_samples = (rows * cols) // (vector_height * sampling_ratio * vector_width * sampling_ratio)
training_set = np.zeros((vector_height * vector_width, total_training_samples), np.int32)
k = 0
for r in range(0, rows, vector_height * sampling_ratio):
for c in range(0, cols, vector_width*sampling_ratio):
training_vector = image[r:r + vector_height, c:c + vector_width]
training_set[:, k] = training_vector.flatten()
k += 1
# + [markdown] slideshow={"slide_type": "slide"}
# ## Step 2: Derivation of the reproduction levels
# Once the training set is available, the reproduction levels can be derived by applying the generalised Lloyd-Max algorithm (see the references listed above for more details). Accordingly, the initial reproduction levels will be set equal to some vectors belonging to the training set. It is important to note at this point that the choice of the initial value for the reproduction level will primarily impact on the convergence speed of the Lloyd-Max algorithm. So if we had more information about the image statistics (e.g. we know that the image has a bimodal histogram) we could reduce the number of iterations by properly selecting the values associated with the two peaks in the histogram. For this example we'll select the initial value by sampling the training set calculated by a factor $r = \frac{|TS|}{2^{qb}}$, where $qb$ in our example is equal to 8 bits per vector and $|\cdot|$ denotes the number of vectors included in the training set. Let's denote the set of initial reproduction levels as $L_{init}$, the Llyod-Max algorithm will take $L_{init}$ as input parameter along with the training set vectors. The output of the algorithm will be the set of reproduction levels $L_{final}$ containing all reproduction levels which minimise the overall Mean Square Error (MSE) between the vectors in *TS* and their vector quantised counterparts.
#
# We can summarise the generalised Lloyd-Max algorithm with the following sequence of ordered steps:
# 1. Set $L_{final} = L_{init}$.
# 1. For each vector $v_i$ in the training set, find the reproduction level $l_i \in L_{final}$ which minimises the square error $e^2 = (v_i - l_i)^2$.
# 1. Add the value of $e^2$ to variable $SE$ which stores the overall square error for the current iteration.
# 1. Update $L_{final}$ as $L_{final} = L_{final} / H$, where $H$ denotes a 1D array having each $i$-th element containing the number of times $l_i$ has been selected as the closest reproduction level for a given $v_i$ in the training set. If a given $l_i$ has never been selected, then substitute it by randomly choosing another vector from the training set.
# 1. If $SE$ hasn't decreased by a factor $\epsilon$ stop, else go to Step 2.
#
# The following Python code cell implements such iterative procedure.
# + slideshow={"slide_type": "slide"}
ts_sampling_ratio = total_training_samples // vq_levels
reproduction_levels = training_set[:, ts_sampling_ratio-1::ts_sampling_ratio].astype(np.float64)
last_iteration_mse = 1e6
epsilon = 1e-3
iteration = 0
delta_mse = 1.0
print("Step\tMSE\tvariation")
while delta_mse > epsilon:
levels_accumulator = np.zeros((vector_height * vector_width, vq_levels), np.float64)
levels_hit_cnt = np.zeros(vq_levels, np.int32)
MSE = 0.0
# Step 2: For each vector vi in the training set, find the reproduction level li which minimises
# the square error
for i in range(total_training_samples):
V = training_set[:, i]
dV = np.dot(V.T, V) + np.sum(np.square(reproduction_levels), axis=0) - 2*np.dot(V.T, reproduction_levels)
square_error = np.min(dV)
l_start_idx = np.argmin(dV)
levels_accumulator[:, l_start_idx] += V
levels_hit_cnt[l_start_idx] += 1
MSE += square_error
MSE /= total_training_samples * vector_height * vector_width
# Step 3: Update Lfinal as Lfinal = Lfinal / H
for i in range(vq_levels):
if levels_hit_cnt[i]:
reproduction_levels[:, i] = levels_accumulator[:, i] / levels_hit_cnt[i]
else:
random_idx = max(1, int(rnd.random()*total_training_samples))
reproduction_levels[:, i] = training_set[:, random_idx]
delta_mse = (last_iteration_mse - MSE) / MSE
print(f"{iteration}\t{MSE}\t{delta_mse}")
iteration += 1
last_iteration_mse = MSE
# + [markdown] slideshow={"slide_type": "skip"}
# Worth noting from the code cell above how the selection of $l_i$ from Step 2 of the generalised Lloyd-Max algorithm is implemented. In principle, finding $l_i$ which minimises the square error with the current vector $v_i$ can be done by looping through all reproduction levels, compute such square error and pick the one which minimises it. However, we can compact the code by noting that what we're doing is indeed the following:
#
# $$
# \large
# e^2 = \lvert\lvert L_{final} - v_i\rvert\rvert^2 = L_{final}\cdot L_{final}^t - 2*L_{final}^t\cdot v_i + v_i\cdot v_i^t,
# $$
#
# where superscript $^t$ denotes the transpose operator and $\lvert\lvert \cdot\rvert\rvert^2$ is the $L^2$ norm. Given that our data are stored in **numpy** arrays, dot products and element wise operations such as sum and subtraction are easily implemented and built in as either overloaded operators or interfaces.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Step 3: Actual vector quantisation over input image
# Now that the optimal reproduction levels have been found by the generalised Lloyd-Max algorithm, it is time to perform actual vector quantisation. The processing is similar to what we did above when deriving the optimal reproduction levels. In fact, this time we'll loop through all vectors associated with the *cameraman.tif* image and, for each one, $v_i$, the reproduction level $l_i$ from $L_{final}$ which minimises the square error $e^2 = (v_i - l_i)^2$, will be selected. Vector $l_i$ will be then placed at the same spatial location of $v_i$ and the process can move to the next $v_i$.
# + slideshow={"slide_type": "slide"}
reproduction_levels = np.round(reproduction_levels).astype(np.int32)
square_sum_level = np.sum(np.square(reproduction_levels), axis=0)
image_vq = np.zeros((rows, cols), np.uint8)
for r in range(0, rows, vector_height):
for c in range(0, cols, vector_width):
V = image[r:r + vector_height, c:c + vector_width].flatten()
dV = np.dot(V.T, V) + square_sum_level - 2*np.dot(V.T, reproduction_levels)
l_start_idx = np.argmin(dV)
image_vq[r:r + vector_height, c:c + vector_width] =\
np.reshape(reproduction_levels[:, l_start_idx], (vector_height, vector_width))
# + [markdown] slideshow={"slide_type": "slide"}
# ## Comparison with scalar quantisation
# The last operation we need to perform is to quantise *cameraman.tif* with a quantiser having $qb = 4$, that is 4 bits per pixel. Over the images obtained with scalar and vector quantisation, we'll then compute the Peak-Signal-to-Noise-Ratio (PSNR) and express it in decibel [dB] according to the following formula:
#
# $$
# PSNR(I,\hat{I}) = 10\cdot\log_{10}\left(\frac{M^2}{E\left[\lvert\lvert I - \hat{I}\rvert\rvert^2\right]}\right) [dB],
# $$
#
# where $\hat{I}$ denotes the image quantised with either scalar or vector quantisation, $M$ is the maximum value allowed for image $I$, that is 255 with an 8 bit per pixel image and finally $E[\cdot]$ denotes the expectation operator.
# + slideshow={"slide_type": "slide"}
sq_bits = vq_bits / vector_height / vector_width
Q = 256 // 2**sq_bits
image_sq = np.round(image / Q).astype(np.int32) * Q
mse_vq = np.mean(np.square(image - image_vq))
mse_sq = np.mean(np.square(image - image_sq))
psnr_vq = 10.0*np.log10(255.0**2 / mse_vq)
psnr_sq = 10.0*np.log10(255.0**2 / mse_sq)
plt.figure(1)
plt.figure(figsize=(20,20))
plt.subplot(1, 3, 1), plt.imshow(image, cmap='gray'), plt.title('Original image')
plt.subplot(1, 3, 2), plt.imshow(image_vq, cmap='gray'), plt.title(f"Vector quantised image (PSNR = {psnr_vq:.2f} [dB])")
plt.subplot(1, 3, 3), plt.imshow(image_sq, cmap='gray'), plt.title(f"Scalar quantised image (PSNR = {psnr_sq:.2f} [dB])");
# + [markdown] slideshow={"slide_type": "skip"}
# As we may note, the image resulting from scalar quantisation shows noticeable banding artefacts. Quality is significantly better for the vector quantised image which not only improves the PSNR by almost 4 dBs but also shows less artefacts.
#
# This example shows a compelling case for the use of vector quantisation: in fact, the vectors considered (i.e. pair of adjacent image pixels) show a correlation which would make their distribution on a scatter plot aligned to the 45 degree straight line. Such a correlation is efficiently exploited by vector quantisation whereby the generalised Lloyd-Max algorithm places the reproduction levels along the joint probability mass function. Scalar quantisation doesn't consider this pair-based correlation, hence places all reproduction levels as to span all possible range of values (even those which would never appear in the image statistics).
#
# # Concluding remarks
# We have presented a simple implementation of the generalised Lloyd-Max algorithm with application to image coding via vector quantisation. We have verified that vector quantisation is indeed a better alternative to scalar quantisation when the input data show some degree of correlation (or redundancy). Accordingly, if the transmitter (i.e. the encoder) can bear some additional complexity, vector quantisation can constitute an attractive alternative. Worth noting that we didn't considered to apply entropy coding on top of the resulting quantisation cells indexes: this would still reduce the coding rate given there will be some inter symbol redundancy to exploit with a coding scheme such as run-length encoding.
#
# It is also worth to mention that sometimes vector quantisation is referred as palette coding and a good example of design for the case of screen content and RGB images is the palette mode from the H.265/HEVC (V3) and H.266/VVC standards.
#
# We shall also provide the reader with some ideas on the extension of the vector quantisation scheme presented in this tutorial:
# * Consider colour images. Some design choices and aspects to address would be wether the input data are considered in the RGB or a YCbCr colour space. The former might save in complexity since no colour transform is required but would not allow for an effective perceptual quantisation. Another aspect is whether to treat each image plane separately or jointly. The latter might bring benefits in terms of coding efficiency.
# * Consider region based vector quantisation. Here the images is broken up into square regions and a different codebook is derived for each region. This will allow for parallel encoding and decoding, along with a more content adaptive coding scheme which, in this case, would get closer the palette mode of the H.265/HEVC and H.266/VVC standards.
#
# Finally, although we pointed out at the encoder's complexity as a limiting factor for vector quantisation, we should remind that in case of a region-based approach, GPU implementation of k-means algorithms (another way of optimising the codebook) will speed up compression. At the receiver side, the decoding process is a simple read from the bitstream and look up operation to write the pixels to the output buffer.
| quantisation/vector-quantisation/vq-image.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Import pre-processing libs
import numpy as np
import pandas as pd
from sklearn.preprocessing import OneHotEncoder, LabelEncoder
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
# Import model
from sklearn.linear_model import LogisticRegression
# Import post-processing libs
import matplotlib.pyplot as plt
from matplotlib.colors import ListedColormap
from sklearn.metrics import confusion_matrix
import pickle
# -
###################### 1- Import Data ######################
filename = ""
dataset = pd.read_csv(filename) # Check file extension before using this function
X = dataset.iloc[:,:-1].values
y = dataset.iloc[:, 1:].values
# +
###################### 2- Preprocessing ######################
# Split data
test_train_ratio = 0.2
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = test_train_ratio)
# -
###################### 3- Training ######################
model = LogisticRegression()
model.fit(X_train, y_train)
# +
###################### 4- Testing ######################
#model_score = model.score(X_test, y_test)
y_pred = model.predict(X_test)
cm = confusion_matrix(y_test, y_pred)
# +
###################### 5- Visualization ######################
# Visualising the Training set results
X_set, y_set = X_train, y_train
X1, X2 = np.meshgrid(np.arange(start = X_set[:, 0].min() - 1, stop = X_set[:, 0].max() + 1, step = 0.01),
np.arange(start = X_set[:, 1].min() - 1, stop = X_set[:, 1].max() + 1, step = 0.01))
plt.contourf(X1, X2, model.predict(np.array([X1.ravel(), X2.ravel()]).T).reshape(X1.shape),
alpha = 0.75, cmap = ListedColormap(('red', 'blue')))
plt.xlim(X1.min(), X1.max())
plt.ylim(X2.min(), X2.max())
for i, j in enumerate(np.unique(y_set)):
plt.scatter(X_set[y_set == j, 0], X_set[y_set == j, 1],
c = ListedColormap(('yellow', 'black'))(i), label = j)
plt.title('Model fitting (Training set)')
plt.xlabel('X')
plt.ylabel('y')
plt.legend()
plt.show()
# Visualising the Test set results
from matplotlib.colors import ListedColormap
X_set, y_set = X_test, y_test
X1, X2 = np.meshgrid(np.arange(start = X_set[:, 0].min() - 1, stop = X_set[:, 0].max() + 1, step = 0.01),
np.arange(start = X_set[:, 1].min() - 1, stop = X_set[:, 1].max() + 1, step = 0.01))
plt.contourf(X1, X2, model.predict(np.array([X1.ravel(), X2.ravel()]).T).reshape(X1.shape),
alpha = 0.75, cmap = ListedColormap(('red', 'blue')))
plt.xlim(X1.min(), X1.max())
plt.ylim(X2.min(), X2.max())
for i, j in enumerate(np.unique(y_set)):
plt.scatter(X_set[y_set == j, 0], X_set[y_set == j, 1],
c = ListedColormap(('yellow', 'black'))(i), label = j)
plt.title('Model fitting (Test set)')
plt.xlabel('X')
plt.ylabel('y')
plt.legend()
plt.show()
# +
###################### 6- Save & Use ######################
values_to_predict = X_test
prediction_result = model.predict([ values_to_predict ])
with open('classifier.pkl', 'wb') as f:
pickle.dump(model, f)
| Machine Learning/Basic Implementation/Classification/Logistic Regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/", "height": 131} colab_type="code" id="7I9ZQXimJCzc" outputId="c93f00bc-b91d-4e1e-c260-84d97dd127df"
# !git clone https://github.com/openai/gpt-2.git
# + colab={"base_uri": "https://localhost:8080/", "height": 510} colab_type="code" id="QO3K8Ep6JvUC" outputId="c3b5158e-e903-489a-a795-456e0809df4b"
# !pip install tensorflow-gpu
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="rpRD47hqK3od" outputId="92218f7d-2ee0-49d7-edc6-a03819cb3bd9"
# %cd gpt-2
# + colab={"base_uri": "https://localhost:8080/", "height": 558} colab_type="code" id="WmZm95VyJ1SJ" outputId="8683c96c-3ab3-46db-d082-5672b99b6ddb"
# !pip install -r requirements.txt
# + colab={"base_uri": "https://localhost:8080/", "height": 292} colab_type="code" id="6O4Q2d_oLHwm" outputId="162560b2-4341-4f29-d319-4c3a7cc1aac5"
# !python download_model.py 345M
# + colab={} colab_type="code" id="hnF91t1lJ9Nu"
# !sed -i 's/top_k=0/top_k=40/g' src/interactive_conditional_samples.py
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="3nVJ6VmYM4T7" outputId="5278213c-18e5-47c3-c08a-64857a8a5ee4"
# %cd src/
# + colab={} colab_type="code" id="cJcjlZKAKmVx"
def generate_text(seed_text="", length=None) -> str:
import os
import numpy as np
import tensorflow as tf
import json
import model, sample, encoder
nsamples = 1
models_dir = "../models"
batch_size = 1
temperature = 1
top_k = 30
seed = None
model_name='117M'
model_name='345M'
models_dir = os.path.expanduser(os.path.expandvars(models_dir))
if batch_size is None:
batch_size = 1
assert nsamples % batch_size == 0
enc = encoder.get_encoder(model_name, models_dir)
hparams = model.default_hparams()
with open(os.path.join(models_dir, model_name, 'hparams.json')) as f:
hparams.override_from_dict(json.load(f))
if length is None:
length = hparams.n_ctx // 2
elif length > hparams.n_ctx:
raise ValueError("Can't get samples longer than window size: %s" % hparams.n_ctx)
with tf.Session(graph=tf.Graph()) as sess:
context = tf.placeholder(tf.int32, [batch_size, None])
np.random.seed(seed)
tf.set_random_seed(seed)
output = sample.sample_sequence(
hparams=hparams, length=length,
context=context,
batch_size=batch_size,
temperature=temperature, top_k=top_k
)
saver = tf.train.Saver()
ckpt = tf.train.latest_checkpoint(os.path.join(models_dir, model_name))
saver.restore(sess, ckpt)
raw_text = seed_text
context_tokens = enc.encode(raw_text)
generated = 0
for _ in range(nsamples // batch_size):
out = sess.run(output, feed_dict={
context: [context_tokens for _ in range(batch_size)]
})[:, len(context_tokens):]
for i in range(batch_size):
generated += 1
text = enc.decode(out[i])
return text
# + colab={} colab_type="code" id="_TFtOdzPZHIz"
def get_future_prediction():
tag_seed = "children life death money career 2021 2022 2030 by the age of 50 "
input_text = tag_seed + "In the year of 2049 I want to"
time_markers = ["Next month I will", "In autumn it will be", "By the end of winter I will", "Next year I have to"]
time_index = 0
default_time_marker = "And then "
new_text = input_text
while len(new_text) - len(input_text) < 500:
new_text += generate_text(new_text, length=40)
period_position = new_text.rfind('.')
new_text = new_text[:period_position+1]
new_text += "\n"
if time_index < len(time_markers):
new_text += time_markers[time_index]
time_index += 1
else:
new_text += default_time_marker
# return new_text[len(tag_seed):]
return new_text
# + colab={"base_uri": "https://localhost:8080/", "height": 187} colab_type="code" id="9lxPAdZqhO-s" outputId="68b86263-949f-4d5a-cdd3-43fe2beb77b5"
new_text = get_future_prediction()
print(new_text)
# -
import telebot
token = '<KEY>'
bot = telebot.TeleBot(token)
| src/GPT_text_generation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Hydrology --- introduction
# > Water cycle, major processes
#
# - toc: true
# - badges: true
# - comments: false
# - categories: [jupyter]
# ## How much water is there? Where?
#
# > youtube: https://youtu.be/2ObMyytxLz8
#
# 
# 
# ## The water cycle
#
# 
# ## Global water distribution
#
# | Water source | Volume (km$^3$) | % of freshwater | % of total water |
# |:--------------------------------------|-----------------------------------|-----------------------|------------------------|
# | Oceans, Seas, & Bays | 1,338,000,000 | -- | 96.54 |
# | Ice caps, Glaciers,<br>& Permanent Snow | 24,064,000 | 68.7 | 1.74 |
# | Groundwater | 23,400,000 | -- | 1.69 |
# |$\quad$Fresh | 10,530,000 | 30.1 | 0.76 |
# |$\quad$Saline | 12,870,000 | -- | 0.93 |
# | Soil Moisture | 16,500 | 0.05 | 0.001 |
# | Ground Ice<br>& Permafrost | 300,000 | 0.86 | 0.022 |
# | Lakes | 176,400 | -- | 0.013 |
# |$\quad$Fresh | 91,000 | 0.26 | 0.007 |
# |$\quad$Saline | 85,400 | -- | 0.006 |
# | Atmosphere | 12,900 | 0.04 | 0.001 |
# | Swamp Water | 11,470 | 0.03 | 0.0008 |
# | Rivers | 2,120 | 0.006 | 0.0002 |
# | Biological Water | 1,120 | 0.003 | 0.0001 |
#
# \* (Percents are rounded, so will not add to 100)
# [https://www.usgs.gov/special-topic/water-science-school/science/fundamentals-water-cycle](https://www.usgs.gov/special-topic/water-science-school/science/fundamentals-water-cycle)
# ## Energy drives the hydrologic cycle
#
# > A key aspect of the hydrologic cycle is the fact that it is driven by energy inputs (primarily from the sun). At the global scale, the system is essentially closed with respect to water; negligible water is entering or leaving the system. In other words, there is no external forcing in terms of a water flux. Systems with no external forcing will generally eventually come to an equilibrium state. So what makes the hydrologic cycle so dynamic? The solar radiative energy input, which is external to the system, drives the hydrologic cycle. Averaged over the globe, 342 W m$^{-2}$ of solar radiative energy is being continuously input to the system at the top of the atmosphere. This energy input must be dissipated, and this is done, to a large extent, via the hydrologic cycle. Due to this fact, the study of hydrology is not isolated to the study of water storage and movement, but also must often include study of energy storage and movements.
#
# Margulis, 2017, "Introduction to Hydrology"
# ## Components of the water cycle
#
# ### Water storage in oceans
#
#
#
#
#
# ### Evaporation / Sublimation
#
# Evaporation $\longrightarrow$ cooling
#
# 
#
# 
#
# 
#
# 
# 
# 
# ### Evapotranspiration
#
# 
# ### Water storage in the atmosphere
#
# Cumulonimbus cloud over Africa
# 
#
# Picture of cumulonimbus taken from the International Space Station, over western Africa near the Senegal-Mali border.
#
# If all of the water in the atmosphere rained down at once, it would only cover the globe to a depth of 2.5 centimeters.
# $$
# \begin{align}
# \text{amount of water in the atmosphere} & \qquad V = 12\, 900\, \text{km}^3 \\
# \text{surface of Earth} & \qquad S = 4 \pi R^2;\quad R=6371\,\text{km}\\
# & \qquad V = S \times h \\
# \text{height} & \qquad h = \frac{V}{S} \simeq 2.5\,\text{cm}
# \end{align}
# $$
#
# Try to calculate this yourself, and click on the button below to check how to do it.
#collapse-hide
# amount of water in the atmosphere
V = 12900 # km^3
# Earth's radius
R = 6371 # km
# surface of Earth = 4 pi Rˆ2
S = 4 * 3.141592 * R**2
# Volume: V = S * h, therefore
# height
h = V / S # in km
h_cm = h * 1e5 # in cm
print(f"The height would be ~ {h_cm:.1f} cm")
#
#
# ### Condensation
#
# ### Precipitation
#
# 
#
# | | Intensity (cm/h) | Median diameter (mm) | Velocity of fall (m/s) | Drops s$^{-1}$ m$^{-2}$ |
# |:-------------------|--------------------------|------------------------------------|---------------------------------------|----------------------------------------|
# | Fog | 0.013 | 0.01 | 0.003 | 67,425,000 |
# | Mist | 0.005 | 0.1 | 0.21 | 27,000 |
# | Drizzle | 0.025 | 0.96 | 4.1 | 151 |
# | Light rain | 0.10 | 1.24 | 4.8 | 280 |
# | Moderate rain | 0.38 | 1.60 | 5.7 | 495 |
# | Heavy rain | 1.52 | 2.05 | 6.7 | 495 |
# | Excessive rain | 4.06 | 2.40 | 7.3 | 818 |
# | Cloudburst | 10.2 | 2.85 | 7.9 | 1,220 |
#
#
# Source: https://www.usgs.gov/special-topic/water-science-school/science/precipitation-and-water-cycle
# ### Water storage in ice and snow
#
# 
#
# 
# ### Snowmelt runoff to streams
# ### Surface runoff
#
# 
#
# 
# ### Streamflow
#
# The Mississippi river basin is very large
# 
#
# The Amazon river basin is **Huge**
# 
#
# 
# ### Lakes and rivers
#
# 
#
# Lake Malawi
# 
#
# 
# ### Infiltration
#
# 
# ### Groundwater storage
#
# 
#
# 
#
# 
#
# Center Pivot irrigation in Nebraska taps the Ogallala Aquifer.
# 
# ### Groundwater flow and discharge
#
# 
#
# 
#
# 
#
# ### Spring
#
# Ein Gedi
# 
#
# Thousand Springs, Idaho
# 
#
#
| _notebooks/2020-02-01-introduction-lecture.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Abstract
#
# In this notebook, I would try to implement the random-forest classification algorithm.
#
# The implementation is inspired by a post from [machinelearningmastery.com](https://machinelearningmastery.com/implement-random-forest-scratch-python/), and reconstructed with a few improvements:
#
# - Use the efficient libraries such as *Pandas* and *Numpy*, instead of basic *list* and *dictionary*.
#
#
# - Visualize the resulting decision tree, with the Pydot framework.
#
#
# - Calculate the **feature importance** of the decision tree, and also the random forest.
#
#
# #### Content
#
# - Dataset
#
#
# - Decision Tree
# - plot decision tree
#
#
# - Random Forest
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
from matplotlib.pylab import rcParams
rcParams['figure.figsize'] = 13,6
rcParams['figure.dpi'] = 100
rcParams['savefig.dpi'] = 100
# -
# ### Dataset
#
# Here we use [a dataset from UCI Machine Learning Repository](https://archive.ics.uci.edu/ml/datasets/Connectionist+Bench+(Sonar,+Mines+vs.+Rocks%29) for testing.
# !ls sonar*
# ! head sonar.all-data -n 3
# +
df_data = pd.read_csv('./sonar.all-data', header=None)
print('Num of rows:', len(df_data))
print('Num of columns:', len(df_data.columns))
# -
# ### Decision Tree
# In this section, we construct/train the decision tree. The main idea of the algorithm is to recursively split the training data into two subgroups. At each split, we evaluate a subset of features, and iterate all values on each feature, to find the **optimal** split that have the minimum **gini impunity**, *i.e.* the groups after spliting are more *uniform* in term of the target feature.
# +
def gini_index(df_groups, target_feature_index):
"""
df_groups: [pd.DataFrame]
"""
total_samples = sum([len(group) for group in df_groups])
gini_value = 0.0
for group in df_groups:
group_size = float(len(group))
if group_size == 0:
continue
class_count = group[target_feature_index].value_counts()
group_gini = sum([(count/group_size) **2 for count in class_count])
# weight the group gini value by its relative size
gini_value += (1 - group_gini) * (group_size / total_samples)
return gini_value
def tree_node_split(df, column_index, split_value):
"""
Split the dataframe based on the values within a specified column,
the ones (with rows) < 'split_value' on the left subnode,
and the one >= 'split_value' on the right subnode.
df: DataFrame
return: two dataframes
"""
left_node = []
right_node = []
for index in range(len(df)):
row = df.iloc[index]
if row[column_index] < split_value:
left_node.append(row)
else:
right_node.append(row)
split_groups = [left_node, right_node]
split_groups = [pd.concat(group, axis=1).T if len(group) else pd.DataFrame() for group in split_groups]
return split_groups
#return (pd.concat(left_node, axis=1).T, pd.concat(right_node, axis=1).T)
def get_best_split(df, n_split_features, target_feature_index):
""" subsample the features,
enumerate each possible value on each feature to find the best split.
"""
import random
import sys
feature_indice = list(set(range(len(df.columns))) - set([target_feature_index]))
sample_features = random.sample(feature_indice, n_split_features)
# the gini_index BEFORE splitting
gini_before_split = gini_index([df], target_feature_index)
# stop splitting when the data is uniform !
if gini_before_split == 0.0:
# put all the data in the left node
ret = {"split_groups": [df, []],
"gini_before_split": 0.0,
"gini_after_split": 0.0,
"left_node_size": len(df),
"right_node_size": 0}
return ret
b_feature_index = None
b_feature_value = None
b_gini_index = sys.maxsize
b_split_groups = None
# split over each feature, on each of the unique feature values
for feature_index in sample_features:
split_values = df[feature_index].unique()
for feature_value in split_values:
split_groups = tree_node_split(df, feature_index, feature_value)
gini_value = gini_index(split_groups, target_feature_index)
if gini_value < b_gini_index:
b_gini_index = gini_value
b_feature_index = feature_index
b_feature_value = feature_value
b_split_groups = split_groups
return {"split_feature": int(b_feature_index),
"split_value": b_feature_value,
"gini_before_split": gini_before_split,
"gini_after_split": b_gini_index,
"split_groups": b_split_groups,
"left_node_size": len(b_split_groups[0]),
'right_node_size': len(b_split_groups[1])}
def weighted_gini_diff(tree):
"""
calculate the weight_feature_importance for all involved splitting features
i.e. n_split_samples * (gini_before_split - gini_after_split)
"""
from collections import defaultdict
gini_diff_dict = defaultdict(list)
def DFS(node):
""" traverse the tree in the way of DFS, and accumulate gini diff """
if isinstance(node, dict):
if node['gini_before_split'] != 0.0:
gini_diff = node['gini_before_split'] - node['gini_after_split']
n_split_samples = node['left_node_size'] + node['right_node_size']
gini_diff_dict[node['split_feature']].append(n_split_samples * gini_diff)
DFS(node['left'])
DFS(node['right'])
# iterate the tree in DFS preorder way
DFS(tree)
return gini_diff_dict
def feature_importance(gini_diff_dict):
from collections import defaultdict
varimp = defaultdict(float)
for feature, value in gini_diff_dict.items():
varimp[feature] = sum(value) / len(value)
return varimp
def make_leaf_node(group, target_feature_index):
if len(group) == 0:
return None
value_counts = group[target_feature_index].value_counts()
# choose the majority class as the label of the node
leaf_label = value_counts.sort_values(ascending=False).index[0]
return leaf_label
def create_node(node, n_split_features,
target_feature_index, depth,
max_tree_depth = 20, min_node_size = 2):
"""
recursively split the node, with certain stopping conditions,
e.g. max_tree_depth, min_node_size etc.
"""
left_split, right_split = node['split_groups']
del node['split_groups']
# case 1). first, check if we reach a non-splitting node
no_split = False
if len(left_split) == 0:
node['left'] = None
node['right'] = make_leaf_node(right_split, target_feature_index)
no_split = True
if len(right_split) == 0:
node['right'] = None
node['left'] = make_leaf_node(left_split, target_feature_index)
no_split = True
# non-splitting node
if no_split:
return
# case 2). check if we reach the maximum depth, then stop splitting
if depth >= max_tree_depth:
node['left'] = make_leaf_node(left_split, target_feature_index)
node['right'] = make_leaf_node(right_split, target_feature_index)
return # early return
# case 3). otherwise, recursively split the left and right node
if len(left_split) <= min_node_size:
node['left'] = make_leaf_node(left_split, target_feature_index)
else:
node['left'] = get_best_split(left_split, n_split_features, target_feature_index)
create_node(node['left'], n_split_features,
target_feature_index, depth + 1,
max_tree_depth, min_node_size)
if len(right_split) <= min_node_size:
node['right'] = make_leaf_node(right_split, target_feature_index)
else:
node['right'] = get_best_split(right_split, n_split_features, target_feature_index)
create_node(node['right'], n_split_features,
target_feature_index, depth + 1,
max_tree_depth, min_node_size)
def build_decision_tree(df, n_split_features, target_feature_index,
max_tree_depth = 10, min_node_size = 2):
root_node = get_best_split(df, n_split_features, target_feature_index)
create_node(root_node, n_split_features, target_feature_index, 1,
max_tree_depth, min_node_size)
return root_node
# +
def predict_one(tree_node, row):
""" Apply the decision tree on a single record
"""
if row[tree_node['split_feature']] < tree_node['split_value']:
if isinstance(tree_node['left'], dict):
return predict(tree_node['left'], row)
else:
# return the label on the leaf node
return tree_node['left']
else:
# check the right sub-tree
if isinstance(tree_node['right'], dict):
return predict(tree_node['right'], row)
else:
return tree_node['right']
def predict_batch(tree, df_data):
predicted = [predict_one(tree, df_data.iloc[i]) \
for i in range(len(df_data))]
return predicted
def get_accuracy(actual, predicted):
correct_count = sum(actual == predicted)
return float(correct_count) * 100 / len(actual)
# -
predict_one(tree, df_data.iloc[10])
get_accuracy(df_data[60], predict_batch(tree, df_data))
# +
gini_diff_dict = weighted_gini_diff(tree)
gini_diff_dict
# -
feature_importance(gini_diff_dict)
# #### Plot Decision Tree
# We can visualize the decision tree, via the Pydot framework.
# +
import pydot
# A global variable to generate a unique ID for each node
NODE_COUNT = 0
def gen_node_id():
global NODE_COUNT
NODE_COUNT += 1
return NODE_COUNT
def subtree_to_node(subtree, graph):
"""
Convert the decision-tree to the representation of node in Pydot
"""
if 'split_feature' in subtree:
subroot = pydot.Node(gen_node_id(),
label = "gini_index {:.4f} - {:.4f} \n split_feature {} \n split_value {} \n left[{}], right[{}]".format(
subtree['gini_before_split'],
subtree['gini_after_split'],
subtree['split_feature'],
subtree['split_value'],
subtree['left_node_size'],
subtree['right_node_size']))
graph.add_node(subroot)
else:
subroot = pydot.Node(gen_node_id(),
label = "gini_index {:.4f} - {:.4f} \n left[{}], right[{}]".format(
subtree['gini_before_split'],
subtree['gini_after_split'],
subtree['left_node_size'],
subtree['right_node_size']))
graph.add_node(subroot)
if subtree['left']:
if isinstance(subtree['left'], dict):
left_node = subtree_to_node(subtree['left'], graph)
else:
# leaf node
left_node = pydot.Node(gen_node_id(),
label = "label {}".format(subtree['left']))
graph.add_node(left_node)
graph.add_edge(pydot.Edge(subroot, left_node))
if subtree['right']:
if isinstance(subtree['right'], dict):
right_node = subtree_to_node(subtree['right'], graph)
else:
# leaf node
right_node = pydot.Node(gen_node_id(),
label = "label {}".format(subtree['right']))
graph.add_node(right_node)
graph.add_edge(pydot.Edge(subroot, right_node))
return subroot
def tree_to_graph(tree):
""" generate a image from the pydot graph """
graph = pydot.Dot(graph_type='digraph')
root = subtree_to_node(tree, graph)
return graph
from IPython.display import Image, display
def show_graph(graph):
plt = Image(graph.create_png())
display(plt)
def plot_tree(tree):
graph = tree_to_graph(tree)
show_graph(graph)
# +
# %%time
tree = build_decision_tree(df_data,
n_split_features = 2,
target_feature_index = 60,
max_tree_depth = 4,
min_node_size = 3)
# -
graph = tree_to_graph(tree)
show_graph(graph)
# +
#tree
# -
# ### Random Forest
#
# Build a list of *decision tree* and do the **emsemble** on the results
# +
import random
def bagging_predict(trees, row):
""" Gather the results given by a list of trees,
take the majority one as the response
"""
predictions = pd.Series([predict_one(tree, row) for tree in trees])
value_counts = predictions.value_counts().sort_values(ascending=False)
return value_counts.index[0]
def subsample(df, ratio):
sample_num = int(len(df) * ratio)
sample_row_indice = random.sample(range(len(df)), sample_num)
return df.iloc[sample_row_indice]
class RandomForest:
def __init__(self, n_trees, n_split_features,
sample_ratio = 1.0, # by default, use all training data
max_tree_depth = 10, min_node_size = 2,
random_seed = 7):
self.n_trees = n_trees
self.n_split_features = n_split_features
self.sample_ratio = sample_ratio
# the split stopping conditions
self.max_tree_depth = max_tree_depth
self.min_node_size = min_node_size
# the list of decision trees to add
self.trees = []
random.seed(random_seed)
def train(self, train_data, target_feature_index):
for i in range(self.n_trees):
samples = subsample(train_data, self.sample_ratio)
tree = build_decision_tree(samples,
n_split_features = self.n_split_features,
target_feature_index = target_feature_index,
max_tree_depth = self.max_tree_depth,
min_node_size = self.min_node_size)
print('build tree #{}'.format(i))
self.trees.append(tree)
def feature_importance(self):
from collections import defaultdict
total_gini_diff = defaultdict(list)
# aggregate the weighted gini diff from each tree
for tree in self.trees:
gini_diff = weighted_gini_diff(tree)
for feature, value in gini_diff.items():
total_gini_diff[feature].extend(value)
return feature_importance(total_gini_diff)
def predict(self, test_data):
predictions = [bagging_predict(self.trees, test_data.iloc[i]) \
for i in range(len(test_data))]
return predictions
# -
rf_model = RandomForest(n_trees=1, n_split_features=3,
max_tree_depth=4, min_node_size=2)
# +
# %%time
target_feature_index=60
rf_model.train(df_data, target_feature_index=target_feature_index)
# -
plot_tree(rf_model.trees[0])
rf_model.feature_importance()
# +
target_feature_index = 60
trees_predictions = rf_model.predict(df_data)
accuracy = get_accuracy(df_data[target_feature_index], trees_predictions)
print('accuracy of ensemble trees: {}'.format(accuracy))
# +
print('accuracy of each decision-tree, without ensemble')
for i in range(len(rf_model.trees)):
single_tree_prediction = predict_batch(rf_model.trees[i], df_data)
accuracy = get_accuracy(df_data[target_feature_index], single_tree_prediction)
print('tree #{} accuracy:{}'.format(i, accuracy))
# -
# ### Misc
ret = get_best_split(df_data, 3, target_feature_index=60)
# +
median = df_data[0].median()
per_75 = 0.0355
per_25 = 0.0135
group_split = tree_node_split(df_data, column_index=0, split_value=per_25)
left_node, right_node = group_split
print('split with median: {}, length: {}'.format(median, len(left_node)))
# -
left_node.head()
gini_index(group_split, target_feature_index=60)
| random-forest/random-forest-implementation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="ur8xi4C7S06n"
# Copyright 2021 Google LLC
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="JAPoU8Sm5E6e"
# <table align="left">
#
# <td>
# <a href="https://colab.research.google.com/github/GoogleCloudPlatform/vertex-ai-samples/blob/master/notebooks/official/matching_engine/intro-swivel.ipynb"">
# <img src="https://cloud.google.com/ml-engine/images/colab-logo-32px.png" alt="Colab logo"> Run in Colab
# </a>
# </td>
# <td>
# <a href="https://github.com/GoogleCloudPlatform/vertex-ai-samples/blob/master/notebooks/official/matching_engine/intro-swivel.ipynb">
# <img src="https://cloud.google.com/ml-engine/images/github-logo-32px.png" alt="GitHub logo">
# View on GitHub
# </a>
# </td>
# </table>
# + [markdown] id="tvgnzT1CKxrO"
# ## Overview
#
# This notebook demonstrate how to train an embedding with Submatrix-wise Vector Embedding Learner ([Swivel](https://arxiv.org/abs/1602.02215)) using Vertex Pipelines. The purpose of the embedding learner is to compute cooccurrences between tokens in a given dataset and to use the cooccurrences to generate embeddings.
#
# Vertex AI provides a pipeline template
# for training with Swivel, so you don't need to design your own pipeline or write
# your own training code.
#
# It will require you provide a bucket where the dataset will be stored.
#
# Note: you may incur charges for training, storage or usage of other GCP products (Dataflow) in connection with testing this SDK.
#
# ### Dataset
#
# You will use the following sample datasets in the public bucket **gs://cloud-samples-data/vertex-ai/matching-engine/swivel**:
#
# 1. **movielens_25m**: A [movie rating dataset](https://grouplens.org/datasets/movielens/25m/) for the items input type that you can use to create embeddings for movies. This dataset is processed so that each line contains the movies that have same rating by the same user. The directory also includes `movies.csv`, which maps the movie ids to their names.
# 2. **wikipedia**: A text corpus dataset created from a [Wikipedia dump](https://dumps.wikimedia.org/enwiki/) that you can use to create word embeddings.
#
# ### Objective
#
# In this notebook, you will learn how to train custom embeddings using Vertex Pipelines and deploy the model for serving. The steps performed include:
#
# 1. **Setup**: Importing the required libraries and setting your global variables.
# 2. **Configure parameters**: Setting the appropriate parameter values for the pipeline job.
# 3. **Train on Vertex Pipelines**: Create a Swivel job to Vertex Pipelines using pipeline template.
# 4. **Deploy on Vertex Prediction**: Importing and deploying the trained model to a callable endpoint.
# 5. **Predict**: Calling the deployed endpoint using online prediction.
# 6. **Cleaning up**: Deleting resources created by this tutorial.
#
# ### Costs
#
# This tutorial uses billable components of Google Cloud:
#
# * Vertex AI
# * Dataflow
# * Cloud Storage
#
# Learn about [Vertex AI
# pricing](https://cloud.google.com/vertex-ai/pricing), [Cloud Storage
# pricing](https://cloud.google.com/storage/pricing), and [Dataflow pricing](https://cloud.google.com/dataflow/pricing), and use the [Pricing
# Calculator](https://cloud.google.com/products/calculator/)
# to generate a cost estimate based on your projected usage.
# + [markdown] id="ze4-nDLfK4pw"
# ### Set up your local development environment
#
# **If you are using Colab or Google Cloud Notebooks**, your environment already meets
# all the requirements to run this notebook. You can skip this step.
# + [markdown] id="gCuSR8GkAgzl"
# **Otherwise**, make sure your environment meets this notebook's requirements.
# You need the following:
#
# * The Google Cloud SDK
# * Git
# * Python 3
# * virtualenv
# * Jupyter notebook running in a virtual environment with Python 3
#
# The Google Cloud guide to [Setting up a Python development
# environment](https://cloud.google.com/python/setup) and the [Jupyter
# installation guide](https://jupyter.org/install) provide detailed instructions
# for meeting these requirements. The following steps provide a condensed set of
# instructions:
#
# 1. [Install and initialize the Cloud SDK.](https://cloud.google.com/sdk/docs/)
#
# 1. [Install Python 3.](https://cloud.google.com/python/setup#installing_python)
#
# 1. [Install
# virtualenv](https://cloud.google.com/python/setup#installing_and_using_virtualenv)
# and create a virtual environment that uses Python 3. Activate the virtual environment.
#
# 1. To install Jupyter, run `pip3 install jupyter` on the
# command-line in a terminal shell.
#
# 1. To launch Jupyter, run `jupyter notebook` on the command-line in a terminal shell.
#
# 1. Open this notebook in the Jupyter Notebook Dashboard.
# + [markdown] id="i7EUnXsZhAGF"
# ### Install additional packages
#
# Install additional package dependencies not installed in your notebook environment, such as google-cloud-aiplatform, tensorboard-plugin-profile. Use the latest major GA version of each package.
# + id="2b4ef9b72d43"
import os
# The Google Cloud Notebook product has specific requirements
IS_GOOGLE_CLOUD_NOTEBOOK = os.path.exists("/opt/deeplearning/metadata/env_version")
# Google Cloud Notebook requires dependencies to be installed with '--user'
USER_FLAG = ""
if IS_GOOGLE_CLOUD_NOTEBOOK:
USER_FLAG = "--user"
# + id="wyy5Lbnzg5fi"
# !pip3 install {USER_FLAG} --upgrade pip
# !pip3 install {USER_FLAG} --upgrade scikit-learn
# !pip3 install {USER_FLAG} --upgrade google-cloud-aiplatform tensorboard-plugin-profile
# !pip3 install {USER_FLAG} --upgrade tensorflow
# + [markdown] id="hhq5zEbGg0XX"
# ### Restart the kernel
#
# After you install the additional packages, you need to restart the notebook kernel so it can find the packages.
# + id="EzrelQZ22IZj"
# Automatically restart kernel after installs
import os
if not os.getenv("IS_TESTING"):
# Automatically restart kernel after installs
import IPython
app = IPython.Application.instance()
app.kernel.do_shutdown(True)
# + [markdown] id="lWEdiXsJg0XY"
# ## Before you begin
# + [markdown] id="BF1j6f9HApxa"
# ### Set up your Google Cloud project
#
# **The following steps are required, regardless of your notebook environment.**
#
# 1. [Select or create a Google Cloud project](https://console.cloud.google.com/cloud-resource-manager). When you first create an account, you get a $300 free credit towards your compute/storage costs.
#
# 1. [Make sure that billing is enabled for your project](https://cloud.google.com/billing/docs/how-to/modify-project).
#
# 1. [Enable the Vertex AI API and Dataflow API](https://console.cloud.google.com/flows/enableapi?apiid=aiplatform.googleapis.com,dataflow.googleapis.com).
#
# 1. If you are running this notebook locally, you will need to install the [Cloud SDK](https://cloud.google.com/sdk).
#
# 1. Enter your project ID in the cell below. Then run the cell to make sure the
# Cloud SDK uses the right project for all the commands in this notebook.
#
# **Note**: Jupyter runs lines prefixed with `!` as shell commands, and it interpolates Python variables prefixed with `$` into these commands.
# + [markdown] id="WReHDGG5g0XY"
# #### Set your project ID
#
# **If you don't know your project ID**, you may be able to get your project ID using `gcloud`.
# + id="oM1iC_MfAts1"
import os
PROJECT_ID = ""
# Get your Google Cloud project ID and project number from gcloud
if not os.getenv("IS_TESTING"):
# shell_output = !gcloud config list --format 'value(core.project)' 2>/dev/null
PROJECT_ID = shell_output[0]
print("Project ID: ", PROJECT_ID)
# shell_output = !gcloud projects list --filter="$(gcloud config get-value project)" --format="value(PROJECT_NUMBER)" 2>/dev/null
PROJECT_NUMBER = shell_output[0]
print("Project number: ", PROJECT_NUMBER)
# + [markdown] id="qJYoRfYng0XZ"
# Otherwise, set your project ID here.
# + id="riG_qUokg0XZ"
if PROJECT_ID == "" or PROJECT_ID is None:
PROJECT_ID = "[your-project-id]" # @param {type:"string"}
# + [markdown] id="0CweX_c7eVSH"
# #### Timestamp
#
# If you are in a live tutorial session, you might be using a shared test account or project. To avoid name collisions between users on resources created, you create a timestamp for each instance session, and append it onto the name of resources you create in this tutorial.
# + id="aK4XnlYSeVSI"
from datetime import datetime
TIMESTAMP = datetime.now().strftime("%Y%m%d%H%M%S")
# + [markdown] id="dr--iN2kAylZ"
# ### Authenticate your Google Cloud account
#
# **If you are using Google Cloud Notebooks**, your environment is already
# authenticated. Skip this step.
# + [markdown] id="sBCra4QMA2wR"
# **If you are using Colab**, run the cell below and follow the instructions
# when prompted to authenticate your account via oAuth.
#
# **Otherwise**, follow these steps:
#
# 1. In the Cloud Console, go to the [**Create service account key**
# page](https://console.cloud.google.com/apis/credentials/serviceaccountkey).
#
# 2. Click **Create service account**.
#
# 3. In the **Service account name** field, enter a name, and
# click **Create**.
#
# 4. In the **Grant this service account access to project** section, click the **Role** drop-down list. Type "Vertex AI"
# into the filter box, and select
# **Vertex AI Administrator**. Type "Storage Object Admin" into the filter box, and select **Storage Object Admin**.
#
# 5. Click *Create*. A JSON file that contains your key downloads to your
# local environment.
#
# 6. Enter the path to your service account key as the
# `GOOGLE_APPLICATION_CREDENTIALS` variable in the cell below and run the cell.
# + id="PyQmSRbKA8r-"
import os
import sys
# If you are running this notebook in Colab, run this cell and follow the
# instructions to authenticate your GCP account. This provides access to your
# Cloud Storage bucket and lets you submit training jobs and prediction
# requests.
# The Google Cloud Notebook product has specific requirements
IS_GOOGLE_CLOUD_NOTEBOOK = os.path.exists("/opt/deeplearning/metadata/env_version")
# If on Google Cloud Notebooks, then don't execute this code
if not IS_GOOGLE_CLOUD_NOTEBOOK:
if "google.colab" in sys.modules:
from google.colab import auth as google_auth
google_auth.authenticate_user()
# If you are running this notebook locally, replace the string below with the
# path to your service account key and run this cell to authenticate your GCP
# account.
elif not os.getenv("IS_TESTING"):
# %env GOOGLE_APPLICATION_CREDENTIALS ''
# + [markdown] id="zgPO1eR3CYjk"
# ### Create a Cloud Storage bucket
#
# **The following steps are required, regardless of your notebook environment.**
#
# When you submit a built-in Swivel job using the Cloud SDK, you need a Cloud Storage bucket for storing the input dataset and pipeline artifacts (the trained model).
#
# Set the name of your Cloud Storage bucket below. It must be unique across all Cloud Storage buckets.
#
# You may also change the `REGION` variable, which is used for operations
# throughout the rest of this notebook. Make sure to [choose a region where Vertex AI services are
# available](https://cloud.google.com/vertex-ai/docs/general/locations#available_regions). You may
# not use a Multi-Regional Storage bucket for training with Vertex AI.
# + id="MzGDU7TWdts_"
BUCKET_NAME = "gs://[your-bucket-name]" # @param {type:"string"}
REGION = "us-central1" # @param {type:"string"}
# + id="cf221059d072"
if BUCKET_NAME == "" or BUCKET_NAME is None or BUCKET_NAME == "gs://[your-bucket-name]":
BUCKET_NAME = "gs://" + PROJECT_ID + "aip-" + TIMESTAMP
# + [markdown] id="-EcIXiGsCePi"
# **Only if your bucket doesn't already exist**: Run the following cell to create your Cloud Storage bucket.
# + id="NIq7R4HZCfIc"
# ! gsutil mb -l $REGION $BUCKET_NAME
# + [markdown] id="ucvCsknMCims"
# Finally, validate access to your Cloud Storage bucket by examining its contents:
# + id="vhOb7YnwClBb"
# ! gsutil ls -al $BUCKET_NAME
# + [markdown] id="d612cd762261"
# ### Service Account
#
# **If you don't know your service account**, try to get your service account using gcloud command by executing the second cell below.
# + id="801acfa0ffbc"
SERVICE_ACCOUNT = "[your-service-account]" # @param {type:"string"}
# + id="e7864c293f9e"
if (
SERVICE_ACCOUNT == ""
or SERVICE_ACCOUNT is None
or SERVICE_ACCOUNT == "[your-service-account]"
):
# Get your GCP project id from gcloud
# shell_output = !gcloud auth list 2>/dev/null
SERVICE_ACCOUNT = shell_output[2].split()[1]
print("Service Account:", SERVICE_ACCOUNT)
# + [markdown] id="1390d2890e4e"
# #### Set service account access for Vertex AI Pipelines
#
# Run the following commands to grant your service account access to read and write pipeline artifacts in the bucket that you created in the previous step -- you only need to run these once per service account.
# + id="0f5ef291f226"
# !gsutil iam ch serviceAccount:{SERVICE_ACCOUNT}:roles/storage.objectCreator $BUCKET_NAME
# !gsutil iam ch serviceAccount:{SERVICE_ACCOUNT}:roles/storage.objectViewer $BUCKET_NAME
# + [markdown] id="I3pGuwT7eVSJ"
# ### Import libraries and define constants
# Define constants used in this tutorial.
# + id="pARPYnT2eVSJ"
SOURCE_DATA_PATH = "{}/swivel".format(BUCKET_NAME)
PIPELINE_ROOT = "{}/pipeline_root".format(BUCKET_NAME)
# + [markdown] id="Vg6BigD5eVSJ"
# Import packages used in this tutorial.
# + id="fiimME4YeVSJ"
import pandas as pd
import tensorflow as tf
from google.cloud import aiplatform
from sklearn.metrics.pairwise import cosine_similarity
# + [markdown] id="Y9Uo3tifg1kx"
# ## Copy and configure the Swivel template
#
# Download the Swivel template and configuration script.
# + id="pRUOFELefqf1"
# !gsutil cp gs://cloud-samples-data/vertex-ai/matching-engine/swivel/pipeline/* .
# + [markdown] id="3ElFBL6BeVSK"
# Change your pipeline configurations:
#
# * pipeline_suffix: Suffix of your pipeline name (lowercase and hyphen are allowed).
# * machine_type: e.g. n1-standard-16.
# * accelerator_count: Number of GPUs in each machine.
# * accelerator_type: e.g. NVIDIA_TESLA_P100, NVIDIA_TESLA_V100.
# * region: e.g. us-east1 (optional, default is us-central1)
# * network_name: e.g., my_network_name (optional, otherwise it uses "default" network).
# + [markdown] id="712fffac0757"
# ### VPC Network peering, subnetwork and private IP address configuration
#
# Executing the following cell will generate two files:
# 1. `swivel_pipeline_basic.json`: The basic template allows public IPs and default network for the Dataflow job, and doesn't require setting up VPC Network peering for Vertex AI and **you will use it in this notebook sample**.
# 1. `swivel_pipeline.json`: This template enables private IPs and subnet configuration for the Dataflow job, also requires setting up VPC Network peering for the Vertex custom training. This template includes the following args:
# * "--subnetwork=regions/%REGION%/subnetworks/%NETWORK_NAME%",
# * "--no_use_public_ips",
# * \"network\": \"projects/%PROJECT_NUMBER%/global/networks/%NETWORK_NAME%\"
#
# **WARNING** In order to specify private IPs and configure VPC network, you need to [set up VPC Network peering for Vertex AI](https://cloud.google.com/vertex-ai/docs/general/vpc-peering#overview) for your subnetwork (e.g. "default" network on "us-central1") before submitting the following job. This is required for using private IP addresses for DataFlow and Vertex AI.
# + id="190tiY-neVSK"
YOUR_PIPELINE_SUFFIX = "swivel-pipeline-movie" # @param {type:"string"}
MACHINE_TYPE = "n1-standard-16" # @param {type:"string"}
ACCELERATOR_COUNT = 2 # @param {type:"integer"}
ACCELERATOR_TYPE = "NVIDIA_TESLA_V100" # @param {type:"string"}
BUCKET = BUCKET_NAME[5:] # remove "gs://" for the following command.
# !chmod +x swivel_template_configuration*
# !./swivel_template_configuration_basic.sh -pipeline_suffix {YOUR_PIPELINE_SUFFIX} -project_number {PROJECT_NUMBER} -project_id {PROJECT_ID} -machine_type {MACHINE_TYPE} -accelerator_count {ACCELERATOR_COUNT} -accelerator_type {ACCELERATOR_TYPE} -pipeline_root {BUCKET}
# !./swivel_template_configuration.sh -pipeline_suffix {YOUR_PIPELINE_SUFFIX} -project_number {PROJECT_NUMBER} -project_id {PROJECT_ID} -machine_type {MACHINE_TYPE} -accelerator_count {ACCELERATOR_COUNT} -accelerator_type {ACCELERATOR_TYPE} -pipeline_root {BUCKET}
# + [markdown] id="3f12048abec2"
# Both `swivel_pipeline_basic.json` and `swivel_pipeline.json` are generated.
# + [markdown] id="fcM5q2wfeVSK"
# ## Create the Swivel job for MovieLens items embeddings
#
# You will submit the pipeline job by passing the compiled spec to the `create_run_from_job_spec()` method. Note that you are passing a `parameter_values` dict that specifies the pipeline input parameters to use.
# + [markdown] id="af31EtxxeVSK"
# The following table shows the runtime parameters required by the Swivel job:
#
# | Parameter |Data type | Description | Required |
# |----------------------------|----------|--------------------------------------------------------------------|------------------------|
# | `embedding_dim` | int | Dimensions of the embeddings to train. | No - Default is 100 |
# | `input_base` | string | Cloud Storage path where the input data is stored. | Yes |
# | `input_type` | string | Type of the input data. Can be either 'text' (for wikipedia sample) or 'items'(for movielens sample). | Yes |
# | `max_vocab_size` | int | Maximum vocabulary size to generate embeddings for. | No - Default is 409600 |
# |`num_epochs` | int | Number of epochs for training. | No - Default is 20 |
#
# In short, the **items** input type means that each line of your input data should be space-separated item ids. Each line is tokenized by splitting on whitespace. The **text** input type means that each line of your input data should be equivalent to a sentence. Each line is tokenized by lowercasing, and splitting on whitespace.
#
# + id="8cd9e3db9bff"
# Copy the MovieLens sample dataset
# ! gsutil cp -r gs://cloud-samples-data/vertex-ai/matching-engine/swivel/movielens_25m/train/* {SOURCE_DATA_PATH}/movielens_25m
# + id="i9SnVxxleVSK"
# MovieLens items embedding sample
PARAMETER_VALUES = {
"embedding_dim": 100, # <---CHANGE THIS (OPTIONAL)
"input_base": "{}/movielens_25m/train".format(SOURCE_DATA_PATH),
"input_type": "items", # For movielens sample
"max_vocab_size": 409600, # <---CHANGE THIS (OPTIONAL)
"num_epochs": 5, # <---CHANGE THIS (OPTIONAL)
}
# + [markdown] id="9f1cae770338"
# Submit the pipeline to Vertex AI:
# + id="kUV5aYtPeVSK"
# Instantiate PipelineJob object
pl = aiplatform.PipelineJob(
display_name=YOUR_PIPELINE_SUFFIX,
# Whether or not to enable caching
# True = always cache pipeline step result
# False = never cache pipeline step result
# None = defer to cache option for each pipeline component in the pipeline definition
enable_caching=False,
# Local or GCS path to a compiled pipeline definition
template_path="swivel_pipeline_basic.json",
# Dictionary containing input parameters for your pipeline
parameter_values=PARAMETER_VALUES,
# GCS path to act as the pipeline root
pipeline_root=PIPELINE_ROOT,
)
# Submit the Pipeline to Vertex AI
# Optionally you may specify the service account below: submit(service_account=SERVICE_ACCOUNT)
# You must have iam.serviceAccounts.actAs permission on the service account to use it
pl.submit()
# + [markdown] id="xhznZuWceVSL"
# After the job is submitted successfully, you can view its details (including run name that you'll need below) and logs.
#
# ### Use TensorBoard to check the model
#
# You may use the TensorBoard to check the model training process. In order to do that, you need to find the path to the trained model artifact. After the job finishes successfully (~ a few hours), you can view the trained model output path in the [Vertex ML Metadata](https://cloud.google.com/vertex-ai/docs/ml-metadata/introduction) browser. It is going to have the following format:
#
# * {BUCKET_NAME}/pipeline_root/{PROJECT_NUMBER}/swivel-{TIMESTAMP}/EmbTrainerComponent_-{SOME_NUMBER}/model/
#
# You may copy this path for the MODELOUTPUT_DIR below.
#
# Alternatively, you can download a pretrained model to `{SOURCE_DATA_PATH}/movielens_model` and proceed. This pretrained model is for demo purpose and not optimized for production usage.
# + id="cc20afaf0e8a"
# ! gsutil -m cp -r gs://cloud-samples-data/vertex-ai/matching-engine/swivel/models/movielens/model {SOURCE_DATA_PATH}/movielens_model
# + id="c1Na-orVeVSL"
SAVEDMODEL_DIR = os.path.join(SOURCE_DATA_PATH, "movielens_model/model")
LOGS_DIR = os.path.join(SOURCE_DATA_PATH, "movielens_model/tensorboard")
# + [markdown] id="PY5ipT0feVSL"
# When the training starts, you can view the logs in TensorBoard:
# + id="zcswl8-OeVSL"
# If on Google Cloud Notebooks, then don't execute this code.
if not IS_GOOGLE_CLOUD_NOTEBOOK:
if "google.colab" in sys.modules:
# Load the TensorBoard notebook extension.
# %load_ext tensorboard
# + id="sjOzNEQseVSL"
# If on Google Cloud Notebooks, then don't execute this code.
if not IS_GOOGLE_CLOUD_NOTEBOOK:
if "google.colab" in sys.modules:
# %tensorboard --logdir $LOGS_DIR
# + [markdown] id="o3PbqO2IeVSL"
# For **Google Cloud Notebooks**, you can do the following:
#
# 1. Open Cloud Shell from the Google Cloud Console.
# 2. Install dependencies: `pip3 install tensorflow tensorboard-plugin-profile`
# 3. Run the following command: `tensorboard --logdir {LOGS_DIR}`. You will see a message "TensorBoard 2.x.0 at http://localhost:<PORT>/ (Press CTRL+C to quit)" as the output. Take note of the port number.
# 4. You can click on the Web Preview button and view the TensorBoard dashboard and profiling results. You need to configure Web Preview's port to be the same port as you receive from step 3.
# + [markdown] id="KE7pltkVeVSM"
# ## Deploy the embedding model for online serving
#
# To deploy the trained model, you will perform the following steps:
# * Create a model endpoint (if needed).
# * Upload the trained model to Model resource.
# * Deploy the Model to the endpoint.
# + id="9-rHi00XeVSM"
ENDPOINT_NAME = "swivel_embedding" # <---CHANGE THIS (OPTIONAL)
MODEL_VERSION_NAME = "movie-tf2-cpu-2.4" # <---CHANGE THIS (OPTIONAL)
# + id="udJ7mBk-eVSM"
aiplatform.init(project=PROJECT_ID, location=REGION)
# + id="P16dMukCeVSM"
# Create a model endpoint
endpoint = aiplatform.Endpoint.create(display_name=ENDPOINT_NAME)
# Upload the trained model to Model resource
model = aiplatform.Model.upload(
display_name=MODEL_VERSION_NAME,
artifact_uri=SAVEDMODEL_DIR,
serving_container_image_uri="us-docker.pkg.dev/vertex-ai/prediction/tf2-cpu.2-4:latest",
)
# Deploy the Model to the Endpoint
model.deploy(
endpoint=endpoint,
machine_type="n1-standard-2",
)
# + [markdown] id="9360794f914f"
# ### Load the movie ids and titles for querying embeddings
# + id="14fd3ab852a1"
# !gsutil cp gs://cloud-samples-data/vertex-ai/matching-engine/swivel/movielens_25m/movies.csv ./movies.csv
# + id="24af0e015685"
movies = pd.read_csv("movies.csv")
print(f"Movie count: {len(movies.index)}")
movies.head()
# + id="8aefb986c6c8"
# Change to your favourite movies.
query_movies = [
"Lion King, The (1994)",
"Aladdin (1992)",
"Star Wars: Episode IV - A New Hope (1977)",
"Star Wars: Episode VI - Return of the Jedi (1983)",
"Terminator 2: Judgment Day (1991)",
"Aliens (1986)",
"Godfather, The (1972)",
"Goodfellas (1990)",
]
# + id="f5aedea722d0"
def get_movie_id(title):
return list(movies[movies.title == title].movieId)[0]
input_items = [str(get_movie_id(title)) for title in query_movies]
# + [markdown] id="LDaCXinreVSM"
# ### Look up embedding by making an online prediction request
# + id="3JTz0i_ieVSM"
predictions = endpoint.predict(instances=input_items)
embeddings = predictions.predictions
print(len(embeddings))
# + [markdown] id="a1c5f572dec7"
# Explore movie embedding similarities:
# + id="8c81e656816a"
for idx1 in range(0, len(input_items) - 1, 2):
item1 = input_items[idx1]
title1 = query_movies[idx1]
print(title1)
print("==================")
embedding1 = embeddings[idx1]
for idx2 in range(0, len(input_items)):
item2 = input_items[idx2]
embedding2 = embeddings[idx2]
similarity = round(cosine_similarity([embedding1], [embedding2])[0][0], 5)
title1 = query_movies[idx1]
title2 = query_movies[idx2]
print(f" - Similarity to '{title2}' = {similarity}")
print()
# + [markdown] id="G8pgaNbveVSM"
# ## Create the Swivel job for Wikipedia text embedding (Optional)
#
# This section shows you how to create embeddings for the movies in the wikipedia dataset using Swivel. You need to do the following steps:
# 1. Configure the swivel template (using the **text** input_type) and create a pipeline job.
# 2. Run the following item embedding exploration code.
#
# The following cell overwrites `swivel_pipeline_template.json`; the new pipeline template file is almost identical, but it's labeled with your new pipeline suffix to distinguish it. This job will take **a few hours**.
# + id="02054a96f564"
# Copy the wikipedia sample dataset
# ! gsutil -m cp -r gs://cloud-samples-data/vertex-ai/matching-engine/swivel/wikipedia/* {SOURCE_DATA_PATH}/wikipedia
# + id="haz9gXnjeVSM"
YOUR_PIPELINE_SUFFIX = "my-first-pipeline-wiki" # @param {type:"string"}
# !./swivel_template_configuration.sh -pipeline_suffix {YOUR_PIPELINE_SUFFIX} -project_id {PROJECT_ID} -machine_type {MACHINE_TYPE} -accelerator_count {ACCELERATOR_COUNT} -accelerator_type {ACCELERATOR_TYPE} -pipeline_root {BUCKET}
# + id="2guNOxgjeVSN"
# wikipedia text embedding sample
PARAMETER_VALUES = {
"embedding_dim": 100, # <---CHANGE THIS (OPTIONAL)
"input_base": "{}/wikipedia".format(SOURCE_DATA_PATH),
"input_type": "text", # For wikipedia sample
"max_vocab_size": 409600, # <---CHANGE THIS (OPTIONAL)
"num_epochs": 20, # <---CHANGE THIS (OPTIONAL)
}
# + [markdown] id="wmcx4EADeVSN"
# **Submit the pipeline job through `aiplatform.PipelineJob` object.**
#
# After the job finishes successfully (~**a few hours**), you can view the trained model in your CLoud Storage browser. It is going to have the following format:
#
# * {BUCKET_NAME}/{PROJECT_NUMBER}/swivel-{TIMESTAMP}/EmbTrainerComponent_-{SOME_NUMBER}/model/
#
# You may copy this path for the MODELOUTPUT_DIR below. For demo purpose, you can download a pretrained model to `{SOURCE_DATA_PATH}/wikipedia_model` and proceed. This pretrained model is for demo purpose and not optimized for production usage.
# + id="ed94bd036502"
# ! gsutil -m cp -r gs://cloud-samples-data/vertex-ai/matching-engine/swivel/models/wikipedia/model {SOURCE_DATA_PATH}/wikipedia_model
# + id="eALPe9CAeVSN"
SAVEDMODEL_DIR = os.path.join(SOURCE_DATA_PATH, "wikipedia_model/model")
embedding_model = tf.saved_model.load(SAVEDMODEL_DIR)
# + [markdown] id="995NzGSAeVSN"
# ### Explore the trained text embeddings
#
# Load the SavedModel to lookup embeddings for items. Note the following:
# * The SavedModel expects a list of string inputs.
# * Each string input is treated as a list of space-separated tokens.
# * If the input is text, the string input is lowercased with punctuation removed.
# * An embedding is generated for each input by looking up the embedding of each token in the input and computing the average embedding per string input.
# * The embedding of an out-of-vocabulary (OOV) token is a vector of zeros.
#
# For example, if the input is ['horror', 'film', 'HORROR! Film'], the output will be three embedding vectors, where the third is the average of the first two.
# + id="1oyMaSvFeVSN"
input_items = ["horror", "film", '"HORROR! Film"', "horror-film"]
output_embeddings = embedding_model(input_items)
horror_film_embedding = tf.math.reduce_mean(output_embeddings[:2], axis=0)
# Average of embeddings for 'horror' and 'film' equals that for '"HORROR! Film"'
# since preprocessing cleans punctuation and lowercases.
assert tf.math.reduce_all(tf.equal(horror_film_embedding, output_embeddings[2])).numpy()
# Embedding for '"HORROR! Film"' equal that for 'horror-film' since the
# latter contains a hyphenation and thus is a separate token.
assert not tf.math.reduce_all(
tf.equal(output_embeddings[2], output_embeddings[3])
).numpy()
# + id="XMF_iHuFeVSN"
# Change input_items with your own item tokens
input_items = ["apple", "orange", "hammer", "nails"]
output_embeddings = embedding_model(input_items)
for idx1 in range(len(input_items)):
item1 = input_items[idx1]
embedding1 = output_embeddings[idx1].numpy()
for idx2 in range(idx1 + 1, len(input_items)):
item2 = input_items[idx2]
embedding2 = output_embeddings[idx2].numpy()
similarity = round(cosine_similarity([embedding1], [embedding2])[0][0], 5)
print(f"Similarity between '{item1}' and '{item2}' = {similarity}")
# + [markdown] id="71mBtTJ2eVSN"
# You can use the [TensorBoard Embedding Projector](https://www.tensorflow.org/tensorboard/tensorboard_projector_plugin) to graphically represent high dimensional embeddings, which can be helpful in examining and understanding your embeddings.
# + [markdown] id="TpV-iwP9qw9c"
# ## Cleaning up
#
# To clean up all Google Cloud resources used in this project, you can [delete the Google Cloud
# project](https://cloud.google.com/resource-manager/docs/creating-managing-projects#shutting_down_projects) you used for the tutorial.
#
# Otherwise, you can delete the individual resources you created in this tutorial:
#
# + id="5GBb7X1reVSN"
# Delete endpoint resource
# If force is set to True, all deployed models on this Endpoint will be undeployed first.
endpoint.delete(force=True)
# Delete model resource
MODEL_RESOURCE_NAME = model.resource_name
# ! gcloud ai models delete $MODEL_RESOURCE_NAME --region $REGION --quiet
# Delete Cloud Storage objects that were created
# ! gsutil -m rm -r $SOURCE_DATA_PATH
| notebooks/official/matching_engine/intro-swivel.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # MonaLIA Pair-Wise Odds Ratio between categories (DL training set)
#
# Experiment to explore whether a probability of appearance of one concept of a context-similar pair can improve the probability score of the second concept in the same image. For example, if a high classification probability score for a bateau could influence the score of the category mer. To achieve this, we used a logistic regression approach.
#
# The idea is to build a pairwise regression predictor of appearance of the category A of a pair (A, B) based on the presence of category B in the Joconde dataset metadata. Both dependent variable (category A) and predictor (category B) are binary labels. The regression estimates the log-odds of observing category A when category B is present compared to situations when category B is not present.
#
# $$log (odds(A)) = β_0+ β_1*B, where β_1=log (\frac{odds(A|B)}{odds(A no B)})$$
#
# These estimates are dependent on the dataset. If a different dataset is used it might lead to a different value of β1. Binary indicator model compares situations when category B is present or not. But because machine learning models predict categories with continuous probability scores S(A) and S(B), we want to reuse the regression parameters to predict an adjustment of probability score of category A of pair (A, B) based on a difference of probability score of category B compared to a baseline P(B)base that category B is present.
#
# $$log (odds(A)_{adj} ) = β_0+ β_1*(S(B) - P(B)_{base} ),$$
#
# where $S(B)$ is a classification prediction score of B. $P(B)_{base}$ can be calculated as a frequency of this category in the dataset. Thus, we consider how much the prediction score for B is higher (or lower) than the one obtained purely by chance. For the Joconde dataset we estimated P(B)base by direct counting of concepts in the annotations in the training set (to be noticed, that the training set is better balanced than the entire dataset).
#
# We assume that an unadjusted estimate of $S(A)$ corresponds to the baseline probability $P(B)_{base}$ and the adjustment could be made by using the actual prediction score $S(B)$. The adjusted odds of A thus become:
#
# $$odds(A)_{adj}= \frac{S(A)}{1-S(A)} * e^{β_1*(S(B)-P(B)_{base} ) }$$
#
# In case of $S(B) = P(B)_{base}$, there will be no adjustments to the score. This approach also considers that when the score of label B is lower than the average, it will reduce the probability of label A. This approach might impact recall and precision not symmetrically which can lead to subjective decisions.
#
# For example, the presence of a boat often implies some body of water, however the sea might be present on a painting without any boat. We thus considered an additional threshold for the lower values of $P(B)_{base}$, below which we do not consider adjustments. This modification will adjust the prediction score of category A when the label B is present and will leave $S(A)$ unadjusted when $P(B)_{base}$ is low.
#
# Method of adjusting the probability scores of category A in the context-bases pair (A, B) and evaluated the adjustments on the test set:
#
# 1. Estimate the odds ratio from a logistic regression on the metadata of the dataset
# 2. Estimate the baseline probability for category B on the same data
# 3. Calculate the adjusted prediction score for category A from the estimated odd ratio coefficient and the difference between the estimated baseline and prediction score for category B.
# 4. Evaluate the adjusted prediction using standard metrics
#
#
# #### This notebook reqires SPARQL engine with Joconde dataset and classification model parameter file
#
#
# Prediction scores obtained by running a pre-trained deep learning model
# +
from __future__ import print_function
import os
import sys
import numpy as np
import pandas as pd
import SPARQLWrapper
import matplotlib.pyplot as plt
import seaborn as sns
import statsmodels.api as sm
from sklearn import metrics
from IPython.display import display, HTML
from textwrap import wrap
import torch
import torch.nn as nn
import torchvision.datasets as dset
import torchvision.transforms as transforms
import torchvision.models
# +
# Import MonaLIA library from the package in the subfolder of the notebook folder
module_path = os.path.abspath(os.path.join('../..'))
if module_path not in sys.path:
sys.path.append(module_path)
import importlib
import MonaLIA.model.train as model
from MonaLIA.util import metadata_helpers as metadata
from MonaLIA.util import visualization_helpers as vis_helpers
from MonaLIA.data.image_dataset import JocondeDataset
importlib.reload(metadata)
importlib.reload(vis_helpers)
# -
print('SPARQLWrapper ver.', SPARQLWrapper.__version__)
print('Pandas ver.', pd.__version__)
wds_Joconde_Corese = 'http://localhost:8080/sparql'
# ## Query annotations
# +
descr_path = 'C:/Datasets/Joconde/Forty classes'
dataset_description_file = os.path.join(descr_path, 'dataset1.csv')
annotations_df = pd.read_csv(dataset_description_file, na_filter=False)
print(annotations_df.shape)
annotations_df.head()
# +
def my_tokenizer(s):
return list(filter(None, set(s.split('+'))))
annotations_df.label = annotations_df.label.apply(my_tokenizer)
# -
annotations_df.shape
# ## Create indicators columns
# +
train_annotations_df = annotations_df.loc[annotations_df.usage == 'train']
annotations_dummies_df = pd.get_dummies(train_annotations_df.explode('label').label).groupby(level=0).sum()
annotations_dummies_df.columns = annotations_dummies_df.columns.str.replace(' ', '_')
annotations_dummies_df.head()
# -
train_annotations_df = train_annotations_df.merge(annotations_dummies_df,
left_index=True,
right_index=True)
print(annotations_df.shape)
corr_df = annotations_dummies_df.corr()
# +
fig=plt.figure(figsize=(12, 12))
cmap_div = sns.diverging_palette(22, 130, s=99, n=16, as_cmap=True)
mask = corr_df >= 0.5
ax = sns.heatmap(corr_df , cmap=cmap_div,
mask = mask,
vmin=-0.5, vmax=0.5 , center =0.0,
square=True, cbar_kws={"shrink": 0.8})
ax.set_xticklabels(ax.get_xticklabels(), rotation=90)
ax.xaxis.set_ticks_position('top')
# +
fig=plt.figure(figsize=(12, 4))
plt.scatter(x = corr_df[corr_df < 0.5].rank(),
y = corr_df[corr_df < 0.5])
# +
top_corr_df = corr_df.stack().reset_index()
top_corr_df.columns = ['label_A', 'label_B', 'cor']
top_corr_df = top_corr_df[(top_corr_df.cor > 0.1) & (top_corr_df.cor < 0.5)]
print(top_corr_df.shape)
top_corr_df.sort_values(by='cor', ascending=False)
# +
label_A = 'mer'#'cheval'#, 'mer'
label_B = 'bateau' #'voiture à attelage'#'bateau'
label_A_ = label_A.replace(' ', '_')
label_B_ = label_B.replace(' ', '_')
# +
from collections import namedtuple
Label_Stats = namedtuple('Label_Stats', ['Probs', 'Joint_prob', 'Cond_probs',
'Variance', 'Odds',
'Covariance', 'Correlation',
'PWOR'])
def calculate_stats (df, label_A, label_B, prnt=False):
p_A = df[label_A].sum() / df.shape[0]
p_B = df[label_B].sum() / df.shape[0]
p_AB = (df[[label_A, label_B]].sum(axis=1) == 2).sum() / df.shape[0]
p_A_given_B = p_AB / p_B
p_B_given_A = p_AB / p_A
var_A = p_A * (1-p_A)
var_B = p_B * (1-p_B)
odds_A = p_A /(1 - p_A)
odds_B = p_B /(1 - p_B)
cov_AB = p_AB - p_A*p_B
cor_AB = cov_AB / ( np.sqrt(var_A) * np.sqrt(var_B) )
pwor_AB = odds_A * ((1 - p_B_given_A) / p_B_given_A)
pwor_BA = odds_B * ((1 - p_A_given_B) / p_A_given_B)
if (prnt):
print('count(%s) = %d' % ( label_A, df[label_A].sum()) )
print('count(%s) = %d' % ( label_B, df[label_B].sum()) )
print()
print('P(%s) = %f' % (label_A, p_A))
print('P(%s) = %f' % (label_B, p_B))
print('P(%s,%s) = %f' % (label_A, label_B, p_AB))
print()
print('P(%s|%s) = %f' % (label_A, label_B, p_A_given_B))
print('P(%s|%s) = %f' % (label_B, label_A, p_B_given_A))
print()
print('Odds(%s) = %f' % (label_A, odds_A))
print('Odds(%s) = %f' % (label_B, odds_B))
print()
print('Var(%s) = %f' % (label_A, var_A))
print('Var(%s) = %f' % (label_B, var_B))
print()
print('Cov(%s,%s) = %f' % (label_A, label_B, cov_AB))
print()
print('Cor(%s,%s) = %f' % (label_A, label_B, cor_AB))
print()
print('PWOR(%s|%s) = %f' % (label_A, label_B, pwor_AB))
print('PWOR(%s|%s) = %f' % (label_B, label_A, pwor_BA))
stats = Label_Stats( (p_A, p_B), p_AB, (p_A_given_B, p_B_given_A), (var_A, var_B), (odds_A, odds_B), cov_AB, cor_AB,
(pwor_AB, pwor_BA))
return stats
stats = calculate_stats(train_annotations_df, label_A_, label_B_, prnt=True)
# +
formula = '%s ~ %s' % (label_A_, label_B_)
res = sm.formula.glm(formula, family=sm.families.Binomial(), data=train_annotations_df).fit()
print(res.summary())
# -
pd.crosstab(train_annotations_df.mer, train_annotations_df.bateau, margins=True)
# ### Extract test set data from the annotations and from the KB
# +
test_annotations_df = annotations_df.loc[annotations_df.usage == 'test']
test_annotations_dummies_df = pd.get_dummies(test_annotations_df.explode('label').label).groupby(level=0).sum()
test_annotations_dummies_df.columns = annotations_dummies_df.columns.str.replace(' ', '_')
test_annotations_df = test_annotations_df.merge(test_annotations_dummies_df,
left_index=True,
right_index=True)
test_annotations_df.shape
# -
stats_test = calculate_stats(test_annotations_df, label_A_, label_B_, prnt=True)
query_image_for_test_set = '''
prefix rdf: <http://www.w3.org/1999/02/22-rdf-syntax-ns#>
PREFIX rdfs: <http://www.w3.org/2000/01/rdf-schema#>
prefix skos: <http://www.w3.org/2004/02/skos/core#>
prefix jcl: <http://jocondelab.iri-research.org/ns/jocondelab/>
#prefix dc: <http://purl.org/dc/elements/1.1/>
prefix ml: <http://ns.inria.fr/monalia/>
select ?ref
?imageURL
?title
#?repr
#?target_labels
#?actual_labels
#?classifier_label
#?score
where
{
VALUES(?ref) {("%s")}
?notice jcl:noticeRef ?ref;
jcl:noticeImage [ jcl:noticeImageIsMain true ; jcl:noticeImageUrl ?imageURL].
optional {?notice jcl:noticeTitr ?title.}
}
''' #% (test_annotations_df.ref[:100].str.cat(sep = ' ' ).replace(' ', '") ("') )
# +
# #%time test_ext_df = metadata.sparql_service_to_dataframe(wds_Joconde_Corese, query_image_for_test_set)
#print(test_ext_df.shape)
# +
chunk_size = 100
ref_chunk_list = [test_annotations_df.ref[i:i+chunk_size] for i in range(0,test_annotations_df.ref.size, chunk_size)]
test_df = pd.DataFrame()
for i, ref_chunk in enumerate(ref_chunk_list):
print(ref_chunk.size * (i+1), end=', ')
chunk_query_str = query_image_for_test_set % (ref_chunk.str.cat(sep = ' ' ).replace(' ', '") ("') )
test_df = pd.concat([test_df, metadata.sparql_service_to_dataframe(wds_Joconde_Corese, chunk_query_str)], ignore_index=True)
print('Done')
# -
test_annotations_df = test_annotations_df.merge(test_df, on='ref')
test_annotations_df.shape
# ### Run query to get image URI and Title
# +
images_root = 'C:/Joconde/joconde'
multi_label = True
model_image_size = 299
dataset_mean = [0.5, 0.5, 0.5]
dataset_std = [0.5, 0.5, 0.5]
batch_size = 4
test_trans = transforms.Compose([
transforms.Resize(model_image_size),
transforms.CenterCrop(model_image_size),
transforms.ToTensor(),
transforms.Normalize(mean = dataset_mean,
std = dataset_std),
])
test_set = JocondeDataset(dataset_description_file,
images_root,
dataset_name = '40_classes',
label_column='label',
multiple_labels = multi_label,
filter_dict= {'usage': ['test']},
add_columns=['ref', 'repr'],
transform=test_trans)
test_loader = torch.utils.data.DataLoader(dataset=test_set,
batch_size=batch_size,
shuffle=False,
num_workers=2)
print('Test', test_set)
print(' Labels:', test_set.labels_count)
print()
# -
# ### Run model to obtain the scores
model_name = 'inception_v3'
model_checkpoint_file = os.path.abspath('../../MonaLIA/output/inception_v3_Joconde_40_classes.1000.no_sched.checkpoint.pth.tar')
checkpoint = torch.load(model_checkpoint_file)
print(checkpoint.keys())
# +
use_cuda = torch.cuda.is_available()
device = torch.device('cuda:0' if use_cuda else 'cpu')
print('Using cuda? ', use_cuda)
net = model.load_net(model_name = model_name, class_count=len(checkpoint['classes']))
#net.load_state_dict(torch.load(model_param_file))
net.load_state_dict(checkpoint['state_dict'])
net = net.to(device)
activation = torch.sigmoid
decision = model.decision_by_class_threshold
decision_param = checkpoint['threshold'].round(2)
scores = model.predict(net, test_loader, activation, decision, decision_param)
# +
index_A = checkpoint['classes'].index(label_A)
index_B = checkpoint['classes'].index(label_B)
scores_A_B = scores[:, [index_A, index_B]].cpu().numpy()
# +
def agg_scores(x):
_x = x#.sort_values(by='classifier_label' , ascending=True, inplace=False)
return pd.Series(dict(#targets = _x.iloc[0].target_labels,
pred_dict = dict(zip(_x['classifier_label'], _x['score'].astype(float)))
#predictions = list(_x['classifier_label']),
#scores = list(_x['score'])
))
def adjust_scores(logit_res, score_A, score_B, base_prob_B = 0.5 ):
score_A = score_A - 0.0000001 # to prevent division by 0 for score = 1.0
odds_pred_A = score_A / (1-score_A)
pwor_A_given_B = np.exp(logit_res.params[1] * (score_B - base_prob_B))
odds_pred_A_new = odds_pred_A * pwor_A_given_B
score_A_new = odds_pred_A_new / (1+ odds_pred_A_new)
return score_A_new
def adjust_scores_with_th(logit_res, score_A, score_B, base_prob_B = 0.5, th=0.5 ):
score_A = score_A - 0.0000001 # to prevent division by 0 for score = 1.0
odds_pred_A = score_A / (1-score_A)
if (score_B > th):
#score_B = score_B - 0.0000001 if score_B > base_prob_B else 0.0
pwor_A_given_B = np.exp(logit_res.params[1] * (score_B - base_prob_B))
odds_pred_A_new = odds_pred_A * pwor_A_given_B
score_A_new = odds_pred_A_new / (1+ odds_pred_A_new)
else:
score_A_new = score_A
return score_A_new
# +
test_df_ = pd.DataFrame.from_dict({'ref' : test_df.ref,
'pred_dict' : [dict(zip( [label_A, label_B] , x )) for x in scores_A_B]})
test_df_['adjusted_score_'+label_A] = test_df_.apply(lambda x: adjust_scores_with_th(res,
x.pred_dict[label_A],
x.pred_dict[label_B],
base_prob_B=0.85,
th=0.5), axis=1 )
test_df_ = test_df_.set_index('ref') \
.merge(test_annotations_df.set_index('ref').loc[:, [label_A_, label_B_]] ,
left_index=True,
right_index=True)
print(test_df_.shape)
test_df_.head()
# +
y_true = np.array(test_df_.loc[:, [label_A_, label_B_]] , dtype = np.dtype('B'))
y_scores = pd.DataFrame.from_dict({label_A: test_df_.pred_dict.apply(lambda x: x[label_A]),
label_B: test_df_.pred_dict.apply(lambda x: x[label_B])}).to_numpy()
AP = metrics.average_precision_score(y_true= y_true,
y_score=y_scores,
average= None)
print('original AP', dict(zip([label_A, label_B], AP)))
y_scores_adj = pd.DataFrame.from_dict({label_A: test_df_['adjusted_score_'+label_A],
label_B: test_df_.pred_dict.apply(lambda x: x[label_B])}).to_numpy()
AP_adj = metrics.average_precision_score(y_true= y_true,
y_score=y_scores_adj,
average= None)
print('adjusted AP', dict(zip([label_A, label_B], AP_adj)))
# +
fig, (ax1, ax2) = plt.subplots(1,2)
fig.set_size_inches((12,6))
precision_A, recall_A, th = metrics.precision_recall_curve(y_true[:, 0],
y_scores[:, 0])
ax1.plot(recall_A, precision_A)
precision_A_adj, recall_A_adj, _ = metrics.precision_recall_curve(y_true[:, 0],
y_scores_adj[:, 0])
ax1.plot(recall_A_adj, precision_A_adj, color='red')
ax1.set_xlim(0.0, 1.05)
ax1.set_ylim(0.0, 1.05)
ax1.set_xlabel('Recall')
ax1.set_ylabel('Precision')
ax1.set_title('Precision-Recall curve: {0} AP={1:0.2f}, adjusted AP={2:0.2f}'.format(label_A, AP[0], AP_adj[0]))
precision_B, recall_B, _ = metrics.precision_recall_curve(y_true[:, 1],
y_scores[:, 1])
ax2.plot(recall_B, precision_B, color='b')
#ax2.fill_between(recall_B, precision_B, alpha=0.2, color='b', step='post')
ax2.set_xlim(0.0, 1.05)
ax2.set_ylim(0.0, 1.05)
ax2.set_xlabel('Recall')
ax2.set_ylabel('Precision')
ax2.set_title(' Precision-Recall curve: {0} AP={1:0.2f}'.format(label_B, AP[1]))
# +
threshold_A = 0.9
threshold_B = 0.85
y_pred = (y_scores > np.array([threshold_A, threshold_B])).astype(dtype = np.dtype('B'))
report = metrics.classification_report(y_true= y_true,
y_pred= y_pred,
target_names = [label_A, label_B])
#print(report)
print()
y_pred_adj = (y_scores_adj > np.array([threshold_A, threshold_B])).astype(dtype = np.dtype('B'))
report_adj = metrics.classification_report(y_true= y_true,
y_pred= y_pred_adj,
target_names = [label_A, label_B])
#print(report_adj)
# +
fig, (ax1, ax2) = plt.subplots(1,2)
fig.set_size_inches((12,8))
cm = metrics.multilabel_confusion_matrix(y_true, y_pred)
metrics.ConfusionMatrixDisplay(cm[0], display_labels=['~' + label_A, label_A]).plot(ax=ax1, cmap='Blues', values_format='d')
ax1.set_title('Before adjustment')
ax1.text(1.5, 3.1, report, horizontalalignment='right', verticalalignment='bottom', fontsize=12)
cm_adj = metrics.multilabel_confusion_matrix(y_true, y_pred_adj)
metrics.ConfusionMatrixDisplay(cm_adj[0], display_labels=['~' + label_A, label_A,]).plot(ax=ax2, cmap='Blues', values_format='d')
ax2.set_title('After adjustment')
ax2.text(1.5, 3.1, report_adj, horizontalalignment='right', verticalalignment='bottom', fontsize=12)
plt.delaxes(fig.axes[2])
plt.delaxes(fig.axes[2])
# +
def gather_row_annotation(row):
return '\n'.join( wrap(row.title, 30 ) +
[''] +
wrap(row.repr, 30 ) +
['',
'<a target="_blank" href="https://www.pop.culture.gouv.fr/notice/joconde/%s">%s</a>' % (row.ref, row.ref),
#'',
# row.imagePath
])
def aggregate_group_by(x): # clean up
_x = x#.sort_values(by='classifier_label' , ascending=True, inplace=False)
return pd.Series(dict(imageURL = _x.iloc[0].imageURL,
info = gather_row_annotation(_x.iloc[0]),
actuals= _x.iloc[0]['actual_labels'].replace('+', '\n'),
targets= _x.iloc[0]['target_labels'].replace('+', '\n'),
#predictions = '\n'.join(_x['classifier_label']),
predictions = '\n'.join(_x.iloc[0]['pred_dict'].keys()),
#scores = '\n'.join(_x['score'])
scores = '\n'.join([str(v.round(4)) for v in _x.iloc[0]['pred_dict'].values()])
))
formatters_dict={'imageURL': vis_helpers.image_url_formatter,
'info': vis_helpers.label_formatter,
'repr': vis_helpers.repr_formatter,
'predictions': vis_helpers.label_formatter,
'classifier_label': vis_helpers.label_formatter,
'scores': vis_helpers.label_formatter,
'actuals': vis_helpers.label_formatter,
'targets': vis_helpers.label_formatter}
# +
test_view_df = test_df.set_index('ref') \
.merge(test_annotations_df.loc[:, ['ref','label', 'terms', 'repr']] , on='ref') \
.merge(test_df_.reset_index(), on='ref' )
test_view_df.rename( columns={'label': 'target_labels',
'terms': 'actual_labels'}, inplace=True)
test_view_df.target_labels = test_view_df.target_labels.apply(lambda x: '+'.join(x))
# +
pd.set_option('display.max_colwidth', None)
pd.set_option('colheader_justify', 'center')
pd.set_option('precision', 4)
test_view_df.fillna('', inplace=True)
print(test_view_df.groupby('ref').size().shape)
print('Sample of 20 images that annotated with either "%s" or "%s"' % (label_A, label_B ) )
HTML(test_view_df.groupby('ref') \
.apply(aggregate_group_by) \
.merge(test_df_.loc[(test_df_[label_A] + test_df_[label_B]) > 0 , ['adjusted_score_'+label_A_]], on='ref')[:20] \
.to_html(
formatters=formatters_dict,
escape=False,
index=False))
# +
test_view_df[label_A_+'_p'] = test_view_df.pred_dict.apply(lambda x: 1 if x[label_A] > threshold_A else 0)
test_view_df[label_A_+'_ap'] = test_view_df['adjusted_score_'+label_A_].apply(lambda x: 1 if x > threshold_A else 0)
print('Changed classification for "%s"' % (label_A) )
test_view_df_flipped = test_view_df.loc[ test_view_df[label_A+'_p'] != test_view_df[label_A_+'_ap'], :]
#print('Classification False Positives for "%s"' % (label_A) )
#test_view_df_flipped = test_view_df_flipped.loc[ (test_view_df_flipped[label_A] == 0) & (test_view_df_flipped[label_A+'_p'] == 1), :]
# +
pd.set_option('display.max_colwidth', None)
pd.set_option('colheader_justify', 'center')
pd.set_option('precision', 4)
#test_view_df_flipped.fillna('', inplace=True)
print(test_view_df_flipped.groupby('ref').size().shape)
HTML(test_view_df_flipped.groupby('ref') \
.apply(aggregate_group_by) \
.reset_index() \
.set_index('ref') \
#.merge(test_df_.loc[:, ['adjusted_score_A']], left_index=True, right_index=True) \
.merge(test_view_df_flipped.loc[:, ['ref', 'adjusted_score_'+label_A, label_A, label_A+'_p', label_A+'_ap']].set_index('ref'), left_index=True, right_index=True) \
.drop_duplicates() \
.sort_values(by=[label_A, label_A+'_p', label_A+'_ap'])
.to_html(
formatters=formatters_dict,
escape=False,
index=False,
show_dimensions=True))
# -
#save to a file
test_view_df_flipped.groupby('ref') \
.apply(aggregate_group_by) \
.reset_index() \
.set_index('ref') \
.merge(test_view_df_flipped.loc[:, ['ref', 'adjusted_score_'+label_A, label_A, label_A+'_p', label_A+'_ap']].set_index('ref'), left_index=True, right_index=True) \
.drop_duplicates() \
.sort_values(by=[label_A, label_A+'_p', label_A+'_ap']) \
.to_html( buf='results of adjusting scores of mer by bateau.html',
formatters=formatters_dict,
escape=False,
index=False,
show_dimensions=True)
# # Scrapbook
test_df_['score_'+ label_A_] = test_df_.pred_dict.apply(lambda x: x[label_A])
test_df_['score_'+ label_B_] = test_df_.pred_dict.apply(lambda x: x[label_B])
# +
import statsmodels.api as sm
res = sm.formula.glm("mer ~ score_mer", family=sm.families.Binomial(),
data=test_df_).fit()
print(res.summary())
# -
print("Coefficeients")
print(res.params)
print()
print("p-Values")
print(res.pvalues)
print()
print("Dependent variables")
print(res.model.endog_names)
x = pd.DataFrame.from_dict({'bateau': [0.0, 0.1, 0.2,0.3, 0.4, 0.5, 0.6, 0.7, 0.8, 0.9, 1.0]})
res_pred = pd.Series(res.predict(x, transform=True)) #.apply(lambda s: 1 / (1+ np.exp(-s)))
plt.scatter(x = x,
y = res_pred)
| Notebooks 3.0/Pair-Wise Odds Ratio Study/MonaLIA PWOR Study-Train set-model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import glob
import pandas as pd
path = '/network/group/aopp/predict/TIP016_PAXTON_RPSPEEDY/ML4L/ECMWF_files/raw/joined_ML_data_w_skt/'
files = sorted(glob.glob(path+'*.pkl'))
f=files[0]
df = pd.read_pickle(f)
df.columns
files[-1]
files[-1]
| legacy/legacy_notebooks/.ipynb_checkpoints/03.Merge_hourly_files-checkpoint.ipynb |